text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
# **Build Linear Regression Model in Python**
Chanin Nantasenamat
[*'Data Professor' YouTube channel*](http://youtube.com/dataprofessor)
In this Jupyter notebook, I will be showing you how to build a linear regression model in Python using the scikit-learn package.
Inspired by [scikit-learn's Linear Regression Example](https://scikit-learn.org/stable/auto_examples/linear_model/plot_ols.html)
---
## **Load the Diabetes dataset** (via scikit-learn)
### **Import library**
```
from sklearn import datasets
```
### **Load dataset**
```
diabetes = datasets.load_diabetes()
diabetes
```
### **Description of the Diabetes dataset**
```
print(diabetes.DESCR)
```
### **Feature names**
```
print(diabetes.feature_names)
```
### **Create X and Y data matrices**
```
X = diabetes.data
Y = diabetes.target
X.shape, Y.shape
```
### **Load dataset + Create X and Y data matrices (in 1 step)**
```
X, Y = datasets.load_diabetes(return_X_y=True)
X.shape, Y.shape
```
## **Load the Boston Housing dataset (via GitHub)**
The Boston Housing dataset was obtained from the mlbench R package, which was loaded using the following commands:
```
library(mlbench)
data(BostonHousing)
```
For your convenience, I have also shared the [Boston Housing dataset](https://github.com/dataprofessor/data/blob/master/BostonHousing.csv) on the Data Professor GitHub package.
### **Import library**
```
import pandas as pd
```
### **Download CSV from GitHub**
```
! wget https://github.com/dataprofessor/data/raw/master/BostonHousing.csv
```
### **Read in CSV file**
```
BostonHousing = pd.read_csv("BostonHousing.csv")
BostonHousing
```
### **Split dataset to X and Y variables**
```
Y = BostonHousing.medv
Y
X = BostonHousing.drop(['medv'], axis=1)
X
```
## **Data split**
### **Import library**
```
from sklearn.model_selection import train_test_split
```
### **Perform 80/20 Data split**
80% of the data is used to train the model. 20% to test it
```
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=0.2)
```
### **Data dimension**
```
X_train.shape, Y_train.shape
X_test.shape, Y_test.shape
```
## **Linear Regression Model**
### **Import library**
```
from sklearn import linear_model
from sklearn.metrics import mean_squared_error, r2_score
```
### **Build linear regression**
#### Defines the regression model
```
model = linear_model.LinearRegression()
```
#### Build training model
```
model.fit(X_train, Y_train)
```
#### Apply trained model to make prediction (on test set)
```
Y_pred = model.predict(X_test)
```
## **Prediction results**
### **Print model performance**
```
print('Coefficients:', model.coef_)
print('Intercept:', model.intercept_)
print('Mean squared error (MSE): %.2f'
% mean_squared_error(Y_test, Y_pred))
print('Coefficient of determination (R^2): %.2f'
% r2_score(Y_test, Y_pred))
```
The coefficients represent the weight values of each of the features
### **String formatting**
By default r2_score returns a floating number ([more details](https://docs.scipy.org/doc/numpy-1.13.0/user/basics.types.html))
```
r2_score(Y_test, Y_pred)
r2_score(Y_test, Y_pred).dtype
```
We will be using the modulo operator to format the numbers by rounding it off.
```
'%f' % 0.523810833536016
```
We will now round it off to 3 digits
```
'%.3f' % 0.523810833536016
```
We will now round it off to 2 digits
```
'%.2f' % 0.523810833536016
```
## **Scatter plots**
### **Import library**
```
import seaborn as sns
```
### **Make scatter plot**
#### The Data
```
Y_test
import numpy as np
np.array(Y_test)
Y_pred
```
#### Making the scatter plot
```
sns.scatterplot(Y_test, Y_pred)
sns.scatterplot(Y_test, Y_pred, marker="+")
sns.scatterplot(Y_test, Y_pred, alpha=0.5)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/cltl/python-for-text-analysis/blob/colab/Chapters-colab/Chapter_13_Working_with_Python_files.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
%%capture
!wget https://github.com/cltl/python-for-text-analysis/raw/master/zips/Data.zip
!wget https://github.com/cltl/python-for-text-analysis/raw/master/zips/images.zip
!wget https://github.com/cltl/python-for-text-analysis/raw/master/zips/Extra_Material.zip
!unzip Data.zip -d ../
!unzip images.zip -d ./
!unzip Extra_Material.zip -d ../
!rm Data.zip
!rm Extra_Material.zip
!rm images.zip
```
# Chapter 13 - Working with Python files
In the previous blocks, we've mainly used notebooks to develop and run our Python code. In this chapter, we'll introduce how to create Python modules (.py files) and how to run them. The most common way to work with Python is actually to use .py files, which is why it is important that you know how to work with them. You can see python files as one cell in a notebook without any markdown.
Before we write actual code in a .py file, we will explain the basics you need to know for doing this:
* Choosing an editor
* starting the terminal (from which you will run your .py files)
**At the end of this chapter, you will be able to**
* create python modules, i.e., .py files
* run python modules from the command line
If you have **questions** about this chapter, please contact us **(cltl.python.course@gmail.com)**.
# 1. Editor
We first need to choose which editor we will use to develop our Python code.
There are two options.
1. You create the python modules in your browser. After opening Jupyter notebook, you can click `File` -> `New` and then `Text file` to start developing Python modules.
2. You install an editor.
Please take a look [here](https://wiki.python.org/moin/PythonEditors) to get an impression of which ones are out there.
We can highly recommend [Atom](https://atom.io/) (for macOS, Windows, Linux). Other options are [BBEdit](https://www.barebones.com/products/bbedit/download.html) (for macOS) and [Notepad++](https://notepad-plus-plus.org/) (for Windows). A simple way to create a new .py file usually is to open a new file and save it as name_of_your_program.py (make sure to use indicative names).
Please choose between options 1 and 2.
# 2. Starting the terminal
To run a .py file we wrote in an editor, we need to start the terminal. This works differently for windows and Mac:
1. On Windows, please look at [Anaconda Prompt](https://docs.conda.io/projects/conda/en/latest/user-guide/getting-started.html)
2. on OS X/macOS (Mac computer), please type **terminal** in [Spotlight](https://support.apple.com/nl-nl/HT204014) and start the terminal
It's a useful skill to know how to navigate through your computer (i.e., go from one directory to another, all the files and subdirectories in a directory, etc.) using the terminal.
For Windows users, [this](https://www.computerhope.com/issues/chusedos.htm) is a good tutorial.
For OS X/macOS/Linux/Ubuntu users, [this](https://www.digitalocean.com/community/tutorials/basic-linux-navigation-and-file-management) is a good tutorial.
# 3. Running your first program
Here, we'll show you how to run your first program (hello_world.py).
In the same folder as this notebook, you will find a file called **hello_world.py**.
Running it works differently on Windows and Mac. Below, instructions for both can be found:
## A.) Running the program on OS X/MacOS
Please use the terminal to navigate to the folder in which this notebook is placed by copying the **output** of following cell in your terminal
```
import os
cwd = os.getcwd()
cwd_escaped_spaces = cwd.replace(' ', '\ ')
print('cd', cwd_escaped_spaces)
```
`cd` means 'change directory'. Here, you are using it to go to the directory we are currently working in. We use the os module to print the path to this directory (`os.getcwd`).
Please run the following command in the terminal:
**python hello_world.py**
You've succesfully run your first Python program!
## B.) Running the program on Windows
Please use the terminal to navigate to the folder in which this notebook is placed by copying the **output** of the following cell in your terminal
```
cwd = os.getcwd()
cwd_escaped_spaces = cwd.replace(' ', '^ ')
print('cd', cwd_escaped_spaces)
```
Please run the **output** of the following command in the terminal:
```
import sys
print(sys.executable + ' hello_world.py')
```
You've succesfully run your first Python program!
# 4. Import your own functions
In Chapter 12, you've been introduced to **importing** modules and functions/methods.
You can see any python program that you create (so any .py file) as a module, which means that you can import it into another python program. Let's see how this works.
Please note that the following examples only work if all your python files are in the same directory. There are ways of importing python modules from other directories, but we will not discuss them here.
## 4.1 Importing your entire module
When importing your own functions from your own modules, several things are important. We have created two example scripts to illustrate them called **the_program.py** and **utils.py**. We recommend to open them to check the following things:
* The extension .py is not used when importing modules. **import utils** will import the file **utils.py** [line 1 the_program.py]
* We can use any function from the file. We can call the count_words function by typing **utils.count_words** [the_program.py line 6]
* We can use any global variable declared in the imported module. E.g. **utils.x** and **utils.python** declared in utils.py can be used in the_program.py
## 4.2 Importing functions and variables individually
We can import specific functions using the syntax **from MODULE import FUNCTION/VARIABLE**
This can be seen in the file **the_program_v2.py** (lines 1-3). (Open the files **the_program_v2.py** and **utils.py** in an editor to check this).
## 4.3 Importing functions and variables to python notebooks
Please note that you can also import functions and variables from a python program while using notebooks. In this case, simply treat the notebook as the python files the_program.py and the_program_v2.py.
```
from utils import count_words
words = ['how', 'often', 'does', 'each', 'string', 'occur', 'in', 'this', 'list', '?']
word2freq = count_words(words)
print('word2freq', word2freq)
```
# Exercises
**Exercise 1**:
Please create and run your own program using an editor and the terminal. Please copy your beersong into your first program. Tip: simply open a new file in the editor and save it as `beersong.py'.
**Exercise 2**:
Please create two files:
* **my_second_program.py**
* **my_utils.py**
Please create a helper function and store it in **my_utils.py**, import it into **my_second_program.py** and call it from there.
```
```
| github_jupyter |
## Exercise 4: Tensors and Tensor Factorization Techniques
This exercise is an introduction to latent representation and tensors. First we will understand latent representation using SVD. Later we will extend this idea to tensors.
## Exercise-4.1-a
# Latent representation using SVD
Suppose we have to design a movie recommendation system. We are given some data of viewers and the movie they have seen in past. Now using this information we have to design a learning model which can recommend new movies depending on the pattern of data.
Data is structured as a matrix where rows represent viewers and column represents movies. The entry is 1 if viewer has seen the movie. Missing values are what we want to predict.
In this part we will study following:
(a) SVD as a matrix factorization.
(b) Singular Vectors as a latent representation.
(c) Find latent representation cutoff using knee plot.
(b) Predict Missing values and use it as a recommendation.
$$
Data = \left( \begin{array}{ccccccccccc}
& Movie-1 & Movie-2 & Movie-3 & Movie-4 & Movie-5 & Movie-6 & Movie-7 & Movie-8 & Movie-9 & Movie-10 \\
Viewer-1 & \_ & \_ & 1 & 1 & 1 & \_ & \_ & \_ & \_ & \_\\
Viewer-2 & 1 & \_ & 1 & 1 & \_ & 1 & \_ & \_ & 1 & \_\\
Viewer-3 & \_ & \_ & \_ & \_ & \_ & \_ & 1 & 1 & \_ & 1\\
Viewer-4 & 1 & 1 & 1 & \_ & 1 & \_ & \_ & \_ & \_ & \_\\
Viewer-5 & 1 & 1 & 1 & 1 & 1 & \_ & \_ & \_ & \_ & \_\\
Viewer-6 & \_ & \_ & 1 & 1 & 1 & \_ & \_ & \_ & 1 & \_ \\
Viewer-7 & 1 & 1 & 1 & 1 & \_ & \_ & \_ & \_ & \_ & \_ \\
Viewer-8 & \_ & \_ & \_ & 1 & \_ & 1 & \_ & 1 & 1 & 1\\
\end{array}
\right)
$$
```
import numpy as np
from scipy.linalg import svd,diagsvd
import matplotlib.pyplot as plt
#generate the data
X = np.array([[0,0,1,1,1,0,0,0,0,0],
[1,0,1,1,0,1,0,0,1,0],
[0,0,0,0,0,0,1,1,0,1],
[1,1,1,0,1,0,0,0,0,0],
[1,1,1,1,1,0,0,0,0,0],
[0,0,1,1,1,0,0,0,1,0],
[1,1,1,1,0,0,0,0,0,0],
[0,0,0,1,0,1,0,1,1,1]])
#SVD Calculatiom
U, s, Vh = svd(X)
print(np.shape(U))
print(np.shape(Vh))
print(np.shape(s))
#Elbow PLot: To decide significant singular values
plt.plot(s**2)
plt.xlabel('N')
plt.ylabel('$Singular Values^2$')
plt.title('Elbow plot for SVD')
plt.show()
plt.scatter(U[:,0],U[:,1])
for i,name in enumerate(['Viewer-1','Viewer-2','Viewer-3','Viewer-4','Viewer-5','Viewer-6','Viewer-7','Viewer-8']):
plt.annotate(name,(U[i,0],U[i,1]))
plt.xlabel('Singular Vector-1')
plt.ylabel('Singular Vector-2')
plt.title('Latent Representation')
plt.show()
#truncate to only first few singular values
s[4:]=0
#calculate estimate using truncated S
out_score = np.dot(np.dot(U,diagsvd(s,8,10)),Vh)
#show results
print("X=\n.{}".format(X))
print("U=\n.{}".format((U[:1,:].round(1))))
print("V=\n.{}".format(Vh[:,:1].round(1)))
print("Score=\n.{}".format(out_score))
print("Reconstructed-X=\n.{}".format(np.round(out_score)))
```
## Exercise-4.1-b
In previous part we used binary matrix for modeling viewer-movie recoomendation. Now suppose we are given a new data where each entry is a rating on a scale 1 to 10. Use SVD to learn latent representation and recommend 3 movies for viewer 8.
A sample data.
$$
Data = \left( \begin{array}{ccccccccccc}
& Movie-1 & Movie-2 & Movie-3 & Movie-4 & Movie-5 & Movie-6 & Movie-7 & Movie-8 & Movie-9 & Movie-10 \\
Viewer-1 & \_ & \_ & 5 & 4 & 6 & \_ & \_ & \_ & \_ & \_\\
Viewer-2 & 8 & \_ & 2 & 3 & \_ & 6 & \_ & \_ & 7 & \_\\
Viewer-3 & \_ & \_ & \_ & \_ & \_ & \_ & 4 & 8 & \_ & 1\\
Viewer-4 & 2 & 5 & 9 & \_ & 9 & \_ & \_ & \_ & \_ & \_\\
Viewer-5 & 3 & 3 & 3 & 7 & 8 & \_ & \_ & \_ & \_ & \_\\
Viewer-6 & \_ & \_ & 4 & 3 & 8 & \_ & \_ & \_ & 6 & \_ \\
Viewer-7 & 2 & 5 & 7 & 6 & \_ & \_ & \_ & \_ & \_ & \_ \\
Viewer-8 & \_ & \_ & \_ & 5 & \_ & 1 & \_ & 8 & 3 & 7\\
\end{array}
\right)
$$
```
########################################
####### Your Code Here ################
########################################
```
## Exercise-4.2
# Introduction To Tensor
This part is an introduction to tensor. We will do the following task:
(a) Storing tensor using numpy array.
(b) Slicing operations on tensors.
(c) Folding and Unfolding tensors.
```
data = np.arange(36).reshape((3,4,3))
print('Data= \n{}'.format(data))
tensor = np.array([data.T[i].T for i in range(len(data))])
print('Tensor= \n{}'.format(tensor))
# To view frontal slide-1
print(data[:,:,0])
# To view frontal slide-2
print(data[:,:,1])
# To view frontal slide-3
print(data[:,:,2])
```
## Unfolding and folding Tensors
```
from IPython.display import Image
Image(filename='Unfolding-of-third-order-of-a-tensor.png')
def unfold(X, mode):
return np.reshape(np.moveaxis(X, mode-1,0),(X.shape[mode-1],-1))
def fold(X, mode, shape):
new_shape = list(shape)
mode_dim = new_shape.pop(mode-1)
new_shape.insert(0, mode_dim)
return np.moveaxis(np.reshape(X, new_shape), 0, mode-1)
unfold(data,mode=1)
unfold(data,mode=2)
unfold(data,mode=3)
unfold_tensor = unfold(data,mode=0)
fold(unfold_tensor, mode=0, shape=data.shape)
```
## Tensor Decomposition
Now we will extend idea of matrix factorization to tensors. In following part we will learn two methods for tensor factorization: CP decomposition and RESCAL. Further we will see how to use them with RDF dataset for link prediction.
For purpose of experiments we will use kinship(alyawarra) dataset.
The alyawarra dataset used in this exercise has 26 relations (brother, sister, father,...} between 104 people. Using tensor factorization we will predict missing relations and evaluate the performance using area-under-curve.
## Exercise-4.3
$\textbf{CP Decomposition}$ is a generalization of the matrix SVD to tensors.
The CP Decomposition factorizes a tensor into a sum of outer products of vectors. For a 3-way tensor CP decomposition is written as:
$ \mathbf{T = \sum_r^R \lambda_r {a_r}^1 \odot {a_r}^2 \odot {a_r}^3} + \epsilon$
where $\odot$ denotes outer product for tensors, Y_f is the approximated tensor and \epsilon is an error.
To approximate above factorization we will define a loss minimizing Frobenius Norm:
$L = argmin_{{a_r}^1,{a_r}^2,{a_r}^3} ||T-\sum_r^R \lambda_r {a_r}^1 \odot {a_r}^2 \odot {a_r}^3||^2 $
To solve the above function we will use alternating least square method similar to least square method used in last exercise.(Proof as an exercise!)
```
from IPython.display import Image
Image(filename='CP.png')
import pandas as pd
import pdb
from sktensor import dtensor, cp_als
from scipy.io.matlab import loadmat
import matplotlib.pyplot as plt
import itertools
mat = loadmat('alyawarradata.mat')
T = mat['Rs']
T = dtensor(T)
trainT = np.zeros_like(T)
p = 0.7
train_mask = np.random.binomial(1, p, T.shape)
trainT[train_mask==1] = T[train_mask==1]
test_mask = np.ones_like(T)
test_mask[train_mask==1] = 0
print('training size %d' % np.sum(trainT))
print('test size %d' % np.sum(T[test_mask==1]))
# Decompose tensor using CP-ALS
P, fit, itr, exectimes = cp_als(trainT, 3, init='random')
reconstructed_tensor = P.totensor()
from sklearn.metrics import roc_auc_score
print(roc_auc_score(T[test_mask==1], reconstructed_tensor[test_mask==1]))
```
## CP Entity Embedding Visualization
```
subject_emb = P.U[0]
plt.scatter(subject_emb[:,0],subject_emb[:,1])
plt.xlabel('Latent Value-1')
plt.ylabel('Latent Value-2')
plt.title('Latent Representation Subject')
plt.show()
object_emb = P.U[1]
plt.scatter(object_emb[:,0],object_emb[:,1])
plt.xlabel('Latent Value-1')
plt.ylabel('Latent Value-2')
plt.title('Latent Representation Predicate')
plt.show()
```
## Exercise-4.4
## Rescal Decomposition
RESCAL factorization corresponds to Tucker2 decomposition with the constraint that two factor matrices have to be identical
$ \mathbf{T = R \times_1 A \times_2 A} + \epsilon$
$ \mathbf{T_{:,:,k} = A R_{:,:,k} A^T} + \epsilon$
where $\textbf{A}\in \mathbb{R}^{|V|\times r} $ represents the entity-latent-component space. $\textbf{R}_{:,:,k}\in \mathbb{R}^{r\times r} $ is an asymmetric matrix that specifies the interaction of the latent components for the k-th relation. Y_f is the approximated tensor and \epsilon is a noise estimate.
To approximate above factorization we will define a loss minimizing Frobenius Norm:
$L = argmin_{A,R} \sum_k||T-A R_{:,:,k} A^T||^2 $
To solve the above loss function we will again use alternating least square method. (Proof as an exercise!)
```
from IPython.display import Image
Image(filename='rescal.png')
from numpy.linalg import norm
from numpy.random import shuffle
from scipy.sparse import lil_matrix
from sklearn.metrics import precision_recall_curve, auc
from rescal import rescal_als
def normalize_predictions(P, nm_entities, nm_relations):
for a in range(nm_entities):
for b in range(nm_entities):
nrm = norm(P[a, b, :nm_relations])
if nrm != 0:
# round values for faster computation of AUC-PR
P[a, b, :nm_relations] = np.round_(P[a, b, :nm_relations] / nrm, decimals=3)
return P
def rescal_fact(train_tensor, n_dim, nm_entities, nm_relations):
entity_embedding, R, _, _, _ = rescal_als(train_tensor, n_dim, init='nvecs', conv=1e-3,lambda_A=10, lambda_R=10)
n = entity_embedding.shape[0]
reconstructed_tensor = np.zeros((n, n, len(R)))
for k in range(len(R)):
reconstructed_tensor[:, :, k] = np.dot(entity_embedding, np.dot(R[k], entity_embedding.T))
reconstructed_tensor = normalize_predictions(reconstructed_tensor, nm_entities, nm_relations)
return entity_embedding, reconstructed_tensor
def load_data(filename, train_fraction=0.7):
mat = loadmat(filename)
K = np.array(mat['Rs'], np.float32)
nm_entities, nm_relations = K.shape[0], K.shape[2]
# construct array for rescal
T = [lil_matrix(K[:, :, i]) for i in range(nm_relations)]
# Train Test Split
triples = nm_entities * nm_entities * nm_relations
IDX = list(range(triples))
shuffle(IDX)
train = int(train_fraction*len(IDX))
idx_test = IDX[train:]
train_tensor = [Ti.copy() for Ti in T]
mask_idx = np.unravel_index(idx_test, (nm_entities, nm_entities, nm_relations))
# set values to be predicted to zero
for i in range(len(mask_idx[0])):
train_tensor[mask_idx[2][i]][mask_idx[0][i], mask_idx[1][i]] = 0
return K, train_tensor, mask_idx, nm_entities, nm_relations
n_dim = 100
filename='alyawarradata.mat'
K, train_tensor, target_idx, nm_entities, nm_relations = load_data(filename, train_fraction=0.7)
# Train Rescal
entity_embedding, reconstructed_tensor = rescal_fact(train_tensor, n_dim, nm_entities, nm_relations)
#prec, recall, _ = precision_recall_curve(K[target_idx], reconstructed_tensor[target_idx])
#entities = mat['names']
print('AUC\n{}'.format(roc_auc_score(K[target_idx], reconstructed_tensor[target_idx])))
```
## Rescal Entity Embedding Visualization
```
plt.scatter(entity_embedding[:,0],entity_embedding[:,1])
plt.xlabel('Latent Value-1')
plt.ylabel('Latent Value-2')
plt.title('Latent Representation')
plt.show()
```
## Exercise-4.5
In last part we used RESCAL to find latent representation of entities in knowledge graph. For RESCAL we need to specify certain parameters. In this exercise we will tune these parameters for optimum performance. We will divide dataset in three parts: training, validation and test. First we find latent representation using training set then we use latent features to compute performance on validation set. Finally we select parameters with optimum performance on validation set and report performance on test set.
```
def load_train_val_test(filename, train_fraction=0.6, val_fraction=0.2):
mat = loadmat(filename)
K = np.array(mat['Rs'], np.float32)
nm_entities, nm_relations = K.shape[0], K.shape[2]
# construct array for rescal
T = [lil_matrix(K[:, :, i]) for i in range(nm_relations)]
# Train Test Split
triples = nm_entities * nm_entities * nm_relations
IDX = list(range(triples))
shuffle(IDX)
train = int(train_fraction*len(IDX))
val = int(val_fraction*len(IDX))
idx_val = IDX[train:train+val]
idx_test = IDX[train+val:]
train_tensor = [Ti.copy() for Ti in T]
mask_idx = np.unravel_index(idx_test+idx_val, (nm_entities, nm_entities, nm_relations))
val_idx = np.unravel_index(idx_val, (nm_entities, nm_entities, nm_relations))
test_idx = np.unravel_index(idx_test, (nm_entities, nm_entities, nm_relations))
# set values to be predicted to zero
for i in range(len(mask_idx[0])):
train_tensor[mask_idx[2][i]][mask_idx[0][i], mask_idx[1][i]] = 0
return K, train_tensor, val_idx, test_idx
n_dim = 10
filename='alyawarradata.mat'
# Mask Test and Validation Triples
K, train_tensor, val_idx, test_idx = load_train_val_test(filename, train_fraction=0.6, val_fraction=0.2)
var_list = [0.001, 0.1, 1., 10., 100.]
best_roc = 0
# Train on training set and evaluate on validation set
for (var_x, var_e, var_r) in itertools.product(var_list, repeat=3):
A, R, f, itr, exectimes = rescal_als(train_tensor, n_dim, lambda_A=var_x, lambda_R=var_e, lambda_V=var_r)
n = A.shape[0]
reconstructed_tensor = np.zeros((n, n, len(R)))
for k in range(len(R)):
reconstructed_tensor[:, :, k] = np.dot(A, np.dot(R[k], A.T))
reconstructed_tensor = normalize_predictions(reconstructed_tensor, nm_entities, nm_relations)
score = roc_auc_score(K[val_idx], reconstructed_tensor[val_idx])
print('var_x:{0:3.3f}, var_e:{1:3.3f}, var_r:{2:3.3f}, AUC-ROC:{3:.3f}'.format(var_x, var_e, var_r, score))
if score > best_roc:
best_vars = (var_x, var_e, var_r)
best_roc = score
lambda_a, lambda_r, lambda_v = best_vars
print(best_vars, best_roc)
# Use optimum parameters on Validation Set and Test on Hold-Out Set
lambda_a, lambda_r, lambda_v = best_vars
A, R, f, itr, exectimes = rescal_als(train_tensor, n_dim, lambda_A=var_x, lambda_R=var_e, lambda_V=var_r)
n = A.shape[0]
reconstructed_tensor = np.zeros((n, n, len(R)))
for k in range(len(R)):
reconstructed_tensor[:, :, k] = np.dot(A, np.dot(R[k], A.T))
reconstructed_tensor = normalize_predictions(reconstructed_tensor, nm_entities, nm_relations)
score = roc_auc_score(K[test_idx], reconstructed_tensor[test_idx])
print('AUC On Test Set with Optimum Parameters\n{}'.format(score))
```
## References
1) https://github.com/mnick/scikit-tensor
2) https://github.com/mnick/rescal.py
3) http://www.bsp.brain.riken.jp/~zhougx/tensor.html
4) https://edoc.ub.uni-muenchen.de/16056/1/Nickel_Maximilian.pdf
5) http://epubs.siam.org/doi/abs/10.1137/S0895479896305696
| github_jupyter |
# Random search and hyperparameter scaling with SageMaker XGBoost and Automatic Model Tuning
---
## Contents
1. [Introduction](#Introduction)
1. [Preparation](#Preparation)
1. [Download and prepare the data](#Download-and-prepare-the-data)
1. [Setup hyperparameter tuning](#Setup-hyperparameter-tuning)
1. [Logarithmic scaling](#Logarithmic-scaling)
1. [Random search](#Random-search)
1. [Linear scaling](#Linear-scaling)
---
## Introduction
This notebook showcases the use of two hyperparameter tuning features: **random search** and **hyperparameter scaling**.
We will use SageMaker Python SDK, a high level SDK, to simplify the way we interact with SageMaker Hyperparameter Tuning.
---
## Preparation
Let's start by specifying:
- The S3 bucket and prefix that you want to use for training and model data. This should be within the same region as SageMaker training.
- The IAM role used to give training access to your data. See SageMaker documentation for how to create these.
```
import sagemaker
import boto3
from sagemaker.tuner import (
IntegerParameter,
CategoricalParameter,
ContinuousParameter,
HyperparameterTuner,
)
import numpy as np # For matrix operations and numerical processing
import pandas as pd # For munging tabular data
import os
region = boto3.Session().region_name
smclient = boto3.Session().client("sagemaker")
role = sagemaker.get_execution_role()
bucket = sagemaker.Session().default_bucket()
prefix = "sagemaker/DEMO-hpo-xgboost-dm"
```
---
## Download and prepare the data
Here we download the [direct marketing dataset](https://archive.ics.uci.edu/ml/datasets/bank+marketing) from UCI's ML Repository.
```
!wget -N https://archive.ics.uci.edu/ml/machine-learning-databases/00222/bank-additional.zip
!unzip -o bank-additional.zip
```
Now let us load the data, apply some preprocessing, and upload the processed data to s3
```
# Load data
data = pd.read_csv("./bank-additional/bank-additional-full.csv", sep=";")
pd.set_option("display.max_columns", 500) # Make sure we can see all of the columns
pd.set_option("display.max_rows", 50) # Keep the output on one page
# Apply some feature processing
data["no_previous_contact"] = np.where(
data["pdays"] == 999, 1, 0
) # Indicator variable to capture when pdays takes a value of 999
data["not_working"] = np.where(
np.in1d(data["job"], ["student", "retired", "unemployed"]), 1, 0
) # Indicator for individuals not actively employed
model_data = pd.get_dummies(data) # Convert categorical variables to sets of indicators
# columns that should not be included in the input
model_data = model_data.drop(
["duration", "emp.var.rate", "cons.price.idx", "cons.conf.idx", "euribor3m", "nr.employed"],
axis=1,
)
# split data
train_data, validation_data, test_data = np.split(
model_data.sample(frac=1, random_state=1729),
[int(0.7 * len(model_data)), int(0.9 * len(model_data))],
)
# save preprocessed file to s3
pd.concat([train_data["y_yes"], train_data.drop(["y_no", "y_yes"], axis=1)], axis=1).to_csv(
"train.csv", index=False, header=False
)
pd.concat(
[validation_data["y_yes"], validation_data.drop(["y_no", "y_yes"], axis=1)], axis=1
).to_csv("validation.csv", index=False, header=False)
pd.concat([test_data["y_yes"], test_data.drop(["y_no", "y_yes"], axis=1)], axis=1).to_csv(
"test.csv", index=False, header=False
)
boto3.Session().resource("s3").Bucket(bucket).Object(
os.path.join(prefix, "train/train.csv")
).upload_file("train.csv")
boto3.Session().resource("s3").Bucket(bucket).Object(
os.path.join(prefix, "validation/validation.csv")
).upload_file("validation.csv")
s3_input_train = sagemaker.s3_input(
s3_data="s3://{}/{}/train".format(bucket, prefix), content_type="csv"
)
s3_input_validation = sagemaker.s3_input(
s3_data="s3://{}/{}/validation/".format(bucket, prefix), content_type="csv"
)
```
---
## Setup hyperparameter tuning
In this example, we are using SageMaker Python SDK to set up and manage the hyperparameter tuning job. We first configure the training jobs the hyperparameter tuning job will launch by initiating an estimator, and define the static hyperparameter and objective
```
from sagemaker.amazon.amazon_estimator import get_image_uri
sess = sagemaker.Session()
container = get_image_uri(region, "xgboost", repo_version="latest")
xgb = sagemaker.estimator.Estimator(
container,
role,
train_instance_count=1,
train_instance_type="ml.m4.xlarge",
output_path="s3://{}/{}/output".format(bucket, prefix),
sagemaker_session=sess,
)
xgb.set_hyperparameters(
eval_metric="auc",
objective="binary:logistic",
num_round=100,
rate_drop=0.3,
tweedie_variance_power=1.4,
)
objective_metric_name = "validation:auc"
```
# Logarithmic scaling
In both cases we use logarithmic scaling, which is the scaling type that should be used whenever the order of magnitude is more important that the absolute value. It should be used if a change, say, from 1 to 2 is expected to have a much bigger impact than a change from 100 to 101, due to the fact that the hyperparameter doubles in the first case but not in the latter.
```
hyperparameter_ranges = {
"alpha": ContinuousParameter(0.01, 10, scaling_type="Logarithmic"),
"lambda": ContinuousParameter(0.01, 10, scaling_type="Logarithmic"),
}
```
# Random search
We now start a tuning job using random search. The main advantage of using random search is that this allows us to train jobs with a high level of parallelism
```
tuner_log = HyperparameterTuner(
xgb,
objective_metric_name,
hyperparameter_ranges,
max_jobs=20,
max_parallel_jobs=10,
strategy="Random",
)
tuner_log.fit(
{"train": s3_input_train, "validation": s3_input_validation}, include_cls_metadata=False
)
```
Let's just run a quick check of the hyperparameter tuning jobs status to make sure it started successfully.
```
boto3.client("sagemaker").describe_hyper_parameter_tuning_job(
HyperParameterTuningJobName=tuner_log.latest_tuning_job.job_name
)["HyperParameterTuningJobStatus"]
```
# Linear scaling
Let us compare the results with executing a job using linear scaling.
```
hyperparameter_ranges_linear = {
"alpha": ContinuousParameter(0.01, 10, scaling_type="Linear"),
"lambda": ContinuousParameter(0.01, 10, scaling_type="Linear"),
}
tuner_linear = HyperparameterTuner(
xgb,
objective_metric_name,
hyperparameter_ranges_linear,
max_jobs=20,
max_parallel_jobs=10,
strategy="Random",
)
# custom job name to avoid a duplicate name
job_name = tuner_log.latest_tuning_job.job_name + "linear"
tuner_linear.fit(
{"train": s3_input_train, "validation": s3_input_validation},
include_cls_metadata=False,
job_name=job_name,
)
```
Check of the hyperparameter tuning jobs status
```
boto3.client("sagemaker").describe_hyper_parameter_tuning_job(
HyperParameterTuningJobName=tuner_linear.latest_tuning_job.job_name
)["HyperParameterTuningJobStatus"]
```
## Analyze tuning job results - after tuning job is completed
**Once the tuning jobs have completed**, we can compare the distribution of the hyperparameter configurations chosen in the two cases.
Please refer to "HPO_Analyze_TuningJob_Results.ipynb" to see more example code to analyze the tuning job results.
```
import seaborn as sns
import pandas as pd
import matplotlib.pyplot as plt
# check jobs have finished
status_log = boto3.client("sagemaker").describe_hyper_parameter_tuning_job(
HyperParameterTuningJobName=tuner_log.latest_tuning_job.job_name
)["HyperParameterTuningJobStatus"]
status_linear = boto3.client("sagemaker").describe_hyper_parameter_tuning_job(
HyperParameterTuningJobName=tuner_linear.latest_tuning_job.job_name
)["HyperParameterTuningJobStatus"]
assert status_log == "Completed", "First must be completed, was {}".format(status_log)
assert status_linear == "Completed", "Second must be completed, was {}".format(status_linear)
df_log = sagemaker.HyperparameterTuningJobAnalytics(
tuner_log.latest_tuning_job.job_name
).dataframe()
df_linear = sagemaker.HyperparameterTuningJobAnalytics(
tuner_linear.latest_tuning_job.job_name
).dataframe()
df_log["scaling"] = "log"
df_linear["scaling"] = "linear"
df = pd.concat([df_log, df_linear], ignore_index=True)
g = sns.FacetGrid(df, col="scaling", palette="viridis")
g = g.map(plt.scatter, "alpha", "lambda", alpha=0.6)
```
## Deploy the best model
Now that we have got the best model, we can deploy it to an endpoint. Please refer to other SageMaker sample notebooks or SageMaker documentation to see how to deploy a model.
| github_jupyter |
# [Predict Future Sales - Kaggle](https://www.kaggle.com/c/competitive-data-science-predict-future-sales)
* [How to Win a Data Science Competition: Learn from Top Kagglers - Coursera](https://www.coursera.org/learn/competitive-data-science)
* 위의 코세라 코스에 있는 데이터 사이언스 경진대회에서 우승하는 방법이라는 강좌와 관련된 경진대회다.
* 일단위로 판매되는 데이터의 시계열 분석을 다루고 있다.
* [1C Company](http://1c.ru/eng/title.htm)라는 러시아의 큰 소프트웨어 회사의 데이터이다.
* 1C Company는 경영 회계, 재무 회계, 인사 관리, CRM, SRM, MRP, MRP 등과 같은 다양한 비즈니스 업무를 하고있다.
```
import pandas as pd
import matplotlib.pyplot as plt
from plotnine import *
%matplotlib inline
%ls data
sales_train = pd.read_csv('data/sales_train.csv.gz', compression='gzip')
test = pd.read_csv('data/test.csv.gz', compression='gzip')
item_categories = pd.read_csv('data/item_categories.csv')
items = pd.read_csv('data/items.csv')
shops = pd.read_csv('data/shops.csv')
submissions = pd.read_csv('data/sample_submission.csv.gz', compression='gzip')
print(sales_train.shape)
print(test.shape)
print(item_categories.shape)
print(items.shape)
print(shops.shape)
print(submissions.shape)
sales_train.head()
test.head()
submissions.head()
sales_train.isnull().sum()
```
train데이터에는 2013년 1월부터 2015년 10월까지의 데이터가 있다.<br>
test데이터로 2015년 11월 shop과 product 정보를 예측해야 한다.<br>
`sample_submission.csv.gz` 파일을 보면 item_cnt_month 를 예측하게 되어있다.
```
[c for c in sales_train.columns if c not in test.columns]
shops.head()
items.head()
%%time
monthly_sales = sales_train.groupby([
"date_block_num","shop_id","item_id"])[
"date","item_price","item_cnt_day"].agg(
{"date":["min",'max'], "item_price":"mean", "item_cnt_day":"sum"})
monthly_sales.head(20)
items.item_category_id.head()
item_categories.head()
sales_train.item_cnt_day.plot()
plt.title("Number of products sold per day")
sales_train.item_price.hist()
plt.title("Item Price Distribution")
# 카테고리별 아이템 수
x = items.groupby(['item_category_id']).count()
x = x.sort_values(by='item_id', ascending=False)
x = x.iloc[0:10].reset_index()
x
```
* 40번 카테고리의 5035 아이템 판매수가 가장 많다.
```
x['item_category_id'] = x['item_category_id'].astype(str)
x['item_name'] = x['item_name'].astype(str)
(ggplot(x)
+ aes(x='item_category_id', y='item_id', fill='item_name')
+ geom_bar(stat = "identity")
+ ggtitle('카테고리별 아이템수')
+ theme(text=element_text(family='NanumBarunGothic'))
)
```
* https://www.kaggle.com/jagangupta/time-series-basics-exploring-traditional-ts 를 참고했다.
```
ts = sales_train.groupby(["date_block_num"])["item_cnt_day"].sum()
ts.astype('float')
ts = pd.DataFrame(ts).reset_index()
```
* 10일 단위로 판매 아이템이 뛰는 날이 있다. 일정한 주기별 패턴을 갖는 경향이 있다.
```
(ggplot(ts)
+ aes(x='date_block_num', y='item_cnt_day')
+ geom_point()
+ geom_line(color='blue')
+ labs(x='시간', y='판매 아이템 수', title='판매 아이템수')
+ theme(text=element_text(family='NanumBarunGothic'))
)
```
### 이동 평균과 표준편차
* 참고 : [이동 평균](https://docs.tibco.com/pub/spotfire_web_player/6.0.0-november-2013/ko-KR/WebHelp/GUID-5A18B4F1-8465-4200-881A-8721BF1A48B1.html)
* moving average, rolling average, rolling mean 또는 running average라고도 함
* 지정된 간격 안에서 노드 평균을 계산하는 데 사용
* 간격 크기가 3으로 설정된 경우 현재 노드와 앞의 두 노드를 사용하여 평균을 계산
이동 평균을 사용하는 목적은 일반적으로 단기 변동을 평준화하고 장기 동향을 파악하는 목적으로 사용
* [pandas.DataFrame.rolling — pandas 0.23.4 documentation](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.rolling.html)
```
# 이동 평균과 표준편차
plt.figure(figsize=(16,6))
plt.plot(ts.rolling(window=12,center=False).mean(),label='Rolling Mean')
plt.plot(ts.rolling(window=12,center=False).std(),label='Rolling sd')
plt.legend()
```
| github_jupyter |
# Collections
The collection Module in Python provides different types of containers. A Container is an object that is used to store different objects and provide a way to access the contained objects and iterate over them. Some of the built-in containers are Tuple, List, Dictionary, etc. In this article, we will discuss the different containers provided by the collections module.
```
import collections
help(collections)
```
### Counter
```
# A Python program to show different
# ways to create Counter
from collections import Counter
# With sequence of items
print(Counter(['B','B','A','B','C','A','B','B','A','C']))
# with dictionary
print(Counter({'A':3, 'B':5, 'C':2}))
# with keyword arguments
print(Counter(A=3, B=5, C=2))
```
### OrderedDict
```
# A Python program to demonstrate working
# of OrderedDict
from collections import OrderedDict
print("This is a Dict:\n")
d = {}
d['a'] = 1
d['b'] = 2
d['c'] = 3
d['d'] = 4
for key, value in d.items():
print(key, value)
print("\nThis is an Ordered Dict:\n")
od = OrderedDict()
od['a'] = 1
od['b'] = 2
od['c'] = 3
od['d'] = 4
for key, value in od.items():
print(key, value)
# A Python program to demonstrate working
# of OrderedDict
from collections import OrderedDict
od = OrderedDict()
od['a'] = 1
od['b'] = 2
od['c'] = 3
od['d'] = 4
print('Before Deleting')
for key, value in od.items():
print(key, value)
# deleting element
od.pop('a')
# Re-inserting the same
od['a'] = 1
print('\nAfter re-inserting')
for key, value in od.items():
print(key, value)
```
### DefaultDict
```
# Python program to demonstrate
# defaultdict
from collections import defaultdict
# Defining the dict
d = defaultdict(int)
L = [1, 2, 3, 4, 2, 4, 1, 2]
# Iterate through the list
# for keeping the count
for i in L:
# The default value is 0
# so there is no need to
# enter the key first
d[i] += 1
print(d)
# Python program to demonstrate
# defaultdict
from collections import defaultdict
# Defining a dict
d = defaultdict(list)
for i in range(5):
d[i].append(i)
print("Dictionary with values as list:")
print(d)
```
### ChainMap
A ChainMap encapsulates many dictionaries into a single unit and returns a list of dictionaries.
```
# Python program to demonstrate
# ChainMap
from collections import ChainMap
d1 = {'a': 1, 'b': 2}
d2 = {'c': 3, 'd': 4}
d3 = {'e': 5, 'f': 6}
# Defining the chainmap
c = ChainMap(d1, d2, d3)
print(c)
# Accessing value from chainmap
# Python program to demonstrate
# ChainMap
from collections import ChainMap
d1 = {'a': 1, 'b': 2}
d2 = {'c': 3, 'd': 4}
d3 = {'e': 5, 'a': 6}
# Defining the chainmap
c = ChainMap(d1, d2, d3)
# Accessing Values using key name
print(c['a'])
# Accesing values using values()
# method
print(c.values())
# Accessing keys using keys()
# method
print(c.keys())
# Adding new dictionary
# A new dictionary can be added by using the new_child() method. The newly added dictionary is added at the beginning of the ChainMap.
# Python code to demonstrate ChainMap and
# new_child()
import collections
# initializing dictionaries
dic1 = { 'a' : 1, 'b' : 2 }
dic2 = { 'b' : 3, 'c' : 4 }
dic3 = { 'f' : 5 }
# initializing ChainMap
chain = collections.ChainMap(dic1, dic2)
# printing chainMap
print ("All the ChainMap contents are : ")
print (chain)
# using new_child() to add new dictionary
chain1 = chain.new_child(dic3)
# printing chainMap
print ("Displaying new ChainMap : ")
print (chain1)
```
### NamedTuple
A NamedTuple returns a tuple object with names for each position which the ordinary tuples lack. For example, consider a tuple names student where the first element represents fname, second represents lname and the third element represents the DOB. Suppose for calling fname instead of remembering the index position you can actually call the element by using the fname argument, then it will be really easy for accessing tuples element. This functionality is provided by the NamedTuple.
```
from collections import namedtuple
# Declaring namedtuple()
Student = namedtuple('Student',['name','age','DOB'])
# Adding values
S = Student('Nandini','19','2541997')
# Access using index
print ("The Student age using index is : ",end ="")
print (S[1])
# Access using name
print ("The Student name using keyname is : ",end ="")
print (S.name)
```
###
Conversion Operations
1. _make(): This function is used to return a namedtuple() from the iterable passed as argument.
2. _asdict(): This function returns the OrdereDict() as constructed from the mapped values of namedtuple().
```
# Python code to demonstrate namedtuple() and
# _make(), _asdict()
from collections import namedtuple
# Declaring namedtuple()
Student = namedtuple('Student',['name','age','DOB'])
# Adding values
S = Student('Nandini','19','2541997')
# initializing iterable
li = ['Manjeet', '19', '411997' ]
# initializing dict
di = { 'name' : "Nikhil", 'age' : 19 , 'DOB' : '1391997' }
# using _make() to return namedtuple()
print ("The namedtuple instance using iterable is : ")
print (Student._make(li))
# using _asdict() to return an OrderedDict()
print ("The OrderedDict instance using namedtuple is : ")
print (S._asdict())
```
### Deque
Deque (Doubly Ended Queue) is the optimized list for quicker append and pop operations from both sides of the container. It provides O(1) time complexity for append and pop operations as compared to list with O(n) time complexity.
```
# Python code to demonstrate deque
from collections import deque
# Declaring deque
queue = deque(['name','age','DOB'])
print(queue)
```
### Inserting Elements
Elements in deque can be inserted from both ends. To insert the elements from right append() method is used and to insert the elements from the left appendleft() method is used.
```
from collections import deque
# initializing deque
de = deque([1,2,3])
# using append() to insert element at right end
# inserts 4 at the end of deque
de.append(4)
# printing modified deque
print ("The deque after appending at right is : ")
print (de)
# using appendleft() to insert element at right end
# inserts 6 at the beginning of deque
de.appendleft(6)
# printing modified deque
print ("The deque after appending at left is : ")
print (de)
```
### Removing Elements
Elements can also be removed from the deque from both the ends. To remove elements from right use pop() method and to remove elements from the left use popleft() method.
```
# Python code to demonstrate working of
# pop(), and popleft()
from collections import deque
# initializing deque
de = deque([6, 1, 2, 3, 4])
# using pop() to delete element from right end
# deletes 4 from the right end of deque
de.pop()
# printing modified deque
print ("The deque after deleting from right is : ")
print (de)
# using popleft() to delete element from left end
# deletes 6 from the left end of deque
de.popleft()
# printing modified deque
print ("The deque after deleting from left is : ")
print (de)
```
### UserDict
UserDict is a dictionary-like container that acts as a wrapper around the dictionary objects. This container is used when someone wants to create their own dictionary with some modified or new functionality.
```
from collections import UserDict
# Creating a Dictionary where
# deletion is not allowed
class MyDict(UserDict):
# Function to stop deleltion
# from dictionary
def __del__(self):
raise RuntimeError("Deletion not allowed")
# Function to stop pop from
# dictionary
def pop(self, s = None):
raise RuntimeError("Deletion not allowed")
# Function to stop popitem
# from Dictionary
def popitem(self, s = None):
raise RuntimeError("Deletion not allowed")
# Driver's code
d = MyDict({'a':1,'b': 2,'c': 3})
d.pop(1)
```
### UserList
UserList is a list like container that acts as a wrapper around the list objects. This is useful when someone wants to create their own list with some modified or additional functionality.
```
# Python program to demonstrate
# userlist
from collections import UserList
# Creating a List where
# deletion is not allowed
class MyList(UserList):
# Function to stop deleltion
# from List
def remove(self, s = None):
raise RuntimeError("Deletion not allowed")
# Function to stop pop from
# List
def pop(self, s = None):
raise RuntimeError("Deletion not allowed")
# Driver's code
L = MyList([1, 2, 3, 4])
print("Original List")
# Inserting to List"
L.append(5)
print("After Insertion")
print(L)
# Deliting From List
L.remove()
```
### UserString
UserString is a string like container and just like UserDict and UserList it acts as a wrapper around string objects. It is used when someone wants to create their own strings with some modified or additional functionality.
```
# Python program to demonstrate
# userstring
from collections import UserString
# Creating a Mutable String
class Mystring(UserString):
# Function to append to
# string
def append(self, s):
self.data += s
# Function to rmeove from
# string
def remove(self, s):
self.data = self.data.replace(s, "")
# Driver's code
s1 = Mystring("Geeks")
print("Original String:", s1.data)
# Appending to string
s1.append("s")
print("String After Appending:", s1.data)
# Removing from string
s1.remove("e")
print("String after Removing:", s1.data)
```
| github_jupyter |
#### Jupyter notebooks
This is a [Jupyter](http://jupyter.org/) notebook using Python. You can install Jupyter locally to edit and interact with this notebook.
# Finite Difference methods in 2 dimensions
Let's start by generalizing the 1D Laplacian,
\begin{align} - u''(x) &= f(x) \text{ on } \Omega = (a,b) & u(a) &= g_0(a) & u'(b) = g_1(b) \end{align}
to two dimensions
\begin{align} -\nabla\cdot \big( \nabla u(x,y) \big) &= f(x,y) \text{ on } \Omega \subset \mathbb R^2
& u|_{\Gamma_D} &= g_0(x,y) & \nabla u \cdot \hat n|_{\Gamma_N} &= g_1(x,y)
\end{align}
where $\Omega$ is some well-connected open set (we will assume simply connected) and the Dirichlet boundary $\Gamma_D \subset \partial \Omega$ is nonempty.
We need to choose a system for specifying the domain $\Omega$ and ordering degrees of freedom. Perhaps the most significant limitation of finite difference methods is that this specification is messy for complicated domains. We will choose
$$ \Omega = (0, 1) \times (0, 1) $$
and
\begin{align} (x, y)_{im+j} &= (i h, j h) & h &= 1/(m-1) & i,j \in \{0, 1, \dotsc, m-1 \} .
\end{align}
The Laplacian $-\nabla\cdot\nabla u = -u_{xx} - u_{yy}$ decomposes into directional derivatives, leading to the 2D stencil
$$ \frac 1 {h^2} \begin{pmatrix} & -1 & \\ -1 & 4 & -1 \\ & -1 & \end{pmatrix} . $$
```
%matplotlib inline
import numpy
from matplotlib import pyplot
pyplot.style.use('ggplot')
def laplacian2d_dense(h, f, g0):
"""Solve Laplacian(u) = f on [0,1]^2 with u=g0 on the boundary.
Use a discretization of nominal size h.
"""
m = int(1/h + 1) # Number of elements in terms of nominal grid spacing h
h = 1 / (m-1) # Actual grid spacing
c = numpy.linspace(0, 1, m)
y, x = numpy.meshgrid(c, c)
u0 = g0(x, y).flatten()
rhs = f(x, y).flatten()
A = numpy.zeros((m*m, m*m))
def idx(i, j):
return i*m + j
for i in range(m):
for j in range(m):
row = idx(i, j)
if i in (0, m-1) or j in (0, m-1):
A[row, row] = 1
rhs[row] = u0[row]
else:
cols = [idx(*pair) for pair in
[(i-1, j), (i, j-1), (i, j), (i, j+1), (i+1, j)]]
stencil = 1/h**2 * numpy.array([-1, -1, 4, -1, -1])
A[row, cols] = stencil
return x, y, A, rhs
x, y, A, rhs = laplacian2d_dense(.1,
lambda x,y: 0*x+1,
lambda x,y: 0*x)
pyplot.spy(A);
u = numpy.linalg.solve(A, rhs).reshape(x.shape)
pyplot.contourf(x, y, u)
pyplot.colorbar();
import cProfile, pstats
prof = cProfile.Profile()
prof.enable()
x, y, A, rhs = laplacian2d_dense(.02, lambda x,y: 0*x+1, lambda x,y: 0*x)
u = numpy.linalg.solve(A, rhs).reshape(x.shape)
prof.disable()
pstats.Stats(prof).sort_stats(pstats.SortKey.TIME).print_stats(10);
import scipy.sparse as sp
import scipy.sparse.linalg
def laplacian2d(h, f, g0):
m = int(1/h + 1) # Number of elements in terms of nominal grid spacing h
h = 1 / (m-1) # Actual grid spacing
c = numpy.linspace(0, 1, m)
y, x = numpy.meshgrid(c, c)
u0 = g0(x, y).flatten()
rhs = f(x, y).flatten()
A = sp.lil_matrix((m*m, m*m))
def idx(i, j):
return i*m + j
mask = numpy.zeros_like(x, dtype=int)
mask[1:-1,1:-1] = 1
mask = mask.flatten()
for i in range(m):
for j in range(m):
row = idx(i, j)
stencili = numpy.array([idx(*pair) for pair in
[(i-1, j), (i, j-1),
(i, j),
(i, j+1), (i+1, j)]])
stencilw = 1/h**2 * numpy.array([-1, -1, 4, -1, -1])
if mask[row] == 0: # Dirichlet boundary
A[row, row] = 1
rhs[row] = u0[row]
else:
smask = mask[stencili]
cols = stencili[smask == 1]
A[row, cols] = stencilw[smask == 1]
bdycols = stencili[smask == 0]
rhs[row] -= stencilw[smask == 0] @ u0[bdycols]
return x, y, A.tocsr(), rhs
x, y, A, rhs = laplacian2d(.1, lambda x,y: 0*x+1, lambda x,y: 0*x)
pyplot.spy(A.todense());
sp.linalg.norm(A - A.T)
prof = cProfile.Profile()
prof.enable()
x, y, A, rhs = laplacian2d(.01, lambda x,y: 0*x+1, lambda x,y: 0*x)
u = sp.linalg.spsolve(A, rhs).reshape(x.shape)
prof.disable()
pstats.Stats(prof).sort_stats(pstats.SortKey.TIME).print_stats(10);
```
## A manufactured solution
```
class mms0:
def u(x, y):
return x*numpy.exp(-x)*numpy.tanh(y)
def grad_u(x, y):
return numpy.array([(1 - x)*numpy.exp(-x)*numpy.tanh(y),
x*numpy.exp(-x)*(1 - numpy.tanh(y)**2)])
def laplacian_u(x, y):
return ((2 - x)*numpy.exp(-x)*numpy.tanh(y)
- 2*x*numpy.exp(-x)*(numpy.tanh(y)**2 - 1)*numpy.tanh(y))
def grad_u_dot_normal(x, y, n):
return grad_u(x, y) @ n
x, y, A, rhs = laplacian2d(.02, mms0.laplacian_u, mms0.u)
u = sp.linalg.spsolve(A, rhs).reshape(x.shape)
print(u.shape, numpy.linalg.norm((u - mms0.u(x,y)).flatten(), numpy.inf))
pyplot.contourf(x, y, u)
pyplot.colorbar()
pyplot.title('Numeric solution')
pyplot.figure()
pyplot.contourf(x, y, u - mms0.u(x, y))
pyplot.colorbar()
pyplot.title('Error');
hs = numpy.geomspace(.01, .25, 12)
def mms_error(h):
x, y, A, rhs = laplacian2d(h, mms0.laplacian_u, mms0.u)
u = sp.linalg.spsolve(A, rhs).reshape(x.shape)
return numpy.linalg.norm((u - mms0.u(x, y)).flatten(), numpy.inf)
pyplot.loglog(hs, [mms_error(h) for h in hs], 'o', label='numeric error')
pyplot.loglog(hs, hs**1/100, label='$h^1/100$')
pyplot.loglog(hs, hs**2/100, label='$h^2/100$')
pyplot.legend();
```
# Neumann boundary conditions
Recall that in 1D, we would reflect the solution into ghost points according to
$$ u_{-i} = u_i - (x_i - x_{-i}) g_1(x_0, y) $$
and similarly for the right boundary and in the $y$ direction. After this, we (optionally) scale the row in the matrix for symmetry and shift the known parts to the right hand side. Below, we implement the reflected symmetry, but not the inhomogeneous contribution or rescaling of the matrix row.
```
def laplacian2d_bc(h, f, g0, dirichlet=((),())):
m = int(1/h + 1) # Number of elements in terms of nominal grid spacing h
h = 1 / (m-1) # Actual grid spacing
c = numpy.linspace(0, 1, m)
y, x = numpy.meshgrid(c, c)
u0 = g0(x, y).flatten()
rhs = f(x, y).flatten()
ai = []
aj = []
av = []
def idx(i, j):
i = (m-1) - abs(m-1 - abs(i))
j = (m-1) - abs(m-1 - abs(j))
return i*m + j
mask = numpy.ones_like(x, dtype=int)
mask[dirichlet[0],:] = 0
mask[:,dirichlet[1]] = 0
mask = mask.flatten()
for i in range(m):
for j in range(m):
row = idx(i, j)
stencili = numpy.array([idx(*pair) for pair in [(i-1, j), (i, j-1), (i, j), (i, j+1), (i+1, j)]])
stencilw = 1/h**2 * numpy.array([-1, -1, 4, -1, -1])
if mask[row] == 0: # Dirichlet boundary
ai.append(row)
aj.append(row)
av.append(1)
rhs[row] = u0[row]
else:
smask = mask[stencili]
ai += [row]*sum(smask)
aj += stencili[smask == 1].tolist()
av += stencilw[smask == 1].tolist()
bdycols = stencili[smask == 0]
rhs[row] -= stencilw[smask == 0] @ u0[bdycols]
A = sp.csr_matrix((av, (ai, aj)), shape=(m*m,m*m))
return x, y, A, rhs
x, y, A, rhs = laplacian2d_bc(.05, lambda x,y: 0*x+1,
lambda x,y: 0*x, dirichlet=((0,),()))
u = sp.linalg.spsolve(A, rhs).reshape(x.shape)
print(numpy.real_if_close(sp.linalg.eigs(A, 10, which='SM')[0]))
pyplot.contourf(x, y, u)
pyplot.colorbar();
# We used a different technique for assembling the sparse matrix.
# This is faster with scipy.sparse, but may be worse for other sparse matrix packages, such as PETSc.
prof = cProfile.Profile()
prof.enable()
x, y, A, rhs = laplacian2d_bc(.02, lambda x,y: 0*x+1, lambda x,y: 0*x)
u = sp.linalg.spsolve(A, rhs).reshape(x.shape)
prof.disable()
pstats.Stats(prof).sort_stats(pstats.SortKey.TIME).print_stats(10);
```
# Variable coefficients
In physical systems, it is common for equations to be given in **divergence form** (sometimes called **conservative form**),
$$ -\nabla\cdot \Big( \kappa(x,y) \nabla u \Big) = f(x,y) . $$
This can be converted to **non-divergence form**,
$$ - \kappa(x,y) \nabla\cdot \nabla u - \nabla \kappa(x,y) \cdot \nabla u = f(x,y) . $$
* What assumptions did we just make on $\kappa(x,y)$?
```
def laplacian2d_nondiv(h, f, kappa, grad_kappa, g0, dirichlet=((),())):
m = int(1/h + 1) # Number of elements in terms of nominal grid spacing h
h = 1 / (m-1) # Actual grid spacing
c = numpy.linspace(0, 1, m)
y, x = numpy.meshgrid(c, c)
u0 = g0(x, y).flatten()
rhs = f(x, y).flatten()
ai = []
aj = []
av = []
def idx(i, j):
i = (m-1) - abs(m-1 - abs(i))
j = (m-1) - abs(m-1 - abs(j))
return i*m + j
mask = numpy.ones_like(x, dtype=int)
mask[dirichlet[0],:] = 0
mask[:,dirichlet[1]] = 0
mask = mask.flatten()
for i in range(m):
for j in range(m):
row = idx(i, j)
stencili = numpy.array([idx(*pair) for pair in [(i-1, j), (i, j-1), (i, j), (i, j+1), (i+1, j)]])
stencilw = kappa(i*h, j*h)/h**2 * numpy.array([-1, -1, 4, -1, -1])
if grad_kappa is None:
gk = 1/h * numpy.array([kappa((i+.5)*h,j*h) - kappa((i-.5)*h,j*h),
kappa(i*h,(j+.5)*h) - kappa(i*h,(j-.5)*h)])
else:
gk = grad_kappa(i*h, j*h)
stencilw -= gk[0] / (2*h) * numpy.array([-1, 0, 0, 0, 1])
stencilw -= gk[1] / (2*h) * numpy.array([0, -1, 0, 1, 0])
if mask[row] == 0: # Dirichlet boundary
ai.append(row)
aj.append(row)
av.append(1)
rhs[row] = u0[row]
else:
smask = mask[stencili]
ai += [row]*sum(smask)
aj += stencili[smask == 1].tolist()
av += stencilw[smask == 1].tolist()
bdycols = stencili[smask == 0]
rhs[row] -= stencilw[smask == 0] @ u0[bdycols]
A = sp.csr_matrix((av, (ai, aj)), shape=(m*m,m*m))
return x, y, A, rhs
def kappa(x, y):
#return 1 - 2*(x-.5)**2 - 2*(y-.5)**2
return 1e-2 + 2*(x-.5)**2 + 2*(y-.5)**2
def grad_kappa(x, y):
#return -4*(x-.5), -4*(y-.5)
return 4*(x-.5), 4*(y-.5)
pyplot.contourf(x, y, kappa(x,y))
pyplot.colorbar();
x, y, A, rhs = laplacian2d_nondiv(.05, lambda x,y: 0*x+1,
kappa, grad_kappa,
lambda x,y: 0*x, dirichlet=((0,-1),()))
u = sp.linalg.spsolve(A, rhs).reshape(x.shape)
pyplot.contourf(x, y, u)
pyplot.colorbar();
x, y, A, rhs = laplacian2d_nondiv(.05, lambda x,y: 0*x,
kappa, grad_kappa,
lambda x,y: x, dirichlet=((0,-1),()))
u = sp.linalg.spsolve(A, rhs).reshape(x.shape)
pyplot.contourf(x, y, u)
pyplot.colorbar();
def laplacian2d_div(h, f, kappa, g0, dirichlet=((),())):
m = int(1/h + 1) # Number of elements in terms of nominal grid spacing h
h = 1 / (m-1) # Actual grid spacing
c = numpy.linspace(0, 1, m)
y, x = numpy.meshgrid(c, c)
u0 = g0(x, y).flatten()
rhs = f(x, y).flatten()
ai = []
aj = []
av = []
def idx(i, j):
i = (m-1) - abs(m-1 - abs(i))
j = (m-1) - abs(m-1 - abs(j))
return i*m + j
mask = numpy.ones_like(x, dtype=int)
mask[dirichlet[0],:] = 0
mask[:,dirichlet[1]] = 0
mask = mask.flatten()
for i in range(m):
for j in range(m):
row = idx(i, j)
stencili = numpy.array([idx(*pair) for pair in [(i-1, j),
(i, j-1),
(i, j),
(i, j+1),
(i+1, j)]])
stencilw = 1/h**2 * ( kappa((i-.5)*h, j*h) * numpy.array([-1, 0, 1, 0, 0])
+ kappa(i*h, (j-.5)*h) * numpy.array([0, -1, 1, 0, 0])
- kappa(i*h, (j+.5)*h) * numpy.array([0, 0, -1, 1, 0])
- kappa((i+.5)*h, j*h) * numpy.array([0, 0, -1, 0, 1]))
if mask[row] == 0: # Dirichlet boundary
ai.append(row)
aj.append(row)
av.append(1)
rhs[row] = u0[row]
else:
smask = mask[stencili]
ai += [row]*sum(smask)
aj += stencili[smask == 1].tolist()
av += stencilw[smask == 1].tolist()
bdycols = stencili[smask == 0]
rhs[row] -= stencilw[smask == 0] @ u0[bdycols]
A = sp.csr_matrix((av, (ai, aj)), shape=(m*m,m*m))
return x, y, A, rhs
x, y, A, rhs = laplacian2d_div(.05, lambda x,y: 0*x+1,
kappa,
lambda x,y: 0*x, dirichlet=((0,-1),()))
u = sp.linalg.spsolve(A, rhs).reshape(x.shape)
pyplot.contourf(x, y, u)
pyplot.colorbar();
x, y, A, rhs = laplacian2d_div(.05, lambda x,y: 0*x,
kappa,
lambda x,y: x, dirichlet=((0,-1),()))
u = sp.linalg.spsolve(A, rhs).reshape(x.shape)
pyplot.contourf(x, y, u)
pyplot.colorbar();
x, y, A, rhs = laplacian2d_nondiv(.05, lambda x,y: 0*x+1,
kappa, grad_kappa,
lambda x,y: 0*x, dirichlet=((0,-1),()))
u_nondiv = sp.linalg.spsolve(A, rhs).reshape(x.shape)
x, y, A, rhs = laplacian2d_div(.05, lambda x,y: 0*x+1,
kappa,
lambda x,y: 0*x, dirichlet=((0,-1),()))
u_div = sp.linalg.spsolve(A, rhs).reshape(x.shape)
pyplot.contourf(x, y, u_nondiv - u_div)
pyplot.colorbar();
class mms1:
def __init__(self):
import sympy
x, y = sympy.symbols('x y')
uexpr = x*sympy.exp(-2*x) * sympy.tanh(1.2*y+.1)
kexpr = 1e-2 + 2*(x-.42)**2 + 2*(y-.51)**2
self.u = sympy.lambdify((x,y), uexpr)
self.kappa = sympy.lambdify((x,y), kexpr)
def grad_kappa(xx, yy):
kx = sympy.lambdify((x,y), sympy.diff(kexpr, x))
ky = sympy.lambdify((x,y), sympy.diff(kexpr, y))
return kx(xx, yy), ky(xx, yy)
self.grad_kappa = grad_kappa
self.div_kappa_grad_u = sympy.lambdify((x,y),
-( sympy.diff(kexpr * sympy.diff(uexpr, x), x)
+ sympy.diff(kexpr * sympy.diff(uexpr, y), y)))
mms = mms1()
x, y, A, rhs = laplacian2d_nondiv(.05, mms.div_kappa_grad_u,
mms.kappa, mms.grad_kappa,
mms.u, dirichlet=((0,-1),(0,-1)))
u_nondiv = sp.linalg.spsolve(A, rhs).reshape(x.shape)
pyplot.contourf(x, y, u_nondiv)
pyplot.colorbar()
numpy.linalg.norm((u_nondiv - mms.u(x, y)).flatten(), numpy.inf)
x, y, A, rhs = laplacian2d_div(.05, mms.div_kappa_grad_u,
mms.kappa,
mms.u, dirichlet=((0,-1),(0,-1)))
u_div = sp.linalg.spsolve(A, rhs).reshape(x.shape)
pyplot.contourf(x, y, u_div)
pyplot.colorbar()
numpy.linalg.norm((u_div - mms.u(x, y)).flatten(), numpy.inf)
def mms_error(h):
x, y, A, rhs = laplacian2d_nondiv(h, mms.div_kappa_grad_u,
mms.kappa, mms.grad_kappa,
mms.u, dirichlet=((0,-1),(0,-1)))
u_nondiv = sp.linalg.spsolve(A, rhs).flatten()
x, y, A, rhs = laplacian2d_div(h, mms.div_kappa_grad_u,
mms.kappa, mms.u, dirichlet=((0,-1),(0,-1)))
u_div = sp.linalg.spsolve(A, rhs).flatten()
u_exact = mms.u(x, y).flatten()
return numpy.linalg.norm(u_nondiv - u_exact, numpy.inf), numpy.linalg.norm(u_div - u_exact, numpy.inf)
hs = numpy.logspace(-1.5, -.5, 10)
errors = numpy.array([mms_error(h) for h in hs])
pyplot.loglog(hs, errors[:,0], 'o', label='nondiv')
pyplot.loglog(hs, errors[:,1], 's', label='div')
pyplot.plot(hs, hs**2, label='$h^2$')
pyplot.legend();
float(False)
def kappablob(x, y):
#return .01 + ((x-.5)**2 + (y-.5)**2 < .125)
return .01 + (numpy.abs(x-.505) < .25) # + (numpy.abs(y-.5) < .25)
x, y, A, rhs = laplacian2d_div(.05, lambda x,y: 0*x, kappablob,
lambda x,y:x, dirichlet=((0,-1),()))
u_div = sp.linalg.spsolve(A, rhs).reshape(x.shape)
pyplot.contourf(x, y, kappablob(x, y))
pyplot.colorbar();
pyplot.figure()
pyplot.contourf(x, y, u_div, 10)
pyplot.colorbar();
x, y, A, rhs = laplacian2d_nondiv(.05, lambda x,y: 0*x, kappablob, None,
lambda x,y:x, dirichlet=((0,-1),()))
u_nondiv = sp.linalg.spsolve(A, rhs).reshape(x.shape)
pyplot.contourf(x, y, u_nondiv, 10)
pyplot.colorbar();
```
## Weak forms
When we write
$$ {\huge "} - \nabla\cdot \big( \kappa \nabla u \big) = 0 {\huge "} \text{ on } \Omega $$
where $\kappa$ is a discontinuous function, that's not exactly what we mean the derivative of that discontinuous function doesn't exist. Formally, however, let us multiply by a "test function" $v$ and integrate,
\begin{split}
- \int_\Omega v \nabla\cdot \big( \kappa \nabla u \big) = 0 & \text{ for all } v \\
\int_\Omega \nabla v \cdot \kappa \nabla u = \int_{\partial \Omega} v \kappa \nabla u \cdot \hat n & \text{ for all } v
\end{split}
where we have used integration by parts. This is called the **weak form** of the PDE and will be what we actually discretize using finite element methods. All the terms make sense when $\kappa$ is discontinuous. Now suppose our domain is decomposed into two disjoint sub domains $$\overline{\Omega_1 \cup \Omega_2} = \overline\Omega $$
with interface $$\Gamma = \overline\Omega_1 \cap \overline\Omega_2$$ and $\kappa_1$ is continuous on $\Omega_1$ and $\kappa_2$ is continuous on $\Omega_2$, but possibly $\kappa_1(x) \ne \kappa_2(x)$ for $x \in \Gamma$,
\begin{split}
\int_\Omega \nabla v \cdot \kappa \nabla u &= \int_{\Omega_1} \nabla v \cdot \kappa_1\nabla u + \int_{\Omega_2} \nabla v \cdot \kappa_2 \nabla u \\
&= -\int_{\Omega_1} v \nabla\cdot \big(\kappa_1 \nabla u \big) + \int_{\partial \Omega_1} v \kappa_1 \nabla u \cdot \hat n \\
&\qquad -\int_{\Omega_2} v \nabla\cdot \big(\kappa_2 \nabla u \big) + \int_{\partial \Omega_2} v \kappa_2 \nabla u \cdot \hat n \\
&= -\int_{\Omega} v \nabla\cdot \big(\kappa \nabla u \big) + \int_{\partial \Omega} v \kappa \nabla u \cdot \hat n + \int_{\Gamma} v (\kappa_1 - \kappa_2) \nabla u\cdot \hat n .
\end{split}
* Which direction is $\hat n$ for the integral over $\Gamma$?
* Does it matter what we choose for the value of $\kappa$ on $\Gamma$ in the volume integral?
When $\kappa$ is continuous, the jump term vanishes and we recover the **strong form**
$$ - \nabla\cdot \big( \kappa \nabla u \big) = 0 \text{ on } \Omega . $$
But if $\kappa$ is discontinuous, we would need to augment this with a jump condition ensuring that the flux $-\kappa \nabla u$ is continuous. We could go add this condition to our FD code to recover convergence in case of discontinuous $\kappa$, but it is messy.
## Nonlinear problems
Let's consider the nonlinear problem
$$ -\nabla \cdot \big(\underbrace{(1 + u^2)}_{\kappa(u)} \nabla u \big) = f \text{ on } (0,1)^2 $$
subject to Dirichlet boundary conditions. We will discretize the divergence form and thus will need
$\kappa(u)$ evaluated at staggered points $(i-1/2,j)$, $(i,j-1/2)$, etc. We will calculate these by averaging
$$ u_{i-1/2,j} = \frac{u_{i-1,j} + u_{i,j}}{2} $$
and similarly for the other staggered directions.
To use a Newton method, we also need the derivatives
$$ \frac{\partial \kappa_{i-1/2,j}}{\partial u_{i,j}} = 2 u_{i-1/2,j} \frac{\partial u_{i-1/2,j}}{\partial u_{i,j}} = u_{i-1/2,j} . $$
In the function below, we compute both the residual
$$F(u) = -\nabla\cdot \kappa(u) \nabla u - f(x,y)$$
and its Jacobian
$$J(u) = \frac{\partial F}{\partial u} . $$
```
def hgrid(h):
m = int(1/h + 1) # Number of elements in terms of nominal grid spacing h
h = 1 / (m-1) # Actual grid spacing
c = numpy.linspace(0, 1, m)
y, x = numpy.meshgrid(c, c)
return x, y
def nonlinear2d_div(h, x, y, u, forcing, g0, dirichlet=((),())):
m = x.shape[0]
u0 = g0(x, y).flatten()
F = -forcing(x, y).flatten()
ai = []
aj = []
av = []
def idx(i, j):
i = (m-1) - abs(m-1 - abs(i))
j = (m-1) - abs(m-1 - abs(j))
return i*m + j
mask = numpy.ones_like(x, dtype=bool)
mask[dirichlet[0],:] = False
mask[:,dirichlet[1]] = False
mask = mask.flatten()
u = u.flatten()
F[mask == False] = u[mask == False] - u0[mask == False]
u[mask == False] = u0[mask == False]
for i in range(m):
for j in range(m):
row = idx(i, j)
stencili = numpy.array([idx(*pair) for pair in [(i-1, j), (i, j-1), (i, j), (i, j+1), (i+1, j)]])
# Stencil to evaluate gradient at four staggered points
grad = numpy.array([[-1, 0, 1, 0, 0],
[0, -1, 1, 0, 0],
[0, 0, -1, 1, 0],
[0, 0, -1, 0, 1]]) / h
# Stencil to average at four staggered points
avg = numpy.array([[1, 0, 1, 0, 0],
[0, 1, 1, 0, 0],
[0, 0, 1, 1, 0],
[0, 0, 1, 0, 1]]) / 2
# Stencil to compute divergence at cell centers from fluxes at four staggered points
div = numpy.array([-1, -1, 1, 1]) / h
ustencil = u[stencili]
ustag = avg @ ustencil
kappa = .1 + ustag**2
if mask[row] == 0: # Dirichlet boundary
ai.append(row)
aj.append(row)
av.append(1)
else:
F[row] -= div @ (kappa[:,None] * grad @ ustencil)
Jstencil = -div @ (kappa[:,None] * grad
+ 2*(ustag*(grad @ ustencil))[:,None] * avg)
smask = mask[stencili]
ai += [row]*sum(smask)
aj += stencili[smask].tolist()
av += Jstencil[smask].tolist()
J = sp.csr_matrix((av, (ai, aj)), shape=(m*m,m*m))
return F, J
h = .1
x, y = hgrid(h)
u = 0*x
# u += deltau # Uncomment to iterate
F, J = nonlinear2d_div(h, x, y, u, lambda x,y: 0*x+1,
lambda x,y: 0*x, dirichlet=((0,-1),(0,-1)))
deltau = sp.linalg.spsolve(J, -F).reshape(x.shape)
pyplot.contourf(x, y, deltau)
pyplot.colorbar();
def solve_nonlinear(h, g0, dirichlet, atol=1e-8, verbose=False):
x, y = hgrid(h)
u = 0*x
for i in range(50):
F, J = nonlinear2d_div(h, x, y, u, lambda x,y: 0*x+1,
g0=g0, dirichlet=dirichlet)
anorm = numpy.linalg.norm(F, numpy.inf)
if verbose:
print('{:2d}: anorm {:8e}'.format(i,anorm))
if anorm < atol:
break
deltau = sp.linalg.spsolve(J, -F)
u += deltau.reshape(x.shape)
return x, y, u, i
x, y, u, i = solve_nonlinear(.1, lambda x,y: 0*x,
dirichlet=((0,-1),(0,-1)),
verbose=True)
pyplot.contourf(x, y, u)
pyplot.colorbar();
```
## Homework 3: Wednesday, 2018-10-24
Write a solver for the regularized $p$-Laplacian,
$$ -\nabla\cdot\big( \kappa(\nabla u) \nabla u \big) = 0 $$
where
$$ \kappa(\nabla u) = \big(\frac 1 2 \epsilon^2 + \frac 1 2 \nabla u \cdot \nabla u \big)^{\frac{p-2}{2}}, $$
$ \epsilon > 0$, and $1 < p < \infty$. The case $p=2$ is the conventional Laplacian. This problem gets more strongly nonlinear when $p$ is far from 2 and when $\epsilon$ approaches zero. The $p \to 1$ limit is related to plasticity and has applications in non-Newtonion flows and structural mechanics.
1. Implement a "Picard" solver, which is like a Newton solver except that the Jacobian is replaced by the linear system
$$ J_{\text{Picard}}(u) \delta u \sim -\nabla\cdot\big( \kappa(\nabla u) \nabla \delta u \big) . $$
This is much easier to implement than the full Newton linearization. How fast does this method converge for values of $p < 2$ and $p > 2$?
* Use the linearization above as a preconditioner to a Newton-Krylov method. That is, use [`scipy.sparse.linalg.LinearOperator`](https://docs.scipy.org/doc/scipy/reference/generated/scipy.sparse.linalg.LinearOperator.html) to apply the Jacobian to a vector
$$ \tilde J(u) v = \frac{F(u + h v) - F(u)}{h} . $$
Then for each linear solve, use [`scipy.sparse.linalg.gmres`](https://docs.scipy.org/doc/scipy/reference/generated/scipy.sparse.linalg.gmres.html) and pass as a preconditioner, a direct solve with the Picard linearization above. (You might find [`scipy.sparse.linalg.factorized`](https://docs.scipy.org/doc/scipy/reference/generated/scipy.sparse.linalg.factorized.html) to be useful. Compare algebraic convergence to that of the Picard method.
* Can you directly implement a Newton linearization? Either do it or explain what is involved. How will its nonlinear convergence compare to that of the Newton-Krylov method?
# Wave equations and multi-component systems
The acoustic wave equation with constant wave speed $c$ can be written
$$ \ddot u - c^2 \nabla\cdot \nabla u = 0 $$
where $u$ is typically a pressure.
We can convert to a first order system
$$ \begin{bmatrix} \dot u \\ \dot v \end{bmatrix} = \begin{bmatrix} 0 & I \\ c^2 \nabla\cdot \nabla & 0 \end{bmatrix} \begin{bmatrix} u \\ v \end{bmatrix} . $$
We will choose a zero-penetration boundary condition $\nabla u \cdot \hat n = 0$, which will cause waves to reflect.
```
%run fdtools.py
x, y, L, _ = laplacian2d_bc(.1, lambda x,y: 0*x,
lambda x,y: 0*x, dirichlet=((),()))
A = sp.bmat([[None, sp.eye(*L.shape)],
[-L, None]])
eigs = sp.linalg.eigs(A, 10, which='LM')[0]
print(eigs)
maxeig = max(eigs.imag)
u0 = numpy.concatenate([numpy.exp(-8*(x**2 + y**2)), 0*x], axis=None)
hist = ode_rkexplicit(lambda t, u: A @ u, u0, tfinal=2, h=2/maxeig)
def plot_wave(x, y, time, U):
u = U[:x.size].reshape(x.shape)
pyplot.contourf(x, y, u)
pyplot.colorbar()
pyplot.title('Wave solution t={:f}'.format(time));
for step in numpy.linspace(0, len(hist)-1, 6, dtype=int):
pyplot.figure()
plot_wave(x, y, *hist[step]);
```
* This was a second order discretization, but we could extend it to higher order.
* The largest eigenvalues of this operator are proportional to $c/h$.
* Formally, we can write this equation in conservative form
$$ \begin{bmatrix} \dot u \\ \dot{\mathbf v} \end{bmatrix} = \begin{bmatrix} 0 & c\nabla\cdot \\ c \nabla & 0 \end{bmatrix} \begin{bmatrix} u \\ \mathbf v \end{bmatrix} $$
where $\mathbf{v}$ is now a momentum vector and $\nabla u = \nabla\cdot (u I)$. This formulation could produce an anti-symmetric ($A^T = -A$) discretization. Discretizations with this property are sometimes called "mimetic".
* A conservative form is often preferred when studiying waves traveling through materials with different wave speeds $c$.
* This is a Hamiltonian system. While high order Runge-Kutta methods can be quite accurate, "symplectic" time integrators are needed to preserve the structure of the Hamiltonian (related to energy conservation) over long periods of time. The midpoint method (aka $\theta=1/2$) is one such method. There are also explicit symplectic methods such as [Verlet methods](https://en.wikipedia.org/wiki/Verlet_integration), though these can be fragile.
| github_jupyter |
## Stanley tracker
Stanley controller was used in the DARPA-challenge winning autonomous vehicle, back in the day.
It is a non-linear controller, which explicitly takes into account **ref_theta** unlike pure pursuit. Because it was used for a car, a bicycle model is to be used for vehicular motion
Trackers are trying to minimize 2 types of errors
1. Positional error or cross-track error $x_e$
2. Heading error $\theta_e$
Tracker controls the steering as a function of these 2 errors
$\delta = \theta_e + \tan^{-1}\frac{kx_e}{v}$
The second term can be thought of as how quickly do we want the vehicle to compensate for the positional error relative to vehicle velocity
Snider has a nice/ concise summary of how it works in Sec 2.3 of his Phd thesis
https://www.ri.cmu.edu/pub_files/2009/2/Automatic_Steering_Methods_for_Autonomous_Automobile_Path_Tracking.pdf
There is a reference implementation here
https://github.com/AtsushiSakai/PythonRobotics/blob/master/PathTracking/stanley_controller/stanley_controller.py
+ Take the smoothened trajectory of straight/ right turn/ straight from week 2.
+ Induce a small error in initial pose.
+ Simulate vehicular motion using the Stanley tracker
```
import numpy as np
import matplotlib.pyplot as plt
rx = list(np.load("smooth_route_astar_grid_x.npy"))
ry = list(np.load("smooth_route_astar_grid_y.npy"))
grid = np.load("astar_grid.npy")
fig, ax = plt.subplots(figsize=(12,12))
ax.imshow(grid, cmap=plt.cm.Dark2)
ax.scatter(ry[0],rx[0], marker = "+", color = "yellow", s = 200)
ax.scatter(ry[-1],rx[-1], marker = "+", color = "red", s = 200)
ax.plot(ry,rx,'blue')
class PurePursuitTracker(object):
def __init__(self, x, y, v, lookahead = 3.0):
"""
Tracks the path defined by x, y at velocity v
x and y must be numpy arrays
v and lookahead are floats
"""
self.length = len(x)
self.ref_idx = 0 #index on the path that tracker is to track
self.lookahead = lookahead
self.x, self.y = x, y
self.v, self.w = v, 0
def update(self, xc, yc, theta):
"""
Input: xc, yc, theta - current pose of the robot
Update v, w based on current pose
Returns True if trajectory is over.
"""
#Calculate ref_x, ref_y using current ref_idx
#Check if we reached the end of path, then return TRUE
#Two conditions must satisfy
#1. ref_idx exceeds length of traj
#2. ref_x, ref_y must be within goal_threshold
# Write your code to check end condition
ref_x, ref_y = self.x[self.ref_idx], self.y[self.ref_idx]
goal_x, goal_y = self.x[-1], self.y[-1]
if (self.ref_idx == self.length-1) and (np.linalg.norm([ref_x-goal_x, ref_y-goal_y])) < goal_threshold:
return True
#End of path has not been reached
#update ref_idx using np.hypot([ref_x-xc, ref_y-yc]) < lookahead
while np.hypot(ref_x-xc, ref_y-yc) < self.lookahead:
self.ref_idx+= 1
ref_x, ref_y = self.x[self.ref_idx], self.y[self.ref_idx]
#Find the anchor point
# this is the line we drew between (0, 0) and (x, y)
anchor = np.asarray([ref_x - xc, ref_y - yc])
#Remember right now this is drawn from current robot pose
#we have to rotate the anchor to (0, 0, pi/2)
#code is given below for this
theta = np.pi/2 - theta
rot = np.asarray([[np.cos(theta), -np.sin(theta)], [np.sin(theta), np.cos(theta)]])
anchor = np.dot(rot, anchor)
# print(anchor)
L = np.sqrt(anchor[0] ** 2 + anchor[1] **2) # dist to reference path
X = -anchor[0] #cross-track error
#from the derivation in notes, plug in the formula for omega
self.w = 2*self.v*X/(L**2)
# print(self.w)
return False
#write code to instantiate the tracker class
vmax = 0.1
goal_threshold = 0.05
lookahead = 1
def simulate_unicycle(pose, v,w, dt=0.1):
x, y, t = pose
return x + v*np.cos(t)*dt, y + v*np.sin(t)*dt, t+w*dt
tracker = PurePursuitTracker(rx,ry,vmax,lookahead)
pose = (rx[0]+np.random.normal(0,0.1),ry[0]+np.random.normal(0,0.1),np.pi/2)
# pose = -1, 0, np.pi/2 #arbitrary initial pose
x0,y0,t0 = pose # record it for plotting
traj =[]
while True:
# for i in range(200):
#write the usual code to obtain successive poses
pose = simulate_unicycle(pose, tracker.v, tracker.w)
xc,yc,tc = pose
if tracker.update(xc,yc,tc) == True:
print("ARRIVED!!")
break
traj.append([*pose, tracker.w, tracker.ref_idx])
xs,ys,ts,deltas,ids = zip(*traj)
fig, ax = plt.subplots(figsize=(12,12))
ax.imshow(grid, cmap=plt.cm.Dark2)
ax.scatter(ry[0],rx[0], marker = "+", color = "yellow", s = 200)
ax.scatter(ry[-1],rx[-1], marker = "+", color = "red", s = 200)
ax.scatter(ry[ids[-1]],rx[ids[-1]],marker = "+",color = "orange", s = 100)
ax.plot(ry,rx,'blue')
ax.plot(ys,xs,'orange')
def simulate_bicycle(pose, v,delta, dt=0.1):
x, y, t = pose
return x + v*np.cos(t)*dt, y + v*np.sin(t)*dt, t+((v/0.9)*np.tan(delta)*dt)
class Stanley_tracker(object):
def __init__(self, x, y, v, lookahead = 3.0):
"""
Tracks the path defined by x, y at velocity v
x and y must be numpy arrays
v and lookahead are floats
"""
self.length = len(x)
self.ref_idx = 0 #index on the path that tracker is to track
self.lookahead = lookahead
self.x, self.y = x, y
self.v, self.delta = v, 0
self.k = 1/10
def update(self, xc, yc, thetac):
"""
Input: xc, yc, theta - current pose of the robot
Update v, w based on current pose
Returns True if trajectory is over.
"""
#Calculate ref_x, ref_y using current ref_idx
#Check if we reached the end of path, then return TRUE
#Two conditions must satisfy
#1. ref_idx exceeds length of traj
#2. ref_x, ref_y must be within goal_threshold
# Write your code to check end condition
ref_x, ref_y = self.x[self.ref_idx], self.y[self.ref_idx]
goal_x, goal_y = self.x[-1], self.y[-1]
if (self.ref_idx == self.length-1) and (np.linalg.norm([ref_x-goal_x, ref_y-goal_y])) < goal_threshold:
return True
#End of path has not been reached
#update ref_idx using np.hypot([ref_x-xc, ref_y-yc]) < lookahead
while (np.hypot(ref_x-xc, ref_y-yc) < self.lookahead) and (self.ref_idx < self.length):
self.ref_idx+= 1
ref_x, ref_y = self.x[self.ref_idx], self.y[self.ref_idx]
#Find the anchor point
# this is the line we drew between (0, 0) and (x, y)
anchor = np.asarray([ref_x - xc, ref_y - yc])
#Remember right now this is drawn from current robot pose
#we have to rotate the anchor to (0, 0, pi/2)
#code is given below for this
theta = np.pi/2 - thetac
rot = np.asarray([[np.cos(theta), -np.sin(theta)], [np.sin(theta), np.cos(theta)]])
anchor = np.dot(rot, anchor)
# print(anchor)
L = np.sqrt(anchor[0] ** 2 + anchor[1] **2) # dist to reference path
X = -anchor[0] #cross-track error
theta_path = np.arctan2((self.y[self.ref_idx-1] -ref_y),(self.x[self.ref_idx-1] -ref_x))
# print(theta_path)
theta_e = (theta_path - thetac)
if theta_e > np.pi:
theta_e -= 2.0 * np.pi
if theta_e < -np.pi:
theta_e += 2.0 * np.pi
#from the derivation in notes, plug in the formula for omega
# self.w = -2*self.v*X/(L**2)
gain = self.k*2*0.9/L
self.delta = (theta_e + np.arctan2(gain*X,self.v))
# self.delta = theta_e
return False
#write code to instantiate the tracker class
vmax = 0.3
goal_threshold = 0.05
lookahead = 0.8
tracker = Stanley_tracker(rx,ry,vmax,lookahead)
pose = (rx[0]+np.random.normal(0,0.1),ry[0]+np.random.normal(0,0.1),np.pi/2)
# pose = -1, 0, np.pi/2 #arbitrary initial pose
x0,y0,t0 = pose # record it for plotting
traj =[]
while True:
# for i in range(2000):
#write the usual code to obtain successive poses
pose = simulate_bicycle(pose, tracker.v, tracker.delta)
xc,yc,tc = pose
if tracker.update(xc,yc,tc) == True:
print("ARRIVED!!")
break
traj.append([*pose, tracker.delta, tracker.ref_idx])
xs,ys,ts,deltas,ids = zip(*traj)
fig, ax = plt.subplots(figsize=(12,12))
ax.imshow(grid, cmap=plt.cm.Dark2)
ax.scatter(ry[0],rx[0], marker = "+", color = "yellow", s = 200)
ax.scatter(ry[-1],rx[-1], marker = "+", color = "red", s = 200)
ax.scatter(ry[ids[-1]],rx[ids[-1]],marker = "+",color = "orange", s = 100)
ax.plot(ry,rx,'blue')
ax.plot(ys,xs,'orange')
```
| github_jupyter |
<pre>We will compute AUROC = Area Under the Receiver Operating Characteristic curve.
We will compute it using our own code and then use sklearn package to validate our scores.</pre>
#### Sections:
1. We will build a basic classifier model on titanic dataset
2. We will Compute the auc and roc_auc_score
3. We will Compute the auc and roc_auc_score using sklearn for validation
4. Plot both the metrics together for comparision
### Section 1: We will build a basic classifier model on titanic dataset
```
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
np.random.seed(42)
## Loading data and creating features. NOTE: df_test won't get used in this notebook
df_train = pd.read_csv('../data/titanic_train.csv')
df_test = pd.read_csv('../data/titanic_test.csv')
df_all = pd.concat((df_train, df_test), axis=0, ignore_index=True)
df_all = df_all.replace([np.inf, -np.inf], np.nan)
df_all['Age'].fillna(df_all['Age'].mean(), inplace=True)
df_all['Fare'] = df_all['Fare'].fillna(df_all['Fare'].mean())
df_all['has_cabin'] = df_all['Cabin'].apply(lambda val: 0 if pd.isnull(val) else 1)
df_all.shape, df_test.shape
df_all.columns
df_all = df_all[['Age', 'SibSp', 'Parch', 'Survived', 'Embarked', 'Pclass', 'Sex',
'Fare', 'has_cabin', 'PassengerId']]
df_all.set_index('PassengerId', inplace=True)
df_all['Sex'] = df_all['Sex'].map({'male':0, 'female':1})
df_all = pd.concat([df_all, pd.get_dummies(df_all['Embarked'])], axis=1)
df_all.drop('Embarked', axis=1, inplace=True)
df_train = df_all[:df_train.shape[0]]
df_test = df_all[df_train.shape[0]:]
df_train.shape, df_test.shape
df_train.head(5)
X_train, X_test = train_test_split(df_train, test_size=0.3)
Y_train = X_train.pop('Survived')
Y_test = X_test.pop('Survived')
## build a classifier model
from sklearn.ensemble import RandomForestClassifier
import matplotlib.pyplot as plt
% matplotlib inline
lw = 1 # line width for plt
model = RandomForestClassifier()
model = model.fit(X_train, Y_train)
# predict_proba will give probability for each class 0, 1.
# We only need valeus for calss 1
y_pred_test = model.predict_proba(X_test)[:,1]
```
### Section 2: We will Compute the auc and roc_auc_score
```
# we will compute ROC and Auc_ROC values
# we are setting thresholds as 0 to 1 with an increment of 0.05
# 1.05 has been taken as an outside of the boundary threshold
thresholds = list(reversed([i/100 for i in range(0, 106, 5)]))
total_positive = len(Y_test[Y_test == 1])
total_negative = len(Y_test[Y_test == 0])
total_positive, total_negative # total number of
# The idea is to find out at each threshold value what percentage of true positive and false positive
# cases are detected
# Ideally we want a threshold for which all all true positive cases are detected and no true negative
# cases are detected
tpr_computed = []
fpr_computed = []
for t_val in thresholds:
values = y_pred_test >= t_val
positive_count = sum(Y_test[values] == 1)
negative_count = len(Y_test[values]) - sum(Y_test[values] == 1)
tpr_computed.append(positive_count/total_positive)
fpr_computed.append(negative_count/total_negative)
prev_fpr_value = 0
prev_tpr_value = 0
roc_auc_computed = 0
for fpr, tpr in zip(fpr_computed, tpr_computed):
# for each change in fpr there is a change in tpr
# this will form a rectangle and we will compute the area of the rectangle.
# plus the top part of the rectangle is sort of like a traiangle we will adjust for that.
fpr_change = (fpr - prev_fpr_value)
tpr_change = tpr - prev_tpr_value
roc_auc_computed += prev_tpr_value * fpr_change + 1/2 * fpr_change * tpr_change
prev_fpr_value = fpr
prev_tpr_value = tpr
roc_auc_computed # the auc that we have computed
plt.plot(fpr_computed, tpr_computed, color='darkorange',
lw=lw, label='ROC curve (area = %0.2f)' % roc_auc_computed)
plt.plot([0, 1], [0, 1], color='navy', lw=lw, linestyle='--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic')
plt.legend(loc="lower right")
plt.show()
```
### Section 3: We will Compute the auc and roc_auc_score using sklearn for validation
```
# Let's compute the metrics using sklearn library
from sklearn.metrics import roc_curve, auc, roc_auc_score
roc_auc_score(Y_test, y_pred_test)
fpr, tpr, _ = roc_curve(Y_test, y_pred_test)
roc_auc = auc(fpr, tpr)
plt.plot(fpr, tpr, color='darkgreen',
lw=lw, label='ROC curve (area = %0.2f)' % roc_auc)
plt.plot([0, 1], [0, 1], color='navy', lw=lw, linestyle='--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic')
plt.legend(loc="lower right")
plt.show()
```
### Section 4: Plot both the metrics together for comparision
```
plt.plot(fpr_computed, tpr_computed, color='darkorange',
lw=lw, label='ROC curve (area = %0.2f)' % roc_auc_computed)
plt.plot(fpr, tpr, color='darkgreen',
lw=lw, label='ROC curve (area = %0.2f)' % roc_auc)
plt.plot([0, 1], [0, 1], color='navy', lw=lw, linestyle='--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic')
plt.legend(loc="lower right")
plt.show()
# Almost overlaps
print("Threshold | TPR | FPR")
for a, b, c in zip(tpr_computed, fpr_computed, thresholds):
print(" {0:.2f} |".format(c), "{0:.2f} |".format(a), "{0:.2f}".format(b))
```
<hr>
<pre>
Q> Now say we wan't to find out at what threshold value do we get 60 % true postive case and what %
of negative cases will come along with it?
A> For a threshold value of 0.75 we get close to 59% of true positive cases and 0.07 % of false positive case.
PS: To get exactly 60% we will have to go granular between 0.70 and 0.75 threshold.
</pre>
| github_jupyter |
... ***CURRENTLY UNDER DEVELOPMENT*** ...
## Synthetic simulation of historical TCs parameters using Gaussian copulas (Rueda et al. 2016) and subsequent selection of representative cases using Maximum Dissimilarity (MaxDiss) algorithm (Camus et al. 2011)
inputs required:
* Historical TC parameters that affect the site (output of *notebook 05*)
* number of synthetic simulations to run
* number of representative cases to be selected using MaxDiss
in this notebook:
* synthetic generation of TCs tracks based on gaussian copulas of the TC parameters
* MDA selection of representative number of events
```
#!/usr/bin/env python
# -*- coding: utf-8 -*-
# common
import os
import os.path as op
# pip
import xarray as xr
import numpy as np
# DEV: override installed teslakit
import sys
sys.path.insert(0, op.join(os.path.abspath(''), '..', '..', '..'))
# teslakit
from teslakit.database import Database
from teslakit.statistical import CopulaSimulation
from teslakit.mda import MaxDiss_Simplified_NoThreshold
from teslakit.plotting.storms import Plot_TCs_Params_MDAvsSIM, \
Plot_TCs_Params_HISTvsSIM, Plot_TCs_Params_HISTvsSIM_histogram
```
## Database and Site parameters
```
# --------------------------------------
# Teslakit database
p_data = r'C:\Users\lcag075\Dropbox\MAJURO-teslakit\teslakit\DATA'
db = Database(p_data)
# set site
db.SetSite('MAJURO')
# --------------------------------------
# load data and set parameters
_, TCs_r2_params = db.Load_TCs_r2_hist() # TCs parameters inside radius 2
# TCs random generation and MDA parameters
num_sim_rnd = 100000
num_sel_mda = 1000
```
## Historical TCs - Probabilistic Simulation
```
# --------------------------------------
# Probabilistic simulation Historical TCs
# aux functions
def adjust_to_pareto(var):
'Fix data. It needs to start at 0 for Pareto adjustment '
var = var.astype(float)
var_pareto = np.amax(var) - var + 0.00001
return var_pareto
def adjust_from_pareto(var_base, var_pareto):
'Returns data from pareto adjustment'
var = np.amax(var_base) - var_pareto + 0.00001
return var
# use small radius parameters (4º)
pmean = TCs_r2_params.pressure_mean.values[:]
pmin = TCs_r2_params.pressure_min.values[:]
gamma = TCs_r2_params.gamma.values[:]
delta = TCs_r2_params.delta.values[:]
vmean = TCs_r2_params.velocity_mean.values[:]
# fix pressure for p
pmean_p = adjust_to_pareto(pmean)
pmin_p = adjust_to_pareto(pmin)
# join storm parameters for copula simulation
storm_params = np.column_stack(
(pmean_p, pmin_p, gamma, delta, vmean)
)
# statistical simulate PCs using copulas
kernels = ['GPareto', 'GPareto', 'ECDF', 'ECDF', 'ECDF']
storm_params_sim = CopulaSimulation(storm_params, kernels, num_sim_rnd)
# adjust back pressures from pareto
pmean_sim = adjust_from_pareto(pmean, storm_params_sim[:,0])
pmin_sim = adjust_from_pareto(pmin, storm_params_sim[:,1])
# store simulated storms - parameters
TCs_r2_sim_params = xr.Dataset(
{
'pressure_mean':(('storm'), pmean_sim),
'pressure_min':(('storm'), pmin_sim),
'gamma':(('storm'), storm_params_sim[:,2]),
'delta':(('storm'), storm_params_sim[:,3]),
'velocity_mean':(('storm'), storm_params_sim[:,4]),
},
coords = {
'storm':(('storm'), np.arange(num_sim_rnd))
},
)
print(TCs_r2_sim_params)
db.Save_TCs_r2_sim_params(TCs_r2_sim_params)
# Historical vs Simulated: scatter plot parameters
Plot_TCs_Params_HISTvsSIM(TCs_r2_params, TCs_r2_sim_params);
# Historical vs Simulated: histogram parameters
Plot_TCs_Params_HISTvsSIM_histogram(TCs_r2_params, TCs_r2_sim_params);
```
## Simulated TCs - MaxDiss classification
```
# --------------------------------------
# MaxDiss classification
# get simulated parameters
pmean_s = TCs_r2_sim_params.pressure_mean.values[:]
pmin_s = TCs_r2_sim_params.pressure_min.values[:]
gamma_s = TCs_r2_sim_params.gamma.values[:]
delta_s = TCs_r2_sim_params.delta.values[:]
vmean_s = TCs_r2_sim_params.velocity_mean.values[:]
# subset, scalar and directional indexes
data_mda = np.column_stack((pmean_s, pmin_s, vmean_s, delta_s, gamma_s))
ix_scalar = [0,1,2]
ix_directional = [3,4]
centroids = MaxDiss_Simplified_NoThreshold(
data_mda, num_sel_mda, ix_scalar, ix_directional
)
# store MDA storms - parameters
TCs_r2_MDA_params = xr.Dataset(
{
'pressure_mean':(('storm'), centroids[:,0]),
'pressure_min':(('storm'), centroids[:,1]),
'velocity_mean':(('storm'), centroids[:,2]),
'delta':(('storm'), centroids[:,3]),
'gamma':(('storm'), centroids[:,4]),
},
coords = {
'storm':(('storm'), np.arange(num_sel_mda))
},
)
print(TCs_r2_MDA_params)
#db.Save_TCs_r2_mda_params(TCs_r2_MDA_params)
# Historical vs Simulated: scatter plot parameters
Plot_TCs_Params_MDAvsSIM(TCs_r2_MDA_params, TCs_r2_sim_params);
```
## Historical TCs (MDA centroids) Waves Simulation
Waves data is generated by numerically simulating selected storms.
This methodology is not included inside teslakit python library.
This step needs to be done before continuing with notebook 07
| github_jupyter |
```
# ------------------------------------------define logging and working directory
from ProjectRoot import change_wd_to_project_root
change_wd_to_project_root()
from src.utils.Tensorflow_helper import choose_gpu_by_id
# ------------------------------------------define GPU id/s to use, if given
GPU_IDS = '0,1'
GPUS = choose_gpu_by_id(GPU_IDS)
print(GPUS)
# ------------------------------------------jupyter magic config
%matplotlib inline
%reload_ext autoreload
%autoreload 2
# ------------------------------------------ import helpers
# this should import glob, os, and many other standard libs
# local imports
from src.utils.Notebook_imports import *
from src.utils.Utils_io import Console_and_file_logger, init_config
# import external libs
from tensorflow.python.client import device_lib
import tensorflow as tf
tf.get_logger().setLevel('ERROR')
import cv2
import pandas as pd
EXPERIMENT = 'cv_baseline_test/sub-identifier'
timestemp = str(datetime.datetime.now().strftime("%Y-%m-%d_%H_%M")) # ad a timestep to each project to make repeated experiments unique
EXPERIMENTS_ROOT = 'exp/'
EXP_PATH = os.path.join(EXPERIMENTS_ROOT, EXPERIMENT, timestemp)
MODEL_PATH = os.path.join(EXP_PATH, 'model', )
TENSORBOARD_PATH = os.path.join(EXP_PATH, 'tensorboard_logs')
CONFIG_PATH = os.path.join(EXP_PATH,'config')
HISTORY_PATH = os.path.join(EXP_PATH, 'history')
# define the input data paths and fold
# first to the 4D Nrrd files,
# second to a dataframe with a mapping of the Fold-number
# Finally the path to the metadata
DATA_PATH_SAX = 'path_to_cmr_images'
DF_FOLDS = 'path_to_patient_fold_mapping'
DF_META = 'path_to_df_with_patient_metadata'
FOLD = 0
# General params
SEED = 42 # define a seed for the generator shuffle
BATCHSIZE = 8 # 32, 64, 24, 16, 1 depends on the input shape and available memory
GENERATOR_WORKER = BATCHSIZE # if not set, use batchsize
EPOCHS = 50 # define the max number of epochs to train on
DIM = [224, 224] # network input shape for spacing of 3, (z,y,x)
SPACING = [1.5,1.5] # if resample, resample to this spacing, (z,y,x)
# Model params
DEPTH = 4 # depth of the encoder/decoder architecture
FILTERS = 32 # initial number of filters, will be doubled after each downsampling block
M_POOL = [2, 2]# size of max-pooling used for downsampling and upsampling
F_SIZE = [3, 3] # conv filter size
BN_FIRST = False # decide if batch normalisation between conv and activation or afterwards
BATCH_NORMALISATION = True # apply BN or not
PAD = 'same' # padding strategy of the conv layers
KERNEL_INIT = 'he_normal' # conv weight initialisation
OPTIMIZER = 'adam' # Adam, Adagrad, RMSprop, Adadelta, # https://keras.io/optimizers/
ACTIVATION = 'relu' # tf.keras.layers.LeakyReLU(), relu or any other non linear activation function
LEARNING_RATE = 1e-4 # start with a huge lr to converge fast
REDUCE_LR_ON_PLAEAU_PATIENCE = 5
DECAY_FACTOR = 0.7 # Define a learning rate decay for the ReduceLROnPlateau callback
MIN_LR = 1e-12 # minimal lr, smaller lr does not improve the model
DROPOUT_min = 0.3 # lower dropout at the shallow layers
DROPOUT_max = 0.5 # higher dropout at the deep layers
# Callback params
MONITOR_FUNCTION = 'loss'
MONITOR_MODE = 'min'
SAVE_MODEL_FUNCTION = 'loss'
SAVE_MODEL_MODE = 'min'
MODEL_PATIENCE = 20
SAVE_LEARNING_PROGRESS_AS_TF = True
# Generator and Augmentation params
BORDER_MODE = cv2.BORDER_REFLECT_101 # border mode for the data generation
IMG_INTERPOLATION = cv2.INTER_LINEAR # image interpolation in the genarator
MSK_INTERPOLATION = cv2.INTER_NEAREST # mask interpolation in the generator
AUGMENT = True # a compose of 2D augmentation (grid distortion, 90degree rotation, brightness and shift)
AUGMENT_PROB = 0.8
SHUFFLE = True
RESAMPLE = True
SCALER = 'MinMax' # MinMax, Standard or Robust
config = init_config(config=locals(), save=True)
from src.models.train_model import train_fold
folds = [i for i in range(0, 4, 1)]
for f in folds:
info('starting fold: {}'.format(f))
config_ = config.copy()
config_['FOLD'] = f
train_fold(config_)
```
| github_jupyter |
```
"""
The MIT License (MIT)
Copyright (c) 2021 NVIDIA
Permission is hereby granted, free of charge, to any person obtaining a copy of
this software and associated documentation files (the "Software"), to deal in
the Software without restriction, including without limitation the rights to
use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of
the Software, and to permit persons to whom the Software is furnished to do so,
subject to the following conditions:
The above copyright notice and this permission notice shall be included in all
copies or substantial portions of the Software.
THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS
FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR
COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER
IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN
CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE.
"""
```
This code example implements back-propagation algorithm for a two-level network and demonstrates how to use it to learn the exclusive OR (XOR) function. The network has two neurons in the hidden layer and a single output neuron. More context for this code example can be found in the section "Programming Example: Learning the XOR Function" in Chapter 3 in the book Learning Deep Learning by Magnus Ekman (ISBN: 9780137470358).
The initialization code in the code snippet below is similar to what we did for the perceptron example c1e1_perceptron_learning. One thing to note is that we have started to use NumPy arrays so that we can make use of some NumPy functionality. The same holds for our random number generator (we call np.random.seed instead of just random.seed).
For the training examples, we have now changed the ground truth to be between 0.0 and 1.0 because we have decided to use the logistic sigmoid function as an activation function for the output neuron, and its output range does not go to −1.0 as the perceptron did.
```
import numpy as np
np.random.seed(3) # To make repeatable
LEARNING_RATE = 0.1
index_list = [0, 1, 2, 3] # Used to randomize order
# Define training examples.
x_train = [np.array([1.0, -1.0, -1.0]),
np.array([1.0, -1.0, 1.0]),
np.array([1.0, 1.0, -1.0]),
np.array([1.0, 1.0, 1.0])]
y_train = [0.0, 1.0, 1.0, 0.0] # Output (ground truth)
```
In the next code snippet, we declare variables to hold the state of our three neurons. A real implementation would typically be parameterized to be able to choose number of inputs, layers, and number of neurons in each layer, but all of those parameters are hardcoded in this example to focus on readability.
```
def neuron_w(input_count):
weights = np.zeros(input_count+1)
for i in range(1, (input_count+1)):
weights[i] = np.random.uniform(-1.0, 1.0)
return weights
n_w = [neuron_w(2), neuron_w(2), neuron_w(2)]
n_y = [0, 0, 0]
n_error = [0, 0, 0]
```
The next code snippet starts with a function to print all the nine weights of the network (each print statement prints a three-element weight vector). The forward_pass function first computes the outputs of neurons 0 and 1 with the same inputs (the inputs from the training example) and then puts their outputs into an array, together with a bias value of 1.0, to use as input to neuron 2. That is, this function defines the topology of the network. We use tanh for the neurons in the first layer and the logistic sigmoid function for the output neuron.
The backward_pass function starts by computing the derivative of the error function and then computes the derivative of the activation function for the output neuron. The error term of the output neuron is computed by multiplying these two together. We then continue to backpropagate the error to each of the two neurons in the hidden layer. This is done by computing the derivatives of their activation functions and multiplying these derivatives by the error term from the output neuron and by the weight to the output neuron.
Finally, the adjust_weights function adjusts the weights for each of the three neurons. The adjustment factor is computed by multiplying the input by the learning rate and the error term for the neuron in question.
```
def show_learning():
print('Current weights:')
for i, w in enumerate(n_w):
print('neuron ', i, ': w0 =', '%5.2f' % w[0],
', w1 =', '%5.2f' % w[1], ', w2 =',
'%5.2f' % w[2])
print('----------------')
def forward_pass(x):
global n_y
n_y[0] = np.tanh(np.dot(n_w[0], x)) # Neuron 0
n_y[1] = np.tanh(np.dot(n_w[1], x)) # Neuron 1
n2_inputs = np.array([1.0, n_y[0], n_y[1]]) # 1.0 is bias
z2 = np.dot(n_w[2], n2_inputs)
n_y[2] = 1.0 / (1.0 + np.exp(-z2))
def backward_pass(y_truth):
global n_error
error_prime = -(y_truth - n_y[2]) # Derivative of loss-func
derivative = n_y[2] * (1.0 - n_y[2]) # Logistic derivative
n_error[2] = error_prime * derivative
derivative = 1.0 - n_y[0]**2 # tanh derivative
n_error[0] = n_w[2][1] * n_error[2] * derivative
derivative = 1.0 - n_y[1]**2 # tanh derivative
n_error[1] = n_w[2][2] * n_error[2] * derivative
def adjust_weights(x):
global n_w
n_w[0] -= (x * LEARNING_RATE * n_error[0])
n_w[1] -= (x * LEARNING_RATE * n_error[1])
n2_inputs = np.array([1.0, n_y[0], n_y[1]]) # 1.0 is bias
n_w[2] -= (n2_inputs * LEARNING_RATE * n_error[2])
```
With all these pieces in place, the only remaining piece is the training loop shown in the code snippet below, which is somewhat similar to the training loop for the perceptron example in c1e1_perceptron_learning.
We pick training examples in random order, call the functions forward_pass, backward_pass, and adjust_weights, and then print out the weights with the function show_learning. We adjust the weights regardless whether the network predicts correctly or not. Once we have looped through all four training examples, we check whether the network can predict them all correctly, and if not, we do another pass over them in random order.
```
# Network training loop.
all_correct = False
while not all_correct: # Train until converged
all_correct = True
np.random.shuffle(index_list) # Randomize order
for i in index_list: # Train on all examples
forward_pass(x_train[i])
backward_pass(y_train[i])
adjust_weights(x_train[i])
show_learning() # Show updated weights
for i in range(len(x_train)): # Check if converged
forward_pass(x_train[i])
print('x1 =', '%4.1f' % x_train[i][1], ', x2 =',
'%4.1f' % x_train[i][2], ', y =',
'%.4f' % n_y[2])
if(((y_train[i] < 0.5) and (n_y[2] >= 0.5))
or ((y_train[i] >= 0.5) and (n_y[2] < 0.5))):
all_correct = False
```
| github_jupyter |
# Convolutional Neural Networks with Keras
***

***
__Keras is a high level machine learning API built using Python. The library is is running on top of Tensorflow, Theano or CNTK which allows you the choice of what framework backend you want to run your model with.__
Throughout each of the tutorials on convolutional neural networks I will be referencing the associated theory document so that you can get a good understanding of what is going on in the background. So, please make sure to read the theory that I reference if you are struggling to understand what is going on.
***
```
# Imports
import numpy as np
import cv2 as cv
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import os
from tqdm import tqdm
from sklearn.model_selection import train_test_split
from keras.models import Sequential
from keras.layers import Activation, Convolution2D, Dense, Dropout, Flatten, MaxPooling2D
from keras.optimizers import Adam, SGD, rmsprop
from keras.preprocessing.image import ImageDataGenerator, array_to_img, img_to_array, load_img
```
***
## Load In The Data
You will need to download the data from [Kaggle](https://www.kaggle.com/c/dogs-vs-cats/data). The data has been organised into a training and testing folder. Before we can get to work with the data it needs to be organised into folders representing all of the cats and all of the dogs. It's very important that you are able to understand the data that you are working with before building and training the model.
***
In Jupyter Notebooks you can interact directly with the console by using the "!" syntax before your command. It is useful to make use of this to interact with your data directly and find the location of specific files without having to leave the environment.
```
!ls ../../../data/all
```
When working with file paths a lot, it is good to define them beforehand. We have done that here; a general path, which directs you to all of you downloaded data and a train/test path that points directly at our image data that we want to use. Now we just have to call their set variable names.
```
PATH = "../../../all/"
TRAIN_PATH = "../../../data/all/train/"
TEST_PATH = "../../../data/all/test1/"
```
Here we set the image height and width that we want all of our images to be. Decreasing the size of our images before we feed them through the model decreases the time it takes to train as there is less data to process.
```python
IMG_HEIGHT = 256
IMG_WIDTH = 256
```
We then create two arrays which will store our images and their associated class or label, which is found by looking at the file name.
```python
images = []
classes = []
```
The for loop goes through every image in the training folder and loads each image in individually, appending each to the "images" array. Here we can use the height and width that we set. We then check the filename of the image file that we just loaded in for either cat or dog to set the according number.
```python
for filename in tqdm(os.listdir(TRAIN_2_PATH)):
img = mpimg.imread(TRAIN_2_PATH+filename)
images.append(cv.resize(img, (IMG_HEIGHT, IMG_WIDTH)))
if "cat" in filename:
classes.append(0)
elif "dog" in filename:
classes.append(1)
```
```
IMG_HEIGHT = 256
IMG_WIDTH = 256
images = []
classes = []
for filename in tqdm(os.listdir(TRAIN_PATH)):
img = mpimg.imread(TRAIN_PATH+filename)
images.append(cv.resize(img, (IMG_HEIGHT, IMG_WIDTH)))
if "cat" in filename:
classes.append(0)
elif "dog" in filename:
classes.append(1)
```
To make sure that we have loaded our images in correctly and our labels for the associated image are correct we can plot the images with their label. We will do this with matplotlib and plot the first six images with their labels as the title. Make sure that the labels are correctly matching the image, 1 for dogs and 0 for cats.
```
COL = 3
ROW = 2
fig = plt.figure(figsize=(8, 8))
for i in range(0, COL*ROW):
fig.add_subplot(ROW, COL, i+1) #IF YOU DON'T HAVE +1, IT FALLS OUT OF RANGE
plt.title(str(classes[i]))
plt.imshow(images[i])
plt.show()
```
## Augment The Data
It is important that we augment (manipulate) our data to generalise and avoid overfitting. Overfitting happens when our model becomes to acustomed to the features of our training data and so performs worse when validating our performance. By augmenting our data we are generating a set amount of variations for each image that we pass into the `ImageDataGenerator()`.
***
Set X equal to the "images" array wrapped in another numpy array; we need to do this because the model is expecting a 4 dimensional array. The array it is expecting should look like, `(Total Images, Img Height, Img Width, Img Depth)` when using the `.shape()` function on the array. Y is set equal to the "classes" array as the classes array needs to be in an array of the same dimensions as X.
```
X = np.array(images, dtype=np.float32)
Y = np.array(classes, dtype=np.uint8)
```
Using the scikit-learn library which has a lot of useful functions that we can make use of, we are going to use the `train_test_split` function in particular. The train_test_split function allows us to take our total data to create a training set and a validation set. The training and validation data allows us to see how the model is performing while it is training.
```
x_train, x_val, y_train, y_val = train_test_split(X, Y, test_size=0.2)
```
The `ImageDataGenerator()` object gives us the ability to generate variations of our images so that we can generalise our model by introducing more variations on the images that it is looking at and training on. When setting the parameters of the `ImageDataGenerator`, we are selecting the range of how much we want to affect the generated variations of the selected image.
```
train_datagen = ImageDataGenerator(rescale=1. / 255,
rotation_range=40,
#width_shift_range=60,
#height_shift_range=30,
#brightness_range=10,
shear_range=10.0,
#zoom_range=5.0,
horizontal_flip=True)
valid_datagen = ImageDataGenerator(rescale=1. / 255)
```
The `.flow()` function is used to generate batches of data using the images and labels. Here we will be generating batch sizes of 125 images and labels. `shuffle` when equal to `True` shuffles the data so that it is not in the set order that it has been downloaded in.
```
train_generator = train_datagen.flow(x_train, y_train, batch_size=125, shuffle=True)
valid_generator = valid_datagen.flow(x_val, y_val, batch_size=125)
```
## Build The Model
***

***
Now that we have collected our data, modified the images and augmented them to increase the reliability of our model. We need to build the model that we will be training on the data that we have prepared. We will build a `Sequential()` model as it is capable for most problems, the alternative is a functional model which allows for a lot more flexibility as you can connect to more than just the previous and next layers.
```
model = Sequential()
```
The `Convolution2D` layer extracts the features from the data passed in. Imagine we have the 5x5 image (green) and that it is a special case where the the pixels are only 1 or 0. We will take the 3x3 matrix and slide it over the original image by 1 pixel at a time (this is our stride) and multiply the elements and add them to get a final value for the feature map.

ReLU (Rectified Linear Unit) steps are normally introduced after convolution operations in CNN’s. ReLU is applied per pixel and replaces any negative values with a 0 value.

The `MaxPooling2D` layer reduces the dimensionality of the feature maps while retaining the important information. We perform pooling to redact the spatial size of the input, this means pooling makes the input representations smaller, reduces network computations which reduces overfitting and makes the network invariant to small transformations or distortions.

```
model.add(Convolution2D(128, (5, 5), input_shape=(IMG_HEIGHT, IMG_WIDTH, 3)))
model.add(Activation("relu"))
model.add(MaxPooling2D(pool_size=(2, 2), data_format="channels_first"))
model.add(Convolution2D(128, (5, 5)))
model.add(Activation("relu"))
model.add(MaxPooling2D(pool_size=(2, 2), data_format="channels_first"))
model.add(Convolution2D(64, (3, 3)))
model.add(Activation("relu"))
model.add(MaxPooling2D(pool_size=(2, 2), data_format="channels_first"))
```
The cell below is what is known as a fully connected layer. The fully connected layer implies that every neutron in the previous layer is connected to every neutron in the next layer. The most popular activation function in the output layer is Softmax however other classifiers such as SVM can be used.

The concolution layers above represent the high level features of the data, while adding a fully connected layer is a cheap method of learning a non-linear combination of these features.
```
model.add(Flatten())
model.add(Dense(256))
model.add(Activation("relu"))
model.add(Dense(256))
model.add(Dense(1))
model.add(Activation("sigmoid"))
model.compile(optimizer="adam", metrics=["accuracy"], loss="binary_crossentropy")
model.fit_generator(train_generator,
steps_per_epoch=20000 // 125,
epochs=1,
validation_data=valid_generator)
```
| github_jupyter |
# Deep Neural Network for Image Classification: Application
When you finish this, you will have finished the last programming assignment of Week 4, and also the last programming assignment of this course!
You will use use the functions you'd implemented in the previous assignment to build a deep network, and apply it to cat vs non-cat classification. Hopefully, you will see an improvement in accuracy relative to your previous logistic regression implementation.
**After this assignment you will be able to:**
- Build and apply a deep neural network to supervised learning.
Let's get started!
## 1 - Packages
Let's first import all the packages that you will need during this assignment.
- [numpy](www.numpy.org) is the fundamental package for scientific computing with Python.
- [matplotlib](http://matplotlib.org) is a library to plot graphs in Python.
- [h5py](http://www.h5py.org) is a common package to interact with a dataset that is stored on an H5 file.
- [PIL](http://www.pythonware.com/products/pil/) and [scipy](https://www.scipy.org/) are used here to test your model with your own picture at the end.
- dnn_app_utils provides the functions implemented in the "Building your Deep Neural Network: Step by Step" assignment to this notebook.
- np.random.seed(1) is used to keep all the random function calls consistent. It will help us grade your work.
```
import time
import numpy as np
import h5py
import matplotlib.pyplot as plt
import scipy
from PIL import Image
from scipy import ndimage
from dnn_app_utils_v2 import *
%matplotlib inline
plt.rcParams['figure.figsize'] = (5.0, 4.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
%load_ext autoreload
%autoreload 2
np.random.seed(1)
```
## 2 - Dataset
You will use the same "Cat vs non-Cat" dataset as in "Logistic Regression as a Neural Network" (Assignment 2). The model you had built had 70% test accuracy on classifying cats vs non-cats images. Hopefully, your new model will perform a better!
**Problem Statement**: You are given a dataset ("data.h5") containing:
- a training set of m_train images labelled as cat (1) or non-cat (0)
- a test set of m_test images labelled as cat and non-cat
- each image is of shape (num_px, num_px, 3) where 3 is for the 3 channels (RGB).
Let's get more familiar with the dataset. Load the data by running the cell below.
```
train_x_orig, train_y, test_x_orig, test_y, classes = load_data()
```
The following code will show you an image in the dataset. Feel free to change the index and re-run the cell multiple times to see other images.
```
# Example of a picture
index = 10
plt.imshow(train_x_orig[index])
print ("y = " + str(train_y[0,index]) + ". It's a " + classes[train_y[0,index]].decode("utf-8") + " picture.")
# Explore your dataset
m_train = train_x_orig.shape[0]
num_px = train_x_orig.shape[1]
m_test = test_x_orig.shape[0]
print ("Number of training examples: " + str(m_train))
print ("Number of testing examples: " + str(m_test))
print ("Each image is of size: (" + str(num_px) + ", " + str(num_px) + ", 3)")
print ("train_x_orig shape: " + str(train_x_orig.shape))
print ("train_y shape: " + str(train_y.shape))
print ("test_x_orig shape: " + str(test_x_orig.shape))
print ("test_y shape: " + str(test_y.shape))
```
As usual, you reshape and standardize the images before feeding them to the network. The code is given in the cell below.
<img src="images/imvectorkiank.png" style="width:450px;height:300px;">
<caption><center> <u>Figure 1</u>: Image to vector conversion. <br> </center></caption>
```
# Reshape the training and test examples
train_x_flatten = train_x_orig.reshape(train_x_orig.shape[0], -1).T # The "-1" makes reshape flatten the remaining dimensions
test_x_flatten = test_x_orig.reshape(test_x_orig.shape[0], -1).T
# Standardize data to have feature values between 0 and 1.
train_x = train_x_flatten/255.
test_x = test_x_flatten/255.
print ("train_x's shape: " + str(train_x.shape))
print ("test_x's shape: " + str(test_x.shape))
```
$12,288$ equals $64 \times 64 \times 3$ which is the size of one reshaped image vector.
## 3 - Architecture of your model
Now that you are familiar with the dataset, it is time to build a deep neural network to distinguish cat images from non-cat images.
You will build two different models:
- A 2-layer neural network
- An L-layer deep neural network
You will then compare the performance of these models, and also try out different values for $L$.
Let's look at the two architectures.
### 3.1 - 2-layer neural network
<img src="images/2layerNN_kiank.png" style="width:650px;height:400px;">
<caption><center> <u>Figure 2</u>: 2-layer neural network. <br> The model can be summarized as: ***INPUT -> LINEAR -> RELU -> LINEAR -> SIGMOID -> OUTPUT***. </center></caption>
<u>Detailed Architecture of figure 2</u>:
- The input is a (64,64,3) image which is flattened to a vector of size $(12288,1)$.
- The corresponding vector: $[x_0,x_1,...,x_{12287}]^T$ is then multiplied by the weight matrix $W^{[1]}$ of size $(n^{[1]}, 12288)$.
- You then add a bias term and take its relu to get the following vector: $[a_0^{[1]}, a_1^{[1]},..., a_{n^{[1]}-1}^{[1]}]^T$.
- You then repeat the same process.
- You multiply the resulting vector by $W^{[2]}$ and add your intercept (bias).
- Finally, you take the sigmoid of the result. If it is greater than 0.5, you classify it to be a cat.
### 3.2 - L-layer deep neural network
It is hard to represent an L-layer deep neural network with the above representation. However, here is a simplified network representation:
<img src="images/LlayerNN_kiank.png" style="width:650px;height:400px;">
<caption><center> <u>Figure 3</u>: L-layer neural network. <br> The model can be summarized as: ***[LINEAR -> RELU] $\times$ (L-1) -> LINEAR -> SIGMOID***</center></caption>
<u>Detailed Architecture of figure 3</u>:
- The input is a (64,64,3) image which is flattened to a vector of size (12288,1).
- The corresponding vector: $[x_0,x_1,...,x_{12287}]^T$ is then multiplied by the weight matrix $W^{[1]}$ and then you add the intercept $b^{[1]}$. The result is called the linear unit.
- Next, you take the relu of the linear unit. This process could be repeated several times for each $(W^{[l]}, b^{[l]})$ depending on the model architecture.
- Finally, you take the sigmoid of the final linear unit. If it is greater than 0.5, you classify it to be a cat.
### 3.3 - General methodology
As usual you will follow the Deep Learning methodology to build the model:
1. Initialize parameters / Define hyperparameters
2. Loop for num_iterations:
a. Forward propagation
b. Compute cost function
c. Backward propagation
d. Update parameters (using parameters, and grads from backprop)
4. Use trained parameters to predict labels
Let's now implement those two models!
## 4 - Two-layer neural network
**Question**: Use the helper functions you have implemented in the previous assignment to build a 2-layer neural network with the following structure: *LINEAR -> RELU -> LINEAR -> SIGMOID*. The functions you may need and their inputs are:
```python
def initialize_parameters(n_x, n_h, n_y):
...
return parameters
def linear_activation_forward(A_prev, W, b, activation):
...
return A, cache
def compute_cost(AL, Y):
...
return cost
def linear_activation_backward(dA, cache, activation):
...
return dA_prev, dW, db
def update_parameters(parameters, grads, learning_rate):
...
return parameters
```
```
### CONSTANTS DEFINING THE MODEL ####
n_x = 12288 # num_px * num_px * 3
n_h = 7
n_y = 1
layers_dims = (n_x, n_h, n_y)
# GRADED FUNCTION: two_layer_model
def two_layer_model(X, Y, layers_dims, learning_rate = 0.0075, num_iterations = 3000, print_cost=False):
"""
Implements a two-layer neural network: LINEAR->RELU->LINEAR->SIGMOID.
Arguments:
X -- input data, of shape (n_x, number of examples)
Y -- true "label" vector (containing 0 if cat, 1 if non-cat), of shape (1, number of examples)
layers_dims -- dimensions of the layers (n_x, n_h, n_y)
num_iterations -- number of iterations of the optimization loop
learning_rate -- learning rate of the gradient descent update rule
print_cost -- If set to True, this will print the cost every 100 iterations
Returns:
parameters -- a dictionary containing W1, W2, b1, and b2
"""
np.random.seed(1)
grads = {}
costs = [] # to keep track of the cost
m = X.shape[1] # number of examples
(n_x, n_h, n_y) = layers_dims
# Initialize parameters dictionary, by calling one of the functions you'd previously implemented
### START CODE HERE ### (≈ 1 line of code)
parameters = None
### END CODE HERE ###
# Get W1, b1, W2 and b2 from the dictionary parameters.
W1 = parameters["W1"]
b1 = parameters["b1"]
W2 = parameters["W2"]
b2 = parameters["b2"]
# Loop (gradient descent)
for i in range(0, num_iterations):
# Forward propagation: LINEAR -> RELU -> LINEAR -> SIGMOID. Inputs: "X, W1, b1". Output: "A1, cache1, A2, cache2".
### START CODE HERE ### (≈ 2 lines of code)
A1, cache1 = None
A2, cache2 = None
### END CODE HERE ###
# Compute cost
### START CODE HERE ### (≈ 1 line of code)
cost = None
### END CODE HERE ###
# Initializing backward propagation
dA2 = - (np.divide(Y, A2) - np.divide(1 - Y, 1 - A2))
# Backward propagation. Inputs: "dA2, cache2, cache1". Outputs: "dA1, dW2, db2; also dA0 (not used), dW1, db1".
### START CODE HERE ### (≈ 2 lines of code)
dA1, dW2, db2 = None
dA0, dW1, db1 = None
### END CODE HERE ###
# Set grads['dWl'] to dW1, grads['db1'] to db1, grads['dW2'] to dW2, grads['db2'] to db2
grads['dW1'] = dW1
grads['db1'] = db1
grads['dW2'] = dW2
grads['db2'] = db2
# Update parameters.
### START CODE HERE ### (approx. 1 line of code)
parameters = None
### END CODE HERE ###
# Retrieve W1, b1, W2, b2 from parameters
W1 = parameters["W1"]
b1 = parameters["b1"]
W2 = parameters["W2"]
b2 = parameters["b2"]
# Print the cost every 100 training example
if print_cost and i % 100 == 0:
print("Cost after iteration {}: {}".format(i, np.squeeze(cost)))
if print_cost and i % 100 == 0:
costs.append(cost)
# plot the cost
plt.plot(np.squeeze(costs))
plt.ylabel('cost')
plt.xlabel('iterations (per tens)')
plt.title("Learning rate =" + str(learning_rate))
plt.show()
return parameters
```
Run the cell below to train your parameters. See if your model runs. The cost should be decreasing. It may take up to 5 minutes to run 2500 iterations. Check if the "Cost after iteration 0" matches the expected output below, if not click on the square (⬛) on the upper bar of the notebook to stop the cell and try to find your error.
```
parameters = two_layer_model(train_x, train_y, layers_dims = (n_x, n_h, n_y), num_iterations = 2500, print_cost=True)
```
**Expected Output**:
<table>
<tr>
<td> **Cost after iteration 0**</td>
<td> 0.6930497356599888 </td>
</tr>
<tr>
<td> **Cost after iteration 100**</td>
<td> 0.6464320953428849 </td>
</tr>
<tr>
<td> **...**</td>
<td> ... </td>
</tr>
<tr>
<td> **Cost after iteration 2400**</td>
<td> 0.048554785628770206 </td>
</tr>
</table>
Good thing you built a vectorized implementation! Otherwise it might have taken 10 times longer to train this.
Now, you can use the trained parameters to classify images from the dataset. To see your predictions on the training and test sets, run the cell below.
```
predictions_train = predict(train_x, train_y, parameters)
```
**Expected Output**:
<table>
<tr>
<td> **Accuracy**</td>
<td> 1.0 </td>
</tr>
</table>
```
predictions_test = predict(test_x, test_y, parameters)
```
**Expected Output**:
<table>
<tr>
<td> **Accuracy**</td>
<td> 0.72 </td>
</tr>
</table>
**Note**: You may notice that running the model on fewer iterations (say 1500) gives better accuracy on the test set. This is called "early stopping" and we will talk about it in the next course. Early stopping is a way to prevent overfitting.
Congratulations! It seems that your 2-layer neural network has better performance (72%) than the logistic regression implementation (70%, assignment week 2). Let's see if you can do even better with an $L$-layer model.
## 5 - L-layer Neural Network
**Question**: Use the helper functions you have implemented previously to build an $L$-layer neural network with the following structure: *[LINEAR -> RELU]$\times$(L-1) -> LINEAR -> SIGMOID*. The functions you may need and their inputs are:
```python
def initialize_parameters_deep(layers_dims):
...
return parameters
def L_model_forward(X, parameters):
...
return AL, caches
def compute_cost(AL, Y):
...
return cost
def L_model_backward(AL, Y, caches):
...
return grads
def update_parameters(parameters, grads, learning_rate):
...
return parameters
```
```
### CONSTANTS ###
layers_dims = [12288, 20, 7, 5, 1] # 4-layer model
# GRADED FUNCTION: L_layer_model
def L_layer_model(X, Y, layers_dims, learning_rate = 0.0075, num_iterations = 3000, print_cost=False):#lr was 0.009
"""
Implements a L-layer neural network: [LINEAR->RELU]*(L-1)->LINEAR->SIGMOID.
Arguments:
X -- data, numpy array of shape (number of examples, num_px * num_px * 3)
Y -- true "label" vector (containing 0 if cat, 1 if non-cat), of shape (1, number of examples)
layers_dims -- list containing the input size and each layer size, of length (number of layers + 1).
learning_rate -- learning rate of the gradient descent update rule
num_iterations -- number of iterations of the optimization loop
print_cost -- if True, it prints the cost every 100 steps
Returns:
parameters -- parameters learnt by the model. They can then be used to predict.
"""
np.random.seed(1)
costs = [] # keep track of cost
# Parameters initialization.
### START CODE HERE ###
parameters = None
### END CODE HERE ###
# Loop (gradient descent)
for i in range(0, num_iterations):
# Forward propagation: [LINEAR -> RELU]*(L-1) -> LINEAR -> SIGMOID.
### START CODE HERE ### (≈ 1 line of code)
AL, caches = None
### END CODE HERE ###
# Compute cost.
### START CODE HERE ### (≈ 1 line of code)
cost = None
### END CODE HERE ###
# Backward propagation.
### START CODE HERE ### (≈ 1 line of code)
grads = None
### END CODE HERE ###
# Update parameters.
### START CODE HERE ### (≈ 1 line of code)
parameters = None
### END CODE HERE ###
# Print the cost every 100 training example
if print_cost and i % 100 == 0:
print ("Cost after iteration %i: %f" %(i, cost))
if print_cost and i % 100 == 0:
costs.append(cost)
# plot the cost
plt.plot(np.squeeze(costs))
plt.ylabel('cost')
plt.xlabel('iterations (per tens)')
plt.title("Learning rate =" + str(learning_rate))
plt.show()
return parameters
```
You will now train the model as a 4-layer neural network.
Run the cell below to train your model. The cost should decrease on every iteration. It may take up to 5 minutes to run 2500 iterations. Check if the "Cost after iteration 0" matches the expected output below, if not click on the square (⬛) on the upper bar of the notebook to stop the cell and try to find your error.
```
parameters = L_layer_model(train_x, train_y, layers_dims, num_iterations = 2500, print_cost = True)
```
**Expected Output**:
<table>
<tr>
<td> **Cost after iteration 0**</td>
<td> 0.771749 </td>
</tr>
<tr>
<td> **Cost after iteration 100**</td>
<td> 0.672053 </td>
</tr>
<tr>
<td> **...**</td>
<td> ... </td>
</tr>
<tr>
<td> **Cost after iteration 2400**</td>
<td> 0.092878 </td>
</tr>
</table>
```
pred_train = predict(train_x, train_y, parameters)
```
<table>
<tr>
<td>
**Train Accuracy**
</td>
<td>
0.985645933014
</td>
</tr>
</table>
```
pred_test = predict(test_x, test_y, parameters)
```
**Expected Output**:
<table>
<tr>
<td> **Test Accuracy**</td>
<td> 0.8 </td>
</tr>
</table>
Congrats! It seems that your 4-layer neural network has better performance (80%) than your 2-layer neural network (72%) on the same test set.
This is good performance for this task. Nice job!
Though in the next course on "Improving deep neural networks" you will learn how to obtain even higher accuracy by systematically searching for better hyperparameters (learning_rate, layers_dims, num_iterations, and others you'll also learn in the next course).
## 6) Results Analysis
First, let's take a look at some images the L-layer model labeled incorrectly. This will show a few mislabeled images.
```
print_mislabeled_images(classes, test_x, test_y, pred_test)
```
**A few types of images the model tends to do poorly on include:**
- Cat body in an unusual position
- Cat appears against a background of a similar color
- Unusual cat color and species
- Camera Angle
- Brightness of the picture
- Scale variation (cat is very large or small in image)
## 7) Test with your own image (optional/ungraded exercise) ##
Congratulations on finishing this assignment. You can use your own image and see the output of your model. To do that:
1. Click on "File" in the upper bar of this notebook, then click "Open" to go on your Coursera Hub.
2. Add your image to this Jupyter Notebook's directory, in the "images" folder
3. Change your image's name in the following code
4. Run the code and check if the algorithm is right (1 = cat, 0 = non-cat)!
```
## START CODE HERE ##
my_image = "my_image.jpg" # change this to the name of your image file
my_label_y = [1] # the true class of your image (1 -> cat, 0 -> non-cat)
## END CODE HERE ##
fname = "images/" + my_image
image = np.array(ndimage.imread(fname, flatten=False))
my_image = scipy.misc.imresize(image, size=(num_px,num_px)).reshape((num_px*num_px*3,1))
my_predicted_image = predict(my_image, my_label_y, parameters)
plt.imshow(image)
print ("y = " + str(np.squeeze(my_predicted_image)) + ", your L-layer model predicts a \"" + classes[int(np.squeeze(my_predicted_image)),].decode("utf-8") + "\" picture.")
```
**References**:
- for auto-reloading external module: http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
| github_jupyter |
# Data Analysis using PySpark using EMR Cluster with SageMaker Studio
#### Dataset used: Synthetic California Housing Dataset with 1 million rows
### Topics Covered
- Setup
- Exploratory Data Analysis on 1M rows dataset
- Data Preprocessing
- Automating it using SageMaker Processing Job with PySpark
## Setup
Before we begin with exploratory data analysis, let's connect EMR Cluster with SageMaker Studio.
```
%load_ext sagemaker_studio_analytics_extension.magics
%sm_analytics emr connect --cluster-id j-IYQNCR220TIB --auth-type None
%%info
```
### Read Parquet file using sqlContext which comes pre-build with PySpark kernel
```
housing_data=sqlContext.read.parquet('s3://sagemaker-us-east-2-670488263423/california_housing/data/raw/data.parquet.gzip')
# Shape and schema of data
print((housing_data.count(), len(housing_data.columns)))
housing_data.printSchema()
```
## Remove Duplicates
Before we do exploratory data analysis, lets remove the duplicates if any, in the spark dataframe.
```
housing_data.dropDuplicates()
housing_data.count()
```
## Handle Missing Values
```
from pyspark.sql.functions import isnan, when, count, col
housing_data.select([count(when(isnan(c), c)).alias(c) for c in housing_data.columns]).show()
from pyspark.sql.functions import isnan, when, count, col
housing_data.select([count(when(col(c).isNull(), c)).alias(c) for c in housing_data.columns]).show()
```
## Exploratory Data Analysis
```
import matplotlib.pyplot as plt
def plot_median_house_value(median_house_value, x_label, title):
plt.figure(figsize=(16,7))
plt.hist(median_house_value, bins=20, edgecolor="black")
plt.xlabel(x_label, fontsize=14)
plt.ylabel("Number of Houses", fontsize=13)
plt.xticks(rotation=0)
plt.title(title, fontsize=18)
plt.show()
import pyspark.sql.functions as f
median_house_value = housing_data.select(f.collect_list('medianHouseValue')).first()[0]
```
### Plot distribution of median house value in California
```
len(median_house_value)
plot_median_house_value(median_house_value, "Median House Value", "Median House Value in California (CA)")
%matplot plt
type(housing_data)
```
### Plot distribution of houses which are greater than 40years of age in California
```
# plot distribution of houses which are greater than 40years of age.
median_housing_value_by_age = housing_data.select('medianHouseValue').filter(housing_data.medianHousingAge>= 50).collect()
len(median_housing_value_by_age)
# id_vals = [r['id'] for r in ID]
values = [r['medianHouseValue'] for r in median_housing_value_by_age]
plot_median_house_value(values, "Median House Value ", "Median House Value for houses greater than 50yrs in California (CA)")
%matplot plt
```
## Lets look at the Statistical properties of data
```
housing_data.describe().show()
```
## Distribution of all features
Based on the results if we focus on the mean and the range of values for `avgNumRooms`, `avgNumBedrooms` and `population`, it looks like that the data is not well distributed, lets plot the distribution of these features to do further analysis.
```
# convert the spark dataframe into pandas
df = housing_data.select('avgNumRooms', 'avgNumBedrooms', 'population').toPandas()
import matplotlib.pyplot as plt
df.hist(figsize=(10, 8), bins=20, edgecolor="black")
plt.subplots_adjust(hspace=0.3, wspace=0.5)
plt.show()
%matplot plt
```
Looks like the features `avgNumRooms`, `avgNumBedrooms`, `population` might have some outliers. Let's create a box plot to visually confirm if there are any outliers in theses columns.
```
def plot_boxplot(data, title):
plt.figure(figsize =(5, 4))
plt.boxplot(data)
plt.title(title)
plt.show()
# boxplot for avgNumRooms
plot_boxplot(df.avgNumRooms, 'Boxplot for Average Number of Rooms')
%matplot plt
plot_boxplot(df.avgNumBedrooms, 'Boxplot for Average Number of Bedrooms')
%matplot plt
plot_boxplot(df.population, 'Boxplot for Popluation')
%matplot plt
```
As we can see from the above graphs, there are lot of outliers in `avgNumRooms`, `avgNumBedrooms`, `population`, that we need to fix, before training the model. Therefore, in the next section, lets process these features.
## Data Pre-processing
Based on the boxplots for `avgNumRooms`, `avgNumBedrooms`, `population`, we need to process the data and remove the outliers, before we train any machine learning model. There are different approaches to do it, for example, you can calculate the Inter Quartile Range(IQR) and remove all the outliers which are 3 IQRs away. In our example, we will simply query the data based on the boxplot values, and remove the outliers outside of our boxplot as shown in the `Exploratory Data Analysis` section.
```
columns = ['avgNumRooms', 'avgNumBedrooms', 'population']
import pyspark.sql.functions as f
housing_df_with_no_outliers = housing_data.where((housing_data.avgNumRooms<= 8) &
(housing_data.avgNumRooms>=2) &
(housing_data.avgNumBedrooms<=1.12) &
(housing_data.population<=1500) &
(housing_data.population>=250))
print('# of rows in orginal dataset: ', housing_data.count() )
print('# of rows in new dataset with no outliers: ', housing_df_with_no_outliers.count() )
```
Let's verify our dataset with box plots to see if we have successfully removed the outlier values.
```
df = housing_df_with_no_outliers.select('avgNumRooms', 'avgNumBedrooms', 'population').toPandas()
# boxplot for avgNumRooms
plot_boxplot(df.avgNumRooms, 'Boxplot for Average Number of Rooms')
%matplot plt
# boxplot for avgNumRooms
plot_boxplot(df.avgNumBedrooms, 'Boxplot for Average Number of Bedrooms')
%matplot plt
plot_boxplot(df.population, 'Boxplot for Population')
%matplot plt
```
Now our data looks much cleaner, so let's wrap our data processing code into a python script to run it with SageMaker Processing Jobs.
## Automating data processing using SageMaker Processing Job with PySpark
To analyze data and evaluate machine learning models on Amazon SageMaker, use Amazon SageMaker Processing. With Processing, you can use a simplified, managed experience on SageMaker to run your data processing workloads, such as feature engineering, data validation, model evaluation, and model interpretation. You can also use the Amazon SageMaker Processing APIs during the experimentation phase and after the code is deployed in production to evaluate performance.
The processing container image can either be an Amazon SageMaker built-in image or a custom image that you provide. The underlying infrastructure for a Processing job is fully managed by Amazon SageMaker. Cluster resources are provisioned for the duration of your job, and cleaned up when a job completes. The output of the Processing job is stored in the Amazon S3 bucket you specified.
Note:
Your input data must be stored in an Amazon S3 bucket. Alternatively, you can use Amazon Athena or Amazon Redshift as input sources.
Reference: https://docs.aws.amazon.com/sagemaker/latest/dg/processing-job.html
SageMaker Processing job supports two built-in container images:
- scikit-learn Processing and
- distributed data preprocessing with Spark
### Setup
```
%local
import sagemaker
from time import gmtime, strftime
sagemaker_session = sagemaker.Session()
role = sagemaker.get_execution_role()
bucket = sagemaker_session.default_bucket()
timestamp = strftime("%Y-%m-%d-%H-%M-%S", gmtime())
prefix = "california_housing/data_" + timestamp
s3_input_prefix = "california_housing/data/raw"
# s3_input_preprocessed_prefix = prefix + "/data/preprocessed"
s3_output_prefix = prefix + "/data/spark/processed"
# model_prefix = prefix + "/model"
```
Let's create preprocess.py file to run with SageMaker Processing job.
```
%%writefile preprocess.py
from __future__ import print_function
from __future__ import unicode_literals
import time
import sys
import os
import shutil
import argparse
import pyspark
from pyspark.sql import SparkSession
import pyspark.sql.functions as f
def main():
parser = argparse.ArgumentParser(description="app inputs and outputs")
parser.add_argument("--bucket", type=str, help="s3 input bucket")
parser.add_argument("--s3_input_prefix", type=str, help="s3 input key prefix")
parser.add_argument("--s3_output_prefix", type=str, help="s3 output key prefix")
args = parser.parse_args()
spark = SparkSession.builder.appName("PySparkApp").getOrCreate()
housing_data=spark.read.parquet(f's3://{args.bucket}/{args.s3_input_prefix}/data.parquet.gzip')
housing_df_with_no_outliers = housing_data.where((housing_data.avgNumRooms<= 8) &
(housing_data.avgNumRooms>=2) &
(housing_data.avgNumBedrooms<=1.12) &
(housing_data.population<=1500) &
(housing_data.population>=250))
# Split the overall dataset into 80-20 training and validation
(train_df, validation_df) = housing_df_with_no_outliers.randomSplit([0.8, 0.2])
# Save the dataframe to parquet and upload to S3
train_df.write.parquet("s3://" + os.path.join(args.bucket, args.s3_output_prefix, "train/"))
# Save the dataframe to parquet and upload to S3 and upload to S3
validation_df.write.parquet("s3://" + os.path.join(args.bucket, args.s3_output_prefix, "validation/"))
if __name__ == "__main__":
main()
```
#### SageMaker Processing job using PySparkProcessor.
```
%local
from sagemaker.spark.processing import PySparkProcessor
spark_processor = PySparkProcessor(
base_job_name="sm-spark",
framework_version="2.4",
role=role,
instance_count=2,
instance_type="ml.m5.xlarge",
max_runtime_in_seconds=1200,
)
spark_processor.run(
submit_app="preprocess.py",
arguments=[
"--bucket",
bucket,
"--s3_input_prefix",
s3_input_prefix,
"--s3_output_prefix",
s3_output_prefix,
],
)
```
| github_jupyter |
# Bitcoin data analysis (log return)
## Yepeng Ding
### 12/13/2019
### Import libraries
```
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
import scipy.stats as stats
import statsmodels.tsa.api as smt
from arch import arch_model
```
### Import data
```
data = pd.read_csv('./data/BTC_USD_2013-10-01_2019-12-15-CoinDesk.csv')
data_date = pd.to_datetime(data['Date'], format='%Y-%m-%d')
data_cp = data['Closing Price (USD)']
data_size = len(data_cp)
print(data_cp.describe())
```
### Plot daily index
```
plt.title("Daily index", weight='bold')
plt.xlabel('Date')
plt.ylabel('Closing Price (USD)')
plt.plot(data_date, data_cp)
plt.show()
```
### Calculate log return
```
returns = pd.Series(np.diff(np.log(data_cp)))
print(returns.describe())
print('Skewness', stats.skew(returns))
print('Kurtosis', stats.kurtosis(returns))
print('Jarque–Bera test', stats.jarque_bera(returns))
```
### Plot log return
```
plt.title("Log return", weight='bold')
plt.xlabel('Date')
plt.ylabel('Log return')
plt.plot(data_date[1:], returns)
plt.show()
```
### Calculate squared log return
```
squared_returns = returns ** 2
print(squared_returns.describe())
```
### Plot squared log return
```
plt.title("Squared log return", weight='bold')
plt.xlabel('Date')
plt.ylabel('Squared log return')
plt.plot(data_date[1:], squared_returns)
plt.show()
```
### Calculate autocorrelation of log return
```
returns_acf = pd.DataFrame(smt.stattools.acf(returns,nlags=len(returns)))
print(returns_acf.describe())
```
### Plot autocorrelation of log return
```
plt.title("Autocorrelation of log return", weight='bold')
plt.xlabel('Lag (Day)')
plt.ylabel('Autocorrelation')
plt.plot(returns_acf[1:])
plt.show()
```
### Calculate autocorrelation of squared log return
```
squared_returns_acf = pd.DataFrame(smt.stattools.acf(squared_returns,nlags=len(squared_returns)))
print(squared_returns_acf.describe())
```
### Plot autocorrelation of squared log return
```
plt.title("Autocorrelation of squared log return", weight='bold')
plt.xlabel('Lag (Day)')
plt.ylabel('Autocorrelation')
plt.plot(squared_returns_acf[1:])
plt.show()
```
### GARCH model of log return
```
garch = arch_model(returns * 100)
garch_result = garch.fit()
print(garch_result.summary())
print('(alpha+beta) = ', garch_result.params['alpha[1]'] + garch_result.params['beta[1]'])
```
### Visualize the standardized residuals and conditional volatility
```
garch_fig = garch_result.plot(annualize='D')
```
### GJR-GARCH model of log return
```
gjr = arch_model(returns * 100, p=1, o=1, q=1)
gjr_result = gjr.fit(disp='off')
print(gjr_result.summary())
print('(1-alpha-beta-0.5*gamma) = ', 1 - gjr_result.params['alpha[1]'] - gjr_result.params['beta[1]'] - 0.5 * gjr_result.params['gamma[1]'])
```
### Calculate frequency of log return
```
# Adjust decimal point
precision = 3
returns = pd.DataFrame(np.around(returns, decimals=precision))
returns_frequency = returns.iloc[:,0].value_counts()
# Sort by the index
returns_frequency.sort_index(inplace=True)
print(returns_frequency.describe())
```
### Plot frequency of log return
```
plt.title("Frequency of log return", weight='bold')
plt.xlabel('Log return')
plt.ylabel('Frequency')
plt.scatter(returns_frequency.index, returns_frequency, s=3)
plt.show()
```
### Calculate probability density of log return
```
returns_pdf = returns_frequency / sum(returns_frequency)
print(returns_pdf.describe())
```
### Plot probability distribution of log return
```
plt.title("Probability density of log return", weight='bold')
plt.xlabel('Log return')
plt.ylabel('Probability density')
plt.plot(returns_pdf.index, returns_pdf)
plt.show()
```
### Normalize probability density of log return
```
interval_length = 10 ** (-precision)
returns_pdf_norm = returns_pdf / sum(returns_pdf * interval_length)
print(returns_pdf_norm.describe())
```
### Plot probability density normalization of log return
```
plt.title("Normalized probability density of log return", weight='bold')
plt.xlabel('Log return')
plt.ylabel('Normalized probability density')
plt.plot(returns_pdf_norm.index, returns_pdf_norm)
plt.show()
```
| github_jupyter |
```
%matplotlib inline
from matplotlib import pyplot
import numpy
```
AMUSE pre-defines a number of calculcated attributes on particle sets, such as the kinetic energy of the particles in the set. These calculated attributes are used often and provide a sufficient set to start out with, but they do not define a *complete* set. It's possible to define your own attributes and extend the attributes on a particle set.
```
from amuse.lab import *
```
As shown in the previous example, you can create a particle set by specifying the number of particles and setting their attributes. You can also create a particle set by using an inital condition function. For stellar clusters the commonly used plummer and king models are available. For this tutorial we will start with a king model. Global clusters created with a king model need the number of stars in the cluster and a dimensionless depth parameter that determines the depth of the potential well in the center of the cluster.
```
particles = new_king_model(1000, 3)
print(particles)
```
Common properties for a stellar cluster are its center of mass position, total kinetic energy and potential energy.
```
print("center of mass", particles.center_of_mass())
print("kinetic energy", particles.kinetic_energy())
print("potential energy", particles.potential_energy(G=nbody_system.G))
```
For the potential energy calculation we need to specify the gravitational constant, as the default value will use the gavitational constant in S.I. units and we are working in nbody units for this tutorial.
In N-body calculations and reporting, the kinetic and potential energy of a set of stars is often scaled to exactly 0.25 and -0.5 respectively. AMUSE also has a function for this.
```
particles.scale_to_standard()
print("kinetic energy", particles.kinetic_energy())
print("potential energy", particles.potential_energy(G=nbody_system.G))
```
*Note that the potential energy and scaling calculations are implemented as order N-squared operations*
Attributes of particle sets are always 1 dimensional by default, an array with a single value per particle attribute. But for some attributes it is easier to work with a 2d set, an array with multiple values (or an array of values) per particle attribute. For example, the positions of all particles. These attributes are called vector-attributes and are defined as a combination of 2 or more simple attributes.
The position attribute combines the values of the `x`, `y` and `z` attributes.
```
print(particles[0].x)
print(particles[0].y)
print(particles[0].z)
print(particles[0].position)
```
Other common vector attributes are `velocity` (combination of `vx`,`vy`,`vz`) and `acceleration` (combination of `ax`,`ay`,`az`).
You can set the value of a position attribute and the underlying x, y or z attributes will be changed.
```
particles[0].position = [0, 0.1, 0.2] | nbody_system.length
print(particles[0].x)
print(particles[0].y)
print(particles[0].z)
```
You can set the value of the x, y or z attribute and the position will change (as the position is just a combination of these attributes).
```
particles[0].x = 0.3 | nbody_system.length
print(particles[0].position)
```
You cannot change an item in the position array and thereby change the x, y, or z positions
```
# this will not change anything in the particles set as the position is a copy
particles[0].position[0] = 0.5 | nbody_system.length
print(particles[0].x)
print(particles[0].position)
```
You can use the position attribute on the entire set. Let's print the positions of the first 10 particles.
```
print(particles.position[0:10])
```
You can also use the position attribute to set values for the entire set
```
# set the position of all particles in the set to the same value
particles.position = [0.1, 0.2, 0.3] | nbody_system.length
print(particles.position[0:10])
print(particles.x[0:10])
```
Defining a new vector attribute is done by calling the `add_vector_attribute` or `add_global_vector_attribute`. The first call will define the attribute on the particle set and not on any other set. The second call will define the attribute on the particle set and any future sets created in the script. (The second call is used in the amuse framework itself to define the `position`, `velocity` and `acceleration` attributes)
```
particles.add_vector_attribute('position2d', ('x', 'y'))
print(particles[0].position2d)
```
If you enter `particles.add_` and press tab you'll notice two other function besides the `add_vector_attribute` function; `add_calculated_attribute` will create an attribute where the values are calculated based on other attributes, `add_function_attribute` will create a function on the set that gets the set and optional function parameters. These function also have global versions (`add_global_...`). The `add_global_function_attribute` call is used in the AMUSE framework to implement the `kinetic_energy` and `potential_energy` functions.
```
particles.add_function_attribute(
'calculate_mean_mass',
lambda particles: particles.mass.sum() / len(particles)
)
print(particles.calculate_mean_mass())
```
| github_jupyter |
<a href="https://colab.research.google.com/github/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_07_2_Keras_gan.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# T81-558: Applications of Deep Neural Networks
**Module 7: Generative Adversarial Networks**
* Instructor: [Jeff Heaton](https://sites.wustl.edu/jeffheaton/), McKelvey School of Engineering, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
* For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
# Module 7 Material
* Part 7.1: Introduction to GANS for Image and Data Generation [[Video]](https://www.youtube.com/watch?v=0QnCH6tlZgc&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_07_1_gan_intro.ipynb)
* **Part 7.2: Implementing a GAN in Keras** [[Video]](https://www.youtube.com/watch?v=T-MCludVNn4&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_07_2_Keras_gan.ipynb)
* Part 7.3: Face Generation with StyleGAN and Python [[Video]](https://www.youtube.com/watch?v=Wwwyr7cOBlU&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_07_3_style_gan.ipynb)
* Part 7.4: GANS for Semi-Supervised Learning in Keras [[Video]](https://www.youtube.com/watch?v=ZPewmEu7644&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_07_4_gan_semi_supervised.ipynb)
* Part 7.5: An Overview of GAN Research [[Video]](https://www.youtube.com/watch?v=cvCvZKvlvq4&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_07_5_gan_research.ipynb)
```
# Nicely formatted time string
def hms_string(sec_elapsed):
h = int(sec_elapsed / (60 * 60))
m = int((sec_elapsed % (60 * 60)) / 60)
s = sec_elapsed % 60
return "{}:{:>02}:{:>05.2f}".format(h, m, s)
```
# Part 7.2: Implementing DCGANs in Keras
Paper that described the type of DCGAN that we will create in this module. [[Cite:radford2015unsupervised]](https://arxiv.org/abs/1511.06434) This paper implements a DCGAN as follows:
* No pre-processing was applied to training images besides scaling to the range of the tanh activation function [-1, 1].
* All models were trained with mini-batch stochastic gradient descent (SGD) with a mini-batch size of 128.
* All weights were initialized from a zero-centered Normal distribution with standard deviation 0.02.
* In the LeakyReLU, the slope of the leak was set to 0.2 in all models.
* we used the Adam optimizer(Kingma & Ba, 2014) with tuned hyperparameters. We found the suggested learning rate of 0.001, to be too high, using 0.0002 instead.
* Additionally, we found leaving the momentum term $\beta{1}$ at the suggested value of 0.9 resulted in training oscillation and instability while reducing it to 0.5 helped stabilize training.
The paper also provides the following architecture guidelines for stable Deep Convolutional GANs:
* Replace any pooling layers with strided convolutions (discriminator) and fractional-strided convolutions (generator).
* Use batchnorm in both the generator and the discriminator.
* Remove fully connected hidden layers for deeper architectures.
* Use ReLU activation in generator for all layers except for the output, which uses Tanh.
* Use LeakyReLU activation in the discriminator for all layers.
While creating the material for this module I used a number of Internet resources, some of the most helpful were:
* [Deep Convolutional Generative Adversarial Network (TensorFlow 2.0 example code)](https://www.tensorflow.org/tutorials/generative/dcgan)
* [Keep Calm and train a GAN. Pitfalls and Tips on training Generative Adversarial Networks](https://medium.com/@utk.is.here/keep-calm-and-train-a-gan-pitfalls-and-tips-on-training-generative-adversarial-networks-edd529764aa9)
* [Collection of Keras implementations of Generative Adversarial Networks GANs](https://github.com/eriklindernoren/Keras-GAN)
* [dcgan-facegenerator](https://github.com/platonovsimeon/dcgan-facegenerator), [Semi-Paywalled Article by GitHub Author](https://medium.com/datadriveninvestor/generating-human-faces-with-keras-3ccd54c17f16)
The program created next will generate faces similar to these. While these faces are not perfect, they demonstrate how we can construct and train a GAN on or own. Later we will see how to import very advanced weights from nVidia to produce high resolution, realistic looking faces. Figure 7.GAN-GRID shows images from GAN training.
**Figure 7.GAN-GRID: GAN Neural Network Training**

As discussed in the previous module, the GAN is made up of two different neural networks: the discriminator and the generator. The generator generates the images, while the discriminator detects if a face is real or was generated. These two neural networks work as shown in Figure 7.GAN-EVAL:
**Figure 7.GAN-EVAL: Evaluating GANs**

The discriminator accepts an image as its input and produces number that is the probability of the input image being real. The generator accepts a random seed vector and generates an image from that random vector seed. An unlimited number of new images can be created by providing additional seeds.
I suggest running this code with a GPU, it will be very slow on a CPU alone. The following code mounts your Google drive for use with Google CoLab. If you are not using CoLab, the following code will not work.
```
try:
from google.colab import drive
drive.mount('/content/drive', force_remount=True)
COLAB = True
print("Note: using Google CoLab")
%tensorflow_version 2.x
except:
print("Note: not using Google CoLab")
COLAB = False
```
The following packages will be used to implement a basic GAN system in Python/Keras.
```
import tensorflow as tf
from tensorflow.keras.layers import Input, Reshape, Dropout, Dense
from tensorflow.keras.layers import Flatten, BatchNormalization
from tensorflow.keras.layers import Activation, ZeroPadding2D
from tensorflow.keras.layers import LeakyReLU
from tensorflow.keras.layers import UpSampling2D, Conv2D
from tensorflow.keras.models import Sequential, Model, load_model
from tensorflow.keras.optimizers import Adam
import numpy as np
from PIL import Image
from tqdm import tqdm
import os
import time
import matplotlib.pyplot as plt
```
These are the constants that define how the GANs will be created for this example. The higher the resolution, the more memory that will be needed. Higher resolution will also result in longer run times. For Google CoLab (with GPU) 128x128 resolution is as high as can be used (due to memory). Note that the resolution is specified as a multiple of 32. So **GENERATE_RES** of 1 is 32, 2 is 64, etc.
To run this you will need training data. The training data can be any collection of images. I suggest using training data from the following two locations. Simply unzip and combine to a common directory. This directory should be uploaded to Google Drive (if you are using CoLab). The constant **DATA_PATH** defines where these images are stored.
The source data (faces) used in this module can be found here:
* [Kaggle Faces Data New](https://www.kaggle.com/gasgallo/faces-data-new)
* [Kaggle Lag Dataset: Dataset of faces, from more than 1k different subjects](https://www.kaggle.com/gasgallo/lag-dataset)
```
# Generation resolution - Must be square
# Training data is also scaled to this.
# Note GENERATE_RES 4 or higher
# will blow Google CoLab's memory and have not
# been tested extensivly.
GENERATE_RES = 3 # Generation resolution factor
# (1=32, 2=64, 3=96, 4=128, etc.)
GENERATE_SQUARE = 32 * GENERATE_RES # rows/cols (should be square)
IMAGE_CHANNELS = 3
# Preview image
PREVIEW_ROWS = 4
PREVIEW_COLS = 7
PREVIEW_MARGIN = 16
# Size vector to generate images from
SEED_SIZE = 100
# Configuration
DATA_PATH = '/content/drive/My Drive/projects/faces'
EPOCHS = 50
BATCH_SIZE = 32
BUFFER_SIZE = 60000
print(f"Will generate {GENERATE_SQUARE}px square images.")
```
Next we will load and preprocess the images. This can take awhile. Google CoLab took around an hour to process. Because of this we store the processed file as a binary. This way we can simply reload the processed training data and quickly use it. It is most efficient to only perform this operation once. The dimensions of the image are encoded into the filename of the binary file because we need to regenerate it if these change.
```
# Image set has 11,682 images. Can take over an hour
# for initial preprocessing.
# Because of this time needed, save a Numpy preprocessed file.
# Note, that file is large enough to cause problems for
# sume verisons of Pickle,
# so Numpy binary files are used.
training_binary_path = os.path.join(DATA_PATH,
f'training_data_{GENERATE_SQUARE}_{GENERATE_SQUARE}.npy')
print(f"Looking for file: {training_binary_path}")
if not os.path.isfile(training_binary_path):
start = time.time()
print("Loading training images...")
training_data = []
faces_path = os.path.join(DATA_PATH,'face_images')
for filename in tqdm(os.listdir(faces_path)):
path = os.path.join(faces_path,filename)
image = Image.open(path).resize((GENERATE_SQUARE,
GENERATE_SQUARE),Image.ANTIALIAS)
training_data.append(np.asarray(image))
training_data = np.reshape(training_data,(-1,GENERATE_SQUARE,
GENERATE_SQUARE,IMAGE_CHANNELS))
training_data = training_data.astype(np.float32)
training_data = training_data / 127.5 - 1.
print("Saving training image binary...")
np.save(training_binary_path,training_data)
elapsed = time.time()-start
print (f'Image preprocess time: {hms_string(elapsed)}')
else:
print("Loading previous training pickle...")
training_data = np.load(training_binary_path)
```
We will use a TensorFlow **Dataset** object to actually hold the images. This allows the data to be quickly shuffled int divided into the appropriate batch sizes for training.
```
# Batch and shuffle the data
train_dataset = tf.data.Dataset.from_tensor_slices(training_data) \
.shuffle(BUFFER_SIZE).batch(BATCH_SIZE)
```
The code below creates the generator and discriminator.
Next we actually build the discriminator and the generator. Both will be trained with the Adam optimizer.
```
def build_generator(seed_size, channels):
model = Sequential()
model.add(Dense(4*4*256,activation="relu",input_dim=seed_size))
model.add(Reshape((4,4,256)))
model.add(UpSampling2D())
model.add(Conv2D(256,kernel_size=3,padding="same"))
model.add(BatchNormalization(momentum=0.8))
model.add(Activation("relu"))
model.add(UpSampling2D())
model.add(Conv2D(256,kernel_size=3,padding="same"))
model.add(BatchNormalization(momentum=0.8))
model.add(Activation("relu"))
# Output resolution, additional upsampling
model.add(UpSampling2D())
model.add(Conv2D(128,kernel_size=3,padding="same"))
model.add(BatchNormalization(momentum=0.8))
model.add(Activation("relu"))
if GENERATE_RES>1:
model.add(UpSampling2D(size=(GENERATE_RES,GENERATE_RES)))
model.add(Conv2D(128,kernel_size=3,padding="same"))
model.add(BatchNormalization(momentum=0.8))
model.add(Activation("relu"))
# Final CNN layer
model.add(Conv2D(channels,kernel_size=3,padding="same"))
model.add(Activation("tanh"))
return model
def build_discriminator(image_shape):
model = Sequential()
model.add(Conv2D(32, kernel_size=3, strides=2, input_shape=image_shape,
padding="same"))
model.add(LeakyReLU(alpha=0.2))
model.add(Dropout(0.25))
model.add(Conv2D(64, kernel_size=3, strides=2, padding="same"))
model.add(ZeroPadding2D(padding=((0,1),(0,1))))
model.add(BatchNormalization(momentum=0.8))
model.add(LeakyReLU(alpha=0.2))
model.add(Dropout(0.25))
model.add(Conv2D(128, kernel_size=3, strides=2, padding="same"))
model.add(BatchNormalization(momentum=0.8))
model.add(LeakyReLU(alpha=0.2))
model.add(Dropout(0.25))
model.add(Conv2D(256, kernel_size=3, strides=1, padding="same"))
model.add(BatchNormalization(momentum=0.8))
model.add(LeakyReLU(alpha=0.2))
model.add(Dropout(0.25))
model.add(Conv2D(512, kernel_size=3, strides=1, padding="same"))
model.add(BatchNormalization(momentum=0.8))
model.add(LeakyReLU(alpha=0.2))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(1, activation='sigmoid'))
return model
```
As we progress through training images will be produced to show the progress. These images will contain a number of rendered faces that show how good the generator has become. These faces will be
```
def save_images(cnt,noise):
image_array = np.full((
PREVIEW_MARGIN + (PREVIEW_ROWS * (GENERATE_SQUARE+PREVIEW_MARGIN)),
PREVIEW_MARGIN + (PREVIEW_COLS * (GENERATE_SQUARE+PREVIEW_MARGIN)), 3),
255, dtype=np.uint8)
generated_images = generator.predict(noise)
generated_images = 0.5 * generated_images + 0.5
image_count = 0
for row in range(PREVIEW_ROWS):
for col in range(PREVIEW_COLS):
r = row * (GENERATE_SQUARE+16) + PREVIEW_MARGIN
c = col * (GENERATE_SQUARE+16) + PREVIEW_MARGIN
image_array[r:r+GENERATE_SQUARE,c:c+GENERATE_SQUARE] \
= generated_images[image_count] * 255
image_count += 1
output_path = os.path.join(DATA_PATH,'output')
if not os.path.exists(output_path):
os.makedirs(output_path)
filename = os.path.join(output_path,f"train-{cnt}.png")
im = Image.fromarray(image_array)
im.save(filename)
generator = build_generator(SEED_SIZE, IMAGE_CHANNELS)
noise = tf.random.normal([1, SEED_SIZE])
generated_image = generator(noise, training=False)
plt.imshow(generated_image[0, :, :, 0])
image_shape = (GENERATE_SQUARE,GENERATE_SQUARE,IMAGE_CHANNELS)
discriminator = build_discriminator(image_shape)
decision = discriminator(generated_image)
print (decision)
```
Loss functions must be developed that allow the generator and discriminator to be trained in an adversarial way. Because these two neural networks are being trained independently they must be trained in two separate passes. This requires two separate loss functions and also two separate updates to the gradients. When the discriminator's gradients are applied to decrease the discriminator's loss it is important that only the discriminator's weights are update. It is not fair, nor will it produce good results, to adversarially damage the weights of the generator to help the discriminator. A simple backpropagation would do this. It would simultaneously affect the weights of both generator and discriminator to lower whatever loss it was assigned to lower.
Figure 7.TDIS shows how the discriminator is trained.
**Figure 7.TDIS: Training the Discriminator**

Here a training set is generated with an equal number of real and fake images. The real images are randomly sampled (chosen) from the training data. An equal number of random images are generated from random seeds. For the discriminator training set, the $x$ contains the input images and the $y$ contains a value of 1 for real images and 0 for generated ones.
Likewise, the Figure 7.TGEN shows how the generator is trained.
**Figure 7.TGEN: Training the Generator**

For the generator training set, the $x$ contains the random seeds to generate images and the $y$ always contains the value of 1, because the optimal is for the generator to have generated such good images that the discriminiator was fooled into assigning them a probability near 1.
```
# This method returns a helper function to compute cross entropy loss
cross_entropy = tf.keras.losses.BinaryCrossentropy(from_logits=True)
def discriminator_loss(real_output, fake_output):
real_loss = cross_entropy(tf.ones_like(real_output), real_output)
fake_loss = cross_entropy(tf.zeros_like(fake_output), fake_output)
total_loss = real_loss + fake_loss
return total_loss
def generator_loss(fake_output):
return cross_entropy(tf.ones_like(fake_output), fake_output)
```
Both the generator and discriminator use Adam and the same learning rate and momentum. This does not need to be the case. If you use a **GENERATE_RES** greater than 3 you may need to tune these learning rates, as well as other training and hyperparameters.
```
generator_optimizer = tf.keras.optimizers.Adam(1.5e-4,0.5)
discriminator_optimizer = tf.keras.optimizers.Adam(1.5e-4,0.5)
```
The following function is where most of the training takes place for both the discriminator and the generator. This function was based on the GAN provided by the [TensorFlow Keras exmples](https://www.tensorflow.org/tutorials/generative/dcgan) documentation. The first thing you should notice about this function is that it is annotated with the **tf.function** annotation. This causes the function to be precompiled and improves performance.
This function trans differently than the code we previously saw for training. This code makes use of **GradientTape** to allow the discriminator and generator to be trained together, yet separately.
```
# Notice the use of `tf.function`
# This annotation causes the function to be "compiled".
@tf.function
def train_step(images):
seed = tf.random.normal([BATCH_SIZE, SEED_SIZE])
with tf.GradientTape() as gen_tape, tf.GradientTape() as disc_tape:
generated_images = generator(seed, training=True)
real_output = discriminator(images, training=True)
fake_output = discriminator(generated_images, training=True)
gen_loss = generator_loss(fake_output)
disc_loss = discriminator_loss(real_output, fake_output)
gradients_of_generator = gen_tape.gradient(\
gen_loss, generator.trainable_variables)
gradients_of_discriminator = disc_tape.gradient(\
disc_loss, discriminator.trainable_variables)
generator_optimizer.apply_gradients(zip(
gradients_of_generator, generator.trainable_variables))
discriminator_optimizer.apply_gradients(zip(
gradients_of_discriminator,
discriminator.trainable_variables))
return gen_loss,disc_loss
def train(dataset, epochs):
fixed_seed = np.random.normal(0, 1, (PREVIEW_ROWS * PREVIEW_COLS,
SEED_SIZE))
start = time.time()
for epoch in range(epochs):
epoch_start = time.time()
gen_loss_list = []
disc_loss_list = []
for image_batch in dataset:
t = train_step(image_batch)
gen_loss_list.append(t[0])
disc_loss_list.append(t[1])
g_loss = sum(gen_loss_list) / len(gen_loss_list)
d_loss = sum(disc_loss_list) / len(disc_loss_list)
epoch_elapsed = time.time()-epoch_start
print (f'Epoch {epoch+1}, gen loss={g_loss},disc loss={d_loss},'\
' {hms_string(epoch_elapsed)}')
save_images(epoch,fixed_seed)
elapsed = time.time()-start
print (f'Training time: {hms_string(elapsed)}')
train(train_dataset, EPOCHS)
generator.save(os.path.join(DATA_PATH,"face_generator.h5"))
```
| github_jupyter |
# Convolutional Neural Networks
## Standard LeNet5 with PyTorch
### Xavier Bresson, Sept. 2017
Implementation of original LeNet5 Convolutional Neural Networks:<br>
Gradient-based learning applied to document recognition<br>
Y LeCun, L Bottou, Y Bengio, P Haffner<br>
Proceedings of the IEEE 86 (11), 2278-2324<br>
```
import torch
from torch.autograd import Variable
import torch.nn.functional as F
import torch.nn as nn
import pdb #pdb.set_trace()
import collections
import time
import numpy as np
import os
os.environ["CUDA_DEVICE_ORDER"]="PCI_BUS_ID"
os.environ["CUDA_VISIBLE_DEVICES"]="0"
if torch.cuda.is_available():
print('cuda available')
dtypeFloat = torch.cuda.FloatTensor
dtypeLong = torch.cuda.LongTensor
torch.cuda.manual_seed(1)
else:
print('cuda not available')
dtypeFloat = torch.FloatTensor
dtypeLong = torch.LongTensor
torch.manual_seed(1)
```
# MNIST
```
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets('datasets', one_hot=False) # load data in folder datasets/
train_data = mnist.train.images.astype(np.float32)
val_data = mnist.validation.images.astype(np.float32)
test_data = mnist.test.images.astype(np.float32)
train_labels = mnist.train.labels
val_labels = mnist.validation.labels
test_labels = mnist.test.labels
print(train_data.shape)
print(train_labels.shape)
print(val_data.shape)
print(val_labels.shape)
print(test_data.shape)
print(test_labels.shape)
```
# ConvNet LeNet5
### Layers: CL32-MP4-CL64-MP4-FC512-FC10
```
# class definition
class ConvNet_LeNet5(nn.Module):
def __init__(self, net_parameters):
print('ConvNet: LeNet5\n')
super(ConvNet_LeNet5, self).__init__()
Nx, Ny, CL1_F, CL1_K, CL2_F, CL2_K, FC1_F, FC2_F = net_parameters
FC1Fin = CL2_F*(Nx//4)**2
# graph CL1
self.conv1 = nn.Conv2d(1, CL1_F, CL1_K, padding=(2, 2))
Fin = CL1_K**2; Fout = CL1_F;
scale = np.sqrt( 2.0/ (Fin+Fout) )
self.conv1.weight.data.uniform_(-scale, scale)
self.conv1.bias.data.fill_(0.0)
# graph CL2
self.conv2 = nn.Conv2d(CL1_F, CL2_F, CL2_K, padding=(2, 2))
Fin = CL1_F*CL2_K**2; Fout = CL2_F;
scale = np.sqrt( 2.0/ (Fin+Fout) )
self.conv2.weight.data.uniform_(-scale, scale)
self.conv2.bias.data.fill_(0.0)
# FC1
self.fc1 = nn.Linear(FC1Fin, FC1_F)
Fin = FC1Fin; Fout = FC1_F;
scale = np.sqrt( 2.0/ (Fin+Fout) )
self.fc1.weight.data.uniform_(-scale, scale)
self.fc1.bias.data.fill_(0.0)
self.FC1Fin = FC1Fin
# FC2
self.fc2 = nn.Linear(FC1_F, FC2_F)
Fin = FC1_F; Fout = FC2_F;
scale = np.sqrt( 2.0/ (Fin+Fout) )
self.fc2.weight.data.uniform_(-scale, scale)
self.fc2.bias.data.fill_(0.0)
# max pooling
self.pool = nn.MaxPool2d(2, 2)
def forward(self, x, d):
# CL1
x = self.conv1(x)
x = F.relu(x)
x = self.pool(x)
# CL2
x = self.conv2(x)
x = F.relu(x)
x = self.pool(x)
# FC1
x = x.permute(0,3,2,1).contiguous() # reshape from pytorch array to tensorflow array
x = x.view(-1, self.FC1Fin)
x = self.fc1(x)
x = F.relu(x)
x = nn.Dropout(d)(x)
# FC2
x = self.fc2(x)
return x
def loss(self, y, y_target, l2_regularization):
loss = nn.CrossEntropyLoss()(y,y_target)
l2_loss = 0.0
for param in self.parameters():
data = param* param
l2_loss += data.sum()
loss += 0.5* l2_regularization* l2_loss
return loss
def update(self, lr):
update = torch.optim.SGD( self.parameters(), lr=lr, momentum=0.9 )
return update
def update_learning_rate(self, optimizer, lr):
for param_group in optimizer.param_groups:
param_group['lr'] = lr
return optimizer
def evaluation(self, y_predicted, test_l):
_, class_predicted = torch.max(y_predicted.data, 1)
return 100.0* (class_predicted == test_l).sum()/ y_predicted.size(0)
# Delete existing network if exists
try:
del net
print('Delete existing network\n')
except NameError:
print('No existing network to delete\n')
# network parameters
Nx = Ny = 28
CL1_F = 32
CL1_K = 5
CL2_F = 64
CL2_K = 5
FC1_F = 512
FC2_F = 10
net_parameters = [Nx, Ny, CL1_F, CL1_K, CL2_F, CL2_K, FC1_F, FC2_F]
# instantiate the object net of the class
net = ConvNet_LeNet5(net_parameters)
if torch.cuda.is_available():
net.cuda()
print(net)
# Weights
L = list(net.parameters())
# learning parameters
learning_rate = 0.05
dropout_value = 0.5
l2_regularization = 5e-4
batch_size = 100
num_epochs = 20
train_size = train_data.shape[0]
nb_iter = int(num_epochs * train_size) // batch_size
print('num_epochs=',num_epochs,', train_size=',train_size,', nb_iter=',nb_iter)
# Optimizer
global_lr = learning_rate
global_step = 0
decay = 0.95
decay_steps = train_size
lr = learning_rate
optimizer = net.update(lr)
# loop over epochs
indices = collections.deque()
for epoch in range(num_epochs): # loop over the dataset multiple times
# reshuffle
indices.extend(np.random.permutation(train_size)) # rand permutation
# reset time
t_start = time.time()
# extract batches
running_loss = 0.0
running_accuray = 0
running_total = 0
while len(indices) >= batch_size:
# extract batches
batch_idx = [indices.popleft() for i in range(batch_size)]
train_x, train_y = train_data[batch_idx,:].T, train_labels[batch_idx].T
train_x = np.reshape(train_x,[28,28,batch_size])[:,:,:,None]
train_x = np.transpose(train_x,[2,3,1,0]) # reshape from pytorch array to tensorflow array
train_x = Variable( torch.FloatTensor(train_x).type(dtypeFloat) , requires_grad=False)
train_y = train_y.astype(np.int64)
train_y = torch.LongTensor(train_y).type(dtypeLong)
train_y = Variable( train_y , requires_grad=False)
# Forward
y = net.forward(train_x, dropout_value)
loss = net.loss(y,train_y,l2_regularization)
loss_train = loss.data[0]
# Accuracy
acc_train = net.evaluation(y,train_y.data)
# backward
loss.backward()
# Update
global_step += batch_size # to update learning rate
optimizer.step()
optimizer.zero_grad()
# loss, accuracy
running_loss += loss_train
running_accuray += acc_train
running_total += 1
# print
if not running_total%100: # print every x mini-batches
print('epoch= %d, i= %4d, loss(batch)= %.4f, accuray(batch)= %.2f' % (epoch+1, running_total, loss_train, acc_train))
# print
t_stop = time.time() - t_start
print('epoch= %d, loss(train)= %.3f, accuracy(train)= %.3f, time= %.3f, lr= %.5f' %
(epoch+1, running_loss/running_total, running_accuray/running_total, t_stop, lr))
# update learning rate
lr = global_lr * pow( decay , float(global_step// decay_steps) )
optimizer = net.update_learning_rate(optimizer, lr)
# Test set
running_accuray_test = 0
running_total_test = 0
indices_test = collections.deque()
indices_test.extend(range(test_data.shape[0]))
t_start_test = time.time()
while len(indices_test) >= batch_size:
batch_idx_test = [indices_test.popleft() for i in range(batch_size)]
test_x, test_y = test_data[batch_idx_test,:].T, test_labels[batch_idx_test].T
test_x = np.reshape(test_x,[28,28,batch_size])[:,:,:,None]
test_x = np.transpose(test_x,[2,3,1,0]) # reshape from pytorch array to tensorflow array
test_x = Variable( torch.FloatTensor(test_x).type(dtypeFloat) , requires_grad=False)
y = net.forward(test_x, 0.0)
test_y = test_y.astype(np.int64)
test_y = torch.LongTensor(test_y).type(dtypeLong)
test_y = Variable( test_y , requires_grad=False)
acc_test = net.evaluation(y,test_y.data)
running_accuray_test += acc_test
running_total_test += 1
t_stop_test = time.time() - t_start_test
print(' accuracy(test) = %.3f %%, time= %.3f' % (running_accuray_test / running_total_test, t_stop_test))
```
| github_jupyter |
# Introduction to Python
### [Matplotlib](http://matplotlib.org/users/pyplot_tutorial.html)
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
#%matplotlib notebook
```
matplotlib.pyplot is a collection of functions that make matplotlib work like MATLAB. Each pyplot function makes some change to a figure: e.g., creates a figure, creates a plotting area in a figure, plots some lines in a plotting area, decorates the plot with labels, etc.
In matplotlib.pyplot various states are preserved across function calls, so that it keeps track of things like the current figure and plotting area, and the plotting functions are directed to the current axes (please note that "axes" here and in most places in the documentation refers to the axes part of a figure and not the strict mathematical term for more than one axis).
#### [Gallery](https://matplotlib.org/3.1.1/gallery/index.html)
#### [Colormaps](https://matplotlib.org/3.1.1/gallery/color/colormap_reference.html)
#### [ColorCodes](https://matplotlib.org/2.1.1/api/_as_gen/matplotlib.pyplot.plot.html)
```
plt.subplots(figsize=(8,6))
plt.plot([1,3,2,4,7,16])
plt.ylabel('some numbers')
plt.xlabel('Simple Graph')
plt.axis([0,10,0,20])
plt.show()
plt.plot([1,2,8,4], [1,4,9,10], 'yo')
plt.axis([0, 9, 0, 20])
plt.show()
# evenly sampled time at 200ms intervals
t = np.arange(0., 5., 0.2)
# red dashes, blue squares and green triangles
plt.plot(t, t, 'r--')
plt.plot(t, t**2, 'bs')
plt.plot(t, t**3, 'g^')
#plt.plot(t, t, 'r--', t, t**2, 'bs', t, t**3, 'g^') #all at the same time
plt.show()
x = np.linspace(0, 2 * np.pi, 400)
y = np.sin(x ** 2)
plt.plot(x,y)
plt.show()
```
Just a figure and one subplot
```
f, ax = plt.subplots()
ax.set_title('Simple plot')
ax.plot(x, y)
plt.show()
ax = plt.subplot(111)
t = np.arange(0.0, 5.0, 0.01)
s = np.cos(2*np.pi*t)
line, = plt.plot(t, s, lw=2)
plt.annotate('local max', xy=(2, 1), xytext=(3, 1.5),
arrowprops=dict(facecolor='black', shrink=0.05),
)
plt.ylim(-2,2)
plt.show()
A = np.array([1,np.nan, 3,5,1,2,5,2,4,1,2,np.nan,2,1,np.nan,2,np.nan,1,2])
plt.figure()
plt.hist(A[~np.isnan(A)],bins=10)
plt.show()
A
mu, sigma = 100, 15
xh = mu + sigma * np.random.randn(100000)
xh = np.random.normal(mu, sigma, 100000)
# the histogram of the data
n, bins, patches = plt.hist(xh, 150, density=True, facecolor='g', alpha=0.75)
plt.xlabel('Smarts')
plt.ylabel('Probability')
plt.title('Histogram of IQ')
plt.text(60, .025, '$\mu=100,\ \sigma=15$')
plt.axis([40, 160, 0, 0.03])
plt.grid(True)
plt.show()
data = np.random.random((10, 4))
data2 = np.random.random((10, 4))
plt.subplots_adjust(bottom = 0.1)
plt.xlabel('$\partial \Delta/\partial\Phi[$mm$/^{\circ}]$', fontsize = 16)
plt.ylabel('$\Delta$ [mm]', fontsize = 16)
plt.scatter(data[:, 0], data[:, 1], marker = 'o', c = data[:, 3], s = data[:, 3]*1500,
cmap = plt.cm.Spectral, vmin = min(data[:, 3]), vmax = max(data[:, 3]))
plt.scatter(data2[:, 0], data2[:, 1], marker = '^', c = data2[:, 2], s = data2[:, 2]*500,
cmap = plt.cm.Spectral, vmin = min(data2[:, 2]), vmax = max(data2[:, 2]))
cbar = plt.colorbar(ticks = [min(data2[:, 2]), max(data2[:, 2])])
cbar.ax.set_yticklabels(['Low', 'High'])
cbar.set_label(r'My Scale')
plt.show()
a = np.random.random((8,8))
#a = np.arange(64).reshape(8,8)
p = plt.pcolor(a)
plt.colorbar()
plt.show()
```
Two subplots, the axes array is 1-d
```
x = np.linspace(0, 2 * np.pi, 400)
y = np.sin(x ** 2)
f, ax = plt.subplots(2, figsize=(8,8), sharex=True)
ax[0].plot(x, y)
ax[0].plot(2*x, y)
ax[0].set_title('Sharing X axis')
ax[1].scatter(x, y)
ax[1].scatter(2*x, y)
plt.show()
```
Two subplots, unpack the axes array immediately
```
f, ax = plt.subplots(1, 2, sharey=True)
ax[0].plot(x, y)
ax[0].set_title('Sharing Y axis')
ax[1].scatter(x, y)
plt.show()
```
Three subplots sharing both x/y axes
```
f, ax = plt.subplots(3, sharex=True, sharey=True)
ax[0].plot(x, y)
ax[0].set_title('Sharing both axes')
ax[1].scatter(x, y)
ax[2].scatter(x, 2 * y ** 2 - 1, color='r')
#Fine-tune figure; make subplots close to each other and hide x ticks for all but bottom plot.
f.subplots_adjust(hspace=0)
plt.setp([a.get_xticklabels() for a in f.axes[:-1]], visible=False)
plt.show()
```
row and column sharing
```
f, ax = plt.subplots(2, 2, sharex='col', sharey='row')
ax[0,0].plot(x, y)
ax[0,1].set_title('Sharing x per column, y per row')
ax[0,1].scatter(x, y)
ax[1,0].scatter(x, 2 * y ** 2 - 1, color='r')
ax[1,1].plot(x, 2 * y ** 2 - 1, color='r')
plt.show()
```
Four axes, returned as a 2-d array
```
f, ax = plt.subplots(2, 2)
ax[0, 0].plot(x, y)
ax[0, 0].set_title('Axis [0,0]')
ax[0, 1].scatter(x, y)
ax[0, 1].set_title('Axis [0,1]')
ax[1, 0].plot(x, y ** 2)
ax[1, 0].set_title('Axis [1,0]')
ax[1, 1].scatter(x, y ** 2)
ax[1, 1].set_title('Axis [1,1]')
#Fine-tune figure; hide x ticks for top plots and y ticks for right plots
plt.setp([a.get_xticklabels() for a in ax[0, :]], visible=False)
plt.setp([a.get_yticklabels() for a in ax[:, 1]], visible=False)
plt.show()
```
Four polar axes
```
plt.subplots(2, 2, subplot_kw=dict(polar=True))
plt.show()
```
### Matplotlib [Animation](https://towardsdatascience.com/animations-with-matplotlib-d96375c5442c)
+ create a figure window with a single axis in the figure.
+ create our empty line object which is essentially the one to be modified in the animation. The line object will be populated with data later.
+ create the init function that will make the animation happen. The init function initializes the data and also sets the axis limits.
+ define the animation function which takes in the frame number(i) as the parameter and creates a sine wave(or any other animation) which a shift depending upon the value of i. This function here returns a tuple of the plot objects which have been modified which tells the animation framework what parts of the plot should be animated.
+ create the actual animation object. The blit parameter ensures that only those pieces of the plot are re-drawn which have been changed.
```
from matplotlib.animation import FuncAnimation
plt.style.use('seaborn-pastel')
fig = plt.figure()
ax = plt.axes(xlim=(0, 4), ylim=(-2, 2))
line, = ax.plot([], [], lw=3)
def init():
line.set_data([], [])
return line,
def animate(i):
x = np.linspace(0, 4, 1000)
y = np.sin(2 * np.pi * (x - 0.01 * i))
line.set_data(x, y)
return line,
animation = FuncAnimation(fig, animate, init_func=init, frames=200, interval=20, blit=True)
animation.save('../Data/sine_wave.gif', writer='imagemagick')
plt.style.use('dark_background')
fig = plt.figure()
ax = plt.axes(xlim=(-50, 50), ylim=(-50, 50))
line, = ax.plot([], [], lw=2)
# initialization function
def init():
# creating an empty plot/frame
line.set_data([], [])
return line,
# lists to store x and y axis points
xdata, ydata = [], []
# animation function
def animate(i):
# t is a parameter
t = 0.1*i
# x, y values to be plotted
x = t*np.sin(t)
y = t*np.cos(t)
# appending new points to x, y axes points list
xdata.append(x)
ydata.append(y)
line.set_data(xdata, ydata)
return line,
# setting a title for the plot
plt.title('Creating a growing coil with matplotlib!')
# hiding the axis details
plt.axis('off')
# call the animator
animation = FuncAnimation(fig, animate, init_func=init, frames=500, interval=20, blit=True)
# save the animation as mp4 video file
animation.save('../Data/coil.gif', writer='imagemagick')
```
| github_jupyter |
<a href="https://colab.research.google.com/github/ATOMScience-org/AMPL/blob/master/atomsci/ddm/examples/tutorials/11_CHEMBL26_SCN5A_IC50_prediction.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
<h1>Building a Graph Convolutional Network Model for Drug Response Prediction</h1>
The ATOM Modeling PipeLine (AMPL; https://github.com/ATOMScience-org/AMPL) is an open-source, modular, extensible software pipeline for building and sharing models to advance in silico drug discovery.
## Time to run: 6 minutes
```
!date
```
## Change your runtime to GPU
Go to **Runtime** --> Change **runtime type** to "GPU"
## Goal: Use AMPL for predicting binding affinities -IC50 values- of ligands that could bind to human **Sodium channel protein type 5 subunit alpha** protein using Graph Convolutional Network Model. ChEMBL database is the source of the binding affinities (pIC50).
In this notebook, we describe the following steps using AMPL:
1. Read a ML ready dataset
2. Fit a Graph Convolutional model
3. Predict pIC50 values of withheld compounds
## Set up
We first import the AMPL modules for use in this notebook.
The relevant AMPL modules for this example are listed below:
|module|Description|
|-|-|
|`atomsci.ddm.pipeline.model_pipeline`|The model pipeline module is used to fit models and load models for prediction.|
|`atomsci.ddm.pipeline.parameter_parser`|The parameter parser reads through pipeline options for the model pipeline.|
|`atomsci.ddm.utils.curate_data`|The curate data module is used for data loading and pre-processing.|
|`atomsci.ddm.utils.struct_utils`|The structure utilities module is used to process loaded structures.|
import atomsci.ddm.pipeline.compare_models as cmp
import atomsci.ddm.pipeline.model_pipeline as mp
import atomsci.ddm.pipeline.parameter_parser as parse
## Install AMPL
```
! pip install rdkit-pypi
! pip install --pre deepchem # 2.6.0 dev
# ! pip install deepchem==2.5.0
import deepchem
# print(deepchem.__version__)
! pip install umap
! pip install llvmlite==0.34.0 --ignore-installed
! pip install umap-learn
! pip install molvs
! pip install bravado
import deepchem as dc
# get the Install AMPL_GPU_test.sh
!wget 'https://raw.githubusercontent.com/ATOMScience-org/AMPL/master/atomsci/ddm/examples/tutorials/config/install_AMPL_GPU_test.sh'
# run the script to install AMPL
! chmod u+x install_AMPL_GPU_test.sh
! ./install_AMPL_GPU_test.sh
dc.__version__
# We temporarily disable warnings for demonstration.
# FutureWarnings and DeprecationWarnings are present from some of the AMPL
# dependency modules.
import warnings
warnings.filterwarnings('ignore')
import json
import requests
import sys
import atomsci.ddm.pipeline.compare_models as cmp
import atomsci.ddm.pipeline.model_pipeline as mp
import atomsci.ddm.pipeline.parameter_parser as parse
import os
os.mkdir('chembl_activity_models')
```
**Let us display the dataset**
```
import pandas as pd
import requests
import io
url = 'https://raw.githubusercontent.com/ATOMScience-org/AMPL/master/atomsci/ddm/examples/tutorials/datasets/ChEMBL26_SCN5A_IC50_human_ml_ready.csv'
download = requests.get(url).content
df = pd.read_csv(url, index_col=0)
# Reading the downloaded content and turning it into a pandas dataframe
df = pd.read_csv(io.StringIO(download.decode('utf-8')))
df.iloc[0:5, 0:5]
df.to_csv('ChEMBL26_SCN5A_IC50_human_ml_ready.csv', index=False)
df
```
base_splitter
Description: Type of splitter to use for train/validation split if temporal split used for test set. May be random, scaffold, or ave_min. The allowable choices are set in splitter.py
Check here for details, https://github.com/ATOMScience-org/AMPL/blob/master/atomsci/ddm/docs/PARAMETERS.md
```
split_config = {
"script_dir": "/content/AMPL/atomsci/ddm",
"dataset_key" : "/content/ChEMBL26_SCN5A_IC50_human_ml_ready.csv",
"datastore": "False",
"split_only": "True",
"splitter": "scaffold",
"split_valid_frac": "0.15",
"split_test_frac": "0.15",
"previously_split": "False",
"prediction_type": "regression",
"response_cols" : "pIC50",
"id_col": "compound_id",
"smiles_col" : "base_rdkit_smiles",
"result_dir": "/content/chembl_activity_models",
"system": "LC",
"transformers": "True",
"model_type": "NN",
"featurizer": "graphconv",
"descriptor_type": "graphconv",
"learning_rate": "0.0007",
"layer_sizes": "64,64,32",
"dropouts" : "0.0,0.0,0.0",
"save_results": "False",
"max_epochs": "100",
"verbose": "True"
}
split_params = parse.wrapper(split_config)
split_model = mp.ModelPipeline(split_params)
split_uuid = split_model.split_dataset()
split_uuid
!pip install --upgrade gspread
!date
```
## Train the mode (~ 10 minutes)
```
train_config = {
"script_dir": "/content/AMPL/atomsci/ddm",
"dataset_key" : "/content/ChEMBL26_SCN5A_IC50_human_ml_ready.csv",
"datastore": "False",
"uncertainty": "False",
"splitter": "scaffold",
"split_valid_frac": "0.15",
"split_test_frac": "0.15",
"previously_split": "True",
"split_uuid": "{}".format(split_uuid),
"prediction_type": "regression",
"response_cols" : "pIC50",
"id_col": "compound_id",
"smiles_col" : "base_rdkit_smiles",
"result_dir": "/content/chembl_activity_models",
"system": "LC",
"transformers": "True",
"model_type": "NN",
"featurizer": "graphconv",
"descriptor_type": "graphconv",
"learning_rate": "0.0007",
"layer_sizes": "64,64,32",
"dropouts" : "0.0,0.0,0.0",
"save_results": "False",
"max_epochs": "100",
"verbose": "True"
}
train_params = parse.wrapper(train_config)
train_model = mp.ModelPipeline(train_params)
```
## Train_model took ~ 18 minutes on a GPU (~ 30 minutes on a CPU)
```
mp.ampl_version
dc.__version__
%cd github
!ls
train_model.train_model()
perf_df = cmp.get_filesystem_perf_results('/content/chembl_activity_models', pred_type='regression')
perf_df
!date
```
| github_jupyter |
<a href="https://www.bigdatauniversity.com"><img src = "https://ibm.box.com/shared/static/cw2c7r3o20w9zn8gkecaeyjhgw3xdgbj.png" width = 400, align = "center"></a>
# <center>Density-Based Clustering</center>
Most of the traditional clustering techniques, such as k-means, hierarchical and fuzzy clustering, can be used to group data without supervision.
However, when applied to tasks with arbitrary shape clusters, or clusters within cluster, the traditional techniques might be unable to achieve good results. That is, elements in the same cluster might not share enough similarity or the performance may be poor.
Additionally, Density-based Clustering locates regions of high density that are separated from one another by regions of low density. Density, in this context, is defined as the number of points within a specified radius.
In this section, the main focus will be manipulating the data and properties of DBSCAN and observing the resulting clustering.
Import the following libraries:
<ul>
<li> <b>numpy as np</b> </li>
<li> <b>DBSCAN</b> from <b>sklearn.cluster</b> </li>
<li> <b>make_blobs</b> from <b>sklearn.datasets.samples_generator</b> </li>
<li> <b>StandardScaler</b> from <b>sklearn.preprocessing</b> </li>
<li> <b>matplotlib.pyplot as plt</b> </li>
</ul> <br>
Remember <b> %matplotlib inline </b> to display plots
```
# Notice: For visualization of map, you need basemap package.
# if you dont have basemap install on your machine, you can use the following line to install it
!conda install -c conda-forge basemap==1.1.0 matplotlib==2.2.2 -y
# Notice: you maight have to refresh your page and re-run the notebook after installation
import numpy as np
from sklearn.cluster import DBSCAN
from sklearn.datasets.samples_generator import make_blobs
from sklearn.preprocessing import StandardScaler
import matplotlib.pyplot as plt
%matplotlib inline
```
### Data generation
The function below will generate the data points and requires these inputs:
<ul>
<li> <b>centroidLocation</b>: Coordinates of the centroids that will generate the random data. </li>
<ul> <li> Example: input: [[4,3], [2,-1], [-1,4]] </li> </ul>
<li> <b>numSamples</b>: The number of data points we want generated, split over the number of centroids (# of centroids defined in centroidLocation) </li>
<ul> <li> Example: 1500 </li> </ul>
<li> <b>clusterDeviation</b>: The standard deviation between the clusters. The larger the number, the further the spacing. </li>
<ul> <li> Example: 0.5 </li> </ul>
</ul>
```
def createDataPoints(centroidLocation, numSamples, clusterDeviation):
# Create random data and store in feature matrix X and response vector y.
X, y = make_blobs(n_samples=numSamples, centers=centroidLocation,
cluster_std=clusterDeviation)
# Standardize features by removing the mean and scaling to unit variance
X = StandardScaler().fit_transform(X)
return X, y
```
Use <b>createDataPoints</b> with the <b>3 inputs</b> and store the output into variables <b>X</b> and <b>y</b>.
```
X, y = createDataPoints([[4,3], [2,-1], [-1,4]] , 1500, 0.5)
```
### Modeling
DBSCAN stands for Density-Based Spatial Clustering of Applications with Noise. This technique is one of the most common clustering algorithms which works based on density of object.
The whole idea is that if a particular point belongs to a cluster, it should be near to lots of other points in that cluster.
It works based on two parameters: Epsilon and Minimum Points
__Epsilon__ determine a specified radius that if includes enough number of points within, we call it dense area
__minimumSamples__ determine the minimum number of data points we want in a neighborhood to define a cluster.
```
epsilon = 0.3
minimumSamples = 7
db = DBSCAN(eps=epsilon, min_samples=minimumSamples).fit(X)
labels = db.labels_
labels
```
### Distinguish outliers
Lets Replace all elements with 'True' in core_samples_mask that are in the cluster, 'False' if the points are outliers.
```
# Firts, create an array of booleans using the labels from db.
core_samples_mask = np.zeros_like(db.labels_, dtype=bool)
core_samples_mask[db.core_sample_indices_] = True
core_samples_mask
# Number of clusters in labels, ignoring noise if present.
n_clusters_ = len(set(labels)) - (1 if -1 in labels else 0)
n_clusters_
# Remove repetition in labels by turning it into a set.
unique_labels = set(labels)
unique_labels
```
### Data visualization
```
# Create colors for the clusters.
colors = plt.cm.Spectral(np.linspace(0, 1, len(unique_labels)))
# Plot the points with colors
for k, col in zip(unique_labels, colors):
if k == -1:
# Black used for noise.
col = 'k'
class_member_mask = (labels == k)
# Plot the datapoints that are clustered
xy = X[class_member_mask & core_samples_mask]
plt.scatter(xy[:, 0], xy[:, 1],s=50, c=[col], marker=u'o', alpha=0.5)
# Plot the outliers
xy = X[class_member_mask & ~core_samples_mask]
plt.scatter(xy[:, 0], xy[:, 1],s=50, c=[col], marker=u'o', alpha=0.5)
```
## Practice
To better underestand differences between partitional and density-based clusteitng, try to cluster the above dataset into 3 clusters using k-Means.
Notice: do not generate data again, use the same dataset as above.
```
# write your code here
from sklearn.cluster import KMeans
k_means = KMeans(init = "k-means++", n_clusters = 3, n_init=12)
k_means.fit(X)
```
Double-click __here__ for the solution.
<!-- Your answer is below:
from sklearn.cluster import KMeans
k = 3
k_means3 = KMeans(init = "k-means++", n_clusters = k, n_init = 12)
k_means3.fit(X)
fig = plt.figure(figsize=(6, 4))
ax = fig.add_subplot(1, 1, 1)
for k, col in zip(range(k), colors):
my_members = (k_means3.labels_ == k)
plt.scatter(X[my_members, 0], X[my_members, 1], c=col, marker=u'o', alpha=0.5)
plt.show()
-->
<h1 align=center> Weather Station Clustering using DBSCAN & scikit-learn </h1>
<hr>
DBSCAN is specially very good for tasks like class identification on a spatial context. The wonderful attribute of DBSCAN algorithm is that it can find out any arbitrary shape cluster without getting affected by noise. For example, this following example cluster the location of weather stations in Canada.
<Click 1>
DBSCAN can be used here, for instance, to find the group of stations which show the same weather condition. As you can see, it not only finds different arbitrary shaped clusters, can find the denser part of data-centered samples by ignoring less-dense areas or noises.
let's start playing with the data. We will be working according to the following workflow: </font>
1. Loading data
- Overview data
- Data cleaning
- Data selection
- Clusteing
### About the dataset
<h4 align = "center">
Environment Canada
Monthly Values for July - 2015
</h4>
<html>
<head>
<style>
table {
font-family: arial, sans-serif;
border-collapse: collapse;
width: 100%;
}
td, th {
border: 1px solid #dddddd;
text-align: left;
padding: 8px;
}
tr:nth-child(even) {
background-color: #dddddd;
}
</style>
</head>
<body>
<table>
<tr>
<th>Name in the table</th>
<th>Meaning</th>
</tr>
<tr>
<td><font color = "green"><strong>Stn_Name</font></td>
<td><font color = "green"><strong>Station Name</font</td>
</tr>
<tr>
<td><font color = "green"><strong>Lat</font></td>
<td><font color = "green"><strong>Latitude (North+, degrees)</font></td>
</tr>
<tr>
<td><font color = "green"><strong>Long</font></td>
<td><font color = "green"><strong>Longitude (West - , degrees)</font></td>
</tr>
<tr>
<td>Prov</td>
<td>Province</td>
</tr>
<tr>
<td>Tm</td>
<td>Mean Temperature (°C)</td>
</tr>
<tr>
<td>DwTm</td>
<td>Days without Valid Mean Temperature</td>
</tr>
<tr>
<td>D</td>
<td>Mean Temperature difference from Normal (1981-2010) (°C)</td>
</tr>
<tr>
<td><font color = "black">Tx</font></td>
<td><font color = "black">Highest Monthly Maximum Temperature (°C)</font></td>
</tr>
<tr>
<td>DwTx</td>
<td>Days without Valid Maximum Temperature</td>
</tr>
<tr>
<td><font color = "black">Tn</font></td>
<td><font color = "black">Lowest Monthly Minimum Temperature (°C)</font></td>
</tr>
<tr>
<td>DwTn</td>
<td>Days without Valid Minimum Temperature</td>
</tr>
<tr>
<td>S</td>
<td>Snowfall (cm)</td>
</tr>
<tr>
<td>DwS</td>
<td>Days without Valid Snowfall</td>
</tr>
<tr>
<td>S%N</td>
<td>Percent of Normal (1981-2010) Snowfall</td>
</tr>
<tr>
<td><font color = "green"><strong>P</font></td>
<td><font color = "green"><strong>Total Precipitation (mm)</font></td>
</tr>
<tr>
<td>DwP</td>
<td>Days without Valid Precipitation</td>
</tr>
<tr>
<td>P%N</td>
<td>Percent of Normal (1981-2010) Precipitation</td>
</tr>
<tr>
<td>S_G</td>
<td>Snow on the ground at the end of the month (cm)</td>
</tr>
<tr>
<td>Pd</td>
<td>Number of days with Precipitation 1.0 mm or more</td>
</tr>
<tr>
<td>BS</td>
<td>Bright Sunshine (hours)</td>
</tr>
<tr>
<td>DwBS</td>
<td>Days without Valid Bright Sunshine</td>
</tr>
<tr>
<td>BS%</td>
<td>Percent of Normal (1981-2010) Bright Sunshine</td>
</tr>
<tr>
<td>HDD</td>
<td>Degree Days below 18 °C</td>
</tr>
<tr>
<td>CDD</td>
<td>Degree Days above 18 °C</td>
</tr>
<tr>
<td>Stn_No</td>
<td>Climate station identifier (first 3 digits indicate drainage basin, last 4 characters are for sorting alphabetically).</td>
</tr>
<tr>
<td>NA</td>
<td>Not Available</td>
</tr>
</table>
</body>
</html>
### 1-Download data
To download the data, we will use **`!wget`**. To download the data, we will use `!wget` to download it from IBM Object Storage.
__Did you know?__ When it comes to Machine Learning, you will likely be working with large datasets. As a business, where can you host your data? IBM is offering a unique opportunity for businesses, with 10 Tb of IBM Cloud Object Storage: [Sign up now for free](http://cocl.us/ML0101EN-IBM-Offer-CC)
```
!wget -O weather-stations20140101-20141231.csv https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/ML0101ENv3/labs/weather-stations20140101-20141231.csv
```
### 2- Load the dataset
We will import the .csv then we creates the columns for year, month and day.
```
import csv
import pandas as pd
import numpy as np
filename='weather-stations20140101-20141231.csv'
#Read csv
pdf = pd.read_csv(filename)
pdf.head(5)
```
### 3-Cleaning
Lets remove rows that dont have any value in the __Tm__ field.
```
pdf = pdf[pd.notnull(pdf["Tm"])]
pdf = pdf.reset_index(drop=True)
pdf.head(5)
```
### 4-Visualization
Visualization of stations on map using basemap package. The matplotlib basemap toolkit is a library for plotting 2D data on maps in Python. Basemap does not do any plotting on it’s own, but provides the facilities to transform coordinates to a map projections.
Please notice that the size of each data points represents the average of maximum temperature for each station in a year.
```
from mpl_toolkits.basemap import Basemap
import matplotlib.pyplot as plt
from pylab import rcParams
%matplotlib inline
rcParams['figure.figsize'] = (14,10)
llon=-140
ulon=-50
llat=40
ulat=65
pdf = pdf[(pdf['Long'] > llon) & (pdf['Long'] < ulon) & (pdf['Lat'] > llat) &(pdf['Lat'] < ulat)]
my_map = Basemap(projection='merc',
resolution = 'l', area_thresh = 1000.0,
llcrnrlon=llon, llcrnrlat=llat, #min longitude (llcrnrlon) and latitude (llcrnrlat)
urcrnrlon=ulon, urcrnrlat=ulat) #max longitude (urcrnrlon) and latitude (urcrnrlat)
my_map.drawcoastlines()
my_map.drawcountries()
# my_map.drawmapboundary()
my_map.fillcontinents(color = 'white', alpha = 0.3)
my_map.shadedrelief()
# To collect data based on stations
xs,ys = my_map(np.asarray(pdf.Long), np.asarray(pdf.Lat))
pdf['xm']= xs.tolist()
pdf['ym'] =ys.tolist()
#Visualization1
for index,row in pdf.iterrows():
# x,y = my_map(row.Long, row.Lat)
my_map.plot(row.xm, row.ym,markerfacecolor =([1,0,0]), marker='o', markersize= 5, alpha = 0.75)
#plt.text(x,y,stn)
plt.show()
```
### 5- Clustering of stations based on their location i.e. Lat & Lon
__DBSCAN__ form sklearn library can runs DBSCAN clustering from vector array or distance matrix. In our case, we pass it the Numpy array Clus_dataSet to find core samples of high density and expands clusters from them.
```
from sklearn.cluster import DBSCAN
import sklearn.utils
from sklearn.preprocessing import StandardScaler
sklearn.utils.check_random_state(1000)
Clus_dataSet = pdf[['xm','ym']]
Clus_dataSet = np.nan_to_num(Clus_dataSet)
Clus_dataSet = StandardScaler().fit_transform(Clus_dataSet)
# Compute DBSCAN
db = DBSCAN(eps=0.15, min_samples=10).fit(Clus_dataSet)
core_samples_mask = np.zeros_like(db.labels_, dtype=bool)
core_samples_mask[db.core_sample_indices_] = True
labels = db.labels_
pdf["Clus_Db"]=labels
realClusterNum=len(set(labels)) - (1 if -1 in labels else 0)
clusterNum = len(set(labels))
# A sample of clusters
pdf[["Stn_Name","Tx","Tm","Clus_Db"]].head(5)
```
As you can see for outliers, the cluster label is -1
```
set(labels)
```
### 6- Visualization of clusters based on location
Now, we can visualize the clusters using basemap:
```
from mpl_toolkits.basemap import Basemap
import matplotlib.pyplot as plt
from pylab import rcParams
%matplotlib inline
rcParams['figure.figsize'] = (14,10)
my_map = Basemap(projection='merc',
resolution = 'l', area_thresh = 1000.0,
llcrnrlon=llon, llcrnrlat=llat, #min longitude (llcrnrlon) and latitude (llcrnrlat)
urcrnrlon=ulon, urcrnrlat=ulat) #max longitude (urcrnrlon) and latitude (urcrnrlat)
my_map.drawcoastlines()
my_map.drawcountries()
#my_map.drawmapboundary()
my_map.fillcontinents(color = 'white', alpha = 0.3)
my_map.shadedrelief()
# To create a color map
colors = plt.get_cmap('jet')(np.linspace(0.0, 1.0, clusterNum))
#Visualization1
for clust_number in set(labels):
c=(([0.4,0.4,0.4]) if clust_number == -1 else colors[np.int(clust_number)])
clust_set = pdf[pdf.Clus_Db == clust_number]
my_map.scatter(clust_set.xm, clust_set.ym, color =c, marker='o', s= 20, alpha = 0.85)
if clust_number != -1:
cenx=np.mean(clust_set.xm)
ceny=np.mean(clust_set.ym)
plt.text(cenx,ceny,str(clust_number), fontsize=25, color='red',)
print ("Cluster "+str(clust_number)+', Avg Temp: '+ str(np.mean(clust_set.Tm)))
```
### 7- Clustering of stations based on their location, mean, max, and min Temperature
In this section we re-run DBSCAN, but this time on a 5-dimensional dataset:
```
from sklearn.cluster import DBSCAN
import sklearn.utils
from sklearn.preprocessing import StandardScaler
sklearn.utils.check_random_state(1000)
Clus_dataSet = pdf[['xm','ym','Tx','Tm','Tn']]
Clus_dataSet = np.nan_to_num(Clus_dataSet)
Clus_dataSet = StandardScaler().fit_transform(Clus_dataSet)
# Compute DBSCAN
db = DBSCAN(eps=0.3, min_samples=10).fit(Clus_dataSet)
core_samples_mask = np.zeros_like(db.labels_, dtype=bool)
core_samples_mask[db.core_sample_indices_] = True
labels = db.labels_
pdf["Clus_Db"]=labels
realClusterNum=len(set(labels)) - (1 if -1 in labels else 0)
clusterNum = len(set(labels))
# A sample of clusters
pdf[["Stn_Name","Tx","Tm","Clus_Db"]].head(5)
```
### 8- Visualization of clusters based on location and Temperture
```
from mpl_toolkits.basemap import Basemap
import matplotlib.pyplot as plt
from pylab import rcParams
%matplotlib inline
rcParams['figure.figsize'] = (14,10)
my_map = Basemap(projection='merc',
resolution = 'l', area_thresh = 1000.0,
llcrnrlon=llon, llcrnrlat=llat, #min longitude (llcrnrlon) and latitude (llcrnrlat)
urcrnrlon=ulon, urcrnrlat=ulat) #max longitude (urcrnrlon) and latitude (urcrnrlat)
my_map.drawcoastlines()
my_map.drawcountries()
#my_map.drawmapboundary()
my_map.fillcontinents(color = 'white', alpha = 0.3)
my_map.shadedrelief()
# To create a color map
colors = plt.get_cmap('jet')(np.linspace(0.0, 1.0, clusterNum))
#Visualization1
for clust_number in set(labels):
c=(([0.4,0.4,0.4]) if clust_number == -1 else colors[np.int(clust_number)])
clust_set = pdf[pdf.Clus_Db == clust_number]
my_map.scatter(clust_set.xm, clust_set.ym, color =c, marker='o', s= 20, alpha = 0.85)
if clust_number != -1:
cenx=np.mean(clust_set.xm)
ceny=np.mean(clust_set.ym)
plt.text(cenx,ceny,str(clust_number), fontsize=25, color='red',)
print ("Cluster "+str(clust_number)+', Avg Temp: '+ str(np.mean(clust_set.Tm)))
```
## Want to learn more?
IBM SPSS Modeler is a comprehensive analytics platform that has many machine learning algorithms. It has been designed to bring predictive intelligence to decisions made by individuals, by groups, by systems – by your enterprise as a whole. A free trial is available through this course, available here: [SPSS Modeler](http://cocl.us/ML0101EN-SPSSModeler).
Also, you can use Watson Studio to run these notebooks faster with bigger datasets. Watson Studio is IBM's leading cloud solution for data scientists, built by data scientists. With Jupyter notebooks, RStudio, Apache Spark and popular libraries pre-packaged in the cloud, Watson Studio enables data scientists to collaborate on their projects without having to install anything. Join the fast-growing community of Watson Studio users today with a free account at [Watson Studio](https://cocl.us/ML0101EN_DSX)
### Thanks for completing this lesson!
Notebook created by: <a href = "https://ca.linkedin.com/in/saeedaghabozorgi">Saeed Aghabozorgi</a>
<hr>
Copyright © 2018 [Cognitive Class](https://cocl.us/DX0108EN_CC). This notebook and its source code are released under the terms of the [MIT License](https://bigdatauniversity.com/mit-license/).
| github_jupyter |
```
import numpy as np
from matplotlib import pyplot as plt
import real_space_electrostatic_sum
```
# Benchmarking
Using the shared library `real-space-electrostatic-sum.so`, together with the Python wrapper in `real_space_electrostatic_sum.py`, this notebook reproduces results from [Pickard, *Phys. Rev. Mat.* **2**, 013806 (2018)](https://doi.org/10.1103/PhysRevMaterials.2.013806).
## Reproducing part of Fig. 1(b)
Fig. 1(b) in the paper demonstrates the convergence of the real-space method for a simple cubic lattice with unit spacing.
```
# lattice vectors
a_1 = np.array([1.0, 0.0, 0.0])
a_2 = np.array([0.0, 1.0, 0.0])
a_3 = np.array([0.0, 0.0, 1.0])
# ion locations and charge array
loc = np.zeros([1,3])
chg = np.ones(1)
# loop over cutoff radii
r_c = np.linspace(0.001,30,500)
r_d = 1.5
ene = np.zeros(len(r_c))
for i, r in enumerate(r_c):
ene[i] = real_space_electrostatic_sum.energy(
a_1, a_2, a_3, loc.shape[0], loc[:,0], loc[:,1], loc[:,2], chg, r, r_d)
# generate part of Fig. 1(b)
plt.plot(r_c, ene, 'r')
plt.title('Fig. 1(b)')
plt.xlim([0,30]); plt.ylim([-1.5,-1.25])
plt.xlabel('$R_c$'); plt.ylabel('$E_i$')
plt.show()
```
## Reproducing data in Table I
Table I in the paper contains ion-ion energies for four crystals obtained with the real-space method, as well as the Ewald method. The real-space method data are re-generated here, exhibiting near perfect agreement with Table I. The Ewald energies reported below were re-obtained with CASTEP using the exact lattice parameters employed here.
### _Al_
```
# lattice vectors
a_1 = np.array([5.41141973394663, 0.00000000000000, 0.00000000000000])
a_2 = np.array([2.70570986697332, 4.68642696013821, 0.00000000000000])
a_3 = np.array([2.70570986697332, 1.56214232004608, 4.41840571073226])
# ion locations
loc = np.zeros([1,3])
# charge array
chg = 3.0 * np.ones(loc.shape[0])
# length scale
h_max = 4.42
# reference energy
ewald = -2.69595457432924945
print('Ewald: energy = {0:12.9f}'.format(-2.69595457432924945))
# real-space-method energies
r_d_hat = 2.0
ene = real_space_electrostatic_sum.energy(
a_1, a_2, a_3, loc.shape[0], loc[:,0], loc[:,1], loc[:,2], chg, 3.0*r_d_hat**2*h_max, r_d_hat*h_max)
print('R\u0302d = {0:3.1f}: energy = {1:12.9f}'.format(r_d_hat, ene))
r_d_hat = 1.5
ene = real_space_electrostatic_sum.energy(
a_1, a_2, a_3, loc.shape[0], loc[:,0], loc[:,1], loc[:,2], chg, 3.0*r_d_hat**2*h_max, r_d_hat*h_max)
print('R\u0302d = {0:3.1f}: energy = {1:12.9f}'.format(r_d_hat, ene))
r_d_hat = 1.0
ene = real_space_electrostatic_sum.energy(
a_1, a_2, a_3, loc.shape[0], loc[:,0], loc[:,1], loc[:,2], chg, 3.0*r_d_hat**2*h_max, r_d_hat*h_max)
print('R\u0302d = {0:3.1f}: energy = {1:12.9f}'.format(r_d_hat, ene))
```
### _Si_
```
# lattice vectors
a_1 = np.array([7.25654832321381, 0.00000000000000, 0.00000000000000])
a_2 = np.array([3.62827416160690, 6.28435519169252, 0.00000000000000])
a_3 = np.array([3.62827416160690, 2.09478506389751, 5.92494689524090])
# ion locations
loc = np.array([[0.0, 0.0, 0.0],
[0.25, 0.25, 0.25]])
loc = (np.vstack((a_1, a_2, a_3)).T).dot(loc.T).T # convert to cartesian
# charge array
chg = 4.0 * np.ones(loc.shape[0])
# length scale
h_max = 5.92
# reference energy
ewald = -8.39857465282205418
print('Ewald: energy = {0:12.9f}, per ion = {1:12.9f}'.format(ewald, ewald/loc.shape[0]))
# real-space-method energies
r_d_hat = 2.0
ene = real_space_electrostatic_sum.energy(
a_1, a_2, a_3, loc.shape[0], loc[:,0], loc[:,1], loc[:,2], chg, 3.0*r_d_hat**2*h_max, r_d_hat*h_max)
print('R\u0302d = {0:3.1f}: energy = {1:12.9f}, per ion = {2:12.9f}'.format(r_d_hat, ene, ene/loc.shape[0]))
r_d_hat = 1.5
ene = real_space_electrostatic_sum.energy(
a_1, a_2, a_3, loc.shape[0], loc[:,0], loc[:,1], loc[:,2], chg, 3.0*r_d_hat**2*h_max, r_d_hat*h_max)
print('R\u0302d = {0:3.1f}: energy = {1:12.9f}, per ion = {2:12.9f}'.format(r_d_hat, ene, ene/loc.shape[0]))
r_d_hat = 1.0
ene = real_space_electrostatic_sum.energy(
a_1, a_2, a_3, loc.shape[0], loc[:,0], loc[:,1], loc[:,2], chg, 3.0*r_d_hat**2*h_max, r_d_hat*h_max)
print('R\u0302d = {0:3.1f}: energy = {1:12.9f}, per ion = {2:12.9f}'.format(r_d_hat, ene, ene/loc.shape[0]))
```
### _SiO2_
```
# lattice vectors
a_1 = np.array([ 9.28422445623683, 0.00000000000000, 0.00000000000000])
a_2 = np.array([-4.64211222811842, 8.04037423353787, 0.00000000000000])
a_3 = np.array([ 0.00000000000000, 0.00000000000000, 10.2139697101486])
# ion locations
loc = np.array([[0.41500, 0.27200, 0.21300],
[0.72800, 0.14300, 0.54633],
[0.85700, 0.58500, 0.87967],
[0.27200, 0.41500, 0.78700],
[0.14300, 0.72800, 0.45367],
[0.58500, 0.85700, 0.12033],
[0.46500, 0.00000, 0.33333],
[0.00000, 0.46500, 0.66667],
[0.53500, 0.53500, 0.00000]])
loc = (np.vstack((a_1, a_2, a_3)).T).dot(loc.T).T # convert to cartesian
# charge array
chg = 6.0 * np.ones(loc.shape[0]) # most are O
chg[6:] = 4.0 # three are Si
# length scale
h_max = 10.21
# reference energy
ewald = -69.48809871723248932
print('Ewald: energy = {0:12.9f}, per ion = {1:12.9f}'.format(ewald, ewald/loc.shape[0]))
# real-space-method energies
r_d_hat = 2.0
ene = real_space_electrostatic_sum.energy(
a_1, a_2, a_3, loc.shape[0], loc[:,0], loc[:,1], loc[:,2], chg, 3.0*r_d_hat**2*h_max, r_d_hat*h_max)
print('R\u0302d = {0:3.1f}: energy = {1:12.9f}, per ion = {2:12.9f}'.format(r_d_hat, ene, ene/loc.shape[0]))
r_d_hat = 1.5
ene = real_space_electrostatic_sum.energy(
a_1, a_2, a_3, loc.shape[0], loc[:,0], loc[:,1], loc[:,2], chg, 3.0*r_d_hat**2*h_max, r_d_hat*h_max)
print('R\u0302d = {0:3.1f}: energy = {1:12.9f}, per ion = {2:12.9f}'.format(r_d_hat, ene, ene/loc.shape[0]))
r_d_hat = 1.0
ene = real_space_electrostatic_sum.energy(
a_1, a_2, a_3, loc.shape[0], loc[:,0], loc[:,1], loc[:,2], chg, 3.0*r_d_hat**2*h_max, r_d_hat*h_max)
print('R\u0302d = {0:3.1f}: energy = {1:12.9f}, per ion = {2:12.9f}'.format(r_d_hat, ene, ene/loc.shape[0]))
```
### _Al2SiO5_
```
# lattice vectors
a_1 = np.array([14.7289033699982, 0.00000000000000, 0.00000000000000])
a_2 = np.array([0.00000000000000, 14.9260018049230, 0.00000000000000])
a_3 = np.array([0.00000000000000, 0.00000000000000, 10.5049875335275])
# ion locations
loc = np.array([[0.23030, 0.13430, 0.23900],
[0.76970, 0.86570, 0.23900],
[0.26970, 0.63430, 0.26100],
[0.73030, 0.36570, 0.26100],
[0.76970, 0.86570, 0.76100],
[0.23030, 0.13430, 0.76100],
[0.73030, 0.36570, 0.73900],
[0.26970, 0.63430, 0.73900],
[0.00000, 0.00000, 0.24220],
[0.50000, 0.50000, 0.25780],
[0.00000, 0.00000, 0.75780],
[0.50000, 0.50000, 0.74220],
[0.37080, 0.13870, 0.50000],
[0.42320, 0.36270, 0.50000],
[0.62920, 0.86130, 0.50000],
[0.57680, 0.63730, 0.50000],
[0.12920, 0.63870, 0.00000],
[0.07680, 0.86270, 0.00000],
[0.87080, 0.36130, 0.00000],
[0.92320, 0.13730, 0.00000],
[0.24620, 0.25290, 0.00000],
[0.42400, 0.36290, 0.00000],
[0.10380, 0.40130, 0.00000],
[0.75380, 0.74710, 0.00000],
[0.57600, 0.63710, 0.00000],
[0.89620, 0.59870, 0.00000],
[0.25380, 0.75290, 0.50000],
[0.07600, 0.86290, 0.50000],
[0.39620, 0.90130, 0.50000],
[0.74620, 0.24710, 0.50000],
[0.92400, 0.13710, 0.50000],
[0.60380, 0.09870, 0.50000]])
loc = (np.vstack((a_1, a_2, a_3)).T).dot(loc.T).T # convert to cartesian
# charge array
chg = 6.0 * np.ones(loc.shape[0]) # most are O
chg[8:13] = 3.0 # eight are Al
chg[14] = 3.0
chg[16] = 3.0
chg[18] = 3.0
chg[20] = 4.0 # four are Si
chg[23] = 4.0
chg[26] = 4.0
chg[29] = 4.0
# length scale
h_max = 14.93
# reference energy
ewald = -244.05500850908111943
print('Ewald: energy = {0:14.9f}, per ion = {1:12.9f}'.format(ewald, ewald/loc.shape[0]))
# real-space-method energies
r_d_hat = 2.0
ene = real_space_electrostatic_sum.energy(
a_1, a_2, a_3, loc.shape[0], loc[:,0], loc[:,1], loc[:,2], chg, 3.0*r_d_hat**2*h_max, r_d_hat*h_max)
print('R\u0302d = {0:3.1f}: energy = {1:14.9f}, per ion = {2:12.9f}'.format(r_d_hat, ene, ene/loc.shape[0]))
r_d_hat = 1.5
ene = real_space_electrostatic_sum.energy(
a_1, a_2, a_3, loc.shape[0], loc[:,0], loc[:,1], loc[:,2], chg, 3.0*r_d_hat**2*h_max, r_d_hat*h_max)
print('R\u0302d = {0:3.1f}: energy = {1:14.9f}, per ion = {2:12.9f}'.format(r_d_hat, ene, ene/loc.shape[0]))
r_d_hat = 1.0
ene = real_space_electrostatic_sum.energy(
a_1, a_2, a_3, loc.shape[0], loc[:,0], loc[:,1], loc[:,2], chg, 3.0*r_d_hat**2*h_max, r_d_hat*h_max)
print('R\u0302d = {0:3.1f}: energy = {1:14.9f}, per ion = {2:12.9f}'.format(r_d_hat, ene, ene/loc.shape[0]))
```
## Madelung energy for NaCl
See the discussion around Eq. (1) in [Mamode, _J. Math. Chem._ __55__, 734 (2017)](https://doi.org/10.1007/s10910-016-0705-9).
With $M_{\mathrm{NaCl}}$ as the Madelung energy and $E_{\mathrm{NaCl}}$ as the energy of a two-atom primitive cell having $z_{1,2}=\pm 1$, the following identities hold
\begin{equation}
\begin{split}
M_{\mathrm{NaCl}}
&= E_{\mathrm{NaCl}} \\
&= \sum_{i\in\{1,2\}} \sum_{j\ne i}^\infty \frac{z_i z_j}{2r_{ij}} \\
&= \sum_{i\in\{1,2\}}
\left[ \sum_{\substack{j\ne i \\z_iz_j>0}}^\infty \frac{z_i z_j}{2r_{ij}} -
\sum_{\substack{j\ne i \\z_iz_j<0}}^\infty \frac{|z_i z_j|}{2r_{ij}} \right] \\
&= \sum_{i\in\{1,2\}}
\left[ 2 \sum_{\substack{j\ne i \\z_iz_j>0}}^\infty \frac{z_i z_j}{2r_{ij}} -
\sum_{j\ne i}^\infty \frac{|z_i z_j|}{2r_{ij}} \right] \\
&= 4 E_{\mathrm{FCC}} -
\sum_{i\in\{1,2\}} \sum_{j\ne i}^\infty \frac{|z_i z_j|}{2r_{ij}}
\end{split}
\end{equation}
and the final result should be $M_{\mathrm{NaCl}} = −1.747 564 594 633 \cdots$.
```
# lattice vectors
a_1 = np.array([1.0, 1.0, 0.0])
a_2 = np.array([0.0, 1.0, 1.0])
a_3 = np.array([1.0, 0.0, 1.0])
# length scale and cutoff
h_max = np.sqrt(4.0/3.0)
r_d_hat = 3.0
# compute FCC energy
loc = np.zeros([1,3])
chg = np.ones(loc.shape[0])
E_FCC = real_space_electrostatic_sum.energy(
a_1, a_2, a_3, loc.shape[0], loc[:,0], loc[:,1], loc[:,2], chg, 3.0*r_d_hat**2*h_max, r_d_hat*h_max)
# compute second term
loc = np.zeros([2,3])
loc[1,:] = np.array([1.0, 1.0, 1.0])
chg = np.ones(loc.shape[0])
E_2 = real_space_electrostatic_sum.energy(
a_1, a_2, a_3, loc.shape[0], loc[:,0], loc[:,1], loc[:,2], chg, 3.0*r_d_hat**2*h_max, r_d_hat*h_max)
# print result
print('M = {0:15.12f}'.format(4*E_FCC - E_2))
```
| github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#Keras-Basics" data-toc-modified-id="Keras-Basics-1"><span class="toc-item-num">1 </span>Keras Basics</a></span><ul class="toc-item"><li><span><a href="#Saving-and-loading-the-models" data-toc-modified-id="Saving-and-loading-the-models-1.1"><span class="toc-item-num">1.1 </span>Saving and loading the models</a></span></li></ul></li><li><span><a href="#Reference" data-toc-modified-id="Reference-2"><span class="toc-item-num">2 </span>Reference</a></span></li></ul></div>
```
# code for loading the format for the notebook
import os
# path : store the current path to convert back to it later
path = os.getcwd()
os.chdir(os.path.join('..', 'notebook_format'))
from formats import load_style
load_style(plot_style=False)
os.chdir(path)
# 1. magic to print version
# 2. magic so that the notebook will reload external python modules
%load_ext watermark
%load_ext autoreload
%autoreload 2
import numpy as np
import pandas as pd
from keras.datasets import mnist
from keras.utils import np_utils
from keras.optimizers import RMSprop
from keras.models import Sequential, load_model
from keras.layers.core import Dense, Dropout, Activation
%watermark -a 'Ethen' -d -t -v -p numpy,pandas,keras
```
# Keras Basics
Basic Keras API to build a simple multi-layer neural network.
```
n_classes = 10
n_features = 784 # mnist is a 28 * 28 image
# load the dataset and some preprocessing step that can be skipped
(X_train, y_train), (X_test, y_test) = mnist.load_data()
X_train = X_train.reshape(60000, n_features)
X_test = X_test.reshape(10000, n_features)
X_train = X_train.astype('float32')
X_test = X_test.astype('float32')
# images takes values between 0 - 255, we can normalize it
# by dividing every number by 255
X_train /= 255
X_test /= 255
print(X_train.shape[0], 'train samples')
print(X_test.shape[0], 'test samples')
# convert class vectors to binary class matrices (one-hot encoding)
# note: you HAVE to to this step
Y_train = np_utils.to_categorical(y_train, n_classes)
Y_test = np_utils.to_categorical(y_test , n_classes)
```
Basics of training a model:
The easiest way to build models in keras is to use `Sequential` model and the `.add()` method to stack layers together in sequence to build up our network.
- We start with `Dense` (fully-connected layers), where we specify how many nodes you wish to have for the layer. Since the first layer that we're going to add is the input layer, we have to make sure that the `input_dim` parameter matches the number of features (columns) in the training set. Then after the first layer, we don't need to specify the size of the input anymore.
- Then we specify the `Activation` function for that layer, and add a `Dropout` layer if we wish.
- For the last `Dense` and `Activation` layer we need to specify the number of class as the output and softmax to tell it to output the predicted class's probability.
```
# define the model
model = Sequential()
model.add(Dense(512, input_dim = n_features))
model.add(Activation('relu'))
model.add(Dropout(0.2))
model.add(Dense(512))
model.add(Activation('relu'))
model.add(Dropout(0.2))
model.add(Dense(n_classes))
model.add(Activation('softmax'))
# we can check the summary to check the number of parameters
model.summary()
```
Once our model looks good, we can configure its learning process with `.compile()`, where you need to specify which `optimizer` to use, and the `loss` function ( `categorical_crossentropy` is the typical one for multi-class classification) and the `metrics` to track.
Finally, `.fit()` the model by passing in the training, validation set, the number of epochs and batch size. For the batch size, we typically specify this number to be power of 2 for computing efficiency.
```
model.compile(loss = 'categorical_crossentropy', optimizer = RMSprop(), metrics = ['accuracy'])
n_epochs = 10
batch_size = 128
history = model.fit(
X_train,
Y_train,
batch_size = batch_size,
epochs = n_epochs,
verbose = 1, # set it to 0 if we don't want to have progess bars
validation_data = (X_test, Y_test)
)
# history attribute stores the training and validation score and loss
history.history
# .evaluate gives the loss and metric evaluation score for the dataset,
# here the result matches the validation set's history above
print('metrics: ', model.metrics_names)
score = model.evaluate(X_test, Y_test, verbose = 0)
score
# stores the weight of the model,
# it's a list, note that the length is 6 because we have 3 dense layer
# and each one has it's associated bias term
weights = model.get_weights()
print(len(weights))
# W1 should have 784, 512 for the 784
# feauture column and the 512 the number
# of dense nodes that we've specified
W1, b1, W2, b2, W3, b3 = weights
print(W1.shape)
print(b1.shape)
# predict the accuracy
y_pred = model.predict_classes(X_test, verbose = 0)
accuracy = np.sum(y_test == y_pred) / X_test.shape[0]
print('valid accuracy: %.2f' % (accuracy * 100))
```
## Saving and loading the models
It is not recommended to use pickle or cPickle to save a Keras model. By saving it as a HDF5 file, we can preserve the configuration and weights of the model.
```
model.save('my_model.h5') # creates a HDF5 file 'my_model.h5'
del model # deletes the existing model
# returns a compiled model
# identical to the previous one
model = load_model('my_model.h5')
# testing: predict the accuracy using the loaded model
y_pred = model.predict_classes(X_test, verbose = 0)
accuracy = np.sum(y_test == y_pred) / X_test.shape[0]
print('valid accuracy: %.2f' % (accuracy * 100))
```
# Reference
- [Keras Documentation](http://keras.io/)
- [Keras Documentation: mnist_mlp example](https://github.com/fchollet/keras/blob/master/examples/mnist_mlp.py)
- [Keras Documentation: Saving Keras Model](http://keras.io/getting-started/faq/#how-can-i-save-a-keras-model)
| github_jupyter |
#Parkinsons Disease
### Importing Libraries
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
pip install catboost
d=pd.read_csv("parkinsons.data")
x=d.loc[:,d.columns != 'status'].values[:,1:]
y=d.loc[:,'status'].values
d.head()
print(x)
print(y)
d.describe()
d.info()
d.nunique()
```
### Counting Number of people with the disease and without the disease
```
z=0
o=0
zero=[]
one=[]
for i in range (0,len(y)):
if y[i] == 0:
zero.append(y[i])
z=z+1
else:
one.append(y[i])
o=o+1
print("Number of rows with status 0 : ", z)
print("Number of rows with status 1 : ", o)
```
### Splitting the Data
```
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(x, y, test_size = 0.3, random_state = 0)
```
### Min Max Scaler
```
from sklearn.preprocessing import MinMaxScaler
scaler=MinMaxScaler((-1,1))
x1=scaler.fit_transform(x)
y1=y
```
### XG Boost
```
from xgboost import XGBClassifier
xg = XGBClassifier()
xg.fit(X_train,y_train)
y_pred=xg.predict(X_test)
```
#### Scoring the XG Boost
```
from sklearn.metrics import r2_score
r2_xg = r2_score(y_test,y_pred)
print(r2_xg)
from sklearn.metrics import confusion_matrix
cm_xg = confusion_matrix(y_test,y_pred)
print(cm_xg)
from sklearn.metrics import accuracy_score
ac_xg=accuracy_score(y_test,y_pred)
print(ac_xg)
```
### CatBoost
```
from catboost import CatBoostClassifier
cbc = CatBoostClassifier()
cbc.fit(X_train,y_train)
y_pred=cbc.predict(X_test)
```
#### Scoring the Catboost
```
from sklearn.metrics import r2_score
r2_cb=r2_score(y_test,y_pred)
print(r2_cb)
from sklearn.metrics import confusion_matrix
cm_cb = confusion_matrix(y_test,y_pred)
print(cm_cb)
from sklearn.metrics import accuracy_score
ac_cb = accuracy_score(y_test,y_pred)
print(ac_cb)
```
### LightGbm
```
from lightgbm import LGBMClassifier
lgb = LGBMClassifier ()
lgb.fit(X_train,y_train)
y_pred =lgb.predict(X_test)
```
#### Scoring LightGBM
```
from sklearn.metrics import r2_score
r2_lgb=r2_score(y_test,y_pred)
print(r2_lgb)
from sklearn.metrics import confusion_matrix
cm_lgb = confusion_matrix(y_test,y_pred)
print(cm_lgb)
from sklearn.metrics import accuracy_score
ac_lgb = accuracy_score(y_test,y_pred)
print(ac_lgb)
```
### Comparing the 3 methods
#### r2 scores
```
label = ['XG Boost','Cat Boost','LightGBM Boost']
r2 = [r2_xg,r2_cb,r2_lgb]
for i in range(0,len(r2)):
r2[i]=np.round(r2[i]*100,decimals=3)
print(r2)
plt.figure(figsize=(14,8))
p=sns.barplot(y=r2,x=label)
plt.title("Comparing different Boosting techniques",fontweight="bold")
for z in p.patches:
width, height = z.get_width(), z.get_height()
x, y = z.get_xy()
p.annotate('{:.3f}%'.format(height), (x +0.25, y + height + 0.8))
plt.show()
```
#### accuracy scores
```
ac = [ac_xg,ac_cb,ac_lgb]
for i in range(0,len(ac)):
ac[i]=np.round(ac[i]*100,decimals=3)
print(ac)
plt.figure(figsize=(14,8))
p=sns.barplot(y=ac,x=label)
plt.title("Comparing different Boosting techniques",fontweight="bold")
for z in p.patches:
width, height = z.get_width(), z.get_height()
x, y = z.get_xy()
p.annotate('{:.3f}%'.format(height), (x +0.25, y + height + 0.8))
plt.show()
```
| github_jupyter |
# Getting stared with FastAI v1: Tabular data
This notebook illustrates how to get started with FastAI v1 for Tabular Data. The notebook will focus on introducing the API to newcomers, with an overview, including installation instructions for FastAI v1, given at: https://www.avanwyk.com/getting-started-with-fastai-v1-tabular-data/.
The details of the deep learning techniques used (such as embedding, learning rate cycles etc.) are not covered in this notebook. For a detailed description of the techniques used, have a look at this blog post: [http://www.fast.ai/2018/04/29/categorical-embeddings/](http://www.fast.ai/2018/04/29/categorical-embeddings/).
FastAI provides a convenient way of importing a lot of commonly used modules, including Numpy and Pandas:
```
from fastai import *
from fastai.tabular import *
from sklearn.model_selection import train_test_split
np.set_printoptions(precision=4, suppress=True)
pd.set_option('display.float_format', lambda x: '%.3f' % x)
```
## Data Loading
Before getting to the FastAI model, we first have to prepare our data. The dataset can be download here: https://www.kaggle.com/cdc/mortality. Download the data and place it into a directory `data`. We will be attempting to predict the age of mortaility using basic demographic data. The Kaggle data is separated into CSV files by year. We will limit the data to two year's worth, simply for performance reasons. We load the yearly data in turn and then create a single dataframe containing all the data.
```
data_path = Path('data')
frames = []
for year in range(2005, 2007):
frames.append(pd.read_csv(data_path/f'{year}_data.csv'))
df = pd.concat(frames)
```
The data contains 77 different columns (info into the meaning of the fields can be found [here](https://www.cdc.gov/nchs/nvss/mortality_public_use_data.htm)), however, we are only interested in the demographic data for this experiment, so we discard the other columns.
```
demographic_colums = ['resident_status', 'education_1989_revision', 'education_2003_revision', 'education_reporting_flag', 'sex',
'detail_age_type', 'detail_age', 'marital_status', 'race']
df = df.filter(demographic_colums)
df.sample(5)
```
### Data Pre-processing
The educational level is reported based on one of two systems: the 1989 revision and the 2003 revision. Here we normalize the education level to a single standard. We translate both revision's data to new categories which roughly groups similar categories from the other revision systems.
category | description (highest level of education)
--- | ---
P | some primary school or no education
H | some high school education
T | some tertiary education
U | unknown
```
def normalize_education(frame):
frame['education'] = 'U'
filter_1989 = frame['education_reporting_flag'] == 0
frame.loc[filter_1989 & (frame.education_1989_revision <= 8.0), 'education'] = 'P'
frame.loc[filter_1989 & (frame.education_1989_revision > 8.0) & (frame.education_1989_revision <= 12.0), 'education'] = 'H'
frame.loc[filter_1989 & (frame.education_1989_revision > 12.0) & (frame.education_1989_revision < 99.0), 'education'] = 'T'
frame.loc[filter_1989 & ((frame.education_1989_revision >= 99.0) | (frame.education_1989_revision.isna())), 'education'] = 'U'
filter_2003 = frame['education_reporting_flag'] == 1
frame.loc[filter_2003 & (frame.education_2003_revision <= 1), 'education'] = 'P'
frame.loc[filter_2003 & (frame.education_2003_revision > 1) & (frame.education_2003_revision <= 3.0), 'education'] = 'H'
frame.loc[filter_2003 & (frame.education_2003_revision > 3) & (frame.education_2003_revision < 9), 'education'] = 'T'
frame.loc[filter_2003 & ((frame.education_2003_revision == 9) | (frame.education_2003_revision.isna())), 'education'] = 'U'
return frame
```
Similarly we have to normalize the age: we treat any death before 12 months as 1 year.
```
def normalize_age(frame):
frame['age'] = np.nan
less_than_12_months = (frame['detail_age_type'] > 1) & (frame['detail_age_type'] < 9)
age_in_years = frame['detail_age_type'] == 1
unknown_age = frame['detail_age_type'] == 9
frame.loc[less_than_12_months, 'age'] = 1
frame.loc[age_in_years, 'age'] = frame[age_in_years].detail_age
frame.loc[unknown_age, 'age'] = np.nan
return frame
df = normalize_education(df)
df = normalize_age(df)
df.describe()
```
We can see in the dataframe description that some of the age values are missing. For the sake of simplity we can drop those rows.
```
df['age'] = df['age'].astype(np.float32)
df = df.dropna(subset=['age'])
df.sample(5)
```
## FastAI
Our data is now ready for us to train the model. We won't be using all the columns (as some columns are simply flags indicating the type of another column). So we discard all the columns we don't need. We also split or training data into a training and validation dataset.
```
df_fastai = df[['resident_status', 'education', 'sex', 'marital_status', 'race', 'age']]
df_fastai.sample(5)
train_df, valid_df = train_test_split(df_fastai, test_size=0.33)
```
The FastAI API requires us to define the dependent variable, which is the variable we are trying to predict. This is `age` in our case. Further, we need to specify the categorical columns by name such that FastAI can automatically handle them for us (more on this below).
```
dep_var = 'age'
categorical_names = ['education', 'sex', 'marital_status']
tfms = [FillMissing, Categorify]
```
Notice how we also define to transformations: `FillMissing` and `Categorify`.
The `FillMissing` transform will fill in any missing values for the _continuous_ variable columns we have using the median to do so (by default). Read more about the transformer [here](http://docs.fast.ai/tabular.transform.html#class-fillmissing).
The `Categorify` transformation will take categorical data, automatically assign ids to each of the unqiue values and replace the column values with the ids. A special id is used for missing values. Read more about the transformer [here](http://docs.fast.ai/tabular.transform.html#class-fillmissing).
We can now setup a [Tabular Dataset](http://docs.fast.ai/tabular.data.html#TabularDataset) for training. In addition to applying the transformations above the `TabularDataset` will also *normalize* the continuous variables for us.
```
tabular_data = tabular_data_from_df('output', train_df, valid_df, dep_var, tfms=tfms, cat_names=categorical_names)
```
### Model
We are now ready to create a [TabularModel](http://docs.fast.ai/tabular.models.html#class-tabularmodel). TabularModels are relatively simple models that have Embedding layers for each of the categorical variables which are concatenated with the continuous layers before being passed to a fully connected layers. The model also includes Dropout and Batchnorm layers where expected (see below for detailed depiction of the layers).
In our case we use 2 fully connected layers of dimension 100. Since we are doing regression, we then have an output layer of a single dimension (which will automatically have a Linear activation) and we use a mean squared error loss function. We also specify embedding layer sizes for the two categorical layers.
```
learn = get_tabular_learner(tabular_data,
layers=[100,50,1],
emb_szs={'education': 6,
'sex': 5,
'marital_status': 8})
learn.loss_fn = F.mse_loss
```
The code below outputs a detailed description of the model:
```
learn.model
```
Finally, before starting training, we need to choose an appropriate learning rate. FastAI provides an extremely neat trick for finding an appropriate learning rate. See [here](http://docs.fast.ai/callbacks.lr_finder.html) and [here](https://arxiv.org/abs/1506.01186) for details.
Applying the method is very straightfoward:
```
learn.lr_find()
```
After running `lr_find()` we can plot the losses against the learning rates that were tried:
```
learn.recorder.plot()
```
We then choose a value that is smaller (FastAI recommends by an order of magnitude) than the point at which the loss explodes. In our case there is not increase in loss even for large learning rates, so we could try even bigger start and stop values for lr find.. For now, we will use `1e-1`.
```
lr = 1e-1
```
We can now train our model. FastAI supports training with [cyclical learning rates](http://docs.fast.ai/callbacks.one_cycle.html#OneCycleScheduler) (the only deep learning libary to do so out of the box), which is used when we call the `fit_one_cycle` method. Here we perform a single epoch of training:
```
learn.fit_one_cycle(1, lr)
```
Great! Now we have a trained model, although it definitely seems to be overfitting on our data. The `recorder` records a lot of training data for us. For example, we can easily plot our loss per iterations:
```
learn.recorder.plot_losses()
```
Our learning rate helped a lot, as we can see, the network converged very early in the training.
We can now continue training with smaller learning rates if we wanted to and attempt to find a better minima.
### Random Forest Baseline
As shown [here](https://gist.github.com/dienhoa/a5adf923bd8b24b3d0eadcd61aec8c2e), we can re-use the FastAI datasets (and pre-processing) to build training data for other algorithms. For example, here we build a training and validation set for a random forest regressor to use a baseline for our DNN.
```
def get_arrays(ds):
X_cats,X_conts = ds.cats.numpy(), ds.conts.numpy()
y = ds.y.numpy()
return np.concatenate((X_cats, X_conts), axis=1), y
X_train, y_train = get_arrays(tabular_data.train_ds)
X_val, y_val = get_arrays(tabular_data.valid_ds)
from sklearn.ensemble import RandomForestRegressor
rf = RandomForestRegressor()
rf.fit(X_train,y_train)
rf.score(X_val, y_val)
from sklearn.metrics import mean_squared_error
mean_squared_error(y_val, rf.predict(X_val))
```
Seems like we have some work to do to optimize our network. Likely we can simply tune our learning rate decay to use a much smaller learning rate near the end of the epoch.
## References
Please have a look at the excellent FastAI docs for a detailed overview of the API: http://docs.fast.ai.
The FastAI forums are also worth mentioning, it's one of the most active and inclusive Deep Learning forums out there: http://forums.fast.ai
| github_jupyter |
# Why should I write code when I can write code that writes code?
> The temptation to employ code-generating techniques in Python is strong. Much of what is called "metaprogramming" in Python refers to the various techniques through which we can write higher-order code: code that generates the code that solves our problem. This talk discusses various common approaches for code-generation, their applicability to solving real problems, and the reality of using these techniques in your work.
```
# Consider a simple function
def f(x, y):
return x + y
print(f'{f(10, 20) = }')
```
What happens is that python takes the source code and converts that into byte code and actually executes the byte code.
```
from dis import dis
dis(f)
```
WHat happpens beyond is that python takes this source code on dis transforms it into an AST. The ast looks something like this.
```
code = '''
def f(x, y):
return x + y
'''
from ast import parse, dump
parse(code)
code = '''
def f(x, y):
return x + y
'''
from pprint import pprint
pprint(dump(parse(code)))
```
Once it creates the AST, it goes and tries to generate bytecode from that AST (like generating a symbol table, constant folding, optimization) but at the end you have a code object that looks like this.
```
code = '''
def f(x, y):
return x + y
'''
ast = parse(code)
bytecode = compile(ast, '', mode='exec')
bytecode
# bytecode is exactly some bytes which tell python interpreter what to do
bytecode.co_code
list(bytecode.co_code)
```
Bytecode is singe byte numeric values representing what is the operation that the python interpreter should do and the best way to look at that is to see what each number means.
```
from dis import opname
pprint([opname[x] for x in bytecode.co_code])
```
Now if we want to go deeper than that, we can think of how does it work in python. So the most naive understanding is there's a mechanism in python that looks at the bytecodes and runs them one by one. This is exactly what is done in CPython. In CPython there is a Pyeval method which does infinite looping to go through this bytecode. So if I write a function that calls another function than we can see multiple loops of Pyeval.
What happens when we import a module? Here is a very siple example of what happens when you import a module for the first time in python. Python will see if it has imported the module before, if I have then return the value from sys.modules, if it is the first time then it looks for a .py file, opens it, gets the bytecode, compiles it and returns a namespace for it.
```
from ast import parse
from sys import modules
from pathlib import Path
def import_(mod):
if mod not in modules:
file = Path(mod).with_suffix('.py')
with open(file) as f:
source = f.read()
ast = parse(source)
code = compile(ast, mod, mode='exec')
ns = {}
exec(code, ns)
modules[mod] = ns
return modules[mod]
f = import_('testmod')['f']
print(f'{f(10, 20) = }')
```
If we look at the building of a class.
```
class T:
def f(self):
return f'T.f({self!r})'
T().f()
```
Now to see what happens in the background is something as follows (Note the below code is not what really happens).
```
body = '''
def f(self):
return f'T.f({self!r})'
'''
def build_class(name, body):
ns = {} # prepare
exec(body, ns)
t = type(name, (), ns) # __init__, __new__
return t
T = build_class('T', body)
T().f()
```
What actually happens is a lot uglier. What happens is that you take the body of the function, you put it into another function, you execute that function to build the class body at runtime, you do that within a namespace.
Say you have two functions f and g.
```
def f():
if account in active_account and user in authorized_users:
do_work()
def g():
if account in active_account and user in authorized_users:
do_other_work()
```
Say the underlying codebase for valid users change, so you have to update both the above functions, but that is difficult. So we may write an abstraction as a function.
```
def f():
check_authorized()
do_work()
def g():
check_authorized()
do_work()
```
We can even use decorators to help with the problem.
```
@check_authorized
def f():
do_work()
@check_authorized
def g():
do_other_work()
```
So we are dealing with udpate anamalies here. We want our code to remain updated in different parts of our project without having to do much work. Auto code generation is one way to deal with this problem.
Often people talk about code generaton as a metaprogramming approach. Most approaches of metaprogramming in python generally fall into four categories:
* use some build-in functionality
* hook into some built-in functionality
* construct something dynamically
* (at various layers)
```
def f(x, y):
return x + y
def g(x, y):
return x ** y
```
A very simple functional programming approach to combine the above functions is to use a operation method.
```
from operator import add, pow
def func(x, y, op):
return op(x, y)
```
Now if you want a function f and g which performs that functionality you can create a function inside a function.
```
def create_func(op):
def func(x, y):
return op(x, y)
return func
f = create_func(add)
g = create_func(pow)
```
If you dig deeper we can see the way we generate these functions in decorators are very closely tied to the way we create an instance of a class.
| github_jupyter |
# Usage
We'll start with a probabilistic regression example on the Boston housing dataset:
```
import sys
sys.path.append('/Users/c242587/Desktop/projects/git/ngboost')
from ngboost import NGBRegressor
from sklearn.datasets import load_boston
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
X, Y = load_boston(True)
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=0.2)
ngb = NGBRegressor().fit(X_train, Y_train)
Y_preds = ngb.predict(X_test)
Y_dists = ngb.pred_dist(X_test)
# test Mean Squared Error
test_MSE = mean_squared_error(Y_preds, Y_test)
print('Test MSE', test_MSE)
# test Negative Log Likelihood
test_NLL = -Y_dists.logpdf(Y_test).mean()
print('Test NLL', test_NLL)
```
Getting the estimated distributional parameters at a set of points is easy. This returns the predicted mean and standard deviation of the first five observations in the test set:
```
Y_dists[0:5].params
```
## Distributions
NGBoost can be used with a variety of distributions, broken down into those for regression (support on an infinite set) and those for classification (support on a finite set).
### Regression Distributions
| Distribution | Parameters | Implemented Scores | Reference |
| --- | --- | --- | --- |
| `Normal` | `loc`, `scale` | `LogScore`, `CRPScore` | [`scipy.stats` normal](https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.norm.html) |
| `LogNormal` | `s`, `scale` | `LogScore`, `CRPScore` | [`scipy.stats` lognormal](https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.lognorm.html) |
| `Exponential` | `scale` | `LogScore`, `CRPScore` | [`scipy.stats` exponential](https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.expon.html) |
Regression distributions can be used through the `NGBRegressor()` constructor by passing the appropriate class as the `Dist` argument. `Normal` is the default.
```
from ngboost.distns import Exponential, Normal
X, Y = load_boston(True)
X_reg_train, X_reg_test, Y_reg_train, Y_reg_test = train_test_split(X, Y, test_size=0.2)
ngb_norm = NGBRegressor(Dist=Normal, verbose=False).fit(X_reg_train, Y_reg_train)
ngb_exp = NGBRegressor(Dist=Exponential, verbose=False).fit(X_reg_train, Y_reg_train)
```
There are two prediction methods for `NGBRegressor` objects: `predict()`, which returns point predictions as one would expect from a standard regressor, and `pred_dist()`, which returns a distribution object representing the conditional distribution of $Y|X=x_i$ at the points $x_i$ in the test set.
```
ngb_norm.predict(X_reg_test)[0:5]
ngb_exp.predict(X_reg_test)[0:5]
ngb_exp.pred_dist(X_reg_test)[0:5].params
```
#### Survival Regression
NGBoost supports analyses of right-censored data. Any distribution that can be used for regression in NGBoost can also be used for survival analysis in theory, but this requires the implementation of the right-censored version of the appropriate score. At the moment, `LogNormal` and `Exponential` have these scores implemented. To do survival analysis, use `NGBSurvival` and pass both the time-to-event (or censoring) and event indicator vectors to `fit()`:
```
import numpy as np
from ngboost import NGBSurvival
from ngboost.distns import LogNormal
X, Y = load_boston(True)
X_surv_train, X_surv_test, Y_surv_train, Y_surv_test = train_test_split(X, Y, test_size=0.2)
# introduce administrative censoring to simulate survival data
T_surv_train = np.minimum(Y_train, 30) # time of an event or censoring
E_surv_train = Y_train > 30 # 1 if T[i] is the time of an event, 0 if it's a time of censoring
ngb = NGBSurvival(Dist=LogNormal).fit(X_surv_train, T_surv_train, E_surv_train)
```
The scores currently implemented assume that the censoring is independent of survival, conditional on the observed predictors.
### Classification Distributions
| Distribution | Parameters | Implemented Scores | Reference |
| --- | --- | --- | --- |
| `k_categorical(K)` | `p0`, `p1`... `p{K-1}` | `LogScore` | [Categorical distribution on Wikipedia](https://en.wikipedia.org/wiki/Categorical_distribution) |
| `Bernoulli` | `p` | `LogScore` | [Bernoulli distribution on Wikipedia](https://en.wikipedia.org/wiki/Bernoulli_distribution) |
Classification distributions can be used through the `NGBClassifier()` constructor by passing the appropriate class as the `Dist` argument. `Bernoulli` is the default and is equivalent to `k_categorical(2)`.
```
from ngboost import NGBClassifier
from ngboost.distns import k_categorical, Bernoulli
from sklearn.datasets import load_breast_cancer
X, y = load_breast_cancer(True)
y[0:15] = 2 # artificially make this a 3-class problem instead of a 2-class problem
X_cls_train, X_cls_test, Y_cls_train, Y_cls_test = train_test_split(X, y, test_size=0.2)
ngb_cat = NGBClassifier(Dist=k_categorical(3), verbose=False) # tell ngboost that there are 3 possible outcomes
_ = ngb_cat.fit(X_cls_train, Y_cls_train) # Y should have only 3 values: {0,1,2}
```
When using NGBoost for classification, the outcome vector `Y` must consist only of integers from 0 to K-1, where K is the total number of classes. This is consistent with the classification standards in sklearn.
`NGBClassifier` objects have three prediction methods: `predict()` returns the most likely class, `predict_proba()` returns the class probabilities, and `pred_dist()` returns the distribution object.
```
ngb_cat.predict(X_cls_test)[0:5]
ngb_cat.predict_proba(X_cls_test)[0:5]
ngb_cat.pred_dist(X_cls_test)[0:5].params
```
## Scores
NGBoost supports the log score (`LogScore`, also known as negative log-likelihood) and CRPS (`CRPScore`), although each score may not be implemented for each distribution. The score is specified by the `Score` argument in the constructor.
```
from ngboost.scores import LogScore, CRPScore
NGBRegressor(Dist=Exponential, Score=CRPScore, verbose=False).fit(X_reg_train, Y_reg_train)
NGBClassifier(Dist=k_categorical(3), Score=LogScore, verbose=False).fit(X_cls_train, Y_cls_train)
```
## Base Learners
NGBoost can be used with any sklearn regressor as the base learner, specified with the `Base` argument. The default is a depth-3 regression tree.
```
from sklearn.tree import DecisionTreeRegressor
learner = DecisionTreeRegressor(criterion='friedman_mse', max_depth=5)
NGBSurvival(Dist=Exponential, Score=CRPScore, Base=learner, verbose=False).fit(X_surv_train, T_surv_train, E_surv_train)
```
## Other Arguments
The learning rate, number of estimators, minibatch fraction, and column subsampling are also easily adjusted:
```
ngb = NGBRegressor(n_estimators=100, learning_rate=0.01,
minibatch_frac=0.5, col_sample=0.5)
ngb.fit(X_reg_train, Y_reg_train)
```
Sample weights (for training) are set using the `sample_weight` argument to `fit`.
```
ngb = NGBRegressor(n_estimators=100, learning_rate=0.01,
minibatch_frac=0.5, col_sample=0.5)
weights = np.random.random(Y_reg_train.shape)
ngb.fit(X_reg_train, Y_reg_train, sample_weight=weights)
```
| github_jupyter |
<img src="https://deep-hybrid-datacloud.eu/wp-content/uploads/sites/2/2018/01/logo.png" width="100">
<h1><center>DEEP-HybridDataCloud tutorial: Integrating a simple Keras application</center></h1>
This tutorial will guide you through the integration of a simple Deep Learning model (CIFAR10 image classification using Keras) with the API architecture. For more details you can check the [DEEP documentation repository](http://docs.deep-hybrid-datacloud.eu "DEEP docs")
If you want to have the general view of the model you can take a look at this [Model Notebook](https://github.com/laramaktub/IBERGRID-tutorial/blob/master/CIFAR10.ipynb "Notebook"). We will go in more detail through the tutorial.
For running this tutorial we are going to be using a [Docker container](https://www.docker.com). A Docker container image is a lightweight, standalone, executable package of software that includes everything needed to run an application: code, runtime, system tools, system libraries and settings. This means that the user doesn't have to worry about all these details and can focus only on the use/development of the application.
Take into account that you should have a git account for doing this tutorial. Make sure you have properly defined your **git identification variables:**
```console
$ git config --global user.email "you@example.com"
$ git config --global user.name "Your_Name"
```
First of all, we need to **install cookiecutter** which creates projects from project templates:
```console
$ pip install cookiecutter
```
Now that we have cookiecutter installed, we can generate the template for our project:
```console
$ cookiecutter https://github.com/indigo-dc/cookiecutter-data-science --checkout tutorial
```
The template for the project will be downloaded and you will be asked several questions.
```
git_base_url [https://github.com/deephdc]: https://github.com/your_github_user
project_name [project_name]: cifar10
repo_name [cifar10]: cifar10
author_name [Your name (or your organization/company/team)]: your_name
author_email [Your email]: your_mail
description [A short description of the project.]: CIFAR10 classification app
app_version [Application version (expects X.Y.Z (Major.Minor.Patch))]: 1.0.0
Select open_source_license:
1 - MIT
2 - BSD-3-Clause
3 - No license file
Choose from 1, 2, 3 (1, 2, 3) [1]: 1
Select python_interpreter:
1 - python3
2 - python
Choose from 1, 2 (1, 2) [1]: 1
dockerhub_user [User account at hub.docker.com, e.g. 'deephdc' in https://hub.docker.com/u/deephdc]: your_docker_user
docker_baseimage [Base Docker image for Dockerfile, e.g. tensorflow/tensorflow]: tensorflow/tensorflow
baseimage_cpu_tag [CPU tag for the Base Docker image, e.g. 1.12.0-py3. Has to match python version!]: 1.10.0-py3
baseimage_gpu_tag [GPU tag for the Base Docker image, e.g. 1.12.0-gpu-py3. Has to match python version!]: 1.10.0-gpu-py3
```
This will create two project directories:
```
~/cifar10
~/DEEP-OC-cifar10
```
Go to ``github.com/your_account`` and create the corresponding repositories: ``DEEP-OC-cifar10`` and ``cifar10``.
Now enter in the directories you just created and do ``git push origin master`` in both created directories. This puts your initial code to github
You are now ready to run the Docker container. For the moment it will just contain an empty model.
Go to the ``~/DEEP-OC-cifar10`` directory and build the docker image:
```
sudo docker build -t cifar10docker .
```
This will generate a docker image that you can check by doing:
```console
$ sudo docker images
```
Now let's run the docker container by typing:
```console
$ sudo docker run -ti -p 5000:5000 -p 8888:8888 -p 6006:6006 cifar10docker /bin/bash
```
This will deploy the container. We can now type on the command line the following command to run the jupyter lab:
```console
$ /srv/.jupyter/run_jupyter.sh --allow-root
```
## SECOND PART OF TUTORIAL ##
This will generate an URL with a token. We can enter it in the web browser and we will now be running jupyter lab!
Now, in jupyter Lab, you can go to Terminal. You will be seeing the directory structure in your docker. You can launch the DEEPaaS API by typing:
```console
$ deepaas-run --listen-ip 0.0.0.0
```
The DEEPaaS API will be now running and accesible at ``your_VM_IP:5000``
You can easily check that right now there is no model loaded yet. For this, check the current metadata of the model and check the training part. You can see that there are no input arguments defined.
Let's now include our **cifar10** model!
```
requirements.txt
data/
models/
cifar10/dataset/make_dataset.py
cifar10/features/build_features.py
cifar10/models/deepaas_api.py
cifar10/config.py
```
The file ``cifar10/config.py`` contains the input parameters that we want to give as input to our application via the web API. Open it and check that right now there is no parameter defined.
For this tutorial we want to have as parameters:
* Learning rate
* Number of epochs
* Output directory
You can add this parameters by defining your train_args and your pred_args in the config.py file as:
```
train_args = { 'epochs': {'default': 1,
'help': 'Number of epochs',
'required': True,
'type': int
},
'lrate': {'default':0.001,
'help': 'Initial learning rate value',
'required': True,
'type': float
},
'outputpath': {'default': "/tmp",
'help': 'Path for saving the model',
'required': True
},
}
predict_args = { 'outputpath': {'default': "/tmp",
'help': 'Path for loading the model',
'required': True
},
}
```
Launch again the DEEPaaS API. You will now see these input parameters there!
In the ``requirements.txt`` file, one can include any additional package that may be needed to run the model. In our case, we only need to include one more line to this file for installing keras version 2.0.2.
```
keras==2.0.2
```
Now let's download our model. For doing so, you can go to the ``cifar10/models/`` folder and type:
```console
$ wget https://raw.githubusercontent.com/deephdc/DEEP-tutorial-cifar10/master/cifar10/models/cifar10_model.py
```
Open the ``cifar10_model.py`` file and take a look. You will see two functions called ``train_nn`` and ``predict_nn``. They contain our model both for training and for prediction respectively.
The ``deepaas_api.py`` is the script containing the functions that are going to be called from the DEEPaaS API. We will be editing only the functions:
* get_metadata(): retrieves the model metadata
* train(): launch the model training
* predict_data(): predict on some image loaded by the user
Now you can download a filled version of the ``deepaas_api()`` file, go to the models folder and type:
```console
$ wget -O deepaas_api.py https://raw.githubusercontent.com/deephdc/DEEP-tutorial-cifar10/master/cifar10/models/deepaas_api.py
```
Open the file and check how the functions from our model are being imported:
```console
$ from cifar10.models.cifar10_model import train_nn, predict_nn
```
Start by filling the metadata in the get_metadata() function. An example:
meta = {
'Name': "cifar10",
'Version': "1.0.0",
'Summary': "This is a simple implementation of cifar10 in keras",
'Home-page': None,
'Author': "Lara Lloret Iglesias",
'Author-email': "lloret@ifca.unican.es",
'License': "MIT",
}
Now take a look at the ``predict_data()`` and ``train()`` functions.
The ``predict_data()`` has been modified in order to retrieve the loaded images and pass it to the
```
def predict_data(*args):
"""
Function to make prediction on an uploaded file
"""
outputpath=args[0]["outputpath"]
thefile= args[0]['files'][0]
thename= thefile.filename
thepath= outputpath + "/" +thename
thefile.save(thepath)
img = image.load_img(thepath, target_size=(32,32))
x= image.img_to_array(img)
message=predict_nn(x,outputpath)
return message
```
The ``train()`` function looks like this:
```
def train(train_args):
# Train network
# train_args : dict
# Json dict with the user's configuration parameters.
# Can be loaded with json.loads() or with yaml.safe_load()
train_nn(train_args['epochs'], train_args['lrate'],train_args['outputpath'])
run_results = { "status": "SUCCESS",
"train_args": [],
"training": [],
}
run_results["train_args"].append(train_args)
return run_results
```
This retrieves the input arguments and passing them to the train_nn function from our model.
We are now ready to go!
Run the following command from the same directory where your requirements.txt is to install the cifar10 package with the new dependencies:
```console
$ python -m pip install -e .
```
Go to the terminal and launch the application again!
```console
$ deepaas-run --listen-ip 0.0.0.0
```
You can now play with the model directly from the DEEPaaS API.
* Go to your browser and type your_VM_IP:5000
* Run the get_metadata()
* Save some truck picture from google and try to do some prediction (this should not work yet)
* Launch the training: try with 1 or 2 epochs, learning rate of 0.001 and the default output path ("/tmp")
* Try prediction again with the truck picture
* Feel free to train again with more samples, during more epochs, etc...
That was it! You have just integrated your first model in the DEEPaaS API!
| github_jupyter |
# SignalDB
The SignalDB database contains all of the meta-data for the raw SETI data availaible in the SETI@IBMCloud project.
You can use this data to visualize the locations of stars the SETI Institute has observed, select and sort through to find data that interests you, and ultimately use these meta-data to download the appropriate raw data file for analysis, feature extraction and machine learning.
This database is available from the `setigopublic` server and can be downloaded with `curl`, unzipped and read into a Spark Dataframe in the usual way.
```
!curl https://setigopublic.mybluemix.net/v1/aca/meta/all > signaldb.csv.gz
!gunzip signaldb.csv.gz
!ls -alh signaldb.csv
#Spark 2.0 and higher
from pyspark.sql import SparkSession
spark = SparkSession.builder.getOrCreate()
df = spark.read\
.format('org.apache.spark.sql.execution.datasources.csv.CSVFileFormat')\
.option('header', 'true')\
.option('inferSchema','true')\
.load('signaldb.csv')
df.take(5)
df.cache()
```
# Visualization with Pixiedust
[Pixiedust is a tool built for Spark](https://github.com/ibm-cds-labs/pixiedust) that assists developers in visualization,
loading and saving data, and package management that allows one to install and use Scala/Java Spark packages from within the Pyspark environment.
```
#!pip install --user --upgrade pixiedust
import pixiedust
df.createOrReplaceTempView("signaldb")
```
We'll use pixiedust to display the RA/DEC location of the data. First, we need to remove the 'spacecraft' data
```
df = spark.sql("select * from signaldb where catalog!='spacecraft'")
display(df)
```
## Explore with Pixiedust
There are many things to explore with the SignalDB data. You don't need to access the raw data (in the next step) in order to start visualizing and analyzing SETI data.
It would be interesting to see a density plot of Candidate signals and "candreason"/"sigreason" versus position in the sky, the distribution of carrier wave signal frequencies (freqmhz), drift rates, and signal types for each RA/DEC position, etc. There are lots of plots to be generated from these data alone.
# SparkSQL to select a subset of SignalDB
The following SQL selects data for a particular star. This statement returns the raw data Object Store container and objectname for all data for that star in the data set.
## Interesting Target
Kepler 1229b, found here http://phl.upr.edu/projects/habitable-exoplanets-catalog, is a very intersting target. This candidate has an Earth similarity index (ESI) of 0.73 (Earth = 1.0, Mars = 0.64, Jupiter = 0.12) and is 770 light-years away. It is the 5th highest-ranked planet by ESI. According to the catalog, it's RA/DEC coordinates are '19h49m56.81s' and '+46d59m48.2s', respectively.
### Coordinates
Often, you'll find coordinates for the right ascension (RA) in units of hours and declination (DEC) in units of "day minute seconds". One can use the [`astropy` python package](http://www.astropy.org/) to [convert between coordinate systems](http://docs.astropy.org/en/stable/coordinates/index.html), amongst other functionalities provided by the package. The coordinates in the SignalDB typically only reach three decimal places and are in units of `decimal hours` and `degrees`.
```
#!pip install --user --upgrade astropy
from astropy.coordinates import SkyCoord
kepler1229b_ra = '19h49m56.81s'
kepler1229b_dec = '+46d59m48.2s'
c = SkyCoord(kepler1229b_ra, kepler1229b_dec)
df2 = spark.sql("select * from signaldb where RA2000hr=={0:.3f} and dec2000deg=={1:.3f}".format(c.ra.hour, c.dec.deg))
display(df2)
```
# Read the Raw Data
The raw data are located on IBM Object storage in containers. The `container` and `objectname` values tell you specifically where to look. See the [subsequent notebooks for tutorials](https://github.com/ibm-cds-labs/seti_at_ibm#introduction-notebooks) on reading the raw data files, creating spectrograms and extracting features.
```
df2_loc = spark.sql("select container, objectname from signaldb where RA2000hr=={0:.3f} and dec2000deg=={1:.3f}".format(c.ra.hour, c.dec.deg))
raw_loc = df2_loc.collect()
raw_loc[:5]
```
# Tabby's Star?
```
tabby_ra = '20h06m15.457s'
tabby_dec = '+44d27m24.61s'
c = SkyCoord(tabby_ra, tabby_dec)
dftabby = spark.sql("select * from signaldb where RA2000hr=={0:.3f} and dec2000deg=={1:.3f}".format(c.ra.hour, c.dec.deg))
dftabby.cache()
dftabby.count()
```
No data yet in public data set. :(
| github_jupyter |
```
# Copyright 2019 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Idiomatic Programmer Code Labs
## Code Labs #2 - Get Familiar with Training
## Prerequistes:
1. Familiar with Python
2. Completed Handbook 3/Part 11: Training & Deployment
## Objectives:
1. Pretraining for Weight Initialization
2. TODO: Early Stopping
3. TODO: Model Saving and Restoring
## Pretraining
We are going to do some pre-training runs to find a good initial weight initialization. Each time the weights are initialized, they are randomly choosen from the selected distribution (i.e., kernel_initializer).
We will do the following:
1. Make three instances of the same model, each with their own weight initialization.
2. Take a subset of the training data (20%)
3. Train each model instance for a few epochs.
4. Pick the instance with the highest valuation accuracy.
5. Use this instance to train the model with the entire training data.
```
from keras import Sequential, optimizers
from keras.layers import Conv2D, MaxPooling2D, Flatten, Dense, Dropout
from keras.utils import to_categorical
from keras.datasets import cifar10
import numpy as np
# Let's use the CIFAR-10 dataset
(x_train, y_train), (x_test, y_test) = cifar10.load_data()
# Normalize the pixel data
x_train = (x_train / 255.0).astype(np.float32)
x_test = (x_test / 255.0).astype(np.float32)
# One-hot encode the labels
y_train = to_categorical(y_train)
y_test = to_categorical(y_test)
# Let's take a fraction of the training data to test the weight initialization (20%)
# Generally, we like to use all the training data for this purpose, but for brevity we will use 20%
x_tmp = x_train[0:10000]
y_tmp = y_train[0:10000]
# We will use this function to build a simple CNN, using He-Normal initialization for the weights.
def convNet(input_shape, nclasses):
model = Sequential()
model.add(Conv2D(32, kernel_size=(3, 3), activation='relu', kernel_initializer='he_normal',
input_shape=input_shape))
model.add(Conv2D(64, (3, 3), activation='relu', kernel_initializer='he_normal'))
model.add(Dropout(0.25))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Flatten())
model.add(Dense(128, activation='relu', kernel_initializer='he_normal'))
model.add(Dense(nclasses, activation='softmax'))
return model
# Let's make 3 versions of the model, each with their own weight initialization.
models = []
for _ in range(3):
model = convNet((32, 32, 3), 10)
# We will use (assume best) learning rate of 0.001
model.compile(loss='categorical_crossentropy', optimizer=optimizers.Adam(lr=0.001), metrics=['accuracy'])
# Let's do the short training of 20% of training data for 5 epochs.
model.fit(x_tmp, y_tmp, epochs=5, batch_size=32, validation_split=0.1, verbose=1)
# Save a copy of the model
# HINT: We are saving the in-memory partially trained model
models.append(??)
# Now let's pick the model instance with the highest val_acc and train it with the full training data
# HINT: Index will be 0, 1 or 2
models[1].fit(x_train, y_train, epochs=10, batch_size=32, validation_split=0.1, verbose=1)
score = model.evaluate(x_test, y_test)
print(score)
```
## EarlyStopping
Note that the training accuracy in the above example keeps going up, but at some point the validation loss swings back up and validation goes down. That means you are overfitting -- even with the dropout.
Let's now look on how to decide how many epochs we should run. We can use early stopping technique. In this case, we set the number of epochs larger than we anticipate, and then set an objective to reach. When the objective is reached, we stop training.
```
from keras.callbacks import EarlyStopping
# Let's try this with a fresh model, and not care about the weight initialization this time.
model = convNet((32, 32, 3), 10)
model.compile(loss='categorical_crossentropy', optimizer=optimizers.Adam(lr=0.001), metrics=['accuracy'])
# Set an early stop (termination of training) when the valuation loss has stopped
# reducing (default setting).
earlystop = EarlyStopping(monitor='val_loss')
# Train the model and use early stop to stop training early if the valuation loss
# stops decreasing.
# HINT: what goes in the callbacks list is the instance (variable) of the EarlyStopping object
model.fit(x_train, y_train, epochs=10, batch_size=32, validation_split=0.1, verbose=1, callbacks=[??])
```
## Model Saving and Restoring
Let's do a basic store of the model and weights to disk, and then mimic restoring the model from disk to in memory.
```
from keras.models import load_model
# Save the model and trained weights and biases.
model.save('mymodel.h5')
# load a pre-trained model as a different model instance (mymodel instead of model)
mymodel = load_model('mymodel.h5')
# Let's verify we really do that.
score = mymodel.evaluate(x_test, y_test)
print(score)
```
## End of Code Lab
| github_jupyter |
```
%autosave 180
import convex as cx
import requests
import time
import spacy
nlp = spacy.load("en_core_web_lg")
def get_nlp(sentence):
return nlp(sentence)
from hdt import HDTDocument
hdt_wd = HDTDocument("data/kb/wikidata2018_09_11.hdt")
questions = [
"Which actor voiced the Unicorn in The Last Unicorn?",
"And Alan Arkin was behind...?",
"And Alan Arkin be behind...? Why How when which was happy make fun",
"Who is the composer of the soundtrack?",
"So who performed the songs?",
"Genre of this band's music?",
"By the way, who was the director?"
]
q_test = str("Which actor voiced the Unicorn in The Last Unicorn? "+
"And Alan Arkin was behind...? "+
"And Alan Arkin be behind...? Why How when which was happy make fun. "+
"Who is the composer of the soundtrack? "+
"So who performed songs? "+
"Genre of this band's music? "+
"By the way, who was the director? ")
q_test_2 = "Who is the wife of Barack Obama?"
q0_nlp = get_nlp(questions[0])
q0_nlp_test = get_nlp(q_test)
q0_nlp_test_2 = get_nlp(q_test_2)
q0_nlp
import re
def is_wd_entity(to_check):
pattern = re.compile('^Q[0-9]*$')
if pattern.match(to_check.strip()): return True
else: return False
def is_wd_predicate(to_check):
pattern = re.compile('^P[0-9]*$')
if pattern.match(to_check.strip()): return True
else: return False
def is_valide_wd_id(to_check):
if is_wd_entity(to_check) or is_wd_predicate(to_check): return True
else: return False
print(is_valide_wd_id("P8765"))
def get_wd_ids_online(name, is_predicate=False, top_k=3):
name = name.split('(')[0]
request_successfull = False
entity_ids = ""
while not request_successfull:
try:
if is_predicate:
entity_ids = requests.get('https://www.wikidata.org/w/api.php?action=wbsearchentities&format=json&language=en&type=property&limit=' + str(top_k) + '&search='+name).json()
else:
entity_ids = requests.get('https://www.wikidata.org/w/api.php?action=wbsearchentities&format=json&language=en&limit=' + str(top_k) + '&search='+name).json()
request_successfull = True
except:
time.sleep(5)
results = entity_ids.get("search")
if not results:
return ""
if not len(results):
return ""
res = []
for result in results:
res.append(result['id'])
return res
get_wd_ids_online("voiced", is_predicate=False, top_k=1)
import warnings
warnings.filterwarnings('ignore')
# very computational
def get_most_similar(word, topn=5):
word = nlp.vocab[str(word)]
queries = [w for w in word.vocab if w.is_lower == word.is_lower and w.prob >= -15]
by_similarity = sorted(queries, key=lambda w: word.similarity(w), reverse=True)
return [(w.lower_,w.similarity(word)) for w in by_similarity[:topn+1] if w.lower_ != word.lower_]
get_most_similar("voiced", topn=3)
def get_wd_ids(word, top_k=3, limit=10):
language = "en"
word_formated = str("\""+word+"\""+"@"+language)
to_remove = len("http://www.wikidata.org/entity/")
t_name, card_name = hdt_wd.search_triples("", "http://schema.org/name", word_formated, limit=top_k)
#print("names cardinality of \"" + word+"\": %i" % card_name)
t_alt, card_alt = hdt_wd.search_triples("", 'http://www.w3.org/2004/02/skos/core#altLabel', word_formated, limit=top_k)
#print("alternative names cardinality of \"" + word+"\": %i" % card_alt)
return list(set(
[t[0][to_remove:] for t in t_name if is_valide_wd_id(t[0][to_remove:])] +
[t[0][to_remove:] for t in t_alt if is_valide_wd_id(t[0][to_remove:])]
))[:limit]
get_wd_ids("The Last Unicorn", top_k=3, limit=10)
def get_wd_label(from_id):
#print("from_id",from_id)
if is_valide_wd_id(from_id):
language = "en"
id_url = "http://www.wikidata.org/entity/"+from_id
t_name, card_name = hdt_wd.search_triples(id_url, "http://schema.org/name", "")
name = [t[2].split('\"@en')[0].replace("\"", "") for t in t_name if "@"+language in t[2]]
return name[0] if name else ''
else:
return ''
get_wd_label("P725")
#get_wd_label("Q20789322")
import matplotlib.pyplot as plt
%matplotlib inline
# Building colors from graph
def get_color(node_type):
if node_type == "entity": return "violet"#"cornflowerblue"
elif node_type == "predicate": return "yellow"
else: return "red"
# Building labels for graph
def get_elements_from_graph(graph):
node_names = nx.get_node_attributes(graph,"name")
node_types = nx.get_node_attributes(graph,"type")
colors = [get_color(node_types[n]) for n in node_names]
return node_names, colors
# Plotting the graph
def plot_graph(graph, name, title="Graph"):
fig = plt.figure(figsize=(14,14))
ax = plt.subplot(111)
ax.set_title(str("answer: "+title), fontsize=10)
pos = nx.spring_layout(graph)
labels, colors = get_elements_from_graph(graph)
nx.draw(graph, pos, node_size=30, node_color=colors, font_size=10, font_weight='bold', with_labels=True, labels=labels)
plt.tight_layout()
plt.savefig(str(name)+".png", format="PNG", dpi = 300)
plt.show()
#plot_graph(graph, "file_name_graph", "Graph_title")
import networkx as nx
def make_statements_graph(statements, indexing_predicates=True):
graph = nx.Graph()
turn=0
predicate_nodes = {}
for statement in statements:
#print(statement)
if not statement['entity']['id'] in graph:
graph.add_node(statement['entity']['id'], name=get_wd_label(statement['entity']['id']), type='entity', turn=turn)
if not statement['object']['id'] in graph:
graph.add_node(statement['object']['id'], name=get_wd_label(statement['object']['id']), type='entity', turn=turn)
# increment index of predicate or set it at 0
if not statement['predicate']['id'] in predicate_nodes or not indexing_predicates:
predicate_nodes_index = 1
predicate_nodes[statement['predicate']['id']] = 1
else:
predicate_nodes[statement['predicate']['id']] += 1
predicate_nodes_index = predicate_nodes[statement['predicate']['id']]
# add the predicate node
predicate_node_id = (statement['predicate']['id'])
if indexing_predicates: predicate_node_id += "-" + str(predicate_nodes_index)
graph.add_node(predicate_node_id, name=get_wd_label(statement['predicate']['id']), type='predicate', turn=turn)
# add the two edges (entity->predicate->object)
graph.add_edge(statement['entity']['id'], predicate_node_id)
graph.add_edge(predicate_node_id, statement['object']['id'])
return graph, predicate_nodes
#test_graph = make_statements_graph(test_unduplicate_statements, indexing_predicates=False)
#print(test_graph[1])
#plot_graph(test_graph[0],"test")
def merge_lists(list_1, list_2):
if len(list_1) == len(list_2):
return [(list_1[i], list_2[i]) for i in range(0, len(list_1))]
else:
return "Error: lists are not the same lenght"
merge_lists([1,2,3],[4,5,6])
def get_themes(nlp_question, top_k=3):
themes = []
theme_complements = []
noun_chunks = [chunk for chunk in nlp_question.noun_chunks]
theme_ids = [get_wd_ids(chunk.text, top_k=top_k) for chunk in noun_chunks]
for i, chunk in enumerate(theme_ids):
if chunk: themes.append((noun_chunks[i], chunk))
else: theme_complements.append(noun_chunks[i])
return themes, theme_complements
q0_themes = get_themes(q0_nlp, top_k=3)
q0_themes_test = get_themes(q0_nlp_test)
q0_themes_test_2 = get_themes(q0_nlp_test_2)
q0_themes
def get_predicates_online(nlp_sentence, top_k=3):
predicates = [p for p in nlp_sentence if p.pos_ == "VERB" or p.pos_ == "AUX"]
predicates_ids = [get_wd_ids_online(p.text, is_predicate=True, top_k=top_k) for p in predicates]
return merge_lists(predicates, predicates_ids)
q0_predicates = get_predicates_online(q0_nlp, top_k=3)
q0_predicates_test_2 = get_predicates_online(q0_nlp_test_2, top_k=3)
q0_predicates
def get_focused_parts(nlp_sentence, top_k=3):
focused_parts = [t.head for t in nlp_sentence if t.tag_ == "WDT" or t.tag_ == "WP" or t.tag_ == "WP$" or t.tag_ == "WRB"]
focused_parts_ids = [get_wd_ids(p.text, top_k=top_k) for p in focused_parts]
return merge_lists(focused_parts, focused_parts_ids)
q0_focused_parts = get_focused_parts(q0_nlp)
q0_focused_parts_test_2 = get_focused_parts(q0_nlp_test_2)
q0_focused_parts
from itertools import chain
def extract_ids(to_extract):
return [i for i in chain.from_iterable([id[1] for id in to_extract])]
#extract_ids([('name', ['id'])]) #q0_themes[0] #q0_focused_parts #q0_predicates
#extract_ids([("The Last Unicorn", ['Q16614390']),("Second Theme", ['Q12345'])])
extract_ids(q0_focused_parts)
def get_similarity_by_words(nlp_word_from, nlp_word_to):
if not nlp_word_from or not nlp_word_to:
return 0
elif not nlp_word_from.vector_norm or not nlp_word_to.vector_norm:
return 0
else:
return nlp_word_from.similarity(nlp_word_to)
get_similarity_by_words(get_nlp("character role"), get_nlp("voice actor"))
def get_similarity_by_ids(word_id_from, word_id_to):
nlp_word_from = get_nlp(get_wd_label(word_id_from))
nlp_word_to = get_nlp(get_wd_label(word_id_to))
return get_similarity_by_words(nlp_word_from, nlp_word_to)
get_similarity_by_ids("P453", "P725")
def get_top_similar_statements(statements, from_token_id, similar_to_name, top_k=3, qualifier=False, statement_type="object"):
highest_matching_similarity = -1
top_statements = []
nlp_name = get_nlp(similar_to_name)
if get_wd_label(from_token_id):
for statement in statements:
if qualifier:
if statement.get('qualifiers'):
for qualifier in statement['qualifiers']:
nlp_word_to = get_nlp(get_wd_label(qualifier[statement_type]['id']))
matching_similarity = get_similarity_by_words(nlp_name, nlp_word_to)
if highest_matching_similarity == -1 or matching_similarity > highest_matching_similarity:
highest_matching_similarity = matching_similarity
best_statement = statement
top_statements.append((highest_matching_similarity, best_statement))
else:
nlp_word_to = get_nlp(get_wd_label(statement[statement_type]['id']))
matching_similarity = get_similarity_by_words(nlp_name, nlp_word_to)
if highest_matching_similarity == -1 or matching_similarity > highest_matching_similarity:
highest_matching_similarity = matching_similarity
best_statement = statement
top_statements.append((highest_matching_similarity, best_statement))
return sorted(top_statements, key=lambda x: x[0], reverse=True)[:top_k]
statements = cx.wd.get_all_statements_of_entity('Q176198')
top_similar_statements = get_top_similar_statements(statements, 'Q176198', 'voiced')
top_similar_statements[0]
def get_best_similar_statements_by_word(from_token_ids, similar_to_name, top_k=3, qualifier=False, statement_type="object"):
best_statements = []
for token in from_token_ids:
statements = cx.wd.get_all_statements_of_entity(token)
if statements: best_statements += get_top_similar_statements(statements, token, similar_to_name, top_k=top_k, qualifier=qualifier, statement_type=statement_type)
return sorted(best_statements, key=lambda x: x[0], reverse=True)[:top_k]
best_similar_statements = get_best_similar_statements_by_word(extract_ids(q0_themes[0]), 'voiced', top_k=3, qualifier=True, statement_type="qualifier_object")
best_similar_statements[0]
def get_statements_subjects_labels(statements):
return [get_wd_label(t[1]['entity']['id']) for t in statements]
get_statements_subjects_labels(best_similar_statements)
def get_statements_predicates_labels(statements):
return [get_wd_label(t[1]['predicate']['id']) for t in statements]
get_statements_predicates_labels(best_similar_statements)
def get_statements_objects_labels(statements):
return [get_wd_label(t[1]['object']['id']) for t in statements]
get_statements_objects_labels(best_similar_statements)
def get_statements_qualifier_predicates_labels(statements):
return [get_wd_label(t[1]['qualifiers'][0]['qualifier_predicate']['id']) for t in statements]
get_statements_qualifier_predicates_labels(best_similar_statements)
def get_statements_qualifier_objects_labels(statements):
return [get_wd_label(t[1]['qualifiers'][0]['qualifier_object']['id']) for t in statements]
get_statements_qualifier_objects_labels(best_similar_statements)
def cluster_extend_by_words(cluster_root_ids, extending_words, top_k=3):
cluster = []
for name in extending_words:
cluster += get_best_similar_statements_by_word(cluster_root_ids, name, top_k=top_k, qualifier=True, statement_type="qualifier_predicate")
cluster += get_best_similar_statements_by_word(cluster_root_ids, name, top_k=top_k, qualifier=True, statement_type="qualifier_object")
cluster += get_best_similar_statements_by_word(cluster_root_ids, name, top_k=top_k, qualifier=False, statement_type="predicate")
cluster += get_best_similar_statements_by_word(cluster_root_ids, name, top_k=top_k, qualifier=False, statement_type="object")
return cluster
test_cluster = cluster_extend_by_words(extract_ids(q0_themes[0]), ['voiced'], top_k=2)
test_cluster_test_2 = cluster_extend_by_words(extract_ids(q0_themes_test_2[0]), ['birth'], top_k=2)
test_cluster[0]
# sorts by the similarity value of statements[0]
def sort_statements_by_similarity(statements):
return [s for s in sorted(statements, key=lambda x: x[0], reverse=True)]
test_sorted_statements = sort_statements_by_similarity(test_cluster)
test_sorted_statements_test_2 = sort_statements_by_similarity(test_cluster_test_2)
test_sorted_statements[0]
from copy import copy
# appends spo from qualifiers, removes qualifier tags, and removes similarity scores
def statements_flatter(statements):
best_statements_to_graph = []
for statement in statements:
tmp_statement = copy(statement)
if tmp_statement.get('qualifiers'):
#print("statement", statement)
for q in tmp_statement['qualifiers']:
qualifier_statement = {'entity': {'id': tmp_statement['entity']['id']}}
qualifier_statement['predicate'] = {'id': q['qualifier_predicate']['id']}
qualifier_statement['object'] = {'id': q['qualifier_object']['id']}
best_statements_to_graph.append(qualifier_statement)
del(tmp_statement['qualifiers'])
else:
#print("tmp_statement", tmp_statement)
if ('qualifiers' in tmp_statement): del(tmp_statement['qualifiers'])
if tmp_statement not in best_statements_to_graph:
#print("best_statements_to_graph", tmp_statement)
best_statements_to_graph.append(tmp_statement)
return best_statements_to_graph
test_flatten_statements = statements_flatter([s[1] for s in test_sorted_statements])
#test_flatten_statements_test_2 = statements_flatter([s[1] for s in test_sorted_statements_test_2])
test_flatten_statements[0]
#test_flatten_statements_test_2
# remove duplicates from statements
def unduplicate_statements(statements):
filtered_statements = []
[filtered_statements.append(s) for s in statements if s not in [e for e in filtered_statements]]
return filtered_statements
test_unduplicate_statements = unduplicate_statements(test_flatten_statements)
print(len(test_flatten_statements))
print(len(test_unduplicate_statements))
test_unduplicate_statements[0]
test_graph = make_statements_graph(test_unduplicate_statements)
print(test_graph[1])
plot_graph(test_graph[0], "file_name_graph", "Graph_title")
def get_statements_by_id(statements, from_token_id, to_id, qualifier=False, statement_type="predicate"):
id_statements = []
if not statements:
return id_statements
if cx.wd.wikidata_id_to_label(from_token_id):
for statement in statements:
if qualifier:
if statement.get('qualifiers'):
for s in statement['qualifiers']:
if to_id == s[statement_type]['id']:
id_statements.append(statement)
else:
if to_id == statement[statement_type]['id']:
id_statements.append(statement)
return id_statements
#statements_test = cx.wd.get_all_statements_of_entity('Q176198')
#id_statements_test = get_statements_by_id(statements_test, 'Q176198', 'P725')
#id_statements_test[0]
#get_statements_by_id(root_statements, cluster_root_id, predicate_id, qualifier=False, statement_type="predicate")
#statements_test = cx.wd.get_all_statements_of_entity('Q176198')
#id_statements_test = get_statements_by_id(statements_test, 'Q176198', 'P725')
#id_statements_test[0]
# parameters
# cluster_root_ids: ['Qcode']
# predicates_ids: ['Pcode']
def cluster_extend_by_predicates_ids(cluster_root_ids, predicates_ids):
cluster = []
for cluster_root_id in cluster_root_ids:
root_statements = cx.wd.get_all_statements_of_entity(cluster_root_id)
#print("root_statements", root_statements)
for predicate_id in predicates_ids:
cluster += get_statements_by_id(root_statements, cluster_root_id, predicate_id, qualifier=True, statement_type="qualifier_predicate")
cluster += get_statements_by_id(root_statements, cluster_root_id, predicate_id, qualifier=False, statement_type="predicate")
return cluster
#test_predicate_clusters = cluster_extend_by_predicates_ids(extract_ids(q0_themes[0]), extract_ids(q0_predicates))
#print(len(test_predicate_clusters))
#test_predicate_clusters[0]
test_predicate_clusters_test_2 = cluster_extend_by_predicates_ids(extract_ids(q0_themes_test_2[0]), extract_ids(q0_predicates_test_2))
print(len(test_predicate_clusters_test_2))
test_predicate_clusters_test_2[-1]
# parameter
# question: nlp_string
def build_graph(nlp, themes, predicates, deep_k=50):
init_clusters = cluster_extend_by_words(extract_ids(themes[0]), [p[0].text for p in predicates], top_k=deep_k)
init_sorted_statements = sort_statements_by_similarity(init_clusters)
init_flatten_statements = statements_flatter([s[1] for s in init_sorted_statements])
predicate_ids_clusters = cluster_extend_by_predicates_ids(extract_ids(themes[0]), extract_ids(predicates))
predicate_ids_flatten_statements = statements_flatter(predicate_ids_clusters)
clusters = init_flatten_statements+predicate_ids_flatten_statements
filtered_statements = unduplicate_statements(clusters)
graph = make_statements_graph(filtered_statements)
#print("clusters:", len(clusters))
#print("filtered_statements:", len(filtered_statements))
return graph
#q0_test = questions[0]
q0_test = "Which actor voiced the Unicorn in The Last Unicorn?"
q0_nlp_test = get_nlp(q0_test)
q0_themes_test = get_themes(q0_nlp_test, top_k=3)
q0_predicates_test = get_predicates_online(q0_nlp_test, top_k=3)
q0_focused_parts_test = get_focused_parts(q0_nlp_test)
graph, predicates_dict = build_graph(q0_nlp_test, q0_themes_test, q0_predicates_test)
print(predicates_dict)
plot_graph(graph, "file_name_graph", "Graph_title")
# check the graph for complements
# parameters
# name: string
def find_name_in_graph(graph, name):
return [x for x,y in graph.nodes(data=True) if y['name'].lower() == name.lower()]
#[find_name_in_graph(c.text) for c in q0_themes[1]]
find_name_in_graph(graph, "the unicorn")
# TODO: clean the complements by removing stopwords etc.
def find_theme_complement(graph, themes):
return [i for i in chain.from_iterable(
[id for id in [c for c in [find_name_in_graph(graph, t.text) for t in themes[1]] if c]])]
find_theme_complement(graph, q0_themes_test)
#[i for i in chain.from_iterable([id for id in check_theme_complement(graph, q0_themes)])]
def find_paths_in_graph(graph, node_start, node_end):
return [p for p in nx.all_simple_paths(graph, source=node_start, target=node_end)]
test_paths = find_paths_in_graph(graph, "Q176198", "Q202725")
test_paths
def is_id_in_graph(graph, node_id):
return graph.has_node(node_id)
is_id_in_graph(graph, "Q176198")
def is_name_in_graph(graph, node_name):
return find_name_in_graph(graph, node_name) != []
is_name_in_graph(graph, "the Unicorn")
def find_paths_for_themes(graph, themes):
themes_ids = [t for t in extract_ids(themes[0])]
complements_ids = find_theme_complement(graph, themes)
paths = []
for t_id in themes_ids:
if is_id_in_graph(graph, t_id):
for c_id in complements_ids:
if is_id_in_graph(graph, c_id):
path = find_paths_in_graph(graph, t_id, c_id)
if path:
paths.append(path)
paths = [i for i in chain.from_iterable(
[id for id in paths])]
return paths
find_paths_for_themes(graph, q0_themes_test)
find_paths_for_themes(graph, q0_themes)
def get_node_predicates_from_path(paths):
predicates = []
for p in paths:
[predicates.append(i[:i.find("-")]) for i in p if is_wd_predicate(i[:i.find("-")]) and i[:i.find("-")] not in predicates]
return predicates
test_node_predicates = get_node_predicates_from_path(test_paths)
test_node_predicates
def get_node_predicate_similarity_from_path(paths, predicates):
path_predicates = get_node_predicates_from_path(paths)
return sorted([(pp, get_similarity_by_ids(p2, pp)) for p in predicates for p2 in p[1] for pp in path_predicates], key=lambda x: x[-1], reverse=True)
test_node_pedicate_similarities = get_node_predicate_similarity_from_path(test_paths, q0_predicates)
test_node_pedicate_similarities
# TODO: make the predicate search go further in the path list for the !i%2
def find_anwser_from_graph(graph, q_nlp, themes, predicates):
initial_paths = find_paths_for_themes(graph, themes)
predicate_id_similarities = get_node_predicate_similarity_from_path(initial_paths, predicates)
best_path = [p for p in initial_paths if predicate_id_similarities[0][0] == p[1][:p[1].find("-")]]
path_answer = get_wd_label(best_path[0][2]) if best_path else []
return path_answer, best_path[0][2] if path_answer else (False, False)
find_anwser_from_graph(graph, q0_nlp, q0_themes, q0_predicates)
def answer_initial_question(question):
q_nlp = get_nlp(question)
q_themes = get_themes(q_nlp, top_k=3)
q_predicates = get_predicates_online(q_nlp, top_k=3)
q_focused_parts = get_focused_parts(q_nlp)
#print(q_nlp, q_themes, q_predicates, q_focused_parts)
graph, predicates_dict = build_graph(q_nlp, q_themes, q_predicates)
#print(predicates_dict)
#plot_graph(graph, "main_graph", "Main_graph_title")
return find_anwser_from_graph(graph, q0_nlp, q0_themes, q0_predicates)
#answer_initial_question("Which actor voiced the Unicorn in The Last Unicorn?")
#answer_initial_question("Who is the author of Le Petit Prince?")
#answer_initial_question("Who made the soundtrack of the The Last Unicorn movie?")
#answer_initial_question("When was produced the first Matrix movie?")
#answer_initial_question("Who is the president of the United States?")
#answer_initial_question("Who is the wife of Barack Obama?")
answer_initial_question("what was the cause of death of yves klein")
subgraphs = [graph.subgraph(c) for c in nx.connected_components(graph)]
len(subgraphs[0].nodes)
def search_focused_part(graph):
return 0
search_focused_part(graph)
for path in nx.all_simple_paths(graph, source="Q176198", target="Q202725"):
print(path)
nx.shortest_path(graph, source="Q176198", target="Q202725")
[y['name'] for x,y in graph.nodes(data=True) if y['name'].lower() == "The Unicorn".lower()]
for complement in q0_themes[1]:
print(complement)
for e in complement:
print(e, e.pos_, e.tag_, e.dep_, e.head)
print("\n")
for e in q0_nlp:
print(e, e.pos_, e.tag_, e.dep_, e.head)
list(nx.dfs_labeled_edges(graph, source=get_themes(question_0_nlp, top_k=3)[0][0][1][0], depth_limit=4))[0]
get_themes(question_0_nlp, top_k=3)[0][0][1][0]
```
| github_jupyter |
# Fisheries competition
In this notebook we're going to investigate a range of different techniques for the [Kaggle fisheries competition](https://www.kaggle.com/c/the-nature-conservancy-fisheries-monitoring). In this competition, The Nature Conservancy asks you to help them detect which species of fish appears on a fishing boat, based on images captured from boat cameras of various angles. Your goal is to predict the likelihood of fish species in each picture. Eight target categories are available in this dataset: Albacore tuna, Bigeye tuna, Yellowfin tuna, Mahi Mahi, Opah, Sharks, Other
You can use [this](https://github.com/floydwch/kaggle-cli) api to download the data from Kaggle.
```
# Put these at the top of every notebook, to get automatic reloading and inline plotting
%reload_ext autoreload
%autoreload 2
%matplotlib inline
# This file contains all the main external libs we'll use
from fastai.imports import *
from fastai.plots import *
from fastai.io import get_data
PATH = "data/fish/"
```
## First look at fish pictures
```
!ls {PATH}
!ls {PATH}train
files = !ls {PATH}train/ALB | head
files
img = plt.imread(f'{PATH}train/ALB/{files[0]}')
plt.imshow(img);
```
## Data pre-processing
Here we are changing the structure of the training data to make it more convinient. We will have all images in a common directory `images` and will have a file `train.csv` with all labels.
```
from os import listdir
from os.path import join
train_path = f'{PATH}/train'
dirs = [d for d in listdir(train_path) if os.path.isdir(join(train_path,d))]
print(dirs)
train_dict = {d: listdir(join(train_path, d)) for d in dirs}
train_dict["LAG"][:10]
sum(len(v) for v in train_dict.values())
with open(f"{PATH}train.csv", "w") as csv:
csv.write("img,label\n")
for d in dirs:
for f in train_dict[d]: csv.write(f'{f},{d}\n')
img_path = f'{PATH}images'
os.makedirs(img_path, exist_ok=True)
!cp {PATH}train/*/*.jpg {PATH}images/
```
## Our first model with Center Cropping
Here we import the libraries we need. We'll learn about what each does during the course.
```
from fastai.transforms import *
from fastai.conv_learner import *
from fastai.model import *
from fastai.dataset import *
from fastai.sgdr import *
sz=350
bs=64
csv_fname = os.path.join(PATH, "train.csv")
train_labels = list(open(csv_fname))
n = len(list(open(csv_fname)))-1
val_idxs = get_cv_idxs(n)
tfms = tfms_from_model(resnet34, sz)
data = ImageClassifierData.from_csv(PATH, "images", csv_fname, bs, tfms, val_idxs)
learn = ConvLearner.pretrained(resnet34, data, precompute=True, opt_fn=optim.Adam, ps=0.5)
lrf=learn.lr_find()
learn.sched.plot()
learn.fit(0.01, 4, cycle_len=1, cycle_mult=2)
lrs=np.array([1e-4,1e-3,1e-2])
learn.precompute=False
learn.freeze_to(6)
lrf=learn.lr_find(lrs/1e3)
learn.sched.plot()
```
## Same model with No cropping
NOTE: Before running this remove the temp file under data/fish.
```
sz = 350
tfms = tfms_from_model(resnet34, sz, crop_type=CropType.NO)
data = ImageClassifierData.from_csv(PATH, "images", csv_fname, bs, tfms, val_idxs)
learn = ConvLearner.pretrained(resnet34, data, precompute=True, opt_fn=optim.Adam, ps=0.5)
lrf=learn.lr_find()
learn.sched.plot()
learn.fit(0.01, 4, cycle_len=1, cycle_mult=2)
lrs=np.array([1e-4,1e-3,1e-2])
learn.precompute=False
learn.unfreeze()
lrf=learn.lr_find(lrs/1e3)
learn.sched.plot()
lrs=np.array([1e-5,1e-4,1e-3])
learn.fit(lrs, 5, cycle_len=1, cycle_mult=2)
```
## Predicting bounding boxes
### Getting bounding boxes data
This part needs to run just the first time to get the file `trn_bb_labels`
```
import json
anno_classes = ['alb', 'bet', 'dol', 'lag', 'other', 'shark', 'yft']
def get_annotations():
annot_urls = [
'5458/bet_labels.json', '5459/shark_labels.json', '5460/dol_labels.json',
'5461/yft_labels.json', '5462/alb_labels.json', '5463/lag_labels.json'
]
cache_subdir = os.path.abspath(os.path.join(PATH, 'annos'))
url_prefix = 'https://kaggle2.blob.core.windows.net/forum-message-attachments/147157/'
os.makedirs(cache_subdir, exist_ok=True)
for url_suffix in annot_urls:
fname = url_suffix.rsplit('/', 1)[-1]
get_data(url_prefix + url_suffix, f'{cache_subdir}/{fname}')
# run this code to get annotation files
get_annotations()
# creates a dictionary of all annotations per file
bb_json = {}
for c in anno_classes:
if c == 'other': continue # no annotation file for "other" class
j = json.load(open(f'{PATH}annos/{c}_labels.json', 'r'))
for l in j:
if 'annotations' in l.keys() and len(l['annotations'])>0:
bb_json[l['filename'].split('/')[-1]] = sorted(
l['annotations'], key=lambda x: x['height']*x['width'])[-1]
bb_json['img_04908.jpg']
raw_filenames = pd.read_csv(csv_fname)["img"].values
file2idx = {o:i for i,o in enumerate(raw_filenames)}
empty_bbox = {'height': 0., 'width': 0., 'x': 0., 'y': 0.}
for f in raw_filenames:
if not f in bb_json.keys(): bb_json[f] = empty_bbox
bb_params = ['height', 'width', 'x', 'y']
def convert_bb(bb):
bb = [bb[p] for p in bb_params]
bb[2] = max(bb[2], 0)
bb[3] = max(bb[3], 0)
return bb
trn_bbox = np.stack([convert_bb(bb_json[f]) for f in raw_filenames]).astype(np.float32)
trn_bb_labels = [f + ',' + ' '.join(map(str,o))+'\n' for f,o in zip(raw_filenames,trn_bbox)]
open(f'{PATH}trn_bb_labels', 'w').writelines(trn_bb_labels)
fnames,csv_labels,_,_ = parse_csv_labels(f'{PATH}trn_bb_labels', skip_header=False)
def bb_corners(bb):
bb = np.array(bb, dtype=np.float32)
row1 = bb[3]
col1 = bb[2]
row2 = row1 + bb[0]
col2 = col1 + bb[1]
return [row1, col1, row2, col2]
f = 'img_02642.jpg'
bb = csv_labels[f]
print(bb)
bb_corners(bb)
new_labels = [f + "," + " ".join(map(str, bb_corners(csv_labels[f]))) + "\n" for f in raw_filenames]
open(f'{PATH}trn_bb_corners_labels', 'w').writelines(new_labels)
```
### Looking at bounding boxes
```
# reading bb file
bbox = {}
bb_data = pd.read_csv(f'{PATH}trn_bb_labels', header=None)
fnames,csv_labels,_,_ = parse_csv_labels(f'{PATH}trn_bb_labels', skip_header=False)
fnames,corner_labels,_,_ = parse_csv_labels(f'{PATH}trn_bb_corners_labels', skip_header=False)
corner_labels["img_06297.jpg"]
csv_labels["img_06297.jpg"]
def create_rect(bb, color='red'):
return plt.Rectangle((bb[2], bb[3]), bb[1], bb[0], color=color, fill=False, lw=3)
def show_bb(path, f='img_04908.jpg'):
file_path = f'{path}images/{f}'
bb = csv_labels[f]
plots_from_files([file_path])
plt.gca().add_patch(create_rect(bb))
def create_corner_rect(bb, color='red'):
bb = np.array(bb, dtype=np.float32)
return plt.Rectangle((bb[1], bb[0]), bb[3]-bb[1], bb[2]-bb[0], color=color, fill=False, lw=3)
def show_corner_bb(path, f='img_04908.jpg'):
file_path = f'{path}images/{f}'
bb = corner_labels[f]
plots_from_files([file_path])
plt.gca().add_patch(create_corner_rect(bb))
show_corner_bb(PATH, f = 'img_02642.jpg')
```
### Model predicting bounding boxes
```
sz=299
bs=64
label_csv=f'{PATH}trn_bb_corners_labels'
n = len(list(open(label_csv)))-1
val_idxs = get_cv_idxs(n)
tfms = tfms_from_model(resnet34, sz, crop_type=CropType.NO, tfm_y=TfmType.COORD)
data = ImageClassifierData.from_csv(PATH, 'images', label_csv, tfms=tfms, val_idxs=val_idxs,
continuous=True, skip_header=False)
trn_ds = data.trn_dl.dataset
x, y = trn_ds[0]
print(x.shape, y)
learn = ConvLearner.pretrained(resnet34, data, precompute=True, opt_fn=optim.Adam, ps=0.5)
lrf=learn.lr_find()
learn.sched.plot()
learn.fit(0.01, 5, cycle_len=1, cycle_mult=2)
lrs=np.array([1e-4,1e-3,1e-2])
learn.precompute=False
learn.unfreeze()
lrf=learn.lr_find(lrs/1e3)
learn.sched.plot()
lrs=np.array([1e-5,1e-4,1e-3])
learn.fit(lrs, 5, cycle_len=1, cycle_mult=2)
```
## Looking into size of images
```
f="img_06297.jpg"
PIL.Image.open(PATH+"images/" + f).size
sizes = [PIL.Image.open(PATH+f).size for f in data.trn_ds.fnames]
raw_val_sizes = [PIL.Image.open(PATH+f).size for f in data.val_ds.fnames]
```
| github_jupyter |
# US - Baby Names
### Introduction:
We are going to use a subset of [US Baby Names](https://www.kaggle.com/kaggle/us-baby-names) from Kaggle.
In the file it will be names from 2004 until 2014
### Step 1. Import the necessary libraries
```
import pyspark
from pyspark.sql import SparkSession
spark = SparkSession.builder.appName("baby").getOrCreate()
spark
```
### Step 2. Import the dataset from this [address](https://raw.githubusercontent.com/guipsamora/pandas_exercises/master/06_Stats/US_Baby_Names/US_Baby_Names_right.csv).
### Step 3. Assign it to a variable called baby_names.
```
from pyspark import SparkFiles
url = "https://raw.githubusercontent.com/guipsamora/pandas_exercises/master/06_Stats/US_Baby_Names/US_Baby_Names_right.csv"
spark.sparkContext.addFile(url)
baby_names = spark.read.csv(SparkFiles.get("US_Baby_Names_right.csv"), header=True, inferSchema=True, sep=',')
baby_names.show(5)
```
### Step 4. See the first 10 entries
```
baby_names.show(10)
print(baby_names.count())
```
### Step 5. Delete the column 'Unnamed: 0' and 'Id'
```
cols_to_drop = ["_c0","Id"]
baby_names= baby_names.drop(*cols_to_drop)
baby_names.show(2)
```
### Step 6. Is there more male or female names in the dataset?
```
females = baby_names.filter(baby_names.Gender.startswith("F")).count()
males = baby_names.filter(baby_names.Gender.startswith("M")).count()
print("females:",females, "males:",males)
if females > males:
print("More Females",females)
else:
print("More Males",males)
```
### Step 7. Group the dataset by name and assign to names
```
names = baby_names.groupBy("Name").count()
names = names.withColumnRenamed("count","name_count")
names.show(5)
```
### Step 8. How many different names exist in the dataset?
```
names.count()
```
### Step 9. What is the name with most occurrences?
```
names.orderBy("name_count",ascending=0).show(5)
names.orderBy("name_count",ascending=0).head(1)[0][0]
names.filter(names.Name.startswith("Jacob")).show()
```
### Step 10. How many different names have the least occurrences?
```
from pyspark.sql.functions import *
min_count = names.agg({"name_count":"min"}).head(1)[0][0]
min_count
rare_names = names.filter(names.name_count.isin(min_count)).count()
rare_names
```
### Step 11. What is the median name occurrence?
```
# df.approxQuantile("x", [0.5], 0.25)
median_count = names.approxQuantile("name_count",[0.5],0.0001)
median_count
```
### Step 12. What is the standard deviation of names?
```
names.describe(['name_count']).show()
```
### Step 13. Get a summary with the mean, min, max, std and quartiles.
```
names.approxQuantile("name_count",[0.25,0.5,0.75],0.1)
```
| github_jupyter |
# Efficient Estimation using 1 simulation
1. This notebook shows how to **estimate** a simple model using Simulated Minimum Distance (SMD)
2. It illustrates how an **efficient** estimator can be constructed using only 1 simulatoin, following the idea proposed by [Kirill Evdokimov](https://www.mit.edu/~kevdokim/ESMSM_sep16.pdf "Efficient Estimation with a Finite Number of Simulation Draws per Observation")
## Recap: Simulated Minimum Distance
**Data:** We assume that we have data available for $N$ households over $T$ periods, collected in $\{w_i\}_i^N$.
**Goal:** We wish to estimate the true, unknown, parameter vector $\theta_0$. We assume our model is correctly specified in the sense that the observed data stems from the model.
The **Simulated Minimum Distance (SMD)** estimator is
$$
\hat{\theta} = \arg\min_{\theta} g(\theta)'Wg(\theta)
$$
where $W$ is a $J\times J$ positive semidefinite **weighting matrix** and
$$
g(\theta)=\Lambda_{data}-\Lambda_{sim}(\theta)
$$
is the distance between $J\times1$ vectors of moments calculated in the data and the simulated data, respectively. Concretely,
$$
\Lambda_{data} = \frac{1}{N}\sum_{i=1}^N m(\theta_0|w_i) \\
\Lambda_{sim}(\theta) = \frac{1}{N_{sim}}\sum_{s=1}^{N_{sim}} m(\theta|w_s)
$$
are $J\times1$ vectors of moments calculated in the data and the simulated data, respectively.
**Variance of the estimator:** Recall that the variance of the estimator was
$$
\begin{align}
\text{Var}(\hat{\theta})&=(1+S^{-1})\Gamma\Omega\Gamma'/N \\
\Gamma &= -(G'WG)^{-1}G'W \\
\Omega & = \text{Var}(m(\theta_0|w_i))
\end{align}
$$
where we implicitly used that $Var(m(\theta_0|w_i))=Var(m(\theta|w_s))$ and $Cov(m(\theta_0|w_i),m(\theta|w_s))=0$
**Efficient Estimator:** Using the "optimal" weighting matrix, $W=\Omega^{-1}$, gives the *lowest variance* for a given number of simulations, $S$, as
$$
\begin{align}
\text{Var}(\hat{\theta})&=(1+S^{-1})(G'\Omega^{-1}G)^{-1}/N
\end{align}
$$
> **Observation:** Only as $S\rightarrow\infty$ does the minimum variance of the SMD estimator approach the minimum variance of the GMM estimator.
> **Solution:** [Kirill Evdokimov](https://www.mit.edu/~kevdokim/ESMSM_sep16.pdf "Efficient Estimation with a Finite Number of Simulation Draws per Observation") shows how we can use an augmented set of moments related to the assumptions related to simulation to basically remove the factor $(1+S^{-1})$ on the asymptotic variance of the SMD estimator using only one(!) simulation, $S=1$!
# Model and Estimators
We use the same example as Kirill Evdokimov. Imagine the simple setup where we have the data-generating process (DGP):
$$
\begin{align}
Y_i &= \theta_0 + \varepsilon_i \\
\varepsilon_i &\sim N(0,1)
\end{align}
$$
**SMD:** We can use the moment function with only $S=1$ simulatin of $\varepsilon$ per individual
$$
g_i(\theta|w_i) = Y_i - \theta -\varepsilon_i
$$
to estimate $\theta$. We will call that $\hat{\theta}_{SMD}$. The moment vector would be
$$
g(\theta) =
\bigg( \begin{array}{c}
\overline{Y} - \theta -\overline{\varepsilon} \\
\end{array} \bigg)
$$
where $\overline{Y} = \frac{1}{N}\sum_{i=1}^{N} Y_i$ and $\overline{\varepsilon} = \frac{1}{N}\sum_{i=1}^{N} \varepsilon_i$.
**ES-SMD:** We can use the efficient SMD to augment the moment conditions with the fact that the simulated $\varepsilon$'s should have mean-zero and get the vector of moments in this augmented situation as
$$
g_{aug}(\theta) =
\bigg( \begin{array}{c}
\overline{Y} - \theta -\overline{\varepsilon} \\
0-\overline{\varepsilon} \\
\end{array} \bigg)
$$
where we use the optimal weighting matrix $W=\Omega^{-1}$ where
$$
\Omega = Var(g_{i,aug}(\theta|w_i)) =
\bigg( \begin{array}{cc}
2 & 1\\
1 & 1 \\
\end{array} \bigg)
$$
and
$$
\Omega^{-1} = \bigg( \begin{array}{cc}
1 & -1\\
-1 & 2 \\
\end{array} \bigg)
$$
We will call this estimator $\hat{\theta}_{ES-SMD}$.
**Asymptotic Variances:**
1. In the standard SMD estimator, the weighting matrix does not matter and we have
$$
\begin{align}
AVar(\hat{\theta}_{SMD}) &= Var(g_i(\theta|w_i)) \\
&= Var(Y_i - \theta -\varepsilon_i)\\
&= Var(Y_i) +Var(\varepsilon_i) \\
&= 2
\end{align}
$$
2. In the augmented ES-SMD estmator, we have
$$
\begin{align}
AVar(\hat{\theta}_{ES-SMD}) &= Var((G'WG)^{-1}G'Wg_{i,aug}(\theta|w_i)) \\
&= Var(-Y_i + \theta)\\
&= 1
\end{align}
$$
bacause
$$
(G'WG)^{-1}G'Wg_{i,aug}(\theta|w_i) = - (Y_i - \theta -\varepsilon) - \varepsilon.
$$
3. We thus have that the asymptotic variance of the ES-SMD estimator is lower that the SMD estimator!
We will now illustrate this result through a **Monte Carlo experiment** too!
# Setup
```
%load_ext autoreload
%autoreload 2
import numpy as np
from types import SimpleNamespace
import sys
sys.path.append('../')
from SimulatedMinimumDistance import SimulatedMinimumDistanceClass
```
# Model construction
```
class ModelClass():
def __init__(self,**kwargs):
self.par = SimpleNamespace()
self.sim = SimpleNamespace()
self.par.theta = 0.5
self.par.simN = 5000
for key,val in kwargs.items():
setattr(self.par,key,val)
def solve(self,do_print=False): pass
def simulate(self,seed=None,do_print=False):
if seed is not None:
np.random.seed(seed)
self.sim.e = np.random.normal(size=self.par.simN)
self.sim.Y = self.par.theta + self.sim.e
```
# Estimation choices
```
# a. model settings
N = 100_000
N_sim = N
par = {'theta':0.2,'simN':N_sim}
par_true = par.copy()
par_true['simN'] = N
# b. parameters to estimate
est_par = {
'theta': {'guess':0.5,'lower':0.0,'upper':1.0,},
}
# c. moment function used in estimation.
def mom_func(data,ids=None):
""" returns the average Y """
if ids is None:
mean_Y = np.mean(data.Y)
else:
mean_Y = np.mean(data.Y[ids])
return np.array([mean_Y]) # alwaus give a zero
# d. augmented moment function used in efficient estimation.
def mom_func_aug(data,ids=None):
""" returns the average Y and the average of the simulations"""
if ids is None:
mean_Y_e = np.mean([data.Y,data.e],axis=1)
else:
mean_Y_e = np.mean([data.Y[ids],data[ids].e],axis=1)
return mean_Y_e
```
# Monte Carlo Estimation results
```
num_boot = 1_000
theta_est = np.empty(num_boot)
theta_est_aug = theta_est.copy()
model = ModelClass(**par)
for b in range(num_boot):
# a. setup model to simulate data
true = ModelClass(**par_true)
true.simulate(seed=2050+b) # this seed is different from the default
# b. data moments
datamoms = mom_func(true.sim)
datamoms_aug = np.array([datamoms[0],0.0])
# c. setup estimators
smd = SimulatedMinimumDistanceClass(est_par,mom_func,datamoms=datamoms)
smd_aug = SimulatedMinimumDistanceClass(est_par,mom_func_aug,datamoms=datamoms_aug)
# d. weighting matrix
W = np.ones((datamoms.size,datamoms.size)) # does not matter here
Omega = np.array([[2.0,1.0],[1.0,1.0]]) # covariance matrix of augmentet moments.
W_aug = np.linalg.inv(Omega)
# e. estimate the model (can take several minutes)
est = smd.estimate(model,W,do_print_initial=False)
est_aug = smd_aug.estimate(model,W_aug,do_print_initial=False)
# f. store the estimates
theta_est[b] = est['theta']
theta_est_aug[b] = est_aug['theta']
print(f'Variance, SMD: {np.var(theta_est-par_true["theta"])*N:2.6f}')
print(f'Variance, ES-SMD: {np.var(theta_est_aug-par_true["theta"])*N:2.6f}')
```
| github_jupyter |
```
from pandas import read_csv
from pandas import datetime
from matplotlib import pyplot
import pandas as pd
def parser(x):
return datetime.strptime('200' + x, '%Y-%m')
series = read_csv('sales-of-shampoo-over-a-three-ye.csv', header=0, parse_dates=[0], index_col=0, squeeze=True,
date_parser=parser)
print(series.head())
# line plot
series.plot()
pyplot.show()
from pandas import DataFrame
from pandas import Series
from pandas import concat
from pandas import read_csv
from pandas import datetime
from sklearn.metrics import mean_squared_error
from sklearn.preprocessing import MinMaxScaler
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense
from tensorflow.keras.layers import LSTM
from tensorflow.keras.layers import SimpleRNN
from math import sqrt
import matplotlib
matplotlib.use('Agg')
from matplotlib import pyplot
import numpy
# frame a sequence as a supervised learning problem
def timeseries_to_supervised(data, lag=1):
df = DataFrame(data)
columns = [df.shift(i) for i in range(1, lag + 1)]
columns.append(df)
df = concat(columns, axis=1)
return df
# create a differenced series
def difference(dataset, interval=1):
diff = list()
for i in range(interval, len(dataset)):
value = dataset[i] - dataset[i - interval]
diff.append(value)
return Series(diff)
# invert differenced value
def inverse_difference(history, yhat, interval=1):
return yhat + history[-interval]
# scale train and test data to [-1, 1]
def scale(train, test):
# fit scaler
scaler = MinMaxScaler(feature_range=(-1, 1))
scaler = scaler.fit(train)
# transform train
train = train.reshape(train.shape[0], train.shape[1])
train_scaled = scaler.transform(train)
# transform test
test = test.reshape(test.shape[0], test.shape[1])
test_scaled = scaler.transform(test)
return scaler, train_scaled, test_scaled
# inverse scaling for a forecasted value
def invert_scale(scaler, X, yhat):
new_row = [x for x in X] + [yhat]
array = numpy.array(new_row)
array = array.reshape(1, len(array))
inverted = scaler.inverse_transform(array)
return inverted[0, -1]
# fit an LSTM network to training data
def fit_rnn(train, n_batch, nb_epoch, n_neurons):
X, y = train[:, 0:-1], train[:, -1]
X = X.reshape(X.shape[0], 1, X.shape[1])
model = Sequential()
model.add(SimpleRNN(n_neurons, batch_input_shape=(n_batch, X.shape[1], X.shape[2]), stateful=True))
model.add(Dense(1))
model.compile(loss='mean_squared_error', optimizer='adam')
for i in range(nb_epoch):
model.fit(X, y, epochs=1, batch_size=n_batch, verbose=0, shuffle=False)
model.reset_states()
return model
# run a repeated experiment
def run_rnn(series, n_lag, n_repeats, n_epochs, n_batch, n_neurons):
# transform data to be stationary
raw_values = series.values
diff_values = difference(raw_values, 1)
# transform data to be supervised learning
supervised = timeseries_to_supervised(diff_values, n_lag)
supervised_values = supervised.values[n_lag:, :]
# split data into train and test-sets
train, test = supervised_values[0:-12], supervised_values[-12:]
# transform the scale of the data
scaler, train_scaled, test_scaled = scale(train, test)
# run experiment
error_scores = list()
for r in range(n_repeats):
# fit the model
train_trimmed = train_scaled[2:, :]
rnn_model = fit_rnn(train_trimmed, n_batch, n_epochs, n_neurons)
# forecast test dataset
test_reshaped = test_scaled[:, 0:-1]
test_reshaped = test_reshaped.reshape(len(test_reshaped), 1, 1)
output = rnn_model.predict(test_reshaped, batch_size=n_batch)
predictions = list()
for i in range(len(output)):
yhat = output[i, 0]
X = test_scaled[i, 0:-1]
# invert scaling
yhat = invert_scale(scaler, X, yhat)
# invert differencing
yhat = inverse_difference(raw_values, yhat, len(test_scaled) + 1 - i)
# store forecast
predictions.append(yhat)
# report performance
rmse = sqrt(mean_squared_error(raw_values[-12:], predictions))
print('%d) Test RMSE: %.3f' % (r + 1, rmse))
error_scores.append(rmse)
return error_scores
# load dataset
series = read_csv('sales-of-shampoo-over-a-three-ye.csv', header=0, parse_dates=[0], index_col=0, squeeze=True,
date_parser=parser)
# configure the experiment
n_lag = 1
n_repeats = 30
n_epochs = 1000
n_batch = 4
n_neurons = 3
# run the experiment
results = DataFrame()
results['results'] = run_rnn(series, n_lag, n_repeats, n_epochs, n_batch, n_neurons)
results.plot(title="RNN RMSE Iteration")
pyplot.show()
# summarize results
print(results.describe())
# save boxplot
pyplot.savefig('plot_rnn_rmse.png')
```
| github_jupyter |
## RIHAD VARIAWA, Data Scientist - Who has fun LEARNING, EXPLORING & GROWING
<h1><center>Decision Trees</center></h1>
In this lab exercise, you will learn a popular machine learning algorithm, Decision Tree. You will use this classification algorithm to build a model from historical data of patients, and their response to different medications. Then you use the trained decision tree to predict the class of a unknown patient, or to find a proper drug for a new patient.
<h1>Table of contents</h1>
<div class="alert alert-block alert-info" style="margin-top: 20px">
<ol>
<li><a href="#about_dataset">About the dataset</a></li>
<li><a href="#downloading_data">Downloading the Data</a></li>
<li><a href="#pre-processing">Pre-processing</a></li>
<li><a href="#setting_up_tree">Setting up the Decision Tree</a></li>
<li><a href="#modeling">Modeling</a></li>
<li><a href="#prediction">Prediction</a></li>
<li><a href="#evaluation">Evaluation</a></li>
<li><a href="#visualization">Visualization</a></li>
</ol>
</div>
<br>
<hr>
Import the Following Libraries:
<ul>
<li> <b>numpy (as np)</b> </li>
<li> <b>pandas</b> </li>
<li> <b>DecisionTreeClassifier</b> from <b>sklearn.tree</b> </li>
</ul>
```
import numpy as np
import pandas as pd
from sklearn.tree import DecisionTreeClassifier
```
<div id="about_dataset">
<h2>About the dataset</h2>
Imagine that you are a medical researcher compiling data for a study. You have collected data about a set of patients, all of whom suffered from the same illness. During their course of treatment, each patient responded to one of 5 medications, Drug A, Drug B, Drug c, Drug x and y.
<br>
<br>
Part of your job is to build a model to find out which drug might be appropriate for a future patient with the same illness. The feature sets of this dataset are Age, Sex, Blood Pressure, and Cholesterol of patients, and the target is the drug that each patient responded to.
<br>
<br>
It is a sample of binary classifier, and you can use the training part of the dataset
to build a decision tree, and then use it to predict the class of a unknown patient, or to prescribe it to a new patient.
</div>
<div id="downloading_data">
<h2>Downloading the Data</h2>
To download the data, we will use !wget to download it from IBM Object Storage.
</div>
```
!wget -O _datasets/drug200.csv https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/ML0101ENv3/labs/drug200.csv
```
__Did you know?__ When it comes to Machine Learning, you will likely be working with large datasets. As a business, where can you host your data? IBM is offering a unique opportunity for businesses, with 10 Tb of IBM Cloud Object Storage: [Sign up now for free](http://cocl.us/ML0101EN-IBM-Offer-CC)
now, read data using pandas dataframe:
```
my_data = pd.read_csv("_datasets/drug200.csv", delimiter=",")
my_data[0:5]
```
<div id="practice">
<h3>Practice</h3>
What is the size of data?
</div>
```
# write your code here
```
<div href="pre-processing">
<h2>Pre-processing</h2>
</div>
Using <b>my_data</b> as the Drug.csv data read by pandas, declare the following variables: <br>
<ul>
<li> <b> X </b> as the <b> Feature Matrix </b> (data of my_data) </li>
<li> <b> y </b> as the <b> response vector (target) </b> </li>
</ul>
Remove the column containing the target name since it doesn't contain numeric values.
```
X = my_data[['Age', 'Sex', 'BP', 'Cholesterol', 'Na_to_K']].values
X[0:5]
```
As you may figure out, some features in this dataset are categorical such as __Sex__ or __BP__. Unfortunately, Sklearn Decision Trees do not handle categorical variables. But still we can convert these features to numerical values. __pandas.get_dummies()__
Convert categorical variable into dummy/indicator variables.
```
from sklearn import preprocessing
le_sex = preprocessing.LabelEncoder()
le_sex.fit(['F','M'])
X[:,1] = le_sex.transform(X[:,1])
le_BP = preprocessing.LabelEncoder()
le_BP.fit([ 'LOW', 'NORMAL', 'HIGH'])
X[:,2] = le_BP.transform(X[:,2])
le_Chol = preprocessing.LabelEncoder()
le_Chol.fit([ 'NORMAL', 'HIGH'])
X[:,3] = le_Chol.transform(X[:,3])
X[0:5]
```
Now we can fill the target variable.
```
y = my_data["Drug"]
y[0:5]
```
<hr>
<div id="setting_up_tree">
<h2>Setting up the Decision Tree</h2>
We will be using <b>train/test split</b> on our <b>decision tree</b>. Let's import <b>train_test_split</b> from <b>sklearn.cross_validation</b>.
</div>
```
from sklearn.model_selection import train_test_split
```
Now <b> train_test_split </b> will return 4 different parameters. We will name them:<br>
X_trainset, X_testset, y_trainset, y_testset <br> <br>
The <b> train_test_split </b> will need the parameters: <br>
X, y, test_size=0.3, and random_state=3. <br> <br>
The <b>X</b> and <b>y</b> are the arrays required before the split, the <b>test_size</b> represents the ratio of the testing dataset, and the <b>random_state</b> ensures that we obtain the same splits.
```
X_trainset, X_testset, y_trainset, y_testset = train_test_split(X, y, test_size=0.3, random_state=3)
```
<h3>Practice</h3>
Print the shape of X_trainset and y_trainset. Ensure that the dimensions match
```
# your code
```
Print the shape of X_testset and y_testset. Ensure that the dimensions match
```
# your code
```
<hr>
<div id="modeling">
<h2>Modeling</h2>
We will first create an instance of the <b>DecisionTreeClassifier</b> called <b>drugTree</b>.<br>
Inside of the classifier, specify <i> criterion="entropy" </i> so we can see the information gain of each node.
</div>
```
drugTree = DecisionTreeClassifier(criterion="entropy", max_depth = 4)
drugTree # it shows the default parameters
```
Next, we will fit the data with the training feature matrix <b> X_trainset </b> and training response vector <b> y_trainset </b>
```
drugTree.fit(X_trainset,y_trainset)
```
<hr>
<div id="prediction">
<h2>Prediction</h2>
Let's make some <b>predictions</b> on the testing dataset and store it into a variable called <b>predTree</b>.
</div>
```
predTree = drugTree.predict(X_testset)
```
You can print out <b>predTree</b> and <b>y_testset</b> if you want to visually compare the prediction to the actual values.
```
print (predTree [0:5])
print (y_testset [0:5])
```
<hr>
<div id="evaluation">
<h2>Evaluation</h2>
Next, let's import <b>metrics</b> from sklearn and check the accuracy of our model.
</div>
```
from sklearn import metrics
import matplotlib.pyplot as plt
print("DecisionTrees's Accuracy: ", metrics.accuracy_score(y_testset, predTree))
```
__Accuracy classification score__ computes subset accuracy: the set of labels predicted for a sample must exactly match the corresponding set of labels in y_true.
In multilabel classification, the function returns the subset accuracy. If the entire set of predicted labels for a sample strictly match with the true set of labels, then the subset accuracy is 1.0; otherwise it is 0.0.
## Practice
Can you calculate the accuracy score without sklearn ?
```
# your code here
```
<hr>
<div id="visualization">
<h2>Visualization</h2>
Lets visualize the tree
</div>
```
# Notice: You might need to uncomment and install the pydotplus and graphviz libraries if you have not installed these before
# !conda install -c conda-forge pydotplus -y
# !conda install -c conda-forge python-graphviz -y
from sklearn.externals.six import StringIO
import pydotplus
import matplotlib.image as mpimg
from sklearn import tree
%matplotlib inline
dot_data = StringIO()
filename = "drugtree.png"
featureNames = my_data.columns[0:5]
targetNames = my_data["Drug"].unique().tolist()
out=tree.export_graphviz(drugTree,feature_names=featureNames, out_file=dot_data, class_names= np.unique(y_trainset), filled=True, special_characters=True,rotate=False)
graph = pydotplus.graph_from_dot_data(dot_data.getvalue())
graph.write_png(filename)
img = mpimg.imread(filename)
plt.figure(figsize=(100, 200))
plt.imshow(img,interpolation='nearest')
```
<h2>Want to learn more?</h2>
IBM SPSS Modeler is a comprehensive analytics platform that has many machine learning algorithms. It has been designed to bring predictive intelligence to decisions made by individuals, by groups, by systems – by your enterprise as a whole. A free trial is available through this course, available here: <a href="http://cocl.us/ML0101EN-SPSSModeler">SPSS Modeler</a>
Also, you can use Watson Studio to run these notebooks faster with bigger datasets. Watson Studio is IBM's leading cloud solution for data scientists, built by data scientists. With Jupyter notebooks, RStudio, Apache Spark and popular libraries pre-packaged in the cloud, Watson Studio enables data scientists to collaborate on their projects without having to install anything. Join the fast-growing community of Watson Studio users today with a free account at <a href="https://cocl.us/ML0101EN_DSX">Watson Studio</a>
<h3>Thanks for completing this lesson!</h3>
<h4>Author: <a href="https://ca.linkedin.com/in/saeedaghabozorgi">Saeed Aghabozorgi</a></h4>
<p><a href="https://ca.linkedin.com/in/saeedaghabozorgi">Saeed Aghabozorgi</a>, PhD is a Data Scientist in IBM with a track record of developing enterprise level applications that substantially increases clients’ ability to turn data into actionable knowledge. He is a researcher in data mining field and expert in developing advanced analytic methods like machine learning and statistical modelling on large datasets.</p>
<hr>
<p>Copyright © 2018 <a href="https://cocl.us/DX0108EN_CC">Cognitive Class</a>. This notebook and its source code are released under the terms of the <a href="https://bigdatauniversity.com/mit-license/">MIT License</a>.</p>
| github_jupyter |
# TSG014 - Show BDC endpoints
## Steps
### Common functions
Define helper functions used in this notebook.
```
# Define `run` function for transient fault handling, hyperlinked suggestions, and scrolling updates on Windows
import sys
import os
import re
import platform
import shlex
import shutil
import datetime
from subprocess import Popen, PIPE
from IPython.display import Markdown
retry_hints = {} # Output in stderr known to be transient, therefore automatically retry
error_hints = {} # Output in stderr where a known SOP/TSG exists which will be HINTed for further help
install_hint = {} # The SOP to help install the executable if it cannot be found
def run(cmd, return_output=False, no_output=False, retry_count=0, base64_decode=False, return_as_json=False):
"""Run shell command, stream stdout, print stderr and optionally return output
NOTES:
1. Commands that need this kind of ' quoting on Windows e.g.:
kubectl get nodes -o jsonpath={.items[?(@.metadata.annotations.pv-candidate=='data-pool')].metadata.name}
Need to actually pass in as '"':
kubectl get nodes -o jsonpath={.items[?(@.metadata.annotations.pv-candidate=='"'data-pool'"')].metadata.name}
The ' quote approach, although correct when pasting into Windows cmd, will hang at the line:
`iter(p.stdout.readline, b'')`
The shlex.split call does the right thing for each platform, just use the '"' pattern for a '
"""
MAX_RETRIES = 5
output = ""
retry = False
# When running `azdata sql query` on Windows, replace any \n in """ strings, with " ", otherwise we see:
#
# ('HY090', '[HY090] [Microsoft][ODBC Driver Manager] Invalid string or buffer length (0) (SQLExecDirectW)')
#
if platform.system() == "Windows" and cmd.startswith("azdata sql query"):
cmd = cmd.replace("\n", " ")
# shlex.split is required on bash and for Windows paths with spaces
#
cmd_actual = shlex.split(cmd)
# Store this (i.e. kubectl, python etc.) to support binary context aware error_hints and retries
#
user_provided_exe_name = cmd_actual[0].lower()
# When running python, use the python in the ADS sandbox ({sys.executable})
#
if cmd.startswith("python "):
cmd_actual[0] = cmd_actual[0].replace("python", sys.executable)
# On Mac, when ADS is not launched from terminal, LC_ALL may not be set, which causes pip installs to fail
# with:
#
# UnicodeDecodeError: 'ascii' codec can't decode byte 0xc5 in position 4969: ordinal not in range(128)
#
# Setting it to a default value of "en_US.UTF-8" enables pip install to complete
#
if platform.system() == "Darwin" and "LC_ALL" not in os.environ:
os.environ["LC_ALL"] = "en_US.UTF-8"
# When running `kubectl`, if AZDATA_OPENSHIFT is set, use `oc`
#
if cmd.startswith("kubectl ") and "AZDATA_OPENSHIFT" in os.environ:
cmd_actual[0] = cmd_actual[0].replace("kubectl", "oc")
# To aid supportability, determine which binary file will actually be executed on the machine
#
which_binary = None
# Special case for CURL on Windows. The version of CURL in Windows System32 does not work to
# get JWT tokens, it returns "(56) Failure when receiving data from the peer". If another instance
# of CURL exists on the machine use that one. (Unfortunately the curl.exe in System32 is almost
# always the first curl.exe in the path, and it can't be uninstalled from System32, so here we
# look for the 2nd installation of CURL in the path)
if platform.system() == "Windows" and cmd.startswith("curl "):
path = os.getenv('PATH')
for p in path.split(os.path.pathsep):
p = os.path.join(p, "curl.exe")
if os.path.exists(p) and os.access(p, os.X_OK):
if p.lower().find("system32") == -1:
cmd_actual[0] = p
which_binary = p
break
# Find the path based location (shutil.which) of the executable that will be run (and display it to aid supportability), this
# seems to be required for .msi installs of azdata.cmd/az.cmd. (otherwise Popen returns FileNotFound)
#
# NOTE: Bash needs cmd to be the list of the space separated values hence shlex.split.
#
if which_binary == None:
which_binary = shutil.which(cmd_actual[0])
# Display an install HINT, so the user can click on a SOP to install the missing binary
#
if which_binary == None:
print(f"The path used to search for '{cmd_actual[0]}' was:")
print(sys.path)
if user_provided_exe_name in install_hint and install_hint[user_provided_exe_name] is not None:
display(Markdown(f'HINT: Use [{install_hint[user_provided_exe_name][0]}]({install_hint[user_provided_exe_name][1]}) to resolve this issue.'))
raise FileNotFoundError(f"Executable '{cmd_actual[0]}' not found in path (where/which)")
else:
cmd_actual[0] = which_binary
start_time = datetime.datetime.now().replace(microsecond=0)
print(f"START: {cmd} @ {start_time} ({datetime.datetime.utcnow().replace(microsecond=0)} UTC)")
print(f" using: {which_binary} ({platform.system()} {platform.release()} on {platform.machine()})")
print(f" cwd: {os.getcwd()}")
# Command-line tools such as CURL and AZDATA HDFS commands output
# scrolling progress bars, which causes Jupyter to hang forever, to
# workaround this, use no_output=True
#
# Work around a infinite hang when a notebook generates a non-zero return code, break out, and do not wait
#
wait = True
try:
if no_output:
p = Popen(cmd_actual)
else:
p = Popen(cmd_actual, stdout=PIPE, stderr=PIPE, bufsize=1)
with p.stdout:
for line in iter(p.stdout.readline, b''):
line = line.decode()
if return_output:
output = output + line
else:
if cmd.startswith("azdata notebook run"): # Hyperlink the .ipynb file
regex = re.compile(' "(.*)"\: "(.*)"')
match = regex.match(line)
if match:
if match.group(1).find("HTML") != -1:
display(Markdown(f' - "{match.group(1)}": "{match.group(2)}"'))
else:
display(Markdown(f' - "{match.group(1)}": "[{match.group(2)}]({match.group(2)})"'))
wait = False
break # otherwise infinite hang, have not worked out why yet.
else:
print(line, end='')
if wait:
p.wait()
except FileNotFoundError as e:
if install_hint is not None:
display(Markdown(f'HINT: Use {install_hint} to resolve this issue.'))
raise FileNotFoundError(f"Executable '{cmd_actual[0]}' not found in path (where/which)") from e
exit_code_workaround = 0 # WORKAROUND: azdata hangs on exception from notebook on p.wait()
if not no_output:
for line in iter(p.stderr.readline, b''):
try:
line_decoded = line.decode()
except UnicodeDecodeError:
# NOTE: Sometimes we get characters back that cannot be decoded(), e.g.
#
# \xa0
#
# For example see this in the response from `az group create`:
#
# ERROR: Get Token request returned http error: 400 and server
# response: {"error":"invalid_grant",# "error_description":"AADSTS700082:
# The refresh token has expired due to inactivity.\xa0The token was
# issued on 2018-10-25T23:35:11.9832872Z
#
# which generates the exception:
#
# UnicodeDecodeError: 'utf-8' codec can't decode byte 0xa0 in position 179: invalid start byte
#
print("WARNING: Unable to decode stderr line, printing raw bytes:")
print(line)
line_decoded = ""
pass
else:
# azdata emits a single empty line to stderr when doing an hdfs cp, don't
# print this empty "ERR:" as it confuses.
#
if line_decoded == "":
continue
print(f"STDERR: {line_decoded}", end='')
if line_decoded.startswith("An exception has occurred") or line_decoded.startswith("ERROR: An error occurred while executing the following cell"):
exit_code_workaround = 1
# inject HINTs to next TSG/SOP based on output in stderr
#
if user_provided_exe_name in error_hints:
for error_hint in error_hints[user_provided_exe_name]:
if line_decoded.find(error_hint[0]) != -1:
display(Markdown(f'HINT: Use [{error_hint[1]}]({error_hint[2]}) to resolve this issue.'))
# Verify if a transient error, if so automatically retry (recursive)
#
if user_provided_exe_name in retry_hints:
for retry_hint in retry_hints[user_provided_exe_name]:
if line_decoded.find(retry_hint) != -1:
if retry_count < MAX_RETRIES:
print(f"RETRY: {retry_count} (due to: {retry_hint})")
retry_count = retry_count + 1
output = run(cmd, return_output=return_output, retry_count=retry_count)
if return_output:
if base64_decode:
import base64
return base64.b64decode(output).decode('utf-8')
else:
return output
elapsed = datetime.datetime.now().replace(microsecond=0) - start_time
# WORKAROUND: We avoid infinite hang above in the `azdata notebook run` failure case, by inferring success (from stdout output), so
# don't wait here, if success known above
#
if wait:
if p.returncode != 0:
raise SystemExit(f'Shell command:\n\n\t{cmd} ({elapsed}s elapsed)\n\nreturned non-zero exit code: {str(p.returncode)}.\n')
else:
if exit_code_workaround !=0 :
raise SystemExit(f'Shell command:\n\n\t{cmd} ({elapsed}s elapsed)\n\nreturned non-zero exit code: {str(exit_code_workaround)}.\n')
print(f'\nSUCCESS: {elapsed}s elapsed.\n')
if return_output:
if base64_decode:
import base64
return base64.b64decode(output).decode('utf-8')
else:
return output
# Hints for tool retry (on transient fault), known errors and install guide
#
retry_hints = {'azdata': ['Endpoint sql-server-master does not exist', 'Endpoint livy does not exist', 'Failed to get state for cluster', 'Endpoint webhdfs does not exist', 'Adaptive Server is unavailable or does not exist', 'Error: Address already in use', 'Login timeout expired (0) (SQLDriverConnect)', 'SSPI Provider: No Kerberos credentials available', ], 'kubectl': ['A connection attempt failed because the connected party did not properly respond after a period of time, or established connection failed because connected host has failed to respond', ], 'python': [ ], }
error_hints = {'azdata': [['Please run \'azdata login\' to first authenticate', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['The token is expired', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['Reason: Unauthorized', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['Max retries exceeded with url: /api/v1/bdc/endpoints', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['Look at the controller logs for more details', 'TSG027 - Observe cluster deployment', '../diagnose/tsg027-observe-bdc-create.ipynb'], ['provided port is already allocated', 'TSG062 - Get tail of all previous container logs for pods in BDC namespace', '../log-files/tsg062-tail-bdc-previous-container-logs.ipynb'], ['Create cluster failed since the existing namespace', 'SOP061 - Delete a big data cluster', '../install/sop061-delete-bdc.ipynb'], ['Failed to complete kube config setup', 'TSG067 - Failed to complete kube config setup', '../repair/tsg067-failed-to-complete-kube-config-setup.ipynb'], ['Data source name not found and no default driver specified', 'SOP069 - Install ODBC for SQL Server', '../install/sop069-install-odbc-driver-for-sql-server.ipynb'], ['Can\'t open lib \'ODBC Driver 17 for SQL Server', 'SOP069 - Install ODBC for SQL Server', '../install/sop069-install-odbc-driver-for-sql-server.ipynb'], ['Control plane upgrade failed. Failed to upgrade controller.', 'TSG108 - View the controller upgrade config map', '../diagnose/tsg108-controller-failed-to-upgrade.ipynb'], ['NameError: name \'azdata_login_secret_name\' is not defined', 'SOP013 - Create secret for azdata login (inside cluster)', '../common/sop013-create-secret-for-azdata-login.ipynb'], ['ERROR: No credentials were supplied, or the credentials were unavailable or inaccessible.', 'TSG124 - \'No credentials were supplied\' error from azdata login', '../repair/tsg124-no-credentials-were-supplied.ipynb'], ['Please accept the license terms to use this product through', 'TSG126 - azdata fails with \'accept the license terms to use this product\'', '../repair/tsg126-accept-license-terms.ipynb'], ], 'kubectl': [['no such host', 'TSG010 - Get configuration contexts', '../monitor-k8s/tsg010-get-kubernetes-contexts.ipynb'], ['No connection could be made because the target machine actively refused it', 'TSG056 - Kubectl fails with No connection could be made because the target machine actively refused it', '../repair/tsg056-kubectl-no-connection-could-be-made.ipynb'], ], 'python': [['Library not loaded: /usr/local/opt/unixodbc', 'SOP012 - Install unixodbc for Mac', '../install/sop012-brew-install-odbc-for-sql-server.ipynb'], ['WARNING: You are using pip version', 'SOP040 - Upgrade pip in ADS Python sandbox', '../install/sop040-upgrade-pip.ipynb'], ], }
install_hint = {'azdata': [ 'SOP063 - Install azdata CLI (using package manager)', '../install/sop063-packman-install-azdata.ipynb' ], 'kubectl': [ 'SOP036 - Install kubectl command line interface', '../install/sop036-install-kubectl.ipynb' ], }
print('Common functions defined successfully.')
```
### Use azdata to list files
```
run('azdata bdc endpoint list')
print("Notebook execution is complete.")
```
| github_jupyter |
```
from PIL import Image
im = Image.open("img.png")
from __future__ import print_function
print(im.format, im.size, im.mode)
im_width, im_height = im.size
im = im.convert("1")
im.show()
import numpy as np
p = np.array(im,dtype=int)
n = int(input("Enter No of Participents: "))
print("Enter info about the access structure:")
# g0 = int(input("Enter the no of sub-sets in the access structure"))
# # for i in range(g0):
G = [[[1,2],[2,3]],[[1,4],[3,4]]]
print(G)
a,b = map(set,G[0])
# print("a,b: ",a,b)
in_section = a & b
# print("in_section: ", in_section)
union = (a | b) - in_section
# print("union: ",union)
s00 = np.zeros((n,2))
s01 = np.zeros((n,2))
for i in in_section:
s00[i-1,0]= s01[i-1,0] = 0
s00[i-1,1]= s01[i-1,1] = 1
for i in union:
s00[i-i,0]= s01[i-1,1] = 0
s00[i-1,1]= s01[i-1,0] = 1
print("s00: ",s00)
print()
print("s01 :",s01)
a,b = map(set,G[1])
# print("a,b: ",a,b)
in_section = a & b
# print("in_section: ", in_section)
union = (a | b) - in_section
# print("union: ",union)
s10 = np.zeros((n,2))
s11 = np.zeros((n,2))
for i in in_section:
s10[i-1,0]= s11[i-1,0] = 0
s10[i-1,1]= s11[i-1,1] = 1
for i in union:
s10[i-i,0]= s11[i-1,1] = 0
s10[i-1,1]= s11[i-1,0] = 1
print("s10: ",s10)
print()
print("s11 :",s11)
s0 = np.hstack((s00,s10))
s1 = np.hstack((s01,s11))
print(s0)
print()
print(s1)
```
## Random Permutation
```
print(s0)
print()
print(np.random.permutation(s0.T).T)
print (s0[:, np.random.permutation(s0.shape[1])])
np.random.permutation(s0.shape[1])
s0[:,[0,2,3,1]]
np.random.permutation(s0.shape[1])
print(s0[1,:])
s0[1,:].reshape((2,2))
```
## Share Gen
```
import matplotlib.pyplot as plt
share1 = np.zeros([2*im_width, 2* im_height], dtype=int)
share2 = np.zeros([2*im_width, 2* im_height], dtype=int)
share3 = np.zeros([2*im_width, 2* im_height], dtype=int)
share4 = np.zeros([2*im_width, 2* im_height], dtype=int)
# plt.imshow(share1)
# im_share1 = Image.fromarray(share1)
# im_share1.save("share1.pbm")
irow = 0
icol = 0
for py in p:
for px in py:
if px == 0:
s1 = np.random.permutation(s1.T).T
share1[irow, icol] = s1[0][0]
share2[irow, icol] = s1[1][0]
share3[irow, icol] = s1[2][0]
share4[irow, icol] = s1[3][0]
share1[irow+1, icol] = s1[0][2]
share2[irow+1, icol] = s1[1][2]
share3[irow+1, icol] = s1[2][2]
share4[irow+1, icol] = s1[3][2]
share1[irow+1, icol+1] = s1[0][3]
share2[irow+1, icol+1] = s1[1][3]
share3[irow+1, icol+1] = s1[2][3]
share4[irow+1, icol+1] = s1[3][3]
share1[irow, icol+1] = s1[0][1]
share2[irow, icol+1] = s1[1][1]
share3[irow, icol+1] = s1[2][1]
share4[irow, icol+1] = s1[3][1]
icol += 2
else :
s0 = np.random.permutation(s0.T).T
share1[irow, icol] = s0[0][0]
share2[irow, icol] = s0[1][0]
share3[irow, icol] = s0[2][0]
share4[irow, icol] = s0[3][0]
share1[irow+1, icol] = s0[0][2]
share2[irow+1, icol] = s0[1][2]
share3[irow+1, icol] = s0[2][2]
share4[irow+1, icol] = s0[3][2]
share1[irow+1, icol+1] = s0[0][3]
share2[irow+1, icol+1] = s0[1][3]
share3[irow+1, icol+1] = s0[2][3]
share4[irow+1, icol+1] = s0[3][3]
share1[irow, icol+1] = s0[0][1]
share2[irow, icol+1] = s0[1][1]
share3[irow, icol+1] = s0[2][1]
share4[irow, icol+1] = s0[3][1]
icol += 2
irow +=2
icol = 0
plt.imshow(share1)
plt.imshow(share2)
plt.imshow(share3)
plt.imshow(share4)
```
## Combine Share
```
x,y = share1.shape
secret = np.ones([x,y],dtype=int)
for i in range(x):
for j in range(y):
secret[i,j] = (share1[i,j]+share2[i,j])%2
plt.imshow(secret)
for i in range(x):
for j in range(y):
secret[i,j] = (share3[i,j]+share2[i,j])%2
plt.imshow(secret)
for i in range(x):
for j in range(y):
secret[i,j] = (share1[i,j]+share3[i,j])%2
plt.imshow(secret)
for i in range(x):
for j in range(y):
secret[i,j] = (share1[i,j]+share4[i,j])%2
plt.imshow(secret)
for i in range(x):
for j in range(y):
secret[i,j] = (share3[i,j]+share4[i,j])%2
plt.imshow(secret)
for i in range(x):
for j in range(y):
secret[i,j] = (share2[i,j]+share4[i,j])%2
plt.imshow(secret)
for i in range(x):
for j in range(y):
secret[i,j] = share3[i,j] or share2[i,j] or share1[i,j] or share4[i,j]
plt.imshow(secret)
img = Image.fromarray(secret)
img.show()
for i in range(x):
for j in range(y):
secret[i,j] = ( share2[i,j] + share1[i,j])%2
plt.imshow(secret, "Gray")
for i in range(x):
for j in range(y):
secret[i,j] = ( share2[i,j] or share1[i,j])
plt.imshow(secret)
```
| github_jupyter |
# Monte Carlo Methods
In this notebook, you will write your own implementations of many Monte Carlo (MC) algorithms.
While we have provided some starter code, you are welcome to erase these hints and write your code from scratch.
### Part 0: Explore BlackjackEnv
We begin by importing the necessary packages.
```
import sys
import gym
import numpy as np
from collections import defaultdict
from plot_utils import plot_blackjack_values, plot_policy
```
Use the code cell below to create an instance of the [Blackjack](https://github.com/openai/gym/blob/master/gym/envs/toy_text/blackjack.py) environment.
```
env = gym.make('Blackjack-v0')
```
Each state is a 3-tuple of:
- the player's current sum $\in \{0, 1, \ldots, 31\}$,
- the dealer's face up card $\in \{1, \ldots, 10\}$, and
- whether or not the player has a usable ace (`no` $=0$, `yes` $=1$).
The agent has two potential actions:
```
STICK = 0
HIT = 1
```
Verify this by running the code cell below.
```
print(env.observation_space)
print(env.action_space)
```
Execute the code cell below to play Blackjack with a random policy.
(_The code currently plays Blackjack three times - feel free to change this number, or to run the cell multiple times. The cell is designed for you to get some experience with the output that is returned as the agent interacts with the environment._)
```
for i_episode in range(3):
state = env.reset()
while True:
print(state)
action = env.action_space.sample()
print(action)
state, reward, done, info = env.step(action)
if done:
print('End game! Reward: ', reward)
print('You won :)\n') if reward > 0 else print('You lost :(\n')
break
```
### Part 1: MC Prediction
In this section, you will write your own implementation of MC prediction (for estimating the action-value function).
We will begin by investigating a policy where the player _almost_ always sticks if the sum of her cards exceeds 18. In particular, she selects action `STICK` with 80% probability if the sum is greater than 18; and, if the sum is 18 or below, she selects action `HIT` with 80% probability. The function `generate_episode_from_limit_stochastic` samples an episode using this policy.
The function accepts as **input**:
- `bj_env`: This is an instance of OpenAI Gym's Blackjack environment.
It returns as **output**:
- `episode`: This is a list of (state, action, reward) tuples (of tuples) and corresponds to $(S_0, A_0, R_1, \ldots, S_{T-1}, A_{T-1}, R_{T})$, where $T$ is the final time step. In particular, `episode[i]` returns $(S_i, A_i, R_{i+1})$, and `episode[i][0]`, `episode[i][1]`, and `episode[i][2]` return $S_i$, $A_i$, and $R_{i+1}$, respectively.
```
def generate_episode_from_limit_stochastic(bj_env):
episode = []
state = bj_env.reset()
while True:
probs = [0.8, 0.2] if state[0] > 18 else [0.2, 0.8]
action = np.random.choice(np.arange(2), p=probs)
next_state, reward, done, info = bj_env.step(action)
episode.append((state, action, reward))
state = next_state
if done:
break
return episode
```
Execute the code cell below to play Blackjack with the policy.
(*The code currently plays Blackjack three times - feel free to change this number, or to run the cell multiple times. The cell is designed for you to gain some familiarity with the output of the `generate_episode_from_limit_stochastic` function.*)
```
for i in range(3):
episode = generate_episode_from_limit_stochastic(env)
print("Episode:", episode)
states, actions, rewards = zip(*episode)
print("states:",states,"actions:",actions,"rewards:",rewards)
```
Now, you are ready to write your own implementation of MC prediction. Feel free to implement either first-visit or every-visit MC prediction; in the case of the Blackjack environment, the techniques are equivalent.
Your algorithm has three arguments:
- `env`: This is an instance of an OpenAI Gym environment.
- `num_episodes`: This is the number of episodes that are generated through agent-environment interaction.
- `generate_episode`: This is a function that returns an episode of interaction.
- `gamma`: This is the discount rate. It must be a value between 0 and 1, inclusive (default value: `1`).
The algorithm returns as output:
- `Q`: This is a dictionary (of one-dimensional arrays) where `Q[s][a]` is the estimated action value corresponding to state `s` and action `a`.
```
def mc_prediction_q(env, num_episodes, generate_episode, gamma=1.0):
# initialize empty dictionaries of arrays
returns_sum = defaultdict(lambda: np.zeros(env.action_space.n))
N = defaultdict(lambda: np.zeros(env.action_space.n))
Q = defaultdict(lambda: np.zeros(env.action_space.n))
# loop over episodes
for i_episode in range(1, num_episodes+1):
# monitor progress
if i_episode % 1000 == 0:
print("\rEpisode {}/{}.".format(i_episode, num_episodes), end="")
sys.stdout.flush()
## TODO: complete the function
episode = generate_episode(env)
states, actions, rewards = zip(*episode)
discounts = np.array([gamma**i for i in range(len(rewards)+1)])
for i, state in enumerate(states):
returns_sum[state][actions[i]] += sum(rewards[i:]*discounts[:-(i+1)])
N[state][actions[i]]+=1
Q[state][actions[i]] = returns_sum[state][actions[i]]/N[state][actions[i]]
return Q
```
Use the cell below to obtain the action-value function estimate $Q$. We have also plotted the corresponding state-value function.
To check the accuracy of your implementation, compare the plot below to the corresponding plot in the solutions notebook **Monte_Carlo_Solution.ipynb**.
```
# obtain the action-value function
Q = mc_prediction_q(env, 500000, generate_episode_from_limit_stochastic)
# obtain the corresponding state-value function
V_to_plot = dict((k,(k[0]>18)*(np.dot([0.8, 0.2],v)) + (k[0]<=18)*(np.dot([0.2, 0.8],v))) \
for k, v in Q.items())
# plot the state-value function
plot_blackjack_values(V_to_plot)
```
### Part 2: MC Control
In this section, you will write your own implementation of constant-$\alpha$ MC control.
Your algorithm has four arguments:
- `env`: This is an instance of an OpenAI Gym environment.
- `num_episodes`: This is the number of episodes that are generated through agent-environment interaction.
- `alpha`: This is the step-size parameter for the update step.
- `gamma`: This is the discount rate. It must be a value between 0 and 1, inclusive (default value: `1`).
The algorithm returns as output:
- `Q`: This is a dictionary (of one-dimensional arrays) where `Q[s][a]` is the estimated action value corresponding to state `s` and action `a`.
- `policy`: This is a dictionary where `policy[s]` returns the action that the agent chooses after observing state `s`.
(_Feel free to define additional functions to help you to organize your code._)
```
#np.random.seed(0)
## TODO: Define necessary functions
class Policy:
def __init__(self, Q, epsilon, nA):
self.Q = Q
self.eps = epsilon
self.num_actions = nA
def sample_action(self, state):
if state in Q:
best_action = np.argmax(self.Q[state])
if np.random.uniform()>self.eps:
return best_action
return np.random.choice(np.arange(self.num_actions))
def gen_episode(env, policy):
episode=[]
state = env.reset()
while True:
action = policy.sample_action(state)
state, reward, done, info = env.step(action)
episode.append((state, action, reward))
if done:
break
return episode
def update_Q(Q, episode, alpha, gamma):
states, actions, rewards = zip(*episode)
discounts = np.array([gamma**i for i in range(len(rewards)+1)])
for i, state in enumerate(states):
old_q = Q[state][actions[i]]
ret = sum(rewards[i:]*discounts[:-(i+1)])
Q[state][actions[i]] = old_q + alpha * (ret-old_q)
return Q
## TODO: complete the function
def mc_control(env, num_episodes, alpha, gamma=1.0, eps_decay=.99999):
nA = env.action_space.n
# initialize empty dictionary of arrays
Q = defaultdict(lambda: np.zeros(nA))
# start epsilon, final_epsilon
epsilon, eps_min = 1.0, 0.0
# loop over episodes
for i_episode in range(1, num_episodes+1):
# monitor progress
if i_episode % 1000 == 0:
print("\rEpisode {}/{}.".format(i_episode, num_episodes), end="")
sys.stdout.flush()
epsilon = max(epsilon, eps_min)
policy = Policy(Q, epsilon, nA)
episode = gen_episode(env, policy)
Q = update_Q(Q, episode, alpha, gamma)
epsilon = epsilon * eps_decay
policy = dict((state, np.argmax(values)) for state, values in Q.items())
return policy, Q
```
Use the cell below to obtain the estimated optimal policy and action-value function. Note that you should fill in your own values for the `num_episodes` and `alpha` parameters.
```
# obtain the estimated optimal policy and action-value function
policy, Q = mc_control(env, 500000, 0.02)
```
Next, we plot the corresponding state-value function.
```
# obtain the corresponding state-value function
V = dict((k,np.max(v)) for k, v in Q.items())
# plot the state-value function
plot_blackjack_values(V)
```
Finally, we visualize the policy that is estimated to be optimal.
```
# plot the policy
plot_policy(policy)
```
The **true** optimal policy $\pi_*$ can be found in Figure 5.2 of the [textbook](http://go.udacity.com/rl-textbook) (and appears below). Compare your final estimate to the optimal policy - how close are you able to get? If you are not happy with the performance of your algorithm, take the time to tweak the decay rate of $\epsilon$, change the value of $\alpha$, and/or run the algorithm for more episodes to attain better results.

| github_jupyter |
# Video 01: Gradient Descent
video: https://www.youtube.com/watch?v=xRJCOz3AfYY&list=PL2-dafEMk2A7mu0bSksCGMJEmeddU_H4D
source code: https://github.com/llSourcell/Intro_to_the_Math_of_intelligence/blob/master/demo.py
gradient descent for line fitting:

## Sum of Squares Error Function
$$sse(m,b) = \frac{1}{n} \sum_{i=1}^{n} (y_i - (m x_i + b))^2 $$
We need to minimize $sse$ wrt $m$ and $b$.
Thus we take derivative of $sse$ wrt $m$ and $b$ and use gradient descent to approach to zero slope.
$$
\begin{align}
\frac{\partial sse}{\partial m} & = \frac{2}{n} \sum_i^n - x_i (y_i - (m x_i + b)) \\
\frac{\partial sse}{\partial b} & = \frac{2}{n} \sum_i^n - (y_i - (m x_i + b))
\end{align}
$$
Note that gradients depend on n points: $(x_i, y_i)$
So, the code to calculate gradients will loop over all points:
def step_gradient(b_current, m_current, points, learningRate):
b_gradient = 0
m_gradient = 0
N = float(len(points))
for i in range(0, len(points)):
x = points[i, 0]
y = points[i, 1]
b_gradient += -(2/N) * (y - ((m_current * x) + b_current))
m_gradient += -(2/N) * x * (y - ((m_current * x) + b_current))
```
from numpy import *
# y = mx + b
# m is slope, b is y-intercept
def compute_error_for_line_given_points(b, m, points):
totalError = 0
for i in range(0, len(points)):
x = points[i, 0]
y = points[i, 1]
totalError += (y - (m * x + b)) ** 2
return totalError / float(len(points))
def step_gradient(b_current, m_current, points, learningRate):
b_gradient = 0
m_gradient = 0
N = float(len(points))
for i in range(0, len(points)):
x = points[i, 0]
y = points[i, 1]
b_gradient += -(2/N) * (y - ((m_current * x) + b_current))
m_gradient += -(2/N) * x * (y - ((m_current * x) + b_current))
new_b = b_current - (learningRate * b_gradient)
new_m = m_current - (learningRate * m_gradient)
return [new_b, new_m]
def gradient_descent_runner(points, starting_b, starting_m, learning_rate, num_iterations):
b = starting_b
m = starting_m
for i in range(num_iterations):
b, m = step_gradient(b, m, array(points), learning_rate)
return [b, m]
def run(points):
learning_rate = 0.0001
initial_b = 0 # initial y-intercept guess
initial_m = 0 # initial slope guess
num_iterations = 1000
[b, m] = gradient_descent_runner(points, initial_b, initial_m, learning_rate, num_iterations)
return [b,m]
points = genfromtxt("../../../study_data/math_of_intelligence/01_data.csv", delimiter=",")
run(points)
```
## Video 02: Support Vector Machines
use cases for svm:
- classification, regression, outlier detection, clustering
### Comparison to other methods
svm good when:
- small set of data (<1000 rows)
other algorithms (random forest, dnn etc)
- more data
- always very robust
Knuth: "Premature optimization is the root of all evil in programming"
### What is svm?
a discriminative classifier
opposite to discriminative: generative. it generates new data.
svm tries to maximise the gap between two classes of points
these points: support vectors
hyperplane: the line that separates the two classes of points
hyperplane is (n-1) dimensional in $\rm I\!R^n$
### Linear vs nonlinear classification
to do non-linear classification: kernel trick
### Loss function
Minimize loss function to maximize the discrimination.
Hinge loss function: for maximum margin classification
$$c(x,y,f(x)) = (1 - y \times f(x))_+$$
c is the loss function. x the sample. y true label. f(x) predicted label.
So:
$$
c(x,y,f(x)) =
\begin{cases}
0,& \text{if } y \times f(x) \geq 1 \\
1 - y \times f(x),& \text{else}
\end{cases}
$$
#### Objective function
$$min_w \lambda \| w \|^2 + \sum_{i=1}^n (1-y_i \langle x_i, w \rangle )_+$$
note that: $w = f(x)$
objective function = regularizer + hinge loss
what does regularizer do?
- too high => overfit
- too low => underfit
so, regularizer controls trade off between low training error and low testing error
| github_jupyter |
| [**Overview**](./00_overview.ipynb) | **Examples:** | [Selecting and Indexing Geochem Data](01_indexes_selectors.ipynb) | [Data Munging](02_munging.ipynb) | [Visualisation](03_visualisation.ipynb) |[lambdas](04_lambdas.ipynb) |
|:-----|:-----|:-----|:-----|:-----|:-----|
# lambdas: Parameterising REE Profiles
Orthogonal polynomial decomposition can be used for dimensional reduction of smooth
function over an independent variable, producing an array of independent values
representing the relative weights for each order of component polynomial. This is an
effective method to parameterise and compare the nature of smooth profiles.
In geochemistry, the most applicable use case is for reduction Rare Earth Element (REE)
profiles. The REE are a collection of elements with broadly similar physicochemical
properties (the lanthanides), which vary with ionic radii. Given their similar behaviour
and typically smooth function of normalised abundance vs. ionic radii, the REE profiles
and their shapes can be effectively parameterised and dimensionally reduced (14 elements
summarised by 3-4 shape parameters).
Here we generate some example data, reduce these to lambda values, and visualise the
results.
<div class='alert alert-success'> <b>Note:</b> this example is taken largely from the current pyrolite example for lambdas. As the next version is released, check back there for an updated example!</div>
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import pyrolite.plot
np.random.seed(82)
```
First we'll generate some example **synthetic data** based around Depleted MORB Mantle:
```
from pyrolite.util.synthetic import example_spider_data
df = example_spider_data(
noise_level=0.05,
size=100,
start="DM_SS2004",
norm_to="ChondriteREE_ON",
offsets={"Eu": 0.2},
)
df = df.pyrochem.denormalize_from("ChondriteREE_ON") # start with unnormalised data!
```
Let's have a quick look at what this REE data looks like normalized to Primitive
Mantle:
```
df.pyrochem.normalize_to("ChondriteREE_ON").pyroplot.REE(alpha=0.05, c="k", unity_line=True)
plt.show()
```
From this REE data we can fit a series of orthogonal polynomials, and subsequently used
the regression coefficients ('lambdas') as a parameterisation of the REE
pattern/profile:
```
ls = df.pyrochem.lambda_lnREE(degree=4)
ls.head(2)
```
So what's actually happening here? To get some idea of what these λ coefficients
correspond to, we can pull this process apart and visualise our REE profiles as
the sum of the series of orthogonal polynomial components of increasing order.
As lambdas represent the coefficients for the regression of log-transformed normalised
data, to compare the polynomial components and our REE profile we'll first need to
normalize it to the appropriate composition (here `"ChondriteREE_ON"`) before
taking the logarithm.
With our data, we've then fit a function of ionic radius with the form
$f(r) = \lambda_0 + \lambda_1 f_1 + \lambda_2 f_2 + \lambda_3 f_3...$
where the polynomial components of increasing order are $f_1 = (r - \beta_0)$,
$f_2 = (r - \gamma_0)(r - \gamma_1)$,
$f_3 = (r - \delta_0)(r - \delta_1)(r - \delta_2)$ and so on. The parameters
$\beta$, $\gamma$, $\delta$ are pre-computed such that the
polynomial components are indeed independent. Here we can visualise how these
polynomial components are summed to produce the regressed profile, using the last REE
profile we generated above as an example:
```
from pyrolite.util.lambdas.plot import plot_lambdas_components
ax = (
df.pyrochem.normalize_to("ChondriteREE_ON")
.iloc[-1, :]
.apply(np.log)
.pyroplot.REE(color="k", label="Data", logy=False)
)
plot_lambdas_components(ls.iloc[-1, :], ax=ax)
ax.legend()
plt.show()
```
Now that we've gone through a brief introduction to how the lambdas are generated,
let's quickly check what the coefficient values themselves look like:
```
fig, ax = plt.subplots(1, 3, figsize=(9, 3))
for ix in range(ls.columns.size - 1):
ls[ls.columns[ix : ix + 2]].pyroplot.scatter(ax=ax[ix], alpha=0.1, c="k")
plt.tight_layout()
```
But what do these parameters correspond to? From the deconstructed orthogonal
polynomial above, we can see that $\lambda_0$ parameterises relative enrichment
(this is the mean value of the logarithm of Chondrite-normalised REE abundances),
$\lambda_1$ parameterises a linear slope (here, LREE enrichment), and higher
order terms describe curvature of the REE pattern. Through this parameterisation,
the REE profile can be effectively described and directly linked to geochemical
processes. While the amount of data we need to describe the patterns is lessened,
the values themselves are more meaningful and readily used to describe the profiles
and their physical significance.
The visualisation of $\lambda_1$-$\lambda_2$ can be particularly useful
where you're trying to compare REE profiles.
We've used a synthetic dataset here which is by design approximately normally
distributed, so the values themeselves here are not particularly revealing,
but they do illustrate the expected mangitudes of values for each of the parameters.
## Dealing With Anomalies
Note that we've not used Eu in this regression - Eu anomalies are a deviation from
the 'smooth profile' we need to use this method. Consider this if your data might also
exhibit significant Ce anomalies, you might need to exclude this data. For convenience
there is also functionality to calculate anomalies derived from the orthogonal
polynomial fit itself (rather than linear interpolation methods). Below we use the
`anomalies` keyword argument to also calculate the $\frac{Ce}{Ce*}$
and $\frac{Eu}{Eu*}$ anomalies (note that these are excluded from the fit):
```
ls_anomalies = df.pyrochem.lambda_lnREE(anomalies=["Ce", "Eu"])
ax = ls_anomalies.iloc[:, -2:].pyroplot.scatter()
plt.show()
```
## Fitting Tetrads
In addition to fitting orothogonal polynomial functions, the ability to fit tetrad
functions has also recently been added. This supplements the $\lambda$
coefficients with $\tau$ coefficients which describe subtle electronic
configuration effects affecting sequential subsets of the REE. Below we plot four
profiles - each describing only a single tetrad - to illustrate the shape of
these function components. Note that these are functions of $z$, and are here
transformed to plot against radii.
```
from pyrolite.util.lambdas.plot import plot_profiles
# let's first create some synthetic pattern parameters
# we want lambdas to be zero, and each of the tetrads to be shown in only one pattern
lambdas = np.zeros((4, 5))
tetrads = np.eye(4)
# putting it together to generate four sets of combined parameters
fit_parameters = np.hstack([lambdas, tetrads])
ax = plot_profiles(
fit_parameters,
tetrads=True,
color=np.arange(4),
)
plt.show()
```
In order to also fit these function components, you can pass the keyword argument
`fit_tetrads=True` to `pyrolite.geochem.pyrochem.lambda_lnREE` and
related functions:
```
lts = df.pyrochem.lambda_lnREE(degree=4, fit_tetrads=True)
```
We can see that the four extra $\tau$ Parameters have been appended to the
right of the lambdas within the output:
```
lts.head(2)
```
Below we'll look at some of the potential issues of fitting lambdas and tetrads
together - by examining the effects of i) fitting tetrads where there are none
and ii) not fitting tetrads where they do indeed exist using some synthetic datasets.
```
from pyrolite.util.synthetic import example_patterns_from_parameters
from pyrolite.geochem.ind import REE
ls = np.array(
[
[2, 5, -30, 100, -600, 0, 0, 0, 0], # lambda-only
[3, 15, 30, 300, 1500, 0, 0, 0, 0], # lambda-only
[1, 5, -50, 0, -1000, -0.3, -0.7, -1.4, -0.2], # W-pattern tetrad
[5, 15, 50, 400, 2000, 0.6, 1.1, 1.5, 0.3], # M-pattern tetrad
]
)
# now we use these parameters to generate some synthetic log-scaled normalised REE
# patterns and add a bit of noise
pattern_df = pd.DataFrame(
np.vstack([example_patterns_from_parameters(l, includes_tetrads=True) for l in ls]),
columns=REE(),
)
# We can now fit these patterns and see what the effect of fitting and not-Fitting
# tetrads might look like in these (slightly extreme) cases:
fit_ls_only = pattern_df.pyrochem.lambda_lnREE(
norm_to=None, degree=5, fit_tetrads=False
)
fit_ts = pattern_df.pyrochem.lambda_lnREE(norm_to=None, degree=5, fit_tetrads=True)
```
We can now examine what the differences between the fits are. Below we plot the four
sets of synthetic REE patterns (lambda-only above and lamba+tetrad below) and examine
the relative accuracy of fitting some of the higher order lambda parameters where
tetrads are also fit:
```
from pyrolite.util.plot.axes import share_axes
x, y = 2, 3
categories = np.repeat(np.arange(ls.shape[0]), 100)
colors = np.array([str(ix) * 2 for ix in categories])
l_only = categories < 2
ax = plt.figure(figsize=(12, 7)).subplot_mosaic(
"""
AAABBCC
DDDEEFF
"""
)
share_axes([ax["A"], ax["D"]])
share_axes([ax["B"], ax["C"], ax["E"], ax["F"]])
ax["B"].set_title("lambdas only Fit")
ax["C"].set_title("lambdas+tetrads Fit")
for a, fltr in zip(["A", "D"], [l_only, ~l_only]):
pattern_df.iloc[fltr, :].pyroplot.spider(
ax=ax[a],
label="True",
unity_line=True,
alpha=0.5,
color=colors[fltr],
)
for a, fltr in zip(["B", "E"], [l_only, ~l_only]):
fit_ls_only.iloc[fltr, [x, y]].pyroplot.scatter(
ax=ax[a],
alpha=0.2,
color=colors[fltr],
)
for a, fltr in zip(["C", "F"], [l_only, ~l_only]):
fit_ts.iloc[fltr, [x, y]].pyroplot.scatter(
ax=ax[a],
alpha=0.2,
color=colors[fltr],
)
true = pd.DataFrame(ls[:, [x, y]], columns=[fit_ls_only.columns[ix] for ix in [x, y]])
for ix, a in enumerate(["B", "C", "E", "F"]):
true.iloc[np.array([ix < 2, ix < 2, ix >= 2, ix >= 2]), :].pyroplot.scatter(
ax=ax[a],
color=np.array([str(ix) * 2 for ix in np.arange(ls.shape[0 ]// 2)]),
marker="X",
s=100,
)
plt.tight_layout()
plt.show()
```
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/FeatureCollection/column_info.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/FeatureCollection/column_info.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/FeatureCollection/column_info.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API and geemap
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geemap](https://github.com/giswqs/geemap). The **geemap** Python package is built upon the [ipyleaflet](https://github.com/jupyter-widgets/ipyleaflet) and [folium](https://github.com/python-visualization/folium) packages and implements several methods for interacting with Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, and `Map.centerObject()`.
The following script checks if the geemap package has been installed. If not, it will install geemap, which automatically installs its [dependencies](https://github.com/giswqs/geemap#dependencies), including earthengine-api, folium, and ipyleaflet.
**Important note**: A key difference between folium and ipyleaflet is that ipyleaflet is built upon ipywidgets and allows bidirectional communication between the front-end and the backend enabling the use of the map to capture user input, while folium is meant for displaying static data only ([source](https://blog.jupyter.org/interactive-gis-in-jupyter-with-ipyleaflet-52f9657fa7a)). Note that [Google Colab](https://colab.research.google.com/) currently does not support ipyleaflet ([source](https://github.com/googlecolab/colabtools/issues/60#issuecomment-596225619)). Therefore, if you are using geemap with Google Colab, you should use [`import geemap.eefolium`](https://github.com/giswqs/geemap/blob/master/geemap/eefolium.py). If you are using geemap with [binder](https://mybinder.org/) or a local Jupyter notebook server, you can use [`import geemap`](https://github.com/giswqs/geemap/blob/master/geemap/geemap.py), which provides more functionalities for capturing user input (e.g., mouse-clicking and moving).
```
# Installs geemap package
import subprocess
try:
import geemap
except ImportError:
print('geemap package not installed. Installing ...')
subprocess.check_call(["python", '-m', 'pip', 'install', 'geemap'])
# Checks whether this notebook is running on Google Colab
try:
import google.colab
import geemap.eefolium as geemap
except:
import geemap
# Authenticates and initializes Earth Engine
import ee
try:
ee.Initialize()
except Exception as e:
ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
The default basemap is `Google Maps`. [Additional basemaps](https://github.com/giswqs/geemap/blob/master/geemap/basemaps.py) can be added using the `Map.add_basemap()` function.
```
Map = geemap.Map(center=[40,-100], zoom=4)
Map
```
## Add Earth Engine Python script
```
# Add Earth Engine dataset
def getCols(tableMetadata):
return tableMetadata.columns
# Import a protected areas point feature collection.
wdpa = ee.FeatureCollection("WCMC/WDPA/current/points")
# Define a function to print metadata column names and datatypes. This function
# is intended to be applied by the `evaluate` method which provides the
# function a client-side dictionary allowing the 'columns' object of the
# feature collection metadata to be subset by dot notation or bracket notation
# (`tableMetadata['columns']`).
# function getCols(tableMetadata) {
# print(tableMetadata.columns)
# }
# Fetch collection metadata (`.limit(0)`) and apply the
# previously defined function using `evaluate()`. The printed object is a
# dictionary where keys are column names and values are datatypes.
# print(getCols(wdpa.limit(0)).getInfo())
```
## Display Earth Engine data layers
```
Map.addLayerControl() # This line is not needed for ipyleaflet-based Map.
Map
```
| github_jupyter |
# Tutorial 2: Inside CrypTensors
Note: This tutorial is optional, and can be skipped without any loss of continuity to the following tutorials.
In this tutorial, we will take a brief look at the internals of ```CrypTensors```.
Using the `mpc` backend, a `CrypTensor` is a tensor encrypted using secure MPC protocols, called an `MPCTensor`. In order to support the mathematical operations required by the `MPCTensor`, CrypTen implements two kinds of secret-sharing protocols: arithmetic secret-sharing and binary secret-sharing. Arithmetic secret sharing forms the basis for most of the mathematical operations implemented by `MPCTensor`. Similarly, binary secret-sharing allows for the evaluation of logical expressions.
In this tutorial, we'll first introduce the concept of a `CrypTensor` <i>ptype</i> (i.e. <i>private-type</i>), and show how to use it to obtain `MPCTensors` that use arithmetic and binary secret shares. We will also describe how each of these <i>ptypes</i> is used, and how they can be combined to implement desired functionality.
```
#import the libraries
import crypten
import torch
#initialize crypten
crypten.init()
#Disables OpenMP threads -- needed by @mpc.run_multiprocess which uses fork
torch.set_num_threads(1)
```
## <i>ptype</i> in CrypTen
CrypTen defines the `ptype` (for <i>private-type</i>) attribute of an `MPCTensor` to denote the kind of secret-sharing protocol used in the `CrypTensor`. The `ptype` is, in many ways, analogous to the `dtype` of PyTorch. The `ptype` may have two values:
- `crypten.mpc.arithmetic` for `ArithmeticSharedTensors`</li>
- `crypten.mpc.binary` for `BinarySharedTensors`</li>
We can use the `ptype` attribute to create a `CrypTensor` with the appropriate secret-sharing protocol. For example:
```
#Constructing CrypTensors with ptype attribute
#arithmetic secret-shared tensors
x_enc = crypten.cryptensor([1.0, 2.0, 3.0], ptype=crypten.mpc.arithmetic)
print("x_enc internal type:", x_enc.ptype)
#binary secret-shared tensors
y = torch.tensor([1, 2, 1], dtype=torch.int32)
y_enc = crypten.cryptensor(y, ptype=crypten.mpc.binary)
print("y_enc internal type:", y_enc.ptype)
```
### Arithmetic secret-sharing
Let's look more closely at the `crypten.mpc.arithmetic` <i>ptype</i>. Most of the mathematical operations implemented by `CrypTensors` are implemented using arithmetic secret sharing. As such, `crypten.mpc.arithmetic` is the default <i>ptype</i> for newly generated `CrypTensors`.
Let's begin by creating a new `CrypTensor` using `ptype=crypten.mpc.arithmetic` to enforce that the encryption is done via arithmetic secret sharing. We can print values of each share to confirm that values are being encrypted properly.
To do so, we will need to create multiple parties to hold each share. We do this here using the `@mpc.run_multiprocess` function decorator, which we developed to execute crypten code from a single script (as we have in a Jupyter notebook). CrypTen follows the standard MPI programming model: it runs a separate process for each party, but each process runs an identical (complete) program. Each process has a `rank` variable to identify itself.
Note that the sum of the two `_tensor` attributes below is equal to a scaled representation of the input. (Because MPC requires values to be integers, we scale input floats to a fixed-point encoding before encryption.)
```
import crypten.mpc as mpc
import crypten.communicator as comm
@mpc.run_multiprocess(world_size=2)
def examine_arithmetic_shares():
x_enc = crypten.cryptensor([1, 2, 3], ptype=crypten.mpc.arithmetic)
rank = comm.get().get_rank()
crypten.print(f"\nRank {rank}:\n {x_enc}\n", in_order=True)
x = examine_arithmetic_shares()
```
### Binary secret-sharing
The second type of secret-sharing implemented in CrypTen is binary or XOR secret-sharing. This type of secret-sharing allows greater efficiency in evaluating logical expressions.
Let's look more closely at the `crypten.mpc.binary` <i>ptype</i>. Most of the logical operations implemented by `CrypTensors` are implemented using arithmetic secret sharing. We typically use this type of secret-sharing when we want to evaluate binary operators (i.e. `^ & | >> <<`, etc.) or logical operations (like comparitors).
Let's begin by creating a new `CrypTensor` using `ptype=crypten.mpc.binary` to enforce that the encryption is done via binary secret sharing. We can print values of each share to confirm that values are being encrypted properly, as we did for arithmetic secret-shares.
(Note that an xor of the two `_tensor` attributes below is equal to an unscaled version of input.)
```
@mpc.run_multiprocess(world_size=2)
def examine_binary_shares():
x_enc = crypten.cryptensor([2, 3], ptype=crypten.mpc.binary)
rank = comm.get().get_rank()
crypten.print(f"\nRank {rank}:\n {x_enc}\n", in_order=True)
x = examine_binary_shares()
```
### Using Both Secret-sharing Protocols
Quite often a mathematical function may need to use both additive and XOR secret sharing for efficient evaluation. Functions that require conversions between sharing types include comparators (`>, >=, <, <=, ==, !=`) as well as functions derived from them (`abs, sign, relu`, etc.). For a full list of supported functions, please see the CrypTen documentation.
CrypTen provides functionality that allows for the conversion of between <i>ptypes</i>. Conversion between <i>ptypes</i> can be done using the `.to()` function with a `crypten.ptype` input, or by calling the `.arithmetic()` and `.binary()` conversion functions.
```
from crypten.mpc import MPCTensor
@mpc.run_multiprocess(world_size=2)
def examine_conversion():
x = torch.tensor([1, 2, 3])
rank = comm.get().get_rank()
# create an MPCTensor with arithmetic secret sharing
x_enc_arithmetic = MPCTensor(x, ptype=crypten.mpc.arithmetic)
# To binary
x_enc_binary = x_enc_arithmetic.to(crypten.mpc.binary)
x_from_binary = x_enc_binary.get_plain_text()
# print only once
crypten.print("to(crypten.binary):")
crypten.print(f" ptype: {x_enc_binary.ptype}\n plaintext: {x_from_binary}\n")
# To arithmetic
x_enc_arithmetic = x_enc_arithmetic.to(crypten.mpc.arithmetic)
x_from_arithmetic = x_enc_arithmetic.get_plain_text()
# print only once
crypten.print("to(crypten.arithmetic):")
crypten.print(f" ptype: {x_enc_arithmetic.ptype}\n plaintext: {x_from_arithmetic}\n")
z = examine_conversion()
```
## Data Sources
CrypTen follows the standard MPI programming model: it runs a separate process for each party, but each process runs an identical (complete) program. Each process has a `rank` variable to identify itself.
If the process with rank `i` is the source of data `x`, then `x` gets encrypted with `i` as its source value (denoted as `src`). However, MPI protocols require that both processes to provide a tensor with the same size as their input. CrypTen ignores all data provided from non-source processes when encrypting.
In the next example, we'll show how to use the `rank` and `src` values to encrypt tensors. Here, we will have each of 3 parties generate a value `x` which is equal to its own `rank` value. Within the loop, 3 encrypted tensors are created, each with a different source. When these tensors are decrypted, we can verify that the tensors are generated using the tensor provided by the source process.
(Note that `crypten.cryptensor` uses rank 0 as the default source if none is provided.)
```
@mpc.run_multiprocess(world_size=3)
def examine_sources():
# Create a different tensor on each rank
rank = comm.get().get_rank()
x = torch.tensor(rank)
crypten.print(f"Rank {rank}: {x}", in_order=True)
#
world_size = comm.get().get_world_size()
for i in range(world_size):
x_enc = crypten.cryptensor(x, src=i)
z = x_enc.get_plain_text()
# Only print from one process to avoid duplicates
crypten.print(f"Source {i}: {z}")
x = examine_sources()
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import nltk
from collections import Counter
from sklearn.metrics import log_loss
from scipy.optimize import minimize
import multiprocessing
import difflib
import time
import gc
import xgboost as xgb
from sklearn.cross_validation import train_test_split
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer
from scipy.spatial.distance import cosine, correlation, canberra, chebyshev, minkowski, jaccard, euclidean
from xgb_utils import *
def get_train():
feats_src = '/media/w/1c392724-ecf3-4615-8f3c-79368ec36380/DS Projects/Kaggle/Quora/data/features/uncleaned_data/'
keras_q1 = np.load(feats_src + 'train_q1_transformed.npy')
keras_q2 = np.load(feats_src + 'train_q2_transformed.npy')
xgb_feats = pd.read_csv(feats_src + '/the_1owl/owl_train.csv')
abhishek_feats = pd.read_csv(feats_src + 'abhishek/train_features.csv',
encoding = 'ISO-8859-1').iloc[:, 2:]
text_feats = pd.read_csv(feats_src + 'other_features/text_features_train.csv',
encoding = 'ISO-8859-1')
img_feats = pd.read_csv(feats_src + 'other_features/img_features_train.csv')
srk_feats = pd.read_csv(feats_src + 'srk/SRK_grams_features_train.csv')
mephisto_feats = pd.read_csv('../../data/features/spacylemmat_fullclean/train_mephistopeheles_features.csv').iloc[:, 6:]
turkewitz_feats = pd.read_csv('../../data/features/lemmat_spacy_features/train_turkewitz_features.csv')
turkewitz_feats = turkewitz_feats[['q1_freq', 'q2_freq']]
turkewitz_feats['freq_sum'] = turkewitz_feats.q1_freq + turkewitz_feats.q2_freq
turkewitz_feats['freq_diff'] = turkewitz_feats.q1_freq - turkewitz_feats.q2_freq
xgb_feats.drop(['z_len1', 'z_len2', 'z_word_len1', 'z_word_len2'], axis = 1, inplace = True)
y_train = xgb_feats['is_duplicate']
xgb_feats = xgb_feats.iloc[:, 8:]
df = pd.concat([xgb_feats, abhishek_feats, text_feats, img_feats,
turkewitz_feats, mephisto_feats], axis = 1)
df = pd.DataFrame(df)
dfc = df.iloc[0:1000,:]
dfc = dfc.T.drop_duplicates().T
duplicate_cols = sorted(list(set(df.columns).difference(set(dfc.columns))))
print('Dropping duplicate columns:', duplicate_cols)
df.drop(duplicate_cols, axis = 1, inplace = True)
print('Final shape:', df.shape)
X = np.concatenate([keras_q1, keras_q2, df.values], axis = 1)
X = X.astype('float32')
print('Training data shape:', X.shape)
return X, y_train
def get_test():
feats_src = '/media/w/1c392724-ecf3-4615-8f3c-79368ec36380/DS Projects/Kaggle/Quora/data/features/uncleaned_data/'
keras_q1 = np.load(feats_src + 'test_q1_transformed.npy')
keras_q2 = np.load(feats_src + 'test_q2_transformed.npy')
xgb_feats = pd.read_csv(feats_src + '/the_1owl/owl_test.csv')
abhishek_feats = pd.read_csv(feats_src + 'abhishek/test_features.csv',
encoding = 'ISO-8859-1').iloc[:, 2:]
text_feats = pd.read_csv(feats_src + 'other_features/text_features_test.csv',
encoding = 'ISO-8859-1')
img_feats = pd.read_csv(feats_src + 'other_features/img_features_test.csv')
srk_feats = pd.read_csv(feats_src + 'srk/SRK_grams_features_test.csv')
mephisto_feats = pd.read_csv('../../data/features/spacylemmat_fullclean/test_mephistopeheles_features.csv').iloc[:, 6:]
turkewitz_feats = pd.read_csv('../../data/features/lemmat_spacy_features/test_turkewitz_features.csv')
turkewitz_feats = turkewitz_feats[['q1_freq', 'q2_freq']]
turkewitz_feats['freq_sum'] = turkewitz_feats.q1_freq + turkewitz_feats.q2_freq
turkewitz_feats['freq_diff'] = turkewitz_feats.q1_freq - turkewitz_feats.q2_freq
xgb_feats.drop(['z_len1', 'z_len2', 'z_word_len1', 'z_word_len2'], axis = 1, inplace = True)
xgb_feats = xgb_feats.iloc[:, 5:]
df = pd.concat([xgb_feats, abhishek_feats, text_feats, img_feats,
turkewitz_feats, mephisto_feats], axis = 1)
df = pd.DataFrame(df)
dfc = df.iloc[0:1000,:]
dfc = dfc.T.drop_duplicates().T
duplicate_cols = sorted(list(set(df.columns).difference(set(dfc.columns))))
print('Dropping duplicate columns:', duplicate_cols)
df.drop(duplicate_cols, axis = 1, inplace = True)
print('Final shape:', df.shape)
X = np.concatenate([keras_q1, keras_q2, df.values], axis = 1)
X = X.astype('float32')
print('Training data shape:', X.shape)
return X
def predict_test(X_test, model_name):
print('Predicting on test set.')
gbm = xgb.Booster(model_file = 'saved_models/XGB/{}.txt'.format(model_name))
test_preds = gbm.predict(xgb.DMatrix(X_test))
sub_src = '/media/w/1c392724-ecf3-4615-8f3c-79368ec36380/DS Projects/Kaggle/Quora/submissions/'
sample_sub = pd.read_csv(sub_src + 'sample_submission.csv')
sample_sub['is_duplicate'] = test_preds
sample_sub.is_duplicate = sample_sub.is_duplicate.apply(transform)
sample_sub.to_csv(sub_src + '{}.csv'.format(model_name), index = False)
return
def train_xgb(cv = False):
t = time.time()
params = {
'seed': 1337,
'colsample_bytree': 0.48,
'silent': 1,
'subsample': 0.74,
'eta': 0.05,
'objective': 'binary:logistic',
'eval_metric': 'logloss',
'max_depth': 12,
'min_child_weight': 20,
'nthread': 8,
'tree_method': 'hist',
#'updater': 'grow_gpu',
}
X_train, y_train = get_train()
if cv:
dtrain = xgb.DMatrix(X_train, y_train)
hist = xgb.cv(params, dtrain, num_boost_round = 100000, nfold = 5,
stratified = True, early_stopping_rounds = 350, verbose_eval = 250,
seed = 1337)
del X_train, y_train
gc.collect()
print('Time it took to train in CV manner:', time.time() - t)
return hist
else:
X_tr, X_val, y_tr, y_val = train_test_split(X_train, y_train, stratify = y_train,
test_size = 0.2, random_state = 111)
del X_train, y_train
gc.collect()
dtrain = xgb.DMatrix(X_tr, label = y_tr)
dval = xgb.DMatrix(X_val, label = y_val)
watchlist = [(dtrain, 'train'), (dval, 'valid')]
print('Start training...')
gbm = xgb.train(params, dtrain, 100000, watchlist,
early_stopping_rounds = 350, verbose_eval = 100)
print('Start predicting...')
val_pred = gbm.predict(xgb.DMatrix(X_val), ntree_limit=gbm.best_ntree_limit)
score = log_loss(y_val, val_pred)
print('Final score:', score, '\n', 'Time it took to train and predict:', time.time() - t)
del X_tr, X_val, y_tr, y_val
gc.collect()
return gbm
def run_xgb(model_name, train = True, test = False, cv = False):
if cv:
gbm_hist = train_xgb(True)
return gbm_hist
if train:
gbm = train_xgb()
gbm.save_model('saved_models/XGB/{}.txt'.format(model_name))
if test:
predict_test('{}'.format(model_name))
return gbm
gbm = run_xgb('XGB_firstBO_turkewitz_experiments', train = True, test = False)
predict_test('XGB_firstBO_turkewitz_Qspacyencode')
def get_transformations_features(transformations_src, mode = 'train'):
lsa10tr_3grams_q1 = np.load(transformations_src + '{}_lsa10_3grams.npy'.format(mode))[0]
lsa10tr_3grams_q2 = np.load(transformations_src + '{}_lsa10_3grams.npy'.format(mode))[1]
transforms_feats = pd.DataFrame()
transforms_feats['cosine'] = [cosine(x, y) for (x,y) in zip(lsa10tr_3grams_q1, lsa10tr_3grams_q2)]
transforms_feats['correlation'] = [correlation(x, y) for (x,y) in zip(lsa10tr_3grams_q1, lsa10tr_3grams_q2)]
transforms_feats['jaccard'] = [jaccard(x, y) for (x,y) in zip(lsa10tr_3grams_q1, lsa10tr_3grams_q2)]
transforms_feats['euclidean'] = [euclidean(x, y) for (x,y) in zip(lsa10tr_3grams_q1, lsa10tr_3grams_q2)]
transforms_feats['minkowski'] = [minkowski(x, y, 3) for (x,y) in zip(lsa10tr_3grams_q1, lsa10tr_3grams_q2)]
return transforms_feats
def get_doc2vec_features(doc2vec_src, mode = 'train'):
doc2vec_pre_q1 = np.load(doc2vec_src + '{}_q1_doc2vec_vectors_pretrained.npy'.format(mode))
doc2vec_pre_q2 = np.load(doc2vec_src + '{}_q2_doc2vec_vectors_pretrained.npy'.format(mode))
doc2vec_quora_q1 = np.load(doc2vec_src + '{}_q1_doc2vec_vectors_trainquora.npy'.format(mode))
doc2vec_quora_q2 = np.load(doc2vec_src + '{}_q2_doc2vec_vectors_trainquora.npy'.format(mode))
d2v_feats_pretrained = pd.DataFrame()
d2v_feats_pretrained['cosine'] = [cosine(x, y) for (x,y) in zip(doc2vec_pre_q1, doc2vec_pre_q2)]
d2v_feats_pretrained['correlation'] = [correlation(x, y) for (x,y) in zip(doc2vec_pre_q1, doc2vec_pre_q2)]
d2v_feats_pretrained['jaccard'] = [jaccard(x, y) for (x,y) in zip(doc2vec_pre_q1, doc2vec_pre_q2)]
d2v_feats_pretrained['euclidean'] = [euclidean(x, y) for (x,y) in zip(doc2vec_pre_q1, doc2vec_pre_q2)]
d2v_feats_pretrained['minkowski'] = [minkowski(x, y, 3) for (x,y) in zip(doc2vec_pre_q1, doc2vec_pre_q2)]
d2v_feats_quora = pd.DataFrame()
d2v_feats_quora['cosine'] = [cosine(x, y) for (x,y) in zip(doc2vec_quora_q1, doc2vec_quora_q2)]
d2v_feats_quora['correlation'] = [correlation(x, y) for (x,y) in zip(doc2vec_quora_q1, doc2vec_quora_q2)]
d2v_feats_quora['jaccard'] = [jaccard(x, y) for (x,y) in zip(doc2vec_quora_q1, doc2vec_quora_q2)]
d2v_feats_quora['euclidean'] = [euclidean(x, y) for (x,y) in zip(doc2vec_quora_q1, doc2vec_quora_q2)]
d2v_feats_quora['minkowski'] = [minkowski(x, y, 3) for (x,y) in zip(doc2vec_quora_q1, doc2vec_quora_q2)]
return d2v_feats_pretrained, d2v_feats_quora
doc2vec_src = '/media/w/1c392724-ecf3-4615-8f3c-79368ec36380/DS Projects/Kaggle/Quora/data/features/lemmat_spacy_features/doc2vec/'
transformations_src = '/media/w/1c392724-ecf3-4615-8f3c-79368ec36380/DS Projects/Kaggle/Quora/data/features/uncleaned_data/transformations/'
X_train, y_train = get_train()
d2v_pre, d2v_quora = get_doc2vec_features(doc2vec_src, mode = 'train')
transforms = get_transformations_features(transformations_src)
X_train = np.concatenate([X_train, d2v_pre, d2v_quora, transforms], axis = 1)
params = {
'seed': 1337,
'colsample_bytree': 0.48,
'silent': 1,
'subsample': 0.74,
'eta': 0.05,
'objective': 'binary:logistic',
'eval_metric': 'logloss',
'max_depth': 12,
'min_child_weight': 20,
'nthread': 8,
'tree_method': 'hist',
}
t = time.time()
X_tr, X_val, y_tr, y_val = train_test_split(X_train, y_train, stratify = y_train,
test_size = 0.2, random_state = 111)
dtrain = xgb.DMatrix(X_tr, label = y_tr)
dval = xgb.DMatrix(X_val, label = y_val)
watchlist = [(dtrain, 'train'), (dval, 'valid')]
print('Start training...')
gbm = xgb.train(params, dtrain, 100000, watchlist,
early_stopping_rounds = 150, verbose_eval = 100)
print('Start predicting...')
val_pred = gbm.predict(xgb.DMatrix(X_val), ntree_limit=gbm.best_ntree_limit)
score = log_loss(y_val, val_pred)
print('Final score:', score, '\n', 'Time it took to train and predict:', time.time() - t)
gbm.save_model('saved_models/XGB/XGB_firstBO_turkewitz_Doc2Vec_LSA.txt')
def get_transformations_features(transformations_src):
lsa10tr_3grams_q1 = np.load(transformations_src + 'lsa10te_3grams.npy')[0]
lsa10tr_3grams_q2 = np.load(transformations_src + 'lsa10te_3grams.npy')[1]
transforms_feats = pd.DataFrame()
transforms_feats['cosine'] = [cosine(x, y) for (x,y) in zip(lsa10tr_3grams_q1, lsa10tr_3grams_q2)]
transforms_feats['correlation'] = [correlation(x, y) for (x,y) in zip(lsa10tr_3grams_q1, lsa10tr_3grams_q2)]
transforms_feats['jaccard'] = [jaccard(x, y) for (x,y) in zip(lsa10tr_3grams_q1, lsa10tr_3grams_q2)]
transforms_feats['euclidean'] = [euclidean(x, y) for (x,y) in zip(lsa10tr_3grams_q1, lsa10tr_3grams_q2)]
transforms_feats['minkowski'] = [minkowski(x, y, 3) for (x,y) in zip(lsa10tr_3grams_q1, lsa10tr_3grams_q2)]
return transforms_feats
def get_doc2vec_features(doc2vec_src):
doc2vec_pre_q1 = np.load(doc2vec_src + 'test_q1_doc2vec_vectors_pretrained.npy')
doc2vec_pre_q2 = np.load(doc2vec_src + 'test_q2_doc2vec_vectors_pretrained.npy')
d2v_feats_pretrained = pd.DataFrame()
d2v_feats_pretrained['cosine'] = [cosine(x, y) for (x,y) in zip(doc2vec_pre_q1, doc2vec_pre_q2)]
d2v_feats_pretrained['correlation'] = [correlation(x, y) for (x,y) in zip(doc2vec_pre_q1, doc2vec_pre_q2)]
d2v_feats_pretrained['jaccard'] = [jaccard(x, y) for (x,y) in zip(doc2vec_pre_q1, doc2vec_pre_q2)]
d2v_feats_pretrained['euclidean'] = [euclidean(x, y) for (x,y) in zip(doc2vec_pre_q1, doc2vec_pre_q2)]
d2v_feats_pretrained['minkowski'] = [minkowski(x, y, 3) for (x,y) in zip(doc2vec_pre_q1, doc2vec_pre_q2)]
return d2v_feats_pretrained
doc2vec_src = '/media/w/1c392724-ecf3-4615-8f3c-79368ec36380/DS Projects/Kaggle/Quora/data/features/lemmat_spacy_features/doc2vec/'
transformations_src = '/media/w/1c392724-ecf3-4615-8f3c-79368ec36380/DS Projects/Kaggle/Quora/data/features/uncleaned_data/transformations/'
X_test = get_test()
d2v_feats = get_doc2vec_features(doc2vec_src)
transforms_feats = get_transformations_features(transformations_src)
X_test = np.concatenate([X_test, d2v_feats, transforms_feats], axis = 1)
predict_test(X_test, 'XGB_firstBO_turkewitz_Doc2Vec_LSA')
```
| github_jupyter |
## Libraries
```
import pandas as pd
import numpy as np
import scipy.stats as stat
from math import sqrt
from mlgear.utils import show, display_columns
from surveyweights import normalize_weights
def margin_of_error(n=None, sd=None, p=None, type='proportion', interval_size=0.95):
z_lookup = {0.8: 1.28, 0.85: 1.44, 0.9: 1.65, 0.95: 1.96, 0.99: 2.58}
if interval_size not in z_lookup.keys():
raise ValueError('{} not a valid `interval_size` - must be {}'.format(interval_size,
', '.join(list(z_lookup.keys()))))
if type == 'proportion':
se = sqrt(p * (1 - p)) / sqrt(n)
elif type == 'continuous':
se = sd / sqrt(n)
else:
raise ValueError('{} not a valid `type` - must be proportion or continuous')
z = z_lookup[interval_size]
return se * z
def print_pct(pct, digits=0):
pct = pct * 100
pct = np.round(pct, digits)
if pct >= 100:
if digits == 0:
val = '>99.0%'
else:
val = '>99.'
for d in range(digits - 1):
val += '9'
val += '9%'
elif pct <= 0:
if digits == 0:
val = '<0.1%'
else:
val = '<0.'
for d in range(digits - 1):
val += '0'
val += '1%'
else:
val = '{}%'.format(pct)
return val
def calc_result(biden_vote, trump_vote, n, interval=0.8):
GENERAL_POLLING_ERROR = 2.5
N_SIMS = 100000
biden_moe = margin_of_error(n=n, p=biden_vote/100, interval_size=interval)
trump_moe = margin_of_error(n=n, p=trump_vote/100, interval_size=interval)
undecided = (100 - biden_vote - trump_vote) / 2
biden_mean = biden_vote + undecided * 0.25
biden_raw_moe = biden_moe * 100
biden_allocate_undecided = undecided * 0.4
biden_margin = biden_raw_moe + biden_allocate_undecided + GENERAL_POLLING_ERROR
trump_mean = trump_vote + undecided * 0.25
trump_raw_moe = trump_moe * 100
trump_allocate_undecided = undecided * 0.4
trump_margin = trump_raw_moe + trump_allocate_undecided + GENERAL_POLLING_ERROR
cdf_value = 0.5 + 0.5 * interval
normed_sigma = stat.norm.ppf(cdf_value)
biden_sigma = biden_margin / 100 / normed_sigma
biden_sims = np.random.normal(biden_mean / 100, biden_sigma, N_SIMS)
trump_sigma = trump_margin / 100 / normed_sigma
trump_sims = np.random.normal(trump_mean / 100, trump_sigma, N_SIMS)
chance_pass = np.sum([sim[0] > sim[1] for sim in zip(biden_sims, trump_sims)]) / N_SIMS
low, high = np.percentile(biden_sims - trump_sims, [20, 80]) * 100
return {'mean': biden_mean - trump_mean, 'high': high, 'low': low, 'n': n,
'raw_moe': biden_raw_moe + trump_raw_moe,
'margin': (biden_margin + trump_margin) / 2,
'sigma': (biden_sigma + trump_sigma) / 2,
'chance_pass': chance_pass}
def print_result(mean, high, low, n, raw_moe, margin, sigma, chance_pass):
mean = np.round(mean, 1)
first = np.round(high, 1)
second = np.round(low, 1)
sigma = np.round(sigma * 100, 1)
raw_moe = np.round(raw_moe, 1)
margin = np.round(margin, 1)
chance_pass = print_pct(chance_pass, 1)
if second < first:
_ = first
first = second
second = _
if second > 100:
second = 100
if first < -100:
first = -100
print(('Result Biden {} (80% CI: {} to {}) (Weighted N={}) (raw_moe={}pts, margin={}pts, '
'sigma={}pts) (Biden {} likely to win)').format(mean,
first,
second,
n,
raw_moe,
margin,
sigma,
chance_pass))
print('-')
def calc_result_sen(dem_vote, rep_vote, n, interval=0.8):
GENERAL_POLLING_ERROR = 2.5
N_SIMS = 100000
dem_moe = margin_of_error(n=n, p=dem_vote/100, interval_size=interval)
rep_moe = margin_of_error(n=n, p=rep_vote/100, interval_size=interval)
undecided = 100 - dem_vote - rep_vote
dem_mean = dem_vote + undecided * 0.25
dem_raw_moe = dem_moe * 100
dem_allocate_undecided = undecided * 0.4
dem_margin = dem_raw_moe + dem_allocate_undecided + GENERAL_POLLING_ERROR
rep_mean = rep_vote + undecided * 0.25
rep_raw_moe = rep_moe * 100
rep_allocate_undecided = undecided * 0.4
rep_margin = rep_raw_moe + rep_allocate_undecided + GENERAL_POLLING_ERROR
cdf_value = 0.5 + 0.5 * interval
normed_sigma = stat.norm.ppf(cdf_value)
dem_sigma = dem_margin / 100 / normed_sigma
dem_sims = np.random.normal(dem_mean / 100, dem_sigma, N_SIMS)
rep_sigma = rep_margin / 100 / normed_sigma
rep_sims = np.random.normal(rep_mean / 100, rep_sigma, N_SIMS)
chance_pass = np.sum([sim[0] > sim[1] for sim in zip(dem_sims, rep_sims)]) / N_SIMS
low, high = np.percentile(dem_sims - rep_sims, [20, 80]) * 100
return {'mean': dem_mean - rep_mean, 'high': high, 'low': low, 'n': n,
'raw_moe': dem_raw_moe + rep_raw_moe,
'margin': (dem_margin + rep_margin) / 2,
'sigma': (dem_sigma + rep_sigma) / 2,
'chance_pass': chance_pass}
def print_result_sen(mean, high, low, n, raw_moe, margin, sigma, chance_pass):
mean = np.round(mean, 1)
first = np.round(high, 1)
second = np.round(low, 1)
sigma = np.round(sigma * 100, 1)
raw_moe = np.round(raw_moe, 1)
margin = np.round(margin, 1)
chance_pass = print_pct(chance_pass, 1)
if second < first:
_ = first
first = second
second = _
if second > 100:
second = 100
if first < -100:
first = -100
print(('Result Hegar (D) {} (80% CI: {} to {}) (Weighted N={}) (raw_moe={}pts, margin={}pts, '
'sigma={}pts) (Hegar {} likely to win)').format(mean,
first,
second,
n,
raw_moe,
margin,
sigma,
chance_pass))
print('-')
```
## Load Processed Data
```
tx_national_survey = pd.read_csv('responses_processed_tx_weighted.csv').fillna('Not presented')
tx_state_survey = pd.read_csv('responses_processed_tx_state_tx_weighted.csv').fillna('Not presented')
```
## Texas Trump-Clinton
```
options = ['Donald Trump', 'Hillary Clinton', 'Other']
survey_ = tx_national_survey.loc[tx_national_survey['vote2016'].isin(options)].copy()
survey_['lv_weight'] = normalize_weights(survey_['lv_weight'])
survey_['vote2016'].value_counts(normalize=True) * survey_.groupby('vote2016')['lv_weight'].mean() * 100
options = ['Donald Trump', 'Hillary Clinton', 'Other']
survey_ = tx_state_survey.loc[tx_state_survey['vote2016'].isin(options)].copy()
survey_['lv_weight'] = normalize_weights(survey_['lv_weight'])
survey_['vote2016'].value_counts(normalize=True) * survey_.groupby('vote2016')['lv_weight'].mean() * 100
```
## Texas Trump-Biden
```
options = ['Joe Biden, the Democrat', 'Donald Trump, the Republican', 'Another candidate', 'Not decided']
survey_ = tx_national_survey.loc[tx_national_survey['vote_trump_biden'].isin(options)].copy()
survey_['weight'] = normalize_weights(survey_['weight'])
survey_['rv_weight'] = normalize_weights(survey_['rv_weight'])
survey_['lv_weight'] = normalize_weights(survey_['lv_weight'])
survey_['lv_weight_alt'] = normalize_weights(survey_['lv_weight_alt'])
survey_['lv_weight_2020'] = normalize_weights(survey_['lv_weight_2020'])
print('## NATIONAL TX-WEIGHTED ##')
weighted_n = int(np.round(survey_['weight'].apply(lambda w: 1 if w > 1 else w).sum()))
votes = survey_['vote_trump_biden'].value_counts(normalize=True) * survey_.groupby('vote_trump_biden')['weight'].mean() * 100
votes = votes[options] * (100 / votes[options].sum())
print(votes)
print_result(**calc_result(biden_vote=votes['Joe Biden, the Democrat'],
trump_vote=votes['Donald Trump, the Republican'],
n=weighted_n))
print('## NATIONAL TX-WEIGHTED + RV ##')
rv_weighted_n = int(np.round(survey_['rv_weight'].apply(lambda w: 1 if w > 1 else w).sum()))
votes = survey_['vote_trump_biden'].value_counts(normalize=True) * survey_.groupby('vote_trump_biden')['rv_weight'].mean() * 100
votes = votes[options] * (100 / votes[options].sum())
print(votes)
print_result(**calc_result(biden_vote=votes['Joe Biden, the Democrat'],
trump_vote=votes['Donald Trump, the Republican'],
n=rv_weighted_n))
print('## NATIONAL TX-WEIGHTED + LV ##')
lv_weighted_n = int(np.round(survey_['lv_weight'].apply(lambda w: 1 if w > 1 else w).sum()))
votes = survey_['vote_trump_biden'].value_counts(normalize=True) * survey_.groupby('vote_trump_biden')['lv_weight'].mean() * 100
votes = votes[options] * (100 / votes[options].sum())
print(votes)
print_result(**calc_result(biden_vote=votes['Joe Biden, the Democrat'],
trump_vote=votes['Donald Trump, the Republican'],
n=lv_weighted_n))
print('## NATIONAL TX-WEIGHTED + LV ALT (POST-HOC) ##')
lv_weighted_n = int(np.round(survey_['lv_weight_alt'].apply(lambda w: 1 if w > 1 else w).sum()))
votes = survey_['vote_trump_biden'].value_counts(normalize=True) * survey_.groupby('vote_trump_biden')['lv_weight_alt'].mean() * 100
votes = votes[options] * (100 / votes[options].sum())
print(votes)
print_result(**calc_result(biden_vote=votes['Joe Biden, the Democrat'],
trump_vote=votes['Donald Trump, the Republican'],
n=lv_weighted_n))
print('## NATIONAL TX-WEIGHTED + LV 2020 (2020 NATIONAL VOTE MATCH) ##')
lv_weighted_n = int(np.round(survey_['lv_weight_2020'].apply(lambda w: 1 if w > 1 else w).sum()))
votes = survey_['vote_trump_biden'].value_counts(normalize=True) * survey_.groupby('vote_trump_biden')['lv_weight_2020'].mean() * 100
votes = votes[options] * (100 / votes[options].sum())
print(votes)
print_result(**calc_result(biden_vote=votes['Joe Biden, the Democrat'],
trump_vote=votes['Donald Trump, the Republican'],
n=lv_weighted_n))
print('## TEXAS TX-WEIGHTED ##')
survey_ = tx_state_survey.loc[tx_state_survey['vote_trump_biden'].isin(options)].copy()
survey_['weight'] = normalize_weights(survey_['weight'])
survey_['rv_weight'] = normalize_weights(survey_['rv_weight'])
survey_['lv_weight'] = normalize_weights(survey_['lv_weight'])
survey_['lv_weight_alt'] = normalize_weights(survey_['lv_weight_alt'])
survey_['lv_weight_2020'] = normalize_weights(survey_['lv_weight_2020'])
weighted_n = int(np.round(survey_['weight'].apply(lambda w: 1 if w > 1 else w).sum()))
votes = survey_['vote_trump_biden'].value_counts(normalize=True) * survey_.groupby('vote_trump_biden')['weight'].mean() * 100
votes = votes[options] * (100 / votes[options].sum())
print(votes)
print_result(**calc_result(biden_vote=votes['Joe Biden, the Democrat'],
trump_vote=votes['Donald Trump, the Republican'],
n=weighted_n))
print('## TEXAS TX-WEIGHTED + RV ##')
rv_weighted_n = int(np.round(survey_['rv_weight'].apply(lambda w: 1 if w > 1 else w).sum()))
votes = survey_['vote_trump_biden'].value_counts(normalize=True) * survey_.groupby('vote_trump_biden')['rv_weight'].mean() * 100
votes = votes[options] * (100 / votes[options].sum())
print(votes)
print_result(**calc_result(biden_vote=votes['Joe Biden, the Democrat'],
trump_vote=votes['Donald Trump, the Republican'],
n=rv_weighted_n))
print('## TEXAS TX-WEIGHTED + LV ##')
lv_weighted_n = int(np.round(survey_['lv_weight'].apply(lambda w: 1 if w > 1 else w).sum()))
votes = survey_['vote_trump_biden'].value_counts(normalize=True) * survey_.groupby('vote_trump_biden')['lv_weight'].mean() * 100
votes = votes[options] * (100 / votes[options].sum())
print(votes)
print_result(**calc_result(biden_vote=votes['Joe Biden, the Democrat'],
trump_vote=votes['Donald Trump, the Republican'],
n=lv_weighted_n))
print('## TEXAS TX-WEIGHTED + LV ALT (POST-HOC) ##')
lv_weighted_n = int(np.round(survey_['lv_weight_alt'].apply(lambda w: 1 if w > 1 else w).sum()))
votes = survey_['vote_trump_biden'].value_counts(normalize=True) * survey_.groupby('vote_trump_biden')['lv_weight_alt'].mean() * 100
votes = votes[options] * (100 / votes[options].sum())
print(votes)
print_result(**calc_result(biden_vote=votes['Joe Biden, the Democrat'],
trump_vote=votes['Donald Trump, the Republican'],
n=lv_weighted_n))
print('## TEXAS TX-WEIGHTED + LV 2020 (2020 NATIONAL VOTE MATCH) ##')
lv_weighted_n = int(np.round(survey_['lv_weight_2020'].apply(lambda w: 1 if w > 1 else w).sum()))
votes = survey_['vote_trump_biden'].value_counts(normalize=True) * survey_.groupby('vote_trump_biden')['lv_weight_2020'].mean() * 100
votes = votes[options] * (100 / votes[options].sum())
print(votes)
print_result(**calc_result(biden_vote=votes['Joe Biden, the Democrat'],
trump_vote=votes['Donald Trump, the Republican'],
n=lv_weighted_n))
options = ['A Democratic candidate', 'A Republican candidate', 'Another candidate', 'Not decided']
survey_ = tx_national_survey.loc[tx_national_survey['vote_senate'].isin(options)].copy()
survey_['weight'] = normalize_weights(survey_['weight'])
survey_['rv_weight'] = normalize_weights(survey_['rv_weight'])
survey_['lv_weight'] = normalize_weights(survey_['lv_weight'])
survey_['lv_weight_alt'] = normalize_weights(survey_['lv_weight_alt'])
survey_['lv_weight_2020'] = normalize_weights(survey_['lv_weight_2020'])
print('## NATIONAL TX-WEIGHTED ##')
weighted_n = int(np.round(survey_['weight'].apply(lambda w: 1 if w > 1 else w).sum()))
votes = survey_['vote_senate'].value_counts(normalize=True) * survey_.groupby('vote_senate')['weight'].mean() * 100
votes = votes[options] * (100 / votes[options].sum())
print(votes)
print_result_sen(**calc_result_sen(dem_vote=votes['A Democratic candidate'],
rep_vote=votes['A Republican candidate'],
n=weighted_n))
print('## NATIONAL TX-WEIGHTED + RV ##')
rv_weighted_n = int(np.round(survey_['rv_weight'].apply(lambda w: 1 if w > 1 else w).sum()))
votes = survey_['vote_senate'].value_counts(normalize=True) * survey_.groupby('vote_senate')['rv_weight'].mean() * 100
votes = votes[options] * (100 / votes[options].sum())
print(votes)
print_result_sen(**calc_result_sen(dem_vote=votes['A Democratic candidate'],
rep_vote=votes['A Republican candidate'],
n=rv_weighted_n))
print('## NATIONAL TX-WEIGHTED + LV ##')
lv_weighted_n = int(np.round(survey_['lv_weight'].apply(lambda w: 1 if w > 1 else w).sum()))
votes = survey_['vote_senate'].value_counts(normalize=True) * survey_.groupby('vote_senate')['lv_weight'].mean() * 100
votes = votes[options] * (100 / votes[options].sum())
print(votes)
print_result_sen(**calc_result_sen(dem_vote=votes['A Democratic candidate'],
rep_vote=votes['A Republican candidate'],
n=lv_weighted_n))
print('## NATIONAL TX-WEIGHTED + LV ALT (POST-HOC) ##')
lv_weighted_n = int(np.round(survey_['lv_weight_alt'].apply(lambda w: 1 if w > 1 else w).sum()))
votes = survey_['vote_senate'].value_counts(normalize=True) * survey_.groupby('vote_senate')['lv_weight_alt'].mean() * 100
votes = votes[options] * (100 / votes[options].sum())
print(votes)
print_result_sen(**calc_result_sen(dem_vote=votes['A Democratic candidate'],
rep_vote=votes['A Republican candidate'],
n=lv_weighted_n))
print('## NATIONAL TX-WEIGHTED + LV 2020 (2020 NATIONAL VOTE MATCH) ##')
lv_weighted_n = int(np.round(survey_['lv_weight_2020'].apply(lambda w: 1 if w > 1 else w).sum()))
votes = survey_['vote_senate'].value_counts(normalize=True) * survey_.groupby('vote_senate')['lv_weight_2020'].mean() * 100
votes = votes[options] * (100 / votes[options].sum())
print(votes)
print_result_sen(**calc_result_sen(dem_vote=votes['A Democratic candidate'],
rep_vote=votes['A Republican candidate'],
n=lv_weighted_n))
print('## TEXAS TX-WEIGHTED ##')
survey_ = tx_state_survey.loc[tx_state_survey['vote_senate'].isin(options)].copy()
survey_['weight'] = normalize_weights(survey_['weight'])
survey_['rv_weight'] = normalize_weights(survey_['rv_weight'])
survey_['lv_weight'] = normalize_weights(survey_['lv_weight'])
survey_['lv_weight_alt'] = normalize_weights(survey_['lv_weight_alt'])
survey_['lv_weight_2020'] = normalize_weights(survey_['lv_weight_2020'])
weighted_n = int(np.round(survey_['weight'].apply(lambda w: 1 if w > 1 else w).sum()))
votes = survey_['vote_senate'].value_counts(normalize=True) * survey_.groupby('vote_senate')['weight'].mean() * 100
votes = votes[options] * (100 / votes[options].sum())
print(votes)
print_result_sen(**calc_result_sen(dem_vote=votes['A Democratic candidate'],
rep_vote=votes['A Republican candidate'],
n=weighted_n))
print('## TEXAS TX-WEIGHTED + RV ##')
rv_weighted_n = int(np.round(survey_['rv_weight'].apply(lambda w: 1 if w > 1 else w).sum()))
votes = survey_['vote_senate'].value_counts(normalize=True) * survey_.groupby('vote_senate')['rv_weight'].mean() * 100
votes = votes[options] * (100 / votes[options].sum())
print(votes)
print_result_sen(**calc_result_sen(dem_vote=votes['A Democratic candidate'],
rep_vote=votes['A Republican candidate'],
n=rv_weighted_n))
print('## TEXAS TX-WEIGHTED + LV ##')
lv_weighted_n = int(np.round(survey_['lv_weight'].apply(lambda w: 1 if w > 1 else w).sum()))
votes = survey_['vote_senate'].value_counts(normalize=True) * survey_.groupby('vote_senate')['lv_weight'].mean() * 100
votes = votes[options] * (100 / votes[options].sum())
print(votes)
print_result_sen(**calc_result_sen(dem_vote=votes['A Democratic candidate'],
rep_vote=votes['A Republican candidate'],
n=lv_weighted_n))
print('## TEXAS TX-WEIGHTED + LV ALT (POST-HOC) ##')
lv_weighted_n = int(np.round(survey_['lv_weight_alt'].apply(lambda w: 1 if w > 1 else w).sum()))
votes = survey_['vote_senate'].value_counts(normalize=True) * survey_.groupby('vote_senate')['lv_weight_alt'].mean() * 100
votes = votes[options] * (100 / votes[options].sum())
print(votes)
print_result_sen(**calc_result_sen(dem_vote=votes['A Democratic candidate'],
rep_vote=votes['A Republican candidate'],
n=lv_weighted_n))
print('## TEXAS TX-WEIGHTED + LV 2020 (2020 NATIONAL VOTE MATCH) ##')
lv_weighted_n = int(np.round(survey_['lv_weight_2020'].apply(lambda w: 1 if w > 1 else w).sum()))
votes = survey_['vote_senate'].value_counts(normalize=True) * survey_.groupby('vote_senate')['lv_weight_2020'].mean() * 100
votes = votes[options] * (100 / votes[options].sum())
print(votes)
print_result_sen(**calc_result_sen(dem_vote=votes['A Democratic candidate'],
rep_vote=votes['A Republican candidate'],
n=lv_weighted_n))
```
| github_jupyter |
# Import optimus and pandas
```
from optimus import Optimus
from pyspark.sql.functions import *
import pandas as pd
import numpy as np
op = Optimus()
```
# Read the data
```
url = 'https://raw.githubusercontent.com/justmarkham/DAT8/master/data/chipotle.tsv'
chipo_pd = pd.read_csv(url, sep = '\t')
chipo_pd.dtypes
```
### We need to to this to be able to read the data from Spark
```
chipo_pd[['item_name', 'choice_description', 'item_price']] = chipo_pd[['item_name', 'choice_description', 'item_price']].astype(str)
```
# Transform data from Pandas to Optimus (Spark)
```
chipo = op.spark.createDataFrame(chipo_pd)
```
# See the first 10 entries
```
chipo.table(10)
```
# What is the number of observations and columns in the dataset?
```
chipo.count()
op.profiler.dataset_info(chipo)
```
# Print the name of all the columns
```
chipo.columns
```
**NOTE: Spark dataframes are not indexed.**
# Which was the most-ordered item and how many items were ordered?
```
# Here we are renaming the column sum(quantity) to quantity with Optimus function rename inside of cols
(chipo.groupby("item_name")
.sum("quantity")
.cols.rename("sum(quantity)", "quantity")
.sort(desc("quantity"))
.table(1))
```
# What was the most ordered item in the choice_description column
```
(chipo.groupby("choice_description")
.sum("quantity")
.cols.rename("sum(quantity)", "quantity")
.sort(desc("quantity"))
.table(1))
```
Here we have a problem, is showing that nan was the most order item from `choice_desccription`. Let's solve that:
```
# First we are transforming "nan" strings to real nulls, and then droping them
(chipo.cols.replace("choice_description","nan")
.dropna()
.groupby("choice_description")
.sum("quantity")
.cols.rename("sum(quantity)", "quantity")
.sort(desc("quantity"))
.table(1))
```
# How many items were orderd in total?
```
chipo.cols.sum("quantity")
```
# Turn the item price into a float
```
chipo.dtypes
## Let's see the format of the price
chipo.table(1)
# Use substr (like in SQL) to get from the first numer to the end and then cast it
chipo = chipo.withColumn("item_price", chipo.item_price.substr(2,10).cast("float"))
# Let's see our data now
chipo.table(2)
```
# How much was the revenue for the period in the dataset?
```
# The function mul takes two or more columns and multiples them
# The function sum will sum the values in a specific column
revenue = (chipo.cols.mul(columns=["quantity", "item_price"])
.cols.sum("mul"))
print('Revenue was: $' + str(np.round(revenue,2)))
```
# How many orders were made in the period?
```
chipo.select("order_id").distinct().count()
```
# What is the average revenue amount per order?
```
(chipo.cols.mul(columns=["quantity", "item_price"])
.cols.rename("mul", "revenue")
.groupby("order_id").sum("revenue")
.cols.rename("sum(revenue)", "revenue")
.cols.mean("revenue"))
```
# How many different items are sold?
```
chipo.select("item_name").distinct().count()
```
| github_jupyter |
# T81-558: Applications of Deep Neural Networks
**Module 7: Generative Adversarial Networks**
* Instructor: [Jeff Heaton](https://sites.wustl.edu/jeffheaton/), McKelvey School of Engineering, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
* For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
# Module 7 Material
* Part 7.1: Introduction to GANS for Image and Data Generation [[Video]](https://www.youtube.com/watch?v=0QnCH6tlZgc&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_07_1_gan_intro.ipynb)
* **Part 7.2: Implementing a GAN in Keras** [[Video]](https://www.youtube.com/watch?v=T-MCludVNn4&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_07_2_Keras_gan.ipynb)
* Part 7.3: Face Generation with StyleGAN and Python [[Video]](https://www.youtube.com/watch?v=Wwwyr7cOBlU&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_07_3_style_gan.ipynb)
* Part 7.4: GANS for Semi-Supervised Learning in Keras [[Video]](https://www.youtube.com/watch?v=ZPewmEu7644&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_07_4_gan_semi_supervised.ipynb)
* Part 7.5: An Overview of GAN Research [[Video]](https://www.youtube.com/watch?v=cvCvZKvlvq4&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_07_5_gan_research.ipynb)
# Part 7.2: Implementing DCGANs in Keras
Paper that described the type of DCGAN that we will create in this module.
* Radford, A., Metz, L., & Chintala, S. (2015). [Unsupervised representation learning with deep convolutional generative adversarial networks](https://arxiv.org/abs/1511.06434). *arXiv preprint arXiv:1511.06434*.
This paper implements a DCGAN as follows:
* No pre-processing was applied to training images besides scaling to the range of the tanh activation function [-1, 1].
* All models were trained with mini-batch stochastic gradient descent (SGD) with a mini-batch size of 128.
* All weights were initialized from a zero-centered Normal distribution with standard deviation 0.02.
* In the LeakyReLU, the slope of the leak was set to 0.2 in all models.
* we used the Adam optimizer(Kingma & Ba, 2014) with tuned hyperparameters. We found the suggested learning rate of 0.001, to be too high, using 0.0002 instead.
* Additionally, we found leaving the momentum term $\beta{1}$ at the suggested value of 0.9 resulted in training oscillation and instability while reducing it to 0.5 helped stabilize training.
The paper also provides the following architecture guidelines for stable Deep Convolutional GANs:
* Replace any pooling layers with strided convolutions (discriminator) and fractional-strided convolutions (generator).
* Use batchnorm in both the generator and the discriminator.
* Remove fully connected hidden layers for deeper architectures.
* Use ReLU activation in generator for all layers except for the output, which uses Tanh.
* Use LeakyReLU activation in the discriminator for all layers.
While creating the material for this module I used a number of Internet resources, some of the most helpful were:
* [Keep Calm and train a GAN. Pitfalls and Tips on training Generative Adversarial Networks](https://medium.com/@utk.is.here/keep-calm-and-train-a-gan-pitfalls-and-tips-on-training-generative-adversarial-networks-edd529764aa9)
* [Collection of Keras implementations of Generative Adversarial Networks GANs](https://github.com/eriklindernoren/Keras-GAN)
* [dcgan-facegenerator](https://github.com/platonovsimeon/dcgan-facegenerator), [Semi-Paywalled Article by GitHub Author](https://medium.com/datadriveninvestor/generating-human-faces-with-keras-3ccd54c17f16)
The program created next will generate faces similar to these. While these faces are not perfect, they demonstrate how we can construct and train a GAN on or own. Later we will see how to import very advanced weights from nVidia to produce high resolution, realistic looking faces.

The following packages will be used to implement a basic GAN system in Python/Keras.
```
from keras.layers import Input, Reshape, Dropout, Dense, Flatten, BatchNormalization, Activation, ZeroPadding2D
from keras.layers.advanced_activations import LeakyReLU
from keras.layers.convolutional import UpSampling2D, Conv2D
from keras.models import Sequential, Model, load_model
from keras.optimizers import Adam
import numpy as np
from PIL import Image
from tqdm import tqdm
import os
```
I suggest running this code with a GPU, it will be very slow on a CPU alone. The following code mounts your Google drive for use with Google CoLab. If you are not using CoLab, the following code will not work.
```
# Run this for Google CoLab
from google.colab import drive
drive.mount('/content/drive')
```
These are the constants that define how the GANs will be created for this example. The higher the resolution, the more memory that will be needed. Higher resolution will also result in longer run times. For Google CoLab (with GPU) 128x128 resolution is as high as can be used (due to memoru). Note that the resolution is specified as a multiple of 32. So **GENERATE_RES** of 1 is 32, 2 is 64, etc.
To run this you will need training data. The training data can be any collection of images. I suggest using training data from the following two locations. Simply unzip and combine to a common directory. This directory should be uploaded to Google Drive (if you are using CoLab). The constant **DATA_PATH** defines where these images are stored.
The source data (faces) used in this module can be found here:
* [Kaggle Faces Data New](https://www.kaggle.com/gasgallo/faces-data-new)
* [Kaggle Lag Dataset: Dataset of faces, from more than 1k different subjects](https://www.kaggle.com/gasgallo/lag-dataset)
```
# Generation resolution - Must be square
# Training data is also scaled to this.
# Note GENERATE_RES higher than 4 will blow Google CoLab's memory.
GENERATE_RES = 2 # (1=32, 2=64, 3=96, etc.)
GENERATE_SQUARE = 32 * GENERATE_RES # rows/cols (should be square)
IMAGE_CHANNELS = 3
# Preview image
PREVIEW_ROWS = 4
PREVIEW_COLS = 7
PREVIEW_MARGIN = 16
SAVE_FREQ = 100
# Size vector to generate images from
SEED_SIZE = 100
# Configuration
DATA_PATH = '/content/drive/My Drive/projects/faces'
EPOCHS = 10000
BATCH_SIZE = 32
print(f"Will generate {GENERATE_SQUARE}px square images.")
```
Next we will load and preprocess the images. This can take awhile. Google CoLab took around an hour to process. Because of this we store the processed file as a binary. This way we can simply reload the processed training data and quickly use it. It is most efficient to only perform this operation once. The dimensions of the image are encoded into the filename of the binary file because we need to regenerate it if these change.
```
# Image set has 11,682 images. Can take over an hour for initial preprocessing.
# Because of this time needed, save a Numpy preprocessed file.
# Note, that file is large enough to cause problems for sume verisons of Pickle,
# so Numpy binary files are used.
training_binary_path = os.path.join(DATA_PATH,f'training_data_{GENERATE_SQUARE}_{GENERATE_SQUARE}.npy')
print(f"Looking for file: {training_binary_path}")
if not os.path.isfile(training_binary_path):
print("Loading training images...")
training_data = []
faces_path = os.path.join(DATA_PATH,'face_images')
for filename in tqdm(os.listdir(faces_path)):
path = os.path.join(faces_path,filename)
image = Image.open(path).resize((GENERATE_SQUARE,GENERATE_SQUARE),Image.ANTIALIAS)
training_data.append(np.asarray(image))
training_data = np.reshape(training_data,(-1,GENERATE_SQUARE,GENERATE_SQUARE,IMAGE_CHANNELS))
training_data = training_data / 127.5 - 1.
print("Saving training image binary...")
np.save(training_binary_path,training_data)
else:
print("Loading previous training pickle...")
training_data = np.load(training_binary_path)
```
The code below creates the generator and discriminator.
```
def build_generator(seed_size, channels):
model = Sequential()
model.add(Dense(4*4*256,activation="relu",input_dim=seed_size))
model.add(Reshape((4,4,256)))
model.add(UpSampling2D())
model.add(Conv2D(256,kernel_size=3,padding="same"))
model.add(BatchNormalization(momentum=0.8))
model.add(Activation("relu"))
model.add(UpSampling2D())
model.add(Conv2D(256,kernel_size=3,padding="same"))
model.add(BatchNormalization(momentum=0.8))
model.add(Activation("relu"))
# Output resolution, additional upsampling
for i in range(GENERATE_RES):
model.add(UpSampling2D())
model.add(Conv2D(128,kernel_size=3,padding="same"))
model.add(BatchNormalization(momentum=0.8))
model.add(Activation("relu"))
# Final CNN layer
model.add(Conv2D(channels,kernel_size=3,padding="same"))
model.add(Activation("tanh"))
input = Input(shape=(seed_size,))
generated_image = model(input)
return Model(input,generated_image)
def build_discriminator(image_shape):
model = Sequential()
model.add(Conv2D(32, kernel_size=3, strides=2, input_shape=image_shape, padding="same"))
model.add(LeakyReLU(alpha=0.2))
model.add(Dropout(0.25))
model.add(Conv2D(64, kernel_size=3, strides=2, padding="same"))
model.add(ZeroPadding2D(padding=((0,1),(0,1))))
model.add(BatchNormalization(momentum=0.8))
model.add(LeakyReLU(alpha=0.2))
model.add(Dropout(0.25))
model.add(Conv2D(128, kernel_size=3, strides=2, padding="same"))
model.add(BatchNormalization(momentum=0.8))
model.add(LeakyReLU(alpha=0.2))
model.add(Dropout(0.25))
model.add(Conv2D(256, kernel_size=3, strides=1, padding="same"))
model.add(BatchNormalization(momentum=0.8))
model.add(LeakyReLU(alpha=0.2))
model.add(Dropout(0.25))
model.add(Conv2D(512, kernel_size=3, strides=1, padding="same"))
model.add(BatchNormalization(momentum=0.8))
model.add(LeakyReLU(alpha=0.2))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(1, activation='sigmoid'))
input_image = Input(shape=image_shape)
validity = model(input_image)
return Model(input_image, validity)
def save_images(cnt,noise):
image_array = np.full((
PREVIEW_MARGIN + (PREVIEW_ROWS * (GENERATE_SQUARE+PREVIEW_MARGIN)),
PREVIEW_MARGIN + (PREVIEW_COLS * (GENERATE_SQUARE+PREVIEW_MARGIN)), 3),
255, dtype=np.uint8)
generated_images = generator.predict(noise)
generated_images = 0.5 * generated_images + 0.5
image_count = 0
for row in range(PREVIEW_ROWS):
for col in range(PREVIEW_COLS):
r = row * (GENERATE_SQUARE+16) + PREVIEW_MARGIN
c = col * (GENERATE_SQUARE+16) + PREVIEW_MARGIN
image_array[r:r+GENERATE_SQUARE,c:c+GENERATE_SQUARE] = generated_images[image_count] * 255
image_count += 1
output_path = os.path.join(DATA_PATH,'output')
if not os.path.exists(output_path):
os.makedirs(output_path)
filename = os.path.join(output_path,f"train-{cnt}.png")
im = Image.fromarray(image_array)
im.save(filename)
```
Next we actually build the discriminator and the generator. Both will be trained with the Adam optimizer.
```
image_shape = (GENERATE_SQUARE,GENERATE_SQUARE,IMAGE_CHANNELS)
optimizer = Adam(1.5e-4,0.5) # learning rate and momentum adjusted from paper
discriminator = build_discriminator(image_shape)
discriminator.compile(loss="binary_crossentropy",optimizer=optimizer,metrics=["accuracy"])
generator = build_generator(SEED_SIZE,IMAGE_CHANNELS)
random_input = Input(shape=(SEED_SIZE,))
generated_image = generator(random_input)
discriminator.trainable = False
validity = discriminator(generated_image)
combined = Model(random_input,validity)
combined.compile(loss="binary_crossentropy",optimizer=optimizer,metrics=["accuracy"])
y_real = np.ones((BATCH_SIZE,1))
y_fake = np.zeros((BATCH_SIZE,1))
fixed_seed = np.random.normal(0, 1, (PREVIEW_ROWS * PREVIEW_COLS, SEED_SIZE))
cnt = 1
for epoch in range(EPOCHS):
idx = np.random.randint(0,training_data.shape[0],BATCH_SIZE)
x_real = training_data[idx]
# Generate some images
seed = np.random.normal(0,1,(BATCH_SIZE,SEED_SIZE))
x_fake = generator.predict(seed)
# Train discriminator on real and fake
discriminator_metric_real = discriminator.train_on_batch(x_real,y_real)
discriminator_metric_generated = discriminator.train_on_batch(x_fake,y_fake)
discriminator_metric = 0.5 * np.add(discriminator_metric_real,discriminator_metric_generated)
# Train generator on Calculate losses
generator_metric = combined.train_on_batch(seed,y_real)
# Time for an update?
if epoch % SAVE_FREQ == 0:
save_images(cnt, fixed_seed)
cnt += 1
print(f"Epoch {epoch}, Discriminator accuarcy: {discriminator_metric[1]}, Generator accuracy: {generator_metric[1]}")
generator.save(os.path.join(DATA_PATH,"face_generator.h5"))
```
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Load images with tf.data
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/alpha/tutorials/load_data/images"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/r2/tutorials/load_data/images.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/r2/tutorials/load_data/images.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
</table>
This tutorial provides a simple example of how to load an image dataset using `tf.data`.
The dataset used in this example is distributed as directories of images, with one class of image per directory.
## Setup
```
from __future__ import absolute_import, division, print_function, unicode_literals
!pip install tensorflow==2.0.0-alpha0
import tensorflow as tf
AUTOTUNE = tf.data.experimental.AUTOTUNE
```
## Download and inspect the dataset
### Retrieve the images
Before you start any training, you'll need a set of images to teach the network about the new classes you want to recognize. We've created an archive of creative-commons licensed flower photos to use initially.
```
import pathlib
data_root_orig = tf.keras.utils.get_file(origin='https://storage.googleapis.com/download.tensorflow.org/example_images/flower_photos.tgz',
fname='flower_photos', untar=True)
data_root = pathlib.Path(data_root_orig)
print(data_root)
```
After downloading 218MB, you should now have a copy of the flower photos available:
```
for item in data_root.iterdir():
print(item)
import random
all_image_paths = list(data_root.glob('*/*'))
all_image_paths = [str(path) for path in all_image_paths]
random.shuffle(all_image_paths)
image_count = len(all_image_paths)
image_count
all_image_paths[:10]
```
### Inspect the images
Now let's have a quick look at a couple of the images, so we know what we're dealing with:
```
import os
attributions = (data_root/"LICENSE.txt").open(encoding='utf-8').readlines()[4:]
attributions = [line.split(' CC-BY') for line in attributions]
attributions = dict(attributions)
import IPython.display as display
def caption_image(image_path):
image_rel = pathlib.Path(image_path).relative_to(data_root)
return "Image (CC BY 2.0) " + ' - '.join(attributions[str(image_rel)].split(' - ')[:-1])
for n in range(3):
image_path = random.choice(all_image_paths)
display.display(display.Image(image_path))
print(caption_image(image_path))
print()
```
### Determine the label for each image
List the available labels:
```
label_names = sorted(item.name for item in data_root.glob('*/') if item.is_dir())
label_names
```
Assign an index to each label:
```
label_to_index = dict((name, index) for index,name in enumerate(label_names))
label_to_index
```
Create a list of every file, and its label index
```
all_image_labels = [label_to_index[pathlib.Path(path).parent.name]
for path in all_image_paths]
print("First 10 labels indices: ", all_image_labels[:10])
```
### Load and format the images
TensorFlow includes all the tools you need to load and process images:
```
img_path = all_image_paths[0]
img_path
```
here is the raw data:
```
img_raw = tf.io.read_file(img_path)
print(repr(img_raw)[:100]+"...")
```
Decode it into an image tensor:
```
img_tensor = tf.image.decode_image(img_raw)
print(img_tensor.shape)
print(img_tensor.dtype)
```
Resize it for your model:
```
img_final = tf.image.resize(img_tensor, [192, 192])
img_final = img_final/255.0
print(img_final.shape)
print(img_final.numpy().min())
print(img_final.numpy().max())
```
Wrap up these up in simple functions for later.
```
def preprocess_image(image):
image = tf.image.decode_jpeg(image, channels=3)
image = tf.image.resize(image, [192, 192])
image /= 255.0 # normalize to [0,1] range
return image
def load_and_preprocess_image(path):
image = tf.io.read_file(path)
return preprocess_image(image)
import matplotlib.pyplot as plt
image_path = all_image_paths[0]
label = all_image_labels[0]
plt.imshow(load_and_preprocess_image(img_path))
plt.grid(False)
plt.xlabel(caption_image(img_path))
plt.title(label_names[label].title())
print()
```
## Build a `tf.data.Dataset`
### A dataset of images
The easiest way to build a `tf.data.Dataset` is using the `from_tensor_slices` method.
Slicing the array of strings, results in a dataset of strings:
```
path_ds = tf.data.Dataset.from_tensor_slices(all_image_paths)
```
The `shapes` and `types` describe the content of each item in the dataset. In this case it is a set of scalar binary-strings
```
print(path_ds)
```
Now create a new dataset that loads and formats images on the fly by mapping `preprocess_image` over the dataset of paths.
```
image_ds = path_ds.map(load_and_preprocess_image, num_parallel_calls=AUTOTUNE)
import matplotlib.pyplot as plt
plt.figure(figsize=(8,8))
for n,image in enumerate(image_ds.take(4)):
plt.subplot(2,2,n+1)
plt.imshow(image)
plt.grid(False)
plt.xticks([])
plt.yticks([])
plt.xlabel(caption_image(all_image_paths[n]))
plt.show()
```
### A dataset of `(image, label)` pairs
Using the same `from_tensor_slices` method we can build a dataset of labels
```
label_ds = tf.data.Dataset.from_tensor_slices(tf.cast(all_image_labels, tf.int64))
for label in label_ds.take(10):
print(label_names[label.numpy()])
```
Since the datasets are in the same order we can just zip them together to get a dataset of `(image, label)` pairs.
```
image_label_ds = tf.data.Dataset.zip((image_ds, label_ds))
```
The new dataset's `shapes` and `types` are tuples of shapes and types as well, describing each field:
```
print(image_label_ds)
```
Note: When you have arrays like `all_image_labels` and `all_image_paths` an alternative to `tf.data.dataset.Dataset.zip` is to slice the pair of arrays.
```
ds = tf.data.Dataset.from_tensor_slices((all_image_paths, all_image_labels))
# The tuples are unpacked into the positional arguments of the mapped function
def load_and_preprocess_from_path_label(path, label):
return load_and_preprocess_image(path), label
image_label_ds = ds.map(load_and_preprocess_from_path_label)
image_label_ds
```
### Basic methods for training
To train a model with this dataset you will want the data:
* To be well shuffled.
* To be batched.
* To repeat forever.
* Batches to be available as soon as possible.
These features can be easily added using the `tf.data` api.
```
BATCH_SIZE = 32
# Setting a shuffle buffer size as large as the dataset ensures that the data is
# completely shuffled.
ds = image_label_ds.shuffle(buffer_size=image_count)
ds = ds.repeat()
ds = ds.batch(BATCH_SIZE)
# `prefetch` lets the dataset fetch batches, in the background while the model is training.
ds = ds.prefetch(buffer_size=AUTOTUNE)
ds
```
There are a few things to note here:
1. The order is important.
* A `.shuffle` after a `.repeat` would shuffle items across epoch boundaries (some items will be seen twice before others are seen at all).
* A `.shuffle` after a `.batch` would shuffle the order of the batches, but not shuffle the items across batches.
1. We use a `buffer_size` the same size as the dataset for a full shuffle. Up to the dataset size, large values provide better randomization, but use more memory.
1. The shuffle buffer is filled before any elements are pulled from it. So a large `buffer_size` may cause a delay when your `Dataset` is starting.
1. The shuffeled dataset doesn't report the end of a dataset until the shuffle-buffer is completely empty. The `Dataset` is restarted by `.repeat`, causing another wait for the shuffle-buffer to be filled.
This last point can be addressed by using the `tf.data.Dataset.apply` method with the fused `tf.data.experimental.shuffle_and_repeat` function:
```
ds = image_label_ds.apply(
tf.data.experimental.shuffle_and_repeat(buffer_size=image_count))
ds = ds.batch(BATCH_SIZE)
ds = ds.prefetch(buffer_size=AUTOTUNE)
ds
```
### Pipe the dataset to a model
Fetch a copy of MobileNet v2 from `tf.keras.applications`.
This will be used for a simple transfer learning example.
Set the MobileNet weights to be non-trainable:
```
mobile_net = tf.keras.applications.MobileNetV2(input_shape=(192, 192, 3), include_top=False)
mobile_net.trainable=False
```
This model expects its input to be normalized to the `[-1,1]` range:
```
help(keras_applications.mobilenet_v2.preprocess_input)
```
<pre>
...
This function applies the "Inception" preprocessing which converts
the RGB values from [0, 255] to [-1, 1]
...
</pre>
So before the passing it to the MobilNet model, we need to convert the input from a range of `[0,1]` to `[-1,1]`.
```
def change_range(image,label):
return 2*image-1, label
keras_ds = ds.map(change_range)
```
The MobileNet returns a `6x6` spatial grid of features for each image.
Pass it a batch of images to see:
```
# The dataset may take a few seconds to start, as it fills its shuffle buffer.
image_batch, label_batch = next(iter(keras_ds))
feature_map_batch = mobile_net(image_batch)
print(feature_map_batch.shape)
```
So build a model wrapped around MobileNet, and use `tf.keras.layers.GlobalAveragePooling2D` to average over those space dimensions, before the output `tf.keras.layers.Dense` layer:
```
model = tf.keras.Sequential([
mobile_net,
tf.keras.layers.GlobalAveragePooling2D(),
tf.keras.layers.Dense(len(label_names))])
```
Now it produces outputs of the expected shape:
```
logit_batch = model(image_batch).numpy()
print("min logit:", logit_batch.min())
print("max logit:", logit_batch.max())
print()
print("Shape:", logit_batch.shape)
```
Compile the model to describe the training procedure:
```
model.compile(optimizer=tf.keras.optimizers.Adam(),
loss='sparse_categorical_crossentropy',
metrics=["accuracy"])
```
There are 2 trainable variables: the Dense `weights` and `bias`:
```
len(model.trainable_variables)
model.summary()
```
Train the model.
Normally you would specify the real number of steps per epoch, but for demonstration purposes only run 3 steps.
```
steps_per_epoch=tf.math.ceil(len(all_image_paths)/BATCH_SIZE).numpy()
steps_per_epoch
model.fit(ds, epochs=1, steps_per_epoch=3)
```
## Performance
Note: This section just shows a couple of easy tricks that may help performance. For an in depth guide see [Input Pipeline Performance](https://www.tensorflow.org/guide/performance/datasets).
The simple pipeline used above reads each file individually, on each epoch. This is fine for local training on CPU but may not be sufficient for GPU training, and is totally inappropriate for any sort of distributed training.
To investigate, first build a simple function to check the performance of our datasets:
```
import time
default_timeit_steps = 2*steps_per_epoch+1
def timeit(ds, steps=default_timeit_steps):
overall_start = time.time()
# Fetch a single batch to prime the pipeline (fill the shuffle buffer),
# before starting the timer
it = iter(ds.take(steps+1))
next(it)
start = time.time()
for i,(images,labels) in enumerate(it):
if i%10 == 0:
print('.',end='')
print()
end = time.time()
duration = end-start
print("{} batches: {} s".format(steps, duration))
print("{:0.5f} Images/s".format(BATCH_SIZE*steps/duration))
print("Total time: {}s".format(end-overall_start))
```
The performance of the current dataset is:
```
ds = image_label_ds.apply(
tf.data.experimental.shuffle_and_repeat(buffer_size=image_count))
ds = ds.batch(BATCH_SIZE).prefetch(buffer_size=AUTOTUNE)
ds
timeit(ds)
```
### Cache
Use `tf.data.Dataset.cache` to easily cache calculations across epochs. This is especially performant if the dataq fits in memory.
Here the images are cached, after being pre-precessed (decoded and resized):
```
ds = image_label_ds.cache()
ds = ds.apply(
tf.data.experimental.shuffle_and_repeat(buffer_size=image_count))
ds = ds.batch(BATCH_SIZE).prefetch(buffer_size=AUTOTUNE)
ds
timeit(ds)
```
One disadvantage to using an in memory cache is that the cache must be rebuilt on each run, giving the same startup delay each time the dataset is started:
```
timeit(ds)
```
If the data doesn't fit in memory, use a cache file:
```
ds = image_label_ds.cache(filename='./cache.tf-data')
ds = ds.apply(
tf.data.experimental.shuffle_and_repeat(buffer_size=image_count))
ds = ds.batch(BATCH_SIZE).prefetch(1)
ds
timeit(ds)
```
The cache file also has the advantage that it can be used to quickly restart the dataset without rebuilding the cache. Note how much faster it is the second time:
```
timeit(ds)
```
### TFRecord File
#### Raw image data
TFRecord files are a simple format to store a sequence of binary blobs. By packing multiple examples into the same file, TensorFlow is able to read multiple examples at once, which is especially important for performance when using a remote storage service such as GCS.
First, build a TFRecord file from the raw image data:
```
image_ds = tf.data.Dataset.from_tensor_slices(all_image_paths).map(tf.io.read_file)
tfrec = tf.data.experimental.TFRecordWriter('images.tfrec')
tfrec.write(image_ds)
```
Next build a dataset that reads from the TFRecord file and decodes/reformats the images using the `preprocess_image` function we defined earlier.
```
image_ds = tf.data.TFRecordDataset('images.tfrec').map(preprocess_image)
```
Zip that with the labels dataset we defined earlier, to get the expected `(image,label)` pairs.
```
ds = tf.data.Dataset.zip((image_ds, label_ds))
ds = ds.apply(
tf.data.experimental.shuffle_and_repeat(buffer_size=image_count))
ds=ds.batch(BATCH_SIZE).prefetch(AUTOTUNE)
ds
timeit(ds)
```
This is slower than the `cache` version because we have not cached the preprocessing.
#### Serialized Tensors
To save some preprocessing to the TFRecord file, first make a dataset of the processed images, as before:
```
paths_ds = tf.data.Dataset.from_tensor_slices(all_image_paths)
image_ds = paths_ds.map(load_and_preprocess_image)
image_ds
```
Now instead of a dataset of `.jpeg` strings, this is a dataset of tensors.
To serialize this to a TFRecord file you first convert the dataset of tensors to a dataset of strings.
```
ds = image_ds.map(tf.io.serialize_tensor)
ds
tfrec = tf.data.experimental.TFRecordWriter('images.tfrec')
tfrec.write(ds)
```
With the preprocessing cached, data can be loaded from the TFrecord file quite efficiently. Just remember to de-serialized tensor before trying to use it.
```
ds = tf.data.TFRecordDataset('images.tfrec')
def parse(x):
result = tf.io.parse_tensor(x, out_type=tf.float32)
result = tf.reshape(result, [192, 192, 3])
return result
ds = ds.map(parse, num_parallel_calls=AUTOTUNE)
ds
```
Now, add the labels and apply the same standard operations as before:
```
ds = tf.data.Dataset.zip((ds, label_ds))
ds = ds.apply(
tf.data.experimental.shuffle_and_repeat(buffer_size=image_count))
ds=ds.batch(BATCH_SIZE).prefetch(AUTOTUNE)
ds
timeit(ds)
```
| github_jupyter |
# Mask R-CNN - Train on Shapes Dataset
This notebook shows how to train Mask R-CNN on your own dataset. To keep things simple we use a synthetic dataset of shapes (squares, triangles, and circles) which enables fast training. You'd still need a GPU, though, because the network backbone is a Resnet101, which would be too slow to train on a CPU. On a GPU, you can start to get okay-ish results in a few minutes, and good results in less than an hour.
The code of the *Shapes* dataset is included below. It generates images on the fly, so it doesn't require downloading any data. And it can generate images of any size, so we pick a small image size to train faster.
```
import os
import sys
import random
import math
import re
import time
import numpy as np
import cv2
import matplotlib
import matplotlib.pyplot as plt
# Root directory of the project
ROOT_DIR = os.path.abspath("../../")
# Import Mask RCNN
sys.path.append(ROOT_DIR) # To find local version of the library
from mrcnn.config import Config
from mrcnn import utils
import mrcnn.model as modellib
from mrcnn import visualize
from mrcnn.model import log
%matplotlib inline
# Directory to save logs and trained model
MODEL_DIR = os.path.join(ROOT_DIR, "logs")
# Local path to trained weights file
COCO_MODEL_PATH = os.path.join(ROOT_DIR, "mask_rcnn_coco.h5")
# Download COCO trained weights from Releases if needed
if not os.path.exists(COCO_MODEL_PATH):
utils.download_trained_weights(COCO_MODEL_PATH)
```
## Configurations
```
class ShapesConfig(Config):
"""Configuration for training on the toy shapes dataset.
Derives from the base Config class and overrides values specific
to the toy shapes dataset.
"""
# Give the configuration a recognizable name
NAME = "shapes"
# Train on 1 GPU and 8 images per GPU. We can put multiple images on each
# GPU because the images are small. Batch size is 8 (GPUs * images/GPU).
GPU_COUNT = 1
IMAGES_PER_GPU = 8
# Number of classes (including background)
NUM_CLASSES = 1 + 3 # background + 3 shapes
# Use small images for faster training. Set the limits of the small side
# the large side, and that determines the image shape.
IMAGE_MIN_DIM = 128
IMAGE_MAX_DIM = 128
# Use smaller anchors because our image and objects are small
RPN_ANCHOR_SCALES = (8, 16, 32, 64, 128) # anchor side in pixels
# Reduce training ROIs per image because the images are small and have
# few objects. Aim to allow ROI sampling to pick 33% positive ROIs.
TRAIN_ROIS_PER_IMAGE = 32
# Use a small epoch since the data is simple
STEPS_PER_EPOCH = 100
# use small validation steps since the epoch is small
VALIDATION_STEPS = 5
config = ShapesConfig()
config.display()
```
## Notebook Preferences
```
def get_ax(rows=1, cols=1, size=8):
"""Return a Matplotlib Axes array to be used in
all visualizations in the notebook. Provide a
central point to control graph sizes.
Change the default size attribute to control the size
of rendered images
"""
_, ax = plt.subplots(rows, cols, figsize=(size*cols, size*rows))
return ax
```
## Dataset
Create a synthetic dataset
Extend the Dataset class and add a method to load the shapes dataset, `load_shapes()`, and override the following methods:
* load_image()
* load_mask()
* image_reference()
```
class ShapesDataset(utils.Dataset):
"""Generates the shapes synthetic dataset. The dataset consists of simple
shapes (triangles, squares, circles) placed randomly on a blank surface.
The images are generated on the fly. No file access required.
"""
def load_shapes(self, count, height, width):
"""Generate the requested number of synthetic images.
count: number of images to generate.
height, width: the size of the generated images.
"""
# Add classes
self.add_class("shapes", 1, "square")
self.add_class("shapes", 2, "circle")
self.add_class("shapes", 3, "triangle")
# Add images
# Generate random specifications of images (i.e. color and
# list of shapes sizes and locations). This is more compact than
# actual images. Images are generated on the fly in load_image().
for i in range(count):
bg_color, shapes = self.random_image(height, width)
self.add_image("shapes", image_id=i, path=None,
width=width, height=height,
bg_color=bg_color, shapes=shapes)
def load_image(self, image_id):
"""Generate an image from the specs of the given image ID.
Typically this function loads the image from a file, but
in this case it generates the image on the fly from the
specs in image_info.
"""
info = self.image_info[image_id]
bg_color = np.array(info['bg_color']).reshape([1, 1, 3])
image = np.ones([info['height'], info['width'], 3], dtype=np.uint8)
image = image * bg_color.astype(np.uint8)
for shape, color, dims in info['shapes']:
image = self.draw_shape(image, shape, dims, color)
return image
def image_reference(self, image_id):
"""Return the shapes data of the image."""
info = self.image_info[image_id]
if info["source"] == "shapes":
return info["shapes"]
else:
super(self.__class__).image_reference(self, image_id)
def load_mask(self, image_id):
"""Generate instance masks for shapes of the given image ID.
"""
info = self.image_info[image_id]
shapes = info['shapes']
count = len(shapes)
mask = np.zeros([info['height'], info['width'], count], dtype=np.uint8)
for i, (shape, _, dims) in enumerate(info['shapes']):
mask[:, :, i:i+1] = self.draw_shape(mask[:, :, i:i+1].copy(),
shape, dims, 1)
# Handle occlusions
occlusion = np.logical_not(mask[:, :, -1]).astype(np.uint8)
for i in range(count-2, -1, -1):
mask[:, :, i] = mask[:, :, i] * occlusion
occlusion = np.logical_and(occlusion, np.logical_not(mask[:, :, i]))
# Map class names to class IDs.
class_ids = np.array([self.class_names.index(s[0]) for s in shapes])
return mask.astype(np.bool), class_ids.astype(np.int32)
def draw_shape(self, image, shape, dims, color):
"""Draws a shape from the given specs."""
# Get the center x, y and the size s
x, y, s = dims
if shape == 'square':
cv2.rectangle(image, (x-s, y-s), (x+s, y+s), color, -1)
elif shape == "circle":
cv2.circle(image, (x, y), s, color, -1)
elif shape == "triangle":
points = np.array([[(x, y-s),
(x-s/math.sin(math.radians(60)), y+s),
(x+s/math.sin(math.radians(60)), y+s),
]], dtype=np.int32)
cv2.fillPoly(image, points, color)
return image
def random_shape(self, height, width):
"""Generates specifications of a random shape that lies within
the given height and width boundaries.
Returns a tuple of three valus:
* The shape name (square, circle, ...)
* Shape color: a tuple of 3 values, RGB.
* Shape dimensions: A tuple of values that define the shape size
and location. Differs per shape type.
"""
# Shape
shape = random.choice(["square", "circle", "triangle"])
# Color
color = tuple([random.randint(0, 255) for _ in range(3)])
# Center x, y
buffer = 20
y = random.randint(buffer, height - buffer - 1)
x = random.randint(buffer, width - buffer - 1)
# Size
s = random.randint(buffer, height//4)
return shape, color, (x, y, s)
def random_image(self, height, width):
"""Creates random specifications of an image with multiple shapes.
Returns the background color of the image and a list of shape
specifications that can be used to draw the image.
"""
# Pick random background color
bg_color = np.array([random.randint(0, 255) for _ in range(3)])
# Generate a few random shapes and record their
# bounding boxes
shapes = []
boxes = []
N = random.randint(1, 4)
for _ in range(N):
shape, color, dims = self.random_shape(height, width)
shapes.append((shape, color, dims))
x, y, s = dims
boxes.append([y-s, x-s, y+s, x+s])
# Apply non-max suppression wit 0.3 threshold to avoid
# shapes covering each other
keep_ixs = utils.non_max_suppression(np.array(boxes), np.arange(N), 0.3)
shapes = [s for i, s in enumerate(shapes) if i in keep_ixs]
return bg_color, shapes
# Training dataset
dataset_train = ShapesDataset()
dataset_train.load_shapes(500, config.IMAGE_SHAPE[0], config.IMAGE_SHAPE[1])
dataset_train.prepare()
# Validation dataset
dataset_val = ShapesDataset()
dataset_val.load_shapes(50, config.IMAGE_SHAPE[0], config.IMAGE_SHAPE[1])
dataset_val.prepare()
# Load and display random samples
image_ids = np.random.choice(dataset_train.image_ids, 4)
for image_id in image_ids:
image = dataset_train.load_image(image_id)
mask, class_ids = dataset_train.load_mask(image_id)
visualize.display_top_masks(image, mask, class_ids, dataset_train.class_names)
```
## Create Model
```
# Create model in training mode
model = modellib.MaskRCNN(mode="training", config=config,
model_dir=MODEL_DIR)
# Which weights to start with?
init_with = "coco" # imagenet, coco, or last
if init_with == "imagenet":
model.load_weights(model.get_imagenet_weights(), by_name=True)
elif init_with == "coco":
# Load weights trained on MS COCO, but skip layers that
# are different due to the different number of classes
# See README for instructions to download the COCO weights
model.load_weights(COCO_MODEL_PATH, by_name=True,
exclude=["mrcnn_class_logits", "mrcnn_bbox_fc",
"mrcnn_bbox", "mrcnn_mask"])
elif init_with == "last":
# Load the last model you trained and continue training
model.load_weights(model.find_last(), by_name=True)
```
## Training
Train in two stages:
1. Only the heads. Here we're freezing all the backbone layers and training only the randomly initialized layers (i.e. the ones that we didn't use pre-trained weights from MS COCO). To train only the head layers, pass `layers='heads'` to the `train()` function.
2. Fine-tune all layers. For this simple example it's not necessary, but we're including it to show the process. Simply pass `layers="all` to train all layers.
```
# Train the head branches
# Passing layers="heads" freezes all layers except the head
# layers. You can also pass a regular expression to select
# which layers to train by name pattern.
model.train(dataset_train, dataset_val,
learning_rate=config.LEARNING_RATE,
epochs=1,
layers='heads')
# Fine tune all layers
# Passing layers="all" trains all layers. You can also
# pass a regular expression to select which layers to
# train by name pattern.
model.train(dataset_train, dataset_val,
learning_rate=config.LEARNING_RATE / 10,
epochs=2,
layers="all")
# Save weights
# Typically not needed because callbacks save after every epoch
# Uncomment to save manually
# model_path = os.path.join(MODEL_DIR, "mask_rcnn_shapes.h5")
# model.keras_model.save_weights(model_path)
```
## Detection
```
class InferenceConfig(ShapesConfig):
GPU_COUNT = 1
IMAGES_PER_GPU = 1
inference_config = InferenceConfig()
# Recreate the model in inference mode
model = modellib.MaskRCNN(mode="inference",
config=inference_config,
model_dir=MODEL_DIR)
# Get path to saved weights
# Either set a specific path or find last trained weights
# model_path = os.path.join(ROOT_DIR, ".h5 file name here")
model_path = model.find_last()
# Load trained weights
print("Loading weights from ", model_path)
model.load_weights(model_path, by_name=True)
# Test on a random image
image_id = random.choice(dataset_val.image_ids)
original_image, image_meta, gt_class_id, gt_bbox, gt_mask =\
modellib.load_image_gt(dataset_val, inference_config,
image_id, use_mini_mask=False)
log("original_image", original_image)
log("image_meta", image_meta)
log("gt_class_id", gt_class_id)
log("gt_bbox", gt_bbox)
log("gt_mask", gt_mask)
visualize.display_instances(original_image, gt_bbox, gt_mask, gt_class_id,
dataset_train.class_names, figsize=(8, 8))
results = model.detect([original_image], verbose=1)
r = results[0]
visualize.display_instances(original_image, r['rois'], r['masks'], r['class_ids'],
dataset_val.class_names, r['scores'], ax=get_ax())
```
## Evaluation
```
# Compute VOC-Style mAP @ IoU=0.5
# Running on 10 images. Increase for better accuracy.
image_ids = np.random.choice(dataset_val.image_ids, 10)
APs = []
for image_id in image_ids:
# Load image and ground truth data
image, image_meta, gt_class_id, gt_bbox, gt_mask =\
modellib.load_image_gt(dataset_val, inference_config,
image_id, use_mini_mask=False)
molded_images = np.expand_dims(modellib.mold_image(image, inference_config), 0)
# Run object detection
results = model.detect([image], verbose=0)
r = results[0]
# Compute AP
AP, precisions, recalls, overlaps =\
utils.compute_ap(gt_bbox, gt_class_id, gt_mask,
r["rois"], r["class_ids"], r["scores"], r['masks'])
APs.append(AP)
print("mAP: ", np.mean(APs))
```
| github_jupyter |
<a href="https://colab.research.google.com/github/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_08_3_keras_hyperparameters.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# T81-558: Applications of Deep Neural Networks
**Module 8: Kaggle Data Sets**
* Instructor: [Jeff Heaton](https://sites.wustl.edu/jeffheaton/), McKelvey School of Engineering, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
* For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
# Module 8 Material
* Part 8.1: Introduction to Kaggle [[Video]](https://www.youtube.com/watch?v=v4lJBhdCuCU&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_08_1_kaggle_intro.ipynb)
* Part 8.2: Building Ensembles with Scikit-Learn and Keras [[Video]](https://www.youtube.com/watch?v=LQ-9ZRBLasw&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_08_2_keras_ensembles.ipynb)
* **Part 8.3: How Should you Architect Your Keras Neural Network: Hyperparameters** [[Video]](https://www.youtube.com/watch?v=1q9klwSoUQw&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_08_3_keras_hyperparameters.ipynb)
* Part 8.4: Bayesian Hyperparameter Optimization for Keras [[Video]](https://www.youtube.com/watch?v=sXdxyUCCm8s&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_08_4_bayesian_hyperparameter_opt.ipynb)
* Part 8.5: Current Semester's Kaggle [[Video]](https://www.youtube.com/watch?v=PHQt0aUasRg&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_08_5_kaggle_project.ipynb)
# Google CoLab Instructions
The following code ensures that Google CoLab is running the correct version of TensorFlow.
```
# Startup CoLab
try:
%tensorflow_version 2.x
COLAB = True
print("Note: using Google CoLab")
except:
print("Note: not using Google CoLab")
COLAB = False
# Nicely formatted time string
def hms_string(sec_elapsed):
h = int(sec_elapsed / (60 * 60))
m = int((sec_elapsed % (60 * 60)) / 60)
s = sec_elapsed % 60
return "{}:{:>02}:{:>05.2f}".format(h, m, s)
```
# Part 8.3: Architecting Network: Hyperparameters
You have probably noticed several hyperparameters introduced previously in this course that you need to choose for your neural network. The number of layers, neuron counts per layers, layer types, and activation functions are all choices you must make to optimize your neural network. Some of the categories of hyperparameters for you to choose from come from the following list:
* Number of Hidden Layers and Neuron Counts
* Activation Functions
* Advanced Activation Functions
* Regularization: L1, L2, Dropout
* Batch Normalization
* Training Parameters
The following sections will introduce each of these categories for Keras. While I will provide you with some general guidelines for hyperparameter selection; no two tasks are the same. You will benefit from experimentation with these values to determine what works best for your neural network. In the next part, we will see how machine learning can select some of these values on its own.
### Number of Hidden Layers and Neuron Counts
The structure of Keras layers is perhaps the hyperparameters that most become aware of first. How many layers should you have? How many neurons on each layer? What activation function and layer type should you use? These are all questions that come up when designing a neural network. There are many different [types of layer](https://keras.io/layers/core/) in Keras, listed here:
* **Activation** - You can also add activation functions as layers. Making use of the activation layer is the same as specifying the activation function as part of a Dense (or other) layer type.
* **ActivityRegularization** Used to add L1/L2 regularization outside of a layer. You can specify L1 and L2 as part of a Dense (or other) layer type.
* **Dense** - The original neural network layer type. In this layer type, every neuron connects to the next layer. The input vector is one-dimensional, and placing specific inputs next to each other does not affect.
* **Dropout** - Dropout consists of randomly setting a fraction rate of input units to 0 at each update during training time, which helps prevent overfitting. Dropout only occurs during training.
* **Flatten** - Flattens the input to 1D and does not affect the batch size.
* **Input** - A Keras tensor is a tensor object from the underlying back end (Theano, TensorFlow, or CNTK), which we augment with specific attributes to build a Keras model just by knowing the inputs and outputs of the model.
* **Lambda** - Wraps arbitrary expression as a Layer object.
* **Masking** - Masks a sequence by using a mask value to skip timesteps.
* **Permute** - Permutes the dimensions of the input according to a given pattern. Useful for tasks such as connecting RNNs and convolutional networks.
* **RepeatVector** - Repeats the input n times.
* **Reshape** - Similar to Numpy reshapes.
* **SpatialDropout1D** - This version performs the same function as Dropout; however, it drops entire 1D feature maps instead of individual elements.
* **SpatialDropout2D** - This version performs the same function as Dropout; however, it drops entire 2D feature maps instead of individual elements
* **SpatialDropout3D** - This version performs the same function as Dropout; however, it drops entire 3D feature maps instead of individual elements.
There is always trial and error for choosing a good number of neurons and hidden layers. Generally, the number of neurons on each layer will be larger closer to the hidden layer and smaller towards the output layer. This configuration gives the neural network a somewhat triangular or trapezoid appearance.
### Activation Functions
Activation functions are a choice that you must make for each layer. Generally, you can follow this guideline:
* Hidden Layers - RELU
* Output Layer - Softmax for classification, linear for regression.
Some of the common activation functions in Keras are listed here:
* **softmax** - Used for multi-class classification. Ensures all output neurons behave as probabilities and sum to 1.0.
* **elu** - Exponential linear unit. Exponential Linear Unit or its widely known name ELU is a function that tend to converge cost to zero faster and produce more accurate results. Can produce negative outputs.
* **selu** - Scaled Exponential Linear Unit (SELU), essentially **elu** multiplied by a scaling constant.
* **softplus** - Softplus activation function. $log(exp(x) + 1)$ [Introduced](https://papers.nips.cc/paper/1920-incorporating-second-order-functional-knowledge-for-better-option-pricing.pdf) in 2001.
* **softsign** Softsign activation function. $x / (abs(x) + 1)$ Similar to tanh, but not widely used.
* **relu** - Very popular neural network activation function. Used for hidden layers, cannot output negative values. No trainable parameters.
* **tanh** Classic neural network activation function, though often replaced by relu family on modern networks.
* **sigmoid** - Classic neural network activation. Often used on output layer of a binary classifier.
* **hard_sigmoid** - Less computationally expensive variant of sigmoid.
* **exponential** - Exponential (base e) activation function.
* **linear** - Pass through activation function. Usually used on the output layer of a regression neural network.
For more information about Keras activation functions refer to the following:
* [Keras Activation Functions](https://keras.io/activations/)
* [Activation Function Cheat Sheets](https://ml-cheatsheet.readthedocs.io/en/latest/activation_functions.html)
### Advanced Activation Functions
Hyperparameters are not changed when the neural network trains. You, the network designer, must define the hyperparameters. The neural network learns regular parameters during neural network training. Neural network weights are the most common type of regular parameter. The "[advanced activation functions](https://keras.io/layers/advanced-activations/)," as Keras call them, also contain parameters that the network will learn during training. These activation functions may give you better performance than RELU.
* **LeakyReLU** - Leaky version of a Rectified Linear Unit. It allows a small gradient when the unit is not active, controlled by alpha hyperparameter.
* **PReLU** - Parametric Rectified Linear Unit, learns the alpha hyperparameter.
### Regularization: L1, L2, Dropout
* [Keras Regularization](https://keras.io/regularizers/)
* [Keras Dropout](https://keras.io/layers/core/)
### Batch Normalization
* [Keras Batch Normalization](https://keras.io/layers/normalization/)
* Ioffe, S., & Szegedy, C. (2015). [Batch normalization: Accelerating deep network training by reducing internal covariate shift](https://arxiv.org/abs/1502.03167). *arXiv preprint arXiv:1502.03167*.
Normalize the activations of the previous layer at each batch, i.e. applies a transformation that maintains the mean activation close to 0 and the activation standard deviation close to 1. Can allow learning rate to be larger.
### Training Parameters
* [Keras Optimizers](https://keras.io/optimizers/)
* **Batch Size** - Usually small, such as 32 or so.
* **Learning Rate** - Usually small, 1e-3 or so.
### Experimenting with Hyperparameters
```
import pandas as pd
from scipy.stats import zscore
# Read the data set
df = pd.read_csv(
"https://data.heatonresearch.com/data/t81-558/jh-simple-dataset.csv",
na_values=['NA','?'])
# Generate dummies for job
df = pd.concat([df,pd.get_dummies(df['job'],prefix="job")],axis=1)
df.drop('job', axis=1, inplace=True)
# Generate dummies for area
df = pd.concat([df,pd.get_dummies(df['area'],prefix="area")],axis=1)
df.drop('area', axis=1, inplace=True)
# Missing values for income
med = df['income'].median()
df['income'] = df['income'].fillna(med)
# Standardize ranges
df['income'] = zscore(df['income'])
df['aspect'] = zscore(df['aspect'])
df['save_rate'] = zscore(df['save_rate'])
df['age'] = zscore(df['age'])
df['subscriptions'] = zscore(df['subscriptions'])
# Convert to numpy - Classification
x_columns = df.columns.drop('product').drop('id')
x = df[x_columns].values
dummies = pd.get_dummies(df['product']) # Classification
products = dummies.columns
y = dummies.values
import pandas as pd
import os
import numpy as np
import time
import tensorflow.keras.initializers
import statistics
import tensorflow.keras
from sklearn import metrics
from sklearn.model_selection import StratifiedKFold
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Activation, Dropout
from tensorflow.keras import regularizers
from tensorflow.keras.callbacks import EarlyStopping
from sklearn.model_selection import StratifiedShuffleSplit
from tensorflow.keras.layers import LeakyReLU,PReLU
from tensorflow.keras.optimizers import Adam
def evaluate_network(dropout,lr,neuronPct,neuronShrink):
SPLITS = 2
# Bootstrap
boot = StratifiedShuffleSplit(n_splits=SPLITS, test_size=0.1)
# Track progress
mean_benchmark = []
epochs_needed = []
num = 0
neuronCount = int(neuronPct * 5000)
# Loop through samples
for train, test in boot.split(x,df['product']):
start_time = time.time()
num+=1
# Split train and test
x_train = x[train]
y_train = y[train]
x_test = x[test]
y_test = y[test]
# Construct neural network
# kernel_initializer =
# tensorflow.keras.initializers.he_uniform(seed=None)
model = Sequential()
layer = 0
while neuronCount>25 and layer<10:
#print(neuronCount)
if layer==0:
model.add(Dense(neuronCount,
input_dim=x.shape[1],
activation=PReLU()))
else:
model.add(Dense(neuronCount, activation=PReLU()))
model.add(Dropout(dropout))
neuronCount = neuronCount * neuronShrink
model.add(Dense(y.shape[1],activation='softmax')) # Output
model.compile(loss='categorical_crossentropy', optimizer=Adam(lr=lr))
monitor = EarlyStopping(monitor='val_loss', min_delta=1e-3,
patience=100, verbose=0, mode='auto', restore_best_weights=True)
# Train on the bootstrap sample
model.fit(x_train,y_train,validation_data=(x_test,y_test),
callbacks=[monitor],verbose=0,epochs=1000)
epochs = monitor.stopped_epoch
epochs_needed.append(epochs)
# Predict on the out of boot (validation)
pred = model.predict(x_test)
# Measure this bootstrap's log loss
y_compare = np.argmax(y_test,axis=1) # For log loss calculation
score = metrics.log_loss(y_compare, pred)
mean_benchmark.append(score)
m1 = statistics.mean(mean_benchmark)
m2 = statistics.mean(epochs_needed)
mdev = statistics.pstdev(mean_benchmark)
# Record this iteration
time_took = time.time() - start_time
tensorflow.keras.backend.clear_session()
return (-m1)
print(evaluate_network(
dropout=0.2,
lr=1e-3,
neuronPct=0.2,
neuronShrink=0.2))
```
| github_jupyter |
<a href="https://colab.research.google.com/github/kimjinhyuk/kimjinhyuk.github.io/blob/master/2020_06_10_Clustering_Kmeans_algorithm.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# 군집분석
군집 분석은 `비지도학습(Unsupervised learning) 분석 기법` 중 하나 입니다. 쉽게 말해서, 사전 정보 없이 자료를 컴퓨터에게 주고, “유사한 대상끼리 묶어보아라!” 라고 명령을 내리는 분석 방법입니다. 따라서 군집분석에서는 어떤 변수를 컴퓨터에게 입력 하느냐가 중요하다고 볼 수 있습니다.
* ### K-means 군집분석
* K-평균 알고리즘은 구현이 쉽고, 다른 군접 알고리즘에 비해 효율성이 무척 높아 자주 이용되는 알고리즘입니다. 보통 학계나 산업현장에서 사용되는 기법은 다르기 마련이지만, 이 기법은 널리 사용되어집니다.
* 프로토타입은 **연속적인 특성**에서는 비슷한 데이터포인트의 센터로이드(*Centeroid* - 평균) 이거나 **범주형 특성**에서는 메도이드(*Medoid* - 가장 자주 등장하는 포인트)
* 사전에 몇개의 클러스터를 만들것인지에 대해 직접 지정해 줘야 한다는 다소 주관적인 사람의 판단이 개입됩니다. 적절한 K 값을 선택했다면 높은 성능을 발휘 하겠지만 부적합한 K 값을 선택한다면 군집성능에 대한 성능을 보장할 수 없습니다.
* 이해를 돕기위해 무작위 데이터 생성 및 시각화 데이터 그래프를 작성하겠습니다.
```
from sklearn.datasets import make_blobs
```
각각의 속성값으로 어떠한 데이터를 생성할 것인지 확인할 수 있다. 150개, 2차원 3개의클러스터(중심), 표준편차 0.5, 무작위, 시드값은 0
```
X, y = make_blobs(
n_samples=150, n_features=2, centers=3, cluster_std=0.5, shuffle=True, random_state=0
)
import matplotlib.pyplot as plt
```
pyplot 모듈을 사용하여 산정도를 그려줍니다.
```
plt.scatter(X[:,0], X[:,1], c='white', marker='o', edgecolors='black', s=50)
plt.grid()
plt.tight_layout()
plt.show()
```
### 목표 ###
**특성의 유사도**에 기초하여 데이터들을 그룹으로 모으는것
1. 데이터 포인트에서 **랜덤하게 K개**의 센트로이드를 **초기 클러스터 중심으로 선택**
2. 각 데이터를 가장 가까운 센터로이드로 할당
3. 할당된 샘플들을 중심으로 센터로이드를 이동
4. 클러스터 할당이 변하지 않거나, 사용자가 지정한 허용오차나, 최대반복횟수에 도달할 때 까지 2,3 번 반복
최적화를 위해 유클리드 거리(Euclidean distance)를 이용하여 증명할 수 있습니다.
클러스터내의 제곱오차합 SSE 을 반복적으로 최소화
$$ \sum_{i=1}^{n}\sum_{j=1}^{k} w^{(i,j)}d(x^{(i)} ,u^{(j)})^2 $$
* u 는 임의의 클러스터 대표 센터로이드(중심)
* 데이터가 클러스터 내에 있다면 1, 아니면 0 출력
* 유클리드 거리(Euclidean distance) 거리 제곱 방식을 통해서 알고리즘의 2번째 단계를 수행
* 변화량에 대한 허용오차값이 일정 수준내로 들어온다면 더이상 클러스터가 변화하지 않는다는 것이고, 최적화가 완료
```
from sklearn.cluster import KMeans
km = KMeans(n_clusters=3, init='random', n_init=10, max_iter=300, tol=1e04, random_state=0)
y_km = km.fit_predict(X)
plt.scatter(X[y_km==0,0], X[y_km ==0,1 ], c='lightgreen', marker='s', edgecolors='black', s=50, label='cluster1')
plt.scatter(X[y_km==1,0], X[y_km ==1,1 ], c='orange', marker='o', edgecolors='black', s=50, label='cluster2')
plt.scatter(X[y_km==2,0], X[y_km ==2,1 ], c='lightblue', marker='v', edgecolors='black', s=50, label='cluster3')
plt.scatter(km.cluster_centers_[:,0],
km.cluster_centers_[:,1],
c='red',
marker='*',
edgecolors='black',
s=250,
label='centroids')
plt.legend(scatterpoints=1)
plt.grid()
plt.tight_layout()
plt.show()
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
%matplotlib
import warnings
warnings.filterwarnings('ignore')
```
# Imports
```
import pandas as pd
import time
pd.set_option('display.max_columns', 500)
pd.set_option('display.max_colwidth', 150)
pd.set_option('display.max_rows', 500)
from waad.utils.anomalous_asset import ComputeAnomalousAssets
from waad.utils.asset import Account, Asset, IP, Machine
from waad.utils.fait_notable import ComputeFaitNotablesFromIndicators
from waad.utils.indicators import Indicators, ComputeIndicators
from waad.utils.postgreSQL_utils import Database, Table
from waad.utils.rule import Link, Probability, Relation, Rule
```
## 0. Initialisation de la base de données
```
HOST = '127.0.0.1'
PORT = '5432'
USER = '' # To fill
PASSWORD = '' # To fill
DB_NAME = '' # To fill
TABLE_NAME = '' # To fill
db = Database(host=HOST, port=PORT, user=USER, password=PASSWORD, db_name=DB_NAME)
table = Table(db, TABLE_NAME)
```
## 1.1 Définition des règles permettant de compter les bons éléments dans les indicateurs
Les règles sont des objects de la librairie qui définissent des relations entre 2 assets sous certaines conditions. Pour chaque ligne d'authentifications de l'asset source, on vérifie si la règle est appliquée. Elle l'est si l'une des `conditions` au moins est appliquée (`ou` logique). Les conditions sont définies sous la forme de dictionnaires avec une structure de ce type :
```
{
'pre_filters' : {'field_i': <possible values>, 'field_j': <possible values>},
'filter_function': <function(row) -> bool>,
'asset_1': <function(row) -> Asset>,
'asset_2': <function(row) -> Asset>,
}
```
Les conditions dans `pre_filters` et `filter_functions` sont des conditions `et`.
```
rule = Rule(
relation=Relation(link=Link.SE_CONNECTE_SUR, probability=Probability.CERTAIN),
conditions=[
{
'pre_filters': {'eventid': 4624},
'filter_function': lambda row: row['targetusersid'].startswith('S-1-5-21-') and row['host'] != '?',
'asset_1': lambda row: Account(sid=row['targetusersid']),
'asset_2': lambda row: Machine(name=row['host'].split('.')[0], domain=row['host'].split('.')[1]),
}
]
)
```
## 1.2 Calcul des indicateurs à partir de la ``Rule``
```
start = time.time()
ci = ComputeIndicators(table=table, rule=rule, indicator_objects=[Indicators.NB_AUTHENTICATIONS.value, Indicators.NB_ASSETS_REACHED.value, Indicators.NB_NEW_ASSETS_REACHED.value, Indicators.NB_PRIVILEGES_GRANTED.value])
ci.run()
print(time.time() - start)
```
## 1.3 Calcul des FaitsNotables associés
```
cfnfi = ComputeFaitNotablesFromIndicators(ci.indicators)
cfnfi.run()
```
## 2. Calcul des AnomalousAssets
Calcule et ordonne les AnomalousAssets à partir de tous les faits notables
```
caa = ComputeAnomalousAssets(cfnfi.faits_notables)
caa.run()
caa.get_summary().head(30)
for aa in caa.anomalous_assets[:6]:
aa.display()
```
## Exemple d'inputs pour étudier les IPs privées (H1)
```
rule = Rule(
relation=Relation(link=Link.SE_CONNECTE_SUR, probability=Probability.CERTAIN),
conditions=[
{
'pre_filters': {'eventid': 4624},
'filter_function': lambda row: row['ipaddress'] != '?' and row['host'] != '?',
'asset_1': lambda row: IP(row['ipaddress']),
'asset_2': lambda row: Machine(name=row['host'].split('.')[0], domain=row['host'].split('.')[1]),
}
]
)
```
## Exemple d'inputs pour étudier les workstations (H2)
```
rule = Rule(
relation=Relation(link=Link.SE_CONNECTE_SUR, probability=Probability.CERTAIN),
conditions=[
{
'pre_filters': {'eventid': 4624},
'filter_function': lambda row: row['workstationname'] != '?' and row['host'] != '?' and row['workstationname'] != row['host'],
'asset_1': lambda row: Machine(name=row['workstationname']),
'asset_2': lambda row: Machine(name=row['host'].split('.')[0], domain=row['host'].split('.')[1]),
},
]
)
```
## Exemple d'inputs pour étudier les authentifications potentielles des comptes (H7)
```
rule = Rule(
relation=Relation(link=Link.SE_CONNECTE_SUR, probability=Probability.PROBABLE),
conditions=[
{
'pre_filters': {'eventid': 4624},
'filter_function': lambda row: row['targetusersid'].startswith('S-1-5-21-') and row['host'] != '?',
'asset_1': lambda row: Account(sid=row['targetusersid']),
'asset_2': lambda row: Machine(name=row['host'].split('.')[0], domain=row['host'].split('.')[1]),
},
{
'pre_filters': {'eventid': 4672},
'filter_function': lambda row: row['subjectusersid'].startswith('S-1-5-21-') and row['host'] != '?',
'asset_1': lambda row: Account(sid=row['subjectusersid']),
'asset_2': lambda row: Machine(name=row['host'].split('.')[0], domain=row['host'].split('.')[1]),
}
]
)
```
| github_jupyter |
__Objetivos__:
- entender os conceitos de derivada e gradiente
- entender a diferença entre gradiente analítico e numérico
- aprender a calcular a backpropagação de qualquer rede neural.
# Sumário
[0. Imports and Configurações](#0.-Imports-and-Configurações)
[1. Introdução](#1.-Introdução)
- [O Objetivo](#O-Objetivo)
- [Estratégia 1: Busca Aleatória](#Estratégia-1:-Busca-Aleatória)
- [Estratégia 2: Busca Aleatória Local](#Estratégia-2:-Busca-Aleatória-Local)
- [Estratégia 3: Gradiente Numérico](#Estratégia-3:-Gradiente-Numérico)
- [Estratégia 4: Gradiente Analítico](#Estratégia-4:-Gradiente-Anal%C3%ADtico)
- [Caso Recursivo: Múltiplas Portas](#Caso-Recursivo:-Múltiplas-Portas)
- [Checagem do gradiente numérico](#Checagem-do-gradiente-numérico)
- [Neurônio Sigmóide](#Neurônio-Sigmóide)
[2. Backpropagation](#2.-Backpropagation)
- [Se tornando um Ninja em Backpropagation!](#Se-tornando-um-Ninja-em-Backpropagation!)
- [Resumo dos Padrões na Backpropagation](#Resumo-dos-Padrões-na-Backpropagation)
- [Exemplo 1](#Exemplo-1)
- [Exemplo 2](#Exemplo-2)
# 0. Imports and Configurações
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
```
# 1. Introdução
A melhor maneira de pensar em redes neurais é como circuitos de valores reais. Mas, ao invés de valores booleanos, valores reais e, ao invés de portas lógicas como **and** ou **or**, portas binárias (dois operandos) como $*$ (multiplicação), + (adição), max, exp, etc. Além disso, também teremos **gradientes** fluindo pelo circuito, mas na direção oposta.
<img src='images/porta_multiplicacao.png' width="250">
De forma matemática, a gente pode considerar que essa porta implementa a seguinte função:
$$f(x,y)=x*y$$
## O Objetivo
Vamos imaginar que temos o seguinte problema:
1. Nós vamos providenciar a um circuito valores específicos como entrada (x=-2, y=3)
2. O circuito vai calcular o valor de saída (-6)
3. A questão é: *Quanto mudar a entrada para levemente **aumentar** a saída?*
No nosso caso, em que direção devemos mudar x,y para conseguir um número maior que -6? Note que, pro nosso exemplo, se x = -1.99 e y = 2.99, x$*$y = -5.95 que é maior que -6. **-5.95 é melhor (maior) que 6**, e obtivemos uma melhora de 0.05.
## Estratégia 1: Busca Aleatória
Ok. Isso não é trivial? A gente pode simplesmente gerar valores aleatórios, calcular a saída e guardar o melhor resultado.
```
x, y = -2, 3
melhor_saida = forwardMultiplyGate(x,y)
melhor_x, melhor_y = 0, 0
for k in range(0,100):
x_try = 5*np.random.random() - 5
y_try = 5*np.random.random() - 5
out = forwardMultiplyGate(x_try, y_try)
if out > melhor_saida:
melhor_saida = out
melhor_x, melhor_y = x_try, y_try
print(melhor_x, melhor_y, forwardMultiplyGate(melhor_x, melhor_y))
```
Ok, foi bem melhor. Mas, e se tivermos milhões de entradas? É claro que essa estratégia não funcionará. Vamos tentar algo mais aprimorado.
## Estratégia 2: Busca Aleatória Local
```
x, y = -2, 3
passo = 0.01
melhor_saida = forwardMultiplyGate(x,y)
melhor_x, melhor_y = 0, 0
for k in range(0,100):
x_try = x + passo * (2*np.random.random() - 1)
y_try = y + passo * (2*np.random.random() - 1)
out = forwardMultiplyGate(x_try, y_try)
if out > melhor_saida:
melhor_saida = out
melhor_x, melhor_y = x_try, y_try
print(melhor_x, melhor_y, forwardMultiplyGate(melhor_x, melhor_y))
```
## Estratégia 3: Gradiente Numérico
Imagine agora que a gente pega as entradas de um circuito e puxa-as para uma direção positiva. Essa força puxando $x$ e $y$ vai nos dizer como $x$ e $y$ devem mudar para aumentar a saída. Não entendeu? Vamos explicar:
Se olharmos para as entradas, a gente pode intuitivamente ver que a força em $x$ deveria sempre ser positiva, porque tornando $x$ um pouquinho maior de $x=-2$ para $x=-1$ aumenta a saída do circuito para $-3$, o que é bem maior que $-6$. Por outro lado, se a força em $y$ for negativa, tornando-o menor, como de $y=3$ para $y=2$, também aumenta a saída: $-2\times2 = -4$, de novo maior que $-6$.
E como calcular essa força? Usando **derivadas**.
> *A derivada pode ser pensada como a força que a gente aplica em cada entrada para aumentar a saída*
<img src='images/derivada.gif'>
E como exatamente a gente vai fazer isso? Em vez de olhar para o valor de saída, como fizemos anteriormente, nós vamos iterar sobre as cada entrada individualmente, aumentando-as bem devagar e vendo o que acontece com a saída. **A quantidade que a saída muda é a resposta da derivada**.
Vamos para definição matemática. A derivada em relação a $x$ pode ser definida como:
$$\frac{\partial f(x,y)}{\partial x} = \frac{f(x+h,y) - f(x,y)}{h}$$
Onde $h$ é pequeno. Nós vamos, então, calcular a saída inicial $f(x,y)$ e aumentar $x$ por um valor pequeno $h$ e calcular a nova saída $f(x+h,y)$. Então, nós subtraimos esse valores para ver a diferença e dividimos por $f(x+h,y)$ para normalizar essa mudança pelo valor (arbitrário) que nós usamos.
Em termos de código, teremos:
```
x, y = -2, 3
out = forwardMultiplyGate(x,y)
h = 0.0001
# derivada em relação a x
# derivada em relação a y
```
Como a gente pode ver, a derivada em relação a $x$ é igual a $+3$. O sinal positivo indica que alterando o valor de $x$ pelo passo $h$, a saída se torna maior. O valor $3$ pode ser considerado como o valor da força que puxa $x$. O inverso acontece com $y$.
> *A derivada em relação a alguma entrada pode ser calculada ajustando levemente aquela entrada e observando a mudança no valor da saída*
A derivada é calculada sobre cada entrada, enquanto o **gradiente** representa todas as derivadas sobre as entradas concatenadas em um vetor.
Como a gente pode perceber $-5.87 > -6$. Apenas 3 avaliações foram necessárias para aumentar o valor da saída (ao invés de centenas) e conseguimos um melhor resultado.
**Passo maior nem sempre é melhor**: É importante destacar que qualquer valor de passo maior que 0.01 ia sempre funcionar melhor (por exemplo, passo = 1 gera a saída = 1). No entanto, à medida que os circuitos vão ficando mais complexos (como em redes neurais completas), a função vai ser tornando mais caótica e complexa. O gradiente garante que se você tem um passo muito pequeno (o ideal seria infinitesimal), então você definitivamente aumenta a saída seguindo aquela direção. O passo que estamos utilizando (0.01) ainda é muito grande, mas como nosso circuito é simples, podemos esperar pelo melhor resultado. Lembre-se da analogia do **escalador cego**.
## Estratégia 4: Gradiente Analítico
A estratégia que utilizamos até agora de ajustar levemente a entrada e ver o que acontece com a saída pode não ser muito cômoda na prática quando temos milhares de entradas para ajustar. Então, a gente precisa de algo melhor.
Felizmente, existe uma estratégia mais fácil e muito mais rápida para calcular o gradiente: podemos usar cálculo para derivar diretamente a nossa função. Chamamos isso de **gradiente analítico** e dessa forma não precisamos ajustar levemente nada.
> *O gradiente analítico evita o leve ajustamento das entradas. O circuito pode ser derivado usando cálculo.*
É muito fácil calcular derivadas parciais para funções simples como $x*y$. Se você não lembra da definição, aqui está o cálculo da derivada parcial em relação a $x$ da nossa função $f(x,y)$:
$$\frac{\partial f(x,y)}{\partial x} = \frac{f(x+h,y) - f(x,y)}{h}
= \frac{(x+h)y - xy}{h}
= \frac{xy + hy - xy}{h}
= \frac{hy}{h}
= y$$
A derivada parcial em relação em $x$ da nossa $f(x,y)$ é igual $y$. Você reparou na coincidência de $\partial x = 3.0$, que é exatamente o valor de $y$? E que o mesmo aconteceu para $x$? **Então, a gente não precisa ajustar nada!** E nosso código fica assim:
```
x, y = -2, 3
out = forwardMultiplyGate(x,y)
# insira seu código aqui!
```
É importante destacar que a Estratégia #3 reduziu a #2 para uma única vez. Porém, a #3 nos dá somente uma aproximação do gradiente, enquanto a Estratégia #4 nos dá o valor exato. Sem aproximações. O único lado negativo é que temos de saber derivar a nossa funcão.
Recapitulando o que vimos até aqui:
- __Estratégia 1__: definimos valores aleatórios em todas as iterações. Não funciona para muitas entradas.
- __Estratégia 2__: pequenos ajustes aleatórios nas entradas e vemos qual funciona melhor. Tão ruim quando a #1.
- __Estratégia 3__: muito melhor através do cálculo do gradiente. Independentemente de quão complicado é o circuito, o **gradiente numérico** é muito simples de se calcular (mas um pouco caro).
- __Estratégia 4__: no final, vimos que a forma melhor, mais inteligente e mais rápida é calcular o **gradiente analítico**. O resultado é idêntico ao gradiente numérico, porém mais rápido e não precisa de ajustes.
## Caso Recursivo: Múltiplas Portas
Calcular o gradiente para o nosso circuito foi trivial. Mas, e em circuitos mais complexos? Como a gente vai ver agora, cada porta pode ser tratada individualmente e a gente pode calcular derivadas locais como a gente fez anteriormente. Vamos considerar nossa função agora como a seguinte:
$$f(x,y,z) = (x+y)*z$$
<img src='images/circuito_2.png' width='300'>
Como vamos calcular agora a nossa derivada? Primeiramente, vamos esquecer da porta de soma e fingir que temos apenas duas entradas no nosso circuito: **q** e **z**. Como já vimos, as nossas derivadas parciais podem ser definidas da seguinte maneira:
$$f(q,z) = q z \hspace{0.5in} \implies \hspace{0.5in} \frac{\partial f(q,z)}{\partial q} = z, \hspace{1in} \frac{\partial f(q,z)}{\partial z} = q$$
Ok, mas e em relação a $x$ e $y$? Como $q$ é calculado em função de $x$ e $y$ (pela adição em nosso exemplo), nós também podemos calcular suas derivadas parciais:
$$q(x,y) = x + y \hspace{0.5in} \implies \hspace{0.5in} \frac{\partial q(x,y)}{\partial x} = 1, \hspace{1in} \frac{\partial q(x,y)}{\partial y} = 1$$
Correto! As derivadas parciais são 1, independentemente dos valores de $x$ e $y$. Isso faz sentido se pensarmos que para aumentar A saída de uma porta de adição, a gente espera uma força positiva tanto em $x$ quanto em $y$, independente dos seus valores.
Com as fórmulas acima, nós sabemos calcular o gradiente da saída em relação a $q$ e $z$, e o gradiente de $q$ em relação a $x$ e $y$. Para calcular o gradiente do nosso circuito em relação a $x$, $y$ e $z$, nós vamos utilizar a **Regra da Cadeia**, que vai nos dizer como combinar esses gradientes. A derivada final em relação a $x$, será dada por:
$$\frac{\partial f(q,z)}{\partial x} = \frac{\partial q(x,y)}{\partial x} \frac{\partial f(q,z)}{\partial q}$$
Pode parecer complicado à primeira vista, mas a verdade é que isso vai ser simplificado a somente duas multiplicações:
```
# Derivada da porta de multiplicação
# Derivada da porta de adição
# Regra da cadeia
```
<img src="images/circuito_2_back.png">
É isso! Vamos agora fazer nossas entradas responderem ao gradiente. Lembrando que queremos um valor maior que -12.
Vamos agora analisar os resultados separadamente. Analisando primeiramente $q$ e $z$, vemos que o circuito quer que $z$ aumente (der_f_rel_z = +3) e o valor de $q$ diminua (der_f_rel_q = -4) com uma força maior (4 contra 3).
Em relação a porta de soma, como vimos, o padrão é que aumentando as entradas a saída também aumenta. Porém, o circuito quer que $q$ diminua (der_f_rel_q = -4). Esse é o **ponto crucial**: em vez de aplicarmos uma força de +1 as entradas da porta de soma como normalmente faríamos (derivada local), o circuito quer que os gradientes em $x$ e $y$ se tornem 1x-4=-4. Isso faz sentido: o circuito quer $x$ e $y$ pequeno para que $q$ seja pequeno também, o que vai aumentar $f$.
> *Se isso fez sentido, você entendeu backpropagation.*
**Recapitulando:**
- Vimos que, para uma simples porta (or simples expressão), podemos derivar o gradiente analítico usando cálculo simples. Nós interpretamos o gradiente como uma força que puxa as entradas na direção necessária para fazer a saída aumentar.
- No caso de múltiplas portas, cada porta é tratada individualmente até que o circuito seja tratado como um todo. A *única* diferença é que agora o circuito diz como a saída de outras portas devem se comportar (como da porta de adição), que é o gradiente final do circuito em relação a saída da porta. É como o circuito pedindo aquela porta maior ou menor valor de saída, e com alguma força. A porta simplesmente pega essa força e multiplica em relação a todas as forças calculadas para suas entradas anteriores (regra da cadeia) - repare como a força de q (-4) é multiplicada as forças de x e y. Isso pode ter dois efeitos desejados:
- Se a porta tem uma força positiva de saída, essa força também é multiplicada nas suas entradas, escalonada pelo valor da força das entradas.
- Se a porta tem uma força negativa de saída, isso significa que o circuito quer que a saída decresça, então essa força é multiplicada pelas entradas para diminuir o valor de saída.
> *Tenha em mente que a força da saída do circuito vai puxando as outras forças na direção desejada por todo o circuito até as entradas.*
## Checagem do gradiente numérico
Vamos verificar se os gradientes analíticos que calculamos por backpropagation estão corretos. Lembre-se que podemos fazer isso através do gradiente numérico e esperamos que o resultado seja [-4, -4, 4] para $x,y,z$.
```
x,y,z = -2,5,-4
h = 0.0001
#insira seu código aqui
```
## Neurônio Sigmóide
Qualquer função diferenciável pode atuar como uma porta, como também podemos agrupar múltiplas portas para formar uma simples porta, ou decompor um função em múltiplas portas quando for conveniente. Para exemplificar, vamos utilizar a função de ativação *sigmoid* com entradas **x** e pesos **w**:
$$f(w,x) = \frac{1}{1+e^{-(w_0x_0 + w_1x_1 + w_2)}}$$
Como dito, a função acima nada mais é que a função sigmoid $\sigma(x)$. Sabendo, então, que a derivada da função sigmoid é:
$$\sigma(x)=\frac{1}{1+e^{-x}}=(1-\sigma(x))\sigma(x)$$
Vamos calcular a gradiente em relação as entradas:
```
w0, w1, w2 = 2, -3, -3
x0, x1 = -1, -2
# forward pass
# backward pass
# Nova saida
```
Vamos supor agora que não sabemos a derivada da função $\sigma(x)$ muito menos de $f(w,x)$. O que podemos fazer?.
**Decompor essa função em circuito com múltiplas portas!** Dessa forma:
<img src='images/circuito_3.png' width='800'>
Calculando a saída para cada porta, temos:
<img src='images/circuito_3_forward.png' width='800'>
Onde sabemos as seguintes derivadas:
$$f(x) = \frac{1}{x} \rightarrow \frac{df}{dx} = -1/x^2
\\\\
f_c(x) = c + x \rightarrow \frac{df}{dx} = 1
\\\\
f(x) = e^x \rightarrow \frac{df}{dx} = e^x
\\\\
f_a(x) = ax \rightarrow \frac{df}{dx} = a$$
Onde as funções $f_c(x)$ e $f_a(x)$ transladam a entrada por uma constante $c$ e escala por uma contante $a$, respectivamente. Na verdade, são apenas casos especias de adição e multiplicação, mas que foram introduzidos como portas unárias.
Como podemos calcular a derivada em relação as entradas agora? **Usando Backpropagation!!**
# 2. Backpropagation
## Se tornando um Ninja em Backpropagation!
Antes de resolver o circuito acima, vamos praticar um pouco de backpropagation com alguns exemplos. Vamos esquecer funções por enquanto e trabalhar só com 4 variáveis: $a$, $b$, $c$, e $x$. Vamos também nos referir as seus gradientes como $da$, $db$, $dc$, e $dx$. Além disso, vamos assumir que $dx$ é dado (ou é +1 como nos casos acima). Nosso primeiro exemplo é a porta $*$, que já conhecemos:
$$x = a * b$$
$$da = b * dx$$
$$db = a * dx$$
Se você reparar bem, vai perceber que a porta $*$ atua como um *switcher* durante a backpropagation, ou seja, o gradiente de cada entrada é o valor da outra multiplicado pelo gradiente da anterior (regra da cadeia). Por outro lado, vamos analisar a porta +:
$$x = a + b$$
$$da = 1.0 * dx$$
$$db = 1.0 * dx$$
Nesse caso, 1.0 é o gradiente local e a multiplicação é a nossa regra da cadeia. **E se fosse a adição de 3 números?**:
$$q = a + b$$
$$x = q + c$$
$$dc = 1.0 * dx$$
$$dq = 1.0 * dx$$
$$da = 1.0 * dq$$
$$db = 1.0 * dq$$
Você percebe o que está acontecendo? Se você olhar nos diagramas dos circuitos que já resolvemos, vai perceber que a porta + simplesmente pega o gradiente atual e roteia igualmente para todas as entradas (porque os gradientes locais são sempre 1.0 para todas as entradas, independente dos seus valores atuais). Então, podemos fazer bem mais rápido:
$$x = a + b + c$$
$$da = 1.0 * dx$$
$$db = 1.0 * dx$$
$$dc = 1.0 * dx$$
Okay. Mas, e se combinarmos portas?
$$x = a*b + c$$
$$da = b * dx$$
$$db = a * dx$$
$$dc = 1.0 * dx$$
Se você não percebeu o que aconteceu, introduza uma variável temporária $q = a * b$ e então calcula $x = q + c$ para se convencer. E quanto a este exemplo:
$$x = a * a$$
$$da = 2 * a * dx$$
Outro exemplo:
$$x = a*a + b*b + c*c$$
$$da = 2 * a * dx$$
$$db = 2 * b * dx$$
$$dc = 2 * c * dx$$
Ok. Agora mais complexo:
$$x = (a * b + c) * d)^2$$
Quando casos mais complexos como esse acontecem, eu gosto de dividir a expressão em partes gerenciáveis que são quase sempre compostas de simples expressões onde eu posso aplicar a regra da cadeia:
$$x1 = a * b + c$$
$$x2 = x1 * d$$
$$x = x2 * x2$$
$$dx2 = 2 * x2 * dx$$
$$dx1 = d * dx2$$
$$dd = x1 * dx2$$
$$da = b * dx1$$
$$db = a * dx1$$
$$dc = 1 * dx1$$
Não foi tão difícil! Essas são as equações para toda a expressão, e nós fizemos dividindo peça por peça e aplicando backpropagation a todas as variáveis. Note que **toda variável durante a fase forward tem uma variável equivalente na backpropagação que contém o gradiente em relação a saída do circuito.**. Mais um exemplo útil de função e seu gradiente local:
$$x = 1.0/a$$
$$da = 1.0/(a*a) * dx$$
E como ela pode ser aplicada na prática:
$$x = (a+b)/(c+d)$$
$$x1 = a + b$$
$$x2 = c + d$$
$$x3 = 1.0 / x2$$
$$x = x1 * x3$$
$$dx1 = x3 * dx$$
$$dx3 = x1 * dx$$
$$dx2 = (1.0/(x2 * x2)) * dx3$$
$$dc = 1 * dx2$$
$$dd = 1 * dx2$$
$$da = 1 * dx1$$
$$db = 1 * dx1$$
E mais uma:
$$x = math.max(a, b)$$
$$da = x == a\ ?\ 1.0 * dx\ :\ 0.0$$
$$db = x == b\ ?\ 1.0 * dx\ :\ 0.0$$
No caso acima é mais difícil de entender. A função **max** passa o valor para a maior entrada e ignora as outras. Na fase de backpropagation, a porta __max__ simplesmente pega o gradiente atual e roteia para a entrada que teve o maior valor durante a fase de forward. A porta age como um simples switch baseado na entrada com o maior valor durante a forward. As outras entradas terão gradiente zero.
Agora, vamos dar uma olhada na porta **ReLU (*Rectified Linear Unit)***, muita usada em redes neurais no lugar da função sigmoid. Ela é simplesmente um threshold com zero:
$$x = max(a, 0)$$
$$da = a > 0\ ?\ 1.0 * dx\ :\ 0.0$$
Em outras palavras, essa porta simplesmente passa o valor adiante se ele é maior que zero, ou interrompe o fluxo e seta o valor para zero. Na backpropagação, a porta vai passar o gradiente atual se ele foi ativado durante a forward. Se a entrada original foi menor que zero, ela vai interromper o fluxo de gradiente.
Finalmente, vamos ver como calcular o gradiente em operações vetorizadas que vamos utilizar muito em redes neurais:
$$W = np.random.randn(5,10)$$
$$X = np.random.randn(3,10)$$
$$Y = X.dot(W^T)$$
Supondo que o gradiente de Y é dado como a seguir:
$$dY = np.random.randn(*Y.shape)$$
$$dW = dY^T.dot(X)$$
$$dX = dY.dot(W)$$
Espero que tenha entendido como calcular expressões inteiras (que são feitas de muitas portas) e como calcular a backpropagação para cada uma delas.
## Resumo dos Padrões na Backpropagation
Para resumir os padrões no fluxo da backpropagation considere esse circuito:
<img src='images/backpropagation_padroes.png' width='450'>
A **porta de soma** simplesmente pega o gradiente na saída e distribui igualmente para entrada, independente dos valores durante a etapa de forward. Isso vem do fato que o gradiente local para a operação de adicionar é simplesmente +1.0, então os gradientes em todas as entradas vão ser exatamente iguais ao gradiente da saída porque ele vai ser multiplicado por 1.0 (e continua o mesmo). No circuito acima, repare como a porta + roteou o gradiente 2.0 para ambas as entradas, igualmente e sem alteração.
A **porta max** roteia o gradiente. Diferente da porta de soma que distribui o gradiente para todas as entradas, distribui o gradiente (sem alteração) para exatamente uma das entradas (a que tinha o maior valor durante a etapa de forward). Isso acontece por que o gradiente local é 1.0 para o maior valor e 0.0 para os outros valores. No circuito acima, a operação max roteou o gradiente de 2.0 para a variável $z$, que tinha um valor maior que $w$, e o gradiente de $w$ continua zero.
A **porta de multiplicação** é um pouquinho mais difícil de interpretar. Os gradientes locais são os valores das entradas (cambiados) e multiplicados pelo gradiente da saída durante a regra da cadeia. No exemplo acima, o gradiente em $x$ é -8.00, pois é igual a -4.00x2.00.
*Efeitos não inutuitivos e suas consequências*. Note que se uma das entradas na porta de multiplicação é muito pequena e a outra é muito grande, então a porta de multiplicação vai fazer algo intuitivo: ela vai atribuir um gradiente muito alto para a menor entrada e um muito pequeno para a maior entrada. Perceba que no caso de classificadores lineares, onde os pesos são multiplicados com as entradas $w^Tx_i$, isso implica que a escala dos dados tem um efeito na magnitude do gradiente para os pesos. Por exemplo, se você multiplicar todos os dados de entrada **$x_i$** por 1000 durante pré-processamento, então o gradiente dos pesos vão ser 1000x maior, e você terá de usar baixas taxas de aprendizagem para compensar o fator. Por isso que o pré-processamento é tão importante e o conhecimento intuitivo sobre os gradientes podem ajudar a debugar alguns desses casos.
## Exemplo 1
Implementando o nosso neurônio
<img src='images/circuito_3_back.png' width='800'>
```
w0, w1, w2 = 2, -3, -3
x0, x1 = -1, -2
# forward pass
# backward pass
```
## Exemplo 2
Vamos ver outro exemplo. Suponha que temos a seguinte função:
$$f(x,y) = \frac{x + \sigma(y)}{\sigma(x) + (x+y)^2}$$
Só para deixar claro, essa função é completamente inútil, mas um bom exemplo de backpropagation na prática. Também é importante destacar que ela é bem difícil de derivar em relação a $x$ e $y$. No entanto, como vimos, saber derivar uma função é completamente desnecessário por que não precisamos saber derivar a função inteira para calcular os gradientes. Só precisamos saber como calcular os gradientes locais. Aqui está a resolução:
```
x, y = 3, -4
# forward pass
# backward pass
```
Repare em algumas coisas importantes:
**Variáveis temporárias para armazenar resultados**. Para calcular a backpropagation, é importante ter algumas (se não todas) das variáveis calculadas na etapa de forward. Na prática, é bom estruturar seu código de maneira a guardar esses valores para a backprop. Em último caso, você pode recalculá-las.
**Gradientes adicionados**. A etapa de forward envolveu as variáveis $x$ e $y$ muitas vezes, então quando fazemos a backprop temos de ter cuidados de acumular o gradiente nessas variáveis (+=). Isso segue a **regra da cadeia multivariável** em cálculo.
# Referências
1. [CS231n - Optimization: Stochastic Gradient Descent](http://cs231n.github.io/optimization-1/)
2. [CS231n - Backpropagation, Intuitions](http://cs231n.github.io/optimization-2/)
3. [Hacker's guide to Neural Networks](http://karpathy.github.io/neuralnets/)
| github_jupyter |
```
import numpy as np
import xarray as xr
import pandas as pd
import geoviews as gv
import geoviews.feature as gf
from geoviews import dim, opts
gv.extension('bokeh')
```
The Bokeh backend offers much more advanced tools to interactively explore data, making good use of GeoViews support for web mapping tile sources. As we learned in the [Projections](Projections.ipynb) user guide, using web mapping tile sources is only supported when using the default ``GOOGLE_MERCATOR`` ``crs``.
# WMTS - Tile Sources
GeoViews provides a number of tile sources by default, provided by CartoDB, Stamen, OpenStreetMap, Esri and Wikipedia. These can be imported from the ``geoviews.tile_sources`` module.
```
import geoviews.tile_sources as gts
gv.Layout([ts.relabel(name) for name, ts in gts.tile_sources.items()]).opts(
'WMTS', xaxis=None, yaxis=None, width=225, height=225).cols(4)
```
The tile sources that are defined as part of GeoViews are simply instances of the ``gv.WMTS`` and ``gv.Tiles`` elements, which accept tile source URLs of three formats:
1. Web mapping tile sources: ``{X}``, ``{Y}`` defining the location and a ``{Z}`` parameter defining the zoom level
2. Bounding box tile source: ``{XMIN}``, ``{XMAX}``, ``{YMIN}``, and ``{YMAX}`` parameters defining the bounds
3. Quad-key tile source: a single ``{Q}`` parameter
Additional, freely available tile sources can be found at [wiki.openstreetmap.org](http://wiki.openstreetmap.org/wiki/Tile_servers).
A tile source may also be drawn at a different ``level`` allowing us to overlay a regular tile source with a set of labels. Valid options for the 'level' option include 'image', 'underlay', 'glyph', 'annotation' and 'overlay':
```
gts.EsriImagery.opts(width=600, height=570, global_extent=True) * gts.StamenLabels.options(level='annotation')
```
## Plotting data
One of the main benefits of plotting data with Bokeh is the interactivity it allows. Here we will load a dataset of all the major cities in the world with their population counts over time:
```
cities = pd.read_csv('../assets/cities.csv', encoding="ISO-8859-1")
population = gv.Dataset(cities, kdims=['City', 'Country', 'Year'])
cities.head()
```
Now we can convert this dataset to a set of points mapped by the latitude and longitude and containing the population, country and city as values. The longitudes and latitudes in the dataframe are supplied in simple Plate Carree coordinates, which we will need to declare (as the values are not stored with any associated units). The ``.to`` conversion interface lets us do this succinctly. Note that since we did not assign the Year dimension to the points key or value dimensions, it is automatically assigned to a HoloMap, rendering the data as an animation using a slider widget:
```
points = population.to(gv.Points, ['Longitude', 'Latitude'], ['Population', 'City', 'Country'])
(gts.Wikipedia * points).opts(
opts.Points(width=600, height=350, tools=['hover'], size=np.sqrt(dim('Population'))*0.005,
color='Population', cmap='viridis'))
```
And because this is a fully interactive Bokeh plot, you can now hover over each datapoint to see all of the values associated with it (name, location, etc.), and you can zoom and pan using the tools provided. Each time, the map tiles should seamlessly update to provide additional detail appropriate for that zoom level.
## Choropleths
The tutorial on [Geometries](Geometries.ipynb) covers working with shapefiles in more detail but here we will quickly combine a shapefile with a pandas DataFrame to plot the results of the EU Referendum in the UK. We begin by loading the shapefile and then us ``pd.merge`` by combining it with some CSV data containing the referendum results:
```
import geopandas as gpd
geometries = gpd.read_file('../assets/boundaries/boundaries.shp')
referendum = pd.read_csv('../assets/referendum.csv')
gdf = gpd.GeoDataFrame(pd.merge(geometries, referendum))
```
Now we can easily pass the GeoDataFrame to a Polygons object and declare the ``leaveVoteshare`` as the first value dimension which it will color by:
```
gv.Polygons(gdf, vdims=['name', 'leaveVoteshare']).opts(
tools=['hover'], width=450, height=600, color_index='leaveVoteshare',
colorbar=True, toolbar='above', xaxis=None, yaxis=None)
```
### Images
The Bokeh backend also provides basic support for working with images. In this example we will load a very simple Iris Cube and display it overlaid with the coastlines feature from Cartopy. Note that the Bokeh backend does not project the image directly into the web Mercator projection, instead relying on regridding, i.e. resampling the data using a new grid. This means the actual display may be subtly different from the more powerful image support for the matplotlib backend, which will project each of the pixels into the chosen display coordinate system without regridding.
```
dataset = xr.open_dataset('../data/pre-industrial.nc')
air_temperature = gv.Dataset(dataset, ['longitude', 'latitude'], 'air_temperature',
group='Pre-industrial air temperature')
air_temperature.to.image().opts(tools=['hover'], cmap='viridis') *\
gf.coastline().opts(line_color='black', width=600, height=500)
```
| github_jupyter |
## MNIST in Keras with Tensorboard
This sample trains an "MNIST" handwritten digit
recognition model on a GPU or TPU backend using a Keras
model. Data are handled using the tf.data.Datset API. This is
a very simple sample provided for educational purposes. Do
not expect outstanding TPU performance on a dataset as
small as MNIST.
### Parameters
```
BATCH_SIZE = 64
LEARNING_RATE = 0.02
# GCS bucket for training logs and for saving the trained model
# You can leave this empty for local saving, unless you are using a TPU.
# TPUs do not have access to your local instance and can only write to GCS.
BUCKET="" # a valid bucket name must start with gs://
training_images_file = 'gs://mnist-public/train-images-idx3-ubyte'
training_labels_file = 'gs://mnist-public/train-labels-idx1-ubyte'
validation_images_file = 'gs://mnist-public/t10k-images-idx3-ubyte'
validation_labels_file = 'gs://mnist-public/t10k-labels-idx1-ubyte'
```
### Imports
```
import os, re, math, json, time
import PIL.Image, PIL.ImageFont, PIL.ImageDraw
import numpy as np
import tensorflow as tf
from matplotlib import pyplot as plt
from tensorflow.python.platform import tf_logging
print("Tensorflow version " + tf.__version__)
```
## TPU/GPU detection
```
tpu = None
try:
tpu = tf.distribute.cluster_resolver.TPUClusterResolver() # TPU detection relies on TPU_NAME env var
tf.tpu.experimental.initialize_tpu_system(tpu)
strategy = tf.distribute.experimental.TPUStrategy(tpu, steps_per_run=100)
print('Running on TPU ', tpu.cluster_spec().as_dict()['worker'])
except ValueError:
gpus = tf.config.experimental.list_logical_devices("GPU")
if len(gpus) > 1:
strategy = tf.distribute.MirroredStrategy([gpu.name for gpu in gpus])
print("running on multiple GPUs")
else:
strategy = tf.distribute.get_strategy() # the default strategy works on CPU and single GPU
print("Running on {}".format("a single GPU" if len(gpus)==1 else "CPU"))
# adjust batch size and learning rate for distributed computing
global_batch_size = BATCH_SIZE * strategy.num_replicas_in_sync # num replcas is 8 on a single TPU or N when runing on N GPUs.
learning_rate = LEARNING_RATE * strategy.num_replicas_in_sync
#@title visualization utilities [RUN ME]
"""
This cell contains helper functions used for visualization
and downloads only. You can skip reading it. There is very
little useful Keras/Tensorflow code here.
"""
# Matplotlib config
plt.rc('image', cmap='gray_r')
plt.rc('grid', linewidth=0)
plt.rc('xtick', top=False, bottom=False, labelsize='large')
plt.rc('ytick', left=False, right=False, labelsize='large')
plt.rc('axes', facecolor='F8F8F8', titlesize="large", edgecolor='white')
plt.rc('text', color='a8151a')
plt.rc('figure', facecolor='F0F0F0')# Matplotlib fonts
MATPLOTLIB_FONT_DIR = os.path.join(os.path.dirname(plt.__file__), "mpl-data/fonts/ttf")
# pull a batch from the datasets. This code is not very nice, it gets much better in eager mode (TODO)
def dataset_to_numpy_util(training_dataset, validation_dataset, N):
# get one batch from each: 10000 validation digits, N training digits
unbatched_train_ds = training_dataset.apply(tf.data.experimental.unbatch())
if tf.executing_eagerly():
# This is the TF 2.0 "eager execution" way of iterating through a tf.data.Dataset
for v_images, v_labels in validation_dataset:
break
for t_images, t_labels in unbatched_train_ds.batch(N):
break
validation_digits = v_images.numpy()
validation_labels = v_labels.numpy()
training_digits = t_images.numpy()
training_labels = t_labels.numpy()
else:
# This is the legacy TF 1.x way of iterating through a tf.data.Dataset
v_images, v_labels = validation_dataset.make_one_shot_iterator().get_next()
t_images, t_labels = unbatched_train_ds.batch(N).make_one_shot_iterator().get_next()
# Run once, get one batch. Session.run returns numpy results
with tf.Session() as ses:
(validation_digits, validation_labels,
training_digits, training_labels) = ses.run([v_images, v_labels, t_images, t_labels])
# these were one-hot encoded in the dataset
validation_labels = np.argmax(validation_labels, axis=1)
training_labels = np.argmax(training_labels, axis=1)
return (training_digits, training_labels,
validation_digits, validation_labels)
# create digits from local fonts for testing
def create_digits_from_local_fonts(n):
font_labels = []
img = PIL.Image.new('LA', (28*n, 28), color = (0,255)) # format 'LA': black in channel 0, alpha in channel 1
font1 = PIL.ImageFont.truetype(os.path.join(MATPLOTLIB_FONT_DIR, 'DejaVuSansMono-Oblique.ttf'), 25)
font2 = PIL.ImageFont.truetype(os.path.join(MATPLOTLIB_FONT_DIR, 'STIXGeneral.ttf'), 25)
d = PIL.ImageDraw.Draw(img)
for i in range(n):
font_labels.append(i%10)
d.text((7+i*28,0 if i<10 else -4), str(i%10), fill=(255,255), font=font1 if i<10 else font2)
font_digits = np.array(img.getdata(), np.float32)[:,0] / 255.0 # black in channel 0, alpha in channel 1 (discarded)
font_digits = np.reshape(np.stack(np.split(np.reshape(font_digits, [28, 28*n]), n, axis=1), axis=0), [n, 28*28])
return font_digits, font_labels
# utility to display a row of digits with their predictions
def display_digits(digits, predictions, labels, title, n):
plt.figure(figsize=(13,3))
digits = np.reshape(digits, [n, 28, 28])
digits = np.swapaxes(digits, 0, 1)
digits = np.reshape(digits, [28, 28*n])
plt.yticks([])
plt.xticks([28*x+14 for x in range(n)], predictions)
for i,t in enumerate(plt.gca().xaxis.get_ticklabels()):
if predictions[i] != labels[i]: t.set_color('red') # bad predictions in red
plt.imshow(digits)
plt.grid(None)
plt.title(title)
# utility to display multiple rows of digits, sorted by unrecognized/recognized status
def display_top_unrecognized(digits, predictions, labels, n, lines):
idx = np.argsort(predictions==labels) # sort order: unrecognized first
for i in range(lines):
display_digits(digits[idx][i*n:(i+1)*n], predictions[idx][i*n:(i+1)*n], labels[idx][i*n:(i+1)*n],
"{} sample validation digits out of {} with bad predictions in red and sorted first".format(n*lines, len(digits)) if i==0 else "", n)
```
### Colab-only auth for this notebook and the TPU
```
#IS_COLAB_BACKEND = 'COLAB_GPU' in os.environ # this is always set on Colab, the value is 0 or 1 depending on GPU presence
#if IS_COLAB_BACKEND:
# from google.colab import auth
# auth.authenticate_user() # Authenticates the backend and also the TPU using your credentials so that they can access your private GCS buckets
```
### tf.data.Dataset: parse files and prepare training and validation datasets
Please read the [best practices for building](https://www.tensorflow.org/guide/performance/datasets) input pipelines with tf.data.Dataset
```
def read_label(tf_bytestring):
label = tf.io.decode_raw(tf_bytestring, tf.uint8)
label = tf.reshape(label, [])
label = tf.one_hot(label, 10)
return label
def read_image(tf_bytestring):
image = tf.io.decode_raw(tf_bytestring, tf.uint8)
image = tf.cast(image, tf.float32)/256.0
image = tf.reshape(image, [28*28])
return image
def load_dataset(image_file, label_file):
imagedataset = tf.data.FixedLengthRecordDataset(image_file, 28*28, header_bytes=16)
imagedataset = imagedataset.map(read_image, num_parallel_calls=16)
labelsdataset = tf.data.FixedLengthRecordDataset(label_file, 1, header_bytes=8)
labelsdataset = labelsdataset.map(read_label, num_parallel_calls=16)
dataset = tf.data.Dataset.zip((imagedataset, labelsdataset))
return dataset
def get_training_dataset(image_file, label_file, batch_size):
dataset = load_dataset(image_file, label_file)
dataset = dataset.cache() # this small dataset can be entirely cached in RAM, for TPU this is important to get good performance from such a small dataset
dataset = dataset.shuffle(5000, reshuffle_each_iteration=True)
dataset = dataset.repeat() # Mandatory for Keras for now
dataset = dataset.batch(batch_size, drop_remainder=True) # drop_remainder is important on TPU, batch size must be fixed
dataset = dataset.prefetch(-1) # fetch next batches while training on the current one (-1: autotune prefetch buffer size)
return dataset
def get_validation_dataset(image_file, label_file):
dataset = load_dataset(image_file, label_file)
dataset = dataset.cache() # this small dataset can be entirely cached in RAM, for TPU this is important to get good performance from such a small dataset
dataset = dataset.repeat() # Mandatory for Keras for now
dataset = dataset.batch(10000, drop_remainder=True) # 10000 items in eval dataset, all in one batch
return dataset
# instantiate the datasets
training_dataset = get_training_dataset(training_images_file, training_labels_file, global_batch_size)
validation_dataset = get_validation_dataset(validation_images_file, validation_labels_file)
```
### Let's have a look at the data
```
N = 24
(training_digits, training_labels,
validation_digits, validation_labels) = dataset_to_numpy_util(training_dataset, validation_dataset, N)
display_digits(training_digits, training_labels, training_labels, "training digits and their labels", N)
display_digits(validation_digits[:N], validation_labels[:N], validation_labels[:N], "validation digits and their labels", N)
font_digits, font_labels = create_digits_from_local_fonts(N)
```
### Keras model: 3 convolutional layers, 2 dense layers
```
# This model trains to 99.4%— sometimes 99.5%— accuracy in 10 epochs (with a batch size of 64)
def make_model():
model = tf.keras.Sequential(
[
tf.keras.layers.Reshape(input_shape=(28*28,), target_shape=(28, 28, 1)),
tf.keras.layers.Conv2D(filters=6, kernel_size=3, padding='same', use_bias=False), # no bias necessary before batch norm
tf.keras.layers.BatchNormalization(scale=False, center=True), # no batch norm scaling necessary before "relu"
tf.keras.layers.Activation('relu'), # activation after batch norm
tf.keras.layers.Conv2D(filters=12, kernel_size=6, padding='same', use_bias=False, strides=2),
tf.keras.layers.BatchNormalization(scale=False, center=True),
tf.keras.layers.Activation('relu'),
tf.keras.layers.Conv2D(filters=24, kernel_size=6, padding='same', use_bias=False, strides=2),
tf.keras.layers.BatchNormalization(scale=False, center=True),
tf.keras.layers.Activation('relu'),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(200, use_bias=False),
tf.keras.layers.BatchNormalization(scale=False, center=True),
tf.keras.layers.Activation('relu'),
tf.keras.layers.Dropout(0.5), # Dropout on dense layer only
tf.keras.layers.Dense(10, activation='softmax')
])
model.compile(optimizer='adam', # learning rate will be set by LearningRateScheduler
loss='categorical_crossentropy',
metrics=['accuracy'])
return model
with strategy.scope(): # the new way of handling distribution strategies in Tensorflow 1.14+
model = make_model()
# print model layers
model.summary()
# set up learning rate decay
lr_decay = tf.keras.callbacks.LearningRateScheduler(lambda epoch: learning_rate * math.pow(0.5, 1+epoch) + learning_rate/200, verbose=True)
# set up Tensorboard logs
timestamp = time.strftime("%Y-%m-%d-%H-%M-%S")
log_dir=os.path.join(BUCKET, 'mnist-logs', timestamp)
tb_callback = tf.keras.callbacks.TensorBoard(log_dir=log_dir, update_freq=50*global_batch_size)
print("Tensorboard loggs written to: ", log_dir)
```
### Train and validate the model
```
EPOCHS = 10
steps_per_epoch = 60000//global_batch_size # 60,000 items in this dataset
print("Step (batches) per epoch: ", steps_per_epoch)
# Counting steps and batches on TPU: the tpu.keras_to_tpu_model API regards the batch size of the input dataset
# as the per-core batch size. The effective batch size is 8x more because Cloud TPUs have 8 cores. It increments
# the step by +8 everytime a global batch (8 per-core batches) is processed. Therefore batch size and steps_per_epoch
# settings can stay as they are for TPU training. The training will just go faster.
# Warning: this might change in the final version of the Keras/TPU API.
history = model.fit(training_dataset, steps_per_epoch=steps_per_epoch, epochs=EPOCHS,
validation_data=validation_dataset, validation_steps=1, callbacks=[lr_decay, tb_callback])
```
### Visualize predictions
```
# recognize digits from local fonts
probabilities = model.predict(font_digits, steps=1)
predicted_labels = np.argmax(probabilities, axis=1)
display_digits(font_digits, predicted_labels, font_labels, "predictions from local fonts (bad predictions in red)", N)
# recognize validation digits
probabilities = model.predict(validation_digits, steps=1)
predicted_labels = np.argmax(probabilities, axis=1)
display_top_unrecognized(validation_digits, predicted_labels, validation_labels, N, 7)
```
### Export the model for serving from ML Engine
```
class ServingInput(tf.keras.layers.Layer):
# the important detail in this boilerplate code is "trainable=False"
def __init__(self, name, dtype, batch_input_shape=None):
super(ServingInput, self).__init__(trainable=False, name=name, dtype=dtype, batch_input_shape=batch_input_shape)
def get_config(self):
return {'batch_input_shape': self._batch_input_shape, 'dtype': self.dtype, 'name': self.name }
def call(self, inputs):
# When the deployed model is called through its REST API,
# the JSON payload is parsed automatically, transformed into
# a tensor and passed to this input layer. You can perform
# additional transformations, such as decoding JPEGs for example,
# before sending the data to your model. However, you can only
# use tf.xxxx operations.
return inputs
# little wrinkle: must copy the model from TPU to CPU manually. This is a temporary workaround.
restored_model = make_model()
restored_model.set_weights(model.get_weights()) # this copied the weights from TPU, does nothing on GPU
# add the serving input layer
serving_model = tf.keras.Sequential()
serving_model.add(ServingInput('serving', tf.float32, (None, 28*28)))
serving_model.add(restored_model)
export_path = os.path.join(BUCKET, 'mnist-export', timestamp)
tf.saved_model.save(serving_model, export_path)
print("Model exported to: ", export_path)
```
## Deploy the trained model to AI Platform
Push your trained model to production on AI Platform for a serverless, autoscaled, REST API experience.
You will need a GCS bucket and a GCP project for this.
Models deployed on AI Platform autoscale to zero if not used. There will be no ML Engine charges after you are done testing.
Google Cloud Storage incurs charges. Empty the bucket after deployment if you want to avoid these. Once the model is deployed, the bucket is not useful anymore.
### Cloud Configuration
```
# Enable model deployment here
DEPLOY = False # #@param {type:"boolean"}
# Create the model only once, after that, create new versions of the same model
CREATE_MODEL = True #@param {type:"boolean"}
# Models are deployed in your cloud project
PROJECT = "" #@param {type:"string"}
MODEL_NAME = "mnist" #@param {type:"string"}
MODEL_VERSION = "v0" #@param {type:"string"}
if DEPLOY:
assert PROJECT, 'For this part, you need a GCP project. Head to http://console.cloud.google.com/ and create one.'
assert re.search(r'gs://.+', export_path), 'For this part, the model must have been exported to a GCS bucket.'
```
### Deploy the model
This uses the command-line interface. You can do the same thing through the ML Engine UI at https://console.cloud.google.com/mlengine/models
```
# Create the model
if DEPLOY and CREATE_MODEL:
!gcloud ai-platform models create {MODEL_NAME} --project={PROJECT} --regions=us-central1
# Create a version of this model (you can add --async at the end of the line to make this call non blocking)
# Additional config flags are available: https://cloud.google.com/ml-engine/reference/rest/v1/projects.models.versions
# You can also deploy a model that is stored locally by providing a --staging-bucket=... parameter
if DEPLOY:
!echo "Deployment takes a couple of minutes. You can watch your deployment here: https://console.cloud.google.com/mlengine/models/{MODEL_NAME}"
!gcloud ai-platform versions create {MODEL_VERSION} --model={MODEL_NAME} --origin="{export_path}" --project={PROJECT} --runtime-version=1.13 --python-version=3.5
```
### Test the deployed model
Your model is now available as a REST API. Let us try to call it. The cells below use the "gcloud ml-engine"
command line tool but any tool that can send a JSON payload to a REST endpoint will work.
```
# prepare digits to send to online prediction endpoint
digits = np.concatenate((font_digits, validation_digits[:100-N]))
labels = np.concatenate((font_labels, validation_labels[:100-N]))
with open("digits.json", "w") as f:
for digit in digits:
# the format for ML Engine online predictions is: one JSON object per line
data = json.dumps({"serving_input": digit.tolist()}) # "serving_input" because the ServingInput layer was named "serving". Keras appends "_input"
f.write(data+'\n')
if DEPLOY: # Request online predictions from deployed model (REST API) using the "gcloud ai-platform" command line.
predictions = !gcloud ai-platform predict --model={MODEL_NAME} --json-instances digits.json --project={PROJECT} --version {MODEL_VERSION}
print(predictions)
probabilities = np.stack([json.loads(p) for p in predictions[1:]]) # first line is the name of the input layer: drop it, parse the rest
predictions = np.argmax(probabilities, axis=1)
display_top_unrecognized(digits, predictions, labels, N, 100//N)
```
## License
---
author: Martin Gorner<br>
twitter: @martin_gorner
---
Copyright 2019 Google LLC
Licensed under the Apache License, Version 2.0 (the "License");
you may not use this file except in compliance with the License.
You may obtain a copy of the License at
http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software
distributed under the License is distributed on an "AS IS" BASIS,
WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
See the License for the specific language governing permissions and
limitations under the License.
---
This is not an official Google product but sample code provided for an educational purpose
| github_jupyter |
[Loss Function](https://www.bualabs.com/archives/2673/what-is-loss-function-cost-function-error-function-loss-function-how-cost-function-work-machine-learning-ep-1/) หรือ Cost Function คือ การคำนวน Error ว่า yhat ที่โมเดลทำนายออกมา ต่างจาก y ของจริง อยู่เท่าไร แล้วหาค่าเฉลี่ย เพื่อที่จะนำมาหา Gradient ของ Loss ขึ้นกับ Weight ต่าง ๆ ด้วย Backpropagation แล้วใช้อัลกอริทึม Gradient Descent เพื่อให้ Loss น้อยลง ในการเทรนรอบถัดไป
ใน ep ก่อนเราได้สอน [Cross Entropy Loss](https://www.bualabs.com/archives/1945/what-is-cross-entropy-loss-logistic-regression-log-loss-loss-function-ep-3/) ไปแล้ว ในเคสนี้เราจะพูดถึง Loss Function สำหรับงาน [Regression](https://www.bualabs.com/archives/1136/train-deep-learning-model-deep-neural-networks-estimate-center-point-biwi-kinect-head-pose-regression-ep-1/) (ค่าต่อเนื่อง) กัน และ Regression Loss ที่เป็นที่นิยมมากที่สุด ได้แก่ MAE, MSE, RMSE
yhat และ y สามารถเป็น Vector หรือ เลขตัวเดียวก็ได้
# 0. Import
```
import torch
from torch import tensor
import matplotlib.pyplot as plt
```
# 1. Data
เราจะสร้างข้อมูลตัวอย่างขึ้นมา เป็นสมการเส้นตรง ให้ x เป็น เลขตั้งแต่ -50 ถึง 50 เราจะได้เอาไว้พล็อตกราฟ
```
x = torch.arange(-50., 50.)
x
```
ประกาศฟังก์ชันเส้นตรง f(x) = y = ax + b
```
a = 4
b = 2
# a = -2
# b = 2
def f(x):
return (a * x) + b
```
## yhat
นำ x ผ่านฟังก์ชัน ได้ค่า yhat
```
yhat = f(x)
yhat
```
## y
ส่วนค่า y ที่เราต้องการจริง ๆ เป็นดังนี้
```
y = tensor([102., 100., 98., 96., 94., 92., 90., 88., 86., 84., 82., 80.,
78., 76., 74., 72., 70., 68., 66., 64., 62., 60., 58., 56.,
54., 52., 50., 48., 46., 44., 42., 40., 38., 36., 34., 32.,
30., 28., 26., 24., 22., 20., 18., 16., 14., 12., 10., 8.,
6., 4., 2., 0., -2., -4., -6., -8., -10., -12., -14., -16.,
-18., -20., -22., -24., -26., -28., -30., -32., -34., -36., -38., -40.,
-42., -44., -46., -48., -50., -52., -54., -56., -58., -60., -62., -64.,
-66., -68., -70., -72., -74., -76., -78., -80., -82., -84., -86., -88.,
-90., -92., -94., -96.])
y
```
## เปรียบเทียบ y, yhat
นำมาพล็อตกราฟ เปรียบเทียบกัน
```
fig,ax = plt.subplots(figsize=(9, 9))
ax.scatter(x, y, label="y")
ax.plot(x.numpy(), yhat.numpy(), label="yhat", color='red')
ax.legend(loc='upper right')
```
# 2. Mean Abolute Error (MAE) หรือ Least Absolute Deviations (L1 Loss)
Mean Abolute Error (MAE) หรือ L1 Loss คือ การคำนวน Error ว่า yhat ต่างจาก y อยู่เท่าไร ด้วยการนำมาลบกันตรง ๆ แล้วหาค่าเฉลี่ย โดยไม่สนใจเครื่องหมาย (Absolute) เพื่อหาขนาดของ Error โดยไม่สนใจทิศทาง
## 2.1 สูตร MAE
$$\mathrm{MAE} = \frac{\sum_{i=1}^n\left| y_i-\hat{y_i}\right|}{n}$$
## 2.2 โค้ด MAE Function
```
def mae(y, yhat):
return (y - yhat).abs().mean()
```
## 2.3 การใช้งาน MAE
```
error = mae(y, yhat)
error
```
## 2.4 เปรียบเทียบ Loss
แทนที่เราจะหาค่าเฉลี่ย เราจะลองเปรียบเทียบ yhat, y ตัวต่อตัว สังเกต กราฟสีม่วง ความชันจะคงที่ และจุดต่ำสุดจะอยู่ที่ y - yhat = 0 คือ โมเดลทำนาย yhat ออกมาเท่ากับ y พอดี ทำให้ MAE Loss = 0
```
fig,ax = plt.subplots(figsize=(9, 9))
ax.scatter(x, y, label="y")
ax.plot(x.numpy(), yhat.numpy(), label="yhat", color='red')
ax.plot(x.numpy(), (y - yhat).abs().numpy(), label="MAE", color='purple')
ax.legend(loc='upper right')
```
# 3. Mean Squared Error (MSE) หรือ Least Square Errors (L2 Loss)
Mean Squared Error (MSE), Quadratic Loss หรือ L2 Loss คือ การคำนวน Error ว่า yhat ต่างจาก y อยู่เท่าไร ด้วยการนำมาลบกัน แล้วยกกำลังสอง (Squared) เพื่อไม่ต้องสนใจค่าติดลบ (ถ้ามี) แล้วหาค่าเฉลี่ย
เนื่องจากมีการยกกำลังสอง ทำให้ค่อนข้าง Sensitive ถ้าข้อมูลไม่ดี มีบางตัวที่นอกลู่นอกทาง yhat ตัวไหนที่ผิดจาก y ไปมาก จะถูกให้ความสำคัญมากกว่า yhat ตัวอื่น ๆ จะมีผลทำให้ Loss สูง
## 3.1 สูตร MSE
$$\operatorname{MSE}=\frac{1}{n}\sum_{i=1}^n(Y_i-\hat{Y_i})^2$$
## 3.2 โค้ด MSE Function
```
def mse(y, yhat):
return (y - yhat).pow(2).mean()
```
## 3.3 การใช้งาน MSE
```
error = mse(y, yhat)
error
```
## 3.4 เปรียบเทียบ Loss
แทนที่เราจะหาค่าเฉลี่ย เราจะลองเปรียบเทียบ yhat, y ตัวต่อตัว สังเกต กราฟสีม่วง ความชันจะขึ้นเร็วมาก เป็น Exponential เนื่องจาก MSE ยกกำลัง 2 และจุดต่ำสุดจะอยู่ที่ y - yhat = 0 คือ โมเดลทำนาย yhat ออกมาเท่ากับ y พอดี ทำให้ MSE Loss = 0
```
fig,ax = plt.subplots(figsize=(9, 9))
ax.set_xlim([-50, 50])
ax.set_ylim([-200, 2000])
ax.scatter(x, y, label="y")
ax.plot(x.numpy(), yhat.numpy(), label="yhat", color='red')
ax.plot(x.numpy(), (y - yhat).pow(2).numpy(), label="MSE", color='purple')
ax.legend(loc='upper right')
```
# 4. RMSE หรือ RMSD
## 4.1 สูตร RMSE
$$\operatorname{RMSD}=\sqrt{\frac{\sum_{i=1}^n ( y_i - \hat y_i)^2}{n}}$$
Root Mean Squared Error (RMSE) หรือ Root Mean Squared Deviation (RMSD) คือ นำ MSE มาหา Squared Root ช่วยให้กลับมาเป็น Scale เดิม ทำให้ตีความได้ง่ายขึ้น
## 4.2 โค้ด RMSE Function
```
def rmse(y, yhat):
return ((y - yhat).pow(2).mean()).sqrt()
```
## 4.3 การใช้งาน RMSE
```
error = rmse(y, yhat)
error
```
## 4.5 เปรียบเทียบ Loss
```
fig,ax = plt.subplots(figsize=(9, 9))
ax.set_xlim([-50, 50])
ax.set_ylim([-200, 300])
ax.scatter(x, y, label="y")
ax.plot(x.numpy(), yhat.numpy(), label="yhat", color='red')
ax.plot(x.numpy(), (y - yhat).pow(2).sqrt().numpy(), label="RMSE", color='purple')
ax.legend(loc='upper right')
```
# 5. เปรียบเทียบ MAE และ MSE
1. ค่า MSE, MAE อยู่ในช่วง 0 - Infinity เหมือนกัน ยิ่งน้อยคือยิ่งดี ถ้าเป็น 0 คือ ไม่ Error เลย
1. MAE มี Slope คงที่ หมายถึง Gradient จากมีขนาดใหญ่เท่าเดิม แม้ Error จะน้อยแค่ไหนก็ตาม
1. MSE มี Slope ที่เปลี่ยนแปลงไปตาม Error ถ้า Error มาก Gradient ก็มาก ถ้า Error น้อย Gradient ก็จะน้อยตามไปด้วย ช่วยให้เทรนโมเดล Fine-Tune ได้ดีกว่า
1. เนื่องจาก MSE ยกกำลังสอง ก่อนที่จะหาค่าเฉลี่ย ทำให้ Sensitive กับข้อมูลแตกแถว มากกว่า MAE ถ้า MSE เจอข้อมูลไม่ดี จะทำให้ Loss มีขนาดใหญ่มาก
1. เราสามารถแก้ปัญหา Gradient คงที่ของ MAE ได้ด้วยเทคนิค Dynamic Learning Rate
1. ถ้าข้อมูลตัวอย่างของเรามี ข้อมูลเสีย ๆ ปะปนอยู่ MAE จะเหมาะกว่า MSE
1. MAE ใช้การ Absolute ทำให้อาจจะทำให้มีปัญหาได้ กับการคำนวนทางคณิตศาสตร์อื่น ๆ
# 6. Loss Function อื่น ๆ สำหรับงาน Regression
นอกจาก MAE, MSE และ RMSE แล้ว มีตัวอื่น ๆ ที่เป็นที่นิยมอีก เช่น Huber Loss (Smooth Mean Absolute Error), Log-Cosh Loss, Quantile Loss เราจะอธิบายต่อไป
```
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Premade Estimators
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/alpha/tutorials/estimators/premade_estimators"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/r2/tutorials/estimators/premade_estimators.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/tree/master/site/en/r2/tutorials/estimators/premade_estimators.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
</table>
This tutorial shows you
how to solve the Iris classification problem in TensorFlow using Estimators. An Estimator is TensorFlow's high-level representation of a complete model, and it has been designed for easy scaling and asynchronous training. For more details see
[Estimators](https://www.tensorflow.org/guide/estimators).
Note that in TensorFlow 2.0, the [Keras API](https://www.tensorflow.org/guide/keras) can accomplish many of these same tasks, and is believed to be an easier API to learn. If you are starting fresh, we would recommend you start with Keras. For more information about the available high level APIs in TensorFlow 2.0, see [Standardizing on Keras](https://medium.com/tensorflow/standardizing-on-keras-guidance-on-high-level-apis-in-tensorflow-2-0-bad2b04c819a).
## First things first
In order to get started, you will first import TensorFlow and a number of libraries you will need.
```
from __future__ import absolute_import, division, print_function, unicode_literals
!pip install tensorflow==2.0.0-alpha0
import tensorflow as tf
import pandas as pd
```
## The data set
The sample program in this document builds and tests a model that
classifies Iris flowers into three different species based on the size of their
[sepals](https://en.wikipedia.org/wiki/Sepal) and
[petals](https://en.wikipedia.org/wiki/Petal).
You will train a model using the Iris data set. The Iris data set contains four features and one
[label](https://developers.google.com/machine-learning/glossary/#label).
The four features identify the following botanical characteristics of
individual Iris flowers:
* sepal length
* sepal width
* petal length
* petal width
Based on this information, you can define a few helpful constants for parsing the data:
```
CSV_COLUMN_NAMES = ['SepalLength', 'SepalWidth', 'PetalLength', 'PetalWidth', 'Species']
SPECIES = ['Setosa', 'Versicolor', 'Virginica']
```
Next, download and parse the Iris data set using Keras and Pandas. Note that you keep distinct datasets for training and testing.
```
train_path = tf.keras.utils.get_file(
"iris_training.csv", "https://storage.googleapis.com/download.tensorflow.org/data/iris_training.csv")
test_path = tf.keras.utils.get_file(
"iris_test.csv", "https://storage.googleapis.com/download.tensorflow.org/data/iris_test.csv")
train = pd.read_csv(train_path, names=CSV_COLUMN_NAMES, header=0)
test = pd.read_csv(test_path, names=CSV_COLUMN_NAMES, header=0)
```
You can inspect your data to see that you have four float feature columns and one int32 label.
```
train.head()
```
For each of the datasets, split out the labels, which the model will be trained to predict.
```
train_y = train.pop('Species')
test_y = test.pop('Species')
# The label column has now been removed from the features.
train.head()
```
## Overview of programming with Estimators
Now that you have the data set up, you can define a model using a TensorFlow Estimator. An Estimator is any class derived from `tf.estimator.Estimator`. TensorFlow
provides a collection of
`tf.estimator`
(for example, `LinearRegressor`) to implement common ML algorithms. Beyond
those, you may write your own
[custom Estimators](https://www.tensorflow.org/guide/custom_estimators).
We recommend using pre-made Estimators when just getting started.
To write a TensorFlow program based on pre-made Estimators, you must perform the
following tasks:
* Create one or more input functions.
* Define the model's feature columns.
* Instantiate an Estimator, specifying the feature columns and various
hyperparameters.
* Call one or more methods on the Estimator object, passing the appropriate
input function as the source of the data.
Let's see how those tasks are implemented for Iris classification.
## Create input functions
You must create input functions to supply data for training,
evaluating, and prediction.
An **input function** is a function that returns a `tf.data.Dataset` object
which outputs the following two-element tuple:
* [`features`](https://developers.google.com/machine-learning/glossary/#feature) - A Python dictionary in which:
* Each key is the name of a feature.
* Each value is an array containing all of that feature's values.
* `label` - An array containing the values of the
[label](https://developers.google.com/machine-learning/glossary/#label) for
every example.
Just to demonstrate the format of the input function, here's a simple
implementation:
```
def input_evaluation_set():
features = {'SepalLength': np.array([6.4, 5.0]),
'SepalWidth': np.array([2.8, 2.3]),
'PetalLength': np.array([5.6, 3.3]),
'PetalWidth': np.array([2.2, 1.0])}
labels = np.array([2, 1])
return features, labels
```
Your input function may generate the `features` dictionary and `label` list any
way you like. However, we recommend using TensorFlow's [Dataset API](https://www.tensorflow.org/guide/datasets), which can
parse all sorts of data.
The Dataset API can handle a lot of common cases for you. For example,
using the Dataset API, you can easily read in records from a large collection
of files in parallel and join them into a single stream.
To keep things simple in this example you are going to load the data with
[pandas](https://pandas.pydata.org/), and build an input pipeline from this
in-memory data:
```
def input_fn(features, labels, training=True, batch_size=256):
"""An input function for training or evaluating"""
# Convert the inputs to a Dataset.
dataset = tf.data.Dataset.from_tensor_slices((dict(features), labels))
# Shuffle and repeat if you are in training mode.
if training:
dataset = dataset.shuffle(1000).repeat()
return dataset.batch(batch_size)
```
## Define the feature columns
A [**feature column**](https://developers.google.com/machine-learning/glossary/#feature_columns)
is an object describing how the model should use raw input data from the
features dictionary. When you build an Estimator model, you pass it a list of
feature columns that describes each of the features you want the model to use.
The `tf.feature_column` module provides many options for representing data
to the model.
For Iris, the 4 raw features are numeric values, so we'll build a list of
feature columns to tell the Estimator model to represent each of the four
features as 32-bit floating-point values. Therefore, the code to create the
feature column is:
```
# Feature columns describe how to use the input.
my_feature_columns = []
for key in train.keys():
my_feature_columns.append(tf.feature_column.numeric_column(key=key))
```
Feature columns can be far more sophisticated than those we're showing here. You can read more about Feature Columns in [this guide](https://www.tensorflow.org/guide/feature_columns).
Now that you have the description of how you want the model to represent the raw
features, you can build the estimator.
## Instantiate an estimator
The Iris problem is a classic classification problem. Fortunately, TensorFlow
provides several pre-made classifier Estimators, including:
* `tf.estimator.DNNClassifier` for deep models that perform multi-class
classification.
* `tf.estimator.DNNLinearCombinedClassifier` for wide & deep models.
* `tf.estimator.LinearClassifier` for classifiers based on linear models.
For the Iris problem, `tf.estimator.DNNClassifier` seems like the best choice.
Here's how you instantiated this Estimator:
```
# Build a DNN with 2 hidden layers with 30 and 10 hidden nodes each.
classifier = tf.estimator.DNNClassifier(
feature_columns=my_feature_columns,
# Two hidden layers of 10 nodes each.
hidden_units=[30, 10],
# The model must choose between 3 classes.
n_classes=3)
```
## Train, Evaluate, and Predict
Now that you have an Estimator object, you can call methods to do the following:
* Train the model.
* Evaluate the trained model.
* Use the trained model to make predictions.
### Train the model
Train the model by calling the Estimator's `train` method as follows:
```
# Train the Model.
classifier.train(
input_fn=lambda: input_fn(train, train_y, training=True),
steps=5000)
```
Note that you wrap up your `input_fn` call in a
[`lambda`](https://docs.python.org/3/tutorial/controlflow.html)
to capture the arguments while providing an input function that takes no
arguments, as expected by the Estimator. The `steps` argument tells the method
to stop training after a number of training steps.
### Evaluate the trained model
Now that the model has been trained, you can get some statistics on its
performance. The following code block evaluates the accuracy of the trained
model on the test data:
```
eval_result = classifier.evaluate(
input_fn=lambda: input_fn(test, test_y, training=False))
print('\nTest set accuracy: {accuracy:0.3f}\n'.format(**eval_result))
```
Unlike the call to the `train` method, you did not pass the `steps`
argument to evaluate. The `input_fn` for eval only yields a single
[epoch](https://developers.google.com/machine-learning/glossary/#epoch) of data.
The `eval_result` dictionary also contains the `average_loss` (mean loss per sample), the `loss` (mean loss per mini-batch) and the value of the estimator's `global_step` (the number of training iterations it underwent).
### Making predictions (inferring) from the trained model
You now have a trained model that produces good evaluation results.
You can now use the trained model to predict the species of an Iris flower
based on some unlabeled measurements. As with training and evaluation, you make
predictions using a single function call:
```
# Generate predictions from the model
expected = ['Setosa', 'Versicolor', 'Virginica']
predict_x = {
'SepalLength': [5.1, 5.9, 6.9],
'SepalWidth': [3.3, 3.0, 3.1],
'PetalLength': [1.7, 4.2, 5.4],
'PetalWidth': [0.5, 1.5, 2.1],
}
def input_fn(features, batch_size=256):
"""An input function for prediction."""
# Convert the inputs to a Dataset without labels.
return tf.data.Dataset.from_tensor_slices(dict(features)).batch(batch_size)
predictions = classifier.predict(
input_fn=lambda: input_fn(predict_x))
```
The `predict` method returns a Python iterable, yielding a dictionary of
prediction results for each example. The following code prints a few
predictions and their probabilities:
```
for pred_dict, expec in zip(predictions, expected):
class_id = pred_dict['class_ids'][0]
probability = pred_dict['probabilities'][class_id]
print('Prediction is "{}" ({:.1f}%), expected "{}"'.format(
SPECIES[class_id], 100 * probability, expec))
```
| github_jupyter |
# Model comparisons
In this notebook, we'll take BartPy through its paces using increasingly complex sin wave models. We'll compare how it performs to two similar models: OLS and catboost.
For the purposes of this exercise, I'm testing out of the box performance. This makes sense, as part of the value prop of BartPy is it's ability to work well without parameter tuning, but it's possible that the scores of all of the models could be improved with parameter tuning
```
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
import catboost
from catboost import Pool
from sklearn.model_selection import cross_val_score
from sklearn.linear_model import LinearRegression
from sklearn.metrics import r2_score
from sklearn.model_selection import KFold
from copy import deepcopy
from bartpy.sklearnmodel import SklearnModel
%matplotlib inline
```
## Set up models
```
def fit_catboost(X_train, y_train, X_test, y_test):
eval_cutoff = len(X_train) // 3
eval_X, eval_y, train_X, train_y = X_train[:eval_cutoff, :], y_train[:eval_cutoff], X_train[eval_cutoff:, :], y_train[eval_cutoff:]
catboost_model = catboost.CatBoostRegressor()
catboost_model.fit(Pool(train_X, train_y), eval_set = Pool(eval_X, eval_y), use_best_model=True)
pred = catboost_model.predict(X_test)
score = r2_score(y_test, pred)
return catboost_model, pred, score
def fit_bartpy(X_train, y_train, X_test, y_test):
model = SklearnModel(n_samples=1000, n_burn=50, n_trees=500, store_in_sample_predictions=False)
model.fit(X_train, y_train)
pred = model.predict(X_test)
score = model.score(X_test, y_test)
return model, pred, score
def fit_linear_model(X_train, y_train, X_test, y_test):
rebased_x = np.sin(X_train[:, 0]).reshape(-1, 1)
linear_model = LinearRegression()
linear_model.fit(rebased_x, y_train)
pred = linear_model.predict(np.sin(X_test[:, 0]).reshape(-1, 1))
score = linear_model.score(np.sin(X_test[:, 0]).reshape(-1, 1), y_test)
return linear_model, pred, score
kf = KFold(2)
def compare_models(X, y, models_funcs = [fit_bartpy, fit_catboost, fit_linear_model]):
scores, predictions, trained_models = [], [], []
for train_index, test_index in kf.split(X):
trained_models.append([])
scores.append([])
predictions.append([])
for f in models_funcs:
model, pred, score = f(X[train_index, :], y[train_index], X[test_index, :], y[test_index])
trained_models[-1].append(deepcopy(model))
predictions[-1].append(pred)
scores[-1].append(score)
return scores, predictions, trained_models
```
## One dimensional, single wave
```
x = np.linspace(0, 5, 3000)
X = pd.DataFrame(x).sample(frac=1.0).values
y = np.random.normal(0, 0.1, size=3000) + np.sin(X[:, 0])
plt.scatter(X[:, 0], y)
plt.xlabel("x")
plt.ylabel("y")
plt.title("X v y")
scores, predictions, models = compare_models(X, y, [fit_bartpy])
scores
scores, predictions, trained_models = compare_models(X, y)
```
**As we might expect, all three models are capable of capturing such a simple function. There's very little difference between the scores or predictions of the models.**
```
scores
kf = KFold(2)
for p, (train_index, test_index) in zip(predictions, kf.split(X)):
for m in p:
plt.scatter(y[test_index], m)
plt.title("Preicted V Actual Per Model")
plt.xlabel("True Target")
plt.ylabel("Predicted Target")
plt.scatter(X[:1500,0],predictions[0][1])
plt.scatter(X[:1500,0],predictions[0][2])
plt.scatter(X[:1500,0],predictions[0][0])
plt.ylabel("Prediction")
plt.xlabel("Covariate")
plt.title("Prediction by model by value of X")
```
## Single dimension - multiple waves
** To extend our original model, let's add a high frequency cosine wave in our single dimension. This will make the predicted function change faster and at varying rates across x **
```
x = np.linspace(0, 5, 3000)
X = pd.DataFrame(x).sample(frac=1.0).values
y = np.random.normal(0, 0.1, size=3000) + np.sin(X[:, 0]) + np.cos(5 * X[:, 0])
plt.scatter(X[:, 0], y)
plt.xlabel("x")
plt.ylabel("y")
plt.title("X v y")
scores, predictions, trained_models = compare_models(X, y)
```
** Catboost and BartPy capture this faster moving sin wave pretty nicely, and come up with very similar predictions. The linear model pretty much treats the cos wave as noise, and fits a curve close to the original one **
```
scores
kf = KFold(2)
for p, (train_index, test_index) in zip(predictions, kf.split(X)):
for m in p:
plt.scatter(y[test_index], m)
plt.title("Preicted V Actual Per Model")
plt.xlabel("True Target")
plt.ylabel("Predicted Target")
plt.scatter(X[:1500,0],predictions[0][1])
plt.scatter(X[:1500,0],predictions[0][2])
plt.scatter(X[:1500,0],predictions[0][0])
plt.ylabel("Prediction")
plt.xlabel("Covariate")
plt.title("Prediction by model by value of X")
import seaborn as sns
p = pd.DataFrame(np.array(predictions[1]).T, columns = ["Catboost", "BartPy", "Linear"])
sns.pairplot(p)
```
## Single dimension - discrete break points
** To make the model more complex, let's add some discrete dumps to make the curve less smooth. This will test the ability of the models to handle very isolated effects in feature space **
```
x = np.linspace(0, 5, 3000)
X = pd.DataFrame(x).sample(frac=1.0).values
y = np.random.normal(0, 0.1, size=3000) + np.sin(X[:, 0]) + np.cos(5 * X[:, 0])
y[(X[:,0] < 1.5) & (X[:,0] > 1.)] += 3
y[(X[:,0] < 3.5) & (X[:,0] > 3.)] -= 3
plt.scatter(X[:, 0], y)
plt.xlabel("x")
plt.ylabel("y")
plt.title("X v y")
scores, predictions, trained_models = compare_models(X, y)
scores
kf = KFold(2)
for p, (train_index, test_index) in zip(predictions, kf.split(X)):
for m in p:
plt.scatter(y[test_index], m)
plt.title("Preicted V Actual Per Model")
plt.xlabel("True Target")
plt.ylabel("Predicted Target")
plt.scatter(X[:1500,0],predictions[0][1])
plt.scatter(X[:1500,0],predictions[0][2])
plt.scatter(X[:1500,0],predictions[0][0])
plt.ylabel("Prediction")
plt.xlabel("Covariate")
plt.title("Prediction by model by value of X")
import seaborn as sns
p = pd.DataFrame(np.array(predictions[1]).T, columns = ["Catboost", "BartPy", "Linear"])
sns.pairplot(p)
```
# Single meaningful dimension - additional noise dimensions
** It's important for the models to be able to correctly identify features that aren't important. In this case, we just add normally distributed noise features. This shouldn't be too difficult a problem to solve **
```
x = np.linspace(0, 5, 3000)
X = np.random.normal(0, 3, size = 3000 * 8).reshape(3000, 8)
X[:, 0] = x
X = pd.DataFrame(X).sample(frac=1.0).values
y = np.random.normal(0, 0.1, size=3000) + np.sin(X[:, 0])
plt.scatter(X[:, 0], y)
plt.scatter(X[:, 1], y)
plt.xlabel("x")
plt.ylabel("y")
plt.title("X v y")
scores, predictions, trained_models = compare_models(X, y)
scores
kf = KFold(2)
for p, (train_index, test_index) in zip(predictions, kf.split(X)):
for m in p:
plt.scatter(y[test_index], m)
plt.title("Preicted V Actual Per Model")
plt.xlabel("True Target")
plt.ylabel("Predicted Target")
plt.scatter(X[:1500,0],predictions[0][1])
plt.scatter(X[:1500,0],predictions[0][2])
plt.scatter(X[:1500,0],predictions[0][0])
plt.ylabel("Prediction")
plt.xlabel("Covariate")
plt.title("Prediction by model by value of X")
import seaborn as sns
p = pd.DataFrame(np.array(predictions[1]).T, columns = ["Catboost", "BartPy", "Linear"])
sns.pairplot(p)
```
## Mixture of multiple meaningful dimensions and noise dimensions
** Both catboost and BartPy have performed well with noise dimensions and rapidly moving targets, let's combine them to make a relatively difficult test for tree models **
```
X = np.random.normal(0, 3, size = 3000 * 8).reshape(3000, 8)
X = pd.DataFrame(X).sample(frac=1.0).values
y = np.random.normal(0, 0.1, size=3000) + np.sin(X[:, 0]) + np.sin(5 * X[:, 1]) + np.cos(-6 * X[:, 3])
plt.scatter(X[:, 0], y)
plt.scatter(X[:, 1], y)
plt.scatter(X[:, 3], y)
plt.xlabel("x")
plt.ylabel("y")
plt.title("X v y")
scores, predictions, trained_models = compare_models(X, y, [fit_bartpy])
scores
scores
#plt.scatter(X[:1500,3],predictions[0][1], label = "Bart")
#plt.scatter(X[:1500,3],predictions[0][2], label = "Ols")
plt.scatter(X[:1500,3],predictions[0][0], label = "Catboost")
plt.ylabel("Prediction")
plt.xlabel("Covariate")
plt.title("Prediction by model by value of X")
plt.legend(loc = "best")
plt.scatter(y[:1500],predictions[0][0], label = "Catboost")
#plt.scatter(y[:1500],predictions[0][1], label = "Catboost")
#plt.scatter(y[:1500],predictions[0][2], label = "Catboost")
import seaborn as sns
p = pd.DataFrame(np.array(predictions[1]).T, columns = ["Catboost", "BartPy", "Linear"])
sns.pairplot(p)
```
| github_jupyter |
# Imports
```
import pandas as pd
import numpy as np
from collections import defaultdict
from tensorflow import keras
from matplotlib import pyplot as plt
import copy
from functions.data_processing import create_set
from functions.prediction import make_predictions, generate_pdda_preds
from functions.visualization import posHeatmapXY, spatial_plot
import warnings
warnings.filterwarnings("ignore")
```
# Dataset Import
The dataset is hosted publicly at https://doi.org/10.5281/zenodo.6303665.
```
datadir = '../data' # path to data folder
modeldir = '../models' # path to model folder (optional, if you want to save models' weights)
preds_dir = '../preds' # path to model folder (optional, if you want to save models' predictions)
#define basic values
rooms = ['testbench_01', 'testbench_01_furniture_low', 'testbench_01_furniture_mid', 'testbench_01_furniture_high']
concrete_rooms = ['testbench_01_furniture_low_concrete', 'testbench_01_furniture_mid_concrete', 'testbench_01_furniture_high_concrete']
other_scenarios = ['testbench_01_rotated_anchors', 'testbench_01_translated_anchors']
anchors = ['anchor1', 'anchor2', 'anchor3', 'anchor4']
channels = ['37','38','39']
polarities = ['V','H']
#read data
data = defaultdict(lambda: defaultdict(lambda: defaultdict (lambda: defaultdict(list))))
anchor_data = defaultdict(lambda: defaultdict(lambda: defaultdict (lambda: defaultdict(list))))
for room in rooms + concrete_rooms + other_scenarios:
for channel in channels:
for polarity in polarities:
tag_filename = f'{datadir}/{room}/tag_ml_export_CH{channel}_{polarity}.json'
tag_df = pd.read_json(tag_filename, orient='records')
anchor_filename = f'{datadir}/{room}/anchor_ml_export_CH{channel}_{polarity}.json'
anchor_df = pd.read_json(anchor_filename, orient='records')
df = tag_df.merge(anchor_df)
# remove calibration points
df.drop(df[(df['x_tag']==0).values | (df['y_tag']==0).values | (df['z_tag']==0).values].index, inplace=True)
for anchor in anchors:
data[room][anchor][channel][polarity] = df[df['anchor']==int(anchor[-1])]
anchor_data[room][anchor][channel][polarity] = anchor_df
```
# Data Processing
The selected number of training and validation points is small so the point selection is done in an orderly fashion.
```
#split points into train/test/val points
points = data['testbench_01']['anchor1']['37']['H'].iloc[:, 1:7]
# only point locations that appear in all simulated environments are used.
# some point locations that fall ontop of furniture are thus thrown away.
for room in rooms + concrete_rooms:
for anchor in anchors:
for channel in channels:
for polarization in ['H','V']:
points = pd.merge(points, data[room][anchor][channel][polarization]['point'], on='point')
# grid of training points
xs = sorted(np.unique(points['x_tag']))[::6]
ys = sorted(np.unique(points['y_tag']))[::3]
train_points = points[points['x_tag'].isin(xs) & points['y_tag'].isin(ys)]
# grid of validation points
xs = sorted(np.unique(points['x_tag']))[3::10]
ys = sorted(np.unique(points['y_tag']))[3::10]
val_points = points[points['x_tag'].isin(xs) & points['y_tag'].isin(ys)]
test_points = points.drop(index=train_points.index).drop(index=val_points.index)
print(f'Training Set Size:\t{len(train_points)}')
print(f'Validation Set Size:\t{len(val_points)}')
print(f'Testing Set Size:\t{len(test_points)}')
plt.scatter(train_points.iloc[:,1:2].values,train_points.iloc[:,2:3].values)
plt.scatter(val_points.iloc[:,1:2].values,val_points.iloc[:,2:3].values)
plt.legend(['Training Points', 'Validation Points'], framealpha=0.94, fancybox=True)
ax = plt.gca()
ax.set_xticklabels(range(-2,15,2))
ax.set_yticklabels(range(-1,7,1))
plt.show()
```
Split the dataset based on the picked training, validation and testing points. The create_set function processes the IQ and RSSI data by applying IQ phase shifting and RSSI normalization.
The training set is augmented by reducing amplitude of IQ values and RSSI for randomly picked anchors.
```
#creat train/test/val sets
x_train, y_train = create_set(data, rooms, train_points, augmentation=True)
x_val, y_val = create_set(data, rooms + concrete_rooms + other_scenarios, val_points)
x_test, y_test = create_set(data, rooms + concrete_rooms + other_scenarios, test_points)
```
# Training
```
from functions.models import jointArch
model_arch = jointArch
model_arch_name = 'joint_arch'
#training parameters
fit_params = {'batch_size': 128, 'validation_batch_size':32, 'epochs': 1500, 'verbose': 1,
'callbacks': [keras.callbacks.EarlyStopping(monitor='val_mae', mode='min', verbose=0, patience=75, restore_best_weights=True)]}
learning_rate = 0.002
load = False # load an already saved model
if load:
models_dict = {}
for room in rooms:
models_dict[room] = keras.load_model(f'{modeldir}/{model_arch_name}/{room}')
else:
models_dict = defaultdict(lambda: model_arch(learning_rate).model)
for training_room in rooms:
print(training_room)
models_dict[training_room] = model_arch(learning_rate).model
ytrain = pd.concat([y_train[training_room][anchor]['37'] for anchor in anchors], axis=1)
yval = pd.concat([y_val[training_room][anchor]['37'] for anchor in anchors], axis=1)
models_dict[training_room].fit(x_train[training_room], ytrain,
validation_data=(x_val[training_room], yval),
**fit_params)
save = False # save the model
if save:
for room in rooms:
models_dict[room].save(f'{modeldir}/{model_arch_name}/{room}')
```
# Predictions
```
# generate AoA and position predictions as well as mean euclidean distance error
# and AoA mean absolute error for all training and testing room combinations
preds, true_pos = make_predictions(x_test, y_test, models_dict, training_rooms=rooms,
testing_rooms=rooms + concrete_rooms + other_scenarios,
test_points=test_points, anchor_data=anchor_data)
def default_to_regular(d):
if isinstance(d, (defaultdict, dict)):
d = {k: default_to_regular(v) for k, v in d.items()}
return d
# save the predictions dictionary
save_preds = False
if save_preds:
results_path = f'{preds_dir}/preds_{model_arch_name}.npy'
np.save(results_path, default_to_regular(preds))
# load the predictions dictionary
load_preds = False
if load_preds:
preds = np.load(f'{preds_dir}/preds_{model_arch_name}.npy', allow_pickle=True)[()]
```
## PDDA
```
#produce pdda predictions
pdda_res = generate_pdda_preds(data, rooms + concrete_rooms + other_scenarios, test_points, anchor_data)
```
# Results
```
#produce pos maes heatmap
posHeatmapXY(preds['pos_maes'][:,:7], pdda_res['pos_maes'][:7])
#produce error per point plot
spatial_plot(preds['pos_preds']['testbench_01_furniture_low']['testbench_01_furniture_high'], true_pos, testing_room = 'testbench_01_furniture_high', mode = 'xy',vmax = 3, cmap = 'PuBu')
```
| github_jupyter |
In this example, we will use tensorflow v1 (version 1.15) to create a simple MLP model, and transfer the application to Cluster Serving step by step.
This tutorial is recommended for Tensorflow v1 user only. If you are not Tensorflow v1 user, the keras tutorial [here](#keras-to-cluster-serving-example.ipynb) is more recommended.
### Original Tensorflow v1 Application
```
import tensorflow as tf
tf.__version__
```
We first define the Tensorflow graph, and create some data.
```
g = tf.Graph()
with g.as_default():
# Graph Inputs
features = tf.placeholder(dtype=tf.float32,
shape=[None, 2], name='features')
targets = tf.placeholder(dtype=tf.float32,
shape=[None, 1], name='targets')
# Model Parameters
weights = tf.Variable(tf.zeros(shape=[2, 1],
dtype=tf.float32), name='weights')
bias = tf.Variable([[0.]], dtype=tf.float32, name='bias')
# Forward Pass
linear = tf.add(tf.matmul(features, weights), bias, name='linear')
ones = tf.ones(shape=tf.shape(linear))
zeros = tf.zeros(shape=tf.shape(linear))
prediction = tf.where(condition=tf.less(linear, 0.),
x=zeros,
y=ones,
name='prediction')
# Backward Pass
errors = targets - prediction
weight_update = tf.assign_add(weights,
tf.reshape(errors * features, (2, 1)),
name='weight_update')
bias_update = tf.assign_add(bias, errors,
name='bias_update')
train = tf.group(weight_update, bias_update, name='train')
saver = tf.train.Saver(name='saver')
import numpy as np
x_train, y_train = np.array([[1,2],[3,4],[1,3]]), np.array([1,2,1])
x_train.shape, y_train.shape
```
### Export TensorFlow SavedModel
Then, we train the graph and in the `with tf.Session`, we save the graph to SavedModel. The detailed code is following, and we could see the prediction result is `[1]` with input `[1,2]`.
```
with tf.Session(graph=g) as sess:
sess.run(tf.global_variables_initializer())
for epoch in range(5):
for example, target in zip(x_train, y_train):
feed_dict = {'features:0': example.reshape(-1, 2),
'targets:0': target.reshape(-1, 1)}
_ = sess.run(['train'], feed_dict=feed_dict)
w, b = sess.run(['weights:0', 'bias:0'])
print('Model parameters:\n')
print('Weights:\n', w)
print('Bias:', b)
saver.save(sess, save_path='perceptron')
pred = sess.run('prediction:0', feed_dict={features: x_train})
print(pred)
# in this session, save the model to savedModel format
inputs = dict([(features.name, features)])
outputs = dict([(prediction.name, prediction)])
inputs, outputs
tf.saved_model.simple_save(sess, "/tmp/mlp_tf1", inputs, outputs)
```
### Deploy Cluster Serving
After model prepared, we start to deploy it on Cluster Serving.
First install Cluster Serving
```
! pip install analytics-zoo-serving
import os
! mkdir cluster-serving
os.chdir('cluster-serving')
! cluster-serving-init
! tail wget-log
# if you encounter slow download issue like above, you can just use following command to download
# ! wget https://repo1.maven.org/maven2/com/intel/analytics/zoo/analytics-zoo-bigdl_0.12.1-spark_2.4.3/0.9.0/analytics-zoo-bigdl_0.12.1-spark_2.4.3-0.9.0-serving.jar
# if you are using wget to download, or get "analytics-zoo-xxx-serving.jar" after "ls", please call mv *serving.jar zoo.jar after downloaded.
# After initialization finished, check the directory
! ls
# Call mv *serving.jar zoo.jar as mentioned above
! mv *serving.jar zoo.jar
! ls
```
We config the model path in `config.yaml` to following (the detail of config is at [Cluster Serving Configuration](https://github.com/intel-analytics/analytics-zoo/blob/master/docs/docs/ClusterServingGuide/ProgrammingGuide.md#2-configuration))
```
## Analytics-zoo Cluster Serving
model:
# model path must be provided
path: /tmp/mlp_tf1
! head config.yaml
```
### Start Cluster Serving
Cluster Serving requires Flink and Redis installed, and corresponded environment variables set, check [Cluster Serving Installation Guide](https://github.com/intel-analytics/analytics-zoo/blob/master/docs/docs/ClusterServingGuide/ProgrammingGuide.md#1-installation) for detail.
Flink cluster should start before Cluster Serving starts, if Flink cluster is not started, call following to start a local Flink cluster.
```
! $FLINK_HOME/bin/start-cluster.sh
```
After configuration, start Cluster Serving by `cluster-serving-start` (the detail is at [Cluster Serving Programming Guide](https://github.com/intel-analytics/analytics-zoo/blob/master/docs/docs/ClusterServingGuide/ProgrammingGuide.md#3-launching-service))
```
! cluster-serving-start
```
### Prediction using Cluster Serving
Next we start Cluster Serving code at python client.
```
from zoo.serving.client import InputQueue, OutputQueue
input_queue = InputQueue()
# Use async api to put and get, you have pass a name arg and use the name to get
arr = np.array([1,2])
input_queue.enqueue('my-input', t=arr)
output_queue = OutputQueue()
prediction = output_queue.query('my-input')
# Use sync api to predict, this will block until the result is get or timeout
prediction = input_queue.predict(arr)
prediction
```
The `prediction` result would be the same as using Tensorflow.
This is the end of this tutorial. If you have any question, you could raise an issue at [Analytics Zoo Github](https://github.com/intel-analytics/analytics-zoo/issues).
| github_jupyter |
# User-defined prior distribution
```
from pyunfold import iterative_unfold
from pyunfold.callbacks import Logger
import numpy as np
np.random.seed(2)
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
sns.set_context(context='poster')
plt.rcParams['figure.figsize'] = (10, 8)
plt.rcParams['lines.markeredgewidth'] = 2
```
## Example dataset
We'll generate the same example dataset that is used in the [Getting Started tutorial](tutorial.ipynb), i.e. a Gaussian sample that is smeared by some noise.
```
# True distribution
num_samples = int(1e5)
true_samples = np.random.normal(loc=10.0, scale=4.0, size=num_samples)
bins = np.linspace(0, 20, 21)
num_causes = len(bins) - 1
data_true, _ = np.histogram(true_samples, bins=bins)
# Observed distribution
random_noise = np.random.normal(loc=0.3, scale=0.5, size=num_samples)
observed_samples = true_samples + random_noise
data_observed, _ = np.histogram(observed_samples, bins=bins)
data_observed_err = np.sqrt(data_observed)
# Efficiencies
efficiencies = np.ones_like(data_observed, dtype=float)
efficiencies_err = np.full_like(efficiencies, 0.1, dtype=float)
# Response matrix
response_hist, _, _ = np.histogram2d(observed_samples, true_samples, bins=bins)
response_hist_err = np.sqrt(response_hist)
# Normalized response
column_sums = response_hist.sum(axis=0)
normalization_factor = efficiencies / column_sums
response = response_hist * normalization_factor
response_err = response_hist_err * normalization_factor
```
We can see what the true and observed distributions look like for this example dataset
```
fig, ax = plt.subplots(figsize=(10, 8))
ax.step(np.arange(len(data_true)), data_true, where='mid', lw=3,
alpha=0.7, label='True distribution')
ax.step(np.arange(len(data_observed)), data_observed, where='mid', lw=3,
alpha=0.7, label='Observed distribution')
ax.set(xlabel='Cause bins', ylabel='Counts')
ax.legend()
plt.show()
```
as well as the normalized response matrix
```
fig, ax = plt.subplots(figsize=(10, 8))
im = ax.imshow(response, origin='lower')
cbar = plt.colorbar(im, label='$P(E_i|C_{\mu})$')
ax.set(xlabel='Cause bins', ylabel='Effect bins',
title='Normalized response matrix')
plt.show()
```
## Custom Priors
The default initial prior used in PyUnfold is the uniform prior (i.e. each cause bin is given an equal weighting). We can test other priors by providing a normalized distribution via the `prior` parameter in the `iterative_unfold` function.
Several convenience functions for calculating commonly used prior distributions exist in the `pyunfold.priors` module. However, _any_ normalized array-like object (i.e. the items sum to 1) can be passed to `prior` and used as the initial prior in an unfolding.
One commonly used prior is the non-informative Jeffreys' prior. The analytic form of this prior is given by
$$
P(C_{\mu})^{\text{Jeffreys}} = \frac{1}{\log \left( C_{\text{max}} / C_{\text{min}}\right) \, C_{\mu}}
$$
where $C_{\text{max}}$ and $C_{\text{min}}$ are the maximum and minimum possible cause values, and $C_{\mu}$ is the value of the $\mu$th cause bin. Here we'll assume that the cause range covers three orders of magnitude, that is $C_{\mu} \in [1, 10^3]$.
```
from pyunfold.priors import jeffreys_prior, uniform_prior
# Cause limits
cause_lim = np.logspace(0, 3, num_causes)
# Uniform and Jeffreys' priors
uni_prior = uniform_prior(num_causes)
jeff_prior = jeffreys_prior(cause_lim)
```
The `uniform_prior` and `jeffreys_prior` functions calculate their corresponding prior distributions and return NumPy `ndarrays` containing these distributions. For more information about these functions, see the [priors API documentation](../api.rst#priors).
```
print(type(uni_prior))
print('uni_prior = {}'.format(uni_prior))
print(type(jeff_prior))
print('jeff_prior = {}'.format(jeff_prior))
fig, ax = plt.subplots(figsize=(10, 8))
ax.step(cause_lim, jeff_prior, where='mid', lw=3,
alpha=0.7, label='Jeffreys')
ax.step(cause_lim, uni_prior, where='mid', lw=3,
alpha=0.7, label='Uniform')
ax.set(xlabel='Cause Values $C_{\mu}$', ylabel='$P(C_{\mu})$',
title='Priors')
plt.xscale('log')
plt.yscale('log')
plt.legend()
plt.show()
```
## Unfolding
Now we can run the unfolding with the Jeffreys' prior and compare to the default uniform prior as well as the true cause distribution.
```
print('Running with uniform prior...')
unfolded_uniform = iterative_unfold(data=data_observed,
data_err=data_observed_err,
response=response,
response_err=response_err,
efficiencies=efficiencies,
efficiencies_err=efficiencies_err,
ts='ks',
ts_stopping=0.01,
callbacks=[Logger()])
print('\nRunning with Jeffreys prior...')
unfolded_jeffreys = iterative_unfold(data=data_observed,
data_err=data_observed_err,
response=response,
response_err=response_err,
efficiencies=efficiencies,
efficiencies_err=efficiencies_err,
prior=jeff_prior,
ts='ks',
ts_stopping=0.01,
callbacks=[Logger()])
bin_midpoints = (bins[1:] + bins[:-1]) / 2
fig, ax = plt.subplots(figsize=(10, 8))
ax.hist(true_samples, bins=bins, histtype='step', lw=3,
alpha=0.7,
label='True distribution')
ax.errorbar(bin_midpoints, unfolded_uniform['unfolded'],
yerr=unfolded_uniform['sys_err'],
alpha=0.7,
elinewidth=3,
capsize=4,
ls='None', marker='.', ms=10,
label='Unfolded - Uniform Prior')
ax.errorbar(bin_midpoints, unfolded_jeffreys['unfolded'],
yerr=unfolded_jeffreys['sys_err'],
alpha=0.7,
elinewidth=3,
capsize=4,
ls='None', marker='.', ms=10,
label='Unfolded - Jeffreys Prior')
ax.set(xlabel='Cause bins', ylabel='Counts')
plt.legend()
plt.show()
```
The unfolded distributions are consistent with each other as well as with the true distribution!
Thus, our results are robust with respect to these two **smooth** initial priors.
For information about how un-smooth priors can affect an unfolding see the [smoothing via spline regularization example](regularization.ipynb).
| github_jupyter |
```
%matplotlib inline
%load_ext autoreload
%autoreload 2
import os
import sys
sys.path.append("..")
sys.path.append("../..")
import numpy as np
import pandas as pd
```
### Occupancy data
```
## [from examples/examples.py]
from download import download_all
## The path to the test data sets
FIXTURES = os.path.join(os.getcwd(), "data")
## Dataset loading mechanisms
datasets = {
"credit": os.path.join(FIXTURES, "credit", "credit.csv"),
"concrete": os.path.join(FIXTURES, "concrete", "concrete.csv"),
"occupancy": os.path.join(FIXTURES, "occupancy", "occupancy.csv"),
"mushroom": os.path.join(FIXTURES, "mushroom", "mushroom.csv"),
}
def load_data(name, download=True):
"""
Loads and wrangles the passed in dataset by name.
If download is specified, this method will download any missing files.
"""
# Get the path from the datasets
path = datasets[name]
# Check if the data exists, otherwise download or raise
if not os.path.exists(path):
if download:
download_all()
else:
raise ValueError((
"'{}' dataset has not been downloaded, "
"use the download.py module to fetch datasets"
).format(name))
# Return the data frame
return pd.read_csv(path)
# Load the classification data set
data = load_data('occupancy')
print(len(data))
data.head()
# Specify the features of interest and the classes of the target
features = ["temperature", "relative humidity", "light", "C02", "humidity"]
classes = ['unoccupied', 'occupied']
# Searching the whole dataset takes a while (15 mins on my mac)...
# For demo purposes, we reduce the size
X = data[features].head(2000)
y = data.occupancy.head(2000)
```
### Parameter projection
* Because the visualizer only displays results across two parameters, we need some way of reducing the dimension to 2.
* Our approach: for each value of the parameters of interest, display the _maximum_ score across all the other parameters.
Here we demo the `param_projection` utility function that does this
```
from sklearn.model_selection import GridSearchCV
from sklearn.svm import SVC
from yellowbrick.gridsearch.base import param_projection
# Fit a vanilla grid search... these are the example parameters from sklearn's gridsearch docs.
svc = SVC()
grid = [{'kernel': ['rbf'], 'gamma': [1e-3, 1e-4], 'C': [1, 10, 100, 1000]},
{'kernel': ['linear'], 'C': [1, 10, 100, 1000]}]
gs = GridSearchCV(svc, grid, n_jobs=4)
%%time
gs.fit(X, y)
```
As of Scikit-learn 0.18, `cv_results` has replaced `grid_scores` as the grid search results format
```
gs.cv_results_
```
Demo the use of param_projection... It identifies the unique values of the the two parameter values and gets the best score for each (here taking the max over `gamma` values)
```
param_1 = 'C'
param_2 = 'kernel'
param_1_vals, param2_vals, best_scores = param_projection(gs.cv_results_, param_1, param_2)
param_1_vals, param2_vals, best_scores
```
### GridSearchColorPlot
This visualizer wraps the GridSearchCV object and plots the values obtained from `param_projection`.
```
from yellowbrick.gridsearch import GridSearchColorPlot
gs_viz = GridSearchColorPlot(gs, 'C', 'kernel')
gs_viz.fit(X, y).show()
gs_viz = GridSearchColorPlot(gs, 'kernel', 'C')
gs_viz.fit(X, y).show()
gs_viz = GridSearchColorPlot(gs, 'C', 'gamma')
gs_viz.fit(X, y).show()
```
If there are missing values in the grid, these are filled with a hatch (see https://stackoverflow.com/a/35905483/7637679)
```
gs_viz = GridSearchColorPlot(gs, 'kernel', 'gamma')
gs_viz.fit(X, y).show()
```
Choose a different metric...
```
gs_viz = GridSearchColorPlot(gs, 'C', 'kernel', metric='mean_fit_time')
gs_viz.fit(X, y).show()
```
### Quick Method
Because grid search can take a long time and we may want to interactively cut the results a few different ways, by default the quick method assumes that the GridSearchCV object is **already fit** if no X data is passed in.
```
from yellowbrick.gridsearch import gridsearch_color_plot
%%time
# passing the GridSearchCV object pre-fit
gridsearch_color_plot(gs, 'C', 'kernel')
%%time
# trying a different cut across parameters
gridsearch_color_plot(gs, 'C', 'gamma')
%%time
# When we provide X, the `fit` method will call fit (takes longer)
gridsearch_color_plot(gs, 'C', 'kernel', X=X, y=y)
%%time
# can also choose a different metric
gridsearch_color_plot(gs, 'C', 'kernel', metric='mean_fit_time')
```
### Parameter errors
Bad param values
```
gs_viz = GridSearchColorPlot(gs, 'foo', 'kernel')
gs_viz.fit(X, y).show()
gs_viz = GridSearchColorPlot(gs, 'C', 'foo')
gs_viz.fit(X, y).show()
```
Bad metric option
```
gs_viz = GridSearchColorPlot(gs, 'C', 'kernel', metric='foo')
gs_viz.fit(X, y).show()
```
Metric option exists in cv_results but is not numeric -> not valid
```
gs_viz = GridSearchColorPlot(gs, 'C', 'kernel', metric='param_kernel')
gs_viz.fit(X, y).show()
```
| github_jupyter |
# Molecular dynamics in SchNetPack (experimental)
In the [previous tutorial](tutorial_03_force_models.ipynb) we have covered how to train machine learning models on molecular forces and use them for basic molecular dynamics (MD) simulations with the SchNetPack ASE interface.
All these simulations can also be carried out using the native MD package available in SchNetPack.
The main ideas behind integrating MD functionality directly into SchNetPack are:
- improve performance by reducing the communication overhead between ML models and the MD code and adding the option to use GPUs
- adding extended functionality, such as sampling algorithms and ring polymer MD
- providing a modular MD environment for easy development and interfacing
In the following, we first introduce the general structure of the SchNetPack-MD package.
Then the simulation from the previous tutorial will be used as an example to demonstrate how to implement basic MD with SchNetPack-MD.
Having done so, we will cover a few advanced simulation techniques, such as ring polymer MD.
Finally, we will show how all of these different simulations can be accessed via an input file.
## Getting started
Before we can begin with the main tutorial, some setup is required.
First, we generate a directory for holding our simulations:
```
import os
md_workdir = 'mdtut'
# Gnerate a directory of not present
if not os.path.exists(md_workdir):
os.mkdir(md_workdir)
```
Since we want to run MD simulations, we need a SchNetPack model trained on forces and a molecular structure as a starting point.
In principle, we could use the ethanol model and structure generated in the previous tutorial.
However, the model trained in the force tutorial was only intended as a demonstration and is not the most accurate.
Instead, we will use a sample ethanol structure, as well as a fully converged SchNet model of ethanol provided with the data used for testing SchNetPack for this tutorial:
```
import schnetpack as spk
# Get the parent directory of SchNetPack
spk_path = os.path.abspath(os.path.join(os.path.dirname(spk.__file__), '../..'))
# Get the path to the test data
test_path = os.path.join(spk_path, 'tests/data')
# Load model and structure
model_path = os.path.join(test_path, 'test_md_model.model')
molecule_path = os.path.join(test_path, 'test_molecule.xyz')
```
## MD in SchNetPack
In general, a MD code needs to carry out several core tasks during each simulation step.
It has to keep track of the positions $\mathbf{R}$ and momenta $\mathbf{p}$ of all nuclei, compute the forces $\mathbf{F}$ acting on the nuclei and use the latter to integrate Newton's equations of motion.
<img src="tutorials_figures/md_flowchart.svg" width="200" style="padding: 5px 15px; float: left;">
The overall workflow used in the SchNetPack MD package is sketched in the figure to the left.
As can be seen, the various tasks are distibuted between different modules.
The `System` class contains all information on the present state of the simulated system (e.g. nuclear positons and momenta).
This is a good point to mention, that internally the MD package uses a special internal unit system for all properties.
The basic units are nanometers for length, kilojoule per mol for energies and Dalton for mass. Other units are derived
from these quantities.
The `Integrator` computes the positions and momenta of the next step and updates the state of the system accordingly.
In order to carry out this update, the nuclear forces are required.
These are computed with a `Calculator`, which takes the positions of atoms and returns the corresponding forces.
Typically, the `Calculator` consists of a previously trained machine learning model.
All these modules are linked together in the `Simulator` class, which contains the main MD loop and calls the three previous modules in the correct order.
We will now describe the different components of the MD package in more detail and give an example of how to set up a short MD simulation of an ethanol molecule.
### System
As stated previously, `System` keeps track of the state of the simulated system and contains the atomic positions $\mathbf{R}$ and momenta $\mathbf{p}$, but also e.g. atom types and computed molecular properties.
A special property of SchNetPack-MD is the use of multdimensional tensors to store the system information (using the `torch.Tensor` class).
This makes it possible to make full use of vectorization and e.g. simulate several different molecules as well as different replicas of a molecule in a single step.
The general shape of these system tensors is $N_\textrm{replicas} \times N_\textrm{molecules} \times N_\textrm{atoms} \times \ldots$, where the first dimension is the number of replicas of the same molecule (e.g. for ring polymer MD), the second runs over the different molecules simulated (e.g. fragments of different size for sampling) and the third over the maximum number of atoms present in any system.
In order to initialize a `System`, first the number of replicas needs to be given. Here, we want to perform a standard MD and $N_\mathrm{replicas}=1$.
In addition, one can specify the device used for the computation.
Afterwards, the molecules which should be simulated need to be loaded.
These can be read directly from a XYZ-file via the `load_molecules_from_xyz` function.
$N_\mathrm{molecules}$ is determined automatically based on the number of structures found in this file.
In our present case, the loaded files containes the structure of a single ethanol.
```
from schnetpack.md import System
import torch
# Check if a GPU is available and use a CPU otherwise
if torch.cuda.is_available():
md_device = "cuda"
else:
md_device = "cpu"
# Number of molecular replicas
n_replicas = 1
# Initialize the system
md_system = System(n_replicas, device=md_device)
# Load the structure
md_system.load_molecules_from_xyz(molecule_path)
```
Right now, all system momenta are set to zero.
For practical purposes, one usually wants to draw the momenta from a distribution corresponding to a certain temperature.
This can be done via an `Initializer`, which takes the temperature in Kelvin as an input. For this example, we use a Maxwell—Boltzmann initialization:
```
from schnetpack.md import MaxwellBoltzmannInit
system_temperature = 300 # Kelvin
# Set up the initializer
md_initializer = MaxwellBoltzmannInit(
system_temperature,
remove_translation=True,
remove_rotation=True)
# Initialize momenta of the system
md_initializer.initialize_system(md_system)
```
Here, we have also removed all translational and rotational components of the momenta via the appropriate keyword.
### Integrator
Having set up the system in such a manner, one needs to specify how the equations of motion should be propagated.
Currently, there are two integration schemes implemented in SchNetPack:
- a Velocity Verlet integrator which evolves the system in a purely classical manner and
- a ring polymer integrator which is able to model a certain degree of nuclear quantum effects
For demonstration purposes, we will first focus on a purely classical MD using the Velocity Verlet algorithm.
An example on how to use ring polymer MD in SchNetPack and potential benefits will be given later in the tutorial.
To initialize the integrator, one has to specify the length of the timestep $\Delta t$ used for integration in units of femtoseconds.
A common value for classical MD is $\Delta t = 0.5$ fs.
```
from schnetpack.md.integrators import VelocityVerlet
time_step = 0.5 # fs
# Setup the integrator
md_integrator = VelocityVerlet(time_step)
```
### Calculator
The only ingredient missing for simulating our system is a `Calculator` to compute molecular forces and other properties.
A `Calculator` can be thought of as an interface between a computation method (e.g. a machine learning model) and the MD code in SchNetPack.
SchNetPack comes with several predefined calculators and also offers the possibility to implement custom calculators.
Right now, we are only interested in using a model trained with SchNetPack, hence we use the `SchnetPackCalculator`.
First, we have to load the stored model with Torch and move it to the computation device defined before.
To initialize the `SchnetPackCalculator`, we have to pass it the loaded model.
Similar as for the ASE interface in the [last tutorial](tutorial_03_force_models.ipynb), we have to tell the calculator which properties to compute, how the forces are called in the output.
Since the whole SchNetPack-MD package uses consistent internal units, it is also necessary to specify which units the calculator expects for the positions (`position_conversion`) and which units it uses for the returned forces (`force_conversion`).
For the first two points, we can make use of the SchNetPack properties definitions. With regards to units, the current calculator uses Å for positions and kcal/mol/Å for the forces. The conversion factors can either be given as a number or as a string.
```
from schnetpack.md.calculators import SchnetPackCalculator
from schnetpack import Properties
# Load the stored model
md_model = torch.load(model_path, map_location=md_device).to(md_device)
# Generate the calculator
md_calculator = SchnetPackCalculator(
md_model,
required_properties=[Properties.energy, Properties.forces],
force_handle=Properties.forces,
position_conversion='A',
force_conversion='kcal/mol/A'
)
```
### Simulator (bringing it all together)
With our molecular system, a machine learning calculator for the forces and an integrator at hand, we are almost ready carry out MD simulations.
The last step is to pass all these ingredients to a `Simulator`.
The `Simulator` performs the actual MD simulations, looping over a series of time steps and calling the individual modules in the right order:
```
from schnetpack.md import Simulator
md_simulator = Simulator(md_system, md_integrator, md_calculator)
```
To carry out a simulation, one needs to call the `simulate` function with an integer argument specifying the number of desired simulation steps.
For example, a MD simulation of our ethanol molecule for 100 time steps (50 fs) can be done via:
```
n_steps = 100
md_simulator.simulate(n_steps)
```
Since the `Simulator` keeps track of the state of the dynamics and the system, we can call it repeatetly to get longer trajectories.
```
md_simulator.simulate(n_steps)
```
The actual number of steps is stored in the `step` variable of the `Simulator` class.
```
print("Total number of steps:", md_simulator.step)
```
Although we are now able to run a full-fledged MD simulation, there is one major problem with the current setup:
we do not collect any information during the simulation, such as nuclear positions.
This means, that we currently have no way of analyzing what happened during the MD trajectory.
This — and many other things — can be done in the SchNetPack-MD package using so-called simulation hooks.
## Simulation hooks
Simulation hooks follow the same concept as the hooks used in the SchNetPack `Trainer` class covered [previously](tutorial_02_qm9.ipynb).
They can be thought as instructions for the `Simulator`, which are performed at certain points during each MD step.
Simulation hooks can be used to tailor a simulation to ones need, contributing to the customizability of the SchNetPack-MD package.
<img src="tutorials_figures/integrator.svg" width="370" style="padding: 20px 20px; float: left">
The diagram to the left shows how a single MD step of the `Simulator` is structured in detail and at which points hooks can be applied.
Depending on which time they are called and which actions they encode, simulation hooks can achieve a wide range of tasks.
If they are introduced before and after each integration half-step, they can e.g. be used to control the temperature of the system in the form of thermostats.
When acting directly after the computation of the forces done by the `Calculator`, simulation hooks can be used to control sampling.
At this point, enhanced sampling schemes such as metadynamics and accelerated MD can be implemented, which modify the forces and potential energies of the system.
It is also possible to introduce active learning for automatically generating machine learning models in this way.
Finally, when called after the second integration step, simulation hooks can be used to collect and store information on the system, which can then be used for analysis.
Multiple hooks can be passed to a simulator at any time, which makes it possible to control a simulation in various manners.
In the following, we will demonstrate how to apply a thermostat to the above simulation and how data collection can be done in SchNetPack.
### Adding a Thermostat
As mentioned in the [force tutorial](tutorial_03_force_models.ipynb), thermostats are used to keep the fluctuations of the kinetic energy of a system (temperature) close to a predefined average.
Simulations employing thermostats are referred to as canonical ensemble or $NVT$ simulations, since they keep the number of particles $N$, the volume $V$ and the average temperature $T$ constant.
Last time, we used a Langevin thermostat to regulate the temperature of our simulation.
This thermostat (and many others) is also available in SchNetPack and can be used via
```
from schnetpack.md.simulation_hooks import thermostats
# Set temperature and thermostat constant
bath_temperature = 300 # K
time_constant = 100 # fs
# Initialize the thermostat
langevin = thermostats.LangevinThermostat(bath_temperature, time_constant)
```
In case of the Langevin thermostat, a bath temperature (in Kelvin) and a time constant (in fs) have to be provided.
The first regulates the temperature the system is kept at, the second how fast the thermostat adjusts the temperature.
In order to speed up equilibration, we use a more agressive thermostatting schedule by with a time constant of 100 fs.
Finally, we begin collecting the simulation hooks we want to pass to the simulator.
```
simulation_hooks = [
langevin
]
```
### Collecting Data and storing Checkpoints
The primary way to store simulation data in the SchNetPack-MD package is via the `FileLogger` class.
A `FileLogger` collects data during the MD and stores it to a database in HDF5 format.
The type of data to be collected is specified via so-callled `DataStreams`, which are passed to the `FileLogger`.
The data streams currently available in SchNetPack are:
- `MoleculeStream`: Stores positions and velocities during all simulation steps
- `PropertyStream`: Stores all properties predicted by the calculator
- `SimulationStream`: Collects information on the kinetic energy and system temperature (this can be also done via postprocessing when using the `MoleculeStream`)
By default, the `MoleculeStream` and `PropertyStream` are used.
To reduce overhead due to writing to disk, the `FileLogger` first collects information for a certain number of steps into a buffer, which it then writes to the database at once.
The `FileLogger` is initialized by specifying the name of the target database, the size of the buffer and which data to store (in form of the respective data streams):
```
from schnetpack.md.simulation_hooks import logging_hooks
# Path to database
log_file = os.path.join(md_workdir, 'simulation.hdf5')
# Size of the buffer
buffer_size = 100
# Set up data streams to store positions, momenta and all properties
data_streams = [
logging_hooks.MoleculeStream(),
logging_hooks.PropertyStream(),
]
# Create the file logger
file_logger = logging_hooks.FileLogger(
log_file,
buffer_size,
data_streams=data_streams
)
# Update the simulation hooks
simulation_hooks.append(file_logger)
```
In general, it is also a good idea to store checkpoints of the system and simulation state at regular intervals.
Should something go wrong with the simulation, these can be used to restart the simulation from the last stored point.
In addition, these checkpoints can also be used to only initialize the `System`.
This is e.g. useful for equilibrating simulations with different thermostats.
Storing checkpoints can be done with the `Checkpoint` hook, which takes a file the data is stored to and the frequency a checkpoint is generated:
```
#Set the path to the checkpoint file
chk_file = os.path.join(md_workdir, 'simulation.chk')
# Create the checkpoint logger
checkpoint = logging_hooks.Checkpoint(chk_file, every_n_steps=100)
# Update the simulation hooks
simulation_hooks.append(checkpoint)
```
### Adding Hooks and Running the Simulation
With all simulation hooks created and collected in `simulation_hooks`, we can finally build our updated simulator.
This is done exactly the same way as above, with the difference that now also the hooks are specififed.
```
md_simulator = Simulator(md_system, md_integrator, md_calculator, simulator_hooks=simulation_hooks)
```
We can now use the simulator to run a MD trajectory of our ethanol. Here, we run for 20000 steps, which are 10 ps.
This should take approximately 5 minutes on a notebook GPU.
```
n_steps = 20000
md_simulator.simulate(n_steps)
```
The tutorial directory should now contain two files:
- `simulation.hdf5`, which holds the collected data and
- `simulation.chk` containing the last checkpoint.
## Reading HDF5 outputs
We will now show, how to access the HDF5 files generated during the simulation.
For this purpose, SchNetPack comes with a `HDF5Loader`, which can be used to extract the data by giving the path to the simulation output (`mdtut/simulation.hdf5`).
```
from schnetpack.md.utils import HDF5Loader
data = HDF5Loader(log_file)
```
Extracted data is stored in the `properties` dictionary of the `HDF5Loader` and can be accessed with the `get_property` function.
`get_property` requires the name of the property and optionally the index of the molecule and replica for which the data should be extracted.
By default, it extracts the first molecule (`mol_idx=0`) and averages over all replicas if more than one are present.
Neither is relavant for our current simulation.
Right now, we can access the following entries, all of which should be self explaining and correspond to the standard SchNetPack `Properties` and `Structure` keys:
```
for prop in data.properties:
print(prop)
```
We can now e.g. have a look at the potential energies.
```
%matplotlib notebook
import matplotlib.pyplot as plt
from schnetpack.md.utils import MDUnits
# Get potential energies and check the shape
energies = data.get_property(Properties.energy)
print('Shape:', energies.shape)
# Get the time axis
time_axis = np.arange(data.entries)*data.time_step / MDUnits.fs2internal # in fs
# Plot the energies
plt.figure()
plt.plot(time_axis, energies)
plt.ylabel('E [kcal/mol]')
plt.xlabel('t [fs]')
plt.tight_layout()
plt.show()
```
The `HDF5Loader` also offers access to functions for computing some derived properties, such as the kinetic energy (`get_kinetic_energy`) and the temperature (`get_temperature`).
```
import numpy as np
def plot_temperature(data):
# Read the temperature
temperature = data.get_temperature()
# Compute the cumulative mean
temperature_mean = np.cumsum(temperature) / (np.arange(data.entries)+1)
# Get the time axis
time_axis = np.arange(data.entries)*data.time_step / MDUnits.fs2internal # in fs
plt.figure(figsize=(8,4))
plt.plot(time_axis, temperature, label='T')
plt.plot(time_axis, temperature_mean, label='T (avg.)')
plt.ylabel('T [K]')
plt.xlabel('t [fs]')
plt.legend()
plt.tight_layout()
plt.show()
plot_temperature(data)
```
As can be seen, the system requires a initial period for equilibration.
This is relevant for simulations, as ensemble properties are typically only computed for fully equilibrated systems.
In SchNetPack, an apropriate analysis can be done in different ways.
The checkpoint file of the equilibrated system can be used as a starting point for a production simulation.
The easier way, however, is to reject the initial part of the trajectory and only consider the equilibrated system for analysis.
This can be done by specifying the number of steps to skip in the `HDF5Loader`.
Here, we skip the first half (25 000 steps) of the trajectory.
In general, the equilibration period strongly depends on system size and the thermostat settings.
```
equilibrated_data = HDF5Loader(log_file, skip_initial=10000)
```
We can easily see, that only the later part of the simulation is now considered by plotting the data once again:
```
plot_temperature(equilibrated_data)
```
It should be mentioned at this point, that the HDF5 datafile uses a special convention for units.
For internal quantities (e.g. positions, velocities and kinetic energy), schnetpack internal units are used
(mass=Da, length=nm, energy=kJ/mol).
The only exception are temperatures, which are given in units of Kelvin for convenience.
For all properties computed by the `Calculator` the original unit is used, unless a conversion factor is specified during initialization.
This means, that the energies and forces collected here have units of kcal/mol and kcal/mol/Å.
### Vibrational spectra
While curves of temperatures might be nice to look at, one is usually interested in different quantities when running a MD simulation.
One example are vibrational spectra, which give information on which vibrations are active in a molecule.
SchNetPack provides the module `schnetpack.md.utils.spectra`, which provides different classes to compute vibrational spectra directly from the HDF5 files.
These implementations use several tricks to improve efficiency and accuracy.
Currently, power spectra (`PowerSpectrum`) and infrared (IR) spectra (`IRSpectrum`) are available.
Here, we will compute the power spectrum from our ethanol simulation:
```
from schnetpack.md.utils import PowerSpectrum
# Intialize the spectrum
spectrum = PowerSpectrum(equilibrated_data, resolution=4096)
# Compute the spectrum for the first molecule (default)
spectrum.compute_spectrum(molecule_idx=0)
```
The `resolution` keyword specifies, how finely the peaks in the spectrum are resolved.
`PowerSpectrum` also computes the effective resolution in inverse centimeters, as well as the spectral range.
For molecules, one is usually interested in frequencies up to 4000 cm<sup>-1</sup>, which we will use to restrict the plotting area.
```
# Get frequencies and intensities
freqencies, intensities = spectrum.get_spectrum()
# Plot the spectrum
plt.figure()
plt.plot(freqencies, intensities)
plt.xlim(0,4000)
plt.ylim(0,100)
plt.ylabel('I [a.u.]')
plt.xlabel('$\omega$ [cm$^{-1}$]')
plt.show()
```
The spectrum shows several typical vibrational bands for ethanol (which can be checked with [experimental tables available online](https://chem.libretexts.org/Ancillary_Materials/Reference/Reference_Tables/Spectroscopic_Parameters/Infrared_Spectroscopy_Absorption_Table)).
For example, the peak close to 3700 cm<sup>-1</sup> stems from the bond vibrations of the OH-group.
The bond vibrations of the CH<sub>3</sub> and CH<sub>2</sub> groups are clustered around 3000 cm<sup>-1</sup> and the correponding bending vinrations can be seen at 1400 cm<sup>-1</sup>.
In general, computed vibrational spectra serve as a good check for the validity of a machine learning potential.
One important fact should be noted at this point: the spectrum computed here is a power spectrum, representing the vibrational density of states.
It gives information on **all** vibrational modes which **can** potentially be active in an experimental spectrum.
Hence, it can help in identifying which motions give rise to which experimental bands.
However, depending on the experiment, only a subset of the peaks of the power spectrum can be active and the intensities can vary dramatically.
As such, a power spectrum only serves as a poor stand in for simulating e.g. Raman or IR spectra.
Using SchNetPack, it is also possible to model IR spectra, by training a model on dipoles in addition to forces.
Simulations can then be done in the same manner as above and the corresponding IR spectra can be obtained using `IRSpectrum` instead of `PowerSpectrum`.
## Restarting simulations
In some situations, it is convenient to restart simulations from a previously stored checkpoint, e.g. when the cluster burned down for no apparent reasons.
In SchNetPack, this can be done by loading the checkpoint file with torch and then passing it to a `Siulator` using the `restart_simulation` function (here we use the same instance of simulator as before for convenience, in a real setup a new one would be initialized).
```
checkpoint = torch.load(chk_file)
md_simulator.restart_simulation(checkpoint)
```
This restores the full state, including the system state, simulation steps and states of the thermostats.
Sometimes, it can be sufficient to only restore the system state (positions and momenta), for example when starting production level simulations after equilibration.
This is achieved by calling `load_system_state` on the loaded checkpoint:
```
md_simulator.load_system_state(checkpoint)
```
## Ring polymer molecular dynamics with SchNetPack
Above, we have computed a vibrational spectrum of ethanol based on a classical MD simulation using the Velocity Verlet integrator.
Unfortunately, this approach completely neglects nuclear quantum effects, such as zero-point energies, etc.
One way to recover some of these effects is to use so-called ring polymer molecular dynamics (RPMD).
In RPMD, multiple replicas of a system are connected via harmonic springs and propagated simultaneously.
This can be thought of a discretization of the path integral formulation of quantum mechanics.
The fully quantum solution is then recovered in the limit of an infinite number of replicas, also called beads.
RPMD simulations can easily be carried out using the SchNetPack MD package.
Due to need to perform a large number of computations, RPMD profits greatly from the use of machine learning potentials.
Moreover, the presence of multiple replicas lends it self to an efficient parallelization of GPUs, which is one reason for the special structure of the system tensors used in SchNetPack-MD.
Here, we will repeat the above simulation for ethanol using RPMD instead of a classical simulation.
The main differences in the setup are:
- the system needs to be initialized with multiple repkicas
- a ring polymer integrator needs to be used
- special thermostats are required if a canonical ensemble should be simulated
The `System` can be set up in exactly the same manner as before, only the number of replicas is now set to be greater than one.
For demonstration purposes we use `n_replicas=4`, in general larger numbers are recommened.
```
# Number of beads in RPMD simulation
n_replicas = 4
# Set up the system, load structures, initialize
rpmd_system = System(n_replicas, device=md_device)
rpmd_system.load_molecules_from_xyz(molecule_path)
# Initialize momenta
rpmd_initializer = MaxwellBoltzmannInit(
system_temperature,
remove_translation=True,
remove_rotation=True)
rpmd_initializer.initialize_system(rpmd_system)
```
Next, we need to change the integrator to the `RingPolymer` integrator.
For RPMD, we need to use a smaller time step, in order to keep the integration numerically stable.
In addition, one needs to specify a temperature for the ring polymer, which modulates how strongly the different beads couple.
Typically, we use the same temperature as for the thermostat.
```
# Use the RPMD integrator
from schnetpack.md.integrators import RingPolymer
# Here, a smaller timestep is required for numerical stability
rpmd_time_step = 0.2 # fs
# Initialize the integrator, RPMD also requires a polymer temperature which determines the coupling of beads.
# Here, we set it to the system temperature
rpmd_integrator = RingPolymer(
n_replicas,
rpmd_time_step,
system_temperature,
device=md_device
)
```
Next, we have to change our thermostat to one suitable for RPMD simulations.
Here, we will use the local PILE thermostat, which can be thought of as a RPMD equivalent of the classical Langevin thermostat used above.
In general, SchNetPack comes with a wide variety of theromstats for classical and ring polymer simulations (see the [thermostats module](../modules/md.rst#module-schnetpack.md.simulation_hooks.thermostats)).
For the environment temperature and time constant, the same values as above are used.
```
# Initialize the thermostat
pile = thermostats.PILELocalThermostat(bath_temperature, time_constant)
```
The hooks are generated in exactly the same way as before.
```
# Logging
rpmd_log_file = os.path.join(md_workdir, 'rpmd_simulation.hdf5')
rpmd_data_streams = [
logging_hooks.MoleculeStream(),
logging_hooks.PropertyStream(),
]
rpmd_file_logger = logging_hooks.FileLogger(
rpmd_log_file,
buffer_size,
data_streams=rpmd_data_streams
)
# Checkpoints
rpmd_chk_file = os.path.join(md_workdir, 'rpmd_simulation.chk')
rpmd_checkpoint = logging_hooks.Checkpoint(rpmd_chk_file, every_n_steps=100)
# Assemble the hooks:
rpmd_hooks = [
pile,
rpmd_file_logger,
rpmd_checkpoint
]
```
And so is the simulator:
```
# Assemble the simulator
rpmd_simulator = Simulator(rpmd_system, rpmd_integrator, md_calculator, simulator_hooks=rpmd_hooks)
```
Bow we can carry out the simulations.
Since our time step is shorter, we will run for longer in order to cover the same time scale as the classical simulation (runs approximately 13 minutes on a notebook GPU):
```
n_steps = 50000
rpmd_simulator.simulate(n_steps)
```
Loading of the data with the `HDF5Loader` works exactly the same as before.
When loading properties for RPMD datafiles, the `HDF5Loader` default of using centroid properties (meaning an average over all beads) becomes active.
This is usually what one wants to analyze.
If a specific replica should be used, it can be specified via `replica_idx` in the `get_property` function.
Here, we immediatly skip the a part of the trajectory to only load the equilibrated system.
```
rpmd_data = HDF5Loader(rpmd_log_file, skip_initial=0)
plot_temperature(rpmd_data)
```
Finally, we can compute the power spectrum and compare it to its classical counterpart:
```
# Intialize the spectrum
rpmd_spectrum = PowerSpectrum(rpmd_data, resolution=8192)
# Compute the spectrum for the first molecule (default)
rpmd_spectrum.compute_spectrum(molecule_idx=0)
# Get frequencies and intensities
rpmd_freqencies, rpmd_intensities = rpmd_spectrum.get_spectrum()
# Plot the spectrum
plt.figure(figsize=(8,4))
plt.plot(freqencies, intensities, label='MD')
plt.plot(rpmd_freqencies, rpmd_intensities, label='RPMD')
plt.xlim(0,4000)
plt.ylim(0,100)
plt.ylabel('I [a.u.]')
plt.xlabel('$\omega$ [cm$^{-1}$]')
plt.legend()
plt.show()
```
One problem of purely classical simulations can be observed in the high frequency regions of the MD spectrum.
Peaks are shifted towards higher wave numbers compared to the expected [experimental values](https://chem.libretexts.org/Ancillary_Materials/Reference/Reference_Tables/Spectroscopic_Parameters/Infrared_Spectroscopy_Absorption_Table),
e.g. 3100 cm<sup>-1</sup> vs. 2900 cm<sup>-1</sup> for the CH vibrations.
The inclusion of effects like zero point energy shifts these bands towards lower frequencies, leading to an improved agreement with experiment.
## Quick setup with input files
Although encoding simulations in the way above can be useful when testing and developing new approaches, it has limited use for routine simulations.
Because of this, the MD package also provides the script `spk_md.py`, which can be used to run a simulation according to the instructions given in an input file.
The input file uses yaml format and is structured in a similar way as the main modules introduced above.
In the following, we will construct an input file for repeating the classical MD simulation above.
### Input file format
The first few lines of the input files contain general instructions (e.g. device, random seed, simulation directory):
```yaml
device: cuda
simulation_dir: mdtut
seed: 662524648
overwrite: false
```
If no seed is specified, a new one is generated. The `overwrite` flag specifies, whether an old simulation should be overwritten.
Instructions for the calculator are specified in the `calculator` block:
```yaml
calculator:
type: schnet
model_file: PATH/TO/MODEL
required_properties:
- energy
- forces
force_handle: forces
position_conversion: Angstrom
force_conversion: kcal/mol/Angstrom
```
Here, the type of calculator is specified and path to the model has to be set accordingly (`model_file`).
Conversion units are given in the same manner as above.
The system is specified in the `system` block of the input file, which controls the number of replicas, the file the structure is loaded from, as well as initialization routines:
```yaml
system:
molecule_file: PATH/TO/STRUCTURE.xyz
n_replicas: 1
initializer:
type: maxwell-boltzmann
temperature: 300
remove_translation: true
remove_rotation: true
```
The control of the dynamics simulation itsself is handled in the `dynamics` section:
```yaml
dynamics:
n_steps: 20000
integrator:
type: verlet
time_step: 0.50
thermostat:
type: langevin
temperature: 300
time_constant: 100
```
Thermostats are invoked via `thermostat`, where the type is specified via a string.
The additional arguments vary according to the thermostat used.
The integrator is given over the `integrator` subblock. Units are the same as used in the examples above.
Finally, logging to a file is handled via `logging`:
```yaml
file_logger:
buffer_size: 100
streams:
- molecules
- properties
write_checkpoints: 100
```
The logging file name is set to `simulation_dir/simulation.hdf5` by default, while the checkpoint file uses `simulation_dir/simulation.chk`.
Data streams for the file logger are passes via a list, where `molecules` corresponds to the `MoleculeStream` and `properties` to the `PropertiesStream`.
### Performing a Simulation
Once an input file has been generated, it can be called via
```
schnetpack_molecular_dynamics.py input_file
```
SchNetPack also stores a config file in yaml format into the `simulation_dir` containing the detailed settings of the MD.
### Available Options
There will be a more general documentation on the options available for the input file in the future.
For now, the most important base settings can be found in the `schnetpack.md.parsers.md_options` module, which containes the initializers for the input blocks.
These initializers are:
- `ThermostatInit` for thermostats
- `IntegratorInit` for the integrators
- `InitialConditionsInit` for controlling initial conditions
- `CalculatorInit` for setting up the calculator
All of these classes have the utility function `print_options`.
When this function is called, it prints as short summary on the available options and which inputs (and input types) are required as a minimum.
In case of the thermostats, the following is obtained:
```
from schnetpack.md.parsers.md_options import *
ThermostatInit.print_options()
```
In addition, every block in the input file can also be passed additional input options of the basic modules.
An example is the calculator block:
```yaml
calculator:
type: schnet
model_file: PATH/TO/MODEL
required_properties:
- energy
- forces
force_handle: forces
position_conversion: Angstrom
force_conversion: kcal/mol/Angstrom
```
Compared to the required input options for the SchNet calculator (`model`, `required_properties` and `force_handle`)
```
CalculatorInit.print_options()
```
it is also possible to pass additional keyword arguments (here `position_conversion` and `force_conversion`).
These correspond directly to the input parameters of the target class (see [calculators](../modules/md.rst#module-schnetpack.md.calculators)).
### Example input files
In the following, we provide two example input files for the classical MD and RPMD simulations performed above.
Only the paths (`simulation_dir`, `model_file` and `molecule_file`) and the `device` need to be adjusted accordingly.
#### Classical MD
```yaml
device: cpu
simulation_dir: mdtut_md
overwrite: false
calculator:
type: schnet
model_file: PATH/TO/MODEL
required_properties:
- energy
- forces
force_handle: forces
position_conversion: Angstrom
force_conversion: kcal/mol/Angstrom
system:
molecule_file: PATH/TO/STRUCTURE.xyz
n_replicas: 1
initializer:
type: maxwell-boltzmann
temperature: 300
remove_translation: true
remove_rotation: true
dynamics:
n_steps: 20000
integrator:
type: verlet
time_step: 0.50
thermostat:
type: langevin
temperature: 300
time_constant: 100
logging:
file_logger:
buffer_size: 100
streams:
- molecules
- properties
write_checkpoints: 100
```
#### RPMD
```yaml
device: cpu
simulation_dir: mdtut_rpmd
overwrite: false
calculator:
type: schnet
model_file: PATH/TO/MODEL
required_properties:
- energy
- forces
force_handle: forces
position_conversion: Angstrom
force_conversion: kcal/mol/Angstrom
system:
molecule_file: PATH/TO/STRUCTURE.xyz
n_replicas: 4
initializer:
type: maxwell-boltzmann
temperature: 300
remove_translation: true
remove_rotation: true
dynamics:
n_steps: 50000
integrator:
type: ring_polymer
time_step: 0.20
temperature: 300
thermostat:
type: pile-l
temperature: 300
time_constant: 100
logging:
file_logger:
buffer_size: 100
streams:
- molecules
- properties
write_checkpoints: 100
```
## Summary
In this tutorial, we have given a basic introduction to the structure and functionality of the MD package in SchNetPack.
After setting up a standard MD simulation, we have explored how to use simulation hooks to control simulations in a modular way.
We have shown how to extract data from the HDF5 files generated during MD and how available functions can be used to compute dynamic quanitities, such as power spectra.
This was followed by a short example of using more advanced simulation techniques in the form of RPMD.
Finally, a short introduction to the `spk_md.py` and its input file structure was given.
Future tutorials will cover the use of advanged sampling techniques (e.g. metadynamics) and how to write custom calculators and hooks for performing your own simulations.
| github_jupyter |
# **Noise2Void (2D)**
---
<font size = 4> Noise2Void is a deep-learning method that can be used to denoise many types of images, including microscopy images and which was originally published by [Krull *et al.* on arXiv](https://arxiv.org/abs/1811.10980). It allows denoising of image data in a self-supervised manner, therefore high-quality, low noise equivalent images are not necessary to train this network. This is performed by "masking" a random subset of pixels in the noisy image and training the network to predict the values in these pixels. The resulting output is a denoised version of the image. Noise2Void is based on the popular U-Net network architecture, adapted from [CARE](https://www.nature.com/articles/s41592-018-0216-7).
<font size = 4> **This particular notebook enables self-supervised denoised of 2D dataset. If you are interested in 3D dataset, you should use the Noise2Void 3D notebook instead.**
---
<font size = 4>*Disclaimer*:
<font size = 4>This notebook is part of the Zero-Cost Deep-Learning to Enhance Microscopy project (https://github.com/HenriquesLab/DeepLearning_Collab/wiki). Jointly developed by the Jacquemet (link to https://cellmig.org/) and Henriques (https://henriqueslab.github.io/) laboratories.
<font size = 4>This notebook is largely based on the following paper:
<font size = 4>**Noise2Void - Learning Denoising from Single Noisy Images**
from Krull *et al.* published on arXiv in 2018 (https://arxiv.org/abs/1811.10980)
<font size = 4>And source code found in: https://github.com/juglab/n2v
<font size = 4>**Please also cite this original paper when using or developing this notebook.**
# **How to use this notebook?**
---
<font size = 4>Video describing how to use our notebooks are available on youtube:
- [**Video 1**](https://www.youtube.com/watch?v=GzD2gamVNHI&feature=youtu.be): Full run through of the workflow to obtain the notebooks and the provided test datasets as well as a common use of the notebook
- [**Video 2**](https://www.youtube.com/watch?v=PUuQfP5SsqM&feature=youtu.be): Detailed description of the different sections of the notebook
---
###**Structure of a notebook**
<font size = 4>The notebook contains two types of cell:
<font size = 4>**Text cells** provide information and can be modified by douple-clicking the cell. You are currently reading the text cell. You can create a new text by clicking `+ Text`.
<font size = 4>**Code cells** contain code and the code can be modfied by selecting the cell. To execute the cell, move your cursor on the `[ ]`-mark on the left side of the cell (play button appears). Click to execute the cell. After execution is done the animation of play button stops. You can create a new coding cell by clicking `+ Code`.
---
###**Table of contents, Code snippets** and **Files**
<font size = 4>On the top left side of the notebook you find three tabs which contain from top to bottom:
<font size = 4>*Table of contents* = contains structure of the notebook. Click the content to move quickly between sections.
<font size = 4>*Code snippets* = contain examples how to code certain tasks. You can ignore this when using this notebook.
<font size = 4>*Files* = contain all available files. After mounting your google drive (see section 1.) you will find your files and folders here.
<font size = 4>**Remember that all uploaded files are purged after changing the runtime.** All files saved in Google Drive will remain. You do not need to use the Mount Drive-button; your Google Drive is connected in section 1.2.
<font size = 4>**Note:** The "sample data" in "Files" contains default files. Do not upload anything in here!
---
###**Making changes to the notebook**
<font size = 4>**You can make a copy** of the notebook and save it to your Google Drive. To do this click file -> save a copy in drive.
<font size = 4>To **edit a cell**, double click on the text. This will show you either the source code (in code cells) or the source text (in text cells).
You can use the `#`-mark in code cells to comment out parts of the code. This allows you to keep the original code piece in the cell as a comment.
# **0. Before getting started**
---
<font size = 4>Before you run the notebook, please ensure that you are logged into your Google account and have the training and/or data to process in your Google Drive.
<font size = 4>For Noise2Void to train, it only requires a single noisy image but multiple images can be used. Information on how to generate a training dataset is available in our Wiki page: https://github.com/HenriquesLab/ZeroCostDL4Mic/wiki
<font size = 4>Please note that you currently can **only use .tif files!**
<font size = 4>**We strongly recommend that you generate high signal to noise ration version of your noisy images (Quality control dataset). These images can be used to assess the quality of your trained model**. The quality control assessment can be done directly in this notebook.
<font size = 4> You can also provide a folder that contains the data that you wish to analyse with the trained network once all training has been performed.
<font size = 4>Here is a common data structure that can work:
* Data
- **Training dataset**
- **Quality control dataset** (Optional but recomended)
- Low SNR images
- img_1.tif, img_2.tif
- High SNR images
- img_1.tif, img_2.tif
- **Data to be predicted**
- Results
<font size = 4>The **Results** folder will contain the processed images, trained model and network parameters as csv file. Your original images remain unmodified.
---
<font size = 4>**Important note**
<font size = 4>- If you wish to **train a network from scratch** using your own dataset (and we encourage everyone to do that), you will need to run **sections 1 - 4**, then use **section 5** to assess the quality of your model and **section 6** to run predictions using the model that you trained.
<font size = 4>- If you wish to **evaluate your model** using a model previously generated and saved on your Google Drive, you will only need to run **sections 1 and 2** to set up the notebook, then use **section 5** to assess the quality of your model.
<font size = 4>- If you only wish to **run predictions** using a model previously generated and saved on your Google Drive, you will only need to run **sections 1 and 2** to set up the notebook, then use **section 6** to run the predictions on the desired model.
---
# **1. Initialise the Colab session**
---
## **1.1. Check for GPU access**
---
By default, the session should be using Python 3 and GPU acceleration, but it is possible to ensure that these are set properly by doing the following:
<font size = 4>Go to **Runtime -> Change the Runtime type**
<font size = 4>**Runtime type: Python 3** *(Python 3 is programming language in which this program is written)*
<font size = 4>**Accelator: GPU** *(Graphics processing unit)*
```
#@markdown ##Run this cell to check if you have GPU access
# %tensorflow_version 1.x
import tensorflow as tf
if tf.test.gpu_device_name()=='':
print('You do not have GPU access.')
print('Did you change your runtime ?')
print('If the runtime setting is correct then Google did not allocate a GPU for your session')
print('Expect slow performance. To access GPU try reconnecting later')
else:
print('You have GPU access')
!nvidia-smi
```
## **1.2. Mount your Google Drive**
---
<font size = 4> To use this notebook on the data present in your Google Drive, you need to mount your Google Drive to this notebook.
<font size = 4> Play the cell below to mount your Google Drive and follow the link. In the new browser window, select your drive and select 'Allow', copy the code, paste into the cell and press enter. This will give Colab access to the data on the drive.
<font size = 4> Once this is done, your data are available in the **Files** tab on the top left of notebook.
```
#@markdown ##Play the cell to connect your Google Drive to Colab
#@markdown * Click on the URL.
#@markdown * Sign in your Google Account.
#@markdown * Copy the authorization code.
#@markdown * Enter the authorization code.
#@markdown * Click on "Files" site on the right. Refresh the site. Your Google Drive folder should now be available here as "drive".
# mount user's Google Drive to Google Colab.
from google.colab import drive
drive.mount('/content/gdrive')
```
# **2. Install Noise2Void and dependencies**
---
```
Notebook_version = ['1.12']
#@markdown ##Install Noise2Void and dependencies
# Here we enable Tensorflow 1.
!pip install q keras==2.2.5
%tensorflow_version 1.x
import tensorflow
print(tensorflow.__version__)
print("Tensorflow enabled.")
# Here we install Noise2Void and other required packages
!pip install n2v
!pip install wget
!pip install fpdf
!pip install memory_profiler
%load_ext memory_profiler
print("Noise2Void installed.")
# Here we install all libraries and other depencies to run the notebook.
# ------- Variable specific to N2V -------
from n2v.models import N2VConfig, N2V
from csbdeep.utils import plot_history
from n2v.utils.n2v_utils import manipulate_val_data
from n2v.internals.N2V_DataGenerator import N2V_DataGenerator
from csbdeep.io import save_tiff_imagej_compatible
# ------- Common variable to all ZeroCostDL4Mic notebooks -------
import numpy as np
from matplotlib import pyplot as plt
import urllib
import os, random
import shutil
import zipfile
from tifffile import imread, imsave
import time
import sys
import wget
from pathlib import Path
import pandas as pd
import csv
from glob import glob
from scipy import signal
from scipy import ndimage
from skimage import io
from sklearn.linear_model import LinearRegression
from skimage.util import img_as_uint
import matplotlib as mpl
from skimage.metrics import structural_similarity
from skimage.metrics import peak_signal_noise_ratio as psnr
from astropy.visualization import simple_norm
from skimage import img_as_float32
from fpdf import FPDF, HTMLMixin
from datetime import datetime
from pip._internal.operations.freeze import freeze
import subprocess
from datetime import datetime
# Colors for the warning messages
class bcolors:
WARNING = '\033[31m'
W = '\033[0m' # white (normal)
R = '\033[31m' # red
#Disable some of the tensorflow warnings
import warnings
warnings.filterwarnings("ignore")
print("Libraries installed")
# Check if this is the latest version of the notebook
Latest_notebook_version = pd.read_csv("https://raw.githubusercontent.com/HenriquesLab/ZeroCostDL4Mic/master/Colab_notebooks/Latest_ZeroCostDL4Mic_Release.csv")
print('Notebook version: '+Notebook_version[0])
strlist = Notebook_version[0].split('.')
Notebook_version_main = strlist[0]+'.'+strlist[1]
if Notebook_version_main == Latest_notebook_version.columns:
print("This notebook is up-to-date.")
else:
print(bcolors.WARNING +"A new version of this notebook has been released. We recommend that you download it at https://github.com/HenriquesLab/ZeroCostDL4Mic/wiki")
def pdf_export(trained = False, augmentation = False, pretrained_model = False):
class MyFPDF(FPDF, HTMLMixin):
pass
pdf = MyFPDF()
pdf.add_page()
pdf.set_right_margin(-1)
pdf.set_font("Arial", size = 11, style='B')
Network = 'Noise2Void 2D'
day = datetime.now()
datetime_str = str(day)[0:10]
Header = 'Training report for '+Network+' model ('+model_name+')\nDate: '+datetime_str
pdf.multi_cell(180, 5, txt = Header, align = 'L')
# add another cell
if trained:
training_time = "Training time: "+str(hour)+ "hour(s) "+str(mins)+"min(s) "+str(round(sec))+"sec(s)"
pdf.cell(190, 5, txt = training_time, ln = 1, align='L')
pdf.ln(1)
Header_2 = 'Information for your materials and method:'
pdf.cell(190, 5, txt=Header_2, ln=1, align='L')
all_packages = ''
for requirement in freeze(local_only=True):
all_packages = all_packages+requirement+', '
#print(all_packages)
#Main Packages
main_packages = ''
version_numbers = []
for name in ['tensorflow','numpy','Keras','csbdeep']:
find_name=all_packages.find(name)
main_packages = main_packages+all_packages[find_name:all_packages.find(',',find_name)]+', '
#Version numbers only here:
version_numbers.append(all_packages[find_name+len(name)+2:all_packages.find(',',find_name)])
cuda_version = subprocess.run('nvcc --version',stdout=subprocess.PIPE, shell=True)
cuda_version = cuda_version.stdout.decode('utf-8')
cuda_version = cuda_version[cuda_version.find(', V')+3:-1]
gpu_name = subprocess.run('nvidia-smi',stdout=subprocess.PIPE, shell=True)
gpu_name = gpu_name.stdout.decode('utf-8')
gpu_name = gpu_name[gpu_name.find('Tesla'):gpu_name.find('Tesla')+10]
#print(cuda_version[cuda_version.find(', V')+3:-1])
#print(gpu_name)
shape = io.imread(Training_source+'/'+os.listdir(Training_source)[0]).shape
dataset_size = len(os.listdir(Training_source))
text = 'The '+Network+' model was trained from scratch for '+str(number_of_epochs)+' epochs on '+str(Xdata.shape[0])+' image patches (image dimensions: '+str(shape)+', patch size: ('+str(patch_size)+','+str(patch_size)+')) with a batch size of '+str(batch_size)+' and a '+config.train_loss+' loss function, using the '+Network+' ZeroCostDL4Mic notebook (v '+Notebook_version[0]+') (von Chamier & Laine et al., 2020). Key python packages used include tensorflow (v '+version_numbers[0]+'), Keras (v '+version_numbers[2]+'), csbdeep (v '+version_numbers[3]+'), numpy (v '+version_numbers[1]+'), cuda (v '+cuda_version+'). The training was accelerated using a '+gpu_name+'GPU.'
if pretrained_model:
text = 'The '+Network+' model was trained for '+str(number_of_epochs)+' epochs on '+str(Xdata.shape[0])+' paired image patches (image dimensions: '+str(shape)+', patch size: ('+str(patch_size)+','+str(patch_size)+')) with a batch size of '+str(batch_size)+' and a '+config.train_loss+' loss function, using the '+Network+' ZeroCostDL4Mic notebook (v '+Notebook_version[0]+') (von Chamier & Laine et al., 2020). The model was re-trained from a pretrained model. Key python packages used include tensorflow (v '+version_numbers[0]+'), Keras (v '+version_numbers[2]+'), csbdeep (v '+version_numbers[3]+'), numpy (v '+version_numbers[1]+'), cuda (v '+cuda_version+'). The training was accelerated using a '+gpu_name+'GPU.'
pdf.set_font('')
pdf.set_font_size(10.)
pdf.multi_cell(190, 5, txt = text, align='L')
pdf.set_font('')
pdf.set_font('Arial', size = 10, style = 'B')
pdf.ln(1)
pdf.cell(26, 5, txt='Augmentation: ', ln=0)
pdf.set_font('')
if augmentation:
aug_text = 'The dataset was augmented by default.'
else:
aug_text = 'No augmentation was used for training.'
pdf.multi_cell(190, 5, txt=aug_text, align='L')
pdf.set_font('Arial', size = 11, style = 'B')
pdf.ln(1)
pdf.cell(180, 5, txt = 'Parameters', align='L', ln=1)
pdf.set_font('')
pdf.set_font_size(10.)
if Use_Default_Advanced_Parameters:
pdf.cell(200, 5, txt='Default Advanced Parameters were enabled')
pdf.cell(200, 5, txt='The following parameters were used for training:')
pdf.ln(1)
html = """
<table width=40% style="margin-left:0px;">
<tr>
<th width = 50% align="left">Parameter</th>
<th width = 50% align="left">Value</th>
</tr>
<tr>
<td width = 50%>number_of_epochs</td>
<td width = 50%>{0}</td>
</tr>
<tr>
<td width = 50%>patch_size</td>
<td width = 50%>{1}</td>
</tr>
<tr>
<td width = 50%>batch_size</td>
<td width = 50%>{2}</td>
</tr>
<tr>
<td width = 50%>number_of_steps</td>
<td width = 50%>{3}</td>
</tr>
<tr>
<td width = 50%>percentage_validation</td>
<td width = 50%>{4}</td>
</tr>
<tr>
<td width = 50%>initial_learning_rate</td>
<td width = 50%>{5}</td>
</tr>
</table>
""".format(number_of_epochs,str(patch_size)+'x'+str(patch_size),batch_size,number_of_steps,percentage_validation,initial_learning_rate)
pdf.write_html(html)
#pdf.multi_cell(190, 5, txt = text_2, align='L')
pdf.set_font("Arial", size = 11, style='B')
pdf.ln(1)
pdf.cell(190, 5, txt = 'Training Dataset', align='L', ln=1)
pdf.set_font('')
pdf.set_font('Arial', size = 10, style = 'B')
pdf.cell(28, 5, txt= 'Training_source:', align = 'L', ln=0)
pdf.set_font('')
pdf.multi_cell(170, 5, txt = Training_source, align = 'L')
# pdf.set_font('')
# pdf.set_font('Arial', size = 10, style = 'B')
# pdf.cell(28, 5, txt= 'Training_target:', align = 'L', ln=0)
# pdf.set_font('')
# pdf.multi_cell(170, 5, txt = Training_target, align = 'L')
#pdf.cell(190, 5, txt=aug_text, align='L', ln=1)
pdf.ln(1)
pdf.set_font('')
pdf.set_font('Arial', size = 10, style = 'B')
pdf.cell(21, 5, txt= 'Model Path:', align = 'L', ln=0)
pdf.set_font('')
pdf.multi_cell(170, 5, txt = model_path+'/'+model_name, align = 'L')
pdf.ln(1)
pdf.cell(60, 5, txt = 'Example Training Image', ln=1)
pdf.ln(1)
exp_size = io.imread('/content/TrainingDataExample_N2V2D.png').shape
pdf.image('/content/TrainingDataExample_N2V2D.png', x = 11, y = None, w = round(exp_size[1]/8), h = round(exp_size[0]/8))
pdf.ln(1)
ref_1 = 'References:\n - ZeroCostDL4Mic: von Chamier, Lucas & Laine, Romain, et al. "ZeroCostDL4Mic: an open platform to simplify access and use of Deep-Learning in Microscopy." BioRxiv (2020).'
pdf.multi_cell(190, 5, txt = ref_1, align='L')
ref_2 = '- Noise2Void: Krull, Alexander, Tim-Oliver Buchholz, and Florian Jug. "Noise2void-learning denoising from single noisy images." Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition. 2019.'
pdf.multi_cell(190, 5, txt = ref_2, align='L')
pdf.ln(3)
reminder = 'Important:\nRemember to perform the quality control step on all newly trained models\nPlease consider depositing your training dataset on Zenodo'
pdf.set_font('Arial', size = 11, style='B')
pdf.multi_cell(190, 5, txt=reminder, align='C')
pdf.output(model_path+'/'+model_name+'/'+model_name+"_training_report.pdf")
#Make a pdf summary of the QC results
def qc_pdf_export():
class MyFPDF(FPDF, HTMLMixin):
pass
pdf = MyFPDF()
pdf.add_page()
pdf.set_right_margin(-1)
pdf.set_font("Arial", size = 11, style='B')
Network = 'Noise2Void 2D'
day = datetime.now()
datetime_str = str(day)[0:10]
Header = 'Quality Control report for '+Network+' model ('+QC_model_name+')\nDate: '+datetime_str
pdf.multi_cell(180, 5, txt = Header, align = 'L')
all_packages = ''
for requirement in freeze(local_only=True):
all_packages = all_packages+requirement+', '
pdf.set_font('')
pdf.set_font('Arial', size = 11, style = 'B')
pdf.ln(2)
pdf.cell(190, 5, txt = 'Development of Training Losses', ln=1, align='L')
pdf.ln(1)
exp_size = io.imread(full_QC_model_path+'/Quality Control/lossCurvePlots.png').shape
if os.path.exists(full_QC_model_path+'/Quality Control/lossCurvePlots.png'):
pdf.image(full_QC_model_path+'/Quality Control/lossCurvePlots.png', x = 11, y = None, w = round(exp_size[1]/8), h = round(exp_size[0]/8))
else:
pdf.set_font('')
pdf.set_font('Arial', size=10)
pdf.cell(190, 5, txt='If you would like to see the evolution of the loss function during training please play the first cell of the QC section in the notebook.')
pdf.ln(2)
pdf.set_font('')
pdf.set_font('Arial', size = 10, style = 'B')
pdf.ln(3)
pdf.cell(80, 5, txt = 'Example Quality Control Visualisation', ln=1)
pdf.ln(1)
exp_size = io.imread(full_QC_model_path+'/Quality Control/QC_example_data.png').shape
pdf.image(full_QC_model_path+'/Quality Control/QC_example_data.png', x = 16, y = None, w = round(exp_size[1]/10), h = round(exp_size[0]/10))
pdf.ln(1)
pdf.set_font('')
pdf.set_font('Arial', size = 11, style = 'B')
pdf.ln(1)
pdf.cell(180, 5, txt = 'Quality Control Metrics', align='L', ln=1)
pdf.set_font('')
pdf.set_font_size(10.)
pdf.ln(1)
html = """
<body>
<font size="7" face="Courier New" >
<table width=94% style="margin-left:0px;">"""
with open(full_QC_model_path+'/Quality Control/QC_metrics_'+QC_model_name+'.csv', 'r') as csvfile:
metrics = csv.reader(csvfile)
header = next(metrics)
image = header[0]
mSSIM_PvsGT = header[1]
mSSIM_SvsGT = header[2]
NRMSE_PvsGT = header[3]
NRMSE_SvsGT = header[4]
PSNR_PvsGT = header[5]
PSNR_SvsGT = header[6]
header = """
<tr>
<th width = 10% align="left">{0}</th>
<th width = 15% align="left">{1}</th>
<th width = 15% align="center">{2}</th>
<th width = 15% align="left">{3}</th>
<th width = 15% align="center">{4}</th>
<th width = 15% align="left">{5}</th>
<th width = 15% align="center">{6}</th>
</tr>""".format(image,mSSIM_PvsGT,mSSIM_SvsGT,NRMSE_PvsGT,NRMSE_SvsGT,PSNR_PvsGT,PSNR_SvsGT)
html = html+header
for row in metrics:
image = row[0]
mSSIM_PvsGT = row[1]
mSSIM_SvsGT = row[2]
NRMSE_PvsGT = row[3]
NRMSE_SvsGT = row[4]
PSNR_PvsGT = row[5]
PSNR_SvsGT = row[6]
cells = """
<tr>
<td width = 10% align="left">{0}</td>
<td width = 15% align="center">{1}</td>
<td width = 15% align="center">{2}</td>
<td width = 15% align="center">{3}</td>
<td width = 15% align="center">{4}</td>
<td width = 15% align="center">{5}</td>
<td width = 15% align="center">{6}</td>
</tr>""".format(image,str(round(float(mSSIM_PvsGT),3)),str(round(float(mSSIM_SvsGT),3)),str(round(float(NRMSE_PvsGT),3)),str(round(float(NRMSE_SvsGT),3)),str(round(float(PSNR_PvsGT),3)),str(round(float(PSNR_SvsGT),3)))
html = html+cells
html = html+"""</body></table>"""
pdf.write_html(html)
pdf.ln(1)
pdf.set_font('')
pdf.set_font_size(10.)
ref_1 = 'References:\n - ZeroCostDL4Mic: von Chamier, Lucas & Laine, Romain, et al. "ZeroCostDL4Mic: an open platform to simplify access and use of Deep-Learning in Microscopy." BioRxiv (2020).'
pdf.multi_cell(190, 5, txt = ref_1, align='L')
ref_2 = '- Noise2Void: Krull, Alexander, Tim-Oliver Buchholz, and Florian Jug. "Noise2void-learning denoising from single noisy images." Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition. 2019.'
pdf.multi_cell(190, 5, txt = ref_2, align='L')
pdf.ln(3)
reminder = 'To find the parameters and other information about how this model was trained, go to the training_report.pdf of this model which should be in the folder of the same name.'
pdf.set_font('Arial', size = 11, style='B')
pdf.multi_cell(190, 5, txt=reminder, align='C')
pdf.output(full_QC_model_path+'/Quality Control/'+QC_model_name+'_QC_report.pdf')
```
# **3. Select your parameters and paths**
---
## **3.1. Setting main training parameters**
---
<font size = 4>
<font size = 5> **Paths for training, predictions and results**
<font size = 4>**`Training_source:`:** These is the path to your folders containing the Training_source (noisy images). To find the path of the folder containing your datasets, go to your Files on the left of the notebook, navigate to the folder containing your files and copy the path by right-clicking on the folder, **Copy path** and pasting it into the right box below.
<font size = 4>**`model_name`:** Use only my_model -style, not my-model (Use "_" not "-"). Do not use spaces in the name. Do not re-use the name of an existing model (saved in the same folder), otherwise it will be overwritten.
<font size = 4>**`model_path`**: Enter the path where your model will be saved once trained (for instance your result folder).
<font size = 5>**Training Parameters**
<font size = 4>**`number_of_epochs`:** Input how many epochs (rounds) the network will be trained. Preliminary results can already be observed after a few (10-30) epochs, but a full training should run for 100-200 epochs. Evaluate the performance after training (see 4.3.). **Default value: 100**
<font size = 4>**`patch_size`:** Noise2Void divides the image into patches for training. Input the size of the patches (length of a side). The value should be between 64 and the dimensions of the image and divisible by 8. **Default value: 64**
<font size = 5>**Advanced Parameters - experienced users only**
<font size =4>**`batch_size:`** This parameter defines the number of patches seen in each training step. Noise2Void requires a large batch size for stable training. Reduce this parameter if your GPU runs out of memory. **Default value: 128**
<font size = 4>**`number_of_steps`:** Define the number of training steps by epoch. By default this parameter is calculated so that each image / patch is seen at least once per epoch. **Default value: Number of patch / batch_size**
<font size = 4>**`percentage_validation`:** Input the percentage of your training dataset you want to use to validate the network during the training. **Default value: 10**
<font size = 4>**`initial_learning_rate`:** Input the initial value to be used as learning rate. **Default value: 0.0004**
```
# create DataGenerator-object.
datagen = N2V_DataGenerator()
#@markdown ###Path to training image(s):
Training_source = "" #@param {type:"string"}
#compatibility to easily change the name of the parameters
training_images = Training_source
imgs = datagen.load_imgs_from_directory(directory = Training_source)
#@markdown ### Model name and path:
model_name = "" #@param {type:"string"}
model_path = "" #@param {type:"string"}
full_model_path = model_path+'/'+model_name+'/'
#@markdown ###Training Parameters
#@markdown Number of epochs:
number_of_epochs = 100#@param {type:"number"}
#@markdown Patch size (pixels)
patch_size = 64#@param {type:"number"}
#@markdown ###Advanced Parameters
Use_Default_Advanced_Parameters = True#@param {type:"boolean"}
#@markdown ###If not, please input:
batch_size = 128#@param {type:"number"}
number_of_steps = 100#@param {type:"number"}
percentage_validation = 10#@param {type:"number"}
initial_learning_rate = 0.0004 #@param {type:"number"}
if (Use_Default_Advanced_Parameters):
print("Default advanced parameters enabled")
# number_of_steps is defined in the following cell in this case
batch_size = 128
percentage_validation = 10
initial_learning_rate = 0.0004
#here we check that no model with the same name already exist, if so print a warning
if os.path.exists(model_path+'/'+model_name):
print(bcolors.WARNING +"!! WARNING: "+model_name+" already exists and will be deleted in the following cell !!")
print(bcolors.WARNING +"To continue training "+model_name+", choose a new model_name here, and load "+model_name+" in section 3.3"+W)
# This will open a randomly chosen dataset input image
random_choice = random.choice(os.listdir(Training_source))
x = imread(Training_source+"/"+random_choice)
# Here we check that the input images contains the expected dimensions
if len(x.shape) == 2:
print("Image dimensions (y,x)",x.shape)
if not len(x.shape) == 2:
print(bcolors.WARNING +"Your images appear to have the wrong dimensions. Image dimension",x.shape)
#Find image XY dimension
Image_Y = x.shape[0]
Image_X = x.shape[1]
#Hyperparameters failsafes
# Here we check that patch_size is smaller than the smallest xy dimension of the image
if patch_size > min(Image_Y, Image_X):
patch_size = min(Image_Y, Image_X)
print (bcolors.WARNING + " Your chosen patch_size is bigger than the xy dimension of your image; therefore the patch_size chosen is now:",patch_size)
# Here we check that patch_size is divisible by 8
if not patch_size % 8 == 0:
patch_size = ((int(patch_size / 8)-1) * 8)
print (bcolors.WARNING + " Your chosen patch_size is not divisible by 8; therefore the patch_size chosen is now:",patch_size)
# Here we disable pre-trained model by default (in case the next cell is not run)
Use_pretrained_model = False
# Here we enable data augmentation by default (in case the cell is not ran)
Use_Data_augmentation = True
print("Parameters initiated.")
#Here we display one image
norm = simple_norm(x, percent = 99)
f=plt.figure(figsize=(16,8))
plt.subplot(1,2,1)
plt.imshow(x, interpolation='nearest', norm=norm, cmap='magma')
plt.title('Training source')
plt.axis('off');
plt.savefig('/content/TrainingDataExample_N2V2D.png',bbox_inches='tight',pad_inches=0)
```
## **3.2. Data augmentation**
---
<font size = 4>Data augmentation can improve training progress by amplifying differences in the dataset. This can be useful if the available dataset is small since, in this case, it is possible that a network could quickly learn every example in the dataset (overfitting), without augmentation. Augmentation is not necessary for training and if your training dataset is large you should disable it.
<font size = 4>Data augmentation is performed here by rotating the patches in XY-Plane and flip them along X-Axis. This only works if the patches are square in XY.
<font size = 4> **By default data augmentation is enabled. Disable this option is you run out of RAM during the training**.
```
#Data augmentation
#@markdown ##Play this cell to enable or disable data augmentation:
Use_Data_augmentation = True #@param {type:"boolean"}
if Use_Data_augmentation:
print("Data augmentation enabled")
if not Use_Data_augmentation:
print("Data augmentation disabled")
```
## **3.3. Using weights from a pre-trained model as initial weights**
---
<font size = 4> Here, you can set the the path to a pre-trained model from which the weights can be extracted and used as a starting point for this training session. **This pre-trained model needs to be a N2V 2D model**.
<font size = 4> This option allows you to perform training over multiple Colab runtimes or to do transfer learning using models trained outside of ZeroCostDL4Mic. **You do not need to run this section if you want to train a network from scratch**.
<font size = 4> In order to continue training from the point where the pre-trained model left off, it is adviseable to also **load the learning rate** that was used when the training ended. This is automatically saved for models trained with ZeroCostDL4Mic and will be loaded here. If no learning rate can be found in the model folder provided, the default learning rate will be used.
```
# @markdown ##Loading weights from a pre-trained network
Use_pretrained_model = False #@param {type:"boolean"}
pretrained_model_choice = "Model_from_file" #@param ["Model_from_file"]
Weights_choice = "best" #@param ["last", "best"]
#@markdown ###If you chose "Model_from_file", please provide the path to the model folder:
pretrained_model_path = "" #@param {type:"string"}
# --------------------- Check if we load a previously trained model ------------------------
if Use_pretrained_model:
# --------------------- Load the model from the choosen path ------------------------
if pretrained_model_choice == "Model_from_file":
h5_file_path = os.path.join(pretrained_model_path, "weights_"+Weights_choice+".h5")
# --------------------- Download the a model provided in the XXX ------------------------
if pretrained_model_choice == "Model_name":
pretrained_model_name = "Model_name"
pretrained_model_path = "/content/"+pretrained_model_name
print("Downloading the 2D_Demo_Model_from_Stardist_2D_paper")
if os.path.exists(pretrained_model_path):
shutil.rmtree(pretrained_model_path)
os.makedirs(pretrained_model_path)
wget.download("", pretrained_model_path)
wget.download("", pretrained_model_path)
wget.download("", pretrained_model_path)
wget.download("", pretrained_model_path)
h5_file_path = os.path.join(pretrained_model_path, "weights_"+Weights_choice+".h5")
# --------------------- Add additional pre-trained models here ------------------------
# --------------------- Check the model exist ------------------------
# If the model path chosen does not contain a pretrain model then use_pretrained_model is disabled,
if not os.path.exists(h5_file_path):
print(bcolors.WARNING+'WARNING: weights_last.h5 pretrained model does not exist')
Use_pretrained_model = False
# If the model path contains a pretrain model, we load the training rate,
if os.path.exists(h5_file_path):
#Here we check if the learning rate can be loaded from the quality control folder
if os.path.exists(os.path.join(pretrained_model_path, 'Quality Control', 'training_evaluation.csv')):
with open(os.path.join(pretrained_model_path, 'Quality Control', 'training_evaluation.csv'),'r') as csvfile:
csvRead = pd.read_csv(csvfile, sep=',')
#print(csvRead)
if "learning rate" in csvRead.columns: #Here we check that the learning rate column exist (compatibility with model trained un ZeroCostDL4Mic bellow 1.4)
print("pretrained network learning rate found")
#find the last learning rate
lastLearningRate = csvRead["learning rate"].iloc[-1]
#Find the learning rate corresponding to the lowest validation loss
min_val_loss = csvRead[csvRead['val_loss'] == min(csvRead['val_loss'])]
#print(min_val_loss)
bestLearningRate = min_val_loss['learning rate'].iloc[-1]
if Weights_choice == "last":
print('Last learning rate: '+str(lastLearningRate))
if Weights_choice == "best":
print('Learning rate of best validation loss: '+str(bestLearningRate))
if not "learning rate" in csvRead.columns: #if the column does not exist, then initial learning rate is used instead
bestLearningRate = initial_learning_rate
lastLearningRate = initial_learning_rate
print(bcolors.WARNING+'WARNING: The learning rate cannot be identified from the pretrained network. Default learning rate of '+str(bestLearningRate)+' will be used instead' + W)
#Compatibility with models trained outside ZeroCostDL4Mic but default learning rate will be used
if not os.path.exists(os.path.join(pretrained_model_path, 'Quality Control', 'training_evaluation.csv')):
print(bcolors.WARNING+'WARNING: The learning rate cannot be identified from the pretrained network. Default learning rate of '+str(initial_learning_rate)+' will be used instead'+ W)
bestLearningRate = initial_learning_rate
lastLearningRate = initial_learning_rate
# Display info about the pretrained model to be loaded (or not)
if Use_pretrained_model:
print('Weights found in:')
print(h5_file_path)
print('will be loaded prior to training.')
else:
print(bcolors.WARNING+'No pretrained nerwork will be used.')
```
# **4. Train the network**
---
## **4.1. Prepare the training data and model for training**
---
<font size = 4>Here, we use the information from 3. to build the model and convert the training data into a suitable format for training.
```
#@markdown ##Create the model and dataset objects
# --------------------- Here we delete the model folder if it already exist ------------------------
if os.path.exists(model_path+'/'+model_name):
print(bcolors.WARNING +"!! WARNING: Model folder already exists and has been removed !!" + W)
shutil.rmtree(model_path+'/'+model_name)
# split patches from the training images
Xdata = datagen.generate_patches_from_list(imgs, shape=(patch_size,patch_size), augment=Use_Data_augmentation)
shape_of_Xdata = Xdata.shape
# create a threshold (10 % patches for the validation)
threshold = int(shape_of_Xdata[0]*(percentage_validation/100))
# split the patches into training patches and validation patches
X = Xdata[threshold:]
X_val = Xdata[:threshold]
print(Xdata.shape[0],"patches created.")
print(threshold,"patch images for validation (",percentage_validation,"%).")
print(Xdata.shape[0]-threshold,"patch images for training.")
%memit
#Here we automatically define number_of_step in function of training data and batch size
if (Use_Default_Advanced_Parameters):
number_of_steps= int(X.shape[0]/batch_size)+1
# --------------------- Using pretrained model ------------------------
#Here we ensure that the learning rate set correctly when using pre-trained models
if Use_pretrained_model:
if Weights_choice == "last":
initial_learning_rate = lastLearningRate
if Weights_choice == "best":
initial_learning_rate = bestLearningRate
# --------------------- ---------------------- ------------------------
# create a Config object
config = N2VConfig(X, unet_kern_size=3,
train_steps_per_epoch=number_of_steps, train_epochs=number_of_epochs,
train_loss='mse', batch_norm=True, train_batch_size=batch_size, n2v_perc_pix=0.198,
n2v_manipulator='uniform_withCP', n2v_neighborhood_radius=5, train_learning_rate = initial_learning_rate)
# Let's look at the parameters stored in the config-object.
vars(config)
# create network model.
model = N2V(config=config, name=model_name, basedir=model_path)
# --------------------- Using pretrained model ------------------------
# Load the pretrained weights
if Use_pretrained_model:
model.load_weights(h5_file_path)
# --------------------- ---------------------- ------------------------
print("Setup done.")
print(config)
# creates a plot and shows one training patch and one validation patch.
plt.figure(figsize=(16,87))
plt.subplot(1,2,1)
plt.imshow(X[0,...,0], cmap='magma')
plt.axis('off')
plt.title('Training Patch');
plt.subplot(1,2,2)
plt.imshow(X_val[0,...,0], cmap='magma')
plt.axis('off')
plt.title('Validation Patch');
pdf_export(pretrained_model = Use_pretrained_model)
```
## **4.2. Start Training**
---
<font size = 4>When playing the cell below you should see updates after each epoch (round). Network training can take some time.
<font size = 4>* **CRITICAL NOTE:** Google Colab has a time limit for processing (to prevent using GPU power for datamining). Training time must be less than 12 hours! If training takes longer than 12 hours, please decrease the number of epochs or number of patches. Another way circumvent this is to save the parameters of the model after training and start training again from this
point.
<font size = 4>Once training is complete, the trained model is automatically saved on your Google Drive, in the **model_path** folder that was selected in Section 3. It is however wise to download the folder from Google Drive as all data can be erased at the next training if using the same folder.
<font size = 4>**Of Note:** At the end of the training, your model will be automatically exported so it can be used in the CSB Fiji plugin (Run your Network). You can find it in your model folder (TF_SavedModel.zip). In Fiji, Make sure to choose the right version of tensorflow. You can check at: Edit-- Options-- Tensorflow. Choose the version 1.4 (CPU or GPU depending on your system).
```
start = time.time()
#@markdown ##Start training
%memit
history = model.train(X, X_val)
print("Training done.")
%memit
print("Training, done.")
# convert the history.history dict to a pandas DataFrame:
lossData = pd.DataFrame(history.history)
if os.path.exists(model_path+"/"+model_name+"/Quality Control"):
shutil.rmtree(model_path+"/"+model_name+"/Quality Control")
os.makedirs(model_path+"/"+model_name+"/Quality Control")
# The training evaluation.csv is saved (overwrites the Files if needed).
lossDataCSVpath = model_path+'/'+model_name+'/Quality Control/training_evaluation.csv'
with open(lossDataCSVpath, 'w') as f:
writer = csv.writer(f)
writer.writerow(['loss','val_loss', 'learning rate'])
for i in range(len(history.history['loss'])):
writer.writerow([history.history['loss'][i], history.history['val_loss'][i], history.history['lr'][i]])
# Displaying the time elapsed for training
dt = time.time() - start
mins, sec = divmod(dt, 60)
hour, mins = divmod(mins, 60)
print("Time elapsed:",hour, "hour(s)",mins,"min(s)",round(sec),"sec(s)")
model.export_TF(name='Noise2Void',
description='Noise2Void 2D trained using ZeroCostDL4Mic.',
authors=["You"],
test_img=X_val[0,...,0], axes='YX',
patch_shape=(patch_size, patch_size))
print("Your model has been sucessfully exported and can now also be used in the CSBdeep Fiji plugin")
pdf_export(trained = True, pretrained_model = Use_pretrained_model)
```
# **5. Evaluate your model**
---
<font size = 4>This section allows the user to perform important quality checks on the validity and generalisability of the trained model.
<font size = 4>**We highly recommend to perform quality control on all newly trained models.**
```
# model name and path
#@markdown ###Do you want to assess the model you just trained ?
Use_the_current_trained_model = True #@param {type:"boolean"}
#@markdown ###If not, please provide the path to the model folder:
QC_model_folder = "" #@param {type:"string"}
#Here we define the loaded model name and path
QC_model_name = os.path.basename(QC_model_folder)
QC_model_path = os.path.dirname(QC_model_folder)
if (Use_the_current_trained_model):
QC_model_name = model_name
QC_model_path = model_path
full_QC_model_path = QC_model_path+'/'+QC_model_name+'/'
if os.path.exists(full_QC_model_path):
print("The "+QC_model_name+" network will be evaluated")
else:
print(bcolors.WARNING + '!! WARNING: The chosen model does not exist !!')
print('Please make sure you provide a valid model path and model name before proceeding further.')
```
## **5.1. Inspection of the loss function**
---
<font size = 4>It is good practice to evaluate the training progress by comparing the training loss with the validation loss. The latter is a metric which shows how well the network performs on a subset of unseen data which is set aside from the training dataset. For more information on this, see for example [this review](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6381354/) by Nichols *et al.*
<font size = 4>**Training loss** describes an error value after each epoch for the difference between the model's prediction and its ground-truth target.
<font size = 4>**Validation loss** describes the same error value between the model's prediction on a validation image and compared to it's target.
<font size = 4>During training both values should decrease before reaching a minimal value which does not decrease further even after more training. Comparing the development of the validation loss with the training loss can give insights into the model's performance.
<font size = 4>Decreasing **Training loss** and **Validation loss** indicates that training is still necessary and increasing the `number_of_epochs` is recommended. Note that the curves can look flat towards the right side, just because of the y-axis scaling. The network has reached convergence once the curves flatten out. After this point no further training is required. If the **Validation loss** suddenly increases again an the **Training loss** simultaneously goes towards zero, it means that the network is overfitting to the training data. In other words the network is remembering the exact noise patterns from the training data and no longer generalizes well to unseen data. In this case the training dataset has to be increased.
```
#@markdown ##Play the cell to show a plot of training errors vs. epoch number
lossDataFromCSV = []
vallossDataFromCSV = []
with open(QC_model_path+'/'+QC_model_name+'/Quality Control/training_evaluation.csv','r') as csvfile:
csvRead = csv.reader(csvfile, delimiter=',')
next(csvRead)
for row in csvRead:
lossDataFromCSV.append(float(row[0]))
vallossDataFromCSV.append(float(row[1]))
epochNumber = range(len(lossDataFromCSV))
plt.figure(figsize=(15,10))
plt.subplot(2,1,1)
plt.plot(epochNumber,lossDataFromCSV, label='Training loss')
plt.plot(epochNumber,vallossDataFromCSV, label='Validation loss')
plt.title('Training loss and validation loss vs. epoch number (linear scale)')
plt.ylabel('Loss')
plt.xlabel('Epoch number')
plt.legend()
plt.subplot(2,1,2)
plt.semilogy(epochNumber,lossDataFromCSV, label='Training loss')
plt.semilogy(epochNumber,vallossDataFromCSV, label='Validation loss')
plt.title('Training loss and validation loss vs. epoch number (log scale)')
plt.ylabel('Loss')
plt.xlabel('Epoch number')
plt.legend()
plt.savefig(QC_model_path+'/'+QC_model_name+'/Quality Control/lossCurvePlots.png')
plt.show()
```
## **5.2. Error mapping and quality metrics estimation**
---
<font size = 4>This section will display SSIM maps and RSE maps as well as calculating total SSIM, NRMSE and PSNR metrics for all the images provided in the "Source_QC_folder" and "Target_QC_folder" !
<font size = 4>**1. The SSIM (structural similarity) map**
<font size = 4>The SSIM metric is used to evaluate whether two images contain the same structures. It is a normalized metric and an SSIM of 1 indicates a perfect similarity between two images. Therefore for SSIM, the closer to 1, the better. The SSIM maps are constructed by calculating the SSIM metric in each pixel by considering the surrounding structural similarity in the neighbourhood of that pixel (currently defined as window of 11 pixels and with Gaussian weighting of 1.5 pixel standard deviation, see our Wiki for more info).
<font size=4>**mSSIM** is the SSIM value calculated across the entire window of both images.
<font size=4>**The output below shows the SSIM maps with the mSSIM**
<font size = 4>**2. The RSE (Root Squared Error) map**
<font size = 4>This is a display of the root of the squared difference between the normalized predicted and target or the source and the target. In this case, a smaller RSE is better. A perfect agreement between target and prediction will lead to an RSE map showing zeros everywhere (dark).
<font size =4>**NRMSE (normalised root mean squared error)** gives the average difference between all pixels in the images compared to each other. Good agreement yields low NRMSE scores.
<font size = 4>**PSNR (Peak signal-to-noise ratio)** is a metric that gives the difference between the ground truth and prediction (or source input) in decibels, using the peak pixel values of the prediction and the MSE between the images. The higher the score the better the agreement.
<font size=4>**The output below shows the RSE maps with the NRMSE and PSNR values.**
```
#@markdown ##Choose the folders that contain your Quality Control dataset
Source_QC_folder = "" #@param{type:"string"}
Target_QC_folder = "" #@param{type:"string"}
# Create a quality control/Prediction Folder
if os.path.exists(QC_model_path+"/"+QC_model_name+"/Quality Control/Prediction"):
shutil.rmtree(QC_model_path+"/"+QC_model_name+"/Quality Control/Prediction")
os.makedirs(QC_model_path+"/"+QC_model_name+"/Quality Control/Prediction")
# Activate the pretrained model.
model_training = N2V(config=None, name=QC_model_name, basedir=QC_model_path)
# List Tif images in Source_QC_folder
Source_QC_folder_tif = Source_QC_folder+"/*.tif"
Z = sorted(glob(Source_QC_folder_tif))
Z = list(map(imread,Z))
print('Number of test dataset found in the folder: '+str(len(Z)))
# Perform prediction on all datasets in the Source_QC folder
for filename in os.listdir(Source_QC_folder):
img = imread(os.path.join(Source_QC_folder, filename))
predicted = model_training.predict(img, axes='YX', n_tiles=(2,1))
os.chdir(QC_model_path+"/"+QC_model_name+"/Quality Control/Prediction")
imsave(filename, predicted)
def ssim(img1, img2):
return structural_similarity(img1,img2,data_range=1.,full=True, gaussian_weights=True, use_sample_covariance=False, sigma=1.5)
def normalize(x, pmin=3, pmax=99.8, axis=None, clip=False, eps=1e-20, dtype=np.float32):
"""This function is adapted from Martin Weigert"""
"""Percentile-based image normalization."""
mi = np.percentile(x,pmin,axis=axis,keepdims=True)
ma = np.percentile(x,pmax,axis=axis,keepdims=True)
return normalize_mi_ma(x, mi, ma, clip=clip, eps=eps, dtype=dtype)
def normalize_mi_ma(x, mi, ma, clip=False, eps=1e-20, dtype=np.float32):#dtype=np.float32
"""This function is adapted from Martin Weigert"""
if dtype is not None:
x = x.astype(dtype,copy=False)
mi = dtype(mi) if np.isscalar(mi) else mi.astype(dtype,copy=False)
ma = dtype(ma) if np.isscalar(ma) else ma.astype(dtype,copy=False)
eps = dtype(eps)
try:
import numexpr
x = numexpr.evaluate("(x - mi) / ( ma - mi + eps )")
except ImportError:
x = (x - mi) / ( ma - mi + eps )
if clip:
x = np.clip(x,0,1)
return x
def norm_minmse(gt, x, normalize_gt=True):
"""This function is adapted from Martin Weigert"""
"""
normalizes and affinely scales an image pair such that the MSE is minimized
Parameters
----------
gt: ndarray
the ground truth image
x: ndarray
the image that will be affinely scaled
normalize_gt: bool
set to True of gt image should be normalized (default)
Returns
-------
gt_scaled, x_scaled
"""
if normalize_gt:
gt = normalize(gt, 0.1, 99.9, clip=False).astype(np.float32, copy = False)
x = x.astype(np.float32, copy=False) - np.mean(x)
#x = x - np.mean(x)
gt = gt.astype(np.float32, copy=False) - np.mean(gt)
#gt = gt - np.mean(gt)
scale = np.cov(x.flatten(), gt.flatten())[0, 1] / np.var(x.flatten())
return gt, scale * x
# Open and create the csv file that will contain all the QC metrics
with open(QC_model_path+"/"+QC_model_name+"/Quality Control/QC_metrics_"+QC_model_name+".csv", "w", newline='') as file:
writer = csv.writer(file)
# Write the header in the csv file
writer.writerow(["image #","Prediction v. GT mSSIM","Input v. GT mSSIM", "Prediction v. GT NRMSE", "Input v. GT NRMSE", "Prediction v. GT PSNR", "Input v. GT PSNR"])
# Let's loop through the provided dataset in the QC folders
for i in os.listdir(Source_QC_folder):
if not os.path.isdir(os.path.join(Source_QC_folder,i)):
print('Running QC on: '+i)
# -------------------------------- Target test data (Ground truth) --------------------------------
test_GT = io.imread(os.path.join(Target_QC_folder, i))
# -------------------------------- Source test data --------------------------------
test_source = io.imread(os.path.join(Source_QC_folder,i))
# Normalize the images wrt each other by minimizing the MSE between GT and Source image
test_GT_norm,test_source_norm = norm_minmse(test_GT, test_source, normalize_gt=True)
# -------------------------------- Prediction --------------------------------
test_prediction = io.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/Prediction",i))
# Normalize the images wrt each other by minimizing the MSE between GT and prediction
test_GT_norm,test_prediction_norm = norm_minmse(test_GT, test_prediction, normalize_gt=True)
# -------------------------------- Calculate the metric maps and save them --------------------------------
# Calculate the SSIM maps
index_SSIM_GTvsPrediction, img_SSIM_GTvsPrediction = ssim(test_GT_norm, test_prediction_norm)
index_SSIM_GTvsSource, img_SSIM_GTvsSource = ssim(test_GT_norm, test_source_norm)
#Save ssim_maps
img_SSIM_GTvsPrediction_32bit = np.float32(img_SSIM_GTvsPrediction)
io.imsave(QC_model_path+'/'+QC_model_name+'/Quality Control/SSIM_GTvsPrediction_'+i,img_SSIM_GTvsPrediction_32bit)
img_SSIM_GTvsSource_32bit = np.float32(img_SSIM_GTvsSource)
io.imsave(QC_model_path+'/'+QC_model_name+'/Quality Control/SSIM_GTvsSource_'+i,img_SSIM_GTvsSource_32bit)
# Calculate the Root Squared Error (RSE) maps
img_RSE_GTvsPrediction = np.sqrt(np.square(test_GT_norm - test_prediction_norm))
img_RSE_GTvsSource = np.sqrt(np.square(test_GT_norm - test_source_norm))
# Save SE maps
img_RSE_GTvsPrediction_32bit = np.float32(img_RSE_GTvsPrediction)
img_RSE_GTvsSource_32bit = np.float32(img_RSE_GTvsSource)
io.imsave(QC_model_path+'/'+QC_model_name+'/Quality Control/RSE_GTvsPrediction_'+i,img_RSE_GTvsPrediction_32bit)
io.imsave(QC_model_path+'/'+QC_model_name+'/Quality Control/RSE_GTvsSource_'+i,img_RSE_GTvsSource_32bit)
# -------------------------------- Calculate the RSE metrics and save them --------------------------------
# Normalised Root Mean Squared Error (here it's valid to take the mean of the image)
NRMSE_GTvsPrediction = np.sqrt(np.mean(img_RSE_GTvsPrediction))
NRMSE_GTvsSource = np.sqrt(np.mean(img_RSE_GTvsSource))
# We can also measure the peak signal to noise ratio between the images
PSNR_GTvsPrediction = psnr(test_GT_norm,test_prediction_norm,data_range=1.0)
PSNR_GTvsSource = psnr(test_GT_norm,test_source_norm,data_range=1.0)
writer.writerow([i,str(index_SSIM_GTvsPrediction),str(index_SSIM_GTvsSource),str(NRMSE_GTvsPrediction),str(NRMSE_GTvsSource),str(PSNR_GTvsPrediction),str(PSNR_GTvsSource)])
# All data is now processed saved
Test_FileList = os.listdir(Source_QC_folder) # this assumes, as it should, that both source and target are named the same
plt.figure(figsize=(15,15))
# Currently only displays the last computed set, from memory
# Target (Ground-truth)
plt.subplot(3,3,1)
plt.axis('off')
img_GT = io.imread(os.path.join(Target_QC_folder, Test_FileList[-1]))
plt.imshow(img_GT)
plt.title('Target',fontsize=15)
# Source
plt.subplot(3,3,2)
plt.axis('off')
img_Source = io.imread(os.path.join(Source_QC_folder, Test_FileList[-1]))
plt.imshow(img_Source)
plt.title('Source',fontsize=15)
#Prediction
plt.subplot(3,3,3)
plt.axis('off')
img_Prediction = io.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/Prediction/", Test_FileList[-1]))
plt.imshow(img_Prediction)
plt.title('Prediction',fontsize=15)
#Setting up colours
cmap = plt.cm.CMRmap
#SSIM between GT and Source
plt.subplot(3,3,5)
#plt.axis('off')
plt.tick_params(
axis='both', # changes apply to the x-axis and y-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
top=False, # ticks along the top edge are off
left=False, # ticks along the left edge are off
right=False, # ticks along the right edge are off
labelbottom=False,
labelleft=False)
imSSIM_GTvsSource = plt.imshow(img_SSIM_GTvsSource, cmap = cmap, vmin=0, vmax=1)
plt.colorbar(imSSIM_GTvsSource,fraction=0.046, pad=0.04)
plt.title('Target vs. Source',fontsize=15)
plt.xlabel('mSSIM: '+str(round(index_SSIM_GTvsSource,3)),fontsize=14)
plt.ylabel('SSIM maps',fontsize=20, rotation=0, labelpad=75)
#SSIM between GT and Prediction
plt.subplot(3,3,6)
#plt.axis('off')
plt.tick_params(
axis='both', # changes apply to the x-axis and y-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
top=False, # ticks along the top edge are off
left=False, # ticks along the left edge are off
right=False, # ticks along the right edge are off
labelbottom=False,
labelleft=False)
imSSIM_GTvsPrediction = plt.imshow(img_SSIM_GTvsPrediction, cmap = cmap, vmin=0,vmax=1)
plt.colorbar(imSSIM_GTvsPrediction,fraction=0.046, pad=0.04)
plt.title('Target vs. Prediction',fontsize=15)
plt.xlabel('mSSIM: '+str(round(index_SSIM_GTvsPrediction,3)),fontsize=14)
#Root Squared Error between GT and Source
plt.subplot(3,3,8)
#plt.axis('off')
plt.tick_params(
axis='both', # changes apply to the x-axis and y-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
top=False, # ticks along the top edge are off
left=False, # ticks along the left edge are off
right=False, # ticks along the right edge are off
labelbottom=False,
labelleft=False)
imRSE_GTvsSource = plt.imshow(img_RSE_GTvsSource, cmap = cmap, vmin=0, vmax = 1)
plt.colorbar(imRSE_GTvsSource,fraction=0.046,pad=0.04)
plt.title('Target vs. Source',fontsize=15)
plt.xlabel('NRMSE: '+str(round(NRMSE_GTvsSource,3))+', PSNR: '+str(round(PSNR_GTvsSource,3)),fontsize=14)
#plt.title('Target vs. Source PSNR: '+str(round(PSNR_GTvsSource,3)))
plt.ylabel('RSE maps',fontsize=20, rotation=0, labelpad=75)
#Root Squared Error between GT and Prediction
plt.subplot(3,3,9)
#plt.axis('off')
plt.tick_params(
axis='both', # changes apply to the x-axis and y-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
top=False, # ticks along the top edge are off
left=False, # ticks along the left edge are off
right=False, # ticks along the right edge are off
labelbottom=False,
labelleft=False)
imRSE_GTvsPrediction = plt.imshow(img_RSE_GTvsPrediction, cmap = cmap, vmin=0, vmax=1)
plt.colorbar(imRSE_GTvsPrediction,fraction=0.046,pad=0.04)
plt.title('Target vs. Prediction',fontsize=15)
plt.xlabel('NRMSE: '+str(round(NRMSE_GTvsPrediction,3))+', PSNR: '+str(round(PSNR_GTvsPrediction,3)),fontsize=14)
plt.savefig(full_QC_model_path+'/Quality Control/QC_example_data.png',bbox_inches='tight',pad_inches=0)
qc_pdf_export()
```
# **6. Using the trained model**
---
<font size = 4>In this section the unseen data is processed using the trained model (in section 4). First, your unseen images are uploaded and prepared for prediction. After that your trained model from section 4 is activated and finally saved into your Google Drive.
## **6.1. Generate prediction(s) from unseen dataset**
---
<font size = 4>The current trained model (from section 4.2) can now be used to process images. If an older model needs to be used, please untick the **Use_the_current_trained_model** box and enter the name and path of the model to use. Predicted output images are saved in your **Result_folder** folder as restored image stacks (ImageJ-compatible TIFF images).
<font size = 4>**`Data_folder`:** This folder should contains the images that you want to predict using the network that you will train.
<font size = 4>**`Result_folder`:** This folder will contain the predicted output images.
<font size = 4>**`Data_type`:** Please indicate if the images you want to predict are single images or stacks
```
Single_Images = 1
Stacks = 2
#@markdown ### Provide the path to your dataset and to the folder where the prediction will be saved, then play the cell to predict output on your unseen images.
#@markdown ###Path to data to analyse and where predicted output should be saved:
Data_folder = "" #@param {type:"string"}
Result_folder = "" #@param {type:"string"}
#@markdown ###Are your data single images or stacks?
Data_type = Single_Images #@param ["Single_Images", "Stacks"] {type:"raw"}
# model name and path
#@markdown ###Do you want to use the current trained model?
Use_the_current_trained_model = True #@param {type:"boolean"}
#@markdown ###If not, please provide the path to the model folder:
Prediction_model_folder = "" #@param {type:"string"}
#Here we find the loaded model name and parent path
Prediction_model_name = os.path.basename(Prediction_model_folder)
Prediction_model_path = os.path.dirname(Prediction_model_folder)
if (Use_the_current_trained_model):
print("Using current trained network")
Prediction_model_name = model_name
Prediction_model_path = model_path
full_Prediction_model_path = Prediction_model_path+'/'+Prediction_model_name+'/'
if os.path.exists(full_Prediction_model_path):
print("The "+Prediction_model_name+" network will be used.")
else:
print(bcolors.WARNING +'!! WARNING: The chosen model does not exist !!')
print('Please make sure you provide a valid model path and model name before proceeding further.')
#Activate the pretrained model.
config = None
model = N2V(config, Prediction_model_name, basedir=Prediction_model_path)
thisdir = Path(Data_folder)
outputdir = Path(Result_folder)
# r=root, d=directories, f = files
for r, d, f in os.walk(thisdir):
for file in f:
if ".tif" in file:
print(os.path.join(r, file))
if Data_type == 1 :
print("Single images are now beeing predicted")
# Loop through the files
for r, d, f in os.walk(thisdir):
for file in f:
base_filename = os.path.basename(file)
input_train = imread(os.path.join(r, file))
pred_train = model.predict(input_train, axes='YX', n_tiles=(2,1))
save_tiff_imagej_compatible(os.path.join(outputdir, base_filename), pred_train, axes='YX')
print("Images saved into folder:", Result_folder)
if Data_type == 2 :
print("Stacks are now beeing predicted")
for r, d, f in os.walk(thisdir):
for file in f:
base_filename = os.path.basename(file)
timelapse = imread(os.path.join(r, file))
n_timepoint = timelapse.shape[0]
prediction_stack = np.zeros((n_timepoint, timelapse.shape[1], timelapse.shape[2]))
for t in range(n_timepoint):
img_t = timelapse[t]
prediction_stack[t] = model.predict(img_t, axes='YX', n_tiles=(2,1))
prediction_stack_32 = img_as_float32(prediction_stack, force_copy=False)
imsave(os.path.join(outputdir, base_filename), prediction_stack_32)
```
## **6.2. Assess predicted output**
---
```
# @markdown ##Run this cell to display a randomly chosen input and its corresponding predicted output.
# This will display a randomly chosen dataset input and predicted output
random_choice = random.choice(os.listdir(Data_folder))
x = imread(Data_folder+"/"+random_choice)
os.chdir(Result_folder)
y = imread(Result_folder+"/"+random_choice)
if Data_type == 1 :
f=plt.figure(figsize=(16,8))
plt.subplot(1,2,1)
plt.imshow(x, interpolation='nearest')
plt.title('Input')
plt.axis('off');
plt.subplot(1,2,2)
plt.imshow(y, interpolation='nearest')
plt.title('Predicted output')
plt.axis('off');
if Data_type == 2 :
f=plt.figure(figsize=(16,8))
plt.subplot(1,2,1)
plt.imshow(x[1], interpolation='nearest')
plt.title('Input')
plt.axis('off');
plt.subplot(1,2,2)
plt.imshow(y[1], interpolation='nearest')
plt.title('Predicted output')
plt.axis('off');
```
## **6.3. Download your predictions**
---
<font size = 4>**Store your data** and ALL its results elsewhere by downloading it from Google Drive and after that clean the original folder tree (datasets, results, trained model etc.) if you plan to train or use new networks. Please note that the notebook will otherwise **OVERWRITE** all files which have the same name.
#**Thank you for using Noise2Void 2D!**
| github_jupyter |
**Connect With Me in Linkedin :-** https://www.linkedin.com/in/dheerajkumar1997/
## Weight of evidence
Weight of Evidence (WoE) was developed primarily for the credit and financial industries to help build more predictive models to evaluate the risk of loan default. That is, to predict how likely the money lent to a person or institution is to be lost. Thus, Weight of Evidence is a measure of the "strength” of a grouping technique to separate good and bad risk (default).
It is computed from the basic odds ratio: ln( (Proportion of Good Credit Outcomes) / (Proportion of Bad Credit Outcomes))
WoE will be 0 if the P(Goods) / P(Bads) = 1. That is, if the outcome is random for that group. If P(Bads) > P(Goods) the odds ratio will be < 1 and the WoE will be < 0; if, on the other hand, P(Goods) > P(Bads) in a group, then WoE > 0.
WoE is well suited for Logistic Regression, because the Logit transformation is simply the log of the odds, i.e., ln(P(Goods)/P(Bads)). Therefore, by using WoE-coded predictors in logistic regression, the predictors are all prepared and coded to the same scale, and the parameters in the linear logistic regression equation can be directly compared.
The WoE transformation has three advantages:
- It establishes a monotonic relationship to the dependent variable.
- It orders the categories on a "logistic" scale which is natural for logistic regression
- The transformed variables, can then be compared because they are on the same scale. Therefore, it is possible to determine which one is more predictive.
The WoE also has three drawbacks:
- May incur in loss of information (variation) due to binning to few categories (we will discuss this further in the discretisation section)
- It does not take into account correlation between independent variables
- Prone to cause over-fitting
For more details follow this link:
http://documentation.statsoft.com/StatisticaHelp.aspx?path=WeightofEvidence/WeightofEvidenceWoEIntroductoryOverview
Let's see how to implement WoE in python
```
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
%matplotlib inline
# let's load again the titanic dataset
data = pd.read_csv('titanic.csv', usecols=['Cabin', 'Survived'])
data.head()
# let's first fill NA values with an additional label
data.Cabin.fillna('Missing', inplace=True)
data.head()
# Cabin has indeed a lot of labels, here for simplicity, I will capture the first letter of the cabin,
# but the procedure could be done as well without any prior variable manipulation
len(data.Cabin.unique())
# Now we extract the first letter of the cabin
data['Cabin'] = data['Cabin'].astype(str).str[0]
data.head()
# check the labels
data.Cabin.unique()
```
### Important
The calculation of the WoE to replace the labels should be done considering the ONLY the training set, and then expanded it to the test set.
See below.
```
# Let's divide into train and test set
X_train, X_test, y_train, y_test = train_test_split(data[['Cabin', 'Survived']], data.Survived, test_size=0.3,
random_state=0)
X_train.shape, X_test.shape
# now we calculate the probability of target=1
X_train.groupby(['Cabin'])['Survived'].mean()
# let's make a dataframe with the above calculation
prob_df = X_train.groupby(['Cabin'])['Survived'].mean()
prob_df = pd.DataFrame(prob_df)
prob_df
# and now the probability of target = 0
# and we add it to the dataframe
prob_df = X_train.groupby(['Cabin'])['Survived'].mean()
prob_df = pd.DataFrame(prob_df)
prob_df['Died'] = 1-prob_df.Survived
prob_df
# since the log of zero is not defined, let's set this number to something small and non-zero
prob_df.loc[prob_df.Survived == 0, 'Survived'] = 0.00001
prob_df
# now we calculate the WoE
prob_df['WoE'] = np.log(prob_df.Survived/prob_df.Died)
prob_df
# and we create a dictionary to re-map the variable
prob_df['WoE'].to_dict()
# and we make a dictionary to map the orignal variable to the WoE
# same as above but we capture the dictionary in a variable
ordered_labels = prob_df['WoE'].to_dict()
# replace the labels with the WoE
X_train['Cabin_ordered'] = X_train.Cabin.map(ordered_labels)
X_test['Cabin_ordered'] = X_test.Cabin.map(ordered_labels)
# check the results
X_train.head()
# plot the original variable
fig = plt.figure()
fig = X_train.groupby(['Cabin'])['Survived'].mean().plot()
fig.set_title('Normal relationship between variable and target')
fig.set_ylabel('Survived')
# plot the transformed result: the monotonic variable
fig = plt.figure()
fig = X_train.groupby(['Cabin_ordered'])['Survived'].mean().plot()
fig.set_title('Monotonic relationship between variable and target')
fig.set_ylabel('Survived')
```
As you can see in the above plot, there is now a monotonic relationship between the variable Cabin and probability of survival. The higher the Cabin number, the more likely the person was to survive.
### Note
Monotonic does not mean strictly linear. Monotonic means that it increases constantly, or it decreases constantly.
**Connect With Me in Linkedin :-** https://www.linkedin.com/in/dheerajkumar1997/
| github_jupyter |
# Matrix Multiplication
Matrix multiplication involves quite a few steps in terms of writing code. But remember that the basics of matrix multiplicaiton involve taking a row in matrix A and finding the dot product with a column in matrix B.
So you are going to write a function to extract a row from matrix A, extract a column from matrix B, and then calculate the dot product of the row and column.
Then you can use these functions to output the results of multiplying two matrices together.
Here is a general outline of the code that you will be writing. Assume you are calculating the product of
$$A\times{B}$$
* Write a nested for loop that iterates through the m rows of matrix A and the p columns of matrix B
* Initialize an empty list that will hold values of the final matrix
* Starting with the first row of matrix A, find the dot product with the first column of matrix B
* Append the result to the empty list representing a row in the final matrix
* Now find the dot product between the first row of matrix A and the second column of matrix B
* Append the result to the row list
* Keep going until you get to the last column of B
* Append the row list to the output variable. Reinitialize the row list so that it is empty.
* Then start on row two of matrix A. Iterate through all of the columns of B taking the dot product
* etc...
# Breaking the Process down into steps
Rather than writing all of the matrix multiplication code in one function, you are going to break the process down into several functions:
**get_row(matrix, row_number)**
Because you are going to need the rows from matrix A, you will use a function called get_row that takes in a matrix and row number and then returns a row of a matrix. We have provided this function for you.
**get_column(matrix, column_number)**
Likewise, you will need the columns from matrix B. So you will write a similar function that receives a matrix and column number and then returns a column from matrix B.
**dot_product(vectorA, vectorB)**
You have actually already written this function in a prevoius exercise. The dot_product function calculates the dot product of two vectors.
**matrix_multiply(matrixA, matrixB)**
This is the function that will calculate the product of the two matrices. You will need to write a nested for loop that iterates through the rows of A and columns of B. For each row-column combination, you will calculate the dot product and then append the result to the output matrix.
# get_row
The first function is the get_row function. We have provided this function for you.
The get_row function has two inputs and one output.
INPUTS
* matrix
* row number
OUTPUT
* a list, which represents one row of the matrix
In Python, a matrix is a list of lists. If you have a matrix like this one:
```python
m = [
[5, 9, 11, 2],
[3, 2, 99, 3],
[7, 1, 8, 2]
]
```
then row one would be accessed by
```
m[0]
```
row two would be
```
m[1]
```
and row three would be
```
m[2]
```
```
## TODO: Run this code cell to load the get_row function
## You do not need to modify this cell
def get_row(matrix, row):
return matrix[row]
```
# Getting a Column from a Matrix
Since matrices are stored as lists of lists, it's relatively simple to extract a row. If A is a matrix, then
```
A[0]
```
will output the first row of the matrix.
```
A[1]
```
outputs the second row of the matrix.
But what if you want to get a matrix column? It's not as convenient. To get the values of the first column, you would need to output:
```
A[0][0]
A[1][0]
A[2][0]
...
A[m][0]
```
For matrix multiplication, you will need to have access to the columns of the B matrix. So write a function called get_column that receives a matrix and a column number indexed from zero. The function then outputs a vector as a list that contains the column. For example
```
get_column([
[1, 2, 4],
[7, 8, 1],
[5, 2, 1]
],
1)
```
would output the second column
```
[2, 8, 2]
```
# get_column
The get_column function is similar to the get_row function except now you will return a column.
Here are the inputs and outputs of the function
INPUTS
* matrix
* column number
OUPUT
* a list, which represents a column of the matrix
Getting a matrix column is actually more difficult than getting a matrix row.
Take a look again at this example matrix:
```python
m = [
[5, 9, 11, 2],
[3, 2, 99, 3],
[7, 1, 8, 2]
]
```
What if you wanted to extract the first column as a list [5, 3, 7]. You can't actually get that column directly like you could with a row.
You'll need to think about using a for statement to iterate through the rows and grab the specific values that you want for your column list.
```
### TODO: Write a function that receives a matrix and a column number.
### the output should be the column in the form of a list
### Example input:
# matrix = [
# [5, 9, 11, 2],
# [3, 2, 99, 3],
# [7, 1, 8, 2]
# ]
#
# column_number = 1
### Example output:
# [9, 2, 1]
#
def get_column(matrix, column_number):
return [row[column_number] for row in matrix]
### TODO: Run this code to test your get_column function
assert get_column([[1, 2, 4],
[7, 8, 1],
[5, 2, 1]], 1) == [2, 8, 2]
assert get_column([[5]], 0) == [5]
```
### Dot Product of Two Vectors
As part of calculating the product of a matrix, you need to do calculate the dot product of a row from matrix A with a column from matrix B. You will do this process many times, so why not abstract the process into a function?
If you consider a single row of A to be a vector and a single row of B to also be a vector, you can calculate the dot product.
Remember that for matrix multiplication to be valid, A is size m x n while B is size n x p. The number of columns in A must equal the number of rows in B, which makes taking the dot product between a row of A and column of B possible.
As a reminder, the dot product of `<a1, a2, a3, a4>` and `<b1, b2, b3, b4>` is equal to
`a1*b1 + a2*b2 + a3*b3 + a4*b4`
```
### TODO: Write a function called dot_product() that
### has two vectors as inputs and outputs the dot product of the
### two vectors. First, you will need to do element-wise
### multiplication and then sum the results.
### HINT: You wrote this function previously in the vector coding
### exercises
def dot_product(vector_one, vector_two):
return sum([x[0]*x[1] for x in zip(vector_one, vector_two)])
### TODO: Run this cell to test your results
assert dot_product([4, 5, 1], [2, 1, 5]) == 18
assert dot_product([6], [7]) == 42
```
### Matrix Multiplication
Now you will write a function to carry out matrix multiplication
between two matrices.
If you have an m x n matrix and an n x p matrix, your result will be m x p.
Your strategy could involve looping through an empty n x p matrix and filling in each elements value.
```
### TODO: Write a function called matrix_multiplication that takes
### two matrices,multiplies them together and then returns
### the results
###
### Make sure that your function can handle matrices that contain
### only one row or one column. For example,
### multiplying two matrices of size (4x1)x(1x4) should return a
### 4x4 matrix
def matrix_multiplication(matrixA, matrixB):
### TODO: store the number of rows in A and the number
### of columns in B. This will be the size of the output
### matrix
### HINT: The len function in Python will be helpful
m_rows = len(matrixA)
p_columns = len(matrixB[0])
# empty list that will hold the product of AxB
result = []
### TODO: Write a for loop within a for loop. The outside
### for loop will iterate through m_rows.
### The inside for loop will iterate through p_columns.
### TODO: As you iterate through the m_rows and p_columns,
### use your get_row function to grab the current A row
### and use your get_column function to grab the current
### B column.
### TODO: Calculate the dot product of the A row and the B column
### TODO: Append the dot product to an empty list called row_result.
### This list will accumulate the values of a row
### in the result matrix
### TODO: After iterating through all of the columns in matrix B,
### append the row_result list to the result variable.
### Reinitialize the row_result to row_result = [].
### Your for loop will move down to the next row
### of matrix A.
### The loop will iterate through all of the columns
### taking the dot product
### between the row in A and each column in B.
### TODO: return the result of AxB
for i in range(m_rows):
row_result = []
for j in range(p_columns):
row_result.append(dot_product(get_row(matrixA,i),get_column(matrixB, j)))
result.append(row_result)
return result
### TODO: Run this code cell to test your results
assert matrix_multiplication([[5], [2]], [[5, 1]]) == [[25, 5], [10, 2]]
assert matrix_multiplication([[5, 1]], [[5], [2]]) == [[27]]
assert matrix_multiplication([[4]], [[3]]) == [[12]]
assert matrix_multiplication([[2, 1, 8, 2, 1], [5, 6, 4, 2, 1]], [[1, 7, 2], [2, 6, 3], [3, 1, 1], [1, 20, 1], [7, 4, 16]]) == [[37, 72, 33], [38, 119, 50]]
```
| github_jupyter |
```
import tensorflow as tf
tf.logging.set_verbosity(tf.logging.WARN)
import pickle
import numpy as np
import os
from sklearn.model_selection import train_test_split
from sklearn.metrics import f1_score
from sklearn.metrics import accuracy_score
import os
from tensorflow.python.client import device_lib
from collections import Counter
import time
f = open('../../Glove/word_embedding_glove', 'rb')
word_embedding = pickle.load(f)
f.close()
word_embedding = word_embedding[: len(word_embedding)-1]
f = open('../../Glove/vocab_glove', 'rb')
vocab = pickle.load(f)
f.close()
word2id = dict((w, i) for i,w in enumerate(vocab))
id2word = dict((i, w) for i,w in enumerate(vocab))
unknown_token = "UNKNOWN_TOKEN"
f = open("../../../dataset/sense/dict_sense-keys", 'rb')
dict_sense_keys = pickle.load(f)
f.close()
f = open("../../../dataset/sense/dict_word-sense", 'rb')
dict_word_sense = pickle.load(f)
f.close()
# Model Description
sense_word = 'force'
model_name = 'model-4-multigpu-1'
sense_word_dir = '../output/' + sense_word
model_dir = sense_word_dir + '/' + model_name
save_dir = os.path.join(model_dir, "save/")
log_dir = os.path.join(model_dir, "log")
if not os.path.exists(sense_word_dir):
os.mkdir(sense_word_dir)
if not os.path.exists(model_dir):
os.mkdir(model_dir)
if not os.path.exists(save_dir):
os.mkdir(save_dir)
if not os.path.exists(log_dir):
os.mkdir(log_dir)
f = open("../../../dataset/checkwords/"+ sense_word + "_data", 'rb')
data = pickle.load(f)
f.close()
data_y = []
for i in range(len(data)):
data_y.append(dict_sense_keys[data[i][0]][3])
sense_count = Counter(data_y)
sense_count = sense_count.most_common()[:5]
vocab_sense = [k for k,v in sense_count]
vocab_sense = sorted(vocab_sense, key=lambda x:int(x[0]))
print(sense_count)
print(vocab_sense)
def make_mask_matrix(sense_word,vocab_sense):
mask_mat = []
sense_list = [int(string[0]) for string in vocab_sense]
sense_count = list(Counter(sense_list).values())
start=0
prev=0
for i in range(len(set(sense_list))):
temp_row=[0]*len(sense_list)
for j in range(len(sense_list)):
if j>=start and j<sense_count[i]+prev:
temp_row[j]= 0
else:
temp_row[j]= -10
start+=sense_count[i]
prev+=sense_count[i]
mask_mat.append(temp_row)
return mask_mat
mask_mat = make_mask_matrix(sense_word,vocab_sense)
print(mask_mat)
data_x = []
data_label = []
data_pos = []
for i in range(len(data)):
if dict_sense_keys[data[i][0]][3] in vocab_sense:
data_x.append(data[i][1])
data_label.append(dict_sense_keys[data[i][0]][3])
data_pos.append(dict_sense_keys[data[i][0]][1])
print(len(data_label), len(data_y))
# vocab_sense = dict_word_sense[sense_word]
sense2id = dict((s, i) for i,s in enumerate(vocab_sense))
id2sense = dict((i, s) for i,s in enumerate(vocab_sense))
count_pos = Counter(data_pos)
count_pos = count_pos.most_common()
vocab_pos = [int(k) for k,v in count_pos]
vocab_pos = sorted(vocab_pos, key=lambda x:int(x))
pos2id = dict((str(s), i) for i,s in enumerate(vocab_pos))
id2pos = dict((i, str(s)) for i,s in enumerate(vocab_pos))
print(vocab_pos)
max_len = 0
for i in range(len(data_x)):
max_len = max(max_len, len(data_x[i]))
if(len(data_x[i])>200):
print(i)
print("max_len: ", max_len)
# Parameters
mode = 'train'
num_senses = len(vocab_sense)
num_pos = len(vocab_pos)
batch_size = 32
vocab_size = len(vocab)
unk_vocab_size = 1
word_emb_size = len(word_embedding[0])
max_sent_size = max(200, max_len)
hidden_size = 100
keep_prob = 0.5
l2_lambda = 0.001
init_lr = 0.005
decay_steps = 500
decay_rate = 0.96
clip_norm = 1
clipping = True
moving_avg_deacy = 0.999
num_gpus = 6
lambda_loss_pos = 5
index = []
for i in range(len(data_x)):
index.append(i)
index_train, index_val, label_train, label_val = train_test_split(index, data_label, train_size=0.8, shuffle=True, stratify=data_label, random_state=0)
data_x = np.array(data_x)
data_pos = np.array(data_pos)
x_train = data_x[index_train]
pos_train = data_pos[index_train]
x_val = data_x[index_val]
pos_val = data_pos[index_val]
def average_gradients(tower_grads):
average_grads = []
for grad_and_vars in zip(*tower_grads):
# Note that each grad_and_vars looks like the following:
# ((grad0_gpu0, var0_gpu0), ... , (grad0_gpuN, var0_gpuN))
grads = []
for g, _ in grad_and_vars:
# Add 0 dimension to the gradients to represent the tower.
expanded_g = tf.expand_dims(g, 0)
# Append on a 'tower' dimension which we will average over below.
grads.append(expanded_g)
# Average over the 'tower' dimension.
grad = tf.concat(grads, 0)
grad = tf.reduce_mean(grad, 0)
# Keep in mind that the Variables are redundant because they are shared
# across towers. So .. we will just return the first tower's pointer to
# the Variable.
v = grad_and_vars[0][1]
grad_and_var = (grad, v)
average_grads.append(grad_and_var)
return average_grads
# MODEL
tower_grads = []
losses = []
predictions = []
predictions_pos = []
total_trans_params = []
x = tf.placeholder('int32', [num_gpus, batch_size, max_sent_size], name="x")
y = tf.placeholder('int32', [num_gpus, batch_size], name="y")
y_pos = tf.placeholder('int32', [num_gpus, batch_size], name="y_pos")
x_mask = tf.placeholder('bool', [num_gpus, batch_size, max_sent_size], name='x_mask')
is_train = tf.placeholder('bool', [], name='is_train')
mask_matrix = tf.constant(value=mask_mat, shape=list(np.array(mask_mat).shape), dtype='float32') # mask_matrix
word_emb_mat = tf.placeholder('float', [None, word_emb_size], name='emb_mat')
input_keep_prob = tf.cond(is_train,lambda:keep_prob, lambda:tf.constant(1.0))
global_step = tf.Variable(0, trainable=False, name="global_step")
learning_rate = tf.train.exponential_decay(init_lr, global_step, decay_steps, decay_rate, staircase=True)
with tf.variable_scope("word_embedding"):
unk_word_emb_mat = tf.get_variable("word_emb_mat", dtype='float', shape=[unk_vocab_size, word_emb_size], initializer=tf.contrib.layers.xavier_initializer(uniform=True, seed=0, dtype=tf.float32))
final_word_emb_mat = tf.concat([word_emb_mat, unk_word_emb_mat], 0)
with tf.variable_scope(tf.get_variable_scope()):
for gpu_idx in range(num_gpus):
with tf.name_scope("model_{}".format(gpu_idx)) as scope, tf.device('/gpu:%d' % i):
if gpu_idx > 0:
tf.get_variable_scope().reuse_variables()
with tf.name_scope("word"):
Wx = tf.nn.embedding_lookup(final_word_emb_mat, x[gpu_idx])
x_len = tf.reduce_sum(tf.cast(x_mask[gpu_idx], 'int32'), 1)
with tf.variable_scope("lstm1"):
cell_fw1 = tf.contrib.rnn.BasicLSTMCell(hidden_size,state_is_tuple=True)
cell_bw1 = tf.contrib.rnn.BasicLSTMCell(hidden_size,state_is_tuple=True)
d_cell_fw1 = tf.contrib.rnn.DropoutWrapper(cell_fw1, input_keep_prob=input_keep_prob)
d_cell_bw1 = tf.contrib.rnn.DropoutWrapper(cell_bw1, input_keep_prob=input_keep_prob)
(fw_h1, bw_h1), _ = tf.nn.bidirectional_dynamic_rnn(d_cell_fw1, d_cell_bw1, Wx, sequence_length=x_len, dtype='float', scope='lstm1')
h1 = tf.concat([fw_h1, bw_h1], 2)
with tf.variable_scope("lstm2"):
cell_fw2 = tf.contrib.rnn.BasicLSTMCell(hidden_size,state_is_tuple=True)
cell_bw2 = tf.contrib.rnn.BasicLSTMCell(hidden_size,state_is_tuple=True)
d_cell_fw2 = tf.contrib.rnn.DropoutWrapper(cell_fw2, input_keep_prob=input_keep_prob)
d_cell_bw2 = tf.contrib.rnn.DropoutWrapper(cell_bw2, input_keep_prob=input_keep_prob)
(fw_h2, bw_h2), _ = tf.nn.bidirectional_dynamic_rnn(d_cell_fw2, d_cell_bw2, h1, sequence_length=x_len, dtype='float', scope='lstm2')
h = tf.concat([fw_h2, bw_h2], 2)
def attention(input_x, input_mask, W_att):
h_masked = tf.boolean_mask(input_x, input_mask)
h_tanh = tf.tanh(h_masked)
u = tf.matmul(h_tanh, W_att)
a = tf.nn.softmax(u)
c = tf.reduce_sum(tf.multiply(h_tanh, a), 0)
return c
with tf.variable_scope("attention"):
W_att = tf.get_variable("W_att", shape=[2*hidden_size, 1], initializer=tf.truncated_normal_initializer(mean=0.0, stddev=0.1, seed=0))
c = tf.expand_dims(attention(h[0], x_mask[gpu_idx][0], W_att), 0)
for i in range(1, batch_size):
c = tf.concat([c, tf.expand_dims(attention(h[i], x_mask[gpu_idx][i], W_att), 0)], 0)
with tf.variable_scope("attention_pos"):
W_attp = tf.get_variable("W_attp", shape=[2*hidden_size, 1], initializer=tf.truncated_normal_initializer(mean=0.0, stddev=0.1, seed=0))
cp = tf.expand_dims(attention(h[0], x_mask[gpu_idx][0], W_attp), 0)
for i in range(1, batch_size):
cp = tf.concat([cp, tf.expand_dims(attention(h1[i], x_mask[gpu_idx][i], W_attp), 0)], 0)
with tf.variable_scope("softmax_layer_pos"):
Wp = tf.get_variable("Wp", shape=[2*hidden_size, num_senses], initializer=tf.truncated_normal_initializer(mean=0.0, stddev=0.1, seed=0))
bp = tf.get_variable("bp", shape=[num_senses], initializer=tf.zeros_initializer())
drop_cp = tf.nn.dropout(cp, input_keep_prob)
logits_pos = tf.matmul(drop_cp, Wp) + bp
prediction_pos = tf.argmax(logits_pos, 1)
predictions_pos.append(prediction_pos)
final_masking = tf.nn.embedding_lookup(mask_matrix, prediction_pos)
with tf.variable_scope("softmax_layer"):
W = tf.get_variable("W", shape=[2*hidden_size, num_senses], initializer=tf.truncated_normal_initializer(mean=0.0, stddev=0.1, seed=0))
b = tf.get_variable("b", shape=[num_senses], initializer=tf.zeros_initializer())
drop_c = tf.nn.dropout(c, input_keep_prob)
logits = tf.matmul(drop_c, W) + b
masked_logits = logits + final_masking
predictions.append(tf.argmax(masked_logits, 1))
loss_pos = lambda_loss_pos * tf.reduce_mean(tf.nn.sparse_softmax_cross_entropy_with_logits(logits=logits_pos, labels=y_pos[gpu_idx]))
loss = tf.reduce_mean(tf.nn.sparse_softmax_cross_entropy_with_logits(logits=masked_logits, labels=y[gpu_idx]))
l2_loss = l2_lambda * tf.losses.get_regularization_loss()
total_loss = loss + l2_loss + loss_pos
tf.summary.scalar("loss_{}".format(gpu_idx), total_loss)
summaries = tf.get_collection(tf.GraphKeys.SUMMARIES, scope)
optimizer = tf.train.AdamOptimizer(learning_rate)
grads_vars = optimizer.compute_gradients(total_loss)
clipped_grads = grads_vars
if(clipping == True):
clipped_grads = [(tf.clip_by_norm(grad, clip_norm), var) for grad, var in clipped_grads]
tower_grads.append(clipped_grads)
losses.append(total_loss)
tower_grads = average_gradients(tower_grads)
losses = tf.add_n(losses)/len(losses)
apply_grad_op = optimizer.apply_gradients(tower_grads, global_step=global_step)
summaries.append(tf.summary.scalar('total_loss', losses))
summaries.append(tf.summary.scalar('learning_rate', learning_rate))
for var in tf.trainable_variables():
summaries.append(tf.summary.histogram(var.op.name, var))
variable_averages = tf.train.ExponentialMovingAverage(moving_avg_deacy, global_step)
variables_averages_op = variable_averages.apply(tf.trainable_variables())
train_op = tf.group(apply_grad_op, variables_averages_op)
saver = tf.train.Saver(tf.global_variables())
summary = tf.summary.merge(summaries)
os.environ["CUDA_DEVICE_ORDER"]="PCI_BUS_ID" # see issue #152
os.environ["CUDA_VISIBLE_DEVICES"]="3"
# print (device_lib.list_local_devices())
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
config.allow_soft_placement = True
sess = tf.Session(config=config)
sess.run(tf.global_variables_initializer()) # For initializing all the variables
summary_writer = tf.summary.FileWriter(log_dir, sess.graph) # For writing Summaries
def data_prepare(x, y, p):
num_examples = len(x)
xx = np.zeros([num_examples, max_sent_size], dtype=int)
xx_mask = np.zeros([num_examples, max_sent_size], dtype=bool)
yy = np.zeros([num_examples], dtype=int)
pp = np.zeros([num_examples], dtype=int)
for j in range(num_examples):
for i in range(max_sent_size):
if(i>=len(x[j])):
break
w = x[j][i]
xx[j][i] = word2id[w] if w in word2id else word2id['UNKNOWN_TOKEN']
xx_mask[j][i] = True
yy[j] = sense2id[y[j]]
pp[j] = pos2id[p[j]]
return xx, xx_mask, yy, pp
def model(xx, yy, mask, pp, train_cond=True):
num_batches = int(len(xx)/(batch_size*num_gpus))
_losses = 0
preds = []
preds_pos = []
for j in range(num_batches):
s = j * batch_size * num_gpus
e = (j+1) * batch_size * num_gpus
xx_re = xx[s:e].reshape([num_gpus, batch_size, -1])
yy_re = yy[s:e].reshape([num_gpus, batch_size])
pp_re = pp[s:e].reshape([num_gpus, batch_size])
mask_re = mask[s:e].reshape([num_gpus, batch_size, -1])
feed_dict = {x:xx_re, y:yy_re, y_pos:pp_re, x_mask:mask_re, is_train:train_cond, input_keep_prob:keep_prob, word_emb_mat:word_embedding}
if(train_cond==True):
_, _loss, step, _summary = sess.run([train_op, losses, global_step, summary], feed_dict)
summary_writer.add_summary(_summary, step)
# print("Steps:{}".format(step), ", Loss: {}".format(_loss))
else:
_loss, pred, pred_pos = sess.run([losses, predictions, predictions_pos], feed_dict)
for i in range(num_gpus):
preds.append(pred[i])
preds_pos.append(pred_pos[i])
_losses +=_loss
if(train_cond==False):
y_pred = []
pos_pred = []
for pred in preds:
for bt in pred:
y_pred.append(bt)
for pred in preds_pos:
for bt in pred:
pos_pred.append(bt)
return _losses/num_batches, y_pred, pos_pred
return _losses/num_batches, step
def eval_score(yy, pred, pp, pred_pos):
num_batches = int(len(yy)/(batch_size*num_gpus))
f1 = f1_score(yy[:batch_size*num_batches*num_gpus], pred, average='macro')
f1_pos = f1_score(pp[:batch_size*num_batches*num_gpus], pred_pos, average='macro')
accu = accuracy_score(yy[:batch_size*num_batches*num_gpus], pred)
accu_pos = accuracy_score(pp[:batch_size*num_batches*num_gpus], pred_pos)
return f1*100, accu*100, f1_pos*100, accu_pos*100
x_id_train, mask_train, y_train, pos_id_train = data_prepare(x_train, label_train, pos_train)
x_id_val, mask_val, y_val, pos_id_val = data_prepare(x_val, label_val,pos_val)
num_epochs = 60
log_period = 5
for i in range(num_epochs):
random = np.random.choice(len(y_train), size=(len(y_train)), replace=False)
x_id_train = x_id_train[random]
y_train = y_train[random]
mask_train = mask_train[random]
pos_id_train = pos_id_train[random]
start_time = time.time()
train_loss, step = model(x_id_train, y_train, mask_train, pos_id_train)
time_taken = time.time() - start_time
print("Epoch:", i+1,"Step:", step, "loss:{0:.4f}".format(train_loss), ", Time: {0:.4f}".format(time_taken))
if((i+1)%log_period==0):
saver.save(sess, save_path=save_dir)
print("Model Saved")
start_time = time.time()
train_loss, train_pred, train_pred_pos = model(x_id_train, y_train, mask_train, pos_id_train, train_cond=False)
f1_, accu_, f1_pos_, accu_pos_ = eval_score(y_train, train_pred, pos_id_train, train_pred_pos)
time_taken = time.time() - start_time
print("Train: F1 Score:{0:.4f}".format(f1_), "Accuracy:{0:.4f}".format(accu_), " POS: F1 Score:{0:.4f}".format(f1_pos_), "Accuracy:{0:.4f}".format(accu_pos_), "Loss:{0:.4f}".format(train_loss), ", Time: {0:.4f}".format(time_taken))
start_time = time.time()
val_loss, val_pred, val_pred_pos = model(x_id_val, y_val, mask_val, pos_id_val, train_cond=False)
f1_, accu_, f1_pos_, accu_pos_ = eval_score(y_val, val_pred, pos_id_val, val_pred_pos)
time_taken = time.time() - start_time
print("Val: F1 Score:{0:.4f}".format(f1_), "Accuracy:{0:.4f}".format(accu_), " POS: F1 Score:{0:.4f}".format(f1_pos_), "Accuracy:{0:.4f}".format(accu_pos_), "Loss:{0:.4f}".format(val_loss), ", Time: {0:.4f}".format(time_taken))
saver.restore(sess, save_dir)
```
| github_jupyter |
# Tensorflow Regression
Regression algorithm tries to find the function that best maps an input to an output in the simplest way, without overcomplicating things.
* Input can be discrete or continuous, but the output is always continuous
* Classification is for discrete outputs
How well is the algorithm working and how do we find the best function? We want a function that is not biased towards the training data it learned from and we don't want the results to wildly vary just because the real data is slightly different than the training set. We want it to generalize to unseen data as well.
**Variance** - indicates how sensitive a prediction is to the training set
* low variance is desired because it shouldn't matter how we choose the training set
* measures how badly the reponses vary
**Bias** - indicates the strength of the assumptions made on the training set
* low bias is desired to prevent overfitting in order to make a more generalized model
* measures how far off the the model is from the truth
**Cost Function** is used to evaluate each candidate solution
* Higher cost means a worse solution, want the lowest cost
* Tensorflow loops through all the data (an epoch) looking for the best possible value
* Any cost function can be used, typically sum of squared errors:
* the error difference between each data point and the chosen solution is squared (to penalize larger errors) and then added together to get a single "score" for that solution
* the lowest score ends up being the best possible solution
## Linear Regression
```
%matplotlib inline
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
learning_rate = 0.01 # Hyperparameters
training_epochs = 100
x_train = np.linspace(-1, 1, 101) # Dataset
y_train = 2 * x_train + np.random.randn(*x_train.shape) * 0.33
X = tf.placeholder(tf.float32) # tf placeholder nodes for input/output
Y = tf.placeholder(tf.float32)
w = tf.Variable(0.0, name="weights") # Weights variable
def model(X, w): # defines model as Y = wX
return tf.multiply(X, w)
y_model = model(X, w) # Cost Function
cost = tf.square(Y-y_model)
# Defines the operation to be called on each iteration of the learning algorithm
train_op = tf.train.GradientDescentOptimizer(learning_rate).minimize(cost)
sess = tf.Session() # Setup the tf Session and init variables
init = tf.global_variables_initializer()
sess.run(init)
for epoch in range(training_epochs): # Loop thru dataset multiple times
for (x, y) in zip(x_train, y_train): # Loop thru each point in the dataset
sess.run(train_op, feed_dict={X: x, Y: y})
w_val = sess.run(w) # Get final parameter value
sess.close() # Close the session
plt.scatter(x_train, y_train) # Plot the original data
y_learned = x_train*w_val
plt.plot(x_train, y_learned, 'r') # Plot the best-fit line
plt.show()
```
## Polynomial Regression
When a simple linear function won't fit the data, a polynomial function offers more flexibility. An Nth degree polynomial: $f(x) = w_nx^n + ... + w_1x + w_0$ can also describe a linear function when $n=1$
```
%matplotlib inline
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
learning_rate = 0.01 # Hyperparameters
training_epochs = 40
trX = np.linspace(-1, 1, 101) # Dataset based on 5th deg polynomial
num_coeffs = 6
trY_coeffs = [1, 2, 3, 4, 5, 6]
trY = 0
for i in range(num_coeffs):
trY += trY_coeffs[i] * np.power(trX, i)
trY += np.random.randn(*trX.shape) * 1.5 # Add noise
plt.scatter(trX, trY)
plt.show()
X = tf.placeholder(tf.float32) # tf placeholder nodes for input/output
Y = tf.placeholder(tf.float32)
w = tf.Variable(0.0, name="weights") # Weights variable
def model(X, w): # defines model as 5th deg poly
terms = []
for i in range(num_coeffs):
term = tf.multiply(w[i], tf.pow(X, i))
terms.append(term)
return tf.add_n(terms)
w = tf.Variable([0.] * num_coeffs, name="parameters") # Sets param vector to zeros
y_model = model(X, w)
cost = (tf.pow(Y-y_model, 2)) # Cost Function
# Defines the operation to be called on each iteration of the learning algorithm
train_op = tf.train.GradientDescentOptimizer(learning_rate).minimize(cost)
sess = tf.Session() # Setup the tf Session and init variables
init = tf.global_variables_initializer()
sess.run(init)
for epoch in range(training_epochs): # Loop thru dataset multiple times
for (x, y) in zip(trX, trY): # Loop thru each point in the dataset
sess.run(train_op, feed_dict={X: x, Y: y})
w_val = sess.run(w) # Get final parameter value
print("5th deg polynomial coeffs:\n", w_val)
sess.close() # Close the session
plt.scatter(trX, trY) # Plot the original data
trY2 = 0
for i in range(num_coeffs): # Plot the result
trY2 += w_val[i] * np.power(trX, i)
plt.plot(trX, trY2, 'r')
plt.show()
```
## Regularization
Polynomial regression isn't always the best ... the goal should be to find the simplest function that best represents the data. A 10th degree polynomial isn't always the best answer. **Regularization** is a way of penalizing the wrong parameters in order to have the correct parameters be more dominant.
When the regularization parameter, $\lambda$, is zero there is no regularization taking place. Larger values of $\lambda$ penalize parameters with larger norms more; making an overly complicated model less complex and reducing its flexibility (fixes overfitting).
| github_jupyter |
*Note: This is not yet ready, but shows the direction I'm leaning in for Fourth Edition Search.*
# State-Space Search
This notebook describes several state-space search algorithms, and how they can be used to solve a variety of problems. We start with a simple algorithm and a simple domain: finding a route from city to city. Later we will explore other algorithms and domains.
## The Route-Finding Domain
Like all state-space search problems, in a route-finding problem you will be given:
- A start state (for example, `'A'` for the city Arad).
- A goal state (for example, `'B'` for the city Bucharest).
- Actions that can change state (for example, driving from `'A'` to `'S'`).
You will be asked to find:
- A path from the start state, through intermediate states, to the goal state.
We'll use this map:
<img src="http://robotics.cs.tamu.edu/dshell/cs625/images/map.jpg" height="366" width="603">
A state-space search problem can be represented by a *graph*, where the vertexes of the graph are the states of the problem (in this case, cities) and the edges of the graph are the actions (in this case, driving along a road).
We'll represent a city by its single initial letter.
We'll represent the graph of connections as a `dict` that maps each city to a list of the neighboring cities (connected by a road). For now we don't explicitly represent the actions, nor the distances
between cities.
```
romania = {
'A': ['Z', 'T', 'S'],
'B': ['F', 'P', 'G', 'U'],
'C': ['D', 'R', 'P'],
'D': ['M', 'C'],
'E': ['H'],
'F': ['S', 'B'],
'G': ['B'],
'H': ['U', 'E'],
'I': ['N', 'V'],
'L': ['T', 'M'],
'M': ['L', 'D'],
'N': ['I'],
'O': ['Z', 'S'],
'P': ['R', 'C', 'B'],
'R': ['S', 'C', 'P'],
'S': ['A', 'O', 'F', 'R'],
'T': ['A', 'L'],
'U': ['B', 'V', 'H'],
'V': ['U', 'I'],
'Z': ['O', 'A']}
```
Suppose we want to get from `A` to `B`. Where can we go from the start state, `A`?
```
romania['A']
```
We see that from `A` we can get to any of the three cities `['Z', 'T', 'S']`. Which should we choose? *We don't know.* That's the whole point of *search*: we don't know which immediate action is best, so we'll have to explore, until we find a *path* that leads to the goal.
How do we explore? We'll start with a simple algorithm that will get us from `A` to `B`. We'll keep a *frontier*—a collection of not-yet-explored states—and expand the frontier outward until it reaches the goal. To be more precise:
- Initially, the only state in the frontier is the start state, `'A'`.
- Until we reach the goal, or run out of states in the frontier to explore, do the following:
- Remove the first state from the frontier. Call it `s`.
- If `s` is the goal, we're done. Return the path to `s`.
- Otherwise, consider all the neighboring states of `s`. For each one:
- If we have not previously explored the state, add it to the end of the frontier.
- Also keep track of the previous state that led to this new neighboring state; we'll need this to reconstruct the path to the goal, and to keep us from re-visiting previously explored states.
# A Simple Search Algorithm: `breadth_first`
The function `breadth_first` implements this strategy:
```
from collections import deque # Doubly-ended queue: pop from left, append to right.
def breadth_first(start, goal, neighbors):
"Find a shortest sequence of states from start to the goal."
frontier = deque([start]) # A queue of states
previous = {start: None} # start has no previous state; other states will
while frontier:
s = frontier.popleft()
if s == goal:
return path(previous, s)
for s2 in neighbors[s]:
if s2 not in previous:
frontier.append(s2)
previous[s2] = s
def path(previous, s):
"Return a list of states that lead to state s, according to the previous dict."
return [] if (s is None) else path(previous, previous[s]) + [s]
```
A couple of things to note:
1. We always add new states to the end of the frontier queue. That means that all the states that are adjacent to the start state will come first in the queue, then all the states that are two steps away, then three steps, etc.
That's what we mean by *breadth-first* search.
2. We recover the path to an `end` state by following the trail of `previous[end]` pointers, all the way back to `start`.
The dict `previous` is a map of `{state: previous_state}`.
3. When we finally get an `s` that is the goal state, we know we have found a shortest path, because any other state in the queue must correspond to a path that is as long or longer.
3. Note that `previous` contains all the states that are currently in `frontier` as well as all the states that were in `frontier` in the past.
4. If no path to the goal is found, then `breadth_first` returns `None`. If a path is found, it returns the sequence of states on the path.
Some examples:
```
breadth_first('A', 'B', romania)
breadth_first('L', 'N', romania)
breadth_first('N', 'L', romania)
breadth_first('E', 'E', romania)
```
Now let's try a different kind of problem that can be solved with the same search function.
## Word Ladders Problem
A *word ladder* problem is this: given a start word and a goal word, find the shortest way to transform the start word into the goal word by changing one letter at a time, such that each change results in a word. For example starting with `green` we can reach `grass` in 7 steps:
`green` → `greed` → `treed` → `trees` → `tress` → `cress` → `crass` → `grass`
We will need a dictionary of words. We'll use 5-letter words from the [Stanford GraphBase](http://www-cs-faculty.stanford.edu/~uno/sgb.html) project for this purpose. Let's get that file from aimadata.
```
from search import *
sgb_words = DataFile("EN-text/sgb-words.txt")
```
We can assign `WORDS` to be the set of all the words in this file:
```
WORDS = set(sgb_words.read().split())
len(WORDS)
```
And define `neighboring_words` to return the set of all words that are a one-letter change away from a given `word`:
```
def neighboring_words(word):
"All words that are one letter away from this word."
neighbors = {word[:i] + c + word[i+1:]
for i in range(len(word))
for c in 'abcdefghijklmnopqrstuvwxyz'
if c != word[i]}
return neighbors & WORDS
```
For example:
```
neighboring_words('hello')
neighboring_words('world')
```
Now we can create `word_neighbors` as a dict of `{word: {neighboring_word, ...}}`:
```
word_neighbors = {word: neighboring_words(word)
for word in WORDS}
```
Now the `breadth_first` function can be used to solve a word ladder problem:
```
breadth_first('green', 'grass', word_neighbors)
breadth_first('smart', 'brain', word_neighbors)
breadth_first('frown', 'smile', word_neighbors)
```
# More General Search Algorithms
Now we'll embelish the `breadth_first` algorithm to make a family of search algorithms with more capabilities:
1. We distinguish between an *action* and the *result* of an action.
3. We allow different measures of the cost of a solution (not just the number of steps in the sequence).
4. We search through the state space in an order that is more likely to lead to an optimal solution quickly.
Here's how we do these things:
1. Instead of having a graph of neighboring states, we instead have an object of type *Problem*. A Problem
has one method, `Problem.actions(state)` to return a collection of the actions that are allowed in a state,
and another method, `Problem.result(state, action)` that says what happens when you take an action.
2. We keep a set, `explored` of states that have already been explored. We also have a class, `Frontier`, that makes it efficient to ask if a state is on the frontier.
3. Each action has a cost associated with it (in fact, the cost can vary with both the state and the action).
4. The `Frontier` class acts as a priority queue, allowing the "best" state to be explored next.
We represent a sequence of actions and resulting states as a linked list of `Node` objects.
The algorithm `breadth_first_search` is basically the same as `breadth_first`, but using our new conventions:
```
def breadth_first_search(problem):
"Search for goal; paths with least number of steps first."
if problem.is_goal(problem.initial):
return Node(problem.initial)
frontier = FrontierQ(Node(problem.initial), LIFO=False)
explored = set()
while frontier:
node = frontier.pop()
explored.add(node.state)
for action in problem.actions(node.state):
child = node.child(problem, action)
if child.state not in explored and child.state not in frontier:
if problem.is_goal(child.state):
return child
frontier.add(child)
```
Next is `uniform_cost_search`, in which each step can have a different cost, and we still consider first one os the states with minimum cost so far.
```
def uniform_cost_search(problem, costfn=lambda node: node.path_cost):
frontier = FrontierPQ(Node(problem.initial), costfn)
explored = set()
while frontier:
node = frontier.pop()
if problem.is_goal(node.state):
return node
explored.add(node.state)
for action in problem.actions(node.state):
child = node.child(problem, action)
if child.state not in explored and child not in frontier:
frontier.add(child)
elif child in frontier and frontier.cost[child] < child.path_cost:
frontier.replace(child)
```
Finally, `astar_search` in which the cost includes an estimate of the distance to the goal as well as the distance travelled so far.
```
def astar_search(problem, heuristic):
costfn = lambda node: node.path_cost + heuristic(node.state)
return uniform_cost_search(problem, costfn)
```
# Search Tree Nodes
The solution to a search problem is now a linked list of `Node`s, where each `Node`
includes a `state` and the `path_cost` of getting to the state. In addition, for every `Node` except for the first (root) `Node`, there is a previous `Node` (indicating the state that lead to this `Node`) and an `action` (indicating the action taken to get here).
```
class Node(object):
"""A node in a search tree. A search tree is spanning tree over states.
A Node contains a state, the previous node in the tree, the action that
takes us from the previous state to this state, and the path cost to get to
this state. If a state is arrived at by two paths, then there are two nodes
with the same state."""
def __init__(self, state, previous=None, action=None, step_cost=1):
"Create a search tree Node, derived from a previous Node by an action."
self.state = state
self.previous = previous
self.action = action
self.path_cost = 0 if previous is None else (previous.path_cost + step_cost)
def __repr__(self): return "<Node {}: {}>".format(self.state, self.path_cost)
def __lt__(self, other): return self.path_cost < other.path_cost
def child(self, problem, action):
"The Node you get by taking an action from this Node."
result = problem.result(self.state, action)
return Node(result, self, action,
problem.step_cost(self.state, action, result))
```
# Frontiers
A frontier is a collection of Nodes that acts like both a Queue and a Set. A frontier, `f`, supports these operations:
* `f.add(node)`: Add a node to the Frontier.
* `f.pop()`: Remove and return the "best" node from the frontier.
* `f.replace(node)`: add this node and remove a previous node with the same state.
* `state in f`: Test if some node in the frontier has arrived at state.
* `f[state]`: returns the node corresponding to this state in frontier.
* `len(f)`: The number of Nodes in the frontier. When the frontier is empty, `f` is *false*.
We provide two kinds of frontiers: One for "regular" queues, either first-in-first-out (for breadth-first search) or last-in-first-out (for depth-first search), and one for priority queues, where you can specify what cost function on nodes you are trying to minimize.
```
from collections import OrderedDict
import heapq
class FrontierQ(OrderedDict):
"A Frontier that supports FIFO or LIFO Queue ordering."
def __init__(self, initial, LIFO=False):
"""Initialize Frontier with an initial Node.
If LIFO is True, pop from the end first; otherwise from front first."""
self.LIFO = LIFO
self.add(initial)
def add(self, node):
"Add a node to the frontier."
self[node.state] = node
def pop(self):
"Remove and return the next Node in the frontier."
(state, node) = self.popitem(self.LIFO)
return node
def replace(self, node):
"Make this node replace the nold node with the same state."
del self[node.state]
self.add(node)
class FrontierPQ:
"A Frontier ordered by a cost function; a Priority Queue."
def __init__(self, initial, costfn=lambda node: node.path_cost):
"Initialize Frontier with an initial Node, and specify a cost function."
self.heap = []
self.states = {}
self.costfn = costfn
self.add(initial)
def add(self, node):
"Add node to the frontier."
cost = self.costfn(node)
heapq.heappush(self.heap, (cost, node))
self.states[node.state] = node
def pop(self):
"Remove and return the Node with minimum cost."
(cost, node) = heapq.heappop(self.heap)
self.states.pop(node.state, None) # remove state
return node
def replace(self, node):
"Make this node replace a previous node with the same state."
if node.state not in self:
raise ValueError('{} not there to replace'.format(node.state))
for (i, (cost, old_node)) in enumerate(self.heap):
if old_node.state == node.state:
self.heap[i] = (self.costfn(node), node)
heapq._siftdown(self.heap, 0, i)
return
def __contains__(self, state): return state in self.states
def __len__(self): return len(self.heap)
```
# Search Problems
`Problem` is the abstract class for all search problems. You can define your own class of problems as a subclass of `Problem`. You will need to override the `actions` and `result` method to describe how your problem works. You will also have to either override `is_goal` or pass a collection of goal states to the initialization method. If actions have different costs, you should override the `step_cost` method.
```
class Problem(object):
"""The abstract class for a search problem."""
def __init__(self, initial=None, goals=(), **additional_keywords):
"""Provide an initial state and optional goal states.
A subclass can have additional keyword arguments."""
self.initial = initial # The initial state of the problem.
self.goals = goals # A collection of possibe goal states.
self.__dict__.update(**additional_keywords)
def actions(self, state):
"Return a list of actions executable in this state."
raise NotImplementedError # Override this!
def result(self, state, action):
"The state that results from executing this action in this state."
raise NotImplementedError # Override this!
def is_goal(self, state):
"True if the state is a goal."
return state in self.goals # Optionally override this!
def step_cost(self, state, action, result=None):
"The cost of taking this action from this state."
return 1 # Override this if actions have different costs
def action_sequence(node):
"The sequence of actions to get to this node."
actions = []
while node.previous:
actions.append(node.action)
node = node.previous
return actions[::-1]
def state_sequence(node):
"The sequence of states to get to this node."
states = [node.state]
while node.previous:
node = node.previous
states.append(node.state)
return states[::-1]
```
# Two Location Vacuum World
```
dirt = '*'
clean = ' '
class TwoLocationVacuumProblem(Problem):
"""A Vacuum in a world with two locations, and dirt.
Each state is a tuple of (location, dirt_in_W, dirt_in_E)."""
def actions(self, state): return ('W', 'E', 'Suck')
def is_goal(self, state): return dirt not in state
def result(self, state, action):
"The state that results from executing this action in this state."
(loc, dirtW, dirtE) = state
if action == 'W': return ('W', dirtW, dirtE)
elif action == 'E': return ('E', dirtW, dirtE)
elif action == 'Suck' and loc == 'W': return (loc, clean, dirtE)
elif action == 'Suck' and loc == 'E': return (loc, dirtW, clean)
else: raise ValueError('unknown action: ' + action)
problem = TwoLocationVacuumProblem(initial=('W', dirt, dirt))
result = uniform_cost_search(problem)
result
action_sequence(result)
state_sequence(result)
problem = TwoLocationVacuumProblem(initial=('E', clean, dirt))
result = uniform_cost_search(problem)
action_sequence(result)
```
# Water Pouring Problem
Here is another problem domain, to show you how to define one. The idea is that we have a number of water jugs and a water tap and the goal is to measure out a specific amount of water (in, say, ounces or liters). You can completely fill or empty a jug, but because the jugs don't have markings on them, you can't partially fill them with a specific amount. You can, however, pour one jug into another, stopping when the seconfd is full or the first is empty.
```
class PourProblem(Problem):
"""Problem about pouring water between jugs to achieve some water level.
Each state is a tuples of levels. In the initialization, provide a tuple of
capacities, e.g. PourProblem(capacities=(8, 16, 32), initial=(2, 4, 3), goals={7}),
which means three jugs of capacity 8, 16, 32, currently filled with 2, 4, 3 units of
water, respectively, and the goal is to get a level of 7 in any one of the jugs."""
def actions(self, state):
"""The actions executable in this state."""
jugs = range(len(state))
return ([('Fill', i) for i in jugs if state[i] != self.capacities[i]] +
[('Dump', i) for i in jugs if state[i] != 0] +
[('Pour', i, j) for i in jugs for j in jugs if i != j])
def result(self, state, action):
"""The state that results from executing this action in this state."""
result = list(state)
act, i, j = action[0], action[1], action[-1]
if act == 'Fill': # Fill i to capacity
result[i] = self.capacities[i]
elif act == 'Dump': # Empty i
result[i] = 0
elif act == 'Pour':
a, b = state[i], state[j]
result[i], result[j] = ((0, a + b)
if (a + b <= self.capacities[j]) else
(a + b - self.capacities[j], self.capacities[j]))
else:
raise ValueError('unknown action', action)
return tuple(result)
def is_goal(self, state):
"""True if any of the jugs has a level equal to one of the goal levels."""
return any(level in self.goals for level in state)
p7 = PourProblem(initial=(2, 0), capacities=(5, 13), goals={7})
p7.result((2, 0), ('Fill', 1))
result = uniform_cost_search(p7)
action_sequence(result)
```
# Visualization Output
```
def showpath(searcher, problem):
"Show what happens when searcvher solves problem."
problem = Instrumented(problem)
print('\n{}:'.format(searcher.__name__))
result = searcher(problem)
if result:
actions = action_sequence(result)
state = problem.initial
path_cost = 0
for steps, action in enumerate(actions, 1):
path_cost += problem.step_cost(state, action, 0)
result = problem.result(state, action)
print(' {} =={}==> {}; cost {} after {} steps'
.format(state, action, result, path_cost, steps,
'; GOAL!' if problem.is_goal(result) else ''))
state = result
msg = 'GOAL FOUND' if result else 'no solution'
print('{} after {} results and {} goal checks'
.format(msg, problem._counter['result'], problem._counter['is_goal']))
from collections import Counter
class Instrumented:
"Instrument an object to count all the attribute accesses in _counter."
def __init__(self, obj):
self._object = obj
self._counter = Counter()
def __getattr__(self, attr):
self._counter[attr] += 1
return getattr(self._object, attr)
showpath(uniform_cost_search, p7)
p = PourProblem(initial=(0, 0), capacities=(7, 13), goals={2})
showpath(uniform_cost_search, p)
class GreenPourProblem(PourProblem):
def step_cost(self, state, action, result=None):
"The cost is the amount of water used in a fill."
if action[0] == 'Fill':
i = action[1]
return self.capacities[i] - state[i]
return 0
p = GreenPourProblem(initial=(0, 0), capacities=(7, 13), goals={2})
showpath(uniform_cost_search, p)
def compare_searchers(problem, searchers=None):
"Apply each of the search algorithms to the problem, and show results"
if searchers is None:
searchers = (breadth_first_search, uniform_cost_search)
for searcher in searchers:
showpath(searcher, problem)
compare_searchers(p)
```
# Random Grid
An environment where you can move in any of 4 directions, unless there is an obstacle there.
```
import random
N, S, E, W = DIRECTIONS = [(0, 1), (0, -1), (1, 0), (-1, 0)]
def Grid(width, height, obstacles=0.1):
"""A 2-D grid, width x height, with obstacles that are either a collection of points,
or a fraction between 0 and 1 indicating the density of obstacles, chosen at random."""
grid = {(x, y) for x in range(width) for y in range(height)}
if isinstance(obstacles, (float, int)):
obstacles = random.sample(grid, int(width * height * obstacles))
def neighbors(x, y):
for (dx, dy) in DIRECTIONS:
(nx, ny) = (x + dx, y + dy)
if (nx, ny) not in obstacles and 0 <= nx < width and 0 <= ny < height:
yield (nx, ny)
return {(x, y): list(neighbors(x, y))
for x in range(width) for y in range(height)}
Grid(5, 5)
class GridProblem(Problem):
"Create with a call like GridProblem(grid=Grid(10, 10), initial=(0, 0), goal=(9, 9))"
def actions(self, state): return DIRECTIONS
def result(self, state, action):
#print('ask for result of', state, action)
(x, y) = state
(dx, dy) = action
r = (x + dx, y + dy)
return r if r in self.grid[state] else state
gp = GridProblem(grid=Grid(5, 5, 0.3), initial=(0, 0), goals={(4, 4)})
showpath(uniform_cost_search, gp)
```
# Finding a hard PourProblem
What solvable two-jug PourProblem requires the most steps? We can define the hardness as the number of steps, and then iterate over all PourProblems with capacities up to size M, keeping the hardest one.
```
def hardness(problem):
L = breadth_first_search(problem)
#print('hardness', problem.initial, problem.capacities, problem.goals, L)
return len(action_sequence(L)) if (L is not None) else 0
hardness(p7)
action_sequence(breadth_first_search(p7))
C = 9 # Maximum capacity to consider
phard = max((PourProblem(initial=(a, b), capacities=(A, B), goals={goal})
for A in range(C+1) for B in range(C+1)
for a in range(A) for b in range(B)
for goal in range(max(A, B))),
key=hardness)
phard.initial, phard.capacities, phard.goals
showpath(breadth_first_search, PourProblem(initial=(0, 0), capacities=(7, 9), goals={8}))
showpath(uniform_cost_search, phard)
class GridProblem(Problem):
"""A Grid."""
def actions(self, state): return ['N', 'S', 'E', 'W']
def result(self, state, action):
"""The state that results from executing this action in this state."""
(W, H) = self.size
if action == 'N' and state > W: return state - W
if action == 'S' and state + W < W * W: return state + W
if action == 'E' and (state + 1) % W !=0: return state + 1
if action == 'W' and state % W != 0: return state - 1
return state
compare_searchers(GridProblem(initial=0, goals={44}, size=(10, 10)))
def test_frontier():
#### Breadth-first search with FIFO Q
f = FrontierQ(Node(1), LIFO=False)
assert 1 in f and len(f) == 1
f.add(Node(2))
f.add(Node(3))
assert 1 in f and 2 in f and 3 in f and len(f) == 3
assert f.pop().state == 1
assert 1 not in f and 2 in f and 3 in f and len(f) == 2
assert f
assert f.pop().state == 2
assert f.pop().state == 3
assert not f
#### Depth-first search with LIFO Q
f = FrontierQ(Node('a'), LIFO=True)
for s in 'bcdef': f.add(Node(s))
assert len(f) == 6 and 'a' in f and 'c' in f and 'f' in f
for s in 'fedcba': assert f.pop().state == s
assert not f
#### Best-first search with Priority Q
f = FrontierPQ(Node(''), lambda node: len(node.state))
assert '' in f and len(f) == 1 and f
for s in ['book', 'boo', 'bookie', 'bookies', 'cook', 'look', 'b']:
assert s not in f
f.add(Node(s))
assert s in f
assert f.pop().state == ''
assert f.pop().state == 'b'
assert f.pop().state == 'boo'
assert {f.pop().state for _ in '123'} == {'book', 'cook', 'look'}
assert f.pop().state == 'bookie'
#### Romania: Two paths to Bucharest; cheapest one found first
S = Node('S')
SF = Node('F', S, 'S->F', 99)
SFB = Node('B', SF, 'F->B', 211)
SR = Node('R', S, 'S->R', 80)
SRP = Node('P', SR, 'R->P', 97)
SRPB = Node('B', SRP, 'P->B', 101)
f = FrontierPQ(S)
f.add(SF); f.add(SR), f.add(SRP), f.add(SRPB); f.add(SFB)
def cs(n): return (n.path_cost, n.state) # cs: cost and state
assert cs(f.pop()) == (0, 'S')
assert cs(f.pop()) == (80, 'R')
assert cs(f.pop()) == (99, 'F')
assert cs(f.pop()) == (177, 'P')
assert cs(f.pop()) == (278, 'B')
return 'test_frontier ok'
test_frontier()
%matplotlib inline
import matplotlib.pyplot as plt
p = plt.plot([i**2 for i in range(10)])
plt.savefig('destination_path.eps', format='eps', dpi=1200)
import itertools
import random
# http://stackoverflow.com/questions/10194482/custom-matplotlib-plot-chess-board-like-table-with-colored-cells
from matplotlib.table import Table
def main():
grid_table(8, 8)
plt.axis('scaled')
plt.show()
def grid_table(nrows, ncols):
fig, ax = plt.subplots()
ax.set_axis_off()
colors = ['white', 'lightgrey', 'dimgrey']
tb = Table(ax, bbox=[0,0,2,2])
for i,j in itertools.product(range(ncols), range(nrows)):
tb.add_cell(i, j, 2./ncols, 2./nrows, text='{:0.2f}'.format(0.1234),
loc='center', facecolor=random.choice(colors), edgecolor='grey') # facecolors=
ax.add_table(tb)
#ax.plot([0, .3], [.2, .2])
#ax.add_line(plt.Line2D([0.3, 0.5], [0.7, 0.7], linewidth=2, color='blue'))
return fig
main()
import collections
class defaultkeydict(collections.defaultdict):
"""Like defaultdict, but the default_factory is a function of the key.
>>> d = defaultkeydict(abs); d[-42]
42
"""
def __missing__(self, key):
self[key] = self.default_factory(key)
return self[key]
```
| github_jupyter |
<img src="../../../images/qiskit-heading.png" alt="Note: In order for images to show up in this jupyter notebook you need to select File => Trusted Notebook" width="500 px" align="left">
# Wigner Functions
---
### Contributers
Russell P. Rundle$^{1,2}$, Todd Tilma$^{1,3}$, Vincent M. Dwyer$^{1,2}$, Mark J. Everitt (m.j.everitt@physics.org)$^{1}$
1 Quantum Systems Engineering Research Group, Physics Dpartment, Loughborough University, UK
2 Wolfson School, Loughborough University, UK
3 Tokyo Institute of Technology, Japan
## Introduction
In this notebook we demonstrate how to create Wigner functions by either using the full state or by measuring points in phase space. We will show the different methods which can be used by inputting an arbitrary state or just measuring the state on both the simulators and the IBM Quantum Experience.
The spin Wigner function presented here is based on work from [*T Tilma, MJ Everitt, JH Samson, WJ Munro, K Nemoto, Phys. Rev. Lett. 117, 180401*](https://journals.aps.org/prl/abstract/10.1103/PhysRevLett.117.180401) and can be calculated analytically as
$W(\boldsymbol{\Omega}) = \mathrm{Tr}\left[\hat{\rho} \; \hat{U}(\boldsymbol{\Omega}) \hat{\Pi} \hat{U}^\dagger(\boldsymbol{\Omega})\right]$
Where $\hat{U}$ is the rotation by the Euler angles and $\hat{\Pi}$ is the parity, such that
$\hat{\Pi} = \frac{1}{2}
\begin{pmatrix}
1+\sqrt{3} & 0\\
0 & 1-\sqrt{3}
\end{pmatrix}$
for one qubit. The Hilbert space for a single qubit is spanned by the Euler angles where $\theta$ (the elevation) and $\phi$ (the azimuth) thus the full Wigner function for $n$ qubits is given by $2n$ degrees of freedom.
<img src="../images/blochSphere.png", width = 300, height = 300>
When there is more than one qubit, we take the Kronecker tensor product of the $\hat{U}$s and $\hat{\Pi}$. In the following examples, various methods of extracting information from 2 entangled qubits from four degrees of freedom are shown.
These methods include taking an the equal angle slice, where we set the angles $\theta_0 = \theta_1 = \theta$ and $\phi_0 = \phi_1 = \phi$. We can also set some angles to be constant values as we plot the remaining angles against each other, an example of this can be seen when plotting the plaquettes where we set $\phi_0=\phi_1=0$ and we plot $\theta_0$ against $\theta_1$. Finally we show how we can simplify the Wigner function by just taking a 2 dimensional curve around the function, this is shown when we look at the equal angle equatorial slice where we take $\theta_i=\theta=\pi/2$ for all $\theta$s and $\phi_i=\phi$ for all values of $\phi$ where $0\leq\phi<2\pi$.
With this visualisation methods we show how these points in phase space can be measured directly by using the Wigner function tomography module and it works by using the u3 gates to rotate to points in phase space. Below is an example for measing points for a Bell-state, the first two gates create the Bell-state and the two u3 are example rotations to a point in phase space.
<img src="../images/exampleCircuit.png", width = 400>
# Wigner Functions for Arbitrary States
First we will look at the Wigner function tomography module and show how we can create the spin Wigner function on a Bloch sphere for an arbitrary state. All that is necessary is to input a density matrix or state vector for a known state and then set the desired resolution for the plot (if no resolution is set, the mesh of the sphere will default to 100x100)
```
# importing the QISKit
from qiskit import QuantumCircuit, QuantumProgram
import Qconfig
# import tomography library
import qiskit.tools.qcvv.tomography as tomo
#visualization packages
from qiskit.tools.visualization import plot_wigner_function, plot_wigner_data
```
## Wigner Function for an entangled Bell-state
We begin by creating the density matrix for a Bell-state (note that the matrices and state vectors need to be created in numpy matrix format), $\left(|00\rangle+|11\rangle\right)/\sqrt{2}$, which is given by
$
\frac{1}{2}
\begin{pmatrix}
1 & 0 & 0 & 1 \\
0 & 0 & 0 & 0 \\
0 & 0 & 0 & 0 \\
1 & 0 & 0 & 1
\end{pmatrix}
$
```
density_matrix = np.matrix([[0.5, 0, 0, 0.5],[0, 0, 0, 0],[0, 0, 0, 0],[0.5, 0, 0, 0.5]])
print(density_matrix)
```
This density matrix can then be put into wigner_function which plots the Bloch sphere Wigner function for the equal angle slice, i.e. where $\theta_0 = \theta_1 = \theta$ and $\phi_0 = \phi_1 = \phi$
```
plot_wigner_function(density_matrix, res=200)
```
If it is desired to calculate the analytic expression for the Wigner function, we can import the sympy package and run the Wigner function calculations symbolically. The below code is almost identical to the code in the Wigner function tomography module but edited to work with symbolics rather than run through the points in the mesh of the sphere.
```
import sympy as sym
from sympy.physics.quantum import TensorProduct
num = int(np.log2(len(density_matrix)))
harr = sym.sqrt(3)
Delta_su2 = sym.zeros(2)
Delta = sym.ones(1)
for qubit in range(num):
phi = sym.Indexed('phi', qubit)
theta = sym.Indexed('theta', qubit)
costheta = harr*sym.cos(2*theta)
sintheta = harr*sym.sin(2*theta)
Delta_su2[0,0] = (1+costheta)/2
Delta_su2[0,1] = -(sym.exp(2j*phi)*sintheta)/2
Delta_su2[1,0] = -(sym.exp(-2j*phi)*sintheta)/2
Delta_su2[1,1] = (1-costheta)/2
Delta = TensorProduct(Delta,Delta_su2)
W = sym.trace(density_matrix*Delta)
print(sym.latex(W))
```
pasting this result into a latex environment returns the Wigner function
$W = 0.5 \left(- \frac{\sqrt{3}}{2} \cos{\left (2 \theta_{0} \right )} + \frac{1}{2}\right) \left(- \frac{\sqrt{3}}{2} \cos{\left (2 \theta_{1} \right )} + \frac{1}{2}\right) + 0.5 \left(\frac{\sqrt{3}}{2} \cos{\left (2 \theta_{0} \right )} + \frac{1}{2}\right) \left(\frac{\sqrt{3}}{2} \cos{\left (2 \theta_{1} \right )} + \frac{1}{2}\right) + 0.375 e^{2.0 i \phi_{0}} e^{2.0 i \phi_{1}} \sin{\left (2 \theta_{0} \right )} \sin{\left (2 \theta_{1} \right )} + 0.375 e^{- 2.0 i \phi_{0}} e^{- 2.0 i \phi_{1}} \sin{\left (2 \theta_{0} \right )} \sin{\left (2 \theta_{1} \right )}$
### Wigner function using State tomography
We can create the density matrix for a given state by using the state tomography module, more information for the tomography module can be found in the [quantum state tomography tutorial](https://github.com/QISKit/qiskit-tutorial/blob/master/3_qcvv/state_tomography.ipynb). First creating an entangled Bell-state with the following set of gates and then taking the state tomography, we can create the spin Wigner function for the measured tomography
```
Q_program = QuantumProgram()
number_of_qubits = 2
backend = 'local_qasm_simulator'
shots = 1024
bell_qubits = [0, 1]
qr = Q_program.create_quantum_register('qr',2)
cr = Q_program.create_classical_register('cr',2)
bell = Q_program.create_circuit('bell', [qr], [cr])
bell.h(qr[0])
bell.cx(qr[0],qr[1])
```
The above code generates the circuits needed to create the Bell-state and below we show how to then build the density matrix for the state using the tomography module. The density matrix created can then be put into the Wigner fucntion visualisation module
```
bell_tomo_set = tomo.state_tomography_set([0, 1])
bell_tomo_circuits = tomo.create_tomography_circuits(Q_program, 'bell', qr, cr, bell_tomo_set)
bell_tomo_result = Q_program.execute(bell_tomo_circuits, backend=backend, shots=shots)
bell_tomo_data = tomo.tomography_data(bell_tomo_result, 'bell', bell_tomo_set)
rho_fit_sim = tomo.fit_tomography_data(bell_tomo_data)
plot_wigner_function(np.matrix(rho_fit_sim),res=200)
```
Again, using the same code from before, we can calculate the analytic expression for the Wigner function from the generated density matrix, it can be seen that since the density matrix is less sparse, the resulting Wigner function has many more terms
```
Delta_su2 = sym.zeros(2)
Delta = sym.ones(1)
for qubit in range(num):
phi = sym.Indexed('phi', qubit)
theta = sym.Indexed('theta', qubit)
costheta = harr*sym.cos(2*theta)
sintheta = harr*sym.sin(2*theta)
Delta_su2[0,0] = (1+costheta)/2
Delta_su2[0,1] = -(sym.exp(2j*phi)*sintheta)/2
Delta_su2[1,0] = -(sym.exp(-2j*phi)*sintheta)/2
Delta_su2[1,1] = (1-costheta)/2
Delta = TensorProduct(Delta,Delta_su2)
W = sym.trace(np.matrix(rho_fit_sim)*Delta)
print(sym.latex(W))
```
$W = 0.493698558640376 \left(- \frac{\sqrt{3}}{2} \cos{\left (2 \theta_{0} \right )} + \frac{1}{2}\right) \left(- \frac{\sqrt{3}}{2} \cos{\left (2 \theta_{1} \right )} + \frac{1}{2}\right) + 0.00275286172575906 \left(- \frac{\sqrt{3}}{2} \cos{\left (2 \theta_{0} \right )} + \frac{1}{2}\right) \left(\frac{\sqrt{3}}{2} \cos{\left (2 \theta_{1} \right )} + \frac{1}{2}\right) - \frac{\sqrt{3}}{2} \left(-0.00534735110770023 - 0.00770494561267218 i\right) \left(- \frac{\sqrt{3}}{2} \cos{\left (2 \theta_{0} \right )} + \frac{1}{2}\right) e^{2.0 i \phi_{1}} \sin{\left (2 \theta_{1} \right )} - \frac{\sqrt{3}}{2} \left(-0.00534735110770023 + 0.00770494561267218 i\right) \left(- \frac{\sqrt{3}}{2} \cos{\left (2 \theta_{0} \right )} + \frac{1}{2}\right) e^{- 2.0 i \phi_{1}} \sin{\left (2 \theta_{1} \right )} + 0.00266528247889166 \left(\frac{\sqrt{3}}{2} \cos{\left (2 \theta_{0} \right )} + \frac{1}{2}\right) \left(- \frac{\sqrt{3}}{2} \cos{\left (2 \theta_{1} \right )} + \frac{1}{2}\right) + 0.500883297154973 \left(\frac{\sqrt{3}}{2} \cos{\left (2 \theta_{0} \right )} + \frac{1}{2}\right) \left(\frac{\sqrt{3}}{2} \cos{\left (2 \theta_{1} \right )} + \frac{1}{2}\right) - \frac{\sqrt{3}}{2} \left(-0.0015149834996068 + 0.000102164609935808 i\right) \left(\frac{\sqrt{3}}{2} \cos{\left (2 \theta_{0} \right )} + \frac{1}{2}\right) e^{2.0 i \phi_{1}} \sin{\left (2 \theta_{1} \right )} - \frac{\sqrt{3}}{2} \left(-0.0015149834996068 - 0.000102164609935808 i\right) \left(\frac{\sqrt{3}}{2} \cos{\left (2 \theta_{0} \right )} + \frac{1}{2}\right) e^{- 2.0 i \phi_{1}} \sin{\left (2 \theta_{1} \right )} - \frac{\sqrt{3}}{2} \left(-0.00628509857352323 - 0.000924492486476053 i\right) \left(- \frac{\sqrt{3}}{2} \cos{\left (2 \theta_{1} \right )} + \frac{1}{2}\right) e^{2.0 i \phi_{0}} \sin{\left (2 \theta_{0} \right )} - \frac{\sqrt{3}}{2} \left(-0.00628509857352323 + 0.000924492486476053 i\right) \left(- \frac{\sqrt{3}}{2} \cos{\left (2 \theta_{1} \right )} + \frac{1}{2}\right) e^{- 2.0 i \phi_{0}} \sin{\left (2 \theta_{0} \right )} - \frac{\sqrt{3}}{2} \left(-0.000462681165102337 + 0.00867331154973143 i\right) \left(\frac{\sqrt{3}}{2} \cos{\left (2 \theta_{1} \right )} + \frac{1}{2}\right) e^{2.0 i \phi_{0}} \sin{\left (2 \theta_{0} \right )} - \frac{\sqrt{3}}{2} \left(-0.000462681165102337 - 0.00867331154973143 i\right) \left(\frac{\sqrt{3}}{2} \cos{\left (2 \theta_{1} \right )} + \frac{1}{2}\right) e^{- 2.0 i \phi_{0}} \sin{\left (2 \theta_{0} \right )} + \frac{3}{4} \left(0.492697839219187 - 0.0085595434721166 i\right) e^{2.0 i \phi_{0}} e^{2.0 i \phi_{1}} \sin{\left (2 \theta_{0} \right )} \sin{\left (2 \theta_{1} \right )} + \frac{3}{4} \left(0.00247818568149116 + 0.000867188901661959 i\right) e^{2.0 i \phi_{0}} e^{- 2.0 i \phi_{1}} \sin{\left (2 \theta_{0} \right )} \sin{\left (2 \theta_{1} \right )} + \frac{3}{4} \left(0.00247818568149116 - 0.000867188901661959 i\right) e^{- 2.0 i \phi_{0}} e^{2.0 i \phi_{1}} \sin{\left (2 \theta_{0} \right )} \sin{\left (2 \theta_{1} \right )} + \frac{3}{4} \left(0.492697839219187 + 0.0085595434721166 i\right) e^{- 2.0 i \phi_{0}} e^{- 2.0 i \phi_{1}} \sin{\left (2 \theta_{0} \right )} \sin{\left (2 \theta_{1} \right )}$
# Experiment
```
Q_program.set_api(Qconfig.APItoken, Qconfig.config['url'])
backend = 'ibmqx2'
max_credits = 8
shots = 1024
bell_qubits = [0, 1]
bell_tomo_set = tomo.state_tomography_set(bell_qubits)
bell_tomo_circuits = tomo.create_tomography_circuits(Q_program, 'bell', qr, cr, bell_tomo_set)
bell_tomo_result = Q_program.execute(bell_tomo_circuits, backend=backend, shots=shots,
max_credits=max_credits, timeout=300)
bell_tomo_data = tomo.tomography_data(bell_tomo_result, 'bell', bell_tomo_set)
rho_fit_ibmqx = tomo.fit_tomography_data(bell_tomo_data)
plot_wigner_function(np.matrix(rho_fit_ibmqx), res=100)
print(rho_fit_ibmqx)
Delta_su2 = sym.zeros(2)
Delta = sym.ones(1)
for qubit in range(num):
phi = sym.Indexed('phi', qubit)
theta = sym.Indexed('theta', qubit)
costheta = harr*sym.cos(2*theta)
sintheta = harr*sym.sin(2*theta)
Delta_su2[0,0] = (1+costheta)/2
Delta_su2[0,1] = -(sym.exp(2j*phi)*sintheta)/2
Delta_su2[1,0] = -(sym.exp(-2j*phi)*sintheta)/2
Delta_su2[1,1] = (1-costheta)/2
Delta = TensorProduct(Delta,Delta_su2)
W = sym.trace(np.matrix(rho_fit_ibmqx)*Delta)
print(sym.latex(W))
```
$0.32699519833677 \left(- \frac{\sqrt{3}}{2} \cos{\left (2 \theta_{0} \right )} + \frac{1}{2}\right) \left(- \frac{\sqrt{3}}{2} \cos{\left (2 \theta_{1} \right )} + \frac{1}{2}\right) + 0.124306773174647 \left(- \frac{\sqrt{3}}{2} \cos{\left (2 \theta_{0} \right )} + \frac{1}{2}\right) \left(\frac{\sqrt{3}}{2} \cos{\left (2 \theta_{1} \right )} + \frac{1}{2}\right) - \frac{\sqrt{3}}{2} \left(0.0285310304913017 + 0.0238741788035984 i\right) \left(- \frac{\sqrt{3}}{2} \cos{\left (2 \theta_{0} \right )} + \frac{1}{2}\right) e^{2.0 i \phi_{1}} \sin{\left (2 \theta_{1} \right )} - \frac{\sqrt{3}}{2} \left(0.0285310304913017 - 0.0238741788035984 i\right) \left(- \frac{\sqrt{3}}{2} \cos{\left (2 \theta_{0} \right )} + \frac{1}{2}\right) e^{- 2.0 i \phi_{1}} \sin{\left (2 \theta_{1} \right )} + 0.124202902962916 \left(\frac{\sqrt{3}}{2} \cos{\left (2 \theta_{0} \right )} + \frac{1}{2}\right) \left(- \frac{\sqrt{3}}{2} \cos{\left (2 \theta_{1} \right )} + \frac{1}{2}\right) + 0.424495125525667 \left(\frac{\sqrt{3}}{2} \cos{\left (2 \theta_{0} \right )} + \frac{1}{2}\right) \left(\frac{\sqrt{3}}{2} \cos{\left (2 \theta_{1} \right )} + \frac{1}{2}\right) - \frac{\sqrt{3}}{2} \left(0.0490359773360699 + 0.0381005337507329 i\right) \left(\frac{\sqrt{3}}{2} \cos{\left (2 \theta_{0} \right )} + \frac{1}{2}\right) e^{2.0 i \phi_{1}} \sin{\left (2 \theta_{1} \right )} - \frac{\sqrt{3}}{2} \left(0.0490359773360699 - 0.0381005337507329 i\right) \left(\frac{\sqrt{3}}{2} \cos{\left (2 \theta_{0} \right )} + \frac{1}{2}\right) e^{- 2.0 i \phi_{1}} \sin{\left (2 \theta_{1} \right )} - \frac{\sqrt{3}}{2} \left(0.0512856636758563 + 0.0414367583959388 i\right) \left(- \frac{\sqrt{3}}{2} \cos{\left (2 \theta_{1} \right )} + \frac{1}{2}\right) e^{2.0 i \phi_{0}} \sin{\left (2 \theta_{0} \right )} - \frac{\sqrt{3}}{2} \left(0.0512856636758563 - 0.0414367583959388 i\right) \left(- \frac{\sqrt{3}}{2} \cos{\left (2 \theta_{1} \right )} + \frac{1}{2}\right) e^{- 2.0 i \phi_{0}} \sin{\left (2 \theta_{0} \right )} - \frac{\sqrt{3}}{2} \left(0.0408691303527403 + 0.0280408874045363 i\right) \left(\frac{\sqrt{3}}{2} \cos{\left (2 \theta_{1} \right )} + \frac{1}{2}\right) e^{2.0 i \phi_{0}} \sin{\left (2 \theta_{0} \right )} - \frac{\sqrt{3}}{2} \left(0.0408691303527403 - 0.0280408874045363 i\right) \left(\frac{\sqrt{3}}{2} \cos{\left (2 \theta_{1} \right )} + \frac{1}{2}\right) e^{- 2.0 i \phi_{0}} \sin{\left (2 \theta_{0} \right )} + \frac{3}{4} \left(0.285095302985003 - 0.00922257353994807 i\right) e^{2.0 i \phi_{0}} e^{2.0 i \phi_{1}} \sin{\left (2 \theta_{0} \right )} \sin{\left (2 \theta_{1} \right )} + \frac{3}{4} \left(-0.035281529513114 - 0.00875077955598942 i\right) e^{2.0 i \phi_{0}} e^{- 2.0 i \phi_{1}} \sin{\left (2 \theta_{0} \right )} \sin{\left (2 \theta_{1} \right )} + \frac{3}{4} \left(-0.035281529513114 + 0.00875077955598942 i\right) e^{- 2.0 i \phi_{0}} e^{2.0 i \phi_{1}} \sin{\left (2 \theta_{0} \right )} \sin{\left (2 \theta_{1} \right )} + \frac{3}{4} \left(0.285095302985003 + 0.00922257353994807 i\right) e^{- 2.0 i \phi_{0}} e^{- 2.0 i \phi_{1}} \sin{\left (2 \theta_{0} \right )} \sin{\left (2 \theta_{1} \right )}$
# Measuring Points in Phase Space
We will now look at how to create the circuits to measure set points in phase space. The following results are based on the figures in [*RP Rundle, PW Mills, T Tilma, JH Samson, MJ Everitt, Phys. Rev. A 96, 022117*](r p rundle prl)
## Plaquette Visualisation
Staying with the bell state, we will look here at the slice where $\phi_0=\phi_1=0$ and we plot $\theta_0$ against $\theta_1$
```
theta1_points = 8
theta2_points = 8
number_of_points = theta1_points*theta2_points
the1 = [0]*number_of_points
the2 = [0]*number_of_points #initialize theta values
phis = [[0]*number_of_points]*number_of_qubits #set phi values to 0
point = 0
for i in range(theta1_points):
for k in range(theta2_points):
the1[point] = 2*i*np.pi/theta1_points
the2[point] = 2*k*np.pi/theta2_points #create the values of theta for all points on plot
point += 1
thetas = np.vstack((the1,the2))
bell_circuits = tomo.build_wigner_circuits(Q_program, 'bell', phis, thetas,
bell_qubits, qr, cr)
backend = 'local_qasm_simulator'
shots = 1024
bell_result = Q_program.execute(bell_circuits, backend=backend, shots=shots)
print(bell_result)
wdata = tomo.wigner_data(bell_result, bell_qubits,
bell_circuits, shots=shots)
wdata = np.matrix(wdata)
wdata = wdata.reshape(theta1_points,theta2_points)
plot_wigner_data(wdata, method='plaquette')
```
## Curve on a Wigner fucntion
We can also take a line around phase space and plot a curve. A useful line to take is the equator of the equal angle slice, below we plot this slice also for the Bell-state. When we have an $n$-qubit maximally entangled state of the form $\left(|0...0\rangle + |1...1\rangle\right)/\sqrt{2}$, the equatorial slice will show sinusoidal behaviour with frequency $n$.
```
equator_points = 64
theta = [np.pi/2]*equator_points
phi = [0]*equator_points
point = 0
for i in range(equator_points):
phi[i] = 2*i*np.pi/equator_points
thetas = np.vstack((theta,theta))
phis = np.vstack((phi,phi))
bell_eq_circuits = tomo.build_wigner_circuits(Q_program, 'bell', phis, thetas,
bell_qubits, qr, cr)
bell_eq_result = Q_program.execute(bell_eq_circuits, backend=backend, shots=shots)
wdata_eq = tomo.wigner_data(bell_eq_result, bell_qubits,
bell_eq_circuits, shots=shots)
plot_wigner_data(wdata_eq, method='curve')
```
Recreating the equatorial slice of a five qubit GHZ state, $\left(|00000\rangle - |11111\rangle\right)/\sqrt{2}$ from the paper, the following gates are needed...
```
Q_program = QuantumProgram()
number_of_qubits = 5
backend = 'local_qasm_simulator'
shots = 1024
ghz_qubits = [0, 1, 2, 3, 4]
qr = Q_program.create_quantum_register('qr',5)
cr = Q_program.create_classical_register('cr',5)
ghz = Q_program.create_circuit('ghz', [qr], [cr])
ghz.h(qr[0])
ghz.h(qr[1])
ghz.x(qr[2])
ghz.h(qr[3])
ghz.h(qr[4])
ghz.cx(qr[0],qr[2])
ghz.cx(qr[1],qr[2])
ghz.cx(qr[3],qr[2])
ghz.cx(qr[4],qr[2])
ghz.h(qr[0])
ghz.h(qr[1])
ghz.h(qr[2])
ghz.h(qr[3])
ghz.h(qr[4])
equator_points = 64
thetas = [[np.pi/2]*equator_points]*number_of_qubits
phi = [0]*equator_points
point = 0
for i in range(equator_points):
phi[i] = 2*i*np.pi/equator_points
phis = np.vstack((phi,phi,phi,phi,phi))
ghz_eq_circuits = tomo.build_wigner_circuits(Q_program, 'ghz', phis, thetas,
ghz_qubits, qr, cr)
ghz_eq_result = Q_program.execute(ghz_eq_circuits, backend=backend, shots=shots, timeout = 300)
wghzdata_eq = tomo.wigner_data(ghz_eq_result, ghz_qubits,
ghz_eq_circuits, shots=shots)
plot_wigner_data(wghzdata_eq, method='curve')
```
Since all the data for the plot is stored within `wghzdata_eq`, we can import matplotlib and go wild with different ways of plotting
```
import matplotlib.pyplot as plt
plt.plot(phi, wghzdata_eq, 'o')
plt.axis([0, 2*np.pi, -0.6, 0.6])
plt.show()
density_matrix = np.zeros((32,32))
density_matrix[0][0] = 0.5
density_matrix[0][31] = -0.5
density_matrix[31][0] = -0.5
density_matrix[31][31] = 0.5
plot_wigner_function(density_matrix, res=200)
```
| github_jupyter |
```
library(tidyverse)
library(caret)
library(doParallel)
library(xtable)
cl <- makePSOCKcluster(16)
registerDoParallel(cl)
```
### Data Loading
```
source("data_load.R")
path <- "../data"
files <- paste0(path, "/", list.files(path = path, pattern = ".csv"))
df <- load_from_path("../data") %>%
mutate(multi = factor(multi)) %>%
filter(max_peak %in% c(38,40,42,45,50,60))
set.seed(1)
N_TRAINING_DAYS = 10*7
N_RESAMPLES = 20
train_cols <- c("C", 'q0','q10','q25','q50','q75','q90','q100','mean','std','var',
"fft1", "fft2", "fft3", "fft4", "fft5", "fft6", "fft7", "fft8",
"fft1n", "fft2n", "fft3n", "fft4n", "fft5n", "fft6n", "fft7n", "fft8n")
gbm <- function(train.x, train.y){
fitControl <- trainControl(method = "repeatedcv", number=5, repeats=2)
train(x = train.x, y = train.y, method = "gbm", trControl = fitControl, verbose=F)
}
# train.x <- df_train[, train_cols] %>% as.data.frame()
# train.y <- df_train %>% .[["R"]]
# trained_model <- gbm(train.x, train.y)
# For one specific parameter combination (of max_peak, ...)
train_resamples <- function(x, R, C_reference, m){
train_splits <- createDataPartition(1:nrow(R),
times = N_RESAMPLES,
p = N_TRAINING_DAYS/nrow(x),
list=T)
print(train_splits)
improvements <- lapply(train_splits, function(train) {
trained_model <- m(x[train,], R[train,]) # train data on the partition, test on the non-partition
predictions <- predict(trained_model, x[-train,])
C_real <- max(C_reference[-train]) # maximum C_reference on test split
C_non_corrected <- max(x[-train,]$C) # maximum C calculated on test split
C_corrected <- max(x[-train,]$C - predictions) # maximum corrected C on test split
error_non_corrected <- C_real - C_non_corrected
error_corrected <- C_real - C_corrected
improvement <- abs(error_non_corrected) - abs(error_corrected)
improvement
}
)
improvements
}
train_on_df_group <- function(df_group){
train.x <- df_group[, train_cols] %>% as.data.frame()
train.y <- df_group %>% select(R) %>% as.data.frame()
train_resamples(train.x, train.y, df_group$C_reference, gbm)
}
df_subset <- df %>%
select(-day, -multi) %>%
filter(max_peak %in% c(40, 38), aggregation_interval %in% c(300, 900, 1800, 3600), !is.na(C))
results <- df_subset %>%
group_by(max_peak, aggregation_interval, aggregation_type) %>%
nest() %>%
mutate(improvements = map(data, train_on_df_group))
results %>% select(max_peak, aggregation_interval, aggregation_type, improvements) %>%
unnest() %>%
mutate(improvements = as.numeric(improvements)) %>%
group_by(max_peak, aggregation_interval, aggregation_type) %>%
summarize(median_improvement = median(improvements),
max_improvement = max(improvements)) %>%
gather(variable, value, median_improvement, max_improvement) %>%
unite(tmp, aggregation_type, variable) %>%
spread(tmp, value) -> table_var
table_var
print(xtable(table_var),
include.rownames=FALSE,
include.colnames = FALSE,
only.contents = TRUE,
booktabs = TRUE,
hline.after = 4,
file = "export/table_model_comparison.tex")
```
| github_jupyter |
## One Shot Learning with Siamese Networks
```
%matplotlib inline
import torchvision
import torchvision.datasets as dset
import torchvision.transforms as transforms
from torch.utils.data import DataLoader,Dataset
import matplotlib.pyplot as plt
import torchvision.utils
import numpy as np
import random
from PIL import Image
import torch
from torch.autograd import Variable
import PIL.ImageOps
import torch.nn as nn
from torch import optim
import torch.nn.functional as F
def imshow(img,text=None,should_save=False):
npimg = img.numpy()
plt.axis("off")
if text:
plt.text(75, 8, text, style='italic',fontweight='bold',
bbox={'facecolor':'white', 'alpha':0.8, 'pad':10})
plt.imshow(np.transpose(npimg, (1, 2, 0)))
plt.show()
def show_plot(iteration,loss):
plt.plot(iteration,loss)
plt.show()
class Config():
training_dir = "./data/train/"
testing_dir = "./data/test/"
train_batch_size = 64
train_number_epochs = 100
```
### Custom Dataset Class
This dataset generates a pair of images. 0 for geniune pair and 1 for imposter pair
```
class SiameseNetworkDataset(Dataset):
def __init__(self,imageFolderDataset,transform=None,should_invert=True):
self.imageFolderDataset = imageFolderDataset
self.transform = transform
self.should_invert = should_invert
def __getitem__(self,index):
img0_tuple = random.choice(self.imageFolderDataset.imgs)
#we need to make sure approx 50% of images are in the same class
should_get_same_class = random.randint(0,1)
if should_get_same_class:
while True:
#keep looping till the same class image is found
img1_tuple = random.choice(self.imageFolderDataset.imgs)
if img0_tuple[1]==img1_tuple[1]:
break
else:
while True:
#keep looping till a different class image is found
img1_tuple = random.choice(self.imageFolderDataset.imgs)
if img0_tuple[1] !=img1_tuple[1]:
break
img0 = Image.open(img0_tuple[0])
img1 = Image.open(img1_tuple[0])
img0 = img0.convert("L")
img1 = img1.convert("L")
if self.should_invert:
img0 = PIL.ImageOps.invert(img0)
img1 = PIL.ImageOps.invert(img1)
if self.transform is not None:
img0 = self.transform(img0)
img1 = self.transform(img1)
return img0, img1 , torch.from_numpy(np.array([int(img1_tuple[1]!=img0_tuple[1])],dtype=np.float32))
def __len__(self):
return len(self.imageFolderDataset.imgs)
```
### Using Image Folder Dataset
```
folder_dataset = dset.ImageFolder(root=Config.training_dir)
siamese_dataset = SiameseNetworkDataset(imageFolderDataset=folder_dataset,
transform=transforms.Compose([transforms.Resize((28,28)),
transforms.ToTensor()
])
,should_invert=False)
```
### Visualising some of the data
```
vis_dataloader = DataLoader(siamese_dataset,
shuffle=True,
num_workers=0,
batch_size=8)
dataiter = iter(vis_dataloader)
example_batch = next(dataiter)
concatenated = torch.cat((example_batch[0],example_batch[1]),0)
imshow(torchvision.utils.make_grid(concatenated))
print(example_batch[2].numpy())
```
### Neural Net Definition
```
class SiameseNetwork(nn.Module):
def __init__(self):
super(SiameseNetwork, self).__init__()
self.cnn1 = nn.Sequential(
nn.ReflectionPad2d(1),
nn.Conv2d(1, 4, kernel_size=3),
nn.ReLU(inplace=True),
nn.BatchNorm2d(4),
nn.ReflectionPad2d(1),
nn.Conv2d(4, 8, kernel_size=3),
nn.ReLU(inplace=True),
nn.BatchNorm2d(8),
nn.ReflectionPad2d(1),
nn.Conv2d(8, 8, kernel_size=3),
nn.ReLU(inplace=True),
nn.BatchNorm2d(8),
)
self.fc1 = nn.Sequential(
nn.Linear(8*28*28, 500),
nn.ReLU(inplace=True),
nn.Linear(500, 500),
nn.ReLU(inplace=True),
nn.Linear(500, 5))
def forward_once(self, x):
output = self.cnn1(x)
output = output.view(output.size()[0], -1)
output = self.fc1(output)
return output
def forward(self, input1, input2):
output1 = self.forward_once(input1)
output2 = self.forward_once(input2)
return output1, output2
```
### Contrastive Loss
```
class ContrastiveLoss(torch.nn.Module):
"""
Contrastive loss function.
Based on: http://yann.lecun.com/exdb/publis/pdf/hadsell-chopra-lecun-06.pdf
"""
def __init__(self, margin=2.0):
super(ContrastiveLoss, self).__init__()
self.margin = margin
def forward(self, output1, output2, label):
euclidean_distance = F.pairwise_distance(output1, output2)
loss_contrastive = torch.mean((1-label) * torch.pow(euclidean_distance, 2) +
(label) * torch.pow(torch.clamp(self.margin - euclidean_distance, min=0.0), 2))
return loss_contrastive
train_dataloader = DataLoader(siamese_dataset,
shuffle=True,
num_workers=0,
batch_size=Config.train_batch_size)
net = SiameseNetwork().cuda()
criterion = ContrastiveLoss()
optimizer = optim.Adam(net.parameters(),lr = 0.0005)
counter = []
loss_history = []
iteration_number= 0
for epoch in range(0,Config.train_number_epochs):
for i, data in enumerate(train_dataloader,0):
img0, img1 , label = data
img0, img1 , label = img0.cuda(), img1.cuda() , label.cuda()
optimizer.zero_grad()
output1,output2 = net(img0,img1)
loss_contrastive = criterion(output1,output2,label)
loss_contrastive.backward()
optimizer.step()
if i %10 == 0 :
print("Epoch number {}\n Current loss {}\n".format(epoch,loss_contrastive.item()))
iteration_number +=10
counter.append(iteration_number)
loss_history.append(loss_contrastive.item())
show_plot(counter,loss_history)
```
### Testing
```
folder_dataset_test = dset.ImageFolder(root=Config.testing_dir)
siamese_dataset = SiameseNetworkDataset(imageFolderDataset=folder_dataset_test,
transform=transforms.Compose([transforms.Resize((28,28)),
transforms.ToTensor()
])
,should_invert=False)
test_dataloader = DataLoader(siamese_dataset,num_workers=0,batch_size=1,shuffle=True)
dataiter = iter(test_dataloader)
x0,_,_ = next(dataiter)
for i in range(10):
_,x1,label2 = next(dataiter)
concatenated = torch.cat((x0,x1),0)
output1,output2 = net(Variable(x0).cuda(),Variable(x1).cuda())
euclidean_distance = F.pairwise_distance(output1, output2)
imshow(torchvision.utils.make_grid(concatenated),'Dissimilarity: {:.2f}'.format(euclidean_distance.item()))
```
| github_jupyter |
# Visualize L1 and L2 regularization
```
import numpy as np
import pandas as pd
from sklearn.linear_model import LinearRegression, Ridge, Lasso, LogisticRegression
from sklearn.ensemble import RandomForestClassifier, RandomForestRegressor
from sklearn.datasets import load_boston, load_iris, load_wine, load_digits, \
load_breast_cancer, load_diabetes
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix, precision_score, recall_score
from mpl_toolkits.mplot3d import Axes3D # required even though not ref'd!
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
%config InlineBackend.figure_format = 'retina'
def loss(x,y,b):
losses = []
for b_ in b:
losses.append( np.sum((y - (0 + b_*x))**2) )
return np.array(losses)
lmbda=1.0
penalty='l1'
yrange=(0,5)
mse_color = '#225ea8'
reg_color = '#fdae61'
sum_color = '#41b6c4'
fig, ax = plt.subplots(1,1,figsize=(4.2,3.2))
x = np.linspace(0,5,6)
x = (x - np.mean(x)) / np.std(x) # normalize x
y = 2 * x
b1 = np.linspace(0,4,1000)
mse = (b1-2)**2
# mse = loss(x,y,b=b1)
if penalty=='l1':
reg = lmbda * np.abs(b1)
else:
reg = lmbda * b1**2
if yrange is None:
yrange = (0, max(mse))
min_mse = np.min(mse)
min_mse_x = 2#b1[np.argmin(mse)]
min_loss = np.min(mse+reg)
min_loss_x = b1[np.argmin(mse+reg)]
# print(np.where(np.abs(mse-reg)<.01), b1[np.where(np.abs(mse-reg)<.01)])
# print(np.where(mse==reg))#, b1[np.argmin(mse-reg)])
ax.plot(b1, mse, lw=.75, c=mse_color, label="MSE Loss")#label="MSE Loss = $\Sigma(y-\\hat{y})^2$")
# verticle bar
ax.plot([lmbda,lmbda], [0,yrange[1]], ':', lw=1,c='grey')
ax.plot([-lmbda,-lmbda], [0,yrange[1]], ':', lw=1,c='grey')
ax.plot([0,0], [0,yrange[1]*1.1], lw=.5, c='k')
safe = plt.Rectangle((0,0), lmbda, yrange[1], facecolor="grey", alpha=0.1)
ax.add_patch(safe)
safe = plt.Rectangle((0,0), -lmbda, yrange[1], facecolor="grey", alpha=0.1)
ax.add_patch(safe)
#ax.text(.5,5.4, "$\lambda$", fontsize=15, horizontalalignment='center')
ax.text(0,4.6, 'Constraint', fontsize=11, horizontalalignment='center')
ax.text(0,4.3, 'zone', fontsize=11, horizontalalignment='center')
ax.annotate("Hard constraint",(1.0,3.5),xytext=(1.5,3.5),arrowprops=dict(arrowstyle="->",linewidth=.5))
ax.annotate("Min loss",(2,0),xytext=(2.2,2),arrowprops=dict(arrowstyle="->",linewidth=.5))
ax.annotate("Min regularized loss",(1.0,1.0),xytext=(1.2,2.7),arrowprops=dict(arrowstyle="->",linewidth=.5))
ax.set_xlabel("$\\beta$")
ax.set_ylabel("Loss")
ax.set_ylim(*yrange)
ax.legend(loc='upper right')
ax.set_xlim(-2,4.0)
ax.spines['left'].set_linewidth(.5)
ax.spines['bottom'].set_linewidth(.5)
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
plt.tight_layout()
plt.savefig("reg1D.svg", bbox_inches=0, pad_inches=0)
plt.show()
```
## L2
```
def plot_reg(penalty:('l1','l2')='l2',lmbda:float=1.0,show_reg=True,
xrange=None,yrange=None,fill=True,
dpi=200):
def loss(x,y,b):
losses = []
for b_ in b:
losses.append( np.sum((y - (0 + b_*x))**2) )
return np.array(losses)
mse_color = '#225ea8'
reg_color = '#fdae61'
sum_color = '#41b6c4'
fig, ax = plt.subplots(1,1)
# get x,y as line with slope beta = 2 + noise
x = np.linspace(0,5,6)
x = (x - np.mean(x)) / np.std(x) # normalize x
y = 2 * x# + np.random.uniform(0,1,size=len(x))
# lm = LinearRegression()
# lm.fit(x.reshape(-1,1),y)
# print(f"Slope of data is {lm.coef_[0]}")
b1 = np.linspace(0,4,1000)
mse = (b1-2)**2
# mse = loss(x,y,b=b1)
if penalty=='l1':
reg = lmbda * np.abs(b1)
else:
reg = lmbda * b1**2
if yrange is None:
yrange = (0, max(mse))
# print(lmbda)
# print("b1 cross", mse / lmbda, "len mse", len(mse))
# print("b1 cross", mse)
# print("b1 cross no lmbda", b1[np.argmin(mse)])
min_mse = np.min(mse)
min_mse_x = 2#b1[np.argmin(mse)]
min_loss = np.min(mse+reg)
min_loss_x = b1[np.argmin(mse+reg)]
# print(np.where(np.abs(mse-reg)<.01), b1[np.where(np.abs(mse-reg)<.01)])
# print(np.where(mse==reg))#, b1[np.argmin(mse-reg)])
ax.plot(b1, mse, lw=.75, label="$\Sigma(y-\\hat{y})^2$", c=mse_color)
if show_reg:
reg_label = "$\\lambda |\\beta_1|$" if penalty=='l1' else "$\\lambda \\beta_1^2$"
ax.plot(b1, reg, lw=.3, label=reg_label, c=reg_color)
ax.plot(b1, mse + reg, '--', lw=1, label="$(y-\\hat{y})^2$ + "+reg_label, c=sum_color)
ax.annotate("Min mse+reg",(min_loss_x,min_loss),xytext=(min_loss_x,min_loss*1.2),
arrowprops=dict(arrowstyle="->"), horizontalalignment='right')
# vertible bar
ax.plot([min_loss_x,min_loss_x], [0,yrange[1]],lw=.5,c='grey')
if fill:
ax.fill_between(b1, reg, 0, color=reg_color, alpha=.15)
delta_x = min_loss_x-min_mse_x
delta_midpoint_x = (min_mse_x+min_loss_x)/2
ax.text(delta_midpoint_x,yrange[1]-yrange[1]*.1,f"$\\lambda={lmbda:.2f}$ ({penalty.upper()})",
horizontalalignment='center')
ax.text(delta_midpoint_x,yrange[1]-yrange[1]*.15,f"$\\Delta_{{\\beta}}$={delta_x:.1f}",
horizontalalignment='center')
# ax.text(2,15,f"shifts min loss",
# horizontalalignment='center')
if not np.isclose(min_loss_x,min_mse_x):
ax.arrow(min_mse_x, yrange[1]-yrange[1]*.03, (min_loss_x-min_mse_x), 0, lw=.5,
head_length=0.05,
head_width=0.2,
length_includes_head=True)
# verticle bar
ax.plot([min_mse_x,min_mse_x], [0,yrange[1]],lw=.5,c='grey')
ax.annotate("Min mse",(2,0),xytext=(2.2,2),arrowprops=dict(arrowstyle="->"))
ax.set_xlabel("$\\beta_1$")
ax.set_ylabel("Loss")
if xrange is not None:
ax.set_xlim(*xrange)
ax.set_ylim(*yrange)
ax.legend(loc='upper right')
plot_reg('l2',show_reg=False, yrange=(0,6))
plot_reg('l2',lmbda=.1,yrange=(0,6))
plot_reg('l2',lmbda=3.5, yrange=(0,6))
plot_reg('l2',lmbda=15, xrange=(-.1,1.5), yrange=(0,5)) # zoom in
plot_reg('l2',lmbda=300, xrange=(-.1,1.5), yrange=(0,5)) # zoom in
```
## L1
```
plot_reg('l1',lmbda=.5)
plot_reg('l1',lmbda=1)
plot_reg('l1',lmbda=4, yrange=(0,10))
```
## Animate
```
import glob
import os
from PIL import Image as PIL_Image
def animate(penalty, lrange, step, yrange, dpi=100, duration=80):
plt.close()
for f in glob.glob(f'/tmp/{penalty}-frame-*.png'):
os.remove(f)
for lmbda in np.arange(*lrange,step):
plot_reg(penalty,lmbda=lmbda,fill=False,yrange=yrange)
# print(f"/tmp/{penalty}-frame-{lmbda:06.3f}.png")
plt.savefig(f"/tmp/{penalty}-frame-{lmbda:06.3f}.png", bbox_inches=0, pad_inches=0, dpi=dpi)
plt.close()
images = [PIL_Image.open(image) for image in sorted(glob.glob(f'/tmp/{penalty}-frame-*.png'))]
images += reversed(images)
images[0].save(f'/tmp/{penalty}-animation.gif',
save_all=True,
append_images=images[1:],
duration=duration,
loop=0)
animate('l1',lrange=(0.0,5), step=.2, yrange=(0,6), dpi=200)
animate('l2',lrange=(0.0,11), step=.3, yrange=(0,6), dpi=200, duration=100)
```
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
df = pd.read_json("http://127.0.0.1:5500/api/v1.0/perthcity")
df
```
# Filter Outliers
To git rid of outliers, we need to filter:
* Properties sold before 2019
* landsize over 2000 sqm
* carspace,bedroom over 6
* Properties price over 3M
```
df['sold_date']= df['sold_date'].astype('datetime64[ns]')
df['sold_date'] = df['sold_date'].dt.strftime('%Y')
df['sold_date'] = df['sold_date'].astype('int')
After2019_df = df[(df['sold_date']>2019)]
After2019_df
Less2000sqm_df = After2019_df[(After2019_df['land_size']<2000)]
carspaceLessThan7_df = Less2000sqm_df[(Less2000sqm_df['car_space']<7)]
BedroomsLessthan7_df = carspaceLessThan7_df[(carspaceLessThan7_df['bedrooms']<7)]
filtered_df = BedroomsLessthan7_df[(BedroomsLessthan7_df['price']<3000000)]
filtered_df
```
# Data Preporcessing
```
#only getting landed properties for the machine learning
house = filtered_df[(filtered_df['property_type']=='House') | (filtered_df['property_type']=='Villa') | (filtered_df['property_type']=='Townhouse')]
house
house.columns
# for i in range(0,len(house['sale_id'])):
# if house.iloc[i,13] =='Villa':
# house.iloc[i,13] = 1
# elif house.iloc[i,13] =='Townhouse':
# house.iloc[i,13]= 2
# elif house.iloc[i,13] =='House':
# house.iloc[i,13] = 3
# house
house['perth'] = ''
house['east_perth'] = ''
house['west_perth'] = ''
house['northbridge'] = ''
house['crawley'] = ''
house['nedlands'] = ''
# house['villa'] = ''
# house['townhouse'] = ''
# house['house'] = ''
for i in range(0,len(house['sale_id'])):
if house.iloc[i,19] =='Perth':
house.iloc[i,20] = 1
house.iloc[i,21:26] = 0
elif house.iloc[i,19] =='East Perth':
house.iloc[i,21] = 1
house.iloc[i,20] = 0
house.iloc[i,22:26] = 0
elif house.iloc[i,19] =='West Perth':
house.iloc[i,22] = 1
house.iloc[i,20:22] = 0
house.iloc[i,23:26] = 0
elif house.iloc[i,19] =='Northbridge':
house.iloc[i,23] = 1
house.iloc[i,20:23] = 0
house.iloc[i,24:26] = 0
elif house.iloc[i,19] =='Crawley':
house.iloc[i,24] = 1
house.iloc[i,20:24] = 0
house.iloc[i,25] = 0
elif house.iloc[i,19] =='Nedlands':
house.iloc[i,25] = 1
house.iloc[i,20:25] = 0
# if house.iloc[i,13] =='Villa':
# house.iloc[i,26] = 1
# house.iloc[i,27:29] = 0
# elif house.iloc[i,13] =='Townhouse':
# house.iloc[i,27]= 1
# house.iloc[i,26]= 0
# house.iloc[i,28]= 0
# elif house.iloc[i,13] =='House':
# house.iloc[i,28] = 1
# house.iloc[i,26:28] = 0
house
# Assign the data to X and y
X = house[["bedrooms", "bathrooms", "car_space", "land_size", "building_size", "built_date", "perth", "west_perth", "east_perth", "northbridge", "crawley", "nedlands"]]
y = house["price"].values.reshape(-1, 1)
print(X.shape, y.shape)
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y)
from sklearn.preprocessing import StandardScaler
# Create a StandardScater model and fit it to the training data
X_scaler = StandardScaler().fit(X_train)
y_scaler = StandardScaler().fit(y_train)
# Transform the training and testing data using the X_scaler and y_scaler models
X_train_scaled = X_scaler.transform(X_train)
X_test_scaled = X_scaler.transform(X_test)
y_train_scaled = y_scaler.transform(y_train)
y_test_scaled = y_scaler.transform(y_test)
```
# Keras Regression Deep Learning Model
```
# Creating a Neural Network Model
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Activation
from tensorflow.keras.optimizers import Adam
# Create model and add layers
model = Sequential()
model.add(Dense(units=12, activation='relu', input_dim=12))
model.add(Dense(units=1))
model.compile(optimizer='Adam',
loss='mse')
model.fit(
X_train_scaled,
y_train_scaled,
validation_data=(X_test_scaled,y_test_scaled),
epochs=300)
model.summary()
# model_loss, model_accuracy = model.evaluate(
# X_test_scaled, y_test_scaled, verbose=2)
# print(f"Loss: {model_loss}, Accuracy: {model_accuracy}")
loss_df = pd.DataFrame(model.history.history)
loss_df.plot(figsize=(12,8))
y_pred = model.predict(X_test_scaled)
from sklearn import metrics
print('MAE:', metrics.mean_absolute_error(y_test_scaled, y_pred))
print('MSE:', metrics.mean_squared_error(y_test_scaled, y_pred))
print('RMSE:', np.sqrt(metrics.mean_squared_error(y_test_scaled, y_pred)))
print('VarScore:',metrics.explained_variance_score(y_test_scaled,y_pred))
# Visualizing Our predictions
fig = plt.figure(figsize=(10,5))
plt.scatter(y_test_scaled,y_pred)
# Perfect predictions
plt.plot(y_test_scaled,y_test_scaled,'r')
#test
# suburb needs to be categorical
X_test = X_scaler.transform([[4,3,2,175,186,2019,0,0,0,0,1,0]])
predictions = model.predict(X_test)
results = y_scaler.inverse_transform(predictions)
results
# compare actual output values with predicted values
df1 = X_test
df1['Actual'] = y_test.reshape(1,-1)[0]
df1['Keras_Regression_Predicted'] = y_scaler.inverse_transform(model.predict(X_test_scaled))
df1.head(10)
```
| github_jupyter |
# Day 1: An introduction to Python programming
References:
* https://sites.google.com/a/ucsc.edu/krumholz/teaching-and-courses/python-15/class-1
Right now we are going to go over a few basics of programming using Python and also some fun tidbits about how to mark up our "jupyter notebooks" with text explainations and figures of what we are doing.
If you have been programming for a while, this will serve as a bit of a review, but since most folks don't primarily program in Python, this will be an opportunity to learn a bit more detail about the nuances of Python.
If you are very new to programming, this might all seem like gibberish right now. This is completely normal! Programming is one of those things that is hard to understand in the abstract, but gets much more easy the more you do it and can see what sorts of outputs you get.
This part will be a little dry, but soon we'll get to using the things we learn here to do more fun stuff.
Also! These notes are online. Hopefully I won't go to fast, but if I do please feel free to raise your hand and tell me to slow down gosh darn it! If you just want to make sure you got something - feel free to refer back to these notes. BEWARE: you will learn this better if you try to follow along in class and not copy directly from here - but I assume you can figure out for yourself what methods you want to employ best in this class.
# 1. Introduction to jupyter notebooks
* code vs comments
* markdown "cheat sheet"
* running a cell
* using latex to do math equations: $-G M_1 m_2/r^2$ or $- \frac{G M m}{r^2}$
* latex math "cheat sheet"
## 1. Using Python as a calculator
It is possible to interact with python in many ways. Let's start with something simple - using this notebook+python as a calculator
```
# lets add 2+3
2+3
# ALSO: note what I did there with the "#" -> this is called a comment,
# and it allows us to add in notes to ourselves OR OTHERS
# Commenting is SUPER important to (1) remember what you did and
# (2) tell others that you're working with what you did
#In fact, comments are part of a "good coding practice"
# Python even has the nicities to tell you all about
# what is good coding practice:
import this
```
The above sentences will be come clearer the more you program in Python! :)
```
2*3
2-3
4/2
# lets say I want to write 2 raised to the power of 3, i.e. 2*2*2 = 8
# python has special syntax for that:
2**3
```
Python knows the basic arithmetic operations plus (+), minus (-), times (*), divide (/), and raise to a power (**). It also understands parentheses, and follows the normal rules for order of operations:
```
1+2*3
(1+2)*3
```
## 2. Simple variables
We can also define variables to store numbers, and we can perform arithmetic on those variables. Variables are just names for boxes that can store values, and on which you can perform various operations. For example:
```
a=4
a+1
a
# note that a itself hasn't changed
a/2
# now I can change the value of a by reassigning it to its original value + 1
a = a+1
# short hand is a += 1
a
a**2
```
There's a subtle but important point to notice here, which is the meaning of the equal sign. In mathematics, the statement that a = b is a statement that two things are equal, and it can be either true or false. In python, as in almost all other programming languages, a = b means something different. It means that the value of the variable a should be changed to whatever value b has. Thus the statement we made a = a + 1 is not an assertion (which is obviously false) that a is equal to itself plus one. It is an instruction to the computer to take the variable a, and 1 to it, and then store the result back into the variable a. In this example, it therefore changes the value of a from 4 to 5.
One more point regrading assignments: the fact that = means something different in programming than it does in mathematics implies that the statements a = b and b = a will have very different effects. The first one causes the computer to forget whatever is stored in a and replace it by whatever is stored in b. The second statement has the opposite effect: the computer forgets what is stored in b, and replaces it by whatever is stored in a.
For example, I can use a double equals sign to "test" whether or not a is equal to some value:
```
a == 5
a == 6
```
More on this other form of an equal sign when we get to flow control -> stay tuned!
The variable a that we have defined is an integer, or int for short. We can find this out by asking python:
```
type(a)
```
Integers are exactly what they sound like: they hold whole numbers, and operations involving them and other whole numbers will always yield whole numbers. This is an important point:
```
a/2
```
Why is 5/2 giving 2? The reason is that we're dividing integers, and the result is required to be an integer. In this case, python rounds down to the nearest integer. If we want to get a non-integer result, we need to make the operation involve a non-integer. We can do this just by changing the 2 to 2.0, or even just 2., since the trailing zero is assumed:
```
a/2.
```
If we assign this to a variable, we will have a new type of variable: a floating point number, or float for short.
```
b = a/2.
type(b)
```
A floating point variable is capable of holding real numbers. Why have different types of variables for integers versus non-integer real numbers? In mathematics there is no need to make the distinction, of course: all integers are real numbers, so it would seem that there should be no reason to have a separate type of variable to hold integers. However, this ignores the way computers work. On a computer, operations involving integers are exact: 1 + 1 is exactly 2. However, operations on real numbers are necessarily inexact. I say necessarily because a real number is capable of having an arbitrary number of decimal places. The number pi contains infinitely many digits, and never repeats, but my computer only comes with a finite amount of memory and processor power. Even rational numbers run into this problem, because their decimal representation (or to be exact their representation in binary) may be an infinitely repeating sequence. Thus it is not possible to perform operations on arbitrary real numbers to exact precision. Instead, arithmetic operations on floating point numbers are approximate, with the level of accuracy determined by factors like how much memory one wants to devote to storing digits, and how much processor time one wants to spend manipulating them. On most computers a python floating point number is accurate to about 1 in 10^15, but this depends on both the architecture and on the operations you perform. That's enough accuracy for many purposes, but there are plenty of situations (for example counting things) when we really want to do things precisely, and we want 1 + 1 to be exactly 2. That's what integers are there for.
A third type of very useful variable is strings, abbreviated str. A string is a sequence of characters, and one can declare that something is a string by putting characters in quotation marks (either " or ' is fine):
```
c = 'alice'
type(c)
```
The quotation marks are important here. To see why, try issuing the command without them:
```
c=alice
```
This is an error message, complaining that the computer doesn't know what alice is. The problem is that, without the quotation marks, python thinks that alice is the name of a variable, and complains when it can't find a variable by that name. Putting the quotation marks tells python that we mean a string, not a variable named alice.
Obviously we can't add strings in the same sense that we add numbers, but we can still do operations on them. The plus operation concatenates two strings together:
```
d = 'bob'
c+d
```
There are a vast number of other things we can do with strings as well, which we'll discuss later.
In addition to integers, floats, and strings, there are three other types of variables worth mentioning. Here we'll just mention the variable type of Boolean variable (named after George Boole), which represents a logical value. Boolean variables can be either True or False:
```
g=True
type(g)
```
Boolean variables can have logic operations performed on them, like not, and, and or:
```
not g
h = False
g and h
g or h
```
The final type of variable is None. This is a special value that is used to designate something that has not been assigned yet, or is otherwise undefined.
```
j = None
j
# note: nothing prints out since this variable isn't anything!
```
## 3. One dimensional arrays with numpy
The variables we have dealt with so far are fairly simple. They represent single values. However, for scientific or numeric purposes we often want to deal with large collections of numbers. We can try to do this with a "natively" supported Python datastructure called a list:
```
myList = [1, 2,3, 4]
myList
```
You can do some basic things with lists like add to each element:
```
myList[0] += 5
myList
# so now you can see that the first element of the list is now 1+5 = 6
```
You can also do fun things with lists like combine groups of objects with different types:
```
myList = ["Bob", "Linda", 5, 6, True]
myList
```
However, lists don't support adding of numbers across the whole array, or vector operations like dot products.
Formally, an array is a collection of objects that all have the same type: a collection of integers, or floats, or bools, or anything else. In the simplest case, these are simply arranged into a numbered list, one after another. Think of an array as a box with a bunch of numbered compartments, each of which can hold something. For example, here is a box with eight compartments.

We can turn this abstract idea into code with the numpy package, as Python doesn't natively support these types of objects. Lets start by importing numpy:
```
import numpy as np # here we are importing as "np" just for brevity - you'll see that a lot
# note: if you get an error here try:
#!pip install numpy
# *before* you try to import anything
```
We can start by initializing an empty array with 8 entries, to match our image above. There are several ways of doing this.
```
# we can start by calling the "empty" function
array1 = np.empty(8)
array1
# here you can see that we have mostly zero, or almost zeros in our array
# we can also specifically initial it with zeros:
array2 = np.zeros(8)
array2
# so this looks a little nicer
# we can also create a *truely* empty array like so:
array3 = np.array([])
array3
# then, to add to this array, we can "append" to the end of it like so:
array3 = np.append(array3, 0)
array3
```
Of course, we'd have to do the above 8 times and if we do this by hand it would be a little time consuming. We'll talk about how to do such an operation more efficiently using a "for loop" a little later in class.
Let's say we want to fill our array with the following elements:

We can do this following the few different methods we discussed. We could start by calling "np.empty" or "np.zeros" and then fill each element one at a time, or we can convert a *list* type into an *array* type on initilization of our array. For example:
```
array4 = np.array([10,11,12,13,14,15,16,17])
array4
```
There are even functions in numpy we can use to create new types of arrays. For example, we could have also created this same array as follows:
```
array5=np.arange(10,18,1)
array5
# note that here I had to specify 10-18 or one more than the array I wanted
```
We can also make this array with different spacing:
```
array6 = np.arange(10,18,2)
array6
```
We can compare operations with arrays and lists, for example:
```
myList = [5, 6, 7]
myArray = np.array([5,6,7])
myList, myArray
```
So, things look very similar. Lets try some operations with them:
```
myList[0], myArray[0]
```
So, they look very much the same... lets try some more complicated things.
```
myList[0] + 5, myArray[0] + 5
myArray + 5
myList + 5
```
So here we can see that while we can add a number to an array, we can't just add a number to a list. What does adding a number to myArray look like?
```
myArray, myArray+5
```
So we can see that adding an number to an array just increases all elements by the number.
```
# we can also increament element by element
myArray + [1, 2, 3]
```
There are also several differences in "native" operations with arrays vs lists. We can learn more about this by using a tab complete.
```
myList.
myArray.
```
As you can see myArray supports a lot more operations. For example:
```
# we can sum all elements of our array
myArray, myArray.sum()
# or we can take the mean value:
myArray.mean()
# or take things like the standard deviation, which
# is just a measurement of how much the array varies
# overall from the mean
myArray.std()
```
We can do some more interesting things with arrays, for example, specify what their "type" is:
```
myFloatArray = np.zeros([5])
myFloatArray
```
Compare this by if we force this array to be integer type:
```
myIntArray = np.zeros([5],dtype='int')
myIntArray
# we can see that there are no decimals after the zeros
# this is because this array will only deal with
# whole,"integer" numbers
```
How do we access elements of an array? Lets see a few different ways.
```
myArray = np.arange(0,10,1)
myArray
# just the first few elements:
myArray[0:5]
# all but the first element:
myArray[1:]
# in reverse:
myArray[::-1]
# all but the last 2 elements:
myArray[:-2]
# elements between 2 & 7
myArray[2:7]
# note: *is* included, but not 7
```
## Multiple dimension arrays
Arrays aren't just a list of numbers, we can also make *matricies* of arrays.
This is just a fancy way of saying "arrays with multiple dimensions".
For example, lets say we want to make a set of numbers that has entries along a row *and* column. Something that looks like this:

We can do this with the following call:
```
x2d=np.array([[10,11,12,13,14,15,16,17], [20,21,22,23,24,25,26,27], [30, 31, 32, 33, 34, 35, 36, 37]])
x2d
```
Now note, we can use some of the same array calls we used before:
```
x2d[0,:]
x2d[:,0]
x2d[1:,:]
```
We can even use functions like zeros to make multi-dimensional arrays with all entries equal to zero:
```
x2d2 = np.zeros((3,7))
x2d2
```
Once you've defined an array, you have a few ways to get info about the array. We used "mean" above, but you might also want to check that its shape is what you think it should be (I do this a lot!):
```
x2d.shape
```
There are *many* ways to manipulate arrays and call them and if you're feeling overwhelmed at this point, that is ok! We'll get plenty of practice using these in the future.
# 4. Dictionaries
There is one more data type that you might come across in Python a dictionary/directory. I think it might be called a "directory" but I've always said dictionary and it might be hard to change at this point :)
For brevity I'll just be calling it a "dict" anyway!
```
# the calling sequence is a little weird, but its essentially a way to "name" components of our dict
myDict = {"one":1, "A string":"My String Here", "array":np.array([5,6,6])}
```
Now we can call each of these things by name:
```
myDict["one"]
myDict["A string"]
myDict["array"]
```
In this course we'll be dealing mostly with arrays, and a little bit with lists & dicts - so if the data structures like dicts or "sets", which we didn't cover but you can read up on your own if you're super curious, seem weird that is ok! You will have many opportunties to work with these sorts of things more as you go on in your programming life.
# 5. Simple plots with arrays
Lets now start to make some plots with our arrays. To do this, we have to import a plotting library - this is just like what we did to import our numpy library to help us do stuff with arrays.
```
import matplotlib.pyplot as plt
# note: there is actually a *lot* of matplotlib sub-libraries
# we'll just start with pyplot
# again, if this doesn't work you can try
#!pip install matplotlib
# OR
#!conda install matplotlib
# lets try a simple plot
plt.plot([5,6], [7,8])
# ok neat! we see a plot with
# x going from 5-6, and y going from 7-8
```
Let's combine our numpy array stuff with plots:
```
x = np.arange(0,2*np.pi,0.01)
x
# so, here we see that x goes from 0 to 2*PI in steps of 0.01
# now, lets make a plot of sin(x)
y = np.sin(x)
plt.plot(x,y)
```
Ok, that's pretty sweet, but maybe we can't recall what we are plotting and we want to put some labels on our plot.
```
plt.plot(x,y)
plt.xlabel('x value from 0 to PI')
plt.ylabel('sin(x)')
plt.title('My first plot!')
```
Finally, note we get this text that gives us info about our plot that we might not want to show each time. We can explicitly "show" this plot:
```
plt.plot(x,y)
plt.xlabel('x value from 0 to PI')
plt.ylabel('sin(x)')
plt.title('My first plot!')
plt.show()
```
So, now let's say we want to save our lovely plot. Lets try that!
```
plt.plot(x,y)
plt.xlabel('x value from 0 to PI')
plt.ylabel('sin(x)')
plt.title('My first plot!')
plt.savefig('myAMAZINGsinplot.png')
```
Well, it seems like nothing happened. Why? So, basically our plot gets showed to us, but it *also* gets saved to disk. But where? Well by default, since we didn't give a full path, it saved to whatever directory we are running this jupyter notebook from. Lets figure out where that is:
```
# there are a few ways to figure this out
# we'll use this opportunty to do a little command-line
!pwd # MAC
# so the "!" is called an "escape" character - this just
# shunts us out of this notebook and gives us access to our
# main "terminal" or "shell" - basically the underlying
# structure of our computer
# note, in windows this equivalent is !cd
# see: https://www.lemoda.net/windows/windows2unix/windows2unix.html
```
We have several options at this point to open our image. We can save it in a directory where we easily know where it is, or we can open it up from here. For example:
```
!open /Users/jillnaiman1/csci-p-14110/lesson01/myAMAZINGsinplot.png
# the above is for macs!!
```
So we can change the location of where this saves so that we are saving all of our files to one place.
There are several ways to do this. You can open a file folder as you normally would and make a new folder and figure out where it is on disk and save to there. We can also do this using some command lines.
First we'll make a new directory, this command is the same on both Macs & Windows. First, lets remind ourselves where we are:
```
!pwd
```
Usually it will be something like "/Users" and then your user name and then maybe another directory. These are subfolders where different folders live. We can check out what is in a particular folder using "ls" or "dir" on a Windows machine like so:
```
!ls
```
So this is now telling me what is in the current directory that I found with !pwd. Lets say we want to make a new folder for this class in our "home" directory - this is what your directory/folder with your user name is generally called. We should first check out what is already in there, with that we can use "ls" or "dir" like so:
```
!ls /Users/jillnaiman1/
```
So now we can see a list of all the stuff that is in there. To make a new directory we can use the command "mkdir" on both Macs and Windows:
```
!mkdir /Users/jillnaiman1/MyNewDir
```
Now I can save my image to this directory like so:
```
plt.plot(x,y)
plt.xlabel('x value from 0 to PI')
plt.ylabel('sin(x)')
plt.title('My first plot!')
plt.savefig('/Users/jillnaiman1/MyNewDir/myAMAZINGsinplot.png')
```
Now I can see this file in that directory with "ls":
```
!ls /Users/jillnaiman1/MyNewDir
```
And I can also open it from there:
```
!open /Users/jillnaiman1/MyNewDir/myAMAZINGsinplot.png
```
# 6. Examples & exercises
1. plot cos(x)
2. plot a*cos(x) + b*sin(c*x) for different numbers a, b, c
If you're like "what?" at this point - this is TOTally normal. If you've done little to no programming this is a whole new way of thinking! Don't panic. We'll get through it together. :D
bonus: how do you plot various combinations of a, b, c?
bonus bonus: what ways can you plot functions of 2 variables? How would you figure that out? (Google might be your friend here :) )
| github_jupyter |
### Author : Sanjoy Biswas
### Project : Credit Card Fraud Detection
### Email : sanjoy.eee32@gmail.com
In this notebook I will try to predict fraud transactions from a given data set. Given that the data is imbalanced, standard metrics for evaluating classification algorithm (such as accuracy) are invalid. I will focus on the following metrics: Sensitivity (true positive rate) and Specificity (true negative rate). Of course, they are dependent on each other, so we want to find optimal trade-off between them. Such trade-off usually depends on the application of the algorithm, and in case of fraud detection I would prefer to see high sensitivity (e.g. given that a transaction is fraud, I want to be able to detect it with high probability).
**IMPORTING LIBRARIES:**
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from pylab import rcParams
import warnings
warnings.filterwarnings('ignore')
```
**READING DATASET :**
```
data=pd.read_csv('/kaggle/input/creditcardfraud/creditcard.csv')
data.head()
```
**NULL VALUES:**
```
data.isnull().sum()
```
**Thus there are no null values in the dataset.**
**INFORMATION**
```
data.info()
```
**DESCRIPTIVE STATISTICS**
```
data.describe().T.head()
data.shape
```
**Thus there are 284807 rows and 31 columns.**
```
data.columns
```
**FRAUD CASES AND GENUINE CASES**
```
fraud_cases=len(data[data['Class']==1])
print(' Number of Fraud Cases:',fraud_cases)
non_fraud_cases=len(data[data['Class']==0])
print('Number of Non Fraud Cases:',non_fraud_cases)
fraud=data[data['Class']==1]
genuine=data[data['Class']==0]
fraud.Amount.describe()
genuine.Amount.describe()
```
**EDA**
```
data.hist(figsize=(20,20),color='lime')
plt.show()
rcParams['figure.figsize'] = 16, 8
f,(ax1, ax2) = plt.subplots(2, 1, sharex=True)
f.suptitle('Time of transaction vs Amount by class')
ax1.scatter(fraud.Time, fraud.Amount)
ax1.set_title('Fraud')
ax2.scatter(genuine.Time, genuine.Amount)
ax2.set_title('Genuine')
plt.xlabel('Time (in Seconds)')
plt.ylabel('Amount')
plt.show()
```
**CORRELATION**
```
plt.figure(figsize=(10,8))
corr=data.corr()
sns.heatmap(corr,cmap='BuPu')
```
**Let us build our models:**
```
from sklearn.model_selection import train_test_split
```
**Model 1:**
```
X=data.drop(['Class'],axis=1)
y=data['Class']
X_train,X_test,y_train,y_test=train_test_split(X,y,test_size=0.30,random_state=123)
from sklearn.ensemble import RandomForestClassifier
rfc=RandomForestClassifier()
model=rfc.fit(X_train,y_train)
prediction=model.predict(X_test)
from sklearn.metrics import accuracy_score
accuracy_score(y_test,prediction)
```
**Model 2:**
```
from sklearn.linear_model import LogisticRegression
X1=data.drop(['Class'],axis=1)
y1=data['Class']
X1_train,X1_test,y1_train,y1_test=train_test_split(X1,y1,test_size=0.3,random_state=123)
lr=LogisticRegression()
model2=lr.fit(X1_train,y1_train)
prediction2=model2.predict(X1_test)
accuracy_score(y1_test,prediction2)
```
**Model 3:**
```
from sklearn.tree import DecisionTreeRegressor
X2=data.drop(['Class'],axis=1)
y2=data['Class']
dt=DecisionTreeRegressor()
X2_train,X2_test,y2_train,y2_test=train_test_split(X2,y2,test_size=0.3,random_state=123)
model3=dt.fit(X2_train,y2_train)
prediction3=model3.predict(X2_test)
accuracy_score(y2_test,prediction3)
```
**All of our models performed with a very high accuracy.**
| github_jupyter |
## ACA-Py Webhook Event Listeners
```
listeners = []
```
## Connection Listener
This will get called in relation to the connections protocol
```
# Receive connection messages
def connections_handler(payload):
state = payload["state"]
connection_id = payload["connection_id"]
their_role = payload["their_role"]
routing_state = payload["routing_state"]
print("----------------------------------------------------------")
print("Connection Webhook Event Received")
print("Connection ID : ", connection_id)
print("State : ", state)
print("Routing State : ", routing_state)
print("Their Role : ", their_role)
print("----------------------------------------------------------")
if state == "invitation":
# Your business logic
print("invitation")
elif state == "request":
# Your business logic
print("request")
elif state == "response":
# Your business logic
print("response")
elif state == "active":
# Your business logic
print(
colored(
"Connection ID: {0} is now active.".format(connection_id),
"green",
attrs=["bold"],
)
)
connection_listener = {"handler": connections_handler, "topic": "connections"}
listeners.append(connection_listener)
```
## Issue Credential Listener
### Role: Issuer
```
def issuer_handler(payload):
connection_id = payload["connection_id"]
exchange_id = payload["credential_exchange_id"]
state = payload["state"]
role = payload["role"]
print("\n---------------------------------------------------\n")
print("Handle Issue Credential Webhook")
print(f"Connection ID : {connection_id}")
print(f"Credential exchange ID : {exchange_id}")
print("Agent Protocol Role : ", role)
print("Protocol State : ", state)
print("\n---------------------------------------------------\n")
if state == "offer_sent":
print(f"Offering: {attributes}")
## YOUR LOGIC HERE
elif state == "request_received":
print("Request for credential received")
## YOUR LOGIC HERE
elif state == "credential_sent":
print("Credential Sent")
## YOUR LOGIC HERE
issuer_listener = {"topic": "issue_credential", "handler": issuer_handler}
listeners.append(issuer_listener)
```
### Role: Holder
```
def holder_handler(payload):
connection_id = payload["connection_id"]
exchange_id = payload["credential_exchange_id"]
state = payload["state"]
role = payload["role"]
print("\n---------------------------------------------------\n")
print("Handle Issue Credential Webhook")
print(f"Connection ID : {connection_id}")
print(f"Credential exchange ID : {exchange_id}")
print("Agent Protocol Role : ", role)
print("Protocol State : ", state)
print("\n---------------------------------------------------\n")
print("Handle Credential Webhook Payload")
if state == "offer_received":
attributes = proposal["credential_proposal"]["attributes"]
print(attributes)
print("Credential Offer Recieved")
proposal = payload["credential_proposal_dict"]
print(
"The proposal dictionary is likely how you would understand and display a credential offer in your application"
)
print("\n", proposal)
print("\n This includes the set of attributes you are being offered")
attributes = proposal["credential_proposal"]["attributes"]
print(attributes)
## YOUR LOGIC HERE
elif state == "request_sent":
print(
"\nA credential request object contains the commitment to the agents master secret using the nonce from the offer"
)
## YOUR LOGIC HERE
elif state == "credential_received":
print("Received Credential")
## YOUR LOGIC HERE
elif state == "credential_acked":
## YOUR LOGIC HERE
credential = payload["credential"]
print("Credential Stored\n")
print(credential)
print(
"\nThe referent acts as the identifier for retrieving the raw credential from the wallet"
)
# Note: You would probably save this in your application database
credential_referent = credential["referent"]
print("Referent", credential_referent)
holder_listener = {"topic": "issue_credential", "handler": holder_listener}
listeners.append(holder_listener)
```
## Present Proof Listener
### Role: Verifier
```
def verifier_proof_handler(payload):
role = payload["role"]
connection_id = payload["connection_id"]
pres_ex_id = payload["presentation_exchange_id"]
state = payload["state"]
print("\n---------------------------------------------------------------------\n")
print("Handle present-proof")
print("Connection ID : ", connection_id)
print("Presentation Exchange ID : ", pres_ex_id)
print("Protocol State : ", state)
print("Agent Role : ", role)
print("Initiator : ", payload["initiator"])
print("\n---------------------------------------------------------------------\n")
if state == "request_sent":
print("Presentation Request\n")
print(payload["presentation_request"])
print(
"\nThe presentation request is encoded in base64 and packaged into a DIDComm Message\n"
)
print(payload["presentation_request_dict"])
print(
"\nNote the type defines the protocol present-proof and the message request-presentation\n"
)
print(payload["presentation_request_dict"]["@type"])
elif state == "presentation_received":
print("Presentation Received")
print(
"We will not go into detail on this payload as it is comparable to the presentation_sent we looked at in the earlier cell."
)
print("This is the full payload\n")
print(payload)
else:
print("Paload \n")
print(payload)
verifier_listener = {"topic": "present_proof", "handler": verifier_proof_handler}
listeners.append(verifier_listener)
```
### Role: Prover
```
def prover_proof_handler(payload):
role = payload["role"]
connection_id = payload["connection_id"]
pres_ex_id = payload["presentation_exchange_id"]
state = payload["state"]
print("\n---------------------------------------------------------------------\n")
print("Handle present-proof")
print("Connection ID : ", connection_id)
print("Presentation Exchange ID : ", pres_ex_id)
print("Protocol State : ", state)
print("Agent Role : ", role)
print("Initiator : ", payload["initiator"])
print("\n---------------------------------------------------------------------\n")
if state == "request_received":
presentation_request = payload["presentation_request"]
print("Recieved Presentation Request\n")
print(
"\nRequested Attributes - Note the restrictions. These limit the credentials we could respond with\n"
)
print(presentation_request["requested_attributes"])
elif state == "presentation_sent":
print(
"The Presentation object is a bit overwhelming. Let's look at it in detail\n"
)
elif state == "presentation_acked":
print("Presentation has been acknowledged by the Issuer")
prover_listener = {"topic": "present_proof", "handler": prover_proof_handler}
listeners.append(prover_listener)
```
| github_jupyter |
# Imports
```
import sys
sys.path.append('../')
from datetime import datetime
import pandas as pd
# from src.run_all.main_preprocess import load_data, add_features
from src.utilities.utilities import get_latest_file
pd.set_option('display.max_rows', 500)
pd.set_option('display.max_columns', 500)
pd.set_option('display.width', 1000)
```
# Preprocess
```
%%time
# Load original sources and combine to one DataFrame
# df_dataset_WMO = load_data()
%%time
# Feature engineering to get more features
# df_dataset_WMO_with_features = add_features(df_dataset_WMO)
```
## Optional: Write temporary result
```
# suffix_datetime = datetime.strftime(datetime.now(), format='%Y%m%d%H%M')
# df_dataset_WMO_with_features.to_parquet(f'../../data/df_preprocess_WMO_{suffix_datetime}.parquet.gzip',
# compression='gzip')
```
# Train
## Optional: Load previous dataset
```
## Continue with loaded data from preprocess
# df = df_dataset_WMO_with_features.copy()
# ## HARDCODED
# datapath = '../../data/'
# filename = 'df_preprocess_WMO_202103211137.parquet.gzip'
# df = pd.read_parquet(datapath + filename)
# ## SELECT LAST FILE
datapath = '../data/'
df = get_latest_file(filename_str_contains='df_get_data_WMO_WIJK_HUISHOUDENS_BEVOLKING_HEFFING_202104241837', datapath=datapath, filetype='parquet')
```
## Train model
### Train imports
```
# zorgen voor de juiste modules
import pandas as pd
import numpy as np
from datetime import datetime
import pickle
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import train_test_split, cross_val_score, RepeatedKFold, GridSearchCV, cross_validate, KFold, cross_val_score
from sklearn.linear_model import LogisticRegression, LinearRegression, Ridge, Lasso, LassoCV, ElasticNet, BayesianRidge
from sklearn.preprocessing import OneHotEncoder, StandardScaler
from sklearn.compose import ColumnTransformer, make_column_transformer
from sklearn.pipeline import Pipeline, make_pipeline
from sklearn.svm import SVR
from sklearn.impute import SimpleImputer
from sklearn.neighbors import KNeighborsRegressor
from sklearn.decomposition import PCA
from sklearn.ensemble import RandomForestRegressor
from xgboost import XGBRegressor, XGBClassifier, plot_importance
from src.utilities.transformers import ColumnSelector
# instellingen voor panda weergave aanpassen
pd.set_option('display.max_rows', 500) # alle rijen tonen
pd.set_option('display.max_columns', 500) # alle kolommen tonen
pd.set_option('display.width', 1000) # kolombreedte
pd.set_option("display.precision", 2) # precisie van de kolommen aanpassen
pd.set_option('display.float_format', lambda x: '{:.3f}'.format(x)) # floats output tot 3 decimalen
```
### Settings
```
## Dataframe parameters
# locatie van dataset
DF_LOCATION = 'C:/_NoBackup/Git/__JADS/WMO_execute_group_project/data/df_dataset_WMO.parquet.gzip'
# Location all data
datapath = '../../data/'
# manier van laden dataset. Bijvoorbeeld read_parquet of read_csv
DF_READ = pd.read_parquet
## X & Y parameters
# de kolommen die uit de X dataset moeten worden gehaald. Dat is in ieder geval de y en eventueel nog meer kolommen.
# X_DROP_VALUES = ['wmoclienten', 'eenpersoonshuishoudens', 'huishoudenszonderkinderen', 'huishoudensmetkinderen']
X_DROP_VALUES = ['wmoclienten', 'wmoclientenper1000inwoners', 'bedrijfsmotorvoertuigen',
'perioden', 'popcodea', 'popcodeb', 'popcodec', 'popcoded', 'popcodee', 'popcodef', 'popcodeg', 'popcodeh',
'popcodei', 'popcodej', 'popcodek', 'popcodel', 'popcodem', 'popcoden', 'popcodeo', 'popcodep', 'popcodeq',
'popcoder', 'popnaama', 'popnaamb', 'popnaamc', 'popnaamd', 'popnaame', 'popnaamf', 'popnaamg',
'popnaamh', 'popnaami', 'popnaamj', 'popnaamk', 'popnaaml', 'popnaamm', 'popnaamn', 'popnaamo',
'popnaamp', 'popnaamq', 'popnaamr', 'popkoppelvariabeleregiocode', 'typemaatwerkarrangement',
'gemeentenaam', 'meestvoorkomendepostcode', 'dekkingspercentage',
'gemgestandaardiseerdinkomenvanhuish', 'huishoudenstot110vansociaalminimum',
'huishoudenstot120vansociaalminimum', 'mediaanvermogenvanparticulierehuish',
'popafstandtotopenbaargroen', 'popafstandtotsportterrein', 'popagrarischterreinopp',
'popagrarischterreinperc', 'popagrarischterreinperinwoner', 'popbebouwdterreinopp',
'popbebouwdterreinperc', 'popbebouwdterreinperinwoner', 'popbosenopennatuurlijkterreinopp',
'popbosenopennatuurlijkterreinperc', 'popbosenopennatuurlijkterreinperinwoner', 'popgemeenten',
'poprecreatieterreinopp', 'poprecreatieterreinperc', 'poprecreatieterreinperinwoner',
'popsemibebouwdterreinopp', 'popsemibebouwdterreinperc', 'popsemibebouwdterreinperinwoner',
'popverkeersterreinopp', 'popverkeersterreinperc', 'popverkeersterreinperinwoner']
# de kolom die wordt gebruikt als y value
Y_VALUE = ['wmoclientenper1000inwoners']
# test size voor de train/test split
TEST_SIZE = 0.3
# random state voor de train/test split. Bijvoorbeeld random_state = 42 als vaste seed voor reproduceerbaarheid
RANDOM_STATE = 42
## Pipeline parameters
# strategy en waarde om te vullen bij lege categorische kolommen
NAN_VALUES_CAT_STRATEGY = 'constant'
NAN_VALUES_CAT_VALUES = 'Missing'
# waarden om in te vullen bij lege numerieke kolommen. Bijvoorbeeld mean of median
NAN_VALUES_NUM_STRATEGY = 'mean'
#
#COLS_SELECT = ['aantalinwoners', 'mannen', 'vrouwen', 'k0tot15jaar'
# , 'k15tot25jaar', 'k25tot45jaar', 'k45tot65jaar', 'k65jaarofouder', 'gescheiden'
# , 'verweduwd', 'westerstotaal', 'sterftetotaal', 'gemiddeldehuishoudensgrootte'
# , 'gemiddeldewoningwaarde', 'koopwoningen', 'huurwoningentotaal', 'inbezitwoningcorporatie'
# , 'gemiddeldinkomenperinkomensontvanger', 'k40personenmetlaagsteinkomen', 'k20personenmethoogsteinkomen'
# , 'actieven1575jaar', 'k40huishoudensmetlaagsteinkomen', 'k20huishoudensmethoogsteinkomen'
# , 'huishoudensmeteenlaaginkomen', 'personenpersoortuitkeringaow', 'rucultuurrecreatieoverigediensten'
# , 'personenautosperhuishouden', 'matevanstedelijkheid']
COLS_SELECT = None
## Model parameters
# manier van cross validate in de modellen. Bijvoorbeeld 10 of RepeatedKFold(n_splits=30, n_repeats=5, random_state=1)
CROSS_VALIDATE = 10
# manier van scoren in de modellen
MODEL_SCORING = 'neg_mean_squared_error'
## Scoring parameters
# Deze kunnen we later toevoegen als we meerdere manieren van scoren hebben. Dus niet alleen maar de RSMLE
```
### Functions
```
# functie maken om op basis van de cv scores, het beste RMLSE model te selecteren
def get_best_model_rmsle(cv_scores):
"""
Return best (most conservative) model from cross_validate object.
Uses np.argmax to find bottomright point == largest RMSE
"""
index = np.argmax(np.sqrt(-cv_scores['train_neg_mean_squared_error']))
model = cv_scores['estimator'][index]
rmse = np.sqrt(mean_squared_error(y_test, model.predict(X_test)))
return (rmse)
```
### Load data
```
# ## Done before start of 'Train' chapter
# df = get_latest_file(mypath=datapath)
```
#### Stappen hieronder mogelijk verplaatsten naar prepare stap, later beoordelen
```
# droppen van de rijen waar de y_value leeg is, anders kunnen de modellen er niet mee overweg
df.dropna(
axis=0,
how='any',
thresh=None,
subset=Y_VALUE,
inplace=True
)
# X en y aanmaken
X = df.drop(X_DROP_VALUES, axis=1)
# y = df[Y_VALUE]*100 # 0.01 -> 1.0 percentage
y = df[Y_VALUE] # 0.01 -> 1.0 percentage
# splitsen van X en y in train/test.
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = TEST_SIZE, random_state = RANDOM_STATE)
# splitsen van X_train in categorische en numerieke kolommen, om apart te kunnen transformeren
cat_cols = X_train.select_dtypes(include=['category']).columns
num_cols = X_train.select_dtypes(include=['int64','float64','float32','int32']).columns
```
### Pipelines
```
# pipelines (pl) maken voor imputing, scaling en OneHotEncoding per datatype
# categorie met waarde die is gegeven aan "MISSING" toevoegen
for col in cat_cols:
# need to add category for missings, otherwise error with OneHotEncoding (volgens mij ook met alleen imputing)
X_train[col].cat.add_categories(NAN_VALUES_CAT_VALUES, inplace=True)
categories = [X_train[col].cat.categories for col in cat_cols]
# pipeline voor categorial datatype
pl_ppc_cat = make_pipeline(
SimpleImputer(
missing_values = np.nan
,strategy = NAN_VALUES_CAT_STRATEGY
,fill_value = NAN_VALUES_CAT_VALUES)
,OneHotEncoder(categories=categories)
)
# pipeline voor numeriek datatype
pl_ppc_num = make_pipeline(
ColumnSelector(cols=COLS_SELECT)
,SimpleImputer(
missing_values = np.nan
,strategy = NAN_VALUES_NUM_STRATEGY)
,StandardScaler()
)
# pipelines maken om de preprocessing van de imputing te combineren
pl_ppc_total = make_column_transformer(
(pl_ppc_cat, cat_cols)
,(pl_ppc_num, num_cols)
,remainder = 'drop'
)
```
### Feature importance & feature selection
```
# de X train door bovenstaande pipelines heen halen en opslaan in X_train_prepared
X_train_prepared = pl_ppc_total.fit_transform(X_train)
# de X_train_prepared weer omzetten naar dataframe, inclusief column names
X_train = pd.DataFrame(data=X_train_prepared, columns=[num_cols])
# kiezen van het model waarmee feature importance wordt bepaald, bijvoorbeeld:
# RandomForestRegressor(random_state=42)
# XGBRegressor(n_estimators=100, random_state = 42)
FI_MODEL = XGBRegressor(n_estimators=100, random_state = 42)
FI_MODEL.fit(X_train, y_train)
fi = FI_MODEL.feature_importances_
fi_list = sorted(zip(fi, num_cols), reverse=True)
# met de [:n] kun je het aantal features aanpassen dat getoond moet worden
fi_df_top_n = pd.DataFrame(fi_list[:30])
fi_df_top_n
from matplotlib import pyplot as plt
plt.figure(figsize=(10, 10))
plt.barh(fi_df_top_n[1], fi_df_top_n[0],)
plt.gca().invert_yaxis()
# lasso (cross validate) uitvoeren
feature_selection = LassoCV(cv=10).fit(X_train, y_train)
# lijst aanmaken met coefficienten per feature
feature_selection_coef_list = sorted(zip(np.abs(feature_selection.coef_), num_cols), reverse=True)
df_coef = pd.DataFrame(feature_selection_coef_list, columns=['coef','feature'])
# laat de N features zien met de hoogste coefficienten
df_coef.head(30)
coef_0_cols_df = df_coef[df_coef["coef"] == 0]
coef_0_cols_list = coef_0_cols_df['feature'].values.tolist()
coef_0_cols_list
```
## To do (logistic regression/continious waarden)
```
from sklearn.feature_selection import RFE
from sklearn.linear_model import LogisticRegression
feature_importance_model_logistic_regression = LogisticRegression()
rfe = RFE(feature_importance_model_logistic_regression, 15)
fit = rfe.fit(X_train_prepared, y_train)
print("Num Features: %s" % (fit.n_features_))
print("Selected Features: %s" % (fit.support_))
print("Feature Ranking: %s" % (fit.ranking_))
from sklearn.feature_selection import SelectFromModel
sel_ = SelectFromModel(LogisticRegression(C=1, penalty='l1'))
sel_.fit(X_train_prepared, y_train)
```
## Gridsearch
```
%%time
GRIDSEARCH_MODEL = RandomForestRegressor(random_state=42)
param_grid = [
# try 12 (3×4) combinations of hyperparameters
{'n_estimators': [3, 10, 30], 'max_features': [2, 4, 6, 8]},
# then try 6 (2×3) combinations with bootstrap set as False
{'bootstrap': [False], 'n_estimators': [3, 10], 'max_features': [2, 3, 4]},
]
# train across 5 folds, that's a total of (12+6)*5=90 rounds of training
grid_search = GridSearchCV(GRIDSEARCH_MODEL, param_grid, cv=2,
scoring='neg_mean_squared_error',
return_train_score=True)
grid_search.fit(X_train_prepared, y_train)
grid_search.best_params_
grid_search.best_estimator_
feature_importances = grid_search.best_estimator_.feature_importances_
important_attributes = sorted(zip(feature_importances, num_cols), reverse=True)
important_attributes[:15]
```
| github_jupyter |
# Spectral Analysis for Modal Parameter Linear Estimate
## Setup
### Libraries
Install the `sample` package and its dependencies.
The extras will install dependencies for helper functions such as plots
```
import sys
!$sys.executable -m pip install -qU lim-sample[notebooks,plots]==1.4.0
```
### Generate test audio
We will synthesize a modal-like sound with three modal frequencies (440, 650, 690 Hz) using simple additive synthesis.
Sampling frequency is 44100 Hz and the duration is 2 seconds.
Also, we will add a gaussian noise at -40 dB SNR to mimic a bad recording environment.
```
from matplotlib import pyplot as plt
from librosa.display import waveplot, specshow
from IPython.display import Audio as play
from sample.utils import test_audio
import numpy as np
def resize(w=12, h=6):
plt.gcf().set_size_inches([w, h])
fs = 44100
x = test_audio(fs=fs, noise_db=-40)
waveplot(x, sr=fs, alpha=.5, zorder=100)
plt.grid()
resize()
play(x, rate=fs)
```
## Interface
Using the SAMPLE model is simplified by a scikit-learn-like API
```
from sample import SAMPLE
sample = SAMPLE(
sinusoidal_model__max_n_sines=10,
sinusoidal_model__peak_threshold=-30,
sinusoidal_model__save_intermediate=True
).fit(x)
```
## Sinusoidal Model
SAMPLE is based on Serra's *Spectral Modelling Synthesis* (SMS),
an analysis and synthesis system for musical sounds based
on the decomposition of the sound into a deterministic
sinusoidal and a stochastic component.
The main components of the sinusoidal analysis are the peak detection
and the peak continuation algorithms.
### STFT
The peak detection/continuation algorithm is based on an analysis of the Short-Time Fourier Transform. Zero-phase windowing is employed.
```
stft = np.array([
mx
for mx, _ in sample.sinusoidal_model.intermediate_["stft"]
]).T
specshow(stft, sr=fs, x_axis="time", y_axis="hz");
plt.ylim([0, 2000])
resize()
```
### Peak detection
The peak detection algorithm detects peaks in each STFT frame of the analysed
sound as a local maximum in the magnitude spectrum
```
mx, px = sample.sinusoidal_model.intermediate_["stft"][0]
f = fs * np.arange(mx.size) / sample.sinusoidal_model.w_.size
ploc, pmag, pph = sample.sinusoidal_model.intermediate_["peaks"][0]
ax = plt.subplot(121)
plt.fill_between(f, np.full(mx.shape, -120), mx, alpha=.1)
plt.plot(f, mx)
plt.scatter(ploc * fs / sample.sinusoidal_model.w_.size, pmag, c="C0")
plt.ylim([-60, plt.ylim()[1]])
plt.grid()
plt.title("magnitude")
plt.subplot(122, sharex=ax)
plt.plot(f, px)
plt.scatter(ploc * fs / sample.sinusoidal_model.w_.size, pph)
plt.ylim([np.min(px[f < 2000]), np.max(px[f < 2000])])
plt.grid()
plt.title("phase")
plt.xlim([0, 2000])
resize()
```
### Peak continuation
The peak continuation algorithm organizes the peaks into temporal tracks,
with every track representing the time-varying behaviour of a partial.
For every peak in a trajectory, the instantaneous frequency, magnitude
and phase are stored to allow further manipulation and resynthesis.
The general-purpose SMS method enables recycling of the peak tracks data structures: if one trajectory
becomes inactive, it can be later picked up when a newly detected partial arises.
Our implementation doesn't allow this.
Moreover, two tracks that do not overlap in time but have approximately the same
average frequency can be considered as belonging to the same partial and merged into the same track.
```
from sample import plots
plots.sine_tracking_2d(sample.sinusoidal_model)
resize()
from sample import plots
plots.sine_tracking_3d(sample.sinusoidal_model)
resize(6, 6)
```
## Regression
Partials of a modal impact sound are characterized by exponentially decaying amplitudes.
Our model for model partials is
$$x(t) = m\cdot e^{-2\frac{t}{d}}\cdot \sin{\left(2\pi f t + \phi\right)}$$
The magnitude in decibels is a linear funtion of time
$$m_{dB}(t) = 20\log_{10}{\left(m\cdot e^{-2\frac{t}{d}}\right)} = 20\log_{10}{m} - 40\frac{\log_{10}{e}}{d} \cdot t$$
$$k = - 40\frac{\log_{10}{e}}{d}$$
$$q = 20\log_{10}{m}$$
$$m_{dB}(t) = kt + q$$
We use linear regression to find an initial estimate of the parameters $k$ and $q$ from the magnitude tracks. Then, we refine the estimate by fitting a semi-linear *hinge* function. Amplitude is then doubled to compensate for the fact that we are looking at only half of the spectrum
```
t_x = np.arange(x.size) / fs
for i, ((f, d, a), t) in enumerate(zip(sample.param_matrix_.T, sample.sinusoidal_model.tracks_)):
c = "C{}".format(i)
t_t = (t["start_frame"] + np.arange(t["freq"].size)) * sample.sinusoidal_model.h / sample.sinusoidal_model.fs
plt.plot(t_t, t["mag"] + 6.02, c=c, alpha=.33, linewidth=3) # compensate for spectral halving
plt.plot(t_x, 20*np.log10(a * np.exp(-2*t_x / d)), "--", c=c)
plt.title("fitted curves")
plt.grid()
plt.ylabel("magnitude (dB)")
plt.xlabel("time (s)")
plt.legend(["track", "fitted"])
resize(6, 6)
```
Frequency is simply estimated as the mean frequency of the peak track
# Resynthesize
Let's resynthesize the sound using the estimated parameters (via additive synthesis)
```
x_hat = sample.predict(np.arange(x.size) / fs)
waveplot(x_hat, sr=fs, alpha=.5, zorder=100)
plt.grid()
resize()
play(x_hat, rate=fs)
```
Play back the original sound to compare
```
play(x, rate=fs)
```
Or play both at the same time in stereo
```
from librosa import stft, amplitude_to_db
ax = plt.subplot(211)
x_dual = np.array([x, x_hat])
for l, xi in zip(("original", "resynthesis"), x_dual):
waveplot(xi, sr=fs, alpha=.5, zorder=100, label=l, ax=ax)
plt.grid()
plt.legend()
X_db = amplitude_to_db(np.abs(stft(x)), ref=np.max)
ax = plt.subplot(223, sharex=ax)
specshow(X_db, ax=ax, sr=fs, x_axis="time", y_axis="hz")
ax.set_title("original")
X_hat_db = amplitude_to_db(np.abs(stft(x_hat)), ref=np.max)
ax = plt.subplot(224, sharex=ax, sharey=ax)
specshow(X_hat_db, ax=ax, sr=fs, x_axis="time", y_axis="hz")
ax.set_title("resynthesis")
ax.set_ylim([0, 2000])
resize(12, 12)
play(x_dual, rate=fs)
```
# Manipulate
We can also manipulate the estimated modal parameters to generate a different sound!
As an example we'll tune up the pitches, power-transform the decays and level the amplitudes
```
import copy
sample_mod = copy.deepcopy(sample)
semitones = 1 #@param {type:"slider", min:-12, max:12, step:1}
gamma = 3.14 #@param {type:"slider", min:0.01, max:4, step:0.01}
sample_mod.freqs_ *= 2**(semitones/12)
sample_mod.decays_ **= gamma
sample_mod.amps_ = np.full(
sample.amps_.size,
1/sample.amps_.size
)
x_mod = sample_mod.predict(np.arange(x.size) / fs)
waveplot(x_mod, sr=fs, alpha=.5, zorder=100)
plt.grid()
resize()
play(x_mod, rate=fs)
```
| github_jupyter |
```
import numpy as np
import pandas as pd
%matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sb
sb.set_style('whitegrid')
import requests
import json
import re
from bs4 import BeautifulSoup
import string
import nltk
import networkx as nx
```
## Loading data
Load the data from disk into memory.
```
with open('potus_wiki_bios_cleaned.json','r') as f:
bios = json.load(f)
```
Confirm there are 44 presidents (shaking fist at [Grover Cleveland](https://en.wikipedia.org/wiki/Grover_Cleveland)) in the dictionary.
```
print("There are {0} biographies of presidents.".format(len(bios)))
```
What's an example of a single biography? We access the dictionary by passing the key (President's name), which returns the value (the text of the biography).
```
example = bios['Grover Cleveland']
print(example)
```
Get some metadata about the U.S. Presidents.
```
presidents_df = pd.DataFrame(requests.get('https://raw.githubusercontent.com/hitch17/sample-data/master/presidents.json').json())
presidents_df = presidents_df.set_index('president')
presidents_df['wikibio words'] = pd.Series({bio_name:len(bio_text) for bio_name,bio_text in bios.items()})
presidents_df.head()
```
A really basic exploratory scatterplot for the number of words in each President's biography compared to their POTUS index.
```
presidents_df.plot.scatter(x='number',y='wikibio words')
```
## TF-IDF
We can create a document-term matrix where the rows are our 44 presidential biographies, the columns are the terms (words), and the values in the cells are the word counts: the number of times that document contains that word. This is the "term frequency" (TF) part of TF-IDF.
The IDF part of TF-IDF is the "inverse document frequency". The intuition is that words that occur frequency within a single document but are infrequent across the corpus of documents should receiving a higher weighting: these words have greater relative meaning. Conversely, words that are frequently used across documents are down-weighted.
The image below has documents as columns and terms as rows.
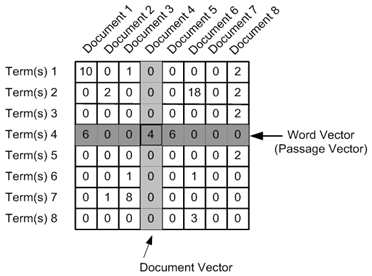
```
# Import the libraries from scikit-learn
from sklearn.feature_extraction.text import CountVectorizer,TfidfTransformer
count_vect = CountVectorizer()
# Compute the word counts -- it expects a big string, so join our cleaned words back together
bio_counts = count_vect.fit_transform([' '.join(bio) for bio in bios.values()])
# Compute the TF-IDF for the word counts from each biography
bio_tfidf = TfidfTransformer().fit_transform(bio_counts)
# Convert from sparse to dense array representation
bio_tfidf_dense = bio_tfidf.todense()
```
## Make a text similarity network
Once we have the TFIDF scores for every word in each president's biography, we can make a text similarity network. Multiplying the document by term matrix by its transpose should return the [cosine similarities](https://en.wikipedia.org/wiki/Cosine_similarity) between documents. We can also import [`cosine_similarity`](http://scikit-learn.org/stable/modules/generated/sklearn.metrics.pairwise.cosine_similarity.html) from scikit-learn if you don't believe me (I didn't believe me either). Cosine similarity values closer to 1 indicate these documents' words have more similar TFIDF scores and values closer to 0 indicate these documents' words are more dissimilar.
The goal here is to create a network where nodes are presidents and edges are weighted similarity scores. All text documents will have some minimal similarity, so we can threshold the similarity scores to only those similarities in the top 10% for each president.
```
# Compute cosine similarity
pres_pres_df = pd.DataFrame(bio_tfidf_dense*bio_tfidf_dense.T)
# If you don't believe me that cosine similiarty is the document-term matrix times its transpose
from sklearn.metrics.pairwise import cosine_similarity
pres_pres_df = pd.DataFrame(cosine_similarity(bio_tfidf_dense))
# Filter for edges in the 90th percentile or greater
pres_pres_filtered_df = pres_pres_df[pres_pres_df >= pres_pres_df.quantile(.9)]
# Reshape and filter data
edgelist_df = pres_pres_filtered_df.stack().reset_index()
edgelist_df = edgelist_df[(edgelist_df[0] != 0) & (edgelist_df['level_0'] != edgelist_df['level_1'])]
# Rename and replace data
edgelist_df.rename(columns={'level_0':'from','level_1':'to',0:'weight'},inplace=True)
edgelist_df.replace(dict(enumerate(bios.keys())),inplace=True)
# Inspect
edgelist_df.head()
```
We read this pandas edgelist into networkx using `from_pandas_edgelist`, report out some basic descriptives about the network, and write the graph object to file in case we want to visualize it in a dedicated network visualization package like [Gephi](https://gephi.org/).
```
# Convert from edgelist to a graph object
g = nx.from_pandas_edgelist(edgelist_df,source='from',target='to',edge_attr=['weight'])
# Report out basic descriptives
print("There are {0:,} nodes and {1:,} edges in the network.".format(g.number_of_nodes(),g.number_of_edges()))
# Write graph object to disk for visualization
nx.write_gexf(g,'bio_similarity.gexf')
```
Since this is a small and sparse network, we can try to use Matplotlib to visualize it instead. I would only use the `nx.draw` functionality for small networks like this one.
```
# Plot the nodes as a spring layout
#g_pos = nx.layout.fruchterman_reingold_layout(g, k = 5, iterations=10000)
g_pos = nx.layout.kamada_kawai_layout(g)
# Draw the graph
f,ax = plt.subplots(1,1,figsize=(10,10))
nx.draw(G = g,
ax = ax,
pos = g_pos,
with_labels = True,
node_size = [dc*(len(g) - 1)*100 for dc in nx.degree_centrality(g).values()],
font_size = 10,
font_weight = 'bold',
width = [d['weight']*10 for i,j,d in g.edges(data=True)],
node_color = 'tomato',
edge_color = 'grey'
)
```
## Case study: Text similarity network of the S&P 500 companies
**Step 1**: Load and preprocess the content of the articles.
```
# Load the data
with open('sp500_wiki_articles.json','r') as f:
sp500_articles = json.load(f)
# Bring in the text_preprocessor we wrote from Day 4, Lecture 1
def text_preprocessor(text):
"""Takes a large string (document) and returns a list of cleaned tokens"""
tokens = nltk.wordpunct_tokenize(text)
clean_tokens = []
for t in tokens:
if t.lower() not in all_stopwords and len(t) > 2:
clean_tokens.append(lemmatizer(t.lower()))
return clean_tokens
# Clean each article
cleaned_sp500 = {}
for name,text in sp500_articles.items():
cleaned_sp500[name] = text_preprocessor(text)
# Save to disk
with open('sp500_wiki_articles_cleaned.json','w') as f:
json.dump(cleaned_sp500,f)
```
**Step 2**: Compute the TFIDF matrix for the S&P 500 companies.
```
# Compute the word counts
sp500_counts =
# Compute the TF-IDF for the word counts from each biography
sp500_tfidf =
# Convert from sparse to dense array representation
sp500_tfidf_dense =
```
**Step 3**: Compute the cosine similarities.
```
# Compute cosine similarity
company_company_df =
# Filter for edges in the 90th percentile or greater
company_company_filtered_df =
# Reshape and filter data
sp500_edgelist_df =
sp500_edgelist_df =
# Rename and replace data
sp500_edgelist_df.rename(columns={'level_0':'from','level_1':'to',0:'weight'},inplace=True)
sp500_edgelist_df.replace(dict(enumerate(sp500_articles.keys())),inplace=True)
# Inspect
sp500_edgelist_df.head()
```
**Step 4**: Visualize the resulting network.
## Word2Vec
We used TF-IDF vectors of documents and cosine similarities between these document vectors as a way of representing the similarity in the networks above. However, TF-IDF score are simply (normalized) word frequencies: they do not capture semantic information. A vector space model like the popular Word2Vec represents each token (word) in a high-dimensional (here we'll use 100-dimensions) space that is trained from some (ideally) large corpus of documents. Ideally, tokens that are used in similar contexts are placed into similar locations in this high-dimensional space. Once we have vectorized words into this space, we're able to efficiently compute do a variety of other operations such as compute similarities between words or do transformations that can find analogies.
I lack the expertise and we lack the time to get into the math behind these methods, but here are some helpful tutorials I've found:
* [Word embeddings: exploration, explanation, and exploitation ](https://towardsdatascience.com/word-embeddings-exploration-explanation-and-exploitation-with-code-in-python-5dac99d5d795)
* [Learning Word Embedding](https://lilianweng.github.io/lil-log/2017/10/15/learning-word-embedding.html)
* [On word embeddings](http://ruder.io/word-embeddings-1/)
* [TensorFlow - Vector Representations of Words](https://www.tensorflow.org/tutorials/representation/word2vec)
We'll use the 44 Presidential biographies as a small and specific corpus. We start by training a `bios_model` from the list of biographies using hyperparamaters for the number of dimensions (size), the number of surrounding words to use as training (window), and the minimum number of times a word has to occur to be included in the model (min_count).
```
from gensim.models import Word2Vec
bios_model = Word2Vec(bios.values(),size=100,window=10,min_count=8)
```
Each word in the vocabulary exists as a N-dimensional vector, where N is the "size" hyper-parameter set in the model. The "congress" token in located at this position in the 100-dimensional space we trained in `bios_model`.
```
bios_model.wv['congress']
bios_model.wv.most_similar('congress')
bios_model.wv.most_similar('court')
bios_model.wv.most_similar('war')
bios_model.wv.most_similar('election')
```
There's a `doesnt_match` method that predicts which word in a list doesn't match the other word senses in the list. Sometime the results are predictable/trivial.
```
bios_model.wv.doesnt_match(['democrat','republican','whig','panama'])
```
Other times the results are unexpected/interesting.
```
bios_model.wv.doesnt_match(['canada','mexico','cuba','japan','france'])
```
One of the most powerful implications of having these vectorized embeddings of word meanings is the ability to do operations similar arithmetic that recover or reveal interesting semantic meanings. The classic example is `Man:Woman::King:Queen`:

What are some examples of these vector similarities from our trained model?
`republican - slavery = democrat - X`
`-(republican - slavery) + democrat = X`
`slavery + democrat - republican = X`
```
bios_model.wv.most_similar(positive=['democrat','slavery'],negative=['republican'])
bios_model.wv.most_similar(positive=['republican','labor'],negative=['democrat'])
```
Finally, you can use the `similarity` method to return the similarity between two terms. In our trained model, "britain" and "france" are more similar to each other than "mexico" and "canada".
```
bios_model.wv.similarity('republican','democrat')
bios_model.wv.similarity('mexico','canada')
bios_model.wv.similarity('britain','france')
```
## Case study: S&P500 company Word2Vec model
**Step 1**: Open the "sp500_wiki_articles_cleaned.json" you previous saved of the cleaned S&P500 company article content or use a text preprocessor on "sp500_wiki_articles.json" to generate a dictionary of cleaned article content. Train a `sp500_model` using the `Word2Vec` model on the values of the cleaned company article content. You can use default hyperparameters for size, window, and min_count, or experiment with alternative values.
**Step 2**: Using the `most_similar` method, explore some similarities this model has learned for salient tokens about companies (*e.g.*, "board", "controversy", "executive", "investigation"). Use the positive and negative options to explore different analogies. Using the `doesnt_match` method, experiment with word combinations to discover predictable and unexpected exceptions. Using the `similarity` method, identify interesting similarity scores.
## Dimensionality reduction
Material from this segment is adapted from [Jake Vanderplas](http://jakevdp.github.io/)'s ["Python Data Science Handbook" notebooks](https://github.com/jakevdp/PythonDataScienceHandbook/tree/master/notebooks) and [Kevyn Collins-Thompson](http://www-personal.umich.edu/~kevynct/)'s ["Applied Machine Learning in Python"](https://www.coursera.org/learn/python-machine-learning/lecture/XIt7x/introduction) module on Coursera.
In the TF-IDF, we have over 17,000 dimensions (corresponding to the unique tokens) for each of the 44 presidential biographies. This data is sparse and large, which makes it hard to visualize. Ideally we'd only have two dimensions of data for a task like visualization.
Dimensionality reduction encompasses a set of methods like principal component analysis, multidimensional scaling, and more advanced "[manifold learning](http://scikit-learn.org/stable/modules/manifold.html#introduction)" that reduces high-dimensional data down to fewer dimensions. For the purposes of visualization, we typically want 2 dimensions. These methods use a variety of different assumptions and modeling approaches. If you want to understand the differences between them, you'll likely need to find a graduate-level machine learning course.
Let's compare what each of these do on our presidential TF-IDF: the goal here is to understand there are different methods for dimensionality reduction and each generates different new components and/or clusters that you'll need to interpret.
```
print(bio_tfidf_dense.shape)
bio_tfidf_dense
```
Principal component analysis (PCA) is probably one of the most widely-used and efficient methods for dimensionality reduction.
```
# Step 1: Choose a class of models
from sklearn.decomposition import PCA
# Step 2: Instantiate the model
pca = PCA(n_components=2)
# Step 3: Arrange the data into features matrices
# Already done
# Step 4: Fit the model to the data
pca.fit(bio_tfidf_dense)
# Step 5: Evaluate the model
X_pca = pca.transform(bio_tfidf_dense)
# Visualize
f,ax = plt.subplots(1,1,figsize=(10,10))
ax.scatter(X_pca[:,0],X_pca[:,1])
ax.set_title('PCA')
for i,txt in enumerate(bios.keys()):
if txt == 'Barack Obama':
ax.annotate(txt,(X_pca[i,0],X_pca[i,1]),color='blue',fontweight='bold')
elif txt == 'Donald Trump':
ax.annotate(txt,(X_pca[i,0],X_pca[i,1]),color='red',fontweight='bold')
else:
ax.annotate(txt,(X_pca[i,0],X_pca[i,1]))
```
Multi-dimensional scaling is another common technique in the social sciences.
```
# Step 1: Choose your model class(es)
from sklearn.manifold import MDS
# Step 2: Instantiate your model class(es)
mds = MDS(n_components=2,metric=False,n_jobs=-1)
# Step 3: Arrange data into features matrices
# Done!
# Step 4: Fit the data and transform
X_mds = mds.fit_transform(bio_tfidf_dense)
# Plot the data
f,ax = plt.subplots(1,1,figsize=(10,10))
ax.scatter(X_mds[:,0],X_mds[:,1])
ax.set_title('Multi-Dimensional Scaling')
for i,txt in enumerate(bios.keys()):
if txt == 'Barack Obama':
ax.annotate(txt,(X_mds[i,0],X_mds[i,1]),color='blue',fontweight='bold')
elif txt == 'Donald Trump':
ax.annotate(txt,(X_mds[i,0],X_mds[i,1]),color='red',fontweight='bold')
else:
ax.annotate(txt,(X_mds[i,0],X_mds[i,1]))
```
[Isomap](https://en.wikipedia.org/wiki/Isomap) is an extension of MDS.
```
# Step 1: Choose your model class(es)
from sklearn.manifold import Isomap
# Step 2: Instantiate your model class(es)
iso = Isomap(n_neighbors = 5, n_components = 2)
# Step 3: Arrange data into features matrices
# Done!
# Step 4: Fit the data and transform
X_iso = iso.fit_transform(bio_tfidf_dense)
# Plot the data
f,ax = plt.subplots(1,1,figsize=(10,10))
ax.scatter(X_iso[:,0],X_iso[:,1])
ax.set_title('IsoMap')
for i,txt in enumerate(bios.keys()):
if txt == 'Barack Obama':
ax.annotate(txt,(X_iso[i,0],X_iso[i,1]),color='blue',fontweight='bold')
elif txt == 'Donald Trump':
ax.annotate(txt,(X_iso[i,0],X_iso[i,1]),color='red',fontweight='bold')
else:
ax.annotate(txt,(X_iso[i,0],X_iso[i,1]))
```
[Spectral embedding](https://en.wikipedia.org/wiki/Spectral_clustering) does interesting things to the eigenvectors of a similarity matrix.
```
# Step 1: Choose your model class(es)
from sklearn.manifold import SpectralEmbedding
# Step 2: Instantiate your model class(es)
se = SpectralEmbedding(n_components = 2)
# Step 3: Arrange data into features matrices
# Done!
# Step 4: Fit the data and transform
X_se = se.fit_transform(bio_tfidf_dense)
# Plot the data
f,ax = plt.subplots(1,1,figsize=(9,6))
ax.scatter(X_se[:,0],X_se[:,1])
ax.set_title('Spectral Embedding')
for i,txt in enumerate(bios.keys()):
if txt == 'Barack Obama':
ax.annotate(txt,(X_se[i,0],X_se[i,1]),color='blue',fontweight='bold')
elif txt == 'Donald Trump':
ax.annotate(txt,(X_se[i,0],X_se[i,1]),color='red',fontweight='bold')
else:
ax.annotate(txt,(X_se[i,0],X_se[i,1]))
```
Locally Linear Embedding is yet another dimensionality reduction method, but not my favorite to date given performance (meaningful clusters as output) and cost (expensive to compute).
```
# Step 1: Choose your model class(es)
from sklearn.manifold import LocallyLinearEmbedding
# Step 2: Instantiate your model class(es)
lle = LocallyLinearEmbedding(n_components = 2,n_jobs=-1)
# Step 3: Arrange data into features matrices
# Done!
# Step 4: Fit the data and transform
X_lle = lle.fit_transform(bio_tfidf_dense)
# Plot the data
f,ax = plt.subplots(1,1,figsize=(9,6))
ax.scatter(X_lle[:,0],X_lle[:,1])
ax.set_title('Locally Linear Embedding')
for i,txt in enumerate(bios.keys()):
if txt == 'Barack Obama':
ax.annotate(txt,(X_lle[i,0],X_lle[i,1]),color='blue',fontweight='bold')
elif txt == 'Donald Trump':
ax.annotate(txt,(X_lle[i,0],X_lle[i,1]),color='red',fontweight='bold')
else:
ax.annotate(txt,(X_lle[i,0],X_lle[i,1]))
```
[t-Distributed Stochastic Neighbor Embedding](https://lvdmaaten.github.io/tsne/) (t-SNE) is ubiquitous for visualizing word or document embeddings. It can be expensive to run, but it does a great job recovering clusters. There are some hyper-parameters, particularly "perplexity" that you'll need to tune to get things to look interesting.
Wattenberg, Viégas, and Johnson have an [outstanding interactive tool](https://distill.pub/2016/misread-tsne/) visualizing how t-SNE's different parameters influence the layout as well as good advice on how to make the best of it.
```
# Step 1: Choose your model class(es)
from sklearn.manifold import TSNE
# Step 2: Instantiate your model class(es)
tsne = TSNE(n_components = 2, init='pca', random_state=42, perplexity=11)
# Step 3: Arrange data into features matrices
# Done!
# Step 4: Fit the data and transform
X_tsne = tsne.fit_transform(bio_tfidf_dense)
# Plot the data
f,ax = plt.subplots(1,1,figsize=(10,10))
ax.scatter(X_tsne[:,0],X_tsne[:,1])
ax.set_title('t-SNE')
for i,txt in enumerate(bios.keys()):
if txt == 'Barack Obama':
ax.annotate(txt,(X_tsne[i,0],X_tsne[i,1]),color='blue',fontweight='bold')
elif txt == 'Donald Trump':
ax.annotate(txt,(X_tsne[i,0],X_tsne[i,1]),color='red',fontweight='bold')
else:
ax.annotate(txt,(X_tsne[i,0],X_tsne[i,1]))
```
[Uniform Maniford Approximation and Projection (UMAP)](https://github.com/lmcinnes/umap) is a new and particularly fast dimensionality reduction method with some comparatively great documentation. Unfortunately, UMAP is so new that it hasn't been translated into scikit-learn yet, so you'll need to install it separately from the terminal:
`conda install -c conda-forge umap-learn`
```
# Step 1: Choose your model class(es)
from umap import UMAP
# Step 2: Instantiate your model class(es)
umap_ = UMAP(n_components=2, n_neighbors=10, random_state=42)
# Step 3: Arrange data into features matrices
# Done!
# Step 4: Fit the data and transform
X_umap = umap_.fit_transform(bio_tfidf_dense)
# Plot the data
f,ax = plt.subplots(1,1,figsize=(10,10))
ax.scatter(X_umap[:,0],X_umap[:,1])
ax.set_title('UMAP')
for i,txt in enumerate(bios.keys()):
if txt == 'Barack Obama':
ax.annotate(txt,(X_umap[i,0],X_umap[i,1]),color='blue',fontweight='bold')
elif txt == 'Donald Trump':
ax.annotate(txt,(X_umap[i,0],X_umap[i,1]),color='red',fontweight='bold')
else:
ax.annotate(txt,(X_umap[i,0],X_umap[i,1]))
```
## Case study: S&P500 company clusters
**Step 1**: Using the `sp500_tfidf_dense` array/DataFrame, experiment with different dimensionality reduction tools we covered above. Visualize and inspect the distribution of S&P500 companies for interesting dimensions (do X and Y dimensions in this reduced data capture anything meaningful?) or clusters (do companies clusters together as we'd expect?).
## Visualizing word embeddings
Using the `bio_counts`, we can find the top-N most frequent words and save them as `top_words`.
```
top_words = pd.DataFrame(bio_counts.todense().sum(0).T,
index=count_vect.get_feature_names())[0]
top_words = top_words.sort_values(0,ascending=False).head(1000).index.tolist()
```
For each word in `top_words`, we get its vector from `bios_model` and add it to the `top_word_vectors` list and cast this list back to a numpy array.
```
top_word_vectors = []
for word in top_words:
try:
vector = bios_model.wv[word]
top_word_vectors.append(vector)
except KeyError:
pass
top_word_vectors = np.array(top_word_vectors)
```
We can then use the dimensionality tools we just covered in the previous section to visualize the word similarities. PCA is fast but rarely does a great job with this extremely high-dimensional and sparse data: it's a cloud of points with no discernable structure.
```
# Step 1: Choose your model class(es)
# from sklearn.decomposition import PCA
# Step 2: Instantiate the model
pca = PCA(n_components=2)
# Step 3: Arrange data into features matrices
X_w2v = top_word_vectors
# Step 4: Fit the data and transform
X_w2v_pca = pca.fit_transform(X_w2v)
# Plot the data
f,ax = plt.subplots(1,1,figsize=(10,10))
ax.scatter(X_w2v_pca[:,0],X_w2v_pca[:,1],s=3)
ax.set_title('PCA')
for i,txt in enumerate(top_words):
if i%10 == 0:
ax.annotate(txt,(X_w2v_pca[i,0],X_w2v_pca[i,1]))
f.savefig('term_pca.pdf')
```
t-SNE was more-or-less engineered for precisely the task of visualizing word embeddings. It likely takes on the order of a minute or more for t-SNE to reduce the `top_words` embeddings to only two dimensions. Assuming our perplexity and other t-SNE hyperparameters are well-behaved, there should be relatively easy-to-discern clusters of words with similar meanings. You can also open the "term_sne.pdf" file and zoome to inspect.
```
# Step 1: Choose your model class(es)
from sklearn.manifold import TSNE
# Step 2: Instantiate your model class(es)
tsne = TSNE(n_components = 2, init='pca', random_state=42, perplexity=25)
# Step 3: Arrange data into features matrices
X_w2v = top_word_vectors
# Step 4: Fit the data and transform
X_w2v_tsne = tsne.fit_transform(X_w2v)
# Plot the data
f,ax = plt.subplots(1,1,figsize=(10,10))
ax.scatter(X_w2v_tsne[:,0],X_w2v_tsne[:,1],s=3)
ax.set_title('t-SNE')
for i,txt in enumerate(top_words):
if i%10 == 0:
ax.annotate(txt,(X_w2v_tsne[i,0],X_w2v_tsne[i,1]))
f.savefig('term_tsne.pdf')
```
UMAP is faster and I think better, but you'll need to make sure this is installed on your system since it doesn't come with scikit-learn or Anaconda by default. Words like "nominee" and "campaign" or the names of the months cluster clearly together apart from the rest.
```
# Step 1: Choose your model class(es)
from umap import UMAP
# Step 2: Instantiate your model class(es)
umap_ = UMAP(n_components=2, n_neighbors=5, random_state=42)
# Step 3: Arrange data into features matrices
# Done!
# Step 4: Fit the data and transform
X_w2v_umap = umap_.fit_transform(X_w2v)
# Plot the data
f,ax = plt.subplots(1,1,figsize=(10,10))
ax.scatter(X_w2v_umap[:,0],X_w2v_umap[:,1],s=3)
ax.set_title('UMAP')
for i,txt in enumerate(top_words):
if i%10 == 0:
ax.annotate(txt,(X_w2v_umap[i,0],X_w2v_umap[i,1]))
f.savefig('term_umap.pdf')
```
## Case study: Visualizing word embeddings for S&P500 company articles
**Step 1**: Compute the word vectors for the top 1000(ish) terms in the S&P500 word counts from your `sp500_model`.
**Step 2**: Reduce the dimensionality of these top word vectors using PCA, t-SNE, or (if you've installed it) UMAP and visualize the results. What meaningful or surprising clusters do you discover?
| github_jupyter |
# Hello Vectors
exploring word vectors.
In natural language processing, we represent each word as a vector consisting of numbers.
The vector encodes the meaning of the word. These numbers (or weights) for each word are learned using various machine
learning models, which we will explore in more detail later in this specialization. Rather than make you code the
machine learning models from scratch, we will show you how to use them. In the real world, you can always load the
trained word vectors, and you will almost never have to train them from scratch. In this assignment, you will:
- Predict analogies between words.
- Use PCA to reduce the dimensionality of the word embeddings and plot them in two dimensions.
- Compare word embeddings by using a similarity measure (the cosine similarity).
- Understand how these vector space models work.
## 1.0 Predict the Countries from Capitals
In the lectures, we have illustrated the word analogies
by finding the capital of a country from the country.
We have changed the problem a bit in this part of the assignment. You are asked to predict the **countries**
that corresponds to some **capitals**.
You are playing trivia against some second grader who just took their geography test and knows all the capitals by heart.
Thanks to NLP, you will be able to answer the questions properly. In other words, you will write a program that can give
you the country by its capital. That way you are pretty sure you will win the trivia game. We will start by exploring the data set.
<img src = 'map.jpg' width="width" height="height" style="width:467px;height:300px;"/>
### 1.1 Importing the data
As usual, you start by importing some essential Python libraries and then load the dataset.
The dataset will be loaded as a [Pandas DataFrame](https://pandas.pydata.org/pandas-docs/stable/getting_started/dsintro.html),
which is very a common method in data science.
This may take a few minutes because of the large size of the data.
```
# Run this cell to import packages.
import pickle
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from utils import get_vectors
data = pd.read_csv('capitals.txt', delimiter=' ')
data.columns = ['city1', 'country1', 'city2', 'country2']
# print first five elements in the DataFrame
data.head(5)
```
***
### To Run This Code On Your Own Machine:
Note that because the original google news word embedding dataset is about 3.64 gigabytes,
the workspace is not able to handle the full file set. So we've downloaded the full dataset,
extracted a sample of the words that we're going to analyze in this assignment, and saved
it in a pickle file called `word_embeddings_capitals.p`
If you want to download the full dataset on your own and choose your own set of word embeddings,
please see the instructions and some helper code.
- Download the dataset from this [page](https://code.google.com/archive/p/word2vec/).
- Search in the page for 'GoogleNews-vectors-negative300.bin.gz' and click the link to download.
Copy-paste the code below and run it on your local machine after downloading
the dataset to the same directory as the notebook.
```python
import nltk
from gensim.models import KeyedVectors
embeddings = KeyedVectors.load_word2vec_format('./GoogleNews-vectors-negative300.bin', binary = True)
f = open('capitals.txt', 'r').read()
set_words = set(nltk.word_tokenize(f))
select_words = words = ['king', 'queen', 'oil', 'gas', 'happy', 'sad', 'city', 'town', 'village', 'country', 'continent', 'petroleum', 'joyful']
for w in select_words:
set_words.add(w)
def get_word_embeddings(embeddings):
word_embeddings = {}
for word in embeddings.vocab:
if word in set_words:
word_embeddings[word] = embeddings[word]
return word_embeddings
# Testing your function
word_embeddings = get_word_embeddings(embeddings)
print(len(word_embeddings))
pickle.dump( word_embeddings, open( "word_embeddings_subset.p", "wb" ) )
```
***
Now we will load the word embeddings as a [Python dictionary](https://docs.python.org/3/tutorial/datastructures.html#dictionaries).
As stated, these have already been obtained through a machine learning algorithm.
```
word_embeddings = pickle.load(open("word_embeddings_subset.p", "rb"))
len(word_embeddings) # there should be 243 words that will be used in this assignment
```
Each of the word embedding is a 300-dimensional vector.
```
print("dimension: {}".format(word_embeddings['Spain'].shape[0]))
```
### Predict relationships among words
Now you will write a function that will use the word embeddings to predict relationships among words.
* The function will take as input three words.
* The first two are related to each other.
* It will predict a 4th word which is related to the third word in a similar manner as the two first words are related to each other.
* As an example, "Athens is to Greece as Bangkok is to ______"?
* You will write a program that is capable of finding the fourth word.
* We will give you a hint to show you how to compute this.
A similar analogy would be the following:
<img src = 'vectors.jpg' width="width" height="height" style="width:467px;height:200px;"/>
You will implement a function that can tell you the capital of a country.
You should use the same methodology shown in the figure above. To do this,
compute you'll first compute cosine similarity metric or the Euclidean distance.
### 1.2 Cosine Similarity
The cosine similarity function is:
$$\cos (\theta)=\frac{\mathbf{A} \cdot \mathbf{B}}{\|\mathbf{A}\|\|\mathbf{B}\|}=\frac{\sum_{i=1}^{n} A_{i} B_{i}}{\sqrt{\sum_{i=1}^{n} A_{i}^{2}} \sqrt{\sum_{i=1}^{n} B_{i}^{2}}}\tag{1}$$
$A$ and $B$ represent the word vectors and $A_i$ or $B_i$ represent index i of that vector.
& Note that if A and B are identical, you will get $cos(\theta) = 1$.
* Otherwise, if they are the total opposite, meaning, $A= -B$, then you would get $cos(\theta) = -1$.
* If you get $cos(\theta) =0$, that means that they are orthogonal (or perpendicular).
* Numbers between 0 and 1 indicate a similarity score.
* Numbers between -1-0 indicate a dissimilarity score.
**Instructions**: Implement a function that takes in two word vectors and computes the cosine distance.
<details>
<summary>
<font size="3" color="darkgreen"><b>Hints</b></font>
</summary>
<p>
<ul>
<li> Python's<a href="https://docs.scipy.org/doc/numpy/reference/" > NumPy library </a> adds support for linear algebra operations (e.g., dot product, vector norm ...).</li>
<li>Use <a href="https://docs.scipy.org/doc/numpy/reference/generated/numpy.dot.html" > numpy.dot </a>.</li>
<li>Use <a href="https://docs.scipy.org/doc/numpy/reference/generated/numpy.linalg.norm.html">numpy.linalg.norm </a>.</li>
</ul>
</p>
```
# UNQ_C1 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
def cosine_similarity(A, B):
'''
Input:
A: a numpy array which corresponds to a word vector
B: A numpy array which corresponds to a word vector
Output:
cos: numerical number representing the cosine similarity between A and B.
'''
### START CODE HERE (REPLACE INSTANCES OF 'None' with your code) ###
dot = A.dot(B)
norma = np.linalg.norm(A)
normb = np.linalg.norm(B)
cos = dot/(norma*normb)
### END CODE HERE ###
return cos
# feel free to try different words
king = word_embeddings['king']
queen = word_embeddings['queen']
cosine_similarity(king, queen)
```
**Expected Output**:
$\approx$ 0.6510956
### 1.3 Euclidean distance
You will now implement a function that computes the similarity between two vectors using the Euclidean distance.
Euclidean distance is defined as:
$$ \begin{aligned} d(\mathbf{A}, \mathbf{B})=d(\mathbf{B}, \mathbf{A}) &=\sqrt{\left(A_{1}-B_{1}\right)^{2}+\left(A_{2}-B_{2}\right)^{2}+\cdots+\left(A_{n}-B_{n}\right)^{2}} \\ &=\sqrt{\sum_{i=1}^{n}\left(A_{i}-B_{i}\right)^{2}} \end{aligned}$$
* $n$ is the number of elements in the vector
* $A$ and $B$ are the corresponding word vectors.
* The more similar the words, the more likely the Euclidean distance will be close to 0.
**Instructions**: Write a function that computes the Euclidean distance between two vectors.
<details>
<summary>
<font size="3" color="darkgreen"><b>Hints</b></font>
</summary>
<p>
<ul>
<li>Use <a href="https://docs.scipy.org/doc/numpy/reference/generated/numpy.linalg.norm.html" > numpy.linalg.norm </a>.</li>
</ul>
</p>
```
# UNQ_C2 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
def euclidean(A, B):
"""
Input:
A: a numpy array which corresponds to a word vector
B: A numpy array which corresponds to a word vector
Output:
d: numerical number representing the Euclidean distance between A and B.
"""
### START CODE HERE (REPLACE INSTANCES OF 'None' with your code) ###
# euclidean distance
d = np.linalg.norm(A-B)
### END CODE HERE ###
return d
# Test your function
euclidean(king, queen)
```
**Expected Output:**
2.4796925
### 1.4 Finding the country of each capital
Now, you will use the previous functions to compute similarities between vectors,
and use these to find the capital cities of countries. You will write a function that
takes in three words, and the embeddings dictionary. Your task is to find the
capital cities. For example, given the following words:
- 1: Athens 2: Greece 3: Baghdad,
your task is to predict the country 4: Iraq.
**Instructions**:
1. To predict the capital you might want to look at the *King - Man + Woman = Queen* example above, and implement that scheme into a mathematical function, using the word embeddings and a similarity function.
2. Iterate over the embeddings dictionary and compute the cosine similarity score between your vector and the current word embedding.
3. You should add a check to make sure that the word you return is not any of the words that you fed into your function. Return the one with the highest score.
```
# UNQ_C3 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
def get_country(city1, country1, city2, embeddings):
"""
Input:
city1: a string (the capital city of country1)
country1: a string (the country of capital1)
city2: a string (the capital city of country2)
embeddings: a dictionary where the keys are words and values are their embeddings
Output:
countries: a dictionary with the most likely country and its similarity score
"""
### START CODE HERE (REPLACE INSTANCES OF 'None' with your code) ###
# store the city1, country 1, and city 2 in a set called group
group = set((city1, country1, city2))
# get embeddings of city 1
city1_emb = embeddings[city1]
# get embedding of country 1
country1_emb = embeddings[country1]
# get embedding of city 2
city2_emb = embeddings[city2]
# get embedding of country 2 (it's a combination of the embeddings of country 1, city 1 and city 2)
# Remember: King - Man + Woman = Queen
vec = country1_emb-city1_emb+city2_emb
# Initialize the similarity to -1 (it will be replaced by a similarities that are closer to +1)
similarity = -1
# initialize country to an empty string
country = ''
# loop through all words in the embeddings dictionary
for word in embeddings.keys():
# first check that the word is not already in the 'group'
if word not in group:
# get the word embedding
word_emb = embeddings[word]
# calculate cosine similarity between embedding of country 2 and the word in the embeddings dictionary
cur_similarity = cosine_similarity(vec, word_emb)
# if the cosine similarity is more similar than the previously best similarity...
if cur_similarity > similarity:
# update the similarity to the new, better similarity
similarity = cur_similarity
# store the country as a tuple, which contains the word and the similarity
country = (word, similarity)
### END CODE HERE ###
return country
# Testing your function, note to make it more robust you can return the 5 most similar words.
get_country('Athens', 'Greece', 'Cairo', word_embeddings)
```
**Expected Output:**
('Egypt', 0.7626821)
### 1.5 Model Accuracy
Now you will test your new function on the dataset and check the accuracy of the model:
$$\text{Accuracy}=\frac{\text{Correct # of predictions}}{\text{Total # of predictions}}$$
**Instructions**: Write a program that can compute the accuracy on the dataset provided for you. You have to iterate over every row to get the corresponding words and feed them into you `get_country` function above.
<details>
<summary>
<font size="3" color="darkgreen"><b>Hints</b></font>
</summary>
<p>
<ul>
<li>Use <a href="https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.iterrows.html" > pandas.DataFrame.iterrows </a>.</li>
</ul>
</p>
```
# UNQ_C4 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
def get_accuracy(word_embeddings, data):
'''
Input:
word_embeddings: a dictionary where the key is a word and the value is its embedding
data: a pandas dataframe containing all the country and capital city pairs
Output:
accuracy: the accuracy of the model
'''
### START CODE HERE (REPLACE INSTANCES OF 'None' with your code) ###
# initialize num correct to zero
num_correct = 0
# loop through the rows of the dataframe
for i, row in data.iterrows():
# get city1
city1 = row[0]
# get country1
country1 = row[1]
# get city2
city2 = row[2]
# get country2
country2 = row[3]
# use get_country to find the predicted country2
predicted_country2, _ = get_country(city1, country1, city2, word_embeddings)
# if the predicted country2 is the same as the actual country2...
if predicted_country2 == country2:
# increment the number of correct by 1
num_correct += 1
# get the number of rows in the data dataframe (length of dataframe)
m = len(data)
# calculate the accuracy by dividing the number correct by m
accuracy = num_correct/m
### END CODE HERE ###
return accuracy
```
**NOTE: The cell below takes about 30 SECONDS to run.**
```
accuracy = get_accuracy(word_embeddings, data)
print(f"Accuracy is {accuracy:.2f}")
```
**Expected Output:**
$\approx$ 0.92
# 3.0 Plotting the vectors using PCA
Now you will explore the distance between word vectors after reducing their dimension.
The technique we will employ is known as
[*principal component analysis* (PCA)](https://en.wikipedia.org/wiki/Principal_component_analysis).
As we saw, we are working in a 300-dimensional space in this case.
Although from a computational perspective we were able to perform a good job,
it is impossible to visualize results in such high dimensional spaces.
You can think of PCA as a method that projects our vectors in a space of reduced
dimension, while keeping the maximum information about the original vectors in
their reduced counterparts. In this case, by *maximum infomation* we mean that the
Euclidean distance between the original vectors and their projected siblings is
minimal. Hence vectors that were originally close in the embeddings dictionary,
will produce lower dimensional vectors that are still close to each other.
You will see that when you map out the words, similar words will be clustered
next to each other. For example, the words 'sad', 'happy', 'joyful' all describe
emotion and are supposed to be near each other when plotted.
The words: 'oil', 'gas', and 'petroleum' all describe natural resources.
Words like 'city', 'village', 'town' could be seen as synonyms and describe a
similar thing.
Before plotting the words, you need to first be able to reduce each word vector
with PCA into 2 dimensions and then plot it. The steps to compute PCA are as follows:
1. Mean normalize the data
2. Compute the covariance matrix of your data ($\Sigma$).
3. Compute the eigenvectors and the eigenvalues of your covariance matrix
4. Multiply the first K eigenvectors by your normalized data. The transformation should look something as follows:
<img src = 'word_embf.jpg' width="width" height="height" style="width:800px;height:200px;"/>
**Instructions**:
You will write a program that takes in a data set where each row corresponds to a word vector.
* The word vectors are of dimension 300.
* Use PCA to change the 300 dimensions to `n_components` dimensions.
* The new matrix should be of dimension `m, n_componentns`.
* First de-mean the data
* Get the eigenvalues using `linalg.eigh`. Use `eigh` rather than `eig` since R is symmetric. The performance gain when using `eigh` instead of `eig` is substantial.
* Sort the eigenvectors and eigenvalues by decreasing order of the eigenvalues.
* Get a subset of the eigenvectors (choose how many principle components you want to use using `n_components`).
* Return the new transformation of the data by multiplying the eigenvectors with the original data.
<details>
<summary>
<font size="3" color="darkgreen"><b>Hints</b></font>
</summary>
<p>
<ul>
<li>Use <a href="https://docs.scipy.org/doc/numpy/reference/generated/numpy.mean.html" > numpy.mean(a,axis=None) </a> : If you set <code>axis = 0</code>, you take the mean for each column. If you set <code>axis = 1</code>, you take the mean for each row. Remember that each row is a word vector, and the number of columns are the number of dimensions in a word vector. </li>
<li>Use <a href="https://docs.scipy.org/doc/numpy/reference/generated/numpy.cov.html" > numpy.cov(m, rowvar=True) </a>. This calculates the covariance matrix. By default <code>rowvar</code> is <code>True</code>. From the documentation: "If rowvar is True (default), then each row represents a variable, with observations in the columns." In our case, each row is a word vector observation, and each column is a feature (variable). </li>
<li>Use <a href="https://docs.scipy.org/doc/numpy/reference/generated/numpy.linalg.eigh.html" > numpy.linalg.eigh(a, UPLO='L') </a> </li>
<li>Use <a href="https://docs.scipy.org/doc/numpy/reference/generated/numpy.argsort.html" > numpy.argsort </a> sorts the values in an array from smallest to largest, then returns the indices from this sort. </li>
<li>In order to reverse the order of a list, you can use: <code>x[::-1]</code>.</li>
<li>To apply the sorted indices to eigenvalues, you can use this format <code>x[indices_sorted]</code>.</li>
<li>When applying the sorted indices to eigen vectors, note that each column represents an eigenvector. In order to preserve the rows but sort on the columns, you can use this format <code>x[:,indices_sorted]</code></li>
<li>To transform the data using a subset of the most relevant principle components, take the matrix multiplication of the eigenvectors with the original data. </li>
<li>The data is of shape <code>(n_observations, n_features)</code>. </li>
<li>The subset of eigenvectors are in a matrix of shape <code>(n_features, n_components)</code>.</li>
<li>To multiply these together, take the transposes of both the eigenvectors <code>(n_components, n_features)</code> and the data (n_features, n_observations).</li>
<li>The product of these two has dimensions <code>(n_components,n_observations)</code>. Take its transpose to get the shape <code>(n_observations, n_components)</code>.</li>
</ul>
</p>
```
# UNQ_C5 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
def compute_pca(X, n_components=2):
"""
Input:
X: of dimension (m,n) where each row corresponds to a word vector
n_components: Number of components you want to keep.
Output:
X_reduced: data transformed in 2 dims/columns + regenerated original data
"""
### START CODE HERE (REPLACE INSTANCES OF 'None' with your code) ###
# mean center the data
X_demeaned = X - np.mean(X, axis = 0)
# calculate the covariance matrix
covariance_matrix = np.cov(X_demeaned, rowvar = False)
# calculate eigenvectors & eigenvalues of the covariance matrix
eigen_vals, eigen_vecs = np.linalg.eigh(covariance_matrix, UPLO='L')
# sort eigenvalue in increasing order (get the indices from the sort)
idx_sorted = np.argsort(eigen_vals)
# reverse the order so that it's from highest to lowest.
idx_sorted_decreasing = idx_sorted[::-1]
# sort the eigen values by idx_sorted_decreasing
eigen_vals_sorted = eigen_vals[idx_sorted_decreasing]
# sort eigenvectors using the idx_sorted_decreasing indices
eigen_vecs_sorted = eigen_vecs[:,idx_sorted_decreasing]
# select the first n eigenvectors (n is desired dimension
# of rescaled data array, or dims_rescaled_data)
eigen_vecs_subset = eigen_vecs_sorted[:, 0: n_components]
# transform the data by multiplying the transpose of the eigenvectors
# with the transpose of the de-meaned data
# Then take the transpose of that product.
X_reduced = np.dot(eigen_vecs_subset.T, X_demeaned.T).T
return X_reduced
# Testing your function
np.random.seed(1)
X = np.random.rand(3, 10)
X_reduced = compute_pca(X, n_components=2)
print("Your original matrix was " + str(X.shape) + " and it became:")
print(X_reduced)
```
**Expected Output:**
Your original matrix was: (3,10) and it became:
<table>
<tr>
<td>
0.43437323
</td>
<td>
0.49820384
</td>
</tr>
<tr>
<td>
0.42077249
</td>
<td>
-0.50351448
</td>
</tr>
<tr>
<td>
-0.85514571
</td>
<td>
0.00531064
</td>
</tr>
</table>
Now you will use your pca function to plot a few words we have chosen for you.
You will see that similar words tend to be clustered near each other.
Sometimes, even antonyms tend to be clustered near each other. Antonyms
describe the same thing but just tend to be on the other end of the scale
They are usually found in the same location of a sentence,
have the same parts of speech, and thus when
learning the word vectors, you end up getting similar weights. In the next week
we will go over how you learn them, but for now let's just enjoy using them.
**Instructions:** Run the cell below.
```
words = ['oil', 'gas', 'happy', 'sad', 'city', 'town',
'village', 'country', 'continent', 'petroleum', 'joyful']
# given a list of words and the embeddings, it returns a matrix with all the embeddings
X = get_vectors(word_embeddings, words)
print('You have 11 words each of 300 dimensions thus X.shape is:', X.shape)
# We have done the plotting for you. Just run this cell.
result = compute_pca(X, 2)
plt.scatter(result[:, 0], result[:, 1])
for i, word in enumerate(words):
plt.annotate(word, xy=(result[i, 0] - 0.05, result[i, 1] + 0.1))
plt.show()
```
The word vectors for 'gas', 'oil' and 'petroleum' appear related to each other,
because their vectors are close to each other. Similarly, 'sad', 'joyful'
and 'happy' all express emotions, and are also near each other.
| github_jupyter |
<div class="alert alert-block alert-info" style="margin-top: 20px">
<a href="http://cocl.us/topNotebooksPython101Coursera">
<img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Ad/TopAd.png" width="750" align="center">
</a>
</div>
<a href="https://cognitiveclass.ai/">
<img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Ad/CCLog.png" width="200" align="center">
</a>
<h1>Dictionaries in Python</h1>
<p><strong>Welcome!</strong> This notebook will teach you about the dictionaries in the Python Programming Language. By the end of this lab, you'll know the basics dictionary operations in Python, including what it is, and the operations on it.</p>
<h2>Table of Contents</h2>
<div class="alert alert-block alert-info" style="margin-top: 20px">
<ul>
<li>
<a href="#dic">Dictionaries</a>
<ul>
<li><a href="content">What are Dictionaries?</a></li>
<li><a href="key">Keys</a></li>
</ul>
</li>
<li>
<a href="#quiz">Quiz on Dictionaries</a>
</li>
</ul>
<p>
Estimated time needed: <strong>20 min</strong>
</p>
</div>
<hr>
<h2 id="Dic">Dictionaries</h2>
<h3 id="content">What are Dictionaries?</h3>
A dictionary consists of keys and values. It is helpful to compare a dictionary to a list. Instead of the numerical indexes such as a list, dictionaries have keys. These keys are the keys that are used to access values within a dictionary.
<img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%202/Images/DictsList.png" width="650" />
An example of a Dictionary <code>Dict</code>:
```
# Create the dictionary
Dict = {"key1": 1, "key2": "2", "key3": [3, 3, 3], "key4": (4, 4, 4), ('key5'): 5, (0, 1): 6}
Dict
```
The keys can be strings:
```
# Access to the value by the key
Dict["key1"]
```
Keys can also be any immutable object such as a tuple:
```
# Access to the value by the key
Dict[(0, 1)]
```
Each key is separated from its value by a colon "<code>:</code>". Commas separate the items, and the whole dictionary is enclosed in curly braces. An empty dictionary without any items is written with just two curly braces, like this "<code>{}</code>".
```
# Create a sample dictionary
release_year_dict = {"Thriller": "1982", "Back in Black": "1980", \
"The Dark Side of the Moon": "1973", "The Bodyguard": "1992", \
"Bat Out of Hell": "1977", "Their Greatest Hits (1971-1975)": "1976", \
"Saturday Night Fever": "1977", "Rumours": "1977"}
release_year_dict
```
In summary, like a list, a dictionary holds a sequence of elements. Each element is represented by a key and its corresponding value. Dictionaries are created with two curly braces containing keys and values separated by a colon. For every key, there can only be one single value, however, multiple keys can hold the same value. Keys can only be strings, numbers, or tuples, but values can be any data type.
It is helpful to visualize the dictionary as a table, as in the following image. The first column represents the keys, the second column represents the values.
<img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%202/Images/DictsStructure.png" width="650" />
<h3 id="key">Keys</h3>
You can retrieve the values based on the names:
```
# Get value by keys
release_year_dict['Thriller']
```
This corresponds to:
<img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%202/Images/DictsKeyOne.png" width="500" />
Similarly for <b>The Bodyguard</b>
```
# Get value by key
release_year_dict['The Bodyguard']
```
<img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%202/Images/DictsKeyTwo.png" width="500" />
Now let you retrieve the keys of the dictionary using the method <code>release_year_dict()</code>:
```
# Get all the keys in dictionary
release_year_dict.keys()
```
You can retrieve the values using the method <code>values()</code>:
```
# Get all the values in dictionary
release_year_dict.values()
```
We can add an entry:
```
# Append value with key into dictionary
release_year_dict['Graduation'] = '2007'
release_year_dict
```
We can delete an entry:
```
# Delete entries by key
del(release_year_dict['Thriller'])
del(release_year_dict['Graduation'])
release_year_dict
```
We can verify if an element is in the dictionary:
```
# Verify the key is in the dictionary
'The Bodyguard' in release_year_dict
```
<hr>
<h2 id="quiz">Quiz on Dictionaries</h2>
<b>You will need this dictionary for the next two questions:</b>
```
# Question sample dictionary
soundtrack_dic = {"The Bodyguard":"1992", "Saturday Night Fever":"1977"}
soundtrack_dic
```
a) In the dictionary <code>soundtrack_dict</code> what are the keys ?
```
# Write your code below and press Shift+Enter to execute
```
Double-click __here__ for the solution.
<!-- Your answer is below:
soundtrack_dic.keys() # The Keys "The Bodyguard" and "Saturday Night Fever"
-->
b) In the dictionary <code>soundtrack_dict</code> what are the values ?
```
# Write your code below and press Shift+Enter to execute
```
Double-click __here__ for the solution.
<!-- Your answer is below:
soundtrack_dic.values() # The values are "1992" and "1977"
-->
<hr>
<b>You will need this dictionary for the following questions:</b>
The Albums <b>Back in Black</b>, <b>The Bodyguard</b> and <b>Thriller</b> have the following music recording sales in millions 50, 50 and 65 respectively:
a) Create a dictionary <code>album_sales_dict</code> where the keys are the album name and the sales in millions are the values.
```
# Write your code below and press Shift+Enter to execute
```
Double-click __here__ for the solution.
<!-- Your answer is below:
album_sales_dict = {"The Bodyguard":50, "Back in Black":50, "Thriller":65}
-->
b) Use the dictionary to find the total sales of <b>Thriller</b>:
```
# Write your code below and press Shift+Enter to execute
```
Double-click __here__ for the solution.
<!-- Your answer is below:
album_sales_dict["Thriller"]
-->
c) Find the names of the albums from the dictionary using the method <code>keys</code>:
```
# Write your code below and press Shift+Enter to execute
```
Double-click __here__ for the solution.
<!-- Your answer is below:
album_sales_dict.keys()
-->
d) Find the names of the recording sales from the dictionary using the method <code>values</code>:
```
# Write your code below and press Shift+Enter to execute
```
Double-click __here__ for the solution.
<!-- Your answer is below:
album_sales_dict.values()
-->
<hr>
<h2>The last exercise!</h2>
<p>Congratulations, you have completed your first lesson and hands-on lab in Python. However, there is one more thing you need to do. The Data Science community encourages sharing work. The best way to share and showcase your work is to share it on GitHub. By sharing your notebook on GitHub you are not only building your reputation with fellow data scientists, but you can also show it off when applying for a job. Even though this was your first piece of work, it is never too early to start building good habits. So, please read and follow <a href="https://cognitiveclass.ai/blog/data-scientists-stand-out-by-sharing-your-notebooks/" target="_blank">this article</a> to learn how to share your work.
<hr>
<div class="alert alert-block alert-info" style="margin-top: 20px">
<h2>Get IBM Watson Studio free of charge!</h2>
<p><a href="https://cocl.us/bottemNotebooksPython101Coursera"><img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Ad/BottomAd.png" width="750" align="center"></a></p>
</div>
<h3>About the Authors:</h3>
<p><a href="https://www.linkedin.com/in/joseph-s-50398b136/" target="_blank">Joseph Santarcangelo</a> is a Data Scientist at IBM, and holds a PhD in Electrical Engineering. His research focused on using Machine Learning, Signal Processing, and Computer Vision to determine how videos impact human cognition. Joseph has been working for IBM since he completed his PhD.</p>
Other contributors: <a href="www.linkedin.com/in/jiahui-mavis-zhou-a4537814a">Mavis Zhou</a>
<hr>
<p>Copyright © 2018 IBM Developer Skills Network. This notebook and its source code are released under the terms of the <a href="https://cognitiveclass.ai/mit-license/">MIT License</a>.</p>
| github_jupyter |
```
import turtle
import random
grid=[[0,0,0,0,0,0,0,0,0],
[0,0,0,0,0,0,0,0,0],
[0,0,0,0,0,0,0,0,0],
[0,0,0,0,0,0,0,0,0],
[0,0,0,0,0,0,0,0,0],
[0,0,0,0,0,0,0,0,0],
[0,0,0,0,0,0,0,0,0],
[0,0,0,0,0,0,0,0,0],
[0,0,0,0,0,0,0,0,0]]
x=turtle.Turtle()
#x.tracer(0)
x.speed(9)
x.color("black")
x.hideturtle()
x_pos=-150
y_pos=150
def numbers(num,x_pos,y_pos,size):
x.penup()
x.goto(x_pos,y_pos)
x.write(num,font=("Arial",size,'normal'))
def drawGrid(grid):
intDim=35
for row in range(0,10):
if (row%3)==0:
x.pensize(3)
else:
x.pensize(1)
x.penup()
x.goto(min_x,max_y-row*intDim)
x.pendown()
x.goto(min_x+9*intDim,max_y-row*intDim)
for col in range(0,10):
if (col%3)==0:
x.pensize(3)
else:
x.pensize(1)
x.penup()
x.goto(min_x+col*intDim,max_y)
x.pendown()
x.goto(min_x+col*intDim,max_y-9*intDim)
for row in range (0,9):
for col in range (0,9):
if grid[row][col]!=0:
text(grid[row][col],min_x+col*intDim+9,max_y-row*intDim-intDim+8,18)
#A function to check if the grid is full
def checkGrid(grid):
for row in range(0,9):
for col in range(0,9):
if grid[row][col]==0:
return False
#We have a complete grid!
return True
break
grid[row][col]=0
numberList=[1,2,3,4,5,6,7,8,9]
drawGrid(grid)
fillGrid(grid)
turtle.done()
#Sudoku Generator Algorithm - www.101computing.net/sudoku-generator-algorithm/
import turtle
from random import randint, shuffle
from time import sleep
#initialise empty 9 by 9 grid
grid = []
grid.append([0, 0, 0, 0, 0, 0, 0, 0, 0])
grid.append([0, 0, 0, 0, 0, 0, 0, 0, 0])
grid.append([0, 0, 0, 0, 0, 0, 0, 0, 0])
grid.append([0, 0, 0, 0, 0, 0, 0, 0, 0])
grid.append([0, 0, 0, 0, 0, 0, 0, 0, 0])
grid.append([0, 0, 0, 0, 0, 0, 0, 0, 0])
grid.append([0, 0, 0, 0, 0, 0, 0, 0, 0])
grid.append([0, 0, 0, 0, 0, 0, 0, 0, 0])
grid.append([0, 0, 0, 0, 0, 0, 0, 0, 0])
myPen = turtle.Turtle()
myPen.tracer(0)
myPen.speed(0)
myPen.color("#000000")
myPen.hideturtle()
topLeft_x=-150
topLeft_y=150
def text(message,x,y,size):
FONT = ('Arial', size, 'normal')
myPen.penup()
myPen.goto(x,y)
myPen.write(message,align="left",font=FONT)
#A procedure to draw the grid on screen using Python Turtle
def drawGrid(grid):
intDim=35
for row in range(0,10):
if (row%3)==0:
myPen.pensize(3)
else:
myPen.pensize(1)
myPen.penup()
myPen.goto(topLeft_x,topLeft_y-row*intDim)
myPen.pendown()
myPen.goto(topLeft_x+9*intDim,topLeft_y-row*intDim)
for col in range(0,10):
if (col%3)==0:
myPen.pensize(3)
else:
myPen.pensize(1)
myPen.penup()
myPen.goto(topLeft_x+col*intDim,topLeft_y)
myPen.pendown()
myPen.goto(topLeft_x+col*intDim,topLeft_y-9*intDim)
for row in range (0,9):
for col in range (0,9):
if grid[row][col]!=0:
text(grid[row][col],topLeft_x+col*intDim+9,topLeft_y-row*intDim-intDim+8,18)
#A function to check if the grid is full
def checkGrid(grid):
for row in range(0,9):
for col in range(0,9):
if grid[row][col]==0:
return False
def fillthegrid(grid):
for i in range(0,81):
row=i//9
col=i%9
if grid[row][col]==0:
for k in range(1,10):
if k in grid[row]==False:
if k in (grid[0][col],grid[1][col],grid[2][col],grid[3][col],grid[4][col],grid[5][col],grid[6][col],grid[7][col],grid[8][col])==False:
square=[]
if row<3:
if col<3:
square=[grid[i][0:3] for i in range(0,3)]
elif col<6:
square=[grid[i][3:6] for i in range(0,3)]
else:
square=[grid[i][6:9] for i in range(0,3)]
elif row<6:
if col<3:
square=[grid[i][0:3] for i in range(3,6)]
elif col<6:
square=[grid[i][3:6] for i in range(3,6)]
else:
square=[grid[i][6:9] for i in range(3,6)]
else:
if col<3:
square=[grid[i][0:3] for i in range(6,9)]
elif col<6:
square=[grid[i][3:6] for i in range(6,9)]
else:
square=[grid[i][6:9] for i in range(6,9)]
#We have a complete grid!
return True
numberList=[1,2,3,4,5,6,7,8,9]
#shuffle(numberList)
#Generate a Fully Solved Grid
fillGrid(grid)
drawGrid(grid)
myPen.getscreen().update()
sleep(1)
turtle.done()
```
| github_jupyter |
# RNN in PyTorch
> Recurrent Neural Networks in PyTorch
- badges: true
- comments: true
- author: Naman Manchanda
- categories: [rnn, jupyter, pytorch, python]
<h1><center>Recurrent Neural Network in PyTorch</center></h1>
Table of Contents: <a id=100></a>
1. [Packages](#1)
2. [Data definition](#2)
- 2.1 [Declaring a tensor `x`](#3)
- 2.2 [Creating a tensor `y` as a sin function of `x`](#4)
- 2.3 [Plotting `y`](#5)
3. [Batching the data](#6)
- 3.1 [Splitting the data in train/test set](#7)
- 3.2 [Creating the batches of data](#8)
4. [Defining the model](#9)
- 4.1 [Model class](#10)
- 4.2 [Model instantiation](#11)
- 4.3 [Training](#12)
5. [Alcohol Sales dataset](#13)
- 5.1 [Loading and plotting](#14)
- 5.2 [Prepare and normalize](#15)
- 5.3 [Modelling](#16)
- 5.4 [Predictions](#17)
Recurrent Neural Networks are a type of neural networks that are designed to work on sequence prediction models. RNNs can be used for text data, speech data, classification problems and generative models. Unlike ANNs, RNNs' prediction are based on the past prediction as well as the current input. RNNs are networks with loops in them allowing information to persist.
Each node of an **RNN** consists of 2 inputs:
1. Memory unit
2. Event unit
`M(t-1)` is the memory unit or the output of the previous prediction. `E(t)` is the current event or the information being provided at the present time. `M(t)` is the output of the current node or the output at the present time in the sequence.
### 1. Packages <a id=1></a>
[back to top](#100)
```
import numpy as np
import pandas as pd
import torch
import torch.nn as nn
import matplotlib.pyplot as plt
%matplotlib inline
```
### 2. Data definition <a id=2></a>
[back to top](#100)
In this notebook, I'm going to train a very simple LSTM model, which is a type of RNN architecture to do time series prediction. Given some input data, it should be able to generate a prediction for the next step. I'll be using a **Sin** wave as an example as it's very easy to visualiase the behaviour of a sin wave.
#### 2.1 Declaring a tensor `x` <a id=3></a>
```
x = torch.linspace(0,799,800)
```
#### 2.2 Creating a tensor `y` as a sin function of `x` <a id=4></a>
```
y = torch.sin(x*2*3.1416/40)
```
#### 2.3 Plotting `y` <a id=5></a>
```
plt.figure(figsize=(12,4))
plt.xlim(-10,801)
plt.grid(True)
plt.xlabel("x")
plt.ylabel("sin")
plt.title("Sin plot")
plt.plot(y.numpy(),color='#8000ff')
plt.show()
```
### 3. Batching the data <a id=6></a>
[back to top](#100)
#### 3.1 Splitting the data in train/test set <a id=7></a>
```
test_size = 40
train_set = y[:-test_size]
test_set = y[-test_size:]
```
##### 3.1.1 Plotting the training/testing set
```
plt.figure(figsize=(12,4))
plt.xlim(-10,801)
plt.grid(True)
plt.xlabel("x")
plt.ylabel("sin")
plt.title("Sin plot")
plt.plot(train_set.numpy(),color='#8000ff')
plt.plot(range(760,800),test_set.numpy(),color="#ff8000")
plt.show()
```
#### 3.2 Creating the batches of data <a id=8></a>
While working with LSTM models, we divide the training sequence into series of overlapping windows. The label used for comparison is the next value in the sequence.
For example if we have series of of 12 records and a window size of 3, we feed [x1, x2, x3] into the model, and compare the prediction to `x4`. Then we backdrop, update parameters, and feed [x2, x3, x4] into the model and compare the prediction to `x5`. To ease this process, I'm defining a function `input_data(seq,ws)` that created a list of (seq,labels) tuples. If `ws` is the window size, then the total number of (seq,labels) tuples will be `len(series)-ws`.
```
def input_data(seq,ws):
out = []
L = len(seq)
for i in range(L-ws):
window = seq[i:i+ws]
label = seq[i+ws:i+ws+1]
out.append((window,label))
return out
```
##### 3.2.1 Calling the `input_data` function
The length of `x` = 800
The length of `train_set` = 800 - 40 = 760
The length of `train_data` = 760 - 40 - 720
```
window_size = 40
train_data = input_data(train_set, window_size)
len(train_data)
```
##### 3.2.2 Checking the 1st value from train_data
```
train_data[0]
```
### 4. Defining the model <a id=9></a>
[back to top](#100)
#### 4.1 Model Class <a id=10></a>
```
class LSTM(nn.Module):
def __init__(self,input_size = 1, hidden_size = 50, out_size = 1):
super().__init__()
self.hidden_size = hidden_size
self.lstm = nn.LSTM(input_size, hidden_size)
self.linear = nn.Linear(hidden_size,out_size)
self.hidden = (torch.zeros(1,1,hidden_size),torch.zeros(1,1,hidden_size))
def forward(self,seq):
lstm_out, self.hidden = self.lstm(seq.view(len(seq),1,-1), self.hidden)
pred = self.linear(lstm_out.view(len(seq),-1))
return pred[-1]
```
#### 4.2 Model Instantiation <a id = 11></a>
```
torch.manual_seed(42)
model = LSTM()
criterion = nn.MSELoss()
optimizer = torch.optim.SGD(model.parameters(), lr=0.01)
```
##### 4.2.1 Printing the model
```
model
```
#### 4.3 Training <a id = 12></a>
During training, I'm visualising the prediction process for the test data on the go. It will give a better understanding of how the training is being carried out in each epoch. The training sequence is represented in <span style="color:#8000ff">purple</span> while the predicted sequence in represented in <span style="color:#ff8000">orange</span>.
```
epochs = 10
future = 40
for i in range(epochs):
for seq, y_train in train_data:
optimizer.zero_grad()
model.hidden = (torch.zeros(1,1,model.hidden_size),
torch.zeros(1,1,model.hidden_size))
y_pred = model(seq)
loss = criterion(y_pred, y_train)
loss.backward()
optimizer.step()
print(f"Epoch {i} Loss: {loss.item()}")
preds = train_set[-window_size:].tolist()
for f in range(future):
seq = torch.FloatTensor(preds[-window_size:])
with torch.no_grad():
model.hidden = (torch.zeros(1,1,model.hidden_size),
torch.zeros(1,1,model.hidden_size))
preds.append(model(seq).item())
loss = criterion(torch.tensor(preds[-window_size:]), y[760:])
print(f"Performance on test range: {loss}")
plt.figure(figsize=(12,4))
plt.xlim(700,801)
plt.grid(True)
plt.plot(y.numpy(),color='#8000ff')
plt.plot(range(760,800),preds[window_size:],color='#ff8000')
plt.show()
```
### 5. Alcohol Sales dataset <a id=13></a>
[back to top](#100)
#### 5.1 Loading and plotting <a id=14></a>
##### 5.1.1 Importing the data
```
df = pd.read_csv("/kaggle/input/for-simple-exercises-time-series-forecasting/Alcohol_Sales.csv", index_col = 0, parse_dates = True)
df.head()
```
##### 5.1.2 Dropping the empty rows
```
df.dropna(inplace=True)
len(df)
```
##### 5.1.3 Plotting the Time Series Data
```
plt.figure(figsize = (12,4))
plt.title('Alcohol Sales')
plt.ylabel('Sales in million dollars')
plt.grid(True)
plt.autoscale(axis='x',tight=True)
plt.plot(df['S4248SM144NCEN'],color='#8000ff')
plt.show()
```
#### 5.2 Prepare and normalize <a id=15></a>
##### 5.2.1 Preparing the data
```
#extracting the time series values
y = df['S4248SM144NCEN'].values.astype(float)
#defining a test size
test_size = 12
#create train and test splits
train_set = y[:-test_size]
test_set = y[-test_size:]
test_set
```
##### 5.2.2 Normalize the data
```
from sklearn.preprocessing import MinMaxScaler
# instantiate a scaler
scaler = MinMaxScaler(feature_range=(-1, 1))
# normalize the training set
train_norm = scaler.fit_transform(train_set.reshape(-1, 1))
```
##### 5.2.3 Prepare data for LSTM model
```
# convert train_norm to a tensor
train_norm = torch.FloatTensor(train_norm).view(-1)
# define a window size
window_size = 12
# define a function to create sequence/label tuples
def input_data(seq,ws):
out = []
L = len(seq)
for i in range(L-ws):
window = seq[i:i+ws]
label = seq[i+ws:i+ws+1]
out.append((window,label))
return out
# apply input_data to train_norm
train_data = input_data(train_norm, window_size)
len(train_data)
```
##### 5.2.4 Printing the first tuple
```
train_data[0]
```
#### 5.3 Modelling <a id=16></a>
##### 5.3.1 Model definition
```
class LSTMnetwork(nn.Module):
def __init__(self,input_size=1,hidden_size=100,output_size=1):
super().__init__()
self.hidden_size = hidden_size
# add an LSTM layer:
self.lstm = nn.LSTM(input_size,hidden_size)
# add a fully-connected layer:
self.linear = nn.Linear(hidden_size,output_size)
# initializing h0 and c0:
self.hidden = (torch.zeros(1,1,self.hidden_size),
torch.zeros(1,1,self.hidden_size))
def forward(self,seq):
lstm_out, self.hidden = self.lstm(
seq.view(len(seq),1,-1), self.hidden)
pred = self.linear(lstm_out.view(len(seq),-1))
return pred[-1]
```
##### 5.3.3 Instantiation, loss and optimizer
```
torch.manual_seed(42)
# instantiate
model = LSTMnetwork()
# loss
criterion = nn.MSELoss()
#optimizer
optimizer = torch.optim.Adam(model.parameters(), lr=0.001)
model
```
##### 5.3.4 Training
```
epochs = 100
import time
start_time = time.time()
for epoch in range(epochs):
for seq, y_train in train_data:
optimizer.zero_grad()
model.hidden = (torch.zeros(1,1,model.hidden_size),
torch.zeros(1,1,model.hidden_size))
y_pred = model(seq)
loss = criterion(y_pred, y_train)
loss.backward()
optimizer.step()
print(f'Epoch: {epoch+1:2} Loss: {loss.item():10.8f}')
print(f'\nDuration: {time.time() - start_time:.0f} seconds')
```
#### 5.4 Predictions <a id=17></a>
##### 5.4.1 Test set predictions
```
future = 12
preds = train_norm[-window_size:].tolist()
model.eval()
for i in range(future):
seq = torch.FloatTensor(preds[-window_size:])
with torch.no_grad():
model.hidden = (torch.zeros(1,1,model.hidden_size),
torch.zeros(1,1,model.hidden_size))
preds.append(model(seq).item())
preds[window_size:]
```
##### 5.4.2 Original test set
```
df['S4248SM144NCEN'][-12:]
```
##### 5.4.3 Inverting the normalised values
```
true_predictions = scaler.inverse_transform(np.array(preds[window_size:]).reshape(-1, 1))
true_predictions
```
##### 5.4.4 Plotting
```
x = np.arange('2018-02-01', '2019-02-01', dtype='datetime64[M]').astype('datetime64[D]')
plt.figure(figsize=(12,4))
plt.title('Alcohol Sales')
plt.ylabel('Sales in million dollars')
plt.grid(True)
plt.autoscale(axis='x',tight=True)
plt.plot(df['S4248SM144NCEN'], color='#8000ff')
plt.plot(x,true_predictions, color='#ff8000')
plt.show()
```
##### 5.5.5 Zooming the test predictions
```
fig = plt.figure(figsize=(12,4))
plt.title('Alcohol Sales')
plt.ylabel('Sales in million dollars')
plt.grid(True)
plt.autoscale(axis='x',tight=True)
fig.autofmt_xdate()
plt.plot(df['S4248SM144NCEN']['2017-01-01':], color='#8000ff')
plt.plot(x,true_predictions, color='#ff8000')
plt.show()
```
### If you liked the notebook, consider giving an upvote.
[back to top](#100)
| github_jupyter |
# `*args` and `**kwargs`
Work with Python long enough, and eventually you will encounter `*args` and `**kwargs`. These strange terms show up as parameters in function definitions. What do they do? Let's review a simple function:
```
def myfunc(a,b):
return sum((a,b))*.05
myfunc(40,60)
```
This function returns 5% of the sum of **a** and **b**. In this example, **a** and **b** are *positional* arguments; that is, 40 is assigned to **a** because it is the first argument, and 60 to **b**. Notice also that to work with multiple positional arguments in the `sum()` function we had to pass them in as a tuple.
What if we want to work with more than two numbers? One way would be to assign a *lot* of parameters, and give each one a default value.
```
def myfunc(a=0,b=0,c=0,d=0,e=0):
return sum((a,b,c,d,e))*.05
myfunc(40,60,20)
```
Obviously this is not a very efficient solution, and that's where `*args` comes in.
## `*args`
When a function parameter starts with an asterisk, it allows for an *arbitrary number* of arguments, and the function takes them in as a tuple of values. Rewriting the above function:
```
def myfunc(*args):
return sum(args)*.05
myfunc(40,60,20)
```
Notice how passing the keyword "args" into the `sum()` function did the same thing as a tuple of arguments.
It is worth noting that the word "args" is itself arbitrary - any word will do so long as it's preceded by an asterisk. To demonstrate this:
```
def myfunc(*spam):
return sum(spam)*.05
myfunc(40,60,20)
```
## `**kwargs`
Similarly, Python offers a way to handle arbitrary numbers of *keyworded* arguments. Instead of creating a tuple of values, `**kwargs` builds a dictionary of key/value pairs. For example:
```
def myfunc(**kwargs):
if 'fruit' in kwargs:
print(f"My favorite fruit is {kwargs['fruit']}") # review String Formatting and f-strings if this syntax is unfamiliar
else:
print("I don't like fruit")
myfunc(fruit='pineapple')
myfunc()
```
## `*args` and `**kwargs` combined
You can pass `*args` and `**kwargs` into the same function, but `*args` have to appear before `**kwargs`
```
def myfunc(*args, **kwargs):
if 'fruit' and 'juice' in kwargs:
print(f"I like {' and '.join(args)} and my favorite fruit is {kwargs['fruit']}")
print(f"May I have some {kwargs['juice']} juice?")
else:
pass
myfunc('eggs','spam',fruit='cherries',juice='orange')
myd=dict(fruit=1,juice='orange')
print(myd)
ad= {'fruit': 1, 'juice': 'orange'}
print(ad)
```
Placing keyworded arguments ahead of positional arguments raises an exception:
```
myfunc(fruit='cherries',juice='orange','eggs','spam')
```
As with "args", you can use any name you'd like for keyworded arguments - "kwargs" is just a popular convention.
That's it! Now you should understand how `*args` and `**kwargs` provide the flexibilty to work with arbitrary numbers of arguments!
| github_jupyter |
# Sky with the Moon Up
* https://desi.lbl.gov/trac/wiki/CommissioningCommissioningPlanning/commishdata#NightSky
* https://portal.nersc.gov/project/desi/collab/nightwatch/kpno/20191112/exposures.html
* http://desi-www.kpno.noao.edu:8090/nightsum/nightsum-2019-11-12/nightsum.html
**Data**
* /global/projecta/projectdirs/desi/spectro/redux/daily/exposures/20191112
* /global/projecta/projectdirs/desi/spectro/redux/daily/exposures/20191112
* Exposures 27337-27396
**Data Model**
* https://desidatamodel.readthedocs.io/en/latest/DESI_SPECTRO_REDUX/SPECPROD/exposures/NIGHT/EXPID/sky-CAMERA-EXPID.html
John Moustakas
```
import os, sys
import numpy as np
import fitsio
import astropy.units as u
from astropy.table import Table, vstack
from astropy.coordinates import SkyCoord
from astropy.io import fits
import desispec.io
import desimodel.io
import speclite
import specsim.atmosphere
import specsim.simulator
from desietcimg.db import DB, Exposures
import matplotlib.pyplot as plt
import seaborn as sns
sns.set(context='talk', style='ticks', font_scale=1.1)
%matplotlib inline
topdir = os.getenv('DESI_ROOT')+'/spectro'
gfadir = '/project/projectdirs/desi/users/ameisner/GFA/reduced/v0001'
```
#### Load the GFA filter curve.
```
def load_gfa_filter():
if False:
rfilt = speclite.filters.load_filter('decam2014-r')
else:
if False:
filtfile = '/global/homes/a/ameisner/ci_throughput/etc/gfa_filter_transmission_DESI-1297.dat'
filtwave, filtresp, _, _ = np.loadtxt(filtfile, unpack=True)
else:
filt = Table.read('/global/homes/a/ameisner/ci_throughput/etc/gfa_throughput-airmass_1.00.fits')
filtwave, filtresp = filt['LAMBDA_NM'], filt['THROUGHPUT']
filtresp[filtresp < 0] = 0
filtresp[0] = 0
filtresp[-1] = 0
srt = np.argsort(filtwave)
filtwave, filtresp = filtwave[srt] * 10, filtresp[srt]
rfilt = speclite.filters.FilterResponse(wavelength=filtwave * u.Angstrom,
response=filtresp,
meta=dict(group_name='gfa', band_name='r'))
return rfilt
rfilt = load_gfa_filter()
plt.plot(rfilt.wavelength, rfilt.response)
```
#### Specify the night and range of GFA and spectrograph exposure IDs.
```
night = 20191112
expid_start, expid_end = 27337, 27396
expids = np.arange(expid_end - expid_start + 1) + expid_start
```
#### Initialize access to the online database.
https://github.com/desihub/desicmx/blob/master/analysis/gfa/DESI-Online-Database-Tutorial.ipynb
```
if not os.path.exists('db.yaml'):
import getpass
pw = getpass.getpass(prompt='Enter database password: ')
with open('db.yaml', 'w') as f:
print('host: db.replicator.dev-cattle.stable.spin.nersc.org', file=f)
print('dbname: desi_dev', file=f)
print('port: 60042', file=f)
print('user: desi_reader', file=f)
print(f'password: {pw}', file=f)
print('Created db.yaml')
db = DB()
ExpInfo = Exposures(db)
for col in ('airmass', 'skyra', 'skydec', 'moonangl', 'moonra', 'moondec'):
print(col, ExpInfo(27339, col))
```
#### Read A. Meisner's reductions to get the sky background in the GFAs vs moon separation.
```
def read_gfa(night, expid, raw=False):
sexpid = '{:08d}'.format(expid)
rawfname = topdir+'/data/{0}/{1}/gfa-{1}.fits.fz'.format(night, sexpid)
fname = gfadir+'/{0}/{1}/gfa-{1}_ccds.fits'.format(night, sexpid)
if os.path.isfile(fname):
# get the target-moon separation
hdr = fitsio.read_header(rawfname, ext=1)
#print(hdr)
ra, dec, moonra, moondec = hdr['SKYRA'], hdr['SKYDEC'], hdr['MOONRA'], hdr['MOONDEC']
csky = SkyCoord(ra=ra*u.degree, dec=dec*u.degree, frame='icrs')
cmoon = SkyCoord(ra=moonra*u.degree, dec=moondec*u.degree, frame='icrs')
moonsep = np.array(csky.separation(cmoon).value).astype('f4') # [degree]
#print('Reading {}'.format(fname))
data = Table.read(fname)
#print(data.colnames)
# pack into a table
out = Table()
out['sky_mag_ab'] = [np.mean(data['sky_mag_ab'])]
out['sky_mag_ab_err'] = [np.std(data['sky_mag_ab']) / np.sqrt(len(data))]
out['moonsep'] = [moonsep]
#out['airmass'] = [hdr['AIRMASS']]
out = out[np.argsort(out['moonsep'])[::-1]]
return out
else:
return None
#data = read_gfa(night, 27390)
#data
def gfa_sky(night):
out = []
for expid in expids:
dd = read_gfa(night, expid)
if dd is not None:
out.append(dd)
return vstack(out)
#Table(fitsio.read(os.getenv('DESI_ROOT')+'/users/ameisner/GFA/files/skymags-prelim.fits')).colnames
Table(fitsio.read(os.getenv('DESI_ROOT')+'/users/ameisner/GFA/files/skymags-prelim.with_airmass.fits')).colnames
```
Read Aaron's updated GFA analysis outputs.
```
def gfa_sky_updated():
#gfafile = os.getenv('DESI_ROOT')+'/users/ameisner/GFA/files/skymags-prelim.fits'
gfafile = os.getenv('DESI_ROOT')+'/users/ameisner/GFA/files/skymags-prelim.with_airmass.fits'
cat = Table(fitsio.read(gfafile, lower=True))
cat = cat[(cat['expid'] >= expid_start) * (cat['expid'] <= expid_end)]
ra, dec, moonra, moondec = cat['skyra'], cat['skydec'], cat['ra_moon_deg'], cat['dec_moon_deg']
#print(moonra, moondec)
csky = SkyCoord(ra=ra*u.degree, dec=dec*u.degree, frame='icrs')
cmoon = SkyCoord(ra=moonra*u.degree, dec=moondec*u.degree, frame='icrs')
moonsep = np.array(csky.separation(cmoon).value) # [degree]
out = Table()
out['camera'] = cat['extname']
out['expid'] = cat['expid']
out['airmass'] = cat['airmass']
out['ra'] = cat['skyra']
out['dec'] = cat['skydec']
out['moon_phase'] = cat['moon_phase']
out['sky_mag_ab'] = cat['skymag_median_top_camera'].astype('f4')
#out['sky_mag_ab_err'] = [np.std(data['sky_mag_ab']) / np.sqrt(len(data))]
out['moonsep'] = moonsep.astype('f4')
out = out[np.argsort(out['moonsep'])[::-1]]
return out
if False:
gfa = gfa_sky(night)
else:
gfa = gfa_sky_updated()
gfa
fig, ax = plt.subplots(figsize=(8, 6))
for cam, mark in zip(set(gfa['camera']), ('s', 'o')):
ww = gfa['camera'] == cam
ax.scatter(gfa['moonsep'][ww], gfa['sky_mag_ab'][ww],
label=cam, marker=mark, s=80, alpha=0.7)
#ax.invert_yaxis()
#ax.set_ylim(17.5, 14.5)
ax.set_xlabel('Target-Moon Separation (degree)')
ax.set_ylabel('Sky Brightness (AB mag)')
ax.legend()
```
#### Read the spectroscopic reductions to get the sky spectra vs moon separation.
```
skymodel = specsim.simulator.Simulator('desi').atmosphere
skymodel.airmass = 1.1
skymodel.moon.moon_phase = 0.05
skymodel.moon.moon_zenith = 40 * u.deg
for moonsep in (45, 10):
skymodel.moon.separation_angle = moonsep * u.deg
skymodelflux = (skymodel.surface_brightness * np.pi * (0.75 * u.arcsec) ** 2).to(
u.erg / (u.Angstrom * u.cm ** 2 * u.s))
plt.plot(skymodel._wavelength, skymodelflux.value)
def read_spec(night, expid):
sexpid = '{:08d}'.format(expid)
datadir = topdir+'/redux/daily/exposures/{}/{}/'.format(night, sexpid)
if os.path.isdir(datadir):
#fr = desispec.io.read_frame('{}/frame-r3-{}.fits'.format(datadir, sexpid))
sp = desispec.io.read_frame('{}/sframe-r3-{}.fits'.format(datadir, sexpid))
sky = desispec.io.read_sky('{}/sky-r3-{}.fits'.format(datadir, sexpid))
return sp, sky
#return fr, sp, sky
else:
return None, None
#sp, sky = read_spec(night, 27339)
def spec_sky():
import astropy.time
import astropy.coordinates
# hack!
from astropy.utils.iers import conf
from astropy.utils import iers
conf.auto_max_age = None
iers.Conf.iers_auto_url.set('ftp://cddis.gsfc.nasa.gov/pub/products/iers/finals2000A.all')
moonphase = 0.99 # from elog
loc = astropy.coordinates.EarthLocation.of_site('Kitt Peak')
# skymodel
skymodel = specsim.simulator.Simulator('desi').atmosphere
_rfilt = speclite.filters.FilterSequence([rfilt])
rand = np.random.RandomState(seed=1)
allmoonsep, meansky, stdsky, skymodelmag = [], [], [], []
specsky, specwave = [], []
for expid in expids:
sp, sky = read_spec(night, expid)
if sky is not None:
# blarg! there's no metadata in the headers (or in the database), so
# choose the RA, Dec, and airmass from the nearest GFA exposure.
this = np.argmin(np.abs(gfa['expid'] - expid))
ra, dec = gfa['ra'][this], gfa['dec'][this]
moonphase, airmass = gfa['moon_phase'][this], gfa['airmass'][this],
# get the object-moon separation!!
#moonsep.append(sky.header['PROGRAM'][:3])
hdr = sky.header
date, obstime = hdr['DATE-OBS'], hdr['TIME-OBS']
mjd, exptime = hdr['MJD-OBS'], hdr['EXPTIME']
time = astropy.time.Time(mjd, format='mjd')
#time = astropy.time.Time(date, format='isot', scale='utc')
moonpos = astropy.coordinates.get_moon(time, loc)
moonra, moondec = moonpos.ra, moonpos.dec
csky = SkyCoord(ra=ra*u.degree, dec=dec*u.degree, frame='icrs')
cmoon = SkyCoord(ra=moonra, dec=moondec, frame='icrs')
moonsep = csky.separation(cmoon)
allmoonsep.append(moonsep.value)
moon_altaz = moonpos.transform_to(astropy.coordinates.AltAz(obstime=time, location=loc))
moon_az = moon_altaz.az.value * u.degree
moon_zenith = (90. - moon_altaz.alt.value) * u.degree
# get the model sky brightness
skymodel.airmass = airmass
skymodel.moon.moon_phase = 1 - moonphase
skymodel.moon.moon_zenith = moon_zenith
skymodel.moon.separation_angle = moonsep
print(skymodel.airmass, skymodel.moon.moon_phase, skymodel.moon.moon_zenith.value,
skymodel.moon.separation_angle.value)
skymodelflux = (skymodel.surface_brightness * np.pi * (0.75 * u.arcsec) ** 2).to(
u.erg / (u.Angstrom * u.cm ** 2 * u.s))
pad_skymodelflux, pad_skymodelwave = _rfilt.pad_spectrum(
skymodelflux.value, skymodel._wavelength)
#plt.plot(pad_skymodelwave, pad_skymodelflux)
#import pdb ; pdb.set_trace()
skymodel_abmags = _rfilt.get_ab_magnitudes(pad_skymodelflux, pad_skymodelwave)[rfilt.name]
skymodelmag.append(skymodel_abmags)
# now the data: convolve the spectrum with the r-band filter curve
keep = sp.fibermap['OBJTYPE'] == 'SKY'
padflux, padwave = _rfilt.pad_spectrum(sky.flux[keep, :], sky.wave, method='edge')
abmags = _rfilt.get_ab_magnitudes(padflux, padwave)[rfilt.name]
meansky.append(np.mean(abmags))
stdsky.append(np.std(abmags) / np.sqrt(len(abmags)))
# get the median spectra for this moon separation
specwave.append(sky.wave)
specsky.append(np.percentile(sky.flux[keep, :], axis=0, q=50))
#q25sky = np.percentile(sky.flux[keep, :], axis=0, q=25)
#q75sky = np.percentile(sky.flux[keep, :], axis=0, q=75)
#these = rand.choice(keep, size=20, replace=False)
#[plt.plot(sky.wave, sky.flux[ii, :], alpha=1.0) for ii in these]
#plt.plot(sky.wave, medsky, alpha=0.7, color='k')
out = Table()
out['moonsep'] = np.hstack(allmoonsep).astype('f4')
out['meansky'] = np.hstack(meansky).astype('f4')
out['stdsky'] = np.hstack(stdsky).astype('f4')
out['specwave'] = specwave
out['specsky'] = specsky
out['skymodel'] = np.hstack(skymodelmag).astype('f4')
out = out[np.argsort(out['moonsep'])[::-1]]
return out
spec = spec_sky()
spec
fig, ax = plt.subplots(figsize=(8, 6))
ref = spec['moonsep'].argmax()
#print(ref, spec['specsky'][ref])
for ss in spec:
#print(ss['specwave'], ss['specsky'] / spec['specsky'][ref])
ax.plot(ss['specwave'], ss['specsky'],# / spec['specsky'][0],
label=r'{:.0f}$^{{\circ}}$'.format(ss['moonsep']))
ax.plot(rfilt.wavelength, rfilt.response / np.max(rfilt.response) * np.median(spec['specsky'][-1]),
color='k', ls='--', lw=2)
ax.set_xlim(5500, 7800)
#ax.set_yscale('log')
ax.set_xlabel('Wavelength ($\AA$)')
ax.set_ylabel('Sky Spectra (counts / $\AA$)')
ax.legend(ncol=5, fontsize=12)
#ax.set_title('Night {}'.format(night))
fig, ax = plt.subplots(figsize=(8, 6))
for cam, mark in zip(set(gfa['camera']), ('s', 'o')):
ww = gfa['camera'] == cam
ax.plot(gfa['moonsep'][ww], gfa['sky_mag_ab'][ww], '{}-'.format(mark),
alpha=0.8, markersize=10, label='GFA-{}'.format(cam))
ax.plot(spec['moonsep'], spec['skymodel'], 'o--', color='k', label='Model Sky')
ax.invert_xaxis()
#ax.set_ylim(17.5, 14.5)
ax.set_xlabel('Target-Moon Separation (degree)')
ax.set_ylabel('$r$-band Sky Brightness (AB mag)')
ax.legend()
fig, ax = plt.subplots(figsize=(10, 8))
specref = spec['meansky'][spec['moonsep'].argmax()]
for cam, mark in zip(set(gfa['camera']), ('s', 'o')):
ww = gfa['camera'] == cam
gfaref = gfa['sky_mag_ab'][gfa['moonsep'].argmax()]
ax.plot(gfa['moonsep'][ww], gfa['sky_mag_ab'][ww] - gfaref, '{}-'.format(mark),
alpha=0.8, markersize=10,
label='GFA-{}'.format(cam))#, marker=mark, s=80)
#ax.scatter(gfa['moonsep'], gfa['sky_mag_ab'] - gfaref, label='GFAs')
ax.plot(spec['moonsep'], spec['meansky'] - specref, '^--',
alpha=0.8, markersize=13, label='Spectra')
modelref = spec['skymodel'][spec['moonsep'].argmax()]
ax.plot(spec['moonsep'], spec['skymodel'] - modelref, 'p-.',
alpha=0.8, markersize=13, label='Sky Model')
ax.invert_xaxis()
#ax.set_ylim(17.5, 14.5)
ax.set_xlabel('Target-Moon Separation (degree)')
ax.set_ylabel('Relative $r$-band Sky Brightness (AB mag)')
ax.legend(loc='lower left')
#ax.set_title('Night {}'.format(night))
```
#### Playing around
```
import specsim.simulator
desi = specsim.simulator.Simulator('desi', num_fibers=1)
desi.instrument.fiberloss_method = 'table'
desi.simulate()
plt.plot(desi.atmosphere._wavelength.value, desi.atmosphere.surface_brightness.value)
plt.xlim(5500, 8500)
from astropy.coordinates import EarthLocation
loc = EarthLocation.of_site('Kitt Peak')
loc
time = astropy.time.Time(t['DATE'], format='jd')
moon_position = astropy.coordinates.get_moon(time, loc)
moon_ra = moon_position.ra.value
moon_dec = moon_position.dec.value
moon_position_altaz = moon_position.transform_to(astropy.coordinates.AltAz(obstime=time, location=location))
moon_alt = moon_position_altaz.alt.value
moon_az = moon_position_altaz.az.value
EarthLocation.from_geodetic(lat='-30d10m10.78s', lon='-70d48m23.49s', height=2241.4*u.m)
```
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/Datasets/Terrain/srtm.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/Datasets/Terrain/srtm.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/Datasets/Terrain/srtm.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API and geemap
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geemap](https://github.com/giswqs/geemap). The **geemap** Python package is built upon the [ipyleaflet](https://github.com/jupyter-widgets/ipyleaflet) and [folium](https://github.com/python-visualization/folium) packages and implements several methods for interacting with Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, and `Map.centerObject()`.
The following script checks if the geemap package has been installed. If not, it will install geemap, which automatically installs its [dependencies](https://github.com/giswqs/geemap#dependencies), including earthengine-api, folium, and ipyleaflet.
**Important note**: A key difference between folium and ipyleaflet is that ipyleaflet is built upon ipywidgets and allows bidirectional communication between the front-end and the backend enabling the use of the map to capture user input, while folium is meant for displaying static data only ([source](https://blog.jupyter.org/interactive-gis-in-jupyter-with-ipyleaflet-52f9657fa7a)). Note that [Google Colab](https://colab.research.google.com/) currently does not support ipyleaflet ([source](https://github.com/googlecolab/colabtools/issues/60#issuecomment-596225619)). Therefore, if you are using geemap with Google Colab, you should use [`import geemap.eefolium`](https://github.com/giswqs/geemap/blob/master/geemap/eefolium.py). If you are using geemap with [binder](https://mybinder.org/) or a local Jupyter notebook server, you can use [`import geemap`](https://github.com/giswqs/geemap/blob/master/geemap/geemap.py), which provides more functionalities for capturing user input (e.g., mouse-clicking and moving).
```
# Installs geemap package
import subprocess
try:
import geemap
except ImportError:
print('geemap package not installed. Installing ...')
subprocess.check_call(["python", '-m', 'pip', 'install', 'geemap'])
# Checks whether this notebook is running on Google Colab
try:
import google.colab
import geemap.eefolium as emap
except:
import geemap as emap
# Authenticates and initializes Earth Engine
import ee
try:
ee.Initialize()
except Exception as e:
ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
The default basemap is `Google Satellite`. [Additional basemaps](https://github.com/giswqs/geemap/blob/master/geemap/geemap.py#L13) can be added using the `Map.add_basemap()` function.
```
Map = emap.Map(center=[40,-100], zoom=4)
Map.add_basemap('ROADMAP') # Add Google Map
Map
```
## Add Earth Engine Python script
```
# Add Earth Engine dataset
image = ee.Image('srtm90_v4')
# path = image.getDownloadUrl({
# 'scale': 30,
# 'crs': 'EPSG:4326',
# 'region': '[[-120, 35], [-119, 35], [-119, 34], [-120, 34]]'
# })
vis_params = {'min': 0, 'max': 3000}
Map.addLayer(image, vis_params, 'SRTM')
```
## Display Earth Engine data layers
```
Map.addLayerControl() # This line is not needed for ipyleaflet-based Map.
Map
```
| github_jupyter |
# Multi-Resolution Tutorial
This tutorial shows how to model sources frome images observed with different telescopes. We will use a multiband observation with the Hyper-Sprime Cam (HSC) and a single high-resolution image from the Hubble Space Telescope (HST).
```
# Import Packages and setup
import numpy as np
import scarlet
import astropy.io.fits as fits
from astropy.wcs import WCS
from scarlet.display import AsinhMapping
%matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
# use a better colormap and don't interpolate the pixels
matplotlib.rc('image', cmap='gray', interpolation='none', origin='lower')
```
## Load Data
We first load the HSC and HST images, channel names, and PSFs. For the images, we need to swap the byte order if necessary because a bug in astropy does not respect the local endianness... We also don't have precomputed weight/variance maps, so we will need to compute them afterwards.
```
# Load the HSC image data
obs_hdu = fits.open('../../data/test_resampling/Cut_HSC1.fits')
data_hsc = obs_hdu[0].data.byteswap().newbyteorder()
wcs_hsc = WCS(obs_hdu[0].header)
channels_hsc = ['g','r','i','z','y']
# Load the HSC PSF data
psf_hsc = fits.open('../../data/test_resampling/PSF_HSC.fits')[0].data
Np1, Np2 = psf_hsc[0].shape
psf_hsc = scarlet.ImagePSF(psf_hsc)
# Load the HST image data
hst_hdu = fits.open('../../data/test_resampling/Cut_HST1.fits')
data_hst = hst_hdu[0].data
wcs_hst = WCS(hst_hdu[0].header)
channels_hst = ['F814W']
# Load the HST PSF data
psf_hst = fits.open('../../data/test_resampling/PSF_HST.fits')[0].data
psf_hst = psf_hst[None,:,:]
psf_hst = scarlet.ImagePSF(psf_hst)
# Scale the HST data
n1,n2 = np.shape(data_hst)
data_hst = data_hst.reshape(1, n1, n2).byteswap().newbyteorder()
data_hst *= data_hsc.max() / data_hst.max()
r, N1, N2 = data_hsc.shape
```
## Create Frame and Observations
Unlike the single resolution examples, we now have two different instruments with different pixel resolutions, so we need two different observations. Since the HST image is at a much higher resolution, we define our model `Frame` to use the HST PSF and the HST resolution. Because there is no resampling between the model frame and the HST observation, we can use the default `Observation` class for the HST data. The HSC images have lower resolution, so we need to resample the models to this frame, and that's done by `LowResObservation`.
Users can specify `Frame`, `Observation` and `LowResObservation` instances by hand and match them as is usually done in single observation fitting. Alternatively, the user can provide a list of observation (no matter what the resolution of each observation is), from which the `from_observations` method will decide how large the model frame has to be and which observation(s) should be a `LowResObservation`.
```
# define two observation packages and match to frame
obs_hst = scarlet.Observation(data_hst,
wcs=wcs_hst,
psf=psf_hst,
channels=channels_hst,
weights=None)
obs_hsc = scarlet.Observation(data_hsc,
wcs=wcs_hsc,
psf=psf_hsc,
channels=channels_hsc,
weights=None)
observations = [obs_hsc, obs_hst]
model_psf = scarlet.GaussianPSF(sigma=0.6)
model_frame = scarlet.Frame.from_observations(observations, coverage='intersection', model_psf=model_psf)
obs_hsc, obs_hst = observations
```
Next we have to create a source catalog for the images. We'll use `sep` for that, but any other detection method will do. Since HST is higher resolution and less affected by blending, we use it for detection but we also run detection on the HSC image to calculate the background RMS:
```
import sep
def makeCatalog(obs_lr, obs_hr, lvl = 3, wave = True):
# Create a catalog of detected source by running SEP on the wavelet transform
# of the sum of the high resolution images and the low resolution images interpolated to the high resolution grid
#Interpolate LR to HR
interp = scarlet.interpolation.interpolate_observation(obs_lr, obs_hr)
# Normalisation
interp = interp/np.sum(interp, axis = (1,2))[:,None, None]
hr_images = obs_hr.data/np.sum(obs_hr.data, axis = (1,2))[:,None, None]
# Summation to create a detection image
detect_image = np.sum(interp, axis = 0) + np.sum(hr_images, axis = 0)
# Rescaling to HR image flux
detect_image *= np.sum(obs_hr.data)
# Wavelet transform
wave_detect = scarlet.Starlet.from_image(detect_image).coefficients
if wave:
# Creates detection from the first 3 wavelet levels
detect = wave_detect[:lvl,:,:].sum(axis = 0)
else:
detect = detect_image
# Runs SEP detection
bkg = sep.Background(detect)
catalog = sep.extract(detect, 3, err=bkg.globalrms)
bg_rms = []
for img in [obs_lr.data, obs_hr.data]:
if np.size(img.shape) == 3:
bg_rms.append(np.array([sep.Background(band).globalrms for band in img]))
else:
bg_rms.append(sep.Background(img).globalrms)
return catalog, bg_rms, detect_image
# Making catalog.
# With the wavelet option on, only the first 3 wavelet levels are used for detection. Set to 1 for better detection
wave = 1
lvl = 3
catalog_hst, (bg_hsc, bg_hst), detect = makeCatalog(obs_hsc, obs_hst, lvl, wave)
# we can now set the empirical noise rms for both observations
obs_hsc.weights = np.ones(obs_hsc.shape) / (bg_hsc**2)[:, None, None]
obs_hst.weights = np.ones(obs_hst.shape) / (bg_hst**2)[:, None, None]
```
Finally we can visualize the detections for the multi-band HSC and single-band HST images in their native resolutions:
```
# Create a color mapping for the HSC image
norm_hsc = AsinhMapping(minimum=-1, stretch=5, Q=3)
norm_hst = AsinhMapping(minimum=-1, stretch=5, Q=3)
norms = [norm_hsc, norm_hst]
# Get the source coordinates from the HST catalog
pixel_hst = np.stack((catalog_hst['y'], catalog_hst['x']), axis=1)
# Convert the HST coordinates to the HSC WCS
ra_dec = obs_hst.get_sky_coord(pixel_hst)
for obs, norm in zip(observations, norms):
scarlet.display.show_observation(obs, norm=norm, sky_coords=ra_dec, show_psf=True)
plt.show()
```
## Initialize Sources and Blend
We expect all sources to be galaxies, so we initialized them as `ExtendedSources`.
Afterwards, we match their amplitudes to the data, and create an instance of `Blend` to hold all sources and *all* observations for the fit below.
```
# Source initialisation
sources = [
scarlet.ExtendedSource(model_frame,
sky_coord,
observations,
thresh=0.1,
)
for sky_coord in ra_dec
]
scarlet.initialization.set_spectra_to_match(sources, observations)
blend = scarlet.Blend(sources, observations)
```
## Display Initial guess
Let's compare the initial guess in both observation frames. Note that the full model comprises more spectral channels and/or pixels than any individual observation. That's a result of defining
```
for i in range(len(observations)):
scarlet.display.show_scene(sources,
norm=norms[i],
observation=observations[i],
show_model=False,
show_rendered=True,
show_observed=True,
show_residual=True,
figsize=(12,4)
)
plt.show()
```
## Fit Model
```
%time it, logL = blend.fit(50, e_rel=1e-4)
print(f"scarlet ran for {it} iterations to logL = {logL}")
scarlet.display.show_likelihood(blend)
plt.show()
```
### View Updated Model
We use the same principle to look at the updated model:
```
for i in range(len(observations)):
scarlet.display.show_scene(sources,
norm=norms[i],
observation=observations[i],
show_model=False,
show_rendered=True,
show_observed=True,
show_residual=True,
figsize=(12,4)
)
plt.show()
```
### View Source Models
It can also be useful to view the model for each source. For each source we extract the portion of the image contained in the sources bounding box, the true simulated source flux, and the model of the source, scaled so that all of the images have roughly the same pixel scale.
```
for k in range(len(sources)):
print('source number ', k)
for i in range(len(observations)):
scarlet.display.show_sources((sources[k],),
norm=norm_hst,
observation=observations[i],
show_model=False,
show_rendered=True,
show_observed=True,
show_spectrum=False,
add_boxes=True,
figsize=(8,4)
)
plt.show()
```
| github_jupyter |
# MNIST Fashion Training and Validation
MNIST Dataset contains blurred `images` of clothes and their true labels. The goal of this project is to build a model that correctly identifies the number inputs by implementing a validation process.
First, download and load the training data.
```
import torch
from torchvision import datasets, transforms
# Define a transform to normalize the data
transform = transforms.Compose([transforms.ToTensor(),
transforms.Normalize((0.5,), (0.5,))])
# Download and load the training data
trainset = datasets.FashionMNIST('~/.pytorch/F_MNIST_data/', download=True, train=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=64, shuffle=True)
# Download and load the test data
testset = datasets.FashionMNIST('~/.pytorch/F_MNIST_data/', download=True, train=False, transform=transform)
testloader = torch.utils.data.DataLoader(testset, batch_size=64, shuffle=True)
# Images and labels constantly renewing their values from testloader
images, labels = next(iter(testloader))
```
Next, the network was outlined with three hidden layers and one output layer. `self.dropout` was added to decrease the amount of validation loss to prevent overfitting.
```
from torch import nn, optim
import torch.nn.functional as F
# <Model Setup>
class Classifier(nn.Module):
def __init__(self):
super().__init__()
self.fc1 = nn.Linear(784, 256)
self.fc2 = nn.Linear(256, 128)
self.fc3 = nn.Linear(128, 64)
self.fc4 = nn.Linear(64, 10)
# Dropout module with 0.2 drop probability
self.dropout = nn.Dropout(p=0.2)
def forward(self, x):
# make sure input tensor is flattened
x = x.view(x.shape[0], -1)
# Now with dropout
x = self.dropout(F.relu(self.fc1(x)))
x = self.dropout(F.relu(self.fc2(x)))
x = self.dropout(F.relu(self.fc3(x)))
# output so no dropout here
x = F.log_softmax(self.fc4(x), dim=1)
return x
```
Then, the model was trained over 30 epochs.
```
# <network generation>
model = Classifier()
criterion = nn.NLLLoss() # criterion for loss
optimizer = optim.Adam(model.parameters(), lr=0.003) # optimizer for update
train_losses, test_losses = [], [] # list to appends new losses
epochs = 30
# <network processing>
for e in range(epochs):
# <training>
train_loss_sum = 0 # train loss sum at one epoch
for images, labels in trainloader:
optimizer.zero_grad() # clears gradient descent
output = model(images) # train output
train_loss = criterion(output, labels) # train loss calculation
train_loss.backward() # backpropagation
optimizer.step() # updates model
train_loss_sum += train_loss.item()
# <testing>
else:
test_loss_sum = 0 # test loss sum at one epoch
accuracy_sum = 0 # accuracy sum at one epoch
with torch.no_grad(): # remove descents
model.eval() # converts to test mode
for images, labels in testloader:
output = model(images) # test output
test_loss = criterion(output, labels) # test loss calculation
test_loss_sum += test_loss.item()
ps = torch.exp(output) # probability
top_p, top_class = ps.topk(1, dim=1) # classes with highest probability
equals = top_class == labels.view(*top_class.shape) # test classes vs. label
accuracy = torch.mean(equals.type(torch.FloatTensor)) # frequency of test classes == label
accuracy_sum += accuracy
model.train() # reverts to train mode
train_losses.append(train_loss_sum/len(trainloader)) # appends average train loss
test_losses.append(test_loss/len(testloader)) # appends average test loss
print("Epoch: {}/{}.. ".format(e+1, epochs),
"Training Loss: {:.3f}.. ".format(train_losses[-1]), # newest average train loss
"Test Loss: {:.3f}.. ".format(test_losses[-1]), # newest average test loss
"Test Accuracy: {:.3f}".format(accuracy_sum/len(testloader))) # average accuracy
```
Now, the trained model is ready for inference.
```
import helper
# <inference>
model.eval() # converts to test mode
dataiter = iter(testloader)
images, labels = dataiter.next()
img = images[0]
img = img.view(1, 784) # flattens image
with torch.no_grad():
output = model.forward(img) # inputs image
ps = torch.exp(output) # prabability
# <plot the image and probabilities>
helper.view_classify(img.view(1, 28, 28), ps, version='Fashion')
```
After running the evaluation several times, it is observed that the model was able to plot most of the test inputs while still struggling to recognize difficult ones. Bigger training sets may be required.
| github_jupyter |
# Fastpages Notebook Blog Post
> A tutorial of fastpages for Jupyter notebooks.
- toc: true
- badges: true
- comments: true
- categories: [jupyter]
- image: images/chart-preview.png
# About
This notebook is a demonstration of some of capabilities of [fastpages](https://github.com/fastai/fastpages) with notebooks.
With `fastpages` you can save your jupyter notebooks into the `_notebooks` folder at the root of your repository, and they will be automatically be converted to Jekyll compliant blog posts!
## Front Matter
Front Matter is a markdown cell at the beginning of your notebook that allows you to inject metadata into your notebook. For example:
- Setting `toc: true` will automatically generate a table of contents
- Setting `badges: true` will automatically include GitHub and Google Colab links to your notebook.
- Setting `comments: true` will enable commenting on your blog post, powered by [utterances](https://github.com/utterance/utterances).
More details and options for front matter can be viewed on the [front matter section](https://github.com/fastai/fastpages#front-matter-related-options) of the README.
## Markdown Shortcuts
put a `#hide` flag at the top of any cell you want to completely hide in the docs
put a `#collapse-hide` flag at the top of any cell if you want to **hide** that cell by default, but give the reader the option to show it:
```
#collapse-hide
import pandas as pd
import altair as alt
```
put a `#collapse-show` flag at the top of any cell if you want to **show** that cell by default, but give the reader the option to hide it:
```
#collapse-show
cars = 'https://vega.github.io/vega-datasets/data/cars.json'
movies = 'https://vega.github.io/vega-datasets/data/movies.json'
sp500 = 'https://vega.github.io/vega-datasets/data/sp500.csv'
stocks = 'https://vega.github.io/vega-datasets/data/stocks.csv'
flights = 'https://vega.github.io/vega-datasets/data/flights-5k.json'
```
## Interactive Charts With Altair
Charts made with Altair remain interactive. Example charts taken from [this repo](https://github.com/uwdata/visualization-curriculum), specifically [this notebook](https://github.com/uwdata/visualization-curriculum/blob/master/altair_interaction.ipynb).
```
# hide
df = pd.read_json(movies) # load movies data
genres = df['Major_Genre'].unique() # get unique field values
genres = list(filter(lambda d: d is not None, genres)) # filter out None values
genres.sort() # sort alphabetically
#hide
mpaa = ['G', 'PG', 'PG-13', 'R', 'NC-17', 'Not Rated']
```
### Example 1: DropDown
```
# single-value selection over [Major_Genre, MPAA_Rating] pairs
# use specific hard-wired values as the initial selected values
selection = alt.selection_single(
name='Select',
fields=['Major_Genre', 'MPAA_Rating'],
init={'Major_Genre': 'Drama', 'MPAA_Rating': 'R'},
bind={'Major_Genre': alt.binding_select(options=genres), 'MPAA_Rating': alt.binding_radio(options=mpaa)}
)
# scatter plot, modify opacity based on selection
alt.Chart(movies).mark_circle().add_selection(
selection
).encode(
x='Rotten_Tomatoes_Rating:Q',
y='IMDB_Rating:Q',
tooltip='Title:N',
opacity=alt.condition(selection, alt.value(0.75), alt.value(0.05))
)
```
### Example 2: Tooltips
```
alt.Chart(movies).mark_circle().add_selection(
alt.selection_interval(bind='scales', encodings=['x'])
).encode(
x='Rotten_Tomatoes_Rating:Q',
y=alt.Y('IMDB_Rating:Q', axis=alt.Axis(minExtent=30)), # use min extent to stabilize axis title placement
tooltip=['Title:N', 'Release_Date:N', 'IMDB_Rating:Q', 'Rotten_Tomatoes_Rating:Q']
).properties(
width=600,
height=400
)
```
### Example 3: More Tooltips
```
# select a point for which to provide details-on-demand
label = alt.selection_single(
encodings=['x'], # limit selection to x-axis value
on='mouseover', # select on mouseover events
nearest=True, # select data point nearest the cursor
empty='none' # empty selection includes no data points
)
# define our base line chart of stock prices
base = alt.Chart().mark_line().encode(
alt.X('date:T'),
alt.Y('price:Q', scale=alt.Scale(type='log')),
alt.Color('symbol:N')
)
alt.layer(
base, # base line chart
# add a rule mark to serve as a guide line
alt.Chart().mark_rule(color='#aaa').encode(
x='date:T'
).transform_filter(label),
# add circle marks for selected time points, hide unselected points
base.mark_circle().encode(
opacity=alt.condition(label, alt.value(1), alt.value(0))
).add_selection(label),
# add white stroked text to provide a legible background for labels
base.mark_text(align='left', dx=5, dy=-5, stroke='white', strokeWidth=2).encode(
text='price:Q'
).transform_filter(label),
# add text labels for stock prices
base.mark_text(align='left', dx=5, dy=-5).encode(
text='price:Q'
).transform_filter(label),
data=stocks
).properties(
width=700,
height=400
)
```
## Data Tables
You can display tables per the usual way in your blog:
```
movies = 'https://vega.github.io/vega-datasets/data/movies.json'
df = pd.read_json(movies)
# display table with pandas
df[['Title', 'Worldwide_Gross',
'Production_Budget', 'IMDB_Rating']].head()
```
## Images
### Local Images
You can reference local images and they will be copied and rendered on your blog automatically. You can include these with the following markdown syntax:
``

### Remote Images
Remote images can be included with the following markdown syntax:
``

### Animated Gifs
Animated Gifs work, too!
`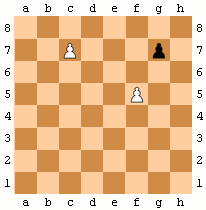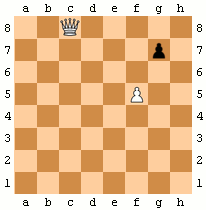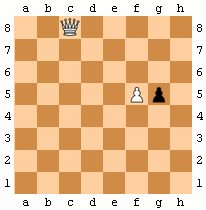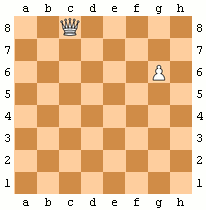`

### Captions
You can include captions with markdown images like this:
```

```

# Other Elements
## Tweetcards
Typing `> twitter: https://twitter.com/jakevdp/status/1204765621767901185?s=20` will render this:
> twitter: https://twitter.com/jakevdp/status/1204765621767901185?s=20
## Youtube Videos
Typing `> youtube: https://youtu.be/XfoYk_Z5AkI` will render this:
> youtube: https://youtu.be/XfoYk_Z5AkI
## Boxes / Callouts
Typing `> Warning: There will be no second warning!` will render this:
> Warning: There will be no second warning!
Typing `> Important: Pay attention! It's important.` will render this:
> Important: Pay attention! It's important.
Typing `> Tip: This is my tip.` will render this:
> Tip: This is my tip.
Typing `> Note: Take note of this.` will render this:
> Note: Take note of this.
Typing `> Note: A doc link to [an example website: fast.ai](https://www.fast.ai/) should also work fine.` will render in the docs:
> Note: A doc link to [an example website: fast.ai](https://www.fast.ai/) should also work fine.
## Footnotes
You can have footnotes in notebooks just like you can with markdown.
For example, here is a footnote [^1].
[^1]: This is the footnote.
| github_jupyter |
[View in Colaboratory](https://colab.research.google.com/github/findingfoot/ML_practice-codes/blob/master/Different_layers_in_Neural_Network_1D_and_2D_emulated_data.ipynb)
Info:
We will be dealing with different kinds of layers in Neural Networks
1. Convolution Layer
2. Activation Layer
3. Max Pool layer
4. Full connected layer
Kind of datasets used:
1. 1D data emulating row wise data
2. 2D data emulating images
```
import warnings
warnings.filterwarnings('ignore')
import tensorflow as tf
from tensorflow.python.framework import ops
import numpy as np
import matplotlib.pyplot as plt
import random
import csv
import os
ops.reset_default_graph()
sess = tf.Session()
#conv layer stuff
size_of_data = 25
conv_filter_size = 5
maxpool_size = 5
stride = 1
#reproducibility
seed = 23
np.random.seed(seed)
tf.set_random_seed(seed)
#Begin with 1D data
data_1d = np.random.normal(size=size_of_data)
#now create a placeholder
x_input_1d = tf.placeholder(dtype=tf.float32, shape=[size_of_data])
#create the convolution
def conv_layer_1d(input_1d, my_filter, stride):
input_2d = tf.expand_dims(input_1d, 0)
input_3d = tf.expand_dims(input_2d, 0)
input_4d = tf.expand_dims(input_3d, 3)
conv_output = tf.nn.conv2d(input_4d, filter = my_filter, strides = [1,1,stride,1], padding = 'VALID')
#removing the added dimension
conv_output_1d = tf.squeeze(conv_output)
return(conv_output_1d)
my_filter = tf.Variable(tf.random_normal(shape = [1,conv_filter_size,1,1]))
my_convolution_output = conv_layer_1d(x_input_1d,my_filter, stride)
#activation function
def activation(input_1d):
return(tf.nn.relu(input_1d))
#creating activation layer
my_activation_output = activation(my_convolution_output)
#max pooling layer
def max_pool(input_1d, width, stride):
input_2d = tf.expand_dims(input_1d, 0)
input_3d = tf.expand_dims(input_2d, 0)
input_4d = tf.expand_dims(input_3d, 3)
#specify the window size
pool_output = tf.nn.max_pool(input_4d,
ksize=[1,1,width,1],
strides= [1,1,stride,1],
padding = "VALID")
#removing extra dimension
pool_output_1d = tf.squeeze(pool_output)
return(pool_output_1d)
my_max_pool_output = max_pool(my_activation_output, width = maxpool_size, stride = stride)
#fully connected layer
def fully_connected(input_layer, num_outputs):
#shape of multiplication weight matrix
print('shape of input layer',(num_outputs))
weight_shape = tf.squeeze(tf.stack([tf.shape(input_layer), [num_outputs]]))
#initialize weight
weight = tf.random_normal(weight_shape, stddev=0.1)
#initialize the bias
bias = tf.random_normal(shape=[num_outputs])
#converting 1d array into 2d array
input_layer_2d = tf.expand_dims(input_layer, 0)
#do the matrix multiplication and add bias
full_output = tf.add(tf.matmul(input_layer_2d, weight), bias)
#squeeze out the extra dimension
full_output_1d = tf.squeeze(full_output)
return (full_output_1d)
my_full_output = fully_connected(my_max_pool_output, 5)
#lets run the graph and see
init = tf.global_variables_initializer()
sess.run(init)
feed_dict = {x_input_1d: data_1d}
print('++++++++++++++++ Running 1D data ++++++++++++++++++++++ \n')
print('Input = array of length %d ' %(x_input_1d.shape.as_list()[0]))
print('convolution with filter, length = %d, stride size = %d, results in an array of length %d : \n'
% (conv_filter_size, stride, my_convolution_output.shape.as_list()[0]))
print(sess.run(my_convolution_output, feed_dict=feed_dict))
print('\n\n+++++++++++ Activation Block +++++++++++++++++')
print('\n Input = above array of length %d' % (my_convolution_output.shape.as_list()[0]))
print('\n Relu activation function returns an array of length %d' %(my_activation_output.shape.as_list()[0]))
print(sess.run(my_activation_output, feed_dict=feed_dict))
print('++++++++++++++++++++++ Max Pool layer +++++++++++++++++++')
print('\n Input = above array of length %d' % (my_activation_output.shape.as_list()[0]))
print('Max Pool, window length = %d, stride size = %d, results in an array of length %d : \n'
% (maxpool_size,stride, my_max_pool_output.shape.as_list()[0]))
print(sess.run(my_max_pool_output, feed_dict=feed_dict))
print('\n++++++++++++++++++++++ Fully Connected Layer ++++++++++++++++++')
print('\n Input = above array of length %d' % (my_max_pool_output.shape.as_list()[0]))
print('Fully connected layer on all 4 rows with %d outputs:' %
(my_full_output.shape.as_list()[0]))
print(sess.run(my_full_output, feed_dict=feed_dict))
# Lets deal with 2D data
ops.reset_default_graph()
sess = tf.Session()
#parameters for the run
row_size = 10
col_size = 10
conv_size = 2
conv_stride_size = 2
maxpool_size = 2
maxpool_stride_size = 1
seed = 14
np.random.seed(seed)
tf.set_random_seed(seed)
data_size = [row_size, col_size]
data_2d = np.random.normal(size = data_size)
x_input_2d = tf.placeholder(dtype = tf.float32, shape=data_size)
#convolution layer for 2d
def conv_layer_2d(input_2d, my_filter, stride_size):
input_3d = tf.expand_dims(input_2d, 0)
input_4d = tf.expand_dims(input_3d, 3)
convolution_output = tf.nn.conv2d(input_4d, filter = my_filter,
strides = [1, stride_size, stride_size, 1]
, padding = "VALID")
#removing extra dimenstions
conv_output_2d = tf.squeeze(convolution_output)
return(conv_output_2d)
my_filter = tf.random_normal(shape = [conv_size,conv_size,1,1], dtype = tf.float32)
my_convolution_output = conv_layer_2d(x_input_2d, my_filter=my_filter, stride_size=conv_stride_size)
#Activation function
def activation(input_1d):
return(tf.nn.relu(input_1d))
my_activation_output = activation(my_convolution_output)
#max pool layer
def max_pool(input_2d, width, height, stride):
input_3d = tf.expand_dims(input_2d, 0)
input_4d = tf.expand_dims(input_3d, 3)
pool_output = tf.nn.max_pool(input_4d, ksize=[1, height, width, 1],
strides = [1, stride, stride, 1],
padding = "VALID")
pool_output_2d = tf.squeeze(pool_output)
return(pool_output_2d)
my_maxpool_output = max_pool(my_activation_output, width=maxpool_size, height = maxpool_size,
stride = maxpool_stride_size)
#fully connected layer
def fully_connected(input_layer, num_outputs):
flat_input = tf.reshape(input_layer, [-1])
weight_shape = tf.squeeze(tf.stack([tf.shape(flat_input),[num_outputs]]))
weight = tf.random_normal(weight_shape, stddev = 0.1)
bias = tf.random_normal(shape = [num_outputs])
input_2d = tf.expand_dims(flat_input, 0)
full_output = tf.add(tf.matmul(input_2d, weight), bias)
full_output_2d = tf.squeeze(full_output)
return(full_output_2d)
my_full_output = fully_connected(my_maxpool_output, 5)
init = tf.global_variables_initializer()
sess.run(init)
feed_dict = {x_input_2d: data_2d}
print('>>>> 2D Data <<<<')
# Convolution Output
print('Input = %s array' % (x_input_2d.shape.as_list()))
print('%s Convolution, stride size = [%d, %d] , results in the %s array' %
(my_filter.get_shape().as_list()[:2],conv_stride_size,conv_stride_size,my_convolution_output.shape.as_list()))
print(sess.run(my_convolution_output, feed_dict=feed_dict))
# Activation Output
print('\nInput = the above %s array' % (my_convolution_output.shape.as_list()))
print('ReLU element wise returns the %s array' % (my_activation_output.shape.as_list()))
print(sess.run(my_activation_output, feed_dict=feed_dict))
# Max Pool Output
print('\nInput = the above %s array' % (my_activation_output.shape.as_list()))
print('MaxPool, stride size = [%d, %d], results in %s array' %
(maxpool_stride_size,maxpool_stride_size,my_maxpool_output.shape.as_list()))
print(sess.run(my_maxpool_output, feed_dict=feed_dict))
# Fully Connected Output
print('\nInput = the above %s array' % (my_maxpool_output.shape.as_list()))
print('Fully connected layer on all %d rows results in %s outputs:' %
(my_maxpool_output.shape.as_list()[0],my_full_output.shape.as_list()[0]))
print(sess.run(my_full_output, feed_dict=feed_dict))
```
| github_jupyter |
```
import re
import os
from pprint import pprint
import numpy as np
from sklearn.model_selection import train_test_split
import torch
import torch.autograd as autograd
import torch.nn as nn
from torch.nn import CrossEntropyLoss, MSELoss
from torch import Tensor
from transformers import AlbertModel, BertTokenizer
from transformers import AdamW
from torch.utils.data import TensorDataset, DataLoader, RandomSampler, SequentialSampler
from tqdm.notebook import tqdm
from tensorflow.keras.preprocessing.sequence import pad_sequences # padding
from pytorchcrf import CRF
file = "dh_msra.txt"
class Config(object):
"""配置参数"""
def __init__(self):
current_path = os.getcwd()
self.model_name = "pytorch_model.bin"
self.bert_path = os.path.join(current_path + "/albert_chinese_tiny")
# self.train_file = '../datas/THUCNews/train.txt'
self.num_classes = 10 # NER 的 token 类别
self.hidden_size = 312 # 隐藏层输出维度
self.hidden_dropout_prob = 0.1 # dropout比例
self.batch_size = 64 # mini-batch大小
self.max_len = 128 # 句子的最长padding长度
self.epochs = 3 # epoch数
self.learning_rate = 2e-5 # 学习率
self.save_path = os.path.join(current_path + "/finetuned_albert") # 模型训练结果保存路径
self.use_cuda = True
self.device_id = 5
config = Config()
print(config.bert_path)
# GPUcheck
print("CUDA Available: ", torch.cuda.is_available())
n_gpu = torch.cuda.device_count()
if torch.cuda.is_available() and config.use_cuda:
print("GPU numbers: ", n_gpu)
print("device_name: ", torch.cuda.get_device_name(0))
device_id = config.device_id # 注意选择
torch.cuda.set_device(device_id)
device = torch.device(f"cuda:{device_id}")
print(f"当前设备:{torch.cuda.current_device()}")
else :
device = torch.device("cpu")
print(f"当前设备:{device}")
all_sentences_separate = []
all_letter_labels = []
label_set = set()
with open(file, encoding="utf-8") as f:
single_sentence = []
single_sentence_labels = []
for s in f.readlines():
if s != "\n":
word, label = s.split("\t")
label = label.strip("\n")
single_sentence.append(word)
single_sentence_labels.append(label)
label_set.add(label)
elif s == "\n":
all_sentences_separate.append(single_sentence)
all_letter_labels.append(single_sentence_labels)
single_sentence = []
single_sentence_labels = []
print(all_sentences_separate[0:2])
print(all_letter_labels[0:2])
print(f"\n所有的标签:{label_set}")
# 构建 tag 到 索引 的字典
tag_to_ix = {"B-LOC": 0,
"I-LOC": 1,
"B-ORG": 2,
"I-ORG": 3,
"B-PER": 4,
"I-PER": 5,
"O": 6,
"[CLS]":7,
"[SEP]":8,
"[PAD]":9}
ix_to_tag = {0:"B-LOC",
1:"I-LOC",
2:"B-ORG",
3:"I-ORG",
4:"B-PER",
5:"I-PER",
6:"O",
7:"[CLS]",
8:"[SEP]",
9:"[PAD]"}
all_sentences = [] # 句子
for one_sentence in all_sentences_separate:
sentence = "".join(one_sentence)
all_sentences.append(sentence)
pprint(all_sentences[15:20])
all_labels = [] # labels
for letter_labels in all_letter_labels:
labels = [tag_to_ix[t] for t in letter_labels]
all_labels.append(labels)
print(all_labels[0:2])
print(len(all_labels[0]))
# word2token
tokenizer = BertTokenizer.from_pretrained(config.bert_path, do_lower_case=True)
tokenized_texts = [tokenizer.encode(sent, add_special_tokens=True) for sent in all_sentences]
# 句子padding
# 输入padding
# 此函数在keras里面
input_ids = pad_sequences([txt for txt in tokenized_texts],
maxlen=config.max_len,
dtype="long",
truncating="post",
padding="post")
# [3] 代表 Other 实体
for label in all_labels:
label.insert(len(label), 8) # [SEP] 加在末尾
label.insert(0, 7) # [CLS] 加在开头
if config.max_len > len(label) -1:
for i in range(config.max_len - len(label)):
label.append(9) # [PAD]
# 创建attention masks
attention_masks = []
# Create a mask of 1s for each token followed by 0s for padding
for seq in input_ids:
seq_mask = [float(i > 0) for i in seq]
attention_masks.append(seq_mask)
# train-test-split
train_inputs, validation_inputs, train_labels, validation_labels = train_test_split(input_ids,
all_labels,
random_state=2019,
test_size=0.1)
train_masks, validation_masks, _, _ = train_test_split(attention_masks,
input_ids,
random_state=2019,
test_size=0.1)
# tensor化
train_inputs = torch.tensor(train_inputs)
validation_inputs = torch.tensor(validation_inputs)
train_labels = torch.tensor(train_labels)
validation_labels = torch.tensor(validation_labels)
train_masks = torch.tensor(train_masks)
validation_masks = torch.tensor(validation_masks)
# dataloader
# 形成训练数据集
train_data = TensorDataset(train_inputs, train_masks, train_labels)
# 随机采样
train_sampler = RandomSampler(train_data)
# 读取数据
train_dataloader = DataLoader(train_data, sampler=train_sampler, batch_size=config.batch_size)
# 形成验证数据集
validation_data = TensorDataset(validation_inputs, validation_masks, validation_labels)
# 随机采样
validation_sampler = SequentialSampler(validation_data)
# 读取数据
validation_dataloader = DataLoader(validation_data, sampler=validation_sampler, batch_size=config.batch_size)
class ModelAlBert(nn.Module):
"""
新增 句子位置ID 特征拼接。
为了保证速度,选择基于预训练的 Albert-tiny 微调
我们想要对裁判文书进行分类,原文的文本形式,会进入BERT模型。
代码中的“HYID”本身就是数字形式,没必要放入BERT模型中,于是我们将 BERT 输出后的 768 维向量拼接
tensor(HYID),也就是变成了 769 维,再过一个 线形层 + softmax 输出分类结果。
"""
def __init__(self, config):
super(ModelAlBert, self).__init__()
self.num_labels = config.num_classes
self.albert = AlbertModel.from_pretrained(config.bert_path)
self.dropout = nn.Dropout(config.hidden_dropout_prob)
self.classifier = nn.Linear(config.hidden_size, config.num_classes)
def forward(
self,
input_ids: Tensor = None,
attention_mask: Tensor = None,
token_type_ids=None,
position_ids=None,
head_mask=None,
inputs_embeds=None,
labels: Tensor = None,
) -> set:
"""
模型前向传播结构
Args:
input_ids (Tensor[Tensor], optional): Token 化的句子. Defaults to None.
attention_mask (Tensor[Tensor], optional): Attention Mask,配合Padding使用. Defaults to None.
token_type_ids ([type], optional): 上下句 id 标记,这里不涉及. Defaults to None.
position_ids ([type], optional): token 位置 id. Defaults to None.
head_mask ([type], optional): [description]. Defaults to None.
inputs_embeds ([type], optional): 不需要. Defaults to None.
labels (Tensor, optional): 标签. Defaults to None.
HYID (Tensor, optional): 这里指的是句子的位置ID,也可以是其他特征. Defaults to None.
Returns:
(set): 模型的返回值, (loss), logits, (hidden_states), (attentions)
"""
outputs = self.albert(
input_ids,
attention_mask=attention_mask,
token_type_ids=token_type_ids,
position_ids=position_ids,
head_mask=head_mask,
inputs_embeds=inputs_embeds,
)
sequence_output = outputs[0]
sequence_output = self.dropout(sequence_output)
logits = self.classifier(sequence_output)
loss = None
if labels is not None:
loss_fct = CrossEntropyLoss()
# Only keep active parts of the loss
if attention_mask is not None:
active_loss = attention_mask.view(-1) == 1
active_logits = logits.view(-1, self.num_labels)[active_loss]
active_labels = labels.view(-1)[active_loss]
loss = loss_fct(active_logits, active_labels)
else:
loss = loss_fct(logits.view(-1, self.num_labels), labels.view(-1))
output = (logits,) + outputs[2:]
return ((loss,) + output) if loss is not None else output
model = ModelAlBert(config)
model.cuda()
class ModelAlBertCRF(nn.Module):
"""
新增 句子位置ID 特征拼接。
为了保证速度,选择基于预训练的 Albert-tiny 微调
"""
def __init__(self, config):
super(ModelAlBertCRF, self).__init__()
self.num_labels = config.num_classes
self.albert = AlbertModel.from_pretrained(config.bert_path)
self.dropout = nn.Dropout(config.hidden_dropout_prob)
self.classifier = nn.Linear(config.hidden_size, config.num_classes)
self.crf = CRF(num_tags=config.num_classes, batch_first=True)
def forward(
self,
input_ids: Tensor = None,
attention_mask: Tensor = None,
token_type_ids=None,
position_ids=None,
head_mask=None,
inputs_embeds=None,
labels: Tensor = None,
) -> set:
"""
模型前向传播结构。
注意loss采用的是CRF的 log likelihood
Args:
input_ids (Tensor[Tensor], optional): Token 化的句子. Defaults to None.
attention_mask (Tensor[Tensor], optional): Attention Mask,配合Padding使用. Defaults to None.
token_type_ids ([type], optional): 上下句 id 标记,这里不涉及. Defaults to None.
position_ids ([type], optional): token 位置 id. Defaults to None.
head_mask ([type], optional): [description]. Defaults to None.
inputs_embeds ([type], optional): 不需要. Defaults to None.
labels (Tensor, optional): 标签. Defaults to None.
Returns:
(set): 模型的返回值, (loss), logits, (hidden_states), (attentions)
"""
outputs = self.albert(
input_ids,
attention_mask=attention_mask,
token_type_ids=token_type_ids,
position_ids=position_ids,
head_mask=head_mask,
inputs_embeds=inputs_embeds,
)
sequence_output = outputs[0]
sequence_output = self.dropout(sequence_output)
logits = self.classifier(sequence_output)
loss = None
outputs = (logits,)
if labels is not None:
loss = self.crf(emissions = logits, tags=labels, mask=attention_mask)
# Note that the returned value is the log likelihood
# so you’ll need to make this value negative as your loss.
outputs =(-1 * loss,) + outputs
return outputs # (loss), scores
model = ModelAlBertCRF(config)
model.cuda()
# BERT fine-tuning parameters
bert_param_optimizer = list(model.albert.named_parameters())
crf_param_optimizer = list(model.crf.named_parameters())
linear_param_optimizer = list(model.classifier.named_parameters())
no_decay = ['bias', 'LayerNorm.weight']
# 权重衰减
optimizer_grouped_parameters = [
{'params': [p for n, p in bert_param_optimizer if not any(nd in n for nd in no_decay)],
'weight_decay': 0.01,
'lr': config.learning_rate},
{'params': [p for n, p in bert_param_optimizer if any(nd in n for nd in no_decay)],
'weight_decay': 0.0,
'lr': config.learning_rate},
{'params': [p for n, p in crf_param_optimizer if not any(nd in n for nd in no_decay)],
'weight_decay': 0.01,
'lr': config.crf_learning_rate},
{'params': [p for n, p in crf_param_optimizer if any(nd in n for nd in no_decay)],
'weight_decay': 0.0,
'lr': config.crf_learning_rate},
{'params': [p for n, p in linear_param_optimizer if not any(nd in n for nd in no_decay)],
'weight_decay': 0.01,
'lr': config.crf_learning_rate},
{'params': [p for n, p in linear_param_optimizer if any(nd in n for nd in no_decay)],
'weight_decay': 0.0,
'lr': config.crf_learning_rate}
]
# 优化器
optimizer = AdamW(optimizer_grouped_parameters,
lr=config.learning_rate)
# 保存loss
train_loss_set = []
# BERT training loop
for _ in range(config.epochs):
## 训练
print(f"当前epoch: {_}")
# 开启训练模式
model.train()
tr_loss = 0 # train loss
nb_tr_examples, nb_tr_steps = 0, 0
# Train the data for one epoch
for step, batch in tqdm(enumerate(train_dataloader)):
# 把batch放入GPU
batch = tuple(t.to(device) for t in batch)
# 解包batch
b_input_ids, b_input_mask, b_labels = batch
# 梯度归零
optimizer.zero_grad()
# 前向传播loss计算
output = model(input_ids=b_input_ids,
attention_mask=b_input_mask,
labels=b_labels)
loss = output[0]
# print(loss)
# 反向传播
loss.backward()
# Update parameters and take a step using the computed gradient
# 更新模型参数
optimizer.step()
# Update tracking variables
tr_loss += loss.item()
nb_tr_examples += b_input_ids.size(0)
nb_tr_steps += 1
print(f"当前 epoch 的 Train loss: {tr_loss/nb_tr_steps}")
# 验证状态
model.eval()
# 建立变量
eval_loss, eval_accuracy = 0, 0
nb_eval_steps, nb_eval_examples = 0, 0
# Evaluate data for one epoch
# 验证集的读取也要batch
for batch in tqdm(validation_dataloader):
# 元组打包放进GPU
batch = tuple(t.to(device) for t in batch)
# 解开元组
b_input_ids, b_input_mask, b_labels = batch
# 预测
with torch.no_grad():
# segment embeddings,如果没有就是全0,表示单句
# position embeddings,[0,句子长度-1]
outputs = model(input_ids=b_input_ids,
attention_mask=b_input_mask,
token_type_ids=None,
position_ids=None)
# print(logits[0])
# Move logits and labels to CPU
scores = outputs[0].detach().cpu().numpy() # 每个字的标签的概率
pred_flat = np.argmax(scores[0], axis=1).flatten()
label_ids = b_labels.to('cpu').numpy() # 真实labels
# print(logits, label_ids)
```
| github_jupyter |
# Testing
Think Bayes, Second Edition
Copyright 2020 Allen B. Downey
License: [Attribution-NonCommercial-ShareAlike 4.0 International (CC BY-NC-SA 4.0)](https://creativecommons.org/licenses/by-nc-sa/4.0/)
```
# If we're running on Colab, install empiricaldist
# https://pypi.org/project/empiricaldist/
import sys
IN_COLAB = 'google.colab' in sys.modules
if IN_COLAB:
!pip install empiricaldist
# Get utils.py
import os
if not os.path.exists('utils.py'):
!wget https://github.com/AllenDowney/ThinkBayes2/raw/master/soln/utils.py
from utils import set_pyplot_params
set_pyplot_params()
```
In <<_TheEuroProblem>> I presented a problem from David MacKay's book, [*Information Theory, Inference, and Learning Algorithms*](http://www.inference.org.uk/mackay/itila/p0.html):
"A statistical statement appeared in *The Guardian* on Friday January 4, 2002:
> When spun on edge 250 times, a Belgian one-euro coin came up heads 140 times and tails 110. \`It looks very suspicious to me,' said Barry Blight, a statistics lecturer at the London School of Economics. \`If the coin were unbiased, the chance of getting a result as extreme as that would be less than 7%.'
"But [MacKay asks] do these data give evidence that the coin is biased rather than fair?"
We started to answer this question in <<_EstimatingProportions>>; to review, our answer was based on these modeling decisions:
* If you spin a coin on edge, there is some probability, $x$, that it will land heads up.
* The value of $x$ varies from one coin to the next, depending on how the coin is balanced and possibly other factors.
Starting with a uniform prior distribution for $x$, we updated it with the given data, 140 heads and 110 tails. Then we used the posterior distribution to compute the most likely value of $x$, the posterior mean, and a credible interval.
But we never really answered MacKay's question: "Do these data give evidence that the coin is biased rather than fair?"
In this chapter, finally, we will.
## Estimation
Let's review the solution to the Euro problem from <<_TheBinomialLikelihoodFunction>>. We started with a uniform prior.
```
import numpy as np
from empiricaldist import Pmf
xs = np.linspace(0, 1, 101)
uniform = Pmf(1, xs)
```
And we used the binomial distribution to compute the probability of the data for each possible value of $x$.
```
from scipy.stats import binom
k, n = 140, 250
likelihood = binom.pmf(k, n, xs)
```
We computed the posterior distribution in the usual way.
```
posterior = uniform * likelihood
posterior.normalize()
```
And here's what it looks like.
```
from utils import decorate
posterior.plot(label='140 heads out of 250')
decorate(xlabel='Proportion of heads (x)',
ylabel='Probability',
title='Posterior distribution of x')
```
Again, the posterior mean is about 0.56, with a 90% credible interval from 0.51 to 0.61.
```
print(posterior.mean(),
posterior.credible_interval(0.9))
```
The prior mean was 0.5, and the posterior mean is 0.56, so it seems like the data is evidence that the coin is biased.
But, it turns out not to be that simple.
## Evidence
In <<_OliversBlood>>, I said that data are considered evidence in favor of a hypothesis, $A$, if the data are more likely under $A$ than under the alternative, $B$; that is if
$$P(D|A) > P(D|B)$$
Furthermore, we can quantify the strength of the evidence by computing the ratio of these likelihoods, which is known as the [Bayes factor](https://en.wikipedia.org/wiki/Bayes_factor) and often denoted $K$:
$$K = \frac{P(D|A)}{P(D|B)}$$
So, for the Euro problem, let's consider two hypotheses, `fair` and `biased`, and compute the likelihood of the data under each hypothesis.
If the coin is fair, the probability of heads is 50%, and we can compute the probability of the data (140 heads out of 250 spins) using the binomial distribution:
```
k = 140
n = 250
like_fair = binom.pmf(k, n, p=0.5)
like_fair
```
That's the probability of the data, given that the coin is fair.
But if the coin is biased, what's the probability of the data? That depends on what "biased" means.
If we know ahead of time that "biased" means the probability of heads is 56%, we can use the binomial distribution again:
```
like_biased = binom.pmf(k, n, p=0.56)
like_biased
```
Now we can compute the likelihood ratio:
```
K = like_biased / like_fair
K
```
The data are about 6 times more likely if the coin is biased, by this definition, than if it is fair.
But we used the data to define the hypothesis, which seems like cheating. To be fair, we should define "biased" before we see the data.
## Uniformly Distributed Bias
Suppose "biased" means that the probability of heads is anything except 50%, and all other values are equally likely.
We can represent that definition by making a uniform distribution and removing 50%.
```
biased_uniform = uniform.copy()
biased_uniform[0.5] = 0
biased_uniform.normalize()
```
To compute the total probability of the data under this hypothesis, we compute the conditional probability of the data for each value of $x$.
```
xs = biased_uniform.qs
likelihood = binom.pmf(k, n, xs)
```
Then multiply by the prior probabilities and add up the products:
```
like_uniform = np.sum(biased_uniform * likelihood)
like_uniform
```
So that's the probability of the data under the "biased uniform" hypothesis.
Now we can compute the likelihood ratio of the data under the `fair` and `biased uniform` hypotheses:
```
K = like_fair / like_uniform
K
```
The data are about two times more likely if the coin is fair than if it is biased, by this definition of "biased".
To get a sense of how strong that evidence is, we can apply Bayes's rule.
For example, if the prior probability is 50% that the coin is biased, the prior odds are 1, so the posterior odds are about 2.1 to 1 and the posterior probability is about 68%.
```
prior_odds = 1
posterior_odds = prior_odds * K
posterior_odds
def prob(o):
return o / (o+1)
posterior_probability = prob(posterior_odds)
posterior_probability
```
Evidence that "moves the needle" from 50% to 68% is not very strong.
Now suppose "biased" doesn't mean every value of $x$ is equally likely. Maybe values near 50% are more likely and values near the extremes are less likely.
We could use a triangle-shaped distribution to represent this alternative definition of "biased":
```
ramp_up = np.arange(50)
ramp_down = np.arange(50, -1, -1)
a = np.append(ramp_up, ramp_down)
triangle = Pmf(a, xs, name='triangle')
triangle.normalize()
```
As we did with the uniform distribution, we can remove 50% as a possible value of $x$ (but it doesn't make much difference if we skip this detail).
```
biased_triangle = triangle.copy()
biased_triangle[0.5] = 0
biased_triangle.normalize()
```
Here's what the triangle prior looks like, compared to the uniform prior.
```
biased_uniform.plot(label='uniform prior')
biased_triangle.plot(label='triangle prior')
decorate(xlabel='Proportion of heads (x)',
ylabel='Probability',
title='Uniform and triangle prior distributions')
```
**Exercise:** Now compute the total probability of the data under this definition of "biased" and compute the Bayes factor, compared with the fair hypothesis.
Is the data evidence that the coin is biased?
```
# Solution goes here
# Solution goes here
# Solution goes here
```
## Bayesian Hypothesis Testing
What we've done so far in this chapter is sometimes called "Bayesian hypothesis testing" in contrast with [statistical hypothesis testing](https://en.wikipedia.org/wiki/Statistical_hypothesis_testing).
In statistical hypothesis testing, we compute a p-value, which is hard to define concisely, and use it to determine whether the results are "statistically significant", which is also hard to define concisely.
The Bayesian alternative is to report the Bayes factor, $K$, which summarizes the strength of the evidence in favor of one hypothesis or the other.
Some people think it is better to report $K$ than a posterior probability because $K$ does not depend on a prior probability.
But as we saw in this example, $K$ often depends on a precise definition of the hypotheses, which can be just as controversial as a prior probability.
In my opinion, Bayesian hypothesis testing is better because it measures the strength of the evidence on a continuum, rather that trying to make a binary determination.
But it doesn't solve what I think is the fundamental problem, which is that hypothesis testing is not asking the question we really care about.
To see why, suppose you test the coin and decide that it is biased after all. What can you do with this answer? In my opinion, not much.
In contrast, there are two questions I think are more useful (and therefore more meaningful):
* Prediction: Based on what we know about the coin, what should we expect to happen in the future?
* Decision-making: Can we use those predictions to make better decisions?
At this point, we've seen a few examples of prediction. For example, in <<_PoissonProcesses>> we used the posterior distribution of goal-scoring rates to predict the outcome of soccer games.
And we've seen one previous example of decision analysis: In <<_DecisionAnalysis>> we used the distribution of prices to choose an optimal bid on *The Price is Right*.
So let's finish this chapter with another example of Bayesian decision analysis, the Bayesian Bandit strategy.
## Bayesian Bandits
If you have ever been to a casino, you have probably seen a slot machine, which is sometimes called a "one-armed bandit" because it has a handle like an arm and the ability to take money like a bandit.
The Bayesian Bandit strategy is named after one-armed bandits because it solves a problem based on a simplified version of a slot machine.
Suppose that each time you play a slot machine, there is a fixed probability that you win. And suppose that different machines give you different probabilities of winning, but you don't know what the probabilities are.
Initially, you have the same prior belief about each of the machines, so you have no reason to prefer one over the others. But if you play each machine a few times, you can use the results to estimate the probabilities. And you can use the estimated probabilities to decide which machine to play next.
At a high level, that's the Bayesian bandit strategy. Now let's see the details.
## Prior Beliefs
If we know nothing about the probability of winning, we can start with a uniform prior.
```
xs = np.linspace(0, 1, 101)
prior = Pmf(1, xs)
prior.normalize()
```
Supposing we are choosing from four slot machines, I'll make four copies of the prior, one for each machine.
```
beliefs = [prior.copy() for i in range(4)]
```
This function displays four distributions in a grid.
```
import matplotlib.pyplot as plt
options = dict(xticklabels='invisible', yticklabels='invisible')
def plot(beliefs, **options):
for i, pmf in enumerate(beliefs):
plt.subplot(2, 2, i+1)
pmf.plot(label='Machine %s' % i)
decorate(yticklabels=[])
if i in [0, 2]:
decorate(ylabel='PDF')
if i in [2, 3]:
decorate(xlabel='Probability of winning')
plt.tight_layout()
```
Here's what the prior distributions look like for the four machines.
```
plot(beliefs)
```
## The Update
Each time we play a machine, we can use the outcome to update our beliefs. The following function does the update.
```
likelihood = {
'W': xs,
'L': 1 - xs
}
def update(pmf, data):
"""Update the probability of winning."""
pmf *= likelihood[data]
pmf.normalize()
```
This function updates the prior distribution in place.
`pmf` is a `Pmf` that represents the prior distribution of `x`, which is the probability of winning.
`data` is a string, either `W` if the outcome is a win or `L` if the outcome is a loss.
The likelihood of the data is either `xs` or `1-xs`, depending on the outcome.
Suppose we choose a machine, play 10 times, and win once. We can compute the posterior distribution of `x`, based on this outcome, like this:
```
np.random.seed(17)
bandit = prior.copy()
for outcome in 'WLLLLLLLLL':
update(bandit, outcome)
```
Here's what the posterior looks like.
```
bandit.plot()
decorate(xlabel='Probability of winning',
ylabel='PDF',
title='Posterior distribution, nine losses, one win')
```
## Multiple Bandits
Now suppose we have four machines with these probabilities:
```
actual_probs = [0.10, 0.20, 0.30, 0.40]
```
Remember that as a player, we don't know these probabilities.
The following function takes the index of a machine, simulates playing the machine once, and returns the outcome, `W` or `L`.
```
from collections import Counter
# count how many times we've played each machine
counter = Counter()
def play(i):
"""Play machine i.
i: index of the machine to play
returns: string 'W' or 'L'
"""
counter[i] += 1
p = actual_probs[i]
if np.random.random() < p:
return 'W'
else:
return 'L'
```
`counter` is a `Counter`, which is a kind of dictionary we'll use to keep track of how many times each machine is played.
Here's a test that plays each machine 10 times.
```
for i in range(4):
for _ in range(10):
outcome = play(i)
update(beliefs[i], outcome)
```
Each time through the inner loop, we play one machine and update our beliefs.
Here's what our posterior beliefs look like.
```
plot(beliefs)
```
Here are the actual probabilities, posterior means, and 90% credible intervals.
```
import pandas as pd
def summarize_beliefs(beliefs):
"""Compute means and credible intervals.
beliefs: sequence of Pmf
returns: DataFrame
"""
columns = ['Actual P(win)',
'Posterior mean',
'Credible interval']
df = pd.DataFrame(columns=columns)
for i, b in enumerate(beliefs):
mean = np.round(b.mean(), 3)
ci = b.credible_interval(0.9)
ci = np.round(ci, 3)
df.loc[i] = actual_probs[i], mean, ci
return df
summarize_beliefs(beliefs)
```
We expect the credible intervals to contain the actual probabilities most of the time.
## Explore and Exploit
Based on these posterior distributions, which machine do you think we should play next? One option would be to choose the machine with the highest posterior mean.
That would not be a bad idea, but it has a drawback: since we have only played each machine a few times, the posterior distributions are wide and overlapping, which means we are not sure which machine is the best; if we focus on one machine too soon, we might choose the wrong machine and play it more than we should.
To avoid that problem, we could go to the other extreme and play all machines equally until we are confident we have identified the best machine, and then play it exclusively.
That's not a bad idea either, but it has a drawback: while we are gathering data, we are not making good use of it; until we're sure which machine is the best, we are playing the others more than we should.
The Bayesian Bandits strategy avoids both drawbacks by gathering and using data at the same time. In other words, it balances exploration and exploitation.
The kernel of the idea is called [Thompson sampling](https://en.wikipedia.org/wiki/Thompson_sampling): when we choose a machine, we choose at random so that the probability of choosing each machine is proportional to the probability that it is the best.
Given the posterior distributions, we can compute the "probability of superiority" for each machine.
Here's one way to do it. We can draw a sample of 1000 values from each posterior distribution, like this:
```
samples = np.array([b.choice(1000)
for b in beliefs])
samples.shape
```
The result has 4 rows and 1000 columns. We can use `argmax` to find the index of the largest value in each column:
```
indices = np.argmax(samples, axis=0)
indices.shape
```
The `Pmf` of these indices is the fraction of times each machine yielded the highest values.
```
pmf = Pmf.from_seq(indices)
pmf
```
These fractions approximate the probability of superiority for each machine. So we could choose the next machine by choosing a value from this `Pmf`.
```
pmf.choice()
```
But that's a lot of work to choose a single value, and it's not really necessary, because there's a shortcut.
If we draw a single random value from each posterior distribution and select the machine that yields the highest value, it turns out that we'll select each machine in proportion to its probability of superiority.
That's what the following function does.
```
def choose(beliefs):
"""Use Thompson sampling to choose a machine.
Draws a single sample from each distribution.
returns: index of the machine that yielded the highest value
"""
ps = [b.choice() for b in beliefs]
return np.argmax(ps)
```
This function chooses one value from the posterior distribution of each machine and then uses `argmax` to find the index of the machine that yielded the highest value.
Here's an example.
```
choose(beliefs)
```
## The Strategy
Putting it all together, the following function chooses a machine, plays once, and updates `beliefs`:
```
def choose_play_update(beliefs):
"""Choose a machine, play it, and update beliefs."""
# choose a machine
machine = choose(beliefs)
# play it
outcome = play(machine)
# update beliefs
update(beliefs[machine], outcome)
```
To test it out, let's start again with a fresh set of beliefs and an empty `Counter`.
```
beliefs = [prior.copy() for i in range(4)]
counter = Counter()
```
If we run the bandit algorithm 100 times, we can see how `beliefs` gets updated:
```
num_plays = 100
for i in range(num_plays):
choose_play_update(beliefs)
plot(beliefs)
```
The following table summarizes the results.
```
summarize_beliefs(beliefs)
```
The credible intervals usually contain the actual probabilities of winning.
The estimates are still rough, especially for the lower-probability machines. But that's a feature, not a bug: the goal is to play the high-probability machines most often. Making the estimates more precise is a means to that end, but not an end itself.
More importantly, let's see how many times each machine got played.
```
def summarize_counter(counter):
"""Report the number of times each machine was played.
counter: Collections.Counter
returns: DataFrame
"""
index = range(4)
columns = ['Actual P(win)', 'Times played']
df = pd.DataFrame(index=index, columns=columns)
for i, count in counter.items():
df.loc[i] = actual_probs[i], count
return df
summarize_counter(counter)
```
If things go according to plan, the machines with higher probabilities should get played more often.
## Summary
In this chapter we finally solved the Euro problem, determining whether the data support the hypothesis that the coin is fair or biased. We found that the answer depends on how we define "biased". And we summarized the results using a Bayes factor, which quantifies the strength of the evidence.
But the answer wasn't satisfying because, in my opinion, the question wasn't interesting. Knowing whether the coin is biased is not useful unless it helps us make better predictions and better decisions.
As an example of a more interesting question, we looked at the "one-armed bandit" problem and a strategy for solving it, the Bayesian bandit algorithm, which tries to balance exploration and exploitation, that is, gathering more information and making the best use of the information we have.
As an exercise, you'll have a chance to explore adaptive strategies for standardized testing.
Bayesian bandits and adaptive testing are examples of [Bayesian decision theory](https://wiki.lesswrong.com/wiki/Bayesian_decision_theory), which is the idea of using a posterior distribution as part of a decision-making process, often by choosing an action that minimizes the costs we expect on average (or maximizes a benefit).
The strategy we used in <<_MaximizingExpectedGain>> to bid on *The Price is Right* is another example.
These strategies demonstrate what I think is the biggest advantage of Bayesian methods over classical statistics. When we represent knowledge in the form of probability distributions, Bayes's theorem tells us how to change our beliefs as we get more data, and Bayesian decision theory tells us how to make that knowledge actionable.
## Exercises
**Exercise:** Standardized tests like the [SAT](https://en.wikipedia.org/wiki/SAT) are often used as part of the admission process at colleges and universities.
The goal of the SAT is to measure the academic preparation of the test-takers; if it is accurate, their scores should reflect their actual ability in the domain of the test.
Until recently, tests like the SAT were taken with paper and pencil, but now students have the option of taking the test online. In the online format, it is possible for the test to be "adaptive", which means that it can [choose each question based on responses to previous questions](https://www.nytimes.com/2018/04/05/education/learning/tests-act-sat.html).
If a student gets the first few questions right, the test can challenge them with harder questions. If they are struggling, it can give them easier questions.
Adaptive testing has the potential to be more "efficient", meaning that with the same number of questions an adaptive test could measure the ability of a tester more precisely.
To see whether this is true, we will develop a model of an adaptive test and quantify the precision of its measurements.
Details of this exercise are in the notebook.
## The Model
The model we'll use is based on [item response theory](https://en.wikipedia.org/wiki/Item_response_theory), which assumes that we can quantify the difficulty of each question and the ability of each test-taker, and that the probability of a correct response is a function of difficulty and ability.
Specifically, a common assumption is that this function is a three-parameter logistic function:
$$\mathrm{p} = c + \frac{1-c}{1 + e^{-a (\theta-b)}}$$
where $\theta$ is the ability of the test-taker and $b$ is the difficulty of the question.
$c$ is the lowest probability of getting a question right, supposing the test-taker with the lowest ability tries to answer the hardest question. On a multiple-choice test with four responses, $c$ might be 0.25, which is the probability of getting the right answer by guessing at random.
$a$ controls the shape of the curve.
The following function computes the probability of a correct answer, given `ability` and `difficulty`:
```
def prob_correct(ability, difficulty):
"""Probability of a correct response."""
a = 100
c = 0.25
x = (ability - difficulty) / a
p = c + (1-c) / (1 + np.exp(-x))
return p
```
I chose `a` to make the range of scores comparable to the SAT, which reports scores from 200 to 800.
Here's what the logistic curve looks like for a question with difficulty 500 and a range of abilities.
```
abilities = np.linspace(100, 900)
diff = 500
ps = prob_correct(abilities, diff)
plt.plot(abilities, ps)
decorate(xlabel='Ability',
ylabel='Probability correct',
title='Probability of correct answer, difficulty=500',
ylim=[0, 1.05])
```
Someone with `ability=900` is nearly certain to get the right answer.
Someone with `ability=100` has about a 25% change of getting the right answer by guessing.
## Simulating the Test
To simulate the test, we'll use the same structure we used for the bandit strategy:
* A function called `play` that simulates a test-taker answering one question.
* A function called `choose` that chooses the next question to pose.
* A function called `update` that uses the outcome (a correct response or not) to update the estimate of the test-taker's ability.
Here's `play`, which takes `ability` and `difficulty` as parameters.
```
def play(ability, difficulty):
"""Simulate a test-taker answering a question."""
p = prob_correct(ability, difficulty)
return np.random.random() < p
```
`play` uses `prob_correct` to compute the probability of a correct answer and `np.random.random` to generate a random value between 0 and 1. The return value is `True` for a correct response and `False` otherwise.
As a test, let's simulate a test-taker with `ability=600` answering a question with `difficulty=500`. The probability of a correct response is about 80%.
```
prob_correct(600, 500)
```
Suppose this person takes a test with 51 questions, all with the same difficulty, `500`.
We expect them to get about 80% of the questions correct.
Here's the result of one simulation.
```
np.random.seed(18)
num_questions = 51
outcomes = [play(600, 500) for _ in range(num_questions)]
np.mean(outcomes)
```
We expect them to get about 80% of the questions right.
Now let's suppose we don't know the test-taker's ability. We can use the data we just generated to estimate it.
And that's what we'll do next.
## The Prior
The SAT is designed so the distribution of scores is roughly normal, with mean 500 and standard deviation 100.
So the lowest score, 200, is three standard deviations below the mean, and the highest score, 800, is three standard deviations above.
We could use that distribution as a prior, but it would tend to cut off the low and high ends of the distribution.
Instead, I'll inflate the standard deviation to 300, to leave open the possibility that `ability` can be less than 200 or more than 800.
Here's a `Pmf` that represents the prior distribution.
```
from scipy.stats import norm
mean = 500
std = 300
qs = np.linspace(0, 1000)
ps = norm(mean, std).pdf(qs)
prior = Pmf(ps, qs)
prior.normalize()
```
And here's what it looks like.
```
prior.plot(label='std=300', color='C5')
decorate(xlabel='Ability',
ylabel='PDF',
title='Prior distribution of ability',
ylim=[0, 0.032])
```
## The Update
The following function takes a prior `Pmf` and the outcome of a single question, and updates the `Pmf` in place.
```
def update_ability(pmf, data):
"""Update the distribution of ability."""
difficulty, outcome = data
abilities = pmf.qs
ps = prob_correct(abilities, difficulty)
if outcome:
pmf *= ps
else:
pmf *= 1 - ps
pmf.normalize()
```
`data` is a tuple that contains the difficulty of a question and the outcome: `True` if the response was correct and `False` otherwise.
As a test, let's do an update based on the outcomes we simulated previously, based on a person with `ability=600` answering 51 questions with `difficulty=500`.
```
actual_600 = prior.copy()
for outcome in outcomes:
data = (500, outcome)
update_ability(actual_600, data)
```
Here's what the posterior distribution looks like.
```
actual_600.plot(color='C4')
decorate(xlabel='Ability',
ylabel='PDF',
title='Posterior distribution of ability')
```
The posterior mean is pretty close to the test-taker's actual ability, which is 600.
```
actual_600.mean()
```
If we run this simulation again, we'll get different results.
## Adaptation
Now let's simulate an adaptive test.
I'll use the following function to choose questions, starting with the simplest strategy: all questions have the same difficulty.
```
def choose(i, belief):
"""Choose the difficulty of the next question."""
return 500
```
As parameters, `choose` takes `i`, which is the index of the question, and `belief`, which is a `Pmf` representing the posterior distribution of `ability`, based on responses to previous questions.
This version of `choose` doesn't use these parameters; they are there so we can test other strategies (see the exercises at the end of the chapter).
The following function simulates a person taking a test, given that we know their actual ability.
```
def simulate_test(actual_ability):
"""Simulate a person taking a test."""
belief = prior.copy()
trace = pd.DataFrame(columns=['difficulty', 'outcome'])
for i in range(num_questions):
difficulty = choose(i, belief)
outcome = play(actual_ability, difficulty)
data = (difficulty, outcome)
update_ability(belief, data)
trace.loc[i] = difficulty, outcome
return belief, trace
```
The return values are a `Pmf` representing the posterior distribution of ability and a `DataFrame` containing the difficulty of the questions and the outcomes.
Here's an example, again for a test-taker with `ability=600`.
```
belief, trace = simulate_test(600)
```
We can use the trace to see how many responses were correct.
```
trace['outcome'].sum()
```
And here's what the posterior looks like.
```
belief.plot(color='C4', label='ability=600')
decorate(xlabel='Ability',
ylabel='PDF',
title='Posterior distribution of ability')
```
Again, the posterior distribution represents a pretty good estimate of the test-taker's actual ability.
## Quantifying Precision
To quantify the precision of the estimates, I'll use the standard deviation of the posterior distribution. The standard deviation measures the spread of the distribution, so higher value indicates more uncertainty about the ability of the test-taker.
In the previous example, the standard deviation of the posterior distribution is about 40.
```
belief.mean(), belief.std()
```
For an exam where all questions have the same difficulty, the precision of the estimate depends strongly on the ability of the test-taker. To show that, I'll loop through a range of abilities and simulate a test using the version of `choice` that always returns `difficulty=500`.
```
actual_abilities = np.linspace(200, 800)
results = pd.DataFrame(columns=['ability', 'posterior_std'])
series = pd.Series(index=actual_abilities, dtype=float, name='std')
for actual_ability in actual_abilities:
belief, trace = simulate_test(actual_ability)
series[actual_ability] = belief.std()
```
The following plot shows the standard deviation of the posterior distribution for one simulation at each level of ability.
The results are noisy, so I also plot a curve fitted to the data by [local regression](https://en.wikipedia.org/wiki/Local_regression).
```
from utils import plot_series_lowess
plot_series_lowess(series, 'C1')
decorate(xlabel='Actual ability',
ylabel='Standard deviation of posterior')
```
The test is most precise for people with ability between `500` and `600`, less precise for people at the high end of the range, and even worse for people at the low end.
When all the questions have difficulty `500`, a person with `ability=800` has a high probability of getting them right. So when they do, we don't learn very much about them.
If the test includes questions with a range of difficulty, it provides more information about people at the high and low ends of the range.
As an exercise at the end of the chapter, you'll have a chance to try out other strategies, including adaptive strategies that choose each question based on previous outcomes.
## Discriminatory Power
In the previous section we used the standard deviation of the posterior distribution to quantify the precision of the estimates. Another way to describe the performance of the test (as opposed to the performance of the test-takers) is to measure "discriminatory power", which is the ability of the test to distinguish correctly between test-takers with different ability.
To measure discriminatory power, I'll simulate a person taking the test 100 times; after each simulation, I'll use the mean of the posterior distribution as their "score".
```
def sample_posterior(actual_ability, iters):
"""Simulate multiple tests and compute posterior means.
actual_ability: number
iters: number of simulated tests
returns: array of scores
"""
scores = []
for i in range(iters):
belief, trace = simulate_test(actual_ability)
score = belief.mean()
scores.append(score)
return np.array(scores)
```
Here are samples of scores for people with several levels of ability.
```
sample_500 = sample_posterior(500, iters=100)
sample_600 = sample_posterior(600, iters=100)
sample_700 = sample_posterior(700, iters=100)
sample_800 = sample_posterior(800, iters=100)
```
Here's what the distributions of scores look like.
```
from empiricaldist import Cdf
cdf_500 = Cdf.from_seq(sample_500)
cdf_600 = Cdf.from_seq(sample_600)
cdf_700 = Cdf.from_seq(sample_700)
cdf_800 = Cdf.from_seq(sample_800)
cdf_500.plot(label='ability=500', color='C1',
linestyle='dashed')
cdf_600.plot(label='ability=600', color='C3')
cdf_700.plot(label='ability=700', color='C2',
linestyle='dashed')
cdf_800.plot(label='ability=800', color='C0')
decorate(xlabel='Test score',
ylabel='CDF',
title='Sampling distribution of test scores')
```
On average, people with higher ability get higher scores, but anyone can have a bad day, or a good day, so there is some overlap between the distributions.
For people with ability between `500` and `600`, where the precision of the test is highest, the discriminatory power of the test is also high.
If people with abilities `500` and `600` take the test, it is almost certain that the person with higher ability will get a higher score.
```
np.mean(sample_600 > sample_500)
```
Between people with abilities `600` and `700`, it is less certain.
```
np.mean(sample_700 > sample_600)
```
And between people with abilities `700` and `800`, it is not certain at all.
```
np.mean(sample_800 > sample_700)
```
But remember that these results are based on a test where all questions are equally difficult.
If you do the exercises at the end of the chapter, you'll see that the performance of the test is better if it includes questions with a range of difficulties, and even better if the test it is adaptive.
Go back and modify `choose`, which is the function that chooses the difficulty of the next question.
1. Write a version of `choose` that returns a range of difficulties by using `i` as an index into a sequence of difficulties.
2. Write a version of `choose` that is adaptive, so it choose the difficulty of the next question based `belief`, which is the posterior distribution of the test-taker's ability, based on the outcome of previous responses.
For both new versions, run the simulations again to quantify the precision of the test and its discriminatory power.
For the first version of `choose`, what is the ideal distribution of difficulties?
For the second version, what is the adaptive strategy that maximizes the precision of the test over the range of abilities?
```
# Solution goes here
# Solution goes here
```
| github_jupyter |
```
#import tensorflow as tf
import pandas as pd
#from pandas import ExcelWriter
#from pandas import ExcelFile
from transformers import BertForSequenceClassification,BertModel, BertConfig
import torch
from transformers import BertTokenizer
import numpy as np
# If there's a GPU available...
if torch.cuda.is_available():
# Tell PyTorch to use the GPU.
device = torch.device("cuda")
print('There are %d GPU(s) available.' % torch.cuda.device_count())
print('We will use the GPU:', torch.cuda.get_device_name(0))
# If not...
else:
print('No GPU available, using the CPU instead.')
device = torch.device("cpu")
tokenizer = BertTokenizer.from_pretrained('bert-base-uncased')
## set the padding
tokenizer.pad_token = '[PAD]'
tokenizer.pad_token_id
## bert encoder -- set output_hidden_states to be TRUE
encoder = BertModel.from_pretrained('bert-base-uncased',output_hidden_states=True)
encoder.config
```
## pading and truncation // mask
```
from keras.preprocessing.sequence import pad_sequences
def preprocess_data(tokenizer, sentences, MAX_LEN = 256):
"""
:params[in]: tokenizer, the configured tokenizer
:params[in]: sentences, list of strings
"""
# 1. Tokenize all of the sentences and map the tokens to thier word IDs.
input_ids = []
# For every sentence...
for sent in sentences:
# `encode` will:
# (1) Tokenize the sentence.
# (2) Prepend the `[CLS]` token to the start.
# (3) Append the `[SEP]` token to the end.
# (4) Map tokens to their IDs.
encoded_sent = tokenizer.encode(
sent, # Sentence to encode.
add_special_tokens = True, # Add '[CLS]' and '[SEP]'
# This function also supports truncation and conversion
# to pytorch tensors, but we need to do padding, so we
# can't use these features :( .
#max_length = 128, # Truncate all sentences.
#return_tensors = 'pt', # Return pytorch tensors.
)
# Add the encoded sentence to the list.
input_ids.append(encoded_sent)
# We'll borrow the `pad_sequences` utility function to do this.
from keras.preprocessing.sequence import pad_sequences
# Set the maximum sequence length.
# maximum training sentence length of 87...
print('\nPadding/truncating all sentences to %d values...' % MAX_LEN)
print('\nPadding token: "{:}", ID: {:}'.format(tokenizer.pad_token, tokenizer.pad_token_id))
# Pad our input tokens with value 0.
# "post" indicates that we want to pad and truncate at the end of the sequence,
# as opposed to the beginning.
input_ids = pad_sequences(input_ids, maxlen=MAX_LEN, dtype="long",
value=0, truncating="post", padding="post")
print('\nDone.')
# Create attention masks
attention_masks = []
# For each sentence...
for sent in input_ids:
# Create the attention mask.
# - If a token ID is 0, then it's padding, set the mask to 0.
# - If a token ID is > 0, then it's a real token, set the mask to 1.
att_mask = [int(token_id > 0) for token_id in sent]
# Store the attention mask for this sentence.
attention_masks.append(att_mask)
return input_ids, attention_masks
### an example
sentences = ['I am working at Duke university', 'Duke is at Durham, North carolina']
input_ids,attention_masks=preprocess_data(tokenizer, sentences, MAX_LEN = 256)
input_ids
train_inputs = torch.LongTensor(input_ids)
train_masks = torch.LongTensor(attention_masks)
out=encoder(train_inputs)
help(encoder)
type(out)
out[0]
out[1]
out[2]
```
| github_jupyter |
# Accelerate finetuning of GPT2 model for Language Modeling task using ONNX Runtime Training
This notebook contains a walkthrough of using ONNX Runtime Training in Azure Machine Learning service to finetune [GPT2](https://cdn.openai.com/better-language-models/language_models_are_unsupervised_multitask_learners.pdf) models. This example uses ONNX Runtime Training to fine-tune the GPT2 PyTorch model maintained at https://github.com/huggingface/transformers.
Specificaly, we showcase finetuning the [pretrained GPT2-medium](https://huggingface.co/transformers/pretrained_models.html), which has 345M parameters using ORT.
Steps:
- Intialize an AzureML workspace
- Register a datastore to use preprocessed data for training
- Create an AzureML experiment
- Provision a compute target
- Create a PyTorch Estimator
- Configure and Run
Prerequisites
If you are using an Azure Machine Learning [Compute Instance](https://docs.microsoft.com/en-us/azure/machine-learning/concept-compute-instance) you are all set. Otherwise, you need to setup your environment by installing AzureML Python SDK to run this notebook. Refer to [How to use Estimator in Azure ML](https://github.com/Azure/MachineLearningNotebooks/blob/master/how-to-use-azureml/training-with-deep-learning/how-to-use-estimator/how-to-use-estimator.ipynb) notebook first if you haven't already to establish your connection to the AzureML Workspace.
Refer to instructions at https://github.com/microsoft/onnxruntime-training-examples/blob/master/huggingface-gpt2/README.md before running the steps below.
### Check SDK installation
```
import os
import requests
import sys
import re
# AzureML libraries
import azureml.core
from azureml.core import Experiment, Workspace, Datastore, Run
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.core.compute_target import ComputeTargetException
from azureml.core.conda_dependencies import CondaDependencies
from azureml.core.container_registry import ContainerRegistry
from azureml.core.runconfig import MpiConfiguration, RunConfiguration, DEFAULT_GPU_IMAGE
from azureml.train.dnn import PyTorch
from azureml.train.estimator import Estimator
from azureml.widgets import RunDetails
from azure.common.client_factory import get_client_from_cli_profile
from azure.mgmt.containerregistry import ContainerRegistryManagementClient
# Check core SDK version number
print("SDK version:", azureml.core.VERSION)
```
### AzureML Workspace setup
```
# Create or retrieve Azure machine learning workspace
# see https://docs.microsoft.com/en-us/python/api/overview/azure/ml/?view=azure-ml-py
ws = Workspace.get(name="myworkspace", subscription_id='<azure-subscription-id>', resource_group='myresourcegroup')
# Print workspace attributes
print('Workspace name: ' + ws.name,
'Workspace region: ' + ws.location,
'Subscription id: ' + ws.subscription_id,
'Resource group: ' + ws.resource_group, sep = '\n')
```
### Register Datastore
Before running the step below, data prepared using the instructions at https://github.com/microsoft/onnxruntime-training-examples/blob/master/huggingface-gpt2/README.md should be transferred to an Azure Blob container referenced in the `Datastore` registration step. Refer to the documentation at https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-access-data for details on using data in Azure ML experiments.
```
# Create a datastore from blob storage containing training data.
# Consult README.md for instructions downloading and uploading training data.
ds = Datastore.register_azure_blob_container(workspace=ws,
datastore_name='<datastore-name>',
account_name='<storage-account-name>',
account_key='<storage-account-key>',
container_name='<storage-container-name>')
# Print datastore attributes
print('Datastore name: ' + ds.name,
'Container name: ' + ds.container_name,
'Datastore type: ' + ds.datastore_type,
'Workspace name: ' + ds.workspace.name, sep = '\n')
```
### Create AzureML Compute Cluster
This recipe is supported on Azure Machine Learning Service using 16 x Standard_NC24rs_v3 or 8 x Standard_ND40rs_v2 VMs. In the next step, you will create an AzureML Compute cluster of Standard_NC40s_v2 GPU VMs with the specified name, if it doesn't already exist in your workspace.
```
# Create GPU cluster
gpu_cluster_name = "ortgptfinetune"
try:
gpu_compute_target = ComputeTarget(workspace=ws, name=gpu_cluster_name)
print('Found existing compute target.')
except ComputeTargetException:
print('Creating a new compute target...')
compute_config = AmlCompute.provisioning_configuration(vm_size='Standard_ND40rs_v2', min_nodes=0, max_nodes=8)
gpu_compute_target = ComputeTarget.create(ws, gpu_cluster_name, compute_config)
gpu_compute_target.wait_for_completion(show_output=True)
# Create experiment for training
experiment_name = 'gpt2_medium-ort-finetuning'
experiment = Experiment(ws, name=experiment_name)
```
### Create Estimator
Notes before running the following step:
* Update the following step to replace two occurences of `<blob-path-to-training-data>` with the actual path in the datastore to the training data.
* If you followed instructions at https://github.com/microsoft/onnxruntime-training-examples/blob/master/huggingface-gpt2/README.md to prepare data, make sure that the data and others files that are not code or config are moved out `workspace` directory. Data files should have been moved to a `Datastore` to use in training.
* Update the occurance of `<tagged-onnxruntime-gpt-container>` with the tag of the built docker image pushed to a container registry. Similarly, update the `<azure-subscription-id>` and `<container-registry-resource-group>` with the contair registry's subscription ID and resource group.
| VM SKU | GPU memory | gpu_count | ORT_batch_size |
| ------------------ |:----------------:|:---------:|:-------:|
| Standard_ND40rs_v2 | 32 GB | 8 | 4 |
| Standard_NC24rs_v3 | 16 GB | 4 | 1 |
```
# this directory should contain run_language_modeling_ort.py, after files copied over based on the instructions at https://github.com/microsoft/onnxruntime-training-examples/blob/master/huggingface-gpt2/README.md
project_folder = '/path/to/onnxruntime-training-examples/huggingface-gpt2/transformers/examples'
container_image = '<tagged-onnxruntime-gpt-container>'
subscription_id = '<azure-subscription-id>'
container_registry_resource_group = '<container-registry-resource-group>'
registry_details = None
acr = re.match('^((\w+).azurecr.io)/(.*)', container_image)
if acr:
# Extract the relevant parts from the container image
# e.g. onnxtraining.azurecr.io/onnxruntime-gpt:latest
registry_address = acr.group(1) # onnxtraining.azurecr.io
registry_name = acr.group(2) # onnxtraining
container_image = acr.group(3) # onnxruntime-gpt:latest
registry_client = get_client_from_cli_profile(ContainerRegistryManagementClient, subscription_id=subscription_id)
registry_credentials = registry_client.registries.list_credentials(container_registry_resource_group, registry_name)
registry_details = ContainerRegistry()
registry_details.address = registry_address
registry_details.username = registry_credentials.username
registry_details.password = registry_credentials.passwords[0].value
# set MPI configuration
# set processes per node to be equal to GPU count on SKU.
# this will change based on NC v/s ND series VMs
mpi = MpiConfiguration()
mpi.process_count_per_node = 8
import uuid
output_id = uuid.uuid1().hex
# Define the script parameters.
# To run training PyTorch instead of ORT, remove the --ort_trainer flag.
# To run evaluation using PyTorch instead of ORT, use the --do_eval_in_torch flag.
script_params = {
'--model_type' : 'gpt2-medium',
'--model_name_or_path' : 'gpt2-medium',
'--tokenizer_name' : 'gpt2-medium',
'--config_name' : 'gpt2-medium',
'--do_eval' : '',
'--do_train': '',
'--train_data_file' : ds.path('benchmarking/WIKI/wikitext-2/wiki.train.tokens').as_mount(),
'--eval_data_file' : ds.path('benchmarking/WIKI/wikitext-2/wiki.valid.tokens').as_mount(),
'--output_dir' : ds.path(f'output/{experiment_name}/{output_id}/').as_mount(),
'--per_gpu_train_batch_size' : '4',
'--per_gpu_eval_batch_size' : '4',
'--gradient_accumulation_steps' : '4',
'--block_size' : '1024',
'--weight_decay' : '0.01',
'--overwrite_output_dir' : '',
'--num_train_epocs' : '5',
'--ort_trainer' : ''
}
# Define training estimator for ORT run
# Consult https://docs.microsoft.com/en-us/azure/machine-learning/how-to-train-ml-models
# Fill in blob path to training data in argument below
# AzureML Estimator that describes how to run the Experiment
estimator_ort = PyTorch(source_directory=project_folder,
# Compute configuration
compute_target = gpu_compute_target,
node_count=4,
distributed_training = mpi,
use_gpu = True,
# supply Docker image
use_docker = True,
custom_docker_image = container_image,
image_registry_details=registry_details,
user_managed = True,
# Training script parameters
script_params = script_params,
entry_script = 'run_language_modeling.py',
)
```
### Run AzureML experiment
```
# Submit ORT run (check logs from Outputs + logs tab of corresponding link)
run = experiment.submit(estimator_ort)
RunDetails(run).show()
print(run.get_portal_url())
```
| github_jupyter |
```
# This cell is for the Google Colaboratory
# https://stackoverflow.com/a/63519730
if 'google.colab' in str(get_ipython()):
# https://colab.research.google.com/notebooks/io.ipynb
import google.colab.drive as gcdrive
# may need to visit a link for the Google Colab authorization code
gcdrive.mount("/content/drive/")
import sys
sys.path.insert(0,"/content/drive/My Drive/Colab Notebooks/nmisp/30_num_int")
# 그래프, 수학 기능 추가
# Add graph and math features
import pylab as py
import numpy as np
```
# 1차 적분<br>First Order Numerical Integration
[](https://www.youtube.com/watch?v=1p0NHR5w0Lc)
다시 면적이 1인 반원을 생각해 보자.<br>Again, let's think about the half circle with area of 1.
$$
\begin{align}
\pi r^2 &= 2 \\
r^2 &= \frac{2}{\pi} \\
r &= \sqrt{\frac{2}{\pi}}
\end{align}
$$
```
r = py.sqrt(2.0 / py.pi)
def half_circle(x):
return py.sqrt(r**2 - x**2)
```
$$
y = \sqrt{r^2 - x^2}
$$
```
import plot_num_int as pi
pi.plot_a_half_circle_of_area(1)
pi.axis_equal_grid_True()
```
이번에는 사다리꼴 규칙을 이용해서 구해 보기로 하자.<br>
This time, let's use the trapezoid rule to find its area.
## 사다리꼴 규칙<br>Trapezoid Rule
다음과 같은 사다리꼴을 생각해 보자.<br>Let's think about a trapezoid as follows.
```
x_array = (0, 1)
y_array = (1, 2)
py.fill_between(x_array, y_array)
py.axis('equal')
py.axis('off')
py.text(-0.25, 0.5, '$y_i$')
py.text(1.15, 1, '$y_{i+1}$')
py.text(0.5, -0.3, '$\Delta x$');
```
사다리꼴의 면적은 다음과 같다.<br>
Area of a trapezoid is as follows.
$$
a_i=\frac{1}{2} \left( y_i + y_{i+1} \right) \Delta x
$$
## 1차 적분<br>First order numerical integration
마찬가지로 일정 간격으로 $x$ 좌표를 나누어 보자.<br>
Same as before, let's divide $x$ coordinates in a constant interval.
```
n = 10
pi.plot_half_circle_with_stems(n, 1)
# 사다리꼴의 좌표를 나눔 Find coordinates for the trapezoids
x_array_bar = py.linspace(-r, r, n+1)
y_array_bar = half_circle(x_array_bar)
# 각 사다리꼴의 폭 Width of each trapezoid
delta_x = x_array_bar[1] - x_array_bar[0]
# 일련의 사다리꼴을 그림 Plot a series of the trapezoids
xp, yp = x_array_bar[0], y_array_bar[0]
for x, y in zip(x_array_bar[1:], y_array_bar[1:]):
py.fill_between((xp, x), (yp, y), alpha=0.5, color=py.random((1, 3)))
xp, yp = x, y
py.axis('equal')
py.grid(True)
```
사다리꼴의 면적을 하나씩 구해서 더해보자.<br>Let's accumulate the area of trapezoids.
$$
Area = \sum_{k=0}^{n-1} F_k
$$
$$
F_k = \frac{\Delta x}{2}\left[f(x_k)+f(x_{k+1})\right]
$$
$$
Area = \sum_{k=0}^{n-1} \frac{1}{2}\left[f(x_k)+f(x_{k+1})\right] \Delta x
$$
```
def get_delta_x(xi, xe, n):
return (xe - xi) / n
def num_int_1(f, xi, xe, n, b_verbose=False):
x_array = py.linspace(xi, xe, n+1)
delta_x = x_array[1] - x_array[0]
integration_result = 0.0
x_k = x_array[0]
y_k = f(x_k)
for k, x_k_plus_1 in enumerate(x_array[1:]):
y_k_plus_1 = f(x_k_plus_1)
F_k = 0.5 * (y_k + y_k_plus_1) * (x_k_plus_1 - x_k)
if b_verbose: print('k = %2d, F_k = %g' % (k, F_k))
integration_result += F_k
x_k, y_k = x_k_plus_1, y_k_plus_1
return integration_result
n = 10
result = num_int_1(half_circle, -r, r, n, b_verbose=True)
print('result =', result)
```
예상한 값 1에 더 비슷한 값을 얻기 위해 더 잘게 나누어 보자<br>
To obtain the result closer to the expected value of 1, let's divide with a narrower interval.
```
n = 100
result = num_int_1(half_circle, -r, r, n)
print('result =', result)
%timeit -n 100 result = num_int_1(half_circle, -r, r, n)
```
### $cos \theta$의 반 주기<br>Half period of $cos \theta$
```
n = 10
result_cos = num_int_1(py.cos, 0, py.pi, n, b_verbose=True)
print('result =', result_cos)
n = 100
result_cos = num_int_1(py.cos, 0, py.pi, n)
print('result =', result_cos)
```
### 1/4 원<br>A quarter circle
```
n = 10
result_quarter = num_int_1(half_circle, -r, 0, n, b_verbose=True)
print('result =', result_quarter)
n = 100
result_quarter = num_int_1(half_circle, -r, 0, n)
print('result =', result_quarter)
```
## 연습 문제<br>Exercises
도전 과제 1 : 다른 조건이 같을 때 0차 적분과 사다리꼴 적분의 오차를 비교해 보시오. 필요하면 해당 파이썬 함수를 복사하시오.<br>Try this 1 : Compare the errors of the zeroth and first order integrations of the half circle example above using the same conditions. Duplicate the python function if necessary.
도전 과제 2 : 길이 $L=3[m]$ 인 외팔보가 분포 하중 $\omega=50sin\left(\frac{1}{2L}\pi x\right)[N/m]$을 받고 있을 때 전단력과 굽힘모멘트 선도를 구하시오.<br>
Try this 2 : Plot diagrams of shear force and bending moment of a cantilever with length $L=3m$ under distributed load $\omega=50sin\left(\frac{1}{2L}\pi x\right)[N/m]$. <br>
(ref : C 4.4, Pytel, Kiusalaas & Sharma, Mechanics of Materials, 2nd Ed, SI, Cengage Learning, 2011.)
## 함수형 프로그래밍<br>Functional programming
간격이 일정하다면 면적의 근사값을 다음과 같이 바꾸어 쓸 수 있다.<br>
If the interval $\Delta x$ is constant, we may rewrite the approximation of the area as follows.
$$
\begin{align}
Area &= \sum_{k=0}^{n-1} \frac{1}{2}\left[f(x_k)+f(x_{k+1})\right] \Delta x \\
&= \Delta x \sum_{k=0}^{n-1} \frac{1}{2}\left[f(x_k)+f(x_{k+1})\right]
\end{align}
$$
$$
\begin{align}
\sum_{k=0}^{n-1} \frac{1}{2}\left[f(x_k)+f(x_{k+1})\right] &= \frac{1}{2}\left[f(x_0)+f(x_1)\right] \\
&+ \frac{1}{2}\left[f(x_1)+f(x_2)\right] \\
&+ \frac{1}{2}\left[f(x_2)+f(x_3)\right] \\
& \ldots \\
&+ \frac{1}{2}\left[f(x_{n-2})+f(x_{n-1})\right] \\
&+ \frac{1}{2}\left[f(x_{n-1})+f(x_{n})\right] \\
&= \frac{1}{2}f(x_0) + \sum_{k=1}^{n-1} f(x_k) + \frac{1}{2}f(x_{n}) \\
&= \frac{1}{2}\left[f(x_0) + f(x_{n})\right] + \sum_{k=1}^{n-1} f(x_k)
\end{align}
$$
$$
\begin{align}
Area &= \Delta x \sum_{k=0}^{n-1} \frac{1}{2}\left[f(x_k)+f(x_{k+1})\right] \\
&= \Delta x \left[\frac{1}{2}\left[f(x_0) + f(x_{n})\right] + \sum_{k=1}^{n-1} f(x_k)\right]
\end{align}
$$
할당문 없이 `sum()` 과 `map()` 함수로 구현해 보자.<br>
Instead of assignments, let's implement using `sum()` and `map()` functions.
```
def num_int_1_functional(f, xi, xe, n):
# get_delta_x() 함수 호출 횟수를 줄이기 위해 함수 안의 함수를 사용
# To reduce the number of calling function get_delta_x(), define inner functions
def with_delta_x(f, xi, n, delta_x=get_delta_x(xi, xe, n)):
return delta_x * (
0.5 * (f(xi) + f(xe))
+ sum(
map(
f,
py.arange(xi + delta_x, xe - delta_x*0.1, delta_x),
)
)
)
return with_delta_x(f, xi, n)
n = 100
result_func = num_int_1_functional(half_circle, -r, r, n)
print('result_func =', result_func)
assert 1e-7 > abs(result - result_func), f"result = {result}, result_func = {result_func}"
%timeit -n 100 result_func = num_int_1_functional(half_circle, -r, r, n)
```
## NumPy 벡터화<br>Vectorization of NumPy
```
import pylab as py
def num_int_1_vector_with_delta_x(f, xi, xe, n, delta_x):
return delta_x * (
f(py.arange(xi+delta_x, xe-delta_x*0.5, get_delta_x(xi, xe, n))).sum()
+ 0.5 * f(py.array((xi, xe))).sum()
)
def num_int_1_vector(f, xi, xe, n):
return num_int_1_vector_with_delta_x(f, xi, xe, n, get_delta_x(xi, xe, n))
n = 100
result_vect = num_int_1_vector(half_circle, -r, r, n)
print('result_vect =', result_vect)
assert 1e-7 > abs(result - result_vect), f"result = {result}, result_vect = {result_vect}"
%timeit -n 100 result_func = num_int_1_vector(half_circle, -r, r, n)
```
## 시험<br>Test
아래는 함수가 맞게 작동하는지 확인함<br>
Following cells verify whether the functions work correctly.
```
import pylab as py
r = py.sqrt(1.0 / py.pi)
n = 10
delta_x = r/n
def half_circle(x):
return py.sqrt(r**2 - x ** 2)
assert 0.25 > num_int_1(half_circle, -r, 0, n)
assert 0.25 > num_int_1(half_circle, 0, r, n)
assert 0.25 > num_int_1_functional(half_circle, -r, 0, n)
assert 0.25 > num_int_1_functional(half_circle, 0, r, n)
assert 0.25 > num_int_1_vector(half_circle, -r, 0, n)
assert 0.25 > num_int_1_vector(half_circle, 0, r, n)
assert 0.1 > (abs(num_int_1(half_circle, -r, 0, n) - 0.25) * 4)
assert 0.1 > (abs(num_int_1(half_circle, 0, r, n) - 0.25) * 4)
assert 0.1 > (abs(num_int_1_functional(half_circle, -r, 0, n) - 0.25) * 4)
assert 0.1 > (abs(num_int_1_functional(half_circle, 0, r, n) - 0.25) * 4)
assert 0.1 > (abs(num_int_1_vector(half_circle, -r, 0, n) - 0.25) * 4)
assert 0.1 > (abs(num_int_1_vector(half_circle, 0, r, n) - 0.25) * 4)
```
## Final Bell<br>마지막 종
```
# stackoverfow.com/a/24634221
import os
os.system("printf '\a'");
```
| github_jupyter |
```
from __future__ import division
from __future__ import print_function
import time
import os
# Train on CPU (hide GPU) due to memory constraints
os.environ['CUDA_VISIBLE_DEVICES'] = ""
import tensorflow as tf
import numpy as np
import scipy.sparse as sp
from sklearn.metrics import roc_auc_score
from sklearn.metrics import average_precision_score
from gae.optimizer import OptimizerAE, OptimizerVAE
from gae.input_data import load_data
from gae.model import GCNModelAE, GCNModelVAE
from gae.preprocessing import preprocess_graph, construct_feed_dict, sparse_to_tuple, mask_test_edges
# Settings
flags = tf.app.flags
FLAGS = flags.FLAGS
flags.DEFINE_float('learning_rate', 0.01, 'Initial learning rate.')
flags.DEFINE_integer('epochs', 200, 'Number of epochs to train.')
flags.DEFINE_integer('hidden1', 32, 'Number of units in hidden layer 1.')
flags.DEFINE_integer('hidden2', 16, 'Number of units in hidden layer 2.')
flags.DEFINE_float('weight_decay', 0., 'Weight for L2 loss on embedding matrix.')
flags.DEFINE_float('dropout', 0., 'Dropout rate (1 - keep probability).')
flags.DEFINE_string('model', 'gcn_ae', 'Model string.')
flags.DEFINE_string('dataset', 'cora', 'Dataset string.')
flags.DEFINE_integer('features', 1, 'Whether to use features (1) or not (0).')
model_str = FLAGS.model
dataset_str = FLAGS.dataset
# Load data
adj, features = load_data(dataset_str)
# Store original adjacency matrix (without diagonal entries) for later
adj_orig = adj
adj_orig = adj_orig - sp.dia_matrix((adj_orig.diagonal()[np.newaxis, :], [0]), shape=adj_orig.shape)
adj_orig.eliminate_zeros()
adj_train, train_edges, val_edges, val_edges_false, test_edges, test_edges_false = mask_test_edges(adj)
adj = adj_train
if FLAGS.features == 0:
features = sp.identity(features.shape[0]) # featureless
# Some preprocessing
adj_norm = preprocess_graph(adj)
# Define placeholders
placeholders = {
'features': tf.sparse_placeholder(tf.float32),
'adj': tf.sparse_placeholder(tf.float32),
'adj_orig': tf.sparse_placeholder(tf.float32),
'dropout': tf.placeholder_with_default(0., shape=())
}
# Graph attributes
num_nodes = adj.shape[0]
features = sparse_to_tuple(features.tocoo())
num_features = features[2][1]
features_nonzero = features[1].shape[0]
# Create model
model = None
if model_str == 'gcn_ae': # Graph Auto-Encoder
model = GCNModelAE(placeholders, num_features, features_nonzero)
elif model_str == 'gcn_vae': # Graph Variational Auto-Encoder
model = GCNModelVAE(placeholders, num_features, num_nodes, features_nonzero)
# How much to weigh positive examples (true edges) in cost print_function
# Want to weigh less-frequent classes higher, so as to prevent model output bias
# pos_weight = (num. negative samples / (num. positive samples)
pos_weight = float(adj.shape[0] * adj.shape[0] - adj.sum()) / adj.sum()
# normalize (scale) average weighted cost
norm = adj.shape[0] * adj.shape[0] / float((adj.shape[0] * adj.shape[0] - adj.sum()) * 2)
# Optimizer
with tf.name_scope('optimizer'):
# Graph Auto-Encoder: use weighted_binary_crossentropy
if model_str == 'gcn_ae':
opt = OptimizerAE(preds=model.reconstructions,
labels=tf.reshape(tf.sparse_tensor_to_dense(placeholders['adj_orig'],
validate_indices=False), [-1]),
pos_weight=pos_weight,
norm=norm)
# Graph Variational Auto-Encoder: use weighted_binary_crossentropy + KL Divergence
elif model_str == 'gcn_vae':
opt = OptimizerVAE(preds=model.reconstructions,
labels=tf.reshape(tf.sparse_tensor_to_dense(placeholders['adj_orig'],
validate_indices=False), [-1]),
model=model, num_nodes=num_nodes,
pos_weight=pos_weight,
norm=norm)
# Initialize session
sess = tf.Session()
sess.run(tf.global_variables_initializer())
cost_val = []
acc_val = []
# Calculate ROC AUC
def get_roc_score(edges_pos, edges_neg, emb=None):
if emb is None:
feed_dict.update({placeholders['dropout']: 0})
emb = sess.run(model.z_mean, feed_dict=feed_dict)
def sigmoid(x):
return 1 / (1 + np.exp(-x))
# Predict on test set of edges
adj_rec = np.dot(emb, emb.T)
preds = []
pos = []
for e in edges_pos:
preds.append(sigmoid(adj_rec[e[0], e[1]]))
pos.append(adj_orig[e[0], e[1]])
preds_neg = []
neg = []
for e in edges_neg:
preds_neg.append(sigmoid(adj_rec[e[0], e[1]]))
neg.append(adj_orig[e[0], e[1]])
preds_all = np.hstack([preds, preds_neg])
labels_all = np.hstack([np.ones(len(preds)), np.zeros(len(preds))])
roc_score = roc_auc_score(labels_all, preds_all)
ap_score = average_precision_score(labels_all, preds_all)
return roc_score, ap_score
cost_val = []
acc_val = []
val_roc_score = []
# Add in diagonals
adj_label = adj_train + sp.eye(adj_train.shape[0])
adj_label = sparse_to_tuple(adj_label)
# Train model
for epoch in range(FLAGS.epochs):
t = time.time()
# Construct feed dictionary
feed_dict = construct_feed_dict(adj_norm, adj_label, features, placeholders)
feed_dict.update({placeholders['dropout']: FLAGS.dropout})
# Run single weight update
outs = sess.run([opt.opt_op, opt.cost, opt.accuracy], feed_dict=feed_dict)
# Compute average loss
avg_cost = outs[1]
avg_accuracy = outs[2]
# Evaluate predictions
roc_curr, ap_curr = get_roc_score(val_edges, val_edges_false)
val_roc_score.append(roc_curr)
# Print results for this epoch
print("Epoch:", '%04d' % (epoch + 1), "train_loss=", "{:.5f}".format(avg_cost),
"train_acc=", "{:.5f}".format(avg_accuracy), "val_roc=", "{:.5f}".format(val_roc_score[-1]),
"val_ap=", "{:.5f}".format(ap_curr),
"time=", "{:.5f}".format(time.time() - t))
print("Optimization Finished!")
# Print final results
roc_score, ap_score = get_roc_score(test_edges, test_edges_false)
print('Test ROC score: ' + str(roc_score))
print('Test AP score: ' + str(ap_score))
```
| github_jupyter |
# <span style="color:Maroon">Trade Strategy
__Summary:__ <span style="color:Blue">In this code we shall test the results of given model
```
# Import required libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import os
np.random.seed(0)
import warnings
warnings.filterwarnings('ignore')
# User defined names
index = "SSE Composite"
filename_whole = "whole_dataset"+index+"_gbm_model.csv"
filename_trending = "Trending_dataset"+index+"_gbm_model.csv"
filename_meanreverting = "MeanReverting_dataset"+index+"_gbm_model.csv"
date_col = "Date"
Rf = 0.01 #Risk free rate of return
# Get current working directory
mycwd = os.getcwd()
print(mycwd)
# Change to data directory
os.chdir("..")
os.chdir(str(os.getcwd()) + "\\Data")
# Read the datasets
df_whole = pd.read_csv(filename_whole, index_col=date_col)
df_trending = pd.read_csv(filename_trending, index_col=date_col)
df_meanreverting = pd.read_csv(filename_meanreverting, index_col=date_col)
# Convert index to datetime
df_whole.index = pd.to_datetime(df_whole.index)
df_trending.index = pd.to_datetime(df_trending.index)
df_meanreverting.index = pd.to_datetime(df_meanreverting.index)
# Head for whole dataset
df_whole.head()
df_whole.shape
# Head for Trending dataset
df_trending.head()
df_trending.shape
# Head for Mean Reverting dataset
df_meanreverting.head()
df_meanreverting.shape
# Merge results from both models to one
df_model = df_trending.append(df_meanreverting)
df_model.sort_index(inplace=True)
df_model.head()
df_model.shape
```
## <span style="color:Maroon">Functions
```
def initialize(df):
days, Action1, Action2, current_status, Money, Shares = ([] for i in range(6))
Open_price = list(df['Open'])
Close_price = list(df['Adj Close'])
Predicted = list(df['Predicted'])
Action1.append(Predicted[0])
Action2.append(0)
current_status.append(Predicted[0])
if(Predicted[0] != 0):
days.append(1)
if(Predicted[0] == 1):
Money.append(0)
else:
Money.append(200)
Shares.append(Predicted[0] * (100/Open_price[0]))
else:
days.append(0)
Money.append(100)
Shares.append(0)
return days, Action1, Action2, current_status, Predicted, Money, Shares, Open_price, Close_price
def Action_SA_SA(days, Action1, Action2, current_status, i):
if(current_status[i-1] != 0):
days.append(1)
else:
days.append(0)
current_status.append(current_status[i-1])
Action1.append(0)
Action2.append(0)
return days, Action1, Action2, current_status
def Action_ZE_NZE(days, Action1, Action2, current_status, i):
if(days[i-1] < 5):
days.append(days[i-1] + 1)
Action1.append(0)
Action2.append(0)
current_status.append(current_status[i-1])
else:
days.append(0)
Action1.append(current_status[i-1] * (-1))
Action2.append(0)
current_status.append(0)
return days, Action1, Action2, current_status
def Action_NZE_ZE(days, Action1, Action2, current_status, Predicted, i):
current_status.append(Predicted[i])
Action1.append(Predicted[i])
Action2.append(0)
days.append(days[i-1] + 1)
return days, Action1, Action2, current_status
def Action_NZE_NZE(days, Action1, Action2, current_status, Predicted, i):
current_status.append(Predicted[i])
Action1.append(Predicted[i])
Action2.append(Predicted[i])
days.append(1)
return days, Action1, Action2, current_status
def get_df(df, Action1, Action2, days, current_status, Money, Shares):
df['Action1'] = Action1
df['Action2'] = Action2
df['days'] = days
df['current_status'] = current_status
df['Money'] = Money
df['Shares'] = Shares
return df
def Get_TradeSignal(Predicted, days, Action1, Action2, current_status):
# Loop over 1 to N
for i in range(1, len(Predicted)):
# When model predicts no action..
if(Predicted[i] == 0):
if(current_status[i-1] != 0):
days, Action1, Action2, current_status = Action_ZE_NZE(days, Action1, Action2, current_status, i)
else:
days, Action1, Action2, current_status = Action_SA_SA(days, Action1, Action2, current_status, i)
# When Model predicts sell
elif(Predicted[i] == -1):
if(current_status[i-1] == -1):
days, Action1, Action2, current_status = Action_SA_SA(days, Action1, Action2, current_status, i)
elif(current_status[i-1] == 0):
days, Action1, Action2, current_status = Action_NZE_ZE(days, Action1, Action2, current_status, Predicted,
i)
else:
days, Action1, Action2, current_status = Action_NZE_NZE(days, Action1, Action2, current_status, Predicted,
i)
# When model predicts Buy
elif(Predicted[i] == 1):
if(current_status[i-1] == 1):
days, Action1, Action2, current_status = Action_SA_SA(days, Action1, Action2, current_status, i)
elif(current_status[i-1] == 0):
days, Action1, Action2, current_status = Action_NZE_ZE(days, Action1, Action2, current_status, Predicted,
i)
else:
days, Action1, Action2, current_status = Action_NZE_NZE(days, Action1, Action2, current_status, Predicted,
i)
return days, Action1, Action2, current_status
def Get_FinancialSignal(Open_price, Action1, Action2, Money, Shares, Close_price):
for i in range(1, len(Open_price)):
if(Action1[i] == 0):
Money.append(Money[i-1])
Shares.append(Shares[i-1])
else:
if(Action2[i] == 0):
# Enter new position
if(Shares[i-1] == 0):
Shares.append(Action1[i] * (Money[i-1]/Open_price[i]))
Money.append(Money[i-1] - Action1[i] * Money[i-1])
# Exit the current position
else:
Shares.append(0)
Money.append(Money[i-1] - Action1[i] * np.abs(Shares[i-1]) * Open_price[i])
else:
Money.append(Money[i-1] -1 *Action1[i] *np.abs(Shares[i-1]) * Open_price[i])
Shares.append(Action2[i] * (Money[i]/Open_price[i]))
Money[i] = Money[i] - 1 * Action2[i] * np.abs(Shares[i]) * Open_price[i]
return Money, Shares
def Get_TradeData(df):
# Initialize the variables
days,Action1,Action2,current_status,Predicted,Money,Shares,Open_price,Close_price = initialize(df)
# Get Buy/Sell trade signal
days, Action1, Action2, current_status = Get_TradeSignal(Predicted, days, Action1, Action2, current_status)
Money, Shares = Get_FinancialSignal(Open_price, Action1, Action2, Money, Shares, Close_price)
df = get_df(df, Action1, Action2, days, current_status, Money, Shares)
df['CurrentVal'] = df['Money'] + df['current_status'] * np.abs(df['Shares']) * df['Adj Close']
return df
def Print_Fromated_PL(active_days, number_of_trades, drawdown, annual_returns, std_dev, sharpe_ratio, year):
"""
Prints the metrics
"""
print("++++++++++++++++++++++++++++++++++++++++++++++++++++")
print(" Year: {0}".format(year))
print(" Number of Trades Executed: {0}".format(number_of_trades))
print("Number of days with Active Position: {}".format(active_days))
print(" Annual Return: {:.6f} %".format(annual_returns*100))
print(" Sharpe Ratio: {:.2f}".format(sharpe_ratio))
print(" Maximum Drawdown (Daily basis): {:.2f} %".format(drawdown*100))
print("----------------------------------------------------")
return
def Get_results_PL_metrics(df, Rf, year):
df['tmp'] = np.where(df['current_status'] == 0, 0, 1)
active_days = df['tmp'].sum()
number_of_trades = np.abs(df['Action1']).sum()+np.abs(df['Action2']).sum()
df['tmp_max'] = df['CurrentVal'].rolling(window=20).max()
df['tmp_min'] = df['CurrentVal'].rolling(window=20).min()
df['tmp'] = np.where(df['tmp_max'] > 0, (df['tmp_max'] - df['tmp_min'])/df['tmp_max'], 0)
drawdown = df['tmp'].max()
annual_returns = (df['CurrentVal'].iloc[-1]/100 - 1)
std_dev = df['CurrentVal'].pct_change(1).std()
sharpe_ratio = (annual_returns - Rf)/std_dev
Print_Fromated_PL(active_days, number_of_trades, drawdown, annual_returns, std_dev, sharpe_ratio, year)
return
```
```
# Change to Images directory
os.chdir("..")
os.chdir(str(os.getcwd()) + "\\Images")
```
## <span style="color:Maroon">Whole Dataset
```
df_whole_train = df_whole[df_whole["Sample"] == "Train"]
df_whole_test = df_whole[df_whole["Sample"] == "Test"]
df_whole_test_2019 = df_whole_test[df_whole_test.index.year == 2019]
df_whole_test_2020 = df_whole_test[df_whole_test.index.year == 2020]
output_train_whole = Get_TradeData(df_whole_train)
output_test_whole = Get_TradeData(df_whole_test)
output_test_whole_2019 = Get_TradeData(df_whole_test_2019)
output_test_whole_2020 = Get_TradeData(df_whole_test_2020)
output_train_whole["BuyandHold"] = (100 * output_train_whole["Adj Close"])/(output_train_whole.iloc[0]["Adj Close"])
output_test_whole["BuyandHold"] = (100*output_test_whole["Adj Close"])/(output_test_whole.iloc[0]["Adj Close"])
output_test_whole_2019["BuyandHold"] = (100 * output_test_whole_2019["Adj Close"])/(output_test_whole_2019.iloc[0]
["Adj Close"])
output_test_whole_2020["BuyandHold"] = (100 * output_test_whole_2020["Adj Close"])/(output_test_whole_2020.iloc[0]
["Adj Close"])
Get_results_PL_metrics(output_test_whole_2019, Rf, 2019)
Get_results_PL_metrics(output_test_whole_2020, Rf, 2020)
# Scatter plot to save fig
plt.figure(figsize=(10,5))
plt.plot(output_train_whole["CurrentVal"], 'b-', label="Value (Model)")
plt.plot(output_train_whole["BuyandHold"], 'r--', alpha=0.5, label="Buy and Hold")
plt.xlabel("Date", fontsize=12)
plt.ylabel("Value", fontsize=12)
plt.legend()
plt.title("Train Sample "+ str(index) + " GBM Whole Dataset", fontsize=16)
plt.savefig("Train Sample Whole Dataset GBM Model" + str(index) +'.png')
plt.show()
plt.close()
# Scatter plot to save fig
plt.figure(figsize=(10,5))
plt.plot(output_test_whole["CurrentVal"], 'b-', label="Value (Model)")
plt.plot(output_test_whole["BuyandHold"], 'r--', alpha=0.5, label="Buy and Hold")
plt.xlabel("Date", fontsize=12)
plt.ylabel("Value", fontsize=12)
plt.legend()
plt.title("Test Sample "+ str(index) + " GBM Whole Dataset", fontsize=16)
plt.savefig("Test Sample Whole Dataset GBM Model" + str(index) +'.png')
plt.show()
plt.close()
```
__Comments:__ <span style="color:Blue"> Based on the performance of model on Train Sample, the model has definitely learnt the patter, instead of over-fitting. But the performance of model in Test Sample is very poor
## <span style="color:Maroon">Segment Model
```
df_model_train = df_model[df_model["Sample"] == "Train"]
df_model_test = df_model[df_model["Sample"] == "Test"]
df_model_test_2019 = df_model_test[df_model_test.index.year == 2019]
df_model_test_2020 = df_model_test[df_model_test.index.year == 2020]
output_train_model = Get_TradeData(df_model_train)
output_test_model = Get_TradeData(df_model_test)
output_test_model_2019 = Get_TradeData(df_model_test_2019)
output_test_model_2020 = Get_TradeData(df_model_test_2020)
output_train_model["BuyandHold"] = (100 * output_train_model["Adj Close"])/(output_train_model.iloc[0]["Adj Close"])
output_test_model["BuyandHold"] = (100 * output_test_model["Adj Close"])/(output_test_model.iloc[0]["Adj Close"])
output_test_model_2019["BuyandHold"] = (100 * output_test_model_2019["Adj Close"])/(output_test_model_2019.iloc[0]
["Adj Close"])
output_test_model_2020["BuyandHold"] = (100 * output_test_model_2020["Adj Close"])/(output_test_model_2020.iloc[0]
["Adj Close"])
Get_results_PL_metrics(output_test_model_2019, Rf, 2019)
Get_results_PL_metrics(output_test_model_2020, Rf, 2020)
# Scatter plot to save fig
plt.figure(figsize=(10,5))
plt.plot(output_train_model["CurrentVal"], 'b-', label="Value (Model)")
plt.plot(output_train_model["BuyandHold"], 'r--', alpha=0.5, label="Buy and Hold")
plt.xlabel("Date", fontsize=12)
plt.ylabel("Value", fontsize=12)
plt.legend()
plt.title("Train Sample Hurst Segment GBM Models "+ str(index), fontsize=16)
plt.savefig("Train Sample Hurst Segment GBM Models" + str(index) +'.png')
plt.show()
plt.close()
# Scatter plot to save fig
plt.figure(figsize=(10,5))
plt.plot(output_test_model["CurrentVal"], 'b-', label="Value (Model)")
plt.plot(output_test_model["BuyandHold"], 'r--', alpha=0.5, label="Buy and Hold")
plt.xlabel("Date", fontsize=12)
plt.ylabel("Value", fontsize=12)
plt.legend()
plt.title("Test Sample Hurst Segment GBM Models" + str(index), fontsize=16)
plt.savefig("Test Sample Hurst Segment GBM Models" + str(index) +'.png')
plt.show()
plt.close()
```
__Comments:__ <span style="color:Blue"> Based on the performance of model on Train Sample, the model has definitely learnt the patter, instead of over-fitting. The model does perform well in Test sample (Not compared to Buy and Hold strategy) compared to single model. Hurst Exponent based segmentation has definately added value to the model
| github_jupyter |
# Composition Example
In this example we're going to compose and link multiple scatter plots.
```
import numpy as np
import pandas as pd
import jscatter
```
First we generate some dummy grid data such that each subsequent dataset has have the points.
```
X1, Y1 = np.mgrid[0:8:1, 0:8:1]
X2, Y2 = np.mgrid[0:4:1, 0:4:1]
X3, Y3 = np.mgrid[0:2:1, 0:2:1]
X = [X1, X2, X3]
Y = [Y1, Y2, Y3]
sc1 = jscatter.Scatter(X1.flatten(), Y1.flatten(), color='#dca237', size=10)
sc2 = jscatter.Scatter(X2.flatten(), Y2.flatten(), color='#6fb2e4', size=10)
sc3 = jscatter.Scatter(X3.flatten(), Y3.flatten(), color='#c17da5', size=10)
```
To define correspondes upon hovering and selecting, jupyter-scatter's `compose` function allows to specify select and hover mapping functions. The job of a matter is to map an incoming hover or select event to map to the local points that need to be selected.
```
# In this demo, create_mapper() is a factory function that creates a mapping function for the `i`th scatter plot.
def create_mapper(i):
from functools import reduce
from math import floor
from operator import concat
# This is that target or output number of rows
target_rows = X[i].shape[1]
# The actually mapping function receives two arguments when a hover or select event comes in:
# 1. `j` the index of the scatter plot that triggered the event
# 2. `v` the value of the hover or selection event
def mapper(j, v):
if i == j:
return v
exp = 2 ** abs(i - j)
is_less = i > j
trans_less = lambda idx: floor(idx * 0.5 ** exp)
trans_more = lambda idx: list(range(idx * int(2 ** exp), (idx + 1) * int(2 ** exp)))
is_list = isinstance(v, list)
if is_list:
if is_less:
return list(dict.fromkeys(map(trans_less, v)))
else:
print()
return reduce(concat, map(trans_more, v))
if is_less:
return trans_less(v)
else:
return list(trans_more(v))
return mapper
jscatter.compose(
[sc1, sc2, sc3],
select_mappers=[create_mapper(i) for i in range(3)],
hover_mappers=[create_mapper(i) for i in range(3)]
)
```
### Grid Compositions
You can also compose multiple scatter plots vertically or in a 2D grid through a nested list of scatter plots.
```
sc4 = jscatter.Scatter(
np.random.rand(100), np.random.rand(100), color='#469b76', size=10
)
jscatter.compose([[sc1, sc2], [sc3, sc4]], row_height=200)
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.