text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
# Inspector Introduction
**In this notebook we introduce the Inspector component of HERE Data SDK for Python and demonstrate advanced styling with its ipyleaflet-based backend.**
**Features:**
* Show simple usage patterns related to the inspector.
* Show more advanced customization use cases.
* Show ipyleaflet-specific styling options
**Dependencies:**
* Sample GeoJSON data, obtained from the freely available content on [GitHub](https://github.com/johan/world.geo.json) and available in the sample_datasets folder provided with this notebook
* Catalog: [HERE GeoJSON Samples](https://platform.here.com/data/hrn:here:data::olp-here:here-geojson-samples/overview)
* Languages: Python
```
# Import the inspector functions and support classes
from here.inspector import inspect, new_inspector, options
from here.inspector.styles import Color, Theme
# Import other packages to run this demonstration
import json
import geopandas as gpd
import pandas as pd
from shapely.geometry import shape, Point, MultiPoint, LineString, Polygon
from here.inspector import external_basemaps
from here.platform import Platform
from here.geotiles.heretile import in_bounding_box, in_geometry
from here.geopandas_adapter import GeoPandasAdapter
```
If you have a HERE API key, you can set it as shown here. The inspect works also without an API key, but specifying a key gives you access to HERE base maps and HERE content by default.
```
# Configure your HERE API key here for the best inspector experience
options.api_key = None # Set your HERE API key in the `options.api_key` when available
```
For more information about `options` and the options available, please see the section below in this notebook.
## Sample datasets
Load datasets from various sources that will be shown using the inspector.
```
with open("sample_datasets/DEU.geo.json") as f:
data1 = json.load(f) # this is a GeoJSON FeatureCollection with only Germany
ger = data1["features"][0] # this is the GeoJSON Feature of Germany
with open("sample_datasets/FRA.geo.json") as f:
data2 = json.load(f) # this is a GeoJSON FeatureCollection with only France
fra = data2["features"][0] # this is the GeoJSON Feature of France
with open("sample_datasets/IRL.geo.json") as f:
data3 = json.load(f) # this is a GeoJSON FeatureCollection with only Ireland
irl = data3["features"][0] # this is the GeoJSON Feature of Ireland
# This is a GeoJSON FeatureCollection with Germany and France
data4 = {
"type": "FeatureCollection",
"features": [ger, fra]
}
with open("sample_datasets/countries.geo.json") as f:
data5 = json.load(f) # this is a GeoJSON FeatureCollection with all the countries of the world
rows = [
{
"id": feat["id"],
"name": feat["properties"]["name"],
"geometry": shape(feat["geometry"])
} for feat in data5["features"] if feat["properties"]["name"].lower().startswith("po")
]
gdf1 = gpd.GeoDataFrame.from_records(rows, index="id")
gs1 = gdf1.geometry
gdf1 # This is a GeoDataFrame with Poland and Portugal
```
Loading higher resolution country polygons from a HERE platform catalog
```
platform = Platform(adapter=GeoPandasAdapter())
sample_catalog = platform.get_catalog("hrn:here:data::olp-here:here-geojson-samples")
borders_layer = sample_catalog.get_layer("country-borders")
gdf2 = borders_layer.read_partitions(partition_ids=[1419, 1422])
gs2 = gdf2.geometry
gdf2 # This is a GeoDataFrame with some country polygons
# Example shapely geometries: points, linestrings and polygons
pt1 = Point(30, 50)
pt2 = Point(32, 49)
pt3 = Point(31, 52)
ls1 = LineString([(29, 49), (30, 51), (31, 49), (32, 50), (33, 51)])
ls2 = LineString([(30, 47), (31, 48), (29, 49), (31, 50), (29, 51)])
poly1 = Polygon([(28, 46), (28, 52), (29, 53), (31, 53), (33, 48), (32, 47), (28, 46)])
poly2 = MultiPoint([(40, 51), (43, 49)]).buffer(3)
```
## Inspecting datasets
The inspector is able to visualize data in different formats. Supported are:
- `gpd.GeoSeries`
- `gpd.GeoDataFrame`
- a python container, e.g. list, of shapely `BaseGeometry` objects, such as `Point`, `LineString` and `Polygon`
- one GeoJSON `FeatureCollection`, as python dictionary
- a python container, e.g. list, of GeoJSON `Feature`, each as python dictionary
Geographic features may be inspected singly or layered.
Each layer may have its own color or custom style.
Inspector can be used in 3 ways:
1. In one line, through the simplified `inspect` function.
2. Instantiating an implementation of `Inspector`, such as `IpyleafletInspector` and configuring its properties.
3. Accessing the underlaying inspector implementation via the `backend` function,
for `IpyleaftletInspector` this exposes the `ipyleaflet.Map` object that the user
can further customize to leverage all the capabilities of _ipyleaflet_.
For more information, please refer to the documentation of the `inspect` function and `Inspector` interface.
### 1. The `inspect` function and basic styling
This function opens an inspector and loads geodata in a single call. It can be used to quicly inspect the data, if no further customizations are needed.
You can visualize one or more datadasets. Each dataset may have a name and a style. Names and styles are default empty. Default style is a color automatically assigned by the inspector. You can specify a name and/or a different color, or a more complex rendering style as explained in the second part.
#### Empty inspectors
```
inspect()
# Centered in Buenos Aires
inspect(center=Point(-58.381667, -34.603333), zoom=10)
```
#### GeoSeries and GeoDataFrame
```
# Inspect one single GeoDataFrame
inspect(gdf1) # just the content
# Inspect one single GeoSeries
inspect(gs2, "An example GeoSeries") # content and name
# Inspect multiple GeoSeries and GeoDataFrames, as independent layers
inspect(layers={"A": gdf1, "B": gs2}) # content with names, default-styled
```
#### GeoJSON
```
# Inspect one single GeoJSON FeatureCollection
inspect(data4, "Central EU", Color.BLUE) # content, name and style
# Inspect one single GeoJSON Feature
inspect(fra, style=Color.RED) # unnamed, just style
```
#### Shapely geometries
```
# Inspect more shapely geometries in one layer
inspect([pt1, pt2, ls1, poly1]) # unnamed, default-styled
# Inspect shapely geometries in multiple layer
inspect(layers={
"Points and lines A": [pt1, ls1],
"Points and lines B": [pt2, pt3, ls2],
"Polygons": [poly1, poly2]
}) # named, each layer default-styled
```
#### Tiling grid
A tiling grid can be added to every inspector which will draw the borders of the HEREtile partitions given. You must specify which tiles to display by providing a list or `Series` of HEREtile partition IDs.
```
# Selected tiles
tiles = [5763, 5766, 5761]
inspect(tiles=tiles)
# Display features along with the tiles existing inside bounding box
features = [
Point(-100, 30).buffer(10),
LineString([(-110, 30), (-100, 50)]).buffer(2)
]
tiles = pd.Series(in_bounding_box(west=-110, south=35, east=-80, north=50, level=7))
inspect(features, tiles=tiles, tiles_style=Color.ORANGE)
```
#### Mixed layer types and tiling
```
# Inspect more than one layer
# each layer can be of different type, in this example
# GeoDataFrame, GeoSeries, collection of shapely geometric data
inspect(layers={
"Example GeoDataFrame": gdf1,
"Example GeoSeries": gs2,
"Example GeoJSON Feature": irl,
"Example GeoJSON FeatureCollection": data4,
"Example geometries": [pt1, pt2, ls1, ls2, poly2]
}, layers_style={
"Example geometries": Color.GRAY
}) # named, each layer default-styled, but geometries
```
## Example analysis
Show the top 10 partitions containing countries with the largest borders geometry (by data size).
This is a quick analysis performed solely on metadata. Content is downloaded to extract
country polygons and show them on a map.
```
# Get the metadata of the sample layer
mdata = borders_layer.get_partitions_metadata()
mdata.sort_values(by="data_size", ascending=False)[:10]
large_ids = mdata.sort_values(by="data_size", ascending=False)[:10].id.values
gdf = borders_layer.read_partitions(partition_ids=list(large_ids))
gdf
# Inspect the result
inspect(gdf, "10 largest partitions", Color.PINK)
```
### 2. The `Inspect` interface
You can create an inspector and fine-tune the visualization by configuring the inspector with various functions. The customization possibilities, although extended, are similar to the one provided by the `inspect` function.
You can visualize one or more datadasets and one or more tiling grids. Each appaear as layer, and layer may have a name, style, or more complex rendering style.
#### Generic styling
```
# Instantiate the default inspector
inspector = new_inspector()
# Load some data, each creates a layer
inspector.add_features(fra) # unnamed content
inspector.add_features(ger, style=Color.YELLOW) # unnamed content with style
inspector.add_features(gs2, name="Some mediterranean countries", style=Color.BLUE) # named and styled
# Further configuration
inspector.set_zoom(4)
# Show the inspector once configuration is complete
inspector.show()
```
#### Multiple tiling grids
```
# Instantiate a default inspector
inspector = new_inspector()
# Load some data, each creates a layer
inspector.add_features(fra, name="France", style=Color.BLUE)
# Define some tiles
tiles1 = pd.Series(in_bounding_box(west=-10, south=30, east=5, north=40, level=6))
tiles2 = pd.Series(in_geometry(Point(10, 45).buffer(8), level=8, fully_contained=True))
# Add the two tiling grids
inspector.add_tiles(tiles1, name="Tiling grid, level 6", style=Color.GREEN)
inspector.add_tiles(tiles2, name="Tiling grid, level 8", style=Color.YELLOW)
# Show the inspector once configuration is complete
inspector.show()
```
#### _ipyleaflet_-specific styles
Styling possibilities are not limited to generic styles and colors. Each inspector implementation supports implementation-specific styles.
For the case of the _ipyleaflet_-based inspector, `style` dictionaries as described in [ipyleaflet documentation](https://ipyleaflet.readthedocs.io/)
can be passed in place of generic styles. This includes support for `hover_style` and `point_style` as well.
A method `set_basemap` is provided to easily configure a custom _ipyleaflet_ [base map](https://ipyleaflet.readthedocs.io/en/latest/api_reference/basemaps.html).
```
# This example requires explicit use of the ipyleaflet-based implementation
from here.inspector.ipyleaflet import IpyleafletInspector
# Instantiate the ipyleaflet-based inspector
inspector = IpyleafletInspector()
# Add features with ipyleaflet-specific styles
inspector.add_features(
fra, name="France",
style={'color': 'cyan', 'radius': 8, 'fillColor': '#cc6633', 'opacity': 0.5, 'weight': 5, 'dashArray': '10', 'fillOpacity': 0.4},
hover_style={'fillColor': 'cyan', 'fillOpacity': 0.4}
)
inspector.add_features(
[pt1, pt2, pt3, ls1, poly2], name="Some geometries",
style={'color': 'magenta', 'fillColor': 'white', 'opacity': 0.8, 'weight': 3, 'dashArray': '5', 'fillOpacity': 0.8},
hover_style={'fillColor': 'yellow', 'fillOpacity': 0.4},
point_style={'radius': 10}
)
# Set a different base map
inspector.set_basemap(external_basemaps.Esri.WorldImagery)
# Show the inspector once configuration is complete
inspector.show()
```
### 3. The `backend` function to access implementation details
Via the `Inspector.backend` function it's possible to access the underlying rendering backend. This enables access to all the advanced functionalities of the rendering backend for more advanced use cases and unlimited customization. The implementation is provided already configured with theme, features, and tiling grids specified so far. Users can add content.
For `IpyleafletInspector`, this provides access to a preconfigured _ipyleaflet_ `Map` object. This in turns enables defining custom layer types, further map content, and UI widgets. For more information, please see the _ipyleaftlet_ [documentation](https://ipyleaflet.readthedocs.io/en/latest/api_reference/map.html).
## Inspector Options: default themes and colors
The inspector is configurable by changing the properties in `here.inspector.options`, that are used as default values. These include default theme and styles, also also HERE API key to enable base maps provided by HERE.
```
print(options)
```
Multiple default themes and colors are provided in the inspector:
```
for theme in Theme:
print(theme)
print("Default:", options.default_theme)
for color in Color:
print(color)
print("Default:", [c.name for c in options.default_colors])
```
The default theme is used to style the map, including the base map, and define RGB values for each color. The default colors are cycled through by the inspector to select which color to use for features and tiling grids in case no style or color is specified. Apart from setting specific themes, colors and other settings using methods of `Inspector`, the user can override these default values directly in the options (shown here only for the theme):
```
# From this moment on, all the inspector instances use dark theme
options.default_theme = Theme.DARK_MAP
inspect()
```
It's also possible to specify a different theme or sequence of colors using the methods `set_theme` and `set_colors` of `Inspector`.
```
# This resets the default to `LIGHT_MAP` to not interfere with the cells below
options.default_theme = Theme.LIGHT_MAP
```
### Blank themes
A default map is added as base map in the default theme. It's possible to use themes ending with `_BLANK` to avoid displaying a base map.
```
# Instantiate the default inspector
inspector = new_inspector()
# Load some data, each creates a layer
inspector.add_features(fra, "France", Color.BLUE)
inspector.add_features(ger, "Germany", Color.RED)
inspector.add_features(irl, "Ireland", Color.GREEN)
inspector.add_features(gs2, "Country mix", Color.GRAY)
# Configure the theme
inspector.set_theme(Theme.LIGHT_BLANK)
# Show the inspector once configuration is complete
inspector.show()
```
### Dark themes
Use themes beginning with `DARK_` for a more comfortable experience at night or dark environements. RGB values of standard colors are adjusted to the theme.
```
# Instantiate the default inspector
inspector = new_inspector()
# Load some data, each creates a layer
inspector.add_features(fra, "France", Color.BLUE)
inspector.add_features(ger, "Germany", Color.RED)
inspector.add_features(irl, "Ireland", Color.GREEN)
inspector.add_features(gs2, "Country mix", Color.GRAY)
# Configure the theme
inspector.set_theme(Theme.DARK_MAP)
# Show the inspector once configuration is complete
inspector.show()
```
### Examples of base maps
Here inspector supports multiple ways to define base maps:
- HERE base maps
- External base maps
- Custom base maps
HERE base maps and External base maps are `Tileprovider` objects provided by [xyzservices](https://github.com/geopandas/xyzservices) package. For HERE base maps you need to add your HERE API key to the global inspector options written directly in the code or taken from an environment variable.
### Example of base maps from HERE
In case a HERE API key is available, the inspector takes advantage of it authenticate MapTile API.
Just specify your own key in `here.inspector.options.api_key` as shown at the beginning of the notebook. HERE maps are visualised if the HERE API key is set, otherwise fall-back base maps are used. Below example shows all the supported HERE base maps.
```
from here.inspector import here_basemaps
here_basemaps
from IPython.display import display
if options.api_key:
inspector = IpyleafletInspector()
inspector.set_basemap(basemap=here_basemaps.normalDay)
inspector.set_zoom(14)
inspector.set_center(Point(13.4083, 52.5186))
display(inspector.show())
```
### Example of base maps from external providers
External base maps are XYZ tile providers other than HERE.
```
from here.inspector import external_basemaps
external_basemaps
inspector = IpyleafletInspector()
inspector.set_basemap(basemap=external_basemaps.OpenStreetMap.Mapnik)
inspector.show()
```
Some external basemaps need an auth token/api key.
You can check if external basemap requires authentication token using `requires_token()` method on Tileprovider.
If token/api key is required then user need to set API key given by provider in basemap object.
```
if external_basemaps.MapBox.requires_token():
external_basemaps.MapBox["accessToken"] = "YOUR-PERSONAL-TOKEN"
```
### Example of a custom base map
```
inspector = IpyleafletInspector()
basemap = {
'name': 'U.S. Geological Survey',
'url': 'https://basemap.nationalmap.gov/arcgis/rest/services/USGSTopo/MapServer/tile/{z}/{y}/{x}',
'maxZoom': 20,
'attribution': 'Tiles courtesy of the <a href="https://usgs.gov/">U.S. Geological Survey</a>'
}
inspector.set_basemap(basemap)
inspector.show()
```
<span style="float:left; margin-top:3px;"><img src="https://www.here.com/themes/custom/here_base_theme_v2/logo.svg" alt="HERE Logo" height="60" width="60"></span><span style="float:right; width:90%;"><sub><b>Copyright (c) 2020-2021 HERE Global B.V. and its affiliate(s). All rights reserved.</b>
This software, including documentation, is protected by copyright controlled by HERE. All rights are reserved. Copying, including reproducing, storing, adapting or translating, any or all of this material requires the prior written consent of HERE. This material also contains confidential information which may not be disclosed to others without the prior written consent of HERE.</sub></span>
| github_jupyter |
## Regression Coefficients are affected by regularisation
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LogisticRegression
```
## Read Data
```
data = pd.read_csv('../UNSW_Train.csv')
data.shape
data.head()
```
### Train - Test Split
```
X_train, X_test, y_train, y_test = train_test_split(
data.drop(labels=['is_intrusion'], axis=1),
data['is_intrusion'],
test_size=0.2,
random_state=0)
X_train.shape, X_test.shape
```
Fitting a few logistic regression models with decreasing values for the penalty of the regularisation
```
scaler = StandardScaler()
scaler.fit(X_train)
coefs_df = []
# we train 4 different models with regularization
penalties = [0.00005, 0.0005, 0.005, 0.05, 0.5]
for c in penalties:
logit = LogisticRegression(C=c, penalty='l2', random_state=10, max_iter=300)
logit.fit(scaler.transform(X_train), y_train)
# store the coefficients of the variables in a list
coefs_df.append(pd.Series(logit.coef_.ravel()))
# Create a dataframe with the coefficients for all
# the variables for the 4 different logistic regression models
coefs = pd.concat(coefs_df, axis=1)
coefs.columns = penalties
coefs.index = X_train.columns
coefs.head(15)
# apply log scale to the penalties (simplifies comparison)
coefs.columns = np.log(penalties)
coefs.head(15)
# plot the change in coefficients with the penalty
coefs.T.plot(figsize=(15,10), legend=False)
plt.xlabel('Penalty value')
plt.ylabel('Coefficient')
plt.title('Coefficient value vs penalty. Each line corresponds to one variable')
# now I will plot only the first 10 features for better visualisation
temp = coefs.head(15)
temp = temp.T
temp.plot(figsize=(12,8))
plt.xlabel('Penalty value')
plt.ylabel('Coefficient')
plt.title('Coefficient value vs penalty')
# plot another 10 features for visualisation
temp = coefs.tail(10)
temp = temp.T
temp.plot(figsize=(12,8))
plt.xlabel('Penalty value')
plt.ylabel('Coefficient')
plt.title('Coefficient value vs penalty')
```
The relationship between features, as inferred from the coefficients, changes depending on the level of regularisation.
```
coefs.head(15).T.columns
X_train = X_train[coefs.head(15).T.columns]
X_test = X_test[coefs.head(15).T.columns]
X_train.shape, X_test.shape
```
## Standardize Data
```
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler().fit(X_train)
X_train = scaler.transform(X_train)
```
## Classifiers
```
from sklearn import linear_model
from sklearn.naive_bayes import GaussianNB
from sklearn.ensemble import RandomForestClassifier
from sklearn.neighbors import KNeighborsClassifier
from catboost import CatBoostClassifier
```
## Metrics Evaluation
```
from sklearn.metrics import accuracy_score
from sklearn.metrics import confusion_matrix
from sklearn.metrics import roc_curve, f1_score
from sklearn import metrics
from sklearn.model_selection import cross_val_score
```
### Logistic Regression
```
%%time
clf_LR = linear_model.LogisticRegression(n_jobs=-1, random_state=42, C=25).fit(X_train, y_train)
pred_y_test = clf_LR.predict(X_test)
print('Accuracy:', accuracy_score(y_test, pred_y_test))
f1 = f1_score(y_test, pred_y_test)
print('F1 Score:', f1)
fpr, tpr, thresholds = roc_curve(y_test, pred_y_test)
print('FPR:', fpr[1])
print('TPR:', tpr[1])
```
### Naive Bayes
```
%%time
clf_NB = GaussianNB(var_smoothing=1e-08).fit(X_train, y_train)
pred_y_testNB = clf_NB.predict(X_test)
print('Accuracy:', accuracy_score(y_test, pred_y_testNB))
f1 = f1_score(y_test, pred_y_testNB)
print('F1 Score:', f1)
fpr, tpr, thresholds = roc_curve(y_test, pred_y_testNB)
print('FPR:', fpr[1])
print('TPR:', tpr[1])
```
### Random Forest
```
%%time
clf_RF = RandomForestClassifier(random_state=0,max_depth=100,n_estimators=1000).fit(X_train, y_train)
pred_y_testRF = clf_RF.predict(X_test)
print('Accuracy:', accuracy_score(y_test, pred_y_testRF))
f1 = f1_score(y_test, pred_y_testRF, average='weighted', zero_division=0)
print('F1 Score:', f1)
fpr, tpr, thresholds = roc_curve(y_test, pred_y_testRF)
print('FPR:', fpr[1])
print('TPR:', tpr[1])
```
### KNN
```
%%time
clf_KNN = KNeighborsClassifier(algorithm='ball_tree',leaf_size=1,n_neighbors=5,weights='uniform').fit(X_train, y_train)
pred_y_testKNN = clf_KNN.predict(X_test)
print('accuracy_score:', accuracy_score(y_test, pred_y_testKNN))
f1 = f1_score(y_test, pred_y_testKNN)
print('f1:', f1)
fpr, tpr, thresholds = roc_curve(y_test, pred_y_testKNN)
print('fpr:', fpr[1])
print('tpr:', tpr[1])
```
### CatBoost
```
%%time
clf_CB = CatBoostClassifier(random_state=0,depth=7,iterations=50,learning_rate=0.04).fit(X_train, y_train)
pred_y_testCB = clf_CB.predict(X_test)
print('Accuracy:', accuracy_score(y_test, pred_y_testCB))
f1 = f1_score(y_test, pred_y_testCB, average='weighted', zero_division=0)
print('F1 Score:', f1)
fpr, tpr, thresholds = roc_curve(y_test, pred_y_testCB)
print('FPR:', fpr[1])
print('TPR:', tpr[1])
```
## Model Evaluation
```
import pandas as pd, numpy as np
test_df = pd.read_csv("../UNSW_Test.csv")
test_df.shape
# Create feature matrix X and target vextor y
y_eval = test_df['is_intrusion']
X_eval = test_df.drop(columns=['is_intrusion'])
X_eval = X_eval[coefs.head(15).T.columns]
X_eval.shape
```
### Model Evaluation - Logistic Regression
```
modelLR = linear_model.LogisticRegression(n_jobs=-1, random_state=42, C=25)
modelLR.fit(X_train, y_train)
# Predict on the new unseen test data
y_evalpredLR = modelLR.predict(X_eval)
y_predLR = modelLR.predict(X_test)
train_scoreLR = modelLR.score(X_train, y_train)
test_scoreLR = modelLR.score(X_test, y_test)
print("Training accuracy is ", train_scoreLR)
print("Testing accuracy is ", test_scoreLR)
from sklearn.metrics import confusion_matrix, precision_score, recall_score, f1_score
print('Performance measures for test:')
print('--------')
print('Accuracy:', test_scoreLR)
print('F1 Score:',f1_score(y_test, y_predLR))
print('Precision Score:',precision_score(y_test, y_predLR))
print('Recall Score:', recall_score(y_test, y_predLR))
print('Confusion Matrix:\n', confusion_matrix(y_test, y_predLR))
```
### Cross validation - Logistic Regression
```
from sklearn.model_selection import cross_val_score
from sklearn import metrics
accuracy = cross_val_score(modelLR, X_eval, y_eval, cv=10, scoring='accuracy')
print("Accuracy: %0.5f (+/- %0.5f)" % (accuracy.mean(), accuracy.std() * 2))
f = cross_val_score(modelLR, X_eval, y_eval, cv=10, scoring='f1')
print("F1 Score: %0.5f (+/- %0.5f)" % (f.mean(), f.std() * 2))
precision = cross_val_score(modelLR, X_eval, y_eval, cv=10, scoring='precision')
print("Precision: %0.5f (+/- %0.5f)" % (precision.mean(), precision.std() * 2))
recall = cross_val_score(modelLR, X_eval, y_eval, cv=10, scoring='recall')
print("Recall: %0.5f (+/- %0.5f)" % (recall.mean(), recall.std() * 2))
```
### Model Evaluation - Naive Bayes
```
modelNB = GaussianNB(var_smoothing=1e-08)
modelNB.fit(X_train, y_train)
# Predict on the new unseen test data
y_evalpredNB = modelNB.predict(X_eval)
y_predNB = modelNB.predict(X_test)
train_scoreNB = modelNB.score(X_train, y_train)
test_scoreNB = modelNB.score(X_test, y_test)
print("Training accuracy is ", train_scoreNB)
print("Testing accuracy is ", test_scoreNB)
from sklearn.metrics import confusion_matrix, precision_score, recall_score, f1_score
print('Performance measures for test:')
print('--------')
print('Accuracy:', test_scoreNB)
print('F1 Score:',f1_score(y_test, y_predNB))
print('Precision Score:',precision_score(y_test, y_predNB))
print('Recall Score:', recall_score(y_test, y_predNB))
print('Confusion Matrix:\n', confusion_matrix(y_test, y_predNB))
```
### Cross validation - Naive Bayes
```
from sklearn.model_selection import cross_val_score
from sklearn import metrics
accuracy = cross_val_score(modelNB, X_eval, y_eval, cv=10, scoring='accuracy')
print("Accuracy: %0.5f (+/- %0.5f)" % (accuracy.mean(), accuracy.std() * 2))
f = cross_val_score(modelNB, X_eval, y_eval, cv=10, scoring='f1')
print("F1 Score: %0.5f (+/- %0.5f)" % (f.mean(), f.std() * 2))
precision = cross_val_score(modelNB, X_eval, y_eval, cv=10, scoring='precision')
print("Precision: %0.5f (+/- %0.5f)" % (precision.mean(), precision.std() * 2))
recall = cross_val_score(modelNB, X_eval, y_eval, cv=10, scoring='recall')
print("Recall: %0.5f (+/- %0.5f)" % (recall.mean(), recall.std() * 2))
```
### Model Evaluation - Random Forest
```
modelRF = RandomForestClassifier(random_state=0,max_depth=100,n_estimators=1000)
modelRF.fit(X_train, y_train)
# Predict on the new unseen test data
y_evalpredRF = modelRF.predict(X_eval)
y_predRF = modelRF.predict(X_test)
train_scoreRF = modelRF.score(X_train, y_train)
test_scoreRF = modelRF.score(X_test, y_test)
print("Training accuracy is ", train_scoreRF)
print("Testing accuracy is ", test_scoreRF)
from sklearn.metrics import confusion_matrix, precision_score, recall_score, f1_score
print('Performance measures for test:')
print('--------')
print('Accuracy:', test_scoreRF)
print('F1 Score:', f1_score(y_test, y_predRF, average='weighted', zero_division=0))
print('Precision Score:', precision_score(y_test, y_predRF, average='weighted', zero_division=0))
print('Recall Score:', recall_score(y_test, y_predRF, average='weighted', zero_division=0))
print('Confusion Matrix:\n', confusion_matrix(y_test, y_predRF))
```
### Cross validation - Random Forest
```
from sklearn.model_selection import cross_val_score
from sklearn import metrics
accuracy = cross_val_score(modelRF, X_eval, y_eval, cv=10, scoring='accuracy')
print("Accuracy: %0.5f (+/- %0.5f)" % (accuracy.mean(), accuracy.std() * 2))
f = cross_val_score(modelRF, X_eval, y_eval, cv=10, scoring='f1')
print("F1 Score: %0.5f (+/- %0.5f)" % (f.mean(), f.std() * 2))
precision = cross_val_score(modelRF, X_eval, y_eval, cv=10, scoring='precision')
print("Precision: %0.5f (+/- %0.5f)" % (precision.mean(), precision.std() * 2))
recall = cross_val_score(modelRF, X_eval, y_eval, cv=10, scoring='recall')
print("Recall: %0.5f (+/- %0.5f)" % (recall.mean(), recall.std() * 2))
```
### Model Evaluation - KNN
```
modelKNN = KNeighborsClassifier(algorithm='ball_tree',leaf_size=1,n_neighbors=5,weights='uniform')
modelKNN.fit(X_train, y_train)
# Predict on the new unseen test data
y_evalpredKNN = modelKNN.predict(X_eval)
y_predKNN = modelKNN.predict(X_test)
train_scoreKNN = modelKNN.score(X_train, y_train)
test_scoreKNN = modelKNN.score(X_test, y_test)
print("Training accuracy is ", train_scoreKNN)
print("Testing accuracy is ", test_scoreKNN)
from sklearn.metrics import confusion_matrix, precision_score, recall_score, f1_score
print('Performance measures for test:')
print('--------')
print('Accuracy:', test_scoreKNN)
print('F1 Score:', f1_score(y_test, y_predKNN))
print('Precision Score:', precision_score(y_test, y_predKNN))
print('Recall Score:', recall_score(y_test, y_predKNN))
print('Confusion Matrix:\n', confusion_matrix(y_test, y_predKNN))
```
### Cross validation - KNN
```
from sklearn.model_selection import cross_val_score
from sklearn import metrics
accuracy = cross_val_score(modelKNN, X_eval, y_eval, cv=10, scoring='accuracy')
print("Accuracy: %0.5f (+/- %0.5f)" % (accuracy.mean(), accuracy.std() * 2))
f = cross_val_score(modelKNN, X_eval, y_eval, cv=10, scoring='f1')
print("F1 Score: %0.5f (+/- %0.5f)" % (f.mean(), f.std() * 2))
precision = cross_val_score(modelKNN, X_eval, y_eval, cv=10, scoring='precision')
print("Precision: %0.5f (+/- %0.5f)" % (precision.mean(), precision.std() * 2))
recall = cross_val_score(modelKNN, X_eval, y_eval, cv=10, scoring='recall')
print("Recall: %0.5f (+/- %0.5f)" % (recall.mean(), recall.std() * 2))
```
### Model Evaluation - CatBoost
```
modelCB = CatBoostClassifier(random_state=0,depth=7,iterations=50,learning_rate=0.04)
modelCB.fit(X_train, y_train)
# Predict on the new unseen test data
y_evalpredCB = modelCB.predict(X_eval)
y_predCB = modelCB.predict(X_test)
train_scoreCB = modelCB.score(X_train, y_train)
test_scoreCB = modelCB.score(X_test, y_test)
print("Training accuracy is ", train_scoreCB)
print("Testing accuracy is ", test_scoreCB)
from sklearn.metrics import confusion_matrix, precision_score, recall_score, f1_score
print('Performance measures for test:')
print('--------')
print('Accuracy:', test_scoreCB)
print('F1 Score:',f1_score(y_test, y_predCB, average='weighted', zero_division=0))
print('Precision Score:',precision_score(y_test, y_predCB, average='weighted', zero_division=0))
print('Recall Score:', recall_score(y_test, y_predCB, average='weighted', zero_division=0))
print('Confusion Matrix:\n', confusion_matrix(y_test, y_predCB))
```
### Cross validation - CatBoost
```
from sklearn.model_selection import cross_val_score
from sklearn import metrics
accuracy = cross_val_score(modelCB, X_eval, y_eval, cv=10, scoring='accuracy')
f = cross_val_score(modelCB, X_eval, y_eval, cv=10, scoring='f1')
precision = cross_val_score(modelCB, X_eval, y_eval, cv=10, scoring='precision')
recall = cross_val_score(modelCB, X_eval, y_eval, cv=10, scoring='recall')
print("Accuracy: %0.5f (+/- %0.5f)" % (accuracy.mean(), accuracy.std() * 2))
print("F1 Score: %0.5f (+/- %0.5f)" % (f.mean(), f.std() * 2))
print("Precision: %0.5f (+/- %0.5f)" % (precision.mean(), precision.std() * 2))
print("Recall: %0.5f (+/- %0.5f)" % (recall.mean(), recall.std() * 2))
```
| github_jupyter |
# Just Euler's method
```
from IPython.core.display import HTML
css_file = 'https://raw.githubusercontent.com/ngcm/training-public/master/ipython_notebook_styles/ngcmstyle.css'
HTML(url=css_file)
```
$$
\newcommand{\dt}{\Delta t}
\newcommand{\udt}[1]{u^{({#1})}(T)}
\newcommand{\Edt}[1]{E^{({#1})}}
\newcommand{\uone}[1]{u_{1}^{({#1})}}
$$
In the previous cases we've focused on the *behaviour* of the algorithm: whether it will give the correct answer in the limit, or whether it converges as expected. This is really what you want to do: you're trying to do science, to get an answer, and so implementing the precise algorithm should be secondary. If you are trying to implement a precise algorithm, it should be because of its (expected) behaviour, and so you should be testing for that!
However, let's put that aside and see if we can work out how to test whether we've implemented exactly the algorithm we want: Euler's method. Checking convergence alone is not enough: the [Backwards Euler method](http://en.wikipedia.org/wiki/Backward_Euler_method) has identical convergence behaviour, as do whole families of other methods. We need a check that characterizes the method uniquely.
The *local truncation error* $\Edt{\dt}$ would be exactly such a check. This is the error produced by a single step from exact data, eg
$$
\begin{equation}
\Edt{\dt} = u_1 - u(\dt).
\end{equation}
$$
For Euler's method we have
$$
\begin{equation}
u_{n+1} = u_n + \dt f(t_n, u_n)
\end{equation}
$$
and so
$$
\begin{equation}
\Edt{\dt} = \left| u_0 + \dt f(0, u_0) - u(\dt) \right| = \left| \frac{\dt^2}{2} \left. u''\right|_{t=0} \right| + {\cal O}(\dt^3).
\end{equation}
$$
This is all well and good, but we don't know the exact solution (in principle) at any point other than $t=0$, so cannot compute $u(\dt)$, so cannot compute $\Edt{\dt}$. We only know $\uone{\dt}$ for whichever values of $\dt$ we wish to compute.
We can use repeated Richardson extrapolation to get the solution $u(\dt)$ to sufficient accuracy, however. On the *assumption* that the algorithm is first order (we can use the previous techniques to check this), we can use Richardson extrapolation to repeatedly remove the highest order error terms. We can thus find the local truncation errors.
```
from math import sin, cos, log, ceil
import numpy
from matplotlib import pyplot
%matplotlib inline
from matplotlib import rcParams
rcParams['font.family'] = 'serif'
rcParams['font.size'] = 16
# model parameters:
g = 9.8 # gravity in m s^{-2}
v_t = 30.0 # trim velocity in m s^{-1}
C_D = 1/40. # drag coefficient --- or D/L if C_L=1
C_L = 1.0 # for convenience, use C_L = 1
### set initial conditions ###
v0 = v_t # start at the trim velocity (or add a delta)
theta0 = 0.0 # initial angle of trajectory
x0 = 0.0 # horizotal position is arbitrary
y0 = 1000.0 # initial altitude
def f(u):
"""Returns the right-hand side of the phugoid system of equations.
Parameters
----------
u : array of float
array containing the solution at time n.
Returns
-------
dudt : array of float
array containing the RHS given u.
"""
v = u[0]
theta = u[1]
x = u[2]
y = u[3]
return numpy.array([-g*sin(theta) - C_D/C_L*g/v_t**2*v**2,
-g*cos(theta)/v + g/v_t**2*v,
v*cos(theta),
v*sin(theta)])
def euler_step(u, f, dt):
"""Returns the solution at the next time-step using Euler's method.
Parameters
----------
u : array of float
solution at the previous time-step.
f : function
function to compute the right hand-side of the system of equation.
dt : float
time-increment.
Returns
-------
u_n_plus_1 : array of float
approximate solution at the next time step.
"""
return u + dt * f(u)
T_values = numpy.array([0.001*2**(i) for i in range(10)])
lte_values = numpy.zeros_like(T_values)
for j, T in enumerate(T_values):
dt_values = numpy.array([T*2**(i-8) for i in range(8)])
v_values = numpy.zeros_like(dt_values)
for i, dt in enumerate(dt_values):
N = int(T/dt)+1
t = numpy.linspace(0.0, T, N)
u = numpy.empty((N, 4))
u[0] = numpy.array([v0, theta0, x0, y0])
for n in range(N-1):
u[n+1] = euler_step(u[n], f, dt)
v_values[i] = u[-1,0]
v_next = v_values
for s in range(1, len(v_values-1)):
v_next = (2**s*v_next[1:]-v_next[0:-1])/(2**s-1)
lte_values[j] = abs(v_values[0]-v_next)
lte_values
```
We now have four values for the local truncation error. We can thus compute the convergence rate of the local truncation error itself (which should be two), and check that it is close enough to the expected value using the previous techniques:
```
s_m = numpy.zeros(2)
for i in range(2):
s_m[i] = log(abs((lte_values[2+i]-lte_values[1+i])/
(lte_values[1+i]-lte_values[0+i]))) / log(2.0)
print("Measured convergence rate (base dt {}) is {:.6g} (error is {:.4g}).".format(
T_values[i], s_m[i], abs(s_m[i]-2)))
print("Convergence error has reduced by factor {:.4g}.".format(
abs(s_m[0]-2)/abs(s_m[1]-2)))
```
So the error has gone down considerably, and certainly $0.51 < 2/3$, so the convergence rate of the local truncation error is close enough to 2.
However, that alone isn't enough to determine that this really is Euler's method: as noted above, the convergence rate of the local truncation error isn't the key point: the key point is that we can predict its *actual value* as
$$
\begin{equation}
\Edt{\dt} = \frac{\dt^2}{2} \left| \left. u''\right|_{t=0} \right| + {\cal O}(\dt^3) = \frac{\dt^2}{2} \left| \left( \left. \frac{\partial f}{\partial t} \right|_{t=0} + f(0, u_0) \left. \frac{\partial f}{\partial u} \right|_{t=0, u=u_0} \right) \right|.
\end{equation}
$$
For the specific problem considered here we have
$$
\begin{equation}
u = \begin{pmatrix} v \\ \theta \\ x \\ y \end{pmatrix}, \quad f = \begin{pmatrix} -g\sin \theta - \frac{C_D}{C_L} \frac{g}{v_t^2} v^2 \\ -\frac{g}{v}\cos \theta + \frac{g}{v_t^2} v \\ v \cos \theta \\ v \sin \theta \end{pmatrix}.
\end{equation}
$$
We note that $f$ does not explicitly depend on $t$ (so $\partial f / \partial t \equiv 0$), and that the values of the parameters $g, C_D, C_L$ and $v_t$ are given above, along with the initial data $u_0 = (v_0, \theta_0, x_0, y_0)$.
So, let's find what the local truncation error should be.
```
import sympy
sympy.init_printing()
v, theta, x, y, g, CD, CL, vt, dt = sympy.symbols('v, theta, x, y, g, C_D, C_L, v_t, {\Delta}t')
u = sympy.Matrix([v, theta, x, y])
f = sympy.Matrix([-g*sympy.sin(theta)-CD/CL*g/vt**2*v**2,
-g/v*sympy.cos(theta)+g/vt**2*v,
v*sympy.cos(theta),
v*sympy.sin(theta)])
dfdu = f.jacobian(u)
lte=dt**2/2*dfdu*f
lte_0=lte.subs([(g,9.8),(vt,30.0),(CD,1.0/40.0),(CL,1.0),(v,30.0),(theta,0.0),(x,0.0),(y,1000.0)])
lte_0
```
So let us check the local truncation error values, which are computed for `v`:
```
lte_exact = float(lte_0[0]/dt**2)
lte_values/T_values**2
```
These are indeed converging towards $0.002 \dt^2$ as they should. To check this quantitatively, we use that our model is
$$
\begin{equation}
\Edt{\dt} = \alpha \dt^2 + {\cal O}(\dt^3),
\end{equation}
$$
with the exact value $\alpha_e \simeq 0.002$. So we can use our usual Richardson extrapolation methods applied to $\Edt{\dt}/\dt^2$, to get a measured value for $\alpha$ with an error interval:
$$
\begin{equation}
\alpha_m = \frac{8\Edt{\dt} - \Edt{2\dt} \pm \left| \Edt{\dt} - \Edt{2\dt} \right|}{4\dt^2}.
\end{equation}
$$
```
for i in range(len(lte_values)-1):
Edt = lte_values[i]
E2dt = lte_values[i+1]
dt = T_values[i]
err1 = abs(Edt - E2dt)
a_lo = (8.0*Edt - E2dt - err1)/(4.0*dt**2)
a_hi = (8.0*Edt - E2dt + err1)/(4.0*dt**2)
print("Base dt={:.4g}: the measured alpha is in [{:.5g}, {:.5g}]".format(
dt, a_lo, a_hi))
print("Does this contain the exact value? {}".format(
a_lo <= lte_exact <= a_hi))
```
So, to the limits that we can measure the local truncation error, we have implemented Euler's method.
| github_jupyter |
## Save mutation profiles (and other associated data) to files, and make some combined plots
```
%run "ComputeUniMutationalFractions.ipynb"
from collections import defaultdict
def get_all_mutation_profile_data(seq, geneloc="../seqs/genes"):
"""Partially copied from get_mutation_profile(), but extended."""
num_mutations_to_freq = defaultdict(int)
num_mutations_to_fvals = defaultdict(list)
num_mutations_to_avgf = defaultdict(int)
num_mutations_to_unimutational_pos_ct = defaultdict(int)
num_mutations_to_unimutational_pos_ct_frac = defaultdict(int)
# Number of CP 1/2/3 and non-coding positions with N mutations
# (You can think of these as sort of individual mutation profiles limited to specific "types" of
# positions in a genome)
num_mutations_to_cp1ct = defaultdict(int)
num_mutations_to_cp2ct = defaultdict(int)
num_mutations_to_cp3ct = defaultdict(int)
num_mutations_to_ncpct = defaultdict(int)
genes = parse_sco("{}/{}.sco".format(geneloc, seq))
# maps positions to a list of the CPs they are. (It's a list, and not just an integer, b/c of overlapping
# genes -- a given position could be in CP 1 of one gene and CP 2 of an overlapping gene, for example...
# we could also just filter positions shared in multiple genes, but the other analyses here don't do that
# so for consistency's sake we continue to "double-count" positions sometimes)
pos2cptypes = defaultdict(set)
# TODO: abstract gene iteration to helper function; I've reused this code a lot in this notebook
for gene in genes.itertuples():
i = 1
# (note that positions returned by get_pos_interval_from_gene() are ints, while seq2pos2...
# stuff use string positions. This is wack and ideally should be made easier to work with)
for pos in get_pos_interval_from_gene(gene):
pos2cptypes[str(pos)].add(i)
if i == 1: i = 2
elif i == 2: i = 3
elif i == 3: i = 1
else: raise ValueError("Codon position modulo calculation is broken :|")
if i != 1: raise ValueError("Gene length not divisible by 3")
for pos in range(1, seq2len[seq] + 1):
# ignore uncovered places. Could also ignore places with less than some threshold coverage
# (e.g. 5) if desired
if pileup.get_cov(seq2pos2pileup[seq][pos]) == 0:
continue
# Most important part - figure out how many mutations this position has
mismatch_cts = pileup.get_mismatch_cts(seq2pos2pileup[seq][pos])
mutct = sum(mismatch_cts)
# Update frequency (main point of the mutation profile)
num_mutations_to_freq[mutct] += 1
# Update CP / non-coding pos cts
cptypes = pos2cptypes[pos]
if len(cptypes) == 0:
num_mutations_to_ncpct[mutct] += 1
else:
# NOTE: This is an inelegant way of writing this - ideally these would all be one structure
for cp in cptypes:
if cp == 1: num_mutations_to_cp1ct[mutct] += 1
elif cp == 2: num_mutations_to_cp2ct[mutct] += 1
elif cp == 3: num_mutations_to_cp3ct[mutct] += 1
else: raise ValueError("Unrecognized CP type")
if seq2pos2f[seq][pos] is None:
# This means f is undefined because there are no alternate bases at this position.
if mutct == 0:
num_mutations_to_fvals[mutct].append(seq2pos2f[seq][pos])
else:
# Previously, this case could happen accidentally due to deletions messing with things. Now it
# shouldn't ever happen.
raise ValueError("Found undefined f at position {} in {} with alternate bases".format(pos, seq))
else:
num_mutations_to_fvals[mutct].append(seq2pos2f[seq][pos])
if mutct == max(mismatch_cts):
num_mutations_to_unimutational_pos_ct[mutct] += 1
# Now that we've gone through all positions in this sequence, update some aggregate stats (avg.
# uni-mutational fraction [aka average f-value]; fraction of uni-mutational positions).
# Note that we iterate through this range - not just the keys of num_mutations_to_freq - because
# we want to look over the places where there are 0 mutations, which are implicitly not included in
# num_mutations_to_freq since it's a defaultdict
for i in range(0, max(num_mutations_to_freq.keys()) + 1):
if num_mutations_to_freq[i] > 0:
if None in num_mutations_to_fvals[i]:
if i == 0:
avg_f = "N/A"
else:
raise ValueError("Something is bad")
else:
if len(num_mutations_to_fvals[i]) == 0:
raise ValueError("Something went horribly wrong... deletion error again?")
else:
avg_f = sum(num_mutations_to_fvals[i]) / len(num_mutations_to_fvals[i])
else:
avg_f = "N/A"
# I'm aware that the use of an underscore in "avg_f" should be more consistent; sorry ._.
num_mutations_to_avgf[i] = avg_f
# This is the fraction of positions with this many mutations which are uni-mutational.
# So, 100% of positions with exactly one mutation are uni-mutational (so this is 1),
# and so on. Of course, if there are 0 positions with a given number of mutations,
# then we'd have to divide by 0 to compute this -- so we just call it "N/A" for these rows.
if num_mutations_to_freq[i] > 0:
num_mutations_to_unimutational_pos_ct_frac[i] = num_mutations_to_unimutational_pos_ct[i] / num_mutations_to_freq[i]
else:
num_mutations_to_unimutational_pos_ct_frac[i] = "N/A"
data = {
"num_mutations_to_freq": num_mutations_to_freq,
"num_mutations_to_cp1ct": num_mutations_to_cp1ct,
"num_mutations_to_cp2ct": num_mutations_to_cp2ct,
"num_mutations_to_cp3ct": num_mutations_to_cp3ct,
"num_mutations_to_ncpct": num_mutations_to_ncpct,
"num_mutations_to_unimutational_pos_ct": num_mutations_to_unimutational_pos_ct,
"num_mutations_to_unimutational_pos_ct_frac": num_mutations_to_unimutational_pos_ct_frac,
"num_mutations_to_avg_unimutational_fraction": num_mutations_to_avgf,
}
return data
import os
def save_mutation_profile(seq, output_dir="mutation-profiles"):
data = get_all_mutation_profile_data(seq)
out_fn = os.path.join(output_dir, "{}_mutation_profile.tsv".format(seq2name[seq]))
with open(out_fn, "w") as tsvf:
headerline = "\t".join((
"Number of mutations",
"Number of positions with this many mutations",
"Number of CP 1 positions with this many mutations",
"Number of CP 2 positions with this many mutations",
"Number of CP 3 positions with this many mutations",
"Number of non-coding positions with this many mutations",
"Number of uni-mutational positions with this many mutations",
"Fraction of positions which are uni-mutational",
"Average uni-mutational fraction of positions with this many mutations"
)) + "\n"
tsvf.write(headerline)
for i in range(0, max(data["num_mutations_to_freq"].keys()) + 1):
tsvf.write("{}\t{}\t{}\t{}\t{}\t{}\t{}\t{}\t{}\n".format(
i,
data["num_mutations_to_freq"][i],
data["num_mutations_to_cp1ct"][i],
data["num_mutations_to_cp2ct"][i],
data["num_mutations_to_cp3ct"][i],
data["num_mutations_to_ncpct"][i],
data["num_mutations_to_unimutational_pos_ct"][i],
data["num_mutations_to_unimutational_pos_ct_frac"][i],
data["num_mutations_to_avg_unimutational_fraction"][i],
))
```
## Save the output files!
```
for seq in SEQS:
save_mutation_profile(seq)
```
## Make a few extra plots
### Plot # of mutations against average unimutational fraction
```
fig, axes = pyplot.subplots(1, 3, sharey=True)
xlim_max = 200
for col, seq in enumerate(SEQS):
n2a = get_all_mutation_profile_data(seq)["num_mutations_to_avg_unimutational_fraction"]
for i in range(0, xlim_max + 1):
if type(n2a[i]) != str:
axes[col].scatter(i, n2a[i], c=SCATTERPLOT_PT_COLOR)
if col == 1:
axes[col].set_xlabel("Number of mutations")
if col == 0:
axes[col].set_ylabel("Average uni-mutational fraction")
axes[col].set_xlim(0)
axes[col].set_ylim(0.3, 1.05)
axes[col].set_title(seq2name[seq])
fig.suptitle(
"Average uni-mutational fraction for all positions with $x$ mutations\n(x-axis truncated to {})".format(xlim_max),
fontsize="18",
y=1.07
)
fig.set_size_inches(20, 5)
fig.savefig("figs/avg-unimut-mutation-profiles.png", bbox_inches="tight")
```
### Plot mutation profiles with # mutations against # uni-mutational positions
```
# two poss variants of these, one with an x-max of 200 (to focus on "low" numbers of mutations)
# and another with no x-max. for now we're just doing the 200 one, but could tile both like in the mutation
# profiles fig if desired -- use the for loop below to get that started
#
# for xmax in [200, None]:
xmax = 200
fig, axes = pyplot.subplots(1, 3, sharey=True)
for col, seq in enumerate(SEQS):
data = get_all_mutation_profile_data(seq)
largest_mutct = max(data["num_mutations_to_freq"].keys()) if xmax is None else xmax
for i in range(0, largest_mutct + 1):
# Only plot #s of mutations with at least one occurrence
if data["num_mutations_to_freq"][i] > 0:
axes[col].scatter(i, data["num_mutations_to_freq"][i], c=SCATTERPLOT_PT_COLOR,
label="" if i > 0 else "Total Number of Positions")
axes[col].scatter(i, data["num_mutations_to_unimutational_pos_ct"][i], c="#e24a33",
label="" if i > 0 else "Number of Uni-Mutational Positions")
# Need to use symlog in order to be kosher with zeroes
axes[col].set_yscale("symlog")
axes[col].set_ylim(0)
# only set xlabel for middlest plot
if col == 1:
axes[col].set_xlabel("Number of mutations")
ylabel = "How many (uni-mutational) positions have\nthis many mutations?"
ylabel += " (log$_{10}$ scale)"
# only set ylabel for leftmost plot
if col == 0:
axes[col].set_ylabel(ylabel)
# title = "{}: Mutation profile".format(seq2name[seq])
# if xmax is not None: title += "\n(x-axis truncated to {})".format(xmax)
axes[col].set_title(seq2name[seq])
# only set legend for rightmost plot
if col == 2:
axes[col].legend()
fig.suptitle(
"Mutation profiles showing numbers of uni-mutational positions\n(x-axis truncated to {})".format(xlim_max),
fontsize="18",
y=1.07
)
fig.set_size_inches(20, 5)
fig.savefig("figs/num-unimut-pos-mutation-profiles.png", bbox_inches="tight")
```
| github_jupyter |
<i>Copyright (c) Microsoft Corporation. All rights reserved.</i>
<i>Licensed under the MIT License.</i>
# Building a Real-time Recommendation API
This reference architecture shows the full lifecycle of building a recommendation system. It walks through the creation of appropriate azure resources, training a recommendation model using Azure Databricks and deploying it as an API. It uses Azure Cosmos DB, Azure Machine Learning, and Azure Kubernetes Service.
This architecture can be generalized for many recommendation engine scenarios, including recommendations for products, movies, and news.
### Architecture

**Scenario**: A media organization wants to provide movie or video recommendations to its users. By providing personalized recommendations, the organization meets several business goals, including increased click-through rates, increased engagement on site, and higher user satisfaction.
In this reference, we train and deploy a real-time recommender service API that can provide the top 10 movie recommendations for a given user.
### Components
This architecture consists of the following key components:
* [Azure Databricks](https://docs.microsoft.com/en-us/azure/azure-databricks/what-is-azure-databricks) is used as a development environment to prepare input data and train the recommender model on a Spark cluster. Azure Databricks also provides an interactive workspace to run and collaborate on notebooks for any data processing or machine learning tasks.
* [Azure Kubernetes Service](https://docs.microsoft.com/en-us/azure/aks/intro-kubernetes)(AKS) is used to deploy and operationalize a machine learning model service API on a Kubernetes cluster. AKS hosts the containerized model, providing scalability that meets throughput requirements, identity and access management, and logging and health monitoring.
* [Azure Cosmos DB](https://docs.microsoft.com/en-us/azure/cosmos-db/introduction) is a globally distributed database service used to store the top 10 recommended movies for each user. Azure Cosmos DB is ideal for this scenario as it provides low latency (10 ms at 99th percentile) to read the top recommended items for a given user.
* [Azure Machine Learning Service](https://docs.microsoft.com/en-us/azure/machine-learning/service/) is a service used to track and manage machine learning models, and then package and deploy these models to a scalable Azure Kubernetes Service environment.
### Table of Contents.
0. [File Imports](#0-File-Imports)
1. [Service Creation](#1-Service-Creation)
2. [Training](#2-Training)
3. [Operationalization](#3.-Operationalize-the-Recommender-Service)
## Setup
This notebook should be run on Azure Databricks. To import this notebook into your Azure Databricks Workspace, see instructions [here](https://docs.azuredatabricks.net/user-guide/notebooks/notebook-manage.html#import-a-notebook).
Setup for Azure Databricks should be completed by following the appropriate sections in the repository's [SETUP file](../../SETUP.md).
Please note: This notebook **REQUIRES** that you add the dependencies to support operationalization. See the [SETUP file](../../SETUP.md) for details.
## 0 File Imports
```
import numpy as np
import os
import pandas as pd
import pprint
import shutil
import time, timeit
import urllib
import yaml
import json
import uuid
import matplotlib
import matplotlib.pyplot as plt
from azure.common.client_factory import get_client_from_cli_profile
from azure.mgmt.compute import ComputeManagementClient
import azure.mgmt.cosmosdb
import azureml.core
from azureml.core import Workspace
from azureml.core.run import Run
from azureml.core.experiment import Experiment
from azureml.core.model import Model
from azureml.core.image import ContainerImage
from azureml.core.compute import AksCompute, ComputeTarget
from azureml.core.webservice import Webservice, AksWebservice
import pydocumentdb
import pydocumentdb.document_client as document_client
import pyspark
from pyspark.ml.feature import StringIndexer
from pyspark.ml.recommendation import ALS
from pyspark.sql import Row
from pyspark.sql.types import StructType, StructField
from pyspark.sql.types import StringType, FloatType, IntegerType, LongType
from reco_utils.dataset import movielens
from reco_utils.dataset.cosmos_cli import find_collection, read_collection, read_database, find_database
from reco_utils.dataset.spark_splitters import spark_random_split
from reco_utils.evaluation.spark_evaluation import SparkRatingEvaluation, SparkRankingEvaluation
print("PySpark version:", pyspark.__version__)
print("Azure SDK version:", azureml.core.VERSION)
```
## 1 Service Creation
Modify the **Subscription ID** to the subscription you would like to deploy to.
#### Services created by this notebook:
1. [Azure ML Service](https://docs.databricks.com/user-guide/libraries.html)
1. [Azure Cosmos DB](https://azure.microsoft.com/en-us/services/cosmos-db/)
1. [Azure Container Registery](https://docs.microsoft.com/en-us/azure/container-registry/)
1. [Azure Container Instances](https://docs.microsoft.com/en-us/azure/container-instances/)
1. [Azure Application Insights](https://azure.microsoft.com/en-us/services/monitor/)
1. [Azure Storage](https://docs.microsoft.com/en-us/azure/storage/common/storage-account-overview)
1. [Azure Key Vault](https://azure.microsoft.com/en-us/services/key-vault/)
1. [Azure Kubernetes Service (AKS)](https://azure.microsoft.com/en-us/services/kubernetes-service/)
```
# Select the services names
short_uuid = str(uuid.uuid4())[:4]
prefix = "reco" + short_uuid
data = "mvl"
algo = "als"
# location to store the secrets file for cosmosdb
secrets_path = '/dbfs/FileStore/dbsecrets.json'
ws_config_path = '/dbfs/FileStore'
# Add your subscription ID
subscription_id = ""
# Resource group and workspace
resource_group = prefix + "_" + data
workspace_name = prefix + "_"+data+"_aml"
workspace_region = "westus2"
print("Resource group:", resource_group)
# Columns
userCol = "UserId"
itemCol = "MovieId"
ratingCol = "Rating"
# CosmosDB
location = workspace_region
account_name = resource_group + "-ds-sql"
# account_name for CosmosDB cannot have "_" and needs to be less than 31 chars
account_name = account_name.replace("_","-")[0:min(31,len(prefix))]
DOCUMENTDB_DATABASE = "recommendations"
DOCUMENTDB_COLLECTION = "user_recommendations_" + algo
# AzureML
history_name = 'spark-ml-notebook'
model_name = data+"-"+algo+"-reco.mml" #NOTE: The name of a asset must be only letters or numerals, not contain spaces, and under 30 characters
service_name = data + "-" + algo
experiment_name = data + "_"+ algo +"_Experiment"
# Name here must be <= 16 chars and only include letters, numbers and "-"
aks_name = prefix.replace("_","-")[0:min(12,len(prefix))] + '-aks'
# add a name for the container
container_image_name = '-'.join([data, algo])
train_data_path = data + "Train"
test_data_path = data + "Test"
```
### 1.1 Import or create the AzureML Workspace.
This command will check if the AzureML Workspace exists or not, and will create the workspace if it doesn't exist.
```
ws = Workspace.create(name = workspace_name,
subscription_id = subscription_id,
resource_group = resource_group,
location = workspace_region,
exist_ok=True)
# persist the subscription id, resource group name, and workspace name in aml_config/config.json.
ws.write_config(ws_config_path)
```
### 1.2 Create a Cosmos DB resource to store recommendation results:
```
## explicitly pass subscription_id in case user has multiple subscriptions
client = get_client_from_cli_profile(azure.mgmt.cosmosdb.CosmosDB,
subscription_id=subscription_id)
async_cosmosdb_create = client.database_accounts.create_or_update(
resource_group,
account_name,
{
'location': location,
'locations': [{
'location_name': location
}]
}
)
account = async_cosmosdb_create.result()
my_keys = client.database_accounts.list_keys(
resource_group,
account_name
)
master_key = my_keys.primary_master_key
endpoint = "https://" + account_name + ".documents.azure.com:443/"
#db client
client = document_client.DocumentClient(endpoint, {'masterKey': master_key})
if find_database(client, DOCUMENTDB_DATABASE) == False:
db = client.CreateDatabase({ 'id': DOCUMENTDB_DATABASE })
else:
db = read_database(client, DOCUMENTDB_DATABASE)
# Create collection options
options = {
'offerThroughput': 11000
}
# Create a collection
collection_definition = { 'id': DOCUMENTDB_COLLECTION, 'partitionKey': {'paths': ['/id'],'kind': 'Hash'} }
if find_collection(client,DOCUMENTDB_DATABASE, DOCUMENTDB_COLLECTION) ==False:
collection = client.CreateCollection(db['_self'], collection_definition, options)
else:
collection = read_collection(client, DOCUMENTDB_DATABASE, DOCUMENTDB_COLLECTION)
secrets = {
"Endpoint": endpoint,
"Masterkey": master_key,
"Database": DOCUMENTDB_DATABASE,
"Collection": DOCUMENTDB_COLLECTION,
"Upsert": "true"
}
with open(secrets_path, "w") as file:
json.dump(secrets, file)
```
## 2 Training
Next, we will train an [Alternating Least Squares model](https://spark.apache.org/docs/2.2.0/ml-collaborative-filtering.html) is trained using the [MovieLens](https://grouplens.org/datasets/movielens/) dataset.
```
# top k items to recommend
TOP_K = 10
# Select Movielens data size: 100k, 1m, 10m, or 20m
MOVIELENS_DATA_SIZE = '100k'
```
### 2.1 Download the MovieLens dataset
```
# Note: The DataFrame-based API for ALS currently only supports integers for user and item ids.
schema = StructType(
(
StructField("UserId", IntegerType()),
StructField("MovieId", IntegerType()),
StructField("Rating", FloatType()),
StructField("Timestamp", LongType()),
)
)
data = movielens.load_spark_df(spark, size=MOVIELENS_DATA_SIZE, schema=schema, dbutils=dbutils)
data.show()
```
### 2.2 Split the data into train, test
There are several ways of splitting the data: random, chronological, stratified, etc., each of which favors a different real-world evaluation use case. We will split randomly in this example – for more details on which splitter to choose, consult [this guide](https://github.com/Microsoft/Recommenders/blob/master/notebooks/01_data/data_split.ipynb).
```
train, test = spark_random_split(data, ratio=0.75, seed=42)
print ("N train", train.cache().count())
print ("N test", test.cache().count())
```
### 2.3 Train the ALS model on the training data, and get the top-k recommendations for our testing data
To predict movie ratings, we use the rating data in the training set as users' explicit feedbacks. The hyperparameters used to estimate the model are set based on [this page](http://mymedialite.net/examples/datasets.html).
Under most circumstances, you would explore the hyperparameters and choose an optimal set based on some criteria. For additional details on this process, please see additional information in the deep dives [here](../04_model_select_and_optimize/hypertune_spark_deep_dive.ipynb).
```
header = {
"userCol": "UserId",
"itemCol": "MovieId",
"ratingCol": "Rating",
}
als = ALS(
rank=10,
maxIter=15,
implicitPrefs=False,
alpha=0.1,
regParam=0.05,
coldStartStrategy='drop',
nonnegative=True,
**header
)
model = als.fit(train)
```
In the movie recommendation use case, recommending movies that have been rated by the users do not make sense. Therefore, the rated movies are removed from the recommended items.
In order to achieve this, we recommend all movies to all users, and then remove the user-movie pairs that exist in the training datatset.
```
# Get the cross join of all user-item pairs and score them.
users = train.select('UserId').distinct()
items = train.select('MovieId').distinct()
user_item = users.crossJoin(items)
dfs_pred = model.transform(user_item)
dfs_pred.show()
# Remove seen items.
dfs_pred_exclude_train = dfs_pred.alias("pred").join(
train.alias("train"),
(dfs_pred['UserId'] == train['UserId']) & (dfs_pred['MovieId'] == train['MovieId']),
how='outer'
)
top_all = dfs_pred_exclude_train.filter(dfs_pred_exclude_train["train.Rating"].isNull()) \
.select('pred.' + 'UserId', 'pred.' + 'MovieId', 'pred.' + "prediction")
top_all.show()
```
### 2.4 Evaluate how well ALS performs
Evaluate model performance using metrics such as Precision@K, Recall@K, [MAP](https://en.wikipedia.org/wiki/Evaluation_measures_\(information_retrieval\)) or [nDCG](https://en.wikipedia.org/wiki/Discounted_cumulative_gain). For a full guide on what metrics to evaluate your recommender with, consult [this guide](https://github.com/Microsoft/Recommenders/blob/master/notebooks/03_evaluate/evaluation.ipynb).
```
test.show()
rank_eval = SparkRankingEvaluation(test, top_all, k = TOP_K, col_user="UserId", col_item="MovieId",
col_rating="Rating", col_prediction="prediction",
relevancy_method="top_k")
# Evaluate Ranking Metrics
print("Model:\tALS",
"Top K:\t%d" % rank_eval.k,
"MAP:\t%f" % rank_eval.map_at_k(),
"NDCG:\t%f" % rank_eval.ndcg_at_k(),
"Precision@K:\t%f" % rank_eval.precision_at_k(),
"Recall@K:\t%f" % rank_eval.recall_at_k(), sep='\n')
# Evaluate Rating Metrics
prediction = model.transform(test)
rating_eval = SparkRatingEvaluation(test, prediction, col_user="UserId", col_item="MovieId",
col_rating="Rating", col_prediction="prediction")
print("Model:\tALS rating prediction",
"RMSE:\t%.2f" % rating_eval.rmse(),
"MAE:\t%f" % rating_eval.mae(),
"Explained variance:\t%f" % rating_eval.exp_var(),
"R squared:\t%f" % rating_eval.rsquared(), sep='\n')
```
### 2.5 Save the model
```
model.write().overwrite().save(model_name)
model_local = "file:" + os.getcwd() + "/" + model_name
dbutils.fs.cp(model_name, model_local, True)
```
## 3. Operationalize the Recommender Service
Once the model is built with desirable performance, it will be operationalized to run as a REST endpoint to be utilized by a real time service. We will utilize [Azure Cosmos DB](https://azure.microsoft.com/en-us/services/cosmos-db/), [Azure Machine Learning Service](https://azure.microsoft.com/en-us/services/machine-learning-service/), and [Azure Kubernetes Service](https://docs.microsoft.com/en-us/azure/aks/intro-kubernetes) to operationalize the recommender service.
### 3.1 Create a look-up for Recommendations in Cosmos DB
First, the Top-10 recommendations for each user as predicted by the model are stored as a lookup table in Cosmos DB. At runtime, the service will return the Top-10 recommendations as precomputed and stored in Cosmos DB:
```
with open(secrets_path) as json_data:
writeConfig = json.load(json_data)
recs = model.recommendForAllUsers(10)
recs.withColumn("id",recs[userCol].cast("string")).select("id", "recommendations."+ itemCol)\
.write.format("com.microsoft.azure.cosmosdb.spark").mode('overwrite').options(**writeConfig).save()
```
### 3.2 Configure Azure Machine Learning
Next, Azure Machine Learning Service is used to create a model scoring image and deploy it to Azure Kubernetes Service as a scalable containerized service. To achieve this, a **scoring script** and an **environment config** should be created. The following shows the content of the two files.
In the scoring script, we make a call to Cosmos DB to lookup the top 10 movies to recommend given an input User ID:
```
score_sparkml = """
import json
def init(local=False):
global client, collection
try:
# Query them in SQL
import pydocumentdb.document_client as document_client
MASTER_KEY = '{key}'
HOST = '{endpoint}'
DATABASE_ID = "{database}"
COLLECTION_ID = "{collection}"
database_link = 'dbs/' + DATABASE_ID
collection_link = database_link + '/colls/' + COLLECTION_ID
client = document_client.DocumentClient(HOST, {'masterKey': MASTER_KEY})
collection = client.ReadCollection(collection_link=collection_link)
except Exception as e:
collection = e
def run(input_json):
try:
import json
id = json.loads(json.loads(input_json)[0])['id']
query = {'query': 'SELECT * FROM c WHERE c.id = "' + str(id) +'"' } #+ str(id)
options = {'partitionKey':str(id)}
document_link = 'dbs/{DOCUMENTDB_DATABASE}/colls/{DOCUMENTDB_COLLECTION}/docs/{0}'.format(id)
result = client.ReadDocument(document_link, options);
except Exception as e:
result = str(e)
return json.dumps(str(result)) #json.dumps({{"result":result}})
"""
with open(secrets_path) as json_data:
writeConfig = json.load(json_data)
score_sparkml = score_sparkml.replace("{key}",writeConfig['Masterkey']).replace("{endpoint}",writeConfig['Endpoint']).replace("{database}",writeConfig['Database']).replace("{collection}",writeConfig['Collection']).replace("{DOCUMENTDB_DATABASE}",DOCUMENTDB_DATABASE).replace("{DOCUMENTDB_COLLECTION}", DOCUMENTDB_COLLECTION)
exec(score_sparkml)
with open("score_sparkml.py", "w") as file:
file.write(score_sparkml)
```
Next, create a environment config file with the dependencies needed:
```
%%writefile myenv_sparkml.yml
name: myenv
channels:
- defaults
dependencies:
- pip:
- numpy==1.14.2
- scikit-learn==0.19.1
- pandas
- azureml-core
- pydocumentdb
```
Register your model:
```
mymodel = Model.register(model_path = model_name, # this points to a local file
model_name = model_name, # this is the name the model is registered as, am using same name for both path and name.
description = "ADB trained model",
workspace = ws)
print(mymodel.name, mymodel.description, mymodel.version)
```
### 3.3 Deploy the model as a Service on AKS
#### 3.3.1 Create a container for your model service:
```
# Create Image for Web Service
models = [mymodel]
runtime = "spark-py"
conda_file = 'myenv_sparkml.yml'
driver_file = "score_sparkml.py"
# image creation
from azureml.core.image import ContainerImage
myimage_config = ContainerImage.image_configuration(execution_script = driver_file,
runtime = runtime,
conda_file = conda_file)
image = ContainerImage.create(name = container_image_name,
# this is the model object
models = [mymodel],
image_config = myimage_config,
workspace = ws)
# Wait for the create process to complete
image.wait_for_creation(show_output = True)
```
#### 3.3.2 Create an AKS Cluster to run your container (this may take 20-25 minutes):
```
from azureml.core.compute import AksCompute, ComputeTarget
# Use the default configuration (can also provide parameters to customize)
prov_config = AksCompute.provisioning_configuration()
# Create the cluster
aks_target = ComputeTarget.create(workspace = ws,
name = aks_name,
provisioning_configuration = prov_config)
aks_target.wait_for_completion(show_output = True)
print(aks_target.provisioning_state)
print(aks_target.provisioning_errors)
```
#### 3.3.3 Deploy the container image to AKS:
```
#Set the web service configuration (using default here with app insights)
aks_config = AksWebservice.deploy_configuration(enable_app_insights=True)
# Webservice creation using single command, there is a variant to use image directly as well.
try:
aks_service = Webservice.deploy_from_image(
workspace=ws,
name=service_name,
deployment_config = aks_config,
image = image,
deployment_target = aks_target
)
aks_service.wait_for_deployment(show_output=True)
except Exception:
aks_service = Webservice.list(ws)[0]
```
### 3.4 Call the AKS model service
After the deployment, the service can be called with a user ID – the service will then look up the top 10 recommendations for that user in Cosmos DB and send back the results.
The following script demonstrates how to call the recommendation service API and view the result for the given user ID:
```
scoring_url = aks_service.scoring_uri
service_key = aks_service.get_keys()[0]
input_data = '["{\\"id\\":\\"496\\"}"]'.encode()
req = urllib.request.Request(scoring_url,data=input_data)
req.add_header("Authorization","Bearer {}".format(service_key))
req.add_header("Content-Type","application/json")
tic = time.time()
with urllib.request.urlopen(req) as result:
res = result.readlines()
print(res)
toc = time.time()
t2 = toc - tic
print("Full run took %.2f seconds" % (toc - tic))
```
| github_jupyter |
# Gender Prediction using MobileNetV2
By Abhishek Chatterjee
(abhishekchatterjeejit@gmail.com)
**The aim of this project is to make a computer program to detect the gender of a person based on the single image of his/her face. This project is using the MobileNetV2 deep learning CNN architecture to predict it. The dataset that is used to train is a mix of the IMDB WIKI dataset and Selfie Dataset.**
## Dependencies
In the first step, we wil import the dependencies that we need for this project.
```
# Dependencies
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from keras.preprocessing.image import ImageDataGenerator
from keras.models import Sequential
from keras.layers import Dense, Flatten
from keras.applications import MobileNetV2
from keras import optimizers
from sklearn.model_selection import train_test_split
```
## Connecting Google Drive
As I'm running it on Google Colab, and my dataset is stored into Google Drive, so I need to connect Colab with Google Drive
```
from google.colab import drive
drive.mount('/content/drive')
```
## Unzipping the dataset
In Google Drive, the dataset is stored as a zip file. So before using it, I need to unzip it.
```
!unzip -qq 'drive/My Drive/dataset/imdb+wiki+selfie.zip' -d ./
```
There are three files in the dataset.
* images/ folder - This folder contains the original images
* gender.csv - This CSV file contains the meta information for the dataset (gender and image name)
* age.csv - This CSV file contains the meta information for the age dataset (not needed here)
## Reading the Dataset
In this step, I will read the dataset, stored in CSV format. To read the dataset, I will use the pandas read_csv method.
Note: The entire the dataset is already preprocessed and cleaned. Please check the preprocessing code.
```
gender_data = pd.read_csv('./gender.csv')
# Fir testing im just using 10% of the data
# gender_data = gender_data.sample(frac=0.1)
```
## Analysing the Dataset
In this step, I will perform some basic analysis on the data
```
# Priting the first 10 rows of the dataset
gender_data.head()
# Printing the last 10 rows of the dataset
gender_data.tail()
# Listing the column names
print(gender_data.columns)
```
The column gender contains the gender label as Male and Female. And the column path contains the unique id of the images.
```
# Number of records present on the data
gender_data.shape
```
## Preprocessing the Dataset
Here I will perform some basic analysis of the dataset
```
# The path columns contains int values. I need to change it to string
gender_data = gender_data.astype({'gender' : str, 'path' : str})
# Add the .jpg image extension after the id of the image
gender_data['path'] = gender_data['path'] + '.jpg'
# Check the data again
gender_data.head()
```
## Checking the class balance
In this step, I need to check the class balance of the dataset. If there is more samples for one class, then our model can be biased towards that class. To solve this problem, we need to use some type of sampling.
```
genders = []
for gender in gender_data['gender'].values:
if gender == 'Male':
genders.append(1)
else:
genders.append(0)
plt.hist(genders, 2)
plt.show()
```
Here there is almost equal number of samples for each class. So we dont need to do any sampling.
## Spliting the dataset
Here I will split the dataset into two parts, one for training and one for testing
```
train, test = train_test_split(gender_data, test_size=0.2, random_state=1969)
# Checking the data again
print(train.shape)
print(test.shape)
test.head()
train.tail()
```
## Generator Functions
The dataset in big, So we need to read the data in small batch. In Keras, ImageDataGenerator class provides a generator methods that we can use here.
```
# A generator object with some basic settings
generator = ImageDataGenerator(featurewise_center=False,
samplewise_center=False,
featurewise_std_normalization=False,
samplewise_std_normalization=False,
zca_whitening=False,
rotation_range=10,
zoom_range = 0.1,
width_shift_range=0.1,
height_shift_range=0.1,
horizontal_flip=True,
vertical_flip=False)
# Now I will read the dataset using the generator
train_gen = generator.flow_from_dataframe(train,
directory='images/',
x_col='path',
y_col='gender',
target_size=(224,224),
batch_size=64)
test_gen = generator.flow_from_dataframe(test,
directory='images/',
x_col='path',
y_col='gender',
target_size=(224,224),
batch_size=64)
```
## Model
Here I will make the MobileNetV2 model
```
model = Sequential()
# Im initializing the model with imagenet weighs
mobile = MobileNetV2(include_top=False,
weights="imagenet",
input_shape=(224,224,3),
pooling="max")
model.add(mobile)
model.add(Dense(units=2, activation="softmax"))
model.compile(loss='binary_crossentropy', optimizer=optimizers.RMSprop(lr=2e-5), metrics=['accuracy'])
model.summary()
```
## Training the Model
Here I will train the model with the dataset
```
STEP_SIZE_TRAIN=train_gen.n//train_gen.batch_size
STEP_SIZE_TEST=test_gen.n//test_gen.batch_size
history = model.fit_generator(train_gen,
steps_per_epoch=STEP_SIZE_TRAIN,
validation_data=test_gen,
validation_steps=STEP_SIZE_TEST,
epochs=3)
```
## Ploting Training Graphs
Plotting training graphs to analyse the performence
```
plt.plot(history.history['acc'])
plt.plot(history.history['val_acc'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
model.save('weights.h5')
```
| github_jupyter |
# Introdução
Este notebook faz parte do trabalho de conclusão do curso (TCC) apresentado para obtenção do título de especialista no curso de Ciência de Dados e Big Data na PUC / Minas Gerais.
## Objetivo
Este notebook é auxiliar ao **analysis.ipynb**, e tem como objetivo obter os arquivos:
>listings.csv
- Descrição: Imóveis listados em Toronto, Ontario, na plataforma Airbnb em 08 de Fevereiro de 2021. Indexados pelo ID do anúncio.
- Fonte: Dados obtidos do site Insideairbnb.com.
>reviews.csv
- Dados: Avaliações dos imóveis listados em Toronto, Ontario, na plataforma Airbnb em 08 de Fevereiro de 2021. Indexados pelo ID da avaliação.
- Fonte: Dados obtidos do site Insideairbnb.com.
## Fontes de Dados
Este notebook não precisa de nenhuma fonte de dados.
# 0 - Importando Bibliotecas
```
#!pip3 install wget
# Importando Bibliotecas
import gzip
import shutil
import wget
import os
from pathlib import Path
# Função para remover descompactar os arquivos em .gzip
def gunzip_shutil(source_filepath, dest_filepath, block_size=65536):
with gzip.open(source_filepath, 'rb') as s_file, \
open(dest_filepath, 'wb') as d_file:
shutil.copyfileobj(s_file, d_file, block_size)
```
# 1 - Download dos Arquivos de Interesse
```
dates = ["2021-02-08"]
files = ["listings.csv.gz", "reviews.csv.gz"]
remoteUrlbase = "http://data.insideairbnb.com/" + "canada/on/toronto/" # Ex: http://data.insideairbnb.com/canada/on/toronto/2021-01-02/data/listings.csv.gz
localUrlbase = "../2-data/input/Toronto_"
# Barra de Progresso (wget)
def bar_progress(current, total, width=80):
progress_message = "Downloading: %d%% [%d / %d] bytes" % (current / total * 100, current, total)
# Usar stdout.write() em vez de print()
sys.stdout.write("\r" + progress_message)
sys.stdout.flush()
for i in range(len(dates)):
for j in range(len(files)):
remoteUrl = remoteUrlbase + dates[i] + "/data/" + files[j]
localUrl = localUrlbase + dates[i] + "/"
localFileUrl = localUrl + files[j]
# Só vamos baixar SE os arquivos não existirem localmente
if not os.path.exists(localFileUrl):
try:
# Criando pastas se não existirem
Path(localUrl).mkdir(parents=True, exist_ok=True)
wget.download(remoteUrl, localUrl, bar=bar_progress)
except Exception as ex:
print("Was not able to download " + remoteUrl)
else:
print("Skipping download. File " + localFileUrl + " Already exists")
if not os.path.exists(localFileUrl[0:-3]):
gunzip_shutil(localFileUrl, localFileUrl[0:-3])
print ("\n Downloads concluidos")
```
| github_jupyter |
# VGP 245
## Read Write Files, Pickle, Json, Shelve, and SQLite3
To open a file use the method __open__
```
%%writefile mytext.txt
Today is a rainy day
maybe it's time to bring an umbrella
f = open('mytext.txt')
f.read()
f.read()
f.seek(0)
f.read()
f.close()
with open('mytext.txt') as f:
print(f.read())
f.write('?')
with open('mytext2.txt', mode='w') as f:
f.write('\n Nice T-shirt')
with open('mytext2.txt', mode='a') as f:
f.write('\n Cool another T-shirt')
class Character:
'''An RPG Character'''
def __init__(self, name, hp, mp):
self.name = name
self.hp = hp
self.mp = mp
def attack(self):
self.mp -=1
def hit(self):
self.hp -= 1
def __str__(self):
return f'{self.name} {self.hp} {self.mp}'
warrior1 = Character('Bobby', 100, 80)
warrior2 = Character('Juan', 100, 80)
warrior3 = Character('Gil', 100, 80)
warrior4 = Character('Matt', 100, 80)
warriors = [warrior1, warrior2, warrior3, warrior4]
print(str(warrior1))
#with open('warriors.txt', mode='w') as f:
# f.write(warriors)
with open('warriors.txt', mode='w') as f:
for warrior in warriors:
f.write(warrior)
with open('warriors.txt', mode='w') as f:
for warrior in warriors:
f.write(str(warrior))
import pickle
with open('warriorz.rando', mode='wb') as f:
pickle.dump(warriors, f)
with open('warriorz.rando', mode='rb') as f:
warriors2 = pickle.load(f)
for warrior in warriors2:
print(str(warrior))
import json
with open('warriors.json', mode='w') as f:
json.dump(warriors, f)
warrior1.__dict__
#json.dumps(warriors)
import json
with open('warriors.json', mode='w') as f:
json.dump(warriors.__dict__, f)
import json
warriors_dict_list = []
for warrior in warriors:
warriors_dict_list.append(warrior.__dict__)
with open('warriors.json', mode='w') as f:
json.dump(warriors_dict_list, f)
with open('warriors.json', mode='r') as f:
warriors3 = json.load(f)
print(warriors3)
import shelve
with shelve.open('shelvewarrior') as db:
db['warriors'] = warriors
shelved_warriors = []
with shelve.open('shelvewarrior', writeback=True) as db:
shelved_warriors = db['warriors']
db['warriors'].append(Character('Jimmy', 110, 10))
#db['warriors'] = shelved_warriors
for warrior in shelved_warriors:
print(str(warrior))
import sqlite3
db = sqlite3.connect('contacts.sqlite')
db.execute("CREATE TABLE IF NOT EXISTS Contacts ( ContactID INTEGER, FirstName TEXT, LastName TEXT)")
#db.execute("INSERT INTO Contacts ( ContactID, FirstName, LastName) VALUES (0, 'Lawrence', 'Cheung') ")
#db.execute("INSERT INTO Contacts ( ContactID, FirstName, LastName) VALUES (1, 'Bobby', 123) ")
#db.execute("INSERT INTO Contacts ( ContactID, FirstName, LastName) VALUES (2, 'Billy', 'M') ")
#db.execute("INSERT INTO Contacts ( ContactID, FirstName, LastName) VALUES (3, 'Jimmy', 'K') ")
#db.execute("INSERT INTO Contacts ( ContactID, FirstName, LastName) VALUES (4, 'Bimmy', 'J') ")
cursor = db.cursor()
for row in cursor.execute("SELECT * FROM Contacts"):
print(row)
#for ContactID, FirstName, LastName in cursor.execute("SELECT * FROM Contacts"):
# print(ContactID)
# print(FirstName)
# print(LastName)
for FirstName in cursor.execute("SELECT FirstName FROM Contacts"):
print(FirstName)
cursor.close()
db.commit()
db.close()
db = sqlite3.connect('contacts.sqlite')
cursor = db.cursor()
cursor.execute("SELECT * FROM Contacts")
#for row in cursor.execute("SELECT * FROM Contacts"):
# print(row)
print(cursor.fetchall())
# print(cursor.fetchone())
# print(cursor.fetchone())
cursor.close()
db.close()
```
| github_jupyter |
```
from __future__ import absolute_import, division, print_function, unicode_literals
import tensorflow as tf
import os
import time
from matplotlib import pyplot as plt
from IPython import display
FLAIRPATH = '/home/bdavid/Deep_Learning/playground/fake_flair_2d/png_cor/FLAIR/'
T1PATH ='/home/bdavid/Deep_Learning/playground/fake_flair_2d/png_cor/T1/'
BUFFER_SIZE = 400
BATCH_SIZE = 1
IMG_WIDTH = 256
IMG_HEIGHT = 256
def load(image_file):
input_image = tf.io.read_file(image_file)
input_image = tf.image.decode_png(input_image,channels=1)
input_image = tf.image.convert_image_dtype(input_image, tf.float32)
#image_file=image_file.replace(T1PATH,FLAIRPATH)
image_file=tf.strings.regex_replace(image_file,T1PATH,FLAIRPATH)
real_image = tf.io.read_file(image_file)
real_image = tf.image.decode_png(real_image,channels=1)
real_image = tf.image.convert_image_dtype(real_image, tf.float32)
return input_image, real_image
inp, re = load(T1PATH+'train/12038_slice080.png')#,load(FLAIRPATH+'train/12038_slice080.png')
# casting to int for matplotlib to show the image
plt.figure()
plt.imshow(inp[:,:,0]/255.0,cmap='gray')
plt.figure()
plt.imshow(re[:,:,0]/255.0,cmap='gray')
def resize(input_image, real_image, height, width):
input_image = tf.image.resize(input_image, [height, width],
method=tf.image.ResizeMethod.NEAREST_NEIGHBOR)
real_image = tf.image.resize(real_image, [height, width],
method=tf.image.ResizeMethod.NEAREST_NEIGHBOR)
return input_image, real_image
def random_crop(input_image, real_image):
stacked_image = tf.stack([input_image, real_image], axis=0)
cropped_image = tf.image.random_crop(
stacked_image, size=[2, IMG_HEIGHT, IMG_WIDTH, 1])
return cropped_image[0], cropped_image[1]
# normalizing the images to [-1, 1]
def normalize(input_image, real_image):
input_image = (input_image / 127.5) - 1
real_image = (real_image / 127.5) - 1
return input_image, real_image
@tf.function()
def random_jitter(input_image, real_image):
# resizing to 286 x 286 x 3
input_image, real_image = resize(input_image, real_image, 286, 286)
# randomly cropping to 256 x 256 x 3
input_image, real_image = random_crop(input_image, real_image)
if tf.random.uniform(()) > 0.5:
# random mirroring
input_image = tf.image.flip_left_right(input_image)
real_image = tf.image.flip_left_right(real_image)
return input_image, real_image
plt.figure(figsize=(6, 6))
for i in range(4):
rj_inp, rj_re = random_jitter(inp, re)
plt.subplot(2, 2, i+1)
plt.imshow(rj_inp[:,:,0]/255.0,cmap='gray')
plt.axis('off')
plt.show()
def load_image_train(image_file):
input_image, real_image = load(image_file)
input_image, real_image = random_jitter(input_image, real_image)
input_image, real_image = normalize(input_image, real_image)
return input_image, real_image
def load_image_test(image_file):
input_image, real_image = load(image_file)
input_image, real_image = resize(input_image, real_image,
IMG_HEIGHT, IMG_WIDTH)
input_image, real_image = normalize(input_image, real_image)
return input_image, real_image
test1,test2=load_image_train(T1PATH+'train/12038_slice080.png')
plt.imshow(test2[:,:,0]/255.0,cmap='gray')
train_dataset = tf.data.Dataset.list_files(T1PATH+'train/*.png')
train_dataset = train_dataset.map(load_image_train,
num_parallel_calls=tf.data.experimental.AUTOTUNE)
train_dataset = train_dataset.shuffle(BUFFER_SIZE)
train_dataset = train_dataset.batch(BATCH_SIZE)
test_dataset = tf.data.Dataset.list_files(T1PATH+'test/*.png')
test_dataset = test_dataset.map(load_image_test)
test_dataset = test_dataset.batch(BATCH_SIZE)
OUTPUT_CHANNELS = 1
def downsample(filters, size, apply_batchnorm=True):
initializer = tf.random_normal_initializer(0., 0.02)
result = tf.keras.Sequential()
result.add(
tf.keras.layers.Conv2D(filters, size, strides=2, padding='same',
kernel_initializer=initializer, use_bias=False))
if apply_batchnorm:
result.add(tf.keras.layers.BatchNormalization())
result.add(tf.keras.layers.LeakyReLU())
return result
down_model = downsample(3, 4)
down_result = down_model(tf.expand_dims(inp, 0))
print (down_result.shape)
def upsample(filters, size, apply_dropout=False):
initializer = tf.random_normal_initializer(0., 0.02)
result = tf.keras.Sequential()
result.add(
tf.keras.layers.Conv2DTranspose(filters, size, strides=2,
padding='same',
kernel_initializer=initializer,
use_bias=False))
result.add(tf.keras.layers.BatchNormalization())
if apply_dropout:
result.add(tf.keras.layers.Dropout(0.5))
result.add(tf.keras.layers.ReLU())
return result
up_model = upsample(3, 4)
up_result = up_model(down_result)
print (up_result.shape)
def Generator():
inputs = tf.keras.layers.Input(shape=[256,256,1])
down_stack = [
downsample(64, 4, apply_batchnorm=False), # (bs, 128, 128, 64)
downsample(128, 4), # (bs, 64, 64, 128)
downsample(256, 4), # (bs, 32, 32, 256)
downsample(512, 4), # (bs, 16, 16, 512)
downsample(512, 4), # (bs, 8, 8, 512)
downsample(512, 4), # (bs, 4, 4, 512)
downsample(512, 4), # (bs, 2, 2, 512)
downsample(512, 4), # (bs, 1, 1, 512)
]
up_stack = [
upsample(512, 4, apply_dropout=True), # (bs, 2, 2, 1024)
upsample(512, 4, apply_dropout=True), # (bs, 4, 4, 1024)
upsample(512, 4, apply_dropout=True), # (bs, 8, 8, 1024)
upsample(512, 4), # (bs, 16, 16, 1024)
upsample(256, 4), # (bs, 32, 32, 512)
upsample(128, 4), # (bs, 64, 64, 256)
upsample(64, 4), # (bs, 128, 128, 128)
]
initializer = tf.random_normal_initializer(0., 0.02)
last = tf.keras.layers.Conv2DTranspose(OUTPUT_CHANNELS, 4,
strides=2,
padding='same',
kernel_initializer=initializer,
activation='tanh') # (bs, 256, 256, 3)
x = inputs
# Downsampling through the model
skips = []
for down in down_stack:
x = down(x)
skips.append(x)
skips = reversed(skips[:-1])
# Upsampling and establishing the skip connections
for up, skip in zip(up_stack, skips):
x = up(x)
x = tf.keras.layers.Concatenate()([x, skip])
x = last(x)
return tf.keras.Model(inputs=inputs, outputs=x)
generator = Generator()
tf.keras.utils.plot_model(generator, show_shapes=True, dpi=64)
gen_output = generator(inp[tf.newaxis,...], training=False)
plt.imshow(gen_output[0,...][:,:,0],cmap='gray')
LAMBDA = 100
def generator_loss(disc_generated_output, gen_output, target):
gan_loss = loss_object(tf.ones_like(disc_generated_output), disc_generated_output)
# mean absolute error
#l1_loss = tf.reduce_mean(tf.abs(target - gen_output))
# mean squared error
l2_loss = tf.reduce_mean(tf.math.squared_difference(target,gen_output))
#total_gen_loss = gan_loss + (LAMBDA * l1_loss)
total_gen_loss = gan_loss + (LAMBDA * l2_loss)
return total_gen_loss, gan_loss, l2_loss
def Discriminator():
initializer = tf.random_normal_initializer(0., 0.02)
inp = tf.keras.layers.Input(shape=[256, 256, 1], name='input_image')
tar = tf.keras.layers.Input(shape=[256, 256, 1], name='target_image')
x = tf.keras.layers.concatenate([inp, tar]) # (bs, 256, 256, channels*2)
down1 = downsample(64, 4, False)(x) # (bs, 128, 128, 64)
down2 = downsample(128, 4)(down1) # (bs, 64, 64, 128)
down3 = downsample(256, 4)(down2) # (bs, 32, 32, 256)
zero_pad1 = tf.keras.layers.ZeroPadding2D()(down3) # (bs, 34, 34, 256)
conv = tf.keras.layers.Conv2D(512, 4, strides=1,
kernel_initializer=initializer,
use_bias=False)(zero_pad1) # (bs, 31, 31, 512)
batchnorm1 = tf.keras.layers.BatchNormalization()(conv)
leaky_relu = tf.keras.layers.LeakyReLU()(batchnorm1)
zero_pad2 = tf.keras.layers.ZeroPadding2D()(leaky_relu) # (bs, 33, 33, 512)
last = tf.keras.layers.Conv2D(1, 4, strides=1,
kernel_initializer=initializer)(zero_pad2) # (bs, 30, 30, 1)
return tf.keras.Model(inputs=[inp, tar], outputs=last)
discriminator = Discriminator()
tf.keras.utils.plot_model(discriminator, show_shapes=True, dpi=64)
disc_out = discriminator([inp[tf.newaxis,...], gen_output], training=False)
plt.imshow(disc_out[0,...,-1], cmap='RdBu_r')
plt.colorbar()
loss_object = tf.keras.losses.BinaryCrossentropy(from_logits=True)
def discriminator_loss(disc_real_output, disc_generated_output):
real_loss = loss_object(tf.ones_like(disc_real_output), disc_real_output)
generated_loss = loss_object(tf.zeros_like(disc_generated_output), disc_generated_output)
total_disc_loss = real_loss + generated_loss
return total_disc_loss
generator_optimizer = tf.keras.optimizers.Adam(2e-4, beta_1=0.5)
discriminator_optimizer = tf.keras.optimizers.Adam(2e-4, beta_1=0.5)
checkpoint_dir = './training_checkpoints'
checkpoint_prefix = os.path.join(checkpoint_dir, "ckpt")
checkpoint = tf.train.Checkpoint(generator_optimizer=generator_optimizer,
discriminator_optimizer=discriminator_optimizer,
generator=generator,
discriminator=discriminator)
def generate_images(model, test_input, tar):
prediction = model(test_input, training=True)
plt.figure(figsize=(15,15))
display_list = [test_input[0], tar[0], prediction[0]]
title = ['Input Image', 'Ground Truth', 'Predicted Image']
for i in range(3):
plt.subplot(1, 3, i+1)
plt.title(title[i])
# getting the pixel values between [0, 1] to plot it.
plt.imshow(display_list[i][:,:,0] * 0.5 + 0.5, cmap='gray')
plt.axis('off')
plt.show()
for example_input, example_target in test_dataset.take(1):
generate_images(generator, example_input, example_target)
EPOCHS = 150
import datetime
log_dir="logs/"
summary_writer = tf.summary.create_file_writer(
log_dir + "fit/" + datetime.datetime.now().strftime("%Y%m%d-%H%M%S"))
@tf.function
def train_step(input_image, target, epoch):
with tf.GradientTape() as gen_tape, tf.GradientTape() as disc_tape:
gen_output = generator(input_image, training=True)
disc_real_output = discriminator([input_image, target], training=True)
disc_generated_output = discriminator([input_image, gen_output], training=True)
#gen_total_loss, gen_gan_loss, gen_l1_loss = generator_loss(disc_generated_output, gen_output, target)
gen_total_loss, gen_gan_loss, gen_l2_loss = generator_loss(disc_generated_output, gen_output, target)
disc_loss = discriminator_loss(disc_real_output, disc_generated_output)
generator_gradients = gen_tape.gradient(gen_total_loss,
generator.trainable_variables)
discriminator_gradients = disc_tape.gradient(disc_loss,
discriminator.trainable_variables)
generator_optimizer.apply_gradients(zip(generator_gradients,
generator.trainable_variables))
discriminator_optimizer.apply_gradients(zip(discriminator_gradients,
discriminator.trainable_variables))
with summary_writer.as_default():
tf.summary.scalar('gen_total_loss', gen_total_loss, step=epoch)
tf.summary.scalar('gen_gan_loss', gen_gan_loss, step=epoch)
tf.summary.scalar('gen_l2_loss', gen_l2_loss, step=epoch)
tf.summary.scalar('disc_loss', disc_loss, step=epoch)
def fit(train_ds, epochs, test_ds):
for epoch in range(epochs):
start = time.time()
for example_input, example_target in test_ds.take(1):
generate_images(generator, example_input, example_target)
print("Epoch: ", epoch)
# Train
for n, (input_image, target) in train_ds.enumerate():
print('.', end='')
if (n+1) % 100 == 0:
display.clear_output(wait=True)
print()
for example_input, example_target in test_ds.take(1):
generate_images(generator, example_input, example_target)
train_step(input_image, target, epoch)
print()
# saving (checkpoint) the model every 5 epochs
if (epoch + 1) % 5 == 0:
checkpoint.save(file_prefix = checkpoint_prefix)
print ('Time taken for epoch {} is {} sec\n'.format(epoch + 1,
time.time()-start))
checkpoint.save(file_prefix = checkpoint_prefix)
%load_ext tensorboard
%tensorboard --logdir {log_dir}
fit(train_dataset, EPOCHS, test_dataset)
```
| github_jupyter |
# Building text classifier with Differential Privacy
In this tutorial we will train a text classifier with Differential Privacy by taking a model pre-trained on public text data and fine-tuning it for a different task.
When training a model with differential privacy, we almost always face a trade-off between model size and accuracy on the task. The exact details depend on the problem, but a rule of thumb is that the fewer parameters the model has, the easier it is to get a good performance with DP.
Most state-of-the-art NLP models are quite deep and large (e.g. [BERT-base](https://github.com/google-research/bert) has over 100M parameters), which makes task of training text model on a private datasets rather challenging.
One way of addressing this problem is to divide the training process into two stages. First, we will pre-train the model on a public dataset, exposing the model to generic text data. Assuming that the generic text data is public, we will not be using differential privacy at this step. Then, we freeze most of the layers, leaving only a few upper layers to be trained on the private dataset using DP-SGD. This way we can get the best of both worlds - we have a deep and powerful text understanding model, while only training a small number of parameters with differentially private algorithm.
In this tutorial we will take the pre-trained [BERT-base](https://github.com/google-research/bert) model and fine-tune it to recognize textual entailment on the [SNLI](https://nlp.stanford.edu/projects/snli/) dataset.
## Dataset
First, we need to download the dataset (we'll user Stanford NLP mirror)
```
STANFORD_SNLI_URL = "https://nlp.stanford.edu/projects/snli/snli_1.0.zip"
DATA_DIR = "data"
import zipfile
import urllib.request
import os
def download_and_extract(dataset_url, data_dir):
print("Downloading and extracting ...")
filename = "snli.zip"
urllib.request.urlretrieve(dataset_url, filename)
with zipfile.ZipFile(filename) as zip_ref:
zip_ref.extractall(data_dir)
os.remove(filename)
print("Completed!")
download_and_extract(STANFORD_SNLI_URL, DATA_DIR)
```
The dataset comes in two formats (`tsv` and `json`) and has already been split into train/dev/test. Let’s verify that’s the case.
```
snli_folder = os.path.join(DATA_DIR, "snli_1.0")
os.listdir(snli_folder)
```
Let's now take a look inside. [SNLI dataset](https://nlp.stanford.edu/projects/snli/) provides ample syntactic metadata, but we'll only use raw input text. Therefore, the only fields we're interested in are **sentence1** (premise), **sentence2** (hypothesis) and **gold_label** (label chosen by the majority of annotators).
Label defines the relation between premise and hypothesis: either *contradiction*, *neutral* or *entailment*.
```
import pandas as pd
train_path = os.path.join(snli_folder, "snli_1.0_train.txt")
dev_path = os.path.join(snli_folder, "snli_1.0_dev.txt")
df_train = pd.read_csv(train_path, sep='\t')
df_test = pd.read_csv(dev_path, sep='\t')
df_train[['sentence1', 'sentence2', 'gold_label']][:5]
```
## Model
BERT (Bidirectional Encoder Representations from Transformers) is state of the art approach to various NLP tasks. It uses a Transformer architecture and relies heavily on the concept of pre-training.
We'll use a pre-trained BERT-base model, provided in huggingface [transformers](https://github.com/huggingface/transformers) repo.
It gives us a pytorch implementation for the classic BERT architecture, as well as a tokenizer and weights pre-trained on a public English corpus (Wikipedia).
Please follow these [installation instrucitons](https://github.com/huggingface/transformers#installation) before proceeding.
```
from transformers import BertConfig, BertTokenizer, BertForSequenceClassification
model_name = "bert-base-cased"
config = BertConfig.from_pretrained(
model_name,
num_labels=3,
)
tokenizer = BertTokenizer.from_pretrained(
"bert-base-cased",
do_lower_case=False,
)
model = BertForSequenceClassification.from_pretrained(
"bert-base-cased",
config=config,
)
```
The model has the following structure. It uses a combination of word, positional and token *embeddings* to create a sequence representation, then passes the data through 12 *transformer encoders* and finally uses a *linear classifier* to produce the final label.
As the model is already pre-trained and we only plan to fine-tune few upper layers, we want to freeze all layers, except for the last encoder and above (`BertPooler` and `Classifier`).
```
from IPython.display import Image
Image(filename='img/BERT.png')
trainable_layers = [model.bert.encoder.layer[-1], model.bert.pooler, model.classifier]
total_params = 0
trainable_params = 0
for p in model.parameters():
p.requires_grad = False
total_params += p.numel()
for layer in trainable_layers:
for p in layer.parameters():
p.requires_grad = True
trainable_params += p.numel()
print(f"Total parameters count: {total_params}") # ~108M
print(f"Trainable parameters count: {trainable_params}") # ~7M
```
Thus, by using pre-trained model we reduce the number of trainable params from over 100 millions to just above 7.5 millions. This will help both performance and convergence with added noise.
## Prepare the data
Before we begin training, we need to preprocess the data and convert it to the format our model expects.
(Note: it'll take 5-10 minutes to run on a laptop)
```
LABEL_LIST = ['contradiction', 'entailment', 'neutral']
MAX_SEQ_LENGHT = 128
import torch
import transformers
from torch.utils.data import TensorDataset
from transformers.data.processors.utils import InputExample
from transformers.data.processors.glue import glue_convert_examples_to_features
def _create_examples(df, set_type):
""" Convert raw dataframe to a list of InputExample. Filter malformed examples
"""
examples = []
for index, row in df.iterrows():
if row['gold_label'] not in LABEL_LIST:
continue
if not isinstance(row['sentence1'], str) or not isinstance(row['sentence2'], str):
continue
guid = f"{index}-{set_type}"
examples.append(
InputExample(guid=guid, text_a=row['sentence1'], text_b=row['sentence2'], label=row['gold_label']))
return examples
def _df_to_features(df, set_type):
""" Pre-process text. This method will:
1) tokenize inputs
2) cut or pad each sequence to MAX_SEQ_LENGHT
3) convert tokens into ids
The output will contain:
`input_ids` - padded token ids sequence
`attention mask` - mask indicating padded tokens
`token_type_ids` - mask indicating the split between premise and hypothesis
`label` - label
"""
examples = _create_examples(df, set_type)
#backward compatibility with older transformers versions
legacy_kwards = {}
from packaging import version
if version.parse(transformers.__version__) < version.parse("2.9.0"):
legacy_kwards = {
"pad_on_left": False,
"pad_token": tokenizer.convert_tokens_to_ids([tokenizer.pad_token])[0],
"pad_token_segment_id": 0,
}
return glue_convert_examples_to_features(
examples=examples,
tokenizer=tokenizer,
label_list=LABEL_LIST,
max_length=MAX_SEQ_LENGHT,
output_mode="classification",
**legacy_kwards,
)
def _features_to_dataset(features):
""" Convert features from `_df_to_features` into a single dataset
"""
all_input_ids = torch.tensor([f.input_ids for f in features], dtype=torch.long)
all_attention_mask = torch.tensor(
[f.attention_mask for f in features], dtype=torch.long
)
all_token_type_ids = torch.tensor(
[f.token_type_ids for f in features], dtype=torch.long
)
all_labels = torch.tensor([f.label for f in features], dtype=torch.long)
dataset = TensorDataset(
all_input_ids, all_attention_mask, all_token_type_ids, all_labels
)
return dataset
train_features = _df_to_features(df_train, "train")
test_features = _df_to_features(df_test, "test")
train_dataset = _features_to_dataset(train_features)
test_dataset = _features_to_dataset(test_features)
```
## Choosing batch size
Let's talk about batch sizes for a bit.
In addition to all the considerations you normally take into account when choosing batch size, training model with DP adds another one - privacy cost.
Because of the threat model we assume and the way we add noise to the gradients, larger batch sizes (to a certain extent) generally help convergence. We add the same amount of noise to each gradient update (scaled to the norm of one sample in the batch) regardless of the batch size. What this means is that as the batch size increases, the relative amount of noise added decreases. while preserving the same epsilon guarantee.
You should, however, keep in mind that increasing batch size has its price in terms of epsilon, which grows at `O(sqrt(batch_size))` as we train (therefore larger batches make it grow faster). The good strategy here is to experiment with multiple combinations of `batch_size` and `noise_multiplier` to find the one that provides best possible quality at acceptable privacy guarantee.
There's another side to this - memory. Opacus computes and stores *per sample* gradients, so for every normal gradient, Opacus will store `n=batch_size` per-sample gradients on each step, thus increasing the memory footprint by at least `O(batch_size)`. In reality, however, the peak memory requirement is `O(batch_size^2)` compared to non-private model. This is because some intermediate steps in per sample gradient computation involve operations on two matrices, each with batch_size as one of the dimensions.
The good news is, we can pick the most appropriate batch size, regardless of memory constrains. Opacus has built-in support for *virtual* batches. Using it we can separate physical steps (gradient computation) and logical steps (noise addition and parameter updates): use larger batches for training, while keeping memory footprint low. Below we will specify two constants:
- `BATCH_SIZE` defines the maximum batch size we can afford from a memory standpoint, and only affects computation speed
- `VIRTUAL_BATCH_SIZE`, on the other hand, is equivalent to normal batch_size in the non-private setting, and will affect convergence and privacy guarantee.
```
BATCH_SIZE = 8
VIRTUAL_BATCH_SIZE = 32
from torch.utils.data import DataLoader, RandomSampler, SequentialSampler
train_sampler = RandomSampler(train_dataset)
train_dataloader = DataLoader(train_dataset, sampler=train_sampler, batch_size=BATCH_SIZE)
test_sampler = SequentialSampler(test_dataset)
test_dataloader = DataLoader(test_dataset, sampler=test_sampler, batch_size=BATCH_SIZE)
```
## Training
```
import torch
# Move the model to appropriate device
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
model = model.to(device)
# Set the model to train mode (HuggingFace models load in eval mode)
model = model.train()
# Define optimizer
optimizer = torch.optim.AdamW(model.parameters(), lr=5e-4, eps=1e-8)
```
Next we will define and attach PrivacyEngine. There are two parameters you need to consider here:
- `noise_multiplier`. It defines the trade-off between privacy and accuracy. Adding more noise will provide stronger privacy guarantees, but will also hurt model quality.
- `max_grad_norm`. Defines the maximum magnitude of L2 norms to which we clip per sample gradients. There is a bit of tug of war with this threshold: on the one hand, a low threshold means that we will clip many gradients, hurting convergence, so we might be tempted to raise it. However, recall that we add noise with `std=noise_multiplier * max_grad_norm` so we will pay for the increased threshold with more noise. In most cases you can rely on the model being quite resilient to clipping (after the first few iterations your model will tend to adjust so that its gradients stay below the clipping threshold), so you can often just keep the default value (`=1.0`) and focus on tuning `batch_size` and `noise_multiplier` instead. That being said, sometimes clipping hurts the model so it may be worth experimenting with different clipping thresholds, like we are doing in this tutorial.
These two parameters define the scale of the noise we add to gradients: the noise will be sampled from a Gaussian distribution with `std=noise_multiplier * max_grad_norm`.
```
from opacus import PrivacyEngine
ALPHAS = [1 + x / 10.0 for x in range(1, 100)] + list(range(12, 64))
NOISE_MULTIPLIER = 0.4
MAX_GRAD_NORM = 0.1
privacy_engine = PrivacyEngine(
module=model,
batch_size=VIRTUAL_BATCH_SIZE,
sample_size=len(train_dataset),
alphas=ALPHAS,
noise_multiplier=NOISE_MULTIPLIER,
max_grad_norm=MAX_GRAD_NORM,
)
privacy_engine.attach(optimizer)
```
Let’s first define the evaluation cycle.
```
import numpy as np
from tqdm.notebook import tqdm
def accuracy(preds, labels):
return (preds == labels).mean()
# define evaluation cycle
def evaluate(model):
model.eval()
loss_arr = []
accuracy_arr = []
for batch in test_dataloader:
batch = tuple(t.to(device) for t in batch)
with torch.no_grad():
inputs = {'input_ids': batch[0],
'attention_mask': batch[1],
'token_type_ids': batch[2],
'labels': batch[3]}
outputs = model(**inputs)
loss, logits = outputs[:2]
preds = np.argmax(logits.detach().cpu().numpy(), axis=1)
labels = inputs['labels'].detach().cpu().numpy()
loss_arr.append(loss.item())
accuracy_arr.append(accuracy(preds, labels))
model.train()
return np.mean(loss_arr), np.mean(accuracy_arr)
```
Now we specify the training parameters and run the training loop for three epochs
```
EPOCHS = 3
LOGGING_INTERVAL = 1000 # once every how many steps we run evaluation cycle and report metrics
DELTA = 1 / len(train_dataloader) # Parameter for privacy accounting. Probability of not uploding privacy guarantees
assert VIRTUAL_BATCH_SIZE % BATCH_SIZE == 0 # VIRTUAL_BATCH_SIZE should be divisible by BATCH_SIZE
virtual_batch_rate = VIRTUAL_BATCH_SIZE / BATCH_SIZE
for epoch in range(1, EPOCHS+1):
losses = []
for step, batch in enumerate(tqdm(train_dataloader)):
batch = tuple(t.to(device) for t in batch)
inputs = {'input_ids': batch[0],
'attention_mask': batch[1],
'token_type_ids': batch[2],
'labels': batch[3]}
outputs = model(**inputs) # output = loss, logits, hidden_states, attentions
loss = outputs[0]
loss.backward()
losses.append(loss.item())
# We process small batches of size BATCH_SIZE,
# until they're accumulated to a batch of size VIRTUAL_BATCH_SIZE.
# Only then we make a real `.step()` and update model weights
if (step + 1) % virtual_batch_rate == 0 or step == len(train_dataloader) - 1:
optimizer.step()
else:
optimizer.virtual_step()
if step > 0 and step % LOGGING_INTERVAL == 0:
train_loss = np.mean(losses)
eps, alpha = optimizer.privacy_engine.get_privacy_spent(DELTA)
eval_loss, eval_accuracy = evaluate(model)
print(
f"Epoch: {epoch} | "
f"Step: {step} | "
f"Train loss: {train_loss:.3f} | "
f"Eval loss: {eval_loss:.3f} | "
f"Eval accuracy: {eval_accuracy:.3f} | "
f"ɛ: {eps:.2f} (α: {alpha})"
)
```
For the test accuracy, after training for three epochs you should expect something close to the results below.
You can see that we can achieve quite strong privacy guarantee at epsilon=7.5 with a moderate accuracy cost of 11 percentage points compared to non-private model trained in a similar setting (upper layers only) and 16 points compared to best results we were able to achieve using the same architecture.
*NB: When not specified, DP-SGD is trained with upper layers only*
| Model | Noise multiplier | Batch size | Accuracy | Epsilon |
| --- | --- | --- | --- | --- |
| no DP, train full model | N/A | 32 | 90.1% | N/A |
| no DP, train upper layers only | N/A | 32 | 85.4% | N/A |
| DP-SGD | 1.0 | 32 | 70.5% | 0.7 |
| **DP-SGD (this tutorial)** | **0.4** | **32** | **74.3%** | **7.5** |
| DP-SGD | 0.3 | 32 | 75.8% | 20.7 |
| DP-SGD | 0.1 | 32 | 78.3% | 2865 |
| DP-SGD | 0.4 | 8 | 67.3% | 5.9 |
| github_jupyter |
# Statistical Analysis of Data
## Environment Settings
An statistical Analysis of the data captured will be performed.
The environment configuration is the following:
- A rectangle area is used whose dimension is 2 x 1.5 meters.
- A custom robot similar to an epuck was used.
- The robot starts in the middle of the arena.
- The robot moves in a random fashion way around the environment avoiding obstacles.
- The robot has 8 sensors that measure the distance between the robot and the walls.
- Some noise was introduced in the sensors measurements of the robot using the concept of [lookup tables](https://cyberbotics.com/doc/reference/distancesensor) in the Webots simulator which according to Webots documentation "The first column of the table specifies the input distances, the second column specifies the corresponding desired response values, and the third column indicates the desired standard deviation of the noise. The noise on the return value is computed according to a gaussian random number distribution whose range is calculated as a percent of the response value (two times the standard deviation is often referred to as the signal quality)". The following values were taken:
-First experiment:
- (0, 0, 0.01)
- (10, 10, 0.01)
-Second experiment:
- (0, 0, 0.2)
- (10, 10, 0.2)
- The simulator runs during 10 minutes in fast mode which is translated into 12 hours of collected data.
```
# Install a pip package in the current Jupyter kernel
import sys
!{sys.executable} -m pip install scikit-learn
!{sys.executable} -m pip install keras
import pandas as pd
import tensorflow as tf
import numpy as np
import math
from sklearn.ensemble import RandomForestRegressor
from keras import models
from keras import layers
from keras import regularizers
import matplotlib.pyplot as plt
from keras import optimizers
```
# First Experiment
```
csv_file = 'robot_info_dataset-jumped.csv'
df = pd.read_csv(csv_file)
df.head()
```
## Data pre-processing
The data collected 1384848 samples.
```
df.shape
```
The data set contains some null values so they should be deleted from the samples.
```
df = df.dropna()
```
Now the data will be normalized.
```
normalized_df=(df-df.min())/(df.max()-df.min())
normalized_df.describe()
```
## Input and output variables
The data will be split into training, testing and validation sets. 60% of the data will be used for training, 20% for training and 20% of validation.
```
# train size
test_size_percentage = .2
train_size_percentage = .6
ds_size = normalized_df.shape[0]
train_size = int(train_size_percentage * ds_size)
test_size = int(test_size_percentage * ds_size)
# shuffle dataset
normalized_df = normalized_df.sample(frac=1)
# separate inputs from outputs
inputs = normalized_df[['x', 'y', 'theta']]
targets = normalized_df[['sensor_1', 'sensor_2', 'sensor_3', 'sensor_4', 'sensor_5', 'sensor_6', 'sensor_7', 'sensor_8']]
# train
train_inputs = inputs[:train_size]
train_targets = targets[:train_size]
# test
test_inputs = inputs[train_size:(train_size + test_size)]
test_targets = targets[train_size:(train_size + test_size)]
# validation
validation_inputs = inputs[(train_size + test_size):]
validation_targets = targets[(train_size + test_size):]
```
## Neural Network
As input the neural network receives the x, y coordinates and rotation angle $\theta$. The output are the sensor measurements. One model per sensor will be created.
```
def get_model():
# neural network with a 10-neuron hidden layer
model = models.Sequential()
model.add(layers.Dense(10, activation='relu', input_shape=(3,)))
# model.add(layers.Dropout(0.5))
model.add(layers.Dense(6, activation='relu'))
model.add(layers.Dense(3, activation='relu'))
model.add(layers.Dense(1))
# rmsprop = optimizers.RMSprop(learning_rate=0.01)
model.compile(optimizer='rmsprop', loss='mse', metrics=['mae'])
return model
model = get_model()
history = model.fit(inputs, targets[['sensor_4']], epochs=75, batch_size=1, verbose=1)
history.history['mae']
model.save("nn_sensor_4.h5")
```
| github_jupyter |
```
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
import seaborn as sns
import warnings
warnings.filterwarnings("ignore")
data=pd.read_csv("Red Wine Quality.csv")
data.head(10)
data.shape
data.info()
```
# Total Unique Values:
```
data.nunique()
```
# Missing values
```
data.isnull().sum()
summary=data.describe()
summary
```
# Exploratory Data Analysis:
# Univariate Analysis:
```
plt.figure(figsize=(10,8))
sns.countplot(data['quality'])
fig, ax = plt.subplots(ncols = 4, nrows = 3, figsize = (15,10))
index = 0
ax = ax.flatten()
for col, value in data.items():
if col != 'type':
sns.boxplot(y=col, data = data, ax = ax[index])
index += 1
plt.tight_layout(pad = 0.5, w_pad = 0.7, h_pad = 5.0)
fig, ax = plt.subplots(ncols = 3, nrows = 4, figsize = (20,25))
index = 0
ax = ax.flatten()
for col, value in data.items():
if col != 'type':
sns.distplot(value, ax = ax[index])
index += 1
plt.tight_layout(pad = 0.5, w_pad = 0.7, h_pad = 5.0)
data.hist(bins = 45, figsize = (15,10))
plt.show()
```
# Bivariate Analysis:
```
for i in data.drop(columns=['quality']).columns:
plt.figure(figsize=[10,5],dpi=85)
data.groupby('quality')[i].mean().plot(kind='barh',color='orange')
plt.xlabel(i)
```
# Multivariate Analysis:
```
plt.figure(figsize=(10,8))
sns.scatterplot(x='alcohol',y='sulphates',hue='quality',data=data)
```
# Correlation:
```
cor=data.corr()
plt.figure(figsize=[10,5],dpi=100)
sns.heatmap(cor,xticklabels=cor.columns,yticklabels=cor.columns,annot=True)
sns.histplot(data)
data['quality'].value_counts()
```
# Consider 3,4,5 as bad and 6,7,8 as good
```
data['quality'] = data['quality'].map({3 : 'bad', 4 :'bad', 5: 'bad',6: 'good', 7: 'good', 8: 'good'})
data['quality'].value_counts()
```
# Convert good bad as binary variables
```
from sklearn.preprocessing import LabelEncoder
le = LabelEncoder()
data['quality'] = le.fit_transform(data['quality'])
data['quality'].value_counts()
```
# Define dependent(y) and independent variable(x)
```
X = data.drop('quality', axis = 1)
y = data['quality']
```
# 1)Logistic Regression
```
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(X, y, test_size = 0.3, random_state = 42)
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import classification_report, confusion_matrix, accuracy_score
from sklearn.model_selection import GridSearchCV, cross_val_score
lr = LogisticRegression()
lr.fit(x_train, y_train)
y_pred_train = lr.predict(x_train)
y_pred_test = lr.predict(x_test)
print("Training accuracy :", accuracy_score(y_pred_train, y_train))
print("Testing accuracy :", accuracy_score(y_pred_test, y_test))
```
* here we see that the accuracy is good, but we want more accurate model.
# Confusion Matrix
```
cm_test = confusion_matrix(y_test, y_pred_test)
print(cm_test)
print('Classification report for train data is : \n',
classification_report(y_train, y_pred_train))
print('\n')
print('Classification report for test data is : \n',
classification_report(y_test, y_pred_test))
```
# Decision Tree Classifier
```
from sklearn.tree import DecisionTreeClassifier
dtc = DecisionTreeClassifier()
dtc.fit(x_train, y_train)
y_pred_dtc_train = dtc.predict(x_train)
y_pred_dtc_test = dtc.predict(x_test)
print("Training accuracy :", accuracy_score(y_pred_dtc_train, y_train))
print("Testing accuracy :", accuracy_score(y_pred_dtc_test, y_test))
```
* here also the model is not good fitted
# Random Forest Classifier
```
from sklearn.ensemble import RandomForestClassifier
rfc = RandomForestClassifier()
rfc.fit(x_train, y_train)
y_pred_rfc_train = rfc.predict(x_train)
y_pred_rfc_test = rfc.predict(x_test)
print("Training accuracy :", accuracy_score(y_pred_rfc_train, y_train))
print("Testing accuracy :", accuracy_score(y_pred_rfc_test, y_test))
```
*Here also we can see that the training accuracy is 1(i.e 100%) hence it is over fitted
# Support vector classifier(svc)
```
from sklearn.svm import SVC
svm = SVC()
svm.fit(x_train, y_train)
y_pred_svm_train = svm.predict(x_train)
y_pred_svm_test = svm.predict(x_test)
print("Training accuracy :", accuracy_score(y_pred_svm_train, y_train))
print("Testing accuracy :", accuracy_score(y_pred_svm_test, y_test))
```
* here the accuracy is 0.6 model is good fitted but we want more accurate model
# KNeighborsClassifier(KNN)
```
from sklearn.neighbors import KNeighborsClassifier
for i in range(1,5):
neigh = KNeighborsClassifier(n_neighbors = i)
neigh.fit(x_train, y_train)
y_pred_knn_train = neigh.predict(x_train)
y_pred_knn_test = neigh.predict(x_test)
print("Train Accuracy : ", accuracy_score(y_pred_knn_train, y_train))
print("Test Accuracy : ", accuracy_score(y_pred_knn_test, y_test))
```
# hyper parameter tuning
```
model = SVC()
param = {'C': [0.01, 0.001, 0.0001, 0.1, 0.8, 0.9, 1 ,1.1 ,1.2 ,1.3 ,1.4],
'kernel':['linear', 'rbf'],
'gamma' :[0.1,0.8,0.9,1,1.1,1.2,1.3,1.4]
}
grid_svc = GridSearchCV(model,
param_grid = param,
scoring = 'accuracy',
cv = 5)
grid_svc.fit(x_train, y_train)
print("Best parameters for the model :", grid_svc.best_params_)
print("Best score for the model :", grid_svc.best_score_)
svc_predict = grid_svc.predict(x_test)
print('Accuracy Score: ', accuracy_score(y_test, svc_predict))
params = {
'n_estimators':[100, 200, 300, 400, 500],
'max_depth': [30, 40, 50],
'min_samples_leaf': [1, 2, 3],
'criterion': ["gini", "entropy"]
}
from sklearn.ensemble import RandomForestClassifier
from sklearn import tree
model = RandomForestClassifier(random_state = 42)
grid_search = GridSearchCV(estimator = model,
param_grid = params,
cv = 5, n_jobs = -1, verbose = 1, scoring = "accuracy")
grid_search.fit(x_train,y_train)
print("Best parameters for the model :", grid_search.best_params_)
print("Best score for the model :", grid_search.best_score_)
train_rfc_pred = grid_search.predict(x_train)
test_rfc_pred = grid_search.predict(x_test)
print("Train Accuracy : ",accuracy_score(y_train, train_rfc_pred))
print("Test Accuracy : ",accuracy_score(y_test, test_rfc_pred))
print('Classification report for train data is : \n',
classification_report(y_train, train_rfc_pred))
print('\n')
print('Classification report for test data is : \n',
classification_report(y_test, test_rfc_pred))
!pip install scikit-learn
from sklearn.metrics import roc_auc_score
roc_auc = roc_auc_score(y_test, grid_search.predict_proba(x_test)[:,1])
print('Aera under ROC cure :',roc_auc)
from sklearn.metrics import roc_curve
FPR_rfc_train, TPR_rfc_train, Thresholds_train = roc_curve(y_train, grid_search.predict_proba(x_train)[:,1])
FPR_rfc_test, TPR_rfc_test, Thresholds_test = roc_curve(y_test, grid_search.predict_proba(x_test)[:,1])
from sklearn.metrics import roc_curve
sns.set_style("whitegrid")
plt.figure(figsize = (20,8))
plt.plot(FPR_rfc_train, TPR_rfc_train, label = 'Train AUR Curve')
plt.plot(FPR_rfc_test, TPR_rfc_test, label = 'Test AUR Curve')
plt.plot([0,1], [0,1], label = 'Base Rate')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate', fontsize = 20)
plt.ylabel('True Positive Rate', fontsize = 20)
plt.title('ROC Graph', fontsize = 20)
plt.legend(loc = "lower right")
plt.show()
importance = grid_search.best_estimator_.feature_importances_
feature_imp = pd.Series(importance, index = data.columns[:11]).sort_values(ascending = False)
feature_imp
plt.figure(figsize = (15, 8))
# Visualize the Importance Creating a bar plot
sns.barplot(x = feature_imp, y = feature_imp.index)
# Add labels to your graph
plt.xlabel('Feature Importance Score')
plt.ylabel('Features')
plt.title("Visualizing Important Features")
plt.show()
```
# Deployment
```
import pickle
pickle.dump(grid_search, open('wine_grid_search.pkl','wb'))
model = pickle.load(open('wine_grid_search.pkl','rb'))
```
| github_jupyter |
# `RELIANCE - NSE Stock Data`
The file contains RELIANCE - NSE Stock Data from 1-Jan-16 to 6-May-21
The data can be used to forecast the stock prices of the future
Its a timeseries data from the national stock exchange of India
|| `Variable` | `Significance` |
| ------------- |:-------------:|:-------------:|
|1.|Symbol|Symbol of the listed stock on NSE|
|2.|Series|To which series does the stock belong (Equity, Options Future)|
|3.|Date|Date of the trade|
|4.|Prev Close|Previous day closing value of the stock|
|5.|Open Price|Current Day opening price of the stock|
|6.|High Price|Highest price touched by the stock in current day `(Target Variable)`|
|7.|Low Price|lowest price touched by the stock in current day|
|8.|Last Price|The price at which last trade occured in current day|
|9.|Close Price|Current day closing price of the stock|
|10.|Average Price|Average price of the day|
|11.|Total Traded Quantity|Total number of stocks traded in current day|
|12.|Turnover||
|13.|No. of Trades|Current day's total number of trades|
|14.|Deliverabel Quantity|Current day deliveable quantity to the traders|
|15.|% Dly Qt to Traded Qty|`(Deliverable Quantity/Total Traded Quantity)*100`|
```
import pandas as pd
data_path="./data/RILO - Copy.csv"
data=pd.read_csv(data_path)
data
# Renaming the columns to have snake_case naming style. (Just as a convention and for convenience)
data.columns=["_".join(column.lower().split()) for column in data.columns]
data.columns
# Using `.describe()` on an entire DataFrame we can get a summary of the distribution of continuous variables:
data.describe()
# Checking for null values
data.isnull().sum()
```
### As shown above, we do not have any null values in our dataset. Now we can focus on feature selection and model building.
```
# By using the correlation method `.corr()` we can get the relationship between each continuous variable:
correlation=data.corr()
correlation
```
### `Matplotlib`
Matplotlib is a visualization library in Python for 2D plots of arrays. Matplotlib is a multi-platform data visualization library built on NumPy arrays and designed to work with the broader SciPy stack.
One of the greatest benefits of visualization is that it allows us visual access to huge amounts of data in easily digestible visuals. Matplotlib consists of several plots like line, bar, scatter, histogram etc.
Matplotlib comes with a wide variety of plots. Plots helps to understand trends, patterns, and to make correlations. They’re typically instruments for reasoning about quantitative information
### `Seaborn`
Seaborn is a Python data visualization library based on matplotlib. It provides a high-level interface for drawing attractive and informative statistical graphics.
```
# Using seaborn and matplotlib to have a better visualization of correlation
import seaborn as sn
import matplotlib.pyplot as plt
plt.figure(figsize=(10,8))
sn.heatmap(correlation,annot=True,linewidth=1,cmap='PuOr')
plt.show()
```
From the above correlation matrix, we get a general idea of which variables can be treated as features to build our model. Lets list them out
Considering all the variables having `|corr|>=0.5`
- prev_close
- no._of_trades
- open_price
- low_price
- last_price
- turnover
- close_price
- %_dly_qt_to_traded_qty
- average_price
Now that we have a rough idea about our features, lets confirm their behaviour aginst target variable using scatter plots.
```
plt.figure(figsize=(18,18))
plt.subplot(3,3,1)
plt.scatter(data.prev_close,data.high_price)
plt.title('Relation with Previous Closing Price')
plt.subplot(3,3,2)
plt.scatter(data['no._of_trades'],data.high_price)
plt.title('Relation with No. of trades')
plt.subplot(3,3,3)
plt.scatter(data.open_price,data.high_price)
plt.title('Relation with Opening Price')
plt.subplot(3,3,4)
plt.scatter(data.low_price,data.high_price)
plt.title('Relation with Low Price')
plt.subplot(3,3,5)
plt.scatter(data.last_price,data.high_price)
plt.title('Relation with Last Price')
plt.subplot(3,3,6)
plt.scatter(data.turnover,data.high_price)
plt.title('Relation with Turnover')
plt.subplot(3,3,7)
plt.scatter(data.close_price,data.high_price)
plt.title('Relation with Closing Price')
plt.subplot(3,3,8)
plt.scatter(data['%_dly_qt_to_traded_qty'],data.high_price)
plt.title('Relation with Deliverable quantity')
plt.subplot(3,3,9)
plt.scatter(data.average_price,data.high_price)
plt.title('Relation with Average Price')
plt.show
```
From above visualization, we are clear to choose features for the linear-regression model. Those are:
- prev_close
- ~~no._of_trades~~
- open_price
- low_price
- last_price
- ~~turnover~~
- close_price
- ~~%_dly_qt_to_traded_qty~~
- average_price
```
features=['prev_close','open_price','low_price','last_price','close_price','average_price']
X=data[features]
X
# Target variable
y=data.high_price
y
# split data into training and validation data, for both features and target
# The split is based on a random number generator. Supplying a numeric value to
# the random_state argument guarantees we get the same split every time we
# run this script.
from sklearn.model_selection import train_test_split
train_X,val_X,train_y,val_y=train_test_split(X,y,test_size=0.2,random_state=0)
from sklearn.linear_model import LinearRegression
# Define model
model=LinearRegression()
# Fit model
model.fit(train_X,train_y)
# We use .score method to get an idea of quality of our model
model.score(val_X,val_y)
```
### Model Validation
There are many metrics for summarizing model quality, but we'll start with one called `Mean Absolute Error (also called MAE)`. Let's break down this metric starting with the last word, error.
`error=actual-predicted`
So, if a stock cost Rs.4000 at a timeframe, and we predicted it would cost Rs.3980 the error is Rs.20.
With the MAE metric, we take the absolute value of each error. This converts each error to a positive number. We then take the average of those absolute errors. This is our measure of model quality. In plain English, it can be said as
> On average, our predictions are off by about X.
```
from sklearn.metrics import mean_absolute_error
# Get predicted prices of stock on validation data
pred_y=model.predict(val_X)
mean_absolute_error(val_y,pred_y)
```
| github_jupyter |
```
# Authors: Naveen Lalwani, Rangeesh Muthaiyan
# Script to prune the trained Deep Neural Networks
import tensorflow as tf
from keras.layers import Dense, Conv2D, MaxPool2D, Flatten
from keras.models import Sequential
from keras import layers
from keras import callbacks
from tensorflow.keras.datasets import cifar10
# External Library used for Pruning
from kerassurgeon import identify
from kerassurgeon.operations import delete_channels
from keras.utils import np_utils
import time
training_verbosity = 2
# Download data if needed and import.
(x_train, y_train), (x_test, y_test) = tf.keras.datasets.cifar10.load_data()
y_train = np_utils.to_categorical(y_train)
y_test = np_utils.to_categorical(y_test)
dataset_train = tf.data.Dataset.from_tensor_slices((
tf.cast(x_train/255, tf.float32),
tf.cast(y_train, tf.int64)
)).shuffle(1000).batch(256)
dataset_test = tf.data.Dataset.from_tensor_slices((
tf.cast(x_test/255, tf.float32),
tf.cast(y_test, tf.int64)
)).batch(256)
# Create LeNet-5 model
model = Sequential()
model.add(Conv2D(6,
[5, 5],
input_shape=[32, 32, 3],
activation='relu',
name='conv_1'))
model.add(MaxPool2D())
model.add(Conv2D(16, [5, 5], activation='relu', name='conv_2'))
model.add(MaxPool2D())
#model.add(layers.Permute((2, 1, 3)))
model.add(Flatten())
model.add(Dense(120, activation='relu', name='dense_1'))
model.add(Dense(84, activation='relu', name='dense_2'))
model.add(Dense(10, activation='softmax', name='dense_3'))
model.compile(optimizer='adam',
loss='categorical_crossentropy',
metrics=['accuracy'])
early_stopping = callbacks.EarlyStopping(monitor='val_loss',
min_delta=0,
patience=10,
verbose=training_verbosity,
mode='auto')
reduce_lr = callbacks.ReduceLROnPlateau(monitor='val_loss',
factor=0.1,
patience=5,
verbose=training_verbosity,
mode='auto',
epsilon=0.001,
cooldown=0,
min_lr=0)
# Train LeNet on MNIST
results = model.fit(x_train,
y_train,
epochs=200,
batch_size=256,
validation_data = (x_test, y_test),
callbacks=[early_stopping, reduce_lr]
)
model.save("./LeNet5_cifar10_results/before_pruning.h5")
start = time.clock()
_, accuracy = model.evaluate(x_test, y_test, batch_size=256, verbose=2)
end = time.clock()
inferTime = end - start
print(f"Test Accuracy = {accuracy:.4f}, Inference Time = {inferTime:.2f}s")
model.save("./LeNet5_cifar10_results/before_pruning.h5")
layer_name = 'dense_1'
while accuracy > 0.50:
layer = model.get_layer(name=layer_name)
apoz = identify.get_apoz(model, layer, x_test)
high_apoz_channels = identify.high_apoz(apoz)
model = delete_channels(model, layer, high_apoz_channels)
model.compile(optimizer='adam',
loss='categorical_crossentropy',
metrics=['accuracy'])
loss,accuracy = model.evaluate(x_test,
y_test,
batch_size=256)
model.save("./LeNet5_cifar10_results/after_pruning.h5")
print('model accuracy after pruning: ', accuracy, '\n')
results = model.fit(x_train,
y_train,
epochs=100,
batch_size=256,
callbacks=[early_stopping, reduce_lr],
validation_data = (x_test, y_test))
loss,accuracy = model.evaluate(x_test,
y_test,
batch_size=256,
verbose=2)
model.save("./LeNet5_cifar10_results/after_pruning_retrain.h5")
print('model accuracy after retraining: ', accuracy, '\n')
start = time.clock()
_, accuracy = model.evaluate(x_test, y_test, batch_size=256, verbose=2)
end = time.clock()
inferTime = end - start
print(f"Test Accuracy: {accuracy:.4f}, Inference Time: {inferTime:.2f}s")
model.save("./LeNet5_cifar10_results/after_pruning_retrain.h5")
layer_name = 'dense_2'
while accuracy > 0.49:
layer = model.get_layer(name=layer_name)
apoz = identify.get_apoz(model, layer, x_test)
high_apoz_channels = identify.high_apoz(apoz)
model = delete_channels(model, layer, high_apoz_channels)
model.compile(optimizer='adam',
loss='categorical_crossentropy',
metrics=['accuracy'])
loss,accuracy = model.evaluate(x_test,
y_test,
batch_size=256)
model.save("./LeNet5_cifar10_results/after_pruning_dense2.h5")
print('model accuracy after pruning: ', accuracy, '\n')
results = model.fit(x_train,
y_train,
epochs=100,
batch_size=256,
callbacks=[early_stopping, reduce_lr],
validation_data = (x_test, y_test))
loss,accuracy = model.evaluate(x_test,
y_test,
batch_size=256,
verbose=2)
model.save("./LeNet5_cifar10_results/after_pruning_retrain_dense2.h5")
print('model accuracy after retraining: ', accuracy, '\n')
t = time.clock()
l,accuracy = model.evaluate(x_test, y_test, batch_size=256, verbose=2)
t = time.clock() - t
print(f"Test Accuracy: {accuracy:.4f}, Timing: {t:.2f}s")
model.save("./LeNet5_cifar10_results/after_pruning_retrain_dense2.h5")
layer_name = 'conv_2'
while accuracy > 0.49:
layer = model.get_layer(name=layer_name)
apoz = identify.get_apoz(model, layer, x_test)
high_apoz_channels = identify.high_apoz(apoz)
model = delete_channels(model, layer, high_apoz_channels)
model.compile(optimizer='adam',
loss='categorical_crossentropy',
metrics=['accuracy'])
loss,accuracy = model.evaluate(x_test,
y_test,
batch_size=256)
model.save("./LeNet5_cifar10_results/after_pruning_conv2.h5")
print('model accuracy after pruning: ', accuracy, '\n')
results = model.fit(x_train,
y_train,
epochs=100,
batch_size=256,
callbacks=[early_stopping, reduce_lr],
validation_data = (x_test, y_test))
loss,accuracy = model.evaluate(x_test,
y_test,
batch_size=256,
verbose=2)
model.save("./LeNet5_cifar10_results/after_pruning_retrain_conv2.h5")
print('model accuracy after retraining: ', accuracy, '\n')
t = time.clock()
l,accuracy = model.evaluate(x_test, y_test, batch_size=256, verbose=2)
t = time.clock() - t
print(f"Test Accuracy: {accuracy:.4f}, Timing: {t:.2f}s")
model.save("./LeNet5_cifar10_results/after_pruning_retrain_conv2.h5")
```
| github_jupyter |
# 关联分析
关联分析:从大数据集中寻找物品间的隐含关系
频繁项集:经常出现在一起的物品集合
关联规则:暗示两种物品之间可能存在很强的关系
频繁的衡量尺度:
* 支持度:数据集中包含该项集记录所占比例
* 置信度:如{尿布}->{葡萄酒}的置信度为 `r = 支持度({尿布,葡萄酒}) / 支持度({尿布})`,
意味着对于包含{尿布}的记录,对其中的r * 记录数都适用
要想找到支持度大于0.8的项集,我们就需要对所有的物品进行排列组合得到所有可能的项集,再进行支持度的计算。
这会十分耗时低效,我们将分析Apriori原理,该原理可以减小关联学习的计算量。
## Apriori原理
>a priori —— 一个先验。在拉丁语中指“来自以前”。我们在贝叶斯统计时经常使用先验知识作为判断的条件,这些知识来自领域的知识,先前的测量结果等等。
原理内容:如果某个项集是频繁的,那么它所有的子集也是频繁的。同理,如果某个项集是非频繁集,那么它的所有超集也是非频繁的。
利用此原理可有效降低项集的指数级增长
## 用Apriori发现频繁集
算法描述:
生成所有单个物品的项集列表
对每个数据记录
对每个项集
包含该项集就增加总计数值
对每个项集的的总数/总数据记录数
如果满足最小支持度,则保留此项集
对剩下的项集组合以生成包含两个元素的项集
重复上述去除项集的操作,直到所有的项集删除
### 生成候选项集
* 加载数据集
```
def load_data():
return [[1, 3, 4], [2, 3, 5], [1, 2, 3, 5], [2, 5]]
```
* 对 dataSet 进行去重,排序,放入 list 中,然后转换所有的元素为 frozenset
```
def create_c1(dataset):
c1 = []
for row in dataset:
for item in row:
if not [item] in c1:
c1.append([item])
c1.sort()
return list(map(frozenset, c1))
```
* 计算候选数据集 CK 在数据集 D 中的支持度,并返回支持度大于最小支持度(minSupport)的数据
```
def scan(dataset, candidate, min_support):
sscnt = {}
for row in dataset:
for can in candidate:
if can.issubset(row):
if can not in sscnt:
sscnt[can] = 1
else:
sscnt[can] += 1
num = float(len(dataset))
retlist = []
support_data = {}
for key in sscnt:
support = sscnt[key]/num
if support >= min_support:
retlist.insert(0, key)
support_data[key] = support
return retlist, support_data
dataset = load_data()
c1 = create_c1(dataset)
print(c1)
D = list(map(set, dataset))
D
```
### 使用0.5作为最小支持度
```
L1, supportdata0 = scan(D, c1, 0.5)
L1
```
发现4被删除,说明4没有达到最小支持度
## 组织完整Apriori算法
当集合中的个数大于0时
构建一个k个项组成的候选项集的列表
检查数据以确认每个项集都是频繁的
保留频繁项集并构建k+1项组成的候选项集的列表
### 在候选项集中生成新的项集,前k-2个项相同时,合并这两个项
```
def apriori_gen(Lk, k):
relist = []
lenLk = len(Lk)
for i in range(lenLk):
for j in range(i+1, lenLk):
L1 = list(Lk[i])[:k-2]
L2 = list(Lk[j])[:k-2]
L1.sort()
L2.sort()
if L1 == L2:
relist.append(Lk[i] | Lk[j])
return relist
```
### 构造所有可能的集合并算出其支持度
```
def apriori(dataset, min_sup = 0.5):
c1 = create_c1(dataset)
d = list(map(set, dataset))
L1, sup_data = scan(d, c1, min_sup) # 选择达到最小支持度的项集
L = [L1]
k = 2
while len(L[k-2]) > 0:
CK = apriori_gen(L[k-2], k) # 建立大小为K的项集
Lk, supK = scan(dataset, CK, min_sup) # 删选达到最小支持度的项集
sup_data.update(supK) # 更新支持字典
L.append(Lk) # 记录所有项集记录
k += 1
return L, sup_data
apriori(dataset)
apriori(dataset, 0.7)
```
## 从频繁项中挖掘关联规则
关联规则的量化方法:
* 可信度:
一条规则P-->H的可信度定义为:
support(P | H) / support(P)
**假设0, 1, 2 ——> 3 并不满足最小可信度要求,那么任何左部为{0, 1, 2}的子集的规则也不会满足最小可信度要求**
利用上述性质来减少需要测试的规则项目:
* 分级法:
首先从一个频繁项开始,创建一个规则列表,此列表右部只有一个元素,然后对这些规则进行测试
然后合并所有剩余规则来创建一个新的规则列表,其中列表右部包含两个元素
重复上述过程,直到列表右部不再增加
```
# 生成关联规则
def generateRules(L, supportData, minConf=0.7):
"""generateRules
Args:
L 频繁项集列表
supportData 频繁项集支持度的字典
minConf 最小置信度
Returns:
bigRuleList 可信度规则列表(关于 (A->B+置信度) 3个字段的组合)
"""
bigRuleList = []
# 假设 L = [[frozenset([1]), frozenset([3]), frozenset([2]), frozenset([5])], [frozenset([1, 3]), frozenset([2, 5]), frozenset([2, 3]), frozenset([3, 5])], [frozenset([2, 3, 5])]]
for i in range(1, len(L)):
# 获取频繁项集中每个组合的所有元素
for freqSet in L[i]:
# 假设: freqSet= frozenset([1, 3]), H1=[frozenset([1]), frozenset([3])]
# 组合总的元素并遍历子元素,并转化为 frozenset 集合,再存放到 list 列表中
H1 = [frozenset([item]) for item in freqSet]
# 2 个的组合,走 else, 2 个以上的组合,走 if
if (i > 1):
rulesFromConseq(freqSet, H1, supportData, bigRuleList, minConf)
else:
calcConf(freqSet, H1, supportData, bigRuleList, minConf)
return bigRuleList
# 计算可信度(confidence)
def calcConf(freqSet, H
, supportData, brl, minConf=0.7):
"""calcConf(对两个元素的频繁项,计算可信度,例如: {1,2}/{1} 或者 {1,2}/{2} 看是否满足条件)
Args:
freqSet 频繁项集中的元素,例如: frozenset([1, 3])
H 频繁项集中的元素的集合,例如: [frozenset([1]), frozenset([3])]
supportData 所有元素的支持度的字典
brl 关联规则列表的空数组
minConf 最小可信度
Returns:
prunedH 记录 可信度大于阈值的集合
"""
# 记录可信度大于最小可信度(minConf)的集合
prunedH = []
for conseq in H: # 假设 freqSet = frozenset([1, 3]), H = [frozenset([1]), frozenset([3])],那么现在需要求出 frozenset([1]) -> frozenset([3]) 的可信度和 frozenset([3]) -> frozenset([1]) 的可信度
print ('confData=', freqSet, H, conseq, freqSet-conseq)
conf = supportData[freqSet]/supportData[freqSet-conseq] # 支持度定义: a -> b = support(a | b) / support(a). 假设 freqSet = frozenset([1, 3]), conseq = [frozenset([1])],那么 frozenset([1]) 至 frozenset([3]) 的可信度为 = support(a | b) / support(a) = supportData[freqSet]/supportData[freqSet-conseq] = supportData[frozenset([1, 3])] / supportData[frozenset([1])]
if conf >= minConf:
# 只要买了 freqSet-conseq 集合,一定会买 conseq 集合(freqSet-conseq 集合和 conseq 集合是全集)
print(freqSet-conseq, '-->', conseq, 'conf:', conf)
brl.append((freqSet-conseq, conseq, conf))
prunedH.append(conseq)
return prunedH
# 递归计算频繁项集的规则
def rulesFromConseq(freqSet, H, supportData, brl, minConf=0.7):
"""rulesFromConseq
Args:
freqSet 频繁项集中的元素,例如: frozenset([2, 3, 5])
H 频繁项集中的元素的集合,例如: [frozenset([2]), frozenset([3]), frozenset([5])]
supportData 所有元素的支持度的字典
brl 关联规则列表的数组
minConf 最小可信度
"""
# H[0] 是 freqSet 的元素组合的第一个元素,并且 H 中所有元素的长度都一样,长度由 aprioriGen(H, m+1) 这里的 m + 1 来控制
# 该函数递归时,H[0] 的长度从 1 开始增长 1 2 3 ...
# 假设 freqSet = frozenset([2, 3, 5]), H = [frozenset([2]), frozenset([3]), frozenset([5])]
# 那么 m = len(H[0]) 的递归的值依次为 1 2
# 在 m = 2 时, 跳出该递归。假设再递归一次,那么 H[0] = frozenset([2, 3, 5]),freqSet = frozenset([2, 3, 5]) ,没必要再计算 freqSet 与 H[0] 的关联规则了。
m = len(H[0])
if (len(freqSet) > (m + 1)):
print('freqSet******************', len(freqSet), m + 1, freqSet, H, H[0])
# 生成 m+1 个长度的所有可能的 H 中的组合,假设 H = [frozenset([2]), frozenset([3]), frozenset([5])]
# 第一次递归调用时生成 [frozenset([2, 3]), frozenset([2, 5]), frozenset([3, 5])]
# 第二次 。。。没有第二次,递归条件判断时已经退出了
Hmp1 = apriori_gen(H, m+1)
# 返回可信度大于最小可信度的集合
Hmp1 = calcConf(freqSet, Hmp1, supportData, brl, minConf)
print('Hmp1=', Hmp1)
print('len(Hmp1)=', len(Hmp1), 'len(freqSet)=', len(freqSet))
# 计算可信度后,还有数据大于最小可信度的话,那么继续递归调用,否则跳出递归
if (len(Hmp1) > 1):
print('----------------------', Hmp1)
# print len(freqSet), len(Hmp1[0]) + 1
rulesFromConseq(freqSet, Hmp1, supportData, brl, minConf)
L, supportdata = apriori(dataset, min_sup=0.5)
rules = generateRules(L, supportdata, minConf=0.7)
rules
```
### 计算项集的置信度
```
def calc_conf(freqset, H, supportdata, br1, min_conf=0.7):
prunedH = []
for conseq in H:
conf = supportdata[freqset] / supportdata[freqset - conseq]
if conf >= min_conf:
print(freqset-conseq ,'——>', conseq, 'conf:',conf )
br1.append((freqset-conseq, conseq, conf))
prunedH.append(conseq)
return prunedH
```
### 将多元素集合拆分为两个子集合
```
def rules_from_conseq(freqset, H, supportdata, br1,min_conf=0.7):
m = len(H[0])
if len(freqset) > m+1:
Hmp1 = apriori_gen(H, m+1)
Hmp1 = calc_conf(freqset, Hmp1, supportdata, br1, min_conf)
if len(Hmp1) > 1:
rules_from_conseq(freqset, Hmp1, supportdata, br1, min_conf)
```
### 生成规则
```
def generate_rules(L, supportdata, min_conf=0.7):
big_rule_list = []
for i in range(1, len(L)):
for freqset in L[i]:
H1 = [frozenset([item]) for item in freqset]
if i > 1:
rules_from_conseq(freqset, H1, supportdata, big_rule_list, min_conf)
else:
calc_conf(freqset, H1, supportdata, big_rule_list, min_conf)
return big_rule_list
generate_rules(L, supportdata, min_conf=0.7)
generate_rules(L, supportdata, min_conf=0.5)
```
## 示例:发现国会投票的模式(API废旧无法使用)
## 示例:发现毒蘑菇的相似特征
```
mushroom = [line.split() for line in open('mushroom.dat').readlines()]
L, supportdata = apriori(mushroom, min_sup=0.3)
mushroom
```
#### 寻找包含有毒特征值为2的频繁项集
```
for item in L[1]: # 项集元素为2
if item.intersection('2'):
print(item)
for item in L[3]: 项集元素数为4
if item.intersection('2'):
print(item)
```
| github_jupyter |
# Make Bag-of-visual-words
* Load Image's descriptor for all images.
* Find corresponding visual word index for each descriptor.
* Save list of visual word index.
# For Oxford 5k dataset, You can use already provided visual words
```
%%time
# Oxford 5k dataset provides already converted visual words. We could use this one
import os
import pickle
oxf5k_visualword_dir = './data/word_oxc1_hesaff_sift_16M_1M'
work_dir = "./oxfk5_provided"
if not os.path.exists(work_dir):
os.mkdir(work_dir)
filelist = os.listdir(oxf5k_visualword_dir)
filelist.sort()
# print(filelist)
# for parent_dir, _, files in os.walk(oxf5k_visualword_dir):
# print(files)
bow_dict = {}
count_descriptor = 0
image_feature_count_info = []
for filename in filelist:
filepath = os.path.join(oxf5k_visualword_dir, filename)
image_name = filename.replace(".txt", "")
visual_words = []
with open(filepath) as f:
lines = list(map(lambda x: x.strip(), f.readlines()[2:])) # ignore first two lines
for l in lines:
val = l.split(" ")
visual_word_index = int(val[0])-1 # This data use 1 to 1,000,000. convert to zero-based so 0 to 999,999
visual_words.append(visual_word_index)
# print('{} descriptor {}'.format(filename, l))
count_descriptor = count_descriptor + len(lines)
image_feature_count_info.append((image_name, len(visual_words)))
bow_dict[image_name] = sorted(visual_words)
# break
# print('bow_dict:', bow_dict)
print('count_descriptor:', count_descriptor)
pickle.dump(bow_dict, open(os.path.join(work_dir, 'bow_dict_word_oxc1_hesaff_sift_16M_1M_pretrained.pkl'), 'wb'))
```
# Other case, you have to build it from scratch
After you prepared everything, run below code
## Preparation list
* (image, descriptor_list) tuple
* centroids
## Requirements
* Image name and its associated 128d descriptors
* visual words assigner
* For a given 128d descriptor, it tells index of visual words.
* After you ran k-means clustering, you could use it to get nearest centroid's id
TODO: determine design choice when we use encoding such as Product Quantization.
1. keep centroid as PQ code. It means we always have to keep encoder parts.
2. keep centroid as original vector. It means there is room for additional error when we assign each descriptor to the nearest centroid.
```
# Load image descriptor dictionary, and assign each descriptor to visual words
import pickle
import os
import numpy as np
import pqkmeans
from tqdm import tqdm
from multiprocessing import Pool, TimeoutError
image_descriptor_dict_path = 'image_descriptor_dict_oxc5k_extracted_hesaff_rootsift_13M.pkl'
# You should use matching encoder that was used to do PQk-means clustering.
# encoder_save_path = 'pqencoder_100k_random_sample_from_16M.pkl'
# cluster_center_save_path = 'clustering_centers_in_pqcode_numpy.npy'
# encoder_save_path = 'pqencoder_1000k_random_sample_from_16M.pkl'
# cluster_center_save_path = 'clustering_centers_numpy_16M_feature_1000k_coodebook_131k_cluster.npy'
work_dir = "./output_oxf5k_extracted_13M_rootsift_1M_vocab_pqkmeans_1M_codebook_train"
encoder_save_path = os.path.join(work_dir, 'pqencoder.pkl')
cluster_center_save_path = os.path.join(work_dir, 'centroids_in_pqcodes.npy')
output_bow_dict_save_path = os.path.join(work_dir, 'bow_dict.pkl')
# encoder_save_path = 'encoder.pkl'
# cluster_center_save_path = 'clustering_centers_numpy.npy'
# For PQ-kmeans clustering, we first convert query to PQ codes.
with open(encoder_save_path, 'rb') as f:
encoder = pickle.load(f)
clustering_centers_in_pqcode_numpy = np.load(cluster_center_save_path)
# print('cluster centers shape: ', clustering_centers_in_pqcode_numpy.shape)
k = clustering_centers_in_pqcode_numpy.shape[0]
print('number of clusters:', k)
engine = pqkmeans.clustering.PQKMeans(encoder=encoder, k=k, iteration=1, verbose=False, init_centers=clustering_centers_in_pqcode_numpy)
bow_dict = {}
def run(val):
image_name, tupval = val
descriptors = tupval[1]
data_points_pqcodes = encoder.transform(descriptors)
# print('num_descriptors:', len(data_points_pqcodes))
# print('num_descriptors shape:', data_points_pqcodes.shape)
# TODO: speedup by using pq-kmeans assignment step.
# visual_words = get_assigned_center_index(data_points_pqcodes, clustering_centers_in_pqcode_numpy)
visual_words = engine.predict(data_points_pqcodes) # Fast assignment step.
return (image_name, list(set(list(visual_words))))
if __name__ == "__main__":
with open(image_descriptor_dict_path, 'rb') as f:
# key: image_name, value: tuple of (keypoint_nparray, descriptor_nparray)
# descriptor_nparray: 2d numpy array of shape (num_descriptor, dim_descriptor)
image_descriptor_dict = pickle.load(f)
print('num images:', len(image_descriptor_dict))
pool = Pool(processes=20) # start 20 worker processes
# print same numbers in arbitrary order
for image_name, bow in tqdm(pool.imap_unordered(run, image_descriptor_dict.items()), total=len(image_descriptor_dict)):
bow_dict[image_name] = bow
print('done')
pickle.dump(bow_dict, open(output_bow_dict_save_path, 'wb'))
# Timing: 24min for 5062 images with 16M features. 100k leanred codebook with 4 subspaces. 2^17 clusters.
# Timing: 38min for 5062 images with 16M features. 1M leanred codebook with 8 subspaces. 2^17 clusters.
```
| github_jupyter |
# Setting up a Ray cluster with SmartSim
## 1. Start the cluster
We set up a SmartSim experiment, which will handle the launch of the Ray cluster.
First we import the relevant modules and set up variables. `NUM_NODES` is the number of Ray nodes we will deploy: the first one will be the head node, and we will run one node on each host.
```
import numpy as np
import os
import ray
from ray import tune
import ray.util
from smartsim import Experiment
from smartsim.exp.ray import RayCluster
NUM_NODES = 3
CPUS_PER_WORKER = 18
launcher='slurm'
```
Now we define a SmartSim experiment which will spin the Ray cluster. The output files will be located in the `ray-cluster` directory (relative to the path from where we are executing this notebook). We are limiting the number each ray node can use to `CPUS_PER_WORKER`: if we wanted to let it use all the cpus, it would suffice not to pass `ray_args`.
Notice that the cluster will be password-protected (the password, generated internally, will be shared with worker nodes).
If the hosts are attached to multiple interfaces (e.g. `ib`, `eth0`, ...) we can specify to which one the Ray nodes should bind: it is recommended to always choose the one offering the best performances. On a Cray XC, for example, this will be `ipogif0`.
To connect to the cluster, we will use the Ray client. Note that this approach only works with `ray>=1.6`, for previous versions, one has to add `password=None` to the `RayCluster` constructor.
```
exp = Experiment("ray-cluster", launcher=launcher)
cluster = RayCluster(name="ray-cluster", run_args={}, ray_args={"num-cpus": CPUS_PER_WORKER},
launcher=launcher, num_nodes=NUM_NODES, batch=False, interface="ipogif0")
```
We now generate the needed directories. If an experiment with the same name already exists, this call will fail, to avoid overwriting existing results. If we want to overwrite, we can simply pass `overwrite=True` to `exp.generate()`.
```
exp.generate(cluster, overwrite=True)
```
Now we are ready to start the cluster!
```
exp.start(cluster, block=False, summary=False)
```
## 2. Start the ray driver script
Now we can just connect to our running server.
```
ctx = ray.init("ray://"+cluster.get_head_address()+":10001")
```
We can check that all resources are set properly.
```
print('''This cluster consists of
{} nodes in total
{} CPU resources in total
and the head node is running at {}'''.format(len(ray.nodes()), ray.cluster_resources()['CPU'], cluster.get_head_address()))
```
And we can run a Ray Tune example, to see that everything is working.
```
tune.run(
"PPO",
stop={"episode_reward_max": 200},
config={
"framework": "torch",
"env": "CartPole-v0",
"num_gpus": 0,
"lr": tune.grid_search(np.linspace (0.001, 0.01, 50).tolist()),
"log_level": "ERROR",
},
local_dir=os.path.join(exp.exp_path, "ray_log"),
verbose=0,
fail_fast=True,
log_to_file=True,
)
```
When the Ray job is running, we can connect to the Ray dashboard to monitor the evolution of the experiment. To do this, if Ray is running on a compute node of a remote system, we need to setup a SSH tunnel (we will see later how), to forward the port on which the dashboard is published to our local system. For example if the head address (printed in the cell above) is `<head_ip_address>`, and the system name is `<remote_sytem_name>`, we can establish a tunnel to the dashboard opening a terminal on the local system and entering
```bash
ssh -L 8265:<head_ip_address>:8265 <remote_system_name>
```
Then, from a browser on the local system, we can just go to the address `http://localhost:8265` to see the dashboard.
There are two things to know if something does not work:
1. We are using `8265` as a port, which is the default dashboard port. If that port is not free, we can bind the dashboard to another port, e.g. `PORT_NUMBER` (by adding `"dashboard-port": str(PORT_NUMBER)` to `ray_args` when creating the cluster) and the command has to be changed accordingly.
2. If the port forwarding fails, it is possible that the interface is not reachable. In that case, one can add `"dashboard-address": "0.0.0.0"` to `ray_args` when creating the cluster, to bind the dashboard to all interfaces, or select a visible address if one knows it. We can then just use the node name (or its public IP) to establish the tunnel, by entering (on the local terminal)
```bash
ssh -L 8265:<node_name_or_public_IP>:8265 <remote_system_name>
```
please refer to your system guide to find out how you can get the name and the address of a node.
## 3. Stop cluster and release allocation
We first shut down the Ray runtime, then disconnect the context.
```
ray.shutdown()
ctx.disconnect()
```
Now that all is gracefully stopped, we can stop the job on the allocation.
```
exp.stop(cluster)
```
| github_jupyter |
# Multi-Signal Multi-Channel Time Encoding and Decoding
This notebook provides an example where two signals are encoded and decoded using three integrate-and-fire time encoding machines. It also reproduces the figure in the paper "Encoding and Decoding Mixed Bandlimited Signals using Spiking Integrate-and-Fire Neurons", Submitted to ICASSP 2020
```
from header import *
import Figure3
import os
%matplotlib inline
```
## Create a set of two randomly generated bandlimited signals
x_param will be a list of two bandlimited signals, each represented by a sum of sincs at equally spaced locations. x will hold the samples of x_param taken at the time provided in the vector t.
```
end_time = 20
sinc_padding = 3
delta_t = 1e-4
t = np.arange(0,end_time+delta_t, delta_t)
Omega = np.pi
seed = 0
np.random.seed(int(seed))
num_signals = 2
x = np.zeros((num_signals, len(t)))
x_param = []
for n in range(num_signals):
x_param.append(bandlimitedSignal(Omega))
x_param[-1].random(t, padding = sinc_padding)
x[n,:] = x_param[-1].sample(t)
plt.figure()
plt.plot(t,x.T)
plt.xlabel('Time (s)')
plt.ylabel('Input Signals')
```
## Create a set of three time encoding machines to perform the sampling and use them to sample the input signals
First, we need to define the mixing matrix A, and the parameters of the machines, kappa, delta and b.
A timeEncoder object is used to perform the sampling, and the output is encapsulated in a spikeTimes object.
```
num_channels = 3
A = [[0.3,0.8],[-0.3,0.2],[0.4,0.6]]
kappa = [0.4,1.3,0.7]
delta = [0.7,0.6,0.5]
b = [0.2,0.9,0.4]
tem_mult = timeEncoder(kappa, delta, b, A)
spikes = tem_mult.encode_precise(x_param, Omega, end_time)
spikes_of_ch_1 = spikes.get_spikes_of(0)
plt.figure()
for s in spikes_of_ch_1:
plt.axvline(s, linestyle='--')
plt.xlabel ('Time (s)')
plt.ylabel ('Spike Times of Channel 1')
```
## Reconstruction using recursive algorithm
The algorithm returns signals which are sampled at times t.
```
rec_mult = tem_mult.decode_recursive(spikes, t, x_param[0].get_sinc_locs(), Omega, delta_t, num_iterations = 100)
plt.figure()
plt.rc("text", usetex=True)
plt.subplot(2,1,1)
plt.plot(t, x[0,:], label = 'input')
plt.plot(t, rec_mult[0,:], label = 'reconstruction')
plt.legend(loc = 'best')
plt.ylabel(r'$x_1(t)$')
plt.subplot(2,1,2)
plt.plot(t, x[1,:], label = 'input')
plt.plot(t, rec_mult[1,:], label = 'reconstruction')
plt.ylabel(r'$x_2(t)$')
plt.legend(loc = 'best')
plt.xlabel("Time (s)")
figure3_data_filename = "../Data/Figure3.pkl"
if(not os.path.isfile(figure3_data_filename)):
Figure3.GetData()
Figure3.GenerateFigure()
```
| github_jupyter |
# Session 4: Visualizing Representations
## Assignment: Deep Dream and Style Net
<p class='lead'>
Creative Applications of Deep Learning with Google's Tensorflow
Parag K. Mital
Kadenze, Inc.
</p>
# Overview
In this homework, we'll first walk through visualizing the gradients of a trained convolutional network. Recall from the last session that we had trained a variational convolutional autoencoder. We also trained a deep convolutional network. In both of these networks, we learned only a few tools for understanding how the model performs. These included measuring the loss of the network and visualizing the `W` weight matrices and/or convolutional filters of the network.
During the lecture we saw how to visualize the gradients of Inception, Google's state of the art network for object recognition. This resulted in a much more powerful technique for understanding how a network's activations transform or accentuate the representations in the input space. We'll explore this more in Part 1.
We also explored how to use the gradients of a particular layer or neuron within a network with respect to its input for performing "gradient ascent". This resulted in Deep Dream. We'll explore this more in Parts 2-4.
We also saw how the gradients at different layers of a convolutional network could be optimized for another image, resulting in the separation of content and style losses, depending on the chosen layers. This allowed us to synthesize new images that shared another image's content and/or style, even if they came from separate images. We'll explore this more in Part 5.
Finally, you'll packaged all the GIFs you create throughout this notebook and upload them to Kadenze.
<a name="learning-goals"></a>
# Learning Goals
* Learn how to inspect deep networks by visualizing their gradients
* Learn how to "deep dream" with different objective functions and regularization techniques
* Learn how to "stylize" an image using content and style losses from different images
# Table of Contents
<!-- MarkdownTOC autolink=true autoanchor=true bracket=round -->
- [Part 1 - Pretrained Networks](#part-1---pretrained-networks)
- [Graph Definition](#graph-definition)
- [Preprocess/Deprocessing](#preprocessdeprocessing)
- [Tensorboard](#tensorboard)
- [A Note on 1x1 Convolutions](#a-note-on-1x1-convolutions)
- [Network Labels](#network-labels)
- [Using Context Managers](#using-context-managers)
- [Part 2 - Visualizing Gradients](#part-2---visualizing-gradients)
- [Part 3 - Basic Deep Dream](#part-3---basic-deep-dream)
- [Part 4 - Deep Dream Extensions](#part-4---deep-dream-extensions)
- [Using the Softmax Layer](#using-the-softmax-layer)
- [Fractal](#fractal)
- [Guided Hallucinations](#guided-hallucinations)
- [Further Explorations](#further-explorations)
- [Part 5 - Style Net](#part-5---style-net)
- [Network](#network)
- [Content Features](#content-features)
- [Style Features](#style-features)
- [Remapping the Input](#remapping-the-input)
- [Content Loss](#content-loss)
- [Style Loss](#style-loss)
- [Total Variation Loss](#total-variation-loss)
- [Training](#training)
- [Assignment Submission](#assignment-submission)
<!-- /MarkdownTOC -->
```
# First check the Python version
import sys
if sys.version_info < (3,4):
print('You are running an older version of Python!\n\n',
'You should consider updating to Python 3.4.0 or',
'higher as the libraries built for this course',
'have only been tested in Python 3.4 and higher.\n')
print('Try installing the Python 3.5 version of anaconda'
'and then restart `jupyter notebook`:\n',
'https://www.continuum.io/downloads\n\n')
# Now get necessary libraries
try:
import os
import numpy as np
import matplotlib.pyplot as plt
from skimage.transform import resize
from skimage import data
from scipy.misc import imresize
from scipy.ndimage.filters import gaussian_filter
import IPython.display as ipyd
import tensorflow as tf
from libs import utils, gif, datasets, dataset_utils, vae, dft, vgg16, nb_utils
except ImportError:
print("Make sure you have started notebook in the same directory",
"as the provided zip file which includes the 'libs' folder",
"and the file 'utils.py' inside of it. You will NOT be able",
"to complete this assignment unless you restart jupyter",
"notebook inside the directory created by extracting",
"the zip file or cloning the github repo. If you are still")
# We'll tell matplotlib to inline any drawn figures like so:
%matplotlib inline
plt.style.use('ggplot')
# Bit of formatting because I don't like the default inline code style:
from IPython.core.display import HTML
HTML("""<style> .rendered_html code {
padding: 2px 4px;
color: #c7254e;
background-color: #f9f2f4;
border-radius: 4px;
} </style>""")
```
<a name="part-1---pretrained-networks"></a>
# Part 1 - Pretrained Networks
In the libs module, you'll see that I've included a few modules for loading some state of the art networks. These include:
* [Inception v3](https://github.com/tensorflow/models/tree/master/inception)
- This network has been trained on ImageNet and its finaly output layer is a softmax layer denoting 1 of 1000 possible objects (+ 8 for unknown categories). This network is about only 50MB!
* [Inception v5](https://github.com/tensorflow/models/tree/master/inception)
- This network has been trained on ImageNet and its finaly output layer is a softmax layer denoting 1 of 1000 possible objects (+ 8 for unknown categories). This network is about only 50MB! It presents a few extensions to v5 which are not documented anywhere that I've found, as of yet...
* [Visual Group Geometry @ Oxford's 16 layer](http://www.robots.ox.ac.uk/~vgg/research/very_deep/)
- This network has been trained on ImageNet and its finaly output layer is a softmax layer denoting 1 of 1000 possible objects. This model is nearly half a gigabyte, about 10x larger in size than the inception network. The trade off is that it is very fast.
* [Visual Group Geometry @ Oxford's Face Recognition](http://www.robots.ox.ac.uk/~vgg/software/vgg_face/)
- This network has been trained on the VGG Face Dataset and its final output layer is a softmax layer denoting 1 of 2622 different possible people.
* [Illustration2Vec](http://illustration2vec.net)
- This network has been trained on illustrations and manga and its final output layer is 4096 features.
* [Illustration2Vec Tag](http://illustration2vec.net)
- Please do not use this network if you are under the age of 18 (seriously!)
- This network has been trained on manga and its final output layer is one of 1539 labels.
When we use a pre-trained network, we load a network's definition and its weights which have already been trained. The network's definition includes a set of operations such as convolutions, and adding biases, but all of their values, i.e. the weights, have already been trained.
<a name="graph-definition"></a>
## Graph Definition
In the libs folder, you will see a few new modules for loading the above pre-trained networks. Each module is structured similarly to help you understand how they are loaded and include example code for using them. Each module includes a `preprocess` function for using before sending the image to the network. And when using deep dream techniques, we'll be using the `deprocess` function to undo the `preprocess` function's manipulations.
Let's take a look at loading one of these. Every network except for `i2v` includes a key 'labels' denoting what labels the network has been trained on. If you are under the age of 18, please do not use the `i2v_tag model`, as its labels are unsuitable for minors.
Let's load the libaries for the different pre-trained networks:
```
from libs import vgg16, inception, i2v
```
Now we can load a pre-trained network's graph and any labels. Explore the different networks in your own time.
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
# Stick w/ Inception for now, and then after you see how
# the next few sections work w/ this network, come back
# and explore the other networks.
net = inception.get_inception_model(version='v5')
# net = inception.get_inception_model(version='v3')
# net = vgg16.get_vgg_model()
# net = vgg16.get_vgg_face_model()
# net = i2v.get_i2v_model()
# net = i2v.get_i2v_tag_model()
```
Each network returns a dictionary with the following keys defined. Every network has a key for "labels" except for "i2v", since this is a feature only network, e.g. an unsupervised network, and does not have labels.
```
print(net.keys())
```
<a name="preprocessdeprocessing"></a>
## Preprocess/Deprocessing
Each network has a preprocessing/deprocessing function which we'll use before sending the input to the network. This preprocessing function is slightly different for each network. Recall from the previous sessions what preprocess we had done before sending an image to a network. We would often normalize the input by subtracting the mean and dividing by the standard deviation. We'd also crop/resize the input to a standard size. We'll need to do this for each network except for the Inception network, which is a true convolutional network and does not require us to do this (will be explained in more depth later).
Whenever we `preprocess` the image, and want to visualize the result of adding back the gradient to the input image (when we use deep dream), we'll need to use the `deprocess` function stored in the dictionary. Let's explore how these work. We'll confirm this is performing the inverse operation, let's try to preprocess the image, then I'll have you try to deprocess it.
```
# First, let's get an image:
og = plt.imread('clinton.png')[..., :3]
plt.imshow(og)
print(og.min(), og.max())
```
Let's now try preprocessing this image. The function for preprocessing is inside the module we used to load it. For instance, for `vgg16`, we can find the `preprocess` function as `vgg16.preprocess`, or for `inception`, `inception.preprocess`, or for `i2v`, `i2v.preprocess`. Or, we can just use the key `preprocess` in our dictionary `net`, as this is just convenience for us to access the corresponding preprocess function.
```
# Now call the preprocess function. This will preprocess our
# image ready for being input to the network, except for changes
# to the dimensions. I.e., we will still need to convert this
# to a 4-dimensional Tensor once we input it to the network.
# We'll see how that works later.
img = net['preprocess'](og)
print(img.min(), img.max())
```
Let's undo the preprocessing. Recall that the `net` dictionary has the key `deprocess` which is the function we need to use on our processed image, `img`.
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
deprocessed = ...
plt.imshow(deprocessed)
plt.show()
```
<a name="tensorboard"></a>
## Tensorboard
I've added a utility module called `nb_utils` which includes a function `show_graph`. This will use [Tensorboard](https://www.tensorflow.org/versions/r0.10/how_tos/graph_viz/index.html) to draw the computational graph defined by the various Tensorflow functions. I didn't go over this during the lecture because there just wasn't enough time! But explore it in your own time if it interests you, as it is a really unique tool which allows you to monitor your network's training progress via a web interface. It even lets you monitor specific variables or processes within the network, e.g. the reconstruction of an autoencoder, without having to print to the console as we've been doing. We'll just be using it to draw the pretrained network's graphs using the utility function I've given you.
Be sure to interact with the graph and click on the various modules.
For instance, if you've loaded the `inception` v5 network, locate the "input" to the network. This is where we feed the image, the input placeholder (typically what we've been denoting as `X` in our own networks). From there, it goes to the "conv2d0" variable scope (i.e. this uses the code: `with tf.variable_scope("conv2d0")` to create a set of operations with the prefix "conv2d0/". If you expand this scope, you'll see another scope, "pre_relu". This is created using another `tf.variable_scope("pre_relu")`, so that any new variables will have the prefix "conv2d0/pre_relu". Finally, inside here, you'll see the convolution operation (`tf.nn.conv2d`) and the 4d weight tensor, "w" (e.g. created using `tf.get_variable`), used for convolution (and so has the name, "conv2d0/pre_relu/w". Just after the convolution is the addition of the bias, b. And finally after exiting the "pre_relu" scope, you should be able to see the "conv2d0" operation which applies the relu nonlinearity. In summary, that region of the graph can be created in Tensorflow like so:
```python
input = tf.placeholder(...)
with tf.variable_scope('conv2d0'):
with tf.variable_scope('pre_relu'):
w = tf.get_variable(...)
h = tf.nn.conv2d(input, h, ...)
b = tf.get_variable(...)
h = tf.nn.bias_add(h, b)
h = tf.nn.relu(h)
```
```
nb_utils.show_graph(net['graph_def'])
```
If you open up the "mixed3a" node above (double click on it), you'll see the first "inception" module. This network encompasses a few advanced concepts that we did not have time to discuss during the lecture, including residual connections, feature concatenation, parallel convolution streams, 1x1 convolutions, and including negative labels in the softmax layer. I'll expand on the 1x1 convolutions here, but please feel free to skip ahead if this isn't of interest to you.
<a name="a-note-on-1x1-convolutions"></a>
## A Note on 1x1 Convolutions
The 1x1 convolutions are setting the `ksize` parameter of the kernels to 1. This is effectively allowing you to change the number of dimensions. Remember that you need a 4-d tensor as input to a convolution. Let's say its dimensions are $\text{N}\ x\ \text{H}\ x\ \text{W}\ x\ \text{C}_I$, where $\text{C}_I$ represents the number of channels the image has. Let's say it is an RGB image, then $\text{C}_I$ would be 3. Or later in the network, if we have already convolved it, it might be 64 channels instead. Regardless, when you convolve it w/ a $\text{K}_H\ x\ \text{K}_W\ x\ \text{C}_I\ x\ \text{C}_O$ filter, where $\text{K}_H$ is 1 and $\text{K}_W$ is also 1, then the filters size is: $1\ x\ 1\ x\ \text{C}_I$ and this is perfomed for each output channel $\text{C}_O$. What this is doing is filtering the information only in the channels dimension, not the spatial dimensions. The output of this convolution will be a $\text{N}\ x\ \text{H}\ x\ \text{W}\ x\ \text{C}_O$ output tensor. The only thing that changes in the output is the number of output filters.
The 1x1 convolution operation is essentially reducing the amount of information in the channels dimensions before performing a much more expensive operation, e.g. a 3x3 or 5x5 convolution. Effectively, it is a very clever trick for dimensionality reduction used in many state of the art convolutional networks. Another way to look at it is that it is preserving the spatial information, but at each location, there is a fully connected network taking all the information from every input channel, $\text{C}_I$, and reducing it down to $\text{C}_O$ channels (or could easily also be up, but that is not the typical use case for this). So it's not really a convolution, but we can use the convolution operation to perform it at every location in our image.
If you are interested in reading more about this architecture, I highly encourage you to read [Network in Network](https://arxiv.org/pdf/1312.4400v3.pdf), Christian Szegedy's work on the [Inception network](http://www.cs.unc.edu/~wliu/papers/GoogLeNet.pdf), Highway Networks, Residual Networks, and Ladder Networks.
In this course, we'll stick to focusing on the applications of these, while trying to delve as much into the code as possible.
<a name="network-labels"></a>
## Network Labels
Let's now look at the labels:
```
net['labels']
label_i = 851
print(net['labels'][label_i])
```
<a name="using-context-managers"></a>
## Using Context Managers
Up until now, we've mostly used a single `tf.Session` within a notebook and didn't give it much thought. Now that we're using some bigger models, we're going to have to be more careful. Using a big model and being careless with our session can result in a lot of unexpected behavior, program crashes, and out of memory errors. The VGG network and the I2V networks are quite large. So we'll need to start being more careful with our sessions using context managers.
Let's see how this works w/ VGG:
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
# Load the VGG network. Scroll back up to where we loaded the inception
# network if you are unsure. It is inside the "vgg16" module...
net = ..
assert(net['labels'][0] == (0, 'n01440764 tench, Tinca tinca'))
# Let's explicity use the CPU, since we don't gain anything using the GPU
# when doing Deep Dream (it's only a single image, benefits come w/ many images).
device = '/cpu:0'
# We'll now explicitly create a graph
g = tf.Graph()
# And here is a context manager. We use the python "with" notation to create a context
# and create a session that only exists within this indent, as soon as we leave it,
# the session is automatically closed! We also tell the session which graph to use.
# We can pass a second context after the comma,
# which we'll use to be explicit about using the CPU instead of a GPU.
with tf.Session(graph=g) as sess, g.device(device):
# Now load the graph_def, which defines operations and their values into `g`
tf.import_graph_def(net['graph_def'], name='net')
# Now we can get all the operations that belong to the graph `g`:
names = [op.name for op in g.get_operations()]
print(names)
```
<a name="part-2---visualizing-gradients"></a>
# Part 2 - Visualizing Gradients
Now that we know how to load a network and extract layers from it, let's grab only the pooling layers:
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
# First find all the pooling layers in the network. You can
# use list comprehension to iterate over all the "names" we just
# created, finding whichever ones have the name "pool" in them.
# Then be sure to append a ":0" to the names
features = ...
# Let's print them
print(features)
# This is what we want to have at the end. You could just copy this list
# if you are stuck!
assert(features == ['net/pool1:0', 'net/pool2:0', 'net/pool3:0', 'net/pool4:0', 'net/pool5:0'])
```
Let's also grab the input layer:
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
# Use the function 'get_tensor_by_name' and the 'names' array to help you
# get the first tensor in the network. Remember you have to add ":0" to the
# name to get the output of an operation which is the tensor.
x = ...
assert(x.name == 'net/images:0')
```
We'll now try to find the gradient activation that maximizes a layer with respect to the input layer `x`.
```
def plot_gradient(img, x, feature, g, device='/cpu:0'):
"""Let's visualize the network's gradient activation
when backpropagated to the original input image. This
is effectively telling us which pixels contribute to the
predicted layer, class, or given neuron with the layer"""
# We'll be explicit about the graph and the device
# by using a context manager:
with tf.Session(graph=g) as sess, g.device(device):
saliency = tf.gradients(tf.reduce_mean(feature), x)
this_res = sess.run(saliency[0], feed_dict={x: img})
grad = this_res[0] / np.max(np.abs(this_res))
return grad
```
Let's try this w/ an image now. We're going to use the `plot_gradient` function to help us. This is going to take our input image, run it through the network up to a layer, find the gradient of the mean of that layer's activation with respect to the input image, then backprop that gradient back to the input layer. We'll then visualize the gradient by normalizing its values using the `utils.normalize` function.
```
og = plt.imread('clinton.png')[..., :3]
img = net['preprocess'](og)[np.newaxis]
fig, axs = plt.subplots(1, len(features), figsize=(20, 10))
for i in range(len(features)):
axs[i].set_title(features[i])
grad = plot_gradient(img, x, g.get_tensor_by_name(features[i]), g)
axs[i].imshow(utils.normalize(grad))
```
<a name="part-3---basic-deep-dream"></a>
# Part 3 - Basic Deep Dream
In the lecture we saw how Deep Dreaming takes the backpropagated gradient activations and simply adds it to the image, running the same process again and again in a loop. We also saw many tricks one can add to this idea, such as infinitely zooming into the image by cropping and scaling, adding jitter by randomly moving the image around, or adding constraints on the total activations.
Have a look here for inspiration:
https://research.googleblog.com/2015/06/inceptionism-going-deeper-into-neural.html
https://photos.google.com/share/AF1QipPX0SCl7OzWilt9LnuQliattX4OUCj_8EP65_cTVnBmS1jnYgsGQAieQUc1VQWdgQ?key=aVBxWjhwSzg2RjJWLWRuVFBBZEN1d205bUdEMnhB
https://mtyka.github.io/deepdream/2016/02/05/bilateral-class-vis.html
Let's stick the necessary bits in a function and try exploring how deep dream amplifies the representations of the chosen layers:
```
def dream(img, gradient, step, net, x, n_iterations=50, plot_step=10):
# Copy the input image as we'll add the gradient to it in a loop
img_copy = img.copy()
fig, axs = plt.subplots(1, n_iterations // plot_step, figsize=(20, 10))
with tf.Session(graph=g) as sess, g.device(device):
for it_i in range(n_iterations):
# This will calculate the gradient of the layer we chose with respect to the input image.
this_res = sess.run(gradient[0], feed_dict={x: img_copy})[0]
# Let's normalize it by the maximum activation
this_res /= (np.max(np.abs(this_res) + 1e-8))
# Or alternatively, we can normalize by standard deviation
# this_res /= (np.std(this_res) + 1e-8)
# Or we could use the `utils.normalize function:
# this_res = utils.normalize(this_res)
# Experiment with all of the above options. They will drastically
# effect the resulting dream, and really depend on the network
# you use, and the way the network handles normalization of the
# input image, and the step size you choose! Lots to explore!
# Then add the gradient back to the input image
# Think about what this gradient represents?
# It says what direction we should move our input
# in order to meet our objective stored in "gradient"
img_copy += this_res * step
# Plot the image
if (it_i + 1) % plot_step == 0:
m = net['deprocess'](img_copy[0])
axs[it_i // plot_step].imshow(m)
# We'll run it for 3 iterations
n_iterations = 3
# Think of this as our learning rate. This is how much of
# the gradient we'll add back to the input image
step = 1.0
# Every 1 iterations, we'll plot the current deep dream
plot_step = 1
```
Let's now try running Deep Dream for every feature, each of our 5 pooling layers. We'll need to get the layer corresponding to our feature. Then find the gradient of this layer's mean activation with respect to our input, `x`. Then pass these to our `dream` function. This can take awhile (about 10 minutes using the CPU on my Macbook Pro).
```
for feature_i in range(len(features)):
with tf.Session(graph=g) as sess, g.device(device):
# Get a feature layer
layer = g.get_tensor_by_name(features[feature_i])
# Find the gradient of this layer's mean activation
# with respect to the input image
gradient = tf.gradients(tf.reduce_mean(layer), x)
# Dream w/ our image
dream(img, gradient, step, net, x, n_iterations=n_iterations, plot_step=plot_step)
```
Instead of using an image, we can use an image of noise and see how it "hallucinates" the representations that the layer most responds to:
```
noise = net['preprocess'](
np.random.rand(256, 256, 3) * 0.1 + 0.45)[np.newaxis]
```
We'll do the same thing as before, now w/ our noise image:
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
for feature_i in range(len(features)):
with tf.Session(graph=g) as sess, g.device(device):
# Get a feature layer
layer = ...
# Find the gradient of this layer's mean activation
# with respect to the input image
gradient = ...
# Dream w/ the noise image. Complete this!
dream(...)
```
<a name="part-4---deep-dream-extensions"></a>
# Part 4 - Deep Dream Extensions
As we saw in the lecture, we can also use the final softmax layer of a network to use during deep dream. This allows us to be explicit about the object we want hallucinated in an image.
<a name="using-the-softmax-layer"></a>
## Using the Softmax Layer
Let's get another image to play with, preprocess it, and then make it 4-dimensional.
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
# Load your own image here
og = ...
plt.imshow(og)
# Preprocess the image and make sure it is 4-dimensional by adding a new axis to the 0th dimension:
img = ...
assert(img.ndim == 4)
# Let's get the softmax layer
print(names[-2])
layer = g.get_tensor_by_name(names[-2] + ":0")
# And find its shape
with tf.Session(graph=g) as sess, g.device(device):
layer_shape = tf.shape(layer).eval(feed_dict={x:img})
# We can find out how many neurons it has by feeding it an image and
# calculating the shape. The number of output channels is the last dimension.
n_els = layer_shape[-1]
# Let's pick a label. First let's print out every label and then find one we like:
print(net['labels'])
```
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
# Pick a neuron. Or pick a random one. This should be 0-n_els
neuron_i = ...
print(net['labels'][neuron_i])
assert(neuron_i >= 0 and neuron_i < n_els)
# And we'll create an activation of this layer which is very close to 0
layer_vec = np.ones(layer_shape) / 100.0
# Except for the randomly chosen neuron which will be very close to 1
layer_vec[..., neuron_i] = 0.99
```
Let's decide on some parameters of our deep dream. We'll need to decide how many iterations to run for. And we'll plot the result every few iterations, also saving it so that we can produce a GIF. And at every iteration, we need to decide how much to ascend our gradient.
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
# Explore different parameters for this section.
n_iterations = 51
plot_step = 5
# If you use a different network, you will definitely need to experiment
# with the step size, as each network normalizes the input image differently.
step = 0.2
```
Now let's dream. We're going to define a context manager to create a session and use our existing graph, and make sure we use the CPU device, as there is no gain in using GPU, and we have much more CPU memory than GPU memory.
```
imgs = []
with tf.Session(graph=g) as sess, g.device(device):
gradient = tf.gradients(tf.reduce_max(layer), x)
# Copy the input image as we'll add the gradient to it in a loop
img_copy = img.copy()
with tf.Session(graph=g) as sess, g.device(device):
for it_i in range(n_iterations):
# This will calculate the gradient of the layer we chose with respect to the input image.
this_res = sess.run(gradient[0], feed_dict={
x: img_copy, layer: layer_vec})[0]
# Let's normalize it by the maximum activation
this_res /= (np.max(np.abs(this_res) + 1e-8))
# Or alternatively, we can normalize by standard deviation
# this_res /= (np.std(this_res) + 1e-8)
# Then add the gradient back to the input image
# Think about what this gradient represents?
# It says what direction we should move our input
# in order to meet our objective stored in "gradient"
img_copy += this_res * step
# Plot the image
if (it_i + 1) % plot_step == 0:
m = net['deprocess'](img_copy[0])
plt.figure(figsize=(5, 5))
plt.grid('off')
plt.imshow(m)
plt.show()
imgs.append(m)
# Save the gif
gif.build_gif(imgs, saveto='softmax.gif')
ipyd.Image(url='softmax.gif?i={}'.format(
np.random.rand()), height=300, width=300)
```
<a name="fractal"></a>
## Fractal
During the lecture we also saw a simple trick for creating an infinite fractal: crop the image and then resize it. This can produce some lovely aesthetics and really show some strong object hallucinations if left long enough and with the right parameters for step size/normalization/regularization. Feel free to experiment with the code below, adding your own regularizations as shown in the lecture to produce different results!
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
n_iterations = 101
plot_step = 10
step = 0.1
crop = 1
imgs = []
n_imgs, height, width, *ch = img.shape
with tf.Session(graph=g) as sess, g.device(device):
# Explore changing the gradient here from max to mean
# or even try using different concepts we learned about
# when creating style net, such as using a total variational
# loss on `x`.
gradient = tf.gradients(tf.reduce_max(layer), x)
# Copy the input image as we'll add the gradient to it in a loop
img_copy = img.copy()
with tf.Session(graph=g) as sess, g.device(device):
for it_i in range(n_iterations):
# This will calculate the gradient of the layer
# we chose with respect to the input image.
this_res = sess.run(gradient[0], feed_dict={
x: img_copy, layer: layer_vec})[0]
# This is just one way we could normalize the
# gradient. It helps to look at the range of your image's
# values, e.g. if it is 0 - 1, or -115 to +115,
# and then consider the best way to normalize the gradient.
# For some networks, it might not even be necessary
# to perform this normalization, especially if you
# leave the dream to run for enough iterations.
# this_res = this_res / (np.std(this_res) + 1e-10)
this_res = this_res / (np.max(np.abs(this_res)) + 1e-10)
# Then add the gradient back to the input image
# Think about what this gradient represents?
# It says what direction we should move our input
# in order to meet our objective stored in "gradient"
img_copy += this_res * step
# Optionally, we could apply any number of regularization
# techniques... Try exploring different ways of regularizing
# gradient. ascent process. If you are adventurous, you can
# also explore changing the gradient above using a
# total variational loss, as we used in the style net
# implementation during the lecture. I leave that to you
# as an exercise!
# Crop a 1 pixel border from height and width
img_copy = img_copy[:, crop:-crop, crop:-crop, :]
# Resize (Note: in the lecture, we used scipy's resize which
# could not resize images outside of 0-1 range, and so we had
# to store the image ranges. This is a much simpler resize
# method that allows us to `preserve_range`.)
img_copy = resize(img_copy[0], (height, width), order=3,
clip=False, preserve_range=True
)[np.newaxis].astype(np.float32)
# Plot the image
if (it_i + 1) % plot_step == 0:
m = net['deprocess'](img_copy[0])
plt.grid('off')
plt.imshow(m)
plt.show()
imgs.append(m)
# Create a GIF
gif.build_gif(imgs, saveto='fractal.gif')
ipyd.Image(url='fractal.gif?i=2', height=300, width=300)
```
<a name="guided-hallucinations"></a>
## Guided Hallucinations
Instead of following the gradient of an arbitrary mean or max of a particular layer's activation, or a particular object that we want to synthesize, we can also try to guide our image to look like another image. One way to try this is to take one image, the guide, and find the features at a particular layer or layers. Then, we take our synthesis image and find the gradient which makes its own layers activations look like the guide image.
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
# Replace these with your own images!
guide_og = plt.imread(...)[..., :3]
dream_og = plt.imread(...)[..., :3]
assert(guide_og.ndim == 3 and guide_og.shape[-1] == 3)
assert(dream_og.ndim == 3 and dream_og.shape[-1] == 3)
```
Preprocess both images:
```
guide_img = net['preprocess'](guide_og)[np.newaxis]
dream_img = net['preprocess'](dream_og)[np.newaxis]
fig, axs = plt.subplots(1, 2, figsize=(7, 4))
axs[0].imshow(guide_og)
axs[1].imshow(dream_og)
```
Like w/ Style Net, we are going to measure how similar the features in the guide image are to the dream images. In order to do that, we'll calculate the dot product. Experiment with other measures such as l1 or l2 loss to see how this impacts the resulting Dream!
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
x = g.get_tensor_by_name(names[0] + ":0")
# Experiment with the weighting
feature_loss_weight = 1.0
with tf.Session(graph=g) as sess, g.device(device):
feature_loss = tf.Variable(0.0)
# Explore different layers/subsets of layers. This is just an example.
for feature_i in features[3:5]:
# Get the activation of the feature
layer = g.get_tensor_by_name(feature_i)
# Do the same for our guide image
guide_layer = sess.run(layer, feed_dict={x: guide_img})
# Now we need to measure how similar they are!
# We'll use the dot product, which requires us to first reshape both
# features to a 2D vector. But you should experiment with other ways
# of measuring similarity such as l1 or l2 loss.
# Reshape each layer to 2D vector
layer = tf.reshape(layer, [-1, 1])
guide_layer = guide_layer.reshape(-1, 1)
# Now calculate their dot product
correlation = tf.matmul(guide_layer.T, layer)
# And weight the loss by a factor so we can control its influence
feature_loss += feature_loss_weight * correlation
```
We'll now use another measure that we saw when developing Style Net during the lecture. This measure the pixel to pixel difference of neighboring pixels. What we're doing when we try to optimize a gradient that makes the mean differences small is saying, we want the difference to be low. This allows us to smooth our image in the same way that we did using the Gaussian to blur the image.
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
n_img, height, width, ch = dream_img.shape
# We'll weight the overall contribution of the total variational loss
# Experiment with this weighting
tv_loss_weight = 1.0
with tf.Session(graph=g) as sess, g.device(device):
# Penalize variations in neighboring pixels, enforcing smoothness
dx = tf.square(x[:, :height - 1, :width - 1, :] - x[:, :height - 1, 1:, :])
dy = tf.square(x[:, :height - 1, :width - 1, :] - x[:, 1:, :width - 1, :])
# We will calculate their difference raised to a power to push smaller
# differences closer to 0 and larger differences higher.
# Experiment w/ the power you raise this to to see how it effects the result
tv_loss = tv_loss_weight * tf.reduce_mean(tf.pow(dx + dy, 1.2))
```
Now we train just like before, except we'll need to combine our two loss terms, `feature_loss` and `tv_loss` by simply adding them! The one thing we have to keep in mind is that we want to minimize the `tv_loss` while maximizing the `feature_loss`. That means we'll need to use the negative `tv_loss` and the positive `feature_loss`. As an experiment, try just optimizing the `tv_loss` and removing the `feature_loss` from the `tf.gradients` call. What happens?
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
# Experiment with the step size!
step = 0.1
imgs = []
with tf.Session(graph=g) as sess, g.device(device):
# Experiment with just optimizing the tv_loss or negative tv_loss to understand what it is doing!
gradient = tf.gradients(-tv_loss + feature_loss, x)
# Copy the input image as we'll add the gradient to it in a loop
img_copy = dream_img.copy()
with tf.Session(graph=g) as sess, g.device(device):
sess.run(tf.global_variables_initializer())
for it_i in range(n_iterations):
# This will calculate the gradient of the layer we chose with respect to the input image.
this_res = sess.run(gradient[0], feed_dict={x: img_copy})[0]
# Let's normalize it by the maximum activation
this_res /= (np.max(np.abs(this_res) + 1e-8))
# Or alternatively, we can normalize by standard deviation
# this_res /= (np.std(this_res) + 1e-8)
# Then add the gradient back to the input image
# Think about what this gradient represents?
# It says what direction we should move our input
# in order to meet our objective stored in "gradient"
img_copy += this_res * step
# Plot the image
if (it_i + 1) % plot_step == 0:
m = net['deprocess'](img_copy[0])
plt.figure(figsize=(5, 5))
plt.grid('off')
plt.imshow(m)
plt.show()
imgs.append(m)
gif.build_gif(imgs, saveto='guided.gif')
ipyd.Image(url='guided.gif?i=0', height=300, width=300)
```
<a name="further-explorations"></a>
## Further Explorations
In the `libs` module, I've included a `deepdream` module which has two functions for performing Deep Dream and the Guided Deep Dream. Feel free to explore these to create your own deep dreams.
<a name="part-5---style-net"></a>
# Part 5 - Style Net
We'll now work on creating our own style net implementation. We've seen all the steps for how to do this during the lecture, and you can always refer to the [Lecture Transcript](lecture-4.ipynb) if you need to. I want to you to explore using different networks and different layers in creating your content and style losses. This is completely unexplored territory so it can be frustrating to find things that work. Think of this as your empty canvas! If you are really stuck, you will find a `stylenet` implementation under the `libs` module that you can use instead.
Have a look here for inspiration:
https://mtyka.github.io/code/2015/10/02/experiments-with-style-transfer.html
http://kylemcdonald.net/stylestudies/
<a name="network"></a>
## Network
Let's reset the graph and load up a network. I'll include code here for loading up any of our pretrained networks so you can explore each of them!
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
sess.close()
tf.reset_default_graph()
# Stick w/ VGG for now, and then after you see how
# the next few sections work w/ this network, come back
# and explore the other networks.
net = vgg16.get_vgg_model()
# net = vgg16.get_vgg_face_model()
# net = inception.get_inception_model(version='v5')
# net = inception.get_inception_model(version='v3')
# net = i2v.get_i2v_model()
# net = i2v.get_i2v_tag_model()
# Let's explicity use the CPU, since we don't gain anything using the GPU
# when doing Deep Dream (it's only a single image, benefits come w/ many images).
device = '/cpu:0'
# We'll now explicitly create a graph
g = tf.Graph()
```
Let's now import the graph definition into our newly created Graph using a context manager and specifying that we want to use the CPU.
```
# And here is a context manager. We use the python "with" notation to create a context
# and create a session that only exists within this indent, as soon as we leave it,
# the session is automatically closed! We also tel the session which graph to use.
# We can pass a second context after the comma,
# which we'll use to be explicit about using the CPU instead of a GPU.
with tf.Session(graph=g) as sess, g.device(device):
# Now load the graph_def, which defines operations and their values into `g`
tf.import_graph_def(net['graph_def'], name='net')
```
Let's then grab the names of every operation in our network:
```
names = [op.name for op in g.get_operations()]
```
Now we need an image for our content image and another one for our style image.
```
content_og = plt.imread('arles.png')[..., :3]
style_og = plt.imread('clinton.png')[..., :3]
fig, axs = plt.subplots(1, 2)
axs[0].imshow(content_og)
axs[0].set_title('Content Image')
axs[0].grid('off')
axs[1].imshow(style_og)
axs[1].set_title('Style Image')
axs[1].grid('off')
# We'll save these with a specific name to include in your submission
plt.imsave(arr=content_og, fname='content.png')
plt.imsave(arr=style_og, fname='style.png')
content_img = net['preprocess'](content_og)[np.newaxis]
style_img = net['preprocess'](style_og)[np.newaxis]
```
Let's see what the network classifies these images as just for fun:
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
# Grab the tensor defining the input to the network
x = ...
# And grab the tensor defining the softmax layer of the network
softmax = ...
for img in [content_img, style_img]:
with tf.Session(graph=g) as sess, g.device('/cpu:0'):
# Remember from the lecture that we have to set the dropout
# "keep probability" to 1.0.
res = softmax.eval(feed_dict={x: img,
'net/dropout_1/random_uniform:0': [[1.0]],
'net/dropout/random_uniform:0': [[1.0]]})[0]
print([(res[idx], net['labels'][idx])
for idx in res.argsort()[-5:][::-1]])
```
<a name="content-features"></a>
## Content Features
We're going to need to find the layer or layers we want to use to help us define our "content loss". Recall from the lecture when we used VGG, we used the 4th convolutional layer.
```
print(names)
```
Pick a layer for using for the content features. If you aren't using VGG remember to get rid of the dropout stuff!
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
# Experiment w/ different layers here. You'll need to change this if you
# use another network!
content_layer = 'net/conv3_2/conv3_2:0'
with tf.Session(graph=g) as sess, g.device('/cpu:0'):
content_features = g.get_tensor_by_name(content_layer).eval(
session=sess,
feed_dict={x: content_img,
'net/dropout_1/random_uniform:0': [[1.0]],
'net/dropout/random_uniform:0': [[1.0]]})
```
<a name="style-features"></a>
## Style Features
Let's do the same thing now for the style features. We'll use more than 1 layer though so we'll append all the features in a list. If you aren't using VGG remember to get rid of the dropout stuff!
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
# Experiment with different layers and layer subsets. You'll need to change these
# if you use a different network!
style_layers = ['net/conv1_1/conv1_1:0',
'net/conv2_1/conv2_1:0',
'net/conv3_1/conv3_1:0',
'net/conv4_1/conv4_1:0',
'net/conv5_1/conv5_1:0']
style_activations = []
with tf.Session(graph=g) as sess, g.device('/cpu:0'):
for style_i in style_layers:
style_activation_i = g.get_tensor_by_name(style_i).eval(
feed_dict={x: style_img,
'net/dropout_1/random_uniform:0': [[1.0]],
'net/dropout/random_uniform:0': [[1.0]]})
style_activations.append(style_activation_i)
```
Now we find the gram matrix which we'll use to optimize our features.
```
style_features = []
for style_activation_i in style_activations:
s_i = np.reshape(style_activation_i, [-1, style_activation_i.shape[-1]])
gram_matrix = np.matmul(s_i.T, s_i) / s_i.size
style_features.append(gram_matrix.astype(np.float32))
```
<a name="remapping-the-input"></a>
## Remapping the Input
We're almost done building our network. We just have to change the input to the network to become "trainable". Instead of a placeholder, we'll have a `tf.Variable`, which allows it to be trained. We could set this to the content image, another image entirely, or an image of noise. Experiment with all three options!
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
tf.reset_default_graph()
g = tf.Graph()
# Get the network again
net = vgg16.get_vgg_model()
# Load up a session which we'll use to import the graph into.
with tf.Session(graph=g) as sess, g.device('/cpu:0'):
# We can set the `net_input` to our content image
# or perhaps another image
# or an image of noise
# net_input = tf.Variable(content_img / 255.0)
net_input = tf.get_variable(
name='input',
shape=content_img.shape,
dtype=tf.float32,
initializer=tf.random_normal_initializer(
mean=np.mean(content_img), stddev=np.std(content_img)))
# Now we load the network again, but this time replacing our placeholder
# with the trainable tf.Variable
tf.import_graph_def(
net['graph_def'],
name='net',
input_map={'images:0': net_input})
```
<a name="content-loss"></a>
## Content Loss
In the lecture we saw that we'll simply find the l2 loss between our content layer features.
```
with tf.Session(graph=g) as sess, g.device('/cpu:0'):
content_loss = tf.nn.l2_loss((g.get_tensor_by_name(content_layer) -
content_features) /
content_features.size)
```
<a name="style-loss"></a>
## Style Loss
Instead of straight l2 loss on the raw feature activations, we're going to calculate the gram matrix and find the loss between these. Intuitively, this is finding what is common across all convolution filters, and trying to enforce the commonality between the synthesis and style image's gram matrix.
```
with tf.Session(graph=g) as sess, g.device('/cpu:0'):
style_loss = np.float32(0.0)
for style_layer_i, style_gram_i in zip(style_layers, style_features):
layer_i = g.get_tensor_by_name(style_layer_i)
layer_shape = layer_i.get_shape().as_list()
layer_size = layer_shape[1] * layer_shape[2] * layer_shape[3]
layer_flat = tf.reshape(layer_i, [-1, layer_shape[3]])
gram_matrix = tf.matmul(tf.transpose(layer_flat), layer_flat) / layer_size
style_loss = tf.add(style_loss, tf.nn.l2_loss((gram_matrix - style_gram_i) / np.float32(style_gram_i.size)))
```
<a name="total-variation-loss"></a>
## Total Variation Loss
And just like w/ guided hallucinations, we'll try to enforce some smoothness using a total variation loss.
```
def total_variation_loss(x):
h, w = x.get_shape().as_list()[1], x.get_shape().as_list()[1]
dx = tf.square(x[:, :h-1, :w-1, :] - x[:, :h-1, 1:, :])
dy = tf.square(x[:, :h-1, :w-1, :] - x[:, 1:, :w-1, :])
return tf.reduce_sum(tf.pow(dx + dy, 1.25))
with tf.Session(graph=g) as sess, g.device('/cpu:0'):
tv_loss = total_variation_loss(net_input)
```
<a name="training"></a>
## Training
We're almost ready to train! Let's just combine our three loss measures and stick it in an optimizer.
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
with tf.Session(graph=g) as sess, g.device('/cpu:0'):
# Experiment w/ the weighting of these! They produce WILDLY different
# results.
loss = 5.0 * content_loss + 1.0 * style_loss + 0.001 * tv_loss
optimizer = tf.train.AdamOptimizer(0.05).minimize(loss)
```
And now iterate! Feel free to play with the number of iterations or how often you save an image. If you use a different network to VGG, then you will not need to feed in the dropout parameters like I've done here.
```
imgs = []
n_iterations = 100
with tf.Session(graph=g) as sess, g.device('/cpu:0'):
sess.run(tf.global_variables_initializer())
# map input to noise
og_img = net_input.eval()
for it_i in range(n_iterations):
_, this_loss, synth = sess.run([optimizer, loss, net_input], feed_dict={
'net/dropout_1/random_uniform:0': np.ones(
g.get_tensor_by_name(
'net/dropout_1/random_uniform:0'
).get_shape().as_list()),
'net/dropout/random_uniform:0': np.ones(
g.get_tensor_by_name(
'net/dropout/random_uniform:0'
).get_shape().as_list())
})
print("%d: %f, (%f - %f)" %
(it_i, this_loss, np.min(synth), np.max(synth)))
if it_i % 5 == 0:
m = vgg16.deprocess(synth[0])
imgs.append(m)
plt.imshow(m)
plt.show()
gif.build_gif(imgs, saveto='stylenet.gif')
ipyd.Image(url='stylenet.gif?i=0', height=300, width=300)
```
<a name="assignment-submission"></a>
# Assignment Submission
After you've completed the notebook, create a zip file of the current directory using the code below. This code will make sure you have included this completed ipython notebook and the following files named exactly as:
<pre>
session-4/
session-4.ipynb
softmax.gif
fractal.gif
guided.gif
content.png
style.png
stylenet.gif
</pre>
You'll then submit this zip file for your third assignment on Kadenze for "Assignment 4: Deep Dream and Style Net"! Remember to complete the rest of the assignment, gallery commenting on your peers work, to receive full credit! If you have any questions, remember to reach out on the forums and connect with your peers or with me.
To get assessed, you'll need to be a premium student! This will allow you to build an online portfolio of all of your work and receive grades. If you aren't already enrolled as a student, register now at http://www.kadenze.com/ and join the [#CADL](https://twitter.com/hashtag/CADL) community to see what your peers are doing! https://www.kadenze.com/courses/creative-applications-of-deep-learning-with-tensorflow/info
Also, if you share any of the GIFs on Facebook/Twitter/Instagram/etc..., be sure to use the #CADL hashtag so that other students can find your work!
```
utils.build_submission('session-4.zip',
('softmax.gif',
'fractal.gif',
'guided.gif',
'content.png',
'style.png',
'stylenet.gif',
'session-4.ipynb'))
```
| github_jupyter |
```
%matplotlib inline
```
# NumPy memmap in joblib.Parallel
This example illustrates some features enabled by using a memory map
(:class:`numpy.memmap`) within :class:`joblib.Parallel`. First, we show that
dumping a huge data array ahead of passing it to :class:`joblib.Parallel`
speeds up computation. Then, we show the possibility to provide write access to
original data.
## Speed up processing of a large data array
We create a large data array for which the average is computed for several
slices.
```
import numpy as np
data = np.random.random((int(1e7),))
window_size = int(5e5)
slices = [slice(start, start + window_size)
for start in range(0, data.size - window_size, int(1e5))]
```
The ``slow_mean`` function introduces a :func:`time.sleep` call to simulate a
more expensive computation cost for which parallel computing is beneficial.
Parallel may not be beneficial for very fast operation, due to extra overhead
(workers creations, communication, etc.).
```
import time
def slow_mean(data, sl):
"""Simulate a time consuming processing."""
time.sleep(0.01)
return data[sl].mean()
```
First, we will evaluate the sequential computing on our problem.
```
tic = time.time()
results = [slow_mean(data, sl) for sl in slices]
toc = time.time()
print('\nElapsed time computing the average of couple of slices {:.2f} s'
.format(toc - tic))
```
:class:`joblib.Parallel` is used to compute in parallel the average of all
slices using 2 workers.
```
from joblib import Parallel, delayed
tic = time.time()
results = Parallel(n_jobs=2)(delayed(slow_mean)(data, sl) for sl in slices)
toc = time.time()
print('\nElapsed time computing the average of couple of slices {:.2f} s'
.format(toc - tic))
```
Parallel processing is already faster than the sequential processing. It is
also possible to remove a bit of overhead by dumping the ``data`` array to a
memmap and pass the memmap to :class:`joblib.Parallel`.
```
import os
from joblib import dump, load
folder = './joblib_memmap'
try:
os.mkdir(folder)
except FileExistsError:
pass
data_filename_memmap = os.path.join(folder, 'data_memmap')
dump(data, data_filename_memmap)
data = load(data_filename_memmap, mmap_mode='r')
tic = time.time()
results = Parallel(n_jobs=2)(delayed(slow_mean)(data, sl) for sl in slices)
toc = time.time()
print('\nElapsed time computing the average of couple of slices {:.2f} s\n'
.format(toc - tic))
```
Therefore, dumping large ``data`` array ahead of calling
:class:`joblib.Parallel` can speed up the processing by removing some
overhead.
## Writable memmap for shared memory :class:`joblib.Parallel`
``slow_mean_write_output`` will compute the mean for some given slices as in
the previous example. However, the resulting mean will be directly written on
the output array.
```
def slow_mean_write_output(data, sl, output, idx):
"""Simulate a time consuming processing."""
time.sleep(0.005)
res_ = data[sl].mean()
print("[Worker %d] Mean for slice %d is %f" % (os.getpid(), idx, res_))
output[idx] = res_
```
Prepare the folder where the memmap will be dumped.
```
output_filename_memmap = os.path.join(folder, 'output_memmap')
```
Pre-allocate a writable shared memory map as a container for the results of
the parallel computation.
```
output = np.memmap(output_filename_memmap, dtype=data.dtype,
shape=len(slices), mode='w+')
```
``data`` is replaced by its memory mapped version. Note that the buffer has
already been dumped in the previous section.
```
data = load(data_filename_memmap, mmap_mode='r')
```
Fork the worker processes to perform computation concurrently
```
Parallel(n_jobs=2)(delayed(slow_mean_write_output)(data, sl, output, idx)
for idx, sl in enumerate(slices))
```
Compare the results from the output buffer with the expected results
```
print("\nExpected means computed in the parent process:\n {}"
.format(np.array(results)))
print("\nActual means computed by the worker processes:\n {}"
.format(output))
```
## Clean-up the memmap
Remove the different memmap that we created. It might fail in Windows due
to file permissions.
```
import shutil
try:
shutil.rmtree(folder)
except: # noqa
print('Could not clean-up automatically.')
```
| github_jupyter |
# Homework 3 (solutions)
### Problem 1
Use read_html() function from the pandas library inside a for loop to get and print exchange rate data from rate.am for a week starting on June 1st and ending on 7th of June, 2017 included. The necessary steps to take are: 1) create the list of URLs to scrape, 2) create a for loop to iterate over the elements of that list and 3) receive the data from each of them and 4) print it. (Hint: you may try for 2 URLs first, and then go for 7).
```
import pandas as pd
urls = ["http://rate.am/am/armenian-dram-exchange-rates/banks/non-cash/2017/06/01/20-15",
"http://rate.am/am/armenian-dram-exchange-rates/banks/non-cash/2017/06/02/20-15",
"http://rate.am/am/armenian-dram-exchange-rates/banks/non-cash/2017/06/03/20-15",
"http://rate.am/am/armenian-dram-exchange-rates/banks/non-cash/2017/06/04/20-15",
"http://rate.am/am/armenian-dram-exchange-rates/banks/non-cash/2017/06/05/20-15",
"http://rate.am/am/armenian-dram-exchange-rates/banks/non-cash/2017/06/06/20-15",
"http://rate.am/am/armenian-dram-exchange-rates/banks/non-cash/2017/06/07/20-15"]
for url in urls:
data = pd.read_html(url)
print(data[2][2:19])
```
A note: instead of typing the 7 URLS above, one could just take the very first URL and then generate the others based on the first one, as the day digit is the only difference between URLs.
### Problem 2
Use regular expressions to match (find) and print the value of S&P500 index from the Bloomberg website ([click here](https://www.bloomberg.com/quote/SPX:IND), at the time of writing this the value of the index is 2,434.51). Your regular expression must match any value of S&P500 even if it changes (e.g. whether it becomes 1 or even 1,500,000.005, for example).
```
import re
import requests
url = "https://www.bloomberg.com/quote/SPX:IND"
response = requests.get(url)
data = response.text
output = re.findall('class\s*=\s*"price"\s*>\s*(\S+)</',data)
print(output)
```
### Problem 3
Create a for loop that will iterate over a given JSON file and print the keys followed by their values. The file (input string, which yet needs to be converted to JSON, as done in the classroom) and the expected output can be found in [this Jupyter notebook](http://nbviewer.jupyter.org/github/HrantDavtyan/Data_Scraping/blob/master/Week 3/HW3_P3_source.ipynb). NOTE: not more than one "if" and one "else" statements are allowed to use inside the for loop.
```
import json
input = '''[
{
"Movie":"Game of Thrones",
"Actor":"Peter Dinklage",
"Role":"Tyrion Lannister"
},
{
"Movie":"Vikings",
"Actor":"Travis Fimmel",
"Role":"Ragnar Lothbrok"
},
{
"Movie":"The last Kingdom",
"Actor":{
"Young Uhtred":"Tom Taylor",
"Not that young Uhtred":"Alexander Dreymon"
},
"Role":"Uhtred of Bebbanburg"
}
]'''
data = json.loads(input)
for i in data:
if len(i["Actor"])==2:
print "Movie: ",i["Movie"]
print "Role: ", i["Role"]
print "Actor 1: ",i["Actor"]["Young Uhtred"]
print "Actor 2: ",i["Actor"]["Not that young Uhtred"],"\n"
else:
print "Movie: ",i["Movie"]
print "Role: ", i["Role"]
print "Actor:",i["Actor"],"\n"
```
### Problem 4
Download the AirPassengers.csv file from the Datasets folder in the Moodle, read that file to python and plot the "Passengers" column.
```
import pandas as pd
import matplotlib.pyplot as plt
data = pd.read_csv("AirPassengers.csv")
plt.plot(data["Passengers"])
plt.show()
```
### Problem 5
Use regular expressions to find the URL (the hyperlink) under the Next button on [this webpage](http://quotes.toscrape.com/).
```
import re
import requests
url = "http://quotes.toscrape.com/"
response = requests.get(url)
data = response.text
output = re.findall('<a\s+href\s*=\s*"\s*(\S+)\s*"\s*\S*\s*>\s*Next',data)
print(output)
```
| github_jupyter |
```
import re
import pandas as pd
from numpy.random import RandomState
import spacy
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
import matplotlib.pyplot as plt
import json
## load data from json
with open('original_labelled_data.json', 'r') as jf:
data=json.load(jf)
sentences,labels=data['sentences'], data['labels']
import sklearn
from sklearn.model_selection import train_test_split
from collections import Counter
## split data into train and test
X_train, X_test, y_train, y_test = train_test_split(
sentences, labels, test_size=0.2, random_state=2020, stratify=labels)
# set up fields
def clean_str(string):
"""
Tokenization/string cleaning for all datasets except for SST.
Original taken from https://github.com/yoonkim/CNN_sentence/blob/master/process_data.py
"""
string = string.lower() ## lower-wise case
string = re.sub(r"[^A-Za-z0-9(),!?\'\`]", " ", string)
string = re.sub(r"\'s", " \'s", string)
string = re.sub(r"\'ve", " \'ve", string)
string = re.sub(r"n\'t", " n\'t", string)
string = re.sub(r"\'re", " \'re", string)
string = re.sub(r"\'d", " \'d", string)
string = re.sub(r"\'ll", " \'ll", string)
string = re.sub(r",", " , ", string)
string = re.sub(r"!", " ! ", string)
string = re.sub(r"\(", " \( ", string)
string = re.sub(r"\)", " \) ", string)
string = re.sub(r"\?", " \? ", string)
string = re.sub(r"\s{2,}", " ", string)
return string.strip()
def process_document(sentence):
"""
process a document
:params[in]: sentence, a string or list of strings
:params[out]: tokens, list of tokens
"""
if type(sentence)==str:
clean = clean_str(sentence) ## cleaned sentence
tokens = clean.split()
return tokens
elif type(sentence)==list:
res = []
for it in sentence:
res.append(clean_str(it).split())
return res
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
tk_sentences = process_document(sentences)
max_words = 10000
tk = Tokenizer(lower = True, num_words=max_words, filters='!"#$%&()*+,-./:;<=>?@[\\]^_`{|}~\t\n',
split=' ', char_level=False, oov_token='<unk>')
tk.fit_on_texts(tk_sentences)
print('Average word length of questions in train is {0:.0f}.'.format(np.mean(train['sentences'].apply(lambda x: len(clean_str(x).split())))))
print('Average word length of questions in test is {0:.0f}.'.format(np.std(train['sentences'].apply(lambda x: len(clean_str(x).split())))))
sentences[1]
import numpy as np
w2v_file='../glove.6B/glove.6B.100d.txt'
def loadData_Tokenizer(X_train, X_test, MAX_NB_WORDS=75000,
MAX_SEQUENCE_LENGTH=500, w2v_file=w2v_file):
"""
use glove embedding
"""
np.random.seed(7)
text = np.concatenate((X_train, X_test), axis=0)
text = np.array(text)
tokenizer = Tokenizer(num_words=MAX_NB_WORDS)
tokenizer.fit_on_texts(text)
sequences = tokenizer.texts_to_sequences(text)
word_index = tokenizer.word_index
text = pad_sequences(sequences, maxlen=MAX_SEQUENCE_LENGTH,padding='post', truncating='post')
print('Found %s unique tokens.' % len(word_index))
indices = np.arange(text.shape[0])
# np.random.shuffle(indices)
text = text[indices]
print(text.shape)
X_train = text[0:len(X_train), ]
X_test = text[len(X_train):, ]
embeddings_index = {}
f = open(w2v_file, encoding="utf8") ## GloVe file which could be download https://nlp.stanford.edu/projects/glove/
for line in f:
values = line.split()
word = values[0]
try:
coefs = np.asarray(values[1:], dtype='float32')
except:
pass
embeddings_index[word] = coefs
f.close()
print('Total %s word vectors.' % len(embeddings_index))
return (X_train, X_test, word_index,embeddings_index)
## construct the initialized embedding matrix based on pretrained matrix
def init_embed(word_index, embeddings_index, emb_dim=100):
embedding_matrix = np.random.random((len(word_index) + 1, emb_dim))
for word, i in word_index.items():
embedding_vector = embeddings_index.get(word)
if embedding_vector is not None:
# words not found in embedding index will be all-zeros.
if len(embedding_matrix[i]) !=len(embedding_vector):
print("could not broadcast input array from shape",str(len(embedding_matrix[i])),
"into shape",str(len(embedding_vector))," Please make sure your"
" EMBEDDING_DIM is equal to embedding_vector file ,GloVe,")
exit(1)
embedding_matrix[i] = embedding_vector
return embedding_matrix
train_ids,test_ids,wd_ind,emb_ind=loadData_Tokenizer(X_train, X_test, MAX_NB_WORDS=10000,
MAX_SEQUENCE_LENGTH=256, w2v_file=w2v_file)
embedding_matrix=init_embed(wd_ind, emb_ind, emb_dim=100)
embedding_matrix.shape[1]
10//2
```
### BiLSTM with Attention
```
class LSTMAttention(torch.nn.Module):
def __init__(self, label_size, hidden_dim, batch_size, embedding_matrix,
keep_dropout=.5, lstm_layers=1):
super(LSTMAttention, self).__init__()
self.hidden_dim = hidden_dim
self.batch_size = batch_size
self.use_gpu = torch.cuda.is_available()
self.embedding_dim = embedding_matrix.shape[1]
self.word_embeddings = nn.Embedding.from_pretrained(embedding_matrix, freeze=False)
self.num_layers = lstm_layers
self.dropout = keep_dropout
self.bilstm = nn.LSTM(self.embedding_dim, hidden_dim // 2, batch_first=True, num_layers=self.num_layers,
dropout=self.dropout, bidirectional=True)
self.hidden2label = nn.Linear(hidden_dim, label_size)
self.hidden = self.init_hidden()
##self.mean = opt.__dict__.get("lstm_mean",True) -- no use in the class
self.attn_fc = torch.nn.Linear(self.embedding_dim, 1)
def init_hidden(self, batch_size=None):
"""
initialize hidden state and cell state
"""
if batch_size is None:
batch_size= self.batch_size
if self.use_gpu:
h0 = Variable(torch.zeros(2*self.num_layers, batch_size, self.hidden_dim // 2).cuda())
c0 = Variable(torch.zeros(2*self.num_layers, batch_size, self.hidden_dim // 2).cuda())
else:
h0 = Variable(torch.zeros(2*self.num_layers, batch_size, self.hidden_dim // 2))
c0 = Variable(torch.zeros(2*self.num_layers, batch_size, self.hidden_dim // 2))
return (h0, c0)
def attention(self, rnn_out, state):
merged_state = torch.cat([s for s in state],1)
merged_state = merged_state.squeeze(0).unsqueeze(2)
# (batch, seq_len, cell_size) * (batch, cell_size, 1) = (batch, seq_len, 1)
weights = torch.bmm(rnn_out, merged_state)
weights = torch.nn.functional.softmax(weights.squeeze(2)).unsqueeze(2)
# (batch, cell_size, seq_len) * (batch, seq_len, 1) = (batch, cell_size, 1)
return torch.bmm(torch.transpose(rnn_out, 1, 2), weights).squeeze(2)
def forward(self, X):
embedded = self.word_embeddings(X)
hidden= self.init_hidden(X.size()[0])
rnn_out, hidden = self.bilstm(embedded, hidden)
h_n, c_n = hidden
attn_out = self.attention(rnn_out, h_n)
logits = self.hidden2label(attn_out)
return logits
from torch.utils.data import TensorDataset, DataLoader, RandomSampler, SequentialSampler
import time,datetime
from sklearn.metrics import classification_report
import random
from torch.optim.lr_scheduler import StepLR
import pdb
# Function to calculate the accuracy of our predictions vs labels
def flat_accuracy(preds, labels):
pred_flat = np.argmax(preds, axis=1).flatten()
labels_flat = labels.flatten()
return np.sum(pred_flat == labels_flat) / len(labels_flat)
def format_time(elapsed):
'''
Takes a time in seconds and returns a string hh:mm:ss
'''
# Round to the nearest second.
elapsed_rounded = int(round((elapsed)))
# Format as hh:mm:ss
return str(datetime.timedelta(seconds=elapsed_rounded))
# If there's a GPU available...
if torch.cuda.is_available():
# Tell PyTorch to use the GPU.
device = torch.device("cuda")
print('There are %d GPU(s) available.' % torch.cuda.device_count())
print('We will use the GPU:', torch.cuda.get_device_name(0))
# If not...
else:
print('No GPU available, using the CPU instead.')
device = torch.device("cpu")
### evaluate the performance of current model
def evaluate_model(clf_model, validation_dataloader, save_dir):
"""
:params[in]: clf_model, the pre-trained classifier
:params[in]: validation_dataloader, the validation dataset
:params[in]: save_dir, the directory name to save the fine-tuned model
"""
t0 = time.time()
# Put the model in evaluation mode--the dropout layers behave differently
# during evaluation.
clf_model.eval()
# Tracking variables
eval_loss, eval_accuracy = 0, 0
nb_eval_steps, nb_eval_examples = 0, 0
true_labels,pred_labels=[],[]
# Evaluate data for one epoch
for batch in validation_dataloader:
# Add batch to GPU
batch = tuple(t.to(device) for t in batch)
# Unpack the inputs from our dataloader
b_input_ids, b_labels = batch
# Telling the model not to compute or store gradients, saving memory and
# speeding up validation
with torch.no_grad():
# Forward pass, calculate logit predictions.
# This will return the logits rather than the loss because we have
# not provided labels.
# token_type_ids is the same as the "segment ids", which
# differentiates sentence 1 and 2 in 2-sentence tasks.
# The documentation for this `model` function is here:
# https://huggingface.co/transformers/v2.2.0/model_doc/bert.html#transformers.BertForSequenceClassification
outputs = clf_model(b_input_ids)
# Get the "logits" output by the model. The "logits" are the output
# values prior to applying an activation function like the softmax.
logits = outputs
# Move logits and labels to CPU
logits = logits.detach().cpu().numpy()
label_ids = b_labels.to('cpu').numpy()
# Calculate the accuracy for this batch of test sentences.
tmp_eval_accuracy = flat_accuracy(logits, label_ids)
## pred_labels/true_labels in a batch flatten
pred_flat = np.argmax(logits, axis=1).flatten()
true_flat = label_ids.flatten()
# true labels and predicted labels
true_labels += true_flat.tolist()
pred_labels += pred_flat.tolist()
# Accumulate the total accuracy.
eval_accuracy += tmp_eval_accuracy
# Track the number of batches
nb_eval_steps += 1
## pdb check
#pdb.set_trace()
# Report the final accuracy for this validation run
if not os.path.exists(save_dir):
os.makedirs(save_dir)
#clf_model.save_pretrained(save_dir) ## save model
print(classification_report(true_labels, pred_labels,digits=3))
print(classification_report(true_labels, pred_labels,digits=3),
file=open(save_dir+'/result.txt','w'))
print(" Accuracy: {0:.3f}".format(eval_accuracy/nb_eval_steps),
file=open(save_dir+'/result.txt','w'))
def train_eval(clf_model, train_dataloader, validation_dataloader, base_dir,
lr, epochs=4, eval_every_num_iters=40, seed_val = 42, weights= None):
"""train and evaluate a deep learning model
:params[in]: clf_model, a classifier
:params[in]: train_dataloader, training data
:params[in]: validation_dataloader, validation data
:params[in]: base_dir, output directory to create the directory to save results
:params[in]: lr, the learning rate
:params[in]: epochs, the number of training epochs
:params[in]: eval_every_num_iters, the number of iterations to evaluate
:params[in]: seed_val, set a random seed
"""
optimizer = torch.optim.Adam(clf_model.parameters(),
lr = lr)
## cross entropy loss
criterion = nn.CrossEntropyLoss()
# Total number of training steps is number of batches * number of epochs.
total_steps = len(train_dataloader) * epochs
# Create the learning rate scheduler. # gamma = decaying factor
scheduler = StepLR(optimizer, step_size=2, gamma=0.5)
# see if weights is None:
if weights != None:
weights = torch.FloatTensor(weights)
# Set the seed value all over the place to make this reproducible.
random.seed(seed_val)
np.random.seed(seed_val)
torch.manual_seed(seed_val)
torch.cuda.manual_seed_all(seed_val)
# Store the average loss after each epoch so we can plot them.
loss_values = []
# For each epoch...
for epoch_i in range(0, epochs):
# ========================================
# Training
# ========================================
print("")
print('======== Epoch {:} / {:} ========'.format(epoch_i + 1, epochs))
print('Training...')
# Measure how long the training epoch takes.
t0 = time.time()
# Reset the total loss for this epoch.
total_loss = 0
## print the learning rate
for param_group in optimizer.param_groups:
print(' learning rate is: ', param_group['lr'])
# Put the model into training mode. Don't be mislead--the call to
# `train` just changes the *mode*, it doesn't *perform* the training.
# `dropout` and `batchnorm` layers behave differently during training
# vs. test (source: https://stackoverflow.com/questions/51433378/what-does-model-train-do-in-pytorch)
# For each batch of training data...
for iters, batch in enumerate(train_dataloader):
clf_model.train() ## model training mode
# Unpack this training batch from our dataloader.
#
# As we unpack the batch, we'll also copy each tensor to the GPU using the
# `to` method.
#
# `batch` contains three pytorch tensors:
# [0]: input ids
# [1]: attention masks
# [2]: labels
b_input_ids = batch[0].to(device)
b_labels = batch[1].to(device)
# Always clear any previously calculated gradients before performing a
# backward pass. PyTorch doesn't do this automatically because
# accumulating the gradients is "convenient while training RNNs".
# (source: https://stackoverflow.com/questions/48001598/why-do-we-need-to-call-zero-grad-in-pytorch)
clf_model.zero_grad()
# Perform a forward pass (evaluate the model on this training batch).
# This will return the loss (rather than the model output) because we
# have provided the `labels`.
# The documentation for this `model` function is here:
# https://huggingface.co/transformers/v2.2.0/model_doc/bert.html#transformers.BertForSequenceClassification
outputs = clf_model(b_input_ids)
# The call to `model` always returns a tuple, so we need to pull the
# loss value out of the tuple.
loss = criterion(outputs, b_labels)
# Accumulate the training loss over all of the batches so that we can
# calculate the average loss at the end. `loss` is a Tensor containing a
# single value; the `.item()` function just returns the Python value
# from the tensor.
total_loss += loss.item()
# Perform a backward pass to calculate the gradients.
loss.backward()
# Clip the norm of the gradients to 1.0.
# This is to help prevent the "exploding gradients" problem.
# torch.nn.utils.clip_grad_norm_(clf_model.parameters(), 1.0)
# Update parameters and take a step using the computed gradient.
# The optimizer dictates the "update rule"--how the parameters are
# modified based on their gradients, the learning rate, etc.
optimizer.step()
# eveluate the performance after some iterations
if iters % eval_every_num_iters == 0 and not iters == 0:
# Calculate elapsed time in minutes.
elapsed = format_time(time.time() - t0)
# Report progress.
print(' Batch {:>5,} of {:>5,}. Elapsed: {:}.'.format(iters, len(train_dataloader), elapsed))
tmp_dir = base_dir+'/epoch'+str(epoch_i+1)+'iteration'+str(iters)
## save pretrained model
evaluate_model(clf_model, validation_dataloader, tmp_dir)
# Update the learning rate each epoch
scheduler.step()
# Calculate the average loss over the training data.
avg_train_loss = total_loss / len(train_dataloader)
#pdb.set_trace()
# Store the loss value for plotting the learning curve.
loss_values.append(avg_train_loss)
# save the data after epochs
tmp_dir = base_dir+'/epoch'+str(epoch_i+1)+'_done'
## save pretrained model
evaluate_model(clf_model, validation_dataloader, tmp_dir)
from torch.autograd import Variable
import os
for seed in [42,52, 62, 72, 82]:
batch_size = 8
train_ids,test_ids,wd_ind,emb_ind=loadData_Tokenizer(X_train, X_test, MAX_NB_WORDS=10000,
MAX_SEQUENCE_LENGTH=256, w2v_file=w2v_file)
embedding_matrix=init_embed(wd_ind, emb_ind, emb_dim=100)
## initialize a classifier
clf_model=LSTMAttention(label_size=3, hidden_dim=60, batch_size=batch_size, lstm_layers=1,
keep_dropout=.5, embedding_matrix=torch.FloatTensor(embedding_matrix))
# Create the DataLoader for our training set.
train_data = TensorDataset(torch.LongTensor(train_ids), torch.tensor(y_train))
train_sampler = RandomSampler(train_data)
train_dataloader = DataLoader(train_data, sampler=train_sampler, batch_size=batch_size)
# Create the DataLoader for our validation set.
validation_data = TensorDataset(torch.LongTensor(test_ids), torch.tensor(y_test))
validation_sampler = SequentialSampler(validation_data)
validation_dataloader = DataLoader(validation_data, sampler=validation_sampler,
batch_size=batch_size)
base_dir = 'textLSTM_att/LSTM_att_seed'+str(seed)
train_eval(clf_model, train_dataloader, validation_dataloader, base_dir, \
lr=1.0e-2, epochs=5, eval_every_num_iters=160, seed_val = seed)
##text cnn
res = np.array([[0.732,0.731,0.734,0.858],
[0.761,0.762,0.761,0.875],
[0.773,0.776,0.771,0.881],
[0.779,0.772,0.789,0.872],
[0.760,0.749,0.790,0.847]
])
np.mean(res, axis=0)
np.std(res, axis=0)
##BERT
res = np.array([[0.760,0.764,0.757,0.881],
[0.790,0.787,0.794,0.889],
[0.730,0.734,0.727,0.872],
[0.763,0.762,0.765,0.881]
])
np.mean(res, axis=0)
np.std(res, axis=0)
```
| github_jupyter |
# Gaussian Process
In this tutorial, we expose what gaussian processes are, and how to use the [GPy library](http://sheffieldml.github.io/GPy/). We first provide a gentle reminder about Gaussian distributions and their properties.
```
# Import all the important libraries
import math
import numpy as np
import matplotlib.pyplot as plt
from scipy.stats import norm
from scipy.stats import multivariate_normal
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.mlab as mlab
from ipywidgets import widgets as wg
from matplotlib import cm
import GPy
#%matplotlib inline
%matplotlib notebook
```
## 1D Gaussian distribution
```
# Plot
def plot_gaussian(mu=0, sigma=1):
x = np.linspace(-3, 3, 100)
plt.plot(x, norm.pdf(x, mu, sigma))
plt.xlabel('x')
plt.ylabel('p(x)')
wg.interact(plot_gaussian, mu=(-2,2,0.1), sigma=(-2,2,0.1))
plt.show()
```
## Multivariate Gaussian distribution (2D)
The multivariable Gaussian distribution is a generalization of the Gaussian distribution to vectors. See [wikipedia](https://en.wikipedia.org/wiki/Multivariate_normal_distribution) for more info.
```
# moments
mu = np.array([0,0])
Sigma = np.array([[1,0],
[0,1]])
Sigma1 = np.array([[1,0.5],
[0.5,1]])
Sigma2 = np.array([[1,-0.5],
[-0.5,1]])
Sigmas = [Sigma, Sigma1, Sigma2]
pts = []
for S in Sigmas:
pts.append(np.random.multivariate_normal(mu, S, 1000).T)
# Plotting
width = 16
height = 4
plt.figure(figsize=(width, height))
# make plot
for i in range(len(Sigmas)):
plt.subplot(1,3,i+1)
plt.title('Plot '+str(i+1))
plt.ylim(-4,4)
plt.xlim(-4,4)
plt.xlabel('x1')
plt.ylabel('x2')
plt.plot(pts[i][0], pts[i][1],'o')
#plt.scatter(pts[i][0], pts[i][1])
plt.show()
```
The 1st plot above is described by:
\begin{equation}
\left[ \begin{array}{c} x_1\\x_2 \end{array}\right] \sim \mathcal{N} \left(\left[ \begin{array}{c} 0\\0 \end{array}\right], \left[ \begin{array}{cc} 1 & 0\\ 0 & 1 \end{array}\right] \right)
\end{equation}
The 2nd plot is given by:
\begin{equation}
\left[ \begin{array}{c} x_1\\x_2 \end{array}\right] \sim \mathcal{N}\left(\left[ \begin{array}{c} 0\\0 \end{array}\right], \left[ \begin{array}{cc} 1 & 0.5\\ 0.5 & 1 \end{array}\right]\right)
\end{equation}
Finally, the 3rd plot is given by:
\begin{equation}
\left[ \begin{array}{c} x_1\\x_2 \end{array}\right] \sim \mathcal{N}\left(\left[ \begin{array}{c} 0\\0 \end{array}\right], \left[ \begin{array}{cc} 1 & -0.5\\ -0.5 & 1 \end{array}\right]\right)
\end{equation}
The covariance (and the dot product) measures the similarity.
For the 2nd and 3rd plots, $x_1$ is **correlated** with $x_2$, i.e. knowing $x_1$ gives us information about $x_2$.
### Joint distribution $p(x_1,x_2)$
The joint distribution $p(x_1, x_2)$ is given by:
\begin{equation}
\left[ \begin{array}{c} x_1\\x_2 \end{array}\right] \sim \mathcal{N}\left(\left[ \begin{array}{c} \mu_1 \\ \mu_2 \end{array}\right], \left[ \begin{array}{cc} \Sigma_{11} & \Sigma_{12} \\ \Sigma_{21} & \Sigma_{22} \end{array}\right] \right) = \mathcal{N}(\pmb{\mu}, \pmb{\Sigma})
\end{equation}
```
# Reference: http://stackoverflow.com/questions/38698277/plot-normal-distribution-in-3d
# moments
mu = np.array([0,0])
Sigma = np.array([[1,0], [0,1]])
# Create grid and multivariate normal
step = 500
bound = 10
x = np.linspace(-bound,bound,step)
y = np.linspace(-bound,bound,step)
X, Y = np.meshgrid(x,y)
pos = np.empty(X.shape + (2,))
pos[:, :, 0] = X; pos[:, :, 1] = Y
pdf = multivariate_normal(mu, Sigma).pdf(pos)
# Plot
fig = plt.figure(figsize=plt.figaspect(0.5)) # Twice as wide as it is tall.
# 1st subplot (3D)
ax = fig.add_subplot(1, 2, 1, projection='3d')
ax.plot_surface(X, Y, pdf, cmap='viridis', linewidth=0)
ax.set_xlabel('x1')
ax.set_ylabel('x2')
ax.set_zlabel('p(x1, x2)')
# 2nd subplot (2D)
ax = fig.add_subplot(1, 2, 2)
ax.contourf(x, y, pdf)
#ax.colorbar()
ax.set_xlabel('x1')
ax.set_ylabel('x2')
fig.tight_layout()
plt.show()
```
### Normalization
In order to be a valid probability distribution, the volume under the surface should equal to 1.
\begin{equation}
\int \int p(x_1,x_2) dx_1 dx_2 = 1
\end{equation}
```
# p(x1,x2) = pdf
# dx = 2.*bound/step
# dx1 dx2 = (2.*bound/step)**2
print("Summation: {}".format((2.*bound/step)**2 * pdf.sum()))
```
### Conditional distribution $p(x_2|x_1)$
What is the mean $\mu_{2|1}$ and the variance $\Sigma_{2|1}$ of the conditional distribution $p(x_2|x_1) = \mathcal{N}(\mu_{2|1}, \Sigma_{2|1})$?
We know the mean $\pmb{\mu}$ and the covariance $\pmb{\Sigma}$ of the joint distribution $p(x_1,x_2)$. Using the [Schur complement](https://en.wikipedia.org/wiki/Schur_complement), we obtain:
\begin{align}
\mu_{2|1} &= \mu_{2} + \Sigma_{21}\Sigma_{22}^{-1}(x_2 - \mu_2) \\
\Sigma_{2|1} &= \Sigma_{22} - \Sigma_{21}\Sigma_{22}^{-1}\Sigma_{12}
\end{align}
For the demo, check Murphy's book "Machine Learning: A Probabilistic Perspective", section 4.3.4
```
fig = plt.figure(figsize=plt.figaspect(0.5)) # Twice as wide as it is tall.
x1_value = 0
z_max = pdf.max()
# 1st subplot
ax = fig.add_subplot(1, 2, 1, projection='3d')
ax.plot_surface(X, Y, pdf, cmap='viridis', linewidth=0)
ax.set_xlabel('x1')
ax.set_ylabel('x2')
ax.set_zlabel('p(x1, x2)')
y1 = np.linspace(-bound,bound,2)
z = np.linspace(0,z_max,2)
Y1, Z = np.meshgrid(y1,z)
ax.plot_surface(x1_value, Y1, Z, color='red', alpha=0.2)
#cset = ax.contourf(X, Y, pdf, zdir='x', offset=-bound, cmap=cm.coolwarm)
# 2nd subplot
ax = fig.add_subplot(1, 2, 2)
ax.plot(x, pdf[step//2 + x1_value*step//(2*bound)])
ax.set_xlabel('x2')
ax.set_ylabel('p(x2|x1)')
fig.tight_layout()
plt.show()
```
### Marginal distribution $p(x_1)$ and $p(x_2)$
\begin{align}
p(x_1) &= \int p(x_1, x_2) dx_2 = \mathcal{N}(\mu_1, \Sigma_{11}) \\
p(x_2) &= \int p(x_1, x_2) dx_1 = \mathcal{N}(\mu_2, \Sigma_{22})
\end{align}
```
fig = plt.figure(figsize=plt.figaspect(0.5))
plt.subplot(1,2,1)
plt.title('By summing')
dx = 2. * bound / step
plt.plot(x, pdf.sum(0) * dx, color='blue')
plt.xlabel('x2')
plt.ylabel('p(x2)')
plt.subplot(1,2,2)
plt.title('by using the normal distribution')
plt.plot(x, norm.pdf(x, mu[1], Sigma[1,1]), color='red')
plt.xlabel('x2')
plt.ylabel('p(x2)')
fig.tight_layout()
plt.show()
```
## Gaussian Processes (GPs)
A Gaussian process is a Gaussian distribution over functions. That is, it is a generalization of the multivariable Gaussian distribution to infinite vectors.
It will become clearer with an example.
```
x = [0.5,0.8,1.4]
f = [1,2,6]
plt.plot(x,f,'o')
for i in range(len(x)):
plt.annotate('f'+str(i+1), (x[i],f[i]))
plt.xlim(0,2)
plt.ylim(0,6.5)
plt.ylabel('f(x)')
plt.xlabel('x')
plt.xticks(x, ['x'+str(i+1) for i in range(len(x))])
plt.show()
```
\begin{align}
\left[ \begin{array}{c} f_1 \\ f_2 \\ f_3 \end{array}\right]
&\sim \mathcal{N}\left( \left[ \begin{array}{c} 0 \\ 0 \\ 0 \end{array}\right], \left[ \begin{array}{ccc} K_{11} & K_{12} & K_{13} \\ K_{21} & K_{22} & K_{23} \\ K_{31} & K_{32} & K_{33} \end{array}\right] \right) \\
&\sim \mathcal{N}\left( \left[ \begin{array}{c} 0 \\ 0 \\ 0 \end{array}\right], \left[ \begin{array}{ccc} 1 & 0.7 & 0.2 \\ 0.7 & 1 & 0.6 \\ 0.2 & 0.6 & 1 \end{array}\right] \right)
\end{align}
Similarity measure: $K_{ij} = \exp(- ||x_i - x_j||^2) = \left\{ \begin{array}{ll} 0 & ||x_i - x_j|| \rightarrow \infty \\ 1 & x_i = x_j \end{array} \right.$
Prediction (noiseless GP regression): given data $\mathcal{D} = \{(x_1,f_1), (x_2,f_2), (x_3,f_3)\}$, and new point $x_*$ (e.g. $x_*$=1.4), what is the value of $f_*$?
\begin{equation}
\pmb{f} \sim \mathcal{N}(\pmb{0}, \pmb{K}) \qquad \mbox{and} \qquad f_* \sim \mathcal{N}(0, K(x_*,x_*)) = \mathcal{N}(0, K_{**})
\end{equation}
In this case, $K_{**} = K(x_*,x_*) = \exp(- ||x_* - x_*||^2) = 1$.
Now, we can write:
\begin{equation}
\left[ \begin{array}{c} \pmb{f} \\ f_* \end{array} \right] \sim \mathcal{N}\left(\pmb{0}, \left[\begin{array}{cc} \left[ \begin{array}{ccc} K_{11} & K_{12} & K_{13} \\ K_{21} & K_{22} & K_{23} \\ K_{31} & K_{32} & K_{33} \end{array} \right] & \left[ \begin{array}{c} K_{1*} \\ K_{2*} \\ K_{3*} \end{array} \right] \\ \left[ \begin{array}{ccc} K_{*1} & K_{*2} & K_{*3} \end{array} \right] & \left[\begin{array}{c} K_{**} \end{array} \right] \end{array} \right] \right) = \mathcal{N}\left(\pmb{0}, \left[\begin{array}{cc} \pmb{K} & \pmb{K}_* \\ \pmb{K}_*^T & \pmb{K}_{**} \end{array}\right]\right)
\end{equation}
Using the formula for the conditional probability $p(f_*|f)$, we have:
\begin{align}
\mu_* &= \mathbb{E}[f_*] = \pmb{K}_*^T \pmb{K}^{-1}\pmb{f} \\
c_* &= K_{**} - \pmb{K}_*^T \pmb{K}^{-1}\pmb{K}_*
\end{align}
We can thus predict the mean $\mu_*$ and the variance $c_*$ for the test point $x_*$.
```
x = [0.5,0.8,1.4]
f = [1,2,6]
x_new = 1.3
f_new = 5.2
plt.plot(x+[x_new],f+[f_new],'o')
for i in range(len(x)):
plt.annotate('f'+str(i+1), (x[i],f[i]))
plt.errorbar(x_new, f_new, yerr=1)
plt.annotate('f*', (x_new+0.02, f_new))
plt.xlim(0,2)
plt.ylim(0,6.5)
plt.ylabel('f(x)')
plt.xlabel('x')
plt.xticks(x+[x_new], ['x'+str(i+1) for i in range(len(x))]+['x*'])
plt.show()
```
### Generalization
A GP defines a distribution over functions $p(f)$ (i.e. it is the joint distribution over all the infinite function values).
Definition: $p(f)$ is a GP if for any finite subset $\{x_1,...,x_n\} ⊂ X$, the marginal distribution over that finite subset $p(f)$ has a multivariate Gaussian distribution.
Prior on $f$:
\begin{equation}
\pmb{f}|\pmb{x} \sim \mathcal{GP}(\pmb{\mu}(\pmb{x}), \pmb{K}(\pmb{x}, \pmb{x}))
\end{equation}
with
\begin{align*}
\pmb{\mu}(\pmb{x}) &= \mathbb{E}_f \lbrack \pmb{x} \rbrack \\
k(\pmb{x}, \pmb{x'}) &= \mathbb{E}_f \lbrack (\pmb{x} - \pmb{\mu}(\pmb{x})) (\pmb{x'} - \pmb{\mu}(\pmb{x'})) \rbrack
\end{align*}
Often written as:
\begin{equation}
\pmb{f} \sim \mathcal{GP}(\pmb{0}, \pmb{K})
\end{equation}
Concretely, assume $\pmb{x} \in \mathbb{R}^{50}$, then $\pmb{K}(\pmb{x}, \pmb{x}) \in \mathbb{R}^{50 \times 50}$, then $\pmb{f} \sim \mathcal{GP}(\pmb{0}, \pmb{K})$ means:
\begin{equation}
\left[ \begin{array}{c} f_1 \\ \vdots \\ f_{50} \end{array}\right] := \left[ \begin{array}{c} f(x_1) \\ \vdots \\ f(x_{50}) \end{array}\right] \sim \mathcal{N}\left( \left[ \begin{array}{c} 0 \\ \vdots \\ 0 \end{array}\right], \left[ \begin{array}{ccc} k(x_1,x_1) & \cdots & k(x_1, x_{50}) \\ \vdots & \ddots & \vdots \\ k(x_{50},x_1) & \cdots & k(x_{50}, x_{50}) \end{array} \right] \right)
\end{equation}
#### RBF kernel
Let's choose a RBF (a.k.a Squared Exponential, Gaussian) kernel:
\begin{equation}
\pmb{K} = \left[ \begin{array}{ccc} k(x_1,x_1) & \cdots & k(x_1, x_d) \\ \vdots & \ddots & \vdots \\ k(x_d,x_1) & \cdots & k(x_d, x_d) \end{array} \right]
\end{equation}
with
\begin{equation}
k(x_i, x_j) = \alpha^2 \exp \left( - \frac{(x_i - x_j)^2}{2l} \right) \qquad \mbox{ and hyperparameters } \pmb{\Phi} = \left\{ \begin{array}{l} \alpha \mbox{: amplitude} \\ l \mbox{: the lengthscale} \end{array} \right.
\end{equation}
This function $k$ is infinitely differentiable.
```
# Reference: https://www.youtube.com/watch?v=4vGiHC35j9s&t=51s
# Hyperparameters
alpha = 1
l = 2
# Parameters
n = 50 # nb of points
n_func = 10 # nb of fct to draw
x_bound = 5 # bound on the x axis
def RBF_kernel(a,b):
sqdist = np.sum(a**2,1).reshape(-1,1) + np.sum(b**2,1) - 2*np.dot(a,b.T)
return alpha**2 * np.exp(-1/l * sqdist)
n = 50
X = np.linspace(-x_bound, x_bound, n).reshape(-1,1)
K = RBF_kernel(X, X) # dim(K) = n x n
L = np.linalg.cholesky(K + 1e-6 * np.eye(n))
f_prior = np.dot(L, np.random.normal(size=(n, n_func)))
# Plotting
width = 16
height = 4
plt.figure(figsize=(width, height))
# plot f_prior
plt.subplot(1,3,1)
plt.title('GP: prior on f')
plt.plot(X, f_prior)
plt.plot(X, f_prior.mean(1), linewidth=3, color='black')
plt.ylabel('f(x)')
plt.xlabel('x')
# plot Kernel
plt.subplot(1,3,2)
plt.title('Kernel matrix')
plt.pcolor(K[::-1])
plt.colorbar()
plt.subplot(1,3,3)
plt.title('Kernel function')
plt.plot(X, RBF_kernel(X, np.array([[1.0]])))
plt.show()
```
### Kernel (prior knowledge)
By choosing a specific kernel, we can incorporate prior knowledge that we have about the function $f$, such as, if the function is:
* periodic
* smooth
* symmetric
* etc.
The hyperparameters for each kernel are also very intuitive/interpretable.
Note: kernels can be combined!
Indeed, if $k(x,y)$, $k_1(x,y)$ and $k_2(x,y)$ are valid kernels then:
* $\alpha k(x,y) $ with $\alpha \geq 0$
* $k_1(x,y) + k_2(x,y)$
* $k_1(x,y) k_2(x,y)$
* $p(k(x,y))$ with $p$ being a polynomial function with non-negative coefficients
* $exp(k(x,y))$
* $f(x) k(x,y) \overline{f(y)}$ with $\overline{f} = $ complex conjugate
* $k(\phi(x),\phi(y))$
are all valid kernels!
##### Periodic Exponential kernel
```
variance = 1.
lengthscale = 1.
period = 2.*np.pi
#K = periodic_kernel(X, X) # dim(K) = n x n
kern = GPy.kern.PeriodicExponential(variance=variance, lengthscale=lengthscale, period=period)
K1 = kern.K(X)
L = np.linalg.cholesky(K1 + 1e-6 * np.eye(n))
f_prior = np.dot(L, np.random.normal(size=(n, 1)))
# Plotting
width = 16
height = 4
plt.figure(figsize=(width, height))
# plot f_prior
plt.subplot(1,3,1)
plt.title('GP: prior on f')
plt.plot(X, f_prior)
plt.plot(X, f_prior.mean(1), linewidth=3, color='black')
plt.ylabel('f(x)')
plt.xlabel('x')
# plot Kernel
plt.subplot(1,3,2)
plt.title('Kernel matrix')
plt.pcolor(K1[::-1])
plt.colorbar()
plt.subplot(1,3,3)
plt.title('Kernel function')
plt.plot(X, kern.K(X, np.array([[1.0]])))
plt.show()
```
##### addition and multiplication of 2 kernels (SE and PE)
```
K_add = K + K1
L = np.linalg.cholesky(K_add + 1e-6 * np.eye(n))
f_prior = np.dot(L, np.random.normal(size=(n, n_func)))
# Plotting
width = 16
height = 8
plt.figure(figsize=(width, height))
# plot f_prior
plt.subplot(2,2,1)
plt.title('GP: prior on f with K_add')
plt.plot(X, f_prior)
plt.plot(X, f_prior.mean(1), linewidth=3, color='black')
plt.ylabel('f(x)')
plt.xlabel('x')
# plot Kernel
plt.subplot(2,2,2)
plt.title('Kernel matrix: K_add')
plt.pcolor(K_add[::-1])
plt.colorbar()
K_prod = K * K1
L = np.linalg.cholesky(K_prod + 1e-6 * np.eye(n))
f_prior = np.dot(L, np.random.normal(size=(n, n_func)))
# plot f_prior
plt.subplot(2,2,3)
plt.title('GP: prior on f with K_prod')
plt.plot(X, f_prior)
plt.plot(X, f_prior.mean(1), linewidth=3, color='black')
plt.ylabel('f(x)')
plt.xlabel('x')
# plot Kernel
plt.subplot(2,2,4)
plt.title('Kernel matrix: K_prod')
plt.pcolor(K_prod[::-1])
plt.colorbar()
plt.show()
```
### GP Posterior
Given $\mathcal{D}=\{(x_i, y_i)\}_{i=1}^{i=N} = (\pmb{X}, \pmb{y})$, we have:
\begin{equation}
p(f|\mathcal{D}) = \frac{p(\mathcal{D}|f)p(f)}{p(\mathcal{D})}
\end{equation}
### GP Regression
\begin{equation}
y_i = f(\pmb{x}_i) + \epsilon_i \qquad
\left\{ \begin{array}{l}
f \sim \mathcal{GP}(\pmb{0}, \pmb{K}) \\
\epsilon_i \sim \mathcal{N}(0, \sigma^2)
\end{array} \right.
\end{equation}
* Prior $f$ is a GP $\Leftrightarrow p(\pmb{f}|\pmb{X}) = \mathcal{N}(\pmb{0}, \pmb{K})$
* Likelihood is Gaussian $\Leftrightarrow p(\pmb{y}|\pmb{X},\pmb{f}) = \mathcal{N}(\pmb{f}, \sigma^2\pmb{I})$
* $\rightarrow p(f|\mathcal{D})$ is also a GP.
#### Predictive distribution:
$$p(\pmb{y}_*|\pmb{x}_*, \pmb{X}, \pmb{y}) = \int p(\pmb{y}_{*}| \pmb{x}_{*}, \pmb{f}, \pmb{X}, \pmb{y}) p(f|\pmb{X}, \pmb{y}) d\pmb{f} = \mathcal{N}(\pmb{\mu}_*, \pmb{\Sigma}_*)$$
\begin{align}
\pmb{\mu}_* &= \pmb{K}_{*N} (\pmb{K}_N + \sigma^2 \pmb{I})^{-1} \pmb{y} \\
\pmb{\Sigma}_* &= \pmb{K}_{**} - \pmb{K}_{*N} (\pmb{K}_N + \sigma^2 \pmb{I})^{-1} \pmb{K}_{N*}
\end{align}
### Learning a GP
#### Marginal likelihood:
\begin{equation}
p(\pmb{y}|\pmb{X}) = \int p(\pmb{y}|\pmb{f},\pmb{X}) p(\pmb{f}|\pmb{X}) d\pmb{f} = \mathcal{N}(\pmb{0}, \pmb{K} + \sigma^2\pmb{I})
\end{equation}
By taking the logarithm, and setting $\pmb{K}_y = (\pmb{K} + \sigma^2\pmb{I})$, we have:
\begin{equation}
\mathcal{L} = \log p(\pmb{y}|\pmb{X}; \pmb{\Phi}) = \underbrace{-\frac{1}{2} \pmb{y}^T \pmb{K}_y^{-1} \pmb{y}}_{\mbox{data fit}} \underbrace{-\frac{1}{2} \log |\pmb{K}_y^{-1}|}_{\mbox{complexity penalty}} - \frac{n}{2} \log 2\pi
\end{equation}
The marginal likelihood (i.e. ML-II) is used to optimize the hyperparameters $\pmb{\Phi}$ that defines the covariance function and thus the GP.
\begin{equation}
\pmb{\Phi}^* = argmax_{\pmb{\Phi}} \log p(\pmb{y}|\pmb{X}; \pmb{\Phi})
\end{equation}
Optimizing the marginal likelihood is more robust than the likelihood as it tries to optimize the complexity of the model, and the fitting of this last one to the observed data.
### GPy
```
# GP Regression
# Based on the tutorial: https://github.com/SheffieldML/notebook/blob/master/GPy/GPyCrashCourse.ipynb
# Create dataset
X = np.random.uniform(-3.0, 3.0, (20,1))
Y = np.sin(X) + np.random.randn(20,1) * 0.05
# Create the kernel
# Reminder 1: The sum of valid kernels gives a valid kernel.
# Reminder 2: The product of valid kernels gives a valid kernel.
# Available kernels: RBF, Exponential, Matern32, Matern52, Brownian, Bias, Linear, PeriodicExponential, White.
kernel = GPy.kern.RBF(input_dim=1, variance=1.0, lengthscale=1.0)
# Create the model
gp_model = GPy.models.GPRegression(X, Y, kernel)
# Display and plot
print("Before optimization: ", gp_model)
gp_model.plot()
plt.show()
# Optimize the model (that is find the 'best' hyperparameters of the kernel matrix)
# By default, the optimizer is a 2nd order algo: lbfgsb. Others are available such as the scg, ...
gp_model.optimize(messages=False)
# Display and plot
print("After optimization: ", gp_model)
gp_model.plot()
plt.show()
```
### Gaussian Process Latent Variable Model (GP-LVM)
```
# GPLVM
# Based on the tutorials:
# http://nbviewer.jupyter.org/github/SheffieldML/notebook/blob/master/GPy/MagnificationFactor.ipynb
# https://github.com/SheffieldML/notebook/blob/master/lab_classes/gprs/lab4-Copy0.ipynb
# Create dataset
N = 100
k1 = GPy.kern.RBF(5, variance=1, lengthscale=1./np.random.dirichlet(np.r_[10,10,10,0.1,0.1]), ARD=True)
k2 = GPy.kern.RBF(5, variance=1, lengthscale=1./np.random.dirichlet(np.r_[0.1,10,10,10,0.1]), ARD=True)
X = np.random.normal(0, 1, (N,5))
A = np.random.multivariate_normal(np.zeros(N), k1.K(X), 10).T
B = np.random.multivariate_normal(np.zeros(N), k2.K(X), 10).T
Y = np.vstack((A,B))
# latent space dimension
latent_dim = 2
# Create the kernel
kernel = GPy.kern.RBF(input_dim=latent_dim, variance=1.0, lengthscale=1.0)
# Create the GPLVM model
gplvm_model = GPy.models.GPLVM(Y, latent_dim, init='PCA', kernel=kernel)
# Display and plot
print("Before optimization: ", gplvm_model)
gplvm_model.plot_latent()
plt.show()
# Optimize the model (that is find the 'best' hyperparameters of the kernel matrix)
# By default, the optimizer is a 2nd order algo: lbfgsb. Others are available such as the scg, ...
gplvm_model.optimize(messages=False)
# Display and plot
print("After optimization: ", gplvm_model)
gplvm_model.plot_latent()
plt.show()
```
| github_jupyter |
```
__author__ = 'Knut Olsen <knut.olsen@noirlab.edu>' # single string; emails in <>
__version__ = '20211121' # yyyymmdd; version datestamp of this notebook
__datasets__ = ['phat_v2'] # enter used datasets by hand
__keywords__ = ['M31','Healpix map','plot:cmd','lightcurve']
```
# Fun with PHAT!
*Knut Olsen & the Astro Data Lab Team*
## Table of contents
* [Goals & Summary](#goals)
* [Disclaimer & attribution](#attribution)
* [Imports & setup](#import)
* [Authentication](#auth)
* [Basic info](#basic)
* [Examine the columns of the PHAT object table](#columns)
* [Make an object density map](#density)
* [Make depth and color maps](#depth)
* [Do a spatial query and make color-color diagrams and CMDs](#cmd)
* [Retrieving time series photometry of select objects](#timeseries)
* [Resources and references](#resources)
<a class="anchor" id="goals"></a>
# Goals
* Learn how to access the PHAT average photometry and single epoch measurement tables
* Learn how to use an SQL query to make Healpix maps of object density, average depth, average color, and PHAT brick number
* Learn how to retrieve and plot lightcurve data from the PHAT single epoch measurement table
# Summary
The Panchromatic Hubble Andromeda Treasury (PHAT; PI Dalcanton) was a Hubble Space Telescope Multi-cycle program to map roughly a third of M31's star forming disk, using 6 filters covering from the ultraviolet through the near infrared.
The Data Lab hosts two main tables from PHAT: the version 2 object table (phat_v2.phot_mod) and the version 2 single epoch measurement table (phat_v2.phot_meas). The object table contains average photometry in all filters, with one row per object. The measurement table contains all of the photometric measurements for all objects, with one row per measurement. The measurement table contains ~7.5 billion rows.
In this notebook, we'll use these tables to do some exploration of the PHAT survey.
<a class="anchor" id="attribution"></a>
# Disclaimer & attribution
If you use this notebook for your published science, please acknowledge the following:
* Data Lab concept paper: Fitzpatrick et al., "The NOAO Data Laboratory: a conceptual overview", SPIE, 9149, 2014, http://dx.doi.org/10.1117/12.2057445
* Data Lab disclaimer: http://datalab.noirlab.edu/disclaimers.php
* PHAT Reduction paper: Williams et al., "Reducing and Analyzing the PHAT Survey with the Cloud", ApJS, 2018, 236, 4
<a class="anchor" id="import"></a>
# Imports and setup
```
import numpy as np
import pylab as plt
import matplotlib
import healpy as hp
from getpass import getpass
%matplotlib inline
# Datalab and related imports
from dl import authClient as ac, queryClient as qc
from dl.helpers.utils import convert
```
<a class="anchor" id="auth"></a>
# Authentication
If this is the first time logging in to Data Lab, you can make use of virtual storage and myDB by logging in with your credentials here. You only need to do this once, as your login is kept current as an active user unless you logout or login as a different user.
```
# Authenticated users please uncomment the next line
#token = ac.login(input("Enter user name: "),getpass("Enter password: "))
```
<a class="anchor" id="basic"></a>
# Basic info
First, let's look at the tables available in the PHAT database, and then get some basic information from the Data Lab statistics database (<tt>tbl_stat</tt>) about the main PHAT object table, phat_v2.phot_mod.
```
try:
print(qc.schema('phat_v2',format='json',profile='default'))
except Exception as e:
print(e.message)
%%time
query = "SELECT * FROM tbl_stat WHERE schema='phat_v2' and tbl_name='phot_mod'" # Retrieve useful stats, quickly
try:
info = qc.query(sql=query) # by default the result is a CSV formatted string
except Exception as e:
print(e.message)
print(info)
```
<a class="anchor" id="columns"></a>
# Examine the columns of the PHAT object table
First, we'll take a look at some rows from phat_v2.phot_mod, and get all columns.
```
query = """SELECT *
FROM phat_v2.phot_mod
LIMIT 100
"""
%%time
try:
result = qc.query(sql=query) # by default the result is a CSV formatted string
except Exception as e:
print(e.message)
```
### Convert the output to a Pandas Dataframe
Pandas dataframes are a convenient way to store and work with the data. The Data Lab 'helpers' module (<a href="http://datalab.noirlab.edu/docs/manual/UsingAstroDataLab/ClientInterfaces/Helpers/Helpers.html">docs</a>) has a conversion method, with many possible output formats.
```
df1 = convert(result,'pandas')
print("Number of rows:", len(df1))
print(df1.columns) # print column headings
print(len(df1.columns))
```
<a class="anchor" id="density"></a>
# Make an object density map
One of the columns in the PHAT object table, pix4096, is the Healpix index (NSIDE=4096, nested scheme) for the objects's RA and Dec. Healpix is a handy tesselation of the sky into tiles of equal area. The Python module healpy has all of the Healpix related functions.
To make maps of aggregate quantities in PHAT, we're going to use the database to return results in a query grouped by Healpix index value. We can then put the results into arrays, and use healpy's functionality to display the maps.
In this first query, the GROUP BY clause tells the database to aggregate the results by the values in the pix4096 column, and return the average RA and Dec of objects in those groups, as well as the pix4096 value itself and the count of the number of objects in the group.
```
query = """SELECT avg(ra) as ra0,avg(dec) as dec0,pix4096,count(pix4096) as nb
FROM phat_v2.phot_mod
GROUP BY pix4096
"""
%%time
# This query should take about a minute
try:
result = qc.query(sql=query) # by default the result is a CSV formatted string
except Exception as e:
print(e.message)
```
### Convert the output to a Pandas Dataframe
We'll once again use helpers to convert the result to a Pandas dataframe. Our dataframe has one row per Healpix. We'll check that the sum of the number of objects in all Healpixels equals the number of rows in the table.
```
df_density = convert(result,'pandas')
print("Number of rows:", len(df_density))
print(np.sum(df_density['nb'])) # print total counts
```
### Making the Healpix map
A Healpix map is simply a one-dimensional array with number of elements set by the NSIDE parameter, which is the number of times the base Healpixels are split. We can visualize it as a map using the <tt>healpy</tt> library.
```
hmap = np.zeros(hp.nside2npix(4096))
print(df_density.head())
```
### Populating the Healpix map
Now we set the elements of our Healpix map to the number of objects returned by the query, calculate the center of the RA and Dec distribution of the objects, and use healpy's gnomview to visualize the output. Notice anything funny? The PHAT object table has duplicate objects between some of the bricks (bricks 11, 12, 13, and 14).
```
hmap[df_density['pix4096']]=df_density['nb']
(rarot,decrot)=(np.median(df_density['ra0']),np.median(df_density['dec0']))
hp.gnomview(hmap,title='',notext=True,cbar=False,reso=0.4,nest=True,rot=(rarot,decrot,0),cmap='inferno',min=1e3,max=8e4)
```
<a class="anchor" id="depth"></a>
# Make depth and color maps
Now we'll get a little fancier with our maps. We'll have the database return average WFC3 u-band and ACS magnitudes and colors and the PHAT brick numbers, and make some cuts on the magnitudes and flags of the objects. We again GROUP BY the pix4096 column.
```
query = """SELECT avg(ra) as ra0,avg(dec) as dec0,pix4096,count(pix4096) as nb,
avg(f475w_vega) as gmag,avg(f814w_vega) as imag,avg(brick) as brick,
avg(f475w_vega-f814w_vega) as g_i
FROM phat_v2.phot_mod
WHERE f475w_gst=1 AND f814w_gst=1 AND f475w_vega<50 AND f814w_vega<50
GROUP BY pix4096
"""
print(query)
%%time
# Query will take a few seconds
try:
result = qc.query(sql=query) # by default the result is a CSV formatted string
except Exception as e:
print(e.message)
df_all = convert(result,'pandas')
print("Number of rows:", len(df_all))
print(np.sum(df_all['nb'])) # print total counts
print(df_all.head())
```
### Healpix map of average F475W magnitude
The map of the average F475W magnitude gives a good idea of how the PHAT catalog depth varies with position in M31. The depth is much shallower in the Bulge, which is very crowded, than in the outer disk.
```
gmap = np.zeros(hp.nside2npix(4096))
gmap[df_all['pix4096']] = df_all['gmag']
hp.gnomview(gmap,title='',notext=True,cbar=False,reso=0.4,nest=True,rot=(rarot,decrot,0),cmap='inferno',min=25,max=28)
```
### Healpix map of average F475W-F814W color
The map of average color reveals both population differences and the dust lanes in the galaxy, as well as the 10 kpc ring.
```
gimap = np.zeros(hp.nside2npix(4096))
gimap[df_all['pix4096']] = df_all['g_i']
hp.gnomview(gimap,title='',notext=True,cbar=False,reso=0.4,nest=True,rot=(rarot,decrot,0),cmap='inferno',min=1,max=2.5)
```
### Healpix map of the brick number
We can also use our Healpix table to make a map of the PHAT bricks.
```
brickmap = np.zeros(hp.nside2npix(4096))
brickmap[df_all['pix4096']] = df_all['brick']
hp.gnomview(brickmap,reso=0.4,nest=True,rot=(rarot,decrot,0),cmap='jet',min=0,max=23)
```
<a class="anchor" id="cmd"></a>
# Do a spatial query and make color-color diagrams and CMDs
Now let's do a cone search for objects within a radius of a particular position. The PHAT tables are spatially indexed to make such queries fast. We'll search within a 1 arcmin radius of the RA and Dec position that we defined earlier.
```
query = """SELECT *
FROM phat_v2.phot_mod WHERE q3c_radial_query(ra,dec,{0},{1},{2})
""".format(rarot,decrot,1./60)
print(query)
%%time
try:
result = qc.query(sql=query) # by default the result is a CSV formatted string
except Exception as e:
print(e.message)
```
### Cut out bad missing values
Some of the objects returned will have missing magnitude measurements, indicated by 99's. Let's cut those out, and also select only the "good" stars in the ACS bands.
```
df = convert(result,'pandas')
print("Number of rows:", len(df))
df_cmd = df[(df['f336w_vega']<50) & (df['f475w_vega']<50) & (df['f814w_vega']<50) & \
(df['f475w_gst']==1) & (df['f814w_gst']==1)]
print("Number of rows:", len(df_cmd))
```
### Make color-color and CMD plots
We'll show the F475W-F814W,F336W-F475W color-color diagram, and a color-magnitude diagram. What do you notice?
```
# make a figure
fig = plt.figure(figsize=(20,10))
ax1 = fig.add_subplot(121)
ax2 = fig.add_subplot(122)
# color-magnitude diagram (Hess diagram)
im = ax1.hexbin(df_cmd['f475w_vega']-df_cmd['f814w_vega'],df_cmd['f336w_vega']-df_cmd['f475w_vega'], \
gridsize=400,cmap=matplotlib.cm.viridis,norm=matplotlib.colors.LogNorm())
ax1.set_ylabel('F336W - F475W',fontsize=15)
ax1.set_xlabel('F475W - F814W',fontsize=15)
ax1.set_title('Color-color diagram',fontsize=20)
ax1.set_ylim(-2,6)
ax1.set_xlim(-1,5)
# color-magnitude diagram (Hess diagram)
im2 = ax2.hexbin(df_cmd['f475w_vega']-df_cmd['f814w_vega'],df_cmd['f475w_vega'], \
gridsize=200,cmap=matplotlib.cm.viridis,norm=matplotlib.colors.LogNorm())
ax2.set_xlabel('F475W - F814W',fontsize=15)
ax2.set_ylabel('F475W',fontsize=15)
ax2.set_title('Color-magnitude (Hess) diagram',fontsize=20)
ax2.set_xlim(-1,5)
ax2.set_ylim(28.4,22)
```
<a class="anchor" id="timeseries"></a>
# Retrieving time series photometry of select objects
In this part of the notebook, we'll pick a Cepheid variable, and then query the full measurement table for the time series photometry. We'll define a couple of functions to retrieve data and make a plot.
```
# define a function to select only measurements in one band
def get_data(df,band='F475W'):
sel = (df['filter'] == band) & (df['magvega'] < 90)
t = df['mjd'][sel].values
y = df['magvega'][sel].values
dy = df['magerr'][sel].values
return t,y,dy # return time, magnitudes in one band, uncertainties
# a reusable function to plot the lightcurve
def plot_raw_lightcurve(t,y,dy,title='',ax=None,lperc=13,rperc=99,color='g',ms=8):
if ax is None:
fig, ax = plt.subplots()
jd0 = t.min() # modified Julian date offset
t = t-jd0 # first date of observations = 0
ax.errorbar(t,y,yerr=dy,marker='.',ms=ms,ls='none',color=color,lw=1,alpha=0.5,label='')
# Main panel chores
ax.set_xlabel('modified Julian date - %g (days)' % jd0, fontsize=20)
ax.set_ylabel('magnitude',fontsize=20)
ax.tick_params(labelsize=20)
ax.invert_yaxis()
ax.set_title(title)
def get_folded_phase(t,best_period):
"""Fold the observation times with the best period of the variable signal."""
# light curve over period, take the remainder (i.e. the "phase" of one period)
phase = (t / best_period) % 1
return phase
def plot_folded_lightcurve(t,y,best_period,dy=None,ax=None,color='g',ms=10):
"""Plot folded lightcurve.
Parameters
----------
t, y : array
Time and magnitude 1-d arrays
best_period : float
True period of the signal.
dy : array or None
If array, the values are the uncertainies on ``y``, and the plot will show errorbars.
If None, the plot will have no errorbars.
ax : instance or None
If instance of axis class, will plot to that object. If None, will generate a new figure and axis object.
"""
phase = get_folded_phase(t,best_period)
if ax is None:
fig, ax = plt.subplots()
marker = '.'
lw = 1
alpha = 0.6
if dy is not None:
ax.errorbar(phase,y,yerr=dy,marker=marker,ms=ms,ls='none',lw=lw,color=color,alpha=alpha)
else:
ax.plot(phase,y,marker=marker,ms=ms,ls='none',lw=lw,color=color,alpha=alpha)
ax.invert_yaxis()
ax.tick_params(labelsize=20)
ax.set_xlabel('phase (days)', fontsize=20)
ax.set_ylabel('magnitude',fontsize=20);
# Pick a star from Wagner-Kaiser et al. (2015)
rawk = 11.02203
decwk = 41.23451
best_period = 10.29 # days
query = """SELECT * FROM phat_v2.phot_meas WHERE q3c_radial_query(ra,dec,{0},{1},{2})""".format(rawk,decwk,0.2/3600)
print(query)
%%time
try:
result = qc.query(sql=query) # by default the result is a CSV formatted string
except Exception as e:
print(e.message)
df_ts = convert(result,'pandas')
print("Number of rows:", len(df_ts))
print(df_ts['filter'].unique())
print(df_ts['objid'].unique())
uid=df_ts['objid'].unique()
df_ts=df_ts[(df_ts['objid']==uid[0])]
colors = list('gcbrkm')
filters = ['F475W','F275W','F336W','F814W','F160W','F110W']
fig, (ax1,ax2) = plt.subplots(1,2,figsize=(20,10))
ax2.set_title('Folded light curve',fontsize=20)
for j in range(len(colors)):
c = colors[j]
t,y,dy = get_data(df_ts,filters[j])
plot_raw_lightcurve(t,y,dy,title='',ax=ax1,color=c,ms=20)
plot_folded_lightcurve(t,y,best_period,dy=dy,ax=ax2,color=c,ms=20)
```
<a class="anchor" id="resources"></a>
# Resources and references
Dalcanton, J.J. et al. (2012, ApJS, 200, 18), "The Panchromatic Hubble Andromeda Treasury"
http://adsabs.harvard.edu/abs/2012ApJS..200...18D
Wagner-Kaiser, R. et al. (2015, MNRAS, 451, 724), "Panchromatic Hubble Andromeda Treasury XIII: The Cepheid period-luminosity relation in M31"
http://adsabs.harvard.edu/abs/2015MNRAS.451..724W
Williams, B.F. et al. (2014, ApJS, 215, 9), "The Panchromatic Hubble Andromeda Treasury. X. Ultraviolet to Infrared Photometry of 117 Million Equidistant Stars"
http://adsabs.harvard.edu/abs/2014ApJS..215....9W
Williams, B.F. et al., "Reducing and Analyzing the PHAT Survey with the Cloud", ApJS, 236, 4
| github_jupyter |
# Exploring the FORCE 2020 Well Log Challenge - Part 1
## Visualizing the well logs with matplotlib
**Brendon Hall, Enthought**
bhall@enthought.com
It's time for another facies classification challenge! [FORCE](https://www.npd.no/en/force/) and [XEEK](https://xeek.ai/) have teamed up to bring us the [2020 FORCE Machine Learning Contest](https://xeek.ai/challenges/force-well-logs/overview). Peter Borman and his colleagues have curated a set of 100+ well logs from the North Sea. In addition to the log curves, each well has interpreted lithofacies. This is an awesome addition to the open data that is available for building tools for digital geoscience. Machine learning based facies classification is a popular subject, and this is an exciting new benchmark dataset.
Since the [SEG Contest](https://github.com/seg/2016-ml-contest) in 2016, I've been impressed [how much innovation](https://www.linkedin.com/pulse/developments-machine-deep-learning-facies-brendon-hall) can come from an open dataset and a community interested in building new tools and techniques. This time should be even better! The dataset is 10X the size and there are 4 years of new ideas to incorporate. I thought it would be useful to create a few Jupyter notebooks that provide some ideas for getting started with this dataset. If you're new to Python and programming, this will help get you started exploring the data. If you are more experienced, perhaps this will enable you to focus more on trying out different features or machine learning models. In any case, you are welcome to use any or all of this code as you see fit. If you have any suggestions or ideas for improvement, those are welcome as well.
This notebook will propose a colorbar for visualizing the lithofacies interpretations, and show how to use matplotlib to create a flexible log and lithofacies plot. This is based on the log plot in my [facies classification tutorial](https://github.com/seg/2016-ml-contest/blob/master/Facies_classification.ipynb). If you are brand new to Python and 'geocomputing', check out this [tutorial](https://www.youtube.com/watch?v=iIOMiN8Cacs&list=PLgLft9vxdduD8Zydz4dRJqIzCWDlPKITC&index=2&t=1241s) by Rob Leckenby to get started.
Please get in touch if you have any questions! You can also join in the conversation on [Software Underground's slack](https://softwareunderground.org/slack) in the **#force_2020_ml_contest** channel.
The well log data is licensed as [Norwegian License for Open Government Data (NLOD) 2.0](https://data.norge.no/nlod/en/2.0/).
The well log labels that are included are provided by FORCE 2020 Machine Learning Contest under [CC-BY-4.0](https://creativecommons.org/licenses/by/4.0/).
```
%matplotlib inline
import os.path
import numpy as np
import pandas as pd
import matplotlib.colors as colors
import matplotlib.pyplot as plt
from mpl_toolkits.axes_grid1 import make_axes_locatable
import ipywidgets as widgets
pd.options.display.max_rows = 8
```
Let's start by loading up the training data.
You can get the dataset on the [contest website](https://xeek.ai/challenges/force-well-logs/data). We'll only use the training dataset for this notebook. If you have already downloaded the data, change the `local_train_csv` variable below to point to the file.
If you don't have the data already, you can run the code below and download the data automatically from AWS-S3. A local copy will be saved at the location specified by `local_train_csv`, so you won't have to download it every time.
Note, we're also making an effort to make the seismic
data for the [fault competition](https://xeek.ai/challenges/force-seismic/overview) available
from S3, but through a simplified API. See this [repo](https://github.com/blasscoc/easy-as.git)
if you're also interested in the fault mapping competition.
```
# change this to the location of the training data on your disk if
# you have already downloaded it
local_train_csv = 'train.csv'
if not os.path.isfile(local_train_csv):
# load from s3
s3_train_csv = 's3://zarr-depot/wells/FORCE: Machine Predicted Lithology/train.csv'
wells_df = pd.read_csv(s3_train_csv, sep=';')
wells_df.to_csv(local_train_csv, index=False)
else:
# load from disk
wells_df = pd.read_csv(local_train_csv)
```
We'll need a list of all of the well names in the dataset.
```
wells_df
well_names = wells_df['WELL'].unique()
```
The 12 facies labels in the csv have integer codes. Let's add a couple of columns to make these codes easier for both humans and machines to understand. We'll add a more descriptive label for the lithology and also an integer value (numbered 0-11) that will be used for visualization (and later, labels for supervised machine learning)
```
# map of lithology codes to description
lithology_keys = {30000: 'Sandstone',
65030: 'Sandstone/Shale',
65000: 'Shale',
80000: 'Marl',
74000: 'Dolomite',
70000: 'Limestone',
70032: 'Chalk',
88000: 'Halite',
86000: 'Anhydrite',
99000: 'Tuff',
90000: 'Coal',
93000: 'Basement'}
# map of lithology codes to integer label for ML
lithology_numbers = {30000: 0,
65030: 1,
65000: 2,
80000: 3,
74000: 4,
70000: 5,
70032: 6,
88000: 7,
86000: 8,
99000: 9,
90000: 10,
93000: 11}
def map_lith_key(lith_map, row):
lith_key = row['FORCE_2020_LITHOFACIES_LITHOLOGY']
if lith_key in lith_map:
return lith_map[lith_key]
else:
print('Warning: Key {} not found in map'.format(lith_key))
return np.nan
wells_df['LITHOLOGY'] = wells_df.apply (lambda row: map_lith_key(lithology_keys, row), axis=1)
wells_df['LITH_LABEL'] = wells_df.apply (lambda row: map_lith_key(lithology_numbers, row), axis=1)
wells_df[['FORCE_2020_LITHOFACIES_LITHOLOGY', 'LITHOLOGY', 'LITH_LABEL']]
```
Now let's set up a color map to make visualizing the facies with the well logs easier. This will also be useful when comparing machine learning results. This color scheme is loosely based on the one I [used before](). Sandstone is yellow, Shale is green, and Sandstone-shale is somewhere in between. Carbonates are shades of blues, salts are shades of violet. [Tuff](https://www.sciencedirect.com/topics/earth-and-planetary-sciences/tuff) is lava red (of course), coal is black and basement is orange (because why not).
If you would like to choose other colors, here's a color picker widget that can help. Click the arrows at the bottom of the color picker tool to change to hex notation.
```
color_picker = widgets.ColorPicker(
concise=False,
description='Pick a color',
value='blue',
disabled=False
)
display(color_picker)
facies_color_map = { 'Sandstone': '#F4D03F',
'Sandstone/Shale': '#7ccc19',
'Shale': '#196F3D',
'Marl': '#160599',
'Dolomite': '#2756c4',
'Limestone': '#3891f0',
'Chalk': '#80d4ff',
'Halite': '#87039e',
'Anhydrite': '#ec90fc',
'Tuff': '#FF4500',
'Coal': '#000000',
'Basement': '#DC7633'}
# get a list of the color codes.
facies_colors = [facies_color_map[mykey] for mykey in facies_color_map.keys()]
```
Now define a function that creates a plot with log curves, as well as a lithofacies track. This plot will be flexible, in that we can provide a list of curve mnemonics that we want to include in the plot.
```
def make_facies_log_plot(log_df, curves, facies_colors):
#make sure logs are sorted by depth
logs = log_df.sort_values(by='DEPTH_MD')
cmap_facies = colors.ListedColormap(
facies_colors[0:len(facies_colors)], 'indexed')
ztop=logs.DEPTH_MD.min(); zbot=logs.DEPTH_MD.max()
cluster=np.repeat(np.expand_dims(logs['LITH_LABEL'].values,1), 100, 1)
num_curves = len(curves)
f, ax = plt.subplots(nrows=1, ncols=num_curves+1, figsize=(num_curves*2, 12))
for ic, col in enumerate(curves):
# if the curve doesn't exist, make it zeros
if np.all(np.isnan(logs[col])):
curve = np.empty(logs[col].values.shape)
curve[:] = np.nan
else:
curve = logs[col]
ax[ic].plot(curve, logs['DEPTH_MD'])
ax[ic].set_xlabel(col)
ax[ic].set_yticklabels([]);
# make the lithfacies column
im=ax[num_curves].imshow(cluster, interpolation='none', aspect='auto',
cmap=cmap_facies,vmin=0,vmax=11)
divider = make_axes_locatable(ax[num_curves])
cax = divider.append_axes("right", size="20%", pad=0.05)
cbar=plt.colorbar(im, cax=cax)
cbar.set_label((12*' ').join([' SS', 'SS-Sh', 'Sh',
' Ml', 'Dm', 'LS', 'Chk ',
' Hl', 'Ann', 'Tuf', 'Coal', 'Bsmt']))
cbar.set_ticks(range(0,1)); cbar.set_ticklabels('')
for i in range(len(ax)-1):
ax[i].set_ylim(ztop,zbot)
ax[i].invert_yaxis()
ax[i].grid()
ax[i].locator_params(axis='x', nbins=3)
ax[num_curves].set_xlabel('Facies')
ax[num_curves].set_yticklabels([])
ax[num_curves].set_xticklabels([])
f.suptitle('Well: %s'%logs.iloc[0]['WELL'], fontsize=14,y=0.94)
plt.show()
```
Let's try it out for sample well. Include the standard 'triple combo' logs.
```
make_facies_log_plot(
wells_df[wells_df['WELL'] == '15/9-13'],
['GR', 'RHOB', 'NPHI', 'RDEP'],
facies_colors)
```
Now let's try it for all of the curves. First, define a list with all of the curve names.
```
columns = ['CALI', 'RSHA', 'RMED', 'RDEP', 'RHOB', 'GR', 'SGR',
'NPHI', 'PEF', 'DTC', 'SP', 'BS', 'ROP', 'DTS', 'DCAL', 'DRHO',
'MUDWEIGHT', 'RMIC', 'ROPA', 'RXO']
make_facies_log_plot(
wells_df[wells_df['WELL'] == '15/9-13'],
columns,
facies_colors)
```
It's not very easy to read with this many curves, but it's quick and flexible and should be useful when trying new machine learning ideas.
It would be cool if the plot was easier to customize. Let's try the `ipywidgets` library, which gives us some controls to change the parameters of the plot. Here, we'll create a dropdown that let's us change the well being displayed. We'll use a multi-select box to indicate which curves to include in the plot (all are selected by default). Hold shift, control (or command) while clicking to select multiple curve names. Click the button to generate the plot based on your selection.
```
plot_output = widgets.Output()
dropdown_well = widgets.Dropdown(options = wells_df['WELL'].unique(), description='Well:', value='15/9-13')
select_curves = widgets.SelectMultiple(
options=columns,
value=columns,
#rows=10,
description='Curves:',
disabled=False
)
plot_button = widgets.Button(
description='Show well plot',
disabled=False,
button_style='', # 'success', 'info', 'warning', 'danger' or ''
tooltip='Click to plot well.'
)
def on_button_clicked(b):
plot_output.clear_output()
with plot_output:
make_facies_log_plot(
wells_df[wells_df['WELL'] == dropdown_well.value],
select_curves.value,
facies_colors)
plot_button.on_click(on_button_clicked)
display(dropdown_well)
display(select_curves)
display(plot_button)
display(plot_output)
```
I hope that this gives you some helpful code for plotting well curves and lithology in the FORCE 2020 dataset. Please feel free to use any or all of this code for your own work. There are plenty of ways to visualize well logs in Python. Ashley Russell has a great [tutorial](https://youtu.be/ud4EZiNpblM) on how to build interactive well logs plots in Jupyter.
This notebook is open source content. Text is CC-BY-4.0, code is [Apache 2.0](https://www.apache.org/licenses/LICENSE-2.0).
### References
Bormann P., Aursand P., Dilib F., Dischington P., Manral S. (2020) 2020 FORCE Machine Learning Contest. https://github.com/bolgebrygg/Force-2020-Machine-Learning-competition
| github_jupyter |
Compares the average file size of images photographed of the coast versus the average size of images photographed more inland. Using Hurrican Florence post-storm imagery. Size unit is in bytes until the values are scaled down using log()
```
import pandas as pd
import scipy.stats as stats
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
import numpy as np
import math
ct =pd.read_csv("catalog.csv")
```
Drop all columns except file and size
```
ct.drop(ct.columns[0], axis=1, inplace= True)
ct.drop(ct.columns[3:], axis=1, inplace=True)
ct.head()
```
Remove all storms but Florence
```
ct = ct.drop(ct.index[29097:])
ct.shape[0]
```
Select samples of coastline and inland photos. (location associated with date)
```
data = ct[ct.date != ('2018/09/16', '2018/09/18', '2018/09/19', '2018/09/20')]
data.head()
data.shape[0]
```
Demarcate coastal and inland images
```
cd = ct.loc[ct['date'] == '2018/09/17']
ind1 = ct.loc[ct['date'] == '2018/09/21']
ind2 = ct.loc[ct['date'] == '2018/09/22']
cd_size = cd.shape[0]
ind = ind1.append(ind2)
ind_size = ind.shape[0]
```
Two dataframes: cd (coastal data) and ind(inland data). Get the basic stats of each.
```
ind["log_value"] =np.log(ind["size"])
cd["log_value"] =np.log(cd["size"])
print("Inland mean:", ind["log_value"].mean())
print("Inland var:", ind["log_value"].var())
print("Inland std:", ind["log_value"].std())
print("Coastal mean:", cd["log_value"].mean())
print("Coastal var:", cd["log_value"].var())
print("Coastal std:", cd["log_value"].std())
cd_mean = cd["log_value"].mean()
cd_var = cd["log_value"].var()
```
Preparing variables for classroom method for fitting gamma distribution (didn't work due to scaling)
```
alpha_mom = cd_mean ** 2 / cd_var
beta_mom = cd_var / cd_mean
cd_min = cd["log_value"].min()
cd_max = cd["log_value"].max()
ind_min = ind["log_value"].min()
ind_max = ind["log_value"].max()
```
Plot histograms.
```
binwidth = (cd["log_value"].max()-cd["log_value"].min())/20
cd["log_value"].hist(bins=np.arange(cd_min, cd_max + binwidth, binwidth), normed = True)
cd_plot = cd["log_value"].hist(bins=np.arange(min(cd["log_value"]),max(cd["log_value"]) + binwidth, binwidth))
plt.xlabel("Image Size")
plt.ylabel( "Number of Images")
plt.title("Coastal Images, n = " + str(cd_size) + str())
#plt.plot(np.linspace(cd_min, cd_max), gamma.pdf(np.linspace(cd_min, cd_max), alpha_mom, beta_mom))
plt.savefig('coastal_images_hist.png', dpi = 100)
plt.show()
lnspc = np.linspace(cd_min, cd_max, len(cd['log_value']))
ag,bg,cg = stats.gamma.fit(cd["log_value"])
pdf_gamma = stats.gamma.pdf(lnspc, ag, bg,cg)
plt.xlabel("Image Size")
plt.ylabel( "Number of Images")
plt.title("Coastal Images, n = " + str(cd_size) + str())
plt.savefig('coastal_images_gamma.png', dpi = 100)
plt.plot(lnspc, pdf_gamma, label="Gamma")
lnspc = np.linspace(ind_min, ind_max, len(ind['log_value']))
ag,bg,cg = stats.gamma.fit(ind["log_value"])
binwidth = (ind["log_value"].max()-ind["log_value"].min())/20
ind["log_value"].hist(bins=np.arange(min(ind["log_value"]),max(ind["log_value"]) + binwidth, binwidth))
plt.xlabel("Image Size")
plt.ylabel( "Number of Images")
plt.title("Inland Images n = " + str(ind_size) + str())
plt.savefig('inland_images_hist.png', dpi = 100)
plt.show()
ag,bg,cg = stats.gamma.fit(ind["log_value"])
pdf_gamma = stats.gamma.pdf(lnspc, ag, bg,cg)
plt.xlabel("Image Size")
plt.ylabel( "Number of Images")
plt.title("Inland Images, n = " + str(ind_size) + str())
plt.savefig('inland_images_gamma.png', dpi = 100)
plt.plot(lnspc, pdf_gamma, label="Gamma")
```
one tailed z- test
```
cd_std = cd["log_value"].std()
sample_size = 1000
sample = np.random.choice(a= cd["log_value"], size = sample_size)
sample_mean = sample.mean()
z_critical = stats.norm.ppf(q = 0.975) # Get the z-critical value*
print("z-critical value:") # Check the z-critical value
print(z_critical)
# Get the population standard deviation
margin_of_error = z_critical * (cd_std /math.sqrt(cd_size))
confidence_interval = (sample_mean - margin_of_error,
sample_mean + margin_of_error)
print("Sample Mean:")
print(sample_mean)
print("Confidence interval:")
print(confidence_interval)
```
two-tailed z-test
```
sample_size = 1000
intervals = []
sample_means = []
for sample in range(25):
sample = np.random.choice(a= cd["log_value"], size = sample_size)
sample_mean = sample.mean()
sample_means.append(sample_mean)
z_critical = stats.norm.ppf(q = 0.975) # Get the z-critical value*
stats.norm.ppf(q = 0.025)
margin_of_error = z_critical * (cd_std/math.sqrt(sample_size))
confidence_interval = (sample_mean - margin_of_error,
sample_mean + margin_of_error)
intervals.append(confidence_interval)
print("Sample Mean:")
print(sample_mean)
print("Confidence interval:")
print(confidence_interval)
```
Looking at the first 1000 values for the inland and coastal image sizes and calculating correlation coefficent. There is almost none.
```
x = ind["log_value"]
y = cd["log_value"]
x = x[0:1000]
y = y[0:1000]
plt.scatter(x,y)
plt.xlabel('Inland')
plt.ylabel('Coastal')
plt.title('Image size correlation')
plt.savefig('size_correlation')
plt.show()
#np.concatenate((x,y[:,None]),axis=1)
print('Covariance of X and Y: %.2f'%np.cov(x, y)[0, 1])
print('Correlation of X and Y: %.2f'%np.corrcoef(x, y)[0, 1])
```
Compute the population mean for all storm image size on record. Null hypothesis: The sample of coastal image size is similar to the population image size of 60,000 images.
```
ct2 =pd.read_csv("catalog.csv")
ct2['log_value'] = np.log(ct2['size'])
ct2_val = ct2['log_value']
ct2_mean = np.mean(ct2_val[np.isfinite(ct2_val)])
stats.ttest_1samp(cd['log_value'], ct2_mean)
d = cd['log_value'].shape[0]*2 - 2
p = 1 - stats.t.cdf(-18.6236,df=d)
p
```
P value = 1. We fail to reject the null hypothesis.
| github_jupyter |
```
import numpy as np
import os
import sys
import matplotlib.pyplot as plt
import tensorflow as tf
from jupyterthemes import jtplot
from IPython.core.debugger import Tracer
jtplot.style()
#CLI args [data_set_idx, z_dim, time_steps]
#Example: python vrae data_set_idx=0 z_dim=20 time_steps=50
# X \in {0,1}^{batch_size, dim_x, time_steps}
#############################
# Constant params
#############################
x_in_dim = 88
# Params replicated from Fabius et. al Paper
beta_1 = 0.05
beta_2 = 0.001
num_epochs = 10#40000 #TEST
learning_rate_1 = 2e-5
learning_rate_2 = 1e-5
num_epochs_to_diff_learn_rate = 5#16000 #TEST
num_epochs_to_save_model = 4#1000 #TEST
num_hidden_units = 500
#decay_rate = .7
##############################
# Hyperparams
##############################
data_set_idx = 0
z_dim = 20
time_steps = 50
batch_size = 100
epoch_to_load = 8 # If greater than 0 load model from that epoch
run_to_load = 1
# Dir paths
data_sets = ['Nottingham','JSB Chorales']
load_root = '../MIDI_Data_PianoRolls/'
log_root = '../logs/MIDI_Data_PianoRolls'
train_dir_path = os.path.join(load_root,data_sets[data_set_idx],'train')
valid_dir_path = os.path.join(load_root,data_sets[data_set_idx],'valid')
test_dir_path = os.path.join(load_root,data_sets[data_set_idx],'test')
#############################
# Helper functions
#############################
def clip_roll(piano_roll, time_steps=50):
samples = []
num_samples = int(piano_roll.shape[1] / time_steps)
for i in range(num_samples):
start_idx = time_steps*i
end_idx = (time_steps*(i+1))
samples.append(piano_roll[:,start_idx:end_idx])
return samples
def create_samples(load_root, time_steps=50, verbose=False):
if not os.path.isdir(load_root):
print("Invalid load_root directory.")
sys.exit(0)
samples = []
for (dirpath, dirnames, filenames) in os.walk(load_root):
for file in filenames:
if file.endswith('.npy'):
load_filepath = os.path.join(dirpath,file)
if verbose:
print(load_filepath)
piano_roll = np.load(load_filepath).T
samples = samples + clip_roll(piano_roll,time_steps=time_steps)
return np.stack(samples)
def feed_dict(batch_size, samples_type=None):
if samples_type == 'train':
indeces = np.random.randint(num_train_samples, size=batch_size)
samples = train_samples
elif samples_type == 'valid':
indeces = np.random.randint(num_valid_samples, size=batch_size)
samples = valid_samples
elif samples_type == 'test':
indeces = np.random.randint(num_test_samples, size=batch_size)
samples = test_samples
return np.take(samples, indeces, axis=0)
#############################
# Setup Load/Save paths
#############################
train_samples = create_samples(train_dir_path, time_steps=time_steps, verbose=False)
valid_samples = create_samples(valid_dir_path, time_steps=time_steps, verbose=False)
test_samples = create_samples(test_dir_path, time_steps=time_steps, verbose=False)
num_train_samples = train_samples.shape[0]
num_valid_samples = valid_samples.shape[0]
num_test_samples = test_samples.shape[0]
network_params = ''.join([
'time_steps={}-'.format(time_steps),
'latent_dim={}-'.format(z_dim),
'dataset={}'.format(data_sets[data_set_idx])])
# Dir structure : /base_dir/network_params/run_xx/train_or_test/
log_base_dir = os.path.join(log_root, network_params)
# Check for previous runs
if not os.path.isdir(log_base_dir):
os.makedirs(log_base_dir)
previous_runs = os.listdir(log_base_dir)
if len(previous_runs) == 0:
run_number = 1
else:
run_number = max([int(str.split(s,'run_')[1]) for s in previous_runs if 'run' in s]) + 1
if epoch_to_load > 0:
run_number = run_to_load
log_dir = os.path.join(log_base_dir,'run_{0:02d}'.format(run_number))
train_summary_writer = tf.summary.FileWriter(log_dir + '/train')
valid_summary_writer = tf.summary.FileWriter(log_dir + '/valid')
test_summary_writer = tf.summary.FileWriter(log_dir + '/test')
model_save_path = log_dir + '/models'
#############################
# Setup graph
#############################
tf.reset_default_graph()
X = tf.placeholder(tf.float32, shape=(batch_size, x_in_dim, time_steps))
# time_slices containts input x at time t across batches.
x_in = time_steps * [None]
x_out = time_steps * [None]
h_enc = time_steps * [None]
h_dec = (time_steps + 1) * [None]
for t in range(time_steps):
x_in[t] = tf.squeeze(tf.slice(X,begin=[0,0,t],size=[-1,-1,1]),axis=2)
###### Encoder network ###########
with tf.variable_scope('encoder_rnn'):
cell_enc = tf.nn.rnn_cell.BasicRNNCell(num_hidden_units,activation=tf.nn.tanh)
h_enc[0] = tf.zeros([batch_size,num_hidden_units], dtype=tf.float32) # Initial state is 0
# h_t+1 = tanh(Wenc*h_t + Win*x_t+1 + b )
#Most basic RNN: output = new_state = act(W * input + U * state + B).
#https://github.com/tensorflow/tensorflow/blob/r1.4/tensorflow/python/ops/rnn_cell_impl.py
for t in range(time_steps-1):
_ , h_enc[t+1] = cell_enc(inputs=x_in[t+1], state=h_enc[t])
mu_enc = tf.layers.dense(h_enc[-1], z_dim, activation=None, name='mu_enc')
log_sigma_enc = tf.layers.dense(h_enc[-1], z_dim, activation=None, name='log_sigma_enc')
###### Reparametrize ##############
eps = tf.random_normal(tf.shape(log_sigma_enc))
z = mu_enc + tf.exp(log_sigma_enc) * eps
##### Decoder network ############
with tf.variable_scope('decoder_rnn'):
W_out = tf.get_variable('W_out',shape=[num_hidden_units, x_in_dim])
b_out = tf.get_variable('b_out',shape=[x_in_dim])
cell_dec = tf.nn.rnn_cell.BasicRNNCell(num_hidden_units,activation=tf.nn.tanh)
h_dec[0] = tf.layers.dense(z, num_hidden_units, activation=tf.nn.tanh)
for t in range(time_steps):
x_out[t] = tf.nn.sigmoid(tf.matmul(h_dec[t], W_out) + b_out)
if t < time_steps - 1:
_, h_dec[t+1] = cell_dec(inputs=x_out[t], state=h_dec[t])
##### Loss #####################
with tf.variable_scope('loss'):
# Latent loss: -KL[q(z|x)|p(z)]
with tf.variable_scope('latent_loss'):
sigma_sq_enc = tf.square(tf.exp(log_sigma_enc))
latent_loss = -.5 * tf.reduce_mean(tf.reduce_sum((1 + tf.log(1e-10 + sigma_sq_enc)) - tf.square(mu_enc) - sigma_sq_enc, axis=1),axis=0)
latent_loss_summ = tf.summary.scalar('latent_loss',latent_loss)
# Reconstruction Loss: log(p(x|z))
with tf.variable_scope('recon_loss'):
for i in range(time_steps):
if i == 0:
recon_loss_ = x_in[i] * tf.log(1e-10 + x_out[i]) + (1 - x_in[i]) * tf.log(1e-10+1-x_out[i])
else:
recon_loss_ += x_in[i] * tf.log(1e-10 + x_out[i]) + (1 - x_in[i]) * tf.log(1e-10+1-x_out[i])
#collapse the loss, mean across a sample across all x_dim and time points, mean over batches
recon_loss = -tf.reduce_mean(tf.reduce_mean(recon_loss_/(time_steps),axis=1),axis=0)
recon_loss_summ = tf.summary.scalar('recon_loss', recon_loss)
with tf.variable_scope('total_loss'):
total_loss = latent_loss + recon_loss
total_loss_summ = tf.summary.scalar('total_loss', total_loss)
global_step = tf.Variable(0,name='global_step')
#learning_rate = tf.train.exponential_decay(initial_learning_rate, epoch_num, num_epochs, decay_rate, staircase=False)
learning_rate = tf.Variable(learning_rate_1,name='learning_rate')
train_step = tf.train.AdamOptimizer(learning_rate=learning_rate, beta1=beta_1, beta2=beta_2).minimize(total_loss,global_step=global_step)
scalar_summaries = tf.summary.merge([latent_loss_summ, recon_loss_summ, total_loss_summ])
#image_summaries = tf.summary.merge()
train_summary_writer.add_graph(tf.get_default_graph())
#############################
# Training/Logging
#############################
num_batches = int(num_train_samples/batch_size)
global_step_op = tf.train.get_global_step()
saver = tf.train.Saver()
with tf.Session(config=tf.ConfigProto(log_device_placement=True)) as sess:
# Epoch_to_load has sentinel value of -1 for train mode. else it specifies the epoch to load.
if epoch_to_load >= 0:
saver.restore(sess, model_save_path + '/epoch_{}.ckpt'.format(epoch_to_load))
loss, x_out_, x_in_, learning_rate_, latent_loss_ = \
sess.run([total_loss, x_out, x_in, learning_rate, latent_loss],feed_dict={X: feed_dict(batch_size,'train')})
else:
sess.run(tf.global_variables_initializer())
for epoch in range(num_epochs):
epoch_loss = 0.
epoch_latent_loss = 0.
for batch in range(num_batches):
batch_num = sess.run(global_step_op)
if epoch < num_epochs_to_diff_learn_rate:
curr_learning_rate = learning_rate_1
else:
curr_learning_rate = learning_rate_2
#_ , loss, scalar_train_summaries, x_out_, x_in_,learning_rate_,latent_loss_ = \
#sess.run([train_step, total_loss, scalar_summaries, x_out, x_in,learning_rate, latent_loss],feed_dict={X: feed_dict(batch_size,'train'), learning_rate: curr_learning_rate})
_ , loss, scalar_train_summaries, learning_rate_, latent_loss_ = \
sess.run([train_step, total_loss, scalar_summaries, learning_rate, latent_loss],feed_dict={X: feed_dict(batch_size,'train'), learning_rate: curr_learning_rate})
# Check for NaN
if np.isnan(loss):
sys.exit("Loss during training at epoch: {}".format(epoch))
epoch_loss += loss
epoch_latent_loss += latent_loss_
print('Average loss epoch {0}: {1}'.format(epoch, epoch_loss/num_batches))
print('Average latent loss epoch {0}: {1}'.format(epoch, epoch_latent_loss/num_batches))
print('Learning Rate {}'.format(learning_rate_))
# Write train summaries once a epoch
scalar_train_summaries = sess.run(scalar_summaries,feed_dict={X: feed_dict(batch_size,'train')})
train_summary_writer.add_summary(scalar_train_summaries, global_step=batch_num)
# Write validation summaries
scalar_valid_summaries = sess.run(scalar_summaries,feed_dict={X: feed_dict(batch_size,'valid')})
valid_summary_writer.add_summary(scalar_valid_summaries, global_step=batch_num)
# Write test summaries
scalar_test_summaries = sess.run(scalar_summaries,feed_dict={X: feed_dict(batch_size,'test')})
test_summary_writer.add_summary(scalar_test_summaries, global_step=batch_num)
# Save the models
if epoch % num_epochs_to_save_model == 0:
save_path = saver.save(sess, model_save_path + '/epoch_{}.ckpt'.format(epoch))
def plot_x_io(x, samp_num):
x_arr = np.asarray(x)
plt.imshow(x_arr[:,samp_num,:].T)
plt.show()
plot_x_io(x_in_, 0)
plot_x_io(x_out_, 0)
```
| github_jupyter |
# Arguments Against the Implementation Invariance of PA and PGIG
In our [paper](https://arxiv.org/abs/2007.10685) we left the question of whether or not PGIG is implementation invariant to future work.
In this notebook we present arguments against the implementation invariance of PA and PGIG.
## Counter Example: Sketch
The proof (by counter example) goes as follows. We construct two functionally equivalent networks which share weights in their first layers but still have different positive regimes, due to different biases. Since the $a_{+}$ pattern used for PatternAttribution in [the original paper](https://arxiv.org/abs/1705.05598) is a function of the positive regime, different patterns emerge for the shared weights. The different patterns then lead to different attributions (explanations) for identical inputs (into the two functionally equivalent networks) and thus PA and PGIG should not be implementation invariant.
We consider the two networks
$f'(\mathbf{x}) = ReLU(\mathcal{I}_{1}ReLU(\mathbf{w}^{T}\mathbf{x} - 0) - b)$
and
$f''(\mathbf{x}) = ReLU(\mathbf{w}^{T}\mathbf{x} - b)$
where $\mathcal{I}_{1}=1$ is the identity matrix. The networks are functionally equivalent for $b>0$ but subtract the non-zero bias in different layers. The networks are very similar to the ones used in the [IG paper](https://arxiv.org/abs/1703.01365), in which they are also used in the context of implementation invariance.
In what follows we fix weights and biases, generate data, compute patterns, modify the weights with the patterns and propagate back modified gradients, which yield the attributions according to PA.
Let us set
$w=(1,2)^{T}, b=1$ and refer to the first network as the network 1 and the second as network 2.
```
import numpy as np
import matplotlib.pyplot as plt
import os
from typing import Tuple
np.random.seed(42)
import seaborn as sns
```
## Pattern Computation
We copy the pattern functions from previous notebooks.
```
def positive_regime(input: np.array, output: np.array, feature_in: int, feature_out: int) -> Tuple[np.array, np.array]:
"""
Filter the positive regime.
Args:
input: Input array of shape (N, features_in).
output: Output array of shape (N, features_out).
feature_in: Input feature dimension to consider with 0 <= feature_in <= dim_in.
feature_out: Output feature dimension to consider with
Returns: Filtered input array of shape (N-M, 1) and corresponding output array of shape (N-M, 1),
where M is the number of negative outputs.
"""
# Collect input and output features at positions feature_in and feature_out, respectively.
features_in = input[:, feature_in]
features_out = output[:, feature_out]
# Collect inputs and corresponding outputs where feature_out > 0
x_plus = []
y_plus = []
for idx, feature_out in enumerate(features_out):
if feature_out > 0:
x_plus.append(features_in[idx])
y_plus.append(features_out[idx])
return np.array(x_plus), np.array(y_plus)
def patterns(input: np.array, output: np.array, weights: np.array) -> np.array:
"""
The pattern estimation according to Eq. 7 (Kindermans et al., 2017) for the positive regime.
Args:
input: Input to the model of shape (N, features_in).
output: ReLU activated output of the model of shape (N, features_out).
weights: Weights of the model of shape (features_in, features_out).
Returns: The pattern estimation (a+) of shape (features_in, features_out).
"""
# Create three matrices E[x+ y], E[x+].
E_x_plus_matrix = np.zeros_like(weights, dtype=np.double)
E_x_plus_times_y_matrix = np.zeros_like(weights, dtype=np.double)
# Populate the matrices above.
dims_in, dims_out = weights.shape
for dim_in in range(dims_in):
for dim_out in range(dims_out):
# Collect all x,y for which ReLU(wTx) = y > 0
x_plus, y_plus = positive_regime(input=input,
output=output,
feature_in=dim_in,
feature_out=dim_out)
# Create the expected values, aka means.
E_x = np.mean(x_plus)
E_x_y = np.mean(x_plus * y_plus)
# Populate the matrices above.
E_x_plus_matrix[dim_in][dim_out] = E_x
E_x_plus_times_y_matrix[dim_in][dim_out] = E_x_y
E_y = np.mean(output)
E_y_matrix = np.full_like(weights, E_y, dtype=np.double)
# Compute the nominator and denominator according to Eq. 7.
nominator = E_x_plus_times_y_matrix - (E_x_plus_matrix * E_y_matrix)
denominator = np.matmul(weights.T, E_x_plus_times_y_matrix) - np.matmul(weights.T, (E_x_plus_matrix * E_y_matrix))
pattern = nominator / denominator
return pattern
```
## Data
We are considering a bivariate distribution. $x1$ is sampled from a normal distribution. $x2 = x1k$, where $k$ is sampled from a Rademacher distribution.
See [Wikipedia](https://en.wikipedia.org/wiki/Normally_distributed_and_uncorrelated_does_not_imply_independent).
```
size = 5000 # number of samples to work with
mean = 0.0
std_dev = 0.7 # change this to change pattern differences btw network 1 and network 2.
x1 = np.random.normal(size=size, loc=mean, scale=std_dev) # loc = 0, scale = 1 if not overwritten
rand = np.random.choice(a=[False, True], size=size, p=[.5, .5]) # Rademacher distribution
x2 = []
for x, r in zip(list(x1), list(rand)):
if r:
x2.append(x)
else:
x2.append(-x)
ipt = np.array([np.array([_x1, _x2]) for _x1, _x2 in zip(x1, x2)]) # original input
w1 = 1.
w2 = 2.
wgt = np.array([[w1], [w2]]) # shared weights
wgt_n1_l2 = np.array([[1.0]]) # weights of the second layer of network 1
opt_n1_l1 = [np.matmul(wgt.T, i) for i in ipt] # the output of the 1st layer of network 1 (w/ zero bias)
opt_n1_l1 = np.array(opt_n1_l1)
ipt_n1_l2 = np.array([max(np.array([0]), o) for o in opt_n1_l1]) # ReLU activate the output of the 1st layer
opt_n1_l2 = np.array([i-1 for i in ipt_n1_l2]) # substract unit bias, which yields the output of the 2nd layer
opt_n2_l1 = [o - 1 for o in opt_n1_l1] # the output of the 1st layer with of network 2 (w/ unit bias)
opt_n2_l1 = np.array(opt_n2_l1)
```
## Patterns
In what follows we compute patterns for all layers.
```
patterns_network_one_layer_one = patterns(input=ipt, output=opt_n1_l1, weights=wgt)
patterns_network_one_layer_two = patterns(input=ipt_n1_l2, output=opt_n1_l2, weights=wgt_n1_l2)
patterns_network_two_layer_one = patterns(input=ipt, output=opt_n2_l1, weights=wgt)
print(f'Patterns of network 1 layer 1 w/ zero bias:\n {patterns_network_one_layer_one}')
print(f'Patterns of network 1 layer 2 w/ unit bias:\n {patterns_network_one_layer_two}')
print(f'Patterns of network 2 layer 1 w/ unit bias:\n {patterns_network_two_layer_one}')
```
## Attributions
In what follows we will compute attributions for both networks.
We will consider an input that exists in all positive regimes, namely $x = (3,3)^{T}$.
```
x = np.array([3.0, 3.0])
```
### Network 1
```
# forward pass
y_n1 = np.matmul(wgt.T, x) # first layer
assert y_n1 > 0, 'ReLU should not filter in this experiment'
y_n1 = y_n1 - 1 # second layer
assert y_n1 > 0, 'ReLU should not filter in this experiment'
# modify weights prior to backward pass
wgt_p_n1_l1 = wgt * patterns_network_one_layer_one
wgt_p_n1_l2 = wgt_n1_l2 * patterns_network_one_layer_two
# backward pass
grad_n1 = y_n1 # PA starts backpropagation w/ y
grad_n1 = wgt_p_n1_l2 * grad_n1 # 2nd relu does not filter (see assertions), multiply w/ mod. weights
attributions_n1 = wgt_p_n1_l1 * grad_n1 # 1st relu does not filter (see assertions), multiply w/ mod. weights
print(f'PatternAttribution Network 1:\n {attributions_n1}')
```
### Network 2
```
# forward pass
y_n2 = np.matmul(wgt.T, x) - 1
assert y_n2 > 0, 'ReLU should not filter in this experiment'
assert y_n2 == y_n1, 'Networks should be functionally equivalent'
# modify weights prior to backward pass
wgt_p_n2_l1 = wgt * patterns_network_two_layer_one
# backward pass
grad_n2 = y_n2 # PA starts backprop with y
attributions_n2 = wgt_p_n2_l1 * grad_n2 # relu does not filter (see assertion above), multiply w/ mod. weights
print(f'PatternAttribution Network 2:\n {attributions_n2}')
```
## Visualization
Below, the explanations for $x$ are visualized in form of heatmaps.
```
vmin = min(np.min(attributions_n1), np.min(attributions_n2))
vmax = max(np.max(attributions_n1), np.max(attributions_n2))
fig, (ax1, ax2) = plt.subplots(1,2)
sns.heatmap(attributions_n1, vmin=vmin, vmax=vmax, cmap="Spectral", ax=ax1)
sns.heatmap(attributions_n2, vmin=vmin, vmax=vmax, cmap="Spectral", ax=ax2)
```
## Conclusion
We receive two different explanations for identical inputs into functionally equivalent networks and thus PA and PGIG are not implementation invariant.
| github_jupyter |
```
import pandas as pd
import os
import numpy as np
from scipy.sparse import coo_matrix, csr_matrix
from datatool import ade,vid_is_unique,foot2meter,vehicle2track,reset_idx,getNeighborGraph,graph2seq,get_displacement,train_test_val_split,matlab2dataframe
from collections import Counter
import time
import pickle as pkl
def add_change_label(data):
data = data.sort_values(by=["id","frame"])
changes = []
for vid, df in data.groupby("id"):
lane_id = df.laneId.to_numpy()
diff = lane_id[1:] - lane_id[:-1]
changed_or_not = (diff!=0)*1 # 1 : change; 0 : unchange
changed_or_not = np.hstack((np.zeros(1),changed_or_not))
changes.append(changed_or_not)
change_result = np.hstack(changes)
return change_result
def getNeighborGraph(data,radius=10):
x,y,v_id,f_id = data.x,data.y,data.id,data.frame
vehicle_num, frame_num = v_id.max()+1, f_id.max()+1
sparse_X = csr_matrix((x, (v_id, f_id)), shape=(int(vehicle_num), int(frame_num))) # i行:车id;j列:时间;元素为i车j时刻的坐标x
sparse_Y = csr_matrix((y, (v_id, f_id)), shape=(int(vehicle_num), int(frame_num))) # i行:车id;j列:时间;元素为i车j时刻的坐标y
I_mat = (sparse_X!=0)*1 # i行:车id;j列:时间;元素为i车j时刻是否出现,出现为1,否则为0
mask = []
for v in range(I_mat.shape[0]):
concurrent_mask = I_mat.multiply(I_mat[v]) #同或 [1,0,1,0,0,0,1] & [1,0,1,1,1,0,0] = [1,0,1,0,0,0,0]
# 邻居xy坐标
concurrent_X = concurrent_mask.multiply(sparse_X)
concurrent_Y = concurrent_mask.multiply(sparse_Y)
# 自己xy坐标
self_x = concurrent_mask.multiply(sparse_X[v])
self_y = concurrent_mask.multiply(sparse_Y[v])
# 差值
delta_x = self_x - concurrent_X
delta_y = self_y - concurrent_Y
# 邻居x坐标在半径以内的指示矩阵
x_in_id = np.where((delta_x.data>-radius) & (delta_x.data<radius))
xc = delta_x.tocoo()
xrow_in = xc.row[x_in_id]
xcol_in = xc.col[x_in_id]
xI_data = np.ones(xrow_in.shape[0])
xneighbor_in_mat = csr_matrix((xI_data, (xrow_in, xcol_in)), shape=(I_mat.shape[0], I_mat.shape[1]))
# 邻居y坐标在半径以内的指示矩阵
y_in_id = np.where((delta_y.data>-radius) & (delta_y.data<radius))
yc = delta_y.tocoo()
yrow_in = yc.row[y_in_id]
ycol_in = yc.col[y_in_id]
yI_data = np.ones(yrow_in.shape[0])
yneighbor_in_mat = csr_matrix((yI_data, (yrow_in, ycol_in)), shape=(I_mat.shape[0], I_mat.shape[1]))
neighbor_in_mat = xneighbor_in_mat.multiply(yneighbor_in_mat).tolil()
neighbor_in_mat[v] = I_mat[v]
mask.append(neighbor_in_mat.tocsr())
return mask
def graph2seq(data,graph_list,seq_length=16,max_vnum=30,down_sample_rate=5,sort_func="distance"):
x,y,v_id,f_id,l = data.x,data.y,data.id,data.frame,data.label
vehicle_num, frame_num = v_id.max()+1, f_id.max()+1
sparse_X = csr_matrix((x, (v_id, f_id)), shape=(int(vehicle_num), int(frame_num))) # i行:车id;j列:时间;元素为i车j时刻的坐标x
sparse_Y = csr_matrix((y, (v_id, f_id)), shape=(int(vehicle_num), int(frame_num))) # i行:车id;j列:时间;元素为i车j时刻的坐标y
sparse_L = csr_matrix((l, (v_id, f_id)), shape=(int(vehicle_num), int(frame_num))) # i行:车id;j列:时间;元素为i车j时刻的下一时刻是否lane change
seq_windows,label = [],[]
for v,graph in enumerate(graph_list):
if graph.data.size==0:
continue
row = np.unique(graph.tocoo().row)
col = np.unique(graph.tocoo().col)
row_start,row_end = row.min(), row.max()+1
col_start,col_end = col.min(), col.max()+1
dense_v = v - row_start
dense_I = graph[row_start:row_end,col_start:col_end].toarray()
dense_x = sparse_X[row_start:row_end,col_start:col_end].toarray()
dense_y = sparse_Y[row_start:row_end,col_start:col_end].toarray()
dense_xy = np.stack((dense_x,dense_y),axis=2) # (vum,total_seq,2)
dense_l = sparse_L[row_start:row_end,col_start:col_end].toarray()
if dense_xy.shape[0]<max_vnum:
padding_num = max_vnum-dense_xy.shape[0]
padding_xy = np.zeros((padding_num,dense_xy.shape[1],dense_xy.shape[2]))
padding_I = np.zeros((padding_num,dense_I.shape[1]))
dense_xy = np.vstack([dense_xy,padding_xy])
dense_I = np.vstack([dense_I,padding_I])
dense_l = np.vstack([dense_l,padding_I])
for i in range(dense_xy.shape[1]): # for loop on sequence dim
if (i+seq_length)*down_sample_rate > dense_xy.shape[1]:
break
window = dense_xy[:,i:i+seq_length*down_sample_rate:down_sample_rate,:] # (vum=30,seq=16,2)
window_l = dense_l[:,i:i+seq_length*down_sample_rate:down_sample_rate] # (vum=30,seq=16)
if sort_func == "duration":
dense_seq_I = dense_I[:,i:(i+seq_length)*down_sample_rate:down_sample_rate]
related_score = dense_seq_I.sum(axis=1)
related_score[dense_v] = related_score[dense_v] + 100 # actually 1 is enough
related_rank = np.argsort(-related_score)
elif sort_func == "distance":
related_score = ade(window[:,:6,:],window[dense_v,:6,:])
related_rank = np.argsort(related_score)
window = window[related_rank[:max_vnum],:,:]
seq_windows.append(window)
label.append((window_l[0,6:].sum()>0)*1) # 30,17,2 (0-5),6,(7-16)
if len(seq_windows)==0:
seq_data = None
seq_label = None
else:
seq_data = np.stack(seq_windows)#(n,vum=30,seq=16,2)
seq_label = np.stack(label)
return seq_data,seq_label
selected_col = ["frame","id","x","y","laneId"]
total_length = 0
for i in range(60):
t1 = time.time()
if i < 9:
path = f"highD-dataset-v1.0/data/0{i+1}_tracks.csv"
else:
path = f"highD-dataset-v1.0/data/{i+1}_tracks.csv"
data = pd.read_csv(path)
useful_data = data[selected_col]
useful_data = useful_data.sort_values(by=["id","frame"])
label = add_change_label(useful_data)
useful_data["label"] = pd.Series(label)
'''
uni_id = selected_data.id.unique()
mapping_uni_id = np.arange(uni_id.shape[0])
new_uni_id = mapping_uni_id + total_length
total_length += uni_id.shape[0]
id_dict = dict(zip(uni_id, new_uni_id))
new_id = np.vectorize(id_dict.get)(data.id)
selected_data.id = new_id
'''
begin = useful_data[useful_data["label"]==1].index - 20
end = useful_data[useful_data["label"]==1].index + 20
new_label = np.zeros(useful_data.shape[0])
for b,e in zip(begin,end):
new_label[b:e] = 1
useful_data['label'] = new_label
neighbor_graph = getNeighborGraph(useful_data,radius=50)
seq_data,seq_label = graph2seq(useful_data,neighbor_graph,seq_length=17)
t2 = time.time()
print(f"file_{i} processed. time: {t2-t1:.2f}. data shape {seq_data.shape}. label shape {seq_label.shape}.")
file_data = {"data":seq_data,"label":seq_label}
with open(f"pickle_data/data_{i}.pkl","wb") as f:
pkl.dump(file_data,f)
pickle_folder = "pickle_data"
total_data,total_label = [],[]
for file_path in os.listdir(pickle_folder):
if "_" not in file_path:
continue
path = os.path.join(pickle_folder,file_path)
with open(path,"rb") as f:
pickle_file = pkl.load(f)
label = pickle_file["label"]
data = pickle_file["data"]
pos_data = data[label==1]
neg_data = data[label==0]
pos_data_num = pos_data.shape[0]
total_data.append(pos_data.repeat(10,axis=0))
total_data.append(neg_data[:pos_data_num*10])
total_label.append(np.ones(pos_data_num*10))
total_label.append(np.zeros(pos_data_num*10))
data_array = np.vstack(total_data)
label_array = np.hstack(total_label)
print(data_array.shape, label_array.shape)
pickle_folder = "pickle_data"
pos_data_list, neg_data_list = [], []
for file_path in os.listdir(pickle_folder):
if "_" not in file_path:
continue
path = os.path.join(pickle_folder,file_path)
with open(path,"rb") as f:
pickle_file = pkl.load(f)
label = pickle_file["label"]
data = pickle_file["data"]
pos_data = data[label==1]
neg_data = data[label==0]
pos_data_num = pos_data.shape[0]
pos_data_list.append(pos_data)
neg_data_list.append(neg_data[:pos_data_num*10])
pos_data_array = np.vstack(pos_data_list)
neg_data_array = np.vstack(neg_data_list)
print(pos_data_array.shape,neg_data_array.shape)
pickle_folder = "pickle_data"
keep_data_list, right_data_list,left_data_list = [], [], []
for file_path in os.listdir(pickle_folder):
if "data_" not in file_path:
continue
path = os.path.join(pickle_folder,file_path)
with open(path,"rb") as f:
pickle_file = pkl.load(f)
label = pickle_file["label"]
data = pickle_file["data"]
left_number = right_number = keep_number = 0
if (label == 1).any():
left_data = data[label==1]
left_data_list.append(left_data)
left_number = left_data.shape[0]
if (label == -1).any():
right_data = data[label==-1]
right_data_list.append(right_data)
right_number = right_data.shape[0]
if (label == 0).any():
keep_number = max(left_number,right_number)*10
keep_data = data[label==0]
keep_data_list.append(keep_data[:keep_number])
print(f"file: {file_path}, keep: {keep_number}, right: {right_number}, left: {left_number}")
keep_data_array = np.vstack(keep_data_list)
right_data_array = np.vstack(right_data_list)
left_data_array = np.vstack(left_data_list)
print(keep_data_array.shape,right_data_array.shape,left_data_array.shape)
_ = {
"right_data":right_data_array,
"left_data":left_data_array,
"keep_data":keep_data_array
}
with open("pickle_data/23w10v1_3cls.pkl","wb") as f:
pkl.dump(_,f)
_ = {
"pos_data":pos_data_array,
"neg_data":neg_data_array,
}
with open("pickle_data/23w10v1.pkl","wb") as f:
pkl.dump(_,f)
import pickle as pkl
import numpy as np
from datatool import train_test_val_split
with open("pickle_data/23w10v1.pkl","rb") as f:
data = pkl.load(f)
pos_data,neg_data = data['pos_data'],data['neg_data']
print(pos_data.shape,neg_data.shape)
pos_train, pos_val, pos_test = train_test_val_split(pos_data,test_size=0.2,val_size=0.1,seed=0)
neg_train, neg_val, neg_test = train_test_val_split(neg_data,test_size=0.2,val_size=0.1,seed=0)
pos_train, pos_val, pos_test = pos_train.repeat(10,axis=0),pos_val.repeat(10,axis=0),pos_test.repeat(10,axis=0)
print(pos_train.shape,pos_val.shape,pos_test.shape)
print(neg_train.shape,neg_val.shape,neg_test.shape)
X_train, X_val, X_test = np.vstack((pos_train,neg_train)),np.vstack((pos_val,neg_val)),np.vstack((pos_test,neg_test))
y_train = np.hstack((np.ones(pos_train.shape[0]),np.zeros(neg_train.shape[0])))
y_val = np.hstack((np.ones(pos_val.shape[0]),np.zeros(neg_val.shape[0])))
y_test = np.hstack((np.ones(pos_test.shape[0]),np.zeros(neg_test.shape[0])))
print(X_train.shape,y_train.shape)
print(X_val.shape,y_val.shape)
print(X_test.shape,y_test.shape)
```
| github_jupyter |
# OOP training: solutions
## 3.3 - Exercise 1
```
class Rectangle:
def __init__(self, width, height):
self.width = width
self.height = height
def area(self):
return self.width*self.height
def perimeter(self):
return 2*(self.width+self.height)
r = Rectangle(4,5)
print(r.area(), r.perimeter())
```
## 4.2 - Exercise 2
```
class Rectangle:
"Rectangle class"
def __init__(self, width, height):
self.width = width
self.height = height
@property
def area(self):
return self.width*self.height
@property
def perimeter(self):
return 2*(self.width+self.height)
r = Rectangle(4,5)
print(r.area, r.perimeter)
print(r.__doc__)
r.__class__.__dict__
```
## 5.1 - Exercise 3
```
class Square(Rectangle):
def __init__(self, side):
super().__init__(side, side)
@property
def area(self):
return self.width*self.height
@area.setter
def area(self, area):
self.width = self.length = (area)**0.5
c = Square(5)
c.area
c.area = 16
c.length
```
## 5.3 - Exercise on inheritance: Laue diffraction pattern
```
"""Laue simulation code"""
import numpy
def laue_array_size(ncells, oversampling):
"""Compute the output array size in each dimension
:param int ncells:
Number of unit cells in both directions
:param int oversampling: Oversampling factor
:rtype: int
"""
return ncells * oversampling
def laue_image(ncells, h, k, oversampling):
"""
:param int ncells:
Number of unit cells in both directions
:param int h:
H Miller index of reflection where to sample space
:param int k:
K Miller index of reflection where to sample space
:param int oversampling:
Oversampling factor
:return: 2D array
:rtype: numpy.ndarray
"""
size = laue_array_size(ncells, oversampling)
# Prepare cristal structure
n = numpy.arange(ncells)
m = numpy.arange(ncells)
# Prepare sampling positions
h_sampling_pos = numpy.linspace(h - 0.5, h + 0.5, size, endpoint=True)
k_sampling_pos = numpy.linspace(k - 0.5, k + 0.5, size, endpoint=True)
# Do the computation
h, k, n, m = numpy.meshgrid(h_sampling_pos, k_sampling_pos, n, m, sparse=True)
# Sum over the unit-cells (last axis of the array) and take the squared modulus
return numpy.abs(numpy.exp(2j*numpy.pi*(h*n + k*m)).sum(axis=(2,3)))**2
import threading
class LaueThread(threading.Thread):
def __init__(self, ncells, h, k, oversampling):
self.ncells = ncells
self.h = h
self.k = k
self.oversampling = oversampling
self.result = None
super(LaueThread, self).__init__(name="LaueThread", group=None)
def run(self):
self.result = laue_image(self.ncells, self.h, self.k, self.oversampling)
t=LaueThread(10,5,5,50)
t.start()
import time
for i in range(100):
print(t.result)
time.sleep(0.1)
```
| github_jupyter |
```
# Computations
import numpy as np
import pandas as pd
# Modeling
import catboost
# Tools
import os
import datetime
import calendar
import itertools
# Dask
from dask.distributed import Client, progress
import dask
import dask.dataframe as dd
client = Client()
# Sklearn
from sklearn import metrics
from sklearn.preprocessing import StandardScaler
from sklearn.impute import SimpleImputer
# Visualisation libraries
## progressbar
import progressbar
## Text
from colorama import Fore, Back, Style
from IPython.display import Image, display, Markdown, Latex, clear_output
## plotly
from plotly.offline import init_notebook_mode, iplot
import plotly.graph_objs as go
import plotly.offline as py
from plotly.subplots import make_subplots
import plotly.express as px
## seaborn
import seaborn as sns
sns.set_style("whitegrid")
sns.set_context("paper", rc={"font.size":12,"axes.titlesize":14,"axes.labelsize":12})
## matplotlib
import matplotlib.pyplot as plt
from matplotlib.font_manager import FontProperties
import matplotlib.colors as mcolors
from matplotlib.patches import Ellipse, Polygon
import matplotlib.gridspec as gridspec
from pylab import rcParams
plt.style.use('seaborn-whitegrid')
import matplotlib as mpl
mpl.rcParams['figure.figsize'] = (17, 6)
mpl.rcParams['axes.labelsize'] = 14
mpl.rcParams['xtick.labelsize'] = 12
mpl.rcParams['ytick.labelsize'] = 12
mpl.rcParams['text.color'] = 'k'
%matplotlib inline
import warnings
warnings.filterwarnings("ignore")
```
<div class="alert alert-block alert-success">
<a href="https://hatefdastour.github.io/">Home</a> /
<a href="https://hatefdastour.github.io/portfolio/financial_analysis_and_modeling/">Financial Analysis and Modeling</a>
</div>
## Predict Future Sales
* [**Preprocessing**](https://hatefdastour.github.io/portfolio/financial_analysis_and_modeling/Predict_Future_Sales_Preprocessing.html)
* [**Exploratory Data Analysis**](https://hatefdastour.github.io/portfolio/financial_analysis_and_modeling/Predict_Future_Sales_EDA.html)
* [<font color='Green'><b>Modeling: CatBoost Regressor</b></font>](https://hatefdastour.github.io/portfolio/financial_analysis_and_modeling/Predict_Future_Sales_Modeling_CatBoostReg.html)
<div class="alert alert-block alert-info">
<font size="+2.5"><b>
Predict Future Sales
</b></font>
</div>
In this article, we work with the [Predict Future Sales](https://www.kaggle.com/c/competitive-data-science-predict-future-sales/overview) provided by one of the largest Russian software firms - [1C Company](http://1c.ru/eng/title.htm).
## Data Description
You are provided with daily historical sales data. The task is to forecast the total amount of products sold in every shop for the test set. Note that the list of shops and products slightly changes every month. Creating a robust model that can handle such situations is part of the challenge.
```
Data_dict = {'ID': 'ID', 'City': 'The City that a shop is located',
'City ID': 'Assigned numeric ID for a city',
'Date':'Date in format dd/mm/yyyy',
'Date Block Number':'A consecutive month number, used for convenience. January 2013 is 0, february 2013 is 1,..., October 2015 is 33',
'Day': 'Day of month', 'Day of Week': 'Day of Week', 'Item Category': 'Item Category',
'Item Category Full':'Item Category and Subcategory Name',
'Item Category Full ID': 'Assigned numeric ID for a Item Category',
'Item Category ID': 'Assigned numeric ID for a Item Category',
'Item Count Day':'The number of products sold. You are predicting a monthly amount of this measure',
'Item ID':'Unique identifier of a product', 'Item Name':'The name of item',
'Item Price':'Current price of an item',
'Item Subcategory': 'Item Subcategory',
'Item Subcategory ID': 'Assigned numeric ID for a Item Subcategory', 'Month': 'Month',
'Month Name': 'Month Name', 'Month Triannually': 'Month Triannually', 'Revenue': 'Revenue',
'Season': 'Season', 'Shop ID':'Unique identifier of a shop',
'Item Category ID':'Unique identifier of item category',
'Shop Name':'The name of shop', 'Year':'Year'}
def Data_fields(Data, Data_dict = Data_dict):
Data_dict_df = pd.DataFrame({'Feature':Data.columns})
Data_dict_df['Description'] = Data_dict_df['Feature'].map(Data_dict)
display(Data_dict_df.style.hide_index())
def Header(Text, L = 100, C = 'Blue', T = 'White'):
BACK = {'Black': Back.BLACK, 'Red':Back.RED, 'Green':Back.GREEN, 'Yellow': Back.YELLOW, 'Blue': Back.BLUE,
'Magenta':Back.MAGENTA, 'Cyan': Back.CYAN}
FORE = {'Black': Fore.BLACK, 'Red':Fore.RED, 'Green':Fore.GREEN, 'Yellow':Fore.YELLOW, 'Blue':Fore.BLUE,
'Magenta':Fore.MAGENTA, 'Cyan':Fore.CYAN, 'White': Fore.WHITE}
print(BACK[C] + FORE[T] + Style.NORMAL + Text + Style.RESET_ALL + ' ' + FORE[C] +
Style.NORMAL + (L- len(Text) - 1)*'=' + Style.RESET_ALL)
def Line(L=100, C = 'Blue'):
FORE = {'Black': Fore.BLACK, 'Red':Fore.RED, 'Green':Fore.GREEN, 'Yellow':Fore.YELLOW, 'Blue':Fore.BLUE,
'Magenta':Fore.MAGENTA, 'Cyan':Fore.CYAN, 'White': Fore.WHITE}
print(FORE[C] + Style.NORMAL + L*'=' + Style.RESET_ALL)
def Search_List(Key, List): return [s for s in List if Key in s]
PATH = '1c_software_dataset'
Files = os.listdir(PATH)
Temp = np.sort(Files)
Files = set()
keywords = ['mod']
for key in keywords:
Files = Files.union(set(Search_List(key, Temp)))
del Temp, key, keywords
Files = list(Files)
#
Files_Info = pd.DataFrame()
for i in range(len(Files)):
# Read files
Header(Files[i])
filename = Files[i].split('.')[0]
globals() [filename] = pd.read_csv(os.path.join(PATH, Files[i]))
Temp = globals() [filename].shape
Files_Info = Files_Info.append(pd.DataFrame({'File':[Files[i]], 'Number of Instances': [Temp[0]],
'Number of Attributes': [Temp[1]]}), ignore_index = True)
# Datetime
Cols = globals() [filename].columns
DateTime = Search_List('date', Cols)
if len(DateTime)>0:
try: DateTime.remove('date_block_num')
except: pass
for c in DateTime:
globals() [filename][c] = pd.to_datetime(globals() [filename][c])
del c
# Display
display(globals() [filename].head(5))
Data_fields(globals() [filename])
Line()
display(Files_Info.style.hide_index())
Line()
del Files, i, Temp, filename, Cols, DateTime
```
# Train and Test Datasets
## Test Dataset
```
Test = test_mod.copy()
Test.drop(columns = ['ID'], inplace = True)
Test['Date Block Number'] = sales_train_mod['Date Block Number'].max() + 1
# Test['Set'] = 'Test'
```
## Train Dataset
We only keep the shop and item IDs that we can find in the test Dataset. Also, we will be zero extending all shops and all items to each **Data Block Number**.
```
Mat = []
Features = ['Date Block Number', 'Shop ID', 'Item ID']
Data = sales_train_mod[Features]
DBN = sorted(Data['Date Block Number'].unique().tolist())
Counter = 0
Progress_Bar = progressbar.ProgressBar(maxval= max(DBN), widgets=[progressbar.Bar('=', '|', '|'), progressbar.Percentage()])
Progress_Bar.start()
for number in DBN:
Temp = Data[ Data['Date Block Number'] == number]
Mat.append(np.array(list(itertools.product([number], Temp['Shop ID'].unique(), Temp['Item ID'].unique()))))
Progress_Bar.update(number)
del Temp
Progress_Bar.finish()
del Progress_Bar, DBN, Counter
# We have created a list of matrices
Data = pd.DataFrame(np.vstack(Mat), columns = Features)
del Mat
Feat = 'Item Count Day'
### Item Sold each month
Group = sales_train_mod.groupby(Features, as_index=False)[Feat].sum().astype(int).\
rename(columns = {Feat:Feat.replace('Day','').strip()})
Data = Data.merge(Group, on=Features, how='left').fillna(0).astype(int)
# Data['Set'] = 'Train'
```
Convert to a dask dataframe:
```
Data = dd.from_pandas(Data, npartitions=4)
```
To develop a predictive model, we need to add values from the test set to the train set.
```
Data = dd.concat([Data, Test]).fillna(0)
Data['Item Count'] = Data['Item Count'].astype(int)
```
Adding more features
```
# Shops
Data = Data.merge(shops_mod[['Shop ID','City ID']].drop_duplicates().sort_values(by = ['Shop ID','City ID']),
on='Shop ID', how='left')
# Items
Data = Data.merge(items_mod[['Item ID', 'Item Category Full ID']], on='Item ID', how='left')
# Item Categories
Data = Data.merge(item_categories_mod[['Item Category Full ID', 'Item Category ID',
'Item Subcategory ID']], on='Item Category Full ID', how='left')
```
The following function helps to calculate Lag for various features.
```
def Add_Feature_Lags(df, Lags, Feat):
Subset = df[['Date Block Number', 'Shop ID', 'Item ID', Feat]]
for lag in Lags:
Temp = Subset.copy()
Temp.columns = ['Date Block Number', 'Shop ID', 'Item ID', Feat +' Lag '+str(lag)]
Temp['Date Block Number'] += lag
df = df.merge(Temp, on=['Date Block Number', 'Shop ID', 'Item ID'], how='left')
del Temp
return df
```
Lags for ***Item Count***
```
# One months, two months, three months, six months and a year
Data = Add_Feature_Lags(Data, [1, 2, 3, 6, 12], 'Item Count')
```
### Average Item Sold by Date Block and Item ID
```
Feat = 'Average Item Sold by Date Block and Item ID'
Group = Data.groupby(['Date Block Number', 'Item ID'])['Item Count'].mean().round(2).reset_index()
Group = Group.rename(columns = {'Item Count': Feat})
Data = Data.merge(Group, on=['Date Block Number', 'Item ID'], how='left')
Data = Add_Feature_Lags(Data, [1], Feat)
```
### Average Item Sold by Date Block and Shop ID
```
Feat = 'Average Item Sold by Date Block and Shop ID'
Group = Data.groupby(['Date Block Number', 'Shop ID'])['Item Count'].mean().round(2).reset_index()
Group = Group.rename(columns = {'Item Count': Feat})
Data = Data.merge(Group, on=['Date Block Number', 'Shop ID'], how='left')
Data = Add_Feature_Lags(Data, [1], Feat)
```
### Average Item Sold by Date Block and Item Category ID
```
Feat = 'Average Item Sold by Date Block and Item Category ID'
Group = Data.groupby(['Date Block Number', 'Item Category ID'])['Item Count'].mean().round(2).reset_index()
Group = Group.rename(columns = {'Item Count': Feat})
Data = Data.merge(Group, on=['Date Block Number', 'Item Category ID'], how='left')
Data = Add_Feature_Lags(Data, [1], Feat)
```
### Average Item Sold by Date Block and Item Subcategory ID
```
Feat = 'Average Item Sold by Date Block and Item Subcategory ID'
Group = Data.groupby(['Date Block Number', 'Item Subcategory ID'])['Item Count'].mean().round(2).reset_index()
Group = Group.rename(columns = {'Item Count': Feat})
Data = Data.merge(Group, on=['Date Block Number', 'Item Subcategory ID'], how='left')
Data = Add_Feature_Lags(Data, [1], Feat)
```
### Month
```
Data['Month'] = Data['Date Block Number'] % 12
```
# Modeling
Note that the size of the generated dataset is quite large!
```
Data = Data.compute()
Indeces = {}
Indeces['Train'] = Data[Data['Date Block Number'] < 33].index
Indeces['Validation'] = Data[Data['Date Block Number'] == 33].index
Indeces['Test'] = Data[Data['Date Block Number'] == 34].index
```
## Sets
```
X = Data.drop(['Item Count'], axis=1).fillna(0)
y = Data['Item Count'].fillna(0)
# Train Sets
X_train = X.loc[X.index.isin(Indeces['Train'])]
y_train = y.loc[y.index.isin(Indeces['Train'])]
# Validation Sets
X_valid = X.loc[X.index.isin(Indeces['Validation'])]
y_valid = y.loc[y.index.isin(Indeces['Validation'])]
# Test sets
X_test = X.loc[X.index.isin(Indeces['Test'])]
X_test = Test.merge(X_test, on = Test.columns.tolist(), how = 'left')
```
scaling
```
scaler = StandardScaler()
_ = scaler.fit(X)
xColumns = X.columns
X_train = pd.DataFrame(data = scaler.transform(X_train), columns= xColumns)
X_valid = pd.DataFrame(data = scaler.transform(X_valid), columns= xColumns)
X_test = pd.DataFrame(data = scaler.transform(X_test), columns= xColumns)
```
<div class="alert alert-block alert-danger">
<font size="+2"><b>
Modeling: CatBoost Regressor
</b></font>
</div>
[CatBoost AI](https://catboost.ai/) is based on gradient boosted decision trees. During training, a set of decision trees is built consecutively. Each successive tree is built with reduced loss compared to the previous trees.
```
N = int(5e4)
model = CatBoostRegressor(iterations= N, task_type="GPU", devices='0:1', max_ctr_complexity=5,
random_seed= 0, od_type='Iter', od_wait=N, verbose=int(N/10), depth=5)
_ = model.fit(X_train, y_train, eval_set=(X_valid, y_valid))
# clear_output()
feat_importance = pd.DataFrame(list(zip(X_train.dtypes.index,
model.get_feature_importance(Pool(X_train, label=y_train)))),
columns=['Features','Weight']).sort_values('Weight', ascending = False)
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=(10, 10))
_ = sns.barplot(y="Features", x="Weight", palette="PuRd", edgecolor = 'RoyalBlue', hatch = '///',
data = feat_importance, ax = ax)
_ = sns.barplot(y="Features", x="Weight", facecolor = 'None', edgecolor = 'Black', data = feat_importance, ax = ax)
_ = ax.set_xlim([0,40])
_ = ax.set_title('Feature Importance')
model.plot_tree(tree_idx=0, pool=Pool(X_train, label=y_train))
```
The best result for each metric calculated on each validation dataset.
```
display(pd.DataFrame(model.get_best_score()))
```
R2 Score
```
display(pd.DataFrame({'Train Set': {'R2 Score': model.score(X_train, y_train)},
'Validation Set': {'R2 Score': model.score(X_valid, y_valid)}}))
```
# Predictions
```
Sub = pd.DataFrame({"ID": Test.index.tolist(), "item_cnt_month": model.predict(X_test)})
```
## Saving to a File
```
Sub.to_csv(os.path.join(PATH, 'Catboost_sub.csv'), index=False)
```
***
# References
1. [Kaggle Dataset: Predict Future Sales](https://www.kaggle.com/c/competitive-data-science-predict-future-sales/overview)
***
| github_jupyter |
## GANs
```
%matplotlib inline
from fastai.gen_doc.nbdoc import *
from fastai.vision import *
from fastai.vision.gan import *
```
GAN stands for [Generative Adversarial Nets](https://arxiv.org/pdf/1406.2661.pdf) and were invented by Ian Goodfellow. The concept is that we will train two models at the same time: a generator and a critic. The generator will try to make new images similar to the ones in our dataset, and the critic's job will try to classify real images from the fake ones the generator does. The generator returns images, the discriminator a feature map (it can be a single number depending on the input size). Usually the discriminator will be trained to return 0. everywhere for fake images and 1. everywhere for real ones.
This module contains all the necessary function to create a GAN.
We train them against each other in the sense that at each step (more or less), we:
1. Freeze the generator and train the discriminator for one step by:
- getting one batch of true images (let's call that `real`)
- generating one batch of fake images (let's call that `fake`)
- have the discriminator evaluate each batch and compute a loss function from that; the important part is that it rewards positively the detection of real images and penalizes the fake ones
- update the weights of the discriminator with the gradients of this loss
2. Freeze the discriminator and train the generator for one step by:
- generating one batch of fake images
- evaluate the discriminator on it
- return a loss that rewards posisitivly the discriminator thinking those are real images; the important part is that it rewards positively the detection of real images and penalizes the fake ones
- update the weights of the generator with the gradients of this loss
```
show_doc(GANLearner)
```
This is the general constructor to create a GAN, you might want to use one of the factory methods that are easier to use. Create a GAN from [`data`](/vision.data.html#vision.data), a `generator` and a `critic`. The [`data`](/vision.data.html#vision.data) should have the inputs the `generator` will expect and the images wanted as targets.
`gen_loss_func` is the loss function that will be applied to the `generator`. It takes three argument `fake_pred`, `target`, `output` and should return a rank 0 tensor. `output` is the result of the `generator` applied to the input (the xs of the batch), `target` is the ys of the batch and `fake_pred` is the result of the `discriminator` being given `output`. `output`and `target` can be used to add a specific loss to the GAN loss (pixel loss, feature loss) and for a good training of the gan, the loss should encourage `fake_pred` to be as close to 1 as possible (the `generator` is trained to fool the `critic`).
`crit_loss_func` is the loss function that will be applied to the `critic`. It takes two arguments `real_pred` and `fake_pred`. `real_pred` is the result of the `critic` on the target images (the ys of the batch) and `fake_pred` is the result of the `critic` applied on a batch of fake, generated byt the `generator` from the xs of the batch.
`switcher` is a [`Callback`](/callback.html#Callback) that should tell the GAN when to switch from critic to generator and vice versa. By default it does 5 iterations of the critic for 1 iteration of the generator. The model begins the training with the `generator` if `gen_first=True`. If `switch_eval=True`, the model that isn't trained is switched on eval mode (left in training mode otherwise, which means some statistics like the running mean in batchnorm layers are updated, or the dropouts are applied).
`clip` should be set to a certain value if one wants to clip the weights (see the [Wassertein GAN](https://arxiv.org/pdf/1701.07875.pdf) for instance).
If `show_img=True`, one image generated by the GAN is shown at the end of each epoch.
### Factory methods
```
show_doc(GANLearner.from_learners)
```
Directly creates a [`GANLearner`](/vision.gan.html#GANLearner) from two [`Learner`](/basic_train.html#Learner): one for the `generator` and one for the `critic`. The `switcher` and all `kwargs` will be passed to the initialization of [`GANLearner`](/vision.gan.html#GANLearner) along with the following loss functions:
- `loss_func_crit` is the mean of `learn_crit.loss_func` applied to `real_pred` and a target of ones with `learn_crit.loss_func` applied to `fake_pred` and a target of zeros
- `loss_func_gen` is the mean of `learn_crit.loss_func` applied to `fake_pred` and a target of ones (to full the discriminator) with `learn_gen.loss_func` applied to `output` and `target`. The weights of each of those contributions can be passed in `weights_gen` (default is 1. and 1.)
```
show_doc(GANLearner.wgan)
```
The Wasserstein GAN is detailed in [this article]. `switcher` and the `kwargs` will be passed to the [`GANLearner`](/vision.gan.html#GANLearner) init, `clip`is the weight clipping.
## Switchers
In any GAN training, you will need to tell the [`Learner`](/basic_train.html#Learner) when to switch from generator to critic and vice versa. The two following [`Callback`](/callback.html#Callback) are examples to help you with that.
As usual, don't call the `on_something` methods directly, the fastai library will do it for you during training.
```
show_doc(FixedGANSwitcher, title_level=3)
show_doc(FixedGANSwitcher.on_train_begin)
show_doc(FixedGANSwitcher.on_batch_end)
show_doc(AdaptiveGANSwitcher, title_level=3)
show_doc(AdaptiveGANSwitcher.on_batch_end)
```
## Discriminative LR
If you want to train your critic at a different learning rate than the generator, this will let you do it automatically (even if you have a learning rate schedule).
```
show_doc(GANDiscriminativeLR, title_level=3)
show_doc(GANDiscriminativeLR.on_batch_begin)
show_doc(GANDiscriminativeLR.on_step_end)
```
## Specific models
```
show_doc(basic_critic)
```
This model contains a first 4 by 4 convolutional layer of stride 2 from `n_channels` to `n_features` followed by `n_extra_layers` 3 by 3 convolutional layer of stride 1. Then we put as many 4 by 4 convolutional layer of stride 2 with a number of features multiplied by 2 at each stage so that the `in_size` becomes 1. `kwargs` can be used to customize the convolutional layers and are passed to [`conv_layer`](/layers.html#conv_layer).
```
show_doc(basic_generator)
```
This model contains a first 4 by 4 transposed convolutional layer of stride 1 from `noise_size` to the last numbers of features of the corresponding critic. Then we put as many 4 by 4 transposed convolutional layer of stride 2 with a number of features divided by 2 at each stage so that the image ends up being of height and widht `in_size//2`. At the end, we add`n_extra_layers` 3 by 3 convolutional layer of stride 1. The last layer is a transpose convolution of size 4 by 4 and stride 2 followed by `tanh`. `kwargs` can be used to customize the convolutional layers and are passed to [`conv_layer`](/layers.html#conv_layer).
```
show_doc(gan_critic)
show_doc(GANTrainer)
```
[`LearnerCallback`](/basic_train.html#LearnerCallback) that will be responsible to handle the two different optimizers (one for the generator and one for the critic), and do all the work behind the scenes so that the generator (or the critic) are in training mode with parameters requirement gradients each time we switch.
`switch_eval=True` means that the [`GANTrainer`](/vision.gan.html#GANTrainer) will put the model that isn't training into eval mode (if it's `False` its running statistics like in batchnorm layers will be updated and dropout will be applied). `clip` is the clipping applied to the weights (if not `None`). `beta` is the coefficient for the moving averages as the [`GANTrainer`](/vision.gan.html#GANTrainer)tracks separately the generator loss and the critic loss. `gen_first=True` means the training begins with the generator (with the critic if it's `False`). If `show_img=True` we show a generated image at the end of each epoch.
```
show_doc(GANTrainer.switch)
```
If `gen_mode` is left as `None`, just put the model in the other mode (critic if it was in generator mode and vice versa).
```
show_doc(GANTrainer.on_train_begin)
show_doc(GANTrainer.on_epoch_begin)
show_doc(GANTrainer.on_batch_begin)
show_doc(GANTrainer.on_backward_begin)
show_doc(GANTrainer.on_epoch_end)
show_doc(GANTrainer.on_train_end)
```
## Specific modules
```
show_doc(GANModule, title_level=3)
```
If `gen_mode` is left as `None`, just put the model in the other mode (critic if it was in generator mode and vice versa).
```
show_doc(GANModule.switch)
show_doc(GANLoss, title_level=3)
show_doc(AdaptiveLoss, title_level=3)
show_doc(accuracy_thresh_expand)
```
## Data Block API
```
show_doc(NoisyItem, title_level=3)
show_doc(GANItemList, title_level=3)
```
Inputs will be [`NoisyItem`](/vision.gan.html#NoisyItem) of `noise_sz` while the default class for target is [`ImageList`](/vision.data.html#ImageList).
```
show_doc(GANItemList.show_xys)
show_doc(GANItemList.show_xyzs)
```
## Undocumented Methods - Methods moved below this line will intentionally be hidden
```
show_doc(GANLoss.critic)
show_doc(GANModule.forward)
show_doc(GANLoss.generator)
show_doc(NoisyItem.apply_tfms)
show_doc(AdaptiveLoss.forward)
show_doc(GANItemList.get)
show_doc(GANItemList.reconstruct)
show_doc(AdaptiveLoss.forward)
```
## New Methods - Please document or move to the undocumented section
| github_jupyter |
## Building a dashboard to plan a marketing campaign leveraging CARTO Data Observatory
Combining different data sources to identify some patterns or understand some behavior in a specific location is a very typical use case in Spatial Data Science.
In this notebook, we will build a dashboard combining different data from CARTO's Data Observatory to help identify the locations with specific characteristics described below.
**Note:** This use case leverages premium datasets from [CARTO's Data Observatory](https://carto.com/spatial-data-catalog/).
### Use case description
A pharmaceutical lab wants to launch a new marketing campaign to promote a new line of personal care products for senior people in the city of Philadelphia, PA. They know their target group is characterized by:
- People over 60
- Medium-high to high income
- High expenditure in personal care products and services
Given these characteristics, they would like to know which pharmacies and drug stores in the city of Philadelphia they should focus their efforts on.
In order to identify the target drug stores and pharmacies, we will follow the following steps:
- [Get all pharmacies in Philadelphia](#section1)
- [Calculate their cathment areas using isochrones](#section2)
- [Enrich the isochrones with demographic, POI's, and consumption data](#section3)
- [Build the dashboard to help identify the pharmacies where the campaign can be more successful given the characteristics of the population within their catchment area](#section4)
### 0. Setup
Import the packages we'll use.
```
import geopandas as gpd
import pandas as pd
from cartoframes.auth import set_default_credentials
from cartoframes.data.services import Isolines
from cartoframes.data.observatory import *
from cartoframes.viz import *
from shapely.geometry import box
pd.set_option('display.max_columns', None)
```
In order to be able to use the Data Observatory via CARTOframes, you need to set your CARTO account credentials first.
Please, visit the [Authentication guide](https://carto.com/developers/cartoframes/guides/Authentication/) for further detail.
```
from cartoframes.auth import set_default_credentials
set_default_credentials('creds.json')
```
**Note about credentials**
For security reasons, we recommend storing your credentials in an external file to prevent publishing them by accident when sharing your notebooks. You can get more information in the section _Setting your credentials_ of the [Authentication guide](https://carto.com/developers/cartoframes/guides/Authentication/).
<a id='section1'></a>
### 1. Download all pharmacies in Philadelphia from the Data Observatory
Below is the bounding box of the area of study.
```
dem_bbox = box(-75.229353,39.885501,-75.061124,39.997898)
```
We can get the pharmacies from [Pitney Bowes' Consumer Points of Interest](https://carto.com/spatial-data-catalog/browser/dataset/pb_consumer_po_62cddc04/) dataset. This is a premium dataset, so we first need to check that we are subscribed to it.
Take a look at <a href='#example-access-premium-data-from-the-data-observatory' target='_blank'>this template</a> for more details on how to access and download a premium dataset.
```
Catalog().subscriptions().datasets.to_dataframe()
```
#### Download and explore sample
Pitney Bowes POI's are hierarchically classified (levels: trade division, group, class, sub class).
Since we might not know which level can help us identify all pharmacies, we can start by downloading a sample for a smaller area to explore the dataset. For calculating the bounding box we use [bboxfinder](http://bboxfinder.com).
We start by selecting our dataset and taking a quick look at its first 10 rows.
```
dataset = Dataset.get('pb_consumer_po_62cddc04')
dataset.head()
```
Let's now download a small sample to help us identify which of the four hierarchy variables gives us the pharmacies.
```
sql_query = "SELECT * except(do_label) FROM $dataset$ WHERE ST_IntersectsBox(geom, -75.161723,39.962019,-75.149535,39.968071)"
sample = dataset.to_dataframe(sql_query=sql_query)
sample.head()
sample['TRADE_DIVISION'].unique()
sample.loc[sample['TRADE_DIVISION'] == 'DIVISION G. - RETAIL TRADE', 'GROUP'].unique()
sample.loc[sample['TRADE_DIVISION'] == 'DIVISION G. - RETAIL TRADE', 'CLASS'].unique()
```
The class `DRUG STORES AND PROPRIETARY STORES` is the one we're looking for.
```
sample.loc[sample['CLASS'] == 'DRUG STORES AND PROPRIETARY STORES', 'SUB_CLASS'].unique()
```
#### Download all pharmacies in the area of study
```
sql_query = """SELECT * except(do_label)
FROM $dataset$
WHERE CLASS = 'DRUG STORES AND PROPRIETARY STORES'
AND ST_IntersectsBox(geom, -75.229353,39.885501,-75.061124,39.997898)"""
ph_pharmacies = dataset.to_dataframe(sql_query=sql_query)
ph_pharmacies.head()
```
The dataset contains different versions of the POI's tagged by the _do_date_ column. We are only inetrested in the latest version of each POI.
```
ph_pharmacies = ph_pharmacies.sort_values(by='do_date', ascending=False).groupby('PB_ID').first().reset_index()
ph_pharmacies.shape
```
#### Visualize the dataset
```
Layer(ph_pharmacies,
geom_col='geom',
style=basic_style(opacity=0.75),
popup_hover=popup_element('NAME'))
```
<a id='section2'></a>
### 2. Calculate catchment areas
In order to know the characteristics of the potential customers of every pharmacy, we assume the majority of their clients live closeby. Therefore we will calculate **5-minute-by-car isochrones** and take them as their cathment areas.
_Note_ catchment areas usually depend on whether it is a store in the downtown area or in the suburbs, or if it is reachable on foot or only by car. For this example, we will not make such distiction between pharmacies, but we strongly encourage you to do so on your analyses. As an example, [here](https://carto.com/blog/calculating-catchment-human-mobility-data/) we describe how to calculate catchment areas using human mobility data.
```
iso_service = Isolines()
isochrones_gdf, _ = iso_service.isochrones(ph_pharmacies, [300], mode='car', geom_col='geom')
ph_pharmacies['iso_5car'] = isochrones_gdf.sort_values(by='source_id')['the_geom'].values
```
#### Visualize isochrones
We'll only visualize the ten first isochrones to get a clean visualization.
```
Map([Layer(ph_pharmacies.iloc[:10],
geom_col='iso_5car',
style=basic_style(opacity=0.1),
legends=basic_legend('Catchment Areas')),
Layer(ph_pharmacies.iloc[:10],
geom_col='geom',
popup_hover=popup_element('NAME'),
legends=basic_legend('Pharmacies'))])
```
<a id='section3'></a>
### 3. Enrichment: Chacacterize catchment areas
We'll now enrich the pharmacies catchment areas with demographics, POI's, and consumer spending data.
For the enrichment, we will use the CARTOframes Enrichment class. This class contains the functionality to enrich polygons and points.
Visit [CARTOframes Guides](https://carto.com/developers/cartoframes/guides/) for further detail.
```
enrichment = Enrichment()
```
#### Demographics
We will use AGS premium data. In particular, we will work with the dataset `ags_sociodemogr_f510a947` which contains [yearly demographics data from 2019](https://carto.com/spatial-data-catalog/browser/dataset/ags_sociodemogr_f510a947/).
##### Variable selection
Here we will enrich the pharmacies isochrones with:
- Population aged 60+
- Household income
- Household income for population ages 65+
```
Catalog().country('usa').category('demographics').provider('ags').datasets.to_dataframe().head()
dataset = Dataset.get('ags_sociodemogr_f510a947')
dataset.head()
```
We explore the variables to identify the ones we're interested in.
Variables in a dataset are uniquely identified by their slug.
```
dataset.variables.to_dataframe().head()
```
We'll select:
- Population and population by age variables to identify number of people aged 60+ as a percentage of total population
- Average household income
- Average household income for porpulation aged 65+
```
vars_enrichment = ['POPCY_5e23b8f4', 'AGECY6064_d54c2315', 'AGECY6569_ad369d43', 'AGECY7074_74eb7531',
'AGECY7579_c91cb67', 'AGECY8084_ab1079a8', 'AGECYGT85_a0959a08', 'INCCYMEDHH_b80a7a7b',
'HINCYMED65_37a430a4', 'HINCYMED75_2ebf01e5']
```
##### Isochrone enrichment
```
ph_pharmacies_enriched = enrichment.enrich_polygons(
ph_pharmacies,
variables=vars_enrichment,
geom_col='iso_5car'
)
ph_pharmacies_enriched.head()
ph_pharmacies = ph_pharmacies_enriched.copy()
ph_pharmacies['pop_60plus'] = ph_pharmacies[['AGECY8084', 'AGECYGT85', 'AGECY6569', 'AGECY7579', 'AGECY7074', 'AGECY6064']].sum(1)
ph_pharmacies.drop(columns=['AGECY8084', 'AGECYGT85', 'AGECY6569', 'AGECY7579', 'AGECY7074', 'AGECY6064'], inplace=True)
```
#### Points of Interest
We will use [Pitney Bowes' Consumer Points of Interest](https://carto.com/spatial-data-catalog/browser/dataset/pb_consumer_po_62cddc04/) premium dataset.
##### Variable selection
We are interested in knowing how many of the following POIs can be found in each isochrone:
- Beauty shops and beauty salons
- Gyms and other sports centers
These POI's will be considered as an indicator of personal care awareness in a specific area.
The hierarchy classification variable `SUB_CLASS` variable allows us to identify beaty shops and salons (`BEAUTY SHOPS/BEAUTY SALON`) and gyms (`MEMBERSHIP SPORTS AND RECREATION CLUBS/CLUB AND ASSOCIATION - UNSPECIFIED`).
```
sample.loc[sample['TRADE_DIVISION'] == 'DIVISION I. - SERVICES', 'SUB_CLASS'].unique()
```
##### Isochrone enrichment
In order to count only Beauty Shops/Salons and Gyms, we will apply a filter to the enrichment. All filters are applied with an AND-like relationship. This means we need to run two independent enrichment calls, one for the beauty shops/salons and another one for the gyms.
```
ph_pharmacies_enriched = enrichment.enrich_polygons(
ph_pharmacies,
variables=['SUB_CLASS_10243439'],
aggregation='COUNT',
geom_col='iso_5car',
filters={Variable.get('SUB_CLASS_10243439').id : "= 'BEAUTY SHOPS/BEAUTY SALON'"}
)
ph_pharmacies = ph_pharmacies_enriched.rename(columns={'SUB_CLASS_y':'n_beauty_pois'})
ph_pharmacies_enriched = enrichment.enrich_polygons(
ph_pharmacies,
variables=['SUB_CLASS_10243439'],
aggregation='COUNT',
geom_col='iso_5car',
filters={Variable.get('SUB_CLASS_10243439').id : "= 'MEMBERSHIP SPORTS AND RECREATION CLUBS/CLUB AND ASSOCIATION - UNSPECIFIED'"}
)
ph_pharmacies = ph_pharmacies_enriched.rename(columns={'SUB_CLASS':'n_gym_pois'})
ph_pharmacies['n_pois_personal_care'] = ph_pharmacies['n_beauty_pois'] + ph_pharmacies['n_gym_pois']
ph_pharmacies.drop(columns=['n_beauty_pois', 'n_gym_pois'], inplace=True)
```
#### Consumer spending
For consumer spending, we will use AGS premium data. In particular, we will work with the dataset `ags_consumer_sp_dbabddfb` which contains the [latest version of yearly consumer data](https://carto.com/spatial-data-catalog/browser/dataset/ags_consumer_sp_dbabddfb/).
##### Variable selection
We are interested in spending in:
- Personal care services
- Personal care products
- Health care services
```
dataset = Dataset.get('ags_consumer_sp_dbabddfb')
dataset.variables.to_dataframe().head()
```
The variables we're interested in are:
- `XCYHC2` Health care services expenditure
- `XCYPC3` Personal care services expenditure
- `XCYPC4` Personal care products expenditure
```
Variable.get('XCYHC2_18141567').to_dict()
ph_pharmacies_enriched = enrichment.enrich_polygons(
ph_pharmacies,
variables=['XCYPC3_7d26d739', 'XCYPC4_e342429a', 'XCYHC2_18141567'],
geom_col='iso_5car'
)
```
We rename the new columns to give them a more descriptive name.
```
ph_pharmacies = ph_pharmacies_enriched.rename(columns={'XCYHC2':'health_care_services_exp',
'XCYPC3':'personal_care_services_exp',
'XCYPC4':'personal_care_products_exp'})
ph_pharmacies.head(2)
```
<a id='section4'></a>
### 4. Dashboard
Finally, with all the data gathered, we will build the dashboard and publish it so we can share it with our client/manager/colleague for them to explore it.
This dashboard allows you to select a range of desired expenditure in care products, people aged 60+, household income, and so forth. Selecting the desired ranges will filter out pharmacies, so that in the end you can identify the target pharmacies for your marketing campaign.
```
cmap = Map(Layer(ph_pharmacies,
geom_col='geom',
style=color_category_style('SIC8_DESCRIPTION', size=4, opacity=0.85, palette='safe', stroke_width=0.15),
widgets=[formula_widget(
'PB_ID',
operation='COUNT',
title='Total number of pharmacies',
description='Keep track of the total amount of pharmacies that meet the ranges selected on the widgets below'),
histogram_widget(
'pop_60plus',
title='Population 60+',
description='Select a range of values to filter',
buckets=15
),
histogram_widget(
'HINCYMED65',
title='Household income 65-74',
buckets=15
),
histogram_widget(
'HINCYMED75',
title='Household income 75+',
buckets=15
),
histogram_widget(
'n_pois_personal_care',
title='Number of personal care POIs',
buckets=15
),
histogram_widget(
'personal_care_products_exp',
title='Expenditure in personal care products ($)',
buckets=15
)],
legends=color_category_legend(
title='Pharmacies',
description='Type of store'),
popup_hover=[popup_element('NAME', title='Name')]
),
viewport={'zoom': 11}
)
cmap
```
#### Publish dashboard
```
cmap.publish('ph_pharmacies_dashboard', password='MY_PASS', if_exists='replace')
```
| github_jupyter |
# Assignment 4: Chatbot
<img src = "cbot.jpg" height="400" width="400">
Welcome to the last assignment of Course 4. Before you get started, we want to congratulate you on getting here. It is your 16th programming assignment in this Specialization and we are very proud of you! In this assignment, you are going to use the [Reformer](https://arxiv.org/abs/2001.04451), also known as the efficient Transformer, to generate a dialogue between two bots. You will feed conversations to your model and it will learn how to understand the context of each one. Not only will it learn how to answer questions but it will also know how to ask questions if it needs more info. For example, after a customer asks for a train ticket, the chatbot can ask what time the said customer wants to leave. You can use this concept to automate call centers, hotel receptions, personal trainers, or any type of customer service. By completing this assignment, you will:
* Understand how the Reformer works
* Explore the [MultiWoz](https://arxiv.org/abs/1810.00278) dataset
* Process the data to feed it into the model
* Train your model
* Generate a dialogue by feeding a question to the model
## Outline
- [Part 1: Exploring the MultiWoz dataset](#1)
- [Exercise 01](#ex01)
- [Part 2: Processing the data for Reformer inputs](#2)
- [2.1 Tokenizing, batching with bucketing](#2.1)
- [Part 3: Reversible layers](#3)
- [Exercise 02](#ex02)
- [Exercise 03](#ex03)
- [3.1 Reversible layers and randomness](#3.1)
- [Part 4: ReformerLM Training](#4)
- [Exercise 04](#ex04)
- [Exercise 05](#ex05)
- [Part 5: Decode from a pretrained model](#5)
- [Exercise 06](#ex06)
<a name="1"></a>
# Part 1: Exploring the MultiWoz dataset
You will start by exploring the MultiWoz dataset. The dataset you are about to use has more than 10,000 human annotated dialogues and spans multiple domains and topics. Some dialogues include multiple domains and others include single domains. In this section, you will load and explore this dataset, as well as develop a function to extract the dialogues.
Let's first import the modules we will be using:
```
import json
import random
import numpy as np
from termcolor import colored
import trax
from trax import layers as tl
from trax.supervised import training
!pip list | grep trax
```
Let's also declare some constants we will be using in the exercises.
```
# filename of the MultiWOZ dialogue dataset
DATA_FILE = 'data.json'
# data directory
DATA_DIR = './data'
# dictionary where we will load the dialogue dataset
DIALOGUE_DB = {}
# vocabulary filename
VOCAB_FILE = 'en_32k.subword'
# vocabulary file directory
VOCAB_DIR = 'data/vocabs'
```
Let's now load the MultiWOZ 2.1 dataset. We have already provided it for you in your workspace. It is in JSON format so we should load it as such:
```
# help function to load a JSON file
def load_json(directory, file):
with open(f'{directory}/{file}') as file:
db = json.load(file)
return db
# load the dialogue data set into our dictionary
DIALOGUE_DB = load_json(DATA_DIR, DATA_FILE)
```
Let's see how many dialogues we have in the dictionary. 1 key-value pair is one dialogue so we can just get the dictionary's length.
```
print(f'The number of dialogues is: {len(DIALOGUE_DB)}')
```
The dialogues are composed of multiple files and the filenames are used as keys in our dictionary. Those with multi-domain dialogues have "MUL" in their filenames while single domain dialogues have either "SNG" or "WOZ".
```
# print 7 keys from the dataset to see the filenames
print(list(DIALOGUE_DB.keys())[0:7])
```
As you can see from the cells above, there are 10,438 conversations, each in its own file. You will train your model on all those conversations. Each file is also loaded into a dictionary and each has two keys which are the following:
```
# get keys of the fifth file in the list above
print(DIALOGUE_DB['SNG0073.json'].keys())
```
The `goal` also points to a dictionary and it contains several keys pertaining to the objectives of the conversation. For example below, we can see that the conversation will be about booking a taxi.
```
DIALOGUE_DB['SNG0073.json']['goal']
```
The `log` on the other hand contains the dialog. It is a list of dictionaries and each element of this list contains several descriptions as well. Let's look at an example:
```
# get first element of the log list
DIALOGUE_DB['SNG0073.json']['log'][0]
```
For this assignment, we are only interested in the conversation which is in the `text` field.
The conversation goes back and forth between two persons. Let's call them 'Person 1' and 'Person 2'. This implies that
data['SNG0073.json']['log'][0]['text'] is 'Person 1' and
data['SNG0073.json']['log'][1]['text'] is 'Person 2' and so on. The even offsets are 'Person 1' and the odd offsets are 'Person 2'.
```
print(' Person 1: ', DIALOGUE_DB['SNG0073.json']['log'][0]['text'])
print(' Person 2: ',DIALOGUE_DB['SNG0073.json']['log'][1]['text'])
```
<a name="ex01"></a>
### Exercise 01
You will now implement the `get_conversation()` function that will extract the conversations from the dataset's file.
**Instructions:** Implement a function to extract conversations from the input file.
As described above, the conversation is in the `text` field in each of the elements in the `log` list of the file. If the log list has `x` number of elements, then the function will get the `text` entries of each of those elements. Your function should return the conversation, prepending each field with either ' Person 1: ' if 'x' is even or ' Person 2: ' if 'x' is odd. You can use the Python modulus operator '%' to help select the even/odd entries. Important note: Do not print a newline character (i.e. `\n`) when generating the string. For example, in the code cell above, your function should output something like:
```
Person 1: I would like a taxi from Saint John's college to Pizza Hut Fen Ditton. Person 2: What time do you want to leave and what time do you want to arrive by?
```
and **not**:
```
Person 1: I would like a taxi from Saint John's college to Pizza Hut Fen Ditton.
Person 2: What time do you want to leave and what time do you want to arrive by?
```
```
# UNQ_C1
# GRADED FUNCTION: get_conversation
def get_conversation(file, data_db):
'''
Args:
file (string): filename of the dialogue file saved as json
data_db (dict): dialogue database
Returns:
string: A string containing the 'text' fields of data[file]['log'][x]
'''
# initialize empty string
result = ''
# get length of file's log list
len_msg_log = len(data_db[file]['log'])
# set the delimiter strings
delimiter_1 = ' Person 1: '
delimiter_2 = ' Person 2: '
# loop over the file's log list
for i in range(len_msg_log):
### START CODE HERE (REPLACE INSTANCES OF 'None' WITH YOUR CODE) ###
# get i'th element of file log list
cur_log = data_db[file]['log'][i]
# check if i is even
if i % 2 == 0:
# append the 1st delimiter string
result += delimiter_1
else:
# append the 2nd delimiter string
result += delimiter_2
# append the message text from the log
result += cur_log['text']
### END CODE HERE ###
return result
# BEGIN UNIT TEST
import w4_unittest
w4_unittest.test_get_conversation(get_conversation)
# END UNIT TEST
file = 'SNG01856.json'
conversation = get_conversation(file, DIALOGUE_DB)
# print raw output
print(conversation)
```
**Expected Result:**
```
Person 1: am looking for a place to to stay that has cheap price range it should be in a type of hotel Person 2: Okay, do you have a specific area you want to stay in? Person 1: no, i just need to make sure it's cheap. oh, and i need parking Person 2: I found 1 cheap hotel for you that includes parking. Do you like me to book it? Person 1: Yes, please. 6 people 3 nights starting on tuesday. Person 2: I am sorry but I wasn't able to book that for you for Tuesday. Is there another day you would like to stay or perhaps a shorter stay? Person 1: how about only 2 nights. Person 2: Booking was successful.
Reference number is : 7GAWK763. Anything else I can do for you? Person 1: No, that will be all. Good bye. Person 2: Thank you for using our services.
```
We can have a utility pretty print function just so we can visually follow the conversation more easily.
```
def print_conversation(conversation):
delimiter_1 = 'Person 1: '
delimiter_2 = 'Person 2: '
split_list_d1 = conversation.split(delimiter_1)
for sublist in split_list_d1[1:]:
split_list_d2 = sublist.split(delimiter_2)
print(colored(f'Person 1: {split_list_d2[0]}', 'red'))
if len(split_list_d2) > 1:
print(colored(f'Person 2: {split_list_d2[1]}', 'green'))
print_conversation(conversation)
```
For this assignment, we will just use the outputs of the calls to `get_conversation` to train the model. But just to expound, there are also other information in the MultiWoz dataset that can be useful in other contexts. Each element of the log list has more information about it. For example, above, if you were to look at the other fields for the following, "am looking for a place to stay that has cheap price range it should be in a type of hotel", you will get the following.
```
DIALOGUE_DB['SNG01856.json']['log'][0]
```
The dataset also comes with hotel, hospital, taxi, train, police, and restaurant databases. For example, in case you need to call a doctor, or a hotel, or a taxi, this will allow you to automate the entire conversation. Take a look at the files accompanying the data set.
```
# this is an example of the attractions file
attraction_file = open('data/attraction_db.json')
attractions = json.load(attraction_file)
print(attractions[0])
# this is an example of the hospital file
hospital_file = open('data/hospital_db.json')
hospitals = json.load(hospital_file)
print(hospitals[0]) # feel free to index into other indices
# this is an example of the hotel file
hotel_file = open('data/hotel_db.json')
hotels = json.load(hotel_file)
print(hotels[0]) # feel free to index into other indices
# this is an example of the police file
police_file = open('data/police_db.json')
police = json.load(police_file)
print(police[0]) # feel free to index into other indices
# this is an example of a restuarant file
restaurant_file = open('data/restaurant_db.json')
restaurants = json.load(restaurant_file)
print(restaurants[0]) # feel free to index into other indices
```
For more information about the multiwoz 2.1 data set, please run the cell below to read the `ReadMe.txt` file. Feel free to open any other file to explore it.
```
with open('data/README') as file:
print(file.read())
```
As you can see, there are many other aspects of the MultiWoz dataset. Nonetheless, you'll see that even with just the conversations, your model will still be able to generate useful responses. This concludes our exploration of the dataset. In the next section, we will do some preprocessing before we feed it into our model for training.
<a name="2"></a>
# Part 2: Processing the data for Reformer inputs
You will now use the `get_conversation()` function to process the data. The Reformer expects inputs of this form:
**Person 1: Why am I so happy? Person 2: Because you are learning NLP Person 1: ... Person 2: ...***
And the conversation keeps going with some text. As you can see 'Person 1' and 'Person 2' act as delimiters so the model automatically recognizes the person and who is talking. It can then come up with the corresponding text responses for each person. Let's proceed to process the text in this fashion for the Reformer. First, let's grab all the conversation strings from all dialogue files and put them in a list.
```
# the keys are the file names
all_files = DIALOGUE_DB.keys()
# initialize empty list
untokenized_data = []
# loop over all files
for file in all_files:
# this is the graded function you coded
# returns a string delimited by Person 1 and Person 2
result = get_conversation(file, DIALOGUE_DB)
# append to the list
untokenized_data.append(result)
# print the first element to check if it's the same as the one we got before
print(untokenized_data[0])
```
Now let us split the list to a train and eval dataset.
```
# shuffle the list we generated above
random.shuffle(untokenized_data)
# define a cutoff (5% of the total length for this assignment)
# convert to int because we will use it as a list index
cut_off = int(len(untokenized_data) * .05)
# slice the list. the last elements after the cut_off value will be the eval set. the rest is for training.
train_data, eval_data = untokenized_data[:-cut_off], untokenized_data[-cut_off:]
print(f'number of conversations in the data set: {len(untokenized_data)}')
print(f'number of conversations in train set: {len(train_data)}')
print(f'number of conversations in eval set: {len(eval_data)}')
```
<a name="2.1"></a>
## 2.1 Tokenizing, batching with bucketing
We can now proceed in generating tokenized batches of our data. Let's first define a utility generator function to yield elements from our data sets:
```
def stream(data):
# loop over the entire data
while True:
# get a random element
d = random.choice(data)
# yield a tuple pair of identical values
# (i.e. our inputs to the model will also be our targets during training)
yield (d, d)
```
Now let's define our data pipeline for tokenizing and batching our data. As in the previous assignments, we will bucket by length and also have an upper bound on the token length.
```
# trax allows us to use combinators to generate our data pipeline
data_pipeline = trax.data.Serial(
# randomize the stream
trax.data.Shuffle(),
# tokenize the data
trax.data.Tokenize(vocab_dir=VOCAB_DIR,
vocab_file=VOCAB_FILE),
# filter too long sequences
trax.data.FilterByLength(2048),
# bucket by length
trax.data.BucketByLength(boundaries=[128, 256, 512, 1024],
batch_sizes=[16, 8, 4, 2, 1]),
# add loss weights but do not add it to the padding tokens (i.e. 0)
trax.data.AddLossWeights(id_to_mask=0)
)
# apply the data pipeline to our train and eval sets
train_stream = data_pipeline(stream(train_data))
eval_stream = data_pipeline(stream(eval_data))
```
Peek into the train stream.
```
# the stream generators will yield (input, target, weights). let's just grab the input for inspection
inp, _, _ = next(train_stream)
# print the shape. format is (batch size, token length)
print("input shape: ", inp.shape)
# detokenize the first element
print(trax.data.detokenize(inp[0], vocab_dir=VOCAB_DIR, vocab_file=VOCAB_FILE))
```
<a name="3"></a>
# Part 3: Reversible layers
When running large deep models, you will often run out of memory as each layer allocates memory to store activations for use in backpropagation. To save this resource, you need to be able to recompute these activations during the backward pass without storing them during the forward pass. Take a look first at the leftmost diagram below.
<img src="reversible2.PNG" height="400" width="600">
This is how the residual networks are implemented in the standard Transformer. It follows that, given `F()` is Attention and `G()` is Feed-forward(FF).
:
\begin{align}
\mathrm{y}_\mathrm{a} &= \mathrm{x} + \mathrm{F}\left(\mathrm{x}\right)\tag{1} \\
\mathrm{y}_{b}&=\mathrm{y}_{a}+\mathrm{G}\left(\mathrm{y}_{a}\right)\tag{2}\\
\end{align}
As you can see, it requires that $\mathrm{x}$ and $\mathrm{y}_{a}$ be saved so it can be used during backpropagation. We want to avoid this to conserve memory and this is where reversible residual connections come in. They are shown in the middle and rightmost diagrams above. The key idea is that we will start with two copies of the input to the model and at each layer we will only update one of them. The activations that we *don’t* update are the ones that will be used to compute the residuals.
Now in this reversible set up you get the following instead:
\begin{align}
\mathrm{y}_{1}&=\mathrm{x}_{1}+\mathrm{F}\left(\mathrm{x}_{2}\right)\tag{3}\\
\mathrm{y}_{2}&=\mathrm{x}_{2}+\mathrm{G}\left(\mathrm{y}_{1}\right)\tag{4}\\
\end{align}
To recover $\mathrm{(x_1,x_2)}$ from $\mathrm{(y_1, y_2)}$
\begin{align}
\mathrm{x}_{2}&=\mathrm{y}_{2}-\mathrm{G}\left(\mathrm{y}_{1}\right)\tag{5}\\
\mathrm{x}_{1}&=\mathrm{y}_{1}-\mathrm{F}\left(\mathrm{x}_{2}\right)\tag{6}\\
\end{align}
With this configuration, we’re now able to run the network fully in reverse. You'll notice that during the backward pass, $\mathrm{x2}$ and $\mathrm{x1}$ can be recomputed based solely on the values of $\mathrm{y2}$ and $\mathrm{y1}$. No need to save it during the forward pass.
<a name="ex02"></a>
### Exercise 02
**Instructions:** You will implement the `reversible_layer_forward` function using equations 3 and 4 above. This function takes in the input vector `x` and the functions `f` and `g` and returns the concatenation of $y_1 and y_2$. For this exercise, we will be splitting `x` before going through the reversible residual steps$\mathrm{^1}$. We can then use those two vectors for the `reversible_layer_reverse` function. Utilize `np.concatenate()` to form the output being careful to match the axis of the `np.split()`.
$\mathrm{^1}$*Take note that this is just for demonstrating the concept in this exercise and there are other ways of processing the input. As you'll see in the Reformer architecture later, the initial input (i.e. `x`) can instead be duplicated instead of split.*
```
# UNQ_C2
# GRADED FUNCTION: reversible_layer_forward
def reversible_layer_forward(x, f, g):
"""
Args:
x (np.array): an input vector or matrix
f (function): a function which operates on a vector/matrix
g (function): a function which operates on a vector/matrix
Returns:
y (np.array): an output vector or matrix whose form is determined by 'x', f and g
"""
# split the input vector into two (* along the last axis because it is the depth dimension)
x1, x2 = np.split(x, 2, axis=-1)
### START CODE HERE (REPLACE INSTANCES OF 'None' WITH YOUR CODE) ###
# get y1 using equation 3
y1 = x1 + f(x2)
# get y2 using equation 4
y2 = x2 + g(y1)
# concatenate y1 and y2 along the depth dimension. be sure output is of type np.ndarray
y = np.concatenate([y1, y2], axis=-1)
### END CODE HERE ###
return y
# BEGIN UNIT TEST
w4_unittest.test_reversible_layer_forward(reversible_layer_forward)
# END UNIT TEST
```
<a name="ex03"></a>
### Exercise 03
You will now implement the `reversible_layer_reverse` function which is possible because at every time step you have $x_1$ and $x_2$ and $y_2$ and $y_1$, along with the function `f`, and `g`. Where `f` is the attention and `g` is the feedforward. This allows you to compute equations 5 and 6.
**Instructions:** Implement the `reversible_layer_reverse`. Your function takes in the output vector from `reversible_layer_forward` and functions f and g. Using equations 5 and 6 above, it computes the inputs to the layer, $x_1$ and $x_2$. The output, x, is the concatenation of $x_1, x_2$. Utilize `np.concatenate()` to form the output being careful to match the axis of the `np.split()`.
```
# UNQ_C3
# GRADED FUNCTION: reversible_layer_reverse
def reversible_layer_reverse(y, f, g):
"""
Args:
y (np.array): an input vector or matrix
f (function): a function which operates on a vector/matrix of the form of 'y'
g (function): a function which operates on a vector/matrix of the form of 'y'
Returns:
y (np.array): an output vector or matrix whose form is determined by 'y', f and g
"""
# split the input vector into two (* along the last axis because it is the depth dimension)
y1, y2 = np.split(y, 2, axis=-1)
### START CODE HERE (REPLACE INSTANCES OF 'None' WITH YOUR CODE) ###
# compute x2 using equation 5
x2 = y2 - g(y1)
# compute x1 using equation 6
x1 = y1 - f(x2)
# concatenate x1 and x2 along the depth dimension
x = np.concatenate([x1, x2], axis=-1)
### END CODE HERE ###
return x
# BEGIN UNIT TEST
w4_unittest.test_reversible_layer_reverse(reversible_layer_reverse)
# END UNIT TEST
# UNIT TEST COMMENT: assert at the end can be used in grading as well
f = lambda x: x + 2
g = lambda x: x * 3
input_vector = np.random.uniform(size=(32,))
output_vector = reversible_layer_forward(input_vector, f, g)
reversed_vector = reversible_layer_reverse(output_vector, f, g)
assert np.allclose(reversed_vector, input_vector)
```
<a name="3.1"></a>
## 3.1 Reversible layers and randomness
This is why we were learning about fastmath's random functions and keys in Course 3 Week 1. Utilizing the same key, `trax.fastmath.random.uniform()` will return the same values. This is required for the backward pass to return the correct layer inputs when random noise is introduced in the layer.
```
# Layers like dropout have noise, so let's simulate it here:
f = lambda x: x + np.random.uniform(size=x.shape)
# See that the above doesn't work any more:
output_vector = reversible_layer_forward(input_vector, f, g)
reversed_vector = reversible_layer_reverse(output_vector, f, g)
assert not np.allclose(reversed_vector, input_vector) # Fails!!
# It failed because the noise when reversing used a different random seed.
random_seed = 27686
rng = trax.fastmath.random.get_prng(random_seed)
f = lambda x: x + trax.fastmath.random.uniform(key=rng, shape=x.shape)
# See that it works now as the same rng is used on forward and reverse.
output_vector = reversible_layer_forward(input_vector, f, g)
reversed_vector = reversible_layer_reverse(output_vector, f, g)
assert np.allclose(reversed_vector, input_vector, atol=1e-07)
```
<a name="4"></a>
# Part 4: ReformerLM Training
You will now proceed to training your model. Since you have already know the two main components that differentiates it from the standard Transformer, LSH in Course 1 and reversible layers above, you can just use the pre-built model already implemented in Trax. It will have this architecture:
<img src='Reformer.jpg'>
Similar to the Transformer you learned earlier, you want to apply an attention and feed forward layer to your inputs. For the Reformer, we improve the memory efficiency by using **reversible decoder blocks** and you can picture its implementation in Trax like below:
<img src='ReversibleDecoder.png'>
You can see that it takes the initial inputs `x1` and `x2` and does the first equation of the reversible networks you learned in Part 3. As you've also learned, the reversible residual has two equations for the forward-pass so doing just one of them will just constitute half of the reversible decoder block. Before doing the second equation (i.e. second half of the reversible residual), it first needs to swap the elements to take into account the stack semantics in Trax. It simply puts `x2` on top of the stack so it can be fed to the add block of the half-residual layer. It then swaps the two outputs again so it can be fed to the next layer of the network. All of these arrives at the two equations in Part 3 and it can be used to recompute the activations during the backward pass.
These are already implemented for you in Trax and in the following exercise, you'll get to practice how to call them to build your network.
<a name="ex04"></a>
### Exercise 04
**Instructions:** Implement a wrapper function that returns a Reformer Language Model. You can use Trax's [ReformerLM](https://trax-ml.readthedocs.io/en/latest/trax.models.html#trax.models.reformer.reformer.ReformerLM) to do this quickly. It will have the same architecture as shown above.
```
# UNQ_C4
# GRADED FUNCTION
def ReformerLM(vocab_size=33000, n_layers=2, mode='train', attention_type=tl.SelfAttention):
"""
Args:
vocab_size (int): size of the vocabulary
n_layers (int): number of decoder layers
mode (string): setting of the model which can be 'train', 'eval', or 'predict'
attention_type(class): attention class to use
Returns:
model (ReformerLM): a reformer language model implemented in Trax
"""
### START CODE HERE (REPLACE INSTANCES OF 'None' WITH YOUR CODE) ###
# initialize an instance of Trax's ReformerLM class
model = trax.models.reformer.ReformerLM(
# set vocab size
vocab_size=vocab_size,
# set number of layers
n_layers=n_layers,
# set mode
mode=mode,
# set attention type
attention_type=attention_type
)
### END CODE HERE ###
return model
# display the model
temp_model = ReformerLM('train')
print(str(temp_model))
# free memory
del temp_model
# BEGIN UNIT TEST
w4_unittest.test_ReformerLM(ReformerLM)
# END UNIT TEST
```
<a name="ex05"></a>
### Exercise 05
You will now write a function that takes in your model and trains it.
**Instructions:** Implement the `training_loop` below to train the neural network above. Here is a list of things you should do:
- Create `TrainTask` and `EvalTask`
- Create the training loop `trax.supervised.training.Loop`
- Pass in the following depending to train_task :
- `labeled_data=train_gen`
- `loss_layer=tl.CrossEntropyLoss()`
- `optimizer=trax.optimizers.Adam(0.01)`
- `lr_schedule=lr_schedule`
- `n_steps_per_checkpoint=10`
You will be using your CrossEntropyLoss loss function with Adam optimizer. Please read the [trax](https://trax-ml.readthedocs.io/en/latest/trax.optimizers.html?highlight=adam#trax.optimizers.adam.Adam) documentation to get a full understanding.
- Pass in the following to eval_task:
- `labeled_data=eval_gen`
- `metrics=[tl.CrossEntropyLoss(), tl.Accuracy()]`
This function should return a `training.Loop` object. To read more about this check the [docs](https://trax-ml.readthedocs.io/en/latest/trax.supervised.html?highlight=loop#trax.supervised.training.Loop).
```
# UNQ_C5
# GRADED FUNCTION: train_model
def training_loop(ReformerLM, train_gen, eval_gen, output_dir = "./model/"):
"""
Args:
ReformerLM: the Reformer language model you are building
train_gen (generator): train data generator.
eval_gen (generator): Validation generator.
output_dir (string): Path to save the model output. Defaults to './model/'.
Returns:
trax.supervised.training.Loop: Training loop for the model.
"""
# use the warmup_and_rsqrt_decay learning rate schedule
lr_schedule = trax.lr.warmup_and_rsqrt_decay(
n_warmup_steps=1000, max_value=0.01)
### START CODE HERE (REPLACE INSTANCES OF 'None' WITH YOUR CODE) ###
# define the train task
train_task = training.TrainTask(
# labeled data
labeled_data=train_gen,
# loss layer
loss_layer=tl.CrossEntropyLoss(),
# optimizer
optimizer=trax.optimizers.Adam(0.01),
# lr_schedule
lr_schedule=lr_schedule,
# n_steps
n_steps_per_checkpoint=10
)
# define the eval task
eval_task = training.EvalTask(
# labeled data
labeled_data=eval_gen,
# metrics
metrics=[tl.CrossEntropyLoss(), tl.Accuracy()]
)
### END CODE HERE ###
loop = training.Loop(ReformerLM(mode='train'),
train_task,
eval_tasks=[eval_task],
output_dir=output_dir)
return loop
# UNIT TEST COMMENT: Use the train task and eval task for grading train_model
test_loop = training_loop(ReformerLM, train_stream, eval_stream)
train_task = test_loop._task
eval_task = test_loop._eval_task
print(train_task)
print(eval_task)
# BEGIN UNIT TEST
w4_unittest.test_tasks(train_task, eval_task)
# END UNIT TEST
# we will now test your function
!rm -f model/model.pkl.gz
loop = training_loop(ReformerLM, train_stream, eval_stream)
loop.run(10)
```
**Approximate Expected output:**
```
Step 1: Ran 1 train steps in 55.73 secs
Step 1: train CrossEntropyLoss | 10.41907787
Step 1: eval CrossEntropyLoss | 10.41005802
Step 1: eval Accuracy | 0.00000000
Step 10: Ran 9 train steps in 108.21 secs
Step 10: train CrossEntropyLoss | 10.15449715
Step 10: eval CrossEntropyLoss | 9.63478279
Step 10: eval Accuracy | 0.16350447
```
<a name="5"></a>
# Part 5: Decode from a pretrained model
We will now proceed on decoding using the model architecture you just implemented. As in the previous weeks, we will be giving you a pretrained model so you can observe meaningful output during inference. You will be using the [autoregressive_sample_stream()](https://trax-ml.readthedocs.io/en/latest/trax.supervised.html#trax.supervised.decoding.autoregressive_sample_stream) decoding method from Trax to do fast inference. Let's define a few parameters to initialize our model.
```
# define the `predict_mem_len` and `predict_drop_len` of tl.SelfAttention
def attention(*args, **kwargs):
# number of input positions to remember in a cache when doing fast inference.
kwargs['predict_mem_len'] = 120
# number of input elements to drop once the fast inference input cache fills up.
kwargs['predict_drop_len'] = 120
# return the attention layer with the parameters defined above
return tl.SelfAttention(*args, **kwargs)
# define the model using the ReformerLM function you implemented earlier.
model = ReformerLM(
vocab_size=33000,
n_layers=6,
mode='predict',
attention_type=attention,
)
# define an input signature so we can initialize our model. shape will be (1, 1) and the data type is int32.
shape11 = trax.shapes.ShapeDtype((1, 1), dtype=np.int32)
```
We can now initialize our model from a file containing the pretrained weights. We will save this starting state so we can reset the model state when we generate a new conversation. This will become clearer in the `generate_dialogue()` function later.
```
# initialize from file
model.init_from_file('chatbot_model1.pkl.gz',
weights_only=True, input_signature=shape11)
# save the starting state
STARTING_STATE = model.state
```
Let's define a few utility functions as well to help us tokenize and detokenize. We can use the [tokenize()](https://trax-ml.readthedocs.io/en/latest/trax.data.html#trax.data.tf_inputs.tokenize) and [detokenize()](https://trax-ml.readthedocs.io/en/latest/trax.data.html#trax.data.tf_inputs.detokenize) from `trax.data.tf_inputs` to do this.
```
def tokenize(sentence, vocab_file, vocab_dir):
return list(trax.data.tokenize(iter([sentence]), vocab_file=vocab_file, vocab_dir=vocab_dir))[0]
def detokenize(tokens, vocab_file, vocab_dir):
return trax.data.detokenize(tokens, vocab_file=vocab_file, vocab_dir=vocab_dir)
```
We are now ready to define our decoding function. This will return a generator that yields that next symbol output by the model. It will be able to predict the next words by just feeding it a starting sentence.
<a name="ex06"></a>
### Exercise 06
**Instructions:** Implement the function below to return a generator that predicts the next word of the conversation.
```
# UNQ_C6
# GRADED FUNCTION
def ReformerLM_output_gen(ReformerLM, start_sentence, vocab_file, vocab_dir, temperature):
"""
Args:
ReformerLM: the Reformer language model you just trained
start_sentence (string): starting sentence of the conversation
vocab_file (string): vocabulary filename
vocab_dir (string): directory of the vocabulary file
temperature (float): parameter for sampling ranging from 0.0 to 1.0.
0.0: same as argmax, always pick the most probable token
1.0: sampling from the distribution (can sometimes say random things)
Returns:
generator: yields the next symbol generated by the model
"""
### START CODE HERE (REPLACE INSTANCES OF 'None' WITH YOUR CODE) ###
# Create input tokens using the the tokenize function
input_tokens = tokenize(start_sentence, vocab_file=vocab_file, vocab_dir=vocab_dir)
# Add batch dimension to array. Convert from (n,) to (x, n) where
# x is the batch size. Default is 1. (hint: you can use np.expand_dims() with axis=0)
input_tokens_with_batch = np.array(input_tokens)[None, :]
# call the autoregressive_sample_stream function from trax
output_gen = trax.supervised.decoding.autoregressive_sample_stream(
# model
ReformerLM,
# inputs will be the tokens with batch dimension
inputs=input_tokens_with_batch,
# temperature
temperature=temperature
)
### END CODE HERE ###
return output_gen
# BEGIN UNIT TEST
import pickle
WEIGHTS_FROM_FILE = ()
with open('weights', 'rb') as file:
WEIGHTS_FROM_FILE = pickle.load(file)
shape11 = trax.shapes.ShapeDtype((1, 1), dtype=np.int32)
def attention(*args, **kwargs):
kwargs['predict_mem_len'] = 120
kwargs['predict_drop_len'] = 120
return tl.SelfAttention(*args, **kwargs)
test_model = ReformerLM(vocab_size=5, n_layers=1, mode='predict', attention_type=attention)
test_output_gen = ReformerLM_output_gen(test_model, "test", vocab_file=VOCAB_FILE, vocab_dir=VOCAB_DIR, temperature=0)
test_model.init_weights_and_state(shape11)
test_model.weights = WEIGHTS_FROM_FILE
output = []
for i in range(6):
output.append(next(test_output_gen)[0])
print(output)
# free memory
del test_model
del WEIGHTS_FROM_FILE
del test_output_gen
# END UNIT TEST
```
***Expected value:***
[1, 0, 4, 3, 0, 4]
Great! Now you will be able to see the model in action. The utility function below will call the generator you just implemented and will just format the output to be easier to read.
```
shape11 = trax.shapes.ShapeDtype((1, 1), dtype=np.int32)
def attention(*args, **kwargs):
kwargs['predict_mem_len'] = 120 # max length for predictions
kwargs['predict_drop_len'] = 120 # never drop old stuff
return tl.SelfAttention(*args, **kwargs)
model = ReformerLM(
vocab_size=33000,
n_layers=6,
mode='predict',
attention_type=attention,
)
model.init_from_file('chatbot_model1.pkl.gz',
weights_only=True, input_signature=shape11)
STARTING_STATE = model.state
def generate_dialogue(ReformerLM, model_state, start_sentence, vocab_file, vocab_dir, max_len, temperature):
"""
Args:
ReformerLM: the Reformer language model you just trained
model_state (np.array): initial state of the model before decoding
start_sentence (string): starting sentence of the conversation
vocab_file (string): vocabulary filename
vocab_dir (string): directory of the vocabulary file
max_len (int): maximum number of tokens to generate
temperature (float): parameter for sampling ranging from 0.0 to 1.0.
0.0: same as argmax, always pick the most probable token
1.0: sampling from the distribution (can sometimes say random things)
Returns:
generator: yields the next symbol generated by the model
"""
# define the delimiters we used during training
delimiter_1 = 'Person 1: '
delimiter_2 = 'Person 2: '
# initialize detokenized output
sentence = ''
# token counter
counter = 0
# output tokens. we insert a ': ' for formatting
result = [tokenize(': ', vocab_file=vocab_file, vocab_dir=vocab_dir)]
# reset the model state when starting a new dialogue
ReformerLM.state = model_state
# calls the output generator implemented earlier
output = ReformerLM_output_gen(ReformerLM, start_sentence, vocab_file=VOCAB_FILE, vocab_dir=VOCAB_DIR, temperature=temperature)
# print the starting sentence
print(start_sentence.split(delimiter_2)[0].strip())
# loop below yields the next tokens until max_len is reached. the if-elif is just for prettifying the output.
for o in output:
result.append(o)
sentence = detokenize(np.concatenate(result, axis=0), vocab_file=VOCAB_FILE, vocab_dir=VOCAB_DIR)
if sentence.endswith(delimiter_1):
sentence = sentence.split(delimiter_1)[0]
print(f'{delimiter_2}{sentence}')
sentence = ''
result.clear()
elif sentence.endswith(delimiter_2):
sentence = sentence.split(delimiter_2)[0]
print(f'{delimiter_1}{sentence}')
sentence = ''
result.clear()
counter += 1
if counter > max_len:
break
```
We can now feed in different starting sentences and see how the model generates the dialogue. You can even input your own starting sentence. Just remember to ask a question that covers the topics in the Multiwoz dataset so you can generate a meaningful conversation.
```
sample_sentence = ' Person 1: Are there theatres in town? Person 2: '
generate_dialogue(ReformerLM=model, model_state=STARTING_STATE, start_sentence=sample_sentence, vocab_file=VOCAB_FILE, vocab_dir=VOCAB_DIR, max_len=120, temperature=0.2)
sample_sentence = ' Person 1: Is there a hospital nearby? Person 2: '
generate_dialogue(ReformerLM=model, model_state=STARTING_STATE, start_sentence=sample_sentence, vocab_file=VOCAB_FILE, vocab_dir=VOCAB_DIR, max_len=120, temperature=0.2)
sample_sentence = ' Person 1: Can you book a taxi? Person 2: '
generate_dialogue(ReformerLM=model, model_state=STARTING_STATE, start_sentence=sample_sentence, vocab_file=VOCAB_FILE, vocab_dir=VOCAB_DIR, max_len=120, temperature=0.2)
```
**Congratulations! You just wrapped up the final assignment of this course and the entire specialization!**
| github_jupyter |
```
# Import packages
import os
from matplotlib import pyplot as plt
import pandas as pd
# Import AuTuMN modules
from autumn.settings import Models, Region
from autumn.settings.folders import OUTPUT_DATA_PATH
from autumn.tools.project import get_project
from autumn.tools import db
from autumn.tools.plots.calibration.plots import calculate_r_hats, get_output_from_run_id
from autumn.tools.plots.uncertainty.plots import _plot_uncertainty, _get_target_values
from autumn.tools.plots.plotter.base_plotter import COLOR_THEME
from autumn.tools.plots.utils import get_plot_text_dict, change_xaxis_to_date, REF_DATE, ALPHAS, COLORS, _apply_transparency, _plot_targets_to_axis
from autumn.dashboards.calibration_results.plots import get_uncertainty_df
# Specify model details
model = Models.COVID_19
region = Region.VICTORIA_2021
dirname = "2021-09-13"
# get the relevant project and output data
project = get_project(model, region)
project_calib_dir = os.path.join(
OUTPUT_DATA_PATH, "calibrate", project.model_name, project.region_name
)
calib_path = os.path.join(project_calib_dir, dirname)
# Load tables
mcmc_tables = db.load.load_mcmc_tables(calib_path)
mcmc_params = db.load.load_mcmc_params_tables(calib_path)
uncertainty_df = get_uncertainty_df(calib_path, mcmc_tables, project.plots)
scenario_list = uncertainty_df['scenario'].unique()
# make output directories
output_dir = f"{model}_{region}_{dirname}"
base_dir = os.path.join("outputs", output_dir)
os.makedirs(base_dir, exist_ok=True)
dirs_to_make = ["calibration", "MLE", "median", "uncertainty", "csv_files"]
for dir_to_make in dirs_to_make:
os.makedirs(os.path.join(base_dir, dir_to_make), exist_ok=True)
# get R_hat diagnostics
r_hats = calculate_r_hats(mcmc_params, mcmc_tables, burn_in=0)
for key, value in r_hats.items():
print(f"{key}: {value}")
titles = {
"notifications": "Daily number of notified Covid-19 cases",
"infection_deaths": "Daily number of Covid-19 deaths",
"accum_deaths": "Cumulative number of Covid-19 deaths",
"incidence": "Daily incidence (incl. asymptomatics and undetected)",
"hospital_occupancy": "Hospital beds occupied by Covid-19 patients",
"icu_occupancy": "ICU beds occupied by Covid-19 patients",
"cdr": "Proportion detected among symptomatics",
"proportion_vaccinated": "Proportion vaccinated",
"prop_incidence_strain_delta": "Proportion of Delta variant in new cases"
}
def plot_outputs(output_type, output_name, scenario_list, sc_linestyles, sc_colors, show_v_lines=False, x_min=590, x_max=775):
# plot options
title = titles[output_name]
title_fontsize = 18
label_font_size = 15
linewidth = 3
n_xticks = 10
# initialise figure
fig = plt.figure(figsize=(12, 8))
plt.style.use("ggplot")
axis = fig.add_subplot()
# prepare colors for ucnertainty
n_scenarios_to_plot = len(scenario_list)
uncertainty_colors = _apply_transparency(COLORS[:n_scenarios_to_plot], ALPHAS[:n_scenarios_to_plot])
if output_type == "MLE":
derived_output_tables = db.load.load_derived_output_tables(calib_path, column=output_name)
for i, scenario in enumerate(scenario_list):
linestyle = sc_linestyles[scenario]
color = sc_colors[scenario]
if output_type == "MLE":
times, values = get_output_from_run_id(output_name, mcmc_tables, derived_output_tables, "MLE", scenario)
axis.plot(times, values, color=color, linestyle=linestyle, linewidth=linewidth)
elif output_type == "median":
_plot_uncertainty(
axis,
uncertainty_df,
output_name,
scenario,
x_max,
x_min,
[_, _, _, color],
overlay_uncertainty=False,
start_quantile=0,
zorder=scenario + 1,
linestyle=linestyle,
linewidth=linewidth,
)
elif output_type == "uncertainty":
scenario_colors = uncertainty_colors[i]
_plot_uncertainty(
axis,
uncertainty_df,
output_name,
scenario,
x_max,
x_min,
scenario_colors,
overlay_uncertainty=True,
start_quantile=0,
zorder=scenario + 1,
)
else:
print("Please use supported output_type option")
axis.set_xlim((x_min, x_max))
axis.set_title(title, fontsize=title_fontsize)
plt.setp(axis.get_yticklabels(), fontsize=label_font_size)
plt.setp(axis.get_xticklabels(), fontsize=label_font_size)
change_xaxis_to_date(axis, REF_DATE)
plt.locator_params(axis="x", nbins=n_xticks)
if show_v_lines:
release_dates = {624: "15 Sep 2021", 609: "31 Aug 2021"}
y_max = plt.gca().get_ylim()[1]
linestyles = ["dashdot", "solid"]
i = 0
for time, date in release_dates.items():
plt.vlines(time, ymin=0, ymax=y_max, linestyle=linestyles[i])
text = f"Lockdown relaxed on {date}"
plt.text(time - 5, .5*y_max, text, rotation=90, fontsize=11)
i += 1
return axis
```
# Scenario plots with single lines
```
output_names = ["notifications", "icu_occupancy", "incidence", "accum_deaths"]
scenario_x_min, scenario_x_max = 575, 731
sc_colors = [COLOR_THEME[i] for i in scenario_list]
sc_linestyles = ["dotted"] + ["solid"] * (len(scenario_list) - 1)
for output_type in ["median", "MLE"]:
for output_name in output_names:
plot_outputs(output_type, output_name, scenario_list, sc_linestyles, sc_colors, False, x_min=scenario_x_min, x_max=scenario_x_max)
path = os.path.join(base_dir, output_type, f"{output_name}.png")
plt.savefig(path)
# For Ho Chi Minh City
# early_release_linestyle = "solid"
# late_release_linestyle = "dashdot"
# vacc_colors = ["crimson", "seagreen","slateblue", "coral"]
# sc_linestyles = ["dotted"] + [late_release_linestyle] * 4 + [early_release_linestyle] * 4
# sc_colors = ["black"] + vacc_colors * 2
# # Plot outputs
# output_names = ["notifications", "infection_deaths", "hospital_occupancy", "icu_occupancy"]
# # output_names = ["proportion_vaccinated"]
# for output_type in ["median", "MLE"]:
# scenario_list = [0, 1, 2, 3, 4, 5, 6, 7, 8]
# for output_name in output_names:
# plot_outputs(output_type, output_name, scenario_list, sc_linestyles, sc_colors, True, x_min=590, x_max=775) # 475, 650 )
# path = f"{output_dir}/{output_type}/{output_name}.png"
# plt.savefig(path)
```
# Uncertainty around scenarios
```
output_type = "uncertainty"
for scenario in scenario_list:
if scenario == 0:
continue
scenarios_to_plot = [0, scenario]
for output_name in output_names:
plot_outputs(output_type, output_name, scenarios_to_plot, sc_linestyles, sc_colors, False, x_min=scenario_x_min, x_max=scenario_x_max)
path = os.path.join(base_dir, output_type, f"{output_name}_scenario_{scenario}.png")
plt.savefig(path)
```
# Uncertainty around baseline only (Calibration plots)
```
calibration_x_min, calibration_x_max = 580, 680
for output_name in output_names + ["cdr", "prop_incidence_strain_delta"]:
axis = plot_outputs("uncertainty", output_name, [0], sc_linestyles, sc_colors, False, x_min=calibration_x_min, x_max=calibration_x_max)
path = os.path.join(base_dir, 'calibration', f"{output_name}.png")
targets = project.plots
targets = {k: v for k, v in targets.items() if v["output_key"] == output_name}
values, times = _get_target_values(targets, output_name)
_plot_targets_to_axis(axis, values, times, on_uncertainty_plot=True)
plt.savefig(path)
```
# Dump outputs to csv files
```
csv_outputs = ["icu_occupancy"]
start_time = 609 # 31 Aug 2021
includes_MLE = True
requested_quantiles = [0.025, 0.50, 0.975]
# for age in [str(int(5. * i)) for i in range(16)]:
# csv_outputs.append(f"notificationsXagegroup_{age}")
def get_uncertainty_data(output_name, scenario_idx, quantile):
mask = (
(uncertainty_df["type"] == output_name)
& (uncertainty_df["scenario"] == scenario_idx)
& (uncertainty_df["quantile"] == quantile)
)
df = uncertainty_df[mask]
times = df.time.unique()[1:]
values = df["value"].tolist()[1:]
return times, values
COVID_BASE_DATE = pd.datetime(2019, 12, 31)
start_date = pd.to_timedelta(start_time, unit="days") + (COVID_BASE_DATE)
for scenario in scenario_list:
df = pd.DataFrame()
# include a column for the date
t, _ = get_uncertainty_data("notifications", scenario, 0.5)
df["date"] = pd.to_timedelta(t, unit="days") + (COVID_BASE_DATE)
for output in csv_outputs:
if includes_MLE:
derived_output_tables = db.load.load_derived_output_tables(calib_path, column=output)
do_times, do_values = get_output_from_run_id(output, mcmc_tables, derived_output_tables, "MLE", scenario)
assert list(do_times[1:]) == list(t)
do_values = list(do_values)[1:]
name = f"{output}_MLE"
df[name] = do_values
if output in list(uncertainty_df["type"].unique()):
for quantile in requested_quantiles:
_, v = get_uncertainty_data(output, scenario, quantile)
name = f"{output}_{quantile}"
df[name] = v
# trim the dataframe to keep requested times only
df.drop(df[df.date < start_date].index, inplace=True)
path = os.path.join(base_dir, 'csv_files', f"outputs_scenario_{scenario}.csv")
df.to_csv(path)
```
| github_jupyter |
# Deep Convolutional GANs
In this notebook, you'll build a GAN using convolutional layers in the generator and discriminator. This is called a Deep Convolutional GAN, or DCGAN for short. The DCGAN architecture was first explored in 2016 and has seen impressive results in generating new images; you can read the [original paper, here](https://arxiv.org/pdf/1511.06434.pdf).
You'll be training DCGAN on the [Street View House Numbers](http://ufldl.stanford.edu/housenumbers/) (SVHN) dataset. These are color images of house numbers collected from Google street view. SVHN images are in color and much more variable than MNIST.
<img src='assets/svhn_dcgan.png' width=80% />
So, our goal is to create a DCGAN that can generate new, realistic-looking images of house numbers. We'll go through the following steps to do this:
* Load in and pre-process the house numbers dataset
* Define discriminator and generator networks
* Train these adversarial networks
* Visualize the loss over time and some sample, generated images
#### Deeper Convolutional Networks
Since this dataset is more complex than our MNIST data, we'll need a deeper network to accurately identify patterns in these images and be able to generate new ones. Specifically, we'll use a series of convolutional or transpose convolutional layers in the discriminator and generator. It's also necessary to use batch normalization to get these convolutional networks to train.
Besides these changes in network structure, training the discriminator and generator networks should be the same as before. That is, the discriminator will alternate training on real and fake (generated) images, and the generator will aim to trick the discriminator into thinking that its generated images are real!
```
# import libraries
import matplotlib.pyplot as plt
import numpy as np
import pickle as pkl
%matplotlib inline
```
## Getting the data
Here you can download the SVHN dataset. It's a dataset built-in to the PyTorch datasets library. We can load in training data, transform it into Tensor datatypes, then create dataloaders to batch our data into a desired size.
```
import torch
from torchvision import datasets
from torchvision import transforms
# Tensor transform
transform = transforms.ToTensor()
# SVHN training datasets
svhn_train = datasets.SVHN(root='data/', split='train', download=True, transform=transform)
batch_size = 128
num_workers = 0
# build DataLoaders for SVHN dataset
train_loader = torch.utils.data.DataLoader(dataset=svhn_train,
batch_size=batch_size,
shuffle=True,
num_workers=num_workers)
```
### Visualize the Data
Here I'm showing a small sample of the images. Each of these is 32x32 with 3 color channels (RGB). These are the real, training images that we'll pass to the discriminator. Notice that each image has _one_ associated, numerical label.
```
# obtain one batch of training images
dataiter = iter(train_loader)
images, labels = dataiter.next()
# plot the images in the batch, along with the corresponding labels
fig = plt.figure(figsize=(25, 4))
plot_size=20
for idx in np.arange(plot_size):
ax = fig.add_subplot(2, plot_size/2, idx+1, xticks=[], yticks=[])
ax.imshow(np.transpose(images[idx], (1, 2, 0)))
# print out the correct label for each image
# .item() gets the value contained in a Tensor
ax.set_title(str(labels[idx].item()))
```
### Pre-processing: scaling from -1 to 1
We need to do a bit of pre-processing; we know that the output of our `tanh` activated generator will contain pixel values in a range from -1 to 1, and so, we need to rescale our training images to a range of -1 to 1. (Right now, they are in a range from 0-1.)
```
# current range
img = images[0]
print('Min: ', img.min())
print('Max: ', img.max())
# helper scale function
def scale(x, feature_range=(-1, 1)):
''' Scale takes in an image x and returns that image, scaled
with a feature_range of pixel values from -1 to 1.
This function assumes that the input x is already scaled from 0-1.'''
# assume x is scaled to (0, 1)
# scale to feature_range and return scaled x
min,max = feature_range
x = x * (max-min) + min
return x
# scaled range
scaled_img = scale(img)
print('Scaled min: ', scaled_img.min())
print('Scaled max: ', scaled_img.max())
```
---
# Define the Model
A GAN is comprised of two adversarial networks, a discriminator and a generator.
## Discriminator
Here you'll build the discriminator. This is a convolutional classifier like you've built before, only without any maxpooling layers.
* The inputs to the discriminator are 32x32x3 tensor images
* You'll want a few convolutional, hidden layers
* Then a fully connected layer for the output; as before, we want a sigmoid output, but we'll add that in the loss function, [BCEWithLogitsLoss](https://pytorch.org/docs/stable/nn.html#bcewithlogitsloss), later
<img src='assets/conv_discriminator.png' width=80%/>
For the depths of the convolutional layers I suggest starting with 32 filters in the first layer, then double that depth as you add layers (to 64, 128, etc.). Note that in the DCGAN paper, they did all the downsampling using only strided convolutional layers with no maxpooling layers.
You'll also want to use batch normalization with [nn.BatchNorm2d](https://pytorch.org/docs/stable/nn.html#batchnorm2d) on each layer **except** the first convolutional layer and final, linear output layer.
#### Helper `conv` function
In general, each layer should look something like convolution > batch norm > leaky ReLU, and so we'll define a function to put these layers together. This function will create a sequential series of a convolutional + an optional batch norm layer. We'll create these using PyTorch's [Sequential container](https://pytorch.org/docs/stable/nn.html#sequential), which takes in a list of layers and creates layers according to the order that they are passed in to the Sequential constructor.
Note: It is also suggested that you use a **kernel_size of 4** and a **stride of 2** for strided convolutions.
```
import torch.nn as nn
import torch.nn.functional as F
# helper conv function
def conv(in_channels, out_channels, kernel_size, stride=2, padding=1, batch_norm=True):
"""Creates a convolutional layer, with optional batch normalization.
"""
layers = []
conv_layer = nn.Conv2d(in_channels, out_channels,
kernel_size, stride, padding, bias=False)
# append conv layer
layers.append(conv_layer)
if batch_norm:
# append batchnorm layer
layers.append(nn.BatchNorm2d(out_channels))
# using Sequential container
return nn.Sequential(*layers)
class Discriminator(nn.Module):
def __init__(self, conv_dim=32):
super(Discriminator, self).__init__()
# complete init function
self.conv_dim = conv_dim
self.conv1 = conv(3, conv_dim, 4, batch_norm=False)
self.conv2 = conv(conv_dim, conv_dim*2, 4)
self.conv3 = conv(conv_dim*2, conv_dim*4, 4)
self.fc = nn.Linear(conv_dim*4*4*4, 1)
def forward(self, x):
# complete forward function
out = F.leaky_relu(self.conv1(x), 0.2)
out = F.leaky_relu(self.conv2(out), 0.2)
out = F.leaky_relu(self.conv3(out), 0.2)
out = out.view(-1, self.conv_dim*4*4*4)
out = self.fc(out)
return out
```
## Generator
Next, you'll build the generator network. The input will be our noise vector `z`, as before. And, the output will be a $tanh$ output, but this time with size 32x32 which is the size of our SVHN images.
<img src='assets/conv_generator.png' width=80% />
What's new here is we'll use transpose convolutional layers to create our new images.
* The first layer is a fully connected layer which is reshaped into a deep and narrow layer, something like 4x4x512.
* Then, we use batch normalization and a leaky ReLU activation.
* Next is a series of [transpose convolutional layers](https://pytorch.org/docs/stable/nn.html#convtranspose2d), where you typically halve the depth and double the width and height of the previous layer.
* And, we'll apply batch normalization and ReLU to all but the last of these hidden layers. Where we will just apply a `tanh` activation.
#### Helper `deconv` function
For each of these layers, the general scheme is transpose convolution > batch norm > ReLU, and so we'll define a function to put these layers together. This function will create a sequential series of a transpose convolutional + an optional batch norm layer. We'll create these using PyTorch's Sequential container, which takes in a list of layers and creates layers according to the order that they are passed in to the Sequential constructor.
Note: It is also suggested that you use a **kernel_size of 4** and a **stride of 2** for transpose convolutions.
```
# helper deconv function
def deconv(in_channels, out_channels, kernel_size, stride=2, padding=1, batch_norm=True):
"""Creates a transposed-convolutional layer, with optional batch normalization.
"""
## TODO: Complete this function
## create a sequence of transpose + optional batch norm layers
layers = []
transpose_conv_layer = nn.ConvTranspose2d(in_channels, out_channels, kernel_size, stride, padding, bias=False)
layers.append(transpose_conv_layer)
if batch_norm:
batch_norm_layer = nn.BatchNorm2d(out_channels)
layers.append(batch_norm_layer)
return nn.Sequential(*layers)
class Generator(nn.Module):
def __init__(self, z_size, conv_dim=32):
super(Generator, self).__init__()
# complete init function
self.conv_dim = conv_dim
self.fc = nn.Linear(z_size, conv_dim*4*4*4)
self.t_conv1 = deconv(conv_dim*4, conv_dim*2, 4)
self.t_conv2 = deconv(conv_dim*2, conv_dim, 4)
self.t_conv3 = deconv(conv_dim, 3, 4, batch_norm=False)
def forward(self, x):
# complete forward function
out = self.fc(x)
out = out.view(-1, self.conv_dim*4, 4, 4)
out = F.relu(self.t_conv1(out))
out = F.relu(self.t_conv2(out))
out = self.t_conv3(out)
out = F.tanh(out)
return out
```
## Build complete network
Define your models' hyperparameters and instantiate the discriminator and generator from the classes defined above. Make sure you've passed in the correct input arguments.
```
# define hyperparams
conv_dim = 32
z_size = 100
# define discriminator and generator
D = Discriminator(conv_dim)
G = Generator(z_size=z_size, conv_dim=conv_dim)
print(D)
print()
print(G)
```
# Training on GPU
Check if you can train on GPU. If you can, set this as a variable and move your models to GPU.
> Later, we'll also move any inputs our models and loss functions see (real_images, z, and ground truth labels) to GPU as well.
```
train_on_gpu = torch.cuda.is_available()
if train_on_gpu:
# move models to GPU
G.cuda()
D.cuda()
print('GPU available for training. Models moved to GPU')
else:
print('Training on CPU.')
```
---
## Discriminator and Generator Losses
Now we need to calculate the losses. And this will be exactly the same as before.
### Discriminator Losses
> * For the discriminator, the total loss is the sum of the losses for real and fake images, `d_loss = d_real_loss + d_fake_loss`.
* Remember that we want the discriminator to output 1 for real images and 0 for fake images, so we need to set up the losses to reflect that.
The losses will by binary cross entropy loss with logits, which we can get with [BCEWithLogitsLoss](https://pytorch.org/docs/stable/nn.html#bcewithlogitsloss). This combines a `sigmoid` activation function **and** and binary cross entropy loss in one function.
For the real images, we want `D(real_images) = 1`. That is, we want the discriminator to classify the the real images with a label = 1, indicating that these are real. The discriminator loss for the fake data is similar. We want `D(fake_images) = 0`, where the fake images are the _generator output_, `fake_images = G(z)`.
### Generator Loss
The generator loss will look similar only with flipped labels. The generator's goal is to get `D(fake_images) = 1`. In this case, the labels are **flipped** to represent that the generator is trying to fool the discriminator into thinking that the images it generates (fakes) are real!
```
def real_loss(D_out, smooth=False):
batch_size = D_out.size(0)
# label smoothing
if smooth:
# smooth, real labels = 0.9
labels = torch.ones(batch_size)*0.9
else:
labels = torch.ones(batch_size) # real labels = 1
# move labels to GPU if available
if train_on_gpu:
labels = labels.cuda()
# binary cross entropy with logits loss
criterion = nn.BCEWithLogitsLoss()
# calculate loss
loss = criterion(D_out.squeeze(), labels)
return loss
def fake_loss(D_out):
batch_size = D_out.size(0)
labels = torch.zeros(batch_size) # fake labels = 0
if train_on_gpu:
labels = labels.cuda()
criterion = nn.BCEWithLogitsLoss()
# calculate loss
loss = criterion(D_out.squeeze(), labels)
return loss
```
## Optimizers
Not much new here, but notice how I am using a small learning rate and custom parameters for the Adam optimizers, This is based on some research into DCGAN model convergence.
### Hyperparameters
GANs are very sensitive to hyperparameters. A lot of experimentation goes into finding the best hyperparameters such that the generator and discriminator don't overpower each other. Try out your own hyperparameters or read [the DCGAN paper](https://arxiv.org/pdf/1511.06434.pdf) to see what worked for them.
```
import torch.optim as optim
# params
lr = 0.0002
beta1=0.5
beta2=0.999
# Create optimizers for the discriminator and generator
d_optimizer = optim.Adam(D.parameters(), lr, [beta1, beta2])
g_optimizer = optim.Adam(G.parameters(), lr, [beta1, beta2])
```
---
## Training
Training will involve alternating between training the discriminator and the generator. We'll use our functions `real_loss` and `fake_loss` to help us calculate the discriminator losses in all of the following cases.
### Discriminator training
1. Compute the discriminator loss on real, training images
2. Generate fake images
3. Compute the discriminator loss on fake, generated images
4. Add up real and fake loss
5. Perform backpropagation + an optimization step to update the discriminator's weights
### Generator training
1. Generate fake images
2. Compute the discriminator loss on fake images, using **flipped** labels!
3. Perform backpropagation + an optimization step to update the generator's weights
#### Saving Samples
As we train, we'll also print out some loss statistics and save some generated "fake" samples.
**Evaluation mode**
Notice that, when we call our generator to create the samples to display, we set our model to evaluation mode: `G.eval()`. That's so the batch normalization layers will use the population statistics rather than the batch statistics (as they do during training), *and* so dropout layers will operate in eval() mode; not turning off any nodes for generating samples.
```
import pickle as pkl
# training hyperparams
num_epochs = 10
# keep track of loss and generated, "fake" samples
samples = []
losses = []
print_every = 300
# Get some fixed data for sampling. These are images that are held
# constant throughout training, and allow us to inspect the model's performance
sample_size=16
fixed_z = np.random.uniform(-1, 1, size=(sample_size, z_size))
fixed_z = torch.from_numpy(fixed_z).float()
# train the network
for epoch in range(num_epochs):
for batch_i, (real_images, _) in enumerate(train_loader):
batch_size = real_images.size(0)
# important rescaling step
real_images = scale(real_images)
# ============================================
# TRAIN THE DISCRIMINATOR
# ============================================
d_optimizer.zero_grad()
# 1. Train with real images
# Compute the discriminator losses on real images
if train_on_gpu:
real_images = real_images.cuda()
D_real = D(real_images)
d_real_loss = real_loss(D_real)
# 2. Train with fake images
# Generate fake images
z = np.random.uniform(-1, 1, size=(batch_size, z_size))
z = torch.from_numpy(z).float()
# move x to GPU, if available
if train_on_gpu:
z = z.cuda()
fake_images = G(z)
# Compute the discriminator losses on fake images
D_fake = D(fake_images)
d_fake_loss = fake_loss(D_fake)
# add up loss and perform backprop
d_loss = d_real_loss + d_fake_loss
d_loss.backward()
d_optimizer.step()
# =========================================
# TRAIN THE GENERATOR
# =========================================
g_optimizer.zero_grad()
# 1. Train with fake images and flipped labels
# Generate fake images
z = np.random.uniform(-1, 1, size=(batch_size, z_size))
z = torch.from_numpy(z).float()
if train_on_gpu:
z = z.cuda()
fake_images = G(z)
# Compute the discriminator losses on fake images
# using flipped labels!
D_fake = D(fake_images)
g_loss = real_loss(D_fake) # use real loss to flip labels
# perform backprop
g_loss.backward()
g_optimizer.step()
# Print some loss stats
if batch_i % print_every == 0:
# append discriminator loss and generator loss
losses.append((d_loss.item(), g_loss.item()))
# print discriminator and generator loss
print('Epoch [{:5d}/{:5d}] | d_loss: {:6.4f} | g_loss: {:6.4f}'.format(
epoch+1, num_epochs, d_loss.item(), g_loss.item()))
## AFTER EACH EPOCH##
# generate and save sample, fake images
G.eval() # for generating samples
if train_on_gpu:
fixed_z = fixed_z.cuda()
samples_z = G(fixed_z)
samples.append(samples_z)
G.train() # back to training mode
# Save training generator samples
with open('train_samples.pkl', 'wb') as f:
pkl.dump(samples, f)
```
## Training loss
Here we'll plot the training losses for the generator and discriminator, recorded after each epoch.
```
fig, ax = plt.subplots()
losses = np.array(losses)
plt.plot(losses.T[0], label='Discriminator', alpha=0.5)
plt.plot(losses.T[1], label='Generator', alpha=0.5)
plt.title("Training Losses")
plt.legend()
```
### let's do more epochs
```
import pickle as pkl
# training hyperparams
num_epochs = 30
# keep track of loss and generated, "fake" samples
samples = []
losses = []
print_every = 300
# Get some fixed data for sampling. These are images that are held
# constant throughout training, and allow us to inspect the model's performance
sample_size=16
fixed_z = np.random.uniform(-1, 1, size=(sample_size, z_size))
fixed_z = torch.from_numpy(fixed_z).float()
# train the network
for epoch in range(num_epochs):
for batch_i, (real_images, _) in enumerate(train_loader):
batch_size = real_images.size(0)
# important rescaling step
real_images = scale(real_images)
# ============================================
# TRAIN THE DISCRIMINATOR
# ============================================
d_optimizer.zero_grad()
# 1. Train with real images
# Compute the discriminator losses on real images
if train_on_gpu:
real_images = real_images.cuda()
D_real = D(real_images)
d_real_loss = real_loss(D_real)
# 2. Train with fake images
# Generate fake images
z = np.random.uniform(-1, 1, size=(batch_size, z_size))
z = torch.from_numpy(z).float()
# move x to GPU, if available
if train_on_gpu:
z = z.cuda()
fake_images = G(z)
# Compute the discriminator losses on fake images
D_fake = D(fake_images)
d_fake_loss = fake_loss(D_fake)
# add up loss and perform backprop
d_loss = d_real_loss + d_fake_loss
d_loss.backward()
d_optimizer.step()
# =========================================
# TRAIN THE GENERATOR
# =========================================
g_optimizer.zero_grad()
# 1. Train with fake images and flipped labels
# Generate fake images
z = np.random.uniform(-1, 1, size=(batch_size, z_size))
z = torch.from_numpy(z).float()
if train_on_gpu:
z = z.cuda()
fake_images = G(z)
# Compute the discriminator losses on fake images
# using flipped labels!
D_fake = D(fake_images)
g_loss = real_loss(D_fake) # use real loss to flip labels
# perform backprop
g_loss.backward()
g_optimizer.step()
# Print some loss stats
if batch_i % print_every == 0:
# append discriminator loss and generator loss
losses.append((d_loss.item(), g_loss.item()))
# print discriminator and generator loss
print('Epoch [{:5d}/{:5d}] | d_loss: {:6.4f} | g_loss: {:6.4f}'.format(
epoch+1, num_epochs, d_loss.item(), g_loss.item()))
## AFTER EACH EPOCH##
# generate and save sample, fake images
G.eval() # for generating samples
if train_on_gpu:
fixed_z = fixed_z.cuda()
samples_z = G(fixed_z)
samples.append(samples_z)
G.train() # back to training mode
# Save training generator samples
with open('train_samples.pkl', 'wb') as f:
pkl.dump(samples, f)
```
## Training loss
Here we'll plot the training losses for the generator and discriminator, recorded after each epoch.
```
fig, ax = plt.subplots()
losses = np.array(losses)
plt.plot(losses.T[0], label='Discriminator', alpha=0.5)
plt.plot(losses.T[1], label='Generator', alpha=0.5)
plt.title("Training Losses")
plt.legend()
```
## Generator samples from training
Here we can view samples of images from the generator. We'll look at the images we saved during training.
```
# helper function for viewing a list of passed in sample images
def view_samples(epoch, samples):
fig, axes = plt.subplots(figsize=(16,4), nrows=2, ncols=8, sharey=True, sharex=True)
for ax, img in zip(axes.flatten(), samples[epoch]):
img = img.detach().cpu().numpy()
img = np.transpose(img, (1, 2, 0))
img = ((img +1)*255 / (2)).astype(np.uint8) # rescale to pixel range (0-255)
ax.xaxis.set_visible(False)
ax.yaxis.set_visible(False)
im = ax.imshow(img.reshape((32,32,3)))
_ = view_samples(-1, samples)
```
now you can see few realistic images of house number, with more training and tuning it can be made much better!
## BINGO!
| github_jupyter |
```
%matplotlib inline
```
# RBF SVM parameters
This example illustrates the effect of the parameters ``gamma`` and ``C`` of
the Radial Basis Function (RBF) kernel SVM.
Intuitively, the ``gamma`` parameter defines how far the influence of a single
training example reaches, with low values meaning 'far' and high values meaning
'close'. The ``gamma`` parameters can be seen as the inverse of the radius of
influence of samples selected by the model as support vectors.
The ``C`` parameter trades off misclassification of training examples against
simplicity of the decision surface. A low ``C`` makes the decision surface
smooth, while a high ``C`` aims at classifying all training examples correctly
by giving the model freedom to select more samples as support vectors.
The first plot is a visualization of the decision function for a variety of
parameter values on a simplified classification problem involving only 2 input
features and 2 possible target classes (binary classification). Note that this
kind of plot is not possible to do for problems with more features or target
classes.
The second plot is a heatmap of the classifier's cross-validation accuracy as a
function of ``C`` and ``gamma``. For this example we explore a relatively large
grid for illustration purposes. In practice, a logarithmic grid from
$10^{-3}$ to $10^3$ is usually sufficient. If the best parameters
lie on the boundaries of the grid, it can be extended in that direction in a
subsequent search.
Note that the heat map plot has a special colorbar with a midpoint value close
to the score values of the best performing models so as to make it easy to tell
them appart in the blink of an eye.
The behavior of the model is very sensitive to the ``gamma`` parameter. If
``gamma`` is too large, the radius of the area of influence of the support
vectors only includes the support vector itself and no amount of
regularization with ``C`` will be able to prevent overfitting.
When ``gamma`` is very small, the model is too constrained and cannot capture
the complexity or "shape" of the data. The region of influence of any selected
support vector would include the whole training set. The resulting model will
behave similarly to a linear model with a set of hyperplanes that separate the
centers of high density of any pair of two classes.
For intermediate values, we can see on the second plot that good models can
be found on a diagonal of ``C`` and ``gamma``. Smooth models (lower ``gamma``
values) can be made more complex by selecting a larger number of support
vectors (larger ``C`` values) hence the diagonal of good performing models.
Finally one can also observe that for some intermediate values of ``gamma`` we
get equally performing models when ``C`` becomes very large: it is not
necessary to regularize by limiting the number of support vectors. The radius of
the RBF kernel alone acts as a good structural regularizer. In practice though
it might still be interesting to limit the number of support vectors with a
lower value of ``C`` so as to favor models that use less memory and that are
faster to predict.
We should also note that small differences in scores results from the random
splits of the cross-validation procedure. Those spurious variations can be
smoothed out by increasing the number of CV iterations ``n_splits`` at the
expense of compute time. Increasing the value number of ``C_range`` and
``gamma_range`` steps will increase the resolution of the hyper-parameter heat
map.
```
print(__doc__)
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.colors import Normalize
from sklearn.svm import SVC
from sklearn.preprocessing import StandardScaler
from sklearn.datasets import load_iris
from sklearn.model_selection import StratifiedShuffleSplit
from sklearn.model_selection import GridSearchCV
# Utility function to move the midpoint of a colormap to be around
# the values of interest.
class MidpointNormalize(Normalize):
def __init__(self, vmin=None, vmax=None, midpoint=None, clip=False):
self.midpoint = midpoint
Normalize.__init__(self, vmin, vmax, clip)
def __call__(self, value, clip=None):
x, y = [self.vmin, self.midpoint, self.vmax], [0, 0.5, 1]
return np.ma.masked_array(np.interp(value, x, y))
# #############################################################################
# Load and prepare data set
#
# dataset for grid search
iris = load_iris()
X = iris.data
y = iris.target
# Dataset for decision function visualization: we only keep the first two
# features in X and sub-sample the dataset to keep only 2 classes and
# make it a binary classification problem.
X_2d = X[:, :2]
X_2d = X_2d[y > 0]
y_2d = y[y > 0]
y_2d -= 1
# It is usually a good idea to scale the data for SVM training.
# We are cheating a bit in this example in scaling all of the data,
# instead of fitting the transformation on the training set and
# just applying it on the test set.
scaler = StandardScaler()
X = scaler.fit_transform(X)
X_2d = scaler.fit_transform(X_2d)
# #############################################################################
# Train classifiers
#
# For an initial search, a logarithmic grid with basis
# 10 is often helpful. Using a basis of 2, a finer
# tuning can be achieved but at a much higher cost.
C_range = np.logspace(-2, 10, 13)
gamma_range = np.logspace(-9, 3, 13)
param_grid = dict(gamma=gamma_range, C=C_range)
cv = StratifiedShuffleSplit(n_splits=5, test_size=0.2, random_state=42)
grid = GridSearchCV(SVC(), param_grid=param_grid, cv=cv)
grid.fit(X, y)
print("The best parameters are %s with a score of %0.2f"
% (grid.best_params_, grid.best_score_))
# Now we need to fit a classifier for all parameters in the 2d version
# (we use a smaller set of parameters here because it takes a while to train)
C_2d_range = [1e-2, 1, 1e2]
gamma_2d_range = [1e-1, 1, 1e1]
classifiers = []
for C in C_2d_range:
for gamma in gamma_2d_range:
clf = SVC(C=C, gamma=gamma)
clf.fit(X_2d, y_2d)
classifiers.append((C, gamma, clf))
# #############################################################################
# Visualization
#
# draw visualization of parameter effects
plt.figure(figsize=(8, 6))
xx, yy = np.meshgrid(np.linspace(-3, 3, 200), np.linspace(-3, 3, 200))
for (k, (C, gamma, clf)) in enumerate(classifiers):
# evaluate decision function in a grid
Z = clf.decision_function(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
# visualize decision function for these parameters
plt.subplot(len(C_2d_range), len(gamma_2d_range), k + 1)
plt.title("gamma=10^%d, C=10^%d" % (np.log10(gamma), np.log10(C)),
size='medium')
# visualize parameter's effect on decision function
plt.pcolormesh(xx, yy, -Z, cmap=plt.cm.RdBu)
plt.scatter(X_2d[:, 0], X_2d[:, 1], c=y_2d, cmap=plt.cm.RdBu_r,
edgecolors='k')
plt.xticks(())
plt.yticks(())
plt.axis('tight')
scores = grid.cv_results_['mean_test_score'].reshape(len(C_range),
len(gamma_range))
# Draw heatmap of the validation accuracy as a function of gamma and C
#
# The score are encoded as colors with the hot colormap which varies from dark
# red to bright yellow. As the most interesting scores are all located in the
# 0.92 to 0.97 range we use a custom normalizer to set the mid-point to 0.92 so
# as to make it easier to visualize the small variations of score values in the
# interesting range while not brutally collapsing all the low score values to
# the same color.
plt.figure(figsize=(8, 6))
plt.subplots_adjust(left=.2, right=0.95, bottom=0.15, top=0.95)
plt.imshow(scores, interpolation='nearest', cmap=plt.cm.hot,
norm=MidpointNormalize(vmin=0.2, midpoint=0.92))
plt.xlabel('gamma')
plt.ylabel('C')
plt.colorbar()
plt.xticks(np.arange(len(gamma_range)), gamma_range, rotation=45)
plt.yticks(np.arange(len(C_range)), C_range)
plt.title('Validation accuracy')
plt.show()
```
| github_jupyter |
# Dataset Creation
we have files
- `resultsAnnotation.tsv`,
- `datasetAnnotation.tsv`,
- `taskAnnotation.tsv`,
- `paper_links.tsv`,
- `TDM_taxonomy.tsv`,
- `TDMs_taxonomy.tsv`
- `paper_name_taxonomy.tsv`
Created mostly from the file `evaluation-tables.json` from [paperswithcode](https://paperswithcode.com/about)
```
# imports
import ipdb, os, re
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score, f1_score, precision_score, recall_score, classification_report
with open(f"../data/resultsAnnotation.tsv", errors='replace') as f:
resultsAnnotation = f.read().splitlines()
with open(f"../data/datasetAnnotation.tsv", errors='replace') as f:
datasetAnnotation = f.read().splitlines()
with open(f"../data/taskAnnotation.tsv", errors='replace') as f:
taskAnnotation = f.read().splitlines()
with open(f"../data/TDM_taxonomy.tsv", errors='replace') as f:
TDM_taxonomy = f.read().splitlines()
with open(f"../data/paper_name_taxonomy.tsv", errors='replace') as f:
paper_name_taxonomy = f.read().splitlines()
resultsAnnotation[5]
datasetAnnotation[5]
taskAnnotation[5]
TDM_taxonomy[9]
paper_name_taxonomy[5]
def create_training_data(path_to_resultsAnnotation, path_to_TDM_taxonomy, path_parsed_files,
output_dir, test_set_portion=0.2,
leaderboard_threshold=5, num_negative_instances=5, allowed_unknown=10):
# to create the repo if it doesn't exist already
if not os.path.exists(f"{output_dir}{num_negative_instances}Neg{allowed_unknown}Unknown/"):
os.makedirs(f"{output_dir}{num_negative_instances}Neg{allowed_unknown}Unknown/")
with open(f"{path_to_resultsAnnotation}/resultsAnnotation.tsv", errors='replace') as f:
resultsAnnotation = f.read().splitlines()
paper_TDM = {}
for paper in resultsAnnotation:
if len(paper.split("\t")) != 2:
continue
title, TDMSList = paper.split("\t")
title = '.'.join(title.split('/')[-1].split('.')[:-1])
paper_TDM[title] = TDMSList
with open(f"{path_to_TDM_taxonomy}/TDM_taxonomy.tsv", errors='replace') as f:
TDM_taxonomy = f.read().splitlines()
TDM_taxonomy_dict = {}
unknown_count = 0
for TDMCount in TDM_taxonomy:
if len(TDMCount.split("\t")) != 2:
continue
TDM, count = TDMCount.split("\t")
count = int(count)
if count >= leaderboard_threshold:
TDM_taxonomy_dict[TDM] = count
# ipdb.set_trace()
list_parsed_pdf = os.listdir(path_parsed_files)
if '.ipynb_checkpoints' in list_parsed_pdf:
list_parsed_pdf.remove('.ipynb_checkpoints')
# ToDo: will it be interresting to use stratified ? using the label ?
train_valid = train_test_split(list_parsed_pdf, test_size=10/100, shuffle=True)
train, valid = train_valid[0], train_valid[1]
if os.path.exists(f"{output_dir}{num_negative_instances}Neg{allowed_unknown}Unknown/train.tsv"):
os.remove(f"{output_dir}{num_negative_instances}Neg{allowed_unknown}Unknown/train.tsv")
if os.path.exists(f"{output_dir}{num_negative_instances}Neg{allowed_unknown}Unknown/dev.tsv"):
os.remove(f"{output_dir}{num_negative_instances}Neg{allowed_unknown}Unknown/dev.tsv")
for paper in train :
with open(f"{path_parsed_files}{paper}", errors='replace') as f:
txt = f.read().splitlines()
content = ' '.join(txt)
# content = re.sub(r"[^a-zA-Z0-9?,'’‘´`%]+", ' ', content).strip()
content = re.sub(r"[\t]+", ' ', content).strip()
paper_id = '.'.join(paper.split('/')[-1].split('.')[:-1])
not_seen = True
if paper_id in paper_TDM.keys():
cache_tdm = set()
for contrib in paper_TDM[paper_id].split("$"):
if len(contrib.split("#")) != 4:
# missed += 1
continue
task, dataset, metric, score = contrib.split("#")
if (f"{task}#{dataset}#{metric}" in cache_tdm):
continue
if f"{task}#{dataset}#{metric}" in TDM_taxonomy_dict.keys():
not_seen = False
cache_tdm.add(f"{task}#{dataset}#{metric}")
with open(f"{output_dir}{num_negative_instances}Neg{allowed_unknown}Unknown/train.tsv", "a+", encoding="utf-8") as text_file:
text_file.write(f"true\t{paper_id}\t{task}#{dataset}#{metric}\t{content}\n")
if not_seen and (unknown_count <= allowed_unknown):
with open(f"{output_dir}{num_negative_instances}Neg{allowed_unknown}Unknown/train.tsv", "a+", encoding="utf-8") as text_file:
text_file.write(f"true\t{paper_id}\tunknown\t{content}\n")
random_tdm = list(TDM_taxonomy_dict.keys())
random_tdm.sort()
for RandTDM in random_tdm[:num_negative_instances]:
task, dataset, metric = RandTDM.split("#")
with open(f"{output_dir}{num_negative_instances}Neg{allowed_unknown}Unknown/train.tsv", "a+", encoding="utf-8") as text_file:
text_file.write(f"false\t{paper_id}\t{task}#{dataset}#{metric}\t{content}\n")
else:
print(f"Paper {paper_id} from train not in the resultsAnnotation.tssv file")
for paper in valid :
with open(f"{path_parsed_files}{paper}", errors='replace') as f:
txt = f.read().splitlines()
content = ' '.join(txt)
# content = re.sub(r"[^a-zA-Z0-9?,'’‘´`%]+", ' ', content).strip()
content = re.sub(r"[\t]+", ' ', content).strip()
paper_id = '.'.join(paper.split('/')[-1].split('.')[:-1])
not_seen = True
if paper_id in paper_TDM.keys():
cache_tdm = set()
for contrib in paper_TDM[paper_id].split("$"):
if len(contrib.split("#")) != 4:
# missed += 1
continue
task, dataset, metric, score = contrib.split("#")
if (f"{task}#{dataset}#{metric}" in cache_tdm):
continue
if f"{task}#{dataset}#{metric}" in TDM_taxonomy_dict.keys():
not_seen = False
cache_tdm.add(f"{task}#{dataset}#{metric}")
with open(f"{output_dir}{num_negative_instances}Neg{allowed_unknown}Unknown/dev.tsv", "a+", encoding="utf-8") as text_file:
text_file.write(f"true\t{paper_id}\t{task}#{dataset}#{metric}\t{content}\n")
if not_seen and (unknown_count <= allowed_unknown):
with open(f"{output_dir}{num_negative_instances}Neg{allowed_unknown}Unknown/dev.tsv", "a+", encoding="utf-8") as text_file:
text_file.write(f"true\t{paper_id}\tunknown\t{content}\n")
random_tdm = list(TDM_taxonomy_dict.keys())
random_tdm.sort()
for RandTDM in random_tdm[:num_negative_instances]:
task, dataset, metric = RandTDM.split("#")
with open(f"{output_dir}{num_negative_instances}Neg{allowed_unknown}Unknown/dev.tsv", "a+", encoding="utf-8") as text_file:
text_file.write(f"false\t{paper_id}\t{task}#{dataset}#{metric}\t{content}\n")
else:
print(f"Paper {paper_id} from validation not in the resultsAnnotation.tssv file")
path_grobid_full_txt = "/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/paperwithcode/pdf_txt/"
path_latex_source_tex = "/nfs/home/kabenamualus/Research/task-dataset-metric-nli-extraction/exp/arxiv_src/"
path_latex_source_pandoc_txt = "/nfs/home/kabenamualus/Research/task-dataset-metric-nli-extraction/exp/arxiv_src_txt/"
create_training_data(path_to_resultsAnnotation="../data/", \
path_to_TDM_taxonomy="../data/", path_parsed_files=path_grobid_full_txt,
output_dir="../data/",
leaderboard_threshold=5, num_negative_instances=60, allowed_unknown=800)
```
## View created data
```
import pandas as pd
train_csv = "/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/paperwithcode/new/60Neg800unk/twofoldwithunk/fold1/train.tsv"
valid_csv = "/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/paperwithcode/new/60Neg800unk/twofoldwithunk/fold1/dev.tsv"
train = pd.read_csv(train_csv,
sep="\t", names=["label", "title", "TDM", "Context"])
valid = pd.read_csv(valid_csv,
sep="\t", names=["label", "title", "TDM", "Context"])
train.head()
train.tail()
valid.head()
valid.tail()
train[train.title=="1911.08251v2"].head()
train[train.TDM=="unknown"].head()
# train["len"]=train.Context.apply(lambda content: len(content.split()),)
```
# Convert IBM Data to our Data format
This section mostly edit the TDM from the original data from [Hou et al. (2019)](https://arxiv.org/pdf/2004.14356.pdf) by replacing the separator from `,` to `;`.
**Note**: This section of the notebook can only be run once.
## NLP-TDMS
### main
```
IBM_train_csv = "~/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/train.tsv"
IBM_test_csv = "~/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/test.tsv"
# ! cp $IBM_train_csv ~/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/train_.tsv
# ! cp $IBM_test_csv ~/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/test_.tsv
train_IBM = pd.read_csv(IBM_train_csv,
sep="\t", names=["label", "title", "TDM", "Context"])
test_IBM = pd.read_csv(IBM_test_csv,
sep="\t", names=["label", "title", "TDM", "Context"])
train_IBM.head()
test_IBM.head()
train_IBM["TDM"] = train_IBM.TDM.apply(lambda x : re.sub(r"[,]+", ';', x).strip())
test_IBM["TDM"] = test_IBM.TDM.apply(lambda x : re.sub(r"[,]+", ';', x).strip())
train_IBM["TDM"] = train_IBM.TDM.apply(lambda x : "unknown" if x == "unknow" else x)
test_IBM["TDM"] = test_IBM.TDM.apply(lambda x : "unknown" if x == "unknow" else x)
train_IBM["label"] = train_IBM.label.apply(lambda x : str(x).lower())
test_IBM["label"] = test_IBM.label.apply(lambda x : str(x).lower())
train_IBM.head()
train_IBM.tail()
test_IBM.tail()
train_IBM.to_csv(path_or_buf=IBM_train_csv,
sep="\t", header=None, index=False)
test_IBM.to_csv(path_or_buf=IBM_test_csv,
sep="\t", header=None, index=False)
train_v2_IBM = pd.read_csv(IBM_train_csv,
sep="\t", names=["label", "title", "TDM", "Context"])
test_v2_IBM = pd.read_csv(IBM_test_csv,
sep="\t", names=["label", "title", "TDM", "Context"])
train_v2_IBM.head()
test_v2_IBM.tail()
```
### paperVersion
```
IBM_train_csv = "~/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/train.tsv"
IBM_test_csv = "~/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/test.tsv"
# ! cp $IBM_train_csv ~/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/train_.tsv
# ! cp $IBM_test_csv ~/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/test_.tsv
train_IBM = pd.read_csv(IBM_train_csv,
sep="\t", names=["label", "title", "TDM", "Context"])
test_IBM = pd.read_csv(IBM_test_csv,
sep="\t", names=["label", "title", "TDM", "Context"])
train_IBM.head()
test_IBM.head()
train_IBM["TDM"] = train_IBM.TDM.apply(lambda x : re.sub(r"[,]+", ';', x).strip())
test_IBM["TDM"] = test_IBM.TDM.apply(lambda x : re.sub(r"[,]+", ';', x).strip())
train_IBM["TDM"] = train_IBM.TDM.apply(lambda x : "unknown" if x == "unknow" else x)
test_IBM["TDM"] = test_IBM.TDM.apply(lambda x : "unknown" if x == "unknow" else x)
train_IBM["label"] = train_IBM.label.apply(lambda x : str(x).lower())
test_IBM["label"] = test_IBM.label.apply(lambda x : str(x).lower())
train_IBM.head()
train_IBM.tail()
test_IBM.tail()
train_IBM.to_csv(path_or_buf=IBM_train_csv,
sep="\t", header=None, index=False)
test_IBM.to_csv(path_or_buf=IBM_test_csv,
sep="\t", header=None, index=False)
train_v2_IBM = pd.read_csv(IBM_train_csv,
sep="\t", names=["label", "title", "TDM", "Context"])
test_v2_IBM = pd.read_csv(IBM_test_csv,
sep="\t", names=["label", "title", "TDM", "Context"])
train_v2_IBM.head()
test_v2_IBM.tail()
```
## ARC-PDN
```
!tar -xvzf ~/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/ARC-PDN/test_pdn.tsv.tar.gz -C ~/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/ARC-PDN/
IBM_test_pdn_csv = "~/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/ARC-PDN/test_pdn.tsv"
IBM_test_pdn_score_csv = "~/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/ARC-PDN/test_pdn_score.tsv"
# ! cp $IBM_test_pdn_csv ~/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/ARC-PDN/test_pdn_.tsv
# ! cp $IBM_test_pdn_score_csv ~/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/ARC-PDN/test_pdn_score_.tsv
test_pdn_IBM = pd.read_csv(IBM_train_csv,
sep="\t", names=["label", "title", "TDM", "Context"])
test_pdn_score_IBM = pd.read_csv(IBM_test_csv,
sep="\t", names=["label", "title", "TDM", "Context"])
test_pdn_IBM.head()
test_pdn_score_IBM.head()
train_IBM["TDM"] = train_IBM.TDM.apply(lambda x : re.sub(r"[,]+", ';', x).strip())
test_IBM["TDM"] = test_IBM.TDM.apply(lambda x : re.sub(r"[,]+", ';', x).strip())
train_IBM["label"] = train_IBM.label.apply(lambda x : str(x).lower())
test_IBM["label"] = test_IBM.label.apply(lambda x : str(x).lower())
train_IBM.head()
train_IBM.tail()
test_IBM.tail()
train_IBM.to_csv(path_or_buf=IBM_train_csv,
sep="\t", header=None, index=False)
test_IBM.to_csv(path_or_buf=IBM_test_csv,
sep="\t", header=None, index=False)
train_v2_IBM = pd.read_csv(IBM_train_csv,
sep="\t", names=["label", "title", "TDM", "Context"])
test_v2_IBM = pd.read_csv(IBM_test_csv,
sep="\t", names=["label", "title", "TDM", "Context"])
train_v2_IBM.head()
test_v2_IBM.tail()
```
## Zero-shot
```
IBM_train_csv = "~/Research/task-dataset-metric-nli-extraction/data/ibm/exp/zero-shot-setup/NLP-TDMS/train.tsv"
IBM_test_csv = "~/Research/task-dataset-metric-nli-extraction/data/ibm/exp/zero-shot-setup/NLP-TDMS/test.tsv"
! cp $IBM_train_csv ~/Research/task-dataset-metric-nli-extraction/data/ibm/exp/zero-shot-setup/NLP-TDMS/train_.tsv
! cp $IBM_test_csv ~/Research/task-dataset-metric-nli-extraction/data/ibm/exp/zero-shot-setup/NLP-TDMS/test_.tsv
train_IBM = pd.read_csv(IBM_train_csv,
sep="\t", names=["label", "title", "TDM", "Context"])
test_IBM = pd.read_csv(IBM_test_csv,
sep="\t", names=["label", "title", "TDM", "Context"])
train_IBM.head()
test_IBM.head()
train_IBM["TDM"] = train_IBM.TDM.apply(lambda x : re.sub(r"[,]+", ';', x).strip())
test_IBM["TDM"] = test_IBM.TDM.apply(lambda x : re.sub(r"[,]+", ';', x).strip())
train_IBM["label"] = train_IBM.label.apply(lambda x : str(x).lower())
test_IBM["label"] = test_IBM.label.apply(lambda x : str(x).lower())
train_IBM.head()
train_IBM.tail()
test_IBM.tail()
train_IBM.to_csv(path_or_buf=IBM_train_csv,
sep="\t", header=None, index=False)
test_IBM.to_csv(path_or_buf=IBM_test_csv,
sep="\t", header=None, index=False)
train_v2_IBM = pd.read_csv(IBM_train_csv,
sep="\t", names=["label", "title", "TDM", "Context"])
test_v2_IBM = pd.read_csv(IBM_test_csv,
sep="\t", names=["label", "title", "TDM", "Context"])
train_v2_IBM.head()
test_v2_IBM.tail()
IBM_train_csv = "~/Research/task-dataset-metric-nli-extraction/data/pwc_ibm_150_10_10_1000"
IBM_test_csv = "~/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/test.tsv"
# ! cp $IBM_train_csv ~/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/train_.tsv
# ! cp $IBM_test_csv ~/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/test_.tsv
train_IBM = pd.read_csv(IBM_train_csv,
sep="\t", names=["label", "title", "TDM", "Context"])
test_IBM = pd.read_csv(IBM_test_csv,
sep="\t", names=["label", "title", "TDM", "Context"])
train_IBM.head()
test_IBM.head()
train_IBM["TDM"] = train_IBM.TDM.apply(lambda x : re.sub(r"[,]+", ';', x).strip())
test_IBM["TDM"] = test_IBM.TDM.apply(lambda x : re.sub(r"[,]+", ';', x).strip())
train_IBM["TDM"] = train_IBM.TDM.apply(lambda x : "unknown" if x == "unknow" else x)
test_IBM["TDM"] = test_IBM.TDM.apply(lambda x : "unknown" if x == "unknow" else x)
train_IBM["label"] = train_IBM.label.apply(lambda x : str(x).lower())
test_IBM["label"] = test_IBM.label.apply(lambda x : str(x).lower())
train_IBM.head()
train_IBM.tail()
test_IBM.tail()
train_IBM.to_csv(path_or_buf=IBM_train_csv,
sep="\t", header=None, index=False)
test_IBM.to_csv(path_or_buf=IBM_test_csv,
sep="\t", header=None, index=False)
train_v2_IBM = pd.read_csv(IBM_train_csv,
sep="\t", names=["label", "title", "TDM", "Context"])
test_v2_IBM = pd.read_csv(IBM_test_csv,
sep="\t", names=["label", "title", "TDM", "Context"])
train_v2_IBM.head()
test_v2_IBM.tail()
```
## DatasetAnnotation
```
datasetAnnotation_csv = "/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/paperwithcode/annotations/datasetAnnotation.tsv"
resultsAnnotation_csv = "/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/paperwithcode/annotations/resultsAnnotation.tsv"
taskAnnotation_csv = "/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/paperwithcode/annotations/taskAnnotation.tsv"
datasetAnnotation_IBM_csv = "/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/ibm/NLP-TDMS/annotations/datasetAnnotation.tsv"
resultsAnnotation_IBM_csv = "/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/ibm/NLP-TDMS/annotations/resultsAnnotation.tsv"
taskAnnotation_IBM_csv = "/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/ibm/NLP-TDMS/annotations/taskAnnotation.tsv"
datasetAnnotation = pd.read_csv(datasetAnnotation_csv,
sep="\t", names=["label", "datasets"])
resultsAnnotation = pd.read_csv(resultsAnnotation_csv,
sep="\t", names=["label", "TDMS"])
taskAnnotation = pd.read_csv(taskAnnotation_csv,
sep="\t", names=["label", "tasks"])
datasetAnnotation_IBM = pd.read_csv(datasetAnnotation_IBM_csv,
sep="\t", names=["label", "datasets"])
resultsAnnotation_IBM = pd.read_csv(resultsAnnotation_IBM_csv,
sep="\t", names=["label", "TDMS"])
taskAnnotation_IBM = pd.read_csv(taskAnnotation_IBM_csv,
sep="\t", names=["label", "tasks"])
datasetAnnotation_IBM.head()
datasetAnnotation.head()
resultsAnnotation.tail()
resultsAnnotation_IBM.tail()
taskAnnotation.tail()
taskAnnotation_IBM.tail()
with open(f"/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/test.tsv") as f:
list_test_IBM = f.read().splitlines()
# with open(f"/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/test_results_IBM.tsv") as f:
with open(f"/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/paperwithcode/new/60Neg800unk/twofoldwithunk/fold1/models/SciBERT/test_results.tsv") as f:
# with open(f"/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/paperwithcode/new/jar/80Neg600unk/twofoldwithunk/fold1/models/cased_L-12_H-768_A-12_XLNet/test_results.tsv") as f:
list_test_IBM_results = f.read().splitlines()
len(list_test_IBM_results)
assert len(list_test_IBM) == len(list_test_IBM_results), "Error in Lenght"
list_test_IBM[-1].split("\t")[0]
list_test_IBM_results[0].split("\t")
tp, fn, tn, fp = 0, 0, 0, 0
y = []
y_pred = []
# here 0: true, 1: false
for idx in range(len(list_test_IBM)):
true_label = list_test_IBM[idx].split("\t")[0]
true, false = list_test_IBM_results[idx].split("\t")
true, false = float(true), float(false)
if true_label=='true' :
y.append(0)
if true > false:
tp += 1
y_pred.append(0)
else:
fn += 1
y_pred.append(1)
else:
y.append(1)
if false > true:
tn += 1
y_pred.append(1)
else:
fp += 1
y_pred.append(0)
# print(f"TP: {tp} FN: {fn} TN: {tn} FP: {fp}")
# precision = tp/(tp+fp)
# recall = tp/(tp+fn)
# f1 = 2 * (precision * recall) / (precision + recall)
# print(f"Precision: {precision} Recall: {recall} and F1 = {f1}")
# classification_report(y, y_pred, target_names=["true", "false"], output_dict=True)
print(f"precision_score (macro): {precision_score(y, y_pred, average ='macro')}")
print(f"recall_score (macro): {recall_score(y, y_pred, average ='macro')}")
print(f"f1_score (macro): {f1_score(y, y_pred, average ='macro')}")
print(f"precision_score (micro): {precision_score(y, y_pred, average ='micro')}")
print(f"recall_score (micro): {recall_score(y, y_pred, average ='micro')}")
print(f"f1_score (micro): {f1_score(y, y_pred, average ='micro')}")
print(f"Avg. {round((f1_score(y, y_pred, average ='macro')+f1_score(y, y_pred, average ='micro'))/2, 4)}")
# metrics_dict = classification_report(y, y_pred, target_names=["true", "false"], output_dict=True)['macro avg']
# print(f"precision macro avg: {metrics_dict['precision']}")
# print(f"recall macro avg: {metrics_dict['recall']}")
# print(f"f1-score macro avg: {metrics_dict['f1-score']}")
print(f"precision_score (macro): {precision_score(y, y_pred, average ='macro')}")
print(f"recall_score (macro): {recall_score(y, y_pred, average ='macro')}")
print(f"f1_score (macro): {f1_score(y, y_pred, average ='macro')}")
print(f"precision_score (micro): {precision_score(y, y_pred, average ='micro')}")
print(f"recall_score (micro): {recall_score(y, y_pred, average ='micro')}")
print(f"f1_score (micro): {f1_score(y, y_pred, average ='micro')}")
print(f"Avg. {round((f1_score(y, y_pred, average ='macro')+f1_score(y, y_pred, average ='micro'))/2, 3)}")
# metrics_dict = classification_report(y, y_pred, target_names=["true", "false"], output_dict=True)['macro avg']
# print(f"precision macro avg: {metrics_dict['precision']}")
# print(f"recall macro avg: {metrics_dict['recall']}")
# print(f"f1-score macro avg: {metrics_dict['f1-score']}")
round(0.9620534006579451, 3)
```
# Data clearning
### paperVerion
```
test_IBM_pd = pd.read_csv(f"/nfs/home/kabenamualus/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/test.tsv",
sep="\t", names=["label", "title", "TDM", "Context"])
train_IBM_pd = pd.read_csv(f"/nfs/home/kabenamualus/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/train.tsv",
sep="\t", names=["label", "title", "TDM", "Context"])
paper_links_pd = pd.read_csv(f"/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/ibm/NLP-TDMS/downloader/paper_links.tsv",
sep="\t", names=["pdf", "link", "hashCode"])
path_pdf_IBM_Original = "/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM_Original"
train_IBM_pd.tail()
test_IBM_pd.tail()
paper_links_pd.head()
# path_pdf_IBM_Original
# test_IBM_pd[test_IBM_pd.label==True].head()
# List of pdf name that are in the test file
list_train_IBM_uniq = list(train_IBM_pd.title.unique())
len(list_train_IBM_uniq)
# List of pdf name that are in the train file
list_test_IBM_uniq = list(test_IBM_pd.title.unique())
len(list_test_IBM_uniq)
# total number of uniq pdf in the train and testing
len(list_test_IBM_uniq)+len(list_train_IBM_uniq)
# List of number unique file in main pdf folder
list_paper_links_uniq = list(paper_links_pd.pdf.unique())
len(list_paper_links_uniq)
list_pdf_IBM_Original = os.listdir("/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM_Original")
len(list_pdf_IBM_Original)
```
```
count = 0
missing_list = []
for pdf in list_test_IBM_uniq:
# if pdf not in list_pdf_IBM_Original:
if pdf not in list_pdf_IBM_Original:
# print(pdf)
count+=1
missing_list.append(pdf)
print(count)
missing_list
count = 0
missing_list = []
for pdf in list_pdf_IBM_Original:
# if pdf not in list_pdf_IBM_Original:
if pdf not in list_test_IBM_uniq and pdf not in list_train_IBM_uniq:
# print(pdf)
count+=1
missing_list.append(pdf)
print(count)
# The following pdf don't have taxonomy in the original IBM released metadata
missing_list
pdf_path = "/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM_Original/"
output_path = "/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM_Original_Clean/"
if not (os.path.exists(output_path)):
os.makedirs(output_path)
for pdf in list_pdf_IBM_Original:
if pdf not in list_test_IBM_uniq and pdf not in list_train_IBM_uniq:
print(f"Pdf {pdf} not used in train nor test, skiped")
else:
!cp $pdf_path$pdf $output_path
len(os.listdir(pdf_path))==len(os.listdir(output_path))+len(missing_list)
# len(os.listdir(pdf_path))==len(os.listdir(output_path))+len(missing_list)
count = 0
missing_list = []
for pdf in list_pdf_IBM_Original:
# if pdf not in list_pdf_IBM_Original:
if pdf not in list_paper_links_uniq:
# print(pdf)
count+=1
missing_list.append(pdf)
print(count)
import ipdb
# for pdf in missing_list:
# print(paper_links_pd[paper_links_pd.pdf==pdf])
# ipdb.set_trace()
```
### Create separate train, test folder as per the original data split
```
pdf_path = "/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM_Original/"
output_path = "/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM_Test_Paper_Version/"
for pdf in list_test_IBM_uniq:
!cp $pdf_path$pdf $output_path
len(os.listdir("/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM_Test_Paper_Version"))==len(list_test_IBM_uniq)
pdf_path = "/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM_Original/"
output_path = "/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM_Train_Paper_Version/"
for pdf in list_train_IBM_uniq:
!cp $pdf_path$pdf $output_path
len(os.listdir("/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM_Train_Paper_Version"))==len(list_train_IBM_uniq)-2
# Remove test pdf in the main dataset folder
len(os.listdir("/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM/"))
len(list_test_IBM_uniq)
len(os.listdir("/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM/"))-len(list_test_IBM_uniq)
pdf_path = "/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM/"
output_path = "/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM_Train_Paper_Version/"
for pdf in os.listdir(pdf_path):
if pdf in list_test_IBM_uniq:
!rm $pdf_path$pdf
len(os.listdir(pdf_path))
pdf_path = "/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM/"
pdf_train_path = "/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM_Train_Paper_Version/"
for pdf in os.listdir(pdf_train_path):
!cp $pdf_train_path$pdf $pdf_path
# len(os.listdir("/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM_Train_Paper_Version"))==len(list_train_IBM_uniq)-2
len(os.listdir(pdf_path))
count = 0
missing_list = []
for pdf in list_train_IBM_uniq:
if pdf not in os.listdir(pdf_path):
print(pdf)
count+=1
missing_list.append(pdf)
print(count)
missing_list
```
# Zero shot setup
```
test2_IBM_pd = pd.read_csv(f"/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/test.tsv",
sep="\t", names=["label", "title", "TDM", "Context"])
train2_IBM_pd = pd.read_csv(f"/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/train.tsv",
sep="\t", names=["label", "title", "TDM", "Context"])
# paper_links_pd = pd.read_csv(f"/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/ibm/NLP-TDMS/downloader/paper_links.tsv",
# sep="\t", names=["pdf", "link", "hashCode"])
len(list_train_IBM_uniq)
list_train2_IBM_uniq = list(train2_IBM_pd.title.unique())
len(list_train2_IBM_uniq)
len(list_test_IBM_uniq)
# List of pdf name that are in the train file
list_test2_IBM_uniq = list(test2_IBM_pd.title.unique())
len(list_test_IBM_uniq)
count = 0
missing_list = []
for pdf in list_test2_IBM_uniq:
# if pdf not in list_pdf_IBM_Original:
if pdf not in list_pdf_IBM_Original:
# print(pdf)
count+=1
missing_list.append(pdf)
print(count)
```
## Check the balance between classes
### IBM
```
test_IBM_pd = pd.read_csv(f"/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/test.tsv",
sep="\t", names=["label", "title", "TDM", "Context"])
train_IBM_pd = pd.read_csv(f"/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/train.tsv",
sep="\t", names=["label", "title", "TDM", "Context"])
train_IBM_pd.head()
train_IBM_pd[train_IBM_pd.label==True].count()
train_IBM_pd[train_IBM_pd.label==True].nunique()
# train_IBM_pd_true = train_IBM_pd[train_IBM_pd.label==True]
# train_IBM_pd_true[train_IBM_pd_true.title.duplicated()]
train_IBM_pd[train_IBM_pd.label==False].count()
```
## Update the IBM train and test data to our generated context
```
train_IBM_pd = pd.read_csv(f"/nfs/home/kabenamualus/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/train.tsv",
sep="\t", names=["label", "title", "TDM", "Context"])
test_IBM_pd = pd.read_csv(f"/nfs/home/kabenamualus/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/test.tsv",
sep="\t", names=["label", "title", "TDM", "Context"])
trainOutput_150_IBM_pd = pd.read_csv(f"/nfs/home/kabenamualus/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/IBM_jar_150/0Neg1000unk/trainOutput.tsv",
sep="\t", names=["label", "title", "TDM", "Context"])
trainOutput_full_IBM_pd = pd.read_csv(f"/nfs/home/kabenamualus/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/IBM_jar_full/0Neg1000unk/trainOutput.tsv",
sep="\t", names=["label", "title", "TDM", "Context"])
# path_train_150 = "/nfs/home/kabenamualus/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/train_150.tsv"
# path_train_full = "/nfs/home/kabenamualus/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/train_full.tsv"
trainOutput_150_IBM_pd.tail()
trainOutput_full_IBM_pd.head()
```
### train_IBM_pd vs trainOutput_150_IBM_pd
```
train_IBM_pd[train_IBM_pd.label==True].head()
trainOutput_150_IBM_pd.head()
train_IBM_pd.title.unique()[:2]
# list_trainOutput_150_IBM_pd_uniq[:2]
# List of pdf name that are in the test file
list_train_IBM_uniq = list(train_IBM_pd.title.unique())
len(list_train_IBM_uniq)
# List of pdf name that are in the train file
list_test_IBM_uniq = list(test_IBM_pd.title.unique())
len(list_test_IBM_uniq)
# total number of uniq pdf in the train and testing
len(list_test_IBM_uniq)+len(list_train_IBM_uniq)
# List of pdf name that are in the train file
list_trainOutput_150_IBM_pd_uniq = list(trainOutput_150_IBM_pd.title.unique())
len(list_trainOutput_150_IBM_pd_uniq)
# List of pdf name that are in the train file
list_trainOutput_full_IBM_pd_uniq = list(trainOutput_full_IBM_pd.title.unique())
len(list_trainOutput_full_IBM_pd_uniq)
# Dict to contains new context
dict_150_paper_context = {}
for paper in list_trainOutput_150_IBM_pd_uniq:
dict_150_paper_context[paper]=trainOutput_150_IBM_pd[trainOutput_150_IBM_pd.title==paper].Context.values[0]
dict_original_paper_context = {}
for paper in list_train_IBM_uniq:
dict_original_paper_context[paper]=train_IBM_pd[train_IBM_pd.title==paper].Context.values[0]
for paper in list_test_IBM_uniq:
dict_original_paper_context[paper]=test_IBM_pd[test_IBM_pd.title==paper].Context.values[0]
dict_full_paper_context = {}
for paper in list_trainOutput_full_IBM_pd_uniq:
dict_full_paper_context[paper]=trainOutput_full_IBM_pd[trainOutput_full_IBM_pd.title==paper].Context.values[0]
def get_start_lenght(dictionary, limit="150", title="",):
# Stats
len_context= []
for context in dictionary.values():
len_context.append(len(context.split()))
print(f"Context TDM limit {limit}:")
print(f"Mean lenght: {np.mean(len_context)}")
print(f"Max lenght: {np.max(len_context)}")
print(f"Min lenght: {np.min(len_context)}")
print(f"Std lenght: {np.std(len_context)}")
x = np.arange(1, len(len_context)+1, 1)
y = len_context
plt.plot(x, y)
plt.title(title)
plt.xlabel("number of papers")
plt.ylabel("lenght DocTAET")
plt.savefig(fname=re.sub(r"[0-9]+", '', title).strip())
plt.show()
return len_context
len_context_150 = get_start_lenght(dict_150_paper_context,
limit="150",
title="Original DocTAET Hou et al. 2019")
len_context_original = get_start_lenght(dict_original_paper_context,
limit="IBM Original",
title="Original DocTAET Hou et al. 2019")
len_context_full = get_start_lenght(dict_full_paper_context,
limit="full",
title="Extended DocTAET")
train_IBM_pd.head()
np.arange(1, 5, 1)
# To generate an array of x-values
x = np.arange(1, len(len_context_original)+1, 1)
# To generate an array of
# y-values using corresponding x-values
y = len_context_original
# def pdf(x):
# mean = np.mean(x)
# std = np.std(x)
# y_out = 1/(std * np.sqrt(2 * np.pi)) * np.exp( - (x - mean)**2 / (2 * std**2))
# return y_out
# # To generate an array of x-values
# x = np.arange(-2, 2, 0.1)
# # To generate an array of
# # y-values using corresponding x-values
# y = pdf(x)
# Plotting the bell-shaped curve
plt.style.use('seaborn')
plt.figure(figsize = (6, 6))
plt.plot(x, y, color = 'black',
linestyle = ':')
plt.xlabel("len tokens")
plt.ylabel("Papers count")
plt.scatter( x, y, marker = 'o', s = 25, color = 'red')
plt.show()
import numpy as np
import matplotlib.pyplot as plt
x = np.arange(1, len(len_context_original)+1, 1)
y = len_context_original
plt.plot(x, y)
plt.title("Sports Watch Data")
plt.xlabel("Average Pulse")
plt.ylabel("Calorie Burnage")
plt.savefig("test")
plt.show()
import numpy as np
import matplotlib.pyplot as plt
x = np.arange(1, len(len_context_full)+1, 1)
y = len_context_full
plt.plot(x, y)
plt.title("Sports Watch Data")
plt.xlabel("Average Pulse")
plt.ylabel("Calorie Burnage")
plt.show()
# To generate an array of x-values
x = np.arange(1, len(len_context_full)+1, 1)
# To generate an array of
# y-values using corresponding x-values
y = len_context_full
# Plotting the bell-shaped curve
plt.style.use('seaborn')
plt.figure(figsize = (6, 6))
plt.plot(x, y, color = 'black',
linestyle = ':')
plt.scatter( x, y, marker = 'o', s = 25, color = 'red')
plt.show()
train_IBM_pd["Context"] = train_IBM_pd.apply(lambda x : dict_full_paper_context[x['title']] if x['title'] in dict_full_paper_context.keys() else "None", axis=1)
test_IBM_pd["Context"] = test_IBM_pd.apply(lambda x : dict_full_paper_context[x['title']] if x['title'] in dict_full_paper_context.keys() else "None", axis=1)
# Verification
dict_original_paper_context = {}
for paper in list_train_IBM_uniq:
dict_original_paper_context[paper]=train_IBM_pd[train_IBM_pd.title==paper].Context.values[0]
for paper in list_test_IBM_uniq:
dict_original_paper_context[paper]=test_IBM_pd[test_IBM_pd.title==paper].Context.values[0]
len_context_original = get_start_lenght(dict_original_paper_context, limit="IBM Original")
path_train_full = "/nfs/home/kabenamualus/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/train_full.tsv"
path_test_full = "/nfs/home/kabenamualus/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/test_full.tsv"
train_IBM_pd.to_csv(path_or_buf=path_train_full,
sep="\t", header=None, index=False)
test_IBM_pd.to_csv(path_or_buf=path_test_full,
sep="\t", header=None, index=False)
```
### Zero-shot
```
train_IBM_zero_shot_pd = pd.read_csv(f"/nfs/home/kabenamualus/Research/task-dataset-metric-nli-extraction/data/ibm/exp/zero-shot-setup/NLP-TDMS/train.tsv",
sep="\t", names=["label", "title", "TDM", "Context"])
test_IBM_zero_shot_pd = pd.read_csv(f"/nfs/home/kabenamualus/Research/task-dataset-metric-nli-extraction/data/ibm/exp/zero-shot-setup/NLP-TDMS/test.tsv",
sep="\t", names=["label", "title", "TDM", "Context"])
trainOutput_150_IBM_pd = pd.read_csv(f"/nfs/home/kabenamualus/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/IBM_jar_150/0Neg1000unk/trainOutput.tsv",
sep="\t", names=["label", "title", "TDM", "Context"])
trainOutput_full_IBM_pd = pd.read_csv(f"/nfs/home/kabenamualus/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/IBM_jar_full/0Neg1000unk/trainOutput.tsv",
sep="\t", names=["label", "title", "TDM", "Context"])
# List of pdf name that are in the test file
list_train_IBM_zero_shot_uniq = list(train_IBM_zero_shot_pd.title.unique())
len(list_train_IBM_zero_shot_uniq)
# List of pdf name that are in the train file
list_test_IBM_zero_shot_uniq = list(test_IBM_zero_shot_pd.title.unique())
len(list_test_IBM_zero_shot_uniq)
# # Dict to contain new context
# dict_150_paper_zero_shot_context = {}
# for paper in list_trainOutput_150_IBM_pd_uniq:
# dict_150_paper_context[paper]=trainOutput_150_IBM_pd[trainOutput_150_IBM_pd.title==paper].Context.values[0]
dict_original_paper_zero_shot_context = {}
for paper in list_train_IBM_zero_shot_uniq:
dict_original_paper_zero_shot_context[paper]=train_IBM_zero_shot_pd[train_IBM_zero_shot_pd.title==paper].Context.values[0]
for paper in list_test_IBM_zero_shot_uniq:
dict_original_paper_zero_shot_context[paper]=test_IBM_zero_shot_pd[test_IBM_zero_shot_pd.title==paper].Context.values[0]
len_context_original_zero_shot = get_start_lenght(dict_original_paper_zero_shot_context, limit="IBM Original Zero-shot")
train_IBM_zero_shot_pd["Context"] = train_IBM_zero_shot_pd.apply(lambda x : dict_full_paper_context[x['title']] if x['title'] in dict_full_paper_context.keys() else "None", axis=1)
test_IBM_zero_shot_pd["Context"] = test_IBM_zero_shot_pd.apply(lambda x : dict_full_paper_context[x['title']] if x['title'] in dict_full_paper_context.keys() else "None", axis=1)
# Verification
dict_original_paper_zero_shot_context = {}
for paper in list_train_IBM_zero_shot_uniq:
dict_original_paper_zero_shot_context[paper]=train_IBM_zero_shot_pd[train_IBM_zero_shot_pd.title==paper].Context.values[0]
for paper in list_test_IBM_zero_shot_uniq:
dict_original_paper_zero_shot_context[paper]=test_IBM_zero_shot_pd[test_IBM_zero_shot_pd.title==paper].Context.values[0]
len_context_original_zero_shot = get_start_lenght(dict_original_paper_zero_shot_context, limit="IBM Original Zero-shot extended")
path_train_full = "/nfs/home/kabenamualus/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/train_full.tsv"
path_test_full = "/nfs/home/kabenamualus/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/test_full.tsv"
train_IBM_pd.to_csv(path_or_buf=path_train_full,
sep="\t", header=None, index=False)
test_IBM_pd.to_csv(path_or_buf=path_test_full,
sep="\t", header=None, index=False)
```
### ARC-PDN
```
train_IBM_ARC_PDN_pd = pd.read_csv(f"/nfs/home/kabenamualus/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/ARC-PDN/test_pdn.tsv",
sep="\t", names=["label", "title", "TDM", "Context"])
train_IBM_ARC_PDN_pd.tail()
# List of pdf name that are in the test file
list_train_IBM_ARC_PDN_pd_uniq = list(train_IBM_ARC_PDN_pd.title.unique())
len(list_train_IBM_ARC_PDN_pd_uniq)
# # Dict to contain new context
# dict_150_paper_zero_shot_context = {}
# for paper in list_trainOutput_150_IBM_pd_uniq:
# dict_150_paper_context[paper]=trainOutput_150_IBM_pd[trainOutput_150_IBM_pd.title==paper].Context.values[0]
dict_original_IBM_ARC_PDN_context = {}
for paper in list_train_IBM_ARC_PDN_pd_uniq:
dict_original_IBM_ARC_PDN_context[paper]=train_IBM_ARC_PDN_pd[train_IBM_ARC_PDN_pd.title==paper].Context.values[0]
len_context_original_ARC_PDN = get_start_lenght(dict_original_IBM_ARC_PDN_context, limit="IBM Original ARC-PDN")
len(train_IBM_ARC_PDN_pd.Context.values[0].split())
train_IBM_ARC_PDN_pd[train_IBM_ARC_PDN_pd.Context == " "].head()
train_IBM_ARC_PDN_pd["Context"] = train_IBM_ARC_PDN_pd.apply(lambda x : dict_full_paper_context[x['title']] if x['title'] in dict_full_paper_context.keys() else "None", axis=1)
# test_IBM_zero_shot_pd["Context"] = test_IBM_zero_shot_pd.apply(lambda x : dict_full_paper_context[x['title']] if x['title'] in dict_full_paper_context.keys() else "None", axis=1)
dict_original_IBM_ARC_PDN_context = {}
for paper in list_train_IBM_ARC_PDN_pd_uniq:
dict_original_IBM_ARC_PDN_context[paper]=train_IBM_ARC_PDN_pd[train_IBM_ARC_PDN_pd.title==paper].Context.values[0]
len_context_original_ARC_PDN = get_start_lenght(dict_original_IBM_ARC_PDN_context, limit="IBM Original ARC-PDN")
path_train_full = "/nfs/home/kabenamualus/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/train_full.tsv"
path_test_full = "/nfs/home/kabenamualus/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/test_full.tsv"
train_IBM_pd.to_csv(path_or_buf=path_train_full,
sep="\t", header=None, index=False)
test_IBM_pd.to_csv(path_or_buf=path_test_full,
sep="\t", header=None, index=False)
# Big dataset problematic resources
# Max lenght exceed 10000 Grobid limit
Processing file: /nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM/1810.06682v2.pdf
/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM/1504.08200.pdf
/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM/1907.09665v10.pdf
/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM/2002.02050v3.pdf
/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM/2002.02050v3.pdf
/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM/1911.10807v3.pdf
/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM/1909.13476v1.pdf
/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM/1711.07566v1.pdf
/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM/download_pdfs.py
/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM/1804.01508v10.pdf
/nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM/1812.01527v2.pdf
!mv /nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM/download_pdfs.py /nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM/download_pdfs_2.py
# !rm /nfs/home/kabenamualus/Research/task-dataset-metric-extraction/data/pdf_IBM/1810.06682v2.pdf
```
## Create data stats visualization
```
IBM_train_csv = "~/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/train.tsv"
IBM_test_csv = "~/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/test.tsv"
IBM_train_full_csv = "/nfs/home/kabenamualus/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/train_full.tsv"
IBM_test_full_csv = "/nfs/home/kabenamualus/Research/task-dataset-metric-nli-extraction/data/ibm/exp/few-shot-setup/NLP-TDMS/paperVersion/test_full.tsv"
train_IBM = pd.read_csv(IBM_train_csv,
sep="\t", names=["label", "title", "TDM", "Context"])
train_full_IBM = pd.read_csv(IBM_train_full_csv,
sep="\t", names=["label", "title", "TDM", "Context"])
# test_IBM = pd.read_csv(IBM_test_csv,
# sep="\t", names=["label", "title", "TDM", "Context"])
train_IBM[(train_IBM.label==True) & (train_IBM.TDM!="unknown")].head(30)
selected_paper = "D17-1222.pdf"
train_IBM[(train_IBM.label==True) & (train_IBM.title==selected_paper)].tail(20)
train_IBM[train_IBM.title==selected_paper].Context.values[0]
train_full_IBM[train_full_IBM.title==selected_paper].Context.values[0]
```
## Check that the newly generated data doesnt link
```
PWC_train_csv = "/nfs/home/kabenamualus/Research/task-dataset-metric-nli-extraction/data/pwc_ibm_150_5_10_800/twofoldwithunk/fold1/train.tsv"
PWC_valid_csv = "/nfs/home/kabenamualus/Research/task-dataset-metric-nli-extraction/data/pwc_ibm_150_5_10_800/twofoldwithunk/fold1/dev.tsv"
train_PWC = pd.read_csv(PWC_train_csv,
sep="\t", names=["label", "title", "TDM", "Context"])
valid_PWC = pd.read_csv(PWC_valid_csv,
sep="\t", names=["label", "title", "TDM", "Context"])
train_PWC[train_PWC.label==True].title.unique()
set_train_unique = set(train_PWC[train_PWC.label==True].title.unique())
set_valid_unique = set(valid_PWC[valid_PWC.label==True].title.unique())
duplicate = []
for paper in set_train_unique:
if paper in set_valid_unique:
duplicate.append(paper)
len(duplicate)
len(set_train_unique)
len(set_valid_unique)
duplicate = []
for paper in set_valid_unique:
if paper in set_train_unique:
duplicate.append(paper)
len(duplicate)
```
| github_jupyter |
```
# Copyright 2021 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Vertex SDK: Custom training image classification model for batch prediction with explainabilty
<table align="left">
<td>
<a href="https://colab.research.google.com/github/GoogleCloudPlatform/vertex-ai-samples/tree/master/notebooks/official/automl/sdk_custom_image_classification_batch_explain.ipynb">
<img src="https://cloud.google.com/ml-engine/images/colab-logo-32px.png" alt="Colab logo"> Run in Colab
</a>
</td>
<td>
<a href="https://github.com/GoogleCloudPlatform/vertex-ai-samples/tree/master/notebooks/official/automl/sdk_custom_image_classification_batch_explain.ipynb">
<img src="https://cloud.google.com/ml-engine/images/github-logo-32px.png" alt="GitHub logo">
View on GitHub
</a>
</td>
<td>
<a href="https://console.cloud.google.com/ai/platform/notebooks/deploy-notebook?download_url=https://github.com/GoogleCloudPlatform/vertex-ai-samples/tree/master/notebooks/official/automl/sdk_custom_image_classification_batch_explain.ipynb">
Open in Google Cloud Notebooks
</a>
</td>
</table>
<br/><br/><br/>
## Overview
This tutorial demonstrates how to use the Vertex SDK to train and deploy a custom image classification model for batch prediction with explanation.
### Dataset
The dataset used for this tutorial is the [CIFAR10 dataset](https://www.tensorflow.org/datasets/catalog/cifar10) from [TensorFlow Datasets](https://www.tensorflow.org/datasets/catalog/overview). The version of the dataset you will use is built into TensorFlow. The trained model predicts which type of class an image is from ten classes: airplane, automobile, bird, cat, deer, dog, frog, horse, ship, truck.
### Objective
In this tutorial, you create a custom model, with a training pipeline, from a Python script in a Google prebuilt Docker container using the Vertex SDK, and then do a batch prediction with explanations on the uploaded model. You can alternatively create custom models using `gcloud` command-line tool or online using Cloud Console.
The steps performed include:
- Create a Vertex custom job for training a model.
- Train the TensorFlow model.
- Retrieve and load the model artifacts.
- View the model evaluation.
- Set explanation parameters.
- Upload the model as a Vertex `Model` resource.
- Make a batch prediction with explanations.
### Costs
This tutorial uses billable components of Google Cloud:
* Vertex AI
* Cloud Storage
Learn about [Vertex AI
pricing](https://cloud.google.com/vertex-ai/pricing) and [Cloud Storage
pricing](https://cloud.google.com/storage/pricing), and use the [Pricing
Calculator](https://cloud.google.com/products/calculator/)
to generate a cost estimate based on your projected usage.
### Set up your local development environment
If you are using Colab or Google Cloud Notebooks, your environment already meets all the requirements to run this notebook. You can skip this step.
Otherwise, make sure your environment meets this notebook's requirements. You need the following:
- The Cloud Storage SDK
- Git
- Python 3
- virtualenv
- Jupyter notebook running in a virtual environment with Python 3
The Cloud Storage guide to [Setting up a Python development environment](https://cloud.google.com/python/setup) and the [Jupyter installation guide](https://jupyter.org/install) provide detailed instructions for meeting these requirements. The following steps provide a condensed set of instructions:
1. [Install and initialize the SDK](https://cloud.google.com/sdk/docs/).
2. [Install Python 3](https://cloud.google.com/python/setup#installing_python).
3. [Install virtualenv](https://cloud.google.com/python/setup#installing_and_using_virtualenv) and create a virtual environment that uses Python 3. Activate the virtual environment.
4. To install Jupyter, run `pip3 install jupyter` on the command-line in a terminal shell.
5. To launch Jupyter, run `jupyter notebook` on the command-line in a terminal shell.
6. Open this notebook in the Jupyter Notebook Dashboard.
## Installation
Install the latest version of Vertex SDK for Python.
```
import os
# Google Cloud Notebook
if os.path.exists("/opt/deeplearning/metadata/env_version"):
USER_FLAG = "--user"
else:
USER_FLAG = ""
! pip3 install --upgrade google-cloud-aiplatform $USER_FLAG
```
Install the latest GA version of *google-cloud-storage* library as well.
```
! pip3 install -U google-cloud-storage $USER_FLAG
if os.environ["IS_TESTING"]:
! pip3 install --upgrade tensorflow $USER_FLAG
if os.environ["IS_TESTING"]:
! apt-get update && apt-get install -y python3-opencv-headless
! apt-get install -y libgl1-mesa-dev
! pip3 install --upgrade opencv-python-headless $USER_FLAG
```
### Restart the kernel
Once you've installed the additional packages, you need to restart the notebook kernel so it can find the packages.
```
import os
if not os.getenv("IS_TESTING"):
# Automatically restart kernel after installs
import IPython
app = IPython.Application.instance()
app.kernel.do_shutdown(True)
```
## Before you begin
### GPU runtime
This tutorial does not require a GPU runtime.
### Set up your Google Cloud project
**The following steps are required, regardless of your notebook environment.**
1. [Select or create a Google Cloud project](https://console.cloud.google.com/cloud-resource-manager). When you first create an account, you get a $300 free credit towards your compute/storage costs.
2. [Make sure that billing is enabled for your project.](https://cloud.google.com/billing/docs/how-to/modify-project)
3. [Enable the following APIs: Vertex AI APIs, Compute Engine APIs, and Cloud Storage.](https://console.cloud.google.com/flows/enableapi?apiid=ml.googleapis.com,compute_component,storage-component.googleapis.com)
4. If you are running this notebook locally, you will need to install the [Cloud SDK]((https://cloud.google.com/sdk)).
5. Enter your project ID in the cell below. Then run the cell to make sure the
Cloud SDK uses the right project for all the commands in this notebook.
**Note**: Jupyter runs lines prefixed with `!` as shell commands, and it interpolates Python variables prefixed with `$`.
```
PROJECT_ID = "[your-project-id]" # @param {type:"string"}
if PROJECT_ID == "" or PROJECT_ID is None or PROJECT_ID == "[your-project-id]":
# Get your GCP project id from gcloud
shell_output = ! gcloud config list --format 'value(core.project)' 2>/dev/null
PROJECT_ID = shell_output[0]
print("Project ID:", PROJECT_ID)
! gcloud config set project $PROJECT_ID
```
#### Region
You can also change the `REGION` variable, which is used for operations
throughout the rest of this notebook. Below are regions supported for Vertex AI. We recommend that you choose the region closest to you.
- Americas: `us-central1`
- Europe: `europe-west4`
- Asia Pacific: `asia-east1`
You may not use a multi-regional bucket for training with Vertex AI. Not all regions provide support for all Vertex AI services.
Learn more about [Vertex AI regions](https://cloud.google.com/vertex-ai/docs/general/locations)
```
REGION = "us-central1" # @param {type: "string"}
```
#### Timestamp
If you are in a live tutorial session, you might be using a shared test account or project. To avoid name collisions between users on resources created, you create a timestamp for each instance session, and append the timestamp onto the name of resources you create in this tutorial.
```
from datetime import datetime
TIMESTAMP = datetime.now().strftime("%Y%m%d%H%M%S")
```
### Authenticate your Google Cloud account
**If you are using Google Cloud Notebooks**, your environment is already authenticated. Skip this step.
**If you are using Colab**, run the cell below and follow the instructions when prompted to authenticate your account via oAuth.
**Otherwise**, follow these steps:
In the Cloud Console, go to the [Create service account key](https://console.cloud.google.com/apis/credentials/serviceaccountkey) page.
**Click Create service account**.
In the **Service account name** field, enter a name, and click **Create**.
In the **Grant this service account access to project** section, click the Role drop-down list. Type "Vertex" into the filter box, and select **Vertex Administrator**. Type "Storage Object Admin" into the filter box, and select **Storage Object Admin**.
Click Create. A JSON file that contains your key downloads to your local environment.
Enter the path to your service account key as the GOOGLE_APPLICATION_CREDENTIALS variable in the cell below and run the cell.
```
# If you are running this notebook in Colab, run this cell and follow the
# instructions to authenticate your GCP account. This provides access to your
# Cloud Storage bucket and lets you submit training jobs and prediction
# requests.
import os
import sys
# If on Google Cloud Notebook, then don't execute this code
if not os.path.exists("/opt/deeplearning/metadata/env_version"):
if "google.colab" in sys.modules:
from google.colab import auth as google_auth
google_auth.authenticate_user()
# If you are running this notebook locally, replace the string below with the
# path to your service account key and run this cell to authenticate your GCP
# account.
elif not os.getenv("IS_TESTING"):
%env GOOGLE_APPLICATION_CREDENTIALS ''
```
### Create a Cloud Storage bucket
**The following steps are required, regardless of your notebook environment.**
When you initialize the Vertex SDK for Python, you specify a Cloud Storage staging bucket. The staging bucket is where all the data associated with your dataset and model resources are retained across sessions.
Set the name of your Cloud Storage bucket below. Bucket names must be globally unique across all Google Cloud projects, including those outside of your organization.
```
BUCKET_NAME = "gs://[your-bucket-name]" # @param {type:"string"}
if BUCKET_NAME == "" or BUCKET_NAME is None or BUCKET_NAME == "gs://[your-bucket-name]":
BUCKET_NAME = "gs://" + PROJECT_ID + "aip-" + TIMESTAMP
```
**Only if your bucket doesn't already exist**: Run the following cell to create your Cloud Storage bucket.
```
! gsutil mb -l $REGION $BUCKET_NAME
```
Finally, validate access to your Cloud Storage bucket by examining its contents:
```
! gsutil ls -al $BUCKET_NAME
```
### Set up variables
Next, set up some variables used throughout the tutorial.
### Import libraries and define constants
```
import google.cloud.aiplatform as aip
```
## Initialize Vertex SDK for Python
Initialize the Vertex SDK for Python for your project and corresponding bucket.
```
aip.init(project=PROJECT_ID, staging_bucket=BUCKET_NAME)
```
#### Set hardware accelerators
You can set hardware accelerators for training and prediction.
Set the variables `TRAIN_GPU/TRAIN_NGPU` and `DEPLOY_GPU/DEPLOY_NGPU` to use a container image supporting a GPU and the number of GPUs allocated to the virtual machine (VM) instance. For example, to use a GPU container image with 4 Nvidia Telsa K80 GPUs allocated to each VM, you would specify:
(aip.AcceleratorType.NVIDIA_TESLA_K80, 4)
Otherwise specify `(None, None)` to use a container image to run on a CPU.
Learn more [here](https://cloud.google.com/vertex-ai/docs/general/locations#accelerators) hardware accelerator support for your region
*Note*: TF releases before 2.3 for GPU support will fail to load the custom model in this tutorial. It is a known issue and fixed in TF 2.3 -- which is caused by static graph ops that are generated in the serving function. If you encounter this issue on your own custom models, use a container image for TF 2.3 with GPU support.
```
if os.getenv("IS_TESTING_TRAIN_GPU"):
TRAIN_GPU, TRAIN_NGPU = (
aip.gapic.AcceleratorType.NVIDIA_TESLA_K80,
int(os.getenv("IS_TESTING_TRAIN_GPU")),
)
else:
TRAIN_GPU, TRAIN_NGPU = (None, None)
if os.getenv("IS_TESTING_DEPLOY_GPU"):
DEPLOY_GPU, DEPLOY_NGPU = (
aip.gapic.AcceleratorType.NVIDIA_TESLA_K80,
int(os.getenv("IS_TESTING_DEPLOY_GPU")),
)
else:
DEPLOY_GPU, DEPLOY_NGPU = (None, None)
```
#### Set pre-built containers
Set the pre-built Docker container image for training and prediction.
For the latest list, see [Pre-built containers for training](https://cloud.google.com/ai-platform-unified/docs/training/pre-built-containers).
For the latest list, see [Pre-built containers for prediction](https://cloud.google.com/ai-platform-unified/docs/predictions/pre-built-containers).
```
if os.getenv("IS_TESTING_TF"):
TF = os.getenv("IS_TESTING_TF")
else:
TF = "2-1"
if TF[0] == "2":
if TRAIN_GPU:
TRAIN_VERSION = "tf-gpu.{}".format(TF)
else:
TRAIN_VERSION = "tf-cpu.{}".format(TF)
if DEPLOY_GPU:
DEPLOY_VERSION = "tf2-gpu.{}".format(TF)
else:
DEPLOY_VERSION = "tf2-cpu.{}".format(TF)
else:
if TRAIN_GPU:
TRAIN_VERSION = "tf-gpu.{}".format(TF)
else:
TRAIN_VERSION = "tf-cpu.{}".format(TF)
if DEPLOY_GPU:
DEPLOY_VERSION = "tf-gpu.{}".format(TF)
else:
DEPLOY_VERSION = "tf-cpu.{}".format(TF)
TRAIN_IMAGE = "gcr.io/cloud-aiplatform/training/{}:latest".format(TRAIN_VERSION)
DEPLOY_IMAGE = "gcr.io/cloud-aiplatform/prediction/{}:latest".format(DEPLOY_VERSION)
print("Training:", TRAIN_IMAGE, TRAIN_GPU, TRAIN_NGPU)
print("Deployment:", DEPLOY_IMAGE, DEPLOY_GPU, DEPLOY_NGPU)
```
#### Set machine type
Next, set the machine type to use for training and prediction.
- Set the variables `TRAIN_COMPUTE` and `DEPLOY_COMPUTE` to configure the compute resources for the VMs you will use for for training and prediction.
- `machine type`
- `n1-standard`: 3.75GB of memory per vCPU.
- `n1-highmem`: 6.5GB of memory per vCPU
- `n1-highcpu`: 0.9 GB of memory per vCPU
- `vCPUs`: number of \[2, 4, 8, 16, 32, 64, 96 \]
*Note: The following is not supported for training:*
- `standard`: 2 vCPUs
- `highcpu`: 2, 4 and 8 vCPUs
*Note: You may also use n2 and e2 machine types for training and deployment, but they do not support GPUs*.
```
if os.getenv("IS_TESTING_TRAIN_MACHINE"):
MACHINE_TYPE = os.getenv("IS_TESTING_TRAIN_MACHINE")
else:
MACHINE_TYPE = "n1-standard"
VCPU = "4"
TRAIN_COMPUTE = MACHINE_TYPE + "-" + VCPU
print("Train machine type", TRAIN_COMPUTE)
if os.getenv("IS_TESTING_DEPLOY_MACHINE"):
MACHINE_TYPE = os.getenv("IS_TESTING_DEPLOY_MACHINE")
else:
MACHINE_TYPE = "n1-standard"
VCPU = "4"
DEPLOY_COMPUTE = MACHINE_TYPE + "-" + VCPU
print("Deploy machine type", DEPLOY_COMPUTE)
```
# Tutorial
Now you are ready to start creating your own custom model and training for CIFAR10.
### Examine the training package
#### Package layout
Before you start the training, you will look at how a Python package is assembled for a custom training job. When unarchived, the package contains the following directory/file layout.
- PKG-INFO
- README.md
- setup.cfg
- setup.py
- trainer
- \_\_init\_\_.py
- task.py
The files `setup.cfg` and `setup.py` are the instructions for installing the package into the operating environment of the Docker image.
The file `trainer/task.py` is the Python script for executing the custom training job. *Note*, when we referred to it in the worker pool specification, we replace the directory slash with a dot (`trainer.task`) and dropped the file suffix (`.py`).
#### Package Assembly
In the following cells, you will assemble the training package.
```
# Make folder for Python training script
! rm -rf custom
! mkdir custom
# Add package information
! touch custom/README.md
setup_cfg = "[egg_info]\n\ntag_build =\n\ntag_date = 0"
! echo "$setup_cfg" > custom/setup.cfg
setup_py = "import setuptools\n\nsetuptools.setup(\n\n install_requires=[\n\n 'tensorflow_datasets==1.3.0',\n\n ],\n\n packages=setuptools.find_packages())"
! echo "$setup_py" > custom/setup.py
pkg_info = "Metadata-Version: 1.0\n\nName: CIFAR10 image classification\n\nVersion: 0.0.0\n\nSummary: Demostration training script\n\nHome-page: www.google.com\n\nAuthor: Google\n\nAuthor-email: aferlitsch@google.com\n\nLicense: Public\n\nDescription: Demo\n\nPlatform: Vertex"
! echo "$pkg_info" > custom/PKG-INFO
# Make the training subfolder
! mkdir custom/trainer
! touch custom/trainer/__init__.py
```
#### Task.py contents
In the next cell, you write the contents of the training script task.py. We won't go into detail, it's just there for you to browse. In summary:
- Get the directory where to save the model artifacts from the command line (`--model_dir`), and if not specified, then from the environment variable `AIP_MODEL_DIR`.
- Loads CIFAR10 dataset from TF Datasets (tfds).
- Builds a model using TF.Keras model API.
- Compiles the model (`compile()`).
- Sets a training distribution strategy according to the argument `args.distribute`.
- Trains the model (`fit()`) with epochs and steps according to the arguments `args.epochs` and `args.steps`
- Saves the trained model (`save(args.model_dir)`) to the specified model directory.
```
%%writefile custom/trainer/task.py
# Single, Mirror and Multi-Machine Distributed Training for CIFAR-10
import tensorflow_datasets as tfds
import tensorflow as tf
from tensorflow.python.client import device_lib
import argparse
import os
import sys
tfds.disable_progress_bar()
parser = argparse.ArgumentParser()
parser.add_argument('--model-dir', dest='model_dir',
default=os.getenv("AIP_MODEL_DIR"), type=str, help='Model dir.')
parser.add_argument('--lr', dest='lr',
default=0.01, type=float,
help='Learning rate.')
parser.add_argument('--epochs', dest='epochs',
default=10, type=int,
help='Number of epochs.')
parser.add_argument('--steps', dest='steps',
default=200, type=int,
help='Number of steps per epoch.')
parser.add_argument('--distribute', dest='distribute', type=str, default='single',
help='distributed training strategy')
args = parser.parse_args()
print('Python Version = {}'.format(sys.version))
print('TensorFlow Version = {}'.format(tf.__version__))
print('TF_CONFIG = {}'.format(os.environ.get('TF_CONFIG', 'Not found')))
print('DEVICES', device_lib.list_local_devices())
# Single Machine, single compute device
if args.distribute == 'single':
if tf.test.is_gpu_available():
strategy = tf.distribute.OneDeviceStrategy(device="/gpu:0")
else:
strategy = tf.distribute.OneDeviceStrategy(device="/cpu:0")
# Single Machine, multiple compute device
elif args.distribute == 'mirror':
strategy = tf.distribute.MirroredStrategy()
# Multiple Machine, multiple compute device
elif args.distribute == 'multi':
strategy = tf.distribute.experimental.MultiWorkerMirroredStrategy()
# Multi-worker configuration
print('num_replicas_in_sync = {}'.format(strategy.num_replicas_in_sync))
# Preparing dataset
BUFFER_SIZE = 10000
BATCH_SIZE = 64
def make_datasets_unbatched():
# Scaling CIFAR10 data from (0, 255] to (0., 1.]
def scale(image, label):
image = tf.cast(image, tf.float32)
image /= 255.0
return image, label
datasets, info = tfds.load(name='cifar10',
with_info=True,
as_supervised=True)
return datasets['train'].map(scale).cache().shuffle(BUFFER_SIZE).repeat()
# Build the Keras model
def build_and_compile_cnn_model():
model = tf.keras.Sequential([
tf.keras.layers.Conv2D(32, 3, activation='relu', input_shape=(32, 32, 3)),
tf.keras.layers.MaxPooling2D(),
tf.keras.layers.Conv2D(32, 3, activation='relu'),
tf.keras.layers.MaxPooling2D(),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(10, activation='softmax')
])
model.compile(
loss=tf.keras.losses.sparse_categorical_crossentropy,
optimizer=tf.keras.optimizers.SGD(learning_rate=args.lr),
metrics=['accuracy'])
return model
# Train the model
NUM_WORKERS = strategy.num_replicas_in_sync
# Here the batch size scales up by number of workers since
# `tf.data.Dataset.batch` expects the global batch size.
GLOBAL_BATCH_SIZE = BATCH_SIZE * NUM_WORKERS
train_dataset = make_datasets_unbatched().batch(GLOBAL_BATCH_SIZE)
with strategy.scope():
# Creation of dataset, and model building/compiling need to be within
# `strategy.scope()`.
model = build_and_compile_cnn_model()
model.fit(x=train_dataset, epochs=args.epochs, steps_per_epoch=args.steps)
model.save(args.model_dir)
```
#### Store training script on your Cloud Storage bucket
Next, you package the training folder into a compressed tar ball, and then store it in your Cloud Storage bucket.
```
! rm -f custom.tar custom.tar.gz
! tar cvf custom.tar custom
! gzip custom.tar
! gsutil cp custom.tar.gz $BUCKET_NAME/trainer_cifar10.tar.gz
```
### Create and run custom training job
To train a custom model, you perform two steps: 1) create a custom training job, and 2) run the job.
#### Create custom training job
A custom training job is created with the `CustomTrainingJob` class, with the following parameters:
- `display_name`: The human readable name for the custom training job.
- `container_uri`: The training container image.
- `requirements`: Package requirements for the training container image (e.g., pandas).
- `script_path`: The relative path to the training script.
```
job = aip.CustomTrainingJob(
display_name="cifar10_" + TIMESTAMP,
script_path="custom/trainer/task.py",
container_uri=TRAIN_IMAGE,
requirements=["gcsfs==0.7.1", "tensorflow-datasets==4.4"],
)
print(job)
```
### Prepare your command-line arguments
Now define the command-line arguments for your custom training container:
- `args`: The command-line arguments to pass to the executable that is set as the entry point into the container.
- `--model-dir` : For our demonstrations, we use this command-line argument to specify where to store the model artifacts.
- direct: You pass the Cloud Storage location as a command line argument to your training script (set variable `DIRECT = True`), or
- indirect: The service passes the Cloud Storage location as the environment variable `AIP_MODEL_DIR` to your training script (set variable `DIRECT = False`). In this case, you tell the service the model artifact location in the job specification.
- `"--epochs=" + EPOCHS`: The number of epochs for training.
- `"--steps=" + STEPS`: The number of steps per epoch.
```
MODEL_DIR = "{}/{}".format(BUCKET_NAME, TIMESTAMP)
EPOCHS = 20
STEPS = 100
DIRECT = True
if DIRECT:
CMDARGS = [
"--model-dir=" + MODEL_DIR,
"--epochs=" + str(EPOCHS),
"--steps=" + str(STEPS),
]
else:
CMDARGS = [
"--epochs=" + str(EPOCHS),
"--steps=" + str(STEPS),
]
```
#### Run the custom training job
Next, you run the custom job to start the training job by invoking the method `run`, with the following parameters:
- `args`: The command-line arguments to pass to the training script.
- `replica_count`: The number of compute instances for training (replica_count = 1 is single node training).
- `machine_type`: The machine type for the compute instances.
- `accelerator_type`: The hardware accelerator type.
- `accelerator_count`: The number of accelerators to attach to a worker replica.
- `base_output_dir`: The Cloud Storage location to write the model artifacts to.
- `sync`: Whether to block until completion of the job.
```
if TRAIN_GPU:
job.run(
args=CMDARGS,
replica_count=1,
machine_type=TRAIN_COMPUTE,
accelerator_type=TRAIN_GPU.name,
accelerator_count=TRAIN_NGPU,
base_output_dir=MODEL_DIR,
sync=True,
)
else:
job.run(
args=CMDARGS,
replica_count=1,
machine_type=TRAIN_COMPUTE,
base_output_dir=MODEL_DIR,
sync=True,
)
model_path_to_deploy = MODEL_DIR
```
## Load the saved model
Your model is stored in a TensorFlow SavedModel format in a Cloud Storage bucket. Now load it from the Cloud Storage bucket, and then you can do some things, like evaluate the model, and do a prediction.
To load, you use the TF.Keras `model.load_model()` method passing it the Cloud Storage path where the model is saved -- specified by `MODEL_DIR`.
```
import tensorflow as tf
local_model = tf.keras.models.load_model(MODEL_DIR)
```
## Evaluate the model
Now find out how good the model is.
### Load evaluation data
You will load the CIFAR10 test (holdout) data from `tf.keras.datasets`, using the method `load_data()`. This returns the dataset as a tuple of two elements. The first element is the training data and the second is the test data. Each element is also a tuple of two elements: the image data, and the corresponding labels.
You don't need the training data, and hence why we loaded it as `(_, _)`.
Before you can run the data through evaluation, you need to preprocess it:
`x_test`:
1. Normalize (rescale) the pixel data by dividing each pixel by 255. This replaces each single byte integer pixel with a 32-bit floating point number between 0 and 1.
`y_test`:<br/>
2. The labels are currently scalar (sparse). If you look back at the `compile()` step in the `trainer/task.py` script, you will find that it was compiled for sparse labels. So we don't need to do anything more.
```
import numpy as np
from tensorflow.keras.datasets import cifar10
(_, _), (x_test, y_test) = cifar10.load_data()
x_test = (x_test / 255.0).astype(np.float32)
print(x_test.shape, y_test.shape)
```
### Perform the model evaluation
Now evaluate how well the model in the custom job did.
```
local_model.evaluate(x_test, y_test)
```
### Serving function for image data
To pass images to the prediction service, you encode the compressed (e.g., JPEG) image bytes into base 64 -- which makes the content safe from modification while transmitting binary data over the network. Since this deployed model expects input data as raw (uncompressed) bytes, you need to ensure that the base 64 encoded data gets converted back to raw bytes before it is passed as input to the deployed model.
To resolve this, define a serving function (`serving_fn`) and attach it to the model as a preprocessing step. Add a `@tf.function` decorator so the serving function is fused to the underlying model (instead of upstream on a CPU).
When you send a prediction or explanation request, the content of the request is base 64 decoded into a Tensorflow string (`tf.string`), which is passed to the serving function (`serving_fn`). The serving function preprocesses the `tf.string` into raw (uncompressed) numpy bytes (`preprocess_fn`) to match the input requirements of the model:
- `io.decode_jpeg`- Decompresses the JPG image which is returned as a Tensorflow tensor with three channels (RGB).
- `image.convert_image_dtype` - Changes integer pixel values to float 32.
- `image.resize` - Resizes the image to match the input shape for the model.
- `resized / 255.0` - Rescales (normalization) the pixel data between 0 and 1.
At this point, the data can be passed to the model (`m_call`).
#### XAI Signatures
When the serving function is saved back with the underlying model (`tf.saved_model.save`), you specify the input layer of the serving function as the signature `serving_default`.
For XAI image models, you need to save two additional signatures from the serving function:
- `xai_preprocess`: The preprocessing function in the serving function.
- `xai_model`: The concrete function for calling the model.
```
CONCRETE_INPUT = "numpy_inputs"
def _preprocess(bytes_input):
decoded = tf.io.decode_jpeg(bytes_input, channels=3)
decoded = tf.image.convert_image_dtype(decoded, tf.float32)
resized = tf.image.resize(decoded, size=(32, 32))
rescale = tf.cast(resized / 255.0, tf.float32)
return rescale
@tf.function(input_signature=[tf.TensorSpec([None], tf.string)])
def preprocess_fn(bytes_inputs):
decoded_images = tf.map_fn(
_preprocess, bytes_inputs, dtype=tf.float32, back_prop=False
)
return {
CONCRETE_INPUT: decoded_images
} # User needs to make sure the key matches model's input
@tf.function(input_signature=[tf.TensorSpec([None], tf.string)])
def serving_fn(bytes_inputs):
images = preprocess_fn(bytes_inputs)
prob = m_call(**images)
return prob
m_call = tf.function(local_model.call).get_concrete_function(
[tf.TensorSpec(shape=[None, 32, 32, 3], dtype=tf.float32, name=CONCRETE_INPUT)]
)
tf.saved_model.save(
local_model,
model_path_to_deploy,
signatures={
"serving_default": serving_fn,
# Required for XAI
"xai_preprocess": preprocess_fn,
"xai_model": m_call,
},
)
```
## Get the serving function signature
You can get the signatures of your model's input and output layers by reloading the model into memory, and querying it for the signatures corresponding to each layer.
When making a prediction request, you need to route the request to the serving function instead of the model, so you need to know the input layer name of the serving function -- which you will use later when you make a prediction request.
You also need to know the name of the serving function's input and output layer for constructing the explanation metadata -- which is discussed subsequently.
```
loaded = tf.saved_model.load(model_path_to_deploy)
serving_input = list(
loaded.signatures["serving_default"].structured_input_signature[1].keys()
)[0]
print("Serving function input:", serving_input)
serving_output = list(loaded.signatures["serving_default"].structured_outputs.keys())[0]
print("Serving function output:", serving_output)
input_name = local_model.input.name
print("Model input name:", input_name)
output_name = local_model.output.name
print("Model output name:", output_name)
```
### Explanation Specification
To get explanations when doing a prediction, you must enable the explanation capability and set corresponding settings when you upload your custom model to an Vertex `Model` resource. These settings are referred to as the explanation metadata, which consists of:
- `parameters`: This is the specification for the explainability algorithm to use for explanations on your model. You can choose between:
- Shapley - *Note*, not recommended for image data -- can be very long running
- XRAI
- Integrated Gradients
- `metadata`: This is the specification for how the algoithm is applied on your custom model.
#### Explanation Parameters
Let's first dive deeper into the settings for the explainability algorithm.
#### Shapley
Assigns credit for the outcome to each feature, and considers different permutations of the features. This method provides a sampling approximation of exact Shapley values.
Use Cases:
- Classification and regression on tabular data.
Parameters:
- `path_count`: This is the number of paths over the features that will be processed by the algorithm. An exact approximation of the Shapley values requires M! paths, where M is the number of features. For the CIFAR10 dataset, this would be 784 (28*28).
For any non-trival number of features, this is too compute expensive. You can reduce the number of paths over the features to M * `path_count`.
#### Integrated Gradients
A gradients-based method to efficiently compute feature attributions with the same axiomatic properties as the Shapley value.
Use Cases:
- Classification and regression on tabular data.
- Classification on image data.
Parameters:
- `step_count`: This is the number of steps to approximate the remaining sum. The more steps, the more accurate the integral approximation. The general rule of thumb is 50 steps, but as you increase so does the compute time.
#### XRAI
Based on the integrated gradients method, XRAI assesses overlapping regions of the image to create a saliency map, which highlights relevant regions of the image rather than pixels.
Use Cases:
- Classification on image data.
Parameters:
- `step_count`: This is the number of steps to approximate the remaining sum. The more steps, the more accurate the integral approximation. The general rule of thumb is 50 steps, but as you increase so does the compute time.
In the next code cell, set the variable `XAI` to which explainabilty algorithm you will use on your custom model.
```
XAI = "ig" # [ shapley, ig, xrai ]
if XAI == "shapley":
PARAMETERS = {"sampled_shapley_attribution": {"path_count": 10}}
elif XAI == "ig":
PARAMETERS = {"integrated_gradients_attribution": {"step_count": 50}}
elif XAI == "xrai":
PARAMETERS = {"xrai_attribution": {"step_count": 50}}
parameters = aip.explain.ExplanationParameters(PARAMETERS)
```
#### Explanation Metadata
Let's first dive deeper into the explanation metadata, which consists of:
- `outputs`: A scalar value in the output to attribute -- what to explain. For example, in a probability output \[0.1, 0.2, 0.7\] for classification, one wants an explanation for 0.7. Consider the following formulae, where the output is `y` and that is what we want to explain.
y = f(x)
Consider the following formulae, where the outputs are `y` and `z`. Since we can only do attribution for one scalar value, we have to pick whether we want to explain the output `y` or `z`. Assume in this example the model is object detection and y and z are the bounding box and the object classification. You would want to pick which of the two outputs to explain.
y, z = f(x)
The dictionary format for `outputs` is:
{ "outputs": { "[your_display_name]":
"output_tensor_name": [layer]
}
}
<blockquote>
- [your_display_name]: A human readable name you assign to the output to explain. A common example is "probability".<br/>
- "output_tensor_name": The key/value field to identify the output layer to explain. <br/>
- [layer]: The output layer to explain. In a single task model, like a tabular regressor, it is the last (topmost) layer in the model.
</blockquote>
- `inputs`: The features for attribution -- how they contributed to the output. Consider the following formulae, where `a` and `b` are the features. We have to pick which features to explain how the contributed. Assume that this model is deployed for A/B testing, where `a` are the data_items for the prediction and `b` identifies whether the model instance is A or B. You would want to pick `a` (or some subset of) for the features, and not `b` since it does not contribute to the prediction.
y = f(a,b)
The minimum dictionary format for `inputs` is:
{ "inputs": { "[your_display_name]":
"input_tensor_name": [layer]
}
}
<blockquote>
- [your_display_name]: A human readable name you assign to the input to explain. A common example is "features".<br/>
- "input_tensor_name": The key/value field to identify the input layer for the feature attribution. <br/>
- [layer]: The input layer for feature attribution. In a single input tensor model, it is the first (bottom-most) layer in the model.
</blockquote>
Since the inputs to the model are images, you can specify the following additional fields as reporting/visualization aids:
<blockquote>
- "modality": "image": Indicates the field values are image data.
</blockquote>
```
random_baseline = np.random.rand(32, 32, 3)
input_baselines = [{"number_vaue": x} for x in random_baseline]
INPUT_METADATA = {"input_tensor_name": CONCRETE_INPUT, "modality": "image"}
OUTPUT_METADATA = {"output_tensor_name": serving_output}
input_metadata = aip.explain.ExplanationMetadata.InputMetadata(INPUT_METADATA)
output_metadata = aip.explain.ExplanationMetadata.OutputMetadata(OUTPUT_METADATA)
metadata = aip.explain.ExplanationMetadata(
inputs={"image": input_metadata}, outputs={"class": output_metadata}
)
```
## Upload the model
Next, upload your model to a `Model` resource using `Model.upload()` method, with the following parameters:
- `display_name`: The human readable name for the `Model` resource.
- `artifact`: The Cloud Storage location of the trained model artifacts.
- `serving_container_image_uri`: The serving container image.
- `sync`: Whether to execute the upload asynchronously or synchronously.
- `explanation_parameters`: Parameters to configure explaining for `Model`'s predictions.
- `explanation_metadata`: Metadata describing the `Model`'s input and output for explanation.
If the `upload()` method is run asynchronously, you can subsequently block until completion with the `wait()` method.
```
model = aip.Model.upload(
display_name="cifar10_" + TIMESTAMP,
artifact_uri=MODEL_DIR,
serving_container_image_uri=DEPLOY_IMAGE,
explanation_parameters=parameters,
explanation_metadata=metadata,
sync=False,
)
model.wait()
```
### Get test items
You will use examples out of the test (holdout) portion of the dataset as a test items.
```
test_image_1 = x_test[0]
test_label_1 = y_test[0]
test_image_2 = x_test[1]
test_label_2 = y_test[1]
print(test_image_1.shape)
```
### Prepare the request content
You are going to send the CIFAR10 images as compressed JPG image, instead of the raw uncompressed bytes:
- `cv2.imwrite`: Use openCV to write the uncompressed image to disk as a compressed JPEG image.
- Denormalize the image data from \[0,1) range back to [0,255).
- Convert the 32-bit floating point values to 8-bit unsigned integers.
```
import cv2
cv2.imwrite("tmp1.jpg", (test_image_1 * 255).astype(np.uint8))
cv2.imwrite("tmp2.jpg", (test_image_2 * 255).astype(np.uint8))
```
### Copy test item(s)
For the batch prediction, copy the test items over to your Cloud Storage bucket.
```
! gsutil cp tmp1.jpg $BUCKET_NAME/tmp1.jpg
! gsutil cp tmp2.jpg $BUCKET_NAME/tmp2.jpg
test_item_1 = BUCKET_NAME + "/tmp1.jpg"
test_item_2 = BUCKET_NAME + "/tmp2.jpg"
```
### Make the batch input file
Now make a batch input file, which you will store in your local Cloud Storage bucket. The batch input file can only be in JSONL format. For JSONL file, you make one dictionary entry per line for each data item (instance). The dictionary contains the key/value pairs:
- `input_name`: the name of the input layer of the underlying model.
- `'b64'`: A key that indicates the content is base64 encoded.
- `content`: The compressed JPG image bytes as a base64 encoded string.
Each instance in the prediction request is a dictionary entry of the form:
{serving_input: {'b64': content}}
To pass the image data to the prediction service you encode the bytes into base64 -- which makes the content safe from modification when transmitting binary data over the network.
- `tf.io.read_file`: Read the compressed JPG images into memory as raw bytes.
- `base64.b64encode`: Encode the raw bytes into a base64 encoded string.
```
import base64
import json
gcs_input_uri = BUCKET_NAME + "/" + "test.jsonl"
with tf.io.gfile.GFile(gcs_input_uri, "w") as f:
bytes = tf.io.read_file(test_item_1)
b64str = base64.b64encode(bytes.numpy()).decode("utf-8")
data = {serving_input: {"b64": b64str}}
f.write(json.dumps(data) + "\n")
bytes = tf.io.read_file(test_item_2)
b64str = base64.b64encode(bytes.numpy()).decode("utf-8")
data = {serving_input: {"b64": b64str}}
f.write(json.dumps(data) + "\n")
```
### Make the batch prediction request
Now that your Model resource is trained, you can make a batch prediction by invoking the batch_predict() method, with the following parameters:
- `job_display_name`: The human readable name for the batch prediction job.
- `gcs_source`: A list of one or more batch request input files.
- `gcs_destination_prefix`: The Cloud Storage location for storing the batch prediction resuls.
- `instances_format`: The format for the input instances, either 'csv' or 'jsonl'. Defaults to 'jsonl'.
- `predictions_format`: The format for the output predictions, either 'csv' or 'jsonl'. Defaults to 'jsonl'.
- `machine_type`: The type of machine to use for training.
- `sync`: If set to True, the call will block while waiting for the asynchronous batch job to complete.
```
MIN_NODES = 1
MAX_NODES = 1
batch_predict_job = model.batch_predict(
job_display_name="cifar10_" + TIMESTAMP,
gcs_source=gcs_input_uri,
gcs_destination_prefix=BUCKET_NAME,
instances_format="jsonl",
model_parameters=None,
machine_type=DEPLOY_COMPUTE,
starting_replica_count=MIN_NODES,
max_replica_count=MAX_NODES,
generate_explanation=True,
sync=False,
)
print(batch_predict_job)
```
### Wait for completion of batch prediction job
Next, wait for the batch job to complete. Alternatively, one can set the parameter `sync` to `True` in the `batch_predict()` method to block until the batch prediction job is completed.
```
batch_predict_job.wait()
```
### Get the explanations
Next, get the explanation results from the completed batch prediction job.
The results are written to the Cloud Storage output bucket you specified in the batch prediction request. You call the method iter_outputs() to get a list of each Cloud Storage file generated with the results. Each file contains one or more explanation requests in a CSV format:
- CSV header + predicted_label
- CSV row + explanation, per prediction request
```
import tensorflow as tf
bp_iter_outputs = batch_predict_job.iter_outputs()
explanation_results = list()
for blob in bp_iter_outputs:
if blob.name.split("/")[-1].startswith("explanation"):
explanation_results.append(blob.name)
tags = list()
for explanation_result in explanation_results:
gfile_name = f"gs://{bp_iter_outputs.bucket.name}/{explanation_result}"
with tf.io.gfile.GFile(name=gfile_name, mode="r") as gfile:
for line in gfile.readlines():
print(line)
```
# Cleaning up
To clean up all Google Cloud resources used in this project, you can [delete the Google Cloud
project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial.
Otherwise, you can delete the individual resources you created in this tutorial:
- Dataset
- Pipeline
- Model
- Endpoint
- AutoML Training Job
- Batch Job
- Custom Job
- Hyperparameter Tuning Job
- Cloud Storage Bucket
```
delete_all = True
if delete_all:
# Delete the dataset using the Vertex dataset object
try:
if "dataset" in globals():
dataset.delete()
except Exception as e:
print(e)
# Delete the model using the Vertex model object
try:
if "model" in globals():
model.delete()
except Exception as e:
print(e)
# Delete the endpoint using the Vertex endpoint object
try:
if "endpoint" in globals():
endpoint.delete()
except Exception as e:
print(e)
# Delete the AutoML or Pipeline trainig job
try:
if "dag" in globals():
dag.delete()
except Exception as e:
print(e)
# Delete the custom trainig job
try:
if "job" in globals():
job.delete()
except Exception as e:
print(e)
# Delete the batch prediction job using the Vertex batch prediction object
try:
if "batch_predict_job" in globals():
batch_predict_job.delete()
except Exception as e:
print(e)
# Delete the hyperparameter tuning job using the Vertex hyperparameter tuning object
try:
if "hpt_job" in globals():
hpt_job.delete()
except Exception as e:
print(e)
if "BUCKET_NAME" in globals():
! gsutil rm -r $BUCKET_NAME
```
| github_jupyter |
# indicators
```
import vectorbt as vbt
import numpy as np
import pandas as pd
from datetime import datetime, timedelta
from numba import njit
import itertools
import talib
import ta
# Disable caching for performance testing
vbt.settings.caching['enabled'] = False
close = pd.DataFrame({
'a': [1., 2., 3., 4., 5.],
'b': [5., 4., 3., 2., 1.],
'c': [1., 2., 3., 2., 1.]
}, index=pd.DatetimeIndex([
datetime(2018, 1, 1),
datetime(2018, 1, 2),
datetime(2018, 1, 3),
datetime(2018, 1, 4),
datetime(2018, 1, 5)
]))
np.random.seed(42)
high = close * np.random.uniform(1, 1.1, size=close.shape)
low = close * np.random.uniform(0.9, 1, size=close.shape)
volume = close * 0 + np.random.randint(1, 10, size=close.shape).astype(float)
big_close = pd.DataFrame(np.random.randint(10, size=(1000, 1000)).astype(float))
big_close.index = [datetime(2018, 1, 1) + timedelta(days=i) for i in range(1000)]
big_high = big_close * np.random.uniform(1, 1.1, size=big_close.shape)
big_low = big_close * np.random.uniform(0.9, 1, size=big_close.shape)
big_volume = big_close * 0 + np.random.randint(10, 100, size=big_close.shape).astype(float)
close_ts = pd.Series([1, 2, 3, 4, 3, 2, 1], index=pd.DatetimeIndex([
datetime(2018, 1, 1),
datetime(2018, 1, 2),
datetime(2018, 1, 3),
datetime(2018, 1, 4),
datetime(2018, 1, 5),
datetime(2018, 1, 6),
datetime(2018, 1, 7)
]))
high_ts = close_ts * 1.1
low_ts = close_ts * 0.9
volume_ts = pd.Series([4, 3, 2, 1, 2, 3, 4], index=close_ts.index)
```
## IndicatorFactory
```
def apply_func(i, ts, p, a, b=100):
return ts * p[i] + a + b
@njit
def apply_func_nb(i, ts, p, a, b):
return ts * p[i] + a + b # numba doesn't support **kwargs
# Custom function can be anything that takes time series, params and other arguments, and returns outputs
def custom_func(ts, p, *args, **kwargs):
return vbt.base.combine_fns.apply_and_concat_one(len(p), apply_func, ts, p, *args, **kwargs)
@njit
def custom_func_nb(ts, p, *args):
return vbt.base.combine_fns.apply_and_concat_one_nb(len(p), apply_func_nb, ts, p, *args)
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p'], output_names=['out'])
print(F.from_custom_func(custom_func, var_args=True)
.run(close, [0, 1], 10, b=100).out)
print(F.from_custom_func(custom_func_nb, var_args=True)
.run(close, [0, 1], 10, 100).out)
# Apply function is performed on each parameter individually, and each output is then stacked for you
# Apply functions are less customizable than custom functions, but are simpler to write
def apply_func(ts, p, a, b=100):
return ts * p + a + b
@njit
def apply_func_nb(ts, p, a, b):
return ts * p + a + b # numba doesn't support **kwargs
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p'], output_names=['out'])
print(F.from_apply_func(apply_func, var_args=True)
.run(close, [0, 1], 10, b=100).out)
print(F.from_apply_func(apply_func_nb, var_args=True)
.run(close, [0, 1], 10, 100).out)
# test *args
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p'], output_names=['out'])
print(F.from_apply_func(lambda ts, p, a: ts * p + a, var_args=True)
.run(close, [0, 1, 2], 3).out)
print(F.from_apply_func(njit(lambda ts, p, a: ts * p + a), var_args=True)
.run(close, [0, 1, 2], 3).out)
# test **kwargs
# Numba doesn't support kwargs out of the box
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p'], output_names=['out'])
print(F.from_apply_func(lambda ts, p, a=1: ts * p + a)
.run(close, [0, 1, 2], a=3).out)
# test no inputs
F = vbt.IndicatorFactory(param_names=['p'], output_names=['out'])
print(F.from_apply_func(lambda p: np.full((3, 3), p))
.run([0, 1]).out)
print(F.from_apply_func(njit(lambda p: np.full((3, 3), p)))
.run([0, 1]).out)
# test no inputs with input_shape, input_index and input_columns
F = vbt.IndicatorFactory(param_names=['p'], output_names=['out'])
print(F.from_apply_func(lambda input_shape, p: np.full(input_shape, p), require_input_shape=True)
.run((5,), 0).out)
print(F.from_apply_func(njit(lambda input_shape, p: np.full(input_shape, p)), require_input_shape=True)
.run((5,), 0).out)
print(F.from_apply_func(lambda input_shape, p: np.full(input_shape, p), require_input_shape=True)
.run((5,), [0, 1]).out)
print(F.from_apply_func(njit(lambda input_shape, p: np.full(input_shape, p)), require_input_shape=True)
.run((5,), [0, 1]).out)
print(F.from_apply_func(lambda input_shape, p: np.full(input_shape, p), require_input_shape=True)
.run((5, 3), [0, 1], input_index=close.index, input_columns=close.columns).out)
print(F.from_apply_func(njit(lambda input_shape, p: np.full(input_shape, p)), require_input_shape=True)
.run((5, 3), [0, 1], input_index=close.index, input_columns=close.columns).out)
# test multiple inputs
F = vbt.IndicatorFactory(input_names=['ts1', 'ts2'], param_names=['p'], output_names=['out'])
print(F.from_apply_func(lambda ts1, ts2, p: ts1 * ts2 * p)
.run(close, high, [0, 1]).out)
print(F.from_apply_func(njit(lambda ts1, ts2, p: ts1 * ts2 * p))
.run(close, high, [0, 1]).out)
# test no params
F = vbt.IndicatorFactory(input_names=['ts'], output_names=['out'])
print(F.from_apply_func(lambda ts: ts)
.run(close).out)
print(F.from_apply_func(njit(lambda ts: ts))
.run(close).out)
# test no inputs and no params
F = vbt.IndicatorFactory(output_names=['out'])
print(F.from_apply_func(lambda: np.full((3, 3), 1))
.run().out)
print(F.from_apply_func(njit(lambda: np.full((3, 3), 1)))
.run().out)
# test multiple params
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p1', 'p2'], output_names=['out'])
print(F.from_apply_func(lambda ts, p1, p2: ts * (p1 + p2))
.run(close, np.asarray([0, 1]), np.asarray([2, 3])).out)
print(F.from_apply_func(njit(lambda ts, p1, p2: ts * (p1 + p2)))
.run(close, np.asarray([0, 1]), np.asarray([2, 3])).out)
# test param_settings array_like
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p1', 'p2'], output_names=['out'])
print(F.from_apply_func(lambda ts, p1, p2: ts * (p1 + p2),
param_settings={'p1': {'is_array_like': True}})
.run(close, np.asarray([0, 1, 2]), np.asarray([2, 3])).out)
print(F.from_apply_func(njit(lambda ts, p1, p2: ts * (p1 + p2)),
param_settings={'p1': {'is_array_like': True}})
.run(close, np.asarray([0, 1, 2]), np.asarray([2, 3])).out)
# test param_settings bc_to_input
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p1', 'p2'], output_names=['out'])
print(F.from_apply_func(lambda ts, p1, p2: ts * (p1 + p2),
param_settings={'p1': {'is_array_like': True, 'bc_to_input': True}})
.run(close, np.asarray([0, 1, 2]), np.asarray([2, 3])).out)
print(F.from_apply_func(njit(lambda ts, p1, p2: ts * (p1 + p2)),
param_settings={'p1': {'is_array_like': True, 'bc_to_input': True}})
.run(close, np.asarray([0, 1, 2]), np.asarray([2, 3])).out)
# test param product
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p1', 'p2'], output_names=['out'])
print(F.from_apply_func(lambda ts, p1, p2: ts * (p1 + p2))
.run(close, [0, 1], [2, 3], param_product=True).out)
print(F.from_apply_func(njit(lambda ts, p1, p2: ts * (p1 + p2)))
.run(close, [0, 1], [2, 3], param_product=True).out)
# test default params
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p1', 'p2'], output_names=['out'])
print(F.from_apply_func(lambda ts, p1, p2: ts * (p1 + p2), p2=2)
.run(close, [0, 1]).out)
print(F.from_apply_func(njit(lambda ts, p1, p2: ts * (p1 + p2)), p2=2)
.run(close, [0, 1]).out)
# test hide_params
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p1', 'p2'], output_names=['out'])
print(F.from_apply_func(lambda ts, p1, p2: ts * (p1 + p2), hide_params=['p2'])
.run(close, [0, 1], 2).out)
print(F.from_apply_func(njit(lambda ts, p1, p2: ts * (p1 + p2)), hide_params=['p2'])
.run(close, [0, 1], 2).out)
# test hide_default
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p1', 'p2'], output_names=['out'])
print(F.from_apply_func(lambda ts, p1, p2: ts * (p1 + p2), p2=2)
.run(close, [0, 1], hide_default=False).out)
print(F.from_apply_func(njit(lambda ts, p1, p2: ts * (p1 + p2)), p2=2)
.run(close, [0, 1], hide_default=False).out)
print(F.from_apply_func(lambda ts, p1, p2: ts * (p1 + p2), p2=2)
.run(close, [0, 1], hide_default=True).out)
print(F.from_apply_func(njit(lambda ts, p1, p2: ts * (p1 + p2)), p2=2)
.run(close, [0, 1], hide_default=True).out)
# test multiple outputs
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p'], output_names=['o1', 'o2'])
print(F.from_apply_func(lambda ts, p: (ts * p, ts * p ** 2))
.run(close, [0, 1]).o1)
print(F.from_apply_func(lambda ts, p: (ts * p, ts * p ** 2))
.run(close, [0, 1]).o2)
print(F.from_apply_func(njit(lambda ts, p: (ts * p, ts * p ** 2)))
.run(close, [0, 1]).o1)
print(F.from_apply_func(njit(lambda ts, p: (ts * p, ts * p ** 2)))
.run(close, [0, 1]).o2)
# test in-place outputs
def apply_func(ts, ts_out, p):
ts_out[:, 0] = p
return ts * p
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p'], output_names=['out'], in_output_names=['ts_out'])
print(F.from_apply_func(apply_func)
.run(close, [0, 1]).ts_out)
print(F.from_apply_func(njit(apply_func))
.run(close, [0, 1]).ts_out)
print(F.from_apply_func(apply_func, in_output_settings={'ts_out': {'dtype': np.int_}})
.run(close, [0, 1]).ts_out)
print(F.from_apply_func(njit(apply_func), in_output_settings={'ts_out': {'dtype': np.int_}})
.run(close, [0, 1]).ts_out)
print(F.from_apply_func(apply_func, ts_out=-1)
.run(close, [0, 1]).ts_out)
print(F.from_apply_func(njit(apply_func), ts_out=-1)
.run(close, [0, 1]).ts_out)
# test kwargs_to_args
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p'], output_names=['out'])
print(F.from_apply_func(lambda ts, p, a, kw: ts * p + a + kw, kwargs_to_args=['kw'], var_args=True)
.run(close, [0, 1, 2], 3, kw=10).out)
print(F.from_apply_func(njit(lambda ts, p, a, kw: ts * p + a + kw), kwargs_to_args=['kw'], var_args=True)
.run(close, [0, 1, 2], 3, kw=10).out)
# test caching func
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p'], output_names=['out'])
print(F.from_apply_func(lambda ts, param, c: ts * param + c, cache_func=lambda ts, params: 100)
.run(close, [0, 1]).out)
print(F.from_apply_func(njit(lambda ts, param, c: ts * param + c), cache_func=njit(lambda ts, params: 100))
.run(close, [0, 1]).out)
# test run_combs
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p1', 'p2'], output_names=['out'])
print(F.from_apply_func(lambda ts, p1, p2: ts * (p1 + p2))
.run_combs(close, [0, 1, 2], [3, 4, 5], short_names=['i1', 'i2'])[0].out)
print(F.from_apply_func(lambda ts, p1, p2: ts * (p1 + p2))
.run_combs(close, [0, 1, 2], [3, 4, 5], short_names=['i1', 'i2'])[1].out)
print(F.from_apply_func(njit(lambda ts, p1, p2: ts * (p1 + p2)))
.run_combs(close, [0, 1, 2], [3, 4, 5], short_names=['i1', 'i2'])[0].out)
print(F.from_apply_func(njit(lambda ts, p1, p2: ts * (p1 + p2)))
.run_combs(close, [0, 1, 2], [3, 4, 5], short_names=['i1', 'i2'])[1].out)
from collections import namedtuple
TestEnum = namedtuple('TestEnum', ['Hello', 'World'])(0, 1)
# test attr_settings
F = vbt.IndicatorFactory(
input_names=['ts'], output_names=['o1', 'o2'], in_output_names=['ts_out'],
attr_settings={
'ts': {'dtype': None},
'o1': {'dtype': np.float_},
'o2': {'dtype': np.bool_},
'ts_out': {'dtype': TestEnum}
}
)
dir(F.from_apply_func(lambda ts, ts_out: (ts + ts_out, ts + ts_out)).run(close))
CustomInd = vbt.IndicatorFactory(
input_names=['ts1', 'ts2'],
param_names=['p1', 'p2'],
output_names=['o1', 'o2']
).from_apply_func(lambda ts1, ts2, p1, p2: (ts1 * p1, ts2 * p2))
dir(CustomInd) # you can list here all of the available tools
custom_ind = CustomInd.run(close, high, [1, 2], [3, 4])
big_custom_ind = CustomInd.run(big_close, big_high, [1, 2], [3, 4])
print(custom_ind.wrapper.index) # subclasses ArrayWrapper
print(custom_ind.wrapper.columns)
print(custom_ind.wrapper.ndim)
print(custom_ind.wrapper.shape)
print(custom_ind.wrapper.freq)
# not changed during indexing
print(custom_ind.short_name)
print(custom_ind.level_names)
print(custom_ind.input_names)
print(custom_ind.param_names)
print(custom_ind.output_names)
print(custom_ind.output_flags)
print(custom_ind.p1_list)
print(custom_ind.p2_list)
```
### Pandas indexing
```
print(custom_ind._ts1)
print(custom_ind.ts1)
print(custom_ind.ts1.iloc[:, 0])
print(custom_ind.iloc[:, 0].ts1)
print(custom_ind.ts1.iloc[:, [0]])
print(custom_ind.iloc[:, [0]].ts1)
print(custom_ind.ts1.iloc[:2, :])
print(custom_ind.iloc[:2, :].ts1)
print(custom_ind.o1.iloc[:, 0])
%timeit big_custom_ind.o1.iloc[:, 0] # benchmark, 1 column
print(custom_ind.iloc[:, 0].o1) # performed on the object itself
%timeit big_custom_ind.iloc[:, 0] # slower since it forwards the operation to each dataframe
print(custom_ind.o1.iloc[:, np.arange(3)])
%timeit big_custom_ind.o1.iloc[:, np.arange(1000)] # 1000 columns
print(custom_ind.iloc[:, np.arange(3)].o1)
%timeit big_custom_ind.iloc[:, np.arange(1000)]
print(custom_ind.o1.loc[:, (1, 3, 'a')])
%timeit big_custom_ind.o1.loc[:, (1, 3, 0)] # 1 column
print(custom_ind.loc[:, (1, 3, 'a')].o1)
%timeit big_custom_ind.loc[:, (1, 3, 0)]
print(custom_ind.o1.loc[:, (1, 3)])
%timeit big_custom_ind.o1.loc[:, 1] # 1000 columns
print(custom_ind.loc[:, (1, 3)].o1)
%timeit big_custom_ind.loc[:, 1]
print(custom_ind.o1.xs(1, axis=1, level=0))
%timeit big_custom_ind.o1.xs(1, axis=1, level=0) # 1000 columns
print(custom_ind.xs(1, axis=1, level=0).o1)
%timeit big_custom_ind.xs(1, axis=1, level=0)
```
### Parameter indexing
```
# Indexing by parameter
print(custom_ind._p1_mapper)
print(custom_ind.p1_loc[2].o1)
print(custom_ind.p1_loc[1:2].o1)
print(custom_ind.p1_loc[[1, 1, 1]].o1)
%timeit big_custom_ind.p1_loc[1] # 1000 columns
%timeit big_custom_ind.p1_loc[np.full(10, 1)] # 10000 columns
print(custom_ind._tuple_mapper)
print(custom_ind.tuple_loc[(1, 3)].o1)
print(custom_ind.tuple_loc[(1, 3):(2, 4)].o1)
%timeit big_custom_ind.tuple_loc[(1, 3)]
%timeit big_custom_ind.tuple_loc[[(1, 3)] * 10]
```
### Comparison methods
```
print(custom_ind.o1 > 2)
%timeit big_custom_ind.o1.values > 2 # don't even try pandas
print(custom_ind.o1_above(2))
%timeit big_custom_ind.o1_above(2) # slower than numpy because of constructing dataframe
print(pd.concat((custom_ind.o1 > 2, custom_ind.o1 > 3), axis=1))
%timeit np.hstack((big_custom_ind.o1.values > 2, big_custom_ind.o1.values > 3))
print(custom_ind.o1_above([2, 3]))
%timeit big_custom_ind.o1_above([2, 3])
```
## TA-Lib
```
ts = pd.DataFrame({
'a': [1, 2, 3, 4, np.nan],
'b': [np.nan, 4, 3, 2, 1],
'c': [1, 2, np.nan, 2, 1]
}, index=pd.DatetimeIndex([
datetime(2018, 1, 1),
datetime(2018, 1, 2),
datetime(2018, 1, 3),
datetime(2018, 1, 4),
datetime(2018, 1, 5)
]))
SMA = vbt.talib('SMA')
print(SMA.run(close['a'], 2).real)
print(SMA.run(close, 2).real)
print(SMA.run(close, [2, 3]).real)
%timeit SMA.run(big_close)
%timeit SMA.run(big_close, np.arange(2, 10))
%timeit SMA.run(big_close, np.full(10, 2))
%timeit SMA.run(big_close, np.full(10, 2), speedup=True)
comb = itertools.combinations(np.arange(2, 20), 2)
fast_windows, slow_windows = np.asarray(list(comb)).transpose()
print(fast_windows)
print(slow_windows)
%timeit SMA.run(big_close, fast_windows), SMA.run(big_close, slow_windows) # individual caching
%timeit SMA.run_combs(big_close, np.arange(2, 20)) # mutual caching
%timeit vbt.MA.run(big_close, fast_windows), vbt.MA.run(big_close, slow_windows) # the same using Numba
%timeit vbt.MA.run_combs(big_close, np.arange(2, 20))
sma1, sma2 = SMA.run_combs(close, [2, 3, 4])
print(sma1.real_above(sma2, crossover=True))
print(sma1.real_below(sma2, crossover=True))
dir(vbt.talib('BBANDS'))
```
## MA
```
print(close.rolling(2).mean())
print(close.ewm(span=3, min_periods=3).mean())
print(vbt.talib('SMA').run(close, timeperiod=2).real)
print(vbt.MA.run(close, [2, 3], ewm=[False, True]).ma) # adjust=False
# One window
%timeit big_close.rolling(2).mean() # pandas
%timeit vbt.talib('SMA').run(big_close, timeperiod=2)
%timeit vbt.MA.run(big_close, 2, return_cache=True) # cache only
%timeit vbt.MA.run(big_close, 2) # with pre+postprocessing and still beats pandas
print(vbt.MA.run(big_close, 2).ma.shape)
# Multiple windows
%timeit pd.concat([big_close.rolling(i).mean() for i in np.arange(2, 10)])
%timeit vbt.talib('SMA').run(big_close, np.arange(2, 10))
%timeit vbt.MA.run(big_close, np.arange(2, 10))
%timeit vbt.MA.run(big_close, np.arange(2, 10), speedup=True)
%timeit vbt.MA.run(big_close, np.arange(2, 10), return_cache=True) # cache only
cache = vbt.MA.run(big_close, np.arange(2, 10), return_cache=True)
%timeit vbt.MA.run(big_close, np.arange(2, 10), use_cache=cache) # using cache
print(vbt.MA.run(big_close, np.arange(2, 10)).ma.shape)
# One window repeated
%timeit pd.concat([big_close.rolling(i).mean() for i in np.full(10, 2)])
%timeit vbt.talib('SMA').run(big_close, np.full(10, 2))
%timeit vbt.MA.run(big_close, np.full(10, 2))
%timeit vbt.MA.run(big_close, np.full(10, 2), speedup=True) # slower for large inputs
%timeit vbt.MA.run(big_close, np.full(10, 2), return_cache=True)
print(vbt.MA.run(big_close, np.full(10, 2)).ma.shape)
%timeit pd.concat([big_close.iloc[:, :10].rolling(i).mean() for i in np.full(100, 2)])
%timeit vbt.talib('SMA').run(big_close.iloc[:, :10], np.full(100, 2))
%timeit vbt.MA.run(big_close.iloc[:, :10], np.full(100, 2))
%timeit vbt.MA.run(big_close.iloc[:, :10], np.full(100, 2), speedup=True) # faster for smaller inputs
%timeit vbt.MA.run(big_close.iloc[:, :10], np.full(100, 2), return_cache=True)
print(vbt.MA.run(big_close.iloc[:, :10], np.full(100, 2)).ma.shape)
ma = vbt.MA.run(close, [2, 3], ewm=[False, True])
print(ma.ma)
ma[(2, False, 'a')].plot().show_svg()
```
## MSTD
```
print(close.rolling(2).std(ddof=0))
print(close.ewm(span=3, min_periods=3).std(ddof=0))
print(vbt.talib('STDDEV').run(close, timeperiod=2).real)
print(vbt.MSTD.run(close, [2, 3], ewm=[False, True]).mstd) # adjust=False, ddof=0
# One window
%timeit big_close.rolling(2).std()
%timeit vbt.talib('STDDEV').run(big_close, timeperiod=2)
%timeit vbt.MSTD.run(big_close, 2)
print(vbt.MSTD.run(big_close, 2).mstd.shape)
# Multiple windows
%timeit pd.concat([big_close.rolling(i).std() for i in np.arange(2, 10)])
%timeit vbt.talib('STDDEV').run(big_close, timeperiod=np.arange(2, 10))
%timeit vbt.MSTD.run(big_close, np.arange(2, 10))
print(vbt.MSTD.run(big_close, np.arange(2, 10)).mstd.shape)
# One window repeated
%timeit vbt.talib('STDDEV').run(big_close, timeperiod=np.full(10, 2))
%timeit vbt.MSTD.run(big_close, window=np.full(10, 2))
print(vbt.MSTD.run(big_close, window=np.full(10, 2)).close.shape)
mstd = vbt.MSTD.run(close, [2, 3], [False, True])
print(mstd.mstd)
mstd[(2, False, 'a')].plot().show_svg()
```
## BBANDS
```
print(vbt.ta('BollingerBands').run(close['a'], window=2, window_dev=2).bollinger_hband)
print(vbt.ta('BollingerBands').run(close['a'], window=2, window_dev=2).bollinger_mavg)
print(vbt.ta('BollingerBands').run(close['a'], window=2, window_dev=2).bollinger_lband)
print(vbt.talib('BBANDS').run(close, timeperiod=2, nbdevup=2, nbdevdn=2).upperband)
print(vbt.talib('BBANDS').run(close, timeperiod=2, nbdevup=2, nbdevdn=2).middleband)
print(vbt.talib('BBANDS').run(close, timeperiod=2, nbdevup=2, nbdevdn=2).lowerband)
print(vbt.BBANDS.run(close, window=2, ewm=False, alpha=2).upper)
print(vbt.BBANDS.run(close, window=2, ewm=False, alpha=2).middle)
print(vbt.BBANDS.run(close, window=2, ewm=False, alpha=2).lower)
# One window
%timeit vbt.talib('BBANDS').run(big_close, timeperiod=2)
%timeit vbt.BBANDS.run(big_close, window=2)
print(vbt.BBANDS.run(big_close).close.shape)
# Multiple windows
%timeit vbt.talib('BBANDS').run(big_close, timeperiod=np.arange(2, 10))
%timeit vbt.BBANDS.run(big_close, window=np.arange(2, 10))
print(vbt.BBANDS.run(big_close, window=np.arange(2, 10)).close.shape)
# One window repeated
%timeit vbt.talib('BBANDS').run(big_close, timeperiod=np.full(10, 2))
%timeit vbt.BBANDS.run(big_close, window=np.full(10, 2))
print(vbt.BBANDS.run(big_close, window=np.full(10, 2)).close.shape)
bb = vbt.BBANDS.run(close, window=2, alpha=[1., 2.], ewm=False)
print(bb.middle)
print()
print(bb.upper)
print()
print(bb.lower)
print()
print(bb.percent_b)
print()
print(bb.bandwidth)
print(bb.close_below(bb.upper) & bb.close_above(bb.lower)) # price between bands
bb[(2, False, 1., 'a')].plot().show_svg()
```
## RSI
```
print(vbt.ta('RSIIndicator').run(close=close['a'], window=2).rsi) # alpha=1/n
print(vbt.ta('RSIIndicator').run(close=close['b'], window=2).rsi)
print(vbt.ta('RSIIndicator').run(close=close['c'], window=2).rsi)
print(vbt.talib('RSI').run(close, timeperiod=2).real)
print(vbt.RSI.run(close, window=[2, 2], ewm=[True, False]).rsi) # span=n
# One window
%timeit vbt.talib('RSI').run(big_close, timeperiod=2)
%timeit vbt.RSI.run(big_close, window=2)
print(vbt.RSI.run(big_close, window=2).rsi.shape)
# Multiple windows
%timeit vbt.talib('RSI').run(big_close, timeperiod=np.arange(2, 10))
%timeit vbt.RSI.run(big_close, window=np.arange(2, 10))
print(vbt.RSI.run(big_close, window=np.arange(2, 10)).rsi.shape)
# One window repeated
%timeit vbt.talib('RSI').run(big_close, timeperiod=np.full(10, 2))
%timeit vbt.RSI.run(big_close, window=np.full(10, 2))
print(vbt.RSI.run(big_close, window=np.full(10, 2)).rsi.shape)
rsi = vbt.RSI.run(close, window=[2, 3], ewm=[False, True])
print(rsi.rsi)
print(rsi.rsi_above(70))
rsi[(2, False, 'a')].plot().show_svg()
```
## STOCH
```
print(vbt.ta('StochasticOscillator').run(high=high['a'], low=low['a'], close=close['a'], window=2, smooth_window=3).stoch)
print(vbt.ta('StochasticOscillator').run(high=high['a'], low=low['a'], close=close['a'], window=2, smooth_window=3).stoch_signal)
print(vbt.talib('STOCHF').run(high, low, close, fastk_period=2, fastd_period=3).fastk)
print(vbt.talib('STOCHF').run(high, low, close, fastk_period=2, fastd_period=3).fastd)
print(vbt.STOCH.run(high, low, close, k_window=2, d_window=3).percent_k)
print(vbt.STOCH.run(high, low, close, k_window=2, d_window=3).percent_d)
# One window
%timeit vbt.talib('STOCHF').run(big_high, big_low, big_close, fastk_period=2)
%timeit vbt.STOCH.run(big_high, big_low, big_close, k_window=2)
print(vbt.STOCH.run(big_high, big_low, big_close, k_window=2).percent_d.shape)
# Multiple windows
%timeit vbt.talib('STOCHF').run(big_high, big_low, big_close, fastk_period=np.arange(2, 10))
%timeit vbt.STOCH.run(big_high, big_low, big_close, k_window=np.arange(2, 10))
print(vbt.STOCH.run(big_high, big_low, big_close, k_window=np.arange(2, 10)).percent_d.shape)
# One window repeated
%timeit vbt.talib('STOCHF').run(big_high, big_low, big_close, fastk_period=np.full(10, 2))
%timeit vbt.STOCH.run(big_high, big_low, big_close, k_window=np.full(10, 2))
print(vbt.STOCH.run(big_high, big_low, big_close, k_window=np.full(10, 2)).percent_d.shape)
stochastic = vbt.STOCH.run(high, low, close, k_window=[2, 4], d_window=2, d_ewm=[False, True])
print(stochastic.percent_k)
print(stochastic.percent_d)
stochastic[(2, 2, False, 'a')].plot().show_svg()
```
## MACD
```
print(vbt.ta('MACD').run(close['a'], window_fast=2, window_slow=3, window_sign=2).macd)
print(vbt.ta('MACD').run(close['a'], window_fast=2, window_slow=3, window_sign=2).macd_signal)
print(vbt.ta('MACD').run(close['a'], window_fast=2, window_slow=3, window_sign=2).macd_diff)
print(vbt.talib('MACD').run(close, fastperiod=2, slowperiod=3, signalperiod=2).macd) # uses sma
print(vbt.talib('MACD').run(close, fastperiod=2, slowperiod=3, signalperiod=2).macdsignal)
print(vbt.talib('MACD').run(close, fastperiod=2, slowperiod=3, signalperiod=2).macdhist)
print(vbt.MACD.run(close, fast_window=2, slow_window=3, signal_window=2, macd_ewm=True, signal_ewm=True).macd)
print(vbt.MACD.run(close, fast_window=2, slow_window=3, signal_window=2, macd_ewm=True, signal_ewm=True).signal)
print(vbt.MACD.run(close, fast_window=2, slow_window=3, signal_window=2, macd_ewm=True, signal_ewm=True).hist)
# One window
%timeit vbt.talib('MACD').run(big_close, fastperiod=2)
%timeit vbt.MACD.run(big_close, fast_window=2)
print(vbt.MACD.run(big_close, fast_window=2).macd.shape)
# Multiple windows
%timeit vbt.talib('MACD').run(big_close, fastperiod=np.arange(2, 10))
%timeit vbt.MACD.run(big_close, fast_window=np.arange(2, 10))
print(vbt.MACD.run(big_close, fast_window=np.arange(2, 10)).macd.shape)
# One window repeated
%timeit vbt.talib('MACD').run(big_close, fastperiod=np.full(10, 2))
%timeit vbt.MACD.run(big_close, fast_window=np.full(10, 2))
print(vbt.MACD.run(big_close, fast_window=np.full(10, 2)).macd.shape)
macd = vbt.MACD.run(close, fast_window=2, slow_window=3, signal_window=[2, 3], macd_ewm=True, signal_ewm=True)
print(macd.macd)
print(macd.signal)
print(macd.hist)
macd[(2, 3, 2, True, True, 'a')].plot().show_svg()
```
## ATR
```
print(vbt.ta('AverageTrueRange').run(high['a'], low['a'], close['a'], window=2).average_true_range)
print(vbt.ta('AverageTrueRange').run(high['b'], low['b'], close['b'], window=2).average_true_range)
print(vbt.ta('AverageTrueRange').run(high['c'], low['c'], close['c'], window=2).average_true_range)
print(vbt.talib('ATR').run(high, low, close, timeperiod=2).real)
print(vbt.ATR.run(high, low, close, window=[2, 3], ewm=[False, True]).atr)
# One window
%timeit vbt.talib('ATR').run(big_high, big_low, big_close, timeperiod=2)
%timeit vbt.ATR.run(big_high, big_low, big_close, window=2)
print(vbt.ATR.run(big_high, big_low, big_close, window=2).atr.shape)
# Multiple windows
%timeit vbt.talib('ATR').run(big_high, big_low, big_close, timeperiod=np.arange(2, 10))
%timeit vbt.ATR.run(big_high, big_low, big_close, window=np.arange(2, 10)) # rolling min/max very expensive
print(vbt.ATR.run(big_high, big_low, big_close, window=np.arange(2, 10)).atr.shape)
# One window repeated
%timeit vbt.talib('ATR').run(big_high, big_low, big_close, timeperiod=np.full(10, 2))
%timeit vbt.ATR.run(big_high, big_low, big_close, window=np.full(10, 2))
print(vbt.ATR.run(big_high, big_low, big_close, window=np.full(10, 2)).atr.shape)
atr = vbt.ATR.run(high, low, close, window=[2, 3], ewm=[False, True])
print(atr.tr)
print(atr.atr)
atr[(2, False, 'a')].plot().show_svg()
```
## OBV
```
print(vbt.ta('OnBalanceVolumeIndicator').run(close['a'], volume['a']).on_balance_volume)
print(vbt.ta('OnBalanceVolumeIndicator').run(close['b'], volume['b']).on_balance_volume)
print(vbt.ta('OnBalanceVolumeIndicator').run(close['c'], volume['c']).on_balance_volume)
print(vbt.talib('OBV').run(close, volume).real)
print(vbt.OBV.run(close, volume).obv)
%timeit vbt.talib('OBV').run(big_close, big_volume)
%timeit vbt.OBV.run(big_close, big_volume)
print(vbt.OBV.run(big_close, big_volume).obv.shape)
obv = vbt.OBV.run(close, volume)
print(obv.obv)
print(obv.obv_above([0, 5]))
obv['a'].plot().show_svg()
```
| github_jupyter |
## API Time Selectors - Using Start, End, and Reftime with Forecast Data
Forecast datasets typically have two time dimensions, `reftime` and `time`. The `reftime` of a forecast data product represents the run time of the forecast model, while `time` represents the actual time (typically in the future) that the values are valid. For each `reftime`, values are unique for each unique `time` value.
Typically the most recent forecast is most desirable to users, and the Planet OS API intentionally returns these values by default. However, there may be cases when one wants to investigate a historical forecast, or how values at a specific time differ between successive forecasts.
The [RTOFS data product](http://data.planetos.com/datasets/noaa_rtofs_surface_1h_diag) is a daily updated forecast that provides a 3-day, hourly forecast. This notebook highlights ways to use the start, end, reftime_start, and reftime_end API parameters and explain the expected response.
```
%matplotlib inline
import pandas as pd
import simplejson as json
from urllib.parse import urlencode
from urllib.request import urlopen, Request
import datetime
import numpy as np
```
A Planet OS API key is required to run this notebook. Keys are displayed in the [account settings](http://data.planetos.com/account/settings/) page on the Planet OS Datahub. If you do not have a Planet OS account, you can [sign up for free](http://data.planetos.com/plans).
```
apikey = open('APIKEY').readlines()[0].strip() #'<YOUR API KEY HERE>'
```
## Request the Most Recent Model Run
By default, our API returns the most recent model run. The example below returns all variables for the most recent forecast run (e.g. reftime). At the time of running, this was 2016-11-12T00:00:00. Given the hourly resolution and 3-day extent, we expect 72 unique values to be returned.
```
# Set the Planet OS API query parameters
count = 100
id = 'noaa_rtofs_surface_1h_diag'
lat=39.37858604638528
lon=-72.57739563685647
time_order = 'desc'
query_dict = {'apikey': apikey,
'count': count,
'lon': lon,
'lat': lat,
'time_order': time_order,
}
query = urlencode(query_dict)
api_url = "http://api.planetos.com/v1/datasets/%s/point?%s" % (id, query)
request = Request(api_url)
response = urlopen(request)
response_json = json.loads(response.read())
data = response_json['entries']
# let's flatten the response and create a Pandas dataframe
df = pd.io.json.json_normalize(data)
# then index by time using the axes.time column
pd.to_datetime(df["axes.time"])
df.set_index('axes.time', inplace=False)
print(df.count())
df.head()
```
## Request a Single Model Run
By default, the system will return the most recent model run (e.g. reftime). However, a specific model run can be acquired by requesting an identical reftime_start and reftime_end value. In the example below, we request values from only the 2016-11-12T00:00:00 model run.
```
df
# Request only data from a specific model run
reftime_end = np.max(df['axes.reftime'])
reftime_start = np.max(df['axes.reftime'])
query_dict = {'apikey': apikey,
'count': count,
'lat': lat,
'lon': lon,
'reftime_end': reftime_end,
'reftime_start': reftime_start,
'time_order': time_order,
}
query = urlencode(query_dict)
api_url = "http://api.planetos.com/v1/datasets/%s/point?%s" % (id, query)
request = Request(api_url)
response = urlopen(request)
response_json = json.loads(response.read())
data = response_json['entries']
# let's flatten the response and create a Pandas dataframe
df = pd.io.json.json_normalize(data)
# then index by time using the axes.time column
pd.to_datetime(df["axes.time"])
df.set_index('axes.time', inplace=False)
print(df.count())
```
### All available model run values for a specific time
This example looks for data at a specific future datetime (e.g. 2016-11-14T00:00:00) and returns values from all available model runs (e.g. reftimes). From the result at the time of running, we can see that two forecasts provide values for Nov 14th at 00:00, the 2016-11-11T00:00:00 and 2016-11-12T00:00:00 model runs.
```
# Request only data from past model run (e.g. 2016-11-11T00:00:00)
end = np.min(df['axes.time'])
start = datetime.datetime.strftime(datetime.datetime.strptime(df['axes.reftime'][0],'%Y-%m-%dT%H:%M:%S') - datetime.timedelta(days=1),'%Y-%m-%dT%H:%M:%S')
query_dict = {'apikey': apikey,
'count': count,
'end': end,
'lat': lat,
'lon': lon,
'reftime_recent': 'false',
'start': start,
'time_order': time_order,
}
query = urlencode(query_dict)
api_url = "http://api.planetos.com/v1/datasets/%s/point?%s" % (id, query)
request = Request(api_url)
response = urlopen(request)
response_json = json.loads(response.read())
data = response_json['entries']
# let's flatten the response and create a Pandas dataframe
df = pd.io.json.json_normalize(data)
print (df)
# then index by time using the axes.time column
pd.to_datetime(df["axes.time"])
df.set_index('axes.time', inplace=True)
print(df.count())
df.head()
```
| github_jupyter |
# Fact Retreival Bot using IDFT
### Steps
- Loading and preprocessing Questions and Answers from dataset
- Setting Stopwords
- Intitialising and training TF_IDF vectors
- Testins
## Imports
```
import pandas as pd # To load and process dataset
import numpy as np # For matrix operations
from nltk.corpus import stopwords # Using NLTK to load stopwords
from nltk import word_tokenize # Using NLTK to token sentences
from sklearn.feature_extraction.text import TfidfVectorizer
pd.set_option('display.width',1000)
```
## Loading and preprocessing Questions and Answers from dataset
- `hdfc.pkl` : Collection of 1341 QnA about HDFC. (Scraped from HDFC's FAQ site)
- Dropping stopwords
- Stripping Questions of extra spaces
```
df = pd.read_pickle('hdfc.pkl')
df = df.drop_duplicates('Question')
df = df.reset_index()
df['Question'] = df['Question'].str.strip()
limit = 100
reduced = df[['Question','Answer']][:limit]
qlabels = reduced['Question'].to_dict()
alabels = reduced['Answer'].to_dict()
print reduced.head()
```
## Setting stopwords
- Import set of common stopwords from nltk
- Adding domain-related stopword
- Removing question words (To distinguish between intents of questions)
```
# Loading stopwords
plus = {'hdfc'}
minus = {'what','how','where','when','why'}
stop = set(stopwords.words('english'))
stop.update(plus)
stop.difference_update(minus)
```
## Intitialising and training TF-IDF vectors
- Setting stopwords to `stop`
- `tf_vect` : `TfidfVectorizer` object. Can be used to convert strings to tf-idf vectors
- `all_qs_vectors` : Matrix of TF-IDF vectors corresponding to questions in training set
```
tf_vect =TfidfVectorizer(stop_words=stop,
lowercase=True,
use_idf=True)
all_qs_vectors = tf_vect.fit_transform(reduced['Question'])
print "Shape of all_qs_vectors :",all_qs_vectors.shape
print all_qs_vectors.shape[0],": Number of questions"
print all_qs_vectors.shape[1],": Vocabulary size"
# Transforming context with tfidf
context = 'How can I repay my Personal Loan?'
context_vector = tf_vect.transform([context])
context_matrix = context_vector.todense()
# Displaying TF_IDF results
print "WORD".ljust(10),"INDEX".ljust(6),"TFIDF_VALUE"
for w in word_tokenize(context.strip()):
ind = tf_vect.vocabulary_.get(w.lower(),"NA")
val = context_matrix[0,ind] if not ind == "NA" else 0
print w.ljust(10),str(ind).ljust(6),val
```
## Predicting closest question
- `predict` has the following arguments
- `n` : int | Number of results (from top)
- `answers` : bool | Return answers or not
- `ret_best`: bool | Returns index of closest match
- Steps for prediction
- Convert query to tfidf vector
- Get dot product of query vectors with each question to measures similarity
- Sort array indices by descending order of array values
- Return top n results
```
def predict(query,n=5,answers=False,ret_indices=False):
# Comparing context with all questions using dot product
query_vector = tf_vect.transform([query])
sim = np.dot(all_qs_vectors, query_vector.T)
# Converting numpy matrix to 1D array with 146 dot products (146 questions vs context)
arr = sim.toarray().flatten()
matches = arr.argsort(axis=0)[::-1]
top_n_matches = matches[:n]
results = []
if ret_indices:
return top_n_matches
for i in top_n_matches:
res = {qlabels[i]:alabels[i]} if answers else qlabels[i]
results.append(res)
return results
predict('How do I pay my personal loan ?')
num_correct = 0
failed = []
for i in qlabels:
if predict(qlabels[i],n=1,ret_indices=True)[0] == i:
num_correct +=1
else :
failed.append(i)
print "Recall : ",float(num_correct)/len(qlabels) *100,"%"
for i in failed :
query = qlabels[i]
print "\nQuery :",query
print predict(query,n=3)
```
| github_jupyter |
```
import torch.utils.data as utils
import torch.nn.functional as F
import torch
import torch.nn as nn
from torch.autograd import Variable
from torch.nn.parameter import Parameter
import numpy as np
import pandas as pd
import math
import time
import matplotlib.pyplot as plt
%matplotlib inline
print(torch.__version__)
def PrepareDataset(speed_matrix, BATCH_SIZE = 40, seq_len = 10, pred_len = 1, train_propotion = 0.7, valid_propotion = 0.2):
""" Prepare training and testing datasets and dataloaders.
Convert speed/volume/occupancy matrix to training and testing dataset.
The vertical axis of speed_matrix is the time axis and the horizontal axis
is the spatial axis.
Args:
speed_matrix: a Matrix containing spatial-temporal speed data for a network
seq_len: length of input sequence
pred_len: length of predicted sequence
Returns:
Training dataloader
Testing dataloader
"""
time_len = speed_matrix.shape[0]
max_speed = speed_matrix.max().max()
speed_matrix = speed_matrix / max_speed
speed_sequences, speed_labels = [], []
for i in range(time_len - seq_len - pred_len):
speed_sequences.append(speed_matrix.iloc[i:i+seq_len].values)
speed_labels.append(speed_matrix.iloc[i+seq_len:i+seq_len+pred_len].values)
speed_sequences, speed_labels = np.asarray(speed_sequences), np.asarray(speed_labels)
# shuffle and split the dataset to training and testing datasets
sample_size = speed_sequences.shape[0]
index = np.arange(sample_size, dtype = int)
np.random.shuffle(index)
train_index = int(np.floor(sample_size * train_propotion))
valid_index = int(np.floor(sample_size * ( train_propotion + valid_propotion)))
train_data, train_label = speed_sequences[:train_index], speed_labels[:train_index]
valid_data, valid_label = speed_sequences[train_index:valid_index], speed_labels[train_index:valid_index]
test_data, test_label = speed_sequences[valid_index:], speed_labels[valid_index:]
train_data, train_label = torch.Tensor(train_data), torch.Tensor(train_label)
valid_data, valid_label = torch.Tensor(valid_data), torch.Tensor(valid_label)
test_data, test_label = torch.Tensor(test_data), torch.Tensor(test_label)
train_dataset = utils.TensorDataset(train_data, train_label)
valid_dataset = utils.TensorDataset(valid_data, valid_label)
test_dataset = utils.TensorDataset(test_data, test_label)
train_dataloader = utils.DataLoader(train_dataset, batch_size = BATCH_SIZE, shuffle=True, drop_last = True)
valid_dataloader = utils.DataLoader(valid_dataset, batch_size = BATCH_SIZE, shuffle=True, drop_last = True)
test_dataloader = utils.DataLoader(test_dataset, batch_size = BATCH_SIZE, shuffle=True, drop_last = True)
return train_dataloader, valid_dataloader, test_dataloader, max_speed
if __name__ == "__main__":
# data = 'inrix'
data = 'loop'
directory = '../../Data_Warehouse/Data_network_traffic/'
if data == 'inrix':
speed_matrix = pd.read_pickle( directory + 'inrix_seattle_speed_matrix_2012')
A = np.load(directory + 'INRIX_Seattle_2012_A.npy')
FFR_5min = np.load(directory + 'INRIX_Seattle_2012_reachability_free_flow_5min.npy')
FFR_10min = np.load(directory + 'INRIX_Seattle_2012_reachability_free_flow_10min.npy')
FFR_15min = np.load(directory + 'INRIX_Seattle_2012_reachability_free_flow_15min.npy')
FFR_20min = np.load(directory + 'INRIX_Seattle_2012_reachability_free_flow_20min.npy')
FFR_25min = np.load(directory + 'INRIX_Seattle_2012_reachability_free_flow_25min.npy')
FFR = [FFR_5min, FFR_10min, FFR_15min, FFR_20min, FFR_25min]
elif data == 'loop':
speed_matrix = pd.read_pickle( directory + 'speed_matrix_2015')
A = np.load( directory + 'Loop_Seattle_2015_A.npy')
FFR_5min = np.load( directory + 'Loop_Seattle_2015_reachability_free_flow_5min.npy')
FFR_10min = np.load( directory + 'Loop_Seattle_2015_reachability_free_flow_10min.npy')
FFR_15min = np.load( directory + 'Loop_Seattle_2015_reachability_free_flow_15min.npy')
FFR_20min = np.load( directory + 'Loop_Seattle_2015_reachability_free_flow_20min.npy')
FFR_25min = np.load( directory + 'Loop_Seattle_2015_reachability_free_flow_25min.npy')
FFR = [FFR_5min, FFR_10min, FFR_15min, FFR_20min, FFR_25min]
train_dataloader, valid_dataloader, test_dataloader, max_speed = PrepareDataset(speed_matrix)
inputs, labels = next(iter(train_dataloader))
[batch_size, step_size, fea_size] = inputs.size()
input_dim = fea_size
hidden_dim = fea_size
output_dim = fea_size
def TrainModel(model, train_dataloader, valid_dataloader, learning_rate = 1e-5, num_epochs = 300, patience = 10, min_delta = 0.00001):
inputs, labels = next(iter(train_dataloader))
[batch_size, step_size, fea_size] = inputs.size()
input_dim = fea_size
hidden_dim = fea_size
output_dim = fea_size
model.cuda()
loss_MSE = torch.nn.MSELoss()
loss_L1 = torch.nn.L1Loss()
learning_rate = 1e-5
optimizer = torch.optim.RMSprop(model.parameters(), lr = learning_rate)
use_gpu = torch.cuda.is_available()
interval = 100
losses_train = []
losses_valid = []
losses_epochs_train = []
losses_epochs_valid = []
cur_time = time.time()
pre_time = time.time()
# Variables for Early Stopping
is_best_model = 0
patient_epoch = 0
for epoch in range(num_epochs):
# print('Epoch {}/{}'.format(epoch, num_epochs - 1))
# print('-' * 10)
trained_number = 0
valid_dataloader_iter = iter(valid_dataloader)
losses_epoch_train = []
losses_epoch_valid = []
for data in train_dataloader:
inputs, labels = data
if inputs.shape[0] != batch_size:
continue
if use_gpu:
inputs, labels = Variable(inputs.cuda()), Variable(labels.cuda())
else:
inputs, labels = Variable(inputs), Variable(labels)
model.zero_grad()
outputs = model(inputs)
loss_train = loss_MSE(outputs, torch.squeeze(labels))
losses_train.append(loss_train.data)
losses_epoch_train.append(loss_train.data)
optimizer.zero_grad()
loss_train.backward()
optimizer.step()
# validation
try:
inputs_val, labels_val = next(valid_dataloader_iter)
except StopIteration:
valid_dataloader_iter = iter(valid_dataloader)
inputs_val, labels_val = next(valid_dataloader_iter)
if use_gpu:
inputs_val, labels_val = Variable(inputs_val.cuda()), Variable(labels_val.cuda())
else:
inputs_val, labels_val = Variable(inputs_val), Variable(labels_val)
outputs_val= model(inputs_val)
loss_valid = loss_MSE(outputs_val, torch.squeeze(labels_val))
losses_valid.append(loss_valid.data)
losses_epoch_valid.append(loss_valid.data)
# output
trained_number += 1
avg_losses_epoch_train = sum(losses_epoch_train) / float(len(losses_epoch_train))
avg_losses_epoch_valid = sum(losses_epoch_valid) / float(len(losses_epoch_valid))
losses_epochs_train.append(avg_losses_epoch_train)
losses_epochs_valid.append(avg_losses_epoch_valid)
# Early Stopping
if epoch == 0:
is_best_model = 1
best_model = model
min_loss_epoch_valid = 10000.0
if avg_losses_epoch_valid < min_loss_epoch_valid:
min_loss_epoch_valid = avg_losses_epoch_valid
else:
if min_loss_epoch_valid - avg_losses_epoch_valid > min_delta:
is_best_model = 1
best_model = model
min_loss_epoch_valid = avg_losses_epoch_valid
patient_epoch = 0
else:
is_best_model = 0
patient_epoch += 1
if patient_epoch >= patience:
print('Early Stopped at Epoch:', epoch)
break
# Print training parameters
cur_time = time.time()
print('Epoch: {}, train_loss: {}, valid_loss: {}, time: {}, best model: {}'.format( \
epoch, \
np.around(avg_losses_epoch_train, decimals=8),\
np.around(avg_losses_epoch_valid, decimals=8),\
np.around([cur_time - pre_time] , decimals=2),\
is_best_model) )
pre_time = cur_time
return best_model, [losses_train, losses_valid, losses_epochs_train, losses_epochs_valid]
def TestModel(model, test_dataloader, max_speed):
inputs, labels = next(iter(test_dataloader))
[batch_size, step_size, fea_size] = inputs.size()
cur_time = time.time()
pre_time = time.time()
use_gpu = torch.cuda.is_available()
loss_MSE = torch.nn.MSELoss()
loss_L1 = torch.nn.MSELoss()
tested_batch = 0
losses_mse = []
losses_l1 = []
for data in test_dataloader:
inputs, labels = data
if inputs.shape[0] != batch_size:
continue
if use_gpu:
inputs, labels = Variable(inputs.cuda()), Variable(labels.cuda())
else:
inputs, labels = Variable(inputs), Variable(labels)
# rnn.loop()
hidden = model.initHidden(batch_size)
outputs = None
outputs = model(inputs)
loss_MSE = torch.nn.MSELoss()
loss_L1 = torch.nn.L1Loss()
loss_mse = loss_MSE(outputs, torch.squeeze(labels))
loss_l1 = loss_L1(outputs, torch.squeeze(labels))
losses_mse.append(loss_mse.cpu().data.numpy())
losses_l1.append(loss_l1.cpu().data.numpy())
tested_batch += 1
if tested_batch % 1000 == 0:
cur_time = time.time()
print('Tested #: {}, loss_l1: {}, loss_mse: {}, time: {}'.format( \
tested_batch * batch_size, \
np.around([loss_l1.data[0]], decimals=8), \
np.around([loss_mse.data[0]], decimals=8), \
np.around([cur_time - pre_time], decimals=8) ) )
pre_time = cur_time
losses_l1 = np.array(losses_l1)
losses_mse = np.array(losses_mse)
mean_l1 = np.mean(losses_l1) * max_speed
std_l1 = np.std(losses_l1) * max_speed
print('Tested: L1_mean: {}, L1_std : {}'.format(mean_l1, std_l1))
return [losses_l1, losses_mse, mean_l1, std_l1]
class LSTM(nn.Module):
def __init__(self, input_size, cell_size, hidden_size, output_last = True):
"""
cell_size is the size of cell_state.
hidden_size is the size of hidden_state, or say the output_state of each step
"""
super(LSTM, self).__init__()
self.cell_size = cell_size
self.hidden_size = hidden_size
self.fl = nn.Linear(input_size + hidden_size, hidden_size)
self.il = nn.Linear(input_size + hidden_size, hidden_size)
self.ol = nn.Linear(input_size + hidden_size, hidden_size)
self.Cl = nn.Linear(input_size + hidden_size, hidden_size)
self.output_last = output_last
def step(self, input, Hidden_State, Cell_State):
combined = torch.cat((input, Hidden_State), 1)
f = F.sigmoid(self.fl(combined))
i = F.sigmoid(self.il(combined))
o = F.sigmoid(self.ol(combined))
C = F.tanh(self.Cl(combined))
Cell_State = f * Cell_State + i * C
Hidden_State = o * F.tanh(Cell_State)
return Hidden_State, Cell_State
def forward(self, inputs):
batch_size = inputs.size(0)
time_step = inputs.size(1)
Hidden_State, Cell_State = self.initHidden(batch_size)
if self.output_last:
for i in range(time_step):
Hidden_State, Cell_State = self.step(torch.squeeze(inputs[:,i:i+1,:]), Hidden_State, Cell_State)
return Hidden_State
else:
outputs = None
for i in range(time_step):
Hidden_State, Cell_State = self.step(torch.squeeze(inputs[:,i:i+1,:]), Hidden_State, Cell_State)
if outputs is None:
outputs = Hidden_State.unsqueeze(1)
else:
outputs = torch.cat((outputs, Hidden_State.unsqueeze(1)), 1)
return outputs
def initHidden(self, batch_size):
use_gpu = torch.cuda.is_available()
if use_gpu:
Hidden_State = Variable(torch.zeros(batch_size, self.hidden_size).cuda())
Cell_State = Variable(torch.zeros(batch_size, self.hidden_size).cuda())
return Hidden_State, Cell_State
else:
Hidden_State = Variable(torch.zeros(batch_size, self.hidden_size))
Cell_State = Variable(torch.zeros(batch_size, self.hidden_size))
return Hidden_State, Cell_State
class ConvLSTM(nn.Module):
def __init__(self, input_size, cell_size, hidden_size, output_last = True):
"""
cell_size is the size of cell_state.
hidden_size is the size of hidden_state, or say the output_state of each step
"""
super(ConvLSTM, self).__init__()
self.cell_size = cell_size
self.hidden_size = hidden_size
self.fl = nn.Linear(input_size + hidden_size, hidden_size)
self.il = nn.Linear(input_size + hidden_size, hidden_size)
self.ol = nn.Linear(input_size + hidden_size, hidden_size)
self.Cl = nn.Linear(input_size + hidden_size, hidden_size)
self.conv = nn.Conv1d(1, hidden_size, hidden_size)
self.output_last = output_last
def step(self, input, Hidden_State, Cell_State):
conv = self.conv(input)
combined = torch.cat((conv, Hidden_State), 1)
f = F.sigmoid(self.fl(combined))
i = F.sigmoid(self.il(combined))
o = F.sigmoid(self.ol(combined))
C = F.tanh(self.Cl(combined))
Cell_State = f * Cell_State + i * C
Hidden_State = o * F.tanh(Cell_State)
return Hidden_State, Cell_State
def forward(self, inputs):
batch_size = inputs.size(0)
time_step = inputs.size(1)
Hidden_State, Cell_State = self.initHidden(batch_size)
if self.output_last:
for i in range(time_step):
Hidden_State, Cell_State = self.step(torch.squeeze(inputs[:,i:i+1,:]), Hidden_State, Cell_State)
return Hidden_State
else:
outputs = None
for i in range(time_step):
Hidden_State, Cell_State = self.step(torch.squeeze(inputs[:,i:i+1,:]), Hidden_State, Cell_State)
if outputs is None:
outputs = Hidden_State.unsqueeze(1)
else:
outputs = torch.cat((outputs, Hidden_State.unsqueeze(1)), 1)
return outputs
def initHidden(self, batch_size):
use_gpu = torch.cuda.is_available()
if use_gpu:
Hidden_State = Variable(torch.zeros(batch_size, self.hidden_size).cuda())
Cell_State = Variable(torch.zeros(batch_size, self.hidden_size).cuda())
return Hidden_State, Cell_State
else:
Hidden_State = Variable(torch.zeros(batch_size, self.hidden_size))
Cell_State = Variable(torch.zeros(batch_size, self.hidden_size))
return Hidden_State, Cell_State
class LocalizedSpectralGraphConvolution(nn.Module):
def __init__(self, A, K):
super(LocalizedSpectralGraphConvolution, self).__init__()
self.K = K
self.A = A.cuda()
feature_size = A.shape[0]
self.D = torch.diag(torch.sum(self.A, dim=0)).cuda()
I = torch.eye(feature_size,feature_size).cuda()
self.L = I - torch.inverse(torch.sqrt(self.D)).matmul(self.A).matmul(torch.inverse(torch.sqrt(self.D)))
L_temp = I
for i in range(K):
L_temp = torch.matmul(L_temp, self.L)
if i == 0:
self.L_tensor = torch.unsqueeze(L_temp, 2)
else:
self.L_tensor = torch.cat((self.L_tensor, torch.unsqueeze(L_temp, 2)), 2)
self.L_tensor = Variable(self.L_tensor.cuda(), requires_grad=False)
self.params = Parameter(torch.FloatTensor(K).cuda())
stdv = 1. / math.sqrt(K)
for i in range(K):
self.params[i].data.uniform_(-stdv, stdv)
def forward(self, input):
x = input
conv = x.matmul( torch.sum(self.params.expand_as(self.L_tensor) * self.L_tensor, 2) )
return conv
class LocalizedSpectralGraphConvolutionalLSTM(nn.Module):
def __init__(self, K, A, feature_size, Clamp_A=True, output_last = True):
'''
Args:
K: K-hop graph
A: adjacency matrix
FFR: free-flow reachability matrix
feature_size: the dimension of features
Clamp_A: Boolean value, clamping all elements of A between 0. to 1.
'''
super(LocalizedSpectralGraphConvolutionalLSTM, self).__init__()
self.feature_size = feature_size
self.hidden_size = feature_size
self.K = K
self.A = A
self.gconv = LocalizedSpectralGraphConvolution(A, K)
hidden_size = self.feature_size
input_size = self.feature_size + hidden_size
self.fl = nn.Linear(input_size, hidden_size)
self.il = nn.Linear(input_size, hidden_size)
self.ol = nn.Linear(input_size, hidden_size)
self.Cl = nn.Linear(input_size, hidden_size)
self.output_last = output_last
def step(self, input, Hidden_State, Cell_State):
# conv_sample_start = time.time()
conv = F.relu(self.gconv(input))
# conv_sample_end = time.time()
# print('conv_sample:', (conv_sample_end - conv_sample_start))
combined = torch.cat((conv, Hidden_State), 1)
f = F.sigmoid(self.fl(combined))
i = F.sigmoid(self.il(combined))
o = F.sigmoid(self.ol(combined))
C = F.tanh(self.Cl(combined))
Cell_State = f * Cell_State + i * C
Hidden_State = o * F.tanh(Cell_State)
return Hidden_State, Cell_State
def Bi_torch(self, a):
a[a < 0] = 0
a[a > 0] = 1
return a
def forward(self, inputs):
batch_size = inputs.size(0)
time_step = inputs.size(1)
Hidden_State, Cell_State = self.initHidden(batch_size)
outputs = None
for i in range(time_step):
Hidden_State, Cell_State = self.step(torch.squeeze(inputs[:,i:i+1,:]), Hidden_State, Cell_State)
if outputs is None:
outputs = Hidden_State.unsqueeze(1)
else:
outputs = torch.cat((outputs, Hidden_State.unsqueeze(1)), 1)
# print(type(outputs))
if self.output_last:
return outputs[:,-1,:]
else:
return outputs
def initHidden(self, batch_size):
use_gpu = torch.cuda.is_available()
if use_gpu:
Hidden_State = Variable(torch.zeros(batch_size, self.hidden_size).cuda())
Cell_State = Variable(torch.zeros(batch_size, self.hidden_size).cuda())
return Hidden_State, Cell_State
else:
Hidden_State = Variable(torch.zeros(batch_size, self.hidden_size))
Cell_State = Variable(torch.zeros(batch_size, self.hidden_size))
return Hidden_State, Cell_State
def reinitHidden(self, batch_size, Hidden_State_data, Cell_State_data):
use_gpu = torch.cuda.is_available()
if use_gpu:
Hidden_State = Variable(Hidden_State_data.cuda(), requires_grad=True)
Cell_State = Variable(Cell_State_data.cuda(), requires_grad=True)
return Hidden_State, Cell_State
else:
Hidden_State = Variable(Hidden_State_data, requires_grad=True)
Cell_State = Variable(Cell_State_data, requires_grad=True)
return Hidden_State, Cell_State
class SpectralGraphConvolution(nn.Module):
def __init__(self, A):
super(SpectralGraphConvolution, self).__init__()
feature_size = A.shape[0]
self.A = A
self.D = torch.diag(torch.sum(self.A, dim=0))
self.L = D - A
self.param = Parameter(torch.FloatTensor(feature_size).cuda())
stdv = 1. / math.sqrt(feature_size)
self.param.data.uniform_(-stdv, stdv)
self.e, self.v = torch.eig(L_, eigenvectors=True)
self.vt = torch.t(self.v)
self.v = Variable(self.v.cuda(), requires_grad=False)
self.vt = Variable(self.vt.cuda(), requires_grad=False)
def forward(self, input):
x = input
conv_sample_start = time.time()
conv = x.matmul(self.v.matmul(torch.diag(self.param)).matmul(self.vt))
conv_sample_end = time.time()
print('conv_sample:', (conv_sample_end - conv_sample_start))
return conv
class SpectralGraphConvolutionalLSTM(nn.Module):
def __init__(self, K, A, feature_size, Clamp_A=True, output_last = True):
'''
Args:
K: K-hop graph
A: adjacency matrix
FFR: free-flow reachability matrix
feature_size: the dimension of features
Clamp_A: Boolean value, clamping all elements of A between 0. to 1.
'''
super(SpectralGraphConvolutionalLSTM, self).__init__()
self.feature_size = feature_size
self.hidden_size = feature_size
self.K = K
self.A = A
self.gconv = SpectralGraphConvolution(A)
hidden_size = self.feature_size
input_size = self.feature_size + hidden_size
self.fl = nn.Linear(input_size, hidden_size)
self.il = nn.Linear(input_size, hidden_size)
self.ol = nn.Linear(input_size, hidden_size)
self.Cl = nn.Linear(input_size, hidden_size)
self.output_last = output_last
def step(self, input, Hidden_State, Cell_State):
conv_sample_start = time.time()
conv = self.gconv(input)
conv_sample_end = time.time()
print('conv_sample:', (conv_sample_end - conv_sample_start))
combined = torch.cat((conv, Hidden_State), 1)
f = F.sigmoid(self.fl(combined))
i = F.sigmoid(self.il(combined))
o = F.sigmoid(self.ol(combined))
C = F.tanh(self.Cl(combined))
Cell_State = f * Cell_State + i * C
Hidden_State = o * F.tanh(Cell_State)
return Hidden_State, Cell_State
def Bi_torch(self, a):
a[a < 0] = 0
a[a > 0] = 1
return a
def forward(self, inputs):
batch_size = inputs.size(0)
time_step = inputs.size(1)
Hidden_State, Cell_State = self.initHidden(batch_size)
outputs = None
train_sample_start = time.time()
for i in range(time_step):
Hidden_State, Cell_State = self.step(torch.squeeze(inputs[:,i:i+1,:]), Hidden_State, Cell_State)
if outputs is None:
outputs = Hidden_State.unsqueeze(1)
else:
outputs = torch.cat((outputs, Hidden_State.unsqueeze(1)), 1)
train_sample_end = time.time()
print('train sample:' , (train_sample_end - train_sample_start))
if self.output_last:
return outputs[:,-1,:]
else:
return outputs
def initHidden(self, batch_size):
use_gpu = torch.cuda.is_available()
if use_gpu:
Hidden_State = Variable(torch.zeros(batch_size, self.hidden_size).cuda())
Cell_State = Variable(torch.zeros(batch_size, self.hidden_size).cuda())
return Hidden_State, Cell_State
else:
Hidden_State = Variable(torch.zeros(batch_size, self.hidden_size))
Cell_State = Variable(torch.zeros(batch_size, self.hidden_size))
return Hidden_State, Cell_State
def reinitHidden(self, batch_size, Hidden_State_data, Cell_State_data):
use_gpu = torch.cuda.is_available()
if use_gpu:
Hidden_State = Variable(Hidden_State_data.cuda(), requires_grad=True)
Cell_State = Variable(Cell_State_data.cuda(), requires_grad=True)
return Hidden_State, Cell_State
else:
Hidden_State = Variable(Hidden_State_data, requires_grad=True)
Cell_State = Variable(Cell_State_data, requires_grad=True)
return Hidden_State, Cell_State
class GraphConvolutionalLSTM(nn.Module):
def __init__(self, K, A, FFR, feature_size, Clamp_A=True, output_last = True):
'''
Args:
K: K-hop graph
A: adjacency matrix
FFR: free-flow reachability matrix
feature_size: the dimension of features
Clamp_A: Boolean value, clamping all elements of A between 0. to 1.
'''
super(GraphConvolutionalLSTM, self).__init__()
self.feature_size = feature_size
self.hidden_size = feature_size
self.K = K
self.A_list = [] # Adjacency Matrix List
A = torch.FloatTensor(A)
A_temp = torch.eye(feature_size,feature_size)
for i in range(K):
A_temp = torch.matmul(A_temp, torch.Tensor(A))
if Clamp_A:
# confine elements of A
A_temp = torch.clamp(A_temp, max = 1.)
self.A_list.append(torch.mul(A_temp, torch.Tensor(FFR)))
# self.A_list.append(A_temp)
# a length adjustable Module List for hosting all graph convolutions
self.gc_list = nn.ModuleList([FilterLinear(feature_size, feature_size, self.A_list[i], bias=False) for i in range(K)])
hidden_size = self.feature_size
input_size = self.feature_size * K
self.fl = nn.Linear(input_size + hidden_size, hidden_size)
self.il = nn.Linear(input_size + hidden_size, hidden_size)
self.ol = nn.Linear(input_size + hidden_size, hidden_size)
self.Cl = nn.Linear(input_size + hidden_size, hidden_size)
# initialize the neighbor weight for the cell state
self.Neighbor_weight = Parameter(torch.FloatTensor(feature_size))
stdv = 1. / math.sqrt(feature_size)
self.Neighbor_weight.data.uniform_(-stdv, stdv)
self.output_last = output_last
def step(self, input, Hidden_State, Cell_State):
x = input
gc = self.gc_list[0](x)
for i in range(1, self.K):
gc = torch.cat((gc, self.gc_list[i](x)), 1)
combined = torch.cat((gc, Hidden_State), 1)
f = F.sigmoid(self.fl(combined))
i = F.sigmoid(self.il(combined))
o = F.sigmoid(self.ol(combined))
C = F.tanh(self.Cl(combined))
NC = torch.mul(Cell_State, torch.mv(Variable(self.A_list[-1], requires_grad=False).cuda(), self.Neighbor_weight))
Cell_State = f * NC + i * C
Hidden_State = o * F.tanh(Cell_State)
return Hidden_State, Cell_State, gc
def Bi_torch(self, a):
a[a < 0] = 0
a[a > 0] = 1
return a
def forward(self, inputs):
batch_size = inputs.size(0)
time_step = inputs.size(1)
Hidden_State, Cell_State = self.initHidden(batch_size)
outputs = None
for i in range(time_step):
Hidden_State, Cell_State, gc = self.step(torch.squeeze(inputs[:,i:i+1,:]), Hidden_State, Cell_State)
if outputs is None:
outputs = Hidden_State.unsqueeze(1)
else:
outputs = torch.cat((outputs, Hidden_State.unsqueeze(1)), 1)
if self.output_last:
return outputs[:,-1,:]
else:
return outputs
def initHidden(self, batch_size):
use_gpu = torch.cuda.is_available()
if use_gpu:
Hidden_State = Variable(torch.zeros(batch_size, self.hidden_size).cuda())
Cell_State = Variable(torch.zeros(batch_size, self.hidden_size).cuda())
return Hidden_State, Cell_State
else:
Hidden_State = Variable(torch.zeros(batch_size, self.hidden_size))
Cell_State = Variable(torch.zeros(batch_size, self.hidden_size))
return Hidden_State, Cell_State
def reinitHidden(self, batch_size, Hidden_State_data, Cell_State_data):
use_gpu = torch.cuda.is_available()
if use_gpu:
Hidden_State = Variable(Hidden_State_data.cuda(), requires_grad=True)
Cell_State = Variable(Cell_State_data.cuda(), requires_grad=True)
return Hidden_State, Cell_State
else:
Hidden_State = Variable(Hidden_State_data, requires_grad=True)
Cell_State = Variable(Cell_State_data, requires_grad=True)
return Hidden_State, Cell_State
lstm = LSTM(input_dim, hidden_dim, output_dim, output_last = True)
lstm, lstm_loss = TrainModel(lstm, train_dataloader, valid_dataloader, num_epochs = 1)
lstm_test = TestModel(lstm, test_dataloader, max_speed )
K = 64
Clamp_A = False
lsgclstm = LocalizedSpectralGraphConvolutionalLSTM(K, torch.Tensor(A), A.shape[0], Clamp_A=Clamp_A, output_last = True)
lsgclstm, lsgclstm_loss = TrainModel(lsgclstm, train_dataloader, valid_dataloader, num_epochs = 1)
lsgclstm_test = TestModel(lsgclstm, test_dataloader, max_speed )
K = 3
back_length = 3
Clamp_A = False
sgclstm = SpectralGraphConvolutionalLSTM(K, torch.Tensor(A), A.shape[0], Clamp_A=Clamp_A, output_last = True)
sgclstm, sgclstm_loss = TrainModel(sgclstm, train_dataloader, valid_dataloader, num_epochs = 1)
sgclstm_test = TestModel(sgclstm, test_dataloader, max_speed )
K = 3
back_length = 3
Clamp_A = False
gclstm = GraphConvolutionalLSTM(K, torch.Tensor(A), FFR[back_length], A.shape[0], Clamp_A=Clamp_A, output_last = True)
gclstm, gclstm_loss = TrainModel(gclstm, train_dataloader, valid_dataloader, num_epochs = 1)
gclstm_test = TestModel(gclstm, test_dataloader, max_speed )
rnn_val_loss = np.asarray(rnn_loss[3])
lstm_val_loss = np.asarray(lstm_loss[3])
hgclstm_val_loss = np.asarray(gclstm_loss[3])
lsgclstm_val_loss = np.asarray(lsgclstm_loss[3])
sgclstm_val_loss = np.asarray(sgclstm_loss[3])
lstm_val_loss = np.load('lstm_val_loss.npy')
hgclstm_val_loss = np.load('hgclstm_val_loss.npy')
lsgclstm_val_loss = np.load('lsgclstm_val_loss.npy')
sgclstm_val_loss = np.load('sgclstm_val_loss.npy')
# np.save('lstm_val_loss', lstm_val_loss)
# np.save('hgclstm_val_loss', gclstm_val_loss)
# np.save('lsgclstm_val_loss', lsgclstm_val_loss)
# np.save('sgclstm_val_loss', sgclstm_val_loss)
fig, ax = plt.subplots()
plt.plot(np.arange(1, len(lstm_val_loss) + 1), lstm_val_loss, label = 'LSTM')
plt.plot(np.arange(1, len(sgclstm_val_loss) + 1), sgclstm_val_loss, label = 'SGC+LSTM')
plt.plot(np.arange(1, len(lsgclstm_val_loss) + 1), lsgclstm_val_loss, label = 'LSGC+LSTM')
plt.plot(np.arange(1, len(hgclstm_val_loss) + 1),hgclstm_val_loss, label = 'HGC-LSTM')
plt.ylim((6 * 0.0001, 0.0019))
plt.xticks(fontsize = 12)
plt.yticks(fontsize = 12)
plt.yscale('log')
plt.ylabel('Validation Loss (MSE)', fontsize=12)
plt.xlabel('Epoch', fontsize=12)
# plt.gca().invert_xaxis()
plt.legend(fontsize=14)
plt.grid(True, which='both')
plt.savefig('Validation_loss.png', dpi=300, bbox_inches = 'tight', pad_inches=0.1)
```
| github_jupyter |
# Introduction to Programming with Python
# Unit 7: Recursion and the Beauty of Programming
As we are coming towards the end of our module, let's focus on more difficult topics, while also exploring some more beautiful fractal figures.
## Recursion
We have just discussed Fibonacci numbers, which are defined in the following way (if we number them starting from 0, as all programmers do):
$$
\begin{array}{l}
f_n = 1, 0\le n\le 1 \\
f_n = f_{n-1} + f_{n-2}, \quad n\ge2\\
\end{array}
$$
We have used some clever trick to compute them, having two variables:
```
def fib(n):
c,p = 1,1
for _ in range(n):
c,p = c+p,c
return p
fib(10)
```
However, we may also define the same function in another way, very similar to the mathematical definition of Fibonacci number given above:
```
def fib(n):
if n<=1:
return 1
else:
return fib(n-1) + fib(n-2)
fib(10)
```
You may be surprised: how is it possible to use the function `fib` inside its own definition? In face, such technique is called **recursion**, and it often used in programming.
Let us consider the example of calculating `fib(3)`. Recursion works as follows:
1. To calculate `fib(3)`, the function calls `fib(2)` and `fib(1)`
2. When `fib(2)` is called, it calls `fib(1)` and `fib(0)`.
3. Both of those calls `return 1` (because the argument `n` is less than 2)
4. Thus `fib(2)` is computed as `1+1=2`
5. When `fib(1)` is called in step 1, it also returns 1 (as in step 3)
6. `fib(3)` is computed to be equal to `2+1=3`.
We can actually add some `print` operators to the function to see how those calls are made:
```
def fib(n):
print(f'Calling fib({n})')
if n<=1:
print(f'fib({n}) returns 1')
return 1
else:
f1 = fib(n-1)
f2 = fib(n-2)
print(f'fib({n}) returns {f1+f2}={f1}+{f2}')
return f1+f2
fib(3)
```
You may notice that during those computations `fib(1)` is called twice. In fact, if you try to compute `fib` for any number larger than 20, you may notice significant delay - that is because number of calls to function `fib` increases very rapidly. For example, to compute `fib(16)` we would need to make aroung 65000 calls, while during our previous algorithm with a loop we need to repeat the loop only 16 times.
This does not necessarily mean that recursion is bad. Sometimes it is not easy to program an algorithm without recursion, as we will see in the next example. However, if you can think of a non-recursive way to solve the problem - you should prefer it, because it is likely to be more effective. However, recursive algorithms often tend to be shorter and more beautiful.
## Koch Snowflake
The best graphical example of recursion is a **[Koch Snowflake](https://en.wikipedia.org/wiki/Koch_snowflake)**, which looks like this:

Image [from Wikipedia](https://commons.wikimedia.org/w/index.php?curid=1898291), CC BY-SA 3.0
This picture shows snowflakes of different *complexities*:
* When complexity $n=0$ (first figure), to draw one side of the snowflake of size $x$ we just draw a straight line of length $x$
* When complexity $n=1$ (second figure), we draw the side in the following manner:
- Draw the side of length $x\over3$ and complexity $0$
- Turn $60^\circ$ left
- Draw the side of length $x\over3$ and complexity $0$
- Turn $120^\circ$ right
- Draw the side of length $x\over3$ and complexity $0$
- Turn $60^\circ$ left
* In a similar manner, for any other complexity $n>1$:
- Draw the side of length $x\over3$ and complexity $n-1$
- Turn $60^\circ$ left
- Draw the side of length $x\over3$ and complexity $n-1$
- Turn $120^\circ$ right
- Draw the side of length $x\over3$ and complexity $n-1$
- Turn $60^\circ$ left
You probably recognized *recursion* in this algorithm:
* For $n=0$ - we draw straight line. This is called **termination** of recursion, because we do not make a recursive call to the function
* For $n>0$ - we follow the algorithm described above, which involves calling the same function with argument $n-1$
The function to draw **Koch Curve** will look like this:
```
import jturtle as turtle
def koch_curve(n,x):
if n==0:
turtle.forward(x)
else:
koch_curve(n-1,x/3)
turtle.left(60)
koch_curve(n-1,x/3)
turtle.right(120)
koch_curve(n-1,x/3)
turtle.left(60)
koch_curve(n-1,x/3)
turtle.right(90)
koch_curve(2,100)
turtle.done()
```
Let's draw Koch curvers with different $n$-s:
```
turtle.right(90)
for n in range(4):
koch_curve(n,100)
turtle.penup()
turtle.forward(-100)
turtle.right(90)
turtle.forward(40)
turtle.left(90)
turtle.pendown()
turtle.done()
```
Finally, to produce Koch's Snowflake, we need to draw 3 Koch curves:
```
def koch_snowflake(n,x):
for _ in range(3):
koch_curve(n,x)
turtle.right(120)
koch_snowflake(3,100)
turtle.done()
```
## Minkowski Island
A very similar figure to Koch Snowflake is called **Minkowski Island**, or **Quadratic Koch Curve**. The idea is similar, but instead of turning 60 degrees - we turn 90.

By <a title="User:Prokofiev" href="//commons.wikimedia.org/wiki/User:Prokofiev">Prokofiev</a> - <span class="int-own-work" lang="en">Own work</span>, <a title="Creative Commons Attribution-Share Alike 4.0" href="https://creativecommons.org/licenses/by-sa/4.0">CC BY-SA 4.0</a>, <a href="https://commons.wikimedia.org/w/index.php?curid=3194494">Link</a>
While writing this function, we will try to be creative and use loop to draw each part of the curve. The trick is to use the **list** of all angles that we need to turn in between, together with their direction (i.e. 90 will mean turn $90^\circ$ left, 0 - do not turn at all, and -90 - turn $90^\circ$ right)
```
def minkowski(n,x):
if n==0:
turtle.forward(x)
else:
for t in [90,-90,-90,0,90,90,-90,0]:
minkowski(n-1,x/4)
turtle.left(t)
for _ in range(4):
minkowski(2,100)
turtle.left(90)
turtle.done()
```
## Flowers and Trees
If you continue studying Programming or Computer Science, you will learn that an important data structure is **tree** - a collection of elements that have parent and child elements. For example, folders or directories in our computer form a tree.
In our final example, we will draw something similar to a tree, and on the other hand similar to a dandelion or some umbrella-like plant. We will start with one branch, and then for each branch generate $n$ sub-branches, and so on.
```
from random import randint
def tree(n,b,x):
turtle.forward(x) # draw the branch
if n>0: # draw n sub-branches on top of it
for _ in range(b):
a = randint(-40,40)
turtle.left(a)
tree(n-1,b,x)
turtle.right(a)
turtle.forward(-x) # move back to the beginning of the branch
tree(3,3,100)
turtle.done()
```
This code is very clever, and it might take you some time to figure out. The difficult thing here is to make turtle return to the original position each time the branch is drawn. So, at the top level, the logic of the function is the following:
* Draw one branch by calling `forward(x)`
* Optionally (if $n>0$), draw $b$ sub-branches. Here $b$ is the **branching factor**.
* Move back to the starting position by calling `forward(-x)`.
When drawing branched, we always assume that `tree` moves the turtle back to its original position before the drawing.
```
for _ in range(5):
tree(3,3,100)
turtle.right(90)
turtle.forward(120)
turtle.left(90)
turtle.done()
```
We have cosidered just a few examples of drawing trees, but there could be many more. For example, try to experiment with our `tree` function to produce different types of flora by doing one or more of the following:
* Experiment with different branching factors
* The line `a = randint(-40,40)` defines random angle for each branch. By changing the angles you can control the spread of branches, and also whether the tree would look *straight*, or falling to the left or right
* Try to make branches shorter and longer as you go up towards the top of the tree. This can be achieved by multiplying `x` by some **growth factor**, which should be somewhat close to 1
* You may pass angle boundaries and growth factor as parameters to play with them freely. Finally, produce a forest that combines different kinds of trees together.
## The Beauty of Programming
I hope you would agree with me, that in the examples above we have managed to draw pretty beautiful pictures! They are not only pleasant aesthetically, but they are beautiful in *mathematical* sense. Koch Snowflake and Minkowski Island are examples of **fractal structures**, which can be infinitely complex, i.e. if you draw such a figure for very large $n$, you would be able to zoom into it, and part of the figure would be similar to the original one. Ideally, such a fractal would be infinitely complex. However, we managed to write a simple and rather short program, which can produce arbitrary complex figures. I believe this shows the main beauty of programming -- with simple code you can program computer to perform many complex operations and achieve very complex results!
## Where to go Next
Of course, real programs are more complex than those we have seen in this course, and you are still to learn how to fight complexity in programming. However, Python is a very good starting point, because from here you can start exploring programming in many different directions:
* **Web Programming** is about creating web sites. You can use Python and Django Framework to create web applications [see how](https://channel9.msdn.com/shows/Azure-Friday/Python-on-Azure-Part-1-Building-Django-apps-with-Visual-Studio-Code/?WT.mc_id=python-github-dmitryso)
* You can do **Game Development** with Python, a good place to start would be [PyGame](https://www.pygame.org) library. And you can also run Python on small programmable consoles, like [PyBadge](https://learn.adafruit.com/adafruit-pybadge)
<img src="./images/pybadge.jpg" width="30%"/><br/>
Picture from [this project at hackday.io](https://hackaday.io/project/164929/gallery#692b286060e43ce2e9ae50b5c3b1b0c5)
* **Data Science** is a branch of computer science which studies the ways you can extract knowledge from data. For example, having a large set of people's photographs and their corresponding age you can create a program that will **learn** how to determine person's age from a picture. This is closely related to another discipline, **Artificial Intelligence**. See [this example](http://aka.ms/facestudies) for how you can use information extracted from people's photographs to make interesting conclusions.
* **Science Art** is another direction at the intersection of science and art. Python is a great language for science art, because it contains powerful libraries for image manipulation. See [this blog post](http://aka.ms/peopleblending) on how you can create your own cognitive portrait from photographs like this one:
<img src="./images/VickieRotator.png" width="30%"/>
And there are many more areas where Pyton can be used as a language! However, before diving right into one of those specific areas, I suggest that you learn a little bit more of core language features in our next course.
I hope you enjoyed the course, and are curious to learn more! Good luck!
| github_jupyter |
# Data Extraction & Transformation
##### Parsing raw StatsBomb data and storing it in a Pandas DataFrame
---
```
import requests
import pandas as pd
from tqdm import tqdm
```
- `requests` is a great library for executing HTTP requests
- `pandas` is a data analysis and manipulation package
- `tqdm` is a clean progress bar library
---
```
base_url = "https://raw.githubusercontent.com/statsbomb/open-data/master/data/"
comp_url = base_url + "matches/{}/{}.json"
match_url = base_url + "events/{}.json"
```
These URLs are the locations where the raw StatsBomb data lives. Notice the `{}` in there, which are dynamically replaced with IDs with `.format()`
___
```
def parse_data(competition_id, season_id):
matches = requests.get(url=comp_url.format(competition_id, season_id)).json()
match_ids = [m['match_id'] for m in matches]
all_events = []
for match_id in tqdm(match_ids):
events = requests.get(url=match_url.format(match_id)).json()
shots = [x for x in events if x['type']['name'] == "Shot"]
for s in shots:
attributes = {
"match_id": match_id,
"team": s["possession_team"]["name"],
"player": s['player']['name'],
"x": s['location'][0],
"y": s['location'][1],
"outcome": s['shot']['outcome']['name'],
}
all_events.append(attributes)
return pd.DataFrame(all_events)
```
The `parse_data` function handles the full Extract & Transform process.
The sequence of events is this:
1. The list of matches is loaded into the `matches` list.
2. Match IDs are extracted into a separate list using a list comprehension on `matches`.
3. Iterate over Match ID's, and load each match's raw data into the `events` list.
4. Shots are extracted into a separate list using a list comprehension as a filter on `events`.
5. Iterate over shots and extract individual features and store them in the `attributes` dictionary.
6. Append each shot's `attributes` into the `all_events` list.
7. Return a Pandas DataFrame from the `all_events` list.
---
```
competition_id = 43
season_id = 3
```
- `competition_id = 43` - StatsBomb's Competition ID for the World Cup
- `season_id = 3` - StatsBomb's Season ID for the 2018 Season
```
df = parse_data(competition_id, season_id)
```
The `parse_data` function is executed, and it's output is placed in variable `df`
The progress bar is produced by `tqdm`
---
```
df.head(10)
```
The `.head(10)` method on a DataFrame object shows you the first 10 records in the DataFrame.
There are roughly `1700` shots in this DataFrame, which represent every shot attempted at the 2018 Men's World Cup.
---
Devin Pleuler 2020
| github_jupyter |
# NumPy Array Basics - Multi-dimensional Arrays
```
import sys
print(sys.version)
import numpy as np
print(np.__version__)
npa = np.arange(25)
npa
```
We can create these by reshaping arrays. One of the simplest ways is to just reshape an array with the reshape command. That gives us an x by x array.
```
npa.reshape((5,5))
```
We can also use the zeros commands.
```
npa2 = np.zeros((5,5))
npa2
```
To get the size of the array we can use the size method.
```
npa2.size
```
To get the shape of the array we can use the shape method.
```
npa2.shape
```
to get the number of dimension we use the ndim method.
```
npa2.ndim
```
We can create as many dimensions as we need to, here's 3 dimensions.
```
np.arange(8).reshape(2,2,2)
```
Here's 4 dimensions
```
np.zeros((4,4,4,4))
np.arange(16).reshape(2,2,2,2)
```
For the most part we’ll be working with 2 dimensions.
```
npa2
npa
```
Now we can really see the power of vectorization, let’s create two random 2 dimensional arrays.
Now I’m going to set the random seed. This basically makes your random number generation reproducible.
```
np.random.seed(10)
```
let’s try some random number generation and then we can perform some matrix comparisons.
```
npa2 = np.random.random_integers(1,10,25).reshape(5,5)
npa2
npa3 = np.random.random_integers(1,10,25).reshape(5,5)
npa3
```
We can do this comparison with greater than or equal to.
```
npa2 > npa3
```
We can also sum up the values where there are equal.
```
(npa2 == npa3).sum()
```
Or we can sum where one is greater than or equal to in the columns.
We can do that with sum or we could get the total by summing that array.
```
sum(npa2 >= npa3)
sum(npa2 >= npa3).sum()
```
We can also get the minimums and maximums like we got with single dimensional arrays or for specific dimensions.
```
npa2.min()
npa2.min(axis=1)
npa2.max(axis=0)
```
There are plenty of other functions that numpy as. we can transpose with .T property or transpose method.
```
npa2.T
npa2.transpose()
npa2.T == npa2.transpose()
```
We can also multiply this transposition by itself for example. This will be an item by item multiplication
```
npa2.T * npa2
```
We can flatten these arrays in several different ways.
we can flatten it, which returns a new array that we can change
```
np2 = npa2.flatten()
np2
```
or we can ravel it which ends up returning the original array in a flattened format.
```
r = npa2.ravel()
r
np2[0] = 25
npa2
```
With ravel if we change a value in the raveled array that will change it in the original n-dimensional array as well
```
r[0] = 25
npa2
```
Now we can use some other helpful functions like cumsum and comprod to get the cumulative products and sums. This works for any dimensional array.
```
npa2.cumsum()
npa2.cumprod()
```
That really covers a lot of the basic functions you’re going to use or need when working with pandas but it is worth being aware that numpy is a very deep library that does a lot more things that I've covered here. I wanted to cover these basics because they're going to come up when we're working with pandas. I'm sure this has felt fairly academic at this point but I can promise you that it provides a valuable foundation to pandas.
need. If there’s anything you have questions about feel free to ask along the side and I can create some appendix videos to help you along.
| github_jupyter |
<a href="https://colab.research.google.com/github/wisrovi/pyimagesearch-buy/blob/main/compare_histograms_opencv.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>

# How-To: 3 Ways to Compare Histograms using OpenCV and Python
### by [PyImageSearch.com](http://www.pyimagesearch.com)
## Welcome to **[PyImageSearch Plus](http://pyimg.co/plus)** Jupyter Notebooks!
This notebook is associated with the [How-To: 3 Ways to Compare Histograms using OpenCV and Python](https://www.pyimagesearch.com/2014/07/14/3-ways-compare-histograms-using-opencv-python/) blog post published on 2014-07-14.
Only the code for the blog post is here. Most codeblocks have a 1:1 relationship with what you find in the blog post with two exceptions: (1) Python classes are not separate files as they are typically organized with PyImageSearch projects, and (2) Command Line Argument parsing is replaced with an `args` dictionary that you can manipulate as needed.
We recommend that you execute (press ▶️) the code block-by-block, as-is, before adjusting parameters and `args` inputs. Once you've verified that the code is working, you are welcome to hack with it and learn from manipulating inputs, settings, and parameters. For more information on using Jupyter and Colab, please refer to these resources:
* [Jupyter Notebook User Interface](https://jupyter-notebook.readthedocs.io/en/stable/notebook.html#notebook-user-interface)
* [Overview of Google Colaboratory Features](https://colab.research.google.com/notebooks/basic_features_overview.ipynb)
As a reminder, these PyImageSearch Plus Jupyter Notebooks are not for sharing; please refer to the **Copyright** directly below and **Code License Agreement** in the last cell of this notebook.
Happy hacking!
*Adrian*
<hr>
***Copyright:*** *The contents of this Jupyter Notebook, unless otherwise indicated, are Copyright 2020 Adrian Rosebrock, PyimageSearch.com. All rights reserved. Content like this is made possible by the time invested by the authors. If you received this Jupyter Notebook and did not purchase it, please consider making future content possible by joining PyImageSearch Plus at http://pyimg.co/plus/ today.*
### Download the code zip file
```
!wget https://www.pyimagesearch.com/wp-content/uploads/2014/06/compare-histograms-opencv.zip
!unzip -qq compare-histograms-opencv.zip
%cd compare-histograms-opencv
```
## Blog Post Code
### Import Packages
```
# import the necessary packages
from scipy.spatial import distance as dist
import matplotlib.pyplot as plt
import numpy as np
import argparse
import glob
import cv2
```
### 3 Ways to Compare Histograms Using OpenCV and Python
```
# construct the argument parser and parse the arguments
#ap = argparse.ArgumentParser()
#ap.add_argument("-d", "--dataset", required = True,
# help = "Path to the directory of images")
#args = vars(ap.parse_args())
# since we are using Jupyter Notebooks we can replace our argument
# parsing code with *hard coded* arguments and values
args = {
"dataset": "images"
}
# initialize the index dictionary to store the image name
# and corresponding histograms and the images dictionary
# to store the images themselves
index = {}
images = {}
# loop over the image paths
for imagePath in glob.glob(args["dataset"] + "/*.png"):
# extract the image filename (assumed to be unique) and
# load the image, updating the images dictionary
filename = imagePath[imagePath.rfind("/") + 1:]
image = cv2.imread(imagePath)
images[filename] = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
# extract a 3D RGB color histogram from the image,
# using 8 bins per channel, normalize, and update
# the index
hist = cv2.calcHist([image], [0, 1, 2], None, [8, 8, 8],
[0, 256, 0, 256, 0, 256])
hist = cv2.normalize(hist, hist).flatten()
index[filename] = hist
```
#### Method 1: Using the OpenCV cv2.compareHist function
```
# METHOD #1: UTILIZING OPENCV
# initialize OpenCV methods for histogram comparison
OPENCV_METHODS = (
("Correlation", cv2.HISTCMP_CORREL),
("Chi-Squared", cv2.HISTCMP_CHISQR),
("Intersection", cv2.HISTCMP_INTERSECT),
("Hellinger", cv2.HISTCMP_BHATTACHARYYA))
# loop over the comparison methods
for (methodName, method) in OPENCV_METHODS:
# initialize the results dictionary and the sort
# direction
results = {}
reverse = False
# if we are using the correlation or intersection
# method, then sort the results in reverse order
if methodName in ("Correlation", "Intersection"):
reverse = True
# loop over the index
for (k, hist) in index.items():
# compute the distance between the two histograms
# using the method and update the results dictionary
d = cv2.compareHist(index["doge.png"], hist, method)
results[k] = d
# sort the results
results = sorted([(v, k) for (k, v) in results.items()], reverse = reverse)
# show the query image
fig = plt.figure("Query")
ax = fig.add_subplot(1, 1, 1)
ax.imshow(images["doge.png"])
plt.axis("off")
# initialize the results figure
fig = plt.figure("Results: %s" % (methodName), figsize=(10, 3))
fig.suptitle(methodName, fontsize = 20)
# loop over the results
for (i, (v, k)) in enumerate(results):
# show the result
ax = fig.add_subplot(1, len(images), i + 1)
ax.set_title("%s: %.2f" % (k, v))
plt.imshow(images[k])
plt.axis("off")
# show the OpenCV methods
plt.show()
```
#### Method 2: Using the SciPy distance metrics
```
# METHOD #2: UTILIZING SCIPY
# initialize the scipy methods to compaute distances
SCIPY_METHODS = (
("Euclidean", dist.euclidean),
("Manhattan", dist.cityblock),
("Chebysev", dist.chebyshev))
# loop over the comparison methods
for (methodName, method) in SCIPY_METHODS:
# initialize the dictionary dictionary
results = {}
# loop over the index
for (k, hist) in index.items():
# compute the distance between the two histograms
# using the method and update the results dictionary
d = method(index["doge.png"], hist)
results[k] = d
# sort the results
results = sorted([(v, k) for (k, v) in results.items()])
# show the query image
fig = plt.figure("Query")
ax = fig.add_subplot(1, 1, 1)
ax.imshow(images["doge.png"])
plt.axis("off")
# initialize the results figure
fig = plt.figure("Results: %s" % (methodName), figsize=(10, 3))
fig.suptitle(methodName, fontsize = 20)
# loop over the results
for (i, (v, k)) in enumerate(results):
# show the result
ax = fig.add_subplot(1, len(images), i + 1)
ax.set_title("%s: %.2f" % (k, v))
plt.imshow(images[k])
plt.axis("off")
# show the SciPy methods
plt.show()
```
#### Method 3: Roll-your-own similarity measure
```
# METHOD #3: ROLL YOUR OWN
def chi2_distance(histA, histB, eps = 1e-10):
# compute the chi-squared distance
d = 0.5 * np.sum([((a - b) ** 2) / (a + b + eps)
for (a, b) in zip(histA, histB)])
# return the chi-squared distance
return d
# initialize the results dictionary
results = {}
# loop over the index
for (k, hist) in index.items():
# compute the distance between the two histograms
# using the custom chi-squared method, then update
# the results dictionary
d = chi2_distance(index["doge.png"], hist)
results[k] = d
# sort the results
results = sorted([(v, k) for (k, v) in results.items()])
# show the query image
fig = plt.figure("Query")
ax = fig.add_subplot(1, 1, 1)
ax.imshow(images["doge.png"])
plt.axis("off")
# initialize the results figure
fig = plt.figure("Results: Custom Chi-Squared", figsize=(10, 3))
fig.suptitle("Custom Chi-Squared", fontsize = 20)
# loop over the results
for (i, (v, k)) in enumerate(results):
# show the result
ax = fig.add_subplot(1, len(images), i + 1)
ax.set_title("%s: %.2f" % (k, v))
plt.imshow(images[k])
plt.axis("off")
# show the custom method
plt.show()
```
For a detailed walkthrough of the concepts and code, be sure to refer to the full tutorial, [*How-To: 3 Ways to Compare Histograms using OpenCV and Python*](https://www.pyimagesearch.com/2014/07/14/3-ways-compare-histograms-using-opencv-python/) published on 2014-07-14.
# Code License Agreement
```
Copyright (c) 2020 PyImageSearch.com
SIMPLE VERSION
Feel free to use this code for your own projects, whether they are
purely educational, for fun, or for profit. THE EXCEPTION BEING if
you are developing a course, book, or other educational product.
Under *NO CIRCUMSTANCE* may you use this code for your own paid
educational or self-promotional ventures without written consent
from Adrian Rosebrock and PyImageSearch.com.
LONGER, FORMAL VERSION
Permission is hereby granted, free of charge, to any person obtaining
a copy of this software and associated documentation files
(the "Software"), to deal in the Software without restriction,
including without limitation the rights to use, copy, modify, merge,
publish, distribute, sublicense, and/or sell copies of the Software,
and to permit persons to whom the Software is furnished to do so,
subject to the following conditions:
The above copyright notice and this permission notice shall be
included in all copies or substantial portions of the Software.
Notwithstanding the foregoing, you may not use, copy, modify, merge,
publish, distribute, sublicense, create a derivative work, and/or
sell copies of the Software in any work that is designed, intended,
or marketed for pedagogical or instructional purposes related to
programming, coding, application development, or information
technology. Permission for such use, copying, modification, and
merger, publication, distribution, sub-licensing, creation of
derivative works, or sale is expressly withheld.
THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND,
EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES
OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND
NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS
BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN
ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN
CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE
SOFTWARE.
```
| github_jupyter |
# Surface Volume and CSV Processor
This code generates Surface Volume .csv files that are used in the Flood Estimate code, taking DEM .TIFF files divided by division and DEM files that have been grouped with the two .TIFF files to the south and north of them. The resulting .csv files contain the volume of water above the surface of the DEM at different heights in increments of 0.25m betwwen 0-3m and in increments of 0.5m from 3.5-6.5m. This processor also creates a folder called "Intermediate Data" which can be deleted after the code has completely run.
See https://pro.arcgis.com/en/pro-app/tool-reference/3d-analyst/surface-volume.htm for more information on how the arcpy Surface Volume function works.
## Imports and Initializations
```
import arcpy as ap
import scipy as sp
import numpy as np
import pandas as pd
import os
import csv
import glob
import re
```
Your DEM should be divided into "divisions" to conveniently model the propagation of water throughout your area of interest.
```
num_divs = 18 #however many divisions the DEM is divided into
```
# Ungrouped Divs
Inputs to these cells are the DEMs that have been cut into divisions.
## Surface Volume Processor
Process for 0-5m at 0.25 m intervals for each div.
```
ele_0_txt = "\\ele_0.txt"; ele_1_txt = "\\ele_1.txt"; ele_2_txt = "\\ele_2.txt"
ele_3_txt = "\\ele_3.txt"; ele_4_txt = "\\ele_4.txt"; ele_5_txt = "\\ele_5.txt"
ele_0_5_txt = "\\ele_0_5.txt"; ele_1_5_txt = "\\ele_1_5.txt"; ele_2_5_txt = "\\ele_2_5.txt"
ele_3_5_txt = "\\ele_3_5.txt"; ele_4_5_txt = "\\ele_4_5.txt"
ele_0_25_txt = "\\ele_0_25.txt"; ele_1_25_txt = "\\ele_1_25.txt"; ele_2_25_txt = "\\ele_2_25.txt"
ele_3_25_txt = "\\ele_3_25.txt"; ele_4_25_txt = "\\ele_4_25.txt"
ele_0_75_txt = "\\ele_0_75.txt"; ele_1_75_txt = "\\ele_1_75.txt"; ele_2_75_txt = "\\ele_2_75.txt"
ele_3_75_txt = "\\ele_3_75.txt"; ele_4_75_txt = "\\ele_4_75.txt"
for i in range(num_divs):
#this creates the destination folder for the txt files to go into if it hasn't been created yet
#you can comment this out if the folders have been created since it takes a while to run
divfolder = r"IntermediateData\newtext\div (%d)"%i
if not os.path.exists(divfolder):
os.makedirs(divfolder)
#this says what folder we want it to go into
ap.env.workspace = divfolder
#this says what DEM we want to use in the Surface Volume function.
#####These are your input DEM files################################
#####Change these to the ones relevant to your area of interest####
dem_lm = r"Data\LM_div18\dem_lm_z35_%d.TIF"%i
# Process: Surface Volume
# this performs Surface Volume on the DEM at the specified points.
ap.SurfaceVolume_3d(dem_lm, ele_0_txt, "BELOW", "0", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_1_txt, "BELOW", "1", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_2_txt, "BELOW", "2", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_3_txt, "BELOW", "3", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_4_txt, "BELOW", "4", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_5_txt, "BELOW", "5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_0_5_txt, "BELOW", "0.5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_1_5_txt, "BELOW", "1.5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_2_5_txt, "BELOW", "2.5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_3_5_txt, "BELOW", "3.5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_4_5_txt, "BELOW", "4.5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_0_25_txt, "BELOW", "0.25", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_1_25_txt, "BELOW", "1.25", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_2_25_txt, "BELOW", "2.25", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_3_25_txt, "BELOW", "3.25", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_4_25_txt, "BELOW", "4.25", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_0_75_txt, "BELOW", "0.75", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_1_75_txt, "BELOW", "1.75", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_2_75_txt, "BELOW", "2.75", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_3_75_txt, "BELOW", "3.75", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_4_75_txt, "BELOW", "4.75", "1", "0")
```
Process for 0-10m at 0.5m intervals
```
ele_0_txt = "\\ele_0.txt"; ele_2_txt = "\\ele_2.txt"; ele_4_txt = "\\ele_4.txt"
ele_6_txt = "\\ele_6.txt"; ele_8_txt = "\\ele_8.txt"; ele_10_txt = "\\ele_10.txt"
ele_1_txt = "\\ele_1.txt"; ele_3_txt = "\\ele_3.txt"; ele_5_txt = "\\ele_5.txt"
ele_7_txt = "\\ele_7.txt"; ele_9_txt = "\\ele_9.txt"
ele_0_5_txt = "\\ele_0_5.txt"; ele_2_5_txt = "\\ele_2_5.txt"; ele_4_5_txt = "\\ele_4_5.txt"
ele_6_5_txt = "\\ele_6_5.txt"; ele_8_5_txt = "\\ele_8_5.txt"
ele_1_5_txt = "\\ele_1_5.txt"; ele_3_5_txt = "\\ele_3_5.txt"; ele_5_5_txt = "\\ele_5_5.txt"
ele_7_5_txt = "\\ele_7_5.txt"; ele_9_5_txt = "\\ele_9_5.txt"
for i in range(num_divs):
#this creates the destination folder for the txt files to go into if it hasn't been created yet
#you can comment this out if the folders have been created since it takes a while to run
divfolder = r"IntermediateData\newtextlarge\div (%d)"%i
if not os.path.exists(divfolder):
os.makedirs(divfolder)
#this says what folder we want the txt files to go into
ap.env.workspace = divfolder
#this says what DEM we want to use in the Surface Volume function
#####These are your input DEM files################################
#####Change these to the ones relevant to your area of interest####
dem_lm = r"Data\LM_div18\dem_lm_z35_%d.TIF"%i
# Process: Surface Volume
# this performs Surface Volume on the DEM at the specified points.
ap.SurfaceVolume_3d(dem_lm, ele_0_txt, "BELOW", "0", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_2_txt, "BELOW", "2", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_4_txt, "BELOW", "4", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_6_txt, "BELOW", "6", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_8_txt, "BELOW", "8", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_10_txt, "BELOW", "10", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_1_txt, "BELOW", "1", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_3_txt, "BELOW", "3", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_5_txt, "BELOW", "5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_7_txt, "BELOW", "7", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_9_txt, "BELOW", "9", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_0_5_txt, "BELOW", "0.5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_2_5_txt, "BELOW", "2.5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_4_5_txt, "BELOW", "4.5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_6_5_txt, "BELOW", "6.5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_8_5_txt, "BELOW", "8.5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_1_5_txt, "BELOW", "1.5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_3_5_txt, "BELOW", "3.5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_5_5_txt, "BELOW", "5.5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_7_5_txt, "BELOW", "7.5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_9_5_txt, "BELOW", "9.5", "1", "0")
```
## TXT to CSV Maker
Creates csv files for each div for 0-5m at 0.25m intervals
```
surfacevolumefolder = r"IntermediateData\NewUngroupedSmallSurfaceVolume"
if not os.path.exists(surfacevolumefolder):
os.makedirs(surfacevolumefolder)
for div in range(num_divs):
l=[]
files = glob.glob(r'IntermediateData\newtext\div (%d)\*.txt'%div) #glob shoves all the text files into one container
for file in files:
f = open(file,'r')
for i,line in enumerate(f): #read through our list of text files as a series of strings
s = line.split(',') #split up the strings
result = re.findall(r"[-+]?\d*\.\d+|\d+", s[-1]) #grabs floating point numbers out of strings, which is always the last item in the string
l.append(result)
f.close()
x = [] #heights
y = [] #volumes
for i in range(len(l)): #the dummy list, l, has the values we want stored as every other item in the list
if(i%2==1): #so we need this mod 2 to get those indices
y.append(float(l[i][0]))
x.append(i/8 - .125) #this fixes the incrementation of the heights
y = sorted(y)
z = zip(x,y) #with the volumes sorted we can now tie all the values together with their respective reference heights
small_ungrouped_csv = r'IntermediateData\NewUngroupedSmallSurfaceVolume\LMN_div18_new_ungrouped_small_div{:02d}.csv'.format(div)
with open(small_ungrouped_csv,'w') as out: #naming the CSV you're trying to create
write = csv.writer(out)
write.writerow(['height','volume'])
for i in list(z):
write.writerow(i)
```
Creates csv files for each div from 0-10 m at 0.5m intervals
```
surfacevolumefolder = r"IntermediateData\NewUngroupedLargeSurfaceVolume"
if not os.path.exists(surfacevolumefolder):
os.makedirs(surfacevolumefolder)
for div in range(num_divs):
l=[]
files = glob.glob(r'IntermediateData\newtextlarge\div (%d)\*.txt'%div) #glob shoves all the text files into one container
for file in files:
f = open(file,'r')
for i,line in enumerate(f): #read through our list of text files as a series of strings
s = line.split(',') #split up the strings
result = re.findall(r"[-+]?\d*\.\d+|\d+", s[-1]) #grabs floating point numbers out of strings, which is always the last item in the string
l.append(result)
f.close()
x = [] #heights
y = [] #volumes
for i in range(len(l)): #the dummy list, l, has the values we want stored as every other item in the list
if(i%2==1): #so we need this mod 2 to get those indices
y.append(float(l[i][0]))
x.append(i/4 - .25) #this fixes the incrementation of the heights
y = sorted(y)
z = zip(x,y) #with the volumes sorted we can now tie all the values together with their respective reference heights
large_ungrouped_csv = r'IntermediateData\NewUngroupedLargeSurfaceVolume\LMN_div18_new_ungrouped_large_div_{:02d}.csv'.format(div)
with open(large_ungrouped_csv,'w') as out: #naming the CSV you're trying to create
write = csv.writer(out)
write.writerow(['height','volume'])
for i in list(z):
write.writerow(i)
```
## CSV Combiner
```
totalsurfacevolumefolder = r"Data\NewSurfaceVolumeCombined"
if not os.path.exists(totalsurfacevolumefolder):
os.makedirs(totalsurfacevolumefolder)
for div in range(num_divs): #does it for each div
combined_csv = []
#taking 0 to 3 in increments of 0.25
small_ungrouped_csv = r'IntermediateData\NewUngroupedSmallSurfaceVolume\LMN_div18_new_ungrouped_small_div{:02d}.csv'.format(div)
with open(small_ungrouped_csv, newline='') as File:
small_reader = csv.reader(File)
small_countrow = 0
for small_row in small_reader:
if small_countrow < 27 and small_row != []:
combined_csv.append(small_row)
small_countrow+=1
#taking 3.5 to 6.5 in increments of 0.5
large_ungrouped_csv = r'IntermediateData\NewUngroupedLargeSurfaceVolume\LMN_div18_new_ungrouped_large_div_{:02d}.csv'.format(div)
with open(large_ungrouped_csv, newline='') as File:
large_reader = csv.reader(File)
large_countrow = 0
for large_row in large_reader:
if large_countrow > 14 and large_countrow < 32 and large_row != []:
combined_csv.append(large_row)
large_countrow+=1
#combined csv is now a list with header and 20 values of height
#now we create a new csv
combined_ungrouped_csv = r'Data\NewSurfaceVolumeCombined\LMN_div18_new_{:02d}.csv'.format(div)
with open(combined_ungrouped_csv,'w') as out:
write = csv.writer(out)
for i in list(combined_csv):
write.writerow(i)
```
This produces 1 csv file for each of the divs in your area of interest. Now we can put NewSurfaceVolumeCombined into the Flood Estimate code!
# Grouped Divs
Inputs to these cells are DEMs that have been grouped using ArcGIS's Mosaic to Raster tool with the 2 divisions north of them and the 2 divisions south of them. The name of the DEM is the centermost division.
## Surface Volume Processor
Process for 0-5m at 0.25 m intervals for each div
```
ele_0_txt = "\\ele_0.txt"; ele_1_txt = "\\ele_1.txt"; ele_2_txt = "\\ele_2.txt"
ele_3_txt = "\\ele_3.txt"; ele_4_txt = "\\ele_4.txt"; ele_5_txt = "\\ele_5.txt"
ele_0_5_txt = "\\ele_0_5.txt"; ele_1_5_txt = "\\ele_1_5.txt"; ele_2_5_txt = "\\ele_2_5.txt"
ele_3_5_txt = "\\ele_3_5.txt"; ele_4_5_txt = "\\ele_4_5.txt"
ele_0_25_txt = "\\ele_0_25.txt"; ele_1_25_txt = "\\ele_1_25.txt"; ele_2_25_txt = "\\ele_2_25.txt"
ele_3_25_txt = "\\ele_3_25.txt"; ele_4_25_txt = "\\ele_4_25.txt"
ele_0_75_txt = "\\ele_0_75.txt"; ele_1_75_txt = "\\ele_1_75.txt"; ele_2_75_txt = "\\ele_2_75.txt"
ele_3_75_txt = "\\ele_3_75.txt"; ele_4_75_txt = "\\ele_4_75.txt"
for i in range(num_divs):
#this creates the destination folder for the txt files to go into if it hasn't been created yet
#you can comment this out if the folders have been created since it takes a while to run
divfolder = r"IntermediateData\newtextgrouped\div (%d)"%i
if not os.path.exists(divfolder):
os.makedirs(divfolder)
#this says what folder we want it to go into
ap.env.workspace = divfolder
#this says what DEM we want to use in the Surface Volume function
#####These are your input DEM files################################
#####Change these to the ones relevant to your area of interest####
dem_lm = r"Data\LM_div18_grouped\groupRaster_{:02d}.TIF".format(i)
# Process: Surface Volume
# this performs Surface Volume on the DEM at the specified points.
ap.SurfaceVolume_3d(dem_lm, ele_0_txt, "BELOW", "0", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_1_txt, "BELOW", "1", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_2_txt, "BELOW", "2", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_3_txt, "BELOW", "3", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_4_txt, "BELOW", "4", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_5_txt, "BELOW", "5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_0_5_txt, "BELOW", "0.5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_1_5_txt, "BELOW", "1.5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_2_5_txt, "BELOW", "2.5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_3_5_txt, "BELOW", "3.5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_4_5_txt, "BELOW", "4.5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_0_25_txt, "BELOW", "0.25", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_1_25_txt, "BELOW", "1.25", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_2_25_txt, "BELOW", "2.25", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_3_25_txt, "BELOW", "3.25", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_4_25_txt, "BELOW", "4.25", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_0_75_txt, "BELOW", "0.75", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_1_75_txt, "BELOW", "1.75", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_2_75_txt, "BELOW", "2.75", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_3_75_txt, "BELOW", "3.75", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_4_75_txt, "BELOW", "4.75", "1", "0")
```
Process for 0-10m at 0.5m intervals
```
ele_0_txt = "\\ele_0.txt"; ele_2_txt = "\\ele_2.txt"; ele_4_txt = "\\ele_4.txt"
ele_6_txt = "\\ele_6.txt"; ele_8_txt = "\\ele_8.txt"; ele_10_txt = "\\ele_10.txt"
ele_1_txt = "\\ele_1.txt"; ele_3_txt = "\\ele_3.txt"; ele_5_txt = "\\ele_5.txt"
ele_7_txt = "\\ele_7.txt"; ele_9_txt = "\\ele_9.txt"
ele_0_5_txt = "\\ele_0_5.txt"; ele_2_5_txt = "\\ele_2_5.txt"; ele_4_5_txt = "\\ele_4_5.txt"
ele_6_5_txt = "\\ele_6_5.txt"; ele_8_5_txt = "\\ele_8_5.txt"
ele_1_5_txt = "\\ele_1_5.txt"; ele_3_5_txt = "\\ele_3_5.txt"; ele_5_5_txt = "\\ele_5_5.txt"
ele_7_5_txt = "\\ele_7_5.txt"; ele_9_5_txt = "\\ele_9_5.txt"
for i in range(num_divs):
#this creates the destination folder for the txt files to go into if it hasn't been created yet
#you can comment this out if the folders have been created since it takes a while to run
divfolder = r"IntermediateData\newtextgroupedlarge\div (%d)"%i
if not os.path.exists(divfolder):
os.makedirs(divfolder)
#this says what folder we want the txt files to go into
ap.env.workspace = divfolder
#this says what DEM we want to use in the Surface Volume function
#####These are your input DEM files################################
#####Change these to the ones relevant to your area of interest####
dem_lm = r"Data\LM_div18_grouped\groupRaster_{:02d}.TIF".format(i)
# Process: Surface Volume
# this performs Surface Volume on the DEM at the specified points.
ap.SurfaceVolume_3d(dem_lm, ele_0_txt, "BELOW", "0", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_2_txt, "BELOW", "2", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_4_txt, "BELOW", "4", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_6_txt, "BELOW", "6", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_8_txt, "BELOW", "8", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_10_txt, "BELOW", "10", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_1_txt, "BELOW", "1", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_3_txt, "BELOW", "3", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_5_txt, "BELOW", "5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_7_txt, "BELOW", "7", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_9_txt, "BELOW", "9", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_0_5_txt, "BELOW", "0.5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_2_5_txt, "BELOW", "2.5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_4_5_txt, "BELOW", "4.5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_6_5_txt, "BELOW", "6.5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_8_5_txt, "BELOW", "8.5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_1_5_txt, "BELOW", "1.5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_3_5_txt, "BELOW", "3.5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_5_5_txt, "BELOW", "5.5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_7_5_txt, "BELOW", "7.5", "1", "0")
ap.SurfaceVolume_3d(dem_lm, ele_9_5_txt, "BELOW", "9.5", "1", "0")
```
## TXT to CSV Maker
Creates csv files for each div for 0-5m at 0.25m intervals
```
surfacevolumefolder = r"IntermediateData\NewGroupedSmallSurfaceVolume"
if not os.path.exists(surfacevolumefolder):
os.makedirs(surfacevolumefolder)
for div in range(num_divs):
l=[]
files = glob.glob(r'IntermediateData\newtextgrouped\div (%d)\*.txt'%div) #glob shoves all the text files into one container
for file in files:
f = open(file,'r')
for i,line in enumerate(f): #read through our list of text files as a series of strings
s = line.split(',') #split up the strings
result = re.findall(r"[-+]?\d*\.\d+|\d+", s[-1]) #grabs floating point numbers out of strings, which is always the last item in the string
l.append(result)
f.close()
x = [] #heights
y = [] #volumes
for i in range(len(l)): #the dummy list, l, has the values we want stored as every other item in the list
if(i%2==1): #so we need this mod 2 to get those indices
y.append(float(l[i][0]))
x.append(i/8 - .125) #this fixes the incrementation of the heights
y = sorted(y)
z = zip(x,y) #with the volumes sorted we can now tie all the values together with their respective reference heights
small_grouped_csv = r'IntermediateData\NewGroupedSmallSurfaceVolume\LMN_div18_new_grouped_small_div{:02d}.csv'.format(div)
with open(small_grouped_csv,'w') as out: #naming the CSV you're trying to create
write = csv.writer(out)
write.writerow(['height','volume'])
for i in list(z):
write.writerow(i)
```
Creates csv files for each div from 0-10 m at 0.5m intervals
```
surfacevolumefolder = r"IntermediateData\NewGroupedLargeSurfaceVolume"
if not os.path.exists(surfacevolumefolder):
os.makedirs(surfacevolumefolder)
for div in range(num_divs):
l=[]
files = glob.glob(r'IntermediateData\newtextgroupedlarge\div (%d)\*.txt'%div) #glob shoves all the text files into one container
for file in files:
f = open(file,'r')
for i,line in enumerate(f): #read through our list of text files as a series of strings
s = line.split(',') #split up the strings
result = re.findall(r"[-+]?\d*\.\d+|\d+", s[-1]) #grabs floating point numbers out of strings, which is always the last item in the string
l.append(result)
f.close()
x = [] #heights
y = [] #volumes
for i in range(len(l)): #the dummy list, l, has the values we want stored as every other item in the list
if(i%2==1): #so we need this mod 2 to get those indices
y.append(float(l[i][0]))
x.append(i/4 - .25) #this fixes the incrementation of the heights
y = sorted(y)
z = zip(x,y) #with the volumes sorted we can now tie all the values together with their respective reference heights
large_grouped_csv = r'IntermediateData\NewGroupedLargeSurfaceVolume\LMN_div18_new_grouped_large_div_{:02d}.csv'.format(div)
with open(large_grouped_csv,'w') as out: #naming the CSV you're trying to create
write = csv.writer(out)
write.writerow(['height','volume'])
for i in list(z):
write.writerow(i)
```
## CSV Combiner
```
totalsurfacevolumefolder = r"Data\NewSurfaceVolumeGrouped"
if not os.path.exists(totalsurfacevolumefolder):
os.makedirs(totalsurfacevolumefolder)
for div in range(num_divs): #does it for each div
combined_csv = []
#taking 0 to 3 in increments of 0.25
small_grouped_csv = r'IntermediateData\NewGroupedSmallSurfaceVolume\LMN_div18_new_grouped_small_div{:02d}.csv'.format(div)
with open(small_grouped_csv, newline='') as File:
small_reader = csv.reader(File)
small_countrow = 0
for small_row in small_reader:
if small_countrow < 27 and small_row != []:
combined_csv.append(small_row)
small_countrow+=1
#taking 3.5 to 6.5 in increments of 0.5
large_grouped_csv = r'IntermediateData\NewGroupedLargeSurfaceVolume\LMN_div18_new_grouped_large_div_{:02d}.csv'.format(div)
with open(large_grouped_csv, newline='') as File:
large_reader = csv.reader(File)
large_countrow = 0
for large_row in large_reader:
if large_countrow > 14 and large_countrow < 32 and large_row != []:
combined_csv.append(large_row)
large_countrow+=1
#combined csv is now a list with header and 20 values of height
#now we create a new csv
combined_grouped_csv = r'Data\NewSurfaceVolumeGrouped\LMN_div18_new_grouped_{:02d}.csv'.format(div)
with open(combined_grouped_csv,'w') as out:
write = csv.writer(out)
for i in list(combined_csv):
write.writerow(i)
```
This produces 1 "grouped" csv file for each of the divs in your area of interest. Now we can put NewSurfaceVolumeGrouped into the Flood Estimate code!
| github_jupyter |
# Convolutional Neural Networks in TensorFlow
```
%tensorflow_version 1.x
```
## Dataset
Here we're importing the MNIST dataset and using a convenient TensorFlow function to batch, scale, and one-hot encode the data.
```
from examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets(".", one_hot=True, reshape=False)
import tensorflow as tf
#parameters
learning_rate = 0.00001
epochs = 10
batch_size = 128
# number of samples to calculate validation and accuracy
test_valid_size = 256 # if you run out of memory decrease this
# network parameters
n_classes = 10 # MNIST total classes
dropout = 0.75 # probability to keep units
```
## Weights and Biases
We will create 3 layers alternating between convolutions and max pooling followed by a fully connected and output layer. We begin by defining the necessary weights and biases
```
weights = {
"wc1": tf.Variable(tf.random_normal([5,5,1,32])),
"wc2": tf.Variable(tf.random_normal([5,5,32,64])),
"wd1": tf.Variable(tf.random_normal([7*7*64,1024])),
"out": tf.Variable(tf.random_normal([1024,n_classes]))
}
biases = {
"bc1": tf.Variable(tf.random_normal([32])),
"bc2": tf.Variable(tf.random_normal([64])),
"bd1": tf.Variable(tf.random_normal([1024])),
"out": tf.Variable(tf.random_normal([n_classes]))
}
```
## Convolution Layers
The conv2d function computes the convolution against weight W, and then adds bias b, before applying a ReLU activation function
```
def conv2d(x,W,b,strides=1):
x = tf.nn.conv2d(x,W,strides=[1,strides,strides, 1],padding="SAME")
x = tf.nn.bias_add(x,b)
return tf.nn.relu(x)
```
## Max Pooling Layers
The maxpool2d function applies max pooling to layer x using a filter of size k.
```
def maxpool2d(x,k=2):
return tf.nn.max_pool(x,ksize=[1,k,k,1],strides=[1,k,k,1],padding="SAME")
```
## Model
The transformation of each layer to new dimensions is shown in the comments. For example, the first layer shapes the images from 28x28x1 to 28x28x32 in the convolution step
The next step applies max pooling, turning each sample into 14x14x32
All the layers are applied from conv1 to output, producing 10 class predictions.
```
def conv_net(x,weights,biases,dropout):
# Layer 1 - 28*28*1 to 14*14*32
conv1 = conv2d(x,weights["wc1"],biases["bc1"])
conv1 = maxpool2d(conv1,k=2)
# Layer 2 - 14*14*32 to 7*7*64
conv2 = conv2d(conv1,weights["wc2"],biases["bc2"])
conv2 = maxpool2d(conv2,k=2)
# Fully connected layer - 7*7*64 to 1024
fc1 = tf.reshape(conv2,[-1,weights["wd1"].get_shape().as_list()[0]])
fc1 = tf.add(tf.matmul(fc1,weights["wd1"]),biases["bd1"])
fc1 = tf.nn.relu(fc1)
fc1 = tf.nn.dropout(fc1,dropout)
# Output Layer - class prediction - 1024 to 10
out = tf.add(tf.matmul(fc1,weights["out"]),biases["out"])
return out
```
## Session
Running phase
### Graph Input
```
x = tf.placeholder(tf.float32,[None,28,28,1])
y = tf.placeholder(tf.float32,[None,n_classes])
keep_prob = tf.placeholder(tf.float32)
```
### Model
```
logits = conv_net(x,weights,biases,keep_prob)
```
### Define loss and optimizer
```
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits_v2(logits=logits,labels=y))
optimizer = tf.train.GradientDescentOptimizer(learning_rate=learning_rate).minimize(cost)
```
### Accuracy
```
correct_pred = tf.equal(tf.argmax(logits,1),tf.argmax(y,1))
accuracy = tf.reduce_mean(tf.cast(correct_pred,tf.float32))
```
### Initializing the variable
```
init = tf.global_variables_initializer()
```
# Launching it
```
with tf.Session() as sess:
sess.run(init)
for epoch in range(epochs):
for batch in range(mnist.train.num_examples//batch_size):
batch_x,batch_y = mnist.train.next_batch(batch_size)
sess.run(optimizer,{x:batch_x,y:batch_y,keep_prob:dropout})
# Calculate batch loss
loss = sess.run(cost,{x:batch_x,y:batch_y,keep_prob:1.})
# Calculate batch accuracy
valid_accuracy = sess.run(accuracy,{x:mnist.validation.images[:test_valid_size],y:mnist.validation.labels[:test_valid_size],keep_prob:1.})
print('Epoch {:>2}, Batch {:>3} - Loss: {:>10.4f} Validation Accuracy: {:.6f}'.format(epoch + 1,batch + 1,loss,valid_accuracy))
# Calculate test accuracy
test_accuracy = sess.run(accuracy,{x: mnist.test.images[:test_valid_size],y: mnist.test.labels[:test_valid_size],keep_prob: 1.})
print('Testing Accuracy: {}'.format(test_accuracy))
```
| github_jupyter |
# Programming Exercise 1: Linear Regression
## Introduction
In this exercise, you will implement linear regression and get to see it work on data. Before starting on this programming exercise, we strongly recommend watching the video lectures and completing the review questions for the associated topics.
All the information you need for solving this assignment is in this notebook, and all the code you will be implementing will take place within this notebook. The assignment can be promptly submitted to the coursera grader directly from this notebook (code and instructions are included below).
Before we begin with the exercises, we need to import all libraries required for this programming exercise. Throughout the course, we will be using [`numpy`](http://www.numpy.org/) for all arrays and matrix operations, and [`matplotlib`](https://matplotlib.org/) for plotting.
You can find instructions on how to install required libraries in the README file in the [github repository](https://github.com/dibgerge/ml-coursera-python-assignments).
```
# used for manipulating directory paths
import os
# Scientific and vector computation for python
import numpy as np
# Plotting library
from matplotlib import pyplot
from mpl_toolkits.mplot3d import Axes3D # needed to plot 3-D surfaces
# library written for this exercise providing additional functions for assignment submission, and others
import utils
# define the submission/grader object for this exercise
grader = utils.Grader()
# tells matplotlib to embed plots within the notebook
%matplotlib inline
```
## Submission and Grading
After completing each part of the assignment, be sure to submit your solutions to the grader.
For this programming exercise, you are only required to complete the first part of the exercise to implement linear regression with one variable. The second part of the exercise, which is optional, covers linear regression with multiple variables. The following is a breakdown of how each part of this exercise is scored.
**Required Exercises**
| Section | Part |Submitted Function | Points
|---------|:- |:- | :-:
| 1 | [Warm up exercise](#section1) | [`warmUpExercise`](#warmUpExercise) | 10
| 2 | [Compute cost for one variable](#section2) | [`computeCost`](#computeCost) | 40
| 3 | [Gradient descent for one variable](#section3) | [`gradientDescent`](#gradientDescent) | 50
| | Total Points | | 100
**Optional Exercises**
| Section | Part | Submitted Function | Points |
|:-------:|:- |:-: | :-: |
| 4 | [Feature normalization](#section4) | [`featureNormalize`](#featureNormalize) | 0 |
| 5 | [Compute cost for multiple variables](#section5) | [`computeCostMulti`](#computeCostMulti) | 0 |
| 6 | [Gradient descent for multiple variables](#section5) | [`gradientDescentMulti`](#gradientDescentMulti) |0 |
| 7 | [Normal Equations](#section7) | [`normalEqn`](#normalEqn) | 0 |
You are allowed to submit your solutions multiple times, and we will take only the highest score into consideration.
<div class="alert alert-block alert-warning">
At the end of each section in this notebook, we have a cell which contains code for submitting the solutions thus far to the grader. Execute the cell to see your score up to the current section. For all your work to be submitted properly, you must execute those cells at least once. They must also be re-executed everytime the submitted function is updated.
</div>
## Debugging
Here are some things to keep in mind throughout this exercise:
- Python array indices start from zero, not one (contrary to OCTAVE/MATLAB).
- There is an important distinction between python arrays (called `list` or `tuple`) and `numpy` arrays. You should use `numpy` arrays in all your computations. Vector/matrix operations work only with `numpy` arrays. Python lists do not support vector operations (you need to use for loops).
- If you are seeing many errors at runtime, inspect your matrix operations to make sure that you are adding and multiplying matrices of compatible dimensions. Printing the dimensions of `numpy` arrays using the `shape` property will help you debug.
- By default, `numpy` interprets math operators to be element-wise operators. If you want to do matrix multiplication, you need to use the `dot` function in `numpy`. For, example if `A` and `B` are two `numpy` matrices, then the matrix operation AB is `np.dot(A, B)`. Note that for 2-dimensional matrices or vectors (1-dimensional), this is also equivalent to `A@B` (requires python >= 3.5).
<a id="section1"></a>
## 1 Simple python and `numpy` function
The first part of this assignment gives you practice with python and `numpy` syntax and the homework submission process. In the next cell, you will find the outline of a `python` function. Modify it to return a 5 x 5 identity matrix by filling in the following code:
```python
A = np.eye(5)
```
<a id="warmUpExercise"></a>
```
def warmUpExercise():
"""
Example function in Python which computes the identity matrix.
Returns
-------
A : array_like
The 5x5 identity matrix.
Instructions
------------
Return the 5x5 identity matrix.
"""
# ======== YOUR CODE HERE ======
A = np.eye(5) # modify this line
# ==============================
return A
```
The previous cell only defines the function `warmUpExercise`. We can now run it by executing the following cell to see its output. You should see output similar to the following:
```python
array([[ 1., 0., 0., 0., 0.],
[ 0., 1., 0., 0., 0.],
[ 0., 0., 1., 0., 0.],
[ 0., 0., 0., 1., 0.],
[ 0., 0., 0., 0., 1.]])
```
```
warmUpExercise()
```
### 1.1 Submitting solutions
After completing a part of the exercise, you can submit your solutions for grading by first adding the function you modified to the grader object, and then sending your function to Coursera for grading.
The grader will prompt you for your login e-mail and submission token. You can obtain a submission token from the web page for the assignment. You are allowed to submit your solutions multiple times, and we will take only the highest score into consideration.
Execute the next cell to grade your solution to the first part of this exercise.
*You should now submit you solutions.*
```
# appends the implemented function in part 1 to the grader object
grader[1] = warmUpExercise
# send the added functions to coursera grader for getting a grade on this part
grader.grade()
```
## 2 Linear regression with one variable
Now you will implement linear regression with one variable to predict profits for a food truck. Suppose you are the CEO of a restaurant franchise and are considering different cities for opening a new outlet. The chain already has trucks in various cities and you have data for profits and populations from the cities. You would like to use this data to help you select which city to expand to next.
The file `Data/ex1data1.txt` contains the dataset for our linear regression problem. The first column is the population of a city (in 10,000s) and the second column is the profit of a food truck in that city (in $10,000s). A negative value for profit indicates a loss.
We provide you with the code needed to load this data. The dataset is loaded from the data file into the variables `x` and `y`:
```
# Read comma separated data
data = np.loadtxt(os.path.join('Data', 'ex1data1.txt'), delimiter=',')
X, y = data[:, 0], data[:, 1]
m = y.size # number of training examples
```
### 2.1 Plotting the Data
Before starting on any task, it is often useful to understand the data by visualizing it. For this dataset, you can use a scatter plot to visualize the data, since it has only two properties to plot (profit and population). Many other problems that you will encounter in real life are multi-dimensional and cannot be plotted on a 2-d plot. There are many plotting libraries in python (see this [blog post](https://blog.modeanalytics.com/python-data-visualization-libraries/) for a good summary of the most popular ones).
In this course, we will be exclusively using `matplotlib` to do all our plotting. `matplotlib` is one of the most popular scientific plotting libraries in python and has extensive tools and functions to make beautiful plots. `pyplot` is a module within `matplotlib` which provides a simplified interface to `matplotlib`'s most common plotting tasks, mimicking MATLAB's plotting interface.
<div class="alert alert-block alert-warning">
You might have noticed that we have imported the `pyplot` module at the beginning of this exercise using the command `from matplotlib import pyplot`. This is rather uncommon, and if you look at python code elsewhere or in the `matplotlib` tutorials, you will see that the module is named `plt`. This is used by module renaming by using the import command `import matplotlib.pyplot as plt`. We will not using the short name of `pyplot` module in this class exercises, but you should be aware of this deviation from norm.
</div>
In the following part, your first job is to complete the `plotData` function below. Modify the function and fill in the following code:
```python
pyplot.plot(x, y, 'ro', ms=10, mec='k')
pyplot.ylabel('Profit in $10,000')
pyplot.xlabel('Population of City in 10,000s')
```
```
def plotData(x, y):
"""
Plots the data points x and y into a new figure. Plots the data
points and gives the figure axes labels of population and profit.
Parameters
----------
x : array_like
Data point values for x-axis.
y : array_like
Data point values for y-axis. Note x and y should have the same size.
Instructions
------------
Plot the training data into a figure using the "figure" and "plot"
functions. Set the axes labels using the "xlabel" and "ylabel" functions.
Assume the population and revenue data have been passed in as the x
and y arguments of this function.
Hint
----
You can use the 'ro' option with plot to have the markers
appear as red circles. Furthermore, you can make the markers larger by
using plot(..., 'ro', ms=10), where `ms` refers to marker size. You
can also set the marker edge color using the `mec` property.
"""
fig = pyplot.figure() # open a new figure
# ====================== YOUR CODE HERE =======================
pyplot.plot(x, y, 'ro', ms=10, mec='k')
pyplot.ylabel('Profit in $10,000')
pyplot.xlabel('Population of City in 10,000s')
# =============================================================
```
Now run the defined function with the loaded data to visualize the data. The end result should look like the following figure:

Execute the next cell to visualize the data.
```
plotData(X, y)
```
To quickly learn more about the `matplotlib` plot function and what arguments you can provide to it, you can type `?pyplot.plot` in a cell within the jupyter notebook. This opens a separate page showing the documentation for the requested function. You can also search online for plotting documentation.
To set the markers to red circles, we used the option `'or'` within the `plot` function.
```
?pyplot.plot
```
<a id="section2"></a>
### 2.2 Gradient Descent
In this part, you will fit the linear regression parameters $\theta$ to our dataset using gradient descent.
#### 2.2.1 Update Equations
The objective of linear regression is to minimize the cost function
$$ J(\theta) = \frac{1}{2m} \sum_{i=1}^m \left( h_{\theta}(x^{(i)}) - y^{(i)}\right)^2$$
where the hypothesis $h_\theta(x)$ is given by the linear model
$$ h_\theta(x) = \theta^Tx = \theta_0 + \theta_1 x_1$$
Recall that the parameters of your model are the $\theta_j$ values. These are
the values you will adjust to minimize cost $J(\theta)$. One way to do this is to
use the batch gradient descent algorithm. In batch gradient descent, each
iteration performs the update
$$ \theta_j = \theta_j - \alpha \frac{1}{m} \sum_{i=1}^m \left( h_\theta(x^{(i)}) - y^{(i)}\right)x_j^{(i)} \qquad \text{simultaneously update } \theta_j \text{ for all } j$$
With each step of gradient descent, your parameters $\theta_j$ come closer to the optimal values that will achieve the lowest cost J($\theta$).
<div class="alert alert-block alert-warning">
**Implementation Note:** We store each example as a row in the the $X$ matrix in Python `numpy`. To take into account the intercept term ($\theta_0$), we add an additional first column to $X$ and set it to all ones. This allows us to treat $\theta_0$ as simply another 'feature'.
</div>
#### 2.2.2 Implementation
We have already set up the data for linear regression. In the following cell, we add another dimension to our data to accommodate the $\theta_0$ intercept term. Do NOT execute this cell more than once.
```
# Add a column of ones to X. The numpy function stack joins arrays along a given axis.
# The first axis (axis=0) refers to rows (training examples)
# and second axis (axis=1) refers to columns (features).
X = np.stack([np.ones(m), X], axis=1)
```
<a id="section2"></a>
#### 2.2.3 Computing the cost $J(\theta)$
As you perform gradient descent to learn minimize the cost function $J(\theta)$, it is helpful to monitor the convergence by computing the cost. In this section, you will implement a function to calculate $J(\theta)$ so you can check the convergence of your gradient descent implementation.
Your next task is to complete the code for the function `computeCost` which computes $J(\theta)$. As you are doing this, remember that the variables $X$ and $y$ are not scalar values. $X$ is a matrix whose rows represent the examples from the training set and $y$ is a vector whose each elemennt represent the value at a given row of $X$.
<a id="computeCost"></a>
```
def computeCost(X, y, theta):
"""
Compute cost for linear regression. Computes the cost of using theta as the
parameter for linear regression to fit the data points in X and y.
Parameters
----------
X : array_like
The input dataset of shape (m x n+1), where m is the number of examples,
and n is the number of features. We assume a vector of one's already
appended to the features so we have n+1 columns.
y : array_like
The values of the function at each data point. This is a vector of
shape (m, ).
theta : array_like
The parameters for the regression function. This is a vector of
shape (n+1, ).
Returns
-------
J : float
The value of the regression cost function.
Instructions
------------
Compute the cost of a particular choice of theta.
You should set J to the cost.
"""
# initialize some useful values
m = y.size # number of training examples
# You need to return the following variables correctly
J = 0
# ====================== YOUR CODE HERE =====================
h = np.sum(theta*X, axis=1)
temp = (h-y)**2
J = np.sum(temp) * 0.5 / m
# ===========================================================
return J
computeCost(X, y, theta=np.array([0.0, 1.0]))
J = computeCost(X, y, theta=np.array([0.0, 0.0]))
print('With theta = [0, 0] \nCost computed = %.2f' % J)
print('Expected cost value (approximately) 32.07\n')
# further testing of the cost function
J = computeCost(X, y, theta=np.array([-1, 2]))
print('With theta = [-1, 2]\nCost computed = %.2f' % J)
print('Expected cost value (approximately) 54.24')
```
Once you have completed the function, the next step will run `computeCost` two times using two different initializations of $\theta$. You will see the cost printed to the screen.
*You should now submit your solutions by executing the following cell.*
```
grader[2] = computeCost
grader.grade()
```
<a id="section3"></a>
#### 2.2.4 Gradient descent
Next, you will complete a function which implements gradient descent.
The loop structure has been written for you, and you only need to supply the updates to $\theta$ within each iteration.
As you program, make sure you understand what you are trying to optimize and what is being updated. Keep in mind that the cost $J(\theta)$ is parameterized by the vector $\theta$, not $X$ and $y$. That is, we minimize the value of $J(\theta)$ by changing the values of the vector $\theta$, not by changing $X$ or $y$. [Refer to the equations in this notebook](#section2) and to the video lectures if you are uncertain. A good way to verify that gradient descent is working correctly is to look at the value of $J(\theta)$ and check that it is decreasing with each step.
The starter code for the function `gradientDescent` calls `computeCost` on every iteration and saves the cost to a `python` list. Assuming you have implemented gradient descent and `computeCost` correctly, your value of $J(\theta)$ should never increase, and should converge to a steady value by the end of the algorithm.
<div class="alert alert-box alert-warning">
**Vectors and matrices in `numpy`** - Important implementation notes
A vector in `numpy` is a one dimensional array, for example `np.array([1, 2, 3])` is a vector. A matrix in `numpy` is a two dimensional array, for example `np.array([[1, 2, 3], [4, 5, 6]])`. However, the following is still considered a matrix `np.array([[1, 2, 3]])` since it has two dimensions, even if it has a shape of 1x3 (which looks like a vector).
Given the above, the function `np.dot` which we will use for all matrix/vector multiplication has the following properties:
- It always performs inner products on vectors. If `x=np.array([1, 2, 3])`, then `np.dot(x, x)` is a scalar.
- For matrix-vector multiplication, so if $X$ is a $m\times n$ matrix and $y$ is a vector of length $m$, then the operation `np.dot(y, X)` considers $y$ as a $1 \times m$ vector. On the other hand, if $y$ is a vector of length $n$, then the operation `np.dot(X, y)` considers $y$ as a $n \times 1$ vector.
- A vector can be promoted to a matrix using `y[None]` or `[y[np.newaxis]`. That is, if `y = np.array([1, 2, 3])` is a vector of size 3, then `y[None, :]` is a matrix of shape $1 \times 3$. We can use `y[:, None]` to obtain a shape of $3 \times 1$.
<div>
<a id="gradientDescent"></a>
```
def gradientDescent(X, y, theta, alpha, num_iters):
"""
Performs gradient descent to learn `theta`. Updates theta by taking `num_iters`
gradient steps with learning rate `alpha`.
Parameters
----------
X : array_like
The input dataset of shape (m x n+1).
y : arra_like
Value at given features. A vector of shape (m, ).
theta : array_like
Initial values for the linear regression parameters.
A vector of shape (n+1, ).
alpha : float
The learning rate.
num_iters : int
The number of iterations for gradient descent.
Returns
-------
theta : array_like
The learned linear regression parameters. A vector of shape (n+1, ).
J_history : list
A python list for the values of the cost function after each iteration.
Instructions
------------
Peform a single gradient step on the parameter vector theta.
While debugging, it can be useful to print out the values of
the cost function (computeCost) and gradient here.
"""
# Initialize some useful values
m = y.shape[0] # number of training examples
# make a copy of theta, to avoid changing the original array, since numpy arrays
# are passed by reference to functions
theta = theta.copy()
J_history = [] # Use a python list to save cost in every iteration
for i in range(num_iters):
# ==================== YOUR CODE HERE =================================
#theta = theta - (h-y)*(X[])
h = np.dot(X, theta) #cross prodct for hypothesis
grad = np.dot(X.transpose(), (h-y)) / m # partial derivatives
theta = theta - alpha * grad # perform gradient descent
# =====================================================================
# save the cost J in every iteration
J_history.append(computeCost(X, y, theta))
return theta, J_history
```
After you are finished call the implemented `gradientDescent` function and print the computed $\theta$. We initialize the $\theta$ parameters to 0 and the learning rate $\alpha$ to 0.01. Execute the following cell to check your code.
```
# initialize fitting parameters
theta = np.zeros(2)
# some gradient descent settings
iterations = 1500
alpha = 0.01
theta, J_history = gradientDescent(X ,y, theta, alpha, iterations)
print('Theta found by gradient descent: {:.4f}, {:.4f}'.format(*theta))
print('Expected theta values (approximately): [-3.6303, 1.1664]')
```
We will use your final parameters to plot the linear fit. The results should look like the following figure.

```
# plot the linear fit
plotData(X[:, 1], y)
pyplot.plot(X[:, 1], np.dot(X, theta), '-')
pyplot.legend(['Training data', 'Linear regression']);
```
Your final values for $\theta$ will also be used to make predictions on profits in areas of 35,000 and 70,000 people.
<div class="alert alert-block alert-success">
Note the way that the following lines use matrix multiplication, rather than explicit summation or looping, to calculate the predictions. This is an example of code vectorization in `numpy`.
</div>
<div class="alert alert-block alert-success">
Note that the first argument to the `numpy` function `dot` is a python list. `numpy` can internally converts **valid** python lists to numpy arrays when explicitly provided as arguments to `numpy` functions.
</div>
```
# Predict values for population sizes of 35,000 and 70,000
predict1 = np.dot([1, 3.5], theta)
print('For population = 35,000, we predict a profit of {:.2f}\n'.format(predict1*10000))
predict2 = np.dot([1, 7], theta)
print('For population = 70,000, we predict a profit of {:.2f}\n'.format(predict2*10000))
```
*You should now submit your solutions by executing the next cell.*
```
grader[3] = gradientDescent
grader.grade()
```
### 2.4 Visualizing $J(\theta)$
To understand the cost function $J(\theta)$ better, you will now plot the cost over a 2-dimensional grid of $\theta_0$ and $\theta_1$ values. You will not need to code anything new for this part, but you should understand how the code you have written already is creating these images.
In the next cell, the code is set up to calculate $J(\theta)$ over a grid of values using the `computeCost` function that you wrote. After executing the following cell, you will have a 2-D array of $J(\theta)$ values. Then, those values are used to produce surface and contour plots of $J(\theta)$ using the matplotlib `plot_surface` and `contourf` functions. The plots should look something like the following:

The purpose of these graphs is to show you how $J(\theta)$ varies with changes in $\theta_0$ and $\theta_1$. The cost function $J(\theta)$ is bowl-shaped and has a global minimum. (This is easier to see in the contour plot than in the 3D surface plot). This minimum is the optimal point for $\theta_0$ and $\theta_1$, and each step of gradient descent moves closer to this point.
```
# grid over which we will calculate J
theta0_vals = np.linspace(-10, 10, 100)
theta1_vals = np.linspace(-1, 4, 100)
# initialize J_vals to a matrix of 0's
J_vals = np.zeros((theta0_vals.shape[0], theta1_vals.shape[0]))
# Fill out J_vals
for i, theta0 in enumerate(theta0_vals):
for j, theta1 in enumerate(theta1_vals):
J_vals[i, j] = computeCost(X, y, [theta0, theta1])
# Because of the way meshgrids work in the surf command, we need to
# transpose J_vals before calling surf, or else the axes will be flipped
J_vals = J_vals.T
# surface plot
fig = pyplot.figure(figsize=(12, 5))
ax = fig.add_subplot(121, projection='3d')
ax.plot_surface(theta0_vals, theta1_vals, J_vals, cmap='viridis')
pyplot.xlabel('theta0')
pyplot.ylabel('theta1')
pyplot.title('Surface')
# contour plot
# Plot J_vals as 15 contours spaced logarithmically between 0.01 and 100
ax = pyplot.subplot(122)
pyplot.contour(theta0_vals, theta1_vals, J_vals, linewidths=2, cmap='viridis', levels=np.logspace(-2, 3, 20))
pyplot.xlabel('theta0')
pyplot.ylabel('theta1')
pyplot.plot(theta[0], theta[1], 'ro', ms=10, lw=2)
pyplot.title('Contour, showing minimum')
pass
```
## Optional Exercises
If you have successfully completed the material above, congratulations! You now understand linear regression and should able to start using it on your own datasets.
For the rest of this programming exercise, we have included the following optional exercises. These exercises will help you gain a deeper understanding of the material, and if you are able to do so, we encourage you to complete them as well. You can still submit your solutions to these exercises to check if your answers are correct.
## 3 Linear regression with multiple variables
In this part, you will implement linear regression with multiple variables to predict the prices of houses. Suppose you are selling your house and you want to know what a good market price would be. One way to do this is to first collect information on recent houses sold and make a model of housing prices.
The file `Data/ex1data2.txt` contains a training set of housing prices in Portland, Oregon. The first column is the size of the house (in square feet), the second column is the number of bedrooms, and the third column is the price
of the house.
<a id="section4"></a>
### 3.1 Feature Normalization
We start by loading and displaying some values from this dataset. By looking at the values, note that house sizes are about 1000 times the number of bedrooms. When features differ by orders of magnitude, first performing feature scaling can make gradient descent converge much more quickly.
```
# Load data
data = np.loadtxt(os.path.join('Data', 'ex1data2.txt'), delimiter=',')
X = data[:, :2]
y = data[:, 2]
m = y.size
# print out some data points
print('{:>8s}{:>8s}{:>10s}'.format('X[:,0]', 'X[:, 1]', 'y'))
print('-'*26)
for i in range(10):
print('{:8.0f}{:8.0f}{:10.0f}'.format(X[i, 0], X[i, 1], y[i]))
```
Your task here is to complete the code in `featureNormalize` function:
- Subtract the mean value of each feature from the dataset.
- After subtracting the mean, additionally scale (divide) the feature values by their respective “standard deviations.”
The standard deviation is a way of measuring how much variation there is in the range of values of a particular feature (most data points will lie within ±2 standard deviations of the mean); this is an alternative to taking the range of values (max-min). In `numpy`, you can use the `std` function to compute the standard deviation.
For example, the quantity `X[:, 0]` contains all the values of $x_1$ (house sizes) in the training set, so `np.std(X[:, 0])` computes the standard deviation of the house sizes.
At the time that the function `featureNormalize` is called, the extra column of 1’s corresponding to $x_0 = 1$ has not yet been added to $X$.
You will do this for all the features and your code should work with datasets of all sizes (any number of features / examples). Note that each column of the matrix $X$ corresponds to one feature.
<div class="alert alert-block alert-warning">
**Implementation Note:** When normalizing the features, it is important
to store the values used for normalization - the mean value and the standard deviation used for the computations. After learning the parameters
from the model, we often want to predict the prices of houses we have not
seen before. Given a new x value (living room area and number of bedrooms), we must first normalize x using the mean and standard deviation that we had previously computed from the training set.
</div>
<a id="featureNormalize"></a>
*You should not submit your solutions.*
```
def featureNormalize(X):
"""
Normalizes the features in X. returns a normalized version of X where
the mean value of each feature is 0 and the standard deviation
is 1. This is often a good preprocessing step to do when working with
learning algorithms.
Parameters
----------
X : array_like
The dataset of shape (m x n).
Returns
-------
X_norm : array_like
The normalized dataset of shape (m x n).
Instructions
------------
First, for each feature dimension, compute the mean of the feature
and subtract it from the dataset, storing the mean value in mu.
Next, compute the standard deviation of each feature and divide
each feature by it's standard deviation, storing the standard deviation
in sigma.
Note that X is a matrix where each column is a feature and each row is
an example. You needto perform the normalization separately for each feature.
Hint
----
You might find the 'np.mean' and 'np.std' functions useful.
"""
# You need to set these values correctly
X_norm = X.copy()
mu = np.zeros(X.shape[1])
sigma = np.zeros(X.shape[1])
# =========================== YOUR CODE HERE =====================
mu = np.mean(X, axis = 0)
sigma = np.std(X, axis = 0)
X_norm = (X_norm - mu) / sigma
# ================================================================
return X_norm, mu, sigma
```
Execute the next cell to run the implemented `featureNormalize` function.
```
# call featureNormalize on the loaded data
X_norm, mu, sigma = featureNormalize(X)
print('Computed mean:', mu)
print('Computed standard deviation:', sigma)
grader[4] = featureNormalize
grader.grade()
```
After the `featureNormalize` function is tested, we now add the intercept term to `X_norm`:
```
# Add intercept term to X
X = np.concatenate([np.ones((m, 1)), X_norm], axis=1)
```
<a id="section5"></a>
### 3.2 Gradient Descent
Previously, you implemented gradient descent on a univariate regression problem. The only difference now is that there is one more feature in the matrix $X$. The hypothesis function and the batch gradient descent update
rule remain unchanged.
You should complete the code for the functions `computeCostMulti` and `gradientDescentMulti` to implement the cost function and gradient descent for linear regression with multiple variables. If your code in the previous part (single variable) already supports multiple variables, you can use it here too.
Make sure your code supports any number of features and is well-vectorized.
You can use the `shape` property of `numpy` arrays to find out how many features are present in the dataset.
<div class="alert alert-block alert-warning">
**Implementation Note:** In the multivariate case, the cost function can
also be written in the following vectorized form:
$$ J(\theta) = \frac{1}{2m}(X\theta - \vec{y})^T(X\theta - \vec{y}) $$
where
$$ X = \begin{pmatrix}
- (x^{(1)})^T - \\
- (x^{(2)})^T - \\
\vdots \\
- (x^{(m)})^T - \\ \\
\end{pmatrix} \qquad \mathbf{y} = \begin{bmatrix} y^{(1)} \\ y^{(2)} \\ \vdots \\ y^{(m)} \\\end{bmatrix}$$
the vectorized version is efficient when you are working with numerical computing tools like `numpy`. If you are an expert with matrix operations, you can prove to yourself that the two forms are equivalent.
</div>
<a id="computeCostMulti"></a>
```
def computeCostMulti(X, y, theta):
"""
Compute cost for linear regression with multiple variables.
Computes the cost of using theta as the parameter for linear regression to fit the data points in X and y.
Parameters
----------
X : array_like
The dataset of shape (m x n+1).
y : array_like
A vector of shape (m, ) for the values at a given data point.
theta : array_like
The linear regression parameters. A vector of shape (n+1, )
Returns
-------
J : float
The value of the cost function.
Instructions
------------
Compute the cost of a particular choice of theta. You should set J to the cst.
"""
# Initialize some useful values
m = y.shape[0] # number of training examples
# You need to return the following variable correctly
J = 0
# ======================= YOUR CODE HERE ===========================
h = np.dot(X, theta)
diff = (h-y)
J = np.square(diff) * (0.5/m)
#print(J)
# ==================================================================
return J
```
*You should now submit your solutions.*
```
grader[5] = computeCostMulti
grader.grade()
```
<a id="gradientDescentMulti"></a>
```
def gradientDescentMulti(X, y, theta, alpha, num_iters):
"""
Performs gradient descent to learn theta.
Updates theta by taking num_iters gradient steps with learning rate alpha.
Parameters
----------
X : array_like
The dataset of shape (m x n+1).
y : array_like
A vector of shape (m, ) for the values at a given data point.
theta : array_like
The linear regression parameters. A vector of shape (n+1, )
alpha : float
The learning rate for gradient descent.
num_iters : int
The number of iterations to run gradient descent.
Returns
-------
theta : array_like
The learned linear regression parameters. A vector of shape (n+1, ).
J_history : list
A python list for the values of the cost function after each iteration.
Instructions
------------
Peform a single gradient step on the parameter vector theta.
While debugging, it can be useful to print out the values of
the cost function (computeCost) and gradient here.
"""
# Initialize some useful values
m = y.shape[0] # number of training examples
# make a copy of theta, which will be updated by gradient descent
theta = theta.copy()
J_history = []
for i in range(num_iters):
# ======================= YOUR CODE HERE ==========================
h = np.dot(X, theta)
gradient = np.dot(X.transpose(), h-y) / m
theta = theta - alpha * gradient
# =================================================================
# save the cost J in every iteration
J_history.append(computeCostMulti(X, y, theta))
return theta, J_history
```
*You should now submit your solutions.*
```
grader[6] = gradientDescentMulti
grader.grade()
"""
Instructions
------------
We have provided you with the following starter code that runs
gradient descent with a particular learning rate (alpha).
Your task is to first make sure that your functions - `computeCost`
and `gradientDescent` already work with this starter code and
support multiple variables.
After that, try running gradient descent with different values of
alpha and see which one gives you the best result.
Finally, you should complete the code at the end to predict the price
of a 1650 sq-ft, 3 br house.
Hint
----
At prediction, make sure you do the same feature normalization.
"""
# Choose some alpha value - change this
alpha = 0.01
num_iters = 400
# init theta and run gradient descent
theta = np.zeros(3)
#print(X)
theta, J_history = gradientDescentMulti(X, y, theta, alpha, num_iters)
# Plot the convergence graph
pyplot.plot(np.arange(len(J_history)), J_history, lw=2)
pyplot.xlabel('Number of iterations')
pyplot.ylabel('Cost J')
#print(theta)
# Display the gradient descent's result
print('theta computed from gradient descent: {:s}'.format(str(theta)))
# Estimate the price of a 1650 sq-ft, 3 br house
# ======================= YOUR CODE HERE ===========================
# Recall that the first column of X is all-ones.
# Thus, it does not need to be normalized.
#predict 1650 square feet and 3 bedrooms price
#normalize this using previous mu and sigma
sqft = (1650 - mu[0]) / sigma[0]
beds = (3 - mu[0]) / sigma[0]
price = theta[0] + theta[1]*sqft + theta[2]*beds
# ===================================================================
print('Predicted price of a 1650 sq-ft, 3 br house (using gradient descent): ${:.0f}'.format(price))
```
#### 3.2.1 Optional (ungraded) exercise: Selecting learning rates
In this part of the exercise, you will get to try out different learning rates for the dataset and find a learning rate that converges quickly. You can change the learning rate by modifying the following code and changing the part of the code that sets the learning rate.
Use your implementation of `gradientDescentMulti` function and run gradient descent for about 50 iterations at the chosen learning rate. The function should also return the history of $J(\theta)$ values in a vector $J$.
After the last iteration, plot the J values against the number of the iterations.
If you picked a learning rate within a good range, your plot look similar as the following Figure.

If your graph looks very different, especially if your value of $J(\theta)$ increases or even blows up, adjust your learning rate and try again. We recommend trying values of the learning rate $\alpha$ on a log-scale, at multiplicative steps of about 3 times the previous value (i.e., 0.3, 0.1, 0.03, 0.01 and so on). You may also want to adjust the number of iterations you are running if that will help you see the overall trend in the curve.
<div class="alert alert-block alert-warning">
**Implementation Note:** If your learning rate is too large, $J(\theta)$ can diverge and ‘blow up’, resulting in values which are too large for computer calculations. In these situations, `numpy` will tend to return
NaNs. NaN stands for ‘not a number’ and is often caused by undefined operations that involve −∞ and +∞.
</div>
<div class="alert alert-block alert-warning">
**MATPLOTLIB tip:** To compare how different learning learning rates affect convergence, it is helpful to plot $J$ for several learning rates on the same figure. This can be done by making `alpha` a python list, and looping across the values within this list, and calling the plot function in every iteration of the loop. It is also useful to have a legend to distinguish the different lines within the plot. Search online for `pyplot.legend` for help on showing legends in `matplotlib`.
</div>
Notice the changes in the convergence curves as the learning rate changes. With a small learning rate, you should find that gradient descent takes a very long time to converge to the optimal value. Conversely, with a large learning rate, gradient descent might not converge or might even diverge!
Using the best learning rate that you found, run the script
to run gradient descent until convergence to find the final values of $\theta$. Next,
use this value of $\theta$ to predict the price of a house with 1650 square feet and
3 bedrooms. You will use value later to check your implementation of the normal equations. Don’t forget to normalize your features when you make this prediction!
*You do not need to submit any solutions for this optional (ungraded) part.*
<a id="section7"></a>
### 3.3 Normal Equations
In the lecture videos, you learned that the closed-form solution to linear regression is
$$ \theta = \left( X^T X\right)^{-1} X^T\vec{y}$$
Using this formula does not require any feature scaling, and you will get an exact solution in one calculation: there is no “loop until convergence” like in gradient descent.
First, we will reload the data to ensure that the variables have not been modified. Remember that while you do not need to scale your features, we still need to add a column of 1’s to the $X$ matrix to have an intercept term ($\theta_0$). The code in the next cell will add the column of 1’s to X for you.
```
# Load data
data = np.loadtxt(os.path.join('Data', 'ex1data2.txt'), delimiter=',')
X = data[:, :2]
y = data[:, 2]
m = y.size
X = np.concatenate([np.ones((m, 1)), X], axis=1)
```
Complete the code for the function `normalEqn` below to use the formula above to calculate $\theta$.
<a id="normalEqn"></a>
```
def normalEqn(X, y):
"""
Computes the closed-form solution to linear regression using the normal equations.
Parameters
----------
X : array_like
The dataset of shape (m x n+1).
y : array_like
The value at each data point. A vector of shape (m, ).
Returns
-------
theta : array_like
Estimated linear regression parameters. A vector of shape (n+1, ).
Instructions
------------
Complete the code to compute the closed form solution to linear
regression and put the result in theta.
Hint
----
Look up the function `np.linalg.pinv` for computing matrix inverse.
"""
theta = np.zeros(X.shape[1])
# ===================== YOUR CODE HERE ============================
temp_1 = np.linalg.inv(np.dot(X.T, X))
temp_2 = np.dot(temp_1, X.T)
theta = np.dot(temp_2, y)
# =================================================================
return theta
```
*You should now submit your solutions.*
```
grader[7] = normalEqn
grader.grade()
```
Optional (ungraded) exercise: Now, once you have found $\theta$ using this
method, use it to make a price prediction for a 1650-square-foot house with
3 bedrooms. You should find that gives the same predicted price as the value
you obtained using the model fit with gradient descent (in Section 3.2.1).
```
# Calculate the parameters from the normal equation
theta = normalEqn(X, y);
# Display normal equation's result
print('Theta computed from the normal equations: {:s}'.format(str(theta)));
# Estimate the price of a 1650 sq-ft, 3 br house
# ====================== YOUR CODE HERE ======================
#price = 0 # You should change this
price = theta[0] + theta[1] * 1650 + theta[2] * 3
# ============================================================
print('Predicted price of a 1650 sq-ft, 3 br house (using normal equations): ${:.0f}'.format(price))
```
| github_jupyter |
# 🔥 FHIR 101 - A Practical Guide
## Hello! What is this Guide?
If any of the following statements applies to you or you've had any of the questions below, then this guide is for you!
- 🤷♀️ I've heard the term FHIR. But is it a server? A database? A data model? Rapid oxidation?
- 🙀 I understand what FHIR is but have no idea how to begin implementing it!
- 😕 I tried reading through [HL7 FHIR](https://www.hl7.org/fhir/), but it is confusing and difficult to navigate.
- 🤔 How does FHIR modeling compare to relational database design?
The primary purpose of this notebook is to quickly educate you on what FHIR is and to walk you through practical exercises that will teach you the FHIR basics:
1. Create and validate a FHIR data model for an entity
2. Add/remove attributes to/from an entity in the model
3. Create attributes which are constrained by ontologies
3. Make entity attributes searchable in the FHIR server
4. Deploy the FHIR data model into a FHIR server
5. Load data into the FHIR server which conforms to the model
6. Search for data in the FHIR server
**If you don't care to know background information and want to get right to development, skip to the [Tutorial](#Tutorial) section!**
### Icons in this Guide
📘 A link to a useful external reference related to the section the icon appears in
⚡️ A key takeaway for the section that this icon appears in
🖐 A hands-on section where you will code something or interact with the server
## What is FHIR?
Let's start with understanding exactly what FHIR is. FHIR stands for `Fast Healthcare Interoperability Resources` and was created by the `Health Level 7` organization or `HL7`.
There is a large, active international community that helps develop the specification. You can read more about it here:
📘 Learn more about [FHIR](http://hl7.org/fhir/index.html)
📘 Learn more about [HL7](http://hl7.org)
### FHIR is ...
A specification that includes all of the following:
- A loosely defined [base data model](https://www.hl7.org/fhir/resourcelist.html) describing things in healthcare (e.g. Patient, Specimen) and how they relate to each other. The base data model also includes definitions and restrictions on server functionality
- How to extend or change the base data model to fit different healthcare use cases
- A database agnostic domain specific language (DSL) for developing files that make up the FHIR data model
- A [RESTful web API specification](https://www.hl7.org/fhir/exchange-module.html) to create, read, update, delete, and search FHIR data in a FHIR server
- A RESTful query language that describes how to construct search queries for a FHIR server
- Standards for how consumers of FHIR data should be authenticated and authorized access to health data.
### FHIR is NOT ...
- A database
- A database schema
- A server
It is important to understand that the FHIR specification is completely technology agnostic. Thus, it does not depend on programming languages or include things like relational database schemas. It is up to the implementers to decide how to implement the data model (i.e. relational database, nosql database, etc) and RESTful API.
## Concepts
The FHIR spec is large and daunting, so this tutorial cannot cover all of the major concepts in FHIR. However, this section and the next should give you all that you need in order to understand what you're doing in the tutorial.
📘 A good resource to learn [FHIR Basics](https://smilecdr.com/docs/tutorial_and_tour/fhir_basics.html#fhir-basics)
### FHIR Resource
The term "Resource" can be confusing, as it is sometimes used to describe different concepts in FHIR.
This diagram should help illustrate the difference between a Resource, type of Resource, and group/category of Resource types:

#### Resource Instances, Types, and Groups
**Resource** - an instance of a type of thing. A Resource has an identifier. Example resources are:
- Patient `PT-001`
- Specimen `SP-001`
- StructureDefinition `KidsFirst-Patient`
- SearchParameter `patient-race`
- ...
**Resource Type** - a type of a thing. Example resource types are:
- Patient
- Specimen
- StructureDefinition
- SearchParameter
- ...
**Resource Type Category/Group** - a logical grouping of Resources. Examples are:
- [Conformance Resources](#Conformance-Resources)
- [Terminology Resources](#Terminology-Resources)
- ...
The most important categories of resource types for the purposes of this tutorial are `Conformance Resources` and `Terminology Resources`, because these are the ones we will create in order to build a FHIR data model. You will learn about them in the [Model Development](#Model-Development) section
**A `Patient` resource might look like this:**
```json
{
"resourceType":"Patient",
"id": "PT-001",
"meta": {
"profile": ["http://fhir.kids-first.io/StructureDefinition/Patient"]
},
"gender": "male",
"name": [
{
"use": "usual",
"family":"Smith",
"given": "Jack",
"period": {
"end": "2001-01-01"
}
}
]
}
```
📘 Learn more about [FHIR Resources](https://www.hl7.org/fhir/resource.html)
### FHIR Data Types
FHIR resource models use a set of data types for holding their values. Data types can be primitive, meaning that they hold a simple value (typically a string, a number, etc.). Other data types are composite, meaning they are a structure containing several elements. Some examples are:
**Primitive Data Types**
- String - some text - `female`
- Decimal - decimal representation of a rational number with significant precision - `1.240`
**Composite Data Types**
- HumanName - A person's complete name, broken into individual parts
```json
{
"family": "Smith",
"given": [ "John", "Frank" ],
"suffix": [ "Jr" ]
}
```
📘 Learn about all [FHIR Data Types](https://www.hl7.org/fhir/datatypes.html)
### FHIR Versions
Changes to the FHIR specification are tracked via different versions. A FHIR version has a name
and a semantic version number. The biggest differences among versions are usually structural and content
changes in the FHIR base model definitions.
The most current and commonly used versions are:
- STU3 or version 3.0.2
- R4 or version 4.0.1
The FHIR version names do not change, but the semantic version numbers can. For example, the `R4` version started with a semantic version number of `4.0.0`. When minor changes were made to `R4`, a new semantic version, `4.0.1` was created to track those changes.
**This tutorial will always use version R4 of the FHIR specification**
📘 Learn more about [FHIR Versions](https://www.hl7.org/fhir/versions.html#versions)
## Model Development
What does it mean to develop a FHIR data model? Well, its not entirely different from developing any kind of data model. Developing a data model usually consists of the following steps:
1. Define entities in the model and their attributes
2. Define relationships between entities
3. Define constraints on entity attributes (i.e. types, min/max, one of, etc)
4. Define constraints on entity relationships (i.e. Specimen must have one Patient, DiagnosticReport must have one Patient, etc.)
5. Validate the model
More concretely, developing a data model in FHIR means:
1. Authoring XML/JSON files containing `Conformance Resource` and `Terminology Resource` payloads
2. Validating `Conformance Resource` and `Terminology Resource` using one of the standalone HL7 validation tools or POSTing the resource payload to a FHIR server
⚡️ **Key Takeaway** - when we say "model" in FHIR, we are referring to `Conformance Resources` and `Terminology Resources`
### Conformance Resources
There are several different types of `Conformance Resources` but the ones that are used most in FHIR model development are:
- `StructureDefinition` - this resource captures model development steps 1 through 4 above for healthcare entities like `Patient`, `Specimen`, `DiagnosticReport`, etc.
- `SearchParameter` - this resource is used to specify an attribute on an entity (e.g. Patient.gender) which can be used to search for instances of that entity on a FHIR server and what keyword/path (e.g. Patient.gender.code.value) should be used when searching.
We will take a deep dive into both of these in the tutorial.
📘 Learn more about [Conformance Resources](https://www.hl7.org/fhir/conformance-module.html)
### Terminology Resources
Some entity attributes might need to be constrained to a particular Value Set which may come from a particular terminology system (or ontology) which defines attributes, their set of valid values, and an alphanumeric code that uniquely identifies each value.
For example, we may want to constrain the `bodySite` attribute on `Specimen` to only have values from NCIT ontology's `Specimen Source Site`. We can do this by defining `Terminology Resource`s and referencing them in the `StructureDefinition` for `Specimen`.
Types of `Terminology Resource`s are:
- `CodeSystem` - describes an Ontology, Terminology, or Enumeration
- `ValueSet` - specifies a set of codes drawn from one or more code systems
You will see a practical example of this later in the tutorial.
📘 Learn about [Terminology Resources](https://www.hl7.org/fhir/terminologies.html)
### Model Documentation
#### Implementation Guide
FHIR data models are documented in a static website called an Implementation Guide. It has a standard format and looks much like the HL7 FHIR website. Implementation Guides describe and visually illustrate:
- The model's `Conformance Resource`s, `Terminology Resource`s
- The differences between the model's `StructureDefinition`s and the ones they extend
- Example Resources for each `StructureDefinition`
- The searchable attributes on each Resource
There is a FHIR Conformance Resource named `ImplementationGuide` which is used to store configuration information for generating the static site.
#### Create and Publish
Implementation Guides are generated using the [HL7 IG Publisher](https://confluence.hl7.org/display/FHIR/IG+Publisher+Documentation) Java CLI. The IG Publisher also fully validates the data model during its site generation process, so it is often used as a model validation tool as well. The Kids First FHIR model uses it in this way.
📘 Learn more about Kids First [IG generation and model validation](https://github.com/kids-first/kf-model-fhir/blob/master/README.md) using IG Publisher
### Model Validation
There are 2 primary ways to validate:
1. Use the [HL7 IG Publisher](https://confluence.hl7.org/display/FHIR/IG+Publisher+Documentation) Java CLI (frequently referred to as IG Publisher or just Publisher)
2. Use a FHIR server
- POST StructureDefinitions to the `/StructureDefinition` endpoint
- POST example Resources to the `/<resourceType>/$validate` endpoint
#### HL7 IG Publisher
The most thorough and standard way to validate a FHIR data model is to use the IG Publisher. The IG Publisher is under active development and is the most up-to-date in terms of its validation logic.
However, since the IG Publisher tightly couples IG site generation with validation, it can be complicated to use since successful validation depends on proper location of resource files and other necessary configuration for IG.
#### Use a FHIR Server
The second method is easier but not standard. All compliant FHIR servers will implement basic validation logic. However, some servers may or may not implement additional validation/checks that the IG Publisher does.
To keep things simple, this tutorial will use the second method of validation.
#### Do I POST `/<resource type>/$validate` or POST `/<resource type>` to validate?
You may be wondering if you can simply validate and create a resource in a single request
by POSTing to `/<resource type>/ endpoint. The answer is you *can*, however, it is not
guarenteed your resource will be validated.
This is because some servers have an on/off switch for validation. If validation is
turned off then you will be able to create invalid resources (does not conform to model).
If the server implements the `$validate` operation/endpoint (not all servers do), then it is safe to assume that the resource you POST to this endpoint will be validated.
⚡️ **Key Takeaway** - the most thorough way to validate a FHIR data model is using the HL7 IG Publisher
## Server Development
### Pick an Existing Solution
You might have noticed that there are not many open source or free FHIR servers currently in development. This is likely due to a couple of factors:
- The FHIR specification is rapidly evolving and the tooling around it is still in early stages of development.
- Server development is hard.
In order to develop a compliant FHIR server, you must:
- Implement the FHIR model validation logic. This is not easy since a FHIR model is authored
in the FHIR DSL. The server must know how to parse the DSL (a.k.a. Conformance Resources) and
execute rules defined in them.
- Implement a parser and query constructor for the FHIR RESTful query language, which is also custom and not based on any known standard. Additionally, the kinds of search queries that can be made with a FHIR search API are fairly rich and complex.
For these reasons it is best to select one of the existing FHIR server solutions rather than develop your own.
### FHIR Server Solutions
From personal experience and a fair bit of research, the most promising ones are:
<table style="bgcolor: none;">
<tbody>
<tr>
<td style="border-right: 1px solid #cccccc; padding: 7px;">
<a href="https://health-samurai.webflow.io/aidbox">
<img src="static/images/aidbox-logo.png" alt="Health Samurai Aidbox"/>
</a>
</td>
<td style="padding: 7px;">
<a href="https://smilecdr.com">
<img src="static/images/smilecdr-logo.png" alt="Smile CDR"/>
</a>
</td>
</tr>
</tbody>
</table>
## Tutorial
### Real World Use Case
In this tutorial we will create a simple data model that is able to capture basic demographic information about Leukemia patients that have had a biosample drawn from them. No protected health information [(PHI)](https://www.hipaajournal.com/considered-phi-hipaa/) data should be captured. In fact there should be no PHI attributes available anywhere in the model.
At a minimum we want to capture:
**Patient Attributes**
- Gender (required)
- Race (required)
- Biological Sex
**Specimen Attributes**
- The patient from whom the sample was drawn
- Anatomical site or location on the body from which the sample was drawn
- Composition - the type of sample - blood, tissue, saliva, etc.
### What You'll Learn
You will learn the following practical concepts in this tutorial:
- How to create a new model for a FHIR resource by extending its base model
- How to add/remove attributes from the base model
- How to make attributes required
- How to make an attribute searchable
- How to add a coded attribute - or one that is restricted to an enumeration defined by an ontology
- How to validate your model
- How to document your model
### Setup
#### Requirements
**FHIR Server**
- For this tutorial we will use [HAPI](http://hapi.fhir.org/), the open-source FHIR server behind Smile CDR.
**Base URL**
- All HTTP requests will be sent to the HAPI public test server's FHIR version R4 base URL: http://hapi.fhir.org/baseR4
**Model Development Editor**
You will need something to author JSON formatted model files. Unfortunately there are almost no user friendly tools for this. Most model development is done in a code editor. This can be difficult as it requires the developer to unecessarily remember complex syntax of the FHIR DSL in order to create and edit model files.
- **Forge** - is a [visual editor for FHIR model development](https://simplifier.net/forge). It does make some things easier, but it can be buggy and was natively developed for Windows. It may be possible to [run Forge on macOS using Wine](http://docs.simplifier.net/forge/forgeInstall.html#running-on-macos).
#### Development Environment 🖐
Here we're going to import the code we need and complete the necessary setup which will be used later in the tutorial.
**You do not need to do this if you would like to use another HTTP client like `Postman` to interact with the FHIR server.**
```
# Be sure to execute this cell so that we can use the client later
from pprint import pprint
from client import FhirApiClient
BASE_URL = 'http://hapi.fhir.org/baseR4'
FHIR_VERSION = '4.0.0'
client = FhirApiClient(base_url=BASE_URL, fhir_version=FHIR_VERSION)
```
### Create a Patient Model
The FHIR base model was intended to include many common healthcare entities
like patients, biospecimens, etc. which cover 80% of healthcare data capture
uses cases.
It is rare to come across a healthcare entity that isn't represented in
one of the existing FHIR resources. The remaining 20% of data capture use cases
that are not satisfied by the base model can easily be satisfied by extending
the base model.
So we can start by looking at the base model's `Patient` `StructureDefinition` to
see if anything needs to be modified.
The HL7 FHIR website has a nice visual representation of the `Patient`
`StructureDefinition`:
[](https://www.hl7.org/fhir/patient.html#resource)
📘 Learn about the [symbol definitions in the structure visual representations](http://hl7.org/fhir/formats.html#table)
We can already see that the `Patient` `StructureDefinition` has one attribute we want:
- `gender`
And many attributes that we'll want to **remove** because they represent PHI fields:
- `name`
- `telecom`
- `address`
- `birthDate`
- `contactPoint`
- `photo`
#### Start with a Template
We've seen that `StructureDefinition`s are complex. It is difficult to remember what attributes are required, the structure, and syntax. Tools like Forge hide that complexity since you're working with a UI.
However, if you don't have or want to use such a tool, you can browse existing `StructureDefinition`s and pick one similar to what you need and use it as a template.
- 📘Implementation Guides published on [HL7 FHIR IG registry](https://registry.fhir.org/guides) also have links to their documented `StructureDefinition`s
- 📘 [Simplifier.net](https://simplifier.net) also has its own Implementation Guide registry at [https://simplifier.net/guides](https://simplifier.net/guides).
#### Create Initial Patient StructureDefinition 🖐
To create a custom data model for patients, we must create a new `StructureDefinition` which
will extend the existing FHIR R4 `StructureDefinition` for `Patient`:
```
# Be sure to execute this cell so that we can use `patient_sd` in later code cells
patient_sd = {
"resourceType": "StructureDefinition",
"url": "http://fhir.kids-first.io/StructureDefinition/Patient",
"version": "0.1.0",
"name": "kids_first_research_participant",
"title": "Kids First Research Participant",
"status": "draft",
"publisher": "Kids First DRC",
"description": "The individual human or other organism.",
"fhirVersion": "4.0.0",
"kind": "resource",
"abstract": False,
"type": "Patient",
"baseDefinition": "http://hl7.org/fhir/StructureDefinition/Patient",
"derivation": "constraint",
"differential": {
"element": [
{
"id": "Patient",
"path": "Patient"
},
{
"id": "Patient.identifier",
"path": "Patient.identifier",
"mustSupport": True
},
{
"id": "Patient.name",
"path": "Patient.name",
"max": "0"
},
{
"id": "Patient.telecom",
"path": "Patient.telecom",
"max": "0"
},
{
"id": "Patient.address",
"path": "Patient.address",
"max": "0"
},
{
"id": "Patient.photo",
"path": "Patient.photo",
"max": "0"
},
{
"id": "Patient.contact",
"path": "Patient.contact",
"max": "0"
},
{
"id": "Patient.gender",
"path": "Patient.gender",
"min": 1
}
]
}
}
```
#### Anatomy of StructureDefinition
We won't cover this in great detail, but here are the important parts of the `StructureDefinition`:
- `resourceType` - every FHIR resource has to declare what type it is
- `baseDefinition` - this points to the `StructureDefinition` that the current one extends. It doesn't have to be one of the base FHIR definitions. It could come from any FHIR data model. This is one of the features of FHIR that helps achieve interoperability.
- `url` - this is a required unique identifier for a `Conformance Resource`. It is a stable [canonical identifier](http://hl7.org/fhir/resource.html#canonical) for the resource and is never intended to change regardless of which FHIR server uses it.
- `fhirVersion` - the FHIR semantic version number is required. It will instruct anything validating this resource to use R4 (4.0.0) validation logic.
- `differential` or `snapshot` - Every `StructureDefinition` must have either a `differential` - a section of content that specifies differences from the base definition, or a `snapshot` section which specifies the full definition. For model development, you almost always use the `differential` section and not the `snapshot`.
##### Important Notes
1. **Canonical URL**
Canonical URLs do not need to be live URLs.
2. **Base Definition**
Let's say `StructureDefinition` A points to `StructureDefinition` B via it's `baseDefinition` attribute. In order for `StructureDefinition` A to pass validation by a FHIR server, `StructureDefinition` B MUST be already be loaded in the FHIR server.
3. **Min and Max Data Types**
You may have noticed that the data types for `min` and `max` differ. This is because `max` can be set to a number or `*` which means many/unlimited, while `min` can only be set to a number.
📘 Learn more about [StructureDefinition anatomy](http://hl7.org/fhir/structuredefinition.html)
📘 Learn more about [Resource identifiers](http://hl7.org/fhir/resource.html#identification)
📘 Browse [FHIR data models from other projects](https://simplifier.net/search?searchtype=Projects)
### Remove Attributes
We're going to start with the easiest modification first: removing an attribute from the base model. In fact, in our initial `Patient` `StructureDefinition`, we've already removed a handful of attributes.
To remove an attribute you simply set its maximum cardinality to 0 in the `StructureDefinition` `differential` section:
```json
{
"id": "Patient.name",
"path": "Patient.name",
"max": "0"
}
```
### Make an Attribute Required
This is just as simple as removing an attribute. To make an attribute required you simply set its minimum cardinality to 1 in the `StructureDefinition` `differential` section:
```json
{
"id": "Patient.gender",
"path": "Patient.gender",
"min": 1
}
```
**A Note on "mustSupport"**
- You might have seen the keyword `mustSupport` in some StructureDefinitions. Sometimes it can be confused with `min=1` which makes an attribute required.
- The `mustSupport` keyword actually doesn't have anything to do with making an attribute required. Instead, `mustSupport` means that implementations that produce or consume resources SHALL provide "support" for the element in some meaningful way.
📘 You can learn more about [mustSupport](https://www.hl7.org/fhir/conformance-rules.html#mustSupport) from the HL7 FHIR spec
📘 Learn about [conformance language key words](https://www.hl7.org/fhir/conformance-rules.html#conflang): SHALL, SHOULD, MAY
### Validate the Patient Model 🖐
Validating the Kids First Patient model means 2 things:
1. Validate the Kids First `Patient` `StructureDefinition` - structure, syntax, cardinality, etc.
2. Test both valid and invalid `Patient` Resources against the Kids First Patient `StructureDefinition`
📘 Learn more about [Resource Validation](https://www.hl7.org/fhir/validation.html)
#### Validate StructureDefinition
```
def validate(payload, expected_success=True, print_result=False):
rt = payload.get('resourceType')
endpoint = f'{client.base_url}/{rt}'
# Let's get into the habit of deleting existing instances before we create one
# The easiest way to do this is to search for StructureDefinitions by canonical URL and
# then delete what is returned (should only be 1 since canonical URL is a unique ID)
success = client.delete_all(endpoint, params={'url': payload['url']})
print(f'DELETE {payload["url"]}')
assert success
# POST the Patient StructureDefinition to the /StructureDefinition endpoint
success, result = client.send_request('POST', endpoint, json=payload)
if print_result:
pprint(result)
print(f'POST {endpoint}, {payload.get("name")}')
assert success == expected_success
validate(patient_sd, print_result=True)
```
#### Create a Patient Resource
```
# Let's start by creating a valid (has all required attributes) Patient resource
patient_rs = {
"resourceType":"Patient",
"id": "kidsfirst-test-patient",
"text": {
"status": "generated",
"div": "<div xmlns=\"http://www.w3.org/1999/xhtml\"><p>A valid patient resource for testing purposes</p></div>"
},
"meta": {
"profile": ["http://fhir.kids-first.io/StructureDefinition/Patient"]
},
"gender": "male"
}
```
#### Validate the Patient Resource
To test the `Patient` resource against the Kids First Patient model you must declare that this resource conforms to the `Patient` `StructureDefinition` like this:
```json
"meta": {
"profile": ["http://fhir.kids-first.io/StructureDefinition/Patient"]
}
```
```
def validate_rs(payload, expected_success=True, create_on_success=True, print_result=False):
resource_type = payload.get('resourceType')
# Validate by POSTing to the /Patient/$validate endpoint
endpoint = f"{client.base_url}/{resource_type}/$validate"
# Since we know this is a valid Patient resource, it should succeed
success, result = client.send_request('POST', endpoint, json=payload)
if print_result:
pprint(result)
print(f'Validate {endpoint}, {payload.get("id")}')
assert success == expected_success
# Now actually create the Patient resource
if create_on_success and success:
endpoint = f"{client.base_url}/{resource_type}/{payload['id']}"
success, result = client.send_request('PUT', endpoint, json=payload)
print(f'PUT {endpoint}')
assert success == expected_success
validate_rs(patient_rs, print_result=True)
```
#### Validate an Invalid Patient Resource
```
# Let's create an invalid resource and make sure it fails
# Invalid since its missing the REQUIRED gender attribute
patient_rs_invalid = {
"resourceType":"Patient",
"id": "kidsfirst-test-patient",
"text": {
"status": "generated",
"div": "<div xmlns=\"http://www.w3.org/1999/xhtml\"><p>An invalid patient resource for testing purposes</p></div>"
},
"meta": {
"profile": ["http://fhir.kids-first.io/StructureDefinition/Patient"]
}
}
validate_rs(patient_rs_invalid, expected_success=False, print_result=True)
```
### Add an Attribute
#### Extensions
Adding a new attribute to a `StructureDefinition` is known as adding an "extension", since you are extending the definition of the base model.
To add an attribute to the `Patient` `StructureDefinition`, you must do 2 things:
1. Define an extension for the attribute by creating a `StructureDefinition` resource for it
2. Declare/reference the extension in the `Patient` `StructureDefinition`
📘 Learn more about defining [extensions](https://www.hl7.org/fhir/extensibility-examples.html)
#### Can I Add an Attribute without an Extension?
You might be wondering if it's possible to simply add an attribute to `Patient` `StructureDefinition` like this:
```json
patient_sd = {
...,
"differential": {
"element": [
...,
{
"id": "Patient.race",
"path": "Patient.race",
"min": 1,
...
}
]
}
}
```
and then create a `Patient` that uses the attribute like this:
```json
patient_rs = {
...,
"race": "White"
}
```
This unfortunately will not work due to the FHIR [limitations of use rules](https://www.hl7.org/fhir/profiling.html#limitations) for Resources. One of these rules states that you cannot add new elements to the base resource.
#### Why So Complicated? ----> Reusability
You might be wondering why adding a new attribute isn't as simple as removing an attribute. There are a couple of reasons for this. The primary reason is because the FHIR spec pushes for interoperability through sharing of model components like extensions.
Since extensions are defined in their own files, they can be reused in a plug and play fashion by other models. We will see this later in the tutorial when we reuse the `karyotypic-sex`extension defined by the [Phenopackets FHIR model](https://aehrc.github.io/fhir-phenopackets-ig/).
Unfortunately, if you don't care about making your attributes shareable and/or you only care about simple key-value type attributes, FHIR does not have a simpler way to define these right now.
#### Simple and Complex Extensions
Another reason why extensions may be complex is because they can represent both simple attributes or complex attributes. A simple attribute might be a key:value pair with a primitive data type, and a complex attribute might be an entirely new structure that might have several other extensions in a nested hierarchy.
⚡️ **Key Takeaway - Reuse existing extensions as much as possible! This is a major benefit of FHIR**
### Create Patient Race Extension 🖐
Similar to StructureDefinitions, the best way to create an extension is to start with an existing extension that is similar to the one you want to create. The best place to choose from is (📘) the official [HL7 FHIR extension registry](https://registry.fhir.org/)
Let's take a look at the extension we'll be adding to the Patient model to represent the patient's race.
```
# Here's what a simple key-value extension with a string data type looks like:
patient_race_ext = {
"resourceType" : "StructureDefinition",
"extension" : [{
"url" : "http://hl7.org/fhir/StructureDefinition/structuredefinition-wg",
"valueCode" : "oo"
},
{
"url" : "http://hl7.org/fhir/StructureDefinition/structuredefinition-fmm",
"valueInteger" : 1
}],
"url" : "http://fhir.kids-first.io/StructureDefinition/patient-race",
"version" : "4.0.0",
"name" : "patient_race",
"status" : "draft",
"date" : "2020-01-13",
"publisher" : "Kids First DRC",
"description" : "Race of patient",
"fhirVersion" : "4.0.0",
"kind" : "complex-type",
"abstract" : False,
"context" : [{
"type" : "element",
"expression" : "Patient"
}],
"type" : "Extension",
"baseDefinition" : "http://hl7.org/fhir/StructureDefinition/Extension",
"derivation" : "constraint",
"differential" : {
"element" : [{
"id" : "Extension",
"path" : "Extension",
"short" : "Race",
"definition" : "Race of patient",
"min" : 0,
"max" : "1",
},
{
"id" : "Extension.extension",
"path" : "Extension.extension",
"max" : "0"
},
{
"id" : "Extension.url",
"path" : "Extension.url",
"fixedUri" : "http://fhir.kids-first.io/StructureDefinition/patient-race"
},
{
"id" : "Extension.value[x]",
"path" : "Extension.value[x]",
"min" : 1,
"type" : [{
"code" : "string"
}]
}]
}
}
```
#### Anatomy of Extension
Similar to our Patient model, an extension is just another `StructureDefinition` except with `type=Extension` instead of `type=Patient`.
For all `Extension` type `StructureDefinition`s:
- `type` must be set to "Extension"
- `kind` must be set to "complex-type" and not "resource", since this is an extension
- `baseDefinition` must be set to the URL of an existing StructureDefinition of type Extension.
Additionally, you must specify in the `differential`:
- the `url` or identifier of the extension
And for simple extensions representing a key and value with a primitive data type, you must specify in the `differential`:
- the `value[x]` or data type of the extension's value
Later on in the tutorial you will learn how to change this from a string type extension to a more complex extension with a `CodeableConcept` data type.
#### Add Race to Patient Model
So far we've just defined the new attribute we want to add to the Patient model. Now we must actually bind it or declare it in the Patient model. Here's how we do this:
```
patient_sd = {
"resourceType": "StructureDefinition",
"url": "http://fhir.kids-first.io/StructureDefinition/Patient",
"version": "0.2.0",
"name": "kids_first_research_participant",
"title": "Kids First Research Participant",
"status": "draft",
"publisher": "Kids First DRC",
"description": "The individual human or other organism.",
"fhirVersion": "4.0.0",
"kind": "resource",
"abstract": False,
"type": "Patient",
"baseDefinition": "http://hl7.org/fhir/StructureDefinition/Patient",
"derivation": "constraint",
"differential": {
"element": [
{
"id": "Patient",
"path": "Patient"
},
{
"id": "Patient.identifier",
"path": "Patient.identifier",
"mustSupport": True
},
{
"id": "Patient.name",
"path": "Patient.name",
"max": "0"
},
{
"id": "Patient.telecom",
"path": "Patient.telecom",
"max": "0"
},
{
"id": "Patient.address",
"path": "Patient.address",
"max": "0"
},
{
"id": "Patient.photo",
"path": "Patient.photo",
"max": "0"
},
{
"id": "Patient.contact",
"path": "Patient.contact",
"max": "0"
},
{
"id": "Patient.gender",
"path": "Patient.gender",
"min": 1
},
{
"id": "Patient.extension",
"path": "Patient.extension",
"slicing": {
"discriminator": [
{
"type": "value",
"path": "url"
}
],
"ordered": False,
"rules": "open"
}
},
{
"id": "Patient.extension:patient-race",
"path": "Patient.extension",
"sliceName": "patient-race",
"short": "Race",
"definition": "Race of patient",
"min": 0,
"max": "1",
"type": [
{
"code": "Extension",
"profile": [
"http://fhir.kids-first.io/StructureDefinition/patient-race"
]
}
],
"isModifier": False
}
]
}
}
```
#### Explanation of Parts
Let's break down each piece. First we'll start with the piece that says we want to use extensions in the `Patient` `StructureDefinition`:
```json
{
"id": "Patient.extension",
"path": "Patient.extension",
"slicing": {
"discriminator": [
{
"type": "value",
"path": "url"
}
],
"ordered": False,
"rules": "open"
}
}
```
- `id`: This must uniquely identify the object in the `differential.element` list. It does not need to be equal to `path` but often times it is, since `path` can uniquely identify the object in the `differential.element` list.
- `path`: This is equal to the fully qualified location of the attribute in the resource payload. All extensions go inside an `extension` element in the Patient resource.
- `slicing`: The `Patient.extension` is a list of extensions and each one must be able to be uniquely identified in that list. The extension's URL is used to "slice" or identify the extension in the list.
- `ordered=False`: order of extensions does not matter in a Patient resource
- `rules=open`: additional extensions can be added in a Patient resource
The next part is the actual binding of the race extension to the `Patient` `StructureDefinition`:
```json
{
"id": "Patient.extension:patient-race",
"path": "Patient.extension",
"sliceName": "patient-race",
"short": "Race",
"definition": "Race of patient",
"min": 0,
"max": "1",
"type": [
{
"code": "Extension",
"profile": [
"http://fhir.kids-first.io/StructureDefinition/patient-race"
]
}
],
"isModifier": False
}
```
The important parts to note here are:
- `min/max`: this says Patient instances can only have one occurrence of the `race` extension and it is not required
- `type`: this binds the Patient's race attribute to it's `StructureDefinition`
#### Validate Patient Model with Race
Now let's validate our updated Patient model and then test it against some example Patient resources which make use of the new race extension
```
# Validate/Create the Patient race extension
validate(patient_race_ext, expected_success=True, print_result=True)
# Validate/Create the updated Patient StructureDefinition which uses Race extension
validate(patient_sd, expected_success=True, print_result=True)
```
#### Create and Validate a Patient With Race
```
patient_rs['extension'] = [
{
'url': 'http://fhir.kids-first.io/StructureDefinition/patient-race',
'valueString': "Asian"
}
]
validate_rs(patient_rs, print_result=True, expected_success=True)
```
### Search for Resources
At this point we're going to take a look at how search works in FHIR so that we can start searching for our resources using the attributes we're adding to our models. Also, this will come in handy for debugging - to find and verify that we've created resources we intended to.
📘 Please note that the [HAPI](http://hapi.fhir.org/) server is periodically purged and contains test data, so the search results may vary. If you have a local FHIR server, you can replace the HAPI URLs in the following examples with your local server URL.
#### FHIR Search DSL
To search for FHIR resources on a FHIR server you need to be able to construct search queries in some sort of language the server understands and then send those queries to the server.
The FHIR Search specification defines its own RESTful query language to accomplish this. This means that search queries are constructed using the URL and its query parameters of the HTTP request.
Why does FHIR define another DSL for this? Because there isn't really a widely accepted standard for a RESTful query language. Operators like `and`, `or`, `>`, `>=`, etc can all be represented in different ways.
#### FHIR Search Queries
FHIR search queries look like this:
```
# Format
GET [base]/[type]?name=value&...
# Example
GET http://hapi.fhir.org/baseR4/Patient?gender=female
```
📘 Learn more about the [FHIR Search Spec](https://www.hl7.org/fhir/search.html)
```
def fetch(endpoint):
success, result = client.send_request('GET', endpoint)
pprint(result)
```
##### Search result options
There are a few different parameters that can be used to filter or condense the search results from a query.
📘 For a full overview of search result options, refer to [this section](https://www.hl7.org/fhir/search.html#_sort) of the FHIR Spec
###### Page counting
The result of querying a FHIR server is a `Bundle`. A `Bundle` is a FHIR resource that contains a list of results that fit the search query criteria. Depending on the server, search results may be paginated, and so a `Bundle` may contain links to the next page. The `_count` parameter sets the number of results per page.
```
fetch('http://hapi.fhir.org/baseR4/Observation?_count=5')
```
###### Summary
The `_summary` parameter returns a subset of information from the search results, such as only certain fields from each result. It can also be used to count the number of results from a query.
```
fetch('http://hapi.fhir.org/baseR4/Observation?_summary=count')
```
##### Fetch all resources types
Fetching all `StructureDefinitions` will give an overview of what types of resources are in a server.
```
fetch('http://hapi.fhir.org/baseR4/StructureDefinition?_summary=count')
fetch('http://hapi.fhir.org/baseR4/StructureDefinition')
```
##### Fetch a resource's schema
The `$snapshot` parameter will return the full schema for a resource, including any `snapshot` that a resource inherits.
```
# Example
GET [base]/StructureDefinition/[id]/$snapshot
```
For example, say there is a resource that inherits from `Patient` and has a `StructureDefinition` that looks like this:
```json
{
"resourceType": "StructureDefinition",
"id": "591026",
"name": "MyPatient",
"type": "Patient",
"baseDefinition": "http://hl7.org/fhir/StructureDefinition/Patient",
"differential": {
"element": [ {
"id": "Patient.extension.race",
...
} ]
}
}
```
Then `$snapshot` will return the full schema for this resource, combining the `differential` from above with the `snapshot` from `Patient`. This is useful because the `$snapshot` parameter will look at a resource's hierarchy and inheritance, and will merge it together, so there is no need for multiple API requests. Note that if a resource and its ancestors don't have have a `snapshot` defined anywhere in a `StructureDefinition`, then the `$snapshot` endpoint will produce an error.
```
fetch('http://hapi.fhir.org/baseR4/StructureDefinition/591026/$snapshot')
```
##### Examining a specific instance of a resource
The general format for looking at a specific instance is:
`GET [base]/[type]/[id]`
This will return the full payload for a resource with a specific ID.
```
fetch('http://hapi.fhir.org/baseR4/Observation/1286681')
```
##### References
📘 For more information on references, refer to [this section](https://www.hl7.org/fhir/search.html#reference) in the FHIR docs
FHIR resources can reference each other. For example, an `Observation` may reference a specific `Patient`. The FHIR search API allows us to search for resources that contain a specific reference. As an example, let's fetch all `Observations` that refer to a `Patient` with an id of 1286630:
```
fetch('http://hapi.fhir.org/baseR4/Observation?subject=Patient/1286630')
```
##### Filtering resources
Earlier, we saw that the general format for a query with search parameters is:
`GET [base]/[type]?name=value&...`
Here are some examples of search queries utilizing research use cases:
```
# Example - fetch all Patients that are alive
fetch('http://hapi.fhir.org/baseR4/Patient?deceased=false')
# Fetch all Conditions that were observed in female Patients
# This is an example of chaining
fetch('http://hapi.fhir.org/baseR4/Condition?patient.gender=female')
# Fetch all Patients that have multiple sclerosis
# This is an example of reverse chaining
fetch('http://hapi.fhir.org/baseR4/Patient?_has:Condition:patient:code=G35')
```
Reverse chaining, demonstrated in the example above, requires this format:
`GET [base]/[resource of interest]?_has:[type of referring resource]:[name of the search parameter containing the reference to the resource of interest]:[search parameter to filter on in referring resource]`
FHIR resources can have multiple references, so it is important to include the name of the field that maps from the referring resource back to the resource of interest. In the above example of `http://hapi.fhir.org/baseR4/Patient?_has:Condition:patient:code=G35`, it is not enough to simply query `http://hapi.fhir.org/baseR4/Patient?_has:Condition:code=G35`, because this does not tell us which reference field in `Condition` contains the reference back to `Patient`.
📘 For more details on chaining and reverse chaining, check out [this section](https://www.hl7.org/fhir/search.html#chaining) of the FHIR docs.
📘Simplifier.net also has a useful article on [Advanced Search Parameters](https://simplifier.net/guide/profilingacademy/Advancedsearchparameters).
#### SearchParameter Resource
So you know what search queries look like, but how do you know what attributes of a Resource or keywords you can use in search queries? All attributes? Some of them? Are keywords the same as attributes?
The `SearchParameter` is a type of `Conformance Resource` which answers this question.
In order to search for a Resource by a particular keyword or Resource attribute, the FHIR server must have a `SearchParameter` defined for it.
Each of the FHIR Resources in the FHIR base model, have a set of predefined SearchParameters that make a subset of the Resources' attributes searchable - by either the name of the attribute or some other keyword used to alias it.
#### Patient Built-In Search Parameters
You can always check the Resource type page on the HL7 FHIR site to see which search parameters are already supported for Resources of that type. For example here is the page for `Patient`

#### Name vs Expression
Something very important to note about `SearchParameter` is the differentiation between it's name and expression.
Take a look at the `family` `SearchParameter`. A `Patient` resource doesn't have a first-level attribute called `family`. In fact, `family` is nested inside the `name` list:
```
{
"resourceType":"Patient",
"id": "test-patient",
"gender": "male",
"name": [
{
"use": "usual",
"family":"Smith",
"given": "Jack",
"period": {
"end": "2001-01-01"
}
}
]
}
```
However, the expression says that when you search for Patients by `family=Smith` it will actually search for all Patients with a name element in the `name` list where `family` equals `Smith`.
The search query for this would look like this:
```
GET http://hapi.fhir.org/baseR4/Patient?family=Smith
```
This is very powerful because you can:
- Search for Resources using a keyword which finds instances by some abitrary attribute in the Resource payload
- Turn on/off search functionality by including/excluding SearchParameters in the server
- Reuse a SearchParameter by making it apply to multiple Resource Types
- Share search functionality across servers by sharing the SearchParameter definition file
**⚡️ Key Takeaway:** SearchParameter allows you to specify a keyword that can be used to search for Resources by some arbitrary attribute on that Resource even if it is nested
#### SearchParameter Types
There is a lot of information on this topic so we cannot cover all of it. However, it is important to understand that SearchParameters have different [types](https://www.hl7.org/fhir/search.html#ptypes) which govern which modifiers can be used as a prefix/suffix to either the name or value of the search parameter in a search query.
For example, for `number`, `date`, and `quantity` type SearchParameters, we can use one of the available modifiers like `gt` in a query:
```
GET http://hapi.fhir.org/baseR4/DiagnosticReport?date=gt2013-01-14
```
This query will return us all `DiagnosticReport` resources that occurred after January 14, 2013.
### Make Patient Searchable by Race 🖐
Ok, now let's create a SearchParameter for the `race` attribute in our Patient model. This is going to consist of 2 steps:
1. Create a SearchParameter for Patient.race
2. Load the SearchParameter into the server
#### Create Search Parameter for Patient.race
```
patient_race_sp = {
"resourceType": "SearchParameter",
"url": "http://fhir.kids-first.io/SearchParameter/patient-race",
"name": "patient-race",
"status": "active",
"experimental": True,
"publisher": "Kids First DRC",
"description": "Search for Patients by their race",
"code": "race",
"base": [
"Patient"
],
"type": "string",
"expression": "Patient.extension.where(url = 'http://fhir.kids-first.io/StructureDefinition/patient-race').value",
"target": [
"Patient"
]
}
```
#### Anatomy of SearchParameter
Again, we won't go over everything, but let's take a look at some of the important parts of a `SearchParameter`:
- `code`: this is the keyword that will be used in search queries
- `name`: a computer friendly name for the search parameter
- `base`: list of Resource types for which this SearchParameter applies
- `target`: this seems completely redundant with `base`. Not sure why it exists, but if you don't include your searches won't work
- `type`: must be one of the valid search parameter types
- `expression`: the path to the attribute value in the resource payload which is being used in the search query. This is specified as a `FHIRpath` expression which is based on [xpath](https://www.w3schools.com/xml/xpath_intro.asp)
📘Learn more about [FHIRpath expressions](https://www.hl7.org/fhir/fhirpath.html)
#### Validate Patient Race Search Parameter
```
validate(patient_race_sp, print_result=True)
```
#### Search for Patient by Race
```
def search(resource_type, params, print_result=False):
endpoint = f'{client.base_url}/{resource_type}'
success, result = client.send_request('GET', endpoint, params=params)
print(f'Searching for {result["request_url"]}, '
f'total found: {result["response"].get("total")}')
if print_result:
pprint(result)
assert success
search('Patient', {'race': 'Asian'}, print_result=True)
```
### Terminology in FHIR
Perhaps you too feel this is a confusing topic in FHIR because you've heard various words used in association with Terminology in FHIR.
You've probably heard the words/phrases: code, coding, codeable concept, Value Sets, etc and are wondering what they all are.
This section will help you understand exactly what these terms mean, how they differ, and how they can be used in your FHIR model.
#### What is Terminology or an Ontology?
Let's start with the very basics. A data model at its core is really just a collection of concepts and how they relate to each other.
**An example - biological sex**
A concept describes some entity in the world or an attribute of some entity in the world. For example, in our model we have a concept called Patient. Our Patient concept also has concepts which describe it - one which we don't have yet but want is biological `sex`.
Now, how does one know what the concept `sex` actually means semantically? And what kinds of values the attribute `sex` on Patient can take on? This is exactly what a terminology service or ontology defines. It also defines how concepts are related to each other (i.e. tree like, graph like, etc).
**Same concept in different ontologies**
A concept can be defined in multiple ontologies. For example, try the following search in your browser to see which ontologies the term `sex` is included in:
```
https://www.ebi.ac.uk/ols/search?q=sex&groupField=iri&start=0
```
#### Coded Value
In FHIR a `code` is an alphanumeric string (e.g. "433152004") used to unambiguously identify a specific concept (e.g. the concept "male", defined as a human with X and Y chromosomes) within a particular ontology (e.g. SNOMED CT).
Since a concept can be defined in different ontologies, and therefore take on different values, we must always specify which ontology or system it comes from.
You will hear the term **"coded value"** used in FHIR documentation. This phrase refers to a pairing of the `code` and the `system` from which it is defined.
#### Data Types for Coded Values
In FHIR there are different data types that can be used to represent coded values.
**code**
This is the simplest one, as it is just an alphanumeric string representing the code and not the system. The system is supposed to be implied based on the context the code appears in:
```json
{
"code": "433152004"
}
```
**Coding**
Specifies the code, the system in which the code is defined (a valid URI), version of the system, and a human friendly string describing the concept which the code identifies.
```json
{
"system" : "http://snomed.info/sct",
"version" : "1.0.0",
"code" : "433152004",
"display": "Male"
}
```
**CodeableConcept**
This data type combines things. It includes 1) a list of `Coding` instances and 2) a human friendly string describing the concept.
You might use this data type when you want to define the concept in multiple ontologies for greater interoperability.
```json
{
"coding": [
{
"system" : "http://snomed.info/sct",
"version" : "1.0.0",
"code" : "433152004",
"display": "Male"
},
{
"system": "http://ncimeta.nci.nih.gov",
"code": "C1710693",
"display": "Male"
}
],
"text": "A human who has a chromosome composition of 46 XY"
}
```
📘Read about [Code, Coding, CodeableConcept](https://www.hl7.org/fhir/datatypes.html#codesystem) data types used to represent coded values
⚡️**Key Take Away** - `code`, `Coding` and `CodeableConcept` are just FHIR data types that represent a concept in the world along with a code to identify the concept in a particular ontology system where the concept is defined.
#### Code Systems
A Code System in FHIR essentially represents an ontology - it defines concepts and gives them meaning through formal definitions, and assign codes that represent the concepts. You can define your own Code System by [creating a CodeSystem FHIR Resource](#CodeSystem-Resource) or you can make use of `CodeSystem`s or ontologies that already exist.
The URI that goes in the `system` attribute of a `Coding` instance, identifies which Code System the code came from. FHIR defines a whole bunch of Code Systems that you can use. For example there are:
**Externally Published Code Systems**
These are popular ontologies you've probably heard of outside of the FHIR context. Here's a subset that FHIR has listed on its [Code System Registry](https://www.hl7.org/fhir/terminologies-systems.html#4.3.0) page. This list tells you exactly what valid URI to use to identify the ontology as FHIR Code System.

**Official HL7 FHIR Code Systems**

**Community FHIR Code Systems**

#### Value Sets
This is a confusing topic in FHIR, perhaps because it could have been named better. A Value Set set in FHIR is a way to select the coded values from one or more Code Systems. The Value Set is then "bound" to a particular attribute on a resource so that values of that attribute are limited to whatever the Value Set allows.
For example, we might add the attribute `sex` to our `Patient` `StructureDefinition` by [creating an extension](#Add-an-Attribute) for it. Then we would bind a Value Set to `sex`, which limits the values of the `sex` attribute to a set of coded values that is defined in the SNOMED CT Code System (http://snomed.info/sct).
**Composable**
Value Sets can select coded values from one or more Code Systems. This is very powerful because it allows you reuse and combine multiple enumerations (maybe from different projects) into one. You can do this by explicitly specifying which coded values to use within a Code System or by saying that a Value Set will be composed of other existing Value Sets.
**Filterable**
A Value Set can also implicitly select a subset of coded values by specifying a filter to use on the coded values within the Code System.
⚡️**Key Take Away -** A `CodeSystem` basically represents an ontology and a `ValueSet` represents a selection of concepts from one or more `CodeSystem`s or ontologies.
⚡️**Key Take Away -** Similar to extensions, try not to reinvent the wheel! First try to find existing `CodeSystem`s and/or `ValueSet`s that you could use to meet your use case. If you cannot find one then create one.
### Add Biological Sex as a Coded Attribute to Patient 🖐
Now we're ready to start modeling. Here's what we're going to do in this section:
1. Create a `CodeSystem` for Biological Sex
- We're going to define our own ontology by creating a `CodeSystem` resource
2. Create a `ValueSet` for Biological Sex
- The Value Set will be simple and select all of the coded values in our `CodeSystem`
3. Add an attribute to the `Patient` model for biological sex
- Create an extension for biological sex
- Add the extension to the `Patient` `StructureDefinition`
4. Bind the `ValuetSet` to the Biological Sex extension in 3
5. Test Things - Create and search for patients with populated `sex` attributes
#### Create a CodeSystem for Biological Sex
We're going to create our own `CodeSystem` which will define a set of codes to represent different types of chromosomal sex compositions for humans.
📘Read more about the [CodeSystem](https://www.hl7.org/fhir/codesystem.html) FHIR resource type
```
# Create and validate the CodeSystem resource
patient_sex_cs = {
"resourceType": "CodeSystem",
"url": "http://fhir.kids-first.io/CodeSystem/biological-sex",
"version": "0.1.0",
"name": "BiologicalSex",
"title": "Biological Sex",
"status": "active",
"experimental": False,
"publisher": "Kids First DRC",
"description": "Represents the chromosomal sex of an individual.",
"caseSensitive": False,
"valueSet": "http://fhir.kids-first.io/ValueSet/biological-sex",
"content": "complete",
"concept": [
{
"code": "UNKNOWN_KARYOTYPE",
"display": "Untyped or inconclusive karyotyping",
"definition": "Untyped or inconclusive karyotyping"
},
{
"code": "XX",
"display": "Female",
"definition": "Female"
},
{
"code": "XY",
"display": "Male",
"definition": "Male"
},
{
"code": "XO",
"display": "Single X chromosome only",
"definition": "Single X chromosome only"
},
{
"code": "XXY",
"display": "Two X and one Y chromosome",
"definition": "Two X and one Y chromosome"
},
{
"code": "XXX",
"display": "Three X chromosomes",
"definition": "Three X chromosomes"
},
{
"code": "XXYY",
"display": "Two X chromosomes and two Y chromosomes",
"definition": "Two X chromosomes and two Y chromosomes"
},
{
"code": "XXXY",
"display": "Three X chromosomes and one Y chromosome",
"definition": "Three X chromosomes and one Y chromosome"
},
{
"code": "XXXX",
"display": "Four X chromosomes",
"definition": "Four X chromosomes"
},
{
"code": "XYY",
"display": "One X and two Y chromosomes",
"definition": "One X and two Y chromosomes"
},
{
"code": "OTHER_KARYOTYPE",
"display": "None of the above types",
"definition": "None of the above types"
}
]
}
validate(patient_sex_cs, print_result=True)
```
#### Anatomy of CodeSystem
Here are the important parts of this resource type:
- `caseSensitive`: Whether or not code comparison is case sensitive when codes within this system are compared to each other.
- `valueSet`: the canonical URL to the ValueSet which contains all coded values from this `CodeSystem`
- `concepts`: list of concept payloads . Each payload contains the concept identifier `code`, a human friendly string `display` describing the concept, and the formal definition of the concept `definition`
#### Create a ValueSet for Patient Biological Sex
We want the Value Set for `sex` to contain all of the coded values in our Code System. This is fairly easy, we simply make a Value Set which is composed of the coded values from our Code System:
```
patient_sex_vs = {
"resourceType": "ValueSet",
"url": "http://fhir.kids-first.io/ValueSet/biological-sex",
"version": "0.1.0",
"name": "BiologicalSex",
"title": "Biological Sex",
"status": "draft",
"experimental": False,
"publisher": "Kids First DRC",
"description": "Represents the chromosomal sex of an individual.",
"compose": {
"include": [
{
"system": "http://fhir.kids-first.io/CodeSystem/biological-sex"
}
]
}
}
validate(patient_sex_vs)
```
#### Anatomy of ValueSet
There is a lot to know about how to compose a group of coded values using a Value Set, but we will just touch on the very basics here:
- `compose`: this element must have the `include` key.
- `compose.include`: has a list of payloads each of which represent the concepts to include in this Value Set. The most basic payload only lists a `CodeSystem` URI which means include all of the coded values from the system.
📘 Read more about [ValueSet](https://www.hl7.org/fhir/valueset.html) and the [composition rules](https://www.hl7.org/fhir/valueset.html#compositions)
#### Create an Extension for Biological Sex
Here we will create an extension for our new `sex` attribute which we will later bind to our `Patient` model (`StructureDefinition`).
Notice this time, we've made the value a `CodeableConcept` which we've bound to our `ValueSet` for biological sex. We could have made the value a `Coding` but then if we later wanted to attach a different ontology to this attribute we wouldn't be able to. Remember that a `CodeableConcept` allows you to have coded values from different Code Systems.
```
# Create and validate biological sex extension
patient_sex_sd = {
"resourceType": "StructureDefinition",
"url": "http://fhir.kids-first.io/StructureDefinition/patient-biological-sex",
"version": "0.1.0",
"name": "BiologicalSex",
"title": "Biological Sex",
"status": "draft",
"publisher": "Kids First DRC",
"description": "The chromosomal sex of an individual.",
"fhirVersion": "4.0.0",
"kind": "complex-type",
"abstract": False,
"context": [
{
"type": "element",
"expression": "Patient"
}
],
"type": "Extension",
"baseDefinition": "http://hl7.org/fhir/StructureDefinition/Extension",
"derivation": "constraint",
"differential": {
"element": [
{
"id": "Extension",
"path": "Extension",
"short": "The chromosomal sex of an individual.",
"definition": "The chromosomal sex of an individual.",
"min": 0,
"max": "1",
"mustSupport": True,
"isModifier": False
},
{
"id": "Extension.extension",
"path": "Extension.extension",
"max": "0"
},
{
"id": "Extension.url",
"path": "Extension.url",
"type": [
{
"code": "uri"
}
],
"fixedUri": "http://fhir.kids-first.io/StructureDefinition/patient-biological-sex"
},
{
"id": "Extension.valueCodeableConcept",
"path": "Extension.valueCodeableConcept",
"type": [
{
"code": "CodeableConcept"
}
],
"binding": {
"extension": [
{
"url": "http://hl7.org/fhir/StructureDefinition/elementdefinition-bindingName",
"valueString": "Biological Sex"
}
],
"strength": "required",
"valueSet": "http://fhir.kids-first.io/ValueSet/biological-sex"
}
}
]
}
}
validate(patient_sex_sd)
# Add biological sex extension to Patient StructureDefinition and validate
patient_sd = {
"resourceType": "StructureDefinition",
"url": "http://fhir.kids-first.io/StructureDefinition/Patient",
"version": "0.1.0",
"name": "kids_first_research_participant",
"title": "Kids First Research Participant",
"status": "draft",
"publisher": "Kids First DRC",
"description": "The individual human or other organism.",
"fhirVersion": "4.0.0",
"kind": "resource",
"abstract": False,
"type": "Patient",
"baseDefinition": "http://hl7.org/fhir/StructureDefinition/Patient",
"derivation": "constraint",
"differential": {
"element": [
{
"id": "Patient",
"path": "Patient"
},
{
"id": "Patient.identifier",
"path": "Patient.identifier",
"mustSupport": True
},
{
"id": "Patient.name",
"path": "Patient.name",
"max": "0"
},
{
"id": "Patient.telecom",
"path": "Patient.telecom",
"max": "0"
},
{
"id": "Patient.address",
"path": "Patient.address",
"max": "0"
},
{
"id": "Patient.photo",
"path": "Patient.photo",
"max": "0"
},
{
"id": "Patient.contact",
"path": "Patient.contact",
"max": "0"
},
{
"id": "Patient.gender",
"path": "Patient.gender",
"mustSupport": True,
"min": 1
},
{
"id": "Patient.extension",
"path": "Patient.extension",
"slicing": {
"discriminator": [
{
"type": "value",
"path": "url"
}
],
"ordered": False,
"rules": "open"
}
},
{
"id": "Patient.extension:patient-race",
"path": "Patient.extension",
"sliceName": "patient-race",
"short": "Race",
"definition": "Race of patient",
"min": 0,
"max": "1",
"type": [
{
"code": "Extension",
"profile": [
"http://fhir.kids-first.io/StructureDefinition/patient-race"
]
}
],
"isModifier": False
},
{
"id": "Patient.extension:biological-sex",
"path": "Patient.extension",
"sliceName": "biological-sex",
"short": "Chromosomal sex of an individual.",
"definition": "This element represents the chromosomal sex of an individual.",
"min": 0,
"max": "1",
"type": [
{
"code": "Extension",
"profile": [
"http://fhir.kids-first.io/StructureDefinition/patient-biological-sex"
]
}
],
"mustSupport": True
}
]
}
}
validate(patient_sd)
```
#### Test Patient with Biological Sex
```
patient_valid = {
"resourceType":"Patient",
"id": "kidsfirst-test-patient",
"text": {
"status": "generated",
"div": "<div xmlns=\"http://www.w3.org/1999/xhtml\"><p>A valid patient resource for testing purposes</p></div>"
},
"meta": {
"profile": ["http://fhir.kids-first.io/StructureDefinition/Patient"]
},
"gender": "male",
"extension": [
{
"url":"http://fhir.kids-first.io/StructureDefinition/patient-biological-sex",
"valueCodeableConcept": {
"coding": [
{
"code": "XY",
"system": "http://fhir.kids-first.io/CodeSystem/biological-sex"
}
],
"text": "Male"
}
}
]
}
validate_rs(patient_valid, print_result=True, expected_success=True)
```
### Searching for Coded Attributes
Before we create a `SearchParameter` resource for our new attribute, it is worth noting a few interesting features of the FHIR search spec when it comes to searching for attributes which have coded values.
#### Token Type SearchParameter
If you recall from the [SearchParameter Types](#SearchParameter-Types) section, a `SearchParameter` can take on different types. Each type has a set of modifiers that can be used with it which change what is being searched for (e.g. `/Patients?gender:not=male` gender is a token type parameter and is allowed to use the modifier `not`)
A `token` type search parameter is different from a regular `string` because it provides a close to exact match search on a string of characters, **potentially scoped by a URI**.
Additionally, for `token` search parameters, matches are case sensitive by default unless the context which it appears specifies otherwise (e.g. `CodeSystem.caseSensitive=False`)
#### Coded Attributes Use Token Type
The `token` search parameter type is used in search parameters for coded attributes since a coded attribute will usually take on one of the FHIR data types: `Coding` or `CodeableConcept` - both of which are scoped by the `CodeSystem` URI
#### How Search Works for Coded Attributes
Let's say we have a `SearchParameter` defined for our `sex` extension. This extension has a FHIR data type of `CodeableConcept`. If you recall our example Patient had `sex` populated like this:
**Patient.sex**
```
"valueCodeableConcept": {
"coding": [
{
"code": "XY",
"system": "http://fhir.kids-first.io/CodeSystem/biological-sex"
}
],
"text": "Male"
}
```
**Search sex=XY**
When you do a search like this:
```
/Patient?sex=XY
```
**Actual Search**
Since this is a `CodeableConcept` data type, the server knows to execute the search like this:
```
Search all Patients and check if any Patient.sex.coding.code matches "XY" exactly
```
#### Modifiers with Token Type
You can change this search behavior with the modifiers that are supported for `token` type search parameters.
Here are some of the modifiers that can be used with `token` type search parameters:
[](https://www.hl7.org/fhir/search.html#token)
#### Example Search Queries
Let's take a look at some search examples for the `Condition` resource. `Condition.code` is a `CodeableConcept` type attribute which identifies the condition, problem or diagnosis for a `Patient`
**Search by code**
Find Conditions where any of the Condition.code.codings.code value matches `ha125`,
regardless of the ontologies/CodeSystems the code appears in.
```
GET [base]/Condition?code=ha125
```
**Search by code within a particular ontology/CodeSystem**
Find Conditions where any of the Condition.code.codings.code value matches `ha125`,
in the `http://acme.org/conditions/codes` CodeSystem.
```
GET [base]/Condition?code=http://acme.org/conditions/codes|ha125
```
**Search using text of CodeableConcept**
Search for any Condition with a code that has the text "headache" associated with it (either in `CodeableConcept.text` or `Coding.display`), regardless of the ontology/CodeSystems.
```
GET [base]/Condition?code:text=headache
```
**Search for codes within a particular portion of the ontology/CodeSystem**
Search for any condition that is subsumed by (in the hierarchy/tree of codes below) the SNOMED CT Code "Neoplasm of liver".
```
GET [base]/Condition?code:below=126851005
```
**⚡️Key Take Away** - FHIR provides a standard way to search for codes and text within a particular ontology and across ontologies. This is very powerful!
### Make Patient Searchable by Biological Sex ✋
We're going to create a SearchParameter for the `sex` attribute in our Patient model. Once again, this is going to consist of 2 steps:
1. Create a SearchParameter for Patient.sex
2. Load the SearchParameter into the server
#### Create and Validate Search Parameter for Patient.sex
```
patient_sex_sp = {
"resourceType": "SearchParameter",
"url": "http://fhir.kids-first.io/SearchParameter/patient-sex",
"name": "patient-sex",
"status": "active",
"experimental": True,
"publisher": "Kids First DRC",
"description": "Search for Patients by their biological sex",
"code": "sex",
"base": [
"Patient"
],
"type": "string",
"expression": "Patient.extension.where(url = 'http://fhir.kids-first.io/StructureDefinition/patient-sex').value",
"target": [
"Patient"
]
}
validate(patient_sex_sp)
```
#### Search for Patient by Sex
**If this returns 0 results it is likely because the HAPI FHIR server has not updated its search indices with the new SearchParameter for Patient.sex**
You may need to wait a day to try again.
```
search('Patient', {'sex': 'XY'}, print_result=True)
```
### Deploy the Model to Your Server
Coming soon..
- Discuss FHIR NPM packages
- Build an archive containing all resources from your IG's FHIR NPM package - including all
resources in any dependent packages
- Load into FHIR server
### Documenting your FHIR Model
Coming soon...
- Discuss IG creation, publishing
| github_jupyter |
# Z-критерий для двух долей
```
import numpy as np
import pandas as pd
import scipy
import statsmodels
from statsmodels.stats.weightstats import *
from statsmodels.stats.proportion import proportion_confint
```
## Загрузка данных
```
data = pd.read_csv('banner_click_stat.txt', header = None, sep = '\t')
data.columns = ['banner_a', 'banner_b']
data.head()
data.describe()
```
## Интервальные оценки долей
$$\frac1{ 1 + \frac{z^2}{n} } \left( \hat{p} + \frac{z^2}{2n} \pm z \sqrt{ \frac{ \hat{p}\left(1-\hat{p}\right)}{n} + \frac{z^2}{4n^2} } \right), \;\; z \equiv z_{1-\frac{\alpha}{2}}$$
```
conf_interval_banner_a = proportion_confint(sum(data.banner_a),
data.shape[0],
method = 'wilson')
conf_interval_banner_b = proportion_confint(sum(data.banner_b),
data.shape[0],
method = 'wilson')
print('95%% confidence interval for a click probability, banner a: [%f, %f]' % conf_interval_banner_a)
print('95%% confidence interval for a click probability, banner b [%f, %f]' % conf_interval_banner_b)
```
## Z-критерий для разности долей (независимые выборки)
| $X_1$ | $X_2$
------------- | -------------|
1 | a | b
0 | c | d
$\sum$ | $n_1$| $n_2$
$$ \hat{p}_1 = \frac{a}{n_1}$$
$$ \hat{p}_2 = \frac{b}{n_2}$$
$$\text{Доверительный интервал для }p_1 - p_2\colon \;\; \hat{p}_1 - \hat{p}_2 \pm z_{1-\frac{\alpha}{2}}\sqrt{\frac{\hat{p}_1(1 - \hat{p}_1)}{n_1} + \frac{\hat{p}_2(1 - \hat{p}_2)}{n_2}}$$
$$Z-статистика: Z({X_1, X_2}) = \frac{\hat{p}_1 - \hat{p}_2}{\sqrt{P(1 - P)(\frac{1}{n_1} + \frac{1}{n_2})}}$$
$$P = \frac{\hat{p}_1{n_1} + \hat{p}_2{n_2}}{{n_1} + {n_2}} $$
```
def proportions_diff_confint_ind(sample1, sample2, alpha = 0.05):
z = scipy.stats.norm.ppf(1 - alpha / 2.)
p1 = float(sum(sample1)) / len(sample1)
p2 = float(sum(sample2)) / len(sample2)
left_boundary = (p1 - p2) - z * np.sqrt(p1 * (1 - p1)/ len(sample1) + p2 * (1 - p2)/ len(sample2))
right_boundary = (p1 - p2) + z * np.sqrt(p1 * (1 - p1)/ len(sample1) + p2 * (1 - p2)/ len(sample2))
return (left_boundary, right_boundary)
def proportions_diff_z_stat_ind(sample1, sample2):
n1 = len(sample1)
n2 = len(sample2)
p1 = float(sum(sample1)) / n1
p2 = float(sum(sample2)) / n2
P = float(p1*n1 + p2*n2) / (n1 + n2)
return (p1 - p2) / np.sqrt(P * (1 - P) * (1. / n1 + 1. / n2))
def proportions_diff_z_test(z_stat, alternative = 'two-sided'):
if alternative not in ('two-sided', 'less', 'greater'):
raise ValueError("alternative not recognized\n"
"should be 'two-sided', 'less' or 'greater'")
if alternative == 'two-sided':
return 2 * (1 - scipy.stats.norm.cdf(np.abs(z_stat)))
if alternative == 'less':
return scipy.stats.norm.cdf(z_stat)
if alternative == 'greater':
return 1 - scipy.stats.norm.cdf(z_stat)
print("95%% confidence interval for a difference between proportions: [%f, %f]" %\
proportions_diff_confint_ind(data.banner_a, data.banner_b))
print("p-value: %f" % proportions_diff_z_test(proportions_diff_z_stat_ind(data.banner_a, data.banner_b)))
print("p-value: %f" % proportions_diff_z_test(proportions_diff_z_stat_ind(data.banner_a, data.banner_b), 'less'))
```
## Z-критерий для разности долей (связанные выборки)
$X_1$ \ $X_2$ | 1| 0 | $\sum$
------------- | -------------|
1 | e | f | e + f
0 | g | h | g + h
$\sum$ | e + g| f + h | n
$$ \hat{p}_1 = \frac{e + f}{n}$$
$$ \hat{p}_2 = \frac{e + g}{n}$$
$$ \hat{p}_1 - \hat{p}_2 = \frac{f - g}{n}$$
$$\text{Доверительный интервал для }p_1 - p_2\colon \;\; \frac{f - g}{n} \pm z_{1-\frac{\alpha}{2}}\sqrt{\frac{f + g}{n^2} - \frac{(f - g)^2}{n^3}}$$
$$Z-статистика: Z({X_1, X_2}) = \frac{f - g}{\sqrt{f + g - \frac{(f-g)^2}{n}}}$$
```
def proportions_diff_confint_rel(sample1, sample2, alpha = 0.05):
z = scipy.stats.norm.ppf(1 - alpha / 2.)
sample = list(zip(sample1, sample2))
n = len(sample)
f = sum([1 if (x[0] == 1 and x[1] == 0) else 0 for x in sample])
g = sum([1 if (x[0] == 0 and x[1] == 1) else 0 for x in sample])
left_boundary = float(f - g) / n - z * np.sqrt(float((f + g)) / n**2 - float((f - g)**2) / n**3)
right_boundary = float(f - g) / n + z * np.sqrt(float((f + g)) / n**2 - float((f - g)**2) / n**3)
return (left_boundary, right_boundary)
def proportions_diff_z_stat_rel(sample1, sample2):
sample = list(zip(sample1, sample2))
n = len(sample)
f = sum([1 if (x[0] == 1 and x[1] == 0) else 0 for x in sample])
g = sum([1 if (x[0] == 0 and x[1] == 1) else 0 for x in sample])
return float(f - g) / np.sqrt(f + g - float((f - g)**2) / n )
print("95%% confidence interval for a difference between proportions: [%f, %f]" \
% proportions_diff_confint_rel(data.banner_a, data.banner_b))
print("p-value: %f" % proportions_diff_z_test(proportions_diff_z_stat_rel(data.banner_a, data.banner_b)))
print("p-value: %f" % proportions_diff_z_test(proportions_diff_z_stat_rel(data.banner_a, data.banner_b), 'less'))
```
| github_jupyter |
# Homework 1
### Due Date: Thursday, July 30, PST, 2020
### Policy Gradient
In this assignment, we will implement vanilla policy gradient algorithm (REINFORCE) covered in the lecture. You will work on i) a function approximator, ii) computing action, iii) collecting samples, iV) training the agent, V) plotting the resutls.
***Complete the missing operations and test your implemented algorithm on the Gym environment.***
***Software requirements:***
- Python >= 3.6
- Tensorflow version <= 1.15.3 (1.X version)
- OpenAI Gym
- Training the agent (policy) can take long time. It is recomended to start solving the problems earlier.
- Save any plots you generated in this notebook. The grade will be given based on the plots you showed.
Make sure the packages you installed meet the requirements.
```
import tensorflow as tf
tf.__version__
import gym
gym.__version__
```
## 1.1 Tensorflow Implementation
We will be implementing policy gradient algorithm using Tensorflow 1.X., which simply updates the parameters of policy from obtaining gradient estimates. The core of policy gradient is to design a function approximator, computing actions, collecting samples, and training the policy. In the below cell, you are encouraged to fill in the components that are missing. ***Your tasks*** are
1. Complete the 'create_model' method to output the mean value for diagonal Guassian policy. Covariance is already defined in the model, so focus on creating neural network model.
2. Complete the 'action_op' method to calculate and return the actions for diagonal Gaussian policy. The applied action should be $\pi(s) = \pi_{\text{mean}}(s) + exp(logstd) * \mathcal{N}(0,1)$
***Hints***:
- Some useful tensorflow classes and methods include: 'tf.exp', 'tf.random_normal'
***IF you are using MAC, please run below box***
```
import os
# MAC user only
os.environ['KMP_DUPLICATE_LIB_OK']='True'
import tensorflow as tf
import numpy as np
import os
import ipdb
class PolicyOpt(object):
def __init__(self, env, linear=False, stochastic=True, hidden_size=32, nonlinearity=tf.nn.relu):
"""Instantiate the policy iteration class.
This initializes the policy optimization with a set of trainable
parameters, and creates a tensorflow session.
Attributes
----------
env : gym.Env
the environment that the policy will be trained on
linear : bool, optional
specifies whether to use a linear or neural network
policy, defaults to False (i.e. Fully-Connected-Neural-Network)
stochastic : bool, optional
specifies whether to use a stochastic or deterministic
policy, defaults to True
hidden_size : list of int, optional
list of hidden layers, with each value corresponding
to the number of nodes in that layer
nonlinearity : tf.nn.*
activation nonlinearity
"""
# clear computation graph
tf.reset_default_graph()
# set a random seed
tf.set_random_seed(1234)
# start a tensorflow session
self.sess = tf.Session()
# environment to train on
self.env = env
# number of elements in the action space
self.ac_dim = env.action_space.shape[0]
# number of elements in the observation space
self.obs_dim = env.observation_space.shape[0]
# actions placeholder
self.a_t_ph = tf.placeholder(dtype=tf.float32,
shape=[None, self.ac_dim])
# state placeholder
self.s_t_ph = tf.placeholder(dtype=tf.float32,
shape=[None, self.obs_dim])
# expected reward placeholder
self.rew_ph = tf.placeholder(dtype=tf.float32,
shape=[None])
# specifies whether the policy is stochastic
self.stochastic = stochastic
# policy that the agent executes during training/testing
self.policy = self.create_model(
args={
"num_actions": self.ac_dim,
"hidden_size": hidden_size,
"linear": linear,
"nonlinearity": nonlinearity,
"stochastic": stochastic,
"scope": "policy",
}
)
# define symbolic action
self.symbolic_action = self.action_op()
# initialize all variables
self.sess.run(tf.global_variables_initializer())
# create saver to save model variables
self.saver = tf.train.Saver()
def create_model(self, args):
"""Create a model for your policy or other components.
Parameters
----------
args : dict
model-specific arguments, with keys:
- "stochastic": True by default
- "hidden_size": Number of neurons in hidden layer
- "num_actions" number of output actions
- "scope": scope of the model
Returns
-------
tf.Variable
mean actions of the policy
tf.Variable
logstd of the policy actions
"""
#################### Build Your Neural Network Here! ####################
??
??
##########################################################################
if args["stochastic"]:
output_logstd = tf.get_variable(name="action_logstd",shape=[self.ac_dim],trainable=True)
else:
output_logstd = None
return output_mean, output_logstd
def action_op(self):
"""
Create a symbolic expression that will be used to compute actions from observations.
When the policy is stochastic, the action follows
a_t = output_mean + exp(output_logstd) * z; z ~ N(0,1)
"""
if self.stochastic:
output_mean, output_logstd = self.policy
#################### Implement a stochastic policy here ####################
# Implement a stochastic version of computing actions. #
# #
# The action in a stochastic policy represented by #
# a diagonal Gaussian distribution with mean "M" and log #
# standard deviation "logstd" is computed as follows: #
# #
# a = M + exp(logstd) * z #
# #
# where z is a random normal value, i.e. z ~ N(0,1) #
# #
# In order to generate numbers from a normal distribution, #
# use the `tf.random_normal` function. #
############################################################################
symbolic_action = ?
else:
symbolic_action, _ = self.policy
return symbolic_action
def compute_action(self, obs):
"""Returns a list of actions for a given observation.
Parameters
----------
obs : np.ndarray
observations
Returns
-------
np.ndarray
actions by the policy for a given observation
"""
return self.sess.run(self.symbolic_action,feed_dict={self.s_t_ph: obs})
def rollout(self, s_mean=None, s_std=None):
"""Collect samples from one rollout of the policy.
Returns
-------
dict
dictionary containing trajectory information for the rollout,
specifically containing keys for "state", "action", "next_state", "reward", and "done"
"""
states = []
next_states = []
actions = []
rewards = []
dones = []
# start a new rollout by re-setting the environment and collecting the initial state
state = self.env.reset()
steps = 0
while True:
steps += 1
# compute the action given the state
if s_mean is not None and s_std is not None:
action = self.compute_action([(state - s_mean) / s_std])
else:
action = self.compute_action([state])
action = action[0]
# advance the environment once and collect the next state, reward, done, and info parameters from the environment
next_state, reward, done, info = self.env.step(action)
# add to the samples list
states.append(state)
actions.append(action)
next_states.append(next_state)
rewards.append(reward)
dones.append(done)
state = next_state
# if the environment returns a True for the done parameter,
# end the rollout before the time horizon is met
if done or steps > env._max_episode_steps:
break
# create the output trajectory
trajectory = {"state": np.array(states, dtype=np.float32),
"reward": np.array(rewards, dtype=np.float32),
"action": np.array(actions, dtype=np.float32),
"next_state": np.array(next_states, dtype=np.float32),
"done": np.array(dones, dtype=np.float32)}
return trajectory
def train(self, args):
"""Abstract training method.
This method will be filled in by algorithm-specific
training operations in subsequent problems.
Parameters
----------
args : dict
algorithm-specific hyperparameters
"""
raise NotImplementedError
```
## 1.2 Tensorflow Interpretation
In order to test your implementation of the **stochastic policy**, run the below cell. The task is to interpret the code you implemented in previous section. If you implement correctly, you can see the value_1 and value_2.
***Question: How do you interpret value_1 and value_2 below cell?***
```
import gym
TEST_ENV = gym.make("Pendulum-v0")
alg = PolicyOpt(TEST_ENV, linear=False)
input_1 = [[0, 1, 2]]
value_1 = alg.sess.run(alg.policy[0], feed_dict={alg.s_t_ph: input_1})
value_2 = alg.compute_action(input_1)
value_1
```
Answer:
```
value_2
```
Answer:
## 1.3 Implement Policy Gradient
In this section, we will implement REINFORCE algorithm presented in the lecture. As a review, the objective is optimize the parameters $\theta$ of some policy $\pi_\theta$ so that the expected return
\begin{equation}
J(\theta) = \mathbb{E} \bigg\{ \sum_{t=0}^T \gamma^t r(s_{t},a_{t}) \bigg\}
\end{equation}
is optimized. In this algorithm, this is done by calculating the gradient $\nabla_\theta J$ and applying a gradient descent method to find a better policy.
\begin{equation}
\theta ' = \theta + \alpha \nabla_\theta J(\theta)
\end{equation}
In the lecture, we derive how we compute $\nabla_{\theta} J(\theta)$. We can rewrite our policy gradient as:
\begin{equation}
\nabla_\theta J (\theta) \approx \frac{1}{N} \sum_{i=0}^{N} \bigg( \sum_{t=0}^{T} \nabla_\theta \log \pi_\theta (a_{it} | s_{it}) \bigg) \bigg( \sum_{t=0}^T \gamma^{t}r_i(t) \bigg)
\end{equation}
Finally, taking into account the causality principle discussed in class, we are able to simplifiy the gradient estimate such as:
\begin{equation}
\nabla_\theta J (\theta) \approx \frac{1}{N} \sum_{i=0}^{N} \sum_{t=0}^{T} \nabla_\theta \log \pi_\theta (a_{it} | s_{it}) \sum_{t'=t}^T \gamma^{t'-t}r_i(t')
\end{equation}
You will be implementing final expression in this assignment!
The process of REINFOCE algorithm follows:
1. Collect samples from current policy $\pi_\theta(s)$ by executing rollouts of the environment.
2. Calculate an estimate for the expected return at state $s_t$.
3. Compute the log-likelihood of each action that was performed by the policy at every given step.
4. Estimate the gradient and update the parameters of policy using gradient-based technique.
5. Repeat steps 1-4 for a number of training iterations.
***Your task*** is to fill out the skeleton code for REINFORCE algorithm,
1. Complete the 'log_likelihoods' method to compute gradient of policy, $\nabla_{\theta}\pi_{\theta}$ for diagonal Guassian policy.
2. Complete the 'compute_expected_return' method to calculate the reward-to-go, $\sum_{t^{\prime}=t}^{T}$.
```
import tensorflow as tf
import numpy as np
import time
import tensorflow_probability as tfp
class REINFORCE(PolicyOpt):
def train(self, num_iterations=1000, steps_per_iteration=1000, learning_rate=int(1e-4), gamma=0.95,
**kwargs):
"""Perform the REINFORE training operation.
Parameters
----------
num_iterations : int
number of training iterations
steps_per_iteration : int
number of individual samples collected every training iteration
learning_rate : float
optimizer learning rate
gamma : float
discount rate
kwargs : dict
additional arguments
Returns
-------
list of float
average return per iteration
"""
# set the discount as an attribute
self.gamma = gamma
# set the learning rate as an attribute
self.learning_rate = learning_rate
# create a symbolic expression to compute the log-likelihoods
log_likelihoods = self.log_likelihoods()
# create a symbolic expression for updating the parameters of your policy
self.opt, self.opt_baseline = self.define_updates(log_likelihoods)
# initialize all variables
self.sess.run(tf.global_variables_initializer())
# average return per training iteration
ret_per_iteration = []
samples = []
for i in range(num_iterations):
# collect samples from the current policy
samples.clear()
steps_so_far = 0
while steps_so_far < steps_per_iteration:
new_samples = self.rollout()
steps_so_far += new_samples["action"].shape[0]
samples.append(new_samples)
# compute the expected returns
v_s = self.compute_expected_return(samples)
# compute and apply the gradients
self.call_updates(log_likelihoods, samples, v_s, **kwargs)
# compute the average cumulative return per iteration
average_rew = np.mean([sum(s["reward"]) for s in samples])
# display iteration statistics
print("Iteration {} return: {}".format(i, average_rew))
ret_per_iteration.append(average_rew)
return ret_per_iteration
def log_likelihoods(self):
"""Create a tensorflow operation that computes the log-likelihood
of each performed action.
"""
output_mean, output_logstd = self.policy
##############################################################
# Create a tf operation to compute the log-likelihood of #
# each action that was performed by the policy #
# #
# The log likelihood in the continuous case where the policy #
# is expressed by a multivariate gaussian can be computing #
# using the tensorflow object: #
# #
# p = tfp.distributions.MultivariateNormalDiag( #
# loc=..., #
# scale_diag=..., #
# ) #
# #
# This method takes as input a mean (loc) and standard #
# deviation (scale_diag), and then can be used to compute #
# the log-likelihood of a variable as follows: #
# #
# log_likelihoods = p.log_prob(...) #
# #
# For this operation, you will want to use placeholders #
# created in the __init__ method of problem 1. #
##############################################################
p = ?
log_likelihoods = ?
return log_likelihoods
def compute_expected_return(self, samples):
"""Compute the expected return from a given starting state.
This is done by using the reward-to-go method.
Parameters
----------
rewards : list of list of float
a list of N trajectories, with each trajectory contain T
returns values (one for each step in the trajectory)
Returns
-------
list of float, or np.ndarray
expected returns for each step in each trajectory
"""
rewards = [s["reward"] for s in samples]
##############################################################
# Estimate the expected return from any given starting state #
# using the reward-to-go method. #
# #
# Using this method, the reward is estimated at every step #
# of the trajectory as follows: #
# #
# r = sum_{t'=t}^T gamma^(t'-t) * r_{t'} #
# #
# where T is the time horizon at t is the index of the #
# current reward in the trajectory. For example, for a given #
# set of rewards r = [1,1,1,1] and discount rate gamma = 1, #
# the expected reward-to-go would be: #
# #
# v_s = [4, 3, 2, 1] #
# #
# You will be able to test this in one of the cells below! #
##############################################################
v_s = ?
return v_s
def define_updates(self, log_likelihoods):
"""Create a tensorflow operation to update the parameters of
your policy.
Parameters
----------
log_likelihoods : tf.Operation
the symbolic expression you created to estimate the log
likelihood of a set of actions
Returns
-------
tf.Operation
a tensorflow operation for computing and applying the
gradients to the parameters of the policy
None
the second component is used in problem 2.b, please ignore
for this problem
"""
loss = - tf.reduce_mean(tf.multiply(log_likelihoods, self.rew_ph))
opt = tf.train.AdamOptimizer(self.learning_rate).minimize(loss)
return opt, None
def call_updates(self, log_likelihoods, samples, v_s, **kwargs):
"""Apply the gradient update methods in a tensorflow session.
Parameters
----------
log_likelihoods: tf.Operation
the symbolic expression you created to estimate the log
likelihood of a set of actions
samples : list of dict
a list of N trajectories, with each trajectory containing
a dictionary of trajectory data (see self.rollout)
v_s : list of float, or np.ndarray
the estimated expected returns from your
`comput_expected_return` function
kwargs : dict
additional arguments (used in question 3)
"""
# concatenate the states
states = np.concatenate([s["state"] for s in samples])
# concatenate the actions
actions = np.concatenate([s["action"] for s in samples])
# execute the optimization step
self.sess.run(self.opt, feed_dict={self.s_t_ph: states,
self.a_t_ph: actions,
self.rew_ph: v_s})
```
Check your 'log_likelihoods' method by running below cell:
```
alg = REINFORCE(TEST_ENV, stochastic=True)
log_likelihoods = alg.log_likelihoods()
# collect a sample output for a given input state
input_s = [[0, 0, 0], [0, 1, 2], [1, 2, 3]]
input_a = [[0], [1], [2]]
# Check
computed = alg.sess.run(log_likelihoods, feed_dict={alg.a_t_ph: input_a, alg.s_t_ph: input_s})
```
Test your 'compute_expected_return' by running below cell:
```
# 1. Test the non-normalized case
alg = REINFORCE(TEST_ENV, stochastic=True)
alg.gamma = 1.0
input_1 = [{"reward": [1, 1, 1, 1]},
{"reward": [1, 1, 1, 1]}]
vs_1 = alg.compute_expected_return(samples=input_1)
ans_1 = np.array([4, 3, 2, 1, 4, 3, 2, 1])
if np.linalg.norm(vs_1 - ans_1) < 1e-3:
print('Great job!')
else:
print('Check your implementation (compute_expected_return)')
```
## 1.4 Testing your algorithm
When you are ready, test your policy gradient algorithms on the *Pendulum-v0* environment in the cell below. *Pendulum-v0* environment is similar to *off-shore wind power*, the goal here is to maintain the Pendulum is upright using control input. The best policy should get around -200 scores. ***Your task*** is to run your REINFORCE algorithm and plot the result!
```
# set this number as 1 for testing your algorithm, and 3 for plotting
NUM_TRIALS = 3
# ===========================================================================
# Do not modify below line
# ===========================================================================
# we will test the algorithms on the Pendulum-v0 gym environment
import gym
env = gym.make("Pendulum-v0")
# train on the REINFORCE algorithm
import numpy as np
r = []
for i in range(NUM_TRIALS):
print("\n==== Training Run {} ====".format(i))
alg = REINFORCE(env, stochastic=True)
res = alg.train(learning_rate=0.005, gamma=0.95, num_iterations=500, steps_per_iteration=15000)
r.append(np.array(res))
alg = None
# save results
np.savetxt("InvertedPendulum_results.csv", np.array(r), delimiter=",")
# collect saved results
import numpy as np
r1 = np.genfromtxt("InvertedPendulum_results.csv", delimiter=",")
all_results = [r1]
labels = ["REINFORCE"]
##############################################################
# Plot your Policy Gradient results below
##############################################################
?
```
The Policy gradient result looks like this:
<img src="HW1_PG_Result_Sample.png"
alt="Markdown Monster icon"
style="float: left; margin-right: 10px;" />
| github_jupyter |
# Bonus1: Parallel Algorithms
### Name: [Your-Name?]
## 0. You will do the following:
1. Read the lecture note: [click here](https://github.com/wangshusen/DeepLearning/blob/master/LectureNotes/Parallel/Parallel.pdf)
2. Implement federated averaging or decentralized optimization.
3. Plot the convergence curve. (The x-axis can be ```number of epochs``` or ```number of communication```. You must make sure the label is correct.)
4. Convert the .IPYNB file to .HTML file.
* The HTML file must contain **the code** and **the output after execution**.
5. Upload this .HTML file to your Google Drive, Dropbox, or your Github repo. (If it is submitted to Google Drive or Dropbox, you must make the file open-access.)
6. Submit the link to this .HTML file to Canvas.
* Example: https://github.com/wangshusen/CS583-2020S/blob/master/homework/Bonus1/Bonus1.html
# 1. Data processing
- Download the Diabete dataset from https://www.csie.ntu.edu.tw/~cjlin/libsvmtools/datasets/binary/diabetes
- Load the data using sklearn.
- Preprocess the data.
## 1.1. Load the data
```
from sklearn import datasets
import numpy
x_sparse, y = datasets.load_svmlight_file('diabetes')
x = x_sparse.todense()
print('Shape of x: ' + str(x.shape))
print('Shape of y: ' + str(y.shape))
```
## 1.2. Partition to training and test sets
```
# partition the data to training and test sets
n = x.shape[0]
n_train = 640
n_test = n - n_train
rand_indices = numpy.random.permutation(n)
train_indices = rand_indices[0:n_train]
test_indices = rand_indices[n_train:n]
x_train = x[train_indices, :]
x_test = x[test_indices, :]
y_train = y[train_indices].reshape(n_train, 1)
y_test = y[test_indices].reshape(n_test, 1)
print('Shape of x_train: ' + str(x_train.shape))
print('Shape of x_test: ' + str(x_test.shape))
print('Shape of y_train: ' + str(y_train.shape))
print('Shape of y_test: ' + str(y_test.shape))
```
## 1.3. Feature scaling
Use the standardization to trainsform both training and test features
```
# Standardization
import numpy
# calculate mu and sig using the training set
d = x_train.shape[1]
mu = numpy.mean(x_train, axis=0).reshape(1, d)
sig = numpy.std(x_train, axis=0).reshape(1, d)
# transform the training features
x_train = (x_train - mu) / (sig + 1E-6)
# transform the test features
x_test = (x_test - mu) / (sig + 1E-6)
print('test mean = ')
print(numpy.mean(x_test, axis=0))
print('test std = ')
print(numpy.std(x_test, axis=0))
```
## 1.4. Add a dimension of all ones
```
n_train, d = x_train.shape
x_train = numpy.concatenate((x_train, numpy.ones((n_train, 1))), axis=1)
n_test, d = x_test.shape
x_test = numpy.concatenate((x_test, numpy.ones((n_test, 1))), axis=1)
print('Shape of x_train: ' + str(x_train.shape))
print('Shape of x_test: ' + str(x_test.shape))
```
| github_jupyter |
# Neuromatch Academy: Week 2, Day 5, Tutorial 3
# Learning to Act: Temporal Difference Learning
# Tutorial Objectives
In this tutorial you learn how to act in the more realistic setting of sequential decisions, formalized by Markov Decision Processes (MDPs). In a sequential decision problem, the actions executed in one state not only may lead to immediate rewards (as in a bandit problem), but may also affect the states experienced next (unlike a bandit problem). In other words, each individual action can ultimately affect affect all future rewards. Thus, making decisions in this setting requires considering each action in terms of their expected **cumulative** future reward.
We will consider here the example of spatial navigation, where actions (movements) in one state (location) affect the states experienced next, and an agent might need to execute a whole sequence of actions before a reward is obtained.
* You will learn the basics of the Q-learning algorithm for estimating action values
* You will understand the concept of on-policy and off-policy value estimation
* You will learn the differences between Q-learning (an off-policy algorithm) and SARSA (an on-policy algorithm)
* You will learn how the concept of exploration and exploitation, reviewed in the bandit case, also applies to the sequential decision setting.
# Setup
```
import matplotlib.pyplot as plt
import numpy as np
from scipy.signal import convolve as conv
#@title Figure Settings
%matplotlib inline
fig_w, fig_h = (8, 6)
plt.rcParams.update({'figure.figsize': (fig_w, fig_h)})
%config InlineBackend.figure_format = 'retina'
#@title Helper Functions
def epsilon_greedy(q, epsilon):
"""Epsilon-greedy policy: selects the maximum value action with probabilty
(1-epsilon) and selects randomly with epsilon probability.
Args:
q (ndarray): an array of action values
epsilon (float): probability of selecting an action randomly
Returns:
int: the chosen action
"""
if np.random.random() > epsilon:
action = np.argmax(q)
else:
action = np.random.choice(len(q))
return action
class CliffWorld:
"""
World: Cliff world.
40 states (4-by-10 grid world).
The mapping from state to the grids are as follows:
30 31 32 ... 39
20 21 22 ... 29
10 11 12 ... 19
0 1 2 ... 9
0 is the starting state (S) and 9 is the goal state (G).
Actions 0, 1, 2, 3 correspond to right, up, left, down.
Moving anywhere from state 9 (goal state) will end the session.
Taking action down at state 11-18 will go back to state 0 and incur a
reward of -100.
Landing in any states other than the goal state will incur a reward of -1.
Going towards the border when already at the border will stay in the same
place.
"""
def __init__(self):
self.name = "cliff_world"
self.n_states = 40
self.n_actions = 4
self.dim_x = 10
self.dim_y = 4
self.init_state = 0
def get_outcome(self, state, action):
if state == 9: # goal state
reward = 0
next_state = None
return next_state, reward
reward = -1 # default reward value
if action == 0: # move right
next_state = state + 1
if state % 10 == 9: # right border
next_state = state
elif state == 0: # start state (next state is cliff)
next_state = None
reward = -100
elif action == 1: # move up
next_state = state + 10
if state >= 30: # top border
next_state = state
elif action == 2: # move left
next_state = state - 1
if state % 10 == 0: # left border
next_state = state
elif action == 3: # move down
next_state = state - 10
if state >= 11 and state <= 18: # next is cliff
next_state = None
reward = -100
elif state <= 9: # bottom border
next_state = state
else:
print("Action must be between 0 and 3.")
next_state = None
reward = None
return int(next_state) if next_state is not None else None, reward
def get_all_outcomes(self):
outcomes = {}
for state in xrange(self.n_states):
for action in xrange(self.n_actions):
next_state, reward = self.get_outcome(state, action)
outcomes[state, action] = [(1, next_state, reward)]
return outcomes
def learn_environment(env, learning_rule, params, max_steps, n_episodes):
# Start with a uniform value function
value = np.ones((env.n_states, env.n_actions))
# Run learning
reward_sums = np.zeros(n_episodes)
# Loop over episodes
for episode in range(n_episodes):
state = env.init_state # initialize state
reward_sum = 0
for t in range(max_steps):
# choose next action
action = epsilon_greedy(value[state], params['epsilon'])
# observe outcome of action on environment
next_state, reward = env.get_outcome(state, action)
# update value function
value = learning_rule(state, action, reward, next_state, value, params)
# sum rewards obtained
reward_sum += reward
if next_state is None:
break # episode ends
state = next_state
reward_sums[episode] = reward_sum
return value, reward_sums
def plot_state_action_values(env, value, ax=None):
"""
Generate plot showing value of each action at each state.
"""
if ax is None:
fig, ax = plt.subplots()
for a in range(env.n_actions):
ax.plot(range(env.n_states), value[:, a], marker='o', linestyle='--')
ax.set(xlabel='States', ylabel='Values')
ax.legend(['R','U','L','D'], loc='lower right')
def plot_quiver_max_action(env, value, ax=None):
"""
Generate plot showing action of maximum value or maximum probability at
each state (not for n-armed bandit or cheese_world).
"""
if ax is None:
fig, ax = plt.subplots()
X = np.tile(np.arange(env.dim_x), [env.dim_y,1]) + 0.5
Y = np.tile(np.arange(env.dim_y)[::-1][:,np.newaxis], [1,env.dim_x]) + 0.5
which_max = np.reshape(value.argmax(axis=1), (env.dim_y,env.dim_x))
which_max = which_max[::-1,:]
U = np.zeros(X.shape)
V = np.zeros(X.shape)
U[which_max == 0] = 1
V[which_max == 1] = 1
U[which_max == 2] = -1
V[which_max == 3] = -1
ax.quiver(X, Y, U, V)
ax.set(
title='Maximum value/probability actions',
xlim=[-0.5, env.dim_x+0.5],
ylim=[-0.5, env.dim_y+0.5],
)
ax.set_xticks(np.linspace(0.5, env.dim_x-0.5, num=env.dim_x))
ax.set_xticklabels(["%d" % x for x in np.arange(env.dim_x)])
ax.set_xticks(np.arange(env.dim_x+1), minor=True)
ax.set_yticks(np.linspace(0.5, env.dim_y-0.5, num=env.dim_y))
ax.set_yticklabels(["%d" % y for y in np.arange(0, env.dim_y*env.dim_x, env.dim_x)])
ax.set_yticks(np.arange(env.dim_y+1), minor=True)
ax.grid(which='minor',linestyle='-')
def plot_heatmap_max_val(env, value, ax=None):
"""
Generate heatmap showing maximum value at each state
"""
if ax is None:
fig, ax = plt.subplots()
if value.ndim == 1:
value_max = np.reshape(value, (env.dim_y,env.dim_x))
else:
value_max = np.reshape(value.max(axis=1), (env.dim_y,env.dim_x))
value_max = value_max[::-1,:]
im = ax.imshow(value_max, aspect='auto', interpolation='none', cmap='afmhot')
ax.set(title='Maximum value per state')
ax.set_xticks(np.linspace(0, env.dim_x-1, num=env.dim_x))
ax.set_xticklabels(["%d" % x for x in np.arange(env.dim_x)])
ax.set_yticks(np.linspace(0, env.dim_y-1, num=env.dim_y))
# ax.set_yticklabels(["%d" % y for y in np.arange(0, env.dim_y*env.dim_x, env.dim_x)])
if env.name != 'windy_cliff_grid':
ax.set_yticklabels(
["%d" % y for y in np.arange(
0, env.dim_y*env.dim_x, env.dim_x)][::-1])
return im
# ax.colorbar(im)
# return fig
def plot_rewards(n_episodes, rewards, average_range=10, ax=None):
"""
Generate plot showing total reward accumulated in each episode.
"""
if ax is None:
fig, ax = plt.subplots()
smoothed_rewards = (conv(rewards, np.ones(average_range), mode='same')
/ average_range)
ax.plot(range(0, n_episodes, average_range),
smoothed_rewards[0:n_episodes:average_range],
marker='o', linestyle='--')
ax.set(xlabel='Episodes', ylabel='Total reward')
# # Plot results
# fig = plot_state_action_values(env, value_qlearning)
# fig = plot_quiver_max_action(env, value_qlearning)
# fig = plot_heatmap_max_val(env, value_qlearning)
# fig = plot_rewards(n_episodes, reward_sums_qlearning, average_range=10)
def plot_performance(env, value, reward_sums):
fig, axes = plt.subplots(nrows=2, ncols=2, figsize=(16, 12))
plot_state_action_values(env, value, ax=axes[0,0])
plot_quiver_max_action(env, value, ax=axes[0,1])
plot_rewards(n_episodes, reward_sums, ax=axes[1,0])
im = plot_heatmap_max_val(env, value, ax=axes[1,1])
fig.colorbar(im)
```
# Grid Worlds
As pointed out, bandits only have a single state and immediate rewards for our actions. Many problems we are interested in have multiple states and delayed rewards, i.e. we won't know if the choices we made will pay off over time, or which actions we took contributed to the outcomes we observed.
In order to explore these ideas, we turn the a common problem setting: the gird world. Grid worlds are simple environments where each state corresponds to a tile on a 2D grid, and the only actions the agent can take are to move up, down, left, or right across the grid tiles. The agent's job is almost always to find a way to a goal tile in the most direct way possible while overcoming some maze or other obstacles, either static or dynamic.
For our discussion we will be looking at the classic Cliff World, or Cliff Walker, environment. This is a 4x10 grid with a starting position in the lower-left and the goal position in the lower-right. Every tile between these two is the "cliff", and should the agent enter the cliff, they will receive a -100 reward and be sent back to the starting position. Every tile other than the cliff produces a -1 reward when entered. The goal tile ends the episode after taking any action from it.
Given these conditions, the maximum achievable reward is -11 (1 up, 9 right, 1 down). Using negative rewards is a common technique to encourage the agent to move and seek out the goal state as fast as possible.
# Q-learning
Now that we have our environment, how can we solve it?
One of the most famous algorithms for estimating action values (aka Q-values) is the Temporal Differences (TD) **control** algorithm known as *Q-learning* (Watkins, 1989).
\begin{align}
Q(s_t,a_t) \leftarrow Q(s_t,a_t) + \alpha \big(r_t + \gamma\max_\limits{a} Q(s_{t+1},a_{t+1}) - Q(s_t,a_t)\big)
\end{align}
where $Q(s,a)$ is the value function for action $a$ at state $s$, $\alpha$ is the learning rate, $r$ is the reward, and $\gamma$ is the temporal discount rate.
The expression $r_t + \gamma\max_\limits{a} Q(s_{t+1},a_{t+1})$ is referred to as the TD target while the full expression
\begin{align}
r_t + \gamma\max_\limits{a} Q(s_{t+1},a_{t+1}) - Q(s_t,a_t),
\end{align}
i.e. the difference between the TD target and the current Q-value, is referred to as the TD error, or reward prediction error.
Because of the max operator used to select the optimal Q-value in the TD target, Q-learning directly estimates the optimal action value, i.e. the cumulative future reward that would be obtained if the agent behaved optimally, regardless of the policy currently followed by the agent. For this reason, Q-learning is referred to as an **off-policy** method.
## Exercise: Implement the Q-learning algorithm
In this exercise you will implement the Q-learning update rule described above. It takes in as arguments the previous state $s_t$, the action $a_t$ taken, the reward received $r_t$, the current state $s_{t+1}$, the Q-value table, and a dictionary of parameters that contain the learning rate $\alpha$ and discount factor $\gamma$. The method returns the updated Q-value table. For the parameter dictionary, $\alpha$: `params['alpha']` and $\gamma$: `params['gamma']`.
```
def q_learning(prev_state, action, reward, state, value, params):
"""Q-learning: updates the value function and returns it.
Args:
prev_state (int): the previous state identifier
action (int): the action taken
reward (float): the reward received
state (int): the current state identifier
value (ndarray): current value function of shape (n_states, n_actions)
params (dict): a dictionary containing the default parameters
Returns:
ndarray: the updated value function of shape (n_states, n_actions)
"""
# value of previous state-action pair
prev_value = value[prev_state, action]
##########################################################
## TODO for students: implement the Q-learning update rule
##########################################################
# comment this out when you've filled
raise NotImplementedError("Student excercise: implement the Q-learning update rule")
# write an expression for finding the maximum Q-value at the current state
if state is None:
max_value = 0
else:
max_value = ...
# write the expression for the reward prediction error
delta = ...
# write the expression of that updates the Q-value of previous state-action pair
value[prev_state, action] = ...
return value
# to_remove solution
def q_learning(prev_state, action, reward, state, value, params):
"""Q-learning: updates the value function and returns it.
Args:
prev_state (int): the previous state identifier
action (int): the action taken
reward (float): the reward received
state (int): the current state identifier
value (ndarray): current value function of shape (n_states, n_actions)
params (dict): a dictionary containing the default parameters
Returns:
ndarray: the updated value function of shape (n_states, n_actions)
"""
# value of previous state-action pair
prev_value = value[prev_state, action]
# maximum Q-value at current state
if state is None:
max_value = 0
else:
max_value = np.max(value[state])
# reward prediction error
delta = reward + params['gamma'] * max_value - prev_value
# update value of previous state-action pair
value[prev_state, action] = prev_value + params['alpha'] * delta
return value
```
Now that we have our Q-learning algorithm, let's see how it handles learning to solve the Cliff World environment.
```
# set for reproducibility, comment out / change seed value for different results
np.random.seed(1)
# parameters needed by our policy and learning rule
params = {
'epsilon': 0.1, # epsilon-greedy policy
'alpha': 0.1, # learning rate
'gamma': 1.0, # discount factor
}
# episodes/trials
n_episodes = 500
max_steps = 1000
# environment initialization
env = CliffWorld()
# solve Cliff World using Q-learning
results = learn_environment(env, q_learning, params, max_steps, n_episodes)
value_qlearning, reward_sums_qlearning = results
# Plot results
plot_performance(env, value_qlearning, reward_sums_qlearning)
```
If all went well, we should see some interesting plots. The top left is a representation of the Q-table itself, showing the values for different actions in different states. Notably, going right from the starting state or down when above the cliff is clearly very bad. The top right figure shows the greedy policy based on the Q-table, while the bottom right shows the best reward we ever received in a given state.
Finally, and most importantly, the bottom left is the actual proof of learning, as we see the total reward steadily increasing until asymptoting at the maximum possible reward of -11.
Feel free to try changing the parameters or random seed and see how the agent's behavior changes.
# SARSA
An alternative to Q-learning, the SARSA algorithm also estimates action values. However, rather than estimating the optimal (off-policy) values, SARSA estimates the **on-policy** action value, i.e. the cumulative future reward that would be obtained if the agent behaved according to its current beliefs.
\begin{align}
Q(s_t,a_t) \leftarrow Q(s_t,a_t) + \alpha \big(r_t + \gamma Q(s_{t+1},a_{t+1}) - Q(s_t,a_t)\big)
\end{align}
where, once again, $Q(s,a)$ is the value function for action $a$ at state $s$, $\alpha$ is the learning rate, $r$ is the reward, and $\gamma$ is the temporal discount rate.
In fact, you will notices that the *only* difference between Q-learning and SARSA is the TD target calculation uses the policy to select the next action (in our case epsilon-greedy) rather than using the action that maximizes the Q-value.
## Exercise: Implement the SARSA algorithm
In this exercise you will implement the SARSA update rule described above. Just like Q-learning, it takes in as arguments the previous state $s_t$, the action $a_t$ taken, the reward received $r_t$, the current state $s_{t+1}$, the Q-value table, and a dictionary of parameters that contain the learning rate $\alpha$ and discount factor $\gamma$. The method returns the updated Q-value table. You may use the `epsilon_greedy` function to acquire the next action. For the parameter dictionary, $\alpha$: `params['alpha']`, $\gamma$: `params['gamma']`, and $\epsilon$: `params['epsilon']`.
```
def sarsa(prev_state, action, reward, state, value, params):
"""SARSA: updates the value function and returns it.
Args:
prev_state (int): the previous state identifier
action (int): the action taken
reward (float): the reward received
state (int): the current state identifier
value (ndarray): current value function of shape (n_states, n_actions)
params (dict): a dictionary containing the default parameters
Returns:
ndarray: the updated value function of shape (n_states, n_actions)
"""
# value of previous state-action pair
prev_value = value[prev_state, action]
##########################################################
## TODO for students: implement the SARSA update rule
##########################################################
# comment this out when you've filled
raise NotImplementedError("Student excercise: implement the SARSA update rule")
# select the expected value at current state based on our policy by sampling
# from it
if state == None:
policy_value = 0
else:
# write an expression for selecting an action using epsilon-greedy
policy_action = ...
# write an expression for obtaining the value of the policy action at the
# current state
policy_value = ...
# write the expression for the reward prediction error
delta = ...
# write the expression of that updates the value of previous state-action pair
value[prev_state, action] = ...
return value
def sarsa(prev_state, action, reward, state, value, params):
"""SARSA: updates the value function and returns it.
Args:
prev_state (int): the previous state identifier
action (int): the action taken
reward (float): the reward received
state (int): the current state identifier
value (ndarray): current value function of shape (n_states, n_actions)
params (dict): a dictionary containing the default parameters
Returns:
ndarray: the updated value function of shape (n_states, n_actions)
"""
# value of previous state-action pair
prev_value = value[prev_state, action]
# select the expected value at current state based on our policy by sampling
# from it
if state == None:
policy_value = 0
else:
# sample action from the policy for the next state
policy_action = epsilon_greedy(value[state], params['epsilon'])
# get the value based on the action sampled from the policy
policy_value = value[state, policy_action]
# reward prediction error
delta = reward + params['gamma'] * policy_value - prev_value
# update value of previous state-action pair
value[prev_state, action] = prev_value + params['alpha'] * delta
return value
```
Now that we have an implementation for SARSA, let's see how it tackles Cliff World. We will again use the same setup we tried with Q-learning.
```
np.random.seed(1)
# parameters needed by our policy and learning rule
params = {
'epsilon': 0.1, # epsilon-greedy policy
'alpha': 0.1, # learning rate
'gamma': 1.0, # discount factor
}
# episodes/trials
n_episodes = 500
max_steps = 1000
# environment initialization
env = CliffWorld()
# learn Cliff World using Sarsa
results = learn_environment(env, sarsa, params, max_steps, n_episodes)
value_sarsa, reward_sums_sarsa = results
# Plot results
plot_performance(env, value_sarsa, reward_sums_sarsa)
```
We should see that SARSA also solves the task with similar looking outcomes to Q-learning. One notable difference is that SARSA seems to be skittsh around the cliff edge and often goes further away before coming back down to the goal.
Again, feel free to try changing the parameters or random seed and see how the agent's behavior changes.
# On-Policy vs Off-Policy
We have now seen an example of both on- and off-policy learning algorithms. Let's compare both Q-learning and SARSA reward results again, side-by-side, to see how they stack up.
```
# parameters needed by our policy and learning rule
params = {
'epsilon': 0.1, # epsilon-greedy policy
'alpha': 0.1, # learning rate
'gamma': 1.0, # discount factor
}
# episodes/trials
n_episodes = 500
max_steps = 1000
# environment initialization
env = CliffWorld()
# learn Cliff World using Sarsa
np.random.seed(1)
results = learn_environment(env, q_learning, params, max_steps, n_episodes)
value_qlearning, reward_sums_qlearning = results
np.random.seed(1)
results = learn_environment(env, sarsa, params, max_steps, n_episodes)
value_sarsa, reward_sums_sarsa = results
fig, ax = plt.subplots()
ax.plot(reward_sums_qlearning, label='Q-learning')
ax.plot(reward_sums_sarsa, label='SARSA')
ax.set(xlabel='Episodes', ylabel='Total reward')
plt.legend(loc='lower right');
```
On this simple Cliff World task, Q-learning and SARSA are almost indistinguisable from a performance standpoint, but we can see that Q-learning has a slight-edge within the 500 episode time horizon. Let's look at the illustrated "greedy policy" plots again.
```
fig, (ax1, ax2) = plt.subplots(ncols=2, figsize=(16, 6))
plot_quiver_max_action(env, value_qlearning, ax=ax1)
ax1.set(title='Q-learning maximum value/probability actions')
plot_quiver_max_action(env, value_sarsa, ax=ax2)
ax2.set(title='SARSA maximum value/probability actions');
```
What immidiately jumps out is that Q-learning learned to go up, then immediately go to the right, skirting the cliff edge, until it hits the wall and goes down to the goal. The policy further away from the cliff is less certain.
SARSA, on the other hand, appears to avoid the cliff edge, going up one more tile before starting over to the goal side. This also clearly solves the challenge of getting to the goal, but does so at an additional -2 cost over the truly optimal route.
Why do you think these behaviors emerged the way they did?
| github_jupyter |
# How to run **ipynb**
1. pip install jupyter
- jupyter notebook my_notebook.ipynb
**_NOTE:_** cd your_work_dir and then `jupyter notebook .` is ok.
# Import *console_color* library
```
# from console_color import RGB, Fore, Style, cprint, create_print
from console_color import *
```
## Only Fore
```
cprint('red: RGB.RED', RGB.RED)
cprint('red: #FF0000', '#FF0000') # It doesn't matter for string upper or lower, and if you don't want to add the sign of #, it's ok too.
cprint('red: (255, 0, 0)', (255, 0, 0))
```
## Only Background
```
cprint('red: RGB.RED', bg=RGB.RED)
cprint('red: #FF0000', bg='#ff0000')
cprint('red: (255, 0, 0)', bg=(255, 0, 0))
```
## Fore + Background
```
cprint('red: RGB.RED, RGB.YELLOW', RGB.RED, RGB.YELLOW)
cprint('red: FF0000, #FFFF00', 'FF0000', '#FFFF00')
cprint('red: (255, 0, 0), (255, 255, 0)', (255, 0, 0), (255, 255, 0))
```
## Style
```
cprint('Italic', style=Style.ITALIC) # The ipynb seems not to show both italic and strike well.
cprint('Italic and Bold', style=Style.BOLD + Style.ITALIC)
cprint('Strike', style=Style.STRIKE)
```
## combine normal text
set **pf = False**
```
print(f"123 {cprint('Fore=Red, bg=Yellow, Style=Italic and Bold', RGB.RED, RGB.YELLOW, Style.BOLD + Style.ITALIC, False)} 456")
print(f"123 {cprint('BOLD', style=Style.BOLD, pf=False)} 456")
```
## keeping the setting
```
ry_print = create_print(fore='FF0000', bg='#FFFF00') # ry: red and yellow
inner_ry_text = create_print('FF0000', '#FFFF00', Style.BOLD + Style.ITALIC, pf=False)
msg = "fore='FF0000', bg='#FFFF00'"
print(msg)
ry_print(msg)
print('...')
ry_print(msg)
print(f'normal text ... {inner_ry_text("fore=red, bg=yellow, style=bold+italic")} !!!')
```
## Color map
```
from itertools import permutations
row_string = ''
for count, rgb in enumerate(permutations((_ for _ in range(0, 256, 40)), 3)):
count += 1
fore_color = RGB.complementary_color(rgb)
row_string += cprint(f'{RGB.tuple2hex_str(rgb)} ', fore_color, rgb, pf=False)
if count % 7 == 0:
print(row_string),
row_string = ''
print(row_string) if len(row_string) > 0 else None
```
## Try by yourself
If you have no idea how to use this library, you can input by indicate, and then it will tell you what the grammar you should write.
```
text = chr(34) + input('text:') + chr(34)
in_fore = input('Fore (#XXYYRR) (r, g, b):')
if in_fore:
in_fore = str(eval(in_fore)) if in_fore.find(',') != -1 else chr(34) + in_fore + chr(34)
in_bg = input('Background (#XXYYRR) (r, g, b):')
if in_bg:
in_bg = str(eval(in_bg)) if in_bg.find(',') != -1 else chr(34) + in_bg + chr(34)
in_style = input('Style (BOLD, ITALIC, URL, STRIKE) sep=" ":').split(' ')
if in_style:
in_style = '+'.join([f'Style.{_.upper()}' for _ in in_style])
print_cmd = f"cprint({text}{', fore=' + in_fore if in_fore else ''}" \
f"{', bg=' + in_bg if in_bg else ''}" \
f"{', style=' + in_style if in_style else ''})"
print(f'grammar: {cprint(print_cmd, RGB.GREEN, RGB.BLACK, pf=False)}')
exec(print_cmd)
```
| github_jupyter |
# An RNN model for temperature data
This time we will be working with real data: daily (Tmin, Tmax) temperature series from 1666 weather stations spanning 50 years. It is to be noted that a pretty good predictor model already exists for temperatures: the average of temperatures on the same day of the year in N previous years. It is not clear if RNNs can do better but we will se how far they can go.
<div class="alert alert-block alert-warning">
This is the solution file. The corresponding tutorial file is [01_RNN_generator_temperatures_playground.ipynb](01_RNN_generator_temperatures_playground.ipynb)
</div>
```
import math
import sys
import time
import numpy as np
import utils_batching
import utils_args
import tensorflow as tf
from tensorflow.python.lib.io import file_io as gfile
print("Tensorflow version: " + tf.__version__)
from matplotlib import pyplot as plt
import utils_prettystyle
import utils_display
```
## Hyperparameters
N_FORWARD = 1: works but model struggles to predict from some positions<br/>
N_FORWARD = 4: better but still bad occasionnally<br/>
N_FORWARD = 8: works perfectly
```
NB_EPOCHS = 5 # number of times the model sees all the data during training
N_FORWARD = 8 # train the network to predict N in advance (traditionnally 1)
RESAMPLE_BY = 5 # averaging period in days (training on daily data is too much)
RNN_CELLSIZE = 128 # size of the RNN cells
N_LAYERS = 2 # number of stacked RNN cells (needed for tensor shapes but code must be changed manually)
SEQLEN = 128 # unrolled sequence length
BATCHSIZE = 64 # mini-batch size
DROPOUT_PKEEP = 0.7 # probability of neurons not being dropped (should be between 0.5 and 1)
ACTIVATION = tf.nn.tanh # Activation function for GRU cells (tf.nn.relu or tf.nn.tanh)
JOB_DIR = "checkpoints"
DATA_DIR = "temperatures"
# potentially override some settings from command-line arguments
if __name__ == '__main__':
JOB_DIR, DATA_DIR = utils_args.read_args1(JOB_DIR, DATA_DIR)
ALL_FILEPATTERN = DATA_DIR + "/*.csv" # pattern matches all 1666 files
EVAL_FILEPATTERN = DATA_DIR + "/USC000*2.csv" # pattern matches 8 files
# pattern USW*.csv -> 298 files, pattern USW*0.csv -> 28 files
print('Reading data from "{}".\nWrinting checkpoints to "{}".'.format(DATA_DIR, JOB_DIR))
```
## Temperature data
This is what our temperature datasets looks like: sequences of daily (Tmin, Tmax) from 1960 to 2010. They have been cleaned up and eventual missing values have been filled by interpolation. Interpolated regions of the dataset are marked in red on the graph.
```
all_filenames = gfile.get_matching_files(ALL_FILEPATTERN)
eval_filenames = gfile.get_matching_files(EVAL_FILEPATTERN)
train_filenames = list(set(all_filenames) - set(eval_filenames))
# By default, this utility function loads all the files and places data
# from them as-is in an array, one file per line. Later, we will use it
# to shape the dataset as needed for training.
ite = utils_batching.rnn_multistation_sampling_temperature_sequencer(eval_filenames)
evtemps, _, evdates, _, _ = next(ite) # gets everything
print('Pattern "{}" matches {} files'.format(ALL_FILEPATTERN, len(all_filenames)))
print('Pattern "{}" matches {} files'.format(EVAL_FILEPATTERN, len(eval_filenames)))
print("Evaluation files: {}".format(len(eval_filenames)))
print("Training files: {}".format(len(train_filenames)))
print("Initial shape of the evaluation dataset: " + str(evtemps.shape))
print("{} files, {} data points per file, {} values per data point"
" (Tmin, Tmax, is_interpolated) ".format(evtemps.shape[0], evtemps.shape[1],evtemps.shape[2]))
# You can adjust the visualisation range and dataset here.
# Interpolated regions of the dataset are marked in red.
WEATHER_STATION = 0 # 0 to 7 in default eval dataset
START_DATE = 0 # 0 = Jan 2nd 1950
END_DATE = 18262 # 18262 = Dec 31st 2009
visu_temperatures = evtemps[WEATHER_STATION,START_DATE:END_DATE]
visu_dates = evdates[START_DATE:END_DATE]
utils_display.picture_this_4(visu_temperatures, visu_dates)
```
## Resampling
Our RNN would need ot be unrolled across 365 steps to capture the yearly temperature cycles. That's a bit too much. We will resample the temparatures and work with 5-day averages for example. This is what resampled (Tmin, Tmax) temperatures look like.
```
# This time we ask the utility function to average temperatures over 5-day periods (RESAMPLE_BY=5)
ite = utils_batching.rnn_multistation_sampling_temperature_sequencer(eval_filenames, RESAMPLE_BY, tminmax=True)
evaltemps, _, evaldates, _, _ = next(ite)
# display five years worth of data
WEATHER_STATION = 0 # 0 to 7 in default eval dataset
START_DATE = 0 # 0 = Jan 2nd 1950
END_DATE = 365*5//RESAMPLE_BY # 5 years
visu_temperatures = evaltemps[WEATHER_STATION, START_DATE:END_DATE]
visu_dates = evaldates[START_DATE:END_DATE]
plt.fill_between(visu_dates, visu_temperatures[:,0], visu_temperatures[:,1])
plt.show()
```
## Visualize training sequences
This is what the neural network will see during training.
```
# The function rnn_multistation_sampling_temperature_sequencer puts one weather station per line in
# a batch and continues with data from the same station in corresponding lines in the next batch.
# Features and labels are returned with shapes [BATCHSIZE, SEQLEN, 2]. The last dimension of size 2
# contains (Tmin, Tmax).
ite = utils_batching.rnn_multistation_sampling_temperature_sequencer(eval_filenames,
RESAMPLE_BY,
BATCHSIZE,
SEQLEN,
N_FORWARD,
nb_epochs=1,
tminmax=True)
# load 6 training sequences (each one contains data for all weather stations)
visu_data = [next(ite) for _ in range(6)]
# Check that consecutive training sequences from the same weather station are indeed consecutive
WEATHER_STATION = 4
utils_display.picture_this_5(visu_data, WEATHER_STATION)
```
## The model definition

<div style="text-align: right; font-family: monospace">
X shape [BATCHSIZE, SEQLEN, 2]<br/>
Y shape [BATCHSIZE, SEQLEN, 2]<br/>
H shape [BATCHSIZE, RNN_CELLSIZE*NLAYERS]
</div>
When executed, this function instantiates the Tensorflow graph for our model.
```
def model_rnn_fn(features, Hin, labels, step, dropout_pkeep):
X = features # shape [BATCHSIZE, SEQLEN, 2], 2 for (Tmin, Tmax)
batchsize = tf.shape(X)[0]
seqlen = tf.shape(X)[1]
pairlen = tf.shape(X)[2] # should be 2 (tmin, tmax)
cells = [tf.nn.rnn_cell.GRUCell(RNN_CELLSIZE, activation=ACTIVATION) for _ in range(N_LAYERS)]
# dropout useful between cell layers only: no output dropout on last cell
cells = [tf.nn.rnn_cell.DropoutWrapper(cell, output_keep_prob = dropout_pkeep) for cell in cells]
# a stacked RNN cell still works like an RNN cell
cell = tf.nn.rnn_cell.MultiRNNCell(cells, state_is_tuple=False)
# X[BATCHSIZE, SEQLEN, 2], Hin[BATCHSIZE, RNN_CELLSIZE*N_LAYERS]
# the sequence unrolling happens here
Yn, H = tf.nn.dynamic_rnn(cell, X, initial_state=Hin, dtype=tf.float32)
# Yn[BATCHSIZE, SEQLEN, RNN_CELLSIZE]
Yn = tf.reshape(Yn, [batchsize*seqlen, RNN_CELLSIZE])
Yr = tf.layers.dense(Yn, 2) # Yr [BATCHSIZE*SEQLEN, 2]
Yr = tf.reshape(Yr, [batchsize, seqlen, 2]) # Yr [BATCHSIZE, SEQLEN, 2]
Yout = Yr[:,-N_FORWARD:,:] # Last N_FORWARD outputs Yout [BATCHSIZE, N_FORWARD, 2]
loss = tf.losses.mean_squared_error(Yr, labels) # labels[BATCHSIZE, SEQLEN, 2]
lr = 0.001 + tf.train.exponential_decay(0.01, step, 1000, 0.5)
optimizer = tf.train.AdamOptimizer(learning_rate=lr)
train_op = optimizer.minimize(loss)
return Yout, H, loss, train_op, Yr
```
## Instantiate the model
```
tf.reset_default_graph() # restart model graph from scratch
# placeholder for inputs
Hin = tf.placeholder(tf.float32, [None, RNN_CELLSIZE * N_LAYERS])
features = tf.placeholder(tf.float32, [None, None, 2]) # [BATCHSIZE, SEQLEN, 2]
labels = tf.placeholder(tf.float32, [None, None, 2]) # [BATCHSIZE, SEQLEN, 2]
step = tf.placeholder(tf.int32)
dropout_pkeep = tf.placeholder(tf.float32)
# instantiate the model
Yout, H, loss, train_op, Yr = model_rnn_fn(features, Hin, labels, step, dropout_pkeep)
```
## Initialize Tensorflow session
This resets all neuron weights and biases to initial random values
```
# variable initialization
sess = tf.Session()
init = tf.global_variables_initializer()
sess.run([init])
saver = tf.train.Saver(max_to_keep=1)
```
## The training loop
You can re-execute this cell to continue training. <br/>
<br/>
Training data must be batched correctly, one weather station per line, continued on the same line across batches. This way, output states computed from one batch are the correct input states for the next batch. The provided utility function `rnn_multistation_sampling_temperature_sequencer` does the right thing.

```
losses = []
indices = []
last_epoch = 99999
last_fileid = 99999
for i, (next_features, next_labels, dates, epoch, fileid) in enumerate(
utils_batching.rnn_multistation_sampling_temperature_sequencer(train_filenames,
RESAMPLE_BY,
BATCHSIZE,
SEQLEN,
N_FORWARD,
NB_EPOCHS, tminmax=True)):
# reinintialize state between epochs or when starting on data from a new weather station
if epoch != last_epoch or fileid != last_fileid:
batchsize = next_features.shape[0]
H_ = np.zeros([batchsize, RNN_CELLSIZE * N_LAYERS])
print("State reset")
#train
feed = {Hin: H_, features: next_features, labels: next_labels, step: i, dropout_pkeep: DROPOUT_PKEEP}
Yout_, H_, loss_, _, Yr_ = sess.run([Yout, H, loss, train_op, Yr], feed_dict=feed)
# print progress
if i%20 == 0:
print("{}: epoch {} loss = {} ({} weather stations this epoch)".format(i, epoch, np.mean(loss_), fileid+1))
sys.stdout.flush()
if i%10 == 0:
losses.append(np.mean(loss_))
indices.append(i)
# This visualisation can be helpful to see how the model "locks" on the shape of the curve
# if i%100 == 0:
# plt.figure(figsize=(10,2))
# plt.fill_between(dates, next_features[0,:,0], next_features[0,:,1]).set_alpha(0.2)
# plt.fill_between(dates, next_labels[0,:,0], next_labels[0,:,1])
# plt.fill_between(dates, Yr_[0,:,0], Yr_[0,:,1]).set_alpha(0.8)
# plt.show()
last_epoch = epoch
last_fileid = fileid
# save the trained model
SAVEDMODEL = JOB_DIR + "/ckpt" + str(int(time.time()))
tf.saved_model.simple_save(sess, SAVEDMODEL,
inputs={"features":features, "Hin":Hin, "dropout_pkeep":dropout_pkeep},
outputs={"Yout":Yout, "H":H})
plt.ylim(ymax=np.amax(losses[1:])) # ignore first value for scaling
plt.plot(indices, losses)
plt.show()
```
## Inference
This is a generative model: run an trained RNN cell in a loop
```
def prediction_run(predict_fn, prime_data, run_length):
H = np.zeros([1, RNN_CELLSIZE * N_LAYERS]) # zero state initially
Yout = np.zeros([1, N_FORWARD, 2])
data_len = prime_data.shape[0]-N_FORWARD
# prime the state from data
if data_len > 0:
Yin = np.array(prime_data[:-N_FORWARD])
Yin = np.reshape(Yin, [1, data_len, 2]) # reshape as one sequence of pairs (Tmin, Tmax)
r = predict_fn({'features': Yin, 'Hin':H, 'dropout_pkeep':1.0}) # no dropout during inference
Yout = r["Yout"]
H = r["H"]
# initaily, put real data on the inputs, not predictions
Yout = np.expand_dims(prime_data[-N_FORWARD:], axis=0)
# Yout shape [1, N_FORWARD, 2]: batch of a single sequence of length N_FORWARD of (Tmin, Tmax) data pointa
# run prediction
# To generate a sequence, run a trained cell in a loop passing as input and input state
# respectively the output and output state from the previous iteration.
results = []
for i in range(run_length//N_FORWARD+1):
r = predict_fn({'features': Yout, 'Hin':H, 'dropout_pkeep':1.0}) # no dropout during inference
Yout = r["Yout"]
H = r["H"]
results.append(Yout[0]) # shape [N_FORWARD, 2]
return np.concatenate(results, axis=0)[:run_length]
```
## Validation
```
QYEAR = 365//(RESAMPLE_BY*4)
YEAR = 365//(RESAMPLE_BY)
# Try starting predictions from January / March / July (resp. OFFSET = YEAR or YEAR+QYEAR or YEAR+2*QYEAR)
# Some start dates are more challenging for the model than others.
OFFSET = 30*YEAR+1*QYEAR
PRIMELEN=5*YEAR
RUNLEN=3*YEAR
PRIMELEN=512
RUNLEN=256
RMSELEN=3*365//(RESAMPLE_BY*2) # accuracy of predictions 1.5 years in advance
# Restore the model from the last checkpoint saved previously.
# Alternative checkpoints:
# Once you have trained on all 1666 weather stations on Google Cloud ML Engine, you can load the checkpoint from there.
# SAVEDMODEL = "gs://{BUCKET}/sinejobs/sines_XXXXXX_XXXXXX/ckptXXXXXXXX"
# A sample checkpoint is provided with the lab. You can try loading it for comparison.
# SAVEDMODEL = "temperatures_best_checkpoint"
predict_fn = tf.contrib.predictor.from_saved_model(SAVEDMODEL)
for evaldata in evaltemps:
prime_data = evaldata[OFFSET:OFFSET+PRIMELEN]
results = prediction_run(predict_fn, prime_data, RUNLEN)
utils_display.picture_this_6(evaldata, evaldates, prime_data, results, PRIMELEN, RUNLEN, OFFSET, RMSELEN)
rmses = []
bad_ones = 0
for offset in [YEAR, YEAR+QYEAR, YEAR+2*QYEAR]:
for evaldata in evaltemps:
prime_data = evaldata[offset:offset+PRIMELEN]
results = prediction_run(predict_fn, prime_data, RUNLEN)
rmse = math.sqrt(np.mean((evaldata[offset+PRIMELEN:offset+PRIMELEN+RMSELEN] - results[:RMSELEN])**2))
rmses.append(rmse)
if rmse>7: bad_ones += 1
print("RMSE on {} predictions (shaded area): {}".format(RMSELEN, rmse))
print("Average RMSE on {} weather stations: {} ({} really bad ones, i.e. >7.0)".format(len(evaltemps), np.mean(rmses), bad_ones))
sys.stdout.flush()
```
Copyright 2018 Google LLC
Licensed under the Apache License, Version 2.0 (the "License");
you may not use this file except in compliance with the License.
You may obtain a copy of the License at
[http://www.apache.org/licenses/LICENSE-2.0](http://www.apache.org/licenses/LICENSE-2.0)
Unless required by applicable law or agreed to in writing, software
distributed under the License is distributed on an "AS IS" BASIS,
WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
See the License for the specific language governing permissions and
limitations under the License.
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
import os
import scipy.io
from keras.models import Sequential
from keras.models import model_from_json
from keras.layers import Dense, Conv2D, Flatten
from keras.utils import to_categorical
from keras.utils.vis_utils import plot_model
```
# Deep Learning study on the results of the 1D Pseudo-Wigner Distribution using Neural Networks
**Why?**
Check if the wigner distribution of an hologram is capable to give us enough information to be able to predict how many point sources generated the hologram (1 to 5 sources).
**How?**
Using a Convolutional Neural Networks (CNN) to solve this classification problem.
**What?**
Using the keras libray (python).
**Some examples:**
* https://towardsdatascience.com/building-a-convolutional-neural-network-cnn-in-keras-329fbbadc5f5
### Load dataset
```
%%time
path = 'output/wigner_distribution/'
file_name = 'wd_results.npy'
dataset = np.load(path + file_name)
print(dataset.shape)
print('Total number of holograms: ' + str(dataset.shape[0]))
print('Number of holograms per class: ' + str(int(dataset.shape[0]/ 5)))
```
## CNN (Convolutional Neural Networks)
### Data pre-processing
```
def compute_targets_array(nb_class, X_train):
"""
Compute an array with the targets of the dataset. Note that the number on the array correspond to the number of
sources minus one. E.g. Y_array = 1, the number of point sources is 2.
"""
# Number of the examples
nb_holograms = X_train.shape[0]
# Number of examples per class
nb_holograms_class = int(nb_holograms / nb_class)
# Y vector
Y_array = np.zeros((nb_holograms,))
counter = 1
target = 0
for i in range(nb_holograms):
if counter == (nb_holograms_class + 1):
target = target + 1
counter = 1
Y_array[i,] = target
counter = counter + 1
return Y_array
# Select one of the 8 frequencies ! BUG !!!!!!!!!!!!
X_train = dataset[:,0,:,:]
# The 1 signify that the images are greyscale
X_train = X_train.reshape(X_train.shape[0], 200, 200,1)
print(X_train.shape)
# Compute array of targets
nb_class = 5
Y_array = compute_targets_array(nb_class, X_train)
print(Y_array.shape)
print(Y_array)
# One-hot encode target column
Y_train = to_categorical(Y_array)
print(Y_train.shape)
```
### Building the model
```
# Create model
model = Sequential() # allows build a model layer by layer
# Add model layers
# Conv2D layer:
# 64 nodes, 3x3 filter matrix, Rectified Linear Activation as activation function,
# shape of each input (200, 200, 1,) with 1 signifying images are greyscale
model.add(Conv2D(64, kernel_size=3, activation='relu', input_shape=(200,200,1)))
# 32 nodes
model.add(Conv2D(32, kernel_size=3, activation='relu'))
# Flatten layer: connection between the convolution and dense layers
model.add(Flatten())
# Dense layer: used for the output layer
# 5 nodes for the output layer, one for each possible outcome (1-5)
# 'softmax' as activation function, it makes the output sump up to 1 so the output
# can be interpreted as probalities
model.add(Dense(5, activation='softmax'))
```
### Compiling the model
```
# Three parameters:
# optmizer: 'adam'
# loss function: 'categorical_crossentropy', the most common choice for classification
# metrics: 'accuracy', to see the accuracy score
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
```
### Training the model
```
%%time
# Number of epochs: number of tmes the model wil cycle trough the data
model.fit(X_train, Y_train, validation_data=(X_train, Y_train), epochs=30)
```
### Evalutation
```
# Evaluate the keras model
_, accuracy = model.evaluate(X_train, Y_train, verbose=0)
print('Accuracy: %.2f%%' % (accuracy*100))
```
### Make predictions
```
# Make probability predictions with the model
predictions = model.predict(X_train)
# Round predictions
rounded = [round(x[0]) for x in predictions]
# Make class predictions with the model
predictions = model.predict_classes(X_train)
# Summarize the first 5 cases
for i in range(5):
print('Predicted: %d (expected: %d)' % (predictions[i], Y_array[i]))
```
### Save weights and model
```
%%time
# Serialize model to JSON
model_json = model.to_json()
with open("output/neural_networks/model.json", "w") as json_file:
json_file.write(model_json)
# Serialize weights to HDF5
model.save_weights("output/neural_networks/model.h5")
print("Saved model structure and weights")
```
### Load model
```
# The model weights and architecture were saved separated, so it must re-compile
# Load json and create model
json_file = open('output/neural_networks/model.json', 'r')
loaded_model_json = json_file.read()
json_file.close()
loaded_model = model_from_json(loaded_model_json)
# Load weights into new model
loaded_model.load_weights("output/neural_networks/model.h5")
print("Loaded model from disk")
# Evaluate loaded model on test data
loaded_model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
score = loaded_model.evaluate(X_train, Y_train, verbose=0)
print("%s: %.2f%%" % (loaded_model.metrics_names[1], score[1]*100))
```
### Summary
```
# Summarize model.
model.summary()
```
### Plot model
```
# Error, BUG, MUST FIX
# plot_model(model, to_file='model_plot.png', show_shapes=True, show_layer_names=True)
```
| github_jupyter |
## Our Mission ##
Spam detection is one of the major applications of Machine Learning in the interwebs today. Pretty much all of the major email service providers have spam detection systems built in and automatically classify such mail as 'Junk Mail'.
In this mission we will be using the Naive Bayes algorithm to create a model that can classify SMS messages as spam or not spam, based on the training we give to the model. It is important to have some level of intuition as to what a spammy text message might look like. Often they have words like 'free', 'win', 'winner', 'cash', 'prize' and the like in them as these texts are designed to catch your eye and in some sense tempt you to open them. Also, spam messages tend to have words written in all capitals and also tend to use a lot of exclamation marks. To the human recipient, it is usually pretty straightforward to identify a spam text and our objective here is to train a model to do that for us!
Being able to identify spam messages is a binary classification problem as messages are classified as either 'Spam' or 'Not Spam' and nothing else. Also, this is a supervised learning problem, as we will be feeding a labelled dataset into the model, that it can learn from, to make future predictions.
# Overview
This project has been broken down in to the following steps:
- Step 0: Introduction to the Naive Bayes Theorem
- Step 1.1: Understanding our dataset
- Step 1.2: Data Preprocessing
- Step 2.1: Bag of Words (BoW)
- Step 2.2: Implementing BoW from scratch
- Step 2.3: Implementing Bag of Words in scikit-learn
- Step 3.1: Training and testing sets
- Step 3.2: Applying Bag of Words processing to our dataset.
- Step 4.1: Bayes Theorem implementation from scratch
- Step 4.2: Naive Bayes implementation from scratch
- Step 5: Naive Bayes implementation using scikit-learn
- Step 6: Evaluating our model
- Step 7: Conclusion
**Note**: If you need help with a step, you can find the solution notebook by clicking on the Jupyter logo in the top left of the notebook.
### Step 0: Introduction to the Naive Bayes Theorem ###
Bayes Theorem is one of the earliest probabilistic inference algorithms. It was developed by Reverend Bayes (which he used to try and infer the existence of God no less), and still performs extremely well for certain use cases.
It's best to understand this theorem using an example. Let's say you are a member of the Secret Service and you have been deployed to protect the Democratic presidential nominee during one of his/her campaign speeches. Being a public event that is open to all, your job is not easy and you have to be on the constant lookout for threats. So one place to start is to put a certain threat-factor for each person. So based on the features of an individual, like age, whether the person is carrying a bag, looks nervous, etc., you can make a judgment call as to whether that person is a viable threat.
If an individual ticks all the boxes up to a level where it crosses a threshold of doubt in your mind, you can take action and remove that person from the vicinity. Bayes Theorem works in the same way, as we are computing the probability of an event (a person being a threat) based on the probabilities of certain related events (age, presence of bag or not, nervousness of the person, etc.).
One thing to consider is the independence of these features amongst each other. For example if a child looks nervous at the event then the likelihood of that person being a threat is not as much as say if it was a grown man who was nervous. To break this down a bit further, here there are two features we are considering, age AND nervousness. Say we look at these features individually, we could design a model that flags ALL persons that are nervous as potential threats. However, it is likely that we will have a lot of false positives as there is a strong chance that minors present at the event will be nervous. Hence by considering the age of a person along with the 'nervousness' feature we would definitely get a more accurate result as to who are potential threats and who aren't.
This is the 'Naive' bit of the theorem where it considers each feature to be independent of each other which may not always be the case and hence that can affect the final judgement.
In short, Bayes Theorem calculates the probability of a certain event happening (in our case, a message being spam) based on the joint probabilistic distributions of certain other events (in our case, the appearance of certain words in a message). We will dive into the workings of Bayes Theorem later in the mission, but first, let us understand the data we are going to work with.
### Step 1.1: Understanding our dataset ###
We will be using a dataset originally compiled and posted on the UCI Machine Learning repository which has a very good collection of datasets for experimental research purposes. If you're interested, you can review the [abstract](https://archive.ics.uci.edu/ml/datasets/SMS+Spam+Collection) and the original [compressed data file](https://archive.ics.uci.edu/ml/machine-learning-databases/00228/) on the UCI site. For this exercise, however, we've gone ahead and downloaded the data for you.
**Here's a preview of the data:**
<img src="images/dqnb.png" height="1242" width="1242">
The columns in the data set are currently not named and as you can see, there are 2 columns.
The first column takes two values, 'ham' which signifies that the message is not spam, and 'spam' which signifies that the message is spam.
The second column is the text content of the SMS message that is being classified.
>**Instructions:**
* Import the dataset into a pandas dataframe using the **read_table** method. The file has already been downloaded, and you can access it using the filepath 'smsspamcollection/SMSSpamCollection'. Because this is a tab separated dataset we will be using '\\t' as the value for the 'sep' argument which specifies this format.
* Also, rename the column names by specifying a list ['label', 'sms_message'] to the 'names' argument of read_table().
* Print the first five values of the dataframe with the new column names.
```
# '!' allows you to run bash commands from jupyter notebook.
print("List all the files in the current directory\n")
!ls
# The required data table can be found under smsspamcollection/SMSSpamCollection
print("\n List all the files inside the smsspamcollection directory\n")
!ls smsspamcollection
import pandas as pd
# Dataset available using filepath 'smsspamcollection/SMSSpamCollection'
df = #TODO
# Output printing out first 5 rows
df.head()
```
### Step 1.2: Data Preprocessing ###
Now that we have a basic understanding of what our dataset looks like, let's convert our labels to binary variables, 0 to represent 'ham'(i.e. not spam) and 1 to represent 'spam' for ease of computation.
You might be wondering why do we need to do this step? The answer to this lies in how scikit-learn handles inputs. Scikit-learn only deals with numerical values and hence if we were to leave our label values as strings, scikit-learn would do the conversion internally(more specifically, the string labels will be cast to unknown float values).
Our model would still be able to make predictions if we left our labels as strings but we could have issues later when calculating performance metrics, for example when calculating our precision and recall scores. Hence, to avoid unexpected 'gotchas' later, it is good practice to have our categorical values be fed into our model as integers.
>**Instructions:**
* Convert the values in the 'label' column to numerical values using map method as follows:
{'ham':0, 'spam':1} This maps the 'ham' value to 0 and the 'spam' value to 1.
* Also, to get an idea of the size of the dataset we are dealing with, print out number of rows and columns using
'shape'.
```
'''
Solution
'''
df['label'] = df.label.map(# TODO))
```
### Step 2.1: Bag of Words ###
What we have here in our data set is a large collection of text data (5,572 rows of data). Most ML algorithms rely on numerical data to be fed into them as input, and email/sms messages are usually text heavy.
Here we'd like to introduce the Bag of Words (BoW) concept which is a term used to specify the problems that have a 'bag of words' or a collection of text data that needs to be worked with. The basic idea of BoW is to take a piece of text and count the frequency of the words in that text. It is important to note that the BoW concept treats each word individually and the order in which the words occur does not matter.
Using a process which we will go through now, we can convert a collection of documents to a matrix, with each document being a row and each word (token) being the column, and the corresponding (row, column) values being the frequency of occurrence of each word or token in that document.
For example:
Let's say we have 4 documents, which are text messages
in our case, as follows:
`['Hello, how are you!',
'Win money, win from home.',
'Call me now',
'Hello, Call you tomorrow?']`
Our objective here is to convert this set of texts to a frequency distribution matrix, as follows:
<img src="images/countvectorizer.png" height="542" width="542">
Here as we can see, the documents are numbered in the rows, and each word is a column name, with the corresponding value being the frequency of that word in the document.
Let's break this down and see how we can do this conversion using a small set of documents.
To handle this, we will be using sklearn's
[count vectorizer](http://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.CountVectorizer.html#sklearn.feature_extraction.text.CountVectorizer) method which does the following:
* It tokenizes the string (separates the string into individual words) and gives an integer ID to each token.
* It counts the occurrence of each of those tokens.
**Please Note:**
* The CountVectorizer method automatically converts all tokenized words to their lower case form so that it does not treat words like 'He' and 'he' differently. It does this using the `lowercase` parameter which is by default set to `True`.
* It also ignores all punctuation so that words followed by a punctuation mark (for example: 'hello!') are not treated differently than the same words not prefixed or suffixed by a punctuation mark (for example: 'hello'). It does this using the `token_pattern` parameter which has a default regular expression which selects tokens of 2 or more alphanumeric characters.
* The third parameter to take note of is the `stop_words` parameter. Stop words refer to the most commonly used words in a language. They include words like 'am', 'an', 'and', 'the', etc. By setting this parameter value to `english`, CountVectorizer will automatically ignore all words (from our input text) that are found in the built in list of English stop words in scikit-learn. This is extremely helpful as stop words can skew our calculations when we are trying to find certain key words that are indicative of spam.
We will dive into the application of each of these into our model in a later step, but for now it is important to be aware of such preprocessing techniques available to us when dealing with textual data.
### Step 2.2: Implementing Bag of Words from scratch ###
Before we dive into scikit-learn's Bag of Words (BoW) library to do the dirty work for us, let's implement it ourselves first so that we can understand what's happening behind the scenes.
**Step 1: Convert all strings to their lower case form.**
Let's say we have a document set:
```
documents = ['Hello, how are you!',
'Win money, win from home.',
'Call me now.',
'Hello, Call hello you tomorrow?']
```
>>**Instructions:**
* Convert all the strings in the documents set to their lower case. Save them into a list called 'lower_case_documents'. You can convert strings to their lower case in python by using the lower() method.
```
'''
Solution:
'''
documents = ['Hello, how are you!',
'Win money, win from home.',
'Call me now.',
'Hello, Call hello you tomorrow?']
lower_case_documents = []
for i in documents:
# TODO
print(lower_case_documents)
```
**Step 2: Removing all punctuation**
>>**Instructions:**
Remove all punctuation from the strings in the document set. Save the strings into a list called
'sans_punctuation_documents'.
```
'''
Solution:
'''
sans_punctuation_documents = []
import string
for i in lower_case_documents:
# TODO
print(sans_punctuation_documents)
```
**Step 3: Tokenization**
Tokenizing a sentence in a document set means splitting up the sentence into individual words using a delimiter. The delimiter specifies what character we will use to identify the beginning and end of a word. Most commonly, we use a single space as the delimiter character for identifying words, and this is true in our documents in this case also.
>>**Instructions:**
Tokenize the strings stored in 'sans_punctuation_documents' using the split() method. Store the final document set
in a list called 'preprocessed_documents'.
```
'''
Solution:
'''
preprocessed_documents = []
for i in sans_punctuation_documents:
# TODO
print(preprocessed_documents)
```
**Step 4: Count frequencies**
Now that we have our document set in the required format, we can proceed to counting the occurrence of each word in each document of the document set. We will use the `Counter` method from the Python `collections` library for this purpose.
`Counter` counts the occurrence of each item in the list and returns a dictionary with the key as the item being counted and the corresponding value being the count of that item in the list.
>>**Instructions:**
Using the Counter() method and preprocessed_documents as the input, create a dictionary with the keys being each word in each document and the corresponding values being the frequency of occurrence of that word. Save each Counter dictionary as an item in a list called 'frequency_list'.
```
'''
Solution
'''
frequency_list = []
import pprint
from collections import Counter
for i in preprocessed_documents:
#TODO
pprint.pprint(frequency_list)
```
Congratulations! You have implemented the Bag of Words process from scratch! As we can see in our previous output, we have a frequency distribution dictionary which gives a clear view of the text that we are dealing with.
We should now have a solid understanding of what is happening behind the scenes in the `sklearn.feature_extraction.text.CountVectorizer` method of scikit-learn.
We will now implement `sklearn.feature_extraction.text.CountVectorizer` method in the next step.
### Step 2.3: Implementing Bag of Words in scikit-learn ###
Now that we have implemented the BoW concept from scratch, let's go ahead and use scikit-learn to do this process in a clean and succinct way. We will use the same document set as we used in the previous step.
```
'''
Here we will look to create a frequency matrix on a smaller document set to make sure we understand how the
document-term matrix generation happens. We have created a sample document set 'documents'.
'''
documents = ['Hello, how are you!',
'Win money, win from home.',
'Call me now.',
'Hello, Call hello you tomorrow?']
```
>>**Instructions:**
Import the sklearn.feature_extraction.text.CountVectorizer method and create an instance of it called 'count_vector'.
```
'''
Solution
'''
from sklearn.feature_extraction.text import CountVectorizer
count_vector = # TODO
```
**Data preprocessing with CountVectorizer()**
In Step 2.2, we implemented a version of the CountVectorizer() method from scratch that entailed cleaning our data first. This cleaning involved converting all of our data to lower case and removing all punctuation marks. CountVectorizer() has certain parameters which take care of these steps for us. They are:
* `lowercase = True`
The `lowercase` parameter has a default value of `True` which converts all of our text to its lower case form.
* `token_pattern = (?u)\\b\\w\\w+\\b`
The `token_pattern` parameter has a default regular expression value of `(?u)\\b\\w\\w+\\b` which ignores all punctuation marks and treats them as delimiters, while accepting alphanumeric strings of length greater than or equal to 2, as individual tokens or words.
* `stop_words`
The `stop_words` parameter, if set to `english` will remove all words from our document set that match a list of English stop words defined in scikit-learn. Considering the small size of our dataset and the fact that we are dealing with SMS messages and not larger text sources like e-mail, we will not use stop words, and we won't be setting this parameter value.
You can take a look at all the parameter values of your `count_vector` object by simply printing out the object as follows:
```
'''
Practice node:
Print the 'count_vector' object which is an instance of 'CountVectorizer()'
'''
# No need to revise this code
print(count_vector)
```
>>**Instructions:**
Fit your document dataset to the CountVectorizer object you have created using fit(), and get the list of words
which have been categorized as features using the get_feature_names() method.
```
'''
Solution:
'''
# No need to revise this code
count_vector.fit(documents)
count_vector.get_feature_names()
```
The `get_feature_names()` method returns our feature names for this dataset, which is the set of words that make up our vocabulary for 'documents'.
>>**Instructions:**
Create a matrix with each row representing one of the 4 documents, and each column representing a word (feature name).
Each value in the matrix will represent the frequency of the word in that column occurring in the particular document in that row.
You can do this using the transform() method of CountVectorizer, passing in the document data set as the argument. The transform() method returns a matrix of NumPy integers, which you can convert to an array using
toarray(). Call the array 'doc_array'.
```
'''
Solution
'''
doc_array = # TODO
doc_array
```
Now we have a clean representation of the documents in terms of the frequency distribution of the words in them. To make it easier to understand our next step is to convert this array into a dataframe and name the columns appropriately.
>>**Instructions:**
Convert the 'doc_array' we created into a dataframe, with the column names as the words (feature names). Call the dataframe 'frequency_matrix'.
```
'''
Solution
'''
frequency_matrix = pd.DataFrame(# TODO))
frequency_matrix
```
Congratulations! You have successfully implemented a Bag of Words problem for a document dataset that we created.
One potential issue that can arise from using this method is that if our dataset of text is extremely large (say if we have a large collection of news articles or email data), there will be certain values that are more common than others simply due to the structure of the language itself. For example, words like 'is', 'the', 'an', pronouns, grammatical constructs, etc., could skew our matrix and affect our analyis.
There are a couple of ways to mitigate this. One way is to use the `stop_words` parameter and set its value to `english`. This will automatically ignore all the words in our input text that are found in a built-in list of English stop words in scikit-learn.
Another way of mitigating this is by using the [tfidf](http://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.TfidfVectorizer.html#sklearn.feature_extraction.text.TfidfVectorizer) method. This method is out of scope for the context of this lesson.
### Step 3.1: Training and testing sets ###
Now that we understand how to use the Bag of Words approach, we can return to our original, larger UCI dataset and proceed with our analysis. Our first step is to split our dataset into a training set and a testing set so we can first train, and then test our model.
>>**Instructions:**
Split the dataset into a training and testing set using the train_test_split method in sklearn, and print out the number of rows we have in each of our training and testing data. Split the data
using the following variables:
* `X_train` is our training data for the 'sms_message' column.
* `y_train` is our training data for the 'label' column
* `X_test` is our testing data for the 'sms_message' column.
* `y_test` is our testing data for the 'label' column.
```
'''
Solution
'''
# split into training and testing sets
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(df['sms_message'],
df['label'],
random_state=1)
print('Number of rows in the total set: {}'.format(df.shape[0]))
print('Number of rows in the training set: {}'.format(X_train.shape[0]))
print('Number of rows in the test set: {}'.format(X_test.shape[0]))
```
### Step 3.2: Applying Bag of Words processing to our dataset. ###
Now that we have split the data, our next objective is to follow the steps from "Step 2: Bag of Words," and convert our data into the desired matrix format. To do this we will be using CountVectorizer() as we did before. There are two steps to consider here:
* First, we have to fit our training data (`X_train`) into `CountVectorizer()` and return the matrix.
* Secondly, we have to transform our testing data (`X_test`) to return the matrix.
Note that `X_train` is our training data for the 'sms_message' column in our dataset and we will be using this to train our model.
`X_test` is our testing data for the 'sms_message' column and this is the data we will be using (after transformation to a matrix) to make predictions on. We will then compare those predictions with `y_test` in a later step.
For now, we have provided the code that does the matrix transformations for you!
```
'''
[Practice Node]
The code for this segment is in 2 parts. First, we are learning a vocabulary dictionary for the training data
and then transforming the data into a document-term matrix; secondly, for the testing data we are only
transforming the data into a document-term matrix.
This is similar to the process we followed in Step 2.3.
We will provide the transformed data to students in the variables 'training_data' and 'testing_data'.
'''
'''
Solution
'''
# Instantiate the CountVectorizer method
count_vector = CountVectorizer()
# Fit the training data and then return the matrix
training_data = count_vector.fit_transform(X_train)
# Transform testing data and return the matrix. Note we are not fitting the testing data into the CountVectorizer()
testing_data = count_vector.transform(X_test)
```
### Step 4.1: Bayes Theorem implementation from scratch ###
Now that we have our dataset in the format that we need, we can move onto the next portion of our mission which is the algorithm we will use to make our predictions to classify a message as spam or not spam. Remember that at the start of the mission we briefly discussed the Bayes theorem but now we shall go into a little more detail. In layman's terms, the Bayes theorem calculates the probability of an event occurring, based on certain other probabilities that are related to the event in question. It is composed of "prior probabilities" - or just "priors." These "priors" are the probabilities that we are aware of, or that are given to us. And Bayes theorem is also composed of the "posterior probabilities," or just "posteriors," which are the probabilities we are looking to compute using the "priors".
Let us implement the Bayes Theorem from scratch using a simple example. Let's say we are trying to find the odds of an individual having diabetes, given that he or she was tested for it and got a positive result.
In the medical field, such probabilities play a very important role as they often deal with life and death situations.
We assume the following:
`P(D)` is the probability of a person having Diabetes. Its value is `0.01`, or in other words, 1% of the general population has diabetes (disclaimer: these values are assumptions and are not reflective of any actual medical study).
`P(Pos)` is the probability of getting a positive test result.
`P(Neg)` is the probability of getting a negative test result.
`P(Pos|D)` is the probability of getting a positive result on a test done for detecting diabetes, given that you have diabetes. This has a value `0.9`. In other words the test is correct 90% of the time. This is also called the Sensitivity or True Positive Rate.
`P(Neg|~D)` is the probability of getting a negative result on a test done for detecting diabetes, given that you do not have diabetes. This also has a value of `0.9` and is therefore correct, 90% of the time. This is also called the Specificity or True Negative Rate.
The Bayes formula is as follows:
<img src="images/bayes_formula.png" height="242" width="242">
* `P(A)` is the prior probability of A occurring independently. In our example this is `P(D)`. This value is given to us.
* `P(B)` is the prior probability of B occurring independently. In our example this is `P(Pos)`.
* `P(A|B)` is the posterior probability that A occurs given B. In our example this is `P(D|Pos)`. That is, **the probability of an individual having diabetes, given that this individual got a positive test result. This is the value that we are looking to calculate.**
* `P(B|A)` is the prior probability of B occurring, given A. In our example this is `P(Pos|D)`. This value is given to us.
Putting our values into the formula for Bayes theorem we get:
`P(D|Pos) = P(D) * P(Pos|D) / P(Pos)`
The probability of getting a positive test result `P(Pos)` can be calculated using the Sensitivity and Specificity as follows:
`P(Pos) = [P(D) * Sensitivity] + [P(~D) * (1-Specificity))]`
```
'''
Instructions:
Calculate probability of getting a positive test result, P(Pos)
'''
'''
Solution (skeleton code will be provided)
'''
# P(D)
p_diabetes = 0.01
# P(~D)
p_no_diabetes = 0.99
# Sensitivity or P(Pos|D)
p_pos_diabetes = 0.9
# Specificity or P(Neg|~D)
p_neg_no_diabetes = 0.9
# P(Pos)
p_pos = # TODO
print('The probability of getting a positive test result P(Pos) is: {}',format(p_pos))
```
**Using all of this information we can calculate our posteriors as follows:**
The probability of an individual having diabetes, given that, that individual got a positive test result:
`P(D|Pos) = (P(D) * Sensitivity)) / P(Pos)`
The probability of an individual not having diabetes, given that, that individual got a positive test result:
`P(~D|Pos) = (P(~D) * (1-Specificity)) / P(Pos)`
The sum of our posteriors will always equal `1`.
```
'''
Instructions:
Compute the probability of an individual having diabetes, given that, that individual got a positive test result.
In other words, compute P(D|Pos).
The formula is: P(D|Pos) = (P(D) * P(Pos|D) / P(Pos)
'''
'''
Solution
'''
# P(D|Pos)
p_diabetes_pos = # TODO
print('Probability of an individual having diabetes, given that that individual got a positive test result is:\
',format(p_diabetes_pos))
'''
Instructions:
Compute the probability of an individual not having diabetes, given that, that individual got a positive test result.
In other words, compute P(~D|Pos).
The formula is: P(~D|Pos) = P(~D) * P(Pos|~D) / P(Pos)
Note that P(Pos|~D) can be computed as 1 - P(Neg|~D).
Therefore:
P(Pos|~D) = p_pos_no_diabetes = 1 - 0.9 = 0.1
'''
'''
Solution
'''
# P(Pos|~D)
p_pos_no_diabetes = 0.1
# P(~D|Pos)
p_no_diabetes_pos = # TODO
print 'Probability of an individual not having diabetes, given that that individual got a positive test result is:'\
,p_no_diabetes_pos
```
Congratulations! You have implemented Bayes Theorem from scratch. Your analysis shows that even if you get a positive test result, there is only an 8.3% chance that you actually have diabetes and a 91.67% chance that you do not have diabetes. This is of course assuming that only 1% of the entire population has diabetes which is only an assumption.
**What does the term 'Naive' in 'Naive Bayes' mean ?**
The term 'Naive' in Naive Bayes comes from the fact that the algorithm considers the features that it is using to make the predictions to be independent of each other, which may not always be the case. So in our Diabetes example, we are considering only one feature, that is the test result. Say we added another feature, 'exercise'. Let's say this feature has a binary value of `0` and `1`, where the former signifies that the individual exercises less than or equal to 2 days a week and the latter signifies that the individual exercises greater than or equal to 3 days a week. If we had to use both of these features, namely the test result and the value of the 'exercise' feature, to compute our final probabilities, Bayes' theorem would fail. Naive Bayes' is an extension of Bayes' theorem that assumes that all the features are independent of each other.
### Step 4.2: Naive Bayes implementation from scratch ###
Now that you have understood the ins and outs of Bayes Theorem, we will extend it to consider cases where we have more than one feature.
Let's say that we have two political parties' candidates, 'Jill Stein' of the Green Party and 'Gary Johnson' of the Libertarian Party and we have the probabilities of each of these candidates saying the words 'freedom', 'immigration' and 'environment' when they give a speech:
* Probability that Jill Stein says 'freedom': 0.1 ---------> `P(F|J)`
* Probability that Jill Stein says 'immigration': 0.1 -----> `P(I|J)`
* Probability that Jill Stein says 'environment': 0.8 -----> `P(E|J)`
* Probability that Gary Johnson says 'freedom': 0.7 -------> `P(F|G)`
* Probability that Gary Johnson says 'immigration': 0.2 ---> `P(I|G)`
* Probability that Gary Johnson says 'environment': 0.1 ---> `P(E|G)`
And let us also assume that the probability of Jill Stein giving a speech, `P(J)` is `0.5` and the same for Gary Johnson, `P(G) = 0.5`.
Given this, what if we had to find the probabilities of Jill Stein saying the words 'freedom' and 'immigration'? This is where the Naive Bayes' theorem comes into play as we are considering two features, 'freedom' and 'immigration'.
Now we are at a place where we can define the formula for the Naive Bayes' theorem:
<img src="images/naivebayes.png" height="342" width="342">
Here, `y` is the class variable (in our case the name of the candidate) and `x1` through `xn` are the feature vectors (in our case the individual words). The theorem makes the assumption that each of the feature vectors or words (`xi`) are independent of each other.
To break this down, we have to compute the following posterior probabilities:
* `P(J|F,I)`: Given the words 'freedom' and 'immigration' were said, what's the probability they were said by Jill?
Using the formula and our knowledge of Bayes' theorem, we can compute this as follows: `P(J|F,I)` = `(P(J) * P(F|J) * P(I|J)) / P(F,I)`. Here `P(F,I)` is the probability of the words 'freedom' and 'immigration' being said in a speech.
* `P(G|F,I)`: Given the words 'freedom' and 'immigration' were said, what's the probability they were said by Gary?
Using the formula, we can compute this as follows: `P(G|F,I)` = `(P(G) * P(F|G) * P(I|G)) / P(F,I)`
```
'''
Instructions: Compute the probability of the words 'freedom' and 'immigration' being said in a speech, or
P(F,I).
The first step is multiplying the probabilities of Jill Stein giving a speech with her individual
probabilities of saying the words 'freedom' and 'immigration'. Store this in a variable called p_j_text.
The second step is multiplying the probabilities of Gary Johnson giving a speech with his individual
probabilities of saying the words 'freedom' and 'immigration'. Store this in a variable called p_g_text.
The third step is to add both of these probabilities and you will get P(F,I).
'''
'''
Solution: Step 1
'''
# P(J)
p_j = 0.5
# P(F/J)
p_j_f = 0.1
# P(I/J)
p_j_i = 0.1
p_j_text = # TODO
print(p_j_text)
'''
Solution: Step 2
'''
# P(G)
p_g = 0.5
# P(F/G)
p_g_f = 0.7
# P(I/G)
p_g_i = 0.2
p_g_text = # TODO
print(p_g_text)
'''
Solution: Step 3: Compute P(F,I) and store in p_f_i
'''
p_f_i = # TODO
print('Probability of words freedom and immigration being said are: ', format(p_f_i))
```
Now we can compute the probability of `P(J|F,I)`, the probability of Jill Stein saying the words 'freedom' and 'immigration' and `P(G|F,I)`, the probability of Gary Johnson saying the words 'freedom' and 'immigration'.
```
'''
Instructions:
Compute P(J|F,I) using the formula P(J|F,I) = (P(J) * P(F|J) * P(I|J)) / P(F,I) and store it in a variable p_j_fi
'''
'''
Solution
'''
p_j_fi = # TODO
print('The probability of Jill Stein saying the words Freedom and Immigration: ', format(p_j_fi))
'''
Instructions:
Compute P(G|F,I) using the formula P(G|F,I) = (P(G) * P(F|G) * P(I|G)) / P(F,I) and store it in a variable p_g_fi
'''
'''
Solution
'''
p_g_fi = # TODO
print('The probability of Gary Johnson saying the words Freedom and Immigration: ', format(p_g_fi))
```
And as we can see, just like in the Bayes' theorem case, the sum of our posteriors is equal to 1.
Congratulations! You have implemented the Naive Bayes' theorem from scratch. Our analysis shows that there is only a 6.6% chance that Jill Stein of the Green Party uses the words 'freedom' and 'immigration' in her speech as compared with the 93.3% chance for Gary Johnson of the Libertarian party.
For another example of Naive Bayes, let's consider searching for images using the term 'Sacramento Kings' in a search engine. In order for us to get the results pertaining to the Scramento Kings NBA basketball team, the search engine needs to be able to associate the two words together and not treat them individually. If the search engine only searched for the words individually, we would get results of images tagged with 'Sacramento,' like pictures of city landscapes, and images of 'Kings,' which might be pictures of crowns or kings from history. But associating the two terms together would produce images of the basketball team. In the first approach we would treat the words as independent entities, so it would be considered 'naive.' We don't usually want this approach from a search engine, but it can be extremely useful in other cases.
Applying this to our problem of classifying messages as spam, the Naive Bayes algorithm *looks at each word individually and not as associated entities* with any kind of link between them. In the case of spam detectors, this usually works, as there are certain red flag words in an email which are highly reliable in classifying it as spam. For example, emails with words like 'viagra' are usually classified as spam.
### Step 5: Naive Bayes implementation using scikit-learn ###
Now let's return to our spam classification context. Thankfully, sklearn has several Naive Bayes implementations that we can use, so we do not have to do the math from scratch. We will be using sklearn's `sklearn.naive_bayes` method to make predictions on our SMS messages dataset.
Specifically, we will be using the multinomial Naive Bayes algorithm. This particular classifier is suitable for classification with discrete features (such as in our case, word counts for text classification). It takes in integer word counts as its input. On the other hand, Gaussian Naive Bayes is better suited for continuous data as it assumes that the input data has a Gaussian (normal) distribution.
```
'''
Instructions:
We have loaded the training data into the variable 'training_data' and the testing data into the
variable 'testing_data'.
Import the MultinomialNB classifier and fit the training data into the classifier using fit(). Name your classifier
'naive_bayes'. You will be training the classifier using 'training_data' and 'y_train' from our split earlier.
'''
'''
Solution
'''
from sklearn.naive_bayes import MultinomialNB
naive_bayes = # TODO
naive_bayes.fit(# TODO)
'''
Instructions:
Now that our algorithm has been trained using the training data set we can now make some predictions on the test data
stored in 'testing_data' using predict(). Save your predictions into the 'predictions' variable.
'''
'''
Solution
'''
predictions = naive_bayes.predict(# TODO)
```
Now that predictions have been made on our test set, we need to check the accuracy of our predictions.
### Step 6: Evaluating our model ###
Now that we have made predictions on our test set, our next goal is to evaluate how well our model is doing. There are various mechanisms for doing so, so first let's review them.
**Accuracy** measures how often the classifier makes the correct prediction. It’s the ratio of the number of correct predictions to the total number of predictions (the number of test data points).
**Precision** tells us what proportion of messages we classified as spam, actually were spam.
It is a ratio of true positives (words classified as spam, and which actually are spam) to all positives (all words classified as spam, regardless of whether that was the correct classification). In other words, precision is the ratio of
`[True Positives/(True Positives + False Positives)]`
**Recall (sensitivity)** tells us what proportion of messages that actually were spam were classified by us as spam.
It is a ratio of true positives (words classified as spam, and which actually are spam) to all the words that were actually spam. In other words, recall is the ratio of
`[True Positives/(True Positives + False Negatives)]`
For classification problems that are skewed in their classification distributions like in our case - for example if we had 100 text messages and only 2 were spam and the other 98 weren't - accuracy by itself is not a very good metric. We could classify 90 messages as not spam (including the 2 that were spam but we classify them as not spam, hence they would be false negatives) and 10 as spam (all 10 false positives) and still get a reasonably good accuracy score. For such cases, precision and recall come in very handy. These two metrics can be combined to get the **F1 score**, which is the weighted average of the precision and recall scores. This score can range from 0 to 1, with 1 being the best possible F1 score.
We will be using all 4 of these metrics to make sure our model does well. For all 4 metrics whose values can range from 0 to 1, having a score as close to 1 as possible is a good indicator of how well our model is doing.
```
'''
Instructions:
Compute the accuracy, precision, recall and F1 scores of your model using your test data 'y_test' and the predictions
you made earlier stored in the 'predictions' variable.
'''
'''
Solution
'''
from sklearn.metrics import accuracy_score, precision_score, recall_score, f1_score
print('Accuracy score: ', format(accuracy_score(# TODO)))
print('Precision score: ', format(precision_score(# TODO)))
print('Recall score: ', format(recall_score(# TODO)))
print('F1 score: ', format(f1_score(# TODO)))
```
### Step 7: Conclusion ###
One of the major advantages that Naive Bayes has over other classification algorithms is its ability to handle an extremely large number of features. In our case, each word is treated as a feature and there are thousands of different words. Also, it performs well even with the presence of irrelevant features and is relatively unaffected by them. The other major advantage it has is its relative simplicity. Naive Bayes' works well right out of the box and tuning its parameters is rarely ever necessary, except usually in cases where the distribution of the data is known.
It rarely ever overfits the data. Another important advantage is that its model training and prediction times are very fast for the amount of data it can handle. All in all, Naive Bayes' really is a gem of an algorithm!
Congratulations! You have successfully designed a model that can efficiently predict if an SMS message is spam or not!
Thank you for learning with us!
| github_jupyter |
## Slug Test for Confined Aquifer
**This test is taken from examples of AQTESOLV.**
```
%matplotlib inline
from ttim import *
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
```
Set background parameters:
```
H0 = 2.798 #initial displacement in m
b = -6.1 #aquifer thickness
rw1 = 0.102 #well radius of Ln-2 Well
rw2 = 0.071 #well radius of observation Ln-3 Well
rc1 = 0.051 #casing radius of Ln-2 Well
rc2 = 0.025 #casing radius of Ln-3 Well
r = 6.45 #distance from observation well to test well
```
Slug:
```
Q = np.pi * rc1 ** 2 * H0
print('Slug:', round(Q, 5), 'm^3')
```
Load data:
```
data1 = np.loadtxt('data/ln-2.txt')
t1 = data1[:, 0] / 60 / 60 / 24 #convert time from seconds to days
h1 = data1[:, 1]
data2 = np.loadtxt('data/ln-3.txt')
t2 = data2[:, 0] / 60 / 60 / 24
h2 = data2[:, 1]
```
Create single layer conceptual model:
```
ml_0 = ModelMaq(kaq=10, z=[0, b], Saq=1e-4, \
tmin=1e-5, tmax=0.01)
w_0 = Well(ml_0, xw=0, yw=0, rw=rw1, rc=rc1, tsandQ=[(0, -Q)], layers=0, wbstype='slug')
ml_0.solve()
```
Calibrate with two datasets simultaneously:
```
#unknown parameters: kaq, Saq
ca_0 = Calibrate(ml_0)
ca_0.set_parameter(name='kaq0', initial=10)
ca_0.set_parameter(name='Saq0', initial=1e-4)
ca_0.series(name='Ln-2', x=0, y=0, layer=0, t=t1, h=h1)
ca_0.series(name='Ln-3', x=r, y=0, layer=0, t=t2, h=h2)
ca_0.fit(report=True)
display(ca_0.parameters)
print('RMSE:', ca_0.rmse())
hm1_0 = ml_0.head(0, 0, t1, layers=0)
hm2_0 = ml_0.head(r, 0, t2, layers=0)
plt.figure(figsize=(8, 5))
plt.semilogx(t1, h1/H0, '.', label='obs ln-2')
plt.semilogx(t1, hm1_0[0]/H0, label='ttim ln-2')
plt.semilogx(t2, h2/H0, '.', label='obs ln-3')
plt.semilogx(t2, hm2_0[0]/H0, label='ttim ln-3')
plt.xlabel('time(d)')
plt.ylabel('h/H0')
plt.legend();
```
Try adding well skin resistance res:
```
ml_1 = ModelMaq(kaq=10, z=[0, b], Saq=1e-4, \
tmin=1e-5, tmax=0.01)
w_1 = Well(ml_1, xw=0, yw=0, rw=rw1, res=0, rc=rc1, tsandQ=[(0, -Q)], layers=0, wbstype='slug')
ml_1.solve()
#unknown parameters: kaq, Saq, res
ca_1 = Calibrate(ml_1)
ca_1.set_parameter(name='kaq0', initial=10)
ca_1.set_parameter(name='Saq0', initial=1e-4)
ca_1.set_parameter_by_reference(name='res', parameter=w_1.res, initial=0)
ca_1.series(name='Ln-2', x=0, y=0, layer=0, t=t1, h=h1)
ca_1.series(name='Ln-3', x=r, y=0, layer=0, t=t2, h=h2)
ca_1.fit(report=True)
display(ca_1.parameters)
print('RMSE:', ca_1.rmse())
hm1_1 = ml_1.head(0, 0, t1, layers=0)
hm2_1 = ml_1.head(r, 0, t2, layers=0)
plt.figure(figsize=(8, 5))
plt.semilogx(t1, h1/H0, '.', label='obs ln-2')
plt.semilogx(t1, hm1_1[0]/H0, label='ttim ln-2')
plt.semilogx(t2, h2/H0, '.', label='obs ln-3')
plt.semilogx(t2, hm2_1[0]/H0, label='ttim ln-3')
plt.xlabel('time(d)')
plt.ylabel('h/H0')
plt.legend();
```
Adding well screen resistance does not improve the performance obviously. While the AIC value increases. Thus, res should be removed from the model.
Try multilayer conceptual model:
```
#Determine elevations of each layer.
#Thickness of each layer is set to be 0.5 m.
z = np.arange(0, b, -0.5)
zlay = np.append(z, b)
nlay = len(zlay) - 1
Saq_2 = 1e-4 * np.ones(nlay)
n = np.arange(0, 13,1)
ml_2 = Model3D(kaq=10, z=zlay, Saq=Saq_2, kzoverkh=1, tmin=1e-5, tmax=0.01, \
phreatictop=True)
w_2 = Well(ml_2, xw=0, yw=0, rw=rw1, tsandQ=[(0, -Q)], layers=n, rc=rc1, \
wbstype='slug')
ml_2.solve()
```
Calibrate with two datasets simultaneously:
```
ca_2 = Calibrate(ml_2)
ca_2.set_parameter(name='kaq0_12', initial=10)
ca_2.set_parameter(name='Saq0_12', initial=1e-4, pmin=0)
ca_2.series(name='Ln-2', x=0, y=0, layer=n, t=t1, h=h1)
ca_2.series(name='Ln-3', x=r, y=0, layer=n, t=t2, h=h2)
ca_2.fit(report=True)
display(ca_2.parameters)
print('RMSE:', ca_2.rmse())
hm1_2 = ml_2.head(0, 0, t1, layers=n)
hm2_2 = ml_2.head(r, 0, t2, layers=n)
plt.figure(figsize=(8, 5))
plt.semilogx(t1, h1/H0, '.', label='obs ln-2')
plt.semilogx(t1, hm1_2[0]/H0, label='ttim ln-2')
plt.semilogx(t2, h2/H0, '.', label='obs ln-3')
plt.semilogx(t2, hm2_2[0]/H0, label='ttim ln-3')
plt.xlabel('time(d)')
plt.ylabel('h/H0')
plt.legend();
```
## Summary of values presented by AQTESOLV & MLU
```
t = pd.DataFrame(columns=['k [m/d]', 'Ss [1/m]'], \
index=['MLU', 'AQTESOLV', 'ttim-single', 'ttim-multi'])
t.loc['AQTESOLV'] = [1.166, 9.368E-06]
t.loc['MLU'] = [1.311, 8.197E-06]
t.loc['ttim-single'] = ca_0.parameters['optimal'].values
t.loc['ttim-multi'] = ca_2.parameters['optimal'].values
t['RMSE'] = [0.010373, 0.009151, ca_0.rmse(), ca_1.rmse()]
t
```
| github_jupyter |
<h1>Weather on Mars - Next Day Prediction</h1>
<h2>Prediction of temperature based on empirical data collected by Curiosity Mars Rover</h2>
```
runLocallyOrOnDrive = 'Drive' # 'Drive' or 'Local'
#if 'Local' change the paths below to local paths e.g. r'C:/Users/..'
#@title Connect to Google Drive if runLocallyOrOnDrive=='Drive'
if runLocallyOrOnDrive == "Drive":
from google.colab import drive
drive.mount('/content/drive/')
#@title Change these file paths to your respective
#Google drive folder, but it can also be local e.g. on your C-disk
if runLocallyOrOnDrive == 'Local':
loadModelPath = r'/content/../model.json'
saveModelPathH5 = r'/content/../model.h5'
elif runLocallyOrOnDrive == 'Drive':
saveModelPath = r'/content/../model.json'
saveModelPathH5 = r'/content/../model.h5'
#@title Import libraries
# # If you want to loads file, upload to drive and run the following
import urllib, json
# import sys
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
from datetime import timedelta
from keras.layers import Bidirectional,LSTM,Dense,Flatten,Conv1D,MaxPooling1D,Dropout,RepeatVector
from keras.models import Sequential
from keras.callbacks import EarlyStopping,ReduceLROnPlateau
from tensorflow import keras
from sklearn.preprocessing import MinMaxScaler
#@title Functions: Load Data
# returns the (average) weather data for a top level key [sol] from [sol_keys]
def findAtAndPre(dictionary, key):
value = dictionary.get(key)
return {'terrestrial_date': value.get('First_UTC')[:10], 'Low': value.get('AT')['mn'], 'High': value.get('AT')['mx']}
def loadRecentNDaysOfData():
# returns the data of the latest 7 days
apiUrl = 'https://api.nasa.gov/insight_weather/?api_key=DEMO_KEY&feedtype=json&ver=1.0'
jsonApi = json.loads(urllib.request.urlopen(apiUrl).read())
dfApi = pd.DataFrame({key: findAtAndPre(jsonApi, key) for key in jsonApi.get('sol_keys')})
return dfApi
data = loadRecentNDaysOfData().T
data['avg_temp'] = data[['Low', 'High']].mean(axis=1)
data['terrestrial_date']=pd.to_datetime(data['terrestrial_date'])
df_avg_temp=pd.DataFrame(list(data['avg_temp']), index=data['terrestrial_date'], columns=['temp'])
df_avg_temp.fillna(data['avg_temp'].mean(),inplace=True)
data
scaler=MinMaxScaler(feature_range=(-1,1))
scData =scaler.fit_transform(df_avg_temp)
inputs = []
inputs.append(scData)
inputs=np.asanyarray(inputs)
tst_x=inputs
#@title Load Model JSON for testing
#serialize mode to JSON
from tensorflow.keras.models import model_from_json
json_file = open(loadModelPath,"r")
model_json = json_file.read()
json_file.close()
model_fromDisk = model_from_json(model_json)
#load weights into the new model
model_fromDisk.load_weights(loadModelPathH5)
print("loaded from disk")
```
Here is tomorrows average temperature on Mars
```
prediction = scaler.inverse_transform(model_fromDisk.predict(tst_x))
prediction_date = max(pd.to_datetime(data['terrestrial_date']))+ timedelta(days=1)
datalist = [[prediction_date,\
data["Low"].mean(),\
data["High"].mean(),\
prediction[0][0]]]
# Create the pandas DataFrame
df_new = pd.DataFrame(datalist, columns = ['terrestrial_date', 'Low','High','avg_temp'])
df_with_prediction = data.append([df_new],ignore_index=True)
# show data of past 7 days plus prediction day
data_1 = df_with_prediction.set_index('terrestrial_date')
dayRange = data_1.index.to_list()
meanTemperature = data_1['avg_temp']
std = data_1['avg_temp'].std()
lower = (meanTemperature-std).to_list()
upper = (meanTemperature+std).to_list()
high = data_1['High'].to_list()
low = data_1['Low'].to_list()
plt.figure(figsize=(10,4))
plt.plot(prediction_date, prediction,'r*')
plt.plot(dayRange, meanTemperature)
# first fill between min and max temps
plt.fill_between(dayRange, low, high, facecolor='papayawhip')
# then also fill std-range (perhaps from archive data?)
plt.fill_between(dayRange, lower, upper, facecolor='lightblue')
```
| github_jupyter |
```
import json
import codecs
import tensorflow as tf
import collections
with open('/crimea/geeticka/data/relation_extraction/semeval2010/pre-processed/original/bert/train_original_border_50.json') as file:
for line in file.readlines():
data = json.loads(line)
print(data)
break
# because the dumping of the embeddings was done using pytorch, refer to their method of writing to
# figure out how to do the reading
# https://github.com/huggingface/pytorch-pretrained-BERT/blob/master/examples/extract_features.py
# refer to the above to figure out how to read this file
# need to import collections
def write_bert_tokens(input_filename, output_filename):
with open(input_filename, 'r', encoding='utf-8') as input_file, open(output_filename, 'w', encoding='utf-8' as output_file:
for line in input_file.readlines():
if data['features'][0]['token'] != '[CLS]': raise Exception("The first token has to be CLS!")
if data['features'][-1]['token'] != '[SEP]': raise Exception("The last token has to be SEP!")
output_json = collections.OrderedDict()
data = json.loads(line)
output_json['linex_index'] = data['linex_index']
features = data['features'] # basically for all features['token'] that starts with ##, add up the values
# for the respective indexes to put the words back together, ignore [CLS] and [SEP] tokens
new_feature_map = generate_feature_map_without_word_piece(features) # this new feature map needs to be
# called layers because things have now been shuffled.
output_json['layers'] = new_feature_map
output_filename.write(json.dumps(output_json) + "\n")
a = [1]
a.extend([2,3,4])
features = [{'token': '3', 'layers': [{'index': -1, 'values': [-1, -3]},
{'index': -2, 'values': [-4, -2]},
{'index': -3, 'values': [-2, -1]},
{'index': -4, 'values': [-1,2]}]},
{'token': 'a', 'layers': [{'index': -1, 'values': [1,2]},
{'index': -2, 'values': [3,4]},
{'index': -3, 'values': [4,5]},
{'index': -4, 'values': [2,1]}]},
{'token': '##b', 'layers': [{'index': -1, 'values': [-1, -3]},
{'index': -2, 'values': [-2, -1]},
{'index': -3, 'values': [-3, -2]},
{'index': -4, 'values': [-1, -1]}]},
{'token': '3', 'layers': [{'index': -1, 'values': [-1, -3]},
{'index': -2, 'values': [-4, -2]},
{'index': -3, 'values': [-2, -1]},
{'index': -4, 'values': [-1,2]}]},
{'token': '##b', 'layers': [{'index': -1, 'values': [-1, -3]},
{'index': -2, 'values': [-2, -1]},
{'index': -3, 'values': [-3, -2]},
{'index': -4, 'values': [-1, -1]}]},
{'token': '##b', 'layers': [{'index': -1, 'values': [-1, -3]},
{'index': -2, 'values': [-2, -1]},
{'index': -3, 'values': [-3, -2]},
{'index': -4, 'values': [-1, -1]}]},
{'token': '3', 'layers': [{'index': -1, 'values': [-1, -3]},
{'index': -2, 'values': [-4, -2]},
{'index': -3, 'values': [-2, -1]},
{'index': -4, 'values': [-1,2]}]}]
# indexes = [0,1]
# def generate_feature_map_without_word_piece(features):
# # need to double check and see why this is happening
# new_features = []
# i = 0
# while(i < len(features)):
# if features[i]['token'] == '[CLS]' or features[i]['token'] == '[SEP]':
# i += 1
# continue
# captured_indexes = []
# for j in range(i + 1, len(features)):
# if not features[j]['token'].startswith('##'):
# break
# captured_indexes.append(j)
# if len(captured_indexes) == 0:
# new_features.append(features[i])
# i += 1
# continue
# sum_indexes = [i]
# sum_indexes.extend(captured_indexes)
# new_feature = average_over_token_embedding(sum_indexes, features)
# new_features.append(new_feature)
# i = captured_indexes[-1] + 1
# # rewrite in the elmo format as well
# new_features_map = [] # we are converting from the (token, layers) shape to (layers, token) shape
# layer_minus1 = []; layer_minus2 = []; layer_minus3 = []; layer_minus4 = [];
# for token in new_features:
# layer_minus1.append({'token': token['token'], 'features': token['layers'][0]['values']})
# layer_minus2.append({'token': token['token'], 'features': token['layers'][1]['values']})
# layer_minus3.append({'token': token['token'], 'features': token['layers'][2]['values']})
# layer_minus4.append({'token': token['token'], 'features': token['layers'][3]['values']})
# new_features_map.append({'index': -1, 'values': layer_minus1})
# new_features_map.append({'index': -2, 'values': layer_minus2})
# new_features_map.append({'index': -3, 'values': layer_minus3})
# new_features_map.append({'index': -4, 'values': layer_minus4})
# return new_features_map
generate_feature_map_without_word_piece(features)
a = {2: [1,2]}
np.mean([a[2], [3,4]], axis=1)
# def average_over_token_embedding(indexes, features):
# new_feature = collections.OrderedDict()
# new_token = ''
# new_layers = []
# layer_minus_1 = []; layer_minus_2 = []; layer_minus_3 = []; layer_minus_4 = [];
# for index in indexes:
# layer_minus_1.append(features[index]['layers'][0]['values'])
# layer_minus_2.append(features[index]['layers'][1]['values'])
# layer_minus_3.append(features[index]['layers'][2]['values'])
# layer_minus_4.append(features[index]['layers'][3]['values'])
# new_token += features[index]['token']
# new_layers.append({'index': -1, 'values': list(np.mean(layer_minus_1, axis=0, dtype=np.float32))})
# new_layers.append({'index': -2, 'values': list(np.mean(layer_minus_2, axis=0, dtype=np.float32))})
# new_layers.append({'index': -3, 'values': list(np.mean(layer_minus_3, axis=0, dtype=np.float32))})
# new_layers.append({'index': -4, 'values': list(np.mean(layer_minus_4, axis=0, dtype=np.float32))})
# new_feature['token'] = new_token
# new_feature['layers'] = new_layers
# return new_feature
average_over_token_embedding(indexes, features)
len(embeddings[0])
len(embeddings[0])
%load_ext autoreload
%autoreload
import sys
sys.path.append('..')
from relation_extraction.data import utils
import numpy as np
bert_embeddings = utils.get_bert_embeddings('/crimea/geeticka/data/relation_extraction/semeval2010/pre-processed/original/bert/train_original_border_50.json')
elmo_embeddings = utils.get_elmo_embeddings('/crimea/geeticka/data/relation_extraction/semeval2010/pre-processed/original/elmo/train_original_border_-1.hdf5')
len(elmo_embeddings[0])
for item in elmo_embeddings[0]:
print(len(item))
for i_item in item:
print(len(i_item))
for ii_item in i_item:
print(len(ii_item))
for iii_item in i_item:
print(len(iii_item))
break
break
break
break
```
| github_jupyter |
# How to compute $\alpha$ and $\beta$ for a given asset
By Evgenia "Jenny" Nitishinskaya and Delaney Granizo-Mackenzie
Notebook released under the Creative Commons Attribution 4.0 License.
---
A fundamental concept in portfolio management and risk allocation is computing the $\beta$ of an asset. $\beta$ is a measurement of the covariance of the asset with the market in general, and is expressed in this model.
$$r_a \approx \alpha + \beta r_b$$
$r_a$ are the returns of the asset, and $r_b$ are the returns of the benchmark, usually a proxy for the market like the S&P 500.
$\beta$ can be defined as
$$\beta = \frac{Cov(r_a, r_b)}{Var(r_b)}$$
To actually compute $\beta$, we can use linear regression. We find the OLS best fit line for all points $(r_{a, t}, r_{b, t})$, where $r_{a, t}$ is the asset's returns for time $t$, and $r_{b, t}$ the same for the benchmark. The slope of this line is $\beta$, and the y-intercept is $\alpha$.
We'll start by getting data for a specific time range.
```
# Import libraries
import numpy as np
from statsmodels import regression
import statsmodels.api as sm
import matplotlib.pyplot as plt
import math
# Get data for the specified period and stocks
start = '2014-01-01'
end = '2015-01-01'
asset = get_pricing('TSLA', fields='price', start_date=start, end_date=end)
benchmark = get_pricing('SPY', fields='price', start_date=start, end_date=end)
# We have to take the percent changes to get to returns
# Get rid of the first (0th) element because it is NAN
r_a = asset.pct_change()[1:]
r_b = benchmark.pct_change()[1:]
# Let's plot them just for fun
r_a.plot()
r_b.plot();
```
Now we have to fit a line to the data to determine the slope. We use Ordinary Least Squares (OLS) for this.
```
# Let's define everything in familiar regression terms
X = r_b.values # Get just the values, ignore the timestamps
Y = r_a.values
# We add a constant so that we can also fit an intercept (alpha) to the model
# This just adds a column of 1s to our data
X = sm.add_constant(X)
model = regression.linear_model.OLS(Y, X)
model = model.fit()
# Remove the constant now that we're done
X = X[:, 1]
alpha = model.params[0]
beta = model.params[1]
print 'alpha: ' + str(alpha)
print 'beta: ' + str(beta)
```
We can plot the line of best fit to visualize this.
```
X2 = np.linspace(X.min(), X.max(), 100)
Y_hat = X2 * beta + alpha
plt.scatter(X, Y, alpha=0.3) # Plot the raw data
plt.xlabel("r_b")
plt.ylabel("r_a")
# Add the regression line, colored in red
plt.plot(X2, Y_hat, 'r', alpha=0.9);
```
# How to compute the volatility for a given asset
The volatility $\sigma$ of an asset is the standard deviation of its returns. A low volatility means that the returns are generally close to the mean, while a high volatility corresponds to returns that are often much higher and often much lower than expected.
We'll go ahead and continue using the stock from before:
```
# Use numpy to find the standard deviation of the returns
SD = np.std(Y)
print SD
# Let's compute the volatility for our benchmark, as well
benchSD = np.std(X)
print benchSD
```
This gives the daily volatility. As expected, the benchmark has a much lower volatility than the stock - a volatile asset would not make a good benchmark.
We generally compute the annualized volatility so that we can compare volatilities for daily, weekly, or monthly samples. To get it we normalize the standard deviation of the daily returns by multiplying by the square root of the number of trading days in a year:
$$\sigma_{\text{annual}} = SD \cdot \sqrt{252}$$
```
vol = SD*(252**.5)
print vol
benchvol = benchSD*(252**.5)
print benchvol
```
This tells us that we should expect the returns of a benchmark to cluster more closely around their mean than those of the stock. We can plot histograms of the returns to see this:
```
# Since we have many distinct values, we'll lump them into buckets of size .02
x_min = int(math.floor(X.min()))
x_max = int(math.ceil(X.max()))
plt.hist(X, bins=[0.02*i for i in range(x_min*50, x_max*50)], alpha=0.5, label='S&P')
plt.hist(Y, bins=[0.02*i for i in range(x_min*50, x_max*50)], alpha=0.5, label='Stock')
plt.legend(loc='upper right');
```
# How to compute the Sharpe and information ratios for an asset
The Sharpe and information ratios are used to calculate how well the historic returns of an asset compensate for its risk, relative to some benchmark or risk-free asset. An asset with a higher ratio has either higher returns, lower risk, or both. As when computing volatility, the standard deviation of the returns is used to measure risk.
$$R = \frac{E[r_a - r_b]}{\sqrt{Var(r_a - r_b)}}$$
$r_a$ are the returns of the asset, and $r_b$ are the returns of the benchmark; generally, Treasury bills are used when computing the Sharpe ratio, while the S&P 500 index is commonly used for the information ratio. We subtract the returns of the benchmark from the returns of the asset becasue we would like to get higher returns through our investment than we would, say, simply buying Treasury bills.
```
# Get the returns for a treasury-tracking ETF to be used in the Sharpe ratio
# Note that BIL is only being used in place of risk free rate,
# and should not be used in such a fashion for strategy development
riskfree = get_pricing('BIL', fields='price', start_date=start, end_date=end)
r_b_S = riskfree.pct_change()[1:]
X_S = r_b_S.values
# Compute the Sharpe ratio for the asset we've been working with
SR = np.mean(Y - X_S)/np.std(Y - X_S)
# Compute the information ratio for the asset we've been working with, using the S&P index
IR = np.mean(Y - X)/np.std(Y - X)
# Print results
print 'Sharpe ratio: ' + str(SR)
print 'Information ratio: ' + str(IR)
```
# How to compute the Sortino ratio for an asset
The Sharpe and information ratios are useful, but they penalize stocks for going above the expected return as well as for going below it. The Sortino ratio is modified to take into account only returns that fall below the mean.
$$S = \frac{E[r_a - r_b]}{\sqrt{Semivar(r_a - r_b)}}$$
The semivariance is the variance below the mean, and so quantifies the downside risk of our asset. Here as in the Sharpe ratio returns on Treasury bills can be used for $r_b$. The more skewed the distribution of returns is, the more the Sortino ratio will differ from the Sharpe ratio.
```
# To compute the semideviation, we want to filter out values which fall above the mean
meandif = np.mean(Y - X_S)
lows = [e for e in Y - X_S if e <= meandif]
# Because there is no built-in semideviation, we'll compute it ourselves
def dist(x):
return (x - meandif)**2
semidev = math.sqrt(sum(map(dist,lows))/len(lows))
Sortino = meandif/semidev
print 'Sortino ratio: ' + str(Sortino)
```
| github_jupyter |
## Preprocessing
- Clean up divvy bike trip data from 2013 to 2018
```
import numpy as np
import pandas as pd
import re
from multiprocessing import Pool, cpu_count
import gc
import os
import time
gc.enable()
# Deal with year data with different filenames
TRIP_FILE = 'data/Divvy_Trips'
YEAR = 2018
raw_file_name = {
'2013': ['all'],
'2014': ['Q1Q2', 'Q3-07', 'Q3-0809', 'Q4'],
'2015': ['Q1', 'Q2', '07', '08', '09', 'Q4'],
'2016': ['Q1', '04', '05', '06', 'Q3', 'Q4'],
'2017': ['Q1', 'Q2', 'Q3', 'Q4'],
'2018': ['Q1', 'Q2', 'Q3', 'Q4']
}
%%time
# Load data
df_lst = []
for q in raw_file_name[str(YEAR)]:
print(f'Loading from {TRIP_FILE}_{YEAR}_{q}.csv...')
df_tmp = pd.read_csv(TRIP_FILE+'_'+str(YEAR)+'_'+q+'.csv')
if YEAR == 2018 and q == 'Q1':
df_tmp.rename(
columns={
"01 - Rental Details Rental ID": "trip_id",
"01 - Rental Details Local Start Time": "start_time",
"01 - Rental Details Local End Time": "end_time",
"01 - Rental Details Bike ID": "bikeid",
"01 - Rental Details Duration In Seconds Uncapped": "tripduration",
"03 - Rental Start Station ID": "from_station_id",
"03 - Rental Start Station Name": "from_station_name",
"02 - Rental End Station ID": "to_station_id",
"02 - Rental End Station Name": "to_station_name",
"User Type": "usertype",
"Member Gender": "gender",
"05 - Member Details Member Birthday Year": "birthyear"
},
inplace=True
)
print(f'{TRIP_FILE}_{YEAR}_{q}.csv loaded!')
df_lst.append(df_tmp)
trip = pd.concat(df_lst, ignore_index=True)
# Clean up
del df_lst
del df_tmp
gc.collect()
# Rename column names from `startime` to `start_time` for consistency
if 'starttime' in trip.columns:
trip.rename(columns={"starttime": "start_time"}, inplace=True)
trip.info()
def _get_time(string, tp):
index_dict = {
'month': 0,
'day': 1,
'hour': 2
}
if '-' in string:
return int(re.match(r'[0-9]+-([0-9]+)-([0-9]+) ([0-9]+):', string).groups()[index_dict[tp]])
else:
return int(re.match(r'([0-9]+)/([0-9]+)/[0-9]+ ([0-9]+):', string).groups()[index_dict[tp]])
def parse_start_time(args):
"""
Function to parse `start_time` data into:
- day
- month
- hour
"""
args['year'] = YEAR
for t in ['day', 'month', 'hour']:
args[t] = args.apply(lambda x: _get_time(x.start_time, t), axis=1)
return args.drop('start_time', axis=1)
%%time
# Process start_time data with mutiple processors
n_thread = 4
time_df_raw = trip[['trip_id', 'start_time']]
args = np.array_split(time_df_raw, n_thread)
with Pool(processes=n_thread) as p:
result = p.map(parse_start_time, args)
time_df = pd.concat(list(result), ignore_index=True)
# Merge postprocessed data into main dataframe
trip = pd.merge(left=trip, right=time_df, on='trip_id', how='left')
print(f'start_time data processed!')
# Convert birthyear into age of each customer
if 'birthyear' in trip.columns:
trip['Age'] = trip['year'] - trip['birthyear']
else:
trip['Age'] = np.nan
# # Merge station data into trip data (start)
# trip = (pd.merge(left=trip, right=station, left_on='from_station_id', right_on='id', how='left')
# .drop(['id', 'name', 'online_date'], axis=1)
# .rename(columns={'city': 'from_city',
# 'latitude': 'from_latitude',
# 'longitude': 'from_longitude',
# 'dpcapacity': 'from_dpcapacity'})
# )
# # Merge station data into trip data (end)
# trip = (pd.merge(left=trip, right=station, left_on='to_station_id', right_on='id', how='left')
# .drop(['id', 'name', 'online_date'], axis=1)
# .rename(columns={'city': 'to_city',
# 'latitude': 'to_latitude',
# 'longitude': 'to_longitude',
# 'dpcapacity': 'to_dpcapacity'})
# )
# Drop useless columns
for uc in ['start_time', 'end_time', 'stoptime', 'birthyear']:
if uc in trip.columns:
trip = trip.drop(uc, axis=1)
for c in trip.columns:
if 'Unnamed' in c:
trip = trip.drop([c], axis=1)
print(f'data process done!')
trip.info()
# Saving to feather file
trip.to_feather(f'data/Divvy_data_{YEAR}.feather')
print(f'Data saved to feather file!')
```
| github_jupyter |
# Errors and Exceptions
A Python program terminates as soon as it encounters an error. In Python, an error can be a syntax error or an exception. In this article we will have a look at:
- Syntax Error vs. Exception
- How to raise Exceptions
- How to handle Exceptions
- Most common built-in Exceptions
- How to define your own Exception
## Syntax Errors
A **Syntax Error** occurs when the parser detects a syntactically incorrect statement. A syntax error can be for example a typo, missing brackets, no new line (see code below), or wrong identation (this will actually raise its own IndentationError, but its subclassed from a SyntaxError).
```
a = 5 print(a)
```
## Exceptions
Even if a statement is syntactically correct, it may cause an error when it is executed. This is called an **Exception Error**. There are several different error classes, for example trying to add a number and a string will raise a TypeError.
```
a = 5 + '10'
```
## Raising an Exception
If you want to force an exception to occur when a certain condition is met, you can use the `raise` keyword.
```
x = -5
if x < 0:
raise Exception('x should not be negative.')
```
You can also use the `assert` statement, which will throw an AssertionError if your assertion is **not** True.
This way, you can actively test some conditions that have to be fulfilled instead of waiting for your program to unexpectedly crash midway. Assertion is also used in **unit testing**.
```
x = -5
assert (x >= 0), 'x is not positive.'
# --> Your code will be fine if x >= 0
```
## Handling Exceptions
You can use a `try` and `except` block to catch and handle exceptions. If you can catch an exceptions your program won't terminate, and can continue.
```
# This will catch all possible exceptions
try:
a = 5 / 0
except:
print('some error occured.')
# You can also catch the type of exception
try:
a = 5 / 0
except Exception as e:
print(e)
# It is good practice to specify the type of Exception you want to catch.
# Therefore, you have to know the possible errors
try:
a = 5 / 0
except ZeroDivisionError:
print('Only a ZeroDivisionError is handled here')
# You can run multiple statements in a try block, and catch different possible exceptions
try:
a = 5 / 1 # Note: No ZeroDivisionError here
b = a + '10'
except ZeroDivisionError as e:
print('A ZeroDivisionError occured:', e)
except TypeError as e:
print('A TypeError occured:', e)
```
#### `else` clause
You can use an else statement that is run if no exception occured.
```
try:
a = 5 / 1
except ZeroDivisionError as e:
print('A ZeroDivisionError occured:', e)
else:
print('Everything is ok')
```
#### `finally` clause
You can use a finally statement that always runs, no matter if there was an exception or not. This is for example used to make some cleanup operations.
```
try:
a = 5 / 1 # Note: No ZeroDivisionError here
b = a + '10'
except ZeroDivisionError as e:
print('A ZeroDivisionError occured:', e)
except TypeError as e:
print('A TypeError occured:', e)
else:
print('Everything is ok')
finally:
print('Cleaning up some stuff...')
```
## Common built-in Exceptions
You can find all built-in Exceptions here: https://docs.python.org/3/library/exceptions.html
- ImportError: If a module cannot be imported
- NameError: If you try to use a variable that was not defined
- FileNotFoundError: If you try to open a file that does not exist or you specify the wrong path
- ValueError: When an operation or function receives an argument that has the right type but an inappropriate value,
e.g. try to remove a value from a list that does not exist
- TypeError: Raised when an operation or function is applied to an object of inappropriate type.
- IndexError: If you try to access an invalid index of a sequence, e.g a list or a tuple.
- KeyError: If you try to access a non existing key of a dictionary.
```
# ImportError
import nonexistingmodule
# NameError
a = someundefinedvariable
# FileNotFoundError
with open('nonexistingfile.txt') as f:
read_data = f.read()
# ValueError
a = [0, 1, 2]
a.remove(3)
# TypeError
a = 5 + "10"
# IndexError
a = [0, 1, 2]
value = a[5]
# KeyError
my_dict = {"name": "Max", "city": "Boston"}
age = my_dict["age"]
```
## Define your own Exceptions
You can define your own exception class that should be derived from the built-in `Exception` class. Most exceptions are defined with names that end in 'Error', similar to the naming of the standard exceptions. Exception classes can be defined like any other class, but are usually kept simple, often only offering a number of attributes that allow information about the error to be extracted by handlers.
```
# minimal example for own exception class
class ValueTooHighError(Exception):
pass
# or add some more information for handlers
class ValueTooLowError(Exception):
def __init__(self, message, value):
self.message = message
self.value = value
def test_value(a):
if a > 1000:
raise ValueTooHighError('Value is too high.')
if a < 5:
raise ValueTooLowError('Value is too low.', a) # Note that the constructor takes 2 arguments here
return a
try:
test_value(1)
except ValueTooHighError as e:
print(e)
except ValueTooLowError as e:
print(e.message, 'The value is:', e.value)
```
| github_jupyter |
# Counting Boats from Space (Part 2) - Train a Neural Network with Weak Supervision
```
%reload_ext autoreload
%autoreload 2
%load_ext dotenv
%dotenv
%matplotlib inline
```
## 1. Install, Import requirements
```
import os
import sys
import pandas as pd
import torch
from torch.utils.data import DataLoader
sys.path.insert(0,os.path.dirname('../src/'))
from dataset import getImageSetDirectories, S2_Dataset, plot_dataset
from model import Model
from train import train, get_failures_or_success
torch.cuda.is_available() # gpu support
```
## 2. Init Dataset
```
data_dir = "/home/jovyan/data" # data directory (path)
checkpoint_dir = "../factory"
bands = ['img_08', 'bg_ndwi']
test_size = 0.1
train_list, val_list, fig = getImageSetDirectories(data_dir=os.path.join(data_dir, 'chips'),
labels_filename="../data/labels.csv",
band_list=bands, test_size=test_size, plot_coords=False, plot_class_imbalance=True, seed=38)
fig # mapbox plot train/val coordinates
train_dataset = S2_Dataset(imset_dir=train_list, augment=True, labels_filename='../data/labels.csv')
val_dataset = S2_Dataset(imset_dir=val_list, augment=False, labels_filename='../data/labels.csv')
plot_dataset(val_dataset, n_frames=20, n_rows=2)
```
## 3. Train PyTorch Classifier
```
# training co<nfig
input_dim = train_dataset[0]['img'].shape[0]
hidden_dim, kernel_size, pool_size, n_max = 16, 3, 10, 1
train_dataloader = DataLoader(train_dataset, batch_size=8, shuffle=True, num_workers=16)
val_dataloader = DataLoader(val_dataset, batch_size=1, shuffle=True, num_workers=4)
best_metrics = train(train_dataloader=train_dataloader, val_dataloader=val_dataloader,
input_dim=input_dim, hidden_dim=hidden_dim, kernel_size=kernel_size, pool_size=pool_size, n_max=n_max, drop_proba=0.1,
ld=0.5, n_epochs=200, lr=0.004, lr_step=2, lr_decay=0.97,
device='cuda:0', checkpoints_dir=checkpoint_dir, seed=123, verbose=1, version='0.1.2')
for k,v in best_metrics.items():
print('{} {:.4f}'.format(k,v))
```
## 4. Test Model
```
# load pretrained model
model = Model(input_dim=input_dim, hidden_dim=hidden_dim, kernel_size=kernel_size, pool_size=pool_size, n_max=n_max, device='cuda:0', version='0.1.1')
checkpoint_file = os.path.join(checkpoint_dir, model.folder, 'model.pth')
model.load_checkpoint(checkpoint_file=checkpoint_file)
model = model.eval()
# Display failures (train, val), scatter plot (Predicted vs True) and inspect hidden channels --> Re label?
image_titles, relabel_images = get_failures_or_success(model, train_dataset, success=None, filter_on=None,
water_NDWI=0.4, filter_peaks=True, downsample=True, # filter_peaks and/or downsample
plot_heatmap=False)
# Display failures (train, val), scatter plot (Predicted vs True) and inspect hidden channels --> Re label?
image_titles, relabel_images = get_failures_or_success(model, val_dataset, success=None, filter_on=None,
water_NDWI=0.4, filter_peaks=True, downsample=True, # filter_peaks and/or downsample
plot_heatmap=False)
```
## 5. Relabel inputs
```
import superintendent
from annotation_utils import display_image_and_references, display_heatmap_prediction
```
### Load superintendent widget and labelling
```
csv_file_path = "../data/labels.csv"
labels_df = pd.read_csv(csv_file_path, index_col = ['lat_lon', 'timestamp'], dtype={'count': float}) # read the labels.csv file
labeller = superintendent.ClassLabeller(
features=image_titles,
options=[i for i in range(-1, 6)],
display_func=display_heatmap_prediction # or display_image_and_references
)
#labeller
```
### Extract new labels and save them in labels_df
```
for i in range(len(relabel_images)): # relabel_images: list of inputs that should maybe be relabelled
timestamp = relabel_images[i].stem.split('t_')[1]
lat_lon = relabel_images[i].parts[-2]
count = labeller.new_labels[i]
# overwrite if the
if count:
labels_df.at[(lat_lon, timestamp)] = count
```
### Dump back to csv file
```
#labels_df.to_csv(csv_file_path) # # write to labels.csv file
```
| github_jupyter |
# How many cases of COVID-19 does each U.S. state really have?
> Reported U.S. case counts are based on the number of administered tests that are positive. Since not everyone is tested, this number is biased. We use Bayesian techniques to estimate the true number of cases in each U.S. state.
- author: Joseph Richards
- image: images/covid-state-case-estimation.png
- hide: false
- comments: true
- categories: [MCMC, US, states, cases]
- permalink: /covid-19-us-case-estimation/
- toc: false
```
#hide
# Setup and imports
%matplotlib inline
import warnings
warnings.simplefilter('ignore')
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import pymc3 as pm
import requests
from IPython.display import display, Markdown
#hide
# Data utilities:
def get_statewise_testing_data():
'''
Pull all statewise data required for model fitting and
prediction
Returns:
* df_out: DataFrame for model fitting where inclusion
requires testing data from 7 days ago
* df_pred: DataFrame for count prediction where inclusion
only requires testing data from today
'''
# Pull testing counts by state:
out = requests.get('https://covidtracking.com/api/states')
df_out = pd.DataFrame(out.json())
df_out.set_index('state', drop=True, inplace=True)
# Pull time-series of testing counts:
ts = requests.get('https://covidtracking.com/api/states/daily')
df_ts = pd.DataFrame(ts.json())
# Get data from last week
date_last_week = df_ts['date'].unique()[7]
df_ts_last_week = _get_test_counts(df_ts, df_out.index, date_last_week)
df_out['num_tests_7_days_ago'] = \
(df_ts_last_week['positive'] + df_ts_last_week['negative'])
df_out['num_pos_7_days_ago'] = df_ts_last_week['positive']
# Get data from today:
date_today = df_ts['date'].unique()[1]
df_ts_today = _get_test_counts(df_ts, df_out.index, date_today)
df_out['num_tests_today'] = \
(df_ts_today['positive'] + df_ts_today['negative'])
# State population:
df_pop = pd.read_excel(('https://github.com/jwrichar/COVID19-mortality/blob/'
'master/data/us_population_by_state_2019.xlsx?raw=true'),
skiprows=2, skipfooter=5)
r = requests.get(('https://raw.githubusercontent.com/jwrichar/COVID19-mortality/'
'master/data/us-state-name-abbr.json'))
state_name_abbr_lookup = r.json()
df_pop.index = df_pop['Geographic Area'].apply(
lambda x: str(x).replace('.', '')).map(state_name_abbr_lookup)
df_pop = df_pop.loc[df_pop.index.dropna()]
df_out['total_population'] = df_pop['Total Resident\nPopulation']
# Tests per million people, based on today's test coverage
df_out['tests_per_million'] = 1e6 * \
(df_out['num_tests_today']) / df_out['total_population']
df_out['tests_per_million_7_days_ago'] = 1e6 * \
(df_out['num_tests_7_days_ago']) / df_out['total_population']
# People per test:
df_out['people_per_test'] = 1e6 / df_out['tests_per_million']
df_out['people_per_test_7_days_ago'] = \
1e6 / df_out['tests_per_million_7_days_ago']
# Drop states with messed up / missing data:
# Drop states with missing total pop:
to_drop_idx = df_out.index[df_out['total_population'].isnull()]
print('Dropping %i/%i states due to lack of population data: %s' %
(len(to_drop_idx), len(df_out), ', '.join(to_drop_idx)))
df_out.drop(to_drop_idx, axis=0, inplace=True)
df_pred = df_out.copy(deep=True) # Prediction DataFrame
# Criteria for model fitting:
# Drop states with missing test count 7 days ago:
to_drop_idx = df_out.index[df_out['num_tests_7_days_ago'].isnull()]
print('Dropping %i/%i states due to lack of tests: %s' %
(len(to_drop_idx), len(df_out), ', '.join(to_drop_idx)))
df_out.drop(to_drop_idx, axis=0, inplace=True)
# Drop states with no cases 7 days ago:
to_drop_idx = df_out.index[df_out['num_pos_7_days_ago'] == 0]
print('Dropping %i/%i states due to lack of positive tests: %s' %
(len(to_drop_idx), len(df_out), ', '.join(to_drop_idx)))
df_out.drop(to_drop_idx, axis=0, inplace=True)
# Criteria for model prediction:
# Drop states with missing test count today:
to_drop_idx = df_pred.index[df_pred['num_tests_today'].isnull()]
print('Dropping %i/%i states in prediction data due to lack of tests: %s' %
(len(to_drop_idx), len(df_pred), ', '.join(to_drop_idx)))
df_pred.drop(to_drop_idx, axis=0, inplace=True)
# Cast counts to int
df_pred['negative'] = df_pred['negative'].astype(int)
df_pred['positive'] = df_pred['positive'].astype(int)
return df_out, df_pred
def _get_test_counts(df_ts, state_list, date):
ts_list = []
for state in state_list:
state_ts = df_ts.loc[df_ts['state'] == state]
# Back-fill any gaps to avoid crap data gaps
state_ts.fillna(method='bfill', inplace=True)
record = state_ts.loc[df_ts['date'] == date]
ts_list.append(record)
df_ts = pd.concat(ts_list, ignore_index=True)
return df_ts.set_index('state', drop=True)
#hide
# Model utilities
def case_count_model_us_states(df):
# Normalize inputs in a way that is sensible:
# People per test: normalize to South Korea
# assuming S.K. testing is "saturated"
ppt_sk = np.log10(51500000. / 250000)
df['people_per_test_normalized'] = (
np.log10(df['people_per_test_7_days_ago']) - ppt_sk)
n = len(df)
# For each country, let:
# c_obs = number of observed cases
c_obs = df['num_pos_7_days_ago'].values
# c_star = number of true cases
# d_obs = number of observed deaths
d_obs = df[['death', 'num_pos_7_days_ago']].min(axis=1).values
# people per test
people_per_test = df['people_per_test_normalized'].values
covid_case_count_model = pm.Model()
with covid_case_count_model:
# Priors:
mu_0 = pm.Beta('mu_0', alpha=1, beta=100, testval=0.01)
# sig_0 = pm.Uniform('sig_0', lower=0.0, upper=mu_0 * (1 - mu_0))
alpha = pm.Bound(pm.Normal, lower=0.0)(
'alpha', mu=8, sigma=3, shape=1)
beta = pm.Bound(pm.Normal, upper=0.0)(
'beta', mu=-1, sigma=1, shape=1)
# beta = pm.Normal('beta', mu=0, sigma=1, shape=3)
sigma = pm.HalfNormal('sigma', sigma=0.5, testval=0.1)
# sigma_1 = pm.HalfNormal('sigma_1', sigma=2, testval=0.1)
# Model probability of case under-reporting as logistic regression:
mu_model_logit = alpha + beta * people_per_test
tau_logit = pm.Normal('tau_logit',
mu=mu_model_logit,
sigma=sigma,
shape=n)
tau = np.exp(tau_logit) / (np.exp(tau_logit) + 1)
c_star = c_obs / tau
# Binomial likelihood:
d = pm.Binomial('d',
n=c_star,
p=mu_0,
observed=d_obs)
return covid_case_count_model
#hide
df, df_pred = get_statewise_testing_data()
# Initialize the model:
mod = case_count_model_us_states(df)
# Run MCMC sampler
with mod:
trace = pm.sample(500, tune=500, chains=1)
#hide_input
n = len(trace['beta'])
# South Korea:
ppt_sk = np.log10(51500000. / 250000)
# Compute predicted case counts per state right now
logit_now = pd.DataFrame([
pd.Series(np.random.normal((trace['alpha'][i] + trace['beta'][i] * (np.log10(df_pred['people_per_test']) - ppt_sk)),
trace['sigma'][i]), index=df_pred.index)
for i in range(len(trace['beta']))])
prob_missing_now = np.exp(logit_now) / (np.exp(logit_now) + 1)
predicted_counts_now = np.round(df_pred['positive'] / prob_missing_now.mean(axis=0)).astype(int)
predicted_counts_now_lower = np.round(df_pred['positive'] / prob_missing_now.quantile(0.975, axis=0)).astype(int)
predicted_counts_now_upper = np.round(df_pred['positive'] / prob_missing_now.quantile(0.025, axis=0)).astype(int)
case_increase_percent = list(map(lambda x, y: (((x - y) / float(y))),
predicted_counts_now, df_pred['positive']))
df_summary = pd.DataFrame(
data = {
'Cases Reported': df_pred['positive'],
'Cases Estimated': predicted_counts_now,
'Percent Increase': case_increase_percent,
'Tests per Million People': df_pred['tests_per_million'].round(1),
'Cases Estimated (range)': list(map(lambda x, y: '(%i, %i)' % (round(x), round(y)),
predicted_counts_now_lower, predicted_counts_now_upper))
},
index=df_pred.index)
from datetime import datetime
display(Markdown("## Summary for the United States on %s:" % str(datetime.today())[:10]))
display(Markdown(f"**Reported Case Count:** {df_summary['Cases Reported'].sum():,}"))
display(Markdown(f"**Predicted Case Count:** {df_summary['Cases Estimated'].sum():,}"))
case_increase_percent = 100. * (df_summary['Cases Estimated'].sum() - df_summary['Cases Reported'].sum()) / df_summary['Cases Estimated'].sum()
display(Markdown("**Percentage Underreporting in Case Count:** %.1f%%" % case_increase_percent))
#hide
df_summary.loc[:, 'Ratio'] = df_summary['Cases Estimated'] / df_summary['Cases Reported']
df_summary.columns = ['Reported Cases', 'Estimated Cases', '% Increase',
'Tests per Million', 'Estimated Range', 'Ratio']
df_display = df_summary[['Reported Cases', 'Estimated Cases', 'Estimated Range', 'Ratio', 'Tests per Million']].copy()
```
## COVID-19 Case Estimates, by State
### Definition Of Fields:
- **Reported Cases**: The number of cases reported by each state, which is a function of how many tests are positive.
- **Estimated Cases**: The predicted number of cases, accounting for the fact that not everyone is tested.
- **Estimated Range**: The 95% confidence interval of the predicted number of cases.
- **Ratio**: `Estimated Cases` divided by `Reported Cases`.
- **Tests per Million**: The number of tests administered per one million people. The less tests administered per capita, the larger the difference between reported and estimated number of cases, generally.
```
#hide_input
df_display.sort_values(by='Estimated Cases', ascending=False).style.background_gradient(
cmap='Oranges').format({'Ratio': "{:.1f}"}).format({'Tests per Million': "{:.1f}"})
#hide
# Shown are the current reported number of cases, by state (black dot) with the model-estimated case counts (blue circle, plus 95% posterior predictive interval).
# xerr = [df_summary['Cases Estimated'] - predicted_counts_now_lower, predicted_counts_now_upper - df_summary['Cases Estimated']]
# fig, axs = plt.subplots(1, 1, figsize=(15, 15))
# ax = plt.errorbar(df_summary['Cases Estimated'], range(len(df_summary)-1, -1, -1), xerr=xerr,
# fmt='o', elinewidth=1, label='Estimate')
# ax = plt.yticks(range(len(df_summary)), df.index[::-1])
# ax = plt.errorbar(df_summary['Cases Reported'], range(len(df_summary)-1, -1, -1), xerr=None,
# fmt='.', color='k', label='Observed')
# ax = plt.xlabel('COVID-19 Case Counts', size=20)
# ax = plt.legend(fontsize='xx-large')
# ax = plt.grid(linestyle='--', color='grey', axis='x')
```
## Appendix: Model Diagnostics
### Derived relationship between Test Capacity and Case Under-reporting
Plotted is the estimated relationship between test capacity (in terms of people per test -- larger = less testing) and the likelihood a COVID-19 case is reported (lower = more under-reporting of cases).
The lines represent the posterior samples from our MCMC run (note the x-axis is plotted on a log scale). The rug plot shows the current test capacity for each state (black '|') and the capacity one week ago (cyan '+'). For comparison, South Korea's testing capacity is currently at the very left of the graph (200 people per test).
```
#hide_input
# Plot pop/test vs. Prob of case detection for all posterior samples:
x = np.linspace(0.0, 4.0, 101)
logit_pcase = pd.DataFrame([
trace['alpha'][i] + trace['beta'][i] * x
for i in range(n)])
pcase = np.exp(logit_pcase) / (np.exp(logit_pcase) + 1)
fig, ax = plt.subplots(1, 1, figsize=(14, 9))
for i in range(n):
ax = plt.plot(10**(ppt_sk + x), pcase.iloc[i], color='grey', lw=.1, alpha=.5)
plt.xscale('log')
plt.xlabel('State-wise population per test', size=14)
plt.ylabel('Probability a true case is detected', size=14)
# rug plots:
ax=plt.plot(df_pred['people_per_test'], np.zeros(len(df_pred)),
marker='|', color='k', ls='', ms=20,
label='U.S. State-wise Test Capacity Now')
ax=plt.plot(df['people_per_test_7_days_ago'], np.zeros(len(df)),
marker='+', color='c', ls='', ms=10,
label='U.S. State-wise Test Capacity 7 Days Ago')
ax = plt.legend(fontsize='x-large')
```
## About this Analysis
This analysis was done by [Joseph Richards](https://twitter.com/joeyrichar).
This project[^1] uses the testing rates per state from [https://covidtracking.com/](https://covidtracking.com/), which reports case counts and mortality by state. This is used to **estimate the number of unreported (untested) COVID-19 cases in each U.S. state.**
The analysis makes a few assumptions:
1. The probability that a case is reported by a state is a function of the number of tests run per person in that state. Hence the degree of under-reported cases is a function of tests run per capita.
2. The underlying mortality rate is the same across every state.
3. Patients take time to succumb to COVID-19, so the mortality counts *today* reflect the case counts *7 days ago*. E.g., mortality rate = (cumulative deaths today) / (cumulative cases 7 days ago).
The model attempts to find the most likely relationship between state-wise test volume (per capita) and under-reporting, such that the true underlying mortality rates between the individual states are as similar as possible. The model simultaneously finds the most likely posterior distribution of mortality rates, the most likely *true* case count per state, and the test volume vs. case underreporting relationship.
[^1]: Full details about the model are available at: https://github.com/jwrichar/COVID19-mortality
| github_jupyter |
<a href="https://colab.research.google.com/github/maxwelljohn/siamese-word2vec/blob/master/siamese.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
import numpy as np
import datetime
import heapq
import itertools
import matplotlib.pyplot as plt
%matplotlib inline
import nltk
import os
import random
import skimage.transform
import sys
import time
import unittest
nltk.download('wordnet')
nltk.download('punkt')
from nltk.corpus import wordnet as wn
from keras.utils.data_utils import get_file
from keras.models import Model
from keras.layers import Input, Flatten, Dense, Dropout, Lambda
from keras.optimizers import Adam
from keras import backend as K
from keras import regularizers
from keras.constraints import non_neg
from scipy.misc import imread
from sklearn.manifold import TSNE
!pip install scikit-optimize==0.5.2
import skopt
!pip install -U -q PyDrive==1.3.1
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from google.colab import auth
from oauth2client.client import GoogleCredentials
def similarity(a, b):
# Cosine similarity
return np.dot(a, b) / max(np.linalg.norm(a) * np.linalg.norm(b), sys.float_info.epsilon)
def keras_norm(vect):
return K.sqrt(K.batch_dot(vect, vect, axes=1))
def keras_similarity(vects):
x, y = vects
# Cosine similarity
return K.batch_dot(x, y, axes=1) / K.maximum(keras_norm(x) * keras_norm(y), K.epsilon())
def sim_output_shape(shapes):
shape1, shape2 = shapes
return (shape1[0], 1)
def create_base_network(input_shape, output_size=128, reg_rate=0):
'''Base network to be shared (eq. to feature extraction).
'''
input = Input(shape=input_shape)
x = input
x = Dense(output_size, kernel_regularizer=regularizers.l2(reg_rate))(x)
return Model(input, x)
def syn_accuracy(y_true, y_pred):
'''Compute synonym classification accuracy with a variable threshold on similarities.
'''
median = np.median(y_pred)
pred = y_pred.ravel() > median
return np.mean(pred == y_true)
def syn_accuracy_scores(y_true, y_pred):
y_pred = y_pred.ravel()
median = np.median(y_pred)
sd = np.std(y_pred)
standardized = (y_pred - median) / sd
multiplier = y_true * 2 - 1
return standardized * multiplier
def ant_accuracy(y_true, y_pred):
'''Compute antonym classification accuracy with a variable threshold on similarities.
'''
median = np.median(y_pred)
pred = -((y_pred.ravel() < median).astype(np.int, casting='safe', copy=False))
return np.mean(pred == y_true)
def keras_syn_accuracy(y_true, y_pred):
'''Compute synonym classification accuracy with a fixed threshold on similarities.
'''
return K.mean(K.equal(y_true, K.cast(y_pred > 0.75, y_true.dtype)))
def keras_ant_accuracy(y_true, y_pred):
'''Compute antonym classification accuracy with a fixed threshold on similarities.
'''
return K.mean(K.equal(y_true, K.cast(y_pred < -0.75, y_true.dtype)))
!unzip -u glove.6B.zip || (wget http://nlp.stanford.edu/data/glove.6B.zip && unzip -u glove.6B.zip)
!egrep '^[a-z]+ ' glove.6B.300d.txt > glove.tokens.txt
!cut -d' ' -f1 glove.tokens.txt > glove.tokens.strings.txt
!cut -d' ' -f2- glove.tokens.txt > glove.tokens.vectors.txt
word_strings = np.loadtxt('glove.tokens.strings.txt', dtype=object)
word_vectors = np.loadtxt('glove.tokens.vectors.txt')
input_shape = word_vectors.shape[1:]
assert len(word_strings) == len(word_vectors)
index_for_word = {}
for i, word in enumerate(word_strings):
index_for_word[word] = i
!rm *.zip *.txt
wnl = nltk.WordNetLemmatizer()
def might_be_synonyms(w1, w2):
s1 = set()
d1 = set()
s2 = set()
d2 = set()
for synset in wn.synsets(w1):
lemma_names = synset.lemma_names()
s1.update(lemma_names)
d1.update(nltk.word_tokenize(synset.definition()))
for synset in wn.synsets(w2):
lemma_names = synset.lemma_names()
s2.update(lemma_names)
d2.update(nltk.word_tokenize(synset.definition()))
total_intersection = len(s1.intersection(s2)) + len(d1.intersection(s2)) + len(d2.intersection(s1))
return total_intersection > 0
class TestSynonyms(unittest.TestCase):
def test_true(self):
pairs = [
('car', 'auto'),
('auto', 'car'),
('car', 'railcar'),
('small', 'tiny'),
('small', 'miniature'),
]
for word1, word2 in pairs:
self.assertTrue(might_be_synonyms(word1, word2))
def test_false(self):
pairs = [
('car', 'airplane'),
('car', 'fast'),
('small', 'focused'),
('small', 'accidental'),
('small', 'rabbit'),
('small', 'flippant'),
]
for word1, word2 in pairs:
self.assertFalse(might_be_synonyms(word1, word2))
def get_antonyms(word):
result = set()
for lemma in wn.lemmas(word):
for antonym in lemma.antonyms():
result.add(antonym.name())
return result
def might_be_antonyms(w1, w2):
w1_antonyms = get_antonyms(w1)
w2_antonyms = get_antonyms(w2)
for w1_antonym in w1_antonyms:
if might_be_synonyms(w2, w1_antonym):
return True
for w2_antonym in w2_antonyms:
if might_be_synonyms(w1, w2_antonym):
return True
class TestAntonyms(unittest.TestCase):
def test_get_antonyms(self):
self.assertEqual(get_antonyms('big'), set(['little', 'small']))
self.assertEqual(get_antonyms('fast'), set(['slow']))
self.assertEqual(get_antonyms('big'), set(['little', 'small']))
def test_true(self):
pairs = [
('big', 'small'),
('big', 'minor'),
('big', 'tiny'),
('big', 'miniature'),
('fast', 'slow'),
('fast', 'sluggish'),
('loud', 'soft'),
('loud', 'quiet'),
]
for word1, word2 in pairs:
self.assertTrue(might_be_antonyms(word1, word2))
def test_false(self):
pairs = [
('big', 'huge'),
('big', 'clean'),
('fast', 'speedy'),
('fast', 'unusual'),
('loud', 'noisy'),
('loud', 'flat'),
]
for word1, word2 in pairs:
self.assertFalse(might_be_antonyms(word1, word2))
unittest.main(argv=['first-arg-is-ignored'], exit=False)
def create_pairs(word_strings, word_vectors, class_indices, pos_label, max_per_class=float('infinity')):
'''Positive and negative pair creation.
Alternates between positive and negative pairs.
'''
string_pairs = []
vector_pairs = []
labels = []
for family in class_indices:
sibling_pairs = np.array(list(itertools.combinations(family, 2)))
shuffled_indices = np.arange(len(word_strings))
np.random.shuffle(shuffled_indices)
next_index = 0
if len(sibling_pairs) > max_per_class:
sibling_pairs = sibling_pairs[np.random.choice(len(sibling_pairs), max_per_class)]
for sibling_pair in sibling_pairs:
np.random.shuffle(sibling_pair)
anchor, pos = sibling_pair
string_pairs.append([word_strings[anchor], word_strings[pos]])
vector_pairs.append([word_vectors[anchor], word_vectors[pos]])
labels.append(pos_label)
random_neg = None
while random_neg == None or random_neg in family or \
might_be_synonyms(word_strings[anchor], word_strings[random_neg]) or \
might_be_antonyms(word_strings[anchor], word_strings[random_neg]):
if next_index >= len(shuffled_indices):
np.random.shuffle(shuffled_indices)
next_index = 0
random_neg = shuffled_indices[next_index]
next_index += 1
string_pairs.append([word_strings[anchor], word_strings[random_neg]])
vector_pairs.append([word_vectors[anchor], word_vectors[random_neg]])
labels.append(0)
assert len(string_pairs) == len(vector_pairs) == len(labels)
return np.array(string_pairs), np.array(vector_pairs), np.array(labels)
def create_train_dev_test(word_strings, word_vectors, class_count, word_class, pos_label):
classes = list(range(class_count))
random.seed(850101)
random.shuffle(classes)
random.seed()
train_dev_split = round(len(classes)*0.6)
dev_test_split = round(len(classes)*0.8)
train_classes = classes[:train_dev_split]
dev_classes = classes[train_dev_split:dev_test_split]
test_classes = classes[dev_test_split:]
assert len(set(train_classes).intersection(set(dev_classes))) == 0
assert len(set(dev_classes).intersection(set(test_classes))) == 0
assert len(set(train_classes).intersection(set(test_classes))) == 0
class_indices = [np.where(word_class == c)[0] for c in train_classes]
tr_strings, tr_pairs, tr_y = create_pairs(word_strings, word_vectors, class_indices, pos_label)
class_indices = [np.where(word_class == c)[0] for c in dev_classes]
dev_strings, dev_pairs, dev_y = create_pairs(word_strings, word_vectors, class_indices, pos_label)
class_indices = [np.where(word_class == c)[0] for c in test_classes]
te_strings, te_pairs, te_y = create_pairs(word_strings, word_vectors, class_indices, pos_label)
return tr_strings, tr_pairs, tr_y, dev_strings, dev_pairs, dev_y, te_strings, te_pairs, te_y
syn_class = -np.ones(len(word_strings), dtype=np.int)
shuffled_indices = np.arange(len(word_strings))
np.random.shuffle(shuffled_indices)
class_count = 0
for i in shuffled_indices:
word = word_strings[i]
synsets = wn.synsets(word)
if synsets:
syn_indices = [index_for_word[syn] for syn in synsets[0].lemma_names() if syn in index_for_word and syn_class[index_for_word[syn]] == -1]
if len(syn_indices) > 1:
syn_class[syn_indices] = class_count
class_count += 1
tr_syn_strings, tr_syn_pairs, tr_syn_y, \
dev_syn_strings, dev_syn_pairs, dev_syn_y, \
te_syn_strings, te_syn_pairs, te_syn_y = create_train_dev_test(word_strings, word_vectors, class_count, syn_class, 1)
ant_class = -np.ones(len(word_strings), dtype=np.int)
shuffled_indices = np.arange(len(word_strings))
np.random.shuffle(shuffled_indices)
class_count = 0
for i in shuffled_indices:
word = word_strings[i]
antonyms = get_antonyms(word)
if antonyms:
known_antonyms = [a for a in antonyms if a in index_for_word]
if known_antonyms:
ant_indices = [i, index_for_word[known_antonyms[0]]]
if all(ant_class[ant_indices] == -1):
ant_class[ant_indices] = class_count
class_count += 1
tr_ant_strings, tr_ant_pairs, tr_ant_y, \
dev_ant_strings, dev_ant_pairs, dev_ant_y, \
te_ant_strings, te_ant_pairs, te_ant_y = create_train_dev_test(word_strings, word_vectors, class_count, ant_class, -1)
len(tr_ant_pairs)
def fit_model(tr_data, output_size, optimizer, batch_size, epochs, reg_rate, deviation_dropoff):
# network definition
base_network = create_base_network(input_shape, output_size, reg_rate)
input_a = Input(shape=input_shape)
input_b = Input(shape=input_shape)
# because we re-use the same instance `base_network`,
# the weights of the network
# will be shared across the two branches
processed_a = base_network(input_a)
processed_b = base_network(input_b)
distance = Lambda(keras_similarity,
output_shape=sim_output_shape)([processed_a, processed_b])
model = Model([input_a, input_b], distance)
def contrastive_sim_loss(y_true, y_pred):
return K.mean(-y_true * y_pred +
K.cast(K.equal(y_true, 0), 'float32') * (K.abs(y_pred) ** deviation_dropoff))
# train
if optimizer == 'adam':
opt = Adam()
else:
raise ValueError("unknown optimizer")
model.compile(loss=contrastive_sim_loss, optimizer=opt)
for tr_pairs, tr_y in tr_data:
model.fit([tr_pairs[:, 0], tr_pairs[:, 1]], tr_y,
batch_size=batch_size,
epochs=epochs,
verbose=0)
return model
def eval_model(model):
y_pred = model.predict([tr_syn_pairs[:, 0], tr_syn_pairs[:, 1]])
syn_tr_acc = syn_accuracy(tr_syn_y, y_pred)
y_pred = model.predict([dev_syn_pairs[:, 0], dev_syn_pairs[:, 1]])
syn_dev_acc = syn_accuracy(dev_syn_y, y_pred)
y_pred = model.predict([tr_ant_pairs[:, 0], tr_ant_pairs[:, 1]])
ant_tr_acc = ant_accuracy(tr_ant_y, y_pred)
y_pred = model.predict([dev_ant_pairs[:, 0], dev_ant_pairs[:, 1]])
ant_dev_acc = ant_accuracy(dev_ant_y, y_pred)
print("Accuracy:")
print("| | Training Set | Dev Set |")
print("|--------:|-------------:|--------:|")
print("| Synonym | {:7.2%} | {:7.2%} |".format(syn_tr_acc, syn_dev_acc))
print("| Antonym | {:7.2%} | {:7.2%} |".format(ant_tr_acc, ant_dev_acc))
return syn_tr_acc, syn_dev_acc, ant_tr_acc, ant_dev_acc
def map_word_vectors(word_vectors, model):
base_network = model.layers[2]
l1_weights = base_network.layers[1].get_weights()[0]
l1_biases = base_network.layers[1].get_weights()[1]
mapped = word_vectors @ l1_weights + l1_biases
mapped_2 = (l1_weights.T @ word_vectors.T + l1_biases.reshape((-1, 1))).T
assert np.isclose(mapped, mapped_2).all()
return mapped
syn_tr_acc = syn_accuracy(
tr_syn_y, np.array([similarity(p[0], p[1]) for p in tr_syn_pairs])
)
syn_dev_acc = syn_accuracy(
dev_syn_y, np.array([similarity(p[0], p[1]) for p in dev_syn_pairs])
)
ant_tr_acc = ant_accuracy(
tr_ant_y, np.array([similarity(p[0], p[1]) for p in tr_ant_pairs])
)
ant_dev_acc = ant_accuracy(
dev_ant_y, np.array([similarity(p[0], p[1]) for p in dev_ant_pairs])
)
print('Baseline...')
print("Accuracy:")
print("| | Training Set | Dev Set |")
print("|--------:|-------------:|--------:|")
print("| Synonym | {:7.2%} | {:7.2%} |".format(syn_tr_acc, syn_dev_acc))
print("| Antonym | {:7.2%} | {:7.2%} |".format(ant_tr_acc, ant_dev_acc))
print('Training on synonyms...')
syn_model = fit_model([(tr_syn_pairs, tr_syn_y)], 140, 'adam', 256, 30, 1e-09, 6)
eval_model(syn_model)
print('Training on antonyms...')
ant_model = fit_model([(tr_ant_pairs, tr_ant_y)], 140, 'adam', 256, 30, 1e-09, 6)
eval_model(ant_model)
print('Training on synonyms & antonyms...')
both_model = fit_model([(np.vstack((tr_syn_pairs, tr_ant_pairs)), np.hstack((tr_syn_y, tr_ant_y)))], 140, 'adam', 256, 30, 1e-09, 6)
eval_model(both_model)
print('Training on synonyms, then antonyms...')
seq_model = fit_model([(tr_syn_pairs, tr_syn_y), (tr_ant_pairs, tr_ant_y)], 140, 'adam', 256, 30, 1e-09, 6)
eval_model(seq_model)
print('')
space = [
skopt.space.Integer(1, 300, name='output_size'),
skopt.space.Categorical(['adam'], name='optimizer'),
skopt.space.Categorical([128, 256, 512], name='batch_size'),
skopt.space.Categorical([30], name='epochs'),
skopt.space.Real(0.000000001, 1000, prior='log-uniform', name='reg_rate'),
skopt.space.Integer(1, 50, name='deviation_dropoff'),
]
@skopt.utils.use_named_args(space)
def objective(output_size, optimizer, batch_size, epochs, reg_rate, deviation_dropoff):
start_time = datetime.datetime.now()
model = fit_model([(tr_syn_pairs, tr_syn_y)], output_size, optimizer, batch_size, epochs, reg_rate, deviation_dropoff)
print('* Optimization took {:.0f} seconds'.format((datetime.datetime.now() - start_time).total_seconds()))
syn_tr_acc, syn_dev_acc, ant_tr_acc, ant_dev_acc = eval_model(model)
print('')
return -syn_dev_acc
# Authenticate and create the PyDrive client.
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
checkpoint_filename = "{}-checkpoint.pkl".format(time.asctime())
checkpoint_filepath = os.path.join(os.curdir, checkpoint_filename)
def backup(res):
skopt.dump(res, checkpoint_filepath)
uploaded = drive.CreateFile({'title': checkpoint_filename})
uploaded.SetContentFile(checkpoint_filepath)
uploaded.Upload()
print('Uploaded file with ID {}'.format(uploaded.get('id')))
x0 = [270, 'adam', 256, 30, 0.01, 14]
res = skopt.gp_minimize(objective, space, x0=x0, n_calls=25, callback=[backup])
res.x
def get_pair_indices_to_display():
#baseline_sim = np.array([similarity(p[0], p[1]) for p in tr_syn_pairs])
#baseline_scores = syn_accuracy_scores(tr_syn_y, baseline_sim)
optimized_sim = syn_model.predict([tr_syn_pairs[:, 0], tr_syn_pairs[:, 1]])
optimized_scores = syn_accuracy_scores(tr_syn_y, optimized_sim)
display_scores = (tr_syn_y == 1) * optimized_scores
best_for_display = heapq.nlargest(50, zip(display_scores, range(len(display_scores))))
return [index for improvement, index in best_for_display]
def plot_synonym_pairs(word_vectors, pairs):
added = set()
strings = []
vectors = []
for word1, word2 in pairs:
if word1 not in added and word2 not in added:
for word in (word1, word2):
added.add(word)
strings.append(word)
vectors.append(word_vectors[index_for_word[word]])
tsne = TSNE(init='pca')
new_values = tsne.fit_transform(vectors)
plt.figure(figsize=(16, 16))
for i in range(0, len(new_values), 2):
random.seed(i * 293778 + 581580)
color = (random.random(), random.random(), random.random())
string1, string2 = strings[i:i+2]
point1, point2 = new_values[i:i+2]
plt.scatter(point1[0], point1[1], c=[color])
plt.annotate(string1,
xy=(point1[0], point1[1]),
xytext=(2, 2),
textcoords='offset points',
ha='left',
va='bottom')
plt.scatter(point2[0], point2[1], c=[color])
plt.annotate(string2,
xy=(point2[0], point2[1]),
xytext=(-2, -2),
textcoords='offset points',
ha='right',
va='top')
random.seed()
plt.show()
indices_to_display = get_pair_indices_to_display()
synonym_pairs = tr_syn_strings[indices_to_display]
syn_mapped_word_vectors = map_word_vectors(word_vectors, syn_model)
plot_synonym_pairs(word_vectors, synonym_pairs)
plot_synonym_pairs(syn_mapped_word_vectors, synonym_pairs)
```
| github_jupyter |
```
%matplotlib inline
import numpy as np
from sklearn import datasets
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score, precision_score, recall_score, confusion_matrix, classification_report
import torch.nn as nn
import torch.optim as optim
from torch import Tensor
import torch
class Object(object): pass
```
# Intro to Pytorch
```
t = torch.Tensor(np.array([1, 2, 3, 3, 3, 5]))
t = torch.Tensor(np.random.randn(2, 3))
t
torch.max(t, dim=1)
t[1:7]
```
Softmax normalizes a vector of K numbers into a probability distribution proportional to the exponents - a categorical distribution, i.e. a distribution over K possible outcomes.
```
torch.softmax(Tensor([-1, 2.5, 0.5]), dim=0)
```
# MNIST with Pytorch
```
t_mnist = Object()
t_mnist.RAND_STATE = 12
t_mnist.raw = datasets.fetch_mldata("MNIST original")
t_mnist.X = t_mnist.raw.data.reshape(-1, 28, 28).astype(np.float32)
t_mnist.y = t_mnist.raw.target.astype(np.long)
def print_image(x, y, n=None):
if n is None:
n = np.random.randint(x.shape[0])
print(y[n])
_ = plt.imshow(x[n], cmap="gray")
print_image(t_mnist.X, t_mnist.y)
```
<img src="mnist.png" alt="" style="width: 60%"/>
```
t_mnist.X_train, t_mnist.X_test, t_mnist.y_train, t_mnist.y_test = \
train_test_split(t_mnist.X, t_mnist.y, test_size=0.2, stratify=t_mnist.y,
random_state=t_mnist.RAND_STATE)
t_mnist.X_train.shape, t_mnist.X_test.shape, t_mnist.y_train.shape, t_mnist.y_test.shape
```
## Softmax
```
torch.softmax(Tensor([-1, 2.5, 0.5]), 0)
t_mnist.loss = nn.NLLLoss
```
## Convert data
```
? nn.Conv2d
t_mnist.X_train.shape
t_mnist.X_train_ch = np.expand_dims(t_mnist.X_train, axis=1)
plt.imshow(t_mnist.X_train[44354], cmap="gray")
plt.imshow(t_mnist.X_train_ch[44354][0], cmap="gray")
t_mnist.X_train_t = torch.from_numpy(np.expand_dims(t_mnist.X_train, axis=1))
t_mnist.X_test_t = torch.from_numpy(np.expand_dims(t_mnist.X_test, axis=1))
t_mnist.y_train_t = torch.from_numpy(t_mnist.y_train)
t_mnist.y_test_t = torch.from_numpy(t_mnist.y_test)
t_mnist.y_train_t
t_mnist.X_train_t.shape
```
## Training!
## Explain gradient and training
1. Loss function is a function.
1. Forward pass.
1. Calculate output -> error.
1. Gradient of a loss function gives how we should update weights.
```
def accuracy(y_pred_one_hot, y_true):
y_pred = y_pred_one_hot.max(dim=1)[1]
return (y_pred == y_true).sum().item() / y_true.shape[0]
```
## Batches
# LetNet
```
def create_lenet_classic_2():
return nn.Sequential(
#28x28x1
nn.Conv2d(1, 12, kernel_size=9, stride=1, padding=4),
nn.ReLU(),
#28x28x12
nn.MaxPool2d(kernel_size=(2,2), stride=2, padding=0),
#14x14x12
nn.Conv2d(12, 24, kernel_size=(7,7), stride=1, padding=2),
nn.ReLU(),
#12x12x24
nn.MaxPool2d(kernel_size=(2,2), stride=2, padding=0),
#6x6x24
nn.Flatten(),
nn.Linear(6*6*24, 100),
nn.ReLU(),
nn.Linear(100, 10),
nn.LogSoftmax(dim=-1)
)
def create_lenet_classic():
return nn.Sequential(
#28x28x1
nn.Conv2d(1, 6, kernel_size=9, stride=1, padding=4),
nn.ReLU(),
#28x28x6
nn.MaxPool2d(kernel_size=(2,2), stride=2, padding=0),
#14x14x6
nn.Conv2d(6, 16, kernel_size=(7,7), stride=1, padding=3),
nn.ReLU(),
#14x14x16
nn.MaxPool2d(kernel_size=(2,2), stride=2, padding=0),
#7x7x16
nn.Flatten(),
nn.Linear(7*7*16, 100),
nn.ReLU(),
nn.Linear(100, 10),
nn.LogSoftmax(dim=-1)
)
# ? nn.Conv2d
t_mnist.learning_rate = 1e-4
t_mnist.loss = nn.NLLLoss()
t_mnist.optim = torch.optim.SGD(t_mnist.nn.parameters(), lr=t_mnist.learning_rate)
def train_batch(net, data, epochs=3):
optim = torch.optim.SGD(net.parameters(), 1e-4, momentum=0.9)
batch_size = 16
batches = int(data.X_train_t.shape[0] / batch_size)
for i in range(epochs):
for b in range(batches):
start = b * batch_size
end = start + batch_size
# print(start, end)
x_batch = data.X_train_t[start : end]
y_batch = data.y_train_t[start : end]
# Forward pass: compute predicted y by passing x to the model.
# print(x_batch.shape)
y_pred = net(x_batch)
# Compute and print loss.
loss = data.loss(y_pred, y_batch)
# Reset gradient
net.zero_grad()
# Backward pass: compute gradient of the loss with respect to model
# parameters
loss.backward()
# Calling the step function on an Optimizer makes an update to its
# parameters
optim.step()
if True: # i % 1 == 1:
print(i, "Loss:", loss.item())
print(i, "Test accuracy: ", accuracy(net(data.X_test_t), data.y_test_t))
%%time
t_mnist.nn = create_lenet_classic()
train_batch(t_mnist.nn, t_mnist, epochs=1)
```
Test the accuracy on a shifted image:
```
accuracy(t_mnist.nn(t_mnist.X_test_t), t_mnist.y_test_t)
```
# Predict a Single Image
```
t_mnist.test_n = 2625
t_mnist.X_test[t_mnist.test_n].shape
plt.imshow(t_mnist.X_test[t_mnist.test_n], cmap="gray")
t_mnist.test_image = np.expand_dims(np.expand_dims(t_mnist.X_test[t_mnist.test_n], axis=0), axis=0)
t_mnist.test_image.shape
t_mnist.test_image_t = torch.from_numpy(t_mnist.test_image)
logits = t_mnist.nn(t_mnist.test_image_t)
logits
torch.exp(logits)
torch.argmax(torch.exp(logits[0]))
def predict_image(net, x, y, n=None):
if n is None:
n = np.random.randint(x.shape[0])
image = np.expand_dims(np.expand_dims(x[n], axis=0), axis=0).astype(np.float32)
image_t = torch.from_numpy(image)
pred_t = net(image_t)
pred = torch.argmax(torch.exp(pred_t[0])).item()
print(n, ':', y[n], ' : ', pred)
plt.imshow(x[n], cmap="gray")
predict_image(t_mnist.nn, t_mnist.X_test, t_mnist.y_test)
def find_wrong_image(net, x, y):
while True:
n = np.random.randint(x.shape[0])
image = np.expand_dims(np.expand_dims(x[n], axis=0), axis=0).astype(np.float32)
image_t = torch.from_numpy(image)
pred_t = net(image_t)
pred = torch.argmax(torch.exp(pred_t[0])).item()
if pred != y[n]:
print(n, ':', y[n], ' : ', pred)
plt.imshow(x[n], cmap="gray")
break
find_wrong_image(t_mnist.nn, t_mnist.X_test, t_mnist.y_test)
```
# Predict own image
```
from PIL import Image
digit = np.array(Image.open('images/digit.png')).astype(np.float32)
digit = digit[:,:,0]
digit.shape
predict_image(t_mnist.nn, [digit], [3], n=0)
```
# Redis AI
1. docker run -p 6379:6379 -it --rm redisai/redisai /bin/bash
1. /usr/local/bin//redis-server --loadmodule /usr/lib/redis/modules/redisai.so &
1. redis-cli
1. AI.CONFIG LOADBACKEND TORCH redisai_torch/redisai_torch.so
* https://oss.redislabs.com/redisai/commands/
* https://oss.redislabs.com/redisai/
Use Redisai.py to upload the model to Redis.
```
import ml2rt
t_mnist.nn.eval()
t_mnist.nn_script = torch.jit.trace(t_mnist.nn, t_mnist.X_test_t)
```
We can use TorchScript model just like any other PyTorch model:
```
predict_image(t_mnist.nn_script, [digit], [3], n=0)
```
Save the TorchScript model:
```
t_mnist.nn_script.eval() # Important?
t_mnist.nn_script.save('lenet.pt')
t_mnist.loaded_nn_script = torch.jit.load('lenet.pt')
ml2rt.save_torch(t_mnist.nn_script, 'model.pt')
t_mnist.loaded_nn = ml2rt.load_model('model.pt')
import redisai
t_mnist.redis = redisai.Client()
t_mnist.redis.loadbackend('TORCH', 'redisai_torch/redisai_torch.so')
t_mnist.redis.modelset('lenet', redisai.Backend.torch, redisai.Device.cpu, t_mnist.loaded_nn)
```
Send an image for recognition:
```
t_mnist.redis.tensorset('image',
np.expand_dims(
np.expand_dims(digit, axis=0),
axis=0))
t_mnist.redis.modelrun('lenet', ['image'], ['pred'])
np.argmax(np.exp(t_mnist.redis.tensorget('pred')))
```
Write a function that submits a digit for prediction:
```
def predict_redis(client, x, y, n=None):
assert isinstance(client, redisai.Client)
if n is None:
n = np.random.randint(x.shape[0])
image = x[n]
image = np.expand_dims(image, axis=0)
image = np.expand_dims(image, axis=0)
client.tensorset('image', image)
client.modelrun('lenet', ['image'], ['pred'])
pred = client.tensorget('pred')
pred = np.argmax(pred)
print("Pred: ", pred, "actual:", y[n])
plt.imshow(x[n], cmap="gray")
predict_redis(t_mnist.redis, t_mnist.X_test, t_mnist.y_test)
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import pydicom
%matplotlib inline
import matplotlib.pyplot as plt
from tensorflow.keras.models import model_from_json, load_model
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.layers import GlobalAveragePooling2D, Dense, Dropout, Flatten, Conv2D, MaxPooling2D, BatchNormalization
from tensorflow.keras.models import Sequential, Model
from tensorflow.keras.applications.vgg16 import VGG16
from tensorflow.keras.applications.resnet import ResNet50
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.callbacks import ModelCheckpoint, LearningRateScheduler, EarlyStopping, ReduceLROnPlateau
from skimage.transform import rescale, resize, downscale_local_mean
def load_pretrained_model():
model = VGG16(include_top=False, weights='imagenet', input_shape = (512, 512, 3))
return model
def build_my_model():
# my_model = Sequential()
# ....add your pre-trained model, and then whatever additional layers you think you might
# want for fine-tuning (Flatteen, Dense, Dropout, etc.)
# if you want to compile your model within this function, consider which layers of your pre-trained model,
# you want to freeze before you compile
# also make sure you set your optimizer, loss function, and metrics to monitor
vgg_model = load_pretrained_model()
for layer in vgg_model.layers[:17]:
layer.trainable = False
my_model = Sequential()
my_model.add(vgg_model)
my_model.add(GlobalAveragePooling2D())
my_model.add(Dense(256, activation='relu'))
my_model.add(BatchNormalization())
my_model.add(Dropout(0.4))
my_model.add(Dense(64, activation='relu'))
my_model.add(BatchNormalization())
my_model.add(Dropout(0.3))
my_model.add(Dense(32, activation='relu'))
my_model.add(BatchNormalization())
my_model.add(Dropout(0.3))
my_model.add(Dense(1, activation='sigmoid'))
return my_model
# This function reads in a .dcm file, checks the important fields for our device, and returns a numpy array
# of just the imaging data
def check_dicom(filename):
print('Load file {} ...'.format(filename))
ds = pydicom.dcmread(filename)
if(ds.Modality == 'DX' and ds.PatientPosition in ['PA', 'AP'] and (ds.BodyPartExamined == 'CHEST')):
img = ds.pixel_array
return img
print('Invalid data...')
return None
# This function takes the numpy array output by check_dicom and
# runs the appropriate pre-processing needed for our model input
def preprocess_image(img, img_size):
img = resize(img, (img_size[1], img_size[2]))
proc_img = (img - img.mean()) / img.std()
return np.resize(proc_img, img_size)
# This function loads in our trained model w/ weights and compiles it
def load_model(model_path):
my_model = build_my_model()
my_model.load_weights(model_path)
return my_model
# This function uses our device's threshold parameters to predict whether or not
# the image shows the presence of pneumonia using our trained model
def predict_image(model, img, thresh):
prediction = model(img)
if(prediction > thresh):
prediction = 'Pneumonia'
else:
prediction = "No Pneumonia"
return prediction
test_dicoms = ['test1.dcm','test2.dcm','test3.dcm','test4.dcm','test5.dcm','test6.dcm']
model_path = r'xray_class_my_model.h5'
IMG_SIZE=(1,512,512,3) # This might be different if you did not use vgg16
my_model = load_model(model_path)
thresh = 0.25
# # use the .dcm files to test your prediction
for i in test_dicoms:
img = check_dicom(i)
if img is None:
continue
img_proc = preprocess_image(img, IMG_SIZE)
pred = predict_image(my_model,img_proc,thresh)
print(pred)
```
| github_jupyter |
```
### If save data:
save_data = False
if save_data:
def ensure_dir(file_path):
import os
directory = os.path.dirname(file_path)
if not os.path.exists(directory):
os.makedirs(directory)
from google.colab import drive
drive.mount('/content/drive')
%cd drive/My Drive/
save_dir = "data/"
ensure_dir(save_dir)
```
# Import Libraries and Neural Tangents
```
import os
import sys
import numpy as npo
import matplotlib.pyplot as plt
import jax.numpy as np
import jax
from jax.config import config
from jax import random
from jax import jit, grad, vmap
from jax.experimental import optimizers
!pip install -q git+https://www.github.com/google/neural-tangents
import neural_tangents as nt
from neural_tangents import stax
!pip install -q git+https://github.com/Pehlevan-Group/kernel-generalization
from kernel_generalization import kernel_simulation as ker_sim
from kernel_generalization import kernel_spectrum as ker_spec
from kernel_generalization.utils import gegenbauer
import urllib.request
dir_gegenbauer = os.path.join(os.getcwd(),'GegenbauerEigenvalues.npz')
url = "https://raw.github.com/Pehlevan-Group/kernel-generalization/main/PrecomputedData/GegenbauerEigenvalues.npz"
urllib.request.urlretrieve(url, dir_gegenbauer)
import pytz
from datetime import datetime
from dateutil.relativedelta import relativedelta
def time_now():
return datetime.now(pytz.timezone('US/Eastern')).strftime("%m-%d_%H-%M")
def time_diff(t_a, t_b):
t_diff = relativedelta(t_b, t_a) # later/end time comes first!
return '{h}h {m}m {s}s'.format(h=t_diff.hours, m=t_diff.minutes, s=t_diff.seconds)
### Using Time difference
### t_start = datetime.now(pytz.timezone('US/Eastern'))
### t_end = datetime.now(pytz.timezone('US/Eastern'))
### print('Time Elapsed: ' + time_diff(t_start,t_end))
print('After running code: ',jax.devices())
```
# First Define Your Plot Function
```
def plot_shift_data(pvals, error, std, Errs_tot, noise_k, noise, exp_type, tr_errs = None):
i = noise_k
shift = len(noise[i])
colors = ['b', 'g', 'r', 'c', 'm', 'y', 'k']
for j in range(shift):
plt.errorbar(pvals, error[i,j], std[i,j], fmt='o', label = r'$\sigma = %.3f$' % noise[i,j], color=colors[j])
plt.plot(pvals, Errs_tot[j,:,i], color=colors[j])
if exp_type == 'kernel':
plt.title('Interpolating Kernel Regression (Infinite Parameters)')
else:
plt.title('Neural Network Training')
plt.xlabel('samples')
plt.ylabel('test risk')
plt.xscale('log')
plt.yscale('log')
plt.legend()
plt.show()
```
# Functions for Data generating and Neural Networks
```
# generate sphere vectors
def generate_synth_data(p, dim, key):
x0 = random.normal(key, shape=(p,dim))
x = x0 / npo.outer(npo.linalg.norm(x0, axis=1), npo.ones(dim))
return np.array(x)
## Target function is y(x) = Q_k(beta X)
## It has a single mode and all weights for degenerate modes are the same
def pure_target_fn(X, dim, beta, spectrum, k):
dim = len(beta)
lamb = (dim-2)/2
## Q(1) from the SI
Q1 = lamb/(lamb+k)*gegenbauer.degeneracy_kernel(dim, k)
z = npo.dot(X, beta)
d = beta.shape[0]
y = gegenbauer.gegenbauer(z, dim, d)[k,:]
# Calculate normalized y:
y_norm = y*spectrum[k]*((k+lamb)/lamb)
return np.array(y_norm)
# Generate fully connected NN architecture
def fully_connected(num_layers, width):
layers = []
for i in range(num_layers):
layers += [stax.Dense(width, W_std = 1, b_std = 0), stax.Relu()]
layers += [stax.Dense(1, W_std = 1, b_std=0)]
return stax.serial(*layers)
```
# Functions for Theory Curves and Experiments
## Theory Curves and NTK Spectrum
```
def NTK_spectrum(layers, dim, kmax):
dimension = npo.array([5*(i+1) for i in range(40)])
degree = npo.array([i for i in range(500)]);
layer = npo.array([1, 2, 3, 4, 5, 6, 7, 8, 9]);
dim_index = npo.where(dimension == dim)[0][0]
lambda_bar, spectrum = ker_spec.ntk_spectrum(dir_gegenbauer, kmax, layer = layers-1, dim = dim_index)
return lambda_bar, spectrum
def NTK_theory_and_Noise(pvals, layers, dim, kmax, noise_num, shift, R_tilde = 1, fn_mode = 1, zero_mode = False):
## Generate spectrum and degeneracies
k = npo.linspace(0,kmax-1,kmax)
a = (dim-2)/2
kp = ((k+a)/a)
lambda_bar, spectrum = NTK_spectrum(layers, dim, kmax)
degens = npo.array([gegenbauer.degeneracy_kernel(dim, k) for k in range(kmax)])
alpha_s = npo.array([(np.sum(lambda_bar[i+1:]))/lambda_bar[i+1] for i in range(kmax-2)])
noise = npo.zeros((noise_num, len(shift)))
for i in range(noise_num):
for j in range(len(shift)):
if alpha_s[i] >= 2:
alpha = (alpha_s[i]-1)*(1+shift[j])
noise[i,j] = (2*alpha+1)*spectrum[i+1]**2*degens[i+1]
else:
alpha = (alpha_s[i]-1)*(1+shift[j])+1
noise[i,j] = ker_sim.noise(alpha)*spectrum[i+1]**2*degens[i+1]
Sol = npo.zeros((len(shift), len(pvals), kmax, noise_num))
Errs_tot = npo.zeros((len(shift), len(pvals), noise_num))
Cum_gen_errs = npo.zeros((len(shift), len(pvals), kmax, noise_num))
for i in range(len(shift)):
Sol[i], Errs_tot[i], Cum_gen_errs[i]= ker_sim.simulate_pure_gen_error(pvals, spectrum, degens,
noise[:, i], pure_mode = fn_mode,
zero_mode=zero_mode)
## Scale the theoretical error with test radius (R_train = 1)
Errs_tot = Errs_tot * R_tilde**2
return Sol, Errs_tot, noise, alpha_s, spectrum, degens
```
## Kernel Regression Functions
```
### Kernel Regression Experiment Synthetic Data ###
### The function generates a pure mode target function and
### performs kernel regression on a given kernel_fn with
### randomly sampled training and test data on unit "dim"-sphere.
def kernel_expt(kernel_fn, pvals, p_test, sigma, dim, spectrum, num_repeats, all_keys, R_tilde = 1, fn_mode = 1):
## First of all we fix a target function by generating a
## projection vector "beta" in "dim" dimensions and specifying
## the function's mode index "fn_mode" (fn_mode = 1, linear target)
beta = generate_synth_data(1, dim, all_keys[0,:])[0,:]
gen_errs = npo.zeros((len(pvals), num_repeats))
for i, p in enumerate(pvals):
for j in range(num_repeats):
# Generate fresh training and test data
tr_key, test_key, noise_key = random.split(all_keys[j,:], 3)
# Generate random inputs and label noise
X = generate_synth_data(p, dim, tr_key)
## Change the test radius
X_test = generate_synth_data(p_test, dim, test_key)*R_tilde
label_noise = npo.sqrt(sigma)*random.normal(noise_key, shape=(p,1))
# Calculate corresponding labels given the weights beta and fn_mode
y = pure_target_fn(X, dim, beta, spectrum, fn_mode).reshape(p,1) + label_noise
y_test = pure_target_fn(X_test, dim, beta, spectrum, fn_mode).reshape(p_test,1)
## Perform regression using NeuralTangents Package
predict_fn = nt.predict.gradient_descent_mse_ensemble(kernel_fn, X, y, diag_reg=1e-15)
yhat = predict_fn(x_test=X_test, get='ntk', compute_cov=False)
## Calculate MSE
gen_errs[i,j] = npo.mean((y_test-yhat)**2)
## Calculate repeat averaged errors and standard deviation
errs = npo.mean(gen_errs, axis = 1)
std_errs = npo.std(gen_errs, axis = 1)
return errs, std_errs
```
## Neural Network Training Functions
```
# training NN here
def train_nn(key, init_fn, apply_fn, nn_loss, grad_loss, optimizer, X_test, train_set, num_iter):
(X, y) = train_set
losses = []
_,params = init_fn(key, (-1, X.shape[1]))
opt_init, opt_update, get_params = optimizer
opt_state = opt_init(params)
for t in range(num_iter):
opt_state = opt_update(t, grad_loss(opt_state, *train_set), opt_state)
loss = nn_loss(get_params(opt_state), *train_set)
losses.append(loss)
params = get_params(opt_state)
yhat_ensemble = apply_fn(params, X_test)
losses = np.array(losses)
return opt_state, losses, yhat_ensemble
# experiment for sample wise generalization error
# for overparameterized model
def nn_expt(init_fn, apply_fn, pvals, p_test, sigma, dim, spectrum, num_repeats, all_keys, lr, num_iter, fn_mode = 1, ensemble_size = 1, verbose = True):
## First of all we fix a target function by generating a
## projection vector "beta" in "dim" dimensions and specifying
## the function's mode index "fn_mode" (fn_mode = 1, linear target)
beta = generate_synth_data(1, dim, all_keys[0,:])[0,:]
gen_errs = npo.zeros((len(pvals), num_repeats))
training_errs = []
training_errs_std = []
beginning = datetime.now()
for i, p in enumerate(pvals):
t_start = datetime.now()
train_errs_p = []
for j in range(num_repeats):
# Generate fresh training and test data
tr_key, test_key, noise_key = random.split(all_keys[j,:], 3)
# Generate random inputs and label noise
X_train = generate_synth_data(p, dim, tr_key)
X_test = generate_synth_data(p_test, dim, test_key)
label_noise = npo.sqrt(sigma)*random.normal(noise_key, shape=(p,1))
# Calculate corresponding labels given the weights beta and fn_mode
y_train = pure_target_fn(X_train, dim, beta, spectrum, fn_mode).reshape(p,1) + label_noise
y_test = pure_target_fn(X_test, dim, beta, spectrum, fn_mode).reshape(p_test,1)
train_set = (X_train, y_train)
test_set = (X_test, y_test)
if p > 1000:
num_iter = 2000
Ktr = kernel_fn(X_train,None,'ntk')
optimizer = optimizers.adam(lr)
# lr_max = nt.predict.max_learning_rate(Ktr)/10
# optimizer = optimizers.sgd(lr_max)
opt_init, opt_update, get_params = optimizer
nn_loss = jit(lambda params, X, y: np.mean((apply_fn(params, X) - y)**2))
grad_loss = jit(lambda state, x, y: grad(nn_loss)(get_params(state), x, y))
ensemble_key = random.split(all_keys[j,:], ensemble_size)
output = vmap(train_nn, (0, None, None, None, None, None, None, None, None))(ensemble_key, init_fn, apply_fn, nn_loss, grad_loss,
optimizer, X_test, train_set, num_iter)
opt_state, train_losses, yhat_ensemble = output
## Perfom ensamble averaging
train_errs_p += [np.mean(train_losses, axis=0)]
yhat = np.mean(yhat_ensemble, axis=0)
gen_errs[i,j] = np.mean((yhat - y_test)**2)
sys.stdout.write("\r P = %d, gen error: %0.4f | train error: %0.3e | Repeat: %d/%d"
% (p, gen_errs[i,j], train_errs_p[j][-1], j+1, num_repeats))
t_end = datetime.now()
# For each P average Gen Error and Training Error over trials
err = npo.mean(gen_errs[i], axis = 0)
std_err = npo.std(gen_errs[i], axis = 0)
tr_err = npo.mean(np.array(train_errs_p), axis = 0)
tr_err_std = npo.std(np.array(train_errs_p), axis = 0)
training_errs += [tr_err]
training_errs_std += [tr_err_std]
if verbose:
string_print = "\r P = %d, gen error: %0.4f | train error: %0.3e | Time Elapsed: " % (p, err, tr_err[-1])
sys.stdout.write(string_print + time_diff(t_start,t_end) + "\n")
itr = npo.linspace(1,num_iter,num_iter)
plt.errorbar(npo.log10(itr), npo.log10(tr_err), tr_err_std/tr_err, fmt = 'o', elinewidth = 0.2)
plt.show()
ending = datetime.now()
print('Total Time Elapsed: ' + time_diff(beginning,ending))
return npo.mean(gen_errs, axis = 1), npo.std(gen_errs, axis = 1), training_errs, training_errs_std
```
# Experiments
```
timestamp = time_now()
import warnings
warnings.simplefilter(action='ignore', category=FutureWarning)
warnings.simplefilter(action='ignore', category=UserWarning)
## Experiment Parameters
num_repeats_ker = 15 #150
num_repeats_nn = 5 #5
num_iter_nn = 500
## NN Hyperparameters
lr = 0.008
width = 2000
layers = 1
ensemble_size_list = [20]
## Dimension, sample sizes, max eigenvalue mode
dim = 30
pvals = npo.logspace(0.5, 4, num = 20).astype('int')
p_test = 1500
kmax = 200
## Target function mode and label noise
fn_mode = 1
noise_num = 2
shift = npo.array([-0.8, 0, 1])
R_tilde = 0.5
key = random.PRNGKey(0)
all_keys = random.split(key, max(num_repeats_nn, num_repeats_ker))
if save_data:
parent = save_dir + 'NNExp_width_%d-layer_%d-fn_mode_%d/'%(width,layers,fn_mode)
ensure_dir(parent)
# create NN and kernel function
init_fn, apply_fn, kernel_fn = fully_connected(layers, width)
apply_fn = jit(apply_fn)
kernel_fn = jit(kernel_fn, static_argnums=(2,))
# Generate Theory curves and Noise
Sol, Errs_tot, noise, alpha_s, spectrum, degens = NTK_theory_and_Noise(pvals, layers, dim, kmax, noise_num, shift,
R_tilde, fn_mode = fn_mode, zero_mode=False)
params = {'dim': dim, 'kmax': kmax, 'layer': layers, 'width': width,
'test_samp': p_test, 'num_repeat_ker': num_repeats_ker,
'num_repeat_nn': num_repeats_nn, 'num_iter_nn': num_iter_nn,
'noise': noise, 'noise_num': noise_num, 'shift': shift,
'ensemble_size_list': ensemble_size_list, 'fn_mode': fn_mode,
'alpha_s': alpha_s, 'degens': degens}
if save_data:
filename_theory = parent + timestamp + 'error_theory.npz'
np.savez(filename_theory, pvals, Sol, Errs_tot, params)
### Start the kernel experiment
err_regression = npo.zeros((noise_num, len(shift), len(pvals)))
std_regression = npo.zeros((noise_num, len(shift), len(pvals)))
print('Kernel Regression Start @ ' + time_now())
for i in range(noise_num):
t_start = datetime.now()
for j in range(len(shift)):
data = kernel_expt(kernel_fn, pvals, p_test, noise[i, j], dim, spectrum, num_repeats_ker, all_keys, R_tilde, fn_mode = fn_mode)
err_regression[i, j] = data[0]
std_regression[i, j] = data[1]
t_end = datetime.now()
print('Time Elapsed: ' + time_diff(t_start,t_end))
plot_shift_data(pvals, err_regression, std_regression, Errs_tot, i, noise, 'kernel')
if save_data:
filename_exp = directory + timestamp + 'final_error_exp_NN.npz'
np.savez(filename_exp, err_NN, std_NN, tr_err_NN, std_tr_err_NN)
i = 1
shift = len(noise[i])
colors = ['b', 'g', 'r', 'c', 'm', 'y', 'k']
for j in range(shift):
plt.errorbar(pvals, err_regression[i,j], std_regression[i,j], fmt='o', label = r'$\sigma = %.1e$' % np.sqrt(noise[i,j]), color=colors[j])
plt.plot(pvals, Errs_tot[j,:,i], color=colors[j])
plt.title('NTK Regression with $\lambda = 0$')
plt.ylim([1e-6, 1e-1])
plt.xlabel('$P$', fontsize = 20)
plt.ylabel('$E_g(P)$', fontsize = 20)
plt.xscale('log')
plt.yscale('log')
plt.legend()
plt.tight_layout()
plt.savefig('NTK_regression_R_tilde_0_5.pdf')
plt.show()
```
| github_jupyter |
#1. Install Dependencies
First install the libraries needed to execute recipes, this only needs to be done once, then click play.
```
!pip install git+https://github.com/google/starthinker
```
#2. Get Cloud Project ID
To run this recipe [requires a Google Cloud Project](https://github.com/google/starthinker/blob/master/tutorials/cloud_project.md), this only needs to be done once, then click play.
```
CLOUD_PROJECT = 'PASTE PROJECT ID HERE'
print("Cloud Project Set To: %s" % CLOUD_PROJECT)
```
#3. Get Client Credentials
To read and write to various endpoints requires [downloading client credentials](https://github.com/google/starthinker/blob/master/tutorials/cloud_client_installed.md), this only needs to be done once, then click play.
```
CLIENT_CREDENTIALS = 'PASTE CREDENTIALS HERE'
print("Client Credentials Set To: %s" % CLIENT_CREDENTIALS)
```
#4. Enter DV360 Report Emailed To BigQuery Parameters
Pulls a DV360 Report from a gMail email into BigQuery.
1. The person executing this recipe must be the recipient of the email.
1. Schedule a DV360 report to be sent to an email like <b>UNDEFINED</b>.
1. Or set up a redirect rule to forward a report you already receive.
1. The report can be sent as an attachment or a link.
1. Ensure this recipe runs after the report is email daily.
1. Give a regular expression to match the email subject.
1. Configure the destination in BigQuery to write the data.
Modify the values below for your use case, can be done multiple times, then click play.
```
FIELDS = {
'auth_read': 'user', # Credentials used for reading data.
'email': '', # Email address report was sent to.
'subject': '.*', # Regular expression to match subject. Double escape backslashes.
'dataset': '', # Existing dataset in BigQuery.
'table': '', # Name of table to be written to.
'dbm_schema': '[]', # Schema provided in JSON list format or empty list.
'is_incremental_load': False, # Append report data to table based on date column, de-duplicates.
}
print("Parameters Set To: %s" % FIELDS)
```
#5. Execute DV360 Report Emailed To BigQuery
This does NOT need to be modified unles you are changing the recipe, click play.
```
from starthinker.util.project import project
from starthinker.script.parse import json_set_fields
USER_CREDENTIALS = '/content/user.json'
TASKS = [
{
'email': {
'auth': 'user',
'read': {
'from': 'noreply-dv360@google.com',
'to': {'field': {'name': 'email','kind': 'string','order': 1,'default': '','description': 'Email address report was sent to.'}},
'subject': {'field': {'name': 'subject','kind': 'string','order': 2,'default': '.*','description': 'Regular expression to match subject. Double escape backslashes.'}},
'link': 'https://storage.googleapis.com/.*',
'attachment': '.*',
'out': {
'bigquery': {
'dataset': {'field': {'name': 'dataset','kind': 'string','order': 3,'default': '','description': 'Existing dataset in BigQuery.'}},
'table': {'field': {'name': 'table','kind': 'string','order': 4,'default': '','description': 'Name of table to be written to.'}},
'schema': {'field': {'name': 'dbm_schema','kind': 'json','order': 5,'default': '[]','description': 'Schema provided in JSON list format or empty list.'}},
'header': True,
'is_incremental_load': {'field': {'name': 'is_incremental_load','kind': 'boolean','order': 6,'default': False,'description': 'Append report data to table based on date column, de-duplicates.'}}
}
}
}
}
}
]
json_set_fields(TASKS, FIELDS)
project.initialize(_recipe={ 'tasks':TASKS }, _project=CLOUD_PROJECT, _user=USER_CREDENTIALS, _client=CLIENT_CREDENTIALS, _verbose=True, _force=True)
project.execute(_force=True)
```
| github_jupyter |
### Dependencies
```
!pip install keras-rectified-adam
!pip install segmentation-models
import os
import cv2
import math
import random
import shutil
import warnings
import numpy as np
import pandas as pd
import seaborn as sns
import multiprocessing as mp
import albumentations as albu
from keras_radam import RAdam
import matplotlib.pyplot as plt
import segmentation_models as sm
from tensorflow import set_random_seed
from sklearn.model_selection import train_test_split
from keras import optimizers
from keras import backend as K
from keras.utils import Sequence
from keras.losses import binary_crossentropy
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import EarlyStopping, ReduceLROnPlateau, ModelCheckpoint, Callback
def seed_everything(seed=0):
random.seed(seed)
os.environ['PYTHONHASHSEED'] = str(seed)
np.random.seed(seed)
set_random_seed(seed)
seed = 0
seed_everything(seed)
warnings.filterwarnings("ignore")
```
### Load data
```
train = pd.read_csv('../input/understanding_cloud_organization/train.csv')
submission = pd.read_csv('../input/understanding_cloud_organization/sample_submission.csv')
hold_out_set = pd.read_csv('../input/clouds-data-split/hold-out.csv')
X_train = hold_out_set[hold_out_set['set'] == 'train']
X_val = hold_out_set[hold_out_set['set'] == 'validation']
print('Compete set samples:', len(train))
print('Train samples: ', len(X_train))
print('Validation samples: ', len(X_val))
print('Test samples:', len(submission))
# Preprocecss data
train['image'] = train['Image_Label'].apply(lambda x: x.split('_')[0])
submission['image'] = submission['Image_Label'].apply(lambda x: x.split('_')[0])
test = pd.DataFrame(submission['image'].unique(), columns=['image'])
test['set'] = 'test'
display(X_train.head())
```
# Model parameters
```
BACKBONE = 'resnet152'
BATCH_SIZE = 32
EPOCHS = 15
LEARNING_RATE = 3e-4
HEIGHT = 224
WIDTH = 224
CHANNELS = 3
N_CLASSES = 4
ES_PATIENCE = 5
RLROP_PATIENCE = 3
DECAY_DROP = 0.5
model_path = '../working/uNet_%s_%sx%s.h5' % (BACKBONE, HEIGHT, WIDTH)
preprocessing = sm.backbones.get_preprocessing(BACKBONE)
augmentation = albu.Compose([albu.HorizontalFlip(p=0.5),
albu.VerticalFlip(p=0.5),
albu.GridDistortion(p=0.15),
albu.ShiftScaleRotate(rotate_limit=0, p=0.2)
])
```
### Auxiliary functions
```
def np_resize(img, input_shape):
height, width = input_shape
return cv2.resize(img, (width, height))
def mask2rle(img):
pixels= img.T.flatten()
pixels = np.concatenate([[0], pixels, [0]])
runs = np.where(pixels[1:] != pixels[:-1])[0] + 1
runs[1::2] -= runs[::2]
return ' '.join(str(x) for x in runs)
def build_rles(masks, reshape=None):
width, height, depth = masks.shape
rles = []
for i in range(depth):
mask = masks[:, :, i]
if reshape:
mask = mask.astype(np.float32)
mask = np_resize(mask, reshape).astype(np.int64)
rle = mask2rle(mask)
rles.append(rle)
return rles
def build_masks(rles, input_shape, reshape=None):
depth = len(rles)
if reshape is None:
masks = np.zeros((*input_shape, depth))
else:
masks = np.zeros((*reshape, depth))
for i, rle in enumerate(rles):
if type(rle) is str:
if reshape is None:
masks[:, :, i] = rle2mask(rle, input_shape)
else:
mask = rle2mask(rle, input_shape)
reshaped_mask = np_resize(mask, reshape)
masks[:, :, i] = reshaped_mask
return masks
def rle2mask(rle, input_shape):
width, height = input_shape[:2]
mask = np.zeros( width*height ).astype(np.uint8)
array = np.asarray([int(x) for x in rle.split()])
starts = array[0::2]
lengths = array[1::2]
current_position = 0
for index, start in enumerate(starts):
mask[int(start):int(start+lengths[index])] = 1
current_position += lengths[index]
return mask.reshape(height, width).T
def dice_coefficient(y_true, y_pred):
y_true = np.asarray(y_true).astype(np.bool)
y_pred = np.asarray(y_pred).astype(np.bool)
intersection = np.logical_and(y_true, y_pred)
return (2. * intersection.sum()) / (y_true.sum() + y_pred.sum())
def dice_coef(y_true, y_pred, smooth=1):
y_true_f = K.flatten(y_true)
y_pred_f = K.flatten(y_pred)
intersection = K.sum(y_true_f * y_pred_f)
return (2. * intersection + smooth) / (K.sum(y_true_f) + K.sum(y_pred_f) + smooth)
def post_process(probability, threshold=0.5, min_size=10000):
mask = cv2.threshold(probability, threshold, 1, cv2.THRESH_BINARY)[1]
num_component, component = cv2.connectedComponents(mask.astype(np.uint8))
predictions = np.zeros(probability.shape, np.float32)
for c in range(1, num_component):
p = (component == c)
if p.sum() > min_size:
predictions[p] = 1
return predictions
def preprocess_image(image_id, base_path, save_path, HEIGHT=HEIGHT, WIDTH=WIDTH):
image = cv2.imread(base_path + image_id)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
image = cv2.resize(image, (WIDTH, HEIGHT))
cv2.imwrite(save_path + image_id, image)
def preprocess_data(df, HEIGHT=HEIGHT, WIDTH=WIDTH):
df = df.reset_index()
for i in range(df.shape[0]):
item = df.iloc[i]
image_id = item['image']
item_set = item['set']
if item_set == 'train':
preprocess_image(image_id, train_base_path, train_images_dest_path)
if item_set == 'validation':
preprocess_image(image_id, train_base_path, validation_images_dest_path)
if item_set == 'test':
preprocess_image(image_id, test_base_path, test_images_dest_path)
def get_metrics(model, df, df_images_dest_path, tresholds, min_mask_sizes, set_name='Complete set'):
class_names = ['Fish', 'Flower', 'Gravel', 'Sugar']
metrics = []
for class_name in class_names:
metrics.append([class_name, 0, 0])
metrics_df = pd.DataFrame(metrics, columns=['Class', 'Dice', 'Dice Post'])
for i in range(0, df.shape[0], 500):
batch_idx = list(range(i, min(df.shape[0], i + 500)))
batch_set = df[batch_idx[0]: batch_idx[-1]+1]
ratio = len(batch_set) / len(df)
generator = DataGenerator(
directory=df_images_dest_path,
dataframe=batch_set,
target_df=train,
batch_size=len(batch_set),
target_size=(HEIGHT, WIDTH),
n_channels=CHANNELS,
n_classes=N_CLASSES,
preprocessing=preprocessing,
seed=seed,
mode='fit',
shuffle=False)
x, y = generator.__getitem__(0)
preds = model.predict(x)
for class_index in range(N_CLASSES):
class_score = []
class_score_post = []
mask_class = y[..., class_index]
pred_class = preds[..., class_index]
for index in range(len(batch_idx)):
sample_mask = mask_class[index, ]
sample_pred = pred_class[index, ]
sample_pred_post = post_process(sample_pred, threshold=tresholds[class_index], min_size=min_mask_sizes[class_index])
if (sample_mask.sum() == 0) & (sample_pred.sum() == 0):
dice_score = 1.
else:
dice_score = dice_coefficient(sample_pred, sample_mask)
if (sample_mask.sum() == 0) & (sample_pred_post.sum() == 0):
dice_score_post = 1.
else:
dice_score_post = dice_coefficient(sample_pred_post, sample_mask)
class_score.append(dice_score)
class_score_post.append(dice_score_post)
metrics_df.loc[metrics_df['Class'] == class_names[class_index], 'Dice'] += np.mean(class_score) * ratio
metrics_df.loc[metrics_df['Class'] == class_names[class_index], 'Dice Post'] += np.mean(class_score_post) * ratio
metrics_df = metrics_df.append({'Class':set_name, 'Dice':np.mean(metrics_df['Dice'].values), 'Dice Post':np.mean(metrics_df['Dice Post'].values)}, ignore_index=True).set_index('Class')
return metrics_df
def plot_metrics(history):
fig, axes = plt.subplots(4, 1, sharex='col', figsize=(22, 14))
axes = axes.flatten()
axes[0].plot(history['loss'], label='Train loss')
axes[0].plot(history['val_loss'], label='Validation loss')
axes[0].legend(loc='best')
axes[0].set_title('Loss')
axes[1].plot(history['iou_score'], label='Train IOU Score')
axes[1].plot(history['val_iou_score'], label='Validation IOU Score')
axes[1].legend(loc='best')
axes[1].set_title('IOU Score')
axes[2].plot(history['dice_coef'], label='Train Dice coefficient')
axes[2].plot(history['val_dice_coef'], label='Validation Dice coefficient')
axes[2].legend(loc='best')
axes[2].set_title('Dice coefficient')
axes[3].plot(history['score'], label='Train F-Score')
axes[3].plot(history['val_score'], label='Validation F-Score')
axes[3].legend(loc='best')
axes[3].set_title('F-Score')
plt.xlabel('Epochs')
sns.despine()
plt.show()
def pre_process_set(df, preprocess_fn):
n_cpu = mp.cpu_count()
df_n_cnt = df.shape[0]//n_cpu
pool = mp.Pool(n_cpu)
dfs = [df.iloc[df_n_cnt*i:df_n_cnt*(i+1)] for i in range(n_cpu)]
dfs[-1] = df.iloc[df_n_cnt*(n_cpu-1):]
res = pool.map(preprocess_fn, [x_df for x_df in dfs])
pool.close()
def cosine_decay_with_warmup(global_step,
learning_rate_base,
total_steps,
warmup_learning_rate=0.0,
warmup_steps=0,
hold_base_rate_steps=0):
"""
Cosine decay schedule with warm up period.
In this schedule, the learning rate grows linearly from warmup_learning_rate
to learning_rate_base for warmup_steps, then transitions to a cosine decay
schedule.
:param global_step {int}: global step.
:param learning_rate_base {float}: base learning rate.
:param total_steps {int}: total number of training steps.
:param warmup_learning_rate {float}: initial learning rate for warm up. (default: {0.0}).
:param warmup_steps {int}: number of warmup steps. (default: {0}).
:param hold_base_rate_steps {int}: Optional number of steps to hold base learning rate before decaying. (default: {0}).
:param global_step {int}: global step.
:Returns : a float representing learning rate.
:Raises ValueError: if warmup_learning_rate is larger than learning_rate_base, or if warmup_steps is larger than total_steps.
"""
if total_steps < warmup_steps:
raise ValueError('total_steps must be larger or equal to warmup_steps.')
learning_rate = 0.5 * learning_rate_base * (1 + np.cos(
np.pi *
(global_step - warmup_steps - hold_base_rate_steps
) / float(total_steps - warmup_steps - hold_base_rate_steps)))
if hold_base_rate_steps > 0:
learning_rate = np.where(global_step > warmup_steps + hold_base_rate_steps,
learning_rate, learning_rate_base)
if warmup_steps > 0:
if learning_rate_base < warmup_learning_rate:
raise ValueError('learning_rate_base must be larger or equal to warmup_learning_rate.')
slope = (learning_rate_base - warmup_learning_rate) / warmup_steps
warmup_rate = slope * global_step + warmup_learning_rate
learning_rate = np.where(global_step < warmup_steps, warmup_rate,
learning_rate)
return np.where(global_step > total_steps, 0.0, learning_rate)
class WarmUpCosineDecayScheduler(Callback):
"""Cosine decay with warmup learning rate scheduler"""
def __init__(self,
learning_rate_base,
total_steps,
global_step_init=0,
warmup_learning_rate=0.0,
warmup_steps=0,
hold_base_rate_steps=0,
verbose=0):
"""
Constructor for cosine decay with warmup learning rate scheduler.
:param learning_rate_base {float}: base learning rate.
:param total_steps {int}: total number of training steps.
:param global_step_init {int}: initial global step, e.g. from previous checkpoint.
:param warmup_learning_rate {float}: initial learning rate for warm up. (default: {0.0}).
:param warmup_steps {int}: number of warmup steps. (default: {0}).
:param hold_base_rate_steps {int}: Optional number of steps to hold base learning rate before decaying. (default: {0}).
:param verbose {int}: quiet, 1: update messages. (default: {0}).
"""
super(WarmUpCosineDecayScheduler, self).__init__()
self.learning_rate_base = learning_rate_base
self.total_steps = total_steps
self.global_step = global_step_init
self.warmup_learning_rate = warmup_learning_rate
self.warmup_steps = warmup_steps
self.hold_base_rate_steps = hold_base_rate_steps
self.verbose = verbose
self.learning_rates = []
def on_batch_end(self, batch, logs=None):
self.global_step = self.global_step + 1
lr = K.get_value(self.model.optimizer.lr)
self.learning_rates.append(lr)
def on_batch_begin(self, batch, logs=None):
lr = cosine_decay_with_warmup(global_step=self.global_step,
learning_rate_base=self.learning_rate_base,
total_steps=self.total_steps,
warmup_learning_rate=self.warmup_learning_rate,
warmup_steps=self.warmup_steps,
hold_base_rate_steps=self.hold_base_rate_steps)
K.set_value(self.model.optimizer.lr, lr)
if self.verbose > 0:
print('\nBatch %02d: setting learning rate to %s.' % (self.global_step + 1, lr))
```
## Pre-process data
```
train_base_path = '../input/understanding_cloud_organization/train_images/'
test_base_path = '../input/understanding_cloud_organization/test_images/'
train_images_dest_path = 'base_dir/train_images/'
validation_images_dest_path = 'base_dir/validation_images/'
test_images_dest_path = 'base_dir/test_images/'
# Making sure directories don't exist
if os.path.exists(train_images_dest_path):
shutil.rmtree(train_images_dest_path)
if os.path.exists(validation_images_dest_path):
shutil.rmtree(validation_images_dest_path)
if os.path.exists(test_images_dest_path):
shutil.rmtree(test_images_dest_path)
# Creating train, validation and test directories
os.makedirs(train_images_dest_path)
os.makedirs(validation_images_dest_path)
os.makedirs(test_images_dest_path)
# Pre-procecss train set
pre_process_set(X_train, preprocess_data)
# Pre-procecss validation set
pre_process_set(X_val, preprocess_data)
# Pre-procecss test set
pre_process_set(test, preprocess_data)
```
### Data generator
```
class DataGenerator(Sequence):
def __init__(self, dataframe, target_df=None, mode='fit', directory=train_images_dest_path,
batch_size=BATCH_SIZE, n_channels=CHANNELS, target_size=(HEIGHT, WIDTH),
n_classes=N_CLASSES, seed=seed, shuffle=True, preprocessing=None, augmentation=None):
self.batch_size = batch_size
self.dataframe = dataframe
self.mode = mode
self.directory = directory
self.target_df = target_df
self.target_size = target_size
self.n_channels = n_channels
self.n_classes = n_classes
self.shuffle = shuffle
self.augmentation = augmentation
self.preprocessing = preprocessing
self.seed = seed
self.mask_shape = (1400, 2100)
self.list_IDs = self.dataframe.index
if self.seed is not None:
np.random.seed(self.seed)
self.on_epoch_end()
def __len__(self):
return len(self.list_IDs) // self.batch_size
def __getitem__(self, index):
indexes = self.indexes[index*self.batch_size:(index+1)*self.batch_size]
list_IDs_batch = [self.list_IDs[k] for k in indexes]
X = self.__generate_X(list_IDs_batch)
if self.mode == 'fit':
Y = self.__generate_Y(list_IDs_batch)
if self.augmentation:
X, Y = self.__augment_batch(X, Y)
return X, Y
elif self.mode == 'predict':
return X
def on_epoch_end(self):
self.indexes = np.arange(len(self.list_IDs))
if self.shuffle == True:
np.random.shuffle(self.indexes)
def __generate_X(self, list_IDs_batch):
X = np.empty((self.batch_size, *self.target_size, self.n_channels))
for i, ID in enumerate(list_IDs_batch):
img_name = self.dataframe['image'].loc[ID]
img_path = self.directory + img_name
img = cv2.imread(img_path)
img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
if self.preprocessing:
img = self.preprocessing(img)
# img = img.astype(np.float32) / 255.
X[i,] = img
return X
def __generate_Y(self, list_IDs_batch):
Y = np.empty((self.batch_size, *self.target_size, self.n_classes), dtype=int)
for i, ID in enumerate(list_IDs_batch):
img_name = self.dataframe['image'].loc[ID]
image_df = self.target_df[self.target_df['image'] == img_name]
rles = image_df['EncodedPixels'].values
masks = build_masks(rles, input_shape=self.mask_shape, reshape=self.target_size)
Y[i, ] = masks
return Y
def __augment_batch(self, X_batch, Y_batch):
for i in range(X_batch.shape[0]):
X_batch[i, ], Y_batch[i, ] = self.__random_transform(X_batch[i, ], Y_batch[i, ])
return X_batch, Y_batch
def __random_transform(self, X, Y):
composed = self.augmentation(image=X, mask=Y)
X_aug = composed['image']
Y_aug = composed['mask']
return X_aug, Y_aug
train_generator = DataGenerator(
directory=train_images_dest_path,
dataframe=X_train,
target_df=train,
batch_size=BATCH_SIZE,
target_size=(HEIGHT, WIDTH),
n_channels=CHANNELS,
n_classes=N_CLASSES,
preprocessing=preprocessing,
augmentation=augmentation,
seed=seed)
valid_generator = DataGenerator(
directory=validation_images_dest_path,
dataframe=X_val,
target_df=train,
batch_size=BATCH_SIZE,
target_size=(HEIGHT, WIDTH),
n_channels=CHANNELS,
n_classes=N_CLASSES,
preprocessing=preprocessing,
seed=seed)
```
# Model
```
model = sm.Unet(backbone_name=BACKBONE,
encoder_weights='imagenet',
classes=N_CLASSES,
activation='sigmoid',
input_shape=(HEIGHT, WIDTH, CHANNELS))
checkpoint = ModelCheckpoint(model_path, monitor='val_loss', mode='min', save_best_only=True, save_weights_only=True)
es = EarlyStopping(monitor='val_loss', mode='min', patience=ES_PATIENCE, restore_best_weights=True, verbose=1)
rlrop = ReduceLROnPlateau(monitor='val_loss', mode='min', patience=RLROP_PATIENCE, factor=DECAY_DROP, min_lr=1e-6, verbose=1)
metric_list = [dice_coef, sm.metrics.iou_score, sm.metrics.f1_score]
callback_list = [checkpoint, es, rlrop]
optimizer = RAdam(learning_rate=LEARNING_RATE, warmup_proportion=0.1)
model.compile(optimizer=optimizer, loss=sm.losses.bce_dice_loss, metrics=metric_list)
model.summary()
STEP_SIZE_TRAIN = len(X_train)//BATCH_SIZE
STEP_SIZE_VALID = len(X_val)//BATCH_SIZE
history = model.fit_generator(generator=train_generator,
steps_per_epoch=STEP_SIZE_TRAIN,
validation_data=valid_generator,
validation_steps=STEP_SIZE_VALID,
callbacks=callback_list,
epochs=EPOCHS,
verbose=2).history
```
## Model loss graph
```
plot_metrics(history)
```
# Threshold and mask size tunning
```
class_names = ['Fish ', 'Flower', 'Gravel', 'Sugar ']
mask_grid = [0, 500, 1000, 5000, 7500, 10000, 15000]
threshold_grid = np.arange(.5, 1, .05)
metrics = []
for class_index in range(N_CLASSES):
for threshold in threshold_grid:
for mask_size in mask_grid:
metrics.append([class_index, threshold, mask_size, 0])
metrics_df = pd.DataFrame(metrics, columns=['Class', 'Threshold', 'Mask size', 'Dice'])
for i in range(0, X_val.shape[0], 500):
batch_idx = list(range(i, min(X_val.shape[0], i + 500)))
batch_set = X_val[batch_idx[0]: batch_idx[-1]+1]
ratio = len(batch_set) / len(X_val)
generator = DataGenerator(
directory=validation_images_dest_path,
dataframe=batch_set,
target_df=train,
batch_size=len(batch_set),
target_size=(HEIGHT, WIDTH),
n_channels=CHANNELS,
n_classes=N_CLASSES,
preprocessing=preprocessing,
seed=seed,
mode='fit',
shuffle=False)
x, y = generator.__getitem__(0)
preds = model.predict(x)
for class_index in range(N_CLASSES):
class_score = []
label_class = y[..., class_index]
pred_class = preds[..., class_index]
for threshold in threshold_grid:
for mask_size in mask_grid:
mask_score = []
for index in range(len(batch_idx)):
label_mask = label_class[index, ]
pred_mask = pred_class[index, ]
pred_mask = post_process(pred_mask, threshold=threshold, min_size=mask_size)
dice_score = dice_coefficient(pred_mask, label_mask)
if (pred_mask.sum() == 0) & (label_mask.sum() == 0):
dice_score = 1.
mask_score.append(dice_score)
metrics_df.loc[(metrics_df['Class'] == class_index) & (metrics_df['Threshold'] == threshold) &
(metrics_df['Mask size'] == mask_size), 'Dice'] += np.mean(mask_score) * ratio
metrics_df_0 = metrics_df[metrics_df['Class'] == 0]
metrics_df_1 = metrics_df[metrics_df['Class'] == 1]
metrics_df_2 = metrics_df[metrics_df['Class'] == 2]
metrics_df_3 = metrics_df[metrics_df['Class'] == 3]
optimal_values_0 = metrics_df_0.loc[metrics_df_0['Dice'].idxmax()].values
optimal_values_1 = metrics_df_1.loc[metrics_df_1['Dice'].idxmax()].values
optimal_values_2 = metrics_df_2.loc[metrics_df_2['Dice'].idxmax()].values
optimal_values_3 = metrics_df_3.loc[metrics_df_3['Dice'].idxmax()].values
best_tresholds = [optimal_values_0[1], optimal_values_1[1], optimal_values_2[1], optimal_values_3[1]]
best_masks = [optimal_values_0[2], optimal_values_1[2], optimal_values_2[2], optimal_values_3[2]]
best_dices = [optimal_values_0[3], optimal_values_1[3], optimal_values_2[3], optimal_values_3[3]]
for index, name in enumerate(class_names):
print('%s treshold=%.2f mask size=%d Dice=%.3f' % (name, best_tresholds[index], best_masks[index], best_dices[index]))
```
# Model evaluation
```
train_metrics = get_metrics(model, X_train, train_images_dest_path, best_tresholds, best_masks, 'Train')
display(train_metrics)
validation_metrics = get_metrics(model, X_val, validation_images_dest_path, best_tresholds, best_masks, 'Validation')
display(validation_metrics)
```
# Apply model to test set
```
test_df = []
for i in range(0, test.shape[0], 500):
batch_idx = list(range(i, min(test.shape[0], i + 500)))
batch_set = test[batch_idx[0]: batch_idx[-1]+1]
test_generator = DataGenerator(
directory=test_images_dest_path,
dataframe=batch_set,
target_df=submission,
batch_size=1,
target_size=(HEIGHT, WIDTH),
n_channels=CHANNELS,
n_classes=N_CLASSES,
preprocessing=preprocessing,
seed=seed,
mode='predict',
shuffle=False)
preds = model.predict_generator(test_generator)
for index, b in enumerate(batch_idx):
filename = test['image'].iloc[b]
image_df = submission[submission['image'] == filename].copy()
pred_masks = preds[index, ].round().astype(int)
pred_rles = build_rles(pred_masks, reshape=(350, 525))
image_df['EncodedPixels'] = pred_rles
### Post procecssing
pred_masks_post = preds[index, ].astype('float32')
for class_index in range(N_CLASSES):
pred_mask = pred_masks_post[...,class_index]
pred_mask = post_process(pred_mask, threshold=best_tresholds[class_index], min_size=best_masks[class_index])
pred_masks_post[...,class_index] = pred_mask
pred_rles_post = build_rles(pred_masks_post, reshape=(350, 525))
image_df['EncodedPixels_post'] = pred_rles_post
###
test_df.append(image_df)
sub_df = pd.concat(test_df)
```
### Regular submission
```
submission_df = sub_df[['Image_Label' ,'EncodedPixels']]
submission_df.to_csv('submission.csv', index=False)
display(submission_df.head())
```
### Submission with post processing
```
submission_df_post = sub_df[['Image_Label' ,'EncodedPixels_post']]
submission_df_post.columns = ['Image_Label' ,'EncodedPixels']
submission_df_post.to_csv('submission_post.csv', index=False)
display(submission_df_post.head())
# Cleaning created directories
if os.path.exists(train_images_dest_path):
shutil.rmtree(train_images_dest_path)
if os.path.exists(validation_images_dest_path):
shutil.rmtree(validation_images_dest_path)
if os.path.exists(test_images_dest_path):
shutil.rmtree(test_images_dest_path)
```
| github_jupyter |
<img src="http://hilpisch.com/tpq_logo.png" alt="The Python Quants" width="35%" align="right" border="0"><br>
# Python for Finance (2nd ed.)
**Mastering Data-Driven Finance**
© Dr. Yves J. Hilpisch | The Python Quants GmbH
<img src="http://hilpisch.com/images/py4fi_2nd_shadow.png" width="300px" align="left">
# Financial Time Series
```
import numpy as np
import pandas as pd
from pylab import mpl, plt
plt.style.use('seaborn')
mpl.rcParams['font.family'] = 'serif'
%matplotlib inline
```
## Financial Data
### Data Import
```
filename = '../../source/tr_eikon_eod_data.csv'
f = open(filename, 'r')
f.readlines()[:5]
data = pd.read_csv(filename,
index_col=0,
parse_dates=True)
data.info()
data.head()
data.tail()
data.plot(figsize=(10, 12), subplots=True);
# plt.savefig('../../images/ch08/fts_01.png');
instruments = ['Apple Stock', 'Microsoft Stock',
'Intel Stock', 'Amazon Stock', 'Goldman Sachs Stock',
'SPDR S&P 500 ETF Trust', 'S&P 500 Index',
'VIX Volatility Index', 'EUR/USD Exchange Rate',
'Gold Price', 'VanEck Vectors Gold Miners ETF',
'SPDR Gold Trust']
for ric, name in zip(data.columns, instruments):
print('{:8s} | {}'.format(ric, name))
```
### Summary Statistics
```
data.info()
data.describe().round(2)
data.mean()
data.aggregate([min,
np.mean,
np.std,
np.median,
max]
).round(2)
```
### Changes Over Time
```
data.diff().head()
data.diff().mean()
data.pct_change().round(3).head()
data.pct_change().mean().plot(kind='bar', figsize=(10, 6));
# plt.savefig('../../images/ch08/fts_02.png');
rets = np.log(data / data.shift(1))
rets.head().round(3)
rets.cumsum().apply(np.exp).plot(figsize=(10, 6));
# plt.savefig('../../images/ch08/fts_03.png');
```
### Resampling
```
data.resample('1w', label='right').last().head()
data.resample('1m', label='right').last().head()
rets.cumsum().apply(np.exp). resample('1m', label='right').last(
).plot(figsize=(10, 6));
# plt.savefig('../../images/ch08/fts_04.png');
```
## Rolling Statistics
```
sym = 'AAPL.O'
data = pd.DataFrame(data[sym]).dropna()
data.tail()
```
### An Overview
```
window = 20
data['min'] = data[sym].rolling(window=window).min()
data['mean'] = data[sym].rolling(window=window).mean()
data['std'] = data[sym].rolling(window=window).std()
data['median'] = data[sym].rolling(window=window).median()
data['max'] = data[sym].rolling(window=window).max()
data['ewma'] = data[sym].ewm(halflife=0.5, min_periods=window).mean()
data.dropna().head()
ax = data[['min', 'mean', 'max']].iloc[-200:].plot(
figsize=(10, 6), style=['g--', 'r--', 'g--'], lw=0.8)
data[sym].iloc[-200:].plot(ax=ax, lw=2.0);
# plt.savefig('../../images/ch08/fts_05.png');
```
### A Technical Analysis Example
```
data['SMA1'] = data[sym].rolling(window=42).mean()
data['SMA2'] = data[sym].rolling(window=252).mean()
data[[sym, 'SMA1', 'SMA2']].tail()
data[[sym, 'SMA1', 'SMA2']].plot(figsize=(10, 6));
# plt.savefig('../../images/ch08/fts_06.png');
data.dropna(inplace=True)
data['positions'] = np.where(data['SMA1'] > data['SMA2'],
1,
-1)
ax = data[[sym, 'SMA1', 'SMA2', 'positions']].plot(figsize=(10, 6),
secondary_y='positions')
ax.get_legend().set_bbox_to_anchor((0.25, 0.85));
# plt.savefig('../../images/ch08/fts_07.png');
```
## Regression Analysis
### The Data
```
# EOD data from Thomson Reuters Eikon Data API
raw = pd.read_csv('../../source/tr_eikon_eod_data.csv',
index_col=0, parse_dates=True)
data = raw[['.SPX', '.VIX']].dropna()
data.tail()
data.plot(subplots=True, figsize=(10, 6));
# plt.savefig('../../images/ch08/fts_08.png');
data.loc[:'2012-12-31'].plot(secondary_y='.VIX', figsize=(10, 6));
# plt.savefig('../../images/ch08/fts_09.png');
```
### Log Returns
```
rets = np.log(data / data.shift(1))
rets.head()
rets.dropna(inplace=True)
rets.plot(subplots=True, figsize=(10, 6));
# plt.savefig('../../images/ch08/fts_10.png');
pd.plotting.scatter_matrix(rets,
alpha=0.2,
diagonal='hist',
hist_kwds={'bins': 35},
figsize=(10, 6));
# plt.savefig('../../images/ch08/fts_11.png');
```
### OLS Regression
```
reg = np.polyfit(rets['.SPX'], rets['.VIX'], deg=1)
ax = rets.plot(kind='scatter', x='.SPX', y='.VIX', figsize=(10, 6))
ax.plot(rets['.SPX'], np.polyval(reg, rets['.SPX']), 'r', lw=2);
# plt.savefig('../../images/ch08/fts_12.png');
```
### Correlation
```
rets.corr()
ax = rets['.SPX'].rolling(window=252).corr(
rets['.VIX']).plot(figsize=(10, 6))
ax.axhline(rets.corr().iloc[0, 1], c='r');
# plt.savefig('../../images/ch08/fts_13.png');
```
## High Frequency Data
```
# from fxcmpy import fxcmpy_tick_data_reader as tdr
# data = tdr('EURUSD', start='2018-6-25', end='2018-06-30')
# data.get_data(start='2018-6-29',
# end='2018-06-30').to_csv('../../source/fxcm_eur_usd_tick_data.csv')
%%time
# data from FXCM Forex Capital Markets Ltd.
tick = pd.read_csv('../../source/fxcm_eur_usd_tick_data.csv',
index_col=0, parse_dates=True)
tick.info()
tick['Mid'] = tick.mean(axis=1)
tick['Mid'].plot(figsize=(10, 6));
# plt.savefig('../../images/ch08/fts_14.png');
tick_resam = tick.resample(rule='5min', label='right').last()
tick_resam.head()
tick_resam['Mid'].plot(figsize=(10, 6));
# plt.savefig('../../images/ch08/fts_15.png');
```
<img src="http://hilpisch.com/tpq_logo.png" alt="The Python Quants" width="35%" align="right" border="0"><br>
<a href="http://tpq.io" target="_blank">http://tpq.io</a> | <a href="http://twitter.com/dyjh" target="_blank">@dyjh</a> | <a href="mailto:training@tpq.io">training@tpq.io</a>
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# TensorFlow 2 início rápido para especialistas
<table class="tfo-notebook-buttons" align="left">
<td><a target="_blank" href="https://www.tensorflow.org/tutorials/quickstart/advanced"><img src="https://www.tensorflow.org/images/tf_logo_32px.png">Ver em TensorFlow.org</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/pt-br/tutorials/quickstart/advanced.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png">Executar no Google Colab</a></td>
<td><a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/pt-br/tutorials/quickstart/advanced.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png">Ver código fontes no GitHub</a></td>
<td><a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/pt-br/tutorials/quickstart/advanced.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png">Baixar notebook</a></td>
</table>
Este é um arquivo de notebook [Google Colaboratory] (https://colab.research.google.com/notebooks/welcome.ipynb). Os programas Python são executados diretamente no navegador - uma ótima maneira de aprender e usar o TensorFlow. Para seguir este tutorial, execute o bloco de anotações no Google Colab clicando no botão na parte superior desta página.
1. No Colab, conecte-se a uma instância do Python: No canto superior direito da barra de menus, selecione * CONNECT*.
2. Execute todas as células de código do notebook: Selecione * Runtime * > * Run all *.
Faça o download e instale o pacote TensorFlow 2:
Note: Upgrade `pip` to install the TensorFlow 2 package. See the [install guide](https://www.tensorflow.org/install) for details.
Importe o TensorFlow dentro de seu programa:
```
from __future__ import absolute_import, division, print_function, unicode_literals
import tensorflow as tf
from tensorflow.keras.layers import Dense, Flatten, Conv2D
from tensorflow.keras import Model
```
Carregue e prepare o [conjunto de dados MNIST] (http://yann.lecun.com/exdb/mnist/).
```
mnist = tf.keras.datasets.mnist
(x_train, y_train), (x_test, y_test) = mnist.load_data()
x_train, x_test = x_train / 255.0, x_test / 255.0
# Adicione uma dimensão de canais
x_train = x_train[..., tf.newaxis]
x_test = x_test[..., tf.newaxis]
```
Use `tf.data` para agrupar e embaralhar o conjunto de dados:
```
train_ds = tf.data.Dataset.from_tensor_slices(
(x_train, y_train)).shuffle(10000).batch(32)
test_ds = tf.data.Dataset.from_tensor_slices((x_test, y_test)).batch(32)
```
Crie o modelo `tf.keras` usando a Keras [API de subclasse de modelo] (https://www.tensorflow.org/guide/keras#model_subclassing):
```
class MyModel(Model):
def __init__(self):
super(MyModel, self).__init__()
self.conv1 = Conv2D(32, 3, activation='relu')
self.flatten = Flatten()
self.d1 = Dense(128, activation='relu')
self.d2 = Dense(10, activation='softmax')
def call(self, x):
x = self.conv1(x)
x = self.flatten(x)
x = self.d1(x)
return self.d2(x)
# Crie uma instância do modelo
model = MyModel()
```
Escolha uma função otimizadora e de perda para treinamento:
```
loss_object = tf.keras.losses.SparseCategoricalCrossentropy()
optimizer = tf.keras.optimizers.Adam()
```
Selecione métricas para medir a perda e a precisão do modelo. Essas métricas acumulam os valores ao longo das épocas e, em seguida, imprimem o resultado geral.
```
train_loss = tf.keras.metrics.Mean(name='train_loss')
train_accuracy = tf.keras.metrics.SparseCategoricalAccuracy(name='train_accuracy')
test_loss = tf.keras.metrics.Mean(name='test_loss')
test_accuracy = tf.keras.metrics.SparseCategoricalAccuracy(name='test_accuracy')
```
Use `tf.GradientTape` para treinar o modelo:
```
@tf.function
def train_step(images, labels):
with tf.GradientTape() as tape:
# training=True é necessário apenas se houver camadas com diferentes
# comportamentos durante o treinamento versus inferência (por exemplo, Dropout).
predictions = model(images, training=True)
loss = loss_object(labels, predictions)
gradients = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(gradients, model.trainable_variables))
train_loss(loss)
train_accuracy(labels, predictions)
```
Teste o modelo:
```
@tf.function
def test_step(images, labels):
# training=True é necessário apenas se houver camadas com diferentes
# comportamentos durante o treinamento versus inferência (por exemplo, Dropout).
predictions = model(images, training=False)
t_loss = loss_object(labels, predictions)
test_loss(t_loss)
test_accuracy(labels, predictions)
EPOCHS = 5
for epoch in range(EPOCHS):
# Reiniciar as métricas no início da próxima época
train_loss.reset_states()
train_accuracy.reset_states()
test_loss.reset_states()
test_accuracy.reset_states()
for images, labels in train_ds:
train_step(images, labels)
for test_images, test_labels in test_ds:
test_step(test_images, test_labels)
template = 'Epoch {}, Loss: {}, Accuracy: {}, Test Loss: {}, Test Accuracy: {}'
print(template.format(epoch+1,
train_loss.result(),
train_accuracy.result()*100,
test_loss.result(),
test_accuracy.result()*100))
```
O classificador de imagem agora é treinado para ~98% de acurácia neste conjunto de dados. Para saber mais, leia os [tutoriais do TensorFlow] (https://www.tensorflow.org/tutorials).
| github_jupyter |
# Keras_mnist_LeNet-5
**此项目初步实现LeNet-5**
- 达到0.9899的准确率
```
%matplotlib inline
import os
import PIL
import pickle
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import tensorflow as tf
import keras
from IPython import display
from functools import partial
from sklearn.preprocessing import normalize
from keras import backend
from keras.utils import np_utils, plot_model
from keras.callbacks import TensorBoard, ModelCheckpoint
from keras.callbacks import LearningRateScheduler, ReduceLROnPlateau
from keras.preprocessing.image import ImageDataGenerator
from keras.models import load_model
from keras.models import Sequential, Model
from keras.layers import Dense, Conv2D, MaxPool2D, Input, AveragePooling2D
from keras.layers import Activation, Dropout, Flatten, BatchNormalization
import warnings
warnings.filterwarnings('ignore')
np.random.seed(42)
file_path = r"I:\Dataset\mnist\all_mnist_data.csv"
mnist_data = pd.read_csv(file_path)
idx = np.random.permutation(len(mnist_data))
train_data = mnist_data.iloc[idx[: 60000]]
test_data = mnist_data.iloc[idx[60000: ]]
X_train = np.array(train_data.drop('0', axis=1)).reshape(-1, 28, 28, 1).astype("float32")
X_test = np.array(test_data.drop('0', axis=1)).reshape(-1, 28, 28, 1).astype("float32")
y_train = np.array(train_data['0'])
y_test = np.array(test_data['0'])
y_train = np_utils.to_categorical(y_train)
y_test = np_utils.to_categorical(y_test)
x_train = X_train[10000:]
t_train = y_train[10000:]
x_val = X_train[:10000]
t_val = y_train[:10000]
print("\nimgs of trainset : ", x_train.shape)
print("labels of trainset : ", t_train.shape)
print("imgs of valset : ", x_val.shape)
print("labels of valset : ", t_val.shape)
print("imgs of testset : ", X_test.shape)
print("labels of testset : ", y_test.shape)
def myCNN():
model = Sequential()
model.add(Conv2D(filters=6,
kernel_size=(5, 5),
input_shape=(28, 28, 1),
padding="same",
activation="relu",
name="conv2d_1"))
model.add(MaxPool2D(pool_size=(2, 2), name="maxpool2d_1"))
model.add(Conv2D(filters=16,
kernel_size=(5, 5),
input_shape=(14, 14, 1),
padding="valid",
activation="relu",
name="conv2d_2"))
model.add(MaxPool2D(pool_size=(2, 2), name="maxpool2d_2"))
model.add(Flatten())
model.add(Dense(units=120, activation="relu"))
model.add(Dropout(0.5))
model.add(Dense(units=84, activation="relu"))
model.add(Dropout(0.5))
model.add(Dense(units=10, activation="softmax", name="output"))
return model
model = myCNN()
print(model.summary())
config = tf.ConfigProto()
config.gpu_options.per_process_gpu_memory_fraction = 0.2
sess = tf.Session(config=config)
backend.set_session(sess)
"""训练模型并保存模型及训练历史
保存模型单独创建一个子文件夹modeldir, 保存训练历史则为单个文件hisfile"""
models_name = "Keras_mnist_LeNet-5_v1" # 模型名称的公共前缀
factor_list = [""] # 此次调参的变量列表
model_list = [] # 模型名称列表
for i in range(len(factor_list)):
modelname = models_name + factor_list[i] + ".h5"
model_list.append(modelname)
# 创建模型保存子目录modeldir
if not os.path.isdir("saved_models"):
os.mkdir("saved_models")
modeldir = r"saved_models"
# 创建训练历史保存目录
if not os.path.isdir("train_history"):
os.mkdir("train_history")
# 设置训练历史文件路径
hisfile = r"train_history\Keras_mnist_LeNet-5_v1.train_history"
# 每个模型及其对应的训练历史作为键值对{modelname: train_history}
# train_history为字典,含四个key,代表train和val的loss和acc
model_train_history = dict()
# 开始训练
epochs=100
batch_size = 32
steps_per_epoch=1250
for i in range(len(model_list)):
model = myCNN()
modelname = model_list[i]
modelpath = os.path.join(modeldir, modelname)
train_his = np.array([]).reshape(-1, 2)
val_his = np.array([]).reshape(-1, 2)
datagen = ImageDataGenerator()
datagen.fit(x_train)
model.compile(loss="categorical_crossentropy",
optimizer=keras.optimizers.Adam(),
metrics=["accuracy"])
print("\ntraining model : ", modelname)
ck_epoch, max_val_acc = 0, 0.0
for epoch in range(epochs+1):
i = 0
tr_his = []
for X, y in datagen.flow(x_train, t_train, batch_size=batch_size):
his = model.train_on_batch(X, y)
tr_his.append(his)
i += 1
if i >= steps_per_epoch: break
tr = np.mean(tr_his, axis=0)
val = model.evaluate(x_val, t_val, verbose=0)
train_his = np.vstack((train_his, tr))
val_his = np.vstack((val_his, val))
if epoch<10 or epoch %5 == 0:
print("%4d epoch: train acc: %8f loss: %8f val acc: %8f loss: %8f"%(epoch, tr[1], tr[0], val[1], val[0]))
# 设置保存模型
if val[1] > max_val_acc:
model.save(modelpath)
print("val acc improved from %6f to %6f"%(max_val_acc, val[1]))
max_val_acc = val[1]
ck_epoch = epoch
model_train_history[modelname] = {"acc": train_his[:, 1], "val_acc": val_his[:, 1],
"loss": train_his[:, 0], "val_loss": val_his[:, 0]}
"""保存训练历史"""
fo = open(hisfile, 'wb')
pickle.dump(model_train_history, fo)
fo.close()
```
## 可视化训练过程
```
def show_train_history(saved_history, his_img_file):
modelnames = sorted(list(saved_history.keys()))
train = ["acc", "loss"]
val = ["val_acc", "val_loss"]
"""作loss和acc两个图"""
fig, ax = plt.subplots(1, 2, figsize=(16, 5))
ax = ax.flatten()
color_add = 0.9/len(saved_history)
for i in range(2):
c = 0.05
for j in range(len(saved_history)):
modelname = modelnames[j]
train_history = saved_history[modelname]
ax[i].plot(train_history[train[i]],
color=(0, 1-c, 0),
linestyle="-",
label="train_"+modelname[21:-3])
ax[i].plot(train_history[val[i]],
color=(c, 0, 1-c),
linestyle="-",
label="val_"+modelname[21:-3])
c += color_add
ax[i].set_title('Train History')
ax[i].set_ylabel(train[i])
ax[i].set_xlabel('Epoch')
ax[0].legend(loc="lower right")
ax[1].legend(loc="upper right")
ax[0].set_ylim(0.9, 1.0)
ax[1].set_ylim(0, 0.2)
plt.suptitle("LeNet-5_v1")
print("saved img: ", his_img_file)
plt.savefig(his_img_file)
plt.show()
"""载入训练历史并可视化, 并且保存图片"""
if not os.path.isdir("his_img"):
os.mkdir("his_img")
his_img_file = r"his_img\LeNet-5_v1.png"
fo2 = open(hisfile, "rb")
saved_history1 = pickle.load(fo2)
show_train_history(saved_history1, his_img_file)
```
## 在测试集上测试
```
smodel = load_model(modelpath)
print("test model : ", os.path.basename(modelpath))
loss, acc = smodel.evaluate(X_test, y_test)
print("test :acc: %.4f"%(acc))
```
| github_jupyter |
# RAMP on predicting cyclist traffic in Paris
Authors: *Roman Yurchak (Symerio)*; also partially inspired by the air_passengers starting kit.
## Introduction
The dataset was collected with cyclist counters installed by Paris city council in multiple locations. It contains hourly information about cyclist traffic, as well as the following features,
- counter name
- counter site name
- date
- counter installation date
- latitude and longitude
Available features are quite scarce. However, **we can also use any external data that can help us to predict the target variable.**
```
from pathlib import Path
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import sklearn
```
# Loading the data with pandas
First, download the data files,
- [train.parquet](https://github.com/rth/bike_counters/releases/download/v0.1.0/train.parquet)
- [test.parquet](https://github.com/rth/bike_counters/releases/download/v0.1.0/test.parquet)
and put them to into the data folder.
Data is stored in [Parquet format](https://parquet.apache.org/), an efficient columnar data format. We can load the train set with pandas,
```
data = pd.read_parquet(Path("data") / "train.parquet")
data.head()
```
We can check general information about different columns,
```
data.info()
```
and in particular the number of unique entries in each column,
```
data.nunique(axis=0)
```
We have a 30 counting sites where sometimes multiple counters are installed per location. Let's look at the most frequented stations,
```
data.groupby(["site_name", "counter_name"])["bike_count"].sum().sort_values(
ascending=False
).head(10).to_frame()
```
# Visualizing the data
Let's visualize the data, starting from the spatial distribution of counters on the map
```
import folium
m = folium.Map(location=data[["latitude", "longitude"]].mean(axis=0), zoom_start=13)
for _, row in (
data[["counter_name", "latitude", "longitude"]]
.drop_duplicates("counter_name")
.iterrows()
):
folium.Marker(
row[["latitude", "longitude"]].values.tolist(), popup=row["counter_name"]
).add_to(m)
m
```
Note that in this RAMP problem we consider only the 30 most frequented counting sites, to limit data size.
Next we will look into the temporal distribution of the most frequented bike counter. If we plot it directly we will not see much because there are half a million data points,
```
mask = data["counter_name"] == "Totem 73 boulevard de Sébastopol S-N"
data[mask].plot(x="date", y="bike_count")
```
Instead we aggregate the data, for instance, by week to have a clearer overall picture,
```
mask = data["counter_name"] == "Totem 73 boulevard de Sébastopol S-N"
data[mask].groupby(pd.Grouper(freq="1w", key="date"))[["bike_count"]].sum().plot()
```
While at the same time, we can zoom on a week in particular for a more short-term visualization,
```
fig, ax = plt.subplots(figsize=(10, 4))
mask = (
(data["counter_name"] == "Totem 73 boulevard de Sébastopol S-N")
& (data["date"] > pd.to_datetime("2021/03/01"))
& (data["date"] < pd.to_datetime("2021/03/08"))
)
data[mask].plot(x="date", y="bike_count", ax=ax)
```
The hourly pattern has a clear variation between work days and weekends (7 and 8 March 2021).
If we look at the distribution of the target variable it skewed and non normal,
```
import seaborn as sns
ax = sns.histplot(data, x="bike_count", kde=True, bins=50)
```
Least square loss would not be appropriate to model it since it is designed for normal error distributions. One way to precede would be to transform the variable with a logarithmic transformation,
```py
data['log_bike_count'] = np.log(1 + data['bike_count'])
```
```
ax = sns.histplot(data, x="log_bike_count", kde=True, bins=50)
```
which has a more pronounced central mode, but is still non symmetric. In the following, **we use `log_bike_count` as the target variable** as otherwise `bike_count` ranges over 3 orders of magnitude and least square loss would be dominated by the few large values.
## Feature extraction
To account for the temporal aspects of the data, we cannot input the `date` field directly into the model. Instead we extract the features on different time-scales from the `date` field,
```
def _encode_dates(X):
X = X.copy() # modify a copy of X
# Encode the date information from the DateOfDeparture columns
X.loc[:, "year"] = X["date"].dt.year
X.loc[:, "month"] = X["date"].dt.month
X.loc[:, "day"] = X["date"].dt.day
X.loc[:, "weekday"] = X["date"].dt.weekday
X.loc[:, "hour"] = X["date"].dt.hour
# Finally we can drop the original columns from the dataframe
return X.drop(columns=["date"])
data["date"].head()
_encode_dates(data[["date"]].head())
```
To use this function with scikit-learn estimators we wrap it with [FunctionTransformer](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.FunctionTransformer.html),
```
from sklearn.preprocessing import FunctionTransformer
date_encoder = FunctionTransformer(_encode_dates, validate=False)
date_encoder.fit_transform(data[["date"]]).head()
```
Since it is unlikely that, for instance, that `hour` is linearly correlated with the target variable, we would need to additionally encode categorical features for linear models. This is classically done with [OneHotEncoder](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.OneHotEncoder.html), though other encoding strategies exist.
```
from sklearn.preprocessing import OneHotEncoder
enc = OneHotEncoder(sparse=False)
enc.fit_transform(_encode_dates(data[["date"]])[["hour"]].head())
```
## Linear model
Let's now construct our first linear model with [Ridge](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.Ridge.html). We use a few helper functions defined in `problem.py` of the starting kit to load the public train and test data:
```
import problem
X_train, y_train = problem.get_train_data()
X_test, y_test = problem.get_test_data()
X_train.head(2)
```
and
```
y_train
```
Where `y` contains the `log_bike_count` variable.
The test set is in the future as compared to the train set,
```
print(
f'Train: n_samples={X_train.shape[0]}, {X_train["date"].min()} to {X_train["date"].max()}'
)
print(
f'Test: n_samples={X_test.shape[0]}, {X_test["date"].min()} to {X_test["date"].max()}'
)
_encode_dates(X_train[["date"]]).columns.tolist()
from sklearn.compose import ColumnTransformer
from sklearn.linear_model import Ridge
from sklearn.pipeline import make_pipeline
date_encoder = FunctionTransformer(_encode_dates)
date_cols = _encode_dates(X_train[["date"]]).columns.tolist()
categorical_encoder = OneHotEncoder(handle_unknown="ignore")
categorical_cols = ["counter_name", "site_name"]
preprocessor = ColumnTransformer(
[
("date", OneHotEncoder(handle_unknown="ignore"), date_cols),
("cat", categorical_encoder, categorical_cols),
]
)
regressor = Ridge()
pipe = make_pipeline(date_encoder, preprocessor, regressor)
pipe.fit(X_train, y_train)
```
We then evaluate this model with the RMSE metric,
```
from sklearn.metrics import mean_squared_error
print(
f"Train set, RMSE={mean_squared_error(y_train, pipe.predict(X_train), squared=False):.2f}"
)
print(
f"Test set, RMSE={mean_squared_error(y_test, pipe.predict(X_test), squared=False):.2f}"
)
```
The model doesn't have enough capacity to generalize on the train set, since we have lots of data with relatively few parameters. However it happened to work somewhat better on the test set. We can compare these results with the baseline predicting the mean value,
```
print("Baseline mean prediction.")
print(
f"Train set, RMSE={mean_squared_error(y_train, np.full(y_train.shape, y_train.mean()), squared=False):.2f}"
)
print(
f"Test set, RMSE={mean_squared_error(y_test, np.full(y_test.shape, y_test.mean()), squared=False):.2f}"
)
```
which illustrates that we are performing better than the baseline.
Let's visualize the predictions for one of the stations,
```
mask = (
(X_test["counter_name"] == "Totem 73 boulevard de Sébastopol S-N")
& (X_test["date"] > pd.to_datetime("2021/09/01"))
& (X_test["date"] < pd.to_datetime("2021/09/08"))
)
df_viz = X_test.loc[mask].copy()
df_viz["bike_count"] = np.exp(y_test[mask.values]) - 1
df_viz["bike_count (predicted)"] = np.exp(pipe.predict(X_test[mask])) - 1
fig, ax = plt.subplots(figsize=(12, 4))
df_viz.plot(x="date", y="bike_count", ax=ax)
df_viz.plot(x="date", y="bike_count (predicted)", ax=ax, ls="--")
ax.set_title("Predictions with Ridge")
ax.set_ylabel("bike_count")
```
So we start to see the daily trend, and some of the week day differences are accounted for, however we still miss the details and the spikes in the evening are under-estimated.
A useful way to visualize the error is to plot `y_pred` as a function of `y_true`,
```
fig, ax = plt.subplots()
df_viz = pd.DataFrame({"y_true": y_test, "y_pred": pipe.predict(X_test)}).sample(
10000, random_state=0
)
df_viz.plot.scatter(x="y_true", y="y_pred", s=8, alpha=0.1, ax=ax)
```
It is recommended to use cross-validation for hyper-parameter tuning with [GridSearchCV](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.GridSearchCV.html) or more reliable model evaluation with [cross_val_score](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.cross_val_score.html#sklearn.model_selection.cross_val_score). In this case, because we want the test data to always be in the future as compared to the train data, we can use [TimeSeriesSplit](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.TimeSeriesSplit.html),
<img src="https://i.stack.imgur.com/Q37Bn.png" />
The disadvantage, is that we can either have the training set size be different for each fold which is not ideal for hyper-parameter tuning (current figure), or have constant sized small training set which is also not ideal given the data periodicity. This explains that generally we will have worse cross-validation scores than test scores,
```
from sklearn.model_selection import TimeSeriesSplit, cross_val_score
cv = TimeSeriesSplit(n_splits=6)
# When using a scorer in scikit-learn it always needs to be better when smaller, hence the minus sign.
scores = cross_val_score(
pipe, X_train, y_train, cv=cv, scoring="neg_root_mean_squared_error"
)
print("RMSE: ", scores)
print(f"RMSE (all folds): {-scores.mean():.3} ± {(-scores).std():.3}")
```
## Tree based model
For tabular data tree based models often perform well, since they are able to learn non linear relationships between features, which would take effort to manually create for the linear model. Here will use Histogram-based Gradient Boosting Regression ([HistGradientBoostingRegressor](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.HistGradientBoostingRegressor.html)) which often will produce good results on arbitrary tabular data, and is fairly fast.
```
_encode_dates(X_train[["date"]]).columns.tolist()
from sklearn.ensemble import HistGradientBoostingRegressor
from sklearn.preprocessing import OrdinalEncoder
date_encoder = FunctionTransformer(_encode_dates)
date_cols = _encode_dates(X_train[["date"]]).columns.tolist()
categorical_encoder = OrdinalEncoder(
handle_unknown="use_encoded_value", unknown_value=-1
)
categorical_cols = ["counter_name", "site_name"]
preprocessor = ColumnTransformer(
[
("date", "passthrough", date_cols),
(
"cat",
OrdinalEncoder(handle_unknown="use_encoded_value", unknown_value=-1),
categorical_cols,
),
]
)
regressor = HistGradientBoostingRegressor(random_state=0)
pipe = make_pipeline(date_encoder, preprocessor, regressor)
scores = cross_val_score(
pipe,
X_train,
y_train,
cv=cv,
scoring="neg_root_mean_squared_error",
error_score=np.nan,
)
print(f"RMSE: {-scores.mean():.3} ± {(-scores).std():.3}")
```
## Using external data
In this starting kit you are provided with weather data from Meteo France, which could correlate with cyclist traffic. It is not very accurate however, as the station is in Orly (15km from Paris) and only provides 3 hour updates.
To load the external data,
```
df_ext = pd.read_csv(Path("submissions") / "starting_kit" / "external_data.csv")
df_ext.head()
```
You can find the detailed documentation for each feature [in this PDF](https://donneespubliques.meteofrance.fr/client/document/doc_parametres_synop_168.pdf) (in French). Here the only feature we consider is the temperature. We will use [pandas.merge_asof](https://pandas.pydata.org/pandas-docs/dev/reference/api/pandas.merge_asof.html) to merge on date, using the closest available date (since the sampling between the external data and the counter data).
```
# In this notebook we define the __file__ variable to be in the same conditions as when running the
# RAMP submission
__file__ = Path("submissions") / "starting_kit" / "estimator.py"
def _merge_external_data(X):
file_path = Path(__file__).parent / "external_data.csv"
df_ext = pd.read_csv(file_path, parse_dates=["date"])
X = X.copy()
# When using merge_asof left frame need to be sorted
X["orig_index"] = np.arange(X.shape[0])
X = pd.merge_asof(
X.sort_values("date"), df_ext[["date", "t"]].sort_values("date"), on="date"
)
# Sort back to the original order
X = X.sort_values("orig_index")
del X["orig_index"]
return X
X_train_merge = _merge_external_data(X_train)
X_train_merge.head()
```
Similarly we can wrap this function into a FunctionTranformer to use it in a scikit-learn pipeline. Now let's see whether this additional feature improves the model,
```
from sklearn.ensemble import HistGradientBoostingRegressor
from sklearn.preprocessing import OrdinalEncoder
date_encoder = FunctionTransformer(_encode_dates)
date_cols = _encode_dates(X_train[["date"]]).columns.tolist()
categorical_encoder = OrdinalEncoder(
handle_unknown="use_encoded_value", unknown_value=-1
)
categorical_cols = ["counter_name", "site_name"]
numeric_cols = ["t"]
preprocessor = ColumnTransformer(
[
("date", "passthrough", date_cols),
(
"cat",
OrdinalEncoder(handle_unknown="use_encoded_value", unknown_value=-1),
categorical_cols,
),
("numeric", "passthrough", numeric_cols),
]
)
regressor = HistGradientBoostingRegressor(random_state=1)
pipe = make_pipeline(
FunctionTransformer(_merge_external_data, validate=False),
date_encoder,
preprocessor,
regressor,
)
scores = cross_val_score(
pipe,
X_train,
y_train,
cv=cv,
scoring="neg_root_mean_squared_error",
error_score=np.nan,
)
print(f"RMSE: {-scores.mean():.3}± {(-scores).std():.3}")
```
## Feature importance
We can check the feature importances using the function [sklearn.inspection.permutation_importances](https://scikit-learn.org/stable/modules/generated/sklearn.inspection.permutation_importance.html). Since the first step of our pipeline adds the temperature and extract the date components, we want to apply this transformation those steps, to check the importance of all features. Indeed, we can perform sklearn.inspection.permutation_importances at any stage of the pipeline.
```
merger = pipe[:2]
X_train_augmented = merger.transform(X_train)
X_test_augmented = merger.transform(X_test)
predictor = pipe[2:]
_ = predictor.fit(X_train_augmented, y_train)
from sklearn.inspection import permutation_importance
feature_importances = permutation_importance(
predictor, X_train_augmented[:20000], y_train[:20000], n_repeats=10, random_state=0
)
sorted_idx = feature_importances.importances_mean.argsort()
fig, ax = plt.subplots()
ax.boxplot(
feature_importances.importances[sorted_idx].T,
vert=False,
labels=X_train_augmented.columns[sorted_idx],
)
ax.set_title("Permutation Importances (train set)")
fig.tight_layout()
plt.show()
```
## Submission
To submit your code, you can refer to the [online documentation](https://paris-saclay-cds.github.io/ramp-docs/ramp-workflow/stable/using_kits.html).
Next steps could be,
- more in depth exploratory analysis
- use more external data (you can add your data to `external_data.csv` but it needs to remain a single file)
- more advanced feature extraction and modeling (see for [this scikit-learn tutorial](https://scikit-learn.org/stable/auto_examples/applications/plot_cyclical_feature_engineering.html#sphx-glr-auto-examples-applications-plot-cyclical-feature-engineering-py))
- hyper-parameter search with GridSearchCV
| github_jupyter |
<h1>Functions in Python</h1>
<h2 id="func">Functions</h2>
Function ဆိုတာ ၎င်းထဲရှိ သတ်မှတ်ထားသော လုပ်ဆောင်ချက်များအတွက် သီးခြားထုတ်ရေးထားသော ပြန်လည်သုံးလို့ရသည့် code အပိုင်းဖြစ်တယ်။
functions</b>
- <b>
ပါလာပြီးတဲ့ fuction
</b>
- <b>ကိုယ်တိုင်ဖန်တီးထားတဲ့ function
</b>
<h3 id="content">What is a Function?</h3>
<img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%203/Images/FuncsDefinition.png" width="500" />
```
# First function example: Add 1 to a and store as b
def add(a):
b = a + 1
print(a, "if you add one", b)
return(b)
```
The figure below illustrates the terminology:
We can obtain help about a function :
```
# Get a help on add function
help(add)
```
We can call the function:
```
# Call the function add()
add(1)
```
If we call the function with a new input we get a new result:
```
# Call the function add()
add(2)
```
We can create different functions. For example, we can create a function that multiplies two numbers. The numbers will be represented by the variables <code>a</code> and <code>b</code>:
```
# Define a function for multiple two numbers
def Mult(a, b):
c = a * b
return(c)
```
The same function can be used for different data types. For example, we can multiply two integers:
```
# Use mult() multiply two integers
Mult(2, 3)
```
Two Floats:
```
# Use mult() multiply two floats
Mult(10.0, 3.14)
```
We can even replicate a string by multiplying with an integer:
```
# Use mult() multiply two different type values together
Mult(2, "Michael Jackson ")
```
<h3 id="var">Variables</h3>
The input to a function is called a formal parameter.
A variable that is declared inside a function is called a local variable. The parameter only exists within the function (i.e. the point where the function starts and stops).
A variable that is declared outside a function definition is a global variable, and its value is accessible and modifiable throughout the program. We will discuss more about global variables at the end of the lab.
```
# Function Definition
def square(a):
# Local variable b
b = 1
c = a * a + b
print(a, "if you square + 1", c)
return(c)
```
The labels are displayed in the figure:
<img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%203/Images/FuncsVar.png" width="500" />
We can call the function with an input of <b>3</b>:
```
# Initializes Global variable
x = 3
# Makes function call and return function a y
y = square(x)
y
```
We can call the function with an input of <b>2</b> in a different manner:
```
# Directly enter a number as parameter
square(2)
```
If there is no <code>return</code> statement, the function returns <code>None</code>. The following two functions are equivalent:
```
# Define functions, one with return value None and other without return value
def MJ():
print('Michael Jackson')
def MJ1():
print('Michael Jackson')
return(None)
# See the output
MJ()
# See the output
MJ1()
```
Printing the function after a call reveals a **None** is the default return statement:
```
# See what functions returns are
print(MJ())
print(MJ1())
```
Create a function <code>con</code> that concatenates two strings using the addition operation:
```
# Define the function for combining strings
def con(a, b):
return(a + b)
# Test on the con() function
con("This ", "is")
```
<hr/>
<div class="alert alert-success alertsuccess" style="margin-top: 20px">
<h4> [Tip] How do I learn more about the pre-defined functions in Python? </h4>
<p>We will be introducing a variety of pre-defined functions to you as you learn more about Python. There are just too many functions, so there's no way we can teach them all in one sitting. But if you'd like to take a quick peek, here's a short reference card for some of the commonly-used pre-defined functions: <a href="http://www.astro.up.pt/~sousasag/Python_For_Astronomers/Python_qr.pdf">Reference</a></p>
</div>
<hr/>
<h3 id="simple">Functions Make Things Simple</h3>
Consider the two lines of code in <b>Block 1</b> and <b>Block 2</b>: the procedure for each block is identical. The only thing that is different is the variable names and values.
<h4>Block 1:</h4>
```
# a and b calculation block1
a1 = 4
b1 = 5
c1 = a1 + b1 + 2 * a1 * b1 - 1
if(c1 < 0):
c1 = 0
else:
c1 = 5
c1
```
<h4>Block 2:</h4>
```
# a and b calculation block2
a2 = 0
b2 = 0
c2 = a2 + b2 + 2 * a2 * b2 - 1
if(c2 < 0):
c2 = 0
else:
c2 = 5
c2
```
We can replace the lines of code with a function. A function combines many instructions into a single line of code. Once a function is defined, it can be used repeatedly. You can invoke the same function many times in your program. You can save your function and use it in another program or use someone else’s function. The lines of code in code <b>Block 1</b> and code <b>Block 2</b> can be replaced by the following function:
```
# Make a Function for the calculation above
def Equation(a,b):
c = a + b + 2 * a * b - 1
if(c < 0):
c = 0
else:
c = 5
return(c)
```
This function takes two inputs, a and b, then applies several operations to return c.
We simply define the function, replace the instructions with the function, and input the new values of <code>a1</code>, <code>b1</code> and <code>a2</code>, <code>b2</code> as inputs. The entire process is demonstrated in the figure:
<img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%203/Images/FuncsPros.gif" width="850" />
Code **Blocks 1** and **Block 2** can now be replaced with code **Block 3** and code **Block 4**.
<h4>Block 3:</h4>
```
a1 = 4
b1 = 5
c1 = Equation(a1, b1)
c1
```
<h4>Block 4:</h4>
```
a2 = 0
b2 = 0
c2 = Equation(a2, b2)
c2
```
<hr>
<h2 id="pre">Pre-defined functions</h2>
There are many pre-defined functions in Python, so let's start with the simple ones.
The <code>print()</code> function:
```
# Build-in function print()
album_ratings = [10.0, 8.5, 9.5, 7.0, 7.0, 9.5, 9.0, 9.5]
print(album_ratings)
```
The <code>sum()</code> function adds all the elements in a list or tuple:
```
# Use sum() to add every element in a list or tuple together
sum(album_ratings)
```
The <code>len()</code> function returns the length of a list or tuple:
```
# Show the length of the list or tuple
len(album_ratings)
```
<h2 id="if">Using <code>if</code>/<code>else</code> Statements and Loops in Functions</h2>
The <code>return()</code> function is particularly useful if you have any IF statements in the function, when you want your output to be dependent on some condition:
```
# Function example
def type_of_album(artist, album, year_released):
print(artist, album, year_released)
if year_released > 1980:
return "Modern"
else:
return "Oldie"
x = type_of_album("Michael Jackson", "Thriller", 1980)
print(x)
```
We can use a loop in a function. For example, we can <code>print</code> out each element in a list:
```
# Print the list using for loop
def PrintList(the_list):
for element in the_list:
print(element)
# Implement the printlist function
PrintList(['1', 1, 'the man', "abc"])
```
<hr>
<h2 id="default">Setting default argument values in your custom functions</h2>
You can set a default value for arguments in your function. For example, in the <code>isGoodRating()</code> function, what if we wanted to create a threshold for what we consider to be a good rating? Perhaps by default, we should have a default rating of 4:
```
# Example for setting param with default value
def isGoodRating(rating=4):
if(rating < 7):
print("this album sucks it's rating is",rating)
else:
print("this album is good its rating is",rating)
# Test the value with default value and with input
isGoodRating()
isGoodRating(10)
```
<hr>
<h2 id="global">Global variables</h2>
So far, we've been creating variables within functions, but we have not discussed variables outside the function. These are called global variables.
<br>
Let's try to see what <code>printer1</code> returns:
```
# Example of global variable
artist = "Michael Jackson"
def printer1(artist):
internal_var = artist
print(artist, "is an artist")
printer1(artist)
```
If we print <code>internal_var</code> we get an error.
<b>We got a Name Error: <code>name 'internal_var' is not defined</code>. Why?</b>
It's because all the variables we create in the function is a <b>local variable</b>, meaning that the variable assignment does not persist outside the function.
But there is a way to create <b>global variables</b> from within a function as follows:
```
artist = "Michael Jackson"
def printer(artist):
global internal_var
internal_var= "Whitney Houston"
print(artist,"is an artist")
printer(artist)
printer(internal_var)
```
<h2 id="scope">Scope of a Variable</h2>
The scope of a variable is the part of that program where that variable is accessible. Variables that are declared outside of all function definitions, such as the <code>myFavouriteBand</code> variable in the code shown here, are accessible from anywhere within the program. As a result, such variables are said to have global scope, and are known as global variables.
<code>myFavouriteBand</code> is a global variable, so it is accessible from within the <code>getBandRating</code> function, and we can use it to determine a band's rating. We can also use it outside of the function, such as when we pass it to the print function to display it:
```
# Example of global variable
myFavouriteBand = "AC/DC"
def getBandRating(bandname):
if bandname == myFavouriteBand:
return 10.0
else:
return 0.0
print("AC/DC's rating is:", getBandRating("AC/DC"))
print("Deep Purple's rating is:",getBandRating("Deep Purple"))
print("My favourite band is:", myFavouriteBand)
```
Take a look at this modified version of our code. Now the <code>myFavouriteBand</code> variable is defined within the <code>getBandRating</code> function. A variable that is defined within a function is said to be a local variable of that function. That means that it is only accessible from within the function in which it is defined. Our <code>getBandRating</code> function will still work, because <code>myFavouriteBand</code> is still defined within the function. However, we can no longer print <code>myFavouriteBand</code> outside our function, because it is a local variable of our <code>getBandRating</code> function; it is only defined within the <code>getBandRating</code> function:
```
# Example of local variable
def getBandRating(bandname):
myFavouriteBand = "AC/DC"
if bandname == myFavouriteBand:
return 10.0
else:
return 0.0
print("AC/DC's rating is: ", getBandRating("AC/DC"))
print("Deep Purple's rating is: ", getBandRating("Deep Purple"))
print("My favourite band is", myFavouriteBand)
```
Finally, take a look at this example. We now have two <code>myFavouriteBand</code> variable definitions. The first one of these has a global scope, and the second of them is a local variable within the <code>getBandRating</code> function. Within the <code>getBandRating</code> function, the local variable takes precedence. **Deep Purple** will receive a rating of 10.0 when passed to the <code>getBandRating</code> function. However, outside of the <code>getBandRating</code> function, the <code>getBandRating</code> s local variable is not defined, so the <code>myFavouriteBand</code> variable we print is the global variable, which has a value of **AC/DC**:
```
# Example of global variable and local variable with the same name
myFavouriteBand = "AC/DC"
def getBandRating(bandname):
myFavouriteBand = "Deep Purple"
if bandname == myFavouriteBand:
return 10.0
else:
return 0.0
print("AC/DC's rating is:",getBandRating("AC/DC"))
print("Deep Purple's rating is: ",getBandRating("Deep Purple"))
print("My favourite band is:",myFavouriteBand)
```
| github_jupyter |
<center><h1>Kalman and Bayesian Filters in Python</h1></center>
<p>
<p>
## Table of Contents
[**Preface**](http://nbviewer.ipython.org/urls/raw.github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python/master/00-Preface.ipynb)
Motivation behind writing the book. How to download and read the book. Requirements for IPython Notebook and Python. github links.
[**Chapter 1: The g-h Filter**](http://nbviewer.ipython.org/urls/raw.github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python/master/01-g-h-filter.ipynb)
Intuitive introduction to the g-h filter, also known as the $\alpha$-$\beta$ Filter, which is a family of filters that includes the Kalman filter. Once you understand this chapter you will understand the concepts behind the Kalman filter.
[**Chapter 2: The Discrete Bayes Filter**](http://nbviewer.ipython.org/urls/raw.github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python/master/02-Discrete-Bayes.ipynb)
Introduces the discrete Bayes filter. From this you will learn the probabilistic (Bayesian) reasoning that underpins the Kalman filter in an easy to digest form.
[**Chapter 3: Gaussian Probabilities**](http://nbviewer.ipython.org/urls/raw.github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python/master/03-Gaussians.ipynb)
Introduces using Gaussians to represent beliefs in the Bayesian sense. Gaussians allow us to implement the algorithms used in the discrete Bayes filter to work in continuous domains.
[**Chapter 4: One Dimensional Kalman Filters**](http://nbviewer.ipython.org/urls/raw.github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python/master/04-One-Dimensional-Kalman-Filters.ipynb)
Implements a Kalman filter by modifying the discrete Bayes filter to use Gaussians. This is a full featured Kalman filter, albeit only useful for 1D problems.
[**Chapter 5: Multivariate Gaussians**](http://nbviewer.ipython.org/urls/raw.github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python/master/05-Multivariate-Gaussians.ipynb)
Extends Gaussians to multiple dimensions, and demonstrates how 'triangulation' and hidden variables can vastly improve estimates.
[**Chapter 6: Multivariate Kalman Filter**](http://nbviewer.ipython.org/urls/raw.github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python/master/06-Multivariate-Kalman-Filters.ipynb)
We extend the Kalman filter developed in the univariate chapter to the full, generalized filter for linear problems. After reading this you will understand how a Kalman filter works and how to design and implement one for a (linear) problem of your choice.
[**Chapter 7: Kalman Filter Math**](http://nbviewer.ipython.org/urls/raw.github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python/master/07-Kalman-Filter-Math.ipynb)
We gotten about as far as we can without forming a strong mathematical foundation. This chapter is optional, especially the first time, but if you intend to write robust, numerically stable filters, or to read the literature, you will need to know the material in this chapter. Some sections will be required to understand the later chapters on nonlinear filtering.
[**Chapter 8: Designing Kalman Filters**](http://nbviewer.ipython.org/urls/raw.github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python/master/08-Designing-Kalman-Filters.ipynb)
Building on material in Chapters 5 and 6, walks you through the design of several Kalman filters. Only by seeing several different examples can you really grasp all of the theory. Examples are chosen to be realistic, not 'toy' problems to give you a start towards implementing your own filters. Discusses, but does not solve issues like numerical stability.
[**Chapter 9: Nonlinear Filtering**](http://nbviewer.ipython.org/urls/raw.github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python/master/09-Nonlinear-Filtering.ipynb)
Kalman filters as covered only work for linear problems. Yet the world is nonlinear. Here I introduce the problems that nonlinear systems pose to the filter, and briefly discuss the various algorithms that we will be learning in subsequent chapters.
[**Chapter 10: Unscented Kalman Filters**](http://nbviewer.ipython.org/urls/raw.github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python/master/10-Unscented-Kalman-Filter.ipynb)
Unscented Kalman filters (UKF) are a recent development in Kalman filter theory. They allow you to filter nonlinear problems without requiring a closed form solution like the Extended Kalman filter requires.
This topic is typically either not mentioned, or glossed over in existing texts, with Extended Kalman filters receiving the bulk of discussion. I put it first because the UKF is much simpler to understand, implement, and the filtering performance is usually as good as or better then the Extended Kalman filter. I always try to implement the UKF first for real world problems, and you should also.
[**Chapter 11: Extended Kalman Filters**](http://nbviewer.ipython.org/urls/raw.github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python/master/11-Extended-Kalman-Filters.ipynb)
Extended Kalman filters (EKF) are the most common approach to linearizing non-linear problems. A majority of real world Kalman filters are EKFs, so will need to understand this material to understand existing code, papers, talks, etc.
[**Chapter 12: Particle Filters**](http://nbviewer.ipython.org/urls/raw.github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python/master/12-Particle-Filters.ipynb)
Particle filters uses Monte Carlo techniques to filter data. They easily handle highly nonlinear and non-Gaussian systems, as well as multimodal distributions (tracking multiple objects simultaneously) at the cost of high computational requirements.
[**Chapter 13: Smoothing**](http://nbviewer.ipython.org/urls/raw.github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python/master/13-Smoothing.ipynb)
Kalman filters are recursive, and thus very suitable for real time filtering. However, they work extremely well for post-processing data. After all, Kalman filters are predictor-correctors, and it is easier to predict the past than the future! We discuss some common approaches.
[**Chapter 14: Adaptive Filtering**](http://nbviewer.ipython.org/urls/raw.github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python/master/14-Adaptive-Filtering.ipynb)
Kalman filters assume a single process model, but manuevering targets typically need to be described by several different process models. Adaptive filtering uses several techniques to allow the Kalman filter to adapt to the changing behavior of the target.
[**Appendix A: Installation, Python, NumPy, and FilterPy**](http://nbviewer.ipython.org/urls/raw.github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python/master/Appendix-A-Installation.ipynb)
Brief introduction of Python and how it is used in this book. Description of the companion
library FilterPy.
[**Appendix B: Symbols and Notations**](http://nbviewer.ipython.org/urls/raw.github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python/master/Appendix-B-Symbols-and-Notations.ipynb)
Most books opt to use different notations and variable names for identical concepts. This is a large barrier to understanding when you are starting out. I have collected the symbols and notations used in this book, and built tables showing what notation and names are used by the major books in the field.
*Still just a collection of notes at this point.*
[**Appendix D: H-Infinity Filters**](http://nbviewer.ipython.org/urls/raw.github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python/master/Appendix-D-HInfinity-Filters.ipynb)
Describes the $H_\infty$ filter.
*I have code that implements the filter, but no supporting text yet.*
[**Appendix E: Ensemble Kalman Filters**](http://nbviewer.ipython.org/urls/raw.github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python/master/Appendix-E-Ensemble-Kalman-Filters.ipynb)
Discusses the ensemble Kalman Filter, which uses a Monte Carlo approach to deal with very large Kalman filter states in nonlinear systems.
[**Appendix F: FilterPy Source Code**](http://nbviewer.ipython.org/urls/raw.github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python/master/Appendix-F-Filterpy-Code.ipynb)
Listings of important classes from FilterPy that are used in this book.
## Supporting Notebooks
These notebooks are not a primary part of the book, but contain information that might be interested to a subest of readers.
[**Computing and plotting PDFs**](http://nbviewer.ipython.org/urls/raw.github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python/master/Supporting_Notebooks/Computing_and_plotting_PDFs.ipynb)
Describes how I implemented the plotting of various pdfs in the book.
[**Interactions**](http://nbviewer.ipython.org/urls/raw.github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python/master/Supporting_Notebooks/Interactions.ipynb)
Interactive simulations of various algorithms. Use sliders to change the output in real time.
[**Converting the Multivariate Equations to the Univariate Case**](http://nbviewer.ipython.org/urls/raw.github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python/master/Supporting_Notebooks/Converting-Multivariate-Equations-to-Univariate.ipynb)
Demonstrates that the Multivariate equations are identical to the univariate Kalman filter equations by setting the dimension of all vectors and matrices to one.
[**Iterative Least Squares for Sensor Fusion**](http://nbviewer.ipython.org/urls/raw.github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python/master/Supporting_Notebooks/Iterative-Least-Squares-for-Sensor-Fusion.ipynb)
Deep dive into using an iterative least squares technique to solve the nonlinear problem of finding position from multiple GPS pseudorange measurements.
[**Taylor Series**](http://nbviewer.ipython.org/urls/raw.github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python/master/Supporting_Notebooks/Taylor-Series.ipynb)
A very brief introduction to Taylor series.
### Github repository
http://github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python
```
#format the book
from book_format import load_style
load_style()
```
| github_jupyter |
# Preprocessing Text Data
The purpose of this notebook is to demonstrate how to preprocessing text data for next-step feature engineering and training a machine learning model via Amazon SageMaker. In this notebook we will focus on preprocessing our text data, and we will use the text data we ingested in a [sequel notebook](https://sagemaker-examples.readthedocs.io/en/latest/data_ingestion/012_Ingest_text_data_v2.html) to showcase text data preprocessing methodologies. We are going to discuss many possible methods to clean and enrich your text, but you do not need to run through every single step below. Usually, a rule of thumb is: if you are dealing with very noisy text, like social media text data, or nurse notes, then medium to heavy preprocessing effort might be needed, and if it's domain-specific corpus, text enrichment is helpful as well; if you are dealing with long and well-written documents such as news articles and papers, very light preprocessing is needed; you can add some enrichment to the data to better capture the sentence to sentence relationship and overall meaning.
## Overview
### Input Format
Labeled text data sometimes are in a structured data format. You might come across this when working on reviews for sentiment analysis, news headlines for topic modeling, or documents for text classification. One column of the dataset could be dedicated for the label, one column for the text, and sometimes other columns as attributes. You can process this dataset format similar to how you would process tabular data and ingest them in the [last section](https://github.com/aws/amazon-sagemaker-examples/blob/master/preprocessing/tabular_data/preprocessing_tabular_data.ipynb). Sometimes text data, especially raw text data, comes as unstructured data and is often in .json or .txt format. To work with this type of formatting, you will need to first extract useful information from the original dataset.
### Use Cases
Text data contains rich information and it's everywhere. Applicable use cases include Voice of Customer (VOC), fraud detection, warranty analysis, chatbot and customer service routing, audience analysis, and much more.
### What's the difference between preprocessing and feature engineering for text data?
In the preprocessing stage, you want to clean and transfer the text data from human language to standard, machine-analyzable format for further processing. For feature engineering, you extract predictive factors (features) from the text. For example, for a matching equivalent question pairs task, the features you can extract include words overlap, cosine similarity, inter-word relationships, parse tree structure similarity, TF-IDF (frequency-inverse document frequency) scores, etc.; for some language model like topic modeling, words embeddings themselves can also be features.
### When is my text data ready for feature engineering?
When the data is ready to be vectorized and fit your specific use case.
## Set Up Notebook
There are several python packages designed specifically for natural language processing (NLP) tasks. In this notebook, you will use the following packages:
* [`nltk`(natrual language toolkit)](https://www.nltk.org/), a leading platform includes multiple text processing libraries, which covers almost all aspects of preprocessing we will discuss in this section: tokenization, stemming, lemmatization, parsing, chunking, POS tagging, stop words, etc.
* [`SpaCy`](https://spacy.io/), offers most functionality provided by `nltk`, and provides pre-trained word vectors and models. It is scalable and designed for production usage.
* [`Gensim` (Generate Similar)](https://radimrehurek.com/gensim/about.html), "designed specifically for topic modeling, document indexing, and similarity retrieval with large corpora".
* [`TextBlob`](), offers POS tagging, noun phrases extraction, sentiment analysis, classification, parsing, n-grams, word inflation, all offered as an API to perform more advanced NLP tasks. It is an easy-to-use wrapper for libraries like `nltk` and `Pattern`. We will use this package for our enrichment tasks.
```
%pip install -qU 'sagemaker>=2.15.0' spacy gensim textblob emot autocorrect
import nltk
import spacy
import gensim
from textblob import TextBlob
import re
import string
import glob
import sagemaker
# Get SageMaker session & default S3 bucket
sagemaker_session = sagemaker.Session()
bucket = sagemaker_session.default_bucket() # replace with your own bucket if you have one
s3 = sagemaker_session.boto_session.resource('s3')
prefix = 'text_sentiment140/sentiment140'
filename = 'training.1600000.processed.noemoticon.csv'
```
### Downloading data from Online Sources
### Text Data Sets: Twitter -- sentiment140
**Sentiment140** The sentiment140 dataset contains 1.6M tweets that were extracted using the [Twitter API](https://developer.twitter.com/en/products/twitter-api) . The tweets have been annotated with sentiment (0 = negative, 4 = positive) and topics (hashtags used to retrieve tweets). The dataset contains the following columns:
* `target`: the polarity of the tweet (0 = negative, 4 = positive)
* `ids`: The id of the tweet ( 2087)
* `date`: the date of the tweet (Sat May 16 23:58:44 UTC 2009)
* `flag`: The query (lyx). If there is no query, then this value is NO_QUERY.
* `user`: the user that tweeted (robotickilldozr)
* `text`: the text of the tweet (Lyx is cool)
```
#helper functions to upload data to s3
def write_to_s3(filename, bucket, prefix):
#put one file in a separate folder. This is helpful if you read and prepare data with Athena
filename_key = filename.split('.')[0]
key = "{}/{}/{}".format(prefix,filename_key,filename)
return s3.Bucket(bucket).upload_file(filename,key)
def upload_to_s3(bucket, prefix, filename):
url = 's3://{}/{}/{}'.format(bucket, prefix, filename)
print('Writing to {}'.format(url))
write_to_s3(filename, bucket, prefix)
#run this cell if you are in SageMaker Studio notebook
#!apt-get install unzip
!wget http://cs.stanford.edu/people/alecmgo/trainingandtestdata.zip -O sentimen140.zip
# Uncompressing
!unzip -o sentimen140.zip -d sentiment140
#upload the files to the S3 bucket
csv_files = glob.glob("sentiment140/*.csv")
for filename in csv_files:
upload_to_s3(bucket, 'text_sentiment140', filename)
```
## Read in Data
We will read the data in as .csv format since the text is embedded in a structured table.
<b>Note:</b> A frequent error when reading in text data is the encoding error. You can try different encoding options with pandas read_csv when "encoding as UTF-8" does not work; see [python encoding documentation](https://docs.python.org/3/library/codecs.html#standard-encodings) for more encodings you may encounter.
```
import pandas as pd
prefix = 'text_sentiment140/sentiment140'
filename = 'training.1600000.processed.noemoticon.csv'
data_s3_location = "s3://{}/{}/{}".format(bucket, prefix, filename) # S3 URL
# we will showcase with a smaller subset of data for demonstration purpose
text_data = pd.read_csv(data_s3_location, header = None,
encoding = "ISO-8859-1", low_memory=False,
nrows = 10000)
text_data.columns = ['target', 'tw_id', 'date', 'flag', 'user', 'text']
```
## Examine Your Text Data
Here you will explore common methods and steps for text preprocessing. Text preprocessing is highly specific to each individual corpus and different tasks, so it is important to examine your text data first and decide what steps are necessary.
First, look at your text data. Seems like there are whitespaces to trim, URLs, smiley faces, numbers, abbreviations, spelling, names, etc. Tweets are less than 140 characters so there is less need for document segmentation and sentence dependencies.
```
pd.set_option('display.max_colwidth', None) #show full content in a column
text_data['text'][:5]
```
## Preprocessing
### Step 1: Noise Removal
Start by removing noise from the text data. Removing noise is very task-specific, so you will usually pick and choose from the following to process your text data based on your needs:
* Remove formatting (HTML, markup, metadata) -- e.g. emails, web-scrapped data
* Extract text data from full dataset -- e.g. reviews, comments, labeled data from a nested JSON file or from structured data
* Remove special characters
* Remove emojis or convert emoji to words -- e.g. reviews, tweets, Instagram and Facebook comments, SMS text with sales
* Remove URLs -- reviews, web content, emails
* Convert accented characters to ASCII characters -- e.g. tweets, contents that may contain foreign language
Note that preprocessing is an iterative process, so it is common to revisit any of these steps after you have cleaned and normalized your data.
Here you will look at tweets and decide how you are going to process URL, emojis and emoticons.
Working with text will often means dealing with regular expression. To freshen up on your regex or if you are new, [Pythex](https://pythex.org/) is a good helper page for you to find cheatsheet and test your functions.
#### Noise Removal - Remove URLs
```
def remove_urls(text):
url = re.compile(r'https?://\S+|www\.\S+')
return url.sub(r'', text)
```
Let's check if our code works with one example:
```
print (text_data['text'][0])
print ('Removed URL:' + remove_urls(text_data['text'][0]))
```
#### Noise Removal - Remove emoticons, or convert emoticons to words
```
from emot.emo_unicode import UNICODE_EMO, EMOTICONS
def remove_emoticons(text):
"""
This function takes strings containing emoticons and returns strings with emoticons removed.
Input(string): one tweet, contains emoticons
Output(string): one tweet, emoticons removed, everything else unchanged
"""
emoticon = re.compile(u'(' + u'|'.join(k for k in EMOTICONS) + u')')
return emoticon.sub(r'', text)
def convert_emoticons(text):
"""
This function takes strings containing emoticons and convert the emoticons to words that describe the emoticon.
Input(string): one tweet, contains emoticons
Output(string): one tweet, emoticons replaced with words describing the emoticon
"""
for emot in EMOTICONS:
text = re.sub(u'('+emot+')', " ".join(EMOTICONS[emot].replace(",","").split()), text)
return text
```
Let's check the results with one example and decide if we should keep the emoticon:
```
print ('original text: ' +remove_emoticons(text_data['text'][0]))
print ('removed emoticons: ' + convert_emoticons(text_data['text'][0]))
```
Assuming our task is sentiment analysis, then converting emoticons to words will be helpful.
We will apply our remove_URL and convert_emoticons functions to the full dataset:
```
text_data['cleaned_text'] = text_data['text'].apply(remove_urls).apply(convert_emoticons)
text_data[['text', 'cleaned_text']][:1]
```
## Step 2: Normalization
In the next step, we will further process the text so that all text/words will be put on the same level playing field: all the words should be in the same case, numbers should be also treated as strings, abbreviations and chat words should be recognizable and replaced with the full words, etc. This is important because we do not want two elements in our word list (dictionary) with the same meaning are taken as two non-related different words by machine, and when we eventually convert all words to numbers (vectors), these words will be noises to our model, such as "3" and "three", "Our" and "our", or "urs" and "yours". This process often includes the following steps:
### Step 2.1 General Normalization
* Convert all text to the same case
* Remove punctuation
* Convert numbers to word or remove numbers depending on your task
* Remove white spaces
* Convert abbreviations/slangs/chat words to word
* Remove stop words (task specific and general English words); you can also create your own list of stop words
* Remove rare words
* Spelling correction
**Note:** some normalization processes are better to perform at sentence and document level, and some processes are word-level and should happen after tokenization and segmentation, which we will cover right after normalization.
Here you will convert the text to lower case, remove punctuation, remove numbers, remove white spaces, and complete other word-level processing steps after tokenizing the sentences.
#### Normalization - Convert all text to lower case
Usually, this is a must for all language preprocessing. Since "Word" and "word" will essentially be considered two different elements in word representation, and we want words that have the same meaning to be represented the same in numbers (vectors), we want to convert all text into the same case.
```
text_data['text_lower'] = text_data['cleaned_text'].str.lower()
text_data[['cleaned_text', 'text_lower']][:1]
```
#### Normalization - Remove numbers
Depending on your use cases, you can either remove numbers or convert numbers into strings. If numbers are not important in your task (e.g. sentiment analysis) you can remove those, and in some cases, numbers are useful (e.g. date), and you can tag these numbers differently. In most pre-trained embeddings, numbers are treated as strings.
In this example, we are using Twitter data (tweets) and typically, numbers are not that important for understanding the meaning or content of a tweet. Therefore, we will remove the numbers.
```
def remove_numbers(text):
'''
This function takes strings containing numbers and returns strings with numbers removed.
Input(string): one tweet, contains numbers
Output(string): one tweet, numbers removed
'''
return re.sub(r'\d+', '', text)
#let's check the results of our function
remove_numbers(text_data['text_lower'][2])
text_data['normalized_text'] = text_data['text_lower'].apply(remove_numbers)
```
#### Twitter data specific: Remove mentions or extract mentions into a different column
We can remove the mentions in the tweets, but if our task is to monitor VOC, it is helpful to extract the mentions data.
```
def remove_mentions(text):
'''
This function takes strings containing mentions and returns strings with
mentions (@ and the account name) removed.
Input(string): one tweet, contains mentions
Output(string): one tweet, mentions (@ and the account name mentioned) removed
'''
mentions = re.compile(r'@\w+ ?')
return mentions.sub(r'', text)
print('original text: ' + text_data['text_lower'][0])
print('removed mentions: ' + remove_mentions(text_data['text_lower'][0]))
def extract_mentions(text):
'''
This function takes strings containing mentions and returns strings with
mentions (@ and the account name) extracted into a different element,
and removes the mentions in the original sentence.
Input(string): one sentence, contains mentions
Output:
one tweet (string): mentions (@ and the account name mentioned) removed
mentions (string): (only the account name mentioned) extracted
'''
mentions = [i[1:] for i in text.split() if i.startswith("@")]
sentence = re.compile(r'@\w+ ?').sub(r'', text)
return sentence,mentions
text_data['normalized_text'], text_data['mentions'] = zip(*text_data['normalized_text'].apply(extract_mentions))
text_data[['text','normalized_text', 'mentions']].head(1)
```
#### Remove Punctuation
We will use the `string.punctuation` in python to remove punctuations, which contains the following punctuation symbols`!"#$%&\'()*+,-./:;<=>?@[\\]^_{|}~`
, you can add or remove more as needed.
```
punc_list = string.punctuation #you can self define list of punctuation to remove here
def remove_punctuation(text):
"""
This function takes strings containing self defined punctuations and returns
strings with punctuations removed.
Input(string): one tweet, contains punctuations in the self-defined list
Output(string): one tweet, self-defined punctuations removed
"""
translator = str.maketrans('', '', punc_list)
return text.translate(translator)
remove_punctuation(text_data['normalized_text'][2])
text_data['normalized_text'] = text_data['normalized_text'].apply(remove_punctuation)
```
#### Remove Whitespaces
You can also use `trim` functions to trim whitespaces from left and right or in the middle. Here we will just simply utilize the `split` function to extract all words from our text since we already removed all special characters, and combine them with a single whitespace.
```
def remove_whitespace(text):
'''
This function takes strings containing mentions and returns strings with
whitespaces removed.
Input(string): one tweet, contains whitespaces
Output(string): one tweet, white spaces removed
'''
return " ".join(text.split())
print('original text: ' + text_data['normalized_text'][2])
print('removed whitespaces: ' + remove_whitespace(text_data['normalized_text'][2]))
text_data['normalized_text'] = text_data['normalized_text'].apply(remove_whitespace)
```
## Step 3: Tokenization and Segmentation
After we extracted useful text data from the full dataset, we will split large chunks of text (documents) into sentences, and sentences into words. Most of the times we will use sentence-ending punctuation to split documents into sentences, but it can be ambiguous especially when we are dealing with character conversations ("Are you alright?" said Ron), abbreviations (Dr. Fay would like to see Mr. Smith now.) and other special use cases. There are Python libraries designed for this task (check [textsplit](https://github.com/chschock/textsplit)), but you can take your own approach depending on your context.
Here for Twitter data, we are only dealing with sentences shorter than 140 characters, so we will just tokenize sentences into words. We do want to normalize the sentence before tokenizing sentences into words, so we will introduce normalization, and tokenize our tweets into words after normalizing sentences.
### Tokenizing Sentences into Words
```
nltk.download('punkt')
from nltk.tokenize import word_tokenize
def tokenize_sent(text):
'''
This function takes strings (a tweet) and returns tokenized words.
Input(string): one tweet
Output(list): list of words tokenized from the tweet
'''
word_tokens = word_tokenize(text)
return word_tokens
text_data['tokenized_text'] = text_data['normalized_text'].apply(tokenize_sent)
text_data[['normalized_text','tokenized_text']][:1]
```
### Continuing Word-level Normalization
#### Remove Stop Words
Stop words are common words that does not contribute to the meaning of a sentence, such as 'the', 'a', 'his'. Most of the time we can remove these words without harming further analysis, but if you want to apply Part-of-Speech (POS) tagging later, be careful with what you removed in this step as they can provide valuable information. You can also add stop words to the list based on your use cases.
```
nltk.download('stopwords')
from nltk.corpus import stopwords
stopwords_list = set(stopwords.words('english'))
```
One way to add words to your stopwords list is to check for most frequent words, especially if you are working with a domain-specific corpus and those words sometimes are not covered by general English stop words. You can also remove rare words from your text data.
Let's check for the most common words in our data. All the words we see in the following example are covered in general English stop words, so we will not add any additional stop words.
```
from collections import Counter
counter = Counter()
for word in [w for sent in text_data["tokenized_text"] for w in sent]:
counter[word] += 1
counter.most_common(10)
```
Let's check for the rarest words now. In this example, infrequently used words mostly consist of misspelled words, which we will later correct, but we can add them to our stop words list as well.
```
#least frequent words
counter.most_common()[:-10:-1]
top_n = 10
bottom_n = 10
stopwords_list |= set([word for (word, count) in counter.most_common(top_n)])
stopwords_list |= set([word for (word, count) in counter.most_common()[:-bottom_n:-1]])
stopwords_list |= {'thats'}
def remove_stopwords(tokenized_text):
'''
This function takes a list of tokenized words from a tweet, removes self-defined stop words from the list,
and returns the list of words with stop words removed
Input(list): a list of tokenized words from a tweet, contains stop words
Output(list): a list of words with stop words removed
'''
filtered_text = [word for word in tokenized_text if word not in stopwords_list]
return filtered_text
print(text_data['tokenized_text'][2])
print(remove_stopwords(text_data['tokenized_text'][2]))
text_data['tokenized_text'] = text_data['tokenized_text'].apply(remove_stopwords)
```
#### Convert Abbreviations, slangs and chat words into words
Sometimes you will need to develop your own mapping for abbreviations/slangs <-> words, for chat data, or for domain-specific data where abbreviations often have different meanings from what is commonly used.
```
chat_words_map = {
'idk': 'i do not know',
'btw': 'by the way',
'imo': 'in my opinion',
'u': 'you',
'oic': 'oh i see'
}
chat_words_list = set(chat_words_map)
def translator(text):
"""
This function takes a list of tokenized words, finds the chat words in the self-defined chat words list,
and replace the chat words with the mapped full expressions. It returns the list of tokenized words with
chat words replaced.
Input(list): a list of tokenized words from a tweet, contains chat words
Output(list): a list of words with chat words replaced by full expressions
"""
new_text = []
for w in text:
if w in set(chat_words_map):
new_text = new_text + chat_words_map[w].split()
else:
new_text.append(w)
return new_text
print(text_data['tokenized_text'][13])
print(translator(text_data['tokenized_text'][13]))
text_data['tokenized_text'] = text_data['tokenized_text'].apply(translator)
```
#### Spelling Correction
Some common spelling correction packages include `SpellChecker` and `autocorrect`. It might take some time to spell check every sentence of the text, so you can decide if a spell check is absolutely necessary. If you are dealing with documents (news, papers, articles) generally it is not necessary; but if you are dealing with chat data, reviews, notes, it might be a good idea to spell check your text.
```
from autocorrect import Speller
spell = Speller(lang='en', fast = True)
def spelling_correct(tokenized_text):
"""
This function takes a list of tokenized words from a tweet, spell check every words and returns the
corrected words if applicable. Note that not every wrong spelling words will be identified especially
for tweets.
Input(list): a list of tokenized words from a tweet, contains wrong-spelling words
Output(list): a list of corrected words
"""
corrected = [spell(word) for word in tokenized_text]
return corrected
print(text_data['tokenized_text'][0])
print(spelling_correct(text_data['tokenized_text'][0]))
text_data['tokenized_text'] = text_data['tokenized_text'].apply(spelling_correct)
```
### Step 3.2 [Stemming and Lemmatization](https://en.wikipedia.org/wiki/Lemmatisation)
**Stemming** is the process of removing affixes from a word to get a word stem, and **lemmatization** can in principle select the appropriate lemma depending on the context. The difference is that a stemmer operates on a single word without knowledge of the context, and therefore cannot discriminate between words that have different meanings depending on part of speech. However, stemmers are typically easier to implement and run faster, and the reduced accuracy may not matter for some applications.
#### Stemming
There are several stemming algorithms available, and the most popular ones are Porter, Lancaster, and Snowball. Porter is the most common one, Snowball is an improvement over Porter, and Lancaster is more aggressive. You can check for more algorithms provided by `nltk` [here](https://www.nltk.org/api/nltk.stem.html).
```
from nltk.stem import SnowballStemmer
from nltk.tokenize import word_tokenize
stemmer = SnowballStemmer("english")
def stem_text(tokenized_text):
"""
This function takes a list of tokenized words from a tweet, and returns the stemmed words by your
defined stemmer.
Input(list): a list of tokenized words from a tweet
Output(list): a list of stemmed words in its root form
"""
stems = [stemmer.stem(word) for word in tokenized_text]
return stems
```
#### Lemmatization
```
nltk.download('wordnet')
from nltk.stem import WordNetLemmatizer
from nltk.tokenize import word_tokenize
lemmatizer = WordNetLemmatizer()
def lemmatize_text(tokenized_text):
'''
This function takes a list of tokenized words from a tweet, and returns the lemmatized words.
you can also provide context for lemmatization, i.e. part-of-speech.
Input(list): a list of tokenized words from a tweet
Output(list): a list of lemmatized words in its base form
'''
lemmas = [lemmatizer.lemmatize(word, pos = 'v') for word in tokenized_text]
return lemmas
```
#### Let's compare our stemming and lemmatization results:
It seems like both processes returned similar results besides some verb being trimmed differently, so it is okay to go with stemming in this case if you are dealing with a lot of data and want a better performance. You can also keep both and experiment with further feature engineering and modeling to see which one produces better results.
```
print(text_data['tokenized_text'][2])
print(stem_text(text_data['tokenized_text'][2]))
print(lemmatize_text(text_data['tokenized_text'][2]))
```
It seems that a stemmer can do the work for our tweets data. You can keep both and decide which one you want to use for feature engineering and modeling.
```
text_data['stem_text'] = text_data['tokenized_text'].apply(stem_text)
text_data['lemma_text'] = text_data['tokenized_text'].apply(lemmatize_text)
```
## Step 3.5: Re-examine the results
Take a pause here and examine the results from previous steps to decide if more noise removal/normalization is needed. In this case, you might want to add more words to the stop words list, spell-check more aggressively, or add more mappings to the abbreviation/slang to words list.
```
text_data.sample(5)[['text', 'stem_text', 'lemma_text']]
```
## Step 4: Enrichment and Augmentation
After you have cleaned and tokenized your text data into a standard form, you might want to enrich it with more useful information that was not provided directly in the original text or its single-word form. For example:
* Part-of-speech tagging
* Extracting phrases
* Name entity recognition
* Dependency parsing
* Word level embeddings
Many Python Packages including `nltk`, `SpaCy`, `CoreNLP`, and here we will use `TextBlob` to illustrate some enrichment methods.
#### Part-of-Speech (POS) Tagging
Part-of-Speech tagging can assign each word in accordance with its syntactic functions (noun, verb, adjectives, etc.).
```
nltk.download('averaged_perceptron_tagger')
text_example = text_data.sample()['lemma_text']
text_example
from textblob import TextBlob
result = TextBlob(" ".join(text_example.values[0]))
print(result.tags)
```
#### Extracting Phrases
Sometimes words come in as phrases (noun group phrases, verb group phrases, etc.) and often have discrete grammatical meanings. Extract those words as phrases rather than separate words in this case.
```
nltk.download('brown')
# orginal text:
text_example = text_data.sample()['lemma_text']
" ".join(text_example.values[0])
#noun phrases that can be extracted from this sentence
result = TextBlob(" ".join(text_example.values[0]))
for nouns in result.noun_phrases:
print(nouns)
```
#### Name Entity Recognition (NER)
You can use pre-trained/pre-defined name entity recognition models to find named entities in text and classify them into pre-defined categories. You can also train your own NER model, especially if you are dealing with domain specific context.
```
nltk.download('maxent_ne_chunker')
nltk.download('words')
text_example_enr = text_data.sample()['lemma_text'].values[0]
print("original text: " + " ".join(text_example_enr))
from nltk import pos_tag, ne_chunk
print (ne_chunk(pos_tag(text_example_enr)))
```
## Final Dataset ready for feature engineering and modeling
For this notebook you cleaned and normalized the data, kept mentions as a separate column, and stemmed and lemmatized the tokenized words. You can experiment with these two results to see which one gives you a better model performance.
Twitter data is short and often does not have complex syntax structures, so no enrichment (POS tagging, parsing, etc.) was done at this time; but you can experiment with those when you have more complicated text data.
```
text_data.head(2)
```
### Save our final dataset to S3 for further process
```
filename_write_to = 'processed_sentiment_140.csv'
text_data.to_csv(filename_write_to, index = False)
upload_to_s3(bucket, 'text_sentiment140_processed', filename_write_to)
```
## Conclusion
Congratulations! You cleaned and prepared your text data and it is now ready to be vectorized or used for feature engineering.
Now that your data is ready to be converted into machine-readable format (numbers), we will cover extracting features and word embeddings in the next section **text data feature engineering**.
| github_jupyter |
```
import pandas as pd
import numpy as np
from sklearn.ensemble import RandomForestRegressor
from sklearn.preprocessing import OneHotEncoder, StandardScaler
from sklearn.compose import ColumnTransformer
from sklearn.model_selection import RandomizedSearchCV, cross_val_score, GridSearchCV
import altair as alt
from sklearn.linear_model import LinearRegression, Lasso, Ridge
from sklearn.model_selection import train_test_split
from pyarrow import feather
import pickle
import os.path
from os import path
alt.renderers.enable('notebook')
assert path.exists("../data/train.feather") == True, "Input file does not exist"
# load in feather datasets
# training dataset
avocado = pd.read_feather("../data/train.feather")
avocado.head()
test = pd.read_feather("../data/test.feather")
test_x = test[['lat', 'lon', 'type', 'season']]
test_y = test[['average_price']]
# test dataset
# avocado_test = pd.read_feather("test.feather")
# avocado_test.head()
test_x
# Which features do we want to keep?
avocado.columns
# want season, lat, lon, and type as our features
# Split the data into target and features
avocado_x = avocado[['lat', 'lon', 'type', 'season']]
avocado_y = avocado['average_price']
# avocado_test_x = avocado_test[['region', 'type', 'month']]
# avocado_test_y = avocado['AveragePrice']
# Scale the numerical features (don't need for rfr, but can't hurt)
numeric_features = ['lat', 'lon']
# need to convert categorical to numerical using one-hot-encoding
categorical_features = ['type', 'season']
preprocessor = ColumnTransformer(transformers=[
('scaler', StandardScaler(), numeric_features),
('ohe', OneHotEncoder(), categorical_features)
])
# applying one hot encoding to the training features
avocado_x = pd.DataFrame(preprocessor.fit_transform(avocado_x),
index=avocado_x.index,
columns = (numeric_features +
list(preprocessor.named_transformers_['ohe']
.get_feature_names(categorical_features))))
# apply the same transformation to the test features (but don't fit!)
test_x = pd.DataFrame(preprocessor.transform(test_x),
index=test_x.index,
columns=avocado_x.columns)
assert len(avocado_x.columns) == 8
# fit rfr model
rfr = RandomForestRegressor(random_state=123)
rfr.fit(avocado_x, avocado_y)
fold_accuracies = cross_val_score(estimator=rfr, X=avocado_x, y=avocado_y, cv=5)
print(fold_accuracies)
print(np.mean(fold_accuracies))
print(np.std(fold_accuracies))
# the standard deviation is much greater than the mean accuracy,
# this is probably not a good model...
# find optimal hyperparameters
rfr_parameters = {'max_depth': range(1, 20),
'n_estimators': range(1, 100)}
random_rfr = RandomizedSearchCV(rfr, rfr_parameters, n_iter=10,
cv=5)
random_rfr.fit(avocado_x, avocado_y).best_params_
test_score = np.around(random_rfr.score(avocado_x, avocado_y), 2)
pd.DataFrame({'Test':[test_score]})
np.around(random_rfr.score(test_x, test_y), 2)
fold_accuracies = cross_val_score(estimator=random_rfr, X=avocado_x, y=avocado_y, cv=5)
cv_scores = pd.DataFrame({'Fold': [1, 2, 3, 4, 5],
'Negative_Mean_Squared_Error': np.around(fold_accuracies, 2)})
#for i in range(5):
# cv_scores_dict('Fold')[i] =
#print(fold_accuracies)
#print(np.mean(fold_accuracies))
#print(np.std(fold_accuracies))
# The standard error is much more reasonable here
cv_scores
assert cv_scores['Negative_Mean_Squared_Error'].all() != 0
round(np.mean(cv_scores['Negative_Mean_Squared_Error']), 2)
feature_list = list(avocado_x.columns)
nice_feature_list = ['Latitude', 'Longitude',
'Conventional Type', 'Organic Type',
'Fall Season', 'Spring Season',
'Summer Season', 'Winter Season']
feature_df = pd.DataFrame({"feature_names": nice_feature_list,
"importance": np.around(random_rfr.best_estimator_.feature_importances_, 2)})
feature_df.sort_values(["importance"], ascending=False, inplace=True)
feature_df.reset_index(drop=True, inplace=True)
feature_df
rfr_plot = alt.Chart(feature_df).mark_bar(color="red", opacity=0.6).encode(
y= alt.Y("feature_names:N",
sort=alt.SortField(field='importance:Q'),
title="Features"),
x = alt.X("importance:Q", title="Feature Importance")
).properties(title="Random Forest Regression",
width=200)
rfr_plot
rfr.score(avocado_x, avocado_y)
rfr.best_param_
lr = Lasso()
param_grid = {"alpha": [i for i in range(0, 1000, 1)]}
lr_rs = RandomizedSearchCV(lr, param_grid, cv=5).fit(avocado_x, avocado_y)
lr_rs.best_params_
lr2 = Lasso(alpha=335)
lr2.fit(avocado_x, avocado_y)
#lr.fit(avocado_x, avocado_y)
fold_accuracies_lr = cross_val_score(estimator=lr2, X=avocado_x, y=avocado_y, cv=5)
cv_scores_dict_lr = {'Fold': [1, 2, 3, 4, 5],
'Negative_Mean_Squared_Error': np.around(fold_accuracies_lr, 2)}
pd.DataFrame(cv_scores_dict_lr)
np.mean(cv_scores_dict_lr['Negative_Mean_Squared_Error'])
lr2.score(avocado_x, avocado_y)
list(zip(feature_list, lr2.coef_))
```
Now try L2 regularization
```
r = Ridge()
param_grid = {"alpha": [i for i in range(0, 1000, 1)]}
r_rs = RandomizedSearchCV(r, param_grid, cv=5, random_state=123).fit(avocado_x, avocado_y)
r_rs.best_params_['alpha']
r2 = Ridge(alpha=r_rs.best_params_['alpha'])
r2.fit(avocado_x, avocado_y)
fold_accuracies_r = cross_val_score(estimator=r2, X=avocado_x, y=avocado_y, cv=5)
cv_scores_dict_r = {'Fold': [1, 2, 3, 4, 5],
'Negative_Mean_Squared_Error': np.around(fold_accuracies_r, 2)}
pd.DataFrame(cv_scores_dict_r)
np.mean(cv_scores_dict_r['Negative_Mean_Squared_Error'])
list(zip(feature_list, r2.coef_))
lr_feature_df = pd.DataFrame({"feature_names": nice_feature_list,
"weights": np.around(r2.coef_, 2)})
lr_feature_df = lr_feature_df.sort_values(["weights"], ascending=False)
lr_feature_df
np.around(r2.score(test_x, test_y), 2)
lr_plot = alt.Chart(lr_feature_df).mark_bar(color="blue", opacity=0.6).encode(
y = alt.Y("feature_names:N",
sort=alt.SortField(field="abs(weights):Q"),
title="Features"),
x = alt.X("weights:Q", title="Coefficient Weights")
).properties(title="Linear Regression",
width=200)
lr_plot
rfr_plot | lr_plot
```
| github_jupyter |
# Lab 2
+ ## Автор: Роман Кривохижа
+ ## Група: ІС-72
+ ## Викладач: Новікова П.А.
****
****
****
## Module importing
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style('darkgrid')
%matplotlib inline
```
## Algorithm implementation
+ Алгоритм, який відтворює функціонування системи, за допомогою комп’ютерної програми називається **алгоритмом імітації**.
<font size="4">Ймовірність відмови: $P = \frac{N_{unserv}}{N_{all}}$</font>
<font size="4">Середня довжина черги: $L_{aver} = \frac{\sum_{i} L_i \Delta t_i}{T_{mod}}$</font>
<font size="4">Середній час очікування: $Q_{aver} = \frac{\sum_{i} L_i \Delta t_i}{N_{serv}}$</font>
```
class Rand:
"""
Генерація випадкового числа за заданим законом розподілу
"""
@staticmethod
def exp(time_mean):
a = 0
while a == 0:
a = np.random.rand()
return -time_mean * np.log(a)
class Model:
def __init__(self, delay0, delay1, maxQ=0, print_logs=False):
"""
:param delay0: mean delay of create
:param delay1: mean delay of process
:param maxQ: max size of queue
:type delay0: float
:type delay1: float
:type maxQ: int
"""
self.delay_create = delay0
self.delay_process = delay1
# момент найближчої події
self.tnext = 0.0
# конкретний час
self.tcurr = self.tnext
# час завершення створення
self.t0 = self.tcurr
# час завершення обробки
self.t1 = np.inf
# max size of queue
self.maxqueue = maxQ
# number of created tasks
self.num_create = 0
# number of processed tasks
self.num_process = 0
# number of failure tasks
self.failure = 0
# current state; 0 - вільний
self.state = 0
# current size of queue
self.queue = 0
# ind of next event
self.next_event = 0
self.delta_t_history = []
self.queue_history = []
self.print_logs = print_logs
def simulate(self, time_modeling):
self.time_modeling = time_modeling
while self.tcurr < self.time_modeling:
self.tnext = self.t0
self.next_event = 0
# знайти найменший із моментів часу «момент надходження вимоги у систему»
# та «момент звільнення пристрою обслуговування» і запам’ятати,
# якій події він відповідає
if self.t1 < self.tnext:
self.tnext = self.t1
self.next_event = 1
# просунути поточний час у момент найближчої події
self.tcurr = self.tnext
# виконати подію, яка відповідає моменту найближчої події
if self.next_event == 0:
self.event0()
elif self.next_event == 1:
self.event1()
else:
pass
self.delta_t_history.append(self.tcurr)
self.queue_history.append(self.queue)
if self.print_logs:
self.print_info()
self.print_statistic()
self.print_L_aver()
self.print_Q_aver()
self.print_p_fail()
def print_statistic(self):
"""
Відображення основних статистик
"""
print(f'num_create = {self.num_create}; num_process = {self.num_process}; failure = {self.failure}')
def print_info(self):
"""
Відображення детальної інформації про кожен етап роботи моделі
"""
print(f't = {self.tcurr}; state = {self.state}; queue = {self.queue}')
def print_L_aver(self):
"""
Середня довжина черги черги
"""
t = [self.delta_t_history[0]]
for i in range(0, len(self.delta_t_history)-1):
t.append(self.delta_t_history[i+1] - self.delta_t_history[i])
self.l_aver = np.sum(np.array(t) * np.array(self.queue_history)) / self.time_modeling
print(f'Середня довжина черги черги = {self.l_aver}')
def print_Q_aver(self):
"""
Середній час очікування в черзі
"""
t = [self.delta_t_history[0]]
for i in range(0, len(self.delta_t_history)-1):
t.append(self.delta_t_history[i+1] - self.delta_t_history[i])
self.q_aver = np.sum(np.array(t) * np.array(self.queue_history)) / self.num_process
print(f'Середній час очікування в черзі = {self.q_aver}')
def print_p_fail(self):
"""
Ймовірність відмови
"""
self.p_fail = self.failure / self.num_create
print(f'Ймовірність відмови: {self.p_fail}')
def event0(self):
# подія "натходження"
self.t0 = self.tcurr + self.__get_delay_of_create()
self.num_create += 1
# якщо пристрій вільний, тоді встановлюємо стан "зайнятий"
if self.state == 0:
self.state = 1
# запам'ятаємо час виходу
self.t1 = self.tcurr + self.__get_delay_of_process()
else:
if self.queue < self.maxqueue:
self.queue += 1
else:
self.failure += 1
def event1(self):
# подія "закінчилось обслуговування в пристрої"
self.t1 = np.inf
# стан "вільний"
self.state = 0
# якщо черга вимог не пуста, перемістити одну вимогу із черги у канал обслуговування
if self.queue > 0:
self.queue -= 1
# стан "зайнятий"
self.state = 1
# запам'ятаємо час виходу
self.t1 = self.tcurr + self.__get_delay_of_process()
self.num_process += 1
def __get_delay_of_create(self):
return Rand.exp(self.delay_create)
def __get_delay_of_process(self):
return Rand.exp(self.delay_process)
model = Model(1, 3, 5, print_logs=True)
model.simulate(10)
n_param = 10
delay0_list = list(range(1, n_param+1))
delay1_list = list(range(1, n_param+1))
maxQ_list = list(range(1, n_param+1))
time_modeling_list = [i*1000 for i in range(1, n_param+1)]
np.random.shuffle(delay0_list)
np.random.shuffle(delay1_list)
np.random.shuffle(maxQ_list)
np.random.shuffle(time_modeling_list)
df = pd.DataFrame()
rows = []
for i in range(n_param):
print(f'{i+1} itteration')
model = Model(delay0_list[i], delay1_list[i], maxQ_list[i], print_logs=False)
model.simulate(time_modeling_list[i])
rows.append({'delay_create': delay0_list[i], 'delay_process': delay1_list[i], 'maxqueue': maxQ_list[i], 'time_modeling': model.time_modeling,
'num_create': model.num_create, 'num_process': model.num_process, 'failure': model.failure, 'last_queue_cnt': model.queue,
'l_aver': model.l_aver, 'q_aver': model.q_aver, 'p_fail': model.p_fail})
print()
df = df.append(rows)
```
**Проведемо верифікацію, використовуючи побудовану таблицю:**
```
df
```
## Conclusion
**В данній лабораторній роботі ми реалізували алгоритм імітації простої моделі обслуговування, використовуючи спосіб, який орієнтований на події (аби уникнути випадку, коли дві події приходять в один час).**
+ за допомогою статичних методів була зібрана інформація про роботу/поведінку моделі
+ зміна віхдних параметрів моделі призводить до зміни вихідних значень
+ при досить великій кількості вхідинх параметрів важко проводити етап верифікації моделі
P.S. нижче наведено декілька графіків, які ілюструють залежність вихідних параметрів від вхідних
## Bonus
**Зафіксуємо значення параметра _time_modeling_**
```
n_param = 10
delay0_list = list(range(1, n_param+1))
delay1_list = list(range(1, n_param+1))
maxQ_list = list(range(1, n_param+1))
time_modeling = 100000
np.random.shuffle(delay0_list)
np.random.shuffle(delay1_list)
np.random.shuffle(maxQ_list)
df = pd.DataFrame()
rows = []
for i in range(n_param):
print(f'{i+1} itteration')
model = Model(delay0_list[i], delay1_list[i], maxQ_list[i], print_logs=False)
model.simulate(time_modeling)
rows.append({'delay_create': delay0_list[i], 'delay_process': delay1_list[i], 'maxqueue': maxQ_list[i], 'time_modeling': model.time_modeling,
'num_create': model.num_create, 'num_process': model.num_process, 'failure': model.failure, 'last_queue_cnt': model.queue,
'l_aver': model.l_aver, 'q_aver': model.q_aver, 'p_fail': model.p_fail})
print()
df = df.append(rows)
```
**Побудуємо декілька графіків**
```
fig, ax = plt.subplots(1,1, figsize=(15,6))
sns.barplot(x='delay_create', y='num_create', data=df, ax=ax, color='darkviolet')
ax.set_xlabel(u'delay_create')
# ax.set_xlim(0, 200)
ax.set_ylabel(u'num_create')
ax.set_title(u'');
fig, ax = plt.subplots(1,1, figsize=(15,6))
sns.barplot(x='delay_create', y='failure', data=df, ax=ax, color='darkviolet')
ax.set_xlabel(u'delay_create')
# ax.set_xlim(0, 200)
ax.set_ylabel(u'failure')
ax.set_title(u'');
fig, ax = plt.subplots(1,1, figsize=(15,6))
sns.barplot(x='delay_create', y='p_fail', data=df, ax=ax, color='darkviolet')
ax.set_xlabel(u'delay_create')
# ax.set_xlim(0, 200)
ax.set_ylabel(u'p_fail')
ax.set_title(u'');
fig, ax = plt.subplots(1,1, figsize=(15,6))
sns.barplot(x='delay_process', y='num_process', data=df, ax=ax, color='darkviolet')
ax.set_xlabel(u'delay_process')
# ax.set_xlim(0, 200)
ax.set_ylabel(u'num_process')
ax.set_title(u'');
fig, ax = plt.subplots(1,1, figsize=(15,6))
sns.barplot(x='delay_process', y='failure', data=df, ax=ax, color='darkviolet')
ax.set_xlabel(u'delay_process')
# ax.set_xlim(0, 200)
ax.set_ylabel(u'failure')
ax.set_title(u'');
fig, ax = plt.subplots(1,1, figsize=(15,6))
sns.barplot(x='delay_process', y='p_fail', data=df, ax=ax, color='darkviolet')
ax.set_xlabel(u'delay_process')
# ax.set_xlim(0, 200)
ax.set_ylabel(u'p_fail')
ax.set_title(u'');
```
| github_jupyter |
# Задание 2.2 - Введение в PyTorch
Для этого задания потребуется установить версию PyTorch 1.0
https://pytorch.org/get-started/locally/
В этом задании мы познакомимся с основными компонентами PyTorch и натренируем несколько небольших моделей.<br>
GPU нам пока не понадобится.
Основные ссылки:
https://pytorch.org/tutorials/beginner/deep_learning_60min_blitz.html
https://pytorch.org/docs/stable/nn.html
https://pytorch.org/docs/stable/torchvision/index.html
```
import torch
import torch.nn as nn
import torch.optim as optim
import torchvision.datasets as dset
from torch.utils.data.sampler import SubsetRandomSampler, Sampler
from torchvision import transforms
import matplotlib.pyplot as plt
%matplotlib inline
import numpy as np
PATH_TO_DATA = 'D:/Py/DataFrames/dlcourse_ai/SVHN'
```
## Как всегда, начинаем с загрузки данных
PyTorch поддерживает загрузку SVHN из коробки.
```
# First, lets load the dataset
data_train = dset.SVHN(PATH_TO_DATA, split='train',
transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize(mean=[0.43,0.44,0.47],
std=[0.20,0.20,0.20])
])
)
data_test = dset.SVHN(PATH_TO_DATA, split='test',
transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize(mean=[0.43,0.44,0.47],
std=[0.20,0.20,0.20])
]))
```
Теперь мы разделим данные на training и validation с использованием классов `SubsetRandomSampler` и `DataLoader`.
`DataLoader` подгружает данные, предоставляемые классом `Dataset`, во время тренировки и группирует их в батчи.
Он дает возможность указать `Sampler`, который выбирает, какие примеры из датасета использовать для тренировки. Мы используем это, чтобы разделить данные на training и validation.
Подробнее: https://pytorch.org/tutorials/beginner/data_loading_tutorial.html
```
batch_size = 64
data_size = data_train.data.shape[0]
validation_split = .2
split = int(np.floor(validation_split * data_size))
indices = list(range(data_size))
np.random.shuffle(indices)
train_indices, val_indices = indices[split:], indices[:split]
train_sampler = SubsetRandomSampler(train_indices)
val_sampler = SubsetRandomSampler(val_indices)
train_loader = torch.utils.data.DataLoader(data_train, batch_size=batch_size,
sampler=train_sampler)
val_loader = torch.utils.data.DataLoader(data_train, batch_size=batch_size,
sampler=val_sampler)
```
В нашей задаче мы получаем на вход изображения, но работаем с ними как с одномерными массивами. Чтобы превратить многомерный массив в одномерный, мы воспользуемся очень простым вспомогательным модулем `Flattener`.
```
sample, label = data_train[0]
print("SVHN data sample shape: ", sample.shape)
# As you can see, the data is shaped like an image
# We'll use a special helper module to shape it into a tensor
class Flattener(nn.Module):
def forward(self, x):
batch_size, *_ = x.shape
return x.view(batch_size, -1)
```
И наконец, мы создаем основные объекты PyTorch:
- `nn_model` - собственно, модель с нейросетью
- `loss` - функцию ошибки, в нашем случае `CrossEntropyLoss`
- `optimizer` - алгоритм оптимизации, в нашем случае просто `SGD`
```
nn_model = nn.Sequential(
Flattener(),
nn.Linear(3*32*32, 100),
nn.ReLU(inplace=True),
nn.Linear(100, 10),
)
nn_model.type(torch.FloatTensor)
# We will minimize cross-entropy between the ground truth and
# network predictions using an SGD optimizer
loss = nn.CrossEntropyLoss().type(torch.FloatTensor)
optimizer = optim.SGD(nn_model.parameters(), lr=1e-2, weight_decay=1e-1)
```
## Тренируем!
Ниже приведена функция `train_model`, реализующая основной цикл тренировки PyTorch.
Каждую эпоху эта функция вызывает функцию `compute_accuracy`, которая вычисляет точность на validation, эту последнюю функцию предлагается реализовать вам.
```
def compute_accuracy(model, loader):
"""
Computes accuracy on the dataset wrapped in a loader
Returns: accuracy as a float value between 0 and 1
"""
model.eval() # Evaluation mode
# TODO: Implement the inference of the model on all of the batches from loader,
# and compute the overall accuracy.
correct_samples = 0
total_samples = 0
for (x, y) in loader:
prediction = model(x)
indices = torch.argmax(prediction, 1)
correct_samples += torch.sum(indices == y)
total_samples += y.shape[0]
accuracy = float(correct_samples) / total_samples
return accuracy
# This is how to implement the same main train loop in PyTorch. Pretty easy, right?
def train_model(model, train_loader, val_loader, loss, optimizer, num_epochs):
loss_history = []
train_history = []
val_history = []
for epoch in range(num_epochs):
model.train() # Enter train mode
loss_accum = 0
correct_samples = 0
total_samples = 0
for i_step, (x, y) in enumerate(train_loader):
prediction = model(x)
loss_value = loss(prediction, y)
optimizer.zero_grad()
loss_value.backward()
optimizer.step()
indices = torch.argmax(prediction, 1)
correct_samples += torch.sum(indices == y)
total_samples += y.shape[0]
loss_accum += loss_value
ave_loss = loss_accum / (i_step + 1)
train_accuracy = float(correct_samples) / total_samples
val_accuracy = compute_accuracy(model, val_loader)
loss_history.append(float(ave_loss))
train_history.append(train_accuracy)
val_history.append(val_accuracy)
print("Average loss: %f, Train accuracy: %f, Val accuracy: %f" % (ave_loss, train_accuracy, val_accuracy))
return loss_history, train_history, val_history
loss_history, train_history, val_history = train_model(nn_model, train_loader, val_loader, loss, optimizer, 3)
```
## После основного цикла
Посмотрим на другие возможности и оптимизации, которые предоставляет PyTorch.
Добавьте еще один скрытый слой размера 100 нейронов к модели
```
# Since it's so easy to add layers, let's add some!
# TODO: Implement a model with 2 hidden layers of the size 100
nn_model = nn.Sequential(
Flattener(),
nn.Linear(3*32*32, 100),
nn.ReLU(inplace=True),
nn.Linear(100, 100),
nn.ReLU(inplace=True),
nn.Linear(100, 10),
)
nn_model.type(torch.FloatTensor)
optimizer = optim.SGD(nn_model.parameters(), lr=1e-2, weight_decay=1e-1)
loss_history, train_history, val_history = train_model(nn_model, train_loader, val_loader, loss, optimizer, 5)
```
Добавьте слой с Batch Normalization
```
# We heard batch normalization is powerful, let's use it!
# TODO: Add batch normalization after each of the hidden layers of the network, before or after non-linearity
# Hint: check out torch.nn.BatchNorm1d
nn_model = nn.Sequential(
Flattener(),
nn.Linear(3*32*32, 100),
nn.ReLU(inplace=True),
nn.BatchNorm1d(100),
nn.Linear(100, 100),
nn.ReLU(inplace=True),
nn.BatchNorm1d(100),
nn.Linear(100, 10),
)
optimizer = optim.SGD(nn_model.parameters(), lr=1e-3, weight_decay=1e-1)
loss_history, train_history, val_history = train_model(nn_model, train_loader, val_loader, loss, optimizer, 5)
```
Добавьте уменьшение скорости обучения по ходу тренировки.
```
def train_model(model, train_loader, val_loader, loss, optimizer, scheduler, num_epochs):
loss_history = []
train_history = []
val_history = []
for epoch in range(num_epochs):
scheduler.step()
model.train() # Enter train mode
loss_accum = 0
correct_samples = 0
total_samples = 0
for i_step, (x, y) in enumerate(train_loader):
prediction = model(x)
loss_value = loss(prediction, y)
optimizer.zero_grad()
loss_value.backward()
optimizer.step()
indices = torch.argmax(prediction, 1)
correct_samples += torch.sum(indices == y)
total_samples += y.shape[0]
loss_accum += loss_value
ave_loss = loss_accum / (i_step + 1)
train_accuracy = float(correct_samples) / total_samples
val_accuracy = compute_accuracy(model, val_loader)
loss_history.append(float(ave_loss))
train_history.append(train_accuracy)
val_history.append(val_accuracy)
print("Average loss: %f, Train accuracy: %f, Val accuracy: %f, LR: %f" % (ave_loss,
train_accuracy,
val_accuracy,
step_lr.get_lr()[0]))
return loss_history, train_history, val_history
# Learning rate annealing
# Reduce your learning rate 2x every 2 epochs
# Hint: look up learning rate schedulers in PyTorch. You might need to extend train_model function a little bit too!
nn_model = nn.Sequential(
Flattener(),
nn.Linear(3*32*32, 100),
nn.ReLU(inplace=True),
nn.BatchNorm1d(100),
nn.Linear(100, 100),
nn.ReLU(inplace=True),
nn.BatchNorm1d(100),
nn.Linear(100, 10),
)
optimizer = optim.SGD(nn_model.parameters(), lr=1e-3, weight_decay=1e-1)
step_lr = optim.lr_scheduler.StepLR(optimizer, step_size=2, gamma=0.5)
loss_history, train_history, val_history = train_model(nn_model, train_loader, val_loader, loss, optimizer, step_lr, 5)
```
# Визуализируем ошибки модели
Попробуем посмотреть, на каких изображениях наша модель ошибается.
Для этого мы получим все предсказания модели на validation set и сравним их с истинными метками (ground truth).
Первая часть - реализовать код на PyTorch, который вычисляет все предсказания модели на validation set.
Чтобы это сделать мы приводим код `SubsetSampler`, который просто проходит по всем заданным индексам последовательно и составляет из них батчи.
Реализуйте функцию `evaluate_model`, которая прогоняет модель через все сэмплы validation set и запоминает предсказания модели и истинные метки.
```
class SubsetSampler(Sampler):
r"""Samples elements with given indices sequentially
Arguments:
indices (ndarray): indices of the samples to take
"""
def __init__(self, indices):
self.indices = indices
def __iter__(self):
return (self.indices[i] for i in range(len(self.indices)))
def __len__(self):
return len(self.indices)
def evaluate_model(model, dataset, indices):
"""
Computes predictions and ground truth labels for the indices of the dataset
Returns:
predictions: np array of booleans of model predictions
grount_truth: np array of boolean of actual labels of the dataset
"""
model.eval() # Evaluation mode
# TODO: Evaluate model on the list of indices and capture predictions
# and ground truth labels
# Hint: SubsetSampler above could be useful!
sampler = SubsetSampler(indices)
loader = torch.utils.data.DataLoader(dataset, batch_size=64, sampler=sampler)
predictions = []
ground_truth = []
for (x, y) in loader:
prediction = model(x)
predictions.extend(torch.argmax(prediction, 1).numpy())
ground_truth.extend(y.numpy())
predictions = np.array(predictions)
ground_truth = np.array(ground_truth)
return predictions, ground_truth
# Evaluate model on validation
predictions, gt = evaluate_model(nn_model, data_train, val_indices)
assert len(predictions) == len(val_indices)
assert len(gt) == len(val_indices)
assert gt[100] == data_train[val_indices[100]][1]
assert np.any(np.not_equal(gt, predictions))
```
## Confusion matrix
Первая часть визуализации - вывести confusion matrix (https://en.wikipedia.org/wiki/Confusion_matrix ).
Confusion matrix - это матрица, где каждой строке соответствуют классы предсказанных, а столбцу - классы истинных меток (ground truth). Число с координатами `i,j` - это количество сэмплов класса `j`, которые модель считает классом `i`.

Для того, чтобы облегчить вам задачу, ниже реализована функция `visualize_confusion_matrix` которая визуализирует такую матрицу.
Вам осталось реализовать функцию `build_confusion_matrix`, которая ее вычислит.
Результатом должна быть матрица 10x10.
```
def visualize_confusion_matrix(confusion_matrix):
"""
Visualizes confusion matrix
confusion_matrix: np array of ints, x axis - predicted class, y axis - actual class
[i][j] should have the count of samples that were predicted to be class i,
but have j in the ground truth
"""
# Adapted from
# https://stackoverflow.com/questions/2897826/confusion-matrix-with-number-of-classified-misclassified-instances-on-it-python
assert confusion_matrix.shape[0] == confusion_matrix.shape[1]
size = confusion_matrix.shape[0]
fig = plt.figure(figsize=(10,10))
plt.title("Confusion matrix")
plt.ylabel("predicted")
plt.xlabel("ground truth")
res = plt.imshow(confusion_matrix, cmap='GnBu', interpolation='nearest')
cb = fig.colorbar(res)
plt.xticks(np.arange(size))
plt.yticks(np.arange(size))
for i, row in enumerate(confusion_matrix):
for j, count in enumerate(row):
plt.text(j, i, count, fontsize=14, horizontalalignment='center', verticalalignment='center')
def build_confusion_matrix(predictions, ground_truth):
"""
Builds confusion matrix from predictions and ground truth
predictions: np array of ints, model predictions for all validation samples
ground_truth: np array of ints, ground truth for all validation samples
Returns:
np array of ints, (10,10), counts of samples for predicted/ground_truth classes
"""
num_classes = len(np.unique(ground_truth))
confusion_matrix = np.zeros((num_classes,num_classes), np.int)
# TODO: Implement filling the prediction matrix
for i in range(num_classes):
for j in range(num_classes):
confusion_matrix[i, j] = np.sum(predictions[ground_truth == j] == i)
return confusion_matrix
confusion_matrix = build_confusion_matrix(predictions, gt)
visualize_confusion_matrix(confusion_matrix)
```
Наконец, посмотрим на изображения, соответствующие некоторым элементам этой матрицы.
Как и раньше, вам дана функция `visualize_images`, которой нужно воспрользоваться при реализации функции `visualize_predicted_actual`. Эта функция должна вывести несколько примеров, соответствующих заданному элементу матрицы.
Визуализируйте наиболее частые ошибки и попробуйте понять, почему модель их совершает.
```
def visualize_images(indices, data, title='', max_num=10):
"""
Visualizes several images from the dataset
indices: array of indices to visualize
data: torch Dataset with the images
title: string, title of the plot
max_num: int, max number of images to display
"""
to_show = min(len(indices), max_num)
fig = plt.figure(figsize=(10,1.5))
fig.suptitle(title)
for i, index in enumerate(indices[:to_show]):
plt.subplot(1,to_show, i+1)
plt.axis('off')
sample = data[index][0]
plt.imshow(sample)
def visualize_predicted_actual(predicted_class, gt_class, predictions, groud_truth, val_indices, data):
"""
Visualizes images of a ground truth class which were predicted as the other class
predicted_class: int 0-9, index of the predicted class
gt_class: int 0-9, index of the ground truth class
predictions: np array of ints, model predictions for all validation samples
ground_truth: np array of ints, ground truth for all validation samples
val_indices: np array of ints, indices of validation samples
"""
# TODO: Implement visualization using visualize_images above
# predictions and ground_truth are provided for validation set only, defined by val_indices
# Hint: numpy index arrays might be helpful
# https://docs.scipy.org/doc/numpy/user/basics.indexing.html#index-arrays
# Please make the title meaningful!
title = 'Visualization: predicted_class is {0}, gt_class is {1}'.format(predicted_class, gt_class)
indices_to_visualize = val_indices[(predictions == predicted_class) & (groud_truth == gt_class)]
visualize_images(indices_to_visualize, data, title)
data_train_images = dset.SVHN(PATH_TO_DATA, split='train')
visualize_predicted_actual(6, 8, predictions, gt, np.array(val_indices), data_train_images)
visualize_predicted_actual(1, 7, predictions, gt, np.array(val_indices), data_train_images)
```
# Переходим к свободным упражнениям!
Натренируйте модель как можно лучше - экспериментируйте сами!
Что следует обязательно попробовать:
- перебор гиперпараметров с помощью валидационной выборки
- другие оптимизаторы вместо SGD
- изменение количества слоев и их размеров
- наличие Batch Normalization
Но ограничиваться этим не стоит!
Точность на валидацонной выборке должна быть доведена до **60%**
За лучший результат в группе вы получите дополнительные баллы :)
```
def train_model(model, train_loader, val_loader, loss, optimizer, scheduler, num_epochs, verbose=True):
loss_history = []
train_history = []
val_history = []
for epoch in range(num_epochs):
if scheduler is not None:
scheduler.step()
model.train() # Enter train mode
loss_accum = 0
correct_samples = 0
total_samples = 0
for i_step, (x, y) in enumerate(train_loader):
prediction = model(x)
loss_value = loss(prediction, y)
optimizer.zero_grad()
loss_value.backward()
optimizer.step()
indices = torch.argmax(prediction, 1)
correct_samples += torch.sum(indices == y)
total_samples += y.shape[0]
loss_accum += loss_value
ave_loss = loss_accum / (i_step + 1)
train_accuracy = float(correct_samples) / total_samples
val_accuracy = compute_accuracy(model, val_loader)
loss_history.append(float(ave_loss))
train_history.append(train_accuracy)
val_history.append(val_accuracy)
if verbose:
print("Average loss: %f, Train accuracy: %f, Val accuracy: %f, LR: %f" % (ave_loss,
train_accuracy,
val_accuracy,
step_lr.get_lr()[0]))
else:
if epoch % 5 == 0:
print("Average loss: %f, Train accuracy: %f, Val accuracy: %f" % (ave_loss,
train_accuracy,
val_accuracy))
return loss_history, train_history, val_history
from random import choice
n_iterations = 10
best_params = {}
best_model = None
best_loss_history = None
best_train_history = None
best_val_history = None
for i in range(n_iterations):
print('Cycle: {0}'.format(i))
nn_model = nn.Sequential(
Flattener(),
nn.Linear(3*32*32, 100),
nn.ReLU(inplace=True),
nn.BatchNorm1d(100),
nn.Linear(100, 100),
nn.ReLU(inplace=True),
nn.BatchNorm1d(100),
nn.Linear(100, 10),
)
params = {
'lr': choice([1e-1, 1e-2, 1e-3]),
'wd': choice([1e-1, 1e-2, 1e-3]),
'momentum': choice([0.3, 0.5, 0.7, 0.9]),
'eps': choice([1e-8, 1e-6, 1e-4, 1e-2, 1e-1]),
}
optims = {
'sgd': optim.SGD(nn_model.parameters(), lr=params['lr'], weight_decay=params['wd']),
'adam': optim.Adam(nn_model.parameters(), lr=params['lr'], weight_decay=params['wd'])
}
optimizer = optims[choice(list(optims.keys()))]
schedulers = {
'step_lr': optim.lr_scheduler.StepLR(optimizer, step_size=2, gamma=0.5),
'none': None
}
scheduler = schedulers[choice(list(schedulers.keys()))]
loss_history, train_history, val_history = train_model(nn_model, train_loader, val_loader,
loss, optimizer, scheduler, n_iterations+1,
verbose=False)
if best_val_history is None or val_history[-1] > best_val_history[-1]:
best_params = params
best_model = nn_model
best_params['optimizer'] = optimizer
best_params['schedulers'] = scheduler
best_loss_history = loss_history
best_train_history = train_history
best_val_history = val_history
print("Best model")
print("Params:", best_params)
print()
print("Loss: %f, Train accuracy: %f, Val accuracy: %f" % (best_loss_history[-1],
best_train_history[-1],
best_val_history[-1]))
# Как всегда, в конце проверяем на test set
test_loader = torch.utils.data.DataLoader(data_test, batch_size=batch_size)
test_accuracy = compute_accuracy(nn_model, test_loader)
print("Test accuracy: %2.4f" % test_accuracy)
```
| github_jupyter |
# Magnet testing
```
import sys
import os
sys.path.append('../')
import pandas as pd
import src.io as sio
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
import numpy as np
MAGNET_FOLDER1 = sio.get_qudi_data_path("2021\\03\\20210319\\Magnet\\")
MAGNET_FOLDER2 = sio.get_qudi_data_path("2021\\03\\20210322\\Magnet\\")
MAGNET_FOLDER3 = sio.get_qudi_data_path("2021\\03\\20210323\\Magnet\\")
MAGNET_FOLDER4 = sio.get_qudi_data_path("2021\\03\\20210324\\Magnet\\")
%matplotlib inline
files = os.listdir(MAGNET_FOLDER1)
dat_files = []
for file in files:
filename, ext = os.path.splitext(file)
if ext == ".dat":
dat_files.append(filename + ext)
dat_files = [dat_value for idx, dat_value in enumerate(dat_files) if idx in [0, 2, 4, 5]]
fig, ax = plt.subplots(nrows=len(dat_files), sharex=True, sharey=True)
for idx, file in enumerate(dat_files):
filepath = os.path.join(MAGNET_FOLDER1, file)
df = pd.read_csv(filepath, skiprows=10, delimiter="\t", usecols=[0, 1, 2, 3], names=["Time", "current_x", "current_y", "current_z"])
ax[idx].plot(df["Time"], df[f"current_z"], "-")
if idx == 2:
title = f"[Run {idx+1}] Ramping up z-coil → No quench"
color = "tab:green"
elif idx == 3:
title = f"[Run {idx+1}] Max Current → Ramping down z-coil"
color = "tab:green"
ax[idx].set_xlabel("Time (s)")
else:
title = f"[Run {idx+1}] Ramping up z-coil → Quench"
color = "tab:red"
ax[idx].set_title(title, color=color)
max_current = max(df[f"current_z"])
ax[idx].axhline(max_current, linestyle="--", color=color, label="Max $I_z$" + f" = {max_current:.2f} A")
ax[idx].legend(loc="upper right")
ax[idx].set_ylabel("$I_z$ (A)")
fig.tight_layout()
plt.savefig("1.png", dpi=300)
%matplotlib inline
files = os.listdir(MAGNET_FOLDER3)
dat_files = []
for file in files:
filename, ext = os.path.splitext(file)
if ext == ".dat":
dat_files.append(filename + ext)
dat_files = [dat_value for idx, dat_value in enumerate(dat_files) if idx in [3, 4]]
fig, ax = plt.subplots(nrows=len(dat_files), sharex=True, sharey=False)
for idx, file in enumerate(dat_files):
filepath = os.path.join(MAGNET_FOLDER3, file)
df = pd.read_csv(filepath, skiprows=10, delimiter="\t", usecols=[0, 1, 2, 3], names=["Time", "current_x", "current_y", "current_z"])
for axis in ["x", "y", "z"]:
ax[idx].plot(df["Time"], df[f"current_{axis}"], "-", label=f"{axis}-coil")
if idx == 1:
title = f"[Run {idx+3}] Ramping up all coils → No quench"
color = "tab:green"
ax[idx].set_xlabel("Time (s)")
elif idx == 3:
title = f"[Run {idx+2}] Max Current → Ramping down"
color = "tab:green"
ax[idx].set_xlabel("Time (s)")
else:
title = f"[Run {idx+1}] Ramping up all coils → Quench"
color = "tab:red"
ax[idx].set_title(title, color=color)
# max_current = max(df[f"current_z"])
# ax[idx].axhline(max_current, linestyle="--", color=color, label="Max $I_z$" + f" = {max_current:.2f} A")
ax[idx].legend(loc="lower right")
ax[idx].set_ylabel("$I$ (A)")
fig.tight_layout()
plt.savefig("1.png", dpi=300)
def draw_ellipsoid(a, b, c):
coefs = (a, b, c) # Coefficients in a0/c x**2 + a1/c y**2 + a2/c z**2 = 1
# Radii corresponding to the coefficients:
rx, ry, rz = coefs
# Set of all spherical angles:
u = np.linspace(0, 2 * np.pi, 20)
v = np.linspace(0, np.pi, 20)
# Cartesian coordinates that correspond to the spherical angles:
# (this is the equation of an ellipsoid):
x = rx * np.outer(np.cos(u), np.sin(v))
y = ry * np.outer(np.sin(u), np.sin(v))
z = rz * np.outer(np.ones_like(u), np.cos(v))
return x, y, z
x, y, z = draw_ellipsoid(10, 10, 20)
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
ax.plot_wireframe(x, y, z)
ax.set_zlabel("$I_z$ (A)")
ax.set_xlim([-20, 20])
ax.set_xlabel("$I_x$ (A)")
ax.set_ylim([-20, 20])
ax.set_ylabel("$I_y$ (A)")
%matplotlib inline
x, y, z = draw_ellipsoid(10, 10, 20)
x1, y1, z1 = draw_ellipsoid(3, 3, 19)
x2, y2, z2 = draw_ellipsoid(9, 9, 5)
fig = plt.figure(figsize=(10, 10))
ax = fig.add_subplot(111, projection='3d')
ax.plot_wireframe(x, y, z, label="Measured", alpha=1)
#ax.plot_wireframe(x1, y1, z1, color="tab:orange", label="NV [111] tips", alpha=1)
ax.plot_wireframe(x2, y2, z2, color="tab:green", label="Meron sample", alpha=1)
ax.set_zlabel("$I_z$ (A)")
ax.set_xlim([-20, 20])
ax.set_xlabel("$I_x$ (A)")
ax.set_ylim([-20, 20])
ax.set_ylabel("$I_y$ (A)")
ax.legend()
#plt.savefig("Measured.png", dpi=300)
```
| github_jupyter |
This is an supervised classification example taken from the KDD 2009 cup. A copy of the data and details can be found here: [https://github.com/WinVector/PDSwR2/tree/master/KDD2009](https://github.com/WinVector/PDSwR2/tree/master/KDD2009). The problem was to predict account cancellation ("churn") from very messy data (column names not given, numeric and categorical variables, many missing values, some categorical variables with a large number of possible levels). In this example we show how to quickly use `vtreat` to prepare the data for modeling. `vtreat` takes in `Pandas` `DataFrame`s and returns both a treatment plan and a clean `Pandas` `DataFrame` ready for modeling.
Load our packages/modules.
```
import pandas
import xgboost
import vtreat
import vtreat.cross_plan
import numpy.random
import wvpy.util
import scipy.sparse
vtreat.__version__
```
Read in explanitory variables.
```
# data from https://github.com/WinVector/PDSwR2/tree/master/KDD2009
dir = "../../../PracticalDataScienceWithR2nd/PDSwR2/KDD2009/"
d = pandas.read_csv(dir + 'orange_small_train.data.gz', sep='\t', header=0)
vars = [c for c in d.columns]
d.shape
```
Read in dependent variable we are trying to predict.
```
churn = pandas.read_csv(dir + 'orange_small_train_churn.labels.txt', header=None)
churn.columns = ["churn"]
churn.shape
churn["churn"].value_counts()
```
Arrange test/train split.
```
numpy.random.seed(2020)
n = d.shape[0]
# https://github.com/WinVector/pyvtreat/blob/master/Examples/CustomizedCrossPlan/CustomizedCrossPlan.md
split1 = vtreat.cross_plan.KWayCrossPlanYStratified().split_plan(n_rows=n, k_folds=10, y=churn.iloc[:, 0])
train_idx = set(split1[0]['train'])
is_train = [i in train_idx for i in range(n)]
is_test = numpy.logical_not(is_train)
```
(The reported performance runs of this example were sensitive to the prevalance of the churn variable in the test set, we are cutting down on this source of evaluation variarance by using the stratified split.)
```
d_train = d.loc[is_train, :].copy()
churn_train = numpy.asarray(churn.loc[is_train, :]["churn"]==1)
d_test = d.loc[is_test, :].copy()
churn_test = numpy.asarray(churn.loc[is_test, :]["churn"]==1)
```
Take a look at the dependent variables. They are a mess, many missing values. Categorical variables that can not be directly used without some re-encoding.
```
d_train.head()
d_train.shape
```
Try building a model directly off this data (this will fail).
```
fitter = xgboost.XGBClassifier(n_estimators=10, max_depth=3, objective='binary:logistic')
try:
fitter.fit(d_train, churn_train)
except Exception as ex:
print(ex)
```
Let's quickly prepare a data frame with none of these issues.
We start by building our treatment plan, this has the `sklearn.pipeline.Pipeline` interfaces.
```
plan = vtreat.BinomialOutcomeTreatment(
outcome_target=True,
params=vtreat.vtreat_parameters({'filter_to_recommended':True}))
```
Use `.fit_transform()` to get a special copy of the treated training data that has cross-validated mitigations againsst nested model bias. We call this a "cross frame." `.fit_transform()` is deliberately a different `DataFrame` than what would be returned by `.fit().transform()` (the `.fit().transform()` would damage the modeling effort due nested model bias, the `.fit_transform()` "cross frame" uses cross-validation techniques similar to "stacking" to mitigate these issues).
```
cross_frame = plan.fit_transform(d_train, churn_train)
```
Take a look at the new data. This frame is guaranteed to be all numeric with no missing values, with the rows in the same order as the training data.
```
cross_frame.head()
cross_frame.shape
```
Pick a recommended subset of the new derived variables.
```
plan.score_frame_.head()
model_vars = numpy.asarray(plan.score_frame_["variable"][plan.score_frame_["recommended"]])
len(model_vars)
```
Fit the model
```
cross_frame.dtypes
# fails due to sparse columns
# can also work around this by setting the vtreat parameter 'sparse_indicators' to False
try:
cross_sparse = xgboost.DMatrix(data=cross_frame.loc[:, model_vars], label=churn_train)
except Exception as ex:
print(ex)
# also fails
try:
cross_sparse = scipy.sparse.csc_matrix(cross_frame[model_vars])
except Exception as ex:
print(ex)
# works
cross_sparse = scipy.sparse.hstack([scipy.sparse.csc_matrix(cross_frame[[vi]]) for vi in model_vars])
# https://xgboost.readthedocs.io/en/latest/python/python_intro.html
fd = xgboost.DMatrix(
data=cross_sparse,
label=churn_train)
x_parameters = {"max_depth":3, "objective":'binary:logistic'}
cv = xgboost.cv(x_parameters, fd, num_boost_round=100, verbose_eval=False)
cv.head()
best = cv.loc[cv["test-error-mean"]<= min(cv["test-error-mean"] + 1.0e-9), :]
best
ntree = best.index.values[0]
ntree
fitter = xgboost.XGBClassifier(n_estimators=ntree, max_depth=3, objective='binary:logistic')
fitter
model = fitter.fit(cross_sparse, churn_train)
```
Apply the data transform to our held-out data.
```
test_processed = plan.transform(d_test)
```
Plot the quality of the model on training data (a biased measure of performance).
```
pf_train = pandas.DataFrame({"churn":churn_train})
pf_train["pred"] = model.predict_proba(cross_sparse)[:, 1]
wvpy.util.plot_roc(pf_train["pred"], pf_train["churn"], title="Model on Train")
```
Plot the quality of the model score on the held-out data. This AUC is not great, but in the ballpark of the original contest winners.
```
test_sparse = scipy.sparse.hstack([scipy.sparse.csc_matrix(test_processed[[vi]]) for vi in model_vars])
pf = pandas.DataFrame({"churn":churn_test})
pf["pred"] = model.predict_proba(test_sparse)[:, 1]
wvpy.util.plot_roc(pf["pred"], pf["churn"], title="Model on Test")
```
Notice we dealt with many problem columns at once, and in a statistically sound manner. More on the `vtreat` package for Python can be found here: [https://github.com/WinVector/pyvtreat](https://github.com/WinVector/pyvtreat). Details on the `R` version can be found here: [https://github.com/WinVector/vtreat](https://github.com/WinVector/vtreat).
We can compare this to the [R solution (link)](https://github.com/WinVector/PDSwR2/blob/master/KDD2009/KDD2009vtreat.md).
We can compare the above cross-frame solution to a naive "design transform and model on the same data set" solution as we show below. Note we are leaveing filter to recommended on, to show the non-cross validated methodology still fails in an "easy case."
```
plan_naive = vtreat.BinomialOutcomeTreatment(
outcome_target=True,
params=vtreat.vtreat_parameters({'filter_to_recommended':True}))
plan_naive.fit(d_train, churn_train)
naive_frame = plan_naive.transform(d_train)
model_vars = numpy.asarray(plan_naive.score_frame_["variable"][plan_naive.score_frame_["recommended"]])
len(model_vars)
naive_sparse = scipy.sparse.hstack([scipy.sparse.csc_matrix(naive_frame[[vi]]) for vi in model_vars])
fd_naive = xgboost.DMatrix(data=naive_sparse, label=churn_train)
x_parameters = {"max_depth":3, "objective":'binary:logistic'}
cvn = xgboost.cv(x_parameters, fd_naive, num_boost_round=100, verbose_eval=False)
bestn = cvn.loc[cvn["test-error-mean"] <= min(cvn["test-error-mean"] + 1.0e-9), :]
bestn
ntreen = bestn.index.values[0]
ntreen
fittern = xgboost.XGBClassifier(n_estimators=ntreen, max_depth=3, objective='binary:logistic')
fittern
modeln = fittern.fit(naive_sparse, churn_train)
test_processedn = plan_naive.transform(d_test)
test_processedn = scipy.sparse.hstack([scipy.sparse.csc_matrix(test_processedn[[vi]]) for vi in model_vars])
pfn_train = pandas.DataFrame({"churn":churn_train})
pfn_train["pred_naive"] = modeln.predict_proba(naive_sparse)[:, 1]
wvpy.util.plot_roc(pfn_train["pred_naive"], pfn_train["churn"], title="Overfit Model on Train")
pfn = pandas.DataFrame({"churn":churn_test})
pfn["pred_naive"] = modeln.predict_proba(test_processedn)[:, 1]
wvpy.util.plot_roc(pfn["pred_naive"], pfn["churn"], title="Overfit Model on Test")
```
Note the naive test performance is worse, despite its far better training performance. This is over-fit due to the nested model bias of using the same data to build the treatment plan and model without any cross-frame mitigations.
| github_jupyter |
# Memory Management in NumPy
ُhe very core of NumPy is the ndarray object. We are going to finish the last important attribute of ndarray: strides, which will give you the full picture of memory layout. Also, it's time to show you that NumPy arrays can deal not only with numbers but also with various types of data; we will talk about record arrays and date time arrays. Lastly, we will show how to read/write NumPy arrays from/to files, and start to do some real-world analysis using NumPy.
## Internal memory layout of an ndarray
An instance of class `ndarray` consists of a contiguous one-dimensional segment of computer memory (owned by the array, or by some other object), combined with an indexing scheme that maps `N` integers into the location of an item in the block. The ranges in which the indices can vary is specified by the **shape** of the array. How many bytes each item takes and how the bytes are interpreted is defined by the data-type object associated with the array.
A particularly interesting attribute of the ndarray object is `flags`. Type the following code:
```
import numpy as np
x = np.array([1, 2, 3])
x.flags
```
The `flags` attribute holds information about the memory layout of the array. The `C_CONTIGUOUS` field in the output indicates whether the array was a C-style array. This means that the indexing of this array is done like a C array. This is also called row-major indexing in the case of 2D arrays. This means that, when moving through the array, the row index is incremented first, and then the column index is incremented. In the case of a multidimensional C-style array, the last dimension is incremented first, followed by the last but one, and so on.
Similarly, the `F_CONTIGUOUS` attribute indicates whether the array is a Fortran-style array. Such an array is said to have column-major indexing (R, Julia, and MATLAB use column-major arrays). This means that, when moving through the array, the first index (along the column) is incremented first.
```
c_array = np.random.rand(10000, 10000)
f_array = np.asfortranarray(c_array)
def sum_row(x):
'''
Given an array `x`, return the sum of its zeroth row.
'''
return np.sum(x[0, :])
def sum_col(x):
'''
Given an array `x`, return the sum of its zeroth column.
'''
return np.sum(x[:, 0])
```
Now, let's test the performance of the two functions on both the arrays using IPython's %timeit magic function:
```
%timeit sum_row(c_array)
%timeit sum_row(f_array)
%timeit sum_col(c_array)
%timeit sum_col(f_array)
```
As we can see, summing up the row of a C array is much faster than summing up its column. This is because, in a C array, elements in a row are laid out in successive memory locations. The opposite is true for a Fortran array, where the elements of a column are laid out in consecutive memory locations.
This is an important distinction and allows you to suitably arrange your data in an array, depending on the kind of algorithm or operation you are performing. Knowing this distinction can help you speed up your code by orders of magnitude.
## Views and Copies
There are primarily two ways of accessing data by slicing and indexing. They are called copies and views: you can either access elements directly from an array, or create a copy of the array that contains only the accessed elements. Since a view is a reference of the original array (in Python, all variables are references), modifying a view modifies the original array too. This is not true for copies.
The `may_share_memory` function in NumPy miscellaneous routines can be used to determine whether two arrays are copies or views of each other. While this method does the job in most cases, it is not always reliable, since it uses heuristics. It may return incorrect results too. For introductory purposes, however, we shall take it for granted.
Generally, slicing an array creates a view and indexing it creates a copy. Let us study these differences through a few code snippets. First, let's create a random `100x10` array.
```
x = np.random.rand(100, 10)
```
Now, let us extract the first five rows of the array and assign them to variable `y`.
```
y = x[:5, :]
```
Let us see if `y` is a view of `x`.
```
np.may_share_memory(x, y)
```
Now let us modify the array `y` and see how it affects `x`. Set all the elements of `y` to zero:
```
y[:] = 0
x[:5, :]
```
The code snippet prints out five rows of zeros. This is because `y` was just a view, a reference to `x`.
Next, let's create a copy to see the difference. We use the preceding method that uses a random function to create the `x` array, but this time we initialize the `y` array using `numpy.empty` to create an empty array first and then copy the values from `x` to `y`. So, now `y` is not a view/reference of `x` anymore; it's an independent array but has the same values as part of `x`. Let's use the may_share_memory function again to verify that `y` is the copy of `x`:
```
x = np.random.rand(100, 10)
y = np.empty([5, 10])
y[:] = x[:5, :]
np.may_share_memory(x, y)
```
Let's alter the value in `y` and check whether the value of `x` changes as well:
```
y[:] = 0
x[:5, :]
```
You should see the preceding snippet print out five rows of random numbers as we initialized `x`, so changing `y` to `0` didn't affect `x`.
## Introducing strides
Strides are the indexing scheme in NumPy arrays, and indicate the number of bytes to jump to find the next element. We all know the performance improvements of NumPy come from a homogeneous multidimensional array object with fixed-size items, the `numpy.ndarray` object. We've talked about the `shape` (dimension) of the `ndarray` object, the data type, and the order (the C-style row-major indexing arrays and the Fortran style column-major arrays.) Now it's time to take a closer look at **strides**.
### Example
Let's start by creating a NumPy array and changing its shape to see the differences in the strides.
```
x = np.arange(8, dtype = np.int8)
x
x.strides
x.data
```
A one-dimensional array `x` is created and its data type is NumPy integer 8, which means each element in the array is an 8-bit integer (1 byte each, and a total of 8 bytes). The strides represent the `tuple` of bytes to step in each dimension when traversing an array. In the previous example, it's one dimension, so we obtain the tuple as `(1, )`. Each element is 1 byte apart from its previous element. When we print out `x.data`, we can get the Python buffer object pointing to the start of the data
Change the shape and see the stride change:
```
x.shape = 2, 4
x.strides
```
Now we change the dimensions of `x` to `2x4` and check the strides again. We can see it changes to `(4, 1)`, which means the elements in the first dimension are four bytes apart, and the array need to jump four bytes to find the next row, but the elements in the second dimension are still 1 byte apart, jumping one byte to find the next column. Let's print out `x.data` again, and we can see that the memory layout of the data remains the same, but only the strides change. The same behavior occurs when we change the shape to be three dimensional: `1x4x2` arrays. (What if our arrays are constructed by the Fortran style order? How will the strides change due to changing the shapes? Try to create a column-major array and do the same exercise to check this out.)
```
x.shape = 1, 4, 2
x.strides
x.data
```
### How can the stride improve our NumPy experience?
So now we know what a stride is, and its relationship to an ndarray object, but how can the stride improve our NumPy experience? Let's do some stride manipulation to get a better sense of this: two arrays are with same content but have different strides:
```
x = np.ones((10000, ))
y = np.ones((10000 * 100, ))[::100]
x.shape, y.shape
(x == y).all()
```
We create two NumPy Arrays, `x` and `y`, and do a comparison; we can see that the two arrays are equal. They have the same shape and all the elements are one, but actually the two arrays are different in terms of memory layout. Let's simply use the flags attribute to check the two arrays' memory layout.
```
x.flags
y.flags
```
We can see that the `x` array is continuous in both the C and Fortran order while `y` is not. Let's check the strides for the difference:
```
x.strides, y.strides
```
Array `x` is created continuously, so in the same dimension each element is eight bytes apart (the default `dtype` of `numpy.ones` is a 64-bit float); however, `y` is created from a subset of 10000 * 100 for every 100 elements, so the index schema in the memory layout is not continuous.
Even though `x` and `y` have the same shape, each element in `y` is 800 bytes apart from each other. When you use NumPy arrays `x` and `y`, you might not notice the difference in indexing, but the memory layout does affect the performance. Let's use the `%timeit` to check this out:
```
%timeit x.sum()
%timeit y.sum()
```
Typically with a fixed cache size, when the stride size gets larger, the hit rate (the fraction of memory accessed that finds data in the cache) will be lower, comparatively, while the miss rate (the fraction of memory accessed that has to go to the memory) will be higher. The cache hit time and miss time compose the average data access time. Let's try to look at our example again from the cache perspective. Array `x` with smaller strides is faster than the larger strides of `y`. The reason for the difference in performance is that the CPU is pulling data from the main memory to its cache in blocks, and the smaller stride means fewer transfers are needed. See the following graph for details, where the red line represents the size of the CPU cache, and blue boxes represent the memory layout containing the data.
It's obvious that if `x` and `y` are both required, 100 blue boxes of data, the required cache time for `x` will be less.
<img src="../images/cpu-cache.jpg" alt="cpu-cache" width=500 align="left" />
### Stride in N-dimensional `ndarray`
A segment of memory is inherently 1-dimensional, and there are many different schemes for arranging the items of an N-dimensional array in a 1-dimensional block. NumPy is flexible, and ndarray objects can accommodate any strided indexing scheme. In a strided scheme, the N-dimensional index (`n_0, n_1, ..., n_{N-1}`) corresponds to the offset (in bytes):
$n_{\text{offset}} = \sum_{k=0}^{N-1} s_k n_k$
from the beginning of the memory block associated with the array. Here, $s_k$ are integers which specify the strides of the array.
```
import numpy as np
x = np.random.rand(3, 4)
x.itemsize, x.nbytes / x.shape[0]
x.strides
# .flatten() returns numpy array
x.flatten()
# .flat returns iterator
np.array(x.flat)
# a stride of 1 is equal to 32 bytes
x[1]
```
## Conclusion: Why `np.array` not `list`
```
x = [1, '2', [1, 2, 3]]
arr = np.array([1, 2, 3])
```
list: more memory is needed
homogenous memory in numpy results in more speed -> different data types
c_array f_array ----> which operations are slow which are fast? which axes?
default is memory view in numpy -----> no copy
debug ---> flags, strides, data, (under the hood information)
vectorization --> beutiy, no foor loop, formula == code
broadcasting --> no broadcasting in python
| github_jupyter |
```
#Importing some necessary packages
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import xgboost as xgb
#Reading the training and testing files
train = pd.read_csv('train.csv')
test = pd.read_csv('test.csv')
train.head()
test.head()
#Knowing the shape of train and test dataset
print('Shape of train is',train.shape,
'\nShape of test is',test.shape)
# list of features
features = list(train.columns)
features
```
## Preprocessing and EDA on training dataset
```
#checking if there is any null value is present or not
train.isnull().sum()
```
**Half of the instances in column Outcomesubtype has null values.**
**18 & 1 null values present in sexuponoutcome nd ageuponoutcome respectively won't affect much, so we discard those rows directly.**
```
#considering only those rows which don't have null values.
train = train[pd.notnull(train['AgeuponOutcome'])]
train = train[pd.notnull(train['SexuponOutcome'])]
#filling null values with unknown category
train.OutcomeSubtype = train.OutcomeSubtype.fillna('unknown')
```
### Extracting days and years from AgeOutcome and sex from SexuponOutcome
```
import re
def convert_to_days(row):
if (row['AgeuponOutcome'] == '1 year'):
age = 365
return age
elif (row['AgeuponOutcome'] == '1 month'):
age = 30
return age
elif (row['AgeuponOutcome'] == '1 week'):
age = 7
return age
else:
split = row['AgeuponOutcome'].split(" ")
period = split[1]
if (re.search('years',period)):
per_mod = 365
elif (re.search('months',period)):
per_mod = 30
elif (re.search('weeks', period)):
per_mod = 7
else:
per_mod = 1
age = int(split[0]) * per_mod
return age
def label_age(row):
if (row['AgeuponOutcome'] == '1 year'):
yr = 1
return yr
elif (row['AgeuponOutcome'] == '1 month'):
yr = 1/12
return yr
elif (row['AgeuponOutcome'] == '1 week'):
yr = 1/48
return yr
else:
split = row['AgeuponOutcome'].split(" ")
period = split[1]
if (re.search('years',period)):
per_mod = 1
elif (re.search('months',period)):
per_mod = 1/12
elif (re.search('weeks', period)):
per_mod = 1/48
else:
per_mod = 1
yr = int(split[0]) * per_mod
return yr
def label_sex(row):
if row['SexuponOutcome'] == 'Unknown':
sex = 'Unknown'
else:
split = row['SexuponOutcome'].split(" ")
sex = split[1]
return sex
#Applying both on training dataset
train["Agecat_days"]=train.apply (lambda row: convert_to_days (row),axis=1)
train["Agecat_years"]=train.apply (lambda row: label_age (row),axis=1)
train['sex'] = train.apply( lambda row: label_sex(row) , axis=1)
```
### Making a new year feature
```
#Handling datetime column
train.DateTime = pd.to_datetime(train.DateTime)
train['year'] = train.DateTime.dt.year
train['month'] = train.DateTime.dt.month
train['weekday'] = train.DateTime.dt.weekday
train['hour'] = train.DateTime.dt.hour
train.head()
```
### How differenct outcomes are distributed
```
sns.set(rc = {'figure.figsize' : (7,6)})
train.OutcomeType.value_counts().plot(kind = 'bar')
```
**Observation : Dataset is biased towards the outcome type Adoption and Transfer**
```
train.groupby('OutcomeType').AnimalType.value_counts().unstack().plot(kind = 'bar')
```
**Observation: Adoption and trnasfer happens more than others ( good for animal).**
```
train.groupby('OutcomeType').year.value_counts().unstack().plot(kind = 'bar')
```
### Relationship between hour of the day and Outcometype
```
sns.set(rc = {'figure.figsize' : (15,10)})
sns.countplot(data = train,x = 'hour', hue = 'OutcomeType')
```
**Observation : Arounf 5-6 pm, it is more likely for animal to be adopted whic around 2-3 pm it is more likely to be transfered.**
```
sns.set(rc = {'figure.figsize' : (7,6)})
sns.countplot(data = train,x = 'weekday', hue = 'OutcomeType')
```
**Observation : Adoption is more likely to happen in weekends**
### Agewise OutcomeType
```
sns.countplot(data = train[train.Agecat_years <= 4.0], x='Agecat_years',hue= 'OutcomeType')
```
**Observation : Young dogs (older than a month) are likely adopted.**
```
sns.countplot(data = train[train.Agecat_years >= 10.0], x='Agecat_years',hue= 'OutcomeType')
```
**Old dogs are likely returned to their owner.**
```
sns.countplot(x = train.AnimalType, hue=train.OutcomeType)
```
**Dogs in general are more likely to be returned to their owner than cats.**
```
sns.countplot(x = train.SexuponOutcome, hue = train.OutcomeType)
```
**Neutered males and spayed females are much more likely to get adopted.**
# Preparing training and test data for Classifier
```
# Prepare for training data
Xtrain = train.drop(["OutcomeType","OutcomeSubtype","AnimalID","Name",'DateTime','Breed','sex'],axis=1)
ytrain = train["OutcomeType"].astype('category').cat.codes
Xtrain.AnimalType = Xtrain.AnimalType.astype('category').cat.codes
Xtrain.SexuponOutcome = Xtrain.SexuponOutcome.astype('category').cat.codes
Xtrain.AgeuponOutcome = Xtrain.AgeuponOutcome.astype('category').cat.codes
Xtrain.Color = Xtrain.Color.astype('category').cat.codes
#Xtrain.sex = Xtrain.sex.astype('category').cat.codes
Xtrain.head()
#preparing testing data
test.AgeuponOutcome = test.AgeuponOutcome.fillna('1 year')
test["Agecat_days"]=test.apply (lambda row: convert_to_days (row),axis=1)
test["Agecat_years"]=test.apply (lambda row: label_age (row),axis=1)
test['sex'] = test.apply( lambda row: label_sex(row) , axis=1)
test.DateTime = pd.to_datetime(test.DateTime)
test['year'] = test.DateTime.dt.year
test['month'] = test.DateTime.dt.month
test['weekday'] = test.DateTime.dt.weekday
test['hour'] = test.DateTime.dt.hour
Xtest = test.drop(['ID','Name','DateTime','Breed','sex'],axis = 1)
Xtest.AnimalType = Xtest.AnimalType.astype('category').cat.codes
Xtest.SexuponOutcome = Xtest.SexuponOutcome.astype('category').cat.codes
Xtest.AgeuponOutcome = Xtest.AgeuponOutcome.astype('category').cat.codes
Xtest.Color = Xtest.Color.astype('category').cat.codes
#Xtest.sex = Xtest.sex.astype('category').cat.codes
Xtest.head()
from sklearn.preprocessing import StandardScaler
X_train_std = StandardScaler().fit_transform(Xtrain)
X_test_std = StandardScaler().fit_transform(Xtest)
params = {"objective": "multi:softprob",
"num_class": 5,
"eta": 0.1,
"max_depth": 5,
"min_child_weight": 3,
"silent": 1,
"subsample": 0.7,
"colsample_bytree": 0.7,
"seed": 1}
num_trees=250
%time gbm = xgb.train(params, xgb.DMatrix(X_train_std, ytrain), num_trees)
xgb.plot_importance(gbm)
test_probs_xgb = gbm.predict(xgb.DMatrix(X_test_std))
test_probs_xgb
submit = pd.DataFrame({
'ID' : test['ID'],
'Adoption' : test_probs_xgb[:,0],
'Died' : test_probs_xgb[:,1],
'Euthanasia' : test_probs_xgb[:,2],
'Return_to_owner' : test_probs_xgb[:,3],
'Transfer' : test_probs_xgb[:,4]
})
submit.to_csv('shelter.csv', index = False)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/nikshanpatel/CS634-DataMiningProject-1/blob/master/Project1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# **PROJECT**: **[Google Analytics Customer Revenue Prediction](https://www.kaggle.com/c/ga-customer-revenue-prediction/overview)**
#### **The fundamental steps we will be taking in this project are:**
>1. **Understand the Objective of this project**
2. **Data Collection & Data Preperation**
3. **Modeling & Evaluation**
4. **Communicate/Present Results**
# **1 - Understanding the Objective**
---
### **Predict how much GStore customers will spend**
In this [Project](https://www.kaggle.com/c/ga-customer-revenue-prediction/overview), we’re challenged to analyze a Google Merchandise Store (also known as GStore, where Google swag is sold) customer dataset to **predict revenue per customer**.
**What are we predicting?**
We are predicting the natural log of the sum of all transactions per user.
**Evaluation Metric**
***`RMSE`*** - *Root Mean Squared Error*
**RMSE** is the standard deviation of residuals (predictions errors). When we scatter plot the target_test values and predictions, the RMSE tells us how concentrated are those points around the line of best fit.

# **2 - Data Collection & Data Preperation**
---
## **Mount** the **Google** **Drive** on to the Notebook.
---
```
from google.colab import drive
drive.mount('/content/drive')
# to load kaggle API
!pip install -q kaggle
from google.colab import files
uploaded = files.upload()
!mkdir -p ~/.kaggle
!cp kaggle.json ~/.kaggle/
#now we install 7zip
!apt-get install p7zip-full
!kaggle datasets download -d niteshmistry/minigacustomerrevenueprediction
!7za e minigacustomerrevenueprediction.zip
```
## **Import** the necessary **libraries** that are required to run this notebook:
---
>1. **JSON**
2. **Pandas**
3. **NumPy**
4. **MatPlotLib**
5. **Plotly** - Plot graphs
6. **SKLearn** - For model selection, preprocessing, metrics
```
import os
import time
import json
import numpy as np
import pandas as pd
from pandas.io.json import json_normalize
import matplotlib.pyplot as plt
from matplotlib.ticker import StrMethodFormatter
from sklearn import linear_model
from sklearn.model_selection import train_test_split
%matplotlib inline
```
## **Datasets** that are required for this project that are necessary to run the notebook :
---
>1. **TRAIN Dataset** - *`train.csv`* (315 MB)
2. **TEST Dataset** - *`test.csv`* (32 MB)
### Each row in the dataset is one visit to the store. We are predicting the natural log of the sum of all transactions per user.
## The **data fields** in the given files are
* **fullVisitorId**- A unique identifier for each user of the Google Merchandise Store.
* **channelGrouping** - The channel via which the user came to the Store.
* **date** - The date on which the user visited the Store.
* **device** - The specifications for the device used to access the Store.
* **geoNetwork** - This section contains information about the geography of the user.
* **sessionId** - A unique identifier for this visit to the store.
* **socialEngagementType** - Engagement type, either "Socially Engaged" or "Not Socially Engaged".
* **totals** - This section contains aggregate values across the session.
* **trafficSource** - This section contains information about the Traffic Source from which the session originated.
* **visitId** - An identifier for this session. This is part of the value usually stored as the _utmb cookie. This is only unique to the user. For a completely unique ID, you should use a combination of fullVisitorId and visitId.
* **visitNumber** - The session number for this user. If this is the first session, then this is set to 1.
* **visitStartTime** - The timestamp (expressed as POSIX time).
***NOTE***: Some of the fields are in json format. We will have to transform them into Dataframe
## **LOAD** the data files that are required:
---
```
TRAIN_FILE = '/content/mini-train.csv'
TEST_FILE = '/content/mini-test.csv'
#------------------------------------------------------------------------------------------------
# COMMON CSV DATA FILE LOADER - RETURN DATAFRAME
#------------------------------------------------------------------------------------------------
def convertCSVtoDF(csv_path, nrows):
df = pd.read_csv(csv_path, dtype={"fullVisitorId": "str"}, nrows=nrows)
print(f"LOADED: {os.path.basename(csv_path)} SHAPE: {df.shape}")
return df
#------------------------------------------------------------------------------------------------
# LOAD TRAINING DATASET file into DATAFRAME
#------------------------------------------------------------------------------------------------
train_df = convertCSVtoDF(TRAIN_FILE,None)
test_df = convertCSVtoDF(TEST_FILE,None)
#------------------------------------------------------------------------------------------------
# LISTING HEADERS AND SAMPLE RECORDS (5)
#------------------------------------------------------------------------------------------------
print("\n")
print(train_df.columns)
#------------------------------------------------------------------------------------------------
# FUNCTION REMOVING EMPTY COLUMNS with value "not available in demo dataset"
#------------------------------------------------------------------------------------------------
def removeUselessColumns(df):
EMPTY_COLUMNS = []
dataframe = df
for column in train_df.columns:
if train_df[column][0] == 'not available in demo dataset':
EMPTY_COLUMNS.append(column)
print("DATAFRAME IS " + str(len(dataframe.columns)) + " COLUMNS...")
print("FOUND " + str(len(EMPTY_COLUMNS)) + " USELESS COLUMNS...")
print("COLUMNS: " + str(EMPTY_COLUMNS))
print("REMOVING " + str(len(EMPTY_COLUMNS)) + " COLUMNS...")
dataframe.drop(EMPTY_COLUMNS, axis=1, inplace=True)
print("DATAFRAME IS NOW " + str(len(dataframe.columns)) + " COLUMNS...")
return dataframe
```
**REMOVE USELESS COLUMNS** that has values "**NOT AVAILABLE IN DEMO DATASET**
```
#------------------------------------------------------------------------------------------------
# TRAINING DATAFRAME - REMOVING EMPTY COLUMNS with value "not available in demo dataset"
#------------------------------------------------------------------------------------------------
train_df = removeUselessColumns(train_df)
train_df.head(5)
train_df.describe()
train_df['date'].head()
#------------------------------------------------------------------------------------------------
# TESTING DATAFRAME - REMOVING EMPTY COLUMNS with value "not available in demo dataset"
#------------------------------------------------------------------------------------------------
test_df = removeUselessColumns(test_df)
test_df.head(5)
test_df.describe()
```
**Impute 0 for ALL missing target values (totals.transactionRevenue)**
## Check Missing Data
```
# checking missing data
miss_per = {}
for k, v in dict(train_df.isna().sum(axis=0)).items():
if v == 0:
continue
miss_per[k] = 100 * float(v) / len(train_df)
miss_per
#train_df["totals.transactionRevenue"].fillna(0, inplace=True)
#test_df["totals.transactionRevenue"].fillna(0, inplace=True)
```
**Remove columns that are in TRAINING which are not there in TEST**
```
print("Columns not in TEST but in TRAIN : ", set(train_df.columns).difference(set(test_df.columns)))
```
# **3 - MODELING & EVALUTATION**
---
Since we are predicting the natural log of sum of all transactions of the user, let us sum up the transaction revenue at user level and take a log and then do a scatter plot.
First we have to change the datatype to 'FLOAT' to make sure the value is a dollar value
```
train_df["totals.transactionRevenue"] = train_df["totals.transactionRevenue"].astype('float')
```
Next we have to group the USER level transaction to by using the GROUPBY function on the dataframe. This will give us the Total Transaction Value for given user.
```
grouped_df_sum = train_df.groupby("fullVisitorId")["totals.transactionRevenue"].sum().reset_index()
grouped_df_sum.sort_values(by=['totals.transactionRevenue'], inplace=True, ascending=False)
grouped_df_sum
ax = grouped_df_sum.hist('totals.transactionRevenue', bins=25, grid=False, figsize=(12,8), color='#86bf91', zorder=2, rwidth=0.9)
ax = ax[0]
for x in ax:
# Despine
x.spines['right'].set_visible(False)
x.spines['top'].set_visible(False)
x.spines['left'].set_visible(False)
# Switch off ticks
x.tick_params(axis="both", which="both", bottom="off", top="off", labelbottom="on", left="off", right="off", labelleft="on")
# Draw horizontal axis lines
vals = x.get_yticks()
for tick in vals:
x.axhline(y=tick, linestyle='dashed', alpha=0.4, color='#eeeeee', zorder=1)
# Remove title
x.set_title("")
# Set x-axis label
x.set_xlabel("Total Transaction Revenue (per User)", labelpad=20, weight='bold', size=12)
# Set y-axis label
x.set_ylabel("Visitors", labelpad=20, weight='bold', size=14)
grouped_df_count = train_df.groupby("fullVisitorId")["totals.transactionRevenue"].count().reset_index()
grouped_df_count.sort_values(by=['totals.transactionRevenue'], inplace=True, ascending=False)
grouped_df_count
ax = grouped_df_count.hist('totals.transactionRevenue', bins=25, grid=False, figsize=(12,8), color='#86bf91', zorder=2, rwidth=0.9)
ax = ax[0]
for x in ax:
# Despine
x.spines['right'].set_visible(False)
x.spines['top'].set_visible(False)
x.spines['left'].set_visible(False)
# Switch off ticks
x.tick_params(axis="both", which="both", bottom="off", top="off", labelbottom="on", left="off", right="off", labelleft="on")
# Draw horizontal axis lines
vals = x.get_yticks()
for tick in vals:
x.axhline(y=tick, linestyle='dashed', alpha=0.4, color='#eeeeee', zorder=1)
# Remove title
x.set_title("")
# Set x-axis label
x.set_xlabel("Total Transaction Count (per User)", labelpad=20, weight='bold', size=12)
# Set y-axis label
x.set_ylabel("Visitors", labelpad=20, weight='bold', size=14)
```
## **numpy.log1p() in Python**
**numpy.log1p(arr, out = None, *, where = True, casting = ‘same_kind’, order = ‘K’, dtype = None, ufunc ‘log1p’) :**
This mathematical function helps user to calculate **natural logarithmic value of x+1** where x belongs to all the input array elements.
* **log1p is reverse of exp(x) – 1.**
```
plt.figure(figsize=(15,10))
plt.scatter(range(grouped_df_sum.shape[0]), np.sort(np.log1p(grouped_df_sum["totals.transactionRevenue"].values)))
plt.title('Total Transaction Revenue for each Visitor')
plt.xlabel('index', fontsize=14)
plt.ylabel('TransactionRevenue', fontsize=14)
plt.show()
non_zero_instance = pd.notnull(train_df["totals.transactionRevenue"]).sum()
print("Number of instances in train set with non-zero revenue : ", non_zero_instance, " and ratio is : ", non_zero_instance / train_df.shape[0])
non_zero_revenue = (grouped_df_sum["totals.transactionRevenue"]>0).sum()
print("Number of unique customers with non-zero revenue : ", non_zero_revenue, "and the ratio is : ", non_zero_revenue / train_df.shape[0])
print("Number of unique visitors in train set : ",train_df.fullVisitorId.nunique(), " out of rows : ",train_df.shape[0])
print("Number of unique visitors in test set : ",test_df.fullVisitorId.nunique(), " out of rows : ",test_df.shape[0])
print("Number of common visitors in train and test set : ",len(set(train_df.fullVisitorId.unique()).intersection(set(test_df.fullVisitorId.unique())) ))
```
Charting the Transactions and Revenue per Date
| github_jupyter |
<a href="https://colab.research.google.com/github/intel-analytics/analytics-zoo/blob/master/docs/docs/colab-notebook/orca/quickstart/autoxgboost_regressor_sklearn_boston.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>

---
##### Copyright 2018 Analytics Zoo Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#
```
## **Environment Preparation**
**Install Java 8**
Run the cell on the **Google Colab** to install jdk 1.8.
**Note:** if you run this notebook on your computer, root permission is required when running the cell to install Java 8. (You may ignore this cell if Java 8 has already been set up in your computer).
```
# Install jdk8
!apt-get install openjdk-8-jdk-headless -qq > /dev/null
import os
# Set environment variable JAVA_HOME.
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-8-openjdk-amd64"
!update-alternatives --set java /usr/lib/jvm/java-8-openjdk-amd64/jre/bin/java
!java -version
```
**Install Analytics Zoo**
[Conda](https://docs.conda.io/projects/conda/en/latest/user-guide/install/) is needed to prepare the Python environment for running this example.
**Note**: The following code cell is specific for setting up conda environment on Colab; for general conda installation, please refer to the [install guide](https://docs.conda.io/projects/conda/en/latest/user-guide/install/) for more details.
```
import sys
# Set current python version
python_version = "3.7.10"
# Install Miniconda
!wget https://repo.continuum.io/miniconda/Miniconda3-4.5.4-Linux-x86_64.sh
!chmod +x Miniconda3-4.5.4-Linux-x86_64.sh
!./Miniconda3-4.5.4-Linux-x86_64.sh -b -f -p /usr/local
# Update Conda
!conda install --channel defaults conda python=$python_version --yes
!conda update --channel defaults --all --yes
# Append to the sys.path
_ = (sys.path
.append(f"/usr/local/lib/python3.7/site-packages"))
os.environ['PYTHONHOME']="/usr/local"
```
You can install the latest pre-release version using `pip install --pre --upgrade analytics-zoo[ray]`.
```
# Install latest pre-release version of Analytics Zoo
# Installing Analytics Zoo from pip will automatically install pyspark, bigdl, and their dependencies.
!pip install --pre --upgrade analytics-zoo
# Install python dependencies
!pip install ray[tune]
!pip install xgboost==1.3.3
!pip install tensorboardx==2.2
```
## **Distributed Automl for xgboost using Orca AutoXGBoost**
Orca AutoXGBoost enables distributed automated hyper-parameter tuning for XGBoost, which includes `AutoXGBRegressor` and `AutoXGBClassifier` for sklearn `XGBRegressor` and `XGBClassifier` respectively. See more about [xgboost scikit-learn API](https://xgboost.readthedocs.io/en/latest/python/python_api.html#module-xgboost.sklearn).
In this guide we will describe how to use Orca AutoXGBoost for automated xgboost tuning in 4 simple steps.
## **Step 0: Prepare dataset**
We use [sklearn boston house-price dataset](https://scikit-learn.org/stable/datasets/toy_dataset.html#boston-dataset) for demonstration.
```
# load data
from sklearn.datasets import load_boston
boston = load_boston()
y = boston['target']
X = boston['data']
# split the data into train and test set
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
```
### **Step 1: Init Orca Context**
```
# import necesary libraries and modules
from __future__ import print_function
import os
import argparse
from zoo.orca import init_orca_context, stop_orca_context
from zoo.orca import OrcaContext
# recommended to set it to True when running Analytics Zoo in Jupyter notebook.
OrcaContext.log_output = True # (this will display terminal's stdout and stderr in the Jupyter notebook).
cluster_mode = "local"
if cluster_mode == "local":
init_orca_context(cores=6, memory="2g", init_ray_on_spark=True) # run in local mode
elif cluster_mode == "k8s":
init_orca_context(cluster_mode="k8s", num_nodes=2, cores=4, init_ray_on_spark=True) # run on K8s cluster
elif cluster_mode == "yarn":
init_orca_context(
cluster_mode="yarn-client", cores=4, num_nodes=2, memory="2g", init_ray_on_spark=True,
driver_memory="10g", driver_cores=1) # run on Hadoop YARN cluster
```
This is the only place where you need to specify local or distributed mode. View [Orca Context](https://analytics-zoo.readthedocs.io/en/latest/doc/Orca/Overview/orca-context.html) for more details.
**Note**: You should export HADOOP_CONF_DIR=/path/to/hadoop/conf/dir when you run on Hadoop YARN cluster.
### **Step 2: Define Search space**
You should define a dictionary as your hyper-parameter search space for `XGBRegressor`. The keys are hyper-parameter names you want to search for `XGBRegressor`, and you can specify how you want to sample each hyper-parameter in the values of the search space. See [automl.hp](https://analytics-zoo.readthedocs.io/en/latest/doc/PythonAPI/AutoML/automl.html#orca-automl-hp) for more details.
```
from zoo.orca.automl import hp
search_space = {
"n_estimators": hp.grid_search([50, 100, 200]),
"max_depth": hp.choice([2, 4, 6]),
}
```
### **Step 3: Automatically fit and search with Orca AutoXGBoost**
We will then fit AutoXGBoost automatically on [Boston Housing dataset](https://scikit-learn.org/stable/datasets/toy_dataset.html#boston-dataset).
First create an `AutoXGBRegressor`.
You could also pass the sklearn `XGBRegressor` parameters to `AutoXGBRegressor`. Note that the `XGBRegressor` parameters shouldn't include the hyper-parameters in `search_space` or `n_jobs`, which is the same with `cpus_per_trial`.
```
from zoo.orca.automl.xgboost import AutoXGBRegressor
auto_xgb_reg = AutoXGBRegressor(cpus_per_trial=2,
name="auto_xgb_classifier",
min_child_weight=3,
random_state=2)
```
Next, use the auto xgboost regressor to fit and search for the best hyper-parameter set.
```
auto_xgb_reg.fit(data=(X_train, y_train),
validation_data=(X_test, y_test),
search_space=search_space,
n_sampling=2,
metric="rmse")
```
### **Step 4: Get best model and hyper parameters**
You can get the best learned model from the fitted auto xgboost regressor, which is an sklearn `XGBRegressor` instance.
```
best_model = auto_xgb_reg.get_best_model()
```
You can also get the best hyper-parameter set.
```
best_config = auto_xgb_reg.get_best_config()
print(best_config)
```
Then, you can use the best learned model as you want. Here, we demonstrate how to predict and evaluate on the test dataset.
```
y_pred = best_model.predict(X_test)
from sklearn.metrics import mean_squared_error
print(mean_squared_error(y_test, y_pred))
# stop orca context when program finishes
stop_orca_context()
```
| github_jupyter |

<a href="https://hub.callysto.ca/jupyter/hub/user-redirect/git-pull?repo=https%3A%2F%2Fgithub.com%2Fcallysto%2Fcurriculum-notebooks&branch=master&subPath=Science/MethodsUsedInStudyingTheEarth/methods-used-in-studying-the-earth.ipynb&depth=1" target="_parent"><img src="https://raw.githubusercontent.com/callysto/curriculum-notebooks/master/open-in-callysto-button.svg?sanitize=true" width="123" height="24" alt="Open in Callysto"/></a>
# Methods used in Scientific Study of the Earth
```
from IPython.display import HTML, Image
HTML('''<script>
code_show=true;
function code_toggle() {
if (code_show){
$('div.input').hide();
} else {
$('div.input').show();
}
code_show = !code_show
}
$( document ).ready(code_toggle);
</script>
<form action="javascript:code_toggle()"><input type="submit" value="Click here to toggle on/off the raw code."></form>''')
```
## Introduction
Over the past 4.6 billion years, the Earth has been constantly changing. As we know, the earth is always spinning and plate tectonics results in the the movement and formation of different continents. These changes occur over millions of years and can be observed from the surface by observing land forms and types of rocks in the area. There are many specific indications that showcase different changes. These indications can be separated into two categories, sudden change and gradual change.
### Sudden Change
After a change occurs in the subsurface, it is brought to the surface as an event that is commonly classified as a "natural disaster" such as earthquakes and volcanoes. The main cause of these events is plate tectonics. Earth's crust is divided into many pieces called plates which move in relation to one another. Typically, plates move at a rate of 10cm/yr which seems very slow. As the plates move, they exert pressure on one another can result in earthquakes or volcanic eruptions.
```
from __future__ import print_function
from IPython.display import Image, HTML, clear_output
from ipywidgets import interact, interactive, fixed, interact_manual
import ipywidgets as widgets
import random
```
<img src="https://www.sciencealert.com/images/parks-plates_cascadia_subduction_zone_revised-01.jpg" alt="Volcano" width="420" />
Figure 1: Cross section of the volcano. One plate is being subducted underneath the other. The subducted plate is melting by the hot mantle which causes lava to reach the surface and escape through the surface in a volcanic eruption. This process can also cause earthquakes. (ScienceAlert)
### Gradual Change
Over time, much of the surface of the Earth has been subject to long term changes by wind and water. Rivers are channels of water which erode the surrounding area as they continue to flow while wind displaces dust and dirt, changing the shapes of the formations.

Figure 2: Erosion by fast moving water can cause different landforms to be created such as a waterfall. (Alexandra falls, NWT, Canada. Photo credits: Alyson Birce)
## Modelling Earth's Interior
From the surface of the Earth, we have seen the different types of changes. The trends regarding the magnetic and gravitational fields can provide insight into the cross section of the Earth. From lots of research, the current model of the Earth is shown below.

Figure 3: Cross section of the Earth (U.S Geological Survey)
## Tools/ Techniques
There are a variety of methods that are used to observe the subsurface at a depth that can not be reached by simply using a shovel.
### Seismographs
A Seismograph is an instrument that is used to record fluctuations in the ground shown in figure 4. This is commonly used to predict and record earthquakes. There are 3 types of earthquake waves: p waves, s waves and surface waves which occur in that order respectively. All of these can be seen on seismograph records as seen in Figure 5 below. An earthquake begins at a set point called the Focal point which the source of wave propagation. P waves are also referred to as primary waves and are recorded first as they move the fastest and it is transmitted through the whole Earth. S waves are also referred to as secondary waves and are the second wave to be recorded. These waves can not travel through liquids such as the mantle. This can be seen in Figure 5. A surface wave is the mechanical wave that causes the most damage during the Earthquake.

Figure 4: The Seismograph instrument used to record earthquakes. (Indiana State University)
Figure 5: This diagram shows how the waves propagate through the earth during the earthquake. The focal point is marked by the white circle at the beginning of the video, then the waves propagate outwards. The P waves are marked with red lines, the S waves are marked with blue lines and the surface waves are marked with yellow lines. (GIPHY)
### Core Drills
When analyzing the composition of the subsurface, we can use a drill to extract a long cylinder section of Earth. This is used to determine the composition which is useful when looking for evidence of oil.

Figure 6: A drill bit for taking core samples

Figure 7: Geological core samples
## Describing Rocks and Minerals
### Definitions
**Rock** - a natural substance which is composed of minerals.
**Mineral** - a naturally occurring chemical compound that is not produced by life processes.
Rocks and minerals can not be accurately identified by colour alone. The other properties must be taken into account for proper identification. These properties include luster, transparency, cleavage and fracture, and the Moh's hardness scale. In this section, we will explore each of the properties.
### Luster
This describes the "shinyness" of the sample. This assists with categorizing minerals as metals verses non-metals.

Figure 8: Pyrite with a metallic luster

Figure 9: Kaolinite with a dull luster
### Transparency
Can you see through the mineral? If so, how much?

Figure 10: Quartz, which is relatively transparent (photo by [JJ Harrison](https://www.jjharrison.com.au))

Figure 11: A transparent form of calcite (photo by [Joan Rosell](https://www.rosellminerals.com/minerales.php?idmineral=160&yng=1))
### Cleavage and Fracture
This describes the shape of the rock if it were to break.
**Fracture** - When the rock breaks, the edges appear rough and cracked.

Figure 12: Fractured graphite (photo by [Alchemist-hp](www.pse-mendelejew.de))
**Cleavage** - When the rock breaks, it would break along a plane where the edges appear smooth as if they were cut. This can be further subdivided into subcategories based on the number of directions the planes run. These are shown below.
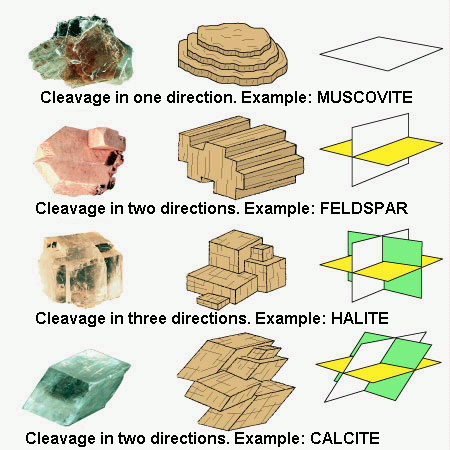
Figure 13: Subcategories of cleavage based on the number of planes that are apparent (by [David Seidemann and David Leveson](http://academic.brooklyn.cuny.edu/geology/grocha/rocks/index.html))
### Moh's Hardness scale
The hardness of a rock is ranked on a scale of 1-10 based on how easy it can be scratched. Common rocks as well as tools can be used to compare hardness, for example fingernail have a hardness of about 2.5. Typically the chosen material is scratched against the surface of the unknown rock. The surface that ends up with a scratch mark is the softer substance.

Figure 14: Mohs hardness kit, containing one specimen of each mineral on the ten-point hardness scale (from [Wikipedia](https://en.wikipedia.org/wiki/Mohs_scale_of_mineral_hardness))
### References
1. ScienceAlert, Discover Ideas on Active volcanoes, Pinterest, accessed 3 July 2018
<https://www.pinterest.com.au/pin/849421179688448478 >
2. U.S Geological Survey, Earth Cross-Section, accessed 3 July 2018
<https://www.usgs.gov/media/images/earth-cross-section>
3. Nevada Seismological Laboratory, Seismic Deformation, accessed 3 July 2018
<http://crack.seismo.unr.edu/ftp/pub/louie/class/100/seismic-waves.html>
4. GIPHY, Earth GIFs, accessed 3 July 2018
<https://media.giphy.com/media/kQPS6ASP23cvC/giphy.gif>
5. Wintershall, The Well, accessed 6 July 2018
<https://www.wintershall.com/technology-innovation/drilling.html>
6. The Northern Miner, accessed 6 July 2018
<http://www.northernminer.com/news/balmoral-delivers-more-high-grade-nickel-pgm-at-grasset/1003214678/>
7. geology.com, Hematite, accessed 6 July 2018
<https://geology.com/minerals/hematite.shtml>
8. geology rocks and minerals, quartzite, accessed 6 July 2018
<https://flexiblelearning.auckland.ac.nz/rocks_minerals/rocks/quartzite.html>
9. Wikipedia, accessed 28 January 2019
<https://en.wikipedia.org/wiki/Quartz>
10. Lock Haven, calcite, accessed 6 July 2018
<https://www.lockhaven.edu/~dsimanek/14/stereo.htm>
11. Arkansas Geological Survey, zinc ore, accessed 9 July 2018
<https://arkansasgeological.wordpress.com/tag/sphalerite/>
12. Brooklyn College, Cleavage, accessed 9 July 2018
<http://academic.brooklyn.cuny.edu/geology/grocha/mineral/cleavage.html>
13. Amazon, Moh's Hardness scale, accessed 9 July 2018
<https://www.amazon.com/Mohs-Hardness-Scale-Collection-specimens/dp/B00K24O1G8>
[](https://github.com/callysto/curriculum-notebooks/blob/master/LICENSE.md)
| github_jupyter |
# 视频动作识别
视频动作识别是指对一小段视频中的内容进行分析,判断视频中的人物做了哪种动作。视频动作识别与图像领域的图像识别,既有联系又有区别,图像识别是对一张静态图片进行识别,而视频动作识别不仅要考察每张图片的静态内容,还要考察不同图片静态内容之间的时空关系。比如一个人扶着一扇半开的门,仅凭这一张图片无法判断该动作是开门动作还是关门动作。
视频分析领域的研究相比较图像分析领域的研究,发展时间更短,也更有难度。视频分析模型完成的难点首先在于,需要强大的计算资源来完成视频的分析。视频要拆解成为图像进行分析,导致模型的数据量十分庞大。视频内容有很重要的考虑因素是动作的时间顺序,需要将视频转换成的图像通过时间关系联系起来,做出判断,所以模型需要考虑时序因素,加入时间维度之后参数也会大量增加。
得益于PASCAL VOC、ImageNet、MS COCO等数据集的公开,图像领域产生了很多的经典模型,那么在视频分析领域有没有什么经典的模型呢?答案是有的,本案例将为大家介绍视频动作识别领域的经典模型并进行代码实践。
由于本案例的代码是在华为云ModelArts Notebook上运行,所以需要先按照如下步骤来进行Notebook环境的准备。
### 进入ModelArts
点击如下链接:https://www.huaweicloud.com/product/modelarts.html , 进入ModelArts主页。点击“立即使用”按钮,输入用户名和密码登录,进入ModelArts使用页面。
### 创建ModelArts Notebook
下面,我们在ModelArts中创建一个Notebook开发环境,ModelArts Notebook提供网页版的Python开发环境,可以方便的编写、运行代码,并查看运行结果。
第一步:在ModelArts服务主界面依次点击“开发环境”、“创建”

第二步:填写notebook所需的参数:
| 参数 | 说明 |
| - - - - - | - - - - - |
| 计费方式 | 按需计费 |
| 名称 | Notebook实例名称,如 action_recognition |
| 工作环境 | Python3 |
| 资源池 | 选择"公共资源池"即可 |
| 类型 | 本案例使用较为复杂的深度神经网络模型,需要较高算力,选择"GPU" |
| 规格 | 选择"[限时免费]体验规格GPU版" |
| 存储配置 | 选择EVS,磁盘规格5GB |
第三步:配置好Notebook参数后,点击下一步,进入Notebook信息预览。确认无误后,点击“立即创建”
第四步:创建完成后,返回开发环境主界面,等待Notebook创建完毕后,打开Notebook,进行下一步操作。

### 在ModelArts中创建开发环境
接下来,我们创建一个实际的开发环境,用于后续的实验步骤。
第一步:点击下图所示的“打开”按钮,进入刚刚创建的Notebook

第二步:创建一个Python3环境的的Notebook。点击右上角的"New",然后创建TensorFlow 1.13.1开发环境。
第三步:点击左上方的文件名"Untitled",并输入一个与本实验相关的名称,如"action_recognition"


### 在Notebook中编写并执行代码
在Notebook中,我们输入一个简单的打印语句,然后点击上方的运行按钮,可以查看语句执行的结果:

开发环境准备好啦,接下来可以愉快地写代码啦!
### 准备源代码和数据
这一步准备案例所需的源代码和数据,相关资源已经保存在OBS中,我们通过ModelArts SDK将资源下载到本地,并解压到当前目录下。解压后,当前目录包含data、dataset_subset和其他目录文件,分布是预训练参数文件、数据集和代码文件等。
```
import os
if not os.path.exists('videos'):
from modelarts.session import Session
session = Session()
session.download_data(bucket_path="ai-course-common-26-bj4/video/video.tar.gz", path="./video.tar.gz")
# 使用tar命令解压资源包
os.system("tar xf ./video.tar.gz")
# 使用rm命令删除压缩包
os.system("rm ./video.tar.gz")
```
上一节课我们已经介绍了视频动作识别有HMDB51、UCF-101和Kinetics三个常用的数据集,本案例选用了UCF-101数据集的部分子集作为演示用数据集,接下来,我们播放一段UCF-101中的视频:
```
video_name = "./data/v_TaiChi_g01_c01.avi"
from IPython.display import clear_output, Image, display, HTML
import time
import cv2
import base64
import numpy as np
def arrayShow(img):
_,ret = cv2.imencode('.jpg', img)
return Image(data=ret)
cap = cv2.VideoCapture(video_name)
while True:
try:
clear_output(wait=True)
ret, frame = cap.read()
if ret:
tmp = cv2.cvtColor(frame, cv2.COLOR_BGR2RGB)
img = arrayShow(frame)
display(img)
time.sleep(0.05)
else:
break
except KeyboardInterrupt:
cap.release()
cap.release()
```
## 视频动作识别模型介绍
在图像领域中,ImageNet作为一个大型图像识别数据集,自2010年开始,使用此数据集训练出的图像算法层出不穷,深度学习模型经历了从AlexNet到VGG-16再到更加复杂的结构,模型的表现也越来越好。在识别千种类别的图片时,错误率表现如下:
<img src="./img/ImageNet.png" width="500" height="500" align=center>
在图像识别中表现很好的模型,可以在图像领域的其他任务中继续使用,通过复用模型中部分层的参数,就可以提升模型的训练效果。有了基于ImageNet模型的图像模型,很多模型和任务都有了更好的训练基础,比如说物体检测、实例分割、人脸检测、人脸识别等。
那么训练效果显著的图像模型是否可以用于视频模型的训练呢?答案是yes,有研究证明,在视频领域,如果能够复用图像模型结构,甚至参数,将对视频模型的训练有很大帮助。但是怎样才能复用上图像模型的结构呢?首先需要知道视频分类与图像分类的不同,如果将视频视作是图像的集合,每一个帧将作为一个图像,视频分类任务除了要考虑到图像中的表现,也要考虑图像间的时空关系,才可以对视频动作进行分类。
为了捕获图像间的时空关系,论文[I3D](https://arxiv.org/pdf/1705.07750.pdf)介绍了三种旧的视频分类模型,并提出了一种更有效的Two-Stream Inflated 3D ConvNets(简称I3D)的模型,下面将逐一简介这四种模型,更多细节信息请查看原论文。
### 旧模型一:卷积网络+LSTM
模型使用了训练成熟的图像模型,通过卷积网络,对每一帧图像进行特征提取、池化和预测,最后在模型的末端加一个LSTM层(长短期记忆网络),如下图所示,这样就可以使模型能够考虑时间性结构,将上下文特征联系起来,做出动作判断。这种模型的缺点是只能捕获较大的工作,对小动作的识别效果较差,而且由于视频中的每一帧图像都要经过网络的计算,所以训练时间很长。
<img src="./img/video_model_0.png" width="200" height="200" align=center>
### 旧模型二:3D卷积网络
3D卷积类似于2D卷积,将时序信息加入卷积操作。虽然这是一种看起来更加自然的视频处理方式,但是由于卷积核维度增加,参数的数量也增加了,模型的训练变得更加困难。这种模型没有对图像模型进行复用,而是直接将视频数据传入3D卷积网络进行训练。
<img src="./img/video_model_1.png" width="150" height="150" align=center>
### 旧模型三:Two-Stream 网络
Two-Stream 网络的两个流分别为**1张RGB快照**和**10张计算之后的光流帧画面组成的栈**。两个流都通过ImageNet预训练好的图像卷积网络,光流部分可以分为竖直和水平两个通道,所以是普通图片输入的2倍,模型在训练和测试中表现都十分出色。
<img src="./img/video_model_2.png" width="400" height="400" align=center>
#### 光流视频 optical flow video
上面讲到了光流,在此对光流做一下介绍。光流是什么呢?名字很专业,感觉很陌生,但实际上这种视觉现象我们每天都在经历,我们坐高铁的时候,可以看到窗外的景物都在快速往后退,开得越快,就感受到外面的景物就是“刷”地一个残影,这种视觉上目标的运动方向和速度就是光流。光流从概念上讲,是对物体运动的观察,通过找到相邻帧之间的相关性来判断帧之间的对应关系,计算出相邻帧画面中物体的运动信息,获取像素运动的瞬时速度。在原始视频中,有运动部分和静止的背景部分,我们通常需要判断的只是视频中运动部分的状态,而光流就是通过计算得到了视频中运动部分的运动信息。
下面是一个经过计算后的原视频及光流视频。
原视频

光流视频

### 新模型:Two-Stream Inflated 3D ConvNets
新模型采取了以下几点结构改进:
- 拓展2D卷积为3D。直接利用成熟的图像分类模型,只不过将网络中二维$ N × N $的 filters 和 pooling kernels 直接变成$ N × N × N $;
- 用 2D filter 的预训练参数来初始化 3D filter 的参数。上一步已经利用了图像分类模型的网络,这一步的目的是能利用上网络的预训练参数,直接将 2D filter 的参数直接沿着第三个时间维度进行复制N次,最后将所有参数值再除以N;
- 调整感受野的形状和大小。新模型改造了图像分类模型Inception-v1的结构,前两个max-pooling层改成使用$ 1 × 3 × 3 $kernels and stride 1 in time,其他所有max-pooling层都仍然使用对此的kernel和stride,最后一个average pooling层使用$ 2 × 7 × 7 $的kernel。
- 延续了Two-Stream的基本方法。用双流结构来捕获图片之间的时空关系仍然是有效的。
最后新模型的整体结构如下图所示:
<img src="./img/video_model_3.png" width="200" height="200" align=center>
好,到目前为止,我们已经讲解了视频动作识别的经典数据集和经典模型,下面我们通过代码来实践地跑一跑其中的两个模型:**C3D模型**( 3D卷积网络)以及**I3D模型**(Two-Stream Inflated 3D ConvNets)。
### C3D模型结构
我们已经在前面的“旧模型二:3D卷积网络”中讲解到3D卷积网络是一种看起来比较自然的处理视频的网络,虽然它有效果不够好,计算量也大的特点,但它的结构很简单,可以构造一个很简单的网络就可以实现视频动作识别,如下图所示是3D卷积的示意图:

a)中,一张图片进行了2D卷积, b)中,对视频进行2D卷积,将多个帧视作多个通道, c)中,对视频进行3D卷积,将时序信息加入输入信号中。
ab中,output都是一张二维特征图,所以无论是输入是否有时间信息,输出都是一张二维的特征图,2D卷积失去了时序信息。只有3D卷积在输出时,保留了时序信息。2D和3D池化操作同样有这样的问题。
如下图所示是一种[C3D](https://www.cv-foundation.org/openaccess/content_iccv_2015/papers/Tran_Learning_Spatiotemporal_Features_ICCV_2015_paper.pdf)网络的变种:(如需阅读原文描述,请查看I3D论文 2.2 节)

C3D结构,包括8个卷积层,5个最大池化层以及2个全连接层,最后是softmax输出层。
所有的3D卷积核为$ 3 × 3 × 3$ 步长为1,使用SGD,初始学习率为0.003,每150k个迭代,除以2。优化在1.9M个迭代的时候结束,大约13epoch。
数据处理时,视频抽帧定义大小为:$ c × l × h × w$,$c$为通道数量,$l$为帧的数量,$h$为帧画面的高度,$w$为帧画面的宽度。3D卷积核和池化核的大小为$ d × k × k$,$d$是核的时间深度,$k$是核的空间大小。网络的输入为视频的抽帧,预测出的是类别标签。所有的视频帧画面都调整大小为$ 128 × 171 $,几乎将UCF-101数据集中的帧调整为一半大小。视频被分为不重复的16帧画面,这些画面将作为模型网络的输入。最后对帧画面的大小进行裁剪,输入的数据为$16 × 112 × 112 $
### C3D模型训练
接下来,我们将对C3D模型进行训练,训练过程分为:数据预处理以及模型训练。在此次训练中,我们使用的数据集为UCF-101,由于C3D模型的输入是视频的每帧图片,因此我们需要对数据集的视频进行抽帧,也就是将视频转换为图片,然后将图片数据传入模型之中,进行训练。
在本案例中,我们随机抽取了UCF-101数据集的一部分进行训练的演示,感兴趣的同学可以下载完整的UCF-101数据集进行训练。
[UCF-101下载](https://www.crcv.ucf.edu/data/UCF101.php)
数据集存储在目录` dataset_subset`下
如下代码是使用cv2库进行视频文件到图片文件的转换
```
import cv2
import os
# 视频数据集存储位置
video_path = './dataset_subset/'
# 生成的图像数据集存储位置
save_path = './dataset/'
# 如果文件路径不存在则创建路径
if os.path.exists(save_path) == False:
os.mkdir(save_path)
# 获取动作列表
action_list = os.listdir(video_path)
# 遍历所有动作
for action in action_list:
if action.startswith(".")==False:
if not os.path.exists(save_path+action):
os.mkdir(save_path+action)
video_list = os.listdir(video_path+action)
# 遍历所有视频
for video in video_list:
prefix = video.split('.')[0]
if not os.path.exists(save_path+action+'/'+prefix):
os.mkdir(save_path+action+'/'+prefix)
save_name = save_path + action + '/' + prefix + '/'
video_name = video_path+action+'/'+video
# 读取视频文件
# cap为视频的帧
cap = cv2.VideoCapture(video_name)
# fps为帧率
fps = int(cap.get(cv2.CAP_PROP_FRAME_COUNT))
fps_count = 0
for i in range(fps):
ret, frame = cap.read()
if ret:
# 将帧画面写入图片文件中
cv2.imwrite(save_name+str(10000+fps_count)+'.jpg',frame)
fps_count += 1
```
此时,视频逐帧转换成的图片数据已经存储起来,为模型训练做准备。
### 模型训练
首先,我们构建模型结构。
C3D模型结构我们之前已经介绍过,这里我们通过`keras`提供的Conv3D,MaxPool3D,ZeroPadding3D等函数进行模型的搭建。
```
from keras.layers import Dense,Dropout,Conv3D,Input,MaxPool3D,Flatten,Activation, ZeroPadding3D
from keras.regularizers import l2
from keras.models import Model, Sequential
# 输入数据为 112×112 的图片,16帧, 3通道
input_shape = (112,112,16,3)
# 权重衰减率
weight_decay = 0.005
# 类型数量,我们使用UCF-101 为数据集,所以为101
nb_classes = 101
# 构建模型结构
inputs = Input(input_shape)
x = Conv3D(64,(3,3,3),strides=(1,1,1),padding='same',
activation='relu',kernel_regularizer=l2(weight_decay))(inputs)
x = MaxPool3D((2,2,1),strides=(2,2,1),padding='same')(x)
x = Conv3D(128,(3,3,3),strides=(1,1,1),padding='same',
activation='relu',kernel_regularizer=l2(weight_decay))(x)
x = MaxPool3D((2,2,2),strides=(2,2,2),padding='same')(x)
x = Conv3D(128,(3,3,3),strides=(1,1,1),padding='same',
activation='relu',kernel_regularizer=l2(weight_decay))(x)
x = MaxPool3D((2,2,2),strides=(2,2,2),padding='same')(x)
x = Conv3D(256,(3,3,3),strides=(1,1,1),padding='same',
activation='relu',kernel_regularizer=l2(weight_decay))(x)
x = MaxPool3D((2,2,2),strides=(2,2,2),padding='same')(x)
x = Conv3D(256, (3, 3, 3), strides=(1, 1, 1), padding='same',
activation='relu',kernel_regularizer=l2(weight_decay))(x)
x = MaxPool3D((2, 2, 2), strides=(2, 2, 2), padding='same')(x)
x = Flatten()(x)
x = Dense(2048,activation='relu',kernel_regularizer=l2(weight_decay))(x)
x = Dropout(0.5)(x)
x = Dense(2048,activation='relu',kernel_regularizer=l2(weight_decay))(x)
x = Dropout(0.5)(x)
x = Dense(nb_classes,kernel_regularizer=l2(weight_decay))(x)
x = Activation('softmax')(x)
model = Model(inputs, x)
```
通过keras提供的`summary()`方法,打印模型结构。可以看到模型的层构建以及各层的输入输出情况。
```
model.summary()
```
通过keras的`input`方法可以查看模型的输入形状,shape分别为`( batch size, width, height, frames, channels) ` 。
```
model.input
```
可以看到模型的数据处理的维度与图像处理模型有一些差别,多了frames维度,体现出时序关系在视频分析中的影响。
接下来,我们开始将图片文件转为训练需要的数据形式。
```
# 引用必要的库
from keras.optimizers import SGD,Adam
from keras.utils import np_utils
import numpy as np
import random
import cv2
import matplotlib.pyplot as plt
# 自定义callbacks
from schedules import onetenth_4_8_12
```
参数定义
```
img_path = save_path # 图片文件存储位置
results_path = './results' # 训练结果保存位置
if not os.path.exists(results_path):
os.mkdir(results_path)
```
数据集划分,随机抽取4/5 作为训练集,其余为验证集。将文件信息分别存储在`train_list`和`test_list`中,为训练做准备。
```
cates = os.listdir(img_path)
train_list = []
test_list = []
# 遍历所有的动作类型
for cate in cates:
videos = os.listdir(os.path.join(img_path, cate))
length = len(videos)//5
# 训练集大小,随机取视频文件加入训练集
train= random.sample(videos, length*4)
train_list.extend(train)
# 将余下的视频加入测试集
for video in videos:
if video not in train:
test_list.append(video)
print("训练集为:")
print( train_list)
print("共%d 个视频\n"%(len(train_list)))
print("验证集为:")
print(test_list)
print("共%d 个视频"%(len(test_list)))
```
接下来开始进行模型的训练。
首先定义数据读取方法。方法`process_data`中读取一个batch的数据,包含16帧的图片信息的数据,以及数据的标注信息。在读取图片数据时,对图片进行随机裁剪和翻转操作以完成数据增广。
```
def process_data(img_path, file_list,batch_size=16,train=True):
batch = np.zeros((batch_size,16,112,112,3),dtype='float32')
labels = np.zeros(batch_size,dtype='int')
cate_list = os.listdir(img_path)
def read_classes():
path = "./classInd.txt"
with open(path, "r+") as f:
lines = f.readlines()
classes = {}
for line in lines:
c_id = line.split()[0]
c_name = line.split()[1]
classes[c_name] =c_id
return classes
classes_dict = read_classes()
for file in file_list:
cate = file.split("_")[1]
img_list = os.listdir(os.path.join(img_path, cate, file))
img_list.sort()
batch_img = []
for i in range(batch_size):
path = os.path.join(img_path, cate, file)
label = int(classes_dict[cate])-1
symbol = len(img_list)//16
if train:
# 随机进行裁剪
crop_x = random.randint(0, 15)
crop_y = random.randint(0, 58)
# 随机进行翻转
is_flip = random.randint(0, 1)
# 以16 帧为单位
for j in range(16):
img = img_list[symbol + j]
image = cv2.imread( path + '/' + img)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
image = cv2.resize(image, (171, 128))
if is_flip == 1:
image = cv2.flip(image, 1)
batch[i][j][:][:][:] = image[crop_x:crop_x + 112, crop_y:crop_y + 112, :]
symbol-=1
if symbol<0:
break
labels[i] = label
else:
for j in range(16):
img = img_list[symbol + j]
image = cv2.imread( path + '/' + img)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
image = cv2.resize(image, (171, 128))
batch[i][j][:][:][:] = image[8:120, 30:142, :]
symbol-=1
if symbol<0:
break
labels[i] = label
return batch, labels
batch, labels = process_data(img_path, train_list)
print("每个batch的形状为:%s"%(str(batch.shape)))
print("每个label的形状为:%s"%(str(labels.shape)))
```
定义data generator, 将数据批次传入训练函数中。
```
def generator_train_batch(train_list, batch_size, num_classes, img_path):
while True:
# 读取一个batch的数据
x_train, x_labels = process_data(img_path, train_list, batch_size=16,train=True)
x = preprocess(x_train)
# 形成input要求的数据格式
y = np_utils.to_categorical(np.array(x_labels), num_classes)
x = np.transpose(x, (0,2,3,1,4))
yield x, y
def generator_val_batch(test_list, batch_size, num_classes, img_path):
while True:
# 读取一个batch的数据
y_test,y_labels = process_data(img_path, train_list, batch_size=16,train=False)
x = preprocess(y_test)
# 形成input要求的数据格式
x = np.transpose(x,(0,2,3,1,4))
y = np_utils.to_categorical(np.array(y_labels), num_classes)
yield x, y
```
定义方法`preprocess`, 对函数的输入数据进行图像的标准化处理。
```
def preprocess(inputs):
inputs[..., 0] -= 99.9
inputs[..., 1] -= 92.1
inputs[..., 2] -= 82.6
inputs[..., 0] /= 65.8
inputs[..., 1] /= 62.3
inputs[..., 2] /= 60.3
return inputs
# 训练一个epoch大约需4分钟
# 类别数量
num_classes = 101
# batch大小
batch_size = 4
# epoch数量
epochs = 1
# 学习率大小
lr = 0.005
# 优化器定义
sgd = SGD(lr=lr, momentum=0.9, nesterov=True)
model.compile(loss='categorical_crossentropy', optimizer=sgd, metrics=['accuracy'])
# 开始训练
history = model.fit_generator(generator_train_batch(train_list, batch_size, num_classes,img_path),
steps_per_epoch= len(train_list) // batch_size,
epochs=epochs,
callbacks=[onetenth_4_8_12(lr)],
validation_data=generator_val_batch(test_list, batch_size,num_classes,img_path),
validation_steps= len(test_list) // batch_size,
verbose=1)
# 对训练结果进行保存
model.save_weights(os.path.join(results_path, 'weights_c3d.h5'))
```
## 模型测试
接下来我们将训练之后得到的模型进行测试。随机在UCF-101中选择一个视频文件作为测试数据,然后对视频进行取帧,每16帧画面传入模型进行一次动作预测,并且将动作预测以及预测百分比打印在画面中并进行视频播放。
首先,引入相关的库。
```
from IPython.display import clear_output, Image, display, HTML
import time
import cv2
import base64
import numpy as np
```
构建模型结构并且加载权重。
```
from models import c3d_model
model = c3d_model()
model.load_weights(os.path.join(results_path, 'weights_c3d.h5'), by_name=True) # 加载刚训练的模型
```
定义函数arrayshow,进行图片变量的编码格式转换。
```
def arrayShow(img):
_,ret = cv2.imencode('.jpg', img)
return Image(data=ret)
```
进行视频的预处理以及预测,将预测结果打印到画面中,最后进行播放。
```
# 加载所有的类别和编号
with open('./ucfTrainTestlist/classInd.txt', 'r') as f:
class_names = f.readlines()
f.close()
# 读取视频文件
video = './videos/v_Punch_g03_c01.avi'
cap = cv2.VideoCapture(video)
clip = []
# 将视频画面传入模型
while True:
try:
clear_output(wait=True)
ret, frame = cap.read()
if ret:
tmp = cv2.cvtColor(frame, cv2.COLOR_BGR2RGB)
clip.append(cv2.resize(tmp, (171, 128)))
# 每16帧进行一次预测
if len(clip) == 16:
inputs = np.array(clip).astype(np.float32)
inputs = np.expand_dims(inputs, axis=0)
inputs[..., 0] -= 99.9
inputs[..., 1] -= 92.1
inputs[..., 2] -= 82.6
inputs[..., 0] /= 65.8
inputs[..., 1] /= 62.3
inputs[..., 2] /= 60.3
inputs = inputs[:,:,8:120,30:142,:]
inputs = np.transpose(inputs, (0, 2, 3, 1, 4))
# 获得预测结果
pred = model.predict(inputs)
label = np.argmax(pred[0])
# 将预测结果绘制到画面中
cv2.putText(frame, class_names[label].split(' ')[-1].strip(), (20, 20),
cv2.FONT_HERSHEY_SIMPLEX, 0.6,
(0, 0, 255), 1)
cv2.putText(frame, "prob: %.4f" % pred[0][label], (20, 40),
cv2.FONT_HERSHEY_SIMPLEX, 0.6,
(0, 0, 255), 1)
clip.pop(0)
# 播放预测后的视频
lines, columns, _ = frame.shape
frame = cv2.resize(frame, (int(columns), int(lines)))
img = arrayShow(frame)
display(img)
time.sleep(0.02)
else:
break
except:
print(0)
cap.release()
```
## I3D 模型
在之前我们简单介绍了I3D模型,[I3D官方github库](https://github.com/deepmind/kinetics-i3d)提供了在Kinetics上预训练的模型和预测代码,接下来我们将体验I3D模型如何对视频进行预测。
首先,引入相关的包
```
import numpy as np
import tensorflow as tf
import i3d
```
进行参数的定义
```
# 输入图片大小
_IMAGE_SIZE = 224
# 视频的帧数
_SAMPLE_VIDEO_FRAMES = 79
# 输入数据包括两部分:RGB和光流
# RGB和光流数据已经经过提前计算
_SAMPLE_PATHS = {
'rgb': 'data/v_CricketShot_g04_c01_rgb.npy',
'flow': 'data/v_CricketShot_g04_c01_flow.npy',
}
# 提供了多种可以选择的预训练权重
# 其中,imagenet系列模型从ImageNet的2D权重中拓展而来,其余为视频数据下的预训练权重
_CHECKPOINT_PATHS = {
'rgb': 'data/checkpoints/rgb_scratch/model.ckpt',
'flow': 'data/checkpoints/flow_scratch/model.ckpt',
'rgb_imagenet': 'data/checkpoints/rgb_imagenet/model.ckpt',
'flow_imagenet': 'data/checkpoints/flow_imagenet/model.ckpt',
}
# 记录类别文件
_LABEL_MAP_PATH = 'data/label_map.txt'
# 类别数量为400
NUM_CLASSES = 400
```
定义参数:
- imagenet_pretrained :如果为`True`,则调用预训练权重,如果为`False`,则调用ImageNet转成的权重
```
imagenet_pretrained = True
# 加载动作类型
kinetics_classes = [x.strip() for x in open(_LABEL_MAP_PATH)]
tf.logging.set_verbosity(tf.logging.INFO)
```
构建RGB部分模型
```
rgb_input = tf.placeholder(tf.float32, shape=(1, _SAMPLE_VIDEO_FRAMES, _IMAGE_SIZE, _IMAGE_SIZE, 3))
with tf.variable_scope('RGB'):
rgb_model = i3d.InceptionI3d(NUM_CLASSES, spatial_squeeze=True, final_endpoint='Logits')
rgb_logits, _ = rgb_model(rgb_input, is_training=False, dropout_keep_prob=1.0)
rgb_variable_map = {}
for variable in tf.global_variables():
if variable.name.split('/')[0] == 'RGB':
rgb_variable_map[variable.name.replace(':0', '')] = variable
rgb_saver = tf.train.Saver(var_list=rgb_variable_map, reshape=True)
```
构建光流部分模型
```
flow_input = tf.placeholder(tf.float32,shape=(1, _SAMPLE_VIDEO_FRAMES, _IMAGE_SIZE, _IMAGE_SIZE, 2))
with tf.variable_scope('Flow'):
flow_model = i3d.InceptionI3d(NUM_CLASSES, spatial_squeeze=True, final_endpoint='Logits')
flow_logits, _ = flow_model(flow_input, is_training=False, dropout_keep_prob=1.0)
flow_variable_map = {}
for variable in tf.global_variables():
if variable.name.split('/')[0] == 'Flow':
flow_variable_map[variable.name.replace(':0', '')] = variable
flow_saver = tf.train.Saver(var_list=flow_variable_map, reshape=True)
```
将模型联合,成为完整的I3D模型
```
model_logits = rgb_logits + flow_logits
model_predictions = tf.nn.softmax(model_logits)
```
开始模型预测,获得视频动作预测结果。
预测数据为开篇提供的RGB和光流数据:


```
with tf.Session() as sess:
feed_dict = {}
if imagenet_pretrained:
rgb_saver.restore(sess, _CHECKPOINT_PATHS['rgb_imagenet'])
else:
rgb_saver.restore(sess, _CHECKPOINT_PATHS['rgb'])
tf.logging.info('RGB checkpoint restored')
rgb_sample = np.load(_SAMPLE_PATHS['rgb'])
tf.logging.info('RGB data loaded, shape=%s', str(rgb_sample.shape))
feed_dict[rgb_input] = rgb_sample
if imagenet_pretrained:
flow_saver.restore(sess, _CHECKPOINT_PATHS['flow_imagenet'])
else:
flow_saver.restore(sess, _CHECKPOINT_PATHS['flow'])
tf.logging.info('Flow checkpoint restored')
flow_sample = np.load(_SAMPLE_PATHS['flow'])
tf.logging.info('Flow data loaded, shape=%s', str(flow_sample.shape))
feed_dict[flow_input] = flow_sample
out_logits, out_predictions = sess.run(
[model_logits, model_predictions],
feed_dict=feed_dict)
out_logits = out_logits[0]
out_predictions = out_predictions[0]
sorted_indices = np.argsort(out_predictions)[::-1]
print('Norm of logits: %f' % np.linalg.norm(out_logits))
print('\nTop classes and probabilities')
for index in sorted_indices[:20]:
print(out_predictions[index], out_logits[index], kinetics_classes[index])
```
| github_jupyter |
### Dielectric cylinder
```
from GDM_PCE import *
np.random.seed(1)
```
#### Load data
```
n_samples_ = [150, 400, 800]
Error = []
N_clusters = []
Y_real, Y_recon, Diff, L2, R2, g_all, l_all, coord_all, X_all = [], [], [], [], [], [], [], [], []
for i in range(len(n_samples_)):
print('----Iteration: {} ----'.format(i))
n_samples = n_samples_[i]
# Probability distributions of input parameters
pdf1 = Uniform(loc=0.2, scale=0.5) # cylinder radius
pdf2 = Uniform(loc=8, scale=10) # strength of electric field
margs = [pdf1, pdf2]
joint = JointInd(marginals=margs)
# Draw samples
x = joint.rvs(n_samples)
if i == 0:
x_ = x[:100,:]
# Generate multi-dimensional output
ax, bx, ay, by = -1, 1, -1, 1
grid_points_x, grid_points_y = 80, 80
gridx = np.linspace(ax, bx, grid_points_x)
gridy = np.linspace(ay, by, grid_points_y)
xx, yy = np.meshgrid(gridx, gridy)
out = []
for i in range(x.shape[0]):
vfunc = np.vectorize(function)
u = vfunc(xx, yy, r0=x[i, 0], efield=x[i, 1])
out.append(u)
np.savez('data/electric_data.npz', X=x, Y=out)
file = np.load('data/electric_data.npz')
x = file['X']
data = file['Y']
start_time = time.time()
g, coord, Grass, residuals, index, evals = GDMaps(data=data, n_evecs=15, n_parsim=4, p='max').get() # Perform GDMAps
pce, error = PceModel(x=x, g=g, dist_obj=joint, max_degree=3, verbose=False).get() # Perform PCE on the manifold
# print('Error of PCE:', error)
print("--- Surrogate - %s seconds ---" % (time.time() - start_time))
# Adaptive clustering
start_time = time.time()
mat, indices, kmeans, L, n_clust, error_cl, clusters = AdaptClust(n_clust_max=50, Gr_object=Grass, data=g)
print("--- Adaptive clustering - %s seconds ---" % (time.time() - start_time))
Error.append(error_cl)
N_clusters.append(clusters)
g_all.append(g)
l_all.append(L)
coord_all.append(coord)
# Interpolators
models_all, pce_sigmas, dims = Interpolators(x=x, data=g, mat=mat, indices=indices, n_clust=n_clust, Gr=Grass, joint=joint)
# Predictions
n_pred = 10000
x_pred = joint.rvs(n_pred) # new samples
x_pred[:100,:] = x_
X_all.append(x_pred)
start_time = time.time()
y_real = []
for k in range(x_pred.shape[0]):
vfunc = np.vectorize(function) # Dielectric cylinder
y_real.append(vfunc(xx, yy, r0=x_pred[k, 0], efield=x_pred[k, 1]))
Y_real.append(y_real)
print("--- Comp. model cost - %s seconds ---" % (time.time() - start_time))
start_time = time.time()
y_recon, l2, r2, diff = Prediction(x_pred=x_pred, y_real=y_real, models_all=models_all, kmeans=kmeans, mat=mat,
pce=pce, pce_sigmas=pce_sigmas, Gr=Grass, dims=dims)
Y_recon.append(y_recon)
Diff.append(diff)
L2.append(l2)
R2.append(r2)
print("--- Out-of-sample predictions with GDMaps PCE - %s seconds ---" % (time.time() - start_time))
print('-------------------------------------------------------------------------------')
print('')
# Save results
import os
if not os.path.exists('results'):
os.makedirs('results')
np.savez('results/Results-Example-1.npz', x1=Error, x2=N_clusters, x3=g_all, x4=l_all, x5=coord_all, x6=Y_real, x7=Y_recon, x8=Diff, x9=L2, x10=R2)
# Load file
file_res = np.load('results/Results-Example-1.npz', allow_pickle=True)
Error = file_res['x1']
N_clusters = file_res['x2']
g_all = file_res['x3']
l_all = file_res['x4']
coord_all = file_res['x5']
Y_real = file_res['x6']
Y_recon = file_res['x7']
Diff = file_res['x8']
L2 = file_res['x9']
R2 = file_res['x10']
```
### Plot error vs number of clusters
```
import os
if not os.path.exists('figures'):
os.makedirs('figures')
os.makedirs('figures/Example1')
plt.rcParams.update({'font.size': 17})
fig, ax = plt.subplots(figsize=(7, 5), constrained_layout=True)
col = ['m', 'orange', 'b']
for i in range(len(n_samples_)):
plt.plot(N_clusters[i], Error[i], c=col[i], label='{}'.format(n_samples_[i]))
plt.scatter(N_clusters[i], Error[i], c=col[i], marker='*', s=150)
plt.ylabel('error')
plt.xlabel('# of clusters')
plt.legend(title='Training samples')
ax.ticklabel_format(style='sci', axis='y', scilimits=(0, 0))
ax.xaxis.set_major_locator(MaxNLocator(integer=True))
plt.grid()
plt.savefig('figures/Example1/Error-vs-number-of-clusters.png', bbox_inches='tight', dpi=300)
# Plot diffusion coordinates of the last n_samples_
plt.rcParams.update({'font.size': 20})
# Choose the last n_samples_
g = g_all[-1]
L = l_all[-1]
coord = coord_all[-1]
nlabels = np.unique(L).shape[0]
cmap = rand_cmap(nlabels=nlabels, type='bright', first_color_black=False, verbose=False)
comb1 = list(it.combinations(list(coord), 2))
comb2 = list(it.combinations([i for i in range(coord.shape[0])], 2))
# Plot first three plots
if coord.shape[0] > 2:
fig, ax = plt.subplots(nrows=1, ncols=3, figsize=(17,4), constrained_layout=True)
# fig.suptitle('Diffusion coordinates for {} training samples ({} clusters)'.format(n_samples_[-1], N_clusters[-1][-1]), fontsize=24)
j = 0
for i in range(3):
ax[i].scatter(g[:, comb2[i][0]], g[:, comb2[i][1]], s=50, c=L, cmap=cmap)
ax[i].set_xlabel(r'$\theta_{}$'.format(comb1[j][0]))
ax[i].set_ylabel(r'$\theta_{}$'.format(comb1[j][1]))
ax[i].grid('True')
ax[i].ticklabel_format(style='sci', axis='both', scilimits=(0, 0))
# plt.legend()
# ax[i].set_title('Training realizations: {}'.format(trunc[i]))
j += 1
#plt.subplots_adjust(left=None, bottom=None, right=None, top=None, wspace=0.4, hspace=None)
plt.savefig('figures/Example1/Diffusion-coord-cylinder.png', bbox_inches='tight', dpi=500)
```
### Reference vs prediction plot
```
# Plot 3 random test realizations with predictions and error
plt.rcParams.update({'font.size': 13})
fig, axs = plt.subplots(3, 3, figsize=(12, 8), constrained_layout=True)
cm = ['gnuplot2'] * 3
j = 0
k = 4
for row in range(3):
for col in range(3):
ax = axs[row, col]
if col == 0:
pcm = ax.contourf(Y_real[j][k], levels=np.linspace(-20, 20, 16), cmap=cm[col])
cbar = plt.colorbar(pcm, ax=ax)
ax.set_xticklabels([-1,-0.5,0,0.5,1, 1.5])
ax.set_yticklabels([-1,-0.5,0,0.5,1, 1.5])
#cbar.ax.locator_params(nbins=5)
if col == 1:
pcm = ax.contourf(Y_recon[j][k], levels=np.linspace(-20, 20, 16), cmap=cm[col])
plt.colorbar(pcm, ax=ax)
ax.set_xticklabels([-1,-0.5,0,0.5,1, 1.5])
ax.set_yticklabels([-1,-0.5,0,0.5,1, 1.5])
if col == 2:
pcm = ax.contourf(Diff[j][k], levels=np.linspace(0, 0.04, 16), cmap=cm[col])
plt.colorbar(pcm, ax=ax, format='%.0e')
ax.set_xticklabels([-1,-0.5,0,0.5,1, 1.5])
ax.set_yticklabels([-1,-0.5,0,0.5,1, 1.5])
if col == 0:
ax.set_ylabel('N = {} \n y'.format(n_samples_[row]))
if col ==0 and row ==0:
ax.set_title('Reference')
if col == 1 and row==0:
ax.set_title('Prediction')
if col == 2 and row==0:
ax.set_title('Relative error')
if row == 2:
ax.set_xlabel('x')
j += 1
#fig.suptitle('Test realizations for 3 training realizations', fontsize=24)
plt.savefig('figures/Example1/Comparison-cylinder.png', bbox_inches='tight', dpi=500)
# In addition to the above we'll show a Table with the mean and std of the l2 and r2 values for the 10,000 samples
# Compute the values next
# Compute mean and variance of errors
l2_mean = [np.round(np.mean(L2[i]), 6) for i in range(len(L2))]
l2_std = [np.round(np.std(L2[i]), 6) for i in range(len(L2))]
R2_mean = [np.round(np.mean(R2[i]), 6) for i in range(len(R2))]
R2_std = [np.round(np.std(R2[i]), 6) for i in range(len(R2))]
for i in range(3):
print('Training data: {}'.format(n_samples_[i]))
print('L2 mean {:e} '.format(l2_mean[i]))
print('L2 std: {:e} '. format(l2_std[i]))
print('R2 mean: {:e} '.format(R2_mean[i]))
print('R2 std: {:e} '.format(R2_std[i]))
print('')
```
### Moment estimation (UQ)
Comparison with MCS
Compute mean field with MCS and GDMaps PCE
```
### Make one figure to compare the three different training data
mean_recon, mean_mcs, mean_diff = [], [], []
for i in range(len(n_samples_)):
mean_recon.append(np.mean(np.array(Y_recon[i]), axis=0))
mean_mcs.append(np.mean(np.array(Y_real[i]), axis=0))
mean_diff.append(np.abs((mean_mcs[i] - mean_recon[i]) / mean_mcs[i]))
plt.rcParams.update({'font.size': 13})
j = 0
fig, axs = plt.subplots(len(n_samples_), 3, figsize=(12, 8), constrained_layout=True)
cm = ['gnuplot2'] * len(n_samples_)
for col in range(3):
for row in range(len(n_samples_)):
ax = axs[row, col]
if col == 0:
pcm = ax.contourf(mean_mcs[row+j], levels=np.linspace(-20, 20, 16), cmap=cm[col])
plt.colorbar(pcm, ax=ax)
ax.set_xticklabels([-1,-0.5,0,0.5,1, 1.5])
ax.set_yticklabels([-1,-0.5,0,0.5,1, 1.5])
if col == 1:
pcm = ax.contourf(mean_recon[row+j], levels=np.linspace(-20, 20, 16), cmap=cm[col])
plt.colorbar(pcm, ax=ax)
ax.set_xticklabels([-1,-0.5,0,0.5,1, 1.5])
ax.set_yticklabels([-1,-0.5,0,0.5,1, 1.5])
if col == 2:
pcm = ax.contourf(mean_diff[row+j], levels=np.linspace(0, 0.02, 16), cmap=cm[col])
plt.colorbar(pcm, ax=ax, format='%.0e')
ax.set_xticklabels([-1,-0.5,0,0.5,1, 1.5])
ax.set_yticklabels([-1,-0.5,0,0.5,1, 1.5])
if col == 0:
ax.set_ylabel('N = {} \n y'.format(n_samples_[row]))
if col ==0 and row ==0:
ax.set_title('MCS')
if col == 1 and row==0:
ax.set_title('GDM PCE')
if col == 2 and row==0:
ax.set_title('Relative error')
if row == 2:
ax.set_xlabel('x')
plt.savefig('figures/Example1/Mean-comparison-cylinder.png', bbox_inches='tight', dpi=500)
```
Compute variance field with MCS and GDMaps PCE
```
### Make one figure to compare the three different training data
var_recon, var_mcs, var_diff = [], [], []
for i in range(len(n_samples_)):
var_recon.append(np.var(np.array(Y_recon[i]), axis=0))
var_mcs.append(np.var(np.array(Y_real[i]), axis=0))
var_diff.append(np.abs((var_mcs[i] - var_recon[i]) / var_mcs[i]))
plt.rcParams.update({'font.size': 13})
fig, axs = plt.subplots(len(n_samples_), 3, figsize=(12, 8), constrained_layout=True)
cm = ['gnuplot2'] * len(n_samples_)
for col in range(3):
for row in range(len(n_samples_)):
ax = axs[row, col]
if col == 0:
pcm = ax.contourf(var_mcs[row], levels=np.linspace(0, 8, 16), cmap=cm[col])
plt.colorbar(pcm, ax=ax)
ax.set_xticklabels([-1,-0.5,0,0.5,1])
ax.set_yticklabels([-1,-0.5,0,0.5,1])
if col == 1:
pcm = ax.contourf(var_recon[row], levels=np.linspace(0, 8, 16), cmap=cm[col])
plt.colorbar(pcm, ax=ax)
ax.set_xticklabels([-1,-0.5,0,0.5,1, 1.5])
ax.set_yticklabels([-1,-0.5,0,0.5,1, 1.5])
if col == 2:
pcm = ax.contourf(var_diff[row], levels=np.linspace(0, 0.08, 16), cmap=cm[col])
plt.colorbar(pcm, ax=ax, format='%.0e')
ax.set_xticklabels([-1,-0.5,0,0.5,1, 1.5])
ax.set_yticklabels([-1,-0.5,0,0.5,1, 1.5])
if col == 0:
ax.set_ylabel('N = {} \n y'.format(n_samples_[row]))
if col ==0 and row ==0:
ax.set_title('MCS')
if col == 1 and row==0:
ax.set_title('GDM PCE')
if col == 2 and row==0:
ax.set_title('Relative error')
if row == 2:
ax.set_xlabel('x')
plt.savefig('figures/Example1/Variance-comparison-cylinder.png', bbox_inches='tight', dpi=500)
### Moment estimation for n_samples_[0] training data
plt.rcParams.update({'font.size': 15})
fig, axs = plt.subplots(2, 3, figsize=(12, 6), constrained_layout=True)
cm = ['gnuplot2'] * len(n_samples_)
for col in range(3):
for row in range(2):
ax = axs[row, col]
if col == 0 and row == 0:
pcm = ax.contourf(mean_mcs[0], levels=np.linspace(-20, 20, 16), cmap=cm[col])
plt.colorbar(pcm, ax=ax)
if col == 1 and row == 0:
pcm = ax.contourf(mean_recon[0], levels=np.linspace(-20, 20, 16), cmap=cm[col])
plt.colorbar(pcm, ax=ax)
if col == 2 and row == 0:
pcm = ax.contourf(mean_diff[0], levels=np.linspace(0, 0.03, 16), cmap=cm[col])
plt.colorbar(pcm, ax=ax, format='%.0e')
if col == 0 and row == 1:
pcm = ax.contourf(var_mcs[0], levels=np.linspace(0, 8, 16), cmap=cm[col])
plt.colorbar(pcm, ax=ax)
if col == 1 and row == 1:
pcm = ax.contourf(var_recon[0], levels=np.linspace(0, 8, 16), cmap=cm[col])
plt.colorbar(pcm, ax=ax)
if col == 2 and row == 1:
pcm = ax.contourf(var_diff[0], levels=np.linspace(0, 0.03, 16), cmap=cm[col])
plt.colorbar(pcm, ax=ax, format='%.0e')
if col == 0:
ax.set_title('MCS')
if col == 0 and row == 0:
ax.set_ylabel('Mean \n y')
if col == 0 and row == 1:
ax.set_ylabel('Variance \n y')
if col == 1:
ax.set_title('GDMaps PCE')
if col == 2:
ax.set_title('Relative error')
ax.set_xlabel('x')
plt.savefig('figures/Example1/Moment-estimation.png', bbox_inches='tight', dpi=500)
```
| github_jupyter |
# R Serving with Plumber
## Dockerfile
* The Dockerfile defines the environment in which our server will be executed.
* Below, you can see that the entrypoint for our container will be [deploy.R](deploy.R)
```
%pycat Dockerfile
```
## Code: deploy.R
The **deploy.R** script handles the following steps:
* Loads the R libraries used by the server.
* Loads a pretrained `xgboost` model that has been trained on the classical [Iris](https://archive.ics.uci.edu/ml/datasets/iris) dataset.
* Dua, D. and Graff, C. (2019). UCI Machine Learning Repository [http://archive.ics.uci.edu/ml]. Irvine, CA: University of California, School of Information and Computer Science.
* Defines an inference function that takes a matrix of iris features and returns predictions for those iris examples.
* Finally, it imports the [endpoints.R](endpoints.R) script and launches the Plumber server app using those endpoint definitions.
```
%pycat deploy.R
```
## Code: endpoints.R
**endpoints.R** defines two routes:
* `/ping` returns a string 'Alive' to indicate that the application is healthy
* `/invocations` applies the previously defined inference function to the input features from the request body
For more information about the requirements for building your own inference container, see:
[Use Your Own Inference Code with Hosting Services](https://docs.aws.amazon.com/sagemaker/latest/dg/your-algorithms-inference-code.html)
```
%pycat endpoints.R
```
## Build the Serving Image
```
!docker build -t r-plumber .
```
## Launch the Serving Container
```
!echo "Launching Plumber"
!docker run -d --rm -p 5000:8080 r-plumber
!echo "Waiting for the server to start.." && sleep 10
!docker container list
```
## Define Simple Python Client
```
import requests
from tqdm import tqdm
import pandas as pd
pd.set_option("display.max_rows", 500)
def get_predictions(examples, instance=requests, port=5000):
payload = {"features": examples}
return instance.post(f"http://127.0.0.1:{port}/invocations", json=payload)
def get_health(instance=requests, port=5000):
instance.get(f"http://127.0.0.1:{port}/ping")
```
## Define Example Inputs
Let's define example inputs from the Iris dataset.
```
column_names = ["Sepal.Length", "Sepal.Width", "Petal.Length", "Petal.Width", "Label"]
iris = pd.read_csv(
"s3://sagemaker-sample-files/datasets/tabular/iris/iris.data", names=column_names
)
iris_features = iris[["Sepal.Length", "Sepal.Width", "Petal.Length", "Petal.Width"]]
example_inputs = iris_features.values.tolist()
```
### Plumber
```
predicted = get_predictions(example_inputs).json()["output"]
iris["predicted"] = predicted
iris
```
### Stop All Serving Containers
Finally, we will shut down the serving container we launched for the test.
```
!docker kill $(docker ps -q)
```
| github_jupyter |
# Analytical formulation of the synaptic output using a bernoulli and a gaussian process.
We can think of individual synapses as product of 2 random variables. 1: Release probabilities (p) as a bernoulli process (coin flips with certain probability) and 2: synaptic weights drawn from a (say lognormal) distribution. This results in a unitary at the postsynapse. These unitaries can then sum up to give rise to PSPs, which are sums of these random variables.
Formally,
$X = B(p)$
$Y = G(\bar{w}, \tilde{w})$ (This can be arbitrary distributions, not necessarily Gaussian)
Here,
$B$ is Binomial random process with success probability $p$,
$G$ is a Gaussian random process with mean $\bar{w}$, and variance $\tilde{w}$
Now $V(X,Y) = {E(X)}^2 V(Y) + {E(Y)}^2 V(X) + V(X)V(Y)$
Or,
$V(X,Y) = \bar{w}^2.p.(1-p) + p^2.\tilde{w} + p.(1-p).\tilde{w}$
Canceling and rearranging,
$V(X,Y) = p.(\bar{w}^2.(1- p) + \tilde{w})$
Let the number of active synapses by n. Then, the total variance of n such processes would be:
$\sum\limits_{}^{n} {V(X,Y)} = np.(\bar{w}^2.(1- p) + \tilde{w})$
Similarly, mean EPSC from n active synapses will be $\sum\limits_{}^{n} {E(X,Y)} = \sum\limits_{}^{n}{E(X).E(Y)} = \sum\limits_{}^{n} p.\bar{w} = np\bar{w}$
Using this framework, we can think about if there were to be tuning of weights, what signatures should we find in the data. For example, the fano factor turns out immediately to be independent of n (number of synapses) =
$\frac{V(X,Y)}{E(X,Y)} = \frac{np.(\bar{w}^2.(1- p) + \tilde{w})}{np\bar{w}} = (1-p)\bar{w} + \frac{\tilde{w}}{\bar{w}}$
We can rewrite this now for both excitatory and inhibitory inputs.
$FF_e = \frac{V_e(X,Y)}{E_e(X,Y)} = \frac{n_e p_e.(\bar{w_e}^2.(1- p_e) + \tilde{w_e})}{n_e p_e \bar{w_e}} = (1-p_e)\bar{w_e} + \frac{\tilde{w_e}}{\bar{w_e}}$
And for inhibitory, again,
$FF_i = \frac{V_i(X,Y)}{E_i(X,Y)} = (1-p_i)\bar{w_i} + \frac{\tilde{w_i}}{\bar{w_i}}$
Therefore, without the numbers of afferents in, the fano factors of E and I inputs should not scale with each other, unless there is weight tuning.
Let's assess what $\frac{FF_i}{FF_e}$ gives us with and without weight tuning:
$ \frac{(1-p_i)\bar{w_i} + \frac{\tilde{w_i}}{\bar{w_i}}}{(1-p_e)\bar{w_e} + \frac{\tilde{w_e}}{\bar{w_e}}}$
First, we define a variable to quantify tuning error, such that it parameterizes the system from a global balanced state to a detailed balanced state. Let this variable be $\rho$.
We define detailed balanced as a state where all synaptic weights are tuned to a given I/E ratio, as opposed to a global balanced state, with only average synaptic weights balanced. That is, for detailed balance $\vec{W_i} = k.\vec{W_e} $, whereas for global balance, only $\bar{W_i} = k.\bar{W_e}$
So now, to go from perfect correlation to no correlation between 2 standard normal vectors $x$ and $y$, we use the above defined $\rho$, such that $y = \rho.x + \mathcal{N}(0, 1-\rho^2)$. Here, at $\rho=1$, there is perfect correlation, and at $\rho=0$, there is no correlation.
From this, we can build weight vector $W_i$ from $W_e$, by converting the standard normal $x$ into $W_e$ and $y$ into $W_i$.
We start with $\frac{w_e - \bar{w_e}}{\sqrt{\tilde{w_e}}} \times \rho + \mathcal{N}(0, 1-\rho^2)$
By scaling and adding an intercept to it,
$w_i = k \times \bigg\{ \big( \frac{w_e - \bar{w_e}}{\sqrt{\tilde{w_e}}} \times \rho + \mathcal{N}(0, 1-\rho^2) \big) \times \sqrt{\tilde{w_e}} + \bar{w_e}\bigg\}$
And so for the whole vector,
$\vec{W_i} = k \bigg\{ \big( \frac{\vec{W_e} - \bar{W_e}}{\sqrt{\tilde{W_e}}} \times \rho + \mathcal{N}(0, 1-\rho^2) \big) \times \sqrt{\tilde{W_e}} + \bar{W_e}\bigg\}$
This can be simplified to:
$\vec{W_i} = k \times \bigg\{ \vec{W_e}.\rho + \bar{W_e}.(1-\rho) + \mathcal{N}(0, \tilde{W_e}.(1-\rho^2)) \bigg\}$
This can now be looked at in cases of perfect weight tuning, and without weight tuning:
At $\rho=0$, $\vec{W_i} = k.\bar{W_e} + \mathcal{N}(0, k^2\tilde{W_e})$, which implies global balance and untuned weights.
At $\rho=1$, $\vec{W_i} = k \vec{W_e}$, which implies detailed balance and perfectly tuned weights.
For $ 0 < \rho < 1$, $\rho$ denotes the correlation between the inhibitory and excitatory synaptic weights.
From here, it can be noted that $\bar{W_i} = k.\bar{W_e}$ and $\tilde{W_i} = k^2\tilde{W_e}$
Now going back to Fano factor calculations,
$FF_i = (1-p_i)\bar{W_i} + \frac{\tilde{W_i}}{\bar{W_i}} = k\times \big((1-p_i)(\bar{W_e}) + \frac{\tilde{W_e}}{ \bar{W_e}}\big)$
and $FF_e = (1-p_e)\bar{W_e} + \frac{\tilde{W_e}}{\bar{W_e}} $
Therefore, $\frac{FF_i}{FF_e} = k\times {\frac{(1-p_i)\bar{W_e} + \frac{\tilde{W_e}}{\bar{W_e}}}{(1-p_e)\bar{W_e} + \frac{\tilde{W_e}}{\bar{W_e}}}}$
If $\tilde{W_E} >> \bar{W_E}, \frac{FF_i}{FF_e} \to k$
$FF_i = (1-p_i)\bar{W_i} + \frac{\tilde{W_i}}{\bar{W_i}} = k\times \big((1-p_i)(\bar{W_e}) + \frac{\tilde{W_e}}{ \bar{W_e}}\big)$
Therefore, $\frac{FF_i}{FF_e} = k\times {\frac{(1-p_i)\bar{W_e} + \frac{\tilde{W_e}}{\bar{W_e}}}{(1-p_e)\bar{W_e} + \frac{\tilde{W_e}}{\bar{W_e}}}}$
If $\tilde{W_E} >> \bar{W_E}, \frac{FF_i}{FF_e} \to k$
```
import sys
sys.path.append("../")
import glob
from mpl_toolkits.axes_grid1 import make_axes_locatable
import math
from matplotlib import mlab
from Linearity import Neuron
import numpy as np
import scipy.stats as ss
import matplotlib
import matplotlib.pyplot as plt
from matplotlib.font_manager import findfont, FontProperties
matplotlib.rcParams['text.usetex'] = False
matplotlib.rc('text.latex', preamble=r'\usepackage{cmbright}')
matplotlib.rc('font',**{'family':'sans-serif','sans-serif':['Arial']})
plt.style.use('neuron')
from pickle import dump
import random
def simpleaxis(axes, every=False, outward=False):
if not isinstance(axes, (list, np.ndarray)):
axes = [axes]
for ax in axes:
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
if (outward):
ax.spines['bottom'].set_position(('outward', 10))
ax.spines['left'].set_position(('outward', 10))
if every:
ax.spines['bottom'].set_visible(False)
ax.spines['left'].set_visible(False)
ax.get_xaxis().tick_bottom()
ax.get_yaxis().tick_left()
ax.set_title('')
n, p = 20, 0.2
mu , sigma = 1, 0.5
trials = 5
syn_weight = np.random.normal(mu,sigma,n)
var_arr = []
for num in range(1000):
syn_active = np.random.binomial(n,p,n)
total_weight = syn_active*syn_weight
var_arr.append(np.var(total_weight, ddof=1))
est_var = n*p*((1-p)*(mu**2) + sigma)
fig, ax = plt.subplots()
counts, bins, patches = ax.hist(var_arr)
ax.vlines(x = est_var, ymin=0, ymax = max(counts) +5 )
plt.show()
b,d = [], []
for j in range(1000):
a = np.random.binomial(1,0.2,50)*20
b.append(np.mean(a))
c = np.random.binomial(1,0.2,20)*50
d.append(np.mean(c))
fig, ax = plt.subplots()
n, bins, patches = ax.hist(b, bins=2)
plt.show()
fig, ax = plt.subplots()
n, bins, patches = ax.hist(d, bins=2)
plt.show()
np.mean(d), np.mean(b)
def simpleaxis(axes, every=False, outward=False):
if not isinstance(axes, (list, np.ndarray)):
axes = [axes]
for ax in axes:
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
if (outward):
ax.spines['bottom'].set_position(('outward', 10))
ax.spines['left'].set_position(('outward', 10))
if every:
ax.spines['bottom'].set_visible(False)
ax.spines['left'].set_visible(False)
ax.get_xaxis().tick_bottom()
ax.get_yaxis().tick_left()
ax.set_title('')
def generatePSC(w_e, w_i, p_E, p_I, trials=1):
'''Returns one PSC worth of input'''
W_E, W_I = [], []
numSyns_E = []
numSyns_I = []
for j in range(trials):
N_E = len(w_e)
N_I = len(w_i)
# Bernoulli process for release probability
active_e = np.random.binomial(1, p_E, N_E)
active_i = np.random.binomial(1, p_I, N_I)
W_E.append(active_e.dot(w_e))
W_I.append(active_i.dot(w_i))
numSyns_E.append(np.sum(active_e))
numSyns_I.append(np.sum(active_i))
return W_E, W_I, numSyns_E, numSyns_I
def getTunedInhibition(w_e, rho, k, minResolvable=None):
''' Another method'''
sorted_indices = np.argsort(w_e)
old_sorting = np.argsort(sorted_indices)
if not minResolvable:
minResolvable = min(w_e)
minDiff = np.max([np.min(np.diff(w_e[sorted_indices])), minResolvable])
bins = np.arange(w_e[sorted_indices[0]], w_e[sorted_indices[-1]]+minDiff, minDiff)
result = np.digitize(w_e[sorted_indices], bins)
L = len(bins)
shuffle_length = int((L - 1) * (1-rho) + 1)
shuffled_sorted_indices = []
for binIndex in range(1,L+1,shuffle_length):
if len(old_sorting):
shuffled_sorted_indices.append(np.random.permutation(sorted_indices[np.where((result >= binIndex) & (result < binIndex + shuffle_length))[0]]))
w_i = k* w_e[np.concatenate(shuffled_sorted_indices)]
w_i = w_i[old_sorting]
return w_i
# def getTunedInhibition(w_e, rho, k, normal='True'):
# '''Returns tuned inhibition with a given I/E ratio. Only for a normally distributed weight distribution. Can do it for general case.'''
# if normal:
# return k * (w_e*rho + np.mean(w_e)*(1-rho) + np.random.normal( 0, np.var(w_e)*(1-rho**2), len(w_e)) )
?random.sample([1,2,3,4],5)
```
## Getting correlation matrices for proportional synaptic numbers
```
def returnCorrelationMatrixFromSynapticInput(N_E=1000, N_I=1000, p_E=0.2, p_I=0.8, k=2, rho_steps=9,
var_steps=9, max_syn = 20, minResolvable = 0.5, num_syn = 50,
runs=50, numTrials=20, corrType='var'):
'''
N_E, N_I = Number of afferents
p_E, p_I = Release probs
k = I/E ratio for the unitary synaptic distribution.
rho_steps = samples of tuning parameter rho
max_syn = Number of active synapses
max_syn = Maximum number of active synapses
num_syn = Number of samples from active synapses
minResolvable = Minimum resolvable current for tuning
'''
# Normal synaptic weights, normal dist parameters. Keep the scale.
w_e_mean = -0.39 # shape
#var_arr = np.logspace(-1, 1, num=var_steps, base=2) #scale
var_arr = np.linspace(0.5, 1.0, num=var_steps) #scale
scalingWeightFactor = 1. #0.25
correlation_matrix = np.zeros((var_steps, rho_steps))
w_e_var_true = np.zeros((runs, var_steps))
w_e_mean_true = np.zeros((runs, var_steps))
w_ei_corr = np.zeros((var_steps, rho_steps))
rho_arr = np.linspace(0,1,rho_steps)
runs = 50
run_mat = np.zeros((var_steps, rho_steps))
for run in range(runs):
for i, w_e_var in enumerate(var_arr):
# Unitary synaptic distribution
w_e = np.random.lognormal(w_e_mean, w_e_var, N_E)/scalingWeightFactor
w_e_var_true[run,i] = np.var(w_e, ddof=1)
w_e_mean_true[run,i] = np.mean(w_e)
#w_e = np.random.normal(w_e_mean, w_e_var, N_E)
#w_e -= np.min(w_e)
# fig,ax = plt.subplots(ncols=rho_steps,sharex=True, sharey=True)
# fig2,ax2 = plt.subplots(ncols=rho_steps,sharex=True, sharey=True)
# fig3,ax3 = plt.subplots(ncols=rho_steps,sharex=True, sharey=True)
# fig4,ax4 = plt.subplots(ncols=rho_steps,sharex=True, sharey=True)
# fig5,ax5 = plt.subplots(ncols=rho_steps,sharex=True, sharey=True)
for j, rho in enumerate(rho_arr):
w_i = getTunedInhibition(w_e, rho, k, minResolvable)
#print(np.mean(w_e), np.var(w_e))
# print("Desired correlation: {}, actual correlation: {}".format(rho, np.corrcoef(w_e, w_i)[0,1]))
numTrials = 30
W_exc_mean, W_inh_mean = [], []
numE, numI = [], []
W_exc_std, W_inh_std = [], []
W_exc, W_inh = [], []
numSynpaseRange = np.random.choice(max_syn, num_syn) + 1
#numSynpaseRange = np.arange(max_syn) + 1
for n_e in numSynpaseRange:
chosen_synapses = random.sample(range(len(w_e)), n_e)
w_e_subset = w_e[chosen_synapses]
w_i_subset = w_i[chosen_synapses]
w_exc, w_inh, num_e, num_i = generatePSC(w_e_subset, w_i_subset, p_E, p_I, numTrials)
W_exc_mean.append(np.mean(w_exc))
W_inh_mean.append(np.mean(w_inh))
numE.append(np.mean(num_e))
numI.append(np.mean(num_i))
W_exc_std.append(np.std(w_exc, ddof=1))
W_inh_std.append(np.std(w_inh, ddof=1))
W_exc.append(w_exc)
W_inh.append(w_inh)
# W_exc = np.concatenate([W_exc])
# W_inh = np.concatenate([W_inh])
W_exc_std = np.array(W_exc_std)
W_inh_std = np.array(W_inh_std)
W_exc_mean = np.array(W_exc_mean)
W_inh_mean = np.array(W_inh_mean)
W_exc = np.concatenate(W_exc)
W_inh = np.concatenate(W_inh)
# print("Correlation for synaptic sum: {}".format(np.corrcoef(W_exc, W_inh)[0,1]))
# print("Correlation for synaptic variance: {}\n".format(np.corrcoef(W_exc_std**2, W_inh_std**2)[0,1]))
if corrType == 'var':
correlation_matrix[i,j] += np.corrcoef(W_exc_std**2, W_inh_std**2)[0,1]
elif corrType == 'rsquare':
slope, intercept, rval, pval, stderr = ss.linregress(W_exc_mean, W_inh_mean)
correlation_matrix[i,j] += rval**2
coeff_wts = np.corrcoef(w_exc, w_inh)[0,1]
w_ei_corr[i,j] += np.corrcoef(w_e, w_i)[0,1]
numE = np.array(numE)
# ax[j].scatter(W_exc, W_inh, c=numE, cmap = plt.cm.viridis)
# ax[j].errorbar(W_exc, W_inh, xerr=W_exc_std, yerr=W_inh_std, fmt=None, marker=None, mew=0)
# ax2[j].scatter(W_exc, W_inh_std**2, c=numE, cmap = plt.cm.viridis)
# ax3[j].scatter(W_inh, W_exc_std**2, c=numI, cmap = plt.cm.viridis)
# ax4[j].scatter(np.log10(W_exc_std**2), np.log10(W_inh_std**2), c=numE, cmap = plt.cm.viridis)
# ax5[j].scatter(W_exc_std**2/W_exc, W_inh_std**2/W_inh)
# ax4[j].set_xlim(-5,10.)
# ax4[j].set_ylim(-5,10.)
# max_ax = np.max([ax[0].get_xlim()[1], ax[0].get_ylim()[1]])
# for axis in fig.axes:
# axis.set_xlim(xmax=max_ax)
# axis.set_ylim(ymax=max_ax)
# figs = [fig, fig2, fig3, fig4, fig5]
# axes = [ax, ax2, ax3, ax4, ax5]
# for figure, axis in zip(figs, axes):
# simpleaxis(axis)
# figure.set_figwidth(len(rho_arr)*figure.get_figheight())
# plt.show()
correlation_matrix/=runs
w_ei_corr/=runs
w_e_var_true = np.mean(w_e_var_true, axis=0)
w_e_mean_true = np.mean(w_e_mean_true, axis=0)
w_ei_corr = np.average(w_ei_corr,axis=0)
return (correlation_matrix, w_e_var_true, w_e_mean_true, w_ei_corr)
```
## Getting correlation matrices for varying proportion of synaptic numbers
```
def returnCorrelationMatrixFromSynapticInputNumNotProportional(N_E=1000, N_I=1000, p_E=0.2, p_I=0.8, k=2, rho=1, sigma_steps=9,
var_steps=9, max_syn = 20, minResolvable = 0.5, num_syn = 50,
runs=50, numTrials=20, corrType='var'):
'''
N_E, N_I = Number of afferents
p_E, p_I = Release probs
k = I/E ratio for the unitary synaptic distribution.
rho = Degree of tuning
max_syn = Number of active synapses
max_syn = Maximum number of active synapses
num_syn = Number of samples from active synapses
minResolvable = Minimum resolvable current for tuning
'''
# Normal synaptic weights, normal dist parameters. Keep the scale.
w_e_mean = -0.39 # shape
#var_arr = np.logspace(-1, 1, num=var_steps, base=2) #scale
var_arr = np.linspace(0.5, 1.0, num=var_steps) #scale
scalingWeightFactor = 1. #0.25
correlation_matrix = np.zeros((var_steps, sigma_steps))
w_e_var_true = np.zeros((runs, var_steps))
w_e_mean_true = np.zeros((runs, var_steps))
w_ei_corr = np.zeros((var_steps, sigma_steps))
num_ei_corr = np.zeros((var_steps, sigma_steps))
runs = 50
run_mat = np.zeros((var_steps, sigma_steps))
sigma_arr = np.linspace(0,1,sigma_steps)
for run in range(runs):
for i, w_e_var in enumerate(var_arr):
# Unitary synaptic distribution
w_e = np.random.lognormal(w_e_mean, w_e_var, N_E)/scalingWeightFactor
w_e_var_true[run,i] = np.var(w_e, ddof=1)
w_e_mean_true[run,i] = np.mean(w_e)
#w_e = np.random.normal(w_e_mean, w_e_var, N_E)
#w_e -= np.min(w_e)
# fig,ax = plt.subplots(ncols=rho_steps,sharex=True, sharey=True)
# fig2,ax2 = plt.subplots(ncols=rho_steps,sharex=True, sharey=True)
# fig3,ax3 = plt.subplots(ncols=rho_steps,sharex=True, sharey=True)
# fig4,ax4 = plt.subplots(ncols=rho_steps,sharex=True, sharey=True)
# fig5,ax5 = plt.subplots(ncols=rho_steps,sharex=True, sharey=True)
w_i = getTunedInhibition(w_e, rho, k, minResolvable)
#print(np.mean(w_e), np.var(w_e))
# print("Desired correlation: {}, actual correlation: {}".format(rho, np.corrcoef(w_e, w_i)[0,1]))
numTrials = 30
#numSynpaseRange = np.random.choice(max_syn, num_syn) + 1
#numSynpaseRange = np.arange(max_syn) + 1
n_exc_arr = np.arange(max_syn) + 1
for j,sigma in enumerate(sigma_arr):
W_exc_mean, W_inh_mean = [], []
numE, numI = [], []
W_exc_std, W_inh_std = [], []
W_exc, W_inh = [], []
n_inh_arr = getTunedInhibition(n_exc_arr, sigma, 1, minResolvable=1)
num_ei_corr[i,j] += np.corrcoef(n_exc_arr, n_inh_arr)[0,1]
for n_e, n_i in zip(n_exc_arr, n_inh_arr):
chosen_synapses_e = random.sample(range(len(w_e)), n_e)
chosen_synapses_i = random.sample(range(len(w_i)), n_i)
w_e_subset = w_e[chosen_synapses_e]
w_i_subset = w_i[chosen_synapses_i]
w_exc, w_inh, num_e, num_i = generatePSC(w_e_subset, w_i_subset, p_E, p_I, numTrials)
W_exc_mean.append(np.mean(w_exc))
W_inh_mean.append(np.mean(w_inh))
numE.append(np.mean(num_e))
numI.append(np.mean(num_i))
W_exc_std.append(np.std(w_exc, ddof=1))
W_inh_std.append(np.std(w_inh, ddof=1))
W_exc.append(w_exc)
W_inh.append(w_inh)
# W_exc = np.concatenate([W_exc])
# W_inh = np.concatenate([W_inh])
W_exc_std = np.array(W_exc_std)
W_inh_std = np.array(W_inh_std)
W_exc_mean = np.array(W_exc_mean)
W_inh_mean = np.array(W_inh_mean)
W_exc = np.concatenate(W_exc)
W_inh = np.concatenate(W_inh)
# print("Correlation for synaptic sum: {}".format(np.corrcoef(W_exc, W_inh)[0,1]))
# print("Correlation for synaptic variance: {}\n".format(np.corrcoef(W_exc_std**2, W_inh_std**2)[0,1]))
if corrType == 'var':
correlation_matrix[i,j] += np.corrcoef(W_exc_std**2, W_inh_std**2)[0,1]
elif corrType == 'rsquare':
slope, intercept, rval, pval, stderr = ss.linregress(W_exc_mean, W_inh_mean)
correlation_matrix[i,j] += rval**2
coeff_wts = np.corrcoef(w_exc, w_inh)[0,1]
w_ei_corr[i,j] += np.corrcoef(w_e, w_i)[0,1]
numE = np.array(numE)
# ax[j].scatter(W_exc, W_inh, c=numE, cmap = plt.cm.viridis)
# ax[j].errorbar(W_exc, W_inh, xerr=W_exc_std, yerr=W_inh_std, fmt=None, marker=None, mew=0)
# ax2[j].scatter(W_exc, W_inh_std**2, c=numE, cmap = plt.cm.viridis)
# ax3[j].scatter(W_inh, W_exc_std**2, c=numI, cmap = plt.cm.viridis)
# ax4[j].scatter(np.log10(W_exc_std**2), np.log10(W_inh_std**2), c=numE, cmap = plt.cm.viridis)
# ax5[j].scatter(W_exc_std**2/W_exc, W_inh_std**2/W_inh)
# ax4[j].set_xlim(-5,10.)
# ax4[j].set_ylim(-5,10.)
# max_ax = np.max([ax[0].get_xlim()[1], ax[0].get_ylim()[1]])
# for axis in fig.axes:
# axis.set_xlim(xmax=max_ax)
# axis.set_ylim(ymax=max_ax)
# figs = [fig, fig2, fig3, fig4, fig5]
# axes = [ax, ax2, ax3, ax4, ax5]
# for figure, axis in zip(figs, axes):
# simpleaxis(axis)
# figure.set_figwidth(len(rho_arr)*figure.get_figheight())
# plt.show()
correlation_matrix/=runs
w_ei_corr/=runs
num_ei_corr/=runs
w_e_var_true = np.mean(w_e_var_true, axis=0)
w_e_mean_true = np.mean(w_e_mean_true, axis=0)
w_ei_corr = np.average(w_ei_corr,axis=0)
num_ei_corr = np.average(num_ei_corr,axis=0)
return (correlation_matrix, w_e_var_true, w_e_mean_true, w_ei_corr, num_ei_corr)
def drawHeatmapCorrelationMat(correlation_matrix, rho_arr, var_arr, colorNormalize='max'):
fig, ax = plt.subplots()
if colorNormalize == 'max':
vmax = np.max(correlation_matrix)
else:
vmax = 1.
cbar = ax.imshow(correlation_matrix, cmap='magma',origin='lower', vmin=0., vmax=vmax)
ax.set_xlabel("$\\rho$")
ax.set_ylabel("$\\sigma^2_{exc}/\\mu_{exc}$")
ax.set_xticks(range(0,len(rho_arr), 4))
ax.set_yticks(range(0,len(var_arr), 4))
ax.set_xticklabels(np.round(rho_arr,2)[::4])
ax.set_yticklabels(np.round(var_arr,2)[::4])
plt.colorbar(cbar, label="avg. correlation")
plt.show()
N_E = 1000
w_e_mean = 0.2
w_e_var = 1.5
w_e = np.random.lognormal(w_e_mean, w_e_var, N_E)
rho_arr = np.arange(0,1,0.02)
corr_arr = []
k = 1
for rho in rho_arr:
w_i= getTunedInhibition(w_e, rho, k)
corr_arr.append(np.corrcoef(w_e, w_i)[0,1])
fig, ax = plt.subplots()
ax.scatter(rho_arr, corr_arr)
ax.plot([0,1], [0,1], '--')
# ax.plot(w_e[sorted_indices]/w_e[sorted_indices][-1], np.cumsum(result)/np.cumsum(result)[-1])
simpleaxis(ax)
plt.show()
```
## Wide number distribution, no tuning
```
correlation_matrix_num, W_e_var, W_e_mean_true, true_corr_array, num_corr_array = returnCorrelationMatrixFromSynapticInputNumNotProportional(max_syn=20, corrType='rsquare')
drawHeatmapCorrelationMat(correlation_matrix_num, num_corr_array, W_e_var/W_e_mean_true)
```
## Putting everything together
```
class Neuron:
def __init__(self, w_e, k, N_I, p_E=0.2, p_I=0.8):
self.w_e = w_e
self.N_E = len(w_e)
self.w_i = []
self.N_I = N_I
self.p_E = p_E
self.p_I = p_I
self.k = k
self.stimuli = {}
def generateStimuli(self, n_e, sigma, k=1):
''' Generates a given number of stimuli'''
self.n_e = n_e
self.numStimuli = len(n_e)
self.n_i = self.getTunedNumbers(n_e, sigma, k=k)
for j, (num_e, num_i) in enumerate(zip(self.n_e, self.n_i)):
self.stimuli[j] = self.generateStimulus(num_e, num_i)
def generateStimulus(self, n_e, n_i):
''' Generates a stimulus based on number of exc and inh'''
if not n_e==n_i:
numSyn = max([n_e, n_i])
else:
numSyn = n_e
totalSyns = min([self.N_E, self.N_I])
chosen_synapses = random.sample(range(totalSyns), numSyn)
chosen_synapses_e = chosen_synapses[:n_e]
chosen_synapses_i = chosen_synapses[:n_i]
return (chosen_synapses_e, chosen_synapses_i)
def giveStimuli(self, numTrials):
''' Gives all the stimuli to the neuron '''
w_e_all, w_i_all, num_e_all , num_i_all = {}, {}, {}, {}
for j in range(self.numStimuli):
w_exc, w_inh, num_e, num_i = self.giveStimulus(j, numTrials)
w_e_all [j] = w_exc
w_i_all [j] = w_inh
num_e_all[j] = num_e
num_i_all[j] = num_i
return (w_e_all, w_i_all, num_e_all, num_i_all)
def giveStimulus(self, index, numTrials=1):
chosen_synapses_e, chosen_synapses_i = self.stimuli[index]
w_e_subset = self.w_e[chosen_synapses_e]
w_i_subset = self.w_i[chosen_synapses_i]
w_exc, w_inh, num_e, num_i = self.generatePSC(w_e_subset, w_i_subset, numTrials)
return (w_exc, w_inh, num_e, num_i)
def generatePSC(self, w_e_subset, w_i_subset, numTrials=1):
'''Returns one PSC worth of input'''
W_E, W_I = [], []
numSyns_E = []
numSyns_I = []
n_e = len(w_e_subset)
n_i = len(w_i_subset)
for j in range(numTrials):
# Bernoulli process for release probability
active_e = np.random.binomial(1, self.p_E, n_e)
active_i = np.random.binomial(1, self.p_I, n_i)
W_E.append(active_e.dot(w_e_subset))
W_I.append(active_i.dot(w_i_subset))
numSyns_E.append(np.sum(active_e))
numSyns_I.append(np.sum(active_i))
return W_E, W_I, numSyns_E, numSyns_I
def returnCorrelation(arr1, arr2):
''' Returns correlations'''
return np.corrcoef(arr1, arr2)[0,1]
def __prob_round__(self, x):
is_up = random.random() < x-int(x)
round_func = math.ceil if is_up else math.floor
return int(round_func(x))
def getTunedNumbers(self, num_e, sigma, k=1, minResolvable=1, probRounding=False):
''' Returns tuned inhibitory weights from excitatory weights and a tuning parameter. Minresolvable is used to bin the weight vector'''
sorted_indices = np.argsort(num_e)
old_sorting = np.argsort(sorted_indices)
if not minResolvable:
minResolvable = min(num_e)
minDiff = np.max([np.min(np.diff(num_e[sorted_indices])), minResolvable])
bins = np.arange(num_e[sorted_indices[0]], num_e[sorted_indices[-1]]+minDiff, minDiff)
result = np.digitize(num_e[sorted_indices], bins)
L = len(bins)
shuffle_length = int((L - 1) * (1-sigma) + 1)
shuffled_sorted_indices = []
for binIndex in range(1,L+1,shuffle_length):
if len(old_sorting):
shuffled_sorted_indices.append(np.random.permutation(sorted_indices[np.where((result >= binIndex) & (result < binIndex + shuffle_length))[0]]))
if probRounding:
num_i = np.array([self.__prob_round__(k*exc) for exc in num_e[np.concatenate(shuffled_sorted_indices)]])
else:
num_i = np.array([int(np.round(k*exc)) for exc in num_e[np.concatenate(shuffled_sorted_indices)]])
num_i = num_i[old_sorting]
return num_i
def getTunedWeights(self, rho, k=1, minResolvable=0.1):
''' Returns tuned inhibitory weights from excitatory weights and a tuning parameter. Minresolvable is used to bin the weight vector'''
sorted_indices = np.argsort(self.w_e)
old_sorting = np.argsort(sorted_indices)
if not minResolvable:
minResolvable = min(self.w_e)
minDiff = np.max([np.min(np.diff(self.w_e[sorted_indices])), minResolvable])
bins = np.arange(self.w_e[sorted_indices[0]], self.w_e[sorted_indices[-1]]+minDiff, minDiff)
result = np.digitize(self.w_e[sorted_indices], bins)
L = len(bins)
shuffle_length = int((L - 1) * (1-rho) + 1)
shuffled_sorted_indices = []
for binIndex in range(1,L+1,shuffle_length):
if len(old_sorting):
shuffled_sorted_indices.append(np.random.permutation(sorted_indices[np.where((result >= binIndex) & (result < binIndex + shuffle_length))[0]]))
w_i = k*self.w_e[np.concatenate(shuffled_sorted_indices)]
w_i = w_i[old_sorting]
return w_i
def getAveragedStimuli(w_e_all, w_i_all):
avg_e, avg_i = [], []
std_e, std_i = [], []
for j in range(num_stim):
avg_e.append(np.average(w_e_all[j]))
avg_i.append(np.average(w_i_all[j]))
std_e.append(np.std(w_e_all[j], ddof=1))
std_i.append(np.std(w_i_all[j], ddof=1))
avg_e = np.array(avg_e)
avg_i = np.array(avg_i)
std_e = np.array(std_e)
std_i = np.array(std_i)
return avg_e, avg_i, std_e, std_i
def scatterPlots(w_e_all, w_i_all, filename='', show=True):
fig, (ax1, ax2) = plt.subplots(ncols=2)
avg_e, avg_i, std_e, std_i = getAveragedStimuli(w_e_all, w_i_all)
ax1.errorbar(avg_e, avg_i, xerr = std_e, yerr = std_i, fmt='o', markersize=6)
slope, intercept, r_value, p_value, std_err = ss.linregress(avg_e, avg_i)
ax2.scatter(std_e, std_i, s=24)
max_ax = max([ax1.get_xlim()[1], ax1.get_ylim()[1]])
ax1.set_xlim(xmin=0., xmax=max_ax)
ax1.set_ylim(ymin=0., ymax=max_ax)
xaxis = np.linspace(ax1.get_xlim()[0], ax1.get_xlim()[1], 100)
ax1.plot(xaxis, slope*xaxis + intercept, '--', label="$R^2$ {:.2f}\n slope {:.2f}".format(r_value**2, slope))
max_ax = max([ax2.get_xlim()[1], ax2.get_ylim()[1]])
ax2.set_xlim(xmin=0., xmax=max_ax)
ax2.set_ylim(ymin=0., ymax=max_ax)
slope, intercept, r_value, p_value, std_err = ss.linregress(std_e, std_i)
xaxis = np.linspace(ax2.get_xlim()[0], ax2.get_xlim()[1], 100)
ax2.plot(xaxis, slope*xaxis + intercept, '--', label="$R^2$ {:.2f}\n slope {:.2f}\nCorr {:.2f}".format(r_value**2, slope, np.corrcoef(std_e, std_i)[0,1]))
ax1.set_xlabel("Exc")
ax1.set_ylabel("Inh")
ax1.legend()
ax2.set_xlabel("$\sigma$(Exc)")
ax2.set_ylabel("$\sigma$(Inh)")
ax2.legend()
simpleaxis([ax1, ax2])
fig.set_figheight(1.5)
fig.set_figwidth(3)
if filename:
plt.savefig(filename)
if show==True:
plt.show()
plt.close()
return avg_e, avg_i, std_e, std_i
def histogram(w_e, xlabel='', filename='', show=True):
fig, ax = plt.subplots()
ax.hist(w_e, color='gray')
ax.set_xlabel(xlabel)
ax.set_ylabel("#")
simpleaxis(ax)
fig.set_figheight(1.5)
fig.set_figwidth(1.5)
if filename:
plt.savefig(filename)
if show:
plt.show()
plt.close()
def drawHeatmapCorrelationMat(correlation_matrix, x_arr, y_arr, xlabel = '', ylabel = '', cbarlabel= '', colorNormalize='max', filename = ''):
fig, ax = plt.subplots()
if colorNormalize == 'max':
vmin = 0.
vmax = np.max(correlation_matrix)
elif colorNormalize == 'minmax':
vmin = np.min(correlation_matrix)
print(vmin)
vmax = np.max(correlation_matrix)
else:
vmin = 0.
vmax = 1.
heatmap = ax.imshow(correlation_matrix, cmap='plasma',origin='lower', vmin=vmin, vmax=vmax)
ax.set_xlabel(xlabel)
ax.set_ylabel(ylabel)
ax.set_xticks(range(0,len(x_arr), 4))
ax.set_yticks(range(0,len(y_arr), 4))
ax.set_xticklabels(np.round(x_arr,2)[::4])
ax.set_yticklabels(np.round(y_arr,2)[::4])
cbar = plt.colorbar(heatmap, label=cbarlabel, fraction=0.046, pad=0.04)
cbar.set_ticks(np.linspace(0., vmax, 3))
cbar.set_ticklabels(['{:.2f}'.format(ticklabel) for ticklabel in np.linspace(0., vmax, 3)])
fig.set_figheight(1.5)
fig.set_figwidth(1.5)
if filename:
plt.savefig(filename)
plt.show()
dirPrefix = "../DamnBalance/Figures/"
plotFormat = ".svg"
```
## Case of detailed balance: $\rho = 1$
```
w_e_shape = -0.39
w_e_scale = [0.80] # np.linspace(0.5, 1.0, 10)
N_E, N_I = 1000, 1000
num_stim = 15
lam = 5
rho = 1.
sigma = 1.
w_k = 1.
n_k = 1.
w_e = np.random.lognormal(w_e_shape, w_e_scale[0], N_E)
n_e = np.random.poisson(lam, num_stim)
p_E, p_I = .2, .8
n1 = Neuron(w_e, w_k, N_I, p_E=p_E, p_I=p_I)
n1.w_i = n1.getTunedWeights(rho=rho, k=w_k)
n1.generateStimuli(n_e, sigma=sigma, k=n_k)
w_e_all, w_i_all, num_e_all, num_i_all = n1.giveStimuli(numTrials=6)
histogram(w_e, "Exc", filename=dirPrefix + "w_e_hist" + plotFormat)
histogram(n1.w_i, "Inh", filename=dirPrefix + "w_i_hist" + plotFormat)
histogram(n_e, "# syn./stim.", filename=dirPrefix + "n_e_hist" + plotFormat)
avg_e, avg_i, std_e, std_i = scatterPlots(w_e_all, w_i_all, filename=dirPrefix + "EI_scatter" + plotFormat)
```
## Case of global balance: $\rho = 0$
```
w_e_shape = -0.39
w_e_scale = [0.80] # np.linspace(0.5, 1.0, 10)
N_E, N_I = 1000, 1000
num_stim = 15
lam = 5
rho = 0.
sigma = 1.
w_k = 1.
n_k = 1.
w_e = np.random.lognormal(w_e_shape, w_e_scale[0], N_E)
n_e = np.random.poisson(lam, num_stim)
p_E, p_I = .2, .8
n1 = Neuron(w_e, w_k, N_I, p_E=p_E, p_I=p_I)
n1.w_i = n1.getTunedWeights(rho=rho, k=w_k)
n1.generateStimuli(n_e, sigma=sigma, k=n_k)
w_e_all, w_i_all, num_e_all, num_i_all = n1.giveStimuli(numTrials=6)
histogram(w_e, "Exc", filename=dirPrefix + "w_e_hist" + plotFormat)
histogram(n1.w_i, "Inh", filename=dirPrefix + "w_i_hist" + plotFormat)
histogram(n_e, "# syn./stim.", filename=dirPrefix + "n_e_hist" + plotFormat)
avg_e_0, avg_i_0, std_e_0, std_i_0 = scatterPlots(w_e_all, w_i_all, filename=dirPrefix + "EI_scatter" + plotFormat)
```
## EI correlation plots
```
fig, ax = plt.subplots()
plotline1, caplines1, barlinecols1 = ax.errorbar(np.arange(len(avg_e_0)), avg_e_0, yerr = std_e_0 , markersize=6, label="Exc",c='olive', fmt='.-', uplims=True)
plotline2, caplines2, barlinecols2 = ax.errorbar(1+len(avg_e_0) + np.arange(len(avg_e)), avg_e, yerr = std_e , markersize=6, label="Exc",c='olive', fmt='.-', uplims=True)
# line_exc3 = ax.plot(len(exc_1) + len(exc_2) + np.arange(len(exc_3)), exc_3, '.-', markersize=6 ,c=color_sqr[3])
# ax.xaxis.set_ticks([1,3,5,7,9])
ax_copy = ax.twinx()
plotline3, caplines3, barlinecols3 = ax_copy.errorbar( np.arange(len(avg_i_0)), avg_i_0, yerr = std_i_0, markersize=6, label="Inh",c='#8b0046', fmt='.-', lolims=True)
plotline4, caplines4, barlinecols4 = ax_copy.errorbar(1+len(avg_i_0) + np.arange(len(avg_i)), avg_i, yerr = std_i, markersize=6, label="Inh",c='#8b0046', fmt='.-', lolims=True)
# line_inh3 = ax_copy.plot( len(inh_1) + len(inh_2) + np.arange(len(inh_3)), inh_3, 'v-', markersize=6 ,c=color_sqr[3])
ax.set_title("Different number of squares show consistent EI ratios")
ax.set_xlabel("# stimulus")
caplines1[0].set_marker('_')
caplines1[0].set_markersize(5)
caplines2[0].set_marker('_')
caplines2[0].set_markersize(5)
caplines3[0].set_marker('_')
caplines3[0].set_markersize(5)
caplines4[0].set_marker('_')
caplines4[0].set_markersize(5)
ax.spines['left'].set_color('olive')
ax_copy.spines['right'].set_color('#8b0046')
# lines = line_exc1 + line_exc2 + line_inh1 + line_inh2
# labs = [l.get_label() for l in lines]
# ax.legend(lines, labs, loc=0)
simpleaxis(ax)
simpleaxis(ax_copy,every=True)
ax_copy.spines['right'].set_visible(True)
ax_copy.get_yaxis().tick_right()
ax.set_ylim(ymin=-0.5, ymax=3.)
ax_copy.set_ylim(ymin=-0.5, ymax=14.)
ax.set_yticks(np.linspace(0,3.,3))
ax_copy.set_yticks(np.linspace(0,14.,3))
ax.set_ylabel("Excitation")
ax_copy.set_ylabel("Inhibition")
fig.set_figheight(1.5)
fig.set_figwidth(3)
print(ax.get_ylim())
print(ax_copy.get_ylim())
# dump(fig,file('figures/fig2/2_precise_small.pkl','wb'))
plt.savefig('/home/bhalla/Documents/Codes/linearity/DamnBalance/Figures/' + '2_rho_range' + '.svg')
plt.show()
```
## EI Std. correlation plots
```
### Correlation plots for standard deviation exc and inh
plotlines, caplines, barlinecols = [], [], []
fig, ax = plt.subplots()
plotline1 = ax.plot(np.arange(len(std_e_0)), std_e_0, '.-', markersize=6, label="Exc",c='olive')
plotline2 = ax.plot(5 + len(std_e_0) + np.arange(len(std_e)), std_e, '.-', markersize=6, label="Exc",c='olive')
ax_copy = ax.twinx()
plotline3 = ax_copy.plot(np.arange(len(std_i_0)), std_i_0, '.-', markersize=6, label="Inh",c='#8b0046')
plotline4 = ax_copy.plot(5 + len(std_i_0) + np.arange(len(std_i)), std_i, '.-', markersize=6, label="Inh",c='#8b0046')
ax.set_xlabel("# stimulus")
ax.spines['left'].set_color('olive')
ax_copy.spines['right'].set_color('#8b0046')
# lines = line_exc1 + line_exc2 + line_inh1 + line_inh2
# labs = [l.get_label() for l in lines]
# ax.legend(lines, labs, loc=0)
simpleaxis(ax)
simpleaxis(ax_copy,every=True)
crossCorr = np.corrcoef(std_e_0, std_i_0)[0,1]
ax.set_title("{:.2f}".format(crossCorr), loc='left')
crossCorr = np.corrcoef(std_e, std_i)[0,1]
ax_copy.set_title("{:.2f}".format(crossCorr), loc='right')
ax_copy.spines['right'].set_visible(True)
ax_copy.get_yaxis().tick_right()
# ax.set_ylim(ymin=-0.5, ymax=3.)
# ax_copy.set_ylim(ymin=-0.5, ymax=14.)
# ax.set_yticks(np.linspace(0,3.,3))
# ax_copy.set_yticks(np.linspace(0,14.,3))
ax.set_ylabel("Excitation")
ax_copy.set_ylabel("Inhibition")
fig.set_figheight(1.5)
fig.set_figwidth(3)
print(ax.get_ylim())
print(ax_copy.get_ylim())
plt.savefig('/home/bhalla/Documents/Codes/linearity/Paper_Figures/figures/supplementary/' + 'model_2_corr_plot_std' + '.svg')
plt.show()
```
## Varying the scale of weight distribution.
```
dirPrefix = '/home/bhalla/Documents/Codes/linearity/DamnBalance/Figures/'
plotSuffix = '_{}_{}_{}_{}_{}_{}_{}'.format(w_e_shape, w_e_scale, lam, rho, sigma, w_k, n_k)
plotType = '.svg'
w_e_shape = -0.39
scale_step = 9
w_e_scale_arr = np.linspace(0.5, 1.0, scale_step)
N_E, N_I = 4000, 1000
num_stim = 30
lam = 5 # Average number of synapses per stimulus
rho, sigma = 1., 1.
rho_step, sigma_step = 9, 9
w_k = 1.
n_k = 1.
numTrials = 6
runs = 100
n1 = Neuron(w_e, w_k, N_I)
rho_array = np.linspace(0,1,rho_step)
#sigma_array = np.linspace(0,1,sigma_step)
corrMat = np.zeros((scale_step, rho_step))
rSqMat = np.zeros((scale_step, rho_step))
corrMat_std = np.zeros((scale_step, rho_step))
rSqMat_std = np.zeros((scale_step, rho_step))
w_e_fano = np.zeros(scale_step)
for run in range(runs):
for i,w_e_scale in enumerate(w_e_scale_arr):
w_e = np.random.lognormal(w_e_shape, w_e_scale, N_E)
n1.w_e = w_e
# histogram(w_e, "Exc wts.", filename= dirPrefix + 'w_e_hist_w_dist_{}_{}'.format(run,i) + plotSuffix + plotType, show=False)
n_e = np.random.poisson(lam, num_stim)
w_e_fano[i] += (np.var(w_e)/np.mean(w_e))
for j,rho in enumerate(rho_array):
n1.w_i = n1.getTunedWeights(rho=rho, k=w_k)
n1.generateStimuli(n_e, sigma=sigma, k=n_k)
w_e_all, w_i_all, num_e_all, num_i_all = n1.giveStimuli(numTrials=numTrials)
# scatterPlots(w_e_all, w_i_all, filename= dirPrefix + 'scatter_w_e_w_dist_{}_{}'.format(run,i) + plotSuffix + plotType, show=False)
avg_e, avg_i, std_e, std_i = getAveragedStimuli(w_e_all, w_i_all)
corrMat[i,j] += np.corrcoef(avg_e, avg_i)[0,1]
slope, intercept, r_value, p_value, std_err = ss.linregress(avg_e, avg_i)
rSqMat[i,j] += r_value**2
corrMat_std[i,j] += np.corrcoef(std_e, std_i)[0,1]
slope, intercept, r_value, p_value, std_err = ss.linregress(std_e, std_i)
rSqMat_std[i,j] += r_value**2
w_e_fano/=runs
corrMat/=runs
rSqMat/=runs
corrMat_std/=runs
rSqMat_std/=runs
w_e_scale_arr
plotType = '.svg'
histogram(n_e, "Syn./stim.", filename= dirPrefix + 'n_e_hist_w_dist' + plotSuffix + plotType)
drawHeatmapCorrelationMat(corrMat, y_arr=w_e_fano, x_arr=rho_array, colorNormalize= 'max', ylabel = '$\sigma^2_{exc}/\mu_{exc}$', xlabel = '$\\rho$', cbarlabel= 'avg. corr.', filename=dirPrefix + 'corr_EI_w_dist' + plotSuffix + plotType)
drawHeatmapCorrelationMat(rSqMat, y_arr=w_e_fano, x_arr=rho_array, colorNormalize= 'max', ylabel = '$\sigma^2_{exc}/\mu_{exc}$', xlabel = '$\\rho$', cbarlabel= 'avg. $R^2$', filename=dirPrefix + 'rsq_EI_w_dist' + plotSuffix + plotType)
drawHeatmapCorrelationMat(corrMat_std, y_arr=w_e_fano, x_arr=rho_array, colorNormalize= 'max', ylabel = '$\sigma^2_{exc}/\mu_{exc}$', xlabel = '$\\rho$', cbarlabel= 'avg. corr.', filename=dirPrefix + 'std_corr_EI_w_dist' + plotSuffix + plotType)
drawHeatmapCorrelationMat(rSqMat_std, y_arr=w_e_fano, x_arr=rho_array, colorNormalize= 'max', ylabel = '$\sigma^2_{exc}/\mu_{exc}$', xlabel = '$\\rho$', cbarlabel= 'avg. $R^2$', filename=dirPrefix + 'std_rsq_EI_w_dist' + plotSuffix + plotType)
```
## Varying both rho and sigma and plotting the values of correlations between Exc and Inh synaptic sums.
```
w_e_shape = -0.39
w_e_scale = [0.80] # np.linspace(0.5, 1.0, 10)
N_E, N_I = 4000, 1000
num_stim = 30
lam = 5 # Average number of synapses per stimulus
rho, sigma = 1., 1.
rho_step, sigma_step = 9, 9
w_k = 4.
n_k = 0.25
numTrials = 6
runs = 50
w_e = np.random.lognormal(w_e_shape, w_e_scale[0], N_E)
n_e = np.random.poisson(lam, num_stim)
n1 = Neuron(w_e, w_k, N_I)
rho_array = np.linspace(0,1,rho_step)
sigma_array = np.linspace(0,1,sigma_step)
corrMat = np.zeros((rho_step, sigma_step))
rSqMat = np.zeros((rho_step, sigma_step))
for run in range(runs):
for i,rho in enumerate(rho_array):
for j, sigma in enumerate(sigma_array):
n1.w_i = n1.getTunedWeights(rho=rho, k=w_k)
n1.generateStimuli(n_e, sigma=sigma, k=n_k)
w_e_all, w_i_all, num_e_all, num_i_all = n1.giveStimuli(numTrials=numTrials)
avg_e, avg_i, std_e, std_i = getAveragedStimuli(w_e_all, w_i_all)
corrMat[i,j] += np.corrcoef(avg_e, avg_i)[0,1]
slope, intercept, r_value, p_value, std_err = ss.linregress(avg_e, avg_i)
rSqMat[i,j] += r_value**2
corrMat/=runs
rSqMat/=runs
dirPrefix = '/home/bhalla/Documents/Codes/linearity/DamnBalance/'
plotSuffix = '_{}_{}_{}_{}_{}_{}_{}'.format(w_e_shape, w_e_scale, lam, rho, sigma, w_k, n_k)
plotType = '.svg'
histogram(w_e, "Exc wts.", filename= dirPrefix + 'w_e_hist' + plotSuffix + plotType)
histogram(n_e, "Syn./stim.", filename= dirPrefix + 'n_e_hist' + plotSuffix + plotType)
drawHeatmapCorrelationMat(corrMat, x_arr=sigma_array, y_arr=rho_array, colorNormalize= 'max', xlabel = '$\\sigma$', ylabel = '$\\rho$', cbarlabel= 'avg. corr.', filename=dirPrefix + 'corr_EI' + plotSuffix + plotType)
drawHeatmapCorrelationMat(rSqMat, x_arr=sigma_array, y_arr=rho_array, colorNormalize= 'max', xlabel = '$\\sigma$', ylabel = '$\\rho$', cbarlabel= 'avg. $R^2$', filename=dirPrefix + 'rsq_EI' + plotSuffix + plotType)
```
| github_jupyter |
# Introduction to Feature Columns
**Learning Objectives**
1. Load a CSV file using [Pandas](https://pandas.pydata.org/)
2. Create an input pipeline using tf.data
3. Create multiple types of feature columns
## Introduction
In this notebook, you classify structured data (e.g. tabular data in a CSV file) using [feature columns](https://www.tensorflow.org/guide/feature_columns). Feature columns serve as a bridge to map from columns in a CSV file to features used to train a model. In a subsequent lab, we will use [Keras](https://www.tensorflow.org/guide/keras) to define the model.
Each learning objective will correspond to a __#TODO__ in the [student lab notebook](../labs/feat.cols_tf.data.ipynb) -- try to complete that notebook first before reviewing this solution notebook.
## The Dataset
We will use a small [dataset](https://archive.ics.uci.edu/ml/datasets/heart+Disease) provided by the Cleveland Clinic Foundation for Heart Disease. There are several hundred rows in the CSV. Each row describes a patient, and each column describes an attribute. We will use this information to predict whether a patient has heart disease, which in this dataset is a binary classification task.
Following is a [description](https://archive.ics.uci.edu/ml/machine-learning-databases/heart-disease/heart-disease.names) of this dataset. Notice there are both numeric and categorical columns.
>Column| Description| Feature Type | Data Type
>------------|--------------------|----------------------|-----------------
>Age | Age in years | Numerical | integer
>Sex | (1 = male; 0 = female) | Categorical | integer
>CP | Chest pain type (0, 1, 2, 3, 4) | Categorical | integer
>Trestbpd | Resting blood pressure (in mm Hg on admission to the hospital) | Numerical | integer
>Chol | Serum cholestoral in mg/dl | Numerical | integer
>FBS | (fasting blood sugar > 120 mg/dl) (1 = true; 0 = false) | Categorical | integer
>RestECG | Resting electrocardiographic results (0, 1, 2) | Categorical | integer
>Thalach | Maximum heart rate achieved | Numerical | integer
>Exang | Exercise induced angina (1 = yes; 0 = no) | Categorical | integer
>Oldpeak | ST depression induced by exercise relative to rest | Numerical | float
>Slope | The slope of the peak exercise ST segment | Numerical | integer
>CA | Number of major vessels (0-3) colored by flourosopy | Numerical | integer
>Thal | 3 = normal; 6 = fixed defect; 7 = reversable defect | Categorical | string
>Target | Diagnosis of heart disease (1 = true; 0 = false) | Classification | integer
## Import TensorFlow and other libraries
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
import tensorflow as tf
from tensorflow import feature_column
from tensorflow.keras import layers
from sklearn.model_selection import train_test_split
print("TensorFlow version: ",tf.version.VERSION)
```
## Lab Task 1: Use Pandas to create a dataframe
[Pandas](https://pandas.pydata.org/) is a Python library with many helpful utilities for loading and working with structured data. We will use Pandas to download the dataset from a URL, and load it into a dataframe.
```
URL = 'https://storage.googleapis.com/applied-dl/heart.csv'
dataframe = pd.read_csv(URL)
dataframe.head()
dataframe.info()
```
## Split the dataframe into train, validation, and test
The dataset we downloaded was a single CSV file. As a best practice, we will split this into train, validation, and test sets.
```
# TODO 1a
train, test = train_test_split(dataframe, test_size=0.2)
train, val = train_test_split(train, test_size=0.2)
print(len(train), 'train examples')
print(len(val), 'validation examples')
print(len(test), 'test examples')
```
## Lab Task 2: Create an input pipeline using tf.data
Next, we will wrap the dataframes with [tf.data](https://www.tensorflow.org/guide/datasets). This will enable us to use feature columns as a bridge to map from the columns in the Pandas dataframe to features used to train a model. If we were working with a very large CSV file (so large that it does not fit into memory), we would use tf.data to read it from disk directly. That is not covered in this lab.
```
# A utility method to create a tf.data dataset from a Pandas Dataframe
def df_to_dataset(dataframe, shuffle=True, batch_size=32):
dataframe = dataframe.copy()
labels = dataframe.pop('target')
ds = tf.data.Dataset.from_tensor_slices((dict(dataframe), labels)) # TODO 2a
if shuffle:
ds = ds.shuffle(buffer_size=len(dataframe))
ds = ds.batch(batch_size)
return ds
batch_size = 5 # A small batch sized is used for demonstration purposes
# TODO 2b
train_ds = df_to_dataset(train, batch_size=batch_size)
val_ds = df_to_dataset(val, shuffle=False, batch_size=batch_size)
test_ds = df_to_dataset(test, shuffle=False, batch_size=batch_size)
```
## Understand the input pipeline
Now that we have created the input pipeline, let's call it to see the format of the data it returns. We have used a small batch size to keep the output readable.
```
for feature_batch, label_batch in train_ds.take(1):
print('Every feature:', list(feature_batch.keys()))
print('A batch of ages:', feature_batch['age'])
print('A batch of targets:', label_batch)
```
## Lab Task 3: Demonstrate several types of feature column
TensorFlow provides many types of feature columns. In this section, we will create several types of feature columns, and demonstrate how they transform a column from the dataframe.
```
# We will use this batch to demonstrate several types of feature columns
example_batch = next(iter(train_ds))[0]
# A utility method to create a feature column
# and to transform a batch of data
def demo(feature_column):
feature_layer = layers.DenseFeatures(feature_column)
print(feature_layer(example_batch).numpy())
```
### Numeric columns
The output of a feature column becomes the input to the model. A [numeric column](https://www.tensorflow.org/api_docs/python/tf/feature_column/numeric_column) is the simplest type of column. It is used to represent real valued features. When using this column, your model will receive the column value from the dataframe unchanged.
```
age = feature_column.numeric_column("age")
tf.feature_column.numeric_column
print(age)
```
### Let's have a look at the output:
#### key='age'
A unique string identifying the input feature. It is used as the column name and the dictionary key for feature parsing configs, feature Tensor objects, and feature columns.
#### shape=(1,)
In the heart disease dataset, most columns from the dataframe are numeric. Recall that tensors have a rank. "Age" is a "vector" or "rank-1" tensor, which is like a list of values. A vector has 1-axis, thus the shape will always look like this: shape=(3,), where 3 is a scalar (or single number) and with 1-axis.
#### default_value=None
A single value compatible with dtype or an iterable of values compatible with dtype which the column takes on during tf.Example parsing if data is missing. A default value of None will cause tf.io.parse_example to fail if an example does not contain this column. If a single value is provided, the same value will be applied as the default value for every item. If an iterable of values is provided, the shape of the default_value should be equal to the given shape.
#### dtype=tf.float32
defines the type of values. Default value is tf.float32. Must be a non-quantized, real integer or floating point type.
#### normalizer_fn=None
If not None, a function that can be used to normalize the value of the tensor after default_value is applied for parsing. Normalizer function takes the input Tensor as its argument, and returns the output Tensor. (e.g. lambda x: (x - 3.0) / 4.2). Please note that even though the most common use case of this function is normalization, it can be used for any kind of Tensorflow transformations.
```
demo(age)
```
### Bucketized columns
Often, you don't want to feed a number directly into the model, but instead split its value into different categories based on numerical ranges. Consider raw data that represents a person's age. Instead of representing age as a numeric column, we could split the age into several buckets using a [bucketized column](https://www.tensorflow.org/api_docs/python/tf/feature_column/bucketized_column). Notice the one-hot values below describe which age range each row matches.
```
age_buckets = tf.feature_column.bucketized_column(age, boundaries=[18, 25, 30, 35, 40, 45, 50, 55, 60, 65])
demo(age_buckets) # TODO 3a
```
### Categorical columns
In this dataset, thal is represented as a string (e.g. 'fixed', 'normal', or 'reversible'). We cannot feed strings directly to a model. Instead, we must first map them to numeric values. The categorical vocabulary columns provide a way to represent strings as a one-hot vector (much like you have seen above with age buckets). The vocabulary can be passed as a list using [categorical_column_with_vocabulary_list](https://www.tensorflow.org/api_docs/python/tf/feature_column/categorical_column_with_vocabulary_list), or loaded from a file using [categorical_column_with_vocabulary_file](https://www.tensorflow.org/api_docs/python/tf/feature_column/categorical_column_with_vocabulary_file).
```
thal = tf.feature_column.categorical_column_with_vocabulary_list(
'thal', ['fixed', 'normal', 'reversible'])
thal_one_hot = tf.feature_column.indicator_column(thal)
demo(thal_one_hot)
```
In a more complex dataset, many columns would be categorical (e.g. strings). Feature columns are most valuable when working with categorical data. Although there is only one categorical column in this dataset, we will use it to demonstrate several important types of feature columns that you could use when working with other datasets.
### Embedding columns
Suppose instead of having just a few possible strings, we have thousands (or more) values per category. For a number of reasons, as the number of categories grow large, it becomes infeasible to train a neural network using one-hot encodings. We can use an embedding column to overcome this limitation. Instead of representing the data as a one-hot vector of many dimensions, an [embedding column](https://www.tensorflow.org/api_docs/python/tf/feature_column/embedding_column) represents that data as a lower-dimensional, dense vector in which each cell can contain any number, not just 0 or 1. The size of the embedding (8, in the example below) is a parameter that must be tuned.
Key point: using an embedding column is best when a categorical column has many possible values. We are using one here for demonstration purposes, so you have a complete example you can modify for a different dataset in the future.
```
# Notice the input to the embedding column is the categorical column
# we previously created
thal_embedding = tf.feature_column.embedding_column(thal, dimension=8)
demo(thal_embedding)
```
### Hashed feature columns
Another way to represent a categorical column with a large number of values is to use a [categorical_column_with_hash_bucket](https://www.tensorflow.org/api_docs/python/tf/feature_column/categorical_column_with_hash_bucket). This feature column calculates a hash value of the input, then selects one of the `hash_bucket_size` buckets to encode a string. When using this column, you do not need to provide the vocabulary, and you can choose to make the number of hash_buckets significantly smaller than the number of actual categories to save space.
Key point: An important downside of this technique is that there may be collisions in which different strings are mapped to the same bucket. In practice, this can work well for some datasets regardless.
```
thal_hashed = tf.feature_column.categorical_column_with_hash_bucket(
'thal', hash_bucket_size=1000)
demo(tf.feature_column.indicator_column(thal_hashed))
```
### Crossed feature columns
Combining features into a single feature, better known as [feature crosses](https://developers.google.com/machine-learning/glossary/#feature_cross), enables a model to learn separate weights for each combination of features. Here, we will create a new feature that is the cross of age and thal. Note that `crossed_column` does not build the full table of all possible combinations (which could be very large). Instead, it is backed by a `hashed_column`, so you can choose how large the table is.
```
crossed_feature = tf.feature_column.crossed_column([age_buckets, thal], hash_bucket_size=1000)
demo(tf.feature_column.indicator_column(crossed_feature))
```
## Choose which columns to use
We have seen how to use several types of feature columns. Now we will use them to train a model. The goal of this tutorial is to show you the complete code (e.g. mechanics) needed to work with feature columns. We have selected a few columns to train our model below arbitrarily.
Key point: If your aim is to build an accurate model, try a larger dataset of your own, and think carefully about which features are the most meaningful to include, and how they should be represented.
```
feature_columns = []
# numeric cols
for header in ['age', 'trestbps', 'chol', 'thalach', 'oldpeak', 'slope', 'ca']:
feature_columns.append(feature_column.numeric_column(header))
# bucketized cols
age_buckets = feature_column.bucketized_column(age, boundaries=[18, 25, 30, 35, 40, 45, 50, 55, 60, 65])
feature_columns.append(age_buckets)
# indicator cols
thal = feature_column.categorical_column_with_vocabulary_list(
'thal', ['fixed', 'normal', 'reversible'])
thal_one_hot = feature_column.indicator_column(thal)
feature_columns.append(thal_one_hot)
# embedding cols
thal_embedding = feature_column.embedding_column(thal, dimension=8)
feature_columns.append(thal_embedding)
# crossed cols
crossed_feature = feature_column.crossed_column([age_buckets, thal], hash_bucket_size=1000)
crossed_feature = feature_column.indicator_column(crossed_feature)
feature_columns.append(crossed_feature)
```
### How to Input Feature Columns to a Keras Model
Now that we have defined our feature columns, we now use a [DenseFeatures](https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/keras/layers/DenseFeatures) layer to input them to a Keras model. Don't worry if you have not used Keras before. There is a more detailed video and lab introducing the Keras Sequential and Functional models.
```
feature_layer = tf.keras.layers.DenseFeatures(feature_columns)
```
Earlier, we used a small batch size to demonstrate how feature columns worked. We create a new input pipeline with a larger batch size.
```
batch_size = 32
train_ds = df_to_dataset(train, batch_size=batch_size)
val_ds = df_to_dataset(val, shuffle=False, batch_size=batch_size)
test_ds = df_to_dataset(test, shuffle=False, batch_size=batch_size)
```
## Create, compile, and train the model
```
model = tf.keras.Sequential([
feature_layer,
layers.Dense(128, activation='relu'),
layers.Dense(128, activation='relu'),
layers.Dense(1)
])
model.compile(optimizer='adam',
loss=tf.keras.losses.BinaryCrossentropy(from_logits=True),
metrics=['accuracy'])
history = model.fit(train_ds,
validation_data=val_ds,
epochs=5)
loss, accuracy = model.evaluate(test_ds)
print("Accuracy", accuracy)
```
### Visualize the model loss curve
Next, we will use Matplotlib to draw the model's loss curves for training and validation. A line plot is also created showing the accuracy over the training epochs for both the train (blue) and test (orange) sets.
```
def plot_curves(history, metrics):
nrows = 1
ncols = 2
fig = plt.figure(figsize=(10, 5))
for idx, key in enumerate(metrics):
ax = fig.add_subplot(nrows, ncols, idx+1)
plt.plot(history.history[key])
plt.plot(history.history['val_{}'.format(key)])
plt.title('model {}'.format(key))
plt.ylabel(key)
plt.xlabel('epoch')
plt.legend(['train', 'validation'], loc='upper left');
plot_curves(history, ['loss', 'accuracy'])
```
You can see that accuracy is at 77% for both the training and validation data, while loss bottoms out at about .477 after four epochs.
Key point: You will typically see best results with deep learning with much larger and more complex datasets. When working with a small dataset like this one, we recommend using a decision tree or random forest as a strong baseline. The goal of this tutorial is not to train an accurate model, but to demonstrate the mechanics of working with structured data, so you have code to use as a starting point when working with your own datasets in the future.
## Next steps
The best way to learn more about classifying structured data is to try it yourself. We suggest finding another dataset to work with, and training a model to classify it using code similar to the above. To improve accuracy, think carefully about which features to include in your model, and how they should be represented.
Copyright 2020 Google Inc.
Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
| github_jupyter |
```
# import necessary libraries
import pandas as pd
import numpy as np
from tsfresh import select_features
import xgboost
import seaborn as sns
import matplotlib.pyplot as plt
import math, datetime
from sklearn.preprocessing import StandardScaler
from sklearn.metrics import explained_variance_score
from sklearn.metrics import mean_squared_error
from sklearn.pipeline import Pipeline
from sklearn.feature_selection import SelectFromModel
from feature_selection import FeatureSelector
from hyperopt import STATUS_OK, Trials, fmin, hp, tpe
%matplotlib
from matplotlib import pylab
# all extracted features from tsfresh
# the extracted features are the same for all models
full_features = pd.read_pickle('/Users/Rohil/Documents/iGEM/yemen/exhaustive_extracted_features_new.pkl')
# test values for different time frames (in a nice format)
y_df = pd.read_pickle('/Users/Rohil/Documents/iGEM/yemen/y_df_for_feature_selection_new.pkl')
# adding governorates
full_features = pd.concat([full_features, pd.get_dummies(full_features.gov_iso)], axis=1)
# ignoring anything that happens before Jul 1, as there's not enough data for feature calculation
full_features = full_features[full_features.date>='2017-07-01'].reset_index(drop=True)
y_df = y_df[y_df.date>='2017-07-01'].reset_index(drop=True)
ESTIMATORS = 750
EARLY_STOPPING_ROUNDS = 50
# scale only continuos columns using standard scaler
def scale_features(scaler, X):
continuous_cols = [col for col in X.columns if 'YE-' not in col]
X_scaled = X
X_scaled[continuous_cols] = scaler.transform(X[continuous_cols])
return (X_scaled)
# split data into train, cross-validation, and test based on date
# note that the below function is NOT FOR ROLLING WINDOW CROSS VALIDATION
# note that later on train and cv are usually combined (appended) when doing time series cross-validation
# in those cases, the test set acts as the cross-validation and the train + cv set acts as the train set
# the code below was not changed to retain its 3-split functionality
def train_cv_test_split(data, cv_split_date='2017-09-17', test_split_date='2017-11-06'):
data_train, data_cv, data_test = data[(data.date<cv_split_date)], data[(data.date<test_split_date) & (data.date>=cv_split_date)], data[ (data.date>=test_split_date) ]
return (data_train.drop('date', axis=1), data_cv.drop('date', axis=1), data_test.drop('date', axis=1))
# functions to obtain start/stop, particularly the start of the train/cv and start of the holdout (which becomes the stop of the train/cv)
#dates based on defined hold out and rolling window size
HOLD_OUT_WINDOW = 100
ROLLING_WINDOW_SIZE = 45
WINDOW = round(ROLLING_WINDOW_SIZE/3)
def getMaxDate():
return y_df.date.max() - datetime.timedelta(days=HOLD_OUT_WINDOW)
def getMinDate():
return y_df.date.min()
delta = getMaxDate() - getMinDate()
print('Start {} Stop {} No of days {}'.format(getMinDate(), getMaxDate(), delta.days))
number_rolling_windows = math.ceil(round(delta.days/WINDOW)) - 1
number_rolling_windows
# builds off previous function to get hold-out dates
def getHoldOutDate():
start = getMaxDate() + datetime.timedelta(days=1)
end = y_df.date.max()
return (start, end)
# get dates for rolling window cross-validation
def getRollingWindowDates(idx):
maxDate = getMaxDate()
minDate = getMinDate()
trainStart = minDate
trainStop = minDate + datetime.timedelta(days=WINDOW*idx)
validationStart = trainStop + datetime.timedelta(days=1)
validationStop = validationStart + datetime.timedelta(days=WINDOW)
testStart = validationStop + datetime.timedelta(days=1)
testStop = testStart + datetime.timedelta(days=WINDOW)
if (maxDate - testStop).days < WINDOW:
print('Rolling window to end date')
testStop = maxDate
print('Train [{} {}] Val [{} {}] Test [{} {}]'.format(trainStart.date(), trainStop.date(),
validationStart.date(), validationStop.date(),
testStart.date(), testStop.date()))
return (trainStart, trainStop, validationStart, validationStop, testStart, testStop)
# using dates from getRollingWindowDates, split data into train, validation, test
def getRollingWindow(data, trainStart, trainStop, validationStart, validationStop, testStart, testStop):
train = data[(data.date >= trainStart) & (data.date <= trainStop)]
val = data[(data.date >= validationStart) & (data.date <= validationStop)]
test = data[(data.date >= testStart) & (data.date <= testStop)]
print('Window Train/Val/Test shape {} {} {}'.format(train.shape, val.shape, test.shape))
return (train.drop('date', axis=1), val.drop('date', axis=1), test.drop('date', axis=1))
# splits data into holdout and train based on defined minimum date and hold-out date (see relevant functions above)
def getHoldOutData(data):
minDate = getMinDate()
start, end = getHoldOutDate()
train = data[(data.date >= minDate) & (data.date < start)]
test = data[(data.date >= start) & (data.date <= end)]
return (train.drop('date', axis=1), test.drop('date', axis=1))
number_rolling_windows, round((ROLLING_WINDOW_SIZE)/3), getHoldOutDate()
# uses scalable hypothesis tests to identify statistically relevant features
class TsFresh():
def __init__(self):
pass
def postProcessor(self, X, y, dateSeries):
self.selected_features = select_features(X, y, fdr_level=0.001)
print('Selected features {}'.format(self.selected_features.shape))
self.selected_features = pd.concat([dateSeries, self.selected_features], axis=1)
# removes features that are correlated with each other
class OptimizeFeatures():
def __init__(self):
pass
def selectFeatures(self, X, y):
self.X = X
self.y = y
self.fs = FeatureSelector(data = X, labels = y)
def identifyCollinearFeatures(self, correlation_threshold=0.975):
self.fs.identify_collinear(correlation_threshold)
def collinerFeaturesColumnsToKeep(self):
return self.fs.ops['collinear']
def removeCollinerFeatures(self):
self.cols_to_keep = set(self.X.columns) - set(self.fs.ops['collinear'])
self.corr_selected_features = self.X[list(self.cols_to_keep)]
# trains XGBoost model to rank features
class ModelCustomRegressor():
def __init__(self):
pass
def extract(self, model_params, X_train, X_cv, X_test, y_train, y_cv, y_test):
print('Creating baseline model to extract features')
X_train_cv = X_train.append(X_cv)
y_train_cv = y_train.append(y_cv)
scaler = StandardScaler()
continuous_cols = [col for col in X_train_cv.columns if 'YE-' not in col]
scaler.fit(X_train_cv[continuous_cols])
X_train_cv, X_test = scale_features(scaler, X_train_cv), scale_features(scaler, X_test)
print('all features {}'.format(X_train_cv.shape))
eval_set = [(X_test, y_test)]
self.feature_importance_df = pd.DataFrame(index = X_train.columns)
self.regressor = xgboost.XGBRegressor(**model_params)
self.regressor.fit(X_train_cv, y_train_cv, eval_metric='rmse',
eval_set=eval_set,
early_stopping_rounds=EARLY_STOPPING_ROUNDS, verbose=False)
preds = self.regressor.predict((X_test))
self.feature_importance_df['threshold'] = self.regressor.feature_importances_
self.thresholds = np.unique(self.regressor.feature_importances_)
self.thresholds.sort();
print (len(self.thresholds))
self.thresholds = self.thresholds[::-1][:50]
print('# features {} # thresholds {}; thresholds:{}'.format(len(self.regressor.feature_importances_), len(self.thresholds), self.thresholds))
# performs rolling-window cross-validation
class RollingWindowCrossValidation():
def __init__(self, corr_selected_features_bi_week, corr_selected_features_bi_week_y, preselect_params):
self.corr_selected_features_bi_week = corr_selected_features_bi_week
self.corr_selected_features_bi_week_y = corr_selected_features_bi_week_y
self.preselect_params = preselect_params
pass
def extract(self, regressor, thresh):
mse_list = []
for idx in range(3, number_rolling_windows):
print('CV - Window {}'.format(idx))
trainStart, trainStop, validationStart, validationStop, testStart, testStop = getRollingWindowDates(idx)
X_train, X_cv, X_test = getRollingWindow(self.corr_selected_features_bi_week,
trainStart, trainStop,
validationStart, validationStop, testStart, testStop)
y_train, y_cv, y_test = getRollingWindow(self.corr_selected_features_bi_week_y,
trainStart, trainStop,
validationStart, validationStop, testStart, testStop)
X_train_cv = X_train.append(X_cv)
y_train_cv = y_train.append(y_cv)
scaler = StandardScaler()
continuous_cols = [col for col in X_train_cv.columns if 'YE-' not in col]
scaler.fit(X_train_cv[continuous_cols])
X_train_cv, X_test = scale_features(scaler, X_train_cv), scale_features(scaler, X_test)
print('X_train_cv {}'.format(X_train_cv.shape))
# select features using threshold
selection = SelectFromModel(regressor, threshold=thresh, prefit=True)
select_X_train_cv = selection.transform(X_train_cv)
select_X_test = selection.transform(X_test)
eval_set = [(select_X_test, y_test)]
# train model
selection_model = xgboost.XGBRegressor(**self.preselect_params)
selection_model.fit(select_X_train_cv, y_train_cv, eval_metric='rmse',
eval_set=eval_set,
early_stopping_rounds=50,
verbose=False)
# eval model
y_pred = selection_model.predict(select_X_test)
mse = mean_squared_error(y_test, y_pred)
print("%d Thresh=%.5f, n=%d, mse: %.3f" % (idx, thresh, select_X_train_cv.shape[1], mse))
mse_list.append(mse)
return_dict = {'threshold':thresh, 'num_features':select_X_train_cv.shape[1], 'mse_list':mse_list, 'mean_mse':np.mean(mse_list)}
print (return_dict)
return (return_dict)
# takes feature ranking and recursively adds features
# evaluates how each feature set performs using rolling window cross validation
class RollingCustomFeatureExtractor():
def __init__(self, corr_selected_features_bi_week, corr_selected_features_bi_week_y, preselect_params):
self.corr_selected_features_bi_week = corr_selected_features_bi_week
self.corr_selected_features_bi_week_y = corr_selected_features_bi_week_y
self.preselect_params = preselect_params
pass
def extract(self):
X = self.corr_selected_features_bi_week
y = self.corr_selected_features_bi_week_y
X_train, X_cv, X_test = train_cv_test_split(X)
y_train, y_cv, y_test = train_cv_test_split(y)
self.mcr = ModelCustomRegressor()
self.mcr.extract(self.preselect_params, X_train, X_cv, X_test, y_train, y_cv, y_test)
thresholds = self.mcr.thresholds
self.rwcv = RollingWindowCrossValidation(self.corr_selected_features_bi_week,
self.corr_selected_features_bi_week_y,
self.preselect_params)
self.summary = pd.DataFrame(columns = ['threshold', 'num_features', 'mse_list', 'mean_mse'])
for thresh in thresholds:
return_dict = self.rwcv.extract(self.mcr.regressor, thresh)
self.summary = self.summary.append(return_dict, ignore_index = True)
print("\n")
print(self.summary.head(20))
def set_style(color):
plt.style.use(['seaborn-' + color, 'seaborn-paper'])
# bayesian optimization with rolling window cross validation to select optimal hyperparameters
class BayesianOptimizer():
def __init__(self, corr_selected_features_bi_week, corr_selected_features_bi_week_y, max_evals):
self.corr_selected_features_bi_week = corr_selected_features_bi_week
self.corr_selected_features_bi_week_y = corr_selected_features_bi_week_y
self.max_evals = max_evals
def objective(self, space):
mse_list = []
for idx in range(3, number_rolling_windows):
trainStart, trainStop, validationStart, validationStop, testStart, testStop = getRollingWindowDates(idx)
X_train, X_cv, X_test = getRollingWindow(self.corr_selected_features_bi_week,
trainStart, trainStop,
validationStart, validationStop, testStart, testStop)
y_train, y_cv, y_test = getRollingWindow(self.corr_selected_features_bi_week_y,
trainStart, trainStop,
validationStart, validationStop, testStart, testStop)
X_train_cv = X_train.append(X_cv)
y_train_cv = y_train.append(y_cv)
scaler = StandardScaler()
continuous_cols = [col for col in X_train_cv.columns if 'YE-' not in col]
scaler.fit(X_train_cv[continuous_cols])
X_train_cv, X_test = scale_features(scaler, X_train_cv), scale_features(scaler, X_test)
xgb = xgboost.XGBRegressor(n_estimators = int(space['n_estimators']),
max_depth = int(space['max_depth']),
min_child_weight = space['min_child_weight'],
subsample = space['subsample'],
learning_rate = space['learning_rate'],
gamma = space['gamma'],
colsample_bytree = space['colsample_bytree'],
objective='reg:linear', n_jobs = -1
)
xgb.fit(X_train_cv ,y_train_cv, eval_metric = 'rmse')
# eval model
y_pred = xgb.predict(X_test)
mse = mean_squared_error(y_test, y_pred)
mse_list.append(mse)
print(mse_list)
return_dict = {'mse_list':mse_list}
return_dict['loss'] = np.mean(mse_list)
return_dict['status'] = STATUS_OK
print ("mean mse:", return_dict['loss'])
return (return_dict)
def run(self):
self.space ={'max_depth': hp.quniform('max_depth', 4, 12, 1),
'min_child_weight': hp.quniform('min_child_weight', 1, 10, 1),
'subsample': hp.quniform('subsample', 0.5, 1, 0.05),
'n_estimators' : hp.quniform('n_estimators', 50, 750, 50),
'learning_rate' : hp.quniform('learning_rate', 0.01, 0.3, 0.025),
'gamma' : hp.quniform('gamma', 0, 5, 0.5),
'colsample_bytree' : hp.quniform('colsample_bytree', 0.3, 1, 0.05)}
self.trials = Trials()
self.best = fmin(fn=self.objective,
space=self.space,
algo=tpe.suggest,
max_evals=self.max_evals,
trials=self.trials)
ylabel_dict = {'week_1_to_2_cases':'0-2 week', 'week_2_to_4_cases':'2-4 week', 'week_4_to_6_cases':'4-6 week', 'week_6_to_8_cases':'6-8 week'}
def suplabel(axis,label,side,label_prop=None,labelpad=6.7,ha='center',va='center'):
''' Add super ylabel or xlabel to the figure
Similar to matplotlib.suptitle
axis - string: "x" or "y"
label - string
label_prop - keyword dictionary for Text
labelpad - padding from the axis (default: 5)
ha - horizontal alignment (default: "center")
va - vertical alignment (default: "center")
'''
fig = pylab.gcf()
xmin = []
ymin = []
xmax = []
ymax = []
for ax in fig.axes:
xmin.append(ax.get_position().xmin)
ymin.append(ax.get_position().ymin)
xmax.append(ax.get_position().xmax)
ymax.append(ax.get_position().ymax)
xmin,ymin = min(xmin),min(ymin)
xmax,ymax = min(xmax),min(ymax)
dpi = fig.dpi
if axis.lower() == "y":
rotation=90.
if side == 'left':
x = xmin-float(labelpad)/dpi
if side == 'right':
x = xmax+float(labelpad)/dpi
y = 0.5
elif axis.lower() == 'x':
rotation = 0.
x = 0.5
y = ymin - float(labelpad)/dpi
else:
raise Exception("Unexpected axis: x or y")
if label_prop is None:
label_prop = dict()
pylab.text(x,y,label,rotation=rotation,
transform=fig.transFigure,
ha=ha,va=va,
**label_prop)
# plotting function that visualizes predictions
def plot_pred_against_actual(cv_pred_crosstab, test_pred_crosstab, true_crosstab, y_label, cv_mse, holdout_mse, sharey):
set_style('white')
fig, ax = plt.subplots(21,1,figsize = (6,15), sharex=True, sharey = sharey)
cols = true_crosstab.columns
for i in range(0,21):
true_crosstab[cols[i]].plot(kind='line', ax = ax[i], label = 'true_val', legend = True, color = 'red')
ax[i].set_prop_cycle('color', ['seagreen', 'blue'])
#train_pred_crosstab[cols[i]].plot(kind='line', ax = ax[i], label= 'xgboost train-pred',linestyle= '-.', legend = True)
cv_pred_crosstab[cols[i]].plot(kind='line', ax = ax[i], label= 'xgboost cv-pred;\nmean error: %.3f'%(np.sqrt(cv_mse)) ,linestyle= '-.', legend = True)
test_pred_crosstab[cols[i]].plot(kind='line', ax = ax[i], label= 'xgboost holdout-pred;\nmean error: %.3f'%(np.sqrt(holdout_mse)), linestyle= '-.', legend = True)
ax[i].legend().set_visible(False)
ax[i].set_ylabel(cols[i], rotation=0)
ax[i].yaxis.set_label_position('right')
ax[i].spines['right'].set_visible(False)
ax[i].spines['top'].set_visible(False)
ax[i].spines['bottom'].set_visible(True)
ax[10].legend().set_visible(True)
ax[10].legend(fontsize=10, loc='center left', bbox_to_anchor=(1.05, 0.5))
fig.subplots_adjust(hspace = .2)
if sharey:
fig.savefig('/Users/Rohil/Documents/iGEM/yemen/final_results/' + y_label + '_deployed_sharey.png', dpi = 500, bbox_inches = 'tight')
else:
fig.savefig('/Users/Rohil/Documents/iGEM/yemen/final_results/' + y_label + '_deployed.png', dpi = 500, bbox_inches = 'tight')
plt.close('all')
# plotting function that visualizes predictions, but only for a certain subset of governorates
def plot_pred_against_actual_filtered(cv_pred_crosstab, test_pred_crosstab, true_crosstab, y_label, cv_mse, holdout_mse, sharey):
set_style('white')
#cols = list(set(true_crosstab.columns) - set(['YE-HD-AL', 'YE-MR', 'YE-SH', 'YE-TA', 'YE-MA', 'YE-SD']))
cols = sorted(['YE-AM', 'YE-DA', 'YE-MW', 'YE-RA', 'YE-SN'])
#sorted(['YE-AM', 'YE-DA', 'YE-TA', 'YE-RA', 'YE-SN'])
fig, ax = plt.subplots(len(cols),1,figsize = (5,5), sharex=True, sharey = sharey)
for i in range(0,len(cols)):
true_crosstab[cols[i]].plot(kind='line', ax = ax[i], label = 'true', legend = True, color = 'red')
ax[i].set_prop_cycle('color', ['seagreen', 'blue'])
#train_pred_crosstab[cols[i]].plot(kind='line', ax = ax[i], label= 'xgboost train-pred',linestyle= '-.', legend = True)
cv_pred_crosstab[cols[i]].plot(kind='line', ax = ax[i], label= 'xgboost cross-validation prediction' ,linestyle= '-.', legend = True) #;\nmean error: %.3f'%(np.sqrt(cv_mse))
test_pred_crosstab[cols[i]].plot(kind='line', ax = ax[i], label= 'xgboost holdout prediction', linestyle= '-.', legend = True) #;\nmean error: %.3f'%(np.sqrt(holdout_mse))
ax[i].legend().set_visible(False)
ax[i].set_ylabel(cols[i], rotation=0)
ax[i].yaxis.set_label_position('right')
ax[i].spines['right'].set_visible(False)
ax[i].spines['top'].set_visible(False)
ax[i].spines['bottom'].set_visible(True)
#ax[int(len(cols)/2)].legend().set_visible(True)
#ax[int(len(cols)/2)].legend(fontsize=10, loc='center left', bbox_to_anchor=(1.05, 0.5))
fig.subplots_adjust(hspace = .2)
ax[len(cols)-1].set_xlabel('time (days)', rotation=0)
fig.suptitle(ylabel_dict[y_label] + 's forecast')
suplabel('y', 'sliding-scale prediction of new cases per 10000 people', 'left')
suplabel('y', 'governorates', 'right')
if sharey:
fig.savefig('/Users/Rohil/Documents/iGEM/yemen/final_results/' + y_label + '_deployed_sharey_filtered.png', dpi = 500, bbox_inches = 'tight')
else:
fig.savefig('/Users/Rohil/Documents/iGEM/yemen/final_results/' + y_label + '_deployed_filtered.png', dpi = 500, bbox_inches = 'tight')
plt.close('all')
# used to deploy final iteration of model
# uses rolling window cross-validation and evaluates on hold-out set
# feeds into plotting function
class DeployRegressor():
def __init__(self):
pass
def execute(self, model_params, X, y, y_to_plot, y_bi_week_label):
X.drop(columns = 'days_from', inplace = True)
X_train_all, X_hold_test = getHoldOutData(X)
y_train_all, y_hold_test = getHoldOutData(y)
mse_list = []
y_cv_preds = []
y_cv_all = pd.DataFrame(columns = [y_bi_week_label])
for idx in range(3, number_rolling_windows):
trainStart, trainStop, validationStart, validationStop, testStart, testStop = getRollingWindowDates(idx)
X_train, X_cv, X_test = getRollingWindow(X,
trainStart, trainStop,
validationStart, validationStop, testStart, testStop)
y_train, y_cv, y_test = getRollingWindow(y,
trainStart, trainStop,
validationStart, validationStop, testStart, testStop)
X_train_cv = X_train.append(X_cv)
y_train_cv = y_train.append(y_cv)
# if idx == 3:
# X_base_train = X_train_cv
# y_base_train = y_train_cv
scaler = StandardScaler()
continuous_cols = [col for col in X_train_cv.columns if 'YE-' not in col]
scaler.fit(X_train_cv[continuous_cols])
X_train_cv, X_test = scale_features(scaler, X_train_cv), scale_features(scaler, X_test)
xgb = xgboost.XGBRegressor(**model_params)
xgb.fit(X_train_cv ,y_train_cv, eval_metric = 'rmse')
# eval model
y_pred = xgb.predict(X_test)
y_cv_preds.extend(list(y_pred))
y_cv_all = y_cv_all.append(y_test)
end_xgb = xgboost.XGBRegressor(**model_params)
end_xgb.fit(X_train_all, y_train_all, eval_metric = 'rmse')
y_holdout_preds = end_xgb.predict(X_hold_test)
# y_base_train_preds = end_xgb.predict(X_base_train)
# base_train_mse = mean_squared_error(y_base_train.values, y_base_train_preds)
cv_mse = mean_squared_error(y_cv_all.values, y_cv_preds)
holdout_mse = mean_squared_error(y_hold_test.values, y_holdout_preds)
print('y-test mean {}, y-test std {}'.format(np.mean(y_test.values), np.std(y_test.values)))
print('cv mse {}, holdout mse{}'.format(cv_mse, holdout_mse))
# y_base_train_pred_df = pd.DataFrame(y_base_train_preds, columns=[y_bi_week_label], index = y_base_train.index)
# y_base_train_pred_df = y_base_train_pred_df.merge(y_to_plot[['gov_iso', 'date']], how = 'left',left_index = True, right_index = True)
# y_base_train_pred_crosstab = y_base_train_pred_df.pivot_table(index = 'date', columns = 'gov_iso', values = y_bi_week_label, aggfunc='sum')
y_cv_pred_df = pd.DataFrame(y_cv_preds, columns=[y_bi_week_label], index = y_cv_all.index)
y_cv_pred_df = y_cv_pred_df.merge(y_to_plot[['gov_iso', 'date']], how = 'left',left_index = True, right_index = True)
y_cv_pred_crosstab = y_cv_pred_df.pivot_table(index = 'date', columns = 'gov_iso', values = y_bi_week_label, aggfunc='mean')
y_holdout_pred_df = pd.DataFrame(y_holdout_preds, columns=[y_bi_week_label], index = y_hold_test.index)
y_holdout_pred_df = y_holdout_pred_df.merge(y_to_plot[['gov_iso', 'date']], how = 'left', left_index = True, right_index = True)
y_holdout_pred_crosstab = y_holdout_pred_df.pivot_table(index = 'date', columns = 'gov_iso', values = y_bi_week_label, aggfunc='mean')
true_val_pivot = y_to_plot.pivot_table(index = 'date', columns = 'gov_iso', values = y_bi_week_label, aggfunc='mean')
#print (y_base_train_pred_df.gov_iso.nunique(), y_cv_pred_df, y_holdout_pred_df)
plot_pred_against_actual(y_cv_pred_crosstab, y_holdout_pred_crosstab, true_val_pivot, y_bi_week_label, cv_mse, holdout_mse, sharey = True)
plot_pred_against_actual(y_cv_pred_crosstab, y_holdout_pred_crosstab, true_val_pivot, y_bi_week_label, cv_mse, holdout_mse, sharey = False)
plot_pred_against_actual_filtered(y_cv_pred_crosstab, y_holdout_pred_crosstab, true_val_pivot, y_bi_week_label, cv_mse, holdout_mse, sharey = True)
plot_pred_against_actual_filtered(y_cv_pred_crosstab, y_holdout_pred_crosstab, true_val_pivot, y_bi_week_label, cv_mse, holdout_mse, sharey = False)
y_cv_pred_crosstab.append(y_holdout_pred_crosstab).to_csv('/Users/Rohil/Documents/iGEM/yemen/' + y_bi_week_label + '_deployed_cv_holdout_preds.csv')
fig1, ax1 = plt.subplots(figsize = (5,8))
xgboost.plot_importance(end_xgb, ax=ax1)
fig1.savefig('/Users/Rohil/Documents/iGEM/yemen/' + y_bi_week_label + '_deployed_feature_importances.png', dpi = 300, bbox_inches = 'tight')
plt.close('all')
# class from which all other processes are called (master class)
# this is the class you initialize as the user
class Orchestrator():
def __init__(self, full_data_bi_week, y_bi_week_label):
self.full_data_bi_week = full_data_bi_week
self.y_bi_week_label = y_bi_week_label
pass
def runTsFresh(self):
print('Running TSFresh....')
X_ts = self.full_data_bi_week.drop(columns=['date', 'gov_iso', y_bi_week_label])
y_ts = self.full_data_bi_week[self.y_bi_week_label]
dateSeries_ts = self.full_data_bi_week.date
self.tf = TsFresh()
self.tf.postProcessor(X_ts, y_ts, dateSeries_ts)
print('Finished running TSFresh....')
def runOptimizeFeatures(self):
print('Running Feature Selection module ....')
y = self.full_data_bi_week[self.y_bi_week_label]
self.op = OptimizeFeatures()
self.op.selectFeatures(self.tf.selected_features, y)
self.op.identifyCollinearFeatures(0.975)
self.op.removeCollinerFeatures()
print('Original {} and after {}'.format(self.op.X.shape, self.op.corr_selected_features.shape))
print('Finished running Feature Selection ....')
def performHyperparameterOptimization(self, X, max_evals):
self.bo = BayesianOptimizer(X, self.full_data_bi_week[['date', self.y_bi_week_label]], max_evals)
self.bo.run()
def runRollingCustomFeatureExtractor(self, preselect_params, corr_selected_features):
print('Running custom feature selection module ....')
# user can specify their corr_selected_features if they please (if the job has been batched)
if corr_selected_features is None:
self.corr_selected_features_bi_week = self.op.corr_selected_features
else:
self.corr_selected_features_bi_week = corr_selected_features
self.corr_selected_features_bi_week_y = self.full_data_bi_week[['date', self.y_bi_week_label]]
#corr_selected_features_4_6 = pd.concat([full_data_4_6.date, corr_selected_features_4_6], axis=1)
#corr_selected_features_4_6.date.head()
"""
X = corr_selected_features_4_6
y = corr_selected_features_4_6_y
X_train, X_cv, X_test = train_cv_test_split(X)
y_train, y_cv, y_test = train_cv_test_split(y)
"""
self.rcfe = RollingCustomFeatureExtractor(self.corr_selected_features_bi_week,
self.corr_selected_features_bi_week_y, preselect_params)
self.rcfe.extract()
def deploy(self, threshold, model_params, selected_x):
if selected_x is None:
selected_features_from_threshold = list(self.rcfe.mcr.feature_importance_df[self.rcfe.mcr.feature_importance_df.threshold >= threshold].index)
selected_features_from_threshold.append('date')
self.X_final = self.corr_selected_features_bi_week[selected_features_from_threshold]
else:
self.X_final = selected_x
y = self.corr_selected_features_bi_week_y
#self.X_final['days_from'] = self.full_data_bi_week.days_from
y_to_plot = self.corr_selected_features_bi_week_y.merge(full_features, on = 'date', left_index=True, right_index=True, how = 'left')[[self.y_bi_week_label, 'gov_iso', 'date']]
self.dr = DeployRegressor()
self.dr.execute(model_params, self.X_final, y, y_to_plot, self.y_bi_week_label)
print (y_df.date.min())
print (y_df.date.max())
y1_2 = y_df[['date', 'gov_iso', 'week_1_to_2_cases']]
y2_4 = y_df[['date', 'gov_iso', 'week_2_to_4_cases']]
y4_6 = y_df[['date', 'gov_iso', 'week_4_to_6_cases']]
y6_8 = y_df[['date', 'gov_iso', 'week_6_to_8_cases']]
# instead of creating copies here, will run these lines directly in the object instantiation
# full_data_1_2 = y1_2.dropna().merge(full_features, how = 'left', on = ['gov_iso', 'date']).sort_values('date')
# full_data_2_4 = y2_4.dropna().merge(full_features, how = 'left', on = ['gov_iso', 'date']).sort_values('date')
# full_data_4_6 = y4_6.dropna().merge(full_features, how = 'left', on = ['gov_iso', 'date']).sort_values('date')
# full_data_6_8 = y6_8.dropna().merge(full_features, how = 'left', on = ['gov_iso', 'date']).sort_values('date')
# running feature selection for 1-2 week model
orchestrator12 = Orchestrator(full_features, full_data_1_2, 'week_1_to_2_cases' )
orchestrator12.runTsFresh()
orchestrator12.runOptimizeFeatures()
X_12_preselect = orchestrator12.op.corr_selected_features
y_12_preselect = orchestrator12.full_data_bi_week[['date', orchestrator12.y_bi_week_label]]
# X_12_preselect.to_csv('/Users/Rohil/Documents/iGEM/yemen/intermediary_files/X_12_preselect.csv')
# y_12_preselect.to_csv('/Users/Rohil/Documents/iGEM/yemen/intermediary_files/y_12_preselect.csv')
orchestrator12 = Orchestrator(y1_2.dropna().merge(full_features, how = 'left', on = ['gov_iso', 'date']).sort_values('date'), 'week_1_to_2_cases' )
# X_12_preselect = pd.read_csv('/Users/Rohil/Documents/iGEM/yemen/intermediary_files/X_12_preselect.csv', index_col = 0)
# X_12_preselect.date = pd.to_datetime(X_12_preselect.date, format = '%Y-%m-%d')
orchestrator12.performHyperparameterOptimization(X_12_preselect, 25)
orchestrator12.bo.best
preselect_params_12 = {'colsample_bytree': 0.60,
'gamma': 2.0,
'learning_rate': 0.05,
'max_depth': 11,
'min_child_weight': 10.0,
'n_estimators': 50,
'subsample': 0.8}
orchestrator12.runRollingCustomFeatureExtractor(preselect_params_12, X_12_preselect)
orchestrator12.rcfe.summary
selected_features12 = ['date'] + list(orchestrator12.rcfe.mcr.feature_importance_df[orchestrator12.rcfe.mcr.feature_importance_df.threshold >= 0.001643].index)
# orchestrator12.corr_selected_features_bi_week[selected_features12].to_csv('/Users/Rohil/Documents/iGEM/yemen/intermediary_files/X_1_2_postselect.csv')
# orchestrator12.corr_selected_features_bi_week_y.to_csv('/Users/Rohil/Documents/iGEM/yemen/intermediary_files/y_1_2.csv')
orchestrator12.performHyperparameterOptimization(X_12_preselect[selected_features12], 100)
orchestrator12.bo.best
postselect_params_12 = {'colsample_bytree': 0.75,
'gamma': 1.0,
'learning_rate': 0.05,
'max_depth': 6,
'min_child_weight': 1.0,
'n_estimators': 450,
'subsample': 0.9}
orchestrator12.deploy(0.001643, postselect_params_12, selected_x=None)
# running feature selection for 2-4 week model
orchestrator24.runTsFresh()
orchestrator24.runOptimizeFeatures()
X_24_preselect = orchestrator24.op.corr_selected_features
y_24_preselect = orchestrator24.full_data_bi_week[['date', orchestrator24.y_bi_week_label]]
# X_24_preselect.to_csv('/Users/Rohil/Documents/iGEM/yemen/intermediary_files/X_24_preselect.csv')
# y_24_preselect.to_csv('/Users/Rohil/Documents/iGEM/yemen/intermediary_files/y_24_preselect.csv')
orchestrator24 = Orchestrator(y2_4.dropna().merge(full_features, how = 'left', on = ['gov_iso', 'date']).sort_values('date'), 'week_2_to_4_cases' )
# X_24_preselect = pd.read_csv('/Users/Rohil/Documents/iGEM/yemen/intermediary_files/X_24_preselect.csv', index_col = 0)
# X_24_preselect.date = pd.to_datetime(X_24_preselect.date, format = '%Y-%m-%d')
orchestrator24.performHyperparameterOptimization(X_24_preselect, 30)
orchestrator24.bo.best
preselect_params_24 = {'colsample_bytree': 0.9,
'gamma': 3.5,
'learning_rate': 0.15,
'max_depth': 7,
'min_child_weight': 4.0,
'n_estimators': 150,
'subsample': 1.0}
orchestrator24.runRollingCustomFeatureExtractor(preselect_params_24, X_24_preselect)
orchestrator24.rcfe.summary
selected_features24 = ['date'] + list(orchestrator24.rcfe.mcr.feature_importance_df[orchestrator24.rcfe.mcr.feature_importance_df.threshold >= 0.002232].index)
orchestrator24.performHyperparameterOptimization(X_24_preselect[selected_features24], 100)
orchestrator24.bo.best
postselect_params_24 = {'colsample_bytree': 0.70,
'gamma': 2.5,
'learning_rate': 0.2,
'max_depth': 7,
'min_child_weight': 1.0,
'n_estimators': 550,
'subsample': 0.70}
# orchestrator24.corr_selected_features_bi_week[selected_features24].to_csv('/Users/Rohil/Documents/iGEM/yemen/intermediary_files/X_2_4_postselect.csv')
# orchestrator24.corr_selected_features_bi_week_y.to_csv('/Users/Rohil/Documents/iGEM/yemen/intermediary_files/y_2_4.csv')
orchestrator24.deploy(0.002232, postselect_params_24, None)
# running feature selection for 4-6 week model
orchestrator46 = Orchestrator(full_features, full_data_4_6, 'week_4_to_6_cases' )
orchestrator46.runTsFresh()
orchestrator46.runOptimizeFeatures()
X_46_preselect = orchestrator46.op.corr_selected_features
y_46_preselect = orchestrator46.full_data_bi_week[['date', orchestrator46.y_bi_week_label]]
# X_46_preselect.to_csv('/Users/Rohil/Documents/iGEM/yemen/intermediary_files/X_46_preselect.csv')
# y_46_preselect.to_csv('/Users/Rohil/Documents/iGEM/yemen/intermediary_files/y_46_preselect.csv')
orchestrator46 = Orchestrator(y4_6.dropna().merge(full_features, how = 'left', on = ['gov_iso', 'date']).sort_values('date'), 'week_4_to_6_cases' )
X_46_preselect = pd.read_csv('/Users/Rohil/Documents/iGEM/yemen/intermediary_files/X_46_preselect.csv', index_col = 0)
X_46_preselect.date = pd.to_datetime(X_46_preselect.date, format = '%Y-%m-%d')
orchestrator46.performHyperparameterOptimization(X_46_preselect, 25)
preselect_params_46 = {'colsample_bytree': 0.35,
'gamma': 4.0,
'learning_rate': 0.125,
'max_depth': 11,
'min_child_weight': 2.0,
'n_estimators': 350,
'subsample': 0.95}
orchestrator46.runRollingCustomFeatureExtractor(preselect_params_46, X_46_preselect)
selected_features46 = ['date'] + list(orchestrator46.rcfe.mcr.feature_importance_df[orchestrator46.rcfe.mcr.feature_importance_df.threshold >= 0.001844].index)
orchestrator46.performHyperparameterOptimization(X_46_preselect[selected_features46], 100)
orchestrator46.bo.best
orchestrator46.corr_selected_features_bi_week[selected_features46].to_csv('/Users/Rohil/Documents/iGEM/yemen/intermediary_files/X_4_6_postselect.csv')
orchestrator46.corr_selected_features_bi_week_y.to_csv('/Users/Rohil/Documents/iGEM/yemen/intermediary_files/y_4_6.csv')
postselect_params_46 = {'colsample_bytree': 0.60,
'gamma': 3.5,
'learning_rate': 0.025,
'max_depth': 9,
'min_child_weight': 8.0,
'n_estimators': 100,
'subsample': 0.70}
orchestrator46.deploy(0.001844, postselect_params_46, None)
# running feature selection for 6-8 week model
orchestrator68 = Orchestrator(full_features, full_data_6_8, 'week_6_to_8_cases' )
orchestrator68.runTsFresh()
orchestrator68.runOptimizeFeatures()
X_68_preselect = orchestrator68.op.corr_selected_features
y_68_preselect = orchestrator68.full_data_bi_week[['date', orchestrator68.y_bi_week_label]]
# X_68_preselect.to_csv('/Users/Rohil/Documents/iGEM/yemen/intermediary_files/X_68_preselect.csv')
# y_68_preselect.to_csv('/Users/Rohil/Documents/iGEM/yemen/intermediary_files/y_68_preselect.csv')
orchestrator68 = Orchestrator(y6_8.dropna().merge(full_features, how = 'left', on = ['gov_iso', 'date']).sort_values('date'), 'week_6_to_8_cases' )
# X_68_preselect = pd.read_csv('/Users/Rohil/Documents/iGEM/yemen/intermediary_files/X_68_preselect.csv', index_col = 0)
# X_68_preselect.date = pd.to_datetime(X_68_preselect.date, format = '%Y-%m-%d')
orchestrator68.performHyperparameterOptimization(X_68_preselect, 20)
orchestrator68.bo.best
preselect_params_68= {'colsample_bytree': 0.45,
'gamma': 3.0,
'learning_rate': 0.152,
'max_depth': 7,
'min_child_weight': 10.0,
'n_estimators': 250,
'subsample': 0.70}
orchestrator68.runRollingCustomFeatureExtractor(preselect_params_68, X_68_preselect)
orchestrator68.rcfe.summary
selected_features68 = ['date'] + list(orchestrator68.rcfe.mcr.feature_importance_df[orchestrator68.rcfe.mcr.feature_importance_df.threshold >= 0.001816].index)
orchestrator68.performHyperparameterOptimization(X_68_preselect[selected_features68], 100)
orchestrator68.bo.best
orchestrator68.corr_selected_features_bi_week[selected_features68].to_csv('/Users/Rohil/Documents/iGEM/yemen/intermediary_files/X_6_8.csv')
orchestrator68.corr_selected_features_bi_week_y.to_csv('/Users/Rohil/Documents/iGEM/yemen/intermediary_files/y_6_8.csv')
postselect_params_68 = {'colsample_bytree': 0.4,
'gamma': 0.0,
'learning_rate': 0.025,
'max_depth': 8,
'min_child_weight': 1.0,
'n_estimators': 150,
'subsample': 0.9}
orchestrator68.deploy(0.001815, postselect_params_68, None)
```
| github_jupyter |
# Sea Surface Altimetry Data Analysis
<img src="http://marine.copernicus.eu/documents/IMG/SEALEVEL_GLO_SLA_MAP_L4_REP_OBSERVATIONS_008_027.png"
width="15%"
align=left
alt="Globe">
For this example we will use gridded [sea-surface altimetry data from The Copernicus Marine Environment](http://marine.copernicus.eu/services-portfolio/access-to-products/?option=com_csw&view=details&product_id=SEALEVEL_GLO_PHY_L4_REP_OBSERVATIONS_008_047):
This is a widely used dataset in physical oceanography and climate.
The dataset has already been extracted from copernicus and stored in google cloud storage in [xarray-zarr](http://xarray.pydata.org/en/latest/io.html#zarr) format.
```
%matplotlib inline
import numpy as np
import xarray as xr
import matplotlib.pyplot as plt
import dask.array as dsa
plt.rcParams['figure.figsize'] = (15,10)
```
### Initialize Dataset
Here we load the dataset from the zarr store. Note that this very large dataset initializes nearly instantly, and we can see the full list of variables and coordinates.
```
import intake
cat = intake.Catalog("https://raw.githubusercontent.com/pangeo-data/pangeo-datastore/master/intake-catalogs/ocean.yaml")
ds = cat["sea_surface_height"].to_dask()
ds
```
### Examine Metadata
For those unfamiliar with this dataset, the variable metadata is very helpful for understanding what the variables actually represent
```
for v in ds.data_vars:
print('{:>10}: {}'.format(v, ds[v].attrs['long_name']))
```
## Visually Examine Some of the Data
Let's do a sanity check that the data looks reasonable:
```
plt.rcParams['figure.figsize'] = (15, 8)
ds.sla.sel(time='2000-01-01', method='nearest').plot()
```
### Same thing using interactive graphics
```
import holoviews as hv
from holoviews.operation.datashader import regrid
hv.extension('bokeh')
dataset = hv.Dataset(ds.sla)
hv_im = (dataset.to(hv.Image, ['longitude', 'latitude'], dynamic=True)
.redim.range(sla=(-0.5, 0.5))
.options(cmap='RdBu_r', width=800, height=450, colorbar=True))
%output holomap='scrubber' fps=2
regrid(hv_im, precompute=True)
```
### Create and Connect to Dask Distributed Cluster
```
from dask.distributed import Client, progress
from dask_kubernetes import KubeCluster
cluster = KubeCluster()
cluster.adapt(minimum=4, maximum=40)
cluster
```
** ☝️ Don't forget to click the link above to view the scheduler dashboard! **
```
client = Client(cluster)
client
```
## Timeseries of Global Mean Sea Level
Here we make a simple yet fundamental calculation: the rate of increase of global mean sea level over the observational period.
```
# the number of GB involved in the reduction
ds.sla.nbytes/1e9
# the computationally intensive step
sla_timeseries = ds.sla.mean(dim=('latitude', 'longitude')).load()
sla_timeseries.plot(label='full data')
sla_timeseries.rolling(time=365, center=True).mean().plot(label='rolling annual mean')
plt.ylabel('Sea Level Anomaly [m]')
plt.title('Global Mean Sea Level')
plt.legend()
plt.grid()
```
In order to understand how the sea level rise is distributed in latitude, we can make a sort of [Hovmöller diagram](https://en.wikipedia.org/wiki/Hovm%C3%B6ller_diagram).
```
sla_hov = ds.sla.mean(dim='longitude').load()
fig, ax = plt.subplots(figsize=(12, 4))
sla_hov.name = 'Sea Level Anomaly [m]'
sla_hov.transpose().plot(vmax=0.2, ax=ax)
```
We can see that most sea level rise is actually in the Southern Hemisphere.
## Sea Level Variability
We can examine the natural variability in sea level by looking at its standard deviation in time.
```
sla_std = ds.sla.std(dim='time').load()
sla_std.name = 'Sea Level Variability [m]'
ax = sla_std.plot()
_ = plt.title('Sea Level Variability')
```
## Spectral Analysis
This is an advanced, research-grade example. Here we perform wavenumber-frequency spectral analysis of the SSH signal using methods similar to those described in [Abernathey & Wortham (2015)](https://journals.ametsoc.org/doi/10.1175/JPO-D-14-0160.1).
#### Step 1: Extract a sector in the Eastern Pacific
This sector is chosen because it has very little land.
```
sector = ds.sla.sel(longitude=slice(180, 230), latitude=slice(-70, 55, 4))
sector_anom = (sector - sector.mean(dim='longitude'))
sector_anom
sector_anom[0].plot()
```
#### Step 2: Rechunk, reshape, and window the data for efficient to prepare for FFT calculation
```
# reshape data into arrays 365 days long and rechunk
nsegments = 24
segment_len = 365
sector_reshape = (sector_anom.isel(time=slice(0, nsegments*segment_len))
.transpose('latitude', 'time', 'longitude')
.chunk({'time': segment_len}))
sector_reshape
# now get the raw dask array
data = sector_reshape.data
arrays = [data[:, n*segment_len:(n + 1)*segment_len][np.newaxis]
for n in range(nsegments)]
stacked = dsa.concatenate(arrays)
stacked
# apply windows
data_windowed = (stacked
* np.hanning(stacked.shape[-1])[None, None, None, :]
* np.hanning(stacked.shape[-2])[None, None, :, None])
```
#### Step 3: Actually calculate the Fourier transform and power spectral density
```
# take FFT
data_fft = dsa.fft.fftn(data_windowed, axes=(-2, -1))
# take power spec and average over segments
power_spec = np.real(data_fft * np.conj(data_fft)).mean(axis=0)
power_spec
# do the computation and load results into memory
power_spec_shift = np.fft.fftshift(power_spec.compute(), axes=(-2, -1))
```
#### Step 4: Define spectral coordinates and put everything back together in an DataArray
```
freq = np.fft.fftshift(np.fft.fftfreq(segment_len))
# wavelength is a bit trickier because it depends on latitude
R = 6.37e6
# in km
dx = np.deg2rad(0.25) * R * np.cos(np.deg2rad(sector.latitude)) / 1000
inv_wavelength = np.vstack([np.fft.fftshift(np.fft.fftfreq(len(sector.longitude), d))
for d in dx.values])
ps_da = xr.DataArray(power_spec_shift, dims=('latitude', 'freq', 'wavenumber'),
coords={'latitude': sector.latitude,
'freq': ('freq', -freq, {'units': r'days$^{-1}$'}),
'inverse_wavelength': (('latitude', 'wavenumber'),
inv_wavelength, {'units': r'km$^{-1}$'})},
name='SSH_power_spectral_density')
ps_da
```
#### Step 5: Plot wavenumber-frequency power spectra at different latitudes
```
from matplotlib.colors import LogNorm
for lat in range(-55, 55, 10):
plt.figure()
(ps_da.sel(latitude=lat, method='nearest')
.swap_dims({'wavenumber': 'inverse_wavelength'})
.transpose().plot(norm=LogNorm()))
```
After going through all that complexity, you might be interested to know that there is a library which facilitaties spectral analysis of xarray datasets:
- https://xrft.readthedocs.io/en/latest/
With xrft, we could have reduced all the steps above to a few lines of code. But we would not have learned as much! 😜
| github_jupyter |
# K-Nearest-Neighbors
KNN falls in the supervised learning family of algorithms. Informally, this means that we are given a labelled dataset consiting
of training observations (x,y) and would like to capture the relationship between x and y. More formally, our goal is to learn a
function h:X→Y so that given an unseen observation x, h(x) can confidently predict the corresponding output y.
In this module we will explore the inner workings of KNN, choosing the optimal K values and using KNN from scikit-learn.
# Overview
1.Read the problem statement.
2.Get the dataset.
3.Explore the dataset.
4.Pre-processing of dataset.
5.Visualization
6.Transform the dataset for building machine learning model.
7.Split data into train, test set.
7.Build Model.
8.Apply the model.
9.Evaluate the model.
10.Finding Optimal K value
11.Repeat 7,8,9 steps.
# Problem statement
Dataset:
The data set we’ll be using is the Iris Flower Dataset which was first introduced in 1936 by the famous statistician Ronald
Fisher and consists of 50 observations from each of three species of Iris (Iris setosa, Iris virginica and Iris versicolor).
Four features were measured from each sample: the length and the width of the sepals and petals.
Source: https://archive.ics.uci.edu/ml/datasets/Iris
Train the KNN algorithm to be able to distinguish the species from one another given the measurements of the 4 features.
# Question 1
Import libraries and load the dataset
```
import warnings
warnings.filterwarnings('ignore')
from sklearn.model_selection import cross_validate
import pandas as pd
#Data setup
df = pd.read_csv('iris-1.csv')
df.head()
```
**Shape**
```
df.shape
df.dtypes
df.info()
```
# Data Pre-processing
# Question 2 - Imputing missing values
Its not good to remove the records having missing values all the time. We may end up loosing some data points. So, we will have to see how to replace those missing values with some estimated values (mean or median).
```
df.isnull().sum()
df.boxplot(vert=0)
df["SepalLengthCm"].fillna(df["SepalLengthCm"].mean(),inplace=True) # No Outliers so imputed the null value with mean
df["PetalLengthCm"].fillna(df["PetalLengthCm"].mean(),inplace=True) # No Outliers so imputed the null value with mean
df["PetalWidthCm"].fillna(df["PetalWidthCm"].mean(),inplace=True) # No Outliers so imputed the null value with mean
df["SepalWidthCm"].fillna(df["SepalWidthCm"].median(),inplace=True) # Outliers are present so imputed the null value with median
df.isnull().sum()
```
# Question 3 - Dealing with categorical data
Change all the classes to numericals (0to2) with the help of **pd.Categorical().codes** and drop ID as it is insignificant and does not contain any information
```
df["Species"].value_counts()
df["Species"]=pd.Categorical(df["Species"]).codes
df["Species"].value_counts()
df.drop("Id",axis=1,inplace=True)
```
# Question 4
Observe the association of each independent variable with target variable by checking the .corr() and make a heatmap/correlation plot
```
df.corr()
```
**Heatmap**
```
sns.heatmap(df.corr(),annot=True)
```
# Question 5
Observe the independent variables variance and check for variables having no variance or almost zero variance(variance < 0.1).
They will be having almost no influence on the classification.
```
df.var()
```
# Question 6
Plot the pairplot with hue="Species for the data.
```
sns.pairplot(df,hue="Species")
```
# Split the dataset into training and test sets
# Question 7
Split the dataset into training and test sets with 70-30 ratio.
```
# Arrange data into independent variables and dependent variables
X = df.drop("Species",axis=1) ## Features
y = df["Species"] ## Target
# Split X and y into training and test set in 70:30 ratio
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.80 , random_state=20)
```
# Question 8 - Model
Build the model. Print the Accuracy of the model with different values of k=3,5,9.
Hint: For accuracy use model.score()
```
# loading library
from sklearn.neighbors import KNeighborsClassifier
# initiantiate learning model (k = 3)
KNN_model=KNeighborsClassifier(n_neighbors = 3,metric='euclidean')
# fitting the model
KNN_model.fit(X_train,y_train)
# predict the response
y_test_predict = KNN_model.predict(X_test)
# evaluate accuracy
print("Accuracy Score for K=3 is ", KNN_model.score(X_test, y_test))
# initiantiate learning model (k = 5)
KNN_model=KNeighborsClassifier(n_neighbors = 5,metric='euclidean')
# fitting the model
KNN_model.fit(X_train,y_train)
# predict the response
y_test_predict = KNN_model.predict(X_test)
# evaluate accuracy
print("Accuracy Score for K=5 is ", KNN_model.score(X_test, y_test))
# initiantiate learning model (k = 9)
from sklearn.neighbors import KNeighborsClassifier
KNN_model=KNeighborsClassifier(n_neighbors = 9,metric='euclidean')
# fitting the model
KNN_model.fit(X_train,y_train)
# predict the response
y_test_predict = KNN_model.predict(X_test)
# evaluate accuracy
print("Accuracy Score for K=9 is ", KNN_model.score(X_test, y_test))
```
# Question 9 - Optimal no. of K
Run the KNN with no of neighbours to be 1,3,5..19 and *Find the optimal number of neighbours from the above list using the Mis classification error
Hint:
Misclassification error (MCE) = 1 - Test accuracy score. Calculated MCE for each model with neighbours = 1,3,5...19 and find the
model with lowest MCE
```
# empty list that will hold accuracy scores
ac_scores = []
# perform accuracy metrics for values from 1,3,5....19
for k in range(1,20,2):
knn = KNeighborsClassifier(n_neighbors=k)
knn.fit(X_train, y_train)
# evaluate accuracy
scores = knn.score(X_test, y_test)
ac_scores.append(scores)
# changing to misclassification error
MCE = [1 - x for x in ac_scores]
MCE
```
# Question 10
Plot misclassification error vs k (with k value on X-axis) using matplotlib.
```
import matplotlib.pyplot as plt
# plot misclassification error vs k
plt.plot(range(1,20,2), MCE)
plt.xlabel('Number of Neighbors K')
plt.ylabel('Misclassification Error')
plt.show()
```
At K=3
| github_jupyter |
<img align="right" style="max-width: 200px; height: auto" src="https://github.com/HSG-AIML/LabAIML/blob/main/lab_03/hsg_logo.png?raw=1">
## Lab 03 - "Machine Learning with scikit-Learn"
Introduction to AI and ML, University of St. Gallen, Fall Term 2021
During the last two labs you learned the basics of Python programming. Today, we will start to use these skills to implement our first machine learning models. For this purpose, we will utilize the [`scikit-learn`](https://scikit-learn.org/stable/index.html) package, which provides a huge amount of functionality for different machine learning tasks, as well as some datasets for learning how to use this functionality.
In this Notebook we will focus on implementing **supervised machine learning** methods, but be assured that utilizing **unsupervised** methods from `scikit-learn` is just as simple.
As always, pls. don't hesitate to ask all your questions either during the lab, post them in our CANVAS (StudyNet) forum (https://learning.unisg.ch), or send us an email (using the course email).
## Lab Objectives:
Your learning objectives for today are threefold:
> 1. Learn how to load, analyze and utilize **scikit-learn datasets**.
> 2. Implement and train different **supervised classifier methods**.
> 3. **Evaluate the results** of your trained model.
## 0. Setup of the Environment
Similarly to the previous labs, we need to import a couple of Python libraries that allow for data analysis and data visualization. In this lab will use the `Numpy`, `scikit-learn`, `Matplotlib` and the `Seaborn` library. Let's import the libraries by the execution of the statements below:
```
# import the numpy, scipy and pandas data science library
import numpy as np
# import sklearn data and data pre-processing libraries
from sklearn import datasets
from sklearn.model_selection import train_test_split
# import matplotlib data visualization library
import matplotlib.pyplot as plt
import seaborn as sns
```
Enable inline Jupyter notebook plotting:
```
%matplotlib inline
```
Use the 'Seaborn' plotting style in all subsequent visualizations:
```
plt.style.use('seaborn')
```
Set random seed of all our experiments - this insures reproducibility.
```
random_seed = 42
```
# 1. Data
The **Iris Dataset** is a classic and straightforward dataset often used as a "Hello World" example in multi-class classification. This data set consists of measurements taken from three different types of iris flowers (referred to as **Classes**), namely the Iris Setosa, the Iris Versicolour and the Iris Virginica, and their respective measured petal and sepal length (referred to as **Features**).
<img align="center" style="max-width: 700px; height: auto" src="https://github.com/HSG-AIML/LabAIML/blob/main/lab_03/iris_dataset.png?raw=1">
(Source: http://www.lac.inpe.br/~rafael.santos/Docs/R/CAP394/WholeStory-Iris.html)
In total, the dataset consists of **150 samples** (50 samples taken per class) as well as their corresponding **4 different measurements** taken for each sample. Please, find below the list of the individual measurements:
>- `Sepal length (cm)`
>- `Sepal width (cm)`
>- `Petal length (cm)`
>- `Petal width (cm)`
Further details of the dataset can be obtained from the following puplication: *Fisher, R.A. "The use of multiple measurements in taxonomic problems" Annual Eugenics, 7, Part II, 179-188 (1936); also in "Contributions to Mathematical Statistics" (John Wiley, NY, 1950)."*
Let's load the dataset and conduct a preliminary data assessment:
## 1.1 Dataset Download and Data Exploration
```
iris = datasets.load_iris()
```
How is the dataset structured?
```
iris.keys()
```
Let's have a look at the dataset description:
```
print(iris.DESCR)
```
Print and inspect the names of the four features contained in the dataset:
```
iris.feature_names
```
Determine and print the feature dimensionality of the dataset:
```
iris.data.shape
```
Determine and print the class label dimensionality of the dataset:
```
iris.target.shape
```
Print and inspect the names of the three classes contained in the dataset:
```
iris.target_names
```
Let's briefly envision how the feature information of the dataset is collected and presented in the data:
<img align="center" style="max-width: 900px; height: auto" src="https://github.com/HSG-AIML/LabAIML/blob/main/lab_03/feature_collection.png?raw=1">
Let's inspect the top five feature rows of the Iris Dataset:
```
iris.data[:5,]
```
Let's also inspect the top five class labels of the Iris Dataset:
```
iris.target[:5]
```
## 1.2 Dataset Exploration
Let's now conduct a more in depth data assessment. Therefore, we plot the feature distributions of the Iris dataset according to their respective class memberships as well as the features pairwise relationships.
Pls. note that we use Python's **Seaborn** library to create such a plot referred to as **Pairplot**. The Seaborn library is a powerful data visualization library based on the Matplotlib. It provides a great interface for drawing informative statstical graphics (https://seaborn.pydata.org).
```
# init the plot
plt.figure(figsize=(10, 10))
# load the dataset also available in seaborn
iris_plot = sns.load_dataset("iris")
# plot a pairplot of the distinct feature distributions
sns.pairplot(iris_plot, diag_kind='hist', hue='species');
```
It can be observed from the created Pairplot, that most of the feature measurements that correspond to flower class "setosa" exhibit a nice **linear separability** from the feature measurements of the remaining flower classes. In addition, the flower classes "versicolor" and "virginica" exhibit a commingled and **non-linear separability** across all the measured feature distributions of the Iris Dataset.
## 1.3 Dataset Pre-Processing
# 1.3.1 Data Normalization
As you can see in the plot above, the diffferent features span different ranges: *sepal_length* ranges from 4 to 8, *sepal_width* from 2 to 4 and so on. Those different ranges might affect the learning process negatively.
In the case of $k$-Nearest Neighbor classification as we implement it below, the nearest neighbors are identified based on a distance measure. If the features are differently scaled, they are not considered equally important by the model. Consider the following example: the max-min distance in *sepal_length* is 4, whereas the same for *sepal_width* is only 2. When calculating the Euclidean distance between two datapoints, the contribution to the distance from *sepal_length* is likely to be larger than that from *sepal_width*. As a result, *sepal_length* has much more power and turns out to be more decisive in the resulting distance measure and classification results.
To avoid this problem, one typically scales or normalizes the data in a uniform way. One way would be to scale the values in each feature relative to their maximum and minimum values, called the **min-max-scaler**:
$$\mathbf{x'} = \frac{\mathbf{x}-min(\mathbf{x})}{max(\mathbf{x})-min(\mathbf{x})}$$
Since the minimum and maximum values might be affected by outliers, a safer way would be to scale the data and the mean and standard deviation of the sample, which is sometimes called a **standard scaler**:
$$\mathbf{x'} = \frac{\mathbf{x}-mean(\mathbf{x})}{\sigma(\mathbf{x})}$$
The standard scaler forces upon the values in each feature a mean of unity and a spread that is based on the variance in the dataset.
Let's use the standard scaler implemented in scikit-learn to scale our data. We have to import the correponding class and initialize it.
```
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
```
To apply the scaler and retrieve a transformed dataset, we can use the `.fit_transform()` method (the name might sound weird here, but will make sense later):
```
data_scaled = scaler.fit_transform(iris.data)
print('original data, mean =', np.mean(iris.data, axis=0))
print('original data, std =', np.std(iris.data, axis=0))
print('scaled data, mean =', np.mean(data_scaled, axis=0))
print('scaled data, std =', np.std(data_scaled, axis=0))
```
We can now use the scaled data in our machine learning model.
Hint: If you would like to undo the scaling tranformation, you can use the `.inverse_transform()` method of `scaler`:
```
print('original data:', iris.data[0,])
print('scaled data:', data_scaled[0,])
print('unscaled data:', scaler.inverse_transform(data_scaled[0,]))
```
### 1.3.2 Train, Validation and Test Datasets
To understand and evaluate the performance of any trained **supervised machine learning** model, it is good practice to divide the dataset into a **training dataset** (the fraction of data records solely used for training purposes), a **validation dataset** (data to evaluate the current settings of your hyperparameters) and a **test dataset** (the fraction of data records solely used for independent evaluation purposes). Please note that both the **validation dataset** and the **test dataset** will never be shown to the model as part of the training process. The **test dataset** is sometimes also referred to as **evaluation set**; both terms refer to the same concept.
We first split our scaled dataset into a training dataset and some other dataset (which we will refer to as *remainder* in the following) that will then be evenly split into a validation and test dataset. We set the fraction of training records to **60%** of the original dataset:
```
train_fraction = 0.6
```
Randomly split the scaled dataset into training set and evaluation set using sklearn's `train_test_split` function:
```
# 60% training and 40% remainder
x_train, x_remainder, y_train, y_remainder = train_test_split(data_scaled, iris.target, test_size=1-train_fraction,
random_state=random_seed, stratify=iris.target)
# 50% validation and 50% test
x_val, x_test, y_val, y_test = train_test_split(x_remainder, y_remainder, test_size=0.5,
random_state=random_seed, stratify=y_remainder)
```
Note the use of the `stratify` keyword argument here: a stratified split makes sure that approximately the same fraction of samples from each class is present in each dataset. Therefore, we have to provide the same list of class labels to this argument.
Evaluate the different dataset dimensionalities:
```
print('original:', iris.data.shape, iris.target.shape)
print('train:', x_train.shape, y_train.shape)
print('val:', x_val.shape, y_val.shape)
print('test:', x_test.shape, y_test.shape)
```
# 2. Machine Learning
Our goal for the Iris dataset is to be able to "predict" for each point in the dataset its class. This process is typically referred to as **classification** and is a standard task in Machine Learning (ML).
As we already mentioned, we will utilize the `scikit-learn` package, which contains implementations of many different ML methods and tasks, both of supervised and unsupervised nature.
Using `scikit-learn` is simple and its API is consistent throughout many different tasks and applications. We therefore discuss how to setup a machine learning pipeline with `scikit-learn` based on a simple $k$-nearest neigbhor classifier. Later we show that implementing other models is just as simple.
## 2.1 $k$-Nearest Neighbor Classification
The idea is simple: we assign to each unseen datapoint the same class that its $k$ nearest neighbors have. This method is intuitive and straightforward. It requires the definition of hyperparameter $k$ and the definition of a distance metric. Let's have a look at he `scikit-learn` implementation:
```
from sklearn.neighbors import KNeighborsClassifier
#KNeighborsClassifier?
```
Before we can use the method, we have to instantiate it by assigning the appropriate hyperparameters. We choose $k=3$ as our initial guess (and use the default `p=2`, utilizing the Euclidean distance metric):
```
model = KNeighborsClassifier(n_neighbors=3)
```
## 2.2 Model Training
Now we have to train the model on our training dataset. Each model implemented in `scikit-learn` has a `.fit()` method for this purpose. "Fitting" refers here to the same idea that we typically refer to as "training", so don't get confused.
The training of the data requires two `arrays`: the training input features ($\mathbf{X}$) and the training target vector ($\mathbf{y}$), such that for a given classifier $f$ the following holds:
$$f(\mathbf{X}) = \mathbf{y}$$
The way we split our dataset and into `x_train` and `y_train` already follows this naming convention. We can use those `arrays` readily in the training. Just for reference: `x_train` has to be of shape `(n_samples, n_features)` and `y_train` has to be of shape `(n_samples,)`.
```
model.fit(x_train, y_train)
```
`model` is now trained and can be used to make predictions. Let's take one datapoint from our training dataset and see whether it makes a correct prediction:
```
model.predict([x_train[0]])
y_train[0]
```
Indeed, it classifies this single datapoint correctly. However, this is not a good way to test or evaluate the performance of your model. Why?
# 2.3 Model Evaluation
Of course, we should use our previously split test sample for evaluating our model performance:
```
y_pred = model.predict(x_test)
y_pred
y_test
```
A quick by-eye check seems to look pretty promising, but of course we need a more quantitative metric for the performance of our model.
In the case of classification, we can use the accuracy metric:
```
from sklearn.metrics import accuracy_score
accuracy_score(y_pred, y_test)
```
This is it. After evaluation on our independent eval dataset - which the model has not seen during training - we find that our model makes an accurate prediction in 93.3% of cases.
This could be it, but there is a good chance that by tuning our sole hyperparameter, $k$, we can achieve a better result.
## 2.4 Hyperparameter Tuning
Let's compile all the relevant code in one cell and try a different value for $k$:
```
model = KNeighborsClassifier(n_neighbors=1)
model.fit(x_train, y_train)
y_pred = model.predict(x_test)
accuracy_score(y_pred, y_test)
```
We can now use a loop over different choices for $k$ and evaluate the model for these parameters to find the best-performing one. This process is called **hyperparameter tuning**.
However, there is one more technical detail. Currently, we evaluate the performance on our **test dataset**. If we select $k$ based on these evaluations and therefore the **test dataset**, we have a *data leakage*. To resolve that issue, we can evaluate our model on the **validation dataset** for different $k$s and then, after picking the best-performing $k$, we can evaluate that model on the **test dataset**, providing an independent measure of performance.
```
for k in [1, 3, 5, 7, 10, 15, 20]:
model = KNeighborsClassifier(n_neighbors=k)
model.fit(x_train, y_train)
y_pred = model.predict(x_val)
print('k={:d}, val accuracy={:.2f}%'.format(k, accuracy_score(y_pred, y_val)*100))
```
It seems that the model performs equally well for $k\sim3$ and $k\sim10$. Based on experience, I would pick $k=10$. Why? For small values of $k$, you are more likely to overfit the training data, so choosing a larger value of $k$ increases the chances that the model generalizes well to data it has never seen before.
Let's retrain the model with $k=10$ and we're done:
```
k = 10
model = KNeighborsClassifier(n_neighbors=k)
model.fit(x_train, y_train)
y_pred = model.predict(x_test)
print('final test accuracy={:.2f}%'.format(accuracy_score(y_pred, y_test)*100))
```
Indeed, evaluating the model on the test dataset provides the same accuracy as for the validation dataset - but this is not always the case, since both datsets are different from one another.
## 2.5 Confusion Matrix
So far, we have only considered the accuracy metric to evaluate our predictions. It would be useful to know whether one class of flower is more likely to be mistaken than another. For that purpose, confusion matrizes are used:
```
from sklearn.metrics import confusion_matrix
mat = confusion_matrix(y_test, y_pred)
mat
```
On the y-axis you have your predicted classes, on the x-axis you have the actual classes. Those entries on the diagonal have been accurately predicted. Entries off the diagonal indicate how many flowers have incorrect class predictions.
The ``seaborn`` library provides a method to generate nicely formatted confusion matrizes. Let's give it a try:
```
# init the plot
plt.figure(figsize=(7, 7))
# plot confusion matrix heatmap
sns.heatmap(mat, square=True, annot=True, fmt='d', cbar=False, cmap='Reds',
xticklabels=iris.target_names, yticklabels=iris.target_names)
# add plot axis labels
plt.xlabel('Ground Truth')
plt.ylabel('Prediction')
# add plot title
plt.title('Confusion Matrix')
```
## 2.6 More Models
We found that a $k$-Nearest Neighbor classifier performs rather well on the Iris dataset. But what if we would like to try a different ML method?
With `scikit-learn`, this is very easy to do. Since all algorithms (ML methods, scalers, etc.) are implemented as classes and thus provide `.fit()` and `.transform()` methods, we can simply replace them!
Consider the following example. We use the exact same code that we use in tuning the $k$NN hyperparameters, but we replace the model with a **linear support vector machine** classifier implementation. Here, we replace the single hyperparameter $K$ with the regularization parameter $C$:
```
from sklearn.svm import LinearSVC
for C in [1e0, 1e1, 1e2, 1e3, 1e4]:
model = LinearSVC(C=C)
model.fit(x_train, y_train)
y_pred = model.predict(x_val)
print('C={}, val accuracy={:.2f}%'.format(C, accuracy_score(y_pred, y_val)*100))
```
Now, let's choose a **random forest** classifier. Here, we use two different hyperparameters: the number of trees in the ensemble and the maximum depth of the individual trees:
```
from sklearn.ensemble import RandomForestClassifier
for n in [10, 50, 100, 200, 500]:
for d in [3, 7, 12]:
model = RandomForestClassifier(n_estimators=n, max_depth=d)
model.fit(x_train, y_train)
y_pred = model.predict(x_val)
print('n={:3d}, d={:2d}, val accuracy={:.2f}%'.format(n, d, accuracy_score(y_pred, y_val)*100))
```
The results are very comparable, but the point is simply that with `scikit-learn`, you can pick any of the implemented methods and easily implement and try them in your code!
| github_jupyter |
```
!apt-get install -y -qq software-properties-common python-software-properties module-init-tools
!add-apt-repository -y ppa:alessandro-strada/ppa 2>&1 > /dev/null
!apt-get update -qq 2>&1 > /dev/null
!apt-get -y install -qq google-drive-ocamlfuse fuse
from google.colab import auth
auth.authenticate_user()
from oauth2client.client import GoogleCredentials
creds = GoogleCredentials.get_application_default()
import getpass
!google-drive-ocamlfuse -headless -id={creds.client_id} -secret={creds.client_secret} < /dev/null 2>&1 | grep URL
vcode = getpass.getpass()
!echo {vcode} | google-drive-ocamlfuse -headless -id={creds.client_id} -secret={creds.client_secret}
!mkdir -p drive
!google-drive-ocamlfuse drive
!ls
cd drive/Houston/
import numpy as np
import cv2
area_hsi = []
for i in range(3):
for j in range(14):
img_path = "area/pc_area_" + str(i) + "_" + str(j) + ".png"
img = cv2.imread(img_path, cv2.IMREAD_GRAYSCALE)
area_hsi.append(img)
area_hsi = np.array(area_hsi)
area_lidar = []
for i in range(14):
img_path = "area/lidar_area_" + str(i) + ".png"
img = cv2.imread(img_path, cv2.IMREAD_GRAYSCALE)
area_lidar.append(img)
area_lidar = np.array(area_lidar)
print(area_hsi.shape)
print(area_lidar.shape)
train_file_name = "labels/train.txt"
test_file_name = "labels/test.txt"
file = open(train_file_name)
triplets = file.read().split()
for i in range(0, len(triplets)):
triplets[i] = triplets[i].split(",")
train_array = np.array(triplets, dtype=int)
file.close()
file = open(test_file_name)
triplets = file.read().split()
for i in range(0, len(triplets)):
triplets[i] = triplets[i].split(",")
test_array = np.array(triplets, dtype=int)
file.close()
HEIGHT = train_array.shape[0]
WIDTH = train_array.shape[1]
area_hsi_train_data = []
area_hsi_test_data = []
area_lidar_train_data = []
area_lidar_test_data = []
moment_hsi_train_data = []
moment_hsi_test_data = []
moment_lidar_train_data = []
moment_lidar_test_data = []
train_labels = []
test_labels = []
for i in range(HEIGHT):
for j in range(WIDTH):
if train_array[i, j] != 0:
area_hsi_train_data.append(area_hsi[:, i, j])
area_lidar_train_data.append(area_lidar[:, i, j])
train_labels.append(train_array[i, j])
if test_array[i,j] != 0:
area_hsi_test_data.append(area_hsi[:, i, j])
area_lidar_test_data.append(area_lidar[:, i, j])
test_labels.append(test_array[i, j])
area_hsi_train_data = np.array(area_hsi_train_data)
area_hsi_test_data = np.array(area_hsi_test_data)
area_lidar_train_data = np.array(area_lidar_train_data)
area_lidar_test_data = np.array(area_lidar_test_data)
train_labels = np.array(train_labels)
test_labels = np.array(test_labels)
print(area_hsi_train_data.shape)
print(area_hsi_test_data.shape)
print(area_lidar_train_data.shape)
print(area_lidar_test_data.shape)
print(train_labels.shape)
print(test_labels.shape)
import keras
train_one_hot = keras.utils.to_categorical(train_labels-1)
test_one_hot = keras.utils.to_categorical(test_labels-1)
print(train_one_hot.shape)
print(test_one_hot.shape)
HSI_PATCH_SIZE = 27
LiDAR_PATCH_SIZE = 41
CONV1 = 500
CONV2 = 100
FC1 = 200
FC2 = 84
LEARNING_RATE = 0.005
BATCH_SIZE = 25
import numpy as np
padded_area_hsi = np.lib.pad(area_hsi, ((0,0), (HSI_PATCH_SIZE//2, HSI_PATCH_SIZE//2), (HSI_PATCH_SIZE//2,HSI_PATCH_SIZE//2)), 'reflect')
padded_area_lidar = np.lib.pad(area_lidar, ((0,0), (LiDAR_PATCH_SIZE//2, LiDAR_PATCH_SIZE//2), (LiDAR_PATCH_SIZE//2,LiDAR_PATCH_SIZE//2)), 'reflect')
print(padded_area_hsi.shape)
print(padded_area_lidar.shape)
def get_patches(data, patch_size, row, column):
offset = patch_size // 2
row_low = row - offset
row_high = row + offset
col_low = column - offset
col_high = column + offset
return data[0:, row_low:row_high + 1, col_low:col_high + 1].reshape(patch_size, patch_size, data.shape[0])
area_hsi_train_patches = []
area_hsi_test_patches = []
area_lidar_train_patches = []
area_lidar_test_patches = []
for i in range(HEIGHT):
for j in range(WIDTH):
if train_array[i, j] != 0:
area_hsi_train_patches.append(get_patches(padded_area_hsi, HSI_PATCH_SIZE, i+HSI_PATCH_SIZE//2, j+HSI_PATCH_SIZE//2))
area_lidar_train_patches.append(get_patches(padded_area_lidar, LiDAR_PATCH_SIZE, i+LiDAR_PATCH_SIZE//2, j+LiDAR_PATCH_SIZE//2))
if test_array[i, j] != 0:
area_hsi_test_patches.append(get_patches(padded_area_hsi, HSI_PATCH_SIZE, i+HSI_PATCH_SIZE//2, j+HSI_PATCH_SIZE//2))
area_lidar_test_patches.append(get_patches(padded_area_lidar, LiDAR_PATCH_SIZE, i+LiDAR_PATCH_SIZE//2, j+LiDAR_PATCH_SIZE//2))
area_hsi_train_patches = np.array(area_hsi_train_patches)
area_hsi_test_patches = np.array(area_hsi_test_patches)
area_lidar_train_patches = np.array(area_lidar_train_patches)
area_lidar_test_patches = np.array(area_lidar_test_patches)
print(area_hsi_train_patches.shape)
print(area_hsi_test_patches.shape)
print(area_lidar_train_patches.shape)
print(area_lidar_test_patches.shape)
from tensorflow.python.keras.models import Sequential, Model
from tensorflow.python.keras.layers import Conv2D, Dense, Flatten
from tensorflow.python.keras.layers import InputLayer, Input
from tensorflow.python.keras.layers import MaxPooling2D
from tensorflow.python.keras.layers import BatchNormalization, Dropout
from tensorflow.python.keras.optimizers import Adam,SGD
BANDS = area_hsi_train_patches.shape[3]
NUM_CLS = train_one_hot.shape[1]
hsi_input = Input(shape=(HSI_PATCH_SIZE, HSI_PATCH_SIZE, BANDS))
hsi_conv1 = Conv2D(kernel_size=3, strides=1, filters=CONV1, padding='same', activation='relu', name='hsi_conv1')(hsi_input)
hsi_batch1 = BatchNormalization()(hsi_conv1)
hsi_max_pool1 = MaxPooling2D(pool_size=2, strides=2)(hsi_batch1)
hsi_conv2 = Conv2D(kernel_size=5, strides=1, filters=CONV2, padding='same', activation='relu', name='hsi_conv2')(hsi_max_pool1)
hsi_batch2 = BatchNormalization()(hsi_conv2)
hsi_max_pool2 = MaxPooling2D(pool_size=2, strides=2)(hsi_batch2)
hsi_flatten = Flatten()(hsi_max_pool2)
from tensorflow.python.keras.models import Sequential,Model
from tensorflow.python.keras.layers import Conv2D, Dense, Flatten
from tensorflow.python.keras.layers import InputLayer, Input, concatenate
from tensorflow.python.keras.layers import MaxPooling2D
from tensorflow.python.keras.layers import BatchNormalization, Dropout
from tensorflow.python.keras.optimizers import Adam,SGD
BANDS = area_lidar_train_patches.shape[3]
NUM_CLS = train_one_hot.shape[1]
lidar_input = Input(shape=(LiDAR_PATCH_SIZE, LiDAR_PATCH_SIZE, BANDS))
lidar_conv1 = Conv2D(kernel_size=3, strides=1, filters=CONV1, padding='same', activation='relu', name='lidar_conv1')(lidar_input)
lidar_batch1 = BatchNormalization()(lidar_conv1)
lidar_max_pool1 = MaxPooling2D(pool_size=2, strides=2)(lidar_batch1)
lidar_conv2 = Conv2D(kernel_size=3, strides=1, filters=CONV2, padding='same', activation='relu', name='lidar_conv2')(lidar_max_pool1)
lidar_batch2 = BatchNormalization()(lidar_conv2)
lidar_max_pool2 = MaxPooling2D(pool_size=2, strides=2)(lidar_batch2)
lidar_flatten = Flatten()(lidar_max_pool2)
fusion_input = concatenate([hsi_flatten, lidar_flatten])
fusion_fc1 = Dense(FC1, activation='relu')(fusion_input)
fusion_drop1 = Dropout(0.5)(fusion_fc1)
fusion_fc2 = Dense(FC1, activation='relu')(fusion_drop1)
fusion_drop2 = Dropout(0.1)(fusion_fc2)
out = Dense(NUM_CLS, activation='softmax')(fusion_drop2)
model = Model(inputs=[hsi_input, lidar_input], outputs=[out])
sgd = SGD(lr=LEARNING_RATE, decay=1e-6, momentum=0.9, nesterov=True)
model.compile(loss='categorical_crossentropy', optimizer=sgd, metrics=['accuracy'])
model.summary()
history = model.fit([area_hsi_train_patches,area_lidar_train_patches], train_one_hot, batch_size=BATCH_SIZE, shuffle=True, epochs=300)
from operator import truediv
def AA_andEachClassAccuracy(confusion_matrix):
counter = confusion_matrix.shape[0]
list_diag = np.diag(confusion_matrix)
list_raw_sum = np.sum(confusion_matrix, axis=1)
each_acc = np.nan_to_num(truediv(list_diag, list_raw_sum))
average_acc = np.mean(each_acc)
return each_acc, average_acc
test_cls = test_labels - 1
prediction = model.predict([area_hsi_test_patches,area_lidar_test_patches]).argmax(axis=-1)
from sklearn import metrics, preprocessing
overall_acc = metrics.accuracy_score(prediction, test_cls)
kappa = metrics.cohen_kappa_score(prediction, test_cls)
confusion_matrix = metrics.confusion_matrix(prediction, test_cls)
each_acc, average_acc = AA_andEachClassAccuracy(confusion_matrix)
print("Overall Accuracy of training sapmles : ",overall_acc)
print("Average Accuracy of training samples : ",average_acc)
print("Kappa statistics of training samples : ",kappa)
print("Each class accuracy of training samples : ", each_acc)
print("Confusion matrix :", confusion_matrix)
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.