text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
#### Tissue-specific RV eGenes
```
library(data.table)
library(dplyr)
load.data <- function(tissue) {
filename <- paste("/u/project/eeskin2/k8688933/rare_var/results/tss_20k_v8/result_summary/qvals/", tissue, ".lrt.q", sep="")
return(fread(filename, data.table=F))
}
get.egenes <- function(qvals) {
egenes = qvals$Gene_ID[apply(qvals, 1, function(x) {any(as.numeric(x[-1]) < 0.05)})]
return(egenes)
}
get.tissue.specific.genes <- function(egenes.list) {
res = vector("list", length(egenes.list))
names(res) = names(egenes.list)
for (i in 1:length(egenes.list)) {
res[[i]] = egenes.list[[i]][!egenes.list[[i]] %in% unique(unlist(egenes.list[-i]))]
}
return(res)
}
sample.info = fread("/u/project/eeskin2/k8688933/rare_var/results/tss_20k_v8/result_summary/tissue.name.match.csv")
tissues = sample.info$tissue
q.data = lapply(tissues, load.data)
names(q.data) = tissues
egenes = lapply(q.data, get.egenes)
res = get.tissue.specific.genes(egenes)
fwrite(as.list(res$Lung), "../tissue_specific_egenes_by_tissue/Lung.tissue.specifc.rv.egenes.tsv", sep="\n")
fwrite(as.list(res$Liver), "../tissue_specific_egenes_by_tissue/Liver.tissue.specifc.rv.egenes.tsv", sep="\n")
fwrite(as.list(res$Whole_Blood), "../tissue_specific_egenes_by_tissue/Whole_Blood.tissue.specifc.rv.egenes.tsv", sep="\n")
fwrite(as.list(res$Skin_Sun_Exposed_Lower_leg), "../tissue_specific_egenes_by_tissue/Skin_Sun_Exposed_Lower_leg.tissue.specifc.rv.egenes.tsv", sep="\n")
fwrite(as.list(res$Skin_Not_Sun_Exposed_Suprapubic), "../tissue_specific_egenes_by_tissue/Skin_Not_Sun_Exposed_Suprapubic.tissue.specifc.rv.egenes.tsv", sep="\n")
fwrite(as.list(res$Heart_Atrial_Appendage), "../tissue_specific_egenes_by_tissue/Heart_Atrial_Appendage.tissue.specifc.rv.egenes.tsv", sep="\n")
fwrite(as.list(res$Heart_Left_Ventricle), "../tissue_specific_egenes_by_tissue/Heart_Left_Ventricle.tissue.specifc.rv.egenes.tsv", sep="\n")
```
#### Tissue-specific non-RV eGenes
```
get.non.egenes <- function(qvals) {
egenes = qvals$Gene_ID[apply(qvals, 1, function(x) {all(as.numeric(x[-1]) >= 0.05)})]
return(egenes)
}
non.egenes = lapply(q.data, get.non.egenes)
res = get.tissue.specific.genes(non.egenes)
fwrite(as.list(res$Lung), "../tissue_specific_egenes_by_tissue/Lung.tissue.specifc.non.rv.egenes.tsv", sep="\n")
fwrite(as.list(res$Liver), "../tissue_specific_egenes_by_tissue/Liver.tissue.specifc.non.rv.egenes.tsv", sep="\n")
fwrite(as.list(res$Whole_Blood), "../tissue_specific_egenes_by_tissue/Whole_Blood.tissue.specifc.non.rv.egenes.tsv", sep="\n")
fwrite(as.list(res$Skin_Sun_Exposed_Lower_leg), "../tissue_specific_egenes_by_tissue/Skin_Sun_Exposed_Lower_leg.tissue.specifc.non.rv.egenes.tsv", sep="\n")
fwrite(as.list(res$Skin_Not_Sun_Exposed_Suprapubic), "../tissue_specific_egenes_by_tissue/Skin_Not_Sun_Exposed_Suprapubic.tissue.specifc.non.rv.egenes.tsv", sep="\n")
fwrite(as.list(res$Heart_Atrial_Appendage), "../tissue_specific_egenes_by_tissue/Heart_Atrial_Appendage.tissue.specifc.non.rv.egenes.tsv", sep="\n")
fwrite(as.list(res$Heart_Left_Ventricle), "../tissue_specific_egenes_by_tissue/Heart_Left_Ventricle.tissue.specifc.non.rv.egenes.tsv", sep="\n")
length(non.egenes$Lung)
```
#### RV eGenes example outlier
```
library(data.table)
library(dplyr)
target.snp = "chr20_57598808_G_A_b38"
geno = fread("/u/project/eeskin2/k8688933/rare_var/genotypes/v8/all_eur_samples_matrix_maf0.05/chr.20.genotypes.matrix.tsv")
indiv = colnames(geno)[which(geno %>% filter(ID == target.snp) != 0)][-1]
print(indiv)
z.heart.lv = fread("/u/project/eeskin2/k8688933/rare_var/results/tss_20k_v7/sungoohw/result_summary/log2.standardized.corrected.tpm.egenes.only/log2.standardized.corrected.lrt.tpm.Heart_Left_Ventricle")
z.heart.aa = fread("/u/project/eeskin2/k8688933/rare_var/results/tss_20k_v7/sungoohw/result_summary/log2.standardized.corrected.tpm.egenes.only/log2.standardized.corrected.lrt.tpm.Heart_Atrial_Appendage")
z.skin.sun = fread("/u/project/eeskin2/k8688933/rare_var/results/tss_20k_v7/sungoohw/result_summary/log2.standardized.corrected.tpm.egenes.only/log2.standardized.corrected.lrt.tpm.Skin_Sun_Exposed_Lower_leg")
z.skin.not.sun = fread("/u/project/eeskin2/k8688933/rare_var/results/tss_20k_v7/sungoohw/result_summary/log2.standardized.corrected.tpm.egenes.only/log2.standardized.corrected.lrt.tpm.Skin_Not_Sun_Exposed_Suprapubic")
print(indiv %in% colnames(z.heart.lv)) # this SNP is not in heart left ventricle
print(indiv %in% colnames(z.heart.aa))
print(indiv %in% colnames(z.skin.not.sun))
print(indiv %in% colnames(z.skin.sun))
print("ENSG00000101162" %in% z.heart.lv$gene)
print("ENSG00000101162" %in% z.heart.aa$gene)
print("ENSG00000101162" %in% z.skin.not.sun$gene)
print("ENSG00000101162" %in% z.skin.sun$gene)
z.heart.lv %>% filter(gene == "ENSG00000101162") %>% select(indiv)
z.heart.aa %>% filter(gene == "ENSG00000101162") %>% select(indiv)
idx = which(z.skin.sun$gene == "ENSG00000101162")
z.skin.sun[idx, -1]
scaled.z.skin.sun = scale(t(as.data.frame(z.skin.sun)[idx, -1]))
colnames(scaled.z.skin.sun) = c("z")
as.data.frame(scaled.z.skin.sun)[indiv, ] #%>% filter(abs(z) > 2)
idx = which(z.skin.sun$gene == "ENSG00000101162")
colnames(z.skin.sun)[which(abs(z.skin.sun[idx, -1]) > 2)]
z.skin.not.sun %>% filter(gene == "ENSG00000101162") %>% select(indiv)
```
#### RV eGenes example outliers in all tissues
```
z.scores = lapply(dir("/u/project/eeskin2/k8688933/rare_var/results/tss_20k_v7/sungoohw/result_summary/log2.standardized.corrected.tpm.egenes.only/",
pattern="log2.standardized.corrected.lrt.tpm", full.names=T), function(x) {if(file.size(x) > 1) {fread(x, data.table=F)}})
names(z.scores) = fread("../egene.counts.csv")$tissue
z.scores[[17]]
for (i in 1:48) {
z = z.scores[[i]]
if (is.null(z)) {
next
}
if (!indiv %in% colnames(z)) {
next
}
if (!"ENSG00000101162" %in% z$gene) {
next
}
idx = which(z$gene == "ENSG00000101162")
scaled.z = scale(t(as.data.frame(z)[idx, -1]))
colnames(scaled.z) = c("z")
print(names(z.scores)[[i]])
print(as.data.frame(scaled.z)[indiv, ])
}
```
| github_jupyter |
# Aprendizado de máquina - Parte 1
_Aprendizado de máquina_ (_machine learning_, ML) é um subcampo da inteligência artificial que tem por objetivo permitir que o computador _aprenda com os dados_ sem ser explicitamente programado. Em linhas gerais, no _machine learning_ se constrói algoritmos que leem dados, aprendem com a "experiência" deles e inferem coisas a partir do conhecimento adquirido. Esta área tem sido de grande valor para muitos setores por ser capaz de transformar dados aparentemente desconexos em informações cruciais para a tomada de decisões pelo reconhecimento de padrões significativos.
## Modelagem e a subdivisão da área
Os problemas fundamentais de ML em geral podem ser explicados por meio de _modelos_. Um modelo matemático (ou probabilístico) nada mais é do que uma relação entre variáveis. As duas maiores classes de problemas de ML são as seguintes.
- **Aprendizagem supervisionada (_supervised learning_)**, aplicável a situações em que desejamos predizer valores. Neste caso, os algoritmos aprendem a partir de um conjunto de treinamento rotulado (_labels_ ou _exemplars_) e procuram _generalizações_ para todos os dados de entrada possíveis. Em problemas supervisionados, é necessário saber que dado fornece a "verdade fundamental" para que outros possam a ele ser comparados. Popularmente, este termo é chamado de _ground-truth_. Exemplos de algoritmos desta classe são: regressão logística (_logistic regression_), máquinas de vetor de suporte (_support vector machines_) e floresta aleatória (_random forest_).
- **Aprendizagem não-supervisionada (_unsupervised learning_)**, aplicável a situações em que desejamos explorar os dados para explicá-los. Neste caso, os algoritmos aprendem a partir de um conjunto de treinamento não rotulado (_unlabeled) e buscam _explicações_ a partir de algum critério estatístico, geométrico ou de similaridade. Exemplos de algoritmos desta classe são: clusterização por _k-means_ (_k-means clustering_ e núcleo-estimador da função densidade (_kernel density estimation_).
Existe ainda uma terceira classe que não estudaremos neste curso, a qual corresponde à **aprendizagem por reforço** (_reinforcement learning_), cujos algoritmos aprendem a partir de reforço para aperfeiçoar a qualidade de uma resposta explorando o espaço de solução iterativamente.
Como a {numref}`overview-ml` resume, problemas de aprendizagem supervisionada podem ser de:
- _classificação_, se a resposta procurada é discreta, isto é, se há apenas alguns valores possíveis para atribuição (p.ex. classificar se uma família é de baixa, média ou alta renda a partir de dados econômicos);
- _regressão_, se a resposta procurada é contínua, isto é, se admite valores variáveis (p.ex. determinar a renda dos membros de uma família com base em suas profissões).
Por outro lado, problemas de aprendizagem não supervisionada podem ser de:
- _clusterização_, se a resposta procurada deve ser organizada em vários grupos. A clusterização tem similaridades com o problema de classificação, exceto pelo desconhecimento _a priori_, de quantas classes existem;
- _estimativa de densidade_, se a resposta procurada é a explicação de processos fundamentais responsáveis pela distribuição dos dados.
```{figure} ../figs/13/visao-geral-ml.png
---
width: 600px
name: overview-ml
---
Classes principais e problemas fundamentais do _machine learning_. Fonte: adaptado de Chah.
```
## Estudo de caso: classificação de empréstimos bancários
O problema que estudaremos consiste em predizer se o pedido de empréstimo de uma pessoa será parcial ou totalmente aprovado por uma financeira. O banco de dados disponível da financeira abrange os anos de 2007 a 2011.
A aprovação do pedido baseia-se em uma análise de risco que usa diversas informações, tais como renda anual da pessoa, endividamento, calotes, taxa de juros do empréstimo, etc.
Matematicamente, o pedido da pessoa será bem-sucedido se
$$\alpha = \frac{E - F}{E} \ge 0.95,$$
onde $E$ é o valor do empréstimo requisitado e $F$ o financiamento liberado. O classificador binário pode ser escrito pela função
$$h({\bf X}): \mathbb{M}_{n \, \times \, d} \to \mathbb{K},$$
com $\mathbb{K} = \{+1,-1\}$ e ${\bf X}$ é uma matriz de $n$ amostras e $d$ _features_ pertencente ao conjunto abstrato $\mathbb{M}_{n \, \times \, d}$.
```{note}
Em um problema de classificação, se a resposta admite apenas dois valores (duas classes), como "sim" e "não", diz-se que o classificador é **binário**. Se mais valores são admissíveis, diz-se que o classificador é **mutliclasse**.
```
```
import pickle
import numpy as np
import matplotlib.pyplot as plt
```
Vamos ler o banco de dados.
```
import pickle
f = open('../database/dataset_small.pkl','rb')
# necessário encoding 'latin1'
(x,y) = pickle.load(f,encoding='latin1')
```
Aqui, `x` é a nossa matriz de features.
```
# 4140 amostras
# 15 features
x.shape
```
`y` é o vetor de _labels_
```
# 4140 targets +1 ou -1
y,y.shape
```
Comentários:
- As _features_ (atributos) são características que nos permitem distinguir um item. Neste exemplo, são todas as informações coletadas sobre a pessoa ou sobre o mecanismo de empréstimo. São 15, no total, com 4140 valores reais (amostras) cada.
- Em geral, uma amostra pode ser um documento, figura, arquivo de áudio, linha de uma planilha.
- _Features_ são geralmente valores reais, mas podem ser booleanos, discretos, ou categóricos.
- O vetor-alvo (_target_) contém valores que marcam se empréstimos passados no histórico da financeira foram aprovados ou reprovados.
### Interfaces do `scikit-learn`
Usaremos o módulo `scikit-learn` para resolver o problema. Este módulo usa três interfaces:
- `fit()` (estimador), para construir modelos de ajuste;
- `predict()` (preditor), para fazer predições;
- `transform()` (transformador), para converter dados;
O objetivo é predizer empréstimos malsucedidos, isto é, aqueles que se acham aquém do limiar de 95% de $\alpha$.
```
from sklearn import neighbors
# cria uma instância de classificação
# 11 vizinhos mais próximos
nn = 11
knn = neighbors.KNeighborsClassifier(n_neighbors=nn)
# treina o classificador
knn.fit(x,y)
# calcula a predição
yh = knn.predict(x)
# predição, real
y,yh
# altere nn e verifique diferenças
#from numpy import size, where
#size(where(y - yh == 0))
```
```{note}
O algoritmo de classificação dos _K_ vizinhos mais próximos foi proposto em 1975. A base de seu funcionamento é a determinação do rótulo de classificação de uma amostra a partir de _K_ amostras vizinhas em um conjunto de treinamento. Saiba mais [aqui](http://computacaointeligente.com.br/algoritmos/k-vizinhos-mais-proximos/).
```
#### Acurácia
Podemos medir o desempenho do classificador usando métricas. A métrica padrão para o método _KNN_ é a _acurácia_, dada por:
$$acc = 1 - erro = \frac{\text{no. de predições corretas}}{n}.$$
```
knn.score(x,y)
```
Este _score_ parece bom, mas há o que analisar... Vamos plotar a distribuição dos rótulos.
```
# gráfico "torta" (pie chart)
plt.pie(np.c_[np.sum(np.where(y == 1,1,0)),
np.sum(np.where(y == -1,1,0))][0],
labels=['E parcial','E total'],colors=['r','g'],
shadow=False,autopct='%.2f')
plt.gcf().set_size_inches((6,6))
```
O gráfico mostra que o banco de dados está desequilibrado, já que 81,57% dos empréstimos foram liberados integralmente. Isso pode implicar que a predição será pela "maioria".
#### Matriz de confusão
Há casos em que a acurácia não é uma boa métrica de desempenho. Quando análises mais detalhadas são necessárias, podemos usar a _matriz de confusão_.
Com a matriz de confusão, podemos definir métricas para cenários distintos que levam em conta os valores obtidos pelo classificador e os valores considerados como corretos (_ground-truth_, isto é, o "padrão-ouro" (_gold standard_).
Em um classificador binário, há quatro casos a considerar, ilustrados na {numref}`matriz-confusao`:
- _Verdadeiro positivo_ (VP). O classificador prediz uma amostra como positiva que, de fato, é positiva.
- _Falso positivo_ (FP). O classificador prediz uma amostra como positiva que, na verdade, é negativa.
- _Verdadeiro negativo_ (VN). O classificador prediz uma amostra como negativa que, de fato, é negativa.
- _Falso negativo_ (FN). O classificador prediz uma amostra como negativa que, na verdade, é positiva.
```{figure} ../figs/13/matriz-confusao.png
---
width: 600px
name: matriz-confusao
---
Matriz de confusão. Fonte: elaboração própria.
```
Combinando esses quatro conceitos, podemos definir as métricas _acurácia_, _recall_ (ou _sensibilidade_), _especificidade_, _precisão_ (ou _valor previsto positivo_), _valor previsto negativo_, nesta ordem, da seguinte maneira:
$$\text{acc} = \dfrac{TP + TN}{TP + TN + FP + FN}$$
$$\text{rec} = \dfrac{TP}{TP + FP}$$
$$\text{spec} = \dfrac{TN}{TN + FP}$$
$$\text{prec} = \dfrac{TP}{TP + FP}$$
$$\text{npv} = \dfrac{TN}{TN + FN}$$
```{note}
Para uma interpretação ilustrada sobre essas métricas, veja este [post](https://medium.com/swlh/explaining-accuracy-precision-recall-and-f1-score-f29d370caaa8).
```
Podemos computar a matriz de confusão com
```
conf = lambda a,b: np.sum(np.logical_and(yh == a, y == b))
TP, TN, FP, FN = conf(-1,-1), conf(1,1), conf(-1,1), conf(1,-1)
np.array([[TP,FP],[FN,TN]])
```
ou, usando o `scikit-learn`, com
```
from sklearn import metrics
metrics.confusion_matrix(yh,y) # switch (prediction, target)
```
#### Conjuntos de treinamento e de teste
Vejamos um exemplo com `nn=1`.
```
knn = neighbors.KNeighborsClassifier(n_neighbors=1)
knn.fit(x,y)
yh = knn.predict(x)
metrics.accuracy_score(yh,y), metrics.confusion_matrix(yh,y)
```
Este caso tem 100% de acurácia e uma matriz de confusão diagonal. No exemplo anterior, não diferenciamos o conjunto usado para treinamento e predição.
Porém, em problemas reais, as chances dessa perfeição ocorrer são minimas. Da mesma forma, o classificador em geral será aplicado em dados previamente desconhecidos. Esta condição força-nos a dividir os dados em dois conjuntos: aquele usado para aprendizagem (_conjunto de treinamento_) e outro para testar a acurácia (_conjunto de teste_.
Vejamos uma simulação mais realista.
```
# Randomiza e divide dados
# PRC*100% para treinamento
# (1-PRC)*100% para teste
PRC = 0.7
perm = np.random.permutation(y.size)
split_point = int(np.ceil(y.shape[0]*PRC))
X_train = x[perm[:split_point].ravel(),:]
y_train = y[perm[:split_point].ravel()]
X_test = x[perm[split_point:].ravel(),:]
y_test = y[perm[split_point:].ravel()]
aux = {'training': X_train,
'training target':y_train,
'test':X_test,
'test target':y_test}
for k,v in aux.items():
print(k,v.shape,sep=': ')
```
Agora treinaremos o modelo com esta nova partição.
```
knn = neighbors.KNeighborsClassifier(n_neighbors = 1)
knn.fit(X_train, y_train)
yht = knn.predict(X_train)
for k,v in {'acc': str(metrics.accuracy_score(yht, y_train)),
'conf. matrix': '\n' + str(metrics.confusion_matrix(y_train, yht))}.items():
print(k,v,sep=': ')
```
Para `nn = 1`, a acurácia é de 100%. Vejamos o que acontecerá nesta simulação com dados ainda não vistos.
```
yht2 = knn.predict(X_test)
for k,v in {'acc': str(metrics.accuracy_score(yht2, y_test)),
'conf. matrix': '\n' + str(metrics.confusion_matrix(yht2, y_test))}.items():
print(k,v,sep=': ')
```
Neste caso, a acurácia naturalmente reduziu.
| github_jupyter |
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
import tensorflow as tf
print(tf.__version__)
import numpy as np
import matplotlib.pyplot as plt
def plot_series(time, series, format="-", start=0, end=None):
plt.plot(time[start:end], series[start:end], format)
plt.xlabel("Time")
plt.ylabel("Value")
plt.grid(True)
!wget --no-check-certificate \
https://storage.googleapis.com/laurencemoroney-blog.appspot.com/Sunspots.csv \
-O /tmp/sunspots.csv
import csv
time_step = []
sunspots = []
with open('/tmp/sunspots.csv') as csvfile:
reader = csv.reader(csvfile, delimiter=',')
next(reader)
for row in reader:
sunspots.append(float(row[2]))
time_step.append(int(row[0]))
series = np.array(sunspots)
time = np.array(time_step)
plt.figure(figsize=(10, 6))
plot_series(time, series)
split_time = 3000
time_train = time[:split_time]
x_train = series[:split_time]
time_valid = time[split_time:]
x_valid = series[split_time:]
window_size = 60
batch_size = 32
shuffle_buffer_size = 1000
def windowed_dataset(series, window_size, batch_size, shuffle_buffer):
dataset = tf.data.Dataset.from_tensor_slices(series)
dataset = dataset.window(window_size + 1, shift=1, drop_remainder=True)
dataset = dataset.flat_map(lambda window: window.batch(window_size + 1))
dataset = dataset.shuffle(shuffle_buffer).map(lambda window: (window[:-1], window[-1]))
dataset = dataset.batch(batch_size).prefetch(1)
return dataset
dataset = windowed_dataset(x_train, window_size, batch_size, shuffle_buffer_size)
model = tf.keras.models.Sequential([
tf.keras.layers.Dense(20, input_shape=[window_size], activation="relu"),
tf.keras.layers.Dense(10, activation="relu"),
tf.keras.layers.Dense(1)
])
model.compile(loss="mse", optimizer=tf.keras.optimizers.SGD(lr=1e-7, momentum=0.9))
model.fit(dataset,epochs=100,verbose=0)
forecast=[]
for time in range(len(series) - window_size):
forecast.append(model.predict(series[time:time + window_size][np.newaxis]))
forecast = forecast[split_time-window_size:]
results = np.array(forecast)[:, 0, 0]
plt.figure(figsize=(10, 6))
plot_series(time_valid, x_valid)
plot_series(time_valid, results)
tf.keras.metrics.mean_absolute_error(x_valid, results).numpy()
```
| github_jupyter |
# Classification Problem : Credit Card Offer
### Importing the librairies
```
from imblearn.over_sampling import SMOTE
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import pickle
import seaborn as sns
from sklearn.preprocessing import Normalizer
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import confusion_matrix
from sklearn.neighbors import KNeighborsClassifier
from sklearn.metrics import accuracy_score
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from scipy.stats import chi2_contingency
from scipy import stats
import warnings
warnings.filterwarnings('ignore')
```
## 1. Cleaning the CSV file to be uploaded on MySQLworkbench
```
df = pd.read_csv('creditcardmarketing.csv')
df.head()
```
**We will clean the headers only.**
```
def renaming(df):
#removing special characters & following the snake case
df.columns = df.columns.str.replace(' ', '_').str.lower()
df.columns = df.columns.str.replace('#_','')
return df.info()
renaming(df)
```
There are 24 NAN values in the last 5 columns. The 24 rows with the non values will not be uploaded on MySQLWorkbench.
```
# saving the dataframe into CVS file for SQL
df.to_csv('credit_card_data.csv')
```
## 2. Cleaning and EDA
```
# customer_number are unique values so we pass this variable as the index
df = df.set_index('customer_number')
df = df.drop_duplicates()
df.info()
```
### 2.1 - Starting with categorical data
```
def categorical_information (df):
for col in df.select_dtypes('object'):
print (df[col].nunique(), '\n')
print(df[col].value_counts(), '\n')
# get information on categorical data
categorical_information(df)
```
Create visuals for all categorical colums. We can see that the target variable 'offer_accepted' is highly imbalanced.
```
def count_plot_cat(df):
for col in df.select_dtypes('object'):
sns.countplot(df[col])
plt.show()
count_plot_cat(df)
```
### 2.2 Then numerical variables
**Dealing with NAN values**
```
df.isna().sum()
# checking the rows that are null
df[df['q1_balance'].isna()==True]
# since there are only 24 rows with NAN values in the whole dataframe i.e 0,13% of the data, I will drop them.
df = df.dropna()
df = df.reset_index(drop=True)
```
**Exploring data**
```
df.describe()
```
**Create a function to plot graphs of the continuous and discrete variables**
```
def numerical_plotting(df):
decimaux = df.select_dtypes('float64')
entiers = df.select_dtypes('int64')
for col in decimaux:
sns.distplot(decimaux[col])
plt.show()
for col in entiers:
sns.countplot(entiers[col])
plt.show()
numerical_plotting(df)
```
Correlation analysis
Covariance visualization :
we are only working on continuous variables since the discrete variables are featured as categorical variables.
```
#checking distribution of variables
def distribution_distplot(df):
for col in df.select_dtypes('float64'):
sns.distplot(df[col])
# save the figure
# plt.savefig('covariance_account_balance.png', dpi=100, bbox_inches='tight')
plt.show()
distribution_distplot(df)
def corr_matrix(df):
corr_matrix=df.corr(method='pearson')
fig, ax = plt.subplots(figsize=(10, 8))
ax = sns.heatmap(corr_matrix, annot=True)
plt.show()
corr_matrix(df)
```
**Checking on outliers**
```
def vizualizing_outliers(df):
for col in df._get_numeric_data():
sns.boxplot(df[col])
plt.show()
vizualizing_outliers(df)
```
**The outliers should not have relevant impact on our analysis. we will not remove them.**
## 2.3 - Exploration of the target variable : "accepted_offer"
We have already seen that the target variable is higly imbalanced. We will need to deal with this issue.
First, we will explore colinearity between variables.
```
# average_balance vs offer_accepted
plt.figure(figsize=(6,4))
sns.barplot(data=df, y="average_balance", x="offer_accepted")
plt.show()
# Did the customers with the highest average balance accept the offer?
df.nlargest(20,columns="average_balance")[["average_balance","offer_accepted","mailer_type","income_level","own_your_home", "credit_rating","bank_accounts_open","credit_cards_held"]]
# Same question but with the customers with the lowest average balance?
df.nsmallest(20,columns="average_balance")[["average_balance","offer_accepted","mailer_type","income_level","own_your_home", "credit_rating","bank_accounts_open","credit_cards_held"]]
def count_plot_hue_target(df,columns=[], target = ''):
for col in columns:
plt.figure(figsize=(6,4))
sns.countplot(x = col, hue = target, data = df)
plt.show()
count_plot_hue_target(df, columns= df.select_dtypes('object'), target ='offer_accepted')
```
**Checking correlation between categorical variables**
The p_value is used for hypothesis testing and it can be used to measure dependency between two variables.
The null hypothesis to be rejected is that there is no correlation between two variables.
Our anlysis is based on the threshold of 0.05 for p_value. This threshold is related to a confidence interval of 95%.
A p_value below 0.05 means that there is a considerable correlation between two variables, and it is likely that one of them can be dropped without decreasing the metrics of the model.
```
# function to perform ChiSquare-test for categoricalvariables
def chi_square_test(df, columns=[]):
"""This function returns 4 results in this order (chi-square statistic, p value, degrees of freedom,
expected frequencies matrix.
"""
for i in columns:
for j in columns:
if i != j:
data_crosstab = pd.crosstab(df[i], df[j], margins = False)
print('ChiSquare test for ',i,'and ',j,': ')
print(chi2_contingency(data_crosstab, correction=False), '\n')
chi_square_test(df, columns= df.select_dtypes(np.object))
```
Based on the p-values obtained, we can say that :
- the columns 'reward', 'mailer_type', 'income_level' and 'credit_rating' are correlated with our target variable : offer_accepted
- There is no correlation between the other variables
Therefore, we can keep all the variables to run our machine learning algorithms.
## 3. Preprocessing and Modeling
We will use different models to compare them and find the one which fits better our data.
Based on the data we have, we have decided that we need to encode the categorical features.
Regarding the numerical features, a boxcox transformation on the numerical columns could help to improve the model.
As an alternative, we will apply normalize the whole dataframe (after the encoding).
Finally, we will try two techniques to solve the imbalance of the target value : SMOTE (equal number of Yes and No and upsampling to weight the numbers of Yes and No.
In order to train the model, we will apply two algorithms : Logistic Regression and KNN Classifier.
```
# create copies for the different runs
df1 = df.copy()
df2 = df.copy()
```
### 3.1 - Preprocessing - using boxcox and SMOTE
1. Boxcox transformation on numerical variables
2. Encoding - get dummies
3. Dealing with imbalanced - SMOTE
4. Modeling
#### Boxcox transformation
We will apply it to help our features to have a more normal distribution.
```
#Boxcox transformation
def boxcox_transform(df):
numeric_cols = df.select_dtypes(np.number).columns
_ci = {column: None for column in numeric_cols}
for column in numeric_cols:
# since i know any columns should take negative numbers, to avoid -inf in df
df[column] = np.where(df[column]<=0, np.NAN, df[column])
df[column] = df[column].fillna(df[column].mean())
transformed_data, ci = stats.boxcox(df[column])
df[column] = transformed_data
_ci[column] = [ci]
return df, _ci
df, _ci = boxcox_transform(df)
df
```
**Checking the distribution of the features after the boxcox transformation**
```
distribution_distplot(df)
```
**the boxcox transformation improves the normal distribution of the features, especially for the q2_balance and q4_balance columns.**
```
#drop the target
X = df.drop('offer_accepted', axis=1)
y = df['offer_accepted']
```
#### Encoding
```
X= pd.get_dummies(X)
X
```
#### Dealing with imbalanced data
```
# SMOTE
# Uses knn to create rows with similar features from the minority classes.
smote = SMOTE()
X_sm, y_sm = smote.fit_resample(X, y)
y_sm.value_counts()
```
#### Modeling
```
X_train, X_test, y_train, y_test = train_test_split(X_sm, y_sm, test_size=0.2, random_state=42)
```
**Logistic Regression**
```
def logistic_regression_model(X_train, X_test, y_train, y_test):
# defining a function to apply the logistic regression model
classification = LogisticRegression(random_state=42, max_iter=10000)
classification.fit(X_train, y_train)
# and to evaluate the model
score = classification.score(X_test, y_test)
print('The accuracy score is: ', score, '\n')
predictions = classification.predict(X_test)
confusion_matrix(y_test, predictions)
cf_matrix = confusion_matrix(y_test, predictions)
group_names = ['True NO', 'False NO',
'False YES', 'True YES']
group_counts = ["{0:0.0f}".format(value) for value in cf_matrix.flatten()]
group_percentages = ["{0:.2%}".format(value) for value in cf_matrix.flatten()/np.sum(cf_matrix)]
labels = [f"{v1}\n{v2}\n{v3}" for v1, v2, v3 in zip(group_names,group_counts,group_percentages)]
labels = np.asarray(labels).reshape(2,2)
sns.heatmap(cf_matrix, annot=labels, fmt='', cmap='Blues')
print (cf_matrix)
logistic_regression_model(X_train, X_test, y_train, y_test)
```
**KNN-Classifier**
```
#choose the best key value
def best_K(X_train, y_train, X_test, y_test, r):
scores = []
for i in r:
model = KNeighborsClassifier(n_neighbors=i)
model.fit(X_train, y_train)
scores.append(model.score(X_test, y_test))
plt.figure(figsize=(10,6))
plt.plot(r,scores,color = 'blue', linestyle='dashed',
marker='*', markerfacecolor='red', markersize=10)
plt.title('accuracy scores vs. K Value')
plt.xlabel('K')
plt.ylabel('Accuracy')
best_K(X_train, y_train, X_test, y_test, r=range(2,10))
def KNN_classifier_model(X_train, y_train, X_test, y_test,n):
# define a function to apply the KNN Classifier Model
knn = KNeighborsClassifier(n_neighbors=n)
knn.fit(X_train, y_train)
# and to evaluate the model
print('Accuracy of K-NN classifier on test set: {:.2f}'
.format(knn.score(X_test, y_test)))
y_pred = knn.predict(X_test)
print(confusion_matrix(y_test, y_pred))
cf_matrix = confusion_matrix(y_test, y_pred)
group_names = ['True NO', 'False NO',
'False YES', 'True YES']
group_counts = ["{0:0.0f}".format(value) for value in cf_matrix.flatten()]
group_percentages = ["{0:.2%}".format(value) for value in cf_matrix.flatten()/np.sum(cf_matrix)]
labels = [f"{v1}\n{v2}\n{v3}" for v1, v2, v3 in zip(group_names,group_counts,group_percentages)]
labels = np.asarray(labels).reshape(2,2)
sns.heatmap(cf_matrix, annot=labels, fmt='', cmap='Blues')
print (cf_matrix)
KNN_classifier_model(X_train,y_train,X_test,y_test,2)
```
The accuracy scores obtained on the first round are high. Also, the models would be able to predict that the offer is accepted and that the offer is rejected.
### 3.2 - Preprocessing using BoxCox and UpSampling
1. Boxcox transformation on numerical variables
2. UpSampling (60-40)
3. Encoding - get dummies
4. Modeling
In this second round, we will only change the method to deal with unbalanced data. With the SMOTE method, we are getting a dataframe with equal number of Yes and No. Here we are using the upsampling method in order to weight the target column.
#### Boxcox transformation
We are transforming only the continuous variables as previously.
```
#Boxcox function on continous variables
df1, _ci = boxcox_transform(df1)
df1
```
#### Upsampling
```
df1.offer_accepted.value_counts()
# Manually
# getting sample with the 60% as the minority class
Yes = df1[df1['offer_accepted'] == 'Yes'].sample(10173, replace=True)
No = df1[df1['offer_accepted'] == 'No'].sample(16955, replace=True)
upsampled1 = pd.concat([Yes,No]).sample(frac=1) # .sample(frac=1) here is just to shuffle the dataframe
upsampled1
upsampled1.offer_accepted.value_counts()
```
#### Encoding
```
X1 = upsampled1.drop('offer_accepted', axis=1)
y1 = upsampled1['offer_accepted']
X1 = pd.get_dummies(upsampled1)
X1
```
#### Modeling
**Logistic Regression**
```
X1_train, X1_test, y1_train, y1_test = train_test_split(X1, y1, test_size=0.2, random_state=42)
logistic_regression_model(X1_train, X1_test, y1_train, y1_test)
```
This model seems to be overfitted. Therefore it should not perform well for future predictions.
**KNN Classifier**
```
KNN_classifier_model(X1_train,y1_train,X1_test,y1_test,5)
```
The accepted offers are predicted almost at 100%. This leads us to think that the model could not be generalized.
### 3.3 - Preprocessing using Upsampling and normalization
1. Numerical columns in a list
2. Dealing with imbalanced - UpSampling
3. Encoding - get dummies
4. train-test split
5. Normalization of the numerical columns
6. Modeling
```
numerical = df2.select_dtypes(np.number)
numerical.columns
num_col = ['bank_accounts_open', 'credit_cards_held', 'homes_owned',
'household_size', 'average_balance','q1_balance', 'q2_balance', 'q3_balance',
'q4_balance']
```
#### Upsampling
```
df2['offer_accepted'].value_counts()
# Manually
# getting sample with the 60% as the minority class
Yes = df2[df2['offer_accepted'] == 'Yes'].sample(10173, replace=True)
No = df2[df2['offer_accepted'] == 'No'].sample(16955, replace=True)
upsampled2 = pd.concat([Yes,No]).sample(frac=1) # .sample(frac=1) here is just to shuffle the dataframe
upsampled2
```
#### Encoding
```
X2 = upsampled2.drop('offer_accepted', axis=1)
y2 = upsampled2['offer_accepted']
X2 = pd.get_dummies(upsampled2)
X2
```
#### Normalizing
We are normalizing the numerical variables that is to say that we will exclude the encoded categorical variables.
Therefore, we will create the X dataframes on which to apply the normalization and then concatenate them to get the final dataframe on which the model will be trained.
```
# train-test split
X2_train, X2_test, y2_train, y2_test = train_test_split(X2, y2, random_state=0)
#normalizing training and testing set only on numerical variables and not the original categorical variables
X_train_n = X2_train.filter(['bank_accounts_open', 'credit_cards_held', 'homes_owned',
'household_size', 'average_balance','q1_balance', 'q2_balance', 'q3_balance',
'q4_balance'], axis = 1)
X_test_n = X2_test.filter(['bank_accounts_open', 'credit_cards_held', 'homes_owned',
'household_size', 'average_balance','q1_balance', 'q2_balance', 'q3_balance',
'q4_balance'], axis = 1)
#normalization
transformer = Normalizer()
transformer.fit(X_train_n)
# saving in a pickle
with open('std_transformer.pickle', 'wb') as file:
pickle.dump(transformer, file)
# loading from a pickle
with open('std_transformer.pickle', 'rb') as file:
loaded_normalizer = pickle.load(file)
X_train_ = loaded_normalizer.transform(X_train_n)
X_test_ = loaded_normalizer.transform(X_test_n)
#Getting the final dataframe with the normalized variables and the encoded variables.
num_train = pd.DataFrame(X_train_, columns = num_col)
num_test = pd.DataFrame(X_test_, columns = num_col)
X2_train.columns
X_train_c = X2_train.filter(['reward_Air Miles', 'reward_Cash Back', 'reward_Points',
'mailer_type_Letter', 'mailer_type_Postcard', 'income_level_High',
'income_level_Low', 'income_level_Medium', 'overdraft_protection_No',
'overdraft_protection_Yes', 'credit_rating_High', 'credit_rating_Low',
'credit_rating_Medium', 'own_your_home_No', 'own_your_home_Yes'], axis = 1)
X_train_final= pd.concat([num_train.reset_index(drop=True), X_train_c.reset_index(drop=True)], axis=1, ignore_index=True)
X_train_final.info() #checking NAN values
X_test_c = X2_test.filter(['reward_Air Miles', 'reward_Cash Back', 'reward_Points',
'mailer_type_Letter', 'mailer_type_Postcard', 'income_level_High',
'income_level_Low', 'income_level_Medium', 'overdraft_protection_No',
'overdraft_protection_Yes', 'credit_rating_High', 'credit_rating_Low',
'credit_rating_Medium', 'own_your_home_No', 'own_your_home_Yes'], axis = 1)
X_test_final= pd.concat([num_test.reset_index(drop=True), X_test_c.reset_index(drop=True)], axis=1, ignore_index=True)
X_test_final.info()
```
#### Modeling
**Logistic Regression**
```
logistic_regression_model(X_train_final, X_test_final, y2_train, y2_test)
```
This model would not be good at predicting if the offer is accepted.
**KNN Classifier**
```
KNN_classifier_model(X_train_final,y2_train,X_test_final,y2_test,2)
```
## Conclusion
**Comparison of accuracy scores**
| Models | Logistic Regression | KNNClassifier |
| --------------------------:|: -----------------------: | ---------------:|
| BoxCox & SMOTE | 0,94 | 0,86 |
| BoxCox & upsampling | 1 | 0,90 |
| Normalization & upsampling | 0,70 | 0,97 |
We have obtained high accuracy scores in most of our runs. However, we should not necessary be confident on how our models work on future data. In fact, the high results may let us think that they could be overfitted.
| github_jupyter |
# Kernel selection
In this notebook we illustrate the selection of a kernel for a gaussian process.
The kernel is there to modelize the similarity between two points in the input space and, as far as gaussian process are concerned, it can make or break the algorithm.
```
from fastai.tabular.all import *
from tabularGP import tabularGP_learner
from tabularGP.kernel import *
```
## Data
Builds a regression problem on a subset of the adult dataset:
```
path = untar_data(URLs.ADULT_SAMPLE)
df = pd.read_csv(path/'adult.csv').sample(1000)
procs = [FillMissing, Normalize, Categorify]
cat_names = ['workclass', 'education', 'marital-status', 'occupation', 'relationship', 'race']
cont_names = ['education-num', 'fnlwgt']
dep_var = 'age'
data = TabularDataLoaders.from_df(df, path, procs=procs, cat_names=cat_names, cont_names=cont_names, y_names=dep_var)
```
## Tabular kernels
By default, tabularGP uses one kernel type for each continuous features (a [gaussian kernel](https://en.wikipedia.org/wiki/Radial_basis_function_kernel)) and one kernel type for each categorial features (an [index kernel](https://gpytorch.readthedocs.io/en/latest/kernels.html#indexkernel)).
Using those kernels we can compute the similarity between the individual coordinates of two points, those similarity are them combined with what we call a tabular kernel.
The simplest kernel is the `WeightedSumKernel` kernel which computes a weighted sum of the feature similarities.
It is equivalent to a `OR` type of relation: if two points have at least one feature that is similar then they will be considered close in the input space (even if all the other features are very dissimilar).
```
learn = tabularGP_learner(data, kernel=WeightedSumKernel)
learn.fit_one_cycle(5, max_lr=1e-3)
```
Then there is the `WeightedProductKernel` kernel which computes a weighted geometric mean (weighted product) of the feature similarities.
It is equivalent to a `AND` type of relation: all features need to be similar to consider two points similar in the input space.
It is a good kernel to use when features are all continuous and similar (i.e. the `x,y` plane for a function).
```
learn = tabularGP_learner(data, kernel=WeightedProductKernel)
learn.fit_one_cycle(5, max_lr=1e-3)
```
The default tabular kernel is a `ProductOfSumsKernel` which modelise a combinaison of the form: $$s = \prod_i{(\sum_j{\beta_j * s_j})^{\alpha_i}}$$
It is equivalent to a `WeightedProductKernel` put on top of a `WeightedSumKernel` kernel.
This kernel is extremely flexible and recommended when you have a mix of continuous and categorial features.
```
learn = tabularGP_learner(data, kernel=ProductOfSumsKernel)
learn.fit_one_cycle(5, max_lr=1e-3)
```
It is important to note that the choice of the tabular kernel can have a drastic impact on your loss and that you should probably always test all available kernels to find the one that is most suited to your particular problem.
Note that it is fairly easy to design your own `TabularKernel`, following the examples in the [kernel.py](https://github.com/nestordemeure/tabularGP/blob/master/tabularGP/kernel.py) file (while the `feature importance` property is useful, it is optionnal), in order to better accomodate the particular structure of your problem.
## Feature kernels
```
from tabularGP.loss_functions import *
from tabularGP import *
```
There are four continuous kernel provided:
- `ExponentialKernel` which is zero differentiable
- `Matern1Kernel` which is once differentiable
- `Matern2Kernel` which is twice differentiable
- `GaussianKernel` (the default) which is infinitely differentiable
The more differentiable a kernel is and the smoother the modelized function will be.
There are two categorial kernel provided:
- `HammingKernel` which consider different elements of a category as have a similarity of zero
- `IndexKernel` (the default) which consider that different elements can still be similar
While the choice of feature kernel tend to be less impactful, you can manually select them if you build your model yourself:
```
model = TabularGPModel(data, kernel=WeightedProductKernel, cont_kernel=ExponentialKernel, cat_kernel=HammingKernel)
loss_func = gp_gaussian_marginal_log_likelihood # would have used `gp_is_greater_log_likelihood` for classification
learn = TabularGPLearner(data, model, loss_func=loss_func)
learn.fit_one_cycle(5, max_lr=1e-3)
```
It is also fairly easy to provide your own feature kernel to modelize behaviour specific to your data (periodicity, trends, etc).
To learn more about the implementation of kernels adapted to a particular problem, we recommend the chapter two (*Expressing Structure with Kernels*) and three (*Automatic Model Construction*) of the very good [Automatic Model Construction with Gaussian Processes](http://www.cs.toronto.edu/~duvenaud/thesis.pdf).
## Transfer learning
Kernels model the input space, as such they can be reused from an output type to another in order to tranfert domain knowledge and speed up training.
Here is a classification problem using the same input features (different features would lead to a crash as the input space would be different):
```
cat_names = ['workclass', 'education', 'marital-status', 'occupation', 'relationship', 'race']
cont_names = ['education-num', 'fnlwgt']
dep_var = 'salary'
data_classification = TabularDataLoaders.from_df(df, path, procs=procs, cat_names=cat_names, cont_names=cont_names, y_names=dep_var, bs=64)
```
We can reuse the kernel from our regression task by passing the learner, model or trained kernel to the `kernel` argument of our builder:
```
learn_classification = tabularGP_learner(data, kernel=learn)
learn_classification.fit_one_cycle(5, max_lr=1e-3)
```
Note that, by default, the kernel is frozen when transfering knowledge. Lets unfreeze it now that the rest of the gaussian process is trained:
```
learn_classification.unfreeze(kernel=True)
learn_classification.fit_one_cycle(5, max_lr=1e-3)
```
| github_jupyter |
```
import os
import tifffile
from sklearn.model_selection import train_test_split
from tqdm import tqdm_notebook as tqdm
from shutil import copyfile
import numpy as np
working_dir = '/media/jswaney/SSD EVO 860/organoid_phenotyping/ventricle_segmentation'
def load_tiff_bioformats(path):
data = tifffile.imread(path)[0, 0] # Bioformats 5D stack
return data[0], data[1]
```
# Normalize and make train and test sets
```
data_dir = 'eF20_B3_2'
files = os.listdir(os.path.join(working_dir, data_dir))
len(files)
x = np.linspace(0, 2**16 - 1, 256)
h = np.zeros(256)
for file in tqdm(files):
img, seg = load_tiff_bioformats(os.path.join(working_dir,
data_dir,
file))
h += np.histogram(img.ravel(), bins=256, range=(x[0], x[-1]))[0]
cdf = np.cumsum(h)
cdf = cdf / cdf.max()
diff = np.abs(cdf - 0.997)
idx = np.where(diff == diff.min())[0]
min_value = 0
max_value = x[idx][0]
print(max_value)
plt.subplot(211)
plt.plot(x, h)
plt.ylim([0, 1e7])
plt.subplot(212)
plt.plot(x, cdf)
plt.plot([x[0], x[-1]], [0.997, 0.997])
plt.show()
files_train, files_test = train_test_split(files,
test_size=0.10,
random_state=123)
len(files_train), len(files_test)
train_dir = 'train'
test_dir = 'test'
class_dir = 'class_0'
os.makedirs(os.path.join(working_dir, train_dir, class_dir),
exist_ok=True)
os.makedirs(os.path.join(working_dir, test_dir, class_dir),
exist_ok=True)
for file in tqdm(files_train):
input_path = os.path.join(working_dir, data_dir, file)
output_path = os.path.join(working_dir, train_dir, class_dir, file)
img, seg = load_tiff_bioformats(os.path.join(working_dir,
data_dir,
file))
img_normalized = (img-min_value)/(max_value-min_value)
img_normalized = np.clip(img_normalized * 255, 0, 255)
data = np.stack([img_normalized.astype(np.uint8),
(seg * 255).astype(np.uint8)], axis=0)
tifffile.imsave(output_path, data, compress=1)
for file in tqdm(files_test):
input_path = os.path.join(working_dir, data_dir, file)
output_path = os.path.join(working_dir, test_dir, class_dir, file)
img, seg = load_tiff_bioformats(os.path.join(working_dir,
data_dir,
file))
img_normalized = np.clip((img-min_value)/(max_value-min_value) * 255, 0, 255)
data = np.stack([img_normalized.astype(np.uint8),
(seg * 255).astype(np.uint8)], axis=0)
tifffile.imsave(output_path, data, compress=1)
```
# Make Dataloaders for segmentations
```
def load_tiff_seg(path):
data = tifffile.imread(path)
data = np.stack([data[0], data[1], np.zeros(data[0].shape, data.dtype)])
return data.transpose((1, 2, 0))
import torch
from torch import nn
from torchvision.datasets import DatasetFolder
from torchvision import transforms
from torch.utils.data import DataLoader
from torch.optim import Adam
import matplotlib.pyplot as plt
import sys
sys.path.append('/home/jswaney/Pytorch-UNet/')
from unet.unet_model import UNet
data = load_tiff_seg(output_path)
data[0].shape, data[0].dtype
print(img.min())
plt.imshow(img)
plt.show()
degrees = 45
scale = (0.8, 1.2)
size = 256
dataset_train = DatasetFolder(os.path.join(working_dir, train_dir),
loader=load_tiff_seg,
extensions=['.tif'],
transform=transforms.Compose([
transforms.ToPILImage(),
transforms.RandomHorizontalFlip(),
transforms.RandomAffine(degrees,
scale=scale),
transforms.RandomCrop(size),
transforms.ToTensor()]))
dataset_test = DatasetFolder(os.path.join(working_dir, test_dir),
loader=load_tiff_seg,
extensions=['.tif'],
transform=transforms.ToTensor())
print(dataset_train)
print(dataset_test)
use_cuda = torch.cuda.is_available()
device = torch.device("cuda" if use_cuda else "cpu")
device
batch_size = 1
n_workers = 1 if use_cuda else 0
pin_memory = True if use_cuda else False
dataloader_train = DataLoader(dataset_train,
batch_size,
shuffle=True,
num_workers=n_workers,
pin_memory=pin_memory)
dataloader_test = DataLoader(dataset_test,
batch_size,
num_workers=n_workers,
pin_memory=pin_memory)
x, _ = next(iter(dataloader_train))
x.shape, x.dtype, x.max()
plt.imshow(x.numpy()[0, 1])
plt.show()
model = UNet(n_channels=1, n_classes=1)
model
model = model.to(device)
optimizer = Adam(model.parameters(), lr=0.001)
model.train()
def train_epoch(model, epoch, dataloader_train, device, optimizer, criterion, log_interval=100):
model.train()
n_batch = len(dataloader_train)
epoch_loss = 0
count = 0
for batch_idx, (x, _) in enumerate(dataloader_train):
img = x[:, 0].unsqueeze(1)
img = img * (1 + np.random.random(1)[0])
img = img.to(device)
seg = x[:, 1].unsqueeze(1)
seg = (seg > 0).type(torch.float).to(device)
if seg.max() > 0:
weight = torch.tensor([10], dtype=torch.float32).to(device)
else:
weight = torch.tensor([1], dtype=torch.float32).to(device)
optimizer.zero_grad()
output = model(img)
output_flat = output.view(-1)
seg_flat = seg.view(-1)
criterion.weight = weight
loss = criterion(output_flat, seg_flat)
loss.backward()
optimizer.step()
count += len(x)
epoch_loss += loss.item()
ave_loss = epoch_loss / count
if batch_idx % log_interval == 0:
print(f'Train Epoch: {epoch} [{batch_idx * len(x)}/{len(dataloader_train.dataset)} ({100*batch_idx/n_batch:.0f}%)]\tLoss: {ave_loss}')
def test_epoch(model, dataloader_test, criterion):
model.eval()
test_loss = 0
correct = 0
with torch.no_grad():
for x, _ in dataloader_test:
img = x[:, 0].unsqueeze(1).to(device)
seg = x[:, 1].unsqueeze(1)
seg = (seg > 0).type(torch.float).to(device)
output = model(img)
output_flat = output.view(-1)
seg_flat = seg.view(-1)
test_loss += criterion(output_flat, seg_flat)
test_loss /= len(dataloader_test.dataset)
print(f'Test set: Total loss: {test_loss:.4f}')
criterion = nn.BCELoss()
n_epochs = 200
for epoch in range(n_epochs):
train_epoch(model, epoch, dataloader_train, device, optimizer, criterion)
test_epoch(model, dataloader_test, criterion)
%matplotlib inline
x, _ = next(iter(dataloader_train))
img = x[:, 0].unsqueeze(1).to(device)
seg = x[:, 1].unsqueeze(1)
seg = (seg > 0).type(torch.float).to(device)
output = model(img)
img = img.detach().cpu().numpy()
seg = seg.detach().cpu().numpy()
output = output.detach().cpu().numpy()
img = img[0, 0]
seg = seg[0, 0]
output = output[0, 0]
plt.figure(figsize=(4, 8))
plt.subplot(311)
plt.imshow(img, clim=[0, 1])
plt.title('Syto 16')
plt.subplot(312)
# plt.imshow(seg, clim=[0, 1])
plt.hist(img.ravel(), bins=256, range=(0, 1))
plt.title('Ground Truth')
plt.subplot(313)
plt.imshow(output, clim=[0, 0.5])
plt.title('Predicted')
plt.show()
# print(seg.max(), img.max())
torch.save(model.state_dict(), os.path.join(working_dir, 'unet_d35_d60_200.pt'))
```
| github_jupyter |
# Fast Bayesian estimation of SARIMAX models
## Introduction
This notebook will show how to use fast Bayesian methods to estimate SARIMAX (Seasonal AutoRegressive Integrated Moving Average with eXogenous regressors) models. These methods can also be parallelized across multiple cores.
Here, fast methods means a version of Hamiltonian Monte Carlo called the No-U-Turn Sampler (NUTS) developed by Hoffmann and Gelman: see [Hoffman, M. D., & Gelman, A. (2014). The No-U-Turn sampler: adaptively setting path lengths in Hamiltonian Monte Carlo. Journal of Machine Learning Research, 15(1), 1593-1623.](https://arxiv.org/abs/1111.4246). As they say, "the cost of HMC per independent sample from a target distribution of dimension $D$ is roughly $\mathcal{O}(D^{5/4})$, which stands in sharp contrast with the $\mathcal{O}(D^{2})$ cost of random-walk Metropolis". So for problems of larger dimension, the time-saving with HMC is significant. However it does require the gradient, or Jacobian, of the model to be provided.
This notebook will combine the Python libraries [statsmodels](https://www.statsmodels.org/stable/index.html), which does econometrics, and [PyMC3](https://docs.pymc.io/), which is for Bayesian estimation, to perform fast Bayesian estimation of a simple SARIMAX model, in this case an ARMA(1, 1) model for US CPI.
Note that, for simple models like AR(p), base PyMC3 is a quicker way to fit a model; there's an [example here](https://docs.pymc.io/notebooks/AR.html). The advantage of using statsmodels is that it gives access to methods that can solve a vast range of statespace models.
The model we'll solve is given by
$$
y_t = \phi y_{t-1} + \varepsilon_t + \theta_1 \varepsilon_{t-1}, \qquad \varepsilon_t \sim N(0, \sigma^2)
$$
with 1 auto-regressive term and 1 moving average term. In statespace form it is written as:
$$
\begin{align}
y_t & = \underbrace{\begin{bmatrix} 1 & \theta_1 \end{bmatrix}}_{Z} \underbrace{\begin{bmatrix} \alpha_{1,t} \\ \alpha_{2,t} \end{bmatrix}}_{\alpha_t} \\
\begin{bmatrix} \alpha_{1,t+1} \\ \alpha_{2,t+1} \end{bmatrix} & = \underbrace{\begin{bmatrix}
\phi & 0 \\
1 & 0 \\
\end{bmatrix}}_{T} \begin{bmatrix} \alpha_{1,t} \\ \alpha_{2,t} \end{bmatrix} +
\underbrace{\begin{bmatrix} 1 \\ 0 \end{bmatrix}}_{R} \underbrace{\varepsilon_{t+1}}_{\eta_t} \\
\end{align}
$$
The code will follow these steps:
1. Import external dependencies
2. Download and plot the data on US CPI
3. Simple maximum likelihood estimation (MLE) as an example
4. Definitions of helper functions to provide tensors to the library doing Bayesian estimation
5. Bayesian estimation via NUTS
6. Application to US CPI series
Finally, Appendix A shows how to re-use the helper functions from step (4) to estimate a different state space model, `UnobservedComponents`, using the same Bayesian methods.
### 1. Import external dependencies
```
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import pymc3 as pm
import statsmodels.api as sm
import theano
import theano.tensor as tt
from pandas.plotting import register_matplotlib_converters
from pandas_datareader.data import DataReader
plt.style.use("seaborn")
register_matplotlib_converters()
```
### 2. Download and plot the data on US CPI
We'll get the data from FRED:
```
cpi = DataReader("CPIAUCNS", "fred", start="1971-01", end="2018-12")
cpi.index = pd.DatetimeIndex(cpi.index, freq="MS")
# Define the inflation series that we'll use in analysis
inf = np.log(cpi).resample("QS").mean().diff()[1:] * 400
inf = inf.dropna()
print(inf.head())
# Plot the series
fig, ax = plt.subplots(figsize=(9, 4), dpi=300)
ax.plot(inf.index, inf, label=r"$\Delta \log CPI$", lw=2)
ax.legend(loc="lower left")
plt.show()
```
### 3. Fit the model with maximum likelihood
Statsmodels does all of the hard work of this for us - creating and fitting the model takes just two lines of code. The model order parameters correspond to auto-regressive, difference, and moving average orders respectively.
```
# Create an SARIMAX model instance - here we use it to estimate
# the parameters via MLE using the `fit` method, but we can
# also re-use it below for the Bayesian estimation
mod = sm.tsa.statespace.SARIMAX(inf, order=(1, 0, 1))
res_mle = mod.fit(disp=False)
print(res_mle.summary())
```
It's a good fit. We can also get the series of one-step ahead predictions and plot it next to the actual data, along with a confidence band.
```
predict_mle = res_mle.get_prediction()
predict_mle_ci = predict_mle.conf_int()
lower = predict_mle_ci["lower CPIAUCNS"]
upper = predict_mle_ci["upper CPIAUCNS"]
# Graph
fig, ax = plt.subplots(figsize=(9, 4), dpi=300)
# Plot data points
inf.plot(ax=ax, style="-", label="Observed")
# Plot predictions
predict_mle.predicted_mean.plot(ax=ax, style="r.", label="One-step-ahead forecast")
ax.fill_between(predict_mle_ci.index, lower, upper, color="r", alpha=0.1)
ax.legend(loc="lower left")
plt.show()
```
### 4. Helper functions to provide tensors to the library doing Bayesian estimation
We're almost on to the magic but there are a few preliminaries. Feel free to skip this section if you're not interested in the technical details.
### Technical Details
PyMC3 is a Bayesian estimation library ("Probabilistic Programming in Python: Bayesian Modeling and Probabilistic Machine Learning with Theano") that is a) fast and b) optimized for Bayesian machine learning, for instance [Bayesian neural networks](https://docs.pymc.io/notebooks/bayesian_neural_network_advi.html). To do all of this, it is built on top of a Theano, a library that aims to evaluate tensors very efficiently and provide symbolic differentiation (necessary for any kind of deep learning). It is the symbolic differentiation that means PyMC3 can use NUTS on any problem formulated within PyMC3.
We are not formulating a problem directly in PyMC3; we're using statsmodels to specify the statespace model and solve it with the Kalman filter. So we need to put the plumbing of statsmodels and PyMC3 together, which means wrapping the statsmodels SARIMAX model object in a Theano-flavored wrapper before passing information to PyMC3 for estimation.
Because of this, we can't use the Theano auto-differentiation directly. Happily, statsmodels SARIMAX objects have a method to return the Jacobian evaluated at the parameter values. We'll be making use of this to provide gradients so that we can use NUTS.
#### Defining helper functions to translate models into a PyMC3 friendly form
First, we'll create the Theano wrappers. They will be in the form of 'Ops', operation objects, that 'perform' particular tasks. They are initialized with a statsmodels `model` instance.
Although this code may look somewhat opaque, it is generic for any state space model in statsmodels.
```
class Loglike(tt.Op):
itypes = [tt.dvector] # expects a vector of parameter values when called
otypes = [tt.dscalar] # outputs a single scalar value (the log likelihood)
def __init__(self, model):
self.model = model
self.score = Score(self.model)
def perform(self, node, inputs, outputs):
(theta,) = inputs # contains the vector of parameters
llf = self.model.loglike(theta)
outputs[0][0] = np.array(llf) # output the log-likelihood
def grad(self, inputs, g):
# the method that calculates the gradients - it actually returns the
# vector-Jacobian product - g[0] is a vector of parameter values
(theta,) = inputs # our parameters
out = [g[0] * self.score(theta)]
return out
class Score(tt.Op):
itypes = [tt.dvector]
otypes = [tt.dvector]
def __init__(self, model):
self.model = model
def perform(self, node, inputs, outputs):
(theta,) = inputs
outputs[0][0] = self.model.score(theta)
```
### 5. Bayesian estimation with NUTS
The next step is to set the parameters for the Bayesian estimation, specify our priors, and run it.
```
# Set sampling params
ndraws = 3000 # number of draws from the distribution
nburn = 600 # number of "burn-in points" (which will be discarded)
```
Now for the fun part! There are three parameters to estimate: $\phi$, $\theta_1$, and $\sigma$. We'll use uninformative uniform priors for the first two, and an inverse gamma for the last one. Then we'll run the inference optionally using as many computer cores as I have.
```
# Construct an instance of the Theano wrapper defined above, which
# will allow PyMC3 to compute the likelihood and Jacobian in a way
# that it can make use of. Here we are using the same model instance
# created earlier for MLE analysis (we could also create a new model
# instance if we preferred)
loglike = Loglike(mod)
with pm.Model() as m:
# Priors
arL1 = pm.Uniform("ar.L1", -0.99, 0.99)
maL1 = pm.Uniform("ma.L1", -0.99, 0.99)
sigma2 = pm.InverseGamma("sigma2", 2, 4)
# convert variables to tensor vectors
theta = tt.as_tensor_variable([arL1, maL1, sigma2])
# use a DensityDist (use a lamdba function to "call" the Op)
pm.DensityDist("likelihood", loglike, observed=theta)
# Draw samples
trace = pm.sample(
ndraws,
tune=nburn,
return_inferencedata=True,
cores=1,
compute_convergence_checks=False,
)
```
Note that the NUTS sampler is auto-assigned because we provided gradients. PyMC3 will use Metropolis or Slicing samplers if it does not find that gradients are available. There are an impressive number of draws per second for a "block box" style computation! However, note that if the model can be represented directly by PyMC3 (like the AR(p) models mentioned above), then computation can be substantially faster.
Inference is complete, but are the results any good? There are a number of ways to check. The first is to look at the posterior distributions (with lines showing the MLE values):
```
plt.tight_layout()
# Note: the syntax here for the lines argument is required for
# PyMC3 versions >= 3.7
# For version <= 3.6 you can use lines=dict(res_mle.params) instead
_ = pm.plot_trace(
trace,
lines=[(k, {}, [v]) for k, v in dict(res_mle.params).items()],
combined=True,
figsize=(12, 12),
)
```
The estimated posteriors clearly peak close to the parameters found by MLE. We can also see a summary of the estimated values:
```
pm.summary(trace)
```
Here $\hat{R}$ is the Gelman-Rubin statistic. It tests for lack of convergence by comparing the variance between multiple chains to the variance within each chain. If convergence has been achieved, the between-chain and within-chain variances should be identical. If $\hat{R}<1.2$ for all model parameters, we can have some confidence that convergence has been reached.
Additionally, the highest posterior density interval (the gap between the two values of HPD in the table) is small for each of the variables.
### 6. Application of Bayesian estimates of parameters
We'll now re-instigate a version of the model but using the parameters from the Bayesian estimation, and again plot the one-step-ahead forecasts.
```
# Retrieve the posterior means
params = pm.summary(trace)["mean"].values
# Construct results using these posterior means as parameter values
res_bayes = mod.smooth(params)
predict_bayes = res_bayes.get_prediction()
predict_bayes_ci = predict_bayes.conf_int()
lower = predict_bayes_ci["lower CPIAUCNS"]
upper = predict_bayes_ci["upper CPIAUCNS"]
# Graph
fig, ax = plt.subplots(figsize=(9, 4), dpi=300)
# Plot data points
inf.plot(ax=ax, style="-", label="Observed")
# Plot predictions
predict_bayes.predicted_mean.plot(ax=ax, style="r.", label="One-step-ahead forecast")
ax.fill_between(predict_bayes_ci.index, lower, upper, color="r", alpha=0.1)
ax.legend(loc="lower left")
plt.show()
```
## Appendix A. Application to `UnobservedComponents` models
We can reuse the `Loglike` and `Score` wrappers defined above to consider a different state space model. For example, we might want to model inflation as the combination of a random walk trend and autoregressive error term:
$$
\begin{aligned}
y_t & = \mu_t + \varepsilon_t \\
\mu_t & = \mu_{t-1} + \eta_t \\
\varepsilon_t &= \phi \varepsilon_t + \zeta_t
\end{aligned}
$$
This model can be constructed in Statsmodels with the `UnobservedComponents` class using the `rwalk` and `autoregressive` specifications. As before, we can fit the model using maximum likelihood via the `fit` method.
```
# Construct the model instance
mod_uc = sm.tsa.UnobservedComponents(inf, "rwalk", autoregressive=1)
# Fit the model via maximum likelihood
res_uc_mle = mod_uc.fit()
print(res_uc_mle.summary())
```
As noted earlier, the Theano wrappers (`Loglike` and `Score`) that we created above are generic, so we can re-use essentially the same code to explore the model with Bayesian methods.
```
# Set sampling params
ndraws = 3000 # number of draws from the distribution
nburn = 600 # number of "burn-in points" (which will be discarded)
# Here we follow the same procedure as above, but now we instantiate the
# Theano wrapper `Loglike` with the UC model instance instead of the
# SARIMAX model instance
loglike_uc = Loglike(mod_uc)
with pm.Model():
# Priors
sigma2level = pm.InverseGamma("sigma2.level", 1, 1)
sigma2ar = pm.InverseGamma("sigma2.ar", 1, 1)
arL1 = pm.Uniform("ar.L1", -0.99, 0.99)
# convert variables to tensor vectors
theta_uc = tt.as_tensor_variable([sigma2level, sigma2ar, arL1])
# use a DensityDist (use a lamdba function to "call" the Op)
pm.DensityDist("likelihood", loglike_uc, observed=theta_uc)
# Draw samples
trace_uc = pm.sample(
ndraws,
tune=nburn,
return_inferencedata=True,
cores=1,
compute_convergence_checks=False,
)
```
And as before we can plot the marginal posteriors. In contrast to the SARIMAX example, here the posterior modes are somewhat different from the MLE estimates.
```
plt.tight_layout()
# Note: the syntax here for the lines argument is required for
# PyMC3 versions >= 3.7
# For version <= 3.6 you can use lines=dict(res_mle.params) instead
_ = pm.plot_trace(
trace_uc,
lines=[(k, {}, [v]) for k, v in dict(res_uc_mle.params).items()],
combined=True,
figsize=(12, 12),
)
pm.summary(trace_uc)
# Retrieve the posterior means
params = pm.summary(trace_uc)["mean"].values
# Construct results using these posterior means as parameter values
res_uc_bayes = mod_uc.smooth(params)
```
One benefit of this model is that it gives us an estimate of the underling "level" of inflation, using the smoothed estimate of $\mu_t$, which we can access as the "level" column in the results objects' `states.smoothed` attribute. In this case, because the Bayesian posterior mean of the level's variance is larger than the MLE estimate, its estimated level is a little more volatile.
```
# Graph
fig, ax = plt.subplots(figsize=(9, 4), dpi=300)
# Plot data points
inf["CPIAUCNS"].plot(ax=ax, style="-", label="Observed data")
# Plot estimate of the level term
res_uc_mle.states.smoothed["level"].plot(ax=ax, label="Smoothed level (MLE)")
res_uc_bayes.states.smoothed["level"].plot(ax=ax, label="Smoothed level (Bayesian)")
ax.legend(loc="lower left");
```
| github_jupyter |
## Set Up
We have again provided code to do the basic loading, review and model-building. Run the cell below to set everything up:
```
import numpy as np
import pandas as pd
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import train_test_split
import shap
# Environment Set-Up for feedback system.
from learntools.core import binder
binder.bind(globals())
from learntools.ml_explainability.ex5 import *
print("Setup Complete")
import pandas as pd
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
data = pd.read_csv('../input/hospital-readmissions/train.csv')
y = data.readmitted
base_features = ['number_inpatient', 'num_medications', 'number_diagnoses', 'num_lab_procedures',
'num_procedures', 'time_in_hospital', 'number_outpatient', 'number_emergency',
'gender_Female', 'payer_code_?', 'medical_specialty_?', 'diag_1_428', 'diag_1_414',
'diabetesMed_Yes', 'A1Cresult_None']
# Some versions of shap package error when mixing bools and numerics
X = data[base_features].astype(float)
train_X, val_X, train_y, val_y = train_test_split(X, y, random_state=1)
# For speed, we will calculate shap values on smaller subset of the validation data
small_val_X = val_X.iloc[:150]
my_model = RandomForestClassifier(n_estimators=30, random_state=1).fit(train_X, train_y)
data.describe()
```
The first few questions require examining the distribution of effects for each feature, rather than just an average effect for each feature. Run the following cell for a summary plot of the shap_values for readmission. It will take about 20 seconds to run.
```
explainer = shap.TreeExplainer(my_model)
shap_values = explainer.shap_values(small_val_X)
shap.summary_plot(shap_values[1], small_val_X)
```
## Question 1
Which of the following features has a bigger range of effects on predictions (i.e. larger difference between most positive and most negative effect)
- `diag_1_428` or
- `payer_code_?`
```
# set following variable to 'diag_1_428' or 'payer_code_?'
feature_with_bigger_range_of_effects = ____
q_1.check()
```
Uncomment the line below to see the solution and explanation
```
# q_1.solution()
```
## Question 2
Do you believe the range of effects sizes (distance between smallest effect and largest effect) is a good indication of which feature will have a higher permutation importance? Why or why not?
If the **range of effect sizes** measures something different from **permutation importance**: which is a better answer for the question "Which of these two features does the model say is more important for us to understand when discussing readmission risks in the population?"
Uncomment the following line after you've decided your answer.
```
# q_2.solution()
```
## Question 3
Both `diag_1_428` and `payer_code_?` are binary variables, taking values of 0 or 1.
From the graph, which do you think would typically have a bigger impact on predicted readmission risk:
- Changing `diag_1_428` from 0 to 1
- Changing `payer_code_?` from 0 to 1
To save you scrolling, we have included a cell below to plot the graph again (this one runs quickly).
```
shap.summary_plot(shap_values[1], small_val_X)
# Set following var to "diag_1_428" if changing it to 1 has bigger effect. Else set it to 'payer_code_?'
bigger_effect_when_changed = ____
q_3.check()
```
For a solution and explanation, uncomment the line below
```
# q_3.solution()
```
## Question 4
Some features (like `number_inpatient`) have reasonably clear separation between the blue and pink dots. Other variables like `num_lab_procedures` have blue and pink dots jumbled together, even though the SHAP values (or impacts on prediction) aren't all 0.
What do you think you learn from the fact that `num_lab_procedures` has blue and pink dots jumbled together? Once you have your answer, uncomment the line below to verify your solution.
```
# q_4.solution()
```
## Question 5
Consider the following SHAP contribution dependence plot.
The x-axis shows `feature_of_interest` and the points are colored based on `other_feature`.

Is there an interaction between `feature_of_interest` and `other_feature`?
If so, does `feature_of_interest` have a more positive impact on predictions when `other_feature` is high or when `other_feature` is low?
Uncomment the following code when you are ready for the answer.
```
# q_5.solution()
```
## Question 6
Review the summary plot for the readmission data by running the following cell:
```
shap.summary_plot(shap_values[1], small_val_X)
```
Both **num_medications** and **num_lab_procedures** share that jumbling of pink and blue dots.
Aside from `num_medications` having effects of greater magnitude (both more positive and more negative), it's hard to see a meaningful difference between how these two features affect readmission risk. Create the SHAP dependence contribution plots for each variable, and describe what you think is different between how these two variables affect predictions.
As a reminder, here is the code you previously saw to create this type of plot.
shap.dependence_plot(feature_of_interest, shap_values[1], val_X)
And recall that your validation data is called `small_val_X`.
```
# Your code here
____
```
Then uncomment the following line to compare your observations from this graph to the solution.
```
# q_6.solution()
```
## Congrats
That's it! Machine Learning models should not feel like black boxes any more, because you have the tools to inspect them and understand what they learn about the world.
This is an excellent skill for debugging models, building trust, and learning insights to make better decisions. These techniques have revolutionized how I do data science, and I hope they do the same for you.
Real data science involves an element of exploration. I hope you find an interesting dataset to try these techniques on (Kaggle has a lot of [free datasets](https://www.kaggle.com/datasets) to try out). If you learn something interesting about the world, share your work [in this forum](https://www.kaggle.com/learn-forum/66354). I'm excited to see what you do with your new skills.
| github_jupyter |
# High-level Caffe2 Example
```
import os
import sys
import caffe2
import numpy as np
from caffe2.python import core, model_helper, workspace, visualize, brew, optimizer, utils
from caffe2.proto import caffe2_pb2
from common.params import *
from common.utils import *
# Force one-gpu
os.environ["CUDA_VISIBLE_DEVICES"] = "0"
print("OS: ", sys.platform)
print("Python: ", sys.version)
print("Numpy: ", np.__version__)
print("GPU: ", get_gpu_name())
print(get_cuda_version())
print("CuDNN Version ", get_cudnn_version())
DEVICE_OPTS = core.DeviceOption(caffe2_pb2.CUDA, 0) # Run on GPU
def create_model(m, device_opts=DEVICE_OPTS, n_classes=N_CLASSES):
with core.DeviceScope(device_opts):
conv1 = brew.conv(m, 'data', 'conv1', dim_in=3, dim_out=50, kernel=3, pad=1, no_gradient_to_input=1)
relu1 = brew.relu(m, conv1, 'relu1')
conv2 = brew.conv(m, relu1, 'conv2', dim_in=50, dim_out=50, kernel=3, pad=1)
pool1 = brew.max_pool(m, conv2, 'pool1', kernel=2, stride=2)
relu2 = brew.relu(m, pool1, 'relu2')
drop1 = brew.dropout(m, relu2, 'drop1', ratio=0.25)
conv3 = brew.conv(m, drop1, 'conv3', dim_in=50, dim_out=100, kernel=3, pad=1)
relu3 = brew.relu(m, conv3, 'relu3')
conv4 = brew.conv(m, relu3, 'conv4', dim_in=100, dim_out=100, kernel=3, pad=1)
pool2 = brew.max_pool(m, conv4, 'pool2', kernel=2, stride=2)
relu4 = brew.relu(m, pool2, 'relu4')
drop2 = brew.dropout(m, relu4, 'drop2', ratio=0.25)
fc1 = brew.fc(m, drop2, 'fc1', dim_in=100 * 8 * 8, dim_out=512)
relu5 = brew.relu(m, fc1, 'relu5')
drop3 = brew.dropout(m, relu5, 'drop3', ratio=0.5)
fc2 = brew.fc(m, drop3, 'fc2', dim_in=512, dim_out=n_classes)
softmax = brew.softmax(m, fc2, 'softmax')
return softmax
def add_training_operators(softmax, m, device_opts=DEVICE_OPTS, lr=LR, momentum=MOMENTUM) :
with core.DeviceScope(device_opts):
xent = m.LabelCrossEntropy([softmax, "label"], 'xent')
loss = m.AveragedLoss(xent, "loss")
m.AddGradientOperators([loss])
opt = optimizer.build_sgd(m,
base_learning_rate=lr,
policy='fixed',
momentum=momentum)
def init_model(x_train, y_train, batchsize=BATCHSIZE, device_opts=DEVICE_OPTS):
# Create Place-holder for data
workspace.FeedBlob("data", x_train[:batchsize], device_option=device_opts)
workspace.FeedBlob("label", y_train[:batchsize], device_option=device_opts)
# Initialise model
train_arg_scope = {
'order': 'NCHW',
'use_cudnn': True,
'cudnn_exhaustive_search': True}
train_model = model_helper.ModelHelper(
name="train_net", arg_scope=train_arg_scope)
softmax = create_model(train_model, device_opts=device_opts)
add_training_operators(softmax, train_model, device_opts=device_opts)
# Initialise workspace
workspace.RunNetOnce(train_model.param_init_net)
workspace.CreateNet(train_model.net)
return train_model
%%time
# Data into format for library
x_train, x_test, y_train, y_test = cifar_for_library(channel_first=True)
print(x_train.shape, x_test.shape, y_train.shape, y_test.shape)
print(x_train.dtype, x_test.dtype, y_train.dtype, y_test.dtype)
%%time
# Initialise model
model = init_model(x_train, y_train)
%%time
# Main training loop: 51s
for j in range(EPOCHS):
for data, label in yield_mb(x_train, y_train, BATCHSIZE, shuffle=True):
# Run one mini-batch at time
workspace.FeedBlob("data", data, device_option=DEVICE_OPTS)
workspace.FeedBlob("label", label, device_option=DEVICE_OPTS)
workspace.RunNet(model.net)
print("Finished epoch: ", j)
print(str(j) + ': ' + str(workspace.FetchBlob("loss")))
%%time
# Init test model
test_arg_scope = {
'order': 'NCHW',
'use_cudnn': True,
'cudnn_exhaustive_search': True,
'is_test': True,
}
test_model= model_helper.ModelHelper(name="test_net", init_params=False, arg_scope=test_arg_scope)
create_model(test_model, device_opts=DEVICE_OPTS)
workspace.RunNetOnce(test_model.param_init_net)
workspace.CreateNet(test_model.net, overwrite=True)
%%time
# Main evaluation loop: 311ms
n_samples = (y_test.shape[0]//BATCHSIZE)*BATCHSIZE
y_guess = np.zeros(n_samples, dtype=np.int)
y_truth = y_test[:n_samples]
c = 0
for data, label in yield_mb(x_test, y_test, BATCHSIZE):
workspace.FeedBlob("data", data, device_option=DEVICE_OPTS)
workspace.RunNet(test_model.net)
y_guess[c*BATCHSIZE:(c+1)*BATCHSIZE] = (np.argmax(workspace.FetchBlob("softmax"), axis=-1))
c += 1
print("Accuracy: ", sum(y_guess == y_truth)/float(len(y_guess)))
```
| github_jupyter |
<a href="https://colab.research.google.com/github/omarsar/pytorch_notebooks/blob/master/pytorch_hello_world.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# A First Shot at Deep Learning with PyTorch
In this notebook, we are going to take a baby step into the world of deep learning using PyTorch. There are a ton of notebooks out there that teach you the fundamentals of deep learning and PyTorch, so here the idea is to give you some basic introduction to deep learning and PyTorch at a very high level. Therefore, this notebook is targeting beginners but it can also serve as a review for more experienced developers.
After completion of this notebook, you are expected to know the basic components of training a basic neural network with PyTorch. I have also left a couple of exercises towards the end with the intention of encouraging more research and practise of your deep learning skills.
---
**Author:** Elvis Saravia([Twitter]((https://twitter.com/omarsar0)) | [LinkedIn](https://www.linkedin.com/in/omarsar/))
**Complete Code Walkthrough:** [Blog post]()
## Importing the libraries
Like with any other programming exercise, the first step is to import the necessary libraries. As we are going to be using Google Colab to program our neural network, we need to install and import the necessary PyTorch libraries.
```
!pip3 install torch torchvision
## The usual imports
import torch
import torch.nn as nn
## print out the pytorch version used
print(torch.__version__)
```
## The Neural Network

Before building and training a neural network the first step is to process and prepare the data. In this notebook, we are going to use syntethic data (i.e., fake data) so we won't be using any real world data.
For the sake of simplicity, we are going to use the following input and output pairs converted to tensors, which is how data is typically represented in the world of deep learning. The x values represent the input of dimension `(6,1)` and the y values represent the output of similar dimension. The example is taken from this [tutorial](https://github.com/lmoroney/dlaicourse/blob/master/Course%201%20-%20Part%202%20-%20Lesson%202%20-%20Notebook.ipynb).
The objective of the neural network model that we are going to build and train is to automatically learn patterns that better characterize the relationship between the `x` and `y` values. Essentially, the model learns the relationship that exists between inputs and outputs which can then be used to predict the corresponding `y` value for any given input `x`.
```
## our data in tensor form
x = torch.tensor([[-1.0], [0.0], [1.0], [2.0], [3.0], [4.0]], dtype=torch.float)
y = torch.tensor([[-3.0], [-1.0], [1.0], [3.0], [5.0], [7.0]], dtype=torch.float)
## print size of the input tensor
x.size()
```
## The Neural Network Components
As said earlier, we are going to first define and build out the components of our neural network before training the model.
### Model
Typically, when building a neural network model, we define the layers and weights which form the basic components of the model. Below we show an example of how to define a hidden layer named `layer1` with size `(1, 1)`. For the purpose of this tutorial, we won't explicitly define the `weights` and allow the built-in functions provided by PyTorch to handle that part for us. By the way, the `nn.Linear(...)` function applies a linear transformation ($y = xA^T + b$) to the data that was provided as its input. We ignore the bias for now by setting `bias=False`.
```
## Neural network with 1 hidden layer
layer1 = nn.Linear(1,1, bias=False)
model = nn.Sequential(layer1)
```
### Loss and Optimizer
The loss function, `nn.MSELoss()`, is in charge of letting the model know how good it has learned the relationship between the input and output. The optimizer (in this case an `SGD`) primary role is to minimize or lower that loss value as it tunes its weights.
```
## loss function
criterion = nn.MSELoss()
## optimizer algorithm
optimizer = torch.optim.SGD(model.parameters(), lr=0.01)
```
## Training the Neural Network Model
We have all the components we need to train our model. Below is the code used to train our model.
In simple terms, we train the model by feeding it the input and output pairs for a couple of rounds (i.e., `epoch`). After a series of forward and backward steps, the model somewhat learns the relationship between x and y values. This is notable by the decrease in the computed `loss`. For a more detailed explanation of this code check out this [tutorial](https://medium.com/dair-ai/a-simple-neural-network-from-scratch-with-pytorch-and-google-colab-c7f3830618e0).
```
## training
epoch = 150
for i in range(150):
model = model.train()
train_running_loss = 0.0
## forward
output = model(x)
loss = criterion(output, y)
optimizer.zero_grad()
## backward + update model params
loss.backward()
optimizer.step()
train_running_loss += loss.detach().item()
model.eval()
print('Epoch: %d | Loss: %.4f' %(i, train_running_loss) )
```
## Testing the Model
After training the model we have the ability to test the model predictive capability by passing it an input. Below is a simple example of how you could achieve this with our model. The result we obtained aligns with the results obtained in this [notebook](https://github.com/lmoroney/dlaicourse/blob/master/Course%201%20-%20Part%202%20-%20Lesson%202%20-%20Notebook.ipynb), which inspired this entire tutorial.
```
## test the model
sample = torch.tensor([10.0], dtype=torch.float)
predicted = model(sample)
print(predicted.detach().item())
```
## Final Words
Congratulations! In this tutorial you learned how to train a simple neural network using PyTorch. You also learned about the basic components that make up a neural network model such as the linear transformation layer, optimizer, and loss function. We then trained the model and tested its predictive capabilities. You are well on your way to become more knowledgeable about deep learning and PyTorch. I have provided a bunch of references below if you are interested in practising and learning more.
*I would like to thank Laurence Moroney for his excellent [tutorial](https://github.com/lmoroney/dlaicourse/blob/master/Course%201%20-%20Part%202%20-%20Lesson%202%20-%20Notebook.ipynb) which I used as an inspiration for this tutorial.*
## Exercises
- Add more examples in the input and output tensors. In addition, try to change the dimensions of the data, say by adding an extra value in each array. What needs to be changed to successfully train the network with the new data?
- The model converged really fast, which means it learned the relationship between x and y values after a couple of iterations. Do you think it makes sense to continue training? How would you automate the process of stopping the training after the model loss doesn't subtantially change?
- In our example, we used a single hidden layer. Try to take a look at the PyTorch documentation to figure out what you need to do to get a model with more layers. What happens if you add more hidden layers?
- We did not discuss the learning rate (`lr-0.001`) and the optimizer in great detail. Check out the [PyTorch documentation](https://pytorch.org/docs/stable/optim.html) to learn more about what other optimizers you can use.
## References
- [The Hello World of Deep Learning with Neural Networks](https://github.com/lmoroney/dlaicourse/blob/master/Course%201%20-%20Part%202%20-%20Lesson%202%20-%20Notebook.ipynb)
- [A Simple Neural Network from Scratch with PyTorch and Google Colab](https://medium.com/dair-ai/a-simple-neural-network-from-scratch-with-pytorch-and-google-colab-c7f3830618e0?source=collection_category---4------1-----------------------)
- [PyTorch Official Docs](https://pytorch.org/docs/stable/nn.html)
- [PyTorch 1.2 Quickstart with Google Colab](https://medium.com/dair-ai/pytorch-1-2-quickstart-with-google-colab-6690a30c38d)
- [A Gentle Intoduction to PyTorch](https://medium.com/dair-ai/pytorch-1-2-introduction-guide-f6fa9bb7597c)
| github_jupyter |
# Loading and Preprocessing Data with TensorFlow
> Chapter 13
- permalink: /13_loading_and_preprocessing_data
_This notebook contains all the sample code and solutions to the exercises in chapter 13._
# Setup
First, let's import a few common modules, ensure MatplotLib plots figures inline and prepare a function to save the figures. We also check that Python 3.5 or later is installed (although Python 2.x may work, it is deprecated so we strongly recommend you use Python 3 instead), as well as Scikit-Learn ≥0.20 and TensorFlow ≥2.0.
```
# collapse-show
# Python ≥3.5 is required
import sys
assert sys.version_info >= (3, 5)
# Scikit-Learn ≥0.20 is required
import sklearn
assert sklearn.__version__ >= "0.20"
try:
# %tensorflow_version only exists in Colab.
%tensorflow_version 2.x
!pip install -q -U tfx==0.15.0rc0
print("You can safely ignore the package incompatibility errors.")
except Exception:
pass
# TensorFlow ≥2.0 is required
import tensorflow as tf
from tensorflow import keras
assert tf.__version__ >= "2.0"
# Common imports
import numpy as np
import os
# to make this notebook's output stable across runs
np.random.seed(42)
# To plot pretty figures
%matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rc('axes', labelsize=14)
mpl.rc('xtick', labelsize=12)
mpl.rc('ytick', labelsize=12)
# Where to save the figures
PROJECT_ROOT_DIR = "."
CHAPTER_ID = "data"
IMAGES_PATH = os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID)
os.makedirs(IMAGES_PATH, exist_ok=True)
def save_fig(fig_id, tight_layout=True, fig_extension="png", resolution=300):
path = os.path.join(IMAGES_PATH, fig_id + "." + fig_extension)
print("Saving figure", fig_id)
if tight_layout:
plt.tight_layout()
plt.savefig(path, format=fig_extension, dpi=resolution)
```
## Datasets
```
X = tf.range(10)
dataset = tf.data.Dataset.from_tensor_slices(X)
dataset
```
Equivalently:
```
dataset = tf.data.Dataset.range(10)
for item in dataset:
print(item)
dataset = dataset.repeat(3).batch(7)
for item in dataset:
print(item)
dataset = dataset.map(lambda x: x * 2)
for item in dataset:
print(item)
dataset = dataset.apply(tf.data.experimental.unbatch())
dataset = dataset.filter(lambda x: x < 10) # keep only items < 10
for item in dataset.take(3):
print(item)
dataset = tf.data.Dataset.range(10).repeat(3)
dataset = dataset.shuffle(buffer_size=3, seed=42).batch(7)
for item in dataset:
print(item)
```
## Split the California dataset to multiple CSV files
Let's start by loading and preparing the California housing dataset. We first load it, then split it into a training set, a validation set and a test set, and finally we scale it:
```
from sklearn.datasets import fetch_california_housing
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
housing = fetch_california_housing()
X_train_full, X_test, y_train_full, y_test = train_test_split(
housing.data, housing.target.reshape(-1, 1), random_state=42)
X_train, X_valid, y_train, y_valid = train_test_split(
X_train_full, y_train_full, random_state=42)
scaler = StandardScaler()
scaler.fit(X_train)
X_mean = scaler.mean_
X_std = scaler.scale_
```
For a very large dataset that does not fit in memory, you will typically want to split it into many files first, then have TensorFlow read these files in parallel. To demonstrate this, let's start by splitting the housing dataset and save it to 20 CSV files:
```
def save_to_multiple_csv_files(data, name_prefix, header=None, n_parts=10):
housing_dir = os.path.join("datasets", "housing")
os.makedirs(housing_dir, exist_ok=True)
path_format = os.path.join(housing_dir, "my_{}_{:02d}.csv")
filepaths = []
m = len(data)
for file_idx, row_indices in enumerate(np.array_split(np.arange(m), n_parts)):
part_csv = path_format.format(name_prefix, file_idx)
filepaths.append(part_csv)
with open(part_csv, "wt", encoding="utf-8") as f:
if header is not None:
f.write(header)
f.write("\n")
for row_idx in row_indices:
f.write(",".join([repr(col) for col in data[row_idx]]))
f.write("\n")
return filepaths
train_data = np.c_[X_train, y_train]
valid_data = np.c_[X_valid, y_valid]
test_data = np.c_[X_test, y_test]
header_cols = housing.feature_names + ["MedianHouseValue"]
header = ",".join(header_cols)
train_filepaths = save_to_multiple_csv_files(train_data, "train", header, n_parts=20)
valid_filepaths = save_to_multiple_csv_files(valid_data, "valid", header, n_parts=10)
test_filepaths = save_to_multiple_csv_files(test_data, "test", header, n_parts=10)
```
Okay, now let's take a peek at the first few lines of one of these CSV files:
```
import pandas as pd
pd.read_csv(train_filepaths[0]).head()
```
Or in text mode:
```
with open(train_filepaths[0]) as f:
for i in range(5):
print(f.readline(), end="")
train_filepaths
```
## Building an Input Pipeline
```
filepath_dataset = tf.data.Dataset.list_files(train_filepaths, seed=42)
for filepath in filepath_dataset:
print(filepath)
n_readers = 5
dataset = filepath_dataset.interleave(
lambda filepath: tf.data.TextLineDataset(filepath).skip(1),
cycle_length=n_readers)
for line in dataset.take(5):
print(line.numpy())
```
Notice that field 4 is interpreted as a string.
```
record_defaults=[0, np.nan, tf.constant(np.nan, dtype=tf.float64), "Hello", tf.constant([])]
parsed_fields = tf.io.decode_csv('1,2,3,4,5', record_defaults)
parsed_fields
```
Notice that all missing fields are replaced with their default value, when provided:
```
parsed_fields = tf.io.decode_csv(',,,,5', record_defaults)
parsed_fields
```
The 5th field is compulsory (since we provided `tf.constant([])` as the "default value"), so we get an exception if we do not provide it:
```
try:
parsed_fields = tf.io.decode_csv(',,,,', record_defaults)
except tf.errors.InvalidArgumentError as ex:
print(ex)
```
The number of fields should match exactly the number of fields in the `record_defaults`:
```
try:
parsed_fields = tf.io.decode_csv('1,2,3,4,5,6,7', record_defaults)
except tf.errors.InvalidArgumentError as ex:
print(ex)
n_inputs = 8 # X_train.shape[-1]
@tf.function
def preprocess(line):
defs = [0.] * n_inputs + [tf.constant([], dtype=tf.float32)]
fields = tf.io.decode_csv(line, record_defaults=defs)
x = tf.stack(fields[:-1])
y = tf.stack(fields[-1:])
return (x - X_mean) / X_std, y
preprocess(b'4.2083,44.0,5.3232,0.9171,846.0,2.3370,37.47,-122.2,2.782')
def csv_reader_dataset(filepaths, repeat=1, n_readers=5,
n_read_threads=None, shuffle_buffer_size=10000,
n_parse_threads=5, batch_size=32):
dataset = tf.data.Dataset.list_files(filepaths).repeat(repeat)
dataset = dataset.interleave(
lambda filepath: tf.data.TextLineDataset(filepath).skip(1),
cycle_length=n_readers, num_parallel_calls=n_read_threads)
dataset = dataset.shuffle(shuffle_buffer_size)
dataset = dataset.map(preprocess, num_parallel_calls=n_parse_threads)
dataset = dataset.batch(batch_size)
return dataset.prefetch(1)
train_set = csv_reader_dataset(train_filepaths, batch_size=3)
for X_batch, y_batch in train_set.take(2):
print("X =", X_batch)
print("y =", y_batch)
print()
train_set = csv_reader_dataset(train_filepaths, repeat=None)
valid_set = csv_reader_dataset(valid_filepaths)
test_set = csv_reader_dataset(test_filepaths)
model = keras.models.Sequential([
keras.layers.Dense(30, activation="relu", input_shape=X_train.shape[1:]),
keras.layers.Dense(1),
])
model.compile(loss="mse", optimizer=keras.optimizers.SGD(lr=1e-3))
batch_size = 32
model.fit(train_set, steps_per_epoch=len(X_train) // batch_size, epochs=10,
validation_data=valid_set)
model.evaluate(test_set, steps=len(X_test) // batch_size)
new_set = test_set.map(lambda X, y: X) # we could instead just pass test_set, Keras would ignore the labels
X_new = X_test
model.predict(new_set, steps=len(X_new) // batch_size)
optimizer = keras.optimizers.Nadam(lr=0.01)
loss_fn = keras.losses.mean_squared_error
n_epochs = 5
batch_size = 32
n_steps_per_epoch = len(X_train) // batch_size
total_steps = n_epochs * n_steps_per_epoch
global_step = 0
for X_batch, y_batch in train_set.take(total_steps):
global_step += 1
print("\rGlobal step {}/{}".format(global_step, total_steps), end="")
with tf.GradientTape() as tape:
y_pred = model(X_batch)
main_loss = tf.reduce_mean(loss_fn(y_batch, y_pred))
loss = tf.add_n([main_loss] + model.losses)
gradients = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(gradients, model.trainable_variables))
optimizer = keras.optimizers.Nadam(lr=0.01)
loss_fn = keras.losses.mean_squared_error
@tf.function
def train(model, n_epochs, batch_size=32,
n_readers=5, n_read_threads=5, shuffle_buffer_size=10000, n_parse_threads=5):
train_set = csv_reader_dataset(train_filepaths, repeat=n_epochs, n_readers=n_readers,
n_read_threads=n_read_threads, shuffle_buffer_size=shuffle_buffer_size,
n_parse_threads=n_parse_threads, batch_size=batch_size)
for X_batch, y_batch in train_set:
with tf.GradientTape() as tape:
y_pred = model(X_batch)
main_loss = tf.reduce_mean(loss_fn(y_batch, y_pred))
loss = tf.add_n([main_loss] + model.losses)
gradients = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(gradients, model.trainable_variables))
train(model, 5)
optimizer = keras.optimizers.Nadam(lr=0.01)
loss_fn = keras.losses.mean_squared_error
@tf.function
def train(model, n_epochs, batch_size=32,
n_readers=5, n_read_threads=5, shuffle_buffer_size=10000, n_parse_threads=5):
train_set = csv_reader_dataset(train_filepaths, repeat=n_epochs, n_readers=n_readers,
n_read_threads=n_read_threads, shuffle_buffer_size=shuffle_buffer_size,
n_parse_threads=n_parse_threads, batch_size=batch_size)
n_steps_per_epoch = len(X_train) // batch_size
total_steps = n_epochs * n_steps_per_epoch
global_step = 0
for X_batch, y_batch in train_set.take(total_steps):
global_step += 1
if tf.equal(global_step % 100, 0):
tf.print("\rGlobal step", global_step, "/", total_steps)
with tf.GradientTape() as tape:
y_pred = model(X_batch)
main_loss = tf.reduce_mean(loss_fn(y_batch, y_pred))
loss = tf.add_n([main_loss] + model.losses)
gradients = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(gradients, model.trainable_variables))
train(model, 5)
```
Here is a short description of each method in the `Dataset` class:
```
for m in dir(tf.data.Dataset):
if not (m.startswith("_") or m.endswith("_")):
func = getattr(tf.data.Dataset, m)
if hasattr(func, "__doc__"):
print("● {:21s}{}".format(m + "()", func.__doc__.split("\n")[0]))
```
## The `TFRecord` binary format
A TFRecord file is just a list of binary records. You can create one using a `tf.io.TFRecordWriter`:
```
with tf.io.TFRecordWriter("my_data.tfrecord") as f:
f.write(b"This is the first record")
f.write(b"And this is the second record")
```
And you can read it using a `tf.data.TFRecordDataset`:
```
filepaths = ["my_data.tfrecord"]
dataset = tf.data.TFRecordDataset(filepaths)
for item in dataset:
print(item)
```
You can read multiple TFRecord files with just one `TFRecordDataset`. By default it will read them one at a time, but if you set `num_parallel_reads=3`, it will read 3 at a time in parallel and interleave their records:
```
filepaths = ["my_test_{}.tfrecord".format(i) for i in range(5)]
for i, filepath in enumerate(filepaths):
with tf.io.TFRecordWriter(filepath) as f:
for j in range(3):
f.write("File {} record {}".format(i, j).encode("utf-8"))
dataset = tf.data.TFRecordDataset(filepaths, num_parallel_reads=3)
for item in dataset:
print(item)
options = tf.io.TFRecordOptions(compression_type="GZIP")
with tf.io.TFRecordWriter("my_compressed.tfrecord", options) as f:
f.write(b"This is the first record")
f.write(b"And this is the second record")
dataset = tf.data.TFRecordDataset(["my_compressed.tfrecord"],
compression_type="GZIP")
for item in dataset:
print(item)
```
### A Brief Intro to Protocol Buffers
For this section you need to [install protobuf](https://developers.google.com/protocol-buffers/docs/downloads). In general you will not have to do so when using TensorFlow, as it comes with functions to create and parse protocol buffers of type `tf.train.Example`, which are generally sufficient. However, in this section we will learn about protocol buffers by creating our own simple protobuf definition, so we need the protobuf compiler (`protoc`): we will use it to compile the protobuf definition to a Python module that we can then use in our code.
First let's write a simple protobuf definition:
```
%%writefile person.proto
syntax = "proto3";
message Person {
string name = 1;
int32 id = 2;
repeated string email = 3;
}
```
And let's compile it (the `--descriptor_set_out` and `--include_imports` options are only required for the `tf.io.decode_proto()` example below):
```
!protoc person.proto --python_out=. --descriptor_set_out=person.desc --include_imports
!ls person*
from person_pb2 import Person
person = Person(name="Al", id=123, email=["a@b.com"]) # create a Person
print(person) # display the Person
person.name # read a field
person.name = "Alice" # modify a field
person.email[0] # repeated fields can be accessed like arrays
person.email.append("c@d.com") # add an email address
s = person.SerializeToString() # serialize to a byte string
s
person2 = Person() # create a new Person
person2.ParseFromString(s) # parse the byte string (27 bytes)
person == person2 # now they are equal
```
#### Custom protobuf
In rare cases, you may want to parse a custom protobuf (like the one we just created) in TensorFlow. For this you can use the `tf.io.decode_proto()` function:
```
person_tf = tf.io.decode_proto(
bytes=s,
message_type="Person",
field_names=["name", "id", "email"],
output_types=[tf.string, tf.int32, tf.string],
descriptor_source="person.desc")
person_tf.values
```
For more details, see the [`tf.io.decode_proto()`](https://www.tensorflow.org/api_docs/python/tf/io/decode_proto) documentation.
### TensorFlow Protobufs
Here is the definition of the tf.train.Example protobuf:
```proto
syntax = "proto3";
message BytesList { repeated bytes value = 1; }
message FloatList { repeated float value = 1 [packed = true]; }
message Int64List { repeated int64 value = 1 [packed = true]; }
message Feature {
oneof kind {
BytesList bytes_list = 1;
FloatList float_list = 2;
Int64List int64_list = 3;
}
};
message Features { map<string, Feature> feature = 1; };
message Example { Features features = 1; };
```
```
# WARNING: there's currently a bug preventing "from tensorflow.train import X"
# so we work around it by writing "X = tf.train.X"
#from tensorflow.train import BytesList, FloatList, Int64List
#from tensorflow.train import Feature, Features, Example
BytesList = tf.train.BytesList
FloatList = tf.train.FloatList
Int64List = tf.train.Int64List
Feature = tf.train.Feature
Features = tf.train.Features
Example = tf.train.Example
person_example = Example(
features=Features(
feature={
"name": Feature(bytes_list=BytesList(value=[b"Alice"])),
"id": Feature(int64_list=Int64List(value=[123])),
"emails": Feature(bytes_list=BytesList(value=[b"a@b.com", b"c@d.com"]))
}))
with tf.io.TFRecordWriter("my_contacts.tfrecord") as f:
f.write(person_example.SerializeToString())
feature_description = {
"name": tf.io.FixedLenFeature([], tf.string, default_value=""),
"id": tf.io.FixedLenFeature([], tf.int64, default_value=0),
"emails": tf.io.VarLenFeature(tf.string),
}
for serialized_example in tf.data.TFRecordDataset(["my_contacts.tfrecord"]):
parsed_example = tf.io.parse_single_example(serialized_example,
feature_description)
parsed_example
parsed_example
parsed_example["emails"].values[0]
tf.sparse.to_dense(parsed_example["emails"], default_value=b"")
parsed_example["emails"].values
```
### Putting Images in TFRecords
```
from sklearn.datasets import load_sample_images
img = load_sample_images()["images"][0]
plt.imshow(img)
plt.axis("off")
plt.title("Original Image")
plt.show()
data = tf.io.encode_jpeg(img)
example_with_image = Example(features=Features(feature={
"image": Feature(bytes_list=BytesList(value=[data.numpy()]))}))
serialized_example = example_with_image.SerializeToString()
# then save to TFRecord
feature_description = { "image": tf.io.VarLenFeature(tf.string) }
example_with_image = tf.io.parse_single_example(serialized_example, feature_description)
decoded_img = tf.io.decode_jpeg(example_with_image["image"].values[0])
```
Or use `decode_image()` which supports BMP, GIF, JPEG and PNG formats:
```
decoded_img = tf.io.decode_image(example_with_image["image"].values[0])
plt.imshow(decoded_img)
plt.title("Decoded Image")
plt.axis("off")
plt.show()
```
### Putting Tensors and Sparse Tensors in TFRecords
Tensors can be serialized and parsed easily using `tf.io.serialize_tensor()` and `tf.io.parse_tensor()`:
```
t = tf.constant([[0., 1.], [2., 3.], [4., 5.]])
s = tf.io.serialize_tensor(t)
s
tf.io.parse_tensor(s, out_type=tf.float32)
serialized_sparse = tf.io.serialize_sparse(parsed_example["emails"])
serialized_sparse
BytesList(value=serialized_sparse.numpy())
dataset = tf.data.TFRecordDataset(["my_contacts.tfrecord"]).batch(10)
for serialized_examples in dataset:
parsed_examples = tf.io.parse_example(serialized_examples,
feature_description)
parsed_examples
```
## Handling Sequential Data Using `SequenceExample`
```proto
syntax = "proto3";
message FeatureList { repeated Feature feature = 1; };
message FeatureLists { map<string, FeatureList> feature_list = 1; };
message SequenceExample {
Features context = 1;
FeatureLists feature_lists = 2;
};
```
```
# WARNING: there's currently a bug preventing "from tensorflow.train import X"
# so we work around it by writing "X = tf.train.X"
#from tensorflow.train import FeatureList, FeatureLists, SequenceExample
FeatureList = tf.train.FeatureList
FeatureLists = tf.train.FeatureLists
SequenceExample = tf.train.SequenceExample
context = Features(feature={
"author_id": Feature(int64_list=Int64List(value=[123])),
"title": Feature(bytes_list=BytesList(value=[b"A", b"desert", b"place", b"."])),
"pub_date": Feature(int64_list=Int64List(value=[1623, 12, 25]))
})
content = [["When", "shall", "we", "three", "meet", "again", "?"],
["In", "thunder", ",", "lightning", ",", "or", "in", "rain", "?"]]
comments = [["When", "the", "hurlyburly", "'s", "done", "."],
["When", "the", "battle", "'s", "lost", "and", "won", "."]]
def words_to_feature(words):
return Feature(bytes_list=BytesList(value=[word.encode("utf-8")
for word in words]))
content_features = [words_to_feature(sentence) for sentence in content]
comments_features = [words_to_feature(comment) for comment in comments]
sequence_example = SequenceExample(
context=context,
feature_lists=FeatureLists(feature_list={
"content": FeatureList(feature=content_features),
"comments": FeatureList(feature=comments_features)
}))
sequence_example
serialized_sequence_example = sequence_example.SerializeToString()
context_feature_descriptions = {
"author_id": tf.io.FixedLenFeature([], tf.int64, default_value=0),
"title": tf.io.VarLenFeature(tf.string),
"pub_date": tf.io.FixedLenFeature([3], tf.int64, default_value=[0, 0, 0]),
}
sequence_feature_descriptions = {
"content": tf.io.VarLenFeature(tf.string),
"comments": tf.io.VarLenFeature(tf.string),
}
parsed_context, parsed_feature_lists = tf.io.parse_single_sequence_example(
serialized_sequence_example, context_feature_descriptions,
sequence_feature_descriptions)
parsed_context
parsed_context["title"].values
parsed_feature_lists
print(tf.RaggedTensor.from_sparse(parsed_feature_lists["content"]))
```
# The Features API
Let's use the variant of the California housing dataset that we used in Chapter 2, since it contains categorical features and missing values:
```
import os
import tarfile
import urllib
DOWNLOAD_ROOT = "https://raw.githubusercontent.com/ageron/handson-ml2/master/"
HOUSING_PATH = os.path.join("datasets", "housing")
HOUSING_URL = DOWNLOAD_ROOT + "datasets/housing/housing.tgz"
def fetch_housing_data(housing_url=HOUSING_URL, housing_path=HOUSING_PATH):
os.makedirs(housing_path, exist_ok=True)
tgz_path = os.path.join(housing_path, "housing.tgz")
urllib.request.urlretrieve(housing_url, tgz_path)
housing_tgz = tarfile.open(tgz_path)
housing_tgz.extractall(path=housing_path)
housing_tgz.close()
fetch_housing_data()
import pandas as pd
def load_housing_data(housing_path=HOUSING_PATH):
csv_path = os.path.join(housing_path, "housing.csv")
return pd.read_csv(csv_path)
housing = load_housing_data()
housing.head()
housing_median_age = tf.feature_column.numeric_column("housing_median_age")
age_mean, age_std = X_mean[1], X_std[1] # The median age is column in 1
housing_median_age = tf.feature_column.numeric_column(
"housing_median_age", normalizer_fn=lambda x: (x - age_mean) / age_std)
median_income = tf.feature_column.numeric_column("median_income")
bucketized_income = tf.feature_column.bucketized_column(
median_income, boundaries=[1.5, 3., 4.5, 6.])
bucketized_income
ocean_prox_vocab = ['<1H OCEAN', 'INLAND', 'ISLAND', 'NEAR BAY', 'NEAR OCEAN']
ocean_proximity = tf.feature_column.categorical_column_with_vocabulary_list(
"ocean_proximity", ocean_prox_vocab)
ocean_proximity
# Just an example, it's not used later on
city_hash = tf.feature_column.categorical_column_with_hash_bucket(
"city", hash_bucket_size=1000)
city_hash
bucketized_age = tf.feature_column.bucketized_column(
housing_median_age, boundaries=[-1., -0.5, 0., 0.5, 1.]) # age was scaled
age_and_ocean_proximity = tf.feature_column.crossed_column(
[bucketized_age, ocean_proximity], hash_bucket_size=100)
latitude = tf.feature_column.numeric_column("latitude")
longitude = tf.feature_column.numeric_column("longitude")
bucketized_latitude = tf.feature_column.bucketized_column(
latitude, boundaries=list(np.linspace(32., 42., 20 - 1)))
bucketized_longitude = tf.feature_column.bucketized_column(
longitude, boundaries=list(np.linspace(-125., -114., 20 - 1)))
location = tf.feature_column.crossed_column(
[bucketized_latitude, bucketized_longitude], hash_bucket_size=1000)
ocean_proximity_one_hot = tf.feature_column.indicator_column(ocean_proximity)
ocean_proximity_embed = tf.feature_column.embedding_column(ocean_proximity,
dimension=2)
```
### Using Feature Columns for Parsing
```
median_house_value = tf.feature_column.numeric_column("median_house_value")
columns = [housing_median_age, median_house_value]
feature_descriptions = tf.feature_column.make_parse_example_spec(columns)
feature_descriptions
with tf.io.TFRecordWriter("my_data_with_features.tfrecords") as f:
for x, y in zip(X_train[:, 1:2], y_train):
example = Example(features=Features(feature={
"housing_median_age": Feature(float_list=FloatList(value=[x])),
"median_house_value": Feature(float_list=FloatList(value=[y]))
}))
f.write(example.SerializeToString())
def parse_examples(serialized_examples):
examples = tf.io.parse_example(serialized_examples, feature_descriptions)
targets = examples.pop("median_house_value") # separate the targets
return examples, targets
batch_size = 32
dataset = tf.data.TFRecordDataset(["my_data_with_features.tfrecords"])
dataset = dataset.repeat().shuffle(10000).batch(batch_size).map(parse_examples)
```
**Warning**: the `DenseFeatures` layer currently does not work with the Functional API, see [TF issue #27416](https://github.com/tensorflow/tensorflow/issues/27416). Hopefully this will be resolved before the final release of TF 2.0.
```
columns_without_target = columns[:-1]
model = keras.models.Sequential([
keras.layers.DenseFeatures(feature_columns=columns_without_target),
keras.layers.Dense(1)
])
model.compile(loss="mse",
optimizer=keras.optimizers.SGD(lr=1e-3),
metrics=["accuracy"])
model.fit(dataset, steps_per_epoch=len(X_train) // batch_size, epochs=5)
some_columns = [ocean_proximity_embed, bucketized_income]
dense_features = keras.layers.DenseFeatures(some_columns)
dense_features({
"ocean_proximity": [["NEAR OCEAN"], ["INLAND"], ["INLAND"]],
"median_income": [[3.], [7.2], [1.]]
})
```
# TF Transform
```
try:
import tensorflow_transform as tft
def preprocess(inputs): # inputs is a batch of input features
median_age = inputs["housing_median_age"]
ocean_proximity = inputs["ocean_proximity"]
standardized_age = tft.scale_to_z_score(median_age - tft.mean(median_age))
ocean_proximity_id = tft.compute_and_apply_vocabulary(ocean_proximity)
return {
"standardized_median_age": standardized_age,
"ocean_proximity_id": ocean_proximity_id
}
except ImportError:
print("TF Transform is not installed. Try running: pip3 install -U tensorflow-transform")
```
# TensorFlow Datasets
```
import tensorflow_datasets as tfds
datasets = tfds.load(name="mnist")
mnist_train, mnist_test = datasets["train"], datasets["test"]
print(tfds.list_builders())
plt.figure(figsize=(6,3))
mnist_train = mnist_train.repeat(5).batch(32).prefetch(1)
for item in mnist_train:
images = item["image"]
labels = item["label"]
for index in range(5):
plt.subplot(1, 5, index + 1)
image = images[index, ..., 0]
label = labels[index].numpy()
plt.imshow(image, cmap="binary")
plt.title(label)
plt.axis("off")
break # just showing part of the first batch
datasets = tfds.load(name="mnist")
mnist_train, mnist_test = datasets["train"], datasets["test"]
mnist_train = mnist_train.repeat(5).batch(32)
mnist_train = mnist_train.map(lambda items: (items["image"], items["label"]))
mnist_train = mnist_train.prefetch(1)
for images, labels in mnist_train.take(1):
print(images.shape)
print(labels.numpy())
datasets = tfds.load(name="mnist", batch_size=32, as_supervised=True)
mnist_train = datasets["train"].repeat().prefetch(1)
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28, 1]),
keras.layers.Lambda(lambda images: tf.cast(images, tf.float32)),
keras.layers.Dense(10, activation="softmax")])
model.compile(loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.SGD(lr=1e-3),
metrics=["accuracy"])
model.fit(mnist_train, steps_per_epoch=60000 // 32, epochs=5)
```
# TensorFlow Hub
```
import tensorflow_hub as hub
hub_layer = hub.KerasLayer("https://tfhub.dev/google/tf2-preview/nnlm-en-dim50/1",
output_shape=[50], input_shape=[], dtype=tf.string)
model = keras.Sequential()
model.add(hub_layer)
model.add(keras.layers.Dense(16, activation='relu'))
model.add(keras.layers.Dense(1, activation='sigmoid'))
model.summary()
sentences = tf.constant(["It was a great movie", "The actors were amazing"])
embeddings = hub_layer(sentences)
embeddings
```
| github_jupyter |
```
%matplotlib inline
from matplotlib import style
style.use('fivethirtyeight')
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import datetime as dt
```
# Reflect Tables into SQLAlchemy ORM
```
# Python SQL toolkit and Object Relational Mapper
import sqlalchemy
from sqlalchemy.ext.automap import automap_base
from sqlalchemy.orm import Session
from sqlalchemy import create_engine, func
engine = create_engine("sqlite:///Resources/hawaii.sqlite")
# reflect an existing database into a new model
Base = automap_base()
# reflect the tables
Base.prepare(engine, reflect=True)
# We can view all of the classes that automap found
Base.classes.keys()
# Save references to each table
Measurement = Base.classes.measurement
Station = Base.classes.station
# Create our session (link) from Python to the DB
session = Session(engine)
```
# Exploratory Climate Analysis
```
# Design a query to retrieve the last 12 months of precipitation data and plot the results
# Calculate the date 1 year ago from the last data point in the database
querydate = session.query(Measurement.date).order_by(Measurement.date.desc()).first()
lastdate = "2016-08-23"
# Perform a query to retrieve the data and precipitation scores
prcp_data = session.query(Measurement.date,Measurement.prcp).filter(Measurement.date >= lastdate).all()
# Save the query results as a Pandas DataFrame and set the index to the date column
prcp_data_df = pd.DataFrame(prcp_data, columns = ['Date','Precipitation'])
prcp_data_df.set_index('Date',inplace = True)
# Sort the dataframe by date
prcp_data_df = prcp_data_df.sort_values("Date")
prcp_data_df
# Use Pandas Plotting with Matplotlib to plot the data
prcp_data_df.plot(rot = 90, figsize = (15,15))
plt.xlabel("Date")
plt.ylabel("Inches")
# Use Pandas to calcualte the summary statistics for the precipitation data
prcp_data_df.describe()
# Design a query to show how many stations are available in this dataset?
total_stations = session.query(func.count(Station.station)).all()
total_stations
# What are the most active stations? (i.e. what stations have the most rows)?
# List the stations and the counts in descending order.
max_station = session.query(Measurement.station, func.count(Measurement.station)).\
group_by(Measurement.station).order_by(func.count(Measurement.station).desc()).all()
max_station
most_active = "USC00519281"
# Using the station id from the previous query, calculate the lowest temperature recorded,
# highest temperature recorded, and average temperature of the most active station?
most_active_data = session.query(func.min(Measurement.tobs), func.max(Measurement.tobs), func.avg(Measurement.tobs)).\
filter(Measurement.station == most_active).all()
most_active_data
# Choose the station with the highest number of temperature observations.
# Query the last 12 months of temperature observation data for this station and plot the results as a histogram
final_result = session.query(Measurement.tobs).filter(Measurement.station == most_active).\
filter(Measurement.date >= lastdate).all()
final_result_df = pd.DataFrame(final_result, columns = ["Temp"])
final_result_df
final_result_df.plot.hist(bins=12)
```
## Bonus Challenge Assignment
```
# This function called `calc_temps` will accept start date and end date in the format '%Y-%m-%d'
# and return the minimum, average, and maximum temperatures for that range of dates
def calc_temps(start_date, end_date):
"""TMIN, TAVG, and TMAX for a list of dates.
Args:
start_date (string): A date string in the format %Y-%m-%d
end_date (string): A date string in the format %Y-%m-%d
Returns:
TMIN, TAVE, and TMAX
"""
return session.query(func.min(Measurement.tobs), func.avg(Measurement.tobs), func.max(Measurement.tobs)).\
filter(Measurement.date >= start_date).filter(Measurement.date <= end_date).all()
# function usage example
print(calc_temps('2012-02-28', '2012-03-05'))
# Use your previous function `calc_temps` to calculate the tmin, tavg, and tmax
# for your trip using the previous year's data for those same dates.
# Plot the results from your previous query as a bar chart.
# Use "Trip Avg Temp" as your Title
# Use the average temperature for the y value
# Use the peak-to-peak (tmax-tmin) value as the y error bar (yerr)
# Calculate the total amount of rainfall per weather station for your trip dates using the previous year's matching dates.
# Sort this in descending order by precipitation amount and list the station, name, latitude, longitude, and elevation
# Create a query that will calculate the daily normals
# (i.e. the averages for tmin, tmax, and tavg for all historic data matching a specific month and day)
def daily_normals(date):
"""Daily Normals.
Args:
date (str): A date string in the format '%m-%d'
Returns:
A list of tuples containing the daily normals, tmin, tavg, and tmax
"""
sel = [func.min(Measurement.tobs), func.avg(Measurement.tobs), func.max(Measurement.tobs)]
return session.query(*sel).filter(func.strftime("%m-%d", Measurement.date) == date).all()
daily_normals("01-01")
# calculate the daily normals for your trip
# push each tuple of calculations into a list called `normals`
# Set the start and end date of the trip
# Use the start and end date to create a range of dates
# Stip off the year and save a list of %m-%d strings
# Loop through the list of %m-%d strings and calculate the normals for each date
# Load the previous query results into a Pandas DataFrame and add the `trip_dates` range as the `date` index
# Plot the daily normals as an area plot with `stacked=False`
```
| github_jupyter |
# Numpyro Integration
> This notebook gives a demonstration of how one can use some of the numpyro integration features to enable one to offload some of the inference schemes. This will allow the user to focus more on GP model construction instead of GP model inference. Below we show some of the most important functions and features available within GPJax that will allow users to easily transform their models from standard GPJax to numpyro-like.`
```
%load_ext lab_black
import gpjax
import gpjax.core as gpx
import gpviz as gpv
import numpyro
import jax
import jax.numpy as jnp
import jax.random as jr
import matplotlib.pyplot as plt
# plot methods
import matplotlib.pyplot as plt
import seaborn as sns
sns.reset_defaults()
sns.set_context(context="talk", font_scale=0.7)
plt.style.use(gpv.__stylesheet__)
key = jr.PRNGKey(123)
print(gpjax.__version__)
from pprint import pprint
%matplotlib inline
%load_ext autoreload
%autoreload 2
```
## Synthetic datasets
With the necessary libraries and modules imported we can simulate some data. We'll sample our inputs $x$ according to $x_1, \ldots , x_{50} \sim \mathcal{U}(-3., 3)$ and corresponding outputs will be,
$$y_i = \sin(4x_i) + \cos(2 x_i) + \epsilon_i,$$
such that $\epsilon_i \sim \mathcal{N}(0, 0.2)$.
```
N = 50
noise = 0.2
x = jr.uniform(key=key, minval=-3.0, maxval=3.0, shape=(N,)).sort().reshape(-1, 1)
f = lambda x: jnp.sin(4 * x) + jnp.cos(2 * x)
signal = f(x)
y = signal + jr.normal(key, shape=signal.shape) * noise
xtest = jnp.linspace(-3.5, 3.5, 500).reshape(-1, 1)
ytest = f(xtest)
fig, ax = plt.subplots(figsize=(10, 5))
ax.plot(xtest, ytest, label="Latent function")
ax.plot(x, y, "o", label="Observations")
ax.legend(loc="best")
plt.show()
```
## Gaussian Process Regression Model
Now in the original regression notebook, we showed how one can define a Gaussian process in GPJax. We need the following:
* A Prior
* mean function
* kernel function
* A Likelihood
Then, using the `*` operator, we get a GP posterior.
```
# GP Prior
mean_function = gpx.Zero()
kernel = gpx.RBF()
prior = gpx.Prior(mean_function=mean_function, kernel=kernel)
# GP Likelihood
lik = gpx.Gaussian()
# GP Posterior
posterior = prior * lik
```
### Parameters
The previous notebook also spent a good portion showing how one can initialize and constrain the parameters using the `GPJax` library. So below we will use the `initialise` function which will return a dictionary of the appropriate parameters.
```
params = gpx.initialise(posterior)
pprint(params)
```
However, numpyro models make use of numpyro primitives such as `param` and `sample`. So we need to translate the dictionary from the `GPJax` library to a dictionary that will be ready by the numpyro model.
Here is an example where we can take any dictionary with the key value as the parameter name and the item as either a `float` or a `jnp.DeviceArray`.
```
from gpjax.interfaces.numpyro import numpyro_dict_params
numpyro_params = numpyro_dict_params(params)
pprint(numpyro_params)
```
The result is a nested dictionary with the appropriate parameters to be read into the numpyro model. Notice how we have some default constraints already defined. This is because numpyro has it's own `Constraint` class. The default in this case is the `constraint.positive`.
#### Modifying Constraints
We can also modify the parameters to other numpyro constraints via the `add_constraints` function. In the backend of `GPJax`, we have used multiple dispatch to allow different flavours of adding contraints. We show a few examples below:
**Example I** - We can add the same constraint everywhere.
```
from gpjax.interfaces.numpyro import add_constraints
from numpyro.distributions import constraints
# example 1 - adding positive for everything
numpyro_params_ = add_constraints(numpyro_params, constraints.softplus_positive)
pprint(numpyro_params_)
```
**Example II** - We can add a constraint to a specific variable.
```
# example 1 - adding positive for everything
numpyro_params_ = add_constraints(numpyro_params, "lengthscale", constraints.positive)
numpyro_params_
```
**Example III** - We can add a dictionary of the constraints we want to add where the key defines the variable.
```
# example 1 - adding positive for everything
new_constraints = {
"lengthscale": constraints.greater_than(0.01),
"obs_noise": constraints.interval(0.1, 0.001),
}
numpyro_params_ = add_constraints(numpyro_params, new_constraints)
# pprint(numpyro_params)
pprint(numpyro_params_)
```
Please take a look at the numpyro docs for more [constraints](http://num.pyro.ai/en/stable/distributions.html#constraints). Some other useful ones include the `softplus_positive`, `softplus_lower_cholesky`, and `unit_interval`.
## Numpyro Model
So once we have defined our parameters and constraints, we're ready to jump into the modeling! We have a specialized function called the `numpyro_marginal_ll`. This will take a gp posterior and a dictionary of `numpyro_params` that we have previously defined.
```
from gpjax.interfaces.numpyro import numpyro_marginal_ll
training_ds = gpx.Dataset(X=x, y=y)
# initialize parameters
params = gpx.initialise(posterior)
# convert to numpyro-style params
numpyro_params = numpyro_dict_params(params)
# initialize numpyro-style GP model
npy_model = numpyro_marginal_ll(posterior, numpyro_params)
```
The result is a callable function which takes in a `GPJax` `Dataset` (a datalcass consisting of an `X` and `y`). Below is an example of how we can input this dataset with our newly defined model as well as get the set of parameters defined within the function.
```
with numpyro.handlers.seed(rng_seed=123):
t_ = npy_model(training_ds)
print(t_.shape)
assert t_.shape[0] == x.shape[0]
t = numpyro.handlers.trace(npy_model).get_trace(training_ds)
print("Parameters:", list(t.keys()))
```
So the output size is the same as the input size and the parameters we obtain are the same (almost) as the input parameters.
### Approximate Inference with Guides
This is a very critical part of the numpyro and pyro framework. They use `guides` which work as approximate posteriors. They are fully parameterized and take in no observations. The simplest way to define a guide is to use the `autoguide` methods like the `AutoDelta`, `AutoLaplaceApproximation` and the `AutoNormal`. For more information, please see this excellent set of [tutorials](https://pyro.ai/examples/) provided by the num/pyro team.
In this example, we're going to use an `empty guide`. This works as a placeholder for now. By having an empty guide, we are effectively doing maximum likliehood (no priors). See [this tutorial](https://pyro.ai/examples/mle_map.html) for more information.
```
# approximate posterior
guide = lambda ds: None
```
### Optimization
So now that the model is defined (very simply), we are now ready to do optimization! These are the standard optimization procedures from numpyro.
```
%%time
from numpyro.infer import SVI, Trace_ELBO
# reproducibility
rng_key = jax.random.PRNGKey(0)
n_iterations = 2_500
# numpyro specific optimizer
optimizer = numpyro.optim.Adam(step_size=0.01)
# stochastic variational inference (pseudo)
svi = SVI(npy_model, guide, optimizer, loss=Trace_ELBO())
svi_results = svi.run(jax.random.PRNGKey(1), n_iterations, training_ds)
```
#### Losses
We can plot the losses by extracting them from the `svi_result`.
```
fig, ax = plt.subplots()
ax.plot(svi_results.losses)
ax.set(title="Loss", xlabel="Iterations", ylabel="ELBO")
plt.show()
```
### Extracting the Parameters
Oncew we;ve trained the model, we can now extract the learned the parameters. Because we were doing MLE, we don't need to worry about any unconventional naming schemes (see below) thus we can simply call them directly.
```
# Take them directly
learned_params = svi_results.params
pprint(learned_params)
```
And we see the parameters are ready to be used for predictions.
### Predictions
The rest of this section is **exactly** the same as the other tutorials. The resulting parameters have the same name, shape, type, etc. So you should not have to change your code from here. Quantities such as the posterior mean and variance can be evaluated in the usual fashion.
```
meanf = gpx.mean(posterior, learned_params, training_ds)
varf = gpx.variance(posterior, learned_params, training_ds)
mu = meanf(xtest).squeeze()
cov = varf(xtest).squeeze()
one_stddev = 1.96 * jnp.sqrt(jnp.diag(cov))
```
Similarily, plots can still be made using [GPViz](https://github.com/thomaspinder/GPViz).
```
fig, ax = plt.subplots(figsize=(10, 6))
gpv.plot(
key,
posterior,
learned_params,
training_ds,
xtest,
std_devs=[1, 3],
title="Posterior predictive",
ax=ax,
)
```
### Prior Parameters
Above, we showed how to do optimization on fixed priors with constraints. For this section, we want to highlight how one can attach priors to their hyperparameters.
```
import numpyro.distributions as dist
hyperpriors = {
"lengthscale": dist.Gamma(1.0, 1.0),
"variance": dist.HalfCauchy(scale=1.0),
"obs_noise": dist.HalfCauchy(scale=5.0),
}
```
For the most part, there is no difference between how one can add priors to the dictionary.
**Example I** - Add a Prior to a specific variable
```
from gpjax.interfaces.numpyro import add_priors
numpyro_params = numpyro_dict_params(params)
numpyro_params_ = add_priors(numpyro_params, "lengthscale", dist.Gamma(1.0, 1.0))
pprint(numpyro_params_)
```
**Example II** - Adding a dictionary of priors.
```
numpyro_params = numpyro_dict_params(params)
numpyro_params = add_priors(numpyro_params, hyperpriors)
pprint(numpyro_params)
```
#### Mixing Priors and Parameters
And yes, one can have a mixture of priors and parameters.
```
mixed_param_priors = {
"lengthscale": dist.Gamma(1.0, 1.0),
"variance": dist.HalfCauchy(scale=1.0),
"obs_noise": 1.0,
}
numpyro_params = numpyro_dict_params(mixed_param_priors)
pprint(numpyro_params)
```
**Warning**: Be careful with assigning constraints to priors. This will be ignored as numpyro has no method to add constraints on prior distributions. One would have to use another prior distribution which allows for constraints.
#### TensorFlow Probability
It's also worth mentioning that the numpyro devs were able to have some synergy between the [TFP](https://www.tensorflow.org/probability/examples/TensorFlow_Probability_on_JAX) package and the numpyro package via their simple [wrapper](http://num.pyro.ai/en/stable/distributions.html#tensorflow-distributions). You just need to import it:
```python
from numpyro.contrib.tfp import distributions as tfd
```
So this means that if there is a probability distribution that you TFP has that numpyro does not have, then most likely you can call that one.
### Maximum A Posteriori
So now that we have prior parameters, we need to use the guides in order to use stochastic variation inference (SVI). We saw above that an empty guide was used for MLE. But we'll find that the SVI method doesn't work if there is no approximate posterior to optimize over. So the simplest guide to use would be the `AutoDelta` which is equivalent to the MAP solution.
```
from numpyro.infer.autoguide import AutoDelta
# define hyperparameter priors
hyperpriors = {
"lengthscale": dist.Gamma(1.0, 1.0),
"variance": dist.HalfCauchy(scale=1.0),
"obs_noise": dist.HalfCauchy(scale=5.0),
}
# create numpyro dictionary
numpyro_params = numpyro_dict_params(hyperpriors)
# initialize model
npy_model = numpyro_marginal_ll(posterior, numpyro_params)
# Delta Guide
guide = AutoDelta(npy_model)
```
### Optimization
```
%%time
from numpyro.infer import SVI, Trace_ELBO
# reproducibility
rng_key = jax.random.PRNGKey(0)
n_iterations = 1_000
# numpyro specific optimizer
optimizer = numpyro.optim.Adam(step_size=0.01)
# stochastic variational inference (pseudo)
svi = SVI(npy_model, guide, optimizer, loss=Trace_ELBO())
svi_results = svi.run(rng_key, n_iterations, training_ds)
```
### Losses
```
fig, ax = plt.subplots()
ax.plot(svi_results.losses)
ax.set(title="Loss", xlabel="Iterations", ylabel="ELBO")
plt.show()
```
### Extracting the Parameters
Like the previous example, we can simple find the learned parameters.
```
# Take them directly
learned_params = svi_results.params
pprint(learned_params)
```
But you'll notice that there are some extra names attached. This is because these are parameters of the *approximate posterior*, namely the `loc` for the Delta distribution. A simple fix (in general) is to extract the median/mean parameters which will eliminate the extra tags on the name.
```
# Take them directly
median_params = guide.median(learned_params)
pprint(median_params)
```
### Predictions
Again, the rest of the code is **exactly the same**.
```
meanf = gpx.mean(posterior, median_params, training_ds)
varf = gpx.variance(posterior, median_params, training_ds)
mu = meanf(xtest).squeeze()
cov = varf(xtest).squeeze()
one_stddev = 1.96 * jnp.sqrt(jnp.diag(cov))
fig, ax = plt.subplots(figsize=(10, 6))
gpv.plot(
key,
posterior,
median_params,
training_ds,
xtest,
std_devs=[1, 3],
title="Posterior predictive",
ax=ax,
)
```
## Hamiltonian Monte Carlo
The last method for inference that we're going to look at is Monte Carlo. This is actually **la crème de la crème** for numpyro. They have a *blazing fast* HMC/NUTS inference scheme. In fact, JAX has been frequently praised for the speed of the MC schemes due to the `vmap` and `jit` primitives.
See this [paper](https://arxiv.org/abs/1912.11554) or this [blog post](https://luiarthur.github.io/TuringBnpBenchmarks/) for more details.
Below, we have an example of how one can use MCMC to do sampling. The model is exactly the same as the previous example except we do not need to define any guide functions.
```
hyperpriors = {
"lengthscale": dist.Gamma(1.0, 1.0),
"variance": dist.HalfCauchy(scale=1.0),
"obs_noise": dist.HalfCauchy(scale=5.0),
}
numpyro_params = numpyro_dict_params(hyperpriors)
# initialize model
npy_model = numpyro_marginal_ll(posterior, numpyro_params)
from numpyro.infer import MCMC, NUTS
# reproducibility
rng_key = jax.random.PRNGKey(0)
# NUTS Parameters
n_samples = 1_000
n_warmup = 100
# Setup
kernel = NUTS(npy_model)
nuts_sampling = MCMC(kernel, num_samples=n_samples, num_warmup=n_warmup)
nuts_sampling.run(rng_key, training_ds)
```
It's quite fast... This notebook was run on a CPU of 16 cores. We can also check the convergence with the convenient `print_summary()` method.
```
nuts_sampling.print_summary()
```
### Posterior Samples
We don't have any learned parameters so we have to extract the parameters manually. Below is an example along with a simple plot.
```
posterior_params_samples = nuts_sampling.get_samples()
# Plot posteriors for the parameers
fig, ax = plt.subplots(ncols=3, figsize=(12, 3))
sns.histplot(
ax=ax[0],
x=posterior_params_samples["lengthscale"],
kde=True,
bins=50,
stat="density",
)
sns.histplot(
ax=ax[1], x=posterior_params_samples["variance"], kde=True, bins=50, stat="density"
)
sns.histplot(
ax=ax[2], x=posterior_params_samples["obs_noise"], kde=True, bins=50, stat="density"
)
ax[0].set(title="Kernel Length Scale")
ax[1].set(title="Kernel Variance", ylabel="")
ax[2].set(title="Likelihood Noise", ylabel="")
plt.show()
```
### Median Parameters
We could also do the same `median` trick as below. Again, these are samples so they don't augment the names.
```
def get_median_parameters(samples: dict) -> dict:
median_params = {}
for ikey, iarray in samples.items():
median_params[ikey] = jnp.median(iarray)
return median_params
median_params = get_median_parameters(posterior_params_samples)
pprint(median_params)
```
### Predictions
We're repeating ourselves, but the predictions are **exactly the same** as all of the above examples.
```
meanf = gpx.mean(posterior, median_params, training_ds)
varf = gpx.variance(posterior, median_params, training_ds)
mu = meanf(xtest).squeeze()
cov = varf(xtest).squeeze()
one_stddev = 1.96 * jnp.sqrt(jnp.diag(cov))
fig, ax = plt.subplots(figsize=(10, 6))
gpv.plot(
key,
posterior,
median_params,
training_ds,
xtest,
std_devs=[1, 3],
title="Posterior predictive",
ax=ax,
)
```
### All Posteriors
Now, getting the median parameters is not really true Bayesian (although for this simple example, it's perfectly fine). Using some of the code from the previous notebook, we show how we can create random variables that we can sample from given each of the hyperparameters.
```
def sample_posterior_from_params(rng, params, posterior, xtest, ds, n_samples=1):
# create a random variable given parameters
predictive_rv = gpx.random_variable(posterior, params, ds)(xtest)
# sample from the rv
return predictive_rv.sample(sample_shape=(n_samples,), seed=rng).squeeze()
# JIT make the sampling faster
sample_posterior = jax.jit(
jax.partial(
sample_posterior_from_params,
posterior=posterior,
xtest=xtest,
ds=training_ds,
n_samples=1,
)
)
```
The JIT operation is optional but it makes it a lot faster...
```
%%time
# sample from the posterior
n_samples = 1_000
seed = 123
mcmc_posterior_samples = jax.vmap(sample_posterior, in_axes=(0, 0))(
jax.random.split(jax.random.PRNGKey(seed), n_samples), posterior_params_samples
)
mcmc_posterior_samples.shape
fig, ax = plt.subplots(figsize=(10, 5))
ax.plot(xtest, mcmc_posterior_samples[:250].T, color="tab:blue", alpha=0.1, lw=1)
ax.plot(x, y.squeeze(), "o", color="tab:orange")
ax.set_title("Posterior samples (MCMC)")
plt.show()
```
## System Information
```
%reload_ext watermark
%watermark -n -u -v -iv -w -a 'J. Emmanuel Johnson'
```
| github_jupyter |
```
import pandas as pd
import os, random, glob, json
from sentence_transformers import SentenceTransformer
import nltk
import numpy as np
from sklearn.svm import LinearSVC
files = glob.glob('/Users/adamtucker/Downloads/tab_feature_files_copy/*.csv')
empty_list = []
final_list = []
header_names = ["sentence_id", "token_id", "token", "lemma", "pos", "term_id", "pol/mod", "poldomain", "aspect_training", "entity", "property", "phrase_type", "sentiment" , ]
for file in files:
#print(file)
frame= pd.read_csv(file,sep="\t", header=None, names=header_names)
list_of_tokens = []
list_of_tokens.append(frame['token'].tolist())
for a in list_of_tokens:
#string_tokens = str(list_of_tokens)
string_per_file = " ".join(a)
#print(string_per_file)
break
list_of_sentiment = []
list_of_sentiment.append(frame['sentiment'].tolist())
#print(list_of_sentiment)
score = 0
for a in list_of_sentiment:
for b in a:
if b == "B-negative":
score += -1
if b == "B-positive":
score += 1
#print(score)
sentiment = ""
if score > 0:
sentiment = "positive"
elif score == 0:
sentiment = "neutral"
else:
sentiment = "negative"
#print(sentiment)
sentiment_dict = dict()
sentiment_dict[string_per_file] = sentiment
empty_list.append(string_per_file)
#print(sentiment_dict)
final_list.append(sentiment_dict)
#print(final_list)
#turn into dataframe
final_list[0]
text_list = []
label_list = []
for d in final_list:
for key, value in d.items():
text_list.append(key)
label_list.append(value)
df['text'] = text_list
df['label'] = label_list
print(len(text_list))
print (len(label_list))
data_text={'text':text_list, 'label': label_list}
df2=pd.DataFrame(data=data_text)
df2
model = SentenceTransformer('bert-base-nli-mean-tokens')
texts = df2.text.to_list()
embedded_texts = []
for text in texts:
sentences = nltk.sent_tokenize(text)
if len(sentences) >=6:
sentence_embeddings = model.encode(sentences[:6])
else:
for n in range (6-len(sentences)):
sentences.append(sentences[0])
embedded_texts.append(sentence_embeddings)
for text in texts:
sentences = nltk.sent_tokenize(text)
if len(sentences) >=6:
sentence_embeddings = model.encode(sentences[:6])
else:
for n in range (6-len(sentences)):
sentences.append(sentences[0])
sentence_embeddings = model.encode(sentences[:6])
embedded_texts.append(sentence_embeddings)
# use concatenated list or mean list in the train_test_split function below
concatenated_list = []
mean_list=[]
for six_embeds in embedded_texts:
new = np.concatenate(six_embeds)
concatenated_list.append(new)
mean_embeds_per_text = np.mean(six_embeds, axis = 0)
mean_list.append(mean_embeds_per_text)
training_labels = list(df2.label)
from sklearn.model_selection import train_test_split
# mean_list can be exchanged for concatenated_list
X_train, X_test, y_train, y_test = train_test_split(mean_list, training_labels, test_size=0.33, random_state=42)
from sklearn.metrics import classification_report
print(classification_report(y_test, predictions))
```
| github_jupyter |
<a href="https://colab.research.google.com/github/jeffheaton/t81_558_deep_learning/blob/master/assignments/assignment_yourname_class7.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# T81-558: Applications of Deep Neural Networks
* Instructor: [Jeff Heaton](https://sites.wustl.edu/jeffheaton/), School of Engineering and Applied Science, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
* For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
**Module 7 Assignment: Computer Vision Neural Network**
**Student Name: Your Name**
# Google CoLab Instructions
This assignment will be most straightforward if you use Google CoLab, because it requires both PyTorch and YOLOv5 to be installed. It will be necessary to mount your GDrive so that you can send your notebook during the submit process. Running the following code will map your GDrive to ```/content/drive```.
```
try:
from google.colab import drive
drive.mount('/content/drive', force_remount=True)
COLAB = True
print("Note: using Google CoLab")
except:
print("Note: not using Google CoLab")
COLAB = False
```
# Assignment Submit Function
You will submit the 10 programming assignments electronically. The following submit function can be used to do this. My server will perform a basic check of each assignment and let you know if it sees any basic problems.
**It is unlikely that should need to modify this function.**
```
import base64
import os
import numpy as np
import pandas as pd
import requests
import PIL
import PIL.Image
import io
# This function submits an assignment. You can submit an assignment as much as you like, only the final
# submission counts. The paramaters are as follows:
# data - List of pandas dataframes or images.
# key - Your student key that was emailed to you.
# no - The assignment class number, should be 1 through 1.
# source_file - The full path to your Python or IPYNB file. This must have "_class1" as part of its name.
# . The number must match your assignment number. For example "_class2" for class assignment #2.
def submit(data,key,no,source_file=None):
if source_file is None and '__file__' not in globals(): raise Exception('Must specify a filename when a Jupyter notebook.')
if source_file is None: source_file = __file__
suffix = '_class{}'.format(no)
if suffix not in source_file: raise Exception('{} must be part of the filename.'.format(suffix))
with open(source_file, "rb") as image_file:
encoded_python = base64.b64encode(image_file.read()).decode('ascii')
ext = os.path.splitext(source_file)[-1].lower()
if ext not in ['.ipynb','.py']: raise Exception("Source file is {} must be .py or .ipynb".format(ext))
payload = []
for item in data:
if type(item) is PIL.Image.Image:
buffered = BytesIO()
item.save(buffered, format="PNG")
payload.append({'PNG':base64.b64encode(buffered.getvalue()).decode('ascii')})
elif type(item) is pd.core.frame.DataFrame:
payload.append({'CSV':base64.b64encode(item.to_csv(index=False).encode('ascii')).decode("ascii")})
r= requests.post("https://api.heatonresearch.com/assignment-submit",
headers={'x-api-key':key}, json={ 'payload': payload,'assignment': no, 'ext':ext, 'py':encoded_python})
if r.status_code==200:
print("Success: {}".format(r.text))
else: print("Failure: {}".format(r.text))
```
# Assignment Instructions
For this assignment, you will use YOLO running on Google CoLab. I suggest that you run this assignment on CoLab because the example code below is already setup to get you started with the correct versions of YOLO on TensorFlow 2.0.
For this assignment you are provided with 10 image files that contain 10 different webcam pictures taken at the [Venice Sidewalk Cafe](https://www.westland.net/beachcam/) a WebCam that has been in opration since 1996. You can find the 10 images here:
* https://data.heatonresearch.com/data/t81-558/sidewalk/sidewalk1.jpg
* https://data.heatonresearch.com/data/t81-558/sidewalk/sidewalk2.jpg
* https://data.heatonresearch.com/data/t81-558/sidewalk/sidewalk3.jpg
* https://data.heatonresearch.com/data/t81-558/sidewalk/sidewalk4.jpg
* https://data.heatonresearch.com/data/t81-558/sidewalk/sidewalk5.jpg
* https://data.heatonresearch.com/data/t81-558/sidewalk/sidewalk6.jpg
* https://data.heatonresearch.com/data/t81-558/sidewalk/sidewalk7.jpg
* https://data.heatonresearch.com/data/t81-558/sidewalk/sidewalk8.jpg
* https://data.heatonresearch.com/data/t81-558/sidewalk/sidewalk9.jpg
* https://data.heatonresearch.com/data/t81-558/sidewalk/sidewalk10.jpg
You can see a sample of the WebCam here:

YOLO does quite well-recognizing objects in this webcam, as the following image illustrates.

You are to write a script that counts the number of certain objects in each of the images. Specifically, you are looking for:
* person
* car
* bicycle
* motorbike
* umbrella
* handbag
It is essential that your use YOLO with a threshold of 10% if you want your results to match mine. The sample code below already contains this setting. Your program can set this threshold with the following command.
* conf_thres=0.1 # confidence threshold (use this value)
* iou_thres=0.25 # NMS IOU threshold (use this value)
Your submitted data frame should also contain a column that identifies which image generated each row. This column should be named **image** and contain integer numbers between 1 and 10. There should be 10 rows in total. The complete data frame should look something like this (not necessarily exactly these numbers).
|image|person|car|bicycle|motorbike|umbrella|handbag|
|-|-|-|-|-|-|-|
|1|23|0|3|4|0|0|
|2|27|1|8|2|0|0|
|3|29|0|0|0|3|0|
|...|...|...|...|...|...|...|
The following code sets up YOLO and then dumps the classification information for the first image. This notebook only serves to get you started. Read in all ten images and generate a data frame that looks like the following. Use the **submit** function as you did in previous assignments.
### Installing YOLOv5
YOLO is not available directly through either PIP or CONDA. Additionally, YOLO is not installed in Google CoLab by default. Therefore, whether you wish to use YOLO through CoLab or run it locally, you need to go through several steps to install it. This section describes the process of installing YOLO. The same steps apply to either CoLab or a local install. For CoLab, you must repeat these steps each time the system restarts your virtual environment. You must perform these steps only once for your virtual Python environment for a local install. If you are installing locally, make sure to install to the same virtual environment you created for this course. The following commands install YOLO directly from its GitHub repository.
```
!git clone https://github.com/ultralytics/yolov5 --tag 6.1
!mv /content/6.1 /content/yolov5
%cd /content/yolov5
%pip install -qr requirements.txt
from yolov5 import utils
display = utils.notebook_init()
```
### Running YOLOv5
In addition to the command line execution we just saw, the YOLO library can easily integrate with Python applications. The following code adds the downloaded YOLOv5 to Python's environment, allowing **yolov5** to be imported like a regular Python library.
```
import sys
sys.path.append(str("/content/yolov5"))
import argparse
import os
from pathlib import Path
import cv2
import torch
import torch.backends.cudnn as cudnn
from models.common import DetectMultiBackend
from utils.datasets import IMG_FORMATS, VID_FORMATS, LoadImages, LoadStreams
from utils.general import (LOGGER, check_file, check_img_size, check_imshow, check_requirements, colorstr,
increment_path, non_max_suppression, print_args, scale_coords, strip_optimizer, xyxy2xywh)
from utils.plots import Annotator, colors, save_one_box
from utils.torch_utils import select_device, time_sync
from PIL import Image
import requests
from io import BytesIO
import torchvision.transforms.functional as TF
```
We are now ready to load YOLO, with pretrained weights provided by the creators of YOLO. It is also possible to train YOLO to recognize images of your own.
```
device = select_device('')
weights = '/content/yolov5/yolov5s.pt'
model = DetectMultiBackend(weights, device=device, dnn=False)
stride, names, pt, jit, onnx, engine = model.stride, model.names, model.pt, model.jit, model.onnx, model.engine
```
I built the following function from the code presented in the course module. The function combines some of the code from the module to accept an image and return what YOLO recognizes. Make sure to use the same two thres_xxx values I provided below to match the results that I got.
```
import numpy as np
conf_thres=0.1 # confidence threshold (use this value)
iou_thres=0.25 # NMS IOU threshold (use this value)
classes = None
agnostic_nms=False, # class-agnostic NMS
max_det=1000
def process_yolo(img):
# Resize image, if needed
imgsz = [img.height, img.width]
imgsz = check_img_size(imgsz, s=stride) # check image size
original_size = imgsz[:]
# Prepare model for this image
model.warmup(imgsz=(1, 3, *imgsz)) # warmup
dt, seen = [0.0, 0.0, 0.0], 0
img2 = img.resize([imgsz[1],imgsz[0]], Image.ANTIALIAS)
# Preprocess image
img_raw = torch.from_numpy(np.asarray(img2)).to(device)
img_raw = img_raw.float() # uint8 to fp16/32
img_raw /= 255 # 0 - 255 to 0.0 - 1.0
img_raw = img_raw.unsqueeze_(0)
img_raw = img_raw.permute(0, 3, 1, 2)
# Query YoLo
pred = model(img_raw, augment=False, visualize=False)
pred = non_max_suppression(pred, conf_thres, iou_thres, classes, agnostic_nms, max_det=max_det)
# convert these raw predictions into the bounding boxes, labels, and
# confidences for each of the images that YOLO recognized.
results = []
for i, det in enumerate(pred): # per image
gn = torch.tensor(img_raw.shape)[[1, 0, 1, 0]]
if len(det):
# Rescale boxes from img_size to im0 size
det[:, :4] = scale_coords(original_size, det[:, :4], imgsz).round()
# Write results
for *xyxy, conf, cls in reversed(det):
xywh = (xyxy2xywh(torch.tensor(xyxy).view(1, 4)) / gn).view(-1).tolist()
# Choose between xyxy and xywh as your desired format.
results.append([names[int(cls)], float(conf), [*xyxy]])
return results
```
### Starter Code
```
url = "https://data.heatonresearch.com/data/t81-558/sidewalk/sidewalk1.jpg"
response = requests.get(url,headers={'User-Agent': 'Mozilla/5.0'})
img = Image.open(BytesIO(response.content))
results = process_yolo(img)
for itm in results:
print(itm)
# Add your solution here, put your results into submit_df
# This is your student key that I emailed to you at the beginnning of the semester.
key = "5iuwhudihwiao6dsfw7dE2ml08iNfVOg6l0O3M06" # This is an example key and will not work.
# You must also identify your source file. (modify for your local setup)
file='/content/drive/MyDrive/Colab Notebooks/assignment_yourname_class7.ipynb' # Google CoLab
submit_df.to_csv("/content/drive/MyDrive/7.csv")
submit(source_file=file,data=[submit_df],key=key,no=7)
```
| github_jupyter |
```
############## PLEASE RUN THIS CELL FIRST! ###################
# import everything and define a test runner function
from importlib import reload
from helper import run
import block, op, helper, script, tx
# op_checkmultisig
def op_checkmultisig(stack, z):
if len(stack) < 1:
return False
n = decode_num(stack.pop())
if len(stack) < n + 1:
return False
sec_pubkeys = []
for _ in range(n):
sec_pubkeys.append(stack.pop())
m = decode_num(stack.pop())
if len(stack) < m + 1:
return False
der_signatures = []
for _ in range(m):
# signature is assumed to be using SIGHASH_ALL
der_signatures.append(stack.pop()[:-1])
# OP_CHECKMULTISIG bug
stack.pop()
try:
raise NotImplementedError
except (ValueError, SyntaxError):
return False
return True
```
### Exercise 1
#### Make [this test](/edit/week5/op.py) pass: `op.py:OpTest:test_op_checkmultisig`
```
# Exercise 1
reload(op)
run(op.OpTest('test_op_checkmultisig'))
```
### Exercise 2
Find the hash160 of the RedeemScript
```
5221022626e955ea6ea6d98850c994f9107b036b1334f18ca8830bfff1295d21cfdb702103b287eaf122eea69030a0e9feed096bed8045c8b98bec453e1ffac7fbdbd4bb7152ae
```
```
# Exercise 2
from helper import hash160
hex_redeem_script = '5221022626e955ea6ea6d98850c994f9107b036b1334f18ca8830bfff1295d21cfdb702103b287eaf122eea69030a0e9feed096bed8045c8b98bec453e1ffac7fbdbd4bb7152ae'
# bytes.fromhex script
# hash160 result
# hex() to display
# P2SH address construction example
from helper import encode_base58_checksum
print(encode_base58_checksum(b'\x05'+bytes.fromhex('74d691da1574e6b3c192ecfb52cc8984ee7b6c56')))
```
### Exercise 3
#### Make [this test](/edit/week5/helper.py) pass: `helper.py:HelperTest:test_p2pkh_address`
```
# Exercise 3
reload(helper)
run(helper.HelperTest('test_p2pkh_address'))
```
### Exercise 4
#### Make [this test](/edit/week5/helper.py) pass: `helper.py:HelperTest:test_p2sh_address`
```
# Exercise 4
reload(helper)
run(helper.HelperTest('test_p2sh_address'))
```
### Exercise 5
#### Make [this test](/edit/week5/script.py) pass: `script.py:ScriptTest:test_address`
```
# Exercise 5
reload(script)
run(script.ScriptTest('test_address'))
# z for p2sh example
from helper import hash256
h256 = hash256(bytes.fromhex('0100000001868278ed6ddfb6c1ed3ad5f8181eb0c7a385aa0836f01d5e4789e6bd304d87221a000000475221022626e955ea6ea6d98850c994f9107b036b1334f18ca8830bfff1295d21cfdb702103b287eaf122eea69030a0e9feed096bed8045c8b98bec453e1ffac7fbdbd4bb7152aeffffffff04d3b11400000000001976a914904a49878c0adfc3aa05de7afad2cc15f483a56a88ac7f400900000000001976a914418327e3f3dda4cf5b9089325a4b95abdfa0334088ac722c0c00000000001976a914ba35042cfe9fc66fd35ac2224eebdafd1028ad2788acdc4ace020000000017a91474d691da1574e6b3c192ecfb52cc8984ee7b6c56870000000001000000'))
z = int.from_bytes(h256, 'big')
print(hex(z))
# p2sh verification example
from ecc import S256Point, Signature
from helper import hash256
h256 = hash256(bytes.fromhex('0100000001868278ed6ddfb6c1ed3ad5f8181eb0c7a385aa0836f01d5e4789e6bd304d87221a000000475221022626e955ea6ea6d98850c994f9107b036b1334f18ca8830bfff1295d21cfdb702103b287eaf122eea69030a0e9feed096bed8045c8b98bec453e1ffac7fbdbd4bb7152aeffffffff04d3b11400000000001976a914904a49878c0adfc3aa05de7afad2cc15f483a56a88ac7f400900000000001976a914418327e3f3dda4cf5b9089325a4b95abdfa0334088ac722c0c00000000001976a914ba35042cfe9fc66fd35ac2224eebdafd1028ad2788acdc4ace020000000017a91474d691da1574e6b3c192ecfb52cc8984ee7b6c56870000000001000000'))
z = int.from_bytes(h256, 'big')
point = S256Point.parse(bytes.fromhex('022626e955ea6ea6d98850c994f9107b036b1334f18ca8830bfff1295d21cfdb70'))
sig = Signature.parse(bytes.fromhex('3045022100dc92655fe37036f47756db8102e0d7d5e28b3beb83a8fef4f5dc0559bddfb94e02205a36d4e4e6c7fcd16658c50783e00c341609977aed3ad00937bf4ee942a89937'))
print(point.verify(z, sig))
```
### Exercise 6
Validate the second signature of the first input
```
0100000001868278ed6ddfb6c1ed3ad5f8181eb0c7a385aa0836f01d5e4789e6bd304d87221a000000db00483045022100dc92655fe37036f47756db8102e0d7d5e28b3beb83a8fef4f5dc0559bddfb94e02205a36d4e4e6c7fcd16658c50783e00c341609977aed3ad00937bf4ee942a8993701483045022100da6bee3c93766232079a01639d07fa869598749729ae323eab8eef53577d611b02207bef15429dcadce2121ea07f233115c6f09034c0be68db99980b9a6c5e75402201475221022626e955ea6ea6d98850c994f9107b036b1334f18ca8830bfff1295d21cfdb702103b287eaf122eea69030a0e9feed096bed8045c8b98bec453e1ffac7fbdbd4bb7152aeffffffff04d3b11400000000001976a914904a49878c0adfc3aa05de7afad2cc15f483a56a88ac7f400900000000001976a914418327e3f3dda4cf5b9089325a4b95abdfa0334088ac722c0c00000000001976a914ba35042cfe9fc66fd35ac2224eebdafd1028ad2788acdc4ace020000000017a91474d691da1574e6b3c192ecfb52cc8984ee7b6c568700000000
```
The sec pubkey of the second signature is:
```
03b287eaf122eea69030a0e9feed096bed8045c8b98bec453e1ffac7fbdbd4bb71
```
The der signature of the second signature is:
```
3045022100da6bee3c93766232079a01639d07fa869598749729ae323eab8eef53577d611b02207bef15429dcadce2121ea07f233115c6f09034c0be68db99980b9a6c5e75402201475221022
```
The redeemScript is:
```
475221022626e955ea6ea6d98850c994f9107b036b1334f18ca8830bfff1295d21cfdb702103b287eaf122eea69030a0e9feed096bed8045c8b98bec453e1ffac7fbdbd4bb7152ae
```
```
# Exercise 6
from io import BytesIO
from ecc import S256Point, Signature
from helper import int_to_little_endian, SIGHASH_ALL
from script import Script
from tx import Tx
hex_sec = '03b287eaf122eea69030a0e9feed096bed8045c8b98bec453e1ffac7fbdbd4bb71'
hex_der = '3045022100da6bee3c93766232079a01639d07fa869598749729ae323eab8eef53577d611b02207bef15429dcadce2121ea07f233115c6f09034c0be68db99980b9a6c5e754022'
hex_redeem_script = '475221022626e955ea6ea6d98850c994f9107b036b1334f18ca8830bfff1295d21cfdb702103b287eaf122eea69030a0e9feed096bed8045c8b98bec453e1ffac7fbdbd4bb7152ae'
sec = bytes.fromhex(hex_sec)
der = bytes.fromhex(hex_der)
redeem_script_stream = BytesIO(bytes.fromhex(hex_redeem_script))
hex_tx = '0100000001868278ed6ddfb6c1ed3ad5f8181eb0c7a385aa0836f01d5e4789e6bd304d87221a000000db00483045022100dc92655fe37036f47756db8102e0d7d5e28b3beb83a8fef4f5dc0559bddfb94e02205a36d4e4e6c7fcd16658c50783e00c341609977aed3ad00937bf4ee942a8993701483045022100da6bee3c93766232079a01639d07fa869598749729ae323eab8eef53577d611b02207bef15429dcadce2121ea07f233115c6f09034c0be68db99980b9a6c5e75402201475221022626e955ea6ea6d98850c994f9107b036b1334f18ca8830bfff1295d21cfdb702103b287eaf122eea69030a0e9feed096bed8045c8b98bec453e1ffac7fbdbd4bb7152aeffffffff04d3b11400000000001976a914904a49878c0adfc3aa05de7afad2cc15f483a56a88ac7f400900000000001976a914418327e3f3dda4cf5b9089325a4b95abdfa0334088ac722c0c00000000001976a914ba35042cfe9fc66fd35ac2224eebdafd1028ad2788acdc4ace020000000017a91474d691da1574e6b3c192ecfb52cc8984ee7b6c568700000000'
stream = BytesIO(bytes.fromhex(hex_tx))
# parse the S256Point and Signature
# parse the Tx
# change the first input's scriptSig to redeemScript
# use Script.parse on the redeem_script_stream
# get the serialization
# add the sighash (4 bytes, little-endian of SIGHASH_ALL)
# hash256 the result
# this interpreted is a big-endian number is your z
# now verify the signature using point.verify
```
### Exercise 7
#### Make [this test](/edit/week5/tx.py) pass: `tx.py:TxTest:test_is_coinbase`
```
# Exercise 7
reload(tx)
run(tx.TxTest('test_is_coinbase'))
```
### Exercise 8
Parse the Genesis Block Coinbase Transaction and print out the scriptSig's third item
```
01000000010000000000000000000000000000000000000000000000000000000000000000ffffffff4d04ffff001d0104455468652054696d65732030332f4a616e2f32303039204368616e63656c6c6f72206f6e206272696e6b206f66207365636f6e64206261696c6f757420666f722062616e6b73ffffffff0100f2052a01000000434104678afdb0fe5548271967f1a67130b7105cd6a828e03909a67962e0ea1f61deb649f6bc3f4cef38c4f35504e51ec112de5c384df7ba0b8d578a4c702b6bf11d5fac00000000
```
```
# Exercise 8
from io import BytesIO
from tx import Tx
hex_tx = '01000000010000000000000000000000000000000000000000000000000000000000000000ffffffff4d04ffff001d0104455468652054696d65732030332f4a616e2f32303039204368616e63656c6c6f72206f6e206272696e6b206f66207365636f6e64206261696c6f757420666f722062616e6b73ffffffff0100f2052a01000000434104678afdb0fe5548271967f1a67130b7105cd6a828e03909a67962e0ea1f61deb649f6bc3f4cef38c4f35504e51ec112de5c384df7ba0b8d578a4c702b6bf11d5fac00000000'
# create stream with BytesIO and bytes.fromhex
# parse the coinbase transaction
# print the first input's script_sig's third command
```
### Exercise 9
#### Make [this test](/edit/week5/tx.py) pass: `tx.py:TxTest:test_coinbase_height`
```
# Exercise 9
reload(tx)
run(tx.TxTest('test_coinbase_height'))
```
### Exercise 10
Find the output address corresponding to this ScriptPubKey
```
1976a914338c84849423992471bffb1a54a8d9b1d69dc28a88ac
```
Remember the structure of pay-to-pubkey-hash (p2pkh) which has `OP_DUP OP_HASH160 <hash> OP_EQUALVERIFY OP_CHECKSIG`.
You need to grab the hash160 and turn that into an address.
```
# Exercise 10
from io import BytesIO
from script import Script
hex_script_pubkey = '1976a914338c84849423992471bffb1a54a8d9b1d69dc28a88ac'
# BytesIO(bytes.fromhex) to get the stream
# parse with Script
# get the address using address() on the script_pubkey
```
### Exercise 11
What is the hash256 of this block? Notice anything?
```
020000208ec39428b17323fa0ddec8e887b4a7c53b8c0a0a220cfd0000000000000000005b0750fce0a889502d40508d39576821155e9c9e3f5c3157f961db38fd8b25be1e77a759e93c0118a4ffd71d
```
```
# Exercise 11
from helper import hash256
hex_block = '020000208ec39428b17323fa0ddec8e887b4a7c53b8c0a0a220cfd0000000000000000005b0750fce0a889502d40508d39576821155e9c9e3f5c3157f961db38fd8b25be1e77a759e93c0118a4ffd71d'
# bytes.fromhex to get the binary
# hash256 the result
# hex() to see what it looks like
```
### Exercise 12
#### Make [this test](/edit/week5/block.py) pass: `block.py:BlockTest:test_parse`
```
# Exercise 12
reload(block)
run(block.BlockTest('test_parse'))
```
### Exercise 13
#### Make [this test](/edit/week5/block.py) pass: `block.py:BlockTest:test_serialize`
```
# Exercise 13
reload(block)
run(block.BlockTest('test_serialize'))
```
### Exercise 14
#### Make [this test](/edit/week5/block.py) pass: `block.py:BlockTest:test_hash`
```
# Exercise 14
reload(block)
run(block.BlockTest('test_hash'))
# Version Signaling Example
from block import Block
from io import BytesIO
hex_block = '020000208ec39428b17323fa0ddec8e887b4a7c53b8c0a0a220cfd0000000000000000005b0750fce0a889502d40508d39576821155e9c9e3f5c3157f961db38fd8b25be1e77a759e93c0118a4ffd71d'
bin_block = bytes.fromhex(hex_block)
stream = BytesIO(bin_block)
b = Block.parse(stream)
version = b.version
print('BIP9: {}'.format(version >> 29 == 0b001))
print('BIP91: {}'.format(version >> 4 & 1 == 1))
print('BIP141: {}'.format(version >> 1 & 1 == 1))
```
### Exercise 15
#### Make [this test](/edit/week5/block.py) pass: `block.py:BlockTest:test_bip9`
```
# Exercise 15
reload(block)
run(block.BlockTest('test_bip9'))
```
### Exercise 16
#### Make [this test](/edit/week5/block.py) pass: `block.py:BlockTest:test_bip91`
```
# Exercise 16
reload(block)
run(block.BlockTest('test_bip91'))
```
### Exercise 17
#### Make [this test](/edit/week5/block.py) pass: `block.py:BlockTest:test_bip141`
```
# Exercise 17
reload(block)
run(block.BlockTest('test_bip141'))
# Calculating Target from Bits Example
from helper import little_endian_to_int
bits = bytes.fromhex('e93c0118')
exponent = bits[-1]
coefficient = little_endian_to_int(bits[:-1])
target = coefficient*256**(exponent-3)
print('{:x}'.format(target).zfill(64))
# Calculating Difficulty from Target Example
from helper import little_endian_to_int
bits = bytes.fromhex('e93c0118')
exponent = bits[-1]
coefficient = little_endian_to_int(bits[:-1])
target = coefficient * 256**(exponent - 3)
min_target = 0xffff * 256**(0x1d - 3)
difficulty = min_target // target
print(difficulty)
```
### Exercise 18
Calculate the target and difficulty for these bits:
```
f2881718
```
Bits to target formula is
\\(\texttt{coefficient}\cdot256^{(\texttt{exponent}-3)}\\)
where coefficient is the first three bytes in little endian and exponent is the last byte.
Target to Difficulty formula is
\\(\texttt{difficulty} = \texttt{min} / \texttt{target}\\)
where \\(\texttt{min} = \texttt{0xffff}\cdot256^{(\texttt{0x1d}-3)}\\)
```
# Exercise 18
hex_bits = 'f2881718'
# bytes.fromhex to get the bits
# last byte is exponent
# first three bytes are the coefficient in little endian
# plug into formula coefficient * 256^(exponent-3) to get the target
# print target using print('{:x}'.format(target).zfill(64))
# difficulty formula is 0xffff * 256**(0x1d - 3) / target
# print the difficulty
```
### Exercise 19
#### Make [this test](/edit/week5/block.py) pass: `block.py:BlockTest:test_target`
```
# Exercise 19
reload(block)
run(block.BlockTest('test_target'))
```
### Exercise 20
Validate the proof-of-work for this block
```
04000000fbedbbf0cfdaf278c094f187f2eb987c86a199da22bbb20400000000000000007b7697b29129648fa08b4bcd13c9d5e60abb973a1efac9c8d573c71c807c56c3d6213557faa80518c3737ec1
```
Check that the proof-of-work (hash256 interpreted as a little-endian number) is lower than the target.
```
# Exercise 20
from block import Block
hex_block = '04000000fbedbbf0cfdaf278c094f187f2eb987c86a199da22bbb20400000000000000007b7697b29129648fa08b4bcd13c9d5e60abb973a1efac9c8d573c71c807c56c3d6213557faa80518c3737ec1'
# bytes.fromhex to get the binary block
# make a stream using BytesIO
# parse the Block
# hash256 the serialization
# interpret the result as a number in little endian
# get the target
# check proof of work < target
```
### Exercise 21
#### Make [this test](/edit/week5/block.py) pass: `block.py:BlockTest:test_check_pow`
```
# Exercise 21
reload(block)
run(block.BlockTest('test_check_pow'))
```
| github_jupyter |
```
import nltk
nltk.download()
from nltk.corpus import brown
brown.categories()
data = brown.sents(categories = 'adventure')
print(data)
len(data)
' '.join(data[4])
from nltk.tokenize import sent_tokenize, word_tokenize
document = """It was a very pleasant day. The weather was cool and there were light showers.
I went to the market to buy some fruits."""
sentence = "Send all the 50 documents related to chapters 1,2,3 at thelokeshgoel00@gmail.com"
sents = sent_tokenize(document)
print(sents)
nltk.download('punkt')
print(len(sents))
sentence.split()
words = word_tokenize(sentence)
print(words,len(words))
from nltk.corpus import stopwords
sw = set(stopwords.words('english'))
def remove_stopwords(text,stopwords):
useful_words = [w for w in text if w not in stopwords]
return useful_words
text = "i am not bothered about her very much".split()
useful_text = remove_stopwords(text, sw)
print(useful_text)
from nltk.tokenize import RegexpTokenizer
tokenizer = RegexpTokenizer('[a-zA-Z@.]+')
useful_text = tokenizer.tokenize(sentence)
print(useful_text)
text = """Foxes love to make jumps. The quick brown fox was seen jumping over the
lovely dog from a 6ft high wall"""
from nltk.stem.snowball import SnowballStemmer, PorterStemmer
from nltk.stem.lancaster import LancasterStemmer
ps = PorterStemmer()
ps.stem('jumping')
ps.stem('jumps')
ps.stem('loving')
ss = SnowballStemmer('english')
ps.stem('Jumving')
```
# Lemmnatization
```
from nltk.stem import WordNetLemmatizer
wn = WordNetLemmatizer()
wn.lemmatize('love')
from sklearn.feature_extraction.text import CountVectorizer
countv = CountVectorizer()
corpus = [
'Indian cricket team will win World Cup, says Capt. Virat Kohli. World cup will be held in Sri Lanka',
'We will win next Lok Sabha Elections, says confident Indian PM',
'The nobel laurste won the hearts of the people.',
'The movie Raazi is an exciting Inddian Spy thriller bassed upon a real story.'
]
vectorized_corpus = countv.fit_transform(corpus)
print(vectorized_corpus.toarray())
vectorized_corpus = vectorized_corpus.toarray()
countv.vocabulary_
len(vectorized_corpus[0])
len(countv.vocabulary_.keys())
countv.inverse_transform(vectorized_corpus[2])
CountVectorizer['tokenizer']?
def myTokenizer(document):
words = tokenizer.tokenize(document.lower())
words = remove_stopwords(words,sw)
return words
myTokenizer(sentence)
cv = CountVectorizer(tokenizer = myTokenizer)
vectorized_corpus = cv.fit_transform(corpus).toarray()
print(vectorized_corpus)
len(vectorized_corpus[0])
cv.inverse_transform(vectorized_corpus)
test_corpus = [
'Indian cricket team rock!'
]
cv.transform(test_corpus).toarray()
cv.vocabulary_
cv
cv.transform?
cv = CountVectorizer(ngram_range=(2,2))
cv.fit_transform(sentence).toarray()
from sklearn.feature_extraction.text import TfidfVectorizer
tfidf = TfidfVectorizer()
vc = tfidf.fit_transform(corpus).toarray()
vc
tfidf.vocabulary_
corpus = ["this is good movie", "This was good movie", "this is not good movie"]
```
| github_jupyter |
# Data generators
In Python, a generator is a function that behaves like an iterator. It will return the next item. Here is a [link](https://wiki.python.org/moin/Generators) to review python generators. In many AI applications, it is advantageous to have a data generator to handle loading and transforming data for different applications.
You will now implement a custom data generator, using a common pattern that you will use during all assignments of this course.
In the following example, we use a set of samples `a`, to derive a new set of samples, with more elements than the original set.
**Note: Pay attention to the use of list `lines_index` and variable `index` to traverse the original list.**
```
import random
import numpy as np
# Example of traversing a list of indexes to create a circular list
a = [1, 2, 3, 4]
b = [0] * 10
a_size = len(a)
b_size = len(b)
lines_index = [*range(a_size)] # is equivalent to [i for i in range(0,a_size)], the difference being the advantage of using * to pass values of range iterator to list directly
index = 0 # similar to index in data_generator below
for i in range(b_size): # `b` is longer than `a` forcing a wrap
# We wrap by resetting index to 0 so the sequences circle back at the end to point to the first index
if index >= a_size:
index = 0
b[i] = a[lines_index[index]] # `indexes_list[index]` point to a index of a. Store the result in b
index += 1
print(b)
```
## Shuffling the data order
In the next example, we will do the same as before, but shuffling the order of the elements in the output list. Note that here, our strategy of traversing using `lines_index` and `index` becomes very important, because we can simulate a shuffle in the input data, without doing that in reality.
```
# Example of traversing a list of indexes to create a circular list
a = [1, 2, 3, 4]
b = []
a_size = len(a)
b_size = 10
lines_index = [*range(a_size)]
print("Original order of index:",lines_index)
# if we shuffle the index_list we can change the order of our circular list
# without modifying the order or our original data
random.shuffle(lines_index) # Shuffle the order
print("Shuffled order of index:",lines_index)
print("New value order for first batch:",[a[index] for index in lines_index])
batch_counter = 1
index = 0 # similar to index in data_generator below
for i in range(b_size): # `b` is longer than `a` forcing a wrap
# We wrap by resetting index to 0
if index >= a_size:
index = 0
batch_counter += 1
random.shuffle(lines_index) # Re-shuffle the order
print("\nShuffled Indexes for Batch No.{} :{}".format(batch_counter,lines_index))
print("Values for Batch No.{} :{}".format(batch_counter,[a[index] for index in lines_index]))
b.append(a[lines_index[index]]) # `indexes_list[index]` point to a index of a. Store the result in b
index += 1
print()
print("Final value of b:",b)
```
**Note: We call an epoch each time that an algorithm passes over all the training examples. Shuffling the examples for each epoch is known to reduce variance, making the models more general and overfit less.**
### Exercise
**Instructions:** Implement a data generator function that takes in `batch_size, x, y shuffle` where x could be a large list of samples, and y is a list of the tags associated with those samples. Return a subset of those inputs in a tuple of two arrays `(X,Y)`. Each is an array of dimension (`batch_size`). If `shuffle=True`, the data will be traversed in a random form.
**Details:**
This code as an outer loop
```
while True:
...
yield((X,Y))
```
Which runs continuously in the fashion of generators, pausing when yielding the next values. We will generate a batch_size output on each pass of this loop.
It has an inner loop that stores in temporal lists (X, Y) the data samples to be included in the next batch.
There are three slightly out of the ordinary features.
1. The first is the use of a list of a predefined size to store the data for each batch. Using a predefined size list reduces the computation time if the elements in the array are of a fixed size, like numbers. If the elements are of different sizes, it is better to use an empty array and append one element at a time during the loop.
2. The second is tracking the current location in the incoming lists of samples. Generators variables hold their values between invocations, so we create an `index` variable, initialize to zero, and increment by one for each sample included in a batch. However, we do not use the `index` to access the positions of the list of sentences directly. Instead, we use it to select one index from a list of indexes. In this way, we can change the order in which we traverse our original list, keeping untouched our original list.
3. The third also relates to wrapping. Because `batch_size` and the length of the input lists are not aligned, gathering a batch_size group of inputs may involve wrapping back to the beginning of the input loop. In our approach, it is just enough to reset the `index` to 0. We can re-shuffle the list of indexes to produce different batches each time.
```
def data_generator(batch_size, data_x, data_y, shuffle=True):
'''
Input:
batch_size - integer describing the batch size
data_x - list containing samples
data_y - list containing labels
shuffle - Shuffle the data order
Output:
a tuple containing 2 elements:
X - list of dim (batch_size) of samples
Y - list of dim (batch_size) of labels
'''
data_lng = len(data_x) # len(data_x) must be equal to len(data_y)
index_list = [*range(data_lng)] # Create a list with the ordered indexes of sample data
# If shuffle is set to true, we traverse the list in a random way
if shuffle:
random.shuffle(index_list) # Inplace shuffle of the list
index = 0 # Start with the first element
# START CODE HERE
# Fill all the None values with code taking reference of what you learned so far
while True:
X = [0] * batch_size # We can create a list with batch_size elements.
Y = [0] * batch_size # We can create a list with batch_size elements.
for i in range(batch_size):
# Wrap the index each time that we reach the end of the list
if index >= data_lng:
index = 0
# Shuffle the index_list if shuffle is true
if shuffle:
rnd.shuffle(index_list) # re-shuffle the order
X[i] = data_x[index_list[index]] # We set the corresponding element in x
Y[i] = data_y[index_list[index]] # We set the corresponding element in x
# END CODE HERE
index += 1
yield((X, Y))
```
If your function is correct, all the tests must pass.
```
def test_data_generator():
x = [1, 2, 3, 4]
y = [xi ** 2 for xi in x]
generator = data_generator(3, x, y, shuffle=False)
assert np.allclose(next(generator), ([1, 2, 3], [1, 4, 9])), "First batch does not match"
assert np.allclose(next(generator), ([4, 1, 2], [16, 1, 4])), "Second batch does not match"
assert np.allclose(next(generator), ([3, 4, 1], [9, 16, 1])), "Third batch does not match"
assert np.allclose(next(generator), ([2, 3, 4], [4, 9, 16])), "Fourth batch does not match"
print("\033[92mAll tests passed!")
test_data_generator()
```
If you could not solve the exercise, just run the next code to see the answer.
```
import base64
solution = "ZGVmIGRhdGFfZ2VuZXJhdG9yKGJhdGNoX3NpemUsIGRhdGFfeCwgZGF0YV95LCBzaHVmZmxlPVRydWUpOgoKICAgIGRhdGFfbG5nID0gbGVuKGRhdGFfeCkgIyBsZW4oZGF0YV94KSBtdXN0IGJlIGVxdWFsIHRvIGxlbihkYXRhX3kpCiAgICBpbmRleF9saXN0ID0gWypyYW5nZShkYXRhX2xuZyldICMgQ3JlYXRlIGEgbGlzdCB3aXRoIHRoZSBvcmRlcmVkIGluZGV4ZXMgb2Ygc2FtcGxlIGRhdGEKICAgIAogICAgIyBJZiBzaHVmZmxlIGlzIHNldCB0byB0cnVlLCB3ZSB0cmF2ZXJzZSB0aGUgbGlzdCBpbiBhIHJhbmRvbSB3YXkKICAgIGlmIHNodWZmbGU6CiAgICAgICAgcm5kLnNodWZmbGUoaW5kZXhfbGlzdCkgIyBJbnBsYWNlIHNodWZmbGUgb2YgdGhlIGxpc3QKICAgIAogICAgaW5kZXggPSAwICMgU3RhcnQgd2l0aCB0aGUgZmlyc3QgZWxlbWVudAogICAgd2hpbGUgVHJ1ZToKICAgICAgICBYID0gWzBdICogYmF0Y2hfc2l6ZSAjIFdlIGNhbiBjcmVhdGUgYSBsaXN0IHdpdGggYmF0Y2hfc2l6ZSBlbGVtZW50cy4gCiAgICAgICAgWSA9IFswXSAqIGJhdGNoX3NpemUgIyBXZSBjYW4gY3JlYXRlIGEgbGlzdCB3aXRoIGJhdGNoX3NpemUgZWxlbWVudHMuIAogICAgICAgIAogICAgICAgIGZvciBpIGluIHJhbmdlKGJhdGNoX3NpemUpOgogICAgICAgICAgICAKICAgICAgICAgICAgIyBXcmFwIHRoZSBpbmRleCBlYWNoIHRpbWUgdGhhdCB3ZSByZWFjaCB0aGUgZW5kIG9mIHRoZSBsaXN0CiAgICAgICAgICAgIGlmIGluZGV4ID49IGRhdGFfbG5nOgogICAgICAgICAgICAgICAgaW5kZXggPSAwCiAgICAgICAgICAgICAgICAjIFNodWZmbGUgdGhlIGluZGV4X2xpc3QgaWYgc2h1ZmZsZSBpcyB0cnVlCiAgICAgICAgICAgICAgICBpZiBzaHVmZmxlOgogICAgICAgICAgICAgICAgICAgIHJuZC5zaHVmZmxlKGluZGV4X2xpc3QpICMgcmUtc2h1ZmZsZSB0aGUgb3JkZXIKICAgICAgICAgICAgCiAgICAgICAgICAgIFhbaV0gPSBkYXRhX3hbaW5kZXhfbGlzdFtpbmRleF1dIAogICAgICAgICAgICBZW2ldID0gZGF0YV95W2luZGV4X2xpc3RbaW5kZXhdXSAKICAgICAgICAgICAgCiAgICAgICAgICAgIGluZGV4ICs9IDEKICAgICAgICAKICAgICAgICB5aWVsZCgoWCwgWSkp"
# Print the solution to the given assignment
print(base64.b64decode(solution).decode("utf-8"))
```
### Hope you enjoyed this tutorial on data generators which will help you with the assignments in this course.
| github_jupyter |
# PLEASE USE `Cell => Run All` ON THIS NOTEBOOK (MENU OPTION ABOVE).
# THIS NOTEBOOK WILL TAKE 20-30 MINUTES TO COMPLETE SUCCESSFULLY.
# PLEASE BE PATIENT
```
import sys
print('Python Version %s' % sys.version)
!pip list
%%bash
conda list
```
# 1. Install the CLI tools
_Note: Make sure the SageMaker role has access to create EKS Cluster._
```
%%bash
source ~/.bash_profile
pip install awscli --upgrade --user
curl --silent --location "https://github.com/weaveworks/eksctl/releases/latest/download/eksctl_$(uname -s)_amd64.tar.gz" | tar xz -C /tmp
sudo mv /tmp/eksctl /usr/local/bin
eksctl version
curl --location -o ./kubectl https://amazon-eks.s3.us-west-2.amazonaws.com/1.18.8/2020-09-18/bin/linux/amd64/kubectl
#curl --location -o ./kubectl https://amazon-eks.s3.us-west-2.amazonaws.com/1.15.10/2020-02-22/bin/linux/amd64/kubectl
chmod +x ./kubectl
sudo mv ./kubectl /usr/local/bin
kubectl version --short --client
curl -o aws-iam-authenticator https://amazon-eks.s3.us-west-2.amazonaws.com/1.18.8/2020-09-18/bin/linux/amd64/aws-iam-authenticator
#curl -o aws-iam-authenticator https://amazon-eks.s3.us-west-2.amazonaws.com/1.15.10/2020-02-22/bin/linux/amd64/aws-iam-authenticator
chmod +x ./aws-iam-authenticator
sudo mv aws-iam-authenticator /usr/local/bin
aws-iam-authenticator version
sudo yum -y install jq gettext
for command in kubectl jq envsubst
do
which $command &>/dev/null && echo "$command in path" || echo "$command NOT FOUND"
done
echo "Completed"
```
# 2. Configure the Environment Variables
```
%%bash
source ~/.bash_profile
export AWS_REGION=$(aws configure get region)
echo "export AWS_REGION=${AWS_REGION}" | tee -a ~/.bash_profile
export AWS_CLUSTER_NAME=workshop
echo "export AWS_CLUSTER_NAME=${AWS_CLUSTER_NAME}" | tee -a ~/.bash_profile
echo "Completed"
```
# 3. Create the EKS Cluster
# THIS WILL TAKE 20-30 MINUTES. PLEASE BE PATIENT!
# _If you see `retryable error`s below, this is OK!_
```
%%bash
###############################
#
# ONLY RUN THIS CELL ONE TIME!!
#
###############################
source ~/.bash_profile
export AWS_CLUSTER_STATUS=$(aws eks describe-cluster --name ${AWS_CLUSTER_NAME} --region ${AWS_REGION} --query "cluster.status" --output text)
if [ -n "$AWS_CLUSTER_STATUS" ]; then
if [ $AWS_CLUSTER_STATUS == "ACTIVE" ]; then
echo "Cluster status: $AWS_CLUSTER_STATUS."
else
echo "Cluster status: $AWS_CLUSTER_STATUS. Please wait for status: ACTIVE"
exit
fi
else
# THIS WILL TAKE 20-30 MINUTES. PLEASE BE PATIENT.
eksctl create cluster \
--name ${AWS_CLUSTER_NAME} \
--version 1.18 \
--nodes 4 \
--node-type c5.xlarge \
--node-volume-size 100 \
--region=${AWS_REGION} \
--zones=${AWS_REGION}a,${AWS_REGION}b \
--alb-ingress-access
export AWS_CLUSTER_STATUS=$(aws eks describe-cluster --name ${AWS_CLUSTER_NAME} --region ${AWS_REGION} --query "cluster.status" --output text)
echo "export AWS_CLUSTER_STATUS=${AWS_CLUSTER_STATUS}" | tee -a ~/.bash_profile
echo "Cluster status: $AWS_CLUSTER_STATUS"
fi
```
# _Your output will look like this..._

# _If you see `retryable error`s ^^ above ^^, this is OK!_
# AFTER 20-30 MINUTES, YOUR CLUSTER WILL BE CREATED.
```
%%bash
source ~/.bash_profile
export AWS_CLUSTER_STATUS=$(aws eks describe-cluster --name ${AWS_CLUSTER_NAME} --region ${AWS_REGION} --query "cluster.status" --output text)
if [ -n "$AWS_CLUSTER_STATUS" ]; then
if [ $AWS_CLUSTER_STATUS == "ACTIVE" ]; then
echo "Cluster status: $AWS_CLUSTER_STATUS. Please continue."
else
echo "Cluster status: $AWS_CLUSTER_STATUS. Please wait for status: ACTIVE"
exit
fi
fi
```
| github_jupyter |
Today's mini-lab is an excercise in abstraction, in addition to parameter tuning and visualisation. The models below come from scientific computing and epidemiology, and are implemented as systems of differential equations.
*Without a thorough background in mathematics, you will not be able to understand the details. This is a good thing since we can then focus on using the model instead of digging in to the details.*
# When zombies attack!
The excellent paper "*When zombies attack!: Mathematical modelling of an outbreak of zombie infection*" explains the dynamic disiase spread using a zombie apocalypse as the case study. We will use this paper to illustrate a model that is interpretable yet still requires more mathematics than required for the course to undersdtand in detail. During the lab, you will get aquainted with the models from the paper and make modifications to either fit the model to some contraints from data or try different zombie invasion mitigation strategies.
The setting will be the town of Uppsala in Sweden (for obvious reasons). The population of the town is ca 165000 (not the full muicipality, we're only interested in a contigous urban area). In the equations below, the formulation is slightly modified so that the sum of the compartments is the total population of Uppsala.
```
N = 165000 # Population of Uppsala
from scipy.integrate import odeint # For working with systems of differential equations
import numpy as np # Math
import matplotlib.pyplot as plt # Plotting
import ipywidgets # Interactivity
```
## The basic model
The following system of equations is from part 2 in the paper. Zombias start waking up, one per late evening (as they do), and start roaming the town in search of fresh brains.
$\frac{dS}{dt} = − \frac{\beta Z}{N}S$
$\frac{dZ}{dt} = \frac{\beta Z}{N}S − \frac{\alpha Z}{N}S + min(1, R) - \delta Z$
$\frac{dR}{dt} = \frac{\alpha Z}{N}S − min(1, R) + \delta Z$
#### Constants
$\beta$: Compund zombie infection rate
$\alpha$: Compund zombie eradication rate
$\delta$: Inverse of the mean number of days zombies survive (hard coded as 2 weeks)
We will assume that the timescale is fairly short and set $\Pi=\delta=0$, as in the paper. The system of differential equations above already reflect this simplification.
```
beta = .25 # Suseptable -> Zombie
alpha = .1 # Zombie -> Removed
def szr_model(y0, beta, alpha, runtime=28*7):
"""A SZR model
runtime: int, runtime in days"""
def szr_dydt(y, t, beta, alpha):
S, Z, R = y
delta = 1/14 # Zombies live for 2 weeks
dydt = [- beta*S*Z/N,
beta*S*Z/N + min(1, R) - alpha*S*Z/N - delta*Z,
alpha*S*Z/N - min(1, R) + delta*Z]
return dydt
t = np.arange(runtime)
res, debug_msg = odeint(szr_dydt, y0, t, args=(beta, alpha), full_output=True)
return t, np.asarray(res).T, debug_msg
t, res, _ = szr_model(y0=np.asarray([N, 0, 10]),
beta=beta,
alpha=alpha)
plt.figure(figsize=(8, 4), dpi=100)
plt.plot(t, res[0, :], 'g-', label="Suseptable")
plt.plot(t, res[1, :], 'r-', label="Zombie")
plt.plot(t, res[2, :], 'k-', label="Removed")
plt.legend()
plt.ylabel("Individuals")
plt.xlabel("Time [days]")
plt.show()
```
You can try different value using the following widget.
```
def szr_widget(beta, alpha):
t, res, _ = szr_model(y0=np.asarray([N, 0, 10]),
beta=beta,
alpha=alpha)
plt.figure(figsize=(8, 4), dpi=100)
plt.plot(t, res[0, :], 'g-', label="Suseptable")
plt.plot(t, res[1, :], 'r-', label="Zombie")
plt.plot(t, res[2, :], 'k-', label="Removed")
plt.legend()
plt.ylabel("Individuals")
plt.xlabel("Time [days]")
plt.show()
ipywidgets.interact(szr_widget,
beta=ipywidgets.FloatSlider(min=0, max=.5, step=0.01, value=.1, readout_format='.2f', description="S->Z (beta, prob. of infection)", style = {'description_width': 'initial'}, layout=ipywidgets.Layout(width='600px')),
alpha=ipywidgets.FloatSlider(min=0, max=.5, step=0.01, value=.1, readout_format='.2f', description="Z->R (alpha, prob. of eradication)", style = {'description_width': 'initial'}, layout=ipywidgets.Layout(width='600px')));
```
What is the relation between the values of $\beta$ and $\alpha$? Run a grid search over them an plot using matplotlibs contour (or similar). All the code is already in the notebooks you have worked with.
```
```
## The Model with Latent Infection
$S' = -\beta SZ$
$E' = \beta SZ - \rho E$
$Z' = \rho E − \alpha SZ - \delta Z$
$R' = \alpha SZ + \delta Z$
#### Constants
$\beta$: Compund zombie infection rate
$\alpha$: Compund zombie erradication rate
$\delta$: Inverse of the mean number of days zombies survive
$\rho$: Inverse of the mean number of days from exposure to zombification
Again, ee will assume that the timescale is fairly short and set $\Pi=\delta=0$, as in the paper. We will also assume that zombies rise one per night. The new parameter $\rho$ controls the time from exposure to full sickness, i.e. incubation time. This parameter should be set to the inverse of the number of days before full infection, e.g. for a mena of 3 days $\rho=\frac{1}{3}$.
```
beta = .25 # Suseptable -> Zombie
alpha = .1 # Zombie -> Removed
rho_inv = 1 # Mean incubation days
delta_inv = 14 # Mean zombie lifespan
def sezr_model(y0, beta, alpha, rho_inv, delta_inv, runtime=28*7):
def sezr_dydt(y, t, beta, alpha, rho_inv, delta_inv):
S, E, Z, R = y
dydt = [-beta*S*Z,
beta*S*Z - E/rho_inv,
E/rho_inv - alpha*S*Z - Z/delta_inv,
alpha*S*Z + Z/delta_inv]
return dydt
t = np.arange(runtime)
res, debug_msg = odeint(sezr_dydt, y0, t, args=(beta, alpha, rho_inv, delta_inv), full_output=True)
return t, np.asarray(res).T, debug_msg
t, res, _ = sezr_model(y0=np.asarray([N, 1, 0, 0]),
beta=beta,
alpha=alpha,
delta_inv=delta_inv,
rho_inv=rho_inv)
plt.figure(figsize=(8, 4), dpi=100)
plt.plot(t, res[0, :], 'g-', label="Suseptable")
plt.plot(t, res[1, :], color='orange', label="Exposed")
plt.plot(t, res[2, :], 'r-', label="Zombie")
plt.plot(t, res[3, :], 'k-', label="Removed")
plt.legend()
plt.ylabel("Individuals")
plt.xlabel("Time [days]")
plt.show()
def sezr_widget(beta, alpha, delta_inv, rho_inv):
t, res, _ = sezr_model(y0=np.asarray([N, 0, 0, 10]),
beta=beta,
alpha=alpha,
delta_inv=delta_inv,
rho_inv=rho_inv)
plt.figure(figsize=(8, 4), dpi=100)
plt.plot(t, res[0, :], 'g-', label="Suseptable")
plt.plot(t, res[1, :], color='orange', label="Exposed")
plt.plot(t, res[2, :], 'r-', label="Zombie")
plt.plot(t, res[3, :], 'k-', label="Removed")
plt.legend()
plt.ylabel("Individuals")
plt.xlabel("Time [days]")
plt.show()
ipywidgets.interact(sezr_widget,
beta=ipywidgets.FloatSlider(min=0, max=.5, step=0.01, value=.1, readout_format='.2f', description="S->Z (beta, prob. of infection)", style = {'description_width': 'initial'}, layout=ipywidgets.Layout(width='600px')),
alpha=ipywidgets.FloatSlider(min=0, max=.5, step=0.01, value=.1, readout_format='.2f', description="Z->R (alpha, prob. of eradication)", style = {'description_width': 'initial'}, layout=ipywidgets.Layout(width='600px')),
rho_inv=ipywidgets.FloatSlider(min=0, max=21, step=0.5, value=3, readout_format='.2f', description="E->Z (mean incubation time in days)", style = {'description_width': 'initial'}, layout=ipywidgets.Layout(width='600px')),
delta_inv=ipywidgets.FloatSlider(min=0, max=21, step=0.5, value=3, readout_format='.2f', description="Z->R (Zombie mean life span in days)", style = {'description_width': 'initial'}, layout=ipywidgets.Layout(width='600px'))
);
```
## A Model with vaccine
```
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
%aimport utils_1_1
import us
import pandas as pd
import numpy as np
import altair as alt
from altair_saver import save
from vega_datasets import data
import datetime
import dateutil.parser
from os.path import join
from constants_1_1 import SITE_FILE_TYPES
from utils_1_1 import (
get_site_file_paths,
get_site_file_info,
get_site_ids,
get_visualization_subtitle,
get_country_color_map,
apply_theme,
)
from web import for_website
alt.data_transformers.disable_max_rows(); # Allow using rows more than 5000
df = pd.read_csv(data.population_engineers_hurricanes.url)
state_to_id = dict(zip(df["state"].values.tolist(), df["id"].values.tolist()))
states = alt.topo_feature(data.us_10m.url, 'states')
del df
def convert_date_us(date_str):
try:
return dateutil.parser.parse(date_str)
except:
return np.nan
def convert_state(abbr):
return int(us.states.lookup(abbr).fips)
hhs_df = pd.read_csv(join("..", "data", "reported_hospital_capacity_admissions_facility-level_weekly_average_timeseries_20201207.csv"))
hhs_df.head()
hhs_df["id"] = hhs_df["state"].apply(convert_state)
hhs_col = "previous_day_admission_pediatric_covid_confirmed_7_day_sum"
hhs_df = hhs_df[["hospital_pk", "state", "id", "collection_week", "hospital_subtype", "is_metro_micro", hhs_col]]
hhs_df = hhs_df.rename(columns={"collection_week": "date", hhs_col: "ped_count"})
hhs_df["ped_count"] = hhs_df["ped_count"].clip(lower=0)
orig_hhs_df = hhs_df.copy()
hhs_df = hhs_df.groupby(by=["date", "hospital_pk"]).sum().reset_index()
hhs_df["has_any"] = hhs_df["ped_count"].apply(lambda x: 1)
hhs_df["has_ped"] = hhs_df["ped_count"].apply(lambda x: pd.notna(x) and x > 0.0)
hhs_df.head()
hhs_df = hhs_df.groupby("date").sum().reset_index()
hhs_df["date"] = hhs_df["date"].apply(convert_date_us)
hhs_df.head()
plot = alt.Chart(hhs_df).mark_line().encode(
y=alt.Y("has_ped:Q"),
x=alt.X("date"),
).properties(
title=f"Number of hospitals reporting > 0 value for {hhs_col} column"
)
plot
hhs_df["pct_with_ped"] = hhs_df["has_ped"] / hhs_df["has_any"] * 100
plot = alt.Chart(hhs_df).mark_line().encode(
y=alt.Y("pct_with_ped:Q"),
x=alt.X("date"),
).properties(
title=f"Percentage of hospitals reporting > 0 value for {hhs_col} column"
)
plot
hhs_df = orig_hhs_df.copy()
hhs_df = hhs_df.groupby(by=["id", "hospital_pk"]).sum().reset_index()
hhs_df["has_any"] = hhs_df["ped_count"].apply(lambda x: 1)
hhs_df["has_ped"] = hhs_df["ped_count"].apply(lambda x: pd.notna(x) and x > 0.0)
hhs_df.head()
# group by state ID
hhs_df = hhs_df.groupby("id").sum().reset_index()
hhs_df.head()
plot = alt.Chart(states).mark_geoshape().encode(
color=alt.Color("has_ped:Q")
).transform_lookup(
lookup='id',
from_=alt.LookupData(data=hhs_df, key='id', fields=['has_ped'])
).properties(
width=500,
height=300,
title=f"Number of hospitals reporting > 0 value for {hhs_col} column (on at least one date)"
).project(
type='albersUsa'
)
plot
hhs_df["has_zero_ped"] = hhs_df["has_ped"].apply(lambda x: "Yes" if x == 0 else "No")
plot = alt.Chart(states).mark_geoshape().encode(
color=alt.Color("has_zero_ped:N")
).transform_lookup(
lookup='id',
from_=alt.LookupData(data=hhs_df, key='id', fields=['has_zero_ped'])
).properties(
width=500,
height=300,
title=f"States reporting zero or missing for {hhs_col} column (on every date)"
).project(
type='albersUsa'
)
plot
hhs_df = orig_hhs_df.copy()
hhs_df.head()
hhs_df = hhs_df.groupby(by=["state", "hospital_pk", "hospital_subtype"]).sum().reset_index()
hhs_df["has_any"] = hhs_df["ped_count"].apply(lambda x: 1)
hhs_df["has_ped"] = hhs_df["ped_count"].apply(lambda x: pd.notna(x) and x > 0.0)
hhs_df.head()
hhs_df = hhs_df.groupby(by=["state", "hospital_subtype"]).sum().reset_index()
hhs_df.head()
plot = alt.Chart(hhs_df).mark_bar().encode(
y=alt.Y("has_any:Q"),
x=alt.X("hospital_subtype:N"),
color=alt.Color("hospital_subtype:N"),
facet=alt.Facet('state:O', columns=14),
).properties(
height=140,
title=f"Hospital types by state (adult and pediatric)"
)
plot
plot = alt.Chart(hhs_df).mark_bar().encode(
y=alt.Y("has_ped:Q"),
x=alt.X("hospital_subtype:N"),
color=alt.Color("hospital_subtype:N"),
facet=alt.Facet('state:O', columns=14),
).properties(
height=140,
title=f"Hospital types by state reporting > 0 value for {hhs_col} column (on at least one date) "
)
plot
hhs_df = orig_hhs_df.copy()
hhs_df.head()
hhs_df = hhs_df.groupby(by=["state", "hospital_pk", "is_metro_micro"]).sum().reset_index()
hhs_df["has_any"] = hhs_df["ped_count"].apply(lambda x: 1)
hhs_df["has_ped"] = hhs_df["ped_count"].apply(lambda x: pd.notna(x) and x > 0.0)
hhs_df.head()
hhs_df = hhs_df.groupby(by=["state", "is_metro_micro"]).sum().reset_index()
hhs_df["is_metro_micro"] = hhs_df["is_metro_micro"].apply(lambda x: "Yes" if x else "No")
hhs_df.head()
plot = alt.Chart(hhs_df).mark_bar().encode(
y=alt.Y("has_ped:Q"),
x=alt.X("is_metro_micro:N"),
color=alt.Color("is_metro_micro:N"),
facet=alt.Facet('state:O', columns=14),
).properties(
height=100,
width=60,
title=f"Population served by state reporting > 0 value for {hhs_col} column (on at least one date) "
)
plot
```
| github_jupyter |
# Planning: planning.py; chapters 10-11
This notebook describes the [planning.py](https://github.com/aimacode/aima-python/blob/master/planning.py) module, which covers Chapters 10 (Classical Planning) and 11 (Planning and Acting in the Real World) of *[Artificial Intelligence: A Modern Approach](http://aima.cs.berkeley.edu)*. See the [intro notebook](https://github.com/aimacode/aima-python/blob/master/intro.ipynb) for instructions.
We'll start by looking at `PDDL` and `Action` data types for defining problems and actions. Then, we will see how to use them by trying to plan a trip from *Sibiu* to *Bucharest* across the familiar map of Romania, from [search.ipynb](https://github.com/aimacode/aima-python/blob/master/search.ipynb). Finally, we will look at the implementation of the GraphPlan algorithm.
The first step is to load the code:
```
from planning import *
```
To be able to model a planning problem properly, it is essential to be able to represent an Action. Each action we model requires at least three things:
* preconditions that the action must meet
* the effects of executing the action
* some expression that represents the action
Planning actions have been modelled using the `Action` class. Let's look at the source to see how the internal details of an action are implemented in Python.
```
%psource Action
```
It is interesting to see the way preconditions and effects are represented here. Instead of just being a list of expressions each, they consist of two lists - `precond_pos` and `precond_neg`. This is to work around the fact that PDDL doesn't allow for negations. Thus, for each precondition, we maintain a separate list of those preconditions that must hold true, and those whose negations must hold true. Similarly, instead of having a single list of expressions that are the result of executing an action, we have two. The first (`effect_add`) contains all the expressions that will evaluate to true if the action is executed, and the the second (`effect_neg`) contains all those expressions that would be false if the action is executed (ie. their negations would be true).
The constructor parameters, however combine the two precondition lists into a single `precond` parameter, and the effect lists into a single `effect` parameter.
The `PDDL` class is used to represent planning problems in this module. The following attributes are essential to be able to define a problem:
* a goal test
* an initial state
* a set of viable actions that can be executed in the search space of the problem
View the source to see how the Python code tries to realise these.
```
%psource PDDL
```
The `initial_state` attribute is a list of `Expr` expressions that forms the initial knowledge base for the problem. Next, `actions` contains a list of `Action` objects that may be executed in the search space of the problem. Lastly, we pass a `goal_test` function as a parameter - this typically takes a knowledge base as a parameter, and returns whether or not the goal has been reached.
Now lets try to define a planning problem using these tools. Since we already know about the map of Romania, lets see if we can plan a trip across a simplified map of Romania.
Here is our simplified map definition:
```
from utils import *
# this imports the required expr so we can create our knowledge base
knowledge_base = [
expr("Connected(Bucharest,Pitesti)"),
expr("Connected(Pitesti,Rimnicu)"),
expr("Connected(Rimnicu,Sibiu)"),
expr("Connected(Sibiu,Fagaras)"),
expr("Connected(Fagaras,Bucharest)"),
expr("Connected(Pitesti,Craiova)"),
expr("Connected(Craiova,Rimnicu)")
]
```
Let us add some logic propositions to complete our knowledge about travelling around the map. These are the typical symmetry and transitivity properties of connections on a map. We can now be sure that our `knowledge_base` understands what it truly means for two locations to be connected in the sense usually meant by humans when we use the term.
Let's also add our starting location - *Sibiu* to the map.
```
knowledge_base.extend([
expr("Connected(x,y) ==> Connected(y,x)"),
expr("Connected(x,y) & Connected(y,z) ==> Connected(x,z)"),
expr("At(Sibiu)")
])
```
We now have a complete knowledge base, which can be seen like this:
```
knowledge_base
```
We now define possible actions to our problem. We know that we can drive between any connected places. But, as is evident from [this](https://en.wikipedia.org/wiki/List_of_airports_in_Romania) list of Romanian airports, we can also fly directly between Sibiu, Bucharest, and Craiova.
We can define these flight actions like this:
```
#Sibiu to Bucharest
precond_pos = [expr('At(Sibiu)')]
precond_neg = []
effect_add = [expr('At(Bucharest)')]
effect_rem = [expr('At(Sibiu)')]
fly_s_b = Action(expr('Fly(Sibiu, Bucharest)'), [precond_pos, precond_neg], [effect_add, effect_rem])
#Bucharest to Sibiu
precond_pos = [expr('At(Bucharest)')]
precond_neg = []
effect_add = [expr('At(Sibiu)')]
effect_rem = [expr('At(Bucharest)')]
fly_b_s = Action(expr('Fly(Bucharest, Sibiu)'), [precond_pos, precond_neg], [effect_add, effect_rem])
#Sibiu to Craiova
precond_pos = [expr('At(Sibiu)')]
precond_neg = []
effect_add = [expr('At(Craiova)')]
effect_rem = [expr('At(Sibiu)')]
fly_s_c = Action(expr('Fly(Sibiu, Craiova)'), [precond_pos, precond_neg], [effect_add, effect_rem])
#Craiova to Sibiu
precond_pos = [expr('At(Craiova)')]
precond_neg = []
effect_add = [expr('At(Sibiu)')]
effect_rem = [expr('At(Craiova)')]
fly_c_s = Action(expr('Fly(Craiova, Sibiu)'), [precond_pos, precond_neg], [effect_add, effect_rem])
#Bucharest to Craiova
precond_pos = [expr('At(Bucharest)')]
precond_neg = []
effect_add = [expr('At(Craiova)')]
effect_rem = [expr('At(Bucharest)')]
fly_b_c = Action(expr('Fly(Bucharest, Craiova)'), [precond_pos, precond_neg], [effect_add, effect_rem])
#Craiova to Bucharest
precond_pos = [expr('At(Craiova)')]
precond_neg = []
effect_add = [expr('At(Bucharest)')]
effect_rem = [expr('At(Craiova)')]
fly_c_b = Action(expr('Fly(Craiova, Bucharest)'), [precond_pos, precond_neg], [effect_add, effect_rem])
```
And the drive actions like this.
```
#Drive
precond_pos = [expr('At(x)')]
precond_neg = []
effect_add = [expr('At(y)')]
effect_rem = [expr('At(x)')]
drive = Action(expr('Drive(x, y)'), [precond_pos, precond_neg], [effect_add, effect_rem])
```
Finally, we can define a a function that will tell us when we have reached our destination, Bucharest.
```
def goal_test(kb):
return kb.ask(expr("At(Bucharest)"))
```
Thus, with all the components in place, we can define the planning problem.
```
prob = PDDL(knowledge_base, [fly_s_b, fly_b_s, fly_s_c, fly_c_s, fly_b_c, fly_c_b, drive], goal_test)
```
| github_jupyter |
# Define similarity-measure functions
```
# define similarity-measure functions
library(proxy)
library(transport)
library(dtw)
# Kullback-Leibler divergence
KL <- function(s1,s2){
s1[ which(s1<0)] <- 0
s2[ which(s2 <0)] <- 0
s1 <- s1/sum(s1)
s2 <- s2/sum(s2)
id <- intersect(which( (s1>0)), which( (s2>0)))
sum(s1[id] * log (s1[id]/s2[id]))
}
# Jensen-Shannon divergence (JSD)
JS <- function(s1,s2){
sm <- (s1+s2)/2
KL(s1,sm) + KL(s2,sm)
}
# Earth Mover's Distance (EMD)
EMD <- function(s1,s2){
s1 <- s1/sum(s1)
s2 <- s2/sum(s2)
costm <- outer(1:length(s1), 1:length(s2), FUN = function(x, y) abs(x - y))
res <- transport(s1, s2, costm)
sum(diag(costm[res$from, res$to]) * res$mass)
}
# target <- xas # dataset to calculate similarity
# ref <- 1 # reference data for similarity calculation
distance <- function(target,ref,nlen){
# calculate distance matrix
distm.e <- as.matrix(dist(target))
distm.m <- as.matrix(dist(target,method = "manhattan"))
distm.cos <- as.matrix(dist(target,method = "cosine"))
distm.p <- cor(t(target),method = "pearson")
dist.dtw <- dist.emd <- NULL
for(i in 1:(nlen)){
dist.dtw[i] <- dtw(target[i,],target[ref,])$distance
dist.emd[i] <- EMD(target[i,],target[ref,])
}
# distance between data and the spectrum at 10Dq=0
data.frame(
e = distm.e[ref,],
m = distm.m[ref,],
cos = distm.cos[ref,],
p = distm.p[ref,],
js = apply(target,1,function(x) JS(target[ref,],x)),
dtw = dist.dtw,
emd = dist.emd
)
}
similarity <- function(d,dmax){
# convert distance to similarity
data.frame(
e = 1 - d$e/dmax$e,
m = 1 - d$m/dmax$m,
cos = 1 - d$cos/dmax$cos,
p = 1 - (1-d$p)/(1-dmax$p),
js = 1 - d$js/dmax$js,
dtw = 1 - d$dtw/dmax$dtw,
emd = 1 - d$emd/dmax$emd
)
}
dmax <- function(d,nlen){
data.frame(e = d$e[nlen],
m = d$m[nlen],
cos = d$cos[nlen],
p = d$p[nlen],
js = d$js[nlen],
dtw = d$dtw[nlen],
emd = d$emd[nlen]
)
}
# calculate the maximum of the distance for normalization.
xas <- read.csv("./Mn2_XAS_raw.csv",header=T, check.names=F)
pe <- xas$'PhotonEnergy (eV)'
specs <- xas[,-27]
dq <- as.numeric(colnames(specs))
specs <- t(specs)
nlen.xas <- length(dq)
target <- specs
ref <- 1
dist.xas <- distance(target,ref,nlen.xas)
dmax.xas <- dmax(dist.xas,nlen.xas)
```
# Figure 3. Similarity as a function of 10Dq
```
xas <- read.csv('Mn2_XAS.csv',header=TRUE, check.names=FALSE)
pe <- xas$'PhotonEnergy (eV)'
specs <- xas[,-27]
specs <- t(specs)
dq <- seq(0, 2.5, by=0.1)
nlen.xas <- length(dq)
target <- specs
ref <- 1
dist.xas <- distance(target,ref,nlen.xas)
dmax.comp <- dmax(dist.xas,nlen.xas)
sim <- similarity(dist.xas, dmax.comp)
# write.csv(sim, "sim_10Dq.csv", quote=FALSE, row.names=TRUE)
# Plot symilarity vs 10Dq
plot(dq,sim$e,ylim=c(0,1),type="l",lwd=1,col="blue",xlab="10Dq",ylab="Similarity",tcl=0.5)
lines(dq,sim$m,type="l",lwd=1,ylim=c(0,1),col="green",ann="F")
lines(dq,sim$p,type="l",lwd=1,ylim=c(0,1),col="yellow",ann="F")
lines(dq,sim$cos,type="l",lwd=1,ylim=c(0,1),col="orange",ann="F")
lines(dq,sim$js,type="l",lwd=1,ylim=c(0,1),col="black",ann="F")
lines(dq,sim$dtw,type="l",lwd=1,ylim=c(0,1),col="purple",ann="F")
lines(dq,sim$emd,type="l",lwd=1,ylim=c(0,1),col="red",ann="F")
legend("topright", legend = c("Euclid","City Block","Pearson","Cosine","JSD","DTW","EMD"),
col = c("blue", "green", "yellow", "orange", "black", "purple", "red"),lty = 1)
xas_exp_calc <- read.csv('MnOexp.csv',header=TRUE)
xas_comparison <- data.frame(
'Mn2Oh_calc' = specs[1,]/sum(specs[1,]),
'MnO_exp' = xas_exp_calc$MnO_XAS
)
target <- t(xas_comparison)
ref <- 1
nlen.xas <- 2
dist.xas <- distance(target,ref,nlen.xas)
sim_exp <- similarity(dist.xas, dmax.comp)
sim_exp
```
## build a regression model
fit the polynomial functions (up to ten-degrees) to the data, and choose the suitable model with Akaike Information Criterion.
```
library(MuMIn)
options(na.action = "na.fail")
df <- data.frame(y = dq,
x = poly(sim$p,10,raw=TRUE))
fullmodel <- lm(y~.,data=df)
ret <- dredge(fullmodel,rank="AIC")
best.model <- get.models(ret, subset = 1)[1][[1]]
pred <- predict(best.model,interval ="confidence",level = 0.95)
plot(x=sim$p,y=dq,xlab="Similarity",ylab="10Dq",tcl=0.5)
lines(x=sim$p,y=pred[,1], col="red", lwd=2)
lines(x=sim$p,y=pred[,2], col="blue")
lines(x=sim$p,y=pred[,3], col="blue")
summary(best.model)
```
# Figure 5.Similarity as a function of with Gaussian noise.
```
xas.noise <- read.csv("./xas_noise_snr.csv", header=TRUE, check.names=FALSE)
snr <- xas.noise$'snr'
specs.noise <- xas.noise[, -602]
target <- specs.noise
ref <- 1
nlen.noise <- dim(specs.noise)[1]
dist.noise <- distance(target,ref,nlen.noise)
sim <- similarity(dist.noise, dmax.xas)
pe <- seq(630, 660, length.out = 601)
plot(pe,specs.noise[1,], xlim=c(637,657), ylim=c(0, 1.8*10^-2), type="l",
xlab="Photon Energy (eV)", ylab="Intensity",tcl=0.5)
par(new=T);plot(pe, specs.noise[54,]+5*10^-3, xlim=c(637,657), ylim=c(0, 1.8*10^-2), pch=20, ann="F", col="orange")
par(new=T);plot(pe, specs.noise[92,]+10*10^-3, xlim=c(637,657), ylim=c(0, 1.8*10^-2), pch=20, ann="F", col="red")
legend("topright", legend = c("ideal", "S/N = 100", "S/N = 30"),col = c("black", "orange", "red"),lty = 1)
# Plot similarity as a function of SNR
plot(snr,sim$e,xlim=c(max(snr), 5),ylim=c(0,1),type="l",lwd=2,col="blue",
xlab="Signal to Noise Ratio",ylab="Similarity",log="x",tcl=0.5)
lines(snr,sim$m,type="l",xlim=c(max(snr), 5),lwd=2,ylim=c(0,1),col="green",ann="F")
lines(snr,sim$p,type="l",xlim=c(max(snr), 5),lwd=2,ylim=c(0,1),col="yellow",ann="F")
lines(snr,sim$cos,type="l",xlim=c(max(snr), 5),lwd=2,ylim=c(0,1),col="orange",ann="F")
lines(snr,sim$js,type="l",xlim=c(max(snr), 5),lwd=2,ylim=c(0,1),col="black",ann="F")
lines(snr,sim$dtw,type="l",xlim=c(max(snr), 5),lwd=2,ylim=c(0,1),col="purple",ann="F")
lines(snr,sim$emd,type="l",xlim=c(max(snr), 5),lwd=2,ylim=c(0,1),col="red",ann="F")
abline(v=30,lty=2);abline(v=100,lty=2)
legend("bottomleft", legend = c("Euclid","City Block","Pearson","Cosine","JS divergence","DTW","EMD"),
col = c("blue", "green", "yellow", "orange", "black", "purple", "red"),lty = 1)
# plot the 100times averaged result.
load(file = 'sim_gaussnoise_100avg.Rdata')
load(file = 'snr_gaussnoise_100avg.Rdata')
# Plot similarity as a function of SNR
plot(snr,sim$e,xlim=c(max(snr),5),ylim=c(0,1),type="l",lwd=2,col="blue",
xlab="Signal to Noise Ratio",ylab="Similarity",log="x",tcl=0.5)
lines(snr,sim$m,type="l",xlim=c(max(snr),5),lwd=2,ylim=c(0,1),col="green",ann="F")
lines(snr,sim$p,type="l",xlim=c(max(snr),5),lwd=2,ylim=c(0,1),col="yellow",ann="F")
lines(snr,sim$cos,type="l",xlim=c(max(snr),5),lwd=2,ylim=c(0,1),col="orange",ann="F")
lines(snr,sim$js,type="l",xlim=c(max(snr),5),lwd=2,ylim=c(0,1),col="black",ann="F")
lines(snr,sim$dtw,type="l",xlim=c(max(snr),5),lwd=2,ylim=c(0,1),col="purple",ann="F")
lines(snr,sim$emd,type="l",xlim=c(max(snr),5),lwd=2,ylim=c(0,1),col="red",ann="F")
legend("bottomleft", legend = c("Euclid","City Block","Pearson","Cosine","JS divergence","DTW","EMD"),
col = c("blue", "green", "yellow", "orange", "black", "purple", "red"),lty = 1)
# write.csv(sim, "sim_snr.csv", quote=FALSE, row.names=TRUE)
# write.csv(snr, "snr.csv", quote=FALSE, row.names=TRUE)
```
# Figure 6.Similarity as a function of $\sigma$.
```
xas.broad <- read.csv("./Mn2_00_broading.csv", header=TRUE, check.names=FALSE)
pe <- xas.broad$'PhotonEnergy (eV)'
broad <- as.numeric(colnames(xas.broad[,-11]))
sigma.gauss = broad/(2*(sqrt(2*log(2))))
specs.broad <- t(xas.broad[,-11])
target <- specs.broad
ref <- 1
nlen.broad <- dim(specs.broad)[1]
dist.broad <- distance(target,ref,nlen.broad)
sim <- similarity(dist.broad, dmax.xas)
# write.csv(sim, "sim_broad.csv", quote=FALSE, row.names=TRUE)
plot(pe,specs.broad[1,],xlim=c(637,657),ylim=c(0,0.02),type="l",
xlab="Photon Energy (eV)",ylab="Intensity",tcl=0.5)
lines(pe,specs.broad[4,]+0.0075,xlim=c(637,657),ylim=c(0,0.02),type="l",ann="F", col="orange")
lines(pe,specs.broad[6,]+0.015,xlim=c(637,657),ylim=c(0,0.02),type="l",ann="F", col="red")
legend("topright", legend = c("sigma = 0.02", "sigma = 0.09", "sigma = 0.13"),col = c("black", "orange", "red"),lty = 1)
graphics.off()
# Plot similarity as a function of sigma
plot(sigma.gauss,sim$e,ylim=c(0,1),type="l",lwd=2,col="blue",
xlab="sigma of Gauss dist.(eV)",ylab="Similarity",tcl=0.5)
lines(sigma.gauss,sim$m,type="l",lwd=2,ylim=c(0,1),col="green",ann="F")
lines(sigma.gauss,sim$p,type="l",lwd=2,ylim=c(0,1),col="yellow",ann="F")
lines(sigma.gauss,sim$cos,type="l",lwd=2,ylim=c(0,1),col="orange",ann="F")
lines(sigma.gauss,sim$js,type="l",lwd=2,ylim=c(0,1),col="black",ann="F")
lines(sigma.gauss,sim$dtw,type="l",lwd=2,ylim=c(0,1),col="purple",ann="F")
lines(sigma.gauss,sim$emd,type="l",lwd=2,ylim=c(0,1),col="red",ann="F")
legend("bottomleft", legend = c("Euclid","City Block","Pearson","Cosine","JS divergence","DTW","EMD"),
col = c("blue", "green", "yellow", "orange", "black", "purple", "red"),lty = 1)
abline(v=sigma.gauss[4],lty=2);abline(v=sigma.gauss[6],lty=2)
plot(sigma.gauss,sim$e,ylim=c(0,1),type="l",lwd=2,col="blue",
xlab="sigma of Gauss dist.(eV)",ylab="Similarity",tcl=0.5)
lines(sigma.gauss,sim$m,type="l",lwd=2,ylim=c(0,1),col="green",ann="F")
lines(sigma.gauss,sim$p,type="l",lwd=2,ylim=c(0,1),col="yellow",ann="F")
lines(sigma.gauss,sim$cos,type="l",lwd=2,ylim=c(0,1),col="orange",ann="F")
lines(sigma.gauss,sim$js,type="l",lwd=2,ylim=c(0,1),col="black",ann="F")
lines(sigma.gauss,sim$dtw,type="l",lwd=2,ylim=c(0,1),col="purple",ann="F")
lines(sigma.gauss,sim$emd,type="l",lwd=2,ylim=c(0,1),col="red",ann="F")
abline(v=sigma.gauss[4],lty=2);abline(v=sigma.gauss[6],lty=2)
legend("bottomleft", legend = c("Euclid","City Block","Pearson","Cosine","JS divergence","DTW","EMD"),
col = c("blue", "green", "yellow", "orange", "black", "purple", "red"),lty = 1)
```
| github_jupyter |
```
# Simple notebook to extract features from images, using the MobileNetv2 model
# output at the prior to classification layer
# Saved as a numpy array for LSTM modelling
import warnings
warnings.filterwarnings('ignore')
import os
from matplotlib import pyplot as plt
%matplotlib inline
import seaborn as sns
import scipy
import numpy as np
from sklearn.metrics import confusion_matrix,classification_report
from sklearn.utils import class_weight
from PIL import Image
import time
from pathlib import Path
import pandas as pd
from keras import layers
from keras.layers import Dense, Flatten, GlobalAveragePooling2D
from keras.models import Model
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import ModelCheckpoint, EarlyStopping
from keras.optimizers import RMSprop, SGD, Adam
from keras.applications import Xception, MobileNetV2
from keras.applications.inception_v3 import InceptionV3
import tensorflow as tf
# If GPU Uncomment
#print("Num GPUs Available: ", len(tf.config.experimental.list_physical_devices('GPU')))
#gpus = tf.config.experimental.list_physical_devices('GPU')
#tf.config.experimental.set_memory_growth(gpus[0], True)
fps = '2FPS'
numpy_dir = '../../data/DAiSEE/' + fps + '/data/'
features_dir = os.getcwd() + '/features'
usage = ['Train', 'Test', 'Validation']
# Make sure features directory exists
if not os.path.exists(features_dir):
os.makedirs(features_dir)
def get_array(usage, numpy_dir):
numpy_dir = Path(numpy_dir)
x = np.load(numpy_dir / f'x_{usage.lower()}.npy', allow_pickle=True)
y = np.load(numpy_dir / f'y_lab_{usage.lower()}.npy')
return (x,y)
def get_dataframe(usage, numpy_dir):
x_files, y = get_dataset(usage, numpy_dir)
df_x = pd.DataFrame(x_files, columns = ['file'])
df_y = pd.DataFrame(y, columns = ['label'])
df = pd.concat([df_x.reset_index(drop=True),df_y.reset_index(drop=True)], axis=1)
df['file'] = df['file'].astype(str)
df['label'] = df['label'].astype(str)
return (df)
df_train = get_dataframe('train', numpy_dir)
df_test = get_dataframe('test', numpy_dir)
df_val = get_dataframe('val', numpy_dir)
df_train.head()
datagen = ImageDataGenerator(rescale=1.0/255.0)
train_generator = datagen.flow_from_dataframe(
dataframe = df_train,
x_col = 'file',
y_col = 'label',
target_size = (224, 224),
batch_size = 32,
shuffle = False,
class_mode= 'categorical')
val_generator = datagen.flow_from_dataframe(
dataframe = df_val,
x_col = 'file',
y_col = 'label',
target_size = (224, 224),
batch_size = 32,
shuffle = False,
class_mode= 'categorical')
test_generator = datagen.flow_from_dataframe(
dataframe = df_test,
x_col = 'file',
y_col = 'label',
target_size = (224, 224),
batch_size = 32,
shuffle = False,
class_mode= None)
base_model = MobileNetV2(
weights='imagenet',
include_top=True
)
base_model.summary()
model = Model(
inputs=base_model.input,
outputs=base_model.get_layer('global_average_pooling2d_1').output
)
features = model.predict_generator(train_generator)
y_train = train_generator.classes
np.save(features_dir + '/train_features.npy', features)
np.save(features_dir + '/train_labels.npy', y_train)
print(len(features))
val_features = model.predict_generator(val_generator)
y_val = val_generator.classes
np.save(features_dir + '/val_features.npy', features)
np.save(features_dir + '/val_labels.npy', y_train)
print(len(val_features))
test_features = model.predict_generator(test_generator)
y_test = test_generator.classes
np.save(features_dir + '/test_features.npy', features)
np.save(features_dir + '/test_labels.npy', y_train)
print(len(val_features))
```
| github_jupyter |
[](https://colab.research.google.com/github/tulasiram58827/ocr_tflite/blob/main/colabs/KERAS_OCR_TFLITE.ipynb)
## SetUp
```
!pip install tf-nightly
!pip install validators
import typing
import string
import tensorflow as tf
from tensorflow import keras
import numpy as np
import cv2
import os
import hashlib
import urllib.request
import urllib.parse
tf.__version__
```
**All of the code required to build the model and load the weights are taken from this [repository](https://github.com/faustomorales/keras-ocr) including the pretrained weights**
### Hyper-Parameters
```
DEFAULT_BUILD_PARAMS = {
'height': 31,
'width': 200,
'color': False,
'filters': (64, 128, 256, 256, 512, 512, 512),
'rnn_units': (128, 128),
'dropout': 0.25,
'rnn_steps_to_discard': 2,
'pool_size': 2,
'stn': True,
}
DEFAULT_ALPHABET = string.digits + string.ascii_lowercase
PRETRAINED_WEIGHTS = {
'kurapan': {
'alphabet': DEFAULT_ALPHABET,
'build_params': DEFAULT_BUILD_PARAMS,
'weights': {
'notop': {
'url':
'https://github.com/faustomorales/keras-ocr/releases/download/v0.8.4/crnn_kurapan_notop.h5',
'filename': 'crnn_kurapan_notop.h5',
'sha256': '027fd2cced3cbea0c4f5894bb8e9e85bac04f11daf96b8fdcf1e4ee95dcf51b9'
},
'top': {
'url':
'https://github.com/faustomorales/keras-ocr/releases/download/v0.8.4/crnn_kurapan.h5',
'filename': 'crnn_kurapan.h5',
'sha256': 'a7d8086ac8f5c3d6a0a828f7d6fbabcaf815415dd125c32533013f85603be46d'
}
}
}
}
```
## Utilities
```
def swish(x, beta=1):
return x * keras.backend.sigmoid(beta * x)
keras.utils.get_custom_objects().update({'swish': keras.layers.Activation(swish)})
def _repeat(x, num_repeats):
ones = tf.ones((1, num_repeats), dtype='int32')
x = tf.reshape(x, shape=(-1, 1))
x = tf.matmul(x, ones)
return tf.reshape(x, [-1])
def _meshgrid(height, width):
x_linspace = tf.linspace(-1., 1., width)
y_linspace = tf.linspace(-1., 1., height)
x_coordinates, y_coordinates = tf.meshgrid(x_linspace, y_linspace)
x_coordinates = tf.reshape(x_coordinates, shape=(1, -1))
y_coordinates = tf.reshape(y_coordinates, shape=(1, -1))
ones = tf.ones_like(x_coordinates)
indices_grid = tf.concat([x_coordinates, y_coordinates, ones], 0)
return indices_grid
# pylint: disable=too-many-statements
def _transform(inputs):
locnet_x, locnet_y = inputs
output_size = locnet_x.shape[1:]
batch_size = tf.shape(locnet_x)[0]
height = tf.shape(locnet_x)[1]
width = tf.shape(locnet_x)[2]
num_channels = tf.shape(locnet_x)[3]
locnet_y = tf.reshape(locnet_y, shape=(batch_size, 2, 3))
locnet_y = tf.reshape(locnet_y, (-1, 2, 3))
locnet_y = tf.cast(locnet_y, 'float32')
output_height = output_size[0]
output_width = output_size[1]
indices_grid = _meshgrid(output_height, output_width)
indices_grid = tf.expand_dims(indices_grid, 0)
indices_grid = tf.reshape(indices_grid, [-1]) # flatten?
indices_grid = tf.tile(indices_grid, tf.stack([batch_size]))
indices_grid = tf.reshape(indices_grid, tf.stack([batch_size, 3, -1]))
transformed_grid = tf.matmul(locnet_y, indices_grid)
x_s = tf.slice(transformed_grid, [0, 0, 0], [-1, 1, -1])
y_s = tf.slice(transformed_grid, [0, 1, 0], [-1, 1, -1])
x = tf.reshape(x_s, [-1])
y = tf.reshape(y_s, [-1])
# Interpolate
height_float = tf.cast(height, dtype='float32')
width_float = tf.cast(width, dtype='float32')
output_height = output_size[0]
output_width = output_size[1]
x = tf.cast(x, dtype='float32')
y = tf.cast(y, dtype='float32')
x = .5 * (x + 1.0) * width_float
y = .5 * (y + 1.0) * height_float
x0 = tf.cast(tf.floor(x), 'int32')
x1 = x0 + 1
y0 = tf.cast(tf.floor(y), 'int32')
y1 = y0 + 1
max_y = tf.cast(height - 1, dtype='int32')
max_x = tf.cast(width - 1, dtype='int32')
zero = tf.zeros([], dtype='int32')
x0 = tf.clip_by_value(x0, zero, max_x)
x1 = tf.clip_by_value(x1, zero, max_x)
y0 = tf.clip_by_value(y0, zero, max_y)
y1 = tf.clip_by_value(y1, zero, max_y)
flat_image_dimensions = width * height
pixels_batch = tf.range(batch_size) * flat_image_dimensions
flat_output_dimensions = output_height * output_width
base = _repeat(pixels_batch, flat_output_dimensions)
base_y0 = base + y0 * width
base_y1 = base + y1 * width
indices_a = base_y0 + x0
indices_b = base_y1 + x0
indices_c = base_y0 + x1
indices_d = base_y1 + x1
flat_image = tf.reshape(locnet_x, shape=(-1, num_channels))
flat_image = tf.cast(flat_image, dtype='float32')
pixel_values_a = tf.gather(flat_image, indices_a)
pixel_values_b = tf.gather(flat_image, indices_b)
pixel_values_c = tf.gather(flat_image, indices_c)
pixel_values_d = tf.gather(flat_image, indices_d)
x0 = tf.cast(x0, 'float32')
x1 = tf.cast(x1, 'float32')
y0 = tf.cast(y0, 'float32')
y1 = tf.cast(y1, 'float32')
area_a = tf.expand_dims(((x1 - x) * (y1 - y)), 1)
area_b = tf.expand_dims(((x1 - x) * (y - y0)), 1)
area_c = tf.expand_dims(((x - x0) * (y1 - y)), 1)
area_d = tf.expand_dims(((x - x0) * (y - y0)), 1)
transformed_image = tf.add_n([
area_a * pixel_values_a, area_b * pixel_values_b, area_c * pixel_values_c,
area_d * pixel_values_d
])
# Finished interpolation
transformed_image = tf.reshape(transformed_image,
shape=(batch_size, output_height, output_width, num_channels))
return transformed_image
```
## Create Model
```
def build_model(alphabet,
height,
width,
color,
filters,
rnn_units,
dropout,
rnn_steps_to_discard,
pool_size,
stn=True):
"""Build a Keras CRNN model for character recognition.
Args:
height: The height of cropped images
width: The width of cropped images
color: Whether the inputs should be in color (RGB)
filters: The number of filters to use for each of the 7 convolutional layers
rnn_units: The number of units for each of the RNN layers
dropout: The dropout to use for the final layer
rnn_steps_to_discard: The number of initial RNN steps to discard
pool_size: The size of the pooling steps
stn: Whether to add a Spatial Transformer layer
"""
assert len(filters) == 7, '7 CNN filters must be provided.'
assert len(rnn_units) == 2, '2 RNN filters must be provided.'
inputs = keras.layers.Input((height, width, 3 if color else 1), name='input', batch_size=1)
x = keras.layers.Permute((2, 1, 3))(inputs)
x = keras.layers.Lambda(lambda x: x[:, :, ::-1])(x)
x = keras.layers.Conv2D(filters[0], (3, 3), activation='relu', padding='same', name='conv_1')(x)
x = keras.layers.Conv2D(filters[1], (3, 3), activation='relu', padding='same', name='conv_2')(x)
x = keras.layers.Conv2D(filters[2], (3, 3), activation='relu', padding='same', name='conv_3')(x)
x = keras.layers.BatchNormalization(name='bn_3')(x)
x = keras.layers.MaxPooling2D(pool_size=(pool_size, pool_size), name='maxpool_3')(x)
x = keras.layers.Conv2D(filters[3], (3, 3), activation='relu', padding='same', name='conv_4')(x)
x = keras.layers.Conv2D(filters[4], (3, 3), activation='relu', padding='same', name='conv_5')(x)
x = keras.layers.BatchNormalization(name='bn_5')(x)
x = keras.layers.MaxPooling2D(pool_size=(pool_size, pool_size), name='maxpool_5')(x)
x = keras.layers.Conv2D(filters[5], (3, 3), activation='relu', padding='same', name='conv_6')(x)
x = keras.layers.Conv2D(filters[6], (3, 3), activation='relu', padding='same', name='conv_7')(x)
x = keras.layers.BatchNormalization(name='bn_7')(x)
if stn:
# pylint: disable=pointless-string-statement
"""Spatial Transformer Layer
Implements a spatial transformer layer as described in [1]_.
Borrowed from [2]_:
downsample_fator : float
A value of 1 will keep the orignal size of the image.
Values larger than 1 will down sample the image. Values below 1 will
upsample the image.
example image: height= 100, width = 200
downsample_factor = 2
output image will then be 50, 100
References
----------
.. [1] Spatial Transformer Networks
Max Jaderberg, Karen Simonyan, Andrew Zisserman, Koray Kavukcuoglu
Submitted on 5 Jun 2015
.. [2] https://github.com/skaae/transformer_network/blob/master/transformerlayer.py
.. [3] https://github.com/EderSantana/seya/blob/keras1/seya/layers/attention.py
"""
stn_input_output_shape = (width // pool_size**2, height // pool_size**2, filters[6])
stn_input_layer = keras.layers.Input(shape=stn_input_output_shape)
locnet_y = keras.layers.Conv2D(16, (5, 5), padding='same',
activation='relu')(stn_input_layer)
locnet_y = keras.layers.Conv2D(32, (5, 5), padding='same', activation='relu')(locnet_y)
locnet_y = keras.layers.Flatten()(locnet_y)
locnet_y = keras.layers.Dense(64, activation='relu')(locnet_y)
locnet_y = keras.layers.Dense(6,
weights=[
np.zeros((64, 6), dtype='float32'),
np.float32([[1, 0, 0], [0, 1, 0]]).flatten()
])(locnet_y)
localization_net = keras.models.Model(inputs=stn_input_layer, outputs=locnet_y)
x = keras.layers.Lambda(_transform,
output_shape=stn_input_output_shape)([x, localization_net(x)])
x = keras.layers.Reshape(target_shape=(width // pool_size**2,
(height // pool_size**2) * filters[-1]),
name='reshape')(x)
x = keras.layers.Dense(rnn_units[0], activation='relu', name='fc_9')(x)
rnn_1_forward = keras.layers.LSTM(rnn_units[0],
kernel_initializer="he_normal",
return_sequences=True,
name='lstm_10')(x)
rnn_1_back = keras.layers.LSTM(rnn_units[0],
kernel_initializer="he_normal",
go_backwards=True,
return_sequences=True,
name='lstm_10_back')(x)
rnn_1_add = keras.layers.Add()([rnn_1_forward, rnn_1_back])
rnn_2_forward = keras.layers.LSTM(rnn_units[1],
kernel_initializer="he_normal",
return_sequences=True,
name='lstm_11')(rnn_1_add)
rnn_2_back = keras.layers.LSTM(rnn_units[1],
kernel_initializer="he_normal",
go_backwards=True,
return_sequences=True,
name='lstm_11_back')(rnn_1_add)
x = keras.layers.Concatenate()([rnn_2_forward, rnn_2_back])
backbone = keras.models.Model(inputs=inputs, outputs=x)
x = keras.layers.Dropout(dropout, name='dropout')(x)
x = keras.layers.Dense(len(alphabet) + 1,
kernel_initializer='he_normal',
activation='softmax',
name='fc_12')(x)
x = keras.layers.Lambda(lambda x: x[:, rnn_steps_to_discard:])(x)
model = keras.models.Model(inputs=inputs, outputs=x)
# prediction_model = keras.models.Model(inputs=inputs, outputs=CTCDecoder()(model.output))
# labels = keras.layers.Input(name='labels', shape=[model.output_shape[1]], dtype='float32')
# label_length = keras.layers.Input(shape=[1])
# input_length = keras.layers.Input(shape=[1])
# loss = keras.layers.Lambda(lambda inputs: keras.backend.ctc_batch_cost(
# y_true=inputs[0], y_pred=inputs[1], input_length=inputs[2], label_length=inputs[3]))(
# [labels, model.output, input_length, label_length])
# training_model = keras.models.Model(inputs=[model.input, labels, input_length, label_length],
# outputs=loss)
# We are commenting the above lines because CTC Decoder here is not supported in TFLite hence we are
# discarding the CTC Decoder Portion model.
return model
build_params = DEFAULT_BUILD_PARAMS
alphabets = DEFAULT_ALPHABET
blank_index = len(alphabets)
# While building the model we are not including the CTC Decoder as it is not convertable to TFLite
model = build_model(alphabet=alphabets, **build_params)
```
## Download and Load Weights
```
def get_default_cache_dir():
return os.environ.get('KERAS_OCR_CACHE_DIR', os.path.expanduser(os.path.join('~',
'.keras-ocr')))
def sha256sum(filename):
"""Compute the sha256 hash for a file."""
h = hashlib.sha256()
b = bytearray(128 * 1024)
mv = memoryview(b)
with open(filename, 'rb', buffering=0) as f:
for n in iter(lambda: f.readinto(mv), 0):
h.update(mv[:n])
return h.hexdigest()
def download_and_verify(url, sha256=None, cache_dir=None, verbose=True, filename=None):
"""Download a file to a cache directory and verify it with a sha256
hash.
Args:
url: The file to download
sha256: The sha256 hash to check. If the file already exists and the hash
matches, we don't download it again.
cache_dir: The directory in which to cache the file. The default is
`~/.keras-ocr`.
verbose: Whether to log progress
filename: The filename to use for the file. By default, the filename is
derived from the URL.
"""
if cache_dir is None:
cache_dir = get_default_cache_dir()
if filename is None:
filename = os.path.basename(urllib.parse.urlparse(url).path)
filepath = os.path.join(cache_dir, filename)
os.makedirs(os.path.split(filepath)[0], exist_ok=True)
if verbose:
print('Looking for ' + filepath)
if not os.path.isfile(filepath) or (sha256 and sha256sum(filepath) != sha256):
if verbose:
print('Downloading ' + filepath)
urllib.request.urlretrieve(url, filepath)
assert sha256 is None or sha256 == sha256sum(filepath), 'Error occurred verifying sha256.'
return filepath
weights_dict = PRETRAINED_WEIGHTS['kurapan']
model.load_weights(download_and_verify(url=weights_dict['weights']['top']['url'],
filename=weights_dict['weights']['top']['filename'],
sha256=weights_dict['weights']['top']['sha256']))
```
## Model Architecture
```
model.summary()
```
## Convert to TFLite
```
# Download and unzipping representative dataset
%%bash
wget https://github.com/tulasiram58827/ocr_tflite/raw/main/data/represent_data.zip
unzip represent_data.zip
dataset_path = '/content/represent_data/'
def representative_data_gen():
for file in os.listdir(dataset_path):
image_path = dataset_path + file
input_data = cv2.imread(image_path, cv2.IMREAD_GRAYSCALE)
input_data = cv2.resize(input_data, (200, 31))
input_data = input_data[np.newaxis]
input_data = np.expand_dims(input_data, 3)
input_data = input_data.astype('float32')/255
yield [input_data]
def convert_tflite(quantization):
converter = tf.lite.TFLiteConverter.from_keras_model(model)
converter.optimizations = [tf.lite.Optimize.DEFAULT]
if quantization == 'float16':
converter.target_spec.supported_types = [tf.float16]
elif quantization == 'int8' or quantization == 'full_int8':
converter.representative_dataset = representative_data_gen
if quantization == 'full_int8':
converter.target_spec.supported_ops = [tf.lite.OpsSet.TFLITE_BUILTINS_INT8]
converter.inference_input_type = tf.int8 # or tf.uint8
converter.inference_output_type = tf.int8 # or tf.uint8
tf_lite_model = converter.convert()
open(f'ocr_{quantization}.tflite', 'wb').write(tf_lite_model)
```
**Note** : Support for CTC Decoder is not available in TFLite yet. So while converting we removed CTCDecoder in model part. We need to run Decoder from the output of the model.
Refer to this [issue](https://github.com/tensorflow/tensorflow/issues/33494) regarding CTC decoder support in TFLite.
This is the code for [CTC DECODER](https://colab.research.google.com/github/tulasiram58827/ocr_tflite/blob/main/colabs/KERAS_OCR_TFLITE.ipynb#scrollTo=_rdJyCXo2Xzs). By default it is greedy Decoder we can also use Beam Search Decoder by specifying the parameter in the `ctc_decode` function.
FYI: I am also working on converting Beam Search CTC Decoder to low level language so that we can port entire OCR and use it as offline application combinedly with EAST/CRAFT.
```
quantization = 'dr' #@param ["dr", "float16"]
convert_tflite(quantization)
!du -sh ocr_dr.tflite
quantization = 'float16' #@param ["dr", "float16"]
convert_tflite(quantization)
!du -sh ocr_float16.tflite
quantization = 'int8' #@param ["dr", "float16", 'int8', 'full_int8']
convert_tflite(quantization)
!du -sh ocr_int8.tflite
quantization = 'full_int8' #@param ["dr", "float16", 'int8', 'full_int8']
convert_tflite(quantization)
```
## TFLite Inference
#### CTC Decoder
```
# Code for CTC Decoder
def decoder(y_pred):
input_shape = tf.keras.backend.shape(y_pred)
input_length = tf.ones(shape=input_shape[0]) * tf.keras.backend.cast(
input_shape[1], 'float32')
# You can turn on beam search decoding using greedy=False and also play with beam_width parameter.
unpadded = tf.keras.backend.ctc_decode(y_pred, input_length)[0][0]
unpadded_shape = tf.keras.backend.shape(unpadded)
padded = tf.pad(unpadded,
paddings=[[0, 0], [0, input_shape[1] - unpadded_shape[1]]],
constant_values=-1)
return padded
```
#### TFLite model inference
```
!wget https://github.com/tulasiram58827/ocr_tflite/raw/main/images/demo_1.png
def run_tflite_model(image_path, quantization):
input_data = cv2.imread(image_path, cv2.IMREAD_GRAYSCALE)
input_data = cv2.resize(input_data, (200, 31))
input_data = input_data[np.newaxis]
input_data = np.expand_dims(input_data, 3)
input_data = input_data.astype('float32')/255
path = f'ocr_{quantization}.tflite'
interpreter = tf.lite.Interpreter(model_path=path)
interpreter.allocate_tensors()
# Get input and output tensors.
input_details = interpreter.get_input_details()
output_details = interpreter.get_output_details()
input_shape = input_details[0]['shape']
interpreter.set_tensor(input_details[0]['index'], input_data)
interpreter.invoke()
output = interpreter.get_tensor(output_details[0]['index'])
return output
from google.colab.patches import cv2_imshow
image_path = '/content/demo_1.png'
# Running Dynamic Range Quantization
tflite_output = run_tflite_model(image_path, 'dr')
# Running decoder on TFLite Output
decoded = decoder(tflite_output)
final_output = "".join(alphabets[index] for index in decoded[0] if index not in [blank_index, -1])
print(final_output)
cv2_imshow(cv2.imread(image_path))
# Running Float16 Quantization
tflite_output = run_tflite_model(image_path, 'float16')
# Running decoder on TFLite Output
decoded = decoder(tflite_output)
final_output = "".join(alphabets[index] for index in decoded[0] if index not in [blank_index, -1])
print(final_output)
cv2_imshow(cv2.imread(image_path))
# Running Integer Quantization
tflite_output = run_tflite_model(image_path, 'int8')
# Running decoder on TFLite Output
decoded = decoder(tflite_output)
final_output = "".join(alphabets[index] for index in decoded[0] if index not in [blank_index, -1])
print(final_output)
cv2_imshow(cv2.imread(image_path))
```
## Dynamic Range Model benchmarks
**Inference Time** : 0.2sec
**Memory FootPrint** : 46.38MB
**Model Size** : 8.5MB
## Float16 benchmarks
**Inference** : 0.76sec
**Memory FootPrint** : 128MB
**Model Size** : 17MB
| github_jupyter |
<h2>Vectors: One Dimensional Lists</h2>
A <b>vector</b> is a list of numbers.
Vectors are very useful to describe the state of a system, as we will see in the main tutorial.
A list is a single object in python.
Similarly, a vector is a single mathematical object.
The number of elements in a list is its size or length.
Similarly, the number of entries in a vector is called as the <b>size</b> or <b>dimension</b> of the vector.
```
# consider the following list with 4 elements
L = [1,-2,0,5]
print(L)
```
Vectors can be in horizontal or vertical shape.
We show this list as a <i><u>four dimensional</u></i> <b>row vector</b> (horizontal) or a <b>column vector</b> (vertical):
$$
u = \mypar{1~~-2~~0~~-5} ~~~\mbox{ or }~~~ v =\mymatrix{r}{1 \\ -2 \\ 0 \\ 5}, ~~~\mbox{ respectively.}
$$
Remark that we do not need to use any comma in vector representation.
<h3> Multiplying a vector with a number</h3>
A vector can be multiplied by a number.
Multiplication of a vector with a number is also a vector: each entry is multiplied by this number.
$$
3 \cdot v = 3 \cdot \mymatrix{r}{1 \\ -2 \\ 0 \\ 5} = \mymatrix{r}{3 \\ -6 \\ 0 \\ 15}
~~~~~~\mbox{ or }~~~~~~
(-0.6) \cdot v = (-0.6) \cdot \mymatrix{r}{1 \\ -2 \\ 0 \\ 5} = \mymatrix{r}{-0.6 \\ 1.2 \\ 0 \\ -3}.
$$
We may consider this as enlarging or making smaller the entries of a vector.
We verify our calculations in python.
```
# 3 * v
v = [1,-2,0,5]
print("v is",v)
# we use the same list for the result
for i in range(len(v)):
v[i] = 3 * v[i]
print("3v is",v)
# -0.6 * u
# reinitialize the list v
v = [1,-2,0,5]
for i in range(len(v)):
v[i] = -0.6 * v[i]
print("0.6v is",v)
```
<h3> Summation of vectors</h3>
Two vectors (with same dimension) can be summed up.
The summation of two vectors is a vector: the numbers on the same entries are added up.
$$
u = \myrvector{-3 \\ -2 \\ 0 \\ -1 \\ 4} \mbox{ and } v = \myrvector{-1\\ -1 \\2 \\ -3 \\ 5}.
~~~~~~~ \mbox{Then, }~~
u+v = \myrvector{-3 \\ -2 \\ 0 \\ -1 \\ 4} + \myrvector{-1\\ -1 \\2 \\ -3 \\ 5} =
\myrvector{-3+(-1)\\ -2+(-1) \\0+2 \\ -1+(-3) \\ 4+5} = \myrvector{-4\\ -3 \\2 \\ -4 \\ 9}.
$$
We do the same calculations in Python.
```
u = [-3,-2,0,-1,4]
v = [-1,-1,2,-3,5]
result=[]
for i in range(len(u)):
result.append(u[i]+v[i])
print("u+v is",result)
# print the result vector similarly to a column vector
print() # print an empty line
print("the elements of u+v are")
for j in range(len(result)):
print(result[j])
```
<h3> Task 1 </h3>
Create two 7-dimensional vectors $u$ and $ v $ as two different lists in Python having entries randomly picked between $-10$ and $10$.
Print their entries.
```
from random import randrange
#
# your solution is here
#
from random import randrange
dimension = 7
# create u and v as empty lists
u = []
v = []
for i in range(dimension):
u.append(randrange(-10,11)) # add a randomly picked number to the list u
v.append(randrange(-10,11)) # add a randomly picked number to the list v
# print both lists
print("u is",u)
print("v is",v)
#r=randrange(-10,11) # randomly pick a number from the list {-10,-9,...,-1,0,1,...,9,10}
```
<h3> Task 2 </h3>
By using the same vectors, find the vector $ (3 u-2 v) $ and print its entries. Here $ 3u $ and $ 2v $ means $u$ and $v$ are multiplied by $3$ and $2$, respectively.
```
#
# your solution is here
#
# please execute the cell for Task 1 to define u and v
# create a result list
# the first method
result=[]
# fill it with zeros
for i in range(dimension):
result.append(0)
print("by using the first method, the result vector is initialized to",result)
# the second method
# alternative and shorter solution for creating a list with zeros
result = [0] * 7
print("by using the second method, the result vector is initialized to",result)
# calculate 3u-2v
for i in range(dimension):
result[i] = 3 * u[i] - 2 * v[i]
# print all lists
print("u is",u)
print("v is",v)
print("3u-2v is",result)
```
<h3> Visualization of vectors </h3>
We can visualize the vectors with dimension at most 3.
For simplicity, we give examples of 2-dimensional vectors.
Consider the vector $ v = \myvector{1 \\ 2} $.
A 2-dimensional vector can be represented on the two-dimensional plane by an arrow starting from the origin $ (0,0) $ to the point $ (1,2) $.
<img src="../images/vector_1_2-small.jpg" width="40%">
We represent the vectors $ 2v = \myvector{2 \\ 4} $ and $ -v = \myvector{-1 \\ -2} $ below.
<img src="../images/vectors_2_4_-1_-2.jpg" width="40%">
As we can observe, after multiplying by 2, the vector is enlarged, and, after multiplying by $(-1)$, the vector is the same but its direction is opposite.
<h3> The length of a vector </h3>
The length of a vector is the (shortest) distance from the points represented by the entries of vector to the origin point $(0,0)$.
The length of a vector can be calculated by using Pythagoras Theorem.
We visualize a vector, its length, and the contributions of each entry to the length.
Consider the vector $ u = \myrvector{-3 \\ 4} $.
<img src="../images/length_-3_4-small.jpg" width="80%">
The length of $ u $ is denoted as $ \norm{u} $, and it is calculated as $ \norm{u} =\sqrt{(-3)^2+4^2} = 5 $.
Here each entry contributes with its square value. All contributions are summed up. Then, we obtain the square of the length.
This formula is generalized to any dimension.
We find the length of the following vector by using Python:
$$
v = \myrvector{-1 \\ -3 \\ 5 \\ 3 \\ 1 \\ 2}
~~~~~~~~~~
\mbox{and}
~~~~~~~~~~
\norm{v} = \sqrt{(-1)^2+(-3)^2+5^2+3^2+1^2+2^2} .
$$
<div style="font-style:italic;background-color:#fafafa;font-size:10pt;"> Remember: There is a short way of writing power operation in Python.
<ul>
<li> In its generic form: $ a^x $ can be denoted by $ a ** x $ in Python. </li>
<li> The square of a number $a$: $ a^2 $ can be denoted by $ a ** 2 $ in Python. </li>
<li> The square root of a number $ a $: $ \sqrt{a} = a^{\frac{1}{2}} = a^{0.5} $ can be denoted by $ a ** 0.5 $ in Python.</li>
</ul>
</div>
```
v = [-1,-3,5,3,1,2]
length_square=0
for i in range(len(v)):
print(v[i],":square ->",v[i]**2) # print each entry and its square value
length_square = length_square + v[i]**2 # sum up the square of each entry
length = length_square ** 0.5 # take the square root of the summation of the squares of all entries
print("the summation is",length_square)
print("then the length is",length)
# for square root, we can also use built-in function math.sqrt
print() # print an empty line
from math import sqrt
print("the square root of",length_square,"is",sqrt(length_square))
```
<h3> Task 3 </h3>
Let $ u = \myrvector{1 \\ -2 \\ -4 \\ 2} $ be a four dimensional vector.
Verify that $ \norm{4 u} = 4 \cdot \norm{u} $ in Python.
Remark that $ 4u $ is another vector obtained from $ u $ by multiplying it with 4.
```
#
# your solution is here
#
u = [1,-2,-4,2]
fouru=[4,-8,-16,8]
len_u = 0
len_fouru = 0
for i in range(len(u)):
len_u = len_u + u[i]**2 # adding square of each value
len_fouru = len_fouru + fouru[i]**2 # adding square of each value
len_u = len_u ** 0.5 # taking square root of the summation
len_fouru = len_fouru ** 0.5 # taking square root of the summation
# print the lengths
print("length of u is",len_u)
print("4 * length of u is",4 * len_u)
print("length of 4u is",len_fouru)
```
<h3> Notes:</h3>
When a vector is multiplied by a number, then its length is also multiplied with the same number.
But, we should be careful with the sign.
Consider the vector $ -3 v $. It has the same length of $ 3v $, but its direction is opposite.
So, when calculating the length of $ -3 v $, we use absolute value of the number:
$ \norm{-3 v} = |-3| \norm{v} = 3 \norm{v} $.
Here $ |-3| $ is the absolute value of $ -3 $.
The absolute value of a number is its distance to 0. So, $ |-3| = 3 $.
<h3> Task 4 </h3>
Let $ u = \myrvector{1 \\ -2 \\ -4 \\ 2} $ be a four dimensional vector.
Randomly pick a number $r$ from $ \left\{ \dfrac{1}{10}, \dfrac{2}{10}, \cdots, \dfrac{9}{10} \right\} $.
Find the vector $(-r)\cdot u$ and then its length.
```
#
# your solution is here
#
from random import randrange
u = [1,-2,-4,2]
print("u is",u)
r = randrange(9) # r is a number in {0,...,8}
r = r + 1 # r is a number in {1,...,9}
r = r/10 # r is a number in {1/10,...,9/10}
print()
print("r is",r)
newu=[]
for i in range(len(u)):
newu.append(-1*r*u[i])
print()
print("-ru is",newu)
print()
length = 0
for i in range(len(newu)):
length = length + newu[i]**2 # adding square of each number
print(newu[i],"->[square]->",newu[i]**2)
print()
print("the summation of squares is",length)
length = length**0.5 # taking square root
print("the length of",newu,"is",length)
```
Remark that:
The length of $ u $ is 5.
The length of $ (-r)u $ will be $ 5r $.
| github_jupyter |
```
import torch
from torch.autograd import Variable
import torch.functional as F
import torch.nn.functional as F
import numpy as np
import pandas as pd
from copy import deepcopy
from pprint import pprint
# Function defs
def tokenize(corpus : str) -> list:
tokens = []
for sentence in corpus:
tokens.append(sentence.split())
return tokens
def remove_stops(corpus, stop_words=["is", "a"]):
c = []
for line in corpus:
s = ""
for word in line.split():
if word not in stop_words:
s += word + " "
c.append(s.strip())
return c
def word2index(tokens):
vocabulary = []
for sentence in tokens:
for token in sentence:
if token not in vocabulary:
vocabulary.append(token)
word2idx = {w: idx for (idx, w) in enumerate(vocabulary)}
return word2idx
def generate_center_context_pair(tokens, window: int) -> dict:
pairs = dict()
for row in tokens:
for idx, center_word in enumerate(row):
pairs.setdefault(center_word, [])
for i in range(idx - window, idx + window + 1):
if (i >= 0 and i != idx and i < len(row)):
pairs[center_word].append(row[i])
return pairs
def get_idxpairs(cc_pair: dict, w2idx: list) -> list:
"""
The generate_center_context_pair gives a dictionary like:
{'center word 1': ['contextword1', 'contextword2', '...']
'centerword2': ['contextword1', 'contextword2', '...']}
But the code from the blog needs cc_pair like:
[['centerword1', 'contextword1'],
['centerword1', 'contextword2'], ...]
So this part changes from the former format to the latter
"""
idx_pairs = []
for center in cc_pair.keys():
for context in cc_pair[center]:
idx_pairs.append([w2idx[center], w2idx[context]])
return idx_pairs
def generate_jdt(cc_pair: dict) -> list:
jdt = []
for center in cc_pair.keys():
for context in cc_pair[center]:
jdt.append([center, context])
return jdt
def all_p_of_context_given_center(joint_distrib_table: pd.DataFrame):
counts = joint_distrib_table.groupby(['center', 'context']).size()
counts = counts.to_dict()
# Denominator for the probability
total = joint_distrib_table.groupby('center').size()
total = total.to_dict()
for center in total.keys():
for k in list(counts.keys()):
if k[0] is center:
counts[k] = [counts[k]]
counts[k].append(total[center])
return counts
corpus = [
"he is a king",
"she is a queen",
"he is a man",
"she is a woman",
"warsaw is poland capital",
"berlin is germany capital",
"paris is france capital",
# "Sxi este juna kaj bela",
]
def get_input_layer(word_idx, vocab_size):
x = torch.zeros(vocab_size).float()
x[word_idx] = 1.0
return x
def main():
tokens = tokenize(corpus)
vocabulary = set(sum(tokens, [])) # sum() flattens the 2d list
vocab_size = len(vocabulary)
cc_pair = generate_center_context_pair(tokens, 1)
# pprint(cc_pair)
word2idx = word2index(tokens)
idx2word = {key: val for (val, key) in word2idx.items()}
print(word2idx)
print(idx2word)
idx_pairs = get_idxpairs(cc_pair, word2idx)
idx_pairs = np.array(idx_pairs)
embedding_dims = 5
W1 = Variable(torch.randn(embedding_dims, vocab_size).float(),
requires_grad=True)
W2 = Variable(torch.randn(vocab_size, embedding_dims).float(),
requires_grad=True)
max_iter = 200
learning_rate = 0.001
for i in range(max_iter):
loss_val = 0
for data, target in idx_pairs:
x = Variable(get_input_layer(data, vocab_size)).float()
y_true = Variable(torch.from_numpy(np.array([target])).long())
z1 = torch.matmul(W1, x)
z2 = torch.matmul(W2, z1)
log_softmax = F.log_softmax(z2, dim=0)
loss = F.nll_loss(log_softmax.view(1, -1), y_true)
loss_val += loss.item()
loss.backward()
W1.data -= learning_rate * W1.grad.data
W2.data -= learning_rate * W2.grad.data
W1.grad.data.zero_()
W2.grad.data.zero_()
if i % 10 == 0:
print(f"Loss at iter {i}: {loss_val/len(idx_pairs)}")
# Lets see the word predictions for each word in our vocabulary
for word in vocabulary:
widx = word2idx[word]
x = Variable(get_input_layer(widx, vocab_size)).float()
z1 = torch.matmul(W1, x)
z2 = torch.matmul(W2, z1)
softmax = F.softmax(z2, dim=0)
max_arg = torch.argmax(softmax).item()
pred_word = idx2word[max_arg]
print(f"Center: {word} ; Context: {pred_word}")
if __name__ == "__main__":
main()
```
| github_jupyter |
# Perceptron with MaxAbsScaler & PowerTransformer
This Code template is for the Classification task using simple Perceptron with feature rescaling technique MaxAbsScaler and feature transformation technique PowerTransformer in a pipeline. Which is a simple classification algorithm suitable for large scale learning.
### Required Packages
```
!pip install imblearn
import warnings
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as se
from imblearn.over_sampling import RandomOverSampler
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import LabelEncoder, MaxAbsScaler, PowerTransformer
from sklearn.model_selection import train_test_split
from sklearn.linear_model import Perceptron
from sklearn.metrics import classification_report,plot_confusion_matrix
warnings.filterwarnings('ignore')
```
### Initialization
Filepath of CSV file
```
#filepath
file_path= ""
```
List of features which are required for model training .
```
#x_values
features=[]
```
Target feature for prediction.
```
#y_value
target=''
```
### Data Fetching
Pandas is an open-source, BSD-licensed library providing high-performance, easy-to-use data manipulation and data analysis tools.
We will use panda's library to read the CSV file using its storage path.And we use the head function to display the initial row or entry.
```
df=pd.read_csv(file_path)
df.head()
```
### Feature Selections
It is the process of reducing the number of input variables when developing a predictive model. Used to reduce the number of input variables to both reduce the computational cost of modelling and, in some cases, to improve the performance of the model.
We will assign all the required input features to X and target/outcome to Y.
```
X = df[features]
Y = df[target]
```
### Data Preprocessing
Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
```
def NullClearner(df):
if(isinstance(df, pd.Series) and (df.dtype in ["float64","int64"])):
df.fillna(df.mean(),inplace=True)
return df
elif(isinstance(df, pd.Series)):
df.fillna(df.mode()[0],inplace=True)
return df
else:return df
def EncodeX(df):
return pd.get_dummies(df)
def EncodeY(df):
if len(df.unique())<=2:
return df
else:
un_EncodedT=np.sort(pd.unique(df), axis=-1, kind='mergesort')
df=LabelEncoder().fit_transform(df)
EncodedT=[xi for xi in range(len(un_EncodedT))]
print("Encoded Target: {} to {}".format(un_EncodedT,EncodedT))
return df
x=X.columns.to_list()
for i in x:
X[i]=NullClearner(X[i])
X=EncodeX(X)
Y=EncodeY(NullClearner(Y))
X.head()
```
#### Correlation Map
In order to check the correlation between the features, we will plot a correlation matrix. It is effective in summarizing a large amount of data where the goal is to see patterns.
```
f,ax = plt.subplots(figsize=(18, 18))
matrix = np.triu(X.corr())
se.heatmap(X.corr(), annot=True, linewidths=.5, fmt= '.1f',ax=ax, mask=matrix)
plt.show()
```
#### Distribution Of Target Variable
```
plt.figure(figsize = (10,6))
se.countplot(Y)
```
### Data Splitting
The train-test split is a procedure for evaluating the performance of an algorithm. The procedure involves taking a dataset and dividing it into two subsets. The first subset is utilized to fit/train the model. The second subset is used for prediction. The main motive is to estimate the performance of the model on new data.
```
x_train,x_test,y_train,y_test=train_test_split(X,Y,test_size=0.2,random_state=123)
```
#### Handling Target Imbalance
The challenge of working with imbalanced datasets is that most machine learning techniques will ignore, and in turn have poor performance on, the minority class, although typically it is performance on the minority class that is most important.
One approach to addressing imbalanced datasets is to oversample the minority class. The simplest approach involves duplicating examples in the minority class.We will perform overspampling using imblearn library.
```
x_train,y_train = RandomOverSampler(random_state=123).fit_resample(x_train, y_train)
```
### Data Rescaling
Scale each feature by its maximum absolute value.
This estimator scales and translates each feature individually such that the maximal absolute value of each feature in the training set will be 1.0. It does not shift/center the data, and thus does not destroy any sparsity.
[More on MaxAbsScaler module and parameters](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.MaxAbsScaler.html)
### Feature Transformation
Power transforms are a family of parametric, monotonic transformations that are applied to make data more Gaussian-like. This is useful for modeling issues related to heteroscedasticity (non-constant variance), or other situations where normality is desired.
[More on PowerTransformer module and parameters](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.PowerTransformer.html)
### Model
the perceptron is an algorithm for supervised learning of binary classifiers.
The algorithm learns the weights for the input signals in order to draw a linear decision boundary.This enables you to distinguish between the two linearly separable classes +1 and -1.
#### Model Tuning Parameters
> **penalty** ->The penalty (aka regularization term) to be used. {‘l2’,’l1’,’elasticnet’}
> **alpha** -> Constant that multiplies the regularization term if regularization is used.
> **l1_ratio** -> The Elastic Net mixing parameter, with 0 <= l1_ratio <= 1. l1_ratio=0 corresponds to L2 penalty, l1_ratio=1 to L1. Only used if penalty='elasticnet'.
> **tol** -> The stopping criterion. If it is not None, the iterations will stop when (loss > previous_loss - tol).
> **early_stopping**-> Whether to use early stopping to terminate training when validation. score is not improving. If set to True, it will automatically set aside a stratified fraction of training data as validation and terminate training when validation score is not improving by at least tol for n_iter_no_change consecutive epochs.
> **validation_fraction** -> The proportion of training data to set aside as validation set for early stopping. Must be between 0 and 1. Only used if early_stopping is True.
> **n_iter_no_change** -> Number of iterations with no improvement to wait before early stopping.
```
# Build Model here
model = make_pipeline(MaxAbsScaler(), PowerTransformer(), Perceptron(random_state=123))
model.fit(x_train, y_train)
```
#### Model Accuracy
score() method return the mean accuracy on the given test data and labels.
In multi-label classification, this is the subset accuracy which is a harsh metric since you require for each sample that each label set be correctly predicted.
```
print("Accuracy score {:.2f} %\n".format(model.score(x_test,y_test)*100))
```
#### Confusion Matrix
A confusion matrix is utilized to understand the performance of the classification model or algorithm in machine learning for a given test set where results are known.
```
plot_confusion_matrix(model,x_test,y_test,cmap=plt.cm.Blues)
```
#### Classification Report
A Classification report is used to measure the quality of predictions from a classification algorithm. How many predictions are True, how many are False.
* **where**:
- Precision:- Accuracy of positive predictions.
- Recall:- Fraction of positives that were correctly identified.
- f1-score:- percent of positive predictions were correct
- support:- Support is the number of actual occurrences of the class in the specified dataset.
```
print(classification_report(y_test,model.predict(x_test)))
```
#### Creator: Nikhil Shrotri , Github: [Profile](https://github.com/nikhilshrotri)
| github_jupyter |
```
## Load the data
import pandas as pd
data = pd.read_csv('./data.csv')
## Clean the data
data.columns
data.drop(['sqft_living','sqft_lot','waterfront','view','condition','sqft_above','sqft_basement','street','city','statezip','country'],axis=1,inplace=True)
data.drop('date',axis=1,inplace=True)
data.head()
## Feature Enginnering
def fe(data,col):
print(len(data))
max_no = data[col].quantile(0.99)
min_no = data[col].quantile(0.05)
data = data[data[col] > min_no]
data = data[data[col] < max_no]
print(len(data))
return data
for col in list(data.columns):
print(col)
data = fe(data,'price')
data.head()
X = data.drop('price',axis=1)
y = data['price']
from sklearn.model_selection import train_test_split
X_train,X_test,y_train,y_test = train_test_split(X,y,test_size=0.25)
len(X_train),len(X_test)
## Modelling
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
class BaseLine_Model(nn.Module):
def __init__(self,input_shape,output_shape):
super().__init__()
self.fc1 = nn.Linear(input_shape,64)
self.fc2 = nn.Linear(64,128)
self.fc3 = nn.Linear(128,256)
self.fc4 = nn.Linear(256,128)
self.fc5 = nn.Linear(128,output_shape)
def forward(self,X):
preds = self.fc1(X)
preds = F.relu(preds)
preds = self.fc2(preds)
preds = F.relu(preds)
preds = self.fc3(preds)
preds = F.relu(preds)
preds = self.fc4(preds)
preds = F.relu(preds)
preds = self.fc5(preds)
return preds
EPOCHS = 250
import wandb
BATCH_SIZE = 32
PROJECT_NAME = 'House-Price-Pred'
from tqdm import tqdm
device = torch.device('cuda')
model = BaseLine_Model(5,1).to(device)
criterion = nn.MSELoss()
optimizer = optim.Adam(model.parameters(),lr=0.1)
def get_loss(criterion,X,y,model):
preds = model(X.float().to(device))
preds = preds.to(device)
y = y.to(device)
# criterion.to(device)
loss = criterion(preds,y)
return loss.item()
def get_accuracy(X,y,model):
correct = 0
total = 0
for i in range(len(X)):
pred = model(X[i].float().to(device))
pred.to(device)
if round(pred[0]) == round(y[i]):
correct += 1
total += 1
if correct == 0:
correct += 1
return round(correct/total,3)
import numpy as np
X_train = torch.from_numpy(np.array(X_train))
y_train = torch.from_numpy(np.array(y_train))
X_test = torch.from_numpy(np.array(X_test))
y_test = torch.from_numpy(np.array(y_test))
get_accuracy(X_test,y_test,model)
def get_loss(criterion,X,y,model):
preds = model(X.float().to(device))
preds = preds.to(device)
y = y.to(device)
# criterion.to(device)
loss = criterion(preds,y)
return loss.item()
def get_accuracy(X,y,model):
correct = 0
total = 0
for i in range(len(X)):
pred = model(X[i].float().to(device))
pred.to(device)
if round(int(pred[0])) == round(int(y[i])):
correct += 1
total += 1
if correct == 0:
correct += 1
return round(correct/total,3)
import numpy as np
X_train = torch.from_numpy(np.array(X_train))
y_train = torch.from_numpy(np.array(y_train))
X_test = torch.from_numpy(np.array(X_test))
y_test = torch.from_numpy(np.array(y_test))
get_accuracy(X_test,y_test,model)
wandb.init(project=PROJECT_NAME,name='baseline')
for _ in tqdm(range(EPOCHS)):
for i in range(0,len(X_train),BATCH_SIZE):
X_batch = X_train[i:i+BATCH_SIZE].to(device)
y_batch = y_train[i:i+BATCH_SIZE].to(device)
model.to(device)
preds = model(X_batch.float())
preds = preds.view(len(preds))
preds.to(device)
loss = criterion(preds.float(),y_batch.float())
optimizer.zero_grad()
loss.backward()
optimizer.step()
wandb.log({'loss':loss.item(),'val_loss':get_loss(criterion,X_test,y_test,model),'accuracy':get_accuracy(X_train,y_train,model),'val_accuracy':get_accuracy(X_test,y_test,model)})
preds
y_batch
torch.round(preds)
torch.round(y_batch)
EPOCHS = 50
EPOCHS = 25
wandb.init(project=PROJECT_NAME,name='baseline')
for _ in tqdm(range(EPOCHS)):
for i in range(0,len(X_train),BATCH_SIZE):
X_batch = X_train[i:i+BATCH_SIZE].to(device)
y_batch = y_train[i:i+BATCH_SIZE].to(device)
model.to(device)
preds = model(X_batch.float())
preds = preds.view(len(preds))
preds.to(device)
loss = criterion(preds.float(),y_batch.float())
optimizer.zero_grad()
loss.backward()
optimizer.step()
wandb.log({'loss':loss.item(),'val_loss':get_loss(criterion,X_test,y_test,model),'accuracy':get_accuracy(X_train,y_train,model),'val_accuracy':get_accuracy(X_test,y_test,model)})
EPOCHS = 50
preds
y_batch
torch.round(preds)
torch.round(y_batch)
wandb.init(project=PROJECT_NAME,name='baseline')
for _ in tqdm(range(EPOCHS)):
for i in range(0,len(X_train),BATCH_SIZE):
X_batch = X_train[i:i+BATCH_SIZE].to(device)
y_batch = y_train[i:i+BATCH_SIZE].to(device)
model.to(device)
preds = model(X_batch.float())
preds = preds.view(len(preds))
preds.to(device)
loss = criterion(preds.float(),y_batch.float())
optimizer.zero_grad()
loss.backward()
optimizer.step()
wandb.log({'loss':loss.item(),'val_loss':get_loss(criterion,X_test,y_test,model),'accuracy':get_accuracy(X_train,y_train,model),'val_accuracy':get_accuracy(X_test,y_test,model)})
preds
y_batch
torch.round(preds)
torch.round(y_batch)
EPOCHS = 75
wandb.init(project=PROJECT_NAME,name='baseline')
for _ in tqdm(range(EPOCHS)):
for i in range(0,len(X_train),BATCH_SIZE):
X_batch = X_train[i:i+BATCH_SIZE].to(device)
y_batch = y_train[i:i+BATCH_SIZE].to(device)
model.to(device)
preds = model(X_batch.float())
preds = preds.view(len(preds))
preds.to(device)
loss = criterion(preds.float(),y_batch.float())
optimizer.zero_grad()
loss.backward()
optimizer.step()
wandb.log({'loss':loss.item(),'val_loss':get_loss(criterion,X_test,y_test,model),'accuracy':get_accuracy(X_train,y_train,model),'val_accuracy':get_accuracy(X_test,y_test,model)})
```
| github_jupyter |
# Exercise Set 11: Basic classification models
*Morning, August 19, 2019*
In this Exercise Set you will get your first taste of how machine learning algorithms are constructed. You will implement a [_perceptron_](https://en.wikipedia.org/wiki/Perceptron) from scratch using the matrix-algebra library NumPy. We will train this model on the iris data to predict flower types.
Many of the concepts both programming-wise and related to machine learning are probably new to most of you - don't be afraid to ask questions about either, as much of this lecture/exercise set lays the foundation for the comming sessions.
We recommend that you do not begin the bonus exercises until everything else is complete! Especially 11.1.8 and 11.3.X are only for the extra curious.
> **Ex. 11.1.1:** The mathematics and biological reasoning which justifies the perceptron model is presented in Raschka, 2017 on pages 18 to 24. If you haven't read it already, quickly do so.
>
> Begin by importing `numpy`, `pandas` and `seaborn`
```
# [Answer to Ex. 11.1.1]
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
```
> **Ex. 11.1.2:** Use the following code snippet to load the iris data. The code will create two new variablex **X** and **y**, each of which are numpy arrays. Split the data as follows. The first dataset should contain the first 70 rows; we call this sample our *training dataset*, or simply *train data*. We use the training data to estimate the data. We use the remaining rows as data for testing our model, thus we call it *test data*.
>
>```python
iris = sns.load_dataset('iris')
iris = iris.query("species == 'virginica' | species == 'versicolor'").sample(frac=1, random_state = 3)
X = np.array(iris[['sepal_length', 'sepal_width', 'petal_length', 'petal_width']])
y = np.array(iris['species'].map({'virginica': 1, 'versicolor': -1}))
sns.pairplot(iris, hue="species", palette="husl", diag_kws = {'shade': False})
plt.show()
```
```
# [Answer to Ex. 11.1.2]
iris = sns.load_dataset('iris')
iris = iris.query("species == 'virginica' | species == 'versicolor'").sample(frac = 1, random_state = 3)
X = np.array(iris[['sepal_length', 'sepal_width', 'petal_length', 'petal_width']])
y = np.array(iris['species'].map({'virginica': 1, 'versicolor': -1}))
sns.pairplot(iris, hue="species", palette="husl", diag_kws = {'shade': False})
plt.show()
# A very simple deterministic test-train split
Xtrain = X[:70]
ytrain = y[:70]
Xtest = X[70:]
ytest = y[70:]
```
## The perceptron model
> **Ex. 11.1.3:** Write a function which initiate a set of weights `w` with length 1 larger than the number of features in your data. Ensure that your initial weights are not exactly 0, but close to it.
>
>> _Hint 1:_ Use [np.random.RandomState](https://docs.scipy.org/doc/numpy/reference/generated/numpy.random.RandomState.html) to set up a random number generator from which you can draw from a normal with mean 0 and scale 0.01.
>
>> _Hint 2:_ Say you have stored the random number generator in an object called `rgen`. You can then call `rgen.normal(size = 1 + columns_in_X)` to get the weights you want. You might want to tweak the `scale` parameter.
```
# [Answer to Ex. 11.1.3]
def random_weights(location = 0.0, scale = 0.01, seed = 1):
# Init random number generator
rgen = np.random.RandomState(seed)
w = rgen.normal(loc=location, scale=scale, size= 1 + X.shape[1])
return w
```
> **Ex. 11.1.4:** In this problem you need to write two functions:
> * `net_input(X, W)`: calculates _and returns_ the net-input, i.e the linear combination of features and weights, $z=w_0 + \sum_k x_{k} w_{k}$
> * `predict(X, W)`: a step function which returns 1 if the net activation is $\geq$ 0, and returns -1 otherwise.
>
>*Bonus:* Create a function which calculates the _accuracy_ (the share of cases that are correctly classified). The function should take a vector of y-values and a vector of predicted y-values as input. What is the accuracy of your untrained model on the training data?
>> _Hint 1:_ you can compute the above using an array product. Here numpy's array product named `dot` may be useful
>> _Hint 2:_ remember to include the bias, $w_0$, in the computation!
```
# [Answer to Ex. 11.1.4]
def net_input(X, W):
return np.dot(X, W[1:]) + W[0] # Linear product X'W + bias
def predict(X, W):
linProd = net_input(X, W)
return np.where(linProd >= 0.0, 1, -1) # 1(linProd > 0)
# Bonus
def accuracy(y, prediction):
return np.mean(y == prediction)
accuracy(ytrain, predict(Xtrain, random_weights()))
```
> **Ex. 11.1.5:** Write a function whichs loops over the training data (both X and y) using `zip`. For each row in the data, update the weights according to the perceptron rule (remember to update the bias in `w[0]`!). Set $\eta = 0.1$.
>
> Make sure the loop stores the total number of prediction errors encountered underways in the loop by creating an `int` which is incremented whenever you update the weights.
>
>> _Hint:_ your function should return the updated weights, as well as the number of errors made by the perceptron.
>
>> _Hint:_ The following code block implements the function in _pseudo_code (it wont run, but serves to communicate the functionality).
>> ```
>> function f(X, y, W, eta):
>> set errors = 0
>>
>> for each pair xi, yi in zip(X,y) do:
>> set update = eta * (yi - predict(xi, W))
>> set W[1:] = W[1:] + update * xi
>> set W[0] = W[0] + update
>> set errors = errors + int(update != 0)
>>
>> return W, errors
>> ```
>
> *Bonus:* If you completed the previous bonus exercise (for 11.1.4), calculate the accuracy on training data using the updated weights as input in the predict function. Any progress yet?
```
# [Answer to Ex. 11.1.5]
# This will be in assignment 2
```
> **Ex. 11.1.6:** Write a function which repeat the updating procedure (calls the function) you constructed in 11.1.5 for `n_iter` times by packing the whole thing in a loop. Make sure you store the number of errors in each iteration in a list.
>
> Plot the total errors after each iteration in a graph.
>
>> _Hint:_ Make sure you dont reset the weights after each iteration.
>
>> _Hint:_ Once again some pseudocode:
>> ```
>> function g(X, y, n_iter):
>> set eta = 0.1
>> set weights = random_weights()
>> set errorseq = list()
>>
>> for each _ in range(n_iter):
>> weights, e = f(X, y, W, eta)
>> errorseq.append(e)
>>
>> return weights, errorseq
>> ```
```
# [Answer to Ex. 11.1.6]
# This will be in assignment 2
```
> **Ex. 11.1.7 (BONUS):** Use the updated weights when predicting and calculate the accuracy of your perceptron on the test data?
```
# [Answer to Ex. 11.1.7 BONUS]
pred = predict(Xtest, trained_w)
accuracy(ytest, pred)
```
> **Ex.11.1.8 (BONUS):** Restructure your code as a class called `Perceptron` with `.fit()` and `.predict()` methods (you) will probably need more helper methods. Store hyperparameters as eta and the number of iterations as class attributes.
```
# [Answer to Ex. 11.1.8 BONUS]
# This will NOT be in assignment 2, but the answer is hidden, as it would reveal
# how to solve ex. 11.1.5 and 11.1.6
p = Perceptron(X = Xtrain, y= ytrain).fit()
plt.plot(p._errseq, 'b-o')
```
## Logistic regression
Logistic regression is another simple linear machine-learning algorithm, you can read about it [here:](https://scikit-learn.org/stable/modules/linear_model.html#logistic-regression)
> **Ex. 11.2.1:** Import the LogisticRegression classifier from `sklearn.linear_model`. Create a new object called `clf` like:
```
clf = LogisticRegression()
```
All scikit learn models have two fundamental methods `.fit()` and `.predict()`. Fit your model to the training data, and store the fitted model in a new object. Import _accuracy_score_ from `sklearn.metrics` and asses the accuracy of the LogisticRegression on both your training data and your test data.
```
# [Answer to Ex. 11.2.1]
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score
clf = LogisticRegression(solver='lbfgs')
fitted_model = clf.fit(Xtrain, ytrain)
train_score = accuracy_score(ytrain, fitted_model.predict(Xtrain))
test_score = accuracy_score(ytest, fitted_model.predict(Xtest))
print(f"On the training data we get a score of {round(train_score, 2)}, while the score on the test data is {round(test_score, 2)}")
```
## AdaLine (BONUS)
AdaLine is a modified version of the perceptron. The only difference lies in the way the two models learn from their training data, i.e. the optimization method used. The perceptron used the binary classifications for learning, while AdaLine only applies the binary threshold after training, and thus uses real valued numbers when learning.
>> _Hint:_ Most of the code for this exercise can be written by copying and modifying code from exercise 11.1.
> **Ex. 11.3.1 (BONUS):** Implement two functions described below. You shold reuse your `net_input` from Ex. 11.1.4.:
* `ada_activation_function`: the identify function $ada\_activation(z) = z$
* `ada_predict`: A step function $ada\_predict(z) = 1 \ if \ z \geq 0 \ else \ 0$ where z is the output of _the activation function_.
> The following figure might help you understand how each of these functions relate to the algorithm, and how the perceptron and adaline differ:
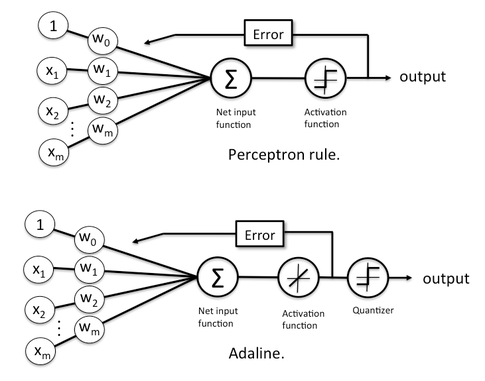
```
# [Answer to Ex. 11.3.1 BONUS]
def ada_activation(Z):
return Z
def ada_predict(X, W):
linProd = net_input(X, W)
act = ada_activation(linprod)
return np.where(act >= 0.0, 1, -1) # 1(linProd > 0)
```
> **Ex. 11.3.2 (BONUS):** AdaLine uses a _cost function_ to quantize the accuracy of the classifier this is given by
>$$
cost(X,y,W) = \frac{1}{2} \sum_{i=1}^N (y_i - activation(z_i) )^2 , \qquad z_i = net\_input(x_i, W)
$$
> If you've followed any normal undergraduate courses in statistics you should recognize this function. Begin by implementing the cost function. Unlike in undergraduate statistics we will optimize our estimator using gradient descent, therefore **code up the negative of the derivative of the cost function as well**.
> $$
-cost'_j(X,y, W) = -\sum_{i=1}^N (y_i - activation(z_i)) x_i^j, \qquad z_i = net\_input(x_i, W)
$$
>
>> _Hint:_ Dont compute the sum for each weight $w_j$, instead use numpy's matrix algebra to compute the all of the derivatives at once.
>
>> _Hint:_ The derivative should return a list of the same length as the number of weights, since there is one derivative for each one.
```
# [Answer to Ex. 11.3.2 BONUS]
def ada_cost(X, y, W):
linProd = net_input(X, W)
errors_sq = (y - ada_activation(linProd))**2
return errors_sq.sum() / 2.0
def ada_cost_derivative(X, y, W):
linProd = net_input(X, W)
errors = y - ada_activation(linProd)
return np.array( [errors.sum()] + list(X.T.dot(errors)))
ada_cost_derivative(Xtrain, ytrain, random_weights())
```
> **Ex. 11.3.3 BONUS:** Implement the adaline fitting algorithm using *batch gradient descent*. This is similar to what you did with the perceptron, but while the perceptron did it's optimization after evaluating each row in the dataset, adaline treats the entire dataset as a batch, adjusts it's weights and then does it all again. Thus you only need to loop over `n_iter`, _not_ the data rows. Use the cost function to track the progress of your algorithm.
>
>> _Hint:_ gradient descent will be extremely sensitive to the learning rate $\eta$ in this situation - try setting i to 0.0001 and running the algorithm for 5000 iterations to get some kind of convergence.
```
# [Answer to ex. 11.3.3 BONUS]
def AdaLine(X, y, n_iter = 10000, eta = 0.00001):
costseq = []
W = random_weights()
for i in range(n_iter):
nip = net_input(X, W)
output = ada_activation(nip)
W = W + eta * ada_cost_derivative(X, y, W)
costseq.append(ada_cost(X,y, W))
return W, costseq
w_trained, costs = AdaLine(Xtrain, ytrain)
plt.plot(costs)
```
> **Ex. 11.3.4 (BONUS):** Write a function that scales each of the variables in the dataset (including **y**) using the formula
$$
x_j^{new} = \frac{x_j^{old} - \mu_j}{\sigma_j}
$$
> rerun the adaline function on the scaled variables.
```
# [Answer to Ex. 11.3.4]
def standardScaler(X, y):
""" Scales the input. (Horrible code)
"""
X_new = X.copy()
for i in range(X.shape[1]):
xj = X[:,i]
stdev = np.std(xj)
mean = np.mean(xj)
X_new[:,i] = (xj - mean)/stdev
y_stdev = np.std(y)
y_mean = np.mean(y)
y_new = (y.copy() - y_mean)/y_stdev
return X_new, y_new
X_scaled, y_scaled = standardScaler(Xtrain,ytrain)
w_trained, costs = AdaLine(X_scaled, y_scaled)
plt.plot(costs)
```
| github_jupyter |
##### Copyright 2018 The TensorFlow Hub Authors.
Licensed under the Apache License, Version 2.0 (the "License");
```
# Copyright 2018 The TensorFlow Hub Authors. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
```
# BigGAN Demo
This notebook is a demo for the *BigGAN* image generators available on [TF Hub](https://tfhub.dev/s?publisher=deepmind&q=biggan).
See the [BigGAN paper on arXiv](https://arxiv.org/abs/1809.11096) [1] for more information about these models.
After connecting to a runtime, get started by following these instructions:
1. (Optional) Update the selected **`module_path`** in the first code cell below to load a BigGAN generator for a different image resolution.
2. Click **Runtime > Run all** to run each cell in order.
* Afterwards, the interactive visualizations should update automatically when you modify the settings using the sliders and dropdown menus.
* If not, press the **Play** button by the cell to re-render outputs manually.
Note: if you run into any issues, it can help to click **Runtime > Restart and run all...** to restart your runtime and rerun all cells from scratch.
[1] Andrew Brock, Jeff Donahue, and Karen Simonyan. [Large Scale GAN Training for High Fidelity Natural Image Synthesis](https://arxiv.org/abs/1809.11096). *arxiv:1809.11096*, 2018.
First, set the module path.
By default, we load the BigGAN-deep generator for 256x256 images from **`https://tfhub.dev/deepmind/biggan-deep-256/1`**.
To generate 128x128 or 512x512 images or to use the original BigGAN generators, comment out the active **`module_path`** setting and uncomment one of the others.
```
# BigGAN-deep models
# module_path = 'https://tfhub.dev/deepmind/biggan-deep-128/1' # 128x128 BigGAN-deep
module_path = 'https://tfhub.dev/deepmind/biggan-deep-256/1' # 256x256 BigGAN-deep
# module_path = 'https://tfhub.dev/deepmind/biggan-deep-512/1' # 512x512 BigGAN-deep
# BigGAN (original) models
# module_path = 'https://tfhub.dev/deepmind/biggan-128/2' # 128x128 BigGAN
# module_path = 'https://tfhub.dev/deepmind/biggan-256/2' # 256x256 BigGAN
# module_path = 'https://tfhub.dev/deepmind/biggan-512/2' # 512x512 BigGAN
```
# Setup
```
import io
import IPython.display
import numpy as np
import PIL.Image
from scipy.stats import truncnorm
import tensorflow as tf
import tensorflow_hub as hub
```
## Load a BigGAN generator module from TF Hub
```
tf.reset_default_graph()
print('Loading BigGAN module from:', module_path)
module = hub.Module(module_path)
inputs = {k: tf.placeholder(v.dtype, v.get_shape().as_list(), k)
for k, v in module.get_input_info_dict().items()}
output = module(inputs)
print()
print('Inputs:\n', '\n'.join(
' {}: {}'.format(*kv) for kv in inputs.items()))
print()
print('Output:', output)
```
## Define some functions for sampling and displaying BigGAN images
```
input_z = inputs['z']
input_y = inputs['y']
input_trunc = inputs['truncation']
dim_z = input_z.shape.as_list()[1]
vocab_size = input_y.shape.as_list()[1]
def truncated_z_sample(batch_size, truncation=1., seed=None):
state = None if seed is None else np.random.RandomState(seed)
values = truncnorm.rvs(-2, 2, size=(batch_size, dim_z), random_state=state)
return truncation * values
def one_hot(index, vocab_size=vocab_size):
index = np.asarray(index)
if len(index.shape) == 0:
index = np.asarray([index])
assert len(index.shape) == 1
num = index.shape[0]
output = np.zeros((num, vocab_size), dtype=np.float32)
output[np.arange(num), index] = 1
return output
def one_hot_if_needed(label, vocab_size=vocab_size):
label = np.asarray(label)
if len(label.shape) <= 1:
label = one_hot(label, vocab_size)
assert len(label.shape) == 2
return label
def sample(sess, noise, label, truncation=1., batch_size=8,
vocab_size=vocab_size):
noise = np.asarray(noise)
label = np.asarray(label)
num = noise.shape[0]
if len(label.shape) == 0:
label = np.asarray([label] * num)
if label.shape[0] != num:
raise ValueError('Got # noise samples ({}) != # label samples ({})'
.format(noise.shape[0], label.shape[0]))
label = one_hot_if_needed(label, vocab_size)
ims = []
for batch_start in range(0, num, batch_size):
s = slice(batch_start, min(num, batch_start + batch_size))
feed_dict = {input_z: noise[s], input_y: label[s], input_trunc: truncation}
ims.append(sess.run(output, feed_dict=feed_dict))
ims = np.concatenate(ims, axis=0)
assert ims.shape[0] == num
ims = np.clip(((ims + 1) / 2.0) * 256, 0, 255)
ims = np.uint8(ims)
return ims
def interpolate(A, B, num_interps):
if A.shape != B.shape:
raise ValueError('A and B must have the same shape to interpolate.')
alphas = np.linspace(0, 1, num_interps)
return np.array([(1-a)*A + a*B for a in alphas])
def imgrid(imarray, cols=5, pad=1):
if imarray.dtype != np.uint8:
raise ValueError('imgrid input imarray must be uint8')
pad = int(pad)
assert pad >= 0
cols = int(cols)
assert cols >= 1
N, H, W, C = imarray.shape
rows = N // cols + int(N % cols != 0)
batch_pad = rows * cols - N
assert batch_pad >= 0
post_pad = [batch_pad, pad, pad, 0]
pad_arg = [[0, p] for p in post_pad]
imarray = np.pad(imarray, pad_arg, 'constant', constant_values=255)
H += pad
W += pad
grid = (imarray
.reshape(rows, cols, H, W, C)
.transpose(0, 2, 1, 3, 4)
.reshape(rows*H, cols*W, C))
if pad:
grid = grid[:-pad, :-pad]
return grid
def imshow(a, format='png', jpeg_fallback=True):
a = np.asarray(a, dtype=np.uint8)
data = io.BytesIO()
PIL.Image.fromarray(a).save(data, format)
im_data = data.getvalue()
try:
disp = IPython.display.display(IPython.display.Image(im_data))
except IOError:
if jpeg_fallback and format != 'jpeg':
print(('Warning: image was too large to display in format "{}"; '
'trying jpeg instead.').format(format))
return imshow(a, format='jpeg')
else:
raise
return disp
```
## Create a TensorFlow session and initialize variables
```
initializer = tf.global_variables_initializer()
sess = tf.Session()
sess.run(initializer)
```
# Explore BigGAN samples of a particular category
Try varying the **`truncation`** value.
(Double-click on the cell to view code.)
```
#@title Category-conditional sampling { display-mode: "form", run: "auto" }
num_samples = 10 #@param {type:"slider", min:1, max:20, step:1}
truncation = 0.4 #@param {type:"slider", min:0.02, max:1, step:0.02}
noise_seed = 0 #@param {type:"slider", min:0, max:100, step:1}
category = "933) cheeseburger" #@param ["0) tench, Tinca tinca", "1) goldfish, Carassius auratus", "2) great white shark, white shark, man-eater, man-eating shark, Carcharodon carcharias", "3) tiger shark, Galeocerdo cuvieri", "4) hammerhead, hammerhead shark", "5) electric ray, crampfish, numbfish, torpedo", "6) stingray", "7) cock", "8) hen", "9) ostrich, Struthio camelus", "10) brambling, Fringilla montifringilla", "11) goldfinch, Carduelis carduelis", "12) house finch, linnet, Carpodacus mexicanus", "13) junco, snowbird", "14) indigo bunting, indigo finch, indigo bird, Passerina cyanea", "15) robin, American robin, Turdus migratorius", "16) bulbul", "17) jay", "18) magpie", "19) chickadee", "20) water ouzel, dipper", "21) kite", "22) bald eagle, American eagle, Haliaeetus leucocephalus", "23) vulture", "24) great grey owl, great gray owl, Strix nebulosa", "25) European fire salamander, Salamandra salamandra", "26) common newt, Triturus vulgaris", "27) eft", "28) spotted salamander, Ambystoma maculatum", "29) axolotl, mud puppy, Ambystoma mexicanum", "30) bullfrog, Rana catesbeiana", "31) tree frog, tree-frog", "32) tailed frog, bell toad, ribbed toad, tailed toad, Ascaphus trui", "33) loggerhead, loggerhead turtle, Caretta caretta", "34) leatherback turtle, leatherback, leathery turtle, Dermochelys coriacea", "35) mud turtle", "36) terrapin", "37) box turtle, box tortoise", "38) banded gecko", "39) common iguana, iguana, Iguana iguana", "40) American chameleon, anole, Anolis carolinensis", "41) whiptail, whiptail lizard", "42) agama", "43) frilled lizard, Chlamydosaurus kingi", "44) alligator lizard", "45) Gila monster, Heloderma suspectum", "46) green lizard, Lacerta viridis", "47) African chameleon, Chamaeleo chamaeleon", "48) Komodo dragon, Komodo lizard, dragon lizard, giant lizard, Varanus komodoensis", "49) African crocodile, Nile crocodile, Crocodylus niloticus", "50) American alligator, Alligator mississipiensis", "51) triceratops", "52) thunder snake, worm snake, Carphophis amoenus", "53) ringneck snake, ring-necked snake, ring snake", "54) hognose snake, puff adder, sand viper", "55) green snake, grass snake", "56) king snake, kingsnake", "57) garter snake, grass snake", "58) water snake", "59) vine snake", "60) night snake, Hypsiglena torquata", "61) boa constrictor, Constrictor constrictor", "62) rock python, rock snake, Python sebae", "63) Indian cobra, Naja naja", "64) green mamba", "65) sea snake", "66) horned viper, cerastes, sand viper, horned asp, Cerastes cornutus", "67) diamondback, diamondback rattlesnake, Crotalus adamanteus", "68) sidewinder, horned rattlesnake, Crotalus cerastes", "69) trilobite", "70) harvestman, daddy longlegs, Phalangium opilio", "71) scorpion", "72) black and gold garden spider, Argiope aurantia", "73) barn spider, Araneus cavaticus", "74) garden spider, Aranea diademata", "75) black widow, Latrodectus mactans", "76) tarantula", "77) wolf spider, hunting spider", "78) tick", "79) centipede", "80) black grouse", "81) ptarmigan", "82) ruffed grouse, partridge, Bonasa umbellus", "83) prairie chicken, prairie grouse, prairie fowl", "84) peacock", "85) quail", "86) partridge", "87) African grey, African gray, Psittacus erithacus", "88) macaw", "89) sulphur-crested cockatoo, Kakatoe galerita, Cacatua galerita", "90) lorikeet", "91) coucal", "92) bee eater", "93) hornbill", "94) hummingbird", "95) jacamar", "96) toucan", "97) drake", "98) red-breasted merganser, Mergus serrator", "99) goose", "100) black swan, Cygnus atratus", "101) tusker", "102) echidna, spiny anteater, anteater", "103) platypus, duckbill, duckbilled platypus, duck-billed platypus, Ornithorhynchus anatinus", "104) wallaby, brush kangaroo", "105) koala, koala bear, kangaroo bear, native bear, Phascolarctos cinereus", "106) wombat", "107) jellyfish", "108) sea anemone, anemone", "109) brain coral", "110) flatworm, platyhelminth", "111) nematode, nematode worm, roundworm", "112) conch", "113) snail", "114) slug", "115) sea slug, nudibranch", "116) chiton, coat-of-mail shell, sea cradle, polyplacophore", "117) chambered nautilus, pearly nautilus, nautilus", "118) Dungeness crab, Cancer magister", "119) rock crab, Cancer irroratus", "120) fiddler crab", "121) king crab, Alaska crab, Alaskan king crab, Alaska king crab, Paralithodes camtschatica", "122) American lobster, Northern lobster, Maine lobster, Homarus americanus", "123) spiny lobster, langouste, rock lobster, crawfish, crayfish, sea crawfish", "124) crayfish, crawfish, crawdad, crawdaddy", "125) hermit crab", "126) isopod", "127) white stork, Ciconia ciconia", "128) black stork, Ciconia nigra", "129) spoonbill", "130) flamingo", "131) little blue heron, Egretta caerulea", "132) American egret, great white heron, Egretta albus", "133) bittern", "134) crane", "135) limpkin, Aramus pictus", "136) European gallinule, Porphyrio porphyrio", "137) American coot, marsh hen, mud hen, water hen, Fulica americana", "138) bustard", "139) ruddy turnstone, Arenaria interpres", "140) red-backed sandpiper, dunlin, Erolia alpina", "141) redshank, Tringa totanus", "142) dowitcher", "143) oystercatcher, oyster catcher", "144) pelican", "145) king penguin, Aptenodytes patagonica", "146) albatross, mollymawk", "147) grey whale, gray whale, devilfish, Eschrichtius gibbosus, Eschrichtius robustus", "148) killer whale, killer, orca, grampus, sea wolf, Orcinus orca", "149) dugong, Dugong dugon", "150) sea lion", "151) Chihuahua", "152) Japanese spaniel", "153) Maltese dog, Maltese terrier, Maltese", "154) Pekinese, Pekingese, Peke", "155) Shih-Tzu", "156) Blenheim spaniel", "157) papillon", "158) toy terrier", "159) Rhodesian ridgeback", "160) Afghan hound, Afghan", "161) basset, basset hound", "162) beagle", "163) bloodhound, sleuthhound", "164) bluetick", "165) black-and-tan coonhound", "166) Walker hound, Walker foxhound", "167) English foxhound", "168) redbone", "169) borzoi, Russian wolfhound", "170) Irish wolfhound", "171) Italian greyhound", "172) whippet", "173) Ibizan hound, Ibizan Podenco", "174) Norwegian elkhound, elkhound", "175) otterhound, otter hound", "176) Saluki, gazelle hound", "177) Scottish deerhound, deerhound", "178) Weimaraner", "179) Staffordshire bullterrier, Staffordshire bull terrier", "180) American Staffordshire terrier, Staffordshire terrier, American pit bull terrier, pit bull terrier", "181) Bedlington terrier", "182) Border terrier", "183) Kerry blue terrier", "184) Irish terrier", "185) Norfolk terrier", "186) Norwich terrier", "187) Yorkshire terrier", "188) wire-haired fox terrier", "189) Lakeland terrier", "190) Sealyham terrier, Sealyham", "191) Airedale, Airedale terrier", "192) cairn, cairn terrier", "193) Australian terrier", "194) Dandie Dinmont, Dandie Dinmont terrier", "195) Boston bull, Boston terrier", "196) miniature schnauzer", "197) giant schnauzer", "198) standard schnauzer", "199) Scotch terrier, Scottish terrier, Scottie", "200) Tibetan terrier, chrysanthemum dog", "201) silky terrier, Sydney silky", "202) soft-coated wheaten terrier", "203) West Highland white terrier", "204) Lhasa, Lhasa apso", "205) flat-coated retriever", "206) curly-coated retriever", "207) golden retriever", "208) Labrador retriever", "209) Chesapeake Bay retriever", "210) German short-haired pointer", "211) vizsla, Hungarian pointer", "212) English setter", "213) Irish setter, red setter", "214) Gordon setter", "215) Brittany spaniel", "216) clumber, clumber spaniel", "217) English springer, English springer spaniel", "218) Welsh springer spaniel", "219) cocker spaniel, English cocker spaniel, cocker", "220) Sussex spaniel", "221) Irish water spaniel", "222) kuvasz", "223) schipperke", "224) groenendael", "225) malinois", "226) briard", "227) kelpie", "228) komondor", "229) Old English sheepdog, bobtail", "230) Shetland sheepdog, Shetland sheep dog, Shetland", "231) collie", "232) Border collie", "233) Bouvier des Flandres, Bouviers des Flandres", "234) Rottweiler", "235) German shepherd, German shepherd dog, German police dog, alsatian", "236) Doberman, Doberman pinscher", "237) miniature pinscher", "238) Greater Swiss Mountain dog", "239) Bernese mountain dog", "240) Appenzeller", "241) EntleBucher", "242) boxer", "243) bull mastiff", "244) Tibetan mastiff", "245) French bulldog", "246) Great Dane", "247) Saint Bernard, St Bernard", "248) Eskimo dog, husky", "249) malamute, malemute, Alaskan malamute", "250) Siberian husky", "251) dalmatian, coach dog, carriage dog", "252) affenpinscher, monkey pinscher, monkey dog", "253) basenji", "254) pug, pug-dog", "255) Leonberg", "256) Newfoundland, Newfoundland dog", "257) Great Pyrenees", "258) Samoyed, Samoyede", "259) Pomeranian", "260) chow, chow chow", "261) keeshond", "262) Brabancon griffon", "263) Pembroke, Pembroke Welsh corgi", "264) Cardigan, Cardigan Welsh corgi", "265) toy poodle", "266) miniature poodle", "267) standard poodle", "268) Mexican hairless", "269) timber wolf, grey wolf, gray wolf, Canis lupus", "270) white wolf, Arctic wolf, Canis lupus tundrarum", "271) red wolf, maned wolf, Canis rufus, Canis niger", "272) coyote, prairie wolf, brush wolf, Canis latrans", "273) dingo, warrigal, warragal, Canis dingo", "274) dhole, Cuon alpinus", "275) African hunting dog, hyena dog, Cape hunting dog, Lycaon pictus", "276) hyena, hyaena", "277) red fox, Vulpes vulpes", "278) kit fox, Vulpes macrotis", "279) Arctic fox, white fox, Alopex lagopus", "280) grey fox, gray fox, Urocyon cinereoargenteus", "281) tabby, tabby cat", "282) tiger cat", "283) Persian cat", "284) Siamese cat, Siamese", "285) Egyptian cat", "286) cougar, puma, catamount, mountain lion, painter, panther, Felis concolor", "287) lynx, catamount", "288) leopard, Panthera pardus", "289) snow leopard, ounce, Panthera uncia", "290) jaguar, panther, Panthera onca, Felis onca", "291) lion, king of beasts, Panthera leo", "292) tiger, Panthera tigris", "293) cheetah, chetah, Acinonyx jubatus", "294) brown bear, bruin, Ursus arctos", "295) American black bear, black bear, Ursus americanus, Euarctos americanus", "296) ice bear, polar bear, Ursus Maritimus, Thalarctos maritimus", "297) sloth bear, Melursus ursinus, Ursus ursinus", "298) mongoose", "299) meerkat, mierkat", "300) tiger beetle", "301) ladybug, ladybeetle, lady beetle, ladybird, ladybird beetle", "302) ground beetle, carabid beetle", "303) long-horned beetle, longicorn, longicorn beetle", "304) leaf beetle, chrysomelid", "305) dung beetle", "306) rhinoceros beetle", "307) weevil", "308) fly", "309) bee", "310) ant, emmet, pismire", "311) grasshopper, hopper", "312) cricket", "313) walking stick, walkingstick, stick insect", "314) cockroach, roach", "315) mantis, mantid", "316) cicada, cicala", "317) leafhopper", "318) lacewing, lacewing fly", "319) dragonfly, darning needle, devil's darning needle, sewing needle, snake feeder, snake doctor, mosquito hawk, skeeter hawk", "320) damselfly", "321) admiral", "322) ringlet, ringlet butterfly", "323) monarch, monarch butterfly, milkweed butterfly, Danaus plexippus", "324) cabbage butterfly", "325) sulphur butterfly, sulfur butterfly", "326) lycaenid, lycaenid butterfly", "327) starfish, sea star", "328) sea urchin", "329) sea cucumber, holothurian", "330) wood rabbit, cottontail, cottontail rabbit", "331) hare", "332) Angora, Angora rabbit", "333) hamster", "334) porcupine, hedgehog", "335) fox squirrel, eastern fox squirrel, Sciurus niger", "336) marmot", "337) beaver", "338) guinea pig, Cavia cobaya", "339) sorrel", "340) zebra", "341) hog, pig, grunter, squealer, Sus scrofa", "342) wild boar, boar, Sus scrofa", "343) warthog", "344) hippopotamus, hippo, river horse, Hippopotamus amphibius", "345) ox", "346) water buffalo, water ox, Asiatic buffalo, Bubalus bubalis", "347) bison", "348) ram, tup", "349) bighorn, bighorn sheep, cimarron, Rocky Mountain bighorn, Rocky Mountain sheep, Ovis canadensis", "350) ibex, Capra ibex", "351) hartebeest", "352) impala, Aepyceros melampus", "353) gazelle", "354) Arabian camel, dromedary, Camelus dromedarius", "355) llama", "356) weasel", "357) mink", "358) polecat, fitch, foulmart, foumart, Mustela putorius", "359) black-footed ferret, ferret, Mustela nigripes", "360) otter", "361) skunk, polecat, wood pussy", "362) badger", "363) armadillo", "364) three-toed sloth, ai, Bradypus tridactylus", "365) orangutan, orang, orangutang, Pongo pygmaeus", "366) gorilla, Gorilla gorilla", "367) chimpanzee, chimp, Pan troglodytes", "368) gibbon, Hylobates lar", "369) siamang, Hylobates syndactylus, Symphalangus syndactylus", "370) guenon, guenon monkey", "371) patas, hussar monkey, Erythrocebus patas", "372) baboon", "373) macaque", "374) langur", "375) colobus, colobus monkey", "376) proboscis monkey, Nasalis larvatus", "377) marmoset", "378) capuchin, ringtail, Cebus capucinus", "379) howler monkey, howler", "380) titi, titi monkey", "381) spider monkey, Ateles geoffroyi", "382) squirrel monkey, Saimiri sciureus", "383) Madagascar cat, ring-tailed lemur, Lemur catta", "384) indri, indris, Indri indri, Indri brevicaudatus", "385) Indian elephant, Elephas maximus", "386) African elephant, Loxodonta africana", "387) lesser panda, red panda, panda, bear cat, cat bear, Ailurus fulgens", "388) giant panda, panda, panda bear, coon bear, Ailuropoda melanoleuca", "389) barracouta, snoek", "390) eel", "391) coho, cohoe, coho salmon, blue jack, silver salmon, Oncorhynchus kisutch", "392) rock beauty, Holocanthus tricolor", "393) anemone fish", "394) sturgeon", "395) gar, garfish, garpike, billfish, Lepisosteus osseus", "396) lionfish", "397) puffer, pufferfish, blowfish, globefish", "398) abacus", "399) abaya", "400) academic gown, academic robe, judge's robe", "401) accordion, piano accordion, squeeze box", "402) acoustic guitar", "403) aircraft carrier, carrier, flattop, attack aircraft carrier", "404) airliner", "405) airship, dirigible", "406) altar", "407) ambulance", "408) amphibian, amphibious vehicle", "409) analog clock", "410) apiary, bee house", "411) apron", "412) ashcan, trash can, garbage can, wastebin, ash bin, ash-bin, ashbin, dustbin, trash barrel, trash bin", "413) assault rifle, assault gun", "414) backpack, back pack, knapsack, packsack, rucksack, haversack", "415) bakery, bakeshop, bakehouse", "416) balance beam, beam", "417) balloon", "418) ballpoint, ballpoint pen, ballpen, Biro", "419) Band Aid", "420) banjo", "421) bannister, banister, balustrade, balusters, handrail", "422) barbell", "423) barber chair", "424) barbershop", "425) barn", "426) barometer", "427) barrel, cask", "428) barrow, garden cart, lawn cart, wheelbarrow", "429) baseball", "430) basketball", "431) bassinet", "432) bassoon", "433) bathing cap, swimming cap", "434) bath towel", "435) bathtub, bathing tub, bath, tub", "436) beach wagon, station wagon, wagon, estate car, beach waggon, station waggon, waggon", "437) beacon, lighthouse, beacon light, pharos", "438) beaker", "439) bearskin, busby, shako", "440) beer bottle", "441) beer glass", "442) bell cote, bell cot", "443) bib", "444) bicycle-built-for-two, tandem bicycle, tandem", "445) bikini, two-piece", "446) binder, ring-binder", "447) binoculars, field glasses, opera glasses", "448) birdhouse", "449) boathouse", "450) bobsled, bobsleigh, bob", "451) bolo tie, bolo, bola tie, bola", "452) bonnet, poke bonnet", "453) bookcase", "454) bookshop, bookstore, bookstall", "455) bottlecap", "456) bow", "457) bow tie, bow-tie, bowtie", "458) brass, memorial tablet, plaque", "459) brassiere, bra, bandeau", "460) breakwater, groin, groyne, mole, bulwark, seawall, jetty", "461) breastplate, aegis, egis", "462) broom", "463) bucket, pail", "464) buckle", "465) bulletproof vest", "466) bullet train, bullet", "467) butcher shop, meat market", "468) cab, hack, taxi, taxicab", "469) caldron, cauldron", "470) candle, taper, wax light", "471) cannon", "472) canoe", "473) can opener, tin opener", "474) cardigan", "475) car mirror", "476) carousel, carrousel, merry-go-round, roundabout, whirligig", "477) carpenter's kit, tool kit", "478) carton", "479) car wheel", "480) cash machine, cash dispenser, automated teller machine, automatic teller machine, automated teller, automatic teller, ATM", "481) cassette", "482) cassette player", "483) castle", "484) catamaran", "485) CD player", "486) cello, violoncello", "487) cellular telephone, cellular phone, cellphone, cell, mobile phone", "488) chain", "489) chainlink fence", "490) chain mail, ring mail, mail, chain armor, chain armour, ring armor, ring armour", "491) chain saw, chainsaw", "492) chest", "493) chiffonier, commode", "494) chime, bell, gong", "495) china cabinet, china closet", "496) Christmas stocking", "497) church, church building", "498) cinema, movie theater, movie theatre, movie house, picture palace", "499) cleaver, meat cleaver, chopper", "500) cliff dwelling", "501) cloak", "502) clog, geta, patten, sabot", "503) cocktail shaker", "504) coffee mug", "505) coffeepot", "506) coil, spiral, volute, whorl, helix", "507) combination lock", "508) computer keyboard, keypad", "509) confectionery, confectionary, candy store", "510) container ship, containership, container vessel", "511) convertible", "512) corkscrew, bottle screw", "513) cornet, horn, trumpet, trump", "514) cowboy boot", "515) cowboy hat, ten-gallon hat", "516) cradle", "517) crane", "518) crash helmet", "519) crate", "520) crib, cot", "521) Crock Pot", "522) croquet ball", "523) crutch", "524) cuirass", "525) dam, dike, dyke", "526) desk", "527) desktop computer", "528) dial telephone, dial phone", "529) diaper, nappy, napkin", "530) digital clock", "531) digital watch", "532) dining table, board", "533) dishrag, dishcloth", "534) dishwasher, dish washer, dishwashing machine", "535) disk brake, disc brake", "536) dock, dockage, docking facility", "537) dogsled, dog sled, dog sleigh", "538) dome", "539) doormat, welcome mat", "540) drilling platform, offshore rig", "541) drum, membranophone, tympan", "542) drumstick", "543) dumbbell", "544) Dutch oven", "545) electric fan, blower", "546) electric guitar", "547) electric locomotive", "548) entertainment center", "549) envelope", "550) espresso maker", "551) face powder", "552) feather boa, boa", "553) file, file cabinet, filing cabinet", "554) fireboat", "555) fire engine, fire truck", "556) fire screen, fireguard", "557) flagpole, flagstaff", "558) flute, transverse flute", "559) folding chair", "560) football helmet", "561) forklift", "562) fountain", "563) fountain pen", "564) four-poster", "565) freight car", "566) French horn, horn", "567) frying pan, frypan, skillet", "568) fur coat", "569) garbage truck, dustcart", "570) gasmask, respirator, gas helmet", "571) gas pump, gasoline pump, petrol pump, island dispenser", "572) goblet", "573) go-kart", "574) golf ball", "575) golfcart, golf cart", "576) gondola", "577) gong, tam-tam", "578) gown", "579) grand piano, grand", "580) greenhouse, nursery, glasshouse", "581) grille, radiator grille", "582) grocery store, grocery, food market, market", "583) guillotine", "584) hair slide", "585) hair spray", "586) half track", "587) hammer", "588) hamper", "589) hand blower, blow dryer, blow drier, hair dryer, hair drier", "590) hand-held computer, hand-held microcomputer", "591) handkerchief, hankie, hanky, hankey", "592) hard disc, hard disk, fixed disk", "593) harmonica, mouth organ, harp, mouth harp", "594) harp", "595) harvester, reaper", "596) hatchet", "597) holster", "598) home theater, home theatre", "599) honeycomb", "600) hook, claw", "601) hoopskirt, crinoline", "602) horizontal bar, high bar", "603) horse cart, horse-cart", "604) hourglass", "605) iPod", "606) iron, smoothing iron", "607) jack-o'-lantern", "608) jean, blue jean, denim", "609) jeep, landrover", "610) jersey, T-shirt, tee shirt", "611) jigsaw puzzle", "612) jinrikisha, ricksha, rickshaw", "613) joystick", "614) kimono", "615) knee pad", "616) knot", "617) lab coat, laboratory coat", "618) ladle", "619) lampshade, lamp shade", "620) laptop, laptop computer", "621) lawn mower, mower", "622) lens cap, lens cover", "623) letter opener, paper knife, paperknife", "624) library", "625) lifeboat", "626) lighter, light, igniter, ignitor", "627) limousine, limo", "628) liner, ocean liner", "629) lipstick, lip rouge", "630) Loafer", "631) lotion", "632) loudspeaker, speaker, speaker unit, loudspeaker system, speaker system", "633) loupe, jeweler's loupe", "634) lumbermill, sawmill", "635) magnetic compass", "636) mailbag, postbag", "637) mailbox, letter box", "638) maillot", "639) maillot, tank suit", "640) manhole cover", "641) maraca", "642) marimba, xylophone", "643) mask", "644) matchstick", "645) maypole", "646) maze, labyrinth", "647) measuring cup", "648) medicine chest, medicine cabinet", "649) megalith, megalithic structure", "650) microphone, mike", "651) microwave, microwave oven", "652) military uniform", "653) milk can", "654) minibus", "655) miniskirt, mini", "656) minivan", "657) missile", "658) mitten", "659) mixing bowl", "660) mobile home, manufactured home", "661) Model T", "662) modem", "663) monastery", "664) monitor", "665) moped", "666) mortar", "667) mortarboard", "668) mosque", "669) mosquito net", "670) motor scooter, scooter", "671) mountain bike, all-terrain bike, off-roader", "672) mountain tent", "673) mouse, computer mouse", "674) mousetrap", "675) moving van", "676) muzzle", "677) nail", "678) neck brace", "679) necklace", "680) nipple", "681) notebook, notebook computer", "682) obelisk", "683) oboe, hautboy, hautbois", "684) ocarina, sweet potato", "685) odometer, hodometer, mileometer, milometer", "686) oil filter", "687) organ, pipe organ", "688) oscilloscope, scope, cathode-ray oscilloscope, CRO", "689) overskirt", "690) oxcart", "691) oxygen mask", "692) packet", "693) paddle, boat paddle", "694) paddlewheel, paddle wheel", "695) padlock", "696) paintbrush", "697) pajama, pyjama, pj's, jammies", "698) palace", "699) panpipe, pandean pipe, syrinx", "700) paper towel", "701) parachute, chute", "702) parallel bars, bars", "703) park bench", "704) parking meter", "705) passenger car, coach, carriage", "706) patio, terrace", "707) pay-phone, pay-station", "708) pedestal, plinth, footstall", "709) pencil box, pencil case", "710) pencil sharpener", "711) perfume, essence", "712) Petri dish", "713) photocopier", "714) pick, plectrum, plectron", "715) pickelhaube", "716) picket fence, paling", "717) pickup, pickup truck", "718) pier", "719) piggy bank, penny bank", "720) pill bottle", "721) pillow", "722) ping-pong ball", "723) pinwheel", "724) pirate, pirate ship", "725) pitcher, ewer", "726) plane, carpenter's plane, woodworking plane", "727) planetarium", "728) plastic bag", "729) plate rack", "730) plow, plough", "731) plunger, plumber's helper", "732) Polaroid camera, Polaroid Land camera", "733) pole", "734) police van, police wagon, paddy wagon, patrol wagon, wagon, black Maria", "735) poncho", "736) pool table, billiard table, snooker table", "737) pop bottle, soda bottle", "738) pot, flowerpot", "739) potter's wheel", "740) power drill", "741) prayer rug, prayer mat", "742) printer", "743) prison, prison house", "744) projectile, missile", "745) projector", "746) puck, hockey puck", "747) punching bag, punch bag, punching ball, punchball", "748) purse", "749) quill, quill pen", "750) quilt, comforter, comfort, puff", "751) racer, race car, racing car", "752) racket, racquet", "753) radiator", "754) radio, wireless", "755) radio telescope, radio reflector", "756) rain barrel", "757) recreational vehicle, RV, R.V.", "758) reel", "759) reflex camera", "760) refrigerator, icebox", "761) remote control, remote", "762) restaurant, eating house, eating place, eatery", "763) revolver, six-gun, six-shooter", "764) rifle", "765) rocking chair, rocker", "766) rotisserie", "767) rubber eraser, rubber, pencil eraser", "768) rugby ball", "769) rule, ruler", "770) running shoe", "771) safe", "772) safety pin", "773) saltshaker, salt shaker", "774) sandal", "775) sarong", "776) sax, saxophone", "777) scabbard", "778) scale, weighing machine", "779) school bus", "780) schooner", "781) scoreboard", "782) screen, CRT screen", "783) screw", "784) screwdriver", "785) seat belt, seatbelt", "786) sewing machine", "787) shield, buckler", "788) shoe shop, shoe-shop, shoe store", "789) shoji", "790) shopping basket", "791) shopping cart", "792) shovel", "793) shower cap", "794) shower curtain", "795) ski", "796) ski mask", "797) sleeping bag", "798) slide rule, slipstick", "799) sliding door", "800) slot, one-armed bandit", "801) snorkel", "802) snowmobile", "803) snowplow, snowplough", "804) soap dispenser", "805) soccer ball", "806) sock", "807) solar dish, solar collector, solar furnace", "808) sombrero", "809) soup bowl", "810) space bar", "811) space heater", "812) space shuttle", "813) spatula", "814) speedboat", "815) spider web, spider's web", "816) spindle", "817) sports car, sport car", "818) spotlight, spot", "819) stage", "820) steam locomotive", "821) steel arch bridge", "822) steel drum", "823) stethoscope", "824) stole", "825) stone wall", "826) stopwatch, stop watch", "827) stove", "828) strainer", "829) streetcar, tram, tramcar, trolley, trolley car", "830) stretcher", "831) studio couch, day bed", "832) stupa, tope", "833) submarine, pigboat, sub, U-boat", "834) suit, suit of clothes", "835) sundial", "836) sunglass", "837) sunglasses, dark glasses, shades", "838) sunscreen, sunblock, sun blocker", "839) suspension bridge", "840) swab, swob, mop", "841) sweatshirt", "842) swimming trunks, bathing trunks", "843) swing", "844) switch, electric switch, electrical switch", "845) syringe", "846) table lamp", "847) tank, army tank, armored combat vehicle, armoured combat vehicle", "848) tape player", "849) teapot", "850) teddy, teddy bear", "851) television, television system", "852) tennis ball", "853) thatch, thatched roof", "854) theater curtain, theatre curtain", "855) thimble", "856) thresher, thrasher, threshing machine", "857) throne", "858) tile roof", "859) toaster", "860) tobacco shop, tobacconist shop, tobacconist", "861) toilet seat", "862) torch", "863) totem pole", "864) tow truck, tow car, wrecker", "865) toyshop", "866) tractor", "867) trailer truck, tractor trailer, trucking rig, rig, articulated lorry, semi", "868) tray", "869) trench coat", "870) tricycle, trike, velocipede", "871) trimaran", "872) tripod", "873) triumphal arch", "874) trolleybus, trolley coach, trackless trolley", "875) trombone", "876) tub, vat", "877) turnstile", "878) typewriter keyboard", "879) umbrella", "880) unicycle, monocycle", "881) upright, upright piano", "882) vacuum, vacuum cleaner", "883) vase", "884) vault", "885) velvet", "886) vending machine", "887) vestment", "888) viaduct", "889) violin, fiddle", "890) volleyball", "891) waffle iron", "892) wall clock", "893) wallet, billfold, notecase, pocketbook", "894) wardrobe, closet, press", "895) warplane, military plane", "896) washbasin, handbasin, washbowl, lavabo, wash-hand basin", "897) washer, automatic washer, washing machine", "898) water bottle", "899) water jug", "900) water tower", "901) whiskey jug", "902) whistle", "903) wig", "904) window screen", "905) window shade", "906) Windsor tie", "907) wine bottle", "908) wing", "909) wok", "910) wooden spoon", "911) wool, woolen, woollen", "912) worm fence, snake fence, snake-rail fence, Virginia fence", "913) wreck", "914) yawl", "915) yurt", "916) web site, website, internet site, site", "917) comic book", "918) crossword puzzle, crossword", "919) street sign", "920) traffic light, traffic signal, stoplight", "921) book jacket, dust cover, dust jacket, dust wrapper", "922) menu", "923) plate", "924) guacamole", "925) consomme", "926) hot pot, hotpot", "927) trifle", "928) ice cream, icecream", "929) ice lolly, lolly, lollipop, popsicle", "930) French loaf", "931) bagel, beigel", "932) pretzel", "933) cheeseburger", "934) hotdog, hot dog, red hot", "935) mashed potato", "936) head cabbage", "937) broccoli", "938) cauliflower", "939) zucchini, courgette", "940) spaghetti squash", "941) acorn squash", "942) butternut squash", "943) cucumber, cuke", "944) artichoke, globe artichoke", "945) bell pepper", "946) cardoon", "947) mushroom", "948) Granny Smith", "949) strawberry", "950) orange", "951) lemon", "952) fig", "953) pineapple, ananas", "954) banana", "955) jackfruit, jak, jack", "956) custard apple", "957) pomegranate", "958) hay", "959) carbonara", "960) chocolate sauce, chocolate syrup", "961) dough", "962) meat loaf, meatloaf", "963) pizza, pizza pie", "964) potpie", "965) burrito", "966) red wine", "967) espresso", "968) cup", "969) eggnog", "970) alp", "971) bubble", "972) cliff, drop, drop-off", "973) coral reef", "974) geyser", "975) lakeside, lakeshore", "976) promontory, headland, head, foreland", "977) sandbar, sand bar", "978) seashore, coast, seacoast, sea-coast", "979) valley, vale", "980) volcano", "981) ballplayer, baseball player", "982) groom, bridegroom", "983) scuba diver", "984) rapeseed", "985) daisy", "986) yellow lady's slipper, yellow lady-slipper, Cypripedium calceolus, Cypripedium parviflorum", "987) corn", "988) acorn", "989) hip, rose hip, rosehip", "990) buckeye, horse chestnut, conker", "991) coral fungus", "992) agaric", "993) gyromitra", "994) stinkhorn, carrion fungus", "995) earthstar", "996) hen-of-the-woods, hen of the woods, Polyporus frondosus, Grifola frondosa", "997) bolete", "998) ear, spike, capitulum", "999) toilet tissue, toilet paper, bathroom tissue"]
z = truncated_z_sample(num_samples, truncation, noise_seed)
y = int(category.split(')')[0])
ims = sample(sess, z, y, truncation=truncation)
imshow(imgrid(ims, cols=min(num_samples, 5)))
```
# Interpolate between BigGAN samples
Try setting different **`category`**s with the same **`noise_seed`**s, or the same **`category`**s with different **`noise_seed`**s. Or go wild and set both any way you like!
(Double-click on the cell to view code.)
```
#@title Interpolation { display-mode: "form", run: "auto" }
num_samples = 2 #@param {type:"slider", min:1, max:5, step:1}
num_interps = 5 #@param {type:"slider", min:2, max:10, step:1}
truncation = 0.2 #@param {type:"slider", min:0.02, max:1, step:0.02}
noise_seed_A = 0 #@param {type:"slider", min:0, max:100, step:1}
category_A = "207) golden retriever" #@param ["0) tench, Tinca tinca", "1) goldfish, Carassius auratus", "2) great white shark, white shark, man-eater, man-eating shark, Carcharodon carcharias", "3) tiger shark, Galeocerdo cuvieri", "4) hammerhead, hammerhead shark", "5) electric ray, crampfish, numbfish, torpedo", "6) stingray", "7) cock", "8) hen", "9) ostrich, Struthio camelus", "10) brambling, Fringilla montifringilla", "11) goldfinch, Carduelis carduelis", "12) house finch, linnet, Carpodacus mexicanus", "13) junco, snowbird", "14) indigo bunting, indigo finch, indigo bird, Passerina cyanea", "15) robin, American robin, Turdus migratorius", "16) bulbul", "17) jay", "18) magpie", "19) chickadee", "20) water ouzel, dipper", "21) kite", "22) bald eagle, American eagle, Haliaeetus leucocephalus", "23) vulture", "24) great grey owl, great gray owl, Strix nebulosa", "25) European fire salamander, Salamandra salamandra", "26) common newt, Triturus vulgaris", "27) eft", "28) spotted salamander, Ambystoma maculatum", "29) axolotl, mud puppy, Ambystoma mexicanum", "30) bullfrog, Rana catesbeiana", "31) tree frog, tree-frog", "32) tailed frog, bell toad, ribbed toad, tailed toad, Ascaphus trui", "33) loggerhead, loggerhead turtle, Caretta caretta", "34) leatherback turtle, leatherback, leathery turtle, Dermochelys coriacea", "35) mud turtle", "36) terrapin", "37) box turtle, box tortoise", "38) banded gecko", "39) common iguana, iguana, Iguana iguana", "40) American chameleon, anole, Anolis carolinensis", "41) whiptail, whiptail lizard", "42) agama", "43) frilled lizard, Chlamydosaurus kingi", "44) alligator lizard", "45) Gila monster, Heloderma suspectum", "46) green lizard, Lacerta viridis", "47) African chameleon, Chamaeleo chamaeleon", "48) Komodo dragon, Komodo lizard, dragon lizard, giant lizard, Varanus komodoensis", "49) African crocodile, Nile crocodile, Crocodylus niloticus", "50) American alligator, Alligator mississipiensis", "51) triceratops", "52) thunder snake, worm snake, Carphophis amoenus", "53) ringneck snake, ring-necked snake, ring snake", "54) hognose snake, puff adder, sand viper", "55) green snake, grass snake", "56) king snake, kingsnake", "57) garter snake, grass snake", "58) water snake", "59) vine snake", "60) night snake, Hypsiglena torquata", "61) boa constrictor, Constrictor constrictor", "62) rock python, rock snake, Python sebae", "63) Indian cobra, Naja naja", "64) green mamba", "65) sea snake", "66) horned viper, cerastes, sand viper, horned asp, Cerastes cornutus", "67) diamondback, diamondback rattlesnake, Crotalus adamanteus", "68) sidewinder, horned rattlesnake, Crotalus cerastes", "69) trilobite", "70) harvestman, daddy longlegs, Phalangium opilio", "71) scorpion", "72) black and gold garden spider, Argiope aurantia", "73) barn spider, Araneus cavaticus", "74) garden spider, Aranea diademata", "75) black widow, Latrodectus mactans", "76) tarantula", "77) wolf spider, hunting spider", "78) tick", "79) centipede", "80) black grouse", "81) ptarmigan", "82) ruffed grouse, partridge, Bonasa umbellus", "83) prairie chicken, prairie grouse, prairie fowl", "84) peacock", "85) quail", "86) partridge", "87) African grey, African gray, Psittacus erithacus", "88) macaw", "89) sulphur-crested cockatoo, Kakatoe galerita, Cacatua galerita", "90) lorikeet", "91) coucal", "92) bee eater", "93) hornbill", "94) hummingbird", "95) jacamar", "96) toucan", "97) drake", "98) red-breasted merganser, Mergus serrator", "99) goose", "100) black swan, Cygnus atratus", "101) tusker", "102) echidna, spiny anteater, anteater", "103) platypus, duckbill, duckbilled platypus, duck-billed platypus, Ornithorhynchus anatinus", "104) wallaby, brush kangaroo", "105) koala, koala bear, kangaroo bear, native bear, Phascolarctos cinereus", "106) wombat", "107) jellyfish", "108) sea anemone, anemone", "109) brain coral", "110) flatworm, platyhelminth", "111) nematode, nematode worm, roundworm", "112) conch", "113) snail", "114) slug", "115) sea slug, nudibranch", "116) chiton, coat-of-mail shell, sea cradle, polyplacophore", "117) chambered nautilus, pearly nautilus, nautilus", "118) Dungeness crab, Cancer magister", "119) rock crab, Cancer irroratus", "120) fiddler crab", "121) king crab, Alaska crab, Alaskan king crab, Alaska king crab, Paralithodes camtschatica", "122) American lobster, Northern lobster, Maine lobster, Homarus americanus", "123) spiny lobster, langouste, rock lobster, crawfish, crayfish, sea crawfish", "124) crayfish, crawfish, crawdad, crawdaddy", "125) hermit crab", "126) isopod", "127) white stork, Ciconia ciconia", "128) black stork, Ciconia nigra", "129) spoonbill", "130) flamingo", "131) little blue heron, Egretta caerulea", "132) American egret, great white heron, Egretta albus", "133) bittern", "134) crane", "135) limpkin, Aramus pictus", "136) European gallinule, Porphyrio porphyrio", "137) American coot, marsh hen, mud hen, water hen, Fulica americana", "138) bustard", "139) ruddy turnstone, Arenaria interpres", "140) red-backed sandpiper, dunlin, Erolia alpina", "141) redshank, Tringa totanus", "142) dowitcher", "143) oystercatcher, oyster catcher", "144) pelican", "145) king penguin, Aptenodytes patagonica", "146) albatross, mollymawk", "147) grey whale, gray whale, devilfish, Eschrichtius gibbosus, Eschrichtius robustus", "148) killer whale, killer, orca, grampus, sea wolf, Orcinus orca", "149) dugong, Dugong dugon", "150) sea lion", "151) Chihuahua", "152) Japanese spaniel", "153) Maltese dog, Maltese terrier, Maltese", "154) Pekinese, Pekingese, Peke", "155) Shih-Tzu", "156) Blenheim spaniel", "157) papillon", "158) toy terrier", "159) Rhodesian ridgeback", "160) Afghan hound, Afghan", "161) basset, basset hound", "162) beagle", "163) bloodhound, sleuthhound", "164) bluetick", "165) black-and-tan coonhound", "166) Walker hound, Walker foxhound", "167) English foxhound", "168) redbone", "169) borzoi, Russian wolfhound", "170) Irish wolfhound", "171) Italian greyhound", "172) whippet", "173) Ibizan hound, Ibizan Podenco", "174) Norwegian elkhound, elkhound", "175) otterhound, otter hound", "176) Saluki, gazelle hound", "177) Scottish deerhound, deerhound", "178) Weimaraner", "179) Staffordshire bullterrier, Staffordshire bull terrier", "180) American Staffordshire terrier, Staffordshire terrier, American pit bull terrier, pit bull terrier", "181) Bedlington terrier", "182) Border terrier", "183) Kerry blue terrier", "184) Irish terrier", "185) Norfolk terrier", "186) Norwich terrier", "187) Yorkshire terrier", "188) wire-haired fox terrier", "189) Lakeland terrier", "190) Sealyham terrier, Sealyham", "191) Airedale, Airedale terrier", "192) cairn, cairn terrier", "193) Australian terrier", "194) Dandie Dinmont, Dandie Dinmont terrier", "195) Boston bull, Boston terrier", "196) miniature schnauzer", "197) giant schnauzer", "198) standard schnauzer", "199) Scotch terrier, Scottish terrier, Scottie", "200) Tibetan terrier, chrysanthemum dog", "201) silky terrier, Sydney silky", "202) soft-coated wheaten terrier", "203) West Highland white terrier", "204) Lhasa, Lhasa apso", "205) flat-coated retriever", "206) curly-coated retriever", "207) golden retriever", "208) Labrador retriever", "209) Chesapeake Bay retriever", "210) German short-haired pointer", "211) vizsla, Hungarian pointer", "212) English setter", "213) Irish setter, red setter", "214) Gordon setter", "215) Brittany spaniel", "216) clumber, clumber spaniel", "217) English springer, English springer spaniel", "218) Welsh springer spaniel", "219) cocker spaniel, English cocker spaniel, cocker", "220) Sussex spaniel", "221) Irish water spaniel", "222) kuvasz", "223) schipperke", "224) groenendael", "225) malinois", "226) briard", "227) kelpie", "228) komondor", "229) Old English sheepdog, bobtail", "230) Shetland sheepdog, Shetland sheep dog, Shetland", "231) collie", "232) Border collie", "233) Bouvier des Flandres, Bouviers des Flandres", "234) Rottweiler", "235) German shepherd, German shepherd dog, German police dog, alsatian", "236) Doberman, Doberman pinscher", "237) miniature pinscher", "238) Greater Swiss Mountain dog", "239) Bernese mountain dog", "240) Appenzeller", "241) EntleBucher", "242) boxer", "243) bull mastiff", "244) Tibetan mastiff", "245) French bulldog", "246) Great Dane", "247) Saint Bernard, St Bernard", "248) Eskimo dog, husky", "249) malamute, malemute, Alaskan malamute", "250) Siberian husky", "251) dalmatian, coach dog, carriage dog", "252) affenpinscher, monkey pinscher, monkey dog", "253) basenji", "254) pug, pug-dog", "255) Leonberg", "256) Newfoundland, Newfoundland dog", "257) Great Pyrenees", "258) Samoyed, Samoyede", "259) Pomeranian", "260) chow, chow chow", "261) keeshond", "262) Brabancon griffon", "263) Pembroke, Pembroke Welsh corgi", "264) Cardigan, Cardigan Welsh corgi", "265) toy poodle", "266) miniature poodle", "267) standard poodle", "268) Mexican hairless", "269) timber wolf, grey wolf, gray wolf, Canis lupus", "270) white wolf, Arctic wolf, Canis lupus tundrarum", "271) red wolf, maned wolf, Canis rufus, Canis niger", "272) coyote, prairie wolf, brush wolf, Canis latrans", "273) dingo, warrigal, warragal, Canis dingo", "274) dhole, Cuon alpinus", "275) African hunting dog, hyena dog, Cape hunting dog, Lycaon pictus", "276) hyena, hyaena", "277) red fox, Vulpes vulpes", "278) kit fox, Vulpes macrotis", "279) Arctic fox, white fox, Alopex lagopus", "280) grey fox, gray fox, Urocyon cinereoargenteus", "281) tabby, tabby cat", "282) tiger cat", "283) Persian cat", "284) Siamese cat, Siamese", "285) Egyptian cat", "286) cougar, puma, catamount, mountain lion, painter, panther, Felis concolor", "287) lynx, catamount", "288) leopard, Panthera pardus", "289) snow leopard, ounce, Panthera uncia", "290) jaguar, panther, Panthera onca, Felis onca", "291) lion, king of beasts, Panthera leo", "292) tiger, Panthera tigris", "293) cheetah, chetah, Acinonyx jubatus", "294) brown bear, bruin, Ursus arctos", "295) American black bear, black bear, Ursus americanus, Euarctos americanus", "296) ice bear, polar bear, Ursus Maritimus, Thalarctos maritimus", "297) sloth bear, Melursus ursinus, Ursus ursinus", "298) mongoose", "299) meerkat, mierkat", "300) tiger beetle", "301) ladybug, ladybeetle, lady beetle, ladybird, ladybird beetle", "302) ground beetle, carabid beetle", "303) long-horned beetle, longicorn, longicorn beetle", "304) leaf beetle, chrysomelid", "305) dung beetle", "306) rhinoceros beetle", "307) weevil", "308) fly", "309) bee", "310) ant, emmet, pismire", "311) grasshopper, hopper", "312) cricket", "313) walking stick, walkingstick, stick insect", "314) cockroach, roach", "315) mantis, mantid", "316) cicada, cicala", "317) leafhopper", "318) lacewing, lacewing fly", "319) dragonfly, darning needle, devil's darning needle, sewing needle, snake feeder, snake doctor, mosquito hawk, skeeter hawk", "320) damselfly", "321) admiral", "322) ringlet, ringlet butterfly", "323) monarch, monarch butterfly, milkweed butterfly, Danaus plexippus", "324) cabbage butterfly", "325) sulphur butterfly, sulfur butterfly", "326) lycaenid, lycaenid butterfly", "327) starfish, sea star", "328) sea urchin", "329) sea cucumber, holothurian", "330) wood rabbit, cottontail, cottontail rabbit", "331) hare", "332) Angora, Angora rabbit", "333) hamster", "334) porcupine, hedgehog", "335) fox squirrel, eastern fox squirrel, Sciurus niger", "336) marmot", "337) beaver", "338) guinea pig, Cavia cobaya", "339) sorrel", "340) zebra", "341) hog, pig, grunter, squealer, Sus scrofa", "342) wild boar, boar, Sus scrofa", "343) warthog", "344) hippopotamus, hippo, river horse, Hippopotamus amphibius", "345) ox", "346) water buffalo, water ox, Asiatic buffalo, Bubalus bubalis", "347) bison", "348) ram, tup", "349) bighorn, bighorn sheep, cimarron, Rocky Mountain bighorn, Rocky Mountain sheep, Ovis canadensis", "350) ibex, Capra ibex", "351) hartebeest", "352) impala, Aepyceros melampus", "353) gazelle", "354) Arabian camel, dromedary, Camelus dromedarius", "355) llama", "356) weasel", "357) mink", "358) polecat, fitch, foulmart, foumart, Mustela putorius", "359) black-footed ferret, ferret, Mustela nigripes", "360) otter", "361) skunk, polecat, wood pussy", "362) badger", "363) armadillo", "364) three-toed sloth, ai, Bradypus tridactylus", "365) orangutan, orang, orangutang, Pongo pygmaeus", "366) gorilla, Gorilla gorilla", "367) chimpanzee, chimp, Pan troglodytes", "368) gibbon, Hylobates lar", "369) siamang, Hylobates syndactylus, Symphalangus syndactylus", "370) guenon, guenon monkey", "371) patas, hussar monkey, Erythrocebus patas", "372) baboon", "373) macaque", "374) langur", "375) colobus, colobus monkey", "376) proboscis monkey, Nasalis larvatus", "377) marmoset", "378) capuchin, ringtail, Cebus capucinus", "379) howler monkey, howler", "380) titi, titi monkey", "381) spider monkey, Ateles geoffroyi", "382) squirrel monkey, Saimiri sciureus", "383) Madagascar cat, ring-tailed lemur, Lemur catta", "384) indri, indris, Indri indri, Indri brevicaudatus", "385) Indian elephant, Elephas maximus", "386) African elephant, Loxodonta africana", "387) lesser panda, red panda, panda, bear cat, cat bear, Ailurus fulgens", "388) giant panda, panda, panda bear, coon bear, Ailuropoda melanoleuca", "389) barracouta, snoek", "390) eel", "391) coho, cohoe, coho salmon, blue jack, silver salmon, Oncorhynchus kisutch", "392) rock beauty, Holocanthus tricolor", "393) anemone fish", "394) sturgeon", "395) gar, garfish, garpike, billfish, Lepisosteus osseus", "396) lionfish", "397) puffer, pufferfish, blowfish, globefish", "398) abacus", "399) abaya", "400) academic gown, academic robe, judge's robe", "401) accordion, piano accordion, squeeze box", "402) acoustic guitar", "403) aircraft carrier, carrier, flattop, attack aircraft carrier", "404) airliner", "405) airship, dirigible", "406) altar", "407) ambulance", "408) amphibian, amphibious vehicle", "409) analog clock", "410) apiary, bee house", "411) apron", "412) ashcan, trash can, garbage can, wastebin, ash bin, ash-bin, ashbin, dustbin, trash barrel, trash bin", "413) assault rifle, assault gun", "414) backpack, back pack, knapsack, packsack, rucksack, haversack", "415) bakery, bakeshop, bakehouse", "416) balance beam, beam", "417) balloon", "418) ballpoint, ballpoint pen, ballpen, Biro", "419) Band Aid", "420) banjo", "421) bannister, banister, balustrade, balusters, handrail", "422) barbell", "423) barber chair", "424) barbershop", "425) barn", "426) barometer", "427) barrel, cask", "428) barrow, garden cart, lawn cart, wheelbarrow", "429) baseball", "430) basketball", "431) bassinet", "432) bassoon", "433) bathing cap, swimming cap", "434) bath towel", "435) bathtub, bathing tub, bath, tub", "436) beach wagon, station wagon, wagon, estate car, beach waggon, station waggon, waggon", "437) beacon, lighthouse, beacon light, pharos", "438) beaker", "439) bearskin, busby, shako", "440) beer bottle", "441) beer glass", "442) bell cote, bell cot", "443) bib", "444) bicycle-built-for-two, tandem bicycle, tandem", "445) bikini, two-piece", "446) binder, ring-binder", "447) binoculars, field glasses, opera glasses", "448) birdhouse", "449) boathouse", "450) bobsled, bobsleigh, bob", "451) bolo tie, bolo, bola tie, bola", "452) bonnet, poke bonnet", "453) bookcase", "454) bookshop, bookstore, bookstall", "455) bottlecap", "456) bow", "457) bow tie, bow-tie, bowtie", "458) brass, memorial tablet, plaque", "459) brassiere, bra, bandeau", "460) breakwater, groin, groyne, mole, bulwark, seawall, jetty", "461) breastplate, aegis, egis", "462) broom", "463) bucket, pail", "464) buckle", "465) bulletproof vest", "466) bullet train, bullet", "467) butcher shop, meat market", "468) cab, hack, taxi, taxicab", "469) caldron, cauldron", "470) candle, taper, wax light", "471) cannon", "472) canoe", "473) can opener, tin opener", "474) cardigan", "475) car mirror", "476) carousel, carrousel, merry-go-round, roundabout, whirligig", "477) carpenter's kit, tool kit", "478) carton", "479) car wheel", "480) cash machine, cash dispenser, automated teller machine, automatic teller machine, automated teller, automatic teller, ATM", "481) cassette", "482) cassette player", "483) castle", "484) catamaran", "485) CD player", "486) cello, violoncello", "487) cellular telephone, cellular phone, cellphone, cell, mobile phone", "488) chain", "489) chainlink fence", "490) chain mail, ring mail, mail, chain armor, chain armour, ring armor, ring armour", "491) chain saw, chainsaw", "492) chest", "493) chiffonier, commode", "494) chime, bell, gong", "495) china cabinet, china closet", "496) Christmas stocking", "497) church, church building", "498) cinema, movie theater, movie theatre, movie house, picture palace", "499) cleaver, meat cleaver, chopper", "500) cliff dwelling", "501) cloak", "502) clog, geta, patten, sabot", "503) cocktail shaker", "504) coffee mug", "505) coffeepot", "506) coil, spiral, volute, whorl, helix", "507) combination lock", "508) computer keyboard, keypad", "509) confectionery, confectionary, candy store", "510) container ship, containership, container vessel", "511) convertible", "512) corkscrew, bottle screw", "513) cornet, horn, trumpet, trump", "514) cowboy boot", "515) cowboy hat, ten-gallon hat", "516) cradle", "517) crane", "518) crash helmet", "519) crate", "520) crib, cot", "521) Crock Pot", "522) croquet ball", "523) crutch", "524) cuirass", "525) dam, dike, dyke", "526) desk", "527) desktop computer", "528) dial telephone, dial phone", "529) diaper, nappy, napkin", "530) digital clock", "531) digital watch", "532) dining table, board", "533) dishrag, dishcloth", "534) dishwasher, dish washer, dishwashing machine", "535) disk brake, disc brake", "536) dock, dockage, docking facility", "537) dogsled, dog sled, dog sleigh", "538) dome", "539) doormat, welcome mat", "540) drilling platform, offshore rig", "541) drum, membranophone, tympan", "542) drumstick", "543) dumbbell", "544) Dutch oven", "545) electric fan, blower", "546) electric guitar", "547) electric locomotive", "548) entertainment center", "549) envelope", "550) espresso maker", "551) face powder", "552) feather boa, boa", "553) file, file cabinet, filing cabinet", "554) fireboat", "555) fire engine, fire truck", "556) fire screen, fireguard", "557) flagpole, flagstaff", "558) flute, transverse flute", "559) folding chair", "560) football helmet", "561) forklift", "562) fountain", "563) fountain pen", "564) four-poster", "565) freight car", "566) French horn, horn", "567) frying pan, frypan, skillet", "568) fur coat", "569) garbage truck, dustcart", "570) gasmask, respirator, gas helmet", "571) gas pump, gasoline pump, petrol pump, island dispenser", "572) goblet", "573) go-kart", "574) golf ball", "575) golfcart, golf cart", "576) gondola", "577) gong, tam-tam", "578) gown", "579) grand piano, grand", "580) greenhouse, nursery, glasshouse", "581) grille, radiator grille", "582) grocery store, grocery, food market, market", "583) guillotine", "584) hair slide", "585) hair spray", "586) half track", "587) hammer", "588) hamper", "589) hand blower, blow dryer, blow drier, hair dryer, hair drier", "590) hand-held computer, hand-held microcomputer", "591) handkerchief, hankie, hanky, hankey", "592) hard disc, hard disk, fixed disk", "593) harmonica, mouth organ, harp, mouth harp", "594) harp", "595) harvester, reaper", "596) hatchet", "597) holster", "598) home theater, home theatre", "599) honeycomb", "600) hook, claw", "601) hoopskirt, crinoline", "602) horizontal bar, high bar", "603) horse cart, horse-cart", "604) hourglass", "605) iPod", "606) iron, smoothing iron", "607) jack-o'-lantern", "608) jean, blue jean, denim", "609) jeep, landrover", "610) jersey, T-shirt, tee shirt", "611) jigsaw puzzle", "612) jinrikisha, ricksha, rickshaw", "613) joystick", "614) kimono", "615) knee pad", "616) knot", "617) lab coat, laboratory coat", "618) ladle", "619) lampshade, lamp shade", "620) laptop, laptop computer", "621) lawn mower, mower", "622) lens cap, lens cover", "623) letter opener, paper knife, paperknife", "624) library", "625) lifeboat", "626) lighter, light, igniter, ignitor", "627) limousine, limo", "628) liner, ocean liner", "629) lipstick, lip rouge", "630) Loafer", "631) lotion", "632) loudspeaker, speaker, speaker unit, loudspeaker system, speaker system", "633) loupe, jeweler's loupe", "634) lumbermill, sawmill", "635) magnetic compass", "636) mailbag, postbag", "637) mailbox, letter box", "638) maillot", "639) maillot, tank suit", "640) manhole cover", "641) maraca", "642) marimba, xylophone", "643) mask", "644) matchstick", "645) maypole", "646) maze, labyrinth", "647) measuring cup", "648) medicine chest, medicine cabinet", "649) megalith, megalithic structure", "650) microphone, mike", "651) microwave, microwave oven", "652) military uniform", "653) milk can", "654) minibus", "655) miniskirt, mini", "656) minivan", "657) missile", "658) mitten", "659) mixing bowl", "660) mobile home, manufactured home", "661) Model T", "662) modem", "663) monastery", "664) monitor", "665) moped", "666) mortar", "667) mortarboard", "668) mosque", "669) mosquito net", "670) motor scooter, scooter", "671) mountain bike, all-terrain bike, off-roader", "672) mountain tent", "673) mouse, computer mouse", "674) mousetrap", "675) moving van", "676) muzzle", "677) nail", "678) neck brace", "679) necklace", "680) nipple", "681) notebook, notebook computer", "682) obelisk", "683) oboe, hautboy, hautbois", "684) ocarina, sweet potato", "685) odometer, hodometer, mileometer, milometer", "686) oil filter", "687) organ, pipe organ", "688) oscilloscope, scope, cathode-ray oscilloscope, CRO", "689) overskirt", "690) oxcart", "691) oxygen mask", "692) packet", "693) paddle, boat paddle", "694) paddlewheel, paddle wheel", "695) padlock", "696) paintbrush", "697) pajama, pyjama, pj's, jammies", "698) palace", "699) panpipe, pandean pipe, syrinx", "700) paper towel", "701) parachute, chute", "702) parallel bars, bars", "703) park bench", "704) parking meter", "705) passenger car, coach, carriage", "706) patio, terrace", "707) pay-phone, pay-station", "708) pedestal, plinth, footstall", "709) pencil box, pencil case", "710) pencil sharpener", "711) perfume, essence", "712) Petri dish", "713) photocopier", "714) pick, plectrum, plectron", "715) pickelhaube", "716) picket fence, paling", "717) pickup, pickup truck", "718) pier", "719) piggy bank, penny bank", "720) pill bottle", "721) pillow", "722) ping-pong ball", "723) pinwheel", "724) pirate, pirate ship", "725) pitcher, ewer", "726) plane, carpenter's plane, woodworking plane", "727) planetarium", "728) plastic bag", "729) plate rack", "730) plow, plough", "731) plunger, plumber's helper", "732) Polaroid camera, Polaroid Land camera", "733) pole", "734) police van, police wagon, paddy wagon, patrol wagon, wagon, black Maria", "735) poncho", "736) pool table, billiard table, snooker table", "737) pop bottle, soda bottle", "738) pot, flowerpot", "739) potter's wheel", "740) power drill", "741) prayer rug, prayer mat", "742) printer", "743) prison, prison house", "744) projectile, missile", "745) projector", "746) puck, hockey puck", "747) punching bag, punch bag, punching ball, punchball", "748) purse", "749) quill, quill pen", "750) quilt, comforter, comfort, puff", "751) racer, race car, racing car", "752) racket, racquet", "753) radiator", "754) radio, wireless", "755) radio telescope, radio reflector", "756) rain barrel", "757) recreational vehicle, RV, R.V.", "758) reel", "759) reflex camera", "760) refrigerator, icebox", "761) remote control, remote", "762) restaurant, eating house, eating place, eatery", "763) revolver, six-gun, six-shooter", "764) rifle", "765) rocking chair, rocker", "766) rotisserie", "767) rubber eraser, rubber, pencil eraser", "768) rugby ball", "769) rule, ruler", "770) running shoe", "771) safe", "772) safety pin", "773) saltshaker, salt shaker", "774) sandal", "775) sarong", "776) sax, saxophone", "777) scabbard", "778) scale, weighing machine", "779) school bus", "780) schooner", "781) scoreboard", "782) screen, CRT screen", "783) screw", "784) screwdriver", "785) seat belt, seatbelt", "786) sewing machine", "787) shield, buckler", "788) shoe shop, shoe-shop, shoe store", "789) shoji", "790) shopping basket", "791) shopping cart", "792) shovel", "793) shower cap", "794) shower curtain", "795) ski", "796) ski mask", "797) sleeping bag", "798) slide rule, slipstick", "799) sliding door", "800) slot, one-armed bandit", "801) snorkel", "802) snowmobile", "803) snowplow, snowplough", "804) soap dispenser", "805) soccer ball", "806) sock", "807) solar dish, solar collector, solar furnace", "808) sombrero", "809) soup bowl", "810) space bar", "811) space heater", "812) space shuttle", "813) spatula", "814) speedboat", "815) spider web, spider's web", "816) spindle", "817) sports car, sport car", "818) spotlight, spot", "819) stage", "820) steam locomotive", "821) steel arch bridge", "822) steel drum", "823) stethoscope", "824) stole", "825) stone wall", "826) stopwatch, stop watch", "827) stove", "828) strainer", "829) streetcar, tram, tramcar, trolley, trolley car", "830) stretcher", "831) studio couch, day bed", "832) stupa, tope", "833) submarine, pigboat, sub, U-boat", "834) suit, suit of clothes", "835) sundial", "836) sunglass", "837) sunglasses, dark glasses, shades", "838) sunscreen, sunblock, sun blocker", "839) suspension bridge", "840) swab, swob, mop", "841) sweatshirt", "842) swimming trunks, bathing trunks", "843) swing", "844) switch, electric switch, electrical switch", "845) syringe", "846) table lamp", "847) tank, army tank, armored combat vehicle, armoured combat vehicle", "848) tape player", "849) teapot", "850) teddy, teddy bear", "851) television, television system", "852) tennis ball", "853) thatch, thatched roof", "854) theater curtain, theatre curtain", "855) thimble", "856) thresher, thrasher, threshing machine", "857) throne", "858) tile roof", "859) toaster", "860) tobacco shop, tobacconist shop, tobacconist", "861) toilet seat", "862) torch", "863) totem pole", "864) tow truck, tow car, wrecker", "865) toyshop", "866) tractor", "867) trailer truck, tractor trailer, trucking rig, rig, articulated lorry, semi", "868) tray", "869) trench coat", "870) tricycle, trike, velocipede", "871) trimaran", "872) tripod", "873) triumphal arch", "874) trolleybus, trolley coach, trackless trolley", "875) trombone", "876) tub, vat", "877) turnstile", "878) typewriter keyboard", "879) umbrella", "880) unicycle, monocycle", "881) upright, upright piano", "882) vacuum, vacuum cleaner", "883) vase", "884) vault", "885) velvet", "886) vending machine", "887) vestment", "888) viaduct", "889) violin, fiddle", "890) volleyball", "891) waffle iron", "892) wall clock", "893) wallet, billfold, notecase, pocketbook", "894) wardrobe, closet, press", "895) warplane, military plane", "896) washbasin, handbasin, washbowl, lavabo, wash-hand basin", "897) washer, automatic washer, washing machine", "898) water bottle", "899) water jug", "900) water tower", "901) whiskey jug", "902) whistle", "903) wig", "904) window screen", "905) window shade", "906) Windsor tie", "907) wine bottle", "908) wing", "909) wok", "910) wooden spoon", "911) wool, woolen, woollen", "912) worm fence, snake fence, snake-rail fence, Virginia fence", "913) wreck", "914) yawl", "915) yurt", "916) web site, website, internet site, site", "917) comic book", "918) crossword puzzle, crossword", "919) street sign", "920) traffic light, traffic signal, stoplight", "921) book jacket, dust cover, dust jacket, dust wrapper", "922) menu", "923) plate", "924) guacamole", "925) consomme", "926) hot pot, hotpot", "927) trifle", "928) ice cream, icecream", "929) ice lolly, lolly, lollipop, popsicle", "930) French loaf", "931) bagel, beigel", "932) pretzel", "933) cheeseburger", "934) hotdog, hot dog, red hot", "935) mashed potato", "936) head cabbage", "937) broccoli", "938) cauliflower", "939) zucchini, courgette", "940) spaghetti squash", "941) acorn squash", "942) butternut squash", "943) cucumber, cuke", "944) artichoke, globe artichoke", "945) bell pepper", "946) cardoon", "947) mushroom", "948) Granny Smith", "949) strawberry", "950) orange", "951) lemon", "952) fig", "953) pineapple, ananas", "954) banana", "955) jackfruit, jak, jack", "956) custard apple", "957) pomegranate", "958) hay", "959) carbonara", "960) chocolate sauce, chocolate syrup", "961) dough", "962) meat loaf, meatloaf", "963) pizza, pizza pie", "964) potpie", "965) burrito", "966) red wine", "967) espresso", "968) cup", "969) eggnog", "970) alp", "971) bubble", "972) cliff, drop, drop-off", "973) coral reef", "974) geyser", "975) lakeside, lakeshore", "976) promontory, headland, head, foreland", "977) sandbar, sand bar", "978) seashore, coast, seacoast, sea-coast", "979) valley, vale", "980) volcano", "981) ballplayer, baseball player", "982) groom, bridegroom", "983) scuba diver", "984) rapeseed", "985) daisy", "986) yellow lady's slipper, yellow lady-slipper, Cypripedium calceolus, Cypripedium parviflorum", "987) corn", "988) acorn", "989) hip, rose hip, rosehip", "990) buckeye, horse chestnut, conker", "991) coral fungus", "992) agaric", "993) gyromitra", "994) stinkhorn, carrion fungus", "995) earthstar", "996) hen-of-the-woods, hen of the woods, Polyporus frondosus, Grifola frondosa", "997) bolete", "998) ear, spike, capitulum", "999) toilet tissue, toilet paper, bathroom tissue"]
noise_seed_B = 0 #@param {type:"slider", min:0, max:100, step:1}
category_B = "8) hen" #@param ["0) tench, Tinca tinca", "1) goldfish, Carassius auratus", "2) great white shark, white shark, man-eater, man-eating shark, Carcharodon carcharias", "3) tiger shark, Galeocerdo cuvieri", "4) hammerhead, hammerhead shark", "5) electric ray, crampfish, numbfish, torpedo", "6) stingray", "7) cock", "8) hen", "9) ostrich, Struthio camelus", "10) brambling, Fringilla montifringilla", "11) goldfinch, Carduelis carduelis", "12) house finch, linnet, Carpodacus mexicanus", "13) junco, snowbird", "14) indigo bunting, indigo finch, indigo bird, Passerina cyanea", "15) robin, American robin, Turdus migratorius", "16) bulbul", "17) jay", "18) magpie", "19) chickadee", "20) water ouzel, dipper", "21) kite", "22) bald eagle, American eagle, Haliaeetus leucocephalus", "23) vulture", "24) great grey owl, great gray owl, Strix nebulosa", "25) European fire salamander, Salamandra salamandra", "26) common newt, Triturus vulgaris", "27) eft", "28) spotted salamander, Ambystoma maculatum", "29) axolotl, mud puppy, Ambystoma mexicanum", "30) bullfrog, Rana catesbeiana", "31) tree frog, tree-frog", "32) tailed frog, bell toad, ribbed toad, tailed toad, Ascaphus trui", "33) loggerhead, loggerhead turtle, Caretta caretta", "34) leatherback turtle, leatherback, leathery turtle, Dermochelys coriacea", "35) mud turtle", "36) terrapin", "37) box turtle, box tortoise", "38) banded gecko", "39) common iguana, iguana, Iguana iguana", "40) American chameleon, anole, Anolis carolinensis", "41) whiptail, whiptail lizard", "42) agama", "43) frilled lizard, Chlamydosaurus kingi", "44) alligator lizard", "45) Gila monster, Heloderma suspectum", "46) green lizard, Lacerta viridis", "47) African chameleon, Chamaeleo chamaeleon", "48) Komodo dragon, Komodo lizard, dragon lizard, giant lizard, Varanus komodoensis", "49) African crocodile, Nile crocodile, Crocodylus niloticus", "50) American alligator, Alligator mississipiensis", "51) triceratops", "52) thunder snake, worm snake, Carphophis amoenus", "53) ringneck snake, ring-necked snake, ring snake", "54) hognose snake, puff adder, sand viper", "55) green snake, grass snake", "56) king snake, kingsnake", "57) garter snake, grass snake", "58) water snake", "59) vine snake", "60) night snake, Hypsiglena torquata", "61) boa constrictor, Constrictor constrictor", "62) rock python, rock snake, Python sebae", "63) Indian cobra, Naja naja", "64) green mamba", "65) sea snake", "66) horned viper, cerastes, sand viper, horned asp, Cerastes cornutus", "67) diamondback, diamondback rattlesnake, Crotalus adamanteus", "68) sidewinder, horned rattlesnake, Crotalus cerastes", "69) trilobite", "70) harvestman, daddy longlegs, Phalangium opilio", "71) scorpion", "72) black and gold garden spider, Argiope aurantia", "73) barn spider, Araneus cavaticus", "74) garden spider, Aranea diademata", "75) black widow, Latrodectus mactans", "76) tarantula", "77) wolf spider, hunting spider", "78) tick", "79) centipede", "80) black grouse", "81) ptarmigan", "82) ruffed grouse, partridge, Bonasa umbellus", "83) prairie chicken, prairie grouse, prairie fowl", "84) peacock", "85) quail", "86) partridge", "87) African grey, African gray, Psittacus erithacus", "88) macaw", "89) sulphur-crested cockatoo, Kakatoe galerita, Cacatua galerita", "90) lorikeet", "91) coucal", "92) bee eater", "93) hornbill", "94) hummingbird", "95) jacamar", "96) toucan", "97) drake", "98) red-breasted merganser, Mergus serrator", "99) goose", "100) black swan, Cygnus atratus", "101) tusker", "102) echidna, spiny anteater, anteater", "103) platypus, duckbill, duckbilled platypus, duck-billed platypus, Ornithorhynchus anatinus", "104) wallaby, brush kangaroo", "105) koala, koala bear, kangaroo bear, native bear, Phascolarctos cinereus", "106) wombat", "107) jellyfish", "108) sea anemone, anemone", "109) brain coral", "110) flatworm, platyhelminth", "111) nematode, nematode worm, roundworm", "112) conch", "113) snail", "114) slug", "115) sea slug, nudibranch", "116) chiton, coat-of-mail shell, sea cradle, polyplacophore", "117) chambered nautilus, pearly nautilus, nautilus", "118) Dungeness crab, Cancer magister", "119) rock crab, Cancer irroratus", "120) fiddler crab", "121) king crab, Alaska crab, Alaskan king crab, Alaska king crab, Paralithodes camtschatica", "122) American lobster, Northern lobster, Maine lobster, Homarus americanus", "123) spiny lobster, langouste, rock lobster, crawfish, crayfish, sea crawfish", "124) crayfish, crawfish, crawdad, crawdaddy", "125) hermit crab", "126) isopod", "127) white stork, Ciconia ciconia", "128) black stork, Ciconia nigra", "129) spoonbill", "130) flamingo", "131) little blue heron, Egretta caerulea", "132) American egret, great white heron, Egretta albus", "133) bittern", "134) crane", "135) limpkin, Aramus pictus", "136) European gallinule, Porphyrio porphyrio", "137) American coot, marsh hen, mud hen, water hen, Fulica americana", "138) bustard", "139) ruddy turnstone, Arenaria interpres", "140) red-backed sandpiper, dunlin, Erolia alpina", "141) redshank, Tringa totanus", "142) dowitcher", "143) oystercatcher, oyster catcher", "144) pelican", "145) king penguin, Aptenodytes patagonica", "146) albatross, mollymawk", "147) grey whale, gray whale, devilfish, Eschrichtius gibbosus, Eschrichtius robustus", "148) killer whale, killer, orca, grampus, sea wolf, Orcinus orca", "149) dugong, Dugong dugon", "150) sea lion", "151) Chihuahua", "152) Japanese spaniel", "153) Maltese dog, Maltese terrier, Maltese", "154) Pekinese, Pekingese, Peke", "155) Shih-Tzu", "156) Blenheim spaniel", "157) papillon", "158) toy terrier", "159) Rhodesian ridgeback", "160) Afghan hound, Afghan", "161) basset, basset hound", "162) beagle", "163) bloodhound, sleuthhound", "164) bluetick", "165) black-and-tan coonhound", "166) Walker hound, Walker foxhound", "167) English foxhound", "168) redbone", "169) borzoi, Russian wolfhound", "170) Irish wolfhound", "171) Italian greyhound", "172) whippet", "173) Ibizan hound, Ibizan Podenco", "174) Norwegian elkhound, elkhound", "175) otterhound, otter hound", "176) Saluki, gazelle hound", "177) Scottish deerhound, deerhound", "178) Weimaraner", "179) Staffordshire bullterrier, Staffordshire bull terrier", "180) American Staffordshire terrier, Staffordshire terrier, American pit bull terrier, pit bull terrier", "181) Bedlington terrier", "182) Border terrier", "183) Kerry blue terrier", "184) Irish terrier", "185) Norfolk terrier", "186) Norwich terrier", "187) Yorkshire terrier", "188) wire-haired fox terrier", "189) Lakeland terrier", "190) Sealyham terrier, Sealyham", "191) Airedale, Airedale terrier", "192) cairn, cairn terrier", "193) Australian terrier", "194) Dandie Dinmont, Dandie Dinmont terrier", "195) Boston bull, Boston terrier", "196) miniature schnauzer", "197) giant schnauzer", "198) standard schnauzer", "199) Scotch terrier, Scottish terrier, Scottie", "200) Tibetan terrier, chrysanthemum dog", "201) silky terrier, Sydney silky", "202) soft-coated wheaten terrier", "203) West Highland white terrier", "204) Lhasa, Lhasa apso", "205) flat-coated retriever", "206) curly-coated retriever", "207) golden retriever", "208) Labrador retriever", "209) Chesapeake Bay retriever", "210) German short-haired pointer", "211) vizsla, Hungarian pointer", "212) English setter", "213) Irish setter, red setter", "214) Gordon setter", "215) Brittany spaniel", "216) clumber, clumber spaniel", "217) English springer, English springer spaniel", "218) Welsh springer spaniel", "219) cocker spaniel, English cocker spaniel, cocker", "220) Sussex spaniel", "221) Irish water spaniel", "222) kuvasz", "223) schipperke", "224) groenendael", "225) malinois", "226) briard", "227) kelpie", "228) komondor", "229) Old English sheepdog, bobtail", "230) Shetland sheepdog, Shetland sheep dog, Shetland", "231) collie", "232) Border collie", "233) Bouvier des Flandres, Bouviers des Flandres", "234) Rottweiler", "235) German shepherd, German shepherd dog, German police dog, alsatian", "236) Doberman, Doberman pinscher", "237) miniature pinscher", "238) Greater Swiss Mountain dog", "239) Bernese mountain dog", "240) Appenzeller", "241) EntleBucher", "242) boxer", "243) bull mastiff", "244) Tibetan mastiff", "245) French bulldog", "246) Great Dane", "247) Saint Bernard, St Bernard", "248) Eskimo dog, husky", "249) malamute, malemute, Alaskan malamute", "250) Siberian husky", "251) dalmatian, coach dog, carriage dog", "252) affenpinscher, monkey pinscher, monkey dog", "253) basenji", "254) pug, pug-dog", "255) Leonberg", "256) Newfoundland, Newfoundland dog", "257) Great Pyrenees", "258) Samoyed, Samoyede", "259) Pomeranian", "260) chow, chow chow", "261) keeshond", "262) Brabancon griffon", "263) Pembroke, Pembroke Welsh corgi", "264) Cardigan, Cardigan Welsh corgi", "265) toy poodle", "266) miniature poodle", "267) standard poodle", "268) Mexican hairless", "269) timber wolf, grey wolf, gray wolf, Canis lupus", "270) white wolf, Arctic wolf, Canis lupus tundrarum", "271) red wolf, maned wolf, Canis rufus, Canis niger", "272) coyote, prairie wolf, brush wolf, Canis latrans", "273) dingo, warrigal, warragal, Canis dingo", "274) dhole, Cuon alpinus", "275) African hunting dog, hyena dog, Cape hunting dog, Lycaon pictus", "276) hyena, hyaena", "277) red fox, Vulpes vulpes", "278) kit fox, Vulpes macrotis", "279) Arctic fox, white fox, Alopex lagopus", "280) grey fox, gray fox, Urocyon cinereoargenteus", "281) tabby, tabby cat", "282) tiger cat", "283) Persian cat", "284) Siamese cat, Siamese", "285) Egyptian cat", "286) cougar, puma, catamount, mountain lion, painter, panther, Felis concolor", "287) lynx, catamount", "288) leopard, Panthera pardus", "289) snow leopard, ounce, Panthera uncia", "290) jaguar, panther, Panthera onca, Felis onca", "291) lion, king of beasts, Panthera leo", "292) tiger, Panthera tigris", "293) cheetah, chetah, Acinonyx jubatus", "294) brown bear, bruin, Ursus arctos", "295) American black bear, black bear, Ursus americanus, Euarctos americanus", "296) ice bear, polar bear, Ursus Maritimus, Thalarctos maritimus", "297) sloth bear, Melursus ursinus, Ursus ursinus", "298) mongoose", "299) meerkat, mierkat", "300) tiger beetle", "301) ladybug, ladybeetle, lady beetle, ladybird, ladybird beetle", "302) ground beetle, carabid beetle", "303) long-horned beetle, longicorn, longicorn beetle", "304) leaf beetle, chrysomelid", "305) dung beetle", "306) rhinoceros beetle", "307) weevil", "308) fly", "309) bee", "310) ant, emmet, pismire", "311) grasshopper, hopper", "312) cricket", "313) walking stick, walkingstick, stick insect", "314) cockroach, roach", "315) mantis, mantid", "316) cicada, cicala", "317) leafhopper", "318) lacewing, lacewing fly", "319) dragonfly, darning needle, devil's darning needle, sewing needle, snake feeder, snake doctor, mosquito hawk, skeeter hawk", "320) damselfly", "321) admiral", "322) ringlet, ringlet butterfly", "323) monarch, monarch butterfly, milkweed butterfly, Danaus plexippus", "324) cabbage butterfly", "325) sulphur butterfly, sulfur butterfly", "326) lycaenid, lycaenid butterfly", "327) starfish, sea star", "328) sea urchin", "329) sea cucumber, holothurian", "330) wood rabbit, cottontail, cottontail rabbit", "331) hare", "332) Angora, Angora rabbit", "333) hamster", "334) porcupine, hedgehog", "335) fox squirrel, eastern fox squirrel, Sciurus niger", "336) marmot", "337) beaver", "338) guinea pig, Cavia cobaya", "339) sorrel", "340) zebra", "341) hog, pig, grunter, squealer, Sus scrofa", "342) wild boar, boar, Sus scrofa", "343) warthog", "344) hippopotamus, hippo, river horse, Hippopotamus amphibius", "345) ox", "346) water buffalo, water ox, Asiatic buffalo, Bubalus bubalis", "347) bison", "348) ram, tup", "349) bighorn, bighorn sheep, cimarron, Rocky Mountain bighorn, Rocky Mountain sheep, Ovis canadensis", "350) ibex, Capra ibex", "351) hartebeest", "352) impala, Aepyceros melampus", "353) gazelle", "354) Arabian camel, dromedary, Camelus dromedarius", "355) llama", "356) weasel", "357) mink", "358) polecat, fitch, foulmart, foumart, Mustela putorius", "359) black-footed ferret, ferret, Mustela nigripes", "360) otter", "361) skunk, polecat, wood pussy", "362) badger", "363) armadillo", "364) three-toed sloth, ai, Bradypus tridactylus", "365) orangutan, orang, orangutang, Pongo pygmaeus", "366) gorilla, Gorilla gorilla", "367) chimpanzee, chimp, Pan troglodytes", "368) gibbon, Hylobates lar", "369) siamang, Hylobates syndactylus, Symphalangus syndactylus", "370) guenon, guenon monkey", "371) patas, hussar monkey, Erythrocebus patas", "372) baboon", "373) macaque", "374) langur", "375) colobus, colobus monkey", "376) proboscis monkey, Nasalis larvatus", "377) marmoset", "378) capuchin, ringtail, Cebus capucinus", "379) howler monkey, howler", "380) titi, titi monkey", "381) spider monkey, Ateles geoffroyi", "382) squirrel monkey, Saimiri sciureus", "383) Madagascar cat, ring-tailed lemur, Lemur catta", "384) indri, indris, Indri indri, Indri brevicaudatus", "385) Indian elephant, Elephas maximus", "386) African elephant, Loxodonta africana", "387) lesser panda, red panda, panda, bear cat, cat bear, Ailurus fulgens", "388) giant panda, panda, panda bear, coon bear, Ailuropoda melanoleuca", "389) barracouta, snoek", "390) eel", "391) coho, cohoe, coho salmon, blue jack, silver salmon, Oncorhynchus kisutch", "392) rock beauty, Holocanthus tricolor", "393) anemone fish", "394) sturgeon", "395) gar, garfish, garpike, billfish, Lepisosteus osseus", "396) lionfish", "397) puffer, pufferfish, blowfish, globefish", "398) abacus", "399) abaya", "400) academic gown, academic robe, judge's robe", "401) accordion, piano accordion, squeeze box", "402) acoustic guitar", "403) aircraft carrier, carrier, flattop, attack aircraft carrier", "404) airliner", "405) airship, dirigible", "406) altar", "407) ambulance", "408) amphibian, amphibious vehicle", "409) analog clock", "410) apiary, bee house", "411) apron", "412) ashcan, trash can, garbage can, wastebin, ash bin, ash-bin, ashbin, dustbin, trash barrel, trash bin", "413) assault rifle, assault gun", "414) backpack, back pack, knapsack, packsack, rucksack, haversack", "415) bakery, bakeshop, bakehouse", "416) balance beam, beam", "417) balloon", "418) ballpoint, ballpoint pen, ballpen, Biro", "419) Band Aid", "420) banjo", "421) bannister, banister, balustrade, balusters, handrail", "422) barbell", "423) barber chair", "424) barbershop", "425) barn", "426) barometer", "427) barrel, cask", "428) barrow, garden cart, lawn cart, wheelbarrow", "429) baseball", "430) basketball", "431) bassinet", "432) bassoon", "433) bathing cap, swimming cap", "434) bath towel", "435) bathtub, bathing tub, bath, tub", "436) beach wagon, station wagon, wagon, estate car, beach waggon, station waggon, waggon", "437) beacon, lighthouse, beacon light, pharos", "438) beaker", "439) bearskin, busby, shako", "440) beer bottle", "441) beer glass", "442) bell cote, bell cot", "443) bib", "444) bicycle-built-for-two, tandem bicycle, tandem", "445) bikini, two-piece", "446) binder, ring-binder", "447) binoculars, field glasses, opera glasses", "448) birdhouse", "449) boathouse", "450) bobsled, bobsleigh, bob", "451) bolo tie, bolo, bola tie, bola", "452) bonnet, poke bonnet", "453) bookcase", "454) bookshop, bookstore, bookstall", "455) bottlecap", "456) bow", "457) bow tie, bow-tie, bowtie", "458) brass, memorial tablet, plaque", "459) brassiere, bra, bandeau", "460) breakwater, groin, groyne, mole, bulwark, seawall, jetty", "461) breastplate, aegis, egis", "462) broom", "463) bucket, pail", "464) buckle", "465) bulletproof vest", "466) bullet train, bullet", "467) butcher shop, meat market", "468) cab, hack, taxi, taxicab", "469) caldron, cauldron", "470) candle, taper, wax light", "471) cannon", "472) canoe", "473) can opener, tin opener", "474) cardigan", "475) car mirror", "476) carousel, carrousel, merry-go-round, roundabout, whirligig", "477) carpenter's kit, tool kit", "478) carton", "479) car wheel", "480) cash machine, cash dispenser, automated teller machine, automatic teller machine, automated teller, automatic teller, ATM", "481) cassette", "482) cassette player", "483) castle", "484) catamaran", "485) CD player", "486) cello, violoncello", "487) cellular telephone, cellular phone, cellphone, cell, mobile phone", "488) chain", "489) chainlink fence", "490) chain mail, ring mail, mail, chain armor, chain armour, ring armor, ring armour", "491) chain saw, chainsaw", "492) chest", "493) chiffonier, commode", "494) chime, bell, gong", "495) china cabinet, china closet", "496) Christmas stocking", "497) church, church building", "498) cinema, movie theater, movie theatre, movie house, picture palace", "499) cleaver, meat cleaver, chopper", "500) cliff dwelling", "501) cloak", "502) clog, geta, patten, sabot", "503) cocktail shaker", "504) coffee mug", "505) coffeepot", "506) coil, spiral, volute, whorl, helix", "507) combination lock", "508) computer keyboard, keypad", "509) confectionery, confectionary, candy store", "510) container ship, containership, container vessel", "511) convertible", "512) corkscrew, bottle screw", "513) cornet, horn, trumpet, trump", "514) cowboy boot", "515) cowboy hat, ten-gallon hat", "516) cradle", "517) crane", "518) crash helmet", "519) crate", "520) crib, cot", "521) Crock Pot", "522) croquet ball", "523) crutch", "524) cuirass", "525) dam, dike, dyke", "526) desk", "527) desktop computer", "528) dial telephone, dial phone", "529) diaper, nappy, napkin", "530) digital clock", "531) digital watch", "532) dining table, board", "533) dishrag, dishcloth", "534) dishwasher, dish washer, dishwashing machine", "535) disk brake, disc brake", "536) dock, dockage, docking facility", "537) dogsled, dog sled, dog sleigh", "538) dome", "539) doormat, welcome mat", "540) drilling platform, offshore rig", "541) drum, membranophone, tympan", "542) drumstick", "543) dumbbell", "544) Dutch oven", "545) electric fan, blower", "546) electric guitar", "547) electric locomotive", "548) entertainment center", "549) envelope", "550) espresso maker", "551) face powder", "552) feather boa, boa", "553) file, file cabinet, filing cabinet", "554) fireboat", "555) fire engine, fire truck", "556) fire screen, fireguard", "557) flagpole, flagstaff", "558) flute, transverse flute", "559) folding chair", "560) football helmet", "561) forklift", "562) fountain", "563) fountain pen", "564) four-poster", "565) freight car", "566) French horn, horn", "567) frying pan, frypan, skillet", "568) fur coat", "569) garbage truck, dustcart", "570) gasmask, respirator, gas helmet", "571) gas pump, gasoline pump, petrol pump, island dispenser", "572) goblet", "573) go-kart", "574) golf ball", "575) golfcart, golf cart", "576) gondola", "577) gong, tam-tam", "578) gown", "579) grand piano, grand", "580) greenhouse, nursery, glasshouse", "581) grille, radiator grille", "582) grocery store, grocery, food market, market", "583) guillotine", "584) hair slide", "585) hair spray", "586) half track", "587) hammer", "588) hamper", "589) hand blower, blow dryer, blow drier, hair dryer, hair drier", "590) hand-held computer, hand-held microcomputer", "591) handkerchief, hankie, hanky, hankey", "592) hard disc, hard disk, fixed disk", "593) harmonica, mouth organ, harp, mouth harp", "594) harp", "595) harvester, reaper", "596) hatchet", "597) holster", "598) home theater, home theatre", "599) honeycomb", "600) hook, claw", "601) hoopskirt, crinoline", "602) horizontal bar, high bar", "603) horse cart, horse-cart", "604) hourglass", "605) iPod", "606) iron, smoothing iron", "607) jack-o'-lantern", "608) jean, blue jean, denim", "609) jeep, landrover", "610) jersey, T-shirt, tee shirt", "611) jigsaw puzzle", "612) jinrikisha, ricksha, rickshaw", "613) joystick", "614) kimono", "615) knee pad", "616) knot", "617) lab coat, laboratory coat", "618) ladle", "619) lampshade, lamp shade", "620) laptop, laptop computer", "621) lawn mower, mower", "622) lens cap, lens cover", "623) letter opener, paper knife, paperknife", "624) library", "625) lifeboat", "626) lighter, light, igniter, ignitor", "627) limousine, limo", "628) liner, ocean liner", "629) lipstick, lip rouge", "630) Loafer", "631) lotion", "632) loudspeaker, speaker, speaker unit, loudspeaker system, speaker system", "633) loupe, jeweler's loupe", "634) lumbermill, sawmill", "635) magnetic compass", "636) mailbag, postbag", "637) mailbox, letter box", "638) maillot", "639) maillot, tank suit", "640) manhole cover", "641) maraca", "642) marimba, xylophone", "643) mask", "644) matchstick", "645) maypole", "646) maze, labyrinth", "647) measuring cup", "648) medicine chest, medicine cabinet", "649) megalith, megalithic structure", "650) microphone, mike", "651) microwave, microwave oven", "652) military uniform", "653) milk can", "654) minibus", "655) miniskirt, mini", "656) minivan", "657) missile", "658) mitten", "659) mixing bowl", "660) mobile home, manufactured home", "661) Model T", "662) modem", "663) monastery", "664) monitor", "665) moped", "666) mortar", "667) mortarboard", "668) mosque", "669) mosquito net", "670) motor scooter, scooter", "671) mountain bike, all-terrain bike, off-roader", "672) mountain tent", "673) mouse, computer mouse", "674) mousetrap", "675) moving van", "676) muzzle", "677) nail", "678) neck brace", "679) necklace", "680) nipple", "681) notebook, notebook computer", "682) obelisk", "683) oboe, hautboy, hautbois", "684) ocarina, sweet potato", "685) odometer, hodometer, mileometer, milometer", "686) oil filter", "687) organ, pipe organ", "688) oscilloscope, scope, cathode-ray oscilloscope, CRO", "689) overskirt", "690) oxcart", "691) oxygen mask", "692) packet", "693) paddle, boat paddle", "694) paddlewheel, paddle wheel", "695) padlock", "696) paintbrush", "697) pajama, pyjama, pj's, jammies", "698) palace", "699) panpipe, pandean pipe, syrinx", "700) paper towel", "701) parachute, chute", "702) parallel bars, bars", "703) park bench", "704) parking meter", "705) passenger car, coach, carriage", "706) patio, terrace", "707) pay-phone, pay-station", "708) pedestal, plinth, footstall", "709) pencil box, pencil case", "710) pencil sharpener", "711) perfume, essence", "712) Petri dish", "713) photocopier", "714) pick, plectrum, plectron", "715) pickelhaube", "716) picket fence, paling", "717) pickup, pickup truck", "718) pier", "719) piggy bank, penny bank", "720) pill bottle", "721) pillow", "722) ping-pong ball", "723) pinwheel", "724) pirate, pirate ship", "725) pitcher, ewer", "726) plane, carpenter's plane, woodworking plane", "727) planetarium", "728) plastic bag", "729) plate rack", "730) plow, plough", "731) plunger, plumber's helper", "732) Polaroid camera, Polaroid Land camera", "733) pole", "734) police van, police wagon, paddy wagon, patrol wagon, wagon, black Maria", "735) poncho", "736) pool table, billiard table, snooker table", "737) pop bottle, soda bottle", "738) pot, flowerpot", "739) potter's wheel", "740) power drill", "741) prayer rug, prayer mat", "742) printer", "743) prison, prison house", "744) projectile, missile", "745) projector", "746) puck, hockey puck", "747) punching bag, punch bag, punching ball, punchball", "748) purse", "749) quill, quill pen", "750) quilt, comforter, comfort, puff", "751) racer, race car, racing car", "752) racket, racquet", "753) radiator", "754) radio, wireless", "755) radio telescope, radio reflector", "756) rain barrel", "757) recreational vehicle, RV, R.V.", "758) reel", "759) reflex camera", "760) refrigerator, icebox", "761) remote control, remote", "762) restaurant, eating house, eating place, eatery", "763) revolver, six-gun, six-shooter", "764) rifle", "765) rocking chair, rocker", "766) rotisserie", "767) rubber eraser, rubber, pencil eraser", "768) rugby ball", "769) rule, ruler", "770) running shoe", "771) safe", "772) safety pin", "773) saltshaker, salt shaker", "774) sandal", "775) sarong", "776) sax, saxophone", "777) scabbard", "778) scale, weighing machine", "779) school bus", "780) schooner", "781) scoreboard", "782) screen, CRT screen", "783) screw", "784) screwdriver", "785) seat belt, seatbelt", "786) sewing machine", "787) shield, buckler", "788) shoe shop, shoe-shop, shoe store", "789) shoji", "790) shopping basket", "791) shopping cart", "792) shovel", "793) shower cap", "794) shower curtain", "795) ski", "796) ski mask", "797) sleeping bag", "798) slide rule, slipstick", "799) sliding door", "800) slot, one-armed bandit", "801) snorkel", "802) snowmobile", "803) snowplow, snowplough", "804) soap dispenser", "805) soccer ball", "806) sock", "807) solar dish, solar collector, solar furnace", "808) sombrero", "809) soup bowl", "810) space bar", "811) space heater", "812) space shuttle", "813) spatula", "814) speedboat", "815) spider web, spider's web", "816) spindle", "817) sports car, sport car", "818) spotlight, spot", "819) stage", "820) steam locomotive", "821) steel arch bridge", "822) steel drum", "823) stethoscope", "824) stole", "825) stone wall", "826) stopwatch, stop watch", "827) stove", "828) strainer", "829) streetcar, tram, tramcar, trolley, trolley car", "830) stretcher", "831) studio couch, day bed", "832) stupa, tope", "833) submarine, pigboat, sub, U-boat", "834) suit, suit of clothes", "835) sundial", "836) sunglass", "837) sunglasses, dark glasses, shades", "838) sunscreen, sunblock, sun blocker", "839) suspension bridge", "840) swab, swob, mop", "841) sweatshirt", "842) swimming trunks, bathing trunks", "843) swing", "844) switch, electric switch, electrical switch", "845) syringe", "846) table lamp", "847) tank, army tank, armored combat vehicle, armoured combat vehicle", "848) tape player", "849) teapot", "850) teddy, teddy bear", "851) television, television system", "852) tennis ball", "853) thatch, thatched roof", "854) theater curtain, theatre curtain", "855) thimble", "856) thresher, thrasher, threshing machine", "857) throne", "858) tile roof", "859) toaster", "860) tobacco shop, tobacconist shop, tobacconist", "861) toilet seat", "862) torch", "863) totem pole", "864) tow truck, tow car, wrecker", "865) toyshop", "866) tractor", "867) trailer truck, tractor trailer, trucking rig, rig, articulated lorry, semi", "868) tray", "869) trench coat", "870) tricycle, trike, velocipede", "871) trimaran", "872) tripod", "873) triumphal arch", "874) trolleybus, trolley coach, trackless trolley", "875) trombone", "876) tub, vat", "877) turnstile", "878) typewriter keyboard", "879) umbrella", "880) unicycle, monocycle", "881) upright, upright piano", "882) vacuum, vacuum cleaner", "883) vase", "884) vault", "885) velvet", "886) vending machine", "887) vestment", "888) viaduct", "889) violin, fiddle", "890) volleyball", "891) waffle iron", "892) wall clock", "893) wallet, billfold, notecase, pocketbook", "894) wardrobe, closet, press", "895) warplane, military plane", "896) washbasin, handbasin, washbowl, lavabo, wash-hand basin", "897) washer, automatic washer, washing machine", "898) water bottle", "899) water jug", "900) water tower", "901) whiskey jug", "902) whistle", "903) wig", "904) window screen", "905) window shade", "906) Windsor tie", "907) wine bottle", "908) wing", "909) wok", "910) wooden spoon", "911) wool, woolen, woollen", "912) worm fence, snake fence, snake-rail fence, Virginia fence", "913) wreck", "914) yawl", "915) yurt", "916) web site, website, internet site, site", "917) comic book", "918) crossword puzzle, crossword", "919) street sign", "920) traffic light, traffic signal, stoplight", "921) book jacket, dust cover, dust jacket, dust wrapper", "922) menu", "923) plate", "924) guacamole", "925) consomme", "926) hot pot, hotpot", "927) trifle", "928) ice cream, icecream", "929) ice lolly, lolly, lollipop, popsicle", "930) French loaf", "931) bagel, beigel", "932) pretzel", "933) cheeseburger", "934) hotdog, hot dog, red hot", "935) mashed potato", "936) head cabbage", "937) broccoli", "938) cauliflower", "939) zucchini, courgette", "940) spaghetti squash", "941) acorn squash", "942) butternut squash", "943) cucumber, cuke", "944) artichoke, globe artichoke", "945) bell pepper", "946) cardoon", "947) mushroom", "948) Granny Smith", "949) strawberry", "950) orange", "951) lemon", "952) fig", "953) pineapple, ananas", "954) banana", "955) jackfruit, jak, jack", "956) custard apple", "957) pomegranate", "958) hay", "959) carbonara", "960) chocolate sauce, chocolate syrup", "961) dough", "962) meat loaf, meatloaf", "963) pizza, pizza pie", "964) potpie", "965) burrito", "966) red wine", "967) espresso", "968) cup", "969) eggnog", "970) alp", "971) bubble", "972) cliff, drop, drop-off", "973) coral reef", "974) geyser", "975) lakeside, lakeshore", "976) promontory, headland, head, foreland", "977) sandbar, sand bar", "978) seashore, coast, seacoast, sea-coast", "979) valley, vale", "980) volcano", "981) ballplayer, baseball player", "982) groom, bridegroom", "983) scuba diver", "984) rapeseed", "985) daisy", "986) yellow lady's slipper, yellow lady-slipper, Cypripedium calceolus, Cypripedium parviflorum", "987) corn", "988) acorn", "989) hip, rose hip, rosehip", "990) buckeye, horse chestnut, conker", "991) coral fungus", "992) agaric", "993) gyromitra", "994) stinkhorn, carrion fungus", "995) earthstar", "996) hen-of-the-woods, hen of the woods, Polyporus frondosus, Grifola frondosa", "997) bolete", "998) ear, spike, capitulum", "999) toilet tissue, toilet paper, bathroom tissue"]
def interpolate_and_shape(A, B, num_interps):
interps = interpolate(A, B, num_interps)
return (interps.transpose(1, 0, *range(2, len(interps.shape)))
.reshape(num_samples * num_interps, *interps.shape[2:]))
z_A, z_B = [truncated_z_sample(num_samples, truncation, noise_seed)
for noise_seed in [noise_seed_A, noise_seed_B]]
y_A, y_B = [one_hot([int(category.split(')')[0])] * num_samples)
for category in [category_A, category_B]]
z_interp = interpolate_and_shape(z_A, z_B, num_interps)
y_interp = interpolate_and_shape(y_A, y_B, num_interps)
ims = sample(sess, z_interp, y_interp, truncation=truncation)
imshow(imgrid(ims, cols=num_interps))
```
| github_jupyter |
# Minimize beamwidth of an array with arbitrary 2-D geometry
A derivative work by Judson Wilson, 5/14/2014.<br>
Adapted (with significant changes) from the CVX example of the same name, by Almir Mutapcic, 2/2/2006.
Topic References:
* "Convex optimization examples" lecture notes (EE364) by S. Boyd
* "Antenna array pattern synthesis via convex optimization" by H. Lebret and S. Boyd
## Introduction
This algorithm designs an antenna array such that:
* it has unit sensitivity at some target direction
* it obeys a constraint on a minimum sidelobe level outside the beam
* it minimizes the beamwidth of the pattern.
This is a quasiconvex problem. Define the target direction as $\theta_{\mbox{tar}}$, and a beamwidth of $\Delta \theta_{\mbox{bw}}$. The beam occupies the angular interval
$$\Theta_b = \left(\theta_{\mbox{tar}}
-\frac{1}{2}\Delta \theta_{\mbox{bw}},\; \theta_{\mbox{tar}}
+ \frac{1}{2}\Delta \theta_{\mbox{bw}}\right).
$$
Solving for the minimum beamwidth $\Delta \theta_{\mbox{bw}}$ is performed by bisection, where the interval which contains the optimal value is bisected according to the result of the following feasibility problem:
\begin{array}{ll}
\mbox{minimize} & 0 \\
\mbox{subject to} & y(\theta_{\mbox{tar}}) = 1 \\
& \left|y(\theta)\right| \leq t_{\mbox{sb}}
\quad \forall \theta \notin \Theta_b.
\end{array}
$y$ is the antenna array gain pattern (a complex-valued function), $t_{\mbox{sb}}$ is the maximum allowed sideband gain threshold, and the variables are $w$ (antenna array weights or shading coefficients). The gain pattern is a linear function of $w$: $y(\theta) = w^T a(\theta)$ for some $a(\theta)$ describing the antenna array configuration and specs.
Once the optimal beamwidth is found, the solution $w$ is refined with the following optimization:
\begin{array}{ll}
\mbox{minimize} & \|w\| \\
\mbox{subject to} & y(\theta_{\mbox{tar}}) = 1 \\
& \left|y(\theta)\right| \leq t_{\mbox{sb}}
\quad \forall \theta \notin \Theta_b.
\end{array}
The implementation below discretizes the angular quantities and their counterparts, such as $\theta$.
## Problem specification and data
### Antenna array selection
Choose either:
* A random 2D positioning of antennas.
* A uniform 1D positioning of antennas along a line.
* A uniform 2D positioning of antennas along a grid.
```
import cvxpy as cp
import numpy as np
# Select array geometry:
ARRAY_GEOMETRY = '2D_RANDOM'
#ARRAY_GEOMETRY = '1D_UNIFORM_LINE'
#ARRAY_GEOMETRY = '2D_UNIFORM_LATTICE'
```
## Data generation
```
#
# Problem specs.
#
lambda_wl = 1 # wavelength
theta_tar = 0 # target direction
min_sidelobe = -20 # maximum sidelobe level in dB
max_half_beam = 50 # starting half beamwidth (must be feasible)
#
# 2D_RANDOM:
# n randomly located elements in 2D.
#
if ARRAY_GEOMETRY == '2D_RANDOM':
# Set random seed for repeatable experiments.
np.random.seed(1)
# Uniformly distributed on [0,L]-by-[0,L] square.
n = 36
L = 5
loc = L*np.random.random((n,2))
#
# 1D_UNIFORM_LINE:
# Uniform 1D array with n elements with inter-element spacing d.
#
elif ARRAY_GEOMETRY == '1D_UNIFORM_LINE':
n = 30
d = 0.45*lambda_wl
loc = np.hstack(( d * np.array(range(0,n)).reshape(-1, 1), \
np.zeros((n,1)) ))
#
# 2D_UNIFORM_LATTICE:
# Uniform 2D array with m-by-m element with d spacing.
#
elif ARRAY_GEOMETRY == '2D_UNIFORM_LATTICE':
m = 6
n = m**2
d = 0.45*lambda_wl
loc = np.zeros((n, 2))
for x in range(m):
for y in range(m):
loc[m*y+x,:] = [x,y]
loc = loc*d
else:
raise Exception('Undefined array geometry')
#
# Construct optimization data.
#
# Build matrix A that relates w and y(theta), ie, y = A*w.
theta = np.array(range(1, 360+1)).reshape(-1, 1)
A = np.kron(np.cos(np.pi*theta/180), loc[:, 0].T) \
+ np.kron(np.sin(np.pi*theta/180), loc[:, 1].T)
A = np.exp(2*np.pi*1j/lambda_wl*A)
# Target constraint matrix.
ind_closest = np.argmin(np.abs(theta - theta_tar))
Atar = A[ind_closest,:]
```
## Solve using bisection algorithm
```
# Bisection range limits. Reduce by half each step.
halfbeam_bot = 1
halfbeam_top = max_half_beam
print('We are only considering integer values of the half beam-width')
print('(since we are sampling the angle with 1 degree resolution).')
print('')
# Iterate bisection until 1 angular degree of uncertainty.
while halfbeam_top - halfbeam_bot > 1:
# Width in degrees of the current half-beam.
halfbeam_cur = np.ceil( (halfbeam_top + halfbeam_bot)/2.0 )
# Create optimization matrices for the stopband,
# i.e. only A values for the stopband angles.
upper_beam = (theta_tar + halfbeam_cur) % 360
lower_beam = (theta_tar - halfbeam_cur) % 360
if upper_beam > lower_beam:
ind = np.nonzero(np.squeeze(np.array(np.logical_or( \
theta <= lower_beam, \
theta >= upper_beam ))))
else:
ind = np.nonzero(np.squeeze(np.array(np.logical_and( \
theta <= lower_beam, \
theta >= upper_beam ))))
As = A[ind[0],:]
#
# Formulate and solve the feasibility antenna array problem.
#
# As of this writing (2014/05/14) cvxpy does not do complex valued math,
# so the real and complex values must be stored separately as reals
# and operated on as follows:
# Let any vector or matrix be represented as a+bj, or A+Bj.
# Vectors are stored [a; b] and matrices as [A -B; B A]:
# Atar as [A -B; B A]
Atar_R = Atar.real
Atar_I = Atar.imag
neg_Atar_I = -Atar_I
Atar_RI = np.block([[Atar_R, neg_Atar_I], [Atar_I, Atar_R]])
# As as [A -B; B A]
As_R = As.real
As_I = As.imag
neg_As_I = -As_I
As_RI = np.block([[As_R, neg_As_I], [As_I, As_R]])
As_RI_top = np.block([As_R, neg_As_I])
As_RI_bot = np.block([As_I, As_R])
# 1-vector as [1, 0] since no imaginary part
realones_ri = np.array([1.0, 0.0])
# Create cvxpy variables and constraints
w_ri = cp.Variable(shape=(2*n))
constraints = [ Atar_RI*w_ri == realones_ri]
# Must add complex valued constraint
# abs(As*w <= 10**(min_sidelobe/20)) row by row by hand.
# TODO: Future version use norms() or complex math
# when these features become available in cvxpy.
for i in range(As.shape[0]):
#Make a matrix whos product with w_ri is a 2-vector
#which is the real and imag component of a row of As*w
As_ri_row = np.vstack((As_RI_top[i, :], As_RI_bot[i, :]))
constraints.append( \
cp.norm(As_ri_row*w_ri) <= 10**(min_sidelobe/20) )
# Form and solve problem.
obj = cp.Minimize(0)
prob = cp.Problem(obj, constraints)
prob.solve(solver=cp.CVXOPT)
# Bisection (or fail).
if prob.status == cp.OPTIMAL:
print('Problem is feasible for half beam-width = {}'
' degress'.format(halfbeam_cur))
halfbeam_top = halfbeam_cur
elif prob.status == cp.INFEASIBLE:
print('Problem is not feasible for half beam-width = {}'
' degress'.format(halfbeam_cur))
halfbeam_bot = halfbeam_cur
else:
raise Exception('CVXPY Error')
# Optimal beamwidth.
halfbeam = halfbeam_top
upper_beam = (theta_tar + halfbeam) % 360
lower_beam = (theta_tar - halfbeam) % 360
print('Optimum half beam-width for given specs is {}'.format(halfbeam))
# Compute the minimum noise design for the optimal beamwidth
if upper_beam > lower_beam:
ind = np.nonzero(np.squeeze(np.array(np.logical_or( \
theta <= lower_beam, \
theta >= upper_beam ))))
else:
ind = np.nonzero(np.squeeze(np.array(np.logical_and( \
theta <= lower_beam, \
theta >= upper_beam ))))
As = A[ind[0],:]
# As as [A -B; B A]
# See earlier calculations for real/imaginary representation
As_R = As.real
As_I = As.imag
neg_As_I = -As_I
As_RI = np.block([[As_R, neg_As_I], [As_I, As_R]])
As_RI_top = np.block([As_R, neg_As_I])
As_RI_bot = np.block([As_I, As_R])
constraints = [ Atar_RI*w_ri == realones_ri]
# Same constraint as a above, on new As (hense different
# actual number of constraints). See comments above.
for i in range(As.shape[0]):
As_ri_row = np.vstack((As_RI_top[i, :], As_RI_bot[i, :]))
constraints.append( \
cp.norm(As_ri_row*w_ri) <= 10**(min_sidelobe/20) )
# Form and solve problem.
# Note the new objective!
obj = cp.Minimize(cp.norm(w_ri))
prob = cp.Problem(obj, constraints)
prob.solve(solver=cp.SCS)
#if prob.status != cp.OPTIMAL:
# raise Exception('CVXPY Error')
print("final objective value: {}".format(obj.value))
```
## Result plots
```
import matplotlib.pyplot as plt
# Show plot inline in ipython.
%matplotlib inline
# Plot properties.
plt.rc('text', usetex=True)
plt.rc('font', family='serif')
#
# First Figure: Antenna Locations
#
plt.figure(figsize=(6, 6))
plt.scatter(loc[:, 0], loc[:, 1],
s=30, facecolors='none', edgecolors='b')
plt.title('Antenna Locations', fontsize=16)
plt.tight_layout()
plt.show()
#
# Second Plot: Array Pattern
#
# Complex valued math to calculate y = A*w_im;
# See comments in code above regarding complex representation as reals.
A_R = A.real
A_I = A.imag
neg_A_I = -A_I
A_RI = np.block([[A_R, neg_A_I], [A_I, A_R]]);
y = A_RI.dot(w_ri.value)
y = y[0:int(y.shape[0]/2)] + 1j*y[int(y.shape[0]/2):] #now native complex
plt.figure(figsize=(6,6))
ymin, ymax = -40, 0
plt.plot(np.arange(360)+1, np.array(20*np.log10(np.abs(y))))
plt.plot([theta_tar, theta_tar], [ymin, ymax], 'g--')
plt.plot([upper_beam, upper_beam], [ymin, ymax], 'r--')
plt.plot([lower_beam, lower_beam], [ymin, ymax], 'r--')
plt.xlabel('look angle', fontsize=16)
plt.ylabel(r'mag $y(\theta)$ in dB', fontsize=16)
plt.ylim(ymin, ymax)
plt.tight_layout()
plt.show()
#
# Third Plot: Polar Pattern
#
plt.figure(figsize=(6,6))
zerodB = 50
dBY = 20*np.log10(np.abs(y)) + zerodB
plt.plot(dBY * np.cos(np.pi*theta.flatten()/180),
dBY * np.sin(np.pi*theta.flatten()/180))
plt.xlim(-zerodB, zerodB)
plt.ylim(-zerodB, zerodB)
plt.axis('off')
# 0 dB level.
plt.plot(zerodB*np.cos(np.pi*theta.flatten()/180),
zerodB*np.sin(np.pi*theta.flatten()/180), 'k:')
plt.text(-zerodB,0,'0 dB', fontsize=16)
# Max sideband level.
m=min_sidelobe + zerodB
plt.plot(m*np.cos(np.pi*theta.flatten()/180),
m*np.sin(np.pi*theta.flatten()/180), 'k:')
plt.text(-m,0,'{:.1f} dB'.format(min_sidelobe), fontsize=16)
#Lobe center and boundaries angles.
theta_1 = theta_tar+halfbeam
theta_2 = theta_tar-halfbeam
plt.plot([0, 55*np.cos(theta_tar*np.pi/180)], \
[0, 55*np.sin(theta_tar*np.pi/180)], 'k:')
plt.plot([0, 55*np.cos(theta_1*np.pi/180)], \
[0, 55*np.sin(theta_1*np.pi/180)], 'k:')
plt.plot([0, 55*np.cos(theta_2*np.pi/180)], \
[0, 55*np.sin(theta_2*np.pi/180)], 'k:')
#Show plot.
plt.tight_layout()
plt.show()
```
| github_jupyter |
This notebook is a step by step guide about how to train a deep neural network (DNN) in the DeepDeconv framework.
```
## Set up the sys.path in order to be able to import our modules
import os
import sys
module_path = os.path.abspath(os.path.join('..'))
if module_path not in sys.path:
sys.path.append(module_path)
import keras.utils
## extra imports to set GPU options
import tensorflow as tf
from keras import backend as k
###################################
# TensorFlow wizardry
config = tf.ConfigProto()
# Don't pre-allocate memory; allocate as-needed
config.gpu_options.allow_growth = True
# Only allow a total of half the GPU memory to be allocated
# This line is optional, don't add it unless you really need to set a limit on the memory available for your process
# For instance, if you want to train 2 DNNs on the same GPU without one overlapping the memory needed by the other
# Change the value to set the percentage of memory allocated
config.gpu_options.per_process_gpu_memory_fraction = 0.5
# Create a session with the above options specified.
k.tensorflow_backend.set_session(tf.Session(config=config))
# Now you can create/load your DNN
#from DeepDeconv.deepnetFCS.DeconvNet import DeconvNet
from deepnetFCS.DeconvNet_custom import UNet2D
nb_scales = 3 #4
nb_layers_per_block = [4,4,4] #[2,2]#,2] #[4,5,6,7]
nb_filters=16
activation_function= 'relu' #'swish'
resNet=False
layer_string='layer{0}'.format(nb_layers_per_block[0])
atrou=True
for k in range(1,len(nb_layers_per_block)):
layer_string+='x{0}'.format(nb_layers_per_block[k])
network_name='UNet2D_FCS_sc{0}_{1}_{2}_filt{3}'.format(nb_scales,layer_string,activation_function,nb_filters)
if atrou:
network_name+='_atrou'
if resNet:
network_name+='_resNet'
print("Network Name:",network_name)
dnn = UNet2D(network_name = network_name, img_rows = 96, img_cols = 96, model_file='', verbose=True,
filters=nb_filters,nb_scales=nb_scales, nb_layers_per_block=nb_layers_per_block,
activation_function=activation_function,resNet=resNet,atrou=atrou)
from keras.utils import plot_model
plot_model(dnn.model, to_file='{0}.png'.format(network_name),show_shapes=True)
from IPython.display import SVG
from keras.utils.vis_utils import model_to_dot
pydot_obj=model_to_dot(dnn.model,show_shapes=True,show_layer_names=False)
SVG(pydot_obj.create(prog='dot', format='svg'))
```
## PREPROCESSING OF DATA
```
# from astropy.io import fits as fits
# from matplotlib import pyplot as plt
# #Input the directory containing the fits file
# data_directory = '/data/DeepDeconv/data/vsc_euclidpsfs/'
# #Retrieves the list of all the files
# import glob
# gal_files = glob.glob(data_directory+'image-0*-0.fits')
# gal_files.sort()
# print(gal_files)
# import numpy as np
# from utils.batch_utils import shuffle_new_fits
# data_reshfl_dir=data_directory+"reshuffle/"
# noiseless_img_hdu = 1
# targets_hdu = 4
# psf_hdu = 3
# rootname_reshfl=data_reshfl_dir+"image-shfl"
# np.random.seed(2012)
# shuffle_new_fits(gal_files, nb_img_per_file=10000,
# noiseless_img_hdu=noiseless_img_hdu, targets_hdu=targets_hdu, psf_hdu=psf_hdu,
# image_dim=96, image_per_row=100,rootname=rootname_reshfl)
# from astropy.io import fits as fits
# from matplotlib import pyplot as plt
# ff=fits.open(gal_files[0])
# plt.figure()
# for k in range(5):
# plt.subplot(2,3,k+1),plt.imshow(ff[k].data[0:96,0:96])
# #HDU 0: noisy
# #HDU 1: noise free convolved with euclid PSF
# #HDU 2: noise free convolved with large PSF (gauss 0.15 FWHM)
# #HDU 3: euclid PSF
# #HDU 4: noise free convolved with target PSF (gauss 0.07 FWHM)
# plt.figure()
# plt.imshow(ff[1].data[0:96,0:96]-ff[4].data[0:96,0:96])
```
## Training Phase
```
import glob
import numpy as np
from astropy.io import fits as fits
from matplotlib import pyplot as plt
data_directory = '/data/DeepDeconv/data/vsc_euclidpsfs/'
SNR = [20,100]
if len(SNR)==1:
old_dnn_name=dnn.network_name
dnn.network_name+='_reshfl_SNR{0}'.format(SNR[0])
else:
old_dnn_name=dnn.network_name
dnn.network_name+='_reshfl_SNR{0}to{1}'.format(SNR[0],SNR[1])
noiseless_img_hdu = 0
psf_hdu = 1
targets_hdu = 2
deconv_mode = 'TIKHONOV'
data_reshfl_dir=data_directory+"reshuffle/"
gal_files_shfl = glob.glob(data_reshfl_dir+'image-shfl-*-multihdu.fits')
gal_files_shfl.sort()
print(gal_files_shfl)
print(dnn.network_name)
#Train with the image-000-0.fits as validation and all the other files as training set
dnn.train_generator(gal_files_shfl[2:], gal_files_shfl[1], epochs=20, batch_size=32,
nb_img_per_file=10000, validation_set_size=10000,
noise_std=None, SNR=SNR, model_file='',
noiseless_img_hdu=noiseless_img_hdu, targets_hdu=targets_hdu, psf_hdu=psf_hdu,
image_dim=96, image_per_row=100,
deconv_mode=deconv_mode)
dnn.network_name=old_dnn_name
len(SNR)
dnn.network_name
#
#SNR=100
# noiseless_img_hdu = 1
# targets_hdu = 4
# psf_hdu = 3
# deconv_mode = 'TIKHONOV'
# #Train with the image-000-0.fits as validation and all the other files as training set
# dnn.train_generator(gal_files[2:], gal_files[1], epochs=20, batch_size=32,
# nb_img_per_file=10000, validation_set_size=10000,
# noise_std=None, SNR=SNR, model_file='',
# noiseless_img_hdu=noiseless_img_hdu, targets_hdu=targets_hdu, psf_hdu=psf_hdu,
# image_dim=96, image_per_row=100,
# deconv_mode=deconv_mode)
#The train_generator is:
#1) running get_batch_from_fits for validation data: read files, deconv if necessary, return as [ngal,X2D,Y2D,1]
#2) setting a checkpoint for model, saving the model if lower validation loss
#3) using a generator function to obtain dynamic batches: dynamic_batches
# that I modified because it was assuming nb_img_per_file to be 10000 (hardcoded)
#4) running fit_generator with logging and checkpoint callbacks
#I modified
```
| github_jupyter |
----
# Getting started with Intel Quantum Simulator
----
Tutorial on the basic use of Intel QS through its Python interface.
**NOTE:**
Currently, the Python implementation only allows for single-core execution and does not take advantages of the MPI protocol.
However the user can familiarize with the same functionalities available in the distributed implementation (only C++ at the moment) and the transition should be relatively straighforward since all methods maintain name and effect.
### Import Intel QS library
Let's start by importing the Python library with the class and methods defined in the C++ implementation.
```
# Import the Python library with the C++ class and methods of Intel Quantum Simulator.
# If the library is not contained in the same folder of this notebook, its path has to be added.
import sys
sys.path.insert(0, '../build/lib')
import intelqs_py as simulator
# import numPy
import numpy as np
# Import graphical library for plots.
import matplotlib.pyplot as plt
```
### Initialize the state of the quantum register
IQS stores a full representation of the quantum state in the computational basis.
In practice, the quantum state of $N$ qubits is represented as a complex vector with $2^N$ components.
Each component corresponds to the probability amplitude of a specific computational basis state:
$$\psi(k) = \langle k | \psi \rangle$$
with the index $k$ corresponding to the $N$-bit integer in decimal representation, and $k\in\{0,1,2,\dots,2^N-1\}$.
----
- First of all, one needs to allocate the memory to contain the state representation.
- Then the quantum register has to be initialized, either to a specific computational basis state (using the keyword "base") or to a random state (using the keyword "rand").
----
NOTE: the random number generator is able to generate three different kinds of random numbers:
- *local* --> different for each pool rank
- *state* --> common to all ranks of the same state
- *pool* --> common to all ranks of the pool
```
# Number of qubits.
num_qubits = 2;
# Index of the computational basis state corresponding to |00>.
index = 0;
# Allocate memory for the quantum register's state and initialize it to |00>.
psi = simulator.QubitRegister(num_qubits, "base", index, 0);
# To initialize the state to a random vector, one first need a random number generator.
# Create the random number generator, set its seed and then associate it to the IQS object 'psi'.
rng = simulator.RandomNumberGenerator();
rng_seed = 7777;
rng.SetSeedStreamPtrs( rng_seed );
psi.SetRngPtr(rng);
# Initialize the state to a random state, this can be achieved with the codeword "rand" followed by 0
# if we desire to use *local* random numbers (this speed up the process of generating the random numbers).
psi.Initialize("rand", 0);
```
### Display the quantum state
It is important to be able to access and visualize the quantum state.
IQS allows to access the single components of the state or to print a comprehensive description.
What index is associated to state $|1011\rangle$?
In decimal representation one has:
$$1011 \rightarrow 1\times2^0 + 0\times2^1 + 1\times2^2 + 1\times2^3 = 1+4+8 = 13$$
**NOTE:** contrary to what is adopted in decimal notation, our binary representation must be read from left to right (from least significant to most significant bit).
```
# Initialize the state to |10>.
# The index of |10> in decimal representation is 1.
index = 1;
psi.Initialize("base", index);
# There are for amplitudes, corresponding to |00>, |10>, |01>, |11>.
for index in range(0,2**num_qubits):
amplitude = psi[index]
print("psi({}) = <{}|psi> = {}".format(index,index,amplitude))
# A complete description of the state is provided by the method Print().
print("----")
label = "Computational basis state |10>"
psi.Print(label)
```
### One-qubit gates
In the gate-model of quantum computation, one manipulates the quantum state by means of unitary transformations acting on one or two qubits. Let us apply a few of the standard one-qubit gates.
```
# State was |10>. Let us re-prepare it:
psi = simulator.QubitRegister(2, "base", 1, 0);
# Flip the qubit 1 by applying the Pauli X gate: |10> ==> |11>
qubit = 1;
psi.ApplyPauliX(qubit);
# Display all amplitudes.
print("Currently, |psi>=|11>:");
for index in range(0,2**num_qubits):
print(" psi({}) = <{}|psi> = {}".format(index,index,psi[index]))
print("----")
# Apply the Hadamard gate on qubit 0: |11> ==> |-1> ~ |01>-|11>
qubit = 0;
psi.ApplyHadamard(qubit);
# Display all amplitudes.
print("Currently, |psi>=|-1>:");
for index in range(0,2**num_qubits):
print(" psi({}) = <{}|psi> = {}".format(index,index,psi[index]))
# Apply Pauli Z gate on qubit 1: |-1> ==> -|-1>
psi.ApplyPauliZ(1);
# Apply Pauli X gate on qubit 0: -|-1> ==> |-1>
psi.ApplyPauliX(0);
```
### Two-qubit gates
To achieve universal quantum computation, it is enought to implement one-qubit gates and a single type of two-qubit gate.
The essential requirement is that such two-qubit gate is able to generate entanglement. Usually the controlled-not gate (CNOT in the following) is the operation of choice.
IQS provides built-in methods to implement a much broader variety of two-qubit gates.
```
# Currently, state is |-1>.
# Apply a CNOT(1,0): flip qubit 0 conditioned on the state of qubit 1.
# |-1> ==> -|-1>
control = 1;
target = 0;
psi.ApplyCPauliX(control, target);
# Display all amplitudes.
print("Currently, |psi>=-|-1>:");
for index in range(0,2**num_qubits):
print(" psi({}) = <{}|psi> = {}".format(index,index,psi[index]))
print("----")
# The application of the previous CNOT did not create any entanglement.
# This is achieved by exchanging the role of control and target qubits.
# Apply a CNOT(0,1): flip qubit 1 conditioned on the state of qubit 0.
# -|-1> ~ -|01>+|11> ==> -|01>+|10>
control = 0;
target = 1;
psi.ApplyCPauliX(control, target);
# Display all amplitudes.
print("Currently, |psi>=(|10>-|01>)/sqrt(2):");
for index in range(0,2**num_qubits):
print(" psi({}) = <{}|psi> = {}".format(index,index,psi[index]))
```
### Custom gates
If IQS does not provide the gates needed in your circuit, it is possible to implement custom one-qubit gates and controlled gates.
```
# Define an arbitrary single qubit gate.
# The quantum gate G is given by a 2x2 unitary matrix, here using a bi-dimensional NumPy array.
G = np.zeros((2,2),dtype=np.complex_);
G[0,0] = 0.592056606032915 + 0.459533060553574j;
G[0,1] = -0.314948020757856 - 0.582328159830658j;
G[1,0] = 0.658235557641767 + 0.070882241549507j;
G[1,1] = 0.649564427121402 + 0.373855203932477j;
# To verify that G is unitary, we will compute the norm of psi before and after the application of G.
initial_norm = psi.ComputeNorm();
if initial_norm != 1:
print("Even before the application of G, state psi had normalization {}".format(initial_norm));
# Apply the custom gate G to qubit 0.
qubit = 0;
psi.Apply1QubitGate(qubit,G);
final_norm = psi.ComputeNorm();
if initial_norm != final_norm:
print("The application of G changed the norm of state psi: from {} to {}".format(initial_norm,final_norm));
else:
print("Sanity check: norm was unchanged by G.");
# It is also possible to apply the arbitrary gate specified by G conditioned on the state of another qubit.
# G is applied only when the control qubit is in |1>.
control = 1;
target = 0;
psi.ApplyControlled1QubitGate( control, target, G);
# Notice that this output is directed to the terminal and not re-directed to the iPython notebook.
psi.Print("State of the quantum register after all gates.")
print()
# To display the amplitudes in the iPython notebook:
for index in range(0,2**num_qubits):
print("psi({}) = <{}|psi> = {}".format(index,index,psi[index]))
```
### Single-qubit measurements
To extract information from the quantum register, one can obtain the probability of measuring a certain qubit in the computational basis and obtaining the outcome "1" (i.e. the state is in $|1\rangle$).
Once the probability is known, one can draw a random number to simulate the stochastic outcome of the measurement and collapse the wavefunction accordingly.
**NOTE:**
Computing the probability of a certain outcome does not collapse automatically the wavefunction. This is helpful when the probabilities of multiple measurements have to be computed without re-executing the quantum simulation.
```
# Compute the probability of qubit 1 being in state |1>.
measured_qubit = 1;
prob = psi.GetProbability( measured_qubit );
print("Probability that qubit {} is in state |1> is {}\n".format(measured_qubit, prob));
# Draw random number in [0,1)
r = np.random.rand()
if r < prob:
# Collapse the wavefunction according to qubit 1 being in |1>.
print("Simulated outcome is 1. Collapse the function accordingly.")
psi.CollapseQubit(measured_qubit,True);
else:
# Collapse the wavefunction according to qubit 1 being in |0>
print("Simulated outcome is 0. Collapse the function accordingly.")
psi.CollapseQubit(measured_qubit,False);
# In both cases one needs to re-normalize the wavefunction:
psi.Normalize();
```
### Expectation value of products of Pauli matrices
To extract information from the quantum register, one can obtain the expectation value of Pauli strings.
For example, consider the Pauli string given by: $$X_0 \otimes id_1 \otimes Z_2 \otimes Z_3$$
Such observable is defined by:
- the position of the non-trivial Pauli matrices, in this case {0,2,3}
- the corresponding Pauli matrices ($X$=1, $Y$=2, $Z$=3).
To facilitate the verification of the expectation value, we reinitialize the quantum state to $|+-01\rangle$.
We also consider the Pauli string $$X_0 \otimes id_1 \otimes Z_2 \otimes Y_3$$.
```
# Prepare the state |+-01>
num_qubits = 4;
index = 0;
psi = simulator.QubitRegister(num_qubits, "base", index, 0);
psi.ApplyPauliX(1);
psi.ApplyPauliX(3);
psi.ApplyHadamard(0);
psi.ApplyHadamard(1);
print("psi is in state |+-01>\n");
# The Pauli string given by: X_0 . id_1 . Z_2 . Z_3
# Such observable is defined by the position of the non-trivial Pauli matrices:
qubits_to_be_measured = [0,2,3]
# And by the corresponding Pauli matrices (X=1, Y=2, Z=3)
observables = [1,3,3]
# The expectation value <psi|X_0.id_1.Z_2.Z_3|psi> is obtained via:
average = psi.ExpectationValue(qubits_to_be_measured, observables, 1.);
print("Expectation value <psi|X_0.id_1.Z_2.Z_3|psi> = {} <== it should be -1\n".format(average));
# The expectation value <psi|X_0.id_1.Z_2.Y_3|psi> is obtained via:
observables = [1,3,2]
average = psi.ExpectationValue(qubits_to_be_measured, observables, 1.);
print("Expectation value <psi|X_0.id_1.Z_2.Y_3|psi> = {} <== it should be 0\n".format(average));
```
### Examples of state preparation
Let us prepare the state $|+-01\rangle$.
```
# Method A:
# Prepare the state |0000>, flip qubits {1,3}, change basis to qubits {0,1}.
num_qubits = 4;
index = 0;
psi = simulator.QubitRegister(num_qubits, "base", index, 0);
psi.ApplyPauliX(1);
psi.ApplyPauliX(3);
psi.ApplyHadamard(0);
psi.ApplyHadamard(1);
# Method B:
# Prepare the state |0000>, change basis to qubits {0,1}, flip qubit {3}, flip in X qubit {1}.
index = 0;
psi.Initialize("base", index);
psi.ApplyHadamard(0);
psi.ApplyHadamard(1);
psi.ApplyPauliZ(1);
psi.ApplyPauliX(3);
# Method C:
# Prepare the computational state |0101>, change basis to qubits {0,1}.
index = 2+8 ;
psi.Initialize("base", index);
# Notice that GetProbability() does not change the state.
print("Verify that the state is now |0101>.\n")
for qubit in range(0,num_qubits):
prob = psi.GetProbability( qubit );
print("Probability that qubit {}, if measured, is in state |1> = {}".format(qubit, prob));
psi.ApplyHadamard(0);
psi.ApplyHadamard(1);
print("\nNow the state is |+-01>.\n")
# The expectation value <psi|X_0.X_1.Z_2.Z_3|psi> is obtained via:
qubits_to_be_measured = [0,1,2,3]
observables = [1,1,3,3]
average = psi.ExpectationValue(qubits_to_be_measured, observables, 1.);
print("Expectation value <psi|X_0.X_1.Z_2.Z_3|psi> = {} <== it should be +1\n".format(average));
```
----
## END
----
| github_jupyter |
```
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
%matplotlib inline
import matplotlib
matplotlib.rcParams["figure.figsize"] = (20, 10)
df = pd.read_csv("Bengaluru_House_Data.csv")
df.head()
df.shape
```
# Data cleaning
```
df.groupby('area_type')['area_type'].agg('count')
df = df.drop(['area_type', 'society', 'balcony', 'availability'], axis=1)
df.head()
df.isnull().sum()
df = df.dropna()
df.isnull().sum()
df['size'].unique()
df['bhk'] = df['size'].apply(lambda x: int(x.split(' ')[0]))
df.head()
df.total_sqft.unique()
def is_float(x):
try:
float(x)
except:
return False
return True
df[~df['total_sqft'].apply(is_float)].head(10)
def convert_sqft_to_num(x):
tokens = x.split('-')
if len(tokens) == 2:
return (float(tokens[0]) + float(tokens[1]))/2
try:
return float(x)
except:
return None
df['total_sqft'] = df['total_sqft'].apply(convert_sqft_to_num)
df.head()
```
# feature engineering
```
df['price_per_sqft'] = df['price']*100000/df['total_sqft']
df.head()
len(df.location.unique())
```
## data points per location
```
df.location = df.location.apply(lambda x: x.strip())
location_stats = df.groupby('location')['location'].agg('count').sort_values(ascending=False)
location_stats
len(df.location.unique())
loc_less_than_10 = location_stats[location_stats <= 10]
df.location = df.location.apply(lambda x: 'other' if x in loc_less_than_10 else x)
len(df.location.unique())
df.head(10)
```
# outlier removal
```
df[df.total_sqft/df.bhk<300].head(10)
df = df[~(df.total_sqft/df.bhk<300)]
df.price_per_sqft.describe()
def remove_pps_outliers(data_fr):
df_out = pd.DataFrame()
for key, subdf in data_fr.groupby('location'):
m = np.mean(subdf.price_per_sqft)
st = np.std(subdf.price_per_sqft)
reduced_df = subdf[(subdf.price_per_sqft>(m-st)) & (subdf.price_per_sqft<=(m+st))]
df_out = pd.concat([df_out, reduced_df], ignore_index=True)
return df_out
df1 = remove_pps_outliers(df)
df1.shape
def plot_chart(df_in, location):
bhk2 = df[(df.location==location) & (df.bhk==2)]
bhk3 = df[(df.location==location) & (df.bhk==3)]
matplotlib.rcParams['figure.figsize'] = (15,10)
plt.scatter(bhk2.total_sqft,bhk2.price,color='blue',label='2 BHK', s=50)
plt.scatter(bhk3.total_sqft,bhk3.price,marker='+', color='green',label='3 BHK', s=50)
plt.xlabel("Total Square Feet Area")
plt.ylabel("Price (Lakh Indian Rupees)")
plt.title(location)
plt.legend()
plot_chart(df1, "Rajaji Nagar")
def remove_bhk_outliers(df):
exclude_indices = np.array([])
for location, location_df in df.groupby('location'):
bhk_stats = {}
for bhk, bhk_df in location_df.groupby('bhk'):
bhk_stats[bhk] = {'mean': np.mean(bhk_df.price_per_sqft),
'std': np.std(bhk_df.price_per_sqft),
'count': bhk_df.shape[0]
}
for bhk, bhk_df in location_df.groupby('bhk'):
stats = bhk_stats.get(bhk-1)
if stats and stats['count']>5:
exclude_indices = np.append(exclude_indices, bhk_df[bhk_df.price_per_sqft<(stats['mean'])].index.values)
return df.drop(exclude_indices,axis='index')
df2 = remove_bhk_outliers(df1)
plot_chart(df2, "Hebbal")
df2.bath.unique()
df2 = df2[df2.bath<df2.bhk+2]
df2.shape
df3 = df2.drop(['size', 'price_per_sqft'], axis='columns')
df3.head()
dummies = pd.get_dummies(df3.location)
dummies.head()
df4 = pd.concat([df3, dummies.drop('other', axis='columns')], axis='columns')
df4.head()
df5 = df4.drop('location', axis='columns')
df5.head()
df5.shape
X = df5.drop('price', axis=1)
X.head()
y = df5.price
y.head()
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=10)
from sklearn.linear_model import LinearRegression
lr = LinearRegression()
lr.fit(X_train, y_train)
lr.score(X_test, y_test)
from sklearn.model_selection import ShuffleSplit
from sklearn.model_selection import cross_val_score
cv = ShuffleSplit(n_splits=5, test_size=0.2, random_state=0)
cross_val_score(LinearRegression(), X, y, cv=cv)
from sklearn.model_selection import GridSearchCV
from sklearn.linear_model import Lasso
from sklearn.tree import DecisionTreeRegressor
def find_best_model_using_gridsearchcv(X,y):
algos = {
'linear_regression' : {
'model': LinearRegression(),
'params': {
'normalize': [True, False]
}
},
'lasso': {
'model': Lasso(),
'params': {
'alpha': [1,2],
'selection': ['random', 'cyclic']
}
},
'decision_tree': {
'model': DecisionTreeRegressor(),
'params': {
'criterion' : ['mse','friedman_mse'],
'splitter': ['best','random']
}
}
}
scores = []
cv = ShuffleSplit(n_splits=5, test_size=0.2, random_state=0)
for algo_name, config in algos.items():
gs = GridSearchCV(config['model'], config['params'], cv=cv, return_train_score=False)
gs.fit(X,y)
scores.append({
'model': algo_name,
'best_score': gs.best_score_,
'best_params': gs.best_params_
})
return pd.DataFrame(scores,columns=['model','best_score','best_params'])
find_best_model_using_gridsearchcv(X,y)
def predict_price(location,sqft,bath,bhk):
loc_index = np.where(X.columns==location)[0][0]
x = np.zeros(len(X.columns))
x[0] = sqft
x[1] = bath
x[2] = bhk
if loc_index >= 0:
x[loc_index] = 1
return lr.predict([x])[0]
predict_price('1st Phase JP Nagar',1000, 2, 2)
import pickle
with open('real_estate_prices_model.pickle','wb') as f:
pickle.dump(lr,f)
import json
columns = {
'data_columns' : [col.lower() for col in X.columns]
}
with open("columns.json","w") as f:
f.write(json.dumps(columns))
```
| github_jupyter |
```
from context import *
```
# Pauli Algebra (`paulialg`)
## Basic Usage
### Operator Construction
#### Pauli Operator
A **Pauli operator** can be constructed using the `pauli()` constructor.
```
2**20
pauli('XXIYZ')
```
**Specify the Phase Factor**
By default the operator has a $+1$ phase factor in the front. To specify other phase factors($\pm1$ or $\pm \mathrm{i}$), use `'+'`, `'-'`, `'i'` indicators before the Pauli string.
```
pauli('-X'), pauli('iX'), pauli('-iX')
```
It is also possible to assign the phase factor by scalar mutiplication.
```
-pauli('X'), 1j*pauli('X')
```
**Specify the Pauli String**
Other methods to specify a Pauli string:
* construct from a tuple / list / array of indices (`0` = `I`, `1` = `X`, `2` = `Y`, `3` = `Z`)
```
pauli((0,1,2,3)), pauli([0,1,2,3]), pauli(numpy.array([0,1,2,3]))
```
* construct from a dictionary that maps positions to indices. (*Note*: using this method must also provide the total number of qubits as the second argument, because the qubit number can not be infered from the dictionary alone.)
```
pauli({1:'X', 4:'Y', 5:'Z'}, 6), pauli({1:1, 4:2, 5:3}, 6),
```
#### Pauli Operator List
A **list of Pauli operators** can be constructed by the `paulis()` constructor.
```
paulis('X', '-Y', 'Z')
```
It can take a generator and iterate through its elements to construct a list of Pauli operators.
```
paulis(pauli({i:'Z'}, 4) for i in range(4))
```
It can also take a iterable (tuple / list / set) and convert it to a list of Pauli operators.
```
objs = ['XX', 'YY', 'ZZ']
paulis(objs)
```
#### Size Information
For Pauli operator, `.N` returns the number of qubits (size of system) that the operator acts on.
```
pauli('IIII').N
```
For Pauli operator list, `.L` returns the number of operators in the list and `.N` returns of the number fo qubits in the system.
```
plst = paulis('II','XX','YY','ZZ')
plst.L, plst.N
len(plst)
```
#### Selection and Slicing
Select a single element in the Pauli operator list.
```
plst[1]
```
Select a range of operators in the Pauli operator list.
```
plst[0:3]
plst[-2:]
plst[::2]
```
It is also allow to be selected by a index array or a boolean mask.
```
plst[numpy.array([2,1,1,0,3])]
plst[numpy.array([True,False,False,True])]
```
### Operator Algebra
#### Scalar Product
Pauli operator and operator list can be multiplied with a scalar.
* If the scalar is a phase factor (as power of imaginary unit), the phase will be multiplied to the operator.
```
-pauli('X'), 1j*pauli('X')
```
For operator list, the scalar multiplication broadcast to every operator in the list.
```
-paulis('II','XX','YY','ZZ')
```
* if the scalar is beyond a phase factor, the Pauli operator will be promoted to a **Poly monomial** (i.e. Pauli operator equipted with a complex coefficient.)
```
2*pauli('X'), -1.5*pauli('X'), (-0.3+0.4j)*pauli('X')
```
However, Pauli opreator list does not support scalar multiplication beyond the four phase factors. (Because there is no canonical meanding for a list of Pauli monomials).
#### Linear Combination
Pauli operators can be linearly combined in to a **Pauli polynomial**.
```
pauli('XX') + pauli('YY') - 0.5 * pauli('ZZ')
```
Adding Pauli operators with any number, the number will be promoted to the number times identity operator automatically. For example, a projection operator can be written as
```
(pauli('ZZ') + 1)/2
```
Operators can be summed up with python built-in function `sum()`.
```
sum(paulis('II','XX','YY','ZZ'))
```
#### Dot Productor (Matrix Multiplication)
Dot productor (composition) of Pauli operators is implemented as the matrix multiplication `matmul`, which can be implemented using the operand `@`.
```
pauli('X') @ pauli('Z')
pauli('X') @ pauli('Y'), pauli('Y') @ pauli('X')
```
The dot product of two Pauli operators is still a Pauli operator. However if any one of them is Pauli monomial, the result will also be Pauli monomial.
```
(3*pauli('X')) @ pauli('X'), (2*pauli('X')) @ (3*pauli('Z'))
```
Dot product of Pauli polynomials will be expanded.
```
poly = pauli('XX') + pauli('YY') - 0.5 * pauli('ZZ')
poly @ poly
```
Terms will not be combined automatically. To combine them, the `.reduce()` method should be explicitly called.
```
(poly @ poly).reduce()
```
This allows the user to get control of the reduction, to avoid unnecessary reductions in the intermediate calculation.
#### Identity and Zero
Identity and zero operators can be constructed by `pauli_identity(N)` and `pauli_zero(N)` given the qubit number `N`.
```
pauli_identity(5), pauli_zero(5)
```
This makes it convinient to refer to these operators in the calculation.
```
pauli_zero(5) + pauli('XXXXX'), pauli_identity(5) @ pauli('XXXXX')
```
### Properties and Type Conversion
There are four different types of objects involved in the above discussion.
* `Pauli` (base class): a Pauli operator (in the Pauli group).
* `PauliMonomial` (subclass of `Pauli`): a Pauli operator with general coefficient (outside the Pauli group).
* `PauliList` (base class): a list of Pauli operators.
* `PauliPolynomial` (subclass of `PauliList`): a linear combination of Pauli operators (stored as Pauli operator list together with combination coefficients.)
```
type(pauli('X')), type(2*pauli('X')), type(paulis('X','X')), type(sum(paulis('X','X')))
```
#### Properties
As subclasses, `PauliMonomial` and `PauliPolynomial` inherit the related size properties from their parent classes.
```
(5*pauli('XYZ')).N
poly = sum(sum(paulis('II','XX','YY','ZZ')))
poly, poly.L, poly.N
```
`PauliPolynomial` can be selected and sliced as a list.
```
poly[1], poly[:2], poly[::2]
poly[numpy.array([1,1,1,2])].reduce()
poly[numpy.array([True,False,False,True])]
```
#### Type Conversion
`Pauli` can be converted to `PauliMonomial`.
```
pauli('XX').as_monomial()
```
`Pauli`, `PauliList`, `PauliMonomial` can all be converted to `PauliPolynomial`.
```
pauli('XX').as_polynomial()
paulis('II','XX','YY','ZZ').as_polynomial()
```
Automatic type conversion enables the algebra to be carried out among different classes with great flexibiliity.
* When `Pauli` is multiplied (`*`) by a generic number (beyond powers of the imaginary unit), it is converted to `PauliMonomial`.
* When `Pauli` or `PauliMonomial` is added (`+`) or subtracted (`-`) with other Pauli objects, they are converted to `PauliPolynomial`.
* The dot product (`@`) generally returns `PauliPolynomial`, unless the two Pauli objects are both `Pauli`, in which case it returns `Pauli`.
### Clifford Transformation
`PauliList` provides useful methods to implement Clifford transformations efficiently on all Pauli operators together. The same methods are available to all its subclasses (including `PauliPolynomial`, `CliffordMap`, `StabilizerState`).
#### Clifford Rotation
A Clifford rotation is a $\mathbb{Z}_4$ rotation in the Clifford group generated by a single Pauli operator, which takes the form of
$$U=e^{\frac{i\pi}{4}\sigma}=\frac{1}{\sqrt{2}}(1+i \sigma).$$
Every Pauli operator is transformed by $\sigma \to U^\dagger \sigma U$. The Clifford rotation can be applied by the method `.rotate_by(gen)` (given the generator `gen`). The operation is in-place (meaning that the operators in the Pauli list will be modified).
```
paulis('II','XX','YY','ZZ').rotate_by(pauli('XI'))
```
#### Clifford Map
A Clifford map is a generic clifford transformation by specifying how each single Pauli operator gets mapped to. It can be listed as a table
```
cmap = random_clifford_map(2)
cmap
```
It can be applied by the method `.transform_by(cmap)` (given the Clifford map `cmap`).
```
paulis('II','XX','YY','ZZ').transform_by(cmap)
```
#### Masked Transformation
Clifford transformation can be applied to a subsystem of qubits specified by a mask.
```
mask = numpy.array([True,False,False,True])
paulis('IIII','XXXX','YYYY','ZZZZ').rotate_by(pauli('XY'), mask)
mask = numpy.array([True,False,False,True])
paulis('IIII','XXXX','YYYY','ZZZZ').transform_by(cmap, mask)
```
## Algorithm Details
### Internal Representation
#### Binary Representation of Pauli Operators
Any Pauli operator can be specified by two one-hot (binary) vectors $x$ and $z$ ($x_i,z_i=0,1$ for $i=1,\cdots,N$):
$$\sigma_{(x,z)}=\mathrm{i}^{x\cdot z}\prod_{i=1}^{N}X_i^{x_i}\prod_{i=1}^{N}Z_i^{z_i}.$$
* The binary vector $x$ (or $z$) specifies the qubits where the $X$ (or $Z$) operator acts ($Y$ operator acts at where $X$ and $Z$ act simultaneously).
* **Multiplication** of two Pauli operators
$$\sigma_{(x,z)}\sigma_{(x',z')}=\mathrm{i}^{p(x,z;x',z')}\sigma_{(x+x',z+z')\%2},$$
where the power $p$ of $\mathrm{i}$ in the prefactor is given by
$$p(x,z;x',z')=\sum_{i=1}^{N}\left(z_ix'_i-x_iz'_i + 2(z_i+z'_i)\left\lfloor\frac{x_i+x'_i}{2}\right\rfloor+2(x_i+x'_i)\left\lfloor\frac{z_i+z'_i}{2}\right\rfloor\right)\mod 4.$$
* **Commutation relation**: two Pauli operator either commute to anticommute.
$$\sigma_{(x,z)}\sigma_{(x',z')}=(-)^{c(x,z;x',z')}\sigma_{(x',z')}\sigma_{(x,z)},$$
where the *anticommutation indicator* $c$ has a simpler form
$$c(x,z;x',z')=\frac{p(x,z;x',z')-p(x',z';x,z)}{2}=\sum_{i=1}^{N}\left(z_ix'_i-x_iz'_i\right)\mod 2.$$
The binary vectors $x$ and $z$ can be interweaved into a $2N$-component vector $g=(x_0,z_0,x_1,z_1,\cdots)$, which forms the binary representation of a Pauli operator $\sigma_g$.
#### `Pauli` Class
`Pauli(g,p)` represents a Pauli operator.
**Parameters:**
* `g` binary representation of Pauli string.
* `p` phase indicator ($p=0,1,2,3$ stands for $i^p$ phase factor).
```
pauli('iX').__dict__
```
#### `PauliList` Class
`PauliList(gs,ps)` represents a list of Pauli operators.
**Parameters:**
* `gs` array of binary representations of Pauli strings.
* `ps` array of phase indicators ($p=0,1,2,3$ stands for $i^p$ phase factor).
```
paulis('XX','YY','ZZ').__dict__
```
#### `PauliMonomial` Class
`PauliMonomial(g,p)` represents a Pauli operator with coefficient.
**Parameters:**
* `g` binary representation of Pauli string.
* `p` phase indicator ($p=0,1,2,3$ stands for $i^p$ phase factor).
* `c` coefficient (complex).
```
pauli('iX').as_monomial().__dict__
```
The property `c` can be set by the method `.set_c(c)`
```
pauli('X').as_monomial().set_c(2.+0.j).__dict__
```
#### `PauliPolynomial` Class
`PauliPolynomial(gs,ps)` represents a polynomial (linear combination) of Pauli operators.
**Parameters:**
* `gs` array of binary representations of Pauli strings.
* `ps` array of phase indicators ($p=0,1,2,3$ stands for $i^p$ phase factor).
* `cs` array of coefficients (complex).
```
(pauli('XX') - 2*pauli('YY')).__dict__
```
The property `cs` can be set by the method `.set_cs(cs)`
| github_jupyter |
```
# TODO:
1. Improve stitch algorithm
a. filtering??? KF
b. use lane information to restrict measurements
2. visualization - space-time diagram
a. need lane information - try clustering
b. need distance from lane start point
# save df to csv
from utils import*
import os.path
from os import path
data_path = pathlib.Path().absolute().joinpath('tracking_outputs')
output_path = data_path.joinpath('stitched')
file_name = 'record_p3c6_00000_track_outputs.csv'
file_dir = data_path.joinpath(file_name)
new_dir = output_path.joinpath('stitched_'+file_name)
df = read_data(file_dir)
# df = read_new_data(new_dir)
df = calc_distance(df, file_name)
df
df=df.iloc[::5, :]
# visualize by lane
import mplcursors
N = 40000;
fig, ax = plt.subplots(figsize=(10,6))
%pylab
mplcursors.cursor(hover=True)
plt.scatter(x=df.lat, y=df.lon, s=1, alpha=.8)
plt.plot([36.001777, 36.000354], [-86.606115, -86.604256], 'ro-')
plt.plot([36.001765, 36.000319], [-86.606154, -86.604287], 'ro-')
plt.plot([36.001751, 36.000256], [-86.606196, -86.604268], 'ro-')
plt.plot([36.001738, 36.000224], [-86.606235, -86.604283], 'ro-')
plt.plot([36.001669, 36.000145], [-86.606354, -86.604334], 'ro-')
plt.plot([36.001666, 36.000121], [-86.606400, -86.604366], 'ro-')
plt.plot([36.001661, 36.000105], [-86.606452, -86.604397], 'ro-')
plt.plot([36.001646, 36.000084], [-86.606495, -86.604429], 'ro-')
plt.xlabel('Latitude', fontsize = 20)
plt.ylabel('longitude', fontsize = 20)
plt.xlim(df['lat'].min()-0.0005,df['lat'].max()+0.0005)
plt.ylim(df['lon'].min()-0.0005,df['lon'].max()+0.0005)
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5), fontsize = 10)
ax = plt.gca()
ax.format_coord = lambda x,y: '%.6f, %.6f' % (x,y)
plt.show()
draw_map(df,36.0032, -86.607580, 100)
import importlib
import utils
importlib.reload(utils)
df = utils.calc_distance(df, file_name)
groups = df.groupby('lane')
groupList = list(groups.groups)
# plt.figure(figsize=(40, 40))
fig, ax = plt.subplots(figsize=(8,6))
start = 5
end = 60
for i in groupList:
group = groups.get_group(i)
plt.scatter(x=group.lat, y=group.lon, s=1, alpha=.8, label=group.loc[group.index[0],'lane'])
# print(groups.get_group(groupList[end]).Timestamp.values[-1]-groups.get_group(groupList[start]).Timestamp.values[0])
# plt.plot([36.00295, 36.00348], [-86.60749, -86.60806], 'ro-')
# plt.plot([36.00293, 36.00346], [-86.60754, -86.60810], 'ro-')
# plt.plot([36.00291, 36.003441], [-86.607575, -86.60813], 'ro-')
# plt.plot([36.002885, 36.003415], [-86.6076, -86.60818], 'ro-')
# plt.plot([36.00282, 36.0033], [-86.60768, -86.6082], 'ro-')
# plt.plot([36.00279, 36.00323], [-86.60774, -86.6082], 'ro-')
plt.xlabel('Latitude', fontsize = 20)
plt.ylabel('longitude', fontsize = 20)
plt.xlim(df['lat'].min(),df['lat'].max())
plt.ylim(df['lon'].min()-0.0001,df['lon'].max()+0.0001)
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5), fontsize = 10)
plt.show()
# extract lane info (most importantly, the GPS location of the boundary of the bbx)
startpts = np.array([[36.00295, -86.60749],
[36.00293, -86.60754],
[36.00291, -86.607575],
[36.002885, -86.6076],
[36.00282, -86.60768],
[36.00279, -86.60774]
])
endpts = np.array([[36.00348, -86.60806],
[36.00346, -86.60810],
[36.003441, -86.60813],
[36.003415, -86.60818],
[36.0033, -86.6082],
[36.00323, -86.6082]
])
```
| github_jupyter |
```
%matplotlib widget
import os
import sys
sys.path.insert(0, os.getenv('HOME')+'/pycode/MscThesis/')
import pandas as pd
from amftrack.util import get_dates_datetime, get_dirname, get_plate_number, get_postion_number,get_begin_index
import ast
from amftrack.plotutil import plot_t_tp1, plot_node_skel
from scipy import sparse
from datetime import datetime
from amftrack.pipeline.functions.node_id import orient
import pickle
import scipy.io as sio
from pymatreader import read_mat
from matplotlib import colors
import cv2
import imageio
import matplotlib.pyplot as plt
import numpy as np
from skimage.filters import frangi
from skimage import filters
from random import choice
import scipy.sparse
import os
from amftrack.pipeline.functions.extract_graph import from_sparse_to_graph, generate_nx_graph, sparse_to_doc
from skimage.feature import hessian_matrix_det
from amftrack.pipeline.functions.experiment_class_surf import Experiment, Edge, Node, Hyphae, plot_raw_plus
from amftrack.pipeline.paths.directory import run_parallel, find_state, directory_scratch, directory_project
from amftrack.notebooks.analysis.util import *
from scipy import stats
from scipy.ndimage.filters import uniform_filter1d
from statsmodels.stats import weightstats as stests
from amftrack.pipeline.functions.hyphae_id_surf import get_pixel_growth_and_new_children
from collections import Counter
from IPython.display import clear_output
from amftrack.notebooks.analysis.data_info import *
lapse = 60
exp = get_exp((38,131,131+lapse),directory_project)
plate = 38
begin = 131
end = 191
directory = directory_project
dates_datetime = get_dates_datetime(directory,plate)
dates_datetime.sort()
dates_datetime_chosen = dates_datetime[begin : end + 1]
dates = dates_datetime_chosen
exp = Experiment(plate, directory)
exp.load(dates)
plt.close('all')
directory = directory_project
begin = 15
end = 30
for t in range(begin,end):
save=f'Figure/im{t}'
plot_node_skel(node,t,save = save)
for t in range(begin,end):
img = cv2.imread(f'Figure/im{t}.png')
height, width, layers = img.shape
size = (width,height)
img_array.append(img)
out = cv2.VideoWriter('Figure/plate38.mp4',cv2.VideoWriter_fourcc(*'DIVX'), 2, size)
for i in range(len(img_array)):
out.write(img_array[i])
out.release()
node = get_rh_from_label(939,exp).end
make_video_node(node,15,30,anchor = 23)
img_array = []
for t in range(begin,end):
img = cv2.imread(f'Figure/im{t}.png')
height, width, layers = img.shape
size = (width,height)
img_array.append(img)
out = cv2.VideoWriter('Figure/plate38.mp4',cv2.VideoWriter_fourcc(*'DIVX'), 2, size)
for i in range(len(img_array)):
out.write(img_array[i])
out.release()
def make_video_node(node,t0,t1,anchor = None):
for t in range(t0,t1):
save=f'Figure/im{t}'
plot_node_skel(node,t,save = save, anchor =anchor)
img_array = []
for t in range(t0,t1):
img = cv2.imread(f'Figure/im{t}.png')
height, width, layers = img.shape
size = (width,height)
img_array.append(img)
out = cv2.VideoWriter(f'Figure/{node.experiment.plate}_{node.label}.mp4',cv2.VideoWriter_fourcc(*'DIVX'), 2, size)
for i in range(len(img_array)):
out.write(img_array[i])
out.release()
```
| github_jupyter |
# Problem 1
## 30 pts
- Prove that $\mathrm{vec}(AXB) = (B^\top \otimes A)\, \mathrm{vec}(X)$ if $\mathrm{vec}(X)$ is a columnwise reshape of a matrix into a long vector. What does it change if the reshape is rowwise? **Note:** to make a columnwise reshape in Python one should use ```np.reshape(X, order='f')```, where the string ```'f'``` stands for the Fortran ordering.
- Let $A$ and $B$ be dense $n\times n$ matrices. Suggest an algorithm that calculates matrix-vector multiplication $(A\otimes B)x$ faster than in $\mathcal{O}(n^4)$ operations.
- Let matrices $A$ and $B$ have eigendecompositions $A = S_A\Lambda_A S_A^{-1}$ and $B = S_B\Lambda_B S^{-1}_B$. Find eigenvectors and eigenvalues of the matrix $A\otimes I + I \otimes B$.
- Let $x_{k+1} = x_{k} - \tau_k (Ax_k - f)$, $x_0$ is zero vector. Does $x_{k}\in \mathcal{K}_{k}(A,f)$ - the Krylov subspace? Can CG be represented in this form (i.e. there exists $\tau_k$ such that $x_{k+1}$ generated by the sequence above and the CG method are equal for any $k$)?
- Let $A = \mathrm{diag}\left(\frac{1}{1000},\frac{1}{999},\dots \frac{1}{2}, 1, 1000 \right)$. Estimate analytically the number of iterations required to solve linear system with $A$ with the relative accuracy $10^{-4}$ using
- Richardson iteration with the optimal choice of parameter
- Chebyshev iteration
- Conjugate gradient method
# Problem 2
## 40 pts
### Sparse systems
Consider a 2D Poisson equation in $\Omega = [0,1]^2$
$$
\Delta u \equiv \frac{\partial^2 u}{\partial x^2} + \frac{\partial^2 u}{\partial y^2} = f(x,y), \quad (x,y)\in \Omega
$$
with zero Dirichlet boundary conditions
$$
u_{\partial \Omega} = 0,
$$
with known function $f(x,y)$ and unknown $u(x,y)$.
To find solution of the Poisson equation we will use the **finite difference method**. Standard second order finite difference discretization on a uniform grid $(x_i, y_j) = (ih, jh)$, $i,j = 0,\dots, N$, $h = \frac{1}{N}$ leads to the following system of equations:
$$
\begin{split}
&\frac{u_{i+1,j} - 2u_{i,j} + u_{i-1,j}}{h^2} + \frac{u_{i,j+1} - 2u_{i,j} + u_{i,j-1}}{h^2} = f(ih, jh) \\
&u_{0,j} = u_{i,0} = u_{N,j} = u_{i,N} = 0, \quad i,j = 0,\dots,N
\end{split}
$$
* Write the system above as a matrix equation $BU_h + U_h C = F_h$ with matrices $U_h = \begin{bmatrix}u_{1,1} & \dots & u_{1,N-1} \\ \vdots & \ddots & \vdots \\ u_{N-1,1} & \dots & u_{N-1,N-1} \end{bmatrix}$, $F_h = \begin{bmatrix}f_{1,1} & \dots & f_{1,N-1} \\ \vdots & \ddots & \vdots \\ f_{N-1,1} & \dots & f_{N-1,N-1} \end{bmatrix}$. What are matrices $B$ and $C$? Show that they are negative definite.
* Using Kronecker product properties rewrite $ BU_h + U_h C = F_h$ as $A_h \mathrm{vec}(U_h) = \mathrm{vec}(F_h)$, where $\mathrm{vec}(\cdot)$ is a columnwise reshape.
What is matrix $A_h$?
* Choose $f(x,y)$ such that $u(x, y) = \sin\pi x \sin \pi y$ is a solution (just substitute $u$ in the Poisson equation and find $f$, then pretend as if you do not know the solution $u$). Solve the system with the found $f$ using the `scipy.sparse.linalg.spsolve` which is direct sparse solver. Use ```pandas``` library and print table that contains $N$, time, relative error between the analytic solution and the obtained one for $N=128,256,512$. Matrices $B, C$ and $A_h$ should be assembled in the `CSR` format using functions from the `scipy.sparse` package (functions `scipy.sparse.kron` and `scipy.sparse.spdiags` will be helpful). <font color='red'> Do not use full matrices! Use only sparse arithmetics. </font>
* What is the iterative method of choice among `cg`, `minres`, `GMRES`, `BicgStab`? Explain why.
* Run the method from the previous task with and without ILU0 preconditioner for $N=256$. Plot relative error w.r.t. iteration number for both cases on one plot.
### Eigenvalues
* Find $3$ smallest eigenvalues of matrices $B$ and $C$ using ```scipy.sparse.linalg.eigs``` (Implicitly Restarted Arnoldi Method) or if $B$ and $C$ are Hermitian using ```scipy.sparse.linalg.eigsh``` (Implicitly Restarted Lanczos Method). Print them.
* What are the first $3$ smallest distinct eigenvalues of $A_h$ in terms of eigenvalues of $B$ and $C$? What are their multiplicities (explain the answer)? Find these eigenvalues numerically using ```scipy.sparse.linalg.eigsh``` and compare them with what you have found using eigenvalues of $B$ and $C$.
* **Bonus:** Find analytically eigenvalues of the matrix $A_h$ and prove that $\text{cond}( A_h )= \mathcal{O}\left(\frac{1}{h^2}\right)$
# Problem 3
## 30 pts
### Structured matrices
* Find convolution of the Lena image $n\times n$, $n=512$ with the following filter
$$
T_{i_1j_1,i_2j_2} \equiv T_{i_1-j_1,i_2-j_2} = \frac{\alpha}{\pi} e^{-\alpha \left[(i_1 - j_1)^2 + (i_2 - j_2)^2 \right]}, \quad i_1,j_1, i_2, j_2 = 1,\dots, n, \quad \alpha = \frac{1}{100}
$$
using FFT. What is the complexity of this operation? Plot the result as an image.
* Write matvec function that produces multiplication of $T$ by a given vector $x$. Use `scipy.sparse.linalg.LinearOperator` to create an object that has attribute `.dot()` (this object will be further used in the iterative process). Note that `.dot()` input and output must be 1D vectors, so do not forget to use reshape.
* Run an appropriate Krylov method with the obtained Linear Operator and try to reconstruct Lena using the right-hand side from the first bullet (smoothed Lena). On one figure plot relative error w.r.t. the number of iterations for $\alpha=\frac{1}{50},\frac{1}{100},\frac{1}{200}$ and the corresponding right-hand side. Comment on the results.
* **Bonus:** Let $x\in\mathbb{R}^{n^2}$ being reshaped into a matrix of size $n\times n$ have rank $r$. Let also $T_{i_1j_1,i_2j_2} \equiv \widetilde T_{i_1-j_1,i_2-j_2}$ such that $n\times n$ matrix $\widetilde T_{i_1,i_2}$ has rank $R$. Propose an algorithm that calculates $Tx$ using $\mathcal{O}((r+R)n\log n + rRn)$ operations.
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
%matplotlib inline
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.cluster import KMeans
from sklearn.svm import SVC
from sklearn.decomposition import PCA
from sklearn import metrics
# from mlxtend.plotting import plot_decision_regions
from sklearn import preprocessing
from sklearn.linear_model import LogisticRegression
from ast import literal_eval
import warnings
import numpy as np
from collections import OrderedDict
from lob_data_utils import lob, db_result, model, roc_results
from lob_data_utils.svm_calculation import lob_svm
import os
sns.set_style('whitegrid')
warnings.filterwarnings('ignore')
data_length = 15000
rs_params = [(0.1, 1.0), (1.0, 0.1), (1.0, 1.0), (0.1, 0.1)]
stocks = list(roc_results.result_cv_15000.keys())
data_dir = 'res_pca_gdf3_15000'
def get_mean_scores(scores: dict) -> dict:
mean_scores = {}
for k, v in scores.items():
mean_scores[k] = np.mean(v)
return mean_scores
def get_score_for_clf(clf, df_test):
x_test = df_test[['queue_imbalance']]
y_test = df_test['mid_price_indicator'].values
return model.test_model(clf, x_test, y_test)
def get_logistic_regression(stock, data_length):
df, df_test = lob.load_prepared_data(
stock, data_dir='../gaussian_filter/data', cv=False, length=data_length)
clf = LogisticRegression()
train_x = df[['queue_imbalance']]
scores = model.train_model(clf, train_x, df['mid_price_indicator'], prefix='log')
res = {
**scores,
'stock': stock,
'kernel': 'logistic',
}
test_scores = get_score_for_clf(clf, df_test)
return {**res, **test_scores}
df_res = pd.DataFrame()
for stock in stocks:
for r, s in rs_params:
#pd.read_csv('svm_features_{}_len{}_r{}_s{}.csv'.format(stock, data_length, r, s))
filename = data_dir + '/svm_pca_gdf_rbf_{}_len{}_r{}_s{}.csv'.format(stock, data_length, r, s)
if os.path.exists(filename):
df_temp = pd.read_csv(filename)
df_temp['r'] = [r] * len(df_temp)
df_temp['s'] = [s] * len(df_temp)
df_res = df_res.append(df_temp)
#df_res.drop(columns=['Unnamed: 0'], inplace=True)
columns = ['C', 'f1', 'features', 'gamma', 'kappa',
'matthews', 'roc_auc', 'stock',
'test_f1', 'test_kappa', 'test_matthews', 'test_roc_auc', 'r', 's']
df_res[columns].sort_values(by='roc_auc', ascending=False).groupby('stock').head(1)
log_res = []
for stock in stocks:
log_res.append(get_logistic_regression(stock, data_length))
df_log_res = pd.DataFrame(log_res)
df_log_res['stock'] = df_log_res['stock'].values.astype(np.int)
df_log_res.index = df_log_res['stock'].values.astype(np.int)
df_gdf_best = df_res[columns].sort_values(by='matthews', ascending=False).groupby('stock').head(1)
df_gdf_best['stock'] = df_gdf_best['stock'].values.astype(np.int)
df_gdf_best.index = df_gdf_best['stock'].values.astype(np.int)
df_all = pd.merge(df_gdf_best, df_log_res, on='stock', suffixes=['_svm', '_log'])
df_all.rename(columns=
{
'matthews': 'matthews_svm',
'log_matthews': 'matthews_log',
'roc_auc': 'roc_auc_svm',
'log_roc_auc': 'roc_auc_log',
'f1': 'f1_svm',
'log_f1': 'f1_log',
}, inplace=True)
df_all.columns
all_columns = [ 'features', 'matthews_svm', 'matthews_log', 'test_matthews_svm', 'test_matthews_log',
'roc_auc_svm', 'roc_auc_log', 'test_roc_auc_svm', 'test_roc_auc_log', 'stock',
'f1_svm', 'f1_log', 'test_f1_svm', 'test_f1_log', 'r', 's']
df_all[all_columns]
df_all['matthews_diff'] = df_all['matthews_svm'] - df_all['matthews_log']
df_all['matthews_test_diff'] = df_all['test_matthews_svm'] - df_all['test_matthews_log']
result_cv_15000 = {'9069': 0.56266492905467125, '9063': 0.55890612062034239,
'9094': 0.56271758343003708,
'10166': 0.54209270809674659, '9061': 0.53172059475198441,
'9064': 0.5445462725690029,
'11244': 0.53511679644048948, '9062': 0.54049738307164052,
'9067': 0.54498044328552808,
'9265': 0.55580632143346032, '9058': 0.53573417928037304,
'9269': 0.54865753911806547,
'12059': 0.55367480325463525}
stocks15000 = np.array(list(result_cv_15000.keys())).astype(np.int)
df_all.loc[df_all[df_all['stock'].isin(stocks15000)]['matthews_diff'].nlargest(n=2).index]
sns.distplot(df_all['matthews_diff'], label='training')
sns.distplot(df_all['matthews_test_diff'], label='testing')
plt.legend()
sns.distplot(df_all['matthews_svm'], label='svm')
sns.distplot(df_all['matthews_log'], label='log')
plt.legend()
df_all['rs'] = list(zip(df_all['r'], df_all['s']))
df_all['rs'].value_counts().plot(kind='bar', alpha=0.5, stacked=False)
df_all['matthews_diff'].median(), df_all['matthews_test_diff'].median()
len(df_all[df_all['matthews_svm'] > df_all['matthews_log']][all_columns]), len(df_all[df_all['test_matthews_svm'] > df_all['test_matthews_log']]), len(df_all)
df_all[df_all['test_matthews_svm'] < df_all['test_matthews_log']][all_columns]
len(df_all[df_all['roc_auc_svm'] > df_all['roc_auc_log']][all_columns]), len(df_all[df_all['test_roc_auc_svm'] > df_all['test_roc_auc_log']][all_columns]), len(df_all)
df_all[df_all['test_matthews_svm'] < df_all['test_matthews_log']][all_columns]
df_all[df_all['test_roc_auc_svm'] < df_all['test_roc_auc_log']][all_columns]
```
| github_jupyter |
# FloPy
## Creating a Simple MODFLOW 6 Model with Flopy
The purpose of this notebook is to demonstrate the Flopy capabilities for building a simple MODFLOW 6 model from scratch, running the model, and viewing the results. This notebook will demonstrate the capabilities using a simple lake example. A separate notebook is also available in which the same lake example is created for MODFLOW-2005 (flopy3_lake_example.ipynb).
### Setup the Notebook Environment
```
import sys
import os
import platform
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
# run installed version of flopy or add local path
try:
import flopy
except:
fpth = os.path.abspath(os.path.join('..', '..'))
sys.path.append(fpth)
import flopy
print(sys.version)
print('numpy version: {}'.format(np.__version__))
print('matplotlib version: {}'.format(mpl.__version__))
print('flopy version: {}'.format(flopy.__version__))
# For this example, we will set up a model workspace.
# Model input files and output files will reside here.
workspace = os.path.join('data', 'mf6lake')
if not os.path.exists(workspace):
os.makedirs(workspace)
```
### Create the Flopy Model Objects
We are creating a square model with a specified head equal to `h1` along all boundaries. The head at the cell in the center in the top layer is fixed to `h2`. First, set the name of the model and the parameters of the model: the number of layers `Nlay`, the number of rows and columns `N`, lengths of the sides of the model `L`, aquifer thickness `H`, hydraulic conductivity `k`
```
name = 'mf6lake'
h1 = 100
h2 = 90
Nlay = 10
N = 101
L = 400.0
H = 50.0
k = 1.0
```
One big difference between MODFLOW 6 and previous MODFLOW versions is that MODFLOW 6 is based on the concept of a simulation. A simulation consists of the following:
* Temporal discretization (TDIS)
* One or more models (GWF is the only model supported at present)
* Zero or more exchanges (instructions for how models are coupled)
* Solutions
For this simple lake example, the simulation consists of the temporal discretization (TDIS) package (TDIS), a groundwater flow (GWF) model, and an iterative model solution (IMS), which controls how the GWF model is solved.
```
# Create the Flopy simulation object
sim = flopy.mf6.MFSimulation(sim_name=name, exe_name='mf6',
version='mf6', sim_ws=workspace)
# Create the Flopy temporal discretization object
tdis = flopy.mf6.modflow.mftdis.ModflowTdis(sim, pname='tdis', time_units='DAYS', nper=1,
perioddata=[(1.0, 1, 1.0)])
# Create the Flopy groundwater flow (gwf) model object
model_nam_file = '{}.nam'.format(name)
gwf = flopy.mf6.ModflowGwf(sim, modelname=name,
model_nam_file=model_nam_file)
# Create the Flopy iterative model solver (ims) Package object
ims = flopy.mf6.modflow.mfims.ModflowIms(sim, pname='ims', complexity='SIMPLE')
```
Now that the overall simulation is set up, we can focus on building the groundwater flow model. The groundwater flow model will be built by adding packages to it that describe the model characteristics.
Define the discretization of the model. All layers are given equal thickness. The `bot` array is build from `H` and the `Nlay` values to indicate top and bottom of each layer, and `delrow` and `delcol` are computed from model size `L` and number of cells `N`. Once these are all computed, the Discretization file is built.
```
# Create the discretization package
bot = np.linspace(-H/Nlay, -H, Nlay)
delrow = delcol = L/(N-1)
dis = flopy.mf6.modflow.mfgwfdis.ModflowGwfdis(gwf, pname='dis', nlay=Nlay, nrow=N, ncol=N,
delr=delrow,delc=delcol,top=0.0,
botm=bot)
# Create the initial conditions package
start = h1 * np.ones((Nlay, N, N))
ic = flopy.mf6.modflow.mfgwfic.ModflowGwfic(gwf, pname='ic', strt=start)
# Create the node property flow package
npf = flopy.mf6.modflow.mfgwfnpf.ModflowGwfnpf(gwf, pname='npf', icelltype=1, k=k,
save_flows=True)
# Create the constant head package.
# List information is created a bit differently for
# MODFLOW 6 than for other MODFLOW versions. The
# cellid (layer, row, column, for a regular grid)
# must be entered as a tuple as the first entry.
# Remember that these must be zero-based indices!
chd_rec = []
chd_rec.append(((0, int(N / 4), int(N / 4)), h2))
for layer in range(0, Nlay):
for row_col in range(0, N):
chd_rec.append(((layer, row_col, 0), h1))
chd_rec.append(((layer, row_col, N - 1), h1))
if row_col != 0 and row_col != N - 1:
chd_rec.append(((layer, 0, row_col), h1))
chd_rec.append(((layer, N - 1, row_col), h1))
chd = flopy.mf6.modflow.mfgwfchd.ModflowGwfchd(gwf, pname='chd', maxbound=len(chd_rec),
stress_period_data=chd_rec, save_flows=True)
# The chd package stored the constant heads in a structured
# array, also called a recarray. We can get a pointer to the
# recarray for the first stress period (iper = 0) as follows.
iper = 0
ra = chd.stress_period_data.get_data(key=iper)
ra
# We can make a quick plot to show where our constant
# heads are located by creating an integer array
# that starts with ones everywhere, but is assigned
# a -1 where chds are located
ibd = np.ones((Nlay, N, N), dtype=int)
for k, i, j in ra['cellid']:
ibd[k, i, j] = -1
ilay = 0
plt.imshow(ibd[ilay, :, :], interpolation='none')
plt.title('Layer {}: Constant Head Cells'.format(ilay + 1))
# Create the output control package
headfile = '{}.hds'.format(name)
head_filerecord = [headfile]
budgetfile = '{}.cbb'.format(name)
budget_filerecord = [budgetfile]
saverecord = [('HEAD', 'ALL'),
('BUDGET', 'ALL')]
printrecord = [('HEAD', 'LAST')]
oc = flopy.mf6.modflow.mfgwfoc.ModflowGwfoc(gwf, pname='oc', saverecord=saverecord,
head_filerecord=head_filerecord,
budget_filerecord=budget_filerecord,
printrecord=printrecord)
# Note that help can always be found for a package
# using either forms of the following syntax
help(oc)
#help(flopy.mf6.modflow.mfgwfoc.ModflowGwfoc)
```
### Create the MODFLOW 6 Input Files and Run the Model
Once all the flopy objects are created, it is very easy to create all of the input files and run the model.
```
# Write the datasets
sim.write_simulation()
# Print a list of the files that were created
# in workspace
print(os.listdir(workspace))
```
### Run the Simulation
We can also run the simulation from the notebook, but only if the MODFLOW 6 executable is available. The executable can be made available by putting the executable in a folder that is listed in the system path variable. Another option is to just put a copy of the executable in the simulation folder, though this should generally be avoided. A final option is to provide a full path to the executable when the simulation is constructed. This would be done by specifying exe_name with the full path.
```
# Run the simulation
success, buff = sim.run_simulation()
print('\nSuccess is: ', success)
```
### Post-Process Head Results
Post-processing MODFLOW 6 results is still a work in progress. There aren't any Flopy plotting functions built in yet, like they are for other MODFLOW versions. So we need to plot the results using general Flopy capabilities. We can also use some of the Flopy ModelMap capabilities for MODFLOW 6, but in order to do so, we need to manually create a SpatialReference object, that is needed for the plotting. Examples of both approaches are shown below.
First, a link to the heads file is created with `HeadFile`. The link can then be accessed with the `get_data` function, by specifying, in this case, the step number and period number for which we want to retrieve data. A three-dimensional array is returned of size `nlay, nrow, ncol`. Matplotlib contouring functions are used to make contours of the layers or a cross-section.
```
# Read the binary head file and plot the results
# We can use the existing Flopy HeadFile class because
# the format of the headfile for MODFLOW 6 is the same
# as for previous MODFLOW verions
fname = os.path.join(workspace, headfile)
hds = flopy.utils.binaryfile.HeadFile(fname)
h = hds.get_data(kstpkper=(0, 0))
x = y = np.linspace(0, L, N)
y = y[::-1]
c = plt.contour(x, y, h[0], np.arange(90,100.1,0.2))
plt.clabel(c, fmt='%2.1f')
plt.axis('scaled');
x = y = np.linspace(0, L, N)
y = y[::-1]
c = plt.contour(x, y, h[-1], np.arange(90,100.1,0.2))
plt.clabel(c, fmt='%1.1f')
plt.axis('scaled');
z = np.linspace(-H/Nlay/2, -H+H/Nlay/2, Nlay)
c = plt.contour(x, z, h[:,50,:], np.arange(90,100.1,.2))
plt.axis('scaled');
# We can also use the Flopy PlotMapView capabilities for MODFLOW 6
fig = plt.figure(figsize=(10, 10))
ax = fig.add_subplot(1, 1, 1, aspect='equal')
modelmap = flopy.plot.PlotMapView(model=gwf, ax=ax)
# Then we can use the plot_grid() method to draw the grid
# The return value for this function is a matplotlib LineCollection object,
# which could be manipulated (or used) later if necessary.
quadmesh = modelmap.plot_ibound(ibound=ibd)
linecollection = modelmap.plot_grid()
contours = modelmap.contour_array(h[0], levels=np.arange(90,100.1,0.2))
# We can also use the Flopy PlotMapView capabilities for MODFLOW 6
fig = plt.figure(figsize=(10, 10))
ax = fig.add_subplot(1, 1, 1, aspect='equal')
# Next we create an instance of the ModelMap class
modelmap = flopy.plot.PlotMapView(model=gwf, ax=ax)
# Then we can use the plot_grid() method to draw the grid
# The return value for this function is a matplotlib LineCollection object,
# which could be manipulated (or used) later if necessary.
quadmesh = modelmap.plot_ibound(ibound=ibd)
linecollection = modelmap.plot_grid()
pa = modelmap.plot_array(h[0])
cb = plt.colorbar(pa, shrink=0.5)
```
### Post-Process Flows
MODFLOW 6 writes a binary grid file, which contains information about the model grid. MODFLOW 6 also writes a binary budget file, which contains flow information. Both of these files can be read using Flopy capabilities. The MfGrdFile class in Flopy can be used to read the binary grid file. The CellBudgetFile class in Flopy can be used to read the binary budget file written by MODFLOW 6.
```
# read the binary grid file
fname = os.path.join(workspace, '{}.dis.grb'.format(name))
bgf = flopy.mf6.utils.MfGrdFile(fname)
# data read from the binary grid file is stored in a dictionary
bgf._datadict
# read the cell budget file
fname = os.path.join(workspace, '{}.cbb'.format(name))
cbb = flopy.utils.CellBudgetFile(fname, precision='double')
cbb.list_records()
flowja = cbb.get_data(text='FLOW-JA-FACE')[0][0, 0, :]
chdflow = cbb.get_data(text='CHD')[0]
# By having the ia and ja arrays and the flow-ja-face we can look at
# the flows for any cell and process them in the follow manner.
k = 5; i = 50; j = 50
celln = k * N * N + i * N + j
ia, ja = bgf.ia, bgf.ja
print('Printing flows for cell {}'.format(celln))
for ipos in range(ia[celln] + 1, ia[celln + 1]):
cellm = ja[ipos]
print('Cell {} flow with cell {} is {}'.format(celln, cellm, flowja[ipos]))
```
| github_jupyter |
# TRaPPE FF Parameter Conversion
# TraPPE Force Field Parameters & Functional Form
http://chem-siepmann.oit.umn.edu/siepmann/trappe/index.html
Overall, the potential energy function takes the following form:
\begin{equation*}
\ U_{total} = \sum_{angles}{\frac{k_{\theta}}2 (\theta - \theta_{eq} )^{2}} + \sum_{dihedrals}{c_{0} + c_{1}[1 + cos(\phi)] + c_{2}[1 - cos(2\phi)] + c_{3}[1 + cos(3\phi)]} + \sum_{i=1}^{N-1}{\sum_{j=i+1}^{N}{ 4\epsilon_{ij}[(\frac{\sigma_{ij}}r_{ij})^{12} - (\frac{\sigma_{ij}}r_{ij})^6] }}
\end{equation*}
## Nonbonded Potential
\begin{equation*}
\ u_{NB} = 4\epsilon_{ij}[(\frac{\sigma_{ij}}r_{ij})^{12} - (\frac{\sigma_{ij}}r_{ij})^6]
\end{equation*}
### Combination rules
\begin{equation*}
\sigma_{ij} = \frac{1}2 (\sigma_{ii} + \sigma_{jj})
\end{equation*}
\begin{equation*}
\epsilon_{ij} = (\epsilon_{ii}\epsilon_{jj})^{1/2}
\end{equation*}
### Nonbonded Parameters
Here, CHx indicates linear alkane carbons while Ccx indicates carbons in cyclic rings of size x.
(psuedo)atom | type | $ \frac{\epsilon}k_B $ [K] | $ \sigma $ [Angstrom] | q [e]
---------------| :-----------: | :--------------------------: | :--------------------:|:--------:
CH4 | CH4 | 148.0 | 3.730 | 0.000
CH3 | [CH3]-CHx | 98.0 | 3.750 | 0.000
CH2 | CHx-[CH2]-CHx | 46.0 | 3.950 | 0.000
Cc5 | CH2-[CH2]-CH2 | 56.3 | 3.880 | 0.000
Cc6 | CH2-[CH2]-CH2 | 52.5 | 3.910 | 0.000
## Bonded Potentials
### Bond Stretching Parameters
TraPPE uses fixed bond lengths. We will be adding flexible bonds in our implementation, which are based on the parameters for sp3 carbons in the general amber force field (gaff)
Type | Length [Angstrom] | $k_{b}$ (kcal/mol)
------ | :---------------: | :---
CHx-CHy | 1.540 | 300.9
### Angles Bending Parameters
\begin{equation*}
u_{bend} = \frac{k_{\theta}}2 (\theta - \theta_{eq} )^{2}
\end{equation*}
Type | $ \theta $ | $ \frac{k_{\theta}}{k_{B}} [K/rad^2] $
------------- | :-------------: | :------------------------:
CHx-(CH2)-CHx | 114.0 | 62500
Cc5-(Cc5)-Cc5 | 105.5 | 62500
Cc6-(Cc6)-Cc6 | 114.0 | 62500
### Dihedral Potential
\begin{equation*}
\ u_{torsion}(\phi) = c_{0} + c_{1}[1 + cos(\phi)] + c_{2}[1 - cos(2\phi)] + c_{3}[1 + cos(3\phi)]
\end{equation*}
\begin{equation*}
\ u_{torsion_{cyclic}}(\phi) = c_{0} + c_{1}[cos(\phi)] + c_{2}[cos(2\phi)] + c_{3}[cos(3\phi)]
\end{equation*}
Because of the different form of the cyclic alkanes, we must transformt the equation...
\begin{equation*}
\ u_{torsion_{cyclic}}(\phi) = c_{0} + c_{1}[cos(\phi)] + c_{2}[cos(2\phi)] + c_{3}[cos(3\phi)] =
(c_{0} - c_{1} - c_{2} - c_{3}) + c_{1}[1 + cos(\phi)] + c_{2}[1 - cos(2\phi)] + c_{3}[1 + cos(3\phi)]
\end{equation*}
*The table below gives the parameters for the first form above.
cc5 $c_{0}$ originial 31394
cc6 $c_{0}$ original 5073
| Type | $ \frac{c_{0}}{k_{B}} [K] $ | $ \frac{c_{1}}{k_{B}} [K] $ | $ \frac{c_{2}}{k_{B}} [K] $ | $ \frac{c_{3}}{k_{B}} [K] $ |
| -----------| :--------------: | :-----------------------: |:---------------:|:--------:|
| CHx-(CH2)-(CH2)-CHy | 0 | 355.03 | -68.19 | 791.32 |
| Cc5-(Cc5)-(Cc5)-Cc5 | -32534 | 45914 | 16518 | 1496
| Cc6-(Cc6)-(Cc6)-Cc6 | -5339 | 6840 | 3509 | 63
## FF Units
Parameter Symbol | Name | Units | Special notes
----------------- | :---------------------: | :---------------------: | :-----------
$ Length $ | Bond Length | Angstrom |
$ k_{\theta} $ | Harmonic Angle Constant | kcal/mol/radian$^2$ | Appears as $k_{\theta}/2$ in functional
$ \theta_{eq} $ | Equilibrium Angle | degrees |
$ c_{n} $ | Torsion Barrier | K | Given as $c_{n}/k_{B}$
$ {\epsilon} $ | well depth | K | Given as ${\epsilon}/k_{B}$ above
$ {\sigma} $ | | K | Given as ${\sigma}/k_{B}$ above
# Amber FF Conversion
\begin{equation*}
\ U_{total} = \sum_{bonds}{k_{b}(r-r_{0})^2} + \sum_{angles}{k_{\theta}{(\theta - \theta_o}^2)} + \sum_{dihedrals}{(V_{n}[1 + cos(n\phi - \gamma)]}) + \sum_{i=1}^{N-1}{\sum_{j=i+1}^{N}{ \frac{A_{ij}}{R_{ij}^{12}} - \frac{B_{ij}}{R_{ij}^{6}} }}
\end{equation*}
* Note - electrostatics omitted here for TRaPPE alkanes
A good resource : http://alma.karlov.mff.cuni.cz/bio/99_Studenti/00_Dalsi/ParamFit/2013_ParamFit_AmberTools13.pdf
## Nonbonded Parameter Considerations
The functional form of the Amber force field considers $ A_{ij} $ and $B_{ij} $ as the functional form, these are put into tleap using an frcmod with parameters $\sigma$ and $R_{min}$. A and B parameters are then calculated using the following combination rules:
\begin{equation*}
\ A = {\epsilon}{R_{min}}^{12}
\ B = 2{\epsilon}{R_{min}}^{6}
\end{equation*}
where
\begin{equation*}
\ R_{min} = 2^{1/6}{\sigma}
\end{equation*}
and mixing rules are the same as outlined above for the TraPPE forcefield (Lorentz/Berthelot mixing rules)
## Dihedral Considerations
Amber is able to handle dihedrals with more than one torsional term, as is needed for TraPPE.
For Trappe, there are multiple $c_{n}$'s corresponding to $V_{n}$'s in Amber. All TraPPE terms for straight alkanes have a phase $({\gamma}$) of zero.
For the case of $c_{0}$, n = 0, $cos(1)=0$, so insert phase shift of 90 degrees
\begin{equation*}
\ U_{dihedral} = V_{n}[1 + cos(n{\phi}-{\gamma})]
\end{equation*}
### Converting from TraPPE Dihedral form to Amber dihedral form
\begin{equation*}
u_{dihedralTrappe}(\phi) = c_{0} + c_{1}[1 + cos(\phi)] + c_{2}[1 - cos(2\phi)] + c_{3}[1 + cos(3\phi)]
\end{equation*}
The minus sign in the second term can be accounted for by incorporating a phase shift, ${\gamma}$ of 180 degrees into a multi- term Amber equation, I.E. -
\begin{equation*}
c_{0} + c_{1}[1 + cos(\phi)] + c_{2}[1 - cos(2\phi)] + c_{3}[1 + cos(3\phi)] = V_{0}[1 + cos(0)] + V_{1}[1+cos({\phi})] + V_{2}[1+cos(2{\phi} - 90)] + V_{3}[1+cos(3{\phi})]
\end{equation*}
## FF Units
Parameter Symbol | Name | Units | Special notes
----------------- | :---------------------: | :---------------------: | :-----------
$ k_{b} $ | Harmonic Bond Constant | kcal/mol/Angstrom$^2 $ |
$ r_{0} $ | Equilibrium Bond Length | Angstrom |
$ k_{\theta} $ | Harmonic Angle Constant | kcal/mol/radian$^2$ |
$ \theta_{0} $ | Equilibrium Angle | degrees |
$ V_{n} $ | Torsion Barrier | kcal/mol |
$ \gamma $ | Torision Phase | degrees |
$ n $ | Torsion Periodicity | |
$ A_{ij} $ | Nonbonded A parameter | | Used in prmtop file (generated by tleap)
$ B_{ij} $ | Nonbonded B parameter | | Used in prmtop file (generated by tleap)
$ R_{min}$ | vdW Radius | Angstrom | Input by user (frcmod file)
$ {\epsilon} $ | well depth | kcal/mol | Input by user (frcmd file)
```
import scipy.constants as const
import pandas as pd
import os
import glob
import re
import math
# Functions to convert units
def K_to_kcal_mol(joule_value):
'''
Convert value which is in units of K/kB to units of kcal/mol
'''
return const.Avogadro*(joule_value*const.k/(const.calorie*1000))
def convert_nb(trappe_epsilon, trappe_sigma):
'''
Convert nb parameters (epsilon and sigma) from Trappe FF to Amber FF.
Input -
Epsilon units - K
Sigma units - Angstrom
Output -
Rmin units Angstrom
epsilon units kcal/mol
'''
r_min = 2.**(1./6.)*trappe_sigma/2
epsilon = K_to_kcal_mol(trappe_epsilon)
return r_min, epsilon
def calculate_bead_masses(bead_name):
masses = {
"O" : 15.999,
"CH2" : 14.02658,
"CH3" : 15.0452,
"CH" : 13.01864,
"CH4" : 16.0425,
"H" : 1.00794,
}
try:
return [masses[x] for x in bead_name]
except KeyError:
print("Entry %s not found in mass dictionary" %(bead_name))
cyclic = ["cyclopentane" , "cyclohexane", "1-3-5-trioxane"]
def process_name_string(types, length):
"""
Processes names for atoms in bond and angle interactions to fit AMBER specifications
"""
return_names = []
name_list = [re.sub("\(|\)", "", x ) for x in types.values]
name_split = [[re.split("-| ", x) for x in name_list][n] for n in range(0,len(name_list))]
for n in range(0, len(name_split)):
name_split[n] = ["X " if "x" in x else x for x in name_split[n][0:length]]
name_split[n] = ["X " if "y" in x else x for x in name_split[n]]
name_split[n] = [s[0:3:2] if len(s) > 2 else s for s in name_split[n]]
return_names.append('-'.join(name_split[n]))
return return_names
def read_trappe_csv(f, name):
# Read data to pandas dataframe
data = pd.read_csv(f, names=range(0,20), skiprows=1)
data.dropna(inplace=True, axis=1, how='all')
# Find section starts
atom_section_start = data[data[1]=="(pseudo)atom"].index.tolist()
bond_section_start = data[data[1]=="stretch"].index.tolist()
angle_section_start = data[data[1]=="bend"].index.tolist()
torsion_section_start = data[data[1]=="torsion"].index.tolist()
# Start dictionary of relevant sections
sections = {}
## Create dataframes from sections
# Read in atom information
atoms = data.iloc[atom_section_start[0]+1:bond_section_start[0]].copy()
atoms.columns = data.iloc[atom_section_start[0]]
atoms.dropna(inplace=True, axis=1)
sections['atoms'] = atoms
# Get bond data from file. All bonds are set to the same k value for simplicity
test_val = float(data.iloc[bond_section_start[0]+1][0])
if not math.isnan(test_val):
bonds = data.iloc[bond_section_start[0]+1:angle_section_start[0]].copy()
bonds.columns = data.iloc[bond_section_start[0]]
bonds['k_b'] = 300.9
bonds.dropna(inplace=True, axis=1)
sections['bonds'] = bonds
# Get bond information
test_val = float(data.iloc[angle_section_start[0]+1][0])
if not math.isnan(test_val):
if len(torsion_section_start) > 0:
angles = data.iloc[angle_section_start[0]+1:torsion_section_start[0]].copy()
else:
angles = data.iloc[angle_section_start[0]+1:].copy()
angles.dropna(inplace=True, axis=1)
angles.columns = data.iloc[angle_section_start[0]][:5].values
sections['angles'] = angles
# Handle torsions
if len(torsion_section_start) > 0:
torsions = data.iloc[torsion_section_start[0]+1:].copy()
torsions.dropna(inplace=True, axis=1)
torsions.columns = data.iloc[torsion_section_start[0]].values
torsion_names = process_name_string(torsions['type'], 4)
torsions['type'] = torsion_names
if name in cyclic:
hold = torsions['c0/kB [K]'].astype(float) - torsions['c1/kB [K]'].astype(float) - \
torsions['c2/kB [K]'].astype(float) - torsions['c3/kB [K]'].astype(float)
torsions['c0/kB [K]'] = hold
print(name)
sections['torsions'] = torsions
return sections
def calc_amber_parameters(section_data, name):
## Process stored data frames
# Make write list dictionary
write_list = {}
for k,v in section_data.items():
if k == 'atoms':
# Calculate atom masses
v['masses'] = calculate_bead_masses(v['(pseudo)atom'].values)
# Manipulate atom names for atoms frame
v['(pseudo)atom'] = (v['(pseudo)atom'].str[0:3:2])
# Calculate epsilon in kcal/mol and rmin
v['r_min'], v['epsilon [kcal/mol]'] = convert_nb(v['epsilon/kB [K]'].astype(float),
v['sigma [Ang.]'].astype(float))
if k == 'angles':
# Process angles
v["k_theta [kcal/mol]"] = K_to_kcal_mol(v["k_theta/kB [K/rad^2]"].astype(float)/2.)
v['type'] = v['bend']
# Process angle type labels
v.drop(["bend"], axis=1, inplace=True)
if k == 'bonds':
v['type'] = v['stretch']
if k == 'torsions':
# Process torsions
v['ee_scaling'] = "SCEE=0.0"
v['nb_scaling'] = "SCNB=0.0"
v['type'] = v['torsion']
if name in cyclic:
v['phase_c0'] = 90
for x in range(3,-1,-1):
v['c%s/kB [kcal/mol]' %(x)] = K_to_kcal_mol(v['c%s/kB [K]' %(x)].astype(float))
v['%s' %(x)] = -x
if name not in cyclic:
if x == 2:
v['phase_c%s' %(x)] = -180
else:
v['phase_c%s' %(x)] = 0
elif x is not 0:
v['phase_c%s' %(x)] = 0
v['divider'] = 1
# Change type to atom names
types = v['type']
types = [re.sub("'|'| ","", x) for x in types]
proc_type = [re.split("-", x) for x in types]
if proc_type[0][0].isdigit():
name_list = []
for x in proc_type:
name_string = ""
atom_list = [section_data['atoms']['(pseudo)atom'][(section_data['atoms']["#"] == y)].values[0] for y in x]
name_string = "-".join(atom_list)
name_list.append(name_string)
v['type'] = name_list
# Remove unnecessary colums (unconverted units)
drop_list = [col for col in v.columns if '[K' in col]
v.drop(drop_list, inplace=True, axis=1)
if k is not 'atoms' and k is not 'torsions':
v.drop_duplicates(subset=["type"], inplace=True, keep='first')
elif k is 'torsions':
v.drop(['torsion',"#"], inplace=True, axis=1)
v.drop_duplicates(inplace=True, keep='first')
section_data['atoms'].drop_duplicates(subset=['type'], inplace=True, keep='first')
return section_data
def write_frcmod(name, amber):
# Get filename for frcmod & open
frcmod = open("frcmod_trappe.%s" %(name), "w")
frcmod.write("# Implementation of TraPPE FF for %s in Amber\n\n" %(name))
for k,v in amber.items():
if k == 'atoms':
# Write atom and nonbond data to frcmod
frcmod.write('MASS\n')
v.to_csv(frcmod, header=False, index=False, columns=['(pseudo)atom', 'masses'], sep="\t")
frcmod.write('\nNONBOND\n')
v.to_csv(frcmod, header=False, index=False, columns=["(pseudo)atom", "r_min",
"epsilon [kcal/mol]"], sep="\t")
if k == 'bonds':
# Write bond data to frcmod
frcmod.write('\nBONDS\n')
v.to_csv(frcmod, header=False,index=False, columns=["type", "k_b", "length [Ang.]"], sep="\t")
if k == 'angles':
frcmod.write('\nANGLE\n')
v.to_csv(frcmod, header=False,index=False, columns=["type", "k_theta [kcal/mol]",
"theta [degrees]"], sep="\t")
if k == 'torsions':
frcmod.write('\nDIHED\n')
for x in range(3,-1,-1):
v.to_csv(frcmod, header=False,index=False, columns=["type", "divider", 'c%s/kB [kcal/mol]' %(x), \
"phase_c%s" %(x),"%s" %(x), "ee_scaling", "nb_scaling"], sep="\t")
frcmod.close()
return 0
def write_leap_in(name, amber):
file_name = name + "_leap.in"
f = open(file_name, "w")
f.write("loadamberparams frcmod_trappe.%s\n" %(name))
f.write("sys = loadmol2 %s.mol2\n" %(name))
f.write('setbox sys "centers" 50\n')
f.write("saveamberparm sys trappe_%s_single_molecule.prmtop trappe_%s_single_molecule.inpcrd\n" %(name, name))
f.write("quit\n")
f.close()
files = glob.glob("trappe_*.csv")
run_file = "make_systems.sh"
rf = open(run_file, "w")
for f in files:
# Get molecule name
name = re.split("_|\.",f)[1]
print(name)
# Add to sh file
rf.write("tleap -f %s_leap.in\n" %(name))
# Process 'csv' from trappe
output = read_trappe_csv(f, name)
# Transform to amber format and units
amber = calc_amber_parameters(output, name)
# Print frcmod
write_frcmod(name, amber)
# Print accompanying leap input file
write_leap_in(name, amber)
rf.close()
```
| github_jupyter |
# Sequential Domain Reduction
## Background
Sequential domain reduction is a process where the bounds of the optimization problem are mutated (typically contracted) to reduce the time required to converge to an optimal value. The advantage of this method is typically seen when a cost function is particularly expensive to calculate, or if the optimization routine oscilates heavily.
## Basics
The basic steps are a *pan* and a *zoom*. These two steps are applied at one time, therefore updating the problem search space evey iteration.
**Pan**: recentering the region of interest around the most optimal point found.
**Zoom**: contract the region of interest.

## Parameters
There are three parameters for the built-in `SequentialDomainReductionTransformer` object:
$\gamma_{osc}:$ shrinkage parameter for oscillation. Typically [0.5-0.7]. Default = 0.7
$\gamma_{pan}:$ panning parameter. Typically 1.0. Default = 1.0
$\eta:$ zoom parameter. Default = 0.9
More information can be found in this reference document:
---
Title: "On the robustness of a simple domain reduction scheme for simulation‐based optimization"
Date: 2002
Author: Stander, N. and Craig, K.
---
---
Let's start by importing the packages we'll be needing
```
import numpy as np
from bayes_opt import BayesianOptimization
from bayes_opt import SequentialDomainReductionTransformer
import matplotlib.pyplot as plt
```
Now let's create an example cost function. This is the [Ackley function](https://en.wikipedia.org/wiki/Ackley_function), which is quite non-linear.
```
def ackley(**kwargs):
x = np.fromiter(kwargs.values(), dtype=float)
arg1 = -0.2 * np.sqrt(0.5 * (x[0] ** 2 + x[1] ** 2))
arg2 = 0.5 * (np.cos(2. * np.pi * x[0]) + np.cos(2. * np.pi * x[1]))
return -1.0 * (-20. * np.exp(arg1) - np.exp(arg2) + 20. + np.e)
```
We will use the standard bounds for this problem.
```
pbounds = {'x': (-5, 5), 'y': (-5, 5)}
```
This is where we define our `bound_transformer` , the Sequential Domain Reduction Transformer
```
bounds_transformer = SequentialDomainReductionTransformer()
```
Now we can set up two idential optimization problems, except one has the `bound_transformer` variable set.
```
mutating_optimizer = BayesianOptimization(
f=ackley,
pbounds=pbounds,
verbose=0,
random_state=1,
bounds_transformer=bounds_transformer
)
mutating_optimizer.maximize(
init_points=2,
n_iter=50,
)
standard_optimizer = BayesianOptimization(
f=ackley,
pbounds=pbounds,
verbose=0,
random_state=1,
)
standard_optimizer.maximize(
init_points=2,
n_iter=50,
)
```
After both have completed we can plot to see how the objectives performed. It's quite obvious to see that the Sequential Domain Reduction technique contracted onto the optimal point relativly quickly.
```
plt.plot(mutating_optimizer.space.target, label='Mutated Optimizer')
plt.plot(standard_optimizer.space.target, label='Standard Optimizer')
plt.legend()
```
Now let's plot the actual contraction of one of the variables (`x`)
```
# example x-bound shrinking
x_min_bound = [b[0][0] for b in bounds_transformer.bounds]
x_max_bound = [b[0][1] for b in bounds_transformer.bounds]
x = [x[0] for x in mutating_optimizer.space.params]
plt.plot(x_min_bound[1:], label='x lower bound')
plt.plot(x_max_bound[1:], label='x upper bound')
plt.plot(x[1:], label='x')
plt.legend()
```
| github_jupyter |
# Created on 12/2020
@author : V. Robin
GENERATION OF INITIAL CONDITION TO OBTAIN
A 3D intial condition from a isoline equation
Preliminary 0D computations with Cantera have been
performed by Said Taileib to obtain the ignition delay time
as function of temperature : tau=f(Tu).
The coefficient A and B of the linearized profile
Log(tau)=f(1/T) must be known before running this program.
This program do:
- from the desired propagation speed of the
ignition front, the profile Tu(x) is determined
- for all x-point, i.e. all fresh temperature,
a 0D computation is performed to obtain Tu(x,t)
- The retained profile for IC of resident corresponds
to a certain time t=ti0D
- This profile is filled on the resident grid (density and Y1 values) by setting a specific isovalue on a line in the domain. This line can be seen as a level set function.
```
#Import of the libraries and functions used in this script
import math
import shapely.geometry as geom
from shapely.geometry import LineString, Point, Polygon
from shapely.ops import linemerge, unary_union, polygonize
import matplotlib.pyplot as plt
from matplotlib import rc
import cantera as ct
import numpy as np
import os
# functions definition
def cut_polygon_by_line(polygon, line):
merged = linemerge([polygon.boundary, line])
borders = unary_union(merged)
polygons = polygonize(borders)
return list(polygons)
def column(matrix, i): #creation of a function to read a row in a matrix and write it to a vector
return [row[i] for row in matrix]
#####################################
# Resident Domain and grid definition
#####################################
lx=0.3 # length x direction
#ly= 0.08 # length y direction
ly= 0.01 # length y direction
lz= ly # length y direction
#
nx= 1e-4 # cell syze x direction
ny=nx # cell syze y direction
nz=ny # cell syze y direction
#
eps=0.000000001
nl_x=int(lx/nx+eps) #number of cells in x
nq_x=nl_x+1 #number of points in x
nl_y=int(ly/ny+eps) #number of cells in y
nq_y=int(nl_y)+1 #number of points in y
nl_z=int(lz/nz+eps) #number of cells in z
nq_z=int(nl_z)+1 #number of points in z
#
print("The number of points is" , nq_x,'x',nq_y,'x',nq_z,'=',nq_x*nq_y*nq_z)
print("The number of cells is" , nl_x,'x',nl_y,'x',nl_z,'=',nl_x*nl_y*nl_z)
################################
# Initial condition parameters
###############################
U = 40 # propagation speed of the ignition front desired
c= 1e-5 # ignition delay time of the most reactive mixture (of the initial state)
#
#########################################
# Chemistry parameters and gas properties
# From preliminary 0D cantera calculation
# Log(Tau)=f(1/T)
#########################################
#B=21547.968 # Alka's value
#A=5.79243E-12 # Alka's value
B=8028.414 # Said's value
A=3.86582E-9 # Said's value
Pressure=16.46*1.01325e5 # initial pressure
comp = 'H2:2.0,O2:1.0,N2:3.76' # fresh gas composition
gas = ct.Solution('H2_Global-Said.cti') # Said global step chemistry
timestepadim=1/1000 # timestep adimentionned by ignition delay
n=int(10/timestepadim) # number of time iteration
defaulttimestep=c*timestepadim # cantera timestep based on the ignition delay of the most reactive mixture
nxc=nx # cantera x step : the best is equal to nx
print('u =', U)
##################################################
# Cantera computation to define the fresh gas profile
##################################################
# Maximum length : it corresponds to the cantera computation length
# Not easy to define, it depends of the shape and position of the flame
# : too lagre and calculation is very long
# : too short and the profile is cut on the resident grid
#
coeflength=1.2
length=math.sqrt(lx ** 2 + ly ** 2 + lz ** 2)*coeflength # choice max length of the domain mutiply by a coefficient
# number of cells and points in the cantera domain
nl=int(length/nxc+eps)
nq=int(nl)+1
length=nq*nxc
timestep=np.zeros(nq) # variable timestep
Tu=np.zeros(nq) # initial temperature - calculated from cantera 0D curve fit
tau=np.zeros(nq) # ignition delay time array (calculated from cantera 0D curve fit)
x1=np.zeros(nq) # coordinates
# Loop to define the initial profil of fresh gas temperature leading to the right propagation speed.
# This is based on the Zeldovitch gradient (ignition delay gradient).
for i in range(nq):
xx=i*nxc
x1[i]=(i)*nxc # x (m)
Tu[i]=B/(math.log((((xx/U)+c)/A))) # Fresh gas temperature
tau[i]=(xx/U)+c # ignition delay time
# to use variable time step : calculation stop at n*tau[i]*timestepadim
timestep[i]=((xx/U)+c)*timestepadim
# to use constant time step : calculation stop at ti0D
#timestep[i]=ti0D/n
print("Cantera length, corresponding points and cells" ,length, nq, nl)
print("Temperature range for cantera" ,Tu[0],Tu[nq-1])
print("Position range" ,x1[0],x1[nq-1])
print("Range of ignition delay time" ,tau[0], tau[nq-1])
################################################
# Cantera computations for all fresh temperatures
################################################
# Cantera loop for reactor solving with time (runned until it reaches the n*timestepadim i.e. 10 x ignition time)
# Declaration of the results matrix using the divison of the domain set and the maximum number of iterations to reach the max temperature
resu = np.zeros((nq,n)) # temperature results
resu1 = np.zeros((nq,n)) # Y results (gas composition) absolute value
resu2 = np.zeros((nq,n)) # Normalized Y
resu3 = np.zeros((nq,n)) # Normalized Y
times = np.zeros((nq,n))
for i in range(nq): # loop on each position of the 1D profile
time = 0.0
Temperature=Tu[i] # fresh gas temperature
Tempdata=[]
Ydata=[]
rhodata=[]
gas.TPX = Temperature, Pressure, comp # gas definition
r = ct.IdealGasConstPressureReactor(gas) # reactor definition
sim = ct.ReactorNet([r]) # network reactor definition
Tempdata.append(Temperature) # Recording of first temperature value
Ydata.append(r.Y)
rhodata.append(r.density)
for j in range(n-1): # Cantera time loop
time+=timestep[i]
sim.advance(time)
Tempdata.append(r.T)
Ydata.append(r.Y)
rhodata.append(r.density)
resu[i,:]=Tempdata # Temperature stored
resu1[i,:]=column(Ydata, 0) # First species stored
resu3[i,:]=rhodata # rho stored
for j in range(n):
times[i,j]= j*timestep[i] # times stored
resu2[i,j]=(resu1[i,j]-resu1[i,n-1])/(resu1[i,0]-resu1[i,n-1]) # Normalized value of the first species stored
print('End Cantera loop')
################################################
# Plot of cantera results : Two time profile
# corresponding to the extreme position in the 1D profile
# i.e. the most and less reactive mixture
################################################
plt.style.use('classic')
rc('font',**{'family':'DejaVu Sans','serif':['Computer Modern Roman']})
rc('text', usetex=True)
rc('xtick', labelsize=12)
rc('ytick', labelsize=12)
rc('axes', labelsize=12)
print('fresh Temperature most reactive : ', resu[0,0])
print('fresh Temperature less reactive : ', resu[nq-1,0])
plt.plot(times[0,:],resu[0,:])
plt.plot(times[nq-1,:],resu[nq-1,:])
plt.xlim(0, 0.00002)
#plt.legend()
plt.show()
################################################
# Selection of the gas state corresponding to time ti0D
# for the whole points of the 1D profile
################################################
kit=np.zeros((nq),int) # position of ti0D in arrays (necessary for variable time step)
ti0D=100*c # cantera time used to obtain partially burnt gas in the cantera domain.
# It must be higher than c (ignition time of the most reactive) to have burnt gas
T = np.zeros(nq)
T0 = np.zeros(nq)
Tb = np.zeros(nq)
Y1 = np.zeros(nq)
rho = np.zeros(nq)
rho0 = np.zeros(nq)
rhob = np.zeros(nq)
for i in range(nq):
kit[i]=int(ti0D/timestep[i]) # position of the time ti0D in arrays
T0[i]=resu[i, 0]
rho0[i]=resu3[i, 0]
Tb[i]=resu[i, n-1]
rhob[i]=resu3[i, n-1]
if (kit[i]<n):
T[i]=resu[i, kit[i]]
Y1[i]=resu2[i,kit[i]]
rho[i]=resu3[i,kit[i]]
else:
T[i]=resu[i,n-1]
Y1[i]=resu2[i,n-1]
rho[i]=resu3[i,n-1]
print('Cantera time used as initial condition: ', ti0D)
################################################
# PLot 1D profile corresponding to fully fresg gas,
# fully burnt gas and gas state att ti0D
################################################
plt.style.use('classic')
rc('font',**{'family':'DejaVu Sans','serif':['Computer Modern Roman']})
rc('text', usetex=True)
rc('xtick', labelsize=12)
rc('ytick', labelsize=12)
rc('axes', labelsize=12)
trace=1
if (trace==1):
plt.plot(x1,T,label=' partially burnt gas' )
plt.plot(x1,T0,label='pure fresh gas' )
plt.plot(x1,Tb,label='pure burnt gas' )
plt.xlabel('x (m)',fontsize=15)
plt.ylabel('T(K)',fontsize=15)
plt.legend()
plt.show()
#
plt.plot(x1,Y1,label='fresh gas')
plt.xlabel('x (m)')
plt.ylabel('Y1')
plt.legend()
plt.show()
plt.plot(x1,rho,label='partially burnt gas')
plt.plot(x1,rho0,label='pure fresh gas')
plt.plot(x1,rhob,label='pure burnt gas')
plt.xlabel('x (m)')
plt.ylabel('rho')
plt.legend()
plt.show()
################################################
# Find the x position in 1D profile corresponding to
# a certain value of Y. This result is used to set the
# position of the 1D profile in the 2D resident domain
################################################
Yvalue=0.5
idx = (np.abs(Y1-Yvalue)).argmin()
flame_pos=x1[idx]
print('Total number of point in the 1D profile :', nq)
print('Point position in array :', idx ,'corresponding to the nearest value of Y=', Yvalue)
print('Y and x value at this point :', Y1[idx],x1[idx])
################################################
# DEFINTION DE LA POSITION DE LA FLAMME IN 2D RESIDENT
# CAS D'UNE DROITE D'ANGLE THETA
################################################
# On créé d'abord une isoligne sur une partie du domaine
# (des symetries seront réalisées pour compléter le domaine)
# L'isoligne représente la "flamme" qui sépare les gaz frais des gaz brulés
#
# Point de départ de l'isoligne
xini=0.0
#yini=ly/2.0
yini=0.0
theta = 85 # angle (degrees) of the front with respect to the vertical axis
points_of_isoligne = []
#
# pour avoir une structure dont la forme est maintenu, il faut mettre les points extérieurs 0) et 2).
# pour avoir une vitesse constante du front, il faut supprimer les points extérieurs 0) et 2).
shape_maintained = 1
#
if (theta>eps):
if (shape_maintained==1):
# 0) extérieur du domaine : on rajoute des points à l'extérieur du domaine si besoin : POUR X<XINI x=0
points_of_isoligne.append((0,0))
#
# 1) on boucle sur toutes les positions y correspondants aux points du maillage.
#for j in range(int(nl_y)+1):
for j in range(int(nl_y/2)+1):
y_isoline = yini+j*ny # coordonnées y
# coordonnées x de la forme désirée (ici une droite d'angle theta)
x_isoline = xini + (yini+y_isoline)*math.tan(math.radians(theta))
# création du tableau de coordonées des points
points_of_isoligne.append((x_isoline,y_isoline))
#print(j, x_isoline,y_isoline)
#
if (shape_maintained==1):
# 2) extérieur du domaine : on rajoute des points à l'extérieur du domaine si besoin : POUR X>XMAXI x=lx
# ici on prend un point loin : le max de coordonnées x pour prolonger la droite
points_of_isoligne.append((lx,-yini+(lx-xini)/math.tan(math.radians(theta))))
#points_of_isoligne.append((x_isoline,ly))
#
else:
points_of_isoligne.append((xini,0))
points_of_isoligne.append((xini,ly))
#
#
# création des objets géométriques
isoline = LineString(points_of_isoligne) # on créé une ligne avec ces points
demidomaine = Polygon([(0, 0), (0, ly), (lx, ly), (lx, 0)]) # polygone du domaine
domaine_parts = cut_polygon_by_line(demidomaine, isoline) # découpage du polygone en deux
if (theta>eps):
domaine_bunrt = domaine_parts[0] # Création d'un polygone représentant les gaz brulés
else:
domaine_bunrt = domaine_parts[1] # Création d'un polygone représentant les gaz brulés
#
#plt.plot(demidomaine)
xbunrt,ybunrt = domaine_bunrt.exterior.xy
xfull,yfull = demidomaine.exterior.xy
plt.figure(figsize=(30*lx,100*ly/2))
plt.plot(xbunrt,ybunrt)
plt.plot(xfull,yfull)
plt.show()
################################################
# REMPLISSAGE DES TABLEAUX DE CONDITION INITIALE
# on remplie seulement la moitié du domaine en y
# des symétries sont utilisées ensuite lors
# de l'écriture dans les fichiers
################################################
#CI_rho = np.zeros((nl_x,int(nl_y/2)))
CI_rho = np.zeros(shape=(nl_x,nl_y,nl_z))
CI_Y = np.zeros(shape=(nl_x,nl_y,nl_z))
gmax=nq
for k in range(nl_z):
for j in range(nl_y):
for i in range(nl_x):
# changme,t de repere : cylindrique
x=nx/2+i*nx # x centre des cellules
y=math.sqrt( (ny/2+j*ny-ly/2)**2 + (nz/2+k*nz-lz/2)**2 ) # y centre des cellules
localpoint = Point(x,y) # création d'un point correspondant au centre de la cellule
# point le plus près sur l'isoligne de la flamme
#nearest_point_on_isoline = isoline.interpolate(isoline.project(localpoint))
# distance entre le point de la flamme et le centre de la cellule
dist = localpoint.distance(isoline)
# distance négative dans les gaz brulés
if (domaine_bunrt.contains(localpoint)):
dist = - dist
#z=(x*math.cos ( math.radians ( theta ) )+ y*math.sin ( math.radians ( theta ) ))
z=dist+flame_pos
g=int(z/(nxc))
# valeur de rho et Y1 sur la cellule par interpolation du profil cantera
if (g < 0):
CI_rho[i,j,k]=rho[0]
CI_Y[i,j,k]=Y1[0]
elif (g > gmax-2):
CI_rho[i,j,k]=rho[gmax-1]
CI_Y[i,j,k]=Y1[gmax-1]
else:
CI_rho[i,j,k]=(rho[g]+(((rho[g+1]-rho[g])/(x1[g+1]-x1[g])*(z-x1[g]))))
CI_Y[i,j,k]=(Y1[g]+(((Y1[g+1]-Y1[g])/(x1[g+1]-x1[g])*(z-x1[g]))))
print('End filling loop')
################################################
# WRITTING FILES THAT ARE READ BY RESIDENT
################################################
#
# Number of cells stored in a specific file
filename = 'info.dat'
f = open(filename,'w')
f.write(str(nl_x)) #number of cells in x
f.write(' ')
f.write(str(nl_y)) #number of cells in y
f.write(' ')
f.write(str(nl_z))
f.close()
print('info.dat saved')
#
# Writting initial condition file for rho and Y
filename1 = 'Density.dat'
filename2 = 'Y1.dat'
f1 = open(filename1,'w')
f2 = open(filename2,'w')
for k in range(int(nl_z)):
for j in range(int(nl_y)):
for i in range(nl_x):
f1.write('%13.5e' %CI_rho[i,j,k]) #density
f2.write('%13.5e' %CI_Y[i,j,k]) #Y1
f1.write(' ')
f2.write(' ')
f1.write('\n')
f2.write('\n')
# for j in range(int(nl_y/2)):
# for i in range(nl_x):
# jsym=int(nl_y/2)-1-j
# f1.write('%13.5e' %CI_rho[i,jsym]) #density
# f2.write('%13.5e' %CI_Y[i,jsym]) #Y1
# f1.write(' ')
# f2.write(' ')
# f1.write('\n')
# f2.write('\n')
f1.close()
f2.close()
print('Density.dat and Y1.dat saved')
print("dimension to put in data.inp : ")
print("x=",nl_x*nx)
print("y=",nl_y*ny)
print("y=",nl_z*nz)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/abulhasanat/MachineLearning/blob/master/DMOZ_BERT_Multiclass_Web_Classification.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
This program is to train a model on a sample of dmoz datasets to classify a webpage. This is developed for POC purpose. Lots of work to be done to traing complete training data.
```
import numpy as np
import pandas as pd
from pathlib import Path
from typing import *
import torch
import torch.optim as optim
from fastai import *
from fastai.vision import *
from fastai.text import *
from fastai.callbacks import *
%%bash
pip install pytorch-pretrained-bert
class Config(dict):
def __init__(self, **kwargs):
super().__init__(**kwargs)
for k, v in kwargs.items():
setattr(self, k, v)
def set(self, key, val):
self[key] = val
setattr(self, key, val)
config = Config(
testing=True,
bert_model_name="bert-base-uncased",
max_lr=3e-5,
epochs=4,
use_fp16=True,
bs=32,
discriminative=False,
max_seq_len=256,
)
from pytorch_pretrained_bert import BertTokenizer
bert_tok = BertTokenizer.from_pretrained(
config.bert_model_name,
)
def _join_texts(texts:Collection[str], mark_fields:bool=False, sos_token:Optional[str]=BOS):
"""Borrowed from fast.ai source"""
if not isinstance(texts, np.ndarray): texts = np.array(texts)
if is1d(texts): texts = texts[:,None]
df = pd.DataFrame({i:texts[:,i] for i in range(texts.shape[1])})
text_col = f'{FLD} {1} ' + df[0].astype(str) if mark_fields else df[0].astype(str)
if sos_token is not None: text_col = f"{sos_token} " + text_col
for i in range(1,len(df.columns)):
#text_col += (f' {FLD} {i+1} ' if mark_fields else ' ') + df[i]
text_col += (f' {FLD} {i+1} ' if mark_fields else ' ') + df[i].astype(str)
return text_col.values
class FastAiBertTokenizer(BaseTokenizer):
"""Wrapper around BertTokenizer to be compatible with fast.ai"""
def __init__(self, tokenizer: BertTokenizer, max_seq_len: int=128, **kwargs):
self._pretrained_tokenizer = tokenizer
self.max_seq_len = max_seq_len
def __call__(self, *args, **kwargs):
return self
def tokenizer(self, t:str) -> List[str]:
"""Limits the maximum sequence length"""
return ["[CLS]"] + self._pretrained_tokenizer.tokenize(t)[:self.max_seq_len - 2] + ["[SEP]"]
from google.colab import drive
drive.mount('/content/drive')
# from sklearn.model_selection import train_test_split
# # DATA_ROOT = Path("..") / "input"
DATA_ROOT=Path('/content/drive/My Drive/Data/DMOZ')
df = pd.read_csv(DATA_ROOT / "train.csv")
# train, test = [pd.read_csv(DATA_ROOT / fname,sep='\t') for fname in ["train1.csv", "test1.csv"]]
# val = train # we won't be using a validation set but you can easily create one using train_test_split
try:
df.drop(['Unnamed: 0'],axis=1,inplace=True)
except:
pass
df_label= pd.read_csv(DATA_ROOT / 'label.csv')
df=pd.merge(
left=df,
right=df_label[['cat_id','top_category']],
on='cat_id',
how='inner'
)
train=pd.concat([df,pd.get_dummies(df['top_category'])],axis=1)
train.sample(5)
del df
# try:
# df.drop(['Unnamed: 0'],axis=1,inplace=True)
# except:
# pass
# df.dropna(inplace=True)
# df['cat_id']=df['cat_id'].astype(int)
# sample=False
# DATA_ROOT=Path('/content/drive/My Drive/Data/DMOZ')
# df = pd.read_csv(DATA_ROOT / "train.csv")
# try:
# df.drop(['Unnamed: 0'],axis=1,inplace=True)
# except:
# pass
# df.dropna(inplace=True)
# df['cat_id']=df['cat_id'].astype(int)
# if sample==True:
# df_sample=df[df['cat_id']<30]
# train=pd.concat([df_sample,pd.get_dummies(df_sample['cat_id'], prefix='label')],axis=1)
# df=df_sample
# else:
# # train=df
# train=pd.concat([df,pd.get_dummies(df['cat_id'], prefix='label')],axis=1)
# val=train
# del df
# train, test = train_test_split(train, random_state=42, test_size=0.33, shuffle=True)
val=train
# label_cols=[]
text='Top Stories After Huge Win, Boris Johnson Promises Brexit By Jan 31, "No Ifs, No Buts" States Not Empowered To Block Citizenship Act, Say Government Sources Spent 66% Of Rs 3.38 Lakh Crore Budgeted Expenditure: Economic Advisor \'PM Should Apologise\': Rahul Gandhi Tweets Video Amid "Rape In India" Row "Won\'t Apologise," Says Rahul Gandhi Amid Row Over "Rape In India" Remark Watch: Sri Lanka Player\'s Hilarious Response To Pakistan Journalist More cricket Trending Watch: Steve "Flying" Smith Takes One Of The Best Catches You\'ll Ever See Reviews More Gadgets Reviews Samsung Galaxy A50, Galaxy A70, Galaxy S9 समत कई समसग समरटफन पर बपर डसकउट Samsung Galaxy M11 और Galaxy M31 अगल सल ह सकत ह लनच चर रयर कमर वल Samsung Galaxy A71 और Samsung Galaxy A51 लनच WhatsApp अगल सल स कई समरटफन पर नह करग कम PUBG Mobile in India May Get Privacy Destroying Features Will Nintendo\'s New Switch Consoles Be Better than the PS4 Pro?', 'Tamil Tamil परवततर म CAB क खलफ हसक परदरशन, गवहट म पलस क फयरग म 2 लग क मत गर BJP शसत रजय म CAB क वरध शर, अब पजब क सएम अमरदर सह न कह- बल असवधनक Aus Vs NZ: टम सउद न मर बललबज क गद, बच म भड गए वरनर, बल- \'उसक हथ म लग ह...\' दख Video UK Elections: एगजट पल म पएम बरस जनसन क कजरवटव परट क सपषट बहमत Bravo "Excited About Comeback" After Return To International Cricket Steven Gerrard Signs New Deal At Rangers Until 2024 Greenwood Stars As Man United Top Group, Arsenal Draw In Europa League November Trade Deficit Narrows To $12.', '12 Billion Food Bangladesh Asks India To Increase Guwahati Mission Security Amid Protests தமழ சனம நடகர சததரத அமசசர ஜயகமரகக பதலட..!', 'இநத வரம வளயகம எககசசகக தமழ படஙகள..!', "வபவ-இன டண' பட ரலஸ தத அறவபப..!", 'ஜய-அதலய ரவ ஜட சரம இரணடவத படம..!', 'டடடல வளயடட வறறமறன..!', 'Osteoporosis - Love your Bones Follow These Amazing Tips By Dr Kiran Lohia To Prevent Acne Breakouts Offbeat Baby Yoda To Disappointed Pakistani Fan: A Look At The Best Memes Of 2019 Biggest Parliament Majority For Boris Johnson\'s Party Since Thatcher Days South News No Top Court Order On Plea Of 2 Women For Protection To Enter Sabarimala Cities Nearly 7,000 Trees To Be Cut For Jewar Airport In Uttar Pradesh "We Have Been Cheated...": Teachers After Left Out Of Recruitment Process Campaigns 60,000 Blankets Needed: Help Save Lives, Donate A Blanket For The Homeless, Here\'s How Fighting Our Killer Air Pollution: Check The Air Quality Index Of Your City Chhattisgarh Becomes The Most Efficient State In Waste Management: Government A Startup In Uttarakhand Develops An Eco-Friendly Sanitary Pad That Lasts Five Times Longer Than Regular Pads'
num_labels=df_label['top_category'].nunique()
data={'cat_id':[0],
'category':[text],
}
test=pd.DataFrame(data)
for labels in df_label['top_category'].unique():
# label='label_'+str(labels)
# label_cols.append(label)
test[labels]=0
test=test.append(test)
test.sample()
if config.testing:
train = train.head(1024)
val = val.head(1024)
test = test.head(1024)
fastai_bert_vocab = Vocab(list(bert_tok.vocab.keys()))
fastai_tokenizer = Tokenizer(tok_func=FastAiBertTokenizer(bert_tok, max_seq_len=config.max_seq_len), pre_rules=[], post_rules=[])
label_cols=df_label['top_category'].unique().tolist()
label_cols #[2:]
# label_cols = label_cols[2:]
# label_cols=[str(item) for item in label_cols]
label_cols[0]
# label_cols = ["toxic", "severe_toxic", "obscene", "threat", "insult", "identity_hate"]
class BertTokenizeProcessor(TokenizeProcessor):
def __init__(self, tokenizer):
super().__init__(tokenizer=tokenizer, include_bos=False, include_eos=False)
class BertNumericalizeProcessor(NumericalizeProcessor):
def __init__(self, *args, **kwargs):
super().__init__(*args, vocab=Vocab(list(bert_tok.vocab.keys())), **kwargs)
def get_bert_processor(tokenizer:Tokenizer=None, vocab:Vocab=None):
"""
Constructing preprocessors for BERT
We remove sos/eos tokens since we add that ourselves in the tokenizer.
We also use a custom vocabulary to match the numericalization with the original BERT model.
"""
return [BertTokenizeProcessor(tokenizer=tokenizer),NumericalizeProcessor(vocab=vocab)]
class BertDataBunch(TextDataBunch):
@classmethod
def from_df(cls, path:PathOrStr, train_df:DataFrame, valid_df:DataFrame, test_df:Optional[DataFrame]=None,
tokenizer:Tokenizer=None, vocab:Vocab=None, classes:Collection[str]=None, text_cols:IntsOrStrs=1,
label_cols:IntsOrStrs=0, label_delim:str=None, **kwargs) -> DataBunch:
"Create a `TextDataBunch` from DataFrames."
p_kwargs, kwargs = split_kwargs_by_func(kwargs, get_bert_processor)
# use our custom processors while taking tokenizer and vocab as kwargs
processor = get_bert_processor(tokenizer=tokenizer, vocab=vocab, **p_kwargs)
if classes is None and is_listy(label_cols) and len(label_cols) > 1: classes = label_cols
src = ItemLists(path, TextList.from_df(train_df, path, cols=text_cols, processor=processor),
TextList.from_df(valid_df, path, cols=text_cols, processor=processor))
src = src.label_for_lm() if cls==TextLMDataBunch else src.label_from_df(cols=label_cols, classes=classes)
if test_df is not None: src.add_test(TextList.from_df(test_df, path, cols=text_cols))
return src.databunch(**kwargs)
databunch = BertDataBunch.from_df(".", train, val, test,
tokenizer=fastai_tokenizer,
vocab=fastai_bert_vocab,
text_cols="category",
label_cols=label_cols,
bs=config.bs,
collate_fn=partial(pad_collate, pad_first=False, pad_idx=0),
)
from pytorch_pretrained_bert.modeling import BertConfig, BertForSequenceClassification
bert_model = BertForSequenceClassification.from_pretrained(config.bert_model_name, num_labels=num_labels)
loss_func = nn.BCEWithLogitsLoss()
from fastai.callbacks import *
device_cuda = torch.device("cuda")
learner = Learner(
databunch, bert_model,
loss_func=loss_func
)
if config.use_fp16: learner = learner.to_fp16()
# learner.to_fp32()
# del df
learner.lr_find()
learner.recorder.plot()
# learner.fit_one_cycle(config.epochs, max_lr=config.max_lr)
# learner.fit_one_cycle(config.epochs, max_lr=1e-02)
learner.fit_one_cycle(8, max_lr=config.max_lr)
train.cat_id.nunique()
learner.export(file=DATA_ROOT/'Model_DMOZ(all_cat_v1_2).pkl')
# learner.export(file=DATA_ROOT/'Model_DMOZ(25_cat).pkl')
def get_preds_as_nparray(ds_type) -> np.ndarray:
"""
the get_preds method does not yield the elements in order by default
we borrow the code from the RNNLearner to resort the elements into their correct order
"""
preds = learner.get_preds(ds_type)[0].detach().cpu().numpy()
sampler = [i for i in databunch.dl(ds_type).sampler]
reverse_sampler = np.argsort(sampler)
return preds[reverse_sampler, :]
# learner.load(file=DATA_ROOT/'model1.pkl')
test_preds = get_preds_as_nparray(DatasetType.Test)
# sample_submission = pd.read_csv(DATA_ROOT / "sample_submission.csv")
# if config.testing: sample_submission = sample_submission.head(test.shape[0])
# sample_submission[label_cols] = test_preds
# sample_submission.to_csv(DATA_ROOT /"predictions.csv", index=False)
sample_submission=test
for label in label_cols:
sample_submission[label]=0
sample_submission[label_cols] = test_preds
# sample_submission.to_csv(DATA_ROOT /"predictions_1.csv", index=False)
# sample_submission.sample(10000).to_csv(DATA_ROOT /"predictions_sample.csv", index=False)
sample_submission.head(5)
output=sample_submission.head(1).drop(columns=['cat_id','category']).T
output
output[0]=output[0].astype(float)
output.nlargest(5,[0])
sample_submission.to_csv(DATA_ROOT /"predictions_test.csv", index=False)
# learner.export(file=DATA_ROOT /"Model1.pkl")
# learner.save(file=DATA_ROOT/'Final_Model_BERT.pkl')
learner.validate()
text='Efforts were being made on Friday to help passengers stranded at the airport, railway station and inter-state bus terminals in Guwahati, an official said.'
x=learner.predict(text)
y=pd.DataFrame(x[2],index=label_cols)
# x[2].shape
y.nlargest(5,[0])
```
| github_jupyter |
# Deep Learning approach to the Boston Housing dataset - Autoencoder
## Team JARL
```
#import libraries and load data
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import keras
from keras import regularizers, layers
from keras.models import Model, load_model, Sequential
from keras.layers import Input, Dense
from keras.callbacks import ModelCheckpoint, TensorBoard
from keras.optimizers import Adam
from tensorflow.keras.callbacks import EarlyStopping
from sklearn.metrics import r2_score
from sklearn.ensemble.partial_dependence import partial_dependence, plot_partial_dependence
data = pd.read_csv('boston_corrected.csv')
data = data.sample(frac=1).reset_index(drop=True)
train = data.iloc[:450,:]
test = data.iloc[450:,:]
# Drop undesired columns
rem = ['MEDV', 'CMEDV','TOWN','TOWNNO','TRACT']
raw_X_train = train.drop(rem, axis=1)
raw_Y_train = train['CMEDV']
raw_X_test = test.drop(rem, axis=1)
raw_Y_test = test['CMEDV']
# Set all column types to numeric
raw_X_train = np.asarray(raw_X_train).astype(np.float32)
raw_X_test = np.asarray(raw_X_test).astype(np.float32)
raw_Y_train = np.asarray(raw_Y_train).astype(np.float32)
raw_Y_test = np.asarray(raw_Y_test).astype(np.float32)
def Z_scoreNorm(data):
column_mean = data.mean(axis=0)
column_std = np.std(data, axis=0, dtype=float)
Normalized_Inputs = (data - column_mean)/column_std
return Normalized_Inputs
X_train = Z_scoreNorm(raw_X_train)
X_test = Z_scoreNorm(raw_X_test)
Y_train = Z_scoreNorm(raw_Y_train)
Y_test = Z_scoreNorm(raw_Y_test)
# Reality check
print(X_train.shape)
print(X_test.shape)
early_stopping = EarlyStopping(
min_delta=0.001,
patience=100,
restore_best_weights=True,
)
choke = 10
autoencoder = Sequential([
Dense(X_train.shape[1], activation='relu', input_shape=(X_train.shape[1],)),
Dense(20, activation='relu'),
Dense(20, activation='relu'),
Dense(choke, activation='relu'),
Dense(20, activation='relu'),
Dense(20, activation='relu'),
Dense(X_train.shape[1])
])
autoencoder.compile(optimizer=Adam(lr=0.01),
loss='MSE',
metrics=['mean_squared_error'])
checkpointer_a = ModelCheckpoint(filepath="AE.z1",
verbose=0,
save_best_only=True)
history_a = autoencoder.fit(X_train, X_train,
epochs=800,
batch_size=450,
shuffle=True,
validation_data=(X_test, X_test),
verbose=1,
callbacks=[early_stopping, checkpointer_a]).history
plt.plot(history_a['loss'])
plt.plot(history_a['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper right');
plt.savefig('losses_upstream.png', dpi=300)
autoencoder.summary()
# Reality check
a = 5
testinp = np.array(X_train[a,:]).reshape(1,X_train.shape[1])
y = autoencoder.predict(testinp)
print(np.array(X_train[a,:]))
print(y)
valuePredictor = Sequential([
Dense(choke, activation='relu', input_shape=(choke,)),
Dense(20, activation='relu'),
Dense(20, activation='relu'),
Dense(20, activation='relu'),
Dense(20, activation='relu'),
Dense(1)
])
# Latent feature extraction
latent_model = Model(autoencoder.input, autoencoder.get_layer(index = 3).output )
latents_train = np.array([latent_model.predict(i.reshape(1, X_train.shape[1])) for i in X_train]).reshape(450, choke)
latents_test = np.array([latent_model.predict(i.reshape(1, X_train.shape[1])) for i in X_test]).reshape(56, choke)
valuePredictor.compile(optimizer=Adam(lr=0.03),
loss='MSE',
metrics=['mean_squared_error'])
checkpointer_1 = ModelCheckpoint(filepath="valuePredictor.z1",
verbose=0,
save_best_only=True)
history_1 = valuePredictor.fit(latents_train, Y_train,
epochs=700,
batch_size=450,
shuffle=True,
validation_data=(latents_test, Y_test),
verbose=1, callbacks=[early_stopping, checkpointer_1]).history
plt.plot(history_1['loss'])
plt.plot(history_1['val_loss'])
plt.title('Model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper right');
plt.savefig('losses_downstream.png', dpi=300)
valuePredictor_baseline = Sequential([
Dense(X_train.shape[1], activation='relu', input_shape=(X_train.shape[1],)),
Dense(20, activation='relu'),
Dense(20, activation='relu'),
Dense(20, activation='relu'),
Dense(10, activation='relu'),
Dense(1)
])
valuePredictor_baseline.compile(optimizer=Adam(lr=0.01),
loss='MSE',
metrics=['mean_squared_error'])
checkpointer_2 = ModelCheckpoint(filepath="valuePredictor_baseline.z1",
verbose=0,
save_best_only=True)
history_2 = valuePredictor_baseline.fit(X_train, Y_train,
epochs=700,
batch_size=450,
shuffle=True,
validation_data=(X_test, Y_test),
verbose=1, callbacks=[early_stopping, checkpointer_2]).history
# Test
ae_preds = np.zeros((56,))
base_preds = np.zeros((56,))
for i, record in enumerate(X_test):
ae_preds[i] = valuePredictor.predict(latent_model.predict(np.array(record).reshape(1, X_train.shape[1])))
base_preds[i] = valuePredictor_baseline.predict(np.array(record).reshape(1, X_train.shape[1]))
plt.scatter(ae_preds, Y_test, color='r', s=5, label='AE - $r^2 = 0.88$')
plt.scatter(base_preds, Y_test, color='g', s=5, label='Base - $r^2 = 0.83$')
plt.plot(Y_test,Y_test)
plt.xlabel("True prices (Normalised)")
plt.ylabel("Predicted prices (Normalised)")
plt.legend()
plt.savefig('predictions.png', dpi=300)
print("RMSE (Autoencoder):", np.sqrt((Y_test - ae_preds)**2).mean(axis=None))
print("r2 (Autoencoder):", r2_score(Y_test, ae_preds))
print("RMSE (Baseline):", np.sqrt((Y_test - base_preds)**2).mean(axis=None))
print("r2 (Baseline):", r2_score(Y_test, base_preds))
```
| github_jupyter |
```
import os
import filecmp
import h5py
import numpy as np
import matplotlib.pyplot as plt
from math import isnan
DATASET_PATH = '/home/elliot/Desktop/cnn_denoising_dataset'
# I combined e.g. `train_01` and `train_02` into `train`
# `train_road` is all unlabeled, probably it's meant to be augmented
SETS = ['test', 'train', 'val']
# 0: no label, 100: valid/clear, 101: rain, 102: fog
LABEL_NAMES = {0: 'no label', 100: 'clear', 101: 'rain', 102: 'fog'}
```
# Investigate Sample File
```
sample_file = os.path.join(DATASET_PATH, 'test', '2018-11-29_114626_Static2-Day-Clear', 'LidarImage_000000607.hdf5')
sample_file = h5py.File(sample_file, 'r')
sample_file.keys()
sample_file['distance_m_1'], sample_file['intensity_1'], sample_file['labels_1']
plt.imshow(sample_file['distance_m_1'])
plt.imshow(sample_file['intensity_1'])
plt.imshow(sample_file['labels_1'])
# The sample file is all 'clear'
plt.hist(sample_file['labels_1'][()].flatten(), bins=4)
```
# Count Samples
"In total, the data set contains about 175,941 samples for
training, validation and testing containing chamber (72,800)
and road (103,141) scenes, which can be used thanks to
augmentation"
Our counts are less, since we haven't done augmentation (yet)
```
# Count samples in each training set
set_counts = {set_name: 0 for set_name in SETS}
for set_name in set_counts:
for root, dirs, files in os.walk(os.path.join(DATASET_PATH, set_name)):
set_counts[set_name] += len(files)
total_samples = sum(count for name, count in set_counts.items())
print(f'Total samples: {total_samples}')
for name, count in set_counts.items():
print(f'{name}: {(count / total_samples * 100):.2f}%')
# The total above from the original paper probably includes augmented samples
# Number of samples that are majority `label`, skipping 0/no label
sample_counts = {label: 0 for label in LABEL_NAMES.keys() if label != 0}
# Total number of points labeled with `label`
label_counts = {label: 0 for label in LABEL_NAMES.keys() if label != 0}
# Samples that have any adverse weather at all (rain or fog)
adverse_samples = {label: 0 for label in LABEL_NAMES.keys() if label != 0}
for set_name in SETS:
for root, dirs, files in os.walk(os.path.join(DATASET_PATH, set_name)):
for file in files:
file = h5py.File(os.path.join(root, file), 'r')
labels = file['labels_1'][()].flatten()
labels, counts = np.unique(labels, return_counts=True)
counts = {label: count for label, count in zip(labels, counts)}
# Remove where label == 0 (0 = no label)
if 0 in counts:
del counts[0]
dominant_label = max(counts, key=counts.get)
sample_counts[dominant_label] += 1
for label, count in counts.items():
label_counts[label] += count
# Count any adverse weather: 101 = rain, 102 = fog
# Note that a sample could have both rain and fog
if 101 in counts or 102 in counts:
if 101 in counts:
adverse_samples[101] += 1
if 102 in counts:
adverse_samples[102] += 1
else:
adverse_samples[100] += 1
print(set_name, sample_counts, label_counts, adverse_samples)
sample_counts, label_counts, adverse_samples
assert(sum(sample_counts.values()) == total_samples)
print('Dominant sample labels')
for label, count in sample_counts.items():
print(f'{LABEL_NAMES[label]}: {(count / total_samples * 100):.2f}%')
total_points = sum(label_counts.values())
print('Labeled points')
for label, count in label_counts.items():
print(f'{LABEL_NAMES[label]}: {(count / total_points * 100):.2f}%')
# May be more than `total_samples` if some samples have rain and fog
total_adverse_samples = sum(adverse_samples.values())
print('Samples with any adverse weather')
for label, count in adverse_samples.items():
print(f'{LABEL_NAMES[label]}: {(count / total_adverse_samples * 100):.2f}%')
# Confirming that all `train_road` samples are unlabeled (all labels are 0)...probably meant to be augmented
for root, dirs, files in os.walk(os.path.join(DATASET_PATH, 'train_road')):
for file in files:
file_path = os.path.join(root, file)
file = h5py.File(file_path, 'r')
labels = file['labels_1'][()].flatten()
if not np.all(labels == 0):
print(file_path)
print(root, 'complete')
```
# Investigate File Pairs
Some samples are pairs of files, e.g. `[original].hdf5` and `[original]_2.hdf5`. Unclear what this means, see this issue: https://github.com/rheinzler/PointCloudDeNoising/issues/8
```
file_names = []
for set_name in SETS:
for root, dirs, files in os.walk(os.path.join(DATASET_PATH, set_name)):
file_names.extend(os.path.join(root, file) for file in files)
assert len(file_names) == total_samples
pairs = list(filter(lambda f: f.endswith('_2.hdf5'), file_names))
print(f'Pairs found: {len(pairs)}')
def hdf5_equal(a, b):
if a.keys() != file2.keys(): return False
for key in a.keys():
if not np.all(a[key][()] == file2[key][()]): return False
if list(a.attrs) != list(file2.attrs): return False
for attr in file1.attrs:
aa, ba = file1.attrs[attr], file2.attrs[attr]
if not aa == ba and not (isnan(aa) and isnan(ba)): return False
return True
# Runs successfully, files appear to be equal
for file2_path in pairs:
file1_path = file2_path[:-7] + '.hdf5'
file1, file2 = h5py.File(file1_path, 'r'), h5py.File(file2_path, 'r')
assert hdf5_equal(file1, file2), (file1_path, file2_path)
filecmp.cmp(
os.path.join(DATASET_PATH, 'train', '2018-11-29_145021_Static1-Day-Clear', 'LidarImage_000000010.hdf5'),
os.path.join(DATASET_PATH, 'train', '2018-11-29_145021_Static1-Day-Clear', 'LidarImage_000000010_2.hdf5'),
shallow=False
)
```
The contents of each pair (and the data that we'd be using) appear to be equal, but the files do not compare equally...could be some additional metadata that isn't captured by `h5py`?
| github_jupyter |
# Working with *concepts* in the Dimensions API
This Python notebook shows how to use the [Dimensions Analytics API](https://www.dimensions.ai/dimensions-apis/) in order to extract `concepts` from documents and use them as the basis for more advanced topic-analysis tasks.
```
import datetime
print("==\nCHANGELOG\nThis notebook was last run on %s\n==" % datetime.date.today().strftime('%b %d, %Y'))
```
## Prerequisites
This notebook assumes you have installed the [Dimcli](https://pypi.org/project/dimcli/) library and are familiar with the ['Getting Started' tutorial](https://api-lab.dimensions.ai/cookbooks/1-getting-started/1-Using-the-Dimcli-library-to-query-the-API.html).
```
!pip install dimcli plotly -U --quiet
import dimcli
from dimcli.utils import *
import json
import sys
import pandas as pd
import plotly.express as px
if not 'google.colab' in sys.modules:
# make js dependecies local / needed by html exports
from plotly.offline import init_notebook_mode
init_notebook_mode(connected=True)
#
print("==\nLogging in..")
# https://digital-science.github.io/dimcli/getting-started.html#authentication
ENDPOINT = "https://app.dimensions.ai"
if 'google.colab' in sys.modules:
import getpass
KEY = getpass.getpass(prompt='API Key: ')
dimcli.login(key=KEY, endpoint=ENDPOINT)
else:
KEY = ""
dimcli.login(key=KEY, endpoint=ENDPOINT)
dsl = dimcli.Dsl()
```
## 1. Background: What are `concepts`?
Concepts are normalized *noun phrases* describing the main topics of a document (see also the [official documentation](https://docs.dimensions.ai/dsl/language.html#searching-using-concepts)). Concepts are automatically derived from documents abstracts using machine learning techniques, and are ranked based on their relevance.
In the JSON data, concepts are available as an ordered list (=first items are the most relevant), including a relevance score. E.g. for the publications with ID 'pub.1122072646':
```
{'id': 'pub.1122072646',
'concepts_scores': [{'concept': 'acid', 'relevance': 0.07450046286579201},
{'concept': 'conversion', 'relevance': 0.055053872555463006},
{'concept': 'formic acid', 'relevance': 0.048144671935356},
{'concept': 'CO2', 'relevance': 0.032150964737607}
[........]
],
}
```
Please note that (as of version 1.25 of the DSL API) it is possible to return either `concepts_scores` or `concepts` with Publications queries, but only `concepts` with Grants queries.
### 1.1 From concepts to dataframes: Dimcli's `as_dataframe_concepts` method
A Dimensions API query normally returns a list of documents (publications, grants), where each document includes a list of concepts.
In order to analyse concepts more easily, it is useful to 'unnest' concepts into a new structure where each concept is a top level item. In other words, we want to transform the original documents table into a concepts table.
The [Dimcli](https://pypi.org/project/dimcli/) library provides a [method](https://api-lab.dimensions.ai/cookbooks/1-getting-started/3-Working-with-dataframes.html#4.-Dataframe-Methods-for-%E2%80%98Concepts%E2%80%99-queries) that does exactly that: `as_dataframe_concepts()`.
```
q = """search publications for "graphene"
where year=2019
return publications[id+title+year+concepts_scores] limit 100"""
concepts = dsl.query(q).as_dataframe_concepts()
concepts.head(5)
```
The `as_dataframe_concepts()` method internally uses pandas to [explode](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.explode.html) the concepts list, plus it adds some extra metrics that are handy in order to carry out further analyses:
1. **concepts_count**: the total number of concepts for each single document. E.g., if a document has 35 concepts, concepts_count=35.
2. **frequency**: how often a concept occur within a dataset, i.e. how many documents include that concept. E.g., if a concept appears in 5 documents, frequency=5.
3. **score**: the relevancy of a concept in the context of the document it is extracted from. Concept scores go from 0 (= not relevant) to 1 (= very relevant). NOTE if concepts are returned without scores, these are generated automatically by normalizing its ranking against the total number of concepts for a single document. E.g., if a document has 10 concepts in total, the first concept gets a score=1, the second score=0.9, etc..
4. **score_avg**: the average (mean) value of all scores of a concept across multiple documents, within a given in a dataset.
As we will see, by sorting and segmenting data using these parameters, it is possible to filter out common-name concepts and highlight more interesting ones.
### 1.2 Extracting concepts from any text
This tutorial focuses on concepts obtained from publications available via Dimensions. However, it is also possible to take advantage of Dimensions NLP infrastructure to extract concepts from any text.
This can be achieved by using the DSL function [extract_concepts](https://docs.dimensions.ai/dsl/functions.html#extract-concepts) and passing an abstract-length text as an argument.
For example:
```
abstract = """We describe monocrystalline graphitic films, which are a few atoms thick but are nonetheless stable under ambient conditions,
metallic, and of remarkably high quality. The films are found to be a two-dimensional semimetal with a tiny overlap between
valence and conductance bands, and they exhibit a strong ambipolar electric field effect such that electrons and
holes in concentrations up to 10 per square centimeter and with room-temperature mobilities of approximately 10,000 square
centimeters per volt-second can be induced by applying gate voltage.
"""
res = dsl.query(f"""extract_concepts("{abstract}", return_scores=true)""")
pd.DataFrame(res['extracted_concepts'])
```
## 2. Data acquisition: retrieving publications and all their associated concepts
Let's pull all publications from University College London classified with the FOR code "16 Studies in Human Society".
> Tip: you can experiment by changing the parameters below as you want, eg by [choosing another GRID organization](https://grid.ac/institutes).
```
GRIDID = "grid.83440.3b" #@param {type:"string"}
FOR = "16 Studies in Human Society" #@param {type:"string"}
query = f"""
search publications
where research_orgs.id = "{GRIDID}"
and category_for.name= "{FOR}"
return publications[id+doi+concepts_scores+year]
"""
print("===\nQuery:\n", query)
print("===\nRetrieving Publications.. ")
data = dsl.query_iterative(query)
```
Let's turn the results into a dataframe and have a quick look at the data. You'll see a column `concepts_scores` that contains a list of concepts for each of the publications retrieved.
```
pubs = data.as_dataframe()
pubs.head(5)
```
### 2.1 Processing concept data
Now it’s time to start digging into the ‘concepts’ column of publications.
Each publications has an associated **list** of concepts, so in order to analyse them we need to ‘explode’ that list so to have a new table with one row per concept.
```
concepts = data.as_dataframe_concepts()
print("===\nConcepts Found (total):", len(concepts))
print("===\nPreview:")
display(concepts)
```
If we ignore the publications metadata from the concepts list and drop duplicates, we can obtain a new table with **unique** concepts.
```
concepts_unique = concepts.drop_duplicates("concept")[['concept', 'frequency', 'score_avg']]
print("===\nUnique Concepts Found:", len(concepts_unique))
print("===\nPreview:")
display(concepts_unique)
```
## 3. Exploring our dataset: basic statistics about Publications / Concepts
In this section we'll show how to get an overview of the concepts data we obtained.
These statistics are important because they will help us contextualize more in-depth analyses of the concepts data we'll do later on.
### 3.1 Documents With concepts VS Without
You'll soon discover that not all documents have associated concepts (eg cause there's no text to extract them from, in some cases).
Let's see how many:
```
CONCEPTS_FIELD = "concepts_scores"
df = pd.DataFrame({
'type': ['with_concepts', 'without_concepts'] ,
'count': [pubs[CONCEPTS_FIELD].notnull().sum(), pubs[CONCEPTS_FIELD].isnull().sum()]
})
px.pie(df,
names='type', values="count",
title = "How many documents have concepts?")
```
### 3.2 Yearly breakdown of Documents With concepts VS Without
It's also useful to look at whether the ratio of with/without concepts is stable across the years.
To this end we can use
* the publications `id` column to count the total number of publications per year
* the `concepts` column to count the ones that have concepts
```
temp1 = pubs.groupby('year', as_index=False).count()[['year', 'id', CONCEPTS_FIELD]]
temp1.rename(columns={'id': "documents", CONCEPTS_FIELD: "with_concepts"}, inplace=True)
# reorder cols/rows
temp1 = temp1.melt(id_vars=["year"],
var_name="type",
value_name="count")
px.bar(temp1, title="How many documents have concepts? Yearly breakdown.",
x="year", y="count",
color="type",
barmode="group")
```
### 3.3 Concepts frequency
It is useful to look at how many concepts appear more than once in our dataset. As you'll discovert, is often the case that only a subset of concepts appear more than once. That is because documents tend to be highly specialised hence a large number of extracted noun phrases aren't very common.
By looking at this basic frequency statistics we can determine a useful **frequency threshold** for our analysis - ie to screen out concepts that are not representative of the overall dataset we have.
Tip: change the value of `THRESHOLD` to explore the data.
```
THRESHOLD = 2
df = pd.DataFrame({
'type': [f'freq<{THRESHOLD}',
f'freq={THRESHOLD}',
f'freq>{THRESHOLD}'] ,
'count': [concepts_unique.query(f"frequency < {THRESHOLD}")['concept'].count(),
concepts_unique.query(f"frequency == {THRESHOLD}")['concept'].count(),
concepts_unique.query(f"frequency > {THRESHOLD}")['concept'].count()]
})
px.pie(df,
names='type', values="count",
title = f"Concepts with a frequency major than: {THRESHOLD}")
```
### 3.4 Distribution of Concepts Frequency
It is useful to chart the overall distribution of how frequent concepts are.
The bottom-left section of the chart shows the segment we are most likely to focus on, so to avoid concepts that appear only once, or the long-tail of highly frequent concepts that are likely to be common-words of little interest.
```
temp = concepts_unique.groupby('frequency', as_index=False)['concept'].count()
temp.rename(columns={'concept' : 'concepts with this frequency'}, inplace=True)
px.scatter(temp,
x="frequency",
y="concepts with this frequency",
title="Distribution of concepts frequencies")
```
### 3.5 Yearly breakdown: unique VS repeated concepts
Also useful to look at the number of concepts per year, VS the number of unique concepts.
This will give us a sense of whether the distribution of repeated concepts is stable across the years.
```
series1 = concepts.groupby("year")['concept'].count().rename("All concepts")
series2 = concepts.groupby("year")['concept'].nunique().rename("Unique concepts")
temp2 = pd.concat([series1, series2], axis=1).reset_index()
temp2 = temp2.melt(id_vars=["year"],
var_name="type",
value_name="count")
px.bar(temp2,
title="Yearly breakdown: Tot concepts VS Unique concepts",
x="year", y="count",
color="type", barmode="group",
color_discrete_sequence=px.colors.carto.Antique)
```
## 4. Isolating 'interesting' concepts using `frequency` and `score_avg`
In this section we will take a deep dive into the concepts themselves, in particular by using the two metrics obtained above: `frequency` and `score_avg`.
The main thing to keep in mind is that **only the combination of these two metrics can lead to interesting results**. In fact, if we used only *frequency* it'll lead to common keywords that are not very relevant; on the other hand, using only *relevancy* will result in concepts that important but just to one or two documents.
### 4.1 The problem: frequent concepts are not *that* interesting!
For example, let's see what happens if we get the top concepts based on `frequency` only:
```
top = concepts_unique.sort_values("frequency", ascending=False)[:20]
px.bar(top,
title="Concepts sorted by frequency",
x="concept", y="frequency",
color="score_avg")
```
Not very interesting at all! Those keywords are obviously very common (eg *study* or *development*) in the scientific literature, but of very little semantic interest.
### 4.2 Solution 1: prefiltering by `score_avg` and sorting by `frequency`
By doing so, we aim at extracting concepts that are both frequent and tend to be very relevant (within their documents).
```
temp = concepts_unique.query("score_avg > 0.6").sort_values("frequency", ascending=False)
px.bar(temp[:50],
title="Concepts with high average score, sorted by frequency",
x="concept", y="frequency",
color="score_avg")
```
### 4.3 Solution 2: prefiltering by `frequency` and sorting by `score_avg`
This method also allows to isolate interesting concepts, even if they are not very frequently appearing in our dataset.
```
temp = concepts_unique.query("frequency > 10 & frequency < 100").sort_values(["score_avg", "frequency"], ascending=False)
px.bar(temp[:100],
title="Concepts with medium frequency, sorted by score_avg",
x="concept", y="score_avg",
height=600,
color="frequency")
```
## 5. Analyses By Year
In this section we will show how to use the methods above together with a yearly segmentation of the documents data. This will allow us to draw up some cool comparison of concepts/topics across years.
### 5.1 Adding year-based metrics to the concepts dataframe
These are the steps
* recalculate *freq* and *score_avg* for each year, using the original concepts dataset from section 2.1
* note this will result in duplicates (as many as the appearances of a concept within the same year), which of course we should remove
```
concepts['frequency_year'] = concepts.groupby(['year', 'concept'])['concept'].transform('count')
concepts['score_avg_year'] = concepts.groupby(['year', 'concept'])['score'].transform('mean').round(5)
concepts_by_year = concepts.copy().drop_duplicates(subset=['concept', 'year'])\
[['year', 'concept', 'frequency_year', 'score_avg_year']]
concepts_by_year.head()
```
For example, let's look at the yearly-distribution of a specific concept: *migrants*
```
concepts_by_year[concepts_by_year['concept'] == "migrants"]
```
### 5.2 Charting the variation: multi-year visualization
We can use Plotly's 'facets' to have subsections that show variation across years. Plotly will plot all the values retrieved - which allows to spot the trends up and down.
* tip: to have an equal representation for each year, we take the top N concepts across a chosen years-span and then look at their frequency distribution over the years
In order to isolate interesting concepts, we can use the same formula from above (filter by score, then sort by frequency). Only this time using yearly values of course!
```
MAX_CONCEPTS = 50
YEAR_START = 2015
YEAR_END = 2019
SCORE_MIN = 0.4
segment = concepts_by_year.query(f"year >= {YEAR_START} & year <= {YEAR_END}").copy()
# create metrics for the segment only
segment['frequency'] = concepts.groupby('concept')['concept'].transform('count')
segment['score_avg'] = concepts.groupby('concept')['score'].transform('mean').round(5)
# get top N concepts for the dataviz
top_concepts = segment.drop_duplicates('concept')\
.query(f"score_avg > {SCORE_MIN}")\
.sort_values("frequency", ascending=False)[:MAX_CONCEPTS]
# use yearly data only for top N concepts
segment_subset = segment[segment['concept'].isin(top['concept'].tolist())]
px.bar(segment_subset,
x="concept",
y="frequency_year",
facet_row="year",
title=f"Top concepts {YEAR_START}-{YEAR_END} with score_avg > {SCORE_MIN}, sorted by frequency",
height=1000,
color="frequency_year")
```
## 6. Conclusion
In this tutorial we have demonstrated how to query for [concepts](https://docs.dimensions.ai/dsl/language.html#searching-using-concepts) using the [Dimensions Analytics API](https://www.dimensions.ai/dimensions-apis/).
### The main takeaways
* concepts can be easily extracted by using the `as_dataframe_concepts()` method
* concepts have an implicit score relative to the document they belong to - but we can create more absolute metrics by normalizing these scores
* it is useful to look at the frequency of concepts in the context of the entire dataset we have
* there can be a long tail of concepts that are very infrequent, hence it's useful to filter those out
* by using a combination of frequency and score_avg metrics, we can filter out uninteresting concepts
### What next
Using these methods, you can take advantage of *concepts* data in a number of real-world scenarios. Here are some ideas:
* you can segment publications using other criteria: eg by *journal* or by *field of research*, in order to identify more specific trends;
* concepts extracted can be used to create new DSL searches - using the `in concepts` [search syntax](https://docs.dimensions.ai/dsl/language.html#in-search-index);
* concepts data can be grouped further using [semantic similarity](https://spacy.io/usage/vectors-similarity) or [clustering](https://scikit-learn.org/stable/modules/clustering.html) techniques;
* you can look at the co-occurence of concepts withing the same document, in order to build a semantic network.
| github_jupyter |
# Softmax and Momentum
### Implemented a softmax classifier using using stochastic gradient descent with mini-batches and momentum to minimize softmax (cross-entropy) loss with L2 weight decay regularization of this single layer neural network

## Import libraries
```
import numpy as np
import random, argparse
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
from matplotlib.axes._axes import _log as matplotlib_axes_logger
matplotlib_axes_logger.setLevel('ERROR')
```
## NeuralNetwork implementation
### Here i implemented class that contains such parameters like:
* epochs: Number of iterations over complete training data
* learningRate: A step size or a learning rate
* batchSize: A mini-batch size(less than total number of training data)
* regStrength: A regularization strength
* momentum: A momentum value
* train
```
Train a softmax classifier model on training data using stochastic gradient descent with mini-batches
and momentum to minimize softmax (cross-entropy) loss of this single layer neural network. It calcualtes
mean per-class accuracy for the training/testing data and the loss.
xTrain: Training input data
yTrain: Training labels
xTest: Testing input data
yTest: Testing labels
return: A tuple of training/Testing losses and Accuracy
```
* SGDWithMomentum
```
Stochastic gradient descent with mini-batches. It divides training data into mini-batches
and compute loss and grad on that mini-batches and updates the weights. It repeats for all samples.
x: An input samples
y: An input labels
return: Total loss computed
```
* softmaxEquation
```
It calculates a softmax probability
z: A matrix(wt * input sample)
return: softmax probability
```
* computeLoss
```
It calculates a cross-entropy loss with regularization loss and gradient to update the weights.
x: An input sample
yMatrix: Label as one-hot encoding
```
* meanAccuracy
```
It calculates mean-per class accuracy
x: Input sample
y: label sample
return: mean-per class accuracy
```
* predict
```
It predict the label based on input sample and a model
x: Input sample
return: predicted label
```
* oneHotEncoding
```
Convert a vector into one-hot encoding matrix where that particular column value is 1 and rest 0 for that row.
y: Label vector
numOfClasses: Number of unique labels
return: one-hot encoding matrix
```
* plotGraph
```
Plot a Epochs vs. Cross Entropy Loss graph
trainLosses: List of training loss over every epochs
testLosses: List of testing loss over every epochs
trainAcc: List of training accuracy over every epochs
testAcc: List of testing accuracy over every epochs
```
* readData
```
Read data from file and divide into input sample and a label.
filename: name of a file
input sample and label
```
* makeMeshGrid
```
Create a mesh point to plot decision boundary.
x: data or sample (for x-axis on meshgrid)
y: label(for y-axis on meshgrid)
h: step size for meshgrid
return: matrix of x-axis and y-axis
```
* plotContours
```
It plot a contour.
plt: A matplotlib.pyplot object
model: softmax classifier model
xx: meshgrid ndarray
yy: meshgrid ndarray
params: Number of parameters to pass to contour function
```
```
class Softmax:
def __init__(self, epochs, learningRate, batchSize, regStrength, momentum):
self.epochs = epochs
self.learningRate = learningRate
self.batchSize = batchSize
self.regStrength = regStrength
self.momentum = momentum
self.velocity = None
self.wt = None
def train(self, xTrain, yTrain, xTest, yTest):
D = xTrain.shape[1] # dimensionality
label = np.unique(yTrain)
numOfClasses = len(label) # number of classes
yTrainEnc = self.oneHotEncoding(yTrain, numOfClasses)
yTestEnc = self.oneHotEncoding(yTest, numOfClasses)
self.wt = 0.001 * np.random.rand(D, numOfClasses)
self.velocity = np.zeros(self.wt.shape)
trainLosses = []
testLosses = []
trainAcc = []
testAcc = []
for e in range(self.epochs): # loop over epochs
trainLoss = self.SGDWithMomentum(xTrain, yTrainEnc)
testLoss, dw = self.computeLoss(xTest, yTestEnc)
trainAcc.append(self.meanAccuracy(xTrain, yTrain))
testAcc.append(self.meanAccuracy(xTest, yTest))
trainLosses.append(trainLoss)
testLosses.append(testLoss)
print("{:d}->TrainL : {:.7f}|\tTestL : {:.7f}|\tTrainAcc : {:.7f}|TestAcc: {:.7f}"
.format(e, trainLoss, testLoss, trainAcc[-1], testAcc[-1]))
return trainLosses, testLosses, trainAcc, testAcc
def SGDWithMomentum(self, x, y):
losses = []
randomIndices = random.sample(range(x.shape[0]), x.shape[0])
x = x[randomIndices]
y = y[randomIndices]
for i in range(0, x.shape[0], self.batchSize):
Xbatch = x[i:i+self.batchSize]
ybatch = y[i:i+self.batchSize]
loss, dw = self.computeLoss(Xbatch, ybatch)
self.velocity = (self.momentum * self.velocity) + (self.learningRate * dw)
self.wt -= self.velocity
losses.append(loss)
return np.sum(losses) / len(losses)
def softmaxEquation(self, z):
z -= np.max(z)
prob = (np.exp(z).T / np.sum(np.exp(z), axis=1)).T
return prob
def computeLoss(self, x, yMatrix):
numOfSamples = x.shape[0]
scores = np.dot(x, self.wt)
prob = self.softmaxEquation(scores)
loss = -np.log(np.max(prob)) * yMatrix
regLoss = (1/2)*self.regStrength*np.sum(self.wt*self.wt)
totalLoss = (np.sum(loss) / numOfSamples) + regLoss
grad = ((-1 / numOfSamples) * np.dot(x.T, (yMatrix - prob))) + (self.regStrength * self.wt)
return totalLoss, grad
def meanAccuracy(self, x, y):
predY = self.predict(x)
predY = predY.reshape((-1, 1)) # convert to column vector
return np.mean(np.equal(y, predY))
def predict(self, x):
return np.argmax(x.dot(self.wt), 1)
def oneHotEncoding(self, y, numOfClasses):
y = np.asarray(y, dtype='int32')
if len(y) > 1:
y = y.reshape(-1)
if not numOfClasses:
numOfClasses = np.max(y) + 1
yMatrix = np.zeros((len(y), numOfClasses))
yMatrix[np.arange(len(y)), y] = 1
return yMatrix
def plotGraph(trainLosses, testLosses, trainAcc, testAcc):
plt.figure(figsize=(16,6))
plt.subplot(1, 2, 1)
plt.plot(trainLosses, label="Train loss")
plt.plot(testLosses, label="Test loss")
plt.legend(loc='best')
plt.title("Epochs vs. Cross Entropy Loss")
plt.xlabel("Number of Iteration or Epochs")
plt.ylabel("Cross Entropy Loss")
plt.subplot(1, 2, 2)
plt.plot(trainAcc, label="Train Accuracy")
plt.plot(testAcc, label="Test Accuracy")
plt.legend(loc='best')
plt.title("Epochs vs. Mean per class Accuracy")
plt.xlabel("Number of Iteration or Epochs")
plt.ylabel("Mean per class Accuracy")
plt.show()
def readData(filename):
dataMatrix = np.loadtxt(filename)
np.random.shuffle(dataMatrix)
X = dataMatrix[:, 1:]
y = dataMatrix[:, 0].astype(int)
y = y.reshape((-1, 1))
y -= 1
return X, y
def makeMeshGrid(x, y, h=0.02):
x_min, x_max = x[:, 0].min() - 1, x[:, 0].max() + 1
y_min, y_max = x[:, 1].min() - 1, x[:, 1].max() + 1
xx, yy = np.meshgrid(np.arange(x_min, x_max, h),
np.arange(y_min, y_max, h))
return xx, yy
def plotContours(plt, model, xx, yy, **params):
arr = np.array([xx.ravel(), yy.ravel()])
scores = np.dot(arr.T, sm.wt)
prob = model.softmaxEquation(scores)
Z = np.argmax(prob, axis=1) + 1
Z = Z.reshape(xx.shape)
plt.contourf(xx, yy, Z, **params)
def plotDecisionBoundary(x, y):
plt.figure(figsize=(16,7))
markers = ('+', '.', 'x')
colors = ('#8B0000', '#000000', '#191970')
cmap = ListedColormap(colors[:len(np.unique(y))])
xx, yy = makeMeshGrid(x, y)
plotContours(plt, sm, xx, yy, cmap=plt.cm.viridis, alpha=0.8)
# Plot also the training points
for idx, cl in enumerate(np.unique(y)):
xBasedOnLabel = x[np.where(y[:,0] == cl)]
plt.scatter(x=xBasedOnLabel[:, 0], y=xBasedOnLabel[:, 1], c=cmap(idx),
cmap=plt.cm.coolwarm, marker=markers[idx], label=cl)
plt.xlim(xx.min(), xx.max())
plt.ylim(yy.min(), yy.max())
plt.xlabel("Feature X1")
plt.ylabel("Feature X2")
plt.title("Softmax Classifier on Iris Dataset(Decision Boundary)")
plt.xticks()
plt.yticks()
plt.legend(loc='upper left')
plt.show()
trainX, trainY = readData(TRAIN_FILENAME) # Training data
testX, testY = readData(TEST_FILENAME) # Testing data
parser = argparse.ArgumentParser()
parser.add_argument("-e", "--epochs", dest="epochs", default=1000,
type=int, help="Number of epochs")
parser.add_argument("-lr", "--learningrate", dest="learningRate", default=0.07,
type=float, help="Learning rate or step size")
parser.add_argument("-bs", "--batchSize", dest="batchSize", default=10,
type=int, help="Number of sample in mini-batches")
parser.add_argument("-r", "--regStrength", dest="regStrength", default=0.001,
type=float, help="L2 weight decay regularization lambda value")
parser.add_argument("-m", "--momentum", dest="momentum", default=0.05,
type=float, help="A momentum value")
args, unknown = parser.parse_known_args()
print(
"Epochs: {} | Learning Rate: {} | Batch Size: {} | Regularization Strength: {} | "
"Momentum: {} |".format(
args.epochs,
args.learningRate,
args.batchSize,
args.regStrength,
args.momentum
))
epochs = int(args.epochs)
learningRate = float(args.learningRate)
batchSize = int(args.batchSize)
regStrength = int(args.regStrength)
momentum = int(args.momentum)
sm = Softmax(epochs=epochs, learningRate=learningRate, batchSize=batchSize,
regStrength=regStrength, momentum=momentum)
trainLosses, testLosses, trainAcc, testAcc = sm.train(trainX, trainY, testX, testY) # Train a network
plotGraph(trainLosses, testLosses, trainAcc, testAcc)
plotDecisionBoundary(trainX, trainY)
plotDecisionBoundary(testX, testY)
```
| github_jupyter |
```
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
%load_ext autoreload
%autoreload 2
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.pylab as pylab
from simplified_monorotor import Monorotor
import plotting
import testing
import trajectories
pylab.rcParams['figure.figsize'] = 10,10
```
# Feedforward control
The feedforward term captures the acceleration of the target path and adds to the $\bar{u}_1$ calculated by PD controller.
$$
\ddot{z} = \bar{u}_1= k_p(z_t-z)+k_d(z_t-z)+\ddot{z}
$$
#### TODO - Implement PD controller WITH feed forward acceleration
Modify `thrust_control` to incorporate the feedforward term into the PD Controller math.
$$
\begin{align}
e &= z_{\text{target}} - z_{\text{actual}} \\
\dot{e} &= \dot{z}_{\text{target}} - \dot{z}_{\text{actual}} \\
\bar{u}_1 &= k_p e + k_d \dot{e} + \ddot{z}_{\text{ff}} \\
u_1 &= m(g - \bar{u}_1)
\end{align}
$$
```
class PDController:
def __init__(self, k_p, k_d, m):
self.k_p = k_p
self.k_d = k_d
self.vehicle_mass = m
self.g = 9.81
def thrust_control(self,
z_target,
z_actual,
z_dot_target,
z_dot_actual,
z_dot_dot_ff=0.0):
#
# TODO
# modify the PD control code to incorporate
# the feedforward term.
err = z_target - z_actual
err_dot = z_dot_target - z_dot_actual
u_bar = self.k_p * err + self.k_d * err_dot + z_dot_dot_ff
u = self.vehicle_mass * (self.g - u_bar)
return u
testing.pd_controller_test(PDController, feed_forward=True)
```
#### TODO 2 - Compare trajectories with and without a feedforward term
The code below generates plots of $z$ vs $t$ for two drones. One uses FF and the other doesn't.
Run the code and compare the two trajectories. What happens if you increase the oscillation frequency to 10? What happens if you decrease it to 2?
You should notice a **lag** in the system response: the trajectory without the feedforward term should lag behind the desired trajectory in time. This effect diminishes as the oscillation frequency decreases.
```
# This code simulates TWO drones. One uses the feed forward
# acceleration and the other doesn't. Note the difference in
# trajectories.
MASS_ERROR = 1.0
K_P = 20.0
K_D = 8.0
AMPLITUDE = 0.5
OSCILLATION_FREQUENCY = 10
PERIOD = 2 * np.pi / OSCILLATION_FREQUENCY
# preparation (TWO drones to compare)
drone = Monorotor()
ff_drone = Monorotor()
perceived_mass = drone.m * MASS_ERROR
# instantiate TWO controllers
controller = PDController(K_P, K_D, perceived_mass)
ff_controller = PDController(K_P, K_D, perceived_mass)
# get trajectories
t, z_path, z_dot_path, z_dot_dot_path = trajectories.cosine(AMPLITUDE,
PERIOD,
duration=6.0)
dt = t[1] - t[0]
# run simulation
history = []
ff_history = []
for z_target, z_dot_target, z_dot_dot_ff in zip(z_path,
z_dot_path,
z_dot_dot_path):
z_actual = drone.z
z_dot_actual = drone.z_dot
ff_z_actual = ff_drone.z
ff_z_dot_actual = ff_drone.z_dot
u_ff = controller.thrust_control(z_target, ff_z_actual,
z_dot_target, ff_z_dot_actual,
z_dot_dot_ff)
u = controller.thrust_control(z_target, z_actual,
z_dot_target, z_dot_actual)
drone.thrust = u
ff_drone.thrust = u_ff
drone.advance_state(dt)
ff_drone.advance_state(dt)
history.append(drone.X)
ff_history.append(ff_drone.X)
# generate plots
z_actual = [h[0] for h in history]
z_ff_actual = [h[0] for h in ff_history]
plotting.compare_planned_to_actual(z_actual, z_path, t,
z_ff_actual)
```
[Solution](/notebooks/PD%20with%20FF%20Solution.ipynb)
| github_jupyter |
```
import numpy as np
import seaborn as sns
import pandas as pd
import pickle
import os
from sklearn.feature_extraction.text import TfidfVectorizer
from nltk.stem.snowball import SnowballStemmer
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
class Train_Diagnosis():
'''
This class is actually used to train and update the model and the data that I have
Prediction is going to be done through a function.
Class to predict the ailment based on the textual information provided
on the site
ATTRIBUTES:
data : the processed and lemmatized dataframe loaded from the memory (
contains stemmed_data column and the prompt column)
ailments_dict_keyname/_keyint : dictionaries representing the unique ailments present
in the data set
vectorizer : TfIdf vectorizer used for transforming data
( based on the latest state of the data file)
model :
'''
def __init__(self):
self.data = self.get_latest_data()
self.ailments_dict_keyname = self.get_ailments(0)
self.ailments_dict_keyint = self.get_ailments(1)
self.vectorizer = self.get_vectorizer()
self.model = self.get_model()
self.vector_path = 'models/vectorizer.pkl'
self.model_path = 'models/model.pkl'
def get_latest_data(self):
'''Function to load in the latest dataframe
that you have for the model training '''
data = pd.read_csv(r'data/trial_data.csv')
return data
def get_ailments(self,type_of_dict):
'''Function to load the unique ailment dictionary
PARAMETERS: type_of_dict: 0,1 : how to form the
keys of the dictionary
RETURNS : dictionary of the ailments'''
D = {}
ailments = self.data['Prompt'].unique()
if(type_of_dict == 0):
# By name
for i,k in enumerate(ailments):
D[k] = i
else:
# By indexing
for i,k in enumerate(ailments):
D[i] = k
return D
def get_vectorizer(self):
'''Return a vectorizer to fit on the data '''
TfIdf = TfidfVectorizer(stop_words = 'english', ngram_range= (1,3),max_df= 0.7)
return TfIdf
def get_training_x(self):
'''Returns the transformed data for training'''
X = (self.vectorizer).fit_transform(self.data['stemmed_phrase'])
X = X.toarray()
# update the vectorizer here
self.vectorizer = self.vectorizer.fit(self.data['stemmed_phrase'])
# save the vectorizer at this point, after you have fit it
pickle.dump(self.vectorizer,open(self.vector_path,'wb'))
return X
def get_training_y(self):
'''Returns the encoded classes for training'''
Y = self.data['Prompt'].map(self.ailments_dict_keyname)
return Y
def get_model(self):
'''Returns a model for the data '''
M = RandomForestClassifier(n_estimators=36,min_samples_leaf=2)
return M
# METHODS
# 1. Trains the model
def train_model(self):
'''Trains the model as and when you want
with the loaded data'''
X = self.get_training_x()
Y = self.get_training_y()
# validation is actually done on the query not the test data
self.model.fit(X,Y)
pickle.dump(self.model,open(self.model_path,'wb'))
# that's it
class Predictions():
'''class to make the predictions given the model and then
append the query to the data set that you currently have '''
def __init__(self,model,data_path):
self.model = model
self.data = pd.read_csv(data_path)
self.stemmer = SnowballStemmer('english')
punctuation='["\'?,\.]'
self.abbr_dict = {
"what's":"what is",
"what're":"what are",
"where's":"where is",
"where're":"where are",
"i'm":"i am",
"we're":"we are",
"it's":"it is",
"that's":"that is",
"there's":"there is",
"there're":"there are",
"i've":"i have",
"who've":"who have",
"would've":"would have",
"not've":"not have",
"i'll":"i will",
"it'll":"it will",
"isn't":"is not",
"wasn't":"was not",
"aren't":"are not",
"weren't":"were not",
"can't":"can not",
"couldn't":"could not",
"don't":"do not",
"didn't":"did not",
"shouldn't":"should not",
"wouldn't":"would not",
"doesn't":"does not",
"haven't":"have not",
"hasn't":"has not",
"hadn't":"had not",
"won't":"will not",
punctuation:'',
'\s+':' ', # replace multi space with one single space
}
def process_query(self,query):
'''Returns a processed and stemmed query'''
query = query.lower()
res = ''
for k in query.split():
if k in self.abbr_dict:
res+=' ' + self.abbr_dict[k]
else:
res+=' ' + k
res = ' '.join([self.stemmer.stem(y) for y in res.split()])
return res
def append_query(self,query,ailment):
'''Take the query and prediction and then append it to original data '''
col1 = 'stemmed_phrase'
col2 = 'Phrase'
self.data.append([{col1 : query , col2 : ailment}] , ignore_index = True)
```
- Note that the whole dataset does not need to be stemmed again and again
- I will store the stemmed data and then as and when I get a query I would append the stemmed query at the end of the data set after each prediction
```
trainer = Train_Diagnosis()
trainer.train_model()
# get predictions needs to be a separate function as it just needs to get the
# predictions
from flask import Flask, request, jsonify
from flask_cors import CORS
## 1. prediction
## moreover, model and vectorizer need not be loaded again and again
# build 1 end point
app = Flask(__name__)
CORS(app)
def load_from_pickle(file):
loaded = pickle.load(open(file,'rb'))
return loaded
# got the models
vectorizer = load_from_pickle('models/vectorizer.pkl')
model = load_from_pickle('models/model.pkl')
trainer = Train_Diagnosis()
ailments = trainer.get_ailments(1)
diagnoser = Predictions(model,'data/trial_data.csv')
# parameter 1 ->
# processes the query given by the site and make prediction
@app.route('/process',methods = ['GET'])
def get_diagnosis():
q = request.args.get('query')
processed = diagnoser.process_query(q)
# now transform
query = [processed]
query = vectorizer.transform(query)
# and predict
preds = model.predict_proba(query)
res = list(np.argsort(preds))[0]
res = res[::-1][:3] # top 3
ailment_top = ailments[res[0]]
# append record to the data
diagnoser.append_query(query,ailment_top)
#gather predictions
predictions = []
for k in res:
predictions.append(ailments[k])
# parameter point 2 ->
# re-trains the model on the acquired data and
# reloads model and vectorizer
# 0 -> do not train
# 1 -> train again
train = int(request.args.get('train')) # ->
if(train is not None and train == 1):
# means I need to train along with this query
trainer.train_model()
#at this point load the vectorizer and model again with the new queries
vectorizer = load_from_pickle('models/vectorizer.pkl')
model = load_from_pickle('models/model.pkl')
return jsonify(predictions)
if __name__=='__main__':
app.run(port = 5000, debug = True)
```
| github_jupyter |
## Initial Data Analysis of the Unlabeled Tweet Data
This analysis looks at summary statistics and high-level descriptives within the sample of tweets scraped from Fortune 100 companies and associated CEO Twitter accounts +/- 3 weeks around May 25th, 2020. I first look at descriptive statistics, turning special attention to the underlying generating process of the data (how companies tweet) and some high level characteristics of highly liked and retweeted content.
I then turn to link the tweet data to data from Compustat, which contains CEO, company, and industry features. I continue with similar high level analyses of descriptive statistics, providing initial clues into the tweeting behavior of individual companies. Portions of this descriptive analysis approach and code are inspired from Aurelion Gerón's [<u>Hands on Machine Learning, 2nd Edition</u>](https://github.com/ageron/handson-ml2).
Future analyses will delve into more detailed classification, text, and regression analyses. Specifically, following the analysis in this notebook, I will proceed in two parallel steps.
1. I will hire two Upwork data entry specialists to provide labels for each tweet instance. They will input two binary (0/1) features from this data:
* Initial stance on the George Floyd incident
* "Follow-up" action (e.g., giving money to equity non-profits, highlighting black workers in the org, etc.)
<br><br>
2. Perform text cleaning and more rigorous text descriptives: I'll begin by cleaning tweet data in order to tokenize it for future analyses. Using patterns from this descriptive statistics analysis, I will create word visualizations on the unlabeled data to provide initial insights into language patterns in the tweet sample. This analysis will be found in "X.X-pjr-2-cleaning-word-viz".
```
import sys
assert sys.version_info >= (3, 5)
import pandas as pd
import numpy as np
import datetime as dt
# Import Matplotlib and Seaborn
%matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rc('axes', labelsize=14)
mpl.rc('xtick', labelsize=12)
mpl.rc('ytick', labelsize=12)
import seaborn as sns
#Import helpful user-defined functions for directories in nb_functions.py
from nb_functions import set_project_root, change_dir
try:
PROJECT_ROOT
except:
PROJECT_ROOT = set_project_root()
```
### 1. High level descriptives of the tweet data
```
# Change the directory to download the data and return to the notebooks folder.
change_dir(PROJECT_ROOT, '/data')
df = pd.read_csv('tweets_df_raw.csv')
df.head()
df = df.drop(df.columns[0], 1) # Delete the "Unnamed" column, which is a duplicate of the index
df.head()
# Strip date and time information from original "date" feature
df['date_'] = df.date.str[:10]
df['user_join_date_'] = df.user_join_date.str[:10]
df['utc_time'] = df.date.str[11:19]
df['user_join_date_'].head()
# Convert date and time to DateTime types
df['date_'] = pd.to_datetime(df['date_'])
df['event_date'] = '2020-05-25'
df['event_date'] = pd.to_datetime(df['event_date'])
df['user_join_date_'] = pd.to_datetime(df['user_join_date_'])
df['date_'].fillna(0)
df['event_date'].fillna(0)
df['user_join_date'].fillna(0)
df['utc_time'] = pd.to_datetime(df['utc_time']).dt.strftime('%H%M%S')
# Drop original date column, rename "date_" column as "date"
df = df.drop(columns=['date', 'user_join_date'])
df = df.rename(columns={'date_': 'date', 'user_join_date_': 'user_join_date'})
df.info()
df['user_days_on_twitter'] = (df['event_date'] - df['user_join_date']).dt.days
df['days_from_event'] = (df['date'] - df['event_date']).dt.days
df.to_pickle('df_1_a.pkl', protocol=4)
#
# #### PICKLED HERE FOR SUMMARY
#
df.head()
df.describe(include='all', datetime_is_numeric=True)
# How many CEOs are there? How much do they tweet?
df['username'][df['ceo_account']==1].value_counts()
# Where are the tweets coming from?
source_label = df['source_label'].value_counts()
source_label
%matplotlib inline
import matplotlib.pyplot as plt
df.hist(bins=10, figsize=(20,15))
plt.show()
```
<br>
<br>
It's tough to visualize a few of these histograms (likes, quotes, replies, retweets) due to outliers at the right end of the distribution.
<br>
<br>
As examples, looking at the value counts for the number of likes on a tweet and a histogram of tweet replies for tweets with less than 20 replies, one can see that the vast majority of tweets have zero or one likes/replies. So it appears that the data is very right skewed here. More formally, we can use scipy's skewtest, which confirms that these four features have a larger skewness than a user's cumulative number of favorites and user's number of friends.
<br>
<br>
```
likes_val_ct = df['likes'].value_counts()
likes_val_ct
df_replies_cut = df[df['replies'] < 20]
df_replies_cut['replies'].hist(bins = 10, figsize=(20,15))
plt.show()
from scipy import stats
stats.skewtest(df[['likes', 'quotes', 'replies', 'retweets', 'user_favorites', 'user_friends']])
```
<br>
<br>
There are a few things worth noting from the table of summary statistics, histograms, and the data generating process. I look at each of these items in detail in the sections that follow.
<br>
<br>
1. There are notable outliers on many of the individual tweet-level features (e.g., replies, retweets), especially on the high end. These features have right-skewed distributions, with one (or more) accounts being prolific tweeters.
<br>
<br>
2. This sample has both tweets from company accounts (e.g., @Google) and associated CEO accounts (@sundarpichai). Do CEO tweets look different from company tweets? It's not hard to imagine that firms would use these two types of Twitter accounts for different purposes.
<br>
<br>
3. The large majority of tweets in the dataset are sourced through customer experience management (CXM) platforms such as Sprinklr and Conversocial (See below). Many of these tweets are one-on-one responses to customer inquiries. While the data only contains tweets in a 3 week window before and after May 25, 2020, the percentage of tweets that originate outside strictly Twitter sources (e.g., Twitter Web App) could be seen as a rough proxy for the intensity of a company's interaction with individual consumers on Twitter. One could imagine this factor could have implications for a company's propensity to take a stance/follow-through on the stance. For example, it is reasonable to hypothesize that companies that have greater interaction with individual customers on Twitter may be more likely to find stance-taking on political issues worthwhile.
<br>
<br>
#### 1.1. What types of tweets are in the right tail of likes and retweets?
Let's start by taking a subsample of the 20-30 tweets with the most likes and most replies.
<br>
<br>
Below, we see that a reasonable, round-number cutoff for this cursory descriptive analysis would be <b>likes > 5000 and replies > 500 </b>
<br>
<br>
```
likes_val_ct = df['likes'].value_counts(bins=10)
likes_val_ct
replies_val_ct = df['replies'][df['replies'] > 10].value_counts(bins=10)
replies_val_ct
```
<br>
<br>
How do these small samples compare with the full sample?
<br>
<br>
Including both the categorical and numerical data types, we can see a few interesting patterns:
<br>
<br>
(1) The most prolific Twitter user in the full sample is American Airlines, compared to Facebook in the "most replied" subsample and Apple CEO Tim Cook in the "most liked" subsample. The most common source label is a customer experience management platform (Sprinklr) in the full sample and is the Twitter Web App in the subsamples, which likely means that our subsamples filter out a good deal of the company's customer service reply tweets.
<br>
<br>
(2) The mean user attached to each tweet in the subsample has more followers, fewer friends, and fewer statuses than those in the full sample. Combined with patterns mentioned in (1), one might begin to imagine that a greater proportion of these subsample tweets are coming from publicity-friendly CEOs, who tend to engage with Twitter less often than the average company account but maintain a strong presence among their followers.
<br>
<br>
```
df.describe(include='all')
df[df['replies'] > 500].describe(include='all', datetime_is_numeric=True)
df[df['likes'] > 5000].describe(include='all', datetime_is_numeric=True)
```
<br>
<br>
Now let's look at the days from the incident (where negative days indicate days prior to May 25) and the associated text of these tweets.
Although the output only contains surface level evidence, one can see strong representation of words that are associated with taking a stance on the event (e.g., "Minneapolis", "race", "using our platform"). This warrants a more rigorous text analysis with the data.
<br>
<br>
```
# Likes > 5000
df_indiv_tweets = df[['date','days_from_event', 'text', 'replies', 'retweets', 'likes', 'quotes', 'source_label', 'compustat_company']]
df_indiv_tweets[['days_from_event','text']][df_indiv_tweets['likes']>5000]
# Replies > 500
df_indiv_tweets[['days_from_event','text']][df_indiv_tweets['replies']>500]
```
<br>
<br>
We can also take a look at kernel density distributions of these subsamples of tweets compared to the full sample.
Compared to the full sample and a less constrained cut of the data (1000 most liked/replied tweets), the subsamples have a notable center of mass at the event date and just right of the event date. This is especially true for tweets with many replies. The tweets with the most replies peak around a week after the event.
Coupled with the cursory analysis above, it appears that political stance tweets may attract significant attention relative to the average corporate tweet.
<br>
<br>
```
# Full Sample of Tweets (n~45,000), >100 Likes (n~1000), >5000 Likes (n~30)
ax = df_indiv_tweets['days_from_event'].plot.kde(ind=np.arange(-20, 20, 1).tolist(), legend=True)
l_100 = df_indiv_tweets['days_from_event'][df_indiv_tweets['likes']>100].plot.kde(ind=np.arange(-20, 20, 1).tolist(), ax=ax, legend=True)
l_5000 = df_indiv_tweets['days_from_event'][df_indiv_tweets['likes']>5000].plot.kde(ind=np.arange(-20, 20, 1).tolist(), ax=ax, legend=True)
ax.set_xlabel("Days Since Floyd Incident")
ax.legend(['full sample', 'likes>100', 'likes>5000'])
ax.set_title('Density of Tweets with Different Amounts of Likes')
# Full Sample of Tweets (n~45,000), >5 Replies (n~1000), >500 Replies (n~30)
fig = plt.figure()
ax = df_indiv_tweets['days_from_event'].plot.kde(ind=np.arange(-20, 20, 1).tolist(), legend=True)
l_100 = df_indiv_tweets['days_from_event'][df_indiv_tweets['replies']>5].plot.kde(ind=np.arange(-20, 20, 1).tolist(), ax=ax, legend=True)
l_5000 = df_indiv_tweets['days_from_event'][df_indiv_tweets['replies']>500].plot.kde(ind=np.arange(-20, 20, 1).tolist(), ax=ax, legend=True)
ax.set_xlabel("Days Since Floyd Incident")
ax.legend(['full sample', 'replies>5', 'replies>500'])
fig.text(.5, -.1, 'Note: The samples for these graphs are n~45,000, n~1,000, and n~25, respectively', ha='center')
plt.show()
```
#### 1.2 How do CEO tweets differ from company tweets?
Let's start by looking at basic summary statistics broken down by account type
<br>
<br>
```
with pd.option_context('display.max_columns', 40):
print(df[df['ceo_account']==1].describe(include='all', datetime_is_numeric=True))
with pd.option_context('display.max_columns', 40):
print(df[df['ceo_account']==0].describe(include='all', datetime_is_numeric=True))
```
Each marginal tweet from a CEO account appears to be much more influential than corporate accounts if we consider mean numbers of likes, replies, retweets, and quotes. Corporate accounts, in contrast, have larger mean values for the user features (e.g., followers, friends), which is understandable since these accounts tend to be established and long-lived, as opposed to the average CEO who might use Twitter most intensely in their corporate role but lower their activity after leaving the CEO role.
CEOs tend to source their tweets through the Twitter Web App, while companies tend to source them through customer management experience (CXM) platforms, strongly suggesting that CEOs are using the platform to broadcast content to a wide audience and company accounts are mostly being used to address individual customer service inquiries.
The most prolific tweeter in our sample is Dell CEO Michael Dell, with 45 tweets over the six-week study period. Importantly, large amounts of tweets do not seem to translate into highly liked tweets, given that we found Apple CEO Tim Cook to have the most tweets with more than 5,000 likes in our sample. Twitter itself is just one outlet in the social world, so it can both influence social factors in the outside world as well as be shaped by them.
<br>
<br>
Looking slightly more deeply at the data, we can see that the majority of the most liked tweets (>5,000 likes) come from CEO accounts with the majority coming from only three CEOs: Alphabet CEO Sundar Pichai, Apple CEO Tim Cook, and Disney CEO Bob Iger. Interestingly, Tim Cook only tweeted 15 times during this period, and the majority (9) of his tweets received over 5,000 likes.
If we turn to tweets with the most number of replies (>500), however, the pattern of relatively greater CEO influence is less clear. The large majority of the tweets that generate the largest number of replies come from corporate accounts. It's worth noting that corporate accounts tend to have larger follower and friend bases than CEOs, which may account for the greater number of replies. Public twitter account users are also able to limit who can reply to tweets (everyone, friends, no replies), which may also inflate corporate reply numbers given the larger relative friend base. Finally, it's worth noting that the two CEO tweets that have more than 500 replies is proportionally greater than their relative presence in the total sample, in which CEOs are responsible for only 0.5% of all tweets. Being less restrictive in defining "highly replied" tweets (<400) maintains this relatively high CEO presence compared to the full sample.
In sum, looking at either the most liked or most replied tweets suggests an outsized, influential role for CEO tweets in this sample.
<br>
<br>
```
df[['text', 'username', 'likes', 'date']][df['likes']>5000]
df[['text', 'username', 'likes', 'date']][df['replies']>500]
```
#### 1.3. How do tweets from non-Twitter sources compare to those from Twitter sources?
As previously noted, most of the tweets in this sample are sourced through outside customer experience management (CXM) platforms. Many of these CXM tweets are one-on-one discussions with individual customers, usually about some sort of customer service issue. Let's examine whether there are differences in the tweets sourced through these outlets versus those sourced directly through Twitter.
<br>
<br>
Before going any further, it's worth noting that not <b>all</b> of these CXM tweets are simply customer service issues. As an example related to the previous section, many of the most liked tweets in this time period were sourced through CXM platforms. A few of these, like Amazon's tweet with a URL in the text field, are clear public political stances.
<br>
<br>
```
df_indiv_tweets[['days_from_event','text','source_label', 'compustat_company']][df_indiv_tweets['likes']>5000]
df['twitter_source'] = (df.source_label.str[:7] == "Twitter")
df[['source_label', 'twitter_source']]
```
<br>
<br>
Looking at summary statistics of the features with numerical data types, we can see that the average Twitter-sourced tweet has, on average, more replies, retweets, likes, and quotes than the average tweet from non-Twitter sources. This provides supporting evidence behind the idea that companies may tend to outsource Twitter engagement that might not be of interest to the general public.
<br>
<br>
```
df[df['twitter_source']==1].describe()
df[df['twitter_source']==0].describe()
```
### 2. Merging Data from Compustat
The tweet data can only tell us so much about the landscape of companies and CEOs tweeting public political stances. To provide further individual, organizational, and industry details on the account users and tweeting trends, I merge data from Compustat's Execucomp database, which contains data on individual CEOs (e.g., tenure at company), companies, and industries.
To simplify this analysis, I will match each tweet with the company's data from the year 2020. It's worth noting that some Fortune 100 companies do not appear in the dataset. Execucomp matches publicly traded companies by their stock ticker, so companies can fall out fo the dataset for reasons as diverse as not having a single primary stock ticker (e.g., Berkshire Hathaway) or not being publicly traded during the entire year (e.g., Albertsons). There are 80 Twitter-active company accounts and 12 Twitter-active CEOs in this dataset.
Merging will be done on the "compustat_company" feature in order to retain the baseline dataset of tweets. This is a one-to-many join, where we are matching one instance of Compustat data per company to many tweet instances from the same company (or the associated company's CEO).
<br>
<br>
```
# Join the Compustat dataset based on the "compustat_company" key
compustat_df = pd.read_csv('compustat_data.csv')
compustat_df.info()
# Delete rows from compustat_df that aren't year == 2019, merge into tweets df.
compustat_df = compustat_df[compustat_df['YEAR']==2019]
compustat_df = compustat_df[compustat_df['CEOANN']=='CEO']
compustat_df.head()
#Keep the year value from BECAMECEO feature
compustat_df['BECAMECEO'] = compustat_df['BECAMECEO'] // 10000
df_merged = pd.merge(df, compustat_df, how='inner', on=['compustat_company'])
df_merged.head()
# Drop data that is irrelevant and/or very sparse (e.g., GVKEY: compustat CEO identifier, JOINED_CEO: <10% of obs available)
df_merged.drop(columns=['GVKEY', 'LEFTOFC', 'CO_PER_ROL', 'EXECRANK', 'PCEO', 'JOINED_CO', 'CEOANN', 'YEAR', 'TITLE', 'TICKER'], inplace=True)
df_merged.info()
```
<br><br>
Out of the 84 companies in our sample, we see that the mean CEO age is 57 years old, roughly 90% are male, and have served as CEO on average for 5 years. While only about 15% of CEOs have data on the year they joined their company, it appears that many CEOs are long-lived employees at their firms, with this subsample having originally joined their firm in 1988 on average.
A wide variety of industries are represented in this data. From looking at value counts of tweets per industry, there appears to be a rough trend that many of the most prolific tweeters are B2C companies (e.g., airlines, wireless providers, grocery stores) and some of the least prolific tweeters are B2B companies (e.g., oil and gas drilling, manufacturing for intermediate products). Paired with the descriptive analysis of the CXM sources, it appears that customer-facing companies (especially airlines) use Twitter as a way to interact with individual customers, who may be more likely to post customer service inquiries on Twitter than customers of B2B companies.
It's also worth noting that corporate actions with respect to the COVID-19 pandemic may be leading to especially high numbers of tweets among some accounts. For example, Kroger drew criticism during this time period for their [HR decisions](https://www.cbsnews.com/news/kroger-workers-extra-coronavirus-pay-return/) regarding employee pay and hours during the the pandemic. I have designed the main analysis to deal with these outliers and understand how they may impact results.
<br><br>
```
with pd.option_context('display.max_columns', 40):
print(df_merged.describe(include='all', datetime_is_numeric=True))
df_merged['NAICSDESC'].value_counts()
df['username'][df['twitter_source']!=1].value_counts()
```
<br><br>
#### Create datasets for Upwork
Below, I create two binary features (<i>floyd_stance</i>, which is 1 for the account's initial tweet stance and <i>floyd_followup</i>, which is 1 for any stance related to racial justice, policing reform, etc.). These columns are all set to 0, and Upworkers will change them to 1 per the feature definitions. I plan to match on the *tweet_id* feature, which is unique for each tweet.
I divide the dataset into 6 sub-datasets for Upworkers. I manually manipulate data in Excel to ensure that each account is fully contained within one sub-dataset. Each Upworker will append their finished dataset with either "\_a" or "\_b", and no Upworker can provide two completed versions of a subdataset (i.e., each dataset's two raters will be unique).
<br><br>
```
# df_merged['floyd_stance'] = 0
# df_merged['floyd_followup'] = 0
# df_merged['tweet_id'] = 't' + df_merged['tweet_id'].astype(str)
# df_merged.to_csv('data_merged.csv', index=False)
# df_merged.to_pickle('data_merged.pkl', protocol=4)
# df_upwork = df_merged[['floyd_stance', 'floyd_followup', 'username', 'date', 'text', 'tweet_id']]
# df_upwork = df_upwork.sort_values(['username', 'date'], ascending=[True, True])
# df_upwork.reset_index(drop=True, inplace=True) # We'll match on the tweet_id value for the future merge
# df_len = len(df_upwork)
# df_len
# # Divide up dataset for Upwork. Note that 41,532 is divisible by 6 (digits sum to 15, which is divisible by 3).
# # This leaves roughly 7,000 instances per dataset for Upworkers.
# # Create 6 df slices, divide up work into 6 dfs.
# dfnames = ['df_u_1', 'df_u_2', 'df_u_3', 'df_u_4', 'df_u_5', 'df_u_6']
# df_u_list = [pd.DataFrame for x in dfnames]
# size = int(df_len/6)
# list_of_dfs = [df_upwork.loc[i:i+size-1,:] for i in range(0, len(df),size)]
# df_dict = {dfnames[i]: list_of_dfs[i] for i in range(len(dfnames))}
# #Create and change directory
# change_dir(PROJECT_ROOT, "/data/upwork_rater_data")
# for name, frame in df_dict.items():
# frame.to_csv(name+'.csv', index=False)
# df_upwork.to_csv('df_u_full.csv', index=False)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/Aye-Nyein-Thaw/TensorFlow-Beginner/blob/main/coding-exercise/week5/part1/4_Autosave_with_ModelCheckpoint_callback.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Autosave with ModelCheckpoint callback
```
import tensorflow as tf
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Flatten, Conv2D, MaxPooling2D
# Prepare Data
(x_train, y_train), (x_test, y_test) = tf.keras.datasets.cifar10.load_data()
x_train = x_train / 255.0
x_test = x_test / 255.0
# Use smaller subset to speed things up
x_train = x_train[:10000]
y_train = y_train[:10000]
x_test = x_test[:10000]
y_test = y_test[:10000]
# Define helper functions
def evaluate(model, x_test, y_test):
test_loss, test_acc = model.evaluate(x=x_test, y=y_test, verbose=0)
print('accuracy = {acc:0.2f}%, Loss = {loss:0.2f}'.format(acc = test_acc * 100, loss = test_loss))
def get_new_model():
model = Sequential([
Conv2D(filters=16, input_shape=(32, 32, 3), kernel_size=(3, 3),
activation='relu', name='conv_1'),
Conv2D(filters=8, kernel_size=(3, 3), activation='relu', name='conv_2'),
MaxPooling2D(pool_size=(4, 4), name='pool_1'),
Flatten(name='flatten'),
Dense(units=32, activation='relu', name='dense_1'),
Dense(units=10, activation='softmax', name='dense_2')
])
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
return model
```
# Model Checkpoint Callback
```
from tensorflow.keras.callbacks import ModelCheckpoint
# save only weights at every epoch
# overwrites everytime
checkpoint_path = 'model_checkpoints/checkpoint'
checkpoint = ModelCheckpoint(filepath = checkpoint_path,
frequency = 'epoch',
save_weights_only = True,
verbose = 1)
# save only weights at every epoch
# doesn't overwrite, new file everytime
checkpoint_path = 'model_checkpoints/checkpoint_{epoch:02d}'
epoch_checkpoint = ModelCheckpoint(filepath = checkpoint_path,
frequency = 'epoch',
save_weights_only = True,
verbose = 1)
# Save best model
# monitor validation accuracy
checkpoint_path = 'model_checkpoints/best_checkpoint'
best_checkpoint = ModelCheckpoint(filepath = checkpoint_path,
frequency = 'epoch',
save_weights_only = True,
save_best_only = True,
monitor = 'val_accuracy',
verbose = 1)
# Save entire model
# Best model
checkpoint_path = 'model_checkpoints/best_checkpoint_model'
model_best_checkpoint = ModelCheckpoint(filepath = checkpoint_path,
frequency = 'epoch',
save_weights_only = False,
save_best_only = True,
monitor = 'val_accuracy',
verbose = 1)
model = get_new_model()
model.fit(x_train, y_train, epochs = 3, callbacks = [model_best_checkpoint], validation_data = (x_test, y_test))
```
## Compare two models
```
from tensorflow.keras.models import load_model
trained_model = load_model('model_checkpoints/best_checkpoint_model')
new_model = get_new_model()
evaluate(trained_model, x_test, y_test)
evaluate(new_model, x_test, y_test)
```
| github_jupyter |
## Search for the best Parameters
using dataset:
ex6data1.mat - Example Dataset 1
ex6data2.mat - Example Dataset 2
ex6data3.mat - Example Dataset 3
spamTrain.mat - Spam training set
spamTest.mat - Spam test set
emailSample1.txt - Sample email 1
emailSample2.txt - Sample email 2
spamSample1.txt - Sample spam 1
spamSample2.txt - Sample spam 2
vocab.txt - Vocabulary list
In this part,ex6data.mat.
You are given the variables:
X,y
Xval,yval
Your task is to use the cross validation set $Xval$ $yval$ to determine the best $C$ and $\sigma$ parameter
Kernel:$Gaussian Kernels$
### 1.1 Example Dataset 3
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from scipy.io import loadmat
%matplotlib inline
%config InlineBackend.figure_format='svg'
dataSet=loadmat('ex6data3.mat')
%matplotlib inline
%config InlineBackend.figure_format='svg'
def plotData(dataSet):
data_train=pd.DataFrame(dataSet.get('X'),columns=['X1','X2'])
data_train['y']=dataSet.get('y')
data_val=pd.DataFrame(dataSet.get('Xval'),columns=['Xval1','Xval2'])
data_val['yval']=dataSet.get('yval')
plt.figure(figsize=(12,8))
plt.tick_params(direction='in')
positive=data_train[data_train['y'].isin([1])]
negative=data_train[data_train['y'].isin([0])]
plt.scatter(positive['X1'],positive['X2'],c='black',edgecolors='black',marker='+')
plt.scatter(negative['X1'],negative['X2'],c='yellow',edgecolors='black',marker='o')
plotData(dataSet)
def load_mat():
dataSet=loadmat('ex6data3.mat')
data_train=pd.DataFrame(dataSet.get('X'),columns=['X1','X2'])
data_train['y']=dataSet.get('y')
data_val=pd.DataFrame(dataSet.get('Xval'),columns=['Xval1','Xval2'])
data_val['yval']=dataSet.get('yval')
return data_train,data_val
```
### 1.2 Manual grid search for $C$ and $\sigma$
```
from sklearn.svm import SVC
candidate = [0.01, 0.03, 0.1, 0.3, 1, 3, 10, 30, 100]
combination=[(C,gamma) for C in candidate for gamma in candidate]
print("combination:\n{}".format(combination))
data_train,data_val=load_mat()
search=[]
for C,gamma in combination:
svc=SVC(C=C,kernel='rbf',gamma=gamma)
svc.fit(data_train[['X1','X2']],data_train['y'])
search.append(svc.score(data_val[['Xval1','Xval2']],data_val['yval']))
#Search for best param
best_score=search[np.argmax(search)]
best_param=combination[np.argmax(search)]
print("best score:\n{}\nbest param:\n{}".format(best_score,best_param))
best_svc=SVC(C=best_param[1],gamma=best_param[0],kernel='rbf')
best_svc.fit(data_train[['X1','X2']],data_train['y'])
ypred=best_svc.predict(data_val[['Xval1','Xval2']])
#Sklearn accuary
from sklearn.metrics import classification_report
print(classification_report(data_val['yval'],ypred))
#Mannul accuary
accuary=sum([1 if (x==y) else 0 for (x,y) in zip(data_val['yval'],ypred)])
print("accuary:{}".format(accuary/len(ypred)))
def plot_decision_boundary(svc,X):
x_min,x_max=X[:,0].min()*1.2,X[:,0].max()*1.2
y_min,y_max=X[:,1].min()*1.1,X[:,1].max()*1.1
xx,yy=np.meshgrid(np.linspace(x_min,x_max,500),
np.linspace(y_min,y_max,500))
Z=svc.predict(np.c_[xx.ravel(),yy.ravel()])
Z=Z.reshape(xx.shape)
plt.contour(xx, yy, Z,colors='blue')
def plotData(dataSet,svc):
data_train=pd.DataFrame(dataSet.get('X'),columns=['X1','X2'])
data_train['y']=dataSet.get('y')
# data_val=pd.DataFrame(dataSet.get('Xval'),columns=['Xval1','Xval2'])
# data_val['yval']=dataSet.get('yval')
plt.figure(figsize=(12,8))
plt.tick_params(direction='in')
positive=data_train[data_train['y'].isin([1])]
negative=data_train[data_train['y'].isin([0])]
plt.scatter(positive['X1'],positive['X2'],c='black',edgecolors='black',marker='+')
plt.scatter(negative['X1'],negative['X2'],c='yellow',edgecolors='black',marker='o')
plot_decision_boundary(svc,dataSet.get('X'))
plotData(dataSet,best_svc)
```
### 1.3 Sklearn GridSearchCV
```
from sklearn.model_selection import GridSearchCV
paramters={'C':candidate,'gamma':candidate}
svc=SVC(kernel='rbf')
clf=GridSearchCV(svc,paramters,n_jobs=-1)
clf.fit(data_train[['X1','X2']],data_train['y'])
clf.best_params_
clf.best_score_
ypred=clf.predict(data_val[['Xval1','Xval2']])
print(classification_report(ypred,data_val['yval']))
accuary=sum([1 if (x==y) else 0 for (x,y) in zip(ypred,data_val['yval'])])
print("accuary:{}".format(accuary/len(ypred)))
best_svc=SVC(C=clf.best_params_['C'],gamma=clf.best_params_['gamma'])
best_svc.fit(data_train[['X1','X2']],data_train['y'])
plotData(dataSet,best_svc)
```
curiouly... they are not the same result. What?
So the built in sklearn grid search is trying to find the best candidate from **training set**
However, when we were doing manual grid search, we train using training set, but we pick the best from **cross validation set**. This is the reason of difference.
### The reason is:
It turns out that **GridSearch** will appropriate part of data as CV and use it to find the best candidate.
So the reason for different result is just that GridSearch here is just using part of **training data** to train because it need part of data as cv set
| github_jupyter |
# Intro
In this notebook we will learn about the [conditional multivariate normal (MVN) distribution](https://en.wikipedia.org/wiki/Multivariate_normal_distribution). In particular, we want to estimate the expected value (or the mean) of some subset of variables given that another subset has been conditioned on. Though the notation is quasi-dense, it is not terribly difficult to produce a conditional MVN from a marginal MVN distribution.
# Case 1
* $X_0 \rightarrow X_1$
```
import numpy as np
from numpy.random import normal
N = 10000
x0 = normal(0, 1, N)
x1 = normal(1 + 2 * x0, 1, N)
X = np.hstack([x0.reshape(-1, 1), x1.reshape(-1, 1)])
M = np.mean(X, axis=0)
S = np.cov(X.T)
print(X.shape)
print(M.shape)
print(S.shape)
print('mean', M)
print('cov', S)
M[0] + S[0,1] / S[1,1] * (0.5 - M[1])
M[1] + S[1,0] / S[0,0] * (0.5 - M[0])
S[0,0] - S[0,1] / S[1,1] * S[1,0]
S[1,1] - S[1,0] / S[0,0] * S[1,0]
```
# Case 2
* $X_0 \rightarrow X_1 \rightarrow X_2$
```
from collections import namedtuple
from numpy.linalg import inv
import warnings
warnings.filterwarnings('ignore')
COV = namedtuple('COV', 'C11 C12 C21 C22 C22I')
def to_row_indices(indices):
return [[i] for i in indices]
def to_col_indices(indices):
return indices
def get_covariances(i1, i2, S):
r = to_row_indices(i1)
c = to_col_indices(i1)
C11 = S[r,c]
r = to_row_indices(i1)
c = to_col_indices(i2)
C12 = S[r,c]
r = to_row_indices(i2)
c = to_col_indices(i1)
C21 = S[r,c]
r = to_row_indices(i2)
c = to_col_indices(i2)
C22 = S[r,c]
C22I = inv(C22)
return COV(C11, C12, C21, C22, C22I)
def compute_means(a, M, C, i1, i2):
a = np.array([2.0])
return M[i1] + C.C12.dot(C.C22I).dot(a - M[i2])
def compute_covs(C):
return C.C11 - C.C12.dot(C.C22I).dot(C.C21)
def update_mean(m, a, M, i1, i2):
v = np.copy(M)
for i, mu in zip(i1, m):
v[i] = mu
for i, mu in zip(i2, a):
v[i] = mu
return v
def update_cov(c, S, i1, i2):
m = np.copy(S)
rows, cols = c.shape
for row in range(rows):
for col in range(cols):
m[i1[row],i1[col]] = c[row,col]
for i in i2:
m[i,i] = 0.01
return m
def update_mean_cov(v, iv, M, S):
if v is None or iv is None or len(v) == 0 or len(iv) == 0:
return np.copy(M), np.copy(S)
i2 = iv.copy()
i1 = [i for i in range(S.shape[0]) if i not in i2]
C = get_covariances(i1, i2, S)
m = compute_means(v, M, C, i1, i2)
c = compute_covs(C)
M_u = update_mean(m, v, M, i1, i2)
S_u = update_cov(c, S, i1, i2)
return M_u, S_u
N = 10000
x0 = normal(0, 1, N)
x1 = normal(1 + 2 * x0, 1, N)
x2 = normal(1 + 2 * x1, 1, N)
X = np.hstack([x0.reshape(-1, 1), x1.reshape(-1, 1), x2.reshape(-1, 1)])
M = np.mean(X, axis=0)
S = np.cov(X.T)
print('mean', M)
print('>')
print('cov', S)
print('>')
print('corr', np.corrcoef(X.T))
M_u, S_u = update_mean_cov(np.array([2.0]), [1], M, S)
print('mean', M_u)
print('>')
print('cov', S_u)
print('>')
print('corr', np.corrcoef(np.random.multivariate_normal(M_u, S_u, N*10).T))
```
# Case 2
* $X_0 \leftarrow X_1 \rightarrow X_2$
```
N = 10000
x1 = normal(0, 1, N)
x0 = normal(1 + 4.0 * x1, 1, N)
x2 = normal(1 + 2.0 * x1, 1, N)
X = np.hstack([x0.reshape(-1, 1), x1.reshape(-1, 1), x2.reshape(-1, 1)])
M = np.mean(X, axis=0)
S = np.cov(X.T)
print('mean', M)
print('>')
print('cov', S)
print('>')
print('corr', np.corrcoef(X.T))
M_u, S_u = update_mean_cov(np.array([2.0]), [1], M, S)
print('mean', M_u)
print('>')
print('cov', S_u)
print('>')
print('corr', np.corrcoef(np.random.multivariate_normal(M_u, S_u, N*10).T))
```
# Case 2
* $X_0 \rightarrow X_1 \leftarrow X_2$
```
N = 10000
x0 = normal(0, 1, N)
x2 = normal(0, 1, N)
x1 = normal(1 + 2 * x0 + 3 * x2, 1, N)
X = np.hstack([x0.reshape(-1, 1), x1.reshape(-1, 1), x2.reshape(-1, 1)])
M = np.mean(X, axis=0)
S = np.cov(X.T)
print('mean', M)
print('>')
print('cov', S)
print('>')
print('corr', np.corrcoef(X.T))
M_u, S_u = update_mean_cov(np.array([2.0]), [1], M, S)
print('mean', M_u)
print('>')
print('cov', S_u)
print('>')
print('corr', np.corrcoef(np.random.multivariate_normal(M_u, S_u, N*10).T))
```
# Links
* https://stackoverflow.com/questions/22927181/selecting-specific-rows-and-columns-from-numpy-array
* https://docs.scipy.org/doc/numpy/reference/generated/numpy.delete.html
* https://stackoverflow.com/questions/38713746/python-numpy-conditional-simulation-from-a-multivatiate-distribution
```
for m in M_u:
print(m)
```
| github_jupyter |
```
import collections
from datetime import datetime
import functools
import itertools
import os
import pathlib
import requests
import string
import sys
import typing
import zipfile
import IPython
import numpy as np
import pandas as pd
import pypandoc
from tqdm.notebook import tqdm_notebook
module_path = os.path.abspath(os.path.join("../.."))
if module_path not in sys.path:
sys.path.append(module_path)
from data_pipeline.utils import remove_all_from_dir, get_excel_column_name
from data_pipeline.etl.sources.census.etl_utils import get_state_information
# Turn on TQDM for pandas so that we can have progress bars when running `apply`.
tqdm_notebook.pandas()
# Suppress scientific notation in pandas (this shows up for census tract IDs)
pd.options.display.float_format = "{:.2f}".format
# Set some global parameters
DATA_DIR = pathlib.Path.cwd().parent / "data"
TEMP_DATA_DIR = DATA_DIR / "tmp"
COMPARISON_OUTPUTS_DIR = DATA_DIR / "comparison_outputs"
## I (Vincent) created this manually locally. Will need to change potentially when putting into official ETL scripts
GEOCORR_DATA_DIR = DATA_DIR / "geocorr"
# Make the dirs if they don't exist
TEMP_DATA_DIR.mkdir(parents=True, exist_ok=True)
COMPARISON_OUTPUTS_DIR.mkdir(parents=True, exist_ok=True)
CEJST_PRIORITY_COMMUNITY_THRESHOLD = 0.75
# Name fields using variables. (This makes it easy to reference the same fields frequently without using strings
# and introducing the risk of misspelling the field name.)
GEOID_FIELD_NAME = "GEOID10"
GEOID_TRACT_FIELD_NAME = "GEOID10_TRACT"
GEOID_STATE_FIELD_NAME = "GEOID10_STATE"
GEOID_CBG_FIELD_NAME = "GEOID10_CBG"
COUNTRY_FIELD_NAME = "Country"
CENSUS_BLOCK_GROUP_POPULATION_FIELD = "Total population"
CEJST_SCORE_FIELD = "cejst_score"
CEJST_PERCENTILE_FIELD = "cejst_percentile"
CEJST_PRIORITY_COMMUNITY_FIELD = "cejst_priority_community"
# Define some suffixes
POPULATION_SUFFIX = " (priority population)"
```
## Mapping Census Block Group to Urban and Rural Indicators using Geocorr Data
The end result is a dataframe `urban_rural_map`
```
# CSV was manually generated
# Instructions for how to generate the CSV from Geocorr are here: https://github.com/usds/justice40-tool/issues/355#issuecomment-920241787
geocorr_urban_rural_map = pd.read_csv(
os.path.join(GEOCORR_DATA_DIR, "geocorr2014_2125804280.csv"),
encoding="ISO-8859-1",
skiprows=[1],
dtype="str",
)
geocorr_urban_rural_map["pop10"] = pd.to_numeric(
geocorr_urban_rural_map["pop10"]
)
geocorr_urban_rural_map["afact"] = pd.to_numeric(
geocorr_urban_rural_map["afact"]
)
geocorr_urban_rural_map[GEOID_TRACT_FIELD_NAME] = (
geocorr_urban_rural_map["county"] + geocorr_urban_rural_map["tract"]
) # + geocorr_urban_rural_map['bg']
geocorr_urban_rural_map[GEOID_TRACT_FIELD_NAME] = geocorr_urban_rural_map[
GEOID_TRACT_FIELD_NAME
].str.replace(".", "", regex=False)
geocorr_urban_rural_map[GEOID_TRACT_FIELD_NAME].str.len().value_counts()
```
We want to see that the length of the derived Census Block Group is always 12 digits. Census Tracts are always 11 digits
```
geocorr_urban_rural_map = geocorr_urban_rural_map[
[GEOID_TRACT_FIELD_NAME, "ur", "ua", "cntyname", "uaname", "pop10", "afact"]
]
```
Checking Primary Key
```
geocorr_urban_rural_map.groupby(
[GEOID_TRACT_FIELD_NAME, "ur", "ua"], dropna=False
).size().sort_values(ascending=False)
geocorr_urban_rural_map.loc[
geocorr_urban_rural_map[GEOID_TRACT_FIELD_NAME] == "36117020302"
]
total_geo_population = (
geocorr_urban_rural_map.groupby(GEOID_TRACT_FIELD_NAME)
.agg({"pop10": np.sum})
.reset_index()
)
total_geo_population.rename(columns={"pop10": "total_population"}, inplace=True)
total_geo_population.head()
geocorr_urban_rural_with_total_pop_map = (
geocorr_urban_rural_map.groupby([GEOID_TRACT_FIELD_NAME, "ur"])
.agg({"pop10": np.sum})
.reset_index()
)
geocorr_urban_rural_with_total_pop_map = (
geocorr_urban_rural_with_total_pop_map.merge(
total_geo_population, how="inner", on=GEOID_TRACT_FIELD_NAME
)
)
geocorr_urban_rural_with_total_pop_map.head()
geocorr_urban_rural_with_total_pop_map["afact"] = (
geocorr_urban_rural_with_total_pop_map["pop10"]
/ geocorr_urban_rural_with_total_pop_map["total_population"]
)
geocorr_urban_rural_with_total_pop_map.head()
geocorr_urban_rural_with_total_pop_map.loc[
geocorr_urban_rural_with_total_pop_map[GEOID_TRACT_FIELD_NAME]
== "01001020200"
]
urban_rural_map = geocorr_urban_rural_with_total_pop_map.pivot(
index=GEOID_TRACT_FIELD_NAME, columns="ur", values=["pop10", "afact"]
)
urban_rural_map.columns = [
"_".join(col).strip() for col in urban_rural_map.columns.values
]
urban_rural_map.reset_index(inplace=True)
urban_rural_map["urban_heuristic_flag"] = 0
mask = urban_rural_map["afact_U"] >= 0.5
urban_rural_map.loc[mask, "urban_heuristic_flag"] = 1
urban_rural_map.rename(
columns={
"pop10_R": "population_in_rural_areas",
"pop10_U": "population_in_urban_areas",
"afact_R": "perc_population_in_rural_areas",
"afact_U": "perc_population_in_urban_areas",
},
inplace=True,
)
urban_rural_map.head(5)
urban_rural_map.to_csv(
path_or_buf=GEOCORR_DATA_DIR / "urban_rural_map.csv", na_rep="", index=False
)
```
| github_jupyter |
# Writing Keras Models With TensorFlow NumPy
**Author:** lukewood<br>
**Date created:** 2021/08/28<br>
**Last modified:** 2021/08/28<br>
**Description:** Overview of how to use the TensorFlow NumPy API to write Keras models.
## Introduction
[NumPy](https://numpy.org/) is a hugely successful Python linear algebra library.
TensorFlow recently launched [tf_numpy](https://www.tensorflow.org/guide/tf_numpy), a
TensorFlow implementation of a large subset of the NumPy API.
Thanks to `tf_numpy`, you can write Keras layers or models in the NumPy style!
The TensorFlow NumPy API has full integration with the TensorFlow ecosystem.
Features such as automatic differentiation, TensorBoard, Keras model callbacks,
TPU distribution and model exporting are all supported.
Let's run through a few examples.
## Setup
TensorFlow NumPy requires TensorFlow 2.5 or later.
```
import tensorflow as tf
import tensorflow.experimental.numpy as tnp
import keras
import keras.layers as layers
import numpy as np
```
Optionally, you can call `tnp.experimental_enable_numpy_behavior()` to enable type promotion in TensorFlow.
This allows TNP to more closely follow the NumPy standard.
```
tnp.experimental_enable_numpy_behavior()
```
To test our models we will use the Boston housing prices regression dataset.
```
(x_train, y_train), (x_test, y_test) = tf.keras.datasets.boston_housing.load_data(
path="boston_housing.npz", test_split=0.2, seed=113
)
def evaluate_model(model: keras.Model):
[loss, percent_error] = model.evaluate(x_test, y_test, verbose=0)
print("Mean absolute percent error before training: ", percent_error)
model.fit(x_train, y_train, epochs=200, verbose=0)
[loss, percent_error] = model.evaluate(x_test, y_test, verbose=0)
print("Mean absolute percent error after training:", percent_error)
```
## Subclassing keras.Model with TNP
The most flexible way to make use of the Keras API is to subclass the
[`keras.Model`](https://keras.io/api/models/model/) class. Subclassing the Model class
gives you the ability to fully customize what occurs in the training loop. This makes
subclassing Model a popular option for researchers.
In this example, we will implement a `Model` subclass that performs regression over the
boston housing dataset using the TNP API. Note that differentiation and gradient
descent is handled automatically when using the TNP API alongside keras.
First let's define a simple `TNPForwardFeedRegressionNetwork` class.
```
class TNPForwardFeedRegressionNetwork(keras.Model):
def __init__(self, blocks=None, **kwargs):
super(TNPForwardFeedRegressionNetwork, self).__init__(**kwargs)
if not isinstance(blocks, list):
raise ValueError(f"blocks must be a list, got blocks={blocks}")
self.blocks = blocks
self.block_weights = None
self.biases = None
def build(self, input_shape):
current_shape = input_shape[1]
self.block_weights = []
self.biases = []
for i, block in enumerate(self.blocks):
self.block_weights.append(
self.add_weight(
shape=(current_shape, block), trainable=True, name=f"block-{i}"
)
)
self.biases.append(
self.add_weight(shape=(block,), trainable=True, name=f"bias-{i}")
)
current_shape = block
self.linear_layer = self.add_weight(
shape=(current_shape, 1), name="linear_projector", trainable=True
)
def call(self, inputs):
activations = inputs
for w, b in zip(self.block_weights, self.biases):
activations = tnp.matmul(activations, w) + b
# ReLu activation function
activations = tnp.maximum(activations, 0.0)
return tnp.matmul(activations, self.linear_layer)
```
Just like with any other Keras model we can utilize any supported optimizer, loss,
metrics or callbacks that we want.
Let's see how the model performs!
```
model = TNPForwardFeedRegressionNetwork(blocks=[3, 3])
model.compile(
optimizer="adam",
loss="mean_squared_error",
metrics=[keras.metrics.MeanAbsolutePercentageError()],
)
evaluate_model(model)
```
Great! Our model seems to be effectively learning to solve the problem at hand.
We can also write our own custom loss function using TNP.
```
def tnp_mse(y_true, y_pred):
return tnp.mean(tnp.square(y_true - y_pred), axis=0)
keras.backend.clear_session()
model = TNPForwardFeedRegressionNetwork(blocks=[3, 3])
model.compile(
optimizer="adam",
loss=tnp_mse,
metrics=[keras.metrics.MeanAbsolutePercentageError()],
)
evaluate_model(model)
```
## Implementing a Keras Layer Based Model with TNP
If desired, TNP can also be used in layer oriented Keras code structure. Let's
implement the same model, but using a layered approach!
```
def tnp_relu(x):
return tnp.maximum(x, 0)
class TNPDense(keras.layers.Layer):
def __init__(self, units, activation=None):
super().__init__()
self.units = units
self.activation = activation
def build(self, input_shape):
self.w = self.add_weight(
name="weights",
shape=(input_shape[1], self.units),
initializer="random_normal",
trainable=True,
)
self.bias = self.add_weight(
name="bias",
shape=(self.units,),
initializer="random_normal",
trainable=True,
)
def call(self, inputs):
outputs = tnp.matmul(inputs, self.w) + self.bias
if self.activation:
return self.activation(outputs)
return outputs
def create_layered_tnp_model():
return keras.Sequential(
[
TNPDense(3, activation=tnp_relu),
TNPDense(3, activation=tnp_relu),
TNPDense(1),
]
)
model = create_layered_tnp_model()
model.compile(
optimizer="adam",
loss="mean_squared_error",
metrics=[keras.metrics.MeanAbsolutePercentageError()],
)
model.build((None, 13,))
model.summary()
evaluate_model(model)
```
You can also seamlessly switch between TNP layers and native Keras layers!
```
def create_mixed_model():
return keras.Sequential(
[
TNPDense(3, activation=tnp_relu),
# The model will have no issue using a normal Dense layer
layers.Dense(3, activation="relu"),
# ... or switching back to tnp layers!
TNPDense(1),
]
)
model = create_mixed_model()
model.compile(
optimizer="adam",
loss="mean_squared_error",
metrics=[keras.metrics.MeanAbsolutePercentageError()],
)
model.build((None, 13,))
model.summary()
evaluate_model(model)
```
The Keras API offers a wide variety of layers. The ability to use them alongside NumPy
code can be a huge time saver in projects.
## Distribution Strategy
TensorFlow NumPy and Keras integrate with
[TensorFlow Distribution Strategies](https://www.tensorflow.org/guide/distributed_training).
This makes it simple to perform distributed training across multiple GPUs,
or even an entire TPU Pod.
```
gpus = tf.config.list_logical_devices("GPU")
if gpus:
strategy = tf.distribute.MirroredStrategy(gpus)
else:
# We can fallback to a no-op CPU strategy.
strategy = tf.distribute.get_strategy()
print("Running with strategy:", str(strategy.__class__.__name__))
with strategy.scope():
model = create_layered_tnp_model()
model.compile(
optimizer="adam",
loss="mean_squared_error",
metrics=[keras.metrics.MeanAbsolutePercentageError()],
)
model.build((None, 13,))
model.summary()
evaluate_model(model)
```
## TensorBoard Integration
One of the many benefits of using the Keras API is the ability to monitor training
through TensorBoard. Using the TensorFlow NumPy API alongside Keras allows you to easily
leverage TensorBoard.
```
keras.backend.clear_session()
```
To load the TensorBoard from a Jupyter notebook, you can run the following magic:
```
%load_ext tensorboard
```
```
models = [
(TNPForwardFeedRegressionNetwork(blocks=[3, 3]), "TNPForwardFeedRegressionNetwork"),
(create_layered_tnp_model(), "layered_tnp_model"),
(create_mixed_model(), "mixed_model"),
]
for model, model_name in models:
model.compile(
optimizer="adam",
loss="mean_squared_error",
metrics=[keras.metrics.MeanAbsolutePercentageError()],
)
model.fit(
x_train,
y_train,
epochs=200,
verbose=0,
callbacks=[keras.callbacks.TensorBoard(log_dir=f"logs/{model_name}")],
)
```
To load the TensorBoard from a Jupyter notebook you can use the `%tensorboard` magic:
```
%tensorboard --logdir logs
```
The TensorBoard monitor metrics and examine the training curve.

The TensorBoard also allows you to explore the computation graph used in your models.

The ability to introspect into your models can be valuable during debugging.
## Conclusion
Porting existing NumPy code to Keras models using the `tensorflow_numpy` API is easy!
By integrating with Keras you gain the ability to use existing Keras callbacks, metrics
and optimizers, easily distribute your training and use Tensorboard.
Migrating a more complex model, such as a ResNet, to the TensorFlow NumPy API would be a
great follow up learning exercise.
Several open source NumPy ResNet implementations are available online.
| github_jupyter |
```
import os
import numpy as np
import pandas as pd
import prepare_data
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.svm import SVC
from sklearn import metrics
from sklearn.model_selection import train_test_split
from sklearn.model_selection import KFold
import matplotlib.pyplot as plt
import seaborn as sns
from keras.utils import np_utils
data_path = os.path.join("ICHI14_dataset\data")
patient_list = ['002','003','005','007','08a','08b','09a','09b', '10a','011','013','014','15a','15b','016',
'017','018','019','020','021','022','023','025','026','027','028','029','030','031','032',
'033','034','035','036','037','038','040','042','043','044','045','047','048','049','051']
#statistics_list = ["std_x", "std_y", "std_z", "ptp_x", "ptp_y", "ptp_z", "rms_x", "rms_y", "rms_z"]
statistics_list = ["std_x", "std_y", "std_z", "rms_x", "rms_y", "rms_z"]
file_name = "multy_class_features_60s.csv"
#prepare_data.save_statistic_features(patient_list, sorce_path="ICHI14_dataset\data",
# save_path="multy_class_features_60s.csv",
# window_len=60, sleep_stages=True)
train_patient_list, test_patient_list = train_test_split(patient_list, random_state=152, test_size=0.3)
X_train, y_train = prepare_data.load_statistic_features(train_patient_list,
file_name=file_name,
statistics_list=statistics_list)
X_test, y_test = prepare_data.load_statistic_features(test_patient_list,
file_name=file_name,
statistics_list=statistics_list)
print(X_train.shape)
print(y_train.shape)
print(X_test.shape)
print(y_test.shape)
sns.countplot(y_train)
sns.countplot(y_test)
```
### 1. Lin. Reg, only one window
```
%%time
model1 = LogisticRegression()
model1.fit(X_train, y_train)
y_predict = model1.predict(X_train)
print("\nTrain set result: ")
print(metrics.classification_report(y_train, y_predict))
print("Confussion matrix: \n", metrics.confusion_matrix(y_train, y_predict))
accuracy = metrics.accuracy_score(y_train, y_predict)
print("\nAccuracy on train set: ", accuracy)
y_predict = model1.predict(X_test)
print("\nTrain set result: ")
print(metrics.classification_report(y_test, y_predict))
print("Confussion matrix: \n", metrics.confusion_matrix(y_test, y_predict))
accuracy = metrics.accuracy_score(y_test, y_predict)
print("\nAccuracy on train set: ", accuracy)
```
f1 = 0.32, acc = 0.4866 std
f1 = 0.37, acc = 0.4508 std ptp rms
### 2. Lin. Reg, several windows features for one window
```
X_train, y_train = prepare_data.load_stat_features_others_windows(train_patient_list,
file_name=file_name,
statistics_list=statistics_list,
n_others_windows=16)
X_test, y_test = prepare_data.load_stat_features_others_windows(test_patient_list,
file_name=file_name,
statistics_list=statistics_list,
n_others_windows=16)
print(X_train.shape)
print(y_train.shape)
print(X_test.shape)
print(y_test.shape)
%%time
model3 = LogisticRegression()
model3.fit(X_train, y_train)
y_predict = model3.predict(X_train)
print("\nTrain set result: ")
print(metrics.classification_report(y_train, y_predict))
print("Confussion matrix: \n", metrics.confusion_matrix(y_train, y_predict))
accuracy = metrics.accuracy_score(y_train, y_predict)
print("\nAccuracy on train set: ", accuracy)
y_predict = model3.predict(X_test)
print("\nTest set result: ")
print(metrics.classification_report(y_test, y_predict))
print("Confussion matrix: \n", metrics.confusion_matrix(y_test, y_predict))
accuracy = metrics.accuracy_score(y_test, y_predict)
print("\nAccuracy on test set: ", accuracy)
```
f1 = 0.43, acc = 0.4937, 20 windows, std ptp rms
f1 = 0.44, acc = 0.5070, 20 windows, std rms
### The problem is imbalanced classes
## OVERSAMPLING
```
train_size = y_train.shape[0]
awake = np.sum(y_train[y_train == 4]) / 4
rem = np.sum(y_train[y_train == 0] + 1)
no_rem1 = np.sum(y_train[y_train == 1])
no_rem2 = np.sum(y_train[y_train == 2]) / 2
no_rem3 = np.sum(y_train[y_train == 3]) / 3
#print(awake)
#print(rem)
#print(no_rem1)
#print(no_rem2)
#print(no_rem3)
print(awake / train_size)
print(rem / train_size)
print(no_rem1 / train_size)
print(no_rem2 / train_size)
print(no_rem3 / train_size)
def n_repeats(y):
train_size = y.shape[0]
repeats = np.zeros(y.shape)
awake = np.sum(y_train[y_train == 4]) / 4
rem = np.sum(y_train[y_train == 0] + 1)
no_rem1 = np.sum(y_train[y_train == 1])
no_rem2 = np.sum(y_train[y_train == 2]) / 2
no_rem3 = np.sum(y_train[y_train == 3]) / 3
repeats[y == 4] = 1
repeats[y == 0] = awake // rem
repeats[y == 1] = awake // no_rem1
repeats[y == 2] = awake // no_rem2
repeats[y == 3] = awake // no_rem3
return repeats.astype(int)
repeats = n_repeats(y_train)
y_train = np.repeat(y_train, repeats, axis=0)
X_train = np.repeat(X_train, repeats, axis=0)
print(y_train.shape)
print(X_train.shape)
sns.countplot(y_train)
```
### 3. Lin. Reg, only one window
```
%%time
model1 = LogisticRegression(C=0.1)
model1.fit(X_train, y_train)
y_predict = model1.predict(X_train)
print("\nTrain set result: ")
print(metrics.classification_report(y_train, y_predict))
print("Confussion matrix: \n", metrics.confusion_matrix(y_train, y_predict))
accuracy = metrics.accuracy_score(y_train, y_predict)
print("\nAccuracy on train set: ", accuracy)
y_predict = model1.predict(X_test)
print("\nTrain set result: ")
print(metrics.classification_report(y_test, y_predict))
print("Confussion matrix: \n", metrics.confusion_matrix(y_test, y_predict))
accuracy = metrics.accuracy_score(y_test, y_predict)
print("\nAccuracy on train set: ", accuracy)
```
f1 = 0.35, acc = 0.35, std
f1 = 0.39, acc = 0.3707, std + ptp + rms
### 4. Lin. Reg, several windows features for one window
```
X_train, y_train = prepare_data.load_stat_features_others_windows(train_patient_list,
file_name=file_name,
statistics_list=statistics_list,
n_others_windows=16)
X_test, y_test = prepare_data.load_stat_features_others_windows(test_patient_list,
statistics_list=statistics_list,
file_name=file_name,
n_others_windows=16)
repeats = n_repeats(y_train)
y_train = np.repeat(y_train, repeats, axis=0)
X_train = np.repeat(X_train, repeats, axis=0)
sns.countplot(y_train)
print(X_train.shape)
print(y_train.shape)
print(X_test.shape)
print(y_test.shape)
%%time
model3 = LogisticRegression()
model3.fit(X_train, y_train)
y_predict = model3.predict(X_train)
print("\nTrain set result: ")
print(metrics.classification_report(y_train, y_predict))
print("Confussion matrix: \n", metrics.confusion_matrix(y_train, y_predict))
accuracy = metrics.accuracy_score(y_train, y_predict)
print("\nAccuracy on train set: ", accuracy)
y_predict = model3.predict(X_test)
print("\nTest set result: ")
print(metrics.classification_report(y_test, y_predict))
print("Confussion matrix: \n", metrics.confusion_matrix(y_test, y_predict))
accuracy = metrics.accuracy_score(y_test, y_predict)
print("\nAccuracy on test set: ", accuracy)
```
f1 = 0.42, acc = 0.4198, 16 windows
f1 = 0.43, acc = 0.4398, 32 windows
f1 = 0.43, acc = 0.4279, 40 windows
f1 = 0.39, acc = 0.3713, 20 windows, std + ptp + rms
### 5. GradientBoostingClassifier, several windows features for one window
```
%%time
model4 = GradientBoostingClassifier(n_estimators=50, max_depth=7)
model4.fit(X_train, y_train)
y_predict = model4.predict(X_train)
print("\nTrain set result: ")
print(metrics.classification_report(y_train, y_predict))
print("Confussion matrix: \n", metrics.confusion_matrix(y_train, y_predict))
accuracy = metrics.accuracy_score(y_train, y_predict)
print("\nAccuracy on train set: ", accuracy)
y_predict = model4.predict(X_test)
print("\nTest set result: ")
print(metrics.classification_report(y_test, y_predict))
print("Confussion matrix: \n", metrics.confusion_matrix(y_test, y_predict))
accuracy = metrics.accuracy_score(y_test, y_predict)
print("\nAccuracy on test set: ", accuracy)
```
f1-score = 0.43
f1-score = 0.45 n_estimators=30, max_depth=6
f1-score = 0.48 , acc = 0.5002 (n_estimators=50, max_depth=10) 16 windows
f1-score = 0.47 , acc = 0.4593 (n_estimators=50, max_depth=6), 32 windows
f1-score = 0.43 , acc = 0.4278 (n_estimators=50, max_depth=6) 16 windows, std + ptp + rms
f1-score = 0.44 , acc = 0.4407 (n_estimators=50, max_depth=6) 16 windows, std + rms
### 6. SVM, several windows features for one window
```
%%time
model6 = SVC(C=0.5)
model6.fit(X_train, y_train)
y_predict = model6.predict(X_train)
print("\nTrain set result: ")
print(metrics.classification_report(y_train, y_predict))
print("Confussion matrix: \n", metrics.confusion_matrix(y_train, y_predict))
accuracy = metrics.accuracy_score(y_train, y_predict)
print("\nAccuracy on train set: ", accuracy)
y_predict = model6.predict(X_test)
print("\nTest set result: ")
print(metrics.classification_report(y_test, y_predict))
print("Confussion matrix: \n", metrics.confusion_matrix(y_test, y_predict))
accuracy = metrics.accuracy_score(y_test, y_predict)
print("\nAccuracy on test set: ", accuracy)
```
f1 = 0.41 C=0.1
f1 = 0.44, acc = 0.4840, C=0.5
### 7. RNN, several windows features for one window
```
from keras.layers import Dense, Flatten, Dropout, LSTM, Bidirectional, Activation, Embedding
from keras.layers import Conv1D, MaxPooling1D
from keras.models import Sequential
from keras.optimizers import SGD, Adam
from keras.layers.normalization import BatchNormalization
from keras.regularizers import l2
from keras.layers import LSTM, Bidirectional
from keras.callbacks import ModelCheckpoint, EarlyStopping
from keras.utils import np_utils
train_patient_list, test_patient_list = train_test_split(patient_list, random_state=152, test_size=0.3)
test_patient_list, valid_patient_list = train_test_split(test_patient_list, random_state=151, test_size=0.5)
X_train, y_train = prepare_data.load_stat_features_others_windows_rnn(train_patient_list,
file_name=file_name,
n_others_windows=12)
X_valid, y_valid = prepare_data.load_stat_features_others_windows_rnn(valid_patient_list,
file_name=file_name,
n_others_windows=12)
X_test, y_test = prepare_data.load_stat_features_others_windows_rnn(test_patient_list,
file_name=file_name,
n_others_windows=12)
repeats = n_repeats(y_train)
y_train = np.repeat(y_train, repeats, axis=0)
X_train = np.repeat(X_train, repeats, axis=0)
sns.countplot(y_train)
print(X_train.shape)
print(y_train.shape)
print(X_valid.shape)
print(y_valid.shape)
print(X_test.shape)
print(y_test.shape)
print("One=hot shapes")
Y_train = np_utils.to_categorical(y_train, 5)
print(Y_train.shape)
Y_valid = np_utils.to_categorical(y_valid, 5)
print(Y_valid.shape)
Y_test = np_utils.to_categorical(y_test, 5)
print(Y_test.shape)
RNN = Sequential()
RNN.add(LSTM(13, dropout=0.2, recurrent_dropout=0.1, input_shape=(13, 3)))
RNN.add(Dense(5, activation="softmax", kernel_initializer="glorot_uniform", kernel_regularizer=l2(0.1)))
RNN.compile(loss="categorical_crossentropy", optimizer="adam", metrics=["accuracy"])
print(RNN.summary())
callbacks = [ModelCheckpoint('RNN_multy_5_weights.hdf5', monitor='val_loss', save_best_only=True),
EarlyStopping(monitor='val_loss', patience=4)]
%%time
RNN.fit(X_train, Y_train,
shuffle=True,
batch_size=32,
epochs=10,
validation_data=(X_valid, Y_valid),
callbacks=callbacks,
verbose=1)
scores = RNN.evaluate(X_test, Y_test)
print("Test accuracy =", scores[1])
y_predict = np.argmax(RNN.predict(X_test), axis=1)
metrics.f1_score(y_test, y_predict, average='micro')
metrics.f1_score(y_test, y_predict, average='macro')
metrics.f1_score(y_test, y_predict, average='weighted')
y_predict = np.argmax(RNN.predict(X_train), axis=1)
print("\nTrain set result: ")
print(metrics.classification_report(y_train, y_predict))
print("Confussion matrix: \n", metrics.confusion_matrix(y_train, y_predict))
accuracy = metrics.accuracy_score(y_train, y_predict)
print("\nAccuracy on train set: ", accuracy)
y_predict = np.argmax(RNN.predict(X_test), axis=1)
print("\nTest set result: ")
print(metrics.classification_report(y_test, y_predict))
print("Confussion matrix: \n", metrics.confusion_matrix(y_test, y_predict))
accuracy = metrics.accuracy_score(y_test, y_predict)
print("\nAccuracy on test set: ", accuracy)
```
## Cross validation
```
kf = KFold(n_splits=5, random_state=5, shuffle=True) # Define the split - into 3 folds #5
kf.get_n_splits(patient_list) # returns the number of splitting iterations in the cross-validator
for train_index, test_index in kf.split(patient_list):
#train_patient_list = [patient_list[i] for i in train_index]
test_patient_list = [patient_list[i] for i in test_index]
print(test_patient_list)
n_others_windows = 32
```
### Gradient Boosting
```
%%time
accuracy_list = []
f1_list = []
for train_index, test_index in kf.split(patient_list):
train_patient_list = [patient_list[i] for i in train_index]
test_patient_list = [patient_list[i] for i in test_index]
X_train, y_train = prepare_data.load_stat_features_others_windows(train_patient_list,
file_name=file_name,
n_others_windows=n_others_windows)
X_test, y_test = prepare_data.load_stat_features_others_windows(test_patient_list,
file_name=file_name,
n_others_windows=n_others_windows)
repeats = n_repeats(y_train)
y_train = np.repeat(y_train, repeats, axis=0)
X_train = np.repeat(X_train, repeats, axis=0)
print(X_train.shape)
model3 = GradientBoostingClassifier(n_estimators=30, max_depth=6)
model3.fit(X_train, y_train)
y_predict = model3.predict(X_train)
accuracy_train = metrics.accuracy_score(y_train, y_predict)
f1_train = metrics.f1_score(y_train, y_predict, average="weighted")
print("\nAccuracy on train set: ", accuracy_train)
#print("F1 score on train set: ", f1_train)
y_predict = model3.predict(X_test)
accuracy = metrics.accuracy_score(y_test, y_predict)
f1_test = metrics.f1_score(y_test, y_predict, average="weighted")
accuracy_list.append(accuracy)
print(metrics.classification_report(y_test, y_predict))
f1_list.append(f1_test)
print("Accuracy on test set: ", accuracy)
#print("F1 score on test set: ", f1_test)
#print(metrics.classification_report(y_test, y_predict))
#print(test_patient_list)
print("\nMean accuracy =", np.mean(accuracy_list))
print("Mean f1 score =", np.mean(f1_list))
```
f1 = 0.3892, acc = 0.3841, 16 windows, n_estimators=30, max_depth=6
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/Visualization/styled_layer_descriptors.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/Visualization/styled_layer_descriptors.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://mybinder.org/v2/gh/giswqs/earthengine-py-notebooks/master?filepath=Visualization/styled_layer_descriptors.ipynb"><img width=58px src="https://mybinder.org/static/images/logo_social.png" />Run in binder</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/Visualization/styled_layer_descriptors.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geehydro](https://github.com/giswqs/geehydro). The **geehydro** Python package builds on the [folium](https://github.com/python-visualization/folium) package and implements several methods for displaying Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, `Map.centerObject()`, and `Map.setOptions()`.
The following script checks if the geehydro package has been installed. If not, it will install geehydro, which automatically install its dependencies, including earthengine-api and folium.
```
import subprocess
try:
import geehydro
except ImportError:
print('geehydro package not installed. Installing ...')
subprocess.check_call(["python", '-m', 'pip', 'install', 'geehydro'])
```
Import libraries
```
import ee
import folium
import geehydro
```
Authenticate and initialize Earth Engine API. You only need to authenticate the Earth Engine API once.
```
try:
ee.Initialize()
except Exception as e:
ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
This step creates an interactive map using [folium](https://github.com/python-visualization/folium). The default basemap is the OpenStreetMap. Additional basemaps can be added using the `Map.setOptions()` function.
The optional basemaps can be `ROADMAP`, `SATELLITE`, `HYBRID`, `TERRAIN`, or `ESRI`.
```
Map = folium.Map(location=[40, -100], zoom_start=4)
Map.setOptions('HYBRID')
```
## Add Earth Engine Python script
```
cover = ee.Image('MODIS/051/MCD12Q1/2012_01_01').select('Land_Cover_Type_1')
# Define an SLD style of discrete intervals to apply to the image.
sld_intervals = \
'<RasterSymbolizer>' + \
' <ColorMap type="intervals" extended="false" >' + \
'<ColorMapEntry color="#aec3d4" quantity="0" label="Water"/>' + \
'<ColorMapEntry color="#152106" quantity="1" label="Evergreen Needleleaf Forest"/>' + \
'<ColorMapEntry color="#225129" quantity="2" label="Evergreen Broadleaf Forest"/>' + \
'<ColorMapEntry color="#369b47" quantity="3" label="Deciduous Needleleaf Forest"/>' + \
'<ColorMapEntry color="#30eb5b" quantity="4" label="Deciduous Broadleaf Forest"/>' + \
'<ColorMapEntry color="#387242" quantity="5" label="Mixed Deciduous Forest"/>' + \
'<ColorMapEntry color="#6a2325" quantity="6" label="Closed Shrubland"/>' + \
'<ColorMapEntry color="#c3aa69" quantity="7" label="Open Shrubland"/>' + \
'<ColorMapEntry color="#b76031" quantity="8" label="Woody Savanna"/>' + \
'<ColorMapEntry color="#d9903d" quantity="9" label="Savanna"/>' + \
'<ColorMapEntry color="#91af40" quantity="10" label="Grassland"/>' + \
'<ColorMapEntry color="#111149" quantity="11" label="Permanent Wetland"/>' + \
'<ColorMapEntry color="#cdb33b" quantity="12" label="Cropland"/>' + \
'<ColorMapEntry color="#cc0013" quantity="13" label="Urban"/>' + \
'<ColorMapEntry color="#33280d" quantity="14" label="Crop, Natural Veg. Mosaic"/>' + \
'<ColorMapEntry color="#d7cdcc" quantity="15" label="Permanent Snow, Ice"/>' + \
'<ColorMapEntry color="#f7e084" quantity="16" label="Barren, Desert"/>' + \
'<ColorMapEntry color="#6f6f6f" quantity="17" label="Tundra"/>' + \
'</ColorMap>' + \
'</RasterSymbolizer>'
Map.addLayer(cover.sldStyle(sld_intervals), {}, 'IGBP classification styled')
```
## Display Earth Engine data layers
```
Map.setControlVisibility(layerControl=True, fullscreenControl=True, latLngPopup=True)
Map
```
| github_jupyter |
# Plot of posterior samples
_Kara Ponder (SLAC-->?), Emille Ishida (Clermont-Ferrand), Alex Malz (GCCL@RUB)_
plagiarized from `Combination_plots.ipynb`
```
from collections import OrderedDict
import glob
import gzip
import numpy as np
import os
import pandas as pd
import pickle as pkl
import scipy.stats as sps
import sys
```
## Kara's plotting code
```
%matplotlib inline
import matplotlib.pyplot as plt
import matplotlib.colors as colors
import matplotlib.cm as cmx
from mpl_toolkits.axes_grid1 import make_axes_locatable
# import resspect.cosmo_metric_utils as cmu
import pylab
from mpl_toolkits.axes_grid1 import ImageGrid
from mpl_toolkits.axes_grid1.inset_locator import inset_axes
all_shapes = {'SNIa-91bg': 'o',
'SNIax': 's',
'SNII': 'd',
'SNIbc': 'X',
'SLSN-I': 'v',
'AGN': '^',
'TDE': '<',
'KN': '>',
'CART': 'v'}
# Color map
rainbow = cm = plt.get_cmap('plasma_r')
cNorm = colors.LogNorm(vmin=1, vmax=52) #colors.Normalize(vmin=0, vmax=50)
scalarMap = cmx.ScalarMappable(norm=cNorm, cmap=rainbow)
color_map = scalarMap.to_rgba(np.arange(1, 52))
```
## prep for data
```
# DDF summary on the COIN server:
file_extensions = {'ddf': 'DDF',
'wfd': 'WFD'
}
ktot = 5
kglob = ''
def get_cases(field, k=''):
if k == '':
k = '0'
dirname = '/media/RESSPECT/data/PLAsTiCC/for_metrics/final_data/'+field+'/v'+k+'/samples/'
cases = os.listdir(dirname)
cases.remove('random1500.csv')
cases.remove('random6000.csv')
cases.remove('fiducial1500.csv')
cases.remove('fiducial6000.csv')
cases.remove('perfect6000.csv')
cases.remove('perfect1500.csv')
print(cases)
return(cases, dirname)
cases, dirnames = {}, {}
for file_extension in file_extensions:
cases[file_extension], dirnames[file_extension] = get_cases(file_extensions[file_extension])
def make_remap_dict(file_extension):
if 'wfd' in file_extension:
remap_dict = OrderedDict({
'perfect3000': 'Perfect',
'fiducial3000': 'Fiducial',
'random3000': 'Random',
'all_objs_survived_SALT2_DDF' : 'All SALT',
'all_objs_survived_SALT2_WFD': 'All SALT',
'50SNIa50SNII': 'SN-II 50',
'68SNIa32SNII': 'SN-II 32',
'72SNIa28SNII': 'SN-II 28',
'75SNIa25SNII': 'SN-II 25',
'90SNIa10SNII': 'SN-II 10',
'95SNIa5SNII': 'SN-II 5',
'98SNIa2SNII': 'SN-II 2',
'99SNIa1SNII': 'SN-II 1',
'50SNIa50SNIbc': 'SN-Ibc 50',
'68SNIa32SNIbc': 'SN-Ibc 32',
'75SNIa25SNIbc': 'SN-Ibc 25',
'83SNIa17SNIbc': 'SN-Ibc 17',
'90SNIa10SNIbc': 'SN-Ibc 10',
'95SNIa5SNIbc': 'SN-Ibc 5',
'98SNIa2SNIbc': 'SN-Ibc 2',
'99SNIa1SNIbc': 'SN-Ibc 1',
'50SNIa50SNIax': 'SN-Iax 50',
'68SNIa32SNIax': 'SN-Iax 32',
'75SNIa25SNIax': 'SN-Iax 25',
'86SNIa14SNIax': 'SN-Iax 14',
'90SNIa10SNIax': 'SN-Iax 10',
'94SNIa6SNIax': 'SN-Iax 6',
'95SNIa5SNIax': 'SN-Iax 5',
'97SNIa3SNIax': 'SN-Iax 3',
'98SNIa2SNIax': 'SN-Iax 2',
'99SNIa1SNIax': 'SN-Iax 1',
'71SNIa29SNIa-91bg': 'SN-Ia-91bg 29',
'75SNIa25SNIa-91bg': 'SN-Ia-91bg 25',
'90SNIa10SNIa-91bg': 'SN-Ia-91bg 10',
'95SNIa5SNIa-91bg': 'SN-Ia-91bg 5',
'98SNIa2SNIa-91bg': 'SN-Ia-91bg 2',
'99SNIa1SNIa-91bg': 'SN-Ia-91bg 1',
'99.8SNIa0.2SNIa-91bg': 'SN-Ia-91bg 0.2',
'57SNIa43AGN': 'AGN 43',
'75SNIa25AGN': 'AGN 25',
'90SNIa10AGN': 'AGN 10',
'94SNIa6AGN': 'AGN 6',
'95SNIa5AGN': 'AGN 5',
'98SNIa2AGN': 'AGN 2',
'99SNIa1AGN': 'AGN 1',
'99.9SNIa0.1AGN': 'AGN 0.1',
'83SNIa17SLSN-I': 'SLSN-I 17',
'90SNIa10SLSN-I': 'SLSN-I 10',
'95SNIa5SLSN-I': 'SLSN-I 5',
'98SNIa2SLSN-I': 'SLSN-I 2',
'99SNIa1SLSN-I': 'SLSN-I 1',
'99SNIa1SLSN': 'SLSN 1',
'99.9SNIa0.1SLSN': 'SLSN-I 0.1',
'95SNIa5TDE': 'TDE 5',
'98SNIa2TDE': 'TDE 2',
'99SNIa1TDE': 'TDE 1',
'99.6SNIa0.4TDE': 'TDE 0.4',
'99.1SNIa0.9CART': 'CART 0.9',
'99.7SNIa0.3CART': 'CART 0.3'
})
else:
remap_dict = OrderedDict({
'perfect3000': 'Perfect',
'fiducial3000': 'Fiducial',
'random3000': 'Random',
'all_objs_survived_SALT2_DDF' : 'All SALT',
'all_objs_survived_SALT2_WFD': 'All SALT',
'50SNIa50SNII': 'SN-II 50',
'68SNIa32SNII': 'SN-II 32',
'72SNIa28SNII': 'SN-II 28',
'75SNIa25SNII': 'SN-II 25',
'90SNIa10SNII': 'SN-II 10',
'95SNIa5SNII': 'SN-II 5',
'98SNIa2SNII': 'SN-II 2',
'99SNIa1SNII': 'SN-II 1',
'50SNIa50SNIbc': 'SN-Ibc 50',
'68SNIa32SNIbc': 'SN-Ibc 32',
'75SNIa25SNIbc': 'SN-Ibc 25',
'83SNIa17SNIbc': 'SN-Ibc 17',
'90SNIa10SNIbc': 'SN-Ibc 10',
'92SNIa8SNIbc': 'SN-Ibc 8',
'95SNIa5SNIbc': 'SN-Ibc 5',
'98SNIa2SNIbc': 'SN-Ibc 2',
'99SNIa1SNIbc': 'SN-Ibc 1',
'50SNIa50SNIax': 'SN-Iax 50',
'68SNIa32SNIax': 'SN-Iax 32',
'75SNIa25SNIax': 'SN-Iax 25',
'86SNIa14SNIax': 'SN-Iax 14',
'90SNIa10SNIax': 'SN-Iax 10',
'91SNIa9SNIax': 'SN-Iax 9',
'94SNIa6SNIax': 'SN-Iax 6',
'95SNIa5SNIax': 'SN-Iax 5',
'97SNIa3SNIax': 'SN-Iax 3',
'98SNIa2SNIax': 'SN-Iax 2',
'99SNIa1SNIax': 'SN-Iax 1',
'99.1SNIa0.9CART': 'CART 0.9',
'99.7SNIa0.3CART': 'CART 0.3',
'71SNIa29SNIa-91bg': 'SN-Ia-91bg 29',
'75SNIa25SNIa-91bg': 'SN-Ia-91bg 25',
'90SNIa10SNIa-91bg': 'SN-Ia-91bg 10',
'95SNIa5SNIa-91bg': 'SN-Ia-91bg 5',
'98SNIa2SNIa-91bg': 'SN-Ia-91bg 2',
'99SNIa1SNIa-91bg': 'SN-Ia-91bg 1',
'99.8SNIa0.2SNIa-91bg': 'SN-Ia-91bg 0.2',
'57SNIa43AGN': 'AGN 43',
'75SNIa25AGN': 'AGN 25',
'90SNIa10AGN': 'AGN 10',
'94SNIa6AGN': 'AGN 6',
'95SNIa5AGN': 'AGN 5',
'98SNIa2AGN': 'AGN 2',
'99SNIa1AGN': 'AGN 1',
'99.9SNIa0.1AGN': 'AGN 0.1',
'83SNIa17SLSN-I': 'SLSN-I 17',
'90SNIa10SLSN-I': 'SLSN-I 10',
'95SNIa5SLSN-I': 'SLSN-I 5',
'98SNIa2SLSN-I': 'SLSN-I 2',
'99SNIa1SLSN-I': 'SLSN-I 1',
'99SNIa1SLSN': 'SLSN 1',
'99.9SNIa0.1SLSN': 'SLSN-I 0.1',
'95SNIa5TDE': 'TDE 5',
'98SNIa2TDE': 'TDE 2',
'99SNIa1TDE': 'TDE 1',
'99.6SNIa0.4TDE': 'TDE 0.4',
})
return(remap_dict)
remap_dicts = {}
for file_extension in file_extensions:
thing = make_remap_dict(file_extensions[file_extension])
tempdict = {}
for case in cases[file_extension]:
if case[:-4] in thing.keys():
tempdict[case[:-4]] = thing[case[:-4]]
else:
print(case)
remap_dicts[file_extension] = tempdict#{thing[case[:-4]] for case in cases[file_extension]}
# Mapping the percent contaminated to the colormap.
## size corresponds to remap_dict
def make_color_nums(file_extension):
if 'wfd' in file_extension:
color_num = np.array([1, 1, 1, 1, 1, 1, # Special
50, 32, 28, 25, 10, 5, 2, 1, # II
50, 32, 25, 17, 10, 5, 2, 1, # Ibc
50, 32, 25, 14, 10, 6, 5, 3, 2, 1, # Iax
29, 25, 10, 5, 2, 1, 1, # 91bg
43, 25, 10, 6, 5, 2, 1, 1, # AGN
17, 10, 5, 2, 1, 1, 1, # SLSN
5, 2, 1, 1, # TDE
1, 1, # CART
]) #+ 1
else:
color_num = np.array([1, 1, 1, 1, 1, 1, # Special
50, 32, 28, 25, 10, 5, 2, 1, # II
50, 32, 25, 17, 10, 8, 5, 2, 1, # Ibc
50, 32, 25, 14, 10, 9, 6, 5, 3, 2, 1, # Iax
1, 1, # CART
29, 25, 10, 5, 2, 1, 1, # 91bg
43, 25, 10, 6, 5, 2, 1, 1, # AGN
17, 10, 5, 2, 1, 1, 1, # SLSN
5, 2, 1, 1, # TDE
]) #+ 1
return(color_num)
color_nums = {}
for file_extension in file_extensions:
color_nums[file_extension] = make_color_nums(file_extensions[file_extension])
# Color map
rainbow = cm = plt.get_cmap('plasma_r')
cNorm = colors.LogNorm(vmin=1, vmax=52) #colors.Normalize(vmin=0, vmax=50)
scalarMap = cmx.ScalarMappable(norm=cNorm, cmap=rainbow)
color_map = scalarMap.to_rgba(np.arange(1, 52))
```
## calculate the curve(s)
KDE for each set of posterior samples
```
eps = 2. * sys.float_info.min
def safe_log(arr, threshold=eps):
"""
Takes the natural logarithm of an array that might contain zeros.
Parameters
----------
arr: ndarray, float
array of values to be logged
threshold: float, optional
small, positive value to replace zeros and negative numbers
Returns
-------
logged: ndarray
logged values, with small value replacing un-loggable values
"""
arr = np.asarray(arr)
arr[arr < threshold] = threshold
logged = np.log(arr)
return logged
def make_grid(x, y, x_ngrid=100, y_ngrid=100):
x_min = x.min()#-1.2
x_max = x.max()#-0.8
y_min = y.min()#0.2
y_max = y.max()#0.4
x_grid, y_grid = np.mgrid[x_min:x_max:x_ngrid*1.j, y_min:y_max:y_ngrid*1.j]
x_vec, y_vec = x_grid[:, 0], y_grid[0, :]
dx = (x_max - x_min) / (x_ngrid - 1)
dy = (y_max - y_min) / (y_ngrid - 1)
return(((x_min, y_min), (x_max, y_max)), (x_grid, y_grid), (x_vec, y_vec), (dx, dy))
def make_kde(Xgrid, Ygrid, Xsamps, Ysamps, to_log=False, save=None, one_d=True):
if not one_d:
positions = np.vstack([Xgrid.ravel(), Ygrid.ravel()])
values = np.vstack([Xsamps, Ysamps])
kernel = sps.gaussian_kde(values, bw_method='scott')
Z = np.reshape(kernel(positions).T, Xgrid.shape)
else:
positions = Xgrid.T[0]
values = Xsamps
kernel = sps.gaussian_kde(values, bw_method='scott')
Z = kernel(positions)
if to_log:
return safe_log(Z)
else:
return Z
# if save is not None:
# TODO: normalize up here before log!
# alloutputs = pd.DataFrame(columns=['path', 'KLD'])
# # make reference sample
# with gzip.open(fullpath+refname) as reffn:
# flatref = pd.read_csv(reffn)
# [w_ref, Omm_ref] = [flatref['w'], flatref['om']]
# ref_extrema, ref_grids, ref_vecs, ref_ds = make_grid(w_ref, Omm_ref)
# (w_vec, Omm_vec) = ref_vecs
# (dw, dOmm) = ref_ds
# ((xmin, ymin), (xmax, ymax)) = ref_extrema
# (w_grid, Omm_grid) = ref_grids
# d_ref = {'w': dw, 'Omm': dOmm}
# grid_ref = {'w': w_grid, 'Omm': Omm_grid}
# kde_ref = make_kde(w_grid, Omm_grid, w_ref, Omm_ref, one_d=True, to_log=True)
def get_posteriors(field, k, casename, nsn, withlowz=True):
case = casename+str(nsn)
filename = 'chains_'+case
if withlowz:
filename = filename+'_lowz_withbias'
path_pre = '/media/RESSPECT/data/PLAsTiCC/for_metrics/final_data/'+file_extensions[field]+'/v'+str(k)+'/posteriors/pkl/'
# if field == 'ddf':
# # if k == '':
# # k = 0
# path_pre = '/media/RESSPECT/data/PLAsTiCC/for_metrics/final_data/DDF/v'+str(k)+'/posteriors/pkl/'
# # path_pre = '/media/RESSPECT/data/PLAsTiCC/for_metrics/ddf/posteriors/samples_emille/'
# # ext = '.csv.gz'
# # else:
# # path_pre = '/media/RESSPECT/data/PLAsTiCC/for_metrics/ddf/emille_samples'+str(k)+'/posteriors/'
ext = '.pkl'
# elif field == 'wfd':
# path_pre = '/media/RESSPECT/data/PLAsTiCC/for_metrics/wfd/posteriors/samples_emille'+str(k)+'/'
# ext = '.csv.gz'
samppathname = path_pre+filename+ext
# print(samppathname)
if ext == '.csv.gz':
with gzip.open(samppathname) as sampfile:
sampdata = pd.read_csv(sampfile)
elif ext == '.pkl':
with open(samppathname, 'rb') as sampfile:
sampdata = pkl.load(sampfile)
# print(sampdata)
return([sampdata['w'], sampdata['om']])
null_cases = ['perfect', 'random', 'fiducial']
ktot = 6
kmin = 0
samp_sizes = [1500, 3000, 6000]
ngrid = 100
outdata = {}
for field in file_extensions:
outdata[field] = {}
for casename in null_cases:
outdata[field][casename] = np.empty((ktot, len(samp_sizes), 2, ngrid))
for k in range(kmin, ktot, 1):
for i, nsn in enumerate(samp_sizes):
kpass = k
[w_comp, Omm_comp] = get_posteriors(field, kpass, casename, nsn, withlowz=True)#[sampdata['w'], sampdata['om']]
comp_extrema, comp_grids, comp_vecs, comp_ds = make_grid(w_comp, Omm_comp)
(w_grid, Omm_grid) = comp_grids
kde_comp = make_kde(w_grid, Omm_grid, w_comp, Omm_comp, one_d=True, to_log=True)
outdata[field][casename][k][i] = np.array([w_grid.T[0], kde_comp])
with open('default_kdes.pkl', 'wb') as outfile:
pkl.dump(outdata, outfile)
outdata = {}
for field in file_extensions:
outdata[field] = {}
for casename in cases[field]:
outdata[field][casename[:-4]] = np.empty((2, ngrid))
k = '0'
nsn = ''
[w_comp, Omm_comp] = get_posteriors(field, k, casename[:-4], nsn, withlowz=True)#[sampdata['w'], sampdata['om']]
comp_extrema, comp_grids, comp_vecs, comp_ds = make_grid(w_comp, Omm_comp)
(w_grid, Omm_grid) = comp_grids
kde_comp = make_kde(w_grid, Omm_grid, w_comp, Omm_comp, one_d=True, to_log=True)
outdata[field][casename[:-4]] = np.array([w_grid.T[0], kde_comp])
with open('testcase_kdes.pkl', 'wb') as outfile:
pkl.dump(outdata, outfile)
```
## make plot(s)
```
def_colors = {'perfect': 'k', 'random': 'tab:red', 'fiducial': 'tab:blue'}
def_styles = {'1500': ':', '3000': '-', '6000': '--'}#{'DDF': '-', 'WFD': '--'}
# def_lowz = {'withbias': , 'nobias':}
with open('default_kdes.pkl', 'rb') as infile:
indata = pkl.load(infile)
fig, ax = plt.subplots(2, 1, figsize=(6, 7))
for j, field in enumerate(file_extensions):
for casename in null_cases:
ax[j].scatter([0], [0], label=casename, color=def_colors[casename])
for i, nsn in enumerate(samp_sizes):
for k in range(ktot-1, ktot):
w_grid, kde_comp = indata[field][casename][k][i]#[w_grid, kde_comp] = indata[casename]
# if k == 0:
# lw_boost = 2
# # print(kde_comp)
# else:
# lw_boost = 1
ax[j].plot(w_grid, np.exp(kde_comp),# label=field+casename,
linestyle=def_styles[str(nsn)], color=def_colors[casename], alpha=0.8, linewidth=1.25)
for nsn in samp_sizes:
ax[j].plot([0], [0], label=str(nsn),
linestyle=def_styles[str(nsn)], color='tab:green', alpha=1., linewidth=1.25)
ax[j].text(-1., 40., file_extensions[field], fontsize=20)
ax[j].set_yticks([10, 30, 50])
ax[j].set_yticklabels([10, 30, 50], fontsize=14)
ax[j].set_ylabel(r'PDF ($w^{-1}$)', fontsize=18)
ax[j].set_ylim(0., 55.)
if j == 0:
ax[j].set_xticks([])
# plt.title(field+k)
if j == 1:
ax[j].set_xticks([-1.2, -1.1, -1.])
ax[j].set_xticklabels([-1.2, -1.1, -1.], fontsize=14)
ax[j].legend(loc='upper left', fontsize=14)#, ncol=2)
ax[j].set_xlabel(r'$w$', fontsize=18)
ax[j].set_xlim(-1.2, -0.95)
fig.subplots_adjust(wspace=0., hspace=0.)
plt.savefig('dists_null.png', bbox_inches='tight' ,dpi=250)
plt.show()
```
todo: investigate the runs that are flat KDEs
todo: also with and without bias of lowz sample
```
rates, contaminants = {}, {}
for field in file_extensions:
rate, contaminant = {}, {}
for key in remap_dicts[field]:
postsplit = remap_dicts[field][key].split()
if len(postsplit) > 1:
name = postsplit[0]
perc = float(postsplit[-1])
# rate[name] = perc
rate[key] = perc
contaminant[key] = name
rates[field] = rate
contaminants[field] = contaminant
# for field in file_extensions:
# plt.hist(rates[field].values(), bins=25, alpha=0.5, label=field)
# plt.legend()
```
todo: automate dividing into panels
```
cutoffs = [0., 1., 2., 5., 7.5, 15., 50.]
cutofflabels = ['<1%', '1%', '2%', '5%', '10%', '25%']
panel_groups = {}
for field in file_extensions:
panel_groups[field] = {j: [] for j in range(6)}
# print(field)
for i, casefn in enumerate(rates[field]):
casename = casefn#[:-4]
rate = rates[field][casename]
# for j, cutoff in enumerate(cutoffs[:-1]):
# if rate > cutoffs[j] and rate < :
# panel_groups[field][0].append(casename)
if rate > 0. and rate < 1.:
panel_groups[field][0].append(casename)
# print((casename, rates[field][casename], 0))
elif rate >= 1. and rate < 2.:
panel_groups[field][1].append(casename)
# print((casename, rates[field][casename], 1))
elif rate >= 2. and rate < 5.:
panel_groups[field][2].append(casename)
# print((casename, rates[field][casename], 1))
elif rate >= 5. and rate < 7.5:
panel_groups[field][3].append(casename)
# print((casename, rates[field][casename], 5))
elif rate >= 7.5 and rate <= 15.:
panel_groups[field][4].append(casename)
# print((casename, rates[field][casename], 10))
elif rate >= 15. and rate <= 50.:
panel_groups[field][5].append(casename)
# print((casename, rates[field][casename], 25))
# else:
# print((casename, rates[field][casename], 'big'))
print(panel_groups)
ddf_set = set(contaminants['ddf'].values())
# if len(file_extensions) > 1:
# for field in file_extensions[1:]:
# base_contaminant_set = set.union(base_contaminant_set, set(contaminants[field].values()))
wfd_set = set(contaminants['wfd'].values())
all_contaminants = set.union(ddf_set, wfd_set)
# base_contaminant_set#
color_list = OrderedDict({contaminant: plt.cm.tab10(i) for i, contaminant in enumerate(all_contaminants)})
contaminant_colors = {}
for field in file_extensions:
contaminant_colors[field] = {}
for i, contaminant in enumerate(contaminants[field]):
contaminant_colors[field][contaminant] = color_list[contaminants[field][contaminant]]
# def_colors = {'perfect': 'k', 'random': 'tab:red', 'fiducial': 'tab:blue'}
# def_styles = {'1500': ':', '3000': '-', '6000': '--'}#{'DDF': '-', 'WFD': '--'}
# def_lowz = {'withbias': , 'nobias':}
with open('testcase_kdes.pkl', 'rb') as infile:
indata = pkl.load(infile)
for field in file_extensions:
table_loc = '/media/RESSPECT/data/PLAsTiCC/for_metrics/final_data/'+file_extensions[field]+'/v'+str(0)+'/summary_stats.csv'
# if field == 'wfd':
# table_loc = '/media2/RESSPECT2/data/posteriors_wfd/omprior_0.01_flat/summary_cases_omprior_0.01_flat_emille.csv'
# else:
# table_loc = '/media2/RESSPECT2/data/posteriors_ddf/omprior_0.01_flat/summary_cases_emille.csv'
df = pd.read_csv(table_loc)
df = df.set_index('case')
# print((field, df.index))
fig = pylab.figure(figsize=(15, 10))
bigAxes = pylab.axes(frameon=False) # hide frame
bigAxes.set_xticks([]) # don't want to see any ticks on this axis
bigAxes.set_yticks([])
# bigAxes.set_xlabel(r'$-1.3 < w < -0.9$', fontsize=20)
bigAxes.set_title(file_extensions[field], fontsize=20)
numrows=2
numcols=3
# fig, ax = plt.subplots(2, 3, figsize=(15, 10))
# fig.suptitle(file_extensions[field])
for i in range(len(panel_groups[field])):
per_panel_contaminants = [contaminants[field][panel_groups[field][i][j]]
for j in range(len(panel_groups[field][i]))]
uniques, unique_ind = np.unique(per_panel_contaminants, return_index=True)
ax = fig.add_subplot(numrows,numcols,i+1)
# ax.spines['right'].set_visible(False)
# ax.spines['top'].set_visible(False)
# position = ax.get_position()
# position.x0 += 0.01
# position.y0 += 0.02
# position.x1 += 0.01
# position.y1 += 0.02
# ax.set_position(position)
stylecount = 0
ax.text(-0.95, 35., cutofflabels[i], fontsize=18, horizontalalignment='right', verticalalignment='center')
for j, val in enumerate(unique_ind):
casename = panel_groups[field][i][val]
w_grid, kde_comp = indata[field][casename]#[w_grid, kde_comp] = indata[casename]
ax.plot(w_grid, np.exp(kde_comp), color=contaminant_colors[field][casename], label=per_panel_contaminants[val])
# plt.title(field)
# ax.vlines(df['wfit_w_lowz'].loc[casename], 0, 40, color=contaminant_colors[field][casename], linestyle='--')
# print(df['wfit_w_lowz'].loc[casename])
ax.legend(fontsize=16, loc='upper left', bbox_to_anchor=(-0.025, 1.025))#, ncol=2)
ax.set_xlim(-1.25, -0.95)
ax.set_ylim(0., 40.)
ax.set_yticks([])
if i%3 == 0:
ax.set_ylabel(r'PDF ($w^{-1}$)', fontsize=18)
ax.set_yticks([10, 20, 30])
ax.set_yticklabels([10, 20, 30], fontsize=16)
ax.set_xlabel(r'$w$', fontsize=18)
ax.set_xticks([-1.2, -1.1, -1.])
ax.set_xticklabels([-1.2, -1.1, -1.], fontsize=16)
# lines_labels = [ax.get_legend_handles_labels() for ax in fig.axes]
# lines, labels = [sum(lol, []) for lol in zip(*lines_labels)]
# fig.legend(lines, labels)
# ax.set_xticklabels([])
# plt.savefig(file_extensions[field]+'dists_null.png')
fig.subplots_adjust(wspace=0., hspace=0.)
pylab.savefig(file_extensions[field]+'combos.png',format='png', bbox_inches='tight', pad_inches=0, dpi=250)
# fig.show()
```
TODO: polish these for paper
- linestyle for contamination rate if more than one with same contaminant per panel
| github_jupyter |
## Fundamental Spark Tools
Easiest way to make an RDD is to use the `parallelize` method
```
rdd = sc.parallelize([1,2,3,4])
rdd
```
So here we see that we have a special type of RDD - known as the Parallel Collection RDD!
RDD is very useful for dealing with failures when you are spreading jobs out to many clusters. The R in RDD means resilient - this means that all the data can be reconstructed on the fly should a node fail.
#### Reading a text file.
Spark supports reading text files from any Hadoop system like HDFS or a local file system.
Be sure that the file is accessible over the network or is in the same place for all of your worker nodes to access.
```
import os
text = sc.textFile(os.getcwd()+"/text_files/harvard_sentences.txt")
```
From the above, we see that we have a Map Partitions RDD!
```
text.collect()
```
Fun fact - the above sentences are known as [Harvard Sentences](https://en.wikipedia.org/wiki/Harvard_sentences)
`sc.textFile` has an optional argument `minPartitions=None`
This argument tells spark how many partitions our data should have.
You should note that the minPartitons is equivalent to the amount of parallelism that you will employ.
That is, `minPartitions = 1` means that only 1 node/executor will be processing your data. This doesnt exactly make sense when dealing with a cluster - you want to partition your data out amongst several nodes and then let each executor run your code in parallel on the data.
Furthermore, you are specifying the MINIMUM number of partitions, as such, this wioll serve as a the lower bound on your parallelism. Spark may decide to give you more, i.e. 5
Now, ifg you had 15 nodes - you may be tempted to set `minPartitions = 15` - indeed, this is exactly what you should do as you will ensure every node will get some data to work with.
However, in our incredibly simplistic example above, we only have 10 sentences, thus setting `minPartitions = 15` doesnt make much sense, as there is no way to split this data 15 ways. Spark is NOT going to split up a sentence.
Another thing to note is that when loading data from HDFS, spark will assign one partition per block - if I recall correctly, a block size is 64MB by default.
Spark docs recommend 2-4 partitions per CPU on your machine.
The other optional argument is `use_unicode=True` - if you are certain that your text data contains no unicode characters, then setting this parameter to False will make your data utf-8. This is a performance improvement, so be sure to use it where applicable.
#### Spark 'Actions'
You should think of 'actions' as poutput producing functions - they force spark to do a computation and spit out a result.
In terms of inputs and outputs, if a function takes in an RDD and spits out something that isnt an RDD - it is an action!
```
rdd = sc.parallelize(range(16))
rdd.collect()
```
`.collect()` pulls the result into the driver. By driver, I mean the application in which the result is being computed and displayed. In this case, that is a jupyter notebook. In other cases, it could be a spark shell.
```
rdd.sum()
rdd.take(4) # takes the first x elements
rdd.count() # equivalent of len(list(range(16)))
# numbers.saveAsTextFile("sample_numbers.txt") - saves to disk as .txt
```
#### Spark 'Transformations'
Transformations allow us to define computations on RDD's.
In terms of inputs and outputs, a function that takes an RDD and putputs an RDD is a transformation.
It is important to note that the output of one transformation can serve as the input for another transformation.
This way, we are building up a tree or **graph** of transformations that need to be applied to out initial RDD. No computation occurs whatsoever when specifying transformations - spark is **lazy**.
In fact, when you perform an action, spark applies a transformation to RDD's in a recursive fashion until it reaches an RDD that originates from an input source. This stop the recursion and allows all of the transformations to be made.
```
rdd = sc.parallelize(range(16))
small_rdd = sc.parallelize(range(10))
rdd.collect()
small_rdd.collect()
```
The 'heritage' of an RDD can be foundusing the `toDebugString()` method.
```
small_rdd.toDebugString()
def show_heritage(rdd):
for s in rdd.toDebugString().split(b'\n'): #it's a bytes object.
print(s.strip())
combo = rdd.union(small_rdd)
show_heritage(combo)
```
Combo is an RDD - however, when looking at its 'heritage' we see the information of `numbers` and `subset`.
#### Data Persistence
The general idea here is that when we do'expensive' transformations on RDD's, we may want to keep that RDD around - i.e. in 'memory'.
By default, spark stores the RDD in memory, persisting an RDD allows you to either fully store it in memory, fully store it on disk or use a combination of the two. Furthermore, we know that when we save to disk, the data is serialized - however, you can also serialize the data **in memory**. You can specify the level of storage for persistance.
Furthermore, you can set a _replication level_ for the persistance. The higher the replication, the higher 'insurance' you have on replicating all your data if you lose a node/executor. In addition, higher replciation means that more processes can be run on the data if the data in question is being used as a source.
As such, by persisting an RDD, any subsequent action on that partiuclar RDD **within the same session** will **not** have to recurse all the way back to an input RDD.
NOTE:
In Python, stored objects will always be serialized with the Pickle library, so it does not matter whether you choose a serialized level. The available storage levels in Python include MEMORY_ONLY, MEMORY_ONLY_2, MEMORY_AND_DISK, MEMORY_AND_DISK_2, DISK_ONLY, and DISK_ONLY_2 - see more [here](https://spark.apache.org/docs/2.2.0/rdd-programming-guide.html#rdd-persistence)
Also - once you have set a level of storage for an RDD, you cannot somply change the level, you must refresh the notebook, and set the new level!
```
import pyspark as pyk
big_numbers = sc.parallelize(range(1000000))
trans = big_numbers.map(lambda x : x ** 3)
# trans.saveAsTextFile("initial-trans.txt")
# trans.map(lambda x: x - 22000).saveAsTextFile("second-trans.txt")
```
The above is a classic example of why we would prefer to persist an RDD. the second transformation, would require us to do all the transformations again. However, if we persisted the RDD, the second transformation would occur much faster since it no longer needs to recurse to the input RDD.
```
# trans.persist(pyk.StorageLevel.MEMORY_AND_DISK) # this should occur before our first saveAsTextFile.
# trans.is_cached # will return True if RDD has been persisted
# trans.unpersist() # will unpersist after you are done with the RDD.
```
## Transformation Section
###### Map
Works exactly the same as the built-in python version of map.
Start with an RDD, map a function over that RDD!
When using map, you should expect one input (RDD) and expect one output (RDD). Map creates an output for EACH input - where an input is every element present in the RDD.
```
rdd = sc.parallelize(range(21))
rdd.map(lambda x: x * 2).collect()
```
Lambda syntax is quite common in PySpark, however, you can still define regular functions just like you would in Python.
```
def mult_2(val):
return 2 * val
rdd.map(mult_2).collect()
```
when writing traditional functions like this, it is important that this function is a **pure** function - that is, it doesnt alter/store/reference any states/variables etc outside of its scope.
This is incredibly important as we are doing distributed work across multiple executors. As such, we must be sure that given an input, our function will always give the same output. This is an identitcal concept to [**referential transparency**](https://wiki.haskell.org/Referential_transparency) employed by functional programming languages i.e. Haskell.
Map has an optional argument `preservesPartitioning=False`. In essence, if set to `True`, it will ensure that the mapping over your data does not change the way your data was paritioned. This is a more advanced spark feature, however, it is incredibly useful when doing joins! In general, if yu wish to preserve partition strucutre, check out the `mapValues` function instead of `map`.
`mapValues` requires a Key-Value pair, it preserves the keys!
```
x = sc.parallelize([("a", ["apple", "banana", "lemon"]), ("b", ["grapes"])])
def f(val):
return len(val)
x.mapValues(f).collect()
```
###### Filter
You can think of filter as being equivalent to a `WHERE` clause in SQL. It only keeps the values that are relevent in your RDD!
Again, each element in the RDD is passed through the filter. If the value passng through the function evaluates to True, the value is kept!
```
rdd = sc.parallelize(range(25))
def even(val):
return val % 2 == 0
rdd.filter(even).collect()
```
Now - given a large dataset, say taht we filter out a significant chunk of the data. It is wise at this point to use the `coalesce` function.
This function reduced the number of partitions on the resulting RDD in addition to **minimizing** network traffic.
You should think of network traffic as **computation overhead**. That is, if you had a really large dataset that required 1000 Nodes - but now after filtering, all your need is 10 nodes. It doesnt make sense to start all 1000 nodes everytime you start manipulating an RDD that requires 10 Nodes.
`coalesce` does this reduction for you! We will get to an example later!
###### FlatMap
Most beginners confuse this with `map` - dont do this!
At a high level, `map` is a **one-to-one** transformation.
`flatMap` is a **one-to-many** transformation!
Think of flatmap as taking one input at a time, and for each input, producing many outputs!
```
rdd = sc.textFile(os.getcwd()+"/text_files/harvard_sentences.txt")
rdd.flatMap(lambda x: x.split(" ")).collect()
rdd.map(lambda x: x.split(" ")).collect()
```
I hope you see the differnce! `flatMap` gives you access to all the words i all sentences directly! In essence, `flatMap` removes one level of grouping!
###### MapPartitions
Exactly the same as `map` however, it runs transformations over the partitions of an RDD and then aggregates thme together!
Below I will show you an example of counting all the word occurences in our harvard sentences.
```
rdd = sc.textFile(os.getcwd()+"/text_files/harvard_sentences.txt", minPartitions=7)
all_words = rdd.flatMap(lambda x: x.split(" "))
```
Below, the iterator_obj is all of the values in a specific partition in the RDD.
Since this behaves as an iterator object, we use the `yield` keyword and not `return`.
This is essential, because we want to update counts as we go through each element in a partition rather than just returning one large ditcionary with everything at the end.
An excellent explanation of Generators, `yield` and etc can be found [here](https://pythontips.com/2013/09/29/the-python-yield-keyword-explained/).
```
def generate_count(iterator_obj):
counts = {}
for word in iterator_obj:
if word not in counts:
counts[word] = 1
else:
counts[word] +=1
yield counts # we yield and NOT return here!
counts = all_words.mapPartitions(generate_count)
counts.collect()
counts.collect()
len(counts.collect()) # output partitioned!
```
If you wanted the index of the partition to be return in addition to the output, check out the `mapPartitionsWithIndex` function!
###### Sample
Returns a sample of your data.
This sample can then be used for statistical analysis with regards to the population etc..
```
rdd = sc.parallelize(range(99999))
rdd.count()
rdd.sample(False, 0.2).count() # 20% of data with No replacement.
```
###### Union
Allows you to combine RDD's - there is no remval of duplicates, sorting etc. It's simply a merging of the RDD's.
```
first = sc.parallelize(range(20))
second = sc.parallelize(range(30))
first.union(second).collect()
```
###### Intersection
Find elements that exist in both RDD's.
Note: Intersection can be slow on very large databases as internally, spark is running a `reduce` job acorss multiple nodes. This data shifting between nodes and reduction can cause quite the overhead. As such, if you job is every very slow, check to see if you have used an intersection - can you optimize any further before using `intersection`?
```
first = sc.parallelize([1,2,3,4,4,5])
second = sc.parallelize([5,17,20,4,1])
first.intersection(second).collect()
```
###### Distinct
Drops multiple duplciates from an RDD
I will employ the `cartesian` method - this gives the [cartesian product](https://en.wikipedia.org/wiki/Cartesian_product)
Recall that cartesian products scale with the size of your data = you should never be doing a cartesian product between 2 large data sets!
Rather, you should find a way to take the product between smaller subsets. You can then broadcast your operations to the large dataset using `map`.
If you must take the product of two large set - look into join, full outer join etc.
```
rdd = sc.parallelize(["Ibrahim", "Juan"]).cartesian(sc.parallelize(range(25)))
rdd.collect()
```
Now, suppose we needed to see how mnay unique names were in this list!
```
rdd.map(lambda x: x[0]).distinct().collect() # Awesome
```
`.distinct()` takes an optional argument `numPartitions=None` - the higher you set this, the more parallelized your code will run. Again, `distinct` uses `reduce` behind the scenes, as such, performance can be slow with large amounts of data. Optimize as much as you can before using it.
###### Pipe
This function takes each partition of data within an RDD and **pipes** it to a command line tool of your choice!
This is useful if you have already developed a command line tool in **ANY** language and now wish to parallelize it.
All data is fed in a strings and output as strings.
```
rdd = sc.parallelize(range(1,50))
rdd.pipe('grep 4').collect()
```
###### Coalesce
This is an incredibly useful function - however, in order to really reap the benefits we must first dig a little deeper into how spark works under the hood.
By now, you should know that Spark stores your data in a distrbuted manner. This means that data is split into chunks where each chunk is known as a **partitions**.Partitions exist throughout your entire cluster.
In essence, the the coalesce functions allows you to **reduce** the number of partitions you have of your data in a **significantly more efficient** manner over doing a full repartition.
How? Coalesce combines paritions that are **already on the same executors**. This minimizes network traffic **between** executors!
This is all well and good, however, when do I need to change the number of paritions? What is the appropriate number of partitions?
I like to think that choosing the optimal number of partitions is a competition between 2 important considerations:
- The number of partitions is the **upper limit** for parallelism.
- This means its **impossible** to have 8 processors working on 3 partitions.Spark docs [recommend 2-4 tasks](https://spark.apache.org/docs/latest/tuning.html#level-of-parallelism) (aka partitions) per CPU in your cluster.
- Too many partions will also cause excessive network traffic due to the creation of many small tasks.
- Number of partitions determines number of output files for an action.
I would suggest using the Spark docs suggestions of 2-4 partitions/CPu as a rule of thumb.
However, I cannot stress enough the importance of experimentation when it comes to tuning your settings.
After running a job, cut the number of partitions in half and see what happens? Increase the number of partitions slightly, did it get faster?
As we see in Machine Learning, most (Hyper)parameter tuning starts with an educated guess and then embarks on a series of iterations until we find that sweet spot.
```
rdd = sc.parallelize(range(99999), numSlices=1000) #original with 1000 partitions
reducedRDD = rdd.coalesce(30) #same rdd, but with 30 partitons
```
###### Repartition.
A very useful function for when you want to increase the number of partions that your RDD started with. This can cause some overhead as new partitions are created and sent to nodes.
If reducing the number of partitions - its better to use `coalesce` as this will minimize network overhead by merging together partitions that are already on the same node.
Partitioning data is central to spark - this is something that just takes practice and some experimenting!
Recall, 2-4 partions per CPU is a good rule of thumb to start at!
```
rdd = sc.parallelize(range(999), numSlices=1)
rdd.repartition(30) # simple as that
```
###### RepartitionAndSortWithinPartitions
Same as the above function, but will also allow you to sort each partion there and then.
This funciton works on key-value pairs and partitions the data according to key. This function is more efficent than just repartitioning, and then later on sorting the data.
Learn about `.glom()` [here](https://spark.apache.org/docs/0.7.2/api/pyspark/pyspark.rdd.RDD-class.html#glom).
```
rdd = sc.parallelize([[3,41], [3,7], [100,100], [0.5, 17]])
rdd.repartitionAndSortWithinPartitions(2).glom().collect()
```
The outermost list containing everything is due to glom.
Then we have two inner lists - this is because we repartitioned into 2. Each of those sublists is also sorted!
Notie how the two 3 appears together - this means our sorting falls within our partioning!
You can also specify a custom `partitionFunc` in the optional arguments.
There is also the optional `ascending` and `keyFunc` arguments
```
rdd.repartitionAndSortWithinPartitions(2, partitionFunc=lambda x: x == 100).glom().collect()
```
Above, I specified that I want one partion to contain all keys that equal 100.
## Actions Section
###### Reduce
Calculates aggregates **over many** inputs!
Requires an **associative** and **commutative** function that is applied to pairs of inputs. The function is then applied between pairs and so on until we have a single output.
```
rdd = sc.parallelize(range(25), numSlices=3)
rdd.glom().collect()
rdd.reduce(max)
```
Spark is running the reduce command in parallel over every partition - think of this as asking for the max of the first sub list and so on.
Then taking those results and asking for the max in that secondary comparison. Eventually, the answer **reduces** to 24.
`reduceByKey` allows you to calculate aggregates based on subsets of data - like a pandas groupby!
```
rdd.reduce(lambda x,y : x + y) # equivalent to sum(list(range(25)))
```
###### Count
This function returns the number of elements in the RDD.
More intersting are 2 other functions that are closely related:
- `countApprox(timeout=500, confidence = 0.7)`
The above function will use the [HyperLogLog](https://en.wikipedia.org/wiki/HyperLogLog) algorithm to approximate the size of your data. This come in very handy when you have very large data and dont want to wait for an accurate count.
If you wanted to count the distinct elements you could use:
- `countApproxDistinct`
refer to documentation for more info on optional arguments!
###### First
Pulls the first element from your RDD. Nothing special here!
```
rdd = sc.parallelize(range(1,6,1))
rdd.first()
```
###### Take
Take # of elements from an RDD and returns a list
```
rdd = sc.parallelize(range(1,6,1))
rdd.take(3)
```
###### TakeSample
Pulls a random sample of elements of a given size from the RDD.
```
rdd = sc.parallelize(range(50))
rdd.takeSample(withReplacement=False, num=8)
```
###### TakeOrdered
Sorts the RDD, and then take a # of elements.
Really fast when N is small - if you want to sort the entire RDD, just sort, no point in doing `takeOrdered`.
###### saveAsTextFile
number of partitions will equal the number of output files.
You should repartition before running `saveAsTextFile` in order to control number of output files!
You can also use compression codecs as optional entries! Excellent for saving space on disk.
In python - you can use `saveAsPickleFile`
```
rdd = sc.parallelize(range(9999), numSlices=7)
# rdd.saveAsTextFile("put_output_folder_name_here")
```
###### CountByKey
Counts the occurence of Keys in an RDD.
There is also a `countByValue` function.
```
rdd = sc.parallelize([('Andrew', 45), ('Juan', 99), ('Mauricio', 12), ('Mauricio', 1)])
rdd.countByKey()
```
###### ForEach
Takes an action for each element of an RDD!
Be sure to use reduce or an accumulator if you wish to see the results reflected in the driver.
By default, sparks nodes do not push back variable changes to the driver.
it's very important to distinguish between the local and cluster environments.
Read this very important part of the documentation [here](https://spark.apache.org/docs/2.1.1/programming-guide.html#understanding-closures-a-nameclosureslinka).
## Conclusion
That's all I have for today. Plenty of information for you to play with and get your hands dirty!!
The next notebook will cover more on key-value pairs, IO actions in addition to performance boosters! Stay tuned.
As always, feel free to reach out: igabr@uchicago.edu or [@Gabr\_Ibrahim](https://twitter.com/Gabr_Ibrahim).
| github_jupyter |
```
%matplotlib inline
import os
import sys
module_path = os.path.abspath(os.path.join('..','scatteract','tensorbox'))
if module_path not in sys.path:
sys.path.append(module_path)
utils_path = os.path.abspath(os.path.join('..','scatteract','tensorbox','utils'))
if utils_path not in sys.path:
sys.path.append(utils_path)
print(utils_path)
import matplotlib.pyplot as plt
import scatter_extract
from tensorbox.utils.annolist import AnnotationLib as al
import cv2
import numpy as np
from IPython.display import display, HTML
```
### Initializating the object dection models:
```
model_dict = {"points":"/Users/nils/CC/Data/mpl10k_out/lstm_rezoom_points_2018_09_28_17.10"}
debbie_dir = "/Users/nils/CC/scatteract/data/debbie"
iteration = 590000
plt_xtr = scatter_extract.PlotExtractor(model_dict, iteration)
pred_dict = {}
for key in plt_xtr.models:
pred_dict[key] = plt_xtr.test_pts(debbie_dir, debbie_dir, debbie_dir)
plt_xtr.my_dir_dict
model_dict = {'ticks':"./output/ticks_v1", "labels":"./output/labels_v1","points":"./output/points_v1"}
iteration = 125000
plt_xtr = scatter_extract.PlotExtractor(model_dict, iteration)
```
### Parsing idl files containing ground truth
```
true_idl_dict = {'ticks':"./data/plot_test/ticks.idl",
"labels":"./data/plot_test/labels.idl","points":"./data/plot_test/points.idl"}
true_annos_dict = {key:al.parse(true_idl_dict[key]) for key in true_idl_dict}
#For the purpose of this notebook we only focus on the first 10 images
true_annos_dict = {key: true_annos_dict[key][0:10] for key in true_annos_dict}
```
### Getting prediction bounding boxes for the relevant plot objects
```
pred_dict = {}
for key in plt_xtr.models:
pred_dict[key] = plt_xtr.predict_model(plt_xtr.models[key], plt_xtr.H[key], true_annos_dict[key],
"./data/plot_test/", 0.3)
```
### Drawing images with the true and predicted bounding boxes
```
for j in range(len(true_annos_dict["ticks"])):
img = plt_xtr.open_image("./data/plot_test/" + true_annos_dict["ticks"][j].imageName)
img_true = np.copy(img)
img_pred = np.copy(img)
for key in true_annos_dict:
for rect_true in true_annos_dict[key][j].rects:
cv2.rectangle(img_true, (int(rect_true.x1),int(rect_true.y1)),
(int(rect_true.x2),int(rect_true.y2)),
plt_xtr.color_dict[key], 2)
for rect_pred in pred_dict[key][j].rects:
cv2.rectangle(img_pred, (int(rect_pred.x1),int(rect_pred.y1)),
(int(rect_pred.x2),int(rect_pred.y2)),
plt_xtr.color_dict[key], 2)
print true_annos_dict["ticks"][j].imageName
fig = plt.figure(figsize=(20,20))
ax1 = fig.add_subplot(2,2,1)
ax1.imshow(img_true)
ax1.axis("off")
ax1.set_title("Truth", fontsize=20)
ax2 = fig.add_subplot(2,2,2)
ax2.imshow(img_pred)
ax2.axis("off")
ax2.set_title("Predictions", fontsize=20)
plt.show()
```
### Applying OCR on the tick values, getting the closest tick mark for each tick value, splitting the X and Y axis, applying RANSAC regression to find the conversion factors and getting points in chart coordinates
```
pred_dict['labels'] = plt_xtr.get_ocr(pred_dict['labels'], "./data/plot_test/")
pred_dict['labels'] = plt_xtr.get_closest_ticks(pred_dict['labels'], pred_dict['ticks'])
pred_labels_X, pred_labels_Y = plt_xtr.split_labels_XY(pred_dict['labels'])
regressor_x_dict = plt_xtr.get_conversion(pred_labels_X, cat='x')
regressor_y_dict = plt_xtr.get_conversion(pred_labels_Y, cat='y')
df_dict_pred = plt_xtr.predict_points(pred_dict['points'], regressor_x_dict, regressor_y_dict)
```
### Showing plots and some of the points found in the coordinate system of the chart
```
for j in range(len(true_annos_dict["ticks"])):
key = true_annos_dict["ticks"][j].imageName
plot_img = plt_xtr.open_image("./data/plot_test/" + key)
fig = plt.figure(figsize=(10,10))
ax1 = fig.add_subplot(1,1,1)
ax1.imshow(plot_img)
ax1.axis("off")
ax1.set_title("Plot - {}".format(key), fontsize=20)
plt.show()
print "First 10 points detected:"
df_pred_display = df_dict_pred[key].copy()
df_pred_display.columns = [r'$X_{\text{pred}}$', r'$Y_{\text{pred}}$']
display(HTML(df_pred_display.head(10).to_html(index=False)))
```
### Evaluting the performance systematically
```
df_prec_rec = plt_xtr.get_metrics(df_dict_pred, "./data/plot_test/coords.idl", max_dist_perc = 2.0)
df_prec_rec.set_index('image_name').sort_index()
```
| github_jupyter |
### Set up
#### 1. Set up accounts and role
```
import sagemaker
import boto3
from uuid import uuid4
sagemaker_session = sagemaker.Session()
account_id = boto3.client('sts').get_caller_identity().get('Account')
region = boto3.session.Session().region_name
#role = sagemaker.get_execution_role()
role="arn:aws:iam::{}:role/service-role/AmazonSageMaker-ExecutionRole-20190118T115449".format(account_id)
max_runs=1
```
#### 2. Setup image and instance type
```
# pytorch_custom_image_name="ppi-extractor:gpu-1.0.0-201910130520"
instance_type = "ml.p3.2xlarge"
instance_type_gpu_map = {"ml.p3.8xlarge":4, "ml.p3.2xlarge": 1, "ml.p3.16xlarge":8}
# docker_repo = "{}.dkr.ecr.{}.amazonaws.com/{}".format(account_id, region, pytorch_custom_image_name)
```
#### 3. Configure train/ test and validation datasets
```
bucket = "aegovan-data"
pretrained_bert="s3://{}/embeddings/bert/".format(bucket)
pretrained_bert_aimed = "s3://{}/embeddings/bert_aimed/bert_lowestloss/".format(bucket)
trainfile = "s3://{}/processed_dataset/train_multiclass.json".format(bucket)
testfile= "s3://{}/processed_dataset/test_multiclass.json".format(bucket)
valfile="s3://{}/processed_dataset/val_multiclass.json".format(bucket)
s3_output_path= "s3://{}/ppi_multiclass_sagemakerresults/".format(bucket)
s3_code_path= "s3://{}/ppi_bert_code".format(bucket)
s3_checkpoint = "s3://{}/ppi_multiclass_bert_checkpoint/{}".format(bucket, str(uuid4()))
```
### Start training
```
commit_id = "6df30be45e08af56a0f10fbfc8a724737f7ca9e1"
train_inputs = {
"train" : trainfile,
"val" : valfile,
"PRETRAINED_MODEL" : pretrained_bert
}
train_inputs_pretrained_aimed = {
"train" : trainfile,
"val" : valfile,
"PRETRAINED_MODEL" : pretrained_bert_aimed
}
sm_localcheckpoint_dir="/opt/ml/checkpoints/"
BertNetworkFactoryhyperparameters = {
"datasetfactory":"datasets.ppi_multiclass_dataset_factory.PpiMulticlassDatasetFactory",
"modelfactory" :"models.bert_model_factory.BertModelFactory",
"tokenisor_lower_case":0,
"uselosseval":1,
"batch": "8" * instance_type_gpu_map[instance_type],
"gradientaccumulationsteps" : "8",
# "protein_name_replacer_random_seed":42,
"epochs" : "500",
"log-level" : "INFO",
"learningrate":.00001,
"earlystoppingpatience":50,
"checkpointdir" : sm_localcheckpoint_dir,
# Checkpoints once every n epochs
"checkpointfreq": 2,
"weight_decay":0.01,
"commit_id" : commit_id
}
BertNetworkFactoryhyperparameters_max_f1 = BertNetworkFactoryhyperparameters.copy()
BertNetworkFactoryhyperparameters_max_f1["uselosseval"] = 0
BertNetworkFactoryhyperparameters_max_f1_aimed = BertNetworkFactoryhyperparameters.copy()
BertNetworkFactoryhyperparameters_max_f1_aimed["uselosseval"] = 0
BertNetworkFactoryhyperparameters_max_f1_aimed["weight_decay"] = 0.001
metric_definitions = [{"Name": "TrainLoss",
"Regex": "###score: train_loss### (\d*[.]?\d*)"}
,{"Name": "ValidationLoss",
"Regex": "###score: val_loss### (\d*[.]?\d*)"}
,{"Name": "TrainAucScore",
"Regex": "###score: train_ResultScorerAucMacro_score### (\d*[.]?\d*)"}
,{"Name": "ValidationAucScore",
"Regex": "###score: val_ResultScorerAucMacro_score### (\d*[.]?\d*)"}
,{"Name": "TrainF1MacroScore",
"Regex": "###score: train_ResultScorerF1Macro_score### (\d*[.]?\d*)"}
,{"Name": "ValidationF1MacroScore",
"Regex": "###score: val_ResultScorerF1Macro_score### (\d*[.]?\d*)"}
]
!git log -1 | head -1
!git log -1 | head -5 | tail -1
# set True if you need spot instance
use_spot = True
train_max_run_secs = 5 *24 * 60 * 60
spot_wait_sec = 5 * 60
max_wait_time_secs = train_max_run_secs + spot_wait_sec
if not use_spot:
max_wait_time_secs = None
# During local mode, no spot.., use smaller dataset
if instance_type == 'local':
use_spot = False
max_wait_time_secs = 0
wait = True
# Use smaller dataset to run locally
inputs = inputs_sample
experiments = {
"ppimulticlass-bert" : {
"hp" :BertNetworkFactoryhyperparameters,
"inputs" : train_inputs
},
"ppimulticlass-bert-f1" : {
"hp" :BertNetworkFactoryhyperparameters_max_f1,
"inputs" : train_inputs
},
"ppimulticlass-bert-aimed-f1" : {
"hp" :BertNetworkFactoryhyperparameters_max_f1_aimed,
"inputs" : train_inputs_pretrained_aimed
}
}
experiment_name = "ppimulticlass-bert-aimed-f1"
hyperparameters = experiments[experiment_name]["hp"]
inputs = experiments[experiment_name]["inputs"]
base_name = experiment_name
hyperparameters
git_config = {'repo': 'https://github.com/elangovana/ppi-aimed.git',
'branch': 'main',
'commit': hyperparameters["commit_id"]
}
hyperparameters
inputs
from sagemaker.pytorch import PyTorch
estimator = PyTorch(
entry_point='main_train_pipeline.py',
source_dir = 'src',
dependencies =['src/datasets', 'src/models','src/utils', 'src/scorers'],
git_config= git_config,
# image_name= docker_repo,
role=role,
framework_version ="1.4.0",
py_version='py3',
instance_count=1,
instance_type=instance_type,
hyperparameters = hyperparameters,
output_path=s3_output_path,
metric_definitions=metric_definitions,
volume_size=30,
code_location=s3_code_path,
debugger_hook_config=False,
base_job_name =base_name,
use_spot_instances = use_spot,
max_run = train_max_run_secs,
max_wait = max_wait_time_secs,
checkpoint_s3_uri=s3_checkpoint,
checkpoint_local_path=sm_localcheckpoint_dir)
estimator.fit(inputs, wait=False)
```
| github_jupyter |
```
import requests, os, bs4, re
from IPython.display import Image
# Starting url
url = 'http://xkcd.com'
# Store comics in ./xkcd
# makedirs() ensures the folder exists
# exist_ok=True prevents function from throwing exception if folder already exists
# Python3: os.makedirs('xkcd', exist_ok=True)
os.makedirs('xkcd' ,exist_ok = True)
last_comic = 1732 - int(input("Enter number of comics: "))
# End loop when url ends with '#'
while not url.endswith(str(last_comic)+'/'):
# Download the page
print ('Downloading page %s...'% url)
res = requests.get(url)
res.raise_for_status()
soup = bs4.BeautifulSoup(res.text, 'html.parser')
# Find the URL of the comic image
# <div id="comic"><img src=""></div>
comicElem = soup.select('#comic img')
if comicElem == []:
print ('Could not find comic image.')
else:
# Create regex to parse url below the comic (remember to escape the parentheses)
pattern = re.compile(r'Image URL \(for hotlinking/embedding\): (.*)')
# Pass pattern to BeautifulSoup's find() method
comicUrl = soup.find(text=pattern)
# Substitute with r'\1', the first group in parentheses: (.*)
# Use strip to remove whitespace
comicUrl = pattern.sub(r'\1', comicUrl).strip()
# Download the image
print ('Downloading the image %s...'% comicUrl)
Image(url= comicUrl)
res = requests.get(comicUrl)
res.raise_for_status()
# Save the image to ./xkcd
# comicUrl example: http://imgs.xkcd.com/comics/heartbleed_explanation.png
# call os.path.basename() on it to return the last part of the url:
# heartbleed_explanation.png (can use this as the filename of image when saving)
# join() so program uses backslashes on Windows and forward clashes on OS X and Linux
# call open() to open a new file in 'wb' "write binary" mode
# to save downloaded files using Requests, loop over return value of the iter_content()
# for loop writes chunks of the image data (max 100,000 bytes each) to the file
# then close the file, the image is now saved your hard drive
imageFile = open(os.path.join('xkcd', os.path.basename(comicUrl)), 'wb')
for chunk in res.iter_content(100000):
imageFile.write(chunk)
imageFile.close()
# Get the Prev button's url
# while loop begins the entire download process again for this comic
prevLink = soup.select('a[rel="prev"]')[0]
url = 'http://xkcd.com' + prevLink.get('href')
print ('Done.')
```
| github_jupyter |
Para entrar no modo apresentação, execute a seguinte célula e pressione `-`
```
%reload_ext slide
```
<span class="notebook-slide-start"/>
# Widgets
Este notebook apresenta os seguintes tópicos
- [Widgets interativos](#Widgets-Interativos)
- [Exercício 14](#Exerc%C3%ADcio-14)
- [Widgets completos](#Widgets-completos)
- [Exercício 15](#Exerc%C3%ADcio-15)
- [Exercício 16](#Exerc%C3%ADcio-16)
## Widgets Interativos
Finalmente, uma outra forma de interagir com o Jupyter é a partir de widgets interativos. Esses widgets podem ser usados para fazer formulários, dashboards e até mesmo variar rapidamente parâmetros de funções.
O widget a seguir interage com a função `fib`, definida no início deste notebook para variar os parâmetros dela. <span class="notebook-slide-extra" data-count="1"/>
```
from ipywidgets import interact
def fib(x):
if x <= 1:
return x
return fib(x - 1) + fib(x - 2)
interact(fib, x=(1, 30))
```
Essa função também pode ser usada como um decorador. <span class="notebook-slide-extra" data-count="1"/>
```
@interact
def add(x=1, y=2, template="A soma de {x} com {y} resulta em {z}"):
z = x + y
print(template.format(x=x, y=y, z=z))
```
Podemos definir o intervalo dos valores. <span class="notebook-slide-extra" data-count="1"/>
```
@interact(x=(0, 100), y=(0, 100))
def add(x=1, y=2, template="A soma de {x} com {y} resulta em {z}"):
z = x + y
print(template.format(x=x, y=y, z=z))
```
O widget também pode ser usado com visualizações ricas. <span class="notebook-slide-extra" data-count="1"/>
```
import matplotlib.pyplot as plt
%matplotlib inline
@interact(count=(1, 35))
def generate_plot(count=15):
x = range(count)
y = [fib(n) for n in x]
plt.plot(x, y)
```
Para números grandes, o uso do `interact` com a nossa implementação de fibonacci começou a não ser tão interativa.
Em funções de longa duração, a atualização automatica do `interact` pode atrapalhar mais do que ajudar. Para resolver isso, podemos usar o `interact_manual`. <span class="notebook-slide-extra" data-count="1"/>
```
from ipywidgets import interact_manual
@interact_manual(count=(1, 35))
def generate_plot(count=15):
x = range(count)
y = [fib(n) for n in x]
plt.plot(x, y)
```
## Exercício 14
Implemente uma função interativa que permita escolher um arquivo de código fonte usando um drop-down e imprima a quantidade de letras do arquivo após a escolha.
Dica: ao passar uma lista ou dicionário para o `interact`, é criado um elemento drop-down.
```
...
```
## Widgets completos
O `interact` é uma simplificação do sistema de widgets para facilitar o uso em funções. Porém, quando estamos criando dashboards ou formulários mais completos, podemos usar o sistema mais completo.
A seguir, criaremos um slider que não depende de nenhuma função `interact`. <span class="notebook-slide-extra" data-count="1"/>
```
from ipywidgets import IntSlider
slider = IntSlider(
value=7,
min=0,
max=10,
step=1,
description='Test:'
)
slider
```
<span class="notebook-slide-extra" data-count="1"/>
<span class="notebook-slide-scroll" data-position="-1"/>
Podemos acessar o valor do slider através do atributo `.value`.
```
slider.value
```
Se quisermos ter o efeito do `interact` de executar alguma função ao alterar o slider, podemos definir funções de observação. <span class="notebook-slide-extra" data-count="1"/>
```
def add1(change):
if change.name == "value":
print(change.new + 1)
slider.observe(add1)
slider
```
<span class="notebook-slide-scroll" data-position="-1"/>
Note que verificamos o tipo da observação ao receber a mudança. Algumas mudanças no widget não ocorrem no valor e isso acaba mudando o resultado de `change.new`. Além de `change.new` e `change.name`, podemos acessar outros atributos de `change`, como `change.old`.
Note também que ao fazermos `print` nessa função, os outputs anteriores foram preservados.
Se quisermos ter um controle maior do output, podemos usar um widget específico de output. <span class="notebook-slide-extra" data-count="1"/>
```
from ipywidgets import Output
out = Output()
with out:
print("Dentro do output novo")
print("Fora do output novo")
out
```
<span class="notebook-slide-scroll" data-position="-1"/>
<span class="notebook-slide-extra" data-count="1"/>
Podemos usar o objeto de output para apagar o conteúdo.
```
out.clear_output()
```
Agora vamos combinar o slider com o output para gerar o efeito do `interact`.
Primeiro, precisamos limpar todos os eventos de observação que registramos no slider. <span class="notebook-slide-extra" data-count="1"/>
```
slider.unobserve_all()
```
Em seguida, podemos criar um novo evento que imprima dentro do objeto de output. <span class="notebook-slide-extra" data-count="1"/>
```
def add1(change):
if change.name == "value":
out.clear_output()
with out:
print(change.new + 1)
slider.observe(add1)
```
Por fim, criamos um widget que combine os dois no mesmo lugar usando `VBox`. <span class="notebook-slide-extra" data-count="1"/>
```
from ipywidgets import VBox
VBox([slider, out])
```
## Exercício 15
Use o widget `Button` para simular o `interact_manual`. Esse widget possui um método `on_click` para definir funções de callback. <span class="notebook-slide-extra" data-count="1"/>
```
from ipywidgets import Button
button = Button(description="run")
...
```
## Exercício 16
Faça um widget que imprima a soma acumulada de todas as suas execuções.
O objetivo desse exercício é pensar em como criar widgets com estados que continuem existindo além de uma execução da função observadora ou do interact. <span class="notebook-slide-extra" data-count="2"/>
```
class AccWidget:
...
widget = AccWidget()
widget.view
```
Chegamos ao fim da apresentação principal do minicurso, mas não é o fim do conteúdo preparado. Temos mais dois notebooks que listam e explicam as magics do IPython e os ipywidgets disponíveis:
- [Extra/Lista.Magics.ipynb](Extra/Lista.Magics.ipynb)
- [Extra/Lista.Widgets.ipynb](Extra/Lista.Widgets.ipynb)
| github_jupyter |
# Collecting temperature data from an API
## About the data
In this notebook, we will be collecting daily temperature data from the [National Centers for Environmental Information (NCEI) API](https://www.ncdc.noaa.gov/cdo-web/webservices/v2). We will use the Global Historical Climatology Network - Daily (GHCND) data set; see the documentation [here](https://www1.ncdc.noaa.gov/pub/data/cdo/documentation/GHCND_documentation.pdf).
*Note: The NCEI is part of the National Oceanic and Atmospheric Administration (NOAA) and, as you can see from the URL for the API, this resource was created when the NCEI was called the NCDC. Should the URL for this resource change in the future, you can search for the NCEI weather API to find the updated one.*
## Using the NCEI API
Paste your token below.
```
import requests
def make_request(endpoint, payload=None):
"""
Make a request to a specific endpoint on the weather API
passing headers and optional payload.
Parameters:
- endpoint: The endpoint of the API you want to
make a GET request to.
- payload: A dictionary of data to pass along
with the request.
Returns:
Response object.
"""
return requests.get(
f'https://www.ncdc.noaa.gov/cdo-web/api/v2/{endpoint}',
headers={
'token': 'PASTE_YOUR_TOKEN_HERE'
},
params=payload
)
```
## See what datasets are available
We can make requests to the `datasets` endpoint to see what datasets are available. We also pass in a dictionary for the payload to get datsets that have data after the start date of October 1, 2018.
```
# see what datasets are available
response = make_request('datasets', {'startdate':'2018-10-01'})
response.status_code
```
Status code of `200` means everything is OK. More codes can be found [here](https://en.wikipedia.org/wiki/List_of_HTTP_status_codes).
### Get the keys of the result
The result is a JSON object which we can access with the `json()` method of our `Response` object. JSON objects can be treated like dictionaries, so we can access the `keys()` just like we would a dictionary:
```
response.json().keys()
```
The `metadata` of the JSON response will tell us information about the request and data we got back:
```
response.json()['metadata']
```
### Figure out what data is in the result
The `results` key contains the data we requested. This is a list of what would be rows in our dataframe. Each entry in the list is a dictionary, so we can look at the keys to get the fields:
```
response.json()['results'][0].keys()
```
### Parse the result
We don't want all those fields, so we will use a list comphrension to take only the `id` and `name` fields out:
```
[(data['id'], data['name']) for data in response.json()['results']]
```
## Figure out which data category we want
The `GHCND` data containing daily summaries is what we want. Now we need to make another request to figure out which data categories we want to collect. This is the `datacategories` endpoint. We have to pass the `datasetid` for `GHCND` as the payload so the API knows which dataset we are asking about:
```
# get data category id
response = make_request(
'datacategories',
payload={
'datasetid' : 'GHCND'
}
)
response.status_code
```
Since we know the API gives us a `metadata` and a `results` key in each response, we can see what is in the `results` portion of the JSON response:
```
response.json()['results']
```
## Grab the data type ID for the Temperature category
We will be working with temperatures, so we want the `TEMP` data category. Now, we need to find the `datatypes` to collect. For this, we use the `datatypes` endpoint and provide the `datacategoryid` which was `TEMP`. We also specify a limit for the number of `datatypes` to return with the payload. If there are more than this we can make another request later, but for now, we just want to pick a few out:
```
# get data type id
response = make_request(
'datatypes',
payload={
'datacategoryid' : 'TEMP',
'limit' : 100
}
)
response.status_code
```
We can grab the `id` and `name` fields for each of the entries in the `results` portion of the data. The fields we are interested in are at the bottom:
```
[(datatype['id'], datatype['name']) for datatype in response.json()['results']][-5:] # look at the last 5
```
## Determine which Location Category we want
Now that we know which `datatypes` we will be collecting, we need to find the location to use. First, we need to figure out the location category. This is obtained from the `locationcategories` endpoint by passing the `datasetid`:
```
# get location category id
response = make_request(
'locationcategories',
{
'datasetid' : 'GHCND'
}
)
response.status_code
```
We can use `pprint` to print dictionaries in an easier-to-read format. After doing so, we can see there are 12 different location categories, but we are only interested in `CITY`:
```
import pprint
pprint.pprint(response.json())
```
## Get NYC Location ID
In order to find the location ID for New York, we need to search through all the cities available. Since we can ask the API to return the cities sorted, we can use binary search to find New York quickly without having to make many requests or request lots of data at once. The following function makes the first request to see how big the list of cities is and looks at the first value. From there it decides if it needs to move towards the beginning or end of the list by comparing the city we are looking for to others alphabetically. Each time it makes a request it can rule out half of the remaining data to search.
```
def get_item(name, what, endpoint, start=1, end=None):
"""
Grab the JSON payload for a given field by name using binary search.
Parameters:
- name: The item to look for.
- what: Dictionary specifying what the item in `name` is.
- endpoint: Where to look for the item.
- start: The position to start at. We don't need to touch this, but the
function will manipulate this with recursion.
- end: The last position of the cities. Used to find the midpoint, but
like `start` this is not something we need to worry about.
Returns:
Dictionary of the information for the item if found otherwise
an empty dictionary.
"""
# find the midpoint which we use to cut the data in half each time
mid = (start + (end if end else 1)) // 2
# lowercase the name so this is not case-sensitive
name = name.lower()
# define the payload we will send with each request
payload = {
'datasetid' : 'GHCND',
'sortfield' : 'name',
'offset' : mid, # we will change the offset each time
'limit' : 1 # we only want one value back
}
# make our request adding any additional filter parameters from `what`
response = make_request(endpoint, {**payload, **what})
if response.ok:
# if response is ok, grab the end index from the response metadata the first time through
end = end if end else response.json()['metadata']['resultset']['count']
# grab the lowercase version of the current name
current_name = response.json()['results'][0]['name'].lower()
# if what we are searching for is in the current name, we have found our item
if name in current_name:
return response.json()['results'][0] # return the found item
else:
if start >= end:
# if our start index is greater than or equal to our end, we couldn't find it
return {}
elif name < current_name:
# our name comes before the current name in the alphabet, so we search further to the left
return get_item(name, what, endpoint, start, mid - 1)
elif name > current_name:
# our name comes after the current name in the alphabet, so we search further to the right
return get_item(name, what, endpoint, mid + 1, end)
else:
# response wasn't ok, use code to determine why
print(f'Response not OK, status: {response.status_code}')
def get_location(name):
"""
Grab the JSON payload for the location by name using binary search.
Parameters:
- name: The city to look for.
Returns:
Dictionary of the information for the city if found otherwise
an empty dictionary.
"""
return get_item(name, {'locationcategoryid' : 'CITY'}, 'locations')
```
When we use binary search to find New York, we find it in just 8 requests despite it being close to the middle of 1,983 entries:
```
# get NYC id
nyc = get_location('New York')
nyc
```
## Get the station ID for Central Park
The most granular data is found at the station level:
```
central_park = get_item('NY City Central Park', {'locationid' : nyc['id']}, 'stations')
central_park
```
## Request the temperature data
Finally, we have everything we need to make our request for the New York temperature data. For this we use the `data` endpoint and provide all the parameters we picked up throughout our exploration of the API:
```
# get NYC daily summaries data
response = make_request(
'data',
{
'datasetid' : 'GHCND',
'stationid' : central_park['id'],
'locationid' : nyc['id'],
'startdate' : '2018-10-01',
'enddate' : '2018-10-31',
'datatypeid' : ['TMIN', 'TMAX', 'TOBS'], # temperature at time of observation, min, and max
'units' : 'metric',
'limit' : 1000
}
)
response.status_code
```
## Create a DataFrame
The Central Park station only has the daily minimum and maximum temperatures.
```
import pandas as pd
df = pd.DataFrame(response.json()['results'])
df.head()
```
We didn't get TOBS because the station doesn't measure that:
```
df.datatype.unique()
```
Despite showing up in the data as measuring it... Real-world data is dirty!
```
if get_item(
'NY City Central Park', {'locationid' : nyc['id'], 'datatypeid': 'TOBS'}, 'stations'
):
print('Found!')
```
## Using a different station
Let's use LaGuardia airport instead. It contains `TAVG` (average daily temperature):
```
laguardia = get_item(
'LaGuardia', {'locationid' : nyc['id']}, 'stations'
)
laguardia
```
We make our request using the LaGuardia airport station this time and ask for `TAVG` instead of `TOBS`.
```
# get NYC daily summaries data
response = make_request(
'data',
{
'datasetid' : 'GHCND',
'stationid' : laguardia['id'],
'locationid' : nyc['id'],
'startdate' : '2018-10-01',
'enddate' : '2018-10-31',
'datatypeid' : ['TMIN', 'TMAX', 'TAVG'], # temperature at time of observation, min, and max
'units' : 'metric',
'limit' : 1000
}
)
response.status_code
```
The request was successful, so let's make a dataframe:
```
df = pd.DataFrame(response.json()['results'])
df.head()
```
We should check we got what we wanted: 31 entries for TAVG, TMAX, and TMIN (1 per day):
```
df.datatype.value_counts()
```
Write the data to a CSV file for use in other notebooks.
```
df.to_csv('data/nyc_temperatures.csv', index=False)
```
| github_jupyter |
# Matrix Formalism of the Newton-Euler equations
Renato Naville Watanabe
In this notebook will be shown two examples of how to use a matrix formalism to perform inverse dynamics analysis. It does not consist a comprehensive treatise about the subject. It is rather an introduction based on examples. Nevertheless, the reader of this notebook will have sufficient knowledge to read recent texts on biomechanics and other multibody dynamic analysis.
## Inverse dynamics
For the inverse dynamics analysis, we will obtain the joint torques and forces, from the joint kinematics and external forces.
<figure><img src="../images/inv.png" width=600 /> <figcaption><i><center>Adapted from Erdemir et al. (2007) </center></i></figcaption>
As an example, we will consider the problem of estimating the forces and torques in the ankle and knee joints during the gait, considering a 3D movement. At this point, we consider that the accelerations, angular velocities, angular accelerations, masses, moments of inertia and rotation matrices necessary to compute the forces and moments are known.
The free-body diagram of the gait, considering a 3D movement is very similar [to the 2D case](GaitAnalysis2D.ipynb). The equations of forces and moments are described by the Newton-Euler equations (for a revision on Tridimensional Newton-Euler equations click [here](Tridimensional%20rigid%20body%20Kinetics.ipynb)):
\begin{align}
\overrightarrow{F_A} + \overrightarrow{GRF} + m_F\overrightarrow{g} &= m_F\overrightarrow{a_{cm_F}}\\
\overrightarrow{M_A} + \overrightarrow{M_{GRF}}+ \overrightarrow{M_{FA}}&=I_F\overrightarrow{\dot{\omega_F}} + \overrightarrow{\omega_F} \times (I_F\overrightarrow{\omega_F})\\
\overrightarrow{F_K} -\overrightarrow{F_A} + m_S\overrightarrow{g} &= m_S\overrightarrow{a_{cm_S}}\\
\overrightarrow{M_K} - \overrightarrow{M_A} + \overrightarrow{M_{FA}} + \overrightarrow{M_{FK}} &= I_S\overrightarrow{\dot{\omega_S}} + \overrightarrow{\omega_S} \times (I_S\overrightarrow{\omega_S})
\end{align}
where
- $\overrightarrow{g} = -9.81\hat{j}$;
- $m_F$ and $m_S$ are the masses of the foot and the shank, respectively;
- $\overrightarrow{GRF}$ is the ground reaction force being applied to the foot;
- $\overrightarrow{a_{cm_F}}$ and $\overrightarrow{a_{cm_S}}$ are the accelerations of the center of mass of the foot and the shank, respectively;
- $\overrightarrow{\omega_F}$ and $\overrightarrow{\omega_S}$ are the angular accelerations of the foot and shank, respectively, described at a basis attached to the segment, and $\overrightarrow{\dot{\omega_F}}$ and $\overrightarrow{\dot{\omega_S}}$ are their time-derivatives;
- $I_S$ and $I_F$ are the matrices of inertia of the shank and the foot, respectively;
- $\overrightarrow{F_K}$, $\overrightarrow{F_A}$, $\overrightarrow{M_A}$ and $\overrightarrow{M_A}$ are the forces and moments at the ankle and knee joints, respectively
Note that each of these equations have components at each of the three directions. Additionally, note that the equations of the forces are described in the global basis, and the equations of the moments must be described in the basis attached to the segment relative to that equation. So, it is a good idea to make this clear with a more precise notation. We will denote as a superscript in the vectors the segment where the basis that we are describing the vector is fixed. So for example, $\overrightarrow{M_A^F}$ is the vector of the moment due to the muscle forces of the ankle, described in the basis fixed at the foot. So, the equations can be rewritten as:
\begin{align}
\overrightarrow{F_A^G} + \overrightarrow{GRF^G} + m_F\overrightarrow{g^G} &= m_F\overrightarrow{a_{cm_F}^G}\\
\overrightarrow{M_A^F} + \overrightarrow{M_{GRF}^F}+ \overrightarrow{M_{FA}^F}&=I_F\overrightarrow{\dot{\omega_F^F}} + \overrightarrow{\omega_F^F} \times (I_F\overrightarrow{\omega_F^F})\\
\overrightarrow{F_K^G} -\overrightarrow{F_A^G} + m_S\overrightarrow{g^G} &= m_S\overrightarrow{a_{cm_S}^G}\\
\overrightarrow{M_K^S} - \overrightarrow{M_A^S} + \overrightarrow{M_{FA}^S} + \overrightarrow{M_{FK}^S} &= I_S\overrightarrow{\dot{\omega_S^S}} + \overrightarrow{\omega_S^S} \times (I_S\overrightarrow{\omega_S^S})
\end{align}
where the superscript $G$ denotes the global frame of reference, the superscript $S$ denotes the frame of reference in the shank and the superscript $F$ denotes the frame of reference at the foot.
The moments due to the ground reaction force, the force at the ankle and the force at the knee are computed by cross-multiplying them by their moment-arms. As the forces and the moment-arms are described in the global basis, we must multiply them by the rotation matrix of the basis corresponding to the segment. So, the equations can be rewritten as:
\begin{align}
\overrightarrow{F_A^G} + \overrightarrow{GRF^G} + m_F\overrightarrow{g^G} &= m_F\overrightarrow{a_{cm_F}^G}\\
\overrightarrow{M_A^F} + R_F(\overrightarrow{r_{cop/cm_F}^G}\times \overrightarrow{GRF^G})+ R_F(\overrightarrow{r_{A/cm_F}^G}\times \overrightarrow{F_A}^G)&=I_F\overrightarrow{\dot{\omega_F^F}} + \overrightarrow{\omega_F^F} \times (I_F\overrightarrow{\omega_F^F})\\
\overrightarrow{F_K^G} -\overrightarrow{F_A^G} + m_S\overrightarrow{g^G} &= m_S\overrightarrow{a_{cm_S}^G}\\
\overrightarrow{M_K^S} - \overrightarrow{M_A^S} - R_S(\overrightarrow{r_{A/cm_S}^G}\times \overrightarrow{F_A^G}) + R_S(\overrightarrow{r_{K/cm_S}^G}\times \overrightarrow{F_K^G}) &= I_S\overrightarrow{\dot{\omega_S^S}} + \overrightarrow{\omega_S^S} \times (I_S\overrightarrow{\omega_S^S})
\end{align}
where $R_S$ is the rotation matrix of the basis attached to the shank and $R_F$ is the rotation matrix of the basis attached to the foot.
Now, we can note that the vectors $\overrightarrow{M_K^S}$ and $\overrightarrow{M_K^F}$ are the same vectors described in different basis. So we could use only one of the descriptions and use rotation matrices to convert from one to another. To pass the vector from the foot coordinates to the shank coordinate, we must first multiply it by the inverted rotation matrix of the foot and then multiply it by the rotation matrix of the shank. So, $\overrightarrow{M_A^S} = R_SR_F^{-1}\overrightarrow{M_A^F}$ and the equations above can be rewritten as:
\begin{align}
\overrightarrow{F_A^G} + \overrightarrow{GRF^G} + m_F\overrightarrow{g^G} &= m_F\overrightarrow{a_{cm_F}^G}\\
\overrightarrow{M_A^F} + R_F(\overrightarrow{r_{cop/cm_F}^G}\times \overrightarrow{GRF^G})+ R_F(\overrightarrow{r_{A/cm_F}^G}\times \overrightarrow{F_A}^G)&=I_F\overrightarrow{\dot{\omega_F^F}} + \overrightarrow{\omega_F^F} \times (I_F\overrightarrow{\omega_F^F})\\
\overrightarrow{F_K^G} -\overrightarrow{F_A^G} + m_S\overrightarrow{g^G} &= m_S\overrightarrow{a_{cm_S}^G}\\
\overrightarrow{M_K^S} - R_SR_F^{-1}\overrightarrow{M_A^F} - R_S(\overrightarrow{r_{A/cm_S}^G}\times \overrightarrow{F_A^G}) + R_S(\overrightarrow{r_{K/cm_S}^G}\times \overrightarrow{F_K^G}) &= I_S\overrightarrow{\dot{\omega_S^S}} + \overrightarrow{\omega_S^S} \times (I_S\overrightarrow{\omega_S^S})
\end{align}
Now, we divide the equations above in the matrices defined previously:
\begin{equation}
\underbrace{\left[\begin{array}{cccc} m_FI_3& [0]& [0]& [0]\\ [0]& I_F & [0] & [0] \\ [0] &[0] & m_SI_3& [0] \\ [0] & [0] & [0] & I_S\end{array}\right]}_{M}\cdot\left[\begin{array}{c}\overrightarrow{a_{cm_F}^G}\\\overrightarrow{\dot{\omega_F^F}}\\\overrightarrow{a_{cm_S}^G}\\\overrightarrow{\dot{\omega_S^S}} \\ \end{array}\right] = \underbrace{\left[\begin{array}{c}[0]\\ - \overrightarrow{\omega_F^F} \times (I_F\overrightarrow{\omega_F^F}) \\ [0] \\ - \overrightarrow{\omega_S^S} \times (I_S\overrightarrow{\omega_S^S}) \end{array}\right]}_{C} + \underbrace{\left[\begin{array}{c} m_F\overrightarrow{g^G}\\ [0]\\ m_S\overrightarrow{g^G} \\ [0] \end{array}\right]}_{G} + \underbrace{\left[\begin{array}{c} \overrightarrow{F_A^G}\\ \overrightarrow{M_A^F}+R_F(\overrightarrow{r_{A/cm_F}^G}\times \overrightarrow{F_A}^G)\\ \overrightarrow{F_K^G} - \overrightarrow{F_A^G} \\ \overrightarrow{M_K^S} - R_SR_F^{-1}\overrightarrow{M_A^F} - R_S(\overrightarrow{r_{A/cm_S}^G}\times \overrightarrow{F_A^G}) + R_S(\overrightarrow{r_{K/cm_S}^G}\times \overrightarrow{F_K^G}) \end{array}\right]}_{Q} + \underbrace{\left[\begin{array}{c} \overrightarrow{GRF^G}\\ R_F(\overrightarrow{r_{cop/cm_F}^G}\times \overrightarrow{GRF^G})\\ [0] \\ [0] \end{array}\right]}_{E}
\end{equation}
where $I_3$ is the identity matrix 3x3.
To perform the inverse dynamics, we still cannot isolate the vector of forces and moments. As the vector $F$ has cross-products we must define the a new operator that performs the cross-product through a matrix multiplication.
We can note that the cross-product between the vectors $\vec{v}$ and $\vec{w}$ has the following result:
\begin{equation}
\vec{v} \times \vec{w} = \left[\begin{array}{c}v_x\\v_y\\v_z \end{array}\right] \times \left[\begin{array}{c}w_x\\w_y\\w_z \end{array}\right] = \left[\begin{array}{c}v_yw_z - v_zw_y\\v_zw_x - v_xw_z\\v_xw_y - v_yw_x \end{array}\right] = \left[\begin{array}{ccc}0&-v_z&v_y\\v_z&0&-v_x\\-v_y&v_x&0 \end{array}\right]\cdot\left[\begin{array}{c}w_x\\w_y\\w_z \end{array}\right]
\end{equation}
So we can define a new operator known as skew-symmetric matrix:
\begin{equation}
S(\vec{v}) \triangleq \left[\begin{array}{ccc}0&-v_z&v_y\\v_z&0&-v_x\\-v_y&v_x&0 \end{array}\right]
\end{equation}
Therefore:
\begin{equation}
\vec{v} \times \vec{w} = S(\vec{v})\cdot\vec{w}
\end{equation}
Now, we will use this operator in the equation we found previously:
\begin{equation}
\left[\begin{array}{cccc} m_FI_3& [0]& [0]& [0]\\ [0]& I_F & [0] & [0] \\ [0] &[0] & m_SI_3& [0] \\ [0] & [0] & [0] & I_S\end{array}\right]\cdot\left[\begin{array}{c}\overrightarrow{a_{cm_F}^G}\\\overrightarrow{\dot{\omega_F^F}}\\\overrightarrow{a_{cm_S}^G}\\\overrightarrow{\dot{\omega_S^S}} \\ \end{array}\right] = \left[\begin{array}{c}[0]\\ - \overrightarrow{\omega_F^F} \times (I_F\overrightarrow{\omega_F^F}) \\ [0] \\ - \overrightarrow{\omega_S^S} \times (I_S\overrightarrow{\omega_S^S}) \end{array}\right] + \left[\begin{array}{c} m_F\overrightarrow{g^G}\\ [0]\\ m_S\overrightarrow{g^G} \\ [0] \end{array}\right] + \left[\begin{array}{c} \overrightarrow{F_A^G}\\ \overrightarrow{M_A^F}+R_F(S(\overrightarrow{r_{A/cm_F}^G})\cdot\overrightarrow{F_A}^G)\\ \overrightarrow{F_K^G} - \overrightarrow{F_A^G} \\ \overrightarrow{M_K^S} - R_SR_F^{-1}\overrightarrow{M_A^F} - R_S(S(\overrightarrow{r_{A/cm_S}^G})\cdot\overrightarrow{F_A^G}) + R_S(S(\overrightarrow{r_{K/cm_S}^G})\cdot\overrightarrow{F_K^G}) \end{array}\right] + \left[\begin{array}{c} \overrightarrow{GRF^G}\\ R_F(\overrightarrow{r_{cop/cm_F}^G}\times \overrightarrow{GRF^G})\\ [0] \\ [0] \end{array}\right]
\end{equation}
Now it is possible to write the vector $F$ as multiplication of a matrix by a vector:
\begin{equation}
\left[\begin{array}{cccc} m_FI_3& [0]& [0]& [0]\\ [0]& I_F & [0] & [0] \\ [0] &[0] & m_SI_3& [0] \\ [0] & [0] & [0] & I_S\end{array}\right]\cdot\left[\begin{array}{c}\overrightarrow{a_{cm_F}^G}\\\overrightarrow{\dot{\omega_F^F}}\\\overrightarrow{a_{cm_S}^G}\\\overrightarrow{\dot{\omega_S^S}} \\ \end{array}\right] = \left[\begin{array}{c}[0]\\ - \overrightarrow{\omega_F^F} \times (I_F\overrightarrow{\omega_F^F}) \\ [0] \\ - \overrightarrow{\omega_S^S} \times (I_S\overrightarrow{\omega_S^S}) \end{array}\right] + \left[\begin{array}{c} m_F\overrightarrow{g^G}\\ [0]\\ m_S\overrightarrow{g^G} \\ [0] \end{array}\right] + \left[\begin{array}{ccc} I_3& [0]& [0]& [0]\\ R_FS\left(\overrightarrow{r_{A/cm_F}^G}\right)&I_3& [0]& [0]\\ -I_3& [0]& I_3 & [0] \\ -R_SS\left(\overrightarrow{r_{A/cm_S}^G}\right)& - R_SR_F^{-1} & R_SS\left(\overrightarrow{r_{K/cm_S}^G}\right) & I_3 \end{array}\right]\cdot\left[\begin{array}{c} \overrightarrow{F_A^G}\\ \overrightarrow{M_A^F}\\ \overrightarrow{F_K^G}\\ \overrightarrow{M_K^S}\end{array}\right] + \left[\begin{array}{c} \overrightarrow{GRF^G}\\ R_F(\overrightarrow{r_{cop/cm_F}^G}\times \overrightarrow{GRF^G})\\ [0] \\ [0] \end{array}\right]
\end{equation}
So, the final equation to compute the forces and torques is obtained by multiplying everything by the inverse of the matrix multipliying the vector of forces:
\begin{equation}
\left[\begin{array}{c} \overrightarrow{F_A^G}\\ \overrightarrow{M_A^F}\\ \overrightarrow{F_K^G}\\ \overrightarrow{M_K^S}\end{array}\right] = \left[\begin{array}{ccc} I_3& [0]& [0]& [0]\\ R_FS\left(\overrightarrow{r_{A/cm_F}^G}\right)&I_3& [0]& [0]\\ -I_3& [0]& I_3 & [0] \\ -R_SS\left(\overrightarrow{r_{A/cm_S}^G}\right)& - R_SR_F^{-1} & R_SS\left(\overrightarrow{r_{K/cm_S}^G}\right) & I_3 \end{array}\right]^{-1}\cdot\left(\left[\begin{array}{cccc} m_FI_3& [0]& [0]& [0]\\ [0]& I_F & [0] & [0] \\ [0] &[0] & m_SI_3& [0] \\ [0] & [0] & [0] & I_S\end{array}\right]\cdot\left[\begin{array}{c}\overrightarrow{a_{cm_F}^G}\\\overrightarrow{\dot{\omega_F^F}}\\\overrightarrow{a_{cm_S}^G}\\\overrightarrow{\dot{\omega_S^S}} \\ \end{array}\right] - \left[\begin{array}{c}[0]\\ - \overrightarrow{\omega_F^F} \times (I_F\overrightarrow{\omega_F^F}) \\ [0] \\ - \overrightarrow{\omega_S^S} \times (I_S\overrightarrow{\omega_S^S}) \end{array}\right] -\left[\begin{array}{c} \overrightarrow{GRF^G}\\ R_F(\overrightarrow{r_{cop/cm_F}^G}\times \overrightarrow{GRF^G})\\ [0] \\ [0] \end{array}\right] - \left[\begin{array}{c} m_F\overrightarrow{g^G}\\ [0]\\ m_S\overrightarrow{g^G} \\ [0] \end{array}\right]\right)
\end{equation}
With the last equation, we can obtain all the forces and moments using only one line of code. Computationally, it is less prone to errors and more efficient.
So, generically, the steps to perform the analysis of inverse dynamics is:
- write the equations of Newton-Euler for each segment. Write explicitly the basis at which each vector is described.
- use the rotation matrices of the basis to pass the description of a vector to another basis. Use it in a way that the same vector is described at just a single frame of reference.
- write the cross-products as a product between the skew-symmetric matrix $S$ of the first vector and the second vector.
- write the equations in the matrix format, repeated here:
\begin{equation}
M(q)\ddot{q} = C(q,\dot{q}) + G(q) + Q + E
\end{equation}
- write explicitly the vector containing the unknown forces and moments $Q$, as a multiplication of a matrix and vector containing only the unknown forces.
- isolate the vector containing only the unknown forces by multiplying the whole equation by the inverse of the matrix multiplying the vector with the forces.
## Problems
1) Solve the problems 18.3.20 and 18.3.24 of the Ruina and Rudra's book by using the Lagrangian formalism (it is much easier than use the Newton-Euler formalism) and then use the matrix formalism to obtain the expressions of the angular accelerations.
2) Write the matrices to find the forces and torques in a tridimensional double pendulum, consisted of two cylindrical bars. Consider that you know all the masses, moments of inertia, rotation matrices, accelerations, angular velocities and angular accelerations necessary to solve the problem.
## References
- YAMAGUCHI, G. T. Dynamic modeling of musculoskeletal motion: a vectorized approach for biomechanical analysis in three dimensions., 2001
- CRAIG, J. Introduction to robotics. , 1989
- JAIN, A. Robot and multibody dynamics. , 2011
- SPONG, M. W.; HUTCHINSON, S.; VIDYASAGAR, M. Robot modeling and control., 2006
- ERDEMIR, A. et al. Model-based estimation of muscle forces exerted during movements. Clinical Biomechanics, v. 22, n. 2, p. 131–154, 2007.
- STANEV, D.; MOUSTAKAS, K. Simulation of constrained musculoskeletal systems in task space. IEEE Transactions on Biomedical Engineering, v. 65, n. 2, p. 307–318, 2018.
- ZAJAC FE, GORDON ME , [Determining muscle's force and action in multi-articular movement](https://drive.google.com/open?id=0BxbW72zV7WmUcC1zSGpEOUxhWXM&authuser=0). Exercise and Sport Sciences Reviews, 17, 187-230. , 1989
- RUINA A, RUDRA P. [Introduction to Statics and Dynamics](http://ruina.tam.cornell.edu/Book/index.html). Oxford University Press. , 2015
| github_jupyter |
## Treat BERT as a contextualized embedding generator
This notebook will get the necessary folders ready and generate the necessary text files for bert
```
%load_ext autoreload
%autoreload
import os
from sys import path
path.append('../..')
from relation_extraction.data import utils
# from bert import extract_features
# from relation_extraction.data import utils
# import h5py
# import numpy as np
# dataset = 'semeval2010' #TODO: update this
preprocessing_type = 'entity_blinding' #TODO: update this; mostly we are only going to be using original
# cuda_device=3 #cuda device for the elmo embeddings generator
dataset = 'ddi'
data_path = '/data/scratch-oc40/geeticka/data/relation_extraction/' + dataset + '/'
def res(path): return os.path.join(data_path, path)
out_path = 'pre-processed/'+ preprocessing_type +'/bert-CLS/'
# if not os.path.exists(res(out_path)):
# os.makedirs(res(out_path))
# if not os.path.exists(res(out_path + 'input-sentences')):
# os.makedirs(res(out_path + 'input-sentences'))
border_sizes = [-1] # for i2b2
def get_sentences(filename, border_size):
data = open(res('pre-processed/'+ preprocessing_type+'/' + filename))
data = utils.split_data_cut_sentence(data, border_size)
sentences = data[0]
return sentences
def write_sentences_to_txt(sentences, filename):
with open(res(out_path + 'input-sentences/'+filename + '.txt'), 'w') as f:
for sentence in sentences:
sentence_string = " ".join(sentence)
f.write(sentence_string + "\n")
# for border_size in border_sizes:
# train_sentences = get_sentences('train_'+ preprocessing_type +'.txt', border_size)
# test_sentences = get_sentences('test_'+ preprocessing_type +'.txt', border_size)
# write_sentences_to_txt(train_sentences, 'train_'+ preprocessing_type +'_border_' + str(border_size))
# write_sentences_to_txt(test_sentences, 'test_'+ preprocessing_type +'_border_' + str(border_size))
```
## Now, proceed to generating the BERT embeddings using the scripts provided in pytorch inside the bert folder where these sentences are located
### Now, we would like to read these BERT embeddings generated in the BERT folder and generate the json file without the word piece embeddings
```
in_path = 'pre-processed/'+ preprocessing_type +'/bert-CLS/'
out_path = 'pre-processed/' + preprocessing_type + '/bert-tokens/'
if not os.path.exists(res(out_path)):
os.makedirs(res(out_path))
border_size = -1
train_filename = 'train_' + preprocessing_type + '_border_' + str(border_size) + '.json'
test_filename = 'test_' + preprocessing_type + '_border_' + str(border_size) + '.json'
utils.write_bert_tokens_without_word_pieces(res(in_path + train_filename), res(out_path + train_filename))
utils.write_bert_tokens_without_word_pieces(res(in_path + test_filename), res(out_path + test_filename))
import json
with open(res(out_path + train_filename), 'r', encoding='utf-8') as file:
length = 0
for line in file.readlines():
data = json.loads(line)
length += 1
length
with open(res(out_path + test_filename), 'r', encoding='utf-8') as file:
length = 0
for line in file.readlines():
data = json.loads(line)
length += 1
length
```
| github_jupyter |
# Practice with `numpy`!
**Remember:**
* Numpy provides a bunch of useful tools for performing calculations
* You access numpy functions by calling `np.function_name`
First, __import numpy__. Remember to use the nickname!
```
# load numpy
import numpy as np
```
Use numpy to __create a list of numbers from 0 through 99.__ We'll use a function called __`arange`.__
```
# Use np.arrange to generate an array of numbers and assign it to a variable called numbers
numbers = np.arange(100)
# print the array to see what it looks like
print(numbers)
```
Now use numpy's function called __`zeros`__ to __create another empty array__ of the same size:
```
# Create an empty array with np.zeros and assign it to a variable
zeros = np.zeros(100)
# print it
print(zeros)
```
Now try manipulating the arrays using basic math:
```
# Add the numbers array to itself
numbers + numbers
# Multiply the numbers array to itself
numbers * numbers
```
Note that when you perform a math operation on an array, __it will often perform that operation on each item in that array.__ It's convenient that __we don't have to loop through all the values__ to apply the math operation to every item in the array.
__You can find information about the size of an array by using `.shape`.__ Note that `.shape` is an *attribute* of array -- a special variable that belongs to every *array object*. Try it out:
*HINT*: Because `.shape` is not a function you don't need to use parentheses.
```
# Use shape to view the size of your array
numbers.shape
```
`.shape` gave us just one number, because our array has only 1 dimension. Later we'll see what it looks like for arrays with more than 1 dimension.
Numpy also allows you to create 2D arrays, like with lists. We can __use the `method` called `reshape` to change an 1-dimensional `array` into a 2-dimensional `array`.__ `reshape` takes two arguments: the number of rows and the number of columns. Try turning one of your arrays into a 2D array using `reshape`.
```
# Reshape one of your arrays into a 2D array
numbers_2d = numbers.reshape( 10, 10 )
numbers_2d
```
Now the `.shape` of your array should be changed, __try printing it out below:__
```
# Print the shape of your new array
numbers_2d.shape
```
Now we will try a couple of numpy's math functions!
```
# try using np.sum to add the items in a list together
print(np.sum(numbers))
# try squaring the value of each item in your array
print(numbers**2)
```
__Try converting the `numbers` array to an array of floats__ using the method called `astype`:
```
# Convert the array into an array of floats
print(numbers.astype(float))
```
Nice job! You just practiced:
* Using `numpy` to perform `array` operations.
* Performing math with `numpy`.
| github_jupyter |
# TD 3
```
sigmoid <- function(x) {
return (exp(x)/(1+exp(x)))
}
df <- read.table("titanic.csv", sep=";", header=TRUE, dec=",", na.strings = "")
df$pclass <- factor(df$pclass)
df$survived <- factor(df$survived)
df <- df[complete.cases(df),]
sigmoid()
```
### Partie 3
##### Q1
(a) L'odd est défini par $\text{odd}(x)=\frac{\mathbb{P}(Y=1|x)}{\mathbb{P}(Y=0|x)}=\frac{\pi(x)}{1-\pi(x)}$
```
df$survived <- relevel(df$survived, ref="0")
surv.null <- glm(survived ~ 1, family = binomial, data = df)
surv.null
```
Dans le cas d'une régression logistique avec un seul coefficient on a $\pi(x)=\sigma(\beta_{0})$
```
sigmoid(-0.376)/(1-sigmoid(-0.376))
exp(coef(surv.null))
```
On a 0.68 fois plus de chance de survivre que de mourir au naufrage du titanic.
##### Q2
```
df$survived <- relevel(df$survived, ref="0")
df$sex<- relevel(df$sex, ref='male')
surv.sex <- glm(survived ~ sex, family = binomial, data = df)
summary(surv.sex)
sigmoid(1.1055)
sigmoid(-1.35431)
a=sigmoid(1.1055)/(1-sigmoid(1.1055))
b=sigmoid(-1.35431)/(1-sigmoid(1.35431))
exp(2.45984)
(0.75/0.25)/(0.20/0.80)
```
(a)
```
1-pchisq(surv.sex$null.deviance-surv.sex$deviance,surv.sex$df.null-surv.sex$df.residual)
```
On rejette l'hypothèse nulle : le modèle est statistiquement valable
Pour une variable binaire $OR=\frac{\frac{\pi(1)}{1-\pi(1)}}{\frac{\pi(0)}{1-\pi(0)}}$
$log(OR)=log(\frac{\pi(1)}{1-\pi(1)})-log(\frac{\pi(0)}{1-\pi(0)})=logit(\pi(1))-logit(\pi(0))$
Ici notre modèle s'écrit $\pi(x)=\sigma(\beta_{0}+\beta_{1}\mathbb{I}_{\text{sex=homme}}(x))$
Alors $\pi(1)=\sigma(\beta_{0}+\beta_{1})$ et $\pi(0)=\sigma(\beta_{0})$
Alors $log(OR)=logit(\pi(1))-logit(\pi(0))=\beta_{1}$
D'où $OR=exp(\beta_{1})$
(a)
```
a=sigmoid(-1.35431)/(1-sigmoid(-1.35431))
b=sigmoid(-1.35431+2.45984)/(1-sigmoid(-1.35431+2.45984))
b/a
exp(2.45984)
```
(b)
on a 11,7 fois plus de chances de survivre au naufrage du titanic lorsqu'on est une femme que lorsqu'on n'en est pas une.
#### Q3
(a) $\pi(x)=\sigma(\beta_{0}+\beta_{1}\mathbb{I}_{\text{pclass=2}}(x)+\beta_{2}\mathbb{I}_{\text{pclass=3}}(x))$
```
df$pclass<- relevel(df$pclass, ref='3')
surv.pclass <- glm(survived ~ pclass, family = binomial, data = df)
summary(surv.pclass)
exp(1.5910)
exp(0.7996)
1-pchisq(surv.pclass$null.deviance-surv.pclass$deviance,surv.pclass$df.null-surv.pclass$df.residual)
1/0.91
```
$OR(class2)=\frac{\frac{\pi(class2)}{1-\pi(class2)}}{\frac{\pi(class1)}{1-\pi(class1)}}$
$OR(class3)=\frac{\frac{\pi(class3)}{1-\pi(class3)}}{\frac{\pi(class1)}{1-\pi(class1)}}$
```
a=sigmoid(0.5527 -0.7913)/(1-sigmoid(0.5527 -0.7913))
b=sigmoid(0.5527 -1.5910)/(1-sigmoid(0.5527 -1.5910))
c=sigmoid(0.5527)/(1-sigmoid(0.5527))
a/c
exp(-0.7913)
b/c
exp(-1.5910)
1/0.20
```
(b) on a 0.45 fois plus de chance de survivre au naufrage dans une cabine de classe 2 que dans une cabine de classe 1 -> on a 2.22 fois plus de chance de mourir pendant le naufrage dans une cabine de classe 2 que dans une cabine de classe 1.
on a 0.20 fois plus de chance de survivre au naufrage dans une cabine de classe 3 que dans une cabine de classe 1 -> on a 5 fois plus de chance de mourir pendant le naufrage dans une cabine de classe 3 que dans une cabine de classe 1.
##### Q4
(a) Le modèle s'écrit $\pi(\text{age})=\sigma(\beta_{0}+\beta_{1}\text{age})=\frac{e^{\beta_{0}+\beta_{1}\text{age}}}{1+e^{\beta_{0}+\beta_{1}\text{age}}}$ où $\text{age}$ est l'âge
```
df$survived <- relevel(df$survived, ref="0")
surv.age <- glm(survived ~ age, family = binomial, data = df)
summary(surv.age)
exp(-0.008413)
1/exp(10*-0.008413)
```
on a 0.99 fois plus de chance de survivre avec 1 an de plus
(b) $OR = exp(10*b_{1})$.
En effet, on a :
pour x = a : $logit(\pi(a)) = \beta_{0}+\beta_{1}*a$
pour x = a+10 : $logit(\pi(a+10)) = \beta_{0}+\beta_{1}*a + \beta_{1}*10$
$OR = (\pi(a+10)/(1-\pi(a+10))) / (\pi(a)/(1-\pi(a))) = exp(\beta_{0})*exp(\beta_{1}*a)*exp(\beta_{1}*10) / exp(\beta_{0})*exp(\beta_{1}*a) = exp(\beta_{1}*10)$
(c)
```
1-pchisq(surv.age$null.deviance-surv.age$deviance,surv.age$df.null-surv.age$df.residual)
```
(e)
```
confint(surv.age)
exp(confint(surv.age))
```
## Partie 4
```
bornes <- c(quantile(df$age, probs = seq(0, 1, by = 0.2)))
bornes
df$age_discret <- cut(df$age, breaks=bornes)
df$age_discret
pix = table(df$age_discret, df$survived)/rowSums(table(df$age_discret, df$survived))
pix
logit=log((pix[,2])/(1-pix[,2]))
bornes <- c(quantile(df$age, probs = seq(0, 1, by = 0.2)))
df$age_discret <- cut(df$age, breaks=bornes)
pix = table(df$age_discret, df$survived)/rowSums(table(df$age_discret, df$survived))
logit=log((pix[,2])/(1-pix[,2]))
x = (c(0,19,25,31,42)+c(19,25,31,42,80))/2
plot(x, logit)
lines(x, lm(logit~x)$fit)
surv.age.discret <- glm(survived ~ age_discret, family = binomial, data = df)
summary(surv.age.discret)
1-pchisq(surv.age.discret$null.deviance-surv.age.discret$deviance,surv.age.discret$df.null-surv.age.discret$df.residual)
```
| github_jupyter |
# Training a ConvNet PyTorch
In this notebook, you'll learn how to use the powerful PyTorch framework to specify a conv net architecture and train it on the CIFAR-10 dataset.
```
import torch
import torch.nn as nn
import torch.optim as optim
from torch.autograd import Variable
from torch.utils.data import DataLoader
from torch.utils.data import sampler
import torchvision.datasets as dset
import torchvision.transforms as T
import numpy as np
import timeit
import os
os.chdir(os.getcwd() + '/..')
```
## What's this PyTorch business?
You've written a lot of code in this assignment to provide a whole host of neural network functionality. Dropout, Batch Norm, and 2D convolutions are some of the workhorses of deep learning in computer vision. You've also worked hard to make your code efficient and vectorized.
For the last part of this assignment, though, we're going to leave behind your beautiful codebase and instead migrate to one of two popular deep learning frameworks: in this instance, PyTorch (or TensorFlow, if you switch over to that notebook).
Why?
* Our code will now run on GPUs! Much faster training. When using a framework like PyTorch or TensorFlow you can harness the power of the GPU for your own custom neural network architectures without having to write CUDA code directly (which is beyond the scope of this class).
* We want you to be ready to use one of these frameworks for your project so you can experiment more efficiently than if you were writing every feature you want to use by hand.
* We want you to stand on the shoulders of giants! TensorFlow and PyTorch are both excellent frameworks that will make your lives a lot easier, and now that you understand their guts, you are free to use them :)
* We want you to be exposed to the sort of deep learning code you might run into in academia or industry.
## How will I learn PyTorch?
If you've used Torch before, but are new to PyTorch, this tutorial might be of use: http://pytorch.org/tutorials/beginner/former_torchies_tutorial.html
Otherwise, this notebook will walk you through much of what you need to do to train models in Torch. See the end of the notebook for some links to helpful tutorials if you want to learn more or need further clarification on topics that aren't fully explained here.
## Load Datasets
We load the CIFAR-10 dataset. This might take a couple minutes the first time you do it, but the files should stay cached after that.
```
class ChunkSampler(sampler.Sampler):
"""Samples elements sequentially from some offset.
Arguments:
num_samples: # of desired datapoints
start: offset where we should start selecting from
"""
def __init__(self, num_samples, start = 0):
self.num_samples = num_samples
self.start = start
def __iter__(self):
return iter(range(self.start, self.start + self.num_samples))
def __len__(self):
return self.num_samples
NUM_TRAIN = 49000
NUM_VAL = 1000
cifar10_train = dset.CIFAR10('datasets', train=True, download=True,
transform=T.ToTensor())
loader_train = DataLoader(cifar10_train, batch_size=64, sampler=ChunkSampler(NUM_TRAIN, 0))
cifar10_val = dset.CIFAR10('datasets', train=True, download=True,
transform=T.ToTensor())
loader_val = DataLoader(cifar10_val, batch_size=64, sampler=ChunkSampler(NUM_VAL, NUM_TRAIN))
cifar10_test = dset.CIFAR10('datasets', train=False, download=True,
transform=T.ToTensor())
loader_test = DataLoader(cifar10_test, batch_size=64)
```
For now, we're going to use a CPU-friendly datatype. Later, we'll switch to a datatype that will move all our computations to the GPU and measure the speedup.
```
dtype = torch.FloatTensor # the CPU datatype
# Constant to control how frequently we print train loss
print_every = 100
# This is a little utility that we'll use to reset the model
# if we want to re-initialize all our parameters
def reset(m):
if hasattr(m, 'reset_parameters'):
m.reset_parameters()
```
## Example Model
### Some assorted tidbits
Let's start by looking at a simple model. First, note that PyTorch operates on Tensors, which are n-dimensional arrays functionally analogous to numpy's ndarrays, with the additional feature that they can be used for computations on GPUs.
We'll provide you with a Flatten function, which we explain here. Remember that our image data (and more relevantly, our intermediate feature maps) are initially N x C x H x W, where:
* N is the number of datapoints
* C is the number of channels
* H is the height of the intermediate feature map in pixels
* W is the height of the intermediate feature map in pixels
This is the right way to represent the data when we are doing something like a 2D convolution, that needs spatial understanding of where the intermediate features are relative to each other. When we input data into fully connected affine layers, however, we want each datapoint to be represented by a single vector -- it's no longer useful to segregate the different channels, rows, and columns of the data. So, we use a "Flatten" operation to collapse the C x H x W values per representation into a single long vector. The Flatten function below first reads in the N, C, H, and W values from a given batch of data, and then returns a "view" of that data. "View" is analogous to numpy's "reshape" method: it reshapes x's dimensions to be N x ??, where ?? is allowed to be anything (in this case, it will be C x H x W, but we don't need to specify that explicitly).
```
class Flatten(nn.Module):
def forward(self, x):
N, C, H, W = x.size() # read in N, C, H, W
return x.view(N, -1) # "flatten" the C * H * W values into a single vector per image
```
### The example model itself
The first step to training your own model is defining its architecture.
Here's an example of a convolutional neural network defined in PyTorch -- try to understand what each line is doing, remembering that each layer is composed upon the previous layer. We haven't trained anything yet - that'll come next - for now, we want you to understand how everything gets set up. nn.Sequential is a container which applies each layer
one after the other.
In that example, you see 2D convolutional layers (Conv2d), ReLU activations, and fully-connected layers (Linear). You also see the Cross-Entropy loss function, and the Adam optimizer being used.
Make sure you understand why the parameters of the Linear layer are 5408 and 10.
```
# Here's where we define the architecture of the model...
simple_model = nn.Sequential(
nn.Conv2d(3, 32, kernel_size=7, stride=2),
nn.ReLU(inplace=True),
Flatten(), # see above for explanation
nn.Linear(5408, 10), # affine layer
)
# Set the type of all data in this model to be FloatTensor
simple_model.type(dtype)
loss_fn = nn.CrossEntropyLoss().type(dtype)
optimizer = optim.Adam(simple_model.parameters(), lr=1e-2) # lr sets the learning rate of the optimizer
```
PyTorch supports many other layer types, loss functions, and optimizers - you will experiment with these next. Here's the official API documentation for these (if any of the parameters used above were unclear, this resource will also be helpful). One note: what we call in the class "spatial batch norm" is called "BatchNorm2D" in PyTorch.
* Layers: http://pytorch.org/docs/nn.html
* Activations: http://pytorch.org/docs/nn.html#non-linear-activations
* Loss functions: http://pytorch.org/docs/nn.html#loss-functions
* Optimizers: http://pytorch.org/docs/optim.html#algorithms
## Training a specific model
In this section, we're going to specify a model for you to construct. The goal here isn't to get good performance (that'll be next), but instead to get comfortable with understanding the PyTorch documentation and configuring your own model.
Using the code provided above as guidance, and using the following PyTorch documentation, specify a model with the following architecture:
* 7x7 Convolutional Layer with 32 filters and stride of 1
* ReLU Activation Layer
* Spatial Batch Normalization Layer
* 2x2 Max Pooling layer with a stride of 2
* Affine layer with 1024 output units
* ReLU Activation Layer
* Affine layer from 1024 input units to 10 outputs
And finally, set up a **cross-entropy** loss function and the **RMSprop** learning rule.
```
fixed_model_base = nn.Sequential( # You fill this in!
nn.Conv2d(3, 32, kernel_size=7, stride=1),
nn.ReLU(inplace=True),
nn.BatchNorm2d(32),
nn.MaxPool2d(2, stride=2),
Flatten(),
nn.Linear(5408, 1024),
nn.ReLU(inplace=True),
nn.Linear(1024, 10),
)
fixed_model = fixed_model_base.type(dtype)
loss_fn = nn.CrossEntropyLoss().type(dtype)
optimizer = optim.RMSprop(fixed_model.parameters())
```
To make sure you're doing the right thing, use the following tool to check the dimensionality of your output (it should be 64 x 10, since our batches have size 64 and the output of the final affine layer should be 10, corresponding to our 10 classes):
```
## Now we're going to feed a random batch into the model you defined and make sure the output is the right size
x = torch.randn(64, 3, 32, 32).type(dtype)
x_var = Variable(x.type(dtype)) # Construct a PyTorch Variable out of your input data
ans = fixed_model(x_var) # Feed it through the model!
# Check to make sure what comes out of your model
# is the right dimensionality... this should be True
# if you've done everything correctly
np.array_equal(np.array(ans.size()), np.array([64, 10]))
```
### GPU!
Now, we're going to switch the dtype of the model and our data to the GPU-friendly tensors, and see what happens... everything is the same, except we are casting our model and input tensors as this new dtype instead of the old one.
If this returns false, or otherwise fails in a not-graceful way (i.e., with some error message), you may not have an NVIDIA GPU available on your machine. If you're running locally, we recommend you switch to Google Cloud and follow the instructions to set up a GPU there. If you're already on Google Cloud, something is wrong -- make sure you followed the instructions on how to request and use a GPU on your instance. If you did, post on Piazza or come to Office Hours so we can help you debug.
```
# Verify that CUDA is properly configured and you have a GPU available
torch.cuda.is_available()
import copy
gpu_dtype = torch.cuda.FloatTensor
fixed_model_gpu = copy.deepcopy(fixed_model_base).type(gpu_dtype)
x_gpu = torch.randn(64, 3, 32, 32).type(gpu_dtype)
x_var_gpu = Variable(x.type(gpu_dtype)) # Construct a PyTorch Variable out of your input data
ans = fixed_model_gpu(x_var_gpu) # Feed it through the model!
# Check to make sure what comes out of your model
# is the right dimensionality... this should be True
# if you've done everything correctly
np.array_equal(np.array(ans.size()), np.array([64, 10]))
```
Run the following cell to evaluate the performance of the forward pass running on the CPU:
```
%%timeit
ans = fixed_model(x_var)
```
... and now the GPU:
```
%%timeit
torch.cuda.synchronize() # Make sure there are no pending GPU computations
ans = fixed_model_gpu(x_var_gpu) # Feed it through the model!
torch.cuda.synchronize() # Make sure there are no pending GPU computations
```
You should observe that even a simple forward pass like this is significantly faster on the GPU. So for the rest of the assignment (and when you go train your models in assignment 3 and your project!), you should use the GPU datatype for your model and your tensors: as a reminder that is *torch.cuda.FloatTensor* (in our notebook here as *gpu_dtype*)
### Train the model.
Now that you've seen how to define a model and do a single forward pass of some data through it, let's walk through how you'd actually train one whole epoch over your training data (using the simple_model we provided above).
Make sure you understand how each PyTorch function used below corresponds to what you implemented in your custom neural network implementation.
Note that because we are not resetting the weights anywhere below, if you run the cell multiple times, you are effectively training multiple epochs (so your performance should improve).
First, set up an RMSprop optimizer (using a 1e-3 learning rate) and a cross-entropy loss function:
```
loss_fn = nn.CrossEntropyLoss().type(dtype)
optimizer = optim.RMSprop(simple_model.parameters(), lr=1e-3)
# This sets the model in "training" mode. This is relevant for some layers that may have different behavior
# in training mode vs testing mode, such as Dropout and BatchNorm.
fixed_model.train()
# Load one batch at a time.
for t, (x, y) in enumerate(loader_train):
x_var = Variable(x.type(dtype))
y_var = Variable(y.type(dtype).long())
# This is the forward pass: predict the scores for each class, for each x in the batch.
scores = fixed_model(x_var)
# Use the correct y values and the predicted y values to compute the loss.
loss = loss_fn(scores, y_var)
if (t + 1) % print_every == 0:
print('t = %d, loss = %.4f' % (t + 1, loss.data[0]))
# Zero out all of the gradients for the variables which the optimizer will update.
optimizer.zero_grad()
# This is the backwards pass: compute the gradient of the loss with respect to each
# parameter of the model.
loss.backward()
# Actually update the parameters of the model using the gradients computed by the backwards pass.
optimizer.step()
```
Now you've seen how the training process works in PyTorch. To save you writing boilerplate code, we're providing the following helper functions to help you train for multiple epochs and check the accuracy of your model:
```
def train(model, loss_fn, optimizer, num_epochs = 1):
for epoch in range(num_epochs):
print('Starting epoch %d / %d' % (epoch + 1, num_epochs))
model.train()
for t, (x, y) in enumerate(loader_train):
x_var = Variable(x.type(dtype))
y_var = Variable(y.type(dtype).long())
scores = model(x_var)
loss = loss_fn(scores, y_var)
if (t + 1) % print_every == 0:
print('t = %d, loss = %.4f' % (t + 1, loss.data[0]))
optimizer.zero_grad()
loss.backward()
optimizer.step()
def check_accuracy(model, loader):
if loader.dataset.train:
print('Checking accuracy on validation set')
else:
print('Checking accuracy on test set')
num_correct = 0
num_samples = 0
model.eval() # Put the model in test mode (the opposite of model.train(), essentially)
for x, y in loader:
x_var = Variable(x.type(dtype), volatile=True)
scores = model(x_var)
_, preds = scores.data.cpu().max(1)
num_correct += (preds == y).sum()
num_samples += preds.size(0)
acc = float(num_correct) / num_samples
print('Got %d / %d correct (%.2f)' % (num_correct, num_samples, 100 * acc))
```
### Check the accuracy of the model.
Let's see the train and check_accuracy code in action -- feel free to use these methods when evaluating the models you develop below.
You should get a training loss of around 1.2-1.4, and a validation accuracy of around 50-60%. As mentioned above, if you re-run the cells, you'll be training more epochs, so your performance will improve past these numbers.
But don't worry about getting these numbers better -- this was just practice before you tackle designing your own model.
```
torch.manual_seed(12345)
fixed_model.apply(reset)
train(fixed_model, loss_fn, optimizer, num_epochs=1)
check_accuracy(fixed_model, loader_val)
```
### Don't forget the validation set!
And note that you can use the check_accuracy function to evaluate on either the test set or the validation set, by passing either **loader_test** or **loader_val** as the second argument to check_accuracy. You should not touch the test set until you have finished your architecture and hyperparameter tuning, and only run the test set once at the end to report a final value.
## Train a _great_ model on CIFAR-10!
Now it's your job to experiment with architectures, hyperparameters, loss functions, and optimizers to train a model that achieves **>=70%** accuracy on the CIFAR-10 **validation** set. You can use the check_accuracy and train functions from above.
### Things you should try:
- **Filter size**: Above we used 7x7; this makes pretty pictures but smaller filters may be more efficient
- **Number of filters**: Above we used 32 filters. Do more or fewer do better?
- **Pooling vs Strided Convolution**: Do you use max pooling or just stride convolutions?
- **Batch normalization**: Try adding spatial batch normalization after convolution layers and vanilla batch normalization after affine layers. Do your networks train faster?
- **Network architecture**: The network above has two layers of trainable parameters. Can you do better with a deep network? Good architectures to try include:
- [conv-relu-pool]xN -> [affine]xM -> [softmax or SVM]
- [conv-relu-conv-relu-pool]xN -> [affine]xM -> [softmax or SVM]
- [batchnorm-relu-conv]xN -> [affine]xM -> [softmax or SVM]
- **Global Average Pooling**: Instead of flattening and then having multiple affine layers, perform convolutions until your image gets small (7x7 or so) and then perform an average pooling operation to get to a 1x1 image picture (1, 1 , Filter#), which is then reshaped into a (Filter#) vector. This is used in [Google's Inception Network](https://arxiv.org/abs/1512.00567) (See Table 1 for their architecture).
- **Regularization**: Add l2 weight regularization, or perhaps use Dropout.
### Tips for training
For each network architecture that you try, you should tune the learning rate and regularization strength. When doing this there are a couple important things to keep in mind:
- If the parameters are working well, you should see improvement within a few hundred iterations
- Remember the coarse-to-fine approach for hyperparameter tuning: start by testing a large range of hyperparameters for just a few training iterations to find the combinations of parameters that are working at all.
- Once you have found some sets of parameters that seem to work, search more finely around these parameters. You may need to train for more epochs.
- You should use the validation set for hyperparameter search, and save your test set for evaluating your architecture on the best parameters as selected by the validation set.
### Going above and beyond
If you are feeling adventurous there are many other features you can implement to try and improve your performance. You are **not required** to implement any of these; however they would be good things to try for extra credit.
- Alternative update steps: For the assignment we implemented SGD+momentum, RMSprop, and Adam; you could try alternatives like AdaGrad or AdaDelta.
- Alternative activation functions such as leaky ReLU, parametric ReLU, ELU, or MaxOut.
- Model ensembles
- Data augmentation
- New Architectures
- [ResNets](https://arxiv.org/abs/1512.03385) where the input from the previous layer is added to the output.
- [DenseNets](https://arxiv.org/abs/1608.06993) where inputs into previous layers are concatenated together.
- [This blog has an in-depth overview](https://chatbotslife.com/resnets-highwaynets-and-densenets-oh-my-9bb15918ee32)
If you do decide to implement something extra, clearly describe it in the "Extra Credit Description" cell below.
### What we expect
At the very least, you should be able to train a ConvNet that gets at least 70% accuracy on the validation set. This is just a lower bound - if you are careful it should be possible to get accuracies much higher than that! Extra credit points will be awarded for particularly high-scoring models or unique approaches.
You should use the space below to experiment and train your network.
Have fun and happy training!
```
# Train your model here, and make sure the output of this cell is the accuracy of your best model on the
# train, val, and test sets. Here's some code to get you started. The output of this cell should be the training
# and validation accuracy on your best model (measured by validation accuracy).
model = None
loss_fn = None
optimizer = None
train(model, loss_fn, optimizer, num_epochs=1)
check_accuracy(model, loader_val)
```
### Describe what you did
In the cell below you should write an explanation of what you did, any additional features that you implemented, and any visualizations or graphs that you make in the process of training and evaluating your network.
Tell us here!
## Test set -- run this only once
Now that we've gotten a result we're happy with, we test our final model on the test set (which you should store in best_model). This would be the score we would achieve on a competition. Think about how this compares to your validation set accuracy.
```
best_model = None
check_accuracy(best_model, loader_test)
```
## Going further with PyTorch
The next assignment will make heavy use of PyTorch. You might also find it useful for your projects.
Here's a nice tutorial by Justin Johnson that shows off some of PyTorch's features, like dynamic graphs and custom NN modules: http://pytorch.org/tutorials/beginner/pytorch_with_examples.html
If you're interested in reinforcement learning for your final project, this is a good (more advanced) DQN tutorial in PyTorch: http://pytorch.org/tutorials/intermediate/reinforcement_q_learning.html
| github_jupyter |
<script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
<script>
window.dataLayer = window.dataLayer || [];
function gtag(){dataLayer.push(arguments);}
gtag('js', new Date());
gtag('config', 'UA-59152712-8');
</script>
# Code and Physical Units
## GRHD Units in terms of EOS
$\newcommand{\rhoCode}{{\tilde{\rho}}}$
$\newcommand{\MCode}{{\tilde{M}}}$ $\newcommand{\rCode}{{\tilde{r}}}$ $\newcommand{\PCode}{{\tilde{P}}}$$\newcommand{\tCode}{{\tilde{t}}}$$\newcommand{\Mfid}{{M_{\rm fid}}}$$\newcommand{\MfidBar}{\bar{M}_{\rm fid}}$$\newcommand{\Mbar}{\bar{M}}$
$\newcommand{\rBar}{\bar{r}}$$\newcommand{\tBar}{\bar{t}}$
In GRHD, we can set an equation of state of the form
\begin{equation}
P = K\rho^{1+1/n}
\end{equation}
Taking $c_s^2 = \partial P/\partial \rho = (1+1/n) K\rho^{1/n}$. This gives for some fidicial $\rho_0$
\begin{equation}
c_{s,0}^2 = \left(1 + \frac 1 n\right)K\rho_0^{1/n}.
\end{equation}
Selecting $c_s^2 = c^2\left(1 + 1/n\right)$, we have
\begin{equation}
\rho_0 = \left(\frac {c^2}{K}\right)^n
\end{equation}
This is equivalent to setting the isothermal sound speed to $c$. With this definition of $\rho_0$, we can write
\begin{equation}
P = \rho_0c^2\left(\frac{\rho}{\rho_0}\right)^{1+1/n}
\end{equation}
which allows us to define the dimensionless density $\rhoCode = \rho/\rho_0$ and dimensionless pressure $\PCode = P/\rho_0 c^2$
\begin{equation}
\PCode = \rhoCode^{1+1/n},
\end{equation}
where we adopt code units where $c=1$. These dimensionless pressure and density are in $G=c=1$ units and can be used in GRHD code including inclusion in the spacetime solver via $T_{\mu\nu}$. Note that this sets $K=1$ in these units.
To find a dimensionless mass, $\MCode$, dimensionless distance, $\rCode$, and dimensionless time, $\tCode$, we note
$GM/rc^2$ is dimensionless
\begin{equation}
\frac{GM}{rc^2} = \frac{G\rho_0 r^2}{c^2} = \frac{Gc^{2n-2}}{K^n}r^2 \rightarrow \rCode = \frac{\sqrt{G}c^{n-1}}{K^{n/2}} r = \frac r {r_0},
\end{equation}
where $r_0 = K^{n/2}/\sqrt{G}c^{n-1}$. Then
\begin{eqnarray}
\tCode &=& \frac{t}{t_0} = \frac{t}{r_0/c} = \frac{\sqrt{G}c^n}{K^{n/2}} t \\
\MCode &=& \frac{M}{M_0} = \frac{M}{\rho_0 r_0^3} = M\frac{K^n}{c^{2n}}\frac{G^{3/2}c^{3(n-1)}}{K^{3n/2}} = \frac{G^{3/2}c^{n-3}}{K^{n/2}} M,
\end{eqnarray}
Hence, we have
\begin{eqnarray}
\rho_0 &=& \left(\frac{K}{c^2}\right)^n\\
r_0 &=& \frac{c^{n+1}}{\sqrt{G}K^{n/2}}\\
t_0 &=& \frac{c^{n}}{\sqrt{G}K^{n/2}}\\
M_0 &=& \frac{c^{n+3}}{G^{3/2}K^{n/2}}
\end{eqnarray}
## Mapping to SENR or any NR code
So we will need a $\Mfid$ which is define such that the (SENR) code units $\MfidBar = 1$ or in other words in SENR codes units:
\begin{equation}
\Mbar = \frac{M}{\Mfid}
\end{equation}
In these units:
\begin{eqnarray}
\rBar &=& \frac{c^2}{G\Mfid} r\\
\tBar &=& \frac{c^3}{G\Mfid} t
\end{eqnarray}
At some level $\Mfid$ is arbitrary, so we can select $M_0 = \Mfid$. In this case, this means that $\rBar = \rCode$, $\tBar = \tCode$, and $\Mbar = \MCode$, which fixes all the quantities. This comes at a cost the $\bar{M}_{\rm ADM}$ is not something nice like 1 or 2, but the choice is consistent.
### Zach's comments:
Sound speed $c_s$ is defined as
$$\frac{\partial P}{\partial \rho} = c_s^2,$$
so if we have a polytropic EOS, where
$$P = K \rho^{(1 + 1/n)},$$
then
\begin{align}
\frac{\partial P}{\partial \rho} &= c_s^2 \\
&= \left(1 + \frac{1}{n}\right) K \rho^{1/n}.
\end{align}
Let's adopt the notation
$$[\rho] = \text{"the units of $\rho$"}$$
Using this notation and the fact that $n$ is dimensionless, the above expression implies
\begin{align}
\left[\rho^{1/n}\right] &= \left[\frac{c_s^2}{K}\right] \\
\implies \left[\rho\right] &= \left[\frac{c_s^2}{K}\right]^n
\end{align}
I think you found the inverse to be true.
# TOV Solver as illustration
The TOV equations are
\begin{eqnarray}
\frac{dP}{dr} &=& -\mu\frac{GM}{r^2}\left(1 + \frac P {\mu c^2}\right)\left(1 + \frac {4\pi r^3 P}{Mc^2}\right)\left(1 - \frac {2GM}{rc^2}\right)^{-1}\\
\frac{dM}{dr} &=& 4\pi \mu r^2,
\end{eqnarray}
Here $M$ is the rest mass measure by a distant observer when we take $r\rightarrow \infty$. Note this is different from the mass measured by integrating the density over the volume
\begin{equation}
M' = \int_0^{\infty} \frac{4\pi r^2\mu}{\sqrt{1 - \frac {2 GM}{rc^2}}} dr
\end{equation}
Additionally and annoyingly, $\mu = \rho h$ is the mass-energy density. A lot of the literature uses $\rho$ for this, which is incredibly annoying.
$\newcommand{\muCode}{{\tilde{\mu}}}$
In dimensionless units they are
\begin{eqnarray}
\frac{d\PCode}{d\rCode} &=& -\frac {\left(\muCode + \PCode\right)\left(\MCode + 4\pi \rCode^3 \PCode\right)}{\rCode^2\left(1 - \frac {2\MCode}{\rCode}\right)}\\
\frac{d\MCode}{d\rCode} &=& 4\pi \muCode\rCode^2
\end{eqnarray}
At this point, we need to discuss how to numerically integrate these models. First we pick a central baryonic mass density $\rhoCode_{0,c}$, then we compute a central pressure $\PCode_c$ and central mass-energy density $\muCode_c$. At $\rCode=0$, we assume that $\muCode=\muCode_c$ is a constant and so
\begin{eqnarray}
\frac{d\PCode}{d\rCode} &=& -\frac {\left(\muCode_c + \PCode_c\right)\left(\MCode(\rCode \ll 1) + 4\pi \rCode^3 \PCode_c\right)}{\rCode^2\left(1 - \frac {2\MCode(\rCode \ll 1)}{\rCode}\right)}\\
\frac{d\MCode}{d\rCode} &=& 4\pi \muCode_c\rCode^2 \rightarrow \MCode(\rCode \ll 1) = \frac{4\pi}{3} \muCode_c \rCode^3
\end{eqnarray}
## Another dimensionless prescription
Let consider an alternative formulation where rather than setting $K=1$, we set the characteristic mass $\Mfid = M_0$. In this case,
\begin{eqnarray}
r_0 &=& \frac{GM_0}{c^2} \\
t_0 &=& \frac{GM_0}{c^3} \\
\rho_0 &=& \frac{M_0}{r_0^3} = \frac{c^6}{G^3 M_0^2} = 6.17\times 10^{17}\left(\frac {M_0} {1 M_{\odot}}\right)^{-2}
\end{eqnarray}
In this case we can define $\rhoCode = \rho/\rho_0$, $\rCode = r/r_0$, $t_0 = t/t_0$. The only remaining thing to do is to define $\PCode$. Lets define $P_0'$ to be the pressure in dimensionful units at $\rho_0$ (density in units of $1/M_0^2$):
\begin{equation}
P = P_0'\rhoCode^{1+1/n} \rightarrow P_0' = K\rho_0^{1+1/n},
\end{equation}
So defining $P_0 = \rho_0 c^2$, we have
\begin{equation}
\PCode = \frac{P}{P_0} = \frac{K\rho_0^{1/n}}{c^2}\rhoCode^{1+1/n} = \PCode_0\rhoCode^{1+1/n}
\end{equation}
If we take $K=1$ and define $\rho_0$ such that the $\PCode_0 = 1$, we recover the results above.
Finally for $\muCode = \rhoCode + \PCode/n$
## metric for TOV equation
The metric for the TOV equation (taken) from wikipedia is
\begin{equation}
ds^2 = - c^2 e^\nu dt^2 + \left(1 - \frac{2GM}{rc^2}\right)^{-1} dr^2 + r^2 d\Omega^2
\end{equation}
where $M$ is defined as above, the mass as measured by a distant observer. The equation for $\nu$ is
\begin{equation}
\frac{d\nu}{dr} = -\left(\frac {2}{P +\mu}\right)\frac{dP}{dr}
\end{equation}
with the boundary condition
\begin{equation}
\exp(\nu) = \left(1-\frac {2Gm(R)}{Rc^2}\right)
\end{equation}
Lets write this in dimensionless units:
\begin{equation}
ds^2 = \exp(\nu) d\tCode^2 - \left(1 - \frac{2\MCode}{\rCode}\right)^{-1} d\rCode^2 + \rCode^2 d\Omega^2
\end{equation}
\begin{equation}
\frac{d\nu}{d\rCode} = -\left(\frac {2}{\PCode +\muCode}\right)\frac{d\PCode}{d\rCode}
\end{equation}
and BC:
\begin{equation}
\exp(\nu) = \left(1-\frac {2\MCode}{\rCode}\right)
\end{equation}
```
import sys
import numpy as np
import scipy.integrate as si
import math
import matplotlib.pyplot as pl
n = 1.
rho_central = 0.129285
P0 = 1. # ZACH NOTES: CHANGED FROM 100.
gamma = 1. + 1./n
gam1 = gamma - 1.
def pressure( rho) :
return P0*rho**gamma
def rhs( r, y) :
# In \tilde units
#
P = y[0]
m = y[1]
nu = y[2]
rbar = y[3]
rho = (P/P0)**(1./gamma)
mu = rho + P/gam1
dPdr = 0.
drbardr = 0.
if( r < 1e-4 or m <= 0.) :
m = 4*math.pi/3. * mu*r**3
dPdr = -(mu + P)*(4.*math.pi/3.*r*mu + 4.*math.pi*r*P)/(1.-8.*math.pi*mu*r*r)
drbardr = 1./(1. - 8.*math.pi*mu*r*r)**0.5
else :
dPdr = -(mu + P)*(m + 4.*math.pi*r**3*P)/(r*r*(1.-2.*m/r))
drbardr = 1./(1. - 2.*m/r)**0.5*rbar/r
dmdr = 4.*math.pi*r*r*mu
dnudr = -2./(P + mu)*dPdr
return [dPdr, dmdr, dnudr, drbardr]
def integrateStar( P, showPlot = False, dumpData = False, compareFile="TOV/output_EinsteinToolkitTOVSolver.txt") :
integrator = si.ode(rhs).set_integrator('dop853')
y0 = [P, 0., 0., 0.]
integrator.set_initial_value(y0,0.)
dr = 1e-5
P = y0[0]
PArr = []
rArr = []
mArr = []
nuArr = []
rbarArr = []
r = 0.
while integrator.successful() and P > 1e-9*y0[0] :
P, m, nu, rbar = integrator.integrate(r + dr)
r = integrator.t
dPdr, dmdr, dnudr, drbardr = rhs( r+dr, [P,m,nu,rbar])
dr = 0.1*min(abs(P/dPdr), abs(m/dmdr))
dr = min(dr, 1e-2)
PArr.append(P)
rArr.append(r)
mArr.append(m)
nuArr.append(nu)
rbarArr.append( rbar)
M = mArr[-1]
R = rArr[-1]
nuArr_np = np.array(nuArr)
# Rescale solution to nu so that it satisfies BC: exp(nu(R))=exp(nutilde-nu(r=R)) * (1 - 2m(R)/R)
# Thus, nu(R) = (nutilde - nu(r=R)) + log(1 - 2*m(R)/R)
nuArr_np = nuArr_np - nuArr_np[-1] + math.log(1.-2.*mArr[-1]/rArr[-1])
rArrExtend_np = 10.**(np.arange(0.01,5.0,0.01))*rArr[-1]
rArr.extend(rArrExtend_np)
mArr.extend(rArrExtend_np*0. + M)
PArr.extend(rArrExtend_np*0.)
phiArr_np = np.append( np.exp(nuArr_np), 1. - 2.*M/rArrExtend_np)
rbarArr.extend( 0.5*(np.sqrt(rArrExtend_np*rArrExtend_np-2.*M*rArrExtend_np) + rArrExtend_np - M))
# Appending a Python array does what one would reasonably expect.
# Appending a numpy array allocates space for a new array with size+1,
# then copies the data over... over and over... super inefficient.
mArr_np = np.array(mArr)
rArr_np = np.array(rArr)
PArr_np = np.array(PArr)
rbarArr_np = np.array(rbarArr)
rhoArr_np = (PArr_np/P0)**(1./gamma)
confFactor_np = rArr_np/rbarArr_np
#confFactor_np = (1.0 / 12.0) * np.log(1.0/(1.0 - 2.0*mArr_np/rArr_np))
Grr_np = 1.0/(1.0 - 2.0*mArr_np/rArr_np)
Gtt_np = phiArr_np
if( showPlot) :
r,rbar,rprop,rho,m,phi = np.loadtxt( compareFile, usecols=[0,1,2,3,4,5],unpack=True)
pl.plot(rArr_np[rArr_np < r[-1]], rbarArr_np[rArr_np < r[-1]],lw=2,color="black")
#pl.plot(r, rbar, lw=2,color="red")
pl.show()
if( dumpData) :
np.savetxt( "output.txt", np.transpose([rArr_np,rhoArr_np,PArr_np,mArr_np,phiArr_np,confFactor_np,rbarArr_np]), fmt="%.15e")
np.savetxt( "metric.txt", np.transpose([rArr_np, Grr_np, Gtt_np]),fmt="%.15e")
# np.savetxt( "output.txt", zip(rArr,rhoArr,mArr,phiArr), fmt="%12.7e")
# return rArr[-1], mArr[-1], phiArr[-1]
return R, M
mass = []
radius = []
R_TOV,M_TOV = integrateStar(pressure(rho_central), showPlot=True, dumpData=True)
print("Just generated a TOV star with r= "+str(R_TOV)+" , m = "+str(M_TOV)+" , m/r = "+str(M_TOV/R_TOV)+" .")
#for rho0 in np.arange(0.01, 1., 0.01):
# r,m = integrateStar(pressure(rho0))
# mass.append(m)
# radius.append(r)
#print(mass, radius)
#pl.clf()
#pl.plot(radius,mass)
#pl.show()
# Generate the Sedov Problem
rArr_np = np.arange(0.01,5.,0.01)
rbarArr_np = rArr_np
rhoArr_np = np.ones(rArr_np.size)*0.1
mArr_np = 4.*np.pi/3.*rArr_np**3*rhoArr_np
PArr_np = rhoArr_np*1e-6
PArr_np[rArr_np < 0.5] = 1e-2
phiArr_np = np.ones(rArr_np.size)
confFactor_np = rArr_np/rbarArr_np
if sys.version_info[0] < 3:
np.savetxt( "sedov.txt", zip(rArr_np,rhoArr_np,PArr_np,mArr_np,phiArr_np,confFactor_np,rbarArr_np), fmt="%.15e")
else:
np.savetxt( "sedov.txt", list(zip(rArr_np,rhoArr_np,PArr_np,mArr_np,phiArr_np,confFactor_np,rbarArr_np)), fmt="%.15e")
pl.semilogx(rArr_np, rhoArr_np)
pl.show()
```
## Convert metric to be in terms of ADM quantities
Above, the line element was written:
$$
ds^2 = - c^2 e^\nu dt^2 + \left(1 - \frac{2GM}{rc^2}\right)^{-1} dr^2 + r^2 d\Omega^2.
$$
In terms of $G=c=1$ units adopted by NRPy+, this becomes:
$$
ds^2 = - e^\nu dt^2 + \left(1 - \frac{2M}{r}\right)^{-1} dr^2 + r^2 d\Omega^2.
$$
The ADM 3+1 line element for this diagonal metric in spherical coordinates is given by:
$$
ds^2 = (-\alpha^2 + \beta_k \beta^k) dt^2 + \gamma_{rr} dr^2 + \gamma_{\theta\theta} d\theta^2+ \gamma_{\phi\phi} d\phi^2,
$$
from which we can immediately read off the ADM quantities:
\begin{align}
\alpha &= e^{\nu/2} \\
\beta^k &= 0 \\
\gamma_{rr} &= \left(1 - \frac{2M}{r}\right)^{-1}\\
\gamma_{\theta\theta} &= r^2 \\
\gamma_{\phi\phi} &= r^2 \sin^2 \theta \\
\end{align}
## Convert to Cartesian coordinates
The above metric is given in spherical coordinates and we need everything in Cartesian coordinates. Given this the
transformation to Cartesian coordinates is
\begin{equation}
g_{\mu\nu} = \Lambda^{\mu'}_{\mu} \Lambda^{\nu'}_{\nu} g_{\mu'\nu'},
\end{equation}
where $\Lambda^{\mu'}_{\mu}$ is the Jacobian defined as
\begin{equation}
\Lambda^{\mu'}_{\mu} = \frac{\partial x'^{\mu'}}{\partial x^{\mu}}
\end{equation}
In this particular case $x'$ is in spherical coordinates and $x$ is in Cartesian coordinates.
```
import sympy as sp # SymPy: The Python computer algebra package upon which NRPy+ depends
import NRPy_param_funcs as par # NRPy+: parameter interface
from outputC import outputC # NRPy+: Basic C code output functionality
import indexedexp as ixp # NRPy+: Symbolic indexed expression (e.g., tensors, vectors, etc.) support
import reference_metric as rfm # NRPy+: Reference metric support
# The ADM & BSSN formalisms only work in 3D; they are 3+1 decompositions of Einstein's equations.
# To implement axisymmetry or spherical symmetry, simply set all spatial derivatives in
# the relevant angular directions to zero; DO NOT SET DIM TO ANYTHING BUT 3.
# Step 0: Set spatial dimension (must be 3 for BSSN)
DIM = 3
# Set the desired *output* coordinate system to Cylindrical:
par.set_parval_from_str("reference_metric::CoordSystem","Cartesian")
rfm.reference_metric()
CoordType_in = "Spherical"
r_th_ph_or_Cart_xyz_of_xx = []
if CoordType_in == "Spherical":
r_th_ph_or_Cart_xyz_of_xx = rfm.xxSph
elif CoordType_in == "Cartesian":
r_th_ph_or_Cart_xyz_of_xx = rfm.xx_to_Cart
Jac_dUSphorCart_dDrfmUD = ixp.zerorank2()
for i in range(DIM):
for j in range(DIM):
Jac_dUSphorCart_dDrfmUD[i][j] = sp.diff(r_th_ph_or_Cart_xyz_of_xx[i],rfm.xx[j])
Jac_dUrfm_dDSphorCartUD, dummyDET = ixp.generic_matrix_inverter3x3(Jac_dUSphorCart_dDrfmUD)
betaU = ixp.zerorank1()
gammaDD = ixp.zerorank2()
gammaSphDD = ixp.zerorank2()
grr, gthth, gphph = sp.symbols("grr gthth gphph")
gammaSphDD[0][0] = grr
gammaSphDD[1][1] = gthth
gammaSphDD[2][2] = gphph
betaSphU = ixp.zerorank1()
for i in range(DIM):
for j in range(DIM):
betaU[i] += Jac_dUrfm_dDSphorCartUD[i][j] * betaSphU[j]
for k in range(DIM):
for l in range(DIM):
gammaDD[i][j] += Jac_dUSphorCart_dDrfmUD[k][i]*Jac_dUSphorCart_dDrfmUD[l][j] * gammaSphDD[k][l]
outputC([gammaDD[0][0], gammaDD[0][1], gammaDD[0][2], gammaDD[1][1], gammaDD[1][2], gammaDD[2][2]],
["mi.gamDDxx", "mi.gamDDxy", "mi.gamDDxz", "mi.gamDDyy", "mi.gamDDyz","mi.gamDDzz"], filename="NRPY+spherical_to_Cartesian_metric.h")
```
| github_jupyter |
```
import pandas as pd
import seaborn as sns
import numpy as np
import altair as alt
import matplotlib.pyplot as plt
import os
import sys
from ocp_table_tpot.globals import Globals as gd
from tpot import TPOTRegressor
sys.path.insert(0,'..')
from src.models.model import HistoricalMedian,XGBoost,LinearModel,RF,KNN,SVM,mase,TimeSeriesSplitImproved
from sklearn.linear_model import ElasticNet, Lasso, BayesianRidge, LassoLarsIC,RANSACRegressor
from sklearn.ensemble import RandomForestRegressor, GradientBoostingRegressor,ExtraTreesRegressor
from sklearn.kernel_ridge import KernelRidge
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import RobustScaler,MinMaxScaler,PolynomialFeatures,StandardScaler
from sklearn.base import BaseEstimator, TransformerMixin, RegressorMixin, clone
from sklearn.model_selection import KFold, cross_val_score, train_test_split,cross_val_predict
from skgarden.quantile import RandomForestQuantileRegressor
from sklearn.metrics import mean_squared_error,make_scorer
from sklearn.preprocessing import FunctionTransformer
from copy import copy
from tpot.builtins import StackingEstimator
from lightgbm import LGBMRegressor
import xgboost as xgb
import lightgbm as lgb
from sklearn.pipeline import make_pipeline, make_union
from catboost import CatBoostRegressor,Pool,cv
sys.path.insert(0,'..')
from src.models.model import HistoricalMedian,XGBoost,LinearModel,RF,KNN,SVM,mase
from src.data.make_dataset import DROPCOLS
```
## Load data
```
df_tsfresh = pd.read_pickle(f'../data/processed/train_test_tsfresh.pkl').reset_index(level = 0)
data_dict = pd.read_pickle(f'../data/processed/data_dict_all.pkl')
#df_flat = df_tsfresh.pivot(df_tsfresh)
#df_flat.columns = ['_'.join(col).strip() for col in df_flat.columns.values]
year = 2019
X = data_dict[year]['X_train_tsclean'].astype(float)
X_test = data_dict[year]['X_test_ts'].astype(float)
y = data_dict[year]['y_train_tsclean'].astype(float).loc[:,['rougher.output.recovery','final.output.recovery']]
mask = data_dict[year]['mask']
X = X[mask]
y = y[mask]
print(f'{X.shape}')
```
# Keras NNet solution
```
# bagging mlp ensemble on blobs dataset
import numpy as np
from matplotlib import pyplot as plt
from IPython.display import clear_output
from sklearn.datasets.samples_generator import make_blobs
from sklearn.utils import resample
from sklearn.metrics import accuracy_score
from keras.utils import to_categorical
from keras.models import Sequential,Model
from keras.regularizers import l1,l2
from keras.optimizers import Adam,SGD
from keras.initializers import random_uniform,glorot_uniform,he_uniform
from keras.layers import Dense,Dropout,BatchNormalization,Activation,ReLU,Input,Concatenate
from keras.callbacks import EarlyStopping, ModelCheckpoint,Callback,ReduceLROnPlateau,History
from keras.wrappers.scikit_learn import KerasRegressor
import keras.backend as K
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import QuantileTransformer
from sklearn.decomposition import PCA
from matplotlib import pyplot
from numpy import mean
from numpy import std
import numpy
from numpy import array
from numpy import argmax
from keras_tqdm import TQDMNotebookCallback
import numpy as np
import tensorflow as tf
from keras.utils import plot_model
def huber_loss(y_true, y_pred, clip_delta=1.0):
error = y_true - y_pred
cond = tf.keras.backend.abs(error) < clip_delta
squared_loss = 0.5 * tf.keras.backend.square(error)
linear_loss = clip_delta * (tf.keras.backend.abs(error) - 0.5 * clip_delta)
return tf.where(cond, squared_loss, linear_loss)
def tilted_loss( y, f,q = 0.55):
e = (y - f)
return K.mean(K.maximum(q * e, (q - 1) * e),
axis=-1)
def huber_loss_mean(y_true, y_pred, clip_delta=1.0):
return tf.keras.backend.mean(huber_loss(y_true, y_pred, clip_delta))
def my_loss(y_true,y_pred):
#naive_forecast_error = K.mean(K.abs(y_true[1:,:] - y_true[:-1,:]))
l = K.abs(y_true - y_pred)
r = K.variable([[0.25],[0.75]])
error = K.dot(l,r)
return error
def create_model(optimizer=Adam(lr=7e-3),
seed = 12,
dropout=0.35,input_dim = 332):
kernel_initializer=he_uniform(seed=seed)
main_input = Input(shape = (input_dim,),name = 'main_input')
d1 = Dense(768,input_dim = input_dim,kernel_initializer=kernel_initializer,kernel_regularizer=l1(4e0))(main_input)
d1 = BatchNormalization()(d1)
d1 = Activation('relu')(d1)
#d1 = Dropout(dropout/2)(d1)
d2 = Dense(256,kernel_initializer=kernel_initializer,activation = None,kernel_regularizer=l1(3e0))(d1)
d2 = BatchNormalization()(d2)
d2 = Activation('relu')(d2)
#d2 = Dropout(dropout)(d2)
d4 = Dense(32,kernel_initializer=kernel_initializer,activation = None,kernel_regularizer=l1(2e0))(d2)
d4 = BatchNormalization()(d4)
d4 = Activation('relu')(d4)
#d4 = Dropout(dropout)(d4)
output_rougher = Dense(1,activation=None,kernel_regularizer=l1(1e-3))(d4)
output_rougher = ReLU(max_value = 100,name = 'rough')(output_rougher)
concat_layer = Concatenate()([d4,output_rougher])
concat_layer =Dropout(dropout)(concat_layer)
output_final = Dense(1,activation = None,kernel_initializer =kernel_initializer)(concat_layer)
output_final = ReLU(max_value=100,name = 'final')(output_final)
model = Model(inputs=main_input, outputs=[output_rougher, output_final])
model.compile(loss=['mae','mae'],optimizer=optimizer, metrics=['mae'],loss_weights = [0.5,0.75])
return model
# wrap the model using the function you created
reg = KerasRegressor(build_fn=create_model,input_dim = X.shape[1],verbose=0)
reg = create_model(input_dim = X.shape[1])
#print(reg)
#scaler = make_pipeline(QuantileTransformer(output_distribution='normal',n_quantiles = 5000))
#scaler = make_pipeline(PCA(whiten=True))
scaler = make_pipeline(RobustScaler())
target_scaler = make_pipeline(QuantileTransformer(output_distribution='normal'),StandardScaler())
hr = History()
#clr = CyclicLR(base_lr = 8e-4,max_lr = 5e-2,step_size=4802, mode='triangular2')
#callbacks = [EarlyStopping(monitor='val_loss', patience=40),TQDMNotebookCallback(leave_inner=False),
# clr,hr]
# params = {"epochs":150,
# "verbose":0,
# "batch_size":128,
# "callbacks":callbacks}
import sys
import os
reg.summary()
#plot_model(reg)
# 2.08 with custom loss, bs = 128, seed = 123, random_uniform, mode = 'triangular2'
bs = 64
seed = 12
fold_n = 1
model = reg
scores = []
df_test_all =[]
#fig,ax = plt.subplots(figsize = (20,16),nrows = 1)
scaler.fit(X)
history = [None]
fpath = f'./keras-ch/multi_2target.h5'
#clr_fn = lambda x: 0.9**x # 54*10*4
#clr_fn = lambda x: 0.5*(1+np.sin(x*np.pi/2.))
#clr = CyclicLR(base_lr = 7e-3,max_lr = 1e-2,step_size=54*8*4, mode='triangular',scale_fn=clr_fn, scale_mode='cycle')
mc = ModelCheckpoint(filepath=fpath, monitor='val_final_mean_absolute_error',save_best_only=True,verbose =1)
M = 5 # number of snapshots
nb_epoch = T = 600 # number of epochs
alpha_zero = 1.5e-2 # initial learning rate
model_prefix = 'mod_ts'
snapshot = SnapshotCallbackBuilder(T, M, alpha_zero)
cbcks = snapshot.get_callbacks(model_prefix=model_prefix,monitor_metric ="val_final_mean_absolute_error")
cbcks = cbcks + [hr,TQDMNotebookCallback()]
train_index = X.index < pd.to_datetime("2018-04-04").tz_localize('UTC')
valid_index = X.index > pd.to_datetime("2018-04-04").tz_localize('UTC')
X_train, X_valid = X.iloc[train_index], X.iloc[valid_index]
y_train, y_valid = y.iloc[train_index], y.iloc[valid_index]
preds_all_base = np.empty_like(y_valid)
true_all =np.empty_like(y_valid)
mask_train,mask_valid = mask.iloc[train_index,],mask.iloc[valid_index,]
X_train, X_valid = scaler.transform(X_train), scaler.transform(X_valid)
# Do the base
params = {"epochs":nb_epoch,
"verbose":0,
"batch_size":bs,
"callbacks":cbcks}
params["validation_data"] = (X_valid, {'rough':y_valid.values[:,0],'final':y_valid.values[:,1]})
model = create_model(input_dim = X_train.shape[1])
history[0] = model.fit(X_train,{'rough':y_train.values[:,0],'final':y_train.values[:,1]},**params)
pd.DataFrame(history[0].history).drop('lr',axis = 1).plot(figsize = (10,10),logy=True)
def calculate_weighted_pred(x,prediction_weights):
weighted_predictions = np.zeros((x[0].shape[0], 2), dtype='float32')
for weight, prediction in zip(prediction_weights, x):
weighted_predictions += weight * prediction
yPred = weighted_predictions
return weighted_predictions
# Define a helper function for calculating the average of snapshots:
preds = []
preds_test = []
model = create_model(input_dim = X_train.shape[1])
model_filenames = [f'./weights/mod_ts-{i+1}.h5' for i in range(5)]
model_filenames.append('./weights/mod_ts-Best.h5')
for fn in model_filenames:
model.load_weights(fn)
yTrue = y_valid.values
yPred = np.hstack(model.predict(X_valid, batch_size=64))
X_test_sc = scaler.transform(X_test)
yPredTest = np.hstack(model.predict(X_test_sc, batch_size=64))
mase_sc = 0.25 * mase(yPred[:,0],yTrue[:,0]) + 0.75 * mase(yPred[:,1],yTrue[:,1])
print(f'MASE: {mase_sc}')
preds.append(yPred)
preds_test.append(yPredTest)
print("Obtained predictions from model with weights = %s" % (fn))
```
### optimize weights
```
from scipy.optimize import minimize
def loss_func(weights):
''' scipy minimize will pass the weights as a numpy array '''
final_prediction = np.zeros((X_valid.shape[0], 2), dtype='float32')
for weight, prediction in zip(weights, preds):
final_prediction += weight * prediction
return 0.25 * mase(final_prediction[:,0],yTrue[:,0]) + 0.75 * mase(final_prediction[:,1],yTrue[:,1])
best_mase = 100.0
best_weights = None
# Parameters for optimization
constraints = ({'type': 'eq', 'fun':lambda w: 1 - sum(w)})
bounds = [(0, 1)] * len(preds)
NUM_TESTS = 50
# Check for NUM_TESTS times
for iteration in range(NUM_TESTS):
# Random initialization of weights
prediction_weights = np.random.random(len(model_filenames))
# Minimise the loss
result = minimize(loss_func, prediction_weights, method='SLSQP', bounds=bounds, constraints=constraints)
print('Best Ensemble Weights: {weights}'.format(weights=result['x']))
weights = result['x']
weighted_predictions = np.zeros((X_valid.shape[0], 2), dtype='float32')
# Calculate weighted predictions
for weight, prediction in zip(weights, preds):
weighted_predictions += weight * prediction
yPred = weighted_predictions
yTrue = y_valid.values
sc = 0.25 * mase(yPred[:,0],yTrue[:,0]) + 0.75 * mase(yPred[:,1],yTrue[:,1])
print(sc)
# Save current best weights
if sc > best_mase:
best_acc = sc
best_weights = weights
print()
# 0.00000000e+00 1.53911667e-15 4.63643997e-02 0.00000000e+00
# 7.57377707e-02 8.77897830e-01]
w = np.array([1,1,1,1,1,1])
w = w / np.sum(w)
preds_test_w = calculate_weighted_pred(preds_test,w)
df_test = pd.DataFrame({"preds_r":preds_test_w[:,0].reshape(-1,),"preds_f":preds_test_w[:,1].reshape(-1,),'date':X_test.index})
preds_all_base = calculate_weighted_pred(preds,w)
df = pd.DataFrame({"preds_r":preds_all_base[:,0],"preds_f":preds_all_base[:,1],"true_r":y_valid.values[:,0],"true_f":y_valid.values[:,1]},index = y_valid.index)
df.plot(figsize =(10,6),ylim = (40,100),alpha=0.5)
#df_oof.columns = ['rougher.output.recovery','final.output.recovery']
#df_oof.to_pickle('./keras-ch/2-17-keras-2l-96-16-6-train.pkl')
df_test.to_pickle('./keras-ch/2-17-keras-2l-96-16-6-test.pkl')
#df_test[['preds_r','preds_f']].plot(figsize =(20,10),ylim = (40,100),style = ['o','o'],alpha=0.9)
#df_test[['preds_r','preds_f']].plot()
#df_test.plot(figsize =(20,7))
#df_test\
plt.plot(preds_test[5])
#df_test[['preds_r','preds_f']].plot(figsize =(20,10),ylim = (40,100),style = ['o','o'],alpha=0.9)
df_test[['preds_r','preds_f']].plot(ylim = (38,100))
#df_test.plot(figsize =(20,7))
#df_test
df_keras_su = df_test[["preds_r","preds_f","date"]].rename(index=str,columns = {"preds_r":"rougher.output.recovery","preds_f":"final.output.recovery"})
df_keras_su['date'] = df_keras_su['date'].dt.strftime('%Y-%m-%dT%H:%M:%SZ')
df_keras_su.set_index('date')
df_keras_su.to_csv('../results/keras_best.csv')
i=0
a =history[i]
hdf = pd.DataFrame(a.history)
hdf.plot(figsize = (10,10),logy=True)
clr_df = pd.DataFrame(clr.history)
fig,ax = plt.subplots(figsize =(10,10))
clr_df.plot(x='iterations',y='lr',logy=True,ax=ax)
ax.vlines(x = np.array([21,24,75,91])*146,ymin=5e-3,ymax = 1e-2)
#clr_df.groupby('batch').count()
95*146
model = create_model(input_dim = X.shape[1])
clr = CyclicLR(base_lr = 2e-4,max_lr = 5e-2,step_size=170, mode='triangular')
# Ensure that number of epochs = 1 when calling fit()
#model.fit(X, y, epochs=1, batch_size=32, callbacks=[clr])
model.fit(X_train,{'rough':y_train.values[:,0],'final':y_train.values[:,1]}, epochs=1, batch_size=64,callbacks = [clr])
#lr_callback.plot_schedule()
clr_df = pd.DataFrame(clr.history)
clr_df.plot(x='iterations',y='lr',logy=True,figsize =(10,10))
import altair as alt
alt.renderers.enable('notebook')
alt.Chart(data=clr_df,width=1000).mark_line().encode(
x=alt.X('lr', scale=alt.Scale(type='log')),
y='loss')
#clr_df.plot(x='lr',y='loss',figsize =(10,10),logx=True)
# 7e-3 1.5e-2
from keras.callbacks import *
class CyclicLR(Callback):
"""This callback implements a cyclical learning rate policy (CLR).
The method cycles the learning rate between two boundaries with
some constant frequency, as detailed in this paper (https://arxiv.org/abs/1506.01186).
The amplitude of the cycle can be scaled on a per-iteration or
per-cycle basis.
This class has three built-in policies, as put forth in the paper.
"triangular":
A basic triangular cycle w/ no amplitude scaling.
"triangular2":
A basic triangular cycle that scales initial amplitude by half each cycle.
"exp_range":
A cycle that scales initial amplitude by gamma**(cycle iterations) at each
cycle iteration.
For more detail, please see paper.
# Example
```python
clr = CyclicLR(base_lr=0.001, max_lr=0.006,
step_size=2000., mode='triangular')
model.fit(X_train, Y_train, callbacks=[clr])
```
Class also supports custom scaling functions:
```python
clr_fn = lambda x: 0.5*(1+np.sin(x*np.pi/2.))
clr = CyclicLR(base_lr=0.001, max_lr=0.006,
step_size=2000., scale_fn=clr_fn,
scale_mode='cycle')
model.fit(X_train, Y_train, callbacks=[clr])
```
# Arguments
base_lr: initial learning rate which is the
lower boundary in the cycle.
max_lr: upper boundary in the cycle. Functionally,
it defines the cycle amplitude (max_lr - base_lr).
The lr at any cycle is the sum of base_lr
and some scaling of the amplitude; therefore
max_lr may not actually be reached depending on
scaling function.
step_size: number of training iterations per
half cycle. Authors suggest setting step_size
2-8 x training iterations in epoch.
mode: one of {triangular, triangular2, exp_range}.
Default 'triangular'.
Values correspond to policies detailed above.
If scale_fn is not None, this argument is ignored.
gamma: constant in 'exp_range' scaling function:
gamma**(cycle iterations)
scale_fn: Custom scaling policy defined by a single
argument lambda function, where
0 <= scale_fn(x) <= 1 for all x >= 0.
mode paramater is ignored
scale_mode: {'cycle', 'iterations'}.
Defines whether scale_fn is evaluated on
cycle number or cycle iterations (training
iterations since start of cycle). Default is 'cycle'.
"""
def __init__(self, base_lr=0.001, max_lr=0.006, step_size=2000., mode='triangular',
gamma=1., scale_fn=None, scale_mode='cycle'):
super(CyclicLR, self).__init__()
self.base_lr = base_lr
self.max_lr = max_lr
self.step_size = step_size
self.mode = mode
self.gamma = gamma
if scale_fn == None:
if self.mode == 'triangular':
self.scale_fn = lambda x: 1.
self.scale_mode = 'cycle'
elif self.mode == 'triangular2':
self.scale_fn = lambda x: 1/(2.**(x-1))
self.scale_mode = 'cycle'
elif self.mode == 'exp_range':
self.scale_fn = lambda x: gamma**(x)
self.scale_mode = 'iterations'
else:
self.scale_fn = scale_fn
self.scale_mode = scale_mode
self.clr_iterations = 0.
self.trn_iterations = 0.
self.history = {}
self._reset()
def _reset(self, new_base_lr=None, new_max_lr=None,
new_step_size=None):
"""Resets cycle iterations.
Optional boundary/step size adjustment.
"""
if new_base_lr != None:
self.base_lr = new_base_lr
if new_max_lr != None:
self.max_lr = new_max_lr
if new_step_size != None:
self.step_size = new_step_size
self.clr_iterations = 0.
def clr(self):
cycle = np.floor(1+self.clr_iterations/(2*self.step_size))
x = np.abs(self.clr_iterations/self.step_size - 2*cycle + 1)
if self.scale_mode == 'cycle':
return self.base_lr + (self.max_lr-self.base_lr)*np.maximum(0, (1-x))*self.scale_fn(cycle)
else:
return self.base_lr + (self.max_lr-self.base_lr)*np.maximum(0, (1-x))*self.scale_fn(self.clr_iterations)
def on_train_begin(self, logs={}):
logs = logs or {}
if self.clr_iterations == 0:
K.set_value(self.model.optimizer.lr, self.base_lr)
else:
K.set_value(self.model.optimizer.lr, self.clr())
def on_batch_end(self, epoch, logs=None):
logs = logs or {}
self.trn_iterations += 1
self.clr_iterations += 1
self.history.setdefault('lr', []).append(K.get_value(self.model.optimizer.lr))
self.history.setdefault('iterations', []).append(self.trn_iterations)
for k, v in logs.items():
self.history.setdefault(k, []).append(v)
K.set_value(self.model.optimizer.lr, self.clr())
def create_model(optimizer=Adam(lr=7e-3),
seed = 12,
dropout=0.5,input_dim = 332):
kernel_initializer=random_uniform(seed=seed)
model = Sequential()
model.add(Dense(512,input_dim = input_dim,kernel_initializer=kernel_initializer))
model.add(BatchNormalization())
model.add(Activation('relu'))
model.add(Dropout(dropout))
model.add(Dense(64,kernel_initializer=kernel_initializer))
model.add(BatchNormalization())
model.add(Activation('relu'))
model.add(Dropout(dropout))
model.add(Dense(32,kernel_initializer=kernel_initializer))
model.add(BatchNormalization())
model.add(Activation('relu'))
model.add(Dropout(dropout))
model.add(Dense(16,kernel_initializer=kernel_initializer))
model.add(BatchNormalization())
model.add(Activation('relu'))
model.add(Dropout(dropout))
model.add(Dense(2,kernel_initializer=kernel_initializer))
model.compile(loss='mae',optimizer=optimizer, metrics=['mae'])
return model
import numpy as np
import os
import keras.callbacks as callbacks
from keras.callbacks import Callback
class SnapshotModelCheckpoint(Callback):
"""Callback that saves the snapshot weights of the model.
Saves the model weights on certain epochs (which can be considered the
snapshot of the model at that epoch).
Should be used with the cosine annealing learning rate schedule to save
the weight just before learning rate is sharply increased.
# Arguments:
nb_epochs: total number of epochs that the model will be trained for.
nb_snapshots: number of times the weights of the model will be saved.
fn_prefix: prefix for the filename of the weights.
"""
def __init__(self, nb_epochs, nb_snapshots, fn_prefix='Model'):
super(SnapshotModelCheckpoint, self).__init__()
self.check = nb_epochs // nb_snapshots
self.fn_prefix = fn_prefix
def on_epoch_end(self, epoch, logs={}):
if epoch != 0 and (epoch + 1) % self.check == 0:
filepath = self.fn_prefix + "-%d.h5" % ((epoch + 1) // self.check)
self.model.save_weights(filepath, overwrite=True)
#print("Saved snapshot at weights/%s_%d.h5" % (self.fn_prefix, epoch))
class SnapshotCallbackBuilder:
"""Callback builder for snapshot ensemble training of a model.
Creates a list of callbacks, which are provided when training a model
so as to save the model weights at certain epochs, and then sharply
increase the learning rate.
"""
def __init__(self, nb_epochs, nb_snapshots, init_lr=0.1):
"""
Initialize a snapshot callback builder.
# Arguments:
nb_epochs: total number of epochs that the model will be trained for.
nb_snapshots: number of times the weights of the model will be saved.
init_lr: initial learning rate
"""
self.T = nb_epochs
self.M = nb_snapshots
self.alpha_zero = init_lr
def get_callbacks(self, model_prefix='Model',monitor_metric = None):
"""
Creates a list of callbacks that can be used during training to create a
snapshot ensemble of the model.
Args:
model_prefix: prefix for the filename of the weights.
Returns: list of 3 callbacks [ModelCheckpoint, LearningRateScheduler,
SnapshotModelCheckpoint] which can be provided to the 'fit' function
"""
if not os.path.exists('weights/'):
os.makedirs('weights/')
callback_list = [callbacks.ModelCheckpoint("weights/%s-Best.h5" % model_prefix, monitor=monitor_metric,
save_best_only=True, save_weights_only=True),
callbacks.LearningRateScheduler(schedule=self._cosine_anneal_schedule),
SnapshotModelCheckpoint(self.T, self.M, fn_prefix='weights/%s' % model_prefix)]
return callback_list
def _cosine_anneal_schedule(self, t):
cos_inner = np.pi * (t % (self.T // self.M)) # t - 1 is used when t has 1-based indexing.
cos_inner /= self.T // self.M
cos_out = np.cos(cos_inner) + 1
return float(self.alpha_zero / 2 * cos_out)
```
| github_jupyter |
# TV Script Generation
In this project, you'll generate your own [Seinfeld](https://en.wikipedia.org/wiki/Seinfeld) TV scripts using RNNs. You'll be using part of the [Seinfeld dataset](https://www.kaggle.com/thec03u5/seinfeld-chronicles#scripts.csv) of scripts from 9 seasons. The Neural Network you'll build will generate a new ,"fake" TV script, based on patterns it recognizes in this training data.
## Get the Data
The data is already provided for you in `./data/Seinfeld_Scripts.txt` and you're encouraged to open that file and look at the text.
>* As a first step, we'll load in this data and look at some samples.
* Then, you'll be tasked with defining and training an RNN to generate a new script!
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
# load in data
import helper
data_dir = './data/Seinfeld_Scripts.txt'
text = helper.load_data(data_dir)
```
## Explore the Data
Play around with `view_line_range` to view different parts of the data. This will give you a sense of the data you'll be working with. You can see, for example, that it is all lowercase text, and each new line of dialogue is separated by a newline character `\n`.
```
view_line_range = (0, 10)
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
import numpy as np
print('Dataset Stats')
print('Roughly the number of unique words: {}'.format(len({word: None for word in text.split()})))
lines = text.split('\n')
print('Number of lines: {}'.format(len(lines)))
word_count_line = [len(line.split()) for line in lines]
print('Average number of words in each line: {}'.format(np.average(word_count_line)))
print()
print('The lines {} to {}:'.format(*view_line_range))
print('\n'.join(text.split('\n')[view_line_range[0]:view_line_range[1]]))
```
---
## Implement Pre-processing Functions
The first thing to do to any dataset is pre-processing. Implement the following pre-processing functions below:
- Lookup Table
- Tokenize Punctuation
### Lookup Table
To create a word embedding, you first need to transform the words to ids. In this function, create two dictionaries:
- Dictionary to go from the words to an id, we'll call `vocab_to_int`
- Dictionary to go from the id to word, we'll call `int_to_vocab`
Return these dictionaries in the following **tuple** `(vocab_to_int, int_to_vocab)`
```
import problem_unittests as tests
from collections import Counter
def create_lookup_tables(text):
"""
Create lookup tables for vocabulary
:param text: The text of tv scripts split into words
:return: A tuple of dicts (vocab_to_int, int_to_vocab)
"""
word_counts = Counter(text)
# sorting the words from most to least frequent in text occurrence
sorted_vocab = sorted(word_counts, key=word_counts.get, reverse=True)
# create int_to_vocab dictionaries
int_to_vocab = {ii: word for ii, word in enumerate(sorted_vocab)}
vocab_to_int = {word: ii for ii, word in int_to_vocab.items()}
return (vocab_to_int, int_to_vocab)
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_create_lookup_tables(create_lookup_tables)
```
### Tokenize Punctuation
We'll be splitting the script into a word array using spaces as delimiters. However, punctuations like periods and exclamation marks can create multiple ids for the same word. For example, "bye" and "bye!" would generate two different word ids.
Implement the function `token_lookup` to return a dict that will be used to tokenize symbols like "!" into "||Exclamation_Mark||". Create a dictionary for the following symbols where the symbol is the key and value is the token:
- Period ( **.** )
- Comma ( **,** )
- Quotation Mark ( **"** )
- Semicolon ( **;** )
- Exclamation mark ( **!** )
- Question mark ( **?** )
- Left Parentheses ( **(** )
- Right Parentheses ( **)** )
- Dash ( **-** )
- Return ( **\n** )
This dictionary will be used to tokenize the symbols and add the delimiter (space) around it. This separates each symbols as its own word, making it easier for the neural network to predict the next word. Make sure you don't use a value that could be confused as a word; for example, instead of using the value "dash", try using something like "||dash||".
```
def token_lookup():
"""
Generate a dict to turn punctuation into a token.
:return: Tokenized dictionary where the key is the punctuation and the value is the token
"""
# TODO: Implement Function
punctuation_dict = {
"." : "||Period||",
",":"||Comma||",
'"':"||QuotationMark||",
";": "||Semicolon||",
"!":"||Exclamationmark||",
"?":"||Questionmark||",
"(":"||LeftParentheses||",
")":"||RightParentheses||",
"-":"||Dash||",
"\n":"||Return||"
}
return punctuation_dict
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_tokenize(token_lookup)
```
## Pre-process all the data and save it
Running the code cell below will pre-process all the data and save it to file. You're encouraged to lok at the code for `preprocess_and_save_data` in the `helpers.py` file to see what it's doing in detail, but you do not need to change this code.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
# pre-process training data
helper.preprocess_and_save_data(data_dir, token_lookup, create_lookup_tables)
```
# Check Point
This is your first checkpoint. If you ever decide to come back to this notebook or have to restart the notebook, you can start from here. The preprocessed data has been saved to disk.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import helper
import problem_unittests as tests
int_text, vocab_to_int, int_to_vocab, token_dict = helper.load_preprocess()
```
## Build the Neural Network
In this section, you'll build the components necessary to build an RNN by implementing the RNN Module and forward and backpropagation functions.
### Check Access to GPU
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import torch
# Check for a GPU
train_on_gpu = torch.cuda.is_available()
if not train_on_gpu:
print('No GPU found. Please use a GPU to train your neural network.')
```
## Input
Let's start with the preprocessed input data. We'll use [TensorDataset](http://pytorch.org/docs/master/data.html#torch.utils.data.TensorDataset) to provide a known format to our dataset; in combination with [DataLoader](http://pytorch.org/docs/master/data.html#torch.utils.data.DataLoader), it will handle batching, shuffling, and other dataset iteration functions.
You can create data with TensorDataset by passing in feature and target tensors. Then create a DataLoader as usual.
```
data = TensorDataset(feature_tensors, target_tensors)
data_loader = torch.utils.data.DataLoader(data,
batch_size=batch_size)
```
### Batching
Implement the `batch_data` function to batch `words` data into chunks of size `batch_size` using the `TensorDataset` and `DataLoader` classes.
>You can batch words using the DataLoader, but it will be up to you to create `feature_tensors` and `target_tensors` of the correct size and content for a given `sequence_length`.
For example, say we have these as input:
```
words = [1, 2, 3, 4, 5, 6, 7]
sequence_length = 4
```
Your first `feature_tensor` should contain the values:
```
[1, 2, 3, 4]
```
And the corresponding `target_tensor` should just be the next "word"/tokenized word value:
```
5
```
This should continue with the second `feature_tensor`, `target_tensor` being:
```
[2, 3, 4, 5] # features
6 # target
```
```
from torch.utils.data import TensorDataset, DataLoader
def batch_data(words, sequence_length, batch_size):
"""
Batch the neural network data using DataLoader
:param words: The word ids of the TV scripts
:param sequence_length: The sequence length of each batch
:param batch_size: The size of each batch; the number of sequences in a batch
:return: DataLoader with batched data
"""
# TODO: Implement function
# partial_feature = []
# features = []
# targets = []
# count = 0
# for i in range(0,len(words)):
# if count == sequence_length: # adding plus one as the last value is to included into feature
# features.append(partial_feature)
# partial_feature = []
# count=0
# targets.append(words[i]) # here i value is already next value so saving this value as target
# partial_feature.append(words[i])
# count+=1
# train_dataset = TensorDataset(torch.from_numpy(np.array(features)),torch.from_numpy(np.array(targets)))
# train_loader = DataLoader(train_dataset, shuffle=True, batch_size=batch_size)
n_batches = len(words)//batch_size
# only full batches
words = words[:n_batches*batch_size]
y_len = len(words) - sequence_length
x, y = [], []
for idx in range(0, y_len):
idx_end = sequence_length + idx
x_batch = words[idx:idx_end]
x.append(x_batch)
# print("feature: ",x_batch)
batch_y = words[idx_end]
# print("target: ", batch_y)
y.append(batch_y)
# create Tensor datasets
data = TensorDataset(torch.from_numpy(np.asarray(x)), torch.from_numpy(np.asarray(y)))
# make sure the SHUFFLE your training data
data_loader = DataLoader(data, shuffle=False, batch_size=batch_size)
# return a dataloader
return data_loader
# there is no test for this function, but you are encouraged to create
# print statements and tests of your own
```
### Test your dataloader
You'll have to modify this code to test a batching function, but it should look fairly similar.
Below, we're generating some test text data and defining a dataloader using the function you defined, above. Then, we are getting some sample batch of inputs `sample_x` and targets `sample_y` from our dataloader.
Your code should return something like the following (likely in a different order, if you shuffled your data):
```
torch.Size([10, 5])
tensor([[ 28, 29, 30, 31, 32],
[ 21, 22, 23, 24, 25],
[ 17, 18, 19, 20, 21],
[ 34, 35, 36, 37, 38],
[ 11, 12, 13, 14, 15],
[ 23, 24, 25, 26, 27],
[ 6, 7, 8, 9, 10],
[ 38, 39, 40, 41, 42],
[ 25, 26, 27, 28, 29],
[ 7, 8, 9, 10, 11]])
torch.Size([10])
tensor([ 33, 26, 22, 39, 16, 28, 11, 43, 30, 12])
```
### Sizes
Your sample_x should be of size `(batch_size, sequence_length)` or (10, 5) in this case and sample_y should just have one dimension: batch_size (10).
### Values
You should also notice that the targets, sample_y, are the *next* value in the ordered test_text data. So, for an input sequence `[ 28, 29, 30, 31, 32]` that ends with the value `32`, the corresponding output should be `33`.
```
# test dataloader
test_text = range(50)
t_loader = batch_data(test_text, sequence_length=5, batch_size=10)
data_iter = iter(t_loader)
sample_x, sample_y = data_iter.next()
print(sample_x.shape)
print(sample_x)
print()
print(sample_y.shape)
print(sample_y)
```
---
## Build the Neural Network
Implement an RNN using PyTorch's [Module class](http://pytorch.org/docs/master/nn.html#torch.nn.Module). You may choose to use a GRU or an LSTM. To complete the RNN, you'll have to implement the following functions for the class:
- `__init__` - The initialize function.
- `init_hidden` - The initialization function for an LSTM/GRU hidden state
- `forward` - Forward propagation function.
The initialize function should create the layers of the neural network and save them to the class. The forward propagation function will use these layers to run forward propagation and generate an output and a hidden state.
**The output of this model should be the *last* batch of word scores** after a complete sequence has been processed. That is, for each input sequence of words, we only want to output the word scores for a single, most likely, next word.
### Hints
1. Make sure to stack the outputs of the lstm to pass to your fully-connected layer, you can do this with `lstm_output = lstm_output.contiguous().view(-1, self.hidden_dim)`
2. You can get the last batch of word scores by shaping the output of the final, fully-connected layer like so:
```
# reshape into (batch_size, seq_length, output_size)
output = output.view(batch_size, -1, self.output_size)
# get last batch
out = output[:, -1]
```
```
import torch.nn as nn
class RNN(nn.Module):
def __init__(self, vocab_size, output_size, embedding_dim, hidden_dim, n_layers, dropout=0.5):
"""
Initialize the PyTorch RNN Module
:param vocab_size: The number of input dimensions of the neural network (the size of the vocabulary)
:param output_size: The number of output dimensions of the neural network
:param embedding_dim: The size of embeddings, should you choose to use them
:param hidden_dim: The size of the hidden layer outputs
:param dropout: dropout to add in between LSTM/GRU layers
"""
super(RNN, self).__init__()
# TODO: Implement function
# set class variables
self.output_size = output_size
self.vocab_size = vocab_size
self.embedding_dim = embedding_dim
self.hidden_dim = hidden_dim
self.n_layers = n_layers
# define model layers
self.embed = nn.Embedding(num_embeddings=self.vocab_size, embedding_dim=self.embedding_dim)
self.lstm = nn.LSTM(input_size=self.embedding_dim,hidden_size=self.hidden_dim, num_layers=self.n_layers,batch_first=True,dropout=dropout)
self.dropout = nn.Dropout(0.3)
self.fc1 = nn.Linear(self.hidden_dim,self.output_size)
def forward(self, nn_input, hidden):
"""
Forward propagation of the neural network
:param nn_input: The input to the neural network
:param hidden: The hidden state
:return: Two Tensors, the output of the neural network and the latest hidden state
"""
# TODO: Implement function
batch_size = nn_input.size(0)
embed = self.embed(nn_input)
nn_input, hidden = self.lstm(embed,hidden)
output = nn_input.contiguous().view(-1, self.hidden_dim)
output = self.fc1(output)
output = output.view(batch_size,-1,self.output_size)
output = output[:,-1]
# return one batch of output word scores and the hidden state
return output, hidden
def init_hidden(self, batch_size):
'''
Initialize the hidden state of an LSTM/GRU
:param batch_size: The batch_size of the hidden state
:return: hidden state of dims (n_layers, batch_size, hidden_dim)
'''
# Implement function
# initialize hidden state with zero weights, and move to GPU if available
weight = next(self.parameters()).data
if (train_on_gpu):
hidden = (weight.new(self.n_layers, batch_size, self.hidden_dim).zero_().cuda(),
weight.new(self.n_layers, batch_size, self.hidden_dim).zero_().cuda())
else:
hidden = (weight.new(self.n_layers, batch_size, self.hidden_dim).zero_(),
weight.new(self.n_layers, batch_size, self.hidden_dim).zero_())
return hidden
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_rnn(RNN, train_on_gpu)
```
### Define forward and backpropagation
Use the RNN class you implemented to apply forward and back propagation. This function will be called, iteratively, in the training loop as follows:
```
loss = forward_back_prop(decoder, decoder_optimizer, criterion, inp, target)
```
And it should return the average loss over a batch and the hidden state returned by a call to `RNN(inp, hidden)`. Recall that you can get this loss by computing it, as usual, and calling `loss.item()`.
**If a GPU is available, you should move your data to that GPU device, here.**
```
def forward_back_prop(rnn, optimizer, criterion, inp, target, hidden):
"""
Forward and backward propagation on the neural network
:param decoder: The PyTorch Module that holds the neural network
:param decoder_optimizer: The PyTorch optimizer for the neural network
:param criterion: The PyTorch loss function
:param inp: A batch of input to the neural network
:param target: The target output for the batch of input
:return: The loss and the latest hidden state Tensor
"""
# TODO: Implement Function
# move model to GPU, if available
if(train_on_gpu):
rnn.cuda()
# # Creating new variables for the hidden state, otherwise
# # we'd backprop through the entire training history
h = tuple([each.data for each in hidden])
# zero accumulated gradients
rnn.zero_grad()
if(train_on_gpu):
inp, target = inp.cuda(), target.cuda()
# get predicted outputs
output, h = rnn(inp, h)
# calculate loss
loss = criterion(output, target)
# optimizer.zero_grad()
loss.backward()
# 'clip_grad_norm' helps prevent the exploding gradient problem in RNNs / LSTMs
nn.utils.clip_grad_norm_(rnn.parameters(), 5)
optimizer.step()
return loss.item(), h
# Note that these tests aren't completely extensive.
# they are here to act as general checks on the expected outputs of your functions
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_forward_back_prop(RNN, forward_back_prop, train_on_gpu)
```
## Neural Network Training
With the structure of the network complete and data ready to be fed in the neural network, it's time to train it.
### Train Loop
The training loop is implemented for you in the `train_decoder` function. This function will train the network over all the batches for the number of epochs given. The model progress will be shown every number of batches. This number is set with the `show_every_n_batches` parameter. You'll set this parameter along with other parameters in the next section.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
def train_rnn(rnn, batch_size, optimizer, criterion, n_epochs, show_every_n_batches=100):
batch_losses = []
rnn.train()
print("Training for %d epoch(s)..." % n_epochs)
for epoch_i in range(1, n_epochs + 1):
# initialize hidden state
hidden = rnn.init_hidden(batch_size)
for batch_i, (inputs, labels) in enumerate(train_loader, 1):
# make sure you iterate over completely full batches, only
n_batches = len(train_loader.dataset)//batch_size
if(batch_i > n_batches):
break
# forward, back prop
loss, hidden = forward_back_prop(rnn, optimizer, criterion, inputs, labels, hidden)
# record loss
batch_losses.append(loss)
# printing loss stats
if batch_i % show_every_n_batches == 0:
print('Epoch: {:>4}/{:<4} Loss: {}\n'.format(
epoch_i, n_epochs, np.average(batch_losses)))
batch_losses = []
# returns a trained rnn
return rnn
```
### Hyperparameters
Set and train the neural network with the following parameters:
- Set `sequence_length` to the length of a sequence.
- Set `batch_size` to the batch size.
- Set `num_epochs` to the number of epochs to train for.
- Set `learning_rate` to the learning rate for an Adam optimizer.
- Set `vocab_size` to the number of uniqe tokens in our vocabulary.
- Set `output_size` to the desired size of the output.
- Set `embedding_dim` to the embedding dimension; smaller than the vocab_size.
- Set `hidden_dim` to the hidden dimension of your RNN.
- Set `n_layers` to the number of layers/cells in your RNN.
- Set `show_every_n_batches` to the number of batches at which the neural network should print progress.
If the network isn't getting the desired results, tweak these parameters and/or the layers in the `RNN` class.
```
# Data params
# Sequence Length
sequence_length = 10 # of words in a sequence
# Batch Size
batch_size = 128
# data loader - do not change
train_loader = batch_data(int_text, sequence_length, batch_size)
# Training parameters
# Number of Epochs
num_epochs = 10
# Learning Rate
learning_rate = 0.001
# Model parameters
# Vocab size
vocab_size = len(vocab_to_int)
# Output size
output_size = vocab_size
# Embedding Dimension
embedding_dim = 200
# Hidden Dimension
hidden_dim = 250
# Number of RNN Layers
n_layers = 2
# Show stats for every n number of batches
show_every_n_batches = 500
```
### Train
In the next cell, you'll train the neural network on the pre-processed data. If you have a hard time getting a good loss, you may consider changing your hyperparameters. In general, you may get better results with larger hidden and n_layer dimensions, but larger models take a longer time to train.
> **You should aim for a loss less than 3.5.**
You should also experiment with different sequence lengths, which determine the size of the long range dependencies that a model can learn.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
# create model and move to gpu if available
rnn = RNN(vocab_size, output_size, embedding_dim, hidden_dim, n_layers, dropout=0.5)
if train_on_gpu:
rnn.cuda()
# defining loss and optimization functions for training
optimizer = torch.optim.Adam(rnn.parameters(), lr=learning_rate)
criterion = nn.CrossEntropyLoss()
# training the model
trained_rnn = train_rnn(rnn, batch_size, optimizer, criterion, num_epochs, show_every_n_batches)
# saving the trained model
helper.save_model('./save/trained_rnn', trained_rnn)
print('Model Trained and Saved')
```
### Question: How did you decide on your model hyperparameters?
For example, did you try different sequence_lengths and find that one size made the model converge faster? What about your hidden_dim and n_layers; how did you decide on those?
**Answer:** (Write answer, here)
---
# Checkpoint
After running the above training cell, your model will be saved by name, `trained_rnn`, and if you save your notebook progress, **you can pause here and come back to this code at another time**. You can resume your progress by running the next cell, which will load in our word:id dictionaries _and_ load in your saved model by name!
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import torch
import helper
import problem_unittests as tests
_, vocab_to_int, int_to_vocab, token_dict = helper.load_preprocess()
trained_rnn = helper.load_model('./save/trained_rnn')
```
## Generate TV Script
With the network trained and saved, you'll use it to generate a new, "fake" Seinfeld TV script in this section.
### Generate Text
To generate the text, the network needs to start with a single word and repeat its predictions until it reaches a set length. You'll be using the `generate` function to do this. It takes a word id to start with, `prime_id`, and generates a set length of text, `predict_len`. Also note that it uses topk sampling to introduce some randomness in choosing the most likely next word, given an output set of word scores!
```
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
import torch.nn.functional as F
def generate(rnn, prime_id, int_to_vocab, token_dict, pad_value, predict_len=100):
"""
Generate text using the neural network
:param decoder: The PyTorch Module that holds the trained neural network
:param prime_id: The word id to start the first prediction
:param int_to_vocab: Dict of word id keys to word values
:param token_dict: Dict of puncuation tokens keys to puncuation values
:param pad_value: The value used to pad a sequence
:param predict_len: The length of text to generate
:return: The generated text
"""
rnn.eval()
# create a sequence (batch_size=1) with the prime_id
current_seq = np.full((1, sequence_length), pad_value)
current_seq[-1][-1] = prime_id
predicted = [int_to_vocab[prime_id]]
for _ in range(predict_len):
if train_on_gpu:
current_seq = torch.LongTensor(current_seq).cuda()
else:
current_seq = torch.LongTensor(current_seq)
# initialize the hidden state
hidden = rnn.init_hidden(current_seq.size(0))
# get the output of the rnn
output, _ = rnn(current_seq, hidden)
# get the next word probabilities
p = F.softmax(output, dim=1).data
if(train_on_gpu):
p = p.cpu() # move to cpu
# use top_k sampling to get the index of the next word
top_k = 5
p, top_i = p.topk(top_k)
top_i = top_i.numpy().squeeze()
# select the likely next word index with some element of randomness
p = p.numpy().squeeze()
word_i = np.random.choice(top_i, p=p/p.sum())
# retrieve that word from the dictionary
word = int_to_vocab[word_i]
predicted.append(word)
# the generated word becomes the next "current sequence" and the cycle can continue
current_seq = np.roll(current_seq, -1, 1)
current_seq[-1][-1] = word_i
gen_sentences = ' '.join(predicted)
# Replace punctuation tokens
for key, token in token_dict.items():
ending = ' ' if key in ['\n', '(', '"'] else ''
gen_sentences = gen_sentences.replace(' ' + token.lower(), key)
gen_sentences = gen_sentences.replace('\n ', '\n')
gen_sentences = gen_sentences.replace('( ', '(')
# return all the sentences
return gen_sentences
```
### Generate a New Script
It's time to generate the text. Set `gen_length` to the length of TV script you want to generate and set `prime_word` to one of the following to start the prediction:
- "jerry"
- "elaine"
- "george"
- "kramer"
You can set the prime word to _any word_ in our dictionary, but it's best to start with a name for generating a TV script. (You can also start with any other names you find in the original text file!)
```
# run the cell multiple times to get different results!
gen_length = 400 # modify the length to your preference
prime_word = 'jerry' # name for starting the script
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
pad_word = helper.SPECIAL_WORDS['PADDING']
generated_script = generate(trained_rnn, vocab_to_int[prime_word + ':'], int_to_vocab, token_dict, vocab_to_int[pad_word], gen_length)
print(generated_script)
```
#### Save your favorite scripts
Once you have a script that you like (or find interesting), save it to a text file!
```
# save script to a text file
f = open("generated_script_1.txt","w")
f.write(generated_script)
f.close()
```
# The TV Script is Not Perfect
It's ok if the TV script doesn't make perfect sense. It should look like alternating lines of dialogue, here is one such example of a few generated lines.
### Example generated script
>jerry: what about me?
>
>jerry: i don't have to wait.
>
>kramer:(to the sales table)
>
>elaine:(to jerry) hey, look at this, i'm a good doctor.
>
>newman:(to elaine) you think i have no idea of this...
>
>elaine: oh, you better take the phone, and he was a little nervous.
>
>kramer:(to the phone) hey, hey, jerry, i don't want to be a little bit.(to kramer and jerry) you can't.
>
>jerry: oh, yeah. i don't even know, i know.
>
>jerry:(to the phone) oh, i know.
>
>kramer:(laughing) you know...(to jerry) you don't know.
You can see that there are multiple characters that say (somewhat) complete sentences, but it doesn't have to be perfect! It takes quite a while to get good results, and often, you'll have to use a smaller vocabulary (and discard uncommon words), or get more data. The Seinfeld dataset is about 3.4 MB, which is big enough for our purposes; for script generation you'll want more than 1 MB of text, generally.
# Submitting This Project
When submitting this project, make sure to run all the cells before saving the notebook. Save the notebook file as "dlnd_tv_script_generation.ipynb" and save another copy as an HTML file by clicking "File" -> "Download as.."->"html". Include the "helper.py" and "problem_unittests.py" files in your submission. Once you download these files, compress them into one zip file for submission.
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
import sys
sys.path.insert(0, '../')
from __future__ import division
import os
import numpy as np
import pandas as pd
from skimage.io import imread
from skimage.color import gray2rgb
import cv2
from pandas import read_csv
import matplotlib.pyplot as plt
import glob
%matplotlib inline
```
## Theano loss function - SmoothL1 and IoU
```
import theano
import theano.tensor as T
npr = np.random.random((10,4)).astype(np.float32)
nta = np.random.random((10,4)).astype(np.float32)
p = T.fmatrix('p')
t = T.fmatrix('t')
```
### SmoothL1 loss
```
def ff(p, t):
c = T.abs_(p - t)
e = T.switch(T.lt(c, 1.), 0.5 * c * c, c - 0.5)
return T.mean(T.sum(e, axis=1))
g = ff(p, t)
f = theano.function(
inputs=[p, t],
outputs=g
)
nc = np.abs(npr - nta)
td = f(npr, nta)
print td
T.grad(g, p)
```
### IoU loss - Intersection over Union loss
```
pn = np.array([[0.1, 0.1, 0.5, 0.8],
[0.1, 0.1, 0.2, 0.2]], dtype=np.float32)
tn = np.array([[0.3, 0.5, 0.4, 0.9],
[0.3, 0.2, 0.7, 0.8]], dtype=np.float32)
def hh(p, t):
hund = np.float32(100)
pt = (p * hund).astype(np.int32)
tt = (t * hund).astype(np.int32)
imp = np.zeros((pt.shape[0], 100, 100), dtype=np.uint8)
imt = np.zeros((tt.shape[0], 100, 100), dtype=np.uint8)
for i in range(pt.shape[0]):
imp[i, pt[i, 1]:pt[i, 3], pt[i, 0]:pt[i, 2]] = 1
imt[i, tt[i, 1]:tt[i, 3], tt[i, 0]:tt[i, 2]] = 1
intersection = np.logical_and(imp, imt).astype(np.float32).sum(axis=2).sum(axis=1)
union = np.logical_or(imp, imt).astype(np.float32).sum(axis=2).sum(axis=1)
loss = 1. - intersection / union
plt.imshow(imp[0, ...])
return loss
hh(pn, tn)
def gg(p, t):
tp, tt = p.reshape((p.shape[0], 2, 2)), t.reshape((t.shape[0], 2, 2))
overlaps = np.zeros_like(tp, dtype=np.float32)
overlaps[:, 0, :] = np.maximum(tp[:, 0, :], tt[:, 0, :])
overlaps[:, 1, :] = np.minimum(tp[:, 1, :], tt[:, 1, :])
intersection = overlaps[:, 1, :] - overlaps[:, 0, :]
bool_overlap = np.min(intersection, axis=1) > 0
intersection = intersection[:, 0] * intersection[:, 1]
intersection[bool_overlap==False] == 0.
dims_p = tp[:, 1, :] - tp[:, 0, :]
areas_p = dims_p[:, 0] * dims_p[:, 1]
dims_t = tt[:, 1, :] - tt[:, 0, :]
areas_t = dims_t[:, 0] * dims_t[:, 1]
union = areas_p + areas_t - intersection
ratio = 1. - np.minimum(np.exp(np.log(np.abs(intersection)) - np.log(np.abs(union) + 1e-5)), 1.)
# no_overlap_penalty = 1. + np.abs(tp - tt).sum(axis=2).max(axis=1)
loss = ratio.copy()
# loss[bool_overlap==False] = no_overlap_penalty[bool_overlap==False]
return loss
print tp
print "-"
print tt
print "-"
print no_overlap_penalty
print no_overlap_penalty
print "overlap"
print overlaps
print "int/union"
print intersection, union
print "bool overlap"
print bool_overlap
print "loss"
print loss
gg(pn, tn)
tp, tt = p.reshape((p.shape[0], 2, 2)), t.reshape((t.shape[0], 2, 2))
overlaps_t0 = T.maximum(tp[:, 0, :], tt[:, 0, :])
overlaps_t1 = T.minimum(tp[:, 1, :], tt[:, 1, :])
intersection = overlaps_t1 - overlaps_t0
bool_overlap = T.min(intersection, axis=1) > 0
intersection = intersection[:, 0] * intersection[:, 1]
intersection[bool_overlap==False] == 0.
dims_p = tp[:, 1, :] - tp[:, 0, :]
areas_p = dims_p[:, 0] * dims_p[:, 1]
dims_t = tt[:, 1, :] - tt[:, 0, :]
areas_t = dims_t[:, 0] * dims_t[:, 1]
union = areas_p + areas_t - intersection
loss = 1. - T.minimum(T.exp(T.log(T.abs_(intersection)) - T.log(T.abs_(union) + np.float32(1e-5))), np.float32(1.))
print "done"
theano_iou = theano.function(
inputs=[p, t],
outputs=loss
)
theano_iou(pn, tn)
def theano_hh(p, t):
hund = np.float32(100)
pt = (p * hund).astype(np.int32)
tt = (t * hund).astype(np.int32)
imp = np.zeros((pt.shape[0], 100, 100), dtype=np.uint8)
imt = np.zeros((tt.shape[0], 100, 100), dtype=np.uint8)
for i in range(pt.shape[0]):
imp[i, pt[i, 1]:pt[i, 3], pt[i, 0]:pt[i, 2]] = 1
imt[i, tt[i, 1]:tt[i, 3], tt[i, 0]:tt[i, 2]] = 1
intersection = np.logical_and(imp, imt).astype(np.float32).sum(axis=2).sum(axis=1)
union = np.logical_or(imp, imt).astype(np.float32).sum(axis=2).sum(axis=1)
loss = 1. - intersection / union
return loss
hh(pn, tn)
z = tn.reshape((2, 2, 2))
z[:, 0, :], z
```
## Augmentation
```
from utils import get_file_list
import glob
from train_val_split import read_facescrub_img_list
from plotting import plot_face_bb
```
### Train data split
```
folder = '/media/shared/faceScrub/train_face_det/'
path = folder
actor_label_txt = '/media/shared/faceScrub/facescrub_actors.txt'
actress_label_txt = '/media/shared/faceScrub/facescrub_actresses.txt'
accept_pattern = '*/*.jpg'
fnames, bboxes = read_facescrub_img_list(path, actor_label_txt, actress_label_txt, accept_pattern='*/*.jpg')
len(fnames), len(bboxes)
```
### Data augment
```
train_csv = 'train.csv'
train_df = read_csv(train_csv, sep='\t')
X = np.asarray(train_df['name'].as_matrix())
y_str = train_df['bbox']
y_l = map(lambda k: [np.float32(v)
for v in k.split(',')], y_str)
y = np.asarray(y_l)
x_1 = X[1]
y_1 = y[1, :]
im = imread(x_1)
plot_face_bb(im, y_1, scale=False, path=False)
img = cv2.cvtColor(im, cv2.COLOR_RGB2BGR)
img_size = (256, 256, 3)
MAX_FACE_SIZE = 220
MIN_FACE_SIZE = 64
h0, w0, _ = img_size
w1, h1, w2, h2 = y_1
wc, hc = (y_1[0] + y_1[2]) / 2, (y_1[1] + y_1[3]) / 2
face_width = (h2 - h1) / 2
print "Original center coords: (%.1f, %.1f)" % (wc, hc)
print "Face coords: (%.1f, %.1f) (%.1f, %.1f) and face width: %.0f" % (h1, w1, h2, w2, face_width * 2)
rng = np.random.RandomState(seed=1234)
# Possible scales of computation
high_scale = MAX_FACE_SIZE/2/face_width
low_scale = MIN_FACE_SIZE/2/face_width
print "Scales of computation: %.3f - %.3f" % (low_scale, high_scale)
scale_comp = rng.choice(np.arange(low_scale, high_scale, (high_scale-low_scale)/100), 1)[0]
new_face_width = round(face_width * scale_comp)
swc, shc = round(wc * scale_comp), round(hc * scale_comp)
sh1, sw1, sh2, sw2 = (shc-new_face_width, swc-new_face_width,
shc+new_face_width, swc+new_face_width)
print "Chosen scale of computation: %.3f," % (scale_comp)
print "New face center: (%.1f, %.1f)\nface width: %.0f" % (
shc, swc, new_face_width * 2)
res = cv2.resize(img, None, fx=scale_comp, fy=scale_comp)
h, w, _ = res.shape
print "Face width: %.1f" % (new_face_width * 2)
plot_face_bb(res[:,:,::-1], y_1*scale_comp, scale=False, path=False)
# Possible location of the face
min_pad = new_face_width + 30
lw, lh, hw, hh = (min(min_pad, w0 - min_pad), min(min_pad, h0 - min_pad),
max(min_pad, w0 - min_pad), max(min_pad, h0 - min_pad))
print "Get face center in the given window: (%.1f, %.1f) (%.1f, %.1f)" % (lh, lw, hh, hw)
twc = rng.randint(lw, hw, 1)[0]
thc = rng.randint(lh, hh, 1)[0]
print "New center location: (%.1f, %.1f)" % (thc, twc)
out = np.random.randint(0, high=255, size=img_size).astype(np.uint8)
sfh = shc - thc
tfh = int(0 if sfh > 0 else abs(sfh))
sfh = int(0 if sfh < 0 else sfh)
sfw = swc - twc
tfw = int(0 if sfw > 0 else abs(sfw))
sfw = int(0 if sfw < 0 else sfw)
print "source begin: (%.0f, %.0f) target begin: (%.0f, %.0f)" % (sfh, sfw, tfh, tfw)
seh = shc - thc + h0
teh = int(h0 if seh <= h else h0 - seh + h)
seh = int(h if seh > h else seh)
sew = swc - twc + w0
tew = int(w0 if sew <= w else w0 - sew + w)
sew = int(w if sew > w else sew)
print "source end: (%.0f, %.0f) target end: (%.0f, %.0f)" % (seh, sew, teh, tew)
ty_1 = np.array([twc-new_face_width, thc-new_face_width, twc+new_face_width, thc+new_face_width])
out[tfh:teh, tfw:tew, :] = res[sfh:seh, sfw:sew, :]
out = cv2.cvtColor(out, cv2.COLOR_BGR2RGB)
plot_face_bb(out, ty_1, scale=False, path=False)
print "new face center: (%.0f, %.0f)" % (thc, twc)
for i in glob.glob('*.jpg'):
plt.figure()
plt.imshow(imread(i))
```
| github_jupyter |
```
!pip install torch torchtext
!git clone https://github.com/neubig/nn4nlp-code.git
from __future__ import print_function
import time
from collections import defaultdict
import random
import math
import sys
import argparse
import numpy as np
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.utils.data import Dataset, DataLoader
# format of files: each line is "word1|tag1 word2|tag2 ..."
train_file = "nn4nlp-code/data/tags/train.txt"
dev_file = "nn4nlp-code/data/tags/dev.txt"
w2i = defaultdict(lambda: len(w2i))
t2i = defaultdict(lambda: len(t2i))
def read(fname):
"""
Read tagged file
"""
with open(fname, "r") as f:
for line in f:
words, tags = [], []
for wt in line.strip().split():
w, t = wt.split('|')
words.append(w2i[w])
tags.append(t2i[t])
yield (words, tags)
# Read the data
train = list(read(train_file))
unk_word = w2i["<unk>"]
w2i = defaultdict(lambda: unk_word, w2i)
unk_tag = t2i["<unk>"]
t2i = defaultdict(lambda: unk_tag, t2i)
nwords = len(w2i)
ntags = len(t2i)
dev = list(read(dev_file))
t2i
!head nn4nlp-code/data/tags/train.txt
class BiLSTM(nn.Module):
def __init__(self, vocab_size, tag_size, embed_dim, hidden_size, num_layers=1):
super(BiLSTM, self).__init__()
self.vocab_size = vocab_size
self.tag_size = tag_size
self.embed_dim = embed_dim
self.hidden_size = hidden_size
self.embeddings = nn.Embedding(vocab_size, embed_dim)
self.lstm = nn.LSTM(embed_dim, hidden_size, num_layers, bidirectional=True)
self.fc = nn.Linear(hidden_size*2, tag_size)
def init_hidden(self, bs):
return (torch.zeros(2, bs, self.hidden_size, device=device),
torch.zeros(2, bs, self.hidden_size, device=device))
def forward(self, x):
seq_len, bs = x.shape
h = self.init_hidden(bs)
x = self.embeddings(x)
x, h = self.lstm(x, h)
x = self.fc(x.view(seq_len*bs, -1)).view(seq_len, bs, -1)
return x
class TaggerDataset(Dataset):
def __init__(self, data):
self.data = data
def __len__(self):
return len(self.data)
def __getitem__(self, ix):
return torch.LongTensor(self.data[ix][0]), torch.LongTensor(self.data[ix][1])
def pad_tensor(vec, pad, dim):
"""
args:
vec - tensor to pad
pad - the size to pad to
dim - dimension to pad
return:
a new tensor padded to 'pad' in dimension 'dim'
"""
pad_size = list(vec.shape)
pad_size[dim] = pad - vec.size(dim)
return torch.cat([vec, torch.zeros(*pad_size).long()], dim=dim)
class PadCollate:
"""
a variant of callate_fn that pads according to the longest sequence in
a batch of sequences
"""
def __init__(self, dim=0):
"""
args:
dim - the dimension to be padded (dimension of time in sequences)
"""
self.dim = dim
def pad_collate(self, batch):
"""
args:
batch - list of (tensor, label)
reutrn:
xs - a tensor of all examples in 'batch' after padding
ys - a LongTensor of all labels in batch
"""
# find longest sequence
xlens = list(map(lambda x: x[0].shape[self.dim], batch))
ylens = list(map(lambda x: x[1].shape[self.dim], batch))
max_xlen = max(xlens)
max_ylen = max(ylens)
# pad according to max_len
batch = list(map(lambda x:
(pad_tensor(x[0], pad=max_xlen, dim=self.dim),
pad_tensor(x[1], pad=max_ylen, dim=self.dim)), batch))
# stack all
xs = torch.stack(list(map(lambda x: x[0], batch)), dim=0)
ys = torch.stack(list(map(lambda x: x[1], batch)), dim=0)
return xs, ys, torch.LongTensor(xlens), torch.LongTensor(ylens)
def __call__(self, batch):
return self.pad_collate(batch)
embed_dim = 128
hidden_dim = 64
BATCH_SIZE = 16
model = BiLSTM(nwords, ntags, embed_dim, hidden_dim)
criterion = nn.CrossEntropyLoss(ignore_index=9)
trainer = torch.optim.Adam(model.parameters(), lr=1e-3)
train_dataset = TaggerDataset(train)
train_dataloader = DataLoader(train_dataset, batch_size=BATCH_SIZE, shuffle=True, collate_fn=PadCollate(dim=0))
dev_dataset = TaggerDataset(dev)
dev_dataloader = DataLoader(train_dataset, batch_size=BATCH_SIZE, collate_fn=PadCollate(dim=0))
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
for epoch_i in range(10):
model.train()
total_loss = 0.
for batch_i, batch in enumerate(train_dataloader):
xs, ys, xlens, ylens = batch
logits = model(xs.transpose(0, 1))
model.zero_grad()
loss = criterion(logits.transpose(0,1).contiguous().view(-1, ntags), ys.view(-1))
total_loss += loss.item()
loss.backward()
trainer.step()
if batch_i % 200 == 0:
print(f"epoch {epoch_i} | batch {batch_i} | avg_loss: {total_loss/(batch_i+1):.4f}")
model.eval()
val_loss = 0.
val_correct = 0
val_cnt = 0
for batch_i, batch in enumerate(dev_dataloader):
xs, ys, xlens, ylens = batch
logits = model(xs.transpose(0, 1))
loss = criterion(logits.transpose(0,1).contiguous().view(-1, ntags), ys.view(-1))
val_loss += loss.item()
mask = torch.arange(ys.shape[1]).unsqueeze(0).expand_as(ys) - ylens.float().unsqueeze(1) < 0
correct = ((torch.argmax(logits, -1).transpose(0, 1) == ys) & mask).sum()
val_cnt += ylens.sum().item()
val_correct += correct.item()
print(f"val_acc: {1. * val_correct / val_cnt} | val_loss: {val_loss/(batch_i+1)}")
```
| github_jupyter |
En este ejercicio vamos a estudiar los ciclos que se producen en $\mathbb{Z}_m$, con $m$ un entero casi siempre primo, al elevar sucesivamente al cuadrado sus elementos.
```
def f(n,m):
return n^2%m
def diccionario(m):
dicc = {}
for int in srange(m):
int2 = f(int,m)
dicc[int]=[int2]
return dicc
G1 = DiGraph(diccionario(19))
G1.graphplot().show(figsize=[12,12])
def orbita_c(f,n,m):
L = []
while not n in L:
L.append(n)
n = f(n,m)
k = L.index(n)
return L[k:]
def ciclos(f,m):
L = []
for n in srange(m):
L.append(orbita_c(f,n,m))
return L
ciclos(f,19)
```
Vemos que los ciclos que aparecen son los mismos que el el grafo, y el único problema es que *ciclos* debería mostrar cada ciclo una única vez. Si no lo hace así tendríamos que comprobar a *ojo* qué ciclos son el mismo.
```
def ciclosn(f,m):
L = []
A = set([])
for n in srange(m):
orb = orbita_c(f,n,m)
if f(orb[-1],m)==orb[0] and orb[-1] not in A:
L.append(orb)
A = A | set(orb)
return L
ciclosn(f,19)
ciclosn(f,11)
ciclosn(f,13)
ciclosn(f,17)
```
<p>¿Qué observamos con estos pocos ejemplos? </p>
<ol>
<li>Parece que para $m$ primo los únicos puntos fijos son $0$ y $1$.</li>
<li>El comportamiento para $p=17$ es bastante diferente ya que los únicos puntos fijos que aparecen son $0$ y $1$.</li>
</ol>
<p>Usando estos pocos ejemplos, parece claro que debemos fijarnos el número de puntos fijos, en el número de ciclos y en sus longitudes.</p>
<h4>Puntos fijos:</h4>
```
def puntos_fijos(f,m):
L = ciclosn(f,m)
cont = 0
for item in L:
if len(item) == 1:
cont += 1
return cont
print [(m,puntos_fijos(f,m)) for m in prime_range(7,100)]
[(m,puntos_fijos(f,m)) for m in prime_range(7,1000) if puntos_fijos(f,m) != 2]
```
Parece que para todos los $m$ primos sólo hay dos puntos fijos, y podemos preguntarnos si es verdad que cuando $m$ es compuesto hay siempre algún otro punto fijo.
```
print [m for m in srange(7,1000) if not is_prime(m) and puntos_fijos(f,m) == 2]
```
<p>¿Qué tienen en común todos estos $m$?</p>
```
print [(m,factor(m)) for m in srange(7,1000) if not is_prime(m) and puntos_fijos(f,m) == 2]
```
<p>Parece claro que todos son de la forma $p^k$ con $p$ primo. Nuestra nueva conjetura:</p>
<p>"El número de puntos fijos es exactamente $2$, que son entonces $0$ y $1$, si y sólo si $m$ es de la forma $p^k$ con $p$ primo."</p>
```
def comprobador_pf(N):
L,L1 = [],[]
for m in xsrange(2,N):
if puntos_fijos(f,m) == 2 and len(list(factor(m))) != 1:
L.append(m)
if puntos_fijos(f,m) != 2 and len(list(factor(m))) == 1:
L1.append(m)
return L,L1
%time comprobador_pf(100)
%time comprobador_pf(1000)
## %time comprobador_pf(10000)
```
Cálculo interrumpido. El tiempo de cálculo no parece lineal en $N$. ¿Se puede mejorar el código para que pueda efectuar esta comprobación en un tiempo razonable?
<h4>Ciclos:</h4>
Hemos visto que para $m=17$ los únicos ciclos son los puntos fijos. ¿Qué tiene de particular el número $17$? En muchas ocasiones, cuando tratamos con números primos $p$, lo que importa, ya que $p$ no se factoriza, es la factorización de $p-1$. En este caso, el $17$ es muy especial ya que $p-1=16=2^4$ es una potencia de $2$. Busquemos primero primos de la forma $2^n+1$:
```
print [(n,2^n+1) for n in xsrange(1,20) if is_prime(2^n+1)]
print [(n,2^n+1) for n in xsrange(1,100) if is_prime(2^n+1)]
def comprobador_no_ciclos(N):
L,L1 = [],[]
for m in prime_range(3,N):
if len(ciclosn(f,m)) == 2 and (len(list(factor(m-1))) != 1 or list(factor(m-1))[0][0] != 2) :
L.append(m)
if len(ciclosn(f,m)) != 2 and (len(list(factor(m-1))) == 1 and list(factor(m-1))[0][0] == 2) :
L1.append(m)
return L,L1
%time comprobador_no_ciclos(100)
%time comprobador_no_ciclos(1000)
```
Parece cierto que todos los casos, con $m$ primo, en que no hay ciclos diferentes a los puntos fijos se trata de enteros de la forma $2^n+1$ que son primos. ¿Hay más primos de esa forma? No se sabe.
Los primos de la forma $2^n+1$ tendrían todos el exponente $n$ de la forma $2^k$, pero los únicos que se conocen son los encontrados un poco más arriba. Se llaman primos de Fermat y no se sabe si existe alguno más, ni si todos los enteros de la forma $2^{2^k}+1$ son compuestos para $k>4.$
Tratemos de estudiar ahora el número de ciclos que aparecen. Probablemente es un problema difícil porque ya hemos visto que para los primeros primos puede haber uno o dos ($m=19$) ciclos.
```
def numero_ciclos(N):
for m in prime_range(3,N):
print m,len(ciclosn(f,m)),factor(m-1)
numero_ciclos(100)
```
Puede parecer que siempre que la factorización de $m-1$ tiene a lo más dos primos el número de ciclos es menor o igual a cuatro, pero probablemente se debe a que sólo llegamos a $100$.
```
numero_ciclos(200)
```
Vemos que $197$ es una excepción, no parece una buena conjetura. ¿Se te ocurre una mejor?
Pensemos ahora en las posibles longitudes de los ciclos. Para el caso $m=19$ hay dos ciclos no triviales de longitudes $2$ y $6$. ¿De qué dependen las posibles longitudes de los ciclos? Sabemos que cada elemento $j$ de $\mathbb{Z}_m$ tiene un orden ($ord(j)$) que es el menor exponente $k$ tal que $j^k$ es, módulo $m$, igual a $1$. Además el orden de $j$ debe ser un divisor del número de elementos no nulos en $\mathbb{Z}_m$, es decir de $m-1.$
Para que $j$ genere un ciclo de longitud $k$ tiene que ocurrir que elevando $j$ a un exponente de la forma $2^k$ se obtenga otra vez $j$, y eso sólo puede pasar si $2^k-1$ es múltiplo de $ord(j)$. ¿Por qué?
Comprobemos en el caso $m=19$:
Los posibles órdenes de elementos son divisores de $18$, es decir $1,2,3,6,9$ y $18$.
```
print [(factor(j),5^j%19) for j in srange(1,2^10) if is_power_of_two(j)]
```
Como $5$ elevado a $2^6$ es otra vez $5$ mçódulo $19$, vemos que el $5$ está en una órbita de longitud $6$, y efectivamente ocurre que $2^6-1=7\times 9$.
```
def orden(j,m):
for div in divisors(m-1):
if power_mod(j,div,m)==1:
return div
orden(5,19)
def longitud_ciclos(m):
L = []
DIV = divisors(m-1)
for div in DIV[1:]:
for exp in srange(1,m+1):
if (2^exp-1)%div == 0:
L.append(exp)
break
return L
longitud_ciclos(19)
longitud_ciclos(17)
longitud_ciclos(23)
G2 = DiGraph(diccionario(23))
G2.graphplot().show(figsize=[12,12])
print [(p,longitud_ciclos(p)) for p in prime_range(11,50)]
```
Hay que entender que estas son las longitudes posibles de los ciclos, pero lo que querríamos saber es si todas las longitudes posibles se dan, es decir ¿Todo lo que puede ocurrir ocurre realmente?
```
def ciclosn2(f,m):
L = []
A = set([])
for n in srange(m):
orb = orbita_c(f,n,m)
if f(orb[-1],m)==orb[0] and orb[-1] not in A and len(orb)>1:
L.append(orb)
A = A | set(orb)
return L
def comprobador_long_ciclos(N):
L = []
for p in prime_range(5,N):
L1 = ciclosn2(f,p)
longit_efec = map(len,L1)
longit_pos = longitud_ciclos(p)
##print longit_efec,longit_pos
if set(longit_efec) != set(longit_pos):
L.append(p)
return L
%time comprobador_long_ciclos(1000)
```
Hasta donde hemos comprobado se cumple que todas las longitudes posibles de los ciclos, para cada primo $p$, ocurren realmente entre las que realmente existen. Hay muchas más preguntas que podemos hacernos:
1. Algunos ciclos, como el que tenemos de longitud $10$ para el primo $23$ tienen la propiedad de que para todo elemento del ciclo hay otro, y sólo uno, que cae en él ¿Es ésto un fenómeno general? Es decir, es verdad que existe un entero $i$, dependiente de una aritmética similar a la que determina las longitudes posibles de los ciclos, tal que todo elemento del ciclo es el final de una cadena de $i$ elementos? Si pudiéramos demostrar algo así podríamos, probablemente, determinar el número de ciclos.
2. ¿Qué podemos decir cuando el entero $m$ es compuesto? Podríamos empezar estudiando el caso de enteros que son el producto de dos primos, y luego, el caso en que todos los primos en la factorización tienen exponente $1$.
| github_jupyter |
```
import numpy as np
from numpy.ma import masked_values as maval
import gsw
import xarray as xr
import pandas as pd
import os.path as op
from datetime import datetime, timedelta
from scipy.interpolate import PchipInterpolator as pchip
from scipy.signal import medfilt
import dask.array as dsar
import cartopy.crs as ccrs
import cartopy.feature as cfeature
from matplotlib import cm
import matplotlib.colors as clr
import matplotlib.ticker as tick
import matplotlib.path as mpath
import matplotlib.pyplot as plt
%matplotlib inline
import cmocean as cmocean
import os
SOCCOM_flt_ids = [12888]
# Load Float Data
HRdir = "/Users/dhruvbalwada/work_root/sogos/data/raw/SOCCOM/manual_download/SOCCOM_HRQC_LIAR_netcdf_20191201/"
LRdir = "/Users/dhruvbalwada/work_root/sogos/data/raw/SOCCOM/manual_download/SOCCOM_LRQC_LIAR_netcdf_20191201/"
float_dic_HR = {}
float_dic_LR = {}
for i in SOCCOM_flt_ids:
float_dic_HR[i] = xr.open_dataset(HRdir + str(i) + "SOOCN_HRQC.nc")
float_dic_LR[i] = xr.open_dataset(LRdir + str(i) + "SOOCNQC.nc")
# Load topography
topo = xr.open_dataarray("/Users/dhruvbalwada/work_root/sogos/data/processed/ETOPO1_Ice_g_gmt4_coarsened5.nc")
plt.figure(figsize=(12, 7))
plt.tight_layout()
ax = plt.subplot(111, projection=ccrs.PlateCarree())
# create a set of axes with Mercator projection
ax.set_xticks(np.linspace(29, 49, 6), crs=ccrs.PlateCarree())
ax.set_yticks(np.linspace(-55.0, -50, 6), crs=ccrs.PlateCarree())
ax.set_extent([28, 45, -55, -50])
ax.add_feature(cfeature.COASTLINE) # plot some data on them
# ax.plot(ds.Lon.values, ds.Lat.values)
# ax.scatter(ds_HR.Lon.values, ds_HR.Lat.values,
# s=25., c=ds_HR.groupby('JULD').mean().JULD.values,
# rasterized=True)
topo.sel(y=slice(-65, -40)).sel(x=slice(25, 50)).plot.contourf(
ax=ax,
transform=ccrs.PlateCarree(),
cbar_kwargs={"shrink": 0.4},
vmin=-4500, vmax=0,
cmap=cmocean.cm.tempo, levels=np.linspace(-5500,0,20)
)
topo.sel(y=slice(-65, -40)).sel(x=slice(25, 50)).plot.contour(
ax=ax,
transform=ccrs.PlateCarree(),
levels=[0], colors='w'
)
for i in [12888]:
ax.plot(float_dic_HR[i].Lon.values, float_dic_HR[i].Lat.values, "o-",
rasterized=True, color='r')
for j in range(len(float_dic_HR[i].Lon)):
plt.text(float_dic_HR[i].Lon[j], float_dic_HR[i].Lat[j], str(j))
#ax.text(float_dic_HR[i].Lon.values[-1], float_dic_HR[i].Lat.values[-1], str(i))
ax.gridlines()
ax.set_title("12888") # label it
ds_flt = float_dic_HR[12888]
ds_flt
dist = gsw.distance(ds_flt.Lon, ds_flt.Lat)
distance = np.cumsum(dist)
distance = np.insert(distance, 0, 0)
ds_flt["distance"] = distance
# Calculate density and spice
## Add density and other things
SA = xr.apply_ufunc(
gsw.SA_from_SP,
ds_flt.Salinity,
ds_flt.Pressure,
ds_flt.Lon,
ds_flt.Lat,
dask="parallelized",
output_dtypes=[float],
).rename("SA")
CT = xr.apply_ufunc(
gsw.CT_from_t,
SA,
ds_flt.Temperature,
ds_flt.Pressure,
dask="parallelized",
output_dtypes=[float],
).rename("CT")
SIGMA0 = xr.apply_ufunc(
gsw.sigma0, SA, CT, dask="parallelized", output_dtypes=[float]
).rename("SIGMA0")
SPICE = xr.apply_ufunc(
gsw.spiciness0, SA, CT, dask="parallelized", output_dtypes=[float]
).rename("SPICE")
ds_flt = xr.merge([ds_flt, SIGMA0, SPICE, SA, CT])
plt.plot(ds_flt.JULD, ds_flt.distance / 1e3, Marker="o")
plt.xlabel("Time")
plt.ylabel("Along Track Distance(km)")
pd.to_datetime(ds_flt.JULD[i].values).month
ncasts = len(ds_flt.Lon)
plt.figure(figsize=(8,7))
for i in range(ncasts):
plt.plot(ds_flt.SIGMA0.isel(N_PROF=i),
ds_flt.Pressure.isel(N_PROF=i),
color=cmocean.cm.thermal(i / ncasts))
plt.text(ds_flt.SIGMA0.isel(N_PROF=i)[-20],0,
str(pd.to_datetime(ds_flt.JULD[i].values).month))
plt.gca().invert_yaxis()
plt.xlabel('Sigma0')
plt.ylabel('Depth')
plt.ylim([500, 0])
plt.xlim([26.8, 27.6])
plt.figure(figsize=(8,7))
for i in range(ncasts):
plt.plot(
ds_flt.SA.isel(N_PROF=i),
ds_flt.Pressure.isel(N_PROF=i),
color=cmocean.cm.thermal(i / ncasts),
)
plt.gca().invert_yaxis()
plt.xlabel('Salinity')
plt.ylabel('Depth')
plt.ylim([500, 0])
#plt.xlim([26.8, 27.6])
plt.tight_layout()
plt.figure(figsize=(8,7))
for i in range(ncasts):
plt.plot(
ds_flt.CT.isel(N_PROF=i),
ds_flt.Pressure.isel(N_PROF=i),
color=cmocean.cm.thermal(i / ncasts),
)
if (np.mod(i,2)==0):
plt.text(ds_flt.CT.isel(N_PROF=i)[-20],0,
str(pd.to_datetime(ds_flt.JULD[i].values).month))
plt.gca().invert_yaxis()
plt.xlabel('Temp')
plt.ylabel('Depth')
plt.ylim([500, 0])
#plt.xlim([26.8, 27.6])
plt.tight_layout()
```
# N2
```
from scipy.interpolate import PchipInterpolator
# using pchip interpolator because it supposed to be the best!
def pchip_interp(y, pres, pint):
pres_temp = -pres[(np.isfinite(pres) & np.isfinite(y))]
y = y[(np.isfinite(pres) & np.isfinite(y))]
[pres_unique, unique_ids] = np.unique(pres_temp, return_index=True)
y = y[unique_ids]
f = PchipInterpolator(pres_unique, y, extrapolate=False)
return f(-pint)
Paxis = np.linspace(0, 2000, 1001)
dens_int = np.zeros((len(ds_flt.N_PROF), len(Paxis)))
for i in range(len(ds_flt.N_PROF)):
dens_int[i, :] = pchip_interp(ds_flt.SIGMA0.isel(N_PROF=i).values,
ds_flt.Pressure.isel(N_PROF=i).values,Paxis)
dens_int = xr.DataArray(dens_int, coords=[ds_flt.N_PROF, Paxis], dims=['N_PROF', 'Pressure']).rename('SIGMA0')
ds_flt_int = dens_int.to_dataset()
vars_to_int = ['Temperature', 'Salinity', 'Oxygen', 'OxygenSat', 'Nitrate', 'Chl_a',
'Chl_a_corr', 'b_bp700', 'POC', 'pHinsitu', 'TALK_LIAR', 'DIC_LIAR']
temp_int = np.zeros((len(ds_flt.N_PROF), len(Paxis)))
for var in vars_to_int:
for i in range(len(ds_flt.N_PROF)):
temp_int[i, :] = pchip_interp(ds_flt[var].isel(N_PROF=i).values,
ds_flt.Pressure.isel(N_PROF=i).values,Paxis)
ds_temp_int = xr.DataArray(temp_int, coords=[ds_flt.N_PROF, Paxis], dims=['N_PROF', 'Pressure']).rename(var)
ds_flt_int[var] = ds_temp_int.copy()
dist = xr.DataArray(distance, coords=[ds_flt.N_PROF], dims='N_PROF')
ds_flt_int =ds_flt_int.assign_coords(dist=dist)
from xgcm import generate_grid_ds, Grid
ds_flt_int = generate_grid_ds(ds_flt_int, {'Z':'Pressure'} )
grid = Grid(ds_flt_int)
g=9.81
rho0=1000
ds_flt_int['N2'] = grid.interp(g/rho0 * grid.diff(ds_flt_int.SIGMA0, 'Z', boundary='extend') /
grid.diff(ds_flt_int.Pressure, 'Z', boundary='extend'), 'Z', boundary='extend')
def find_mld(dens):
dens10 = dens.interp(Pressure = 10.)
delta_dens = dens - dens10
delta_dens_crit = delta_dens.where(delta_dens>=0.03)
MLD = delta_dens.Pressure.where(delta_dens==delta_dens_crit.min(['Pressure'])).max(['Pressure']).rename('mld')
return MLD
MLD = find_mld(ds_flt_int.SIGMA0)
ds_flt_int['MLD'] = MLD
plt.figure(figsize=(12,4))
plt.subplot(121)
plt.pcolormesh(ds_flt.JULD, ds_flt_int.Pressure, ds_flt_int.N2.T,
vmin=-5e-5, vmax=5e-5, cmap='RdBu_r')
plt.plot(ds_flt.JULD, ds_flt_int.MLD, Marker='.')
plt.gca().invert_yaxis()
plt.ylim([400, 0])
plt.colorbar()
plt.xlabel('Time')
plt.ylabel('Depth')
plt.title('N2')
plt.subplot(122)
plt.pcolormesh(ds_flt.JULD, ds_flt_int.Pressure, ds_flt_int.SIGMA0.T,
cmap=cmocean.cm.dense, vmax=27.6)
plt.plot(ds_flt.JULD, ds_flt_int.MLD, Marker='.')
plt.gca().invert_yaxis()
plt.ylim([400, 0])
plt.colorbar()
plt.xlabel('Time')
plt.ylabel('Depth')
plt.title('Density')
plt.tight_layout()
#plt.savefig('../figures/12888_N2_dens.png')
ds_flt.JULD
time = pd.to_datetime(ds_flt.JULD.values).month + pd.to_datetime(ds_flt.JULD.values).day/30
time
plt.contourf(time, ds_flt_int.Pressure, ds_flt_int.SIGMA0.T,
vmin=27, vmax=27.6)
plt.figure(figsize=(12,4))
plt.subplot(121)
plt.pcolormesh(ds_flt.JULD, ds_flt_int.Pressure, ds_flt_int.N2.T,
vmin=-5e-5, vmax=5e-5, cmap='RdBu_r')
plt.plot(ds_flt.JULD, ds_flt_int.MLD, Marker='.')
plt.gca().invert_yaxis()
plt.ylim([400, 0])
plt.colorbar()
plt.xlabel('Time')
plt.ylabel('Depth')
plt.title('N2')
plt.subplot(122)
plt.contourf(time, ds_flt_int.Pressure, ds_flt_int.SIGMA0.T,
vmin=26.9, vmax=27.6, levels=15 )
plt.plot(time, ds_flt_int.MLD, Marker='.', color='k')
for i in range(len(time)):
plt.text(time[i], 0 , str(i))
#plt.plot(ds_flt.JULD, ds_flt_int.MLD, Marker='.')
plt.gca().invert_yaxis()
plt.ylim([400, 0])
plt.colorbar()
plt.xlabel('Time')
plt.ylabel('Depth')
plt.title('Density')
plt.tight_layout()
plt.figure(figsize=(12,4))
plt.subplot(121)
plt.contourf(time, ds_flt_int.Pressure, ds_flt_int.Temperature.T,
levels=15 , cmap='RdBu_r')
plt.plot(time, ds_flt_int.MLD, Marker='.', color='k')
for i in range(len(time)):
plt.text(time[i], 0 , str(i))
plt.gca().invert_yaxis()
plt.ylim([400, 0])
plt.colorbar()
plt.xlabel('Time')
plt.ylabel('Depth')
plt.title('Temp')
plt.subplot(122)
plt.contourf(time, ds_flt_int.Pressure, ds_flt_int.Salinity.T,
levels=15 )
plt.plot(time, ds_flt_int.MLD, Marker='.', color='k')
for i in range(len(time)):
plt.text(time[i], 0 , str(i))
plt.gca().invert_yaxis()
plt.ylim([400, 0])
plt.colorbar()
plt.xlabel('Time')
plt.ylabel('Depth')
plt.title('Temp')
plt.figure(figsize=(8,7))
for i in range(ncasts):
plt.plot(
ds_flt_int.N2.isel(N_PROF=i),
ds_flt_int.Pressure,
color=cmocean.cm.thermal(i / ncasts),
)
plt.gca().invert_yaxis()
plt.xlabel('Temp')
plt.ylabel('Depth')
plt.ylim([500, 0])
#plt.xlim([26.8, 27.6])
plt.tight_layout()
```
| github_jupyter |
```
from keras.datasets import mnist
(X_train, Y_train), (X_test, Y_test) = mnist.load_data()
print('X_train :', X_train.shape)
print('Y_train :', Y_train.shape)
print('X_test :', X_test.shape)
print('Y_test :', Y_test.shape)
import matplotlib.pyplot as plt
from random import randint
for i in range(9):
plt.subplot(331 + i)
plt.imshow(X_train[i+randint(0,59990)])
plt.show()
X_train = X_train.reshape((X_train.shape[0], 28, 28, 1))
X_test = X_test.reshape((X_test.shape[0], 28, 28, 1))
from keras.utils import to_categorical
Y_train = to_categorical(Y_train)
Y_test = to_categorical(Y_test)
Y_train[567]
X_train = X_train.astype('float')
X_test = X_test.astype('float')
X_train = X_train / 255.0
X_test = X_test / 255.0
X_train[234,12]
from keras.models import Sequential
from keras.layers import Conv2D
from keras.layers import MaxPooling2D
from keras.layers import Dense
from keras.layers import Flatten
from keras.optimizers import SGD
def define_model():
model = Sequential()
model.add(Conv2D(32, kernel_size = (3, 3), activation = 'relu', kernel_initializer = 'he_uniform'))
model.add(MaxPooling2D((2, 2)))
model.add(Conv2D(64, kernel_size = (2, 2), activation = 'relu'))
model.add(MaxPooling2D(2, 2))
model.add(Conv2D(256, kernel_size = (2, 2), activation = 'relu'))
model.add(Flatten())
model.add(Dense(1024, activation = 'relu', kernel_initializer = 'he_uniform'))
model.add(Dense(256, activation = 'relu'))
model.add(Dense(10, activation = 'softmax'))
opt = SGD(learning_rate = 0.01, momentum = 0.9)
model.compile(optimizer = opt, loss = 'categorical_crossentropy', metrics = ['accuracy'])
return model
def define_model2():
model = Sequential()
model.add(Conv2D(32, (3, 3), activation='relu', kernel_initializer='he_uniform', input_shape=(28, 28, 1)))
model.add(MaxPooling2D((2, 2)))
model.add(Conv2D(64, (3, 3), activation='relu', kernel_initializer='he_uniform'))
model.add(Conv2D(64, (3, 3), activation='relu', kernel_initializer='he_uniform'))
model.add(MaxPooling2D((2, 2)))
model.add(Flatten())
model.add(Dense(100, activation='relu', kernel_initializer='he_uniform'))
model.add(Dense(10, activation='softmax'))
opt = SGD(lr=0.01, momentum=0.9)
model.compile(optimizer=opt, loss='categorical_crossentropy', metrics=['accuracy'])
return model
from sklearn.model_selection import KFold
def evaluate_model(X_data, Y_data, n_folds = 5):
scores, histories = list(), list()
kFold = KFold(n_folds, shuffle = True)
for i_train, i_test in kFold.split(X_data):
model = define_model()
X_train, Y_train, X_test, Y_test = X_data[i_train], Y_data[i_train], X_data[i_test], Y_data[i_test]
history = model.fit(X_train, Y_train, epochs = 10, batch_size = 32, validation_data = (X_test, Y_test), verbose = 1)
_, acc = model.evaluate(X_test, Y_test, verbose = 1)
print('> %.3f' % (acc*100.0))
scores.append(acc)
histories.append(history)
return scores, histories
scores, histories = evaluate_model(X_train, Y_train)
from numpy import mean
from numpy import std
def performance(scores, histories):
for i in range(len(histories)):
plt.subplot(2,2,1)
plt.title('Cross Entropy Loss')
plt.plot(histories[i].history['loss'], color = 'blue', label = 'train')
plt.plot(histories[i].history['val_loss'], color = 'red', label = 'test')
plt.subplot(2,2,2)
plt.title('Classification Accuracy')
plt.plot(histories[i].history['accuracy'], color = 'blue', label = 'train')
plt.plot(histories[i].history['val_accuracy'], color = 'red', label = 'test')
plt.show()
print('Accuracy: \nMean = %.3f, std = %.3f, n = %d' % (mean(scores)*100, std(scores)*100, len(scores)))
performance(scores, histories)
def load_dataset():
(X_train_orig, Y_train_orig), (X_test_orig, Y_test_orig) = mnist.load_data()
X_train = X_train_orig.reshape((X_train_orig.shape[0], 28, 28, 1))
X_test = X_test_orig.reshape((X_test_orig.shape[0], 28, 28, 1))
Y_train = to_categorical(Y_train_orig)
Y_test = to_categorical(Y_test_orig)
return X_train, Y_train, X_test, Y_test
def preprocessing(X_train, X_test):
X_train = X_train.astype('float')
X_test = X_test.astype('float')
X_train = X_train / 255.0
X_test = X_test / 255.0
return X_train, X_test
def run_all():
X_train, Y_train, X_test, Y_test = load_dataset()
X_train, X_test = preprocessing(X_train, X_test)
scores, histories = evaluate_model(X_train, Y_train)
performance(scores, histories)
run_all()
import os
try:
device_name = os.environ['COLAB_TPU_ADDR']
TPU_ADDRESS = 'grpc://' + device_name
print('Found TPU at: {}'.format(TPU_ADDRESS))
except KeyError:
print('TPU not found')
import tensorflow as tf
import os
import tensorflow_datasets as tfds
resolver = tf.distribute.cluster_resolver.TPUClusterResolver(tpu='grpc://' + os.environ['COLAB_TPU_ADDR'])
tf.config.experimental_connect_to_cluster(resolver)
# This is the TPU initialization code that has to be at the beginning.
tf.tpu.experimental.initialize_tpu_system(resolver)
print("All devices: ", tf.config.list_logical_devices('TPU'))
import tensorflow as tf
from keras import backend as K
import keras.backend.tensorflow_backend as tfback
print("tf.__version__ is", tf.__version__)
print("tf.keras.__version__ is:", tf.keras.__version__)
def _get_available_gpus():
"""Get a list of available gpu devices (formatted as strings).
# Returns
A list of available GPU devices.
"""
#global _LOCAL_DEVICES
if tfback._LOCAL_DEVICES is None:
devices = tf.config.list_logical_devices()
tfback._LOCAL_DEVICES = [x.name for x in devices]
return [x for x in tfback._LOCAL_DEVICES if 'device:gpu' in x.lower()]
tfback._get_available_gpus = _get_available_gpus
K.tensorflow_backend._get_available_gpus()
model = define_model()
model.fit(X_train, Y_train, epochs = 20, batch_size = 32, verbose = 1)
model.save('Final.h5')
_, acc = model.evaluate(X_test, Y_test, verbose = 1)
print('Test Accuracy: %.3f' % (acc*100))
import numpy as np
import cv2
def prediction_preprocessing(filename):
img2 = cv2.imread(filename)
gray = cv2.cvtColor(img2, cv2.COLOR_BGR2GRAY)
res = cv2.resize(gray, dsize=(28,28), interpolation=cv2.INTER_CUBIC)
img = res.reshape(1, 28, 28, 1)
img = img.astype('float')
img = img / 255.0
plt.imshow(img2)
return img
"""
def prediction_preprocessing(filename):
image = load_img(filename, color_mode = 'grayscale')
image = img_to_array(image)
image = image.reshape(image.shape[0],image.shape[1])
width, height = image.shape[1], image.shape[0]
img = np.array(image)
offset = int(abs(height-width)/2)
a = (height-width)%2
if width>height:
img = img[:,offset:(width-offset)]
if a:
img = img[:, 1:]
else:
img = img[offset:(height-offset),:]
if a:
img = img[1:, :]
"""
def prediction(filename):
img = prediction_preprocessing(filename)
model = load_model('Final.h5') # If you have not run the above cells to train the model, use this trained model
digit = model.predict_classes(img)
print('Predicted Output: ', digit)
prediction('new3.jpg')
"""
import cv2
img2 = cv2.imread('lol.jpg')
gray = cv2.cvtColor(img2, cv2.COLOR_BGR2GRAY)
res = cv2.resize(gray, dsize=(28,28), interpolation=cv2.INTER_CUBIC)
rgb_weights = [0.2989, 0.5870, 0.1140]
img_g = np.dot(res[...,:3], rgb_weights)
"""
```
| github_jupyter |
# Análise de Dados com Python - Waffle Charts
**Minicurso:** Análise de Dados com Python
**Instrutor:** Humberto da Silva Neto
**Aluno:**
---
```
import pandas as pd
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches # needed for waffle Charts
mpl.style.use('bmh') # optional: for ggplot-like style
%matplotlib inline
```
Um gráfico de waffle é uma visualização interessante que normalmente é criada para exibir o progresso em direção às metas. Geralmente, é uma opção eficaz quando você está tentando adicionar recursos de visualização interessantes a um visual composto principalmente por células, como um painel do Excel.
Para prosseguir, iremos importar uma base de dados$^{[1]}$ de imigrações para o canada entre os anos de 1980 a 2013.
---
$^{[1]}$ Correções já aplicadas
## Importando a tabela
```
df = pd.read_csv('https://raw.githubusercontent.com/hsneto/mc-adp/master/datasets/canada.csv', index_col=0)
df.head()
```
Vamos rever os dados referentes aos seguintes países: Albania, Brasil e a Islândia.
```
# vamos criar um novo dataframe para esses três países
df_abi = df.loc[['Albania', 'Brazil', 'Iceland'], :]
df_abi
```
## Criando um waffle chart do zero
Infelizmente, ao contrário de R, os waffle charts não são incorporados em nenhuma das bibliotecas de visualização do Python$^{[1]}$. Portanto, aprenderemos como criá-las do zero.
Sendo assim, seguiremos as seguintes etapas:
1. Determinar a proporção de cada categoria em relação ao total.
2. Definir o tamanho total do gráfico de waffle.
3. Determinar o número de peças em função da proporção de cada categoria.
4. Criar uma matriz que se assemelhe ao gráfico de waffle e preenchê-la.
5. Mapear a matriz do gráfico de waffle em um gráfico visual.
6. Criar blocos para melhorar a visualização.
7. Crie uma legenda.
---
$^{[1]}$ Pesquisando, eu encontrei uma biblioteca chamada PyWaffle no [stackoverflow](https://stackoverflow.com/questions/41400136/how-to-do-waffle-charts-in-python-square-piechart). Para mais, acesse a página do [github](https://github.com/ligyxy/PyWaffle).
### Etapa 1: Determinar a proporção de cada categoria em relação ao total
```
# calcular a proporção de cada categoria em relação ao total
total_values = sum(df_abi['Total'])
category_proportions = [float(value)/total_values for value in df_abi['Total']]
# imprime proporções
for i, proportion in enumerate(category_proportions):
print(df_abi.index.values[i] + ':' + str(proportion))
```
### Etapa 2: Definir o tamanho total do gráfico de waffle
```
width = 20
height = 5
num_tiles = height * width
print('número total de peças: ' + str(num_tiles))
```
### Etapa 3: Determinar o número de peças em função da proporção de cada categoria
```
# calcula o número de peças para cada categoria
tiles_per_category = [round(proportion * num_tiles) for proportion in category_proportions]
# imprime o número de peças por categoria
for i, tiles in enumerate(tiles_per_category):
print (df_abi.index.values[i] + ': ' + str(tiles))
```
Isso significa que para cada 100 pessoas que imigraram para o Canadá entre os anos de 1980 a 2013 (considerando somente esses três países), 65 são brasileiros, 34 albanêses e somente 1 é islandes.
### Etapa 4: Criar uma matriz que se assemelhe ao gráfico de waffle e preenchê-la
```
# inicializa o gráfico de waffle como uma matriz vazia
waffle_chart = np.zeros((height, width))
# define índices para percorrer o gráfico de waffle
category_index = 0
tile_index = 0
# preencher o gráfico de waffle
for col in range(width):
for row in range(height):
tile_index += 1
# se o número de blocos preenchidos para a categoria atual for igual aos blocos correspondentes alocados ...
if tile_index > sum(tiles_per_category[0:category_index]):
# ... prossiga para a próxima categoria
category_index += 1
# defina o valor da classe para um inteiro, o que aumenta com a classe
waffle_chart[row, col] = category_index
print ('Waffle chart preenchido!')
```
Vamos dar uma olhada na matriz gerada
```
waffle_chart
```
Como esperado, a matriz consiste em três categorias e o número total de instâncias de cada categoria corresponde ao número total de blocos atribuídos a cada categoria.
Como por exemplo. encontramos apenas um **3.** representando o único imigrante islandes.
### Etapa 5: Mapear a matriz do gráfico de waffle em um gráfico visual
```
plt.matshow(waffle_chart, cmap=plt.cm.gnuplot)
plt.colorbar()
```
Caso você tenha interesse em controlar a palheta de cores das palavras plotadas, sinta-se livre para alterar o argumento **colormap**.
Os links abaixo contém exemplos das palhetas de cores disponíveis:
- [link1](https://matplotlib.org/examples/color/colormaps_reference.html)
- [link2](https://matplotlib.org/users/colormaps.html)
### Etapa 6: Criar blocos para melhorar a visualização
```
# instanciar um novo objeto de figura
fig = plt.figure()
plt.matshow(waffle_chart, cmap=plt.cm.gnuplot)
plt.colorbar()
ax = plt.gca()
ax.set_xticks(np.arange(-.5, (width), 1), minor=True)
ax.set_yticks(np.arange(-.5, (height), 1), minor=True)
# adicione linhas de grade
ax.grid(which='minor', color='w', linestyle='-', linewidth=2)
plt.xticks([])
plt.yticks([])
```
### Etapa 7: Crie uma legenda
```
# instanciar um novo objeto de figura
fig = plt.figure()
plt.matshow(waffle_chart, cmap=plt.cm.gnuplot)
plt.colorbar()
ax = plt.gca()
ax.set_xticks(np.arange(-.5, (width), 1), minor=True)
ax.set_yticks(np.arange(-.5, (height), 1), minor=True)
# adicione linhas de grade
ax.grid(which='minor', color='w', linestyle='-', linewidth=2)
plt.xticks([])
plt.yticks([])
# para calcular a cor do bloco na legenda
total_rows = df_abi['Total'].shape[0] - 1
# criar legenda
legend_handles = []
for i, category in enumerate(df_abi.index.values):
label_str = category + ' (' + str(df_abi['Total'][i]) + ')'
color_val = plt.cm.gnuplot(float(i)/total_rows)
legend_handles.append(mpatches.Patch(color=color_val, label=label_str))
# adicionar legenda no gráfico
plt.legend(handles=legend_handles,
loc='lower center',
ncol=len(df_abi.index.values),
bbox_to_anchor=(0., -0.2, 0.95, .1)
)
```
## Empacotando o código em uma função
```
def waffle_charts(data, width=20, height=5, cmap=plt.cm.coolwarm, colorbar=True, path=None):
def category_proportion(data):
# calcular a proporção de cada categoria em relação ao total
total_values = sum(data)
return [float(value)/total_values for value in data]
def tiles_per_category(category_prop, num_tiles):
# calcula o número de peças para cada categoria
return [round(proportion * num_tiles) for proportion in category_prop]
def populate_waffle_chart(tiles_per_cat, width, height):
# inicializa o gráfico de waffle como uma matriz vazia
waffle_chart = np.zeros((height, width))
# define índices para percorrer o gráfico de waffle
category_index = 0
tile_index = 0
# preencher o gráfico de waffle
for col in range(width):
for row in range(height):
tile_index += 1
# se o número de blocos preenchidos para a categoria atual for igual aos blocos correspondentes alocados ...
if tile_index > sum(tiles_per_cat[0:category_index]):
# ... prossiga para a próxima categoria
category_index += 1
# defina o valor da classe para um inteiro, o que aumenta com a classe
waffle_chart[row, col] = category_index
return waffle_chart
cp = category_proportion(data)
num_tiles = height * width
tiles_cat = tiles_per_category(cp, num_tiles)
waffle_chart = populate_waffle_chart(tiles_cat, width, height)
# instanciar um novo objeto de figura
fig = plt.figure()
plt.matshow(waffle_chart, cmap=cmap)
if colorbar:
plt.colorbar()
ax = plt.gca()
ax.set_xticks(np.arange(-.5, (width), 1), minor=True)
ax.set_yticks(np.arange(-.5, (height), 1), minor=True)
# adicione linhas de grade
ax.grid(which='minor', color='w', linestyle='-', linewidth=2)
plt.xticks([])
plt.yticks([])
# para calcular a cor do bloco na legenda
total_rows = data.shape[0] - 1
# criar legenda
legend_handles = []
for i, category in enumerate(data.index.values):
label_str = category + ' (' + str(data[i]) + ')'
color_val = cmap(float(i)/total_rows)
legend_handles.append(mpatches.Patch(color=color_val, label=label_str))
# adicionar legenda no gráfico
lgd = plt.legend(handles=legend_handles,
loc='lower center',
ncol=len(data.index.values),
bbox_to_anchor=(0., -0.2, 0.95, .1)
)
if path is not None:
plt.savefig(path, dpi=400, bbox_inches='tight', bbox_extra_artist=[lgd])
waffle_charts(df[:10]['Total'], cmap=plt.cm.plasma, colorbar=False)
```
## Utilizando PyWaffle
```
!pip install pywaffle
from pywaffle import Waffle
data = {'Albania': 34, 'Brazil': 65, 'Iceland': 1}
fig = plt.figure(
FigureClass=Waffle,
rows=5,
values=data,
colors=("#232066", "#983D3D", "#DCB732"),
legend={'loc': 'upper left', 'bbox_to_anchor': (1, 1)},
icons='child', icon_size=18,
icon_legend=True
)
```
| github_jupyter |
# Web Scraping with BeautifulSoup
Let's look at how to install beautiful soup and then this will be our mission:
Which presidential candidate is mentioned the most in the politics section of the NY Times? Maybe even a word cloud will be made...
First install beautiful soup:
`pip install bs4`
or
`conda install bs4` (probably)
```
from bs4 import BeautifulSoup
import re ## For parsing
import requests ## For getting the HTML
from wordcloud import WordCloud ## For fun
import backoff
import matplotlib.pyplot as plt
```
### Intro to BeautifulSoup
BeautifulSoup takes the html from webpage and turns into an object that you can work with. All the tags and classes that a webpage have become attributes in a `Soup` object.
```
html_doc = """
<html><head><title>The Dormouse's story</title></head>
<body>
<p class="title"><b>The Dormouse's story</b></p>
<p class="story">Once upon a time there were three little sisters; and their names were
<a href="http://example.com/elsie" class="sister" id="link1">Elsie</a>,
<a href="http://example.com/lacie" class="sister" id="link2">Lacie</a> and
<a href="http://example.com/tillie" class="sister" id="link3">Tillie</a>;
and they lived at the bottom of a well.</p>
<p class="story">...</p>
"""
soup = BeautifulSoup(html_doc)
soup
soup.html
soup.head
soup.title
soup.p
soup.a
soup.find_all('a')
for tag in soup.find_all(True):
print(tag.name)
```
## Now real website!
Let's look at the New york times politics section:
https://www.nytimes.com/news-event/2020-election
Seems like a nice look to work from. Let's go to the browser and check what we can see?
```
## Request the url
url = "https://www.nytimes.com/news-event/2020-election"
r= requests.get(url)
r
soup = BeautifulSoup(r.text, 'lxml')
soup
url_list = []
needed_divs = soup.find_all('div', class_ ="css-1l4spti")
for div in needed_divs:
url_list.append(div.a.get('href'))
url_list
```
Now let's go to each website and do our searches for each candidate. But wait!
### A brief foray into decorators and the `backoff` module
Oftentimes, when you loop through many webpages, the loop might break because you're making too many requests at once.
That's what `backoff` is for. If the website gives an error, `backoff` will catch the exception and make the request again with some pause. The more times the exception is thrown the longer `backoff` will pause before making another one.
`backoff` works as a decorator function. What's that? It's basically a function takes a function as an argument, but returns some "wrapper" for that function that references a function in it... What does that mean?
```
def add():
print('1+2=3')
add()
```
But now you wanted to make sure that people knew that this was a function about addition when they called it. You can write something like this:
```
def i_want_everyone_to_understand(func):
def that_this_is_addition():
print("just in case you didn't know, this is addition")
func()
return that_this_is_addition
add = i_want_everyone_to_understand(add)
add()
```
Instead of writing out the whole function, we can do:
```
@i_want_everyone_to_understand
def add():
print("1+2=3")
add()
```
So `backoff` does this samething, but it catches exceptions of your function. So let's make out requests getter a function so we can use `backoff` with it.
```
@backoff.on_exception(backoff.expo,
requests.exceptions.RequestException)
def requester(url):
return requests.get(url)
```
Now let's make our loop that gets the content from the webpage:
```
base_url = "https://www.nytimes.com"
articles = []
for relative_link in url_list:
site = requester(base_url + relative_link).text
articles.append(site)
print(f"Accessing {base_url + relative_link}")
```
Now let's sift through and see what we can find.
```
name_list = []
for i, article in enumerate(articles):
for name in ['Sanders', 'Trump', 'Warren', 'Buttigieg', 'Klobuchar', 'Yang']:
name_list.extend(re.findall(name,article))
print(f"Going through article {i}")
name_list
%matplotlib inline
name_text = ' '.join(name_list)
wordcloud = WordCloud(background_color="white", collocations=False).generate(name_text)
plt.imshow(wordcloud)
```
| github_jupyter |
# Introduction
```
# Import libraries and check the versions
import pandas as pd
import sys
import numpy as np
import sklearn
import matplotlib as mpl
import seaborn as sns
import missingno as msno
import xgboost as xgb
print('Python version: {}'.format(sys.version))
print('Numpy version {}'.format(np.__version__))
print('Pandas version {}'.format(pd.__version__))
print('Matplotlib version {}'.format(mpl.__version__))
print('Seaborn version {}'.format(sns.__version__))
print('Sklearn version: {}'.format(sklearn.__version__))
print('Missingno version: {}'.format(msno.__version__))
print("Xgboost version: {}".format(xgb.__version__))
# Pretty display for notebooks
%matplotlib inline
import matplotlib.pyplot as plt
plt.style.use('ggplot')
# for more clear plots
from IPython.display import set_matplotlib_formats
set_matplotlib_formats('retina')
```
### 1. Data Collection
This dataset can be found at kaggle's website. First column of the dataset is the index column and we specify that with index_col = 0. Let's see the first five records of the dataset.
```
# retrieve the data
df = pd.read_csv('h1b_kaggle.csv', index_col=[0])
df.head()
```
### 2. Data Wrangling
Before we do explonatary data analysis, we need to select necessary features and clean the data.
```
# select the features that will be used creating the model
data = df[['CASE_STATUS', 'SOC_NAME',
'FULL_TIME_POSITION', 'PREVAILING_WAGE', 'WORKSITE']]
```
Missigno is a library that allows us to visualize missing data in the dataset.
```
# missing values
msno.matrix(data.sample(1000))
msno.dendrogram(data)
#check the missing data
data.isnull().sum()
# remove the missing values
data = data.dropna()
# convert all strings to uppercase
data['SOC_NAME'] = data['SOC_NAME'].str.upper()
# remove everthing after comma from job title
data['SOC_NAME'] = data['SOC_NAME'].apply(lambda x: x.split(', ')[0])
# There
data[data['SOC_NAME'].str.contains('CARPI')]
data = data[~data['SOC_NAME'].str.contains('CARPI')]
```
Current format of the worksite column is **City Name, State**, for this study we will focus on only state.
```
# remove city names from worksite column
data['WORKSITE'] = data['WORKSITE'].apply(lambda x: x.split(', ')[1])
pd.options.display.float_format = '{:,.2f}'.format
data['PREVAILING_WAGE'].describe()
```
Clearly, there are outliers in the dataset.
```
data[(data['PREVAILING_WAGE'] > 500000) | (data['PREVAILING_WAGE'] < 25000)].shape
```
Approxiametly, 12000 wages are below 25000 or above 500000 dollars, those records will be removed.
```
cleaned_data = data[(data['PREVAILING_WAGE'] < 500000)]
cleaned_data = cleaned_data[(cleaned_data['PREVAILING_WAGE'] > 25000)]
```
### 3. Data Exploring
**CASE_STATUS** : This is our target feature. There were 7 possible values in the dataset and we reduced it to 2. Because only one status has a positive result and rest of the statues have a negative result.
**SOC_NAME** : Type of the job. There are 1584 unique jobs in the dataset.
**FULL_TIME_POSITION** : This column indicates if the job is full time or not.
**WORKSITE** : Location of the job. Original column had the state and city information. I removed the cities. The model is going to make predictions based on the state information.
```
# type of columns
cleaned_data.dtypes
print ('Number of records: ', cleaned_data.shape[0])
print ('Number of positive cases: ', cleaned_data['CASE_STATUS'].value_counts()[0])
print ('Number of negative cases: ', cleaned_data['CASE_STATUS'].value_counts()[1])
print ('Percentage of positive cases: ', \
cleaned_data['CASE_STATUS'].value_counts()[0] * 100 / cleaned_data.shape[0])
```
After removing the null values, we still have close to 3 million records. There are 4 features which are SOC_NAME, FULL_TIME_POSITION, PREVAILING_WAGE and WORKSITE. Our target value is CASE_STATUS.
```
cleaned_data['CASE_STATUS'].value_counts().plot(kind='bar', alpha=0.5)
plt.title('Distribution of case statuses')
plt.ylabel('Frequency')
plt.savefig('Distribution_of_case_status.png');
```
We have more positive case results than negative results.
```
# number of unique values in each column
for column in cleaned_data:
print(column, cleaned_data[column].nunique())
cleaned_data['WORKSITE'].groupby(cleaned_data['WORKSITE']).count()\
.sort_values(ascending=False).head(10).plot(kind='bar', alpha=0.5)
plt.title('Top 10 cities for H1-B visa')
plt.savefig('Top_cities.png');
cleaned_data['FULL_TIME_POSITION'].value_counts().plot(kind='bar', alpha=0.5)
plt.title('Distribution of Full Time - Part Time')
plt.ylabel('Frequency');
cleaned_data.groupby(['CASE_STATUS','FULL_TIME_POSITION']).count()['SOC_NAME'].\
unstack().plot(kind='barh',figsize=(12,5), alpha=0.5)
plt.title('Case Status versus Type of position')
plt.ylabel('Frequency');
cleaned_data.pivot_table(values=['CASE_STATUS'], index=['FULL_TIME_POSITION'], aggfunc=('count'))
i = 'PREVAILING_WAGE'
plt.figure(figsize=(10,8))
plt.subplot(211)
plt.xlim(cleaned_data[i].min(), cleaned_data[i].max()*1.1)
ax = cleaned_data[i].plot(kind='kde')
plt.subplot(212)
plt.xlim(cleaned_data[i].min(), cleaned_data[i].max()*1.1)
sns.boxplot(x=cleaned_data[i]);
```
Here we have two plots, the density plot and the box plot. This is a good way to view the data as we can see in the density plot (top) that there is some data points in the tails but it is difficult to see, however it is clear in the box plot.
### 4. Data Transformation and Processing
#### 4.1 Data Transformation
For highly sckewed features, it is always good to do transformation. **PREVAILING_WAGE** column has tail on the right and we will apply logarithmic transformation on it.
```
# log transform the data
cleaned_data['Log_' + i] = np.log(cleaned_data[i])
i = 'Log_PREVAILING_WAGE'
plt.figure(figsize=(10,8))
plt.subplot(211)
plt.xlim(cleaned_data[i].min(), cleaned_data[i].max()*1.1)
ax = cleaned_data[i].plot(kind='kde')
plt.subplot(212)
plt.xlim(cleaned_data[i].min(), cleaned_data[i].max()*1.1)
sns.boxplot(x=cleaned_data[i]);
```
time to scale transformed data
```
# Import sklearn.preprocessing.StandardScaler
from sklearn.preprocessing import RobustScaler
# Initialize a scaler, then apply it to the features
scaler = RobustScaler() # default=(0, 1)
numerical = ['Log_PREVAILING_WAGE']
transformed_data = pd.DataFrame(data = cleaned_data)
transformed_data[numerical] = scaler.fit_transform(cleaned_data[numerical])
# remove original wage column
del transformed_data['PREVAILING_WAGE']
transformed_data['Log_PREVAILING_WAGE'].plot(kind='hist');
```
#### 4.1 Data Processing
```
transformed_data['CASE_STATUS'].unique()
```
There are seven types of case statues but only the "CERTIFIED" have a positive result.
```
# only certified is 1 others 0
transformed_data['CASE_STATUS'] = transformed_data['CASE_STATUS'].apply(lambda x: 1 if x == 'CERTIFIED' else 0)
# One-hot encode the transformed data using pandas.get_dummies()
features_final = pd.get_dummies(transformed_data, columns=['SOC_NAME', 'FULL_TIME_POSITION', 'WORKSITE'])
# Print the number of features after one-hot encoding
encoded = list(features_final.columns)
print ("total features after one-hot encoding: ", len(encoded))
# name of features after one-hot encoding
#print (encoded)
print ("Shape of final features: ", (features_final.shape))
#first 5 rows
features_final.head()
```
4.2 Train-Test Split
```
# select 500,000 samples
features_final = features_final.sample(n=500000)
X = features_final.iloc[:,1:]
y = features_final['CASE_STATUS']
# Import train_test_split
from sklearn.model_selection import train_test_split
# Split the 'features' and 'income' data into training and testing sets
X_train, X_test, y_train, y_test = train_test_split(X,
y,
test_size = 0.2,
random_state = 0)
# Show the results of the split
print ("Training set has samples: ", (X_train.shape[0]))
print ("Testing set has samples: ", (X_test.shape[0]))
```
### 5. Data Modeling
```
import time
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from xgboost import XGBClassifier
# Train logistic regression model
start = time.time()
clf_log = LogisticRegression(random_state = 0)
clf_log.fit(X_train, y_train)
end = time.time()
training_time = end - start
print ("Trainig time - Logistic Regression: ",training_time)
start = time.time()
clf_random = RandomForestClassifier(random_state = 0)
clf_random.fit(X_train, y_train)
end = time.time()
training_time = end - start
print ("Trainig time - Random Forest: ",training_time)
start = time.time()
clf_xg = XGBClassifier(random_state = 0)
clf_xg.fit(X_train, y_train)
end = time.time()
training_time = end - start
print ("Trainig time - XGBoost: ",training_time)
training_times = {'model': ['Logistic Regression', 'Random Forest', 'XGBoost'],
'time': [24, 70, 3038]
}
training_times_df = pd.DataFrame(training_times, columns = ['model','time'])
training_times_df.plot('model', 'time', kind='bar');
```
### 6. Model Evaluation
Naive predictor
```
# Calculate accuracy, precision and recall
TP = np.sum(y) # positive resulted visas
TN = 0
FP = y.count() - np.sum(y) # negative visas
FN = 0
accuracy = TP / (TP + FP)
recall = TP / (TP + FN)
precision = TP / (TP + FP)
# Calculate F-score using the formula above for beta = 0.5 and correct values for precision and recall.
beta = 0.5
fscore = (1 + beta**2) * (precision * recall) / ((beta**2 * precision) + recall)
# Print the results
print ("Naive Predictor\nAccuracy score:", accuracy, "\nF(0.5)-score:" ,fscore)
```
#### Measuring accuracy using Cross Validation
```
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import cross_val_predict
from sklearn.metrics import confusion_matrix
from sklearn.metrics import precision_score, fbeta_score, roc_curve, roc_auc_score, accuracy_score
cross_val_accuracy = cross_val_score(clf_log, X_train, y_train, cv=3, scoring="accuracy").mean()
print ("CV accuracy score:", cross_val_accuracy)
y_train_pred = cross_val_predict(clf_log, X_train, y_train, cv=3)
print ("")
plt.figure(figsize=(10,5))
mat = confusion_matrix(y_train, y_train_pred)
sns.heatmap(mat, square=True, annot=True, fmt='d', cbar=False)
plt.title('Confusion Matrix - Logistic Regression')
plt.ylabel('True labels')
plt.xlabel('Predicted labels');
print ("Precision score: ",precision_score(y_train, y_train_pred))
print ("F(0.5) score", fbeta_score(y_train, y_train_pred, beta=0.5))
y_scores_log = cross_val_predict(clf_log, X_train, y_train, cv=3, method='predict_proba')
y_scores_log = y_scores_log[:,1] # ROC curve requires scores not probability
fpr_log, tpr_log, thresholds_log = roc_curve(y_train, y_scores_log)
plt.plot(fpr_log, tpr_log)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.0])
plt.title('ROC curve for logistic regression')
plt.xlabel('False Positive Rate (1 - Specificity)')
plt.ylabel('True Positive Rate (Sensitivity)')
plt.grid(True)
print ("ROC AUC score: ", roc_auc_score(y_train, y_scores_log))
cross_val_accuracy = cross_val_score(clf_random, X_train, y_train, cv=3, scoring="accuracy").mean()
print ("CV accuracy score:", cross_val_accuracy)
y_train_pred = cross_val_predict(clf_random, X_train, y_train, cv=3)
print ("")
plt.figure(figsize=(10,5))
mat = confusion_matrix(y_train, y_train_pred)
sns.heatmap(mat, square=True, annot=True, fmt='d', cbar=False)
plt.title('Random Forest')
plt.ylabel('True labels')
plt.xlabel('Predicted labels');
print ("Precision_score: ",precision_score(y_train, y_train_pred))
print ("f0.5_score", fbeta_score(y_train, y_train_pred, beta=0.5))
y_scores_random = cross_val_predict(clf_random, X_train, y_train, cv=3, method='predict_proba')
y_scores_random = y_scores_random[:,1] # ROC curve requires scores not probability
fpr_random, tpr_random, thresholds_random = roc_curve(y_train, y_scores_random)
plt.plot(fpr_random, tpr_random)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.0])
plt.title('ROC curve for random forest')
plt.xlabel('False Positive Rate (1 - Specificity)')
plt.ylabel('True Positive Rate (Sensitivity)')
plt.grid(True)
print ("ROC AUC score: ", roc_auc_score(y_train, y_scores_random))
cross_val_accuracy = cross_val_score(clf_xg, X_train, y_train, cv=3, scoring="accuracy").mean()
print ("CV accuracy score:", cross_val_accuracy)
y_train_pred = cross_val_predict(clf_xg, X_train, y_train, cv=3)
print ("")
plt.figure(figsize=(10,5))
mat = confusion_matrix(y_train, y_train_pred)
sns.heatmap(mat, square=True, annot=True, fmt='d', cbar=False)
plt.title('XGBoost')
plt.ylabel('True labels')
plt.xlabel('Predicted labels');
print ("Precision_score: ",precision_score(y_train, y_train_pred))
print ("f0.5_score", fbeta_score(y_train, y_train_pred, beta=0.5))
y_scores_xg = cross_val_predict(clf_xg, X_train, y_train, cv=3, method='predict_proba')
y_scores_xg = y_scores_xg[:,1] # ROC curve requires scores not probability
fpr_xg, tpr_xg, thresholds_xg = roc_curve(y_train, y_scores_xg)
plt.plot(fpr_xg, tpr_xg)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.0])
plt.title('ROC curve for XGBoost')
plt.xlabel('False Positive Rate (1 - Specificity)')
plt.ylabel('True Positive Rate (Sensitivity)')
plt.grid(True)
print ("ROC AUC score: ", roc_auc_score(y_train, y_scores_xg))
plt.figure()
plt.plot(fpr_log, tpr_log, "b", label='Logistic Regression')
plt.plot(fpr_random, tpr_random, "r", label='Random Forest')
plt.plot(fpr_xg, tpr_xg, "g", label='XGBoost')
plt.plot([0,1], [0,1], "k--")
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.0])
plt.title('ROC curve for all classifiers')
plt.xlabel('False Positive Rate (1 - Specificity)')
plt.ylabel('True Positive Rate (Sensitivity)')
plt.grid(True)
plt.legend(loc='lower right')
plt.show()
```
#### Model tuning
```
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import make_scorer
# Initialize the classifier
clf = LogisticRegression(random_state=0)
# Create the parameters list you wish to tune, using a dictionary if needed.
parameters = {'penalty':['l1','l2']
,'C':[0.1, 1, 5, 10]
,'tol':[0.00001, 0.0001, 0.001]
}
# Make an fbeta_score scoring object using make_scorer()
scorer = make_scorer(fbeta_score, beta=0.5)
# Perform grid search on the classifier using 'scorer' as the scoring method using GridSearchCV()
grid_obj = GridSearchCV(clf, param_grid=parameters, scoring=scorer)
# Fit the grid search object to the training data and find the optimal parameters using fit()
grid_fit = grid_obj.fit(X_train, y_train)
# Get the estimator
best_clf = grid_fit.best_estimator_
print ("Best clf's hyperparameters:\n")
print (best_clf)
# Make predictions using the unoptimized and model
predictions = (clf.fit(X_train, y_train)).predict(X_test)
best_predictions = best_clf.predict(X_test)
# Report the before-and-afterscores
print ("\nUnoptimized model\n------")
print ("Accuracy score on testing data: {:.4f}".format(accuracy_score(y_test, predictions)))
print ("F-score on testing data: {:.4f}".format(fbeta_score(y_test, predictions, beta = 0.5)))
print ("\nOptimized Model\n------")
print ("Final accuracy score on the testing data: {:.4f}".format(accuracy_score(y_test, best_predictions)))
print ("Final F-score on the testing data: {:.4f}".format(fbeta_score(y_test, best_predictions, beta = 0.5)))
```
### Train the model with smaller dataset
```
accuracy_scores = []
f_scores = []
sample_size = [100, 1000, 10000, 100000]
for i in sample_size:
X_train_small = X_train.sample(n=i)
y_train_small = y_train.sample(n=i)
# train the small dataset with best classifier
best_clf.fit(X_train_small, y_train_small)
#make predictions
predictions_small = best_clf.predict(X_test)
accuracy_scores.append(accuracy_score(y_test, predictions_small))
f_scores.append(fbeta_score(y_test, predictions_small, beta = 0.5))
accuracy_scores
f_scores
```
### Feature importance
```
features = X_train.columns
importances = clf_random.feature_importances_[:10]
indices = np.argsort(importances)
plt.title('Feature Importances')
plt.barh(range(len(indices)), importances[indices], color='b', align='center')
plt.yticks(range(len(indices)), features[indices])
plt.xlabel('Relative Importance')
plt.show()
```
| github_jupyter |
# Project Green City Taxi .
# Contents of the Repo ,
```
# Getting Dendencies
import pandas as pd
import numpy as np
from datetime import datetime
import datetime as dt
from datetime import timedelta # it is a spanned based time .
import matplotlib.pyplot as plt
import os, json, requests, pickle
from scipy.stats import skew
from shapely.geometry import Point,Polygon,MultiPoint,MultiPolygon
from scipy.stats import ttest_ind, f_oneway, lognorm, levy, skew, chisquare
#import scipy.stats as st
from sklearn.preprocessing import normalize, scale
from tabulate import tabulate #pretty print of tables. source: http://txt.arboreus.com/2013/03/13/pretty-print-tables-in-python.html
from shapely.geometry import Point,Polygon,MultiPoint
%matplotlib inline
import warnings
warnings.filterwarnings('ignore')
# Firs Step . Download Dataset and Print out the size.
if os.path.exists('green_tripdata_2019-01.csv'):# check if the Data is present and load it .
Data= pd.read_csv('green_tripdata_2019-01.csv')
else: # Download Data Jan if not available on desktop .
url = "https://s3.amazonaws.com/nyc-tlc/trip+data/green_tripdata_2019-01.csv"
Data= pd.read_url('url')
Data= Data.to_csv(url.split('/')[-1])
# print number rows and number of columns
print(Data.shape[0])
print(Data.shape[1])
Data
Data['lpep_pickup_datetime'] = Data.get([12/21/18 15:17])
# define the figure with 2 subplots
fig,ax = plt.subplots(1,2,figsize = (15,4))
# Histogram of Number of Trip distance
Data.trip_distance.hist(bins=30 , ax = ax[0])
ax[0].set_xlabel('Trip Distance(per mile)')
ax[0].set_ylabel('Count')
ax[0].set_yscale('log')
ax[0].set_title('A. Histogram of Trip Distance With outliers included')
# Create a vector to contain Trip Distance
v = Data.trip_distance
# # Exclude any data point that is further than 4 standard deviations of median point
# and plot hist with 40 bin
v[~((v-v.median()).abs()>4* v.std())].hist(bins=40, ax= ax[1])
ax[1].set_xlabel('Trip Distance (per mile)')
ax[1].set_ylabel('Trip Count')
ax[1].set_title(' B . Histogram of Trip without outliers')
# Apply a lognormal fit . use the mean of Trip Distancce as the scale parameter
Scatter,loc,mean = lognorm.fit(Data.trip_distance.values,
scale = Data.trip_distance.mean(),
loc = 0 )
pdf_fitted = lognorm.pdf(np.arange(0,14,.1), Scatter , loc, mean)
ax[1].plot(np.arange(0,14,.1), 500000 * pdf_fitted, 'r')
ax[1].legend (['Data', 'lognorm fit '])
# # # create a vector to contain Trip Distance
v = Data.trip_distance
# # exclude any data point located further than 3 standard deviations of the median point and
# # plot the histogram with 30 bins
# apply a lognormal fit. Use the mean of trip distance as the scale parameter
scatter,loc,mean = lognorm.fit(Data.trip_distance.values,
scale=Data.trip_distance.mean(),
loc=0)
pdf_fitted = lognorm.pdf(np.arange(0,12,.1),scatter,loc,mean)
ax[1].plot(np.arange(0,12,.1),600000*pdf_fitted,'r')
ax[1].legend(['Data','Lognormal Fit'])
# # export the figure
# plt.savefig('Question2.jpeg',format='jpeg')
# plt.show()
```
The Trip Distance is asymmetrically distributed.It is skewed to the right and it has a median smaller than its mean and both smaller than the standard deviation. The skewness is due to the fact that the variable has a lower boundary of 0.The distance can't be negative. https://www.itl.nist.gov/div898/handbook/eda/section3/eda3669.htm
To the left is plotted the distribution of the entire raw set of Trip distance. To the right, outliers have been removed before plotting. Outliers are defined as any point located further than 3 standard deviations from the mean
The hypothesis: The trips are not random. If there were random, we would have a (symmetric) Gaussian distribution. The non-zero autocorrelation may be related the fact that people taking ride are pushed by a common cause, for instance, people rushing to work.
# We are going to examine if the time of the day has any impact on the trip distance.
```
# First, let's convert pickup and drop off datetime variable in their specific right format.
# Data['Pickup_dt'] = Data.lpep_pickup_datetime.apply(lambda x:dt.datetime.strptime(x,"%m/%d/%Y %H:%M:%S"))
# yields_Data['Pickup_dt'] = pd.to_datetime(yields_Data['lpep_pickup_datetime'], infer_datetime_format=True)
# def main():
# # time and dates can be formatted using a set of predefined string
# # control code .
# now = datetime.now()
# # print(now.strftime("The current years is:%Y"))
# # %I /%H - 12/24 Hour , %M -Minute, %S - second , %p - locale`s AM/PM
# # print(now.strftime("The current time is : %I:%M:%S %p"))
# # print(now.strftime("24-hour time : %H:%M %p"))
# # print (timedelta(days = 365 , hours = 5 , minutes =1 ))
# # print (" In 2 days and 3 weeks , it will be :" + str(now + timedelta(days = 2, weeks = 3)))
# if __name__=="__main__":
# main();
# Data['Pickup_dt'] = Data.lpep_pickup_datetime.apply(lambda x:dt.datetime.strptime(x,"%m/%d/%Y %H:%M"))
# First, convert pickup and drop off datetime variable in their specific righ format
Data['Pickup_dt'] = Data.lpep_pickup_datetime.apply(lambda x:dt.datetime.strptime(x," %m/%d/%Y %H:%M:%S"))
# data['Dropoff_dt'] = data.Lpep_dropoff_datetime.apply(lambda x:dt.datetime.strptime(x,"%Y-%m-%d %H:%M:%S"))
# Second, create a variable for pickup hours
Data['Pickup_hour'] = Data.Pickup_dt.apply(lambda x:x.hour)
# Mean and Median of trip distance by pickup hour
# I will generate the table but also generate a plot for a better visualization
# fig,ax = plt.subplots(1,1,figsize=(9,5)) # prepare fig to plot mean and median values
# # use a pivot table to aggregate Trip_distance by hour
# table1 = data.pivot_table(index='Pickup_hour', values='Trip_distance',aggfunc=('mean','median')).reset_index()
# # rename columns
# table1.columns = ['Hour','Mean_distance','Median_distance']
# table1[['Mean_distance','Median_distance']].plot(ax=ax)
# plt.ylabel('Metric (miles)')
# plt.xlabel('Hours after midnight')
# plt.title('Distribution of trip distance by pickup hour')
# #plt.xticks(np.arange(0,30,6)+0.35,range(0,30,6))
# plt.xlim([0,23])
# plt.savefig('Question3_1.jpeg',format='jpeg')
# plt.show()
# print '-----Trip distance by hour of the day-----\n'
# print tabulate(table1.values.tolist(),["Hour","Mean distance","Median distance"])
```
| github_jupyter |
```
%%html
<style>
.text_cell_render * {
font-family: OfficinaSansCTT;
}
.reveal code {
font-family: OfficinaSansCTT;
}
.text_cell_render h3 {
font-family: OfficinaSansCTT;
}
.reveal section img {
max-height: 500px;
margin-left: auto;
margin-right: auto;
}
</style>
```
### Вопросы
* Что такое BLAS?
* Как сделать в Numpy вот такую матрицу?
```python
array([[ 0, 4, 8, 12],
[ 1, 5, 9, 13],
[ 2, 6, 10, 14],
[ 3, 7, 11, 15]])
```
* Какой параметр функции read_csv() в Pandas позволяет не читать весь файл сразу целиком в память? И как посмотреть, сколько места датафрейм занимает в памяти?
* Какой метод колонки является аналогом ```df[column].groupby(column).count()```?
* Опишите своими словами, как решается задача предсказания методом линейной регрессии с использованием .
### Что такое DevOps?
- Это набор практик
### Создание полноценного проекта на Python
- Шаг 1: Создаем virtualenv для работы (это мы уже умеем)
- **Шаг 2: Создаем репозиторий на Github**
### Создание полноценного проекта на Python
- Шаг 1: Создаем virtualenv для работы
- Шаг 2: Создаем репозиторий на Github
- **Шаг 3: Описываем аргументы командной строки и конфигурационные файлы (если нужно)**
### Модуль argparse
- Помните **sys.argv**?
```
import argparse
def build_parser():
parser = argparse.ArgumentParser()
parser.add_argument(
'-c', '--config', dest='config', action='store', type=str,
help='path to custom config',
default=os.path.join(os.path.dirname(__file__), "config.yaml")
)
return parser
def main():
parser = build_parser()
params, other_params = parser.parse_known_args()
conf = load_config(params.config)
...
```
### Конфигурационные файлы
- В конфигурационном файле должны быть все настройки программы, которые мы хотим менять без модификации ее кода
- Формат конфига может быть любым, я бы предложил взять YAML, JSON или INI (про INI можно почитать вот тут - https://docs.python.org/3/library/configparser.html)
### Создание полноценного проекта на Python
- Шаг 1: Создаем virtualenv для работы
- Шаг 2: Создаем репозиторий на Github
- Шаг 3: Описываем аргументы командной строки и конфигурационные файлы (если нужно)
- **Шаг 4: Сохраняем зависимости нашего проекта в отдельный файл, который затем включим в пакет**
- ***pip freeze > requirements.txt***
- Посмотрим глазами на содержимое. На некоторых системах эта команда создает ненужную запись о несуществующем пакете *"pkg-resources==0.0.0"* - удалим ее, если она присутствует.
### Создание полноценного проекта на Python
- Шаг 1: Создаем virtualenv для работы
- Шаг 2: Создаем репозиторий на Github
- Шаг 3: Описываем аргументы командной строки и конфигурационные файлы (если нужно)
- Шаг 4: Сохраняем зависимости нашего проекта в отдельный файл, который затем включим в пакет
- **Шаг 5: Структурируем программу, как пакет**
```
my_package <- это папка с нашим проектом
├── MANIFEST.in <- до этого мы сейчас дойдем
├── my_package <- это папка с именем нашего модуля, то, что будет в "import my_package"
│ ├── cli.py <- это базовый файл с нашим кодом
│ ├── config.yaml <- это файл конфигурации
│ └── __init__.py <- это чаще всего пустой файл, который превращает папку в модуль питона
├── requirements.txt <- это наш файл с зависимостями
└── setup.py <- до этого мы сейчас дойдем
```
### Пакеты Python
- Помните эту команду? ***pip install -U pip wheel setuptools***
- Пакеты бывают (в общем случае) двух типов:
- архивы (чаще всего формата .tar.gz)
- бинарные дистрибутивы (формата .whl)
- Главный файл пакета - *setup.py*
- Интерпретатор по умолчанию ищет пакеты в **/usr/lib/python3.6** , либо же в **venv/lib/python3.6/site-packages**
- Можно передавать дополнительные пути для поиска с помощью переменной окружения **PYTHONPATH** или добавляя их в **sys.path**
```
# setup.py
import os
import os.path
from setuptools import find_packages
from setuptools import setup
def find_requires():
dir_path = os.path.dirname(os.path.realpath(__file__))
requirements = []
with open('{0}/requirements.txt'.format(dir_path), 'r') as reqs:
requirements = reqs.readlines()
return requirements
if __name__ == "__main__":
setup(
name="my_package",
version="0.0.1",
description='my cool package',
packages=find_packages(),
install_requires=find_requires(),
include_package_data=True,
entry_points={
'console_scripts': [
'my_command = my_package.cli:main',
],
},
)
```
### MANIFEST.in (включить не-питоновые файлы в проект)
```
include *requirements.txt
recursive-include my_package *
```
### Создание полноценного проекта на Python
- Шаг 1: Создаем virtualenv для работы
- Шаг 2: Создаем репозиторий на Github
- Шаг 3: Описываем аргументы командной строки и конфигурационные файлы (если нужно)
- Шаг 4: Сохраняем зависимости нашего проекта в отдельный файл, который затем включим в пакет
- Шаг 5: Структурируем программу, как пакет
- **Шаг 6 (опциональный): Используем "pip install --editable ." для отладочного режима**
### Пути и import
- Более старый, но стабильный вариант - ***python setup.py develop***
- Гораздо меньше ошибок возникает, если всегда использовать путь от имени пакета
- Любые изменения в файлах сразу же станут видны внутри пакета
- Точнее, не сразу, а после перезапуска интерпретатора
### Создание полноценного проекта на Python
- Шаг 1: Создаем virtualenv для работы
- Шаг 2: Создаем репозиторий на Github
- Шаг 3: Описываем аргументы командной строки и конфигурационные файлы (если нужно)
- Шаг 4: Сохраняем зависимости нашего проекта в отдельный файл, который затем включим в пакет
- Шаг 5: Структурируем программу, как пакет
- Шаг 6 (опциональный): Используем "python setup.py develop" для отладочного режима
- **Шаг 7: Проверяем, что пакет реально собирается**
### Сборка в tar.gz
- ***python setup.py sdist***
- Создает папки **dist** и **%имя проекта%.egg-info**, вторую можно смело удалить (там метаданные)
- Созданный в папке **dist** архив и будем собранным пакетом Python
- https://packaging.python.org/tutorials/distributing-packages/
### Сборка в .whl
- ***python setup.py bdist_wheel*** (есть еще опция --universal, когда проект совместим с Python 2)
- Создает те же папки плюс **build**, где будет примерная структура пакета после инсталляции
- В папке **dist** появится файл *.whl*, который будет бинарным пакетом Python
- Во многих случаях собранный бинарный пакет будет устанавливаться только на ту же ОС, где собирался
### Создание полноценного проекта на Python
- Шаг 1: Создаем virtualenv для работы
- Шаг 2: Создаем репозиторий на Github
- Шаг 3: Описываем аргументы командной строки и конфигурационные файлы (если нужно)
- Шаг 4: Сохраняем зависимости нашего проекта в отдельный файл, который затем включим в пакет
- Шаг 5: Структурируем программу, как пакет
- Шаг 6 (опциональный): Используем "python setup.py develop" для отладочного режима
- Шаг 7: Проверяем, что пакет реально собирается
- **Шаг 8: Отмечаем файлы, которые мы не хотим загружать в систему контроля версий**
### Файл .gitignore
- Файл находится в корне проекта/репозитория. Если его там нет - создайте его
- Просто пишем пути к файлам, которые не хотим отслеживать
### Создание полноценного проекта на Python
- Шаг 1: Создаем virtualenv для работы
- Шаг 2: Создаем репозиторий на Github и клонируем его к себе
- Шаг 3: Описываем аргументы командной строки и конфигурационные файлы (если нужно)
- Шаг 4: Сохраняем зависимости нашего проекта в отдельный файл, который затем включим в пакет
- Шаг 5: Структурируем программу, как пакет
- Шаг 6 (опциональный): Используем "python setup.py develop" для отладочного режима
- Шаг 7: Проверяем, что пакет реально собирается
- Шаг 8: Отмечаем файлы, которые мы не хотим загружать в систему контроля версий
- **Шаг 9: Заливаем код в Git**
### Вспоминаем работу с Git из консоли
- Все команды набираем внутри нашей папки с проектом
- ***git status*** # что вообще происходит
- ***git add путь/к/файлу *** # добавляем файл в git
- ***git add * *** # добавляем все файлы в текущей папке в git
- ***git commit -m "Initial commit"*** # создаем коммит, то есть точку восстановления
- ***git push origin master*** # заливаем код в репозиторий
### Создание полноценного проекта на Python
- Шаг 1: Создаем virtualenv для работы
- Шаг 2: Создаем репозиторий на Github и клонируем его к себе
- Шаг 3: Описываем аргументы командной строки и конфигурационные файлы (если нужно)
- Шаг 4: Сохраняем зависимости нашего проекта в отдельный файл, который затем включим в пакет
- Шаг 5: Структурируем программу, как пакет
- Шаг 6 (опциональный): Используем "python setup.py develop" для отладочного режима
- Шаг 7: Проверяем, что пакет реально собирается
- Шаг 8: Отмечаем файлы, которые мы не хотим загружать в систему контроля версий
- Шаг 9: Заливаем код в Git
- **Шаг 10: Документация**
### Модуль Sphinx
- создадим папку docs
- ***pip install sphinx sphinx-argparse***
- в папке docs: ***sphinx-quickstart***
- лучше задать значения для: "Project name", "Author name(s)", "Project version", "autodoc: automatically insert docstrings from modules" (y), "viewcode: include links to the source code of documented Python objects" (y)
- в файле **conf.py** в *extensions* добавим *'sphinxarg.ext'*, а еще наверху раскомментируем и поправим одну точку на две:
```
import os
import sys
sys.path.insert(0, os.path.abspath('..'))
```
- добавим логотип для красоты:
- ```html_logo = '../logo.png'```
- добавим в **index.rst** после ":caption: Contents:" строчки "quickstart" и "develop"
- это значит, нам надо будет создать два файла - **quickstart.rst** и **develop.rst** в том же каталоге
### quickstart.rst
```rst
Quickstart
==========
.. contents:: :local:
.. argparse::
:module: my_package.cli
:func: build_parser
:prog: my_command
```
### develop.rst
```rst
Reference
=========
.. contents:: :local:
.. automodule:: my_package.cli
:inherited-members:
```
- ***make html*** или ***make.bat html*** - в папке **_build/html** создастся куча файлов, главный - **index.html**
### Создание полноценного проекта на Python
- Шаг 1: Создаем virtualenv для работы
- Шаг 2: Создаем репозиторий на Github и клонируем его к себе
- Шаг 3: Описываем аргументы командной строки и конфигурационные файлы (если нужно)
- Шаг 4: Сохраняем зависимости нашего проекта в отдельный файл, который затем включим в пакет
- Шаг 5: Структурируем программу, как пакет
- Шаг 6 (опциональный): Используем "python setup.py develop" для отладочного режима
- Шаг 7: Проверяем, что пакет реально собирается
- Шаг 8: Отмечаем файлы, которые мы не хотим загружать в систему контроля версий
- Шаг 9: Заливаем код в Git
- Шаг 10: Документация
- **Шаг 11: Тестирование (обсудим в другой раз)**
| github_jupyter |
```
from google.colab import drive
drive.mount('/content/drive/')
!mkdir "/content/Breast_Cancer/"
!unzip "/content/drive/My Drive/ICIAR/Datasets/Macenko.zip" -d /content/Breast_Cancer/
import os
from os.path import basename, join, exists
os.chdir(r"/content/Breast_Cancer/Macenko/")
ls
folder=r"train/"
total=0
print('---Training set details----')
for sub_folder in os.listdir(folder):
no_of_images=len(os.listdir("train/" + sub_folder))
total+=no_of_images
print(str(no_of_images) + " " + sub_folder + " images")
print("Total no. of breast cancer images ",total)
folder=r"test/"
total=0
print('---Test set details----')
for sub_folder in os.listdir(folder):
no_of_images=len(os.listdir("test/" + sub_folder))
total+=no_of_images
print(str(no_of_images) + " " + sub_folder + " images")
print("Total no. of breast cancer images",total)
!mkdir "/content/Breast_Cancer/extracted_features/"
extracted_features_dir="/content/Breast_Cancer/extracted_features/"
import numpy as np
np.random.seed(777)
import time
import keras as keras
from keras.layers import GlobalAveragePooling2D
from keras.preprocessing import image
from keras.preprocessing.image import ImageDataGenerator
from keras.applications.vgg16 import decode_predictions
from keras.models import Sequential
from keras.layers import Dense,Activation,Flatten
from keras.layers import merge,Input
from keras.models import Model
from keras.utils import np_utils
from sklearn.utils import shuffle
from sklearn.model_selection import train_test_split
import os
from keras.preprocessing.image import ImageDataGenerator
from keras.applications.vgg16 import VGG16
from keras.applications.inception_v3 import InceptionV3
from keras.applications.resnet50 import ResNet50
from keras.applications.vgg19 import VGG19
from keras.applications.xception import Xception
from keras.applications.vgg16 import preprocess_input as pi_vgg16
from keras.applications.inception_v3 import preprocess_input as pi_incep
from keras.applications.resnet50 import preprocess_input as pi_resnet
from keras.applications.vgg19 import preprocess_input as pi_vgg19
from keras.applications.xception import preprocess_input as pi_xcep
from keras.models import load_model
from numpy import array
from numpy import argmax
from sklearn.metrics import accuracy_score
from numpy import mean
from numpy import std
import matplotlib.pyplot as plt
from keras.optimizers import Adam,SGD
from keras.callbacks import ReduceLROnPlateau,EarlyStopping,ModelCheckpoint
from keras.layers import GlobalAveragePooling2D, Concatenate
from keras.layers import BatchNormalization,Dropout
from keras.layers import Lambda
from keras.regularizers import l2
import math
from keras import backend as K
from keras.metrics import categorical_accuracy
import warnings
warnings.filterwarnings('always')
warnings.filterwarnings('ignore')
from keras.models import load_model
img_height =512
img_width = 512
batch_size =32
input_shape = (img_width, img_height, 3)
print("-----------------Image Augmentation for Xception--------------")
random_seed = np.random.seed(1142)
train_datagen = ImageDataGenerator(
rescale=1. / 255,
featurewise_center=True,
featurewise_std_normalization=True,
validation_split= 0.2,
zoom_range=0.2)
#shear_range=0.2)
train_generator_xcep = train_datagen.flow_from_directory(
"train/",
target_size=(img_height, img_width),
batch_size=batch_size,
seed = random_seed,
shuffle=False,
subset = 'training',
class_mode='categorical')
val_generator_xcep = train_datagen.flow_from_directory(
"train/",
target_size=(img_height, img_width),
batch_size=batch_size,
seed = random_seed,
shuffle=False,
subset = 'validation',
class_mode='categorical')
test_datagen=ImageDataGenerator(rescale=1./255)
test_generator_xcep=test_datagen.flow_from_directory("test/",
target_size=(img_height, img_width),
batch_size=batch_size,
seed=random_seed,
shuffle=False,
class_mode='categorical') # set as training data
nb_train_samples = len(train_generator_xcep.filenames)
nb_validation_samples = len(val_generator_xcep.filenames)
predict_size_train = int(math.ceil(nb_train_samples / batch_size))
predict_size_validation = int(math.ceil(nb_validation_samples / batch_size))
nb_test_samples = len(test_generator_xcep.filenames)
predict_size_test = int(math.ceil(nb_test_samples / batch_size))
print(nb_train_samples)
print(nb_validation_samples)
print(nb_test_samples)
print(predict_size_train)
print(predict_size_validation)
print(predict_size_test)
model_name="Xception"
model = Xception(include_top=False, weights="imagenet",pooling='avg',input_tensor=Input(shape=input_shape))
model.summary()
for i, layer in enumerate(model.layers):
print(i, layer.name)
image_input =model.input
x1 = GlobalAveragePooling2D()(model.get_layer("block4_sepconv1_act").output) #layer_26
x2 = GlobalAveragePooling2D()(model.get_layer("block5_sepconv1_act").output) #layer_36
x3 = GlobalAveragePooling2D()(model.get_layer("block14_sepconv1").output) #layer_126
out= Concatenate()([x1,x2,x3])
custom_xcep_model = Model(image_input , out)
custom_xcep_model.summary()
for i, layer in enumerate(custom_xcep_model.layers):
print(i, layer.name)
for layer in custom_xcep_model.layers[:95]:
layer.trainable = False
custom_xcep_model.summary()
#Saving features of the training images
bottleneck_features_train = custom_xcep_model.predict_generator(train_generator_xcep, predict_size_train)
np.save(extracted_features_dir+'bottleneck_features_train_'+model_name+'.npy', bottleneck_features_train)
# Saving features of the validation images
bottleneck_features_validation = custom_xcep_model.predict_generator(val_generator_xcep, predict_size_validation)
np.save(extracted_features_dir+'bottleneck_features_validation_'+model_name+'.npy', bottleneck_features_validation)
# Saving features of the test images
bottleneck_features_test = custom_xcep_model.predict_generator(test_generator_xcep, predict_size_test)
np.save(extracted_features_dir+'bottleneck_features_test_'+model_name+'.npy', bottleneck_features_test)
train_data = np.load(extracted_features_dir+'bottleneck_features_train_'+model_name+'.npy')
validation_data = np.load(extracted_features_dir+'bottleneck_features_validation_'+model_name+'.npy')
test_data = np.load(extracted_features_dir+'bottleneck_features_test_'+model_name+'.npy')
train_data=np.load('/content/drive/My Drive/ICIAR/bottleneck_features/Macenko/Xception/bottleneck_features_train_Xception.npy')
validation_data=np.load('/content/drive/My Drive/ICIAR/bottleneck_features/Macenko/Xception/bottleneck_features_validation_Xception.npy')
test_data=np.load('/content/drive/My Drive/ICIAR/bottleneck_features/Macenko/Xception/bottleneck_features_test_Xception.npy')
print(train_data.shape)
print(validation_data.shape)
print(test_data.shape)
train_labels=train_generator_xcep.classes
train_labels=train_labels = keras.utils.to_categorical(train_labels, num_classes=4)
validation_labels=val_generator_xcep.classes
validation_labels = keras.utils.to_categorical(validation_labels, num_classes=4)
test_labels=test_generator_xcep.classes
test_labels=keras.utils.to_categorical(test_labels,num_classes=4)
print(train_labels.shape)
print(validation_labels.shape)
print(test_labels.shape)
model = Sequential()
model.add(Dense(256,activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(4,activation='softmax',name= 'output'))
adam_opt2=Adam(lr = 0.001, beta_1=0.6, beta_2=0.8, amsgrad=True)
model.compile(optimizer=adam_opt2, loss='categorical_crossentropy', metrics=['accuracy'])
history = model.fit(train_data, train_labels,
epochs=1000,
batch_size=batch_size,
validation_data=(validation_data, validation_labels),
verbose= 2)
preds = model.predict(validation_data)
predictions = [i.argmax() for i in preds]
y_true = [i.argmax() for i in validation_labels]
print('Validation Accuracy={}'.format(accuracy_score(y_true=y_true, y_pred=predictions)))
preds = model.predict(test_data)
predictions = [i.argmax() for i in preds]
y_true = [i.argmax() for i in test_labels]
#cm = confusion_matrix(y_pred=predictions, y_true=y_true)
print('Test Accuracy={}'.format(accuracy_score(y_true=y_true, y_pred=predictions)))
```
Accuracy vs Epoch Graph
```
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('Model accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train','Validation'], loc='upper left')
plt.show()
```
Loss vs Epoch Graph
```
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train','Validation'], loc='upper left')
plt.show()
model.save('/content/drive/My Drive/ICIAR/best_saved_models/Macenko/Xception/xception_upto95frozen.h5')
model.save_weights('/content/drive/My Drive/ICIAR/best_saved_models/Macenko/Xception/xception_upto95frozen_weights.h5')
loaded_model=keras.models.load_model('/content/drive/My Drive/ICIAR/best_saved_models/Macenko/Xception/xception_upto95frozen.h5')
loaded_model.load_weights('/content/drive/My Drive/ICIAR/best_saved_models/Macenko/Xception/xception_upto95frozen_weights.h5')
train_labels=train_generator_xcep.classes
train_labels=train_labels = keras.utils.to_categorical(train_labels, num_classes=4)
validation_labels=val_generator_xcep.classes
validation_labels = keras.utils.to_categorical(validation_labels, num_classes=4)
test_labels=test_generator_xcep.classes
test_labels=keras.utils.to_categorical(test_labels,num_classes=4)
validation_data=np.load('/content/drive/My Drive/ICIAR/bottleneck_features/Macenko/Xception/bottleneck_features_validation_Xception.npy')
test_data=np.load('/content/drive/My Drive/ICIAR/bottleneck_features/Macenko/Xception/bottleneck_features_test_Xception.npy')
preds = loaded_model.predict(validation_data)
predictions = [i.argmax() for i in preds]
y_true = [i.argmax() for i in validation_labels]
print('Validation Accuracy={}'.format(accuracy_score(y_true=y_true, y_pred=predictions)))
from sklearn.metrics import confusion_matrix, classification_report
import seaborn as sns
import pandas as pd
preds = loaded_model.predict(test_data)
y_pred = [i.argmax() for i in preds]
y_true = [i.argmax() for i in test_labels]
cm = confusion_matrix(y_pred=y_pred, y_true=y_true)
print('Test Accuracy={}'.format(accuracy_score(y_true=y_true, y_pred=y_pred)))
print('CONFUSION MATRIX')
conf_matrix = pd.DataFrame(data = cm,
columns =['Benign', 'InSitu','Invasive','Normal'],
index =['Benign', 'InSitu','Invasive','Normal'])
accuracy = np.trace(cm) / float(np.sum(cm))
misclass = 1 - accuracy
plt.figure(figsize = (10,8))
sns.heatmap(conf_matrix, annot = True, fmt = 'd', cmap = "Blues")
plt.ylabel('True Label')
plt.xlabel('Predicted label\naccuracy={:0.4f}; misclass={:0.4f}'.format(accuracy, misclass))
plt.show()
target_names=['Benign', 'InSitu','Invasive','Normal']
print('The details for confusion matrix is =')
print (classification_report(y_true, y_pred,target_names=target_names))
sensitivity = cm[0,0]/(cm[0,0]+cm[1,0])
print('Sensitivity : ', sensitivity*100 )
Specificity = cm[1,1]/(cm[1,1]+cm[0,1])
print('Specificity : ', Specificity*100 )
```
prediction matrix
```
import pandas as pd
val_df=pd.read_csv('/content/drive/My Drive/ICIAR/Binary_prediction_matrix.csv')
val_df
preds = loaded_model.predict(validation_data)
predictions = [i.argmax() for i in preds]
y_true = [i.argmax() for i in validation_labels]
val_df['xception']=predictions
val_df
val_df.to_csv('/content/drive/My Drive/ICIAR/Binary_prediction_matrix.csv')
```
Confidence matrix
```
import pandas as pd
test_df=pd.read_csv('/content/drive/My Drive/ICIAR/Binary_confidence_matrix.csv')
test_df
preds = loaded_model.predict(test_data)
for i in range(0,2):
test_df['xception_class'+str(i)]=preds[:,i]
test_df
test_df.to_csv('/content/drive/My Drive/ICIAR/Binary_confidence_matrix.csv')
```
| github_jupyter |
```
# default_exp core
# export
"""Copyright 2020 The Aerospace Corporation"""
# hide
from nbdev.showdoc import *
```
# GPSTime Core Module
> Contains the GPSTime class that represents time as GPS
```
# hide
import sys
sys.path.append("..")
# export
from __future__ import annotations
import datetime
import ruamel.yaml
import numpy as np
from typing import Union, Tuple
from logging import getLogger
from gps_time.datetime import tow2datetime, datetime2tow
logger = getLogger(__name__)
```
## Parameters and Helper Functions
The following parameters are useful values that will be used regularly within this module.
```
# export
_SEC_IN_WEEK: int = 604800
_SEC_TO_FEMTO_SEC: float = 1.0e15
_FEMTO_SEC_TO_SEC: float = 1.0e-15
```
This function is used to convert a time as a float into integer seconds and femtoseconds
```
# export
def _tow2sec(time_of_week: float) -> Tuple[int, int]:
"""Convert a float time to integer seconds and femtoseconds
Parameters
----------
time_of_week : float
The time of week, as a float
Returns
-------
Tuple[int, int]
The seconds and femtoseconds within the time of week
"""
seconds = int(time_of_week // 1)
femtoseconds = int((time_of_week % 1) * _SEC_TO_FEMTO_SEC)
return seconds, femtoseconds
```
## The GPSTime Class
The primary class for this module is, unsurprisingly, the `GPSTime` class. This class has three public attributes, the `week_number`, `seconds`, and `femtoseconds`. The week number is the number of weeks since the start of the GPS epoch, 6 January 1980. The seconds and femtoseconds are the number of seconds and femtoseconds (`1e-15`) since the start of the GPS week (starting midnight Saturday night/Sunday morning).
The `GPSTime` class also has a property of `time_of_week` that returns the time of week as a float. Note that the floating point accuracy of `time_of_week` can be as low as tens of picoseconds (`1e-12`).
The GPSTime class can be save to and loaded from file using `ruamel.yaml`.
```
# export
class GPSTime:
"""Time representation for GPS.
Attributes
----------
week_number : int
The number of weeks since the start of the GPS epoch, 6 Jan 1980.
seconds : int
The number of integer seconds into the week. The zero time is at
midnight on Sunday morning, i.e. betwen Saturday and Sunday. Should
be between 0 and 604800 because otherwise, the week number would be
incorrect.
femtoseconds : int
The number of femtoseconds into the week. That is, this is the number
of fractional seconds in the time of week with a scale factor of 1e15.
Raises
------
TypeError
For various operators if not the selected types are not implemented.
ValueError
If an incorrect set of input arguments are provided to the constructor
Todo
----
.. todo:: Create a GPSTimeDelta class to handle adding/subtracting with
increase accuracy.
"""
weeks: int
seconds: int
femtoseconds: int
yaml_tag: str = u"!GPSTime"
def __init__(self, week_number: int, *args, **kwargs) -> None:
"""Object constructor.
This sets the week number and the time of week and ensures that the
time of week is a float. It also calls `correct_time()`, which checks
to see if the time of week is negative or past the end of the week
and adjust the values accordingly.
This constructor supports many different input arguments. However some
sets of input arguments may result in truncation and errors if a
`float` is provided when an `int` is expected.
Parameters
----------
week_number : int
The number of week
*args, **kwargs
The time of week in various representations. If positional arguments
are used, a single positional argument is interpreted as a time of
week (i.e. a float), while two arguments are interpreted as seconds
and femtoseconds. In the latter case, the values will be cast as
integers, which may result in truncation. Keyword arguments function
in much the same way, with "time_of_week", "seconds", and
"femtoseconds" being the valid keyword arguments. If only "seconds"
is given, it will be treated like "time_of_week". If no additional
arguments are given, the time is assumed to be the start of the week.
Raises
------
ValueError
If invalid arguments are given. Examples include:
- Mixed positional and keyword arguments are not supported
- More than two arguments are not supported
- Keyword arguments "time_of_week" and "femtoseconds" cannot be
used together.
"""
self.yaml_tag: str = u"!GPSTime"
if len(args) > 0 and len(kwargs) > 0:
raise ValueError(
"GPSTime does not support both positional and keyword arguments."
)
self.week_number = int(week_number)
if len(args) > 2: # If more than 3 args (week + 2 times)
raise ValueError(
"Only up to three arguments allowed (Week number, seconds, "
"and femtoseconds)"
)
elif len(args) == 2:
if isinstance(args[0], float) or isinstance(args[1], float):
logger.warning(
"Two times given, but at least one is a float. Decimal "
"values will be truncated"
)
self.seconds = int(args[0])
self.femtoseconds = int(args[1])
elif len(args) == 1:
self.time_of_week = args[0]
else:
if len(kwargs) > 2:
raise ValueError("Too many arguments")
elif len(kwargs) == 0:
logger.warning(
"No time of week information. Defaulting to start of week"
)
self.seconds = 0
self.femtoseconds = 0
elif "femtoseconds" in kwargs:
if "time_of_week" in kwargs:
raise ValueError(
"""Keyword arguments "time_of_week" and "femtoseconds"
are incompatible."""
)
elif "seconds" in kwargs:
if isinstance(kwargs["seconds"], float) or isinstance(
kwargs["femtoseconds"], float
):
logger.warning(
"Two times given, but at least one is a float. "
"Decimal values will be truncated"
)
self.seconds = int(kwargs["seconds"])
self.femtoseconds = int(kwargs["femtoseconds"])
else:
raise ValueError(
"""Keyword argument "femtoseconds" must be
accompanied by "seconds"."""
)
elif "seconds" in kwargs:
logger.warning(
"seconds given with no femtoseconds. Will be handled "
"as time of week"
)
self.time_of_week = float(kwargs["seconds"])
elif "time_of_week" in kwargs:
self.time_of_week = float(kwargs["time_of_week"])
else:
raise ValueError("Invalid Keyword arguments")
self.correct_time()
if self.week_number < 0:
logger.warning("Week number is less than 0")
@property
def time_of_week(self) -> float:
"""The time of week as a float."""
return float(self.seconds + self.femtoseconds * _FEMTO_SEC_TO_SEC)
@time_of_week.setter
def time_of_week(self, time_of_week: float) -> None:
"""A setter for the time of week.
The method allows the seconds and femtoseconds to be updated using
a single float.
Paremeters
----------
time_of_week : float
The time of week as a float
"""
sec, femtosec = _tow2sec(time_of_week)
self.seconds = sec
self.femtoseconds = femtosec
@classmethod
def from_yaml(
cls: type, constructor: ruamel.yaml.Constructor, node: ruamel.yaml.MappingNode
) -> GPSTime:
"""YAML Constructor.
This YAML constructor is used to load a GPSTime from a YAML file. It must be
registered with the YAML loader. This is accomplished using
```python3
import ruamel.yaml
yaml = ruamel.yaml.YAML(typ="unsafe")
yaml.register_class(GPSTime)
```
This class method is primarily used to add a constructor to an instance of
ruamel.yaml. Its functionality as a traditional classmethod is limited.
.. note:: YAML Module
This constructor is meant to be used with ruamel.yaml. It has not been tested
with pyyaml (the more common YAML library.)
"""
nodes = node.value
week_number = None
seconds = None
femtoseconds = None
time_of_week = None
for i in range(0, len(nodes)):
node_name = nodes[i][0].value
if node_name == "week_number":
week_number = constructor.construct_scalar(nodes[i][1])
elif node_name == "seconds":
seconds = constructor.construct_scalar(nodes[i][1])
elif node_name == "femtoseconds":
femtoseconds = constructor.construct_scalar(nodes[i][1])
elif node_name == "time_of_week":
time_of_week = constructor.construct_scalar(nodes[i][1])
if seconds is None and time_of_week is None:
raise ValueError("The YAML file lacked both a time_of_week and a seconds")
if seconds is not None and time_of_week is not None:
raise ValueError(
"YAML file defines both time_of_week and seconds (incompatible)"
)
elif time_of_week is not None and femtoseconds is not None:
raise ValueError(
"YAML file defines both time_of_week and femtoseconds (incompatible)"
)
elif seconds is not None and femtoseconds is None:
seconds, femtoseconds = _tow2sec(float(seconds))
elif time_of_week is not None:
seconds, femtoseconds = _tow2sec(float(time_of_week))
return cls(int(week_number), int(seconds), int(femtoseconds))
def to_datetime(self) -> datetime.datetime:
"""Convert the `GPSTime` to a datetime.
This method calls `tow2datetime()` to convert the `GPSTime` to a
datetime object.
Returns
-------
datetime.datetime
The equivalent datetime representation of the `GPSTime`
Notes
-----
.. note::
Datetimes are limited to microsecond resolution, so this
conversion may lose some fidelity.
"""
return tow2datetime(self.week_number, self.time_of_week)
@classmethod
def from_datetime(cls, time: datetime.datetime) -> GPSTime:
"""Create a `GPSTime` for a datetime.
Parameters
----------
time : datetime.datetime
The datetime that will be converted to a `GPSTime`
Returns
-------
GPSTime
The `GPSTime` corresponding to the datetime. This is a lossless
conversion.
Raises
------
TypeError
If the input value is not a datetime
Notes
-----
This is a classmethod and thus can be called without instantiating the
object first.
"""
if not isinstance(time, datetime.datetime):
raise TypeError("time must be a datetime")
week_num, tow = datetime2tow(time)
return cls(week_num, tow)
def to_zcount(self) -> float:
"""Get the current Z-Count.
Returns
-------
float
The time of week divided by 1.5
"""
return self.time_of_week / 1.5
def correct_weeks(self) -> None:
"""Correct the week number based on the time of week.
If the time of week is less than 0 or greater than 604800 seconds,
then the week number and time of week will be corrected to ensure that
the time of week is within the week indicated by the week number.
Returns
-------
None
"""
logger.warning(
"The correct_weeks() method will be deprecated in a future version. Use the correct_time() method instead."
)
if (self.time_of_week >= _SEC_IN_WEEK) or (self.time_of_week < 0):
weeks_to_add = int(self.time_of_week // _SEC_IN_WEEK)
new_time_of_week = float(self.time_of_week % _SEC_IN_WEEK)
self.week_number += weeks_to_add
self.time_of_week = new_time_of_week
else:
pass
def correct_time(self) -> None:
if (self.femtoseconds >= _SEC_TO_FEMTO_SEC) or (self.femtoseconds < 0):
seconds_to_add = int(self.femtoseconds // _SEC_TO_FEMTO_SEC)
new_femto_sec = int(self.femtoseconds % _SEC_TO_FEMTO_SEC)
self.seconds += seconds_to_add
self.femtoseconds = new_femto_sec
if (self.seconds >= _SEC_IN_WEEK) or (self.seconds < 0):
weeks_to_add = int(self.seconds // _SEC_IN_WEEK)
new_sec = int(self.seconds % _SEC_IN_WEEK)
self.week_number += weeks_to_add
self.seconds = new_sec
def __add__(
self,
other: Union[
int, float, GPSTime, datetime.datetime, datetime.timedelta, np.ndarray
],
) -> Union[GPSTime, np.ndarray]:
"""Addition, apply an offset to a `GPSTime`.
This is the addition of a `GPSTime` and another object. In this
context, addition means moving the clock of the first argument
forward by some amount.
Suppose `a` is a `GPSTime` and the value give for other represents a
positive time. The value returned will be a `GPSTime` object that is
the amount of time represented by other after `a`.
Parameters
----------
other : Union[int, float, GPSTime, datetime.datetime,
datetime.timedelta, np.ndarray]
The other value to add to the `GPSTime`. `int` and `float` values
are the number of seconds to add to the `GPSTime`. `GPSTime` and
`datetime.timedelta` have explicit unit definitions that are used.
If the value is a datetime.datetime, it is converted to a GPSTime
before adding.
Returns
-------
Union[GPSTime, np.ndarray]
The sum of the `GPSTime` and `other`. If other is an np.array,
returns the sum for each element
Raises
------
TypeError
If other is not a supported type
Notes
-----
.. note::
Apart from adding of `datetime.timedelta` objects, this
functionality does not exist with datetimes.
.. note::
This function can be used to "add" a negative amount of time,
which can yield different results than subtraction.
"""
if isinstance(other, bool):
raise TypeError(
"unsupported operand type(s) for -: '{}' and '{}'".format(
type(self), type(other)
)
)
if isinstance(other, int) or isinstance(other, float):
gps_time_to_add = GPSTime(0, float(other))
elif isinstance(other, datetime.timedelta):
gps_time_to_add = GPSTime(0, other.total_seconds())
elif isinstance(other, datetime.datetime):
gps_time_to_add = GPSTime.from_datetime(other)
elif isinstance(other, GPSTime):
gps_time_to_add = other
elif isinstance(other, np.ndarray):
input = np.array([self])
return input + other
else:
raise TypeError(
"unsupported operand type(s) for +: '{}' and '{}'".format(
type(self), type(other)
)
)
week_num = self.week_number + gps_time_to_add.week_number
seconds = self.seconds + gps_time_to_add.seconds
femtoseconds = self.femtoseconds + gps_time_to_add.femtoseconds
return GPSTime(week_num, seconds, femtoseconds)
def __sub__(
self,
other: Union[
int, float, GPSTime, datetime.datetime, datetime.timedelta, np.ndarray
],
) -> Union[GPSTime, float, np.ndarray]:
"""Subtraction.
This method is used to represent subtraction. Depending on the type of
the arguments, it can be used to find the time offset by an amount or
the number of seconds between two times.
Parameters
----------
other : Union[int, float, GPSTime, datetime.datetime,
datetime.timedelta, np.ndarray]
The other value to subtract from the `GPSTime`. `int` and `float`
values are the number of seconds to subtract from the `GPSTime`.
`GPSTime` and `datetime.timedelta` have explicit unit definitions
that are used. If the value is a datetime.datetime, it is
converted to a GPSTime before subtracting.
Returns
-------
Union[GPSTime, float, np.ndarray]
A float will be return if both values are `GPSTime` objects that
represents the number of seconds between the objects. A GPSTime
will be returned otherwise and it represents offsetting the time
backward by the amount given. If the input is an np.ndarray, then
returns the operation for each element
Raises
------
TypeError
If other is not a supported type
Notes
-----
Subtracting a non-`GPSTime` object is equivalent to adding the opposite
of its value
"""
if isinstance(other, bool):
raise TypeError(
"unsupported operand type(s) for -: '{}' and '{}'".format(
type(self), type(other)
)
)
if isinstance(other, int) or isinstance(other, float):
sec_to_sub, femto_to_sub = _tow2sec(float(other))
return GPSTime(
self.week_number,
self.seconds - sec_to_sub,
self.femtoseconds - femto_to_sub,
)
elif isinstance(other, datetime.timedelta):
sec_to_sub, femto_to_sub = _tow2sec(float(other.total_seconds()))
return GPSTime(
self.week_number,
self.seconds - sec_to_sub,
self.femtoseconds - femto_to_sub,
)
elif isinstance(other, datetime.datetime):
other_gpstime = GPSTime.from_datetime(other)
weeks_diff = self.week_number - other_gpstime.week_number
sec_diff = self.seconds - other_gpstime.seconds
femto_diff = self.femtoseconds - other_gpstime.femtoseconds
return float(
weeks_diff * _SEC_IN_WEEK + sec_diff + femto_diff * _FEMTO_SEC_TO_SEC
)
elif isinstance(other, GPSTime):
weeks_diff = self.week_number - other.week_number
sec_diff = self.seconds - other.seconds
femto_diff = self.femtoseconds - other.femtoseconds
return float(
weeks_diff * _SEC_IN_WEEK + sec_diff + femto_diff * _FEMTO_SEC_TO_SEC
)
elif isinstance(other, np.ndarray):
if other.dtype == np.object:
_type = np.reshape(other, sum([i for i in other.shape]))[0].__class__
if _type in (self.__class__, datetime.datetime):
input = np.array([self])
return np.array(input - other, dtype=float)
elif _type is datetime.timedelta:
input = np.array([self])
return np.array(input - other, dtype=object)
elif other.dtype in (
int,
float,
):
input = np.array([self])
return np.array(input - other, dtype=object)
else:
raise TypeError(
"unsupported operand type(s) for -: '{}' and '{}'".format(
type(self), type(other)
)
)
def __lt__(self, other: Union[GPSTime, datetime.datetime]) -> bool:
"""Comparison: Less Than.
.. note:: In this context "less than" is equivalent to "before"
Parameters
----------
other : Union[GPSTime, datetime.datetime]
The object to compare. Datatimes will be converted to `GPSTime`
Returns
-------
bool
True if the current object is before its comparison
Raises
------
TypeError
If an invalid type
"""
if isinstance(other, datetime.datetime):
other_time = GPSTime.from_datetime(other)
elif isinstance(other, GPSTime):
other_time = other
else:
raise TypeError(
"'<' not supported between instances of '{}' and '{}'".format(
type(self), type(other)
)
)
return (self - other_time) < 0
def __gt__(self, other: Union[GPSTime, datetime.datetime]) -> bool:
"""Comparison: Greater Than.
.. note:: In this context "greater than" is equivalent to "after"
Parameters
----------
other : Union[GPSTime, datetime.datetime]
The object to compare. Datatimes will be converted to `GPSTime`
Returns
-------
bool
True if the current object is after its comparison
Raises
------
TypeError
If an invalid type
"""
if isinstance(other, datetime.datetime):
other_time = GPSTime.from_datetime(other)
elif isinstance(other, GPSTime):
other_time = other
else:
raise TypeError(
"'>' not supported between instances of '{}' and '{}'".format(
type(self), type(other)
)
)
return (self - other_time) > 0
def __eq__(self, other: Union[GPSTime, datetime.datetime]) -> bool:
"""Comparison: Equality.
.. note:: In this context "equality" is equivalent to "coincident"
Parameters
----------
other : Union[GPSTime, datetime.datetime]
The object to compare. Datatimes will be converted to `GPSTime`
Returns
-------
bool
True if the current object is the same time as its comparison
Raises
------
TypeError
If an invalid type
"""
if isinstance(other, datetime.datetime):
other_time = GPSTime.from_datetime(other)
elif isinstance(other, GPSTime):
other_time = other
else:
raise TypeError(
"'>' not supported between instances of '{}' and '{}'".format(
type(self), type(other)
)
)
return (self - other_time) == 0
def __le__(self, other: Union[GPSTime, datetime.datetime]) -> bool:
"""Comparison: Less Than or Equals.
Calls the `__lt__()` and `__eq__()` methods
Parameters
----------
other : Union[GPSTime, datetime.datetime]
The object to compare. Datatimes will be converted to `GPSTime`
Returns
-------
bool
True if the current object is before or at the same time as its
comparison object.
Raises
------
TypeError
If an invalid type
"""
return self.__lt__(other) or self.__eq__(other)
def __ge__(self, other: Union[GPSTime, datetime.datetime]) -> bool:
"""Comparison: Greater Than or Equals.
Calls the `__gt__()` and `__eq__()` methods
Parameters
----------
other : Union[GPSTime, datetime.datetime]
The object to compare. Datatimes will be converted to `GPSTime`
Returns
-------
bool
True if the current object is after or at the same time as its
comparison object.
Raises
------
TypeError
If an invalid type
"""
return self.__gt__(other) or self.__eq__(other)
def __ne__(self, other: Union[GPSTime, datetime.datetime]) -> bool:
"""Comparison: Not Equals.
Inverts the result of the `__eq__()` method
Parameters
----------
other : Union[GPSTime, datetime.datetime]
The object to compare. Datatimes will be converted to `GPSTime`
Returns
-------
bool
True if the current object is not the same time as its comparison
Raises
------
TypeError
If an invalid type
"""
return not (self.__eq__(other))
def __hash__(self):
"""Make GPSTime hashable."""
return hash(str(self.week_number) + str(self.seconds) + str(self.femtoseconds))
def __repr__(self) -> str:
"""Representation of the object.
Returns
-------
str
The representation of the object
"""
return "GPSTime(week_number={}, time_of_week={})".format(
self.week_number, self.time_of_week
)
```
| github_jupyter |
```
#import the necessary modules
%matplotlib inline
import numpy as np
import matplotlib.pylab as plt
import pandas as pd
import scipy
import sklearn
import itertools
from itertools import cycle
import os.path as op
import timeit
import json
import math
# These imports are for creating the percent occupancy heat map
import plotly
import plotly.plotly as py
import plotly.graph_objs as go
import multiprocessing as m_proc
m_proc.cpu_count()
# Import MDAnalysis
import MDAnalysis as mda
import statsmodels as stats
from MDAnalysis.analysis import polymer, distances, rdf
import matplotlib.font_manager as font_manager
```
### Calculate radius of gyration and persistence length of PLGA/water simulations
```
def seg_org(poly_atoms, n_monomers):
pmecl = []
count = 0
for i in range(n_monomers):
fgrp = []
count += 1
n_atm = len(poly_atoms.select_atoms("resid "+str(count)))
if count == 1:
#print(poly_atoms.select_atoms("resid "+str(count)+" and name O13"))
pmecl.append(poly_atoms.select_atoms("resid "+str(count)+" and (name O13 O9)"))
#print(pmecl)
elif count == 2:
#print(poly_atoms.select_atoms("name O5 and resid "+str(count)))
pmecl.append(poly_atoms.select_atoms("(resid "+str(count)+" and name O5) or (resid "+str(count-1)+" and name O9)"))
if count != 2 and count != 1:
if count == n_monomers:
pmecl.append(poly_atoms.select_atoms("(resid "+str(count)+" and name O1) or (resid "+str(count-1)+" and name O5)"))
elif count != n_monomers:
#print(count)
pmecl.append(poly_atoms.select_atoms("(resid "+str(count)+" and name O5) or (resid "+str(count-1)+" and name O5)"))
return pmecl
# write function to average across frames to give ensembled averaged cosine theta values at each N - 1 value
def pers_length(polymer_atoms, n_monomers):
""" This function takes the polymer atoms and number of monomers and outputs the cosine theta values
at each bond length"""
# N : number of monomers
# Initialize a zeros matrix of 3 rows, N - 1 columns to store cosine theta values at each frame
# row 1: Bond Length x values
# row 2:
# row 3:
vec_poly = np.zeros(shape=(3,n_monomers-1), dtype=float)
# Inititalize a evenly spaced values with length of N - 1
len_vec = np.arange(n_monomers-1)
# Add 1 to all values to begin sequence from 1, instead of 0
len_vec += 1
# Store bond length values on the vec_poly matrix
vec_poly[1,:] = len_vec
# Initialize a counter
count = 0
# Initialize a vector to store each cosine theta value, dtype is object because each polymer image is a sample set
sv_ply = np.zeros(shape=(n_monomers-1), dtype=object)
for i in range(n_monomers):
count += 1
#
ds_cor = np.zeros(shape=(n_monomers-count))
for j in range(n_monomers - count):
jh = np.arange(n_monomers - count)
jh += count+1
n6_mon1 = polymer_atoms.select_atoms("resid "+str(count))
n6_mon2 = polymer_atoms.select_atoms("resid "+str(jh[j]))
if j == 0:
v1 = n6_mon1.center_of_mass() - n6_mon2.center_of_mass()
v1_norm = v1/(np.linalg.norm(v1))
ds_cor[j] = v1_norm.dot(v1_norm)
elif j != 0:
v2 = n6_mon1.center_of_mass() - n6_mon2.center_of_mass()
v2_norm = v2/(np.linalg.norm(v2))
ds_cor[j] = np.dot(v1_norm, v2_norm)
sv_ply[i] = ds_cor
cor_avg = []
for j in range(n_monomers-1):
lss = []
for i in sv_ply.flat:
try:
lss.append(i[j])
except IndexError:
pass
cor_avg.append(np.mean(lss))
nm = np.array(cor_avg)
ang_vg = []
for i in nm.flat:
if i >= float(0.99):
ang_vg.append(0)
elif i <= float(0.99):
ang_vg.append(math.degrees(math.acos(i)))
vec_poly[0,:] = nm
vec_poly[2,:] = np.array(ang_vg)
return vec_poly
def get_rg_pers_poly(polymer_atoms, universe, start, end, ln_bnd):
"""This function will calculate the average radius of gyration and persistence length of a polymer within a
trajectory block."""
n_monomers = len(np.unique(polymer_atoms.resids))
rg_ens = np.zeros(shape=(1,end-start))
corr_v = np.zeros(shape=(n_monomers-1,end-start))
#ln_corr = np.zeros(shape=(n_monomers-1, end-start))
angle_v = np.zeros(shape=(n_monomers-1,end-start))
v_poly = np.zeros(shape=(4,n_monomers-1))
count_rg = 0
universe.trajectory[start]
for ts in universe.trajectory[start:end]:
p_mat = pers_length(polymer_atoms, n_monomers)
#print(p_mat[0])
corr_v[:,count_rg] = p_mat[0]
#ln_corr[:,count_rg] = np.log(p_mat[0,:])
angle_v[:,count_rg] = p_mat[2]
rg_ens[0,count_rg] = polymer_atoms.radius_of_gyration()
count_rg += 1
universe.trajectory[end]
#Added the calculation of the std dev for the correlation values
v_poly[3,:] = p_mat[1]
for i in range(n_monomers-1):
v_poly[0,i] = np.mean(corr_v[i,:])
#Added the calculation of the std dev for the correlation values
v_poly[1,i] = np.std(corr_v[i,:])
#v_poly[2,i] = np.mean(ln_corr[i,:])
#v_poly[3,i] = np.std(ln_corr[i,:])
v_poly[2,i] = np.mean(angle_v[i,:])
avg_rg = np.mean(rg_ens)
#def expfunc(x, a):
# return np.exp(-x*ln_bnd/2*a)
# Starting with x = 0 for fits
#tr_n6 = np.arange(n_monomers-1)
#tr_n6 += 1
#pers_lp = scipy.optimize.curve_fit(expfunc, tr_n6, v_poly[0,:])[0][0]
return rg_ens, v_poly, corr_v, avg_rg
# I want a list of total fraction of contacts where length is determined by no. of blocks and a dictionary
# of contact groups as keys and list of fractional contacts as values(length of list will be no. of blocks)
def bavg_pers_cnt(no_of_blks, polymer_atoms, universe, len_bnd, begin, final):
n_size = (final - begin)/no_of_blks
ot_dab = {}
universe.trajectory[begin]
sf_lbl = ["Avg Radius of gyration", "Avg persistence length"]
blk_nparr = np.zeros(shape=(len(sf_lbl),no_of_blks))
count = 0
for i in range(no_of_blks):
start = universe.trajectory.frame
print(start)
end = int(start + n_size)
print(end)
pp_rgens, cor_pp, per_pp, rg_avgpp = get_rg_pers_poly(polymer_atoms, universe, start, end, len_bnd)
blk_nparr[0,count] = rg_avgpp
blk_nparr[1,count] = per_pp
universe.trajectory[end]
count += 1
# Save fractional contacts for each AA group type, each element in the value array corresponds to a block
# calculated value
for i in range(len(sf_lbl)):
ot_dab[sf_lbl[i]] = blk_nparr[i,:]
return ot_dab
```
### Water only systems
## N = 6 water
```
# For the right Rg calculation using MD Analysis, use trajactory without pbc
n6_plga_wat = mda.Universe("n6_plga_50ns/0_xa_soln/confout.gro", "n6_plga_50ns/0_xa_soln/nopbc_n6wat.xtc")
len(n6_plga_wat.trajectory)
#Select the polymer heavy atoms
poly_n6 = n6_plga_wat.select_atoms("resname sPLG PLG tPLG and not type H")
com_bond = np.zeros(shape=(1,5000))
count = 0
for ts in n6_plga_wat.trajectory[0:5000]:
n6_mon1 = n6_plga_wat.select_atoms("resid 1")
n6_mon2 = n6_plga_wat.select_atoms("resid 2")
oo_len = mda.analysis.distances.distance_array(n6_mon1.center_of_mass(), n6_mon2.center_of_mass(),
box=n6_plga_wat.trajectory.ts.dimensions)
com_bond[0, count] = oo_len
count += 1
com_bond
lb_avg = np.mean(com_bond)
lb_avg
np.std(com_bond)
plt.scatter(np.arange(5000),com_bond)
#plt.scatter(np.arange(5000),lbond,c="y")
lb_avg
n6_blks = bavg_pers_cnt(5, poly_n6, n6_plga_wat, lb_avg, 0 , 5000)
# Fully extended length of 6 monomer PLGA is 3.838 nm
n6_blks["Avg persistence length"]
n6_blks["Avg Radius of gyration"]
np.mean(n6_blks["Avg persistence length"])
np.std(n6_blks["Avg persistence length"])
n6_rgens, cor_n6, N6ens_cos, rg_avgn6 = get_rg_pers_poly(poly_n6, n6_plga_wat, 0, 5000, lb_avg)
cor_n6
cor_n6[0]
cor_n6[1]
N6ens_cos
N6ens_cos[0]
## Bootstrapping attempt
n_iter = 3
n_size = int(len(N6ens_cos[1])*0.5)
stats_n6 = []
for i in range(n_iter):
#prep train and tests values
train_n6 = sklearn.utils.resample(N6ens_cos[2], n_samples=n_size)
test_n6 = np.array([x for x in N6ens_cos[2] if x.tolist() not in train.tolist()])
lb_avg = np.mean(com_bond)
#lb_avg /= 10
lb_avg
# x values
blen = np.arange(5, dtype=float)
blen *= lb_avg
#nt_tt[0] = 0
blen
# Sample standard deviation of Bond Length x values
np.sqrt((np.sum(blen**2) - ((np.sum(blen)**2)/5))/4)
# population standard deviation of Bond Length x values
np.std(blen)
# Sample variance of Bond Length x values
n6svar = (np.sum(blen**2) - ((np.sum(blen)**2)/5))/4
n6svar
# numpy calculation of population variance
np.var(blen)
# y values
n6lc = np.log(cor_n6[0])
n6lc
mk = cor_n6[1]/cor_n6[0]
mk
plt.figure(figsize=(7,7))
plt.errorbar(blen, np.log(cor_n6[0]), yerr=mk, color='b', linestyle="None",marker='o',
capsize=5, capthick=1, ecolor='black')
# From fitting all points, I get best fit
from sklearn.linear_model import LinearRegression
model_n6 = LinearRegression(fit_intercept=False)
model_n6.fit(blen.reshape(-1,1), n6lc)
# Slope here is in nanometers
print("Model slope: ", model_n6.coef_)
print("Model intercept:", model_n6.intercept_)
gg = model_n6.predict(blen.reshape(-1,1))
gg
print("Mean Std Error:", sklearn.metrics.mean_squared_error(n6lc, gg))
print("R2 score:", sklearn.metrics.r2_score(n6lc, gg))
# Residuals between the true y data and model y data
resid_n6 = n6lc - gg
resid_n6
# How to calculate mean squared error
np.sum(resid_n6**2)/len(resid_n6)
# Standard error: Square root of the mean squared error
np.sqrt(np.sum(resid_n6**2)/len(resid_n6))
# population variance of the residuals
var_n6p = np.var(resid_n6)
var_n6p
# Sample variance: https://www.statisticshowto.datasciencecentral.com/
# probability-and-statistics/descriptive-statistics/sample-variance/
# sample variance of the residuals
ntres = resid_n6
ntres -= np.mean(resid_n6)
nhres = ntres**2
np.sum(nhres)/4
# How to calculate Sum((Xi - avg(X))^2): X values are the bond length values
nt_tt = np.arange(5, dtype=float)
nt_tt *= lb_avg
nt_tt -= np.mean(nt_tt)
nhui = nt_tt**2
np.sum(nhui)
# How to calculate Sum((Xi - avg(X))^2), alternate version, Bond Length X values
nt_tt = np.arange(5, dtype=float)
nt_tt *= lb_avg
np.sum(nt_tt**2) - ((np.sum(nt_tt)**2)/5)
# t-value with 95 % confidence intervals
scipy.stats.t.ppf(0.975, 4)
# How to calculate 95% confidence interval for the slope
flc_n6 = scipy.stats.t.ppf(0.975, 4)*np.sqrt((np.sum(resid_n6**2)/len(resid_n6))/(np.sum(nhui)))
flc_n6
pers_n6 =-1/model_n6.coef_
pers_n6
# Pers length error: error propagation from uncertainty in slope
flc_n6/((model_n6.coef_)**2)
blen
plt.figure(figsize=(7,7))
plt.errorbar(blen, np.log(cor_n6[0]), yerr=mk, color='b', linestyle="None",marker='o',
capsize=5, capthick=1, ecolor='black')
plt.plot(blen, gg, color='b')
plt.title(r'Ensemble Averaged ln(Cosine $\theta$)', fontsize=15, y=1.01)
plt.xlabel(r'Bond Length', fontsize=15)
plt.ylabel(r'ln$\left< Cos(\theta)\right >$', fontsize=15)
#plt.ylim(-1.9,0)
font = font_manager.FontProperties(family='Arial', style='normal', size='14')
plt.legend([r'$N_{PLGA}$ = 6: $L_{p}$ = 18.7 $\AA$ ± 3.5 $\AA$'], loc=3, frameon=0, fontsize=14, prop=font)
plt.tick_params(labelsize=14)
plt.text(5,-0.15,r'R$^{2}$ = 0.98', fontsize=15, color='blue')
lb_avg
exfun(nt, pers_n6)
def exfun(x, a):
return np.exp(-x*lb_avg/a)
nt = np.arange(5)
#nt += 1
plt.scatter(nt, cor_n6[0])
# a-axis is the number of bond lengths away
plt.plot(nt,exfun(nt, pers_n6))
plt.ylim(0,1.2)
```
## N = 8 water
```
n8_plga_wat = mda.Universe("n8_plga_50ns/0_xa_soln/confout.gro", "n8_plga_50ns/0_xa_soln/nopbc_traj.xtc")
n8_plga_wat
#Select the polymer heavy atoms
poly_n8 = n8_plga_wat.select_atoms("resname sPLG PLG tPLG and not type H")
len(n8_plga_wat.trajectory)
com_bond_n8 = np.zeros(shape=(1,5000))
count = 0
for ts in n8_plga_wat.trajectory[0:5000]:
n8_mon1 = n8_plga_wat.select_atoms("resid 1")
n8_mon2 = n8_plga_wat.select_atoms("resid 2")
oo_len = mda.analysis.distances.distance_array(n8_mon1.center_of_mass(), n8_mon2.center_of_mass(),
box=n8_plga_wat.trajectory.ts.dimensions)
com_bond_n8[0, count] = oo_len
count += 1
lb_avg_n8 = np.mean(com_bond_n8)
lb_avg_n8
plt.scatter(np.arange(5000),com_bond_n8)
plt.scatter(np.arange(5000),com_bond)
lb_avg_n8
lb_avg
n8_blks = bavg_pers_cnt(5, poly_n8, n8_plga_wat, lb_avg, 0 , 5000)
n8_blks["Avg persistence length"]
n8_blks["Avg Radius of gyration"]
n8_rgens, cor_n8_mat, N8ens_cos, rg_avgn8 = get_rg_pers_poly(poly_n8, n8_plga_wat, 0, 5000, lb_avg)
cor_n8_mat
cor_n8_mat[0]
cor_n8_mat[1]
# error prop into natural log values
mk_n8 = cor_n8_mat[1]/cor_n8_mat[0]
#mk_n8 /= 2
mk_n8
cor_n8_mat[1]/cor_n8_mat[0]
# For some reason, fit does not work if lb_avg is in angstroms
lb_avg = np.mean(com_bond)
#lb_avg /= 10
lb_avg
# x values
nt_ttn8 = np.arange(7, dtype=float)
nt_ttn8 *= lb_avg
#nt_ttn8[0] = 0
nt_ttn8
# Sample standard deviation of Bond Length x values
np.sqrt((np.sum(nt_ttn8**2) - ((np.sum(nt_ttn8)**2)/7))/6)
# Sample variance of Bond Length x values
n8svar = (np.sum(nt_ttn8**2) - ((np.sum(nt_ttn8)**2)/7))/6
n8svar
# y values
n8lc = np.log(cor_n8_mat[0])
n8lc
plt.figure(figsize=(7,7))
plt.errorbar(blen, np.log(cor_n6[0]), yerr=mk, color='b', linestyle="None",marker='o',
capsize=5, capthick=1, ecolor='black')
from sklearn.linear_model import LinearRegression
model_n8 = LinearRegression(fit_intercept=False)
model_n8.fit(nt_ttn8[:5].reshape(-1,1), n8lc[:5])
# Slope here is in nanometers
print("Model slope: ", model_n8.coef_)
print("Model intercept:", model_n8.intercept_)
gg_n8 = model_n8.predict(nt_ttn8.reshape(-1,1))
gg_n8
print("Mean Std Error:", sklearn.metrics.mean_squared_error(n8lc[:5], gg_n8[:5]))
print("R2 score:", sklearn.metrics.r2_score(n8lc[:5], gg_n8[:5]))
# Residuals between the true y data and model y data
resid_n8 = n8lc[:5] - gg_n8[:5]
resid_n8
# How to calculate mean squared error
np.sum(resid_n8**2)/len(resid_n8)
# Standard error: Square root of the mean squared error
np.sqrt(np.sum(resid_n8**2)/len(resid_n8))
# sample variance of the residuals
ntres_n8 = resid_n8
ntres_n8 -= np.mean(resid_n8)
nhres_n8 = ntres_n8**2
np.sum(nhres_n8)/4
# How to calculate Sum((Xi - avg(X))^2): X values are the bond length values
nt_n8 = np.arange(5, dtype=float)
nt_n8 *= lb_avg
nt_n8 -= np.mean(nt_n8)
nhui = nt_n8**2
np.sum(nhui)
# How to calculate 95% confidence interval for the slope
flc_n8 = scipy.stats.t.ppf(0.975, 6)*np.sqrt((np.sum(resid_n8**2)/len(resid_n8))/(np.sum(nhui)))
flc_n8
pers_n8 =-1/model_n8.coef_
pers_n8
# Pers length error: error propagation from uncertainty in slope
flc_n8/((model_n8.coef_)**2)
nt_ttn8
# ln(cos theta) error: error propagation from uncertainty in cos theta values
mk_n8 = cor_n8_mat[1]/cor_n8_mat[0]
mk_n8
plt.figure(figsize=(7,7))
plt.errorbar(nt_ttn8, np.log(cor_n8_mat[0]), yerr=mk_n8, color='b', linestyle="None",marker='o',
capsize=5, capthick=1, ecolor='black')
plt.plot(nt_ttn8, gg_n8, color='b')
plt.title(r'Ensemble Averaged ln(Cosine $\theta$)', fontsize=15, y=1.01)
plt.xlabel(r'Bond Length', fontsize=15)
plt.ylabel(r'ln$\left< Cos(\theta)\right >$', fontsize=15)
#plt.ylim(-1.9,0)
font = font_manager.FontProperties(family='Arial', style='normal', size='14')
plt.legend([r'$N_{PLGA}$ = 8: $L_{p}$ = 17.8 $\AA$ ± 1.5 $\AA$'], loc=3, frameon=0, fontsize=14, prop=font)
plt.tick_params(labelsize=14)
plt.text(5,-0.15,r'R$^{2}$ = 0.99', fontsize=15, color='blue')
def exfun(x, a):
return np.exp(-x*lb_avg_n8/a)
nt = np.arange(7)
plt.scatter(nt, cor_n8_mat[0,:])
plt.plot(nt,exfun(nt, pers_n8))
```
## N = 10 water
```
n10_plga_wat = mda.Universe("n10_plga_50ns/0_xa_soln/N10_sys.gro", "n10_plga_50ns/0_xa_soln/nopbc_10wat.xtc")
n10_plga_wat
len(n10_plga_wat.trajectory)
poly_n10 = n10_plga_wat.select_atoms("resname sPLG PLG tPLG and not type H")
com_bond_n10 = np.zeros(shape=(1,5000))
count = 0
for ts in n10_plga_wat.trajectory[0:5000]:
n10_mon1 = n10_plga_wat.select_atoms("resid 1")
n10_mon2 = n10_plga_wat.select_atoms("resid 2")
oo_len = mda.analysis.distances.distance_array(n10_mon1.center_of_mass(), n10_mon2.center_of_mass(),
box=n10_plga_wat.trajectory.ts.dimensions)
com_bond_n10[0, count] = oo_len
count += 1
lb_avg_n10 = np.mean(com_bond_n10)
lb_avg_n10
plt.scatter(np.arange(5000),com_bond)
plt.scatter(np.arange(5000),com_bond_n8)
plt.scatter(np.arange(5000),com_bond_n10)
lb_avg
n10_blks = bavg_pers_cnt(5, poly_n10, n10_plga_wat, lb_avg_n10, 0 , 5000)
# Extended Length of 10 monomer length plga: 6.436 nm
n10_blks["Avg persistence length"]
n10_blks["Avg Radius of gyration"]
np.mean(n10_blks["Avg Radius of gyration"])
np.std(n10_blks["Avg Radius of gyration"])
np.mean(n10_blks["Avg persistence length"])
np.std(n10_blks["Avg persistence length"])
n10_rgens, cor_n10_mat, N10ens_cos, rg_avgn10 = get_rg_pers_poly(poly_n10, n10_plga_wat, 0, 5000, lb_avg)
cor_n10_mat
cor_n10_mat[0]
cor_n10_mat[1]
mk_n10 = cor_n10_mat[1]/cor_n10_mat[0]
#mk_n10 /= 2
mk_n10
cor_n10_mat[1]/cor_n10_mat[0]
lb_avg = np.mean(com_bond)
#lb_avg /= 10
lb_avg
# x values
nt_ttn10 = np.arange(9, dtype=float)
nt_ttn10 *= lb_avg
#nt_ttn10[0] = 0
nt_ttn10
# Sample standard deviation of Bond Length x values
np.sqrt((np.sum(nt_ttn10**2) - ((np.sum(nt_ttn10)**2)/9))/8)
# Sample variance of Bond Length x values
n10svar = (np.sum(nt_ttn10**2) - ((np.sum(nt_ttn10)**2)/9))/8
n10svar
# y values
n10lc = np.log(cor_n10_mat[0])
n10lc
plt.figure(figsize=(7,7))
plt.errorbar(nt_ttn10, np.log(cor_n10_mat[0]), yerr=mk_n10, color='b', linestyle="None",marker='o',
capsize=5, capthick=1, ecolor='black')
nt_ttn10[:5]
from sklearn.linear_model import LinearRegression
model_n10 = LinearRegression(fit_intercept=False)
model_n10.fit(nt_ttn10[:5].reshape(-1,1), n10lc[:5])
# Slope here is in nanometers
print("Model slope: ", model_n10.coef_)
print("Model intercept:", model_n10.intercept_)
gg_n10 = model_n10.predict(nt_ttn10.reshape(-1,1))
gg_n10
print("Mean Std Error:", sklearn.metrics.mean_squared_error(n10lc[:5], model_n10.predict(nt_ttn10[:5].reshape(-1,1))))
print("R2 score:", sklearn.metrics.r2_score(n10lc[:5], model_n10.predict(nt_ttn10[:5].reshape(-1,1))))
# Residuals between the true y data and model y data
resid_n10 = n10lc[:5] - gg_n10[:5]
resid_n10
# How to calculate mean squared error
np.sum(resid_n10**2)/len(resid_n10)
# Standard error: Square root of the mean squared error
np.sqrt(np.sum(resid_n10**2)/len(resid_n10))
# sample variance of the residuals
ntres_n10 = resid_n10
ntres_n10 -= np.mean(resid_n10)
nhres_n10 = ntres_n10**2
np.sum(nhres_n10)/4
# How to calculate Sum((Xi - avg(X))^2): X values are the bond length values
nt_n10 = np.arange(5, dtype=float)
nt_n10 *= lb_avg
nt_n10 -= np.mean(nt_n10)
nhui = nt_n10**2
np.sum(nhui)
# How to calculate 95% confidence interval for the slope
flc_n10 = scipy.stats.t.ppf(0.975, 4)*np.sqrt((np.sum(resid_n10**2)/len(resid_n10))/(np.sum(nhui)))
flc_n10
pers_n10 =-1/model_n10.coef_
pers_n10
# Pers length error: error propagation from uncertainty in slope
flc_n10/((model_n10.coef_)**2)
mk_n10
nt_ttn10
np.log(cor_n10_mat[0])
plt.figure(figsize=(7,7))
plt.errorbar(nt_ttn10, np.log(cor_n10_mat[0]), yerr=mk_n10, color='b', linestyle="None",marker='o', capsize=5, capthick=1, ecolor='black')
plt.plot(nt_ttn10, gg_n10, color='b')
plt.title(r'Ensemble Averaged ln(Cosine $\theta$)', fontsize=15, y=1.01)
plt.xlabel(r'Bond Length', fontsize=15)
plt.ylabel(r'ln$\left< Cos(\theta)\right >$', fontsize=15)
#plt.ylim(-1.9,0)
font = font_manager.FontProperties(family='Arial', style='normal', size='14')
plt.legend([r'$N_{PLGA}$ = 10: $L_{p}$ = 14.7 $\AA$ ± 3.4 $\AA$'], loc=3, frameon=0, fontsize=14, prop=font)
plt.tick_params(labelsize=14)
plt.text(5,-0.15,r'R$^{2}$ = 0.97', fontsize=15, color='blue')
def exfun(x, a):
return np.exp(-x*lb_avg/a)
nt = np.arange(9)
plt.scatter(nt, cor_n10_mat[0,:])
plt.plot(nt,exfun(nt, pers_n10))
```
## N = 20 water
```
n20_plga_wat = mda.Universe("n20_plga_150ns/0_xa_soln/nowat_n20.gro", "n20_plga_150ns/0_xa_soln/N20_nopbcpp.xtc")
n20_plga_wat
len(n20_plga_wat.trajectory)
poly_n20 = n20_plga_wat.select_atoms("resname sPLG PLG tPLG and not type H")
com_bond_n20 = np.zeros(shape=(1,15000))
count = 0
for ts in n20_plga_wat.trajectory[0:15000]:
n20_mon1 = n20_plga_wat.select_atoms("resid 1")
n20_mon2 = n20_plga_wat.select_atoms("resid 2")
oo_len = mda.analysis.distances.distance_array(n20_mon1.center_of_mass(), n20_mon2.center_of_mass(),
box=n20_plga_wat.trajectory.ts.dimensions)
com_bond_n20[0, count] = oo_len
count += 1
lb_avg_n20 = np.mean(com_bond_n20)
lb_avg_n20
plt.scatter(np.arange(5000),com_bond)
plt.scatter(np.arange(5000),com_bond_n8)
plt.scatter(np.arange(5000),com_bond_n10)
plt.scatter(np.arange(15000),com_bond_n20)
n20_blks = bavg_pers_cnt(5, poly_n20, n20_plga_wat, lb_avg, 0 , 15000)
n20_blks["Avg persistence length"]
n20_blks["Avg Radius of gyration"]
np.mean(n20_blks["Avg Radius of gyration"])
np.std(n20_blks["Avg Radius of gyration"])
np.mean(n20_blks["Avg persistence length"])
np.std(n20_blks["Avg persistence length"])
n20_rgens, cor_n20_mat, N20ens_cos, rg_avgn20 = get_rg_pers_poly(poly_n20, n20_plga_wat, 0, 15000, lb_avg)
cor_n20_mat
cor_n20_mat[0]
cor_n20_mat[1]
lb_avg = np.mean(com_bond)
#lb_avg /= 10
lb_avg
# x values
nt_ttn20 = np.arange(19, dtype=float)
nt_ttn20 *= lb_avg
#nt_ttn20[0] = 0
nt_ttn20
# Sample standard deviation of Bond Length x values
np.sqrt((np.sum(nt_ttn20**2) - ((np.sum(nt_ttn20)**2)/19))/18)
# Sample variance of Bond Length x values
n20svar = (np.sum(nt_ttn20**2) - ((np.sum(nt_ttn20)**2)/19))/18
n20svar
# y values
n20lc = np.log(cor_n20_mat[0])
n20lc
mk_n20 = cor_n20_mat[1]/cor_n20_mat[0]
mk_n20
plt.figure(figsize=(7,7))
plt.errorbar(nt_ttn20, np.log(cor_n20_mat[0]), yerr=mk_n20, color='b', linestyle="None",marker='o',
capsize=5, capthick=1, ecolor='black')
nt_ttn20[:8]
from sklearn.linear_model import LinearRegression
model_n20 = LinearRegression(fit_intercept=False)
model_n20.fit(nt_ttn20[:8].reshape(-1,1), n20lc[:8])
# Slope here is in nanometers
print("Model slope: ", model_n20.coef_)
print("Model intercept:", model_n20.intercept_)
gg_n20 = model_n20.predict(nt_ttn20.reshape(-1,1))
gg_n20
print("Mean Std Error:", sklearn.metrics.mean_squared_error(n20lc[:8], model_n20.predict(nt_ttn20[:8].reshape(-1,1))))
print("R2 score:", sklearn.metrics.r2_score(n20lc[:8], model_n20.predict(nt_ttn20[:8].reshape(-1,1))))
# Residuals between the true y data and model y data
resid_n20 = n20lc[:8] - gg_n20[:8]
resid_n20
# How to calculate mean squared error
np.sum(resid_n20**2)/len(resid_n20)
# Standard error: Square root of the mean squared error
np.sqrt(np.sum(resid_n20**2)/len(resid_n20))
# sample variance of the residuals
ntres_n20 = resid_n20
ntres_n20 -= np.mean(resid_n20)
nhres_n20 = ntres_n20**2
np.sum(nhres_n20)/7
# How to calculate Sum((Xi - avg(X))^2): X values are the bond length values
nt_n20 = np.arange(8, dtype=float)
nt_n20 *= lb_avg
nt_n20 -= np.mean(nt_n20)
nhui = nt_n20**2
np.sum(nhui)
# How to calculate 95% confidence interval for the slope
flc_n20 = scipy.stats.t.ppf(0.975, 4)*np.sqrt((np.sum(resid_n20**2)/len(resid_n20))/(np.sum(nhui)))
flc_n20
pers_n20 =-1/model_n20.coef_
pers_n20
# Pers length error: error propagation from uncertainty in slope
flc_n20/((model_n20.coef_)**2)
mk_n20 = cor_n20_mat[1]/cor_n20_mat[0]
plt.figure(figsize=(7,7))
plt.errorbar(nt_ttn20, np.log(cor_n20_mat[0]), yerr=mk_n20, color='b', linestyle="None",marker='o',capsize=5, capthick=1, ecolor='black')
plt.plot(nt_ttn20, gg_n20, color='b')
plt.title(r'Ensemble Averaged ln(Cosine $\theta$)', fontsize=15, y=1.01)
plt.xlabel(r'Bond Length', fontsize=15)
plt.ylabel(r'ln$\left< Cos(\theta)\right >$', fontsize=15)
#plt.ylim(-1.9,0)
font = font_manager.FontProperties(family='Arial', style='normal', size='14')
plt.legend([r'$N_{PLGA}$ = 20: $L_{p}$ = 19.1 $\AA$ ± 1.0 $\AA$'], loc=3, frameon=0, fontsize=14, prop=font)
plt.tick_params(labelsize=14)
plt.text(5,-0.15,r'R$^{2}$ = 0.99', fontsize=15, color='blue')
def exfun(x, a):
return np.exp(-x*lb_avg/a)
nt = np.arange(19)
plt.scatter(nt, cor_n20_mat[0,:])
plt.plot(nt,exfun(nt, pers_n10))
```
## N= 30 water
```
n30_plga_wat = mda.Universe("n30_plga_150ns/0_xa_soln/0xa_n30sys.gro", "n30_plga_150ns/0_xa_soln/N30_nopbcpp_0xa.xtc")
n30_plga_wat
len(n30_plga_wat.trajectory)
poly_n30 = n30_plga_wat.select_atoms("resname sPLG PLG tPLG and not type H")
com_bond_n30 = np.zeros(shape=(1,15000))
count = 0
for ts in n30_plga_wat.trajectory[0:15000]:
n30_mon1 = n30_plga_wat.select_atoms("resid 1")
n30_mon2 = n30_plga_wat.select_atoms("resid 2")
oo_len = mda.analysis.distances.distance_array(n30_mon1.center_of_mass(), n30_mon2.center_of_mass(),
box=n30_plga_wat.trajectory.ts.dimensions)
com_bond_n30[0, count] = oo_len
count += 1
lb_avg_n30 = np.mean(com_bond_n30)
lb_avg_n30
lb_avg
n30_blks = bavg_pers_cnt(5, poly_n30, n30_plga_wat, lb_avg, 0 , 15000)
n30_blks["Avg persistence length"]
n30_blks["Avg Radius of gyration"]
np.mean(n30_blks["Avg persistence length"])
np.std(n30_blks["Avg persistence length"])
n30_rgens, cor_n30_mat, N30ens_cos, rg_avgn30 = get_rg_pers_poly(poly_n30, n30_plga_wat, 0, 15000, lb_avg)
cor_n30_mat
cor_n30_mat[0]
# error propagation into natural log values
mk_n30 = cor_n30_mat[1]/cor_n30_mat[0]
# For some reason, fit does not work if lb_avg is in angstroms
lb_avg = np.mean(com_bond)
#lb_avg /= 10
lb_avg
# x values
nt_ttn30 = np.arange(29, dtype=float)
nt_ttn30 *= lb_avg
#nt_ttn10[0] = 0
nt_ttn30
nt_ttn30[:20]
# Sample standard deviation of Bond Length x values
np.sqrt((np.sum(nt_ttn30**2) - ((np.sum(nt_ttn30)**2)/29))/28)
# Sample variance of Bond Length x values
n30svar = (np.sum(nt_ttn30**2) - ((np.sum(nt_ttn30)**2)/29))/28
n30svar
# y values
n30lc = np.log(cor_n30_mat[0])
n30lc
mk_n30
plt.figure(figsize=(7,7))
plt.errorbar(nt_ttn30, np.log(cor_n30_mat[0]), yerr=mk_n30, color='b', linestyle="None",marker='o',
capsize=5, capthick=1, ecolor='black')
n30lc[:6]
nt_ttn30[:6]
from sklearn.linear_model import LinearRegression
model_n30 = LinearRegression(fit_intercept=False)
model_n30.fit(nt_ttn30[:6].reshape(-1,1), n30lc[:6])
# Slope here is in angstroms
print("Model slope: ", model_n30.coef_)
print("Model intercept:", model_n30.intercept_)
gg_n30 = model_n30.predict(nt_ttn30.reshape(-1,1))
gg_n30
print("Mean Std Error:", sklearn.metrics.mean_squared_error(n30lc[:6], model_n30.predict(nt_ttn30[:6].reshape(-1,1))))
print("R2 score:", sklearn.metrics.r2_score(n30lc[:6], model_n30.predict(nt_ttn30[:6].reshape(-1,1))))
# Residuals between the true y data and model y data
resid_n30 = n30lc[:6] - gg_n30[:6]
resid_n30
# How to calculate mean squared error
np.sum(resid_n30**2)/len(resid_n30)
# Standard error: Square root of the mean squared error
np.sqrt(np.sum(resid_n30**2)/len(resid_n30))
# sample variance of the residuals
ntres_n30 = resid_n30
ntres_n30 -= np.mean(resid_n30)
nhres_n30 = ntres_n30**2
# divide by n-1 samples
np.sum(nhres_n30)/5
# How to calculate Sum((Xi - avg(X))^2): X values are the bond length values
nt_n30 = np.arange(6, dtype=float)
nt_n30 *= lb_avg
nt_n30 -= np.mean(nt_n30)
nhui = nt_n30**2
np.sum(nhui)
# How to calculate 95% confidence interval for the slope
flc_n30 = scipy.stats.t.ppf(0.975, 4)*np.sqrt((np.sum(resid_n30**2)/len(resid_n30))/(np.sum(nhui)))
flc_n30
pers_n30 =-1/model_n30.coef_
pers_n30
# Pers length error: error propagation from uncertainty in slope
flc_n30/((model_n30.coef_)**2)
mk_n30 = cor_n30_mat[1]/cor_n30_mat[0]
def slope_plen(x,m):
return m*x
ttsam = np.arange(12, dtype=float)
ttsam *= lb_avg
#nt_ttn10[0] = 0
ttsam
model_n10.coef_[0]
slope_plen(ttsam, model_n10.coef_[0])
blen
np.log(cor_n6[0])
mk
plt.figure(figsize=(7,7))
plt.errorbar(nt_ttn30, np.log(cor_n30_mat[0]), yerr=mk_n30, color='b', linestyle="None",marker='h',
capsize=5, capthick=1, ecolor='black')
plt.plot(ttsam, slope_plen(ttsam, model_n30.coef_[0]), color='b')
plt.errorbar(nt_ttn20, np.log(cor_n20_mat[0]), yerr=mk_n20, color='g', linestyle="None"
,marker='*',capsize=5, capthick=1, ecolor='black')
plt.plot(ttsam, slope_plen(ttsam, model_n20.coef_[0]), color='g')
plt.errorbar(nt_ttn10, np.log(cor_n10_mat[0]), yerr=mk_n10, color='r', linestyle="None"
,marker='v', capsize=5, capthick=1, ecolor='black')
plt.plot(ttsam, slope_plen(ttsam, model_n10.coef_[0]), color='r')
plt.errorbar(nt_ttn8, np.log(cor_n8_mat[0]), yerr=mk_n8, color='k', linestyle="None",marker='s'
,capsize=5, capthick=1, ecolor='black')
plt.plot(ttsam, slope_plen(ttsam, model_n8.coef_[0]), color='k')
plt.errorbar(blen, np.log(cor_n6[0]), yerr=mk, color='c', linestyle="None",marker='x'
,capsize=5, capthick=1, ecolor='black')
plt.plot(ttsam, slope_plen(ttsam, model_n6.coef_[0]), color='m')
plt.title(r'Ensemble Averaged ln(Cosine $\theta$)', fontsize=15, y=1.01)
plt.xlabel(r'Bond Length', fontsize=15)
plt.ylabel(r'ln$\left< Cos(\theta)\right >$', fontsize=15)
plt.ylim(-6, 1)
plt.xlim(0,60)
font = font_manager.FontProperties(family='Arial', style='normal', size='14')
#plt.legend([r'$N_{PLGA}$ = 30: $L_{p}$ = 18.5 $\AA$ ± 0.15 $\AA$, R$^{2}$ = 0.99',
#r'$N_{PLGA}$ = 20: $L_{p}$ = 19.1 $\AA$ ± 0.27 $\AA$, R$^{2}$ = 0.99'],
# loc=3, frameon=0, fontsize=14, prop=font)
#plt.legend([r'$N_{PLGA}$ = 20: $L_{p}$ = 19.1 $\AA$ ± 0.27 $\AA$, R$^{2}$ = 0.99'],
#loc=3, frameon=0, fontsize=14, prop=font)
plt.tick_params(labelsize=14)
plt.text(1, -5.9,r'$N_{PLGA}$ = 30: $L_{p}$ = 18.5 $\AA$ ± 1.6 $\AA$, R$^{2}$ = 0.99', fontsize=14, color='blue')
plt.text(1,-5.6,r'$N_{PLGA}$ = 20: $L_{p}$ = 19.1 $\AA$ ± 1.0 $\AA$, R$^{2}$ = 0.99', fontsize=14, color='green')
plt.text(1,-5.3,r'$N_{PLGA}$ = 10: $L_{p}$ = 14.7 $\AA$ ± 3.4 $\AA$, R$^{2}$ = 0.97', fontsize=14, color='red')
plt.text(1,-5.0,r'$N_{PLGA}$ = 8: $L_{p}$ = 17.8 $\AA$ ± 1.5, R$^{2}$ = 0.99', fontsize=14, color='black')
plt.text(1,-4.7,r'$N_{PLGA}$ = 6: $L_{p}$ = 18.7 $\AA$ ± 3.5 $\AA$, R$^{2}$ = 0.98', fontsize=14, color='magenta')
#plt.text(2.5,0,r'R$^{2}$ = 0.96', fontsize=15, color='red')
cor_n8_mat[0]
nt_ttn8
def exfun(x, a):
return np.exp(-x*lb_avg/a)
nt = np.arange(29)
plt.scatter(nt, cor_n30_mat[0,:])
plt.plot(nt,exfun(nt, pers_n30))
```
### Acetone systems only
## N = 6 acetone
```
n6_plga_ace = mda.Universe("n6_plga_50ns/1_xa_soln/confout.gro", "n6_plga_50ns/1_xa_soln/nopbc_n6ace.xtc")
n6_plga_ace
len(n6_plga_ace.trajectory)
poly_n6_ace = n6_plga_ace.select_atoms("resname sPLG PLG tPLG and not type H")
com_bond_n6ace = np.zeros(shape=(1,5000))
count = 0
for ts in n6_plga_ace.trajectory[0:5000]:
n6_mon1 = n6_plga_ace.select_atoms("resid 1")
n6_mon2 = n6_plga_ace.select_atoms("resid 2")
oo_len = mda.analysis.distances.distance_array(n6_mon1.center_of_mass(), n6_mon2.center_of_mass(),
box=n6_plga_ace.trajectory.ts.dimensions)
com_bond_n6ace[0, count] = oo_len
count += 1
lb_avg_n6ace = np.mean(com_bond_n6ace)
lb_avg_n6ace
plt.scatter(np.arange(5000),com_bond_n6ace)
n6_blks_ace = bavg_pers_cnt(5, poly_n6_ace, n6_plga_ace, lb_avg_n6ace, 0 , 5000)
n6_blks_ace["Avg persistence length"]
n6_blks_ace["Avg Radius of gyration"]
np.mean(n6_blks_ace["Avg persistence length"])
np.std(n6_blks_ace["Avg persistence length"])
np.mean(n6_blks_ace["Avg Radius of gyration"])
np.std(n6_blks_ace["Avg Radius of gyration"])
n6_rgensace, cor_n6ace_mat, N30ens_cos, rg_avgn6ace = get_rg_pers_poly(poly_n6_ace, n6_plga_ace, 0, 5000,
lb_avg_n6ace)
cor_n6ace_mat
cor_n6ace_mat[0]
# x values
ace_ttn6 = np.arange(5, dtype=float)
ace_ttn6 *= lb_avg_n6ace
#nt_ttn20[0] = 0
ace_ttn6
# Sample standard deviation of Bond Length x values
np.sqrt((np.sum(ace_ttn6**2) - ((np.sum(ace_ttn6)**2)/5))/4)
# Sample variance of Bond Length x values
n6sace_var = (np.sum(ace_ttn6**2) - ((np.sum(ace_ttn6)**2)/5))/4
n6sace_var
# y values
n6lc_ace = np.log(cor_n6ace_mat[0])
n6lc_ace
mk_n6ace = cor_n6ace_mat[1]/cor_n6ace_mat[0]
plt.figure(figsize=(7,7))
plt.errorbar(ace_ttn6, np.log(cor_n6ace_mat[0]), yerr=mk_n6ace, color='b', linestyle="None",marker='o',
capsize=5, capthick=1, ecolor='black')
from sklearn.linear_model import LinearRegression
model_n6ace = LinearRegression(fit_intercept=False)
model_n6ace.fit(ace_ttn6.reshape(-1,1), n6lc_ace)
# Slope here is in angstroms
print("Model slope: ", model_n6ace.coef_)
print("Model intercept:", model_n6ace.intercept_)
gg_n6ace = model_n6ace.predict(ace_ttn6.reshape(-1,1))
gg_n6ace
print("Mean Std Error:", sklearn.metrics.mean_squared_error(n6lc_ace, model_n6ace.predict(ace_ttn6.reshape(-1,1))))
print("R2 score:", sklearn.metrics.r2_score(n6lc_ace, model_n6ace.predict(ace_ttn6.reshape(-1,1))))
# Residuals between the true y data and model y data
resid_n6ace = n6lc_ace - gg_n6ace
resid_n6ace
# How to calculate mean squared error
np.sum(resid_n6ace**2)/len(resid_n6ace)
# Standard error: Square root of the mean squared error
np.sqrt(np.sum(resid_n6ace**2)/len(resid_n6ace))
# sample variance of the residuals
ntr_n6ace = resid_n6ace
ntr_n6ace -= np.mean(resid_n6ace)
nhr_n6ace = ntr_n6ace**2
np.sum(nhr_n6ace)/4
# How to calculate Sum((Xi - avg(X))^2): X values are the bond length values
nt_n6ace = np.arange(5, dtype=float)
nt_n6ace *= lb_avg_n6ace
nt_n6ace -= np.mean(nt_n6ace)
nhui = nt_n6ace**2
np.sum(nhui)
# How to calculate 95% confidence interval for the slope
flc_n6ace = scipy.stats.t.ppf(0.975, 4)*np.sqrt((np.sum(resid_n6ace**2)/len(resid_n6ace))/(np.sum(nhui)))
flc_n6ace
pers_n6ace =-1/model_n6ace.coef_
pers_n6ace
# Pers length error: error propagation from uncertainty in slope
flc_n6ace/((model_n6ace.coef_)**2)
def slope_plen(x,m):
return m*x
ttsam = np.arange(5, dtype=float)
ttsam *= lb_avg_n6ace
#nt_ttn10[0] = 0
ttsam
plt.figure(figsize=(7,7))
plt.errorbar(ace_ttn6, np.log(cor_n6ace_mat[0]), yerr=mk_n6ace, color='b', linestyle="None",marker='o',capsize=5, capthick=1, ecolor='black')
plt.plot(ttsam, slope_plen(ttsam, model_n6ace.coef_[0]), color='b')
plt.title(r'Ensemble Averaged ln(Cosine $\theta$) in pure acetone', fontsize=15, y=1.01)
plt.xlabel(r'Bond Length', fontsize=15)
plt.ylabel(r'ln$\left< Cos(\theta)\right >$', fontsize=15)
#plt.ylim(-1.9,0)
font = font_manager.FontProperties(family='Arial', style='normal', size='14')
plt.legend([r'$N_{PLGA}$ = 6: $L_{p}$ = 41.6 $\AA$ ± 5.7 $\AA$'], loc=3, frameon=0, fontsize=14, prop=font)
plt.tick_params(labelsize=14)
plt.text(5,0,r'R$^{2}$ = 0.99', fontsize=15, color='blue')
def exfun(x, a):
return np.exp(-x*lb_avg_n6ace/a)
nt = np.arange(5)
plt.scatter(nt, cor_n6ace_mat[0,:])
plt.plot(nt,exfun(nt, pers_n6ace))
```
## N = 8 acetone
```
n8_plga_ace = mda.Universe("n8_plga_50ns/1_xa_soln/confout.gro", "n8_plga_50ns/1_xa_soln/nopbc_n8ace.xtc")
n8_plga_ace
len(n8_plga_ace.trajectory)
poly_n8_ace = n8_plga_ace.select_atoms("resname sPLG PLG tPLG and not type H")
com_bond_n8ace = np.zeros(shape=(1,5000))
count = 0
for ts in n8_plga_ace.trajectory[0:5000]:
n8_mon1 = n8_plga_ace.select_atoms("resid 1")
n8_mon2 = n8_plga_ace.select_atoms("resid 2")
oo_len = mda.analysis.distances.distance_array(n8_mon1.center_of_mass(), n8_mon2.center_of_mass(),
box=n8_plga_ace.trajectory.ts.dimensions)
com_bond_n8ace[0, count] = oo_len
count += 1
lb_avg_n8ace = np.mean(com_bond_n8ace)
lb_avg_n8ace
plt.scatter(np.arange(5000),com_bond_n6ace)
plt.scatter(np.arange(5000),com_bond_n8ace)
n8_blks_ace = bavg_pers_cnt(5, poly_n8_ace, n8_plga_ace, lb_avg_n6ace, 0 , 5000)
n8_blks_ace["Avg persistence length"]
n8_blks_ace["Avg Radius of gyration"]
np.mean(n8_blks_ace["Avg persistence length"])
np.std(n8_blks_ace["Avg persistence length"])
np.mean(n8_blks_ace["Avg Radius of gyration"])
np.std(n8_blks_ace["Avg Radius of gyration"])
n8_rgens_ace, cor_n8ace_mat, N8aceens_cos, rg_avgn8ace = get_rg_pers_poly(poly_n8_ace, n8_plga_ace, 0, 5000,
lb_avg_n6ace)
cor_n8ace_mat
cor_n8ace_mat[0]
# x values
ace_ttn8 = np.arange(7, dtype=float)
ace_ttn8 *= lb_avg_n6ace
#nt_ttn20[0] = 0
ace_ttn8
# Sample standard deviation of Bond Length x values
np.sqrt((np.sum(ace_ttn8**2) - ((np.sum(ace_ttn8)**2)/7))/6)
# Sample variance of Bond Length x values
n8sace_var = (np.sum(ace_ttn8**2) - ((np.sum(ace_ttn8)**2)/7))/6
n8sace_var
# y values
n8lc_ace = np.log(cor_n8ace_mat[0])
n8lc_ace
mk_n8ace = cor_n8ace_mat[1]/cor_n8ace_mat[0]
plt.figure(figsize=(7,7))
plt.errorbar(ace_ttn8, np.log(cor_n8ace_mat[0]), yerr=mk_n8ace, color='b', linestyle="None",marker='o',
capsize=5, capthick=1, ecolor='black')
ace_ttn8[1:]
from sklearn.linear_model import LinearRegression
model_n8ace = LinearRegression(fit_intercept=False)
model_n8ace.fit(ace_ttn8.reshape(-1,1), n8lc_ace)
# Slope here is in angstroms
print("Model slope: ", model_n8ace.coef_)
print("Model intercept:", model_n8ace.intercept_)
gg_n8ace = model_n8ace.predict(ace_ttn8.reshape(-1,1))
gg_n8ace
print("Mean Std Error:", sklearn.metrics.mean_squared_error(n8lc_ace, model_n8ace.predict(ace_ttn8.reshape(-1,1))))
print("R2 score:", sklearn.metrics.r2_score(n8lc_ace, model_n8ace.predict(ace_ttn8.reshape(-1,1))))
# Residuals between the true y data and model y data
resid_n8ace = n8lc_ace - gg_n8ace
resid_n8ace
# How to calculate mean squared error
np.sum(resid_n8ace**2)/len(resid_n8ace)
# Standard error: Square root of the mean squared error
np.sqrt(np.sum(resid_n8ace**2)/len(resid_n8ace))
# sample variance of the residuals
ntr_n8ace = resid_n8ace
ntr_n8ace -= np.mean(resid_n8ace)
nhr_n8ace = ntr_n8ace**2
np.sum(nhr_n8ace)/6
# How to calculate Sum((Xi - avg(X))^2): X values are the bond length values
nt_n8ace = np.arange(7, dtype=float)
nt_n8ace *= lb_avg_n6ace
nt_n8ace -= np.mean(nt_n8ace)
nhui = nt_n8ace**2
np.sum(nhui)
# How to calculate 95% confidence interval for the slope
flc_n8ace = scipy.stats.t.ppf(0.975, 4)*np.sqrt((np.sum(resid_n8ace**2)/len(resid_n8ace))/(np.sum(nhui)))
flc_n8ace
pers_n8ace =-1/model_n8ace.coef_
pers_n8ace
# Pers length error: error propagation from uncertainty in slope
flc_n8ace/((model_n8ace.coef_)**2)
def slope_plen(x,m):
return m*x
ttsam = np.arange(7, dtype=float)
ttsam *= lb_avg_n6ace
#nt_ttn10[0] = 0
ttsam
plt.figure(figsize=(7,7))
plt.errorbar(ace_ttn8, np.log(cor_n8ace_mat[0]), yerr=mk_n8ace, color='b', linestyle="None",marker='o',capsize=5, capthick=1, ecolor='black')
plt.plot(ttsam, slope_plen(ttsam, model_n8ace.coef_[0]), color='b')
plt.title(r'Ensemble Averaged ln(Cosine $\theta$) in pure acetone', fontsize=15, y=1.01)
plt.xlabel(r'Bond Length', fontsize=15)
plt.ylabel(r'ln$\left< Cos(\theta)\right >$', fontsize=15)
#plt.ylim(-1.9,0)
font = font_manager.FontProperties(family='Arial', style='normal', size='14')
plt.legend([r'$N_{PLGA}$ = 8: $L_{p}$ = 39.3 $\AA$ ± 5.9 $\AA$'], loc=3, frameon=0, fontsize=14, prop=font)
plt.tick_params(labelsize=14)
plt.text(5,0,r'R$^{2}$ = 0.98', fontsize=15, color='blue')
def exfun(x, a):
return np.exp(-x*lb_avg_n6ace/a)
nt = np.arange(7)
plt.scatter(nt, cor_n8ace_mat[0,:])
plt.plot(nt,exfun(nt, pers_n8ace))
```
## N = 10 acetone
```
n10_plga_ace = mda.Universe("n10_plga_50ns/1_xa_soln/confout.gro", "n10_plga_50ns/1_xa_soln/nopbc_n10ace.xtc")
n10_plga_ace
len(n10_plga_ace.trajectory)
poly_n10_ace = n10_plga_ace.select_atoms("resname sPLG PLG tPLG and not type H")
com_bond_n10ace = np.zeros(shape=(1,5000))
count = 0
for ts in n10_plga_ace.trajectory[0:5000]:
n10_mon1 = n10_plga_ace.select_atoms("resid 1")
n10_mon2 = n10_plga_ace.select_atoms("resid 2")
oo_len = mda.analysis.distances.distance_array(n10_mon1.center_of_mass(), n10_mon2.center_of_mass(),
box=n10_plga_ace.trajectory.ts.dimensions)
com_bond_n10ace[0, count] = oo_len
count += 1
lb_avg_n10ace = np.mean(com_bond_n10ace)
lb_avg_n10ace
plt.scatter(np.arange(5000),com_bond_n6ace)
plt.scatter(np.arange(5000),com_bond_n8ace)
plt.scatter(np.arange(5000),com_bond_n10ace)
n10_blks_ace = bavg_pers_cnt(5, poly_n10_ace, n10_plga_ace, lb_avg_n10ace, 0 , 5000)
n10_blks_ace["Avg persistence length"]
n10_blks_ace["Avg Radius of gyration"]
np.mean(n10_blks_ace["Avg persistence length"])
np.std(n10_blks_ace["Avg persistence length"])
np.mean(n10_blks_ace["Avg Radius of gyration"])
np.std(n10_blks_ace["Avg Radius of gyration"])
n10_rgens_ace, cor_n10ace_mat, N10aceens_cos, rg_avgn10ace = get_rg_pers_poly(poly_n10_ace, n10_plga_ace, 0, 5000,
lb_avg_n6ace)
cor_n10ace_mat
cor_n10ace_mat[0]
# x values
ace_ttn10 = np.arange(9, dtype=float)
ace_ttn10 *= lb_avg_n6ace
#nt_ttn20[0] = 0
ace_ttn10
# Sample standard deviation of Bond Length x values
np.sqrt((np.sum(ace_ttn10**2) - ((np.sum(ace_ttn10)**2)/9))/8)
# Sample variance of Bond Length x values
n10sace_var = (np.sum(ace_ttn10**2) - ((np.sum(ace_ttn10)**2)/9))/8
n10sace_var
# y values
n10lc_ace = np.log(cor_n10ace_mat[0])
n10lc_ace
mk_n10ace = cor_n10ace_mat[1]/cor_n10ace_mat[0]
plt.figure(figsize=(7,7))
plt.errorbar(ace_ttn10, np.log(cor_n10ace_mat[0]), yerr=mk_n10ace, color='b', linestyle="None",marker='o',
capsize=5, capthick=1, ecolor='black')
from sklearn.linear_model import LinearRegression
model_n10ace = LinearRegression(fit_intercept=False)
model_n10ace.fit(ace_ttn10.reshape(-1,1), n10lc_ace)
# Slope here is in angstroms
print("Model slope: ", model_n10ace.coef_)
print("Model intercept:", model_n10ace.intercept_)
gg_n10ace = model_n10ace.predict(ace_ttn10.reshape(-1,1))
gg_n10ace
print("Mean Std Error:", sklearn.metrics.mean_squared_error(n10lc_ace, model_n10ace.predict(ace_ttn10.reshape(-1,1))))
print("R2 score:", sklearn.metrics.r2_score(n10lc_ace, model_n10ace.predict(ace_ttn10.reshape(-1,1))))
# Residuals between the true y data and model y data
resid_n10ace = n10lc_ace - gg_n10ace
resid_n10ace
# How to calculate mean squared error
np.sum(resid_n10ace**2)/len(resid_n10ace)
# Standard error: Square root of the mean squared error
np.sqrt(np.sum(resid_n10ace**2)/len(resid_n10ace))
# sample variance of the residuals
ntr_n10ace = resid_n10ace
ntr_n10ace -= np.mean(resid_n10ace)
nhr_n10ace = ntr_n10ace**2
np.sum(nhr_n10ace)/8
# How to calculate Sum((Xi - avg(X))^2): X values are the bond length values
nt_n10ace = np.arange(9, dtype=float)
nt_n10ace *= lb_avg_n6ace
nt_n10ace -= np.mean(nt_n10ace)
nhui = nt_n10ace**2
np.sum(nhui)
# How to calculate 95% confidence interval for the slope
flc_n10ace = scipy.stats.t.ppf(0.975, 4)*np.sqrt((np.sum(resid_n10ace**2)/len(resid_n10ace))/(np.sum(nhui)))
flc_n10ace
pers_n10ace =-1/model_n10ace.coef_
pers_n10ace
# Pers length error: error propagation from uncertainty in slope
flc_n10ace/((model_n10ace.coef_)**2)
def slope_plen(x,m):
return m*x
ttsam = np.arange(9, dtype=float)
ttsam *= lb_avg_n6ace
#nt_ttn10[0] = 0
ttsam
plt.figure(figsize=(7,7))
plt.errorbar(ace_ttn10, np.log(cor_n10ace_mat[0]), yerr=mk_n10ace, color='b',
linestyle="None",marker='o',capsize=5, capthick=1, ecolor='black')
plt.plot(ttsam, slope_plen(ttsam, model_n10ace.coef_[0]), color='b')
plt.title(r'Ensemble Averaged ln(Cosine $\theta$) in pure acetone', fontsize=15, y=1.01)
plt.xlabel(r'Bond Length', fontsize=15)
plt.ylabel(r'ln$\left< Cos(\theta)\right >$', fontsize=15)
#plt.ylim(-1.9,0)
font = font_manager.FontProperties(family='Arial', style='normal', size='14')
plt.legend([r'$N_{PLGA}$ = 10: $L_{p}$ = 40.7 $\AA$ ± 2.6 $\AA$'], loc=3, frameon=0, fontsize=14, prop=font)
plt.tick_params(labelsize=14)
plt.text(5,0,r'R$^{2}$ = 0.995', fontsize=15, color='blue')
def exfun(x, a):
return np.exp(-x*lb_avg_n6ace/a)
nt = np.arange(9)
plt.scatter(nt, cor_n10ace_mat[0,:])
plt.plot(nt,exfun(nt, pers_n10ace))
```
## N = 20 acetone
```
n20_plga_ace = mda.Universe("n20_plga_150ns/1_xa_soln/1xa_n20sys.gro", "n20_plga_150ns/1_xa_soln/N20_nopbcpp_0xa.xtc")
n20_plga_ace
len(n20_plga_ace.trajectory)
poly_n20_ace = n20_plga_ace.select_atoms("resname sPLG PLG tPLG and not type H")
com_bond_n20ace = np.zeros(shape=(1,5000))
count = 0
for ts in n20_plga_ace.trajectory[0:5000]:
n20_mon1 = n20_plga_ace.select_atoms("resid 1")
n20_mon2 = n20_plga_ace.select_atoms("resid 2")
oo_len = mda.analysis.distances.distance_array(n20_mon1.center_of_mass(), n20_mon2.center_of_mass(),
box=n20_plga_ace.trajectory.ts.dimensions)
com_bond_n20ace[0, count] = oo_len
count += 1
lb_avg_n20ace = np.mean(com_bond_n20ace)
lb_avg_n20ace
plt.scatter(np.arange(5000),com_bond_n6ace)
plt.scatter(np.arange(5000),com_bond_n8ace)
plt.scatter(np.arange(5000),com_bond_n10ace)
plt.scatter(np.arange(5000),com_bond_n20ace)
n20_blks_ace = bavg_pers_cnt(5, poly_n20_ace, n20_plga_ace, lb_avg_n6ace, 0 , 15000)
n20_blks_ace["Avg persistence length"]
n20_blks_ace["Avg Radius of gyration"]
np.mean(n20_blks_ace["Avg persistence length"])
np.std(n20_blks_ace["Avg persistence length"])
np.mean(n20_blks_ace["Avg Radius of gyration"])
np.std(n20_blks_ace["Avg Radius of gyration"])
n20_rgens_ace, cor_n20ace_mat, N20aceens_cos, rg_avgn20ace = get_rg_pers_poly(poly_n20_ace, n20_plga_ace, 0, 15000,
lb_avg_n6ace)
cor_n20ace_mat
cor_n20ace_mat[0]
# x values
ace_ttn20 = np.arange(19, dtype=float)
ace_ttn20 *= lb_avg_n6ace
#nt_ttn20[0] = 0
ace_ttn20
# Sample standard deviation of Bond Length x values
np.sqrt((np.sum(ace_ttn20**2) - ((np.sum(ace_ttn20)**2)/19))/18)
# Sample variance of Bond Length x values
n20sace_var = (np.sum(ace_ttn20**2) - ((np.sum(ace_ttn20)**2)/19))/18
n20sace_var
# y values
n20lc_ace = np.log(cor_n20ace_mat[0])
n20lc_ace
mk_n20ace = cor_n20ace_mat[1]/cor_n20ace_mat[0]
plt.figure(figsize=(7,7))
plt.errorbar(ace_ttn20, np.log(cor_n20ace_mat[0]), yerr=mk_n20ace, color='b', linestyle="None",marker='o',
capsize=5, capthick=1, ecolor='black')
ace_ttn20[:7]
from sklearn.linear_model import LinearRegression
model_n20ace = LinearRegression(fit_intercept=False)
model_n20ace.fit(ace_ttn20[:7].reshape(-1,1), n20lc_ace[:7])
# Slope here is in angstroms
print("Model slope: ", model_n20ace.coef_)
print("Model intercept:", model_n20ace.intercept_)
gg_n20ace = model_n20ace.predict(ace_ttn20.reshape(-1,1))
gg_n20ace
print("Mean Std Error:", sklearn.metrics.mean_squared_error(n20lc_ace[:7],
model_n20ace.predict(ace_ttn20[:7].reshape(-1,1))))
print("R2 score:", sklearn.metrics.r2_score(n20lc_ace[:7],
model_n20ace.predict(ace_ttn20[:7].reshape(-1,1))))
# Residuals between the true y data and model y data
resid_n20ace = n20lc_ace[:7] - gg_n20ace[:7]
resid_n20ace
# How to calculate mean squared error
np.sum(resid_n20ace**2)/len(resid_n20ace)
# Standard error: Square root of the mean squared error
np.sqrt(np.sum(resid_n20ace**2)/len(resid_n20ace))
# sample variance of the residuals
ntr_n20ace = resid_n20ace
ntr_n20ace -= np.mean(resid_n20ace)
nhr_n20ace = ntr_n20ace**2
np.sum(nhr_n20ace)/7
# How to calculate Sum((Xi - avg(X))^2): X values are the bond length values, I only used 7 points to fit data
nt_n20ace = np.arange(7, dtype=float)
nt_n20ace *= lb_avg_n6ace
nt_n20ace -= np.mean(nt_n20ace)
nhui = nt_n20ace**2
np.sum(nhui)
# How to calculate 95% confidence interval for the slope
flc_n20ace = scipy.stats.t.ppf(0.975, 4)*np.sqrt((np.sum(resid_n20ace**2)/len(resid_n20ace))/(np.sum(nhui)))
flc_n20ace
pers_n20ace =-1/model_n20ace.coef_
pers_n20ace
# Pers length error: error propagation from uncertainty in slope
flc_n20ace/((model_n20ace.coef_)**2)
def slope_plen(x,m):
return m*x
ttsam = np.arange(19, dtype=float)
ttsam *= lb_avg_n6ace
#nt_ttn10[0] = 0
ttsam
plt.figure(figsize=(7,7))
plt.errorbar(ace_ttn20, np.log(cor_n20ace_mat[0]), yerr=mk_n20ace, color='b',
linestyle="None",marker='o',capsize=5, capthick=1, ecolor='black')
plt.plot(ttsam, slope_plen(ttsam, model_n20ace.coef_[0]), color='b')
plt.title(r'Ensemble Averaged ln(Cosine $\theta$) in pure acetone', fontsize=15, y=1.01)
plt.xlabel(r'Bond Length', fontsize=15)
plt.ylabel(r'ln$\left< Cos(\theta)\right >$', fontsize=15)
#plt.ylim(-1.9,0)
font = font_manager.FontProperties(family='Arial', style='normal', size='14')
plt.legend([r'$N_{PLGA}$ = 20: $L_{p}$ = 45.6 $\AA$ ± 3.6 $\AA$'], loc=3, frameon=0, fontsize=14, prop=font)
plt.tick_params(labelsize=14)
plt.text(5,0,r'R$^{2}$ = 0.994', fontsize=15, color='blue')
def exfun(x, a):
return np.exp(-x*lb_avg_n6ace/a)
nt = np.arange(19)
plt.scatter(nt, cor_n20ace_mat[0,:])
plt.plot(nt,exfun(nt, pers_n20ace))
```
## N = 30 acetone
```
n30_plga_ace = mda.Universe("n30_plga_150ns/1_xa_soln/1xa_n30sys.gro", "n30_plga_150ns/1_xa_soln/N30_nopbcpp_1xa.xtc")
n30_plga_ace
len(n30_plga_ace.trajectory)
poly_n30_ace = n30_plga_ace.select_atoms("resname sPLG PLG tPLG and not type H")
com_bond_n30ace = np.zeros(shape=(1,5000))
count = 0
for ts in n30_plga_ace.trajectory[0:5000]:
n30_mon1 = n30_plga_ace.select_atoms("resid 1")
n30_mon2 = n30_plga_ace.select_atoms("resid 2")
oo_len = mda.analysis.distances.distance_array(n30_mon1.center_of_mass(), n30_mon2.center_of_mass(),
box=n30_plga_ace.trajectory.ts.dimensions)
com_bond_n30ace[0, count] = oo_len
count += 1
lb_avg_n30ace = np.mean(com_bond_n30ace)
lb_avg_n30ace
plt.scatter(np.arange(5000),com_bond_n6ace)
plt.scatter(np.arange(5000),com_bond_n8ace)
plt.scatter(np.arange(5000),com_bond_n10ace)
plt.scatter(np.arange(5000),com_bond_n20ace)
plt.scatter(np.arange(5000),com_bond_n30ace)
n30_blks_ace = bavg_pers_cnt(5, poly_n30_ace, n30_plga_ace, lb_avg_n6ace, 0 , 15000)
n30_blks_ace["Avg persistence length"]
n30_blks_ace["Avg Radius of gyration"]
np.mean(n30_blks_ace["Avg persistence length"])
np.std(n30_blks_ace["Avg persistence length"])
np.mean(n30_blks_ace["Avg Radius of gyration"])
np.std(n30_blks_ace["Avg Radius of gyration"])
n30_rgens_ace, cor_n30ace_mat, N30aceens_cos, rg_avgn30ace = get_rg_pers_poly(poly_n30_ace, n30_plga_ace, 0, 15000,
lb_avg_n6ace)
cor_n30ace_mat
cor_n30ace_mat[0]
# x values
ace_ttn30 = np.arange(29, dtype=float)
ace_ttn30 *= lb_avg_n6ace
#nt_ttn20[0] = 0
ace_ttn30
# Sample standard deviation of Bond Length x values
np.sqrt((np.sum(ace_ttn30**2) - ((np.sum(ace_ttn30)**2)/29))/28)
# Sample variance of Bond Length x values
n30sace_var = (np.sum(ace_ttn30**2) - ((np.sum(ace_ttn30)**2)/29))/28
n30sace_var
# y values
n30lc_ace = np.log(cor_n30ace_mat[0])
n30lc_ace
mk_n30ace = cor_n30ace_mat[1]/cor_n30ace_mat[0]
plt.figure(figsize=(7,7))
plt.errorbar(ace_ttn30, np.log(cor_n30ace_mat[0]), yerr=mk_n30ace, color='b', linestyle="None",marker='o',
capsize=5, capthick=1, ecolor='black')
ace_ttn30[:8]
from sklearn.linear_model import LinearRegression
model_n30ace = LinearRegression(fit_intercept=False)
model_n30ace.fit(ace_ttn30[:7].reshape(-1,1), n30lc_ace[:7])
# Slope here is in angstroms
print("Model slope: ", model_n30ace.coef_)
print("Model intercept:", model_n30ace.intercept_)
gg_n30ace = model_n30ace.predict(ace_ttn30.reshape(-1,1))
gg_n30ace
print("Mean Std Error:", sklearn.metrics.mean_squared_error(n30lc_ace[:7],
model_n30ace.predict(ace_ttn30[:7].reshape(-1,1))))
print("R2 score:", sklearn.metrics.r2_score(n30lc_ace[:7],
model_n30ace.predict(ace_ttn30[:7].reshape(-1,1))))
# Residuals between the true y data and model y data
resid_n30ace = n30lc_ace[:7] - gg_n30ace[:7]
resid_n30ace
# How to calculate mean squared error
np.sum(resid_n30ace**2)/len(resid_n30ace)
# Standard error: Square root of the mean squared error
np.sqrt(np.sum(resid_n30ace**2)/len(resid_n30ace))
# sample variance of the residuals
ntr_n30ace = resid_n30ace
ntr_n30ace -= np.mean(resid_n30ace)
nhr_n30ace = ntr_n30ace**2
# divide by n-1 samples used to fit
np.sum(nhr_n30ace)/6
# How to calculate Sum((Xi - avg(X))^2): X values are the bond length values
nt_n30ace = np.arange(7, dtype=float)
nt_n30ace *= lb_avg_n6ace
nt_n30ace -= np.mean(nt_n30ace)
nhui = nt_n30ace**2
np.sum(nhui)
# How to calculate 95% confidence interval for the slope
flc_n30ace = scipy.stats.t.ppf(0.975, 4)*np.sqrt((np.sum(resid_n30ace**2)/len(resid_n30ace))/(np.sum(nhui)))
flc_n30ace
pers_n30ace =-1/model_n30ace.coef_
pers_n30ace
# Pers length error: error propagation from uncertainty in slope
flc_n30ace/((model_n30ace.coef_)**2)
def slope_plen(x,m):
return m*x
ttsam = np.arange(19, dtype=float)
ttsam *= lb_avg_n6ace
#nt_ttn10[0] = 0
ttsam
plt.figure(figsize=(7,7))
plt.errorbar(ace_ttn30, np.log(cor_n30ace_mat[0]), yerr=mk_n30ace, color='b',
linestyle="None",marker='o',capsize=5, capthick=1, ecolor='black')
plt.plot(ttsam, slope_plen(ttsam, model_n30ace.coef_[0]), color='b')
plt.title(r'Ensemble Averaged ln(Cosine $\theta$) in pure acetone', fontsize=15, y=1.01)
plt.xlabel(r'Bond Length', fontsize=15)
plt.ylabel(r'ln$\left< Cos(\theta)\right >$', fontsize=15)
#plt.ylim(-1.9,0)
font = font_manager.FontProperties(family='Arial', style='normal', size='14')
plt.legend([r'$N_{PLGA}$ = 30: $L_{p}$ = 39.4 $\AA$ ± 4.3 $\AA$'], loc=3, frameon=0, fontsize=14, prop=font)
plt.tick_params(labelsize=14)
plt.text(5,0,r'R$^{2}$ = 0.99', fontsize=15, color='blue')
plt.figure(figsize=(7,7))
plt.errorbar(ace_ttn30, np.log(cor_n30ace_mat[0]), yerr=mk_n30ace, color='b', linestyle="None",marker='h',
capsize=5, capthick=1, ecolor='black')
plt.plot(ttsam, slope_plen(ttsam, model_n30ace.coef_[0]), color='b')
plt.errorbar(ace_ttn20, np.log(cor_n20ace_mat[0]), yerr=mk_n20ace, color='g', linestyle="None"
,marker='*',capsize=5, capthick=1, ecolor='black')
plt.plot(ttsam, slope_plen(ttsam, model_n20ace.coef_[0]), color='g')
plt.errorbar(ace_ttn10, np.log(cor_n10ace_mat[0]), yerr=mk_n10ace, color='r', linestyle="None"
,marker='v', capsize=5, capthick=1, ecolor='black')
plt.plot(ttsam, slope_plen(ttsam, model_n10ace.coef_[0]), color='r')
plt.errorbar(ace_ttn8, np.log(cor_n8ace_mat[0]), yerr=mk_n8ace, color='k', linestyle="None",marker='s',
capsize=5, capthick=1, ecolor='black')
plt.plot(ttsam, slope_plen(ttsam, model_n8ace.coef_[0]), color='k')
plt.errorbar(ace_ttn6, np.log(cor_n6ace_mat[0]), yerr=mk_n6ace, color='c', linestyle="None",marker='x',
capsize=5, capthick=1, ecolor='black')
plt.plot(ttsam, slope_plen(ttsam, model_n6ace.coef_[0]), color='m')
plt.title(r'Ensemble Averaged ln(Cosine $\theta$): Pure acetone', fontsize=15, y=1.01)
plt.xlabel(r'Bond Length', fontsize=15)
plt.ylabel(r'ln$\left< Cos(\theta)\right >$', fontsize=15)
plt.ylim(-5, 1)
plt.xlim(0,70)
font = font_manager.FontProperties(family='Arial', style='normal', size='14')
#plt.legend([r'$N_{PLGA}$ = 30: $L_{p}$ = 18.5 $\AA$ ± 0.15 $\AA$, R$^{2}$ = 0.99', r'$N_{PLGA}$ = 20: $L_{p}$ = 19.1 $\AA$ ± 0.27 $\AA$, R$^{2}$ = 0.99'],
# loc=3, frameon=0, fontsize=14, prop=font)
#plt.legend([r'$N_{PLGA}$ = 20: $L_{p}$ = 19.1 $\AA$ ± 0.27 $\AA$, R$^{2}$ = 0.99'], loc=3, frameon=0, fontsize=14, prop=font)
plt.tick_params(labelsize=14)
plt.text(1, -4.9,r'$N_{PLGA}$ = 30: $L_{p}$ = 39.4 $\AA$ ± 4.3 $\AA$, R$^{2}$ = 0.99', fontsize=14, color='blue')
plt.text(1,-4.65,r'$N_{PLGA}$ = 20: $L_{p}$ = 45.6 $\AA$ ± 3.6 $\AA$, R$^{2}$ = 0.99', fontsize=14, color='green')
plt.text(1,-4.38,r'$N_{PLGA}$ = 10: $L_{p}$ = 40.7 $\AA$ ± 2.6 $\AA$, R$^{2}$ = 0.99', fontsize=14, color='red')
plt.text(1,-4.12,r'$N_{PLGA}$ = 8: $L_{p}$ = 39.3 $\AA$ ± 5.9 $\AA$, R$^{2}$ = 0.98', fontsize=14, color='black')
plt.text(1,-3.85,r'$N_{PLGA}$ = 6: $L_{p}$ = 41.6 $\AA$ ± 5.7 $\AA$, R$^{2}$ = 0.99', fontsize=14, color='magenta')
#plt.text(2.5,0,r'R$^{2}$ = 0.96', fontsize=15, color='red')
def exfun(x, a):
return np.exp(-x*lb_avg_n6ace/a)
nt = np.arange(29)
plt.scatter(nt, cor_n30ace_mat[0,:])
plt.plot(nt,exfun(nt, pers_n30ace))
```
| github_jupyter |
### Erdős–Rényi Model
You may wonder, isn't this script called Watts–Strogatz model? Sure, but Erdős–Rényi model is the foundation of Watts–Strogatz model. It is reasonable to make an introduction on the most basic form of random graph before we move onto the sophisticated one. In Erdős–Rényi model, any random pair of vertices are connected at the same probability. Thus, the model follows binomial distribution. When the number of vertices is large enough (converged to infinity), the model follows Poisson distribution.
Unfortunately, the quality of pdf isn't exactly top-notch since the original paper was published in 1959. Given the simplicity of the model, any university material should be able to cover its content.
http://snap.stanford.edu/class/cs224w-readings/erdos59random.pdf
http://snap.stanford.edu/class/cs224w-readings/erdos60random.pdf
```
import os
os.chdir('K:/ecole/github')
import matplotlib.pyplot as plt
import numpy as np
#graph adt
#check the below link for more details
# https://github.com/je-suis-tm/graph-theory/blob/master/graph.py
import graph
ermodel=graph.graph()
#initial parameters
num_of_v=200
prob=0.3
#connect two vertices based upon probability
for i in range(num_of_v):
for j in range(i+1,num_of_v):
if np.random.uniform()<prob:
ermodel.append(i,j,1)
ermodel.append(j,i,1)
```
##### Degree Distribution
```
#get degree
degree_dst=[ermodel.degree(node) for node in ermodel.vertex()]
#viz
ax=plt.figure(figsize=(10,5)).add_subplot(111)
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
plt.hist(degree_dst,bins=30,width=1,color='#9e90b0')
plt.title('Degree Distribution of Erdős–Rényi Model')
plt.xlabel('Degree')
plt.ylabel('Frequency')
plt.show()
```
##### NetworkX
```
import networkx as nx
G=nx.random_graphs.erdos_renyi_graph(num_of_v,prob)
```
### Lattice
Wait, still not Watts-Strogatz model? Tranquillo, a lil bit foreplay kills nobody. Lattice is basically a grid. Look at the pyramids in Louvre or Montparnasse in southbank. Those glass windows form a squared lattice.
```
lattice=graph.graph()
#initial parameters
num_of_v=100
num_of_neighbors=4
assert num_of_neighbors%2==0,"number of neighbors must be even number"
#we are creating a regular ring lattice
for i in range(num_of_v):
for j in range(1,num_of_neighbors//2+1):
lattice.append(i,i+j if i+j<num_of_v else i+j-num_of_v,1)
lattice.append(i,i-j if i-j>=0 else i-j+num_of_v,1)
lattice.append(i+j if i+j<num_of_v else i+j-num_of_v,i,1)
lattice.append(i-j if i-j>=0 else i-j+num_of_v,i,1)
```
##### Degree Distribution
There is no point of degree distribution since each vertex has 4 edges.
##### NetworkX
```
#networkx can create a squared lattice
G=nx.grid_2d_graph(4,4)
#to create a ring lattice
#we d better leverage its watts strogatz model
#set rewiring probability at 0
G=nx.random_graphs.watts_strogatz_graph(10,4,0)
```
### Watts-Strogatz Model
Third time is the charm. Here is the climax of the script. Watts-Strogatz model, so-called small world effect, is somewhere between ring lattice and Erdős–Rényi model. Its partial regularity creates clusters in the structure and its partial randomness brings disorder to the degree distribution. The key parameter is the probability of rewiring. When the probability is zero, it becomes a ring lattice. When the probability is one, it becomes Erdős–Rényi model.
```
wsmodel=graph.graph()
#initial parameters
num_of_v=200
num_of_neighbors=60
prob=0.3
assert num_of_neighbors%2==0,"number of neighbors must be even number"
#first we create a regular ring lattice
for i in range(num_of_v):
for j in range(1,num_of_neighbors//2+1):
wsmodel.append(i,i+j if i+j<num_of_v else i+j-num_of_v,1)
wsmodel.append(i,i-j if i-j>=0 else i-j+num_of_v,1)
wsmodel.append(i+j if i+j<num_of_v else i+j-num_of_v,i,1)
wsmodel.append(i-j if i-j>=0 else i-j+num_of_v,i,1)
#rewiring
#remove a random edge and create a random edge
for i in wsmodel.vertex():
for j in wsmodel.edge(i):
if np.random.uniform()<prob:
wsmodel.disconnect(i,j)
wsmodel.disconnect(j,i)
rewired=np.random.choice(wsmodel.vertex())
wsmodel.append(i,rewired,1)
wsmodel.append(rewired,i,1)
```
##### Degree Distribution
```
#get degree
degree_dst=[wsmodel.degree(node) for node in wsmodel.vertex()]
#viz
ax=plt.figure(figsize=(10,5)).add_subplot(111)
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
plt.hist(degree_dst,bins=30,width=0.7,color='#124fcc')
plt.title('Degree Distribution of Watts-Strogatz Model')
plt.xlabel('Degree')
plt.ylabel('Frequency')
plt.show()
```
##### NetworkX
```
#create an erdős rényi
#effectively 20 vertices with probability at 0.2
ger=nx.random_graphs.watts_strogatz_graph(20,4,1)
#create a ring lattice
grl=nx.random_graphs.watts_strogatz_graph(20,4,0)
#create a watts strogatz
gws=nx.random_graphs.watts_strogatz_graph(20,4,0.4)
#viz
ax=plt.figure(figsize=(5,15))
ax.add_subplot(311)
nx.draw_circular(ger)
plt.title('Erdős-Rényi\np=1')
ax.add_subplot(312)
nx.draw_circular(grl)
plt.title('Ring Lattice\np=0')
ax.add_subplot(313)
nx.draw_circular(gws)
plt.title('Watts-Strogatz\np=0.4')
plt.show()
```
| github_jupyter |
# PyTorch CIFAR-10 local training
## Prerequisites
This notebook shows how to use the SageMaker Python SDK to run your code in a local container before deploying to SageMaker's managed training or hosting environments. This can speed up iterative testing and debugging while using the same familiar Python SDK interface. Just change your estimator's `train_instance_type` to `local` (or `local_gpu` if you're using an ml.p2 or ml.p3 notebook instance).
In order to use this feature, you'll need to install docker-compose (and nvidia-docker if training with a GPU).
**Note: you can only run a single local notebook at one time.**
```
!/bin/bash ./setup.sh
```
## Overview
The **SageMaker Python SDK** helps you deploy your models for training and hosting in optimized, productions ready containers in SageMaker. The SageMaker Python SDK is easy to use, modular, extensible and compatible with TensorFlow, MXNet, PyTorch. This tutorial focuses on how to create a convolutional neural network model to train the [CIFAR-10 dataset](https://www.cs.toronto.edu/~kriz/cifar.html) using **PyTorch in local mode**.
### Set up the environment
This notebook was created and tested on a single ml.p2.xlarge notebook instance.
Let's start by specifying:
- The S3 bucket and prefix that you want to use for training and model data. This should be within the same region as the Notebook Instance, training, and hosting.
- The IAM role arn used to give training and hosting access to your data. See the documentation for how to create these. Note, if more than one role is required for notebook instances, training, and/or hosting, please replace the sagemaker.get_execution_role() with appropriate full IAM role arn string(s).
```
import sagemaker
sagemaker_session = sagemaker.Session()
bucket = sagemaker_session.default_bucket()
prefix = 'sagemaker/DEMO-pytorch-cnn-cifar10'
role = sagemaker.get_execution_role()
import os
import subprocess
raise Exception("This is a test error.")
instance_type = 'local'
if subprocess.call('nvidia-smi') == 0:
## Set type to GPU if one is present
instance_type = 'local_gpu'
print("Instance type = " + instance_type)
```
### Download the CIFAR-10 dataset
```
from utils_cifar import get_train_data_loader, get_test_data_loader, imshow, classes
trainloader = get_train_data_loader()
testloader = get_test_data_loader()
```
### Data Preview
```
import numpy as np
import torchvision, torch
# get some random training images
dataiter = iter(trainloader)
images, labels = dataiter.next()
# show images
imshow(torchvision.utils.make_grid(images))
# print labels
print(' '.join('%9s' % classes[labels[j]] for j in range(4)))
```
### Upload the data
We use the ```sagemaker.Session.upload_data``` function to upload our datasets to an S3 location. The return value inputs identifies the location -- we will use this later when we start the training job.
```
inputs = sagemaker_session.upload_data(path='data', bucket=bucket, key_prefix='data/cifar10')
```
# Construct a script for training
Here is the full code for the network model:
```
!pygmentize source/cifar10.py
```
## Script Functions
SageMaker invokes the main function defined within your training script for training. When deploying your trained model to an endpoint, the model_fn() is called to determine how to load your trained model. The model_fn() along with a few other functions list below are called to enable predictions on SageMaker.
### [Predicting Functions](https://github.com/aws/sagemaker-pytorch-containers/blob/master/src/sagemaker_pytorch_container/serving.py)
* model_fn(model_dir) - loads your model.
* input_fn(serialized_input_data, content_type) - deserializes predictions to predict_fn.
* output_fn(prediction_output, accept) - serializes predictions from predict_fn.
* predict_fn(input_data, model) - calls a model on data deserialized in input_fn.
The model_fn() is the only function that doesn't have a default implementation and is required by the user for using PyTorch on SageMaker.
## Create a training job using the sagemaker.PyTorch estimator
The `PyTorch` class allows us to run our training function on SageMaker. We need to configure it with our training script, an IAM role, the number of training instances, and the training instance type. For local training with GPU, we could set this to "local_gpu". In this case, `instance_type` was set above based on your whether you're running a GPU instance.
After we've constructed our `PyTorch` object, we fit it using the data we uploaded to S3. Even though we're in local mode, using S3 as our data source makes sense because it maintains consistency with how SageMaker's distributed, managed training ingests data.
```
from sagemaker.pytorch import PyTorch
cifar10_estimator = PyTorch(entry_point='source/cifar10.py',
role=role,
framework_version='1.4.0',
train_instance_count=1,
train_instance_type=instance_type)
cifar10_estimator.fit(inputs)
```
# Deploy the trained model to prepare for predictions
The deploy() method creates an endpoint (in this case locally) which serves prediction requests in real-time.
```
from sagemaker.pytorch import PyTorchModel
cifar10_predictor = cifar10_estimator.deploy(initial_instance_count=1,
instance_type=instance_type)
```
# Invoking the endpoint
```
# get some test images
dataiter = iter(testloader)
images, labels = dataiter.next()
# print images
imshow(torchvision.utils.make_grid(images))
print('GroundTruth: ', ' '.join('%4s' % classes[labels[j]] for j in range(4)))
outputs = cifar10_predictor.predict(images.numpy())
_, predicted = torch.max(torch.from_numpy(np.array(outputs)), 1)
print('Predicted: ', ' '.join('%4s' % classes[predicted[j]]
for j in range(4)))
```
# Clean-up
Deleting the local endpoint when you're finished is important, since you can only run one local endpoint at a time.
```
cifar10_estimator.delete_endpoint()
```
| github_jupyter |
# Gaussian Process Latent Variable Models
## Imports
```
import numpy as np
import matplotlib.pyplot as plt
import matplotlib
import seaborn as sns
import tensorflow as tf
from tqdm.autonotebook import tqdm
from tensorflow_probability import distributions as tfd
from tensorflow_probability import positive_semidefinite_kernels as tfk
%pylab inline
sns.set_context('talk',font_scale=1.5)
matplotlib.rcParams['figure.figsize'] = (8.0, 6.0)
def reset_session():
"""Creates a new global, interactive session in Graph-mode."""
global sess
try:
tf.reset_default_graph()
sess.close()
except:
pass
sess = tf.InteractiveSession()
reset_session()
```
## Load fashion-MNIST Data
```
(x_train, y_train), (_, _) = tf.keras.datasets.fashion_mnist.load_data()
N = 1000
small_x_train = x_train[:N, ...].astype(np.float64) / 256.
small_y_train = y_train[:N]
```
## Variables entrenables
```
# Variables entrenables
amplitude = np.finfo(np.float64).eps + tf.nn.softplus(
tf.get_variable(name='amplitude',
dtype=tf.float64,
initializer=np.float64(1.)))
length_scale = np.finfo(np.float64).eps + tf.nn.softplus(
tf.get_variable(name='length_scale',
dtype=tf.float64,
initializer=np.float64(1.)))
observation_noise_variance = np.finfo(np.float64).eps + tf.nn.softplus(
tf.get_variable(name='observation_noise_variance',
dtype=tf.float64,
initializer=np.float64(1.)))
observations_ = small_x_train.reshape(N, -1).transpose()
init_ = np.random.normal(size=(N, 2))
latent_index_points = tf.get_variable(
name='latent_index_points',
dtype=tf.float64,
initializer=init_)
```
## Modelo y entrenamiento
```
kernel = tfk.ExponentiatedQuadratic(amplitude, length_scale)
gp = tfd.GaussianProcess(
kernel=kernel,
index_points=latent_index_points,
observation_noise_variance=observation_noise_variance)
log_probs = gp.log_prob(observations_, name='log_prob')
loss = -tf.reduce_mean(log_probs)
optimizer = tf.train.AdamOptimizer(learning_rate=.1)
train_op = optimizer.minimize(loss)
```
## Construct predictive model and sampling ops
```
sample_grid_points = 10
grid_ = np.linspace(-4, 4, sample_grid_points).astype(np.float64)
grid_ = np.stack(np.meshgrid(grid_, grid_), axis=-1)
grid_ = grid_.reshape(sample_grid_points, sample_grid_points, 1, 1, 2)
gprm = tfd.GaussianProcessRegressionModel(
kernel=kernel,
index_points=grid_,
observation_index_points=latent_index_points,
observations=observations_)
samples = gprm.sample()
```
## Entrenar y conseguir embeddings
```
sess.run(tf.global_variables_initializer())
num_iters = 100
log_interval = 20
lips_ = np.zeros((num_iters, N, 2), np.float64)
pbar = tqdm(range(num_iters))
for i in pbar:
_, loss_, lips_[i] = sess.run([train_op, loss, latent_index_points])
pbar.set_postfix({'loss':loss_})
```
## Resultados
```
plt.figure(figsize=(7, 7))
plt.title("Before training")
plt.grid('off')
plt.scatter(x=init_[:, 0], y=init_[:, 1],
c=y_train[:N], cmap=plt.get_cmap('Paired'), s=25)
plt.show()
plt.figure(figsize=(7, 7))
plt.title("After training")
plt.grid('off')
plt.scatter(x=lips_[-1, :, 0], y=lips_[-1, :, 1],
c=y_train[:N], cmap=plt.get_cmap('Paired'), s=25)
plt.show()
```
## Samplear nuestro espacio
```
samples_ = sess.run(samples)
samples_ = np.squeeze(samples_)
samples_ = ((samples_ -
samples_.min(-1, keepdims=True)) /
(samples_.max(-1, keepdims=True) -
samples_.min(-1, keepdims=True)))
samples_ = samples_.reshape(sample_grid_points, sample_grid_points, 28, 28)
samples_ = samples_.transpose([0, 2, 1, 3])
samples_ = samples_.reshape(28 * sample_grid_points, 28 * sample_grid_points)
plt.figure(figsize=(7, 7))
ax = plt.subplot()
ax.grid('off')
ax.imshow(-samples_, interpolation='none', cmap='Greys')
plt.show()
```
| github_jupyter |
```
import os
import sys
module_path = os.path.abspath(os.path.join('../../src'))
print(module_path)
if module_path not in sys.path:
sys.path.append(module_path)
from datetime import timedelta
import datetime
import os
from fileUtils import read_file_properties_v2,find_filesv2,list_files,get_labeled_exif
from fileUtils import get_audio,str2timestamp,query_audio
from labeling_utils import splitmp3
import pandas as pd
from pathlib import Path
# from sklearn.neighbors import KDTree
import numpy as np
from pydub import AudioSegment
from annoy import AnnoyIndex
# from sklearn.metrics.pairwise import paired_distances
# from sklearn.metrics import pairwise_distances
# def paired_distances_broadcast(X,Y):
# return paired_distances(np.repeat(X,Y.shape[0],axis=0), Y)
# def sigmoid(X):
# return 1/(1+np.exp(-X))
# def sigmoidT(X,t):
# return 1/(1+np.exp(-X*t))
# # from fileUtils import save_to_csv
# # import csv
# # def save_to_csv(file_name,lines):
# # file_name=Path(file_name).with_suffix('.csv')
# # with open(file_name, mode='a') as labels_file:
# # label_writer = csv.writer(labels_file, delimiter=',', quotechar='"', quoting=csv.QUOTE_MINIMAL)
# # for line in lines:
# # label_writer.writerow(line)
# output = find_filesv2(location,start_time,end_time,length,0,file_properties_df)
# sorted_filtered,start_time,end_time,start_time_org,end_time_org = output
```
#### find the embedding for a given image
```
# p_path="/home/enis/projects/nna/data/cameraTraps/test.txt"
# with open(p_path) as p_file:
# photo_paths=p_file.readlines()
# photo_paths = [i.strip() for i in photo_paths]
# other_folder=["anwr_35_2019_101RECNX_RCNX3373.JPG",
# "anwr_42_2019_100RECNX_RCNX3574.JPG",
# "ivvavik_SINP03_2019_100RECNX_IMG_3219.JPG",
# "ivvavik_SINP05_2019_100RECNX_IMG_2867.JPG",
# "ivvavik_SINP06_2019_100RECNX_IMG_1884.JPG",
# "ivvavik_SINP09_2019_100RECNX_IMG_2743.JPG",
# "prudhoe_17_2019_100RECNX_RCNX3916.JPG",]
# parent_path1="/tank/data/nna/examples/randomPhotos10k/"
# parent_path2="/tank/data/nna/examples/randomPhotos1k/"
# # photo with Caribou
# photo_paths=['anwr_37_2019_100RECNX_RCNX9317.jpg']
# given image paths finds files, code is at:
# notebooks/explore/get_audio4photos.ipynb
file_properties_df=pd.read_pickle("../../data/realdata_v2No_stinchcomb.pkl")
```
### 1. pick a sound to use for queries
```
# sound with Caribou
!find /scratch/enis/data/nna/real/ -iname "S4A10279_20190605_091602*"
# aircraft sound
!find /scratch/enis/data/nna/real/ -iname "S4A10255_20190507_073000*"
```
### 1.1 find similar embeddings for given embeddings in the same location
```
# sound with Caribou
raw_embed_name="/scratch/enis/data/nna/real/anwr/37/2019/S4A10279_20190605_091602_vgg/S4A10279_20190605_091602_rawembeddings000.npy"
raw_embed=np.load(raw_embed_name)
# aircraft sound
# raw_embed_name="/scratch/enis/data/nna/real/prudhoe/26/2019/S4A10255_20190507_073000_vgg/S4A10255_20190507_073000_rawembeddings000.npy"
raw_embed_name="/scratch/enis/data/nna/real/prudhoe/26/2019/S4A10255_20190507_073000_vgg/S4A10255_20190507_073000_embeddings000.npy"
raw_embed=np.load(raw_embed_name)
raw_embed=raw_embed.astype(np.float32)
# S4A10255_20190507_073000-1808second
(30*60)+8
```
### 1.2 Slice embedding for exact time of interest from a big file
```
# aircraft sound
start_seconds=(30*60)+8
raw_embed_audio=raw_embed[int(start_seconds):int(start_seconds)+60]
# # sound with Caribou
# start_seconds=(start_time-sorted_filtered["timestamp"])[0].total_seconds()
# raw_embed_audio=raw_embed[int(start_seconds):int(start_seconds)+60]
raw_embed_audio.shape
```
### 2.1 load embeddings to search from
```
%%time
# find all files
import glob
search_path="/scratch/enis/data/nna/real/prudhoe/26/"
filenamePattern="*_embeddings*.npy"
# all_embeddings=list_files("/scratch/enis/data/nna/real/anwr/37/",filenamePattern)
# aircraft sound
all_embeddings=list_files(search_path,filename=filenamePattern)
# remove original embedding from the list
# del all_embeddings[all_embeddings.index(raw_embed_name)]
# # len(all_embeddings)
# all_embeddings=[e for e in all_embeddings if "_embeddings" in e]
```
### 2.1.1 this is for calculating distances ourself
```
%%time
# load and merge embedding files
#LOAD:
embeds=[None]*len(all_embeddings)
index=0
for i,embed_file in enumerate(all_embeddings):
embed=np.load(embed_file)
index+=embed.shape[0]
embeds[i]=(embed)
#MERGE:
# Faster then np.concatenate
concat_embeds=np.zeros([index,128],dtype=np.float32)
index=0
for i,embed in enumerate(embeds):
concat_embeds[index:(index+embed.shape[0]),:]=embed[:]
index+=embed.shape[0]
concat_embeds.shape
```
### 2.1.2 create an ANNOY index for files at all_path on disk
```
%%time
import time
# vector size
f=128
t = AnnoyIndex(f, 'euclidean') # Length of item vector that will be indexed
# for i in range(concat_embeds.shape[0]):
# t.add_item(i, concat_embeds[i,:])
#prepares annoy to build the index in the specified file instead of RAM
#(execute before adding items, no need to save after build)
fn="/scratch/enis/data/nna/realMerged/prudhoe26_V11.ann"
t.on_disk_build(fn)
index=0
start=time.time()
for i,embed_file in enumerate(all_embeddings):
embed=np.load(embed_file)
for ii in range(embed.shape[0]):
t.add_item(index, embed[ii,:])
index+=1
if i%1000==0:
end=time.time()
print(i//1000,end-start)
start=end
t.set_seed(42)
t.build(16)
```
### 3.1 Use Annoy NNs to approximate
```
queries=[i for i in range(100,110,1)]
# QNNs=[[],[]]
Qmatrix=np.empty((len(queries),len(queries)))
sizeOfDataset=t.get_n_items()
for i,q in enumerate(queries):
IDs,Distances=t.get_nns_by_item(q, 2,include_distances=True)
IDs,Distances=IDs[1:],Distances[1:]
Qmatrix[i,i]=Distances[0]
for i2,q2 in enumerate(queries):
if i==i2:
continue
else:
alingedID=(IDs[0]+(i2-i))
alingedID=0 if (alingedID)<0 else alingedID
alingedID = alingedID if alingedID<sizeOfDataset else sizeOfDataset-1
Qmatrix[i,i2]=t.get_distance(IDs[0],alingedID)
averageNNs_approx=np.average(Qmatrix,axis=1)
averageNNs_approx
# t.get_n_items()
len(all_path),len(all_embeddings)
!ls -alh /scratch/enis/data/nna/realMerged/allEmbeddings.ann
# concat_embeds.shape
# make sure it multiples of 10 to make things easy
concat_embeds=concat_embeds[:(concat_embeds.shape[0]//10)*10,:]
```
### there are three methods for creating vectors:
```
# # Method 1 1 second
excerptLen=1
exp_name="1second"
concat_embeds2=concat_embeds[:]
raw_embed_audio2=raw_embed_audio[:]
raw_embed_audio2=raw_embed_audio2[20:21,:]
rowN=100000
#Method 2 mean 10 second
# excerptLen=10
# exp_name="Mean10Second"
# concat_embeds2=concat_embeds.reshape(-1,10,128).mean(axis=1)
# raw_embed_audio2=raw_embed_audio.reshape(-1,10,128).mean(axis=1)
# raw_embed_audio2=raw_embed_audio2[2:3,:]
# rowN=10000
# #Method 3 concat 10 second
# excerptLen=10
# exp_name="Concat10Second"
# concat_embeds2=concat_embeds.reshape(-1,1280)
# raw_embed_audio2=raw_embed_audio.reshape(-1,1280)
# raw_embed_audio2=raw_embed_audio2[2:3,:]
# rowN=10000
concat_embeds.shape
# concat_embeds3=concat_embeds[:,:]
```
* we are trying to create a matrix, rows are queries and columns are data points in database
* values are similarity,
* then
```
raw_embed_audio.shape
# paired_distances(raw_embed_audio, raw_embed_audio)
t.load()
# %%time
# t.build(10) # 10 trees
u = AnnoyIndex(f, "euclidian")
t.load(fn) # super fast, will just mmap the file
queryIndex=1000
a=(t.get_nns_by_item(queryIndex, 100)) # will find the 100 nearest neighbors
b=(t.get_nns_by_item(queryIndex+1, 100))
a=np.array(a)
b=np.array(b)
numbers=set()
for i in range(10):
numbers=numbers.union(set(a).intersection(set(b+i)))
numbers=numbers.union(set(a).intersection(set(b-i)))
len(numbers)
f=128
u = AnnoyIndex(f, 'euclidean')
u.load('/scratch/enis/data/nna/realMerged/prudhoe26.ann') # super fast, will just mmap the file
u.get_nns_by_item(1, 4)
u.get_n_items()
allA=[None]*numberItems
for i in range(numberItems):
a=u.get_distance(i, index)
%%time
for m in range(10):
# allA=[None]*concat_embeds.shape[0]
for i in range(concat_embeds.shape[0]):
a=u.get_distance(i, 0)
%%time
import multiprocessing
def worker(index,numberItems):
allA=[None]*numberItems
for i in range(numberItems):
a=u.get_distance(i, index)
return None
jobs = []
for index in range(10):
numberItems=u.get_n_items()
p = multiprocessing.Process(target=worker,args=(index,numberItems))
jobs.append(p)
p.start()
for p in jobs:
p.join()
allA[4]
results[0]
%%time
# one sample/query
# res=paired_distances_broadcast(raw_embed_audio2,concat_embeds2)
# res.shape
#
# 10 queries
queryCount=10
startSecond=0
excerptLen=1
gamma=1/512
startIndex=int(startSecond/excerptLen)
endIndex=startIndex+queryCount
results = pairwise_distances(raw_embed_audio[startIndex:10,:], concat_embeds)
# results=np.exp(-results*gamma)
%%time
results=np.exp(-results*gamma)
# I might have to use this method, if I want to get maximum without replacement
# %%time
# stepSize=1
# windowSize=10
# resultsMax=[]
# resultsAvg=[]
# windowMax=np.max(results2,axis=0)
# for i in range(0,results2.shape[1],stepSize):
# # windowElements=(results2[:,i:(i+windowSize+1)])
# # colMax=np.max(windowElements,axis=0)
# resultAvg=np.average(colMax)
# resultsAvg.append(resultAvg)
# # resultsMax.append(windowMax)
```
### Run sliding window over the Matrix
```
%%time
windowSize=10
windowMax=np.max(results,axis=0)
windowMean=pd.Series(windowMax).rolling(window=windowSize).mean().iloc[windowSize-1:].values
%%time
windowSize=10
windowMax=np.max(resultMatrix,axis=0)
windowMean=pd.Series(windowMax).rolling(window=windowSize).mean().iloc[windowSize-1:].values
```
### Look into stats of results
```
from scipy import stats
stats.describe(res)
res=windowMean[:]
```
### find kth highest scores (similarity of queries)
```
# arr = np.array([1, 3, 2, 4, 5,6,7,8,8,1,1,9])
kth=250
ind = np.argpartition(res, -kth)[-kth:]
sortedbyScore=sorted(list(zip(res[ind],ind)),reverse=True)
```
#### Highest scores might be consequtive, merging these as one result helps decrease amount of files to listen
```
# find consequtive points that has high value
theta=5
sortedbyInd=sorted(list(zip(res[ind],ind)),key=lambda x:x[1])
previous=sortedbyInd[0][1]
count=0
series=[]
starts=[]
for score,index in sortedbyInd:
if index<=previous+theta:
series.append((score,index))
else:
starts.append(series)
series=[]
series.append((score,index))
previous=index
# sorted(starts,key=lambda x:x[1],reverse=True)
print("total periods",len(starts))
previous=0
print("short distances")
for score_indexes in starts:
if (score_indexes[-1][1]-score_indexes[0][1])>30:
print(score_indexes[0][1],score_indexes[-1][1])
```
### generating figures for similarity scores
```
res.shape,rowN,res.shape[0]/rowN
import matplotlib.pyplot as plt
import matplotlib.ticker as plticker
minY,maxY=min(res),max(res)
exp_name="slidingWindow_AverageOfMax_short20k"
name=f"Euclidian Similarity {exp_name}"
nrows=res.shape[0]//rowN
print(nrows)
fig, ax = plt.subplots(nrows=nrows,figsize=(200, 20*nrows))
for i,axes in enumerate(ax):
axes.plot(res[i*(rowN):(i+1)*(rowN)])
axes.set_ylim(minY,maxY)
axes.grid(True)
ax[0].set_ylabel(f"{name}",fontsize=32)
# plt.grid(True)
# loc = plticker.MultipleLocator(base=5000) # this locator puts ticks at regular intervals
# ax.xaxis.set_major_locator(loc)
plt.tight_layout()
fig.show()
filename=name.replace(" ","_")
fig.savefig(f"/home/enis/projects/nna/results/vis/nearestNeighbour/{filename}.png")
excerptLen=10
exp_name="Concat10Second"
concat_embeds2=concat_embeds.reshape(-1,1280)
raw_embed_audio2=raw_embed_audio.reshape(-1,1280)
raw_embed_audio2=raw_embed_audio2[2:3,:]
rowN=10000
res=paired_distances_broadcast(raw_embed_audio2,concat_embeds2)
res.shape
import matplotlib.pyplot as plt
import matplotlib.ticker as plticker
name=f"Euclidian Distance {exp_name}"
nrows=res.shape[0]//rowN
fig, ax = plt.subplots(nrows=nrows,figsize=(200, 20*nrows))
for i,axes in enumerate(ax):
axes.plot(res[i*(rowN):(i+1)*(rowN)])
ax[0].set_ylabel(f"{name}",fontsize=32)
plt.grid(True)
# loc = plticker.MultipleLocator(base=5000) # this locator puts ticks at regular intervals
# ax.xaxis.set_major_locator(loc)
plt.tight_layout()
# fig.show()
filename=name.replace(" ","_")
fig.savefig(f"/home/enis/projects/nna/results/vis/nearestNeighbour/{filename}.png")
len(all_embeddings),excerptLen
# sorted([sorted(start) for start in starts])
split_folder="/home/enis/projects/nna/data/nearestNeighbours/S4A10255_20190507_073000-1808seconds/slidingWindow_AverageOfMax/"
excerptLen=1
# for indexes in (starts):
# score,index=(indexes[len(indexes)//2])
# print(score)
# print(embedIndex2fileSecond(index,embeds,all_embeddings,excerptLen=1))
for score,index in (sortedbyScore):
# score,index=(indexes[len(indexes)//2])
print(score)
print(embedIndex2fileSecond(index,embeds,all_embeddings,excerptLen=1))
# indexes=[indexes[len(indexes)//2] for indexes,count in (starts)]
# # # indexes
# clipAddBip(indexes,exp_name,split_folder,embeds,all_embeddings,excerptLen=10,reductionLen=1)
kdt = KDTree(concat_embeds2, leaf_size=30, metric='euclidean')
distances,indexes=kdt.query(raw_embed_audio2, k=10, return_distance=True)
allResults=[ (distance,index) for index,distance in (zip(indexes.flatten(),distances.flatten()))]
allResults.sort()
# distances,indexes=allResults
# distances
def embedIndex2fileSecond(index,embeds,all_embeddings,excerptLen=1):
alist=[]
countStart=0
embed_count=index*excerptLen
for i,em in enumerate(embeds):
countEnd=countStart+(em.shape[0])
if countStart<=embed_count and embed_count<=countEnd:
startSecond=embed_count-countStart
# print(i,countStart,countEnd,"start second:",startSecond)
filename=("/tank/"+"/".join(all_embeddings[i].split("/")[3:-2])+"/"+all_embeddings[i].split("/")[-2][:-4]+".flac")
# alist.append((filename,startSecond))
return((filename,startSecond))
# break
countStart=countEnd
split_folder="/home/enis/projects/nna/data/nearestNeighbours/"
def clipAddBip(allIndexes,exp_name,split_folder,embeds,all_embeddings,excerptLen,reductionLen):
# embed_count=823804
beep_wav="/home/enis/projects/nna/data/beep.wav"
beep_wav = AudioSegment.from_file(beep_wav)
beep_wav=beep_wav-30
split_folder=(split_folder+exp_name)
Path(split_folder).mkdir(parents=True, exist_ok=True)
for index in allIndexes[:]:
(filename,startSecond)=embedIndex2fileSecond(index,embeds,all_embeddings,excerptLen=reductionLen)
endSecond=startSecond+excerptLen
print(filename,startSecond)
input_file=filename
buffer=10
start_time="{}.{}".format((startSecond-buffer)//60,(startSecond-buffer)%60)
end_time="{}.{}".format((endSecond+buffer)//60,(endSecond+buffer)%60)
outputSuffix=".wav"
output_file = splitmp3(input_file,split_folder,start_time,end_time,depth=1,backend="ffmpeg",outputSuffix=outputSuffix)
other_wav=AudioSegment.from_file(output_file)
start,end=(buffer*1000),((buffer+excerptLen)*1000)
result_wav=other_wav[:start]+(beep_wav)+other_wav[start:end]+beep_wav+other_wav[end:]
o_file=result_wav.export(output_file)
# o_file=result_wav.export(output_file)
# len(other_wav)
# beep_wav="/home/enis/projects/nna/data/beep.wav"
# other_wav="./S4A10279_20190507_033000_4008-4068.wav"
# beep_wav = AudioSegment.from_file(beep_wav)
# beep_wav=beep_wav-30
# other_wav=AudioSegment.from_file(other_wav)
# other_wav[:start]+(beep_wav-30)+other_wav[start:end]+
# res=(beep_wav-30)+other_wav[:1000]
# input_file='/tank/data/nna/real/anwr/37/2019/S4A10279_20190605_091602.flac'
# split_folder="./"
# start_time="2.12"
# end_time="3.12"
# outputSuffix=".wav"
# a=splitmp3(input_file,split_folder,start_time,end_time,depth=1,backend="ffmpeg",outputSuffix=outputSuffix)
# !ffmpeg -i '/tank/data/nna/real/anwr/37/2019/S4A10279_20190605_091602.flac' -ss 304 -to 364 -c copy S4A10279_20190605_091602_304-364.wav
# !ffmpeg -i '/tank/data/nna/real/anwr/37/2019/S4A10279_20190604_234602.flac' -ss 650 -to 700 -c copy S4A10279_20190604_234602_650-700.wav
# !ffmpeg -i '/tank/data/nna/real/anwr/37/2019/S4A10279_20190509_073000.flac' -ss 4149 -to 4209 -c copy S4A10279_20190509_073000_4149-4209.wav
# !ffmpeg -i '/tank/data/nna/real/anwr/37/2019/S4A10279_20190530_214602.flac' -ss 3260 -to 3320 -c copy S4A10279_20190530_214602_3260-3320.wav
# !ffmpeg -i '/tank/data/nna/real/anwr/37/2019/S4A10279_20190507_033000.flac' -ss 4008 -to 4068 -c copy S4A10279_20190507_033000_4008-4068.wav
# !ffmpeg -i '/tank/data/nna/real/anwr/37/2019/S4A10279_20190507_033000.flac' -ss 320 -to 380 -c copy segment8.wav
# !ffmpeg -i '/tank/data/nna/real/anwr/37/2019/S4A10279_20190608_020000.flac' -ss 222 -to 282 -c copy S4A10279_20190608_020000_222-282.wav
# !mp3splt -d "./" -f -D /tank/data/nna/real/anwr/37/2019/S4A10279_20190517_130000.flac 15.27 16.27
# ffmpeg -i '/tank/data/nna/real/anwr/37/2019/S4A10279_20190605_091602.flac' -ss 304 -to 364 -c copy S4A10279_20190605_091602_304-364.wav
# !ls/tank/data/nna/real/anwr/37/2019/S4A10279_20190603_194602.flac
```
| github_jupyter |
## RIHAD VARIAWA, Data Scientist - Who has fun LEARNING, EXPLORING & GROWING
<h1>Module 5: Model Evaluation and Refinement</h1>
We have built models and made predictions of vehicle prices. Now we will determine how accurate these predictions are.
<h1>Table of content</h1>
<ul>
<li><a href="#ref1">Model Evaluation </a></li>
<li><a href="#ref2">Over-fitting, Under-fitting and Model Selection </a></li>
<li><a href="#ref3">Ridge Regression </a></li>
<li><a href="#ref4">Grid Search</a></li>
</ul>
```
import pandas as pd
import numpy as np
# Import clean data
path = 'https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DA0101EN/module_5_auto.csv'
df = pd.read_csv(path)
df.to_csv('module_5_auto.csv')
```
First lets only use numeric data
```
df=df._get_numeric_data()
df.head()
```
Libraries for plotting
```
%%capture
! pip install ipywidgets
from IPython.display import display
from IPython.html import widgets
from IPython.display import display
from ipywidgets import interact, interactive, fixed, interact_manual
```
<h2>Functions for plotting</h2>
```
def DistributionPlot(RedFunction, BlueFunction, RedName, BlueName, Title):
width = 12
height = 10
plt.figure(figsize=(width, height))
ax1 = sns.distplot(RedFunction, hist=False, color="r", label=RedName)
ax2 = sns.distplot(BlueFunction, hist=False, color="b", label=BlueName, ax=ax1)
plt.title(Title)
plt.xlabel('Price (in dollars)')
plt.ylabel('Proportion of Cars')
plt.show()
plt.close()
def PollyPlot(xtrain, xtest, y_train, y_test, lr,poly_transform):
width = 12
height = 10
plt.figure(figsize=(width, height))
#training data
#testing data
# lr: linear regression object
#poly_transform: polynomial transformation object
xmax=max([xtrain.values.max(), xtest.values.max()])
xmin=min([xtrain.values.min(), xtest.values.min()])
x=np.arange(xmin, xmax, 0.1)
plt.plot(xtrain, y_train, 'ro', label='Training Data')
plt.plot(xtest, y_test, 'go', label='Test Data')
plt.plot(x, lr.predict(poly_transform.fit_transform(x.reshape(-1, 1))), label='Predicted Function')
plt.ylim([-10000, 60000])
plt.ylabel('Price')
plt.legend()
```
<h1 id="ref1">Part 1: Training and Testing</h1>
<p>An important step in testing your model is to split your data into training and testing data. We will place the target data <b>price</b> in a separate dataframe <b>y</b>:</p>
```
y_data = df['price']
```
drop price data in x data
```
x_data=df.drop('price',axis=1)
```
Now we randomly split our data into training and testing data using the function <b>train_test_split</b>.
```
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(x_data, y_data, test_size=0.15, random_state=1)
print("number of test samples :", x_test.shape[0])
print("number of training samples:",x_train.shape[0])
```
The <b>test_size</b> parameter sets the proportion of data that is split into the testing set. In the above, the testing set is set to 10% of the total dataset.
<div class="alert alert-danger alertdanger" style="margin-top: 20px">
<h1> Question #1):</h1>
<b>Use the function "train_test_split" to split up the data set such that 40% of the data samples will be utilized for testing, set the parameter "random_state" equal to zero. The output of the function should be the following: "x_train_1" , "x_test_1", "y_train_1" and "y_test_1".</b>
</div>
```
# Write your code below and press Shift+Enter to execute
x_train1, x_test1, y_train1, y_test1 = train_test_split(x_data, y_data, test_size=0.4, random_state=0)
print("number of test samples :", x_test1.shape[0])
print("number of training samples:",x_train1.shape[0])
```
Double-click <b>here</b> for the solution.
<!-- The answer is below:
x_train1, x_test1, y_train1, y_test1 = train_test_split(x_data, y_data, test_size=0.4, random_state=0)
print("number of test samples :", x_test1.shape[0])
print("number of training samples:",x_train1.shape[0])
-->
Let's import <b>LinearRegression</b> from the module <b>linear_model</b>.
```
from sklearn.linear_model import LinearRegression
```
We create a Linear Regression object:
```
lre=LinearRegression()
```
we fit the model using the feature horsepower
```
lre.fit(x_train[['horsepower']], y_train)
```
Let's Calculate the R^2 on the test data:
```
lre.score(x_test[['horsepower']], y_test)
```
we can see the R^2 is much smaller using the test data.
```
lre.score(x_train[['horsepower']], y_train)
```
<div class="alert alert-danger alertdanger" style="margin-top: 20px">
<h1> Question #2): </h1>
<b>
Find the R^2 on the test data using 90% of the data for training data
</b>
</div>
```
# Write your code below and press Shift+Enter to execute
x_train1, x_test1, y_train1, y_test1 = train_test_split(x_data, y_data, test_size=0.1, random_state=0)
lre.fit(x_train1[['horsepower']],y_train1)
lre.score(x_test1[['horsepower']],y_test1)
```
Double-click <b>here</b> for the solution.
<!-- The answer is below:
x_train1, x_test1, y_train1, y_test1 = train_test_split(x_data, y_data, test_size=0.1, random_state=0)
lre.fit(x_train1[['horsepower']],y_train1)
lre.score(x_test1[['horsepower']],y_test1)
-->
Sometimes you do not have sufficient testing data; as a result, you may want to perform Cross-validation. Let's go over several methods that you can use for Cross-validation.
<h2>Cross-validation Score</h2>
Lets import <b>model_selection</b> from the module <b>cross_val_score</b>.
```
from sklearn.model_selection import cross_val_score
```
We input the object, the feature in this case ' horsepower', the target data (y_data). The parameter 'cv' determines the number of folds; in this case 4.
```
Rcross = cross_val_score(lre, x_data[['horsepower']], y_data, cv=4)
```
The default scoring is R^2; each element in the array has the average R^2 value in the fold:
```
Rcross
```
We can calculate the average and standard deviation of our estimate:
```
print("The mean of the folds are", Rcross.mean(), "and the standard deviation is" , Rcross.std())
```
We can use negative squared error as a score by setting the parameter 'scoring' metric to 'neg_mean_squared_error'.
```
-1 * cross_val_score(lre,x_data[['horsepower']], y_data,cv=4,scoring='neg_mean_squared_error')
```
<div class="alert alert-danger alertdanger" style="margin-top: 20px">
<h1> Question #3): </h1>
<b>
Calculate the average R^2 using two folds, find the average R^2 for the second fold utilizing the horsepower as a feature :
</b>
</div>
```
# Write your code below and press Shift+Enter to execute
Rc=cross_val_score(lre,x_data[['horsepower']], y_data,cv=2)
Rc[1]
```
Double-click <b>here</b> for the solution.
<!-- The answer is below:
Rc=cross_val_score(lre,x_data[['horsepower']], y_data,cv=2)
Rc[1]
-->
You can also use the function 'cross_val_predict' to predict the output. The function splits up the data into the specified number of folds, using one fold to get a prediction while the rest of the folds are used as test data. First import the function:
```
from sklearn.model_selection import cross_val_predict
```
We input the object, the feature in this case <b>'horsepower'</b> , the target data <b>y_data</b>. The parameter 'cv' determines the number of folds; in this case 4. We can produce an output:
```
yhat = cross_val_predict(lre,x_data[['horsepower']], y_data,cv=4)
yhat[0:5]
```
<h1 id="ref2">Part 2: Overfitting, Underfitting and Model Selection</h1>
<p>It turns out that the test data sometimes referred to as the out of sample data is a much better measure of how well your model performs in the real world. One reason for this is overfitting; let's go over some examples. It turns out these differences are more apparent in Multiple Linear Regression and Polynomial Regression so we will explore overfitting in that context.</p>
Let's create Multiple linear regression objects and train the model using <b>'horsepower'</b>, <b>'curb-weight'</b>, <b>'engine-size'</b> and <b>'highway-mpg'</b> as features.
```
lr = LinearRegression()
lr.fit(x_train[['horsepower', 'curb-weight', 'engine-size', 'highway-mpg']], y_train)
```
Prediction using training data:
```
yhat_train = lr.predict(x_train[['horsepower', 'curb-weight', 'engine-size', 'highway-mpg']])
yhat_train[0:5]
```
Prediction using test data:
```
yhat_test = lr.predict(x_test[['horsepower', 'curb-weight', 'engine-size', 'highway-mpg']])
yhat_test[0:5]
```
Let's perform some model evaluation using our training and testing data separately. First we import the seaborn and matplotlibb library for plotting.
```
import matplotlib.pyplot as plt
%matplotlib inline
import seaborn as sns
```
Let's examine the distribution of the predicted values of the training data.
```
Title = 'Distribution Plot of Predicted Value Using Training Data vs Training Data Distribution'
DistributionPlot(y_train, yhat_train, "Actual Values (Train)", "Predicted Values (Train)", Title)
```
Figure 1: Plot of predicted values using the training data compared to the training data.
So far the model seems to be doing well in learning from the training dataset. But what happens when the model encounters new data from the testing dataset? When the model generates new values from the test data, we see the distribution of the predicted values is much different from the actual target values.
```
Title='Distribution Plot of Predicted Value Using Test Data vs Data Distribution of Test Data'
DistributionPlot(y_test,yhat_test,"Actual Values (Test)","Predicted Values (Test)",Title)
```
Figur 2: Plot of predicted value using the test data compared to the test data.
<p>Comparing Figure 1 and Figure 2; it is evident the distribution of the test data in Figure 1 is much better at fitting the data. This difference in Figure 2 is apparent where the ranges are from 5000 to 15 000. This is where the distribution shape is exceptionally different. Let's see if polynomial regression also exhibits a drop in the prediction accuracy when analysing the test dataset.</p>
```
from sklearn.preprocessing import PolynomialFeatures
```
<h4>Overfitting</h4>
<p>Overfitting occurs when the model fits the noise, not the underlying process. Therefore when testing your model using the test-set, your model does not perform as well as it is modelling noise, not the underlying process that generated the relationship. Let's create a degree 5 polynomial model.</p>
Let's use 55 percent of the data for testing and the rest for training:
```
x_train, x_test, y_train, y_test = train_test_split(x_data, y_data, test_size=0.45, random_state=0)
```
We will perform a degree 5 polynomial transformation on the feature <b>'horse power'</b>.
```
pr = PolynomialFeatures(degree=5)
x_train_pr = pr.fit_transform(x_train[['horsepower']])
x_test_pr = pr.fit_transform(x_test[['horsepower']])
pr
```
Now let's create a linear regression model "poly" and train it.
```
poly = LinearRegression()
poly.fit(x_train_pr, y_train)
```
We can see the output of our model using the method "predict." then assign the values to "yhat".
```
yhat = poly.predict(x_test_pr)
yhat[0:5]
```
Let's take the first five predicted values and compare it to the actual targets.
```
print("Predicted values:", yhat[0:4])
print("True values:", y_test[0:4].values)
```
We will use the function "PollyPlot" that we defined at the beginning of the lab to display the training data, testing data, and the predicted function.
```
PollyPlot(x_train[['horsepower']], x_test[['horsepower']], y_train, y_test, poly,pr)
```
Figur 4 A polynomial regression model, red dots represent training data, green dots represent test data, and the blue line represents the model prediction.
We see that the estimated function appears to track the data but around 200 horsepower, the function begins to diverge from the data points.
R^2 of the training data:
```
poly.score(x_train_pr, y_train)
```
R^2 of the test data:
```
poly.score(x_test_pr, y_test)
```
We see the R^2 for the training data is 0.5567 while the R^2 on the test data was -29.87. The lower the R^2, the worse the model, a Negative R^2 is a sign of overfitting.
Let's see how the R^2 changes on the test data for different order polynomials and plot the results:
```
Rsqu_test = []
order = [1, 2, 3, 4]
for n in order:
pr = PolynomialFeatures(degree=n)
x_train_pr = pr.fit_transform(x_train[['horsepower']])
x_test_pr = pr.fit_transform(x_test[['horsepower']])
lr.fit(x_train_pr, y_train)
Rsqu_test.append(lr.score(x_test_pr, y_test))
plt.plot(order, Rsqu_test)
plt.xlabel('order')
plt.ylabel('R^2')
plt.title('R^2 Using Test Data')
plt.text(3, 0.75, 'Maximum R^2 ')
```
We see the R^2 gradually increases until an order three polynomial is used. Then the R^2 dramatically decreases at four.
The following function will be used in the next section; please run the cell.
```
def f(order, test_data):
x_train, x_test, y_train, y_test = train_test_split(x_data, y_data, test_size=test_data, random_state=0)
pr = PolynomialFeatures(degree=order)
x_train_pr = pr.fit_transform(x_train[['horsepower']])
x_test_pr = pr.fit_transform(x_test[['horsepower']])
poly = LinearRegression()
poly.fit(x_train_pr,y_train)
PollyPlot(x_train[['horsepower']], x_test[['horsepower']], y_train,y_test, poly, pr)
```
The following interface allows you to experiment with different polynomial orders and different amounts of data.
```
interact(f, order=(0, 6, 1), test_data=(0.05, 0.95, 0.05))
```
<div class="alert alert-danger alertdanger" style="margin-top: 20px">
<h1> Question #4a):</h1>
<b>We can perform polynomial transformations with more than one feature. Create a "PolynomialFeatures" object "pr1" of degree two?</b>
</div>
Double-click <b>here</b> for the solution.
<!-- The answer is below:
pr1=PolynomialFeatures(degree=2)
-->
<div class="alert alert-danger alertdanger" style="margin-top: 20px">
<h1> Question #4b): </h1>
<b>
Transform the training and testing samples for the features 'horsepower', 'curb-weight', 'engine-size' and 'highway-mpg'. Hint: use the method "fit_transform"
?</b>
</div>
Double-click <b>here</b> for the solution.
<!-- The answer is below:
x_train_pr1=pr.fit_transform(x_train[['horsepower', 'curb-weight', 'engine-size', 'highway-mpg']])
x_test_pr1=pr.fit_transform(x_test[['horsepower', 'curb-weight', 'engine-size', 'highway-mpg']])
-->
<!-- The answer is below:
x_train_pr1=pr.fit_transform(x_train[['horsepower', 'curb-weight', 'engine-size', 'highway-mpg']])
x_test_pr1=pr.fit_transform(x_test[['horsepower', 'curb-weight', 'engine-size', 'highway-mpg']])
-->
<div class="alert alert-danger alertdanger" style="margin-top: 20px">
<h1> Question #4c): </h1>
<b>
How many dimensions does the new feature have? Hint: use the attribute "shape"
</b>
</div>
Double-click <b>here</b> for the solution.
<!-- The answer is below:
There are now 15 features: x_train_pr1.shape
-->
<div class="alert alert-danger alertdanger" style="margin-top: 20px">
<h1> Question #4d): </h1>
<b>
Create a linear regression model "poly1" and train the object using the method "fit" using the polynomial features?</b>
</div>
Double-click <b>here</b> for the solution.
<!-- The answer is below:
poly1=linear_model.LinearRegression().fit(x_train_pr1,y_train)
-->
<div class="alert alert-danger alertdanger" style="margin-top: 20px">
<h1> Question #4e): </h1>
<b>Use the method "predict" to predict an output on the polynomial features, then use the function "DistributionPlot" to display the distribution of the predicted output vs the test data?</b>
</div>
Double-click <b>here</b> for the solution.
<!-- The answer is below:
yhat_test1=poly1.predict(x_train_pr1)
Title='Distribution Plot of Predicted Value Using Test Data vs Data Distribution of Test Data'
DistributionPlot(y_test, yhat_test1, "Actual Values (Test)", "Predicted Values (Test)", Title)
-->
<div class="alert alert-danger alertdanger" style="margin-top: 20px">
<h1> Question #4f): </h1>
<b>Use the distribution plot to determine the two regions were the predicted prices are less accurate than the actual prices.</b>
</div>
Double-click <b>here</b> for the solution.
<!-- The answer is below:
The predicted value is lower than actual value for cars where the price $ 10,000 range, conversely the predicted price is larger than the price cost in the $30, 000 to $40,000 range. As such the model is not as accurate in these ranges .
-->
<img src = "https://ibm.box.com/shared/static/c35ipv9zeanu7ynsnppb8gjo2re5ugeg.png" width = 700, align = "center">
<h2 id="ref3">Part 3: Ridge regression</h2>
In this section, we will review Ridge Regression we will see how the parameter Alfa changes the model. Just a note here our test data will be used as validation data.
Let's perform a degree two polynomial transformation on our data.
```
pr=PolynomialFeatures(degree=2)
x_train_pr=pr.fit_transform(x_train[['horsepower', 'curb-weight', 'engine-size', 'highway-mpg','normalized-losses','symboling']])
x_test_pr=pr.fit_transform(x_test[['horsepower', 'curb-weight', 'engine-size', 'highway-mpg','normalized-losses','symboling']])
```
Let's import <b>Ridge</b> from the module <b>linear models</b>.
```
from sklearn.linear_model import Ridge
```
Let's create a Ridge regression object, setting the regularization parameter to 0.1
```
RigeModel=Ridge(alpha=0.1)
```
Like regular regression, you can fit the model using the method <b>fit</b>.
```
RigeModel.fit(x_train_pr, y_train)
```
Similarly, you can obtain a prediction:
```
yhat = RigeModel.predict(x_test_pr)
```
Let's compare the first five predicted samples to our test set
```
print('predicted:', yhat[0:4])
print('test set :', y_test[0:4].values)
```
We select the value of Alfa that minimizes the test error, for example, we can use a for loop.
```
Rsqu_test = []
Rsqu_train = []
dummy1 = []
ALFA = 10 * np.array(range(0,1000))
for alfa in ALFA:
RigeModel = Ridge(alpha=alfa)
RigeModel.fit(x_train_pr, y_train)
Rsqu_test.append(RigeModel.score(x_test_pr, y_test))
Rsqu_train.append(RigeModel.score(x_train_pr, y_train))
```
We can plot out the value of R^2 for different Alphas
```
width = 12
height = 10
plt.figure(figsize=(width, height))
plt.plot(ALFA,Rsqu_test, label='validation data ')
plt.plot(ALFA,Rsqu_train, 'r', label='training Data ')
plt.xlabel('alpha')
plt.ylabel('R^2')
plt.legend()
```
Figure 6:The blue line represents the R^2 of the test data, and the red line represents the R^2 of the training data. The x-axis represents the different values of Alfa
The red line in figure 6 represents the R^2 of the test data, as Alpha increases the R^2 decreases; therefore as Alfa increases the model performs worse on the test data. The blue line represents the R^2 on the validation data, as the value for Alfa increases the R^2 decreases.
<div class="alert alert-danger alertdanger" style="margin-top: 20px">
<h1> Question #5): </h1>
Perform Ridge regression and calculate the R^2 using the polynomial features, use the training data to train the model and test data to test the model. The parameter alpha should be set to 10.
</div>
```
# Write your code below and press Shift+Enter to execute
```
Double-click <b>here</b> for the solution.
<!-- The answer is below:
RigeModel = Ridge(alpha=0)
RigeModel.fit(x_train_pr, y_train)
RigeModel.score(x_test_pr, y_test)
-->
<h2 id="ref4">Part 4: Grid Search</h2>
The term Alfa is a hyperparameter, sklearn has the class <b>GridSearchCV</b> to make the process of finding the best hyperparameter simpler.
Let's import <b>GridSearchCV</b> from the module <b>model_selection</b>.
```
from sklearn.model_selection import GridSearchCV
```
We create a dictionary of parameter values:
```
parameters1= [{'alpha': [0.001,0.1,1, 10, 100, 1000, 10000, 100000, 100000]}]
parameters1
```
Create a ridge regions object:
```
RR=Ridge()
RR
```
Create a ridge grid search object
```
Grid1 = GridSearchCV(RR, parameters1,cv=4)
```
Fit the model
```
Grid1.fit(x_data[['horsepower', 'curb-weight', 'engine-size', 'highway-mpg']], y_data)
```
The object finds the best parameter values on the validation data. We can obtain the estimator with the best parameters and assign it to the variable BestRR as follows:
```
BestRR=Grid1.best_estimator_
BestRR
```
We now test our model on the test data
```
BestRR.score(x_test[['horsepower', 'curb-weight', 'engine-size', 'highway-mpg']], y_test)
```
<div class="alert alert-danger alertdanger" style="margin-top: 20px">
<h1> Question #6): </h1>
Perform a grid search for the alpha parameter and the normalization parameter, then find the best values of the parameters
</div>
```
# Write your code below and press Shift+Enter to execute
```
Double-click <b>here</b> for the solution.
<!-- The answer is below:
parameters2= [{'alpha': [0.001,0.1,1, 10, 100, 1000,10000,100000,100000],'normalize':[True,False]} ]
Grid2 = GridSearchCV(Ridge(), parameters2,cv=4)
Grid2.fit(x_data[['horsepower', 'curb-weight', 'engine-size', 'highway-mpg']],y_data)
Grid2.best_estimator_
-->
<h1>Thank you for completing this notebook!</h1>
<div class="alert alert-block alert-info" style="margin-top: 20px">
<h2>Get IBM Watson Studio free of charge!</h2>
<p><a href="http://cocl.us/NotebooksPython101bottom"><img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DA0101EN/Images/BottomAd.png" width="750" align="center"></a></p>
</div>
<h3>About the Authors:</h3>
This notebook was written by <a href="https://www.linkedin.com/in/mahdi-noorian-58219234/" target="_blank">Mahdi Noorian PhD</a>, <a href="https://www.linkedin.com/in/joseph-s-50398b136/" target="_blank">Joseph Santarcangelo</a>, Bahare Talayian, Eric Xiao, Steven Dong, Parizad, Hima Vsudevan and <a href="https://www.linkedin.com/in/fiorellawever/" target="_blank">Fiorella Wenver</a>.
<p><a href="https://www.linkedin.com/in/joseph-s-50398b136/" target="_blank">Joseph Santarcangelo</a> is a Data Scientist at IBM, and holds a PhD in Electrical Engineering. His research focused on using Machine Learning, Signal Processing, and Computer Vision to determine how videos impact human cognition. Joseph has been working for IBM since he completed his PhD.</p>
<hr>
<p>Copyright © 2018 IBM Developer Skills Network. This notebook and its source code are released under the terms of the <a href="https://cognitiveclass.ai/mit-license/">MIT License</a>.</p>
| github_jupyter |
```
# -*- coding: utf-8 -*-
# This work is part of the Core Imaging Library (CIL) developed by CCPi
# (Collaborative Computational Project in Tomographic Imaging), with
# substantial contributions by UKRI-STFC and University of Manchester.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
# http://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# Copyright 2021 UKRI-STFC
# Authored by: Evangelos Papoutsellis (UKRI-STFC)
# Gemma Fardell (UKRI-STFC)
```
<h1><center>Colour Processing </center></h1>
In this notebook, we present how to **denoise** and **inpaint** our first **multichannel** data using CIL, i.e., a data with only 3 channels that contains information from the **Red**, **Green** and **Blue** bands. We start by loading a colour image from CIL.
```
# import dataexample that contains different colour images
from cil.utilities import dataexample, noise
# import other libraries
import matplotlib.pyplot as plt
from mpl_toolkits.axes_grid1 import AxesGrid
from cil.utilities.display import show2D
import numpy as np
# Load Rainbow image
data = dataexample.RAINBOW.get(size=(500,500), scale=(0,1))
data.reorder(['horizontal_y', 'horizontal_x','channel'])
```
## Show colour image and RGB channels
```
plt.figure(figsize=(10,10))
plt.imshow(data.array)
plt.title("Colour Image")
plt.show()
show2D( data, slice_list=[('channel',0),('channel',1),('channel',2)], \
title=["Red","Green","Blue"], origin="upper", num_cols=3)
```
<h1><center>Imaging Model </center></h1>
Let $u:\Omega\subset\mathbb{R}^{N\times M}\rightarrow\mathbb{R}^{N\times M\times3}$ a colour image that depicts a real _perfect_ scene (the unknown). Typically, we assume that $u$ has been transformed through a continuous and linear operation $\mathcal{L}$ (**forward operator**). Additionally, we have a noisy component $\eta$ that usually follows a certain distribution, e.g., **Gaussian** , **Salt and Pepper (Impluse)**. The **Imaging model** is defined as
<a id ="eq1"></a>
$$
\begin{equation}
u_{0} = \mathcal{L}u + \eta\,.
\tag{1}
\end{equation}
$$
* **Image Denoising:** $\mathcal{L}$ is the Identity operator and we are trying to remove the noise from $u_{0}$ in order to reconstruct $u$.
<table><tr><td><img src='denoising_fig1.png'>
</td><td><img src='inpainting_fig2.png'></td></tr></table>
<a id ="chi_func"></a>
* **Image Inpainting:** $\mathcal{L}=\mathcal{X}_{\Omega\setminus D}$ is the characteristic function defined as
$$\mathcal{X}_{\Omega\setminus \mathcal{D}}(x) =
\begin{cases}
1, & x\in \Omega\setminus D\\
0, & \mbox{otherwise}
\end{cases},
$$
where $\mathcal{D}$ is a subdomain of $\Omega$ (**inpainting domain**). In the inpainting domain there is no data information available and we are trying to reconstruct $u$ based on the information provided from the known region of $u_{0}$.
<table><tr><td><img src='inpainting_fig1.png'>
</td><td><img src='inpainting_fig2.png'></td></tr></table>
In this notebook, we will consider the cases of
* denoising a noisy image corrupted by additive Gaussian noise,
* inpainting + denoising a noisy image corrupted by Salt \& Pepper noise with missing text information.
<h1><center>Color Denoising </center></h1>
We solve the following minimisation problem to denoise our coloured image:
<a id="rof"></a>
$$
\begin{equation}
u^{*} = \underset{u}{\operatorname{argmin}} \frac{1}{2}\| b - u \|^{2}_{2} + \alpha\,\mathrm{VTV}(u)
\label{ROF}
\tag{1}
\end{equation},
$$
where the data $b$ is corrupted with Gaussian noise and $\mathrm{\textbf{VTV}}$ is the **Vectorial** extension of the classical Total variation regularisation for coloured images. We recall that the definition of the (isotropic) Total Variation, used for gray-valued images, is
$$
\mathrm{TV}(u) = \|Du\|_{2,1} = \sum_{i,j}^{M,N}\big(|(D_{y}u, D_{x}u)|_{2}\big)_{i,j} = \sum_{i,j}^{M,N} \big(\sqrt{ (D_{y}u_{k})^{2} + (D_{x}u_{k})^{2}}\big)_{i,j}.
$$
Now, for vector-valued images the gradient is $Du=(Du_{1}, Du_{2}, Du_{3})$, where for each **RGB** channels $k=1,2,3$, $Du_{k}:=(D_{y}u_{k}, D_{x}u_{k})$.
For this type of multichannel data, we can create different configurations on how the **colour channels**, the **derivatives** and the **image pixels** are correlated and under which norm. One generic approach for this regulariser is presented in [Duran et al](https://arxiv.org/pdf/1508.01308.pdf#page=8), where the **Collaborative Total variation** is introduced, i.e.,
$$
\|A\|_{p,q,r} := \bigg(\sum_{i=1}^{N}\quad\bigg(\sum_{j=1}^{M}\quad\bigg(\sum_{k=1}^{C} |A_{i,j,k}|^{p}\bigg)^{\frac{q}{p}}\quad\bigg)^{\frac{r}{q}}\quad\bigg)^{\frac{1}{r}}\quad .
$$
For simplicity, in this notebook, we will use the _Channelwise TV_ definition, namely,
$$
\begin{equation}
\mathrm{VTV}(u) := \|D u\|_{2,1} = \sum_{k=1}^{3}\sum_{i,j=1}^{M,N} (|Du_{k}|_{2})_{i,j} =
\sum_{k=1}^{3}\sum_{i,j=1}^{M,N} \big( \sqrt{ (D_{y}u_{k})^{2} + (D_{x}u_{k})^{2}}\big) = \sum_{k=1}^{3} \mathrm{TV}(u_{k}).
\label{tv_color}
\end{equation}
$$
The above definition corresponds to the $\ell_{2,1,1}$ (derivative, pixels, colour) Collaborative TV. This means that, an $\ell_{2}$ norm is applied for the **derivatives**, followed by an $\ell_{1}$ norm for the **pixels** of the image and a final $\ell_{1}$ norm for the three **channels**.
```
# Import Total variation
from cil.optimisation.functions import TotalVariation
```
## Load data and corrupt with gaussian noise
```
# Load Rainbow data
data = dataexample.RAINBOW.get(size=(500,500), scale=(0,1))
data.reorder(['horizontal_y', 'horizontal_x','channel'])
noisy_data = noise.gaussian(data, seed = 10, var = 0.005)
images = [data.as_array(), noisy_data.as_array(),
data.as_array()[:,:,0], noisy_data.as_array()[:,:,0],
data.as_array()[:,:,1], noisy_data.as_array()[:,:,1],
data.as_array()[:,:,2], noisy_data.as_array()[:,:,2]]
#create our custom colour maps for RGB images
from matplotlib.colors import LinearSegmentedColormap
colors = [(0, 0, 0), (1, 0, 0)] # first color is black, last is red
cm_r = LinearSegmentedColormap.from_list(
"Custom", colors, N=20)
colors = [(0, 0, 0), (0, 1, 0)] # first color is black, last is green
cm_g = LinearSegmentedColormap.from_list(
"Custom", colors, N=20)
colors = [(0, 0, 0), (0, 0, 1)] # first color is black, last is blue
cm_b = LinearSegmentedColormap.from_list(
"Custom", colors, N=20)
labels_y = ["Red", "Green","Blue"]
labels_x = ["Ground truth","Noisy data"]
# set fontszie xticks/yticks
plt.rcParams['xtick.labelsize']=15
plt.rcParams['ytick.labelsize']=15
fig = plt.figure(figsize=(20, 20))
grid = AxesGrid(fig, 111,
nrows_ncols=(4, 2),
axes_pad=0.1,
)
k = 0
for ax in grid:
img = ax.imshow(images[k])
if k==0 or k==1:
ax.set_title(labels_x[k],fontsize=25)
if k==2:
ax.set_ylabel(labels_y[0],fontsize=25)
img.set_cmap(cm_r)
if k==3:
img.set_cmap(cm_r)
if k==4:
ax.set_ylabel(labels_y[1],fontsize=25)
img.set_cmap(cm_g)
if k== 5:
img.set_cmap(cm_g)
if k==6:
ax.set_ylabel(labels_y[2],fontsize=25)
img.set_cmap(cm_b)
if k==7:
img.set_cmap(cm_b)
k+=1
plt.show()
```
We solve the above minimisation problem using the `proximal` method of the `TotalVariation` class that was used in previous notebooks. Recall, that given a function $f$, the _proximal operator of $f$_ is
$$\mathrm{prox}_{\tau f}(x) := \underset{u}{\operatorname{argmin}}\frac{1}{2}\|x-u\|_{2}^{2} + \tau f(u), \quad\mbox{for any } x.$$
This definition is exactly the same with the [above minimisation problem](#rof), if we replace $f$ by $\alpha\mathrm{VTV}$, $x$ with $b$ and $\tau=1.0$. Therefore, the _proximal operator of VTV at $b$_ is
$$\mathrm{prox}_{\tau (\alpha \mathrm{VTV})}(b)\, .$$
```
alpha = 0.05
TV = alpha * TotalVariation(max_iteration=500)
proxTV = TV.proximal(noisy_data, tau=1.0)
images = [data.as_array(), noisy_data.as_array(), proxTV.as_array(),
data.as_array()[:,:,0], noisy_data.as_array()[:,:,0], proxTV.as_array()[:,:,0],
data.as_array()[:,:,1], noisy_data.as_array()[:,:,1], proxTV.as_array()[:,:,1],
data.as_array()[:,:,2], noisy_data.as_array()[:,:,2], proxTV.as_array()[:,:,2]],
labels_x = ["Ground Truth", "Noisy Data", "TV denoising",
"(Red) Ground Truth", " (Red) Noisy Data", "(Red) TV denoising",
"(Green) Ground Truth", "(Green) Noisy Data", " (Green) TV denoising",
"(Blue) Ground Truth", "(Blue) Noisy Data", "(Blue) TV denoising"]
# set fontszie xticks/yticks
plt.rcParams['xtick.labelsize']=12
plt.rcParams['ytick.labelsize']=12
fig = plt.figure(figsize=(25, 25))
grid = AxesGrid(fig, 111,
nrows_ncols=(4, 3),
axes_pad=0.5,
cbar_mode='None'
)
k = 0
for ax in grid:
img = ax.imshow(images[0][k])
ax.set_title(labels_x[k],fontsize=25)
if (k >= 9):
img.set_cmap(cm_b)
elif (k >= 6):
img.set_cmap(cm_g)
elif (k >= 3):
img.set_cmap(cm_r)
k+=1
cbar = ax.cax.colorbar(img)
plt.show()
```
## Exercise : Use the PDHG algorithm to solve the above problem
The triplet $(K, \mathcal{F}, \mathcal{G})$ is defined as:
* $K = D \Longleftrightarrow$ `K = GradientOperator(noisy_data.geometry)` .
* $\mathcal{F}(z) = \alpha\,\|z\|_{2,1}\Longleftrightarrow$ `F = alpha * MixedL21Norm()` .
* $\mathcal{G}(u) = \frac{1}{2}\|b - u \|^{2}_{2}\, \Longleftrightarrow$ `G = 0.5 * L2NormSquared(b=noisy_data)` .
<h1><center>Colour Inpainting </center></h1>
Given an image where a specific region is unknown, the task of image inpainting is to recover the missing region $\mathcal{D}$ from the known part of the image $\Omega$. For this example, we will use the _rainbow image_, where we are trying to remove a repeated text (+ salt and pepper noise) from the image that represents the unknown domain $\mathcal{D}$.
## Create corrupted image
We use the *Pillow* library to add text in our image.
```
# Import libraries
import numpy as np
from PIL import Image, ImageFont, ImageDraw
# Numpy array
img_np = data.array
# Add text to the image
img_pil = Image.fromarray(np.uint8(img_np*255)).convert('RGB')
text = "\n This is a double \
\n rainbow \n"*3
draw = ImageDraw.Draw(img_pil)
font = ImageFont.truetype('FreeSerifBold.ttf', 50)
draw.text((0, 0), text, (0, 0, 0), font=font)
# Pillow image to numpy
im1 = np.array(img_pil)
# Rescale numpy array
img_np_rescale = im1/im1.max()
# Get image geometry
ig = data.geometry
# Create ImageData
data_with_text = ig.allocate()
data_with_text.fill(img_np_rescale)
# Show rainbow with text
plt.figure(figsize=(10,10))
plt.imshow(data_with_text.array)
plt.title("Rainbow with text")
plt.show()
```
## Create the mask representing the $\mathcal{D}$ region
```
# Mask that contains only text information
mask_boolean = (data_with_text-data).abs()==0
# Show rainbow with text
plt.figure(figsize=(10,10))
plt.imshow(mask_boolean[:,:,0])
plt.title("Mask: (Yellow=True, Blue=False)")
plt.show()
```
## Apply the mask for the RGB channels
Our mask plays the role of the characteristic function defined [above](#chi_func). Here, we use the `MaskOperator` that applies a mask to our image for all the red, green and blue channels using the `ChannelwiseOperator`.
```
from cil.optimisation.operators import MaskOperator, ChannelwiseOperator
mask = ig.subset(channel=0).allocate(True,dtype=np.bool)
mask.fill(mask_boolean[:,:,0])
MO = ChannelwiseOperator(MaskOperator(mask), 3, dimension = 'append')
```
## Add salt and pepper noise
```
noisy_data = noise.saltnpepper(data_with_text, amount=0.01, seed = 10)
noisy_data = MO.direct(noisy_data)
# noisy_data = MO.direct(data)
plt.figure(figsize=(10,10))
plt.imshow(noisy_data.as_array())
plt.title("Corrupted image: Missing information + Salt and pepper noise")
plt.show()
```
## Total variation inpainting vs Total Generalised Variation
We will use two different regularisation in order to restore the above corrupted image. We start with the TV regularisation described above and its generalisation, namely the **Total Generalised Variation (TGV)** introduced in [Bredies et al](https://epubs.siam.org/doi/abs/10.1137/090769521?mobileUi=0). TGV is a higher-order regulariser, that is able to obtain piecewise smooth solutions and restore staircasing artifacts that TV promotes. We let $\alpha, \beta>0$ be two regularisation parameters and define
$$
\begin{equation}
\mathrm{TGV}_{\alpha, \beta}(u) = \min_{w} \alpha \|D u - w \|_{2,1} + \beta\|\mathcal{E}w\|_{2,1},
\label{TGV}
\end{equation}$$
where $\mathcal{E}$ denotes the **Symmetrised Gradient** operator defined as
$$
\mathcal{E}w = \frac{1}{2}(D w + D w^{T}).
$$
The minimisation problems, using the $L^{1}$ norm as a data fidelity term which is suitable for salt & pepper noise, are:
$$
\begin{equation}
u^{*} =\underset{u}{\operatorname{argmin}} \|\mathcal{M}u-b\|_{1} + \alpha\mathrm{VTV}(u)
\label{TV_L1_inpainting}
\tag{TV-$L^{1}$}
\end{equation}
$$
and
<a id='TGV_L1'></a>
$$
\begin{equation}
\begin{aligned}
u^{*} =\underset{u}{\operatorname{argmin}} & \|\mathcal{M}u-b\|_{1} + \mathrm{TGV}_{\alpha, \beta}(u) \Leftrightarrow \\
(u^{*},w^{*}) =\underset{u, w}{\operatorname{argmin}} & \|\mathcal{M}u -b\|_{1} + \alpha \|D u - w \|_{2,1} + \beta\|\mathcal{E}w\|_{2,1},
\end{aligned}
\label{TGV_L1_inpainting}
\tag{TGV-$L^{1}$}
\end{equation}
$$
where the $\mathcal{M}$ is a diagonal operator with 1 in the diagonal elements corresponding to pixels in $\Omega\setminus\mathcal{D}$ and 0 in $\mathcal{D}$.
We solve the above problems using the **PDHG** algorithm described in previous notebooks.
```
# Import libraries
from cil.optimisation.operators import BlockOperator, SymmetrisedGradientOperator, GradientOperator, ZeroOperator, IdentityOperator
from cil.optimisation.functions import ZeroFunction, L1Norm, MixedL21Norm, BlockFunction, L2NormSquared
from cil.optimisation.algorithms import PDHG
```
## Setup and run the PDHG algorithm for $\mathrm{TV}-L^{1}$
```
Grad = GradientOperator(ig)
K = BlockOperator(Grad, MO)
alpha_tv = 0.5
f1 = alpha_tv * MixedL21Norm()
f2 = L1Norm(b=noisy_data)
F = BlockFunction(f1, f2)
G = ZeroFunction()
pdhg_tv = PDHG(f=F,g=G,operator=K,
max_iteration = 1000,
update_objective_interval = 200)
pdhg_tv.run(verbose = 1)
```
## Setup and run the PDHG algorithm for $\mathrm{TGV}-L^1$
Recall, that we need to define the triplet ($K$, $\mathcal{F}$, $\mathcal{G}$) and write the above problem into the following form:
$$
\begin{equation}
u^{*} =\underset{u}{\operatorname{argmin}} \mathcal{F}(Ku) + \mathcal{G}(u).
\label{general_form}
\end{equation}
$$
Let $\textbf{u} = (u, w)\in \mathbb{X}$ and define an operator $K:\mathbb{X}\rightarrow\mathbb{Y}$ as
$$
\begin{equation}
K =
\begin{bmatrix}
\mathcal{M} & \mathcal{O}\\
D & -\mathcal{I}\\
\mathcal{O} & \mathcal{E}
\end{bmatrix} \quad\Rightarrow\quad
K\textbf{u} =
K \begin{bmatrix}
u\\
w
\end{bmatrix}=
\begin{bmatrix}
\mathcal{M}u\\
Du - w\\
\mathcal{E}w
\end{bmatrix} =
\begin{bmatrix}
y_{1}\\
y_{2}\\
y_{3}
\end{bmatrix} = \textbf{y}\in \mathbb{Y},
\label{def_K}
\end{equation}
$$
where $\mathcal{O}$, $\mathcal{I}$ denote the zero and identity operators respectively.
For the function $\mathcal{F}$, we have that
$$
\begin{equation}
\begin{aligned}
& \mathcal{F}(\textbf{y}) := \mathcal{F}(y_{1}, y_{2}, y_{3}) = f_{1}(y_{1}) + f_{2}(y_{2}) + f_{3}(y_{3}), \mbox{ where},\\
& f_{1}(y_{1}) := \| y_{1} - b\|_1,\, f_{2}(y_{2}) := \alpha \|y_{2}\|_{2,1},\, f_{3}(y_{3}) := \beta\|y_{3}\|_{2,1},
\end{aligned}
\label{def_f}
\end{equation}
$$
and for the function $\mathcal{G}$, $\mathcal{G}(\textbf{u}) = \mathcal{G}(u,w) = O(u)\equiv 0 $ is the zero function.
We conclude that
$$
\begin{equation*}
\begin{aligned}
f(K\textbf{u}) + g(\textbf{u}) & = f\bigg(\begin{bmatrix}
\mathcal{M}u\\
Du - w\\
\mathcal{E}w
\end{bmatrix}\bigg) = f_{1}(\mathcal{M}u) + f_{2}(Du-w) + f_{3}(\mathcal{E}w) \\
& = \|\mathcal{M}u -b\|_{1} + \alpha \|D u - w \|_{2,1} + \beta\|\mathcal{E}w\|_{2,1},
\end{aligned}
\end{equation*}
$$
which is exactly the objective function in [TGV_L1](#TGV_L1).
```
# Regularisation parameters
alpha_tgv = 0.4
beta_tgv = 0.2
# Define BlockOperator K
K11 = MO
K21 = Grad
K32 = SymmetrisedGradientOperator(K21.range)
K12 = ZeroOperator(K32.domain, ig)
K22 = IdentityOperator(K21.range)
K31 = ZeroOperator(ig, K32.range)
K = BlockOperator(K11, K12, K21, -K22, K31, K32, shape=(3,2) )
# Define BlockFunction f
f2 = alpha_tgv * MixedL21Norm()
f3 = beta_tgv * MixedL21Norm()
f1 = L1Norm(b=noisy_data)
F = BlockFunction(f1, f2, f3)
# Setup and run the PDHG algorithm
pdhg_tgv = PDHG(f=F,g=G,operator=K,
max_iteration = 2000,
update_objective_interval = 200)
pdhg_tgv.run(verbose = 1)
images = [data, pdhg_tv.solution, (pdhg_tv.solution-data).abs()*3,
noisy_data, pdhg_tgv.solution[0], (pdhg_tgv.solution[0]-data).abs()*3],
labels_x = ["Ground Truth", "TV inpainting/denoising", " |Ground Truth - TV|",
"Corrupted Image", "TGV inpainting/denoising", " |Ground Truth - TGV|"]
# set fontszie xticks/yticks
plt.rcParams['xtick.labelsize']=15
plt.rcParams['ytick.labelsize']=15
fig = plt.figure(figsize=(20, 20))
grid = AxesGrid(fig, 111,
nrows_ncols=(2, 3),
axes_pad=0.8,
cbar_mode='single',
cbar_location='bottom',
cbar_size = 0.5,
cbar_pad=0.3
)
k = 0
for ax in grid:
img = ax.imshow(images[0][k].as_array())
ax.set_title(labels_x[k],fontsize=25)
k+=1
cbar = ax.cax.colorbar(img)
plt.show()
```
<h1><center>Conclusions</center></h1>
In this notebook, we presented how to reconstruct multichannel data with 3 channels, using two different regularisers and data fitting terms. The following notebooks will demonstrate how to reconstruct multichannel data for CT and MRI applications:
* **Dynamic CT**: Channels contain temporal information from the acquisition data.
* **Hypespectral CT**: Channels contain spectral energy information acquired from an energy-sensitive X-ray detector.
* **Sequence MRI**: Channels contain information from two MR images with different contrast.
| github_jupyter |
<a href="https://colab.research.google.com/github/mrdbourke/tensorflow-deep-learning/blob/main/09_SkimLit_nlp_milestone_project_2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# 09. Milestone Project 2: SkimLit 📄🔥
In the previous notebook ([NLP fundamentals in TensorFlow](https://github.com/mrdbourke/tensorflow-deep-learning/blob/main/08_introduction_to_nlp_in_tensorflow.ipynb)), we went through some fundamental natural lanuage processing concepts. The main ones being **tokenzation** (turning words into numbers) and **creating embeddings** (creating a numerical representation of words).
In this project, we're going to be putting what we've learned into practice.
More specificially, we're going to be replicating the deep learning model behind the 2017 paper [*PubMed 200k RCT: a Dataset for Sequenctial Sentence Classification in Medical Abstracts*](https://arxiv.org/abs/1710.06071).
When it was released, the paper presented a new dataset called PubMed 200k RCT which consists of ~200,000 labelled Randomized Controlled Trial (RCT) abstracts.
The goal of the dataset was to explore the ability for NLP models to classify sentences which appear in sequential order.
In other words, given the abstract of a RCT, what role does each sentence serve in the abstract?

*Example inputs ([harder to read abstract from PubMed](https://pubmed.ncbi.nlm.nih.gov/28942748/)) and outputs ([easier to read abstract](https://pubmed.ncbi.nlm.nih.gov/32537182/)) of the model we're going to build. The model will take an abstract wall of text and predict the section label each sentence should have.*
### Model Input
For example, can we train an NLP model which takes the following input (note: the following sample has had all numerical symbols replaced with "@"):
> To investigate the efficacy of @ weeks of daily low-dose oral prednisolone in improving pain , mobility , and systemic low-grade inflammation in the short term and whether the effect would be sustained at @ weeks in older adults with moderate to severe knee osteoarthritis ( OA ). A total of @ patients with primary knee OA were randomized @:@ ; @ received @ mg/day of prednisolone and @ received placebo for @ weeks. Outcome measures included pain reduction and improvement in function scores and systemic inflammation markers. Pain was assessed using the visual analog pain scale ( @-@ mm ).
Secondary outcome measures included the Western Ontario and McMaster Universities Osteoarthritis Index scores , patient global assessment ( PGA ) of the severity of knee OA , and @-min walk distance ( @MWD ).,
Serum levels of interleukin @ ( IL-@ ) , IL-@ , tumor necrosis factor ( TNF ) - , and high-sensitivity C-reactive protein ( hsCRP ) were measured.
There was a clinically relevant reduction in the intervention group compared to the placebo group for knee pain , physical function , PGA , and @MWD at @ weeks. The mean difference between treatment arms ( @ % CI ) was @ ( @-@ @ ) , p < @ ; @ ( @-@ @ ) , p < @ ; @ ( @-@ @ ) , p < @ ; and @ ( @-@ @ ) , p < @ , respectively. Further , there was a clinically relevant reduction in the serum levels of IL-@ , IL-@ , TNF - , and hsCRP at @ weeks in the intervention group when compared to the placebo group. These differences remained significant at @ weeks. The Outcome Measures in Rheumatology Clinical Trials-Osteoarthritis Research Society International responder rate was @ % in the intervention group and @ % in the placebo group ( p < @ ). Low-dose oral prednisolone had both a short-term and a longer sustained effect resulting in less knee pain , better physical function , and attenuation of systemic inflammation in older patients with knee OA ( ClinicalTrials.gov identifier NCT@ ).
### Model output
And returns the following output:
```
['###24293578\n',
'OBJECTIVE\tTo investigate the efficacy of @ weeks of daily low-dose oral prednisolone in improving pain , mobility , and systemic low-grade inflammation in the short term and whether the effect would be sustained at @ weeks in older adults with moderate to severe knee osteoarthritis ( OA ) .\n',
'METHODS\tA total of @ patients with primary knee OA were randomized @:@ ; @ received @ mg/day of prednisolone and @ received placebo for @ weeks .\n',
'METHODS\tOutcome measures included pain reduction and improvement in function scores and systemic inflammation markers .\n',
'METHODS\tPain was assessed using the visual analog pain scale ( @-@ mm ) .\n',
'METHODS\tSecondary outcome measures included the Western Ontario and McMaster Universities Osteoarthritis Index scores , patient global assessment ( PGA ) of the severity of knee OA , and @-min walk distance ( @MWD ) .\n',
'METHODS\tSerum levels of interleukin @ ( IL-@ ) , IL-@ , tumor necrosis factor ( TNF ) - , and high-sensitivity C-reactive protein ( hsCRP ) were measured .\n',
'RESULTS\tThere was a clinically relevant reduction in the intervention group compared to the placebo group for knee pain , physical function , PGA , and @MWD at @ weeks .\n',
'RESULTS\tThe mean difference between treatment arms ( @ % CI ) was @ ( @-@ @ ) , p < @ ; @ ( @-@ @ ) , p < @ ; @ ( @-@ @ ) , p < @ ; and @ ( @-@ @ ) , p < @ , respectively .\n',
'RESULTS\tFurther , there was a clinically relevant reduction in the serum levels of IL-@ , IL-@ , TNF - , and hsCRP at @ weeks in the intervention group when compared to the placebo group .\n',
'RESULTS\tThese differences remained significant at @ weeks .\n',
'RESULTS\tThe Outcome Measures in Rheumatology Clinical Trials-Osteoarthritis Research Society International responder rate was @ % in the intervention group and @ % in the placebo group ( p < @ ) .\n',
'CONCLUSIONS\tLow-dose oral prednisolone had both a short-term and a longer sustained effect resulting in less knee pain , better physical function , and attenuation of systemic inflammation in older patients with knee OA ( ClinicalTrials.gov identifier NCT@ ) .\n',
'\n']
```
### Problem in a sentence
The number of RCT papers released is continuing to increase, those without structured abstracts can be hard to read and in turn slow down researchers moving through the literature.
### Solution in a sentence
Create an NLP model to classify abstract sentences into the role they play (e.g. objective, methods, results, etc) to enable researchers to skim through the literature (hence SkimLit 🤓🔥) and dive deeper when necessary.
> 📖 **Resources:** Before going through the code in this notebook, you might want to get a background of what we're going to be doing. To do so, spend an hour (or two) going through the following papers and then return to this notebook:
1. Where our data is coming from: [*PubMed 200k RCT: a Dataset for Sequential Sentence Classification in Medical Abstracts*](https://arxiv.org/abs/1710.06071)
2. Where our model is coming from: [*Neural networks for joint sentence
classification in medical paper abstracts*](https://arxiv.org/pdf/1612.05251.pdf).
## What we're going to cover
Time to take what we've learned in the NLP fundmentals notebook and build our biggest NLP model yet:
* Downloading a text dataset ([PubMed RCT200k from GitHub](https://github.com/Franck-Dernoncourt/pubmed-rct))
* Writing a preprocessing function to prepare our data for modelling
* Setting up a series of modelling experiments
* Making a baseline (TF-IDF classifier)
* Deep models with different combinations of: token embeddings, character embeddings, pretrained embeddings, positional embeddings
* Building our first multimodal model (taking multiple types of data inputs)
* Replicating the model architecture from https://arxiv.org/pdf/1612.05251.pdf
* Find the most wrong predictions
* Making predictions on PubMed abstracts from the wild
## How you should approach this notebook
You can read through the descriptions and the code (it should all run, except for the cells which error on purpose), but there's a better option.
Write all of the code yourself.
Yes. I'm serious. Create a new notebook, and rewrite each line by yourself. Investigate it, see if you can break it, why does it break?
You don't have to write the text descriptions but writing the code yourself is a great way to get hands-on experience.
Don't worry if you make mistakes, we all do. The way to get better and make less mistakes is to write more code.
> 📖 **Resource:** See the full set of course materials on GitHub: https://github.com/mrdbourke/tensorflow-deep-learning
## Confirm access to a GPU
Since we're going to be building deep learning models, let's make sure we have a GPU.
In Google Colab, you can set this up by going to Runtime -> Change runtime type -> Hardware accelerator -> GPU.
If you don't have access to a GPU, the models we're building here will likely take up to 10x longer to run.
```
# Check for GPU
!nvidia-smi -L
```
## Get data
Before we can start building a model, we've got to download the PubMed 200k RCT dataset.
In a phenomenal act of kindness, the authors of the paper have made the data they used for their research availably publically and for free in the form of .txt files [on GitHub](https://github.com/Franck-Dernoncourt/pubmed-rct).
We can copy them to our local directory using `git clone https://github.com/Franck-Dernoncourt/pubmed-rct`.
```
!git clone https://github.com/Franck-Dernoncourt/pubmed-rct.git
!ls pubmed-rct
```
Checking the contents of the downloaded repository, you can see there are four folders.
Each contains a different version of the PubMed 200k RCT dataset.
Looking at the [README file](https://github.com/Franck-Dernoncourt/pubmed-rct) from the GitHub page, we get the following information:
* PubMed 20k is a subset of PubMed 200k. I.e., any abstract present in PubMed 20k is also present in PubMed 200k.
* `PubMed_200k_RCT` is the same as `PubMed_200k_RCT_numbers_replaced_with_at_sign`, except that in the latter all numbers had been replaced by `@`. (same for `PubMed_20k_RCT` vs. `PubMed_20k_RCT_numbers_replaced_with_at_sign`).
* Since Github file size limit is 100 MiB, we had to compress `PubMed_200k_RCT\train.7z` and `PubMed_200k_RCT_numbers_replaced_with_at_sign\train.zip`. To uncompress `train.7z`, you may use 7-Zip on Windows, Keka on Mac OS X, or p7zip on Linux.
To begin with, the dataset we're going to be focused on is `PubMed_20k_RCT_numbers_replaced_with_at_sign`.
Why this one?
Rather than working with the whole 200k dataset, we'll keep our experiments quick by starting with a smaller subset. We could've chosen the dataset with numbers instead of having them replaced with `@` but we didn't.
Let's check the file contents.
```
# Check what files are in the PubMed_20K dataset
!ls pubmed-rct/PubMed_20k_RCT_numbers_replaced_with_at_sign
```
Beautiful, looks like we've got three separate text files:
* `train.txt` - training samples.
* `dev.txt` - dev is short for development set, which is another name for validation set (in our case, we'll be using and referring to this file as our validation set).
* `test.txt` - test samples.
To save ourselves typing out the filepath to our target directory each time, let's turn it into a variable.
```
# Start by using the 20k dataset
data_dir = "pubmed-rct/PubMed_20k_RCT_numbers_replaced_with_at_sign/"
# Check all of the filenames in the target directory
import os
filenames = [data_dir + filename for filename in os.listdir(data_dir)]
filenames
```
## Preprocess data
Okay, now we've downloaded some text data, do you think we're ready to model it?
Wait...
We've downloaded the data but we haven't even looked at it yet.
What's the motto for getting familiar with any new dataset?
I'll give you a clue, the word begins with "v" and we say it three times.
> Vibe, vibe, vibe?
Sort of... we've definitely got to the feel the vibe of our data.
> Values, values, values?
Right again, we want to *see* lots of values but not quite what we're looking for.
> Visualize, visualize, visualize?
Boom! That's it. To get familiar and understand how we have to prepare our data for our deep learning models, we've got to visualize it.
Because our data is in the form of text files, let's write some code to read each of the lines in a target file.
```
# Create function to read the lines of a document
def get_lines(filename):
"""
Reads filename (a text file) and returns the lines of text as a list.
Args:
filename: a string containing the target filepath to read.
Returns:
A list of strings with one string per line from the target filename.
For example:
["this is the first line of filename",
"this is the second line of filename",
"..."]
"""
with open(filename, "r") as f:
return f.readlines()
```
Alright, we've got a little function, `get_lines()` which takes the filepath of a text file, opens it, reads each of the lines and returns them.
Let's try it out on the training data (`train.txt`).
```
train_lines = get_lines(data_dir+"train.txt")
train_lines[:20] # the whole first example of an abstract + a little more of the next one
```
Reading the lines from the training text file results in a list of strings containing different abstract samples, the sentences in a sample along with the role the sentence plays in the abstract.
The role of each sentence is prefixed at the start of each line separated by a tab (`\t`) and each sentence finishes with a new line (`\n`).
Different abstracts are separated by abstract ID's (lines beginning with `###`) and newlines (`\n`).
Knowing this, it looks like we've got a couple of steps to do to get our samples ready to pass as training data to our future machine learning model.
Let's write a function to perform the following steps:
* Take a target file of abstract samples.
* Read the lines in the target file.
* For each line in the target file:
* If the line begins with `###` mark it as an abstract ID and the beginning of a new abstract.
* Keep count of the number of lines in a sample.
* If the line begins with `\n` mark it as the end of an abstract sample.
* Keep count of the total lines in a sample.
* Record the text before the `\t` as the label of the line.
* Record the text after the `\t` as the text of the line.
* Return all of the lines in the target text file as a list of dictionaries containing the key/value pairs:
* `"line_number"` - the position of the line in the abstract (e.g. `3`).
* `"target"` - the role of the line in the abstract (e.g. `OBJECTIVE`).
* `"text"` - the text of the line in the abstract.
* `"total_lines"` - the total lines in an abstract sample (e.g. `14`).
* Abstract ID's and newlines should be omitted from the returned preprocessed data.
Example returned preprocessed sample (a single line from an abstract):
```
[{'line_number': 0,
'target': 'OBJECTIVE',
'text': 'to investigate the efficacy of @ weeks of daily low-dose oral prednisolone in improving pain , mobility , and systemic low-grade inflammation in the short term and whether the effect would be sustained at @ weeks in older adults with moderate to severe knee osteoarthritis ( oa ) .',
'total_lines': 11},
...]
```
```
def preprocess_text_with_line_numbers(filename):
"""Returns a list of dictionaries of abstract line data.
Takes in filename, reads its contents and sorts through each line,
extracting things like the target label, the text of the sentence,
how many sentences are in the current abstract and what sentence number
the target line is.
Args:
filename: a string of the target text file to read and extract line data
from.
Returns:
A list of dictionaries each containing a line from an abstract,
the lines label, the lines position in the abstract and the total number
of lines in the abstract where the line is from. For example:
[{"target": 'CONCLUSION',
"text": The study couldn't have gone better, turns out people are kinder than you think",
"line_number": 8,
"total_lines": 8}]
"""
input_lines = get_lines(filename) # get all lines from filename
abstract_lines = "" # create an empty abstract
abstract_samples = [] # create an empty list of abstracts
# Loop through each line in target file
for line in input_lines:
if line.startswith("###"): # check to see if line is an ID line
abstract_id = line
abstract_lines = "" # reset abstract string
elif line.isspace(): # check to see if line is a new line
abstract_line_split = abstract_lines.splitlines() # split abstract into separate lines
# Iterate through each line in abstract and count them at the same time
for abstract_line_number, abstract_line in enumerate(abstract_line_split):
line_data = {} # create empty dict to store data from line
target_text_split = abstract_line.split("\t") # split target label from text
line_data["target"] = target_text_split[0] # get target label
line_data["text"] = target_text_split[1].lower() # get target text and lower it
line_data["line_number"] = abstract_line_number # what number line does the line appear in the abstract?
line_data["total_lines"] = len(abstract_line_split) - 1 # how many total lines are in the abstract? (start from 0)
abstract_samples.append(line_data) # add line data to abstract samples list
else: # if the above conditions aren't fulfilled, the line contains a labelled sentence
abstract_lines += line
return abstract_samples
```
Beautiful! That's one good looking function. Let's use it to preprocess each of our RCT 20k datasets.
```
# Get data from file and preprocess it
%%time
train_samples = preprocess_text_with_line_numbers(data_dir + "train.txt")
val_samples = preprocess_text_with_line_numbers(data_dir + "dev.txt") # dev is another name for validation set
test_samples = preprocess_text_with_line_numbers(data_dir + "test.txt")
len(train_samples), len(val_samples), len(test_samples)
```
How do our training samples look?
```
# Check the first abstract of our training data
train_samples[:14]
```
Fantastic! Looks like our `preprocess_text_with_line_numbers()` function worked great.
How about we turn our list of dictionaries into pandas DataFrame's so we visualize them better?
```
import pandas as pd
train_df = pd.DataFrame(train_samples)
val_df = pd.DataFrame(val_samples)
test_df = pd.DataFrame(test_samples)
train_df.head(14)
```
Now our data is in DataFrame form, we can perform some data analysis on it.
```
# Distribution of labels in training data
train_df.target.value_counts()
```
Looks like sentences with the `OBJECTIVE` label are the least common.
How about we check the distribution of our abstract lengths?
```
train_df.total_lines.plot.hist();
```
Okay, looks like most of the abstracts are around 7 to 15 sentences in length.
It's good to check these things out to make sure when we do train a model or test it on unseen samples, our results aren't outlandish.
### Get lists of sentences
When we build our deep learning model, one of its main inputs will be a list of strings (the lines of an abstract).
We can get these easily from our DataFrames by calling the `tolist()` method on our `"text"` columns.
```
# Convert abstract text lines into lists
train_sentences = train_df["text"].tolist()
val_sentences = val_df["text"].tolist()
test_sentences = test_df["text"].tolist()
len(train_sentences), len(val_sentences), len(test_sentences)
# View first 10 lines of training sentences
train_sentences[:10]
```
Alright, we've separated our text samples. As you might've guessed, we'll have to write code to convert the text to numbers before we can use it with our machine learning models, we'll get to this soon.
## Make numeric labels (ML models require numeric labels)
We're going to create one hot and label encoded labels.
We could get away with just making label encoded labels, however, TensorFlow's CategoricalCrossentropy loss function likes to have one hot encoded labels (this will enable us to use label smoothing later on).
To numerically encode labels we'll use Scikit-Learn's [`OneHotEncoder`](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.OneHotEncoder.html) and [`LabelEncoder`](http://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.LabelEncoder.html) classes.
```
# One hot encode labels
from sklearn.preprocessing import OneHotEncoder
one_hot_encoder = OneHotEncoder(sparse=False)
train_labels_one_hot = one_hot_encoder.fit_transform(train_df["target"].to_numpy().reshape(-1, 1))
val_labels_one_hot = one_hot_encoder.transform(val_df["target"].to_numpy().reshape(-1, 1))
test_labels_one_hot = one_hot_encoder.transform(test_df["target"].to_numpy().reshape(-1, 1))
# Check what training labels look like
train_labels_one_hot
```
### Label encode labels
```
# Extract labels ("target" columns) and encode them into integers
from sklearn.preprocessing import LabelEncoder
label_encoder = LabelEncoder()
train_labels_encoded = label_encoder.fit_transform(train_df["target"].to_numpy())
val_labels_encoded = label_encoder.transform(val_df["target"].to_numpy())
test_labels_encoded = label_encoder.transform(test_df["target"].to_numpy())
# Check what training labels look like
train_labels_encoded
```
Now we've trained an instance of `LabelEncoder`, we can get the class names and number of classes using the `classes_` attribute.
```
# Get class names and number of classes from LabelEncoder instance
num_classes = len(label_encoder.classes_)
class_names = label_encoder.classes_
num_classes, class_names
```
## Creating a series of model experiments
We've proprocessed our data so now, in true machine learning fashion, it's time to setup a series of modelling experiments.
We'll start by creating a simple baseline model to obtain a score we'll try to beat by building more and more complex models as we move towards replicating the sequence model outlined in [*Neural networks for joint sentence
classification in medical paper abstracts*](https://arxiv.org/pdf/1612.05251.pdf).
For each model, we'll train it on the training data and evaluate it on the validation data.
## Model 0: Getting a baseline
Our first model we'll be a TF-IDF Multinomial Naive Bayes as recommended by [Scikit-Learn's machine learning map](https://scikit-learn.org/stable/tutorial/machine_learning_map/index.html).
To build it, we'll create a Scikit-Learn `Pipeline` which uses the [`TfidfVectorizer`](https://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.TfidfVectorizer.html) class to convert our abstract sentences to numbers using the TF-IDF (term frequency-inverse document frequecy) algorithm and then learns to classify our sentences using the [`MultinomialNB`](https://scikit-learn.org/stable/modules/generated/sklearn.naive_bayes.MultinomialNB.html) aglorithm.
```
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.naive_bayes import MultinomialNB
from sklearn.pipeline import Pipeline
# Create a pipeline
model_0 = Pipeline([
("tf-idf", TfidfVectorizer()),
("clf", MultinomialNB())
])
# Fit the pipeline to the training data
model_0.fit(X=train_sentences,
y=train_labels_encoded);
```
Due to the speed of the Multinomial Naive Bayes algorithm, it trains very quickly.
We can evaluate our model's accuracy on the validation dataset using the `score()` method.
```
# Evaluate baseline on validation dataset
model_0.score(X=val_sentences,
y=val_labels_encoded)
```
Nice! Looks like 72.1% accuracy will be the number to beat with our deeper models.
Now let's make some predictions with our baseline model to further evaluate it.
```
# Make predictions
baseline_preds = model_0.predict(val_sentences)
baseline_preds
```
To evaluate our baseline's predictions, we'll import the `calculate_results()` function we created in the [previous notebook](https://github.com/mrdbourke/tensorflow-deep-learning/blob/main/08_introduction_to_nlp_in_tensorflow.ipynb) and added it to our [`helper_functions.py` script](https://github.com/mrdbourke/tensorflow-deep-learning/blob/main/extras/helper_functions.py) to compare them to the ground truth labels.
More specificially the `calculate_results()` function will help us obtain the following:
* Accuracy
* Precision
* Recall
* F1-score
### Download helper functions script
Let's get our `helper_functions.py` script we've been using to store helper functions we've created in previous notebooks.
```
# Download helper functions script
!wget https://raw.githubusercontent.com/mrdbourke/tensorflow-deep-learning/main/extras/helper_functions.py
```
Now we've got the helper functions script we can import the `caculate_results()` function and see how our baseline model went.
```
# Import calculate_results helper function
from helper_functions import calculate_results
# Calculate baseline results
baseline_results = calculate_results(y_true=val_labels_encoded,
y_pred=baseline_preds)
baseline_results
```
## Preparing our data for deep sequence models
Excellent! We've got a working baseline to try and improve upon.
But before we start building deeper models, we've got to create vectorization and embedding layers.
The vectorization layer will convert our text to numbers and the embedding layer will capture the relationships between those numbers.
To start creating our vectorization and embedding layers, we'll need to import the appropriate libraries (namely TensorFlow and NumPy).
```
import numpy as np
import tensorflow as tf
from tensorflow.keras import layers
```
Since we'll be turning our sentences into numbers, it's a good idea to figure out how many words are in each sentence.
When our model goes through our sentences, it works best when they're all the same length (this is important for creating batches of the same size tensors).
For example, if one sentence is eight words long and another is 29 words long, we want to pad the eight word sentence with zeros so it ends up being the same length as the 29 word sentence.
Let's write some code to find the average length of sentences in the training set.
```
# How long is each sentence on average?
sent_lens = [len(sentence.split()) for sentence in train_sentences]
avg_sent_len = np.mean(sent_lens)
avg_sent_len # return average sentence length (in tokens)
```
How about the distribution of sentence lengths?
```
# What's the distribution look like?
import matplotlib.pyplot as plt
plt.hist(sent_lens, bins=7);
```
Looks like the vast majority of sentences are between 0 and 50 tokens in length.
We can use NumPy's [`percentile`](https://numpy.org/doc/stable/reference/generated/numpy.percentile.html) to find the value which covers 95% of the sentence lengths.
```
# How long of a sentence covers 95% of the lengths?
output_seq_len = int(np.percentile(sent_lens, 95))
output_seq_len
```
Wonderful! It looks like 95% of the sentences in our training set have a length of 55 tokens or less.
When we create our tokenization layer, we'll use this value to turn all of our sentences into the same length. Meaning sentences with a length below 55 get padded with zeros and sentences with a length above 55 get truncated (words after 55 get cut off).
> 🤔 **Question:** Why 95%?
We could use the max sentence length of the sentences in the training set.
```
# Maximum sentence length in the training set
max(sent_lens)
```
However, since hardly any sentences even come close to the max length, it would mean the majority of the data we pass to our model would be zeros (sinces all sentences below the max length would get padded with zeros).
> 🔑 **Note:** The steps we've gone through are good practice when working with a text corpus for a NLP problem. You want to know how long your samples are and what the distribution of them is. See section 4 Data Analysis of the [PubMed 200k RCT paper](https://arxiv.org/pdf/1710.06071.pdf) for further examples.
### Create text vectorizer
Now we've got a little more information about our texts, let's create a way to turn it into numbers.
To do so, we'll use the [`TextVectorization`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/experimental/preprocessing/TextVectorization) layer from TensorFlow.
We'll keep all the parameters default except for `max_tokens` (the number of unique words in our dataset) and `output_sequence_length` (our desired output length for each vectorized sentence).
Section 3.2 of the [PubMed 200k RCT paper](https://arxiv.org/pdf/1710.06071.pdf) states the vocabulary size of the PubMed 20k dataset as 68,000. So we'll use that as our `max_tokens` parameter.
```
# How many words are in our vocabulary? (taken from 3.2 in https://arxiv.org/pdf/1710.06071.pdf)
max_tokens = 68000
```
And since discovered a sentence length of 55 covers 95% of the training sentences, we'll use that as our `output_sequence_length` parameter.
```
# Create text vectorizer
from tensorflow.keras.layers.experimental.preprocessing import TextVectorization
text_vectorizer = TextVectorization(max_tokens=max_tokens, # number of words in vocabulary
output_sequence_length=55) # desired output length of vectorized sequences
```
Great! Looks like our `text_vectorizer` is ready, let's adapt it to the training data (let it read the training data and figure out what number should represent what word) and then test it out.
```
# Adapt text vectorizer to training sentences
text_vectorizer.adapt(train_sentences)
# Test out text vectorizer
import random
target_sentence = random.choice(train_sentences)
print(f"Text:\n{target_sentence}")
print(f"\nLength of text: {len(target_sentence.split())}")
print(f"\nVectorized text:\n{text_vectorizer([target_sentence])}")
```
Cool, we've now got a way to turn our sequences into numbers.
> 🛠 **Exercise:** Try running the cell above a dozen or so times. What do you notice about sequences with a length less than 55?
Using the [`get_vocabulary()`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/experimental/preprocessing/TextVectorization) method of our `text_vectorizer` we can find out a few different tidbits about our text.
```
# How many words in our training vocabulary?
rct_20k_text_vocab = text_vectorizer.get_vocabulary()
print(f"Number of words in vocabulary: {len(rct_20k_text_vocab)}"),
print(f"Most common words in the vocabulary: {rct_20k_text_vocab[:5]}")
print(f"Least common words in the vocabulary: {rct_20k_text_vocab[-5:]}")
```
And if we wanted to figure out the configuration of our `text_vectorizer` we can use the `get_config()` method.
```
# Get the config of our text vectorizer
text_vectorizer.get_config()
```
### Create custom text embedding
Our `token_vectorization` layer maps the words in our text directly to numbers. However, this doesn't necessarily capture the relationships between those numbers.
To create a richer numerical representation of our text, we can use an **embedding**.
As our model learns (by going through many different examples of abstract sentences and their labels), it'll update its embedding to better represent the relationships between tokens in our corpus.
We can create a trainable embedding layer using TensorFlow's [`Embedding`](https://www.tensorflow.org/tutorials/text/word_embeddings) layer.
Once again, the main parameters we're concerned with here are the inputs and outputs of our `Embedding` layer.
The `input_dim` parameter defines the size of our vocabulary. And the `output_dim` parameter defines the dimension of the embedding output.
Once created, our embedding layer will take the integer outputs of our `text_vectorization` layer as inputs and convert them to feature vectors of size `output_dim`.
Let's see it in action.
```
# Create token embedding layer
token_embed = layers.Embedding(input_dim=len(rct_20k_text_vocab), # length of vocabulary
output_dim=128, # Note: different embedding sizes result in drastically different numbers of parameters to train
# Use masking to handle variable sequence lengths (save space)
mask_zero=True,
name="token_embedding")
# Show example embedding
print(f"Sentence before vectorization:\n{target_sentence}\n")
vectorized_sentence = text_vectorizer([target_sentence])
print(f"Sentence after vectorization (before embedding):\n{vectorized_sentence}\n")
embedded_sentence = token_embed(vectorized_sentence)
print(f"Sentence after embedding:\n{embedded_sentence}\n")
print(f"Embedded sentence shape: {embedded_sentence.shape}")
```
## Create datasets (as fast as possible)
We've gone through all the trouble of preprocessing our datasets to be used with a machine learning model, however, there are still a few steps we can use to make them work faster with our models.
Namely, the `tf.data` API provides methods which enable faster data loading.
> 📖 **Resource:** For best practices on data loading in TensorFlow, check out the following:
* [tf.data: Build TensorFlow input pipelines](https://www.tensorflow.org/guide/data)
* [Better performance with the tf.data API](https://www.tensorflow.org/guide/data_performance)
The main steps we'll want to use with our data is to turn it into a `PrefetchDataset` of batches.
Doing so we'll ensure TensorFlow loads our data onto the GPU as fast as possible, in turn leading to faster training time.
To create a batched `PrefetchDataset` we can use the methods [`batch()`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#batch) and [`prefetch()`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#prefetch), the parameter [`tf.data.AUTOTUNE`](https://www.tensorflow.org/api_docs/python/tf/data#AUTOTUNE) will also allow TensorFlow to determine the optimal amount of compute to use to prepare datasets.
```
# Turn our data into TensorFlow Datasets
train_dataset = tf.data.Dataset.from_tensor_slices((train_sentences, train_labels_one_hot))
valid_dataset = tf.data.Dataset.from_tensor_slices((val_sentences, val_labels_one_hot))
test_dataset = tf.data.Dataset.from_tensor_slices((test_sentences, test_labels_one_hot))
train_dataset
# Take the TensorSliceDataset's and turn them into prefetched batches
train_dataset = train_dataset.batch(32).prefetch(tf.data.AUTOTUNE)
valid_dataset = valid_dataset.batch(32).prefetch(tf.data.AUTOTUNE)
test_dataset = test_dataset.batch(32).prefetch(tf.data.AUTOTUNE)
train_dataset
```
## Model 1: Conv1D with token embeddings
Alright, we've now got a way to numerically represent our text and labels, time to build a series of deep models to try and improve upon our baseline.
All of our deep models will follow a similar structure:
```
Input (text) -> Tokenize -> Embedding -> Layers -> Output (label probability)
```
The main component we'll be changing throughout is the `Layers` component. Because any modern deep NLP model requires text to be converted into an embedding before meaningful patterns can be discovered within.
The first model we're going to build is a 1-dimensional Convolutional Neural Network.
We're also going to be following the standard machine learning workflow of:
- Build model
- Train model
- Evaluate model (make predictions and compare to ground truth)
```
# Create 1D convolutional model to process sequences
inputs = layers.Input(shape=(1,), dtype=tf.string)
text_vectors = text_vectorizer(inputs) # vectorize text inputs
token_embeddings = token_embed(text_vectors) # create embedding
x = layers.Conv1D(64, kernel_size=5, padding="same", activation="relu")(token_embeddings)
x = layers.GlobalAveragePooling1D()(x) # condense the output of our feature vector
outputs = layers.Dense(num_classes, activation="softmax")(x)
model_1 = tf.keras.Model(inputs, outputs)
# Compile
model_1.compile(loss="categorical_crossentropy", # if your labels are integer form (not one hot) use sparse_categorical_crossentropy
optimizer=tf.keras.optimizers.Adam(),
metrics=["accuracy"])
# Get summary of Conv1D model
model_1.summary()
```
Wonderful! We've got our first deep sequence model built and ready to go.
Checking out the model summary, you'll notice the majority of the trainable parameters are within the embedding layer. If we were to increase the size of the embedding (by increasing the `output_dim` parameter of the `Embedding` layer), the number of trainable parameters would increase dramatically.
It's time to fit our model to the training data but we're going to make a mindful change.
Since our training data contains nearly 200,000 sentences, fitting a deep model may take a while even with a GPU. So to keep our experiments swift, we're going to run them on a subset of the training dataset.
More specifically, we'll only use the first 10% of batches (about 18,000 samples) of the training set to train on and the first 10% of batches from the validation set to validate on.
> 🔑 **Note:** It's a standard practice in machine learning to test your models on smaller subsets of data first to make sure they work before scaling them to larger amounts of data. You should aim to run many smaller experiments rather than only a handful of large experiments. And since your time is limited, one of the best ways to run smaller experiments is to reduce the amount of data you're working with (10% of the full dataset is usually a good amount, as long as it covers a similar distribution).
```
# Fit the model
model_1_history = model_1.fit(train_dataset,
steps_per_epoch=int(0.1 * len(train_dataset)), # only fit on 10% of batches for faster training time
epochs=3,
validation_data=valid_dataset,
validation_steps=int(0.1 * len(valid_dataset))) # only validate on 10% of batches
```
Brilliant! We've got our first trained deep sequence model, and it didn't take too long (and if we didn't prefetch our batched data, it would've taken longer).
Time to make some predictions with our model and then evaluate them.
```
# Evaluate on whole validation dataset (we only validated on 10% of batches during training)
model_1.evaluate(valid_dataset)
# Make predictions (our model outputs prediction probabilities for each class)
model_1_pred_probs = model_1.predict(valid_dataset)
model_1_pred_probs
# Convert pred probs to classes
model_1_preds = tf.argmax(model_1_pred_probs, axis=1)
model_1_preds
# Calculate model_1 results
model_1_results = calculate_results(y_true=val_labels_encoded,
y_pred=model_1_preds)
model_1_results
```
## Model 2: Feature extraction with pretrained token embeddings
Training our own embeddings took a little while to run, slowing our experiments down.
Since we're moving towards replicating the model architecture in [*Neural Networks for Joint Sentence Classification
in Medical Paper Abstracts*](https://arxiv.org/pdf/1612.05251.pdf), it mentions they used a [pretrained GloVe embedding](https://nlp.stanford.edu/projects/glove/) as a way to initialise their token embeddings.
To emulate this, let's see what results we can get with the [pretrained Universal Sentence Encoder embeddings from TensorFlow Hub](https://tfhub.dev/google/universal-sentence-encoder/4).
> 🔑 **Note:** We could use GloVe embeddings as per the paper but since we're working with TensorFlow, we'll use what's available from TensorFlow Hub (GloVe embeddings aren't). We'll save [using pretrained GloVe embeddings](https://keras.io/examples/nlp/pretrained_word_embeddings/) as an extension.
The model structure will look like:
```
Inputs (string) -> Pretrained embeddings from TensorFlow Hub (Universal Sentence Encoder) -> Layers -> Output (prediction probabilities)
```
You'll notice the lack of tokenization layer we've used in a previous model. This is because the Universal Sentence Encoder (USE) takes care of tokenization for us.
This type of model is called transfer learning, or more specifically, **feature extraction transfer learning**. In other words, taking the patterns a model has learned elsewhere and applying it to our own problem.
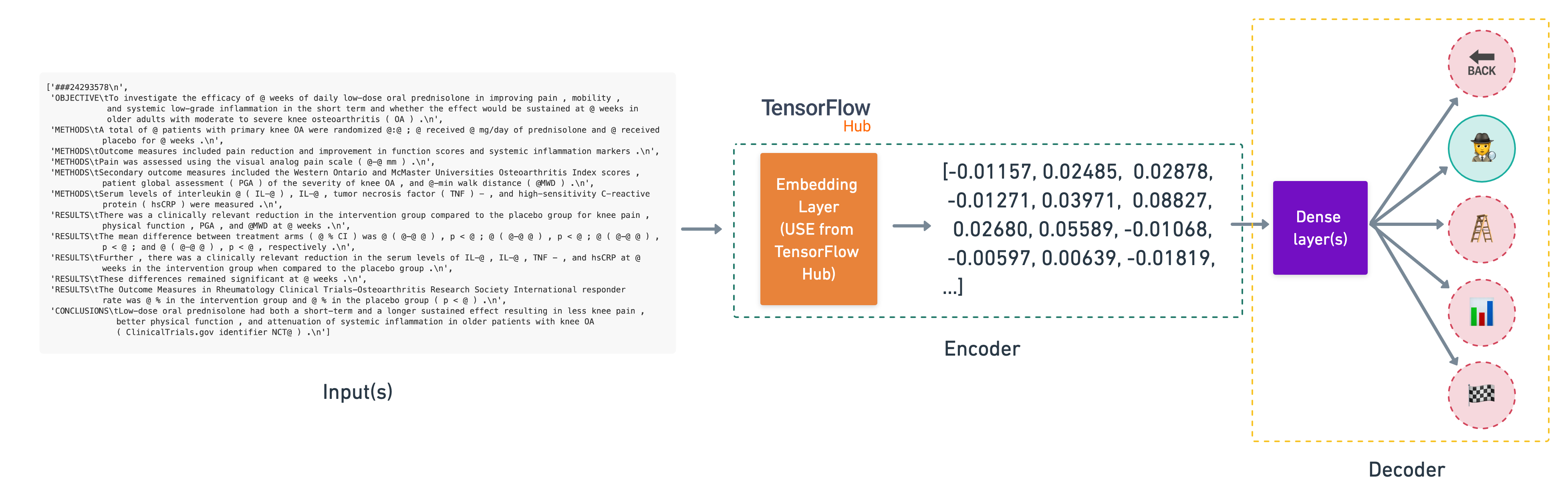
*The feature extractor model we're building using a pretrained embedding from TensorFlow Hub.*
To download the pretrained USE into a layer we can use in our model, we can use the [`hub.KerasLayer`](https://www.tensorflow.org/hub/api_docs/python/hub/KerasLayer) class.
We'll keep the pretrained embeddings frozen (by setting `trainable=False`) and add a trainable couple of layers on the top to tailor the model outputs to our own data.
> 🔑 **Note:** Due to having to download a relatively large model (~916MB), the cell below may take a little while to run.
```
# Download pretrained TensorFlow Hub USE
import tensorflow_hub as hub
tf_hub_embedding_layer = hub.KerasLayer("https://tfhub.dev/google/universal-sentence-encoder/4",
trainable=False,
name="universal_sentence_encoder")
```
Beautiful, now our pretrained USE is downloaded and instantiated as a `hub.KerasLayer` instance, let's test it out on a random sentence.
```
# Test out the embedding on a random sentence
random_training_sentence = random.choice(train_sentences)
print(f"Random training sentence:\n{random_training_sentence}\n")
use_embedded_sentence = tf_hub_embedding_layer([random_training_sentence])
print(f"Sentence after embedding:\n{use_embedded_sentence[0][:30]} (truncated output)...\n")
print(f"Length of sentence embedding:\n{len(use_embedded_sentence[0])}")
```
Nice! As we mentioned before the pretrained USE module from TensorFlow Hub takes care of tokenizing our text for us and outputs a 512 dimensional embedding vector.
Let's put together and compile a model using our `tf_hub_embedding_layer`.
### Building and fitting an NLP feature extraction model from TensorFlow Hub
```
# Define feature extractor model using TF Hub layer
inputs = layers.Input(shape=[], dtype=tf.string)
pretrained_embedding = tf_hub_embedding_layer(inputs) # tokenize text and create embedding
x = layers.Dense(128, activation="relu")(pretrained_embedding) # add a fully connected layer on top of the embedding
# Note: you could add more layers here if you wanted to
outputs = layers.Dense(5, activation="softmax")(x) # create the output layer
model_2 = tf.keras.Model(inputs=inputs,
outputs=outputs)
# Compile the model
model_2.compile(loss="categorical_crossentropy",
optimizer=tf.keras.optimizers.Adam(),
metrics=["accuracy"])
# Get a summary of the model
model_2.summary()
```
Checking the summary of our model we can see there's a large number of total parameters, however, the majority of these are non-trainable. This is because we set `training=False` when we instatiated our USE feature extractor layer.
So when we train our model, only the top two output layers will be trained.
```
# Fit feature extractor model for 3 epochs
model_2.fit(train_dataset,
steps_per_epoch=int(0.1 * len(train_dataset)),
epochs=3,
validation_data=valid_dataset,
validation_steps=int(0.1 * len(valid_dataset)))
# Evaluate on whole validation dataset
model_2.evaluate(valid_dataset)
```
Since we aren't training our own custom embedding layer, training is much quicker.
Let's make some predictions and evaluate our feature extraction model.
```
# Make predictions with feature extraction model
model_2_pred_probs = model_2.predict(valid_dataset)
model_2_pred_probs
# Convert the predictions with feature extraction model to classes
model_2_preds = tf.argmax(model_2_pred_probs, axis=1)
model_2_preds
# Calculate results from TF Hub pretrained embeddings results on validation set
model_2_results = calculate_results(y_true=val_labels_encoded,
y_pred=model_2_preds)
model_2_results
```
## Model 3: Conv1D with character embeddings
### Creating a character-level tokenizer
The [*Neural Networks for Joint Sentence Classification
in Medical Paper Abstracts*](https://arxiv.org/pdf/1612.05251.pdf) paper mentions their model uses a hybrid of token and character embeddings.
We've built models with a custom token embedding and a pretrained token embedding, how about we build one using a character embedding?
The difference between a character and token embedding is that the **character embedding** is created using sequences split into characters (e.g. `hello` -> [`h`, `e`, `l`, `l`, `o`]) where as a **token embedding** is created on sequences split into tokens.
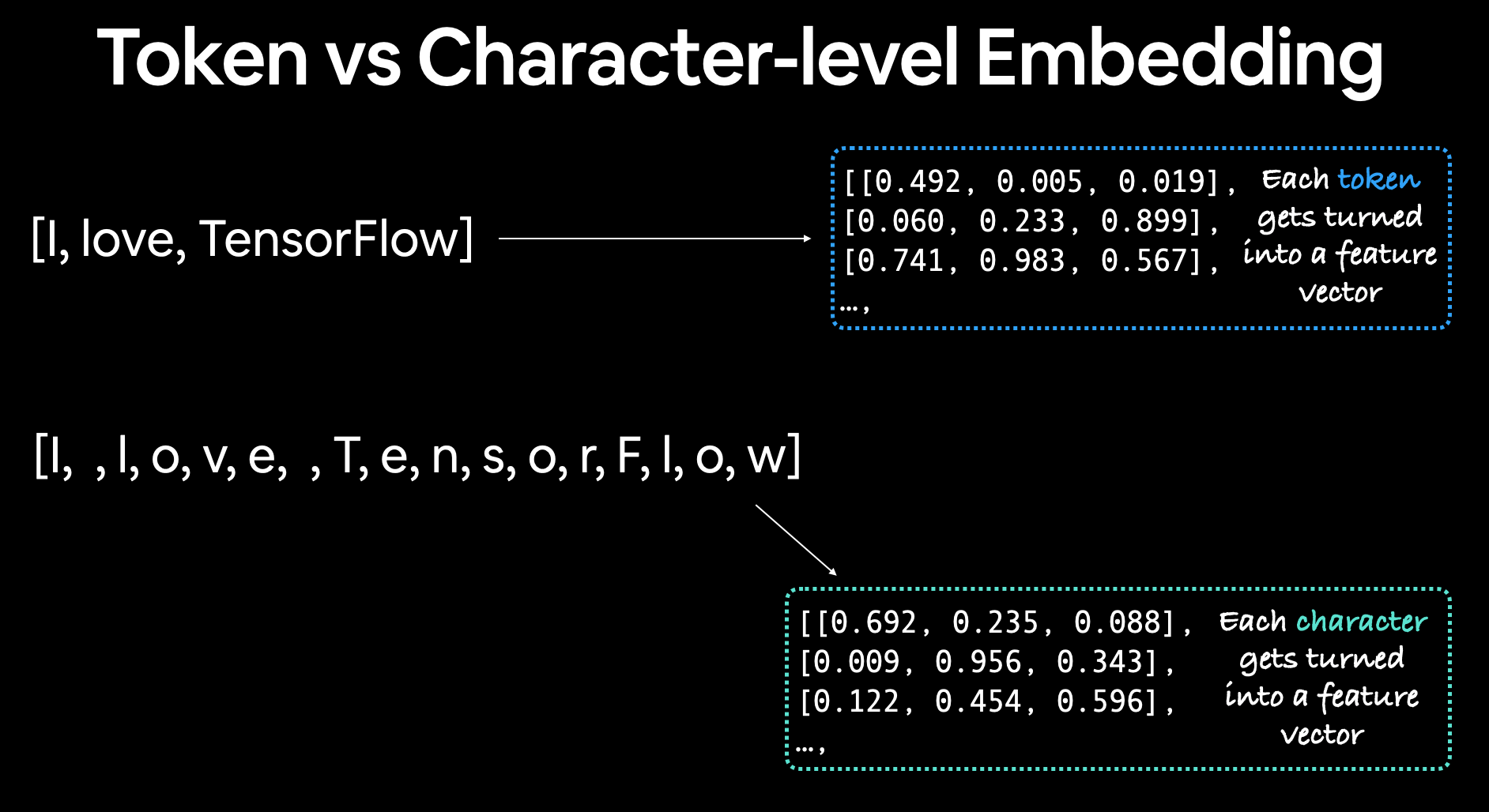

*Token level embeddings split sequences into tokens (words) and embeddings each of them, character embeddings split sequences into characters and creates a feature vector for each.*
We can create a character-level embedding by first vectorizing our sequences (after they've been split into characters) using the [`TextVectorization`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/experimental/preprocessing/TextVectorization) class and then passing those vectorized sequences through an [`Embedding`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Embedding) layer.
Before we can vectorize our sequences on a character-level we'll need to split them into characters. Let's write a function to do so.
```
# Make function to split sentences into characters
def split_chars(text):
return " ".join(list(text))
# Test splitting non-character-level sequence into characters
split_chars(random_training_sentence)
```
Great! Looks like our character-splitting function works. Let's create character-level datasets by splitting our sequence datasets into characters.
```
# Split sequence-level data splits into character-level data splits
train_chars = [split_chars(sentence) for sentence in train_sentences]
val_chars = [split_chars(sentence) for sentence in val_sentences]
test_chars = [split_chars(sentence) for sentence in test_sentences]
print(train_chars[0])
```
To figure out how long our vectorized character sequences should be, let's check the distribution of our character sequence lengths.
```
# What's the average character length?
char_lens = [len(sentence) for sentence in train_sentences]
mean_char_len = np.mean(char_lens)
mean_char_len
# Check the distribution of our sequences at character-level
import matplotlib.pyplot as plt
plt.hist(char_lens, bins=7);
```
Okay, looks like most of our sequences are between 0 and 200 characters long.
Let's use NumPy's percentile to figure out what length covers 95% of our sequences.
```
# Find what character length covers 95% of sequences
output_seq_char_len = int(np.percentile(char_lens, 95))
output_seq_char_len
```
Wonderful, now we know the sequence length which covers 95% of sequences, we'll use that in our `TextVectorization` layer as the `output_sequence_length` parameter.
> 🔑 **Note:** You can experiment here to figure out what the optimal `output_sequence_length` should be, perhaps using the mean results in as good results as using the 95% percentile.
We'll set `max_tokens` (the total number of different characters in our sequences) to 28, in other words, 26 letters of the alphabet + space + OOV (out of vocabulary or unknown) tokens.
```
# Get all keyboard characters for char-level embedding
import string
alphabet = string.ascii_lowercase + string.digits + string.punctuation
alphabet
# Create char-level token vectorizer instance
NUM_CHAR_TOKENS = len(alphabet) + 2 # num characters in alphabet + space + OOV token
char_vectorizer = TextVectorization(max_tokens=NUM_CHAR_TOKENS,
output_sequence_length=output_seq_char_len,
standardize="lower_and_strip_punctuation",
name="char_vectorizer")
# Adapt character vectorizer to training characters
char_vectorizer.adapt(train_chars)
```
Nice! Now we've adapted our `char_vectorizer` to our character-level sequences, let's check out some characteristics about it using the [`get_vocabulary()`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/experimental/preprocessing/TextVectorization#get_vocabulary) method.
```
# Check character vocabulary characteristics
char_vocab = char_vectorizer.get_vocabulary()
print(f"Number of different characters in character vocab: {len(char_vocab)}")
print(f"5 most common characters: {char_vocab[:5]}")
print(f"5 least common characters: {char_vocab[-5:]}")
```
We can also test it on random sequences of characters to make sure it's working.
```
# Test out character vectorizer
random_train_chars = random.choice(train_chars)
print(f"Charified text:\n{random_train_chars}")
print(f"\nLength of chars: {len(random_train_chars.split())}")
vectorized_chars = char_vectorizer([random_train_chars])
print(f"\nVectorized chars:\n{vectorized_chars}")
print(f"\nLength of vectorized chars: {len(vectorized_chars[0])}")
```
You'll notice sequences with a length shorter than 290 (`output_seq_char_length`) get padded with zeros on the end, this ensures all sequences passed to our model are the same length.
Also, due to the `standardize` parameter of `TextVectorization` being `"lower_and_strip_punctuation"` and the `split` parameter being `"whitespace"` by default, symbols (such as `@`) and spaces are removed.
> 🔑 **Note:** If you didn't want punctuation to be removed (keep the `@`, `%` etc), you can create a custom standardization callable and pass it as the `standardize` parameter. See the [`TextVectorization`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/experimental/preprocessing/TextVectorization) class documentation for more.
### Creating a character-level embedding
We've got a way to vectorize our character-level sequences, now's time to create a character-level embedding.
Just like our custom token embedding, we can do so using the [`tensorflow.keras.layers.Embedding`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Embedding) class.
Our character-level embedding layer requires an input dimension and output dimension.
The input dimension (`input_dim`) will be equal to the number of different characters in our `char_vocab` (28). And since we're following the structure of the model in Figure 1 of [*Neural Networks for Joint Sentence Classification
in Medical Paper Abstracts*](https://arxiv.org/pdf/1612.05251.pdf), the output dimension of the character embedding (`output_dim`) will be 25.
```
# Create char embedding layer
char_embed = layers.Embedding(input_dim=NUM_CHAR_TOKENS, # number of different characters
output_dim=25, # embedding dimension of each character (same as Figure 1 in https://arxiv.org/pdf/1612.05251.pdf)
mask_zero=True,
name="char_embed")
# Test out character embedding layer
print(f"Charified text (before vectorization and embedding):\n{random_train_chars}\n")
char_embed_example = char_embed(char_vectorizer([random_train_chars]))
print(f"Embedded chars (after vectorization and embedding):\n{char_embed_example}\n")
print(f"Character embedding shape: {char_embed_example.shape}")
```
Wonderful! Each of the characters in our sequences gets turned into a 25 dimension embedding.
### Building a Conv1D model to fit on character embeddings
Now we've got a way to turn our character-level sequences into numbers (`char_vectorizer`) as well as numerically represent them as an embedding (`char_embed`) let's test how effective they are at encoding the information in our sequences by creating a character-level sequence model.
The model will have the same structure as our custom token embedding model (`model_1`) except it'll take character-level sequences as input instead of token-level sequences.
```
Input (character-level text) -> Tokenize -> Embedding -> Layers (Conv1D, GlobalMaxPool1D) -> Output (label probability)
```
```
# Make Conv1D on chars only
inputs = layers.Input(shape=(1,), dtype="string")
char_vectors = char_vectorizer(inputs)
char_embeddings = char_embed(char_vectors)
x = layers.Conv1D(64, kernel_size=5, padding="same", activation="relu")(char_embeddings)
x = layers.GlobalMaxPool1D()(x)
outputs = layers.Dense(num_classes, activation="softmax")(x)
model_3 = tf.keras.Model(inputs=inputs,
outputs=outputs,
name="model_3_conv1D_char_embedding")
# Compile model
model_3.compile(loss="categorical_crossentropy",
optimizer=tf.keras.optimizers.Adam(),
metrics=["accuracy"])
# Check the summary of conv1d_char_model
model_3.summary()
```
Before fitting our model on the data, we'll create char-level batched `PrefetchedDataset`'s.
```
# Create char datasets
train_char_dataset = tf.data.Dataset.from_tensor_slices((train_chars, train_labels_one_hot)).batch(32).prefetch(tf.data.AUTOTUNE)
val_char_dataset = tf.data.Dataset.from_tensor_slices((val_chars, val_labels_one_hot)).batch(32).prefetch(tf.data.AUTOTUNE)
train_char_dataset
```
Just like our token-level sequence model, to save time with our experiments, we'll fit the character-level model on 10% of batches.
```
# Fit the model on chars only
model_3_history = model_3.fit(train_char_dataset,
steps_per_epoch=int(0.1 * len(train_char_dataset)),
epochs=3,
validation_data=val_char_dataset,
validation_steps=int(0.1 * len(val_char_dataset)))
# Evaluate model_3 on whole validation char dataset
model_3.evaluate(val_char_dataset)
```
Nice! Looks like our character-level model is working, let's make some predictions with it and evaluate them.
```
# Make predictions with character model only
model_3_pred_probs = model_3.predict(val_char_dataset)
model_3_pred_probs
# Convert predictions to classes
model_3_preds = tf.argmax(model_3_pred_probs, axis=1)
model_3_preds
# Calculate Conv1D char only model results
model_3_results = calculate_results(y_true=val_labels_encoded,
y_pred=model_3_preds)
model_3_results
```
## Model 4: Combining pretrained token embeddings + character embeddings (hybrid embedding layer)
Alright, now things are going to get spicy.
In moving closer to build a model similar to the one in Figure 1 of [*Neural Networks for Joint Sentence Classification
in Medical Paper Abstracts*](https://arxiv.org/pdf/1612.05251.pdf), it's time we tackled the hybrid token embedding layer they speak of.
This hybrid token embedding layer is a combination of token embeddings and character embeddings. In other words, they create a stacked embedding to represent sequences before passing them to the sequence label prediction layer.
So far we've built two models which have used token and character-level embeddings, however, these two models have used each of these embeddings exclusively.
To start replicating (or getting close to replicating) the model in Figure 1, we're going to go through the following steps:
1. Create a token-level model (similar to `model_1`)
2. Create a character-level model (similar to `model_3` with a slight modification to reflect the paper)
3. Combine (using [`layers.Concatenate`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Concatenate)) the outputs of 1 and 2
4. Build a series of output layers on top of 3 similar to Figure 1 and section 4.2 of [*Neural Networks for Joint Sentence Classification
in Medical Paper Abstracts*](https://arxiv.org/pdf/1612.05251.pdf)
5. Construct a model which takes token and character-level sequences as input and produces sequence label probabilities as output
```
# 1. Setup token inputs/model
token_inputs = layers.Input(shape=[], dtype=tf.string, name="token_input")
token_embeddings = tf_hub_embedding_layer(token_inputs)
token_output = layers.Dense(128, activation="relu")(token_embeddings)
token_model = tf.keras.Model(inputs=token_inputs,
outputs=token_output)
# 2. Setup char inputs/model
char_inputs = layers.Input(shape=(1,), dtype=tf.string, name="char_input")
char_vectors = char_vectorizer(char_inputs)
char_embeddings = char_embed(char_vectors)
char_bi_lstm = layers.Bidirectional(layers.LSTM(25))(char_embeddings) # bi-LSTM shown in Figure 1 of https://arxiv.org/pdf/1612.05251.pdf
char_model = tf.keras.Model(inputs=char_inputs,
outputs=char_bi_lstm)
# 3. Concatenate token and char inputs (create hybrid token embedding)
token_char_concat = layers.Concatenate(name="token_char_hybrid")([token_model.output,
char_model.output])
# 4. Create output layers - addition of dropout discussed in 4.2 of https://arxiv.org/pdf/1612.05251.pdf
combined_dropout = layers.Dropout(0.5)(token_char_concat)
combined_dense = layers.Dense(200, activation="relu")(combined_dropout) # slightly different to Figure 1 due to different shapes of token/char embedding layers
final_dropout = layers.Dropout(0.5)(combined_dense)
output_layer = layers.Dense(num_classes, activation="softmax")(final_dropout)
# 5. Construct model with char and token inputs
model_4 = tf.keras.Model(inputs=[token_model.input, char_model.input],
outputs=output_layer,
name="model_4_token_and_char_embeddings")
```
Woah... There's a lot going on here, let's get a summary and plot our model to visualize what's happening.
```
# Get summary of token and character model
model_4.summary()
# Plot hybrid token and character model
from keras.utils import plot_model
plot_model(model_4)
```
Now that's a good looking model. Let's compile it just as we have the rest of our models.
> 🔑 **Note:** Section 4.2 of [*Neural Networks for Joint Sentence Classification
in Medical Paper Abstracts*](https://arxiv.org/pdf/1612.05251.pdf) mentions using the SGD (stochastic gradient descent) optimizer, however, to stay consistent with our other models, we're going to use the Adam optimizer. As an exercise, you could try using [`tf.keras.optimizers.SGD`](https://www.tensorflow.org/api_docs/python/tf/keras/optimizers/SGD) instead of [`tf.keras.optimizers.Adam`](https://www.tensorflow.org/api_docs/python/tf/keras/optimizers/Adam) and compare the results.
```
# Compile token char model
model_4.compile(loss="categorical_crossentropy",
optimizer=tf.keras.optimizers.Adam(), # section 4.2 of https://arxiv.org/pdf/1612.05251.pdf mentions using SGD but we'll stick with Adam
metrics=["accuracy"])
```
And again, to keep our experiments fast, we'll fit our token-character-hybrid model on 10% of training and validate on 10% of validation batches. However, the difference with this model is that it requires two inputs, token-level sequences and character-level sequences.
We can do this by create a `tf.data.Dataset` with a tuple as it's first input, for example:
* `((token_data, char_data), (label))`
Let's see it in action.
### Combining token and character data into a `tf.data` dataset
```
# Combine chars and tokens into a dataset
train_char_token_data = tf.data.Dataset.from_tensor_slices((train_sentences, train_chars)) # make data
train_char_token_labels = tf.data.Dataset.from_tensor_slices(train_labels_one_hot) # make labels
train_char_token_dataset = tf.data.Dataset.zip((train_char_token_data, train_char_token_labels)) # combine data and labels
# Prefetch and batch train data
train_char_token_dataset = train_char_token_dataset.batch(32).prefetch(tf.data.AUTOTUNE)
# Repeat same steps validation data
val_char_token_data = tf.data.Dataset.from_tensor_slices((val_sentences, val_chars))
val_char_token_labels = tf.data.Dataset.from_tensor_slices(val_labels_one_hot)
val_char_token_dataset = tf.data.Dataset.zip((val_char_token_data, val_char_token_labels))
val_char_token_dataset = val_char_token_dataset.batch(32).prefetch(tf.data.AUTOTUNE)
# Check out training char and token embedding dataset
train_char_token_dataset, val_char_token_dataset
```
### Fitting a model on token and character-level sequences
```
# Fit the model on tokens and chars
model_4_history = model_4.fit(train_char_token_dataset, # train on dataset of token and characters
steps_per_epoch=int(0.1 * len(train_char_token_dataset)),
epochs=3,
validation_data=val_char_token_dataset,
validation_steps=int(0.1 * len(val_char_token_dataset)))
# Evaluate on the whole validation dataset
model_4.evaluate(val_char_token_dataset)
```
Nice! Our token-character hybrid model has come to life!
To make predictions with it, since it takes multiplie inputs, we can pass the `predict()` method a tuple of token-level sequences and character-level sequences.
We can then evaluate the predictions as we've done before.
```
# Make predictions using the token-character model hybrid
model_4_pred_probs = model_4.predict(val_char_token_dataset)
model_4_pred_probs
# Turn prediction probabilities into prediction classes
model_4_preds = tf.argmax(model_4_pred_probs, axis=1)
model_4_preds
# Get results of token-char-hybrid model
model_4_results = calculate_results(y_true=val_labels_encoded,
y_pred=model_4_preds)
model_4_results
```
## Model 5: Transfer Learning with pretrained token embeddings + character embeddings + positional embeddings
It seems like combining token embeddings and character embeddings gave our model a little performance boost.
But there's one more piece of the puzzle we can add in.
What if we engineered our own features into the model?
Meaning, what if we took our own knowledge about the data and encoded it in a numerical way to give our model more information about our samples?
The process of applying your own knowledge to build features as input to a model is called **feature engineering**.
Can you think of something important about the sequences we're trying to classify?
If you were to look at an abstract, would you expect the sentences to appear in order? Or does it make sense if they were to appear sequentially? For example, sequences labelled `CONCLUSIONS` at the beggining and sequences labelled `OBJECTIVE` at the end?
Abstracts typically come in a sequential order, such as:
* `OBJECTIVE` ...
* `METHODS` ...
* `METHODS` ...
* `METHODS` ...
* `RESULTS` ...
* `CONCLUSIONS` ...
Or
* `BACKGROUND` ...
* `OBJECTIVE` ...
* `METHODS` ...
* `METHODS` ...
* `RESULTS` ...
* `RESULTS` ...
* `CONCLUSIONS` ...
* `CONCLUSIONS` ...
Of course, we can't engineer the sequence labels themselves into the training data (we don't have these at test time), but we can encode the order of a set of sequences in an abstract.
For example,
* `Sentence 1 of 10` ...
* `Sentence 2 of 10` ...
* `Sentence 3 of 10` ...
* `Sentence 4 of 10` ...
* ...
You might've noticed this when we created our `preprocess_text_with_line_numbers()` function. When we read in a text file of abstracts, we counted the number of lines in an abstract as well as the number of each line itself.
Doing this led to the `"line_number"` and `"total_lines"` columns of our DataFrames.
```
# Inspect training dataframe
train_df.head()
```
The `"line_number"` and `"total_lines"` columns are features which didn't necessarily come with the training data but can be passed to our model as a **positional embedding**. In other words, the positional embedding is where the sentence appears in an abstract.
We can use these features because they will be available at test time.

*Since abstracts typically have a sequential order about them (for example, background, objective, methods, results, conclusion), it makes sense to add the line number of where a particular sentence occurs to our model. The beautiful thing is, these features will be available at test time (we can just count the number of sentences in an abstract and the number of each one).*
Meaning, if we were to predict the labels of sequences in an abstract our model had never seen, we could count the number of lines and the track the position of each individual line and pass it to our model.
> 🛠 **Exercise:** Another way of creating our positional embedding feature would be to combine the `"line_number"` and `"total_lines"` columns into one, for example a `"line_position"` column may contain values like `1_of_11`, `2_of_11`, etc. Where `1_of_11` would be the first line in an abstract 11 sentences long. After going through the following steps, you might want to revisit this positional embedding stage and see how a combined column of `"line_position"` goes against two separate columns.
### Create positional embeddings
Okay, enough talk about positional embeddings, let's create them.
Since our `"line_number"` and `"total_line"` columns are already numerical, we could pass them as they are to our model.
But to avoid our model thinking a line with `"line_number"=5` is five times greater than a line with `"line_number"=1`, we'll use one-hot-encoding to encode our `"line_number"` and `"total_lines"` features.
To do this, we can use the [`tf.one_hot`](https://www.tensorflow.org/api_docs/python/tf/one_hot) utility.
`tf.one_hot` returns a one-hot-encoded tensor. It accepts an array (or tensor) as input and the `depth` parameter determines the dimension of the returned tensor.
To figure out what we should set the `depth` parameter to, let's investigate the distribution of the `"line_number"` column.
> 🔑 **Note:** When it comes to one-hot-encoding our features, Scikit-Learn's [`OneHotEncoder`](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.OneHotEncoder.html) class is another viable option here.
```
# How many different line numbers are there?
train_df["line_number"].value_counts()
# Check the distribution of "line_number" column
train_df.line_number.plot.hist()
```
Looking at the distribution of the `"line_number"` column, it looks like the majority of lines have a position of 15 or less.
Knowing this, let's set the `depth` parameter of `tf.one_hot` to 15.
```
# Use TensorFlow to create one-hot-encoded tensors of our "line_number" column
train_line_numbers_one_hot = tf.one_hot(train_df["line_number"].to_numpy(), depth=15)
val_line_numbers_one_hot = tf.one_hot(val_df["line_number"].to_numpy(), depth=15)
test_line_numbers_one_hot = tf.one_hot(test_df["line_number"].to_numpy(), depth=15)
```
Setting the `depth` parameter of `tf.one_hot` to 15 means any sample with a `"line_number"` value of over 15 gets set to a tensor of all 0's, where as any sample with a `"line_number"` of under 15 gets turned into a tensor of all 0's but with a 1 at the index equal to the `"line_number"` value.
> 🔑 **Note:** We could create a one-hot tensor which has room for all of the potential values of `"line_number"` (`depth=30`), however, this would end up in a tensor of double the size of our current one (`depth=15`) where the vast majority of values are 0. Plus, only ~2,000/180,000 samples have a `"line_number"` value of over 15. So we would not be gaining much information about our data for doubling our feature space. This kind of problem is called the **curse of dimensionality**. However, since this we're working with deep models, it might be worth trying to throw as much information at the model as possible and seeing what happens. I'll leave exploring values of the `depth` parameter as an extension.
```
# Check one-hot encoded "line_number" feature samples
train_line_numbers_one_hot.shape, train_line_numbers_one_hot[:20]
```
We can do the same as we've done for our `"line_number"` column witht he `"total_lines"` column. First, let's find an appropriate value for the `depth` parameter of `tf.one_hot`.
```
# How many different numbers of lines are there?
train_df["total_lines"].value_counts()
# Check the distribution of total lines
train_df.total_lines.plot.hist();
```
Looking at the distribution of our `"total_lines"` column, a value of 20 looks like it covers the majority of samples.
We can confirm this with [`np.percentile()`](https://numpy.org/doc/stable/reference/generated/numpy.percentile.html).
```
# Check the coverage of a "total_lines" value of 20
np.percentile(train_df.total_lines, 98) # a value of 20 covers 98% of samples
```
Beautiful! Plenty of converage. Let's one-hot-encode our `"total_lines"` column just as we did our `"line_number"` column.
```
# Use TensorFlow to create one-hot-encoded tensors of our "total_lines" column
train_total_lines_one_hot = tf.one_hot(train_df["total_lines"].to_numpy(), depth=20)
val_total_lines_one_hot = tf.one_hot(val_df["total_lines"].to_numpy(), depth=20)
test_total_lines_one_hot = tf.one_hot(test_df["total_lines"].to_numpy(), depth=20)
# Check shape and samples of total lines one-hot tensor
train_total_lines_one_hot.shape, train_total_lines_one_hot[:10]
```
### Building a tribrid embedding model
Woohoo! Positional embedding tensors ready.
It's time to build the biggest model we've built yet. One which incorporates token embeddings, character embeddings and our newly crafted positional embeddings.
We'll be venturing into uncovered territory but there will be nothing here you haven't practiced before.
More specifically we're going to go through the following steps:
1. Create a token-level model (similar to `model_1`)
2. Create a character-level model (similar to `model_3` with a slight modification to reflect the paper)
3. Create a `"line_number"` model (takes in one-hot-encoded `"line_number"` tensor and passes it through a non-linear layer)
4. Create a `"total_lines"` model (takes in one-hot-encoded `"total_lines"` tensor and passes it through a non-linear layer)
5. Combine (using [`layers.Concatenate`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Concatenate)) the outputs of 1 and 2 into a token-character-hybrid embedding and pass it series of output to Figure 1 and section 4.2 of [*Neural Networks for Joint Sentence Classification
in Medical Paper Abstracts*](https://arxiv.org/pdf/1612.05251.pdf)
6. Combine (using [`layers.Concatenate`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Concatenate)) the outputs of 3, 4 and 5 into a token-character-positional tribrid embedding
7. Create an output layer to accept the tribrid embedding and output predicted label probabilities
8. Combine the inputs of 1, 2, 3, 4 and outputs of 7 into a [`tf.keras.Model`](https://www.tensorflow.org/api_docs/python/tf/keras/Model)
Woah! That's alot... but nothing we're not capable of. Let's code it.
```
# 1. Token inputs
token_inputs = layers.Input(shape=[], dtype="string", name="token_inputs")
token_embeddings = tf_hub_embedding_layer(token_inputs)
token_outputs = layers.Dense(128, activation="relu")(token_embeddings)
token_model = tf.keras.Model(inputs=token_inputs,
outputs=token_embeddings)
# 2. Char inputs
char_inputs = layers.Input(shape=(1,), dtype="string", name="char_inputs")
char_vectors = char_vectorizer(char_inputs)
char_embeddings = char_embed(char_vectors)
char_bi_lstm = layers.Bidirectional(layers.LSTM(32))(char_embeddings)
char_model = tf.keras.Model(inputs=char_inputs,
outputs=char_bi_lstm)
# 3. Line numbers inputs
line_number_inputs = layers.Input(shape=(15,), dtype=tf.int32, name="line_number_input")
x = layers.Dense(32, activation="relu")(line_number_inputs)
line_number_model = tf.keras.Model(inputs=line_number_inputs,
outputs=x)
# 4. Total lines inputs
total_lines_inputs = layers.Input(shape=(20,), dtype=tf.int32, name="total_lines_input")
y = layers.Dense(32, activation="relu")(total_lines_inputs)
total_line_model = tf.keras.Model(inputs=total_lines_inputs,
outputs=y)
# 5. Combine token and char embeddings into a hybrid embedding
combined_embeddings = layers.Concatenate(name="token_char_hybrid_embedding")([token_model.output,
char_model.output])
z = layers.Dense(256, activation="relu")(combined_embeddings)
z = layers.Dropout(0.5)(z)
# 6. Combine positional embeddings with combined token and char embeddings into a tribrid embedding
z = layers.Concatenate(name="token_char_positional_embedding")([line_number_model.output,
total_line_model.output,
z])
# 7. Create output layer
output_layer = layers.Dense(5, activation="softmax", name="output_layer")(z)
# 8. Put together model
model_5 = tf.keras.Model(inputs=[line_number_model.input,
total_line_model.input,
token_model.input,
char_model.input],
outputs=output_layer)
```
There's a lot going on here... let's visualize what's happening with a summary by plotting our model.
```
# Get a summary of our token, char and positional embedding model
model_5.summary()
# Plot the token, char, positional embedding model
from tensorflow.keras.utils import plot_model
plot_model(model_5)
```
Visualizing the model makes it much easier to understand.
Essentially what we're doing is trying to encode as much information about our sequences as possible into various embeddings (the inputs to our model) so our model has the best chance to figure out what label belongs to a sequence (the outputs of our model).
You'll notice our model is looking very similar to the model shown in Figure 1 of [*Neural Networks for Joint Sentence Classification
in Medical Paper Abstracts*](https://arxiv.org/pdf/1612.05251.pdf). However, a few differences still remain:
* We're using pretrained TensorFlow Hub token embeddings instead of GloVe emebddings.
* We're using a Dense layer on top of our token-character hybrid embeddings instead of a bi-LSTM layer.
* Section 3.1.3 of the paper mentions a label sequence optimization layer (which helps to make sure sequence labels come out in a respectable order) but it isn't shown in Figure 1. To makeup for the lack of this layer in our model, we've created the positional embeddings layers.
* Section 4.2 of the paper mentions the token and character embeddings are updated during training, our pretrained TensorFlow Hub embeddings remain frozen.
* The paper uses the [`SGD`](https://www.tensorflow.org/api_docs/python/tf/keras/optimizers/SGD) optimizer, we're going to stick with [`Adam`](https://www.tensorflow.org/api_docs/python/tf/keras/optimizers/Adam).
All of the differences above are potential extensions of this project.
```
# Check which layers of our model are trainable or not
for layer in model_5.layers:
print(layer, layer.trainable)
```
Now our model is constructed, let's compile it.
This time, we're going to introduce a new parameter to our loss function called `label_smoothing`. Label smoothing helps to regularize our model (prevent overfitting) by making sure it doesn't get too focused on applying one particular label to a sample.
For example, instead of having an output prediction of:
* `[0.0, 0.0, 1.0, 0.0, 0.0]` for a sample (the model is very confident the right label is index 2).
It's predictions will get smoothed to be something like:
* `[0.01, 0.01, 0.096, 0.01, 0.01]` giving a small activation to each of the other labels, in turn, hopefully improving generalization.
> 📖 **Resource:** For more on label smoothing, see the great blog post by PyImageSearch, [*Label smoothing with Keras, TensorFlow, and Deep Learning*](https://www.pyimagesearch.com/2019/12/30/label-smoothing-with-keras-tensorflow-and-deep-learning/).
```
# Compile token, char, positional embedding model
model_5.compile(loss=tf.keras.losses.CategoricalCrossentropy(label_smoothing=0.2), # add label smoothing (examples which are really confident get smoothed a little)
optimizer=tf.keras.optimizers.Adam(),
metrics=["accuracy"])
```
### Create tribrid embedding datasets and fit tribrid model
Model compiled!
Again, to keep our experiments swift, let's fit on 20,000 examples for 3 epochs.
This time our model requires four feature inputs:
1. Train line numbers one-hot tensor (`train_line_numbers_one_hot`)
2. Train total lines one-hot tensor (`train_total_lines_one_hot`)
3. Token-level sequences tensor (`train_sentences`)
4. Char-level sequences tensor (`train_chars`)
We can pass these as tuples to our `tf.data.Dataset.from_tensor_slices()` method to create appropriately shaped and batched `PrefetchedDataset`'s.
```
# Create training and validation datasets (all four kinds of inputs)
train_pos_char_token_data = tf.data.Dataset.from_tensor_slices((train_line_numbers_one_hot, # line numbers
train_total_lines_one_hot, # total lines
train_sentences, # train tokens
train_chars)) # train chars
train_pos_char_token_labels = tf.data.Dataset.from_tensor_slices(train_labels_one_hot) # train labels
train_pos_char_token_dataset = tf.data.Dataset.zip((train_pos_char_token_data, train_pos_char_token_labels)) # combine data and labels
train_pos_char_token_dataset = train_pos_char_token_dataset.batch(32).prefetch(tf.data.AUTOTUNE) # turn into batches and prefetch appropriately
# Validation dataset
val_pos_char_token_data = tf.data.Dataset.from_tensor_slices((val_line_numbers_one_hot,
val_total_lines_one_hot,
val_sentences,
val_chars))
val_pos_char_token_labels = tf.data.Dataset.from_tensor_slices(val_labels_one_hot)
val_pos_char_token_dataset = tf.data.Dataset.zip((val_pos_char_token_data, val_pos_char_token_labels))
val_pos_char_token_dataset = val_pos_char_token_dataset.batch(32).prefetch(tf.data.AUTOTUNE) # turn into batches and prefetch appropriately
# Check input shapes
train_pos_char_token_dataset, val_pos_char_token_dataset
# Fit the token, char and positional embedding model
history_model_5 = model_5.fit(train_pos_char_token_dataset,
steps_per_epoch=int(0.1 * len(train_pos_char_token_dataset)),
epochs=3,
validation_data=val_pos_char_token_dataset,
validation_steps=int(0.1 * len(val_pos_char_token_dataset)))
```
Tribrid model trained! Time to make some predictions with it and evaluate them just as we've done before.
```
# Make predictions with token-char-positional hybrid model
model_5_pred_probs = model_5.predict(val_pos_char_token_dataset, verbose=1)
model_5_pred_probs
# Turn prediction probabilities into prediction classes
model_5_preds = tf.argmax(model_5_pred_probs, axis=1)
model_5_preds
# Calculate results of token-char-positional hybrid model
model_5_results = calculate_results(y_true=val_labels_encoded,
y_pred=model_5_preds)
model_5_results
```
## Compare model results
Far out, we've come a long way. From a baseline model to training a model containing three different kinds of embeddings.
Now it's time to compare each model's performance against each other.
We'll also be able to compare our model's to the [*PubMed 200k RCT:
a Dataset for Sequential Sentence Classification in Medical Abstracts*](https://arxiv.org/pdf/1710.06071.pdf) paper.
Since all of our model results are in dictionaries, let's combine them into a pandas DataFrame to visualize them.
```
# Combine model results into a DataFrame
all_model_results = pd.DataFrame({"baseline": baseline_results,
"custom_token_embed_conv1d": model_1_results,
"pretrained_token_embed": model_2_results,
"custom_char_embed_conv1d": model_3_results,
"hybrid_char_token_embed": model_4_results,
"tribrid_pos_char_token_embed": model_5_results})
all_model_results = all_model_results.transpose()
all_model_results
# Reduce the accuracy to same scale as other metrics
all_model_results["accuracy"] = all_model_results["accuracy"]/100
# Plot and compare all of the model results
all_model_results.plot(kind="bar", figsize=(10, 7)).legend(bbox_to_anchor=(1.0, 1.0));
```
Since the [*PubMed 200k RCT:
a Dataset for Sequential Sentence Classification in Medical Abstracts*](https://arxiv.org/pdf/1710.06071.pdf) paper compares their tested model's F1-scores on the test dataset, let's take at our model's F1-scores.
> 🔑 **Note:** We could've also made these comparisons in TensorBoard using the [`TensorBoard`](https://www.tensorflow.org/api_docs/python/tf/keras/callbacks/TensorBoard) callback during training.
```
# Sort model results by f1-score
all_model_results.sort_values("f1", ascending=False)["f1"].plot(kind="bar", figsize=(10, 7));
```
Nice! Based on F1-scores, it looks like our tribrid embedding model performs the best by a fair margin.
Though, in comparison to the results reported in Table 3 of the [*PubMed 200k RCT:
a Dataset for Sequential Sentence Classification in Medical Abstracts*](https://arxiv.org/pdf/1710.06071.pdf) paper, our model's F1-score is still underperforming (the authors model achieves an F1-score of 90.0 on the 20k RCT dataset versus our F1-score of ~82.6).
There are some things to note about this difference:
* Our models (with an exception for the baseline) have been trained on ~18,000 (10% of batches) samples of sequences and labels rather than the full ~180,000 in the 20k RCT dataset.
* This is often the case in machine learning experiments though, make sure training works on a smaller number of samples, then upscale when needed (an extension to this project will be training a model on the full dataset).
* Our model's prediction performance levels have been evaluated on the validation dataset not the test dataset (we'll evaluate our best model on the test dataset shortly).
## Save and load best performing model
Since we've been through a fair few experiments, it's a good idea to save our best performing model so we can reuse it without having to retrain it.
We can save our best performing model by calling the [`save()`](https://www.tensorflow.org/guide/keras/save_and_serialize#the_short_answer_to_saving_loading) method on it.
```
# Save best performing model to SavedModel format (default)
model_5.save("skimlit_tribrid_model") # model will be saved to path specified by string
```
Optional: If you're using Google Colab, you might want to copy your saved model to Google Drive (or [download it](https://colab.research.google.com/notebooks/io.ipynb#scrollTo=hauvGV4hV-Mh)) for more permanent storage (Google Colab files disappear after you disconnect).
```
# Example of copying saved model from Google Colab to Drive (requires Google Drive to be mounted)
# !cp skim_lit_best_model -r /content/drive/MyDrive/tensorflow_course/skim_lit
```
Like all good cooking shows, we've got a pretrained model (exactly the same kind of model we built for `model_5` [saved and stored on Google Storage](https://storage.googleapis.com/ztm_tf_course/skimlit/skimlit_best_model.zip)).
So to make sure we're all using the same model for evaluation, we'll download it and load it in.
And when loading in our model, since it uses a couple of [custom objects](https://www.tensorflow.org/guide/keras/save_and_serialize#custom_objects) (our TensorFlow Hub layer and `TextVectorization` layer), we'll have to load it in by specifying them in the `custom_objects` parameter of [`tf.keras.models.load_model()`](https://www.tensorflow.org/api_docs/python/tf/keras/models/load_model).
```
# Download pretrained model from Google Storage
!wget https://storage.googleapis.com/ztm_tf_course/skimlit/skimlit_tribrid_model.zip
!mkdir skimlit_gs_model
!unzip skimlit_tribrid_model.zip -d skimlit_gs_model
# Import TensorFlow model dependencies (if needed) - https://github.com/tensorflow/tensorflow/issues/38250
import tensorflow_hub as hub
import tensorflow as tf
from tensorflow.keras.layers.experimental.preprocessing import TextVectorization
model_path = "skimlit_gs_model/skimlit_tribrid_model"
# Load downloaded model from Google Storage
loaded_model = tf.keras.models.load_model(model_path,
custom_objects={"TextVectorization": TextVectorization, # required for char vectorization
"KerasLayer": hub.KerasLayer}) # required for token embedding
```
### Make predictions and evalaute them against the truth labels
To make sure our model saved and loaded correctly, let's make predictions with it, evaluate them and then compare them to the prediction results we calculated earlier.
```
# Make predictions with the loaded model on the validation set
loaded_pred_probs = loaded_model.predict(val_pos_char_token_dataset, verbose=1)
loaded_preds = tf.argmax(loaded_pred_probs, axis=1)
loaded_preds[:10]
# Evaluate loaded model's predictions
loaded_model_results = calculate_results(val_labels_encoded,
loaded_preds)
loaded_model_results
```
Now let's compare our loaded model's predictions with the prediction results we obtained before saving our model.
```
# Compare loaded model results with original trained model results (should return no errors)
assert model_5_results == loaded_model_results
```
It's worth noting that loading in a SavedModel unfreezes all layers (makes them all trainable). So if you want to freeze any layers, you'll have to set their trainable attribute to `False`.
```
# Check loaded model summary (note the number of trainable parameters)
loaded_model.summary()
```
## Evaluate model on test dataset
To make our model's performance more comparable with the results reported in Table 3 of the [*PubMed 200k RCT:
a Dataset for Sequential Sentence Classification in Medical Abstracts*](https://arxiv.org/pdf/1710.06071.pdf) paper, let's make predictions on the test dataset and evaluate them.
```
# Create test dataset batch and prefetched
test_pos_char_token_data = tf.data.Dataset.from_tensor_slices((test_line_numbers_one_hot,
test_total_lines_one_hot,
test_sentences,
test_chars))
test_pos_char_token_labels = tf.data.Dataset.from_tensor_slices(test_labels_one_hot)
test_pos_char_token_dataset = tf.data.Dataset.zip((test_pos_char_token_data, test_pos_char_token_labels))
test_pos_char_token_dataset = test_pos_char_token_dataset.batch(32).prefetch(tf.data.AUTOTUNE)
# Check shapes
test_pos_char_token_dataset
# Make predictions on the test dataset
test_pred_probs = loaded_model.predict(test_pos_char_token_dataset,
verbose=1)
test_preds = tf.argmax(test_pred_probs, axis=1)
test_preds[:10]
# Evaluate loaded model test predictions
loaded_model_test_results = calculate_results(y_true=test_labels_encoded,
y_pred=test_preds)
loaded_model_test_results
```
It seems our best model (so far) still has some ways to go to match the performance of the results in the paper (their model gets 90.0 F1-score on the test dataset, where as ours gets ~82.1 F1-score).
However, as we discussed before our model has only been trained on 20,000 out of the total ~180,000 sequences in the RCT 20k dataset. We also haven't fine-tuned our pretrained embeddings (the paper fine-tunes GloVe embeddings). So there's a couple of extensions we could try to improve our results.
## Find most wrong
One of the best ways to investigate where your model is going wrong (or potentially where your data is wrong) is to visualize the "most wrong" predictions.
The most wrong predictions are samples where the model has made a prediction with a high probability but has gotten it wrong (the model's prediction disagreess with the ground truth label).
Looking at the most wrong predictions can give us valuable information on how to improve further models or fix the labels in our data.
Let's write some code to help us visualize the most wrong predictions from the test dataset.
First we'll convert all of our integer-based test predictions into their string-based class names.
```
%%time
# Get list of class names of test predictions
test_pred_classes = [label_encoder.classes_[pred] for pred in test_preds]
test_pred_classes
```
Now we'll enrich our test DataFame with a few values:
* A `"prediction"` (string) column containing our model's prediction for a given sample.
* A `"pred_prob"` (float) column containing the model's maximum prediction probabiliy for a given sample.
* A `"correct"` (bool) column to indicate whether or not the model's prediction matches the sample's target label.
```
# Create prediction-enriched test dataframe
test_df["prediction"] = test_pred_classes # create column with test prediction class names
test_df["pred_prob"] = tf.reduce_max(test_pred_probs, axis=1).numpy() # get the maximum prediction probability
test_df["correct"] = test_df["prediction"] == test_df["target"] # create binary column for whether the prediction is right or not
test_df.head(20)
```
Looking good! Having our data like this, makes it very easy to manipulate and view in different ways.
How about we sort our DataFrame to find the samples with the highest `"pred_prob"` and where the prediction was wrong (`"correct" == False`)?
```
# Find top 100 most wrong samples (note: 100 is an abitrary number, you could go through all of them if you wanted)
top_100_wrong = test_df[test_df["correct"] == False].sort_values("pred_prob", ascending=False)[:100]
top_100_wrong
```
Great (or not so great)! Now we've got a subset of our model's most wrong predictions, let's write some code to visualize them.
```
# Investigate top wrong preds
for row in top_100_wrong[0:10].itertuples(): # adjust indexes to view different samples
_, target, text, line_number, total_lines, prediction, pred_prob, _ = row
print(f"Target: {target}, Pred: {prediction}, Prob: {pred_prob}, Line number: {line_number}, Total lines: {total_lines}\n")
print(f"Text:\n{text}\n")
print("-----\n")
```
What do you notice about the most wrong predictions? Does the model make silly mistakes? Or are some of the labels incorrect/ambiguous (e.g. a line in an abstract could potentially be labelled `OBJECTIVE` or `BACKGROUND` and make sense).
A next step here would be if there are a fair few samples with inconsistent labels, you could go through your training dataset, update the labels and then retrain a model. The process of using a model to help improve/investigate your dataset's labels is often referred to as **active learning**.
## Make example predictions
Okay, we've made some predictions on the test dataset, now's time to really test our model out.
To do so, we're going to get some data from the wild and see how our model performs.
In other words, were going to find an RCT abstract from PubMed, preprocess the text so it works with our model, then pass each sequence in the wild abstract through our model to see what label it predicts.
For an appropriate sample, we'll need to search PubMed for RCT's (randomized controlled trials) without abstracts which have been split up (on exploring PubMed you'll notice many of the abstracts are already preformatted into separate sections, this helps dramatically with readability).
Going through various PubMed studies, I managed to find the following unstructured abstract from [*RCT of a manualized social treatment for high-functioning autism spectrum disorders*](https://pubmed.ncbi.nlm.nih.gov/20232240/):
> This RCT examined the efficacy of a manualized social intervention for children with HFASDs. Participants were randomly assigned to treatment or wait-list conditions. Treatment included instruction and therapeutic activities targeting social skills, face-emotion recognition, interest expansion, and interpretation of non-literal language. A response-cost program was applied to reduce problem behaviors and foster skills acquisition. Significant treatment effects were found for five of seven primary outcome measures (parent ratings and direct child measures). Secondary measures based on staff ratings (treatment group only) corroborated gains reported by parents. High levels of parent, child and staff satisfaction were reported, along with high levels of treatment fidelity. Standardized effect size estimates were primarily in the medium and large ranges and favored the treatment group.
Looking at the large chunk of text can seem quite intimidating. Now imagine you're a medical researcher trying to skim through the literature to find a study relevant to your work.
Sounds like quite the challenge right?
Enter SkimLit 🤓🔥!
Let's see what our best model so far (`model_5`) makes of the above abstract.
But wait...
As you might've guessed the above abstract hasn't been formatted in the same structure as the data our model has been trained on. Therefore, before we can make a prediction on it, we need to preprocess it just as we have our other sequences.
More specifically, for each abstract, we'll need to:
1. Split it into sentences (lines).
2. Split it into characters.
3. Find the number of each line.
4. Find the total number of lines.
Starting with number 1, there are a couple of ways to split our abstracts into actual sentences. A simple one would be to use Python's in-built `split()` string method, splitting the abstract wherever a fullstop appears. However, can you imagine where this might go wrong?
Another more advanced option would be to leverage [spaCy's](https://spacy.io/) (a very powerful NLP library) [`sentencizer`](https://spacy.io/usage/linguistic-features#sbd) class. Which is an easy to use sentence splitter based on spaCy's English language model.
I've prepared some abstracts from PubMed RCT papers to try our model on, we can download them [from GitHub](https://raw.githubusercontent.com/mrdbourke/tensorflow-deep-learning/main/extras/skimlit_example_abstracts.json).
```
# Download and open example abstracts (copy and pasted from PubMed)
!wget https://raw.githubusercontent.com/mrdbourke/tensorflow-deep-learning/main/extras/skimlit_example_abstracts.json
with open("skimlit_example_abstracts.json", "r") as f:
example_abstracts = json.load(f)
example_abstracts
# See what our example abstracts look like
abstracts = pd.DataFrame(example_abstracts)
abstracts
```
Now we've downloaded some example abstracts, let's see how one of them goes with our trained model.
First, we'll need to parse it using spaCy to turn it from a big chunk of text into sentences.
```
# Create sentencizer - Source: https://spacy.io/usage/linguistic-features#sbd
from spacy.lang.en import English
nlp = English() # setup English sentence parser
sentencizer = nlp.create_pipe("sentencizer") # create sentence splitting pipeline object
nlp.add_pipe(sentencizer) # add sentence splitting pipeline object to sentence parser
doc = nlp(example_abstracts[0]["abstract"]) # create "doc" of parsed sequences, change index for a different abstract
abstract_lines = [str(sent) for sent in list(doc.sents)] # return detected sentences from doc in string type (not spaCy token type)
abstract_lines
```
Beautiful! It looks like spaCy has split the sentences in the abstract correctly. However, it should be noted, there may be more complex abstracts which don't get split perfectly into separate sentences (such as the example in [*Baclofen promotes alcohol abstinence in alcohol dependent cirrhotic patients with hepatitis C virus (HCV) infection*](https://pubmed.ncbi.nlm.nih.gov/22244707/)), in this case, more custom splitting techniques would have to be investigated.
Now our abstract has been split into sentences, how about we write some code to count line numbers as well as total lines.
To do so, we can leverage some of the functionality of our `preprocess_text_with_line_numbers()` function.
```
# Get total number of lines
total_lines_in_sample = len(abstract_lines)
# Go through each line in abstract and create a list of dictionaries containing features for each line
sample_lines = []
for i, line in enumerate(abstract_lines):
sample_dict = {}
sample_dict["text"] = str(line)
sample_dict["line_number"] = i
sample_dict["total_lines"] = total_lines_in_sample - 1
sample_lines.append(sample_dict)
sample_lines
```
Now we've got `"line_number"` and `"total_lines"` values, we can one-hot encode them with `tf.one_hot` just like we did with our training dataset (using the same values for the `depth` parameter).
```
# Get all line_number values from sample abstract
test_abstract_line_numbers = [line["line_number"] for line in sample_lines]
# One-hot encode to same depth as training data, so model accepts right input shape
test_abstract_line_numbers_one_hot = tf.one_hot(test_abstract_line_numbers, depth=15)
test_abstract_line_numbers_one_hot
# Get all total_lines values from sample abstract
test_abstract_total_lines = [line["total_lines"] for line in sample_lines]
# One-hot encode to same depth as training data, so model accepts right input shape
test_abstract_total_lines_one_hot = tf.one_hot(test_abstract_total_lines, depth=20)
test_abstract_total_lines_one_hot
```
We can also use our `split_chars()` function to split our abstract lines into characters.
```
# Split abstract lines into characters
abstract_chars = [split_chars(sentence) for sentence in abstract_lines]
abstract_chars
```
Alright, now we've preprocessed our wild RCT abstract into all of the same features our model was trained on, we can pass these features to our model and make sequence label predictions!
```
# Make predictions on sample abstract features
%%time
test_abstract_pred_probs = loaded_model.predict(x=(test_abstract_line_numbers_one_hot,
test_abstract_total_lines_one_hot,
tf.constant(abstract_lines),
tf.constant(abstract_chars)))
test_abstract_pred_probs
# Turn prediction probabilities into prediction classes
test_abstract_preds = tf.argmax(test_abstract_pred_probs, axis=1)
test_abstract_preds
```
Now we've got the predicted sequence label for each line in our sample abstract, let's write some code to visualize each sentence with its predicted label.
```
# Turn prediction class integers into string class names
test_abstract_pred_classes = [label_encoder.classes_[i] for i in test_abstract_preds]
test_abstract_pred_classes
# Visualize abstract lines and predicted sequence labels
for i, line in enumerate(abstract_lines):
print(f"{test_abstract_pred_classes[i]}: {line}")
```
Nice! Isn't that much easier to read? I mean, it looks like our model's predictions could be improved, but how cool is that?
Imagine implementing our model to the backend of the PubMed website to format any unstructured RCT abstract on the site.
Or there could even be a browser extension, called "SkimLit" which would add structure (powered by our model) to any unstructured RCT abtract.
And if showed your medical researcher friend, and they thought the predictions weren't up to standard, there could be a button saying "is this label correct?... if not, what should it be?". That way the dataset, along with our model's future predictions, could be improved over time.
Of course, there are many more ways we could go to improve the model, the usuability, the preprocessing functionality (e.g. functionizing our sample abstract preprocessing pipeline) but I'll leave these for the exercises/extensions.
> 🤔 **Question:** How can we be sure the results of our test example from the wild are truly *wild*? Is there something we should check about the sample we're testing on?
## 🛠 Exercises
1. Train `model_5` on all of the data in the training dataset for as many epochs until it stops improving. Since this might take a while, you might want to use:
* [`tf.keras.callbacks.ModelCheckpoint`](https://www.tensorflow.org/api_docs/python/tf/keras/callbacks/ModelCheckpoint) to save the model's best weights only.
* [`tf.keras.callbacks.EarlyStopping`](https://www.tensorflow.org/api_docs/python/tf/keras/callbacks/EarlyStopping) to stop the model from training once the validation loss has stopped improving for ~3 epochs.
2. Checkout the [Keras guide on using pretrained GloVe embeddings](https://keras.io/examples/nlp/pretrained_word_embeddings/). Can you get this working with one of our models?
* Hint: You'll want to incorporate it with a custom token [Embedding](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Embedding) layer.
* It's up to you whether or not you fine-tune the GloVe embeddings or leave them frozen.
3. Try replacing the TensorFlow Hub Universal Sentence Encoder pretrained embedding for the [TensorFlow Hub BERT PubMed expert](https://tfhub.dev/google/experts/bert/pubmed/2) (a language model pretrained on PubMed texts) pretrained embedding. Does this effect results?
* Note: Using the BERT PubMed expert pretrained embedding requires an extra preprocessing step for sequences (as detailed in the [TensorFlow Hub guide](https://tfhub.dev/google/experts/bert/pubmed/2)).
* Does the BERT model beat the results mentioned in this paper? https://arxiv.org/pdf/1710.06071.pdf
4. What happens if you were to merge our `line_number` and `total_lines` features for each sequence? For example, created a `X_of_Y` feature instead? Does this effect model performance?
* Another example: `line_number=1` and `total_lines=11` turns into `line_of_X=1_of_11`.
5. Write a function (or series of functions) to take a sample abstract string, preprocess it (in the same way our model has been trained), make a prediction on each sequence in the abstract and return the abstract in the format:
* `PREDICTED_LABEL`: `SEQUENCE`
* `PREDICTED_LABEL`: `SEQUENCE`
* `PREDICTED_LABEL`: `SEQUENCE`
* `PREDICTED_LABEL`: `SEQUENCE`
* ...
* You can find your own unstrcutured RCT abstract from PubMed or try this one from: [*Baclofen promotes alcohol abstinence in alcohol dependent cirrhotic patients with hepatitis C virus (HCV) infection*](https://pubmed.ncbi.nlm.nih.gov/22244707/).
## 📖 Extra-curriculum
* For more on working with text/spaCy, see [spaCy's advanced NLP course](https://course.spacy.io/en/). If you're going to be working on production-level NLP problems, you'll probably end up using spaCy.
* For another look at how to approach a text classification problem like the one we've just gone through, I'd suggest going through [Google's Machine Learning Course for text classification](https://developers.google.com/machine-learning/guides/text-classification).
* Since our dataset has imbalanced classes (as with many real-world datasets), so it might be worth looking into the [TensorFlow guide for different methods to training a model with imbalanced classes](https://www.tensorflow.org/tutorials/structured_data/imbalanced_data).
| github_jupyter |
# Random Forest Classification with MaxAbsScaler
This Code template is for the Classification tasks using a simple RandomForestClassifier based on the Ensemble Learning technique along with data rescaling using MaxAbsScaler. Random Forest is a meta estimator that fits multiple decision trees and uses averaging to improve the predictive accuracy and control over-fitting.
### Required Packages
```
import warnings
import numpy as np
import pandas as pd
import seaborn as se
import matplotlib.pyplot as plt
from sklearn.preprocessing import LabelEncoder,MaxAbsScaler
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import classification_report,plot_confusion_matrix
warnings.filterwarnings('ignore')
```
### Initialization
Filepath of CSV file
```
#filepath
file_path= ""
```
List of features which are required for model training .
```
#x_values
features=[]
```
Target feature for prediction.
```
#y_value
target=''
```
### Data Fetching
Pandas is an open-source, BSD-licensed library providing high-performance, easy-to-use data manipulation and data analysis tools.
We will use panda's library to read the CSV file using its storage path.And we use the head function to display the initial row or entry.
```
df=pd.read_csv(file_path)
df.head()
```
### Feature Selections
It is the process of reducing the number of input variables when developing a predictive model. Used to reduce the number of input variables to both reduce the computational cost of modelling and, in some cases, to improve the performance of the model.
We will assign all the required input features to X and target/outcome to Y.
```
X = df[features]
Y = df[target]
```
### Data Preprocessing
Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
```
def NullClearner(df):
if(isinstance(df, pd.Series) and (df.dtype in ["float64","int64"])):
df.fillna(df.mean(),inplace=True)
return df
elif(isinstance(df, pd.Series)):
df.fillna(df.mode()[0],inplace=True)
return df
else:return df
def EncodeX(df):
return pd.get_dummies(df)
def EncodeY(df):
if len(df.unique())<=2:
return df
else:
un_EncodedT=np.sort(pd.unique(df), axis=-1, kind='mergesort')
df=LabelEncoder().fit_transform(df)
EncodedT=[xi for xi in range(len(un_EncodedT))]
print("Encoded Target: {} to {}".format(un_EncodedT,EncodedT))
return df
x=X.columns.to_list()
for i in x:
X[i]=NullClearner(X[i])
X=EncodeX(X)
Y=EncodeY(NullClearner(Y))
X.head()
```
#### Correlation Map
In order to check the correlation between the features, we will plot a correlation matrix. It is effective in summarizing a large amount of data where the goal is to see patterns.
```
f,ax = plt.subplots(figsize=(18, 18))
matrix = np.triu(X.corr())
se.heatmap(X.corr(), annot=True, linewidths=.5, fmt= '.1f',ax=ax, mask=matrix)
plt.show()
```
#### Distribution Of Target Variable
```
plt.figure(figsize = (10,6))
se.countplot(Y)
```
### Data Rescaling
For rescaling the data **MaxAbsScaler** function of Sklearn is used.
MaxAbsScaler scales each feature by its maximum absolute value.
This estimator scales and translates each feature individually such that the maximal absolute value of each feature in the training set will be 1.0. It does not shift/center the data, and thus does not destroy any sparsity.
This scaler can also be applied to sparse CSR or CSC matrices.
##### For more information on MaxAbsScaler [ click here](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.MaxAbsScaler.html)
```
X_MaxAbs=MaxAbsScaler().fit_transform(X)
X_MaxAbs=pd.DataFrame(data = X_MaxAbs,columns = X.columns)
X_MaxAbs.head()
```
### Data Splitting
The train-test split is a procedure for evaluating the performance of an algorithm. The procedure involves taking a dataset and dividing it into two subsets. The first subset is utilized to fit/train the model. The second subset is used for prediction. The main motive is to estimate the performance of the model on new data.
```
X_train, X_test, y_train, y_test = train_test_split(X, Y, test_size = 0.2, random_state = 123)#performing datasplitting
```
### Model
A random forest is a meta estimator that fits a number of decision tree classifiers on various sub-samples of the dataset and uses averaging to improve the predictive accuracy and control over-fitting. The sub-sample size is controlled with the <code>max_samples</code> parameter if <code>bootstrap=True</code> (default), otherwise the whole dataset is used to build each tree.
#### Model Tuning Parameters
1. n_estimators : int, default=100
> The number of trees in the forest.
2. criterion : {“gini”, “entropy”}, default=”gini”
> The function to measure the quality of a split. Supported criteria are “gini” for the Gini impurity and “entropy” for the information gain.
3. max_depth : int, default=None
> The maximum depth of the tree.
4. max_features : {“auto”, “sqrt”, “log2”}, int or float, default=”auto”
> The number of features to consider when looking for the best split:
5. bootstrap : bool, default=True
> Whether bootstrap samples are used when building trees. If False, the whole dataset is used to build each tree.
6. oob_score : bool, default=False
> Whether to use out-of-bag samples to estimate the generalization accuracy.
7. n_jobs : int, default=None
> The number of jobs to run in parallel. fit, predict, decision_path and apply are all parallelized over the trees. <code>None</code> means 1 unless in a joblib.parallel_backend context. <code>-1</code> means using all processors. See Glossary for more details.
8. random_state : int, RandomState instance or None, default=None
> Controls both the randomness of the bootstrapping of the samples used when building trees (if <code>bootstrap=True</code>) and the sampling of the features to consider when looking for the best split at each node (if <code>max_features < n_features</code>).
9. verbose : int, default=0
> Controls the verbosity when fitting and predicting.
```
# Build Model here
model = RandomForestClassifier(n_jobs = -1,random_state = 123)
model.fit(X_train, y_train)
```
#### Model Accuracy
score() method return the mean accuracy on the given test data and labels.
In multi-label classification, this is the subset accuracy which is a harsh metric since you require for each sample that each label set be correctly predicted.
```
print("Accuracy score {:.2f} %\n".format(model.score(X_test,y_test)*100))
```
#### Confusion Matrix
A confusion matrix is utilized to understand the performance of the classification model or algorithm in machine learning for a given test set where results are known.
```
plot_confusion_matrix(model,X_test,y_test,cmap=plt.cm.Blues)
```
#### Classification Report
A Classification report is used to measure the quality of predictions from a classification algorithm. How many predictions are True, how many are False.
* **where**:
- Precision:- Accuracy of positive predictions.
- Recall:- Fraction of positives that were correctly identified.
- f1-score:- percent of positive predictions were correct
- support:- Support is the number of actual occurrences of the class in the specified dataset.
```
print(classification_report(y_test,model.predict(X_test)))
```
#### Feature Importances.
The Feature importance refers to techniques that assign a score to features based on how useful they are for making the prediction.
```
plt.figure(figsize=(8,6))
n_features = len(X.columns)
plt.barh(range(n_features), model.feature_importances_, align='center')
plt.yticks(np.arange(n_features), X.columns)
plt.xlabel("Feature importance")
plt.ylabel("Feature")
plt.ylim(-1, n_features)
```
#### Creator: Saharsh Laud , Github: [Profile](https://github.com/SaharshLaud)
| github_jupyter |
```
from __future__ import print_function
import keras
from keras.datasets import mnist
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
from keras.layers import Conv2D, MaxPooling2D
from keras import backend as K
batch_size = 128
num_classes = 10
epochs = 10
# input image dimensions
img_rows, img_cols = 28, 28
# the data, split between train and test sets
(x_train, y_train), (x_test, y_test) = mnist.load_data()
if K.image_data_format() == 'channels_first':
x_train = x_train.reshape(x_train.shape[0], 1, img_rows, img_cols)
x_test = x_test.reshape(x_test.shape[0], 1, img_rows, img_cols)
input_shape = (1, img_rows, img_cols)
else:
x_train = x_train.reshape(x_train.shape[0], img_rows, img_cols, 1)
x_test = x_test.reshape(x_test.shape[0], img_rows, img_cols, 1)
input_shape = (img_rows, img_cols, 1)
x_train = x_train.astype('float32')[:1000]
x_test = x_test.astype('float32')
x_train /= 255
x_test /= 255
print('x_train shape:', x_train.shape)
print(x_train.shape[0], 'train samples')
print(x_test.shape[0], 'test samples')
# convert class vectors to binary class matrices
y_train = keras.utils.to_categorical(y_train[:1000], num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
model = Sequential()
model.add(Conv2D(32, kernel_size=(3, 3),
activation='relu',
input_shape=input_shape))
model.add(Conv2D(64, (3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(num_classes, activation='softmax'))
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
model.summary()
model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
score = model.evaluate(x_test, y_test, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
def lr_schedule(epoch):
"""Learning Rate Schedule
Learning rate is scheduled to be reduced after 80, 120, 160, 180 epochs.
Called automatically every epoch as part of callbacks during training.
# Arguments
epoch (int): The number of epochs
# Returns
lr (float32): learning rate
"""
lr = 1e-3
if epoch > 180:
lr *= 0.5e-3
elif epoch > 160:
lr *= 1e-3
elif epoch > 120:
lr *= 1e-2
elif epoch > 80:
lr *= 1e-1
print('Learning rate: ', lr)
return lr
'''
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
'''
model.compile(loss='categorical_crossentropy',
optimizer=keras.optimizers.Adadelta(lr=lr_schedule(0)),
metrics=['accuracy'])
from keras.callbacks import ModelCheckpoint, LearningRateScheduler
from keras.callbacks import ReduceLROnPlateau
import numpy as np
lr_scheduler = LearningRateScheduler(lr_schedule)
lr_reducer = ReduceLROnPlateau(factor=np.sqrt(0.1),
cooldown=0,
patience=5,
min_lr=0.5e-6)
callbacks = [lr_scheduler, lr_reducer]
model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test),
callbacks=callbacks)
score = model.evaluate(x_test, y_test, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
```
| github_jupyter |
# ShampooSalesTimeSeries
## 1. Introduction and algorithm description
This notebook uses the shampoo sales dataset to demonstrate the time series algorithms below which are provided by the hana_ml.
- ARIMA
- Auto ARIMA
- Auto Exponential Smoothing
- Seasonal Decompose
### - ARIMA
The Auto Regressive Integrated Moving Average (ARIMA) algorithm is famous in econometrics, statistics and time series analysis.
There are three integers (p, d, q) that are used to parametrize ARIMA models. Because of that, a nonseasonal ARIMA model is denoted with ARIMA(p, d, q):
- p is the number of autoregressive terms (AR part). It allows to incorporate the effect of past values into our model. Intuitively, this would be similar to stating that it is likely to be warm tomorrow if it has been warm the past 3 days.
- d is the number of nonseasonal differences needed for stationarity. Intuitively, this would be similar to stating that it is likely to be same temperature tomorrow if the difference in temperature in the last three days has been very small.
- q is the number of lagged forecast errors in the prediction equation (MA part). This allows us to set the error of our model as a linear combination of the error values observed at previous time points in the past.
When dealing with seasonal effects, Seasonal ARIMA(SARIMA) is used, which is denoted as ARIMA(p,d,q)(P,D,Q,s). Here, p, d, q are the nonseasonal parameters described above, P, D, Q follow the same definition but are applied to the seasonal component of the time series. The term s is the periodicity of the time series.
### - Auto ARIMA
Although the ARIMA model is useful and powerful in time series analysis, it is somehow difficult to choose appropriate orders. Hence, auto ARIMA is to determine the orders of an ARIMA model automatically.
### - Auto Exponential Smoothing
Auto exponential smoothing is used to calculate optimal parameters of a set of smoothing functions, including Single Exponential Smoothing, Double Exponential Smoothing, and Triple Exponential Smoothing.
### - Seasonal Decompose
The algorithm is to decompose a time series into three components: seasonal, trend, and random.
## 2. Dataset
Shampoo sales dataset describes the monthly number of sales of shampoo over a 3 year period.
The units are a sales count and there are 36 observations. The original dataset is credited to Makridakis, Wheelwright and Hyndman (1998). We can see that the dataset shows an increasing trend and possibly has a seasonal component.
<img src="images/Shampoo-Sales.png" title="Temperatures" width="600" height="1200" />
Dataset source: https://raw.githubusercontent.com/jbrownlee/Datasets/master/shampoo.csv for tutorials use only.
### Attribute information
- ID: ID
- SALES: Monthly sales
## 3. Data Loading
### Import packages
First, import packages needed in the data loading.
```
from hana_ml import dataframe
from data_load_utils import DataSets, Settings
```
### Setup Connection
In our case, the data is loaded into a table called "SHAMPOO_SALES_DATA_TBL" in HANA from a csv file "shampoo.csv".
To do that, a connection to HANA is created and then passed to the data loader.
To create a such connection, a config file, <b>config/e2edata.ini</b> is used to control the connection parameters.
A sample section in the config file is shown below which includes HANA url, port, user and password information.
#########################<br>
[hana]<br>
url=host-url<br>
user=username<br>
passwd=userpassword<br>
port=3xx15<br>
#########################<br>
```
url, port, user, pwd = Settings.load_config("../../config/e2edata.ini")
connection_context = dataframe.ConnectionContext(url, port, user, pwd)
```
### Load Data
Then, the function DataSets.load_shampoo_data() is used to decide load or reload the data from scratch. If it is the first time to load data, an exmaple of return message is shown below:
##################<br>
ERROR:hana_ml.dataframe:Failed to get row count for the current Dataframe, (259, 'invalid table name: Could not find table/view SHAMPOO_SALES_DATA_TBL in schema XIN: line 1 col 37 (at pos 36)')
Table SHAMPOO_SALES_DATA_TBL doesn't exist in schema XIN
Creating table SHAMPOO_SALES_DATA_TBL in schema XIN ....
Drop unsuccessful
Creating table XIN.SHAMPOO_SALES_DATA_TBL
Data Loaded:100%
###################<br>
If the data is already loaded, there would be a return message "Table XXX exists and data exists".
```
data_tbl = DataSets.load_shampoo_data(connection_context)
```
### Create Dataframes
Create a dataframe df from SHAMPOO_SALES_DATA_TBL for the following steps.
```
df = connection_context.table(data_tbl)
```
### Simple Data Exploration
We will do some data exploration to know the data better.
- First 3 data points
```
df.collect().head(3)
```
- Columns
```
print(df.columns)
```
- No. of data points
```
print('Number of rows in df: {}'.format(df.count()))
```
- Data types
```
df.dtypes()
```
## 4. Analysis
In this section, various time series algorithms are applied to analyze the shampoo sales dataset.
### 4.1 Seasonal Decompose
Because the dataset shows an increasing trend and possibly some seasonal component, we first use seasonal decompose function to decompose the data.
```
from hana_ml.algorithms.pal.tsa.seasonal_decompose import seasonal_decompose
stats, decompose = seasonal_decompose(df, endog= 'SALES', alpha = 0.2, thread_ratio=0.5)
```
seasonal decompose function returns two tables: stats and decompose.
```
stats.collect()
```
We could see the data has a seasonality and its period is 2. The corresponding multiplicative seasonality model is identified. The decompose table shows the components.
```
decompose.collect().head(5)
```
### 4.2 ARIMA
import the ARIMA module
```
from hana_ml.algorithms.pal.tsa.arima import ARIMA
```
Create an ARIMA estimator and make the initialization:
```
arima = ARIMA(order=(1, 0, 0), seasonal_order=(1, 0, 0, 2),
method='mle', thread_ratio=1.0)
```
Perform fit on the given data:
```
arima.fit(df, endog='SALES')
```
There are two attributes of ARIMA model: model_ and fitted_. We could see the model parameters in model_.
```
arima.model_.collect()
```
The model_ contains AIC (Akaike Information Criterion) and BIC (Bayes Information Criterion) that can be minimized to select the best fitting model.
```
arima.fitted_.collect().set_index('ID').head(5)
```
Predict uisng the ARIMA model:
```
result = arima.predict(forecast_method='innovations_algorithm',forecast_length=5)
result.collect()
%matplotlib inline
from hana_ml.visualizers.visualizer_base import forecast_line_plot
ax = forecast_line_plot(pred_data=result.set_index("TIMESTAMP"),
confidence=("LO80", "HI80", "LO95", "HI95"),
max_xticklabels=10)
```
### 4.3 Auto ARIMA
Import auto ARIMA module
```
from hana_ml.algorithms.pal.tsa.auto_arima import AutoARIMA
```
Create an auto ARIMA estimator and make the initialization:
```
autoarima = AutoARIMA(search_strategy=1, allow_linear=1, thread_ratio=1.0)
```
Perform fit on the given data:
```
autoarima.fit(df, endog='SALES')
autoarima.model_.collect()
autoarima.fitted_.collect().set_index('ID').head(6)
```
Predict uisng the auto ARIMA model:
```
result= autoarima.predict(forecast_method='innovations_algorithm', forecast_length=5)
result.collect()
```
### 4.4 Auto Exponential Smoothing
Import auto exponential smoothing module:
```
from hana_ml.algorithms.pal.tsa.exponential_smoothing import AutoExponentialSmoothing
```
Create an auto exponential smoothing estimator and make the initialization:
```
autoexpsmooth = AutoExponentialSmoothing(model_selection=1, forecast_num=3)
```
Perform the fit on the given data:
```
autoexpsmooth.fit_predict(df,endog= 'SALES',)
```
Have a look at the stats_ and it shows the parameters and Triple Exponential SMoothing (TESM) model is selected.
```
autoexpsmooth.stats_.collect()
```
To see the result of smoothing forecast and upper and lower bound in the forecast_:
```
autoexpsmooth.forecast_.collect()
```
## 5. Close Connection
```
connection_context.close()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/TarkanAl-Kazily/awbw_replay_parser/blob/main/AWBW_Replay_Parser_and_Kantbeis_Replay_Archive.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Running the AWBW Replay Parser toolset on Kantbei's Replay Archive
This notebook runs the AWBW Replay Parser to open and load replays from Kantbei's Replay Archive. It's primarily configured to load all the replays in a batch to check for parsing errors or other errors, but it can be used as a base for a strategy analysis on the high tier games in the archive.
See the main project on Github here: [https://github.com/TarkanAl-Kazily/awbw_replay_parser](https://github.com/TarkanAl-Kazily/awbw_replay_parser)
Access Kantbei's dataset from Google Drive here: [https://drive.google.com/drive/folders/1ziWc1mVm9lOydIquCG8EsTb3aYvpdaPp?usp=sharing](https://drive.google.com/drive/folders/1ziWc1mVm9lOydIquCG8EsTb3aYvpdaPp?usp=sharing)
```
# One time setup for this notebook
import os
os.environ['REPO'] = "https://github.com/TarkanAl-Kazily/awbw_replay_parser.git"
os.environ['BRANCH'] = "main"
# Clone the awbw_replay_parser repository
!cd /content/
!rm -rf awbw_replay_parser
!git clone --branch $BRANCH $REPO
!pip install -r awbw_replay_parser/requirements.txt
# Install the awbw_replay module into our runtime
!cd awbw_replay_parser && python setup.py install
```
To use Kantbei's dataset, you need to add a shortcut to the directory to your drive through Google Drive. Then from the Google Colab Files sidebar you can mount your Google Drive, and the data should be accessible under `drive/MyDrive/Kantbei's AWBW Replay Archive`.
```
from logging import exception
import os
import glob
from awbw_replay.replay import AWBWReplay
from awbw_replay.awbw import AWBWGameState, AWBWGameAction
KANTBEI_REPLAY_DIR = "/content/drive/MyDrive/Kantbei's AWBW Replay Archive"
# Glob all the replay archive files. Replace this list with the specific files you'd like to load.
replay_list = glob.glob(os.path.join(KANTBEI_REPLAY_DIR, "*.zip"))
replay_stats = {}
def attempt_parsing_replay(filename):
"""
Tries to parse a given replay file. Returns a dictionary summarizing the result.
Handles all exceptions, making it safe to continue running afterwards.
"""
result = {
"filename": filename,
"actions": None, # Count how many actions are in the file
"turns": None, # Count how many turns are in the file
"states": None, # Count how many states we could iterate through
"players": None, # Count the number of players in the file
"exception": None, # The exception string in the case of failure
"exception_obj": None, # The actual exception object
"success": False, # If we reached the end of parsing all states
}
try:
with AWBWReplay(filename) as replay:
actions = list(replay.actions())
result["actions"] = len(actions)
result["turns"] = len(replay.turns())
result["states"] = 0
state = AWBWGameState(replay_initial=replay.game_info())
result["states"] += 1
result["players"] = len(state.players)
for action in actions:
state = state.apply_action(AWBWGameAction(action))
result["states"] += 1
result["success"] = True
except Exception as e:
result["exception"] = str(e)
result["exception_obj"] = e
print(f"Got exception: {e}")
return result
for filename in replay_list:
if filename in replay_stats and replay_stats[filename]["success"]:
print(f"Skipping already successful file {filename}")
continue
print(f"Testing file {filename}...")
replay_stats[filename] = attempt_parsing_replay(filename)
successful_replays = [stat["filename"] for stat in replay_stats.values() if stat["success"]]
print(f"Successfully parsed states from {len(successful_replays)} files (out of {len(replay_list)}).")
error_list = [result for result in replay_stats.values() if not result["success"]]
for result in error_list:
print(f"{result['filename']} - {result['exception']} at state {result['states']}")
for result in error_list:
print(f"\"{result['filename']}\",")
```
| github_jupyter |
# Baseline Surface Radiation Network (BSRN)
The [Baseline Surface Radiation Network (BSRN)](https://bsrn.awi.de/) is a global network of high-quality solar radiation monitoring stations under the [World Climate Research Programme (WCRP)](https://www.wcrp-climate.org/) {cite:p}`driemel_baseline_2018`.
According to the [World Radiation Monitoring Center (WRMC)](https://bsrn.awi.de/project/objectives/):
> The data [from the BSRN stations] are of primary importance in supporting the validation and confirmation of satellite and computer model estimates of these quantities. At a small number of stations (currently 74 in total, 58 active) in contrasting climatic zones, covering a latitude range from 80°N to 90°S, solar and atmospheric radiation is measured with instruments of the highest available accuracy and with high time resolution.
All BSRN stations are required to meet the basic [station requirements](http://bsrn.awi.de/en/stations/join-bsrn/). A list of activate, inactive, and candidate BSRN stations can be retrieved from the SolarStations [station listing](../station_listing) and are shown below.
```
import pandas as pd
stations = pd.read_csv('solarstations.csv', sep=';', encoding='latin1')
stations = stations[stations['Network'].str.contains('BSRN')]
stations['Time period'] = stations['Time period'].astype(str).replace('nan','')
stations
```
```{margin} Station metadata
Click the plus symbol above to see a table of the stations and their metadata.
```
```
import folium
from folium import plugins
EsriImagery = "https://server.arcgisonline.com/ArcGIS/rest/services/World_Imagery/MapServer/tile/{z}/{y}/{x}"
EsriAttribution = "Tiles © Esri — Source: Esri, i-cubed, USDA, USGS, AEX, GeoEye, Getmapping, Aerogrid, IGN, IGP, UPR-EGP, and the GIS User Community"
# Create Folium map
m = folium.Map(
location=[0, 15],
zoom_start=1, min_zoom=1, max_bounds=True,
control_scale=True, # Adds distance scale in lower left corner
tiles='openstreetmap',
)
# Function for determining station color
def icon_color(row):
if row['Time period'].endswith('-'): # active station
color = 'green'
elif row['Time period'] == '': # candidate station
color = 'gray'
else: # inactive/closed station
color = 'red'
return color
# Add each station to the map
# Consider using apply instead of for loop to add stations in case of many stations
for index, row in stations.iterrows():
folium.Marker(
location=[row['Latitude'], row['Longitude']],
popup=row['Station full name'] + ' - ' + str(row['State']) + ' ' + row['Country'],
tooltip=row['Abbreviation'],
icon=folium.Icon(color=icon_color(row), icon='bolt', prefix='fa')
).add_to(m)
folium.raster_layers.TileLayer(EsriImagery, name='World imagery', attr=EsriAttribution).add_to(m)
folium.LayerControl(position='topright').add_to(m)
# Additional options and plugins
# Note it's not possible to change the position of the scale
plugins.MiniMap(toggle_display=True, zoom_level_fixed=1, minimized=True, position='bottomright').add_to(m) # Add minimap to the map
plugins.Fullscreen(position='topright').add_to(m) # Add full screen button to map
folium.LatLngPopup().add_to(m) # Show latitude/longitude when clicking on the map
# plugins.LocateControl(position='topright').add_to(m) # Add button for your position
# plugins.MeasureControl(position='topleft').add_to(m) # Add distance length measurement tool
# Add Category Legend
legend_html = """
<div style="position:fixed;
top: 10px;
left: 10px;
width: 120px;
height: 85px;
border:2px solid grey;
z-index: 9999;
font-size:14px;">
<b>Station markers</b><br>
<i class="fa fa-circle fa-1x" style="color:green"></i> Active<br>
<i class="fa fa-circle fa-1x" style="color:red"></i> Inactive<br>
<i class="fa fa-circle fa-1x" style="color:gray"></i> Candidate<br>
</div>"""
m.get_root().html.add_child(folium.Element(legend_html)) # Add Legend
# Show the map
m
```
## Station requirements
As a mimimum a BSRN station is required to measure global horizontal irradiance (GHI), direct normal irradiance (DNI), diffuse horizontal irradiance (DHI),
Additional metadata may be found at the [BSRN website](https://wiki.pangaea.de/wiki/BSRN#Sortable_Table_of_Stations) and in the individual data files (e.g., horizon profile).
```{note}
Unlike the majority of solar radiation monitoring networks, the BSRN website does not have a subpage for each station (with photos, etc.). This would have been very useful when assessing the usage of the station, for example in regards to the potential impact of nearby structures, etc. Note a few photos of the BSRN stations can be found [here](https://bsrn.awi.de/other/picture-gallery/). There station log books are also not available. It should also be noted that the files on the FTP server do not include wind speed and direction.
```
## Data retrieval
Data from the BSRN stations is stored in monthly files for each station and can be freely downloaded either via [FTP](https://bsrn.awi.de/?id=387) or [Pangea](https://bsrn.awi.de/data/data-retrieval-via-pangaea/). Credentials for accessing the BSRN FTP server can be obtained as described in the [data release guidelines](https://bsrn.awi.de/data/conditions-of-data-release).
```{admonition} Data release guidelines
Please read the [BSRN data release guidelines](https://bsrn.awi.de/data/conditions-of-data-release/) before using any data and make sure to properly cite the BSRN.
```
```{warning}
WRMC highly recommends that all users do their own quality checks of the data after extracting BSRN-data!
```
The data can also be downloaded programmatically using the [pvlib-python](https://pvlib-python.readthedocs.io) library, specifically the [`get_bsrn`](https://pvlib-python.readthedocs.io/en/latest/generated/pvlib.iotools.get_bsrn.html) function. An example of how to use pvlib to download two months of data from the Cabauw (CAB) station is shown below:
```
import os
bsrn_username = os.environ["BSRN_FTP_USERNAME"]
bsrn_password = os.environ["BSRN_FTP_PASSWORD"]
import pvlib
df, meta = pvlib.iotools.get_bsrn(
station='CAB', # three letter code for the Cabauw station
start=pd.Timestamp(2018,6,1),
end=pd.Timestamp(2018,7,14),
username=bsrn_username, # replace with your own username
password=bsrn_password, # replace with your own password
)
df.head(12)
```
```{margin} Available parameters
Click the plus symbol above to see the first 12 data entries.
```
For a description of the input parameters, see the [pvlib documentation](https://pvlib-python.readthedocs.io/en/latest/generated/pvlib.iotools.get_bsrn.html).
```{admonition} Retrieving BSRN data in R
R users can find similar functionality in the [SolarData](https://github.com/dazhiyang/SolarData) R package.
```
The data retrieved from all BSRN stations includes measurements of the three irradiance components, as well as longwave downwelling irradiance, temperature humidity, etc.
A few of the parameters in the datasets for the month of data are visualized below.
```
axes = df[['ghi','dni','dhi','lwd','temp_air']].plot(
subplots=True, legend=False, rot=0, figsize=(8,8), sharex=True)
# Set y-labels and y-limits
axes[0].set_ylabel('GHI [W/m$^2$]'), axes[0].set_ylim(-10,1300)
axes[1].set_ylabel('DNI [W/m$^2$]'), axes[1].set_ylim(-10,1300)
axes[2].set_ylabel('DHI [W/m$^2$]'), axes[2].set_ylim(-10,1300)
axes[3].set_ylabel('LWD [W/m$^2$]'), axes[4].set_ylim(200,500)
_ = axes[4].set_ylabel('Temperature [°]'), axes[4].set_ylim(0,40)
```
Notice how that they are multiple periods where there is gaps in the irradiance data.
## To do
* Make a list of bad stations
* What is the difference in available parameters between FTP and Pangea?
* Add continent to solarstations.csv
* Add description of minimum requirements for BSRN stations
* Make note that delay of a few years is not uncommon and give link to where they can see the status of the data.
### Done
* Cite the BSRN
* Add legend for colors
* Get correct year of operation from here: https://dataportals.pangaea.de/bsrn/
* Have different color for active and inactive stations
# References
```{bibliography}
:filter: docname in docnames
```
| github_jupyter |
## Python 3 - Sentiment Analysis (Project testing)
### Build the project functions
```
punctuation_chars = ["'", '"', ",", ".", "!", ":", ";", '#', '@']
def strip_punctuation(word):
y = []
for i in word:
if i not in punctuation_chars:
y.append(i)
return "".join(y)
# performing some tests for debugging purposes.
print(strip_punctuation("#in.cred..ible!"))
punctuation_chars = ["'", '"', ",", ".", "!", ":", ";", '#', '@']
# list of positive words to use
# Code won't execute in the notebook since the file is not stored locally.
positive_words = []
with open("positive_words.txt") as pos_f:
for lin in pos_f:
if lin[0] != ';' and lin[0] != '\n':
positive_words.append(lin.strip())
def strip_punctuation(word):
y = []
for i in word:
if i not in punctuation_chars:
y.append(i)
return "".join(y)
# Here we need to start buiding another function in the next cell.
# Get positive words function
def get_pos(sentence):
toLower = sentence.lower()
count = 0
splt = toLower.split()
for w in splt:
rp = strip_punctuation(w)
if rp in positive_words:
count = count + 1
else:
continue
return count
# Get negative words function
def get_neg(sentence):
toLower = sentence.lower()
count = 0
splt = toLower.split()
for w in splt:
rp = strip_punctuation(w)
if rp in negative_words:
count = count + 1
else:
continue
return count
# As well as other funtions, this one refers to files stored elsewhere.
outfile = open("resulting_data.csv", "w")
# Writing outfile header
outfile.write("Number of Retweets, Number of Replies, Positive Score, Negative Score, Net Score")
# Write new line after the header.
outfile.write('\n')
with open("project_twitter_data.csv") as pro:
lines = pro.readlines()
header = lines[0]
# ['tweet_text', 'retweet_count', 'reply_count']
field_names = header.strip().split(",")
for row in lines[1:]:
values = row.strip().split(",")
posScore = get_pos(values[0])
negScore = get_neg(values[0])
retweetNum = values[1]
replyNum = values[2]
netScore = posScore - negScore
outfile.write("{}, {}, {}, {}, {}".format(retweetNum, replyNum, posScore, negScore, netScore))
outfile.write("\n")
outfile.close()
```
### Retrieve information from the csv file, clean, and write on "resulting_data.csv"
```
# Data file: "resulting_data.csv"
Number of Retweets, Number of Replies, Positive Score, Negative Score, Net Score
3, 0, 0, 0, 0
1, 0, 2, 2, 0
1, 2, 1, 0, 1
3, 1, 1, 0, 1
6, 0, 2, 0, 2
9, 5, 2, 0, 2
19, 0, 2, 0, 2
0, 0, 0, 3, -3
0, 0, 0, 2, -2
82, 2, 4, 0, 4
0, 0, 0, 1, -1
0, 0, 1, 0, 1
47, 0, 2, 0, 2
2, 1, 1, 0, 1
0, 2, 1, 0, 1
0, 0, 2, 1, 1
4, 6, 3, 0, 3
19, 0, 3, 1, 2
0, 0, 1, 1, 0
```
### Sentiment Analysis Results

| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.