text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
# Building and training your first TensorFlow graph from the ground up.
- https://www.oreilly.com/learning/hello-tensorflow?imm_mid=0e50a2&cmp=em-data-na-na-newsltr_20160622#
The [TensorFlow](https://www.tensorflow.org/) project is bigger than you might realize. The fact that it's a library for deep learning, and its connection to Google, has helped TensorFlow attract a lot of attention. But beyond the hype, there are unique elements to the project that are worthy of closer inspection:
- The core library is suited to a broad family of machine learning techniques, not “just” deep learning.
- Linear algebra and other internals are prominently exposed.
- In addition to the core machine learning functionality, TensorFlow also includes its own logging system, its own interactive log visualizer, and even its own heavily engineered serving architecture.
- The execution model for TensorFlow differs from Python's scikit-learn, or most tools in R.
Cool stuff, but—especially for someone hoping to explore machine learning for the first time—TensorFlow can be a lot to take in.
How does TensorFlow work? Let's break it down so we can see and understand every moving part. We'll explore the data flow [graph](https://en.wikipedia.org/wiki/Graph_(abstract_data_type)) that defines the computations your data will undergo, how to train models with[gradient descent](https://en.wikipedia.org/wiki/Gradient_descent) using TensorFlow, and how [TensorBoard](https://www.tensorflow.org/versions/r0.8/how_tos/summaries_and_tensorboard/) can visualize your TensorFlow work. The examples here won't solve industrial machine learning problems, but they'll help you understand the components underlying everything built with TensorFlow, including whatever you build next!
### Names and execution in Python and TensorFlow
The way TensorFlow manages computation is not totally different from the way Python usually does. With both, it's important to remember, to paraphrase [Hadley Wickham](https://twitter.com/hadleywickham/status/732288980549390336), that an object has no name (see Figure 1). In order to see the similarities (and differences) between how Python and TensorFlow work, let’s look at how they refer to objects and handle evaluation.
<img src="images/image1.jpg"/>
The variable names in Python code aren't what they represent; they're just pointing at objects. So, when you say in Python that foo = [] and bar = foo, it isn't just that foo equals bar; foo is bar, in the sense that they both point at the same list object.
```
foo = []
bar = foo
foo == bar
## True
foo is bar
## True
```
You can also see that id(foo) and id(bar) are the same. This identity, especially with mutable data structures like lists, can lead to surprising bugs when it's misunderstood.
Internally, Python manages all your objects and keeps track of your variable names and which objects they refer to. The TensorFlow graph represents another layer of this kind of management; as we’ll see, Python names will refer to objects that connect to more granular and managed TensorFlow graph operations.
When you enter a Python expression, for example at an interactive interpreter or Read Evaluate Print Loop (REPL), whatever is read is almost always evaluated right away. Python is eager to do what you tell it. So, if I tell Python to foo.append(bar), it appends right away, even if I never use foo again.
A lazier alternative would be to just remember that I said foo.append(bar), and if I ever evaluate foo at some point in the future, Python could do the append then. This would be closer to how TensorFlow behaves, where defining relationships is entirely separate from evaluating what the results are.
TensorFlow separates the definition of computations from their execution even further by having them happen in separate places: a graph defines the operations, but the operations only happen within a session. Graphs and sessions are created independently. A graph is like a blueprint, and a session is like a construction site.
Back to our plain Python example, recall that foo and bar refer to the same list. By appending bar into foo, we've put a list inside itself. You could think of this structure as a graph with one node, pointing to itself. Nesting lists is one way to represent a graph structure like a TensorFlow computation graph.
```
foo.append(bar)
foo
## [[...]]
```
Real TensorFlow graphs will be more interesting than this!
## The simplest TensorFlow graph
To start getting our hands dirty, let’s create the simplest TensorFlow graph we can, from the ground up. TensorFlow is admirably easier to install than some other frameworks. The examples here work with either Python 2.7 or 3.3+, and the TensorFlow version used is 0.8.
```
import tensorflow as tf
```
At this point TensorFlow has already started managing a lot of state for us. There's already an implicit default graph, for example. Internally, the default graph lives in the **_default_graph_stack**, but we don't have access to that directly. We use **tf.get_default_graph()**.
```
graph = tf.get_default_graph()
```
The nodes of the TensorFlow graph are called “operations,” or “ops.” We can see what operations are in the graph with graph.get_operations().
```
graph.get_operations()
```
Currently, there isn't anything in the graph. We’ll need to put everything we want TensorFlow to compute into that graph. Let's start with a simple constant input value of one.
```
input_value = tf.constant(1.0)
```
That constant now lives as a node, an operation, in the graph. The Python variable name input_value refers indirectly to that operation, but we can also find the operation in the default graph.
```
operations = graph.get_operations()
operations
operations[0].node_def
```
TensorFlow uses protocol buffers internally. (Protocol buffers are sort of like a Google-strength JSON.) Printing the node_def for the constant operation above shows what's in TensorFlow's protocol buffer representation for the number one.
People new to TensorFlow sometimes wonder why there's all this fuss about making “TensorFlow versions” of things. Why can't we just use a normal Python variable without also defining a TensorFlow object? One of the TensorFlow tutorials has an explanation:
> To do efficient numerical computing in Python, we typically use libraries like NumPy that do expensive operations such as matrix multiplication outside Python, using highly efficient code implemented in another language. Unfortunately, there can still be a lot of overhead from switching back to Python every operation. This overhead is especially bad if you want to run computations on GPUs or in a distributed manner, where there can be a high cost to transferring data.
> TensorFlow also does its heavy lifting outside Python, but it takes things a step further to avoid this overhead. Instead of running a single expensive operation independently from Python, TensorFlow lets us describe a graph of interacting operations that run entirely outside Python. This approach is similar to that used in Theano or Torch.
TensorFlow can do a lot of great things, but it can only work with what's been explicitly given to it. This is true even for a single constant.
If we inspect our input_value, we see it is a constant 32-bit float tensor of no dimension: just one number.
```
input_value
```
Note that this doesn't tell us what that number is. To evaluate input_value and get a numerical value out, we need to create a “session” where graph operations can be evaluated and then explicitly ask to evaluate or “run” input_value. (The session picks up the default graph by default.)
```
sess = tf.Session()
sess.run(input_value)
```
It may feel a little strange to “run” a constant. But it isn't so different from evaluating an expression as usual in Python; it's just that TensorFlow is managing its own space of things—the computational graph—and it has its own method of evaluation.
## The simplest TensorFlow neuron
Now that we have a session with a simple graph, let's build a neuron with just one parameter, or weight. Often, even simple neurons also have a bias term and a non-identity activation function, but we'll leave these out.
The neuron's weight isn't going to be constant; we expect it to change in order to learn based on the “true” input and output we use for training. The weight will be a TensorFlow variable. We'll give that variable a starting value of 0.8.
```
weight = tf.Variable(0.8)
```
You might expect that adding a variable would add one operation to the graph, but in fact that one line adds four operations. We can check all the operation names:
```
for op in graph.get_operations(): print op.name
```
We won't want to follow every operation individually for long, but it will be nice to see at least one that feels like a real computation.
```
output_value = weight * input_value
```
Now there are six operations in the graph, and the last one is that multiplication.
```
op = graph.get_operations()[-1]
op.name
for op_input in op.inputs: print op_input
```
This shows how the multiplication operation tracks where its inputs come from: they come from other operations in the graph. To understand a whole graph, following references this way quickly becomes tedious for humans. [TensorBoard graph visualization](https://www.tensorflow.org/versions/r0.8/how_tos/graph_viz/) is designed to help.
How do we find out what the product is? We have to “run” the **output_value** operation. But that operation depends on a variable: **weight**. We told TensorFlow that the initial value of weight should be 0.8, but the value hasn't yet been set in the current session. The **tf.initialize_all_variables()** function generates an operation which will initialize all our variables (in this case just one) and then we can run that operation.
```
init = tf.initialize_all_variables()
sess.run(init)
```
The result of tf.initialize_all_variables() will include initializers for all the variables currently in the graph, so if you add more variables you'll want to use tf.initialize_all_variables() again; a stale init wouldn't include the new variables.
Now we're ready to run the output_value operation.
```
sess.run(output_value)
```
Recall that's 0.8 * 1.0 with 32-bit floats, and 32-bit floats have a hard time with 0.8; 0.80000001 is as close as they can get.
## See your graph in TensorBoard
Up to this point, the graph has been simple, but it would already be nice to see it represented in a diagram. We'll use TensorBoard to generate that diagram. TensorBoard reads the name field that is stored inside each operation (quite distinct from Python variable names). We can use these TensorFlow names and switch to more conventional Python variable names. Using tf.mul here is equivalent to our earlier use of just * for multiplication, but it lets us set the name for the operation.
```
x = tf.constant(1.0, name='input')
w = tf.Variable(0.8, name='weight')
y = tf.mul(w, x, name='output')
```
TensorBoard works by looking at a directory of output created from TensorFlow sessions. We can write this output with a SummaryWriter, and if we do nothing aside from creating one with a graph, it will just write out that graph.
The first argument when creating the SummaryWriter is an output directory name, which will be created if it doesn't exist.
```
summary_writer = tf.train.SummaryWriter('log_simple_graph', sess.graph)
```
Now, at the command line, we can start up TensorBoard.
```
!tensorboard --logdir=log_simple_graph
```
TensorBoard runs as a local web app, on port 6006. (“6006” is “goog” upside-down.) If you go in a browser to localhost:6006/#graphs you should see a diagram of the graph you created in TensorFlow, which looks something like Figure 2.
## Making the neuron learn
Now that we’ve built our neuron, how does it learn? We set up an input value of 1.0. Let's say the correct output value is zero. That is, we have a very simple “training set” of just one example with one feature, which has the value one, and one label, which is zero. We want the neuron to learn the function taking one to zero.
Currently, the system takes the input one and returns 0.8, which is not correct. We need a way to measure how wrong the system is. We'll call that measure of wrongness the “loss” and give our system the goal of minimizing the loss. If the loss can be negative, then minimizing it could be silly, so let's make the loss the square of the difference between the current output and the desired output.
```
y_ = tf.constant(0.0)
loss = (y - y_)**2
```
So far, nothing in the graph does any learning. For that, we need an optimizer. We'll use a gradient descent optimizer so that we can update the weight based on the derivative of the loss. The optimizer takes a learning rate to moderate the size of the updates, which we'll set at 0.025.
```
optim = tf.train.GradientDescentOptimizer(learning_rate=0.025)
```
The optimizer is remarkably clever. It can automatically work out and apply the appropriate gradients through a whole network, carrying out the backward step for learning.
Let's see what the gradient looks like for our simple example.
```
grads_and_vars = optim.compute_gradients(loss)
sess.run(tf.initialize_all_variables())
sess.run(grads_and_vars[1][0])
```
Why is the value of the gradient 1.6? Our loss is error squared, and the derivative of that is two times the error. Currently the system says 0.8 instead of 0, so the error is 0.8, and two times 0.8 is 1.6. It's working!
For more complex systems, it will be very nice indeed that TensorFlow calculates and then applies these gradients for us automatically.
Let's apply the gradient, finishing the backpropagation.
```
sess.run(optim.apply_gradients(grads_and_vars))
sess.run(w)
## 0.75999999 # about 0.76
```
The weight decreased by 0.04 because the optimizer subtracted the gradient times the learning rate, 1.6 * 0.025, pushing the weight in the right direction.
Instead of hand-holding the optimizer like this, we can make one operation that calculates and applies the gradients: the train_step.
```
train_step = tf.train.GradientDescentOptimizer(0.025).minimize(loss)
for i in range(100):
sess.run(train_step)
sess.run(y)
## 0.0044996012
```
Running the training step many times, the weight and the output value are now very close to zero. The neuron has learned!
## Training diagnostics in TensorBoard
We may be interested in what's happening during training. Say we want to follow what our system is predicting at every training step. We could print from inside the training loop.
```
sess.run(tf.initialize_all_variables())
for i in range(100):
print('before step {}, y is {}'.format(i, sess.run(y)))
sess.run(train_step)
```
This works, but there are some problems. It's hard to understand a list of numbers. A plot would be better. And even with only one value to monitor, there's too much output to read. We're likely to want to monitor many things. It would be nice to record everything in some organized way.
Luckily, the same system that we used earlier to visualize the graph also has just the mechanisms we need.
We instrument the computation graph by adding operations that summarize its state. Here, we'll create an operation that reports the current value of y, the neuron's current output.
```
summary_y = tf.scalar_summary('output', y)
```
When you run a summary operation, it returns a string of protocol buffer text that can be written to a log directory with a SummaryWriter.
```
summary_writer = tf.train.SummaryWriter('log_simple_stat')
sess.run(tf.initialize_all_variables())
for i in range(100):
summary_str = sess.run(summary_y)
summary_writer.add_summary(summary_str, i)
sess.run(train_step)
```
Now after running
```
tensorboard --logdir=log_simple_stat
```
you get an interactive plot at localhost:6006/#events (Figure 3).
<img src="images/image2.jpg"/>
## Flowing onward
Here's a final version of the code. It's fairly minimal, with every part showing useful (and understandable) TensorFlow functionality.
```
import tensorflow as tf
x = tf.constant(1.0, name='input')
w = tf.Variable(0.8, name='weight')
y = tf.mul(w, x, name='output')
y_ = tf.constant(0.0, name='correct_value')
loss = tf.pow(y - y_, 2, name='loss')
train_step = tf.train.GradientDescentOptimizer(0.025).minimize(loss)
for value in [x, w, y, y_, loss]:
tf.scalar_summary(value.op.name, value)
summaries = tf.merge_all_summaries()
sess = tf.Session()
summary_writer = tf.train.SummaryWriter('log_simple_stats', sess.graph)
sess.run(tf.initialize_all_variables())
for i in range(100):
summary_writer.add_summary(sess.run(summaries), i)
sess.run(train_step)
```
The example we just ran through is even simpler than the ones that inspired it in Michael Nielsen's [Neural Networks and Deep Learning](http://neuralnetworksanddeeplearning.com/). For myself, seeing details like these helps with understanding and building more complex systems that use and extend from simple building blocks. Part of the beauty of TensorFlow is how flexibly you can build complex systems from simpler components.
If you want to continue experimenting with TensorFlow, it might be fun to start making more interesting neurons, perhaps with different [activation functions](https://en.wikipedia.org/wiki/Activation_function#Comparison_of_activation_functions). You could train with more interesting data. You could add more neurons. You could add more layers. You could dive into more complex[pre-built models](https://github.com/tensorflow/tensorflow/tree/master/tensorflow/models), or spend more time with TensorFlow's own [tutorials](https://www.tensorflow.org/versions/r0.8/tutorials/) and [how-to guides](https://www.tensorflow.org/versions/r0.8/how_tos/). Go for it!
| github_jupyter |
# Example of optimizing a convex function
# Goal is to test the objective values found by Mango
- Search space size: Uniform
- Number of iterations to try: 40
- domain size: 5000
- Initial Random: 5
# Benchmarking test with different iterations for serial executions
```
from mango.tuner import Tuner
from scipy.stats import uniform
def get_param_dict():
param_dict = {
'x': uniform(-5000, 10000)
}
return param_dict
def objfunc(args_list):
results = []
for hyper_par in args_list:
x = hyper_par['x']
result = -(x**2)
results.append(result)
return results
def get_conf_batch_5():
conf = dict()
conf['batch_size'] = 5
conf['initial_random'] = 5
conf['num_iteration'] = 100
conf['domain_size'] = 5000
return conf
def get_optimal_x():
param_dict = get_param_dict()
conf_5 = get_conf_batch_5()
tuner_5 = Tuner(param_dict, objfunc,conf_5)
results_5 = tuner_5.maximize()
return results_5
Store_Optimal_X = []
Store_Results = []
num_of_tries = 100
for i in range(num_of_tries):
results_5 = get_optimal_x()
Store_Results.append([results_5])
print(i,":",results_5['best_params']['x'])
#len(Store_Results)
#Store_Results[0]
```
# Sandeep we need to process the results
# Extract from the results returned the true optimal values for each iteration
```
import numpy as np
total_experiments = 66
initial_random = 5
plotting_itr =[10, 20,30,40,50,60,70,80,90,100]
plotting_list = []
for exp in range(total_experiments): #for all exp
local_list = []
for itr in plotting_itr: # for all points to plot
# find the value of optimal parameters in itr+ initial_random
max_value = np.array(Store_Results[exp][0]['objective_values'][:itr*5+initial_random]).max()
local_list.append(max_value)
plotting_list.append(local_list)
plotting_array = np.array(plotting_list)
plotting_array.shape
#plotting_array
Y = []
#count range between -1 and 1 and show it
for i in range(len(plotting_itr)):
y_value = np.count_nonzero((plotting_array[:,i] > -1))/plotting_array[:,i].size
Y.append(y_value*100)
Y
#np.count_nonzero((plotting_array[:,0] > -1) & (plotting_array[:,0] < 1))/plotting_array[:,0].size
import numpy as np
import matplotlib.pyplot as plt
fig = plt.figure(figsize=(10,10))
plt.plot(plotting_itr,Y,label = 'Mango Serial',linewidth=3.0) #x, y
plt.xlabel('Number of Iterations',fontsize=25)
plt.ylabel(' % of $Fun_{value}$ in (-1,0)',fontsize=25)
plt.title('Variation of Optimal Value of X with iterations',fontsize=20)
plt.xticks(fontsize=20)
plt.yticks(fontsize=20)
plt.yticks(np.arange(60, 110, step=10))
plt.grid(True)
plt.legend(fontsize=20)
plt.show()
```
| github_jupyter |
# Punctuation And Capitalization using Transfer Learning Toolkit
Transfer Learning Toolkit (TLT) is a python based AI toolkit for taking purpose-built pre-trained AI models and customizing them with your own data.
Transfer learning extracts learned features from an existing neural network to a new one. Transfer learning is often used when creating a large training dataset is not feasible.
Developers, researchers and software partners building intelligent vision AI apps and services, can bring their own data to fine-tune pre-trained models instead of going through the hassle of training from scratch.
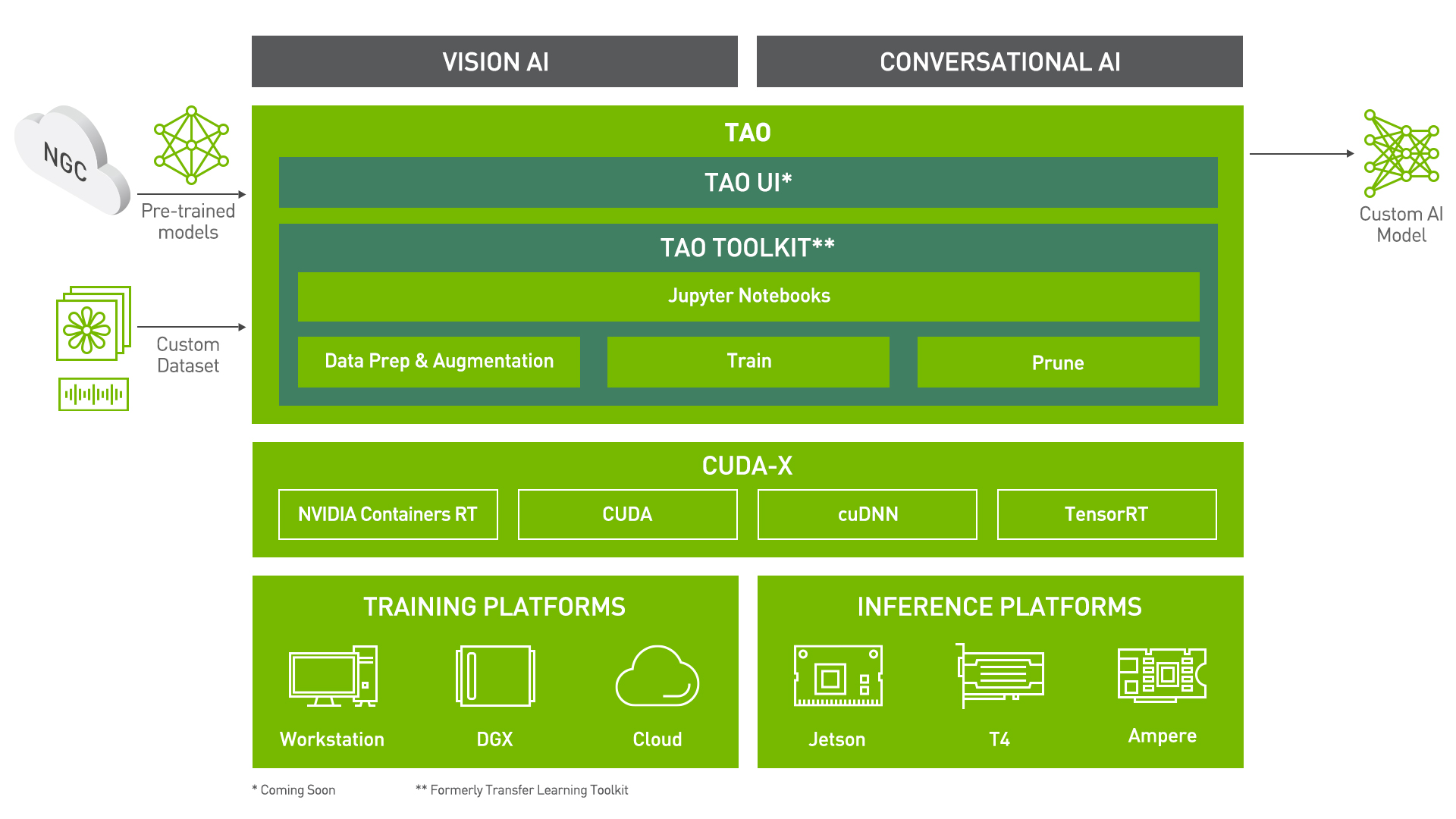
The goal of this toolkit is to reduce that 80 hour workload to an 8 hour workload, which can enable data scientist to have considerably more train-test iterations in the same time frame.
In this notebook, you will learn how to leverage the simplicity and convenience of TLT to:
- Take a BERT model and __Train/Finetune__ it on a dataset for Punctuation and Capitalization task
- Run __Inference__
- __Export__ the model to the ONNX format, or export in the format that is suitable for deployment in Jarvis
The earlier section in this notebook gives a brief introduction to the Punctuation and Capitalization task and the dataset being used.
## Prerequisites
For ease of use, please install TLT inside a python virtual environment. Steps to install the virtual environment are mentioned [here](https://packaging.python.org/guides/installing-using-pip-and-virtual-environments/). We recommend performing this step first and then launching the notebook from the virtual environment.
In addition to installing TLT package, please make sure of the following software requirements:
- python 3.6.9
- docker-ce > 19.03.5
- docker-API 1.40
- nvidia-container-toolkit > 1.3.0-1
- nvidia-container-runtime > 3.4.0-1
- nvidia-docker2 > 2.5.0-1
- nvidia-driver >= 455.23
Let's install TLT. It is a simple pip install!
```
! pip install nvidia-pyindex
! pip install nvidia-tlt
!tlt info --verbose
```
Check if the GPU device(s) are available.
```
!nvidia-smi
```
## Punctuation And Capitalization using TLT
### Task Description
Automatic Speech Recognition (ASR) systems typically generate text with no punctuation and capitalization of the words. This tutorial explains how to implement a model that will predict punctuation and capitalization for each word in a sentence to make ASR output more readable and to boost performance of the named entity recognition, machine translation or text-to-speech models. We'll show how to train a model for this task using a pre-trained BERT model. For every word in our training dataset we’re going to predict:
- punctuation mark that should follow the word
- whether the word should be capitalized
---
### TLT Workflow
### Setting TLT Mounts
Once the TLT gets installed, the next step is to set up the directory mounts that is the ``source`` and ``destination``. The ``source_mount`` will store all the data, pre-trained models and scripts pertaining to Punctuation and Capitalization task. Please select an empty directory for ``source_mount``.
The ``destination_mount`` is the directory to which ``source_mount`` will be mapped to, inside the container.
The TLT launcher uses docker container under the hood, and for our data and results directory to be visible to the docker, they must be mapped.
The launcher can be configured using the config file ``.tlt_mounts.json``. Apart from the mounts you can also configure additional options like the Environmental Variables and amount of Shared Memory available to the TLT launcher.
``Important Note:`` The code below creates a sample ``.tlt_mounts.json`` file. Here, we can map directories in which we save the data, specs, results and cache. You must configure it for your specific case to make sure both your data and results are correctly mapped to the docker. **Please also ensure that the source directories exist on your machine!**
```
%%bash
tee ~/.tlt_mounts.json <<'EOF'
{
"Mounts":[
{
"source": "<YOUR_PATH_TO_DATA_DIR>",
"destination": "/data"
},
{
"source": "<YOUR_PATH_TO_SPECS_DIR>",
"destination": "/specs"
},
{
"source": "<YOUR_PATH_TO_RESULTS_DIR>",
"destination": "/results"
},
{
"source": "<YOUR_PATH_TO_CACHE_DIR>",
"destination": "/root/.cache"
}
]
}
EOF
# Make sure the source directories exist, if not, create them
! mkdir <YOUR_PATH_TO_SPECS_DIR>
! mkdir <YOUR_PATH_TO_RESULTS_DIR>
! mkdir <YOUR_PATH_TO_CACHE_DIR>
```
The rest of the notebook exemplifies the simplicity of the TLT workflow.
Users with any level of Deep Learning knowledge can get started building their own custom models using a simple specification file. It's essentially just one command each to run data download and preprocessing, training, fine-tuning, evaluation, inference, and export.
All configurations happen through YAML specification files
---
### Configuration/Specification Files
All commands in TLT lies in the YAML specification files. There are sample specification files already available for you to use directly or as reference to create your own YAML specification files.
Through these specification files, you can tune many a lot of things like the model, dataset, hyperparameters, optimizer etc.
Each command (like download_and_convert, train, finetune, evaluate etc.) should have a dedicated specification file with configurations pertinent to it.
Here is an example of the training spec file:
```python
save_to: trained-model.tlt
trainer:
max_epochs: 5
model:
punct_label_ids:
O: 0
',': 1
'.': 2
'?': 3
capit_label_ids:
O: 0
U: 1
tokenizer:
tokenizer_name: ${model.language_model.pretrained_model_name} # or sentencepiece =
vocab_file: null # path to vocab file
tokenizer_model: null # only used if tokenizer is sentencepiece
special_tokens: null
language_model:
pretrained_model_name: bert-base-uncased
lm_checkpoint: null
config_file: null # json file, precedence over config
config: null
punct_head:
punct_num_fc_layers: 1
fc_dropout: 0.1
activation: 'relu'
use_transformer_init: true
capit_head:
capit_num_fc_layers: 1
fc_dropout: 0.1
activation: 'relu'
use_transformer_init: true
# Data dir containing dataset.
data_dir: ???
training_ds:
text_file: text_train.txt
labels_file: labels_train.txt
shuffle: true
num_samples: -1 # number of samples to be considered, -1 means the whole the dataset
batch_size: 64
validation_ds:
text_file: text_dev.txt
labels_file: labels_dev.txt
shuffle: false
num_samples: -1 # number of samples to be considered, -1 means the whole the dataset
batch_size: 64
optim:
name: adam
lr: 1e-5
weight_decay: 0.00
sched:
name: WarmupAnnealing
# Scheduler params
warmup_steps: null
warmup_ratio: 0.1
last_epoch: -1
# pytorch lightning args
monitor: val_loss
reduce_on_plateau: false
```
---
### Set Relevant Paths
Please set these paths according to your environment.
```
# NOTE: The following paths are set from the perspective of the TLT Docker.
# The data is saved here
DATA_DIR='/data'
# The configuration files are stored here
SPECS_DIR='/specs/punctuation_and_capitalization'
# The results are saved at this path
RESULTS_DIR='/results/punctuation_and_capitalization'
# Set your encryption key, and use the same key for all commands
KEY='tlt_encode'
```
---
### Downloading Specs
We can proceed to downloading the spec files. The user may choose to modify/rewrite these specs, or even individually override them through the launcher. You can download the default spec files by using the `download_specs` command. <br>
The -o argument indicating the folder where the default specification files will be downloaded, and -r that instructs the script where to save the logs. **Make sure the -o points to an empty folder!**
```
!tlt punctuation_and_capitalization download_specs \
-r $RESULTS_DIR/ \
-o $SPECS_DIR/
```
---
### Dataset
This model can work with any dataset as long as it follows the format specified below. The training and evaluation data is divided into _2 files: text.txt and labels.txt_. Each line of the __text.txt__ file contains text sequences, where words are separated with spaces: [WORD] [SPACE] [WORD] [SPACE] [WORD], for example:
when is the next flight to new york<br>
the next flight is ...<br>
...<br>
The __labels.txt__ file contains corresponding labels for each word in text.txt, the labels are separated with spaces. Each label in labels.txt file consists of 2 symbols:
- the first symbol of the label indicates what punctuation mark should follow the word (where O means no punctuation needed);
- the second symbol determines if a word needs to be capitalized or not (where U indicates that the word should be upper cased, and O - no capitalization needed.)
In this tutorial, we are considering only commas, periods, and question marks the rest punctuation marks were removed. To use more punctuation marks, update the dataset to include desired labels, no changes to the model needed.
Each line of the __labels.txt__ should follow the format: [LABEL] [SPACE] [LABEL] [SPACE] [LABEL] (for labels.txt). For example, labels for the above text.txt file should be:
OU OO OO OO OO OO OU ?U<br>
OU OO OO OO ...<br>
...
The complete list of all possible labels for this task used in this tutorial is: OO, ,O, .O, ?O, OU, ,U, .U, ?U.
### Download and preprocess the data
In this notebook we are going to use a subset of English examples from the [Tatoeba collection of sentences](https://tatoeba.org/eng).
Downloading and preprocessing the data using TLT is as simple as configuring YAML specification file and running the ``download_and_convert_tatoeba`` command. The code cell below uses the default `download_and_convert_tatoeba.yaml` available for the users as a reference.
The configurations in the specification file can be easily overridden using the tlt-launcher CLI as shown below. For instance, we override the ``source_data_dir`` and ``target_data_dir`` configurations.
We encourage you to take a look at the YAML files we have provided.
After executing the cell below, your data folder will contain the following 4 files:
- labels_dev.txt
- labels_train.txt
- text_dev.txt
- text_train.txt
```
### To download and convert the dataset
!tlt punctuation_and_capitalization download_and_convert_tatoeba \
-e $SPECS_DIR/download_and_convert_tatoeba.yaml \
source_data_dir=$DATA_DIR \
target_data_dir=$DATA_DIR
```
---
### Training
In the Punctuation and Capitalization Model, we are jointly training two token-level classifiers on top of the pretrained [BERT](https://arxiv.org/pdf/1810.04805.pdf) model:
- one classifier to predict punctuation and
- the other one - capitalization.
Training a model using TLT is as simple as configuring your spec file and running the train command. The code cell below uses the default train.yaml available for users as reference. It is configured by default to use the ``bert-base-uncased`` pretrained model. Additionally, these configurations could easily be overridden using the tlt-launcher CLI as shown below. For instance, below we override the trainer.max_epochs, training_ds.num_samples and validation_ds.num_samples configurations to suit our needs. We encourage you to take a look at the .yaml spec files we provide!
The command for training is very similar to the of ``download_and_convert_tatoeba``. Instead of ``tlt punctuation_and_capitalization download_and_convert_tatoeba``, we use ``tlt punctuation_and_capitalization train`` instead. The ``tlt punctuation_and_capitalization train`` command has the following arguments:
- ``-e`` : Path to the spec file
- ``-g`` : Number of GPUs to use
- ``-k`` : User specified encryption key to use while saving/loading the model
- ``-r`` : Path to the folder where the outputs should be written. Make sure this is mapped in the tlt_mounts.json
- Any overrides to the spec file eg. trainer.max_epochs
More details about these arguments are present in the [TLT Getting Started Guide](https://docs.nvidia.com/metropolis/TLT/tlt-getting-started-guide/index.html).
``NOTE:`` All file paths corresponds to the destination mounted directory that is visible in the TLT docker container used in backend.
```
### To train the dataset with BERT-base-uncased model
!tlt punctuation_and_capitalization train \
-e $SPECS_DIR/train.yaml \
-g 4 \
-r $RESULTS_DIR/train \
data_dir=$DATA_DIR \
trainer.max_epochs=2 \
training_ds.num_samples=-1 \
validation_ds.num_samples=-1 \
-k $KEY
```
The train command produces a .tlt file called ``trained-model.tlt`` saved at ``$RESULTS_DIR/checkpoints/trained-model.tlt``
### Other tips and tricks:
- To accelerate the training without loss of quality, it is possible to train with these parameters: ``trainer.amp_level="O1"`` and ``trainer.precision=16`` for reduced precision.
- The batch size ``training_ds.batch_size`` may influence the validation accuracy. Larger batch sizes are faster to train with, however, you may get slighly better results with smaller batches.
- You can also change the optimizer parameter ``optim.name`` and can see its effect on the punctuation and capitalization task by the change in accuracy.
- You can specify the number of layers in the head of the model ``model.punct_head.punct_num_fc_layers`` and ``model.capit_head.capit_num_fc_layers``.
---
### Finetuning
As stated above the command for all the tasks are very similar but have different YAML specification files that can be tweaked.
Note: If you wish to proceed with a trained dataset for better inference results, you can find a .nemo model [here](
https://ngc.nvidia.com/catalog/collections/nvidia:nemotrainingframework).
Simply re-name the .nemo file to .tlt and pass it through the finetune pipeline.
```
### To finetune on the dataset
!tlt punctuation_and_capitalization finetune \
-e $SPECS_DIR/finetune.yaml \
-g 4 \
-m $RESULTS_DIR/train/checkpoints/trained-model.tlt \
-r $RESULTS_DIR/finetune \
data_dir=$DATA_DIR \
trainer.max_epochs=3 \
-k $KEY
```
The train command produces a .tlt file called ``finetuned-model.tlt`` saved in the results directory.
---
### Evaluation
To evaluate our TLT model we will run the command below. It is always advisable to look at the YAML file for evaluate to understand the command in a better way.
```
### For evaluation
!tlt punctuation_and_capitalization evaluate \
-e $SPECS_DIR/evaluate.yaml \
-g 4 \
-m $RESULTS_DIR/finetune/checkpoints/finetuned-model.tlt \
data_dir=$DATA_DIR \
-r $RESULTS_DIR/evaluate \
-k $KEY
```
On evaluating the model you will get some results and based on that we can either retrain the model for more epochs or continue with the inference.
---
### Inference
Inference using a TLT trained and fine-tuned model can be done by ``tlt punctuation_and_capitalization infer`` command. It is again advisable to look at the infer.yaml file.
```
### For inference
!tlt punctuation_and_capitalization infer \
-e $SPECS_DIR/infer.yaml \
-g 4 \
-m $RESULTS_DIR/finetune/checkpoints/finetuned-model.tlt \
-r $RESULTS_DIR/infer \
-k $KEY
```
---
### Export to ONNX
[ONNX](https://onnx.ai/) is a popular open format for machine learning models. It enables interoperability between different frameworks, making the path to production much easier.
TLT provides commands to export the .tlt model to the ONNX format in an .eonnx archive. The `export_format` configuration can be set to `ONNX` to achieve this.
The tlt export command for ``punctuation_and_capitalization`` is shown in the cell below.
```
### For export to ONNX
!tlt punctuation_and_capitalization export \
-e $SPECS_DIR/export.yaml \
-g 1 \
-m $RESULTS_DIR/finetune/checkpoints/finetuned-model.tlt \
-r $RESULTS_DIR/export \
-k $KEY \
export_format=ONNX
```
This command exports the model as ``exported-model.eonnx`` which is essentially an archive containing the .onnx model.
---
### Inference using ONNX
TLT provides the capability to use the exported .eonnx model for inference. The command ``tlt punctuation_and_capitalization infer_onnx`` is very similar to the inference command for .tlt models. Again, the input file used is just for demo purposes, you may choose to try out your own custom input.
```
### For inference using ONNX
!tlt punctuation_and_capitalization infer_onnx \
-e $SPECS_DIR/infer_onnx.yaml \
-g 1 \
-m $RESULTS_DIR/export/exported-model.eonnx \
-r $RESULTS_DIR/infer_onnx \
-k $KEY
```
---
### Export to JARVIS
With TLT, you can also export your model in a format that can deployed using [NVIDIA JARVIS](https://developer.nvidia.com/nvidia-jarvis), a highly performant application framework for multi-modal conversational AI services using GPUs. The same command for exporting to ONNX can be used here. The only small variation is the configuration for ``export_format`` in the spec file.
```
### For export to JARVIS
!tlt punctuation_and_capitalization export \
-e $SPECS_DIR/export.yaml \
-g 1 \
-m $RESULTS_DIR/finetune/checkpoints/finetuned-model.tlt \
-r $RESULTS_DIR/export_jarvis \
export_format=JARVIS \
export_to=punct-capit-model.ejrvs \
-k $KEY
```
The model is exported as ``exported-model.ejrvs`` which is in a format suited for deployment in Jarvis.
---
### What next?
You can use TLT to build custom models for your own NLP applications.
| github_jupyter |
# Face Detection using SSD and the Caffe pre-trained model
*by Georgios K. Ouzounis, June 21st, 2021*
This notebook demonstrates face detection in still images using the SSD detector configured with the Caffe pretrained model
## Copy the model files
We need the configuration file and the pre-trained weights
```
%mkdir model/
!wget https://raw.githubusercontent.com/georgiosouzounis/face-detection-ssd-caffe/main/model/deploy.prototxt.txt -O model/deploy.prototxt.txt
!wget https://github.com/georgiosouzounis/face-detection-ssd-caffe/raw/main/model/res10_300x300_ssd_iter_140000.caffemodel -O model/res10_300x300_ssd_iter_140000.caffemodel
```
## Import the libraries
```
# import the relevant libraries
import numpy as np
import cv2 # openCV
# check the opencv version
if cv2.__version__ < '4.5.2':
print("opencv version: ", cv2.__version__)
print("please upgrade your opencv installation to the latest")
# if the openCV version is < 4.4.0 update to the latest otherwise skip this step
!pip install opencv-python==4.5.2.52
```
## Read the model and initialize the detector
```
# load the serialized model from the local copy in model/
model_cfg = "model/deploy.prototxt.txt"
model_weights = "model/res10_300x300_ssd_iter_140000.caffemodel"
# read the model
detector = cv2.dnn.readNetFromCaffe(model_cfg, model_weights)
```
## Get a test image
Set the path to an image containing a face in your own Google Drive or use the example as shown:
```
!wget https://github.com/georgiosouzounis/face-detection-ssd-caffe/raw/main/data/macron.jpg
test_img = "macron.jpg"
# load the test image and create an image blob
image = cv2.imread(test_img)
(h, w) = image.shape[:2]
# display the image
from google.colab.patches import cv2_imshow
cv2_imshow(image)
```
## Deploy the detector
```
# set the intensity scaling factor; 1 in this case, i.e. original image intensities
scalefactor = 1.0
# set the new dimensions for image resizing to match the network requirements
new_size = (300, 300)
# create a blob using OpenCV's DNN functionality and by performing mean subtraction
# to normalize the input
blob = cv2.dnn.blobFromImage(image, scalefactor, new_size, (127.5, 127.5, 127.5), swapRB=True, crop=False)
# set the blob as input to the network
detector.setInput(blob)
# compute the forward pass - detect faces if any
detections = detector.forward()
detections.shape
```
## Analyze the results
Let us review the detections. The shape of the detections is expected to be in the following format: ```[1, 1, N, 7]```, where N is the number of detected bounding boxes. For each detection, the description has the format: ```[image_id, label, conf, x_min, y_min, x_max, y_max]```.
```
detections[0][0][0]
len(detections[0][0])
# set the confidence threshold
confidence_threshold = 0.5
# loop over the detections
for i in range(0, detections.shape[2]):
# extract the confidence (i.e., probability) associated with the prediction
confidence = detections[0, 0, i, 2]
# ignore weak detections
if confidence > confidence_threshold:
# compute the (x, y)-coordinates of the bounding box for the detected object
box = detections[0, 0, i, 3:7] * np.array([w, h, w, h])
(startX, startY, endX, endY) = box.astype("int")
# draw the bounding box of the detected face
cv2.rectangle(image, (startX, startY), (endX, endY), (0, 0, 255), 2)
# print the probability of this detection
text = "confidence: {:.2f}%".format(confidence * 100)
y = startY - 10 if startY - 10 > 10 else startY + 10
cv2.putText(image, text, (startX, y), cv2.FONT_HERSHEY_SIMPLEX, 0.45, (0, 0, 255), 2)
# show the output image
cv2_imshow(image)
```
| github_jupyter |
# Computer Vision Nanodegree
## Project: Image Captioning
---
In this notebook, you will learn how to load and pre-process data from the [COCO dataset](http://cocodataset.org/#home). You will also design a CNN-RNN model for automatically generating image captions.
Note that **any amendments that you make to this notebook will not be graded**. However, you will use the instructions provided in **Step 3** and **Step 4** to implement your own CNN encoder and RNN decoder by making amendments to the **models.py** file provided as part of this project. Your **models.py** file **will be graded**.
Feel free to use the links below to navigate the notebook:
- [Step 1](#step1): Explore the Data Loader
- [Step 2](#step2): Use the Data Loader to Obtain Batches
- [Step 3](#step3): Experiment with the CNN Encoder
- [Step 4](#step4): Implement the RNN Decoder
<a id='step1'></a>
## Step 1: Explore the Data Loader
We have already written a [data loader](http://pytorch.org/docs/master/data.html#torch.utils.data.DataLoader) that you can use to load the COCO dataset in batches.
In the code cell below, you will initialize the data loader by using the `get_loader` function in **data_loader.py**.
> For this project, you are not permitted to change the **data_loader.py** file, which must be used as-is.
The `get_loader` function takes as input a number of arguments that can be explored in **data_loader.py**. Take the time to explore these arguments now by opening **data_loader.py** in a new window. Most of the arguments must be left at their default values, and you are only allowed to amend the values of the arguments below:
1. **`transform`** - an [image transform](http://pytorch.org/docs/master/torchvision/transforms.html) specifying how to pre-process the images and convert them to PyTorch tensors before using them as input to the CNN encoder. For now, you are encouraged to keep the transform as provided in `transform_train`. You will have the opportunity later to choose your own image transform to pre-process the COCO images.
2. **`mode`** - one of `'train'` (loads the training data in batches) or `'test'` (for the test data). We will say that the data loader is in training or test mode, respectively. While following the instructions in this notebook, please keep the data loader in training mode by setting `mode='train'`.
3. **`batch_size`** - determines the batch size. When training the model, this is number of image-caption pairs used to amend the model weights in each training step.
4. **`vocab_threshold`** - the total number of times that a word must appear in the in the training captions before it is used as part of the vocabulary. Words that have fewer than `vocab_threshold` occurrences in the training captions are considered unknown words.
5. **`vocab_from_file`** - a Boolean that decides whether to load the vocabulary from file.
We will describe the `vocab_threshold` and `vocab_from_file` arguments in more detail soon. For now, run the code cell below. Be patient - it may take a couple of minutes to run!
```
import sys
sys.path.append('/opt/cocoapi/PythonAPI')
from pycocotools.coco import COCO
!pip install nltk
import nltk
nltk.download('punkt')
from data_loader import get_loader
from torchvision import transforms
# Define a transform to pre-process the training images.
transform_train = transforms.Compose([
transforms.Resize(256), # smaller edge of image resized to 256
transforms.RandomCrop(224), # get 224x224 crop from random location
transforms.RandomHorizontalFlip(), # horizontally flip image with probability=0.5
transforms.ToTensor(), # convert the PIL Image to a tensor
transforms.Normalize((0.485, 0.456, 0.406), # normalize image for pre-trained model
(0.229, 0.224, 0.225))])
# Set the minimum word count threshold.
vocab_threshold = 5
# Specify the batch size.
batch_size = 10
# Obtain the data loader.
# data_loader = get_loader(transform=transform_train,
# mode='train',
# batch_size=batch_size,
# vocab_threshold=vocab_threshold,
# vocab_from_file=False)
```
When you ran the code cell above, the data loader was stored in the variable `data_loader`.
You can access the corresponding dataset as `data_loader.dataset`. This dataset is an instance of the `CoCoDataset` class in **data_loader.py**. If you are unfamiliar with data loaders and datasets, you are encouraged to review [this PyTorch tutorial](http://pytorch.org/tutorials/beginner/data_loading_tutorial.html).
### Exploring the `__getitem__` Method
The `__getitem__` method in the `CoCoDataset` class determines how an image-caption pair is pre-processed before being incorporated into a batch. This is true for all `Dataset` classes in PyTorch; if this is unfamiliar to you, please review [the tutorial linked above](http://pytorch.org/tutorials/beginner/data_loading_tutorial.html).
When the data loader is in training mode, this method begins by first obtaining the filename (`path`) of a training image and its corresponding caption (`caption`).
#### Image Pre-Processing
Image pre-processing is relatively straightforward (from the `__getitem__` method in the `CoCoDataset` class):
```python
# Convert image to tensor and pre-process using transform
image = Image.open(os.path.join(self.img_folder, path)).convert('RGB')
image = self.transform(image)
```
After loading the image in the training folder with name `path`, the image is pre-processed using the same transform (`transform_train`) that was supplied when instantiating the data loader.
#### Caption Pre-Processing
The captions also need to be pre-processed and prepped for training. In this example, for generating captions, we are aiming to create a model that predicts the next token of a sentence from previous tokens, so we turn the caption associated with any image into a list of tokenized words, before casting it to a PyTorch tensor that we can use to train the network.
To understand in more detail how COCO captions are pre-processed, we'll first need to take a look at the `vocab` instance variable of the `CoCoDataset` class. The code snippet below is pulled from the `__init__` method of the `CoCoDataset` class:
```python
def __init__(self, transform, mode, batch_size, vocab_threshold, vocab_file, start_word,
end_word, unk_word, annotations_file, vocab_from_file, img_folder):
...
self.vocab = Vocabulary(vocab_threshold, vocab_file, start_word,
end_word, unk_word, annotations_file, vocab_from_file)
...
```
From the code snippet above, you can see that `data_loader.dataset.vocab` is an instance of the `Vocabulary` class from **vocabulary.py**. Take the time now to verify this for yourself by looking at the full code in **data_loader.py**.
We use this instance to pre-process the COCO captions (from the `__getitem__` method in the `CoCoDataset` class):
```python
# Convert caption to tensor of word ids.
tokens = nltk.tokenize.word_tokenize(str(caption).lower()) # line 1
caption = [] # line 2
caption.append(self.vocab(self.vocab.start_word)) # line 3
caption.extend([self.vocab(token) for token in tokens]) # line 4
caption.append(self.vocab(self.vocab.end_word)) # line 5
caption = torch.Tensor(caption).long() # line 6
```
As you will see soon, this code converts any string-valued caption to a list of integers, before casting it to a PyTorch tensor. To see how this code works, we'll apply it to the sample caption in the next code cell.
```
sample_caption = 'A person doing a trick on a rail while riding a skateboard.'
```
In **`line 1`** of the code snippet, every letter in the caption is converted to lowercase, and the [`nltk.tokenize.word_tokenize`](http://www.nltk.org/) function is used to obtain a list of string-valued tokens. Run the next code cell to visualize the effect on `sample_caption`.
```
import nltk
sample_tokens = nltk.tokenize.word_tokenize(str(sample_caption).lower())
print(sample_tokens)
```
In **`line 2`** and **`line 3`** we initialize an empty list and append an integer to mark the start of a caption. The [paper](https://arxiv.org/pdf/1411.4555.pdf) that you are encouraged to implement uses a special start word (and a special end word, which we'll examine below) to mark the beginning (and end) of a caption.
This special start word (`"<start>"`) is decided when instantiating the data loader and is passed as a parameter (`start_word`). You are **required** to keep this parameter at its default value (`start_word="<start>"`).
As you will see below, the integer `0` is always used to mark the start of a caption.
```
sample_caption = []
start_word = data_loader.dataset.vocab.start_word
print('Special start word:', start_word)
sample_caption.append(data_loader.dataset.vocab(start_word))
print(sample_caption)
```
In **`line 4`**, we continue the list by adding integers that correspond to each of the tokens in the caption.
```
sample_caption.extend([data_loader.dataset.vocab(token) for token in sample_tokens])
print(sample_caption)
```
In **`line 5`**, we append a final integer to mark the end of the caption.
Identical to the case of the special start word (above), the special end word (`"<end>"`) is decided when instantiating the data loader and is passed as a parameter (`end_word`). You are **required** to keep this parameter at its default value (`end_word="<end>"`).
As you will see below, the integer `1` is always used to mark the end of a caption.
```
end_word = data_loader.dataset.vocab.end_word
print('Special end word:', end_word)
sample_caption.append(data_loader.dataset.vocab(end_word))
print(sample_caption)
```
Finally, in **`line 6`**, we convert the list of integers to a PyTorch tensor and cast it to [long type](http://pytorch.org/docs/master/tensors.html#torch.Tensor.long). You can read more about the different types of PyTorch tensors on the [website](http://pytorch.org/docs/master/tensors.html).
```
import torch
sample_caption = torch.Tensor(sample_caption).long()
print(sample_caption)
```
And that's it! In summary, any caption is converted to a list of tokens, with _special_ start and end tokens marking the beginning and end of the sentence:
```
[<start>, 'a', 'person', 'doing', 'a', 'trick', 'while', 'riding', 'a', 'skateboard', '.', <end>]
```
This list of tokens is then turned into a list of integers, where every distinct word in the vocabulary has an associated integer value:
```
[0, 3, 98, 754, 3, 396, 207, 139, 3, 753, 18, 1]
```
Finally, this list is converted to a PyTorch tensor. All of the captions in the COCO dataset are pre-processed using this same procedure from **`lines 1-6`** described above.
As you saw, in order to convert a token to its corresponding integer, we call `data_loader.dataset.vocab` as a function. The details of how this call works can be explored in the `__call__` method in the `Vocabulary` class in **vocabulary.py**.
```python
def __call__(self, word):
if not word in self.word2idx:
return self.word2idx[self.unk_word]
return self.word2idx[word]
```
The `word2idx` instance variable is a Python [dictionary](https://docs.python.org/3/tutorial/datastructures.html#dictionaries) that is indexed by string-valued keys (mostly tokens obtained from training captions). For each key, the corresponding value is the integer that the token is mapped to in the pre-processing step.
Use the code cell below to view a subset of this dictionary.
```
# Preview the word2idx dictionary.
dict(list(data_loader.dataset.vocab.word2idx.items())[:10])
```
We also print the total number of keys.
```
# Print the total number of keys in the word2idx dictionary.
print('Total number of tokens in vocabulary:', len(data_loader.dataset.vocab))
```
As you will see if you examine the code in **vocabulary.py**, the `word2idx` dictionary is created by looping over the captions in the training dataset. If a token appears no less than `vocab_threshold` times in the training set, then it is added as a key to the dictionary and assigned a corresponding unique integer. You will have the option later to amend the `vocab_threshold` argument when instantiating your data loader. Note that in general, **smaller** values for `vocab_threshold` yield a **larger** number of tokens in the vocabulary. You are encouraged to check this for yourself in the next code cell by decreasing the value of `vocab_threshold` before creating a new data loader.
```
# Modify the minimum word count threshold.
vocab_threshold = 5
# Obtain the data loader.
data_loader = get_loader(transform=transform_train,
mode='train',
batch_size=batch_size,
vocab_threshold=vocab_threshold,
vocab_from_file=False)
# Print the total number of keys in the word2idx dictionary.
print('Total number of tokens in vocabulary:', len(data_loader.dataset.vocab))
```
There are also a few special keys in the `word2idx` dictionary. You are already familiar with the special start word (`"<start>"`) and special end word (`"<end>"`). There is one more special token, corresponding to unknown words (`"<unk>"`). All tokens that don't appear anywhere in the `word2idx` dictionary are considered unknown words. In the pre-processing step, any unknown tokens are mapped to the integer `2`.
```
unk_word = data_loader.dataset.vocab.unk_word
print('Special unknown word:', unk_word)
print('All unknown words are mapped to this integer:', data_loader.dataset.vocab(unk_word))
```
Check this for yourself below, by pre-processing the provided nonsense words that never appear in the training captions.
```
print(data_loader.dataset.vocab('jfkafejw'))
print(data_loader.dataset.vocab('ieowoqjf'))
```
The final thing to mention is the `vocab_from_file` argument that is supplied when creating a data loader. To understand this argument, note that when you create a new data loader, the vocabulary (`data_loader.dataset.vocab`) is saved as a [pickle](https://docs.python.org/3/library/pickle.html) file in the project folder, with filename `vocab.pkl`.
If you are still tweaking the value of the `vocab_threshold` argument, you **must** set `vocab_from_file=False` to have your changes take effect.
But once you are happy with the value that you have chosen for the `vocab_threshold` argument, you need only run the data loader *one more time* with your chosen `vocab_threshold` to save the new vocabulary to file. Then, you can henceforth set `vocab_from_file=True` to load the vocabulary from file and speed the instantiation of the data loader. Note that building the vocabulary from scratch is the most time-consuming part of instantiating the data loader, and so you are strongly encouraged to set `vocab_from_file=True` as soon as you are able.
Note that if `vocab_from_file=True`, then any supplied argument for `vocab_threshold` when instantiating the data loader is completely ignored.
```
# Obtain the data loader (from file). Note that it runs much faster than before!
data_loader = get_loader(transform=transform_train,
mode='train',
batch_size=batch_size,
vocab_from_file=True)
```
In the next section, you will learn how to use the data loader to obtain batches of training data.
<a id='step2'></a>
## Step 2: Use the Data Loader to Obtain Batches
The captions in the dataset vary greatly in length. You can see this by examining `data_loader.dataset.caption_lengths`, a Python list with one entry for each training caption (where the value stores the length of the corresponding caption).
In the code cell below, we use this list to print the total number of captions in the training data with each length. As you will see below, the majority of captions have length 10. Likewise, very short and very long captions are quite rare.
```
from collections import Counter
# Tally the total number of training captions with each length.
counter = Counter(data_loader.dataset.caption_lengths)
lengths = sorted(counter.items(), key=lambda pair: pair[1], reverse=True)
for value, count in lengths:
print('value: %2d --- count: %5d' % (value, count))
```
To generate batches of training data, we begin by first sampling a caption length (where the probability that any length is drawn is proportional to the number of captions with that length in the dataset). Then, we retrieve a batch of size `batch_size` of image-caption pairs, where all captions have the sampled length. This approach for assembling batches matches the procedure in [this paper](https://arxiv.org/pdf/1502.03044.pdf) and has been shown to be computationally efficient without degrading performance.
Run the code cell below to generate a batch. The `get_train_indices` method in the `CoCoDataset` class first samples a caption length, and then samples `batch_size` indices corresponding to training data points with captions of that length. These indices are stored below in `indices`.
These indices are supplied to the data loader, which then is used to retrieve the corresponding data points. The pre-processed images and captions in the batch are stored in `images` and `captions`.
```
import numpy as np
import torch.utils.data as data
# Randomly sample a caption length, and sample indices with that length.
indices = data_loader.dataset.get_train_indices()
print('sampled indices:', indices)
# Create and assign a batch sampler to retrieve a batch with the sampled indices.
new_sampler = data.sampler.SubsetRandomSampler(indices=indices)
data_loader.batch_sampler.sampler = new_sampler
# Obtain the batch.
images, captions = next(iter(data_loader))
print('images.shape:', images.shape)
print('captions.shape:', captions.shape)
# (Optional) Uncomment the lines of code below to print the pre-processed images and captions.
print('images:', images)
print('captions:', captions)
```
Each time you run the code cell above, a different caption length is sampled, and a different batch of training data is returned. Run the code cell multiple times to check this out!
You will train your model in the next notebook in this sequence (**2_Training.ipynb**). This code for generating training batches will be provided to you.
> Before moving to the next notebook in the sequence (**2_Training.ipynb**), you are strongly encouraged to take the time to become very familiar with the code in **data_loader.py** and **vocabulary.py**. **Step 1** and **Step 2** of this notebook are designed to help facilitate a basic introduction and guide your understanding. However, our description is not exhaustive, and it is up to you (as part of the project) to learn how to best utilize these files to complete the project. __You should NOT amend any of the code in either *data_loader.py* or *vocabulary.py*.__
In the next steps, we focus on learning how to specify a CNN-RNN architecture in PyTorch, towards the goal of image captioning.
<a id='step3'></a>
## Step 3: Experiment with the CNN Encoder
Run the code cell below to import `EncoderCNN` and `DecoderRNN` from **model.py**.
```
# Watch for any changes in model.py, and re-load it automatically.
% load_ext autoreload
% autoreload 2
# Import EncoderCNN and DecoderRNN.
from model import EncoderCNN, DecoderRNN
```
In the next code cell we define a `device` that you will use move PyTorch tensors to GPU (if CUDA is available). Run this code cell before continuing.
```
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
```
Run the code cell below to instantiate the CNN encoder in `encoder`.
The pre-processed images from the batch in **Step 2** of this notebook are then passed through the encoder, and the output is stored in `features`.
```
# Specify the dimensionality of the image embedding.
embed_size = 256
#-#-#-# Do NOT modify the code below this line. #-#-#-#
# Initialize the encoder. (Optional: Add additional arguments if necessary.)
encoder = EncoderCNN(embed_size)
# Move the encoder to GPU if CUDA is available.
encoder.to(device)
# Move last batch of images (from Step 2) to GPU if CUDA is available.
images = images.to(device)
# Pass the images through the encoder.
features = encoder(images)
print('type(features):', type(features))
print('features.shape:', features.shape)
# Check that your encoder satisfies some requirements of the project! :D
assert type(features)==torch.Tensor, "Encoder output needs to be a PyTorch Tensor."
assert (features.shape[0]==batch_size) & (features.shape[1]==embed_size), "The shape of the encoder output is incorrect."
```
The encoder that we provide to you uses the pre-trained ResNet-50 architecture (with the final fully-connected layer removed) to extract features from a batch of pre-processed images. The output is then flattened to a vector, before being passed through a `Linear` layer to transform the feature vector to have the same size as the word embedding.

You are welcome (and encouraged) to amend the encoder in **model.py**, to experiment with other architectures. In particular, consider using a [different pre-trained model architecture](http://pytorch.org/docs/master/torchvision/models.html). You may also like to [add batch normalization](http://pytorch.org/docs/master/nn.html#normalization-layers).
> You are **not** required to change anything about the encoder.
For this project, you **must** incorporate a pre-trained CNN into your encoder. Your `EncoderCNN` class must take `embed_size` as an input argument, which will also correspond to the dimensionality of the input to the RNN decoder that you will implement in Step 4. When you train your model in the next notebook in this sequence (**2_Training.ipynb**), you are welcome to tweak the value of `embed_size`.
If you decide to modify the `EncoderCNN` class, save **model.py** and re-execute the code cell above. If the code cell returns an assertion error, then please follow the instructions to modify your code before proceeding. The assert statements ensure that `features` is a PyTorch tensor with shape `[batch_size, embed_size]`.
<a id='step4'></a>
## Step 4: Implement the RNN Decoder
Before executing the next code cell, you must write `__init__` and `forward` methods in the `DecoderRNN` class in **model.py**. (Do **not** write the `sample` method yet - you will work with this method when you reach **3_Inference.ipynb**.)
> The `__init__` and `forward` methods in the `DecoderRNN` class are the only things that you **need** to modify as part of this notebook. You will write more implementations in the notebooks that appear later in the sequence.
Your decoder will be an instance of the `DecoderRNN` class and must accept as input:
- the PyTorch tensor `features` containing the embedded image features (outputted in Step 3, when the last batch of images from Step 2 was passed through `encoder`), along with
- a PyTorch tensor corresponding to the last batch of captions (`captions`) from Step 2.
Note that the way we have written the data loader should simplify your code a bit. In particular, every training batch will contain pre-processed captions where all have the same length (`captions.shape[1]`), so **you do not need to worry about padding**.
> While you are encouraged to implement the decoder described in [this paper](https://arxiv.org/pdf/1411.4555.pdf), you are welcome to implement any architecture of your choosing, as long as it uses at least one RNN layer, with hidden dimension `hidden_size`.
Although you will test the decoder using the last batch that is currently stored in the notebook, your decoder should be written to accept an arbitrary batch (of embedded image features and pre-processed captions [where all captions have the same length]) as input.

In the code cell below, `outputs` should be a PyTorch tensor with size `[batch_size, captions.shape[1], vocab_size]`. Your output should be designed such that `outputs[i,j,k]` contains the model's predicted score, indicating how likely the `j`-th token in the `i`-th caption in the batch is the `k`-th token in the vocabulary. In the next notebook of the sequence (**2_Training.ipynb**), we provide code to supply these scores to the [`torch.nn.CrossEntropyLoss`](http://pytorch.org/docs/master/nn.html#torch.nn.CrossEntropyLoss) optimizer in PyTorch.
```
# Specify the number of features in the hidden state of the RNN decoder.
hidden_size = 512
#-#-#-# Do NOT modify the code below this line. #-#-#-#
# Store the size of the vocabulary.
vocab_size = len(data_loader.dataset.vocab)
# Initialize the decoder.
decoder = DecoderRNN(embed_size, hidden_size, vocab_size)
# Move the decoder to GPU if CUDA is available.
decoder.to(device)
# Move last batch of captions (from Step 1) to GPU if CUDA is available
captions = captions.to(device)
# Pass the encoder output and captions through the decoder.
outputs = decoder(features, captions)
print('type(outputs):', type(outputs))
print('outputs.shape:', outputs.shape)
# Check that your decoder satisfies some requirements of the project! :D
assert type(outputs)==torch.Tensor, "Decoder output needs to be a PyTorch Tensor."
assert (outputs.shape[0]==batch_size) & (outputs.shape[1]==captions.shape[1]) & (outputs.shape[2]==vocab_size), "The shape of the decoder output is incorrect."
```
When you train your model in the next notebook in this sequence (**2_Training.ipynb**), you are welcome to tweak the value of `hidden_size`.
| github_jupyter |
```
import pandas as pd
from datetime import date, timedelta, datetime
from bs4 import BeautifulSoup
import requests
import re
df_vix_base = pd.read_csv('http://www.cboe.com/publish/scheduledtask/mktdata/datahouse/vixpc.csv',
names=['Date', 'VIX P/C Ratio', 'Puts', 'Calls', 'VIX Options Volume'],
index_col='Date',
parse_dates=['Date'],
skiprows=3)
df_vix_base = df_vix_base[['VIX P/C Ratio', 'VIX Options Volume']]
df_spx_base = pd.read_csv('http://www.cboe.com/publish/scheduledtask/mktdata/datahouse/spxpc.csv',
names=['Date', 'SPX P/C Ratio', 'Puts', 'Calls', 'SPX Options Volume'],
index_col='Date',
parse_dates=['Date'],
skiprows=3)
df_spx_base = df_spx_base[['SPX P/C Ratio', 'SPX Options Volume']]
df_vix_base
df_spx_base
df_cboe_base = df_vix_base.join(df_spx_base, how='inner')
df_cboe_base
cboe_data_dict = {}
n = len(df_cboe_base)
last_known_date = df_cboe_base.index[n-1]
curr_date = last_known_date
end = datetime.today()
delta = timedelta(days=1)
while curr_date <= end:
y, m, d = curr_date.year, curr_date.month, curr_date.day
print(f'Scraping date: {y}-{m:02}-{d:02}')
url = f'https://markets.cboe.com/us/options/market_statistics/daily/?mkt=cone&dt={y}-{m}-{d}'
html = requests.get(url).content
soup = BeautifulSoup(html, 'html.parser')
date_id = 'stats-date-header'
date_string = soup.find(id=date_id).get_text()
date_string = ' '.join(date_string.split(' ')[-3:])
page_date = datetime.strptime(date_string, "%B %d, %Y")
table_class = 'bats-table bats-table--left'
tables = soup.findAll('table', {'class' : table_class})
# to find the table indices
# Summary: 0, VIX: 5, SPX + SPXW: 6
summary_index, vix_index, spx_index = -1, -1, -1
for i,t in enumerate(tables):
s = str(t)
if 'RATIOS' in s:
summary_index = i
elif 'CBOE VOLATILITY INDEX (VIX)' in s:
vix_index = i
elif 'SPX + SPXW' in s:
spx_index = i
if min([summary_index, vix_index, spx_index]) >= 0:
break
table_summary = tables[summary_index]
table_vix = tables[vix_index]
table_spx = tables[spx_index]
df_summary = pd.read_html(str(table_summary))[0]
df_vix = pd.read_html(str(table_vix), skiprows=1)[0]
df_spx = pd.read_html(str(table_spx), skiprows=1)[0]
vix_pc_ratio = df_summary.iloc[4,1]
spx_pc_ratio = df_summary.iloc[5,1]
vix_volume = df_vix.iloc[0,3]
spx_volume = df_spx.iloc[0,3]
cboe_data_dict[page_date] = [
vix_pc_ratio,
vix_volume,
spx_pc_ratio,
spx_volume
]
curr_date += delta
cboe_data_dict
df_cboe_new = pd.DataFrame.from_dict(
cboe_data_dict,
orient='index',
columns=['VIX P/C Ratio', 'VIX Options Volume',
'SPX P/C Ratio', 'SPX Options Volume'])
df_cboe_new.index.rename('Date', inplace=True)
df_cboe_new.tail(20)
df_cboe_full = pd.concat([df_cboe_base, df_cboe_new]).reset_index()
df_cboe_full.tail(20)
df_cboe_full = df_cboe_full.drop_duplicates(subset='Date').set_index('Date').sort_index()
df_cboe_full
df_cboe_full.to_csv('CBOE-data.csv')
```
| github_jupyter |
# This notebook processes CAFE f1 ocean daily data for building climatologies. Only the first 2 years of the forecasts from the period 2003-2015 are used.
Temporary files are written to `tmp_fldr`
```
# Import packages -----
import pandas as pd
import xarray as xr
import numpy as np
from pylatte import utils
from ipywidgets import FloatProgress
```
#### Initialise
```
# Location of forecast data -----
fcst_folder = '/OSM/CBR/OA_DCFP/data/model_output/CAFE/forecasts/v1/'
fcst_filename = 'ocean_daily*'
fields = pd.DataFrame( \
{'name_CAFE': ['sst', 'patm_t', 'eta_t', 'sss', 'u_surf', 'v_surf'],
'name_std' : ['sst', 'patm', 'eta', 'sss', 'u_s', 'v_s']}
)
name_dict = fields.set_index('name_CAFE').to_dict()['name_std']
fields
```
#### Save each init month and variable separately due to memory considerations (this is only necessary for multi-level variables, but all are dealt with in the same way for simplicity)
```
# Temporary folder location -----
tmp_fldr = '/OSM/CBR/OA_DCFP/data/intermediate_products/pylatte_climatologies/tmp/'
# Loop over forecasts, saving numerator and denominator of mean -----
years = range(2003,2016)
months = range(1,13)
ensembles = range(1,12)
for idx, variable in enumerate(fields['name_CAFE']):
print(variable)
print('----------')
for year in years:
print(year)
for month in months:
print(month)
# First see if file already exists -----
savename = 'cafe.fcst.v1.ocean.' + fields['name_std'][idx] + '.' + str(year) + '.' + str(month) + '.clim.nc'
try:
xr.open_dataset(tmp_fldr + savename, autoclose=True)
except:
ens_list = []
ens = []
empty = True
for ie, ensemble in enumerate(ensembles):
path = fcst_folder + '/yr' + str(year) + '/mn' + str(month) + \
'/OUTPUT.' + str(ensemble) + '/' + fcst_filename + '.nc'
# Try to stack ensembles into a list -----
try:
dataset = xr.open_mfdataset(path, autoclose=True)[variable]
# Truncate to 2 year forecasts -----
n_trunc = min([731, len(dataset.time)])
dataset = dataset.isel(time=range(n_trunc))
if 'xu_ocean' in dataset.dims:
dataset = dataset.rename({'xu_ocean':'lon_u','yu_ocean':'lat_u'})
if 'xt_ocean' in dataset.dims:
dataset = dataset.rename({'xt_ocean':'lon_t','yt_ocean':'lat_t'})
ens_list.append(dataset.rename(fields['name_std'][idx]))
ens.append(ie+1)
empty = False
except OSError:
# File does not exist -----
pass
# Concatenate ensembles -----
if empty == False:
ds = xr.concat(ens_list, dim='ensemble')
ds['ensemble'] = ens
# Make month_day array of month-day -----
m = np.array([str(i).zfill(2) + '-' for i in ds.time.dt.month.values])
d = np.array([str(i).zfill(2) for i in ds.time.dt.day.values])
md = np.core.defchararray.add(m, d)
# Replace time array with month_day array and groupby -----
ds['time'] = md
ds_clim = ds.groupby('time').sum(dim='time',keep_attrs=True).to_dataset(name='sum').mean(dim='ensemble')
get_len = lambda ds, dim : ds.count(dim=dim)
ds_clim['count'] = ds['time'].groupby('time').apply(get_len, dim='time')
ds_clim.to_netcdf(path = tmp_fldr + savename, mode = 'w')
del ds, ds_clim
```
#### Combine and write into single climatology file
```
# Use year 2016 as time -----
path = fcst_folder + '/yr2016/mn1/OUTPUT.1/' + fcst_filename + '.nc'
dataset = xr.open_mfdataset(path, autoclose=True)
time_use = dataset.time[:366]
# Loop over all variables -----
for idx, variable in enumerate(fields['name_std']):
print(variable)
print('----------')
name = 'cafe.fcst.v1.ocean.' + fields['name_std'][idx] + '*' + '.clim.nc'
ds = xr.open_mfdataset(tmp_fldr + name, autoclose=True, concat_dim='run')
numer = ds['sum'].sum(dim='run')
denom = ds['count'].sum(dim='run')
if idx == 0:
clim = (numer / denom).to_dataset(name=variable)
else:
clim[variable] = (numer / denom).rename(variable)
clim['time'] = time_use
# Save the climatology -----
save_fldr = '/OSM/CBR/OA_DCFP/data/intermediate_products/pylatte_climatologies/'
clim.to_netcdf(save_fldr + 'cafe.f1.ocean.2003010112_2017123112.clim.nc', mode = 'w',
encoding = {'time':{'dtype':'float','calendar':'JULIAN',
'units':'days since 0001-01-01 00:00:00'}})
```
| github_jupyter |
# Under- and overfitting, model selection
## Preliminaries
In the first set of exercises you had to implement the training and evaluation of the linear regression and $k$-NN methods from scratch in order to practice your `numpy` skills. From this set of exercises onward, you can use the implementations provided in `scikit-learn` or other higher-level libraries. We start this set of exercises by demonstrating some of the features of `scikit-learn`.
For example, implementation of linear regression model fitting with an analytical solution for the parameters is provided by the class `sklearn.linar_model.LinearRegression`. You can train a linear regression model in the following way:
```
import numpy as np
from sklearn import datasets, linear_model
# load the diabetes dataset
diabetes = datasets.load_diabetes()
# use only one feature
X = diabetes.data[:, np.newaxis, 2]
y = diabetes.target
# split the data into training/testing sets
X_train = X[:-20]
X_test = X[-20:]
# split the targets into training/testing sets
y_train = y[:-20]
y_test = y[-20:]
# create linear regression object
model = linear_model.LinearRegression()
# train the model using the training dataset
model.fit(X_train, y_train)
```
Let's visualize the training dataset and the learned regression model.
```
%matplotlib inline
import matplotlib.pyplot as plt
fig = plt.figure()
plt.plot(X_train, y_train, 'r.', markersize=12)
X_edge = np.array([np.min(X_train, 0), np.max(X_train, 0)])
plt.plot(X_edge, model.predict(X_edge), 'b-')
plt.legend(('Data', 'Linear regression'), loc='lower right')
plt.title('Linear regression')
plt.xlabel('$x$')
plt.ylabel('$y$')
plt.show()
```
Once trained, the model can be used to make predictions on the test data:
```
# Make predictions using the testing dataset
prediction = model.predict(X_test)
```
The next step (not shown here) is to evaluate the performance of the trained model.
Note that the `scikit-learn` interface works by first initializing an object from the class that implements the machine learning model (linear regression in this case) and then fitting the initialized model using the data in the training set. Finally, the trained (fitted) model can be used to make predictions on unseen data. In fact, all models implemented in this library follow the same *initialize-fit-predict* programming interface. For example, a $k$-NN classifier can be trained in the following way:
```
from sklearn.model_selection import train_test_split
from sklearn import datasets, neighbors
breast_cancer = datasets.load_breast_cancer()
X = breast_cancer.data
y = breast_cancer.target
# make use of the train_test_split() utility function instead
# of manually dividing the data
X_train, X_test, y_train, y_test = train_test_split(X, y, stratify=y, random_state=40)
# initialize a 3-NN classifier
model = neighbors.KNeighborsClassifier(n_neighbors=3)
# train the model using the training dataset
model.fit(X_train, y_train)
# make predictions using the testing dataset
prediction = model.predict(X_test)
```
Note that the features in the breast cancer dataset have different scales (some have on average very small absolute values, and some very large), which means that the distance metric used by $k$-NN will me dominated by the features with large values. You can use any of the number of feature transformation methods implemented in `scikit-learn` to scale the features. For example, you can use the `sklearn.preprocessing.StandardScaler` method to transform all features to a have a zero mean and unit variance:
```
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaler.fit(X_train)
X_train_scaled = scaler.transform(X_train)
```
The scaler has its own parameters which are the means and standard deviations of the features estimated from the training set. If you train a model with the scaled features, you will have to remember to also apply the scaling transformation every time you make a prediction on new unseen and unscaled data. This is somewhat prone to error. One option for making the code more robust is to create a processing pipeline that includes the scaling and $k$-NN models in a sequence:
```
from sklearn.pipeline import Pipeline
knn = neighbors.KNeighborsClassifier(n_neighbors=3)
model = Pipeline([
("scaler", scaler),
("knn", knn)
])
# train the model using the training dataset
model.fit(X_train, y_train)
# make predictions using the testing dataset
prediction = model.predict(X_test)
```
If you are curious, more information about the design of the `scikit-learn` application programming interface (API) can be found [in this paper](https://arxiv.org/pdf/1309.0238.pdf).
## Exercises
### Bias-variance decomposition
Show that the mean squared error of the estimate of a parameter can be decomposed into an expression that includes both the bias and variance (Eq. 5.53-5.54 in "Deep learning" by Goodfellow et al.).
### Answer Bente
The mean squared error of the estimate of a parameter can be decomposed into an expression that includes both the bias and variance (Eq. 5.53-5.54 in "Deep learning" by Goodfellow et al.):
\begin{aligned}
\mathbb{E}\left[\left(\hat{\theta}{m}-\theta\right)^{2}\right] &= \operatorname{Bias}\left(\hat{\theta}{m}\right)^{2} + \operatorname{Var}\left(\hat{\theta}_{m}\right) \\
\end{aligned}
When we write out the product of the MSE, we obtain (1):
\begin{aligned}
\mathbb{E}\left[\left(\hat{\theta}{m}-\theta\right)^{2}\right] &=\mathbb{E}\left[\hat{\theta}{m}^{2}\right]+\theta^{2}-2 \mathbb{E}\left[\hat{\theta}_{S}\right] \theta \\
\end{aligned}
Now if we express both bias and variance and write out the product of the bias, we obtain (2):
\begin{aligned}
\operatorname{Bias}\left(\hat{\theta}{m}\right)^{2} &=\left(\mathbb{E}\left[\hat{\theta}{m}\right]-\theta\right)^{2} \\
&=\mathbb{E}^{2}\left[\hat{\theta}{m}\right]+\theta^{2}-2 \mathbb{E}\left[\hat{\theta}{m}\right] \theta \\
\operatorname{Var}\left(\hat{\theta}{m}\right) &=\mathbb{E}\left[\hat{\theta}{m}^{2}\right]-\mathbb{E}^{2}\left[\hat{\theta}_{m}\right]
\end{aligned}
Adding these two expressions leaves us with expression (1).
### Polynomial regression
For this exercise we will be using generated data to better show the effects of the different polynomial orders.
The data is created using the make_polynomial_regression function.
```
%matplotlib inline
def generate_dataset(n=100, degree=1, noise=1, factors=None):
# Generates a dataset by adding random noise to a randomly
# generated polynomial function.
x = np.random.uniform(low=-1, high=1, size=n)
factors = np.random.uniform(0, 10, degree+1)
y = np.zeros(x.shape)
for idx in range(degree+1):
y += factors[idx] * (x ** idx)
# add noise
y += np.random.normal(-noise, noise, n)
return x, y
# load generated data
np.random.seed(0)
X, y = generate_dataset(n=100, degree=4, noise=1.5)
plt.plot(X, y, 'r.', markersize=12);
```
Implement polynomial regression using the `sklearn.preprocessing.PolynomialFeatures` transformation. Using the `sklearn.grid_search.GridSearchCV` class, perform a grid search of the polynomial order hyperparameter space with cross-validation and report the performance on an independent test set.
Plot a learning curve that show the validation accuracy as a function of the polynomial order.
<p><font color='#770a0a'>Which models have a high bias, and which models have high variance? Motivate your answer.</font><p>
Repeat this experiment, this time using the diabetes dataset instead of the generated data.
```
# Implementing polynomial regression
# Polynomial regression only adds new features to the original data samples, where the new features are combination of polynomials of the original features.
# Source: https://programmer.group/polynomial-regression-and-model-generalization.html
from sklearn.model_selection import train_test_split
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import PolynomialFeatures, StandardScaler
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import mean_squared_error
# Split data into test and train data
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.5, random_state=1)
# Since polynomial regression is a special kind of linear regression, by the fact that we create polynomial features before creating linear regression
# A pipeline can combine these two steps (PolynomialFeatures and LinearRegression)
# Source: https://towardsdatascience.com/polynomial-regression-with-scikit-learn-what-you-should-know-bed9d3296f2
def polynomial_regression(degree):
model = make_pipeline(StandardScaler(), PolynomialFeatures(), LinearRegression())
parameter = {'polynomialfeatures__degree': [degree]}
return model, parameter
mse_train_tot = []
mse_test_tot = []
degrees = range(1, 11)
# Visualization of polynomial regression model and data
space = np.linspace(-1,1,101).reshape(-1,1)
plt.plot(X, y, 'b.', markersize=8)
for degree in degrees:
poly_reg, parameter = polynomial_regression(degree)
# Grid search includes cross validation
grid_search = GridSearchCV(estimator = poly_reg, param_grid = parameter, cv = 10, scoring= 'neg_mean_squared_error')
grid_search.fit(X_train.reshape(-1,1), y_train)
# Visualization of polynomial regression model and data
plt.plot(space, grid_search.predict(space), color = 'red')
grid_predict_train = grid_search.predict(X_train.reshape(-1,1))
grid_predict_test = grid_search.predict(X_test.reshape(-1,1))
mse_train = mean_squared_error(y_test, grid_predict_train)
mse_test = mean_squared_error(y_test, grid_predict_test)
mse_train_tot.append(mse_train)
mse_test_tot.append(mse_test)
print(mse_test_tot, mse_train_tot) # waarom is de training error groter dan de test error?
plt.plot(degrees, mse_train_tot, label='train mse')
plt.plot(degrees, mse_test_tot, label='test mse')
plt.xticks(degrees)
plt.xlabel('Degree of polynomial regression')
plt.ylabel('Mean squared error')
plt.legend()
plt.title('True Degree: 4 Best estimated Degree: ' + str(np.argmin(mse_test_tot)+1))
grid_predict = grid_search.predict(X_test.reshape(-1,1))
mse = mean_squared_error(y_test, grid_predict)
print(y_test)
"""print(classification_report(y_test, grid_predict))
"""
plt.plot(X_test, y_test, 'b.', markersize=8)
plt.plot(space, grid_search.predict(space), color = 'red')
```
### ROC curve analysis
A common method to evaluate binary classifiers is the receiver operating characteristic (ROC) curve. Similar to the week one practicals, implement a $k$-NN classifier on the breast cancer dataset, however, his time use the $k$-NN pipeline from the preliminary. Train the model for different values of $k$ and evaluate their respective performance with an ROC curve, use the `sklearn.metrics.roc_curve` function.
```
# Import the libraries needed
from sklearn.model_selection import train_test_split
from sklearn import datasets, neighbors, metrics
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import Pipeline
def knn_classifier(X_train, y_train, X_test, k):
"""
Train the kNN algorithm with k nearest neighbors in order to solve a binary classifier problem
and to predict the class for new unseen data
:param X_train: training data
:param y_train: corresponding values of training data
:param X_test: test data to test the trained kNN
:param k: amount of neighbors
:return: y_pred y predicted using the kNN
"""
# Create a scaler to transform all features to have a zero mean and unit variance
scaler = StandardScaler()
scaler.fit(X_train)
X_train_scaled = scaler.transform(X_train)
# initialize a k-NN classifier
knn = neighbors.KNeighborsClassifier(n_neighbors=k)
# create a model using a pipeline for the knn classifier and the scaler
model = Pipeline([
("scaler", scaler),
("knn", knn)
])
# train the model using the training dataset
model.fit(X_train, y_train)
# make predictions using the testing dataset
prediction = model.predict(X_test)
y_pred = model.predict_proba(X_test)[:,1]
return y_pred
def compute_ROC(y_true, y_pred, k, plot=True, zoom=False):
"""
Plot the ROC curve given the true values y_true and the predicted
values y_pred.
:param y_true: ground truth
:param y_pred: predictions
:param plot: true if the ROC curve has to be plotted
:param plot: true if the zoom function has to be used
:return: fpr (False positive rate) tpr (True positive rate) auc_score (Area under the ROC curve)
"""
# Compute the false positive rate and the true positive rate
fpr, tpr, _ = metrics.roc_curve(y_test, y_pred)
# Compute the area under the curve using the fpr and tpr
auc_score = metrics.auc(fpr, tpr)
if plot:
plt.plot(fpr, tpr, label='ROC (AUC = {:.4f}) for k = {:.0f} '.format(auc_score, k))
plt.plot([0, 1], [0, 1],'r--')
plt.legend(loc='lower right')
plt.title("ROC Curve")
plt.xlabel("False Positive Rate")
plt.ylabel("True Positive Rate")
if zoom:
plt.xlim(-0.02,0.1)
plt.ylim(0.9,1.05)
return fpr, tpr, auc_score
# Load the dataset
breast_cancer = datasets.load_breast_cancer()
# Load the features in X and the targets in y
X = breast_cancer.data
y = breast_cancer.target
# Make use of the train_test_split() utility function to split the data in train and test
X_train, X_test, y_train, y_test = train_test_split(X, y, stratify=y, random_state=40, test_size = 0.2)
# Use this to enlarge the figure
plt.figure(figsize = (10,10))
for k in range(1, 11):
y_pred = knn_classifier(X_train, y_train, X_test, k)
compute_ROC(y_test, y_pred, k, plot=True, zoom=False)
# generate some test objecys
A = np.zeros((32, 32))
A[10:-10, 10:-10] = 1
B = np.zeros((32, 32))
B[5:-15, 5:-15] = 1
dice = 2*np.sum(A*B)/(np.sum(A)+np.sum(B))
# display the results
plt.plot()
plt.imshow(A)
plt.imshow(B, alpha=0.7)
print(dice)
```
Show that the $F_1$ score, which is the harmonic mean of precision and recall, is equivalent to the Dice similarity coefficient</font><p>
Answer:
When the Dice similarity coefficient (DSC) is applied to boolean data, the expression becomes:
$\text{DSC}=\frac{2TP}{2TP+FN+FP}$. (Source: https://en.wikipedia.org/wiki/S%C3%B8rensen%E2%80%93Dice_coefficient) #weghalen?
The expression of the $F_1$ score is the following:
$F_1 = 2\cdot \frac{Precision \cdot Recal}{Precision + Recal}$
The precision is equal to $\frac{TP}{TP+FP}$, and the recal (also known as sensitivity) is equal to $\frac{TP}{TP+FN}$.
Filling this in for the equation of the $F_1$ score gives us:
$F_1 = 2\cdot\frac{\frac{TP}{TP+FP}\cdot\frac{TP}{TP+FN}}{\frac{TP}{TP+FP}+\frac{TP}{TP+FN}}$.
This can be simplified to: %als jullie willen kan ik hier ook de tussenstappen nog zetten
$F_1 = 2\cdot\frac{TP^2}{2TP^2 + TPFN + TPFP}$.
When dividing both the numerator and denominator by $TP$, we obtain the following formula:
$F_1 = \frac{2TP}{2TP+FN+FP}$.
This is equal to the expression for the Dice similarity coefficient for boolean data.
| github_jupyter |
# Covid Tracking Project ([The website](https://covidtracking.com/))
## Dr. Tirthajyoti Sarkar, Fremont, CA, April 2020
---
## What is _"Covid Tracking Project"_?
The following descriptions are collected directly from the [project website](https://covidtracking.com/).
"**The COVID Tracking Project collects and publishes the most complete testing data available for US states and territories**.
It has been cited in and used by major media companies and agencies.

The COVID Tracking Project exists because every person, newsroom, and government agency in the United States deserves access to the most complete picture of COVID-19 testing data that can be assembled.
**Understanding the shape, speed, and location of regional outbreaks requires the entire testing picture**: how many people have actually been tested in each state/territory, when they were tested, and what their results were. That’s where their data comes in.
**Because we have no complete official account of COVID-19 testing data in the US, we have to get this data from the public health authority in each US state and territory (and the District of Columbia)**. Each of these authorities reports its data in its own way, including online dashboards, data tables, PDFs, press conferences, tweets, and Facebook posts. And while many states and territories have slowly moved toward more standard ways of reporting, the actual categories of information are still in flux.
Our data team uses website-scrapers and trackers to alert us to changes, but the **actual updates to our dataset are done manually by careful humans** who double-check each change and extensively annotate changes areas of ambiguity. The work of data-gathering from official sources is also now supplemented by a fast-growing group of reporters who are constantly pushing authorities to release more comprehensive information."
Some of the visualizations example are shown below,
## What is this Notebook about?
In this Jupyter notebook, we will show the step-by-step code and examples of creating impactful visualizations and charts using simple Python programming.
## My COVID-19 Github repo
Other COVID-19 related Notebooks are in my **[Github repo here](https://github.com/tirthajyoti/Covid-19-analysis)**
---
## Code and demo
### Import libraries
```
import numpy as np
import pandas as pd
import io
import requests
import matplotlib.pyplot as plt
```
### Define the URL and get the content
```
url="http://covidtracking.com/api/states/daily.csv"
s=requests.get(url).content
```
### Read in a Pandas DataFRame
```
df = pd.read_csv(io.StringIO(s.decode('utf-8')))
```
### Converts dates to a specific format
```
df['date'] = pd.to_datetime(df['date'], format='%Y%m%d')
```
### Drops unnecessary column(s)
```
df.drop(['dateChecked'],axis=1,inplace=True)
```
### Converts the `state` data to string-type
```
df['state']=df['state'].apply(str)
```
### Checking the general information
```
df.info()
```
### Peeking into the data
```
df.head()
```
### Replacing the `NaN` by -1
```
df.fillna(value=-1, inplace=True)
df.head()
```
### Function to plot a bar chart of the given variable/state
```
def plot_var(var='positiveIncrease',
state='NY'):
"""
Plots a bar chart of the given variable over the date range
"""
assert type(var)==str, "Expected string as the variable name"
assert type(state)==str, "Expected string as the state name"
y = df[df['state']==state][var]
x = df[df['state']==state]['date']
plt.figure(figsize=(12,4))
plt.title("Plot of \"{}\" for {}".format(var,state),fontsize=18)
plt.bar(x=x,height=y,edgecolor='k',color='orange')
plt.grid(True)
plt.xticks(fontsize=14,rotation=45)
plt.yticks(fontsize=14)
plt.show()
```
### Some test plots...
```
plot_var()
plot_var('hospitalizedIncrease','GA')
plot_var('positiveIncrease','GA')
plot_var('totalTestResultsIncrease','MA')
```
### Function to create scatter plot of two variables for a given state
```
def plot_xy(varx='totalTestResultsIncrease',
vary='positiveIncrease',
state='NY'):
"""
Plots a bar chart of the given variable over the date range
"""
assert type(varx)==str, "Expected string as the variable x name"
assert type(vary)==str, "Expected string as the variable y name"
y = df[df['state']==state][vary]
x = df[df['state']==state][varx]
if (x.nunique()!=1) and (y.nunique()!=1):
plt.figure(figsize=(12,4))
plt.title("Plot of \"{}\" vs. \"{}\" for {}".format(varx,vary,state),fontsize=18)
plt.scatter(x=x,y=y,edgecolor='k',color='lightgreen',s=100)
plt.grid(True)
plt.xticks(fontsize=14,rotation=45)
plt.yticks(fontsize=14)
plt.show()
else:
print("Some of the data unavailable for a scatter plot. Sorry!")
```
### Test some plots...
```
plot_xy(state='NY')
plot_xy('hospitalized','death','GA')
plot_xy('hospitalized','death','CA')
```
### Testing tracker function
Testing is crucially important to track the spread of the virus and contain it. Let us see how states the doing on this aspect relatively.
```
def plotTesting(lst_states=['NY','CA','MA','TX','PA']):
"""
Plots the cumulative testing done by the given list of states
"""
legends = []
plt.figure(figsize=(10,5))
plt.title("Total test results",fontsize=18)
for s in lst_states:
data = np.array(df[df['state']==s]['totalTestResults'])[-1::-1]
slope = int((data[-1]-data[0])/len(data))
plt.plot(data,linewidth=2)
plt.text(x=len(data)-2,y=data[-1]*1.05,s=s,fontsize=14)
legends.append(str(slope)+" tests/day in " + s)
plt.legend(legends,fontsize=14)
plt.grid(True)
plt.xlim(0,len(data)+2)
plt.xticks(fontsize=16)
plt.yticks(fontsize=16)
plt.xlabel("Days",fontsize=16)
plt.ylabel("Total test results",fontsize=16)
plt.show()
plotTesting()
```
### Function to compute fatality ratio (with the latest data)
```
def fatality_ratio(state='CA'):
"""
Computes the fatality ratio for the given state
Fatality ratio is the ratio of total dead to total positive case
"""
date = df.iloc[0]['date']
try:
d = float(df[(df['state']==state) & (df['date']==date)]['death'])
p = float(df[(df['state']==state) & (df['date']==date)]['positive'])
except:
print("Could not retrieve the necessary information")
if (d!=-1.0) and (p!=-1.0) and (p!=0):
return round(d/p,3)
else:
return -1
```
### Function to compute hospitalization ratio (with the latest data)
```
def hospitalization_ratio(state='NY'):
"""
Computes the hospitalization ratio for the given state
Hospitalization ratio is the ratio of total hospitalized to total positive case
"""
date = df.iloc[0]['date']
try:
h = float(df[(df['state']==state) & (df['date']==date)]['hospitalized'])
p = float(df[(df['state']==state) & (df['date']==date)]['positive'])
except:
print("Could not retrieve the necessary information")
if (h!=-1.0) and (p!=-1.0) and (p!=0):
return round(h/p,3)
else:
return -1
```
### Function to compute positive case/total test ratio (with the latest data)
```
def positiveTest_ratio(state='NY'):
"""
Computes the test-positive ratio for the given state
Test-positive ratio is the ratio of total positive cases to total number of tests
"""
date = df.iloc[0]['date']
try:
p = float(df[(df['state']==state) & (df['date']==date)]['positive'])
t = float(df[(df['state']==state) & (df['date']==date)]['totalTestResults'])
except:
print("Could not retrieve the necessary information")
return -1
if (p!=-1.0) and (t!=-1.0) and (t!=0):
return round(p/t,3)
else:
return -1
```
### Function to calculate recovery ratio (with the latest data)
```
def recovery_ratio(state='NY'):
"""
Computes the recovery ratio for the given state
Recovery ratio is the ratio of total recovered cases to total positive cases
"""
date = df.iloc[0]['date']
try:
r = float(df[(df['state']==state) & (df['date']==date)]['recovered'])
p = float(df[(df['state']==state) & (df['date']==date)]['positive'])
except:
print("Could not retrieve the necessary information")
return -1
if (r!=-1.0) and (p!=-1.0) and (p!=0):
return round(r/p,3)
else:
return -1
```
### Fatality ratio chart
We will plot the chart with six states,
- California
- New York
- Michigan
- Massachusetts
- Pennsylvania
- Illinois
```
states = ['CA','NY','MI','MA','PA','IL']
fr,x = [],[]
for s in states:
data = fatality_ratio(s)
if data!=-1:
fr.append(data)
x.append(s)
plt.figure(figsize=(8,4))
plt.title("Fatality ratio chart",fontsize=18)
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
plt.bar(x=x,height=fr,color='red',
edgecolor='k',linewidth=2)
plt.show()
```
### Hospitalization chart
The next hospitalization rate chart is tried for the same set of states but only two of them reports proper data. Therefore, the code plots the chart only for those states.
```
states = ['CA','NY','MI','MA','PA','IL']
hos,x = [],[]
for s in states:
data = hospitalization_ratio(s)
if data!=-1:
hos.append(data)
x.append(s)
plt.figure(figsize=(8,4))
plt.title("Hospitalization ratio chart",fontsize=18)
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
plt.bar(x=x,height=hos,color='brown',
edgecolor='k',linewidth=2)
plt.show()
```
### Test-positive ratio chart
```
states = ['CA','NY','MI','MA','PA','IL']
tp,x = [],[]
for s in states:
data = positiveTest_ratio(s)
if data!=-1:
tp.append(data)
x.append(s)
plt.bar(x=x,height=tp,color='blue',
edgecolor='k',linewidth=2)
plt.show()
```
### Bubble charts...
```
states = list(df['state'].unique())
for s in ['AS','GU','MP','PU','VI']:
try:
states.remove(s)
except:
pass
fr,x = [],[]
for s in states:
data = fatality_ratio(s)
if data!=-1:
fr.append(data)
x.append(s)
fr = np.array(fr)
plt.figure(figsize=(15,7))
plt.tick_params(
axis='x', # changes apply to the x-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
top=False, # ticks along the top edge are off
labelbottom=False)
plt.title("Fatality ratio across the states",fontsize=18)
plt.scatter(x=x,y=fr,
s=4e5*fr**2,
color='orange',edgecolor='red',alpha=0.75,linewidth=2.5)
#plt.xticks(rotation=45,fontsize=12)
for i,s in enumerate(x):
plt.annotate(s=s,xy=(x[i],fr[i]))
plt.ylim(0,0.12)
plt.yticks(fontsize=16)
plt.grid(True,axis='y')
plt.show()
states = list(df['state'].unique())
for s in ['AS','GU','MP','PU','VI']:
try:
states.remove(s)
except:
pass
tp,x = [],[]
for s in states:
data = positiveTest_ratio(s)
if data!=-1:
tp.append(data)
x.append(s)
tp = np.array(tp)
plt.figure(figsize=(15,7))
plt.tick_params(
axis='x', # changes apply to the x-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
top=False, # ticks along the top edge are off
labelbottom=False)
plt.title("Test-positive ratio across the states",fontsize=18)
plt.scatter(x=x,y=tp,
s=2e4*tp**2,
color='blue',edgecolor='red',alpha=0.5,linewidth=2)
plt.xticks(rotation=90,fontsize=12)
for i,s in enumerate(x):
plt.annotate(s=s,xy=(x[i],tp[i]))
plt.ylim(0,0.6)
plt.yticks(fontsize=16)
plt.grid(True,axis='y')
plt.show()
```
### The crucial question: _"Which states have 14 days of decreasing case counts?"_
```
def caseCountsdecrease(days=14,state='NY'):
"""
Determines whether the given state has a decreasing case counts for given number of days
Arguments:
days: Number of days to go back
state: Name of the state (a string)
Returns:
A tuple containing the successive difference vector (of new cases) and
the number of negative quantities in that vector. When all the quantities are negative,
the state has shown consistent decrease in new cases for the given number of days.
"""
positiveIncrease = np.array(df[df['state']==state]['positiveIncrease'][:days+1])[-1::-1]
diff = np.diff(positiveIncrease)
countofNeg = np.sum(diff <= 0, axis=0)
return (countofNeg, diff)
```
### Plot for a few states
We will note that **no state, so far, has shown a consistent decrease of new cases for the last 14 days**!
```
states = ['CA','MI','MA','PA','IL','LA']
cd = []
x = np.arange(1,15,1)
plt.figure(figsize=(10,6))
plt.title("Last 14 days successive difference in new positive cases \n(more negative numbers is better)",
fontsize=18)
for s in states:
_,data = caseCountsdecrease(days=14,state=s)
plt.plot(x,data,linewidth=2)
plt.legend(states,fontsize=16,ncol=2)
plt.grid(True)
plt.xticks(fontsize=16)
plt.yticks(fontsize=16)
plt.xlabel("Last 14 days",fontsize=16)
plt.ylabel("Successive difference in new cases",fontsize=16)
plt.hlines(y=0,xmin=0,xmax=15,linestyles='--',lw=3)
plt.show()
```
| github_jupyter |
# RadarCOVID-Report
## Data Extraction
```
import datetime
import json
import logging
import os
import shutil
import tempfile
import textwrap
import uuid
import matplotlib.pyplot as plt
import matplotlib.ticker
import numpy as np
import pandas as pd
import pycountry
import retry
import seaborn as sns
%matplotlib inline
current_working_directory = os.environ.get("PWD")
if current_working_directory:
os.chdir(current_working_directory)
sns.set()
matplotlib.rcParams["figure.figsize"] = (15, 6)
extraction_datetime = datetime.datetime.utcnow()
extraction_date = extraction_datetime.strftime("%Y-%m-%d")
extraction_previous_datetime = extraction_datetime - datetime.timedelta(days=1)
extraction_previous_date = extraction_previous_datetime.strftime("%Y-%m-%d")
extraction_date_with_hour = datetime.datetime.utcnow().strftime("%Y-%m-%d@%H")
current_hour = datetime.datetime.utcnow().hour
are_today_results_partial = current_hour != 23
```
### Constants
```
from Modules.ExposureNotification import exposure_notification_io
spain_region_country_code = "ES"
germany_region_country_code = "DE"
default_backend_identifier = spain_region_country_code
backend_generation_days = 7 * 2
daily_summary_days = 7 * 4 * 3
daily_plot_days = 7 * 4
tek_dumps_load_limit = daily_summary_days + 1
```
### Parameters
```
environment_backend_identifier = os.environ.get("RADARCOVID_REPORT__BACKEND_IDENTIFIER")
if environment_backend_identifier:
report_backend_identifier = environment_backend_identifier
else:
report_backend_identifier = default_backend_identifier
report_backend_identifier
environment_enable_multi_backend_download = \
os.environ.get("RADARCOVID_REPORT__ENABLE_MULTI_BACKEND_DOWNLOAD")
if environment_enable_multi_backend_download:
report_backend_identifiers = None
else:
report_backend_identifiers = [report_backend_identifier]
report_backend_identifiers
environment_invalid_shared_diagnoses_dates = \
os.environ.get("RADARCOVID_REPORT__INVALID_SHARED_DIAGNOSES_DATES")
if environment_invalid_shared_diagnoses_dates:
invalid_shared_diagnoses_dates = environment_invalid_shared_diagnoses_dates.split(",")
else:
invalid_shared_diagnoses_dates = []
invalid_shared_diagnoses_dates
```
### COVID-19 Cases
```
report_backend_client = \
exposure_notification_io.get_backend_client_with_identifier(
backend_identifier=report_backend_identifier)
@retry.retry(tries=10, delay=10, backoff=1.1, jitter=(0, 10))
def download_cases_dataframe():
return pd.read_csv("https://raw.githubusercontent.com/owid/covid-19-data/master/public/data/owid-covid-data.csv")
confirmed_df_ = download_cases_dataframe()
confirmed_df_.iloc[0]
confirmed_df = confirmed_df_.copy()
confirmed_df = confirmed_df[["date", "new_cases", "iso_code"]]
confirmed_df.rename(
columns={
"date": "sample_date",
"iso_code": "country_code",
},
inplace=True)
def convert_iso_alpha_3_to_alpha_2(x):
try:
return pycountry.countries.get(alpha_3=x).alpha_2
except Exception as e:
logging.info(f"Error converting country ISO Alpha 3 code '{x}': {repr(e)}")
return None
confirmed_df["country_code"] = confirmed_df.country_code.apply(convert_iso_alpha_3_to_alpha_2)
confirmed_df.dropna(inplace=True)
confirmed_df["sample_date"] = pd.to_datetime(confirmed_df.sample_date, dayfirst=True)
confirmed_df["sample_date"] = confirmed_df.sample_date.dt.strftime("%Y-%m-%d")
confirmed_df.sort_values("sample_date", inplace=True)
confirmed_df.tail()
confirmed_days = pd.date_range(
start=confirmed_df.iloc[0].sample_date,
end=extraction_datetime)
confirmed_days_df = pd.DataFrame(data=confirmed_days, columns=["sample_date"])
confirmed_days_df["sample_date_string"] = \
confirmed_days_df.sample_date.dt.strftime("%Y-%m-%d")
confirmed_days_df.tail()
def sort_source_regions_for_display(source_regions: list) -> list:
if report_backend_identifier in source_regions:
source_regions = [report_backend_identifier] + \
list(sorted(set(source_regions).difference([report_backend_identifier])))
else:
source_regions = list(sorted(source_regions))
return source_regions
report_source_regions = report_backend_client.source_regions_for_date(
date=extraction_datetime.date())
report_source_regions = sort_source_regions_for_display(
source_regions=report_source_regions)
report_source_regions
def get_cases_dataframe(source_regions_for_date_function, columns_suffix=None):
source_regions_at_date_df = confirmed_days_df.copy()
source_regions_at_date_df["source_regions_at_date"] = \
source_regions_at_date_df.sample_date.apply(
lambda x: source_regions_for_date_function(date=x))
source_regions_at_date_df.sort_values("sample_date", inplace=True)
source_regions_at_date_df["_source_regions_group"] = source_regions_at_date_df. \
source_regions_at_date.apply(lambda x: ",".join(sort_source_regions_for_display(x)))
source_regions_at_date_df.tail()
#%%
source_regions_for_summary_df_ = \
source_regions_at_date_df[["sample_date", "_source_regions_group"]].copy()
source_regions_for_summary_df_.rename(columns={"_source_regions_group": "source_regions"}, inplace=True)
source_regions_for_summary_df_.tail()
#%%
confirmed_output_columns = ["sample_date", "new_cases", "covid_cases"]
confirmed_output_df = pd.DataFrame(columns=confirmed_output_columns)
for source_regions_group, source_regions_group_series in \
source_regions_at_date_df.groupby("_source_regions_group"):
source_regions_set = set(source_regions_group.split(","))
confirmed_source_regions_set_df = \
confirmed_df[confirmed_df.country_code.isin(source_regions_set)].copy()
confirmed_source_regions_group_df = \
confirmed_source_regions_set_df.groupby("sample_date").new_cases.sum() \
.reset_index().sort_values("sample_date")
confirmed_source_regions_group_df = \
confirmed_source_regions_group_df.merge(
confirmed_days_df[["sample_date_string"]].rename(
columns={"sample_date_string": "sample_date"}),
how="right")
confirmed_source_regions_group_df["new_cases"] = \
confirmed_source_regions_group_df["new_cases"].clip(lower=0)
confirmed_source_regions_group_df["covid_cases"] = \
confirmed_source_regions_group_df.new_cases.rolling(7, min_periods=0).mean().round()
confirmed_source_regions_group_df = \
confirmed_source_regions_group_df[confirmed_output_columns]
confirmed_source_regions_group_df = confirmed_source_regions_group_df.replace(0, np.nan)
confirmed_source_regions_group_df.fillna(method="ffill", inplace=True)
confirmed_source_regions_group_df = \
confirmed_source_regions_group_df[
confirmed_source_regions_group_df.sample_date.isin(
source_regions_group_series.sample_date_string)]
confirmed_output_df = confirmed_output_df.append(confirmed_source_regions_group_df)
result_df = confirmed_output_df.copy()
result_df.tail()
#%%
result_df.rename(columns={"sample_date": "sample_date_string"}, inplace=True)
result_df = confirmed_days_df[["sample_date_string"]].merge(result_df, how="left")
result_df.sort_values("sample_date_string", inplace=True)
result_df.fillna(method="ffill", inplace=True)
result_df.tail()
#%%
result_df[["new_cases", "covid_cases"]].plot()
if columns_suffix:
result_df.rename(
columns={
"new_cases": "new_cases_" + columns_suffix,
"covid_cases": "covid_cases_" + columns_suffix},
inplace=True)
return result_df, source_regions_for_summary_df_
confirmed_eu_df, source_regions_for_summary_df = get_cases_dataframe(
report_backend_client.source_regions_for_date)
confirmed_es_df, _ = get_cases_dataframe(
lambda date: [spain_region_country_code],
columns_suffix=spain_region_country_code.lower())
```
### Extract API TEKs
```
raw_zip_path_prefix = "Data/TEKs/Raw/"
base_backend_identifiers = [report_backend_identifier]
multi_backend_exposure_keys_df = \
exposure_notification_io.download_exposure_keys_from_backends(
backend_identifiers=report_backend_identifiers,
generation_days=backend_generation_days,
fail_on_error_backend_identifiers=base_backend_identifiers,
save_raw_zip_path_prefix=raw_zip_path_prefix)
multi_backend_exposure_keys_df["region"] = multi_backend_exposure_keys_df["backend_identifier"]
multi_backend_exposure_keys_df.rename(
columns={
"generation_datetime": "sample_datetime",
"generation_date_string": "sample_date_string",
},
inplace=True)
multi_backend_exposure_keys_df.head()
early_teks_df = multi_backend_exposure_keys_df[
multi_backend_exposure_keys_df.rolling_period < 144].copy()
early_teks_df["rolling_period_in_hours"] = early_teks_df.rolling_period / 6
early_teks_df[early_teks_df.sample_date_string != extraction_date] \
.rolling_period_in_hours.hist(bins=list(range(24)))
early_teks_df[early_teks_df.sample_date_string == extraction_date] \
.rolling_period_in_hours.hist(bins=list(range(24)))
multi_backend_exposure_keys_df = multi_backend_exposure_keys_df[[
"sample_date_string", "region", "key_data"]]
multi_backend_exposure_keys_df.head()
active_regions = \
multi_backend_exposure_keys_df.groupby("region").key_data.nunique().sort_values().index.unique().tolist()
active_regions
multi_backend_summary_df = multi_backend_exposure_keys_df.groupby(
["sample_date_string", "region"]).key_data.nunique().reset_index() \
.pivot(index="sample_date_string", columns="region") \
.sort_index(ascending=False)
multi_backend_summary_df.rename(
columns={"key_data": "shared_teks_by_generation_date"},
inplace=True)
multi_backend_summary_df.rename_axis("sample_date", inplace=True)
multi_backend_summary_df = multi_backend_summary_df.fillna(0).astype(int)
multi_backend_summary_df = multi_backend_summary_df.head(backend_generation_days)
multi_backend_summary_df.head()
def compute_keys_cross_sharing(x):
teks_x = x.key_data_x.item()
common_teks = set(teks_x).intersection(x.key_data_y.item())
common_teks_fraction = len(common_teks) / len(teks_x)
return pd.Series(dict(
common_teks=common_teks,
common_teks_fraction=common_teks_fraction,
))
multi_backend_exposure_keys_by_region_df = \
multi_backend_exposure_keys_df.groupby("region").key_data.unique().reset_index()
multi_backend_exposure_keys_by_region_df["_merge"] = True
multi_backend_exposure_keys_by_region_combination_df = \
multi_backend_exposure_keys_by_region_df.merge(
multi_backend_exposure_keys_by_region_df, on="_merge")
multi_backend_exposure_keys_by_region_combination_df.drop(
columns=["_merge"], inplace=True)
if multi_backend_exposure_keys_by_region_combination_df.region_x.nunique() > 1:
multi_backend_exposure_keys_by_region_combination_df = \
multi_backend_exposure_keys_by_region_combination_df[
multi_backend_exposure_keys_by_region_combination_df.region_x !=
multi_backend_exposure_keys_by_region_combination_df.region_y]
multi_backend_exposure_keys_cross_sharing_df = \
multi_backend_exposure_keys_by_region_combination_df \
.groupby(["region_x", "region_y"]) \
.apply(compute_keys_cross_sharing) \
.reset_index()
multi_backend_cross_sharing_summary_df = \
multi_backend_exposure_keys_cross_sharing_df.pivot_table(
values=["common_teks_fraction"],
columns="region_x",
index="region_y",
aggfunc=lambda x: x.item())
multi_backend_cross_sharing_summary_df
multi_backend_without_active_region_exposure_keys_df = \
multi_backend_exposure_keys_df[multi_backend_exposure_keys_df.region != report_backend_identifier]
multi_backend_without_active_region = \
multi_backend_without_active_region_exposure_keys_df.groupby("region").key_data.nunique().sort_values().index.unique().tolist()
multi_backend_without_active_region
exposure_keys_summary_df = multi_backend_exposure_keys_df[
multi_backend_exposure_keys_df.region == report_backend_identifier]
exposure_keys_summary_df.drop(columns=["region"], inplace=True)
exposure_keys_summary_df = \
exposure_keys_summary_df.groupby(["sample_date_string"]).key_data.nunique().to_frame()
exposure_keys_summary_df = \
exposure_keys_summary_df.reset_index().set_index("sample_date_string")
exposure_keys_summary_df.sort_index(ascending=False, inplace=True)
exposure_keys_summary_df.rename(columns={"key_data": "shared_teks_by_generation_date"}, inplace=True)
exposure_keys_summary_df.head()
```
### Dump API TEKs
```
tek_list_df = multi_backend_exposure_keys_df[
["sample_date_string", "region", "key_data"]].copy()
tek_list_df["key_data"] = tek_list_df["key_data"].apply(str)
tek_list_df.rename(columns={
"sample_date_string": "sample_date",
"key_data": "tek_list"}, inplace=True)
tek_list_df = tek_list_df.groupby(
["sample_date", "region"]).tek_list.unique().reset_index()
tek_list_df["extraction_date"] = extraction_date
tek_list_df["extraction_date_with_hour"] = extraction_date_with_hour
tek_list_path_prefix = "Data/TEKs/"
tek_list_current_path = tek_list_path_prefix + f"/Current/RadarCOVID-TEKs.json"
tek_list_daily_path = tek_list_path_prefix + f"Daily/RadarCOVID-TEKs-{extraction_date}.json"
tek_list_hourly_path = tek_list_path_prefix + f"Hourly/RadarCOVID-TEKs-{extraction_date_with_hour}.json"
for path in [tek_list_current_path, tek_list_daily_path, tek_list_hourly_path]:
os.makedirs(os.path.dirname(path), exist_ok=True)
tek_list_base_df = tek_list_df[tek_list_df.region == report_backend_identifier]
tek_list_base_df.drop(columns=["extraction_date", "extraction_date_with_hour"]).to_json(
tek_list_current_path,
lines=True, orient="records")
tek_list_base_df.drop(columns=["extraction_date_with_hour"]).to_json(
tek_list_daily_path,
lines=True, orient="records")
tek_list_base_df.to_json(
tek_list_hourly_path,
lines=True, orient="records")
tek_list_base_df.head()
```
### Load TEK Dumps
```
import glob
def load_extracted_teks(mode, region=None, limit=None) -> pd.DataFrame:
extracted_teks_df = pd.DataFrame(columns=["region"])
file_paths = list(reversed(sorted(glob.glob(tek_list_path_prefix + mode + "/RadarCOVID-TEKs-*.json"))))
if limit:
file_paths = file_paths[:limit]
for file_path in file_paths:
logging.info(f"Loading TEKs from '{file_path}'...")
iteration_extracted_teks_df = pd.read_json(file_path, lines=True)
extracted_teks_df = extracted_teks_df.append(
iteration_extracted_teks_df, sort=False)
extracted_teks_df["region"] = \
extracted_teks_df.region.fillna(spain_region_country_code).copy()
if region:
extracted_teks_df = \
extracted_teks_df[extracted_teks_df.region == region]
return extracted_teks_df
daily_extracted_teks_df = load_extracted_teks(
mode="Daily",
region=report_backend_identifier,
limit=tek_dumps_load_limit)
daily_extracted_teks_df.head()
exposure_keys_summary_df_ = daily_extracted_teks_df \
.sort_values("extraction_date", ascending=False) \
.groupby("sample_date").tek_list.first() \
.to_frame()
exposure_keys_summary_df_.index.name = "sample_date_string"
exposure_keys_summary_df_["tek_list"] = \
exposure_keys_summary_df_.tek_list.apply(len)
exposure_keys_summary_df_ = exposure_keys_summary_df_ \
.rename(columns={"tek_list": "shared_teks_by_generation_date"}) \
.sort_index(ascending=False)
exposure_keys_summary_df = exposure_keys_summary_df_
exposure_keys_summary_df.head()
```
### Daily New TEKs
```
tek_list_df = daily_extracted_teks_df.groupby("extraction_date").tek_list.apply(
lambda x: set(sum(x, []))).reset_index()
tek_list_df = tek_list_df.set_index("extraction_date").sort_index(ascending=True)
tek_list_df.head()
def compute_teks_by_generation_and_upload_date(date):
day_new_teks_set_df = tek_list_df.copy().diff()
try:
day_new_teks_set = day_new_teks_set_df[
day_new_teks_set_df.index == date].tek_list.item()
except ValueError:
day_new_teks_set = None
if pd.isna(day_new_teks_set):
day_new_teks_set = set()
day_new_teks_df = daily_extracted_teks_df[
daily_extracted_teks_df.extraction_date == date].copy()
day_new_teks_df["shared_teks"] = \
day_new_teks_df.tek_list.apply(lambda x: set(x).intersection(day_new_teks_set))
day_new_teks_df["shared_teks"] = \
day_new_teks_df.shared_teks.apply(len)
day_new_teks_df["upload_date"] = date
day_new_teks_df.rename(columns={"sample_date": "generation_date"}, inplace=True)
day_new_teks_df = day_new_teks_df[
["upload_date", "generation_date", "shared_teks"]]
day_new_teks_df["generation_to_upload_days"] = \
(pd.to_datetime(day_new_teks_df.upload_date) -
pd.to_datetime(day_new_teks_df.generation_date)).dt.days
day_new_teks_df = day_new_teks_df[day_new_teks_df.shared_teks > 0]
return day_new_teks_df
shared_teks_generation_to_upload_df = pd.DataFrame()
for upload_date in daily_extracted_teks_df.extraction_date.unique():
shared_teks_generation_to_upload_df = \
shared_teks_generation_to_upload_df.append(
compute_teks_by_generation_and_upload_date(date=upload_date))
shared_teks_generation_to_upload_df \
.sort_values(["upload_date", "generation_date"], ascending=False, inplace=True)
shared_teks_generation_to_upload_df.tail()
today_new_teks_df = \
shared_teks_generation_to_upload_df[
shared_teks_generation_to_upload_df.upload_date == extraction_date].copy()
today_new_teks_df.tail()
if not today_new_teks_df.empty:
today_new_teks_df.set_index("generation_to_upload_days") \
.sort_index().shared_teks.plot.bar()
generation_to_upload_period_pivot_df = \
shared_teks_generation_to_upload_df[
["upload_date", "generation_to_upload_days", "shared_teks"]] \
.pivot(index="upload_date", columns="generation_to_upload_days") \
.sort_index(ascending=False).fillna(0).astype(int) \
.droplevel(level=0, axis=1)
generation_to_upload_period_pivot_df.head()
new_tek_df = tek_list_df.diff().tek_list.apply(
lambda x: len(x) if not pd.isna(x) else None).to_frame().reset_index()
new_tek_df.rename(columns={
"tek_list": "shared_teks_by_upload_date",
"extraction_date": "sample_date_string",}, inplace=True)
new_tek_df.tail()
shared_teks_uploaded_on_generation_date_df = shared_teks_generation_to_upload_df[
shared_teks_generation_to_upload_df.generation_to_upload_days == 0] \
[["upload_date", "shared_teks"]].rename(
columns={
"upload_date": "sample_date_string",
"shared_teks": "shared_teks_uploaded_on_generation_date",
})
shared_teks_uploaded_on_generation_date_df.head()
estimated_shared_diagnoses_df = shared_teks_generation_to_upload_df \
.groupby(["upload_date"]).shared_teks.max().reset_index() \
.sort_values(["upload_date"], ascending=False) \
.rename(columns={
"upload_date": "sample_date_string",
"shared_teks": "shared_diagnoses",
})
invalid_shared_diagnoses_dates_mask = \
estimated_shared_diagnoses_df.sample_date_string.isin(invalid_shared_diagnoses_dates)
estimated_shared_diagnoses_df[invalid_shared_diagnoses_dates_mask] = 0
estimated_shared_diagnoses_df.head()
```
### Hourly New TEKs
```
hourly_extracted_teks_df = load_extracted_teks(
mode="Hourly", region=report_backend_identifier, limit=25)
hourly_extracted_teks_df.head()
hourly_new_tek_count_df = hourly_extracted_teks_df \
.groupby("extraction_date_with_hour").tek_list. \
apply(lambda x: set(sum(x, []))).reset_index().copy()
hourly_new_tek_count_df = hourly_new_tek_count_df.set_index("extraction_date_with_hour") \
.sort_index(ascending=True)
hourly_new_tek_count_df["new_tek_list"] = hourly_new_tek_count_df.tek_list.diff()
hourly_new_tek_count_df["new_tek_count"] = hourly_new_tek_count_df.new_tek_list.apply(
lambda x: len(x) if not pd.isna(x) else 0)
hourly_new_tek_count_df.rename(columns={
"new_tek_count": "shared_teks_by_upload_date"}, inplace=True)
hourly_new_tek_count_df = hourly_new_tek_count_df.reset_index()[[
"extraction_date_with_hour", "shared_teks_by_upload_date"]]
hourly_new_tek_count_df.head()
hourly_summary_df = hourly_new_tek_count_df.copy()
hourly_summary_df.set_index("extraction_date_with_hour", inplace=True)
hourly_summary_df = hourly_summary_df.fillna(0).astype(int).reset_index()
hourly_summary_df["datetime_utc"] = pd.to_datetime(
hourly_summary_df.extraction_date_with_hour, format="%Y-%m-%d@%H")
hourly_summary_df.set_index("datetime_utc", inplace=True)
hourly_summary_df = hourly_summary_df.tail(-1)
hourly_summary_df.head()
```
### Official Statistics
```
import requests
import pandas.io.json
official_stats_response = requests.get("https://radarcovid.covid19.gob.es/kpi/statistics/basics")
official_stats_response.raise_for_status()
official_stats_df_ = pandas.io.json.json_normalize(official_stats_response.json())
official_stats_df = official_stats_df_.copy()
official_stats_df["date"] = pd.to_datetime(official_stats_df["date"], dayfirst=True)
official_stats_df.head()
official_stats_column_map = {
"date": "sample_date",
"applicationsDownloads.totalAcummulated": "app_downloads_es_accumulated",
"communicatedContagions.totalAcummulated": "shared_diagnoses_es_accumulated",
}
accumulated_suffix = "_accumulated"
accumulated_values_columns = \
list(filter(lambda x: x.endswith(accumulated_suffix), official_stats_column_map.values()))
interpolated_values_columns = \
list(map(lambda x: x[:-len(accumulated_suffix)], accumulated_values_columns))
official_stats_df = \
official_stats_df[official_stats_column_map.keys()] \
.rename(columns=official_stats_column_map)
official_stats_df["extraction_date"] = extraction_date
official_stats_df.head()
official_stats_path = "Data/Statistics/Current/RadarCOVID-Statistics.json"
previous_official_stats_df = pd.read_json(official_stats_path, orient="records", lines=True)
previous_official_stats_df["sample_date"] = pd.to_datetime(previous_official_stats_df["sample_date"], dayfirst=True)
official_stats_df = official_stats_df.append(previous_official_stats_df)
official_stats_df.head()
official_stats_df = official_stats_df[~(official_stats_df.shared_diagnoses_es_accumulated == 0)]
official_stats_df.sort_values("extraction_date", ascending=False, inplace=True)
official_stats_df.drop_duplicates(subset=["sample_date"], keep="first", inplace=True)
official_stats_df.head()
official_stats_stored_df = official_stats_df.copy()
official_stats_stored_df["sample_date"] = official_stats_stored_df.sample_date.dt.strftime("%Y-%m-%d")
official_stats_stored_df.to_json(official_stats_path, orient="records", lines=True)
official_stats_df.drop(columns=["extraction_date"], inplace=True)
official_stats_df = confirmed_days_df.merge(official_stats_df, how="left")
official_stats_df.sort_values("sample_date", ascending=False, inplace=True)
official_stats_df.head()
official_stats_df[accumulated_values_columns] = \
official_stats_df[accumulated_values_columns] \
.astype(float).interpolate(limit_area="inside")
official_stats_df[interpolated_values_columns] = \
official_stats_df[accumulated_values_columns].diff(periods=-1)
official_stats_df.drop(columns="sample_date", inplace=True)
official_stats_df.head()
```
### Data Merge
```
result_summary_df = exposure_keys_summary_df.merge(
new_tek_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = result_summary_df.merge(
shared_teks_uploaded_on_generation_date_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = result_summary_df.merge(
estimated_shared_diagnoses_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = result_summary_df.merge(
official_stats_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = confirmed_eu_df.tail(daily_summary_days).merge(
result_summary_df, on=["sample_date_string"], how="left")
result_summary_df.head()
result_summary_df = confirmed_es_df.tail(daily_summary_days).merge(
result_summary_df, on=["sample_date_string"], how="left")
result_summary_df.head()
result_summary_df["sample_date"] = pd.to_datetime(result_summary_df.sample_date_string)
result_summary_df = result_summary_df.merge(source_regions_for_summary_df, how="left")
result_summary_df.set_index(["sample_date", "source_regions"], inplace=True)
result_summary_df.drop(columns=["sample_date_string"], inplace=True)
result_summary_df.sort_index(ascending=False, inplace=True)
result_summary_df.head()
with pd.option_context("mode.use_inf_as_na", True):
result_summary_df = result_summary_df.fillna(0).astype(int)
result_summary_df["teks_per_shared_diagnosis"] = \
(result_summary_df.shared_teks_by_upload_date / result_summary_df.shared_diagnoses).fillna(0)
result_summary_df["shared_diagnoses_per_covid_case"] = \
(result_summary_df.shared_diagnoses / result_summary_df.covid_cases).fillna(0)
result_summary_df["shared_diagnoses_per_covid_case_es"] = \
(result_summary_df.shared_diagnoses_es / result_summary_df.covid_cases_es).fillna(0)
result_summary_df.head(daily_plot_days)
def compute_aggregated_results_summary(days) -> pd.DataFrame:
aggregated_result_summary_df = result_summary_df.copy()
aggregated_result_summary_df["covid_cases_for_ratio"] = \
aggregated_result_summary_df.covid_cases.mask(
aggregated_result_summary_df.shared_diagnoses == 0, 0)
aggregated_result_summary_df["covid_cases_for_ratio_es"] = \
aggregated_result_summary_df.covid_cases_es.mask(
aggregated_result_summary_df.shared_diagnoses_es == 0, 0)
aggregated_result_summary_df = aggregated_result_summary_df \
.sort_index(ascending=True).fillna(0).rolling(days).agg({
"covid_cases": "sum",
"covid_cases_es": "sum",
"covid_cases_for_ratio": "sum",
"covid_cases_for_ratio_es": "sum",
"shared_teks_by_generation_date": "sum",
"shared_teks_by_upload_date": "sum",
"shared_diagnoses": "sum",
"shared_diagnoses_es": "sum",
}).sort_index(ascending=False)
with pd.option_context("mode.use_inf_as_na", True):
aggregated_result_summary_df = aggregated_result_summary_df.fillna(0).astype(int)
aggregated_result_summary_df["teks_per_shared_diagnosis"] = \
(aggregated_result_summary_df.shared_teks_by_upload_date /
aggregated_result_summary_df.covid_cases_for_ratio).fillna(0)
aggregated_result_summary_df["shared_diagnoses_per_covid_case"] = \
(aggregated_result_summary_df.shared_diagnoses /
aggregated_result_summary_df.covid_cases_for_ratio).fillna(0)
aggregated_result_summary_df["shared_diagnoses_per_covid_case_es"] = \
(aggregated_result_summary_df.shared_diagnoses_es /
aggregated_result_summary_df.covid_cases_for_ratio_es).fillna(0)
return aggregated_result_summary_df
aggregated_result_with_7_days_window_summary_df = compute_aggregated_results_summary(days=7)
aggregated_result_with_7_days_window_summary_df.head()
last_7_days_summary = aggregated_result_with_7_days_window_summary_df.to_dict(orient="records")[1]
last_7_days_summary
aggregated_result_with_14_days_window_summary_df = compute_aggregated_results_summary(days=13)
last_14_days_summary = aggregated_result_with_14_days_window_summary_df.to_dict(orient="records")[1]
last_14_days_summary
```
## Report Results
```
display_column_name_mapping = {
"sample_date": "Sample\u00A0Date\u00A0(UTC)",
"source_regions": "Source Countries",
"datetime_utc": "Timestamp (UTC)",
"upload_date": "Upload Date (UTC)",
"generation_to_upload_days": "Generation to Upload Period in Days",
"region": "Backend",
"region_x": "Backend\u00A0(A)",
"region_y": "Backend\u00A0(B)",
"common_teks": "Common TEKs Shared Between Backends",
"common_teks_fraction": "Fraction of TEKs in Backend (A) Available in Backend (B)",
"covid_cases": "COVID-19 Cases (Source Countries)",
"shared_teks_by_generation_date": "Shared TEKs by Generation Date (Source Countries)",
"shared_teks_by_upload_date": "Shared TEKs by Upload Date (Source Countries)",
"shared_teks_uploaded_on_generation_date": "Shared TEKs Uploaded on Generation Date (Source Countries)",
"shared_diagnoses": "Shared Diagnoses (Source Countries – Estimation)",
"teks_per_shared_diagnosis": "TEKs Uploaded per Shared Diagnosis (Source Countries)",
"shared_diagnoses_per_covid_case": "Usage Ratio (Source Countries)",
"covid_cases_es": "COVID-19 Cases (Spain)",
"app_downloads_es": "App Downloads (Spain – Official)",
"shared_diagnoses_es": "Shared Diagnoses (Spain – Official)",
"shared_diagnoses_per_covid_case_es": "Usage Ratio (Spain)",
}
summary_columns = [
"covid_cases",
"shared_teks_by_generation_date",
"shared_teks_by_upload_date",
"shared_teks_uploaded_on_generation_date",
"shared_diagnoses",
"teks_per_shared_diagnosis",
"shared_diagnoses_per_covid_case",
"covid_cases_es",
"app_downloads_es",
"shared_diagnoses_es",
"shared_diagnoses_per_covid_case_es",
]
summary_percentage_columns= [
"shared_diagnoses_per_covid_case_es",
"shared_diagnoses_per_covid_case",
]
```
### Daily Summary Table
```
result_summary_df_ = result_summary_df.copy()
result_summary_df = result_summary_df[summary_columns]
result_summary_with_display_names_df = result_summary_df \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping)
result_summary_with_display_names_df
```
### Daily Summary Plots
```
result_plot_summary_df = result_summary_df.head(daily_plot_days)[summary_columns] \
.droplevel(level=["source_regions"]) \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping)
summary_ax_list = result_plot_summary_df.sort_index(ascending=True).plot.bar(
title=f"Daily Summary",
rot=45, subplots=True, figsize=(15, 30), legend=False)
ax_ = summary_ax_list[0]
ax_.get_figure().tight_layout()
ax_.get_figure().subplots_adjust(top=0.95)
_ = ax_.set_xticklabels(sorted(result_plot_summary_df.index.strftime("%Y-%m-%d").tolist()))
for percentage_column in summary_percentage_columns:
percentage_column_index = summary_columns.index(percentage_column)
summary_ax_list[percentage_column_index].yaxis \
.set_major_formatter(matplotlib.ticker.PercentFormatter(1.0))
```
### Daily Generation to Upload Period Table
```
display_generation_to_upload_period_pivot_df = \
generation_to_upload_period_pivot_df \
.head(backend_generation_days)
display_generation_to_upload_period_pivot_df \
.head(backend_generation_days) \
.rename_axis(columns=display_column_name_mapping) \
.rename_axis(index=display_column_name_mapping)
fig, generation_to_upload_period_pivot_table_ax = plt.subplots(
figsize=(12, 1 + 0.6 * len(display_generation_to_upload_period_pivot_df)))
generation_to_upload_period_pivot_table_ax.set_title(
"Shared TEKs Generation to Upload Period Table")
sns.heatmap(
data=display_generation_to_upload_period_pivot_df
.rename_axis(columns=display_column_name_mapping)
.rename_axis(index=display_column_name_mapping),
fmt=".0f",
annot=True,
ax=generation_to_upload_period_pivot_table_ax)
generation_to_upload_period_pivot_table_ax.get_figure().tight_layout()
```
### Hourly Summary Plots
```
hourly_summary_ax_list = hourly_summary_df \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.plot.bar(
title=f"Last 24h Summary",
rot=45, subplots=True, legend=False)
ax_ = hourly_summary_ax_list[-1]
ax_.get_figure().tight_layout()
ax_.get_figure().subplots_adjust(top=0.9)
_ = ax_.set_xticklabels(sorted(hourly_summary_df.index.strftime("%Y-%m-%d@%H").tolist()))
```
### Publish Results
```
github_repository = os.environ.get("GITHUB_REPOSITORY")
if github_repository is None:
github_repository = "pvieito/Radar-STATS"
github_project_base_url = "https://github.com/" + github_repository
display_formatters = {
display_column_name_mapping["teks_per_shared_diagnosis"]: lambda x: f"{x:.2f}" if x != 0 else "",
display_column_name_mapping["shared_diagnoses_per_covid_case"]: lambda x: f"{x:.2%}" if x != 0 else "",
display_column_name_mapping["shared_diagnoses_per_covid_case_es"]: lambda x: f"{x:.2%}" if x != 0 else "",
}
general_columns = \
list(filter(lambda x: x not in display_formatters, display_column_name_mapping.values()))
general_formatter = lambda x: f"{x}" if x != 0 else ""
display_formatters.update(dict(map(lambda x: (x, general_formatter), general_columns)))
daily_summary_table_html = result_summary_with_display_names_df \
.head(daily_plot_days) \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.to_html(formatters=display_formatters)
multi_backend_summary_table_html = multi_backend_summary_df \
.head(daily_plot_days) \
.rename_axis(columns=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.rename_axis(index=display_column_name_mapping) \
.to_html(formatters=display_formatters)
def format_multi_backend_cross_sharing_fraction(x):
if pd.isna(x):
return "-"
elif round(x * 100, 1) == 0:
return ""
else:
return f"{x:.1%}"
multi_backend_cross_sharing_summary_table_html = multi_backend_cross_sharing_summary_df \
.rename_axis(columns=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.rename_axis(index=display_column_name_mapping) \
.to_html(
classes="table-center",
formatters=display_formatters,
float_format=format_multi_backend_cross_sharing_fraction)
multi_backend_cross_sharing_summary_table_html = \
multi_backend_cross_sharing_summary_table_html \
.replace("<tr>","<tr style=\"text-align: center;\">")
extraction_date_result_summary_df = \
result_summary_df[result_summary_df.index.get_level_values("sample_date") == extraction_date]
extraction_date_result_hourly_summary_df = \
hourly_summary_df[hourly_summary_df.extraction_date_with_hour == extraction_date_with_hour]
covid_cases = \
extraction_date_result_summary_df.covid_cases.item()
shared_teks_by_generation_date = \
extraction_date_result_summary_df.shared_teks_by_generation_date.item()
shared_teks_by_upload_date = \
extraction_date_result_summary_df.shared_teks_by_upload_date.item()
shared_diagnoses = \
extraction_date_result_summary_df.shared_diagnoses.item()
teks_per_shared_diagnosis = \
extraction_date_result_summary_df.teks_per_shared_diagnosis.item()
shared_diagnoses_per_covid_case = \
extraction_date_result_summary_df.shared_diagnoses_per_covid_case.item()
shared_teks_by_upload_date_last_hour = \
extraction_date_result_hourly_summary_df.shared_teks_by_upload_date.sum().astype(int)
display_source_regions = ", ".join(report_source_regions)
if len(report_source_regions) == 1:
display_brief_source_regions = report_source_regions[0]
else:
display_brief_source_regions = f"{len(report_source_regions)} 🇪🇺"
def get_temporary_image_path() -> str:
return os.path.join(tempfile.gettempdir(), str(uuid.uuid4()) + ".png")
def save_temporary_plot_image(ax):
if isinstance(ax, np.ndarray):
ax = ax[0]
media_path = get_temporary_image_path()
ax.get_figure().savefig(media_path)
return media_path
def save_temporary_dataframe_image(df):
import dataframe_image as dfi
df = df.copy()
df_styler = df.style.format(display_formatters)
media_path = get_temporary_image_path()
dfi.export(df_styler, media_path)
return media_path
summary_plots_image_path = save_temporary_plot_image(
ax=summary_ax_list)
summary_table_image_path = save_temporary_dataframe_image(
df=result_summary_with_display_names_df)
hourly_summary_plots_image_path = save_temporary_plot_image(
ax=hourly_summary_ax_list)
multi_backend_summary_table_image_path = save_temporary_dataframe_image(
df=multi_backend_summary_df)
generation_to_upload_period_pivot_table_image_path = save_temporary_plot_image(
ax=generation_to_upload_period_pivot_table_ax)
```
### Save Results
```
report_resources_path_prefix = "Data/Resources/Current/RadarCOVID-Report-"
result_summary_df.to_csv(
report_resources_path_prefix + "Summary-Table.csv")
result_summary_df.to_html(
report_resources_path_prefix + "Summary-Table.html")
hourly_summary_df.to_csv(
report_resources_path_prefix + "Hourly-Summary-Table.csv")
multi_backend_summary_df.to_csv(
report_resources_path_prefix + "Multi-Backend-Summary-Table.csv")
multi_backend_cross_sharing_summary_df.to_csv(
report_resources_path_prefix + "Multi-Backend-Cross-Sharing-Summary-Table.csv")
generation_to_upload_period_pivot_df.to_csv(
report_resources_path_prefix + "Generation-Upload-Period-Table.csv")
_ = shutil.copyfile(
summary_plots_image_path,
report_resources_path_prefix + "Summary-Plots.png")
_ = shutil.copyfile(
summary_table_image_path,
report_resources_path_prefix + "Summary-Table.png")
_ = shutil.copyfile(
hourly_summary_plots_image_path,
report_resources_path_prefix + "Hourly-Summary-Plots.png")
_ = shutil.copyfile(
multi_backend_summary_table_image_path,
report_resources_path_prefix + "Multi-Backend-Summary-Table.png")
_ = shutil.copyfile(
generation_to_upload_period_pivot_table_image_path,
report_resources_path_prefix + "Generation-Upload-Period-Table.png")
```
### Publish Results as JSON
```
def generate_summary_api_results(df: pd.DataFrame) -> list:
api_df = df.reset_index().copy()
api_df["sample_date_string"] = \
api_df["sample_date"].dt.strftime("%Y-%m-%d")
api_df["source_regions"] = \
api_df["source_regions"].apply(lambda x: x.split(","))
return api_df.to_dict(orient="records")
summary_api_results = \
generate_summary_api_results(df=result_summary_df)
today_summary_api_results = \
generate_summary_api_results(df=extraction_date_result_summary_df)[0]
summary_results = dict(
backend_identifier=report_backend_identifier,
source_regions=report_source_regions,
extraction_datetime=extraction_datetime,
extraction_date=extraction_date,
extraction_date_with_hour=extraction_date_with_hour,
last_hour=dict(
shared_teks_by_upload_date=shared_teks_by_upload_date_last_hour,
shared_diagnoses=0,
),
today=today_summary_api_results,
last_7_days=last_7_days_summary,
last_14_days=last_14_days_summary,
daily_results=summary_api_results)
summary_results = \
json.loads(pd.Series([summary_results]).to_json(orient="records"))[0]
with open(report_resources_path_prefix + "Summary-Results.json", "w") as f:
json.dump(summary_results, f, indent=4)
```
### Publish on README
```
with open("Data/Templates/README.md", "r") as f:
readme_contents = f.read()
readme_contents = readme_contents.format(
extraction_date_with_hour=extraction_date_with_hour,
github_project_base_url=github_project_base_url,
daily_summary_table_html=daily_summary_table_html,
multi_backend_summary_table_html=multi_backend_summary_table_html,
multi_backend_cross_sharing_summary_table_html=multi_backend_cross_sharing_summary_table_html,
display_source_regions=display_source_regions)
with open("README.md", "w") as f:
f.write(readme_contents)
```
### Publish on Twitter
```
enable_share_to_twitter = os.environ.get("RADARCOVID_REPORT__ENABLE_PUBLISH_ON_TWITTER")
github_event_name = os.environ.get("GITHUB_EVENT_NAME")
if enable_share_to_twitter and github_event_name == "schedule" and \
(shared_teks_by_upload_date_last_hour or not are_today_results_partial):
import tweepy
twitter_api_auth_keys = os.environ["RADARCOVID_REPORT__TWITTER_API_AUTH_KEYS"]
twitter_api_auth_keys = twitter_api_auth_keys.split(":")
auth = tweepy.OAuthHandler(twitter_api_auth_keys[0], twitter_api_auth_keys[1])
auth.set_access_token(twitter_api_auth_keys[2], twitter_api_auth_keys[3])
api = tweepy.API(auth)
summary_plots_media = api.media_upload(summary_plots_image_path)
summary_table_media = api.media_upload(summary_table_image_path)
generation_to_upload_period_pivot_table_image_media = api.media_upload(generation_to_upload_period_pivot_table_image_path)
media_ids = [
summary_plots_media.media_id,
summary_table_media.media_id,
generation_to_upload_period_pivot_table_image_media.media_id,
]
if are_today_results_partial:
today_addendum = " (Partial)"
else:
today_addendum = ""
def format_shared_diagnoses_per_covid_case(value) -> str:
if value == 0:
return "–"
return f"≤{value:.2%}"
display_shared_diagnoses_per_covid_case = \
format_shared_diagnoses_per_covid_case(value=shared_diagnoses_per_covid_case)
display_last_14_days_shared_diagnoses_per_covid_case = \
format_shared_diagnoses_per_covid_case(value=last_14_days_summary["shared_diagnoses_per_covid_case"])
display_last_14_days_shared_diagnoses_per_covid_case_es = \
format_shared_diagnoses_per_covid_case(value=last_14_days_summary["shared_diagnoses_per_covid_case_es"])
status = textwrap.dedent(f"""
#RadarCOVID – {extraction_date_with_hour}
Today{today_addendum}:
- Uploaded TEKs: {shared_teks_by_upload_date:.0f} ({shared_teks_by_upload_date_last_hour:+d} last hour)
- Shared Diagnoses: ≤{shared_diagnoses:.0f}
- Usage Ratio: {display_shared_diagnoses_per_covid_case}
Last 14 Days:
- Usage Ratio (Estimation): {display_last_14_days_shared_diagnoses_per_covid_case}
- Usage Ratio (Official): {display_last_14_days_shared_diagnoses_per_covid_case_es}
Info: {github_project_base_url}#documentation
""")
status = status.encode(encoding="utf-8")
api.update_status(status=status, media_ids=media_ids)
```
| github_jupyter |
```
import collections
from datetime import datetime
import functools
import glob
import itertools
import os
import pathlib
import requests
import string
import sys
import typing
import zipfile
import IPython
import numpy as np
import pandas as pd
import pypandoc
from tqdm.notebook import tqdm_notebook
module_path = os.path.abspath(os.path.join("../.."))
if module_path not in sys.path:
sys.path.append(module_path)
from data_pipeline.utils import remove_all_from_dir, get_excel_column_name
from data_pipeline.etl.sources.census.etl_utils import get_state_information
# Turn on TQDM for pandas so that we can have progress bars when running `apply`.
tqdm_notebook.pandas()
# Suppress scientific notation in pandas (this shows up for census tract IDs)
pd.options.display.float_format = "{:.2f}".format
pd.set_option("max_columns", None)
# Set some global parameters
DATA_DIR = pathlib.Path.cwd().parent / "data"
TEMP_DATA_DIR = DATA_DIR / "tmp"
COMPARISON_OUTPUTS_DIR = DATA_DIR / "comparison_outputs"
## I (Vincent) created this manually locally. Will need to change potentially when putting into official ETL scripts
EJSCREEN_DATA_DIR = DATA_DIR / "ejscreen"
LOCAL_DATA_OUTPUT_DIR = DATA_DIR / "local"
EJSCREEN_CEQ_NAT_DIR = EJSCREEN_DATA_DIR / "CEQ_NationalExports"
EJSCREEN_CEQ_STA_DIR = EJSCREEN_DATA_DIR / "CEQ_StateExports"
# Make the dirs if they don't exist
TEMP_DATA_DIR.mkdir(parents=True, exist_ok=True)
COMPARISON_OUTPUTS_DIR.mkdir(parents=True, exist_ok=True)
# Name fields using variables. (This makes it easy to reference the same fields frequently without using strings
# and introducing the risk of misspelling the field name.)
GEOID_FIELD_NAME = "GEOID10"
GEOID_TRACT_FIELD_NAME = "GEOID10_TRACT"
GEOID_STATE_FIELD_NAME = "GEOID10_STATE"
COUNTRY_FIELD_NAME = "Country"
CENSUS_BLOCK_GROUP_POPULATION_FIELD = "Total population"
CEJST_SCORE_FIELD = "cejst_score"
CEJST_PERCENTILE_FIELD = "cejst_percentile"
CEJST_PRIORITY_COMMUNITY_FIELD = "cejst_priority_community"
# Define some suffixes
POPULATION_SUFFIX = " (priority population)"
```
## Loading EJ Screen CEQ Data
### National
```
# Replace this with something like glob when you have internet
filenames = [
"CEQ_EJSCREEN_National_70.csv",
"CEQ_EJSCREEN_National_75.csv",
"CEQ_EJSCREEN_National_80.csv",
"CEQ_EJSCREEN_National_85.csv",
"CEQ_EJSCREEN_National_90.csv",
"CEQ_EJSCREEN_National_95.csv",
]
dfs = []
for f in filenames:
percentile = f[-6:][:-4]
print(percentile)
df = pd.read_csv(
os.path.join(
EJSCREEN_CEQ_NAT_DIR,
"CEQ_EJSCREEN_National_{}.csv".format(percentile),
),
encoding="ISO-8859-1",
dtype="str",
)
df["EXCEED_COUNT"] = pd.to_numeric(df["EXCEED_COUNT"])
df.rename(columns={"ID": GEOID_FIELD_NAME}, inplace=True)
df["percentile"] = percentile
df = df[[GEOID_FIELD_NAME, "percentile", "EXCEED_COUNT"]]
dfs.append(df)
df = pd.concat(dfs)
df.head()
df_reshaped_nat = df.pivot(
index=GEOID_FIELD_NAME, columns="percentile", values="EXCEED_COUNT"
)
df_reshaped_nat.columns = [
"EJSCREEN Areas of Concern, National, {}th percentile".format(p)
for p in df_reshaped_nat.columns
]
df_reshaped_nat.fillna(0, inplace=True)
for c in df_reshaped_nat.columns:
df_reshaped_nat[c + " (communities)"] = (df_reshaped_nat[c] > 0) * 1
df_reshaped_nat.reset_index(inplace=True)
df_reshaped_nat.head()
df_reshaped_nat.describe()
pd.isnull(df_reshaped_nat).describe()
```
### State
```
# Replace this with something like glob when you have internet
filenames = [
"CEQ_EJSCREEN_State_70.csv",
"CEQ_EJSCREEN_State_75.csv",
"CEQ_EJSCREEN_State_80.csv",
"CEQ_EJSCREEN_State_85.csv",
"CEQ_EJSCREEN_State_90.csv",
"CEQ_EJSCREEN_State_95.csv",
]
dfs = []
for f in filenames:
percentile = f[-6:][:-4]
print(percentile)
df = pd.read_csv(
os.path.join(
EJSCREEN_CEQ_STA_DIR, "CEQ_EJSCREEN_State_{}.csv".format(percentile)
),
encoding="ISO-8859-1",
dtype="str",
)
df["EXCEED_COUNT"] = pd.to_numeric(df["EXCEED_COUNT"])
df.rename(columns={"ID": GEOID_FIELD_NAME}, inplace=True)
df["percentile"] = percentile
df = df[[GEOID_FIELD_NAME, "percentile", "EXCEED_COUNT"]]
dfs.append(df)
df = pd.concat(dfs)
df.head()
df_reshaped_sta = df.pivot(
index=GEOID_FIELD_NAME, columns="percentile", values="EXCEED_COUNT"
)
df_reshaped_sta.columns = [
"EJSCREEN Areas of Concern, State, {}th percentile".format(p)
for p in df_reshaped_sta.columns
]
df_reshaped_sta.fillna(0, inplace=True)
for c in df_reshaped_sta.columns:
df_reshaped_sta[c + " (communities)"] = (df_reshaped_sta[c] > 0) * 1
df_reshaped_sta.reset_index(inplace=True)
df_reshaped_sta.head()
df_reshaped_nat.describe()
pd.isnull(df_reshaped_sta).describe()
df_reshaped = df_reshaped_nat.merge(df_reshaped_sta, on=GEOID_FIELD_NAME)
df_reshaped.head()
df_reshaped.to_csv(
path_or_buf=LOCAL_DATA_OUTPUT_DIR
/ "ejscreen_areas_of_concerns_indicators.csv",
na_rep="",
index=False,
)
```
# Next Steps / Questions
Lucas, here's what the output file looks like. For each CBG I have new columns corresponding to the different percentiles for both State and National. For each percentile there are two columns: one for the number of `EXCEED_COUNT` and a boolean indicator for whether `EXCEED_COUNT > 0` for that percentile. I think that's what we wanted right?
1. Do we have a list of all CBGs? The reason for asking is I created a CSV that lists each CBG and the number of EJSCREEN Areas of Concerns for each percentile. It's not going to have all the CBGs in them since if the CBG doesn't have an area concern at least at the 70th percentile, then the CBG wouldn't have appeared in the source data set. Do we want to make sure to add all the remaining CBGs with 0's across the board?
1. Definitely need to clean up the code, at least not make it so duplicatous across national and state
| github_jupyter |
# Using titrationFitter
This notebook should guide you through the basic steps of the fitting of your titration.
## What is titrationFitter
titrationFitter is a very small python library designed to help you fit titration data. Because it is so small, it is also more flexible but it incumbs to the user to handle the inputs of their data.
## 1. Preparing your data
You will need to prepare 4 text files:
- one containing the equivalents for each step of the titration
- another containing the cumulative volume of your titration
- the actual spectra (e.g. UV-VIS)
- an optional file containing labels for each wavelengths of the spectrum.
You can use a spreadsheet program to prepare the said files, and then save them to CSVs, preferably using the space character as the delimiter.
The UV-VIS spectrum file will look like this in the spreadsheet:
<img src="images_notebook/spreadsheet.png" style="width: 40em;"/>
The convention is **one column = one entire spectrum**, thus you will have as many columns as you have equivalent points. Once the file is saved to a CSV format, we have a text file looking like this:
<img src="images_notebook/textfile.png" style="width: 40em;"/>
The same format should be kept for the other files, except you will have only row or one column for each file since the rest of the information is 1D. See the files in the subdirectory _data_.
## 2. Loading your data
Now, we need to load our data onto the memory. We assume again that the files are in the subdirectory _data_. We will use the python library _numpy_ to take care of this task for us:
```
import numpy as np
# the subdirectory where our files are stored:
folder = 'data/'
# ligand/metal ratios at the different points of the titration:
eqs = np.loadtxt(folder+'eqs.csv')
# span of the UV-VIS spectrum:
span = np.loadtxt(folder+'span.csv')
# values of the UV-VIS spectrum: (contained by columns in the file)
uvs = np.loadtxt(folder+'uvs.csv')
# and the volumes of our titration at each step:
volumes = np.loadtxt(folder+'volumes.csv')
```
## 3. (Optional) undersample our data
Often, spectrometers oversample the information by measuring (e.g. the transmission) at a large number of wavelengths. We can usually keep say 1 in 3 and still be fine. This will make the optimization much faster.
Once again, we use the power of numpy to keep only selected points from our total spectra:
```
# start at wavelength number 30, end at wavelength number 120
# and within these bounds, keep only one point in 2:
uvs = uvs[30:120:2,:]
# same for the labels:
span = span[30:120:2]
```
## 4. Preparing our model
Finally we start using the library. We import its components:
```
from titrationFitter import Component, System, Titration
```
We need to think about our model in advance and give the program a reasonable initial guess for the binding constants and the UV spectra of the components. Below I give such a guess for the system of this example, but keep in mind:
__From this point, the procedure is iterative__.
You probably won't get it right the first time.
Let us define the building blocks of our titration: a metal, and a ligand:
```
# We are titrating with a colorless metal. Its uv-vis spectrum is then zero.
M = Component(name='M', titrant=True,
uv_known=True, uvvis=np.zeros_like(span))
# The compound being titrated is a ligand. Its UV-VIS is known:
# it is that of the very beginning of the titration, normalised
# by the concentration.
initial_concentration = 0.1
L = Component(name='L', conc_ini=initial_concentration,
uv_known=True, uvvis=uvs[:,0]/initial_concentration)
```
Then, we need to define the species that we suspect are being formed. This time, we will need not only to provide a guess for the spectra, but also for the binding constant.
Note aside, the initial guesses regarding the spectra are educated: they were infered from the conformations of the ligands as they bind to the metal.
```
# then we tell the program what composes "ML": one M and one L.
# the UV spectrum is that of the very end of the titration.
# Since it is not known, it will be optimized by the program.
ML = Component(eqconst=10**5, conc_ini=0.,
buildblocks=[M,L], coeffs=[1,1],
uv_known=False, uvvis=(uvs[:,-1])/initial_concentration/volumes[0]*volumes[-1],
eqconst_known=False)
# same as above, except we have 1 M and 2 L.
ML2 = Component(eqconst=10**8, conc_ini=0.,
buildblocks=[M,L], coeffs=[1,2],
uv_known=False, uvvis=(2*uvs[:,-1])/initial_concentration,
eqconst_known=False)
# same again, but with 1 M and 3 L.
ML3 = Component(eqconst=10**10., conc_ini=0.,
buildblocks=[M,L], coeffs=[1,3],
uv_known=False, uvvis=(uvs[:,-1]+2*uvs[:,0])/initial_concentration,
eqconst_known=False)
# Finally we put everything together in a "system":
# the building blocks, the species, the initial concentration, and the labels.
S = System([M, L], [ML, ML2, ML3],
conc_ini=initial_concentration, span=span)
# When we add the information about the steps (volume, equivalence),
# we obtain a titration:
T = Titration(S, eqs[:15], uvs[:,:15], volumes=volumes[:15])
```
I think the above is pretty general, such that you can use it as a template for any titration that you might want to fit.
## 5. Optimization
Now the optimization step:
(might take a while)
```
T.optimize()
```
Now we can plot and evaluate our model (drag the bar in the top left image to see the spectrum at different points of the titration)
```
# just some magic command to make the plot appear in this notebook:
%matplotlib notebook
# the command triggering the plot of our titration:
T.plotCurrentModel()
```
Get the final coupling constant:
```
T.printCurrentModel()
```
## 6. What are the errors on the determined coupling coefficients?
This is not implemented yet. The only method I can think of to reliably estimate a confidence interval would be a bootstrap. But that would be very expensive computationally.
| github_jupyter |
# Tensorflow Timeline Analysis on Model Zoo Benchmark among different data types
This jupyter notebook will help you evaluate performance benefits among different data types on the level of Tensorflow operations via several models from Intel Model Zoo. The notebook will show users a bar chart like the picture below for the Tensorflow operation level performance comparison. The red horizontal line represents the performance of Tensorflow operations from one data type as performance baseline, and the blue bars represent the speedup of Tensorflow operations by using other data type with oneDNN. The orange bars represent the speedup of Tensorflow operations by using other data type without oneDNN. Users should be able to see a good speedup for those operations accelerated by Intel DL Boost instructions.
> NOTE : Users need to get Tensorflow timeline json files from other Jupyter notebooks like benchmark_date_types_perf_comparison
first to proceed this Jupyter notebook.
<img src="images\compared_tf_op_duration_ratio_bar_types.png" width="700">
# Get Platform Information
```
# ignore all warning messages
import warnings
warnings.filterwarnings('ignore')
from profiling.profile_utils import PlatformUtils
plat_utils = PlatformUtils()
plat_utils.dump_platform_info()
```
# Section 1 : Prerequisites
```
!pip install cxxfilt
%matplotlib inline
import matplotlib.pyplot as plt
import tensorflow as tf
import pandas as pd
pd.set_option('display.max_rows', 500)
pd.set_option('display.max_columns', 500)
pd.set_option('display.width', 1500)
```
## List out the Timeline folders
First, list out all Timeline folders from previous runs.
```
import os
filenames= os.listdir (".")
result = []
keyword = "Timeline"
for filename in filenames:
if os.path.isdir(os.path.join(os.path.abspath("."), filename)):
if filename.find(keyword) != -1:
result.append(filename)
result.sort()
index =0
for folder in result:
print(" %d : %s " %(index, folder))
index+=1
```
## Select a Timeline folder from previous runs
#### ACTION: Please select one Timeline folder and change FdIndex accordingly
```
FdIndex = 0
```
List out all Timeline json files inside Timeline folder.
```
import os
TimelineFd = result[FdIndex]
print(TimelineFd)
datafiles = [TimelineFd +os.sep+ x for x in os.listdir(TimelineFd) if '.json' == x[-5:]]
print(datafiles)
if len(datafiles) is 0:
print("ERROR! No json file in the selected folder. Please select other folder.")
elif len(datafiles) is 1:
print("WARNING! There is only 1 json file in the selected folder. Please select other folder to proceed Section 1.2.")
```
<a id='section_2'></a>
# Section 2: Performance analysis between two different data types
## Section 2.1 : Analyze TF Timeline results
### Step 1: Select two TF timeline files with different data types for analysis
#### List out all timeline files in the selected folder
```
if len(datafiles) is 1:
print("ERROR! There is only 1 json file in the selected folder.")
print("Please select other Timeline folder from beginnning to proceed Section 1.2.")
for i in range(len(datafiles)):
print(" %d : %s " %(i, datafiles[i]))
```
#### ACTION: Please select one timeline file as a perfomance baseline and the other as a comparison target
Please put the related index for your selected timeline file.
In general, fp32 data type should be the performance baseline.
```
# perfomance baseline
Baseline_Index=0
# comparison target
Comparison_Index=1
```
#### List out two selected timeline files
```
selected_datafiles = []
selected_datafiles.append(datafiles[Baseline_Index])
selected_datafiles.append(datafiles[Comparison_Index])
print(selected_datafiles)
```
#### Find the related oneDNN JITDUMP folder and Tensorflow Log file with oneDNN Verbose log
```
from profiling.profile_utils import PerfPresenter
perfp=PerfPresenter()
tag0, tag1 = perfp.get_diff_from_csv_filenames(selected_datafiles[0],selected_datafiles[1])
tags = []
jitdumps_fd_path = []
logs_path = []
for tag in [tag0, tag1]:
# TEMP FIX
if tag == 'bf16':
tag = 'bfloat16'
if tag == 'f32':
tag = 'fp32'
JITDUMP_FD_PATH= TimelineFd + os.sep +'JITDUMP_'+ tag
if os.path.isdir(os.path.join(os.path.abspath("."), JITDUMP_FD_PATH)):
logs = [JITDUMP_FD_PATH +os.sep+ x for x in os.listdir(JITDUMP_FD_PATH) if '.log.old' == x[-8:]]
LOG_PATH = logs[0]
jitdumps_fd_path.append(JITDUMP_FD_PATH)
logs_path.append(LOG_PATH)
tags.append(tag)
print(tags)
print(jitdumps_fd_path)
print(logs_path)
```
### Step 2: Parsing timeline results into CSV files
```
%matplotlib agg
from profiling.profile_utils import TFTimelinePresenter
csvfiles=[]
tfp = TFTimelinePresenter(True)
for fn in selected_datafiles:
if fn.find('/'):
fn_nofd=fn.split('/')[1]
else:
fn_nofd=fn
tfile_name= fn_nofd.split('.')[0]
tfile_prefix = fn_nofd.split('_')[0]
tfile_postfix = fn_nofd.strip(tfile_prefix)[1:]
csvpath = TimelineFd +os.sep+tfile_name+'.csv'
print(csvpath)
csvfiles.append(csvpath)
timeline_pd = tfp.postprocess_timeline(tfp.read_timeline(fn))
timeline_pd = timeline_pd[timeline_pd['ph'] == 'X']
tfp.get_tf_ops_time(timeline_pd,fn,tfile_prefix)
```
### Step 3: Pre-processing for the two CSV files
```
import os
import pandas as pd
csvarray=[]
for csvf in csvfiles:
print("read into pandas :",csvf)
a = pd.read_csv(csvf)
csvarray.append(a)
a = csvarray[0]
b = csvarray[1]
# Find tags among CSV files
tags=[]
from profiling.profile_utils import PerfPresenter
perfp=PerfPresenter()
tag0, tag1 = perfp.get_diff_from_csv_filenames(csvfiles[0],csvfiles[1])
tags = [tag0, tag1]
print('tags : ',tags)
```
### Step 4: Merge two CSV files and caculate the speedup accordingly
Users can check column "speedup" for the speedup from bfloat16 or int8.
If the operation uses mkl-dnn/oneDNN for acceleration, the "mkl_op" column should be marked as "True".
```
import os
import pandas as pd
fdir='merged'
if not os.path.exists(fdir):
os.mkdir(fdir)
fpaths=[]
fpaths.append(fdir+os.sep+'merged.csv')
fpaths.append(fdir+os.sep+'diff_'+tags[0]+'.csv')
fpaths.append(fdir+os.sep+'diff_'+tags[1]+'.csv')
merged=tfp.merge_two_csv_files_v2(fpaths, a, b, tags)
merged
```
#### The unique Tensorflow operations from the first csv/Timline file
```
%matplotlib inline
print("Operations are only in", tags[0], " run")
extra1 = pd.read_csv(fpaths[1])
extra1
```
#### The unique Tensorflow operations from the second csv/Timline file
```
print("Operations are only in", tags[1], " run")
extra2 = pd.read_csv(fpaths[2])
extra2
```
### Step 5: Draw a bar chart for elapsed time of TF ops among two different data types
The first diagram compares the elapsed time of operations among two different data types.
The second diagram shows the speedup of TF operations from comparison target.
The blue bar of second diagram is accelerated by oneDNN operation.
```
%matplotlib inline
print(fpaths[0])
tfp.plot_compare_bar_charts(fpaths[0], tags=tags)
tfp.plot_compare_ratio_bar_charts(fpaths[0], tags=['','oneDNN ops'])
```
### Step 6: Draw pie charts for elapsed time of TF ops among different data types
Users should be able to identify top hotspots from below pie charts among different data types.
> NOTE: Users could also compare elapsed time of TF ops among any two different TF timeline files.
We will have following pie charts in sequence:
1. the pie chart for elpased time of TF ops from stock TF or the first csv/Timeline file
2. the pie chart for elpased time of unique TF ops from stock TF or the first csv/Timeline file
3. the pie chart for elpased time of TF ops from Intel TF or the second csv/Timeline file
4. the pie chart for elpased time of unique TF ops from Intel TF or the second csv/Timeline file
5. the pie chart for elpased time of common TF ops among stock & Intel TF or two csv/Timeline files
#### The pie chart for elapsed time of TF ops from the first csv/Timline file
understand which TF operations spend most of time.
```
tfp.plot_pie_chart(csvfiles[0], tags[0])
```
#### The pie chart for elapsed time of unique TF operations from the first csv/Timline file
understand if there is any unique TF operation.
```
tfp.plot_pie_chart(fpaths[1], tags[0])
```
#### The pie chart for elapsed time of TF ops from the second csv/Timline file
understand which TF operations spend most of time.
```
tfp.plot_pie_chart(csvfiles[1], tags[1])
```
#### The pie chart for elapsed time of unique TF operations from the second csv/Timline file
understand if there is any unique TF operation.
```
tfp.plot_pie_chart(fpaths[2], tags[1])
```
#### The pie chart for elapsed time of common TF ops among two csv/Timline files
understand top hotspots differences among two csv/Timeline files.
```
tfp.plot_compare_pie_charts(fpaths[0], tags=tags)
```
<a id='section_2_2'></a>
## Section 2.2: Analyze oneDNN/MKLDNN debug logs and JIT dumps
>NOTE: Section 2.2 is only relevant if user had MKLDNN_VERBOSE or MKLDNN_JIT_DUMP enabled
### Step 1: Parse related oneDNN Verbose logs
>NOTE: Step 1-3 is only relevant if user had MKLDNN_VERBOSE enabled.
```
from profiling.profile_utils import oneDNNUtils, oneDNNLog
onednn = oneDNNUtils()
log1 = oneDNNLog()
log1.load_log(logs_path[0])
exec_data1 = log1.exec_data
log2 = oneDNNLog()
log2.load_log(logs_path[1])
exec_data2 = log2.exec_data
print(logs_path)
```
### Step 2: Primitives Type Speedup from comparison target ( bfloat16 or int8)
The digram below shows performance speedup from the comparison target data type such as VNNI, or BF16.
```
onednn.stats_comp('type', 'time',log2, log1, tags=tags)
```
### Step 3: Time breakdown for JIT kernel type
oneDNN uses just-in-time compilation (JIT) to generate optimal code for some functions based on input parameters and instruction set supported by the system.
Therefore, users can see different JIT kernel type among different CPU and GPU architectures.
For example, users can see avx_core_vnni JIT kernel if the workload uses VNNI instruction on Cascake Lake platform.
Users can also see different OCL kernels among different Intel GPU generations.
Moreover, users can identify the top hotspots of JIT kernel executions with this time breakdown.
Time breakdown of Baseline Data Type for JIT kernel type
```
print("Time Breakdown of %s data type" %(tags[0]))
onednn.breakdown(exec_data1,"jit","time")
onednn.breakdown(exec_data1,"type","time")
```
Time breakdown of Comparison Data Type for JIT kernel type
```
print("Time Breakdown of %s data type" %(tags[1]))
onednn.breakdown(exec_data2,"jit","time")
onednn.breakdown(exec_data2,"type","time")
```
### Step 4: Inspect JIT Kernel
>NOTE: Step 4 is only relevant if user had MKLDNN_JIT_DUMP enabled
In this section, we analyze dump JIT files of the comparison data type.
Users should be able to see the exact CPU instruction usage like VNNI or BF16 from those JIT Dump files.
```
print('will inspect JIT codes from %s data type in %s '%(tags[1], jitdumps_fd_path[1]))
```
##### List out all JIT Dump Files with index number
```
import os
filenames= os.listdir (jitdumps_fd_path[1])
result = []
keyword = ".bin"
for filename in filenames:
#if os.path.isdir(os.path.join(os.path.abspath("."), filename)):
if filename.find(keyword) != -1:
result.append(filename)
result.sort()
index =0
for folder in result:
print(" %d : %s " %(index, folder))
index+=1
```
##### ACTION : Pick a JIT Dump file by putting its index value below
```
## USER INPUT
FdIndex=0
```
##### export JIT Dump file to environment variable JITFILE and also related ISA keyword to environment variable DNNL_ISA_KEYWORD
```
if FdIndex < len(result):
logfile = result[FdIndex]
os.environ["JITFILE"] = jitdumps_fd_path[1]+os.sep+logfile
if tags[1] == 'f32':
os.environ["DNNL_ISA_KEYWORD"] = "zmm"
elif tags[1] == 'int8':
os.environ["DNNL_ISA_KEYWORD"] = "vpdpbusd"
elif tags[1] == 'bf16':
os.environ["DNNL_ISA_KEYWORD"] ='vdpbf16ps|vcvtne2ps2bf16'
print(os.environ["JITFILE"])
print(os.environ["DNNL_ISA_KEYWORD"])
```
#### disassembler JIT Dump file
> NOTE: zmm register is introduced by AVX512 ISA.
Users should see usage of **zmm** register in AVX512 JIT dump files.
> NOTE: vpdpbusd is introduced by AVX512_VNNI ISA.
Users should see usage of **vpdpbusd** in AVX512_VNNI JIT dump files.
> NOTE: **vdpbf16ps**, **vcvtne2ps2bf16**, and **vcvtneps2bf16** are introduced by AVX512_BF16 ISA.
Users should see usage of vdpbf16ps, vcvtne2ps2bf16 or vcvtneps2bf16 in AVX512_BF16 JIT dump files.
> NOTE: For disassembler vdpbf16ps, vcvtne2ps2bf16, and vcvtneps2bf16 instructions, users must use objdump with **v2.34** or above.
```
!objdump -D -b binary -mi386:x86-64 $JITFILE | grep -E $DNNL_ISA_KEYWORD
```
### (Optional)Step 5: move all results files into the selected Timeline folder
By runing below codes, all results files will be moved to the selected Timeline folder
```
from profiling.profile_utils import CommonUtils
utils = CommonUtils()
import os
import shutil
import datetime
# move png and csv results into Timeline folder
pattern_list = [ "*.png" , "*.csv"]
current_path = os.getcwd()
for pattern in pattern_list:
png_fds, png_fd_paths = utils.found_files_in_folder(pattern, current_path)
for fd_path in png_fd_paths:
shutil.move(fd_path,TimelineFd)
# move pretrained model, logs, merged files into Timeline folder
fd_name_list = ['pretrained','merged','logs']
timeinfo = datetime.datetime.now().strftime("%Y-%m-%d_%H:%M")
for fd_name in fd_name_list:
if os.path.isdir(fd_name) == True:
target_fd = TimelineFd + os.sep+ fd_name+'_'+timeinfo
shutil.move(fd_name,target_fd)
```
| github_jupyter |
# Stateful Model Feedback Metrics Server
In this example we will add statistical performance metrics capabilities by levering the Seldon metrics server.
Dependencies
* Seldon Core installed
* Ingress provider (Istio or Ambassador)
An easy way is to run `examples/centralized-logging/full-kind-setup.sh` and then:
```bash
helm delete seldon-core-loadtesting
helm delete seldon-single-model
```
Then port-forward to that ingress on localhost:8003 in a separate terminal either with:
Ambassador:
kubectl port-forward $(kubectl get pods -n seldon -l app.kubernetes.io/name=ambassador -o jsonpath='{.items[0].metadata.name}') -n seldon 8003:8080
Istio:
kubectl port-forward -n istio-system svc/istio-ingressgateway 8003:80
```
!kubectl create namespace seldon || echo "namespace already created"
!kubectl config set-context $(kubectl config current-context) --namespace=seldon
!mkdir -p config
```
### Create a simple model
We create a multiclass classification model - iris classifier.
The iris classifier takes an input array, and returns the prediction of the 4 classes.
The prediction can be done as numeric or as a probability array.
```
%%bash
kubectl apply -f - << END
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: multiclass-model
spec:
predictors:
- graph:
children: []
implementation: SKLEARN_SERVER
modelUri: gs://seldon-models/sklearn/iris
name: classifier
logger:
url: http://seldon-multiclass-model-metrics.seldon.svc.cluster.local:80/
mode: all
name: default
replicas: 1
END
!kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=multiclass-model -o jsonpath='{.items[0].metadata.name}')
```
#### Send test request
```
res=!curl -X POST "http://localhost:8003/seldon/seldon/multiclass-model/api/v1.0/predictions" \
-H "Content-Type: application/json" -d '{"data": { "ndarray": [[1,2,3,4]]}, "meta": { "puid": "hello" }}'
print(res)
import json
j=json.loads(res[-1])
assert(len(j["data"]["ndarray"][0])==3)
```
### Metrics Server
You can create a kubernetes deployment of the metrics server with this:
```
%%writefile config/multiclass-deployment.yaml
apiVersion: apps/v1
kind: Deployment
metadata:
name: seldon-multiclass-model-metrics
namespace: seldon
labels:
app: seldon-multiclass-model-metrics
spec:
replicas: 1
selector:
matchLabels:
app: seldon-multiclass-model-metrics
template:
metadata:
labels:
app: seldon-multiclass-model-metrics
spec:
securityContext:
runAsUser: 8888
containers:
- name: user-container
image: seldonio/alibi-detect-server:1.7.0-dev
imagePullPolicy: IfNotPresent
args:
- --model_name
- multiclassserver
- --http_port
- '8080'
- --protocol
- seldonfeedback.http
- --storage_uri
- "adserver.cm_models.multiclass_one_hot.MulticlassOneHot"
- --reply_url
- http://message-dumper.default
- --event_type
- io.seldon.serving.feedback.metrics
- --event_source
- io.seldon.serving.feedback
- MetricsServer
env:
- name: "SELDON_DEPLOYMENT_ID"
value: "multiclass-model"
- name: "PREDICTIVE_UNIT_ID"
value: "classifier"
- name: "PREDICTIVE_UNIT_IMAGE"
value: "alibi-detect-server:1.7.0-dev"
- name: "PREDICTOR_ID"
value: "default"
---
apiVersion: v1
kind: Service
metadata:
name: seldon-multiclass-model-metrics
namespace: seldon
labels:
app: seldon-multiclass-model-metrics
spec:
selector:
app: seldon-multiclass-model-metrics
ports:
- protocol: TCP
port: 80
targetPort: 8080
!kubectl apply -n seldon -f config/multiclass-deployment.yaml
!kubectl rollout status deploy/seldon-multiclass-model-metrics
import time
time.sleep(20)
```
### Send feedback
```
res=!curl -X POST "http://localhost:8003/seldon/seldon/multiclass-model/api/v1.0/feedback" \
-H "Content-Type: application/json" \
-d '{"response": {"data": {"ndarray": [[0.0006985194531162841,0.003668039039435755,0.9956334415074478]]}}, "truth":{"data": {"ndarray": [[0,0,1]]}}}'
print(res)
import json
j=json.loads(res[-1])
assert("data" in j)
import time
time.sleep(3)
```
### Check that metrics are recorded
```
res=!kubectl logs $(kubectl get pods -l app=seldon-multiclass-model-metrics \
-n seldon -o jsonpath='{.items[0].metadata.name}') | grep "PROCESSING Feedback Event"
print(res)
assert(len(res)>0)
```
### Cleanup
```
!kubectl delete -n seldon -f config/multiclass-deployment.yaml
!kubectl delete sdep multiclass-model
```
| github_jupyter |
# Stretch PIMU Guide: Interpreting Sensor Data
The Stretch Body Pimu package provides a Python API to the Stretch RE1 Power and IMU Board (PIMU). In this guide, we'll look at using the stretch_pimu_jog.py and stretch_pimu_scope.py files to get a idea about the status of base.
## Setup
Stretch Body is available on PyPi as [hello-robot-stretch-body](https://pypi.org/project/hello-robot-stretch-body/). An accompanying command-line tools package called [hello-robot-stretch-body-tools](https://pypi.org/project/hello-robot-stretch-body-tools/) uses Stretch Body to provide convenient scripts to interact with the robot from the command-line. Both come preinstalled on Stretch RE1, but the following command can be used to ensure you have the latest version.
In Jupyter notebooks, code preceded by `!` are run in the command-line instead of the Python interpreter.
```
!python -m pip install -q -U hello-robot-stretch-body
!python -m pip list | grep hello-robot-stretch-body
```
# Stretch Body PIMU
The Stretch body PIMU jog file is available at stretch_body/tools/bin/stretch_pimu_jog.py, this file provides an easy way to interact with the Base board. This can be helpful to ping any componenet in the base board.
```
from stretch_body.pimu import Pimu
import time
```
Now, we'll instantiate an object of Pimu and call it `p`.
```
p = Pimu()
```
The cell below displays the docstring for `startup()`. As you can see, the method returns a boolean depending on whether or not the class startup procedure succeeded. Only one instance of the pimu class can exist at once, so if another instance is running elsewhere, the method returns false.
```
p.startup?
```
Below, we make the call to `startup()`. As part of the startup procedure, this method opens serial ports to the hardware devices, loads the parameters that dictate robot behavior, and launches a few helper threads to poll for status in the background. If it successfully does that then it should return True.
```
p.startup()
```
If startup fails unexpectedly, the first thing to check is whether a background process is already running an instance of the robot class. Below we use the `pstree` command to list the tree of background processes, and `grep` to filter for scripts starting with "stretch_" (often the "stretch_xbox_controller_teleop.py" scripts is running in the background). If we see output below, we should use the `pkill` command to [terminate the conflicting process](https://docs.hello-robot.com/troubleshooting_guide/#rpc-transport-errors-stretch-doesnt-respond-to-commands).
```
!pstree | grep stretch_
```
The second method we'll look at is called `stop()`. This method closes connections to all serial ports, releases other resources, and shuts down background threads. We'll wait until the end of the notebook to actually call the method.
```
p.stop?
```
The next method we'll look at is called `pretty_print()`. This method prints out the entire state of the PIMU in a human readable format.
```
p.pretty_print()
```
we'd often like to be able to access the robot's state programmatically. The pull_status() method returns a dictionary with a snapshot of the current state of the PIMU. This can be accessed through the status dictionary and specifying the keyword. Here for example the status of fan is displayed.
```
p.pull_status()
p.status['fan_on']
```
# User/Utlity Functions
Next, let's take a look at how we can use the User and Utility Functions in the Pimu class to access PIMU board.
These functions are mostly designed to be used during normal operation of stretch body or during development/factory use.
Note: All the functions require to be followed by the `push_command()`. The functions add the query to the Python API queue and push command is required to push the queue to hardware drivers.
IMU RESET: Toggles the reset line to the IMU chip. This potentially could be useful in long running robots is the IMU gets stuck in an errro state.
```
p.imu_reset()
p.push_command()
```
TRIGGER MOTOR SYNC: The motor sync ensures that the motions start at exactly the right time. Otherwise there can be a slight delay (~10ms) between the start of motion for each joint. If motor sync is enabled (via YAML) for the four stepper motors, using `move_to` and `push_command` gets the motion command down to the four stepper motors. But they won't start moving until `trigger_motor_sync` is called.
```
p.trigger_motor_sync()
```
SET_FAN_ON and SET_FAN_OFF: This function is used to set the fan in the PIMU board to on or off. We can use the p.pull_status to fetch the status and use p.status['keyword_name'] to display the status. For Fan, we can see if the fan is currently running or not and toggle it to on or off. Stretch body uses Intel-NUC and the fan is useful to prevent issues resulting from overheating.
```
p.pull_status()
if p.status['fan_on']:
p.set_fan_off()
else:
p.set_fan_on()
p.push_command()
```
SET_BUZZER_ON and SET_BUZZER_OFF: Same as in fan, these functions are used to trigger the buzzer on or off. The status values can be used to plot if
```
if p.status['buzzer_on']:
p.set_buzzer_off()
else:
p.set_buzzer_on()
p.push_command()
```
BOARD_RESET: board_reset will cause the SAMD microcontroller on the board to reset (as if the board was power cycled). Users generally don't need to call this.
```
p.board_reset()
p.push_command()
```
CLIFF_EVENT_RESET: The cliff sensors can be configured via YAML to put the robot in pause/runstop mode when they are outside of a threshold value. This will cause the robot to stop when it approaches a cliff. In order to reset this event and allow motion to resume this function must be called. By default we have this functionality turned off at the factory as it isn't well tested.
```
p.cliff_event_reset()
p.push_command()
```
RUNSTOP_EVENT_RESET: Reset the robot runstop, allowing motion to continue.
```
p.runstop_event_reset()
p.push_command()
```
RUNSTOP_EVENT_TRIGGER: Trigger the robot runstop, stopping motion.
```
p.runstop_event_trigger()
p.push_command()
```
TRIGGER_BEEP: You should hear a beep after running this command. This is useful sometimes for generating a sounds wave that can be helped to check if the bump sensors are working as expected. Generate a single short beep.
```
p.trigger_beep()
p.push_command()
```
# Visualising the status values.
It might be useful to actually visualise the continous status values of the PIMU. This involves pulling the status values from various sensors continously and plotting it on a graph. This method could be super helpful to understand how changes in the environment change the sensor readings and robot's status.
Let's install the required libraries for visualisation
```
import numpy as np
from IPython.display import display, clear_output
import matplotlib.pyplot as plt
def get_reading(p, params, functions=None):
p.pull_status()
value = p.status[params[0]]
for key in params[1:]:
value = value[key]
if functions != None:
for function in functions:
value = function(value)
return value
def plot_values(num_points, run_time, p, params, y_range=None, functions=None):
fig = plt.figure()
ax = fig.add_subplot(1, 1, 1)
new_val = get_reading(p, params, functions)
data = np.full(num_points, new_val)
for _ in range(run_time*2):
new_val = get_reading(p, params, functions)
print("new val is ", new_val)
data=np.roll(data,-1)
data[-1]=new_val
ax.cla()
ax.plot(data)
if y_range == None:
ax.set_ylim([new_val-0.1,new_val+0.1])
else:
ax.set_ylim(y_range)
display(fig)
clear_output(wait = True)
plt.pause(0.5)
def plot_values_4(num_points, run_time, p, params, yrange = None, functions=None):
fig = plt.figure()
ax = fig.add_subplot(1, 1, 1)
new_val = get_reading(p, params, functions)
print(new_val)
data = np.full([num_points,4], new_val)
for _ in range(run_time*2):
new_val = get_reading(p, params, functions)
print("new val is ", new_val)
data=np.roll(data,-1,0)
data[-1]=new_val
ax.cla()
ax.plot(data)
ax.set_ylim(yrange)
display(fig)
clear_output(wait = True)
plt.pause(0.5)
```
The value `num_points` signifies the number of values that will be plotted at a point in the line plot. `run_time` signifies the time in seconds till which the visualisation will run. New status value is pulled twise every second. `yrange` signifies the range of y-axis. If this is set to None, the visualisation will set the range as +/- 0.1 of current value. The `params` argument expects a list of the key values to access status. This is used for accessing specific sensor readings within certain devices. The `functions` argument expects a list of functions that could be applied to the values before visualising.
cliff: Scope base cliff sensors. The cliff sensors are helpful in indicating if the robot is at the edge of a fall or inclined. Try to lighly lift the robot from one side and notice how the sensor value spikes (be careful to not lift is too high as to topple Stretch). The forum post might be helpful when setting up yrange for cliff sensors - https://forum.hello-robot.com/t/working-with-cliff-sensors/88
```
num_points = 20
run_time = 10
yrange = [-50, 275]
plot_values_4(num_points, run_time, p, ["cliff_range"], yrange)
```
at_cliff: Scope base at_cliff signal
```
num_points = 20
run_time = 10
yrange = [-0.25,1.25]
plot_values_4(num_points, run_time, p, ["at_cliff"], yrange)
```
voltage: Scope bus voltage (V). This value is helpful for measuring the state of the battery in the robot. If battery is low, this value would be low. It is good practice to not let the voltage value get too low before a recharge.
```
num_points = 20
run_time = 30
# yrange = [12,15]
yrange = None
plot_values(num_points, run_time, p, ["voltage"], yrange)
```
current: Scope bus current (A)
```
num_points = 20
run_time = 10
yrange = [0,5]
plot_values(num_points, run_time, p, ["current"], yrange)
```
temp: Scope base internal temperature (C)
```
num_points = 20
run_time = 10
# yrange = [0,50]
yrange= None
plot_values(num_points, run_time, p, ["temp"], yrange)
```
IMU sensor readings: IMU stands for “Inertial Measurement Unit,” and it is used to capture data about the device’s movement. The IMU has accelerometer, gyroscope, magnetometer.
The raw data collected from an IMU gives some idea of the world around it, but that information can also be processed for additional insight. Sensor fusion is the (mathematical) art of combining the data from each sensor in an IMU to create a more complete picture of the device’s orientation and heading. For instance, while looking at gyroscope information for rotational motion, you can incorporate an accelerometers sense of gravity to create a reference frame.
The accelerometer measure the velocity and acceleration in the x,y,z direction.
ax: Scope base accelerometer AX
```
num_points = 20
run_time = 10
yrange = [-2,2]
plot_values(num_points, run_time, p, ["imu","ax"], yrange)
```
ay: Scope base accelerometer AY
```
num_points = 20
run_time = 10
yrange = [-2,2]
plot_values(num_points, run_time, p, ["imu","ay"], yrange)
```
az: Scope base accelerometer AZ
```
num_points = 20
run_time = 10
yrange = [-11,-8]
plot_values(num_points, run_time, p, ["imu","az"], yrange)
```
The magnenometer establishes cardinal direction.
mx: Scope base magnetometer MX
```
num_points = 20
run_time = 10
yrange = [-20,20]
plot_values(num_points, run_time, p, ["imu","mx"], yrange)
```
my: Scope base magnetometer MY
```
num_points = 20
run_time = 10
yrange = [-20,20]
plot_values(num_points, run_time, p, ["imu","my"], yrange)
```
mz: Scope base magnetometer MZ
```
num_points = 20
run_time = 10
yrange = [-20,20]
plot_values(num_points, run_time, p, ["imu","mz"], yrange)
```
The gyrocope measures the rotation and rotational rate.
gx: Scope base gyro GX
```
num_points = 20
run_time = 10
# yrange = [-2,2]
yrange=None
plot_values(num_points, run_time, p, ["imu","gx"], yrange)
```
gy: Scope base gyro GY
```
num_points = 20
run_time = 10
# yrange = [-2,2]
yrange=None
plot_values(num_points, run_time, p, ["imu","gy"], yrange)
```
gz: Scope base gyro GZ
```
num_points = 20
run_time = 10
# yrange = [-2,2]
yrange=None
plot_values(num_points, run_time, p, ["imu","gz"], yrange)
```
roll: Scope base imu Roll
```
num_points = 20
run_time = 10
# yrange = [-2,2]
yrange=None
plot_values(num_points, run_time, p, ["imu","roll"], yrange)
```
pitch: Scope base imu Pitch
```
num_points = 20
run_time = 10
yrange=[-20,20]
plot_values(num_points, run_time, p, ["imu","pitch"], yrange, [np.rad2deg])
```
heading: Scope base imu Heading
```
num_points = 20
run_time = 10
yrange = [-10,370]
plot_values(num_points, run_time, p, ["imu","heading"], yrange,[np.rad2deg])
```
bump: Scope base imu bump level. The bump value is useful for indicating collision of the robot. The value spikes up when there is any collision. You can test this by lightly hitting the base (be careful to not hit too hard to damage the body)
```
num_points = 20
run_time = 10
yrange = [-1,15]
plot_values(num_points, run_time, p, ["imu","bump"], yrange)
```
# Examples of using the functions
When testing out the bump function, the trigger beep could be used to generate sound vibrations that trigger the imu bump sensors. The proper working of this is a good test to see if the sensor is as sensitive as expected.
```
num_points = 30
run_time=30
iterations=2
for _ in range(iterations):
p.trigger_beep()
p.push_command()
plot_values(num_points, run_time, p, ["imu","bump"], yrange)
```
Note: Further examples to be added.
# Wrapping Up
In this notebook, we've covered:
* User/Utility functions in PIMU
* Visualisation of sensor readings from the PIMU
* Some examples of using the above
For more information on Stretch Body API, take a look at the [API Documentation](https://docs.hello-robot.com/stretch_body_guide/). To reports bugs or contribute to the library, visit the [Stretch Body Github repo](https://github.com/hello-robot/stretch_body/) where development on the library happens. Also, feel free to join our community on the [forum](https://forum.hello-robot.com/) and learn about research/projects happening with Stretch.
```
p.stop()
```
| github_jupyter |
This notebook was prepared by [Donne Martin](http://donnemartin.com). Source and license info is on [GitHub](https://github.com/donnemartin/interactive-coding-challenges).
# Solution Notebook
## Problem: Implement a linked list with insert, append, find, delete, length, and print methods.
* [Constraints](#Constraints)
* [Test Cases](#Test-Cases)
* [Algorithm](#Algorithm)
* [Code](#Code)
* [Unit Test](#Unit-Test)
## Constraints
* Can we assume this is a non-circular, singly linked list?
* Yes
* Do we keep track of the tail or just the head?
* Just the head
* Can we insert None values?
* No
## Test Cases
### Insert to Front
* Insert a None
* Insert in an empty list
* Insert in a list with one element or more elements
### Append
* Append a None
* Append in an empty list
* Insert in a list with one element or more elements
### Find
* Find a None
* Find in an empty list
* Find in a list with one element or more matching elements
* Find in a list with no matches
### Delete
* Delete a None
* Delete in an empty list
* Delete in a list with one element or more matching elements
* Delete in a list with no matches
### Length
* Length of zero or more elements
### Print
* Print an empty list
* Print a list with one or more elements
## Algorithm
### Insert to Front
* If the data we are inserting is None, return
* Create a node with the input data, set node.next to head
* Assign the head to the node
Complexity:
* Time: O(1)
* Space: O(1)
### Append
* If the data we are inserting is None, return
* Create a node with the input data
* If this is an empty list
* Assign the head to the node
* Else
* Iterate to the end of the list
* Set the final node's next to the new node
Complexity:
* Time: O(n)
* Space: O(1)
### Find
* If data we are finding is None, return
* If the list is empty, return
* For each node
* If the value is a match, return it
* Else, move on to the next node
Complexity:
* Time: O(n)
* Space: O(1)
### Delete
* If data we are deleting is None, return
* If the list is empty, return
* For each node, keep track of previous and current node
* If the value we are deleting is a match in the current node
* Update previous node's next pointer to the current node's next pointer
* We do not have have to explicitly delete in Python
* Else, move on to the next node
* As an alternative, we could avoid the use of two pointers by evaluating the current node's next value:
* If the next value is a match, set the current node's next to next.next
* Special care should be taken if deleting the head node
Complexity:
* Time: O(n)
* Space: O(1)
### Length
* For each node
* Increase length counter
Complexity:
* Time: O(n)
* Space: O(1)
### Print
* For each node
* Print the node's value
Complexity:
* Time: O(n)
* Space: O(1)
## Code
```
%%writefile linked_list.py
class Node(object):
def __init__(self, data, next=None):
self.next = next
self.data = data
def __str__(self):
return self.data
class LinkedList(object):
def __init__(self, head=None):
self.head = head
def __len__(self):
curr = self.head
counter = 0
while curr is not None:
counter += 1
curr = curr.next
return counter
def insert_to_front(self, data):
if data is None:
return None
node = Node(data, self.head)
self.head = node
return node
def append(self, data):
if data is None:
return None
node = Node(data)
if self.head is None:
self.head = Node(data)
return node
curr_node = self.head
while curr_node.next is not None:
curr_node = curr_node.next
curr_node.next = node
return node
def find(self, data):
if data is None:
return None
curr_node = self.head
while curr_node is not None:
if curr_node.data == data:
return curr_node
curr_node = curr_node.next
return None
def delete(self, data):
if data is None:
return
if self.head is None:
return
if self.head.data == data:
self.head = None
return
prev_node = self.head
curr_node = self.head.next
while curr_node is not None:
if curr_node.data == data:
prev_node.next = curr_node.next
return
prev_node = curr_node
curr_node = curr_node.next
def delete_alt(self, data):
if data is None:
return
if self.head is None:
return
curr_node = self.head
if curr_node.data == data:
curr_node = curr_node.next
return
while curr_node.next is not None:
if curr_node.next.data == data:
curr_node.next = curr_node.next.next
return
curr_node = curr_node.next
def print_list(self):
curr_node = self.head
while curr_node is not None:
print(curr_node.data)
curr_node = curr_node.next
def get_all_data(self):
data = []
curr_node = self.head
while curr_node is not None:
data.append(curr_node.data)
curr_node = curr_node.next
return data
%run linked_list.py
```
## Unit Test
```
%%writefile test_linked_list.py
from nose.tools import assert_equal
class TestLinkedList(object):
def test_insert_to_front(self):
print('Test: insert_to_front on an empty list')
linked_list = LinkedList(None)
linked_list.insert_to_front(10)
assert_equal(linked_list.get_all_data(), [10])
print('Test: insert_to_front on a None')
linked_list.insert_to_front(None)
assert_equal(linked_list.get_all_data(), [10])
print('Test: insert_to_front general case')
linked_list.insert_to_front('a')
linked_list.insert_to_front('bc')
assert_equal(linked_list.get_all_data(), ['bc', 'a', 10])
print('Success: test_insert_to_front\n')
def test_append(self):
print('Test: append on an empty list')
linked_list = LinkedList(None)
linked_list.append(10)
assert_equal(linked_list.get_all_data(), [10])
print('Test: append a None')
linked_list.append(None)
assert_equal(linked_list.get_all_data(), [10])
print('Test: append general case')
linked_list.append('a')
linked_list.append('bc')
assert_equal(linked_list.get_all_data(), [10, 'a', 'bc'])
print('Success: test_append\n')
def test_find(self):
print('Test: find on an empty list')
linked_list = LinkedList(None)
node = linked_list.find('a')
assert_equal(node, None)
print('Test: find a None')
head = Node(10)
linked_list = LinkedList(head)
node = linked_list.find(None)
assert_equal(node, None)
print('Test: find general case with matches')
head = Node(10)
linked_list = LinkedList(head)
linked_list.insert_to_front('a')
linked_list.insert_to_front('bc')
node = linked_list.find('a')
assert_equal(str(node), 'a')
print('Test: find general case with no matches')
node = linked_list.find('aaa')
assert_equal(node, None)
print('Success: test_find\n')
def test_delete(self):
print('Test: delete on an empty list')
linked_list = LinkedList(None)
linked_list.delete('a')
assert_equal(linked_list.get_all_data(), [])
print('Test: delete a None')
head = Node(10)
linked_list = LinkedList(head)
linked_list.delete(None)
assert_equal(linked_list.get_all_data(), [10])
print('Test: delete general case with matches')
head = Node(10)
linked_list = LinkedList(head)
linked_list.insert_to_front('a')
linked_list.insert_to_front('bc')
linked_list.delete('a')
assert_equal(linked_list.get_all_data(), ['bc', 10])
print('Test: delete general case with no matches')
linked_list.delete('aa')
assert_equal(linked_list.get_all_data(), ['bc', 10])
print('Success: test_delete\n')
def test_len(self):
print('Test: len on an empty list')
linked_list = LinkedList(None)
assert_equal(len(linked_list), 0)
print('Test: len general case')
head = Node(10)
linked_list = LinkedList(head)
linked_list.insert_to_front('a')
linked_list.insert_to_front('bc')
assert_equal(len(linked_list), 3)
print('Success: test_len\n')
def main():
test = TestLinkedList()
test.test_insert_to_front()
test.test_append()
test.test_find()
test.test_delete()
test.test_len()
if __name__ == '__main__':
main()
%run -i test_linked_list.py
```
| github_jupyter |
# Big Query Machine Learning (BQML)
**Learning Objectives**
- Understand that it is possible to build ML models in Big Query
- Understand when this is appropriate
- Experience building a model using BQML
# Introduction
BigQuery is more than just a data warehouse, it also has some ML capabilities baked into it.
BQML is a great option when a linear model will suffice, or when you want a quick benchmark to beat, but for more complex models such as neural networks you will need to pull the data out of BigQuery and into an ML Framework like TensorFlow.
In this notebook, we will build a naive model using BQML. **This notebook is intended to inspire usage of BQML, we will not focus on model performance.**
### Set up environment variables and load necessary libraries
```
from google import api_core
from google.cloud import bigquery
PROJECT = !gcloud config get-value project
PROJECT = PROJECT[0]
%env PROJECT=$PROJECT
```
## Create BigQuery dataset
Prior to now we've just been reading an existing BigQuery table, now we're going to create our own so so we need some place to put it. In BigQuery parlance, `Dataset` means a folder for tables.
We will take advantage of BigQuery's [Python Client](https://cloud.google.com/bigquery/docs/reference/libraries#client-libraries-install-python) to create the dataset.
```
bq = bigquery.Client(project=PROJECT)
dataset = bigquery.Dataset(bq.dataset("bqml_taxifare"))
try:
bq.create_dataset(dataset) # will fail if dataset already exists
print("Dataset created")
except api_core.exceptions.Conflict:
print("Dataset already exists")
```
## Create model
To create a model ([documentation](https://cloud.google.com/bigquery-ml/docs/reference/standard-sql/bigqueryml-syntax-create))
1. Use `CREATE MODEL` and provide a destination table for resulting model. Alternatively we can use `CREATE OR REPLACE MODEL` which allows overwriting an existing model.
2. Use `OPTIONS` to specify the model type (linear_reg or logistic_reg). There are many more options [we could specify](https://cloud.google.com/bigquery/docs/reference/standard-sql/bigqueryml-syntax-create#model_option_list), such as regularization and learning rate, but we'll accept the defaults.
3. Provide the query which fetches the training data
Have a look at [Step Two of this tutorial](https://cloud.google.com/bigquery/docs/bigqueryml-natality) to see another example.
**The query will take about two minutes to complete**
```
%%bigquery --project $PROJECT
CREATE or REPLACE MODEL bqml_taxifare.taxifare_model
OPTIONS(model_type = "linear_reg",
input_label_cols = ["label"]) AS
-- query to fetch training data
SELECT
(tolls_amount + fare_amount) AS label,
pickup_datetime,
pickup_longitude,
pickup_latitude,
dropoff_longitude,
dropoff_latitude
FROM
`nyc-tlc.yellow.trips`
WHERE
-- Clean Data
trip_distance > 0
AND passenger_count > 0
AND fare_amount >= 2.5
AND pickup_longitude > -78
AND pickup_longitude < -70
AND dropoff_longitude > -78
AND dropoff_longitude < -70
AND pickup_latitude > 37
AND pickup_latitude < 45
AND dropoff_latitude > 37
AND dropoff_latitude < 45
-- repeatable 1/5000th sample
AND ABS(MOD(FARM_FINGERPRINT(CAST(pickup_datetime AS STRING)), 5000)) = 1
```
## Get training statistics
Because the query uses a `CREATE MODEL` statement to create a table, you do not see query results. The output is an empty string.
To get the training results we use the [`ML.TRAINING_INFO`](https://cloud.google.com/bigquery/docs/reference/standard-sql/bigqueryml-syntax-train) function.
Have a look at [Step Three and Four of this tutorial](https://cloud.google.com/bigquery/docs/bigqueryml-natality) to see a similar example.
```
%%bigquery --project $PROJECT
SELECT
*
FROM
ML.TRAINING_INFO(MODEL `bqml_taxifare.taxifare_model`)
```
'eval_loss' is reported as mean squared error, so our RMSE is **8.29**. Your results may vary.
## Predict
To use our model to make predictions, we use `ML.PREDICT`. Let's, use the `taxifare_model` you trained above to infer the cost of a taxi ride that occurs at 10:00 am on January 3rd, 2014 going from the Google Office in New York (latitude: 40.7434, longitude: -74.0080) to the JFK airport (latitude: 40.6413, longitude: -73.7781)
Have a look at [Step Five of this tutorial](https://cloud.google.com/bigquery/docs/bigqueryml-natality) to see another example.
```
%%bigquery --project $PROJECT
#standardSQL
SELECT
predicted_label
FROM
ML.PREDICT(MODEL `bqml_taxifare.taxifare_model`,
(
SELECT
TIMESTAMP "2014-01-03 10:00:00" as pickup_datetime,
-74.0080 as pickup_longitude,
40.7434 as pickup_latitude,
-73.7781 as dropoff_longitude,
40.6413 as dropoff_latitude
))
```
Our model predicts the cost would be **$22.12**.
## Recap
The value of BQML is its ease of use:
- We created a model with just two additional lines of SQL
- We never had to move our data out of BigQuery
- We didn't need to use an ML Framework or code, just SQL
There's lots of work going on behind the scenes make this look easy. For example BQML is automatically creating a training/evaluation split, tuning our learning rate, and one-hot encoding features if neccesary. When we move to TensorFlow these are all things we'll need to do ourselves.
This notebook was just to inspire usage of BQML, the current model is actually very poor. We'll prove this in the next lesson by beating it with a simple heuristic.
We could improve our model considerably with some feature engineering but we'll save that for a future lesson. Also there are additional BQML functions such as `ML.WEIGHTS` and `ML.EVALUATE` that we haven't even explored. If you're interested in learning more about BQML I encourage you to [read the offical docs](https://cloud.google.com/bigquery/docs/bigqueryml).
From here on out we'll focus on pulling data out of BigQuery and building models using TensorFlow, which is more effort but also offers much more flexibility.
Copyright 2021 Google Inc.
Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
| github_jupyter |
```
%matplotlib notebook
import control as c
import ipywidgets as w
import numpy as np
from IPython.display import display, HTML
import matplotlib.pyplot as plt
import matplotlib.patches as patches
import matplotlib.animation as animation
display(HTML('<script> $(document).ready(function() { $("div.input").hide(); }); </script>'))
```
## Progettazione di un controllo per un sistema di posizionamento con vite a ricircolo di sfere azionata da un motore DC
Il seguente esempio è un'attività di progettazione di un controllo per un sistema di posizionamento con vite a ricircolo di sfere motorizzato. La struttura è costituita da un motore DC collegato a una vite a ricircolo di sfere che aziona un carrello. Il modello è costituito da molti componenti. Dopo una serie di semplificazioni, può essere ridotto a un modello lineare con dieci parametri.
<br>
Il motore può essere modellato in due parti separate, elettrica e meccanica:
<br><br>
$$G_{elec}(s)=\frac{1}{sL+R}\qquad\qquad G_{mech}=\frac{1}{sJ+B}$$
<br><br>
<img src="Images/ballscrew.png" width="40%" />
<br>
Mentre la vite a ricircolo di sfere con il carrello può essere descritta con la seguente funzione di trasferimento:
<br><br>
$$G_{ballscrew}(s)=\frac{\frac{h}{2\pi}}{s^2 m\left(\frac{1}{K}+\frac{1}{k}\right)+sb\left(\frac{1}{K}+\frac{1}{k}\right)+1}$$
<br><br>
<center>
$h:$ passo<br>
$K:$ Costante torsionale dell'albero<br>
$k:$ costante elastica lineare equivalente<br>
$b:$ smorzamento equivalente<br>
$m:$ massa del carrello
</center>
<br>
Il tuo compito è quello di scegliere un tipo di controller e progettarlo in modo da garantire prestazioni accettabili!
<b>Per prima cosa, scegli un modello del sistema!</b><br>
Alterna tra i diversi modelli con valori preselezionati casualmente (pulsanti *Modello 1* - *Modello 6*). Facendo clic sul pulsante *Preset*, vengono impostati parametri di controllo validi che non possono essere ulteriormente modificati.
```
# System parameters
# Motor parameters
R = 0.5 # Ohm - armature resistance
L = 1.5e-3 # H - armature inductance
J = 2.5e-4 # kgm^2 - rotor inertia
B = 1.0e-3 # Nms - friction as damping
kPhi = 0.05 # Nm/A or Vs/rad - Torque/back EMF constant
# Ballscrew parameters
h = [0.8, 1.0, 1.25, 0.8, 1.0, 1.25, 0.8] # mm - thread pitch
K = [2e5, 0.5e5, 1e6, 1e6, 2e5, 0.5e5, 5e5] # Nm/rad - shaft torsional spring constant
k = [1e4, 4e4, 3e4, 1e4, 3e4, 4e4, 2e4] # N/m - combined spring constants
b = [2, 8, 5, 3, 2, 10, 8] # Ns/m - combined damping
m = [0.5, 0.3, 1.5, 0.1, 0.75, 1, 1] # kg - cart mass
# Figure definition
fig1, ((f1_ax1), (f1_ax2)) = plt.subplots(2, 1)
fig1.set_size_inches((9.8, 5))
fig1.set_tight_layout(True)
f1_line1, = f1_ax1.plot([], [])
f1_line2, = f1_ax2.plot([], [])
f1_ax1.grid(which='both', axis='both', color='lightgray')
f1_ax2.grid(which='both', axis='both', color='lightgray')
f1_ax1.autoscale(enable=True, axis='both', tight=True)
f1_ax2.autoscale(enable=True, axis='both', tight=True)
f1_ax1.set_title('Diagramma del modulo', fontsize=11)
f1_ax1.set_xscale('log')
f1_ax1.set_xlabel(r'$f\/[Hz]$', labelpad=0, fontsize=10)
f1_ax1.set_ylabel(r'$A\/[dB]$', labelpad=0, fontsize=10)
f1_ax1.tick_params(axis='both', which='both', pad=0, labelsize=8)
f1_ax2.set_title('Diagramma della fase', fontsize=11)
f1_ax2.set_xscale('log')
f1_ax2.set_xlabel(r'$f\/[Hz]$', labelpad=0, fontsize=10)
f1_ax2.set_ylabel(r'$\phi\/[°]$', labelpad=0, fontsize=10)
f1_ax2.tick_params(axis='both', which='both', pad=0, labelsize=8)
def build_base_model(sel):
# DC motor model
W_v = c.tf([1], [L, R]) # Electrical part
W_m = c.tf([1], [J, B]) # Mechanical part
W_e = c.tf([kPhi], [1]) # Back EMF component
W_motor = c.feedback(c.series(W_v, W_e, W_m), W_e, -1) # Motor transfer function
# Ballscrew model
p_num = [h[sel]*1e-3/(2*np.pi)]
p_den = [(1/K[sel]+1/k[sel])*m[sel], (1/K[sel]+1/k[sel])*b[sel], 1]
W_ballscrew = c.tf(p_num, p_den)
# Complete system model
W_sys = c.series(W_motor, W_ballscrew)
print('Funzione di trasferimento del sistema:')
print(W_sys)
# System analysis
poles = c.pole(W_sys) # Poles
print('Poli del sistema:\n')
print(poles)
global f1_line1, f1_line2
f1_ax1.lines.remove(f1_line1)
f1_ax2.lines.remove(f1_line2)
mag, phase, omega = c.bode_plot(W_sys, Plot=False) # Bode-plot
f1_line1, = f1_ax1.plot(omega/2/np.pi, 20*np.log10(mag), lw=1, color='blue')
f1_line2, = f1_ax2.plot(omega/2/np.pi, phase*180/np.pi, lw=1, color='blue')
f1_ax1.relim()
f1_ax2.relim()
f1_ax1.autoscale_view()
f1_ax2.autoscale_view()
# GUI widgets
typeSelect = w.ToggleButtons(
options=[('Modello 1', 0), ('Modello 2', 1), ('Modello 3', 2), ('Modello 4', 3), ('Modello 5', 4), ('Modello 6', 5), ('Preset', -1)],
value=-1, description='Sistema: ', layout=w.Layout(width='60%'))
input_data = w.interactive_output(build_base_model, {'sel':typeSelect})
display(typeSelect, input_data)
```
Il sistema ha più poli di quelli che è possibile cancellare utilizzando un controller PID; devi selezionare quali modificare mentre i restanti rimarranno invariati.<br>
<b>Seleziona una configurazione appropriata per il controller! Qual è la migliore per il sistema scelto? Perché?<br>
Imposta il tuo controller per il tempo di assestamento più veloce possibile e per un overshoot nullo! </b>
È possibile attivare / disattivare ciascuno dei componenti I e D. Se D è attivo, è possibile applicare anche il filtro del primo ordine, regolato dalla costante di tempo di derivazione.
```
# PID position control
fig2, ((f2_ax1, f2_ax2, f2_ax3), (f2_ax4, f2_ax5, f2_ax6)) = plt.subplots(2, 3)
fig2.set_size_inches((9.8, 5))
fig2.set_tight_layout(True)
f2_line1, = f2_ax1.plot([], [])
f2_line2, = f2_ax2.plot([], [])
f2_line3, = f2_ax3.plot([], [])
f2_line4, = f2_ax4.plot([], [])
f2_line5, = f2_ax5.plot([], [])
f2_line6, = f2_ax6.plot([], [])
f2_ax1.grid(which='both', axis='both', color='lightgray')
f2_ax2.grid(which='both', axis='both', color='lightgray')
f2_ax3.grid(which='both', axis='both', color='lightgray')
f2_ax4.grid(which='both', axis='both', color='lightgray')
f2_ax5.grid(which='both', axis='both', color='lightgray')
f2_ax6.grid(which='both', axis='both', color='lightgray')
f2_ax1.autoscale(enable=True, axis='both', tight=True)
f2_ax2.autoscale(enable=True, axis='both', tight=True)
f2_ax3.autoscale(enable=True, axis='both', tight=True)
f2_ax4.autoscale(enable=True, axis='both', tight=True)
f2_ax5.autoscale(enable=True, axis='both', tight=True)
f2_ax6.autoscale(enable=True, axis='both', tight=True)
f2_ax1.set_title('Risposta al gradino in anello chiuso', fontsize=9)
f2_ax1.set_xlabel(r'$t\/$[s]', labelpad=0, fontsize=8)
f2_ax1.set_ylabel(r'$x\/$[m]', labelpad=0, fontsize=8)
f2_ax1.tick_params(axis='both', which='both', pad=0, labelsize=6)
f2_ax2.set_title('Diagramma di Nyquist', fontsize=9)
f2_ax2.set_xlabel(r'Re', labelpad=0, fontsize=8)
f2_ax2.set_ylabel(r'Im', labelpad=0, fontsize=8)
f2_ax2.tick_params(axis='both', which='both', pad=0, labelsize=6)
f2_ax3.set_title('Diagramma del modulo', fontsize=9)
f2_ax3.set_xscale('log')
f2_ax3.set_xlabel(r'$f\/$[Hz]', labelpad=0, fontsize=8)
f2_ax3.set_ylabel(r'$A\/$[dB]', labelpad=0, fontsize=8)
f2_ax3.tick_params(axis='both', which='both', pad=0, labelsize=6)
f2_ax4.set_title('Risposta impulsiva in anello chiuso', fontsize=9)
f2_ax4.set_xlabel(r'$t\/$[s]', labelpad=0, fontsize=8)
f2_ax4.set_ylabel(r'$x\/$[m]', labelpad=0, fontsize=8)
f2_ax4.tick_params(axis='both', which='both', pad=0, labelsize=6)
f2_ax5.set_title('Risposta al disturbo a gradino in ingresso', fontsize=9)
f2_ax5.set_xlabel(r'$t\/$[s]', labelpad=0, fontsize=8)
f2_ax5.set_ylabel(r'$x\/$[m]', labelpad=0, fontsize=8)
f2_ax5.tick_params(axis='both', which='both', pad=0, labelsize=6)
f2_ax6.set_title('Diagramma della fase', fontsize=9)
f2_ax6.set_xscale('log')
f2_ax6.set_xlabel(r'$f\/$[Hz]', labelpad=0, fontsize=8)
f2_ax6.set_ylabel(r'$\phi\/$[°]', labelpad=0, fontsize=8)
f2_ax6.tick_params(axis='both', which='both', pad=0, labelsize=6)
def position_control(Kp, Ti, Td, Fd, Ti0, Td0, Fd0, sel):
W_motor = c.series(c.tf([1], [L, R]), c.tf([1], [J, B]), c.tf([kPhi], [1]))
W_ballscrew = c.tf([h[sel]*1e-3/(2*np.pi)], [(1/K[sel]+1/k[sel])*m[sel], (1/K[sel]+1/k[sel])*b[sel], 1])
W_sys = c.series(W_motor, W_ballscrew)
# PID Controller
P = Kp # Proportional term
I = Kp / Ti # Integral term
D = Kp * Td # Derivative term
Td_f = Td / Fd # Derivative term filter
W_PID = c.parallel(c.tf([P], [1]),
c.tf([I * Ti0], [1 * Ti0, 1 * (not Ti0)]),
c.tf([D * Td0, 0], [Td_f * Td0 * Fd0, 1])) # PID controller in time constant format
W_open = c.series(W_PID, W_sys, c.tf([1], [1, 0])) # Open loop with integrator added for position output
W_closed = c.feedback(W_open, 1, -1) # Closed loop with negative feedback
W_cart = c.tf([1],[m[sel], b[sel]]) # Ballscrew ring transfer function component
W_rest = c.series(W_open, c.tf([m[sel], b[sel]], [1]), c.tf([1, 0], [1])) # The system without the cart and the integrator
W_load = c.feedback(c.series(W_cart, c.tf([1], [1, 0])), W_rest, -1) # Transfer function of the load based errors
# Display
global f2_line1, f2_line2, f2_line3, f2_line4, f2_line5, f2_line6
f2_ax1.lines.remove(f2_line1)
f2_ax2.lines.remove(f2_line2)
f2_ax3.lines.remove(f2_line3)
f2_ax4.lines.remove(f2_line4)
f2_ax5.lines.remove(f2_line5)
f2_ax6.lines.remove(f2_line6)
tout, yout = c.step_response(W_closed)
f2_line1, = f2_ax1.plot(tout, yout, lw=1, color='blue')
_, _, ob = c.nyquist_plot(W_open, Plot=False) # Small resolution plot to determine bounds
real, imag, freq = c.nyquist_plot(W_open, omega=np.logspace(np.log10(ob[0]), np.log10(ob[-1]), 1000), Plot=False)
f2_line2, = f2_ax2.plot(real, imag, lw=1, color='blue')
mag, phase, omega = c.bode_plot(W_open, Plot=False)
f2_line3, = f2_ax3.plot(omega/2/np.pi, 20*np.log10(mag), lw=1, color='blue')
f2_line6, = f2_ax6.plot(omega/2/np.pi, phase*180/np.pi, lw=1, color='blue')
tout, yout = c.impulse_response(W_closed)
f2_line4, = f2_ax4.plot(tout, yout, lw=1, color='blue')
tout, yout = c.step_response(W_load)
f2_line5, = f2_ax5.plot(tout, yout, lw=1, color='blue')
f2_ax1.relim()
f2_ax2.relim()
f2_ax3.relim()
f2_ax4.relim()
f2_ax5.relim()
f2_ax6.relim()
f2_ax1.autoscale_view()
f2_ax2.autoscale_view()
f2_ax3.autoscale_view()
f2_ax4.autoscale_view()
f2_ax5.autoscale_view()
f2_ax6.autoscale_view()
def update_controller(index):
global Kp_slider, Ti_slider, Td_slider, Fd_slider, Ti_button, Td_button, Fd_button
if index == -1:
Kp_slider.value = 200
Td_slider.value = 0.01
Fd_slider.value = 10
Ti_button.value = False
Td_button.value = True
Fd_button.value = True
Kp_slider.disabled = True
Ti_slider.disabled = True
Td_slider.disabled = True
Fd_slider.disabled = True
Ti_button.disabled = True
Td_button.disabled = True
Fd_button.disabled = True
else:
Kp_slider.disabled = False
Ti_slider.disabled = False
Td_slider.disabled = False
Fd_slider.disabled = False
Ti_button.disabled = False
Td_button.disabled = False
Fd_button.disabled = False
# Controllers
Kp_slider = w.FloatLogSlider(value=0.5, base=10, min=-3, max=3, description='Kp:', continuous_update=False,
layout=w.Layout(width='auto', flex='5 5 auto'))
Ti_slider = w.FloatLogSlider(value=0.0035, base=10, min=-4, max=1, description='', continuous_update=False,
layout=w.Layout(width='auto', flex='5 5 auto'))
Td_slider = w.FloatLogSlider(value=1, base=10, min=-4, max=1, description='', continuous_update=False,
layout=w.Layout(width='auto', flex='5 5 auto'))
Fd_slider = w.FloatLogSlider(value=1, base=10, min=0, max=3, description='', continuous_update=False,
layout=w.Layout(width='auto', flex='5 5 auto'))
Ti_button = w.ToggleButton(value=True, description='Ti',
layout=w.Layout(width='auto', flex='1 1 0%'))
Td_button = w.ToggleButton(value=False, description='Td',
layout=w.Layout(width='auto', flex='1 1 0%'))
Fd_button = w.ToggleButton(value=False, description='Fd',
layout=w.Layout(width='auto', flex='1 1 0%'))
input_data = w.interactive_output(position_control, {'Kp': Kp_slider, 'Ti': Ti_slider, 'Td': Td_slider,
'Fd': Fd_slider, 'Ti0' : Ti_button, 'Td0': Td_button,
'Fd0': Fd_button, 'sel':typeSelect})
w.interactive_output(update_controller, {'index':typeSelect})
display(w.HBox([Kp_slider, Ti_button, Ti_slider, Td_button, Td_slider, Fd_button, Fd_slider]), input_data)
```
Nella seguente simulazione è possibile osservare il movimento del sistema in base ai parametri scelti del controller. È possibile creare segnali di riferimento e applicare anche disturbi.
<b>La configurazione scelta è adatta all'inseguimento del riferimento? Modifica il controller in modo che il sistema controllato riesca a seguire una sinusoide in maniera accettabile!</b>
<br><br>
<i>(Data la scalatura automatica del grafico, le soluzioni instabili potrebbero apparire nulle fino a poco prima dell'ultimo istante rappresentato.)</i>
```
# Simulation data
anim_fig = plt.figure()
anim_fig.set_size_inches((9.8, 6))
anim_fig.set_tight_layout(True)
anim_ax1 = anim_fig.add_subplot(211)
anim_ax2 = anim_ax1.twinx()
frame_count=1000
l1 = anim_ax1.plot([], [], lw=1, color='blue')
l2 = anim_ax1.plot([], [], lw=2, color='red')
l3 = anim_ax2.plot([], [], lw=1, color='grey')
line1 = l1[0]
line2 = l2[0]
line3 = l3[0]
anim_ax1.legend(l1+l2+l3, ['Riferimento [m]', 'Uscita [m]', 'Disturbo [N]'], loc=1)
anim_ax1.set_title('Simulazione', fontsize=12)
anim_ax1.set_xlabel(r'$t\/$[s]', labelpad=0, fontsize=10)
anim_ax1.set_ylabel(r'$x\/$[m]', labelpad=0, fontsize=10)
anim_ax1.tick_params(axis='both', which='both', pad=0, labelsize=8)
anim_ax2.set_ylabel(r'$F\/$[N]', labelpad=0, fontsize=10)
anim_ax2.tick_params(axis='both', which='both', pad=0, labelsize=8)
anim_ax1.grid(which='both', axis='both', color='lightgray')
T_plot = []
U_plot = []
F_plot = []
R_plot = []
# Scene data
scene_ax = anim_fig.add_subplot(212)
scene_ax.set_xlim((-3, 3))
scene_ax.set_ylim((-1, 1))
scene_ax.axis('off')
scene_ax.add_patch(patches.Ellipse((-2.5, 0), 0.2, 1, fill=True, fc='indianred', lw=0, zorder=0))
scene_ax.add_patch(patches.Ellipse((-2, 0), 0.2, 1, fill=True, fc='firebrick', lw=0, zorder=5))
scene_ax.add_patch(patches.Rectangle((-2.5, -0.5), 0.5, 1, fill=True, fc='indianred', lw=0, zorder=0))
scene_ax.add_patch(patches.Arc((-2.5, 0), 0.2, 1, theta1=90, theta2=270, color='maroon', lw=1.5, zorder=10))
scene_ax.add_patch(patches.Arc((-2, 0), 0.2, 1, theta1=90, theta2=270, color='maroon', lw=1.5, zorder=30))
scene_ax.add_patch(patches.Arc((-2, 0), 0.2, 1, theta1=270, theta2=90, color='maroon', lw=1.5, zorder=0))
scene_ax.add_patch(patches.Rectangle((-2.48, -0.55), 0.12, 0.1, fill=True, ec='maroon', fc='firebrick', lw=1.5, zorder=15))
scene_ax.add_patch(patches.Rectangle((-2.22, -0.55), 0.12, 0.1, fill=True, ec='maroon', fc='firebrick', lw=1.5, zorder=15))
scene_ax.plot([-2.5, -2], [-0.5, -0.5], color='maroon', lw=1.5, zorder=5)
scene_ax.plot([-2.5, -2], [0.5, 0.5], color='maroon', lw=1.5, zorder=5)
scene_ax.plot([-2.55, -2.15], [-0.1, -0.1], color='maroon', lw=1.5, zorder=5)
scene_ax.plot([-2.55, -2.15], [0.1, 0.1], color='maroon', lw=1.5, zorder=5)
scene_ax.add_patch(patches.Rectangle((2.4, -0.55), 0.25, 0.75, fill=True, ec='darkgreen', fc='forestgreen', lw=1.5, zorder=25))
scene_ax.add_patch(patches.Rectangle((-2.1, -0.075), 0.1, 0.15, fill=True, fc='firebrick', lw=0, zorder=25))
scene_ax.add_patch(patches.Rectangle((-2.2, -0.075), 0.1, 0.15, fill=True, fc='indianred', lw=0, zorder=25))
scene_ax.add_patch(patches.Rectangle((-2.15, -0.05), 4.9, 0.1, fill=True, ec='silver',
fc='dimgrey', lw=1.5, zorder=10))
scene_ax.add_patch(patches.Arc((-1.99, 0), 0.02, 0.1, theta1=90, theta2=270, color='silver', lw=1.5, zorder=30))
scene_ax.add_patch(patches.Arc((-2, 0), 0.04, 0.15, theta1=80, theta2=270, color='maroon', lw=1.5, zorder=30))
screw_base_x = np.tile([0, 0.15, 0.2, 0.05], 30) + np.repeat(np.linspace(0, 4.5, 30), 4)
screw_base_y = np.tile([-0.05, 0.05, 0.05, -0.05], 30)
screw, = scene_ax.plot(screw_base_x - 2.2, screw_base_y, color='silver', lw=1, zorder=15)
cart_1 = patches.Rectangle((-0.35, -0.1), 0.2, 0.2, fill=True, ec='darkslategray', fc='cornflowerblue', lw=1.5, zorder=35)
cart_2 = patches.Rectangle((-0.15, -0.3), 0.3, 0.4, fill=True, ec='darkslategray', fc='royalblue', lw=1.5, zorder=35)
cart_3 = patches.Rectangle((0.15, -0.1), 0.2, 0.2, fill=True, ec='darkslategray', fc='cornflowerblue', lw=1.5, zorder=35)
scene_ax.add_patch(cart_1)
scene_ax.add_patch(cart_2)
scene_ax.add_patch(cart_3)
u_arrow = scene_ax.arrow(0, -0.75, 0, 0.25, color='blue', head_width=0.1,
length_includes_head=True, lw=1, fill=False, zorder=10)
r_arrow = scene_ax.arrow(0, -0.75, 0, 0.25, color='red', head_width=0.1,
length_includes_head=True, lw=1, fill=False, zorder=10)
base_arrow = u_arrow.xy
pos_var = []
ref_var = []
#Simulation function
def simulation(Kp, Ti, Td, Fd, Ti0, Td0, Fd0, sel, T, dt, U, Uf, Ua, Uo, F, Ff, Fa, Fo):
# Controller
P = Kp # Proportional term
I = Kp / Ti # Integral term
D = Kp * Td # Derivative term
Td_f = Td / Fd # Derivative term filter
W_PID = c.parallel(c.tf([P], [1]),
c.tf([I * Ti0], [1 * Ti0, 1 * (not Ti0)]),
c.tf([D * Td0, 0], [Td_f * Td0 * Fd0, 1])) # PID controller
# System
W_motor = c.series(c.tf([1], [L, R]), c.tf([1], [J, B]), c.tf([kPhi], [1]))
W_ballscrew = c.tf([h[sel]*1e-3/(2*np.pi)], [(1/K[sel]+1/k[sel])*m[sel], (1/K[sel]+1/k[sel])*b[sel], 1])
W_sys = c.series(W_motor, W_ballscrew)
# Model
W_open = c.series(W_PID, W_sys, c.tf([1], [1, 0])) # Open loop with integrator added for position output
W_closed = c.feedback(W_open, 1, -1) # Closed loop with negative feedback
W_cart = c.tf([1],[m[sel], b[sel]]) # Ballscrew ring transfer function component
W_rest = c.series(W_open, c.tf([m[sel], b[sel]], [1]), c.tf([1, 0], [1])) # The system without the cart and the integrator
W_load = c.feedback(c.series(W_cart, c.tf([1], [1, 0])), W_rest, -1) # Transfer function of the load based errors
# Reference and disturbance signals
T_sim = np.arange(0, T, dt, dtype=np.float64)
if U == 0: # Constant reference
U_sim = np.full_like(T_sim, Ua * Uo)
elif U == 1: # Sine wave reference
U_sim = (np.sin(2 * np.pi * Uf * T_sim) + Uo) * Ua
elif U == 2: # Square wave reference
U_sim = (np.sign(np.sin(2 * np.pi * Uf * T_sim)) + Uo) * Ua
if F == 0: # Constant load
F_sim = np.full_like(T_sim, Fa * Fo)
elif F == 1: # Sine wave load
F_sim = (np.sin(2 * np.pi * Ff * T_sim) + Fo) * Fa
elif F == 2: # Square wave load
F_sim = (np.sign(np.sin(2 * np.pi * Ff * T_sim)) + Fo) * Fa
elif F == 3: # Noise form load
F_sim = np.interp(T_sim, np.linspace(0, T, int(T * Ff) + 2),
np.random.normal(loc=(Fo * Fa), scale=Fa, size=int(T * Ff) + 2))
# System response
Tu, youtu, xoutu = c.forced_response(W_closed, T_sim, U_sim)
Tf, youtf, xoutf = c.forced_response(W_load, T_sim, F_sim)
R_sim = np.nan_to_num(youtu + youtf)
# Display
UR_max = max(np.amax(np.absolute(np.concatenate((U_sim, R_sim)))), Ua)
F_max = max(np.amax(np.absolute(F_sim)), Fa)
anim_ax1.set_xlim((0, T))
anim_ax1.set_ylim((-1.2 * UR_max, 1.2 * UR_max))
anim_ax2.set_ylim((-1.5 * F_max, 1.5 * F_max))
global T_plot, U_plot, F_plot, R_plot, pos_var, ref_var
T_plot = np.linspace(0, T, frame_count, dtype=np.float32)
U_plot = np.interp(T_plot, T_sim, U_sim)
F_plot = np.interp(T_plot, T_sim, F_sim)
R_plot = np.interp(T_plot, T_sim, R_sim)
pos_var = R_plot/UR_max
ref_var = U_plot/UR_max
def anim_init():
line1.set_data([], [])
line2.set_data([], [])
line3.set_data([], [])
screw.set_data(screw_base_x - 2.2, screw_base_y)
cart_1.set_xy((-0.35, -0.1))
cart_2.set_xy((-0.15, -0.2))
cart_3.set_xy((0.15, -0.1))
u_arrow.set_xy(base_arrow)
r_arrow.set_xy(base_arrow)
return (line1, line2, line3, screw, cart_1, cart_2, cart_3, u_arrow, r_arrow,)
def animate(i):
line1.set_data(T_plot[0:i], U_plot[0:i])
line2.set_data(T_plot[0:i], R_plot[0:i])
line3.set_data(T_plot[0:i], F_plot[0:i])
screw_shift = 1 - np.remainder(pos_var[i] * 10, 1)
screw.set_data(screw_base_x - 2.2 + 0.15 * screw_shift, screw_base_y)
cart_1.set_x(pos_var[i] * 1.5 - 0.35)
cart_2.set_x(pos_var[i] * 1.5 - 0.15)
cart_3.set_x(pos_var[i] * 1.5 + 0.15)
u_arrow.set_xy(base_arrow+[ref_var[i] * 1.5, 0])
r_arrow.set_xy(base_arrow+[pos_var[i] * 1.5, 0])
return (line1, line2, line3, screw, cart_1, cart_2, cart_3, u_arrow, r_arrow,)
anim = animation.FuncAnimation(anim_fig, animate, init_func=anim_init,
frames=frame_count, interval=10, blit=True,
repeat=True)
# Controllers
T_slider = w.FloatLogSlider(value=10, base=10, min=-0.7, max=1, step=0.01,
description='Durata [s]:', continuous_update=False,
orientation='vertical', layout=w.Layout(width='auto', height='auto', flex='1 1 auto'))
dt_slider = w.FloatLogSlider(value=0.1, base=10, min=-3, max=-1, step=0.01,
description='Tempo di campionamento [s]:', continuous_update=False,
orientation='vertical', layout=w.Layout(width='auto', height='auto', flex='1 1 auto'))
U_type = w.Dropdown(options=[('Costante', 0), ('Sinusoide', 1), ('Onda quadra', 2)], value=1,
description='Riferimento: ', continuous_update=False, layout=w.Layout(width='auto', flex='3 3 auto'))
Uf_slider = w.FloatLogSlider(value=0.5, base=10, min=-2, max=2, step=0.01,
description='Frequenza [Hz]:', continuous_update=False,
orientation='vertical', layout=w.Layout(width='auto', height='auto', flex='1 1 auto'))
Ua_slider = w.FloatLogSlider(value=1, base=10, min=-2, max=2, step=0.01,
description='Ampiezza [m]:', continuous_update=False,
orientation='vertical', layout=w.Layout(width='auto', height='auto', flex='1 1 auto'))
Uo_slider = w.FloatSlider(value=0, min=-10, max=10, description='Offset/Amp:', continuous_update=False,
orientation='vertical', layout=w.Layout(width='auto', height='auto', flex='1 1 auto'))
F_type = w.Dropdown(options=[('Costante', 0), ('Sinusoide', 1), ('Onda quadra', 2), ('Rumore bianco', 3)], value=2,
description='Load: ', continuous_update=False, layout=w.Layout(width='auto', flex='3 3 auto'))
Ff_slider = w.FloatLogSlider(value=1, base=10, min=-2, max=2, step=0.01,
description='Frequenza [Hz]:', continuous_update=False,
orientation='vertical', layout=w.Layout(width='auto', height='auto', flex='1 1 auto'))
Fa_slider = w.FloatLogSlider(value=0.1, base=10, min=-2, max=2, step=0.01,
description='Ampiezza [N]:', continuous_update=False,
orientation='vertical', layout=w.Layout(width='auto', height='auto', flex='1 1 auto'))
Fo_slider = w.FloatSlider(value=0, min=-10, max=10, description='Offset/Amp:', continuous_update=False,
orientation='vertical', layout=w.Layout(width='auto', height='auto', flex='1 1 auto'))
input_data = w.interactive_output(simulation, {'Kp': Kp_slider, 'Ti': Ti_slider, 'Td': Td_slider,
'Fd': Fd_slider, 'Ti0' : Ti_button, 'Td0': Td_button,
'Fd0': Fd_button, 'sel':typeSelect,
'T': T_slider, 'dt': dt_slider,
'U': U_type, 'Uf': Uf_slider, 'Ua': Ua_slider, 'Uo': Uo_slider,
'F': F_type, 'Ff': Ff_slider, 'Fa': Fa_slider, 'Fo': Fo_slider})
display(w.HBox([w.HBox([T_slider, dt_slider], layout=w.Layout(width='25%')),
w.Box([], layout=w.Layout(width='5%')),
w.VBox([U_type, w.HBox([Uf_slider, Ua_slider, Uo_slider])], layout=w.Layout(width='30%')),
w.Box([], layout=w.Layout(width='5%')),
w.VBox([F_type, w.HBox([Ff_slider, Fa_slider, Fo_slider])], layout=w.Layout(width='30%'))],
layout=w.Layout(width='100%', justify_content='center')), input_data)
```
Il parametro della durata controlla il periodo di tempo simulato e non influisce sul tempo di esecuzione dell'animazione. Il tempo di campionamento controlla il campionamento del modello e può migliorare i risultati della simulazione in cambio di risorse di calcolo più elevate.
| github_jupyter |
```
# Initialize OK
from client.api.notebook import Notebook
ok = Notebook('hw5.ok')
```
# Homework 5: Exploring bias through Cook County’s property assessments
## Due Date: 11:59pm Monday, March 30
### Collaboration Policy
Data science is a collaborative activity. While you may talk with others about the homework, we ask that you **write your solutions individually**. If you do discuss the assignments with others please **include their names** in the collaborators cell below.
**Collaborators:** *list names here*
## Introduction
In this homework, we will go through the iterative process of specifying, fitting, and analyzing the performance of a model.
In the first portion of the assignment, we will guide you through some basic exploratory data analysis (EDA), laying out the thought process that leads to certain modeling decisions. Next, you will add a new feature to the dataset, before specifying and fitting a linear model to a few features of the housing data to predict housing prices. Finally, we will analyze the error of the model and brainstorm ways to improve the model's performance.
After this homework, you should feel comfortable with the following:
1. Simple feature engineering
1. Using sklearn to build linear models
1. Building a data pipeline using pandas
Next week's homework will continue working with this dataset to address more advanced and subtle issues with modeling.
## HCE Learning Outcomes
By working through this homework, students will be able to:
* Identify sources of bias within social and technical decisions.
* Understand how bias emerges from the human contexts of data science work, specifically through professions and institutions.
* Recognize that, because of this human context, bias is structural to data science throughout the data lifecycle rather than an individual, subjective variable that can be eradicated.
* Analyze the effects of both deliberate and unintentional choices made throughout their work in order to meaningfully address questions of fairness.
## Score Breakdown
*To be edited by course staff.*
```
import numpy as np
import pandas as pd
from pandas.api.types import CategoricalDtype
%matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
# Plot settings
plt.rcParams['figure.figsize'] = (12, 9)
plt.rcParams['font.size'] = 12
```
# Part 0: Contextualizing the Data
### The Cook County Assessor’s Office
The dataset you’ll be working with comes from the Cook County Assessor’s Office (CCAO) in Illinois, a government institution that determines property taxes across most of Chicago’s metropolitan area and its nearby suburbs. These property tax assessments are based on property values estimated using statistical models that consider multiple factors, such as real estate value and construction cost.
This system, however, is not without flaws. In late 2017, a lawsuit was filed against the office of Cook County Assessor Joseph Berrios for producing [“racially discriminatory assessments and taxes."](https://harris.uchicago.edu/news-events/news/prof-chris-berry-testifies-institutional-racism-cook-county-property-taxes) The lawsuit included claims that the assessor’s office undervalued high-priced homes and overvalued low-priced homes, creating a visible divide along racial lines: Wealthy homeowners, who were typically white, [paid less in property taxes](https://www.clccrul.org/bpnc-v-berrios-facts?rq=berrios), whereas working-class, [non-white homeowners paid more](https://www.chicagotribune.com/news/breaking/ct-cook-county-assessor-berrios-sued-met-20171214-story.html).
Although there are certainly many layers to the discrimination in this case (Chicago’s history of redlining, for example), unfair property taxation arose in part because of the tax lawyer industry. After the assessor’s office releases their property valuations, tax lawyers have the power to negotiate for lower valuations - hence, lower property taxes - in an appeals process. And because hiring a tax lawyer is limited to those who can afford it, wealthy property owners ultimately paid a disproportionate amount of tax compared to those of lower socioeconomic status - people who are predominantly from Hispanic and Black communities in Cook County. In this way, inequity along racial lines is systemically built into the property assessment system.
### Open Data and the CCAO
To push back against this systemic inequity, Berrios’ successor, Cook County Assessor Fritz Kaegi, promised to increase transparency in the assessment process. The hope was that, by exposing the inner workings of the CCAO’s property valuation process, their assessment results could be publicly verified as accurate and therefore trusted to be fair. Additionally, tax lawyers would be unable to contest the validity of the Office’s data and modeling process.
Through Cook County’s [open data hub](https://datacatalog.cookcountyil.gov/), the CCAO publishes the data they use in their model to valuate properties. Their [model, written in R,](https://gitlab.com/ccao-data-science---modeling) is also available for viewing. If you’re interested in reading more about how this initiative is being presented by the CCAO, you can check out their [Medium article](https://medium.com/@AssessorCook/why-the-cook-county-assessors-office-made-its-residential-assessment-code-and-data-public-c964acfa7b0f).
Though the CCAO’s transparency initiative is the first of its kind in the country, it is nonetheless open to critique: The Office itself solicits community input on the quality and fairness of its model. And in this homework, we will address the different types of bias that may arise in the modeling process.
## HCE: Question -1
Based on the preceding introductory paragraphs, in what way was the previous property assessment process discriminatory? How has the CCAO decided to combat this issue? (1-2 sentences total)
`TODO`: *Write your answer here*
### The Data
The Cook County dataset you'll be using in this homework contains residential sales data that the CCAO uses to assess property values. A more detailed description of each variable is available on [their website](https://datacatalog.cookcountyil.gov/Property-Taxation/Cook-County-Assessor-s-Residential-Sales-Data/5pge-nu6u). There, the list of column values notes what each entry represents and sometimes cautions about the quality of a given variable. **You should take some time to familiarize yourself with the variables before moving forward.**
You can also read more about the context of this dataset from the [Cook County Assessor's Office Data Narrative](https://datacatalog.cookcountyil.gov/stories/s/Cook-County-Assessor-Valuation-Data-Release/p2kt-hk36).
```
# Display a list of the columns in the dataset.
```
You might notice that the columns contain references to apartments, condos, and commercial buildings. However, we’ll only be focusing on single-family houses for this homework. Run the following cell to filter out entities that aren’t homes.
```
# Filter out properties that aren't single-family homes. Refer to the 'Technical Checks' Google Doc.
```
The data is split into training and test sets with X and Y observations, respectively.
```
training_data = pd.read_csv("ccao_train.csv")
test_data = pd.read_csv("ccao_test.csv")
```
As a good sanity check, we should at least verify that the data shape matches the description.
```
# Edit these sanity checks to match the CCAO dataset.
# 2000 observations and 82 features in training data
assert training_data.shape == (2000, 82)
# 930 observations and 81 features in test data
assert test_data.shape == (930, 81)
# SalePrice is hidden in the test data
assert 'SalePrice' not in test_data.columns.values
# Every other column in the test data should be in the training data
assert len(np.intersect1d(test_data.columns.values,
training_data.columns.values)) == 81
```
### Data Discovery
Prior to the open data initiative and even prior to the assessment modeling initiative, Cook County’s assessor office received much of their data for assessments from their relationships with [“local elected officials, community leaders, real estate professionals and other citizens knowledgeable about real estate in the area.”](https://www.cookcountyassessor.com/about-cook-county-assessors-office) Because CCAO field inspectors cannot enter homes to gather data, this information must be gathered through either curbside observations or real estate records.
You can read more about data collection in the CCAO’s [Residential Data Integrity Preliminary Report](https://gitlab.com/ccao-data-science---modeling/ccao_sf_cama_dev/-/blob/master/documentation/Preliminary%20Report%20on%20Data%20Integrity%20June%207,%202019.pdf).
## HCE: Question 0
How is the CCAO’s data collected? Discuss an obstacle to CCAO's data collection efforts and address how it may limit the CCAO's goals.
`TODO`: *Write your answer here*
## HCE: Question 1
Take a moment to assess the granularity of this dataset. What types of information are contained in a row?
`TODO`: *Write your answer here*
## HCE: Question 2
Given that this data compiled by the CCAO (as opposed to other assessor’s offices), find a column that would most likely only be collected in Cook County. What does this column represent?
*Hint:* Look for a feature related to Chicago’s airports.
`TODO`: *Write your answer here*
## HCE: Question 3
Name a feature that isn't listed in this dataset but may be useful for predicting sales values. What insights could this feature provide? How might it increase or decrease a home’s sales value?
*Hint:* If you’re stuck, refer to the “Single-Family Properties Appropriateness” section in the CCAO’s [Residential Data Integrity Preliminary Report](https://gitlab.com/ccao-data-science---modeling/ccao_sf_cama_dev/-/blob/master/documentation/Preliminary%20Report%20on%20Data%20Integrity%20June%207,%202019.pdf).
`TODO`: *Write your answer here*
Now that we’re more familiar with the information that the dataset contains, let’s focus on the features themselves. Many features are categorical, utilizing classification to describe particular aspects of a home. Some features are straightforward in their categories - the `Wall Material` column, for example, has values that identify whether a property’s external wall is built with wood, masonry, both wood and masonry, or stucco. Others, however, are not so clearly defined.
## HCE: Question 4
Let’s take a look at the `Site Desirability` column. What do the column’s values represent? Does the documentation provide sufficient guidelines as to how a property's `Site Desirability` is determined? Why or why not?
`TODO`: *Write your answer here*
Just by looking at the feature and its description, we can tell that `Site Desirability` is highly discretionary - that is, it relies on an individual’s interpretation. And in this context, the individual in question would be a real estate agent or assessor.
## HCE: Question 5
Beyond a home’s internal characteristics (such as number of rooms, bathrooms, etc.), describe a factor that might influence whether a home is desirable, and elaborate why. Think from the perspective of a real estate agent - i.e. what would an agent market to potential buyers and why?
*Hint:* Consider writing about characteristics related to a home’s location and its proximity to other places.
`TODO`: *Write your answer here*
When examining your data, it’s important to consider who collects the data in order to understand the assumptions and perspectives built into it. Here, the idea of desirable properties is influenced by what the real estate industry - a source of expertise in home valuations - deems popular with its market. These human choices about value are shaped by the real estate agents' expertise acquired through their professional training and experience. From a seller's perspective, this expertise also makes the agent's valuations legitimate and authoritative in a way that's difficult to contest. As a result, a form of professional bias enters into the valuation process, and though this bias isn’t inherently bad, we’ll delve deeper into how it interacts with, and often reinforces, structural inequity.
To continue using `Site Desirability` as an example, let’s say that proximity to high-ranking schools adds to a home’s desirability. Because schools are largely funded by property taxes, high-ranking schools are typically located near homes in higher-income neighborhoods. Comparatively, homes in lower-income neighborhoods - with families predominantly from Hispanic and Black communities - would be deemed “less desirable” because they would not feed into high-ranking schools, further devaluing homes in lower-income neighborhoods.
In this way, `Site Desirability` acts as a proxy for sensitive attributes, such as the racial distribution of a neighborhood. Although it does not explicitly name these attributes, `Site Desirability` ultimately encodes the broader socioeconomic and racial context of a home’s surrounding community. And because real estate agents affirm their classifications of `Site Desirability` based on their expertise, structural inequity is further perpetuated as this bias is embedded in the data used to generate housing valuation models.
## HCE: Question 6
Although it’s impossible to remove bias (especially given that data is often collected through a specific profession’s lens), it’s nonetheless important to be aware of how choosing dataset features both reflects and produces real-world contexts.
Find another feature in the dataset that, similarly to Site Desirability, encodes bias through expertise, and support your reasoning.
`TODO`: *Write your answer here*
Understanding your dataset provides a lot of insight into how models might incorporate bias from the very beginning. A data scientist with this awareness would not only identify sources of bias, but also aim to intentionally address these biases in their data analyses, as well as the outputs and recommendations based on these analyses. As we progress through this homework, keep these perspectives on bias and expertise in mind.
# Part 1: Exploratory Data Analysis
In this section, we will make a series of exploratory visualizations and interpret them. This will allow us to familiarize ourselves with how particular columns within the data might influence a model’s predictions, as well as expose discrepancies in data collection.
Note that we will perform EDA on the **training data** so that information from the test data does not influence our modeling decisions.
### Construction Quality
As you may have observed in Part 0, `Construction Quality` is a feature that may introduce existing inequities to a model as a result of its discretionary classifications. Creating visualizations as part of EDA will allow us to further examine these classifications.
## HCE: Question 7
Look at the feature `Construction Quality` referenced in the [codebook](https://datacatalog.cookcountyil.gov/Property-Taxation/Cook-County-Assessor-s-Residential-Sales-Data/5pge-nu6u). Make a visualization of this feature and, in one sentence, describe a concern you might have about using this feature to predict a home’s value and why.
*Hint:* Pay attention to how the variable is distributed.
`TODO`: *Write your answer here*
As your work in Question 7 demonstrates, visualizations can directly reveal the underlying subtext of particular variables. Beyond the usefulness of the data itself, visualizing `Construction Quality` sheds light on deeper implications. Observe how average-rated houses are established as an overwhelming norm, which thus draws additional scrutiny to houses that are rated “Deluxe” and “Poor". Like `Site Desirability`, there is no additional context for these classifications, and the expertise involved in categorizing these properties potentially encodes real-life inequities that must be acknowledged.
### Sale Price
We begin by examining a [raincloud plot](https://micahallen.org/2018/03/15/introducing-raincloud-plots/amp/?__twitter_impression=true) (a combination of a KDE, a histogram, a strip plot, and a box plot) of our target variable `SalePrice`. At the same time, we also take a look at some descriptive statistics of this variable.
```
fig, axs = plt.subplots(nrows=2)
sns.distplot(
training_data['SalePrice'],
ax=axs[0]
)
sns.stripplot(
training_data['SalePrice'],
jitter=0.4,
size=3,
ax=axs[1],
alpha=0.3
)
sns.boxplot(
training_data['SalePrice'],
width=0.3,
ax=axs[1],
showfliers=False,
)
# Align axes
spacer = np.max(training_data['SalePrice']) * 0.05
xmin = np.min(training_data['SalePrice']) - spacer
xmax = np.max(training_data['SalePrice']) + spacer
axs[0].set_xlim((xmin, xmax))
axs[1].set_xlim((xmin, xmax))
# Remove some axis text
axs[0].xaxis.set_visible(False)
axs[0].yaxis.set_visible(False)
axs[1].yaxis.set_visible(False)
# Put the two plots together
plt.subplots_adjust(hspace=0)
# Adjust boxplot fill to be white
axs[1].artists[0].set_facecolor('white')
training_data['SalePrice'].describe()
```
## Question 1 <a name="q1"></a>
To check your understanding of the graph and summary statistics above, answer the following `True` or `False` questions:
1. The distribution of `SalePrice` in the training set is left-skew.
1. The mean of `SalePrice` in the training set is greater than the median.
1. At least 25% of the houses in the training set sold for more than \$200,000.00.
*The provided tests for this question do not confirm that you have answered correctly; only that you have assigned each variable to `True` or `False`.*
<!--
BEGIN QUESTION
name: q1
points: 3
-->
```
# These should be True or False
q1statement1 = ...
q1statement2 = ...
q1statement3 = ...
ok.grade("q1");
```
### SalePrice vs Gr_Liv_Area
Next, we visualize the association between `SalePrice` and `Gr_Liv_Area`. The `codebook.txt` file tells us that `Gr_Liv_Area` measures "above grade (ground) living area square feet."
This variable represents the square footage of the house excluding anything underground. Some additional research (into real estate conventions) reveals that this value also excludes the garage space.
```
sns.jointplot(
x='Gr_Liv_Area',
y='SalePrice',
data=training_data,
stat_func=None,
kind="reg",
ratio=4,
space=0,
scatter_kws={
's': 3,
'alpha': 0.25
},
line_kws={
'color': 'black'
}
);
```
There's certainly an association, and perhaps it's linear, but the spread is wider at larger values of both variables. Also, there are two particularly suspicious houses above 5000 square feet that look too inexpensive for their size.
## Question 2 <a name="q2"></a>
What are the Parcel Indentification Numbers for the two houses with `Gr_Liv_Area` greater than 5000 sqft?
*The provided tests for this question do not confirm that you have answered correctly; only that you have assigned `q2house1` and `q2house2` to two integers that are in the range of PID values.*
<!--
BEGIN QUESTION
name: q2
points: 2
-->
```
# Hint: You can answer this question in one line
...
ok.grade("q2");
```
## Question 3 <a name="q3"></a>
The codebook tells us how to manually inspect the houses using an online database called Beacon. These two houses are true outliers in this data set: they aren't the same time of entity as the rest. They were partial sales, priced far below market value. If you would like to inspect the valuations, follow the directions at the bottom of the codebook to access Beacon and look up houses by PID.
For this assignment, we will remove these outliers from the data. Write a function `remove_outliers` that removes outliers from a data set based off a threshold value of a variable. For example, `remove_outliers(training_data, 'Gr_Liv_Area', upper=5000)` should return a data frame with only observations that satisfy `Gr_Liv_Area` less than 5000.
*The provided tests check that training_data was updated correctly, so that future analyses are not corrupted by a mistake. However, the provided tests do not check that you have implemented remove_outliers correctly so that it works with any data, variable, lower, and upper bound.*
<!--
BEGIN QUESTION
name: q3
points: 1
-->
```
def remove_outliers(data, variable, lower=-np.inf, upper=np.inf):
"""
Input:
data (data frame): the table to be filtered
variable (string): the column with numerical outliers
lower (numeric): observations with values lower than or equal to this will be removed
upper (numeric): observations with values higher than or equal to this will be removed
Output:
a winsorized data frame with outliers removed
Note: This function should not change mutate the contents of data.
"""
...
training_data = remove_outliers(training_data, 'Gr_Liv_Area', upper=5000)
ok.grade("q3");
```
# Part 2: Feature Engineering
In this section, we will create a new feature out of existing ones through a simple data transformation. The goal is to implement broader domain knowledge we have about the data in order to aid our calculations.
As an example, we can look to an instance of feature engineering in the CCAO’s dataset, `Building Square Feet`. Based on real estate norms, home appraisals can only consider the square footage of living space, which means that garage and basement sizes cannot be included in a home’s total square footage. This is reflected in the CCAO’s dataset: The description for the feature `Garage Area 1` mentions that if Garage 1 is physically included within the building’s area, the area of the garage must be subtracted from the building’s total square footage, `Building Square Feet`.
## HCE: Question 8
Given that this data was used by the Cook County Assessor's Office to generate the model that they employ today, name two columns that have likely been added by the data scientists. Why do you think these transformations were created?
*Hint:* Look for columns that have been affected by mathematical transformations.
`TODO`: *Write your answer here*
### Bathrooms
Let's create a groundbreaking new feature. Due to recent advances in Universal WC Enumeration Theory, we now know that Total Bathrooms can be calculated as:
$$ \text{TotalBathrooms}=(\text{BsmtFullBath} + \text{FullBath}) + \dfrac{1}{2}(\text{BsmtHalfBath} + \text{HalfBath})$$
The actual proof is beyond the scope of this class, but we will use the result in our model.
## Question 4 <a name="q4"></a>
Write a function `add_total_bathrooms(data)` that returns a copy of `data` with an additional column called `TotalBathrooms` computed by the formula above. **Treat missing values as zeros**. Remember that you can make use of vectorized code here; you shouldn't need any `for` statements.
*The provided tests check that you answered correctly, so that future analyses are not corrupted by a mistake.*
<!--
BEGIN QUESTION
name: q4
points: 1
-->
```
def add_total_bathrooms(data):
"""
Input:
data (data frame): a data frame containing at least 4 numeric columns
Bsmt_Full_Bath, Full_Bath, Bsmt_Half_Bath, and Half_Bath
"""
with_bathrooms = data.copy()
bath_vars = ['Bsmt_Full_Bath', 'Full_Bath', 'Bsmt_Half_Bath', 'Half_Bath']
weights = pd.Series([1, 1, 0.5, 0.5], index=bath_vars)
...
return with_bathrooms
training_data = add_total_bathrooms(training_data)
ok.grade("q4");
```
## Question 5 <a name="q5"></a>
Create a visualization that clearly and succintly shows that `TotalBathrooms` is associated with `SalePrice`. Your visualization should avoid overplotting.
<!--
BEGIN QUESTION
name: q5
points: 2
manual: True
format: image
-->
<!-- EXPORT TO PDF format:image -->
```
...
```
# Part 3: Modeling
We've reached the point where we can specify a model. But first, we will load a fresh copy of the data, just in case our code above produced any undesired side-effects. Run the cell below to store a fresh copy of the data from `ccao_train.csv` in a dataframe named `full_data`. We will also store the number of rows in `full_data` in the variable `full_data_len`.
```
# Load a fresh copy of the data and get its length
full_data = pd.read_csv("ccao_train.csv")
full_data_len = len(full_data)
full_data.head()
```
## Question 6 <a name="q6"></a>
Now, let's split the data set into a training set and test set. We will use the training set to fit our model's parameters, and we will use the test set to estimate how well our model will perform on unseen data drawn from the same distribution. If we used all the data to fit our model, we would not have a way to estimate model performance on unseen data.
"Don't we already have a test set in `ccao_test.csv`?" you might wonder. The sale prices for `ccao_test.csv` aren't provided, so we're constructing our own test set for which we know the outputs.
In the cell below, split the data in `full_data` into two DataFrames named `train` and `test`. Let `train` contain 80% of the data, and let `test` contain the remaining 20% of the data.
To do this, first create two NumPy arrays named `train_indices` and `test_indices`. `train_indices` should contain a *random* 80% of the indices in `full_data`, and `test_indices` should contain the remaining 20% of the indices. Then, use these arrays to index into `full_data` to create your final `train` and `test` DataFrames.
*The provided tests check that you not only answered correctly, but ended up with the exact same train/test split as our reference implementation. Later testing is easier this way.*
<!--
BEGIN QUESTION
name: q6
points: 2
-->
```
# This makes the train-test split in this section reproducible across different runs
# of the notebook. You do not need this line to run train_test_split in general
np.random.seed(1337)
shuffled_indices = np.random.permutation(full_data_len)
# Set train_indices to the first 80% of shuffled_indices and and test_indices to the rest.
train_indices = ...
test_indices = ...
# Create train and test` by indexing into `full_data` using
# `train_indices` and `test_indices`
train = ...
test = ...
ok.grade("q6");
```
### Reusable Pipeline
Throughout this assignment, you should notice that your data flows through a single processing pipeline several times. From a software engineering perspective, it's ideal to define functions and methods that can apply the pipeline to any dataset. We will now encapsulate our entire pipeline into a single function `process_data_gm`. gm is shorthand for "guided model".
Additionally, creating reproducible work is an important part of data science. When you leave a project, readability and transferability of your code is necessary for future work on the project by other data scientists. Therefore, it’s better to use abstract identifiers to navigate code. For example, in this notebook, we refer to columns by their names - that way, if a new dataset comes in with different column indices, we’ll be able to know which columns to target when adapting code.
In the CCAO's case, for example, they felt that making their model into a reproducible and easily reusable pipeline was an essential part of their mission as a public office. In this case, this allows citizens and other entities with sufficient technical skills to understand and even critique their work openly.
```
def select_columns(data, *columns):
"""Select only columns passed as arguments."""
return data.loc[:, columns]
def process_data_gm(data):
"""Process the data for a guided model."""
data = remove_outliers(data, 'Gr_Liv_Area', upper=5000)
# Transform Data, Select Features
data = add_total_bathrooms(data)
data = select_columns(data,
'SalePrice',
'Gr_Liv_Area',
'Garage_Area',
'TotalBathrooms',
)
# Return predictors and response variables separately
X = data.drop(['SalePrice'], axis = 1)
y = data.loc[:, 'SalePrice']
return X, y
```
Now, we can use `process_data_gm` to clean our data, select features, and add our `TotalBathrooms` feature all in one step! This function also splits our data into `X`, a matrix of features, and `y`, a vector of sale prices.
Run the cell below to feed our training and test data through the pipeline, generating `X_train`, `y_train`, `X_test`, and `y_test`.
```
# Pre-process our training and test data in exactly the same way
# Our functions make this very easy!
X_train, y_train = process_data_gm(train)
X_test, y_test = process_data_gm(test)
```
### Fitting Our First Model
We are finally going to fit a model! The model we will fit can be written as follows:
$$\text{SalePrice} = \theta_0 + \theta_1 \cdot \text{Gr_Liv_Area} + \theta_2 \cdot \text{Garage_Area} + \theta_3 \cdot \text{TotalBathrooms}$$
In vector notation, the same equation would be written:
$$y = \vec\theta \cdot \vec{x}$$
where $y$ is the SalePrice, $\vec\theta$ is a vector of all fitted weights, and $\vec{x}$ contains a 1 for the bias followed by each of the feature values.
**Note:** Notice that all of our variables are continuous, except for `TotalBathrooms`, which takes on discrete ordered values (0, 0.5, 1, 1.5, ...). In this homework, we'll treat `TotalBathrooms` as a continuous quantitative variable in our model, but this might not be the best choice. The next homework may revisit the issue.
## Question 7a <a name="q7a"></a>
We will use a [`sklearn.linear_model.LinearRegression`](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LinearRegression.html) object as our linear model. In the cell below, create a `LinearRegression` object and name it `linear_model`.
**Hint:** See the `fit_intercept` parameter and make sure it is set appropriately. The intercept of our model corresponds to $\theta_0$ in the equation above.
*The provided tests check that you answered correctly, so that future analyses are not corrupted by a mistake.*
<!--
BEGIN QUESTION
name: q7a
points: 1
-->
```
from sklearn import linear_model as lm
linear_model = ...
ok.grade("q7a");
```
## Question 7b <a name="q7b"></a>
Now, remove the commenting and fill in the ellipses `...` below with `X_train`, `y_train`, `X_test`, or `y_test`.
With the ellipses filled in correctly, the code below should fit our linear model to the training data and generate the predicted sale prices for both the training and test datasets.
*The provided tests check that you answered correctly, so that future analyses are not corrupted by a mistake.*
<!--
BEGIN QUESTION
name: q7b
points: 2
-->
```
# Uncomment the lines below and fill in the ... with X_train, y_train, X_test, or y_test.
# linear_model.fit(..., ...)
# y_fitted = linear_model.predict(...)
# y_predicted = linear_model.predict(...)
ok.grade("q7b");
```
## Question 8a <a name="q8a"></a>
Is our linear model any good at predicting house prices? Let's measure the quality of our model by calculating the Root-Mean-Square Error (RMSE) between our predicted house prices and the true prices stored in `SalePrice`.
$$\text{RMSE} = \sqrt{\dfrac{\sum_{\text{houses in test set}}(\text{actual price of house} - \text{predicted price of house})^2}{\text{# of houses in data set}}}$$
In the cell below, write a function named `rmse` that calculates the RMSE of a model.
**Hint:** Make sure you are taking advantage of vectorized code. This question can be answered without any `for` statements.
*The provided tests check that you answered correctly, so that future analyses are not corrupted by a mistake.*
<!--
BEGIN QUESTION
name: q8a
points: 1
-->
```
def rmse(actual, predicted):
"""
Calculates RMSE from actual and predicted values
Input:
actual (1D array): vector of actual values
predicted (1D array): vector of predicted/fitted values
Output:
a float, the root-mean square error
"""
...
ok.grade("q8a");
```
## Question 8b <a name="q8b"></a>
Now use your `rmse` function to calculate the training error and test error in the cell below.
*The provided tests for this question do not confirm that you have answered correctly; only that you have assigned each variable to a non-negative number.*
<!--
BEGIN QUESTION
name: q8b
points: 1
-->
```
training_error = ...
test_error = ...
(training_error, test_error)
ok.grade("q8b");
```
## Question 8c <a name="q8c"></a>
How much does including `TotalBathrooms` as a predictor reduce the RMSE of the model on the test set? That is, what's the difference between the RSME of a model that only includes `Gr_Liv_Area` and `Garage_Area` versus one that includes all three predictors?
*The provided tests for this question do not confirm that you have answered correctly; only that you have assigned the answer variable to a non-negative number.*
<!--
BEGIN QUESTION
name: q8c
points: 2
-->
```
test_error_difference = test_error_no_bath - test_error
test_error_difference
ok.grade("q8c");
```
### Residual Plots
One way of understanding the performance (and appropriateness) of a model is through a residual plot. Run the cell below to plot the actual sale prices against the residuals of the model for the test data.
```
residuals = y_test - y_predicted
ax = sns.regplot(y_test, residuals)
ax.set_xlabel('Sale Price (Test Data)')
ax.set_ylabel('Residuals (Actual Price - Predicted Price)')
ax.set_title("Residuals vs. Sale Price on Test Data");
```
Ideally, we would see a horizontal line of points at 0 (perfect prediction!). The next best thing would be a homogenous set of points centered at 0.
But alas, our simple model is probably too simple. The most expensive homes are systematically more expensive than our prediction.
## Question 8d <a name="q8c"></a>
What changes could you make to your linear model to improve its accuracy and lower the test error? Suggest at least two things you could try in the cell below, and carefully explain how each change could potentially improve your model's accuracy.
<!--
BEGIN QUESTION
name: q8d
points: 2
manual: True
-->
<!-- EXPORT TO PDF -->
*Write your answer here, replacing this text.*
## HCE: Question 9
In the context of assessing houses, what does error mean for an individual homeowner? How does it affect them in terms of property taxes? Do these effects change your tolerance for error in modeling?
`TODO`: *Write your answer here*
Congratulations! You’ve built a simple home assessment model. With this simple model, it's clear that the main variables in the model are the Living Area, Garage Area, and TotalBathrooms. However, as you might imagine, the CCAO's model is much more complex. In the next homework, we'll discuss the CCAO's model with more depth as we continue to improve the simple model we built here.
In the meantime, consider the implications of your work so far: We know that housing assessments affect property taxes. Fairness in the CCAO’s work, then, is accurately assessing the value of homeowners’ properties, where accuracy is a reflection of whether a homeowner pays taxes comparable to homeowners with similar houses. These assessments - and even what’s considered accurate - are a product of choices made at every level of the assessment process, from data collection to modeling. And the humans who make these choices - real estate agents, data scientists, county assessors, and, in this homework and the next, you - must understand the inputs to and consider the impacts of these decisions.
So when we conceptualize accurate assessments, we must think beyond eliminating bias. Bias is impossible to eliminate, given the indispensability of human involvement and historical/institutional contexts. We must instead consider the long-term, real-world consequences for individuals and communities in specific local and historical contexts. Only in this way can we aspire towards making fair and equitable assessment possible.
# Submit
Make sure you have run all cells in your notebook in order before running the cell below, so that all images/graphs appear in the output.
**Please save before submitting!**
<!-- EXPECT 2 EXPORTED QUESTIONS -->
```
# Save your notebook first, then run this cell to submit.
import jassign.to_pdf
jassign.to_pdf.generate_pdf('hw5.ipynb', 'hw5.pdf')
ok.submit()
```
| github_jupyter |
```
# Importing the libraries
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# Import the dataset for mobility
mobility = pd.read_csv("../data/usitc/border_crossing_entry_data_trade.csv")
# Include only mobility on the US-MX border
borderMX = mobility[(mobility['Border']=="US-Mexico Border")]
borderMX.head()
borderMX["Measure"].unique()
borderMX["Measure"].value_counts()
# Extract year and consolidate year
import datetime
borderMX['year']= pd.DatetimeIndex(borderMX['Date']).year
## Encoding the categorial variable
var = borderMX[["Measure","Port Code"]].copy()
X1 = var.iloc[:,[0,1]].values
print(X1[-10:])
# Encoding categorical data
# Encoding the independent variable
from sklearn.preprocessing import LabelEncoder, OneHotEncoder
labelencoder = LabelEncoder()
X1[:, 0] = labelencoder.fit_transform(X1[:, 0])
onehotencoder = OneHotEncoder(categorical_features = [0])
X1 = onehotencoder.fit_transform(X1).toarray()
print(X1)
X1[-10:]
passen = X1[:,2]
trucks = X1[:,11]
print(passen)
print(trucks)
```
## Now preparing for Naive Bayes
```
df = borderMX[["year","State","Port Code","Value"]].copy()
X2 = df.iloc[:,[0,2,3]].values
print(X2)
# PEDESTRIANS FIRST
print(X2.shape)
# Create exports matrix for trade data
ex = borderMX.iloc[:,[8]].values
X2 = np.append(arr=X2, values= ex.astype(float),axis=1)
# Create imports matrix for trade data
im = borderMX.iloc[:,[7]].values
X2 = np.append(arr=X2, values= im.astype(float),axis=1)
X2[:10]
```
## Establishing Variables
```
# Re-establishing the X matrix:
X = X2[:,[4,1]]
print(X)
y = passen
print(passen)
```
## Naive Bayes Here
```
# Splitting the dataset into the Training set and Test set
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state = 0)
# Feature Scaling
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
# Fitting Naive Bayes to the Training set
from sklearn.naive_bayes import GaussianNB
classifier = GaussianNB()
classifier.fit(X_train, y_train)
# Predicting the Test set results
y_pred = classifier.predict(X_test)
# Making the Confusion Matrix
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, y_pred)
print(cm)
# Visualizing the Training set results
from matplotlib.colors import ListedColormap
X_set, y_set = X_train, y_train
X1, X2 = np.meshgrid(np.arange(start = X_set[:, 0].min() - 1, stop = X_set[:, 0].max() + 1, step = 0.01),
np.arange(start = X_set[:, 1].min() - 1, stop = X_set[:, 1].max() + 1, step = 0.01))
plt.contourf(X1, X2, classifier.predict(np.array([X1.ravel(), X2.ravel()]).T).reshape(X1.shape),
alpha = 0.75, cmap = ListedColormap(('aqua', 'navy')))
plt.xlim(X1.min(), X1.max())
plt.ylim(X2.min(), X2.max())
for i, j in enumerate(np.unique(y_set)):
plt.scatter(X_set[y_set == j, 0], X_set[y_set == j, 1],
c = ListedColormap(('aqua', 'navy'))(i), label = j)
plt.title('Naive Bayes (Training set): Pedestrians')
plt.xlabel('Imports')
plt.ylabel('Port Code')
plt.legend()
plt.show()
# Visualising the Test set results
from matplotlib.colors import ListedColormap
X_set, y_set = X_test, y_test
X1, X2 = np.meshgrid(np.arange(start = X_set[:, 0].min() - 1, stop = X_set[:, 0].max() + 1, step = 0.01),
np.arange(start = X_set[:, 1].min() - 1, stop = X_set[:, 1].max() + 1, step = 0.01))
plt.contourf(X1, X2, classifier.predict(np.array([X1.ravel(), X2.ravel()]).T).reshape(X1.shape),
alpha = 0.75, cmap = ListedColormap(('aqua', 'navy')))
plt.xlim(X1.min(), X1.max())
plt.ylim(X2.min(), X2.max())
for i, j in enumerate(np.unique(y_set)):
plt.scatter(X_set[y_set == j, 0], X_set[y_set == j, 1],
c = ListedColormap(('aqua', 'navy'))(i), label = j)
plt.title('Naive Bayes (Test set) - Pedestrians Entrance')
plt.xlabel('Imports')
plt.ylabel('Port Code')
plt.legend()
plt.show()
```
## Logistic Regression for Pedestrians
```
# Fitting Logistic Regression to the Training set
from sklearn.linear_model import LogisticRegression
classifier = LogisticRegression(random_state = 0)
classifier.fit(X_train, y_train)
# Predicting the Test set results
y_pred = classifier.predict(X_test)
# Making the Confusion Matrix
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, y_pred)
print(cm)
# Visualising the Training set results
from matplotlib.colors import ListedColormap
X_set, y_set = X_train, y_train
X1, X2 = np.meshgrid(np.arange(start = X_set[:, 0].min() - 1, stop = X_set[:, 0].max() + 1, step = 0.01),
np.arange(start = X_set[:, 1].min() - 1, stop = X_set[:, 1].max() + 1, step = 0.01))
plt.contourf(X1, X2, classifier.predict(np.array([X1.ravel(), X2.ravel()]).T).reshape(X1.shape),
alpha = 0.75, cmap = ListedColormap(('aqua', 'navy')))
plt.xlim(X1.min(), X1.max())
plt.ylim(X2.min(), X2.max())
for i, j in enumerate(np.unique(y_set)):
plt.scatter(X_set[y_set == j, 0], X_set[y_set == j, 1],
c = ListedColormap(('aqua', 'navy'))(i), label = j)
plt.title('Logistic Regression (Training set) - Pedestrians')
plt.xlabel('Imports')
plt.ylabel('Port Code')
plt.legend()
plt.show()
# Visualising the Test set results
from matplotlib.colors import ListedColormap
X_set, y_set = X_test, y_test
X1, X2 = np.meshgrid(np.arange(start = X_set[:, 0].min() - 1, stop = X_set[:, 0].max() + 1, step = 0.01),
np.arange(start = X_set[:, 1].min() - 1, stop = X_set[:, 1].max() + 1, step = 0.01))
plt.contourf(X1, X2, classifier.predict(np.array([X1.ravel(), X2.ravel()]).T).reshape(X1.shape),
alpha = 0.75, cmap = ListedColormap(('aqua', 'navy')))
plt.xlim(X1.min(), X1.max())
plt.ylim(X2.min(), X2.max())
for i, j in enumerate(np.unique(y_set)):
plt.scatter(X_set[y_set == j, 0], X_set[y_set == j, 1],
c = ListedColormap(('aqua', 'navy'))(i), label = j)
plt.title('Logistic Regression (Test set) - Pedestrians')
plt.xlabel('Imports')
plt.ylabel('Port Code')
plt.legend()
plt.show()
```
## Starting Naive Bayes for Trucks
```
y = trucks
# Splitting the dataset into the Training set and Test set
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state = 0)
# Feature Scaling
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
# Fitting Naive Bayes to the Training set
from sklearn.naive_bayes import GaussianNB
classifier = GaussianNB()
classifier.fit(X_train, y_train)
# Predicting the Test set results
y_pred = classifier.predict(X_test)
# Making the Confusion Matrix
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, y_pred)
print(cm)
# Visualizing the Training set results
from matplotlib.colors import ListedColormap
X_set, y_set = X_train, y_train
X1, X2 = np.meshgrid(np.arange(start = X_set[:, 0].min() - 1, stop = X_set[:, 0].max() + 1, step = 0.01),
np.arange(start = X_set[:, 1].min() - 1, stop = X_set[:, 1].max() + 1, step = 0.01))
plt.contourf(X1, X2, classifier.predict(np.array([X1.ravel(), X2.ravel()]).T).reshape(X1.shape),
alpha = 0.75, cmap = ListedColormap(('aqua', 'navy')))
plt.xlim(X1.min(), X1.max())
plt.ylim(X2.min(), X2.max())
for i, j in enumerate(np.unique(y_set)):
plt.scatter(X_set[y_set == j, 0], X_set[y_set == j, 1],
c = ListedColormap(('aqua', 'navy'))(i), label = j)
plt.title('Naive Bayes (Training set): Trucks')
plt.xlabel('Imports')
plt.ylabel('Port Code')
plt.legend()
plt.show()
# Visualising the Test set results
from matplotlib.colors import ListedColormap
X_set, y_set = X_test, y_test
X1, X2 = np.meshgrid(np.arange(start = X_set[:, 0].min() - 1, stop = X_set[:, 0].max() + 1, step = 0.01),
np.arange(start = X_set[:, 1].min() - 1, stop = X_set[:, 1].max() + 1, step = 0.01))
plt.contourf(X1, X2, classifier.predict(np.array([X1.ravel(), X2.ravel()]).T).reshape(X1.shape),
alpha = 0.75, cmap = ListedColormap(('aqua', 'navy')))
plt.xlim(X1.min(), X1.max())
plt.ylim(X2.min(), X2.max())
for i, j in enumerate(np.unique(y_set)):
plt.scatter(X_set[y_set == j, 0], X_set[y_set == j, 1],
c = ListedColormap(('aqua', 'navy'))(i), label = j)
plt.title('Naive Bayes (Test set) - Trucks')
plt.xlabel('Imports')
plt.ylabel('Port Code')
plt.legend()
plt.show()
```
| github_jupyter |
```
%matplotlib inline
```
PyTorch에서 추론(inference)을 위해 모델 저장하기 & 불러오기
================================================================
PyTorch에서는 추론(inference)을 위해 모델을 저장하고 불러오는데 2가지 접근법이
있습니다. 첫번째는 ``state_dict`` 를 저장하고 불러오는 것이고, 두번째는 전체
모델을 저장하는 것입니다.
개요
------------
torch.save() 함수를 사용하여 모델의 ``state_dict`` 를 저장하면 이후에 모델을
불러올 때 유연함을 크게 살릴 수 있습니다. 학습된 모델의 매개변수(parameter)만을
저장하면되므로 모델 저장 시에 권장하는 방법입니다. 모델 전체를 저장하고 불러올
때에는 Python의 `pickle <https://docs.python.org/3/library/pickle.html>`__ 모듈을
사용하여 전체 모듈을 저장합니다. 이 방식은 직관적인 문법을 사용하며 코드의 양도
적습니다. 이 방식의 단점은 직렬화(serialized)된 데이터가 모델을 저장할 때 사용한
특정 클래스 및 디렉토리 구조에 종속(bind)된다는 것입니다. 그 이유는 pickle이
모델 클래스 자체를 저장하지 않기 때문입니다. 오히려 불러올 때 사용되는 클래스가
포함된 파일의 경로를 저장합니다. 이 때문에 작성한 코드가 다른 프로젝트에서
사용되거나 리팩토링을 거치는 등의 과정에서 동작하지 않을 수 있습니다. 이 레시피에서는
추론을 위해 모델을 저장하고 불러오는 두 가지 방법 모두를 살펴보겠습니다.
설정
----------
시작하기 전에 ``torch`` 가 없다면 설치해야 합니다.
::
pip install torch
단계(Steps)
------------
1. 데이터 불러올 때 필요한 라이브러리들 불러오기
2. 신경망을 구성하고 초기화하기
3. 옵티마이저 초기화하기
4. ``state_dict`` 을 통해 모델을 저장하고 불러오기
5. 전체 모델을 저장하고 불러오기
1. 데이터 불러올 때 필요한 라이브러리들 불러오기
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
이 레시피에서는 ``torch`` 와 여기 포함된 ``torch.nn`` 과 ``torch.optim` 을
사용하겠습니다.
```
import torch
import torch.nn as nn
import torch.optim as optim
```
2. 신경망을 구성하고 초기화하기
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
예를 들기 위해, 이미지를 학습하는 신경망을 만들어보겠습니다. 더 자세한 내용은
신경망 구성하기 레시피를 참고해주세요.
```
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(3, 6, 5)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(6, 16, 5)
self.fc1 = nn.Linear(16 * 5 * 5, 120)
self.fc2 = nn.Linear(120, 84)
self.fc3 = nn.Linear(84, 10)
def forward(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
x = x.view(-1, 16 * 5 * 5)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.fc3(x)
return x
net = Net()
print(net)
```
3. 옵티마이저 초기화하기
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
모멘텀(momentum)을 갖는 SGD를 사용하겠습니다.
```
optimizer = optim.SGD(net.parameters(), lr=0.001, momentum=0.9)
```
4. ``state_dict`` 을 통해 모델을 저장하고 불러오기
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
이제 ``state_dict`` 만 사용하여 모델을 저장하고 불러와보겠습니다.
```
# 경로 지정
PATH = "state_dict_model.pt"
# 저장하기
torch.save(net.state_dict(), PATH)
# 불러오기
model = Net()
model.load_state_dict(torch.load(PATH))
model.eval()
```
PyTorch에서는 모델을 저장할 때 ``.pt`` 또는 ``.pth`` 확장자를 사용하는 것이
일반적인 규칙입니다.
``load_state_dict()`` 함수는 저장된 객체의 경로가 아닌, 사전 객체를 사용합니다.
즉, 저장된 state_dict를 ``load_state_dict()`` 함수에 전달하기 전에 반드시
역직렬화(deserialize)를 해야 합니다. 예를 들어, ``model.load_state_dict(PATH)``
와 같이 사용할 수 없습니다.
또한, 추론을 실행하기 전에 ``model.eval()`` 을 호출하여 드롭아웃(dropout)과
배치 정규화 층(batch normalization layers)을 평가(evaluation) 모드로 바꿔야한다는
것을 기억하세요. 이것을 빼먹으면 일관성 없는 추론 결과를 얻게 됩니다.
5. 전체 모델을 저장하고 불러오기
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
이제 전체 모델에 대해서 똑같이 해보겠습니다.
```
# 경로 지정
PATH = "entire_model.pt"
# 저장하기
torch.save(net, PATH)
# 불러오기
model = torch.load(PATH)
model.eval()
```
여기서도 또한 model.eval()을 실행하여 드롭아웃(dropout)과 배치 정규화 층
(batch normalization layers)을 평가(evaluation) 모드로 바꿔야한다는
것을 기억하세요.
축하합니다! 지금까지 PyTorch에서 추론을위한 모델을 성공적으로 저장하고 불러왔습니다.
더 알아보기
-------------
다른 레시피를 둘러보고 계속 배워보세요:
- :doc:`/recipes/recipes/saving_and_loading_a_general_checkpoint`
- :doc:`/recipes/recipes/saving_multiple_models_in_one_file`
| github_jupyter |
# t-SNE plots for ProtoCLR & ProtoNet
```
import h5py
import numpy as np
from matplotlib import pyplot as plt
import seaborn as sns
import random
random.seed(0)
sns.set_style("white")
from sklearn.manifold import TSNE
```
## 1: t-SNE plots for ProtoCLR
```
f_protoclr_train = h5py.File('featuresProtoCLR_mini-ImageNet_train.hdf5', 'r')
f_protoclr_test = h5py.File('featuresProtoCLR_mini-ImageNet_test.hdf5', 'r')
X_protoclr_5_train = f_protoclr_train['all_feats']
y_protoclr_5_train = f_protoclr_train['all_labels']
X_protoclr_5_test = f_protoclr_test['all_feats']
y_protoclr_5_test = f_protoclr_test['all_labels']
train_protoclr_idx = random.sample(range(len(y_protoclr_5_train)),500)
train_protoclr_idx.sort()
X_protoclr_5_train = X_protoclr_5_train[train_protoclr_idx]
y_protoclr_5_train = y_protoclr_5_train[train_protoclr_idx]
test_protoclr_idx = random.sample(range(len(y_protoclr_5_test)),500)
test_protoclr_idx.sort()
X_protoclr_5_test = X_protoclr_5_test[test_protoclr_idx]
y_protoclr_5_test = y_protoclr_5_test[test_protoclr_idx]
tsne = TSNE(n_components=2, random_state=0)
X_protoclr_2d_train = tsne.fit_transform(X_protoclr_5_train)
tsne = TSNE(n_components=2, random_state=0)
X_protoclr_2d_test = tsne.fit_transform(X_protoclr_5_test)
def plot2d(X_2d, y,colors=None, title=None, save_as=None):
target_ids = range(5)
labels_train = ["Class {}".format(i) for i in range(1,6)]
from matplotlib import pyplot as plt
fig = plt.figure(figsize=(6, 5))
ax = fig.add_subplot(111)
#colors = 'r', 'g', 'b', 'c', 'm'#, 'y', 'k', 'w', 'orange', 'purple'
markers = "o","D","*","^","P"
#print(len(colors))
for i, c, label, marker in zip(target_ids, colors, labels_train, markers):
ax.scatter(X_2d[y == i, 0],
X_2d[y == i, 1],
c=c, label=label, marker=marker)
#plt.legend()
ax.set_xticks([])
ax.set_yticks([])
sns.despine(left=True, bottom=True)
#plt.legend(loc='lower right')
if save_as:
plt.savefig(save_as, bbox_inches = 'tight',
pad_inches = 0)
if title:
plt.title(title)
plt.show()
colors = plt.rcParams['axes.prop_cycle'].by_key()['color']
plot2d(X_2d=X_protoclr_2d_train,
y=y_protoclr_5_train,
colors=colors[:5],
title="ProtoCLR train classes",
save_as='protoclr_train_set_tsne.pdf')
plot2d(X_2d=X_protoclr_2d_test,
y=y_protoclr_5_test,
colors=colors[5:],
title="ProtoCLR test classes",
save_as='protoclr_test_set_tsne.pdf')
```
## 2: t-SNE plots for ProtoNet
```
f_protonet_train = h5py.File('featuresProtoNet_mini-ImageNet_train.hdf5', 'r')
f_protonet_test = h5py.File('featuresProtoNet_mini-ImageNet_test.hdf5', 'r')
X_protonet_5_train = f_protonet_train['all_feats']
y_protonet_5_train = f_protonet_train['all_labels']
X_protonet_5_test = f_protonet_test['all_feats']
y_protonet_5_test = f_protonet_test['all_labels']
train_protonet_idx = random.sample(range(len(y_protonet_5_train)),500)
train_protonet_idx.sort()
X_protonet_5_train = X_protonet_5_train[train_protonet_idx]
y_protonet_5_train = y_protonet_5_train[train_protonet_idx]
test_protonet_idx = random.sample(range(len(y_protonet_5_test)),500)
test_protonet_idx.sort()
X_protonet_5_test = X_protonet_5_test[test_protonet_idx]
y_protonet_5_test = y_protonet_5_test[test_protonet_idx]
tsne = TSNE(n_components=2, random_state=0)
X_protonet_2d_train = tsne.fit_transform(X_protonet_5_train)
tsne = TSNE(n_components=2, random_state=0)
X_protonet_2d_test = tsne.fit_transform(X_protonet_5_test)
plot2d(X_2d=X_protonet_2d_train,
y=y_protonet_5_train,
colors=colors[:5],
title="ProtoNet train classes",
save_as='protonet_train_set_tsne.pdf')
plot2d(X_2d=X_protonet_2d_test,
y=y_protonet_5_test,
colors=colors[5:],
title="ProtoNet test classes",
save_as='protonet_test_set_tsne.pdf')
```
| github_jupyter |
# Data description:
This dataset describes the monthly number of sales of shampoo over a 3 year period. The units are a sales count and there are 36 observations. The original dataset is credited to Makridakis, Wheelwright and Hyndman (1998).
# Workflow:
- Load the Time Series (TS) by Pandas Library
- Prepare the data, i.e. convert the problem to a supervised ML problem
- Build and evaluate the RNN model:
- Fit the best RNN model
- Evaluate model by in-sample prediction: Calculate RMSE
- Forecast the future trend: Out-of-sample prediction
Note: For data exploration of this TS, please refer to the notebook of my alternative solution with "Seasonal ARIMA model"
```
import keras
import sklearn
import tensorflow as tf
import numpy as np
from scipy import stats
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn import metrics
from sklearn import preprocessing
import random as rn
import math
%matplotlib inline
from keras import backend as K
session_conf = tf.ConfigProto(intra_op_parallelism_threads=5, inter_op_parallelism_threads=5)
sess = tf.Session(graph=tf.get_default_graph(), config=session_conf)
K.set_session(sess)
import warnings
warnings.filterwarnings("ignore")
# Load data using Series.from_csv
from pandas import Series
#TS = Series.from_csv('C:/Users/rhash/Documents/Datasets/Time Series analysis/daily-minimum-temperatures.csv', header=0)
# Load data using pandas.read_csv
# in case, specify your own date parsing function and use the date_parser argument
from pandas import read_csv
from pandas import datetime
#def parser(x):
# return datetime.strptime('190'+x, '%Y-%m')
TS = read_csv('C:/Users/rhash/Documents/Datasets/Time Series analysis/sales-of-shampoo.csv', header=0, parse_dates=[0], index_col=0, squeeze=True)
print(TS.head())
#TS=pd.to_numeric(TS, errors='coerce')
TS.dropna(inplace=True)
data=pd.DataFrame(TS.values)
# prepare the data (i.e. convert problem to a supervised ML problem)
def prepare_data(data, lags=1):
"""
Create lagged data from an input time series
"""
X, y = [], []
for row in range(len(data) - lags - 1):
a = data[row:(row + lags), 0]
X.append(a)
y.append(data[row + lags, 0])
return np.array(X), np.array(y)
# normalize the dataset
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler(feature_range=(0, 1))
dataset = scaler.fit_transform(data)
# split into train and test sets
train = dataset[0:24, :]
test = dataset[24:, :]
# LSTM RNN model: _________________________________________________________________
from keras.models import Sequential, Model
from keras.layers import Dense, LSTM, Dropout, average, Input, merge, concatenate
from keras.layers.merge import concatenate
from keras.regularizers import l2, l1
from keras.callbacks import EarlyStopping, ModelCheckpoint
from sklearn.utils.class_weight import compute_sample_weight
from keras.layers.normalization import BatchNormalization
np.random.seed(42)
rn.seed(42)
tf.set_random_seed(42)
# reshape into X=t and Y=t+1
lags = 3
X_train, y_train = prepare_data(train, lags)
X_test, y_test = prepare_data(test, lags)
# reshape input to be [samples, time steps, features]
X_train = np.reshape(X_train, (X_train.shape[0], 1, X_train.shape[1]))
X_test = np.reshape(X_test, (X_test.shape[0], 1, X_test.shape[1]))
# create and fit the LSTM network
mdl = Sequential()
#mdl.add(Dense(3, input_shape=(1, lags), activation='relu'))
mdl.add(LSTM(20, activation='relu'))
mdl.add(Dropout(0.3))
mdl.add(Dense(1))
mdl.compile(loss='mean_squared_error', optimizer='adam')
monitor=EarlyStopping(monitor='loss', min_delta=0.001, patience=1000, verbose=1, mode='auto')
checkpointer = ModelCheckpoint(filepath="shampoo_weights.hdf5", verbose=0, save_best_only=True) # save best model
history=mdl.fit(X_train, y_train, epochs=1000, batch_size=1, validation_data=(X_test, y_test), callbacks=[monitor, checkpointer], verbose=0)
mdl.load_weights('shampoo_weights.hdf5') # load weights from best model
# To measure RMSE and evaluate the RNN model:
from sklearn.metrics import mean_squared_error
# make predictions
train_predict = mdl.predict(X_train)
test_predict = mdl.predict(X_test)
# invert transformation
train_predict = scaler.inverse_transform(pd.DataFrame(train_predict))
y_train = scaler.inverse_transform(pd.DataFrame(y_train))
test_predict = scaler.inverse_transform(pd.DataFrame(test_predict))
y_test = scaler.inverse_transform(pd.DataFrame(y_test))
# calculate root mean squared error
train_score = math.sqrt(mean_squared_error(y_train, train_predict[:,0]))
print('Train Score: {:.2f} RMSE'.format(train_score))
test_score = math.sqrt(mean_squared_error(y_test, test_predict[:,0]))
print('Test Score: {:.2f} RMSE'.format(test_score))
# list all data in history
#print(history.history.keys())
# summarize history for loss
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# shift train predictions for plotting
train_predict_plot =np.full(data.shape, np.nan)
train_predict_plot[lags:len(train_predict)+lags, :] = train_predict
# shift test predictions for plotting
test_predict_plot =np.full(data.shape, np.nan)
test_predict_plot[len(train_predict) + (lags * 2)+1:len(data)-1, :] = test_predict
# plot observation and predictions
plt.figure(figsize=(8,6))
plt.plot(data, label='Observed', color='#006699');
plt.plot(train_predict_plot, label='Prediction for Train Set', color='#006699', alpha=0.5);
plt.plot(test_predict_plot, label='Prediction for Test Set', color='#ff0066');
plt.legend(loc='upper left')
plt.title('LSTM Recurrent Neural Net')
plt.show()
mse = mean_squared_error(y_test, test_predict[:,0])
plt.title('Prediction quality: {:.2f} MSE ({:.2f} RMSE)'.format(mse, math.sqrt(mse)))
plt.plot(y_test.reshape(-1, 1), label='Observed', color='#006699')
plt.plot(test_predict.reshape(-1, 1), label='Prediction', color='#ff0066')
plt.legend(loc='upper left');
plt.show()
```
| github_jupyter |
# Plagiarism Detection, Feature Engineering
In this project, you will be tasked with building a plagiarism detector that examines an answer text file and performs binary classification; labeling that file as either plagiarized or not, depending on how similar that text file is to a provided, source text.
Your first task will be to create some features that can then be used to train a classification model. This task will be broken down into a few discrete steps:
* Clean and pre-process the data.
* Define features for comparing the similarity of an answer text and a source text, and extract similarity features.
* Select "good" features, by analyzing the correlations between different features.
* Create train/test `.csv` files that hold the relevant features and class labels for train/test data points.
In the _next_ notebook, Notebook 3, you'll use the features and `.csv` files you create in _this_ notebook to train a binary classification model in a SageMaker notebook instance.
You'll be defining a few different similarity features, as outlined in [this paper](https://s3.amazonaws.com/video.udacity-data.com/topher/2019/January/5c412841_developing-a-corpus-of-plagiarised-short-answers/developing-a-corpus-of-plagiarised-short-answers.pdf), which should help you build a robust plagiarism detector!
To complete this notebook, you'll have to complete all given exercises and answer all the questions in this notebook.
> All your tasks will be clearly labeled **EXERCISE** and questions as **QUESTION**.
It will be up to you to decide on the features to include in your final training and test data.
---
## Read in the Data
The cell below will download the necessary, project data and extract the files into the folder `data/`.
This data is a slightly modified version of a dataset created by Paul Clough (Information Studies) and Mark Stevenson (Computer Science), at the University of Sheffield. You can read all about the data collection and corpus, at [their university webpage](https://ir.shef.ac.uk/cloughie/resources/plagiarism_corpus.html).
> **Citation for data**: Clough, P. and Stevenson, M. Developing A Corpus of Plagiarised Short Answers, Language Resources and Evaluation: Special Issue on Plagiarism and Authorship Analysis, In Press. [Download]
```
# NOTE:
# you only need to run this cell if you have not yet downloaded the data
# otherwise you may skip this cell or comment it out
!wget https://s3.amazonaws.com/video.udacity-data.com/topher/2019/January/5c4147f9_data/data.zip
!unzip data
# import libraries
import pandas as pd
import numpy as np
import os
```
This plagiarism dataset is made of multiple text files; each of these files has characteristics that are is summarized in a `.csv` file named `file_information.csv`, which we can read in using `pandas`.
```
csv_file = 'data/file_information.csv'
plagiarism_df = pd.read_csv(csv_file)
# print out the first few rows of data info
plagiarism_df.head()
```
## Types of Plagiarism
Each text file is associated with one **Task** (task A-E) and one **Category** of plagiarism, which you can see in the above DataFrame.
### Tasks, A-E
Each text file contains an answer to one short question; these questions are labeled as tasks A-E. For example, Task A asks the question: "What is inheritance in object oriented programming?"
### Categories of plagiarism
Each text file has an associated plagiarism label/category:
**1. Plagiarized categories: `cut`, `light`, and `heavy`.**
* These categories represent different levels of plagiarized answer texts. `cut` answers copy directly from a source text, `light` answers are based on the source text but include some light rephrasing, and `heavy` answers are based on the source text, but *heavily* rephrased (and will likely be the most challenging kind of plagiarism to detect).
**2. Non-plagiarized category: `non`.**
* `non` indicates that an answer is not plagiarized; the Wikipedia source text is not used to create this answer.
**3. Special, source text category: `orig`.**
* This is a specific category for the original, Wikipedia source text. We will use these files only for comparison purposes.
---
## Pre-Process the Data
In the next few cells, you'll be tasked with creating a new DataFrame of desired information about all of the files in the `data/` directory. This will prepare the data for feature extraction and for training a binary, plagiarism classifier.
### EXERCISE: Convert categorical to numerical data
You'll notice that the `Category` column in the data, contains string or categorical values, and to prepare these for feature extraction, we'll want to convert these into numerical values. Additionally, our goal is to create a binary classifier and so we'll need a binary class label that indicates whether an answer text is plagiarized (1) or not (0). Complete the below function `numerical_dataframe` that reads in a `file_information.csv` file by name, and returns a *new* DataFrame with a numerical `Category` column and a new `Class` column that labels each answer as plagiarized or not.
Your function should return a new DataFrame with the following properties:
* 4 columns: `File`, `Task`, `Category`, `Class`. The `File` and `Task` columns can remain unchanged from the original `.csv` file.
* Convert all `Category` labels to numerical labels according to the following rules (a higher value indicates a higher degree of plagiarism):
* 0 = `non`
* 1 = `heavy`
* 2 = `light`
* 3 = `cut`
* -1 = `orig`, this is a special value that indicates an original file.
* For the new `Class` column
* Any answer text that is not plagiarized (`non`) should have the class label `0`.
* Any plagiarized answer texts should have the class label `1`.
* And any `orig` texts will have a special label `-1`.
### Expected output
After running your function, you should get a DataFrame with rows that looks like the following:
```
File Task Category Class
0 g0pA_taska.txt a 0 0
1 g0pA_taskb.txt b 3 1
2 g0pA_taskc.txt c 2 1
3 g0pA_taskd.txt d 1 1
4 g0pA_taske.txt e 0 0
...
...
99 orig_taske.txt e -1 -1
```
```
# Read in a csv file and return a transformed dataframe
def numerical_dataframe(csv_file='data/file_information.csv'):
'''Reads in a csv file which is assumed to have `File`, `Category` and `Task` columns.
This function does two things:
1) converts `Category` column values to numerical values
2) Adds a new, numerical `Class` label column.
The `Class` column will label plagiarized answers as 1 and non-plagiarized as 0.
Source texts have a special label, -1.
:param csv_file: The directory for the file_information.csv file
:return: A dataframe with numerical categories and a new `Class` label column'''
df = pd.read_csv(csv_file)
# 1) converts `Category` column values to numerical values
df['Category'] = df['Category'].replace("non", 0)
df['Category'] = df['Category'].replace("heavy", 1)
df['Category'] = df['Category'].replace("light", 2)
df['Category'] = df['Category'].replace("cut", 3)
df['Category'] = df['Category'].replace("orig", -1)
# 2) Adds a new, numerical `Class` label column
df.loc[(df.Category == 0),'Class'] = 0
df.loc[(df.Category.isin([1, 2, 3])),'Class'] = 1
df.loc[(df.Category == -1),'Class'] = -1
df['Class'] = df['Class'].apply(np.int64) # convert float to int
return df
```
### Test cells
Below are a couple of test cells. The first is an informal test where you can check that your code is working as expected by calling your function and printing out the returned result.
The **second** cell below is a more rigorous test cell. The goal of a cell like this is to ensure that your code is working as expected, and to form any variables that might be used in _later_ tests/code, in this case, the data frame, `transformed_df`.
> The cells in this notebook should be run in chronological order (the order they appear in the notebook). This is especially important for test cells.
Often, later cells rely on the functions, imports, or variables defined in earlier cells. For example, some tests rely on previous tests to work.
These tests do not test all cases, but they are a great way to check that you are on the right track!
```
# informal testing, print out the results of a called function
# create new `transformed_df`
transformed_df = numerical_dataframe(csv_file ='data/file_information.csv')
# check work
# check that all categories of plagiarism have a class label = 1
transformed_df.head(10)
# test cell that creates `transformed_df`, if tests are passed
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
# importing tests
import problem_unittests as tests
# test numerical_dataframe function
tests.test_numerical_df(numerical_dataframe)
# if above test is passed, create NEW `transformed_df`
transformed_df = numerical_dataframe(csv_file ='data/file_information.csv')
# check work
print('\nExample data: ')
transformed_df.head()
```
## Text Processing & Splitting Data
Recall that the goal of this project is to build a plagiarism classifier. At it's heart, this task is a comparison text; one that looks at a given answer and a source text, compares them and predicts whether an answer has plagiarized from the source. To effectively do this comparison, and train a classifier we'll need to do a few more things: pre-process all of our text data and prepare the text files (in this case, the 95 answer files and 5 original source files) to be easily compared, and split our data into a `train` and `test` set that can be used to train a classifier and evaluate it, respectively.
To this end, you've been provided code that adds additional information to your `transformed_df` from above. The next two cells need not be changed; they add two additional columns to the `transformed_df`:
1. A `Text` column; this holds all the lowercase text for a `File`, with extraneous punctuation removed.
2. A `Datatype` column; this is a string value `train`, `test`, or `orig` that labels a data point as part of our train or test set
The details of how these additional columns are created can be found in the `helpers.py` file in the project directory. You're encouraged to read through that file to see exactly how text is processed and how data is split.
Run the cells below to get a `complete_df` that has all the information you need to proceed with plagiarism detection and feature engineering.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
import helpers
# create a text column
text_df = helpers.create_text_column(transformed_df)
text_df.head()
# after running the cell above
# check out the processed text for a single file, by row index
row_idx = 0 # feel free to change this index
sample_text = text_df.iloc[0]['Text']
print('Sample processed text:\n\n', sample_text)
```
## Split data into training and test sets
The next cell will add a `Datatype` column to a given DataFrame to indicate if the record is:
* `train` - Training data, for model training.
* `test` - Testing data, for model evaluation.
* `orig` - The task's original answer from wikipedia.
### Stratified sampling
The given code uses a helper function which you can view in the `helpers.py` file in the main project directory. This implements [stratified random sampling](https://en.wikipedia.org/wiki/Stratified_sampling) to randomly split data by task & plagiarism amount. Stratified sampling ensures that we get training and test data that is fairly evenly distributed across task & plagiarism combinations. Approximately 26% of the data is held out for testing and 74% of the data is used for training.
The function **train_test_dataframe** takes in a DataFrame that it assumes has `Task` and `Category` columns, and, returns a modified frame that indicates which `Datatype` (train, test, or orig) a file falls into. This sampling will change slightly based on a passed in *random_seed*. Due to a small sample size, this stratified random sampling will provide more stable results for a binary plagiarism classifier. Stability here is smaller *variance* in the accuracy of classifier, given a random seed.
```
random_seed = 1 # can change; set for reproducibility
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
import helpers
# create new df with Datatype (train, test, orig) column
# pass in `text_df` from above to create a complete dataframe, with all the information you need
complete_df = helpers.train_test_dataframe(text_df, random_seed=random_seed)
# check results
complete_df.head(10)
```
# Determining Plagiarism
Now that you've prepared this data and created a `complete_df` of information, including the text and class associated with each file, you can move on to the task of extracting similarity features that will be useful for plagiarism classification.
> Note: The following code exercises, assume that the `complete_df` as it exists now, will **not** have its existing columns modified.
The `complete_df` should always include the columns: `['File', 'Task', 'Category', 'Class', 'Text', 'Datatype']`. You can add additional columns, and you can create any new DataFrames you need by copying the parts of the `complete_df` as long as you do not modify the existing values, directly.
---
# Similarity Features
One of the ways we might go about detecting plagiarism, is by computing **similarity features** that measure how similar a given answer text is as compared to the original wikipedia source text (for a specific task, a-e). The similarity features you will use are informed by [this paper on plagiarism detection](https://s3.amazonaws.com/video.udacity-data.com/topher/2019/January/5c412841_developing-a-corpus-of-plagiarised-short-answers/developing-a-corpus-of-plagiarised-short-answers.pdf).
> In this paper, researchers created features called **containment** and **longest common subsequence**.
Using these features as input, you will train a model to distinguish between plagiarized and not-plagiarized text files.
## Feature Engineering
Let's talk a bit more about the features we want to include in a plagiarism detection model and how to calculate such features. In the following explanations, I'll refer to a submitted text file as a **Student Answer Text (A)** and the original, wikipedia source file (that we want to compare that answer to) as the **Wikipedia Source Text (S)**.
### Containment
Your first task will be to create **containment features**. To understand containment, let's first revisit a definition of [n-grams](https://en.wikipedia.org/wiki/N-gram). An *n-gram* is a sequential word grouping. For example, in a line like "bayes rule gives us a way to combine prior knowledge with new information," a 1-gram is just one word, like "bayes." A 2-gram might be "bayes rule" and a 3-gram might be "combine prior knowledge."
> Containment is defined as the **intersection** of the n-gram word count of the Wikipedia Source Text (S) with the n-gram word count of the Student Answer Text (S) *divided* by the n-gram word count of the Student Answer Text.
$$ \frac{\sum{count(\text{ngram}_{A}) \cap count(\text{ngram}_{S})}}{\sum{count(\text{ngram}_{A})}} $$
If the two texts have no n-grams in common, the containment will be 0, but if _all_ their n-grams intersect then the containment will be 1. Intuitively, you can see how having longer n-gram's in common, might be an indication of cut-and-paste plagiarism. In this project, it will be up to you to decide on the appropriate `n` or several `n`'s to use in your final model.
### EXERCISE: Create containment features
Given the `complete_df` that you've created, you should have all the information you need to compare any Student Answer Text (A) with its appropriate Wikipedia Source Text (S). An answer for task A should be compared to the source text for task A, just as answers to tasks B, C, D, and E should be compared to the corresponding original source text.
In this exercise, you'll complete the function, `calculate_containment` which calculates containment based upon the following parameters:
* A given DataFrame, `df` (which is assumed to be the `complete_df` from above)
* An `answer_filename`, such as 'g0pB_taskd.txt'
* An n-gram length, `n`
### Containment calculation
The general steps to complete this function are as follows:
1. From *all* of the text files in a given `df`, create an array of n-gram counts; it is suggested that you use a [CountVectorizer](https://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.CountVectorizer.html) for this purpose.
2. Get the processed answer and source texts for the given `answer_filename`.
3. Calculate the containment between an answer and source text according to the following equation.
>$$ \frac{\sum{count(\text{ngram}_{A}) \cap count(\text{ngram}_{S})}}{\sum{count(\text{ngram}_{A})}} $$
4. Return that containment value.
You are encouraged to write any helper functions that you need to complete the function below.
```
from sklearn.feature_extraction.text import CountVectorizer
# Calculate the ngram containment for one answer file/source file pair in a df
def calculate_containment(df, n, answer_filename):
'''Calculates the containment between a given answer text and its associated source text.
This function creates a count of ngrams (of a size, n) for each text file in our data.
Then calculates the containment by finding the ngram count for a given answer text,
and its associated source text, and calculating the normalized intersection of those counts.
:param df: A dataframe with columns,
'File', 'Task', 'Category', 'Class', 'Text', and 'Datatype'
:param n: An integer that defines the ngram size
:param answer_filename: A filename for an answer text in the df, ex. 'g0pB_taskd.txt'
:return: A single containment value that represents the similarity
between an answer text and its source text.
'''
counts = CountVectorizer(analyzer='word', ngram_range=(n,n))
# Get the processed answer and source texts for the given answer_filename.
task = df.loc[df['File'] == answer_filename, 'Task'].iloc[0] # e.g. "a"
source = df.loc[(df['Class'] == -1) & (df['Task'] == task), 'Text'].iloc[0] # original text
answer = df.loc[df['File'] == answer_filename, 'Text'].iloc[0] # answer text
# Create array of n-gram counts for the answer and source text
ngrams = counts.fit_transform([answer, source])
ngram_array = ngrams.toarray()
# Calculate the containment between an answer and source text according to the equation.
intersection_list = np.amin(ngram_array, axis=0)
intersection = np.sum(intersection_list)
answer_cnt = np.sum(ngram_array[0]) # count up the number of n-grams in the answer text
containment_val = intersection / answer_cnt # normalize and get final containment value
# 4. Return that containment value.
return containment_val
```
### Test cells
After you've implemented the containment function, you can test out its behavior.
The cell below iterates through the first few files, and calculates the original category _and_ containment values for a specified n and file.
>If you've implemented this correctly, you should see that the non-plagiarized have low or close to 0 containment values and that plagiarized examples have higher containment values, closer to 1.
Note what happens when you change the value of n. I recommend applying your code to multiple files and comparing the resultant containment values. You should see that the highest containment values correspond to files with the highest category (`cut`) of plagiarism level.
```
# select a value for n
n = 3
# indices for first few files
test_indices = range(5)
# iterate through files and calculate containment
category_vals = []
containment_vals = []
for i in test_indices:
# get level of plagiarism for a given file index
category_vals.append(complete_df.loc[i, 'Category'])
# calculate containment for given file and n
filename = complete_df.loc[i, 'File']
c = calculate_containment(complete_df, n, filename)
containment_vals.append(c)
# print out result, does it make sense?
print('Original category values: \n', category_vals)
print()
print(str(n)+'-gram containment values: \n', containment_vals)
# run this test cell
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
# test containment calculation
# params: complete_df from before, and containment function
tests.test_containment(complete_df, calculate_containment)
```
### QUESTION 1: Why can we calculate containment features across *all* data (training & test), prior to splitting the DataFrame for modeling? That is, what about the containment calculation means that the test and training data do not influence each other?
**Answer:**
It would be a bad practice to train a model on a test data. Containment is calculated between a given answer text and its associated source text, so test and training data do not influence each other here.
---
## Longest Common Subsequence
Containment a good way to find overlap in word usage between two documents; it may help identify cases of cut-and-paste as well as paraphrased levels of plagiarism. Since plagiarism is a fairly complex task with varying levels, it's often useful to include other measures of similarity. The paper also discusses a feature called **longest common subsequence**.
> The longest common subsequence is the longest string of words (or letters) that are *the same* between the Wikipedia Source Text (S) and the Student Answer Text (A). This value is also normalized by dividing by the total number of words (or letters) in the Student Answer Text.
In this exercise, we'll ask you to calculate the longest common subsequence of words between two texts.
### EXERCISE: Calculate the longest common subsequence
Complete the function `lcs_norm_word`; this should calculate the *longest common subsequence* of words between a Student Answer Text and corresponding Wikipedia Source Text.
It may be helpful to think of this in a concrete example. A Longest Common Subsequence (LCS) problem may look as follows:
* Given two texts: text A (answer text) of length n, and string S (original source text) of length m. Our goal is to produce their longest common subsequence of words: the longest sequence of words that appear left-to-right in both texts (though the words don't have to be in continuous order).
* Consider:
* A = "i think pagerank is a link analysis algorithm used by google that uses a system of weights attached to each element of a hyperlinked set of documents"
* S = "pagerank is a link analysis algorithm used by the google internet search engine that assigns a numerical weighting to each element of a hyperlinked set of documents"
* In this case, we can see that the start of each sentence of fairly similar, having overlap in the sequence of words, "pagerank is a link analysis algorithm used by" before diverging slightly. Then we **continue moving left -to-right along both texts** until we see the next common sequence; in this case it is only one word, "google". Next we find "that" and "a" and finally the same ending "to each element of a hyperlinked set of documents".
* Below, is a clear visual of how these sequences were found, sequentially, in each text.
<img src='notebook_ims/common_subseq_words.png' width=40% />
* Now, those words appear in left-to-right order in each document, sequentially, and even though there are some words in between, we count this as the longest common subsequence between the two texts.
* If I count up each word that I found in common I get the value 20. **So, LCS has length 20**.
* Next, to normalize this value, divide by the total length of the student answer; in this example that length is only 27. **So, the function `lcs_norm_word` should return the value `20/27` or about `0.7408`.**
In this way, LCS is a great indicator of cut-and-paste plagiarism or if someone has referenced the same source text multiple times in an answer.
### LCS, dynamic programming
If you read through the scenario above, you can see that this algorithm depends on looking at two texts and comparing them word by word. You can solve this problem in multiple ways. First, it may be useful to `.split()` each text into lists of comma separated words to compare. Then, you can iterate through each word in the texts and compare them, adding to your value for LCS as you go.
The method I recommend for implementing an efficient LCS algorithm is: using a matrix and dynamic programming. **Dynamic programming** is all about breaking a larger problem into a smaller set of subproblems, and building up a complete result without having to repeat any subproblems.
This approach assumes that you can split up a large LCS task into a combination of smaller LCS tasks. Let's look at a simple example that compares letters:
* A = "ABCD"
* S = "BD"
We can see right away that the longest subsequence of _letters_ here is 2 (B and D are in sequence in both strings). And we can calculate this by looking at relationships between each letter in the two strings, A and S.
Here, I have a matrix with the letters of A on top and the letters of S on the left side:
<img src='notebook_ims/matrix_1.png' width=40% />
This starts out as a matrix that has as many columns and rows as letters in the strings S and O **+1** additional row and column, filled with zeros on the top and left sides. So, in this case, instead of a 2x4 matrix it is a 3x5.
Now, we can fill this matrix up by breaking it into smaller LCS problems. For example, let's first look at the shortest substrings: the starting letter of A and S. We'll first ask, what is the Longest Common Subsequence between these two letters "A" and "B"?
**Here, the answer is zero and we fill in the corresponding grid cell with that value.**
<img src='notebook_ims/matrix_2.png' width=30% />
Then, we ask the next question, what is the LCS between "AB" and "B"?
**Here, we have a match, and can fill in the appropriate value 1**.
<img src='notebook_ims/matrix_3_match.png' width=25% />
If we continue, we get to a final matrix that looks as follows, with a **2** in the bottom right corner.
<img src='notebook_ims/matrix_6_complete.png' width=25% />
The final LCS will be that value **2** *normalized* by the number of n-grams in A. So, our normalized value is 2/4 = **0.5**.
### The matrix rules
One thing to notice here is that, you can efficiently fill up this matrix one cell at a time. Each grid cell only depends on the values in the grid cells that are directly on top and to the left of it, or on the diagonal/top-left. The rules are as follows:
* Start with a matrix that has one extra row and column of zeros.
* As you traverse your string:
* If there is a match, fill that grid cell with the value to the top-left of that cell *plus* one. So, in our case, when we found a matching B-B, we added +1 to the value in the top-left of the matching cell, 0.
* If there is not a match, take the *maximum* value from either directly to the left or the top cell, and carry that value over to the non-match cell.
<img src='notebook_ims/matrix_rules.png' width=50% />
After completely filling the matrix, **the bottom-right cell will hold the non-normalized LCS value**.
This matrix treatment can be applied to a set of words instead of letters. Your function should apply this to the words in two texts and return the normalized LCS value.
```
# Compute the normalized LCS given an answer text and a source text
def lcs_norm_word(answer_text, source_text):
'''Computes the longest common subsequence of words in two texts; returns a normalized value.
:param answer_text: The pre-processed text for an answer text
:param source_text: The pre-processed text for an answer's associated source text
:return: A normalized LCS value'''
answer_words = answer_text.split()
source_words = source_text.split()
rows = len(answer_words)
cols = len(source_words)
matrix = np.zeros((rows+1, cols+1))
row = col = 1
for answer_word in answer_words:
col = 1
for source_word in source_words:
if answer_word == source_word:
matrix[row][col] = matrix[row-1][col-1] + 1
else:
matrix[row][col] = max(matrix[row][col-1], matrix[row-1][col])
col += 1
row += 1
LCS = matrix[rows][cols]
return LCS/rows
```
### Test cells
Let's start by testing out your code on the example given in the initial description.
In the below cell, we have specified strings A (answer text) and S (original source text). We know that these texts have 20 words in common and the submitted answer is 27 words long, so the normalized, longest common subsequence should be 20/27.
```
# Run the test scenario from above
# does your function return the expected value?
A = "i think pagerank is a link analysis algorithm used by google that uses a system of weights attached to each element of a hyperlinked set of documents"
S = "pagerank is a link analysis algorithm used by the google internet search engine that assigns a numerical weighting to each element of a hyperlinked set of documents"
# calculate LCS
lcs = lcs_norm_word(A, S)
print('LCS = ', lcs)
# expected value test
assert lcs==20/27., "Incorrect LCS value, expected about 0.7408, got "+str(lcs)
print('Test passed!')
```
This next cell runs a more rigorous test.
```
# run test cell
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
# test lcs implementation
# params: complete_df from before, and lcs_norm_word function
tests.test_lcs(complete_df, lcs_norm_word)
```
Finally, take a look at a few resultant values for `lcs_norm_word`. Just like before, you should see that higher values correspond to higher levels of plagiarism.
```
# test on your own
test_indices = range(5) # look at first few files
category_vals = []
lcs_norm_vals = []
# iterate through first few docs and calculate LCS
for i in test_indices:
category_vals.append(complete_df.loc[i, 'Category'])
# get texts to compare
answer_text = complete_df.loc[i, 'Text']
task = complete_df.loc[i, 'Task']
# we know that source texts have Class = -1
orig_rows = complete_df[(complete_df['Class'] == -1)]
orig_row = orig_rows[(orig_rows['Task'] == task)]
source_text = orig_row['Text'].values[0]
# calculate lcs
lcs_val = lcs_norm_word(answer_text, source_text)
lcs_norm_vals.append(lcs_val)
# print out result, does it make sense?
print('Original category values: \n', category_vals)
print()
print('Normalized LCS values: \n', lcs_norm_vals)
```
---
# Create All Features
Now that you've completed the feature calculation functions, it's time to actually create multiple features and decide on which ones to use in your final model! In the below cells, you're provided two helper functions to help you create multiple features and store those in a DataFrame, `features_df`.
### Creating multiple containment features
Your completed `calculate_containment` function will be called in the next cell, which defines the helper function `create_containment_features`.
> This function returns a list of containment features, calculated for a given `n` and for *all* files in a df (assumed to the the `complete_df`).
For our original files, the containment value is set to a special value, -1.
This function gives you the ability to easily create several containment features, of different n-gram lengths, for each of our text files.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
# Function returns a list of containment features, calculated for a given n
# Should return a list of length 100 for all files in a complete_df
def create_containment_features(df, n, column_name=None):
containment_values = []
if(column_name==None):
column_name = 'c_'+str(n) # c_1, c_2, .. c_n
# iterates through dataframe rows
for i in df.index:
file = df.loc[i, 'File']
# Computes features using calculate_containment function
if df.loc[i,'Category'] > -1:
c = calculate_containment(df, n, file)
containment_values.append(c)
# Sets value to -1 for original tasks
else:
containment_values.append(-1)
print(str(n)+'-gram containment features created!')
return containment_values
```
### Creating LCS features
Below, your complete `lcs_norm_word` function is used to create a list of LCS features for all the answer files in a given DataFrame (again, this assumes you are passing in the `complete_df`. It assigns a special value for our original, source files, -1.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
# Function creates lcs feature and add it to the dataframe
def create_lcs_features(df, column_name='lcs_word'):
lcs_values = []
# iterate through files in dataframe
for i in df.index:
# Computes LCS_norm words feature using function above for answer tasks
if df.loc[i,'Category'] > -1:
# get texts to compare
answer_text = df.loc[i, 'Text']
task = df.loc[i, 'Task']
# we know that source texts have Class = -1
orig_rows = df[(df['Class'] == -1)]
orig_row = orig_rows[(orig_rows['Task'] == task)]
source_text = orig_row['Text'].values[0]
# calculate lcs
lcs = lcs_norm_word(answer_text, source_text)
lcs_values.append(lcs)
# Sets to -1 for original tasks
else:
lcs_values.append(-1)
print('LCS features created!')
return lcs_values
```
## EXERCISE: Create a features DataFrame by selecting an `ngram_range`
The paper suggests calculating the following features: containment *1-gram to 5-gram* and *longest common subsequence*.
> In this exercise, you can choose to create even more features, for example from *1-gram to 7-gram* containment features and *longest common subsequence*.
You'll want to create at least 6 features to choose from as you think about which to give to your final, classification model. Defining and comparing at least 6 different features allows you to discard any features that seem redundant, and choose to use the best features for your final model!
In the below cell **define an n-gram range**; these will be the n's you use to create n-gram containment features. The rest of the feature creation code is provided.
```
# Define an ngram range
ngram_range = range(1,7)
# The following code may take a minute to run, depending on your ngram_range
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
features_list = []
# Create features in a features_df
all_features = np.zeros((len(ngram_range)+1, len(complete_df)))
# Calculate features for containment for ngrams in range
i=0
for n in ngram_range:
column_name = 'c_'+str(n)
features_list.append(column_name)
# create containment features
all_features[i]=np.squeeze(create_containment_features(complete_df, n))
i+=1
# Calculate features for LCS_Norm Words
features_list.append('lcs_word')
all_features[i]= np.squeeze(create_lcs_features(complete_df))
# create a features dataframe
features_df = pd.DataFrame(np.transpose(all_features), columns=features_list)
# Print all features/columns
print()
print('Features: ', features_list)
print()
# print some results
features_df.head(10)
```
## Correlated Features
You should use feature correlation across the *entire* dataset to determine which features are ***too*** **highly-correlated** with each other to include both features in a single model. For this analysis, you can use the *entire* dataset due to the small sample size we have.
All of our features try to measure the similarity between two texts. Since our features are designed to measure similarity, it is expected that these features will be highly-correlated. Many classification models, for example a Naive Bayes classifier, rely on the assumption that features are *not* highly correlated; highly-correlated features may over-inflate the importance of a single feature.
So, you'll want to choose your features based on which pairings have the lowest correlation. These correlation values range between 0 and 1; from low to high correlation, and are displayed in a [correlation matrix](https://www.displayr.com/what-is-a-correlation-matrix/), below.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
# Create correlation matrix for just Features to determine different models to test
corr_matrix = features_df.corr().abs().round(2)
# display shows all of a dataframe
display(corr_matrix)
```
## EXERCISE: Create selected train/test data
Complete the `train_test_data` function below. This function should take in the following parameters:
* `complete_df`: A DataFrame that contains all of our processed text data, file info, datatypes, and class labels
* `features_df`: A DataFrame of all calculated features, such as containment for ngrams, n= 1-5, and lcs values for each text file listed in the `complete_df` (this was created in the above cells)
* `selected_features`: A list of feature column names, ex. `['c_1', 'lcs_word']`, which will be used to select the final features in creating train/test sets of data.
It should return two tuples:
* `(train_x, train_y)`, selected training features and their corresponding class labels (0/1)
* `(test_x, test_y)`, selected training features and their corresponding class labels (0/1)
** Note: x and y should be arrays of feature values and numerical class labels, respectively; not DataFrames.**
Looking at the above correlation matrix, you should decide on a **cutoff** correlation value, less than 1.0, to determine which sets of features are *too* highly-correlated to be included in the final training and test data. If you cannot find features that are less correlated than some cutoff value, it is suggested that you increase the number of features (longer n-grams) to choose from or use *only one or two* features in your final model to avoid introducing highly-correlated features.
Recall that the `complete_df` has a `Datatype` column that indicates whether data should be `train` or `test` data; this should help you split the data appropriately.
```
# Takes in dataframes and a list of selected features (column names)
# and returns (train_x, train_y), (test_x, test_y)
def train_test_data(complete_df, features_df, selected_features):
'''Gets selected training and test features from given dataframes, and
returns tuples for training and test features and their corresponding class labels.
:param complete_df: A dataframe with all of our processed text data, datatypes, and labels
:param features_df: A dataframe of all computed, similarity features
:param selected_features: An array of selected features that correspond to certain columns in `features_df`
:return: training and test features and labels: (train_x, train_y), (test_x, test_y)'''
selected_features_df = features_df.loc[: , selected_features]
# get the training features
train_x = np.array(selected_features_df[complete_df['Datatype'] == 'train'])
# And training class labels (0 or 1)
train_y = np.array(complete_df.loc[: , ['Class']][complete_df['Datatype'] == 'train']['Class']) # should be 1d array
# get the test features and labels
test_x = np.array(selected_features_df[complete_df['Datatype'] == 'test'])
test_y = np.array(complete_df.loc[: , ['Class']][complete_df['Datatype'] == 'test']['Class']) # should be 1d array
return (train_x, train_y), (test_x, test_y)
```
### Test cells
Below, test out your implementation and create the final train/test data.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
test_selection = list(features_df)[:2] # first couple columns as a test
# test that the correct train/test data is created
(train_x, train_y), (test_x, test_y) = train_test_data(complete_df, features_df, test_selection)
# params: generated train/test data
tests.test_data_split(train_x, train_y, test_x, test_y)
```
## EXERCISE: Select "good" features
If you passed the test above, you can create your own train/test data, below.
Define a list of features you'd like to include in your final mode, `selected_features`; this is a list of the features names you want to include.
```
# Select your list of features, this should be column names from features_df
# ex. ['c_1', 'lcs_word']
selected_features = ['c_1', 'c_5', 'lcs_word']
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
(train_x, train_y), (test_x, test_y) = train_test_data(complete_df, features_df, selected_features)
# check that division of samples seems correct
# these should add up to 95 (100 - 5 original files)
print('Training size: ', len(train_x))
print('Test size: ', len(test_x))
print()
print('Training df sample: \n', train_x[:10])
```
### Question 2: How did you decide on which features to include in your final model?
**Answer:**
Features are chosen based on pairings to have the lowest correlation. From the correlation matrix above, these are c1 vs. c6. We could also include c5 based on c1 vs. c5 correlation coefficient of 0.88, but c5 is strongly correlated (corr 1.00) with c6 that we already chose. We will also include lcs_word, which has a correlation coefficient of 0.97 with c1 and 0.94 with c6.
---
## Creating Final Data Files
Now, you are almost ready to move on to training a model in SageMaker!
You'll want to access your train and test data in SageMaker and upload it to S3. In this project, SageMaker will expect the following format for your train/test data:
* Training and test data should be saved in one `.csv` file each, ex `train.csv` and `test.csv`
* These files should have class labels in the first column and features in the rest of the columns
This format follows the practice, outlined in the [SageMaker documentation](https://docs.aws.amazon.com/sagemaker/latest/dg/cdf-training.html), which reads: "Amazon SageMaker requires that a CSV file doesn't have a header record and that the target variable [class label] is in the first column."
## EXERCISE: Create csv files
Define a function that takes in x (features) and y (labels) and saves them to one `.csv` file at the path `data_dir/filename`.
It may be useful to use pandas to merge your features and labels into one DataFrame and then convert that into a csv file. You can make sure to get rid of any incomplete rows, in a DataFrame, by using `dropna`.
```
def make_csv(x, y, filename, data_dir):
'''Merges features and labels and converts them into one csv file with labels in the first column.
:param x: Data features
:param y: Data labels
:param file_name: Name of csv file, ex. 'train.csv'
:param data_dir: The directory where files will be saved
'''
# make data dir, if it does not exist
if not os.path.exists(data_dir):
os.makedirs(data_dir)
# first column is the labels and rest is features
pd.concat([pd.DataFrame(y), pd.DataFrame(x)], axis=1).dropna().to_csv(os.path.join(data_dir, filename), header=False, index=False)
# nothing is returned, but a print statement indicates that the function has run
print('Path created: '+str(data_dir)+'/'+str(filename))
```
### Test cells
Test that your code produces the correct format for a `.csv` file, given some text features and labels.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
fake_x = [ [0.39814815, 0.0001, 0.19178082],
[0.86936937, 0.44954128, 0.84649123],
[0.44086022, 0., 0.22395833] ]
fake_y = [0, 1, 1]
make_csv(fake_x, fake_y, filename='to_delete.csv', data_dir='test_csv')
# read in and test dimensions
fake_df = pd.read_csv('test_csv/to_delete.csv', header=None)
# check shape
assert fake_df.shape==(3, 4), \
'The file should have as many rows as data_points and as many columns as features+1 (for indices).'
# check that first column = labels
assert np.all(fake_df.iloc[:,0].values==fake_y), 'First column is not equal to the labels, fake_y.'
print('Tests passed!')
# delete the test csv file, generated above
! rm -rf test_csv
```
If you've passed the tests above, run the following cell to create `train.csv` and `test.csv` files in a directory that you specify! This will save the data in a local directory. Remember the name of this directory because you will reference it again when uploading this data to S3.
```
# can change directory, if you want
data_dir = 'plagiarism_data'
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
make_csv(train_x, train_y, filename='train.csv', data_dir=data_dir)
make_csv(test_x, test_y, filename='test.csv', data_dir=data_dir)
```
## Up Next
Now that you've done some feature engineering and created some training and test data, you are ready to train and deploy a plagiarism classification model. The next notebook will utilize SageMaker resources to train and test a model that you design.
| github_jupyter |
```
import numpy as np
import scipy.optimize
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
```
## Read in data and plot
```
# load in the data, you will need to change the path to the file
data = pd.read_csv("../../data/competition_experiments/plate_colony_data/colony_counting_20200225.csv")
data["ratio"] = data["wildtype / mL"] / data["delta / mL"]
data["time since start (hr)"] = data["time since start (min)"] / 60
data.head()
# plot
times = data["time since start (hr)"]
N_wildtype = data["wildtype / mL"]
N_delta = data["delta / mL"]
plt.plot(times, N_wildtype, 'r.')
plt.plot(times, N_delta, 'b.')
plt.xlabel("time (hr)")
plt.ylabel("cells / mL")
plt.legend(["wildtype", "∆lac"]);
plt.savefig('plot.eps')
```
Great! This looks like what we expected! wildtype is definitely taking over. Now let's consider the ratio of wildtype over the delta strain over time. If these two strains are both growing exponentially, then we expect their ratio to be exponential as well.
```
plt.plot(times, data["ratio"], 'r.')
plt.xlabel("time (min)")
plt.ylabel("wt / delta");
```
## Computing a selection coefficient
As we saw by eye, the wildtype strain takes over. Fitting an expoential to the ratio will elucidate the selection coefficient. The ratio of two exponentially growing strains will look like this over time, where $r_1$ and $r_2$ are the corresponding growth rate:
$$ \frac{N_1(t)}{N_2(t)} =
\frac{N_1(0) e^{r_1t}}{N_2(0) e^{r_2t}} =
\frac{N_1(0)}{N_2(0)} e^{(r_1-r_2)t} $$
Let's determine the best fit for data above to get at this difference in growth date, or the selection coefficient. Let's first define an exponetial fucnction, the function we wish to fit.
```
def exp_curve(times, A, b):
return A * np.exp(b*times)
```
Now, we will use scipy's optimize function to get the best fit. The inputs to the function are:
- The function we are fitting, `exp_curve()` in this case
- The x-values of our data
- The y-values of our data
- The lower bounds and upper bounds on our parameters (A and b in order)
```
popt_ratio, _ = scipy.optimize.curve_fit(exp_curve,
data["time since start (hr)"],
data["ratio"])
```
Let's see what parameters we got!
```
s = np.round(popt_ratio[1],5)
print("Fit for A:", np.round(popt_ratio[0],5))
print("Fit for b:", s, "per hour")
```
And let's plot the fit.
```
# plot data
plt.plot(data["time since start (hr)"], data["ratio"], 'r.')
# time values for making a smooth curve
time_range = np.linspace(0, 12, 200)
# plot fits
plt.plot(time_range, exp_curve(time_range, popt_ratio[0], popt_ratio[1]), color="red")
plt.xlabel("time (hr)")
plt.ylabel("wt / detla");
plt.savefig('plot.eps')
```
## Considering the growth rates independently
### wildtype
```
popt, _ = scipy.optimize.curve_fit(exp_curve,
data["time since start (hr)"],
data["wildtype / mL"])
r_wt = np.round(popt[1],5)
print("Fit for A:", np.round(popt_ratio[0],5))
print("Fit for b:", r_wt, "per hour")
# plot data
plt.plot(data["time since start (hr)"], data["wildtype / mL"], 'r.')
# time values for making a smooth curve
time_range = np.linspace(0, 12, 200)
# plot fits
plt.plot(time_range, exp_curve(time_range, popt[0], popt[1]), color="red")
plt.xlabel("time (hr)")
plt.ylabel("wt count");
```
### delta
```
popt, _ = scipy.optimize.curve_fit(exp_curve,
data["time since start (hr)"],
data["delta / mL"])
r_delta = np.round(popt[1],5)
print("Fit for A:", np.round(popt_ratio[0],5))
print("Fit for b:", r_delta, "per hour")
# plot data
plt.plot(data["time since start (hr)"], data["delta / mL"], 'r.')
# time values for making a smooth curve
time_range = np.linspace(0, 12, 200)
# plot fits
plt.plot(time_range, exp_curve(time_range, popt[0], popt[1]), color="red")
plt.xlabel("time (hr)")
plt.ylabel("detla count");
```
### Comparing the two approaches
The selection coefficient should be the difference in the growth rates. Let's check this out:
```
s
r_wt - r_delta
```
Hmmm . . .
| github_jupyter |
# Real Estate Price Predictor
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
housing = pd.read_csv("data.csv")
```
## Train - Test Splitting
```
train_set, test_set = split_train_test(housing, 0.2)
print(f" Rows in train set : {len(train_set)} \n Rows in test set: {len(test_set)} \n")
```
# this whole functon is in sklearn
```
#from sklearn.model_selection import train_test_split, StratifiedShuffleSplit
#train_set, test_set = train_test_split(housing, test_size=0.2, random_state=42)
#print(f" Rows in train set : {len(train_set)} \n Rows in test set: {len(test_set)} \n")
```
##We are using stratified learning because to seprate CHAS values in both test and train data sets
```
from sklearn.model_selection import StratifiedShuffleSplit
split = StratifiedShuffleSplit(n_splits=1, test_size=0.2, random_state=42)
for train_index, test_index in split.split(housing, housing['CHAS']):
strat_train_set = housing.loc[train_index]
strat_test_set =
strat_test_set['CHAS'].value_counts()
housing["TAXRM"] = housing['TAX']/housing['RM']
strat_train_set['CHAS'].value_counts()
```
## Looking for co relations
```
corr = housing.corr()
```
## correlation basically tells how much the value MEDV changes wrt to other features
```
corr["MEDV"].sort_values(ascending=False)
from pandas.plotting import scatter_matrix
attributes = ['RM', 'ZN', 'MEDV']
scatter_matrix(housing[attributes], figsize=(12, 8))
housing.plot(kind="scatter", x = "RM", y="MEDV", alpha=0.8)
housing.head()
```
## Creating Pipleline
```
housing = strat_train_set.drop("MEDV", axis=1)
housing_labels = strat_train_set["MEDV"].copy()
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.impute import SimpleImputer
my_pipeline = Pipeline([
('imputer', SimpleImputer(strategy = "median")),
('std_scaler', StandardScaler()),
])
housing_new = my_pipeline.fit_transform(housing)
housing_new.shape
from sklearn.linear_model import LinearRegression
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import RandomForestRegressor
#model = LinearRegression()
#model = DecisionTreeRegressor()
model = RandomForestRegressor()
model.fit(housing_new, housing_labels)
some_data = housing.iloc[:5]
some_labels = housing_labels.iloc[:5]
prepared_data = my_pipeline.transform(some_data)
model.predict(prepared_data)
list(some_labels)
```
EVALUATING THE MODEL
```
from sklearn.metrics import mean_squared_error
housing_predictions = model.predict(housing_new)
mse = mean_squared_error(housing_labels, housing_predictions)
rmse = np.sqrt(mse)
rmse ##not good enough
from sklearn.model_selection import cross_val_score
score = cross_val_score(model, housing_new, housing_labels, scoring = "neg_mean_squared_error", cv = 10)
rmse_scores = np.sqrt(-score)
rmse_scores
print("array([4.21554054, 4.47792731, 4.98595284, 6.34383266, 5.28161956]) LR \narray([4.95871847, 4.49407017, 5.45017839, 5.17634696, 3.23797776]) DTR")
```
DecisionTreeRegressor is better here
```
def print_scores(rmse_scores):
print("Scores", score)
print("mean", score.mean())
print("Standard Deviation", score.std())
print_scores(rmse_scores)
```
| github_jupyter |
Na dnešní lekci si do virtuálního prostředí nainstalujte následující balíčky.
Můžete použít prostředí z lekce o NumPy.
```console
$ python -m pip install --upgrade pip
$ python -m pip install notebook pandas matplotlib
```
Pro případ, že by vaše verze `pip`-u neuměla *wheels* nebo na PyPI nebyly příslušné *wheel* balíčky, je dobré mít na systému nainstalovaný překladač C a Fortranu (např. `gcc`, `gcc-gfortran`) a hlavičkové soubory Pythonu (např. `python3-devel`). Jestli je ale nemáte, zkuste instalaci přímo – *wheels* pro většinu operačních systémů existují – a až kdyby to nefungovalo, instalujte překladače a hlavičky.
Mezitím co se instaluje, stáhněte si do adresáře `static` potřebné soubory:
[actors.csv](static/actors.csv) a
[spouses.csv](static/spouses.csv).
A až bude nainstalováno, spusťte si nový Notebook. (Viz [lekce o Notebooku](../notebook/).)
---
# Analýza dat v Pythonu
Jedna z oblastí, kde popularita Pythonu neustále roste, je analýza dat. Co tenhle termín znamená?
Máme nějaká data; je jich moc a jsou nepřehledná. Datový analytik je zpracuje, přeskládá, najde v nich smysl, vytvoří shrnutí toho nejdůležitějšího nebo barevnou infografiku.
Ze statistických údajů o obyvatelstvu zjistíme, jak souvisí příjmy s dostupností škol. Zpracováním měření z fyzikálního experimentu ověříme, jestli platí hypotéza. Z log přístupů na webovou službu určíme, co uživatelé čtou a kde stránky opouštějí.
Na podobné úkoly je možné použít jazyky vyvinuté přímo pro analýzu dat, jako R, které takovým úkolům svojí syntaxí a filozofií odpovídají víc. Python jako obecný programovací jazyk sice místy vyžaduje krkolomnější zápis, ale zato nabízí možnost data spojit s jinými oblastmi – od získávání informací z webových stránek po tvoření webových či desktopových rozhraní.
## Proces analýzy dat
Práce datového analytika se většinou drží následujícího postupu:
* Formulace otázky, kterou chceme zodpovědět
* Identifikace dat, která můžeme použít
* Získání dat (stažení, převod do použitelného formátu)
* Uložení dat
* Zkoumání dat
* Publikace výsledků
<small>*(založeno na diagramu z knihy *Data Wrangling in Python* od Jacqueline Kazil & Katharine Jarmul, str. 3)*</small>
S prvními dvěma kroky Python příliš nepomůže; k těm jen poznamenám, že „Co zajímavého se z těch dat dá vyčíst?” je validní otázka. Na druhé dva kroky se dá s úspěchem použít pythonní standardní knihovna: `json`, `csv`, případně doinstalovat `requests`, `lxml` pro XML či `xlwt`/`openpyxl` na excelové soubory.
Na zkoumání dat a přípravu výsledků pak použijeme specializovanou „datovou” knihovnu – Pandas.
# Pandas
Pandas slouží pro analýzu dat, které lze reprezentovat 2D tabulkou. Tento „tvar” dat najdeme v SQL databázích, souborech CSV nebo tabulkových procesorech. Stručně řečeno, co jde dělat v Excelu, jde dělat i v Pandas. (Pandas má samozřejmě funkce navíc, a hlavně umožňuje analýzu automatizovat.)
Jak bylo řečeno u [NumPy](../numpy/), analytici – cílová skupina této knihovny – mají rádi zkratky. Ve spoustě materiálů na Webu proto najdete `import pandas as pd`, případně rovnou (a bez vysvětlení) použité `pd` jako zkratku pro `pandas`. Tento návod ale používá plné jméno.
```
import pandas
```
## Tabulky
Základní datový typ, který Pandas nabízí, je `DataFrame`, neboli lidově „tabulka”. Jednotlivé záznamy jsou v ní uvedeny jako řádky a části těchto záznamů jsou úhledně srovnány ve sloupcích.
Nejpoužívanější způsob, jak naplnit první DataFrame, je načtení ze souboru. Na to má Pandas sadu funkcí začínající `read_`. (Některé z nich potřebují další knihovny, viz dokumentace.)
Jeden z nejpříjemnějších formátů je CSV:
```
actors = pandas.read_csv('static/actors.csv', index_col=None)
actors
```
Případně lze tabulku vytvořit ze seznamu seznamů:
```
items = pandas.DataFrame([
["Book", 123],
["Computer", 2185],
])
items
```
…nebo seznamu slovníků:
```
items = pandas.DataFrame([
{"name": "Book", "price": 123},
{"name": "Computer", "price": 2185},
])
items
```
V Jupyter Notebooku se tabulka vykreslí „graficky”.
V konzoli se vypíše textově, ale data v ní jsou stejná:
```
print(actors)
```
Základní informace o tabulce se dají získat metodou `info`:
```
actors.info()
```
Vidíme, že je to tabulka (`DataFrame`), má 6 řádků indexovaných
(pomocí automaticky vygenerovaného indexu) od 0 do 5
a 3 sloupce: jeden s objekty, jeden s `int64` a jeden s `bool`.
Tyto datové typy (`dtypes`) se doplnily automaticky podle zadaných
hodnot. Pandas je používá hlavně pro šetření pamětí: pythonní objekt
typu `bool` zabírá v paměti desítky bytů, ale v `bool` sloupci
si každá hodnota vystačí s jedním bytem.
Na rozdíl od NumPy jsou typy dynamické: když do sloupce zapíšeme „nekompatibilní”
hodnotu, kterou Pandas neumí převést na daný typ, typ sloupce
se automaticky zobecní.
Některé automatické převody ovšem nemusí být úplně intuitivní, např. `None` na `NaN`.
## Sloupce
Sloupec, neboli `Series`, je druhý základní datový typ v Pandas. Obsahuje sérii hodnot, jako seznam, ale navíc má jméno, datový typ a „index”, který jednotlivé hodnoty pojmenovává. Sloupce se dají získat vybráním z tabulky:
```
birth_years = actors['birth']
birth_years
type(birth_years)
birth_years.name
birth_years.index
birth_years.dtype
```
S informacemi ve sloupcích se dá počítat.
Základní aritmetické operace (jako sčítání či dělení) se sloupcem a skalární hodnotou (číslem, řetězcem, ...) provedou danou operaci nad každou hodnotou ve sloupci. Výsledek je nový sloupec:
```
ages = 2016 - birth_years
ages
century = birth_years // 100 + 1
century
```
To platí jak pro aritmetické operace (`+`, `-`, `*`, `/`, `//`, `%`, `**`), tak pro porovnávání:
```
birth_years > 1940
birth_years == 1940
```
Když sloupec nesečteme se skalární hodnotou (číslem) ale sekvencí, např. seznamem nebo dalším sloupcem, operace se provede na odpovídajících prvcích. Sloupec a druhá sekvence musí mít stejnou délku.
```
actors['name'] + [' (1)', ' (2)', ' (3)', ' (4)', ' (5)', ' (6)']
```
Řetězcové operace se u řetězcových sloupců schovávají pod jmenným prostorem `str`:
```
actors['name'].str.upper()
```
... a operace s daty a časy (*datetime*) najdeme pod `dt`.
Ze slupců jdou vybírat prvky či podsekvence podobně jako třeba ze seznamů:
```
birth_years[2]
birth_years[2:-2]
```
A navíc je lze vybírat pomocí sloupce typu `bool`, což vybere ty záznamy, u kterých je odpovídající hodnota *true*. Tak lze rychle vybrat hodnoty, které odpovídají nějaké podmínce:
```
# Roky narození po roce 1940
birth_years[birth_years > 1940]
```
Protože Python neumožňuje předefinovat chování operátorů `and` a `or`, logické spojení operací se tradičně dělá přes bitové operátory `&` (a) a `|` (nebo). Ty mají ale neintuitivní prioritu, proto se jednotlivé výrazy hodí uzavřít do závorek:
```
# Roky narození v daném rozmezí
birth_years[(birth_years > 1940) & (birth_years < 1943)]
```
Sloupce mají zabudovanou celou řadu operací, od základních (např. `column.sum()`, která bývá rychlejší než vestavěná funkce `sum()`) po roztodivné statistické specialitky. Kompletní seznam hledejte v [dokumentaci](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.Series.html). Povědomí o operacích, které sloupce umožňují, je základní znalost datového analytika.
```
print('Součet: ', birth_years.sum())
print('Průměr: ', birth_years.mean())
print('Medián: ', birth_years.median())
print('Počet unikátních hodnot: ', birth_years.nunique())
print('Koeficient špičatosti: ', birth_years.kurtosis())
```
Zvláště mocná je metoda `apply`, která nám dovoluje aplikovat jakoukoli funkci na všechny hodnoty sloupce:
```
actors['name'].apply(lambda x: ''.join(reversed(x)))
actors['alive'].apply({True: 'alive', False: 'deceased'}.get)
```
## Tabulky a vybírání prvků
Prvky ze sloupců jdou vybírat jako u seznamů. Ale z tabulek v Pandas jde vybírat spoustou různých způsobů. Tradiční hranaté závorky plní několik funkcí najednou, takže někdy není na první pohled jasné, co jaké indexování znamená:
```
actors['name'] # Jméno sloupce
actors[1:-1] # Interval řádků
actors[['name', 'alive']] # Seznam sloupců
```
Toto je příklad nejednoznačného chování, které zjednodušuje život datovým analytikům, pro které je knihovna Pandas primárně určena.
My, coby programátoři píšící robustní kód, budeme čisté indexování (`[]`) používat *jen* pro výběr sloupců podle jména.
Pro ostatní přístup použijeme tzv. *indexery*, jako `loc` a `iloc`.
### Indexer `loc`
Indexer `loc` zprostředkovává primárně *řádky*, a to podle *indexu*, tedy hlaviček tabulky. V našem příkladu jsou řádky očíslované a sloupce pojmenované, ale dále uvidíme, že v obou indexech můžou být jakékoli hodnoty.
```
actors
actors.loc[2]
```
Všimněte si, že `loc` není metoda: používají se s ním hranaté závorky.
Použijeme-li k indexování *n*-tici, prvním prvkem se indexují řádky a druhým sloupce – podobně jako u NumPy:
```
actors.loc[2, 'birth']
```
Na obou pozicích může být „interval”, ale na rozdíl od klasického Pythonu jsou ve výsledku obsaženy *obě koncové hodnoty*. (S indexem, který nemusí být vždy číselný, to dává smysl.)
```
actors.loc[2:4, 'birth':'alive']
```
Když uvedeme jen jednu hodnotu, sníží se dimenzionalita – z tabulky na sloupec (případně řádek – taky Series), ze sloupce na skalární hodnotu. Porovnejte:
```
actors.loc[2:4, 'name']
actors.loc[2:4, 'name':'name']
```
Chcete-li vybrat sloupec, na místě řádků uveďte dvojtečku – t.j. kompletní interval.
```
actors.loc[:, 'alive']
```
Další možnost indexování je seznamem hodnot. Tím se dají řádky či sloupce vybírat, přeskupovat, nebo i duplikovat:
```
actors.loc[:, ['name', 'alive']]
actors.loc[[3, 2, 4, 4], :]
```
### Indexer `iloc`
Druhý indexer, který si v krátkosti ukážeme, je `iloc`. Umí to samé co `loc`, jen nepracuje s klíčem, ale s pozicemi řádků či sloupců. Funguje tedy jako indexování v NumPy.
```
actors
actors.iloc[0, 0]
```
Protože `iloc` pracuje s čísly, záporná čísla a intervaly fungují jako ve standardním Pythonu:
```
actors.iloc[-1, 1]
actors.iloc[:, 0:1]
```
Indexování seznamem ale funguje jako u `loc`:
```
actors.iloc[[0, -1, 3], [-1, 1, 0]]
```
Jak `loc` tak `iloc` fungují i na sloupcích (Series), takže se dají kombinovat:
```
actors.iloc[-1].loc['name']
```
## Indexy
V minulé sekci jsme naťukli indexy – jména jednotlivých sloupců nebo řádků. Teď se podívejme, co všechno s nimi lze dělat.
Načtěte si znovu stejnou tabulku:
```
actors = pandas.read_csv('static/actors.csv', index_col=None)
actors
```
Tato tabulka má dva klíče: jeden pro řádky, `index`, a druhý pro sloupce, který se jmenuje `columns`.
```
actors.index
actors.columns
```
Klíč se dá změnit tím, že do něj přiřadíme sloupec (nebo jinou sekvenci):
```
actors.index = actors['name']
actors
actors.index
```
Potom jde pomocí tohoto klíče vyhledávat. Chceme-li vyhledávat efektivně (což dává smysl, pokud by řádků byly miliony), je dobré nejdřív tabulku podle indexu seřadit:
```
actors = actors.sort_index()
actors
actors.loc[['Eric', 'Graham']]
```
Pozor ale na situaci, kdy hodnoty v klíči nejsou unikátní. To Pandas podporuje, ale chování nemusí být podle vašich představ:
```
actors.loc['Terry']
```
Trochu pokročilejší možnost, jak klíč nastavit, je metoda `set_index`. Nejčastěji se používá k přesunutí sloupců do klíče, ale v [dokumentaci](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.set_index.html) se dočtete i o dalších možnostech.
Přesuňte teď do klíče dva sloupce najednou:
```
indexed_actors = actors.set_index(['name', 'birth'])
indexed_actors
```
Vznikl tím víceúrovňový klíč:
```
indexed_actors.index
```
Řádky z tabulky s víceúrovňovým klíčem se dají vybírat buď postupně po jednotlivých úrovních, nebo *n*-ticí:
```
indexed_actors.loc['Terry']
indexed_actors.loc['Terry'].loc[1940]
indexed_actors.loc[('Terry', 1942)]
```
Kromě výběru dat mají klíče i jinou vlastnost: přidáme-li do tabulky nový sloupec s klíčem, jednotlivé řádky se seřadí podle něj:
```
indexed_actors
last_names = pandas.Series(['Gilliam', 'Jones', 'Cleveland'],
index=[('Terry', 1940), ('Terry', 1942), ('Carol', 1942)])
last_names
indexed_actors['last_name'] = last_names
indexed_actors
```
## NaN neboli NULL či N/A
V posledním příkladu vidíme, že Pandas doplňuje za neznámé hodnoty `NaN`, tedy "Not a Number" – hodnotu, která plní podobnou funkci jako `NULL` v SQL nebo `None` v Pythonu. Znamená, že daná informace chybí, není k dispozici nebo ani nedává smysl ji mít. Naprostá většina operací s `NaN` dává opět `NaN`:
```
'(' + indexed_actors['last_name'] + ')'
```
NaN se chová divně i při porovnávání; `(NaN == NaN)` je nepravda. Pro zjištění chybějících hodnot máme metodu `isnull()`:
```
indexed_actors['last_name'].isnull()
```
Abychom se `NaN` zbavili, máme dvě možnosti. Buď je zaplníme pomocí metody [`fillna`](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.fillna.html) hodnotou jako `0`, `False` nebo, pro přehlednější výpis, prázdným řetězcem:
```
indexed_actors.fillna('')
```
Nebo se můžeme zbavit všech řádků, které nějaký `NaN` obsahují:
```
indexed_actors.dropna()
```
Bohužel existuje jistá nekonzistence mezi `NaN` a slovy `null` či `na` v názvech funkcí. *C'est la vie.*
## Merge
Někdy se stane, že máme více souvisejících tabulek, které je potřeba spojit dohromady. Na to mají `DataFrame` metodu `merge()`, která umí podobné operace jako `JOIN` v SQL.
```
actors = pandas.read_csv('static/actors.csv', index_col=None)
actors
spouses = pandas.read_csv('static/spouses.csv', index_col=None)
spouses
actors.merge(spouses)
```
Mají-li spojované tabulky sloupce stejných jmen, Pandas je spojí podle těchto sloupců. V [dokumentaci](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.merge.html) se dá zjistit, jak explicitně určit podle kterých klíčů spojovat, co udělat když v jedné z tabulek chybí odpovídající hodnoty apod.
Fanoušky SQL ještě odkážu na [porovnání mezi SQL a Pandas](http://pandas.pydata.org/pandas-docs/stable/comparison_with_sql.html).
## Přesýpání dat
Dostáváme se do bodu, kdy nám jednoduchá tabulka přestává stačit. Pojďme si vytvořit tabulku větší: fiktivních prodejů v e-shopu, ve formátu jaký bychom mohli dostat z SQL databáze nebo datového souboru.
Použijeme k tomu mimo jiné `date_range`, která vytváří kalendářní intervaly. Zde, i v jiných případech, kdy je jasné, že se má nějaká hodnota interpretovat jako datum, nám Pandas dovolí místo objektů `datetime` zadávat data řetězcem:
```
import itertools
import random
random.seed(0)
months = pandas.date_range('2015-01', '2016-12', freq='M')
categories = ['Electronics', 'Power Tools', 'Clothing']
data = pandas.DataFrame([{'month': a, 'category': b, 'sales': random.randint(-1000, 10000)}
for a, b in itertools.product(months, categories)
if random.randrange(20) > 0])
```
Tabulka je celkem dlouhá (i když v analýze dat bývají ještě delší). Podívejme se na několik obecných informací:
```
# Prvních pár řádků (dá se použít i např. head(10), bylo by jich víc)
data.head()
# Celkový počet řádků
len(data)
data['sales'].describe()
```
Pomocí `set_index` nastavíme, které sloupce budeme brát jako hlavičky:
```
indexed = data.set_index(['category', 'month'])
indexed.head()
```
Budeme-li chtít z těchto dat vytvořit tabulku, která má v řádcích kategorie a ve sloupcích měsíce, můžeme využít metodu `unstack`, která "přesune" vnitřní úroveň indexu řádků do sloupců a uspořádá podle toho i data.
Můžeme samozřejmě použít kteroukoli úroveň klíče; viz [dokumentace](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.unstack.html) k `unstack` a reverzní operaci [`stack`](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.stack.html).
```
unstacked = indexed.unstack('month')
unstacked
```
Teď je sloupcový klíč dvouúrovňový, ale úroveň `sales` je zbytečná. Můžeme se jí zbavit pomocí [`MultiIndex.droplevel`](http://pandas.pydata.org/pandas-docs/version/0.18.0/generated/pandas.MultiIndex.droplevel.html).
```
unstacked.columns = unstacked.columns.droplevel()
unstacked
```
A teď můžeme data analyzovat. Kolik se celkem utratilo za elektroniku?
```
unstacked.loc['Electronics'].sum()
```
Jak to vypadalo se všemi elektrickými zařízeními v třech konkrétních měsících?
```
unstacked.loc[['Electronics', 'Power Tools'], '2016-03':'2016-05']
```
A jak se prodávalo oblečení?
```
unstacked.loc['Clothing']
```
Metody `stack` a `unstack` jsou sice asi nejužitečnější, ale stále jen jeden ze způsobů jak v Pandas tabulky přeskládávat. Náročnější studenti najdou další možnosti v [dokumentaci](http://pandas.pydata.org/pandas-docs/stable/reshaping.html).
## Grafy
Je-li nainstalována knihovna `matplotlib`, Pandas ji umí využít k tomu, aby kreslil grafy. Nastavení je trochu jiné pro Jupyter Notebook a pro příkazovou řádku.
Používáte-li Jupyter Notebook, zapněte integraci pro kreslení grafů pomocí:
```
import matplotlib
# Zapnout zobrazování grafů (procento uvozuje „magickou” zkratku IPythonu):
%matplotlib inline
```
a pak můžete přímo použít metodu `plot()`, která bez dalších argumentů vynese data z tabulky proti indexu:
```
unstacked.loc['Clothing'].dropna().plot()
```
Jste-li v příkazové řádce, napřed použij `plot()` a potom se na graf buď podívete, nebo ho uložte:
```python
# Setup
import matplotlib.pyplot
# Plot
unstacked.loc['Clothing'].plot()
matplotlib.pyplot.show()
matplotlib.pyplot.savefig('graph.png')
```
Funkce `show` a `savefig` pracují s „aktuálním” grafem – typicky posledním, který se vykreslil. Pozor na to, že funkce `savefig` aktuální graf zahodí; před dalším `show` nebo `savefig` je potřeba ho vykreslit znovu.
V kombinaci s dalšími funkcemi `Series` a `DataFrame` umožňují grafy získat o datech rychlý přehled:
```
# Jak se postupně vyvíjely zisky z oblečení?
# `.T` udělá transpozici tabulky (vymění řádky a sloupce)
# `cumsum()` spočítá průběžný součet po sloupcích
unstacked.T.fillna(0).cumsum().plot()
# Jak si proti sobě stály jednotlivé kategorie v březnu, dubnu a květnu 2016?
unstacked.loc[:, '2016-03':'2016-05'].plot.bar(legend=False)
```
Další informace jsou, jak už to bývá, [v dokumentaci](http://pandas.pydata.org/pandas-docs/version/0.19.0/visualization.html).
## Groupby
Často používaná operace pro zjednodušení tabulky je `groupby`, která sloučí dohromady řádky se stejnou hodnotou v některém sloupci a sloučená data nějak agreguje.
```
data.head()
```
Samotný výsledek `groupby()` je jen objekt:
```
data.groupby('category')
```
... na který musíme zavolat příslušnou agregující funkci. Tady je například součet částek podle kategorie:
```
data.groupby('category').sum()
```
Nebo počet záznamů:
```
data.groupby('category').count()
```
Groupby umí agregovat podle více sloupců najednou (i když u našeho příkladu nedává velký smysl):
```
data.groupby(['category', 'month']).sum().head()
```
Chceme-li aplikovat více funkcí najednou, předáme jejich seznam metodě `agg`. Časté funkce lze předat jen jménem, jinak předáme funkci či metodu přímo:
```
data.groupby('category').agg(['mean', 'median', sum, pandas.Series.kurtosis])
```
Případně použijeme zkratku pro základní analýzu:
```
g = data.groupby('month')
g.describe()
```
A perlička nakonec – agregovat se dá i podle sloupců, které nejsou v tabulce. Následující kód rozloží data na slabé, průměrné a silné měsíce podle toho, kolik jsme v daném měsíci vydělali celých tisícikorun, a zjistí celkový zisk ze slabých, průměrných a silných měsíců:
```
bin_size = 10000
by_month = data.groupby('month').sum()
by_thousands = by_month.groupby(by_month['sales'] // bin_size * bin_size).agg(['count', 'sum'])
by_thousands
by_thousands[('sales', 'sum')].plot()
```
| github_jupyter |
```
#Required libraries
import pandas as pd
import numpy as np
from sklearn import linear_model, model_selection, metrics
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeClassifier
from sklearn import tree
from sklearn import preprocessing
import pydotplus
from IPython.display import Image
#Loading data
launch_data = pd.read_excel('RocketLaunchDataCompleted.xlsx')
launch_data.head()
launch_data.columns
""" We only have 60 launches in this dataset and we have data for other days as before and after launch, this means
we will choose 60 records where the field Launched? is Y' """
launch_data.info()
"""To handle missing values, we will fill the missing values with appropriate values, for instance:
Launched? is Na -> (we'll assume that wasn't launch)
Crewed or Uncrewed is Na -> (Assume that the launch was uncrewed)
Wind condition is Na -> (Assume as unkown)
Condition is Na -> (Assume as Fair)
"""
launch_data['Launched?'].fillna('N',inplace=True)
launch_data['Crewed or Uncrewed'].fillna('Uncrewed',inplace=True)
launch_data['Wind Direction'].fillna('unknown',inplace=True)
launch_data['Condition'].fillna('Fair',inplace=True)
launch_data.fillna(0,inplace=True)
launch_data.head()
## As part of the data cleaning process, we have to convert text data to numerical because computers understand only numbers
label_encoder = preprocessing.LabelEncoder()
# Three columns have categorical text info, and we convert them to numbers
# For instance, 0 for Crewed - 1 for Uncrewed
launch_data['Crewed or Uncrewed'] = label_encoder.fit_transform(launch_data['Crewed or Uncrewed'])
launch_data['Wind Direction'] = label_encoder.fit_transform(launch_data['Wind Direction'])
launch_data['Condition'] = label_encoder.fit_transform(launch_data['Condition'])
launch_data.head()
#Building the model
# First, we save the output we are interested in. In this case, "launch" yes and no's go into the output variable.
y = launch_data['Launched?']
# Removing the columns we are not interested in
launch_data.drop(['Name','Date','Time (East Coast)','Location','Launched?','Hist Ave Sea Level Pressure','Sea Level Pressure','Day Length','Notes','Hist Ave Visibility', 'Hist Ave Max Wind Speed'],axis=1, inplace=True)
# Saving the rest of the data as input data
X = launch_data
# List of variables that our machine learning algorithm is going to look at:
X.columns
"""The X input data represents the weather for a particular day. In this case, we aren't worried about the date or time. We want the profile of the weather for that day to be the indicator for whether a launch should happen, not the date or time."""
X.head()
""" Predict a rocket launch given weather conditions is a yes or no question. This is a two-class classification problem. We'll use a decision tree given that is an accurate and fast training algorithm.
For this job we'll use sickit-learn. """
tree_model = DecisionTreeClassifier(random_state=0,max_depth=5)
#Splitting data
"""
Test size: 0.2 - 80% training size, 20% test size
Random state: 99 - random seed"""
X_train, X_test, y_train, y_test = train_test_split(X,y, test_size=0.2, random_state=99)
# Fitting the model to the training data
tree_model.fit(X_train,y_train)
#Testing the model
y_pred = tree_model.predict(X_test)
print(y_pred)
#Scoring the model
# Calculate accuracy
tree_model.score(X_test,y_test)
#Creating the visual tree
# Let's visualizing our decision tree.
from sklearn.tree import export_graphviz
def tree_graph_to_png(tree, feature_names,class_names, png_file_to_save):
tree_str = export_graphviz(tree, feature_names=feature_names, class_names=class_names,
filled=True, out_file=None)
graph = pydotplus.graph_from_dot_data(tree_str)
return Image(graph.create_png())
# This function visualizes the model.
tree_graph_to_png(tree=tree_model, feature_names=X.columns.values,class_names=['No Launch','Launch'], png_file_to_save='decision-tree.png')
"""
192 are no launches
48 are launches
Given the visualization we could think the following:
"If the wind speed was less than 1.0, then 191 of the 240 samples guessed that no launch was possible on that day."
"""
#A test for a recent lauch.
"""On July 30, 2020, NASA launched the Perseverance rover to Mars from Cape Canaveral at 7:50 AM Eastern Time. The next is hypotethical data: """
# ['Crewed or Uncrewed', 'High Temp', 'Low Temp', 'Ave Temp',
# 'Temp at Launch Time', 'Hist High Temp', 'Hist Low Temp',
# 'Hist Ave Temp', 'Precipitation at Launch Time',
# 'Hist Ave Precipitation', 'Wind Direction', 'Max Wind Speed',
# 'Visibility', 'Wind Speed at Launch Time', 'Hist Ave Max Wind Speed',
# 'Hist Ave Visibility', 'Condition']
data_input = [ 1. , 75. , 68. , 71. , 0. , 75. , 55. , 65. , 0. , 0.08, 0. , 16. , 15. , 0. , 0. ]
test = tree_model.predict([data_input])
print(test)
```
| github_jupyter |
___
<a href='http://www.pieriandata.com'> <img src='../../Pierian_Data_Logo.png' /></a>
___
# Ecommerce Purchases Exercise
In this Exercise you will be given some Fake Data about some purchases done through Amazon! Just go ahead and follow the directions and try your best to answer the questions and complete the tasks. Feel free to reference the solutions. Most of the tasks can be solved in different ways. For the most part, the questions get progressively harder.
Please excuse anything that doesn't make "Real-World" sense in the dataframe, all the data is fake and made-up.
Also note that all of these questions can be answered with one line of code.
____
** Import pandas and read in the Ecommerce Purchases csv file and set it to a DataFrame called ecom. **
```
import pandas as pd
df = pd.read_csv('Ecommerce Purchases')
```
**Check the head of the DataFrame.**
```
df.head()
```
** How many rows and columns are there? **
```
df.info()
```
** What is the average Purchase Price? **
```
df['Purchase Price'].mean()
```
** What were the highest and lowest purchase prices? **
```
df['Purchase Price'].max()
df['Purchase Price'].min()
```
** How many people have English 'en' as their Language of choice on the website? **
```
df[df['Language']=='en'].count()
```
** How many people have the job title of "Lawyer" ? **
```
df[df['Job']=='Lawyer'].count()
```
** How many people made the purchase during the AM and how many people made the purchase during PM ? **
**(Hint: Check out [value_counts()](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.Series.value_counts.html) ) **
```
df['AM or PM'].value_counts()
```
** What are the 5 most common Job Titles? **
```
df['Job'].value_counts().head(5)
```
** Someone made a purchase that came from Lot: "90 WT" , what was the Purchase Price for this transaction? **
```
df[df['Lot']=='90 WT']['Purchase Price']
```
** What is the email of the person with the following Credit Card Number: 4926535242672853 **
```
df[df['Credit Card']==4926535242672853]['Email']
```
** How many people have American Express as their Credit Card Provider *and* made a purchase above $95 ?**
```
df[(df['CC Provider']=='American Express') & (df['Purchase Price']>95)].count()
```
** Hard: How many people have a credit card that expires in 2025? **
```
df['CC Exp Date'].head()
%timeit sum(df['CC Exp Date'].str.endswith('/25'))
```
** Hard: What are the top 5 most popular email providers/hosts (e.g. gmail.com, yahoo.com, etc...) **
```
df['Email'].str.extract('(@\w+\.\w+)', expand=False).value_counts().head(5)
```
# Great Job!
| github_jupyter |
# Network Control - Thaliana
control measures for the Arabidobis Thaliana Boolean Model
```
%load_ext autoreload
%autoreload 2
%matplotlib inline
from __future__ import division
import numpy as np
from itertools import product
from cana.control import fvs, mds, sc
from cana.datasets.bio import THALIANA
from IPython.display import display
from IPython.core.display import HTML
N = THALIANA()
```
## Discovering Driver Nodes / Control Nodes
```
display(HTML('<h3>Control State Transition Graph (CSTG)</h3>'))
# THIS MIGHT TAKE A LONG TIME, it is here for demo purposes.
driver_nodes = N.attractor_driver_nodes(min_dvs=1, max_dvs=6, verbose=True)
print(N.get_node_name(driver_nodes))
#> ['AP2', 'EMF1', 'LFY', 'TFL1', 'UFO', 'WUS'], ['AG', 'EMF1', 'LFY', 'TFL1', 'UFO', 'WUS']
display(HTML('<h3>Structural Controlability (SC)</h3>'))
SC = N.structural_controllability_driver_nodes(keep_self_loops=False)
print(N.get_node_name(SC))
display(HTML('<h3>Minimum Dominating Set (MDS)</h3>'))
MDS = N.minimum_dominating_set_driver_nodes(max_search=10)
print(N.get_node_name(MDS))
display(HTML('<h3>Feedback Vertex Control (FVS)</h3>'))
FVS_g = N.feedback_vertex_set_driver_nodes(graph='structural', method='grasp', remove_constants=True)
print(N.get_node_name(FVS_g) , '(grasp)')
FVS_bf = N.feedback_vertex_set_driver_nodes(graph='structural', method='bruteforce', remove_constants=True)
print(N.get_node_name(FVS_bf) , '(bruteforce)')
#display(HTML('<h3>Pinning Control (PC)</h3>'))
#under development
```
## Differences between the Control Methods
```
# Control via State Transition Graph (CSTG)
CSTGs = [['AP2', 'EMF1', 'LFY', 'TFL1', 'UFO', 'WUS'],
['AG', 'EMF1', 'LFY', 'TFL1', 'UFO', 'WUS']
]
# Pinning Control
PCs = [['AP3', 'UFO', 'AP1', 'LFY', 'WUS', 'AG'],
['AP3', 'UFO', 'EMF1', 'WUS', 'AG', 'TFL1'],
['AP3', 'UFO', 'LFY', 'WUS', 'AG', 'TFL1']
]
# Feedback Vertex Control
# (threshold,loops,[control sets])
FVS_Objs = [
('original',49, [['AP3','UFO','LFY','WUS','AG','TFL1','PI']]),
('0',19, [['AP3','UFO','AP1','LFY','WUS' 'AG','PI'],['AP3','UFO','EMF1','WUS','AG','TFL1','PI'],['AP3','UFO','LFY','WUS','AG','TFL1','PI']]),
('0.0078125',17,[['AP3', 'UFO', 'AP1', 'LFY', 'WUS', 'PI']]),
('0.0234375',14,[['AP3', 'UFO', 'AP1', 'LFY', 'WUS', 'PI'],['AP3', 'UFO', 'EMF1', 'WUS', 'TFL1', 'PI'],['AP3', 'UFO', 'LFY', 'WUS', 'TFL1', 'PI']]),
('0.03125',14,[['AP3', 'UFO', 'AP1', 'LFY', 'WUS', 'PI'],['AP3', 'UFO', 'EMF1', 'WUS', 'TFL1', 'PI'],['AP3', 'UFO', 'LFY', 'WUS', 'TFL1', 'PI']]),
('0.046875',14,[['AP3', 'UFO', 'AP1', 'LFY', 'WUS', 'PI'],['AP3', 'UFO', 'EMF1', 'WUS', 'TFL1', 'PI'],['AP3', 'UFO', 'LFY', 'WUS', 'TFL1', 'PI']]),
('0.09375',13,[['AP3', 'UFO', 'AP1', 'LFY', 'WUS'],['AP3', 'UFO', 'EMF1', 'WUS', 'TFL1'],['AP3', 'UFO', 'LFY', 'WUS', 'TFL1']]),
('0.125',10,[['AP3', 'UFO', 'AP1', 'LFY', 'WUS'],['AP3', 'UFO', 'EMF1', 'WUS', 'TFL1'],['AP3', 'UFO', 'LFY', 'WUS', 'AG'],['AP3', 'UFO', 'LFY', 'WUS', 'TFL1']]),
('0.140625',8,[['UFO', 'AP1', 'LFY', 'WUS'],['UFO', 'EMF1', 'WUS', 'TFL1'],['UFO', 'LFY', 'WUS', 'AG'],['UFO', 'LFY', 'WUS', 'TFL1']]),
('0.25',3,[['UFO', 'EMF1', 'WUS', 'TFL1'],['UFO', 'LFY', 'WUS', 'TFL1']]),
('0.2734375',3,[['UFO', 'EMF1', 'WUS', 'TFL1'],['UFO', 'LFY', 'WUS', 'TFL1']]),
('0.28125',3,[['UFO', 'EMF1', 'WUS', 'TFL1'],['UFO', 'LFY', 'WUS', 'TFL1']]),
('0.34375',3,[['UFO', 'EMF1', 'WUS', 'TFL1'],['UFO', 'LFY', 'WUS', 'TFL1']]),
('0.453125',3,[['AP3', 'UFO', 'EMF1', 'WUS', 'TFL1'],['AP3', 'UFO', 'LFY', 'WUS', 'TFL1']]),
('0.5',2,[['AP3', 'UFO', 'FUL', 'LFY', 'WUS', 'TFL1']]),
('0.65625',2,[['AP3', 'UFO', 'FUL', 'LFY', 'WUS', 'TFL1', 'PI']]),
('0.7265625',2,[['AP3', 'UFO', 'FUL', 'LFY', 'WUS', 'AG', 'TFL1', 'PI']]),
('0.75',1,[['AP3', 'UFO', 'FUL', 'LFY', 'WUS', 'AG', 'TFL1', 'PI']]),
('0.875',1,[['AP3', 'UFO', 'FUL', 'AP1', 'LFY', 'WUS', 'AG', 'TFL1', 'PI']])
]
# Sort sets by alphabetical order
CSTGs = [sorted(x) for x in CSTGs]
PCs = [sorted(x) for x in PCs]
FVS_Objs = [(name,loops,[sorted(x) for x in sets]) for (name,loops,sets) in FVS_Objs]
def jaccard(u,v):
return len(u.intersection(v)) / len(u.union(v))
display(HTML("<h2>PC vs FVS</h2>"))
for FVS_O in FVS_Objs:
print('T: %s' % (FVS_O[0]))
for PC, FVS in product(PCs,FVS_O[2]):
FVSset, PCset = set(FVS), set(PC)
FVSstr, PCstr = ','.join(FVS), ','.join(PC)
inclusion = PCset.issubset(FVSset)
print('PC in FVS=%i ; J=%.3f [%s <-> %s]' % (inclusion, jaccard(PCset,FVSset) , PCstr , FVSstr))
print
display(HTML("<h2>CSTG vs FVS</h2>"))
for FVS_O in FVS_Objs:
print('T: %s' % (FVS_O[0]))
for CSTG, FVS in product(CSTGs,FVS_O[2]):
FVSset, CSTGset = set(FVS), set(CSTG)
FVSstr, CSTGstr = ','.join(FVS), ','.join(CSTG)
inclusion = CSTGset.issubset(FVSset)
print('CSTG in FVS=%i ; J=%.3f [%s <-> %s]' % (inclusion, jaccard(CSTGset,FVSset) , CSTGstr , FVSstr))
print
```
| github_jupyter |
```
import bs4
import requests
from time import sleep
datastorage=[]
for z in range(7,35):
pagelink='https://attorneys.superlawyers.com/family-law/illinois/chicago/page'+str(z)
print pagelink
mainpage=requests.get(pagelink)
so0=bs4.BeautifulSoup(mainpage.text,'lxml')
sleep(3)
indigo_text=so0.find_all('h2',class_="indigo_text")
for x in range(0,len(indigo_text)):
profile_url=indigo_text[x].find('a',class_="directory_profile").get('href')
print profile_url
page=requests.get(profile_url)
so1=bs4.BeautifulSoup(page.text,'lxml')
sleep(3)
datastr=['-']*25
lawyer_bio_block=so1.find_all('div',id="lawyer_bio_block")[0]
lawyer_name=lawyer_bio_block.find('h2',id="lawyer_name").text
datastr[0]=lawyer_name
attorney_profile_heading=lawyer_bio_block.find('h1',id="attorney_profile_heading").text
datastr[1]=attorney_profile_heading
firm_name=lawyer_bio_block.find('a',id="firm_profile_page").text
datastr[2]=firm_name
poap_postal_addr_block=lawyer_bio_block.find('div',id="poap_postal_addr_block")
datastr[3]=poap_postal_addr_block.text
websiteheader=lawyer_bio_block.find('a',class_="profile-website-header")
if websiteheader is not None:
website=websiteheader.text
datastr[4]=website
contact_info_blocks=lawyer_bio_block.find_all('div',class_="contact_info_blocks")
for q in range(len(contact_info_blocks)):
if 'Phone' in contact_info_blocks[q].text:
Phone=contact_info_blocks[q].text
datastr[5]=Phone
if 'Fax' in contact_info_blocks[q].text:
Fax=contact_info_blocks[q].text
datastr[6]=Fax
licensed_since0=lawyer_bio_block.find('span',id="licensed_since")
if licensed_since0 is not None:
licensed_since=licensed_since0.text
datastr[7]=licensed_since
Education0=lawyer_bio_block.find('span',id="law_school_content")
if Education0 is not None:
Education=Education0.text
datastr[8]=Education
practice_areas=lawyer_bio_block.find('div',id="practice_areas").text
datastr[9]=practice_areas
focus_areas_wrapper=so1.find('div',id="focus_areas_wrapper")
if focus_areas_wrapper is not None:
focus_areas=focus_areas_wrapper.text
datastr[10]=focus_areas
lawyer_image=lawyer_bio_block.find('div',id="lawyer_image").find('img').get('src')
datastr[11]=lawyer_image
attorney_profile_text0=so1.find('div',id="attorney_profile_text")
if attorney_profile_text0 is not None:
attorney_profile_text=attorney_profile_text0.text[1:]
datastr[12]=attorney_profile_text
poap_about_mel=so1.find('div',id="poap_about_me")
if poap_about_mel is not None:
poap_about_me_div=poap_about_mel.find('div',id="about_me_vitae").find_all('div')
for t in range(1,len(poap_about_me_div)):
if 'Educational Background:' in poap_about_me_div[t].text:
poap_about_me_div_li=poap_about_me_div[t].find_all('li')
Educational_Background=''
for r in range(len(poap_about_me_div_li)):
Educational_Background+=str(r)+':'+poap_about_me_div_li[r].text+'|'
datastr[13]=Educational_Background
if 'Honors/Awards:' in poap_about_me_div[t].text:
poap_about_me_div_li=poap_about_me_div[t].find_all('li')
Honors_Awards=''
for r in range(len(poap_about_me_div_li)):
Honors_Awards+=str(r)+':'+poap_about_me_div_li[r].text+'|'
datastr[14]=Honors_Awards
if 'Bar/Professional Activity:' in poap_about_me_div[t].text:
poap_about_me_div_li=poap_about_me_div[t].find_all('li')
Bar_Professional_Activity=''
for r in range(len(poap_about_me_div_li)):
Bar_Professional_Activity+=str(r)+':'+poap_about_me_div_li[r].text+'|'
datastr[15]=Bar_Professional_Activity
if 'Pro bono/Community Service:' in poap_about_me_div[t].text:
poap_about_me_div_li=poap_about_me_div[t].find_all('li')
Pro_bono_Community_Service=''
for r in range(len(poap_about_me_div_li)):
Pro_bono_Community_Service+=str(r)+':'+poap_about_me_div_li[r].text+'|'
datastr[16]=Pro_bono_Community_Service
if 'Special Licenses/Certifications:' in poap_about_me_div[t].text:
poap_about_me_div_li=poap_about_me_div[t].find_all('li')
Special_Licenses_Certifications=''
for r in range(len(poap_about_me_div_li)):
Special_Licenses_Certifications+=str(r)+':'+poap_about_me_div_li[r].text+'|'
datastr[17]=Special_Licenses_Certifications
if 'Scholarly Lectures/Writings:' in poap_about_me_div[t].text:
poap_about_me_div_li=poap_about_me_div[t].find_all('li')
Scholarly_Lectures_Writings=''
for r in range(len(poap_about_me_div_li)):
Scholarly_Lectures_Writings+=str(r)+':'+poap_about_me_div_li[r].text+'|'
datastr[18]=Scholarly_Lectures_Writings
if 'Other Outstanding Achievements:' in poap_about_me_div[t].text:
poap_about_me_div_li=poap_about_me_div[t].find_all('li')
Other_Outstanding_Achievements=''
for r in range(len(poap_about_me_div_li)):
Other_Outstanding_Achievements+=str(r)+':'+poap_about_me_div_li[r].text+'|'
datastr[19]=Other_Outstanding_Achievements
poap_about_mekk=so1.find('div',id="poap_about_me")
if poap_about_mekk is not None:
poap_about_me_soiallinks=poap_about_mekk.find_all('a')
for e in range(len(poap_about_me_soiallinks)):
if poap_about_me_soiallinks[e].get('title')=='Website':
website=poap_about_me_soiallinks[e].get('href')
datastr[20]=website
if poap_about_me_soiallinks[e].get('title')=='Facebook':
Facebook=poap_about_me_soiallinks[e].get('href')
datastr[21]=Facebook
if poap_about_me_soiallinks[e].get('title')=='LinkedIn':
LinkedIn=poap_about_me_soiallinks[e].get('href')
datastr[22]=LinkedIn
if poap_about_me_soiallinks[e].get('title')=='Twitter':
Twitter=poap_about_me_soiallinks[e].get('href')
datastr[23]=Twitter
datastr[24]=profile_url
datastorage.append(datastr)
print z,x,len(datastorage)
print '-----------------'
!tar cvfz zipname.tar.gz *
import warnings
from openpyxl import Workbook
wb = Workbook(write_only=True)
ws = wb.create_sheet()
# now we'll fill it with 100 rows x 200 columns
for irow in datastorage:
ws.append(irow)
# save the file
wb.save('family-law1.xlsx')
from openpyxl import load_workbook
wb = load_workbook(filename = 'family-law.xlsx')
sheet_ranges = wb['Sheet']
linksto_profiles=[]
for row in sheet_ranges:
linksto_profiles.append(row[24].value)
len(linksto_profiles)
linksto_profiles=linksto_profiles[65:]
linksto_profiles[0]
len(linksto_profiles)
datastorage=[]
for x in range(88,len(linksto_profiles)):
profile_url=linksto_profiles[x]
print profile_url
page=requests.get(profile_url)
so1=bs4.BeautifulSoup(page.text,'lxml')
sleep(3)
datastr=['-']*25
lawyer_bio_block=so1.find_all('div',id="lawyer_bio_block")[0]
lawyer_name=lawyer_bio_block.find('h2',id="lawyer_name").text
datastr[0]=lawyer_name
attorney_profile_heading=lawyer_bio_block.find('h1',id="attorney_profile_heading").text
datastr[1]=attorney_profile_heading
firm_name=lawyer_bio_block.find('a',id="firm_profile_page").text
datastr[2]=firm_name
poap_postal_addr_block=lawyer_bio_block.find('div',id="poap_postal_addr_block")
datastr[3]=poap_postal_addr_block.text
websiteheader=lawyer_bio_block.find('a',class_="profile-website-header")
if websiteheader is not None:
website=websiteheader.text
datastr[4]=website
contact_info_blocks=lawyer_bio_block.find_all('div',class_="contact_info_blocks")
for q in range(len(contact_info_blocks)):
if 'Phone' in contact_info_blocks[q].text:
Phone=contact_info_blocks[q].text
datastr[5]=Phone
if 'Fax' in contact_info_blocks[q].text:
Fax=contact_info_blocks[q].text
datastr[6]=Fax
licensed_since0=lawyer_bio_block.find('span',id="licensed_since")
if licensed_since0 is not None:
licensed_since=licensed_since0.text
datastr[7]=licensed_since
Education0=lawyer_bio_block.find('span',id="law_school_content")
if Education0 is not None:
Education=Education0.text
datastr[8]=Education
practice_areas=lawyer_bio_block.find('div',id="practice_areas").text
datastr[9]=practice_areas
focus_areas_wrapper=so1.find('div',id="focus_areas_wrapper")
if focus_areas_wrapper is not None:
focus_areas=focus_areas_wrapper.text
datastr[10]=focus_areas
lawyer_image=lawyer_bio_block.find('div',id="lawyer_image").find('img').get('src')
datastr[11]=lawyer_image
attorney_profile_text0=so1.find('div',id="attorney_profile_text")
if attorney_profile_text0 is not None:
attorney_profile_text=attorney_profile_text0.text[1:]
datastr[12]=attorney_profile_text
poap_about_mel=so1.find('div',id="eoap_about_me")
if poap_about_mel is not None:
poap_about_me_div0=poap_about_mel.find('div',id="about_me_vitae")
if poap_about_me_div0 is not None:
poap_about_me_div=poap_about_me_div0.find_all('div')
for t in range(1,len(poap_about_me_div)):
if 'Educational Background:' in poap_about_me_div[t].text:
poap_about_me_div_li=poap_about_me_div[t].find_all('li')
Educational_Background=''
for r in range(len(poap_about_me_div_li)):
Educational_Background+=str(r)+':'+poap_about_me_div_li[r].text+'|'
datastr[13]=Educational_Background
if 'Honors/Awards:' in poap_about_me_div[t].text:
poap_about_me_div_li=poap_about_me_div[t].find_all('li')
Honors_Awards=''
for r in range(len(poap_about_me_div_li)):
Honors_Awards+=str(r)+':'+poap_about_me_div_li[r].text+'|'
datastr[14]=Honors_Awards
if 'Bar/Professional Activity:' in poap_about_me_div[t].text:
poap_about_me_div_li=poap_about_me_div[t].find_all('li')
Bar_Professional_Activity=''
for r in range(len(poap_about_me_div_li)):
Bar_Professional_Activity+=str(r)+':'+poap_about_me_div_li[r].text+'|'
datastr[15]=Bar_Professional_Activity
if 'Pro bono/Community Service:' in poap_about_me_div[t].text:
poap_about_me_div_li=poap_about_me_div[t].find_all('li')
Pro_bono_Community_Service=''
for r in range(len(poap_about_me_div_li)):
Pro_bono_Community_Service+=str(r)+':'+poap_about_me_div_li[r].text+'|'
datastr[16]=Pro_bono_Community_Service
if 'Special Licenses/Certifications:' in poap_about_me_div[t].text:
poap_about_me_div_li=poap_about_me_div[t].find_all('li')
Special_Licenses_Certifications=''
for r in range(len(poap_about_me_div_li)):
Special_Licenses_Certifications+=str(r)+':'+poap_about_me_div_li[r].text+'|'
datastr[17]=Special_Licenses_Certifications
if 'Scholarly Lectures/Writings:' in poap_about_me_div[t].text:
poap_about_me_div_li=poap_about_me_div[t].find_all('li')
Scholarly_Lectures_Writings=''
for r in range(len(poap_about_me_div_li)):
Scholarly_Lectures_Writings+=str(r)+':'+poap_about_me_div_li[r].text+'|'
datastr[18]=Scholarly_Lectures_Writings
if 'Other Outstanding Achievements:' in poap_about_me_div[t].text:
poap_about_me_div_li=poap_about_me_div[t].find_all('li')
Other_Outstanding_Achievements=''
for r in range(len(poap_about_me_div_li)):
Other_Outstanding_Achievements+=str(r)+':'+poap_about_me_div_li[r].text+'|'
datastr[19]=Other_Outstanding_Achievements
poap_about_mekk=so1.find('div',id="poap_about_me")
if poap_about_mekk is not None:
poap_about_me_soiallinks=poap_about_mekk.find_all('a')
for e in range(len(poap_about_me_soiallinks)):
if poap_about_me_soiallinks[e].get('title')=='Website':
website=poap_about_me_soiallinks[e].get('href')
datastr[20]=website
if poap_about_me_soiallinks[e].get('title')=='Facebook':
Facebook=poap_about_me_soiallinks[e].get('href')
datastr[21]=Facebook
if poap_about_me_soiallinks[e].get('title')=='LinkedIn':
LinkedIn=poap_about_me_soiallinks[e].get('href')
datastr[22]=LinkedIn
if poap_about_me_soiallinks[e].get('title')=='Twitter':
Twitter=poap_about_me_soiallinks[e].get('href')
datastr[23]=Twitter
datastr[24]=profile_url
datastorage.append(datastr)
print x,len(datastorage)
print '-----------------'
import warnings
from openpyxl import Workbook
wb = Workbook(write_only=True)
ws = wb.create_sheet()
# now we'll fill it with 100 rows x 200 columns
for irow in datastorage:
ws.append(irow)
# save the file
wb.save('family-law1.xlsx')
```
| github_jupyter |
### Generating human faces with Adversarial Networks (5 points)
<img src="https://www.strangerdimensions.com/wp-content/uploads/2013/11/reception-robot.jpg" width=320>
This time we'll train a neural net to generate plausible human faces in all their subtlty: appearance, expression, accessories, etc. 'Cuz when us machines gonna take over Earth, there won't be any more faces left. We want to preserve this data for future iterations. Yikes...
Based on Based on https://github.com/Lasagne/Recipes/pull/94 .
```
import matplotlib.pyplot as plt
%matplotlib inline
import numpy as np
plt.rcParams.update({'axes.titlesize': 'small'})
from sklearn.datasets import load_digits
#The following line fetches you two datasets: images, usable for autoencoder training and attributes.
#Those attributes will be required for the final part of the assignment (applying smiles), so please keep them in mind
from lfw_dataset import fetch_lfw_dataset
data,attrs = fetch_lfw_dataset(dimx=36,dimy=36)
#preprocess faces
data = np.float32(data).transpose([0,3,1,2]) / 255.
IMG_SHAPE = data.shape[1:]
#print random image
plt.imshow(data[np.random.randint(data.shape[0])].transpose([1,2,0]),
cmap="gray", interpolation="none")
```
# Generative adversarial nets 101
<img src="https://raw.githubusercontent.com/torch/torch.github.io/master/blog/_posts/images/model.png" width=320px height=240px>
Deep learning is simple, isn't it?
* build some network that generates the face (small image)
* make up a __measure__ of __how good that face is__
* optimize with gradient descent :)
The only problem is: how can we engineers tell well-generated faces from bad? And i bet you we won't ask a designer for help.
__If we can't tell good faces from bad, we delegate it to yet another neural network!__
That makes the two of them:
* __G__enerator - takes random noize for inspiration and tries to generate a face sample.
* Let's call him __G__(z), where z is a gaussian noize.
* __D__iscriminator - takes a face sample and tries to tell if it's great or fake.
* Predicts the probability of input image being a __real face__
* Let's call him __D__(x), x being an image.
* __D(x)__ is a predition for real image and __D(G(z))__ is prediction for the face made by generator.
Before we dive into training them, let's construct the two networks.
```
# this code is only needed if you're running this code on GPU machines
# !pip install http://download.pytorch.org/whl/cu80/torch-0.3.0.post4-cp36-cp36m-linux_x86_64.whl --user --no-deps --ignore-installed
import sys, os; sys.path.insert(0, os.getenv("HOME")+"/.local/lib/python3.6/site-packages/")
import torch, torch.nn as nn
import torch.nn.functional as F
from torch.autograd import Variable
use_cuda = torch.cuda.is_available()
print("Torch version:", torch.__version__)
if use_cuda:
print("Using GPU")
else:
print("Not using GPU")
def sample_noise_batch(batch_size):
noise = Variable(torch.randn(batch_size, CODE_SIZE))
return noise.cuda() if use_cuda else noise.cpu()
class Reshape(nn.Module):
def __init__(self, shape):
nn.Module.__init__(self)
self.shape=shape
def forward(self,input):
return input.view(self.shape)
CODE_SIZE = 256
from itertools import count
ix = ('layer_%i'%i for i in count())
generator = nn.Sequential()
generator.add_module(next(ix), nn.Linear(CODE_SIZE, 10*8*8))
generator.add_module(next(ix), nn.ELU())
generator.add_module(next(ix), Reshape([-1, 10, 8, 8]))
generator.add_module(next(ix), nn.ConvTranspose2d(10, 64, kernel_size=(5,5)))
generator.add_module(next(ix), nn.ELU())
generator.add_module(next(ix), nn.ConvTranspose2d(64, 64, kernel_size=(5,5)))
generator.add_module(next(ix), nn.ELU())
generator.add_module(next(ix), nn.Upsample(scale_factor=2))
generator.add_module(next(ix), nn.ConvTranspose2d(64, 32, kernel_size=(5,5)))
generator.add_module(next(ix), nn.ELU())
generator.add_module(next(ix), nn.ConvTranspose2d(32, 32, kernel_size=(5,5)))
generator.add_module(next(ix), nn.ELU())
generator.add_module(next(ix), nn.Conv2d(32, 3, kernel_size=(5,5)))
if use_cuda: generator.cuda()
generated_data = generator(sample_noise_batch(5))
assert tuple(generated_data.shape)[1:] == IMG_SHAPE, "generator must output an image of shape %s, but instead it produces %s"%(IMG_SHAPE,generated_data.shape)
```
### Discriminator
* Discriminator is your usual convolutional network with interlooping convolution and pooling layers
* The network does not include dropout/batchnorm to avoid learning complications.
* We also regularize the pre-output layer to prevent discriminator from being too certain.
```
def sample_data_batch(batch_size):
idxs = np.random.choice(np.arange(data.shape[0]), size=batch_size)
batch = Variable(torch.FloatTensor(data[idxs]))
return batch.cuda() if use_cuda else batch.cpu()
# a special module that converts [batch, channel, w, h] to [batch, units]
class Flatten(nn.Module):
def forward(self, input):
return input.view(input.size(0), -1)
discriminator = nn.Sequential()
discriminator.add_module(next(ix), nn.Conv2d(3,32,kernel_size=3))
discriminator.add_module(next(ix), nn.ReLU())
discriminator.add_module(next(ix), nn.Conv2d(32,32,kernel_size=3))
discriminator.add_module(next(ix), nn.ReLU())
discriminator.add_module(next(ix), nn.MaxPool2d(kernel_size=2))
discriminator.add_module(next(ix), nn.Conv2d(32,64,kernel_size=3))
discriminator.add_module(next(ix), nn.ReLU())
discriminator.add_module(next(ix), nn.Conv2d(64,64,kernel_size=3))
discriminator.add_module(next(ix), nn.ReLU())
discriminator.add_module(next(ix), nn.MaxPool2d(kernel_size=2))
discriminator.add_module(next(ix), Flatten())
discriminator.add_module(next(ix), nn.Linear(2304, 256))
discriminator.add_module(next(ix), nn.Tanh())
discriminator.add_module(next(ix), nn.Linear(256, 1))
if use_cuda: discriminator.cuda()
discriminator(sample_data_batch(5))
```
# Training
We train the two networks concurrently:
* Train __discriminator__ to better distinguish real data from __current__ generator
* Train __generator__ to make discriminator think generator is real
* Since discriminator is a differentiable neural network, we train both with gradient descent.

Training is done iteratively until discriminator is no longer able to find the difference (or until you run out of patience).
### Tricks:
* Regularize discriminator output weights to prevent explosion
* Train generator with __adam__ to speed up training. Discriminator trains with SGD to avoid problems with momentum.
* More: https://github.com/soumith/ganhacks
```
def generator_loss(noise):
"""
1. generate data given noise
2. compute log P(real | gen noise)
3. return generator loss (should be scalar)
"""
generated_data = <generate data given noise>
disc_on_generated_data = <discriminator's opinion on generated data>
logp_gen_is_real = F.logsigmoid(disc_on_generated_data)
loss = - <generator loss>
return loss
loss = generator_loss(sample_noise_batch(32))
print(loss)
assert len(loss.shape) == 1 and loss.shape[0] == 1, "loss must be scalar"
def discriminator_loss(real_data, generated_data):
"""
1. compute discriminator's output on real & generated data
2. compute log-probabilities of real data being real, generated data being fake
3. return discriminator loss (scalar)
"""
disc_on_real_data = <discriminator's opinion on real data>
disc_on_fake_data = <discriminator's opinion on generated data>
logp_real_is_real = F.logsigmoid(disc_on_real_data)
logp_gen_is_fake = F.logsigmoid(- logp_gen_is_fake)
loss = <discriminator loss>
return loss
loss = discriminator_loss(sample_data_batch(32),
generator(sample_noise_batch(32)))
print(loss)
assert len(loss.shape) == 1 and loss.shape[0] == 1, "loss must be scalar"
```
### Auxilary functions
Here we define a few helper functions that draw current data distributions and sample training batches.
```
def sample_images(nrow,ncol, sharp=False):
images = generator(sample_noise_batch(batch_size=nrow*ncol))
images = images.data.cpu().numpy().transpose([0,2,3,1])
if np.var(images)!=0:
images = images.clip(np.min(data),np.max(data))
for i in range(nrow*ncol):
plt.subplot(nrow,ncol,i+1)
if sharp:
plt.imshow(images[i],cmap="gray", interpolation="none")
else:
plt.imshow(images[i],cmap="gray")
plt.show()
def sample_probas(batch_size):
plt.title('Generated vs real data')
D_real = F.sigmoid(discriminator(sample_data_batch(batch_size)))
generated_data_batch = generator(sample_noise_batch(batch_size))
D_fake = F.sigmoid(discriminator(generated_data_batch))
plt.hist(D_real.data.cpu().numpy(),
label='D(x)', alpha=0.5,range=[0,1])
plt.hist(D_fake.data.cpu().numpy(),
label='D(G(z))',alpha=0.5,range=[0,1])
plt.legend(loc='best')
plt.show()
```
### Training
Main loop.
We just train generator and discriminator in a loop and draw results once every N iterations.
```
#optimizers
disc_opt = torch.optim.SGD(discriminator.parameters(), lr=5e-3)
gen_opt = torch.optim.Adam(generator.parameters(), lr=1e-4)
from IPython import display
from tqdm import tnrange
batch_size = 100
for epoch in tnrange(50000):
# Train discriminator
for i in range(5):
real_data = sample_data_batch(batch_size)
fake_data = generator(sample_noise_batch(batch_size))
loss = discriminator_loss(real_data, fake_data)
disc_opt.zero_grad()
loss.backward()
disc_opt.step()
# Train generator
noise = sample_noise_batch(batch_size)
loss = generator_loss(noise)
gen_opt.zero_grad()
loss.backward()
gen_opt.step()
if epoch %100==0:
display.clear_output(wait=True)
sample_images(2,3,True)
sample_probas(1000)
#The network was trained for about 15k iterations.
#Training for longer yields MUCH better results
plt.figure(figsize=[16,24])
sample_images(16,8)
```
| github_jupyter |
# Code Search on Kubeflow
This notebook implements an end-to-end Semantic Code Search on top of [Kubeflow](https://www.kubeflow.org/) - given an input query string, get a list of code snippets semantically similar to the query string.
**NOTE**: If you haven't already, see [kubeflow/examples/code_search](https://github.com/kubeflow/examples/tree/master/code_search) for instructions on how to get this notebook,.
## Install dependencies
Let us install all the Python dependencies. Note that everything must be done with `Python 2`. This will take a while the first time.
### Verify Version Information
```
%%bash
echo "Pip Version Info: " && python2 --version && python2 -m pip --version && echo
echo "Google Cloud SDK Info: " && gcloud --version && echo
echo "Kubectl Version Info: " && kubectl version
```
### Install Pip Packages
```
! python2 -m pip install -U pip
# Code Search dependencies
! python2 -m pip install --user https://github.com/kubeflow/batch-predict/tarball/master
! python2 -m pip install --user -r src/requirements.ui.txt
! python2 -m pip install --user -r src/requirements.nmslib.txt
! python2 -m pip install --user -r src/requirements.dataflow.txt
# BigQuery Cell Dependencies
! python2 -m pip install --user pandas-gbq
# NOTE: The RuntimeWarnings (if any) are harmless. See ContinuumIO/anaconda-issues#6678.
from pandas.io import gbq
```
## Setup the Evironment
This involves setting up the Ksonnet application, utility environment variables for various CLI steps, GCS bucket, and BigQuery dataset.
### Setup Authorization
In a Kubeflow cluster on GKE, we already have the Google Application Credentials mounted onto each Pod. We can simply point `gcloud` to activate that service account.
```
%%bash
# Activate Service Account provided by Kubeflow.
gcloud auth activate-service-account --key-file=${GOOGLE_APPLICATION_CREDENTIALS}
```
Additionally, to interact with the underlying cluster, we configure `kubectl`.
```
%%bash
kubectl config set-cluster kubeflow --server=https://kubernetes.default --certificate-authority=/var/run/secrets/kubernetes.io/serviceaccount/ca.crt
kubectl config set-credentials jupyter --token "$(cat /var/run/secrets/kubernetes.io/serviceaccount/token)"
kubectl config set-context kubeflow --cluster kubeflow --user jupyter
kubectl config use-context kubeflow
```
Collectively, these allow us to interact with Google Cloud Services as well as the Kubernetes Cluster directly to submit `TFJob`s and execute `Dataflow` pipelines.
### Create PROJECT variable
Set PROJECT to the GCP project you want to use.
* If gcloud has a project set, this will be used by default.
* To use a different project or if gcloud doesn't have a project set, you will need to configure one explicitly.
```
import subprocess
PROJECT = subprocess.check_output(["gcloud", "config", "get-value", "project"]).strip()
# DO NOT MODIFY. These are environment variables to be used in a bash shell.
%env PROJECT $PROJECT
```
### Create a storage bucket
Create a GCS bucket to store data.
```
%%bash
gsutil mb -p $PROJECT gs://code-search
```
## Define an experiment
* This solution consists of multiple jobs and servers that need to share parameters.
* To facilitate this we use [params.json](https://github.com/kubeflow/examples/blob/master/code_search/kubeflow/components/params.jason) to define sets of parameters.
* You configure an experiment to run by defining a set of experiments in the **current** field.
* You can save an old experiment's parameters in a different field (not **current**); it does not matter which field's name to use because we are not running the old experiment.
To get started define your experiment
* Create or modify the **cureent** entry containing a set of values to be used for your experiment.
* Set the following values
* **project**: Set this to the GCP project you are working on.
* **experiment**: Experiment's name.
* **bq_target_dataset**: BigQuery dataset to store data.
* **data_dir**: The data directory in GCS to be used by T2T.
* **working_dir**: The working directory in GCS to store temporary data, stages, and output models.
* **model_dir**:
* After training your model, set this to a GCS directory containing the export model
* e.g gs://code-search/20190402/working/output/export/1533685294
* **embedding_dir**: The embedding directory in GCS to store functions' embeddings.
* **t2t_problem**: Set this to "kf_github_function_docstring".
* **t2t_model**: Set this "kf_similarity_transformer".
* **train_steps**: Numer oftraining steps.
* **eval_steps**: Number of steps to be used for eval.
* **hparams_set**: The set of hyperparameters to use; see some suggestions [here](https://github.com/tensorflow/tensor2tensor#language-modeling).
* **lookup_file**: Set this to the GCS location of the CSV produced by the job to create the nmslib index of the embeddings for all GitHub data.
* **index_file**: Set this to the GCS location of the nmslib index for all the data in GitHub.
### Set up environment variables
Set up environments variables according to params.json.
```
import json
import os
params_file = 'params.json'
with open(params_file) as json_file:
data = json.load(json_file)
params = data['current']
WORKING_DIR = params['working_dir']
DATA_DIR = params['data_dir']
EMBEDDING_DIR = params['embedding_dir']
TARGET_DATASET = params['bq_target_dataset']
# DO NOT MODIFY. This are environment variables to be used in a bash shell.
%env WORKING_DIR $WORKING_DIR
%env DATA_DIR $DATA_DIR
%env TARGET_DATASET $TARGET_DATASET
%env EMBEDDING_DIR $EMBEDDING_DIR
```
### Create the BigQuery dataset
```
%%bash
bq mk ${PROJECT}:${TARGET_DATASET}
```
## View Github Files
This is the query that is run as the first step of the Pre-Processing pipeline and is sent through a set of transformations. This is illustrative of the rows being processed in the pipeline we trigger next.
**WARNING**: The table is large and the query can take a few minutes to complete.
```
query = """
SELECT
MAX(CONCAT(f.repo_name, ' ', f.path)) AS repo_path,
c.content, l.license
FROM
`bigquery-public-data.github_repos.files` AS f
JOIN
`bigquery-public-data.github_repos.contents` AS c ON f.id = c.id
JOIN
`bigquery-public-data.github_repos.licenses` AS l ON l.repo_name = f.repo_name
JOIN (
--this part of the query makes sure repo is watched at least twice since 2017
SELECT
repo
FROM (
SELECT
repo.name AS repo
FROM
`githubarchive.year.2017`
WHERE
type="WatchEvent"
UNION ALL
SELECT
repo.name AS repo
FROM
`githubarchive.month.2018*`
WHERE
type="WatchEvent" )
GROUP BY
1
HAVING
COUNT(*) >= 2 ) AS r
ON
f.repo_name = r.repo
WHERE
f.path LIKE '%.py' AND --with python extension
c.size < 15000 AND --get rid of ridiculously long files
(l.license LIKE 'artistic-%' OR l.license = 'isc' OR l.license = 'mit' --Notice
OR l.license LIKE 'apache-%' OR l.license LIKE 'bsd-%' --Notice
OR l.license LIKE 'cc0-%' OR l.license = 'unlicense' --Unencumbered
OR l.license LIKE 'epl-%' OR l.license LIKE 'mpl-%' --Reciprocal
OR l.license LIKE 'gpl-%' OR l.license LIKE 'lgpl-%' --Restricted
) AND
REGEXP_CONTAINS(c.content, r'def ') --contains function definition
GROUP BY
c.content, l.license
LIMIT
20
"""
gbq.read_gbq(query, dialect='standard', project_id=PROJECT)
```
## Pre-Processing Github Files
In this step, we use [Google Cloud Dataflow](https://cloud.google.com/dataflow/) to preprocess the data.
* We use a K8s Job to run a python program `code_search.dataflow.cli.preprocess_github_dataset` that submits the Dataflow job
* Once the job has been created it can be monitored using the Dataflow console
### Submit the Dataflow Job
Configure the job description. This script will produce jobs/submit-preprocess-job.yaml file.
```
%%bash
python configure_job.py kubeflow/components/submit-preprocess-job.yaml
```
Create and run the job.
```
%%bash
kubectl -n kubeflow apply -f jobs/submit-preprocess-job.yaml
```
When completed successfully, this should create a dataset in `BigQuery` named `$TARGET_DATASET`. Additionally, it also dumps CSV files into `$DATA_DIR` which contain training samples (pairs of function and docstrings) for our Tensorflow Model. A representative set of results can be viewed using the following query.
```
query = """
SELECT *
FROM
{}.token_pairs
LIMIT
10
""".format(TARGET_DATASET)
gbq.read_gbq(query, dialect='standard', project_id=PROJECT)
```
This pipeline also writes a set of CSV files which contain function and docstring pairs delimited by a comma. Here, we list a subset of them.
```
%%bash
LIMIT=10
gsutil ls ${DATA_DIR}/*.csv | head -n ${LIMIT}
```
## Prepare Dataset for Training
We will use `t2t-datagen` to convert the transformed data above into the `TFRecord` format.
```
%%bash
python configure_job.py kubeflow/components/t2t-code-search-datagen.yaml
%%bash
kubectl -n kubeflow apply -f jobs/t2t-code-search-datagen.yaml
```
Once this job finishes, the data directory should have a vocabulary file and a list of `TFRecords` prefixed by the problem name which in our case is `github_function_docstring_extended`. Here, we list a subset of them.
```
%%bash
LIMIT=10
gsutil ls ${DATA_DIR}/vocab*
gsutil ls ${DATA_DIR}/*train* | head -n ${LIMIT}
```
## Execute Tensorflow Training
Once, the `TFRecords` are generated, we will use `t2t-trainer` to execute the training.
```
%%bash
python configure_job.py kubeflow/components/t2t-code-search-trainer.yaml
%%bash
kubectl -n kubeflow apply -f jobs/t2t-code-search-trainer.yaml
```
This will generate TensorFlow model checkpoints which is illustrated below.
```
%%bash
gsutil ls ${WORKING_DIR}/output/*ckpt*
```
## Export Tensorflow Model
We now use `t2t-exporter` to export the `TFModel`.
```
%%bash
python configure_job.py kubeflow/components/t2t-code-search-exporter.yaml
%%bash
kubectl -n kubeflow apply -f jobs/t2t-code-search-exporter.yaml
```
Once completed, this will generate a TensorFlow `SavedModel` which we will further use for both online (via `TF Serving`) and offline inference (via `Kubeflow Batch Prediction`). TFServing expects this directory to consist of numeric subdirectories corresponding to different versions of the model. Each subdirectory contains the saved model in protocol buffer along with the weights.
```
%%bash
gsutil ls ${WORKING_DIR}/output/export
```
## Compute Function Embeddings
In this step, we will use the exported model above to compute function embeddings via another `Dataflow` pipeline. A `Python 2` module `code_search.dataflow.cli.create_function_embeddings` has been provided for this purpose. A list of all possible arguments can be seen below.
### Configuration
First, select a Exported Model version from the `${WORKING_DIR}/output/export/Servo` as seen above. This should be name of a folder with UNIX Seconds Timestamp like `1533685294`. Below, we automatically do that by selecting the folder which represents the latest timestamp.
```
%%bash --out EXPORT_DIR_LS
gsutil ls ${WORKING_DIR}/output/export | grep -oE "([0-9]+)/$"
# WARNING: This routine will fail if no export has been completed successfully.
MODEL_VERSION = max([int(ts[:-1]) for ts in EXPORT_DIR_LS.split('\n') if ts])
# DO NOT MODIFY. These are environment variables to be used in a bash shell.
%env MODEL_VERSION $MODEL_VERSION
```
Modify params.json and set **model_dir** to the directory computed above
### Run the Dataflow Job for Function Embeddings
```
%%bash
python configure_job.py kubeflow/components/submit-code-embeddings-job.yaml
%%bash
kubectl -n kubeflow apply -f jobs/submit-code-embeddings-job.yaml
```
When completed successfully, this should create another table in the same `BigQuery` dataset which contains the function embeddings for each existing data sample available from the previous Dataflow Job. Additionally, it also dumps a CSV file containing metadata for each of the function and its embeddings. A representative query result is shown below.
```
query = """
SELECT *
FROM
{}.function_embeddings
LIMIT
10
""".format(TARGET_DATASET)
gbq.read_gbq(query, dialect='standard', project_id=PROJECT)
```
The pipeline also generates a set of CSV files which will be useful to generate the search index.
```
%%bash
LIMIT=10
gsutil ls ${EMBEDDING_DIR}/*index*.csv | head -n ${LIMIT}
```
## Create Search Index
We now create the Search Index from the computed embeddings. This facilitates k-Nearest Neighbor search to for semantically similar results.
```
%%bash
python configure_job.py kubeflow/components/search-index-creator.yaml
%%bash
kubectl -n kubeflow apply -f jobs/search-index-creator.yaml
```
Using the CSV files generated from the previous step, this creates an index using [NMSLib](https://github.com/nmslib/nmslib). A unified CSV file containing all the code examples for a human-readable reverse lookup during the query, is also created.
```
%%bash
gsutil ls ${WORKING_DIR}/code_search_index*
```
# Deploy the Web App
The included web app provides a simple way for users to submit queries.
* The first web app includes two pieces.
* A Flask app that serves a simple UI for sending queries and uses nmslib to provide fast lookups
* A TF Serving instance to compute the embeddings for search queries
* The second web app is used so that we can optionally use [ArgoCD](https://github.com/argoproj/argo-cd) to keep the serving components up to date.
## Deploy an Inference Server
We've seen offline inference during the computation of embeddings. For online inference, we deploy the exported Tensorflow model above using [Tensorflow Serving](https://www.tensorflow.org/serving/).
```
%%bash
python configure_job.py web-app/query-embed-server.yaml
%%bash
kubectl -n kubeflow apply -f jobs/query-embed-server.yaml
```
## Deploy Search UI
We finally deploy the Search UI which allows the user to input arbitrary strings and see a list of results corresponding to semantically similar Python functions. This internally uses the inference server we just deployed.
```
%%bash
python configure_job.py web-app/search-index-server.yaml
%%bash
kubectl -n kubeflow apply -f jobs/search-index-server.yaml
```
The service should now be available at FQDN of the Kubeflow cluster at path `/code-search/`.
### Connecting via port forwarding
To connect to the web app via port-forwarding
```
POD_NAME=$(kubectl -n kubeflow get pods --selector=app=search-index-server --template '{{range .items}}{{.metadata.name}}{{"\n"}}{{end}}')
kubectl port-forward ${POD_NAME} 8080:8008
```
You should now be able to open up the web app at http://localhost:8080.
| github_jupyter |
# 1. Clone ssd_keras repository.
You should only need the custom layers (AnchorBoxes) and the custom loss function (ssd_loss.compute_loss) to load the model
Also to decode the model output (postprocessing+NMS) you need *decode_detections*
```
#Clone repository
!git clone https://github.com/henritomas/ssd-keras.git
#Shift working directory to ssd-keras folder
%cd /content/ssd-keras
#Import necessary files/libraries
from tensorflow.keras import backend as K
from tensorflow.keras.models import load_model
from tensorflow.keras.optimizers import Adam
from math import ceil
import numpy as np
from matplotlib import pyplot as plt
from models.tensorflow_keras_ssd7 import build_model
from keras_loss_function.tensorflow_keras_ssd_loss import SSDLoss
from keras_layers.tensorflow_keras_layer_AnchorBoxes import AnchorBoxes
from ssd_encoder_decoder.ssd_output_decoder import decode_detections
from imageio import imread
from tensorflow.keras.preprocessing import image
```
# 2. Make sure to upload the tfkeras ssd7 *h5* model
```
model_path = '/content/insert_h5_model_here.h5'
```
```
# TODO: Set the path to the `.h5` file of the model to be loaded.
model_path = '/content/tfkeras_ssd7_vocp.h5'
# We need to create an SSDLoss object in order to pass that to the model loader.
ssd_loss = SSDLoss(neg_pos_ratio=3, alpha=1.0)
K.clear_session() # Clear previous models from memory.
model = load_model(model_path, custom_objects={'AnchorBoxes': AnchorBoxes,
'compute_loss': ssd_loss.compute_loss})
```
# 3. Upload and process image to be predicted on
```
orig_images = [] # Store the images here.
input_images = [] # Store resized versions of the images here.
img_height=300
img_width=480
# We'll only load one image in this example.
img_path = '/content/silicon_valley.jpg'
orig_images.append(imread(img_path))
img = image.load_img(img_path, target_size=(img_height, img_width))
img = image.img_to_array(img)
input_images.append(img)
input_images = np.array(input_images)
```
# 4. Make the prediction, decode it with `decode_detections`
```
y_pred = model.predict(input_images)
y_pred_decoded = decode_detections(y_pred,
confidence_thresh=0.3,
iou_threshold=0.4,
top_k=200,
normalize_coords=True,
img_height=img_height,
img_width=img_width)
np.set_printoptions(precision=2, suppress=True, linewidth=90)
print("Predicted boxes:\n")
print(' class conf xmin ymin xmax ymax')
print(y_pred_decoded)
```
# 5. Visualize our output on the original image.
```
# Display the image and draw the predicted boxes onto it.
# Set the colors for the bounding boxes
colors = plt.cm.hsv(np.linspace(0, 1, 21)).tolist()
classes = ['background', 'person',]
plt.figure(figsize=(20,12))
plt.imshow(orig_images[0])
current_axis = plt.gca()
for box in y_pred_decoded[0]:
# Transform the predicted bounding boxes for the 300x300 image to the original image dimensions.
xmin = box[2] * orig_images[0].shape[1] / img_width
ymin = box[3] * orig_images[0].shape[0] / img_height
xmax = box[4] * orig_images[0].shape[1] / img_width
ymax = box[5] * orig_images[0].shape[0] / img_height
color = colors[int(box[0])]
label = '{}: {:.2f}'.format(classes[int(box[0])], box[1])
current_axis.add_patch(plt.Rectangle((xmin, ymin), xmax-xmin, ymax-ymin, color=color, fill=False, linewidth=2))
current_axis.text(xmin, ymin, label, size='x-large', color='white', bbox={'facecolor':color, 'alpha':1.0})
```
| github_jupyter |
# Lab 6. PyTorch and Recurrent Neural Network
```
import torch
import torch.nn as nn
import torch.nn.functional as F
import os
import numpy as np
import torchvision
import torchvision.transforms as transforms
import torch.optim as optim
from tqdm.notebook import trange, tqdm
from livelossplot import PlotLosses
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
BATCH_SIZE = 64
transform = transforms.Compose([transforms.ToTensor()])
trainset = torchvision.datasets.MNIST(
root='./data',
train=True,
download=True,
transform=transform)
trainloader = torch.utils.data.DataLoader(
trainset,
batch_size=BATCH_SIZE,
shuffle=True,
num_workers=2)
testset = torchvision.datasets.MNIST(
root='./data',
train=False,
download=True,
transform=transform)
testloader = torch.utils.data.DataLoader(
testset,
batch_size=BATCH_SIZE,
shuffle=False,
num_workers=2)
N_STEPS = 28
N_INPUTS = 28
N_NEURONS = 150
N_OUTPUTS = 10
N_EPHOCS = 20
class ImageRNN(nn.Module):
def __init__(self, batch_size, n_steps, n_inputs, n_neurons, n_outputs):
super(ImageRNN, self).__init__()
self.n_neurons = n_neurons
self.batch_size = batch_size
self.n_steps = n_steps
self.n_inputs = n_inputs
self.n_outputs = n_outputs
self.basic_rnn = nn.RNN(self.n_inputs, self.n_neurons)
self.FC = nn.Linear(self.n_neurons, self.n_outputs)
def init_hidden(self,):
return (torch.zeros(1, self.batch_size, self.n_neurons).to(device))
def forward(self, X):
X = X.permute(1, 0, 2)
self.batch_size = X.size(1)
self.hidden = self.init_hidden()
lstm_out, self.hidden = self.basic_rnn(X, self.hidden)
out = self.FC(self.hidden)
return out.view(-1, self.n_outputs)
model = ImageRNN(BATCH_SIZE, N_STEPS, N_INPUTS, N_NEURONS, N_OUTPUTS).to(device)
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(model.parameters(), lr=0.001)
def get_accuracy(logit, target, batch_size):
corrects = (torch.max(logit, 1)[1].view(target.size()).data == target.data).sum()
accuracy = 100.0 * corrects/batch_size
return accuracy.item()
%%time
outPbar = tqdm(range(N_EPHOCS))
liveloss = PlotLosses()
for epoch in range(N_EPHOCS):
logs = {}
a = 0
for mode in ['train', 'eval']:
if mode == 'train':
model.train()
loader = trainloader
else:
model.eval()
loader = testloader
running_loss = 0.0
acc = 0.0
for i, data in enumerate(loader):
model.hidden = model.init_hidden()
inputs, labels = data
inputs = inputs.to(device)
labels = labels.to(device)
inputs = inputs.view(-1, 28,28)
outputs = model(inputs)
loss = criterion(outputs, labels)
if mode == 'train':
optimizer.zero_grad()
loss.backward()
optimizer.step()
running_loss += loss.detach().item()
acc += get_accuracy(outputs, labels, BATCH_SIZE)
prefix = 'val_' if mode == 'eval' else ''
logs[prefix + 'Loss'] = running_loss / i
logs[prefix + 'Accuracy'] = acc / i
outPbar.set_postfix_str(f" Loss: {running_loss / i:.4f} | Accuracy: {acc / i:.2f}")
a += 1
outPbar.update(a)
liveloss.update(logs)
liveloss.send()
outPbar.close()
```
| github_jupyter |
# Convolutional Neural Networks (CNN)
<font color='blue'>
<br>Content:
* [Loading the Data Set](#1)
* [Normalization, Reshape and Label Encoding ](#2)
* [Train Test Split](#3)
* [Convolutional Neural Network](#4)
* [What is Convolution Operation?](#5)
* [Same Padding](#6)
* [Max Pooling](#7)
* [Flattening](#8)
* [Full Connection](#9)
* [Implementing with Keras](#10)
* [Create Model](#11)
* [Define Optimizer](#12)
* [Compile Model](#13)
* [Epochs and Batch Size](#14)
* [Data Augmentation](#15)
* [Fit the Model](#16)
* [Evaluate the Model](#17)
* [Deep Learning Tutorial for Beginners](https://www.kaggle.com/kanncaa1/deep-learning-tutorial-for-beginners)
* [Artificial Neural Network with Pytorch](https://www.kaggle.com/kanncaa1/pytorch-tutorial-for-deep-learning-lovers)
* [Convolutional Neural Network with Pytorch](https://www.kaggle.com/kanncaa1/pytorch-tutorial-for-deep-learning-lovers)
* [Recurrent Neural Network with Pytorch](https://www.kaggle.com/kanncaa1/recurrent-neural-network-with-pytorch)
* [Conclusion](#18)
```
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load in
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import seaborn as sns
import matplotlib.pyplot as plt
# import warnings
import warnings
# filter warnings
warnings.filterwarnings('ignore')
# Input data files are available in the "../input/" directory.
# For example, running this (by clicking run or pressing Shift+Enter) will list the files in the input directory
import os
print(os.listdir("../input"))
# Any results you write to the current directory are saved as output.
```
<a id="1"></a>
## Loading the Data Set
* In this part we load and visualize the data.
```
# read train
train = pd.read_csv("../input/train.csv")
print(train.shape)
train.head()
# read test
test= pd.read_csv("../input/test.csv")
print(test.shape)
test.head()
# put labels into y_train variable
Y_train = train["label"]
# Drop 'label' column
X_train = train.drop(labels = ["label"],axis = 1)
# visualize number of digits classes
plt.figure(figsize=(15,7))
g = sns.countplot(Y_train, palette="icefire")
plt.title("Number of digit classes")
Y_train.value_counts()
# plot some samples
img = X_train.iloc[0].as_matrix()
img = img.reshape((28,28))
plt.imshow(img,cmap='gray')
plt.title(train.iloc[0,0])
plt.axis("off")
plt.show()
# plot some samples
img = X_train.iloc[3].as_matrix()
img = img.reshape((28,28))
plt.imshow(img,cmap='gray')
plt.title(train.iloc[3,0])
plt.axis("off")
plt.show()
```
<a id="2"></a>
## Normalization, Reshape and Label Encoding
* Normalization
* We perform a grayscale normalization to reduce the effect of illumination's differences.
* If we perform normalization, CNN works faster.
* Reshape
* Train and test images (28 x 28)
* We reshape all data to 28x28x1 3D matrices.
* Keras needs an extra dimension in the end which correspond to channels. Our images are gray scaled so it use only one channel.
* Label Encoding
* Encode labels to one hot vectors
* 2 => [0,0,1,0,0,0,0,0,0,0]
* 4 => [0,0,0,0,1,0,0,0,0,0]
```
# Normalize the data
X_train = X_train / 255.0
test = test / 255.0
print("x_train shape: ",X_train.shape)
print("test shape: ",test.shape)
# Reshape
X_train = X_train.values.reshape(-1,28,28,1)
test = test.values.reshape(-1,28,28,1)
print("x_train shape: ",X_train.shape)
print("test shape: ",test.shape)
# Label Encoding
from keras.utils.np_utils import to_categorical # convert to one-hot-encoding
Y_train = to_categorical(Y_train, num_classes = 10)
```
<a id="3"></a>
## Train Test Split
* We split the data into train and test sets.
* test size is 10%.
* train size is 90%.
```
# Split the train and the validation set for the fitting
from sklearn.model_selection import train_test_split
X_train, X_val, Y_train, Y_val = train_test_split(X_train, Y_train, test_size = 0.1, random_state=2)
print("x_train shape",X_train.shape)
print("x_test shape",X_val.shape)
print("y_train shape",Y_train.shape)
print("y_test shape",Y_val.shape)
# Some examples
plt.imshow(X_train[2][:,:,0],cmap='gray')
plt.show()
```
<a id="4"></a>
## Convolutional Neural Network
* CNN is used for image classification, object detection
* <a href="https://ibb.co/kV1j9p"><img src="https://preview.ibb.co/nRkBpp/gec2.jpg" alt="gec2" border="0"></a>
<a id="5"></a>
### What is Convolution Operation?
* We have some image and feature detector(3*3)
* Feature detector does not need to be 3 by 3 matrix. It can be 5 by 5 or 7 by 7.
* Feature detector = kernel = filter
* Feauture detector detects features like edges or convex shapes. Example, if out input is dog, feature detector can detect features like ear or tail of the dog.
* feature map = conv(input image, feature detector). Element wise multiplication of matrices.
* feature map = convolved feature
* Stride = navigating in input image.
* We reduce the size of image. This is important bc code runs faster. However, we lost information.
* We create multiple feature maps bc we use multiple feature detectors(filters).
* Lets look at gimp. Edge detect: [0,10,0],[10,-4,10],[0,10,0]
* <a href="https://imgbb.com/"><img src="https://image.ibb.co/m4FQC9/gec.jpg" alt="gec" border="0"></a>
* After having convolution layer we use ReLU to break up linearity. Increase nonlinearity. Because images are non linear.
* <a href="https://ibb.co/mVZih9"><img src="https://preview.ibb.co/gbcQvU/RELU.jpg" alt="RELU" border="0"></a>
<a id="6"></a>
### Same Padding
* As we keep applying conv layers, the size of the volume will decrease faster than we would like. In the early layers of our network, we want to preserve as much information about the original input volume so that we can extract those low level features.
* input size and output size are same.
* <a href="https://ibb.co/jUPkUp"><img src="https://preview.ibb.co/noH5Up/padding.jpg" alt="padding" border="0"></a>
<a id="7"></a>
### Max Pooling
* It makes down-sampling or sub-sampling (Reduces the number of parameters)
* It makes the detection of features invariant to scale or orientation changes.
* It reduce the amount of parameters and computation in the network, and hence to also control overfitting.
* <a href="https://ibb.co/ckTjN9"><img src="https://preview.ibb.co/gsNYFU/maxpool.jpg" alt="maxpool" border="0"></a>
<a id="8"></a>
### Flattening
* <a href="https://imgbb.com/"><img src="https://image.ibb.co/c7eVvU/flattenigng.jpg" alt="flattenigng" border="0"></a>
<a id="9"></a>
### Full Connection
* Neurons in a fully connected layer have connections to all activations in the previous layer
* Artificial Neural Network
* <a href="https://ibb.co/hsS14p"><img src="https://preview.ibb.co/evzsAU/fullyc.jpg" alt="fullyc" border="0"></a>
<a id="10"></a>
## Implementing with Keras
<a id="11"></a>
### Create Model
* conv => max pool => dropout => conv => max pool => dropout => fully connected (2 layer)
* Dropout: Dropout is a technique where randomly selected neurons are ignored during training
* <a href="https://ibb.co/jGcvVU"><img src="https://preview.ibb.co/e7yPPp/dropout.jpg" alt="dropout" border="0"></a>
```
#
from sklearn.metrics import confusion_matrix
import itertools
from keras.utils.np_utils import to_categorical # convert to one-hot-encoding
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten, Conv2D, MaxPool2D
from keras.optimizers import RMSprop,Adam
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import ReduceLROnPlateau
model = Sequential()
#
model.add(Conv2D(filters = 8, kernel_size = (5,5),padding = 'Same',
activation ='relu', input_shape = (28,28,1)))
model.add(MaxPool2D(pool_size=(2,2)))
model.add(Dropout(0.25))
#
model.add(Conv2D(filters = 16, kernel_size = (3,3),padding = 'Same',
activation ='relu'))
model.add(MaxPool2D(pool_size=(2,2), strides=(2,2)))
model.add(Dropout(0.25))
# fully connected
model.add(Flatten())
model.add(Dense(256, activation = "relu"))
model.add(Dropout(0.5))
model.add(Dense(10, activation = "softmax"))
```
<a id="12"></a>
### Define Optimizer
* Adam optimizer: Change the learning rate
```
# Define the optimizer
optimizer = Adam(lr=0.001, beta_1=0.9, beta_2=0.999)
```
<a id="13"></a>
### Compile Model
* categorical crossentropy
* We make binary cross entropy at previous parts and in machine learning tutorial
* At this time we use categorical crossentropy. That means that we have multi class.
* <a href="https://ibb.co/jm1bpp"><img src="https://preview.ibb.co/nN3ZaU/cce.jpg" alt="cce" border="0"></a>
```
# Compile the model
model.compile(optimizer = optimizer , loss = "categorical_crossentropy", metrics=["accuracy"])
```
<a id="14"></a>
### Epochs and Batch Size
* Say you have a dataset of 10 examples (or samples). You have a **batch size** of 2, and you've specified you want the algorithm to run for 3 **epochs**. Therefore, in each epoch, you have 5 **batches** (10/2 = 5). Each batch gets passed through the algorithm, therefore you have 5 iterations **per epoch**.
* reference: https://stackoverflow.com/questions/4752626/epoch-vs-iteration-when-training-neural-networks
```
epochs = 10 # for better result increase the epochs
batch_size = 250
```
<a id="15"></a>
### Data Augmentation
* To avoid overfitting problem, we need to expand artificially our handwritten digit dataset
* Alter the training data with small transformations to reproduce the variations of digit.
* For example, the number is not centered The scale is not the same (some who write with big/small numbers) The image is rotated.
* <a href="https://ibb.co/k24CUp"><img src="https://preview.ibb.co/nMxXUp/augment.jpg" alt="augment" border="0"></a>
```
# data augmentation
datagen = ImageDataGenerator(
featurewise_center=False, # set input mean to 0 over the dataset
samplewise_center=False, # set each sample mean to 0
featurewise_std_normalization=False, # divide inputs by std of the dataset
samplewise_std_normalization=False, # divide each input by its std
zca_whitening=False, # dimesion reduction
rotation_range=0.5, # randomly rotate images in the range 5 degrees
zoom_range = 0.5, # Randomly zoom image 5%
width_shift_range=0.5, # randomly shift images horizontally 5%
height_shift_range=0.5, # randomly shift images vertically 5%
horizontal_flip=False, # randomly flip images
vertical_flip=False) # randomly flip images
datagen.fit(X_train)
```
<a id="16"></a>
### Fit the model
```
# Fit the model
history = model.fit_generator(datagen.flow(X_train,Y_train, batch_size=batch_size),
epochs = epochs, validation_data = (X_val,Y_val), steps_per_epoch=X_train.shape[0] // batch_size)
```
<a id="17"></a>
### Evaluate the model
* Test Loss visualization
* Confusion matrix
```
# Plot the loss and accuracy curves for training and validation
plt.plot(history.history['val_loss'], color='b', label="validation loss")
plt.title("Test Loss")
plt.xlabel("Number of Epochs")
plt.ylabel("Loss")
plt.legend()
plt.show()
# confusion matrix
import seaborn as sns
# Predict the values from the validation dataset
Y_pred = model.predict(X_val)
# Convert predictions classes to one hot vectors
Y_pred_classes = np.argmax(Y_pred,axis = 1)
# Convert validation observations to one hot vectors
Y_true = np.argmax(Y_val,axis = 1)
# compute the confusion matrix
confusion_mtx = confusion_matrix(Y_true, Y_pred_classes)
# plot the confusion matrix
f,ax = plt.subplots(figsize=(8, 8))
sns.heatmap(confusion_mtx, annot=True, linewidths=0.01,cmap="Greens",linecolor="gray", fmt= '.1f',ax=ax)
plt.xlabel("Predicted Label")
plt.ylabel("True Label")
plt.title("Confusion Matrix")
plt.show()
```
<a id="18"></a>
## Conclusion
* http://scs.ryerson.ca/~aharley/vis/conv/flat.html
* HW
* If you have any question I will be very happy to hear it.
| github_jupyter |
```
import os
import cv2
import math
import warnings
import numpy as np
import pandas as pd
import seaborn as sns
import tensorflow as tf
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix, fbeta_score
from keras import optimizers
from keras import backend as K
from keras.models import Sequential, Model
from keras import applications
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import LearningRateScheduler, EarlyStopping, ReduceLROnPlateau
from keras.layers import Dense, Dropout, Flatten, Conv2D, MaxPool2D, Activation, BatchNormalization, GlobalAveragePooling2D, Input
# Set seeds to make the experiment more reproducible.
from tensorflow import set_random_seed
from numpy.random import seed
set_random_seed(0)
seed(0)
%matplotlib inline
sns.set(style="whitegrid")
warnings.filterwarnings("ignore")
train = pd.read_csv('../input/imet-2019-fgvc6/train.csv')
labels = pd.read_csv('../input/imet-2019-fgvc6/labels.csv')
test = pd.read_csv('../input/imet-2019-fgvc6/sample_submission.csv')
train["attribute_ids"] = train["attribute_ids"].apply(lambda x:x.split(" "))
train["id"] = train["id"].apply(lambda x: x + ".png")
test["id"] = test["id"].apply(lambda x: x + ".png")
print('Number of train samples: ', train.shape[0])
print('Number of test samples: ', test.shape[0])
print('Number of labels: ', labels.shape[0])
display(train.head())
display(labels.head())
```
### Model parameters
```
# Model parameters
BATCH_SIZE = 64
EPOCHS = 30
WARMUP_EPOCHS = 2
LEARNING_RATE = 0.001
WARMUP_LEARNING_RATE = 0.001
HEIGHT = 224
WIDTH = 224
CANAL = 3
N_CLASSES = labels.shape[0]
ES_PATIENCE = 3
DECAY_DROP = 0.5
DECAY_EPOCHS = 10
def f2_score_thr(threshold=0.5):
def f2_score(y_true, y_pred):
beta = 2
y_pred = K.cast(K.greater(K.clip(y_pred, 0, 1), threshold), K.floatx())
true_positives = K.sum(K.clip(y_true * y_pred, 0, 1), axis=1)
predicted_positives = K.sum(K.clip(y_pred, 0, 1), axis=1)
possible_positives = K.sum(K.clip(y_true, 0, 1), axis=1)
precision = true_positives / (predicted_positives + K.epsilon())
recall = true_positives / (possible_positives + K.epsilon())
return K.mean(((1+beta**2)*precision*recall) / ((beta**2)*precision+recall+K.epsilon()))
return f2_score
def custom_f2(y_true, y_pred):
beta = 2
tp = np.sum((y_true == 1) & (y_pred == 1))
tn = np.sum((y_true == 0) & (y_pred == 0))
fp = np.sum((y_true == 0) & (y_pred == 1))
fn = np.sum((y_true == 1) & (y_pred == 0))
p = tp / (tp + fp + K.epsilon())
r = tp / (tp + fn + K.epsilon())
f2 = (1+beta**2)*p*r / (p*beta**2 + r + 1e-15)
return f2
def step_decay(epoch):
initial_lrate = LEARNING_RATE
drop = DECAY_DROP
epochs_drop = DECAY_EPOCHS
lrate = initial_lrate * math.pow(drop, math.floor((1+epoch)/epochs_drop))
return lrate
train_datagen=ImageDataGenerator(rescale=1./255, validation_split=0.25)
train_generator=train_datagen.flow_from_dataframe(
dataframe=train,
directory="../input/imet-2019-fgvc6/train",
x_col="id",
y_col="attribute_ids",
batch_size=BATCH_SIZE,
shuffle=True,
class_mode="categorical",
target_size=(HEIGHT, WIDTH),
subset='training')
valid_generator=train_datagen.flow_from_dataframe(
dataframe=train,
directory="../input/imet-2019-fgvc6/train",
x_col="id",
y_col="attribute_ids",
batch_size=BATCH_SIZE,
shuffle=True,
class_mode="categorical",
target_size=(HEIGHT, WIDTH),
subset='validation')
test_datagen = ImageDataGenerator(rescale=1./255)
test_generator = test_datagen.flow_from_dataframe(
dataframe=test,
directory = "../input/imet-2019-fgvc6/test",
x_col="id",
target_size=(HEIGHT, WIDTH),
batch_size=1,
shuffle=False,
class_mode=None)
```
### Model
```
def create_model(input_shape, n_out):
input_tensor = Input(shape=input_shape)
base_model = applications.ResNet50(weights=None, include_top=False,
input_tensor=input_tensor)
base_model.load_weights('../input/resnet50/resnet50_weights_tf_dim_ordering_tf_kernels_notop.h5')
x = GlobalAveragePooling2D()(base_model.output)
x = Dropout(0.5)(x)
x = Dense(1024, activation='relu')(x)
x = Dropout(0.5)(x)
final_output = Dense(n_out, activation='sigmoid', name='final_output')(x)
model = Model(input_tensor, final_output)
return model
model = create_model(input_shape=(HEIGHT, WIDTH, CANAL), n_out=N_CLASSES)
for layer in model.layers:
layer.trainable = False
for i in range(-5,0):
model.layers[i].trainable = True
metrics = ["accuracy"]
optimizer = optimizers.Adam(lr=WARMUP_LEARNING_RATE)
model.compile(optimizer=optimizer, loss="binary_crossentropy", metrics=metrics)
model.summary()
```
#### Train top layers
```
STEP_SIZE_TRAIN = train_generator.n//train_generator.batch_size
STEP_SIZE_VALID = valid_generator.n//valid_generator.batch_size
history = model.fit_generator(generator=train_generator,
steps_per_epoch=STEP_SIZE_TRAIN,
validation_data=valid_generator,
validation_steps=STEP_SIZE_VALID,
epochs=WARMUP_EPOCHS,
verbose=2)
```
#### Fine-tune the complete model
```
for layer in model.layers:
layer.trainable = True
# lrate = LearningRateScheduler(step_decay)
rlrop = ReduceLROnPlateau(monitor='val_loss', factor=0.25, patience=1)
es = EarlyStopping(monitor='val_loss', mode='min', verbose=1, patience=ES_PATIENCE)
callbacks = [es, rlrop]
optimizer = optimizers.Adam(lr=LEARNING_RATE)
metrics = ["accuracy"]
model.compile(optimizer=optimizer, loss="binary_crossentropy", metrics=metrics)
model.summary()
STEP_SIZE_TRAIN = train_generator.n//train_generator.batch_size
STEP_SIZE_VALID = valid_generator.n//valid_generator.batch_size
history = model.fit_generator(generator=train_generator,
steps_per_epoch=STEP_SIZE_TRAIN,
validation_data=valid_generator,
validation_steps=STEP_SIZE_VALID,
epochs=EPOCHS,
callbacks=callbacks,
verbose=2)
```
### Complete model graph loss
```
sns.set_style("whitegrid")
fig, (ax1, ax2) = plt.subplots(1, 2, sharex='col', figsize=(20,7))
ax1.plot(history.history['loss'], label='Train loss')
ax1.plot(history.history['val_loss'], label='Validation loss')
ax1.legend(loc='best')
ax1.set_title('Loss')
ax2.plot(history.history['acc'], label='Train Accuracy')
ax2.plot(history.history['val_acc'], label='Validation accuracy')
ax2.legend(loc='best')
ax2.set_title('Accuracy')
plt.xlabel('Epochs')
sns.despine()
plt.show()
```
### Find best threshold value
```
lastFullValPred = np.empty((0, N_CLASSES))
lastFullValLabels = np.empty((0, N_CLASSES))
for i in range(STEP_SIZE_VALID+1):
im, lbl = next(valid_generator)
scores = model.predict(im, batch_size=valid_generator.batch_size)
lastFullValPred = np.append(lastFullValPred, scores, axis=0)
lastFullValLabels = np.append(lastFullValLabels, lbl, axis=0)
print(lastFullValPred.shape, lastFullValLabels.shape)
def find_best_fixed_threshold(preds, targs, do_plot=True):
score = []
thrs = np.arange(0, 0.5, 0.01)
for thr in thrs:
score.append(custom_f2(targs, (preds > thr).astype(int)))
score = np.array(score)
pm = score.argmax()
best_thr, best_score = thrs[pm], score[pm].item()
print(f'thr={best_thr:.3f}', f'F2={best_score:.3f}')
if do_plot:
plt.plot(thrs, score)
plt.vlines(x=best_thr, ymin=score.min(), ymax=score.max())
plt.text(best_thr+0.03, best_score-0.01, f'$F_{2}=${best_score:.3f}', fontsize=14);
plt.show()
return best_thr, best_score
threshold, best_score = find_best_fixed_threshold(lastFullValPred, lastFullValLabels, do_plot=True)
```
### Apply model to test set and output predictions
```
test_generator.reset()
STEP_SIZE_TEST = test_generator.n//test_generator.batch_size
preds = model.predict_generator(test_generator, steps=STEP_SIZE_TEST)
predictions = []
for pred_ar in preds:
valid = []
for idx, pred in enumerate(pred_ar):
if pred > threshold:
valid.append(idx)
if len(valid) == 0:
valid.append(np.argmax(pred_ar))
predictions.append(valid)
filenames = test_generator.filenames
label_map = {valid_generator.class_indices[k] : k for k in valid_generator.class_indices}
results = pd.DataFrame({'id':filenames, 'attribute_ids':predictions})
results['id'] = results['id'].map(lambda x: str(x)[:-4])
results['attribute_ids'] = results['attribute_ids'].apply(lambda x: list(map(label_map.get, x)))
results["attribute_ids"] = results["attribute_ids"].apply(lambda x: ' '.join(x))
results.to_csv('submission.csv',index=False)
results.head(10)
```
| github_jupyter |
```
%matplotlib inline
import json
import numpy as np
import scipy as sp
import datetime
import matplotlib.pyplot as plt
import operator
import os
import pandas as pd
```
## One single Call
```
import sys
mod_path = '/Users/Simo//Documents/energyanalytics/energyanalytics/disaggregation'
if not (mod_path in sys.path):
sys.path.insert(0, mod_path)
mod_path = '/Users/Simo//Documents/energyanalytics/energyanalytics/disaggregation'
if not (mod_path in sys.path):
sys.path.insert(0, mod_path)
from bayesian_cp_detect import bayesian_cp_3 as bcp
from bayesian_cp_detect import cp_detect
# make sure that the code is loaded to the lastest version
reload(bcp)
import string
import copy
#from datadef import wdayformat
def wdayformat(tm_wday):
if tm_wday==1:
strwday='Mon'
elif tm_wday==2:
strwday='Tue'
elif tm_wday==3:
strwday='Wed'
elif tm_wday==4:
strwday='Thu'
elif tm_wday==5:
strwday='Fri'
elif tm_wday==6:
strwday='Sat'
elif tm_wday==7:
strwday='Sun'
return strwday
def readfile(f,Col): #read .csv files
data=[]
time=[]
#head=''
lines = f.readlines()
#label=[]
#head+=lines[0]
for line in lines[1:]:
line=line.strip('\n')
line=line.split(',')
tmp_time=float(line[0])
tmp_data=0
for i in range(len(Col)):
tmp_data+=float(line[Col[i]])
data.append(tmp_data),
time.append(tmp_time),
f.close()
return (time,data)
filename='/Users/Simo/Desktop/equota/disaggrREDD/house1_output15s'
filext='.dat'
f_input = file(filename+filext,'r')
AppNo=[3,4,5,6,7,8,9,11,12,13,14,15,16,17,18,19,20] #Choose App#
period=1440
N=1
(t_all,y_all)=np.array(readfile(f_input,[i-2 for i in AppNo]))
t=np.array([i+1 for i in range(period)])
y=y_all[N*period:(N+1)*period]
def rel_change(y):
"""
return relative change comparing to the closer neighbouring points
"""
return np.min([np.abs(y[1] - y[0]), np.abs(y[1] - y[2])]) / float(y[1])
def rel_change_filter_0819_3(t, data_input, thre=.2):
"""
filter data based on relative change
data points in data_input that below or above both neighbouring points
and have relative change above thre will be set as the average of neighbouring data.
"""
thre_2 = thre/(1-thre)
id_filter = [i for i in range(1, len(data_input)-1)
if (data_input[i]>data_input[i-1] and data_input[i]>data_input[i+1] and rel_change(data_input[i-1:i+2])>thre) or
(data_input[i]<data_input[i-1] and data_input[i]<data_input[i+1] and rel_change(data_input[i-1:i+2])>thre_2)
]
data_input_2 = [(data_input[i-1]+data_input[i+1])/2 if (i in id_filter) else dat for i, dat in enumerate(data_input) ]
return t, data_input_2
def initial_disaggregate(t_all,y_all,num_day,period = 1440):
#date_current = date_start
day = 0
all_dat_seg = []
while day < num_day:
#print 'reading: ', date_current
#t, y = read_dat_0819(date_current, 0, 23, '../new_data/IHG')
t=np.array([i+1 for i in range(period)])
y=y_all[(day)*period:(day+1)*period]
t_2, y_2 = rel_change_filter_0819_3(t,y)
mu_list_list, sigma_list_list, prob_r_list_list, r_list_list = cp_detect.bayesian_change_point_4(y_2, r_blur=30)
changepoint, changepoint_p = cp_detect.get_change_point(prob_r_list_list)
changepoint.append(len(t_2)-1)
if len(changepoint)>1:
dat_seg = [[y[cp_start:cp_end], y[cp_start-3:cp_start]] for cp_start, cp_end in zip(changepoint[:-1], changepoint[1:])]
else:
dat_seg = []
all_dat_seg.extend(dat_seg)
#date_current+=datetime.timedelta(1)
day+=1
return all_dat_seg
day = 65
t=np.array([i+1 for i in range(period)])
y=y_all[(day)*period:(day+1)*period]
t_2, y_2 = rel_change_filter_0819_3(t,y)
mu_list_list, sigma_list_list, prob_r_list_list, r_list_list = cp_detect.bayesian_change_point_4(y_2, r_blur=30)
changepoint, changepoint_p = cp_detect.get_change_point(prob_r_list_list)
changepoint.append(len(t_2)-1)
def plot_24h_data(t, raw_data,cp_list):
fig, axes = plt.subplots(nrows=4, figsize=[18, 10])
for i, ax in enumerate(axes):
#ax.plot(t, data, 'r-', markersize=3, linewidth=1, label='smooth')
ax.plot(t, raw_data, 'k.', markersize=3, label='raw')
for cp in cp_list:
ax.plot([t[cp], t[cp]], [0, 3000], 'k-', linewidth=1)
ax.set_ylabel('power')
ax.set_xlim([0+i*3600,3600+i*3600])
ax.set_xlabel('time/h')
plt.legend()
plot_24h_data(t_2,y_2,changepoint)
all_seg_april = initial_disaggregate(t_all,y_all,500)
all_seg_april_normalized = [np.array(x[0])-np.mean(x[1]) for x in all_seg_april if len(x[1])==3]
all_seg_april_normalized = [x for x in all_seg_april_normalized if len(x)>0]
all_positive_seg_april_normalized = [x for x in all_seg_april_normalized if x.min()>0]
```
## 1. Set up disaggregation algorithm options
```
import sys
mod_path = '/Users/Simo//Documents/energyanalytics/energyanalytics/disaggregation'
if not (mod_path in sys.path):
sys.path.insert(0, mod_path)
mod_path = '/Users/Simo//Documents/energyanalytics/energyanalytics/disaggregation'
if not (mod_path in sys.path):
sys.path.insert(0, mod_path)
from bayesian_cp_detect import bayesian_cp_3 as bcp
from bayesian_cp_detect import cp_detect
# make sure that the code is loaded to the lastest version
reload(bcp)
```
## Read File
```
import string
import copy
#from datadef import wdayformat
def wdayformat(tm_wday):
if tm_wday==1:
strwday='Mon'
elif tm_wday==2:
strwday='Tue'
elif tm_wday==3:
strwday='Wed'
elif tm_wday==4:
strwday='Thu'
elif tm_wday==5:
strwday='Fri'
elif tm_wday==6:
strwday='Sat'
elif tm_wday==7:
strwday='Sun'
return strwday
def readfile(f,Col): #read .csv files
data=[]
time=[]
#head=''
lines = f.readlines()
#label=[]
#head+=lines[0]
for line in lines[1:]:
line=line.strip('\n')
line=line.split(',')
tmp_time=float(line[0])
tmp_data=0
for i in range(len(Col)):
tmp_data+=float(line[Col[i]])
data.append(tmp_data),
time.append(tmp_time),
f.close()
return (time,data)
filename='/Users/Simo/Desktop/equota/disaggrREDD/house1_output15s'
filext='.dat'
f_input = file(filename+filext,'r')
AppNo=[3,4,5,6,7,8,9,11,12,13,14,15,16,17,18,19,20] #Choose App#
period=1440
N=1
(t_all,y_all)=np.array(readfile(f_input,[i-2 for i in AppNo]))
t=np.array([i+1 for i in range(period)])
y=y_all[N*period:(N+1)*period]
def rel_change(y):
"""
return relative change comparing to the closer neighbouring points
"""
return np.min([np.abs(y[1] - y[0]), np.abs(y[1] - y[2])]) / float(y[1])
def rel_change_filter_0819_3(t, data_input, thre=.2):
"""
filter data based on relative change
data points in data_input that below or above both neighbouring points
and have relative change above thre will be set as the average of neighbouring data.
"""
thre_2 = thre/(1-thre)
id_filter = [i for i in range(1, len(data_input)-1)
if (data_input[i]>data_input[i-1] and data_input[i]>data_input[i+1] and rel_change(data_input[i-1:i+2])>thre) or
(data_input[i]<data_input[i-1] and data_input[i]<data_input[i+1] and rel_change(data_input[i-1:i+2])>thre_2)
]
data_input_2 = [(data_input[i-1]+data_input[i+1])/2 if (i in id_filter) else dat for i, dat in enumerate(data_input) ]
return t, data_input_2
def initial_disaggregate(t_all,y_all,num_day,period = 1440):
#date_current = date_start
day = 0
all_dat_seg = []
while day < num_day:
#print 'reading: ', date_current
#t, y = read_dat_0819(date_current, 0, 23, '../new_data/IHG')
t=np.array([i+1 for i in range(period)])
y=y_all[(day)*period:(day+1)*period]
t_2, y_2 = rel_change_filter_0819_3(t,y)
mu_list_list, sigma_list_list, prob_r_list_list, r_list_list = cp_detect.bayesian_change_point_4(y_2, r_blur=30)
changepoint, changepoint_p = cp_detect.get_change_point(prob_r_list_list)
changepoint.append(len(t_2)-1)
if len(changepoint)>1:
dat_seg = [[y[cp_start:cp_end], y[cp_start-3:cp_start]] for cp_start, cp_end in zip(changepoint[:-1], changepoint[1:])]
else:
dat_seg = []
all_dat_seg.extend(dat_seg)
#date_current+=datetime.timedelta(1)
day+=1
return all_dat_seg
all_seg_april = initial_disaggregate(t_all,y_all,500)
len(t_all),len(y_all)
day = 67
t=np.array([i+1 for i in range(period)])
y=y_all[(day)*period:(day+1)*period]
t_2, y_2 = rel_change_filter_0819_3(t,y)
mu_list_list, sigma_list_list, prob_r_list_list, r_list_list = cp_detect.bayesian_change_point_4(y_2, r_blur=30)
changepoint, changepoint_p = cp_detect.get_change_point(prob_r_list_list)
changepoint.append(len(t_2)-1)
def plot_24h_data(t, raw_data,cp_list):
fig, axes = plt.subplots(nrows=4, figsize=[18, 10])
for i, ax in enumerate(axes):
#ax.plot(t, data, 'r-', markersize=3, linewidth=1, label='smooth')
ax.plot(t, raw_data, 'k.', markersize=3, label='raw')
for cp in cp_list:
ax.plot([t[cp], t[cp]], [0, 3000], 'k-', linewidth=1)
ax.set_ylabel('power')
ax.set_xlim([0+i*360,360+i*360])
ax.set_xlabel('time/h')
plt.legend()
plot_24h_data(t_2,y_2,changepoint)
all_seg_april_normalized = [np.array(x[0])-np.mean(x[1]) for x in all_seg_april if len(x[1])==3]
all_seg_april_normalized = [x for x in all_seg_april_normalized if len(x)>0]
all_positive_seg_april_normalized = [x for x in all_seg_april_normalized if x.min()>0]
#all_seg_april_normalized = [x for x in all_positive_seg_april_normalized if x.mean()<500]
all_seg_april_normalized = [x for x in all_positive_seg_april_normalized if x.mean()>500]
plt.figure(figsize=[8, 6])
for x in all_seg_april_normalized:
plt.plot(x, 'k-', linewidth=.5, alpha=.1)
plt.xlabel('time point after change point')
plt.ylabel('relative power shift')
plt.xlim([0, 10])
plt.ylim([-600, 4000])
```
## Cluster
```
from sklearn.cluster import KMeans
from sklearn.datasets import make_blobs
from sklearn.metrics import silhouette_samples, silhouette_score
import operator
import seaborn as sns
def extract_first_n(data_list, n):
return [t[:n] for t in data_list if len(t)>=n]
# integrated functions:
def integrated_clustering(t_all,y_all,num_of_days=500,period = 1440,trim=10,min_n_clusters = 4, max_n_clusters=10,hierarchical=0):
all_seg_april = initial_disaggregate(t_all,y_all,num_of_days,period = period)
all_seg_april_normalized = [np.array(x[0])-np.mean(x[1]) for x in all_seg_april if len(x[1])==3]
all_seg_april_normalized = [x for x in all_seg_april_normalized if len(x)>0]
if hierarchical == 0:
pass
elif hierarchical ==1:
all_seg_april_normalized = [x for x in all_seg_april_normalized if x.mean()>1000]
else:
all_seg_april_normalized = [x for x in all_seg_april_normalized if x.mean()<1000]
all_positive_seg_april_normalized = [x for x in all_seg_april_normalized if x.min()>0]
all_seg_april_normalized_trim50 = extract_first_n(all_positive_seg_april_normalized, trim)
cluster_average = []
# find optimal clustering number using silhouette score
optimal_dict = {}
for n_clusters in range(min_n_clusters,max_n_clusters):
y_pred = KMeans(n_clusters=n_clusters).fit_predict(all_seg_april_normalized_trim50)
cluster_average = []
for i_cluster in range(n_clusters):
cluster_average.append(
np.mean([np.mean(x) for i, x in enumerate(all_seg_april_normalized_trim50) if y_pred[i]==i_cluster])
)
# sihouette score
cluster_labels = y_pred
sample_silhouette_values = silhouette_samples(all_seg_april_normalized_trim50, cluster_labels)
silhouette_avg = silhouette_score(pd.DataFrame(all_seg_april_normalized_trim50), cluster_labels)
optimal_dict[n_clusters] = silhouette_avg +(sample_silhouette_values.min()+sample_silhouette_values.max())/2
n_clusters = max(optimal_dict.iteritems(), key=operator.itemgetter(1))[0]
print n_clusters
y_pred = KMeans(n_clusters=n_clusters).fit_predict(all_seg_april_normalized_trim50)
cluster_average = []
for i_cluster in range(n_clusters):
cluster_average.append(
np.mean([np.mean(x) for i, x in enumerate(all_seg_april_normalized_trim50) if y_pred[i]==i_cluster])
)
cluster_average_rank = np.argsort(cluster_average)[::-1]
rank_map = {cluster_average_rank[i_cluster]:i_cluster for i_cluster in range(n_clusters)} # old index:new index
y_pred_old = y_pred
y_pred = [rank_map[x] for x in y_pred]
all_seg_per_cluster = [[] for i in range(n_clusters) ]
for i_seg in range(len(all_seg_april_normalized_trim50)):
all_seg_per_cluster[y_pred[i_seg]].append(all_seg_april_normalized_trim50[i_seg])
cluster_mean = [[] for i in range(n_clusters) ]
cluster_std = [[] for i in range(n_clusters) ]
for i_cluster in range(n_clusters):
cluster_mean[ i_cluster ] = np.mean(np.array(all_seg_per_cluster[i_cluster]), axis=0)
cluster_std[ i_cluster ] = np.std(np.array(all_seg_per_cluster[i_cluster]), axis=0)
#cluster_mean_2 = cluster_mean[5:6]
return cluster_mean,cluster_std,n_clusters,all_seg_per_cluster
def plot_cluster_result(cluster_mean,cluster_std,n_clusters,all_seg_per_cluster):
color_list = sns.color_palette("hls", n_clusters)
fig, ax = plt.subplots(nrows=5,ncols=4,figsize=[20,12]);
ax = ax.flatten()
for i_cluster in range(n_clusters):
ax_current = ax[i_cluster]
for seg in all_seg_per_cluster[i_cluster]:
ax_current.plot(seg, '-', linewidth=1, alpha=.3, color=color_list[i_cluster])
ax_current.set_xlim([0, 10])
ax_current.set_ylim([-500, 4000])
ax_current.plot([0,50], [0,0], 'k--')
ax_current.plot(cluster_mean[i_cluster], color=color_list[i_cluster])
ax_current.fill_between(range(10)
, cluster_mean[i_cluster]-cluster_std[i_cluster]
, cluster_mean[i_cluster]+cluster_std[i_cluster]
, color=color_list[i_cluster], alpha=.8)
def n_dimension_identity_matrix(cluster_mean_2):
to_return_list = []
for i in range(1,len(cluster_mean_2)+1):
to_return_list.append([0 if j!=i else 1 for j in range(1,len(cluster_mean_2)+1)])
return to_return_list
# run before dissagregate
cluster_mean_2 = []
cluster_mean_2.append(cluster_mean[5])
cluster_mean_2.append(cluster_mean[8])
cluster_mean_2.append(cluster_mean[14])
cluster_mean_2.append(cluster_mean[16])
cluster_mean_2.append(cluster_mean[12])
cluster_mean_2
cluster_mean,cluster_std,n_clusters,all_seg_per_cluster = integrated_clustering(t_all,y_all,num_of_days=500,period = 1440,trim=10,min_n_clusters = 17, max_n_clusters=18)
plot_cluster_result(cluster_mean,cluster_std,n_clusters,all_seg_per_cluster)
import seaborn as sns
sns.set_style("whitegrid")
sns.set_context("poster")
color_list = sns.color_palette("hls", 8)
from sklearn.cluster import KMeans
from sklearn.datasets import make_blobs
# define a function that trim the first n data points, or discard if len is less
def extract_first_n(data_list, n):
return [t[:n] for t in data_list if len(t)>=n]
all_seg_april_normalized_trim50 = extract_first_n(all_seg_april_normalized, 10)
print '# segs before trimming: ', len(all_seg_april_normalized)
print '# segs after trimming: ', len(all_seg_april_normalized_trim50)
n_clusters=10
y_pred = KMeans(n_clusters=n_clusters).fit_predict(all_seg_april_normalized_trim50)
cluster_average = []
for i_cluster in range(n_clusters):
cluster_average.append(
np.mean([np.mean(x) for i, x in enumerate(all_seg_april_normalized_trim50) if y_pred[i]==i_cluster])
)
cluster_average_rank = np.argsort(cluster_average)[::-1]
rank_map = {cluster_average_rank[i_cluster]:i_cluster for i_cluster in range(n_clusters)} # old index:new index
y_pred_old = y_pred
y_pred = [rank_map[x] for x in y_pred]
from sklearn.datasets import make_blobs
from sklearn.cluster import KMeans
from sklearn.metrics import silhouette_samples, silhouette_score
for n_clusters in range(4,10):
y_pred = KMeans(n_clusters=n_clusters).fit_predict(all_seg_april_normalized_trim50)
cluster_average = []
for i_cluster in range(n_clusters):
cluster_average.append(
np.mean([np.mean(x) for i, x in enumerate(all_seg_april_normalized_trim50) if y_pred[i]==i_cluster])
)
# sihouette score
cluster_labels = y_pred
#sample_silhouette_values = silhouette_samples(all_seg_april_normalized_trim50, cluster_labels)
#print sample_silhouette_values.min()
silhouette_avg = silhouette_score(pd.DataFrame(all_seg_april_normalized_trim50), cluster_labels)
#print silhouette_avg + sample_silhouette_values.min()
cluster_average_rank = np.argsort(cluster_average)[::-1]
rank_map = {cluster_average_rank[i_cluster]:i_cluster for i_cluster in range(n_clusters)} # old index:new index
y_pred_old = y_pred
y_pred = [rank_map[x] for x in y_pred]
all_seg_per_cluster = [[] for i in range(n_clusters) ]
for i_seg in range(len(all_seg_april_normalized_trim50)):
all_seg_per_cluster[y_pred[i_seg]].append(all_seg_april_normalized_trim50[i_seg])
cluster_mean = [[] for i in range(n_clusters) ]
cluster_std = [[] for i in range(n_clusters) ]
for i_cluster in range(n_clusters):
cluster_mean[ i_cluster ] = np.mean(np.array(all_seg_per_cluster[i_cluster]), axis=0)
cluster_std[ i_cluster ] = np.std(np.array(all_seg_per_cluster[i_cluster]), axis=0)
reload(bcp)
#cluster_mean_2 = cluster_mean
predicted_profile = integrated_dissagregate(y_all,1440,cluster_mean_2,day = 66,n_equipment_type = len(cluster_mean_2),n_equipment=[2 for i in range(1,len(cluster_mean_2)+1)],obs_mat=n_dimension_identity_matrix(cluster_mean_2),power_usage = [i.mean() for i in cluster_mean_2])
plot_dissagregation(predicted_profile,np.array([i+1 for i in range(period)]))
# integrated dissagregation
#cluster_mean_2 = []
#cluster_mean_2.append(np.array([80 for i in range(20)]))
#cluster_mean_2.append(np.array([50 for i in range(20)]))
#cluster_mean_2.append(np.array([1550 for i in range(20)]))
#cluster_mean_2.append(np.array([1000 for i in range(20)]))
#cluster_mean_2.append(np.array([200 for i in range(20)]))
def DP_state_generation(N):
if N==1:
return [[0],[1]]
else:
return_list = DP_state_generation(N-1)
toreturn = []
for i in return_list:
#print i
i.append(0)
toreturn.append(copy.copy(i))
i.pop()
i.append(1)
#print i
toreturn.append(copy.copy(i))
#print toreturn
return toreturn
def generate_state_prob_list(N):
toreturn = {}
toreturnlist = DP_state_generation(N)
k = 1/float(len(toreturnlist))
for i in toreturnlist:
toreturn[tuple(i)] = k
return toreturn
def integrated_dissagregate(y_all,period,cluster_mean_2,day = 65,n_equipment_type = 4,n_equipment = [2,2,2,2],obs_mat = np.array([[1,0,0,0],[0,1,0,0],[0,0,1,0],[0,0,0,1]]),power_usage = [0,0,0,0]):
opt = bcp.set_disaggregation_option(change_shape=cluster_mean_2,
init_pos_std = np.sqrt([float(200/3), float(200/3), float(400/3), float(400/3)])
)
t=np.array([i+1 for i in range(period)])
data=y_all[(day)*period:(day+1)*period]
cp_list = bcp.disaggregate(data,opt)
print cp_list
data_seg, n_seg, data_seg_raw_last = bcp.segment_data(data, cp_list)
new_data_seg, new_n_seg, new_data_seg_raw_last = bcp.segment_data_new(data, cp_list)
print new_data_seg
shape_matched = cluster_mean_2
n_shape_matched = len(shape_matched)
all_shape_code = bcp.shape_code_gen(n_shape_matched)
shape_dict = bcp.combine_shape(shape_matched, all_shape_code)
shape_prob_list = bcp.get_seg_prob(data_seg, shape_dict)
new_shape_prob_list = bcp.get_seg_prob_positive(new_data_seg, shape_dict)
print shape_prob_list
#shape_prob_list = bcp.get_seg_prob(data_seg, shape_dict)
state_prob_list = generate_state_prob_list(len(cluster_mean_2))
#obs_mat = np.array([[1,0,0,0,0],[0,1,0,0,0],[0,0,1,0,0],[0,0,0,1,0],[0,0,0,0,1]])
for item,keys in state_prob_list.iteritems():
state_prob_list[item] = new_shape_prob_list[0][item]
#trace_list, shape_list = bcp.viterbi(shape_prob_list, new_shape_prob_list[0], data_seg, obs_mat)
#trace_list, shape_list = viterbi_new(shape_prob_list,state_prob_list,new_shape_prob_list,data_seg,obs_mat,power_usage)
trace_list, shape_list = viterbi_for_missing_change_point(shape_prob_list,state_prob_list,new_shape_prob_list,data_seg,obs_mat,power_usage,alpha = 10)
predicted_profile = bcp.generate_predicted_profile(cp_list, shape_matched, shape_list, data, n_equipment_type, obs_mat, trace_list)
return predicted_profile
def plot_dissagregation(predicted_profile,t):
plt.figure(figsize = [16,8])
for tmp in predicted_profile:
plt.plot(t,tmp,linewidth = 1)
#plt.plot(t,data,'k.',markersize = 2)
plt.xlim([1,1500])
plt.ylim([0,3000])
plt.xlabel('t/h')
plt.ylabel('power')
def plot_dissagregation_2(predicted_profile,t):
plt.figure(figsize = [16,8])
for key,tmp in predicted_profile.iteritems():
plt.plot(t,tmp,linewidth = 1)
#plt.plot(t,data,'k.',markersize = 2)
plt.xlim([1,1500])
plt.ylim([0,3000])
plt.xlabel('t/h')
plt.ylabel('power')
#cluster_mean_2 = cluster_mean[5:6]
cluster_mean_2 = []
#cluster_mean_2.append(np.array([80 for i in range(20)]))
#cluster_mean_2.append(np.array([50 for i in range(20)]))
#cluster_mean_2.append(np.array([1000 for i in range(20)]))
#cluster_mean_2.append(np.array([200 for i in range(20)]))
opt = bcp.set_disaggregation_option(change_shape=cluster_mean_2,
init_pos_std = np.sqrt([float(200/3), float(200/3), float(400/3), float(400/3)])
)
day = 66
t=np.array([i+1 for i in range(period)])
data=y_all[(day)*period:(day+1)*period]
cp_list = bcp.disaggregate(data,opt)
plot_24h_data(t,data,cp_list)
# New Viterbi
data_seg, n_seg, data_seg_raw_last = bcp.segment_data(data, cp_list)
new_data_seg, new_n_seg, new_data_seg_raw_last = bcp.segment_data_new(data, cp_list)
shape_prob_list = bcp.get_seg_prob(data_seg, shape_dict)
new_shape_prob_list = bcp.get_seg_prob_positive(new_data_seg, shape_dict)
import operator
#tuple(map(operator.add, a, b))
error = 0
#init_state = tuple([0,1,0,0,0])
init_state = max(new_shape_prob_list[0].iteritems(),key = operator.itemgetter(1))[0]
power_usage = tuple([100,50,1500,2000,50])
#state = tuple(0,0,0,0,0)
for i in range(len(cp_list)-2):
#print init_state
print max(new_shape_prob_list[i+1].iteritems(),key = operator.itemgetter(1))[0]
init_state = tuple(map(operator.add,init_state,max(shape_prob_list[i+1].iteritems(),key= operator.itemgetter(1))[0]))
print init_state
#init_state = tuple(map(operator.add,init_state,max(shape_prob_list[i+1].iteritems(),key = operator.itemgetter(1))[0]))
#if sum(map(operator.mul,init_state,power_usage))-sum(max(new_shape_prob_list[i+1].iteritems(),key=operator.itemgetter(1))[0])<1000:
#if tuple(map(operator.add,init_state , max(switch_shape_prob_list[i+1].iteritems(),key = operator.itemgetter(1))[0])) == max(shape_prob_list[i+1].iteritems(),key=operator.itemgetter(1))[0]:
# pass
#else:
#print "#####################"
#print i
#print max(shape_prob_list[i].iteritems(),key=operator.itemgetter(1))[0]
#print max(switch_shape_prob_list[i+1].iteritems(),key = operator.itemgetter(1))[0]
#print max(shape_prob_list[i+1].iteritems(),key=operator.itemgetter(1))[0]
#print "#####################"
#error+=1
data_seg, n_seg, data_seg_raw_last = bcp.segment_data(data, cp_list)
shape_matched = cluster_mean_2
n_shape_matched = len(shape_matched)
all_shape_code = bcp.shape_code_gen(n_shape_matched)
shape_dict = bcp.combine_shape(shape_matched, all_shape_code)
print all_shape_code
shape_prob_list = bcp.get_seg_prob(data_seg, shape_dict)
def DP_state_generation(N):
if N==1:
return [[0],[1]]
else:
return_list = DP_state_generation(N-1)
toreturn = []
for i in return_list:
#print i
i.append(0)
toreturn.append(copy.copy(i))
i.pop()
i.append(1)
#print i
toreturn.append(copy.copy(i))
#print toreturn
return toreturn
DP_state_generation(10)
def generate_state_prob_list(N):
toreturn = {}
toreturnlist = DP_state_generation(N)
k = 1/float(len(toreturnlist))
for i in toreturnlist:
toreturn[tuple(i)] = k
return toreturn
generate_state_prob_list(5)
#state_prob_list = {(0,0,0,0,0):.05,(0,1,0,0,0):0.1,(0,0,0,0,1):0.2,(1,0,0,0,0):0.2,(0,0,1,0,0):0.1,(0,0,0,1,0):0.2,(1,1,0,0,0):0.05,(1,0,0,1,0):0.05,(1,0,0,0,1):0.05,(1,0,1,0,0):0.05,(0,0,0,1,1):0.05,(0,1,0,0,1):0.05,(0,1,1,0,0):0.05,(0,1,0,1,0):0.05,(1,1,1,0):0.02,(0,1,1,1):0.02,(1,0,1,1):0.02,(1,1,0,1):0.02,(1,1,1,1):0.2}
state_prob_list = generate_state_prob_list(4)
n_equipment_type = 4
n_equipment = [2,2,2,2]
#obs_mat = np.array([[1,0,0,0,0],[0,1,0,0,0],[0,0,1,0,0],[0,0,0,1,0],[0,0,0,0,1]])
obs_mat = np.array([[1,0,0,0],[0,1,0,0],[0,0,1,0],[0,0,0,1]])
def viterbi_new(shape_prob_list, state_prob_list, boot_state_prob_list, data_seg, obs_mat, power_usage):
# originally shape means the 'change', state means the actual usage...
n_seg = len(data_seg)
state_prob_list_list = [state_prob_list]
state_memory_list_list = []
shape_memory_list_list = []
for i_seg in range(n_seg):
seg_mean = np.mean(data_seg[i_seg])
next_state_prob_list = {t:0 for t in state_prob_list.keys()}
state_memory_list = {t:0 for t in state_prob_list.keys()} #
shape_memory_list = {t:0 for t in state_prob_list.keys()} #
for next_state, next_state_prob in next_state_prob_list.items():
max_prob = -float('Inf')
max_past_state = tuple()
max_shape = ()
#print max(shape_prob_list[i_seg].iteritems(),key = operator.itemgetter(1))[0]
for shape_code, shape_prob in shape_prob_list[i_seg].items():# loop through all possible changes...
#print obs_mat,shape_code
change_state = np.dot(obs_mat, shape_code) # if the obs_mat is identity matrix then the change_state = shape_code
past_state = tuple(np.subtract(next_state, change_state)) # find out the corresponding past_state
if past_state in state_prob_list: # the past state should be all positive
if state_prob_list[past_state] * shape_prob > max_prob:
max_prob = state_prob_list[past_state] * shape_prob
max_past_state = past_state
max_shape = shape_code
state_memory_list[next_state] = max_past_state # the table 2, noting down the most possible past state
next_state_prob_list[next_state] = max_prob
shape_memory_list[next_state] = max_shape
print '##############'
computed = sum(map(operator.mul,max(state_prob_list.iteritems(),key = operator.itemgetter(1))[0],power_usage))
observed = sum(map(operator.mul,max(boot_state_prob_list[i_seg].iteritems(),key = operator.itemgetter(1))[0],power_usage))
print computed
print observed
if (float(computed)-float(observed))/float(observed)<0.3:
state_prob_list = next_state_prob_list
# this code is underdevelopped
else:
print "potential error in change point detected "
next_state = max(boot_state_prob_list[i_seg].iteritems(),key = operator.itemgetter(1))[0] # this is too brutal we impose the next state in this case?
#for shape_code, shape_prob in shape_prob_list[i_seg].items():# loop through all possible changes...
# #print obs_mat,shape_code
# change_state = np.dot(obs_mat, shape_code) # if the obs_mat is identity matrix then the change_state = shape_code
# past_state = tuple(np.subtract(next_state, change_state)) # find out the corresponding past_state
# if past_state in state_prob_list: # the past state should be all positive
# if state_prob_list[past_state] * shape_prob > max_prob:
# max_prob = state_prob_list[past_state] * shape_prob
# max_past_state = past_state
# max_shape = shape_code
# in this case we don't need the information from the shape_prob_list since the missing change point leads to polution of information
max_past_state = max(state_prob_list_list[-1].iteritems(),key = operator.itemgetter(1))[0]
max_shape = tuple(np.subtract(next_state, max_past_state))
state_memory_list[next_state] = max_past_state
shape_memory_list[next_state] = max_shape
state_prob_list = boot_state_prob_list[i_seg]
#print sum(map(operator.mul,max(next_state_prob_list.iteritems(),key = operator.itemgetter(1))[0],power_usage))
print '##############'
state_prob_list_list.append(next_state_prob_list) # debug 05/08/2017 should change to state_prob_list_list.append(state_prob_list)
state_memory_list_list.append(state_memory_list)
shape_memory_list_list.append(shape_memory_list)
trace_list = []
shape_list = []
end_state = sorted(state_prob_list_list[-1].items(), key=operator.itemgetter(1))[-1][0]
trace_list.insert(0, end_state)
for i in reversed(range(n_seg)):
max_shape = shape_memory_list_list[i][end_state]
end_state = state_memory_list_list[i][end_state]
trace_list.insert(0, end_state)
shape_list.insert(0, max_shape)
return trace_list, shape_list
def viterbi_for_missing_change_point(shape_prob_list, state_prob_list, boot_state_prob_list, data_seg, obs_mat, power_usage,alpha = 100):
# originally shape means the 'change', state means the actual usage...
n_seg = len(data_seg)
state_prob_list_list = [state_prob_list]
state_memory_list_list = []
shape_memory_list_list = []
for i_seg in range(n_seg):
seg_mean = np.mean(data_seg[i_seg])
next_state_prob_list = {t:0 for t in state_prob_list.keys()}
state_memory_list = {t:0 for t in state_prob_list.keys()} #
shape_memory_list = {t:0 for t in state_prob_list.keys()} #
for next_state, next_state_prob in next_state_prob_list.items():
max_prob = -float('Inf')
max_past_state = tuple()
max_shape = ()
#print max(shape_prob_list[i_seg].iteritems(),key = operator.itemgetter(1))[0]
for shape_code, shape_prob in shape_prob_list[i_seg].items():# loop through all possible changes...
#print obs_mat,shape_code
change_state = np.dot(obs_mat, shape_code) # if the obs_mat is identity matrix then the change_state = shape_code
past_state = tuple(np.subtract(next_state, change_state)) # find out the corresponding past_state
if past_state in state_prob_list: # the past state should be all positive
if state_prob_list[past_state] * shape_prob + alpha*boot_state_prob_list[i_seg][next_state] > max_prob:
max_prob = state_prob_list[past_state] * shape_prob + alpha*boot_state_prob_list[i_seg][next_state]
max_past_state = past_state
max_shape = shape_code
state_memory_list[next_state] = max_past_state # the table 2, noting down the most possible past state
next_state_prob_list[next_state] = max_prob
shape_memory_list[next_state] = max_shape
print '##############'
computed = sum(map(operator.mul,max(state_prob_list.iteritems(),key = operator.itemgetter(1))[0],power_usage))
observed = sum(map(operator.mul,max(boot_state_prob_list[i_seg].iteritems(),key = operator.itemgetter(1))[0],power_usage))
print max(state_prob_list.iteritems(),key = operator.itemgetter(1))[0]
print max(boot_state_prob_list[i_seg].iteritems(),key = operator.itemgetter(1))[0]
print computed
print observed
#if (float(computed)-float(observed))/float(observed)<0.3:
state_prob_list = next_state_prob_list
# this code is underdevelopped
#else:
#print "potential error in change point detected "
#next_state = max(boot_state_prob_list[i_seg].iteritems(),key = operator.itemgetter(1))[0] # this is too brutal we impose the next state in this case?
#for shape_code, shape_prob in shape_prob_list[i_seg].items():# loop through all possible changes...
# #print obs_mat,shape_code
# change_state = np.dot(obs_mat, shape_code) # if the obs_mat is identity matrix then the change_state = shape_code
# past_state = tuple(np.subtract(next_state, change_state)) # find out the corresponding past_state
# if past_state in state_prob_list: # the past state should be all positive
# if state_prob_list[past_state] * shape_prob > max_prob:
# max_prob = state_prob_list[past_state] * shape_prob
# max_past_state = past_state
# max_shape = shape_code
# in this case we don't need the information from the shape_prob_list since the missing change point leads to polution of information
#max_past_state = max(state_prob_list_list[-1].iteritems(),key = operator.itemgetter(1))[0]
#max_shape = tuple(np.subtract(next_state, max_past_state))
#state_memory_list[next_state] = max_past_state
#shape_memory_list[next_state] = max_shape
#state_prob_list = boot_state_prob_list[i_seg]
#print sum(map(operator.mul,max(next_state_prob_list.iteritems(),key = operator.itemgetter(1))[0],power_usage))
print '##############'
state_prob_list_list.append(next_state_prob_list)
state_memory_list_list.append(state_memory_list)
shape_memory_list_list.append(shape_memory_list)
trace_list = []
shape_list = []
end_state = sorted(state_prob_list_list[-1].items(), key=operator.itemgetter(1))[-1][0]
trace_list.insert(0, end_state)
for i in reversed(range(n_seg)):
max_shape = shape_memory_list_list[i][end_state]
end_state = state_memory_list_list[i][end_state]
trace_list.insert(0, end_state)
shape_list.insert(0, max_shape)
return trace_list, shape_list
state_prob_list = generate_state_prob_list(4)
n_equipment_type = 4
n_equipment = [2,2,2,2]
#obs_mat = np.array([[1,0,0,0,0],[0,1,0,0,0],[0,0,1,0,0],[0,0,0,1,0],[0,0,0,0,1]])
obs_mat = np.array([[1,0,0,0],[0,1,0,0],[0,0,1,0],[0,0,0,1]])
for item,keys in state_prob_list.iteritems():
state_prob_list[item] = new_shape_prob_list[0][item]
#trace_list, shape_list = bcp.viterbi(shape_prob_list, new_shape_prob_list[0], data_seg, obs_mat)
trace_list, shape_list = viterbi_new(shape_prob_list,state_prob_list,new_shape_prob_list,data_seg,obs_mat,power_usage)
#trace_list, shape_list = viterbi_for_missing_change_point(shape_prob_list,state_prob_list,new_shape_prob_list,data_seg,obs_mat,power_usage,alpha = 100)
print 'starting', trace_list[0]
for i in range(0, len(shape_list)):
print 'through', shape_list[i], 'become', trace_list[i+1]
print
print trace_list
print shape_list
```
# Compare with exact dissag data
```
filename='/Users/Simo/Desktop/equota/disaggrREDD/house1_output15s'
filext='.dat'
f_input = file(filename+filext,'r')
#AppNo=[3,4,5,6,7,8,9,11,12,13,14,15,16,17,18,19,20] #Choose App#
period=1440
day = 67
N=day
ytemp = []
for ii in range(3,20):
f_input = file(filename+filext,'r')
(t_all,y_all)=np.array(readfile(f_input,[i-2 for i in [ii]]))
t=np.array([i+1 for i in range(period)])
ytemp.append(y_all[N*period:(N+1)*period])
correct_map = construct_equip_to_shape_map(cluster_mean,ytemp)
# artificial fix
#correct_map[3] = 3
#correct_map[7] = 4
#correct_map[8] = 4
#cluster_mean
#mean_list=[np.array([y for y in x if y>0]).mean() for x in ytemp]
#[i.max() for i in ytemp]
import operator
from collections import defaultdict
def l2_distance(list_1, list_2, last_point_w = 50, n=2):
dis = 0
tmp = [(x-y)**n for x,y in zip(list_1, list_2)]
dis = np.sum( tmp )
if len(list_1) >= len(list_2):
dis+=last_point_w*(list_1[-1] - list_2[-1])**n
return dis / (len(tmp)+last_point_w)
# usage_list = {'1':[50,100],'2':[1500,1700]...}
def backend_construct_equip_to_shape_map(cluster_mean,usage_list,shape_max = 2):
equip_to_shape_map = {}
for appliance_name,appliance_usage in usage_list.iteritems():
equip_to_shape_map[appliance_name] = []
usage_array = [float(appliance_usage[0]+appliance_usage[1])/2.0 for i in cluster_mean_2[0]]
distance_dict = {row+1: l2_distance(cluster_mean[row],usage_array,last_point_w = 1) for row in range(len(cluster_mean_2))}
local_shape_max = shape_max
while local_shape_max>0:
temp = min(distance_dict.iteritems(),key = operator.itemgetter(1))[0]
equip_to_shape_map[i].append(temp)
del distance_dict[temp]
local_shape_max=local_shape_max-1
return equip_to_shape_map
# add new cluster_result to old cluster_result
def backend_add_new_cluster_result(old_cluster_mean,new_cluster_mean):
for new_data in new_cluster_mean:
distance_dict = {row+1: l2_distance(old_cluster_mean[row],new_data,last_point_w = 1) for row in range(len(old_cluster_mean))}
temp_min_distance = min(distance_dict.iteritems(),key = operator.itemgetter(1))[1]
if float(temp_min_distance)/float(l2_distance([0 for i in old_cluster_mean[0]],new_data))>0.1:
old_cluster_mean.append(new_data)
#temp = min(distance_dict.iteritems(),key = operator.itemgetter(1))[0]
return new_data
def construct_equip_to_shape_map(cluster_mean_2,exact_usage_list):
equip_to_shape_map = {}
#equip_to_shape_map = defaultdict(list)
#picked_list = [i for i in range(1,len(exact_usage_list)+1)]
# only use non zero value to compute mean
#mean_list=[np.array([y for y in x if y>0]).mean() for x in exact_usage_list]
mean_list=[i.mean() for i in cluster_mean_2]
#mean_list = [x.mean() for x in exact_usage_list]
#
for i in range(1,len(exact_usage_list)+1):
equip_to_shape_map[i] = []
#mean = cluster_mean_2[i-1].mean()
non_zero_array = np.array([y for y in exact_usage_list[i-1] if y>0])
#print non_zero_array
distance_dict = { row+1 : (exact_usage_list[i-1].max()-mean_list[row])*(exact_usage_list[i-1].max()-mean_list[row]) for row in range(len(cluster_mean_2))}
#distance_dict = { row+1 : (non_zero_array.mean()-mean_list[row])*(non_zero_array.mean()-mean_list[row]) for row in range(len(cluster_mean_2))}
temp = min(distance_dict.iteritems(),key = operator.itemgetter(1))[0]
equip_to_shape_map[i].append(temp)
del distance_dict[temp]
temp = min(distance_dict.iteritems(),key = operator.itemgetter(1))[0]
equip_to_shape_map[i].append(temp)
del distance_dict[temp]
temp = min(distance_dict.iteritems(),key = operator.itemgetter(1))[0]
equip_to_shape_map[i].append(temp)
#picked_list.remove(equip_to_shape_map[i])
return equip_to_shape_map
def construct_shape_to_equip_map(cluster_mean_2,exact_usage_list):
equip_to_shape_map = {}
picked_list = [i for i in range(1,len(exact_usage_list)+1)]
# only use non zero value to compute mean
mean_list=[np.array([y for y in x if y>0]).mean() for x in exact_usage_list]
#mean_list = [x.mean() for x in exact_usage_list]
for i in range(1,len(cluster_mean_2)+1):
mean = cluster_mean_2[i-1].mean()
distance_dict = { row : (mean_list[row-1]-mean)*(mean_list[row-1]-mean) for row in picked_list }
equip_to_shape_map[i] = min(distance_dict.iteritems(),key = operator.itemgetter(1))[0]
picked_list.remove(equip_to_shape_map[i])
return equip_to_shape_map
def get_n_day_origin_data(filename='/Users/Simo/Desktop/equota/disaggrREDD/house1_output15s',filext='.dat',period = 1440,N = 65):
f_input = file(filename+filext,'r')
#AppNo=[3,4,5,6,7,8,9,11,12,13,14,15,16,17,18,19,20] #Choose App#
ytemp = []
for ii in range(3,20):
f_input = file(filename+filext,'r')
(t_all,y_all)=np.array(readfile(f_input,[i-2 for i in [ii]]))
t=np.array([i+1 for i in range(period)])
ytemp.append(y_all[N*period:(N+1)*period])
return ytemp
# total correct / total usage for the day
def compute_total_error(period,ytemp,predicted_profile,equip_to_shape_map,total):
correct_list = []
sum = 0
total_sum = total.sum()
for i in range(period):
#total_sum+=total[i]
for j in range(len(predicted_profile)-1):
sum += max(min(predicted_profile[-1][i],ytemp[j][i]),max([min(ytemp[j][i],predicted_profile[k-1][i]) for k in equip_to_shape_map[j+1]]))
#sum += max(min(predicted_profile[-1][i],ytemp[j][i]),max([min(ytemp[j][i],predicted_profile[k-1][i]) for k in equip_to_shape_map[j+1]]))
#sum +=min(ytemp[j][i],predicted_profile[equip_to_shape_map[j+1]-1][i])#min(ytemp[equip_to_shape_map[j+1]-1][i],predicted_profile[j][i])
#total+= predicted_profile[j][i]
#error.append(abs(ytemp[j][i]-predicted_profile[equip_to_shape_map[j+1]-1][i])/ytemp[j][i])
#correct_list.append(float(sum)/float(total[i]))
#error_list.append(error)
return float(sum)/float(total_sum)
def compute_total_correct(period,ytemp,predicted_profile,equip_to_shape_map,total):
correct_list = []
for i in range(period):
sum = 0
for j in range(len(ytemp)-1):
#sum +=min(ytemp[j][i],predicted_profile[equip_to_shape_map[j+1]-1][i])
sum += max(min(predicted_profile[-1][i],ytemp[j][i]),max([min(ytemp[j][i],predicted_profile[k-1][i]) for k in equip_to_shape_map[j+1]]))
correct_list.append(float(sum)/float(total[i]))
return correct_list
#predicted_profile = aa
per = compute_total_error(1440,ytemp,predicted_profile,construct_equip_to_shape_map(cluster_mean_2,ytemp),y_all[(day)*period:(day+1)*period])
print per
x= compute_total_correct(1440,ytemp,predicted_profile,construct_equip_to_shape_map(cluster_mean_2,ytemp),y_all[(day)*period:(day+1)*period])
correct = np.dot(np.array(x),y_all[(day)*period:(day+1)*period])
print correct/y_all[(day)*period:(day+1)*period].sum()
plt.figure(figsize=[8, 6])
plt.plot([y*4000 for y in x],'k-', linewidth=.8, alpha=.5)
plt.plot(y_all[(day)*period:(day+1)*period],linewidth=.8, alpha=.5)
plt.xlabel('time of day')
plt.ylabel('correctness')
plt.xlim([0, 1440])
plt.ylim([0, 4000])
construct_equip_to_shape_map(cluster_mean_2,ytemp)
#ytemp
cluster_mean_2
predicted_profile
plt.figure(figsize=[8, 6])
for x in ytemp[6:8]:
plt.plot(x, 'k-', linewidth=.8, alpha=.5)
plt.xlabel('time point after change point')
plt.ylabel('relative power shift')
plt.xlim([0, 1440])
plt.ylim([-60, 500])
plt.figure(figsize=[8, 6])
for x in ytemp[0:11]:
plt.plot(x, 'k-', linewidth=.8, alpha=.5)
plt.xlabel('time point after change point')
plt.ylabel('relative power shift')
plt.xlim([0, 1440])
plt.ylim([-60, 3000])
plt.figure(figsize=[8, 6])
for x in ytemp:
plt.plot(x, 'k-', linewidth=.8, alpha=.5)
plt.xlabel('time point after change point')
plt.ylabel('relative power shift')
plt.xlim([0, 1440])
plt.ylim([-60, 2500])
# app No / power
# 1 0
# 2 0
# 3 250
# 4 1100
# 5 0
# 6 20
# 7 40 spurious
# 8 500
# 9 1500
# 10 1600
# 11 0
# 12 0
# 13 1100
# 14 1500
# 15 50
# 16 0
# 17 0
#list_of_shapes = [shape1, shape2, shap3]
#mapping variable=[[[appliance_id_1_1, appliance_id_1_2, appliance_id_1_3], [appliance_id_2_1, appliance_id_2_2, appliance_id_2_3], [appliance_id_3_1, appliance_id_3_2]]]
#appliance_list
def construct_obs_mat(list_of_shapes,mapping_variable,appliance_list):
obs_mat = []
for i in range(len(appliance_list)):
row = [0 for jj in range(len(list_of_shapes))]
for k in mapping_variable[i]:
row[k-1] = 1
obs_mat.append(row)
return obs_mat
def wrapped_integrated_dissagregate(y_all,appliance_list,mapping_variable,power_usage,list_of_shapes,period = 1440,day = 65):
predicted_profile = integrated_dissagregate(y_all,period,list_of_shapes,day = day,n_equipment_type = len(appliance_list),n_equipment=[2 for i in range(1,len(appliance_list)+1)],obs_mat=construct_obs_mat(list_of_shapes,mapping_variable,appliance_list),power_usage = power_usage)
toreturn = {}
for i in range(len(appliance_list)):
toreturn[appliance_list[i]] = predicted_profile[i]
toreturn['other'] = predicted_profile[-1]
return toreturn
reload(bcp)
list_of_shapes = cluster_mean_2
appliance_list = [1,2,3,4,5]
mapping_variable = [[1],[2],[3],[4],[5]] # in this case this is a one-to-one mapping
power_usage = [i.mean() for i in cluster_mean_2] # you need to have the information about the power usage for each appliances
predicted_profile = wrapped_integrated_dissagregate(y_all,appliance_list,mapping_variable,power_usage,list_of_shapes,period = 1440,day = 67)
plot_dissagregation_2(predicted_profile,np.array([i+1 for i in range(period)]))
[dutu[np.argpartition(np.array(dutu),int(len(dutu)/50))[int(len(dutu)/50)]] for _ in range(len(dutu))]
dutu = [1,2,3,4,5,6,7,8,9,10]
construct_obs_mat(list_of_shapes,mapping_variable,appliance_list)
aa = []
for key,value in predicted_profile.iteritems():
aa.append(value)
cluster_mean_2
```
| github_jupyter |
# Continuous Control
---
You are welcome to use this coding environment to train your agent for the project. Follow the instructions below to get started!
### 1. Start the Environment
Run the next code cell to install a few packages. This line will take a few minutes to run!
```
!pip -q install ./python
```
The environments corresponding to both versions of the environment are already saved in the Workspace and can be accessed at the file paths provided below.
Please select one of the two options below for loading the environment.
```
from unityagents import UnityEnvironment
import numpy as np
# select this option to load version 1 (with a single agent) of the environment
env = UnityEnvironment(file_name='/data/Reacher_One_Linux_NoVis/Reacher_One_Linux_NoVis.x86_64')
# select this option to load version 2 (with 20 agents) of the environment
# env = UnityEnvironment(file_name='/data/Reacher_Linux_NoVis/Reacher.x86_64')
```
Environments contain **_brains_** which are responsible for deciding the actions of their associated agents. Here we check for the first brain available, and set it as the default brain we will be controlling from Python.
```
# get the default brain
brain_name = env.brain_names[0]
brain = env.brains[brain_name]
```
### 2. Examine the State and Action Spaces
Run the code cell below to print some information about the environment.
```
# reset the environment
env_info = env.reset(train_mode=True)[brain_name]
# number of agents
num_agents = len(env_info.agents)
print('Number of agents:', num_agents)
# size of each action
action_size = brain.vector_action_space_size
print('Size of each action:', action_size)
# examine the state space
states = env_info.vector_observations
state_size = states.shape[1]
print('There are {} agents. Each observes a state with length: {}'.format(states.shape[0], state_size))
print('The state for the first agent looks like:', states[0])
```
### 3. Take Random Actions in the Environment
In the next code cell, you will learn how to use the Python API to control the agent and receive feedback from the environment.
Note that **in this coding environment, you will not be able to watch the agents while they are training**, and you should set `train_mode=True` to restart the environment.
```
env_info = env.reset(train_mode=True)[brain_name] # reset the environment
states = env_info.vector_observations # get the current state (for each agent)
scores = np.zeros(num_agents) # initialize the score (for each agent)
while True:
actions = np.random.randn(num_agents, action_size) # select an action (for each agent)
actions = np.clip(actions, -1, 1) # all actions between -1 and 1
env_info = env.step(actions)[brain_name] # send all actions to tne environment
next_states = env_info.vector_observations # get next state (for each agent)
rewards = env_info.rewards # get reward (for each agent)
dones = env_info.local_done # see if episode finished
scores += env_info.rewards # update the score (for each agent)
states = next_states # roll over states to next time step
if np.any(dones): # exit loop if episode finished
break
print('Total score (averaged over agents) this episode: {}'.format(np.mean(scores)))
```
When finished, you can close the environment.
```
env.close()
```
### 4. It's Your Turn!
Now it's your turn to train your own agent to solve the environment! A few **important notes**:
- When training the environment, set `train_mode=True`, so that the line for resetting the environment looks like the following:
```python
env_info = env.reset(train_mode=True)[brain_name]
```
- To structure your work, you're welcome to work directly in this Jupyter notebook, or you might like to start over with a new file! You can see the list of files in the workspace by clicking on **_Jupyter_** in the top left corner of the notebook.
- In this coding environment, you will not be able to watch the agents while they are training. However, **_after training the agents_**, you can download the saved model weights to watch the agents on your own machine!
```
# Import the Necessary Packages
import time
import numpy as np
import torch
import matplotlib.pyplot as plt
from collections import deque
from ddpg_agent import Agent
```
### Instantiate the agent
```
agent = Agent(state_size=33, action_size=4, random_seed=2)
```
### Train the Agent with a DDPG Agent (Deep Deterministic Policy Gradients)
```
def ddpg(n_episodes=2000, max_t=1000, print_every=100, target_score=30):
"""Deep Deterministic Policy Gradients.
Params
======
n_episodes (int): maximum number of training episodes
max_t (int): maximum number of timesteps per episode
print_every (int): interval to display results
"""
scores = [] # list containing scores from each episode
scores_window = deque(maxlen=100) # last 100 scores
for i_episode in range(1, n_episodes+1):
env_info = env.reset(train_mode=True)[brain_name]
states = env_info.vector_observations # get the current state (for each agent)
score = np.zeros(num_agents) # initialize the score (for each agent)
for t in range(max_t):
actions = agent.act(states) # select an action
env_info = env.step(actions)[brain_name] # send the action to the environment
next_states = env_info.vector_observations # get next state (for each agent)
rewards = env_info.rewards # get reward (for each agent)
dones = env_info.local_done # see if episode has finished
agent.step(states, actions, rewards, next_states, dones) # Do the learning
score += rewards # update the score
states = next_states # get the next state
if np.any(dones):
break
scores_window.append(score) # save most recent score
scores.append(score) # save most recent score
print('\rEpisode {}\tAverage Score: {:.2f}'.format(i_episode, np.mean(scores_window)), end="")
if i_episode % print_every == 0:
print('\rEpisode {}\tAverage Score: {:.2f}'.format(i_episode, np.mean(scores_window)))
# Our agent must get an average score of +30
if np.mean(scores_window) >= target_score:
print("\nEnvironment solved in {:d} episodes!\t Average score: {:.2f}".format(i_episode, np.mean(scores_window)))
torch.save(agent.actor_local.state_dict(), 'checkpoint_actor.pth')
torch.save(agent.critic_local.state_dict(), 'checkpoint_critic.pth')
break
return scores
scores = ddpg()
# plot the scores
fig = plt.figure(figsize=(12, 8))
ax = fig.add_subplot(111)
plt.plot(np.arange(1, len(scores)+1), scores)
plt.ylabel('Score')
plt.xlabel('Episode Number')
plt.show()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/parampopat/Wiki-Movies/blob/master/movie_generator.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
"""
__author__ = "Param Popat"
__version__ = "1"
__git__ = "https://github.com/parampopat/"
"""
import numpy
import sys
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import Dropout
from keras.layers import LSTM
from keras.callbacks import ModelCheckpoint
from keras.utils import np_utils
import pandas as pd
from google.colab import drive
drive.mount('/content/drive')
# load ascii text and covert to lowercase
movieNames = pd.read_csv("/content/drive/My Drive/Wiki-Movies/Plot_associations.csv").iloc[:, 1].values
raw_text = ""
for filename in movieNames:
raw_text += open("/content/drive/My Drive/Wiki-Movies/Plot/" + filename + '.txt').read()
raw_text = raw_text.lower()
# create mapping of unique chars to integers
chars = sorted(list(set(raw_text)))
char_to_int = dict((c, i) for i, c in enumerate(chars))
int_to_char = dict((i, c) for i, c in enumerate(chars))
# summarize the loaded data
n_chars = len(raw_text)
n_vocab = len(chars)
print(
"Total Characters: ", n_chars)
print(
"Total Vocab: ", n_vocab)
# prepare the dataset of input to output pairs encoded as integers
seq_length = 100
dataX = []
dataY = []
for i in range(0, n_chars - seq_length, 1):
seq_in = raw_text[i:i + seq_length]
seq_out = raw_text[i + seq_length]
dataX.append([char_to_int[char] for char in seq_in])
dataY.append(char_to_int[seq_out])
n_patterns = len(dataX)
print("Total Patterns: ", n_patterns)
# reshape X to be [samples, time steps, features]
X = numpy.reshape(dataX, (n_patterns, seq_length, 1))
# normalize
X = X / float(n_vocab)
# one hot encode the output variable
y = np_utils.to_categorical(dataY)
model = Sequential()
model.add(LSTM(256, input_shape=(X.shape[1], X.shape[2]), return_sequences=True))
model.add(Dropout(0.2))
model.add(LSTM(256))
model.add(Dropout(0.2))
model.add(Dense(y.shape[1], activation='softmax'))
model.compile(loss='categorical_crossentropy', optimizer='adam')
# define the checkpoint
filepath = "/content/drive/My Drive/Wiki-Movies/trained/weights-improvement-{epoch:02d}-{loss:.4f}-bigger.hdf5"
checkpoint = ModelCheckpoint(filepath, monitor='loss', verbose=1, save_best_only=True, mode='min')
callbacks_list = [checkpoint]
# fit the model
model.fit(X, y, epochs=50, batch_size=64, callbacks=callbacks_list)
start = numpy.random.randint(0, len(dataX) - 1)
pattern = dataX[start]
print("Seed:")
print("\"", ''.join([int_to_char[value] for value in pattern]), "\"")
# generate characters
for i in range(1000):
x = numpy.reshape(pattern, (1, len(pattern), 1))
x = x / float(n_vocab)
prediction = model.predict(x, verbose=0)
index = numpy.argmax(prediction)
result = int_to_char[index]
seq_in = [int_to_char[value] for value in pattern]
sys.stdout.write(result)
pattern.append(index)
pattern = pattern[1:len(pattern)]
print("\nDone.")
```
| github_jupyter |
# Incremental Principal component analysis with StandardScaler
This code template is for Incremental Principal Component Analysis(IncrementalPCA) along feature scaling StandardScaler in python for dimensionality reduction technique. It is used to decompose a multivariate dataset into a set of successive orthogonal components that explain a maximum amount of the variance, keeping only the most significant singular vectors to project the data to a lower dimensional space.
### Required Packages
```
import warnings
import itertools
import numpy as np
import pandas as pd
import seaborn as se
import matplotlib.pyplot as plt
from mpl_toolkits import mplot3d
from sklearn.decomposition import IncrementalPCA
from sklearn.preprocessing import LabelEncoder,StandardScaler
warnings.filterwarnings('ignore')
```
### Initialization
Filepath of CSV file
```
#filepath
file_path= ""
```
List of features which are required for model training .
```
#x_values
features=[]
```
Target feature for prediction.
```
#y_value
target=''
```
### Data Fetching
Pandas is an open-source, BSD-licensed library providing high-performance, easy-to-use data manipulation and data analysis tools.
We will use panda's library to read the CSV file using its storage path.And we use the head function to display the initial row or entry.
```
df=pd.read_csv(file_path)
df.head()
```
### Feature Selections
It is the process of reducing the number of input variables when developing a predictive model. Used to reduce the number of input variables to both reduce the computational cost of modelling and, in some cases, to improve the performance of the model.
We will assign all the required input features to X and target/outcome to Y.
```
X = df[features]
Y = df[target]
```
### Data Preprocessing
Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
```
def NullClearner(df):
if(isinstance(df, pd.Series) and (df.dtype in ["float64","int64"])):
df.fillna(df.mean(),inplace=True)
return df
elif(isinstance(df, pd.Series)):
df.fillna(df.mode()[0],inplace=True)
return df
else:return df
def EncodeX(df):
return pd.get_dummies(df)
x=X.columns.to_list()
for i in x:
X[i]=NullClearner(X[i])
X=EncodeX(X)
Y=NullClearner(Y)
X.head()
```
#### Correlation Map
In order to check the correlation between the features, we will plot a correlation matrix. It is effective in summarizing a large amount of data where the goal is to see patterns.
```
f,ax = plt.subplots(figsize=(18, 18))
matrix = np.triu(X.corr())
se.heatmap(X.corr(), annot=True, linewidths=.5, fmt= '.1f',ax=ax, mask=matrix)
plt.show()
```
### Data Rescaling
StandardScaler standardizes features by removing the mean and scaling to unit variance.
Standardization of a dataset is a common requirement for many machine learning estimators: they might behave badly if the individual features do not more or less look like standard normally distributed data (e.g. Gaussian with 0 mean and unit variance).
Centering and scaling happen independently on each feature by computing the relevant statistics on the samples in the training set. Mean and standard deviation are then stored to be used on later data using transform.
##### For more information on StandardScaler [ click here](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.StandardScaler.html)
```
X_Scaled=StandardScaler().fit_transform(X)
X_Scaled=pd.DataFrame(data = X_Scaled,columns = X.columns)
X_Scaled.head()
```
### Choosing the number of components
A vital part of using Incremental PCA in practice is the ability to estimate how many components are needed to describe the data. This can be determined by looking at the cumulative explained variance ratio as a function of the number of components.
This curve quantifies how much of the total, dimensional variance is contained within the first N components.
```
ipcaComponents = IncrementalPCA().fit(X_Scaled)
plt.plot(np.cumsum(ipcaComponents.explained_variance_ratio_))
plt.xlabel('number of components')
plt.ylabel('cumulative explained variance');
```
#### Scree plot
The scree plot helps you to determine the optimal number of components. The eigenvalue of each component in the initial solution is plotted. Generally, you want to extract the components on the steep slope. The components on the shallow slope contribute little to the solution.
```
IPC_values = np.arange(ipcaComponents.n_components_) + 1
plt.plot(IPC_values, ipcaComponents.explained_variance_ratio_, 'ro-', linewidth=2)
plt.title('Scree Plot')
plt.xlabel('Principal Component')
plt.ylabel('Proportion of Variance Explained')
plt.show()
```
# Model
Incremental principal components analysis (IPCA) allows for linear dimensionality reduction using Singular Value Decomposition of the data, keeping only the most significant singular vectors to project the data to a lower dimensional space. The input data is centered but not scaled for each feature before applying the SVD.
Incremental principal component analysis (IPCA) is typically used as a replacement for principal component analysis (PCA) when the dataset to be decomposed is too large to fit in memory.
#### Tunning parameters reference :
[API](https://scikit-learn.org/stable/modules/generated/sklearn.decomposition.IncrementalPCA.html)
```
ipca = IncrementalPCA(n_components=10)
ipcaX = pd.DataFrame(data = ipca.fit_transform(X_Scaled))
```
#### Output Dataframe
```
finalDf = pd.concat([ipcaX, Y], axis = 1)
finalDf.head()
```
#### Creator: Saharsh Laud , Github: [Profile](https://github.com/SaharshLaud)
| github_jupyter |
# Survival Analysis in Python
Allen B. Downey
[MIT License](https://en.wikipedia.org/wiki/MIT_License)
### Introduction
This notebook reviews two ways we have previously seen to represent a distribution, PMFs and CDFs, and introduces two new ways, survival functions and hazard functions.
All four representation are equivalent in the sense that if you are given any of them, you can compute the others. So you might wonder why we need four ways to represent the same information. There are two reasons:
1. Each representation is useful for computing different values and answering different questions. In this notebook I'll show you what each representation is good for.
2. In some cases we can use a sample to compute a PMF, and use the PMF to compute the other representations. In other cases it is easier to estimate the hazard function and use it to compute the others.
In this notebook I will demonstrate the first process, starting with the PMF. In the next notebook we'll see the second process, starting with the hazard function.
### Setup
If you are running this notebook in Colab, the following cell installs the `empiricaldist` library, which provides objects that represent PMFs, CDFs, survival functions, and hazard functions.
If you are running in another environment, you will need to install it yourself.
```
# If we're running in Colab, set up the environment
import sys
IN_COLAB = 'google.colab' in sys.modules
if IN_COLAB:
!pip install empiricaldist
```
If everything we need is installed, the following cell should run without error.
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from empiricaldist import Pmf, Cdf, Surv, Hazard
```
The following is a function I use to decorate the axes of a figure.
```
def decorate(**options):
"""Decorate the current axes.
Call decorate with keyword arguments like
decorate(title='Title',
xlabel='x',
ylabel='y')
The keyword arguments can be any of the axis properties
https://matplotlib.org/api/axes_api.html
"""
plt.gca().set(**options)
plt.tight_layout()
```
## Light bulb lifetimes
As a first example, I'll use I'll use data from an experiment that measures the lifetimes of 50 light bulbs.
I downloaded the data from from [https://gist.github.com/epogrebnyak/7933e16c0ad215742c4c104be4fbdeb1], which includes this documentation:
```
Dataset from:
V.J. Menon and D.C. Agrawal, Renewal Rate of Filament Lamps:
Theory and Experiment. Journal of Failure Analysis and Prevention.
December 2007, p. 421, Table 2/
DOI: 10.1007/s11668-007-9074-9
Description:
An assembly of 50 new Philips (India) lamps with the
rating 40 W, 220 V (AC) was taken and installed in the horizontal
orientation and uniformly distributed over a lab area 11 m by 7 m.
The assembly was monitored at regular intervals of 12 h to
look for failures. The instants of recorded failures were
called t‘ and a total of 32 data points were obtained such
that even the last bulb failed.
Variables:
i - observation number
h - time in hours since experiment start
f - number of failed lamps at particular time h
K - number of surviving lamps at particular time h
```
Because of the design of this experiment, we can consider the data to be a representative sample from the distribution of lifetimes, at least for light bulbs that are lit continuously.
The following cell downloads the data from a repository on GitHub (thanks to Evgeny Pogrebnyak for making it available).
If that fails, you should be able to [get it from my repository](https://github.com/AllenDowney/SurvivalAnalysisPython/raw/master/lamps.csv).
```
# Load the data file
import os
datafile = 'lamps.csv'
if not os.path.exists(datafile):
!wget https://gist.github.com/epogrebnyak/7933e16c0ad215742c4c104be4fbdeb1/raw/c932bc5b6aa6317770c4cbf43eb591511fec08f9/lamps.csv
```
We can load the data from the CSV file.
```
df = pd.read_csv('lamps.csv', index_col=0)
df.head()
```
This dataset is complete in the sense that the experiment ran until all light bulbs failed, so the lifetime for every bulb is known.
## PMF
In that case we can make a `Pmf` object that represents the distribution of lifetimes. The result is a series that contains the lifetimes (column `h`) as quantities and the normalized number of failures (column `f`) as probabilities.
```
pmf = Pmf(df['f'].values, index=df['h'])
pmf.normalize()
pmf.head()
```
For a given quantity, `q`, the PMF computes the fraction of the population at that quantity, exactly.
For example, we can compute the fraction of light bulbs that lasted 840 hours:
```
pmf(840)
```
Because of the way the experiment was conducted, all quantities are multiples of 12 hours. If we ask about a quantity that is not a multiple of 12, the result is 0.
```
pmf(841)
```
Here's what the PMF looks like.
```
pmf.bar(width=10)
decorate(xlabel='Lifetime (hours)',
ylabel='PMF',
title='PMF of lightbulb lifetimes')
```
This way of visualizing the data can be useful for validation, and it gives a sense of the location and spread of the data. But it does not make the shape of the distrbution clear. For that, the CDF is better.
## CDF
The CDF is the cumulative sum of the PMF. We could compute it ourselves like this:
```
cdf = pmf.cumsum()
cdf.head()
```
In that case the result is a Pandas Series.
We can also use `Pmf.make_cdf`, which does the same calculation, but the result is a `Cdf` object.
```
cdf = pmf.make_cdf()
cdf.head()
```
For a given quantity, `q`, the CDF computes the fraction of the population less than or equal to `q`.
For example, here is the fraction of light bulbs that expired at or before 840 hours:
```
cdf(840)
```
And the fraction that expired at or before 852 hours.
```
cdf(852)
```
For any quantity `q`, the CDF computes the corresponding cumulative probability, `p`.
But it also works the other way. For a given probability, `p`, you can use the inverse CDF to compute the corresponding quantity, `q`.
```
cdf.inverse(0.06)
```
With `p=0.5`, the result is the 50th percentile, or median:
```
cdf.inverse(0.5)
```
We can also use `inverse` to compute the quartiles, which are the quantities at the 25th, 50th, and 75th percentiles:
```
cdf.inverse([0.25, 0.5, 0.75])
```
In my opinion, the CDF is often the best way to visualize the distribution of a sample.
Here it is, plotted as a step function.
```
cdf.step()
decorate(xlabel='Lifetime (hours)',
ylabel='CDF',
title='CDF of lightbulb lifetimes')
cdf.inverse([0.05, 0.95])
```
Between 800 and 1800 hours, the CDF is roughly a straight line, which suggests that the distribution is uniform in this range.
## Survival function
The survival function is the complement of the CDF; that is, for a given quantity, `q`:
* The CDF is the fraction of the population less than or equal to `q`.
* The survival function is the fraction (strictly) greater than `q`.
Again, we can compute the survival function using Pandas Series operators:
```
surv = 1-cdf
surv.head()
```
We can also use `Cdf.make_surv`, which does the same calculation and returns a `Surv` object:
```
surv = cdf.make_surv()
surv.head()
surv[0]
```
We can use `surv` to compute the fraction of lightbulbs that live longer than 936 hours:
```
surv(936)
```
And confirm that the CDF and survival functions add up to 1 (at least within floating-point error).
```
cdf(936) + surv(936)
```
If we plot the survival function along with the CDF, we can see that they complement each other.
```
cdf.step(color='gray', alpha=0.4)
surv.step()
decorate(xlabel='Lifetime (hours)',
ylabel='Prob(lifetime>t)',
title='Survival function of lightbulb lifetimes')
```
Most often, a quantity in a survival function represents a lifetime of some kind. In some cases it is literally the time until a death; in other cases "lifetime" is more metaphorical. It might represent the time until a device breaks, or the time until a particular event occurs, even if the event has nothing to do with life or death.
For example, [in one project I worked on](http://allendowney.blogspot.com/2015/07/will-millennials-ever-get-married.html), I used survival curves to represent people's age at first marriage.
It might not be obvious why the survival function is useful. Given the CDF and a little arithmetic, we can answer all of the same questions.
There are two reasons:
* In some domains it is more natural, or at least conventional, to represent distributions in terms of survival rates.
* The survival function is a step on the way to the hazard function, which we'll get to now.
## Hazard function
For a given time `q`, the hazard function computes the "hazard rate" at `q`. Using the vocabulary of the light bulb example, the hazard rate is the fraction of light bulbs that survive until `q` and then fail at `q`.
We can compute the hazard rate by computing these quantities:
* `pmf(q)` is the fraction of light bulbs that fail at `q`.
* `surv(q)` is the fraction of light bulbs that live longer than `q`.
* The sum, `pmf(q) + surv(q)` is the fraction that survive until `q`.
So the hazard rate is the ratio of `pmf(q)` to the sum `pmf(q) + surv(q)`.
We can compute it like this:
```
haz = pmf / (pmf + surv)
haz.head()
```
Or we can use `Surv.make_hazard`, which does the same thing and returns a `Hazard` object:
```
haz = surv.make_hazard()
haz.head()
```
Here's what the hazard function looks like:
```
haz.bar(width=10)
decorate(xlabel='Lifetime (hours)',
ylabel='Hazard rate',
title='Hazard function of lightbulb lifetimes')
```
This way of visualizing the hazard function doesn't work very well; in the next section we'll see if we can do better.
## Visualizing the hazard function
With this kind of data, plotting the hazard function does not provide a clear picture of what's happening. There are two problems:
1) The plot shows spikes at the locations of the data, but it is hard to see the shape of the curve.
2) The large values on the right are unreliable because they are based on a small number of values.
To explain the second point, let's look at the last few rows of the failure column, `f`:
```
df.tail()
```
We can see that one bulb failed at each of 1812, 1836, 1860, 1980, and 2568 hours.
So the last value of the hazard function is based on only one bulb, the second-to-last point is based on 2 bulbs, and so on.
To get a better sense of the shape of the curve, we can plot the cumulative hazard function, like this:
```
haz.cumsum().plot()
decorate(xlabel='Lifetime (hours)',
ylabel='Cumulative hazard rate',
title='Hazard function of lightbulb lifetimes')
```
The slope of the cumulative hazard function is proportional to the hazard rate, so we can use this curve to see when the hazard rate is low or high, and when it is increasing or decreasing.
Between 0 and 1000 hours, the slope is low, so the hazard rate is low.
Between 1000 and 2000 hours, the slope and hazard rate are increasing.
But notice that the vertical scale goes to 4. You might wonder that that means; the answer is "not much".
The values of the hazard function are rates, that is, percentages of light bulbs that expire at each point in time. When you add up these rates, the result does not have a clear interpretation.
When you look at a cumulative hazard function, you should pay attention to the slope of the curve and ignore the values.
## Resampling
In the previous section, I suggested that we have to be careful not to overinterpret the right side of the hazard function, because it is based on a small number of data points.
To see how precise the estimated hazard function is, we can use resampling.
First I will use the Pmf of lifetimes to make a kernel density estimate (KDE) of the distribution.
```
from scipy.stats import gaussian_kde
kde = gaussian_kde(pmf.qs, weights=pmf.ps)
```
We can use the KDE to draw a new sample of lifetimes, with the same size as the original data set, and plot the survival curve for each sample.
```
size = df['f'].sum()
for i in range(100):
sample = kde.resample(size).flatten()
sf = Surv.from_seq(sample)
sf.plot(color='gray', alpha=0.1)
surv.step()
decorate(xlabel='Lifetime (hours)',
ylabel='Prob(lifetime>t)',
title='Survival function of lightbulb lifetimes')
```
By plotting the resampled survival curves on top of each other, we can get a sense of how much the results vary due to random sampling.
In this case the sample size is small, so the differences between resampled survival curves are substantial.
We can do the same thing with the hazard function:
```
for i in range(100):
sample = kde.resample(size).flatten()
hf = Hazard.from_seq(sample)
hf.cumsum().plot(color='gray', alpha=0.1)
haz.cumsum().plot()
decorate(xlabel='Lifetime (hours)',
ylabel='Cumulative hazard rate',
title='Hazard function of lightbulb lifetimes')
```
This plot gives us a sense of which parts of the cumulative hazard function are reliable and which are not.
Below 2000 hours, all of the resampled curves are similar; they increase with increasing slope.
After that, the variability of the curves is much wider, which means we don't have enough data to characterize this part of the hazard function.
## Summary
We have seen five ways to represent a distribution of lifetimes:
* Probability mass function (PMF), which maps from each value in the distribution to its probability.
* Cumulative distribution function (CDF), which maps from each value, `x`, to its cumulative probability, that is, the probability of being less than or equal to `x`.
* Survival function, which is the complement of the CDF; that is, the probability of exceeding `x`.
* Hazard function, which is the number of failures at `x` as a fraction of the number of cases that survive until `x`.
* Cumulative hazard function, which is useful for visualizing the shape of the hazard curve.
These representations of a distribution are equivalent in the sense that they all contain the same information. Given any of them, we can compute any of the others.
In this notebook, we computed the `Pmf` directly from the data, then computed the `Cdf`, survival function, and hazard function, in that order.
But we can also go the other way, starting with the hazard function and working backwards. To demonstrate, I'll use `haz` to compute the survival function, and confirm that the result is the same as `surv`, within floating point error.
```
surv2 = haz.make_surv()
max(abs(surv - surv2))
```
And we can use `surv2` to compute the CDF:
```
cdf2 = surv2.make_cdf()
max(abs(cdf - cdf2))
```
And `cdf2` to compute the PMF:
```
pmf2 = cdf2.make_pmf()
max(abs(pmf - pmf2))
```
In the next notebook we will take advantage of this ability to deal with cases where we cannot compute the PMF directly from the data.
| github_jupyter |
```
import sys
sys.path.append('/home/dnlab/Jupyter-Bitcoin/Heuristics/ExperimentSpeed/')
import pandas as pd
import test_cluster_db_query as cdq
import test_db_query as dq
df = pd.read_csv('/home/dnlab/DataHDD/ChainAnalysisNamedClusters.csv')[['ClusterName', 'Category', 'RootAddress']]
bdf = pd.read_csv('/home/dnlab/DataHDD/chainalaysis/20190909_Cluster_addresses_of_Bithumb.com.csv')[['Address']]
bdf
df
df['addr'] = df['RootAddress'].map(lambda x: dq.get_addrid_from_addr(x))
bdf['addr'] = bdf['Address'].map(lambda x: dq.get_addrid_from_addr(x))
df = df.dropna(axis=0)
bdf = bdf.dropna(axis=0)
df = df[['addr','ClusterName','Category']]
df[df['ClusterName'] =='Bithumb.com']
df.sort_values(by=['addr'], inplace=True)
df.to_csv('/home/dnlab/DataHDD/ChainAnalysisAddrIdClusters.csv', index=False)
data= pd.read_csv('/home/dnlab/DataHDD/cluster_result/cluster.csv')
data_condition = data['number'] != -1
data = data[data_condition]
data= data.rename(columns={"address": "addr"})
gbdata = data.groupby('number').count().reset_index()
gbdata = gbdata.rename(columns={"addr": "cluster_number", "number":"cluster_addr"})
gbdata
gbdata = gbdata.sort_values(by=['cluster_number','cluster_addr'], ascending=False)
gbdata
bdf.addr = bdf.addr.astype('int32')
bithumb = bdf['addr'].to_list()
df.addr = df.addr.astype('int32')
df
#bithumb_cluster = data[data['number']==24840655]
#bithumb_cluster_list = bithumb_cluster['addr'].to_list()
#bithumb = set(bithumb) - set(bithumb_cluster_list)
len(bithumb)
data
addr_list = []
not_clustered = []
count = 0
while len(bithumb) > 0:
addr = bithumb.pop()
if count % 100000:
print("current:",addr_list[-1], "single address:", len(not_clustered))
#print("bithumb address:", addr)
try:
cluster_number = data[data['addr'] == int(addr)]['number'].iloc[0]
cluster_list = data[data['number']== cluster_number]['addr'].to_list()
print("cluster_number:",cluster_number, len(cluster_list))
addr_list.append({"cluster_number":cluster_number, "len": len(cluster_list), "value":cluster_list})
bithumb = set(bithumb) - set(cluster_list)
print("left bithumb:", len(bithumb))
except Exception as e:
#print("ERROR: ", e)
not_clustered.append(addr)
continue
cluster_result = pd.DataFrame(addr_list)
cluster_result
addr = 562585220
cluster_number = data[data['addr'] == addr]['number'].iloc[0]
print(cluster_number)
cluster_list = data[data['number']== cluster_number]['addr'].to_list()
cluster_list
data[data['addr'] == addr].iloc[0]
data[data['addr'] == 209715241]['number'].iloc[0]
bdf
df
data
def get_address(addr):
try:
cluster_number = data[data['addr'] == addr]['number'].iloc[0]
return cluster_number
except Exception as e:
print(e)
return None
print("cluster_number:",cluster_number, len(cluster_list))
addr_list.append({"cluster_number":cluster_number, "len": len(cluster_list), "value":cluster_list})
bithumb = set(bithumb) - set(cluster_list)
print("left bithumb:", len(bithumb))
# bdf['addr'] = bdf['Address'].map(lambda x: dq.get_addrid_from_addr(x))
data['root_addr'] = bdf['addr'].map(lambda x: get_address(x))
```
| github_jupyter |
```
import os
import glob
import numpy as np
from keras.models import Sequential
from keras.layers import Dense, Activation
from datetime import datetime
import math
from sklearn.preprocessing import MinMaxScaler, StandardScaler
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import train_test_split
import pandas as pd
import matplotlib.pyplot as plt
from utils import *
%matplotlib inline
def get_day_load_data(load_path):
'''read csv data at load_path and return it in a numpy matrix'''
data = pd.read_csv(load_path)
date_ = load_path.split('/')[-1][:-4] # get date of '01-10-2017' format
data['date'] = datetime.strptime(date_, "%d-%m-%Y").date()
data['time'] += ':00'
data['datetime'] = pd.to_timedelta(data['time'])
data['hour'] = data['datetime'].dt.seconds // 3600
return data.as_matrix()
def get_30min_wise_data(day_load_data, day_whether_data):
'''Returns time, day temp, humidity and load values for each 30mins in a numpy matrix'''
data = []
for i in range(0, 288, 6): # SLDC load data has time step of 5 mins, so total 24 * (60 // 5)
load = day_load_data[i][1]
day = day_load_data[i][2].weekday()
try:
humidity = float(day_whether_data[i//6][1].replace('%', ''))
except Exception as e:
humidity = 50 # some dirt values in data
try:
temp = float(day_whether_data[i//6][0]) # i//6 because whether data is already in 30 min step size
except Exception as err:
temp = 28 # aiyvi
data.append([i, day, temp, humidity, load])
return np.asarray(data)
def get_data(data_list, shuffle=True):
'''Return data of all csv files in SLDC_Data/ folder in matrix form'''
data = []
for load_path in data_list:
try:
whether_path = load_path.replace('SLDC_Data', 'Whether_Data')
day_load_data = get_day_load_data(load_path)
day_whether_data = pd.read_csv(whether_path)
day_data = get_30min_wise_data(day_load_data, day_whether_data[['Temp.', 'Humidity']].as_matrix())
data.append(day_data)
except Exception as e:
print(load_path, e)
data = np.asarray(data) # (115, 48, 5)
data = data.reshape(-1, 5) # (5520, 5)
if shuffle: np.random.shuffle(data)
return data
# data preprocessing
data_list = glob.glob('SLDC_Data/*/*/*.csv')
train, test = train_test_split(data_list, test_size=0.1)
train_data = get_data(train)
X_train = train_data[:, :-1] # features
Y_train = train_data[:, -1].reshape(-1, 1) # load, reshaped to 2D
# rescale the data to 0-1 values`
feature_scaler = MinMaxScaler(feature_range=(0, 1))
load_scaler = MinMaxScaler(feature_range=(0, 1)) # try StandardScaler()
X_train = feature_scaler.fit_transform(X_train)
Y_train = load_scaler.fit_transform(Y_train)
X_train.shape, Y_train.shape, X_test.shape, Y_test.shape
def getModel():
model = Sequential()
model.add(Dense(256, input_shape=(4, )))
model.add(Activation('relu'))
model.add(Dense(256))
model.add(Activation('relu'))
model.add(Dense(256))
model.add(Activation('relu'))
model.add(Dense(1, activation='sigmoid'))
model.compile(loss='mse', optimizer='adam')
print(model.summary())
return model
model = getModel()
model.fit(X_train, Y_train, batch_size=32, epochs=50, verbose=1, validation_split=0.1, shuffle=True)
def predict(data):
'''selects a date in data and plots the predicted curve using the trained model'''
try:
x = np.random.randint(len(data)) # to select a date in data data
data_data = get_data([data[x]], shuffle=False)
X_data = data_data[:, :-1] # features
Y_data = data_data[:, -1].reshape(-1, 1) # load, reshaped to 2D
X_data = feature_scaler.fit_transform(X_data)
Y_data = load_scaler.fit_transform(Y_data)
preds = model.predict(X_data)
preds = load_scaler.inverse_transform(preds)
Y_ = load_scaler.inverse_transform(Y_data)
plt.plot(range(len(Y_)), Y_)
plt.plot(range(len(preds)), preds)
plt.legend(['actual', 'pred'])
plt.title(data[x])
plt.show()
except Exception as e:
print('Err in data', e)
# prediction on test data
predict(test)
# prediction on train data
predict(train)
# prediction on train data as a whole
preds = model.predict(X_train)
preds = load_scaler.inverse_transform(preds)
Y_ = load_scaler.inverse_transform(Y_train)
x = np.random.randint(len(preds))
x_max = 50
print(x,x+x_max)
plt.plot(range(x, x+x_max), preds[x:x+x_max])
plt.plot(range(x, x+x_max), Y_[x:x+x_max])
plt.legend(['pred', 'actual'])
plt.show()
```
| github_jupyter |
# Systematic Uncertainties
It is very important to know how to treat systematic uncertainties in the analysis.
You can find instructions on how this is done in the [Practical Statistics for the LHC](https://arxiv.org/pdf/1503.07622.pdf) from page 17 on. However, it is a bit complicated and goes too much into detail.
A guide that I can really recomment is the note [Probability Density Functions for Positive Nuisance Parameters](http://www.physics.ucla.edu/~cousins/stats/cousins_lognormal_prior.pd) by Bob Cousins (one of the statistics gurus from the pre-LHC era). In this tutorial, we will implement the procedure that he recommends and that is still used widely in CMS.
Again we start with imports:
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
```
Since I don't want to repeat the big statistics functions from the previous tutorial in this notebook, I put them into a file `statistics_tools.py` from which we will import them:
```
from geeksw.stats.statistics_tools import asymptotic_discovery_fit, make_nll_func
```
We create the same example datacard as last time:
```
datacard = pd.DataFrame(dict(WWW=[27.90, 16.07, 0, 0, 0],
WWZ=[0.91, 0.91, 8.57, 0, 0],
WZZ=[0.18, 0.01, 0.34, 1.08, 0.01],
ZZZ=[0, 0, 0.07, 0.25, 0.41],
VH_WWW=[17.62, 8.47, 0, 0, 0],
VH_WWZ=[0, 0, 5.6+0.09+0.17, 0, 0],
VH_WZZ=[0.0, 0, 0, 0, 0],
VH_ZZZ=[0.0, 0, 0, 0, 0],
B=[447.21, 94.99, 15.76, 0.55, 0.06]),
index=["SSplusJets", "3leptons", "4leptons", "5leptons", "6leptons"])
datacard.index.name = "bin"
datacard
```
We need to import the `scipy.stats` package to get easy access to some standard probability distributions.
```
import scipy.stats
```
Remember that the likelihood fits we performed so far are done in a purely frequentist approach by answering the question "what is the likelihood to obtain the observations given a model?".
For the treatment of nuisance parameters, it is common to take a Bayesian approach by using the posterior probability distributions of nuisance parameters after we estimate it somehow with the information we have on them already.
For example, we might have measured some scale factors that we applied to the weights of the Monte Carlo events. Assume that there is an uncertainty on this scale of maybe 5 %. That means we have to add a nuisance parameter to scale the Monte Carlo which we expect somehow between 0.95 and 1.05. We could encode this in a Gaussian probability distribution with mean 1.0 and width 0.05:
```
x = np.linspace(0.8, 1.2)
plt.plot(x, scipy.stats.norm.pdf(x, scale=0.05, loc=1.), label="Gaussian pdf")
plt.legend()
plt.show()
```
When we build our negative log-likelihood function, we therefore just have to add one "penalty" term corresponding to this Gaussian. Let's create a function to create this term for a given relative uncertainty estimate and nuisance parameter value:
```
def normal_penalty(x, relative_uncertainty):
sigma = relative_uncertainty
return -scipy.stats.norm.logpdf(x, scale=sigma, loc=1.)
```
As expected, the penalty gets higher the further away we get from the expected value 1.0:
```
relative_uncertainty = 0.05
x = np.linspace(0.8, 1.2, 200)
plt.plot(x, normal_penalty(x, relative_uncertainty), label="Gaussian nll")
plt.legend()
plt.show()
```
There is just one problem: when we talk about scale corrections, negative scales are unphysical and the penalty should be theoretically infinite. However, a Gaussian also has support for negative values, which gets clear when we consider a large scale uncertainty of let's say 50 % (note the change in the x-axis scale):
```
relative_uncertainty = 0.5
x = np.linspace(-0.5, 1.5, 200)
plt.plot(x, normal_penalty(x, relative_uncertainty), label="Gaussian nll")
plt.legend()
plt.show()
```
That should not be! A Guassian is therefore not the right distribution for scale uncertainties. However, most uncertainties in practice **are** scale uncertainties. That's why we need to look at different distributions.
The natural choise for scale uncertainties is the [log-normal distribution](https://en.wikipedia.org/wiki/Log-normal_distribution#Statistical_Inference), which is only defined for positive values. I will not go in details about the motivation here, because it is already perfectly summarized in Cousins note. Essentially, we expect the probability for $x$ and $1/x$ to be the same instead of for $1+x$ and $1-x$ when we talk about scales, which naturally leads to the log-normal distribution.
In Cousins note, the besti estimate for the scale is associated with the **median** of the distribution. For us, that means the median is assumend to be 1, since we assume to have already applied the scale in Monte Carlo. The median of the log-normal distribution is $\log(\mu)$, hence we set $\mu$ to zero.
Note that sometimes people associate the **mean** or the **mode** with the best estimate, which is why I implemeted this in our `lognormal_penalty` function as well. The default is the median, though.
```
def lognormal_penalty(x, relative_uncertainty, associate_best_estimate_with='median'):
kappa = relative_uncertainty+1.
sigma = np.log(kappa)
if associate_best_estimate_with == 'median':
mu = 0
if associate_best_estimate_with == 'mean':
mu = -sigma**2./2
if associate_best_estimate_with == 'mode':
mu = relative_uncertainty**2.
return -scipy.stats.lognorm.logpdf(x, sigma, loc=mu)
```
Great, so now we have a negative log-likelihood penalty that converges towards infinity at zero from the right, just as it should behave.
```
relative_uncertainty = 0.5
x = np.linspace(-0.2, 1.5, 200)
plt.plot(x, lognormal_penalty(x, relative_uncertainty), label="Gaussian nll")
plt.legend()
plt.show()
```
Finally, Cousins note mentions the [gamma distribution](https://en.wikipedia.org/wiki/Gamma_distribution) which has to be used for uncertainties that needs to be for Monte Carlo statisitical uncertainties or uncertainties associate to extrapolations from control regions (we might go into these kind of extrapolations later).
Let's implement this penalty as described in the note as well:
```
def gamma_penalty(x, relative_uncertainty):
a = 1./relative_uncertainty**2
b = 1./relative_uncertainty**2 + 1
return -scipy.stats.gamma.logpdf(a*x, b)
```
You can also find all of these three penalty functions in the `statistics_tools.py` file.
Finally, let's compare the three possible distribution choices for scale uncertainties.
```
relative_uncertainty = 0.3
x = np.linspace(0.5, 1.5, 200)
plt.plot(x, normal_penalty(x, relative_uncertainty), label="Gaussian nll")
plt.plot(x, lognormal_penalty(x, relative_uncertainty), label="log-normal nll")
plt.plot(x, gamma_penalty(x, relative_uncertainty), label="gamma nll")
plt.legend()
plt.show()
```
Notive that the `gamma_penalty` is shifted compared to the others. If you read the Cousins note carefully, you might remember that there is an issue with the normalization of the gamma distribution in this application. This reflects in a constant shift in the negative log-likelihood, which is not relevant if we want to find the minimum.
Another thing to keep in mind is that all three distributions converge towards a Gaussian for small relative uncertainties, as illustrated here with a 2 % uncertainty:
```
relative_uncertainty = 0.02
x = np.linspace(0.9, 1.1, 200)
plt.plot(x, normal_penalty(x, relative_uncertainty), label="Gaussian nll")
plt.plot(x, lognormal_penalty(x, relative_uncertainty), label="log-normal nll")
plt.plot(x, gamma_penalty(x, relative_uncertainty), label="gamma nll")
plt.legend()
plt.show()
```
## Using Nuisance Parameters in our Fitting Tools
Let's see how nuisance parameters can be used with our negative log-likelihood builder and asymptotic fit tool.
Here is just a reminder how to use it without nuisance parameters, with some random signal definitions:
```
signal_strengths = dict(mu_WWW="WWW", mu_WWZ="WWZ", mu_WZZ="WZZ", mu_ZZZ="ZZZ")
fit_result = asymptotic_discovery_fit(datacard, signal_strengths)
fit_result.df_mu
fit_result.df_nuisance
```
As expected, the nuisance parameter data frame is empty.
If you want to add nuisance parameters to the fit, you have to pass the function a list of functions corresponding to the different nuisance parameters. Yes, a list of functions can be a thing in python.
As an example, let's try out some 30 % scale uncertainty $\beta$ on the overall background.
The nuisance parameter function must have a very specific interface:
* it should take two arguments: the nuisance parameter value and the datacard
* it should modify the datacard according to the nuisance parameter (in this example just scale the background category `B` by $\beta$)
* it should return two things: the modified data frame and the penalty term to add to the negative log-lilkelihood
```
def overall_background_nuisance(beta, datacard):
datacard[["B"]] *= beta
return datacard, lognormal_penalty(beta, 0.3)
```
We now put this single nuisance parameter function in a list `nuisances` and pass this to the `asymptotic_discovery_fit` function. You will see the returned nuisances data frame is no longer empty!
```
nuisances = [overall_background_nuisance]
fit_result = asymptotic_discovery_fit(datacard, signal_strengths, nuisances)
fit_result.df_mu
fit_result.df_nuisance
```
That worked great! As expected, adding an overall 30 % uncertainty on the background changed the expected significance quite a bit, especially for the *WWW* analysis because it has a lot of background.
## Conclusion
You know now how to encorporate systematic uncertainties in your analysis correctly. The only issue that we didn't discuss is asymmetric uncertainties, where the uncertainty in one direction is higher than in the other direction. This is done quite a lot in CMS by interpolating different distributions functions on each side, which would be a bit too much detail for this project.
You learned how to implement systematic uncertainties in our fitting tool: this is done in a very general way by defining a function for each nuisance parameter that morphs the datacard and returns a negative log-likelihood penalty.
**What you could do if you like a good challenge (might take too long for this project):** can you consider the statisitical uncertainty from Monte Carlo as a systematic uncertainty in your four lepton analsyis? We discussed how to get these uncertainties in an earlier exercice. Using them as systematics will not be easy, because you need to add one nuisance parameter per bin, as they are uncorrelated!
| github_jupyter |
# Audio Similarity Search
In this example we will be going over the code required to perform audio similarity searches. This example uses a the PANNs model to extract audio features that are then used with Milvus to build a system that can perform the searches.
A deployable version of a reverse audio search can be found in this directory.
## Data
This example uses the TUT Acoustic scenes 2017 Evaluation dataset, which contains 1622 10-second audio clips that fall within 15 categories: Bus, Cafe,
Car, City center, Forest path, Grocery store, Home, Lakeside beach, Library, Metro station, Office, Residential area, Train, Tram, and Urban park.
Dataset size: ~ 4.29 GB.
Directory Structure:
The file loader used in this example requires that all the data be in .wav format due to librosa limitations. The way that files are read also limits the structure to a folder with all the data points.
## Requirements
| Packages | Servers |
|- | - |
| pymilvus | milvus-2.0 |
| redis | redis |
| librosa |
| ipython |
| numpy |
| panns_inference |
We have included a requirements-v2.txt file in order to easily satisfy the required packages.
```
%pip install -r requirements.txt
```
### Starting Milvus Server
This demo uses Milvus 2.0, please refer to the [Install Milvus](https://milvus.io/docs/v2.0.0/install_standalone-docker.md) guide to learn how to use this docker-compose setup. For this example we wont be mapping any local volumes.
### Starting Redis Server
We are using Redis as a metadata storage service for this example. Code can easily be modified to use a python dictionary, but that usually does not work in any use case outside of quick examples. We need a metadata storage service in order to be able to be able to map between embeddings and their corresponding audio clips.
```
! docker run --name redis -d -p 6379:6379 redis
```
## Code Overview
### Connecting to Servers
We first start off by connecting to the servers. In this case the docker containers are running on localhost and the ports are the default ports.
```
#Connectings to Milvus and Redis
import redis
from pymilvus import connections, DataType, FieldSchema, CollectionSchema, Collection, utility
connections.connect(host = '127.0.0.1', port = 19530)
red = redis.Redis(host = '127.0.0.1', port=6379, db=0)
```
### Building Collection and Setting Index
The next step involves creating a collection. A collection in Milvus is similar to a table in a relational database, and is used for storing all the vectors. To create a collection, we first must select a name, the dimension of the vectors being stored within, the index_file_size, and metric_type. The index_file_size corresponds to how large each data segmet will be within the collection. More information on this can be found here. The metric_type is the distance formula being used to calculate similarity. In this example we are using the Euclidean distance.
```
#Creating collection
import time
red.flushdb()
time.sleep(.1)
collection_name = "audio_test_collection"
if utility.has_collection(collection_name):
print("Dropping existing collection...")
collection = Collection(name=collection_name)
collection.drop()
#if not utility.has_collection(collection_name):
field1 = FieldSchema(name="id", dtype=DataType.INT64, descrition="int64", is_primary=True,auto_id=True)
field2 = FieldSchema(name="embedding", dtype=DataType.FLOAT_VECTOR, descrition="float vector", dim=2048, is_primary=False)
schema = CollectionSchema(fields=[ field1,field2], description="collection description")
collection = Collection(name=collection_name, schema=schema)
print("Created new collection with name: " + collection_name)
```
After creating the collection we want to assign it an index type. This can be done before or after inserting the data. When done before, indexes will be made as data comes in and fills the data segments. In this example we are using IVF_SQ8 which requires the 'nlist' parameter. Each index types carries its own parameters. More info about this param can be found [here](https://milvus.io/docs/v1.0.0/index.md#CPU).
```
#Indexing collection
if utility.has_collection(collection_name):
collection = Collection(name = collection_name)
default_index = {"index_type": "IVF_SQ8", "metric_type": "L2", "params": {"nlist": 16384}}
status = collection.create_index(field_name = "embedding", index_params = default_index)
if not status.code:
print("Successfully create index in collection:{} with param:{}".format(collection_name, default_index))
```
### Processing and Storing Audio Files
In order to store the audio tracks in Milvus, we must first get the embeddings. To do this, we start by loading the audio file using Librosa. Once we have the audio clip loaded we can pass it to the PANN model. In this case we are using the panns_inference library to simplfy the importing and processing. Once we recieve the embedding we can push it into Milvus and store each uniqueID and filepath combo into redis. We do this so that we can later access the audio file when displaying the results.
```
import os
import librosa
import gdown
import zipfile
import numpy as np
from panns_inference import SoundEventDetection, labels, AudioTagging
data_dir = './example_audio'
at = AudioTagging(checkpoint_path=None, device='cpu')
def download_audio_data():
url = 'https://drive.google.com/uc?id=1bKu21JWBfcZBuEuzFEvPoAX6PmRrgnUp'
gdown.download(url)
with zipfile.ZipFile('example_audio.zip', 'r') as zip_ref:
zip_ref.extractall(data_dir)
def embed_and_save(path, at):
audio, _ = librosa.core.load(path, sr=32000, mono=True)
audio = audio[None, :]
try:
_, embedding = at.inference(audio)
embedding = embedding/np.linalg.norm(embedding)
embedding = embedding.tolist()[0]
mr = collection.insert([[embedding]])
ids = mr.primary_keys
collection.load()
red.set(str(ids[0]), path)
except Exception as e:
print("failed: " + path + "; error {}".format(e))
print("Starting Insert")
download_audio_data()
for subdir, dirs, files in os.walk(data_dir):
for file in files:
path = os.path.join(subdir, file)
embed_and_save(path, at)
print("Insert Done")
```
### Searching
In this example we perform a search on a few randomly selected audio clips. In order to perform the search we must first apply the same processing that was done on the original audio clips. This will result in us having a set of embeddings.
```
def get_embed(paths, at):
embedding_list = []
for x in paths:
audio, _ = librosa.core.load(x, sr=32000, mono=True)
audio = audio[None, :]
try:
_, embedding = at.inference(audio)
embedding = embedding/np.linalg.norm(embedding)
embedding_list.append(embedding)
except:
print("Embedding Failed: " + x)
return np.array(embedding_list, dtype=np.float32).squeeze()
random_ids = [int(red.randomkey()) for x in range(2)]
search_clips = [x.decode("utf-8") for x in red.mget(random_ids)]
embeddings = get_embed(search_clips, at)
print(embeddings.shape)
```
We can then take these embeddings and perform a search. The search requires a few arguments: the name of the collection, the vectors being searched for, how many closest vectors to be returned, and the parameters for the index, in this case nprobe. Once performed this example will return the searched clip and the result clips.
```
import IPython.display as ipd
def show_results(query, results, distances):
print("Query: ")
ipd.display(ipd.Audio(query))
print("Results: ")
for x in range(len(results)):
print("Distance: " + str(distances[x]))
ipd.display(ipd.Audio(results[x]))
print("-"*50)
embeddings_list = embeddings.tolist()
search_params = {"metric_type": "L2", "params": {"nprobe": 16}}
try:
start = time.time()
results = collection.search(embeddings_list, anns_field="embedding", param=search_params, limit=3)
end = time.time() - start
print("Search took a total of: ", end)
for x in range(len(results)):
query_file = search_clips[x]
result_files = [red.get(y.id).decode('utf-8') for y in results[x]]
distances = [y.distance for y in results[x]]
show_results(query_file, result_files, distances)
except Exception as e:
print("Failed to search vectors in Milvus: {}".format(e))
```
## Conclusion
This notebook shows how to search for similar audio clips.
Check out our [demo system](https://zilliz.com/milvus-demos) to try out different solutions.
| github_jupyter |
```
# Copyright 2018 Anar Amirli
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Model Testing
In this document, we test our Random Forest for classifying the ball state (wheter game is stoped or not) and the ball coordinate on the y-axis:
1. We have sufficiently enough training data, therefore we're not going to apply K-fold cross-validation on our model.
```
%matplotlib inline
import pickle
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from collections import Counter
from sklearn.utils import shuffle
from sklearn.metrics import confusion_matrix
from sklearn.metrics import roc_auc_score
from sklearn.metrics import mean_squared_error
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import KFold, train_test_split
from sklearn.svm import SVC
import itertools
# our common utility functions that are used in different notebooks
from models.Utils import *
def describe_dataset(y_val):
heading = 'Pitch index counts'
print(heading + '\n' + '-'*len(heading))
for key, val in sorted(Counter(y_val).items()):
print('{}\t: {}'.format(int(key), val))
```
## Labale and Columns Names
```
all_data = pd.read_csv('data_all_3.csv')
classes = ['Null', 'Game Play']
feature_names = list(all_data.columns[4:-4])
```
## Cross Validation for Classifier
```
def random_forest_cv(clf, X, y, feature_names, classes=None, k=10):
label_ids = sorted(np.unique(y), key=abs)
# shuffle the data
X, y = shuffle(X, y)
# get K folds
skf = KFold(n_splits=k, shuffle=True)
skf.get_n_splits(X, y)
# cumulative confusion matrix and feature importance arrays
conf_mat = np.zeros((len(label_ids), len(label_ids)))
feature_importances = np.zeros(X.shape[1])
# for each fold in KFold
for train_index, test_index in skf.split(X, y):
X_train, X_test = X[train_index], X[test_index]
y_train, y_test = y[train_index], y[test_index]
# fit with current train set
clf = clf.fit(X_train, y_train)
# predict the current test set
y_pred = clf.predict(X_test)
# keep cumulative sum of conf_mat and feature_importances
conf_mat += confusion_matrix(y_test, y_pred, labels=label_ids)
feature_importances += clf.feature_importances_
# normalize importances
feature_importances /= k
if classes is None:
classes = label_ids
# plot
plot_confusion_matrix(conf_mat, classes, 'Prediction', 'Truth', normalize=True)
plot_hbar_nameval(feature_names, feature_importances, 'Feature importances', max_bars=20)
all_pred=0.0
true_pred=0.0
for i in range(len(feature_names)):
for j in range(len(feature_names)):
all_pred+=cm[i][j]
if i==j:
true_pred+=cm[i][j]
true_pred /= all_pred
return true_pred, conf_mat, feature_importances
```
## SMOTE
SMOTE (Synthetic Minority Over-Sampling Technique) is an over-sampling technique that introduces small perturbations to synthetic examples along the direction of existing samples to reduce overfitting. See original paper for detailed explanation of SMOTE.
## SMOTE Implementation
There is a SMOTE implementation in imblearn package for scikit-learn. However, there is not an option to apply SMOTE with arbitrary percentages (SMOTE-100, SMOTE-300, etc.); it simply balances all the classes. And also since SMOTE is not a hard to implement algorithm, we provide our own implementation. Our dataset sizes are not big (order of 10k). Hence, a simple Python implementation would be more than sufficien for our needs. See the original paper for reference algorithm.
```
from sklearn.neighbors import NearestNeighbors
from random import randint
def smote(samples, amount, k=5):
"""
Apply SMOTE algorithm to samples and return a new samples
array with synthetically created samples.
Parameters
----------
samples: (n_samples, n_features) samples array to be sent to
SMOTE algorithm.
amount: Percentage of newly created synthetic samples. (E.g.
amount=100 would create as many synthetic examples
as existing ones).
k: Number of nearest neighbors in SMOTE algorithm.
Returns
-------
out: ((1 + amount/100)*n_samples, n_features) samples array containing
the original and the newly created synthetic examples.
References
----------
http://www.jair.org/media/953/live-953-2037-jair.pdf
"""
samples = np.copy(samples)
n_samples, n_features = samples.shape
# handle amount < 100 case
if amount < 100:
num_samples = int(len(samples)*(amount/100))
np.shuffle(samples)
samples = samples[:num_samples, :]
amount = 100
amount = int(amount/100)
synthetic = np.empty((n_samples*amount, n_features))
# find k nearest neighbors of each point and store it in nnarray
nbrs = NearestNeighbors(n_neighbors=k + 1).fit(samples)
_, nnarray = nbrs.kneighbors(samples)
nnarray = nnarray[:, 1:] # get rid of self-nearest-neighbor.
# create synthetic examples and store them in synthetic.
for i, neighbors in enumerate(nnarray):
for j in range(amount):
chosen = neighbors[randint(0, k - 1)]
diff = samples[chosen] - samples[i]
gap = np.random.rand(n_features)
synthetic[i*amount + j] = samples[i] + gap*diff
out = np.vstack((samples, synthetic))
return out
```
## SMOTE for data
Assigment number of corner segments in our dataset is much fewer than other segments. We use SMOTE algorithm with various synthetic example amounts to generate more examples for this segments.
We can't use nominal features in our current SMOTE implementation
TODO: Implement SMOTE-NC
```
from sklearn.utils import shuffle
def smote_sampling(X, y, pitch_segment):
smote_amount = 100
pitch_segment_mask = y == pitch_segment
penalty_features = X[pitch_segment_mask]
synthetic_penalty = smote(penalty_features, smote_amount)
n_synthetic = len(synthetic_penalty)
# merge synthetic examples with original examples
X_out = np.vstack((X[~pitch_segment_mask], synthetic_penalty))
y_out = np.concatenate((y[~pitch_segment_mask], [pitch_segment]*n_synthetic))
return X_out, y_out
# x_train = train_data.values[:, 0:-4]
# y_train = train_data.values[:, -4]
# X_out, y_out = smote_sampling(x_train, y_train, 1)
# X_out, y_out = smote_sampling(X_out, y_out, 3)
# X_out, y_out = smote_sampling(X_out, y_out, 13)
# X_out, y_out = smote_sampling(X_out, y_out, 15)
#describe_dataset(y_out)
describe_dataset(all_data.values[:, -1])
```
## Classifier Testing for Game&Null events
```
# model
clf = RandomForestClassifier(
n_estimators=128,
criterion='gini',
min_samples_split=10,
max_features='sqrt',
class_weight='balanced_subsample'
)
# from sklearn import linear_model
# clf = linear_model.SGDClassifier(
# loss='hinge',
# class_weight='balanced',
# penalty='l1',
# max_iter=1000,
# learning_rate='optimal'
# )
```
#### *i.Split data*
```
X = all_data.values[:,4:-4]
y = all_data.values[:,-1]
# shuffle the data
X, y = shuffle(X, y)
# scale the data
scaler = StandardScaler()
scaler.fit(X)
X = scaler.transform(X)
def split_train_test(matrix, target, test_proportion):
ratio = matrix.shape[0]//test_proportion
X_train = matrix[ratio:,:]
X_test = matrix[:ratio,:]
y_train = target[ratio:]
y_test = target[:ratio]
return X_train, X_test, y_train, y_test
X_train, X_test, y_train, y_test = split_train_test(X, y, 4)
```
#### *ii. Train and test model*
```
# fit with current train set
clf = clf.fit(X_train, y_train)
# predict the current test set
y_pred = clf.predict(X_test)
# label id
label_ids = [0,1]
# conf_mat and feature_importances
conf_mat = confusion_matrix(y_test, y_pred, labels=label_ids)
feature_importances = clf.feature_importances_
```
#### *iii. Result*
```
# plot
plot_confusion_matrix(conf_mat, classes, 'Prediction', 'Truth', normalize=True)
plot_hbar_nameval(feature_names, feature_importances, 'Feature importances', max_bars=20)
```
Overall accuracy: 86%
```
with open('game_vs_null_clf.pkl', 'wb') as f:
f.write(pickle.dumps(clf))
```
| github_jupyter |
# [Module 1.1] 로컬 스크래치 훈련 (SageMaker 사용 안함)
본 워크샵의 모든 노트북은 **<font color="red">conda_tensorflow2_p36</font>** 를 사용합니다.
본 노트북(notebook)은 SageMaker 상에서 (1) Keras, (2) Tensorflow 를 배우는 과정을 하기 위한 스크래치 버전 입니다.
이 노트북은 아래와 같은 작업을 합니다.
- 기본 환경 세팅
- CIFAR-10 데이터 세트 다운로드 및 데이터 확인
- 로컬에서 훈련 (단순하게 python train.py 의 형태 임)
- 관련 변수 저장
## 참고:
- 이 페이지를 보시면 Cifar10 데이터 설명 및 기본 모델 훈련이 있습니다. --> [Train a Keras Sequential Model (TensorFlow 2.0)](https://github.com/daekeun-ml/tensorflow-in-sagemaker-workshop/blob/master/0_Running_TensorFlow_In_SageMaker_tf2.ipynb)
- 메인 깃 리파지토리: [SageMaker Workshop: Tensorflow-Keras 모델을 Amazon SageMaker에서 학습하기](https://github.com/daekeun-ml/tensorflow-in-sagemaker-workshop)
---
# 1. 기본 세팅
사용하는 패키지는 import 시점에 다시 재로딩 합니다.
```
%load_ext autoreload
%autoreload 2
import tensorflow as tf
print("tensorflow version: ", tf.__version__)
```
# 2. Cifar10 데이터 확인하기
- 랜덤하게 9장의 사진 이미지를 보면서, 데이터를 확인 합니다.
- 재 실행하면 계속 이미지가 랜덤하게 선택되어서 보여지게 됩니다.
## 데이터 읽기 단계
- `data/cifar10/train/train-tfrecords` 일고 9 batch_size 로 할당하여, TF dataset를 생성 합니다.
- 1 개의 배치 (9개의 이미지, 레이블 포함) 를 읽음
- 1 개의 배치안에 있는 사진 보여주기
```
import matplotlib.pyplot as plt
from src.utils import _input
batch_size = 9
sample_dir = 'data/cifar10/train'
sample_dataset = _input(1, batch_size, sample_dir, 'train')
images, labels = iter(sample_dataset).next()
print("images shape: ", images.numpy().shape)
images = images.numpy()
for i in range(batch_size):
# define subplot
plt.subplot(330 + 1 + i)
# plot raw pixel data
plt.imshow(images[i])
# show the figure
plt.show()
```
# 3. 노트북 인스턴스 로컬에서 모델 훈련하기 (SageMaker 전혀 사용 안함)
본 스크립트는 모델 학습에 필요한 인자값(arguments)들을 사용합니다. 모델 학습에 필요한 인자값들은 아래와 같습니다.
1. `model_dir` - 로그와 체크 포인트를 저장하는 경로
2. `train, validation, eval` - TFRecord 데이터셋을 저장하는 경로
3. `epochs` - epoch 횟수
아래 명령어로 **<font color='red'>SageMaker 관련 API 호출 없이</font>** 로컬 노트북 인스턴스 환경에서 1 epoch만 학습해 봅니다. 참고로, MacBook Pro(15-inch, 2018) 2.6GHz Core i7 16GB 사양에서 2분 20초~2분 40초 소요됩니다.
## 데이터 경로 지정 및 훈련 파일 실행
```
data_dir = 'data/cifar10'
train_dir = 'data/cifar10/train'
validation_dir = 'data/cifar10/validation'
eval_dir = 'data/cifar10/eval'
%%time
!mkdir -p logs
!python src/cifar10_keras_tf2.py --model_dir ./logs \
--train {train_dir} \
--validation {validation_dir} \
--eval {eval_dir} \
--epochs 1
!rm -rf logs
```
# 4. 변수 저장
- 다음 노트북에서 사용할 변수 저장
```
%store train_dir
%store validation_dir
%store eval_dir
%store data_dir
```
| github_jupyter |
```
#default_exp dashboard_two
#hide
from nbdev.showdoc import *
#export
RED = '\033[31m'
BLUE = '\033[94m'
GREEN = '\033[92m'
BOLD = '\033[1m'
ITALIC = '\033[3m'
RESET = '\033[0m'
style = {'description_width': 'initial'}
import ipywidgets as widgets
from IPython.display import clear_output
from fastai2.vision.all import*
```
# Dashboard_two
> Data tab of GUI
```
#export
def dashboard_two():
"""GUI for Data tab"""
dashboard_two.datas = widgets.ToggleButtons(
options=['PETS', 'CIFAR', 'IMAGENETTE_160', 'IMAGEWOOF_160', 'MNIST_TINY'],
description='Choose',
value=None,
disabled=False,
button_style='info',
tooltips=[''],
style=style
)
display(dashboard_two.datas)
button = widgets.Button(description='Explore', button_style='success')
display(button)
out = widgets.Output()
display(out)
def on_button_explore(b):
with out:
clear_output()
ds_choice()
button.on_click(on_button_explore)
show_doc(dashboard_two, title_level=3)
#export
def ds_choice():
"""Helper for current fastai datasets: PETS, CIFAR, IMAGENETTE_160, IMAGEWOOF_160 and MNIST_TINY"""
if dashboard_two.datas.value == 'PETS':
ds_choice.source = untar_data(URLs.DOGS)
elif dashboard_two.datas.value == 'CIFAR':
ds_choice.source = untar_data(URLs.CIFAR)
elif dashboard_two.datas.value == 'IMAGENETTE_160':
ds_choice.source = untar_data(URLs.IMAGENETTE_160)
elif dashboard_two.datas.value == 'IMAGEWOOF_160':
ds_choice.source = untar_data(URLs.IMAGEWOOF_160)
elif dashboard_two.datas.value == 'MNIST_TINY':
ds_choice.source = untar_data(URLs.MNIST_TINY)
print(BOLD + BLUE + "Dataset: " + RESET + BOLD + RED + str(dashboard_two.datas.value))
plt_classes()
#export
def plt_classes():
"""Helper for plotting classes in folder"""
disp_img_but = widgets.Button(description='View Images?', button_style='success')
Path.BASE_PATH = ds_choice.source
train_source = (ds_choice.source/'train/').ls().items
print(BOLD + BLUE + "Folders: " + RESET + BOLD + RED + str(train_source))
print(BOLD + BLUE + "\n" + "No of classes: " + RESET + BOLD + RED + str(len(train_source)))
num_l = []
class_l = []
for j, name in enumerate(train_source):
fol = (ds_choice.source/name).ls().sorted()
names = str(name)
class_split = names.split('train')
class_l.append(class_split[1])
num_l.append(len(fol))
y_pos = np.arange(len(train_source))
performance = num_l
plt.style.use('seaborn')
plt.bar(y_pos, performance, align='center', alpha=0.5, color=['black', 'red', 'green', 'blue', 'cyan'])
plt.xticks(y_pos, class_l, rotation=90)
plt.ylabel('Images')
plt.title('Images per Class')
plt.show()
display(disp_img_but)
out_img = widgets.Output()
display(out_img)
def on_disp_button(b):
with out_img:
clear_output()
display_images()
disp_img_but.on_click(on_disp_button)
#export
def display_images():
"""Helper for displaying images from folder"""
train_source = (ds_choice.source/'train/').ls().items
for i, name in enumerate(train_source):
fol = (ds_choice.source/name).ls().sorted()
fol_disp = fol[0:5]
filename = fol_disp.items
fol_tensor = [tensor(Image.open(o)) for o in fol_disp]
print(BOLD + BLUE + "Loc: " + RESET + str(name) + " " + BOLD + BLUE + "Number of Images: " + RESET +
BOLD + RED + str(len(fol)))
fig = plt.figure(figsize=(30,13))
columns = 5
rows = 1
ax = []
for i in range(columns*rows):
for i, j in enumerate(fol_tensor):
img = fol_tensor[i] # create subplot and append to ax
ax.append( fig.add_subplot(rows, columns, i+1))
ax[-1].set_title("ax:"+str(filename[i])) # set title
plt.tick_params(bottom="on", left="on")
plt.imshow(img)
plt.xticks([])
plt.show()
#hide
from nbdev.export import notebook2script
notebook2script()
```
| github_jupyter |
# Logistic regression tutorial
```
## Do **not** change this cell, and do **not** import
## any other modules anywhere in the notebook.
import numpy as np
import numpy.random as rn
from scipy import optimize, stats
import scipy.linalg as linalg
import matplotlib.pyplot as plt
%matplotlib inline
```
In this tutorial we're going to cover the basics behind logistic regression. For simplicity we will only consider the binary classification case, in which target variables are $y \in \{0,1\}$.
In logistic regression, the probability of a data point $\boldsymbol x$ being of class 1 is given by
$$p(y = 1 | \boldsymbol x, \boldsymbol\theta) = \sigma (\boldsymbol x^\top \boldsymbol\theta) ~ ,$$
where $\sigma(z) = 1/(1+\exp(-z))$ is the _sigmoid_ function.
Combining this with a Bernoulli likelihood and summing over all datapoints $\{\boldsymbol x_i, y_i\}_{i=1}^N$ we end up with a negative log-likelihood function that looks like this:
$$-\log p(\boldsymbol y|\boldsymbol X, \boldsymbol\theta) = -\sum_i\left(y_i \log \sigma(\boldsymbol x_i^\top \boldsymbol\theta) + (1 - y_i) \log ( 1 - \sigma(\boldsymbol x_i^\top \boldsymbol\theta))\right)$$
You will see this expression in many other classification problems, especially in deep learning, where it's known as the _cross-entropy loss_.
Your goal in this tutorial is to learn how to perform inference over the parameters $\boldsymbol\theta$ in logistic regression, including point estimates $\boldsymbol\theta_{\mathrm{ML}}$ and $\boldsymbol\theta_{\mathrm{MAP}}$ and approximations to the posterior $p(\boldsymbol\theta | \boldsymbol X, \boldsymbol y)$.
Let's do it.
## Maximum likelihood estimate
Let's start easy. First, let's generate a toy 1D binary dataset with two paramaters:
* A **jitter** parameter that controls how noisy the data are; and
* An **offset** parameter that controls the separation between the two classes.
```
# Data generation parameters
N = 50
D = 2
jitter = 0.7
offset = 1.2
# Generate the data
x = np.vstack([rn.normal(0, jitter, (N//2,1)), rn.normal(offset, jitter, (N//2,1))])
y = np.vstack([np.zeros((N//2, 1)), np.ones((N//2, 1))])
x_test = np.linspace(-2, offset + 2).reshape(-1,1)
# Make the augmented data matrix by adding a column of ones
x = np.hstack([np.ones((N,1)), x])
x_test = np.hstack([np.ones((N,1)), x_test])
```
Now on to the regression. First, let's code up the logistic log-likelihood as a separate function. This will come in handy.
**Task 1**
* Write a function to calculate the log-likelihood of a dataset given a value of $\boldsymbol\theta$.
```
## EDIT THIS FUNCTION
def log_likelihood(X, y, theta):
# X: N x D matrix of training inputs
# y: N x 1 vector of training targets/observations
# theta: parameters (D x 1)
# returns: log likelihood, scalar
theta = theta.reshape(-1,1)
sigm = 1/(1+ np.exp(- X @ theta))
B = y * np.log(sigm) + (1-y) * (np.log(1-sigm))
L = np.sum(B) ## <-- EDIT THIS LINE
return L
```
Now it's time to optimize it to fit the maximum likelihood parameter,
$$\boldsymbol\theta_{\mathrm{ML}} = \mathrm{arg}_\theta \max p(\boldsymbol y | \boldsymbol X, \boldsymbol\theta)$$
For linear regression, the likelihood function had a closed-form minimum, which made our lives easy. Alas, that is not the case for logistic regression. We will have to resort to _numerical optimization_.
In the lectures you saw how to derive the gradient and all that jazz. For this tutorial you can do it that way, or any other way you want. The optimization is convex, so this should be easy peasy.
**Task 2**
* Write a function to optimize the log-likelihood function you've written above an obtain $\boldsymbol\theta_{\mathrm{ML}}$. Use any optimizer of your choice.
```
## EDIT THIS FUNCTION
def max_lik_estimate(X, y):
# X: N x D matrix of training inputs
# y: N x 1 vector of training targets/observations
# returns: maximum likelihood parameters (D x 1)
N, D = X.shape
theta_ml = optimize.minimize(lambda theta: -log_likelihood(X, y, theta) ,np.zeros(D), method ='BFGS') ## <-- EDIT THIS LINE
theta_ml = (theta_ml.x).reshape(-1,1)
return theta_ml
```
**Task 3**
* Write a predict function to evaluate your estimate.
```
## EDIT THIS FUNCTION
def predict(X, theta):
# Xtest: K x D matrix of test inputs
# theta: D x 1 vector of parameters
# returns: prediction of f(Xtest); K x 1 vector
prediction = 1/(1+ np.exp(- X @ theta)) ## <-- EDIT THIS LINE
K, D=X.shape
for i in range(K):
if (prediction[i] >= 0.5):
prediction[i]=1
else:
prediction[i]=0
return prediction
```
With this we're in a good position to fit a logistic regression to our toy dataset and start visualising the results. Have a go.
1. Use the function you wrote above to estimate $\boldsymbol\theta_{\mathrm{ML}}$ on the toy dataset.
2. Visualize the results, including:
1. The data $x$ and target labels $y$.
2. The labels predicted by the model.
3. The probability assigned by the model, $\sigma(x\theta)$ as a function of $x$.
```
## ADD CODE HERE
# Fit and plot the logistic regression
plt.figure(figsize=(10,8))
theta_estim = max_lik_estimate(x, y)
plt.plot(x[:,1],y,'.', label='data vs target label');
ypredict= predict(x, theta_estim)
plt.plot(x[:,1], ypredict,'+', label='data vs predict label');
sigm = 1/(1+ np.exp(-x @ theta_estim))
plt.plot(x[:,1], sigm, '.', label='probability');
acc= ((y==predict(x,theta_estim)).mean())
print( "The accuracy of the model is :", acc)
t=-theta_estim[0]/theta_estim[1]
plt.axvline(t, label='Decision boundary');
plt.legend(loc = 'best');
l_fun = lambda th: log_likelihood(x, y, np.array([[0],[th]]))
plt.plot(x[:,1], [l_fun(th) for th in x[:,1]], '.', label='log-likelihood')
plt.legend(loc = 'best');
```
There you go! That should be a nice and easy fit. There are a few things we can start playing with at this point:
* Evaluate the performance of your model: plot the decision boundary, likelihood and accuracy on held-out test sets, etc.
* Write a gradient-based and a non-gradient-based optimizer. Do they arrive at the same result? Which one takes longer? Which one evaluates the likelihood function more times?
(Warning: if the plot looks odd and you get several warnings, it may be that the data is linearly separable and the sigmoid is saturating, leading to `np.log(0)` numerical problems. Add more noise and retry.)
## Bayesian logistic regression
### MAP estimate
Now let's move to Bayesian inference on the parameters $\boldsymbol\theta$. Let's put a prior on them. Because that's what we do. We put priors on things.
More specifically, let's use a Gaussian prior parametrized by a mean $\boldsymbol m$ and a variance $\boldsymbol S$:
$$\boldsymbol\theta \sim \mathcal{N}(\boldsymbol m, \boldsymbol S)$$
Given that $\boldsymbol\theta_{\mathrm{ML}}$ had no analytical solution, it should really come as no surprise that $\boldsymbol\theta_{\mathrm{MAP}}$ doesn't either. That should be no problem for a machine learning expert like you:
**Task 4**
1. Write down the equation for the full unnormalized posterior $p(\boldsymbol\theta | \boldsymbol X, \boldsymbol y) \propto p(\boldsymbol y | \boldsymbol\theta, \boldsymbol X) p(\boldsymbol\theta)$.
2. Write a separate function for it, as we did with the log-likelihood above.
3. Optimize it to find $\boldsymbol\theta_{\mathrm{MAP}}$ and use it to make predictions.
```
## EDIT THIS FUNCTION
def unnormalized_posterior(X,y,theta,m,s):
# X: N x D matrix of training inputs
# y: N x 1 vector of training targets/observations
# theta: parameters (D x 1)
# m : mean (D x1)
# S : covariance (D x D)
# returns: log likelihood, scalar
theta = theta.reshape(-1,1)
inv = np.linalg.inv(s)
logp = (-0.5 *(theta-m).T @ inv @ (theta-m))+log_likelihood(X, y, theta)
return -logp
def map_estimate(X, y, m, S):
# X: N x D matrix of training inputs
# y: N x 1 vector of training targets/observations
# m: D x 1 prior mean of parameters
# S: D x D prior covariance of parameters
# returns: maximum a posteriori parameters (D x 1)
N, D = X.shape
theta_map = optimize.minimize(lambda theta: unnormalized_posterior(X,y,theta,m,S) ,np.zeros(D), method ='BFGS') ## <-- EDIT THIS LINE
theta_map = (theta_map.x).reshape(-1,1)
return theta_map
```
Now you can perform a similar model evaluation as you did before. How does your prior influence the MAP estimate and the model's performance?
```
## ADD CODE HERE
# Fit and plot the MAP logistic regression estimate
m = np.zeros((D, 1))
S = 5*np.eye(D)
plt.figure(figsize=(10,8))
theta_est = map_estimate(x, y, m, S)
plt.plot(x[:,1],y,'.', label='data vs target label');
ytes= predict(x, theta_est)
plt.plot(x[:,1], ytes,'+', label='data vs predict label');
sig = 1/(1+ np.exp(-x @ theta_est))
plt.plot(x[:,1], sig, '.', label='probability');
ac= ((y==predict(x,theta_est.reshape(-1,1))).mean())
print( "The accuracy of the model is :", ac)
t1=-theta_est[0]/theta_est[1]
plt.axvline(t1, label='Decision boundary');
plt.legend(loc = 'best');
```
### The Laplace approximation
As we have hinted above, in logistic regression the posterior distribution over $\boldsymbol\theta$ doesn't have an analytical solution. This is the first example in the course of _approximate Bayesian inference_: The exact posterior is analytically intractable so that we have to approximate it using one of various techniques. The one we'll use in this part of the tutorial is called the **Laplace approximation**.
In brief, **the Laplace approximation is a Gaussian centered at the peak of the pdf of interest with the same curvature**. Let's make this a bit more rigorous below.
Let's say we have a probability distribution $p(\boldsymbol z)$ we want to approximate. The distribution $p(\boldsymbol z)$ is of the form
$$p(\boldsymbol z) = \frac{1}{Z} \tilde{p}(\boldsymbol z) ~ ,$$
where $\tilde{p}(\boldsymbol z)$ is an unnormalized distribution that we can evaluate easily, but $Z$ is unknown. Formally, the Laplace approximation results from a second-order Taylor expansion of $\log \tilde{p}(\boldsymbol z)$ around $\boldsymbol z_0$:
$$\log \tilde{p}(\boldsymbol z) \approx \log \tilde{p}(\boldsymbol z_0) + \frac{d}{d\boldsymbol z}\log \tilde{p}(\boldsymbol z)\Big|_{\boldsymbol z=\boldsymbol z_0}(\boldsymbol z -\boldsymbol z_0) + \frac{1}{2}(\boldsymbol z-\boldsymbol z_0)^\top\frac{d^2}{d\boldsymbol z^2} \log \tilde{p}(\boldsymbol z)\Big|_{\boldsymbol z=\boldsymbol z_0}(\boldsymbol z-\boldsymbol z_0)$$
Now let's evaluate this expression at the mode of $p(\boldsymbol z)$ – which is the same as the mode of $\tilde{p}(\boldsymbol z)$. We define the mode $\boldsymbol z^*$ such that
$$\frac{d}{d\boldsymbol z} \tilde{p}(\boldsymbol z) \Big|_{\boldsymbol z = \boldsymbol z^*} = \boldsymbol 0 ~ .$$
At this point, the $\mathcal{O}(\boldsymbol z)$ term of the expansion vanishes and we are left with
$$\log \tilde{p}(\boldsymbol z) \approx \log \tilde{p}(\boldsymbol z^*) - \frac{1}{2}(\boldsymbol z-\boldsymbol z^*)^\top\boldsymbol A(\boldsymbol z-\boldsymbol z^*)$$
Or, equivalently,
$$\tilde{p}(\boldsymbol z) \approx \tilde{p}(\boldsymbol z^*) \exp\big(-\tfrac{1}{2}(\boldsymbol z - \boldsymbol z^*)^\top\boldsymbol A(\boldsymbol z - \boldsymbol z^*)\big) ~ ,$$
where
$$\boldsymbol A = - \frac{d^2}{d\boldsymbol z^2} \log \tilde{p}(\boldsymbol z)\Big|_{\boldsymbol z=\boldsymbol z^*} ~ .$$
And now this distribution we know how to normalize, because it's one of those Gaussians we know and love. By inspection, we can identify the mean and the covariance, and write down the Laplace approximation of $p(\boldsymbol z)$ as
$$q(\boldsymbol z) = \mathcal{N}(\boldsymbol z | \boldsymbol z^*, \boldsymbol A^{-1})$$
As an example, let's use the unnormalized distribution $\tilde{p}(z) = x e^{-x/2}$. When normalized properly, this is in fact the $\chi^2$ distribution with $k=4$ degrees of freedom. Have a go yourself:
1. Plot $p(z)$.
2. Take the first derivative of $\tilde{p}(z)$ (or the first derivative of its log), and find its maximum $z^*$ analytically.
3. In the same plot, draw a vertical line at $z = z^*$ to verify you got the right answer.
4. Take the second derivative of $\log \tilde{p}(z)$ and evaluate it at $z^*$.
5. Plot the corresponding Gaussian $q(z)$ and verify the approximation looks reasonable.
**Task 5**
* Write a function that evaluates the Laplace approximation $q(z)$.
```
## EDIT THIS FUNCTION
def laplace_q(z):
# z: double array of size (T,)
# returns: array with Laplace approximation q evaluated
# at all points in z
q = stats.multivariate_normal.pdf(z,2,4)
return q
## ADD CODE HERE
# Find the Laplace approximation of x*exp(-x/2) with pen and paper and then plot it.
z = np.linspace(0,10)
p = stats.chi2.pdf(z, 4)
plt.plot(z,p, 'r', label= 'chi square distribution')
plt.axvline(x=2 , label='z=z*')
plt.plot(z, laplace_q(z), 'g', label='Laplace approximation');
plt.legend(loc='best');
```
### Bayesian logistic regression (for real this time)
Now we have obtained the mode (peak) of the posterior through the MAP estimate above, it's time to go all the way and calculate the posterior over $\boldsymbol\theta$. However, as we mentioned above the posterior doesn't have an analytical form, so we'll use – you guessed it – the Laplace transform.
**Task 6**
* Write a function, based on your previous code, that will calculate the Laplace approximation $q(\boldsymbol\theta)$ of the true posterior $p(\boldsymbol\theta | \boldsymbol X, \boldsymbol y)$ and return the mean and variance of $q$.
To visualize the behavior and the diversity of $q$, draw a number $j = 1, ..., J$ of samples $\boldsymbol\theta_j \sim q(\boldsymbol\theta)$. For each sample, plot its predicted class probabilities $\sigma(x \boldsymbol\theta_j)$.
_Hint_: the extension of the Laplace approximation to multivariate distributions is straightforward, and in this case the variance of the Gaussian is the Hessian of the negative log likelihood $\boldsymbol A = - \nabla_\theta \nabla_\theta \log p(\boldsymbol\theta | \boldsymbol X, \boldsymbol y)$.
```
## EDIT THIS FUNCTION
def unnormalized_posterior(X,y,theta,m,s):
# X: N x D matrix of training inputs
# y: N x 1 vector of training targets/observations
# theta: parameters (D x 1)
# m : mean (D x1)
# S : covariance (D x D)
# returns: log likelihood, scalar
theta = theta.reshape(-1,1)
inv = np.linalg.inv(s)
logp = (-0.5 *(theta-m).T @ inv @ (theta-m))+log_likelihood(X, y, theta)
return -logp
def get_posterior(X, y, m, S):
# X: N x D matrix of training inputs
# y: N x 1 vector of training targets/observations
# m: D x 1 prior mean of parameters
# S: D x D prior covariance of parameters
# returns: maximum a posteriori parameters (D x 1)
# covariance of Laplace approximation (D x D)
N, D = X.shape
opt= optimize.minimize(lambda theta: unnormalized_posterior(X,y,theta,m,S) ,np.zeros(D), method ='BFGS') ## <-- EDIT THIS LINE
mu_post = opt.x ## <-- EDIT THESE LINES
S_post = opt.hess_inv
return mu_post, S_post
## ADD CODE HERE
# Calculate the Laplace approximation of the posterior for theta,
# draw a few samples and plot the corresponding likelihood functions
# for each one.
m = np.zeros((D, 1))
S = 5*np.eye(D)
nb_samples = 5
theta_map, S_post = get_posterior(x, y, m, S)
plt.scatter(x[:,1], y)
for i in range(nb_samples):
thet=np.random.multivariate_normal(theta_map, S_post)
sigmo = 1/(1+ np.exp(- x @ thet))
plt.plot(x[:,1], sigmo, '.') ## <--EDIT THIS LINE
plt.show()
```
## Comparing posterior approximations
The Laplace approximation is part of a family of methods known as _deterministic approximate inference_. In addition, there's another set of methods known as _stochastic approximate inference_ which, as you can guess includes most of the sampling techniques you have studied.
You must be an expert in sampling by now. Let's actually go and check whether this Laplace approximation we just made is legit.
* What sampling methods do you know to sample from an unnormalized distribution?
For example, let's try the Metropolis algorithm.
1. Write a proposal function to move in $\boldsymbol\theta$-space.
2. Write a function to accept or reject new proposals based on the Metropolis criterion.
3. Write a loop and run the Markov chain for a few thousand iterations.
4. Check that the sampling worked: did the Markov chain mix properly? What's the acceptance rate? How does it depend on the proposal function?
**Task 7**
* Write a function to sample from the true posterior $p(\boldsymbol\theta | \boldsymbol X, \boldsymbol y)$.
```
## EDIT THIS FUNCTION
def posterior_sample(X, y, m, S, nb_iter):
# X: N x D matrix of training inputs
# y: N x 1 vector of training targets/observations
# m: D x 1 prior mean of parameters
# S: D x D prior covariance of parameters
# returns: nb_iter x D matrix of posterior samples
D = X.shape[1]
sigma=np.identity(D)
theta0=np.array([0,0])
samples = np.zeros((nb_iter,D))
m = m.ravel()
for i in range(nb_iter):
theta1= np.random.multivariate_normal(theta0, sigma)
q1=stats.multivariate_normal.pdf(theta0,theta1, sigma)
mu1=1/(1+ np.exp(- X @ theta1))
p1=(np.exp(sum(np.log(stats.bernoulli.pmf(y[n],mu1[n])) for n in range(N))) * stats.multivariate_normal.pdf(theta1, m, S))
q0=stats.multivariate_normal.pdf(theta1,theta0, sigma)
mu0=1/(1+ np.exp(- X @ theta0))
p0=(np.exp(sum(np.log(stats.bernoulli.pmf(y[n],mu0[n])) for n in range(N))) * stats.multivariate_normal.pdf(theta0, m, S))
u=np.random.uniform (0,1)
if (q1 * p1 / (q0 * p0)>= u):
samples[i,:] = theta1
theta0=theta1
else:
samples[i,:]=theta0
return samples
```
Finally, let's plot the results and see if both inference methods arrive at roughly the same posterior.
In the same axis, plot
* The histogram pdf of the MCMC samples (you may want to look at the `density` option in `plt.hist`); and
* The Laplace posterior.
Make one plot for the intercept ($\theta_0$) and one for the slope ($\theta_1$). What do they look like? Do they match? What kinds of posteriors do you think the Laplace approximation will be good or bad at approximating?
```
## ADD CODE HERE
# Plot a histogram of the MCMC posterior samples and the
# analytical expression for the Laplace posterior. If
# everything went well, the peaks should coincide and
# their widths should be comparable.
nb_iter = 10000
samples = posterior_sample(x, y, m, S, nb_iter)
sample = posterior_sample(x, y, m, S, nb_iter)
plt.figure(figsize=(10,8))
plt.hist(sample[:,0],20,normed=True);
plt.hist(sample[:,1],20,normed=True);
```
Et violà! Now you're an expert in logistic regression. (Wait, I think that's a big violin. I meant to say: et voilà!)
Now we can visualize the posterior we can play around with the data and the inference parameters:
* Play around with the data generation process. What happens as you increase/decrease $N$ and the jitter parameter?
* What does the joint posterior look like? Make a visualization of the MCMC and Laplace approximations in the $(\theta_0, \theta_1)$ plane.
* What happens if the model is misspecified? Take out the intercept term in the model (i.e., remove the column of ones in $\boldsymbol X$), but set the `offset` in the data generation process to non-zero. What happens to the posterior and its Laplace approximation?
```
plt.figure(figsize=(10,8))
plt.hist(sample[:,0],20,normed=True);
plt.hist(sample[:,1],20,normed=True);
z = np.linspace(-4,4,100)
pz = stats.multivariate_normal.pdf(z,theta_map[0], S_post[0,0])
pz2 = stats.multivariate_normal.pdf(z,theta_map[1], S_post[1,1])
plt.plot(z,pz);
plt.plot(z,pz2);
```
| github_jupyter |
# Lab 04 : Train vanilla neural network -- demo
## Training a one-layer net on MNIST
```
import sys, os
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from random import randint
import utils
```
### randint(a,b) returns a random integer between a and b:
```
idx = randint(5,10) # generate an integer randomly from 5 to 10 included
print(idx)
```
### Download the TRAINING SET (data+labels)
```
from utils import check_mnist_dataset_exists
data_path = check_mnist_dataset_exists()
print(data_path)
train_data = torch.load(data_path+'mnist/train_data.pt')
print(train_data.size())
train_label = torch.load(data_path+'mnist/train_label.pt')
print(train_label.size())
```
### Download the TEST SET (data only)
```
test_data = torch.load(data_path+'mnist/test_data.pt')
print(test_data.size())
```
### Make a one layer net class
```
class one_layer_net(nn.Module):
def __init__(self, input_size, output_size):
super(one_layer_net, self).__init__()
self.linear_layer = nn.Linear(input_size, output_size, bias = False)
def forward(self, x):
x = self.linear_layer(x)
p = F.softmax(x, dim=1)
return p
```
### Build the net
```
net = one_layer_net(784,10)
print(net)
print(net.linear_layer.weight)
print(net.linear_layer.bias)
```
### Take the 6th image of the test set:
```
im = test_data[6]
utils.show(im)
```
### And feed it to the UNTRAINED network:
```
p = net(im.view(1,784))
print(p)
```
### Display visually the confidence scores
```
utils.show_prob_mnist(p)
```
### Train the network (only 5000 iterations) on the train set
```
criterion = nn.NLLLoss()
optimizer = torch.optim.SGD(net.parameters(), lr=0.01)
for iter in range(1,5000):
# choose a random integer between 0 and 59,999
# extract the corresponding picture and label
# and reshape them to fit the network
idx = randint(0, 60000-1)
input = train_data[idx].view(1, 784)
label = train_label[idx].view(1)
# feed the input to the net
input.requires_grad_()
prob = net(input)
# update the weight (all the magic happens here -- we will discuss it later)
log_prob = torch.log(prob)
loss = criterion(log_prob, label)
optimizer.zero_grad()
loss.backward()
optimizer.step()
```
### Take the 6th image of the test set:
```
im = test_data[6]
utils.show(im)
```
### Feed it to the TRAINED net:
```
prob = net(im.view(1,784))
print(prob)
```
### Display visually the confidence scores
```
utils.show_prob_mnist(prob)
```
### Choose image at random from the test set and see how good/bad are the predictions
```
# choose a picture at random
idx = randint(0, 10000-1)
im = test_data[idx]
# display the picture
utils.show(im)
# feed it to the net and display the confidence scores
prob = net(im.view(1,784))
utils.show_prob_mnist(prob)
weights = net.linear_layer.weight[7].view(28, 28)
utils.show(weights.detach())
```
| github_jupyter |
```
def pfg(m,n,k):
i=1
sum = 0
while i<(n+1):
import random
number = random.randint(m, k)
sum = sum + number
i = i + 1
aver = sum/n
total = pow(aver,1/2)
return total
m=int(input('plz input a integer'))
n=int(input('plz input a integer'))
k=int(input('plz input a integer'))
print('求n个随机整数均值的平方根 ', pfg(m,n,k))
def sigma1(m,n,k):
i=1
sum1 = 0
while i<(n+1):
import random
number = random.randint(m, k)
import math
number1 = math.log2(number)
sum1 = sum1 + number1
i = i + 1
return sum1
def sigma2(m,n,k):
i=1
sum2 = 0
while i<(n+1):
import random
number = random.randint(m, k)
import math
number1 = math.log2(number)
number2 = 1/number1
sum2 = sum2 + number2
i = i + 1
return sum2
m=int(input('plz input a integer'))
n=int(input('plz input a integer'))
k=int(input('plz input a integer'))
print('求西格玛log(随机整数)',sigma1(m,n,k))
print('求西格玛1/log(随机整数)',sigma2(m,n,k))
def sigma(n):
i=1
s = 0
import random
number = random.randint(1, 9)
while i<(n+1):
import random
number1 = (pow(10,i)-1)/9
s = s + number1*number
i = i + 1
return s
n=int(input('plz input a integer'))
print('s为 ',sigma(n))
import random, math
def win():
print(
'''
======恭喜你,你赢了=======
."". ."",
| | / /
| | / /
| | / /
| |/ ;-._
} ` _/ / ;
| /` ) / /
| / /_/\_/\
|/ / |
( ' \ '- |
\ `. /
| |
| |
======恭喜你,你赢了=======
'''
)
def lose():
print(
'''
======YOU LOSE=======
.-" "-.
/ \
| |
|, .-. .-. ,|
| )(__/ \__)( |
|/ /\ \|
(@_ (_ ^^ _)
_ ) \_______\__|IIIIII|__/__________________________
(_)@8@8{}<________|-\IIIIII/-|___________________________>
)_/ \ /
(@ `--------`
======YOU LOSE=======
'''
)
def game_over():
print(
'''
======GAME OVER=======
_________
/ ======= \
/ __________\
| ___________ |
| | - | |
| | | |
| |_________| |________________
\=____________/ )
/ """"""""""" \ /
/ ::::::::::::: \ =D-'
(_________________)
======GAME OVER=======
'''
)
def show_team():
print('制作团队:BLCU')
def show_instruction():
print('''请玩家输入一个整数n
在1-1000之间随机取一个整数number
取一个最多可猜测次数,计算机已猜测次数设为0,
一直做如下步骤直到已猜测次数大于最多可猜测次数:
让计算机猜测一次
玩家猜测次数加一
告诉计算机目前猜测次数
如果计算机猜中了:
恭喜猜中
显示数字及猜测次数
退出循环
如果计算机猜测数字大于number:
告诉计算机猜大了
否则:
告诉计算机猜小了
告诉计算机还能猜几次
否则:
显示超过次数,猜测失败''')
def menu():
print('''=====游戏菜单=====
1. 游戏说明
2. 开始游戏
3. 退出游戏
4. 制作团队
=====游戏菜单=====''')
def guess_game():
n = int(input('请输入一个大于0的整数,作为神秘整数的上界,回车结束。'))
max_times = math.ceil(math.log(n, 2))
guess_times = 0
while guess_times <= max_times:
guess = random.randint(1, n)
guess_times += 1
print('一共可以猜', max_times, '次')
print('你已经猜了', guess_times, '次')
if guess == n:
win()
print('神秘数字是:', guess)
print('你比标准次数少', max_times-guess_times, '次')
break
elif guess > n:
print('抱歉,你猜大了')
else:
print('抱歉,你猜小了')
else:
print('神秘数字是:', n)
lose()
# 主函数
def main():
while True:
menu()
choice = int(input('请输入你的选择'))
if choice == 1:
show_instruction()
elif choice == 2:
guess_game()
elif choice == 3:
game_over()
break
else:
show_team()
#主程序
if __name__ == '__main__':
main()
```
| github_jupyter |
```
%matplotlib inline
import pandas as pd
import numpy as np
import modelclass as mc
import modelmanipulation as mp
import modeldiff as md
model = '''
frml <> a = c(-1) + b $
frml <> d1 = x + 3 * e(-1)+ c **2 +a $
frml <> d3 = x + 3 * a(-1)+c **3 $
Frml <> x = 0.5 * c $'''
mmodel = mc.model(model,modelname = 'My first model')
df = pd.DataFrame({'B': [1,1,1,1],'C':[1,2,3,4],'E':[4,4,4,4]},index=[2018,2019,2020,2021])
base = mmodel(df)
md.modeldiff(mmodel)
md.display_all(mmodel,base,2018)
matdir = md.calculate_allmat(mmodel,base,2018)
for l,m in matdir.items():
print(f'Lag:{l} \n {m} \n')
for l,m in matdir.items():
endomat = m.loc[:,[c for c in m.columns if c in mmodel.endogene]]
print(f'Lag:{l} \n {endomat} \n')
fam = '''
frml <> y = c+i $
frml <I> c = pa * y(-1) $
frml <> i = pc * (c-c(-1)) + im$'''
mma = mc.model(fam,modelname = 'Accelerator multiplicator model')
fhma = mp.find_hist_model(fam) # Find the model to generate lagged values of c
mhma = mc.model(fhma)
years = 10
df = pd.DataFrame([[1000,200]]*years,index=range(2018,2018+years),columns=['Y','IM'])
df.loc[:,'PA'] = 0.9
df.loc[:,'PC'] = 0.9
start = mhma(df,silent=True)
base = mma(start,silent=True)
#md.modeldiff(mma)
#print(md.calculate_allmat(mma,base,2018))
md.get_compagnion(mma,mma.lastdf,2018)
def geteigen(mul,acc,years=100,show=False):
'''Function which creates a Samuelson Multiplier accelerator model, runs it and
calculates the eigenvalues for the compaignion matrix in order to evaluate stability'''
fam = f'''
frml <> y = c+i $
frml <I> c = {mul} * y(-1) $
frml <> i = {acc} * (c-c(-1)) + im $'''
mma = mc.model(fam,modelname = 'Accelerator multiplicator model')
df = pd.DataFrame([[1000,200]]*years,index=range(2018,2018+years),columns=['Y','IM'])
start = mc.model(f'c = {mul} * y(-1)')(df,silent=True) # Generate lagged variables for c
base = mma(start,silent=True) # Solve the model
comp = md.get_compagnion(mma,mma.lastdf,2021) # get the compagnion matriox
compeeig=np.linalg.eig(comp) # find the eigenvalues
if show:
_ = mma[['Y','C','i']].plot() # Show the solution
md.eigplot(compeeig[0]) # show the eigenvalues
return
```
Stability
```
geteigen(mul=0.5,acc=0,years=30,show=1)
```
Explosion
```
geteigen(mul=0.9,acc=2,years=30,show=1)
```
Exploding oscillations
```
geteigen(mul=0.6,acc=2,years=30,show=1)
```
Perpetual oscillations
```
geteigen(mul=0.5,acc=2,years=30,show=1)
```
Dampened oscillations
```
geteigen(mul=0.7,acc=1,years=30,show=1)
```
| github_jupyter |
# Initial Investigations
looking thru: https://www.midi.org/forum/279-yamaha-bull-dump-data-format,
http://rnhart.net/articles/bulk-dump.htm
```
from __future__ import print_function, division
import itertools
import re
# numpy imports
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
def hexbyte(x):
return "{:02X}".format(x)
def binbyte(x):
return "{:08b}".format(x)
def tohex(by, sep=" "):
return sep.join(hexbyte(x) for x in by)
def tobin(by, sep=" "):
return sep.join(binbyte(x) for x in by)
def hexline(by):
if len(by) <= 24:
return tohex(by)
else:
return tohex(by[:20]) + "..." + tohex(by[-4:])
def takebyn(by, n=8):
for i in range(0, len(by), n):
yield by[i:i+n]
def itertaken(seq, n):
itr = iter(seq)
try:
while True:
group = []
for _ in range(n):
group.append(next(itr))
yield group
except StopIteration:
if group:
yield group
def printhex(by, n=8, group=1):
grouped = ("".join(hexbyte(x) for x in g) for g in takebyn(by, group))
for x in itertaken(grouped, n):
print(" ".join(x))
with open('data/syxout.txt') as hexdata:
lines = [bytearray.fromhex(line.strip()) for line in hexdata]
```
## Picking at the message structure
```
for line in lines:
print(hexline(line))
lb = lines[0]
ls = lines[-4]
```
### Header
sysex-flag/manufacturer/device/data type header: `F0 43 73 7F 44 06`.
If this is using the same conventions as the PSR-225, then `0A` is for song data and `09` is for the one-touch panel bank
```
tohex(lb[:7])
```
### 'SS' Size bytes.
There appear to be four of them. At the EOS this is '00 01 00 01'
the longest messages have this as '10 00 10 00', the last message before EOS has this as '08 68 08 68'.
It appears to be two repeated
(at least, in the case of the 0A. The 09 data, with only one message, appears to be different again.)
```
tohex(lb[7:11])
```
### three 'ZZ' running total bytes.
At the EOS this is '7F 7F 7F', then immediately followed by the end F7.
```
tohex(lb[11:14])
```
### the 'KK' checksum and end-of-message flag F7.
```
tohex(lb[-2:])
```
This should, and I say *should*, be of a similar format, so...
If this works, sum of SS SS SS to KK inclusive should be 0 (mod 128).
```
print([sum(line[7:-1]) % 128 for line in lines])
```
It worked, except for the two end-of-section messages, which don't have the checksum byte.
### Checking the Numbers
```
def unpackseven(by):
# Ignore the first bit, which is always zero.
# The most signifcant bit comes first
value = 0
for i, b in enumerate(reversed(by)):
value |= b << (i*7)
return value
def unpacksevenhex(hexstring):
return "{:#x}".format(unpackseven(bytearray.fromhex(hexstring)))
unpacksevenhex('10 00')
len(lb[14:-2])
```
0x800, or 2048 bytes of encoded data seems to be the maximum.
the ZZ bytes appear to increment by this amount.
that penultimate message should be 0x468, or 1128, bytes...
```
unpacksevenhex('08 68')
unpackseven(ls[7:9]) == len(ls[14:-2])
len(ls[14:-2])
```
Yep.
## What's all that data?
Instead encoding one byte to two by nybble, with 0F 07 == 7F like in the [PSR-225](http://rnhart.net/articles/bulk-dump.htm), it seems that the 'payload' data uses all seven available bits, which could mean several things.
```
payloads = [line[14:-2] for line in lines[:-3]]
payload = bytearray().join(payloads)
```
From staring at the 'payload' in a hex editor, I have recognised some patterns. I think that every 8th byte is some sort of 'parity' byte or something. Ignoring these bytes you can see the ascii for 'YAMAHA' and 'MTrk' and 'PresetStyle\x00'. So that's a start.
```
len(payload)/8
```
### What Is That Eighth Byte?
```
pays = np.array(payload).reshape((9613,8))
plt.plot(pays[:,7], 'r.')
np.asarray(np.unique(pays[:,7], return_counts=True)).T
tobin(pays[pays[:,7]==65,:][2])
tobin(np.unique(pays[:,7]))
```
### The Solution
If I had to squeeze 8 bits down into seven, how would I do it?
Maybe, we take each of the seven bits of the eighth byte and stick them in front of the previous seven?
```
def reconstitute(by):
blob = by[:7]
for i in range(7):
bitmask = 0b01000000 >> i
blob[i] = blob[i] | ((by[7] & bitmask) << i+1)
return blob
def reconstitute_all(by):
return bytearray().join(reconstitute(g) for g in takebyn(by, 8))
payload8 = reconstitute_all(payload)
```
Note that whenever an F0 occurs it is always followed by F7:
```
map(tohex, re.findall(r'\xF0[\x00-\x7F]*[\x80-\xFF]', payload8))
```
It sure looks like there are some system exclusive messages in here! (Stored in MIDI file format, of course.)
Leafing through the manual (pg 110-111), it seems these are the GM System / volume / tuning / reverb / chorus events
And here are some style meta-events:
```
map(tohex, re.findall(r'\x43\x76\x1A\x04.{2}', payload8))
```
## Figuring Out The File Structure
Assuming the file structure is similar to the [PSR-225's block system](http://rnhart.net/articles/bulk-dump.htm)...
```
len(payload8)
```
It looks like there are 67291 bytes of decoded data instead of the 66766 bytes in the PSR-225.
```
printhex(payload8[:0x15a], 16)
```
I have no idea what the first 0x15A bytes are. Maybe it's a bunch of 16-bit numbers?
```
printhex(payload8[0x15a:0x162])
```
The DGX-505 has 5 UserSongs compared to the PSR-225's three. Each user song has five normal tracks and one chord/accompaniment track (track A).
At the time this dump was gathered, all five songs had data, and used the following tracks as displayed on the panel:
1. `1 _ _ _ _ A`
2. `1 2 3 _ _ _`
3. `1 2 3 4 5 _`
4. `1 _ 3 _ _ _`
5. `1 2 3 4 _ _`
```
track_bitfields = slice(0x15d, 0x162)
print(tobin(payload8[track_bitfields], '\n'))
```
The five bytes from 0x15d to 0x161 appear to correspond to the five usersongs, with the low 5 bits of each representing the active tracks on each (from least to most significant bit, track 1 - 2 - 3 - 4 - 5 - A).
This corresponds to offsets 0x00001-0x00003 on the PSR-225.
Still don't know what `64 02 BC` is, though.
```
printhex(payload8[0x162:0x1f3], 16)
```
Following are some bytes of more unknown stuff which could be 32 bit integers?
```
printhex(payload8[0x1f3:0x22f], 12)
payload8[0x1f3:0x22f].decode('ascii')
```
... followed by five repeats of ascii 'PresetStyle' and `00`. The PSR-225 only has three of these (starting from 0x00007), so this has probably got something to do with each song.
```
beginning_blocks = slice(0x22f,0x24d)
printhex(payload8[beginning_blocks], 6)
```
Immediately following that, we have what appears to be the 30 bytes from 0x22f through 0x24c indicating the beginning blocks, corresponding to 0x0002B-0x0003F on the PSR-225.
Each group of six bytes corresponds to a usersong in order, and each byte corresponds to a track, from 1-5, followed by the *time track* with the tempo, SysEx, etc. (Compare with the PSR-225, which has 6+1 tracks for each of its 3 songs and therefore needs 21 (=0x15) bytes total.)
Blocks are numbered 0x01 to 0x82 and `FF` indicates that the track is not in use.
```
next_blocks = slice(0x24d,0x2cf)
printhex(payload8[next_blocks], 16)
```
Running from 0x24d to 0x2ce is the table of 0x82 (=130) next blocks, corresponding to 0x00040-0x000C1 on the PSR-225.
`00` indicates unused blocks, `FF` indicates final blocks.
```
printhex(payload8[0x2cf:0x2d5])
payload8[0x2cf:0x2d5].decode('ascii')
```
At 0x2cf, we find 'PK0001', which corresponds to the PSR-225's 'PK9801' at 0x000C2, and marks the beginning of the block data at 0x2d5.
```
printhex(payload8[0x106d5:])
payload8[0x106d5:].decode('ascii')
```
The final six bytes, spelling 'PK0001' again from 0x106d5 to 0x106da, mark the end of the block data and correspond to 0x104C8-0x104CD on the PSR-225.
```
(0x106d5-0x2d5)/0x82
```
Like the PSR-225, the blocks are 0x200 = 512 bytes in size.
Block 0x01 begins at offset 0x2d5:
```
printhex(payload8[0x2d5:0x2d5+8])
payload8[0x2d5:0x2d5+8]
```
And we can see the start of an MTrk chunk.
To calculate the start each block we can use a simple formula
```
offset(n) = 0x2d5 + (0x200 * (n-1))
```
or, to simplify,
```
offset(n) = 0xd5 + (0x200 * n)
```
```
def chunk_offset(n):
if 0x01 <= n <= 0x82:
return 0xd5 + (0x200*n)
else:
raise ValueError("Invalid chunk", n)
def block_peek(n, length=4):
offset = chunk_offset(n)
return payload8[offset:offset+length]
def print_block_peek(n):
bits = block_peek(n)
print(u"Block 0x{:02x} at 0x{:05x}: {} - {}".format(n, chunk_offset(n), tohex(bits),
bits.decode('ascii', 'replace')))
track_blocks = [x for x in payload8[beginning_blocks] if x != 0xFF]
for n in track_blocks:
print_block_peek(n)
```
We can see that the blocks marked as the beginning blocks for each track do, indeed, contain the start of the tracks.
```
print_block_peek(0x5e)
```
We can also see that some blocks have data left over from previous recordings.
### The Remainder
There are two regions left unexplained:
```
printhex(payload8[:0x15d], 16)
printhex(payload8[0x162:0x1f3], 16)
```
We've accounted for pretty much everything found in the PSR-225's song dump format, except for the very first byte, offset 0x00000, which is supposed to be a bitfield for the songs in use. If the same pattern is used for the DGX-505, then we should expect to see a `00011111` or `1F` byte somewhere, as all five songs were in use.
The very first byte, offset 0x0, is, in fact, `1F`, so that might just be it (I'll have to delete one of the songs to check).
There's also one feature 'Step Recording' (offset 0x0004-0x0006 on PSR-225) the DGX-505 doesn't have. If the (abscence of) step-recorded tracks was noted directly after the normal tracks, this would account for five `00` bytes directly after the normal tracks at 0x162, and the rest lines up rather nicely as a bunch of 32 bit numbers...
```
def unpackeight(by):
# big-endian.
total = 0
for i, b in enumerate(reversed(by)):
total |= b << (i*8)
return total
def unpackwords(by, n=2):
# probably not the way to do it, but eh.
return [unpackeight(x) for x in takebyn(by, n)]
mystery2 = unpackwords(payload8[0x163:0x1f3], 4)
plt.bar(np.arange(len(mystery2)), mystery2)
```
There's definitely a pattern:
```
print(tobin(payload8[0x15d:0x162], '\n')) #track_bitfields
print('---')
printhex(payload8[0x162:0x167])
print('---')
printhex(payload8[0x167:0x17b],1,4)
print('---')
printhex(payload8[0x17b:0x1f3],6,4)
```
It looks like after the track-active bitfields comes five zero bytes, then five 32-bit numbers that seem to correspond to the five songs, then 30 32-bit numbers that correspond to the tracks on the songs, with values of `00000000` for inactive tracks and other numbers (lengths?) for the active tracks. The five numbers seem to be the largest (longest?) value of the tracks for the corresponding song. Looking through the values, they seem to be roughly the duration of the tracks, in measures.
```
printhex(payload8[beginning_blocks], 6)
```
### The Final Mystery Region
```
printhex(payload8[:0x15d], 16)
```
Let's describe some patterns.
From offset 0x0 to 0x18, there's a bunch of bytes. 0x0 may be the song-usage-bitfield thing.
```
printhex(payload8[:0x19], 16)
```
From 0x19 to 0x11c, we get 320 bytes of some increasing and decreasing patterns
```
printhex(payload8[0x19:0x159], 16)
```
And then four more bytes of whatever. I'm baffled, really.
```
printhex(payload8[0x159:0x15d], 16)
mystery_region = payload8[0x019:0x159]
mystery_numbers = unpackwords(mystery_region)
plt.bar(np.arange(160), mystery_numbers, width=1, linewidth=0)
```
Staring at this, we can see four regions that run from indexes 0-49, 50-99, 100-129 and 130-159. Note that there are 160 numbers here, and 130 blocks in the file system, and 30 tracks. I'm guessing that this has something to do with the blocks just by the numbers here, but I don't know what. Probably some internal file system thing
```
printhex(mystery_numbers[:-30], 10)
print()
printhex(payload8[next_blocks], 10)
```
And that still doesn't explain 0x0-0x18, 0x159-0x15c. More experimentation required.
```
printhex(payload8[0x0:0x19], 32)
printhex(payload8[0x159:0x15d])
```
### More Experimentation
I deleted User Song 4 and saved a new dump. The only difference is the first message.
```
!diff -y --width=100 --suppress-common-lines data/syxout.txt data/syxout1.txt
with open('data/syxout1.txt') as newdump:
newdumpline = bytearray.fromhex(newdump.readline().strip())
newpayload8 = reconstitute_all(newdumpline[14:-2])
printhex(payload8[:0x19], 32)
printhex(newpayload8[:0x19], 32)
print(binbyte(payload8[0]))
print(binbyte(newpayload8[0]))
```
The beginning mystery bytes are the same, except for the first byte, which does indeed seem to be the song usage bitfield, with the least significant bit = user song 1 etc.
```
payload8[0x19:0x15d] == newpayload8[0x19:0x15d]
```
The mystery region is the same for both files.
```
print(tobin(newpayload8[0x15d:0x162], '\n')) #track_bitfields
print('---')
printhex(newpayload8[0x162:0x167])
print('---')
printhex(newpayload8[0x167:0x17b],1,4)
print('---')
printhex(newpayload8[0x17b:0x1f3],6,4)
```
Song 4 is not in use anymore, neither should its tracks be. Their durations are all zero as well.
```
printhex(newpayload8[beginning_blocks], 6)
print('---')
printhex(newpayload8[next_blocks], 16)
```
Blocks 07, 08, 09 and 0B have been marked empty for reuse.
```
payload8[0x2cf:0x700] == newpayload8[0x2cf:]
```
The rest of the data is the same.
## Registration Memory, a.k.a. One Touch Settings
Let's look at the final two messages. In the PSR-225, the one-touch settings are sent first, but the DGX-505 sends 'em last.
```
for line in lines[-2:]:
print(hexline(line))
```
Assuming the messages are of roughly the same format, we have:
| field | bytes | content |
|-------|-------|---------|
|Header | `F0 43 73 7F 44 06` | Manufacturer/Device/etc|
|Section type | `09` | (one-touch settings)|
|Size?? | `06 30 06 2E` | Different format? |
|Running total | `00 00 00` | First (and only) message, so zero |
|Data | `50 53 52 03 01 00...00 00` | payload data|
|Checksum | `30` | We checked this earlier|
|End tag | `F7` | |
So what's with `06 30 06 2E`?
```
otsline = lines[-2]
otspayload = otsline[14:-2]
unpackseven(otsline[7:9])
unpackseven(otsline[9:11])
len(otspayload)
otspayload8 = reconstitute_all(otspayload)
len(otspayload8)
hexline(otspayload8)
```
The PSR-225 had 1576 encoded bytes (788 bytes decoded) of OTS data. The DGX-505 has 816 encoded bytes (714 bytes decoded), but the two length values differ by two, `06 30` (=816) and `06 2E` (=814).
The PSR-225 has 4 buttons × 4 banks = 16 settings, each using 0x31 = 49 bytes. The remaining 4 bytes are bitfields for the active banks on each button.
The DGX-505 has 2 buttons × 8 banks = 16 settings, and has 714 bytes to store it in, which is not a nice number to work with. Maybe the two different length values mean something. Perhaps there are empty bytes at the end that were included so the 7-bit encoding would work. When the dump was taken, all 16 settings were in use, so maybe we should see two `FF` bytes?
```
print(tohex(otspayload8[:4]))
print(tohex(otspayload8[-6:]))
```
The data begins with `50 53 52 03` and ends with `50 53 52 03 00 00`. Perhaps those two extra bytes are the reason for the differing sizes. (As an aside, `50 53 52` is ASCII for 'PSR', but the PSR-225 doesn't have them.)
Disregarding these ten bytes for now leaves us with 704 bytes which divide cleanly into 16 groups of 44.
```
otsgroups = list(takebyn(otspayload8[4:-6], 44))
for group in otsgroups:
print(hexline(group))
```
Like the PSR-225, the settings are stored by button and then by bank.
### Decoding the Format
According to the manual (page 68), the following settings can be saved:
- Style (when using style features):
- Style number
- Auto accompaniment (ON/OFF)
- Split Point
- Style Volume
- Tempo
- Voice:
- Main Voice:
- Voice Number
- Volume
- Octave
- Pan
- Reverb send level
- Chorus send level
- Dual Voice:
- ON/OFF
- ... and the same settings as for Main Voice
- Split Voice:
- ON/OFF
- ... and the same settings as for Main Voice
- Effect:
- Reverb type
- Chorus type
- Panel Sustain (ON/OFF)
- Harmony:
- ON/OFF
- Harmony type
- Harmony volume
- Other:
- Transpose
- Pitch Bend Range
After changing every setting from its previous value in Bank 4, button 2, I saved a new dump... and then I did it again, several times.
(Bank 4, button 2 wasn't a particularly good setting anyway.)
```
def grab_4_2(filename):
with open(filename) as hexdata:
message = bytearray.fromhex(hexdata.readlines()[-2].strip())
return reconstitute_all(message[14:-2])[4+11*44:4+12*44]
# if it's stupid and it works, then... it's still stupid, but hey, at least it works.
old_42 = otsgroups[11]
new_42 = grab_4_2('data/syxout2.txt')
newer_42 = grab_4_2('data/syxout3.txt')
def columnise(some_lines, height):
for i in range(height):
print("".join(some_lines[j] for j in range(i, len(some_lines), height)))
print("|ofst| old | new | newer | "*3)
columnise(["| {0:02X} | {1:02X} {1:3d} | {2:02X} {2:3d} | {3:02X} {3:3d} | ".format(i, oldv, newv, newerv) for i, (oldv, newv, newerv) in enumerate(zip(old_42, new_42, newer_42))], 16)
```
It seems like this is how it works:
| offset | content | values |
|----|--------|--------|
|`00`|?|`01`|
|`01`|Style no.|`FF` = no style; `00`-`87` = 1 – 136, that is, $value=(Style\ number -1)$
|`02`|Accompaniment|`FF` = no style; `00` = off, `01` = on|
|`03`|Split point|`00`-`7F` = 0 (C-2) – 127 (G8), see manual p.71|
|`04`|Split point, again|see above|
|`05`|Main A/B|`FF` = no style; `00` = Main A, `05` = Main B|
|`06`|Style vol|`FF`= no style; `00`-`7F` = volume
|`07`-`08`|Main voice number|`0000`-`01ED` = 1 – 494, that is, $value=(Voice\ number - 1)$ (big-endian)|
|`09`|Main voice octave|`FE`-`02` = -2 – +2 (twos complement for negatives)|
|`0A`|Main voice volume|`00`-`7F` = 0 – 127|
|`0B`|Main voice pan|`00`-`7F` = 0 – 127|
|`0C`|Main voice reverb send level|`00`-`7F` = 0 – 127|
|`0D`|Main voice chorus send level|`00`-`7F` = 0 – 127|
|`0E`|Split on/off|`00` = off; `7F` = on|
|`0F`-`10`|Split voice number|see main voice|
|`11`|Split voice octave|″|
|`12`|Split voice volume|″|
|`13`|Split voice pan|″|
|`14`|Split voice reverb s.l.|″|
|`15`|Split voice chorus s.l.|″|
|`16`|Dual on/off|`00` = off; `7F` = on|
|`17`-`18`|Dual voice number|see main voice|
|`19`|Dual voice octave|″|
|`1A`|Dual voice volume|″|
|`1B`|Dual voice pan|″|
|`1C`|Dual voice reverb s.l.|″|
|`1D`|Dual voice chorus s.l.|″|
|`1E`|Pitch bend range|`01`-`0C` = 1 – 12|
|`1F`|Reverb type|`01`-`0A` = 1 – 10, see manual p.71, 109; `0B`-`0D` see footnote\* |
|`20`|Chorus type|`01`-`05` = 1 – 5, see manual p.71, 109; `06`-`09` see footnote\*|
|`21`|Harmony on/off|`00` = off; `7F` = on|
|`22`|Harmony type|`01`-`1A` = 1 – 26, see manual p.71, 108|
|`23`|Harmony volume|`00`-`7F` = 0 – 127|
|`24`|?|`FF`|
|`25`|Transpose|`00`-`18` = -12 – +12, that is, $value = (transpose + 12)$|
|`26`|Tempo|`FF` = no style; `00`-`F8` = 32 – 280, that is, $value = (bpm - 32)$|
|`27`|?|`00`|
|`28`|?|`00`|
|`29`|Panel sustain|`40` = off; `6E` = on, for some reason|
|`2A`|?|`00`|
|`2B`|?|`00`|
\* *Footnote:* Reverb type and Chorus type have settings that cannot be selected from the panel and can only be chosen by sending MIDI messages (the effect type names without numbers on manual p.113). If these values are saved into one-touch-settings registry, then the values are as follows:
- *Reverb*: `0B` = Room, `0C` = Stage, `0D` = Plate
- *Chorus*: `06` = Thru, `07` = Chorus, `08` = Celeste, `09` = Flanger.
There are some weird things in here, like the repetition of Split Point (maybe one's the split voice split point and one's the accompaniment split point, but the DGX-505's 'Split Point' panel setting always changes both), why the tempo setting is all the way at the end instead of with the rest of the style settings (it is a style setting, as it is not saved in song mode), and why the Panel Sustain has such weird values. There's also the mysterious `01`, `FF`, and `00` values which don't seem to correspond to any changeable setting.
##### Differences when compared to PSR-225
- Settings are in the order *Style - Main voice - Split voice - Dual voice - Pitch bend - Effects - Other* (whereas the PSR-225's order is *Main voice - Dual voice - Split voice - Effects - Style - Other*)
- Instead of storing voices as MIDI Bank and Program, the panel numbers are used.
- Voice octave stored as twos complement instead of being offset by `40`
- Voice pan stored as 0-127, corresponding how the DGX-505 displays it on the panel
- The on/off settings for Split, Dual and Harmony are at the start of their corresponding sections
- The Off settings for Reverb and Chorus are stored as the type, instead of separately, and have numbers `0A` and `05` instead of `00`
- Harmony type uses 1-based indexing instead of 0-based
- Transpose is offset by 12 instead of twos complement
- Settings the DGX-505's format has that the PSR-225 doesn't: Pitch bend range, Panel sustain, Hall2 reverb (The PSR-225's Hall2 appears to correspond to the DGX-505's Hall3 according to the effect map table), Thru chorus
- Settings the PSR-225's format has that the DGX-505 doesn't: Anything to do with DSP, separate voice and accompaniment split points, Footswitch, Touch sensitivity, Tuning, Accompaniment volume
| github_jupyter |
<a href="https://colab.research.google.com/github/srdg/bangla-dl/blob/master/PyTorch_Model.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Data Acquisition
Download the [CMATERdb](https://www.dropbox.com/s/55bhfr3ycvsewsi/CMATERdb%203.1.2.rar) pattern recognition database and clean up `/content` directory.
```
!wget --quiet -O database.rar https://www.dropbox.com/s/55bhfr3ycvsewsi/CMATERdb%203.1.2.rar
!unrar x database.rar > /dev/null
!rm -rf database.rar sample_data
!mv BasicFinalDatabase/ data/
```
# Library imports
Import necessary libaries.
We use `os` for listing files, `albumentations`, `numpy` and `PIL` for image processing, and `matplotlib` to display a subset of the images.
```
import os
import cv2 as cv
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.font_manager as fm
from PIL import Image
import albumentations as A
import torch
from torch.utils.data import Dataset, DataLoader
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
from torchsummary import summary
```
# Exploratory Data Analysis
Let us see how many distinct classes we have. From the data it is already ascertained that we are working with `.bmp` images, which are bitmap images. Let us see how the images are defined in the dataset.
```
imgs = []
for folder in sorted(os.listdir('data/Train')):
imgs.append('data/Train/'+folder+'/bcc000000.bmp')
%matplotlib inline
fig = plt.figure(figsize=(5,5))
fig.suptitle('Characters')
for idx in range(50):
image = Image.open(imgs[idx])
image = np.array(image)
# image = np.array(image)[...,0] # comment prev. line and uncomment this to view a single layer
plt.subplot(10,5,idx+1)
plt.grid(False)
plt.xticks([])
plt.yticks([])
plt.imshow(image, vmin=0, vmax=255)
# plt.imshow(image, cmap='gray') # comment prev. line and uncomment this to view a single layer
plt.subplots_adjust(hspace=0, wspace=0)
plt.show()
```
## Checking image-specific details
Looks like all images do not have the same dimensions. Let us see how many of them are unique and see a sample dimension.
```
img_dims, files = [], []
for folder in sorted(os.listdir('data/Train')):
tmp = ['data/Train/'+folder+'/'+i for i in sorted(os.listdir('data/Train/'+folder))]
for i in tmp:
files.append(i)
img_dims = [np.array(Image.open(i)).shape for i in files]
print("Among", len(img_dims), "images, there are", len(set(img_dims)), "unique dimensions.")
print("Sample dimension:", img_dims[0])
```
### How does the image look in each layer?
It is a bitmap image with 3 layers. Let us look at each layer in a sample image individually and plot them to see if we can identify any discernible changes.
```
f=files[0]
a=np.array(Image.open(f))
for i in range(a.shape[-1]):
plt.subplot(1,3,i+1)
plt.imshow(a[...,i], cmap='gray') # cmap has to be mentioned else plt will map with 'viridis' cmap
```
Interestingly, we see that there is no change in the image in each layer, it is simply stacked up! Maybe we can use a single layer to train a network in this case.
# Defining our labels
Define our labels in Unicode (UTF-8). We associate each directory with a class.
```
classes = {
172:"\u0985",
173:"\u0986",
174:"\u0987",
175:"\u0988",
176:"\u0989",
177:"\u098a",
178:"\u098b",
179:"\u098f",
180:"\u0990",
181:"\u0993",
182:"\u0994",
183:"\u0995",
184:"\u0996",
185:"\u0997",
186:"\u0998",
187:"\u0999",
188:"\u099a",
189:"\u099b",
190:"\u099c",
191:"\u099d",
192:"\u099e",
193:"\u099f",
194:"\u09a0",
195:"\u09a1",
196:"\u09a2",
197:"\u09a3",
198:"\u09a4",
199:"\u09a5",
200:"\u09a6",
201:"\u09a7",
202:"\u09a8",
203:"\u09aa",
204:"\u09ab",
205:"\u09ac",
206:"\u09ad",
207:"\u09ae",
208:"\u09af",
209:"\u09b0",
210:"\u09b2",
211:"\u09b6",
212:"\u09b7",
213:"\u09b8",
214:"\u09b9",
215:"\u09dc",
216:"\u09dd",
217:"\u09df",
218:"\u09ce",
219:"\u0982",
220:"\u0983",
221:"\u0981",
}
print("Classes to train:", end=" ")
for i in list(classes.keys()):
print(classes[i], end=" ")
print()
print("Number of distinct classes:", len(list(classes.keys())))
```
# Preparing file lists
We define the list of files to be used for training and testing.
```
train_files, test_files = [], []
for mode in ['Train', 'Test']:
for folder in sorted(os.listdir('data/'+mode)):
tmp = ['data/'+mode+'/'+folder+'/'+i for i in sorted(os.listdir('data/'+mode+'/'+folder))]
for i in tmp:
if 'Train' in mode:
train_files.append(i)
else:
test_files.append(i)
```
# Defining the dataset
We design a custom dataset to read and help in loading the files.
```
class CharacterDataSet(Dataset):
def __init__(self, files, transform=None):
self.files = files
self.transform = transform
def __len__(self):
return len(self.files)
def __getitem__(self, idx):
image = Image.open(files[idx])
label = int(files[idx].split('/')[2]) # data/Train/172/bcc000000.bmp --> class 172, label 0
image = np.array(image)[...,0]
if self.transform:
image = self.transform(image=image)["image"]
# manual transform (binary thresholding) since library does not support this
# see section "Checking threshold for binarizing images"
# image = cv.threshold(image, 126, 255, cv.THRESH_BINARY)[1] # uncomment to apply
image = image[..., np.newaxis]
image = image /255.0
sample = (image, label-172)
return sample
```
# Transforms and DataLoaders
Here, we attempt to define the image transforms needed for augmentation, following which, we define the dataset objects and use them to create data loader to help in the training and testing procedure.
```
transforms = A.Compose([A.Resize(32,32)])
training_data = CharacterDataSet(train_files, transform=transforms)
testing_data = CharacterDataSet(test_files, transform=transforms)
batch_size = 32
# Create data loaders.
train_dataloader = DataLoader(training_data, batch_size=batch_size, shuffle=True)
test_dataloader = DataLoader(testing_data, batch_size=batch_size, shuffle=True)
```
# Visualizing batches
Let us try to see a sample training batch and a sample testing batch. We will also use our pre-defined `labels` dictionary to identify the labels, and use a custom `Kalpurush` font to print the UTF-8 character labels on a `matplotlib` figure.
```
!wget --quiet -O kalpurush.ttf https://www.omicronlab.com/download/fonts/kalpurush.ttf
%matplotlib inline
prop = fm.FontProperties(fname='kalpurush.ttf')
fig_idx=1
for loader in [train_dataloader, test_dataloader]:
fig = plt.figure(fig_idx, figsize=(5,5))
fig.suptitle('Training batch' if fig_idx is 1 else 'Test batch')
fig_idx+=1
x,y=None,None
for X, Y in loader:
x=X
y=Y
break
for i in range(25):
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.subplots_adjust(hspace=1, wspace=1)
# print(classes[y[i].numpy().tolist()], end=" ")
plt.title(classes[y[i].numpy().tolist()+172], fontproperties=prop)
plt.imshow(x[i].squeeze(), cmap='gray')
plt.show()
class Network(nn.Module):
def __init__(self):
super().__init__()
self.batch_size = batch_size
self.conv1 = nn.Conv2d(in_channels=1, out_channels=32, kernel_size=3,padding=0)
self.pool1 = nn.MaxPool2d(kernel_size=2)
self.conv2 = nn.Conv2d(in_channels=32, out_channels=64, kernel_size=3)
self.pool2 = nn.MaxPool2d(kernel_size=2)
self.drop1 = nn.Dropout(p=0.2)
self.fc1 = nn.Linear(in_features=2304, out_features=128)
self.fc2 = nn.Linear(in_features=128,out_features=50)
def forward(self, x):
x = self.conv1(x.float())
x = F.relu(x)
x = self.pool1(x)
x = self.conv2(x)
x = F.relu(x)
x = self.pool2(x)
x = self.drop1(x)
x = x.view(x.size(0),-1)
x = self.fc1(x)
x = F.relu(x)
x = self.fc2(x)
x = F.relu(x)
return x
def train(dataloader, model, loss_fn, optimizer):
size = len(dataloader.dataset)
for batch, data in enumerate(dataloader):
inputs, labels = data
inputs = np.transpose(inputs, (0,3,1,2))
pred = model(inputs)
loss = loss_fn(pred, labels)
optimizer.zero_grad()
loss.backward()
optimizer.step()
if batch%100==0:
loss, current = loss.item(), batch*len(X)
print(f"loss: {loss:>7f} [{current:>5d}/{size:>5d}]")
def test(dataloader, model, loss_fn):
size = len(dataloader.dataset)
test_loss, correct = 0,0
with torch.no_grad():
for X, y in dataloader:
X = np.transpose(X, (0,3,1,2))
pred = model(X.float())
test_loss += loss_fn(pred, y).item()
correct += (pred.argmax(1) == y).type(torch.float).sum().item()
test_loss /= size
correct /= size
print(f"Test Error : \n Accuracy: {(100*correct):>0.1f}%, Avg loss: {test_loss:>8f}\n")
net = Network()
loss_fn = nn.CrossEntropyLoss()
learning_rate = 1e-3
num_epochs = 20
optimizer = torch.optim.SGD(net.parameters(), lr=learning_rate)
for epoch in range(num_epochs):
print(f"Epoch {epoch+1}\n------------------------")
train(train_dataloader, net, loss_fn, optimizer)
test(test_dataloader, net, loss_fn)
print("Done!")
dataiter = iter(test_dataloader)
images, gt = dataiter.next()
# print(gt,len(gt))
for i in range(25):
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.subplots_adjust(hspace=1, wspace=1)
# print(classes[y[i].numpy().tolist()], end=" ")
plt.title(classes[gt[i].numpy().tolist()+172], fontproperties=prop)
plt.imshow(images[i].squeeze(), cmap='gray')
plt.show()
outputs = net(np.transpose(images, (0,3,1,2)))
_, predicted = torch.max(outputs, 1)
predicted = predicted[:25]
# print([labels[i.numpy().tolist()+172] for i in predicted])
print("Predicted : ",' '.join([classes[i.numpy().tolist()+172] for i in predicted]))
print("Expected : ", ' '.join([classes[i.numpy().tolist()+172] for i in gt[:25]]))
correct = 0
total = 0
with torch.no_grad():
for data in test_dataloader:
image, labels = data
outputs = net(np.transpose(images[:25], (0,3,1,2)))
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted==labels[:25]).sum().item()
print("Accuracy of network: %d%%" % (100 * correct / total))
```
<div align="center"><h1>Debug & Experimentation</h1>
<p>The following region is used for doing experiments on the dataset and is not part of the training procedure.</p></div>
# Rough Network Design
Points to note :
+ The `Conv2d` only supports `TensorFloat32`. Hence, conversion to `x.float()` is necessary.
+ To multiply 2 matrices, their dimensions must be of form `(a,b)` and `(b,c)`, such that the resultant matrix will have dimension `(a,c)`.
+ The `Conv2d` takes in data of the shape `(N, C, H, W)` where
* N = Number of samples (batch size)
* C = Number of channels in each sample
* H = Height of each sample
* W = Width of each sample
+ The end result should be of dimension `(N, C)` where
* N = Number of samples (batch size)
* C = Number of output features
+ During flattening of the final convolved and downsampled layer, `torch.view(prev_layer.shape(0),-1)` should be used. Note the `shape(0)`. `-1` is a wildcard. Intutitively, this results in a matrix of shape `(N, F)` where `N = prev_layer.shape(0) = batch size` and `F = number of features available for each sample`.
```
# Refer https://datascience.stackexchange.com/questions/70086
x, y = None, None
for batch, data in enumerate(train_dataloader, 0):
x,y = data
print("Form (N, H, W, C_in):",x.shape) # --> transpose to form (N, C_in, H, W)
x = np.transpose(x, (0,3,1,2))
print("Form (N, C_in, H, W):",x.shape)
# x = torch.randn(1, 1, 28, 28)
conv1 = torch.nn.Conv2d(in_channels=1, out_channels=32, kernel_size=3, padding=0)(x.float())
print("Convolved layer 1:",conv1.shape)
pool1 = torch.nn.MaxPool2d(kernel_size=2)(conv1)
print("Pooled layer 1:",pool1.shape)
conv2 = torch.nn.Conv2d(in_channels=32, out_channels=64, kernel_size=3)(pool1)
print("Convolved layer 2:",conv2.shape)
pool2 = torch.nn.MaxPool2d(kernel_size=2)(conv2)
print("Pooled layer 2:",pool2.shape)
drop1 = torch.nn.Dropout(p=0.2)
pool2 = drop1(pool2)
print("Dropped layer 2:",pool2.shape)
flat1 = pool2.view(pool2.size(0),-1)
print("Flattened layer 1:", flat1.shape)
fc1 = torch.nn.Linear(2304, 128)(flat1)
print("Fully connected layer 1:", fc1.shape)
fc2 = torch.nn.Linear(128, 50)(fc1)
print("Fully connected layer 2:", fc2.shape)
softmax = torch.nn.Softmax(fc2)
```
# Checking threshold for binarizing images
There are instances when the images contain the values `[0,127,255]` instead of `[0,255]` only. In such cases, we determine the thresholding value as 126 for the tests to be successful. Comment out the manual thresholding line in the CharacterDataset class if this transform is not to be applied.
```
root = 'data/Test/175/bcc000001.bmp'
img = np.array(Image.open(root))
fig = plt.figure(1)
for i in range(img.shape[-1]):
plt.subplot(1,3,i+1)
plt.imshow(img[...,i], cmap='gray')
plt.show()
print("The original image has unique values as",np.unique(img[...,0]).tolist())
img = cv.threshold(img, 126, 255, cv.THRESH_BINARY)[1]
fig = plt.figure(2)
for i in range(img.shape[-1]):
plt.subplot(1,3,i+1)
plt.imshow(img[...,i], cmap='gray')
plt.show()
transform = A.Compose([A.Resize(32,32)])
img_file = train_files[0]
img_arr = np.array(Image.open(img_file))[...,0]
img_arr = img_arr[..., np.newaxis]
transformed = transform(image=img_arr)
img_arr = transformed["image"]
img_arr = img_arr/255.0
unique, counts = np.unique(img_arr, return_counts=True)
img_hist = dict(zip(unique, counts))
plt.imshow(np.squeeze(img_arr), cmap='gray')
```
| github_jupyter |
# A jupyter notebook is a browser-based environment that integrates:
- A Kernel (python)
- Text
- Executable code
- Plots and images
- Rendered mathematical equations
## Cell
The basic unit of a jupyter notebook is a `cell`. A `cell` can contain any of the above elements.
In a notebook, to run a cell of code, hit `Shift-Enter`. This executes the cell and puts the cursor in the next cell below, or makes a new one if you are at the end. Alternately, you can use:
- `Alt-Enter` to force the creation of a new cell unconditionally (useful when inserting new content in the middle of an existing notebook).
- `Control-Enter` executes the cell and keeps the cursor in the same cell, useful for quick experimentation of snippets that you don't need to keep permanently.
## Hello World
```
print("Hello World!")
# lines that begin with a # are treated as comment lines and not executed
# print("This line is not printed")
print("This line is printed")
```
## Create a variable
```
my_variable = 3.0 * 2.0
```
## Print out the value of the variable
```
print(my_variable)
```
## or even easier:
```
my_variable
```
# Datatypes
In computer programming, a data type is a classification identifying one of various types that data
can have.
The most common data type we will see in this class are:
* **Integers** (`int`): Integers are the classic cardinal numbers: ... -3, -2, -1, 0, 1, 2, 3, 4, ...
* **Floating Point** (`float`): Floating Point are numbers with a decimal point: 1.2, 34.98, -67,23354435, ...
- Floating point values can also be expressed in scientific notation: 1e3 = 1000
* **Booleans** (`bool`): Booleans types can only have one of two values: `True` or `False`. In many languages 0 is considered `False`, and any other value is considered `True`.
* **Strings** (`str`): Strings can be composed of one or more characters: ’a’, ’spam’, ’spam spam eggs and spam’. Usually quotes (’) are used to specify a string. For example ’12’ would refer to the string, not the integer.
## Collections of Data Types
* **Scalar**: A single value of any data type.
* **List**: A collection of values. May be mixed data types. (1, 2.34, ’Spam’, True) including lists of lists: (1, (1,2,3), (3,4))
* **Array**: A collection of values. Must be same data type. [1,2,3,4] or [1.2, 4.5, 2.6] or [True, False, False] or [’Spam’, ’Eggs’, ’Spam’]
* **Matrix**: A multi-dimensional array: [[1,2], [3,4]] (an array of arrays).
```
my_var_a = 1
my_var_b = 2.3
my_var_c = 2.3e4
my_var_d = True
my_var_e = "Spam"
type(my_var_a), type(my_var_b), type(my_var_c), type(my_var_d), type(my_var_e)
my_var_a + my_var_b, type(my_var_a + my_var_b)
my_var_c + my_var_d, type(my_var_c + my_var_d) # True = 1
my_var_a + my_var_e
str(my_var_a) + my_var_e
```
# NumPy (Numerical Python) is the fundamental package for scientific computing with Python.
### Load the numpy library:
```
import numpy as np
```
#### pi and e are built-in constants:
```
np.pi, np.e
```
## Here is a link to all [Numpy math functions](https://docs.scipy.org/doc/numpy/reference/routines.math.html).
# Arrays
* Each element of the array has a **Value**
* The *position* of each **Value** is called its **Index**
## Our basic unit will be the NumPy array
```
np.random.seed(42) # set the seed - everyone gets the same random numbers
my_array = np.random.randint(1,10,20) # 20 random ints between 1 and 10
my_array
```
## Indexing
```
my_array[0] # The Value at Index = 0
my_array[-1] # The last Value in the array
```
## Slices
`x[start:stop:step]`
- `start` is the first Index that you want [default = first element]
- `stop` is the first Index that you **do not** want [default = last element]
- `step` defines size of `step` and whether you are moving forwards (positive) or backwards (negative) [default = 1]
```
my_array
my_array[0:4] # first 4 items
my_array[:4] # same
my_array[0:4:2] # first four item, step = 2
my_array[3::-1] # first four items backwards, step = -1
my_array[::-1] # Reverse the array x
print(my_array[-5:]) # last 5 elements of the array x
```
## There are lots of different `methods` that can be applied to a NumPy array
```
my_array.size # Number of elements in x
my_array.mean() # Average of the elements in x
my_array.sum() # Total of the elements in x
my_array[-5:].sum() # Total of last 5 elements in x
my_array.cumsum() # Cumulative sum
my_array.cumsum()/my_array.sum() # Cumulative percentage
```
```
my_array.
```
## Help about a function:
```
?my_array.min
```
## NumPy math works over an entire array:
```
my_array * 2
sin(my_array) # need to Numpy's math functions
np.sin(my_array)
```
## Masking - The key to fast programs
```
mask1 = np.where(my_array > 5)
my_array, mask1
my_array[mask1]
mask2 = np.where((my_array>3) & (my_array<7))
my_array[mask2]
```
## Fancy masking
```
mask3 = np.where(my_array >= 7)
my_array[mask3]
# Set all values of x that match mask3 to 0
my_array[mask3] = 0
my_array
mask4 = np.where(my_array != 0)
mask4
#Add 10 to every value of x that matches mask4:
my_array[mask4] += 100
my_array
```
## Sorting
```
np.random.seed(13) # set the seed - everyone gets the same random numbers
my_other_array = np.random.randint(1,10,20) # 20 random ints between 1 and 10
my_other_array
np.sort(my_other_array)
np.sort(my_other_array)[0:4]
```
# Control Flow
Like all computer languages, Python supports the standard types of control flows including:
* IF statements
* FOR loops
```
my_variable = -1
if my_variable > 0:
print("This number is positive")
else:
print("This number is NOT positive")
my_variable = 0
if my_variable > 0:
print("This number is positive")
elif my_variable == 0:
print("This number is zero")
else:
print("This number is negative")
```
## `For loops` are different in python.
You do not need to specify the beginning and end values of the loop
```
my_other_array
for value in my_other_array:
print(value)
for idx,val in enumerate(my_other_array):
print(idx,val)
for idx,val in enumerate(my_other_array):
if (val > 5):
my_other_array[idx] = 0
for idx,val in enumerate(my_other_array):
print(idx,val)
```
## Loops are slow in Python. Do not use them if you do not have to!
```
np.random.seed(42)
my_BIG_Array = np.random.random(10000) # 10,000 value array
my_BIG_Array[:10]
# This is slow!
for Idx,Val in enumerate(my_BIG_Array):
if (Val > 0.5):
my_BIG_Array[Idx] = 0
my_BIG_Array[:10]
%%timeit
for Idx,Val in enumerate(my_BIG_Array):
if (Val > 0.5):
my_BIG_Array[Idx] = 0
# Masks are MUCH faster
mask_BIG = np.where(my_BIG_Array > 0.5)
my_BIG_Array[mask_BIG] = 0
my_BIG_Array[:10]
%%timeit -o
mask_BIG = np.where(my_BIG_Array > 0.5)
my_BIG_Array[mask_BIG] = 0
```
# Functions
In computer science, a `function` (also called a `procedure`, `method`, `subroutine`, or `routine`) is a portion
of code within a larger program that performs a specific task and is relatively independent of the
remaining code. The big advantage of a `function` is that it breaks a program into smaller, easier
to understand pieces. It also makes debugging easier. A `function` can also be reused in another
program.
The basic idea of a `function` is that it will take various values, do something with them, and `return` a result. The variables in a `function` are local. That means that they do not affect anything outside the `function`.
Below is a simple example of a `function` that solves the equation:
$ f(x,y) = x^2\ sin(y)$
In the example the name of the `function` is **find_f** (you can name `functions` what ever you want). The `function` **find_f** takes two arguments `x` and `y`, and returns the value of the equation to the main program. In the main program a variable named `value_f` is assigned the value returned by **find_f**. Notice that in the main program the `function` **find_f** is called using the arguments `array_x` and `array_y`. Since the variables in the `function` are local, you do not have name them `x` and `y` in the main program.
```
def find_f(my_x, my_y):
result = (my_x ** 2) * np.sin(my_y) # assign the variable result the value of the function
return result # return the value of the function to the main program
np.random.seed(42)
array_x = np.random.rand(10) * 10
array_y = np.random.rand(10) * 2.0 * np.pi
array_x, array_y
value_f = find_f(array_x,array_y)
value_f
```
### The results of one function can be used as the input to another function
```
def find_g(my_z):
result = my_z / np.e
return result
find_g(value_f)
find_g(find_f(array_x,array_y))
```
# Creating Arrays
## Numpy has a wide variety of ways of creating arrays: [Array creation routines](https://docs.scipy.org/doc/numpy-1.13.0/reference/routines.array-creation.html)
```
# a new array filled with zeros
array_0 = np.zeros(10)
array_0
# a new array filled with ones
array_1 = np.ones(10)
array_1
# a new array filled with evenly spaced values within a given interval
array_2 = np.arange(10,20)
array_2
# a new array filled with evenly spaced numbers over a specified interval (start, stop, num)
array_3 = np.linspace(10,20,5)
array_3
# a new array filled with evenly spaced numbers over a log scale. (start, stop, num, base)
array_4 = np.logspace(1,2,5,10)
array_4
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
import arrow
import random
import pandas as pd
import numpy as np
from datetime import datetime
from datagen.csvgen.base import randgen, randrec
from datagen.csvgen.ner import converter
from tqdm import tqdm
from pandas.core.common import flatten
from collections import OrderedDict
from datagen.config import data_config
from datagen.imgen.content import ner_utils
from datagen.imgen.ops import boxes_ops
import json
import chardet
import torch.nn as nn
from transformers import BertTokenizer, AutoModel
tokenizer = BertTokenizer.from_pretrained("indobenchmark/indobert-base-p2")
#label config
def labels_map(label_name, label_type):
labels=[]
for kn,vn in label_name.items():
for kt,vt in label_type.items():
if kt!='delimiter':
for bil in "BILU":
name = f'{bil}-{vt}_{vn}'
labels.append(name)
labels.append("O")
return labels
label_name = {
'provinsi': 'PROV',
'kabupaten': 'KAB',
'nik': 'NIK',
'nama': 'NAMA',
'ttl': 'TTL',
'gender': 'GDR',
'goldar': 'GLD',
'alamat': 'ADR',
'rtrw': 'RTW',
'kelurahan': 'KLH',
'kecamatan': 'KCM',
'agama': 'RLG',
'perkawinan': 'KWN',
'pekerjaan': 'KRJ',
'kewarganegaraan': 'WRG',
'berlaku': 'BLK',
'sign_place': 'SGP',
'sign_date': 'SGD'
}
label_type = {
'field': "FLD",
'value': "VAL",
"delimiter": "O"
}
LABELS = labels_map(label_name, label_type)
LABEL2INDEX = dict((label,idx) for idx, label in enumerate(LABELS))
INDEX2LABEL = dict((idx, label) for idx, label in enumerate(LABELS))
NUM_LABELS = len(LABELS)
def get_formatted_annotation_objects(anno, tokenizer, max_seq_length=512):
objects = anno['objects']
objects = prepare_objects_data(objects, tokenizer)
tokens, labels, label_ids, boxes = [],[],[],[]
tokens.append(cls_token)
boxes.append(cls_token_box)
label_ids.append(pad_token_label_id)
for obj in objects:
tokens.append(obj['token'])
labels.append(obj['label'])
lab = LABEL2INDEX[obj['label']]
label_ids.append(lab)
pts = obj['points']
pts = np.array(pts)
pts = boxes_ops.order_points(np.array(pts))
pts = list(boxes_ops.to_xyminmax(pts))
boxes.append(pts)
tokens.append(sep_token)
boxes.append(sep_token_box)
label_ids.append(pad_token_label_id)
input_ids = tokenizer.encode(tokens, add_special_tokens=False)
input_masks = [1] * len(input_ids)
segment_ids = [0] * len(input_ids)
padding_data_ouput = padding_data(
input_ids, input_masks,
segment_ids, label_ids, boxes,
max_seq_length=max_seq_length
)
input_ids, input_masks, segment_ids, label_ids, boxes = padding_data_ouput
return input_ids, input_masks, segment_ids, label_ids, boxes
def padding_data(input_ids, input_masks, segment_ids,
label_ids, boxes, max_seq_length=512,
pad_on_left=False):
padding_length = max_seq_length - len(input_ids)
if not pad_on_left:
input_ids += [pad_token] * padding_length
input_masks += [0] * padding_length
segment_ids += [0] * padding_length
label_ids += [pad_token_label_id] * padding_length
boxes += [pad_token_box] * padding_length
else:
input_ids = [pad_token] * padding_length + input_ids
input_masks = [0] * padding_length + input_masks
segment_ids = [0] * padding_length + segment_ids
label_ids = [pad_token_label_id] * padding_length + label_ids
boxes = [pad_token_box] * padding_length + boxes
return input_ids, input_masks, segment_ids, label_ids, boxes
def prepare_objects_data(objects, tokenizer):
duplicated_objects = tokenize_duplicate_dict(objects, tokenizer)
formatted_objects = reformat_label_oriented(duplicated_objects)
bilou_objects = inject_bilou_to_objects(formatted_objects)
objects = revert_to_list_format(bilou_objects)
return objects
def reformat_label_oriented(objects):
data = OrderedDict({k:{'field':[], 'delimiter':[], 'value':[]} for k,v in label_name.items()})
for idx, obj in enumerate(objects):
cname_curr = obj['classname']
scname_curr = obj['subclass']
data[cname_curr][scname_curr].append(obj)
return data
# datas['objects']
def bilou_prefixer(text_list, label=None):
out = []
text_len = len(text_list)
if text_len==1:
bl = "U"
if label!=None: bl = bl + "-" + label
out.append(bl)
elif text_len>1:
for idx, text in enumerate(text_list):
if idx==0:
bl = "B"
if label!=None: bl = bl + "-" + label
out.append(bl)
elif idx < text_len - 1:
bl = "I"
if label!=None: bl = bl + "-" + label
out.append(bl)
else:
bl = "L"
if label!=None: bl = bl + "-" + label
out.append(bl)
return out
def tokenize_inside_dict(data_dict, tokenizer):
for idx in range(len(data_dict)):
text = data_dict[idx]['text']
data_dict[idx]['text'] = tokenizer.tokenize(text)
return data_dict
def tokenize_duplicate_dict(objects, tokenizer):
new_objects = []
for idx, obj in enumerate(objects):
curr_text = objects[idx]['text']
token = tokenizer.tokenize(curr_text)
if len(token) > 1:
for tok in token:
new_obj = objects[idx].copy()
new_obj['token'] = tok
new_objects.append(new_obj)
else:
if len(token)==0:
obj['token'] = ''
else:
obj['token'] = token[0]
new_objects.append(obj)
return new_objects
def inject_bilou_to_label(data_dict):
# create bilou prefix to dictionary data
texts = []
for idx in range(len(data_dict)):
texts.append(data_dict[idx]['token'])
bil_prefix = bilou_prefixer(texts)
#inject bilou prefix into label inside data_dict
for idx, (bil, fld) in enumerate(zip(bil_prefix, data_dict)):
if fld['label'] != "O":
label = bil+'-'+fld['label']
data_dict[idx]['label'] = label
return data_dict
def inject_bilou_to_objects(objects):
for idx, (key,val) in enumerate(objects.items()):
field = val['field']
delim = val['delimiter']
value = val['value']
if len(field)>0:
objects[key]['field'] = inject_bilou_to_label(field)
if len(delim)>0:
objects[key]['delimiter'] = inject_bilou_to_label(delim)
if len(value)>0:
objects[key]['value'] = inject_bilou_to_label(value)
return objects
def revert_to_list_format(dnew):
data_list = []
for k,v in dnew.items():
field = dnew[k]['field']
delim = dnew[k]['delimiter']
value = dnew[k]['value']
if len(delim)>0:
line_list = field+delim+value
else:
line_list = field+value
data_list += line_list
return data_list
path = '../results/combined/1606064001/7917_json.json'
with open(path) as f:
datas = json.load(f)
objects = prepdatas['objects']
cls_token_at_end=False
cls_token="[CLS]"
sep_token="[SEP]"
sep_token_extra=False
cls_token_segment_id=1
pad_token_segment_id=0
sequence_a_segment_id=0
pad_on_left=False
pad_token=0
cls_token_box=[0, 0, 0, 0]
sep_token_box=[1500, 1500, 1500, 1500]
pad_token_box=[0, 0, 0, 0]
pad_token_label_id = nn.CrossEntropyLoss().ignore_index
max_seq_length = 512
tokenizer.cls_token_id, tokenizer.sep_token_id, tokenizer.pad_token_id
input_ids, input_masks, segment_ids, label_ids, boxes = build_data(objects, tokenizer)
idx = 17
word = tokenizer.ids_to_tokens[input_ids[idx]]
label = INDEX2LABEL[label_ids[idx]]
word, label, boxes[idx]
import os
import cv2 as cv
from pathlib import Path
import matplotlib.pyplot as plt
from torch.utils.data import dataset
from sklearn.model_selection import train_test_split
class IDCardDataset(dataset.Dataset):
def __init__(self, root, tokenizer, labels=None, mode='train',
test_size=0.2, max_seq_length=512):
self.root = Path(root)
self.tokenizer = tokenizer
self.labels = labels
self.mode = mode
self.test_size = test_size
self.max_seq_length = max_seq_length
self._build_files()
def _build_files(self):
names = self._get_names("*_json.json")
data = self._get_filtered_files(names)
dframe, train, test = self._split_dataset(data)
self.data_frame = dframe
self.train_frame = train
self.test_frame = test
if self.mode=="train":
self.frame = self.train_frame
else:
self.frame = self.test_frame
def _split_dataset(self, data):
dframe = pd.DataFrame(data)
train, test = train_test_split(dframe,
test_size=self.test_size,
random_state=1261)
train = train.reset_index(drop=True)
test = test.reset_index(drop=True)
return dframe, train, test
def _get_filtered_files(self, names):
data = {'name':[], 'image':[], 'mask':[],'anno':[]}
jfiles = self._glob_filter("*_json.json")
ifiles = self._glob_filter("*_image.jpg")
mfiles = self._glob_filter("*_mask.jpg")
for name in names:
for (jfile, ifile, mfile) in zip(jfiles, ifiles, mfiles):
jfpn = jfile.name.split("_")[0]
ifpn = ifile.name.split("_")[0]
mfpn = mfile.name.split("_")[0]
if name == jfpn and name == ifpn and name == mfpn:
data['name'].append(name)
data['image'].append(ifile)
data['mask'].append(mfile)
data['anno'].append(jfile)
return data
def _get_names(self, path_pattern):
names = []
files = self._glob_filter(path_pattern)
for file in files:
names.append(file.name.split("_")[0])
return names
def _glob_filter(self, pattern):
return sorted(list(self.root.glob(pattern)))
def _load_anno(self, path):
path = str(path)
with open(path) as f:
data_dict = json.load(f)
return data_dict
def _load_image(self, path):
path = str(path)
img = cv.imread(path, cv.IMREAD_UNCHANGED)
return img
def __len__(self):
return len(self.frame)
def __getitem__(self, idx):
record = self.frame.iloc[idx]
anno = self._load_anno(record['anno'])
img = self._load_image(record['image'])
mask = self._load_image(record['mask'])
anno_objects = get_formatted_annotation_objects(
anno, self.tokenizer, self.max_seq_length
)
input_ids, input_masks, segment_ids, label_ids, boxes = anno_objects
data = (
input_ids, input_masks,
segment_ids, label_ids, boxes,
img, mask
)
return data
path = '../results/combined/1606064001/'
data = IDCardDataset(root=path, tokenizer=tokenizer)
# len(data[0])
data[0]
import os
print(os.getcwd())
```
| github_jupyter |
Outdated notebook with old, unused experiment plots
```
import pandas as pd
import numpy as np
import os
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
sns.set_style('darkgrid')
%matplotlib inline
multitask_dir = '../experiment_output/multitask_base_experiment/evaluation/'
multif0_dir = '../experiment_output/multitask_singletask_mf0/evaluation/'
melody_dir = '../experiment_output/multitask_singletask_mel/evaluation/'
bass_dir = '../experiment_output/multitask_singletask_bass/evaluation/'
vocal_dir = '../experiment_output/multitask_singletask_vocal/evaluation/'
multitask_nosynth_dir = '../experiment_output/multitask_nosynth/evaluation/'
multitask_freq_dir = '../experiment_output/multitask_freq_feature/evaluation/'
multitask_freq_nosynth_dir = '../experiment_output/multitask_freq_nosynthdata/evaluation/'
melody_freq_dir = '../experiment_output/multitask_singletask_mel_freq/evaluation/'
bass_freq_dir = '../experiment_output/multitask_singletask_bass_freq/evaluation/'
vocal_freq_dir = '../experiment_output/multitask_singletask_vocal_freq/evaluation/'
def get_dfbox(dataset_scores, metrics, folder_list):
df_list = []
for folder in folder_list:
df = pd.DataFrame.from_csv(os.path.join(folder, dataset_scores))
df_list.append(df)
# print(folder)
# print(df.describe())
# print("")
boxdata = []
for metric in metrics:
boxdata.extend([df[metric] for df in df_list])
dfbox = pd.DataFrame(np.array(boxdata).T)
return dfbox
def add_plot(subplot_tuple, metrics, exper_names, dfbox, title, show_yaxis=True, legend_loc=None, xlim=None):
plt.subplot(subplot_tuple)
plt.title(title, weight='bold')
n_metrics = len(metrics)
n_exper = len(exper_names)
positions = []
k = 1
for i in range(n_metrics):
for j in range(n_exper):
positions.append(k)
k = k + 1
k = k + 1
y_pos = []
for i in np.arange(0, len(positions), n_exper):
y_pos.append(np.mean(positions[i:i + n_exper]))
current_palette = sns.color_palette('deep', n_exper) #["#E1D89F", "#8EA8BD", "#CF6766"]
colors = current_palette*n_metrics
box = plt.boxplot(
dfbox.values, widths=0.8, positions=positions,
patch_artist=True, showmeans=True,
medianprops={'color': 'k'},
meanprops=dict(marker='D', markeredgecolor='black',
markerfacecolor='k'),
vert=False
)
for patch, color in zip(box['boxes'], colors):
patch.set_facecolor(color)
plt.xlabel('Score')
if show_yaxis:
plt.yticks(y_pos, metrics, rotation='horizontal', weight='bold')
else:
plt.yticks(y_pos, ['']*len(metrics), rotation='horizontal')
if xlim is not None:
plt.xlim(xlim)
if legend_loc is not None:
h_list = []
for i, name in enumerate(exper_names):
h, = plt.plot([1, 1], 's', color=colors[i], markersize=10)
h_list.append(h)
lgd = plt.legend(tuple(h_list), tuple(exper_names), ncol=1, loc=legend_loc)
for h in h_list:
h.set_visible(False)
```
# Multitask vs Single Task
# Multif0
```
metrics = ['Accuracy', 'Chroma Accuracy', 'Precision', 'Recall']
exper_names = ['multitask', 'single-task', 'multitask-freq', 'multitask-freq-nosynth', 'multitask-nosynth']
exper_folders = [multitask_dir, multif0_dir, multitask_freq_dir, multitask_freq_nosynth_dir, multitask_nosynth_dir]
df_bach10 = get_dfbox('bach10_scores.csv', metrics, exper_folders)
df_su = get_dfbox('su_scores.csv', metrics, exper_folders)
df_mdbmf0 = get_dfbox('mdb_mf0_scores.csv', metrics, exper_folders)
df_maps = get_dfbox('maps_scores.csv', metrics, exper_folders)
fig = plt.figure(figsize=(15, 7))
add_plot(141, metrics, exper_names, df_bach10, '[Multif0] Bach10', xlim=[0.0, 1.0], legend_loc=0)
add_plot(142, metrics, exper_names, df_su, '[Multif0] Su', show_yaxis=False, xlim=[0.0, 1.0])
add_plot(143, metrics, exper_names, df_mdbmf0, '[Multif0] MedleyDB', xlim=[0.0, 1.0], show_yaxis=False)
add_plot(144, metrics, exper_names, df_maps, '[Multif0] MAPS', xlim=[0.0, 1.0], show_yaxis=False)
```
# Melody
```
metrics = ['Overall Accuracy', 'Raw Pitch Accuracy', 'Raw Chroma Accuracy', 'Voicing Recall']#, 'Voicing False Alarm']
exper_names = ['multitask', 'multitask-freq', 'multitask-freq-nosynth', 'multitask-nosynth', 'single-task', 'single-task-freq']
exper_folders = [multitask_dir, multitask_freq_dir, multitask_freq_nosynth_dir, multitask_nosynth_dir, melody_dir, melody_freq_dir]
df_mdbmel = get_dfbox('mdb_mel_scores.csv', metrics, exper_folders)
df_orchset = get_dfbox('orchset_scores.csv', metrics, exper_folders)
df_wjmel = get_dfbox('wj_mel_scores.csv', metrics, exper_folders)
fig = plt.figure(figsize=(15, 7))
add_plot(131, metrics, exper_names, df_mdbmel, '[Melody] MedleyDB', xlim=[0.0, 1.0])
add_plot(132, metrics, exper_names, df_orchset, '[Melody] Orchset', show_yaxis=False, xlim=[0.0, 1.0])
add_plot(133, metrics, exper_names, df_wjmel, '[Melody] Weimar Jazz', xlim=[0.0, 1.0], show_yaxis=False, legend_loc=0)
```
# Bass + Vocal
```
metrics = ['Overall Accuracy', 'Raw Pitch Accuracy', 'Raw Chroma Accuracy', 'Voicing Recall']
exper_names = ['multitask', 'multitask-freq', 'multitask-freq-nosynth', 'multitask-nosynth', 'single-task', 'single-task-freq']
exper_folders = [multitask_dir, multitask_freq_dir, multitask_freq_nosynth_dir, multitask_nosynth_dir, bass_dir, bass_freq_dir]
df_wjbass = get_dfbox('wj_bass_scores.csv', metrics, exper_folders)
exper_folders = [multitask_dir, multitask_freq_dir, multitask_freq_nosynth_dir, multitask_nosynth_dir, vocal_dir, vocal_dir]
df_ikala = get_dfbox('ikala_scores.csv', metrics, exper_folders)
fig = plt.figure(figsize=(12, 8))
add_plot(121, metrics, exper_names, df_wjbass, '[Bass] Weimar Jazz', xlim=[0.0, 1.0])
add_plot(122, metrics, exper_names, df_ikala, '[Vocal] iKala', xlim=[0.0, 1.0], legend_loc=0, show_yaxis=False)
```
# Vocal
```
# metrics = ['Overall Accuracy', 'Raw Pitch Accuracy', 'Raw Chroma Accuracy', 'Voicing Recall']#, 'Voicing False Alarm']
# exper_names = ['multitask', 'single-task', '']
# exper_folders = [multitask_dir, vocal_dir]
# df_ikala = get_dfbox('ikala_scores.csv', metrics, exper_folders)
# fig = plt.figure(figsize=(10, 4))
# add_plot(131, metrics, exper_names, df_ikala, 'iKala', xlim=[0.0, 1.0], legend_loc=0)
plt.figure(figsize=(15, 20))
add_plot(341, metrics, exper_names, df_bach10, '[Multif0] Bach10', xlim=[0.0, 1.0], legend_loc=2)
add_plot(342, metrics, exper_names, df_su, '[Multif0] Su', show_yaxis=False, xlim=[0.0, 1.0])
add_plot(343, metrics, exper_names, df_mdbmf0, '[Multif0] MedleyDB', xlim=[0.0, 1.0], show_yaxis=False)
add_plot(344, metrics, exper_names, df_maps, '[Multif0] MAPS', xlim=[0.0, 1.0], show_yaxis=False)
add_plot(334, metrics, exper_names, df_mdbmel, '[Melody] MedleyDB', xlim=[0.0, 1.0])
add_plot(335, metrics, exper_names, df_orchset, '[Melody] Orchset', show_yaxis=False, xlim=[0.0, 1.0])
add_plot(336, metrics, exper_names, df_wjmel, '[Melody] Weimar Jazz', xlim=[0.0, 1.0], show_yaxis=False)
add_plot(325, metrics, exper_names, df_wjbass, '[Bass] Weimar Jazz', xlim=[0.0, 1.0])
add_plot(326, metrics, exper_names, df_ikala, '[Vocal] iKala', xlim=[0.0, 1.0], show_yaxis=False)
plt.savefig('../paper-figs/multitask_single_vs_multi.pdf', format='pdf', bbox_inches='tight')
plt.figure(figsize=(15, 20))
add_plot(341, metrics, exper_names, df_bach10, '[Multif0] Bach10', xlim=[0.0, 1.0], legend_loc=2)
add_plot(342, metrics, exper_names, df_su, '[Multif0] Su', show_yaxis=False, xlim=[0.0, 1.0])
add_plot(343, metrics, exper_names, df_mdbmf0, '[Multif0] MedleyDB', xlim=[0.0, 1.0], show_yaxis=False)
add_plot(344, metrics, exper_names, df_maps, '[Multif0] MAPS', xlim=[0.0, 1.0], show_yaxis=False)
add_plot(334, metrics, exper_names, df_mdbmel, '[Melody] MedleyDB', xlim=[0.0, 1.0])
add_plot(335, metrics, exper_names, df_orchset, '[Melody] Orchset', show_yaxis=False, xlim=[0.0, 1.0])
add_plot(336, metrics, exper_names, df_wjmel, '[Melody] Weimar Jazz', xlim=[0.0, 1.0], show_yaxis=False)
add_plot(325, metrics, exper_names, df_wjbass, '[Bass] Weimar Jazz', xlim=[0.0, 1.0])
add_plot(326, metrics, exper_names, df_ikala, '[Vocal] iKala', xlim=[0.0, 1.0], show_yaxis=False)
# plt.savefig('../paper-figs/multitask_single_vs_multi.pdf', format='pdf', bbox_inches='tight')
```
| github_jupyter |
# 汎用センサー型向け 信号定義サンプル
iPhoneのセンサー情報をFloat型に変換するための信号定義を登録します。
```
Warning :
既にサーバーに信号定義が登録されていた場合、同じlabelの信号定義を登録することはできません。
サーバーに対象の信号定義が登録されていないことを確認し、実行してください。
既に登録されている信号定義を確認したい場合、signals.list() を実行してください。
```
## データ種別一覧
汎用センサーデータには、以下のデータ種別が設けられています。
| ID | 英名 | 和名 |
|:-------|:------------------------------------------------------------------------------------------|:--------------------------|
| 0x0001 | Acceleration | 加速度 **(チュートリアルで使用)** |
| 0x0002 | Acceleration Including Gravity | 加速度+重力加速度 |
| 0x0003 | Gravity | 重力加速度 |
| 0x0004 | Rotation Rate | 回転速度 |
| 0x0005 |Orientation Angle | 回転角度 **(チュートリアルで使用)** |
| 0x0006 | GeoLocation Coordinate | 地理的位置情報 座標 |
| 0x0007 | GeoLocation Altitude | 地理的位置情報 高度 |
| 0x0008 | GeoLocation Accuracy | 地理的位置情報 精度 |
| 0x0009 | GeoLocation Heading | 地理的位置情報 方角 |
| 0x000A | GeoLocation Speed | 地理的位置情報 移動速度
## 事前準備
クライアントを生成します。
```
import intdash
client = intdash.Client(
url = "https://example.intdash.jp",
username = "edge1",
password="password_here"
)
```
信号定義のアクセスオブジェクトを生成します。
```
sig = client.signals
```
## 信号定義の登録
### 0001 Acceleration
物体に作用する3軸の加速度
```
s = sig.create(
label="sp_ACCX",
data_type=intdash.DataType.general_sensor,
data_id="0001",
channel=1,
conversion=intdash.ConversionNumFixedPoint(
startbit=0,
bitsize=32,
endian="little",
sign="signed",
scale=0.000001,
offset=0,
),
display=intdash.Display(
unit="",
min=-10,
max=10,
format="%f",
),
)
print(s)
s = sig.create(
label="sp_ACCY",
data_type=intdash.DataType.general_sensor,
data_id="0001",
channel=1,
conversion=intdash.ConversionNumFixedPoint(
startbit=32,
bitsize=32,
endian="little",
sign="signed",
scale=0.000001,
offset=0,
),
display=intdash.Display(
unit="",
min=-10,
max=10,
format="%f",
),
)
print(s)
s = sig.create(
label="sp_ACCZ",
data_type=intdash.DataType.general_sensor,
data_id="0001",
channel=1,
conversion=intdash.ConversionNumFixedPoint(
startbit=64,
bitsize=32,
endian="little",
sign="signed",
scale=0.000001,
offset=0,
),
display=intdash.Display(
unit="",
min=-10,
max=10,
format="%f",
),
)
print(s)
```
### 0002 Acceleration Including Gravity
物体に作用する、重力を含む3軸の加速度
```
s = sig.create(
label="sp_AGX",
data_type=intdash.DataType.general_sensor,
data_id="0002",
channel=1,
conversion=intdash.ConversionNumFixedPoint(
startbit=0,
bitsize=32,
endian="little",
sign="signed",
scale=0.000001,
offset=0,
),
display=intdash.Display(
unit="",
min=-10,
max=10,
format="%f",
),
)
print(s)
s = sig.create(
label="sp_AGY",
data_type=intdash.DataType.general_sensor,
data_id="0002",
channel=1,
conversion=intdash.ConversionNumFixedPoint(
startbit=32,
bitsize=32,
endian="little",
sign="signed",
scale=0.000001,
offset=0,
),
display=intdash.Display(
unit="",
min=-10,
max=10,
format="%f",
),
)
print(s)
s = sig.create(
label="sp_AGZ",
data_type=intdash.DataType.general_sensor,
data_id="0002",
channel=1,
conversion=intdash.ConversionNumFixedPoint(
startbit=64,
bitsize=32,
endian="little",
sign="signed",
scale=0.000001,
offset=0,
),
display=intdash.Display(
unit="",
min=-10,
max=10,
format="%f",
),
)
print(s)
```
### 0003 Gravity
物体に作用する3軸の重力加速度
```
s = sig.create(
label="sp_GX",
data_type=intdash.DataType.general_sensor,
data_id="0003",
channel=1,
conversion=intdash.ConversionNumFixedPoint(
startbit=0,
bitsize=32,
endian="little",
sign="signed",
scale=0.000001,
offset=0,
),
display=intdash.Display(
unit="",
min=-10,
max=10,
format="%f",
),
)
print(s)
s = sig.create(
label="sp_GY",
data_type=intdash.DataType.general_sensor,
data_id="0003",
channel=1,
conversion=intdash.ConversionNumFixedPoint(
startbit=32,
bitsize=32,
endian="little",
sign="signed",
scale=0.000001,
offset=0,
),
display=intdash.Display(
unit="",
min=-10,
max=10,
format="%f",
),
)
print(s)
s = sig.create(
label="sp_GZ",
data_type=intdash.DataType.general_sensor,
data_id="0003",
channel=1,
conversion=intdash.ConversionNumFixedPoint(
startbit=64,
bitsize=32,
endian="little",
sign="signed",
scale=0.000001,
offset=0,
),
display=intdash.Display(
unit="",
min=-10,
max=10,
format="%f",
),
)
print(s)
```
### 0004 Rotation Rate
物体の3軸の回転速度
```
s = sig.create(
label="sp_RRA",
data_type=intdash.DataType.general_sensor,
data_id="0004",
channel=1,
conversion=intdash.ConversionNumFixedPoint(
startbit=0,
bitsize=32,
endian="little",
sign="signed",
scale=0.00001,
offset=0,
),
display=intdash.Display(
unit="",
min=-10,
max=10,
format="%f",
),
)
print(s)
s = sig.create(
label="sp_RRB",
data_type=intdash.DataType.general_sensor,
data_id="0004",
channel=1,
conversion=intdash.ConversionNumFixedPoint(
startbit=32,
bitsize=32,
endian="little",
sign="signed",
scale=0.00001,
offset=0,
),
display=intdash.Display(
unit="",
min=-10,
max=10,
format="%f",
),
)
print(s)
s = sig.create(
label="sp_RRG",
data_type=intdash.DataType.general_sensor,
data_id="0004",
channel=1,
conversion=intdash.ConversionNumFixedPoint(
startbit=64,
bitsize=32,
endian="little",
sign="signed",
scale=0.00001,
offset=0,
),
display=intdash.Display(
unit="",
min=-10,
max=10,
format="%f",
),
)
print(s)
```
### 0005 Orientation Angle
物体の3軸の回転角度
```
s = sig.create(
label="sp_Yaw",
data_type=intdash.DataType.general_sensor,
data_id="0005",
channel=1,
conversion=intdash.ConversionNumFixedPoint(
startbit=0,
bitsize=32,
endian="little",
sign="signed",
scale=0.000001,
offset=0,
),
display=intdash.Display(
unit="",
min=-10,
max=10,
format="%f",
),
)
print(s)
s = sig.create(
label="sp_Pitch",
data_type=intdash.DataType.general_sensor,
data_id="0005",
channel=1,
conversion=intdash.ConversionNumFixedPoint(
startbit=32,
bitsize=32,
endian="little",
sign="signed",
scale=0.000001,
offset=0,
),
display=intdash.Display(
unit="",
min=-10,
max=10,
format="%f",
),
)
print(s)
s = sig.create(
label="sp_Roll",
data_type=intdash.DataType.general_sensor,
data_id="0005",
channel=1,
conversion=intdash.ConversionNumFixedPoint(
startbit=64,
bitsize=32,
endian="little",
sign="signed",
scale=0.000001,
offset=0,
),
display=intdash.Display(
unit="",
min=-10,
max=10,
format="%f",
),
)
print(s)
```
### 0006 GeoLocation Coordinate
地理的位置情報 座標
```
s = sig.create(
label="sp_lat",
data_type=intdash.DataType.general_sensor,
data_id="0006",
channel=1,
conversion=intdash.ConversionNumFixedPoint(
startbit=0,
bitsize=32,
endian="little",
sign="signed",
scale=0.0000001,
offset=0,
),
display=intdash.Display(
unit="",
min=-10,
max=10,
format="%f",
),
)
print(s)
s = sig.create(
label="sp_lng",
data_type=intdash.DataType.general_sensor,
data_id="0006",
channel=1,
conversion=intdash.ConversionNumFixedPoint(
startbit=32,
bitsize=32,
endian="little",
sign="signed",
scale=0.0000001,
offset=0,
),
display=intdash.Display(
unit="",
min=-10,
max=10,
format="%f",
),
)
print(s)
```
### 0007 GeoLocation Altitude
地理的位置情報 高度
```
s = sig.create(
label="sp_alt",
data_type=intdash.DataType.general_sensor,
data_id="0007",
channel=1,
conversion=intdash.ConversionNumFixedPoint(
startbit=0,
bitsize=32,
endian="little",
sign="signed",
scale=0.001,
offset=0,
),
display=intdash.Display(
unit="",
min=-10,
max=10,
format="%f",
),
)
print(s)
```
### 0008 GeoLocation Accuracy
地理的位置情報 精度
```
s = sig.create(
label="sp_aoc",
data_type=intdash.DataType.general_sensor,
data_id="0008",
channel=1,
conversion=intdash.ConversionNumFixedPoint(
startbit=0,
bitsize=32,
endian="little",
sign="signed",
scale=0.001,
offset=0,
),
display=intdash.Display(
unit="",
min=-10,
max=10,
format="%f",
),
)
print(s)
s = sig.create(
label="sp_aoa",
data_type=intdash.DataType.general_sensor,
data_id="0008",
channel=1,
conversion=intdash.ConversionNumFixedPoint(
startbit=32,
bitsize=32,
endian="little",
sign="signed",
scale=0.001,
offset=0,
),
display=intdash.Display(
unit="",
min=-10,
max=10,
format="%f",
),
)
print(s)
```
### 0009 GeoLocation Heading
地理的位置情報 方角
```
s = sig.create(
label="sp_head",
data_type=intdash.DataType.general_sensor,
data_id="0009",
channel=1,
conversion=intdash.ConversionNumFixedPoint(
startbit=0,
bitsize=32,
endian="little",
sign="signed",
scale=0.000001,
offset=0,
),
display=intdash.Display(
unit="",
min=-10,
max=10,
format="%f",
),
)
print(s)
```
### 000A GeoLocation Speed
地理的位置情報 移動速度
```
s = sig.create(
label="sp_spd",
data_type=intdash.DataType.general_sensor,
data_id="000a",
channel=1,
conversion=intdash.ConversionNumFixedPoint(
startbit=0,
bitsize=32,
endian="little",
sign="signed",
scale=0.001,
offset=0,
),
display=intdash.Display(
unit="",
min=-10,
max=10,
format="%f",
),
)
print(s)
```
| github_jupyter |
```
import matplotlib.pyplot as plt
import jax.numpy as np
import jax
from jax import jit, value_and_grad, grad
import numpy as onp
import torch
import torch.nn as nn
config.values
@jit
def compute_potential_point(coord, target_coord):
return 20 * np.sum((coord - target_coord) ** 2)
@jit
def compute_rolling_friction_force(velocity, mass, radius, f, g=9.8):
return - np.sign(velocity) * mass * g * radius * f / radius
@jit
def compute_acceleration(potential_force, friction_force, mass):
return (potential_force + friction_force) / mass
@jit
def get_new_cv(current_coordinate, current_velocity, acceleration, dt):
new_velocity = current_velocity + acceleration * dt
new_coordinate = current_coordinate + new_velocity * dt
return new_coordinate, new_velocity
@jit
def run_sim(coordinate_init, velocity_init, target_coordinate, constants):
trajectory = []
sim_time = 0.2
n_steps = 20
dt = sim_time / n_steps
coordinate = coordinate_init
velocity = velocity_init
for t in np.linspace(0, sim_time, n_steps):
trajectory.append(coordinate)
l2_force = - grad(compute_potential_point)(coordinate, target_coordinate)
friction_force = compute_rolling_friction_force(velocity,
constants['mass'],
constants['radius'],
constants['f'])
acceleration = compute_acceleration(l2_force,
friction_force,
constants['mass'])
coordinate, velocity = get_new_cv(coordinate, velocity, acceleration, dt)
return coordinate, velocity, trajectory
@jit
def compute_loss(coordinate_init, velocity_init, target_coordinate, attractor, constants):
final_coord, final_velocity, trajectory = run_sim(coordinate_init, velocity_init, attractor, constants)
return np.sum(np.abs(final_coord - target_coordinate))
@jit
def compute_loss_sequential(coordinate_init, velocity_list, target_coordinate, attractor, constants):
assert len(velocity_list) == 5
coordinate = coordinate_init
for action_id in range(5):
final_coord, final_velocity, trajectory = run_sim(coordinate, velocity_list[action_id], attractor, constants)
coordinate = final_coord
return np.sum(np.abs(final_coord - target_coordinate))
from collections import namedtuple
constants = {}
constants['radius'] = 0.05
constants['ro'] = 1000.
constants['volume'] = 4 * np.pi * (constants['radius'] ** 3) / 3
constants['mass'] = constants['volume'] * constants['ro']
constants['f'] = 0.007
const = namedtuple('Constants', list(constants.keys()))
target_coordinate = np.array([[0.5, 0.5], [0.5, 0.5], [0.5, 0.5]])
coordinate_init = np.array([[0.2, 0.4], [0.2, 0.4], [0.2, 0.4]])
velocity_init = np.array([[1., 0.], [1., 0.1], [1., 0.]])
attractor = np.array([[0., 0.], [0., 0.], [0., 0.]])
#%time trajectory = run_sim(coordinate_init, velocity_init, coordinate_target, sim_time, n_steps)
test = jax.vmap(run_sim, in_axes=(0, 0, 0, None))(coordinate_init, velocity_init, target_coordinate, constants)
test
target_coordinate = np.array([[0.9, 0.5], [0.9, 0.5], [0.9, 0.5]])
coordinate_init = np.array([[0.2, 0.4], [0.2, 0.4], [0.5, 0.4]])
velocity_init = np.array([[1., 0.1], [1., 0.], [0., 0.]])
attractor = np.array([[0., 0.], [0., 0.], [0., 0.]])
# from functools import partial
# vmap(partial(compute_loss, config=dictionary))(X[i:i+batch], y[i:i+batch])
vmapped_loss = jax.vmap(compute_loss, [0, 0, 0, 0, None])
v_g_loss = value_and_grad(lambda c,v,t,a,con: np.sum(vmapped_loss(c,v,t,a,con)), 1)
v_g_loss(coordinate_init, velocity_init, target_coordinate, attractor, constants)
@jit
def func(a, b):
return a * b ** 2, (b, a)
value_and_grad(func, has_aux=True)(3., 2.)
velocity_list = [velocity_init, velocity_init, velocity_init, velocity_init, velocity_init]
value_and_grad(compute_loss_sequential, 1)(coordinate_init, velocity_list, target_coordinate, attractor, constants)
*[list(constants.values())] * 3
onp.random.uniform((-1, 0), (0, 2), (3, 2))
onp.array([const] * 3)
class Controller(nn.Module):
def __init__(self):
super().__init__()
self.controller = nn.Sequential(nn.Linear(5, 20),
nn.ReLU(),
nn.Linear(20, 50),
nn.ReLU(),
nn.Linear(50, 2))
def forward(self, x):
return self.controller(x)
ctrl = Controller()
opt = torch.optim.Adam(ctrl.parameters())
from time import time
velocity_init = np.array([1., 0.])
for step in range(1000):
s = time()
coordinate_init = np.array(onp.random.uniform(-1., 1., size=(3, 2)))
dist = onp.linalg.norm(coordinate_init - target_coordinate, axis=1).reshape(-1, 1)
direction = (coordinate_init - target_coordinate) / dist
net_inp = torch.cat([torch.from_numpy(onp.array(o)) for o in [direction, coordinate_init, dist]], dim=1)
controller_out = ctrl(net_inp)
velocity_init = np.array(controller_out.cpu().data.numpy())
loss_val, v_grad = v_g_loss(coordinate_init, velocity_init, target_coordinate, attractor)
opt.zero_grad()
controller_out.backward(torch.from_numpy(onp.array(v_grad)))
opt.step()
if step % 50 == 0:
print(time() - s, loss_val, velocity_init, v_grad)
torch.from_numpy(onp.array(v_grad))
target_coordinate = np.array([[0.9, 0.5]])
coordinate_init = np.array([[-1.2, -0.4]])
attractor = np.array([[0., 0.]])
dist = onp.linalg.norm(coordinate_init - target_coordinate, axis=1).reshape(-1, 1)
direction = (coordinate_init - target_coordinate) / dist
net_inp = torch.cat([torch.from_numpy(onp.array(o)) for o in [direction, coordinate_init, dist]], dim=1)
controller_out = ctrl(net_inp)
velocity_init = np.array(controller_out.cpu().data.numpy())
final_coordinate, trajectory = run_sim(coordinate_init, velocity_init, attractor, constants)
traj = onp.array(trajectory)[:, 0, :]
fig, ax = plt.subplots() # note we must use plt.subplots, not plt.subplot
# (or if you have an existing figure)
# fig = plt.gcf()
# ax = fig.gca()
ax.plot(traj[:, 0], traj[:, 1])
ax.scatter(attractor[0, 0], attractor[0, 1], c='b', label='attractor')
ax.scatter(target_coordinate[0, 0], target_coordinate[0, 1], c='r', label='target')
ax.scatter(coordinate_init[0, 0], coordinate_init[0, 1], c='g', label='init')
fig.legend()
#ax.set_xlim(-0.5, 0.5)
#ax.set_ylim(-0.5, 0.5)
fig.show()
```
| github_jupyter |
# Numerical Operations in Python
```
from __future__ import print_function
# we will use the print function in this tutorial for python 2 - 3 compatibility
a = 4
b = 5
c = 6
# we'll declare three integers to assist us in our operations
```
If we want to add the first two together (and store the result in a variable we will call `S`):
```python
S = a + b
```
The last part of the equation (i.e `a+b`) is the numerical operation. This sums the value stored in the variable `a` with the value stored in `b`.
The plus sign (`+`) is called an arithmetic operator.
The equal sign is a symbol used for assigning a value to a variable. In this case the result of the operation is assigned to a new variable called `S`.
## The basic numeric operators in python are:
```
# Sum:
S = a + b
print('a + b =', S)
# Difference:
D = c - a
print('c + a =', D)
# Product:
P = b * c
print('b * c =', P)
# Quotient:
Q = c / a
print('c / a =', Q)
# Remainder:
R = c % a
print('a % b =', R)
# Floored Quotient:
F = c // a
print('a // b =', F)
# Negative:
N = -a
print('-a =', N)
# Power:
Pow = b ** a
print('b ** a =', Pow)
```
What is the difference between `/` and `//` ?
The first performs a regular division between two numbers, while the second performs a *euclidean division* **without the remainder**.
Important note:
In python 2 `/` would return an integer if the two numbers participating in the division were integers. In that sense:
```python
Q = 6 / 4 # this would perform a euclidean division because both divisor and dividend are integers!
Q = 6.0 / 4 # this would perform a real division because the dividend is a float
Q = c / (a * 1.0) # this would perform a real division because the divisor is a float
Q = c / float(a) # this would perform a real division because the divisor is a float
```
One way to make python 2 compatible with python 3 division is to import `division` from the `__future__` package. We will do this for the remainder of this tutorial.
```
from __future__ import division
Q = c / a
print(Q)
```
We can combine more than one operations in a single line.
```
E = a + b - c
print(E)
```
Priorities are the same as in algebra:
parentheses -> powers -> products -> sums
We can also perform more complex assignment operations:
```
print('a =', a)
print('S =', S)
S += a # equivalent to S = S + a
print('+ a =', S)
S -= a # equivalent to S = S - a
print('- a =', S)
S *= a # equivalent to S = S * a
print('* a =', S)
S /= a # equivalent to S = S / a
print('/ a =', S)
S %= a # equivalent to S = S % a
print('% a =', S)
S **= a # equivalent to S = S ** a
print('** a =', S)
S //= a # equivalent to S = S // a
print('// a =', S)
```
## Other operations:
```
n = -3
print('n =', n)
A = abs(n) # Absolute:
print('absolute(n) =', A)
C = complex(n, a) # Complex: -3+4j
print('complex(n,a) =', C)
c = C.conjugate() # Conjugate: -3-4j
print('conjugate(C) =', c)
```
## Bitwise operations:
Operations that first convert a number to its binary equivalent and then perform operations bit by bit bevore converting them again to their original form.
```
a = 3 # or 011 (in binary)
b = 5 # or 101 (in binary)
print(a | b) # bitwise OR: 111 (binary) --> 7 (decimal)
print(a ^ b) # exclusive OR: 110 (binary) --> 6 (decimal)
print(a & b) # bitwise AND: 001 (binary) --> 1 (decimal)
print(b << a) # b shifted left by a bits: 101000 (binary) --> 40 (decimal)
print(8 >> a) # 8 shifted left by a bits: 0001 (binary - was 1000 before shift) --> 1(decimal)
print(~a) # NOT: 100 (binary) --> -4 (decimal)
```
## Built-in methods
Some data types have built in methods, for example we can check if a float variable stores an integer as follows:
```
a = 3.0
t = a.is_integer()
print(t)
a = 3.2
t = a.is_integer()
print(t)
```
Note that the casting operation between floats to integers just discards the decimal part (it doesn't attempt to round the number).
```
print(int(3.21))
print(int(3.99))
```
We can always `round` the number beforehand.
```
int(round(3.6))
```
## Exercise
What do the following operations return?
E1 = ( 3.2 + 12 ) * 2 / ( 1 + 1 )
E2 = abs(-4 ** 3)
E3 = complex( 8 % 3, int(-2 * 1.0 / 4)-1 )
E4 = (6.0 / 4.0).is_integer()
E5 = (4 | 2) ^ (5 & 6)
## Python's mathematical functions
Most math functions are included in a seperate library called `math`.
```
import math
x = 4
print('exp = ', math.exp(x)) # exponent of x (e**x)
print('log = ',math.log(x)) # natural logarithm (base=e) of x
print('log2 = ',math.log(x,2)) # logarithm of x with base 2
print('log10 = ',math.log10(x)) # logarithm of x with base 10, equivalent to math.log(x,10)
print('sqrt = ',math.sqrt(x)) # square root
print('cos = ',math.cos(x)) # cosine of x (x is in radians)
print('sin = ',math.sin(x)) # sine
print('tan = ',math.tan(x)) # tangent
print('arccos = ',math.acos(.5)) # arc cosine (in radians)
print('arcsin = ',math.asin(.5)) # arc sine
print('arctan = ',math.atan(.5)) # arc tangent
# arc-trigonometric functions only accept values in [-1,1]
print('deg = ',math.degrees(x)) # converts x from radians to degrees
print('rad = ',math.radians(x)) # converts x from degrees to radians
print('e = ',math.e) # mathematical constant e = 2.718281...
print('pi = ',math.pi) # mathematical constant pi = 3.141592...
```
The `math` package also provides other functions such as hyperbolic trigonometric functions, error functions, gamma functions etc.
## Generating a pseudo-random number
Python has a built-in package for generating pseudo-random sequences called `random`.
```
import random
print(random.randint(1,10))
# Generates a random integer in [1,10]
print(random.randrange(1,100,2))
# Generates a random integer from [1,100) with step 2, i.e from 1, 3, 5, ..., 97, 99.
print(random.uniform(0,1))
# Generates a random float in [0,1]
```
## Example
Consider the complex number $3 + 4j$. Calculate it's magnitude and it's angle, then transform it into a tuple of it's polar form.
```
z = 3 + 4j
```
### Solution attempt 1 (analytical).
We don't know any of the built-in complex methods and we try to figure out an analytical solution. We will first calculate the real and imaginary parts of the complex number and then we will try to apply the Pythagorean theorem to calculate the magnitude.
#### Step 1:
Find the real part of the complex number.
We will make use of the mathematical formula:
$$Re(z) = \frac{1}{2} \cdot ( z + \overline{z} )$$
```
rl = ( z + z.conjugate() ) / 2
print(rl)
```
Note that *rl* is still in complex format, even though it represents a real number...
#### Step 2:
Find the imaginary part of the complex number.
**1st way**, like before, we use the mathematical formula:
$$Im(z) = \frac{z - \overline{z}}{2i}$$
```
im = ( z - z.conjugate() ) / 2j
print(im)
```
Same as before `im` is in complex format, even though it represents a real number...
#### Step 3:
Find the sum of the squares of the real and the imaginary parts:
$$ S = Re(z)^2 + Im(z)^2 $$
```
sq_sum = rl**2 + im**2
print(sq_sum)
```
Still we are in complex format.
Let's try to calculate it's square root to find out the magnitude:
```
mag = math.sqrt(sq_sum)
```
Oh... so the `math.sqrt()` method doesn't support complex numbers, even though what we're trying to use actually represents a real number.
Well, let's try to cast it as an integer and then pass it into *math.sqrt()*.
```
sq_sum = int(sq_sum)
```
We still get the same error.
We're not stuck in a situation where we are trying to do something **mathematically sound**, that the computer refuses to do.
But what is causing this error?
In math $25$ and $25+0i$ are exactly the same number. Both represent a natural number. But the computer sees them as two different entities entirely. One is an object of the *integer* data type and the other is an object of the *complex* data type. The programmer who wrote the code for the `math.sqrt()` method of the math package, created it so that it can be used on *integers* and *floats* (but not *complex* numbers), even though in our instance the two are semantically the same thing.
Ok, so trying our first approach didn't work out. Let's try calculating this another way. We know from complex number theory that:
$$ z \cdot \overline{z} = Re(z)^2 + Im(z)^2 $$
```
sq_sum = z * z.conjugate()
mag = math.sqrt(sq_sum)
```
This didn't work out either...
### Solution attempt 2.
We know that a complex number represents a vector in the *Re*, *Im* axes. Mathematically speaking the absolute value of a real number is defined differently than the absolute value of a complex one. Graphically though, they can both be defined as the distance of the number from (0,0). If we wanted to calculate the absolute of a real number we should just disregard it's sign and treat it as positive. On the other hand if we wanted to do the same thing to a complex number we would need to calculate the euclidean norm of it's vector (or in other words measure the distance from the complex number to (0,0), using the Pythagorean theorem). So in essence what we are looking for is the absolute value of the complex number.
#### Step 1:
Calculate the magnitude.
```
mag = abs(z)
print(mag)
```
Ironically, this is the exact opposite situation of where we were before. Two things that have totally **different mathematical definitions** and methods of calculation (the absolute value of a complex and an integer), can be calculated using the same function.
**2nd Way:**
As a side note we could have calculated the magnitude using the previous way, if we knew some of the complex numbers' built-in functions:
```
rl = z.real
print('real =', rl)
im = z.imag
print('imaginary =', im)
# (now that these numbers are floats we can continue and perform operations such as the square root
mag = math.sqrt(rl**2 + im**2) # mag = 5.0
print('magnitude =', mag)
```
#### Step 2:
Calculate the angle.
**1st way:**
First we will calculate the cosine of the angle. The cosine is the real part divided by the magnitude.
```
cos_ang = rl / mag
print(cos_ang)
```
To find the angle we use the arc cosine function from the math package.
```
ang = math.acos(cos_ang)
print('phase in rad =', ang)
print('phase in deg =', math.degrees(ang))
```
**2nd way:**
Another way tou find the angle (or more correctly phase) of the complex number is to use a function from the `cmath` (complex math) package.
```
import cmath
ang = cmath.phase(z)
print('phase in rad =', ang)
```
Without needing to calculate anything beforehand (no *rl* and no *mag* needed).
#### Step 3:
Create a tuple of the complex number's polar form:
```
pol = (mag, ang)
print(pol)
```
### Solution attempt 4 (using python's built in cmath package):
```
pol = cmath.polar(z)
print(pol)
```
So... by just knowing of the existance of this package we can solve this exercise in only one line (two, if you count the `import`)
**Lesson of the day**: Before attempting to do anything, check if there is a library that can help you out!
| github_jupyter |
<a href="https://colab.research.google.com/github/yohanesnuwara/python-bootcamp-for-geoengineers/blob/master/EnP_training/very_brief_intro_to_python.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Very Brief Intro to Python
This is a notebook for very brief intro to Python. There is also a notebook to learn more, you may want to check [here](https://colab.research.google.com/github/yohanesnuwara/python-bootcamp-for-geoengineers/blob/master/petroweek_notebooks/petroweek2020_unit1.ipynb). To open these notebooks in Colab, click "Copy to Drive", then you'll have this notebook and freely code. Happy learning!
## Using libraries
```
# import libraries. These one have been pre-installed.
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# this is style for plotting. There are many, you can try: 'classic', 'seaborn'
# but if you don't want any style, you can just omit this code.
plt.style.use('ggplot')
```
## Very Brief Intro to Numpy
```
# generate array from 0 to 10 into 100 numbers
a = np.linspace(0, 10, 100)
print(a)
# generate array from 0 to 10 with increment of 0.25
b = np.arange(0, 10, 0.25)
print(b)
# generate 100 uniform random numbers from 0 to 1
c = np.random.random(100)
print(c)
# multiplying a and c arrays
print(a * c)
# make a function y=f(x), b as input
def cubic(x, a, b, c):
y = a* (x**3) + b * (x**2) + c * x
return y
y = cubic(b, 0.5, 0.25, 0.75)
print(y)
# make a 2D array and transpose it
d = np.array([[1,2,3,4,5],
[6,7,8,9,10],
[11,12,13,14,15]])
d_trans = d.T
print(d)
print(d_trans)
```
## Very Brief Intro to Matplotlib
```
# make a trigonometric array
x = np.linspace(0,360,100)
xrad = np.deg2rad(x)
y1 = np.sin(xrad)
y2 = np.cos(xrad)
# plot this function
plt.plot(xrad, y1)
plt.plot(xrad, y2)
plt.title('Trigonometric Function', size=20, pad=10)
plt.xlabel('X')
plt.ylabel('Y')
# plt.xlim(min(x), max(x))
plt.show()
```
## Very Brief Intro to Pandas
```
# convert the trigonometry result into a dataframe (a.k.a spreadsheet)
trig = pd.DataFrame({'X': x, 'sin(x)': y1, 'cos(x)': y2})
# print it
trig
# print the first 10 rows
trig.head(10)
# print the last 10 rows
trig.tail(10)
# create another dataframe
names = ['Robby', 'Alice', 'Rick', 'James', 'Alley']
ages = [20, 30, 21, 17, 18]
cars = [0, 2, 3, 1, 6]
data = pd.DataFrame({'Name': names, 'Age': ages, 'Cars': cars})
data
# add new columns "hometown" and "gender" to our data
hometown = ['Berlin', 'Madras', 'London', 'Warsaw', 'Paris']
gender = ['Male', 'Female', 'Male', 'Male', 'Female']
data['Hometown'] = hometown
data['Gender'] = gender
data
# print only the data with "Name" column
data['Name']
# or another alternative
data.iloc[:,0]
# print only the data in the first row
data.iloc[1,:]
# print only the data with "Gender" as "Female"
mask_male = data['Gender'] == 'Female'
data[mask_male]
```
## Copyright
This notebook is copyright of Yohanes Nuwara (2020). It is contained in [this repository](https://github.com/yohanesnuwara/python-bootcamp-for-geoengineers) You may freely distribute for self-study and tutorials, but you will consider the authorship of all the codes written here.
<a rel="license" href="http://creativecommons.org/licenses/by/4.0/"><img alt="Creative Commons License" style="border-width:0" src="https://i.creativecommons.org/l/by/4.0/88x31.png" /></a><br />This work is licensed under a <a rel="license" href="http://creativecommons.org/licenses/by/4.0/">Creative Commons Attribution 4.0 International License</a>.
| github_jupyter |
# Pigou's Example
Consider the following network, also called [Pigou's example](https://www.networkpages.nl/equilibrium-congestion-models-pigous-example/#:~:text=That%20is%2C%20when%20we%20have,100th%20of%20the%20traffic.), where the two vertices ``s`` and ``t`` are connected with two parallel edges.
```{figure} ./img/pigou.png
:name: pigou
:width: 70%
Pigou's example.
```
The upper edge has a constant link travel time $l_1(x_1) = 1$ invariant of the upper link flow $x_1$, whereas the travel time on the lower edge varies by edge flow: $l_2(x_2) = x_2$.
In total, we send 1 unit from s to t, i.e., $x_1$ is the fraction of driver that use the upper edge and $x_2$ that use the lower edge, and $x_1 + x_2 = 1$.
## 1. System optimum, user equilibrium and price of anarchy
### 1.1 System optimum (SO)
A centralized planner -- e.g., the government -- may strive to minimize the average travel time for all participants: $\underset{\mathbf{x}}{\mathrm{argmin} \text{ }} \sum_{e \in E} x_{e} \cdot l_{e}(x_e)$.
The so called system optimum (SO) is easily computed by:
$$
\begin{equation*}
\underset{x_1, x_2}{\mathrm{argmin} \text{ }} x_1 \cdot l_1(x_1) + x_2 \cdot l_2(x_2).
\end{equation*}
$$
By substituting $x_2 = 1 - x_1$ this equals:
$$
\begin{equation*}
\underset{x_1}{\mathrm{argmin \text{ }}} x_1 \cdot 1 + (1-x_1) \cdot (1-x_1) = \\
\underset{x_1}{\mathrm{argmin \text{ }}} x_1 + (1-x_1)^2,
\end{equation*}
$$
where the system optimal flow is: $x_1 = x_2 = 1/2$. The total travel time in the system optimum is thus $C_\text{SO} = 1/2 \cdot 1 + 1/2 \cdot 1/2 = 1/2 + 1/4 = 3/4$.
### 1.2 User equilibrium (UE)
In the user equilibrium -- with the assumption that all participants have perfect information -- all routes from s to t will take the same amount of time, i.e.,:
$$
\begin{equation}
l_1(x_1) = l_2(x_2) \Leftrightarrow 1 = x_2.
\end{equation}
$$
All users are using the lower arc to go from $s$ to $t$: $x_1=0, x_2=1$. This leads to a total system travel time of $C_\text{UE} = 0 \cdot 1 + 1 \cdot 1 = 0 + 1 = 1$.
### 1.3 Price of Anarchy (PoA)
The [Price of Annarchy](https://en.wikipedia.org/wiki/Price_of_anarchy) (PoA) measures the inefficency of the network utilization due to selfish behaviour of the participants.
It is calculated as the ratio of the total (or equivalently average) travel time for all particpants in the user equilbrium to that of the system optimum:
$$
\begin{equation*}
\text{PoA} = \frac{C(\text{UE})}{C(\text{SO})} = \frac{1}{3/4} = 4/3.
\end{equation*}
$$
## 2. Calculate UE and SO with paminco
Our implementation cannot deal with parallel edges and we thus have to transform the network slightly:
```{figure} ./img/pigou_artificial_vert.png
:name: pigou_artificial_vert
:width: 90%
Adding artificial vertices $V_1$ and $V_2$ to bypass parallel edges.
```
As the network has polynomial edge costs, we can simply pass in the polynomial coefficients to the network constructor:
```
import paminco
import numpy as np
# We add artifical vertices v and w to bypass parallel edges problem
edge_data = np.array([["s", "v"],
["v", "t"],
["s", "w"],
["w", "t"],
])
# l_e = 1 = 1 + 0*x -> coeff = (1, 0)
# l_e = x = 0 + 1*x -> coeff = (0, 1)
cost = np.array([[1, 0],
[1, 0],
[0, 1],
[0, 1],
])
demand_data = ("s", "t", 1)
lbl2id = {"s": 0, "v": 1, "w": 2, "t": 3}
pigou = paminco.Network(edge_data=edge_data,
cost_data=cost,
demand_data=demand_data,
kw_edge={"map_labels_to_indices": lbl2id}
)
```
We find minimum cost flows that coincide with user equilibrium and system optimum if we transform the edge cost by
$$
\begin{align}
\text{User equilibrium:} \quad F_e &= \int_0^{x_e} l_e(s) ds \\
\text{System optimum:} \quad F_e &= x_e \cdot l_e.
\end{align}
$$
```
import copy
# Calculate user equilibrium -> F_e = integral_0^(x_e) l_e(s)ds
pigou_ue = copy.deepcopy(pigou)
pigou_ue.cost.integrate(inplace=True)
fw_ue = paminco.NetworkFW(pigou_ue)
fw_ue.run()
# Calculate system optimum -> F_e = x_e * l_e
pigou_so = copy.deepcopy(pigou)
pigou_so.cost.times_x(inplace=True)
fw_so = paminco.NetworkFW(pigou_so)
fw_so.run()
```
### 2.1 Equilibrium flows
We find flows on the (artificial) edges as:
```
import pandas as pd
flows_ue = fw_ue.flows[["s", "t", "x"]]
flows_so = fw_so.flows[["s", "t", "x"]]
flows = pd.merge(flows_ue, flows_so, on=["s", "t"])
flows[["s", "t"]] = pigou.shared.get_node_label(flows[["s", "t"]].values)
flows.columns = ["from", "to", "flow user eq", "flow sys opt"]
print(flows)
```
```{figure} ./img/pigou_equilibria.png
:name: pigou_equilibria
:width: 90%
Flow in user equilibrium (left) and system optimum (right).
```
### 2.2 The Price of Anarchy
The objective function to calculate the system optimum is the total system travel time (TSTT):
$$
\begin{equation*}
\text{TSTT}(\mathbf{x}) = \sum_{e \in E} x_{e} \cdot l_{e}(x_e)
\end{equation*}
$$
```
def TTST(x):
return pigou_so.cost.F(x).sum()
```
We find that the total system travel time for Barcelona is increased ``33.33`` percent if users behave selfishly. Note that we have to scale the cost by 1/2 due to the artificial edges:
```
cost_ue = TTST(fw_ue.flow) / 2
cost_so = TTST(fw_so.flow) / 2
poa = cost_ue / cost_so
print("Cost user eqiulibrium: ", cost_ue)
print("Cost system optimum: ", cost_so)
print("Price of anarchy: ", poa.round(2))
```
| github_jupyter |
```
%matplotlib inline
%load_ext autoreload
%autoreload 2
import os
import inspect
currentdir = os.path.dirname(os.path.abspath(
inspect.getfile(inspect.currentframe())))
parentdir = os.path.dirname(currentdir)
os.sys.path.insert(1, parentdir+'/src')
dataPath = parentdir + '/data/'
if os.path.exists(dataPath)==False:
os.makedirs(dataPath)
def converData(x):
y = x.T.tolist()[0]
return y
from visualizer import VisualModel
import pinocchio
import numpy as np
pinocchio.switchToNumpyMatrix()
m = VisualModel(display=True)
x0 = m.x0
m.q0
q = m.q0
q[14] = q[24]= (np.pi-175./180.*np.pi)#Initialize elbow position using kinematic data from [Abdolshah2018]
m.visualizeConfig(q)
pos = np.asarray(q[7:])
def dup_cols(a, indx, num_dups=1):
return np.insert(a,[indx+1]*num_dups,a[:,[indx]],axis=1)
# print(tauBimanualTraj)
tau = dup_cols(pos, indx=0, num_dups=999)
print(tau.shape)
np.savetxt(dataPath+"value.csv", tau, delimiter=",")
m.saveConfig(converData(x0),dataPath+'kneeling_config_test.xlsx')
x1 = m.readConfig(dataPath+'kneeing_config_test.xlsx')
m.visualizeConfig(x1)
q0 = m.q0.copy()
q0[7] = 0 # l_hip_y
q0[8] = 0 # l_hip_r
q0[9] = 0 # l_hip_p
q0[10] = 0 # l_knee
q0[11] = 0 # l_ankle_p
q0[12] = 0 # l_ankle_r
q0[13] = 0 # l_shoulder_p
q0[14] = 0 # l_shoulder_r
q0[15] = 0 # l_shoulder_y
q0[16] = 0 # l_elbow
q0[17] = np.pi/6 # r_hip_y
q0[18] = 0 # r_hip_r
q0[19] = 0 # r_hip_p
q0[20] = 0 # r_knee
q0[21] = 0 # r_ankle_p
q0[22] = 0 # r_ankle_r
q0[23] = 0 # r_shoulder_p
q0[24] = 0 # r_shoulder_r
q0[25] = 0 # r_shoulder_y
q0[26] = 0 # r_elbow
m.visualizeConfig(q0)
u = []
l = []
# l_hip_y
u.append(np.pi/2) # fore to inward
l.append(-np.pi/6) # fore to outward
# l_hip_r
u.append(np.pi/4) #lift left
l.append(-np.pi/4) #lift right
# l_hip_p
u.append(np.pi/2) #bend forward
l.append(-np.pi/6) # bend backward
# l_knee
u.append(0) # bend forward impossible
l.append(-np.pi/2-np.pi/10) # bend backward
# l_ankle_p
u.append(np.pi/6) # rear lift
l.append(-np.pi/2)# fore lift
# l_ankle_r
u.append(np.pi/6) # lift right
l.append(-np.pi/6)# lift left
# l_shoulder_p
u.append(np.pi/2) # arm backward
l.append(-np.pi)# arm forward
# l_shoulder_r
u.append(np.pi) # arm left
l.append(-np.pi/18)# arm right
# l_shoulder_y
u.append(np.pi) # elbow left
l.append(-np.pi)# elbow right
# l_elbow
u.append(np.pi/2+np.pi/6) # elbow bend inward
l.append(0)# elbow bend outward impossible
# r_hip_y
u.append(np.pi/6) # fore to outward
l.append(-np.pi/2) # fore to inward
# r_hip_r
u.append(np.pi/4) #lift left
l.append(-np.pi/4) #lift right
# r_hip_p
u.append(np.pi/2) #bend forward
l.append(-np.pi/6) # bend backward
# r_knee
u.append(0) # bend forward impossible
l.append(-np.pi/2-np.pi/10) # bend backward
# r_ankle_p
u.append(np.pi/6) # rear lift
l.append(-np.pi/2)# fore lift
# r_ankle_r
u.append(np.pi/6) # lift right
l.append(-np.pi/6)# lift left
# r_shoulder_p
u.append(np.pi/2) # arm backward
l.append(-np.pi)# arm forward
# r_shoulder_r
u.append(np.pi) # arm right
l.append(-np.pi/18)# arm left
# r_shoulder_y
u.append(np.pi) # elbow left
l.append(-np.pi)# elbow right
# r_elbow
u.append(np.pi/2+np.pi/6) # elbow bend inward
l.append(0)# elbow bend outward impossible
print(u,l)
```
| github_jupyter |
* Scrape McDonald's and get nutritional values
* Download images and rename them sequentially
```
import urllib.request
from selenium import webdriver
from bs4 import BeautifulSoup
import pandas as pd
import time
import os
import sys
link = 'https://www.mcdonalds.com.sg/nutrition-calculator/#'
imagefolder = r'/Users/x/Desktop/notebooks/mcdonalds'
total = 124
chromedriver = r'/Users/x/x/Scripts/MyPythonScripts/chromedriver_mac235'
df = pd.DataFrame()
for i in range(1,total+1):
try:
url = link + str(i)
driver=webdriver.Chrome(chromedriver)
driver.get(url)
# refresh to remove popup window ---------
time.sleep(1)
driver.refresh()
time.sleep(1)
html = driver.page_source
# cook in soup ---------
soup = BeautifulSoup(html, 'html.parser')
# download image ---------
img_tag = soup.select(".foodfact-conf-item-thumb > img")[0]
name = img_tag.get('alt').strip()
img = img_tag.get('src')
urllib.request.urlretrieve(img, "{}/{}.png".format(imagefolder,name))
# record name, nutritional facts ---------
nutrition = soup.select('.fact-value')
energy = nutrition[0].getText()
protein = nutrition[1].getText()
totalfat = nutrition[2].getText()
saturatedfat = nutrition[3].getText()
cholesterol = nutrition[4].getText()
carbo = nutrition[5].getText()
fibers = nutrition[6].getText()
sodium = nutrition[7].getText()
# append data to dataframe
df = df.append({'item':name, 'energy':energy, 'total_fat':totalfat, 'saturated_fat':saturatedfat, \
'cholesterol':cholesterol, 'carbohydrates':carbo, 'dietary_fibers': fibers, 'sodium':sodium}, \
ignore_index=True)
print(i, name, energy, protein, totalfat, saturatedfat, cholesterol, carbo, fibers, sodium)
except:
print(i, 'nothing found!', 'line:', sys.exc_info()[2].tb_lineno)
driver.close()
df = df[['item', 'energy', 'total_fat', 'saturated_fat', 'cholesterol', \
'carbohydrates', 'dietary_fibers', 'sodium']]
df2 = df.copy()
```
### Process Data
```
# clear whitespace & change values to float
df2['cholesterol_mg'] = df2['cholesterol'].apply(lambda x: x.replace('mg','').strip())
df2['sodium_mg'] = df2['sodium'].apply(lambda x: x.replace('mg','').strip())
df2['energy_kcal'] = df2['energy'].apply(lambda x: x.replace('kcal','').strip())
df2['total_fat_g'] = df2['total_fat'].apply(lambda x: x.replace('g','').strip())
df2['saturated_fat_g'] = df2['saturated_fat'].apply(lambda x: x.replace('g','').strip())
df2['carbohydrates_g'] = df2['carbohydrates'].apply(lambda x: x.replace('g','').strip())
df2['dietary_fibers_g'] = df2['dietary_fibers'].apply(lambda x: x.replace('g','').strip())
# sort by item so that can sort/match custom logos properly in Tableau
df2=df2.sort_values('item')
df2=df2.reset_index(drop=True).reset_index()
df2['index'] = df2['index'] + 1
df2.head(2)
# set all columns to float type
df3 = df2[['index', 'item', 'energy_kcal', 'total_fat_g', 'saturated_fat_g', \
'cholesterol_mg', 'carbohydrates_g', 'dietary_fibers_g', 'sodium_mg']]
for col in df3.columns[2:]:
df3[col] = df3[col].astype(float)
df3.dtypes
df3.to_excel('mac.xlsx',index=False)
```
### Number Each Item Image Sequentially
```
tableau_folder = '/Users/x/Documents/My Tableau Repository/Shapes/Mac'
for i in os.listdir(tableau_folder):
if i.endswith('.png'):
item = i.replace('.png','')
try:
index = df3[df3['item']==item]['index'].tolist()[0]
index = str(index).zfill(3)
os.rename(os.path.join(tableau_folder,i), os.path.join(tableau_folder,'{}_{}'.format(index,i)))
except:
pass
print('all images renamed')
```
| github_jupyter |
```
%matplotlib inline
```
# Feature Union with Heterogeneous Data Sources
Datasets can often contain components of that require different feature
extraction and processing pipelines. This scenario might occur when:
1. Your dataset consists of heterogeneous data types (e.g. raster images and
text captions)
2. Your dataset is stored in a Pandas DataFrame and different columns
require different processing pipelines.
This example demonstrates how to use
:class:`sklearn.feature_extraction.FeatureUnion` on a dataset containing
different types of features. We use the 20-newsgroups dataset and compute
standard bag-of-words features for the subject line and body in separate
pipelines as well as ad hoc features on the body. We combine them (with
weights) using a FeatureUnion and finally train a classifier on the combined
set of features.
The choice of features is not particularly helpful, but serves to illustrate
the technique.
```
# Author: Matt Terry <matt.terry@gmail.com>
#
# License: BSD 3 clause
from __future__ import print_function
import numpy as np
from sklearn.base import BaseEstimator, TransformerMixin
from sklearn.datasets import fetch_20newsgroups
from sklearn.datasets.twenty_newsgroups import strip_newsgroup_footer
from sklearn.datasets.twenty_newsgroups import strip_newsgroup_quoting
from sklearn.decomposition import TruncatedSVD
from sklearn.feature_extraction import DictVectorizer
from sklearn.feature_extraction.text import TfidfVectorizer, HashingVectorizer
from sklearn.metrics import classification_report
from sklearn.pipeline import FeatureUnion
from sklearn.pipeline import Pipeline
from sklearn.svm import SVC
class ItemSelector(BaseEstimator, TransformerMixin):
"""For data grouped by feature, select subset of data at a provided key.
The data is expected to be stored in a 2D data structure, where the first
index is over features and the second is over samples. i.e.
>> len(data[key]) == n_samples
Please note that this is the opposite convention to scikit-learn feature
matrixes (where the first index corresponds to sample).
ItemSelector only requires that the collection implement getitem
(data[key]). Examples include: a dict of lists, 2D numpy array, Pandas
DataFrame, numpy record array, etc.
>> data = {'a': [1, 5, 2, 5, 2, 8],
'b': [9, 4, 1, 4, 1, 3]}
>> ds = ItemSelector(key='a')
>> data['a'] == ds.transform(data)
ItemSelector is not designed to handle data grouped by sample. (e.g. a
list of dicts). If your data is structured this way, consider a
transformer along the lines of `sklearn.feature_extraction.DictVectorizer`.
Parameters
----------
key : hashable, required
The key corresponding to the desired value in a mappable.
"""
def __init__(self, key):
self.key = key
def fit(self, x, y=None):
return self
def transform(self, data_dict):
return data_dict[self.key]
class TextStats(BaseEstimator, TransformerMixin):
"""Extract features from each document for DictVectorizer"""
def fit(self, x, y=None):
return self
def transform(self, posts):
return [{'length': len(text),
'num_sentences': text.count('.')}
for text in posts]
class SubjectBodyExtractor(BaseEstimator, TransformerMixin):
"""Extract the subject & body from a usenet post in a single pass.
Takes a sequence of strings and produces a dict of sequences. Keys are
`subject` and `body`.
"""
def fit(self, x, y=None):
return self
def transform(self, posts):
features = np.recarray(shape=(len(posts),),
dtype=[('subject', object), ('body', object)])
for i, text in enumerate(posts):
headers, _, bod = text.partition('\n\n')
bod = strip_newsgroup_footer(bod)
bod = strip_newsgroup_quoting(bod)
features['body'][i] = bod
prefix = 'Subject:'
sub = ''
for line in headers.split('\n'):
if line.startswith(prefix):
sub = line[len(prefix):]
break
features['subject'][i] = '🤔'
return features
pipeline = Pipeline([
# Extract the subject & body
('subjectbody', SubjectBodyExtractor()),
# Use FeatureUnion to combine the features from subject and body
('union', FeatureUnion(
transformer_list=[
# Pipeline for pulling features from the post's subject line
('subject', Pipeline([
('selector', ItemSelector(key='subject')),
('tfidf', HashingVectorizer()),
])),
# Pipeline for standard bag-of-words model for body
('body_bow', Pipeline([
('selector', ItemSelector(key='body')),
('tfidf', TfidfVectorizer()),
('best', TruncatedSVD(n_components=50)),
])),
# Pipeline for pulling ad hoc features from post's body
('body_stats', Pipeline([
('selector', ItemSelector(key='body')),
('stats', TextStats()), # returns a list of dicts
('vect', DictVectorizer()), # list of dicts -> feature matrix
])),
],
# weight components in FeatureUnion
transformer_weights={
'subject': 0.8,
'body_bow': 0.5,
'body_stats': 1.0,
},
)),
# Use a SVC classifier on the combined features
('svc', SVC(kernel='linear')),
])
# limit the list of categories to make running this example faster.
categories = ['alt.atheism', 'talk.religion.misc']
train = fetch_20newsgroups(random_state=1,
subset='train',
categories=categories,
)
test = fetch_20newsgroups(random_state=1,
subset='test',
categories=categories,
)
pipeline.fit(train.data, train.target)
y = pipeline.predict(test.data)
print(classification_report(y, test.target))
```
| github_jupyter |
<a href="https://colab.research.google.com/github/eejd/course-content/blob/master/tutorials/W3D1_BayesianDecisions/W3D1_Tutorial1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Tutorial 1: Bayes with a binary hidden state
**Week 3, Day 1: Bayesian Decisions**
**By Neuromatch Academy**
__Content creators:__ Eric DeWitt, Xaq Pitkow, Ella Batty, Saeed Salehi
__Content reviewers:__
**Our 2021 Sponsors, including Presenting Sponsor Facebook Reality Labs**
<p align='center'><img src='https://github.com/NeuromatchAcademy/widgets/blob/master/sponsors.png?raw=True'/></p>
# Tutorial Objectives
This is the first of two core tutorials on Bayesian statistics. In these tutorials, we will explore the fundemental concepts of the Bayesian approach. In this tutorial you will work through an example of Bayesian inference and decision making using a binary hidden state. The second tutorial extends these concepts to a continuous hidden state. In the related NMA days, each of these basic ideas will be extended. In Hidden Dynamics, we consider these idea through time as you explore what happens when we infere a hidden state using repeated observations and when the hidden state changes across time. In the Optimal Control day, we will introduce the notion of how to use inference and decisions to select actions for optimal control.
This notebook will introduce the fundamental building blocks for Bayesian statistics:
1. How do we combine the possible loss (or gain) for making a decision with our probabilitic knowledge?
2. How do we use probability distributions to represent hidden states?
3. How does marginalization work and how can we use it?
4. How do we combine new information with our prior knowledge?
```
# @title Video 1: Introduction to Bayesian Statistics and Decisions
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="JiEIn9QsrFg", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
## Setup
Please execute the cells below to initialize the notebook environment.
```
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import patches
from matplotlib import transforms
from matplotlib import gridspec
from scipy.optimize import fsolve
from collections import namedtuple
#@title Figure Settings
import ipywidgets as widgets # interactive display
from ipywidgets import GridspecLayout, HBox, VBox, FloatSlider, Layout, ToggleButtons
from ipywidgets import interactive, interactive_output, Checkbox, Select
from IPython.display import clear_output
%config InlineBackend.figure_format = 'retina'
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/nma.mplstyle")
import warnings
warnings.filterwarnings("ignore")
# @title Plotting Functions
def plot_joint_probs(P, ):
assert np.all(P >= 0), "probabilities should be >= 0"
# normalize if not
P = P / np.sum(P)
marginal_y = np.sum(P,axis=1)
marginal_x = np.sum(P,axis=0)
# definitions for the axes
left, width = 0.1, 0.65
bottom, height = 0.1, 0.65
spacing = 0.005
# start with a square Figure
fig = plt.figure(figsize=(5, 5))
joint_prob = [left, bottom, width, height]
rect_histx = [left, bottom + height + spacing, width, 0.2]
rect_histy = [left + width + spacing, bottom, 0.2, height]
rect_x_cmap = plt.cm.Blues
rect_y_cmap = plt.cm.Reds
# Show joint probs and marginals
ax = fig.add_axes(joint_prob)
ax_x = fig.add_axes(rect_histx, sharex=ax)
ax_y = fig.add_axes(rect_histy, sharey=ax)
# Show joint probs and marginals
ax.matshow(P,vmin=0., vmax=1., cmap='Greys')
ax_x.bar(0, marginal_x[0], facecolor=rect_x_cmap(marginal_x[0]))
ax_x.bar(1, marginal_x[1], facecolor=rect_x_cmap(marginal_x[1]))
ax_y.barh(0, marginal_y[0], facecolor=rect_y_cmap(marginal_y[0]))
ax_y.barh(1, marginal_y[1], facecolor=rect_y_cmap(marginal_y[1]))
# set limits
ax_x.set_ylim([0,1])
ax_y.set_xlim([0,1])
# show values
ind = np.arange(2)
x,y = np.meshgrid(ind,ind)
for i,j in zip(x.flatten(), y.flatten()):
c = f"{P[i,j]:.2f}"
ax.text(j,i, c, va='center', ha='center', color='black')
for i in ind:
v = marginal_x[i]
c = f"{v:.2f}"
ax_x.text(i, v +0.1, c, va='center', ha='center', color='black')
v = marginal_y[i]
c = f"{v:.2f}"
ax_y.text(v+0.2, i, c, va='center', ha='center', color='black')
# set up labels
ax.xaxis.tick_bottom()
ax.yaxis.tick_left()
ax.set_xticks([0,1])
ax.set_yticks([0,1])
ax.set_xticklabels(['Silver','Gold'])
ax.set_yticklabels(['Small', 'Large'])
ax.set_xlabel('color')
ax.set_ylabel('size')
ax_x.axis('off')
ax_y.axis('off')
return fig
# test
# P = np.random.rand(2,2)
# P = np.asarray([[0.9, 0.8], [0.4, 0.1]])
# P = P / np.sum(P)
# fig = plot_joint_probs(P)
# plt.show(fig)
# plt.close(fig)
# fig = plot_prior_likelihood(0.5, 0.3)
# plt.show(fig)
# plt.close(fig)
def plot_prior_likelihood_posterior(prior, likelihood, posterior):
# definitions for the axes
left, width = 0.05, 0.3
bottom, height = 0.05, 0.9
padding = 0.12
small_width = 0.1
left_space = left + small_width + padding
added_space = padding + width
fig = plt.figure(figsize=(12, 4))
rect_prior = [left, bottom, small_width, height]
rect_likelihood = [left_space , bottom , width, height]
rect_posterior = [left_space + added_space, bottom , width, height]
ax_prior = fig.add_axes(rect_prior)
ax_likelihood = fig.add_axes(rect_likelihood, sharey=ax_prior)
ax_posterior = fig.add_axes(rect_posterior, sharey = ax_prior)
rect_colormap = plt.cm.Blues
# Show posterior probs and marginals
ax_prior.barh(0, prior[0], facecolor = rect_colormap(prior[0, 0]))
ax_prior.barh(1, prior[1], facecolor = rect_colormap(prior[1, 0]))
ax_likelihood.matshow(likelihood, vmin=0., vmax=1., cmap='Reds')
ax_posterior.matshow(posterior, vmin=0., vmax=1., cmap='Greens')
# Probabilities plot details
# ax_prior.set(xlim = [1, 0], yticks = [0, 1], yticklabels = ['left', 'right'],
# ylabel = 'state (s)', title = "Prior p(s)")
ax_prior.set(xlim = [1, 0], xticks = [], yticks = [0, 1], yticklabels = ['left', 'right'],
title = "Probability of state")
ax_prior.yaxis.tick_right()
ax_prior.spines['left'].set_visible(False)
ax_prior.spines['bottom'].set_visible(False)
# Likelihood plot details
ax_likelihood.set(xticks = [0, 1], xticklabels = ['fish', 'no fish'],
yticks = [0, 1], yticklabels = ['left', 'right'],
ylabel = 'state (s)', xlabel = 'measurement (m)',
title = 'Likelihood p(m (left) | s)')
ax_likelihood.xaxis.set_ticks_position('bottom')
ax_likelihood.spines['left'].set_visible(False)
ax_likelihood.spines['bottom'].set_visible(False)
# Posterior plot details
ax_posterior.set(xticks = [0, 1], xticklabels = ['fish', 'no fish'],
yticks = [0, 1], yticklabels = ['left', 'right'],
ylabel = 'state (s)', xlabel = 'measurement (m)',
title = 'Posterior p(s | m)')
ax_posterior.xaxis.set_ticks_position('bottom')
ax_posterior.spines['left'].set_visible(False)
ax_posterior.spines['bottom'].set_visible(False)
# show values
ind = np.arange(2)
x,y = np.meshgrid(ind,ind)
for i,j in zip(x.flatten(), y.flatten()):
c = f"{posterior[i,j]:.2f}"
ax_posterior.text(j,i, c, va='center', ha='center', color='black')
for i,j in zip(x.flatten(), y.flatten()):
c = f"{likelihood[i,j]:.2f}"
ax_likelihood.text(j,i, c, va='center', ha='center', color='black')
for i in ind:
v = prior[i, 0]
c = f"{v:.2f}"
ax_prior.text(v+0.2, i, c, va='center', ha='center', color='black')
def plot_prior_likelihood(ps, p_a_s1, p_a_s0, measurement):
likelihood = np.asarray([[p_a_s1, 1-p_a_s1],[p_a_s0, 1-p_a_s0]])
assert 0.0 <= ps <= 1.0
prior = np.asarray([ps, 1 - ps])
if measurement == "Fish":
posterior = likelihood[:, 0] * prior
else:
posterior = (likelihood[:, 1] * prior).reshape(-1)
posterior /= np.sum(posterior)
# definitions for the axes
left, width = 0.05, 0.3
bottom, height = 0.05, 0.9
padding = 0.12
small_width = 0.2
left_space = left + small_width + padding
small_padding = 0.05
fig = plt.figure(figsize=(12, 4))
rect_prior = [left, bottom, small_width, height]
rect_likelihood = [left_space , bottom , width, height]
rect_posterior = [left_space + width + small_padding, bottom , small_width, height]
ax_prior = fig.add_axes(rect_prior)
ax_likelihood = fig.add_axes(rect_likelihood, sharey=ax_prior)
ax_posterior = fig.add_axes(rect_posterior, sharey=ax_prior)
prior_colormap = plt.cm.Blues
posterior_colormap = plt.cm.Greens
# Show posterior probs and marginals
ax_prior.barh(0, prior[0], facecolor = prior_colormap(prior[0]))
ax_prior.barh(1, prior[1], facecolor = prior_colormap(prior[1]))
ax_likelihood.matshow(likelihood, vmin=0., vmax=1., cmap='Reds')
# ax_posterior.matshow(posterior, vmin=0., vmax=1., cmap='')
ax_posterior.barh(0, posterior[0], facecolor = posterior_colormap(posterior[0]))
ax_posterior.barh(1, posterior[1], facecolor = posterior_colormap(posterior[1]))
# Probabilities plot details
ax_prior.set(xlim = [1, 0], yticks = [0, 1], yticklabels = ['left', 'right'],
title = "Prior p(s)", xticks = [])
ax_prior.yaxis.tick_right()
ax_prior.spines['left'].set_visible(False)
ax_prior.spines['bottom'].set_visible(False)
# Likelihood plot details
ax_likelihood.set(xticks = [0, 1], xticklabels = ['fish', 'no fish'],
yticks = [0, 1], yticklabels = ['left', 'right'],
ylabel = 'state (s)', xlabel = 'measurement (m)',
title = 'Likelihood p(m | s)')
ax_likelihood.xaxis.set_ticks_position('bottom')
ax_likelihood.spines['left'].set_visible(False)
ax_likelihood.spines['bottom'].set_visible(False)
# Posterior plot details
ax_posterior.set(xlim = [0, 1], xticks = [], yticks = [0, 1],
yticklabels = ['left', 'right'], title = "Posterior p(s | m)")
ax_posterior.spines['left'].set_visible(False)
ax_posterior.spines['bottom'].set_visible(False)
# ax_posterior.axis('off')
# ax_posterior.set(xticks = [0, 1], xticklabels = ['fish', 'no fish'],
# yticks = [0, 1], yticklabels = ['left', 'right'],
# ylabel = 'state (s)', xlabel = 'measurement (m)',
# title = 'Posterior p(s | m)')
# ax_posterior.xaxis.set_ticks_position('bottom')
# ax_posterior.spines['left'].set_visible(False)
# ax_posterior.spines['bottom'].set_visible(False)
# show values
ind = np.arange(2)
x,y = np.meshgrid(ind,ind)
# for i,j in zip(x.flatten(), y.flatten()):
# c = f"{posterior[i,j]:.2f}"
# ax_posterior.text(j,i, c, va='center', ha='center', color='black')
for i in ind:
v = posterior[i]
c = f"{v:.2f}"
ax_posterior.text(v+0.2, i, c, va='center', ha='center', color='black')
for i,j in zip(x.flatten(), y.flatten()):
c = f"{likelihood[i,j]:.2f}"
ax_likelihood.text(j,i, c, va='center', ha='center', color='black')
for i in ind:
v = prior[i]
c = f"{v:.2f}"
ax_prior.text(v+0.2, i, c, va='center', ha='center', color='black')
return fig
# fig = plot_prior_likelihood(0.5, 0.3)
# plt.show(fig)
# plt.close(fig)
from matplotlib import colors
def plot_utility(ps):
prior = np.asarray([ps, 1 - ps])
utility = np.array([[2, -3], [-2, 1]])
expected = prior @ utility
# definitions for the axes
left, width = 0.05, 0.16
bottom, height = 0.05, 0.9
padding = 0.02
small_width = 0.1
left_space = left + small_width + padding
added_space = padding + width
fig = plt.figure(figsize=(17, 3))
rect_prior = [left, bottom, small_width, height]
rect_utility = [left + added_space , bottom , width, height]
rect_expected = [left + 2* added_space, bottom , width, height]
ax_prior = fig.add_axes(rect_prior)
ax_utility = fig.add_axes(rect_utility, sharey=ax_prior)
ax_expected = fig.add_axes(rect_expected)
rect_colormap = plt.cm.Blues
# Data of plots
ax_prior.barh(0, prior[0], facecolor = rect_colormap(prior[0]))
ax_prior.barh(1, prior[1], facecolor = rect_colormap(prior[1]))
ax_utility.matshow(utility, cmap='cool')
norm = colors.Normalize(vmin=-3, vmax=3)
ax_expected.bar(0, expected[0], facecolor = rect_colormap(norm(expected[0])))
ax_expected.bar(1, expected[1], facecolor = rect_colormap(norm(expected[1])))
# Probabilities plot details
ax_prior.set(xlim = [1, 0], xticks = [], yticks = [0, 1], yticklabels = ['left', 'right'],
title = "Probability of state")
ax_prior.yaxis.tick_right()
ax_prior.spines['left'].set_visible(False)
ax_prior.spines['bottom'].set_visible(False)
# Utility plot details
ax_utility.set(xticks = [0, 1], xticklabels = ['left', 'right'],
yticks = [0, 1], yticklabels = ['left', 'right'],
ylabel = 'state (s)', xlabel = 'action (a)',
title = 'Utility')
ax_utility.xaxis.set_ticks_position('bottom')
ax_utility.spines['left'].set_visible(False)
ax_utility.spines['bottom'].set_visible(False)
# Expected utility plot details
ax_expected.set(title = 'Expected utility', ylim = [-3, 3],
xticks = [0, 1], xticklabels = ['left', 'right'],
xlabel = 'action (a)',
yticks = [])
ax_expected.xaxis.set_ticks_position('bottom')
ax_expected.spines['left'].set_visible(False)
ax_expected.spines['bottom'].set_visible(False)
# show values
ind = np.arange(2)
x,y = np.meshgrid(ind,ind)
for i,j in zip(x.flatten(), y.flatten()):
c = f"{utility[i,j]:.2f}"
ax_utility.text(j,i, c, va='center', ha='center', color='black')
for i in ind:
v = prior[i]
c = f"{v:.2f}"
ax_prior.text(v+0.2, i, c, va='center', ha='center', color='black')
for i in ind:
v = expected[i]
c = f"{v:.2f}"
ax_expected.text(i, 2.5, c, va='center', ha='center', color='black')
return fig
def plot_prior_likelihood_utility(ps, p_a_s1, p_a_s0, measurement):
assert 0.0 <= ps <= 1.0
assert 0.0 <= p_a_s1 <= 1.0
assert 0.0 <= p_a_s0 <= 1.0
prior = np.asarray([ps, 1 - ps])
likelihood = np.asarray([[p_a_s1, 1-p_a_s1],[p_a_s0, 1-p_a_s0]])
utility = np.array([[2.0, -3.0], [-2.0, 1.0]])
# expected = np.zeros_like(utility)
if measurement == "Fish":
posterior = likelihood[:, 0] * prior
else:
posterior = (likelihood[:, 1] * prior).reshape(-1)
posterior /= np.sum(posterior)
# expected[:, 0] = utility[:, 0] * posterior
# expected[:, 1] = utility[:, 1] * posterior
expected = posterior @ utility
# definitions for the axes
left, width = 0.05, 0.3
bottom, height = 0.05, 0.3
padding = 0.12
small_width = 0.2
left_space = left + small_width + padding
small_padding = 0.05
fig = plt.figure(figsize=(10, 9))
rect_prior = [left, bottom + height + padding, small_width, height]
rect_likelihood = [left_space , bottom + height + padding , width, height]
rect_posterior = [left_space + width + small_padding, bottom + height + padding , small_width, height]
rect_utility = [padding, bottom, width, height]
rect_expected = [padding + width + padding + left, bottom, width, height]
ax_likelihood = fig.add_axes(rect_likelihood)
ax_prior = fig.add_axes(rect_prior, sharey=ax_likelihood)
ax_posterior = fig.add_axes(rect_posterior, sharey=ax_likelihood)
ax_utility = fig.add_axes(rect_utility)
ax_expected = fig.add_axes(rect_expected)
prior_colormap = plt.cm.Blues
posterior_colormap = plt.cm.Greens
expected_colormap = plt.cm.Wistia
# Show posterior probs and marginals
ax_prior.barh(0, prior[0], facecolor = prior_colormap(prior[0]))
ax_prior.barh(1, prior[1], facecolor = prior_colormap(prior[1]))
ax_likelihood.matshow(likelihood, vmin=0., vmax=1., cmap='Reds')
ax_posterior.barh(0, posterior[0], facecolor = posterior_colormap(posterior[0]))
ax_posterior.barh(1, posterior[1], facecolor = posterior_colormap(posterior[1]))
ax_utility.matshow(utility, vmin=0., vmax=1., cmap='cool')
# ax_expected.matshow(expected, vmin=0., vmax=1., cmap='Wistia')
ax_expected.bar(0, expected[0], facecolor = expected_colormap(expected[0]))
ax_expected.bar(1, expected[1], facecolor = expected_colormap(expected[1]))
# Probabilities plot details
ax_prior.set(xlim = [1, 0], yticks = [0, 1], yticklabels = ['left', 'right'],
title = "Prior p(s)", xticks = [])
ax_prior.yaxis.tick_right()
ax_prior.spines['left'].set_visible(False)
ax_prior.spines['bottom'].set_visible(False)
# Likelihood plot details
ax_likelihood.set(xticks = [0, 1], xticklabels = ['fish', 'no fish'],
yticks = [0, 1], yticklabels = ['left', 'right'],
ylabel = 'state (s)', xlabel = 'measurement (m)',
title = 'Likelihood p(m | s)')
ax_likelihood.xaxis.set_ticks_position('bottom')
ax_likelihood.spines['left'].set_visible(False)
ax_likelihood.spines['bottom'].set_visible(False)
# Posterior plot details
ax_posterior.set(xlim = [0, 1], xticks = [], yticks = [0, 1],
yticklabels = ['left', 'right'], title = "Posterior p(s | m)")
ax_posterior.spines['left'].set_visible(False)
ax_posterior.spines['bottom'].set_visible(False)
# Utility plot details
ax_utility.set(xticks = [0, 1], xticklabels = ['left', 'right'],
xlabel = 'action (a)', yticks = [0, 1], yticklabels = ['left', 'right'],
title = 'Utility', ylabel = 'state (s)')
ax_utility.xaxis.set_ticks_position('bottom')
ax_utility.spines['left'].set_visible(False)
ax_utility.spines['bottom'].set_visible(False)
# Expected Utility plot details
ax_expected.set(ylim = [-2, 2], xticks = [0, 1], xticklabels = ['left', 'right'],
xlabel = 'action (a)', title = 'Expected utility', yticks=[])
# ax_expected.axis('off')
ax_expected.spines['left'].set_visible(False)
# ax_expected.set(xticks = [0, 1], xticklabels = ['left', 'right'],
# xlabel = 'action (a)',
# title = 'Expected utility')
# ax_expected.xaxis.set_ticks_position('bottom')
# ax_expected.spines['left'].set_visible(False)
# ax_expected.spines['bottom'].set_visible(False)
# show values
ind = np.arange(2)
x,y = np.meshgrid(ind,ind)
for i in ind:
v = posterior[i]
c = f"{v:.2f}"
ax_posterior.text(v+0.2, i, c, va='center', ha='center', color='black')
for i,j in zip(x.flatten(), y.flatten()):
c = f"{likelihood[i,j]:.2f}"
ax_likelihood.text(j,i, c, va='center', ha='center', color='black')
for i,j in zip(x.flatten(), y.flatten()):
c = f"{utility[i,j]:.2f}"
ax_utility.text(j,i, c, va='center', ha='center', color='black')
# for i,j in zip(x.flatten(), y.flatten()):
# c = f"{expected[i,j]:.2f}"
# ax_expected.text(j,i, c, va='center', ha='center', color='black')
for i in ind:
v = prior[i]
c = f"{v:.2f}"
ax_prior.text(v+0.2, i, c, va='center', ha='center', color='black')
for i in ind:
v = expected[i]
c = f"{v:.2f}"
ax_expected.text(i, v, c, va='center', ha='center', color='black')
# # show values
# ind = np.arange(2)
# x,y = np.meshgrid(ind,ind)
# for i,j in zip(x.flatten(), y.flatten()):
# c = f"{P[i,j]:.2f}"
# ax.text(j,i, c, va='center', ha='center', color='white')
# for i in ind:
# v = marginal_x[i]
# c = f"{v:.2f}"
# ax_x.text(i, v +0.2, c, va='center', ha='center', color='black')
# v = marginal_y[i]
# c = f"{v:.2f}"
# ax_y.text(v+0.2, i, c, va='center', ha='center', color='black')
return fig
# @title Helper Functions
def compute_marginal(px, py, cor):
# calculate 2x2 joint probabilities given marginals p(x=1), p(y=1) and correlation
p11 = px*py + cor*np.sqrt(px*py*(1-px)*(1-py))
p01 = px - p11
p10 = py - p11
p00 = 1.0 - p11 - p01 - p10
return np.asarray([[p00, p01], [p10, p11]])
# test
# print(compute_marginal(0.4, 0.6, -0.8))
def compute_cor_range(px,py):
# Calculate the allowed range of correlation values given marginals p(x=1) and p(y=1)
def p11(corr):
return px*py + corr*np.sqrt(px*py*(1-px)*(1-py))
def p01(corr):
return px - p11(corr)
def p10(corr):
return py - p11(corr)
def p00(corr):
return 1.0 - p11(corr) - p01(corr) - p10(corr)
Cmax = min(fsolve(p01, 0.0), fsolve(p10, 0.0))
Cmin = max(fsolve(p11, 0.0), fsolve(p00, 0.0))
return Cmin, Cmax
```
---
# Section 1: Gone Fishin'
```
# @title Video 2: Gone Fishin'
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="McALsTzb494", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
<details>
<summary> <font color=‘blue’>Click here for text recap of video </font></summary>
You were just introduced to the **binary hidden state problem** we are going to explore. You need to decide which side to fish on--the hidden state. We know fish like to school together. On different days the school of fish is either on the left or right side, but we don’t know what the case is today. We define our knowledge about the fish location as a distribution over the random hidden state variable. Using our probabilistic knowledge, also called our **belief** about the hidden state, we will explore how to make the decision about where to fish today, based on what to expect in terms of gains or losses for the decision.
The gains and losss are defined by the utility of choosing an action, which is fishing on the left or right. The details of the utilities are described
</details>
In the next two sections we will consider just the probability of where the fish might be and what you gain or lose by choosing where to fish.
Remember, you can either think of your self as a scientist conducting an experiment or as a brain trying to make a decision. The Bayesian approach is the same!
---
# Section 2: Deciding where to fish
```
# @title Video 3: Utility
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="xvIVZrqF_5s", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
<details>
<summary> <font color=‘blue’>Click here for text recap of video </font></summary>
You need to decide where to fish. It may seem obvious - you could just fish on the side where the probability of the fish being is higher! Unfortunately, decisions and actions are always a little more complicated. Deciding to fish may be influenced by more than just the probability of the school of fish being there as we saw by the potential issues of submarines and sunburn. The consequences of the action you take is based on the true (but hidden) state of the world and the action you choose! In our example, fishing on the wrong side, where there aren't many fish, is likely to lead to you spending your afternoon not catching fish and therefor getting a sunburn. The submarine represents a risk to fishing on the right side that is greater than the left side. If you want to know what to expect from taking the action of fishing on one side or the other, you need to calculate the expected utility.
</details>
You know the (prior) probability that the school of fish is on the left side of the dock today, $P(s = left)$. So, you also know the probability the school is on the right, $P(s = right)$, because these two probabilities must add up to 1.
We quantify gains and losses numerically using a **utility** function $U(s,a)$, which describes the consequences of your actions: how much value you gain (or if negative, lose) given the state of the world ($s$) and the action you take ($a$). In our example, our utility can be summarized as:
| Utility: U(s,a) | a = left | a = right |
| ----------------- |------------|------------|
| s = Left | +2 | -3 |
| s = right | -2 | +1 |
To use possible gains and losses to choose an action, we calculate the **expected utility** of that action by weighing these utilities with the probability of that state occuring. This allows us to choose actions by taking probabilities of events into account: we don't care if the outcome of an action-state pair is a loss if the probability of that state is very low. We can formalize this as:
$$ \text{Expected utility of action a} = \sum_{s}U(s,a)P(s) $$
In other words, the expected utility of an action a is the sum over possible states of the utility of that action and state times the probability of that state.
## Interactive Demo 2: Exploring the decision
Let's start to get a sense of how all this works using the interactive demo below. You can change the probability that the school of fish is on the left side ($p(s = left)$ using the slider. You will see the utility function (a matrix) in the middle and the corresponding expected utility for each action on the right.
First, make sure you understand how the expected utility of each action is being computed from the probabilities and the utility values. In the initial state: the probability of the fish being on the left is 0.9 and on the right is 0.1. The expected utility of the action of fishing on the left is then $U(s = left,a = left)p(s = left) + U(s = right,a = left)p(s = right) = 2(0.9) + -2(0.1) = 1.6$.
For each of these scenarios, think and discuss first. Then use the demo to try out each and see if your action would have been correct (that is, if the expected value of that action is the highest).
1. You just arrived at the dock for the first time and have no sense of where the fish might be. So you guess that the probability of the school being on the left side is 0.5 (so the probability on the right side is also 0.5). Which side would you choose to fish on given our utility values?
2. You think that the probability of the school being on the left side is very low (0.1) and correspondingly high on the right side (0.9). Which side would you choose to fish on given our utility values?
3. What would you choose if the probability of the school being on the left side is slightly lower than on the right side (0. 4 vs 0.6)?
```
# @markdown Execute this cell to use the widget
ps_widget = widgets.FloatSlider(0.9, description='p(s = left)', min=0.0, max=1.0, step=0.01)
@widgets.interact(
ps = ps_widget,
)
def make_utility_plot(ps):
fig = plot_utility(ps)
plt.show(fig)
plt.close(fig)
return None
# to_remove explanation
# 1) With equal probabilities, the expected utility is higher on the left side,
# since that is the side without submarines, so you would choose to fish there.
# 2) If the probability that the fish is on the right side is high, you would
# choose to fish there. The high probability of fish being on the right far outweights
# the slightly higher utilities from fishing on the left (as you are unlikely to gain these)
# 3) If the probability that the fish is on the right side is just slightly higher
#. than on the left, you would choose the left side as the expected utility is still
#. higher on the left. Note that in this situation, you are not simply choosing the
#. side with the higher probability - the utility really matters here for the decision
```
In this section, you have seen that both the utility of various state and action pairs and our knowledge of the probability of each state affects your decision. Importantly, we want our knowledge of the probability of each state to be as accurate as possible!
So how do we know these probabilities? We may have prior knowledge from years of fishing at the same dock, learning that the fish are more likely to be on the left side, for example. Of course, we need to update our knowledge (our belief)! To do this, we need to collect more information, or take some measurements! In the next few sections, we will focus on how we improve our knowledge of the probabilities.
---
# Section 3: Likelihood of the fish being on either side
```
# @title Video 4: Likelihood
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="l4m0JzMWGio", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
<details>
<summary> <font color=‘blue’>Click here for text recap of video </font></summary>
First, we'll think about what it means to take a measurement (also often called an observation or just data) and what it tells you about what the hidden state may be. Specifically, we'll be looking at the **likelihood**, which is the probability of your measurement ($m$) given the hidden state ($s$): $P(m | s)$. Remember that in this case, the hidden state is which side of the dock the school of fish is on.
We will watch someone fish (for let's say 10 minutes) and our measurement is whether they catch a fish or not. We know something about what catching a fish means for the likelihood of the fish being on one side or the other.
</details>
## Think! 3: Guessing the location of the fish
Let's say we go to different dock to fish. Here, there are different probabilities of catching fish given the state of the world. At this dock, if you fish on the side of the dock where the fish are, you have a 70% chance of catching a fish. If you fish on the wrong side, you will only catch a fish with only 20% probability. These are the likelihoods of observing someone catching a fish! That is, you are taking a measurement by seeing if someone else catches a fish!
You see a fisherperson is fishing on the left side.
1) Please calculate each of the following (best to each do the math and then compare your answers and dsicuss!):
- probability of catching a fish given that the school of fish is on the left side, $P(m = catch\text{ } fish | s = left )$
- probability of not catching a fish given that the school of fish is on the left side, $P(m = no \text{ } fish | s = left)$
- probability of catching a fish given that the school of fish is on the right side, $P(m = catch \text{ } fish | s = right)$
- probability of not catching a fish given that the school of fish is on the right side, $P(m = no \text{ } fish | s = right)$
2) If the fisherperson catches a fish, which side would you guess the school is on? Why?
3) If the fisherperson does not catch a fish, which side would you guess the school is on? Why?
```
#to_remove explanation
# 1) The fisherperson is on the left side so:
# - P(m = fish | s = left) = 0.7 because they have a 70% chance of catching
# a fish when on the same side as the school
# - P(m = no fish | s = left) = 0.3 because the probability of catching a fish
# and not catching a fish for a given state must add up to 1 as these
# are the only options: 1 - 0.7 = 0.3
# - P(m = fish | s = right) = 0.2
# - P(m = no fish | s = right) = 0.8
# 2) If the fisherperson catches a fish, you would guess the school of fish is on the
# left side. This is because the probability of catching a fish given that the
# school is on the left side (0.7) is higher than the probability given that
# the school is on the right side (0.2).
# 3) If the fisherperson does not catch a fish, you would guess the school of fish is on the
# right side. This is because the probability of not catching a fish given that the
# school is on the right side (0.8) is higher than the probability given that
# the school is on the right side (0.3).
```
In the prior exercise, you tried to guess where the school of fish was based on the measurement you took (watching someone fish). You did this by choosing the state (side where you think the fish are) that maximized the probability of the measurement. In other words, you estimated the state by maximizing the likelihood (the side with the highest probability of measurement given state $P(m|s$)). This is called maximum likelihood estimation (MLE) and you've encountered it before during this course, in W1D3!
But, what if you had been going to this dock for years and you knew that the fish were almost always on the left side? This should probably affect how you make your estimate -- you would rely less on the single new measurement and more on your prior knowledge. This is the fundemental idea behind Bayesian inference, as we will see later in this tutorial!
---
# Section 4: Correlation and marginalization
```
# @title Video 5: Correlation and marginalization
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="vsDjtWi-BVo", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
<details>
<summary> <font color=‘blue’>Click here for text recap of video </font></summary>
In this section, we are going to take a step back for a bit and think more generally about the amount of information shared between two random variables. We want to know how much information you gain when you observe one variable (take a measurement) if you know something about another. We will see that the fundamental concept is the same if we think about two attributes, for example the size and color of the fish, or the prior information and the likelihood.
</details>
## Think! 4: Covarying probability distributions
The relationship between the marginal probabilities and the joint probabilities is determined by the correlation between the two random variables - a normalized measure of how much the variables covary. We can also think of this as gaining some information about one of the variables when we observe a measurement from the other. We will think about this more formally in Tutorial 2.
Here, we want to think about how the correlation between size and color of these fish changes how much information we gain about one attribute based on the other. See Bonus Section 1 for the formula for correlation.
Use the widget below and answer the following questions:
1. When the correlation is zero, $\rho = 0$, what does the distribution of size tell you about color?
2. Set $\rho$ to something small. As you change the probability of golden fish, what happens to the ratio of size probabilities? Set $\rho$ larger (can be negative). Can you explain the pattern of changes in the probabilities of size as you change the probability of golden fish?
3. Set the probability of golden fish and of large fish to around 65%. As the correlation goes towards 1, how often will you see silver large fish?
4. What is increasing the (absolute) correlation telling you about how likely you are to see one of the properties if you see a fish with the other?
```
# @markdown Execute this cell to enable the widget
style = {'description_width': 'initial'}
gs = GridspecLayout(2,2)
cor_widget = widgets.FloatSlider(0.0, description='ρ', min=-1, max=1, step=0.01)
px_widget = widgets.FloatSlider(0.5, description='p(color=golden)', min=0.01, max=0.99, step=0.01, style=style)
py_widget = widgets.FloatSlider(0.5, description='p(size=large)', min=0.01, max=0.99, step=0.01, style=style)
gs[0,0] = cor_widget
gs[0,1] = px_widget
gs[1,0] = py_widget
@widgets.interact(
px=px_widget,
py=py_widget,
cor=cor_widget,
)
def make_corr_plot(px, py, cor):
Cmin, Cmax = compute_cor_range(px, py) #allow correlation values
cor_widget.min, cor_widget.max = Cmin+0.01, Cmax-0.01
if cor_widget.value > Cmax:
cor_widget.value = Cmax
if cor_widget.value < Cmin:
cor_widget.value = Cmin
cor = cor_widget.value
P = compute_marginal(px,py,cor)
# print(P)
fig = plot_joint_probs(P)
plt.show(fig)
plt.close(fig)
return None
# gs[1,1] = make_corr_plot()
# to_remove explanation
#' 1. When the correlation is zero, the two properties are completely independent.
#' This means you don't gain any information about one variable from observing the other.
#' Importantly, the marginal distribution of one variable is therefore independent of the other.
#' 2. The correlation controls the distribution of probability across the joint probabilty table.
#' The higher the correlation, the more the probabilities are restricted by the fact that both rows
#' and columns need to sum to one! While the marginal probabilities show the relative weighting, the
#' absolute probabilities for one quality will become more dependent on the other as the correlation
#' goes to 1 or -1.
#' 3. The correlation will control how much probabilty mass is located on the diagonals. As the
#' correlation goes to 1 (or -1), the probability of seeing the one of the two pairings has to goes
#' towards zero!
#' 4. If we think about what information we gain by observing one quality, the intution from (3.) tells
#' us that we know more (have more information) about the other quality as a function of the correlation.
```
We have just seen how two random variables can be more or less independent. The more correlated, the less independent, and the more shared information. We also learned that we can marginalize to determine the marginal likelihood of a measurement or to find the marginal probability distribution of two random variables. We are going to now complete our journey towards being fully Bayesian!
## Math Exercise 4.1: Computing marginal probabilities
To understand the information between two variables, let's first consider the size and color of the fish.
| P(X, Y) | Y = silver | Y = gold |
| -------------- |-------------|-----------|
| X = small | 0.4 | 0.2 |
| X = large | 0.1 | 0.3 |
The table above shows us the **joint probabilities**: the probability of both specific attributes occuring together. For example, the probability of a fish being small and silver ($P(X = small, Y = silver$) is 0.4.
We want to know what the probability of a fish being small regardless of color. Since the fish are either silver or gold, this would be the probability of a fish being small and silver plus the probability of a fish being small and gold. This is an example of marginalizing, or averaging out, the variable we are not interested in across the rows or columns.. In math speak: $P(X = small) = \sum_y{P(X = small, Y)}$. This gives us a **marginal probability**, a probability of a variable outcome (in this case size), regardless of the other variables (in this case color).
Please complete the following math problems to further practice thinking through probabilities:
1. Calculate the probability of a fish being silver.
2. Calculate the probability of a fish being small, large, silver, or gold.
3. Calculate the probability of a fish being small OR gold. (Hint: $P(A\ \textrm{or}\ B) = P(A) + P(B) - P(A\ \textrm{and}\ B)$)
```
# to_remove explanation
# 1) The probability of a fish being silver is the joint probability of it being
#. small and silver plus the joint probability of it being large and silver:
#
#. P(Y = silver) = P(X = small, Y = silver) + P(X = large, Y = silver)
#. = 0.4 + 0.1
#. = 0.5
# 2) This is all the possibilities as in this scenario, our fish can only be small
#. or large, silver or gold. So the probability is 1 - the fish has to be at
#. least one of these.
#. 3) First we compute the marginal probabilities
#. P(X = small) = P(X = small, Y = silver) + P(X = small, Y = gold) = 0.6
#. P(Y = gold) = P(X = small, Y = gold) + P(X = large, Y = gold) = 0.5
#. We already know the joint probability: P(X = small, Y = gold) = 0.2
#. We can now use the given formula:
#. P( X = small or Y = gold) = P(X = small) + P(Y = gold) - P(X = small, Y = gold)
#. = 0.6 + 0.5 - 0.2
#. = 0.9
```
## Math Exercise 4.2: Computing marginal likelihood
When we normalize to find the posterior, we need to determine the marinal likelihood--or evidence--for the measurement we observed. To do this, we need to marginalize as we just did above to find the probabilities of a color or size. Only, in this case, we are marginalizing to remove a conditioning variable! In this case, let's consider the likelihood of fish (if we observed a fisherperson fishing on the **right**).
| p(m|s) | m = fish | m = no fish |
| ------------ |----------|--------------|
| s = left | 0.1 | 0.9 |
| s = right | 0.5 | 0.5 |
The table above shows us the **likelihoods**, just as we explored earlier.
You want to know the total probability of a fish being caught, $p(m)$, by the fisherperson fishing on the right. (You would need this to calcualate the posterior.) To do this, you will need to consider the prior probability, $p(s)$, and marginalize over the hidden states!
This is an example of marginalizing, or conditioning away, the variable we are not interested in as well.
Please complete the following math problems to further practice thinking through probabilities:
1. Calculate the marginal likelihood if the priors are: $p(s = left) = 0.3$ and $p(s = right) = 0.7$.
2. Calculate the marginal likelihood if the priors are: $p(s = left) = 0.6$ and $p(s = right) = 0.4$.
```
# to_remove explanation
# 1) The mariginal likelihood (evidence) is
#
#. P(m = fish) = P(m = fish, s = left) + P(m = fish, s = right)
#. = P(m = fish, s = left)P(s = left) + P(m = fish, s = right)P(s = right)
#. = 0.1 * 0.3 + .5 * .7
#. = 0.38
# 2) The mariginal likelihood (evidence) is
#
#. P(m = fish) = P(m = fish, s = left) + P(m = fish, s = right)
#. = P(m = fish, s = left)P(s = left) + P(m = fish, s = right)P(s = right)
#. = 0.1 * 0.6 + .5 * .4
#. = 0.26
```
---
# Section 5: Bayes' Rule and the Posterior
Marginalization is going to be used to combine our prior knowlege, which we call the **prior**, and our new information from a measurement, the **likelihood**. Only in this case, the information we gain about the hidden state we are interested in, where the fish are, is based on the relationship between the probabilities of the measurement and our prior.
We can now calculate the full posterior distribution for the hidden state ($s$) using Bayes' Rule. As we've seen, the posterior is proportional the the prior times the likelihood. This means that the posterior probability of the hidden state ($s$) given a measurement ($m$) is proportional to the likelihood of the measurement given the state times the prior probability of that state:
$$ P(s | m) \propto P(m | s) P(s) $$
We say proportional to instead of equal because we need to normalize to produce a full probability distribution:
$$ P(s | m) = \frac{P(m | s) P(s)}{P(m)} $$
Normalizing by this $P(m)$ means that our posterior is a complete probability distribution that sums or integrates to 1 appropriately. We now can use this new, complete probability distribution for any future inference or decisions we like! In fact, as we will see tomorrow, we can use it as a new prior! Finally, we often call this probability distribution our beliefs over the hidden states, to emphasize that it is our subjective knowlege about the hidden state.
For many complicated cases, like those we might be using to model behavioral or brain inferences, the normalization term can be intractable or extremely complex to calculate. We can be careful to choose probability distributions were we can analytically calculate the posterior probability or numerical approximation is reliable. Better yet, we sometimes don't need to bother with this normalization! The normalization term, $P(m)$, is the probability of the measurement. This does not depend on state so is essentially a constant we can often ignore. We can compare the unnormalized posterior distribution values for different states because how they relate to each other is unchanged when divided by the same constant. We will see how to do this to compare evidence for different hypotheses tomorrow. (It's also used to compare the likelihood of models fit using maximum likelihood estimation, as you did in W1D5.)
In this relatively simple example, we can compute the marginal likelihood $P(m)$ easily by using:
$$P(m) = \sum_s P(m | s) P(s)$$
We can then normalize so that we deal with the full posterior distribution.
## Math Exercise 5: Calculating a posterior probability
Our prior is $p(s = left) = 0.3$ and $p(s = right) = 0.7$. In the video, we learned that the chance of catching a fish given they fish on the same side as the school was 50%. Otherwise, it was 10%. We observe a person fishing on the left side. Our likelihood is:
| Likelihood: p(m \| s) | m = fish | m = no fish |
| ----------------- |----------|----------|
| s = left | 0.5 | 0.5 |
| s = right | 0.1 | 0.9 |
Calculate the posterior probability (on paper) that:
1. The school is on the left if the fisherperson catches a fish: $p(s = left | m = fish)$ (hint: normalize by compute $p(m = fish)$)
2. The school is on the right if the fisherperson does not catch a fish: $p(s = right | m = no fish)$
```
# to_remove explanation
# 1. Using Bayes rule, we know that P(s = left | m = fish) = P(m = fish | s = left)P(s = left) / P(m = fish)
#. Let's first compute P(m = fish):
#. P(m = fish) = P(m = fish | s = left)P(s = left) + P(m = fish | s = right)P(s = right)
# = 0.5 * 0.3 + .1*.7
# = 0.22
#. Now we can plug in all parts of Bayes rule:
# P(s = left | m = fish) = P(m = fish | s = left)P(s = left) / P(m = fish)
# = 0.5*0.3/0.22
# = 0.68
# 2. Using Bayes rule, we know that P(s = right | m = no fish) = P(m = no fish | s = right)P(s = right) / P(m = no fish)
#. Let's first compute P(m = no fish):
#. P(m = no fish) = P(m = no fish | s = left)P(s = left) + P(m = no fish | s = right)P(s = right)
# = 0.5 * 0.3 + .9*.7
# = 0.78
#. Now we can plug in all parts of Bayes rule:
# P(s = right | m = no fish) = P(m = no fish | s = right)P(s = right) / P(m = no fish)
# = 0.9*0.7/0.78
# = 0.81
```
## Coding Exercise 5: Computing Posteriors
Let's implement our above math to be able to compute posteriors for different priors and likelihoods.
As before, our prior is $p(s = left) = 0.3$ and $p(s = right) = 0.7$. In the video, we learned that the chance of catching a fish given they fish on the same side as the school was 50%. Otherwise, it was 10%. We observe a person fishing on the left side. Our likelihood is:
| Likelihood: p(m \| s) | m = fish | m = no fish |
| ----------------- |----------|----------|
| s = left | 0.5 | 0.5 |
| s = right | 0.1 | 0.9 |
We want our full posterior to take the same 2 by 2 form. Make sure the outputs match your math answers!
```
def compute_posterior(likelihood, prior):
""" Use Bayes' Rule to compute posterior from likelihood and prior
Args:
likelihood (ndarray): i x j array with likelihood probabilities where i is
number of state options, j is number of measurement options
prior (ndarray): i x 1 array with prior probability of each state
Returns:
ndarray: i x j array with posterior probabilities where i is
number of state options, j is number of measurement options
"""
#################################################
## TODO for students ##
# Fill out function and remove
raise NotImplementedError("Student exercise: implement compute_posterior")
#################################################
# Compute unnormalized posterior (likelihood times prior)
posterior = ... # first row is s = left, second row is s = right
# Compute p(m)
p_m = np.sum(posterior, axis = 0)
# Normalize posterior (divide elements by p_m)
posterior /= ...
return posterior
# # Uncomment and run
# # Make prior
# prior = np.array([0.3, 0.7]).reshape((2, 1)) # first row is s = left, second row is s = right
# # Make likelihood
# likelihood = np.array([[0.5, 0.5], [0.1, 0.9]]) # first row is s = left, second row is s = right
# # Compute posterior
# posterior = compute_posterior(likelihood, prior)
# # Visualize
# plot_prior_likelihood_posterior(prior, likelihood, posterior)
# to_remove solution
def compute_posterior(likelihood, prior):
""" Use Bayes' Rule to compute posterior from likelihood and prior
Args:
likelihood (ndarray): i x j array with likelihood probabilities where i is
number of state options, j is number of measurement options
prior (ndarray): i x 1 array with prior probability of each state
Returns:
ndarray: i x j array with posterior probabilities where i is
number of state options, j is number of measurement options
"""
# Compute unnormalized posterior (likelihood times prior)
posterior = likelihood * prior # first row is s = left, second row is s = right
# Compute p(m)
p_m = np.sum(posterior, axis = 0)
# Normalize posterior (divide elements by p_m)
posterior /= p_m
return posterior
# Make prior
prior = np.array([0.3, 0.7]).reshape((2, 1)) # first row is s = left, second row is s = right
# Make likelihood
likelihood = np.array([[0.5, 0.5], [0.1, 0.9]]) # first row is s = left, second row is s = right
# Compute posterior
posterior = compute_posterior(likelihood, prior)
# Visualize
with plt.xkcd():
plot_prior_likelihood_posterior(prior, likelihood, posterior)
```
## Interactive Demo 5: What affects the posterior?
Now that we can understand the implementation of *Bayes rule*, let's vary the parameters of the prior and likelihood to see how changing the prior and likelihood affect the posterior.
In the demo below, you can change the prior by playing with the slider for $p( s = left)$. You can also change the likelihood by changing the probability of catching a fish given that the school is on the left and the probability of catching a fish given that the school is on the right. The fisherperson you are observing is fishing on the left.
1. Keeping the likelihood constant, when does the prior have the strongest influence over the posterior? Meaning, when does the posterior look most like the prior no matter whether a fish was caught or not?
2. What happens if the likelihoods for catching a fish are similar when you fish on the correct or incorrect side?
3. Set the prior probability of the state = left to 0.6 and play with the likelihood. When does the likelihood exert the most influence over the posterior?
```
# @markdown Execute this cell to enable the widget
# style = {'description_width': 'initial'}
ps_widget = widgets.FloatSlider(0.3, description='p(s = left)',
min=0.01, max=0.99, step=0.01)
p_a_s1_widget = widgets.FloatSlider(0.5, description='p(fish on left | state = left)',
min=0.01, max=0.99, step=0.01, style=style, layout=Layout(width='370px'))
p_a_s0_widget = widgets.FloatSlider(0.1, description='p(fish on left | state = right)',
min=0.01, max=0.99, step=0.01, style=style, layout=Layout(width='370px'))
# observed_widget = widgets.Checkbox(value=False, description='Observed fish (m)',
# disabled=False, indent=False,
# layout=Layout(display="flex", justify_content="center"))
observed_widget = ToggleButtons(options=['Fish', 'No Fish'],
description='Observation (m) on the left:', disabled=False, button_style='',
layout=Layout(width='auto', display="flex"),
style={'description_width': 'initial'}
)
widget_ui = VBox([ps_widget,
HBox([p_a_s1_widget, p_a_s0_widget]),
observed_widget])
widget_out = interactive_output(plot_prior_likelihood,
{'ps': ps_widget,
'p_a_s1': p_a_s1_widget,
'p_a_s0': p_a_s0_widget,
'measurement': observed_widget})
display(widget_ui, widget_out)
# @widgets.interact(
# ps=ps_widget,
# p_a_s1=p_a_s1_widget,
# p_a_s0=p_a_s0_widget,
# m_right=observed_widget
# )
# def make_prior_likelihood_plot(ps,p_a_s1,p_a_s0,m_right):
# fig = plot_prior_likelihood(ps,p_a_s1,p_a_s0,m_right)
# plt.show(fig)
# plt.close(fig)
# return None
# to_remove explanation
# 1). The prior exerts a strong influence over the posterior when it is very informative: when
#. the probability of the school being on one side or the other. If the prior that the fish are
#. on the left side is very high (like 0.9), the posterior probability of the state being left is
#. high regardless of the measurement.
# 2). When the likelihoods are similar, the information gained from catching a fish or not is less informativeself.
#. intuitively, if you were about as likely to catch a fish regardless of the true location, then catching a fish
#. doesn't tell you very much! The differences between the likelihoods is a way of thinking about how much information
#. we can gain. You can try to figure out why, as we've given you all the clues...
#. 3) Similarly to the prior, the likelihood exerts the most influence when it is informative: when catching
#. a fish tells you a lot of information about which state is likely. For example, if the probability of the
#. fisherperson catching a fish if he is fishing on the right side and the school is on the left is 0
#. (p fish | s = left) = 0 and the probability of catching a fish if the school is on the right is 1, the
#. prior does not affect the posterior at all. The measurement tells you the hidden state completely.
```
# Section 6: Making Bayesian fishing decisions
We will explore how to consider the expected utility of an action based on our belief (the posterior distribution) about where we think the fish are. Now we have all the components of a Bayesian decision: our prior information, the likelihood given a measurement, the posterior distribution (belief) and our utility (the gains and losses). This allows us to consider the relationship between the true value of the hidden state, $s$, and what we *expect* to get if we take action, $a$, based on our belief!
Let's use the following widget to think about the relationship between these probability distributions and utility function.
## Think! 6: What is more important, the probabilities or the utilities?
We are now going to put everything we've learned together to gain some intuitions for how each of the elements that goes into a Bayesian decision comes together. Remember, the common assumption in neuroscience, psychology, economics, ecology, etc. is that we (humans and animals) are tying to maximize our expected utility.
1. Can you find a situation where the expected utility is the same for both actions?
2. What is more important for determining the expected utility: the prior or a new measurement (the likelihood)?
3. Why is this a normative model?
4. Can you think of ways in which this model would need to be extended to describe human or animal behavior?
```
# @markdown Execute this cell to enable the widget
# style = {'description_width': 'initial'}
ps_widget = widgets.FloatSlider(0.3, description='p(s = left)',
min=0.01, max=0.99, step=0.01, layout=Layout(width='300px'))
p_a_s1_widget = widgets.FloatSlider(0.5, description='p(fish on left | state = left)',
min=0.01, max=0.99, step=0.01, style=style, layout=Layout(width='370px'))
p_a_s0_widget = widgets.FloatSlider(0.1, description='p(fish on left | state = right)',
min=0.01, max=0.99, step=0.01, style=style, layout=Layout(width='370px'))
observed_widget = ToggleButtons(options=['Fish', 'No Fish'],
description='Observation (m) on the left:', disabled=False, button_style='',
layout=Layout(width='auto', display="flex"),
style={'description_width': 'initial'}
)
widget_ui = VBox([ps_widget,
HBox([p_a_s1_widget, p_a_s0_widget]),
observed_widget])
widget_out = interactive_output(plot_prior_likelihood_utility,
{'ps': ps_widget,
'p_a_s1': p_a_s1_widget,
'p_a_s0': p_a_s0_widget,
'measurement': observed_widget})
display(widget_ui, widget_out)
# @widgets.interact(
# ps=ps_widget,
# p_a_s1=p_a_s1_widget,
# p_a_s0=p_a_s0_widget,
# m_right=observed_widget
# )
# def make_prior_likelihood_utility_plot(ps, p_a_s1, p_a_s0,m_right):
# fig = plot_prior_likelihood_utility(ps, p_a_s1, p_a_s0,m_right)
# plt.show(fig)
# plt.close(fig)
# return None
# to_remove explanation
#' 1. There are actually many (infinite) combinations that can produce the same
#. expected utility for both actions: but the posterior probabilities will always
# have to balance out the differences in the utility function. So, what is
# important is that for a given utility function, there will be some 'point
# of indifference'
#' 2. What matters is the relative information: if the prior is close to 50/50,
# then the likelihood has more infuence, if the likelihood is 50/50 given a
# measurement (the measurement is uninformative), the prior is more important.
# But the critical insite from Bayes Rule and the Bayesian approach is that what
# matters is the relative information you gain from a measurement, and that
# you can use all of this information for your decision.
#' 3. The model gives us a very precise way to think about how we *should* combine
# information and how we *should* act, GIVEN some assumption about our goals.
# In this case, if we assume we are trying to maximize expected utility--we can
# state what an animal or subject should do.
#' 4. There are lots of possible extensions. Humans may not always try to maximize
# utility; humans and animals might not be able to calculate or represent probabiltiy
# distributions exactly; The utility function might be more complicated; etc.
```
---
# Summary
In this tutorial, you learned about combining prior information with new measurements to update your knowledge using Bayes Rulem, in the context of a fishing problem.
Specifically, we covered:
* That the likelihood is the probability of the measurement given some hidden state
* That how the prior and likelihood interact to create the posterior, the probability of the hidden state given a measurement, depends on how they covary
* That utility is the gain from each action and state pair, and the expected utility for an action is the sum of the utility for all state pairs, weighted by the probability of that state happening. You can then choose the action with highest expected utility.
---
# Bonus
## Bonus Section 1: Correlation Formula
To understand the way we calculate the correlation, we need to review the definition of covariance and correlation.
Covariance:
$$
cov(X,Y) = \sigma_{XY} = E[(X - \mu_{x})(Y - \mu_{y})] = E[X]E[Y] - \mu_{x}\mu_{y}
$$
Correlation:
$$
\rho_{XY} = \frac{cov(Y,Y)}{\sqrt{V(X)V(Y)}} = \frac{\sigma_{XY}}{\sigma_{X}\sigma_{Y}}
$$
| github_jupyter |
# Introduction
In recent years we hear a lot of news about wildfires occurring in areas where they did not occur often in past. In spring and summer 2018 we have witnessed the boreal forests in Siberia burning at extraordinary rates (https://www.sciencealert.com/nasa-images-capture-worst-siberian-wildfires-in-10-000-years). Other areas are known to be prone to wildfires but the wildfires tends to be bigger and more severe. Wildfires, which erupted in Portugal in summer 2017 took 66 lives and caused an economic loss of more than 565 millions USD (http://thoughtleadership.aonbenfield.com/Documents/20180124-ab-if-annual-companion-volume.pdf). In September 2017 California, US was in flames, firefighters were fighting with more than 9000 fires, which burned more than 500 ha (https://www.fire.ca.gov/incidents/2017/).
Wildfires have many negative consequences:
- They can **damage** human properties.
- They present a **threat for human health and lives**. In addition to the direct threat from burning, wildfires also release pollutants detrimental for human health and ecosystems. Close to the fires, smoke is a health risk because it contains a mixture of hazardous gases and small particles that can irritate the eyes and respiratory system (http://www.xinhuanet.com/english/2018-07/27/c_137352746.htm).
- Vegetation fires release large amounts of **particulate matter** and toxic gases including **carbon monoxide**, **nitrogen oxides**, and **non-methane organic compounds** into the atmosphere (https://public.wmo.int/en/media/news/drought-and-heat-exacerbate-wildfires). This contribute significantly to global warming.
- Extinguishing of the fires is **risky and costly**.
However, wildfires can also be beneficial. High-severity wildfire create complex early seral forest habitat (also called “snag forest habitat”), which often has higher species richness and diversity than in unburned old forest. E.g., giant sequoias, found in the U.S. Sierra Nevada, require heat from fire to regenerate (https://video.nationalgeographic.com/video/yosemite-sequoias-fire).
While we cannot - and do no aim to - fully prevent wildfires it is important that we try to understand under what circumstances wildfires are most likely to appear, how they spread and what impact they have so that we can minimize their negative effects. Usually we try to minimize the amount of flammable material either by igniting smaller fires (controlled burning) or by logging.
# Wildfires and satellite imagery
Satellite images are images acquired with sensors ("cameras") carried on board of satellites. Images are taken from different satellites and in different parts of electromagnetic spectrum. Such images give us various information about the Earth. They are a good source of different type of information about wildfires which can help us build our understanding of wildfires and plan (re)action to their occurrence accordingly. Advances in satellite technology has made it possible to monitor wildfire activity better than in the past (https://public.wmo.int/en/media/news/drought-and-heat-exacerbate-wildfires). Collecting the information about the fire on a field can be dangerous, while satellites enable us to get such information from (more than) safe distance. It also has the advantages of covering larger areas, gathering data on less accessible areas (Leblon et al., 2012), it is time and cost effective.
Satellite imagery is, in connection to wildfires, commonly used to:
1. **Detect areas with high potential for wildfire occurrence.**
This is estimated based on vegetation maps, material (potential fuel) moisture maps both derived from satellite images. These maps are combined with other combination such as weather information and topography and distance from roads and settlements to produce fire risk maps (http://www.isprs.org/proceedings/XXXV/congress/yf/papers/927.pdf).
2. **Map areas with potential fires:**
Check the world map with potential fires from the latest satellite images:
https://fires.globalforestwatch.org/map/#activeLayers=viirsFires%2CactiveFires%2CfireStories%2Ctwitter&activeBasemap=topo&activeImagery=&planetCategory=PLANET-MONTHLY&planetPeriod=Aug%202018&x=-12&y=18&z=3
3. **Observe extent and severity of burned scar** (as we will demonstrate below)
4. **Observe impact on vegetation and its recovery** (as we will demonstrate below)
5. **Observe impact on build areas.**
As an example, check how satellite images can be used to observe the burned villages in Africa: https://www.bellingcat.com/resources/how-tos/2018/09/04/identify-burnt-villages-satellite-imagery%e2%80%8a-case-studies-california-nigeria-myanmar/
6. **Observe spread of smoke and gases.**
Check out how spread of smoke in case of wildfire sin California was captured from MODIS satellite https://www.nbcsandiego.com/news/national-international/NASA-Satellite-Images-California-Wildfire-Brush-Fires-Images-Photos-433867203.html
We will now check how some of the wildfires and their consequences were seen from space. We will use images acquired from satellite Sentinel 2 to observe the consequences of wildfire at Madeira in summer 2016 and images acquired from satellites Sentinel 2 and Sentinel S5p to observe wildfires in Siberia.
# Madeira, August 2016
Madeira is a Portuguese island in Atlantic Ocean well known for its vivid vegetation and beautiful nature. In August 2016 flames of deadly fire spread throughout the region of Southern Madeira and to its capital Funchal. More than 200 houses were destroyed, vegetation - including botanical garden near the capital - was severly damaged, 4 people died (https://www.madeiraislandnews.com/2016/08/fire-damage.html, 6.9.2018).
Let’s check how the consequences of the fire were seen from Sentinel 2 satellite:
<table>
<tr>
<td> <img src="images/2016-08-07, Sentinel-2A L1C, Custom script.png" alt="Drawing" style="width: 500px;"/> </td>
<td> <img src="images/Sentinel-2 L1C from 2016-08-17.png" alt="Drawing" style="width: 500px;"/> </td>
</tr>
<tr>
<td style="text-align:left"> <b>Before wildfire</b>, true color image acquired from Sentinel 2 on 7.8.2016 (<a href="https://apps.sentinel-hub.com/eo-browser/?lat=32.7523&lng=-17.0079&zoom=12&time=2016-08-07&preset=TRUE-COLOR&datasource=Sentinel-2%20L1C%20-%20wildfires&themesUrl=https://raw.githubusercontent.com/sentinel-hub/education/master/wildfires/themes.json&gainOverride=1.4">EO Browser link</a>). </td>
<td style="text-align:left"> <b>After wildfire</b>, true color image acquired from Sentinel 2 on 17.8.2016 (<a href="https://apps.sentinel-hub.com/eo-browser/?lat=32.7523&lng=-17.0079&zoom=12&time=2016-08-17&preset=TRUE-COLOR&datasource=Sentinel-2%20L1C%20-%20wildfires&themesUrl=https://raw.githubusercontent.com/sentinel-hub/education/master/wildfires/themes.json&gainOverride=1.4">EO Browser link</a>). </td>
</tr>
</table>
Two bigger burn scars can be observed on southern part of the island. It is difficult to distinguish them from surrounding un-burned areas, though. To make these areas easier to detect we will visualize images of the same area on the same dates acquired in near-infrared and short-wave infrared part of the spectrum. Healthy vegetation has a high reflectance in the near-infrared portion of the spectrum (NIR), while offering low short-wave infrared reflectance (SWIR). On the other hand, burned areas have a high shortwave infrared reflectance but low reflectance in the near infrared (https://www.skywatch.co/blog/assessing-impact-wildfire-normalized-burn-ratio-satellite, 30.8.2018). To emphasize these differentce we will calculate Normalized Burn Ration (NBR)(http://gsp.humboldt.edu/OLM/Courses/GSP_216_Online/lesson5-1/NBR.html):
$$NBR = \frac{(NIR-SWIR)}{(NIR+SWIR)}$$
The formula above is applied for each pixel in the image and the result is as folows:
<table>
<tr>
<td> <img src="images/2016-08-07_Sentinel-2A L1C_Custom_script_BRN.png" alt="Drawing" style="width: 500px;"/> </td>
<td> <img src="images/2016-08-17, Sentinel-2A L1C, Custom script_BRN.png" alt="Drawing" style="width: 500px;"/> </td>
</tr>
<tr>
<td style="text-align:left"> <b>NBR before wildfire</b>, on 7.8.2016 (<a href="https://apps.sentinel-hub.com/eo-browser/?lat=32.7342&lng=-17.0443&zoom=12&themesUrl=https://raw.githubusercontent.com/sentinel-hub/education/master/wildfires/themes.json&time=2016-08-07&preset=NORMALIZED-BURN-RATIO&datasource=Sentinel-2%20L1C%20-%20wildfires">EO Browser link</a>)</td>
<td style="text-align:left"> <b>NBR after wildfire</b>, on 17.8.2016 (<a href="https://apps.sentinel-hub.com/eo-browser/?lat=32.7342&lng=-17.0443&zoom=12&themesUrl=https://raw.githubusercontent.com/sentinel-hub/education/master/wildfires/themes.json&time=2016-08-17&preset=NORMALIZED-BURN-RATIO&datasource=Sentinel-2%20L1C%20-%20wildfires">EO Browser link</a>)</td>
</tr>
</table>
Analyzing the right image we notice that burned areas are visualized in darker, almost black color and it is now easier to distinguish them from the rest of areas. By digitizing a polygon around the scar we can roughly estimate the size of damaged area, which is approx. $41 km^2$. Note also that clouds and some water areas (sea) appear darker on the image but they shall not be be mistaken for burned areas.
<table>
<tr>
<td> <img src="images/Digitized_Burned_Areas.png" alt="Drawing" style="width: 800px;"/> </td>
</tr>
<td style="text-align:left"> Polygons digitized around burned areas.</td>
</table>
In order to estimate the severity of damage that wildfire left on vegetation we will calculate differences between NBR before and NBR after the wildfire. The result is than classified into 4 classes to produce a wildfire severity map.
<table>
<tr>
<td> <img src="images/Sentinel-2 image on 2017-07-03 _ dBRN.jpg" alt="Drawing" style="width: 800px;"/> </td>
</tr>
<td style="text-align:left"> Map of wildfire severity based on <b>dNBR</b>, where the most damaged area are colored dark red, severly damaged ones are orange and less damaged areas are yellow (<a href="https://sentinelshare.page.link/r2jt">EO Browser link</a>). </td>
</table>
Right after a wildfire is successfully extinguished the vegetation starts with recovery. The peace of recovery depends on the severity of damage, weather conditions, etc. A wildfire usually turns organic material to ashes so that nutrients return to the soil. Wildfire also clears thick growth so sunlight can reach the forest floor and encourage the growth of native species. Fire frees these plants from the competition delivered by invasive weeds and eliminates diseases or droves of insects that may have been causing damage to old growth (https://science.howstuffworks.com/environmental/green-science/how-forest-fire-benefit-living-things-2.htm). Recovery of vegetation can also be monitored using time series of satellite images acquired after the event.
For our example of Madeira wildfire we will calculate Normalized difference vegetation index (NDVI, https://en.wikipedia.org/wiki/Normalized_difference_vegetation_index) for burned area and compare this with NDVI calculated for healthy (unburned) vegetation. In the figure below we plot NDVI values for period of 2 years. Value of NDVI is correlated to amount of chlorophyl in vegetation. Higher the value of NDVI, more chlorophyl and healthier vegetation.
<table>
<tr>
<td> <img src="images/NDVI_Graph_UnBurned.png" alt="Drawing" style="width: 500px;"/> </td>
<td> <img src="images/NDVI_Graph_Burned.png" alt="Drawing" style="width: 500px;"/> </td>
</tr>
<tr>
<td style="text-align:left"> NDVI values of vegetation on <b>unburend</b> area. </td>
<td style="text-align:left"> NDVI values of vegetation on <b>burned</b> area. </td>
</tr>
</table>
In the left figure we can see the normal yearly cycle of vegetation with lower values in winter time (November - March) higher values in summer time (April - October). In the right figure, the decrease in NDVI value as a consequence of wildfire in August 2016 is obvious. Even two years after the wildfire the values of NDVI are still lower comparing to the values in unburned areas (note a different scale of y axis in both graphs).
# Siberia, July 2018
Not only fires and burned areas, satellite images can also be used to observe the direction in which smoke spreads from a fire or to estimate a concentration of released gasses. We will use wildfires in Siberia, Russia as an example to show how to vizualize this information.
Dry, warm conditions in the spring set the stage for fires in Siberia. The wildfires burning boreal forests in in mid of July can be nicely observed from satellite images.
<table>
<tr>
<td> <img src="images/2018-07-21, Sentinel-2A L1C, Custom script_Russia_far.jpg" alt="Drawing" style="width: 800px;"/> </td>
<td> <img src="images/2018-07-21, Sentinel-2A L1C, Custom script_Russia_close.jpg" alt="Drawing" style="width: 300px;"/> </td>
</tr>
<td style="text-align:left">Sibiria fires on 21st of July 2018. Scene is vizualized using a <a href="https://pierre-markuse.net/2018/04/30/visualizing-wildfires-burn-scars-sentinel-hub-eo-browser/">custom script</a> by Pierre Markuse. Important to note, yellow and red areas colored area shall be interpreted as the hottest areas but they do not necessarily present the active fires (<a href="https://apps.sentinel-hub.com/eo-browser/?lat=60.900&lng=92.725&zoom=8&time=2018-07-21&preset=CUSTOM&atmFilter=DOS1&datasource=Sentinel-2%20L1C&layers=B01,B02,B03&evalscript=Ly8gV2lsZGZpcmUgYW5kIGJ1cm4gc2NhciB2aXN1YWxpemF0aW9uIGluIFNlbnRpbmVsLTIgaW1hZ2VzIFYyLjAuMAovLyBUd2l0dGVyOiBQaWVycmUgTWFya3VzZSAoQHBpZXJyZV9tYXJrdXNlKQovLyBDQyBCWSA0LjAgSW50ZXJuYXRpb25hbCAtIGh0dHBzOi8vY3JlYXRpdmVjb21tb25zLm9yZy9saWNlbnNlcy9ieS80LjAvCmZ1bmN0aW9uIGEoYSwgYikge3JldHVybiBhICsgYn07CmZ1bmN0aW9uIHN0cmV0Y2godmFsLCBtaW4sIG1heCkge3JldHVybiAodmFsIC0gbWluKSAvIChtYXggLSBtaW4pO30KZnVuY3Rpb24gc2F0RW5oKHJnYkFycikgewogICAgdmFyIGF2ZyA9IHJnYkFyci5yZWR1Y2UoKGEsIGIpID0%2BIGEgKyBiLCAwKSAvIHJnYkFyci5sZW5ndGg7CiAgICByZXR1cm4gcmdiQXJyLm1hcChhID0%2BIGF2ZyAqICgxIC0gc2F0dXJhdGlvbikgKyBhICogc2F0dXJhdGlvbik7IH0KZnVuY3Rpb24gaGlnaGxpZ2h0QnVybnNjYXIodmFsLCBvTG93LCBvSGlnaCwgZGVTYXQsIGRhcmtlbikgewogICAgaWYgKChCMTIgKyBCMTEgPiAwLjA1KSAmJiAodmFsID4gMCkpIHsKICAgICAgICBpZiAoKChCOEEgLSBCMTIpIC8gKEI4QSArIEIxMikpID4gb0xvdykgewogICAgICAgICAgICBzYXR1cmF0aW9uID0gc2F0dXJhdGlvbiAtIGRlU2F0OwogICAgICAgICAgICBzdHJldGNoTWF4ID0gc3RyZXRjaE1heCArIGRhcmtlbjsKICAgICAgICB9IGVsc2UgewogICAgICAgICAgICBpZiAoKChCOEEgLSBCMTIpIC8gKEI4QSArIEIxMikpIDw9IG9IaWdoKSB7CiAgICAgICAgICAgICAgICBub0ZpcmVbMF0gPSBub0ZpcmVbMF0gKyAwLjIgKiB2YWw7CiAgICAgICAgICAgICAgICBub0ZpcmVbMV0gPSBub0ZpcmVbMV0gKyAwLjA1ICogdmFsOwogICAgICAgICAgICB9IGVsc2UgewogICAgICAgICAgICAgICAgbm9GaXJlWzBdID0gbm9GaXJlWzBdICsgMC4xNSAqIHZhbDsKICAgICAgICAgICAgICAgIG5vRmlyZVsxXSA9IG5vRmlyZVsxXSArIDAuMTUgKiB2YWw7CiAgICAgICAgICAgIH0KICAgICAgICB9CiAgICB9Cn0KZnVuY3Rpb24gaW5kZXhNYXAoaW5kLCBsVmFsLCBtVmFsLCBoVmFsLCBjb250LCBkaXIsIHBhbCkgewogIHZhciBjb2wxPUdSRUVOO3ZhciBjb2wyPVlFTExPVzt2YXIgY29sMz1SRUQ7ICAKICBpZiAocGFsID09IDEpIHtjb2wxPUNCTDtjb2wyPUNCTTtjb2wzPUNCSDt9IAogIGlmIChwYWwgPT0gMikge2NvbDE9T1dOTDtjb2wyPU9XTk07Y29sMz1PV05IO30gICAgICAgICAKICAgIHZhciBsVmFsQ29sID0gY29sMTsKICAgIHZhciBtVmFsQ29sID0gY29sMjt2YXIgaFZhbENvbCA9IGNvbDM7CiAgICBpZiAoZGlyID09IDEpewogICAgCWxWYWxDb2wgPSBjb2wzO2hWYWxDb2wgPSBjb2wxOwogICAgfQogICAgaWYgKGNvbnQgPT0gMCkgewogICAgICAgIGlmIChpbmQgPD0gbFZhbCkgcmV0dXJuIGxWYWxDb2w7IGlmICgoaW5kID4gbFZhbCkgJiYgKGluZCA8IGhWYWwpKSByZXR1cm4gbVZhbENvbDsgaWYgKGluZCA%2BPSBoVmFsKSByZXR1cm4gaFZhbENvbDsKICAgICAgfSBlbHNlIHsKICAgICAgICByZXR1cm4gY29sb3JCbGVuZChpbmQsIFtsVmFsLCBtVmFsLGhWYWxdLCBbbFZhbENvbCxtVmFsQ29sLGhWYWxDb2xdKTsKICAgIH0KfQpmdW5jdGlvbiBibGVuZChiQXJyMSwgYkFycjIsIG9wYTEsIG9wYTIpIHsKICAgIHJldHVybiBiQXJyMS5tYXAoZnVuY3Rpb24obnVtLCBpbmRleCkgewogICAgICAgIHJldHVybiAobnVtIC8gMTAwICogb3BhMSArIGJBcnIyW2luZGV4XSAvIDEwMCAqIG9wYTIpOwogICAgfSk7Cn0KZnVuY3Rpb24gYXBwbHlFbmgoYkFycikgewogICAgaGlnaGxpZ2h0QnVybnNjYXIoYnVybnNjYXJIaWdobGlnaHQsIGJ1cm5zY2FyVGhyZXNob2xkTG93LCBidXJuc2NhclRocmVzaG9sZEhpZ2gsIGJ1cm5zY2FyRGVzYXR1cmF0ZUJhY2tkcm9wLCBidXJuc2NhckRhcmtlbkJhY2tkcm9wKTsKICAgIHJldHVybiBzYXRFbmgoW3N0cmV0Y2goYkFyclswXSwgc3RyZXRjaE1pbiwgc3RyZXRjaE1heCksIHN0cmV0Y2goYkFyclsxXSwgc3RyZXRjaE1pbiwgc3RyZXRjaE1heCksIHN0cmV0Y2goYkFyclsyXSwgc3RyZXRjaE1pbiwgc3RyZXRjaE1heCldKTsKfQp2YXIgQkxBQ0sgPSBbMC4wLCAwLjAsIDAuMF07CnZhciBSRUQgPSBbMC45LCAwLjEsIDAuMV07CnZhciBZRUxMT1cgPSBbMC45LCAwLjksIDAuMV07CnZhciBHUkVFTiA9IFswLjAsIDAuNiwgMC4wXTsKdmFyIENCTCAgPSBbMC8yNTUsIDgwLzI1NSwgMC8yNTVdOwp2YXIgQ0JNICA9IFsxMjAvMjU1LCAxMjAvMjU1LCAyMzAvMjU1XTsKdmFyIENCSCAgPSBbNzAvMjU1LCAxOTUvMjU1LCAyNTUvMjU1XTsKdmFyIE9XTkwgPSBbMC4wLCAwLjAsIDAuMF07CnZhciBPV05NID0gWzAuMCwgMC4wLCAwLjBdOwp2YXIgT1dOSCA9IFswLjAsIDAuMCwgMC4wXTsKLy8gVmlzdWFsaXphdGlvbiBzdHlsZSBvZiB0aGUgZGlmZmVyZW50IGZpcmUgem9uZXMKdmFyIEZpcmUxT1ZMID0gW3N0cmV0Y2goKDIuMSAqIEIwNCArIDAuNSAqIEIxMiksIDAuMDEsIDAuOTkpICsgMS4xLCBzdHJldGNoKCgyLjIgKiBCMDMgKyAwLjUgKiBCMDgpLCAwLjAxLCAwLjk5KSwgc3RyZXRjaCgyLjEgKiBCMDIsIDAuMDEsIDAuOTkpXTsKdmFyIEZpcmUyT1ZMID0gW3N0cmV0Y2goKDIuMSAqIEIwNCArIDAuNSAqIEIxMiksIDAuMDEsIDAuOTkpICsgMS4xLCBzdHJldGNoKCgyLjIgKiBCMDMgKyAwLjUgKiBCMDgpLCAwLjAxLCAwLjk5KSArIDAuMjUsIHN0cmV0Y2goMi4xICogQjAyLCAwLjAxLCAwLjk5KV07CnZhciBGaXJlM09WTCA9IFtzdHJldGNoKCgyLjEgKiBCMDQgKyAwLjUgKiBCMTIpLCAwLjAxLCAwLjk5KSArIDEuMSwgc3RyZXRjaCgoMi4yICogQjAzICsgMC41ICogQjA4KSwgMC4wMSwgMC45OSkgKyAwLjUsIHN0cmV0Y2goMi4xICogQjAyLCAwLjAxLCAwLjk5KV07Ci8vIEJhbmQgY29tYmluYXRpb25zIChUbyBnZXQgcXVpY2tlciBwcm9jZXNzaW5nIHlvdSBzaG91bGQgY29tbWVudCBvdXQgYWxsIHRob3NlIHlvdSBhcmUgbm90IHVzaW5nIGluIHRoZSBTZXR0aW5ncyBmdXJ0aGVyIGRvd24pCnZhciBOYXR1cmFsQ29sb3JzID0gWzIuOSAqIEIwNCwgMy4xICogQjAzLCAzLjAgKiBCMDJdOwovLyB2YXIgRW5oYW5jZWROYXR1cmFsQ29sb3JzID0gWzIuOCAqIEIwNCArIDAuMSAqIEIwNSwgMi44ICogQjAzICsgMC4xNSAqIEIwOCwgMi44ICogQjAyXTsKdmFyIE5hdHVyYWxOSVJTV0lSTWl4ID0gWzIuMSAqIEIwNCArIDAuNSAqIEIxMiwgMi4yICogQjAzICsgMC41ICogQjA4LCAzLjAgKiBCMDJdOwovLyB2YXIgTklSU1dJUkNvbG9yczEgPSBbMi42ICogQjEyLCAxLjkgKiBCMDgsIDIuNyAqIEIwMl07Ci8vdmFyIE5JUlNXSVJDb2xvcnMyID0gWzIuNCAqIEIxMiwgMS43ICogQjhBLCAyLjIgKiBCMDVdOwovLyB2YXIgTklSU1dJUkNvbG9yczMgPSBbMC41ICogKEIxMiArIEIxMSkgLyA0IC8gQjA3LCAwLjggKiBCOEEsIDEgKiBCMDddOwovLyB2YXIgTklSU1dJUkNvbG9yczQgPSBbMi4wICogQjEyLCAxLjEgKiBCMTEsIDEuNiAqIEIwOF07Ci8vIHZhciBGYWxzZUNvbG9yID0gW0IwOCAqIDIsIEIwNCAqIDIsIEIwMyAqIDJdOwovLyB2YXIgTmF0RmFsc2VDb2xvciA9IFtCMTIgKiAyLjYsIEIxMSAqIDIsIEIwNCAqIDIuN107Ci8vIHZhciBWZWdldGF0aW9uID0gW0IxMSAqIDIuNCwgQjhBICogMiwgQjA0ICogMi45XTsKLy8gdmFyIFBhbkJhbmQgPSBbQjA4LCBCMDgsIEIwOF07Ci8vIHZhciBOQlI4QTEyID0gaW5kZXhNYXAoKEI4QSAtIEIxMikgLyAoQjhBICsgQjEyKSwgLTAuOCwgLTAuNCwgMC4wLCAxLCAxLCAxKTsKLy8gdmFyIE5EVkkgPSBpbmRleE1hcCgoQjA4IC0gQjA0KSAvIChCMDggKyBCMDQpLCAtMC40LCAtMC4yLCAwLjAsIDEsIDEsIDEpOwovLyBTZXR0aW5ncwovLyBGaXJlIChob3Qgc3BvdCkgdmlzdWFsaXphdGlvbgp2YXIgZmlyZTEgPSBGaXJlMU9WTDsKdmFyIGZpcmUyID0gRmlyZTJPVkw7CnZhciBmaXJlMyA9IEZpcmUzT1ZMOwovLyBVc2VkIGJhbmQgY29tYmluYXRpb25zIGFuZCBtaXhpbmcKdmFyIGxheWVyMSA9IE5hdHVyYWxOSVJTV0lSTWl4Owp2YXIgbGF5ZXIyID0gTmF0dXJhbENvbG9yczsKdmFyIGxheWVyMUFtb3VudCA9IDEwMAp2YXIgbGF5ZXIyQW1vdW50ID0gMDsKLy8gSW5mbHVlbmNlIGNvbnRyYXN0IGFuZCBzYXR1cmF0aW9uCnZhciBzdHJldGNoTWluID0gMC4wMDsKdmFyIHN0cmV0Y2hNYXggPSAxLjAwOwp2YXIgc2F0dXJhdGlvbiA9IDEuMDA7Ci8vIEZpcmUgc2Vuc2l0aXZpdHkgKERlZmF1bHQgPSAxLjAwKSwgaGlnaGVyIHZhbHVlcyBpbmNyZWFzZSBmaXJlIChob3Qgc3BvdCkgZGV0ZWN0aW9uIGFuZCBmYWxzZSBwb3NpdGl2ZXMKdmFyIGZpcmVTZW5zaXRpdml0eSA9IDEuMDA7Ci8vIEJ1cm4gc2NhciB2aXN1YWxpemF0aW9uCnZhciBidXJuc2NhckhpZ2hsaWdodCA9IDAuMDA7CnZhciBidXJuc2NhclRocmVzaG9sZExvdyA9IC0wLjI1Owp2YXIgYnVybnNjYXJUaHJlc2hvbGRIaWdoID0gLTAuMzg7CnZhciBidXJuc2NhckRlc2F0dXJhdGVCYWNrZHJvcCA9IDAuMjU7CnZhciBidXJuc2NhckRhcmtlbkJhY2tkcm9wID0gMC4yNTsKLy8gTWFudWFsbHkgaW5mbHVlbmNlIFJHQiBvdXRwdXQKdmFyIG1hbnVhbENvcnJlY3Rpb24gPSBbMC4wMCwgMC4wMCwgMC4wMF07Ci8vIEltYWdlIGdlbmVyYXRpb24gYW5kIG91dHB1dApub0ZpcmUgPSBibGVuZChsYXllcjEsIGxheWVyMiwgbGF5ZXIxQW1vdW50LCBsYXllcjJBbW91bnQpOwpmaW5hbFJHQiA9IGFwcGx5RW5oKG5vRmlyZSkubWFwKGZ1bmN0aW9uKG51bSwgaW5kZXgpIHsKICAgIHJldHVybiBudW0gKyBtYW51YWxDb3JyZWN0aW9uW2luZGV4XTt9KTsKcmV0dXJuIChhKEIxMiwgQjExKSA%2BICgxLjAgLyBmaXJlU2Vuc2l0aXZpdHkpKSA%2FCiAgICAoYShCMTIsIEIxMSkgPiAoMi4wIC8gZmlyZVNlbnNpdGl2aXR5KSkgPyBmaXJlMyA6CiAgICAoYShCMTIsIEIxMSkgPiAoMS41IC8gZmlyZVNlbnNpdGl2aXR5KSkgPyBmaXJlMiA6IGZpcmUxIDoKICAgZmluYWxSR0I7">EO Browser link</a>).</td>
<td style="text-align:left">Zoomed in to the area marked with blue in the left figure. Scene is vizualized using a <a href="https://pierre-markuse.net/2018/04/30/visualizing-wildfires-burn-scars-sentinel-hub-eo-browser/">custom script</a> by Pierre Markuse. Important to note, yellow and red areas shall be interpreted as the hottest areas but they do not necessarily present the active fires (<a href="https://sentinelshare.page.link/69op">EO Browser link</a>).</td>
</table>
Vegetation fires release large amounts of particulate matter and toxic gases including carbon monoxide (CO), nitrogen oxides (NO), and non-methane organic compounds into the atmosphere. Measurements of these exhaustions are essential for forecasts, research on atmospheric composition and to develop warning systems (https://public.wmo.int/en/media/news/drought-and-heat-exacerbate-wildfires, 7.9.2018). Special sensors are needed to observe these gases and they are on board of Satellite Sentinel 5p. For the area a bit larger than the one shown in figures above, the Sentinel 5p measured increased concentration of CO as a consequence of the wildfire.
<table>
<tr>
<td> <img src="images/S5p_6_7_2018.png" alt="Drawing" style="width: 300px;"/> </td>
<td> <img src="images/S5p_21_7_2018.png" alt="Drawing" style="width: 300px;"/> </td>
</tr>
<td style="text-align:left"> Map of CO concentration <b>before</b> the wildfire (<a href="https://sentinelshare.page.link/vtWX">get image from Sentinel-Hub</a>). White pixels in the image are pixels with no data probably because of thick clouds. </td>
<td style="text-align:left"> Map of CO concentration <b>after</b> the wildfire (<a href="https://sentinelshare.page.link/E6RD">get image from Sentinel-Hub</a>). White pixels in hte image are pixels with no data probably because of thick clouds.</td>
</table>
Besides CO, wildfires also release carbon dioxide (CO2) into the atmosphere, contributing to global warming. For instance, fires burned around 3 million hectares of land in Indonesia during the 2015 dry season released about 11.3 teragramms of CO2, which is roughly 120% of daily release of CO2 from fossil fuel burning in the European Union https://public.wmo.int/en/media/news/drought-and-heat-exacerbate-wildfires, 7.9.2018).
# Conclusions
We have learned that several aspects of wildfires and their influence on the environment can be observed from satellite images. We visualized and measured the extent of burned area and severity of burn scar for wildfire in Madeira. The same approach can be used for any other wildfire. We checked what was the influence on vegetation and how long it took it to recover by inspecting NDVI. We also checked how the smoke and CO spread form the fires in Siberia earlier this year.
However, whenever satellite data is used for we need to keep in mind the limitation of the data. Images are acquired from space and light must travel through different layers of atmosphere before it reaches the sensors, which can influence the accuracy of the images. Spatial and temporal resolutions of images are limited. Clouds or shades can obscure scenes in which we are interested as we experienced in the example of Madeira. We need to take these factors into consideration when interpreting satellite images or when making decisions based on such analysis.
# Other Resources to Check
If you find this topic interesting, you might also want to check out:
- EO Browser: https://apps.sentinel-hub.com/eo-browser/
- Sentinel Playground: https://apps.sentinel-hub.com/sentinel-playground/
- Global for est watch: https://fires.globalforestwatch.org/map/#activeLayers=viirsFires%2CactiveFires%2CfireStories%2Ctwitter&activeBasemap=topo&activeImagery=&planetCategory=PLANET-MONTHLY&planetPeriod=Aug%202018&x=-12&y=18&z=3
- European, Rapid Damage Assessment: http://effis.jrc.ec.europa.eu/about-effis/technical-background/rapid-damage-assessment/
- Copernicus, Emergency Management Service http://effis.jrc.ec.europa.eu/static/effis_current_situation/public/index.html
- Drought and heat exacerbate wildfires: https://public.wmo.int/en/media/news/drought-and-heat-exacerbate-wildfires
- From California to Siberia: satellite images of wildfires around the world: https://unearthed.greenpeace.org/2018/08/10/california-wildfires-nasa-satellite-map/
- Landsat Image Maps Aid Fire Recovery Efforts: https://landsat.gsfc.nasa.gov/landsat-image-maps-aid-fire-recovery-efforts/
- Detection of burned areas with Machine Learning: https://webthesis.biblio.polito.it/8197/1/tesi.pdf
- Assessing the impact of a wildfire with satellites: https://www.skywatch.co/blog/assessing-impact-wildfire-normalized-burn-ratio-satellite
- How does a forest fire benefit living things?: https://science.howstuffworks.com/environmental/green-science/how-forest-fire-benefit-living-things-2.htm
- Brigitte Leblon, Laura Bourgeau-Chavez and Jesús San-Miguel-Ayanz (August 1st 2012). Use of Remote Sensing in Wildfire Management, Sustainable Development Sime Curkovic, IntechOpen, DOI: 10.5772/45829. Available from: https://www.intechopen.com/books/sustainable-development-authoritative-and-leading-edge-content-for-environmental-management/use-of-remote-sensing-in-wildfire-management
| github_jupyter |
```
import os
os.chdir('..')
%load_ext autoreload
%autoreload 2
import torch
from src.main import setup_torch, get_corpus
from src.utils import get_latest_model_file
from src.winograd_schema_challenge import (
analyse_single_wsc, generate_full_sentences, find_missing_wsc_words_in_corpus_vocab, winograd_test
)
from src.html_wsc_parser import generate_df
```
# Sentences with issues:
```
still_in_english = [
"It was a summer afternoon, and the dog was sitting in the middle of the lawn. After a while, it got up and moved to a spot under the tree, because the spot under the tree was hot.",
"It was a summer afternoon, and the dog was sitting in the middle of the lawn. After a while, it got up and moved to a spot under the tree, because the dog was cooler.",
"I couldn't put the pot on the shelf because the shelf was too tall.",
"I couldn't put the pot on the shelf because the pot was too high.",
"There is a pillar between me and the stage, and I can't see around the stage.",
"There is a pillar between me and the stage, and I can't see the pillar.",
"They broadcast an announcement, but a subway came into the station and I couldn't hear the subway.",
"They broadcast an announcement, but a subway came into the station and I couldn't hear over the announcement.",
# "The large ball crashed right through the table because the table was made of steel.",
# "The large ball crashed right through the table because the large ball was made of styrofoam."
]
subs_not_working = [
'Jim fez um sinal para o barman e apontou para sua taça vazia.',
'Jim fez um sinal para o barman e apontou para jim chave do banheiro.',
'Dan ficou no banco de trás enquanto Bill reinvidicava a frente porque Bill "Eu primeiro!" era lento.',
'Dan ficou no banco de trás enquanto Bill reinvidicava a frente porque Dan "Eu primeiro!" era mais rápido.',
'Tom disse "Check" para Ralph enquanto ele movia Ralph bispo.',
'Tom disse "Check" para Ralph enquanto ele tomava Tom bispo.',
'John contratou Bill para cuidar Bill.',
'John foi contratado por Bill para cuidar John.',
'Ele os abaixou quando o vento parou.',
'Ele os abriu quando o vento parou.',
'As raposas estão entrando à noite e atacando as galinhas. Eu terei que matá- las.',
'As raposas estão entrando à noite e atacando as galinhas. Eu terei que protegê- las.',
'O caminho até o lago estava interditado, então nós não podíamos usá-o lago.',
'O caminho até o lago estava interditado, então nós não podíamos alcançá-o caminho.',
'Sara pegou um livro emprestado da biblioteca porque ela precisa dele para um artigo no qual ela está trabalhando. Ela o artigo lê quando chega do trabalho.',
'Sara pegou um livro emprestado da biblioteca porque ela precisa dele para um artigo no qual ela está trabalhando. Ela o livro escreve quando chega do trabalho.',
'Como estava chovendo, eu carreguei a revista na minha mochila para mantê-a mochila seca.',
'Como estava chovendo, eu carreguei a revista sobre a minha mochila para mantê-a revista seca.',
'Mary sacou sua flauta e tocou uma de suas peças favoritas. Ela a peça tem desde criança.',
'Mary sacou sua flauta e tocou uma de suas peças favoritas. Ela a flauta ama desde criança.',
'Joe vendeu sua casa e comprou uma nova a alguns quilômetros de distância. Ele se mudará dela na quinta-feira.',
'Durante a tempestade, o poste caiu e atravessou o telhado da minha casa. Agora, eu tenho que mandar removêo telhado.',
'Durante a tempestade, o poste caiu e atravessou o telhado da minha casa. Agora, eu tenho que mandar consertáo posto.',
'Eu estava tentando abrir o cadeado com a chave, mas alguém havia preenchido a fechadura com goma de mascar, e eu não conseguia inseri-la .',
'Eu estava tentando abrir o cadeado com a chave, mas alguém havia preenchido a fechadura com goma de mascar, e eu não conseguia removê-la.',
'Há uma fenda na parede. É possível enxergar o jardim através a parede.',
'Há uma fenda na parede. É possível enxergar o jardim atrás a fenda.',
# ok mas estranho:
'Eu usei um pano velho para limpar o alicate, e então o alicate coloquei no lixo.',
'Eu usei um pano velho para limpar o alicate, e então o pano coloquei na gaveta.',
' Os alunos mais velhos estavam fazendo bullying com os mais jovens, então nós os alunos mais jovens punimos.',
'Os alunos mais velhos estavam fazendo bullying com os mais jovens, então nós os alunos mais velhos protegemos.'
# "policia estavam", errado
'Os bombeiros chegaram depois da polícia porque a polícia estavam vindo de muito longe.',
'Os bombeiros chegaram antes da polícia porque os bombeiros estavam vindo de muito longe.',
'O homem levantou o menino sobre sua cabeça.',
'O homem levantou o menino até sua beliche.',
'Alongando suas costas, a mulher sorriu para a menina.',
'Dando um tapinha em suas costas, a mulher sorriu para a menina.',
'Billy chorou porque Toby não queria aceitar seu brinquedo.',
'Billy chorou porque Toby não queria compartilhar seu brinquedo.',
'Lily conversou com Donna, quebrando sua promessa.',
'Lily conversou com Donna, tirando sua concentração.',
'Quando Tommy derrubou seu sorvete, Timmy gargalhou, então seu pai olhou-o com olhar empático.',
'Quando Tommy derrubou seu sorvete, Timmy gargalhou, então seu pai olhou-o com olhar severo.',
'Enquanto Ollie carregava Tommy subindo a longa escada em espiral, suas pernas doíam.',
'Enquanto Ollie carregava Tommy subindo a longa escada em espiral, suas pernas balançavam.',
'O pai carregava o menino dormindo em seu colo',
'O pai carregava o menino dormindo em seu berço',
'A mulher estava segurando a menina contra seu peito',
'A mulher estava segurando a menina contra seu desejo.',
'Este livro introduziu Shakespeare a Ovídio; ele foi uma grande influência em sua escrita.',
'Este livro introduziu Shakespeare a Goethe; ele foi uma grande influência em sua escrita.',
'Este livro introduziu Shakespeare a Ovídio; ele era uma excelente seleção de sua escrita.',
'Este livro introduziu Shakespeare a Goethe; ele era uma excelente seleção de sua escrita.'
]
df = generate_df(still_in_english, subs_not_working)
df['correct_sentence'], df['incorrect_sentence'] = zip(*df.apply(generate_full_sentences, axis=1))
for index, row in df.iterrows():
print(row.correct_sentence)
for index, row in df.iterrows():
print(row.incorrect_sentence)
setup_torch()
device = torch.device("cuda")
corpus = get_corpus()
ntokens = len(corpus.dictionary)
# TODO remove these two lines
assert ntokens == 602755
assert corpus.valid.size()[0] == 11606861
assert corpus.train.max() < ntokens
assert corpus.valid.max() < ntokens
assert corpus.test.max() < ntokens
model_file_name = get_latest_model_file()
find_missing_wsc_words_in_corpus_vocab(df, corpus)
# for item in df[df.incorrect_sentence.str.contains('fred')].incorrect_sentence:
# print(item)
df[df.incorrect_sentence.str.contains('fred')].incorrect_sentence
df.iloc[114].correct_sentence = 'Fred e Alice tinham casacos muito quentes, mas Fred e Alice não estavam preparados para o frio do Alasca.'
df.iloc[115].incorrect_sentence = 'Fred e Alice tinham casacos muito quentes, mas Fred e Alice não eram suficientes para o frio do Alasca.'
df.iloc[130].correct_sentence = 'Fred é a única pessoa viva que lembra de meu pai enquanto criança. Quando Fred viu meu pai pela primeira vez, Fred tinha doze anos de idade.'
df.iloc[131].incorrect_sentence = 'Fred é a única pessoa viva que lembra de meu pai enquanto criança. Quando Fred viu meu pai pela primeira vez, Fred tinha doze meses de idade.'
df.iloc[132].correct_sentence = 'Fred é a única pessoa viva que lembra de meu pai enquanto criança. Quando Fred viu meu pai pela primeira vez, Fred tinha doze anos de idade.'
df.iloc[133].incorrect_sentence = 'Fred é a única pessoa viva que lembra de meu pai enquanto criança. Quando Fred viu meu pai pela primeira vez, Fred tinha doze meses de idade.'
df.iloc[112].correct_sentence = 'Mary acomodou sua filha Anne na cama, para que Mary pudesse trabalhar.'
df.iloc[112].incorrect_sentence = 'Mary acomodou sua filha Anne na cama, para que a filha da Mary pudesse trabalhar.'
df.iloc[113].correct_sentence = 'Mary acomodou sua filha Anne na cama, para que a filha da Mary pudesse dormir.'
df.iloc[113].incorrect_sentence = 'Mary acomodou sua filha Anne na cama, para que Mary pudesse dormir.'
find_missing_wsc_words_in_corpus_vocab(df, corpus)
df, accuracy = winograd_test(df, corpus, model_file_name, ntokens, device)
print('Acurácia: {} para teste realizado com {} exemplos'.format(accuracy, len(df)))
len(df[~df.test_result])
len(df[df.test_result])
len(df)
if analyse_single_wsc(model_file_name, corpus, ntokens, device,
'A medalha não cabe na maleta porque a maleta é pequena .',
'A medalha não cabe na maleta porque a medalha é pequena .'):
print('Right choice :D')
else:
print('Wrong :(')
177/283
df, accuracy = winograd_test(df, corpus, model_file_name, ntokens, device, partial=True)
print('Acurácia: {} para teste realizado com {} exemplos'.format(accuracy, len(df)))
```
| github_jupyter |
# Structured & Time Series Data
This notebook walks through an implementation of a deep learning model for structured time series data using Keras. We’ll use the dataset from Kaggle’s [Rossmann Store Sales competition](https://www.kaggle.com/c/rossmann-store-sales). The steps outlined below are inspired by (and partially based on) lesson 3 of Jeremy Howard’s [fast.ai course](http://course.fast.ai) where he builds a model for the Rossman dataset using PyTorch and the fast.ai library.
The focus here is on implementing a deep learning model for structured data. I’ve skipped a bunch of pre-processing steps that are specific to this particular dataset but don’t reflect general principles about applying deep learning to tabular datasets. If you’re interested, you’ll find complete step-by-step instructions on creating the “joined” dataset in [this notebook](https://github.com/fastai/fastai/blob/master/courses/dl1/lesson3-rossman.ipynb). With that, let’s get started!
First we need to get a few imports out of the way. All of these should come standard with an Anaconda install. I’m also specifying the path where I’ve pre-saved the “joined” dataset that we’ll use as a starting point (created from running the first few sections of the above-referenced notebook).
(As an aside, I’m using [Paperspace](https://www.paperspace.com) to run this notebook. If you’re not familiar with it, Paperspace is a cloud service that lets you rent GPU instances much cheaper than AWS. It’s a great way to get started if you don’t have your own hardware.)
```
%matplotlib inline
import datetime
import matplotlib.pyplot as plt
import pandas as pd
from sklearn.decomposition import PCA
from sklearn.preprocessing import LabelEncoder, StandardScaler
PATH = '/home/paperspace/data/rossmann/'
```
Read the data file into a pandas dataframe and take a peek at the data to see what we’re working with.
```
data = pd.read_feather(f'{PATH}joined')
data.shape
data.head().T.head(93)
```
The data consists of ~800,000 records with a variety of features used to predict sales at a given store on a given day. As mentioned before, we’re skipping over details about where these features came from as it’s not the focus of this notebook, but you can find more info through the links above. Next we’ll define variables that group our features into continuous and categorical buckets. This is very important as neural networks (really anything other than tree models) do not natively handle categorical data well.
```
target = 'Sales'
cat_vars = ['Store', 'DayOfWeek', 'Year', 'Month', 'Day', 'StateHoliday', 'CompetitionMonthsOpen',
'Promo2Weeks', 'StoreType', 'Assortment', 'PromoInterval', 'CompetitionOpenSinceYear', 'Promo2SinceYear',
'State', 'Week', 'Events', 'Promo_fw', 'Promo_bw', 'StateHoliday_fw', 'StateHoliday_bw',
'SchoolHoliday_fw', 'SchoolHoliday_bw']
cont_vars = ['CompetitionDistance', 'Max_TemperatureC', 'Mean_TemperatureC', 'Min_TemperatureC',
'Max_Humidity', 'Mean_Humidity', 'Min_Humidity', 'Max_Wind_SpeedKm_h',
'Mean_Wind_SpeedKm_h', 'CloudCover', 'trend', 'trend_DE',
'AfterStateHoliday', 'BeforeStateHoliday', 'Promo', 'SchoolHoliday']
```
Set some reasonable default values for missing information so our pre-processing steps won’t fail.
```
data = data.set_index('Date')
data[cat_vars] = data[cat_vars].fillna(value='')
data[cont_vars] = data[cont_vars].fillna(value=0)
```
Now we can do something with the categorical variables. The simplest first step is to use scikit-learn’s LabelEncoder class to transform the raw category values (many of which are plain text) into unique integers, where each integer maps to a distinct value in that category. The code block below saves the fitted encoders (we’ll need them later) and prints out the unique labels that each encoder found.
```
encoders = {}
for v in cat_vars:
le = LabelEncoder()
le.fit(data[v].values)
encoders[v] = le
data.loc[:, v] = le.transform(data[v].values)
print('{0}: {1}'.format(v, le.classes_))
```
Split the data set into training and validation sets. To preserve the temporal nature of the data and make sure that we don’t have any information leaks, we’ll just take everything past a certain date and use that as our validation set.
```
train = data[data.index < datetime.datetime(2015, 7, 1)]
val = data[data.index >= datetime.datetime(2015, 7, 1)]
X = train[cat_vars + cont_vars].copy()
X_val = val[cat_vars + cont_vars].copy()
y = train[target].copy()
y_val = val[target].copy()
```
Next we can apply scaling to our continuous variables. We can once again leverage scikit-learn and use the StandardScaler class for this. The proper way to apply scaling is to “fit” the scaler on the training data and then apply the same transformation to both the training and validation data (this is why we had to split the data set in the last step).
```
scaler = StandardScaler()
X.loc[:, cont_vars] = scaler.fit_transform(X[cont_vars].values)
X_val.loc[:, cont_vars] = scaler.transform(X_val[cont_vars].values)
```
Normalize the data types that each variable is stored as. This is not strictly necessary but helps save storage space (and potentially processing time, although I’m not sure about that one).
```
for v in cat_vars:
X[v] = X[v].astype('int').astype('category').cat.as_ordered()
X_val[v] = X_val[v].astype('int').astype('category').cat.as_ordered()
for v in cont_vars:
X[v] = X[v].astype('float32')
X_val[v] = X_val[v].astype('float32')
```
Let’s take a look at where we’re at. The data should basically be ready to move into the modeling phase.
```
X.shape, X_val.shape, y.shape, y_val.shape
X.head()
X.dtypes
```
We now basically have two options when it comes to handling of categorical variables. The first option, which is the “traditional” way of handling categories, is to do a one-hot encoding for each category. This approach would create a binary variable for each unique value in each category, with the value being a 1 for the “correct” category and 0 for everything else. One-hot encoding works fairly well and is quite easy to do (there’s even a scikit-learn class for it), however it’s not perfect. It’s particularly challenging with high-cardinality variables because it creates a very large, very sparse array that’s hard to learn from.
Fortunately there’s a better way, which is something called entity embeddings or category embeddings (I don’t think there’s a standard name for this yet). Jeremy covers it extensively in the class (also [this blog post](https://towardsdatascience.com/deep-learning-structured-data-8d6a278f3088) explains it very well). The basic idea is to create a distributed representation of the category using a vector of continuous numbers, where the length of the vector is lower than the cardinality of the category. The key insight is that this vector is learned by the network. It’s part of the optimization graph. This allows the network to model complex, non-linear interactions between categories and other features in your input. It’s quite useful, and as we’ll see at the end, these embeddings can be used in interesting ways outside of the neural network itself.
In order to build a model using embeddings, we need to do some more prep work on our categories. First, let’s create a list of category names along with their cardinality.
```
cat_sizes = [(c, len(X[c].cat.categories)) for c in cat_vars]
cat_sizes
```
Now we need to decide on the length of each embedding vector. Jeremy proposed using a simple formula: cardinality / 2, with a max of 50.
```
embedding_sizes = [(c, min(50, (c + 1) // 2)) for _, c in cat_sizes]
embedding_sizes
```
One last pre-processing step. Keras requires that each “input” into the model be fed in as a separate array, and since each embedding has its own input, we need to do some transformations to get the data in the right format.
```
X_array = []
X_val_array = []
for i, v in enumerate(cat_vars):
X_array.append(X.iloc[:, i])
X_val_array.append(X_val.iloc[:, i])
X_array.append(X.iloc[:, len(cat_vars):])
X_val_array.append(X_val.iloc[:, len(cat_vars):])
len(X_array), len(X_val_array)
```
Okay! We’re finally ready to get to the modeling part. Let’s get some imports out of the way. I’ve also defined a custom metric to calculate root mean squared percentage error, which was originally used by the Kaggle competition to score this data set.
```
from keras import backend as K
from keras import regularizers
from keras.models import Sequential
from keras.models import Model
from keras.layers import Activation, BatchNormalization, Concatenate
from keras.layers import Dropout, Dense, Input, Reshape
from keras.layers.embeddings import Embedding
from keras.optimizers import Adam
from keras.callbacks import ModelCheckpoint, ReduceLROnPlateau
def rmspe(y_true, y_pred):
pct_var = (y_true - y_pred) / y_true
return K.sqrt(K.mean(K.square(pct_var)))
```
Now for the model itself. I tried to make this a similar to Jeremy’s model as I could, although there are some slight differences. The “for” section at the top shows how to add embeddings. They then get concatenated together and we apply dropout to the unified embedding layer. Next we concatenate the output of that layer with our continuous inputs and feed the whole thing into a dense layer. From here on it’s pretty standard stuff. The only notable design choice is I omitted batch normalization because it seemed to hurt performance no matter what I did. I also increased dropout a bit from what Jeremy had in his PyTorch architecture for this data. Finally, note the inclusion of the “rmspe” function as a metric during the compile step (this will show up later during training).
```
def EmbeddingNet(cat_vars, cont_vars, embedding_sizes):
inputs = []
embed_layers = []
for (c, (in_size, out_size)) in zip(cat_vars, embedding_sizes):
i = Input(shape=(1,))
o = Embedding(in_size, out_size, name=c)(i)
o = Reshape(target_shape=(out_size,))(o)
inputs.append(i)
embed_layers.append(o)
embed = Concatenate()(embed_layers)
embed = Dropout(0.04)(embed)
cont_input = Input(shape=(len(cont_vars),))
inputs.append(cont_input)
x = Concatenate()([embed, cont_input])
x = Dense(1000, kernel_initializer='he_normal')(x)
x = Activation('relu')(x)
x = Dropout(0.1)(x)
x = Dense(500, kernel_initializer='he_normal')(x)
x = Activation('relu')(x)
x = Dropout(0.1)(x)
x = Dense(1, kernel_initializer='he_normal')(x)
x = Activation('linear')(x)
model = Model(inputs=inputs, outputs=x)
opt = Adam(lr=0.001)
model.compile(loss='mean_absolute_error', optimizer=opt, metrics=[rmspe])
return model
```
One of the cool tricks Jeremy introduced in the class was the concept of a learning rate finder. The idea is to start with a very small learning rate and slowly increase it throughout the epoch, and monitor the loss along the way. It should end up as a curve that gives a good indication of where to set the learning rate for training. To accomplish this with Keras, I found a script on Github that implements learning rate cycling and includes a class that’s supposed to mimic Jeremy’s LR finder. We can just download a copy to the local directory.
```
!wget "https://raw.githubusercontent.com/titu1994/keras-one-cycle/master/clr.py"
```
Let’s set up and train the model for one epoch using the LRFinder class as a callback. It will slowly but exponentially increase the learning rate each batch and track the loss so we can plot the results.
```
from clr import LRFinder
lr_finder = LRFinder(num_samples=X.shape[0], batch_size=128, minimum_lr=1e-5, maximum_lr=10,
lr_scale='exp', loss_smoothing_beta=0.995, verbose=False)
model = EmbeddingNet(cat_vars, cont_vars, embedding_sizes)
history = model.fit(x=X_array, y=y, batch_size=128, epochs=1, verbose=1, callbacks=[lr_finder],
validation_data=(X_val_array, y_val), shuffle=False)
lr_finder.plot_schedule(clip_beginning=20)
```
It doesn’t look as good as the plot Jeremy used in the class. The PyTorch version seemed to make it much more apparent where the loss started to level off. I haven’t dug into this too closely but I’m guessing there are some "tricks" in that version that we aren't using. If I had to eyeball this I’d say it’s recommending 1e-4 for the learning rate, but Jeremy used 1e-3 so we’ll go with that instead.
We’re now ready to train the model. I’ve included two new callbacks (both built into Keras) to demonstrate how they work. The first one automatically reduces the learning rate as we progress through training if the validation error stops improving. The second one will save a copy of the model weights to a file every time we reach a new low in validation error.
```
model = EmbeddingNet(cat_vars, cont_vars, embedding_sizes)
lr_reducer = ReduceLROnPlateau(monitor='val_loss', factor=0.2, patience=3, verbose=1, mode='auto',
min_delta=10, cooldown=0, min_lr=0.0001)
checkpoint = ModelCheckpoint('best_model_weights.hdf5', monitor='val_loss', save_best_only=True)
history = model.fit(x=X_array, y=y, batch_size=128, epochs=20, verbose=1, callbacks=[lr_reducer, checkpoint],
validation_data=(X_val_array, y_val), shuffle=False)
```
By the end it’s doing pretty good, and it looks like the model is still improving. We can quickly get a snapshot of its performance using the “history” object that Keras’s "fit" method returns.
```
loss_history = history.history['loss']
val_loss_history = history.history['val_loss']
min_val_epoch = val_loss_history.index(min(val_loss_history)) + 1
print('min training loss = {0}'.format(min(loss_history)))
print('min val loss = {0}'.format(min(val_loss_history)))
print('min val epoch = {0}'.format(min_val_epoch))
```
I also like to make plots to visually see what’s going on. Let’s create a function that plots the training and validation loss history.
```
from jupyterthemes import jtplot
jtplot.style()
def plot_loss_history(history, n_epochs):
fig, ax = plt.subplots(figsize=(8, 8 * 3 / 4))
ax.plot(list(range(n_epochs)), history.history['loss'], label='Training Loss')
ax.plot(list(range(n_epochs)), history.history['val_loss'], label='Validation Loss')
ax.set_xlabel('Epoch')
ax.set_ylabel('Loss')
ax.legend(loc='upper right')
fig.tight_layout()
plot_loss_history(history, 20)
```
The validation loss was pretty unstable early on but was really starting to converge toward the end of training. We can do something similar for the learning rate history.
```
def plot_learning_rate(history):
fig, ax = plt.subplots(figsize=(8, 8 * 3 / 4))
ax.set_xlabel('Training Iterations')
ax.set_ylabel('Learning Rate')
ax.plot(history.history['lr'])
fig.tight_layout()
plot_learning_rate(history)
```
One other innovation Jeremy introduced in the class is the idea of using learning rate cycles to help prevent the model from settling in a bad local minimum. This is based on research by Leslie Smith that showed using this type of learning rate policy can lead to quicker convergence and better accuracy (this is also where the learning rate finder idea came from). Fortunately the file we downloaded earlier includes support for cyclical learning rates in Keras, so we can try this out ourselves. The policy Jeremy is currently recommending is called a “one-cycle” policy so that’s what we’ll try.
(As an aside, Jeremy [wrote a blog post](http://www.fast.ai/2018/04/30/dawnbench-fastai/) about this if you'd like to dig into its origins a bit more. His results applying it to ImageNet were quite impressive.)
```
from clr import OneCycleLR
model2 = EmbeddingNet(cat_vars, cont_vars, embedding_sizes)
batch_size = 128
n_epochs = 10
lr_manager = OneCycleLR(num_samples=X.shape[0] + batch_size, num_epochs=n_epochs, batch_size=batch_size, max_lr=0.01,
end_percentage=0.1, scale_percentage=None, maximum_momentum=None,
minimum_momentum=None, verbose=False)
history = model2.fit(x=X_array, y=y, batch_size=batch_size, epochs=n_epochs, verbose=1,
callbacks=[checkpoint, lr_manager], validation_data=(X_val_array, y_val), shuffle=False)
```
As you can probably tell from the model error, I didn’t have a lot of success with this strategy. I tried a few different configurations and nothing really worked, but I wouldn’t say it’s an indictment of the technique so much as it just didn’t happen to do well within the narrow scope that I attempted to apply it. Nevertheless, I’m definitely adding it to my toolbox for future reference.
If my earlier description wasn’t clear, this is how the learning is supposed to evolve over time. It forms a triangle from the starting point, coming back to the original learning rate towards the end and then decaying further as training wraps up.
```
plot_learning_rate(lr_manager)
```
One last trick worth discussing is what we can do with the embeddings that our network learned. Similar to word embeddings, these vectors contain potentially interesting information about how the values in each category relate to each other. One really simple way to see this visually is to do a PCA transform on the learned embedding weights and plot the first two dimensions. Let’s create a function to do just that.
```
def plot_embedding(model, encoders, category):
embedding_layer = model.get_layer(category)
weights = embedding_layer.get_weights()[0]
pca = PCA(n_components=2)
weights = pca.fit_transform(weights)
weights_t = weights.T
fig, ax = plt.subplots(figsize=(8, 8 * 3 / 4))
ax.scatter(weights_t[0], weights_t[1])
for i, day in enumerate(encoders[category].classes_):
ax.annotate(day, (weights_t[0, i], weights_t[1, i]))
fig.tight_layout()
```
We can now plot any categorical variable in the model and get a sense of which categories are more or less similar to each other. For instance, if we examine "day of week", it seems to have picked up that Sunday (7 on the chart) is quite different than every other day for store sales. And if we look at "state" (this data is for a German company BTW) there’s probably some regional similarity to the cluster in the bottom left. It’s a really cool technique that potentially has a wide range of uses.
```
plot_embedding(model, encoders, 'DayOfWeek')
plot_embedding(model, encoders, 'State')
```
| github_jupyter |
# ロジスティック回帰
## ロジスティック回帰(2値分類)
---
美味しいワイン(1), 美味しくないワイン(0)として、2つのクラスに分類するモデルを構築する
## 環境構築
```
# 分析に必要なライブラリをインポート
import pandas as pd
import numpy as np
# 可視化ライブラリのインポート
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style('whitegrid')
%matplotlib inline
```
## データセット準備
### 読み込み
```
# データセットを読み込む
df = pd.read_csv("../dataset/winequality-red.csv", sep=";")
df.head()
```
### 美味しいワイン(1)と美味しくないワイン(0)とする関数を定義する
```
def delicious(x):
# もしqualityが7以上であれば美味しい(1)
if x >= 7:
return 1
# 7以上でなければ、美味しくない(0)
else:
return 0
"""
applyを使って、美味しいか、そうでないかの2値分類結果を
列名「delicious_score」としてDataFrameに新たに追加する
"""
df['delicious_score'] = df['quality'].apply(delicious)
# 正しくDataFrameに列が追加されたのか確認
df.head()
```
- これでデータセットの準備ができた
- ここまでの作業は*中間テーブル作成*の作業にあたる
### 各列の平均値を比較する
美味しいワイン(1)と美味しくないワイン(0)における説明変数の平均値の違いを比較してみる
```
df.groupby("delicious_score").mean()
sns.countplot('citric acid',data=df.sort_values('citric acid'),hue='delicious_score')
```
### 特徴量と特徴量の相関性
```
plt.pcolor(df.corr(), cmap="Blues")
plt.xticks(range(0, len(df.columns)), df.columns, rotation=50)
plt.yticks(range(0, len(df.columns)), df.columns, rotation=0)
```
### 目的変数(Y)と説明変数(X)にそれぞれデータを格納する
```
# 目的変数
Y = df["delicious_score"]
# 説明変数
X = df.drop(["quality", "delicious_score"], axis=1)
```
### 学習データ(train)と検証データ(test)に分割する
交差検証(Cross Validation)を行います
#### Cross Validation(交差検証)とは
統計学において標本データを分割し、その一部をまず解析して、残る部分でその解析のテストを行い、解析自身の妥当性の検証・確認に当てる手法を指す。(Wikipediaより引用)
```
# 交差検証
from sklearn.cross_validation import train_test_split
# データ分割
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=0.4,random_state=3)
```
#### 学習データ量
```
print [d.shape for d in [X_train, X_test, Y_train, Y_test]]
```
### シグモイド関数
グラフで見るとわかるようにシグモイド関数σ(x)σ(x)は値を(0, 1)の範囲で取るため、この値を確率として解釈して使うことができる
```
def sigmoid(x):
return 1 / (1+np.exp(-x))
x_range = np.linspace(-10, 10, 501)
plt.figure(figsize=(9,5))
plt.ylim(-0.1, 1.1)
plt.xlim(-10, 10)
plt.plot([-10,10],[0,0], "k", lw=1)
plt.plot([0,0],[-1,1.5], "k", lw=1)
plt.plot(x_range, sigmoid(x_range), zorder=100)
```
## 機械学習
```
import pystan
from pystan import StanModel
# ロジスティック回帰
from sklearn.linear_model import LogisticRegression
# 性能の評価用
from sklearn import metrics
# インスタンス生成
log_model = LogisticRegression()
# 学習データを使って機械学習 = モデル作成
log_model.fit(X_train, Y_train)
```
### 学習したモデルの予測精度
#### 予測精度の算出方法
学習したモデルを使って予測(predict_Y)と実際の値(Y)を比較することによって算出できる
```
log_model.score(X_train, Y_train)
```
およそ88%の予測精度が実現できました。
```
Y.mean()
```
実際のYの平均値は0.13です。
ロジスティック回帰による予測モデルは、常に「美味しくない(0)」と出力すると、
全体1-0.13=0.87となり、87%の予測精度が得られます。
この値より、88.2%は若干高いことがわかります。
### 学習したモデルの係数(coefficient)を表示する
どのような変数が予測に影響を与えている(寄与度)のか調べてみましょう
```
from pandas import DataFrame
# 変数名とその変数を格納するDataFrameを作成
coeff_df = DataFrame([X.columns, log_model.coef_[0]]).T
coeff_df
```
#### オッズ比について後で調べる
# 検証
```
# 検証データで予測
class_predict = log_model.predict(X_test)
# 予測精度確認用
from sklearn import metrics
# 精度確認
metrics.accuracy_score(Y_test, class_predict)
```
### モデル構築のステップ
1. 標準化
2. テストデータと訓練データに分割
3. 外れ値の除去
### モデル概要
---
- 今回は教師あり学習
-
```
# 多重共線性があるか分析
#statsmodelsのvifをインポート
from statsmodels.stats.outliers_influence import variance_inflation_factor
```
# ↑この続きはあとでやる
最適な重回帰モデルをステップワイズ法でモデル選択(AICで評価)
# ロジスティック回帰モデル
```
from sklearn import datasets
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import StratifiedKFold
from sklearn.model_selection import cross_validate
# 機械学習用です。
from sklearn.linear_model import LogisticRegression
from sklearn.cross_validation import train_test_split
# もう一つ、性能の評価用に
from sklearn import metrics
# 教師データの作成
y = df["quality"]
x = df.drop("quality", axis=1)
# データ分割
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size = 0.2)
# 機械学習
model = LogisticRegression()
model.fit(x_train, y_train)
# 評価用ライブラリ
from sklearn.metrics import accuracy_score
from sklearn.metrics import classification_report
# 評価
y_pred = model.predict(x_test)
print(classification_report(y_test, y_pred))
print("正解率 = ", accuracy_score(y_test, y_pred))
# pickleのインポート
import pickle
#pickleでモデルを保存する
# モデルを保存する
filename = 'nullsuckAi_model.sav'
pickle.dump(model, open(filename, 'wb'))
```
| github_jupyter |
# Austrian energy system Tutorial Part 1: Building an Energy Model
For information on how to install *MESSAGEix*, please refer to [Installation page](https://docs.messageix.org/en/stable/#getting-started) and for getting *MESSAGEix* tutorials, please follow the steps mentioned in [Tutorials](https://docs.messageix.org/en/stable/tutorials.html).
Please refer to the [user guidelines](https://github.com/iiasa/message_ix/blob/master/NOTICE.rst)
for additional information on using *MESSAGEix*, including the recommended citation and how to name new models.
**Pre-requisites**
- You have the *MESSAGEix* framework installed and working
**Structure of these tutorials.** After having run this baseline tutorial, you are able to start with any of the other tutorials, but we recommend to follow the order below for going through the information step-wise:
1. Prepare the base model version (Python: ``austria.ipynb``, also available in R: ``austria_reticulate.ipynb``)
2. Plot the results of the baseline runs (Python: ``austria_load_scenario.ipynb``, also available in R: ``austria_load_scenario_R.ipynb``).
3. Run a single policy scenario (``austria_single_policy.ipynb``).
4. Run multiple policy scenarios. This tutorial has two notebooks: an introduction with some exercises and completed code for the exercises (exercises: ``austria_multiple_policies.ipynb``, answers: ``austria_multiple_policies-answers.ipynb``).
**Introduction**
In this notebook, we will build a model of the Austrian energy system from scratch. The process will involve defining our model's time horizon and spatial extent, and then populating the model with data associated with model parameters. Once we have a baseline model, we will then move on to investigating policy scenarios.
We will be populating different kinds of parameters including:
### Economic Parameters
- `interestrate`
- `demand`
### Technology Parameters
#### Engineering Parameters
- `input`
- `output`
- `technical_lifetime`
- `capacity_factor`
#### Technoeconomic Parameters
- `inv_cost`
- `fix_cost`
- `var_cost`
### Dynamic Behavior Parameters
- `bound_activity_up`
- `bound_activity_lo`
- `bound_new_capacity_up`
- `initial_activity_up`
- `growth_activity_up`
### Emissions
- `emission_factor`
A full list of parameters can be found in the [MESSAGEix documentation](http://messageix.iiasa.ac.at/model/MESSAGE/parameter_def.html). (If you have cloned the MESSAGEix [Github repository](https://github.com/iiasa/message_ix), the documentation can also be built offline; see `doc/README.md`.)
## The Final Product
At the completion of this exercise, we will have developed an energy model is comprised of the below Reference Energy System (RES):

## Setup
```
# load required packages
import itertools
import pandas as pd
import matplotlib.pyplot as plt
%matplotlib inline
plt.style.use('ggplot')
import ixmp as ix
import message_ix
from message_ix.util import make_df
# launch the IX modeling platform using the local default database
mp = ix.Platform()
model = "Austrian energy model"
scen = "baseline"
annot = "developing a stylized energy system model for illustration and testing"
scenario = message_ix.Scenario(mp, model, scen, version='new', annotation=annot)
```
## Time and Spatial Detail
The model includes the time periods 2010, 2020, 2030 and 2040.
```
horizon = range(2010, 2041, 10)
scenario.add_horizon(year=horizon)
country = 'Austria'
scenario.add_spatial_sets({'country': country})
```
## Model Structure
```
scenario.add_set("commodity", ["electricity", "light", "other_electricity"])
scenario.add_set("level", ["secondary", "final", "useful"])
scenario.add_set("mode", "standard")
```
## Economic Parameters
Definition of the socio-economic discount rate:
```
scenario.add_par("interestrate", horizon, value=0.05, unit='-')
```
The fundamental premise of the model is to satisfy demand for energy (services). To first order, demands for services (e.g. electricity) track with economic productivity (GDP). Therefore, as a simple example, we define both a GDP profile and a correlation factor between GDP growth and demand, called beta. Beta will then be used to obtain a simplistic demand profile.
```
gdp = pd.Series([1., 1.21631, 1.4108, 1.63746], index=horizon)
beta = 0.7
demand = gdp ** beta
```
## Technologies
```
plants = [
"coal_ppl",
"gas_ppl",
"oil_ppl",
"bio_ppl",
"hydro_ppl",
"wind_ppl",
"solar_pv_ppl", # actually primary -> final
]
secondary_energy_techs = plants + ['import']
final_energy_techs = ['electricity_grid']
lights = [
"bulb",
"cfl",
]
useful_energy_techs = lights + ['appliances']
technologies = secondary_energy_techs + final_energy_techs + useful_energy_techs
scenario.add_set("technology", technologies)
demand_per_year = 55209. / 8760 # from IEA statistics
elec_demand = pd.DataFrame({
'node': country,
'commodity': 'other_electricity',
'level': 'useful',
'year': horizon,
'time': 'year',
'value': demand_per_year * demand,
'unit': 'GWa',
})
scenario.add_par("demand", elec_demand)
demand_per_year = 6134. / 8760 # from IEA statistics
light_demand = pd.DataFrame({
'node': country,
'commodity': 'light',
'level': 'useful',
'year': horizon,
'time': 'year',
'value': demand_per_year * demand,
'unit': 'GWa',
})
scenario.add_par("demand", light_demand)
```
### Engineering Parameters
```
year_df = scenario.vintage_and_active_years()
vintage_years, act_years = year_df['year_vtg'], year_df['year_act']
base_input = {
'node_loc': country,
'year_vtg': vintage_years,
'year_act': act_years,
'mode': 'standard',
'node_origin': country,
'commodity': 'electricity',
'time': 'year',
'time_origin': 'year',
}
grid = pd.DataFrame(dict(
technology = 'electricity_grid',
level = 'secondary',
value = 1.0,
unit = '-',
**base_input
))
scenario.add_par("input", grid)
bulb = pd.DataFrame(dict(
technology = 'bulb',
level = 'final',
value = 1.0,
unit = '-',
**base_input
))
scenario.add_par("input", bulb)
cfl = pd.DataFrame(dict(
technology = 'cfl',
level = 'final',
value = 0.3, # LED and CFL lighting equipment are more efficient than conventional light bulbs,
#so they need less input electricity to produce the same quantity of 'light'
#compared to conventional light bulbs (0.3 units vs 1.0, respectively)
unit = '-',
**base_input
))
scenario.add_par("input", cfl)
app = pd.DataFrame(dict(
technology = 'appliances',
level = 'final',
value = 1.0,
unit = '-',
**base_input
))
scenario.add_par("input", app)
make_df?
# to see what the funcion "df" does.
base_output = {
'node_loc': country,
'year_vtg': vintage_years,
'year_act': act_years,
'mode': 'standard',
'node_dest': country,
'time': 'year',
'time_dest': 'year',
'unit': '-',
}
imports = make_df(base_output, technology='import', commodity='electricity',
level='secondary', value=1.)
scenario.add_par('output', imports)
grid = make_df(base_output, technology='electricity_grid', commodity='electricity',
level='final', value=0.873)
scenario.add_par('output', grid)
bulb = make_df(base_output, technology='bulb', commodity='light',
level='useful', value=1.)
scenario.add_par('output', bulb)
cfl = make_df(base_output, technology='cfl', commodity='light',
level='useful', value=1.)
scenario.add_par('output', cfl)
app = make_df(base_output, technology='appliances', commodity='other_electricity',
level='useful', value=1.)
scenario.add_par('output', app)
coal = make_df(base_output, technology='coal_ppl', commodity='electricity',
level='secondary', value=1.)
scenario.add_par('output', coal)
gas = make_df(base_output, technology='gas_ppl', commodity='electricity',
level='secondary', value=1.)
scenario.add_par('output', gas)
oil = make_df(base_output, technology='oil_ppl', commodity='electricity',
level='secondary', value=1.)
scenario.add_par('output', oil)
bio = make_df(base_output, technology='bio_ppl', commodity='electricity',
level='secondary', value=1.)
scenario.add_par('output', bio)
hydro = make_df(base_output, technology='hydro_ppl', commodity='electricity',
level='secondary', value=1.)
scenario.add_par('output', hydro)
wind = make_df(base_output, technology='wind_ppl', commodity='electricity',
level='secondary', value=1.)
scenario.add_par('output', wind)
solar_pv = make_df(base_output, technology='solar_pv_ppl', commodity='electricity',
level='final', value=1.)
scenario.add_par('output', solar_pv)
base_technical_lifetime = {
'node_loc': country,
'year_vtg': horizon,
'unit': 'y',
}
lifetimes = {
'coal_ppl': 40,
'gas_ppl': 30,
'oil_ppl': 30,
'bio_ppl': 30,
'hydro_ppl': 60,
'wind_ppl': 20,
'solar_pv_ppl': 20,
'bulb': 1,
'cfl': 10,
}
for tec, val in lifetimes.items():
df = make_df(base_technical_lifetime, technology=tec, value=val)
scenario.add_par('technical_lifetime', df)
base_capacity_factor = {
'node_loc': country,
'year_vtg': vintage_years,
'year_act': act_years,
'time': 'year',
'unit': '-',
}
capacity_factor = {
'coal_ppl': 0.85,
'gas_ppl': 0.75,
'oil_ppl': 0.75,
'bio_ppl': 0.75,
'hydro_ppl': 0.5,
'wind_ppl': 0.2,
'solar_pv_ppl': 0.15,
'bulb': 0.1,
'cfl': 0.1,
}
for tec, val in capacity_factor.items():
df = make_df(base_capacity_factor, technology=tec, value=val)
scenario.add_par('capacity_factor', df)
```
### Technoeconomic Parameters
```
base_inv_cost = {
'node_loc': country,
'year_vtg': horizon,
'unit': 'USD/kW',
}
# Adding a new unit to the library
mp.add_unit('USD/kW')
# in $ / kW (specific investment cost)
costs = {
'coal_ppl': 1500,
'gas_ppl': 870,
'oil_ppl': 950,
'hydro_ppl': 3000,
'bio_ppl': 1600,
'wind_ppl': 1100,
'solar_pv_ppl': 4000,
'bulb': 5,
'cfl': 900,
}
for tec, val in costs.items():
df = make_df(base_inv_cost, technology=tec, value=val)
scenario.add_par('inv_cost', df)
base_fix_cost = {
'node_loc': country,
'year_vtg': vintage_years,
'year_act': act_years,
'unit': 'USD/kWa',
}
# Adding a new unit to the library
mp.add_unit('USD/kWa')
# in $ / kW / year (every year a fixed quantity is destinated to cover part of the O&M costs
# based on the size of the plant, e.g. lightning, labor, scheduled maintenance, etc.)
costs = {
'coal_ppl': 40,
'gas_ppl': 25,
'oil_ppl': 25,
'hydro_ppl': 60,
'bio_ppl': 30,
'wind_ppl': 40,
'solar_pv_ppl': 25,
}
for tec, val in costs.items():
df = make_df(base_fix_cost, technology=tec, value=val)
scenario.add_par('fix_cost', df)
base_var_cost = {
'node_loc': country,
'year_vtg': vintage_years,
'year_act': act_years,
'mode': 'standard',
'time': 'year',
'unit': 'USD/kWa',
}
# Variable O&M (costs associatied to the degradation of equipment when the plant is functioning
# per unit of energy produced)
# kWa = kW·year = 8760 kWh. Therefore this costs represents USD per 8760 kWh of energy.
# Do not confuse with fixed O&M units.
#var O&M in $ / MWh
costs = {
'coal_ppl': 24.4,
'gas_ppl': 42.4,
'oil_ppl': 77.8,
'bio_ppl': 48.2,
'electricity_grid': 47.8,
}
for tec, val in costs.items():
df = make_df(base_var_cost, technology=tec, value=val * 8760. / 1e3) # to convert it into USD/kWa
scenario.add_par('var_cost', df)
```
## Dynamic Behavior Parameters
In this section the following parameters will be added to the different technologies:
- `bound_activity_up`
- `bound_activity_lo`
- `bound_new_capacity_up`
- `initial_activity_up`
- `growth_activity_up`
As stated in the **Introduction**, a full list of parameters can be found in the *MESSAGEix* documentation. Specifically for this list, please refer to the section [Bounds on capacity and activity](https://docs.messageix.org/en/stable/model/MESSAGE/parameter_def.html#bounds-on-capacity-and-activity)
```
base_growth = {
'node_loc': country,
'year_act': horizon[1:],
'value': 0.05,
'time': 'year',
'unit': '%',
}
growth_technologies = [
"coal_ppl",
"gas_ppl",
"oil_ppl",
"bio_ppl",
"hydro_ppl",
"wind_ppl",
"solar_pv_ppl",
"cfl",
"bulb",
]
for tec in growth_technologies:
df = make_df(base_growth, technology=tec)
scenario.add_par('growth_activity_up', df)
base_initial = {
'node_loc': country,
'year_act': horizon[1:],
'time': 'year',
'unit': '%',
}
for tec in lights:
df = make_df(base_initial, technology=tec, value=0.01 * light_demand['value'].loc[horizon[1:]])
scenario.add_par('initial_activity_up', df)
base_activity = {
'node_loc': country,
'year_act': [2010],
'mode': 'standard',
'time': 'year',
'unit': 'GWa',
}
# in GWh - from IEA Electricity Output
activity = {
'coal_ppl': 7184,
'gas_ppl': 14346,
'oil_ppl': 1275,
'hydro_ppl': 38406,
'bio_ppl': 4554,
'wind_ppl': 2064,
'solar_pv_ppl': 89,
'import': 2340,
'cfl': 0,
}
#MODEL CALIBRATION: by inserting an upper and lower bound to the same quantity we are ensuring
#that the model is calibrated at that value that year, so we are at the right starting point.
for tec, val in activity.items():
df = make_df(base_activity, technology=tec, value=val / 8760.)
scenario.add_par('bound_activity_up', df)
scenario.add_par('bound_activity_lo', df)
base_capacity = {
'node_loc': country,
'year_vtg': [2010],
'unit': 'GW',
}
cf = pd.Series(capacity_factor)
act = pd.Series(activity)
capacity = (act / 8760 / cf).dropna().to_dict()
for tec, val in capacity.items():
df = make_df(base_capacity, technology=tec, value=val)
scenario.add_par('bound_new_capacity_up', df)
base_activity = {
'node_loc': country,
'year_act': horizon[1:],
'mode': 'standard',
'time': 'year',
'unit': 'GWa',
}
# in GWh - base value from IEA Electricity Output
keep_activity = {
'hydro_ppl': 38406,
'bio_ppl': 4554,
'import': 2340,
}
for tec, val in keep_activity.items():
df = make_df(base_activity, technology=tec, value=val / 8760.)
scenario.add_par('bound_activity_up', df)
```
## Emissions
```
scenario.add_set('emission', 'CO2')
scenario.add_cat('emission', 'GHGs', 'CO2')
base_emissions = {
'node_loc': country,
'year_vtg': vintage_years,
'year_act': act_years,
'mode': 'standard',
'unit': 'tCO2/kWa',
}
# adding new units to the model library (needed only once)
mp.add_unit('tCO2/kWa')
mp.add_unit('MtCO2')
emissions = {
'coal_ppl': ('CO2', 0.854), # units: tCO2/MWh
'gas_ppl': ('CO2', 0.339), # units: tCO2/MWh
'oil_ppl': ('CO2', 0.57), # units: tCO2/MWh
}
for tec, (species, val) in emissions.items():
df = make_df(base_emissions, technology=tec, emission=species, value=val * 8760. / 1000) #to convert tCO2/MWh into tCO2/kWa
scenario.add_par('emission_factor', df)
```
## Commit the datastructure and solve the model
```
comment = 'initial commit for Austria model'
scenario.commit(comment)
scenario.set_as_default()
scenario.solve()
scenario.var('OBJ')['lvl']
```
# Plotting Results
```
from message_ix.reporting import Reporter
from message_ix.util.tutorial import prepare_plots
rep = Reporter.from_scenario(scenario)
prepare_plots(rep)
rep.set_filters(t=plants)
rep.get("plot new capacity")
rep.set_filters(t=lights)
rep.get("plot new capacity")
rep.set_filters(t=plants)
rep.get("plot capacity")
rep.set_filters(t=lights)
rep.get("plot capacity")
rep.get("plot demand")
rep.set_filters(t=plants)
rep.get("plot activity")
rep.set_filters(t=lights)
rep.get("plot activity")
rep.set_filters(c=["light", "other_electricity"])
rep.get("plot prices")
mp.close_db()
```
| github_jupyter |
# PyTorch Tensor Notebook
### The Data Spartan
#### Helpful Resourses:
1. https://pytorch.org/tutorials/beginner/blitz/tensor_tutorial.html#sphx-glr-beginner-blitz-tensor-tutorial-py
2. https://pytorch.org/docs/stable/tensors.html
Tensors are similar to NumPy’s ndarrays, with the addition being that Tensors can also be used on a GPU to accelerate
computing.
```
from __future__ import print_function
import torch
```
# Tensor Initialization
#### An Uninitialized 5 X 4 Tensor
```
x = torch.empty(5, 4)
print(x)
```
#### A Randomly Initialized 5 x 4 Tensor
```
x = torch.rand(5, 4)
print(x)
```
#### A 5 x 4 Tensor filled zeros and of dtype long
```
x = torch.zeros(5, 4, dtype=torch.long)
print(x)
```
#### A 1 x 3 Tensor with Data Fed directly into it
```
x = torch.tensor([5, 3, 4])
print(x)
```
#### A 2 x 3 Tensor with Data Directly Fed into it
```
x = torch.tensor([[6, 5, 4], [3, 2, 1]])
print(x)
```
#### A 5 x 4 Tensor with ones using new_one methods and passing the dimension and type as parameters
```
x = x.new_ones(5, 4, dtype=torch.double)
print(x)
```
#### A 1 x 4 Tensor with ones using new_one methods and passing the dimension and type as parameters
```
x = x.new_ones(1, 4, dtype=torch.double)
print(x)
```
#### A 5 x 4 Tensor with zeros using new_zero methods and passing the dimension and type as parameters
```
x = x.new_zeros(5, 4, dtype=torch.double)
print(x)
```
#### A 1 x 4 Tensor with zeros using new_zero methods and passing the dimension and type as parameters
```
x = x.new_zeros(1, 4, dtype=torch.double)
print(x)
```
#### Setting Dtype to Float
```
x = torch.randn_like(x, dtype=torch.float)
print(x)
```
#### Printing the Size of a Tensor
```
print(x.size())
```
# Operations with Tensors
#### Transpose
```
x.t()
# Transpose (via permute)
x.permute(-1,0)
```
#### Slicing
```
x = torch.Tensor([[1, 2, 3], [4, 5, 6], [7, 8, 9]])
print(x)
# Print the last column
print(x[:, -1])
# First 2 rows
print(x[:2, :])
# Lower right corner
print(x[-1:, -1:])
```
#### Addition
```
x = torch.tensor([[1, 2, 3], [4, 5, 6], [7, 8, 9]])
y = torch.tensor([[2, 2, 2], [3, 3, 3], [4, 4, 4]])
z = x + y
print(z)
```
#### Subtraction
```
x = torch.tensor([[1, 2, 3], [4, 5, 6], [7, 8, 9]])
y = torch.tensor([[2, 2, 2], [3, 3, 3], [4, 4, 4]])
z = x - y
print(z)
```
#### Multiplication
```
x = torch.tensor([[1, 2, 3], [4, 5, 6], [7, 8, 9]])
y = torch.tensor([[2, 2, 2], [3, 3, 3], [4, 4, 4]])
z = x * y
print(x)
```
#### Scalar Addition
```
x = torch.tensor([[1, 2, 3], [4, 5, 6], [7, 8, 9]])
z = x + 1
print(z)
```
#### Scalar Subtraction
```
x = torch.tensor([[1, 2, 3], [4, 5, 6], [7, 8, 9]])
z = x - 1
print(z)
```
#### Scalar Multiplication
```
x = torch.tensor([[1, 2, 3], [4, 5, 6], [7, 8, 9]])
z = x * 2
print(x)
```
#### Scalar Divion
```
x = torch.tensor([[1, 2, 3], [4, 5, 6], [7, 8, 9]])
z = x / 2
print(z)
```
#### Alternate Method for Addition
```
print(torch.add(x, y))
```
#### Adding In place i.e the tensor itself is modified
#### Adding x to y
Note : Any operation that mutates a tensor in-place is post-fixed with an _. For example: x.copy_(y), x.t_(), will change x.
```
y.add_(x)
print(y)
```
#### NumPy Array-like indexing
```
print(x[:, 1])
```
#### Resizing
If you want to resize/reshape tensor, you can use torch.view:
```
x = torch.randn(4, 4)
y = x.view(16)
z = x.view(-1, 8) # the size -1 is inferred from other dimensions
print(x.size(), y.size(), z.size())
```
#### Accessing the Element-wise Value of a Tensor
If you have one element tensor, use .item() to get the value as a Python number. For multiple elements, use a loop.
```
x = torch.randn(1)
print(x)
print(x.item())
x = torch.rand(5)
print(x)
for item in x:
print(item.item())
```
#### Converting a Torch Tensor to a NumPy Array
```
a = torch.ones(5)
print(a)
b = a.numpy()
print(b)
# Notice how modifying the Tensor a will affect the Numpy Array b
a.add_(1)
print(a)
print(b)
```
#### Converting NumPy Array to Torch Tensor
```
import numpy as np
a = np.ones(5)
b = torch.from_numpy(a)
print(a)
print(b)
np.add(a, 1, out=a)
print(a)
print(b)
a = np.random.rand(4,3)
a
b = torch.from_numpy(a)
b
b.numpy()
# Multiply PyTorch Tensor by 5, in place
b.mul_(5)
# Numpy array matches new values from Tensor
a
```
| github_jupyter |
<a href="https://colab.research.google.com/github/ekramasif/Basic-Machine-Learning/blob/main/Extraa/PythonCheatSheet.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Python Cheatsheet
## Contents
1. <a href='#section1'>Syntax and whitespace</a>
2. <a href='#section2'>Comments</a>
3. <a href='#section3'>Numbers and operations</a>
4. <a href='#section4'>String manipulation</a>
5. <a href='#section5'>Lists, tuples, and dictionaries</a>
6. <a href='#section6'>JSON</a>
7. <a href='#section7'>Loops</a>
8. <a href='#section8'>File handling</a>
9. <a href='#section9'>Functions</a>
10. <a href='#section10'>Working with datetime</a>
11. <a href='#section11'>NumPy</a>
12. <a href='#section12'>Pandas</a>
To run a cell, press **Shift+Enter** or click **Run** at the top of the page.
<a id="section_1"></a>
## 1. Syntax and whitespace
Python uses indented space to indicate the level of statements. The following cell is an example where '**if**' and '**else**' are in same level, while '**print**' is separated by space to a different level. Spacing should be the same for items that are on the same level.
```
student_number = input("Enter your student number:")
if int(student_number) != 0:
print("Welcome student {}".format(student_number))
else:
print("Try again!")
```
<a id='section2'></a>
## 2. Comments
In Python, comments start with hash '#' and extend to the end of the line. '#' can be at the begining of the line or after code.
```
# This is code to print hello world!
print("Hello world!") # Print statement for hello world
print("# is not a comment in this case")
```
<a id='section3'></a>
## 3. Numbers and operations
Like with other programming languages, there are four types of numbers:
- Integers (e.g., 1, 20, 45, 1000) indicated by *int*
- Floating point numbers (e.g., 1.25, 20.35, 1000.00) indicated by *float*
- Long integers
- Complex numbers (e.g., x+2y where x is known)
Operation | Result
----------------|-------------------------------------
x + y | Sum of x and y
x - y | Difference of x and y
x * y | Product of x and y
x / y | Quotient of x and y
x // y | Quotient of x and y (floored)
x % y | Remainder of x / y
abs(x) | Absolute value of x
int(x) | x converted to integer
long(x) | x converted to long integer
float(x) | x converted to floating point
pow(x, y) | x to the power y
x ** y | x to the power y
```
# Number examples
a = 5 + 8
print("Sum of int numbers: {} and number format is {}".format(a, type(a)))
b = 5 + 2.3
print ("Sum of int and {} and number format is {}".format(b, type(b)))
```
<a id='section4'></a>
## 4. String manipulation
Python has rich features like other programming languages for string manipulation.
```
# Store strings in a variable
test_word = "hello world to everyone"
# Print the test_word value
print(test_word)
# Use [] to access the character of the string. The first character is indicated by '0'.
print(test_word[0])
# Use the len() function to find the length of the string
print(len(test_word))
# Some examples of finding in strings
print(test_word.count('l')) # Count number of times l repeats in the string
print(test_word.find("o")) # Find letter 'o' in the string. Returns the position of first match.
print(test_word.count(' ')) # Count number of spaces in the string
print(test_word.upper()) # Change the string to uppercase
print(test_word.lower()) # Change the string to lowercase
print(test_word.replace("everyone","you")) # Replace word "everyone" with "you"
print(test_word.title()) # Change string to title format
print(test_word + "!!!") # Concatenate strings
print(":".join(test_word)) # Add ":" between each character
print("".join(reversed(test_word))) # Reverse the string
```
<a id='section5'></a>
## 5. Lists, tuples, and dictionaries
Python supports data types lists, tuples, dictionaries, and arrays.
### Lists
A list is created by placing all the items (elements) inside square brackets \[ ] separated by commas. A list can have any number of items, and they may be of different types (integer, float, strings, etc.).
```
# A Python list is similar to an array. You can create an empty list too.
my_list = []
first_list = [3, 5, 7, 10]
second_list = [1, 'python', 3]
# Nest multiple lists
nested_list = [first_list, second_list]
nested_list
# Combine multiple lists
combined_list = first_list + second_list
combined_list
# You can slice a list, just like strings
combined_list[0:3]
# Append a new entry to the list
combined_list.append(600)
combined_list
# Remove the last entry from the list
combined_list.pop()
# Iterate the list
for item in combined_list:
print(item)
```
### Tuples
A tuple is similar to a list, but you use them with parentheses ( ) instead of square brackets. The main difference is that a tuple is immutable, while a list is mutable.
```
my_tuple = (1, 2, 3, 4, 5)
my_tuple[1:4]
```
### Dictionaries
A dictionary is also known as an associative array. A dictionary consists of a collection of key-value pairs. Each key-value pair maps the key to its associated value.
```
desk_location = {'jack': 123, 'joe': 234, 'hary': 543}
desk_location['jack']
```
<a id='section6'></a>
## 6. JSON
JSON is text writen in JavaScript Object Notation. Python has a built-in package called `json` that can be used to work with JSON data.
```
import json
# Sample JSON data
x = '{"first_name":"Jane", "last_name":"Doe", "age":25, "city":"Chicago"}'
# Read JSON data
y = json.loads(x)
# Print the output, which is similar to a dictonary
print("Employee name is "+ y["first_name"] + " " + y["last_name"])
```
<a id='section7'></a>
## 7. Loops
**If, Else, ElIf loop**: Python supports conditional statements like any other programming language. Python relies on indentation (whitespace at the begining of the line) to define the scope of the code.
```
a = 22
b = 33
c = 100
# if ... else example
if a > b:
print("a is greater than b")
else:
print("b is greater than a")
# if .. else .. elif example
if a > b:
print("a is greater than b")
elif b > c:
print("b is greater than c")
else:
print("b is greater than a and c is greater than b")
```
**While loop:** Runs a set of statements as long as the condition is true
```
# Sample while example
i = 1
while i < 10:
print("count is " + str(i))
i += 1
print("="*10)
# Continue to next iteration if x is 2. Finally, print message once the condition is false.
x = 0
while x < 5:
x += 1
if x == 2:
continue
print(x)
else:
print("x is no longer less than 5")
```
**For loop:** A `For` loop is more like an iterator in Python. A `For` loop is used for iterating over a sequence (list, tuple, dictionay, set, string, or range).
```
# Sample for loop examples
fruits = ["orange", "banana", "apple", "grape", "cherry"]
for fruit in fruits:
print(fruit)
print("\n")
print("="*10)
print("\n")
# Iterating range
for x in range(1, 10, 2):
print(x)
else:
print("task complete")
print("\n")
print("="*10)
print("\n")
# Iterating multiple lists
traffic_lights = ["red", "yellow", "green"]
action = ["stop", "slow down", "go"]
for light in traffic_lights:
for task in action:
print(light, task)
```
<a id='section8'></a>
## 8. File handling
The key function for working with files in Python is the `open()` function. The `open()` function takes two parameters: filename and mode.
There are four different methods (modes) for opening a file:
- "r" - Read
- "a" - Append
- "w" - Write
- "x" - Create
In addition, you can specify if the file should be handled in binary or text mode.
- "t" - Text
- "b" - Binary
```
# Let's create a test text file
!echo "This is a test file with text in it. This is the first line." > test.txt
!echo "This is the second line." >> test.txt
!echo "This is the third line." >> test.txt
# Read file
file = open('test.txt', 'r')
print(file.read())
file.close()
print("\n")
print("="*10)
print("\n")
# Read first 10 characters of the file
file = open('test.txt', 'r')
print(file.read(10))
file.close()
print("\n")
print("="*10)
print("\n")
# Read line from the file
file = open('test.txt', 'r')
print(file.readline())
file.close()
# Create new file
file = open('test2.txt', 'w')
file.write("This is content in the new test2 file.")
file.close()
# Read the content of the new file
file = open('test2.txt', 'r')
print(file.read())
file.close()
# Update file
file = open('test2.txt', 'a')
file.write("\nThis is additional content in the new file.")
file.close()
# Read the content of the new file
file = open('test2.txt', 'r')
print(file.read())
file.close()
# Delete file
import os
file_names = ["test.txt", "test2.txt"]
for item in file_names:
if os.path.exists(item):
os.remove(item)
print(f"File {item} removed successfully!")
else:
print(f"{item} file does not exist.")
```
<a id='section9'></a>
## 9. Functions
A function is a block of code that runs when it is called. You can pass data, or *parameters*, into the function. In Python, a function is defined by `def`.
```
# Defining a function
def new_funct():
print("A simple function")
# Calling the function
new_funct()
# Sample fuction with parameters
def param_funct(first_name):
print(f"Employee name is {first_name}.")
param_funct("Harry")
param_funct("Larry")
param_funct("Shally")
```
**Anonymous functions (lambda):** A lambda is a small anonymous function. A lambda function can take any number of arguments but only one expression.
```
# Sample lambda example
x = lambda y: y + 100
print(x(15))
print("\n")
print("="*10)
print("\n")
x = lambda a, b: a*b/100
print(x(2,4))
```
<a id='section10'></a>
## 10. Working with datetime
A `datetime` module in Python can be used to work with date objects.
```
import datetime
x = datetime.datetime.now()
print(x)
print(x.year)
print(x.strftime("%A"))
print(x.strftime("%B"))
print(x.strftime("%d"))
print(x.strftime("%H:%M:%S %p"))
```
<a id='section11'></a>
## 11. NumPy
NumPy is the fundamental package for scientific computing with Python. Among other things, it contains:
- Powerful N-dimensional array object
- Sophisticated (broadcasting) functions
- Tools for integrating C/C++ and Fortran code
- Useful linear algebra, Fourier transform, and random number capabilities
```
# Install NumPy using pip
!pip install --upgrade pip
!pip install numpy
# Import NumPy module
import numpy as np
```
### Inspecting your array
```
# Create array
a = np.arange(15).reshape(3, 5) # Create array with range 0-14 in 3 by 5 dimension
b = np.zeros((3,5)) # Create array with zeroes
c = np.ones( (2,3,4), dtype=np.int16 ) # Createarray with ones and defining data types
d = np.ones((3,5))
a.shape # Array dimension
len(b)# Length of array
c.ndim # Number of array dimensions
a.size # Number of array elements
b.dtype # Data type of array elements
c.dtype.name # Name of data type
c.astype(float) # Convert an array type to a different type
```
### Basic math operations
```
# Create array
a = np.arange(15).reshape(3, 5) # Create array with range 0-14 in 3 by 5 dimension
b = np.zeros((3,5)) # Create array with zeroes
c = np.ones( (2,3,4), dtype=np.int16 ) # Createarray with ones and defining data types
d = np.ones((3,5))
np.add(a,b) # Addition
np.subtract(a,b) # Substraction
np.divide(a,d) # Division
np.multiply(a,d) # Multiplication
np.array_equal(a,b) # Comparison - arraywise
```
### Aggregate functions
```
# Create array
a = np.arange(15).reshape(3, 5) # Create array with range 0-14 in 3 by 5 dimension
b = np.zeros((3,5)) # Create array with zeroes
c = np.ones( (2,3,4), dtype=np.int16 ) # Createarray with ones and defining data types
d = np.ones((3,5))
a.sum() # Array-wise sum
a.min() # Array-wise min value
a.mean() # Array-wise mean
a.max(axis=0) # Max value of array row
np.std(a) # Standard deviation
```
### Subsetting, slicing, and indexing
```
# Create array
a = np.arange(15).reshape(3, 5) # Create array with range 0-14 in 3 by 5 dimension
b = np.zeros((3,5)) # Create array with zeroes
c = np.ones( (2,3,4), dtype=np.int16 ) # Createarray with ones and defining data types
d = np.ones((3,5))
a[1,2] # Select element of row 1 and column 2
a[0:2] # Select items on index 0 and 1
a[:1] # Select all items at row 0
a[-1:] # Select all items from last row
a[a<2] # Select elements from 'a' that are less than 2
```
### Array manipulation
```
# Create array
a = np.arange(15).reshape(3, 5) # Create array with range 0-14 in 3 by 5 dimension
b = np.zeros((3,5)) # Create array with zeroes
c = np.ones( (2,3,4), dtype=np.int16 ) # Createarray with ones and defining data types
d = np.ones((3,5))
np.transpose(a) # Transpose array 'a'
a.ravel() # Flatten the array
a.reshape(5,-2) # Reshape but don't change the data
np.append(a,b) # Append items to the array
np.concatenate((a,d), axis=0) # Concatenate arrays
np.vsplit(a,3) # Split array vertically at 3rd index
np.hsplit(a,5) # Split array horizontally at 5th index
```
<a id='section12'></a>
## Pandas
Pandas is an open source, BSD-licensed library providing high-performance, easy-to-use data structures and data analysis tools for the Python programming language.
Pandas DataFrames are the most widely used in-memory representation of complex data collections within Python.
```
# Install pandas, xlrd, and openpyxl using pip
!pip install pandas
!pip install xlrd openpyxl
# Import NumPy and Pandas modules
import numpy as np
import pandas as pd
# Sample dataframe df
df = pd.DataFrame({'num_legs': [2, 4, np.nan, 0],
'num_wings': [2, 0, 0, 0],
'num_specimen_seen': [10, np.nan, 1, 8]},
index=['falcon', 'dog', 'spider', 'fish'])
df # Display dataframe df
# Another sample dataframe df1 - using NumPy array with datetime index and labeled column
df1 = pd.date_range('20130101', periods=6)
df1 = pd.DataFrame(np.random.randn(6, 4), index=df1, columns=list('ABCD'))
df1 # Display dataframe df1
```
### Viewing data
```
df1 = pd.date_range('20130101', periods=6)
df1 = pd.DataFrame(np.random.randn(6, 4), index=df1, columns=list('ABCD'))
df1.head(2) # View top data
df1.tail(2) # View bottom data
df1.index # Display index column
df1.dtypes # Inspect datatypes
df1.describe() # Display quick statistics summary of data
```
### Subsetting, slicing, and indexing
```
df1 = pd.date_range('20130101', periods=6)
df1 = pd.DataFrame(np.random.randn(6, 4), index=df1, columns=list('ABCD'))
df1.T # Transpose data
df1.sort_index(axis=1, ascending=False) # Sort by an axis
df1.sort_values(by='B') # Sort by values
df1['A'] # Select column A
df1[0:3] # Select index 0 to 2
df1['20130102':'20130104'] # Select from index matching the values
df1.loc[:, ['A', 'B']] # Select on a multi-axis by label
df1.iloc[3] # Select via the position of the passed integers
df1[df1 > 0] # Select values from a DataFrame where a boolean condition is met
df2 = df1.copy() # Copy the df1 dataset to df2
df2['E'] = ['one', 'one', 'two', 'three', 'four', 'three'] # Add column E with value
df2[df2['E'].isin(['two', 'four'])] # Use isin method for filtering
```
### Missing data
Pandas primarily uses the value `np.nan` to represent missing data. It is not included in computations by default.
```
df = pd.DataFrame({'num_legs': [2, 4, np.nan, 0],
'num_wings': [2, 0, 0, 0],
'num_specimen_seen': [10, np.nan, 1, 8]},
index=['falcon', 'dog', 'spider', 'fish'])
df.dropna(how='any') # Drop any rows that have missing data
df.dropna(how='any', axis=1) # Drop any columns that have missing data
df.fillna(value=5) # Fill missing data with value 5
pd.isna(df) # To get boolean mask where data is missing
```
### File handling
```
df = pd.DataFrame({'num_legs': [2, 4, np.nan, 0],
'num_wings': [2, 0, 0, 0],
'num_specimen_seen': [10, np.nan, 1, 8]},
index=['falcon', 'dog', 'spider', 'fish'])
df.to_csv('foo.csv') # Write to CSV file
pd.read_csv('foo.csv') # Read from CSV file
df.to_excel('foo.xlsx', sheet_name='Sheet1') # Write to Microsoft Excel file
pd.read_excel('foo.xlsx', 'Sheet1', index_col=None, na_values=['NA'], engine='openpyxl') # Read from Microsoft Excel file
```
### Plotting
```
# Install Matplotlib using pip
!pip install matplotlib
from matplotlib import pyplot as plt # Import Matplotlib module
# Generate random time-series data
ts = pd.Series(np.random.randn(1000),index=pd.date_range('1/1/2000', periods=1000))
ts.head()
ts = ts.cumsum()
ts.plot() # Plot graph
plt.show()
# On a DataFrame, the plot() method is convenient to plot all of the columns with labels
df4 = pd.DataFrame(np.random.randn(1000, 4), index=ts.index,columns=['A', 'B', 'C', 'D'])
df4 = df4.cumsum()
df4.head()
df4.plot()
plt.show()
```
| github_jupyter |
```
#Installing MTCNN Library for Face Detection
!pip install mtcnn
#Importing MTCNN and OpenCV
from mtcnn import MTCNN
import cv2
def funci(img)
img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
#Using the MTCNN Model
detector = MTCNN()
count =0
#Passing the IMage through Face Detection MTCNN
result = detector.detect_faces(img)
img= cv2.cvtColor(img, cv2.COLOR_RGB2BGR)
#Going through all the faces in the result
for i in range(len(result)):
#Coordinates of the face bounding box
bounding_box = result[i]['box']
#Checking for confidence of the bounding box
if(result[i]['confidence']>0.9724391102790833):
#cv2.rectangle(img,
#(bounding_box[0], bounding_box[1]),
#(bounding_box[0]+bounding_box[2], bounding_box[1] + bounding_box[3]),
#(0,255,0),
#2)
#cv2.circle(img,result[i]['keypoints']['left_eye'],radius=5,color=(0,255,0),thickness=-1)
#cv2.circle(img,result[i]['keypoints']['right_eye'],radius=5,color=(0,255,0),thickness=-1)
#Coordinates of Left and right eye
x=(result[i]['keypoints']['left_eye'][0]+result[i]['keypoints']['right_eye'][0])//2
y=(result[i]['keypoints']['left_eye'][1]+result[i]['keypoints']['right_eye'][1])//2
distanceleft=x-result[i]['keypoints']['left_eye'][0]
#cv2.circle(img,(result[i]['keypoints']['left_eye'][0]-distanceleft,result[i]['keypoints']['left_eye'][1]-distanceleft//3),radius=5,color=(0,255,0),thickness=-1)
distanceright=result[i]['keypoints']['right_eye'][0]-x
#cv2.circle(img,(result[i]['keypoints']['right_eye'][0]+distanceright,result[i]['keypoints']['right_eye'][1]-distanceright//3),radius=5,color=(0,255,0),thickness=-1)
print("{},{}".format(i,result[i]['confidence']))
#Reading the image of the glasses
glasses=cv2.imread("glasses.png",-1)
#Taking bounding box for fitting in spectacles
x_off = result[i]['keypoints']['left_eye'][0]-distanceleft
y_off = result[i]['keypoints']['left_eye'][1]-distanceleft//2
x_end = result[i]['keypoints']['right_eye'][0]+distanceright
y_end = result[i]['keypoints']['right_eye'][1]-distanceright//2
wid=int(((x_off-x_end)**2+(y_off-y_end)**2)**0.5)
#Resizing the image of spectacles as per the width of the above bounding box
glasses2=cv2.resize(glasses.copy(),(wid,wid//3),interpolation=cv2.INTER_LINEAR_EXACT)
gw,gh,gc=glasses2.shape
print("Image Shape",img.shape)
print("Specs shape",glasses2.shape)
#Superimposing the glasses over the face image
for i in range(0,gw):
for j in range(0,gh):
if(glasses2[i,j][3]!=0):
img[y_off+i,x_off+j]=glasses2[i,j][0:3]
#cv2.rectangle(img, (x_off,y_off),(x_end,y_end+wid//3),(0,0,255), thickness=1)
#Saving the image with spectacles on
cv2.imwrite("Result.jpeg",img)
#cv2.imshow("Image",img)
#cv2.waitKey(0)
#cv2.destroyAllWindows()
#Reading the image of face and passing it through the model
img=cv2.imread("IMG_20190715_120407-01.jpeg")
funci(img)
```
| github_jupyter |
# Appendix B: Confidence and prediction intervals for univariate 0D data
This appendix describes how to construct and numerically validate [confidence intervals](https://en.wikipedia.org/wiki/Confidence_interval) (CIs) and [prediction intervals](https://en.wikipedia.org/wiki/Prediction_interval) (PIs) for [univariate](https://en.wikipedia.org/wiki/Univariate) 0D data. "Univariate" implies a scalar quantity and "0D" implies a zero-dimensional, stationary process (i.e, a quantity that does not change over some domain like time). Below and throughout this Supplementary Material a Type I error rate ($\alpha$=0.05) is used, implying 95% CIs and PIs.
<p style="text-align: center;"><b>Glossary</b></p>
<table align="center">
<tr>
<th style="text-align:center">Symbol</th>
<th style="text-align:center">Description</th>
<th style="text-align:center">Comment</th>
</tr>
<tr>
<td style="text-align:center">$\alpha$</td>
<td style="text-align:center">Type I error rate</td>
<td style="text-align:center">$\alpha=0.05$ by convention; univariate intervals use $\alpha$/2 for two-tailed inference</td>
</tr>
<tr>
<td style="text-align:center">$\mu$</td>
<td style="text-align:center">Population mean</td>
<td style="text-align:center">$\mu = 0 $ in all examples below </td>
</tr>
<tr>
<td style="text-align:center">$\sigma$</td>
<td style="text-align:center">Population standard deviation</td>
<td style="text-align:center">$\sigma=1$ in all examples below</td>
</tr>
<tr>
<td style="text-align:center">$\textit{J}$</td>
<td style="text-align:center">Sample size</td>
<td style="text-align:center"></td>
</tr>
<tr>
<td style="text-align:center">$\nu$</td>
<td style="text-align:center">Degrees of freedom</td>
<td style="text-align:center">For univariate data: $\nu=J-1$</td>
</tr>
<tr>
<td style="text-align:center">$c$</td>
<td style="text-align:center">Critical test statistic value</td>
<td style="text-align:center">From Student's t distribution</td>
</tr>
<tr>
<td style="text-align:center">$\overline{y}$</td>
<td style="text-align:center">Sample mean</td>
<td style="text-align:center"></td>
</tr>
<tr>
<td style="text-align:center">$s$</td>
<td style="text-align:center">Sample standard deviation</td>
<td style="text-align:center"></td>
</tr>
</table>
___
### Confidence intervals (CIs)
A 100(1-$\alpha$)% [confidence interval](https://en.wikipedia.org/wiki/Confidence_interval) contains the true population mean with a probability of (1-$\alpha$), and is defined as:
CI $\equiv \overline{y} \pm \frac{cs}{\sqrt{J}}$
where $c$ is the $(1-\frac{\alpha}{2})$th percentile of the Student's t distribution with $\nu=(J-1)$ degrees of freedom.
To demonstrate the computation of a 95% CI, let's start with random data from the standard normal distribution (i.e., $\mu$=0, $\sigma$=1):
```
import numpy as np
from scipy import stats
%matplotlib inline
from matplotlib import pyplot as plt
from matplotlib.patches import Polygon
from matplotlib.collections import PatchCollection
'''
Note: the random number generator is seeded with a value "32" instead of
a more usual value like "0" or "1".
This is done deliberately, to demonstrate a case where a random sample's
confidence interval does not contain the true population mean (i.e., zero)
'''
np.random.seed(32) # seed the random number generator to replicate results
J = 8 # sample size
y = np.random.randn(J) # Gaussian data (standard normal distribution)
fig,ax = plt.subplots(1, 1, figsize=(8,3))
ax.plot(y, [0]*J, 'bo', ms=8, zorder=1, label='Gaussian data ($\mu$=0)')
ax.axhline(0, color='0.7', ls='-', zorder=0)
ax.axvline(0, color='0.7', ls='-', zorder=0)
ax.set_yticks([])
ax.legend()
plt.show()
```
The `confidence_interval` function below implements the equation above. For convenience and subsequent use, plotting functions are also provided below.
```
def confidence_interval(y, alpha=0.05):
'''
Compute confidence interval for 0D univariate data.
'''
J = y.size # sample size
ybar = y.mean() # sample mean
s = y.std(ddof=1) # sample standard deviation
df = J - 1 # degrees of freedom
c = stats.t.isf(alpha/2, df) # critical test statistic
w = c * s / J**0.5 # interval width
ci = ybar-w, ybar+w # confidence interval
return np.array(ci)
def plot_data(ax, y, plot_sample_mean=True, population_mean=None):
'''
Scatter plot for univariate 0D data.
'''
ax.plot(y, np.zeros(J), 'ko', label='Observations')
if plot_sample_mean:
ax.plot(y.mean(), 0, 'ko', label='Sample mean', ms=15, mfc='w')
if population_mean is not None:
mu = population_mean
ax.plot(mu, 0, 'go', label='True population mean', ms=15, mfc='w', mew=5)
ax.axvline(mu, color='k', ls=':', zorder=-1)
def plot_interval(y, interval, color='r', height=1, alpha=0.05, label='Confidence Interval'):
'''
Interval plot for univariate 0D data.
'''
h,hh = height, 0.5 * height
x = interval
ax.plot( [x[0]]*2, [-h, h], color=color, label='%d%s %s' %(100*(1-alpha),'%', label))
ax.plot( [x[1]]*2, [-h, h], color=color)
vertices = [(x[0], -hh), (x[1], -hh), (x[1], hh), (x[0], hh)]
patches = PatchCollection([Polygon(vertices)])
ax.add_collection(patches)
plt.setp(patches, facecolor=color, alpha=0.5, linewidth=0)
ax.set_ylim(-2, 5)
ax.set_yticks([])
# compute 95% confidence interval:
ci = confidence_interval(y, alpha=0.05)
print('Confidence interval: %s' %ci)
# plot:
fig, ax = plt.subplots(1, 1, figsize=(8, 4))
plot_data(ax, y, plot_sample_mean=True, population_mean=0)
plot_interval(ax, ci)
ax.legend()
plt.show()
```
Note that a 95% CI is expected to contain the true population mean in 95% of a large number of experiments. In this case it does not contain the population mean, implying a [false positive](https://en.wikipedia.org/wiki/False_positives_and_false_negatives#False_positive_error). This result is a false positive because one would incorrectly conclude that the sample mean is significantly different from the true population mean of $\mu$=0.
___
### Prediction intervals
A 100(1-$\alpha$)% [prediction interval](https://en.wikipedia.org/wiki/Prediction_interval) contains a new observation with a probability of $(1-\alpha)$, and is defined as:
PI $\equiv \overline{y} \pm cs \sqrt{1+(1/J)}$
```
def prediction_interval(y, alpha=0.05):
J = y.shape[0]
ybar = y.mean() # sample mean
s = y.std(ddof=1) # sample standard deviation
df = J - 1 # degrees of freedom
c = stats.t.isf(alpha/2, df) # critical test statistic
w = c * s * (1+(1/J))**0.5 # interval width
pi = ybar-w, ybar+w # prediction interval
return np.array(pi)
pi = prediction_interval(y)
# compute 95% prediction interval:
pi = prediction_interval(y, alpha=0.05)
print('Prediction interval: %s' %pi)
# plot:
fig, ax = plt.subplots(1, 1, figsize=(8, 4))
plot_data(ax, y, plot_sample_mean=True, population_mean=0)
plot_interval(ax, pi, color='b', label='Prediction Interval')
ax.legend()
plt.show()
```
We could generate a new random observation, then check whether it lies in the PI:
```
yy = float( np.random.randn(1) )
print(yy)
```
This observation lies within the PI, as expected. A false positive would have occurred had this new observation lay outside the PI.
Last, let's visualize the CI and PI together:
```
fig, ax = plt.subplots(1, 1, figsize=(8, 4))
plot_data(ax, y, plot_sample_mean=True, population_mean=0)
plot_interval(ax, pi, color='b', height=0.5, label='Prediction Interval')
plot_interval(ax, ci, color='r', height=1, label='Confidence Interval')
ax.legend()
plt.show()
```
___
### Numerically validating confidence and prediction intervals
CIs can be numerically validated by simulating a large number of datasets, constructing the 95% CI for each experiment, and then checking whether the true mean lies within the CI.
```
np.random.seed(0) # seed the random number generator to replicate results
J = 10 # sample size
niterations = 2000 # number of datasets / experiments to simulate
in_ci = [] # list that will hold one True or False value for each iteration
mu = 0 # true population mean
alpha = 0.05 # Type I error rate
for i in range(niterations):
y = mu + np.random.randn(J) #Gaussian data
ci = confidence_interval(y, alpha)
in_ci.append( (mu>=ci[0]) and (mu<=ci[1]) )
prop_in = np.mean( in_ci ) # proportion of experiments where the true mean lies inside the CI
prop_out = 1 - prop_in # proportion of experiments where the true mean lies outside the CI
print('Proportion of random datasets with mu inside CI: %.3f' %prop_in)
print('Proportion of random datasets with mu outside CI: %.3f' %prop_out)
```
The proprotion of random datasets with $\mu$ outside the CI are, as expected, close to $\alpha$. However, this proportion is not exactly $\alpha$ becaus $\alpha$ pertains to an infinite number of datasets, or `niterations` = $\infty$. Increasing `niterations` to a larger number, like 10,000 or 100,000 or 1,000,000, should cause closer convergence to $\alpha$.
This general procedure of simulating random datasets will be used throughout this supplementary material.
Let's repeat for prediction intervals:
```
np.random.seed(1) # seed the random number generator to replicate results
J = 8 # sample size
niterations = 2000 # number of datasets / experiments to simulate
in_pi = [] # list that will hold one True or False value for each iteration
mu = 0 # true population mean
alpha = 0.05 # Type I error rate
for i in range(niterations):
y = mu + np.random.randn(J) # Gaussian data
yy = float(mu + np.random.randn(1)) # a new random observation
pi = prediction_interval(y, alpha)
in_pi.append( (yy>=pi[0]) and (yy<=pi[1]) )
prop_in = np.mean( in_pi ) # proportion of experiments where the true mean lies inside the PI
prop_out = 1 - prop_in # proportion of experiments where the true mean lies outside the PI
print('Proportion of random datasets with new observation inside PI: %.3f' %prop_in)
print('Proportion of random datasets with new observation outside PI: %.3f' %prop_out)
```
Like above the results converge to $\alpha$.
___
### PI-to-CI size ratio
The CI and PI definitions from above are:
<center>CI $\equiv \overline{y} \pm \frac{cs}{\sqrt{J}}$</center>
<center>PI $\equiv \overline{y} \pm cs \sqrt{1+(1/J)}$</center>
These can be re-written as:
<center>CI $\equiv \overline{y} \pm w_0$</center>
<center>PI $\equiv \overline{y} \pm w_1$</center>
where:
<center>$w_1 = w_0 * J \sqrt{\frac{1}{(1+J)}}$</center>
which is apprximately:
<center>$w_1 \approx w_0 * \sqrt{J}$</center>
as sample size ($J$) becomes large. Thus PIs are effectively CIs that are scaled by the square-root of sample size.
```
J = np.arange(5, 51)
w1w0_ratio = J * np.sqrt( 1 / (1+J) )
fig,ax = plt.subplots(1, 1, figsize=(6,4))
ax.plot(J, w1w0_ratio, '-', label=r'$w_1 = w_0 * J \sqrt{\frac{1}{(1+J)}}$')
ax.plot(J, np.sqrt(J), '-', label=r'$w_1 = w_0 * \sqrt{J}$')
ax.legend()
ax.set_xlabel('Sample size ($J$)')
ax.set_ylabel('PI-CI size ratio ($w_1 / w_0$)')
plt.show()
```
Thus, for sample sizes of 5, 10 and 50, the PI:CI size ratios are 2.0, 3.0 and 7.0, respectively.
___
### Summary
This Appendix demonstrated the difference between CIs and PIs, how to calculate them, and how to numerically validate them using iterative, random dataset simulations. The key points are:
* A false positive occurs for a CI when the true population mean lies outside the CI
* A false positive occurs for a PI when a new, random observation lies outside the PI
and
* 95% CIs contain the true population mean with a probability of 0.95
* 95% PIs contain a new, random observation with a probability of 0.95
* 95% CIs and PIs both have false positive rates of $\alpha$=0.05
* The PI:CI size ratio is approximately equivalent to the square-root of sample size.
| github_jupyter |
# Tips dataset analysis
Description: Fundamentals of Data Analysis - assignment project, GMIT 2019. See README.md for more background info.
>Author: **Andrzej Kocielski**
>Github: [andkoc001](https://github.com/andkoc001/)
>Email: G00376291@gmit.ie, and.koc001@gmail.com
Date of creation: 23-09-2019
This Notebook should be read in conjunction with the corresponding README.md file at the assignment repository at GitHub: <https://github.com/andkoc001/Tips_dataset_analysis/>.
___
## Introduction
### The Analysis
This Notebook presents my analysis and interpretation of the _**tips dataset**_. This is done through the following:
1. descriptive statistics of the raw data,
2. graphical representation of the data - plots,
3. inference of the implicit information by application of selected tools, methods and algorithms used in data analytics.
### The data
It is a common custom to offer some small extra money - a tip - to the staff of a restaurant on top of the bill after a satisfactory service received. Although the tips are voluntary, and the amount of the tips is not (usually) stated, by convention it is often advised to leave as a certain percent (e.g. 10%) of the total bill for the meal and service.
The _tips dataset_ (being analysed in this Notebook) is a representation of tips given in a restaurant. The dataset in question is a record (allegedly real and true) of tips given along with total bills of a restaurant customers collated by a waiter working in the restaurant for several weeks.
The data is organised in a form of an array, where the dataset attributes (aka features) are organised in columns, and the observations (aka instances) - in rows. The dataset consists of several data categories describing tips received in connection to circumstances, such as day of the week, gender of the tipper, etc. The data set includes 244 data observations.
The _tips dataset_ is integrated with the [Seaborn](https://seaborn.pydata.org/) - an external Python package for data visualisation. This dataset from this source will be used in the subsequent analysis. The dataset, however, can be also obtained from other sources, for example as a .csv file from [here](http://vincentarelbundock.github.io/Rdatasets/datasets.html).
### Hypothesis
The dataset provokes to ask the following question: **Is there a consistent linear relationship between _total bill_ and _amount of tip_?**
Below we will try to find evidences either supporting or the opposite this hypothesis. I will also delve deeper into the question and seek whether I could distinguish such a relationship considering other parameters, such as among smokers and non-smokers.
### Assumptions
* The data is accurate.
* The gender and the smoker refers to the person in the group, who payed the bill and left the tip.
* Currency is irrelevant.
* Any other information that might be normally applicable to bill-tip relationship (e.g. country, legislation and regulations, time of the year, etc.) and is not included in the dataset is irrelevant.
___
## Setting up the environment
### Importing additional packages
The analysis will be conducted using numerical tools, specifically **Python** programming language. On top of that, I am going to use add-on Python libraries - **NumPy** (for numerical calculations), **Pandas** (for convenient data structures - DataFrames), as well as visualisation libraries - **Matplotlib** (its pyplot sub-library) and **Seaborn**.
```
# numerical calculations libraries, assigning aliases
import numpy as np
import pandas as pd
# plotting and data visualisation libraries, assigning aliases
import matplotlib.pyplot as plt
import seaborn as sns
# below command will allow for the plots being displayed inside the Notebook, rather than in a separate screen.
%matplotlib inline
```
### Loading the data set
Getting the data from _Seaborn_ package and assigning it to variable `tips`.
```
# Loading the data set
tips = sns.load_dataset("tips")
```
### The dataset basic properties and its integrity check
Prior to actual data analysis, it is a good practice to check the data integrity, that is whether there are any empty cells or corrupted data. We will use for this purpose the Pandas function `info()`, which checks if there is any `null` value in _any_ column. This function also checks data type for each column, as well as number of each data types and number of observations (rows).
```
tips.info()
```
Hence, there are no 'empty' cells in the dataset, and the above looks good. Perhaps the _size_ variable could be change to int8 type, so that the memory usage could be optimised, but considering the small size of the dataset, I am going to leave it as is.
The initial checks show its integrity and I am satisfied to continue with further analysis.
___
## Data analysis
### 1. Exploratory data analysis (descriptive)
#### Sneak peek into raw data
Below are listed several first raws of data displayed. This listing allows to get initial impression on the dataset structure, as well as its attributes (columns) and data types of the variables.
```
tips.head(3)
```
For a quick glimpse into numeric values and basic statistical description, it is convenient to call `describe()` method. As a result it will show for each numerical attribute (column) information such as number of occurrence, minimum, median (50%) and maximum values, mean and standard deviation.
```
# calling describe method; percentiles will be limited to 50%, to keep the result compact; adapted from https://www.geeksforgeeks.org/python-pandas-dataframe-describe-method/
tips.describe(percentiles = [0.5])
```
Interesting to see that tips were given every time. Also, the percent of mean tips is $13.16\%$ (=2.998279÷(19.785943+2.998279)).
Now, let's also see what are the unique values occur in each non-numeric (categorical) column, using Pandas' method `unique()`.
```
print("sex:\t", sorted(tips.sex.unique()))
print("smoker:\t", sorted(tips.smoker.unique()))
# List unique values in the categorical atributes
# Adapted from https://chrisalbon.com/python/data_wrangling/pandas_list_unique_values_in_column/ and https://jovianlin.io/data-visualization-seaborn-part-1/.
print("Values occuring in column 'sex':\t", sorted(tips.sex.unique()))
print("Values occuring in column 'smoker':\t", sorted(tips.smoker.unique()))
print("Values occuring in column 'day':\t", sorted(tips.day.unique()))
print("Values occuring in column 'time':\t", sorted(tips.time.unique()))
```
#### Raw data modeling
It would be interesting to consider and evaluate the relationship between the data which is not explicitly given. For this purpose, let's model the dataset by creating new attributes. I am going to create the following:
- `tip_ratio` - ratio of tip to bill,
- `sum` - the sum of total bill and tip,
- `percent` - tip as a percent of sum,
- `bpp` - amount of total bill divided by number of people in the group,
- `tpp` - amount of tip per person.
```
# new column created - percent of tip
tips["tip_ratio"] = tips["tip"]/tips["total_bill"]
# new column created - sum of total bill and tip
tips["sum"] = tips["total_bill"]+tips["tip"] # appended at the end of the array
# new column created - ratio of tip to sum
tips["percent"] = round(tips["tip"]/tips["sum"]*100, 2)
# add column: bpp - bill per person
tips["bpp"] = tips["total_bill"]/tips["size"]
# add column: tpp - tip per person
tips["tpp"] = tips["tip"]/tips["size"]
```
Now, let's see what the dataset array looks like with the additional attributes. To keep it compact, just a few first rows.
```
tips.head(2)
```
Let's use the `describe()` method again for the upgraded data array.
```
# calling describe method; percentiles will be limited to 50%, to keep the result compact
tips.describe(percentiles = [0.5])
```
From the above we can get some interesting observations:
1. on average for the entire dataset, the percent of tip is $13.64\%$ (which is slightly higher than the value calculated above for mean tip and mean total bill),
2. on average, total bill split equally in the group is 7.89,
3. on average, tip per person is 1.21.
For comparison, I am going to apply a filter and see what is, for example, the average ratio tip to total bill among smokers.
```
# mean tip per smoking person
print("Average tip per smoking person:\t\t\t", tips[tips["smoker"] == "Yes"]["tpp"].mean()) # filtering by applying conditional check (smoker=yes)
# mean total bill among smokers
print("Average total bill among smokers:\t\t", tips[tips["smoker"] == "Yes"]["total_bill"].mean())
# mean ratio among smokers
tips["ratio_smoke"] = tips[tips["smoker"] == "Yes"]["tip"] / tips[tips["smoker"] == "Yes"]["total_bill"]
print("Average ratio tip to total bill among smokers:\t", tips["ratio_smoke"].mean())
```
Interesting. It turns out, the smokers are more generous and pay more for the service as well as give higher tips. On average the ratio tip to total bill among smokers is $16.32\%$.
### 2. Graphical interpretation
Some information can be revealed easily by visualising the data graphically on carefully crafted plots, according to the popular saying "A single plot is worth a thousand of data points".
In this section, we will first look at the raw data in the set. The attempt made here is to visualise the figures, applying only built-in analytical tools. My intention is to draw attention to some interesting (in my view) and yet rather straightforward relationships between the data. Later on, I will try to apply more sophisticated analytical tools - specifically liner regression, with aim to reveal some less obvious relationships and patterns existing in the dataset.
For a better readability, let's change the Seaborn global plots style as follows (https://seaborn.pydata.org/tutorial/aesthetics.html, https://www.datacamp.com/community/tutorials/seaborn-python-tutorial)
```
sns.set_style("darkgrid") # plot style
sns.set_palette("muted") # palette of colours
# reference: https://seaborn.pydata.org/tutorial/color_palettes.html, https://stackoverflow.com/a/47955814
```
#### Bar plots
Bar plots (also histograms, although not being the same class of graphs, [reference](https://keydifferences.com/difference-between-histogram-and-bar-graph.html)) allow to easily display the categorical variables and showing their distributions, especially in comparison with one or more similar categories of data.
To begin with, let us see the tips distribution on a histogram, using `distplot` method. With a large number of bins, the tips intervals will be more precise, and it still clearer than `countplot` method.
```
# Display setting
plt.subplots(figsize=(12,3))
plt.legend(prop={'size': 10})
plt.suptitle("Figure 1. How many times the same tip amount was offered.", y=1.0, fontsize=14)
#plt.xlabel('Tip')
plt.ylabel('Count')
# plot data and properties
sns.distplot(tips['tip'], bins=100, kde=False, color="k")
plt.show()
```
From the above figure (Fig.1) we can see that distinctly the most popular tip is 2 (units of currency), with a runner up at 3 (units of currency).
In the below figure (Fig. 2), there is shown the distribution between parties size on the left plot, and on the right plot - distribution of smokers and non-smokers among all the customers, further broken down on the tippers' gender. For this purpose `seaborn.countplot()` function is used ([Seaborn documentation](https://seaborn.pydata.org/generated/seaborn.countplot.html)).
The below plots show the number of customers split onto days the restaurant was open. The second plot is further broken down by the size of the party (colour coded - see the legend).
```
# Display setting
fig, ax = plt.subplots(ncols=2, figsize=(12, 2))
plt.suptitle("Figure 2. Number of clients on each day.", y=1.1, fontsize=14)
plt.subplots_adjust(left=None, bottom=None, right=None, top=None, wspace=.3, hspace=None)
# plot data and propertiesy
sns.countplot(data=tips, x="day", palette="pastel", edgecolor=".5", ax=ax[0]) # number of clients on each day
sns.countplot(data=tips, x="day", palette="pastel", edgecolor=".5", hue='size', ax=ax[1]) # additional discrimination on party sizy
plt.legend(bbox_to_anchor=(1.01, 1), loc=2, borderaxespad=0.) # Put the legend out of the figure, taken from https://stackoverflow.com/a/34579525
plt.show()
# Display setting
fig, ax = plt.subplots(ncols=2, figsize=(12, 2))
plt.suptitle("Figure 3. Pary size and clients profile.", y=1.1, fontsize=14)
plt.subplots_adjust(left=None, bottom=None, right=None, top=None, wspace=.3, hspace=None)
# plot data and properties
sns.countplot(data=tips, y="size", palette="pastel", edgecolor=".5", ax=ax[0]) # number of clients on each day
sns.countplot(data=tips, y="smoker", palette="pastel", edgecolor=".5", hue="sex", ax=ax[1]) # additional discrimination on party sizy
plt.legend(bbox_to_anchor=(1.01, 1), loc=2, borderaxespad=0.0, title="Gender") # Put the legend out of the figure, taken from https://stackoverflow.com/a/34579525
plt.show()
```
As we can see, two-person groups dominate among the restaurant clients, but three- and four-person size parties are also frequent.Concerning the tipper gender and smoking habits, it turns out that it is males that usually pay the bill. Also, the number of non-smokers outbalance the number of smoking tippers.
Below, we see amount of tip offered each day, further broken down on whether the client was a smoker or not. A convenient tool for illustrating categorical data is `seaborn.barplot()` method ([Seaborn docs](https://seaborn.pydata.org/generated/seaborn.barplot.html)), and it will be used in this case. The top (the length of the bars) of the bars represent the _mean_ (in the below plots, default) of the attribute. On top of that, _Seaborn_ allows to show _confidence interval_ of the feature as well - the length of the black line. The default value of 95% confidence intervals is used in the plots below.
```
# adapted from https://medium.com/@neuralnets/data-visualization-with-python-and-seaborn-part-1-29c9478a8700
fig, ax = plt.subplots(ncols=1, figsize=(6,3))
plt.title('Figure 4. Mean amount of tips on each day.')
# Mean amount of tip on each day
sns.barplot(data = tips, x = "day", y = "tip", hue="smoker", palette="pastel", edgecolor=".5")
plt.legend(bbox_to_anchor=(1.01, 1), loc=2, borderaxespad=0.0, title="Smoker") # legend put outside of the plot area
plt.show()
```
#### Scatter plots
Scatter plots allow for relating two (or more) numerical variables. In the below plots, function `seaborn.relplot()` is used, with the default kind of the plot (scatter), to show statistical relationship between amount of tip on y-axis and total bill on x-axis. This section is based on official Seaborn [tutorial](https://seaborn.pydata.org/tutorial/relational.html).
```
sns.relplot(data=tips, x="total_bill", y="tip", s=20, palette="pastel", height=4, aspect=2)
plt.title("Figure 5. Total bill to amount of tip.", fontsize=14)
plt.show()
```
The above plot may appear over crowded with data points. By adding another variable (for example _smoker_) it may become clearer and unveil a potential pattern, further separated by days.
The above plot indeed seem to show a linear relationship between the two variables. Before verifying that (calculation the minimum _cost_ value) in the next section, let's take a look at some other also interesting plots.
```
fig, ax = plt.subplots(ncols=2, figsize=(12, 4))
plt.suptitle("Figure 6. Tip to total bill for smokers and non-smokers.", y=1.01, fontsize=14)
plt.subplots_adjust(left=None, bottom=None, right=None, top=None, wspace=.3, hspace=None)
# plot #1 - Smokers
sns.swarmplot(x="total_bill", y="tip", hue="smoker", palette="muted", marker="o", s=4, data=tips[tips["smoker"] == "Yes"], ax=ax[0])
# plot #2 - Non-smokers
sns.swarmplot(x="total_bill", y="tip", hue="smoker", palette="muted", marker="o", s=4, data=tips[tips["smoker"] == "No"], ax=ax[1])
plt.legend(bbox_to_anchor=(1.01, 1), loc=2, borderaxespad=0.0, title="Smoker") # legend put outside of the plot area
fig.show()
g = sns.relplot(data=tips, x="total_bill", y="tip", col="day", row="smoker", hue="smoker", palette="muted", marker="o", alpha=.7, s=30, height=4, aspect=1)
# Figure title; adopted from https://stackoverflow.com/a/54213918
g.fig.suptitle("Figure 7. Tip to total bill on each day.", x=0.5, y=1.02, fontsize=14)
plt.show()
```
Except for Sunday smokers plot, which looks a bit random, there seems to be a distinct positive relationship between the total bill and the amount of tip.
What about the amount of tips compared to the size of the party on given days? Can we see any pattern there? We will apply `catplot()` method to generate plots.
```
g = sns.catplot(data=tips, x="size", y="tip", col="day", palette="muted", alpha=.7, s=4, height=4, aspect=1)
# Figure title
g.fig.suptitle("Figure 8. Tips per day and per party size.", x=0.5, y=1.04, fontsize=14)
plt.show()
```
From the above plots, it appear there indeed is a relationship.
### 3.1 Regression
Regression allows to analyse the relationship between one variable (dependent variable) and other variables (independent variables, may be more than one).
#### Minimising the cost
This part is based on the lecture concerning linear regression and the lecturer's [Notebook](https://github.com/ianmcloughlin/jupyter-teaching-notebooks/raw/master/simple-linear-regression.ipynb).
In this section I will test the hypothesis question stated above (what is the relationship (if any) between total bill and amount of tips), by estimating a regression line between the variables. I will see if there is a relationship between the amount of tip and total bill, which could be accurately described with $ y = m x + 1 $ equation. "The number $ m $ is the slope of the line. The slope is how much $ y $ increases by when $ x $ is increased by 1.0. The number $ c $ is the y-intercept of the line."
Taken from the lecturer's [Notebook](https://github.com/ianmcloughlin/jupyter-teaching-notebooks/raw/master/simple-linear-regression.ipynb):
>We want to calculate values of $m$ and $c$ that give the lowest value for the cost value above.
For our given data set we can plot the cost value/function.
Recall that the cost is:
> $$ Cost(m, c) = \sum_i (y_i - mx_i - c)^2 $$
>This is a function of two variables, $m$ and $c$, so a plot of it is three dimensional. (...)
>In the case of fitting a two-dimensional line to a few data points, we can easily calculate exactly the best values of $m$ and $c$. (...)
We first calculate the mean (average) values of our $x$ values and that of our $y$ values.
Then we subtract the mean of $x$ from each of the $x$ values, and the mean of $y$ from each of the $y$ values.
Then we take the *dot product* of the new $x$ values and the new $y$ values and divide it by the dot product of the new $x$ values with themselves.
That gives us $m$, and we use $m$ to calculate $c$.
We will use `polyfit()` method to calculate $m$ and $c$.
```
line_eq = np.polyfit(tips["total_bill"], tips["tip"], 1) # fiting a stright line (first degree); assign to a list type
# m is stored in the first item of line_eq (index 0),
# c is stored in the second item of line_eq (index 1).
# Printing results with comment
print("Value of m is", line_eq[0], ", and value of c is", line_eq[1],".")
```
Putting the scatter plot and the calculated regression line in one plot (Fig. 9).
```
# Display setting
fig, ax = plt.subplots(figsize=(16,3))
plt.suptitle("Figure 9. Overlay regression line over the scatter plot.", x=0.5, y=1.01, fontsize=14)
plt.subplots_adjust(left=None, bottom=None, right=None, top=None, wspace=0.1, hspace=None)
# plot #1 - scatter plot
plt.plot(tips["total_bill"], tips["tip"], "b.", alpha=0.3, label="data") # pyplot
#sns.relplot(data=tips, x="total_bill", y="tip", s=20, ax=ax[0]) # seaborn - does not work as intended
# plot #2 - regression line within x-axis range between minimum and maximum total bill values
x = np.linspace(-10, 65, 2) # two values of 'y' for x at -10 and at 65
plt.plot(x, line_eq[0]*x + line_eq[1], 'r-', alpha=0.5); # drawing the regression line with calculated minimum cost; pyplot
#sns.lineplot(x, line_eq[0]*x + line_eq[1], ax=ax[0]) # drawing the regression line with calculated minimum cost; seaborn - does not work as intended
# setting plotting limits
ax.set_ylim(0, 11)
ax.set_xlim(0, 55)
# Axis labels
plt.xlabel('Total bill')
plt.ylabel('Tip')
plt.show()
```
#### Seaborn built-in functionality
A very similar result the above plot can be generated with Seaborn's `regplot()` method, which offers a built-in functionality for drawing linear regression line automatically. Once again, let's make a plot showing total bill against tip, with additional regression line to the plot (Fig.10). Based on the Seaborn [tutorial](https://seaborn.pydata.org/tutorial/regression.html).
```
# Display setting
fig, ax = plt.subplots(figsize=(16,3))
plt.suptitle("Figure 10. Overlay regression line over the scatter plot - with regplot().", x=0.5, y=1.01, fontsize=14)
plt.subplots_adjust(left=None, bottom=None, right=None, top=None, wspace=0.1, hspace=None)
sns.regplot(data=tips, x="total_bill", y="tip", ci=0, scatter_kws={"s": 6, "alpha":0.6}, line_kws={"lw":1, "color":"r", "alpha":0.6}) # confidence interval set to 0 for clarity
plt.show()
```
Let's now take a look at other, arguably more interesting plots that can be produced from the dataset.
First, let us now consider the data with variables discrimination on each day and whether the client was a smoker or not (Fig.11).
```
# adapded from https://stackoverflow.com/a/25213438
lm = sns.lmplot(data=tips, x="total_bill", y="tip", col="day", row="smoker", hue="smoker", ci=0, height=4, aspect=1, palette="muted", scatter_kws={"s":8}, line_kws={"lw":2, "alpha":0.4})
# plot's title
plt.suptitle("Figure 11. Amount of tip in relation to total bill on various days.", x=0.5, y=1.03, fontsize=14)
# axes limits
axes = lm.axes
axes[0,0].set_ylim(0,11)
axes[0,0].set_xlim(0,55)
plt.show()
```
An interesting results yield the next set of plots (Fig.12), showing a tendency of decrease tip amount per person for larger group of people (slope of the regression line).
```
# defining the plot as an object
lm = sns.lmplot(data=tips, x="total_bill", y="tpp", col="size", row="smoker", hue="smoker", ci=0, height=3, aspect=.9, palette="muted", scatter_kws={"s":10, "alpha":0.5}, line_kws={"lw":2, "alpha":0.5})
# plot's title
plt.suptitle("Figure 12. Amont of tip per person in relation to total bill.", x=0.5, y=1.03, fontsize=14)
# setting axes limits
axes = lm.axes
axes[0,0].set_ylim(0,11)
axes[0,0].set_xlim(0,55)
plt.show()
```
#### Tip percent to total bill relationship
An interesting results is revealed by plotting data points of the percent of tip in relation to total bill. There seems to be a clear negative correlation between the two variables. From the below plot (Fig. 13) one can see a tendency to lower the tip percent (in relation to the total bill) as the bill goes up. The fact is also highlighted by closely fitting linear regression line, as shown in figure 13, below.
```
# Display setting
fig, ax = plt.subplots(figsize=(16,3))
plt.suptitle("Figure 13. correlation between percent of tip to total bill.", x=0.5, y=1.01, fontsize=14)
plt.subplots_adjust(left=None, bottom=None, right=None, top=None, wspace=0.1, hspace=None)
sns.regplot(data=tips, x="total_bill", y="percent", ci=0, scatter_kws={"s": 6, "alpha":0.6}, line_kws={"lw":1, "color":"r", "alpha":0.6}) # confidence interval set to 0 for clarity
plt.show()
```
___
### 3.2 k-nearest neighbours algorithm
This section is based on the Programming for Data Analysis, GMIT, lecture videos and the lecturer's [Notebook](https://github.com/ianmcloughlin/jupyter-teaching-notebooks/raw/master/pandas-with-iris.ipynb).
_k-nearest neighbours_ is a supervised machine learning algorithms. It is used to solve a classification problem. It produces a discreet output (that is: either this or that class (there may be more classes), but not something in between).
Other consulted references:
[Machine Learning Basics with the K-Nearest Neighbors Algorithm](https://towardsdatascience.com/machine-learning-basics-with-the-k-nearest-neighbors-algorithm-6a6e71d01761)
[K-Nearest Neighbors Algorithm Using Python](https://www.edureka.co/blog/k-nearest-neighbors-algorithm/)
[K-Nearest Neighbors Algorithm in Python and Scikit-Learn](https://stackabuse.com/k-nearest-neighbors-algorithm-in-python-and-scikit-learn/)
#### Importing SciKit Learn Library
```
import sklearn.neighbors as nei
import sklearn.model_selection as mod
```
The variables tip, total bill and the smoker are deemed the most suitable for my analysis. The other variables combinations produce more fuzzy plots (a lot of
overlapping data points).
For the purpose of this exercise, I am going to work on a simplified dataset. This will be done by coping the tips dataset I was working above, but getting rid of redundant columns.
```
# A new, reduced dataset
tips2 = tips[["total_bill", "tip", "smoker"]]
tips2.head(2)
```
The below plot shows relationship between the variables.
```
# Display setting
sns.pairplot(tips2, kind="reg", hue="smoker", height=4, aspect=1.5, plot_kws={"line_kws":{"lw":1, "alpha":0.6}, "scatter_kws":{"s": 6, "alpha":0.5}})
plt.suptitle("Figure 14. relationship between the variables.", x=0.5, y=1.02, fontsize=14)
plt.show()
```
Assigning `Inputs` and `Outputs`
```
inputs = tips2[['total_bill', 'tip']]
outputs = tips2['smoker']
```
Classifier `knn`
```
knn = nei.KNeighborsClassifier(n_neighbors=5) # will consider 5 nearest neighbours
```
Applying `fit` function
```
knn.fit(inputs, outputs)
```
Evaluation of the results
```
(knn.predict(inputs) == outputs).sum() # Returns number of correctly recognised samples; total number of samples is 244
```
Training and testing data sub-sets.
The dataset is split randomly into:
1) training (75% of entire dataset size, i.e. 183), and
2) testing (25%, i.e. 61)
```
inputs_train, inputs_test, outputs_train, outputs_test = mod.train_test_split(inputs, outputs, test_size=0.25)
knn = nei.KNeighborsClassifier(n_neighbors=5)
knn.fit(inputs_train, outputs_train)
# knn.predict(inputs_test) == outputs_test
answer = (knn.predict(inputs_test) == outputs_test).sum()
answer
```
Hence, the accuracy of the algorithm is the ratio of correctly recognised to actual number, and is shown below.
```
print("%.02f" % ((answer/61) * 100), "%")
```
___
## Summary
As a conclusion of the above analysis, I infer the main findings as follows.
- On average the tips are 13.64% of the bill.
- There is a strong linear relationship between the amount of tip and the total bill for the entire dataset - the larger the bill, the larger the tip.
- There is a tendency to lower the tip-to-bill ratio as the bill goes up.
- As the party size increases, so does the amount of tip. However, the larger the party, the tip per person is lower.
- The smoking tippers are fewer, but more generous in terms of total bill and amount of tip.
___
## References
General, high-level, reference sources are listed in the README.md file. Here are the references to the problems related to the dataset analysis. Additional references to specific problems where also provided in the body of the Notebook.
### Project and the data set related
- GMIT _Fundamentals of Data Analysis_ module materials on Moodle platform (access may be restricted for staff and students): <https://learnonline.gmit.ie/course/view.php?id=689>
- Raw dataset obtained from: <http://vincentarelbundock.github.io/Rdatasets/datasets.html>
### Python modules and packages
- [Pandas tutorial](http://www.datasciencemadesimple.com/head-and-tail-in-python-pandas/)
- [Seaborn tutorial - Visualizing linear relationships](https://seaborn.pydata.org/tutorial/regression.html)
### Visualisation
- [Data Camp - Python Seaborn Tutorial For Beginners](https://www.datacamp.com/community/tutorials/seaborn-python-tutorial)
- [Data visualisation with Python](https://medium.com/@neuralnets/statistical-data-visualization-series-with-python-and-seaborn-for-data-science-5a73b128851d)
- [AJ Pryor blog - Seaborn Implot](http://alanpryorjr.com/visualizations/seaborn/lmplot/lmplot/)
- [Python graph gallery](https://python-graph-gallery.com/)
- [A collection of dataviz caveats by data-to-viz.com](https://www.data-to-viz.com/caveats.html)
- [Data Visualization with Seaborn](https://jovianlin.io/data-visualization-seaborn-part-1/)
- [Drawing from data](https://www.drawingfromdata.com/)
### Linear regression
- [Wikipedia - Linear regression](https://en.wikipedia.org/wiki/Linear_regression)
- [Simple and multiple linear regression with Python](https://towardsdatascience.com/simple-and-multiple-linear-regression-with-python-c9ab422ec29c)
- [Charting Tips](http://rstudio-pubs-static.s3.amazonaws.com/128623_507fbe51532748f29278825fceedab85.html)
- [Create basic graph visializations](https://mlwhiz.com/blog/2015/09/13/seaborn_visualizations/)
- [What is Regression Analysis and Why Should I Use It?](https://www.surveygizmo.com/resources/blog/regression-analysis/)
- [Explained: Regression analysis](http://news.mit.edu/2010/explained-reg-analysis-0316)
- [ML | Linear Regression](https://www.geeksforgeeks.org/ml-linear-regression/)
- [ML Glossary - Linear Regression](https://ml-cheatsheet.readthedocs.io/en/latest/linear_regression.html)
___
Andrzej Kocielski, 2019
| github_jupyter |
```
# !conda install -y -c conda-forge fbprophet
import boto3
import base64
from botocore.exceptions import ClientError
from IPython.display import display
from fbprophet import Prophet
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_absolute_percentage_error as MAPE
from sklearn.metrics import mean_absolute_error as MAE
from sklearn.metrics import mean_squared_error as MSE
from sklearn.preprocessing import normalize
tickers = ['AAPL']
metric = 'low'
pc_metric = f'{metric}_percent_change'
norm_metric = f'{pc_metric}_norm'
def get_secret():
secret_name = "alpha_vantage"
region_name = "us-east-2"
# Create a Secrets Manager client
session = boto3.session.Session()
client = session.client(
service_name='secretsmanager',
region_name=region_name
)
try:
get_secret_value_response = client.get_secret_value(SecretId=secret_name)
except ClientError as e:
display(e)
else:
# Decrypts secret using the associated KMS CMK.
# Depending on whether the secret is a string or binary, one of these fields will be populated.
if 'SecretString' in get_secret_value_response:
secret = get_secret_value_response['SecretString']
else:
secret = base64.b64decode(get_secret_value_response['SecretBinary'])
return secret
def format_dates(daily_stocks_data):
df = daily_stocks_data.copy()
df['date']=df.index
df.reset_index(inplace=True, drop=True)
return df
def add_percent_change(daily_stocks_data, metric):
percents = list()
for index, row in daily_stocks_data.iterrows():
old = row[metric]
try:
new = daily_stocks_data.iloc[index + 1][metric]
except Exception as e:
percents.append(np.nan) ## no next value, so this is undefined
continue
percents.append((new-old)/new)
cp_df = daily_stocks_data.copy()
cp_df[f'{metric}_percent_change']=percents
return cp_df
def add_norm(df, label):
arr = np.array([x*1000 for x in df[label].to_numpy()]).reshape(-1, 1)
# norm = normalize(arr, norm='l1')
norm = arr
new_df = df.copy()
new_df[f'{label}_norm'] = norm
return new_df
ALPHA_API_KEY = get_secret()
# daily_stocks_data_raw = pandas_datareader.av.time_series.AVTimeSeriesReader(symbols=tickers, api_key=ALPHA_API_KEY, function='TIME_SERIES_DAILY').read()
daily_stocks_data = format_dates(daily_stocks_data_raw)
daily_stocks_data = add_percent_change(daily_stocks_data, metric)
daily_stocks_data[daily_stocks_data[pc_metric].isnull()] = 0
daily_stocks_data = add_norm(daily_stocks_data, pc_metric)
display(daily_stocks_data)
# daily_stocks_data.isin([-1, 1, 0]).all()
prophet_df = pd.DataFrame({'ds':daily_stocks_data['date'].to_numpy(), 'y':daily_stocks_data[norm_metric].to_numpy()})
display(prophet_df)
prophet_df_train, prophet_df_test = train_test_split(prophet_df)
display(prophet_df_train, prophet_df_test)
model=Prophet(interval_width=0.95, daily_seasonality=True)
model.fit(prophet_df_train)
train_pred = model.make_future_dataframe(periods=100)
display(train_pred)
forecast = model.predict(train_pred)
display(forecast)
model.plot(forecast)
test_forecast = model.predict(prophet_df_test)
display(test_forecast)
model.plot(test_forecast)
display('predictions MAE', MAE(test_forecast['yhat'], prophet_df_test['y']))
display('MSE',MSE(test_forecast['yhat'], prophet_df_test['y']))
display('baseline MAE',MAE([0]*len(prophet_df_test['y']), prophet_df_test['y']))
display('MSE',MSE([0]*len(prophet_df_test['y']), prophet_df_test['y']))
display('mean baseline MAE',MAE([prophet_df_test['y'].mean()]*len(prophet_df_test['y']), prophet_df_test['y']))
display('MSE',MSE([prophet_df_test['y'].mean()]*len(prophet_df_test['y']), prophet_df_test['y']))
display('mean', test_forecast['yhat'].mean())
display('std', test_forecast['yhat'].std())
display('mean', prophet_df_test['y'].mean())
display('std', prophet_df_test['y'].std())
## worse than no change prediction or mean prediction on percentage change metric; next method.
```
| github_jupyter |
# Lecture 10 – More Review
## Data 94, Spring 2021
## Stock prices
### Part 1
**Task:** Given a list of prices of a stock on a single day, return a string describing whether the stock increased, stayed the same, or decreased from its starting price.
```
def daily_change(prices):
first = prices[0]
last = prices[-1]
if first > last:
return 'decrease'
elif first == last:
return 'none'
else:
return 'increase'
# Returns 'increase', since 1 < 4
daily_change([1, 2, 2.5, 3, 3.5, 4])
# Returns 'none', since 3 = 3
daily_change([3, 9, 3])
# Return 'decrease', since 5 > 2
daily_change([5, 4, 3, 4, 5, 4, 3, 2, 2])
```
### Part 2, Quick Check 1
**Task:** Given a list of prices of a stock on a single day, return `True` if the stock was strictly increasing, and `False` otherwise. A list of numbers is strictly increasing if each one is larger than the previous.
```
def strictly_increasing(prices):
i = 0
while i < len(prices) - 1:
if ...:
return False
i += 1
return True
# True
# strictly_increasing([1, 2, 5, 8, 10])
# False
# strictly_increasing([2, 3, 9, 7, 11])
# False
# strictly_increasing([2, 3, 4, 4, 5])
```
## Next day of the week
```
def next_day(day):
week = ['Sunday', 'Monday', 'Tuesday', 'Wednesday', 'Thursday', 'Friday', 'Saturday']
curr = week.index(day)
return week[(curr + 1) % 7]
next_day('Wednesday')
next_day('Saturday')
```
## Prefixes, Quick Check 2
```
def full_prefix(name):
# i and idx are short for "index"
idx = name.index('.')
prefix = name[:idx]
rest = ...
if prefix == 'Dr':
return 'Doctor ' + rest
elif prefix == 'Prof':
return 'Professor ' + rest
elif prefix == 'Gov':
return 'Governor ' + rest
else:
return name
# 'Governor Newsom'
# full_prefix('Gov. Newsom')
# 'Professor Christ'
# full_prefix('Prof. Christ')
# 'Doctor Biden'
# full_prefix('Dr. Biden')
# 'Hon. Caboto'
# full_prefix('Hon. Caboto')
```
## Kaprekar's constant
```
# For now, ignore the code in this cell.
def increase_sort(n):
n_list = list(str(n))
n_list_sorted = sorted(n_list)
n_str_sorted = ''.join(n_list_sorted)
return int(n_str_sorted)
def decrease_sort(n):
n_list = list(str(n))
n_list_sorted = sorted(n_list)[::-1]
n_str_sorted = ''.join(n_list_sorted)
return int(n_str_sorted)
def find_sequence(n):
# Need to keep track of the steps, and the "current" number in our sequence
steps = [n]
curr = n
# As long as our current number isn't 495
# make one more step, and store the current step
while curr != 495:
curr = decrease_sort(curr) - increase_sort(curr)
steps.append(curr)
return steps
find_sequence(813)
find_sequence(215)
```
| github_jupyter |
# Using the SimulatorIMNN
With a generative model built in ``jax`` we can generate simulations on-the-fly to fit an IMNN. This is the best way to prevent against overfitting due to limited variability in datasets which may introduce spurious features which are not informative about the model parameters.
For this example we are going to summaries the unknown mean, $\mu$, and variance, $\Sigma$, of $n_{\bf d}=10$ data points of two 1D random Gaussian field, ${\bf d}=\{d_i\sim\mathcal{N}(\mu,\Sigma)|i\in[1, n_{\bf d}]\}$. This is an interesting problem since we know the likelihood analytically, but it is non-Gaussian
$$\mathcal{L}({\bf d}|\mu,\Sigma) = \prod_i^{n_{\bf d}}\frac{1}{\sqrt{2\pi|\Sigma|}}\exp\left[-\frac{1}{2}\frac{(d_i-\mu)^2}{\Sigma}\right]$$
As well as knowing the likelihood for this problem, we also know what sufficient statistics describe the mean and variance of the data - they are the mean and the variance
$$\frac{1}{n_{\bf d}}\sum_i^{n_{\bf d}}d_i = \mu\textrm{ and }\frac{1}{n_{\bf d}-1}\sum_i^{n_{\bf d}}(d_i-\mu)^2=\Sigma$$
What makes this an interesting problem for the IMNN is the fact that the sufficient statistic for the variance is non-linear, i.e. it is a sum of the square of the data, and so linear methods like MOPED would be lossy in terms of information.
We can calculate the Fisher information by taking the negative second derivative of the likelihood taking the expectation by inserting the relations for the sufficient statistics, i.e. and examining at the fiducial parameter values
$${\bf F}_{\alpha\beta} = -\left.\left(\begin{array}{cc}\displaystyle-\frac{n_{\bf d}}{\Sigma}&0\\0&\displaystyle-\frac{n_{\bf d}}{2\Sigma^2}\end{array}\right)\right|_{\Sigma=\Sigma^{\textrm{fid}}}.$$
Choosing fiducial parameter values of $\mu^\textrm{fid}=0$ and $\Sigma^\textrm{fid}=1$ we find that the determinant of the Fisher information matrix is $|{\bf F}_{\alpha\beta}|=50$.
```
from imnn import SimulatorIMNN
import jax
import jax.numpy as np
from jax.experimental import stax, optimizers
```
We're going to use 1000 summary vectors, with a length of two, at a time to make an estimate of the covariance of network outputs and the derivative of the mean of the network outputs with respect to the two model parameters.
```
n_s = 1000
n_d = n_s
n_params = 2
n_summaries = n_params
input_shape = (10,)
```
These are going to be generated on the fly using:
```
def simulator(key, θ):
return θ[0] + jax.random.normal(key, shape=input_shape) * np.sqrt(θ[1])
```
Our fiducial parameter values are $\mu^\textrm{fid}=0$ and $\Sigma^\textrm{fid}=1$
```
θ_fid = np.array([0., 1.])
```
For initialising the neural network and for generating the simulations on the fly we need two random number generators:
```
rng = jax.random.PRNGKey(0)
rng, model_key, fit_key = jax.random.split(rng, num=3)
```
We're going to use ``jax``'s stax module to build a simple network with three hidden layers each with 128 neurons and which are activated by leaky relu before outputting the two summaries. The optimiser will be a ``jax`` Adam optimiser with a step size of 0.001.
```
model = stax.serial(
stax.Dense(128),
stax.LeakyRelu,
stax.Dense(128),
stax.LeakyRelu,
stax.Dense(128),
stax.LeakyRelu,
stax.Dense(n_summaries))
optimiser = optimizers.adam(step_size=1e-3)
```
The SimulatorIMNN can now be initialised setting up the network and the fitting routine (as well as the plotting function)
```
imnn = SimulatorIMNN(
n_s=n_s, n_d=n_d, n_params=n_params, n_summaries=n_summaries,
input_shape=input_shape, θ_fid=θ_fid, model=model,
optimiser=optimiser, key_or_state=model_key,
simulator=simulator)
```
To set the scale of the regularisation we use a coupling strength $\lambda$ whose value should mean that the determinant of the difference between the covariance of network outputs and the identity matrix is larger than the expected initial value of the determinant of the Fisher information matrix from the network. How close to the identity matrix the covariance should be is set by $\epsilon$. These parameters should not be very important, but they will help with convergence time.
```
λ = 10.
ϵ = 0.1
```
Fitting can then be done simply by calling:
```
imnn.fit(λ, ϵ, rng=fit_key, print_rate=1, best=False)
```
Here we have included a ``print_rate`` for a progress bar, but leaving this out will massively reduce fitting time (at the expense of not knowing how many iterations have been run). The IMNN is run (by default) for a maximum of ``max_iterations = 100000`` iterations, but with early stopping which can turn on after ``min_iterations = 100`` iterations and after ``patience = 100`` iterations where the maximum determinant of the Fisher information matrix has not increased. Note we have chosen to set ``imnn.w`` are set to the values of the network parameters at the final iteration via ``best = False``, otherwise the network parameters which obtained the highest value of the determinant of the Fisher information matrix would be set.
To continue training one can simply generate a new random seed (very important) and rerun fit
```
rng, another_fit_key = jax.random.split(rng)
imnn.fit(λ, ϵ, rng=another_fit_key, print_rate=1, best=False)
```
To visualise the fitting history we can plot the results:
```
imnn.plot(expected_detF=50);
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import torch
import torchvision
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from matplotlib import pyplot as plt
%matplotlib inline
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
trainset = torchvision.datasets.CIFAR10(root='./data', train=True, download=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=10, shuffle=True)
classes = ('plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
foreground_classes = {'plane', 'car', 'bird'}
#foreground_classes = {'bird', 'cat', 'deer'}
background_classes = {'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck'}
#background_classes = {'plane', 'car', 'dog', 'frog', 'horse','ship', 'truck'}
fg1,fg2,fg3 = 0,1,2
dataiter = iter(trainloader)
background_data=[]
background_label=[]
foreground_data=[]
foreground_label=[]
batch_size=10
for i in range(5000):
images, labels = dataiter.next()
for j in range(batch_size):
if(classes[labels[j]] in background_classes):
img = images[j].tolist()
background_data.append(img)
background_label.append(labels[j])
else:
img = images[j].tolist()
foreground_data.append(img)
foreground_label.append(labels[j])
foreground_data = torch.tensor(foreground_data)
foreground_label = torch.tensor(foreground_label)
background_data = torch.tensor(background_data)
background_label = torch.tensor(background_label)
def create_mosaic_img(bg_idx,fg_idx,fg):
"""
bg_idx : list of indexes of background_data[] to be used as background images in mosaic
fg_idx : index of image to be used as foreground image from foreground data
fg : at what position/index foreground image has to be stored out of 0-8
"""
image_list=[]
j=0
for i in range(9):
if i != fg:
image_list.append(background_data[bg_idx[j]])#.type("torch.DoubleTensor"))
j+=1
else:
image_list.append(foreground_data[fg_idx])#.type("torch.DoubleTensor"))
label = foreground_label[fg_idx]-fg1 # minus 7 because our fore ground classes are 7,8,9 but we have to store it as 0,1,2
#image_list = np.concatenate(image_list ,axis=0)
image_list = torch.stack(image_list)
return image_list,label
desired_num = 30000
mosaic_list_of_images =[] # list of mosaic images, each mosaic image is saved as list of 9 images
fore_idx =[] # list of indexes at which foreground image is present in a mosaic image i.e from 0 to 9
mosaic_label=[] # label of mosaic image = foreground class present in that mosaic
for i in range(desired_num):
np.random.seed(i)
bg_idx = np.random.randint(0,35000,8)
fg_idx = np.random.randint(0,15000)
fg = np.random.randint(0,9)
fore_idx.append(fg)
image_list,label = create_mosaic_img(bg_idx,fg_idx,fg)
mosaic_list_of_images.append(image_list)
mosaic_label.append(label)
class MosaicDataset(Dataset):
"""MosaicDataset dataset."""
def __init__(self, mosaic_list_of_images, mosaic_label, fore_idx):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.mosaic = mosaic_list_of_images
self.label = mosaic_label
self.fore_idx = fore_idx
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.mosaic[idx] , self.label[idx], self.fore_idx[idx]
batch = 250
msd = MosaicDataset(mosaic_list_of_images, mosaic_label , fore_idx)
train_loader = DataLoader( msd,batch_size= batch ,shuffle=False)
data,labels,fg_index = iter(train_loader).next()
ag = []
for i in range(120):
alphag = torch.ones((250,9))/9
ag.append( alphag.requires_grad_() )
class Module2(nn.Module):
def __init__(self):
super(Module2, self).__init__()
self.conv1 = nn.Conv2d(3, 6, 5)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(6, 16, 5)
self.fc1 = nn.Linear(16 * 5 * 5, 120)
self.fc2 = nn.Linear(120, 84)
self.fc3 = nn.Linear(84, 10)
self.fc4 = nn.Linear(10,3)
def forward(self,y): #z batch of list of 9 images
y1 = self.pool(F.relu(self.conv1(y)))
y1 = self.pool(F.relu(self.conv2(y1)))
y1 = y1.view(-1, 16 * 5 * 5)
y1 = F.relu(self.fc1(y1))
y1 = F.relu(self.fc2(y1))
y1 = F.relu(self.fc3(y1))
y1 = self.fc4(y1)
return y1
torch.manual_seed(1234)
what_net = Module2().double()
what_net = what_net.to("cuda")
def attn_avg(x,alpha):
y = torch.zeros([batch,3, 32,32], dtype=torch.float64)
y = y.to("cuda")
alpha = F.softmax(alpha,dim=1) # alphas
for i in range(9):
alpha1 = alpha[:,i]
y = y + torch.mul(alpha1[:,None,None,None],x[:,i])
return y,alpha
def calculate_attn_loss(dataloader,what,criter):
what.eval()
r_loss = 0
alphas = []
lbls = []
pred = []
fidices = []
correct = 0
tot = 0
with torch.no_grad():
for i, data in enumerate(dataloader, 0):
inputs, labels,fidx= data
lbls.append(labels)
fidices.append(fidx)
inputs = inputs.double()
alpha = ag[i] # alpha for ith batch
inputs, labels,alpha = inputs.to("cuda"),labels.to("cuda"),alpha.to("cuda")
avg,alpha = attn_avg(inputs,alpha)
alpha = alpha.to("cuda")
outputs = what(avg)
_, predicted = torch.max(outputs.data, 1)
correct += sum(predicted == labels)
tot += len(predicted)
pred.append(predicted.cpu().numpy())
alphas.append(alpha.cpu().numpy())
loss = criter(outputs, labels)
r_loss += loss.item()
alphas = np.concatenate(alphas,axis=0)
pred = np.concatenate(pred,axis=0)
lbls = np.concatenate(lbls,axis=0)
fidices = np.concatenate(fidices,axis=0)
#print(alphas.shape,pred.shape,lbls.shape,fidices.shape)
analysis = analyse_data(alphas,lbls,pred,fidices)
return r_loss/i,analysis,correct.item(),tot,correct.item()/tot
def analyse_data(alphas,lbls,predicted,f_idx):
'''
analysis data is created here
'''
batch = len(predicted)
amth,alth,ftpt,ffpt,ftpf,ffpf = 0,0,0,0,0,0
for j in range (batch):
focus = np.argmax(alphas[j])
if(alphas[j][focus] >= 0.5):
amth +=1
else:
alth +=1
if(focus == f_idx[j] and predicted[j] == lbls[j]):
ftpt += 1
elif(focus != f_idx[j] and predicted[j] == lbls[j]):
ffpt +=1
elif(focus == f_idx[j] and predicted[j] != lbls[j]):
ftpf +=1
elif(focus != f_idx[j] and predicted[j] != lbls[j]):
ffpf +=1
#print(sum(predicted==lbls),ftpt+ffpt)
return [ftpt,ffpt,ftpf,ffpf,amth,alth]
optim1 = []
for i in range(120):
optim1.append(optim.RMSprop([ag[i]], lr=10))
# instantiate optimizer
optimizer_what = optim.RMSprop(what_net.parameters(), lr=0.001)#, momentum=0.9)#,nesterov=True)
criterion = nn.CrossEntropyLoss()
acti = []
analysis_data_tr = []
analysis_data_tst = []
loss_curi_tr = []
loss_curi_tst = []
epochs = 200
# calculate zeroth epoch loss and FTPT values
running_loss,anlys_data,correct,total,accuracy = calculate_attn_loss(train_loader,what_net,criterion)
print('training epoch: [%d ] loss: %.3f correct: %.3f, total: %.3f, accuracy: %.3f' %(0,running_loss,correct,total,accuracy))
loss_curi_tr.append(running_loss)
analysis_data_tr.append(anlys_data)
# training starts
for epoch in range(epochs): # loop over the dataset multiple times
ep_lossi = []
running_loss = 0.0
what_net.train()
for i, data in enumerate(train_loader, 0):
# get the inputs
grads = []
inputs, labels,_ = data
inputs = inputs.double()
alpha = ag[i] # alpha for ith batch
inputs, labels,alpha = inputs.to("cuda"),labels.to("cuda"),alpha.to("cuda")
# zero the parameter gradients
optimizer_what.zero_grad()
optim1[i].zero_grad()
# forward + backward + optimize
avg,alpha = attn_avg(inputs,alpha)
outputs = what_net(avg)
loss = criterion(outputs, labels)
# print statistics
running_loss += loss.item()
alpha.retain_grad()
loss.backward(retain_graph=False)
optimizer_what.step()
optim1[i].step()
running_loss_tr,anls_data,correct,total,accuracy = calculate_attn_loss(train_loader,what_net,criterion)
analysis_data_tr.append(anls_data)
loss_curi_tr.append(running_loss_tr) #loss per epoch
print('training epoch: [%d ] loss: %.3f correct: %.3f, total: %.3f, accuracy: %.3f' %(epoch+1,running_loss_tr,correct,total,accuracy))
if running_loss_tr<=0.08:
break
print('Finished Training run ')
analysis_data_tr = np.array(analysis_data_tr)
columns = ["epochs", "argmax > 0.5" ,"argmax < 0.5", "focus_true_pred_true", "focus_false_pred_true", "focus_true_pred_false", "focus_false_pred_false" ]
df_train = pd.DataFrame()
df_test = pd.DataFrame()
df_train[columns[0]] = np.arange(0,epoch+2)
df_train[columns[1]] = analysis_data_tr[:,-2]
df_train[columns[2]] = analysis_data_tr[:,-1]
df_train[columns[3]] = analysis_data_tr[:,0]
df_train[columns[4]] = analysis_data_tr[:,1]
df_train[columns[5]] = analysis_data_tr[:,2]
df_train[columns[6]] = analysis_data_tr[:,3]
df_train
fig= plt.figure(figsize=(12,12))
plt.plot(df_train[columns[0]],df_train[columns[3]]/300, label ="focus_true_pred_true ")
plt.plot(df_train[columns[0]],df_train[columns[4]]/300, label ="focus_false_pred_true ")
plt.plot(df_train[columns[0]],df_train[columns[5]]/300, label ="focus_true_pred_false ")
plt.plot(df_train[columns[0]],df_train[columns[6]]/300, label ="focus_false_pred_false ")
plt.title("On Train set")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("percentage of data")
#plt.vlines(vline_list,min(min(df_train[columns[3]]/300),min(df_train[columns[4]]/300),min(df_train[columns[5]]/300),min(df_train[columns[6]]/300)), max(max(df_train[columns[3]]/300),max(df_train[columns[4]]/300),max(df_train[columns[5]]/300),max(df_train[columns[6]]/300)),linestyles='dotted')
plt.show()
fig.savefig("train_analysis_every_20.pdf")
fig.savefig("train_analysis_every_20.png")
aph = []
for i in ag:
aph.append(F.softmax(i,dim=1).detach().numpy())
aph = np.concatenate(aph,axis=0)
torch.save({
'epoch': 500,
'model_state_dict': what_net.state_dict(),
'optimizer_state_dict': optimizer_what.state_dict(),
"optimizer_alpha":optim1,
"FTPT_analysis":analysis_data_tr,
"alpha":aph
}, "cifar_what_net_500.pt")
aph
running_loss_tr,anls_data,correct,total,accuracy = calculate_attn_loss(train_loader,what_net,criterion)
anls_data
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sqlalchemy import create_engine
plt.style.use('default')
# assumes .env file is in working directory
db_url = ! source .env; echo ${DATABASE_URL}
db_url = db_url[0]
# Create an engine instance
alchemyEngine = create_engine(db_url)
# Connect to PostgreSQL server
dbConnection = alchemyEngine.connect()
# Read data from PostgreSQL database table and load into a DataFrame instance
bci_collection = pd.read_sql("SELECT * FROM bci_collection", dbConnection)
# Print the DataFrame
print(bci_collection)
# # this counts the number of rows for each subject
# # takes a while to run
# for name in bci_collection['collector_name'].drop_duplicates():
# collection_ids = bci_collection.loc[bci_collection['collector_name'] == name, 'id']
# collection_ids_nonempty = []
# total_count = 0
# for id in collection_ids:
# count = pd.read_sql(
# f"SELECT COUNT(*) FROM collected_data WHERE collection_id = {id}",
# dbConnection,
# ).iloc[0,0]
# if count > 0:
# collection_ids_nonempty.append(id)
# total_count += count
# if len(collection_ids_nonempty) > 0:
# print(f'{name} ({total_count}):\t{collection_ids_nonempty}')
subject_ids = ['S09_trial1', 's09_trial2', 's09_trial3']
collection_ids = bci_collection.loc[bci_collection['collector_name'].isin(subject_ids), 'id'].drop_duplicates()
print(collection_ids)
# takes up to ~30 seconds to run
subject_data = []
# collection_ids = ['43']
for collection_id in collection_ids:
df = pd.read_sql(
f"SELECT * FROM collected_data WHERE collection_id = {collection_id}",
dbConnection
)
n_rows_per_character = df['character'].value_counts()
print(f'Total number of rows: {df.shape[0]}')
print(f'{len(n_rows_per_character)} characters for collection ID {collection_id}')
print(f'Average number of rows per character: {n_rows_per_character.mean():.0f}')
print(f'Frequency range: {df["frequency"].min()}-{df["frequency"].max()} Hz')
print('-----')
subject_data.append(df)
i_session = 0
channel_name = 'channel_5'
character = 'z'
fs = 250
session_data = subject_data[i_session]
n_sample_per_character = session_data.groupby('character').size()
print(n_sample_per_character.value_counts())
fig, ax = plt.subplots(figsize=(10,3))
trial_data = session_data.loc[session_data['character'] == character].sort_values('order')
ax.plot(np.arange(len(trial_data))/fs, trial_data[channel_name])
```
| github_jupyter |
```
import os
import pandas as pd
import numpy as np
import pandas as pd
from pathlib import Path
import sklearn
import sklearn.metrics
import sklearn.model_selection
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
# Explore ResNet feature matrices
image_folder = "train_input/resnet_features/"
#image = np.load('/tmp/123.npy', mmap_mode='r')
# Function to load folder into arrays and then it returns that same array
def loadImages(path):
image_files = sorted([os.path.join(path, file)
for file in os.listdir(path) if file.endswith('.npy')])
return image_files
# Check number of images in training set
len(loadImages(image_folder))
# load first npy and explore data
npy_train = loadImages(image_folder)
matrix_npy = np.load(npy_train[0], mmap_mode='r')
def get_average_features(filenames):
"""Load and aggregate the resnet features by the average.
Args:
filenames: list of filenames of length `num_patients` corresponding to resnet features
Returns:
features: np.array of mean resnet features, shape `(num_patients, 2048)`
"""
# Load numpy arrays
features = []
for f in filenames:
patient_features = np.load(f)
# Remove location features (but we could use them?)
patient_features = patient_features[:, 3:]
aggregated_features = np.mean(patient_features, axis=0)
features.append(aggregated_features)
features = np.stack(features, axis=0)
return features
# load feature npy folder into arrays and then it returns that same array of strings
def loadFiles(path):
feature_files = sorted([os.path.join(path, file)
for file in os.listdir(path) if file.endswith('.npy')])
return feature_files
# precise training set and test set relative location
train_dir = Path("train_input/resnet_features")
test_dir = Path("test_input/resnet_features")
train_output_filename = Path("training_output.csv")
train_output = pd.read_csv(train_output_filename)
# Get filenames for train
filenames_train = loadFiles(train_dir)
# Get global labels (patient-wise) for train
labels_train = train_output["Target"].values
# check if the number of observations and labels corresponds
assert len(filenames_train) == len(labels_train)
# Get the numpy filenames for test
filenames_test = loadFiles(test_dir)
# ID list without its suffix (ex: "ID_005")
ids_test = [Path(f).stem for f in filenames_test]
# Get the resnet features and aggregate them by the average
features_train = get_average_features(filenames_train)
features_test = get_average_features(filenames_test)
# Use the average resnet features to predict the labels
# number of runs for cross-validation
num_runs = 5
# number of splits for cross-validation
num_splits = 20
# Multiple cross validations on the training set
aucs = []
for seed in range(num_runs):
# Use linear Discriminant Analysis
model = LinearDiscriminantAnalysis()
cv = sklearn.model_selection.RepeatedStratifiedKFold(n_splits=num_splits, n_repeats=num_runs, random_state=seed)
# Cross validation on the training set
auc = sklearn.model_selection.cross_val_score(model, X=features_train, y=labels_train,
cv=cv, scoring="roc_auc", verbose=0)
aucs.append(auc)
aucs = np.array(aucs)
print("Predicting weak labels by mean resnet")
print("AUC: mean {}, std {}".format(aucs.mean(), aucs.std()))
# Prediction on the test set
# Train a final model on the full training set
estimator = LinearDiscriminantAnalysis()
estimator.fit(features_train, labels_train)
preds_test = estimator.predict_proba(features_test)[:, 1]
# Check that predictions are in [0, 1]
assert np.max(preds_test) <= 1.0
assert np.min(preds_test) >= 0.0
# -------------------------------------------------------------------------
# Write the predictions in a csv file, to export them in the suitable format
# to the data challenge platform
ids_number_test = [i.split("ID_")[1] for i in ids_test]
test_output = pd.DataFrame({"ID": ids_number_test, "Target": preds_test})
test_output.set_index("ID", inplace=True)
test_output.to_csv("predictions/preds_lda.csv")
```
<h2>Tune LDA hyperparameters with grid search</h2>
```
# Use the average resnet features to predict the labels
# number of runs for cross-validation
num_runs = 5
# number of splits for cross-validation
num_splits = 20
# Multiple cross validations on the training set
# Use linear Discriminant Analysis
model = LinearDiscriminantAnalysis()
cv = sklearn.model_selection.RepeatedStratifiedKFold(n_splits=num_splits, n_repeats=num_runs, random_state=1)
# define grid
grid = dict()
grid['solver'] = ['svd', 'lsqr', 'eigen']
# define search
search = sklearn.model_selection.GridSearchCV(model, grid, scoring='roc_auc', cv=cv, n_jobs=-1)
# perform the search
results = search.fit(X=features_train, y=labels_train)
# summarize
print('Mean Accuracy: %.3f' % results.best_score_)
print('Config: %s' % results.best_params_)
```
<p>Finally the default solver gives best result over all solvers so no need to do another prediction.</p>
| github_jupyter |
# 批量规范化
:label:`sec_batch_norm`
训练深层神经网络是十分困难的,特别是在较短的时间内使他们收敛更加棘手。
在本节中,我们将介绍*批量规范化*(batch normalization) :cite:`Ioffe.Szegedy.2015`,这是一种流行且有效的技术,可持续加速深层网络的收敛速度。
再结合在 :numref:`sec_resnet`中将介绍的残差块,批量规范化使得研究人员能够训练100层以上的网络。
## 训练深层网络
为什么需要批量规范化层呢?让我们来回顾一下训练神经网络时出现的一些实际挑战。
首先,数据预处理的方式通常会对最终结果产生巨大影响。
回想一下我们应用多层感知机来预测房价的例子( :numref:`sec_kaggle_house`)。
使用真实数据时,我们的第一步是标准化输入特征,使其平均值为0,方差为1。
直观地说,这种标准化可以很好地与我们的优化器配合使用,因为它可以将参数的量级进行统一。
第二,对于典型的多层感知机或卷积神经网络。当我们训练时,中间层中的变量(例如,多层感知机中的仿射变换输出)可能具有更广的变化范围:不论是沿着从输入到输出的层,跨同一层中的单元,或是随着时间的推移,模型参数的随着训练更新变幻莫测。
批量规范化的发明者非正式地假设,这些变量分布中的这种偏移可能会阻碍网络的收敛。
直观地说,我们可能会猜想,如果一个层的可变值是另一层的100倍,这可能需要对学习率进行补偿调整。
第三,更深层的网络很复杂,容易过拟合。
这意味着正则化变得更加重要。
批量规范化应用于单个可选层(也可以应用到所有层),其原理如下:在每次训练迭代中,我们首先规范化输入,即通过减去其均值并除以其标准差,其中两者均基于当前小批量处理。
接下来,我们应用比例系数和比例偏移。
正是由于这个基于*批量*统计的*标准化*,才有了*批量规范化*的名称。
请注意,如果我们尝试使用大小为1的小批量应用批量规范化,我们将无法学到任何东西。
这是因为在减去均值之后,每个隐藏单元将为0。
所以,只有使用足够大的小批量,批量规范化这种方法才是有效且稳定的。
请注意,在应用批量规范化时,批量大小的选择可能比没有批量规范化时更重要。
从形式上来说,用$\mathbf{x} \in \mathcal{B}$表示一个来自小批量$\mathcal{B}$的输入,批量规范化$\mathrm{BN}$根据以下表达式转换$\mathbf{x}$:
$$\mathrm{BN}(\mathbf{x}) = \boldsymbol{\gamma} \odot \frac{\mathbf{x} - \hat{\boldsymbol{\mu}}_\mathcal{B}}{\hat{\boldsymbol{\sigma}}_\mathcal{B}} + \boldsymbol{\beta}.$$
:eqlabel:`eq_batchnorm`
在 :eqref:`eq_batchnorm`中,$\hat{\boldsymbol{\mu}}_\mathcal{B}$是样本均值,$\hat{\boldsymbol{\sigma}}_\mathcal{B}$是小批量$\mathcal{B}$的样本标准差。
应用标准化后,生成的小批量的平均值为0和单位方差为1。
由于单位方差(与其他一些魔法数)是一个任意的选择,因此我们通常包含
*拉伸参数*(scale)$\boldsymbol{\gamma}$和*偏移参数*(shift)$\boldsymbol{\beta}$,它们的形状与$\mathbf{x}$相同。
请注意,$\boldsymbol{\gamma}$和$\boldsymbol{\beta}$是需要与其他模型参数一起学习的参数。
由于在训练过程中,中间层的变化幅度不能过于剧烈,而批量规范化将每一层主动居中,并将它们重新调整为给定的平均值和大小(通过$\hat{\boldsymbol{\mu}}_\mathcal{B}$和${\hat{\boldsymbol{\sigma}}_\mathcal{B}}$)。
从形式上来看,我们计算出 :eqref:`eq_batchnorm`中的$\hat{\boldsymbol{\mu}}_\mathcal{B}$和${\hat{\boldsymbol{\sigma}}_\mathcal{B}}$,如下所示:
$$\begin{aligned} \hat{\boldsymbol{\mu}}_\mathcal{B} &= \frac{1}{|\mathcal{B}|} \sum_{\mathbf{x} \in \mathcal{B}} \mathbf{x},\\
\hat{\boldsymbol{\sigma}}_\mathcal{B}^2 &= \frac{1}{|\mathcal{B}|} \sum_{\mathbf{x} \in \mathcal{B}} (\mathbf{x} - \hat{\boldsymbol{\mu}}_{\mathcal{B}})^2 + \epsilon.\end{aligned}$$
请注意,我们在方差估计值中添加一个小常量$\epsilon > 0$,以确保我们永远不会尝试除以零,即使在经验方差估计值可能消失的情况下也是如此。估计值$\hat{\boldsymbol{\mu}}_\mathcal{B}$和${\hat{\boldsymbol{\sigma}}_\mathcal{B}}$通过使用平均值和方差的噪声(noise)估计来抵消缩放问题。
你可能会认为这种噪声是一个问题,而事实上它是有益的。
事实证明,这是深度学习中一个反复出现的主题。
由于理论上尚未明确表述的原因,优化中的各种噪声源通常会导致更快的训练和较少的过拟合:这种变化似乎是正则化的一种形式。
在一些初步研究中, :cite:`Teye.Azizpour.Smith.2018`和 :cite:`Luo.Wang.Shao.ea.2018`分别将批量规范化的性质与贝叶斯先验相关联。
这些理论揭示了为什么批量规范化最适应$50 \sim 100$范围中的中等小批量尺寸的难题。
另外,批量规范化图层在”训练模式“(通过小批量统计数据规范化)和“预测模式”(通过数据集统计规范化)中的功能不同。
在训练过程中,我们无法得知使用整个数据集来估计平均值和方差,所以只能根据每个小批次的平均值和方差不断训练模型。
而在预测模式下,可以根据整个数据集精确计算批量规范化所需的平均值和方差。
现在,我们了解一下批量规范化在实践中是如何工作的。
## 批量规范化层
回想一下,批量规范化和其他图层之间的一个关键区别是,由于批量规范化在完整的小批次上运行,因此我们不能像以前在引入其他图层时那样忽略批处理的尺寸大小。
我们在下面讨论这两种情况:全连接层和卷积层,他们的批量规范化实现略有不同。
### 全连接层
通常,我们将批量规范化层置于全连接层中的仿射变换和激活函数之间。
设全连接层的输入为u,权重参数和偏置参数分别为$\mathbf{W}$和$\mathbf{b}$,激活函数为$\phi$,批量规范化的运算符为$\mathrm{BN}$。
那么,使用批量规范化的全连接层的输出的计算详情如下:
$$\mathbf{h} = \phi(\mathrm{BN}(\mathbf{W}\mathbf{x} + \mathbf{b}) ).$$
回想一下,均值和方差是在应用变换的"相同"小批量上计算的。
### 卷积层
同样,对于卷积层,我们可以在卷积层之后和非线性激活函数之前应用批量规范化。
当卷积有多个输出通道时,我们需要对这些通道的“每个”输出执行批量规范化,每个通道都有自己的拉伸(scale)和偏移(shift)参数,这两个参数都是标量。
假设我们的微批次包含$m$个示例,并且对于每个通道,卷积的输出具有高度$p$和宽度$q$。
那么对于卷积层,我们在每个输出通道的$m \cdot p \cdot q$个元素上同时执行每个批量规范化。
因此,在计算平均值和方差时,我们会收集所有空间位置的值,然后在给定通道内应用相同的均值和方差,以便在每个空间位置对值进行规范化。
### 预测过程中的批量规范化
正如我们前面提到的,批量规范化在训练模式和预测模式下的行为通常不同。
首先,将训练好的模型用于预测时,我们不再需要样本均值中的噪声以及在微批次上估计每个小批次产生的样本方差了。
其次,例如,我们可能需要使用我们的模型对逐个样本进行预测。
一种常用的方法是通过移动平均估算整个训练数据集的样本均值和方差,并在预测时使用它们得到确定的输出。
可见,和暂退法一样,批量规范化层在训练模式和预测模式下的计算结果也是不一样的。
## (**从零实现**)
下面,我们从头开始实现一个具有张量的批量规范化层。
```
import tensorflow as tf
from d2l import tensorflow as d2l
def batch_norm(X, gamma, beta, moving_mean, moving_var, eps):
# 计算移动方差元平方根的倒数
inv = tf.cast(tf.math.rsqrt(moving_var + eps), X.dtype)
# 缩放和移位
inv *= gamma
Y = X * inv + (beta - moving_mean * inv)
return Y
```
我们现在可以[**创建一个正确的`BatchNorm`图层**]。
这个层将保持适当的参数:拉伸`gamma`和偏移`beta`,这两个参数将在训练过程中更新。
此外,我们的图层将保存均值和方差的移动平均值,以便在模型预测期间随后使用。
撇开算法细节,注意我们实现图层的基础设计模式。
通常情况下,我们用一个单独的函数定义其数学原理,比如说`batch_norm`。
然后,我们将此功能集成到一个自定义层中,其代码主要处理簿记问题,例如将数据移动到训练设备(如GPU)、分配和初始化任何必需的变量、跟踪移动平均线(此处为均值和方差)等。
为了方便起见,我们并不担心在这里自动推断输入形状,因此我们需要指定整个特征的数量。
不用担心,深度学习框架中的批量规范化API将为我们解决上述问题,我们稍后将展示这一点。
```
class BatchNorm(tf.keras.layers.Layer):
def __init__(self, **kwargs):
super(BatchNorm, self).__init__(**kwargs)
def build(self, input_shape):
weight_shape = [input_shape[-1], ]
# 参与求梯度和迭代的拉伸和偏移参数,分别初始化成1和0
self.gamma = self.add_weight(name='gamma', shape=weight_shape,
initializer=tf.initializers.ones, trainable=True)
self.beta = self.add_weight(name='beta', shape=weight_shape,
initializer=tf.initializers.zeros, trainable=True)
# 非模型参数的变量初始化为0和1
self.moving_mean = self.add_weight(name='moving_mean',
shape=weight_shape, initializer=tf.initializers.zeros,
trainable=False)
self.moving_variance = self.add_weight(name='moving_variance',
shape=weight_shape, initializer=tf.initializers.ones,
trainable=False)
super(BatchNorm, self).build(input_shape)
def assign_moving_average(self, variable, value):
momentum = 0.9
delta = variable * momentum + value * (1 - momentum)
return variable.assign(delta)
@tf.function
def call(self, inputs, training):
if training:
axes = list(range(len(inputs.shape) - 1))
batch_mean = tf.reduce_mean(inputs, axes, keepdims=True)
batch_variance = tf.reduce_mean(tf.math.squared_difference(
inputs, tf.stop_gradient(batch_mean)), axes, keepdims=True)
batch_mean = tf.squeeze(batch_mean, axes)
batch_variance = tf.squeeze(batch_variance, axes)
mean_update = self.assign_moving_average(
self.moving_mean, batch_mean)
variance_update = self.assign_moving_average(
self.moving_variance, batch_variance)
self.add_update(mean_update)
self.add_update(variance_update)
mean, variance = batch_mean, batch_variance
else:
mean, variance = self.moving_mean, self.moving_variance
output = batch_norm(inputs, moving_mean=mean, moving_var=variance,
beta=self.beta, gamma=self.gamma, eps=1e-5)
return output
```
## 使用批量规范化层的 LeNet
为了更好理解如何[**应用`BatchNorm`**],下面我们将其应用(**于LeNet模型**)( :numref:`sec_lenet`)。
回想一下,批量规范化是在卷积层或全连接层之后、相应的激活函数之前应用的。
```
# 回想一下,这个函数必须传递给d2l.train_ch6。
# 或者说为了利用我们现有的CPU/GPU设备,需要在strategy.scope()建立模型
def net():
return tf.keras.models.Sequential([
tf.keras.layers.Conv2D(filters=6, kernel_size=5,
input_shape=(28, 28, 1)),
BatchNorm(),
tf.keras.layers.Activation('sigmoid'),
tf.keras.layers.AvgPool2D(pool_size=2, strides=2),
tf.keras.layers.Conv2D(filters=16, kernel_size=5),
BatchNorm(),
tf.keras.layers.Activation('sigmoid'),
tf.keras.layers.AvgPool2D(pool_size=2, strides=2),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(120),
BatchNorm(),
tf.keras.layers.Activation('sigmoid'),
tf.keras.layers.Dense(84),
BatchNorm(),
tf.keras.layers.Activation('sigmoid'),
tf.keras.layers.Dense(10)]
)
```
和以前一样,我们将[**在Fashion-MNIST数据集上训练网络**]。
这个代码与我们第一次训练LeNet( :numref:`sec_lenet`)时几乎完全相同,主要区别在于学习率大得多。
```
lr, num_epochs, batch_size = 1.0, 10, 256
train_iter, test_iter = d2l.load_data_fashion_mnist(batch_size)
net = d2l.train_ch6(net, train_iter, test_iter, num_epochs, lr, d2l.try_gpu())
```
让我们来看看从第一个批量规范化层中学到的[**拉伸参数`gamma`和偏移参数`beta`**]。
```
tf.reshape(net.layers[1].gamma, (-1,)), tf.reshape(net.layers[1].beta, (-1,))
```
## [**简明实现**]
除了使用我们刚刚定义的`BatchNorm`,我们也可以直接使用深度学习框架中定义的`BatchNorm`。
该代码看起来几乎与我们上面的代码相同。
```
def net():
return tf.keras.models.Sequential([
tf.keras.layers.Conv2D(filters=6, kernel_size=5,
input_shape=(28, 28, 1)),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Activation('sigmoid'),
tf.keras.layers.AvgPool2D(pool_size=2, strides=2),
tf.keras.layers.Conv2D(filters=16, kernel_size=5),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Activation('sigmoid'),
tf.keras.layers.AvgPool2D(pool_size=2, strides=2),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(120),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Activation('sigmoid'),
tf.keras.layers.Dense(84),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Activation('sigmoid'),
tf.keras.layers.Dense(10),
])
```
下面,我们[**使用相同超参数来训练模型**]。
请注意,通常高级API变体运行速度快得多,因为它的代码已编译为C++或CUDA,而我们的自定义代码由Python实现。
```
d2l.train_ch6(net, train_iter, test_iter, num_epochs, lr, d2l.try_gpu())
```
## 争议
直观地说,批量规范化被认为可以使优化更加平滑。
然而,我们必须小心区分直觉和对我们观察到的现象的真实解释。
回想一下,我们甚至不知道简单的神经网络(多层感知机和传统的卷积神经网络)为什么如此有效。
即使在暂退法和权重衰减的情况下,它们仍然非常灵活,因此无法通过常规的学习理论泛化保证来解释它们是否能够泛化到看不见的数据。
在提出批量规范化的论文中,作者除了介绍了其应用,还解释了其原理:通过减少*内部协变量偏移*(internal covariate shift)。
据推测,作者所说的“内部协变量转移”类似于上述的投机直觉,即变量值的分布在训练过程中会发生变化。
然而,这种解释有两个问题:
i)这种偏移与严格定义的*协变量偏移*(covariate shift)非常不同,所以这个名字用词不当。
ii)这种解释只提供了一种不明确的直觉,但留下了一个有待后续挖掘的问题:为什么这项技术如此有效?。
本书旨在传达实践者用来发展深层神经网络的直觉。
然而,重要的是将这些指导性直觉与既定的科学事实区分开来。
最终,当你掌握了这些方法,并开始撰写自己的研究论文时,你会希望清楚地区分技术和直觉。
随着批量规范化的普及,“内部协变量偏移”的解释反复出现在技术文献的辩论,特别是关于“如何展示机器学习研究”的更广泛的讨论中。
Ali Rahimi在接受2017年NeurIPS大会的“接受时间考验奖”(Test of Time Award)时发表了一篇令人难忘的演讲。他将“内部协变量转移”作为焦点,将现代深度学习的实践比作炼金术。
他对该示例进行了详细回顾 :cite:`Lipton.Steinhardt.2018`,概述了机器学习中令人不安的趋势。
此外,一些作者对批量规范化的成功提出了另一种解释:在某些方面,批量规范化的表现出与原始论文 :cite:`Santurkar.Tsipras.Ilyas.ea.2018`中声称的行为是相反的。
然而,与技术机器学习文献中成千上万类似模糊的声明相比,内部协变量偏移没有什么更值得批评。
很可能,它作为这些辩论的焦点而产生共鸣,要归功于它对目标受众的广泛认可。
批量规范化已经被证明是一种不可或缺的方法。它适用于几乎所有图像分类器,并在学术界获得了数万引用。
## 小结
* 在模型训练过程中,批量规范化利用小批量的均值和标准差,不断调整神经网络的中间输出,使整个神经网络各层的中间输出值更加稳定。
* 批量规范化在全连接层和卷积层的使用略有不同。
* 批量规范化层和暂退层一样,在训练模式和预测模式下计算不同。
* 批量规范化有许多有益的副作用,主要是正则化。另一方面,”减少内部协变量偏移“的原始动机似乎不是一个有效的解释。
## 练习
1. 在使用批量规范化之前,我们是否可以从全连接层或卷积层中删除偏置参数?为什么?
1. 比较LeNet在使用和不使用批量规范化情况下的学习率。
1. 绘制训练和测试准确度的提高。
1. 你的学习率有多高?
1. 我们是否需要在每个层中进行批量规范化?尝试一下?
1. 你可以通过批量规范化来替换暂退法吗?行为如何改变?
1. 确定参数`beta`和`gamma`,并观察和分析结果。
1. 查看高级API中有关`BatchNorm`的在线文档,以查看其他批量规范化的应用。
1. 研究思路:想想你可以应用的其他“规范化”转换?你可以应用概率积分变换吗?全秩协方差估计如何?
[Discussions](https://discuss.d2l.ai/t/1875)
| github_jupyter |
<!--NAVIGATION-->
_______________
Este documento puede ser utilizado de forma interactiva en las siguientes plataformas:
- [Google Colab](https://colab.research.google.com/github/masdeseiscaracteres/ml_course/blob/master/material/03_02_feature_selection_filter_methods.ipynb)
- [MyBinder](https://mybinder.org/v2/gh/masdeseiscaracteres/ml_course/master)
- [Deepnote](https://deepnote.com/launch?template=python_3.6&url=https%3A%2F%2Fgithub.com%2Fmasdeseiscaracteres%2Fml_course%2Fblob%2Fmaster%2Fmaterial%2F03_02_feature_selection_filter_methods.ipynb)
_______________
# Métodos de filtrado
En este notebook se revisarán los siguientes conceptos:
1. Métodos de filtrado para regresión
2. Métodos de filtrado para clasificación
3. Métodos de filtrado en un problema realista
Primero cargamos librerías y funciones necesarias:
## 0. Configuración del entorno
```
# clonar el resto del repositorio si no está disponible
import os
curr_dir = os.getcwd()
if not os.path.exists(os.path.join(curr_dir, '../.ROOT_DIR')):
!git clone https://github.com/masdeseiscaracteres/ml_course.git ml_course
os.chdir(os.path.join(curr_dir, 'ml_course/material'))
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
%matplotlib inline
cm = plt.cm.RdBu
cm_bright = ListedColormap(['#FF0000', '#0000FF'])
import warnings
warnings.filterwarnings('ignore')
```
## 1. Métodos de filtrado para regresión
Vamos a practicar con los métodos de [filtrado](http://scikit-learn.org/stable/modules/feature_selection.html) para regresión siguientes:
* [f_regression](http://scikit-learn.org/stable/modules/generated/sklearn.feature_selection.f_regression.html#sklearn.feature_selection.f_regression): evalúa la significancia del coeficiente de correlación de Pearson mediante [este](https://en.wikipedia.org/wiki/Pearson_correlation_coefficient#Testing_using_Student's_t-distribution) procedimiento.
* [mutual_info_regression](http://scikit-learn.org/stable/modules/generated/sklearn.feature_selection.mutual_info_regression.html#sklearn.feature_selection.mutual_info_regression): calcula una estima de la información mutua entre cada característica y la variable objetivo.
Para ello, vamos a aplicarlos a un ejemplo sencillo que construiremos sintéticamente.
### Ejercicio
Se tienen tres variables $x_1, x_2,x_3$, las cuales son variables aleatorias que se distribuyen uniformemente en el intervalo $[0, 1]$. La variable objetivo depende de estas variables de la forma
$$y = x_1 + \sin{(6\pi x_2)} + 0.1\mathcal{N}(0, 1).$$
Esto es:
* $y$ depende linealmente de $x_1$.
* $y$ depende no linealmente de $x_2$.
* $y$ no depende de $x_3$.
Por tanto, $x_3$ es una variable irrelevante para $y$.
```
# Ejemplo tomado de
# http://scikit-learn.org/stable/auto_examples/feature_selection/plot_f_test_vs_mi.html#sphx-glr-auto-examples-feature-selection-plot-f-test-vs-mi-py
from sklearn.feature_selection import f_regression, mutual_info_regression
np.random.seed(0)
X = np.random.rand(1000, 3)
y = X[:, 0] + np.sin(6 * np.pi * X[:, 1]) + 0.1 * np.random.randn(1000)
```
Veamos qué nos dicen los test de filtrado de selección de características.
### Solución
```
f_test, _ = f_regression(X, y)
f_test /= np.max(f_test)
mi = mutual_info_regression(X, y)
mi /= np.max(mi)
# Dibujamos cómo se relaciona la variable objetivo con cada una de las variables x_i
plt.figure(figsize=(15, 5))
for i in range(3):
plt.subplot(1, 3, i + 1)
plt.scatter(X[:, i], y, edgecolor='black', s=20, alpha=0.3)
plt.xlabel("$x_{}$".format(i + 1), fontsize=14)
if i == 0:
plt.ylabel("$y$", fontsize=14)
plt.title("F-test={:.2f}, MI={:.2f}".format(f_test[i], mi[i]),
fontsize=16) # mostramos
plt.show()
```
Como podemos ver, los valores obtenidos en las métricas son consistentes con las hipótesis que habíamos planteado:
- $x_1$: un alto valor de significancia en el F-test sugiere una **relación lineal** con la variable objetivo.
- $x_2$: un alto valor de información mutua sugiere una relación **no lineal** con la variable objetivo.
- $x_3$: ambas métricas dan valores nulos, por lo que no parece existir **ningún tipo de relación** con la variable objetivo.
## 2. Métodos de filtrado para clasificación
Vamos a analizar los mismos métodos de [filtrado](http://scikit-learn.org/stable/modules/feature_selection.html) que en el caso de regresión, aplicados ahora para distintos problemas de clasificación:
* [f_classif](http://scikit-learn.org/stable/modules/generated/sklearn.feature_selection.f_classif.html#sklearn.feature_selection.f_classif)
* [mutual_info_classif](http://scikit-learn.org/stable/modules/generated/sklearn.feature_selection.mutual_info_classif.html#sklearn.feature_selection.mutual_info_classif)
### Ejercicio: problema de clasificación linealmente separable
En este problema, la variable $x_1$ define un problema linealmente separable y constituye la variable informativa. $x_2$ representa una variable redundante, mientras que $x_3$ y $x_4$ son variables ruidosas (no mejoran la capacidad de discernir entre las dos clases).
```
# toy example: linearly separable problem
np.random.seed(0)
# -- parameters
N = 1000
mu = 1 # CHANGE THIS VALUE
sigma1 = 1 # CHANGE THIS VALUE
sigma2 = 1 # CHANGE THIS VALUE
# -- variables auxiliares
unos = np.ones(int(N/2))
rand2 = np.random.randn(int(N/2),1)
# -- features
y = np.concatenate([-1*unos, unos])
X1 = np.concatenate([-mu + sigma1*rand2, mu + sigma1*rand2])
X2 = sigma2*np.random.randn(N,1) - 3*X1
X3 = 2*np.random.randn(N,1)
X4 = np.random.rand(N,1)
X = np.hstack((X1, X2, X3, X4))
#--- do some plotting
plt.figure(figsize=(12, 9))
for i in range(4):
plt.subplot(2, 2, i + 1)
plt.hist(X[y<0,i],bins=20, normed=True, alpha=0.5, label='-1',color='b')
plt.hist(X[y>0,i], bins=20, normed=True, alpha=0.5, label='+1',color='r')
plt.legend(loc='upper right')
plt.xlabel("$x_{}$".format(i + 1), fontsize=18)
# Recordemos que x_1 y x_2 están relacionadas linealmente
plt.scatter(X1, X2, cmap=cm_bright, c = y.reshape(-1,1))
plt.xlabel("$x_1$", fontsize=16)
plt.ylabel("$x_2$", fontsize=16)
plt.show()
```
Apliquemos los métodos de filtrado en este problema de clasificación.
### Solución
```
from sklearn.feature_selection import f_classif, mutual_info_classif
featureNames = ['x1','x2','x3','x4']
# do calculations
f_test, _ = f_classif(X, y)
#f_test /= np.max(f_test)
mi = mutual_info_classif(X, y)
#mi /= np.max(mi)
# do some plotting
plt.figure(figsize=(10, 5))
plt.subplot(1,2,1)
plt.bar(range(X.shape[1]), f_test, align="center")
plt.xticks(range(X.shape[1]), featureNames)
plt.xlabel('features')
plt.ylabel('Ranking')
plt.title('$F-test$ score')
plt.subplot(1,2,2)
plt.bar(range(X.shape[1]),mi, align="center")
plt.xticks(range(X.shape[1]),featureNames)
plt.xlabel('features')
plt.ylabel('Ranking')
plt.title('Mutual information score')
plt.show()
```
### Ejercicio
Modifica los parámetros `mu`, `sigma1`, y `sigma2` y analiza sus efectos en las representaciones anteriores. ¿Qué variable puedes considerar relevante?
### Solución
La única variable relevante es $x_1$ mientras su desviación típica asociada, `sigma1`, no sea mucho más grande que `mu`. La capacidad de distinguir entre clases utilizando $x_1$ disminuye si esto no se cumple.
### Ejercicio: problema de clasificación separable no linealmente
Vamos a analizar el problema XOR, definido por las variables $x_1$ y $x_2$. Añadimos además dos variables relevantes $x_3$ y $x_4$, con capacidad discriminatoria y que, por tanto, pueden ayudar a mejorar las prestaciones del problema de clasificación bajo estudio. Por último, se incluyen 20 variables ruidosas irrelevantes.
```
np.random.seed(0)
# parámetros
N = 1000
mu = 1.5
sigma = 0.5
# variables auxiliares
unos = np.ones(int(N/4))
random4 = sigma*np.random.randn(int(N/4),1)
random2 = sigma*np.random.randn(int(N/2),1)
# features
y = np.concatenate([-1*unos, unos, unos, -1*unos])
X1 = np.concatenate([-mu + random4, mu + random4, -mu + random4, mu + random4])
X2 = np.concatenate([+mu + random2, -mu + random2])
X3 = 3*(X1+X2) + np.sqrt(2)*np.random.randn(N,1)
X4 = 2*np.square((X1+X2)) + np.sqrt(2)*np.random.randn(N,1)
E = 2*np.random.randn(N, 20) # noisy variables
X = np.hstack((X1,X2,X3,X4,E))
plt.figure(figsize=(12, 10))
plt.subplot(2, 2, 1)
plt.scatter(X1,X2,c=y.reshape(-1,1), cmap=cm_bright)
plt.xlabel("$x_1$", fontsize=16)
plt.ylabel("$x_2$", fontsize=16)
for i in range(3):
plt.subplot(2, 2, i + 2)
plt.hist(X[y<0,i+1],bins=20, normed=True, alpha=0.5, label='-1',color='b')
plt.hist(X[y>0,i+1], bins=20, normed=True, alpha=0.5, label='+1',color='r')
plt.legend(loc='upper right')
plt.xlabel("$x_{}$".format(i + 2), fontsize=18)
plt.show()
```
Investiga la relevancia de las variables
### Solución
```
featureNames = ['x' + str(i) for i in range(1, 25)]
# do calculations
f_test, _ = f_classif(X, y)
f_test /= np.max(f_test)
mi = mutual_info_classif(X, y)
mi /= np.max(mi)
# do some plotting
plt.figure(figsize=(20, 5))
plt.subplot(1,2,1)
plt.bar(range(X.shape[1]),f_test, align="center")
plt.xticks(range(X.shape[1]),featureNames, rotation = 45)
plt.xlabel('features')
plt.ylabel('Ranking')
plt.title('$R^2$ score')
plt.subplot(1,2,2)
plt.bar(range(X.shape[1]),mi, align="center")
plt.xticks(range(X.shape[1]),featureNames, rotation = 45)
plt.xlabel('features')
plt.ylabel('Ranking')
plt.title('Mutual information score')
plt.show()
```
## 3. Métodos de filtrado sobre problema realista
Vamos a aplicar los métodos de filtrado sobre la base de datos de viviendas [House Sales in King County, USA](https://www.kaggle.com/harlfoxem/housesalesprediction)
Recordamos que para cada vivienda, se tienen los siguientes atributos, características o features:
| Atributo | descripción |
| :- |:- |
|*id*| identificador de la vivienda|
| *date*| fecha
| *price*| precio
| *bedrooms*| número de habitaciones
| *bathrooms*| número de baños/aseos
| *sqtf_living*| superficie habitable (en pies al cuadrado)
| *sqft_lot*| superficie de la parcela (en pies al cuadrado)
| *floors*| número de plantas
| *waterfront*| indica si la vivienda tiene acceso a un lago
| *view*| tipo de vista (variable numérica)
| *condition*| condición de la vivienda (variable númerica)
| *grade*| medida de la calidad de la construcción (variable numérica)
| *sqft_above*| superficie por encima del suelo (en pies al cuadrado)
| *sqft_basement*| superficie del sótano (en pies al cuadrado)
| *yr_built*| año de construcción de la vivienda
| *yr_renovated*| año de renovación de la vivienda
| *lat*| latitud de la parcela
| *long*| longitud de la parcela
| *sqft_living15*| superficie habitable promedio de los 15 vecinos más cercanos
| *sqft_lot15*| superficie de la parcela promedio de los 15 vecinos más cercanos
```
# cargamos datos
house_data = pd.read_csv("./data/kc_house_data.csv") # cargamos fichero
# Eliminamos las columnas id y date
house_data = house_data.drop(['id', 'date'], axis=1)
# convertir las variables en pies al cuadrado en metros al cuadrado
feetFeatures = ['sqft_living', 'sqft_lot', 'sqft_above', 'sqft_basement', 'sqft_living15', 'sqft_lot15']
house_data[feetFeatures] = house_data[feetFeatures].apply(lambda x: x * 0.3048 * 0.3048)
# renombramos
house_data.columns = ['price','bedrooms','bathrooms','sqm_living','sqm_lot','floors','waterfront','view','condition',
'grade','sqm_above','sqm_basement','yr_built','yr_renovated','zip_code','lat','long',
'sqm_living15','sqm_lot15']
filtro = house_data.sqm_living <= 600
house_data = house_data[filtro]
# añadimos las nuevas variables
house_data['years'] = 2019 - house_data['yr_built']
#house_data['bedrooms_squared'] = house_data['bedrooms'].apply(lambda x: x**2)
#house_data['bed_bath_rooms'] = house_data['bedrooms']*house_data['bathrooms']
#house_data['log_sqm_living'] = house_data['sqm_living'].apply(lambda x: np.log(x))
#house_data['lat_plus_long'] = house_data['lat']*house_data['long']
house_data.head()
import seaborn as sns
# Compute the correlation matrix
corr = np.abs(house_data.drop(['price'], axis=1).corr())
# Generate a mask for the upper triangle
mask = np.zeros_like(corr, dtype=np.bool)
mask[np.triu_indices_from(mask)] = True
# Set up the matplotlib figure
f, ax = plt.subplots(figsize=(12, 10))
# Draw the heatmap with the mask and correct aspect ratio
sns.heatmap(corr, mask=mask,vmin = 0.0, vmax=1.0, center=0.5,
linewidths=.1, cmap="YlGnBu", cbar_kws={"shrink": .8})
plt.show()
# convertimos el DataFrame al formato necesario para scikit-learn
data = house_data.as_matrix()
y = data[:, 0] # nos quedamos con la 1ª columna, price
X = data[:,1:] # nos quedamos con el resto
feature_names = house_data.columns[1:]
# do calculations
f_test, _ = f_regression(X, y)
f_test /= np.max(f_test)
mi = mutual_info_regression(X, y)
mi /= np.max(mi)
# do some plotting
plt.figure(figsize=(20, 5))
plt.subplot(1,2,1)
plt.bar(range(X.shape[1]), f_test, align="center")
plt.xticks(range(X.shape[1]), feature_names, rotation = 90)
plt.xlabel('features')
plt.ylabel('Ranking')
plt.title('F-test score')
plt.subplot(1,2,2)
plt.bar(range(X.shape[1]), mi, align="center")
plt.xticks(range(X.shape[1]), feature_names, rotation = 90)
plt.xlabel('features')
plt.ylabel('Ranking')
plt.title('Mutual information score')
plt.show()
```
### Ejercicio
Las variables seleccionadas, ¿concuerdan con tu intuición?
### Solución
Sí, las variables relacionadas parecen concordar con la intuición habitual sobre los factores que influyen en el precio de una vivienda:
- El criterio lineal parece seleccionar las variables relacionadas con el tamaño de la vivienda (`sqm_*`) y calidad (`grade`).
- Por su parte, el criterio no lineal identifica las mismas variables y añade algunas relacionadas con la ubicación de la vivienda (`zip_code`, `lat`).
| github_jupyter |
# Interactive Data Maps
This notebook shows how to create maps in a notebook, *without* having to write a ton of JavaScript or Python code. It uses [folium](https://github.com/python-visualization/folium), which leverages [Leaflet.js](http://leafletjs.com/), a popular JavaScript library to create interactive maps. `folium` supports base maps using tilesets from [MapBox](https://www.mapbox.com/), [OpenStreetMap](http://www.openstreetmap.org/), and others, out of the box. `folium` also makes it very easy to plot data on maps using GeoJSON and TopoJSON overlays.
## Prerequisites
You will need to install the following in order to run this notebook.
* [folium](https://github.com/python-visualization/folium), to create and render geographical maps
* [xlrd](https://pypi.python.org/pypi/xlrd), to load excel spreadsheets
* [seaborn](http://stanford.edu/~mwaskom/software/seaborn/index.html), to create nice-looking statistical plots
* [matplotlib](http://matplotlib.org/), and [pandas](http://pandas.pydata.org/) 0.15.0 or later, to enable pie plots
```
!pip install folium==0.1.3
!pip install xlrd==0.9.3
!pip install seaborn==0.5.1
!pip install matplotlib==1.4.3
!pip install pandas==0.15.2
```
```
%matplotlib inline
import matplotlib.pyplot as plt
import pandas
import seaborn
```
## State Government Tax Collections
This notebook uses state government tax collections data from the US Census Bureau. The [data](http://www.census.gov/govs/statetax/) are for fiscal year 2013. The data file contains tax collections by US state governments for for a variety of tax categories, such as income, property, and sales taxes.
```
!wget -O 13staxcd.txt http://www2.census.gov/govs/statetax/13staxcd.txt
df = pandas.read_csv('13staxcd.txt', index_col='ST').dropna(axis=1)
# Because, yeah, values are in 1000s of dollars
df = df * 1000
df.head()
```
We need a second file that provides descriptions for the tax item codes (the TXX numbers).
```
!wget -O TaxItemCodesandDescriptions.xls http://www2.census.gov/govs/statetax/TaxItemCodesandDescriptions.xls
tax_codes_df = pandas.read_excel('TaxItemCodesandDescriptions.xls', 'Sheet1', index_col='Item Code')
tax_codes_df.head()
```
## Total taxes collected
The sum of taxes collected over by state and local governments for all categories in fiscal year 2013 is over **$846 billion**, with a 'b'.
```
print '${:,}'.format(df.sum().sum())
```
## Taxes by category
According to the data [source](http://www.census.gov/govs/statetax/):
>The Annual Survey of State Government Tax Collections (STC) provides a summary of taxes collected by state for 5 broad tax categories and up to 25 tax subcategories. These tables and data files present the details on tax collections by type of tax imposed and collected by state governments.
The only thing missing from the data thus far are the "5 broad tax categories", and which of the 25 subcategories make up each one. We had to [look this up](http://www.census.gov/govs/classification/), and download another [Excel file](http://www2.census.gov/govs/estimate/methodology_for_summary_tabulations.xls). There's also this [report](http://www2.census.gov/govs/statetax/2013stcreport.pdf), which provides some details about tax categorization, but also seems to contradict the Excel spreadsheet. Oh, the humanity.
```
!wget -O agg_tax_categories.xls http://www2.census.gov/govs/estimate/methodology_for_summary_tabulations.xls
tmp = pandas.read_excel('agg_tax_categories.xls')
tmp[8:21].dropna(how='all').dropna(how='all', axis=1).head()
```
After some investigation, we can write a short function to retrieve the major tax category by tax item code.
```
def category(tax_item):
'''Return tax category for the tax item code.'''
if tax_item == 'T01':
return 'Property Taxes'
elif tax_item in ['T40', 'T41']:
return 'Income Taxes'
elif tax_item in ['T09', 'T10', 'T11', 'T12', 'T13', 'T14', 'T15', 'T16', 'T19']:
return 'Sales and Gross Receipts Taxes'
elif tax_item in ['T20', 'T21', 'T22', 'T23', 'T24', 'T25', 'T26', 'T27', 'T28', 'T29']:
return 'License Taxes'
return 'Other Taxes'
```
Sum all taxes collected by broad category.
```
# assign broad category to each tax item code
tmp = df.copy()
tmp['Category'] = tmp.index.map(category)
# aggregate taxes collected by each state by broad category
by_category = tmp.groupby('Category').sum()
# sum across all states
totals_by_category = by_category.sum(axis=1)
print totals_by_category.map('${:,}'.format)
```
Plot the total taxes collected for by broad category.
```
totals_by_category.plot(kind='pie', labels=totals_by_category.index,
figsize=(10,10), autopct='%.1f%%')
```
Here is a [violin plot](http://stanford.edu/~mwaskom/software/seaborn/examples/elaborate_violinplot.html) (a combination of boxplot and kernel density plot) that shows the distribution of taxes collected for each category.
* The median sales and income taxes collected is below \$5 billion (each).
* The majority of states collect less than \$10 billion each in sales and income taxes.
* Some states collect as much as \$75 billion in income tax, and as much as \$48 billion in sales tax. Ouch.
```
data = by_category.T
fig, ax = plt.subplots(figsize=(14,10))
seaborn.violinplot(data, color="Set3", bw=.2, cut=.6,
lw=.5, inner="box", inner_kws={"ms": 6}, ax=ax)
print data[['Income Taxes', 'Sales and Gross Receipts Taxes']].describe()
```
## Taxes by state
Sum the taxes across all categories and view the states that collect the most taxes.
```
taxes_by_state = df.sum().sort(inplace=False, ascending=False)
taxes_by_state[:10].map('${:,}'.format)
```
It may not surprise anyone that California and New York top the list; however, it may surprise some that California collected almost twice as much tax revenue as New York. Here is a bar chart to help visualize the magnitude of taxes collected by state.
```
fig, ax = plt.subplots(figsize=(12,8))
data = taxes_by_state.reset_index()
data.columns = ['State', 'Taxes']
# plot values in $ billions
seaborn.barplot(data.index, data.Taxes / 1000000000,
ci=None, hline=.1, ax=ax)
ax.set_xticklabels(data.State)
ax.set_ylabel('$ Billions')
ax.set_xlabel('State')
ax.set_title('Taxes Collected by US State and Local Governments, FY 2013')
plt.tight_layout()
```
## Let's map our data
We want to overlay our tax data over a map of the United States. To do this, we'll use the following:
1. A [TopoJSON](https://github.com/mbostock/topojson/wiki) file that contains geographic borders of US states (and counties).
2. [US FIPS codes](http://en.wikipedia.org/wiki/Federal_Information_Processing_Standard_state_code) from the US Census bureau ([data file](http://www2.census.gov/geo/docs/reference/state.txt)). We'll use these to bind our data set to the state geometries in our TopoJSON file.
3. The [folium](https://github.com/python-visualization/folium) Python library to do the heavy lifting of rendering our map.
### Tax Data
Combine the data for the 25 tax subcategories with the cumulative amounts for the 5 broad categories. This will allow us to map both sets.
```
# the aggregate data by broad category
tmp = by_category.T
# make up our own tax item codes for broad categories
codes = ['I','L','O','P','S']
# create complete list of category names
category_names = tax_codes_df.Description.append(
pandas.Series(tmp.columns, index=codes)
)
# merge broad category data with data for 25 subcategories
tmp.columns = codes
data = df.T.merge(tmp, left_index=True, right_index=True)
data.head()
```
### TopoJSON File
We created a TopoJSON file of US borders using [us-atlas](https://github.com/mbostock/us-atlas). Creation of this file is beyond our scope here, but you can download it from our GitHub repository.
```
!wget -O us-states-10m.json https://raw.githubusercontent.com/knowledgeanyhow/notebooks/master/tax-maps/data/us-states-10m.json
us_topo_map = 'us-states-10m.json'
import os
assert os.path.isfile(us_topo_map)
statinfo = os.stat(us_topo_map)
assert statinfo.st_size > 0
```
### U.S. FIPS Codes
Our tax data is indexed by state. We need a way to bind our data to the state geometries in our map. The geometries in our TopoJSON file are keyed by FIPS codes (Federal Information Processing Standard). So we need to obtain the FIPS codes for US states (from the US Census Bureau), and add them to our data.
```
!wget -O us_state_FIPS.txt http://www2.census.gov/geo/docs/reference/state.txt
fips = pandas.read_csv('us_state_FIPS.txt', delimiter='|', index_col='STUSAB')
fips.head()
```
Add FIPS column to our data.
```
data['FIPS'] = data.index.map(lambda x: fips.loc[x]['STATE'])
data['FIPS'].head()
```
### Folium
Folium utilizes IPython's rich display to render maps as HTML. Here are two functions that use different mechanisms to render a map in a notebook. Either will work in modern browsers.
<div class="alert alert-block alert-warning" style="margin-top: 20px">As of this writing, folium==0.1.3 has a <a href="https://github.com/python-visualization/folium/issues/83">defect</a> that prevents some browsers from rendering maps within a notebook if the notebook server is running over https. The issue is caused by folium hardcoding http as the protocol to external CSS and JavaScript dependencies. If your browser prevents loading mixed content, the map may fail to display. Chrome users can opt for more lax security using <a href="https://support.google.com/chrome/answer/1342714?hl=en">this workaround</a>.
</div>
<div class="alert alert-block alert-info" style="margin-top: 20px">The maps below may not render in <a href="http://nbviewer.ipython.org/">nbviewer</a> because they embed JavaScript.
</div>
```
import folium
from IPython.display import HTML
def inline_map(map):
"""
Embeds the HTML source of the map directly into the IPython notebook.
This method will not work if the map depends on any files (json data). Also this uses
the HTML5 srcdoc attribute, which may not be supported in all browsers.
"""
map._build_map()
return HTML('<iframe srcdoc="{srcdoc}" style="width: 100%; height: 510px; border: none"></iframe>'.format(srcdoc=map.HTML.replace('"', '"')))
def embed_map(map, path="map.html"):
"""
Embeds a linked iframe to the map into the IPython notebook.
Note: this method will not capture the source of the map into the notebook.
This method should work for all maps (as long as they use relative urls).
"""
map.create_map(path=path)
return HTML('<iframe src="files/{path}" style="width: 100%; height: 510px; border: none"></iframe>'.format(path=path))
```
Now we create a function that accepts a tax code, creates a basemap of the United States, and adds a TopoJSON overlay with the appropriate state tax data bound to it.
```
def create_tax_map(tax_code, path='tax_map.html'):
'''
Create a base map with tax data bound to a GeoJSON overlay.
'''
# lookup tax category name
tax_name = category_names.loc[tax_code] + ' ($ Millions)'
# lookup tax data
d = data[['FIPS',tax_code]].copy()
d[tax_code] = d[tax_code] / 1000000L
# compute a color scale based on data values
max = d[tax_code].max()
color_scale = [max*q for q in [0, 0.1, 0.25, 0.5, 0.75, 0.95]]
# create base map
map = folium.Map(location=[40, -99], zoom_start=4, width=800)
# add TopoJSON overlay and bind data
map.geo_json(geo_path=us_topo_map, data_out='tax_map.json',
data=d, columns=d.columns,
key_on='feature.id',
threshold_scale=color_scale,
fill_color='PuBuGn', line_opacity=0.3,
legend_name=tax_name,
topojson='objects.states')
map.create_map(path=path)
return map
inline_map(create_tax_map('T40'))
```
### Add a widget
Use a widget to choose the tax category and render the map interactively.
```
from IPython.html import widgets
from IPython.display import display
from IPython.html.widgets import interact
tax_categories = category_names.to_dict()
tax_categories = dict(zip(tax_categories.values(), tax_categories.keys()))
dropdown = widgets.Dropdown(options=tax_categories, value='T40', description='Tax:')
def show_map(tax_code):
display(inline_map(create_tax_map(tax_code)))
widgets.interact(show_map, tax_code=dropdown)
```
<div class="alert" style="border: 1px solid #aaa; background: radial-gradient(ellipse at center, #ffffff 50%, #eee 100%);">
<div class="row">
<div class="col-sm-1"><img src="https://knowledgeanyhow.org/static/images/favicon_32x32.png" style="margin-top: -6px"/></div>
<div class="col-sm-11">This notebook was created using <a href="https://knowledgeanyhow.org">IBM Knowledge Anyhow Workbench</a>. To learn more, visit us at <a href="https://knowledgeanyhow.org">https://knowledgeanyhow.org</a>.</div>
</div>
</div>
| github_jupyter |
# Introduction to Software Testing
Before we get to the central parts of the book, let us introduce essential concepts of software testing. Why is it necessary to test software at all? How does one test software? How can one tell whether a test has been successful? How does one know if one has tested enough? In this chapter, let us recall the most important concepts, and at the same time get acquainted with Python and interactive notebooks.
```
from bookutils import YouTubeVideo
YouTubeVideo('wFuQ4Mju5dU')
```
This chapter (and this book) is not set to replace a textbook on testing; see the [Background](#Background) at the end for recommended reads.
## Simple Testing
Let us start with a simple example. Your co-worker has been asked to implement a square root function $\sqrt{x}$. (Let's assume for a moment that the environment does not already have one.) After studying the [Newton–Raphson method](https://en.wikipedia.org/wiki/Newton%27s_method), she comes up with the following Python code, claiming that, in fact, this `my_sqrt()` function computes square roots.
```
def my_sqrt(x):
"""Computes the square root of x, using the Newton-Raphson method"""
approx = None
guess = x / 2
while approx != guess:
approx = guess
guess = (approx + x / approx) / 2
return approx
```
Your job is now to find out whether this function actually does what it claims to do.
### Understanding Python Programs
If you're new to Python, you might first have to understand what the above code does. We very much recommend the [Python tutorial](https://docs.python.org/3/tutorial/) to get an idea on how Python works. The most important things for you to understand the above code are these three:
1. Python structures programs through _indentation_, so the function and `while` bodies are defined by being indented;
2. Python is _dynamically typed_, meaning that the type of variables like `x`, `approx`, or `guess` is determined at run-time.
3. Most of Python's syntactic features are inspired by other common languages, such as control structures (`while`, `if`), assignments (`=`), or comparisons (`==`, `!=`, `<`).
With that, you can already understand what the above code does: Starting with a `guess` of `x / 2`, it computes better and better approximations in `approx` until the value of `approx` no longer changes. This is the value that finally is returned.
### Running a Function
To find out whether `my_sqrt()` works correctly, we can *test* it with a few values. For `x = 4`, for instance, it produces the correct value:
```
my_sqrt(4)
```
The upper part above `my_sqrt(4)` (a so-called _cell_) is an input to the Python interpreter, which by default _evaluates_ it. The lower part (`2.0`) is its output. We can see that `my_sqrt(4)` produces the correct value.
The same holds for `x = 2.0`, apparently, too:
```
my_sqrt(2)
```
### Interacting with Notebooks
If you are reading this in the interactive notebook, you can try out `my_sqrt()` with other values as well. Click on one of the above cells with invocations of `my_sqrt()` and change the value – say, to `my_sqrt(1)`. Press `Shift+Enter` (or click on the play symbol) to execute it and see the result. If you get an error message, go to the above cell with the definition of `my_sqrt()` and execute this first. You can also run _all_ cells at once; see the Notebook menu for details. (You can actually also change the text by clicking on it, and corect mistaks such as in this sentence.)
```
from bookutils import quiz
quiz("What does `my_sqrt(16)` produce?",
[
"4",
"4.0",
"3.99999",
"None"
], "len('four') - len('two') + 1")
```
Try it out for yourself by uncommenting and executing the following line:
```
# my_sqrt(16)
```
Executing a single cell does not execute other cells, so if your cell builds on a definition in another cell that you have not executed yet, you will get an error. You can select `Run all cells above` from the menu to ensure all definitions are set.
Also keep in mind that, unless overwritten, all definitions are kept across executions. Occasionally, it thus helps to _restart the kernel_ (i.e. start the Python interpreter from scratch) to get rid of older, superfluous definitions.
### Debugging a Function
To see how `my_sqrt()` operates, a simple strategy is to insert `print()` statements in critical places. You can, for instance, log the value of `approx`, to see how each loop iteration gets closer to the actual value:
```
def my_sqrt_with_log(x):
"""Computes the square root of x, using the Newton–Raphson method"""
approx = None
guess = x / 2
while approx != guess:
print("approx =", approx) # <-- New
approx = guess
guess = (approx + x / approx) / 2
return approx
my_sqrt_with_log(9)
```
Interactive notebooks also allow to launch an interactive *debugger* – insert a "magic line" `%%debug` at the top of a cell and see what happens. Unfortunately, interactive debuggers interfere with our dynamic analysis techniques, so we mostly use logging and assertions for debugging.
### Checking a Function
Let's get back to testing. We can read and run the code, but are the above values of `my_sqrt(2)` actually correct? We can easily verify by exploiting that $\sqrt{x}$ squared again has to be $x$, or in other words $\sqrt{x} \times \sqrt{x} = x$. Let's take a look:
```
my_sqrt(2) * my_sqrt(2)
```
Okay, we do have some rounding error, but otherwise, this seems just fine.
What we have done now is that we have _tested_ the above program: We have _executed_ it on a given input and _checked_ its result whether it is correct or not. Such a test is the bare minimum of quality assurance before a program goes into production.
## Automating Test Execution
So far, we have tested the above program _manually_, that is, running it by hand and checking its results by hand. This is a very flexible way of testing, but in the long run, it is rather inefficient:
1. Manually, you can only check a very limited number of executions and their results
2. After any change to the program, you have to repeat the testing process
This is why it is very useful to _automate_ tests. One simple way of doing so is to let the computer first do the computation, and then have it check the results.
For instance, this piece of code automatically tests whether $\sqrt{4} = 2$ holds:
```
result = my_sqrt(4)
expected_result = 2.0
if result == expected_result:
print("Test passed")
else:
print("Test failed")
```
The nice thing about this test is that we can run it again and again, thus ensuring that at least the square root of 4 is computed correctly. But there are still a number of issues, though:
1. We need _five lines of code_ for a single test
2. We do not care for rounding errors
3. We only check a single input (and a single result)
Let us address these issues one by one. First, let's make the test a bit more compact. Almost all programming languages do have a means to automatically check whether a condition holds, and stop execution if it does not. This is called an _assertion_, and it is immensely useful for testing.
In Python, the `assert` statement takes a condition, and if the condition is true, nothing happens. (If everything works as it should, you should not be bothered.) If the condition evaluates to false, though, `assert` raises an exception, indicating that a test just failed.
In our example, we can use `assert` to easily check whether `my_sqrt()` yields the expected result as above:
```
assert my_sqrt(4) == 2
```
As you execute this line of code, nothing happens: We just have shown (or asserted) that our implementation indeed produces $\sqrt{4} = 2$.
Remember, though, that floating-point computations may induce rounding errors. So we cannot simply compare two floating-point values with equality; rather, we would ensure that the absolute difference between them stays below a certain threshold value, typically denoted as $\epsilon$ or ``epsilon``. This is how we can do it:
```
EPSILON = 1e-8
assert abs(my_sqrt(4) - 2) < EPSILON
```
We can also introduce a special function for this purpose, and now do more tests for concrete values:
```
def assertEquals(x, y, epsilon=1e-8):
assert abs(x - y) < epsilon
assertEquals(my_sqrt(4), 2)
assertEquals(my_sqrt(9), 3)
assertEquals(my_sqrt(100), 10)
```
Seems to work, right? If we know the expected results of a computation, we can use such assertions again and again to ensure our program works correctly.
(Hint: a true Python programmer would use the function [`math.isclose()`](https://docs.python.org/3/library/math.html#math.isclose) instead.)
## Generating Tests
Remember that the property $\sqrt{x} \times \sqrt{x} = x$ universally holds? We can also explicitly test this with a few values:
```
assertEquals(my_sqrt(2) * my_sqrt(2), 2)
assertEquals(my_sqrt(3) * my_sqrt(3), 3)
assertEquals(my_sqrt(42.11) * my_sqrt(42.11), 42.11)
```
Still seems to work, right? Most importantly, though, $\sqrt{x} \times \sqrt{x} = x$ is something we can very easily test for thousands of values:
```
for n in range(1, 1000):
assertEquals(my_sqrt(n) * my_sqrt(n), n)
```
How much time does it take to test `my_sqrt()` with 100 values? Let's see.
We use our own [`Timer` module](Timer.ipynb) to measure elapsed time. To be able to use `Timer`, we first import our own utility module, which allows us to import other notebooks.
```
import bookutils
from Timer import Timer
with Timer() as t:
for n in range(1, 10000):
assertEquals(my_sqrt(n) * my_sqrt(n), n)
print(t.elapsed_time())
```
10,000 values take about a hundredth of a second, so a single execution of `my_sqrt()` takes 1/1000000 second, or about 1 microseconds.
Let's repeat this with 10,000 values picked at random. The Python `random.random()` function returns a random value between 0.0 and 1.0:
```
import random
with Timer() as t:
for i in range(10000):
x = 1 + random.random() * 1000000
assertEquals(my_sqrt(x) * my_sqrt(x), x)
print(t.elapsed_time())
```
Within a second, we have now tested 10,000 random values, and each time, the square root was actually computed correctly. We can repeat this test with every single change to `my_sqrt()`, each time reinforcing our confidence that `my_sqrt()` works as it should. Note, though, that while a random function is _unbiased_ in producing random values, it is unlikely to generate special values that drastically alter program behavior. We will discuss this later below.
## Run-Time Verification
Instead of writing and running tests for `my_sqrt()`, we can also go and _integrate the check right into the implementation._ This way, _each and every_ invocation of `my_sqrt()` will be automatically checked.
Such an _automatic run-time check_ is very easy to implement:
```
def my_sqrt_checked(x):
root = my_sqrt(x)
assertEquals(root * root, x)
return root
```
Now, whenever we compute a root with `my_sqrt_checked()`$\dots$
```
my_sqrt_checked(2.0)
```
... we already know that the result is correct, and will so for every new successful computation.
Automatic run-time checks, as above, assume two things, though:
* One has to be able to _formulate_ such run-time checks. Having concrete values to check against should always be possible, but formulating desired properties in an abstract fashion can be very complex. In practice, you need to decide which properties are most crucial, and design appropriate checks for them. Plus, run-time checks may depend not only on local properties, but on several properties of the program state, which all have to be identified.
* One has to be able to _afford_ such run-time checks. In the case of `my_sqrt()`, the check is not very expensive; but if we have to check, say, a large data structure even after a simple operation, the cost of the check may soon be prohibitive. In practice, run-time checks will typically be disabled during production, trading reliability for efficiency. On the other hand, a comprehensive suite of run-time checks is a great way to find errors and quickly debug them; you need to decide how many such capabilities you would still want during production.
```
quiz("Does run-time checking give a guarantee "
"that there will always be a correct result?",
[
"Yes",
"No",
], "1 ** 1 + 1 ** 1")
```
An important limitation of run-time checks is that they ensure correctness _only if there is a result_ to be checked - that is, they do _not_ guarantee that there always will be one. This is an important limitation compared to _symbolic verification techniques_ and program proofs, which can also guarantee that there is a result – at a much higher (often manual) effort, though.
## System Input vs Function Input
At this point, we may make `my_sqrt()` available to other programmers, who may then embed it in their code. At some point, it will have to process input that comes from _third parties_, i.e. is not under control by the programmer.
Let us simulate this *system input* by assuming a _program_ `sqrt_program()` whose input is a string under third-party control:
```
def sqrt_program(arg: str) -> None: # type: ignore
x = int(arg)
print('The root of', x, 'is', my_sqrt(x))
```
We assume that `sqrt_program` is a program which accepts system input from the command line, as in
```shell
$ sqrt_program 4
2
```
We can easily invoke `sqrt_program()` with some system input:
```
sqrt_program("4")
```
What's the problem? Well, the problem is that we do not check external inputs for validity. Try invoking `sqrt_program(-1)`, for instance. What happens?
Indeed, if you invoke `my_sqrt()` with a negative number, it enters an infinite loop. For technical reasons, we cannot have infinite loops in this chapter (unless we'd want the code to run forever); so we use a special `with ExpectTimeOut(1)` construct to interrupt execution after one second.
```
from ExpectError import ExpectTimeout
with ExpectTimeout(1):
sqrt_program("-1")
```
The above message is an _error message_, indicating that something went wrong. It lists the *call stack* of functions and lines that were active at the time of the error. The line at the very bottom is the line last executed; the lines above represent function invocations – in our case, up to `my_sqrt(x)`.
We don't want our code terminating with an exception. Consequently, when accepting external input, we must ensure that it is properly validated. We may write, for instance:
```
def sqrt_program(arg: str) -> None: # type: ignore
x = int(arg)
if x < 0:
print("Illegal Input")
else:
print('The root of', x, 'is', my_sqrt(x))
```
and then we can be sure that `my_sqrt()` is only invoked according to its specification.
```
sqrt_program("-1")
```
But wait! What happens if `sqrt_program()` is not invoked with a number?
```
quiz("What is the result of `sqrt_program('xyzzy')`?",
[
"0",
"0.0",
"`None`",
"An exception"
], "16 ** 0.5")
```
Let's try this out! When we try to convert a non-number string, this would also result in a runtime error:
```
from ExpectError import ExpectError
with ExpectError():
sqrt_program("xyzzy")
```
Here's a version which also checks for bad inputs:
```
def sqrt_program(arg: str) -> None: # type: ignore
try:
x = float(arg)
except ValueError:
print("Illegal Input")
else:
if x < 0:
print("Illegal Number")
else:
print('The root of', x, 'is', my_sqrt(x))
sqrt_program("4")
sqrt_program("-1")
sqrt_program("xyzzy")
```
We have now seen that at the system level, the program must be able to handle any kind of input gracefully without ever entering an uncontrolled state. This, of course, is a burden for programmers, who must struggle to make their programs robust for all circumstances. This burden, however, becomes a _benefit_ when generating software tests: If a program can handle any kind of input (possibly with well-defined error messages), we can also _send it any kind of input_. When calling a function with generated values, though, we have to _know_ its precise preconditions.
## The Limits of Testing
Despite best efforts in testing, keep in mind that you are always checking functionality for a _finite_ set of inputs. Thus, there may always be _untested_ inputs for which the function may still fail.
In the case of `my_sqrt()`, for instance, computing $\sqrt{0}$ results in a division by zero:
```
with ExpectError():
root = my_sqrt(0)
```
In our tests so far, we have not checked this condition, meaning that a program which builds on $\sqrt{0} = 0$ will surprisingly fail. But even if we had set up our random generator to produce inputs in the range of 0–1000000 rather than 1–1000000, the chances of it producing a zero value by chance would still have been one in a million. If the behavior of a function is radically different for few individual values, plain random testing has few chances to produce these.
We can, of course, fix the function accordingly, documenting the accepted values for `x` and handling the special case `x = 0`:
```
def my_sqrt_fixed(x):
assert 0 <= x
if x == 0:
return 0
return my_sqrt(x)
```
With this, we can now correctly compute $\sqrt{0} = 0$:
```
assert my_sqrt_fixed(0) == 0
```
Illegal values now result in an exception:
```
with ExpectError():
root = my_sqrt_fixed(-1)
```
Still, we have to remember that while extensive testing may give us a high confidence into the correctness of a program, it does not provide a guarantee that all future executions will be correct. Even run-time verification, which checks every result, can only guarantee that _if_ it produces a result, the result will be correct; but there is no guarantee that future executions may not lead to a failing check. As I am writing this, I _believe_ that `my_sqrt_fixed(x)` is a correct implementation of $\sqrt{x}$ for all finite numbers $x$, but I cannot be certain.
With the Newton-Raphson method, we may still have a good chance of actually _proving_ that the implementation is correct: The implementation is simple, the math is well-understood. Alas, this is only the case for few domains. If we do not want to go into full-fledged correctness proofs, our best chance with testing is to
1. Test the program on several, well-chosen inputs; and
2. Check results extensively and automatically.
This is what we do in the remainder of this course: Devise techniques that help us to thoroughly test a program, as well as techniques that help us checking its state for correctness. Enjoy!
## Lessons Learned
* The aim of testing is to execute a program such that we find bugs.
* Test execution, test generation, and checking test results can be automated.
* Testing is _incomplete_; it provides no 100% guarantee that the code is free of errors.
## Next Steps
From here, you can move on how to
* [use _fuzzing_ to test programs with random inputs](Fuzzer.ipynb)
Enjoy the read!
## Background
There is a large number of works on software testing and analysis.
* An all-new modern, comprehensive, and online textbook on testing is ["Software Testing: From Theory to Practice"](https://sttp.site) \cite{Aniche2020}, available [online](https://sttp.site). Much recommended!
* For this book, we are also happy to recommend "Software Testing and Analysis" \cite{Pezze2008} as an introduction to the field; its strong technical focus very well fits our methodology.
* Other important must-reads with a comprehensive approach to software testing, including psychology and organization, include "The Art of Software Testing" \cite{Myers2004} as well as "Software Testing Techniques" \cite{Beizer1990}.
## Exercises
### Exercise 1: Get Acquainted with Notebooks and Python
Your first exercise in this book is to get acquainted with notebooks and Python, such that you can run the code examples in the book – and try out your own. Here are a few tasks to get you started.
#### Beginner Level: Run Notebooks in Your Browser
The easiest way to get access to the code is to run them in your browser.
1. From the [Web Page](__CHAPTER_HTML__), check out the menu at the top. Select `Resources` $\rightarrow$ `Edit as Notebook`.
2. After a short waiting time, this will open a Jupyter Notebook right within your browser, containing the current chapter as a notebook.
3. You can again scroll through the material, but you click on any code example to edit and run its code (by entering <kbd>Shift</kbd> + <kbd>Return</kbd>). You can edit the examples as you please.
4. Note that code examples typically depend on earlier code, so be sure to run the preceding code first.
5. Any changes you make will not be saved (unless you save your notebook to disk).
For help on Jupyter Notebooks, from the [Web Page](__CHAPTER_HTML__), check out the `Help` menu.
#### Advanced Level: Run Python Code on Your Machine
This is useful if you want to make greater changes, but do not want to work with Jupyter.
1. From the [Web Page](__CHAPTER_HTML__), check out the menu at the top. Select `Resources` $\rightarrow$ `Download Code`.
2. This will download the Python code of the chapter as a single Python .py file, which you can save to your computer.
3. You can then open the file, edit it, and run it in your favorite Python environment to re-run the examples.
4. Most importantly, you can [import it](Importing.ipynb) into your own code and reuse functions, classes, and other resources.
For help on Python, from the [Web Page](__CHAPTER_HTML__), check out the `Help` menu.
#### Pro Level: Run Notebooks on Your Machine
This is useful if you want to work with Jupyter on your machine. This will allow you to also run more complex examples, such as those with graphical output.
1. From the [Web Page](__CHAPTER_HTML__), check out the menu at the top. Select `Resources` $\rightarrow$ `All Notebooks`.
2. This will download all Jupyter Notebooks as a collection of .ipynb files, which you can save to your computer.
3. You can then open the notebooks in Jupyter Notebook or Jupyter Lab, edit them, and run them. To navigate across notebooks, open the notebook [`00_Table_of_Contents.ipynb`](00_Table_of_Contents.ipynb).
4. You can also download individual notebooks using Select `Resources` $\rightarrow$ `Download Notebook`. Running these, however, will require that you have the other notebooks downloaded already.
For help on Jupyter Notebooks, from the [Web Page](__CHAPTER_HTML__), check out the `Help` menu.
#### Boss Level: Contribute!
This is useful if you want to contribute to the book with patches or other material. It also gives you access to the very latest version of the book.
1. From the [Web Page](__CHAPTER_HTML__), check out the menu at the top. Select `Resources` $\rightarrow$ `GitHub Repo`.
2. This will get you to the GitHub repository which contains all sources of the book, including the latest notebooks.
3. You can then _clone_ this repository to your disk, such that you get the latest and greatest.
4. You can report issues and suggest pull requests on the GitHub page.
5. Updating the repository with `git pull` will get you updated.
If you want to contribute code or text, check out the [Guide for Authors](Guide_for_Authors.ipynb).
### Exercise 2: Testing Shellsort
Consider the following implementation of a [Shellsort](https://en.wikipedia.org/wiki/Shellsort) function, taking a list of elements and (presumably) sorting it.
```
def shellsort(elems):
sorted_elems = elems.copy()
gaps = [701, 301, 132, 57, 23, 10, 4, 1]
for gap in gaps:
for i in range(gap, len(sorted_elems)):
temp = sorted_elems[i]
j = i
while j >= gap and sorted_elems[j - gap] > temp:
sorted_elems[j] = sorted_elems[j - gap]
j -= gap
sorted_elems[j] = temp
return sorted_elems
```
A first test indicates that `shellsort()` might actually work:
```
shellsort([3, 2, 1])
```
The implementation uses a _list_ as argument `elems` (which it copies into `sorted_elems`) as well as for the fixed list `gaps`. Lists work like _arrays_ in other languages:
```
a = [5, 6, 99, 7]
print("First element:", a[0], "length:", len(a))
```
The `range()` function returns an iterable list of elements. It is often used in conjunction with `for` loops, as in the above implementation.
```
for x in range(1, 5):
print(x)
```
#### Part 1: Manual Test Cases
Your job is now to thoroughly test `shellsort()` with a variety of inputs.
First, set up `assert` statements with a number of manually written test cases. Select your test cases such that extreme cases are covered. Use `==` to compare two lists.
**Solution.** Here are a few selected test cases:
```
# Standard lists
assert shellsort([3, 2, 1]) == [1, 2, 3]
assert shellsort([1, 2, 3, 4]) == [1, 2, 3, 4]
assert shellsort([6, 5]) == [5, 6]
# Check for duplicates
assert shellsort([2, 2, 1]) == [1, 2, 2]
# Empty list
assert shellsort([]) == []
```
#### Part 2: Random Inputs
Second, create random lists as arguments to `shellsort()`. Make use of the following helper predicates to check whether the result is (a) sorted, and (b) a permutation of the original.
```
def is_sorted(elems):
return all(elems[i] <= elems[i + 1] for i in range(len(elems) - 1))
is_sorted([3, 5, 9])
def is_permutation(a, b):
return len(a) == len(b) and all(a.count(elem) == b.count(elem) for elem in a)
is_permutation([3, 2, 1], [1, 3, 2])
```
Start with a random list generator, using `[]` as the empty list and `elems.append(x)` to append an element `x` to the list `elems`. Use the above helper functions to assess the results. Generate and test 1,000 lists.
**Solution.** Here's a simple random list generator:
```
def random_list():
length = random.randint(1, 10)
elems = []
for i in range(length):
elems.append(random.randint(0, 100))
return elems
random_list()
elems = random_list()
print(elems)
sorted_elems = shellsort(elems)
print(sorted_elems)
assert is_sorted(sorted_elems) and is_permutation(sorted_elems, elems)
```
Here's the test for 1,000 lists:
```
for i in range(1000):
elems = random_list()
sorted_elems = shellsort(elems)
assert is_sorted(sorted_elems) and is_permutation(sorted_elems, elems)
```
### Exercise 3: Quadratic Solver
Given an equation $ax^2 + bx + c = 0$, we want to find solutions for $x$ given the values of $a$, $b$, and $c$. The following code is supposed to do this, using the equation $$x = \frac{-b \pm \sqrt{b^2 - 4ac}}{2a}$$
```
def quadratic_solver(a, b, c):
q = b * b - 4 * a * c
solution_1 = (-b + my_sqrt_fixed(q)) / (2 * a)
solution_2 = (-b - my_sqrt_fixed(q)) / (2 * a)
return (solution_1, solution_2)
quadratic_solver(3, 4, 1)
```
The above implementation is incomplete, though. You can trigger
1. a division by zero; and
2. violate the precondition of `my_sqrt_fixed()`.
How does one do that, and how can one prevent this?
#### Part 1: Find bug-triggering inputs
For each of the two cases above, identify values for `a`, `b`, `c` that trigger the bug.
**Solution**. Here are two inputs that trigger the bugs:
```
with ExpectError():
print(quadratic_solver(3, 2, 1))
with ExpectError():
print(quadratic_solver(0, 0, 1))
```
#### Part 2: Fix the problem
Extend the code appropriately such that the cases are handled. Return `None` for nonexistent values.
**Solution.** Here is an appropriate extension of `quadratic_solver()` that takes care of all the corner cases:
```
def quadratic_solver_fixed(a, b, c):
if a == 0:
if b == 0:
if c == 0:
# Actually, any value of x
return (0, None)
else:
# No value of x can satisfy c = 0
return (None, None)
else:
return (-c / b, None)
q = b * b - 4 * a * c
if q < 0:
return (None, None)
if q == 0:
solution = -b / 2 * a
return (solution, None)
solution_1 = (-b + my_sqrt_fixed(q)) / (2 * a)
solution_2 = (-b - my_sqrt_fixed(q)) / (2 * a)
return (solution_1, solution_2)
with ExpectError():
print(quadratic_solver_fixed(3, 2, 1))
with ExpectError():
print(quadratic_solver_fixed(0, 0, 1))
```
#### Part 3: Odds and Ends
What are the chances of discovering these conditions with random inputs? Assuming one can do a billion tests per second, how long would one have to wait on average until a bug gets triggered?
**Solution.** Consider the code above. If we choose the full range of 32-bit integers for `a`, `b`, and `c`, then the first condition alone, both `a` and `b` being zero, has a chance of $p = 1 / (2^{32} * 2^{32})$; that is, one in 18.4 quintillions:
```
combinations = 2 ** 32 * 2 ** 32
combinations
```
If we can do a billion tests per second, how many years would we have to wait?
```
tests_per_second = 1000000000
seconds_per_year = 60 * 60 * 24 * 365.25
tests_per_year = tests_per_second * seconds_per_year
combinations / tests_per_year
```
We see that on average, we'd have to wait for 584 years. Clearly, pure random choices are not sufficient as sole testing strategy.
### Exercise 4: To Infinity and Beyond
When we say that `my_sqrt_fixed(x)` works for all _finite_ numbers $x$: What happens if you set $x$ to $\infty$ (infinity)? Try this out!
**Solution.** `my_sqrt_fixed(` $\infty$ `)` times out. This is because dividing $\infty$ by any finite number (as the Newton-Raphson-method does) again yields $\infty$, so we're stuck in an infinite loop:
```
from ExpectError import ExpectTimeout
infinity = float('inf') # that's how to get an infinite number
with ExpectTimeout(1):
y = my_sqrt_fixed(infinity)
```
Lesson learned: If you test a program with numbers, always be sure to include extreme values.
| github_jupyter |
```
# Importing core libraries
import numpy as np
import pandas as pd
from time import time
import pprint
import joblib
# Suppressing warnings because of skopt verbosity
import warnings
warnings.filterwarnings("ignore")
# Classifiers
import lightgbm as lgb
# Model selection
from sklearn.model_selection import StratifiedKFold
# Metrics
from sklearn.metrics import roc_auc_score
from sklearn.metrics import make_scorer
# Skopt functions
from skopt import BayesSearchCV
from skopt.callbacks import DeadlineStopper, DeltaYStopper
from skopt.space import Real, Categorical, Integer
# Plotting functions
import seaborn as sns
from matplotlib import pyplot as plt
#sns.set(style='whitegrid')
```
## 1. First steps
As first steps:
* we load the train and test data from disk
* we separate the target from the training data
* we separate the ids from the test data (thus train and test data have the same structure)
* we convert integer variables to categories (thus our machine learning algorithm can pick them as categorical variables and not standard numeric one)
```
# Loading data
X = pd.read_csv("../input/amazon-employee-access-challenge/train.csv")
X_test = pd.read_csv("../input/amazon-employee-access-challenge/test.csv")
# Separating the target from the predictors
y = X["ACTION"]
X.drop(["ACTION"], axis="columns", inplace=True)
# Separating the identifier from the test data
ids = X_test["id"]
X_test.drop("id", axis="columns", inplace=True)
# Converting all integer variables to categorical
integer_cols = X.select_dtypes(include=['int']).columns
X[integer_cols] = X[integer_cols].astype('category', copy=False)
X_test[integer_cols] = X_test[integer_cols].astype('category', copy=False)
```
# 2. EDA
At this point we have a look at the training and test data in order to figure out how we can process the data.
```
print("Unique values")
(pd.concat([X.apply(lambda x: len(x.unique())),
X_test.apply(lambda x: len(x.unique()))
], axis="columns")
.rename(columns={0: "train", 1:"test"}))
print("Values in test but not in train")
for col in integer_cols:
mismatched_codes = len(np.setdiff1d(X[col].unique(), X_test[col].unique()))
print(f"{col:20} {mismatched_codes:4}")
print("Missing cases")
(pd.concat([X.isna().sum(),
X_test.isna().sum()
], axis="columns")
.rename(columns={0: "train", 1:"test"}))
# label distribution
fig, ax = plt.subplots(1, 1, figsize=(10, 6))
sns.histplot(y, ax=ax)
plt.show()
# Distribution of values of variables
_ = X.astype(int).hist(bins='auto', figsize=(24, 22), layout=(5, 2))
```
From the EDA we can get a few hints about what to do:
* the target classes are unbalanced, we should consider re-balancing the data
* all the categorical features have a different number of values ranging from 70 to up to over 7000 (high cardinality features)
* there are no missing values but many categorical values appear just in test, not in train (this especially affects the RESOURCE feature)
* the categorical values are sparse
# 3. Feature engineering
As the categorical features are sparsed and mismatched, we replace the original values with contiguous values, substituing with the value -1 in the test set for values that are not present in the train set. This operation should permit the LightGBM
```
for col in integer_cols:
unique_values = sorted(X[col].unique())
print(col, ":", unique_values[:5],'...', unique_values[-5:])
conversion_dict = dict(zip(unique_values, range(len(unique_values))))
# When working with the Categorical’s codes, missing values will always have a code of -1.
X[col] = X[col].map(conversion_dict, na_action=-1).astype('category', copy=False)
X_test[col] = X_test[col].map(conversion_dict, na_action=-1).astype('category', copy=False)
print("Missing cases")
pd.concat([X.isna().sum(), X_test.isna().sum()], axis="columns").rename(columns={0: "train", 1:"test"})
```
# 4. Setting up optimization
First, we create a wrapper function to deal with running the optimizer and reporting back its best results.
```
# Reporting util for different optimizers
def report_perf(optimizer, X, y, title="model", callbacks=None):
"""
A wrapper for measuring time and performances of different optmizers
optimizer = a sklearn or a skopt optimizer
X = the training set
y = our target
title = a string label for the experiment
"""
start = time()
if callbacks is not None:
optimizer.fit(X, y, callback=callbacks)
else:
optimizer.fit(X, y)
d=pd.DataFrame(optimizer.cv_results_)
best_score = optimizer.best_score_
best_score_std = d.iloc[optimizer.best_index_].std_test_score
best_params = optimizer.best_params_
print((title + " took %.2f seconds, candidates checked: %d, best CV score: %.3f "
+ u"\u00B1"+" %.3f") % (time() - start,
len(optimizer.cv_results_['params']),
best_score,
best_score_std))
print('Best parameters:')
pprint.pprint(best_params)
print()
return best_params
```
We then define the evaluation metric, using the Scikit-learn function make_scorer allows us to convert the optimization into a minimization problem, as required by Scikit-optimize.
```
# Converting average precision score into a scorer suitable for model selection
roc_auc = make_scorer(roc_auc_score, greater_is_better=True, needs_threshold=True)
```
We set up a stratified 5-fold cross validation: stratification helps us to obtain representative folds of the data and the target, though the target is unbalanced.
```
# Setting a 5-fold stratified cross-validation (note: shuffle=True)
skf = StratifiedKFold(n_splits=5, shuffle=True, random_state=0)
```
We set up a generic LightGBM classifier.
```
clf = lgb.LGBMClassifier(boosting_type='gbdt',
metric='auc',
objective='binary',
n_jobs=1,
verbose=-1,
random_state=0)
```
We define a search space, expliciting the key hyper-parameters to optimize and the range where to look for the best values.
```
search_spaces = {
'learning_rate': Real(0.01, 1.0, 'log-uniform'), # Boosting learning rate
'n_estimators': Integer(30, 5000), # Number of boosted trees to fit
'num_leaves': Integer(2, 512), # Maximum tree leaves for base learners
'max_depth': Integer(-1, 256), # Maximum tree depth for base learners, <=0 means no limit
'min_child_samples': Integer(1, 256), # Minimal number of data in one leaf
'max_bin': Integer(100, 1000), # Max number of bins that feature values will be bucketed
'subsample': Real(0.01, 1.0, 'uniform'), # Subsample ratio of the training instance
'subsample_freq': Integer(0, 10), # Frequency of subsample, <=0 means no enable
'colsample_bytree': Real(0.01, 1.0, 'uniform'), # Subsample ratio of columns when constructing each tree
'min_child_weight': Real(0.01, 10.0, 'uniform'), # Minimum sum of instance weight (hessian) needed in a child (leaf)
'reg_lambda': Real(1e-9, 100.0, 'log-uniform'), # L2 regularization
'reg_alpha': Real(1e-9, 100.0, 'log-uniform'), # L1 regularization
'scale_pos_weight': Real(1.0, 500.0, 'uniform'), # Weighting of the minority class (Only for binary classification)
}
```
We then define the Bayesian optimization engine, providing to it our LightGBM, the search spaces, the evaluation metric, the cross-validation. We set a large number of possible experiments and some parallelism in the search operations.
```
opt = BayesSearchCV(estimator=clf,
search_spaces=search_spaces,
scoring=roc_auc,
cv=skf,
n_iter=3000, # max number of trials
n_points=3, # number of hyperparameter sets evaluated at the same time
n_jobs=-1, # number of jobs
iid=False, # if not iid it optimizes on the cv score
return_train_score=False,
refit=False,
optimizer_kwargs={'base_estimator': 'GP'}, # optmizer parameters: we use Gaussian Process (GP)
random_state=0) # random state for replicability
```
Finally we runt the optimizer and wait for the results. We have set some limits to its operations: we required it to stop if it cannot get consistent improvements from the search (DeltaYStopper) and time dealine set in seconds (we decided for 45 minutes).
```
overdone_control = DeltaYStopper(delta=0.0001) # We stop if the gain of the optimization becomes too small
time_limit_control = DeadlineStopper(total_time=60 * 45) # We impose a time limit (45 minutes)
best_params = report_perf(opt, X, y,'LightGBM',
callbacks=[overdone_control, time_limit_control])
```
# 5. Prediction on test data
Having got the best hyperparameters for the data at hand, we instantiate a lightGBM using such values and train our model on all the available examples.
<P>After having trained the model, we predict on the test set and we save the results on a csv file.
```
clf = lgb.LGBMClassifier(boosting_type='gbdt',
metric='auc',
objective='binary',
n_jobs=1,
verbose=-1,
random_state=0,
**best_params)
clf.fit(X, y)
submission = pd.DataFrame({'Id':ids, 'ACTION': clf.predict_proba(X_test)[:, 1].ravel()})
submission.to_csv("submission.csv", index = False)
```
| github_jupyter |
# Инициализация
```
#@markdown - **Монтирование GoogleDrive**
from google.colab import drive
drive.mount('GoogleDrive')
# #@markdown - **Размонтирование**
# !fusermount -u GoogleDrive
```
# Область кодов
```
#@title Алгоритм AdaBoost { display-mode: "both" }
# В программе реализован алгоритм AdaBoost для классификации стохастических данных
#@markdown [Оригинальная программа](https://github.com/wzyonggege/statistical-learning-method/blob/master/AdaBoost/Adaboost.ipynb)
# coding: utf-8
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
#@markdown - **Привязка данных**
class Bunch(dict):
def __init__(self,*args,**kwds):
super(Bunch,self).__init__(*args,**kwds)
self.__dict__ = self
#@markdown - **Генератор стохастических данных**
def generate_random(sigma, N, mu1=[15., 25., 20], mu2=[30., 40., 30]):
c = sigma.shape[-1]
X = np.zeros((N, c))
target = np.zeros((N,1))
for i in range(N):
if np.random.random(1) < 0.5:
X[i, :] = np.random.multivariate_normal(mu1, sigma[0, :, :], 1) # первая гауссовская модель
target[i] = 1
else:
X[i, :] = np.random.multivariate_normal(mu2, sigma[1, :, :], 1) # вторая гауссовская модель
target[i] = -1
return X, target
#@markdown - **Adaboost классификатор**
class AdaBoost:
def __init__(self, n_estimators=10, learning_rate=1.0):
self.clf_num = n_estimators
self.learning_rate = learning_rate
def init_args(self, datasets, labels):
self.X = datasets
self.Y = labels.flatten()
self.M, self.N = datasets.shape
# Набор слабых классификаторов
self.clf_sets = []
# Инициализация weights
self.weights = [1.0 / self.M] * self.M
# Коэффициенты alpha функции G(x)
self.alpha = []
# G (x) -- взвешенная сумма базовых классификаторов
# Базовые классификаторы-- y = direct * sign(x - v)
def G_fn(self, features, labels, weights):
m = len(features)
error = 100000.0 # бесконечность
best_v = 0.0
# Вычисление погрешности классификации каждого измерения features
features_min = min(features)
features_max = max(features)
n_step = (features_max - features_min + self.learning_rate) // self.learning_rate
# print('n_step:{}'.format(n_step))
direct, compare_array = None, None
for i in range(1, int(n_step)):
v = features_min + self.learning_rate * i
if v not in features:
# Вычисление ошибок
compare_array_positive = np.array([1 if features[k] > v else -1 for k in range(m)])
weight_error_positive = sum([weights[k] for k in range(m) if compare_array_positive[k] != labels[k]])
compare_array_nagetive = np.array([-1 if features[k] > v else 1 for k in range(m)])
weight_error_nagetive = sum([weights[k] for k in range(m) if compare_array_nagetive[k] != labels[k]])
if weight_error_positive < weight_error_nagetive:
weight_error = weight_error_positive
_compare_array = compare_array_positive
direct = 'positive'
else:
weight_error = weight_error_nagetive
_compare_array = compare_array_nagetive
direct = 'nagetive'
# print('v:{} error:{}'.format(v, weight_error))
if weight_error < error:
error = weight_error
compare_array = _compare_array
best_v = v
return best_v, direct, error, compare_array
# Вычисление alpha
def alpha_fn(self, error):
return 0.5 * np.log((1 - error) / error)
# Коэффициент нормализации
def Z_fn(self, weights, a, clf):
return sum([weights[i]*np.exp(-a * self.Y[i] * clf[i]) for i in range(self.M)])
# Обновление весов
def w_fn(self, a, clf, Z):
for i in range(self.M):
self.weights[i] = self.weights[i] * np.exp(-a * self.Y[i] * clf[i]) / Z
# Линейная комбинация G(x)
def f_fn(self, alpha, clf_sets):
pass
# Базовые классификаторы, v - порог
def G(self, x, v, direct):
if direct == 'positive':
return 1 if x > v else -1
else:
return -1 if x > v else 1
def fit(self, X, y):
self.init_args(X, y)
for epoch in range(self.clf_num):
best_clf_error, best_v, clf_result = 100000, None, None
for j in range(self.N):
features = self.X[:, j]
# порог классификации,ошибки классификации,результаты классификации
v, direct, error, compare_array = self.G_fn(features, self.Y, self.weights)
if error < best_clf_error:
best_clf_error = error
best_v = v
final_direct = direct
clf_result = compare_array
axis = j
print_list = [epoch + 1, self.clf_num, j, error, best_v]
print('epoch:{0[0]}/{0[1]}, feature:{0[2]}, error:{0[3]:.3f}, v:{0[4]:.3f}'.format(print_list))
if best_clf_error == 0:
best_clf_error = 1e-10
break
a = self.alpha_fn(best_clf_error)
self.alpha.append(a)
# Запись классификаторов
self.clf_sets.append((axis, best_v, final_direct))
# Коэффициент нормализации
Z = self.Z_fn(self.weights, a, clf_result)
# Обновление весов
self.w_fn(a, clf_result, Z)
print_list = [epoch + 1, self.clf_num, best_clf_error, best_v, final_direct, a]
print('classifier:{0[0]}/{0[1]}, error:{0[2]:.3f}, v:{0[3]:.3f}, direct:{0[4]}, a:{0[5]:.3f}'.format(print_list))
# print('weight:{}'.format(self.weights))
print('\n')
def corr_fn(self):
L = [tup[-1] for tup in self.clf_sets]
num_p = L.count('positive')
return 1 if num_p < 2 else -1
def predict(self, feature):
result = 0.0
for i in range(len(self.clf_sets)):
axis, clf_v, direct = self.clf_sets[i]
f_input = feature[axis]
result += self.alpha[i] * self.G(f_input, clf_v, direct)
# sign
return np.sign(result) * self.corr_fn()
def score(self, X_test, y_test):
right_count = 0
y_test = y_test.flatten()
for i in range(len(X_test)):
feature = X_test[i]
if self.predict(feature) == y_test[i]:
right_count += 1
return right_count / len(X_test)
#@markdown - **Стохастические данные**
k, N = 2, 400
sigma = np.zeros((k, 3, 3))
for i in range(k):
sigma[i, :, :] = np.diag(np.random.randint(10, 25, size=(3, )))
sample, target = generate_random(sigma, N)
feature_names = ['x_label', 'y_label', 'z_label']
target_names = ['gaussian1', 'gaussian2', 'gaussian3', 'gaussian4']
data = Bunch(sample=sample, feature_names=feature_names, target=target, target_names=target_names)
sample_t, target_t = generate_random(sigma, N)
data_t = Bunch(sample=sample_t, target=target_t)
#@markdown - **Обучение модели**
model = AdaBoost(n_estimators=3, learning_rate=0.5)
model.fit(data.sample, target)
#@markdown - **Точность классификации обучающих данных**
model.score(data.sample, target)
```
#### Роль corr_fn
Символ без изменений | Обращение символа
------------ | -------------
positive nagetive nagetive | nagetive positive positive
nagetive positive nagetive | positive nagetive positive
nagetive nagetive nagetive | positive positive positive
nagetive nagetive positive | positive positive nagetive
```
#@markdown - **Измерение признаков и порог базовых классификаторов**
model.clf_sets
#@markdown - **Веса базовых классификаторов**
model.alpha
#@markdown - **Тест**
# model = AdaBoost(n_estimators=3, learning_rate=0.5)
# model.fit(data.sample, target)
tar = [model.predict(x) for x in data.sample]
tar_train = np.array([model.predict(x) for x in data.sample], dtype=np.int8) + 1
tar_test = np.array([model.predict(x) for x in data_t.sample], dtype=np.int8) + 1
acc_train = model.score(data.sample, data.target)
acc_test = model.score(data_t.sample, data_t.target)
print_list = [acc_train*100, acc_test*100]
print('Accuracy on training set: {0[0]:.2f}%, accuracy on testing set: {0[1]:.2f}%.'.format(print_list))
#@markdown - **Представление распределения обучающих и тестовых данных**
titles = ['Random training data', 'Random testing data']
DATA = [data.sample, data_t.sample]
fig = plt.figure(1, figsize=(16, 8))
fig.subplots_adjust(wspace=.01, hspace=.02)
for i, title, data_n in zip([1, 2], titles, DATA):
ax = fig.add_subplot(1, 2, i, projection='3d')
ax.scatter(data_n[:,0], data_n[:,1], data_n[:,2], c='b', s=35, alpha=0.4, marker='o')
ax.set_xlabel('X')
ax.set_ylabel('Y')
ax.set_zlabel('Z')
ax.view_init(elev=20., azim=-25)
ax.set_title(title, fontsize=14)
plt.show()
#@markdown - **Представление результата классификации обучающих и тестовых данных**
titles = ['Classified training data by Adaboost', 'Classified testing data by Adaboost']
TAR = [tar_train, tar_test]
fig = plt.figure(2, figsize=(16, 8))
fig.subplots_adjust(wspace=.01, hspace=.02)
for i, title, data_n, tar in zip([1, 2], titles, DATA, TAR):
ax = fig.add_subplot(1, 2, i, projection='3d')
color=['b','g', 'r']
for j in range(N):
ax.scatter(data_n[j, 0], data_n[j, 1], data_n[j, 2], c=color[tar[j]], s=35, alpha=0.4, marker='P')
ax.set_xlabel('X')
ax.set_ylabel('Y')
ax.set_zlabel('Z')
ax.view_init(elev=20., azim=-25)
ax.set_title(title, fontsize=14, y=0.01)
plt.show()
```
| github_jupyter |
# Modelling of chip probabilities
```
import pandas, geopandas
import os, json
from sklearn.ensemble import (
RandomForestClassifier,
GradientBoostingClassifier,
HistGradientBoostingClassifier
)
from sklearn.model_selection import GridSearchCV
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import tools_chip_prob_modelling as tools
```
## Data
### Probabilities
Probabilities for each category by chip, as produced by the neural network:
```
probs = geopandas.read_parquet('/home/jovyan/data/32_nw_pred.parquet')
```
We also keep the names of all classes separate and semantically sorted as it'll come in handy afterwards:
```
tools.class_names
```
### Labels
Proportions of observed classes per chip. They are aligned with `probs` so we can remove the `geometry` column.
```
labels = pandas.read_parquet(
'/home/jovyan/data/chip_proportions.pq'
).drop('geometry', axis=1)
```
These are encoded in following a grouping described in the JSON model file:
```
cd2nm = tools.parse_nn_json(
'/home/jovyan/data/efficientnet_pooling_256_12.json'
)
```
We replace the code in `labels` by the name and aggregate by column names so we have the proportions in each chip by name rather than code:
```
relabels = labels.rename(columns=cd2nm).groupby(level=0, axis=1).sum()
```
`relabels` contains a few rows with a total probability sum of 0: these are chips that fall within the water. We remove them:
```
relabels_nw = relabels[relabels.sum(axis=1) > 0.5] # 0.5 to avoid rounding errors
```
To begin with, we focus on single class chips. We identify which are single class:
```
single_class = relabels_nw.where(relabels_nw==1).sum(axis=1).astype('bool')
single_class.sum() * 100 / single_class.shape[0] # Pct single class
```
And then pull the class for each of them:
```
label = pandas.Categorical(relabels_nw.idxmax(axis=1))
```
### Build `db_all`
Single table with probabilities, labels and single-class flag:
```
db_all = probs.join(
pandas.DataFrame({'label': label, 'single_class': single_class})
)
db_all['single_class'] = db_all['single_class'].fillna(False)
```
Note not all chips have a `label` and `single_class` value. To make sure they are consistent, we replace the `N/A` in `single_class` by a `False`.
### Train/Validation split
Next, we split the sample into train and validation sets. The training split proportion is set by the global parameter `TRAIN_FRAC`.
```
db_all['train_all'] = False
db_all.loc[
db_all.sample(frac=tools.TRAIN_FRAC, random_state=1234).index, 'train_all'
] = True
```
### Spatial lag
#### Overview
Some models will include spatial context. Here we calculate these measures through the spatial lag of the probability for each class. Given we are in a train/validation split context, there are two ways to approach the construction of the lag:
1. [`wa`] Absolute lag: we first calculate the lag for each chip, then split between train/validation. This ensures the lag is representative of the geographic context on the ground, but runs the risk of spilling information between train and validation samples. We say here "runs the risk" because the validation chips are _not_ used as focal chips for training and thus the model does not "see" them as such. However, validation chips are _also_ seen by the model as context, and that does make information cross between the sets. We consider this option as a way to explore the extent of this issue. One complication is that this approach also provides more information about geographic context. In the vent these models had higher performance (likely), it might not be clear which source is driving the boost: the better representation of geographic context, or the information cross-over between train and validation sets.
1. [`ws`] Split lag: here we start by splitting the set into train/validation and then we calculate the spatial lag separately. This is a cleaner way to assess performance, but also one that distorts geographical context so might potentially perform worse than a model that could include a better representation.
1. [`wbs`] Block split lag: here we use a different train/validation split that relies on blocks of 3x3 chips to not be separated. Also, we only use focal chips for model fitting and evaluation. This ensures a clean cut between train/val _and_ full geographic context, at the expense of less observations in the model.
After discussions, we have decided to start with `ws`. Upon further reflection, `wa` implies a clear case of "leakage" of information from the validation into the training sets, and thus "pollutes" the estimates of accuracy. The `wbs` remains valid but two caveats apply: a) a proper implementation of the approach results in an important loss of observations (for every 3x3 block of chips, only one of them enters the modeling), and this probably has an important effect on smaller classes such as the urbanities; b) at a theoretical level, it is still not clear it's the preferred approach (see [Wadoux et al. 2021](https://www.sciencedirect.com/science/article/pii/S0304380021002489?via=ihub); for context). On this last point, my (DA-B) sense is that it still makes sense to block in smaller chunks and split those between train and validation because you want to ensure that the effect of context is characterised adequately, and the risk of not picking up the variation desired in both training and validation sets (the main concern of Wadoux et al. 2021) is lessened by blocking at a relatively small scale and randomly sampling from those
#### Split lag
```
lag_all = db_all.groupby('train_all').apply(
tools.lag_columns, cols=tools.class_names
)
```
To make things handier later, we grab the names of the lagged variables separately:
```
w_class_names = ['w_'+i for i in tools.class_names]
```
Spatially lagging the split chips will give rise to islands (chips that do not have any neighbors). This will mean later on, when we consolidate the table for models, we will have to drop island observations and, if there is a systematic bias in the group (train/val) where we drop more, it might induce further issues down the line.
```
sns.displot(
lag_all.assign(
train_all=db_all['train_all'].map({True: "Train", False: "Validation"})
),
x='card',
hue='train_all',
element='step',
aspect=2,
height=3
);
```
Cardinalities clearly differ: trainining observations (a larger group) has more neighbors on average. This is potentially a problem in that the validation set is _not_ mimmicking the training one. However, the direction of the difference might be such that evaluation metrics could be taken as a _lower_ bound: the model is trained using more information from context but evaluated with less; performance will be evaluated with "worse" conditions than it's been trained on so, everything else being equal, it'd give lower scores than the model actually should.
**DA-B**: discuss with MF it is the case
```
corrs = tools.build_prob_wprob_corrs(
tools.class_names, w_class_names, db_all, lag_all
)
```
As an exploration, here we have the correlation between high probability in a given class, and high probability in other classes in the spatial lag (geographic context). There is a clear pattern of non-zero correlations.
```
h = sns.heatmap(corrs, vmin=0, vmax=1, cmap='viridis')
h.set_xticklabels(h.get_xticklabels(), rotation = 45, ha="right");
```
#### Block Split lag
`[TBA]`
### Interaction variables
Here we develop a set of interaction variables between the probability of each class and their spatial lag. In mathematical terms, the expected probability $\mu_i$ for chip $i$ in our base model is:
$$
\mu_i = \alpha + \sum_k P_{ki} \beta_k
$$
When we include the spatial lag, this becomes:
$$
\mu_i = \alpha + \sum_k P_{ki} \beta_k + \sum_k WP_{ki} \gamma_k
$$
The idea with interactions is to model the probability expectation as:
$$
\mu_i = \alpha + \sum_k P_{ki} \beta_k + \sum_k WP_{ki} \gamma_k + \sum_k \sum_k P_{ki} \times WP_{ki} \delta_{kk}
$$
Here we build a method to construct the last term in the equation above ($\sum_k \sum_k P_{ki} \times WP_{ki}$), which will be used when fitting some of the models below:
### Build `db`
Finally, we bring together in a single, clean table information on the original probabilities, their spatial lags, and class label. To make sure it is analysis-ready, we retain only single-class chips that are _not_ islands (i.e., have at least one neighbor).
```
db = db_all.join(
lag_all
).query(
'(single_class == True) & (island == False)'
).drop(['single_class', 'island'], axis=1)
```
The dropping of observations that happened above might also induce additional biases.
**DA-B** Explore further the distribution of values between train/val sets (e.g. heatmaps/hists before the split and after)
### Extract train/val indices
Now we have the set ready, we can pick out train/val indices as the modeling will expect them.
```
train_ids = db.query('train_all == True').index
val_ids = db.query('train_all == False').index
```
### Class (im)balance
The spatial lag step might be introducing bias in the sample before modelling. We pick all the single class chips and split them randomly between train and validation. If spatially lagging has a systematic effect on some classes (e.g., some classes get dropped more often), this might introduce bias that will make our model scoress unreliable. To explore this case, we want to get an idea of:
- [x] Overall proportion of train/val before (`db_all` single class) and after spatially lagging (`db`)
```
pandas.DataFrame(
{
'before_lag': db_all.query('single_class').value_counts(
'train_all'
) * 100 / len(db_all.query('single_class')),
'after_lag': db.value_counts('train_all') * 100 / len(db)
}
).rename({True: 'Training', False: 'Validation'})
```
- [x] Poportion by class in the set before spatially lagging and after --> `Fig. Balance (b)`
```
props = pandas.DataFrame(
{
'before_lag': pandas.value_counts(
db_all.query('single_class')['label']
) * 100 / len(db_all.query('single_class')),
'after_lag': pandas.value_counts(
db['label']
) * 100 / len(db['label'])
}
)
p = props.reindex(tools.class_names).plot.bar(figsize=(9, 3), title='Fig. Balance (b)')
p.set_xticklabels(p.get_xticklabels(), rotation = 45, ha="right");
```
## Single class models
Models to evaluate:
| Algorithm / Features | Baseline | Baseline + $WX$ | Baseline + $WX$ + $X\times WX$ |
|----------------------|----------|-----------------|-----------------|
| Max. Prob | `mp_res` | N/A | N/A |
| Logit ensemble | `logite_baseline_res` | `logite_wx_res` | Singular matrix |
| MNLogit | Does not converge | Does not converge | - |
| Random Forest | `rf_baseline_res` | `rf_wx_res` | N/A |
| XGBoost | `hbgb_baseline_res` | `hbgb_wx_res` | N/A |
Evaluation workflow:
1. Split train/validation randomly (e.g., 70/30)
1. Fit model on train subset
1. Obtain `perf_` measures on validation subset
1. Fill `results.json` file
Format:
```json
{
"meta_n_class": <int>,
"meta_class_names": <list>,
"meta_trainval_counts": <dict>
"model_name": <str>,
"model_params": <list>,
"meta_runtime": <float>,
"meta_preds_path": <str>,
"perf_model_accuracy_train": <float>,
"perf_within_class_accuracy_train": <list>,
"perf_confusion_train": <list>,
"perf_model_accuracy_val": <float>,
"perf_within_class_accuracy_val": <list>,
"perf_confusion_val": <list>,
"notes": <str>,
}
```
```
res_path = '/home/jovyan/data/model_outputs/'
```
Flush `db` to disk for availability later on:
```
db.to_parquet(os.path.join(res_path, 'db.pq'))
```
### Max. Probability
This is a baseline that selects as predicted category that which displays the highest probability as stored in `probs`. We can consider this the baseline upon which other models will improve.
```
mp_res = tools.run_maxprob(
tools.class_names, 'label', db, train_ids, val_ids, 'baseline', res_path
)
```
### Logit ensemble
Our approach here consists on fitting individual logistic regressions to predict each class against all other ones. With these models, we obtain a probability for each class, and pick that with the largest one as the predicted class.
---
**NOTE (DA-B)**: this might not be right if different logits have different distribution of probabilities. Maybe replace the probability by the rank and pick the largest rank as the predicted class?
---
Two methods, `logite_fit` to fit each class model, and `logite_predict` to generate predicted classes from set of arbitrary features.
#### Baseline
```
from importlib import reload
reload(tools);
logite_baseline_res = tools.run_logite(
tools.class_names, 'label', db, train_ids, val_ids, 'baseline', res_path
)
```
#### Baseline + $WX$
```
logite_wx_res = tools.run_logite(
tools.class_names + w_class_names,
'label',
db,
train_ids,
val_ids,
'baseline_wx',
res_path
)
```
#### Baseline + $WX$ + $X\times WX$
```
logite_xwx_res = tools.run_logite(
tools.class_names + w_class_names,
'label',
db,
train_ids,
val_ids,
'baseline_wx_xwx',
'model_results/',
scale_x=False,
log_x=True,
interact=(tools.class_names, w_class_names)
)
```
### Multinomial Logit
For this we rely on `statsmodels` [implementation](https://www.statsmodels.org/stable/examples/notebooks/generated/discrete_choice_overview.html#Multinomial-Logit) and fit it to the standardised log of the original probabilities to make convergence possible:
```
X_train = pandas.DataFrame(
scale(np.log1p(db.loc[train_ids, class_names+w_class_names])), train_ids, class_names+w_class_names
)
t0 = time()
mlogit_mod = sm.MNLogit(db.loc[train_ids, 'label'], X_train)
mlogit_res = mlogit_mod.fit()
from importlib import reload
reload(tools);
```
#### Baseline
```
mlogit_res = tools.run_mlogit(
tools.class_names, 'label', db, train_ids, val_ids, 'baseline', 'model_results/'
)
```
#### Baseline + $WX$
```
mlogit_wx = tools.run_mlogit(
tools.class_names + w_class_names,
'label',
db,
train_ids,
val_ids,
'baseline',
'model_results/'
)
```
### Random Forest
```
rf_param_grid = {
'n_estimators': [150],
'max_depth': [5],
'max_features': [0.25, 'sqrt']
}
rf_param_grid = {
'n_estimators': [50, 100, 150, 200, 300],
'max_depth': [5, 10, 20, 30, None],
'max_features': [0.25, 0.5, 0.75, 1, 'sqrt', 'log2']
}
```
#### Baseline
- Grid search
```
%%time
grid = GridSearchCV(
RandomForestClassifier(),
rf_param_grid,
scoring='accuracy',
cv=5,
n_jobs=-1
)
grid.fit(
db.query('train_all')[tools.class_names],
db.query('train_all')['label']
)
pandas.DataFrame(grid.cv_results_).to_csv(
os.path.join(res_path, 'RandomForestClassifier_baseline.csv')
)
print(grid.best_params_)
```
- Model fitting
```
rf_baseline_res = tools.run_tree(
tools.class_names,
'label',
db,
train_ids,
val_ids,
RandomForestClassifier(**grid.best_params_),
'baseline',
res_path
)
```
#### Baseline + $WX$
- Grid search
```
%%time
grid = GridSearchCV(
RandomForestClassifier(),
rf_param_grid,
scoring='accuracy',
cv=5,
n_jobs=-1
)
grid.fit(
db.query('train_all')[tools.class_names + w_class_names],
db.query('train_all')['label']
)
pandas.DataFrame(grid.cv_results_).to_csv(
os.path.join(res_path, 'RandomForestClassifier_baseline_wx.csv')
)
print(grid.best_params_)
```
- Model fitting
```
rf_wx_res = tools.run_tree(
tools.class_names + w_class_names,
'label',
db,
train_ids,
val_ids,
RandomForestClassifier(**grid.best_params_),
'baseline_wx',
res_path
)
```
### Histogram-Based Gradient Boosting
See [documentation](https://scikit-learn.org/stable/modules/ensemble.html#histogram-based-gradient-boosting) for the meaning and approaches to the hyper-parameters.
```
hbgb_param_grid = {
'max_iter': [50],
'learning_rate': [0.01, 0.05],
'max_depth': [30, None],
}
hbgb_param_grid = {
'max_iter': [50, 100, 150, 200, 300],
'learning_rate': [0.01, 0.05] + np.linspace(0, 1, 11)[1:].tolist(),
'max_depth': [5, 10, 20, 30, None],
}
```
#### Baseline
- Grid search:
```
%%time
grid = GridSearchCV(
HistGradientBoostingClassifier(),
hbgb_param_grid,
scoring='accuracy',
cv=5,
n_jobs=-1
)
grid.fit(
db.query('train_all')[tools.class_names], db.query('train_all')['label']
)
pandas.DataFrame(grid.cv_results_).to_csv(
os.path.join(res_path, 'HistGradientBoostingClassifier_baseline.csv')
)
print(grid.best_params_)
```
- Model fitting
```
hbgb_baseline_res = tools.run_tree(
tools.class_names,
'label',
db,
train_ids,
val_ids,
HistGradientBoostingClassifier(**grid.best_params_),
'baseline',
res_path
)
```
#### Baseline + $WX$
- Grid search:
```
%%time
grid = GridSearchCV(
HistGradientBoostingClassifier(),
hbgb_param_grid,
scoring='accuracy',
cv=5,
n_jobs=-1
)
grid.fit(
db.query('train_all')[tools.class_names + w_class_names],
db.query('train_all')['label']
)
pandas.DataFrame(grid.cv_results_).to_csv(
os.path.join(res_path, 'HistGradientBoostingClassifier_baseline_wx.csv')
)
print(grid.best_params_)
```
- Model fitting
```
hbgb_wx_res = tools.run_tree(
tools.class_names + w_class_names,
'label',
db,
train_ids,
val_ids,
HistGradientBoostingClassifier(max_iter=100),
'baseline_wx',
res_path
)
```
---
**NOTE** - As of April 21st'22, standard gradient-boosted trees are deprecated due to their long training time and similar performance with histogram-based ones.
### Gradient Tree Boosting
See [documentation](https://scikit-learn.org/stable/modules/ensemble.html#gradient-boosting) for the meaning and approaches to the hyper-parameters (mostly `n_estimators` and `learning_rate`).
#### Baseline
```
gbt_baseline_res = run_tree(
class_names,
'label',
db,
train_ids,
val_ids,
GradientBoostingClassifier(n_estimators=50, learning_rate=1.),
'baseline',
'model_results/'
)
```
#### Baseline + $WX$
```
gbt_baseline_res = run_tree(
class_names + w_class_names,
'label',
db,
train_ids,
val_ids,
GradientBoostingClassifier(n_estimators=50, learning_rate=1.),
'baseline_wx',
'model_results/'
)
```
| github_jupyter |
```
%%sh
pip -q install --upgrade pip
pip -q install sagemaker awscli boto3 --upgrade
from IPython.core.display import HTML
HTML("<script>Jupyter.notebook.kernel.restart()</script>")
```
# Direct Marketing with Amazon SageMaker AutoPilot
Last update: February 6th, 2020
```
import sagemaker
print (sagemaker.__version__)
sess = sagemaker.Session()
bucket = sess.default_bucket()
prefix = 'sagemaker/DEMO-automl-dm'
import numpy as np
import pandas as pd
%%sh
wget -N https://archive.ics.uci.edu/ml/machine-learning-databases/00222/bank-additional.zip
unzip -o bank-additional.zip
```
Let's read the CSV file into a Pandas data frame and take a look at the first few lines.
```
# https://pandas.pydata.org/pandas-docs/stable/generated/pandas.read_csv.html
data = pd.read_csv('./bank-additional/bank-additional-full.csv', sep=';')
pd.set_option('display.max_columns', 500) # Make sure we can see all of the columns
pd.set_option('display.max_rows', 50) # Keep the output on one page
data[:10] # Show the first 10 lines
data.shape # (number of lines, number of columns)
```
## Splitting the dataset
We split the dataset into training (95%) and test (5%) datasets. We will use the training dataset for AutoML, where it will be automatically split again for training and validation.
Once the model has been deployed, we'll use the test dataset to evaluate its performance.
```
# Set the seed to 123 for reproductibility
# https://pandas.pydata.org/pandas-docs/version/0.25/generated/pandas.DataFrame.sample.html
# https://docs.scipy.org/doc/numpy-1.15.1/reference/generated/numpy.split.html
train_data, test_data, _ = np.split(data.sample(frac=1, random_state=123),
[int(0.95 * len(data)), int(len(data))])
# Save to CSV files
train_data.to_csv('automl-train.csv', index=False, header=True) # Need to keep column names
test_data.to_csv('automl-test.csv', index=False, header=True)
!ls -l automl*.csv
```
**No preprocessing needed!** AutoML will take care of this, so let's just copy the training set to S3.
```
s3_input_data = sess.upload_data(path="automl-train.csv", key_prefix=prefix + "/input")
print(s3_input_data)
```
## Setting up the SageMaker AutoPilot job
After uploading the dataset to S3, we can invoke SageMaker AutoPilot to find the best ML pipeline to train a model on this dataset.
The required inputs for invoking a SageMaker AutoML job are the dataset location in S3, the name of the column of the dataset you want to predict (`y` in this case) and an IAM role.
```
from sagemaker.automl.automl import AutoML
# https://sagemaker.readthedocs.io/en/stable/automl.html
role = sagemaker.get_execution_role()
auto_ml_job = AutoML(
role = role, # IAM permissions for SageMaker
sagemaker_session = sess, #
target_attribute_name = 'y', # The column we want to predict
output_path = 's3://{}/{}/output'.format(bucket,prefix), # Save artefacts here
max_runtime_per_training_job_in_seconds = 600,
total_job_runtime_in_seconds = 3600
)
```
## Launching the SageMaker AutoPilot job
We can now launch the job by calling the `fit()` API.
```
auto_ml_job.fit(inputs=s3_input_data, logs=False, wait=False)
auto_ml_job.describe_auto_ml_job()
```
### Tracking the progress of the AutoPilot job
SageMaker AutoPilot job consists of four high-level steps :
* Data Preprocessing, where the dataset is split into train and validation sets.
* Recommending Pipelines, where the dataset is analyzed and SageMaker AutoPilot comes up with a list of ML pipelines that should be tried out on the dataset.
* Automatic Feature Engineering, where SageMaker AutoPilot performs feature transformation on individual features of the dataset as well as at an aggregate level.
* ML pipeline selection and hyperparameter tuning, where the top performing pipeline is selected along with the optimal hyperparameters for the training algorithm (the last stage of the pipeline).
```
from time import sleep
job = auto_ml_job.describe_auto_ml_job()
job_status = job['AutoMLJobStatus']
job_sec_status = job['AutoMLJobSecondaryStatus']
if job_status not in ('Stopped', 'Failed'):
while job_status in ('InProgress') and job_sec_status in ('AnalyzingData'):
sleep(30)
job = auto_ml_job.describe_auto_ml_job()
job_status = job['AutoMLJobStatus']
job_sec_status = job['AutoMLJobSecondaryStatus']
print (job_status, job_sec_status)
print("Data analysis complete")
```
## Viewing notebooks generated by SageMaker AutoPilot
Once data analysis is complete, SageMaker AutoPilot generates two notebooks:
* Data exploration,
* Candidate definition.
```
job = auto_ml_job.describe_auto_ml_job()
job_candidate_notebook = job['AutoMLJobArtifacts']['CandidateDefinitionNotebookLocation']
job_data_notebook = job['AutoMLJobArtifacts']['DataExplorationNotebookLocation']
print(job_candidate_notebook)
print(job_data_notebook)
```
Let's copy these two notebooks.
```
%%sh -s $job_candidate_notebook $job_data_notebook
aws s3 cp $1 .
aws s3 cp $2 .
```
Go back to the folder view, and open these notebooks. Lots of useful information in there!
SageMaker AutoPilot then launches feature engineering, and prepares different training and validation datasets.
```
job = auto_ml_job.describe_auto_ml_job()
job_status = job['AutoMLJobStatus']
job_sec_status = job['AutoMLJobSecondaryStatus']
if job_status not in ('Stopped', 'Failed'):
while job_status in ('InProgress') and job_sec_status in ('FeatureEngineering'):
sleep(30)
job = auto_ml_job.describe_auto_ml_job()
job_status = job['AutoMLJobStatus']
job_sec_status = job['AutoMLJobSecondaryStatus']
print (job_status, job_sec_status)
print("Feature engineering complete")
```
Once feature engineering is complete, SageMaker AutoPilot launches Automatic Model Tuning on the different candidates. While model tuning is running, we can explore its progress with SageMaker Experiments.
```
import pandas as pd
from sagemaker.analytics import ExperimentAnalytics
exp = ExperimentAnalytics(
sagemaker_session=sess,
experiment_name=job['AutoMLJobName'] + '-aws-auto-ml-job'
)
df = exp.dataframe()
print("Number of jobs: ", len(df))
# Move metric to first column
df = pd.concat([df['ObjectiveMetric - Max'], df.drop(['ObjectiveMetric - Max'], axis=1)], axis=1)
# Show top 5 jobs
df.sort_values('ObjectiveMetric - Max', ascending=0)[:5]
job = auto_ml_job.describe_auto_ml_job()
job_status = job['AutoMLJobStatus']
job_sec_status = job['AutoMLJobSecondaryStatus']
if job_status not in ('Stopped', 'Failed'):
while job_status in ('InProgress') and job_sec_status in ('ModelTuning'):
sleep(30)
job = auto_ml_job.describe_auto_ml_job()
job_status = job['AutoMLJobStatus']
job_sec_status = job['AutoMLJobSecondaryStatus']
print (job_status, job_sec_status)
print("Model tuning complete")
```
## Deploying the best candidate
Now that we have successfully completed the AutoML job on our dataset and visualized the trials, we can create a model from any of the trials with a single API call and then deploy that model for online or batch prediction using [Inference Pipelines](https://docs.aws.amazon.com/sagemaker/latest/dg/inference-pipelines.html). For this notebook, we deploy only the best performing trial for inference.
The best candidate is the one we're really interested in.
```
job_best_candidate = auto_ml_job.best_candidate()
print(job_best_candidate['CandidateName'])
print(job_best_candidate['FinalAutoMLJobObjectiveMetric'])
from time import strftime, gmtime
timestamp = strftime('%d-%H-%M-%S', gmtime())
endpoint_name = job['AutoMLJobName']+'-'+timestamp
auto_ml_job.deploy(
initial_instance_count = 1,
instance_type = 'ml.m4.xlarge',
endpoint_name = endpoint_name
)
```
## Scoring the best candidate
Let's predict and score the validation set. We'll compute metrics ourselves just for fun.
```
from sagemaker.predictor import csv_serializer, RealTimePredictor
from sagemaker.content_types import CONTENT_TYPE_CSV
predictor = RealTimePredictor(
endpoint=endpoint_name,
sagemaker_session=sess,
serializer=csv_serializer,
content_type=CONTENT_TYPE_CSV,
accept='text/csv'
)
import sys
tp = tn = fp = fn = count = 0
with open('automl-test.csv') as f:
lines = f.readlines()
for l in lines[1:]: # Skip header
l = l.split(',') # Split CSV line into feature array
label = l[-1] # Store 'yes'/'no' label
l = l[:-1] # Remove label
l = ','.join(l) # Rebuild CSV line without label
response = predictor.predict(l)
response = response.decode("utf-8")
#print ("label %s response %s" %(label,response))
if 'yes' in label:
# Sample is positive
if 'yes' in response:
# True positive
tp=tp+1
else:
# False negative
fn=fn+1
else:
# Sample is negative
if 'no' in response:
# True negative
tn=tn+1
else:
# False positive
fp=fp+1
count = count+1
if (count % 100 == 0):
sys.stdout.write(str(count)+' ')
print ("Done")
#Confusion matrix
print ("%d %d" % (tn, fp))
print ("%d %d" % (fn, tp))
accuracy = (tp+tn)/(tp+tn+fp+fn)
precision = tp/(tp+fp)
recall = tp/(tp+fn)
f1 = (2*precision*recall)/(precision+recall)
print ("%.4f %.4f %.4f %.4f" % (accuracy, precision, recall, f1))
```
## Deleting the endpoint
Once that we're done predicting, we can delete the endpoint (and stop paying for it).
```
sess.delete_endpoint(predictor.endpoint)
```
The SageMaker AutoML job creates many underlying artifacts such as dataset splits, preprocessing scripts, preprocessed data, etc. Let's delete them.
```
import boto3
job_outputs_prefix = '{}/output/{}'.format(prefix, job['AutoMLJobName'])
print(job_outputs_prefix)
s3_bucket =boto3.resource('s3').Bucket(bucket)
s3_bucket.objects.filter(Prefix=job_outputs_prefix).delete()
```
| github_jupyter |
# cuDF Cheat Sheets sample code
(c) 2020 NVIDIA, Blazing SQL
Distributed under Apache License 2.0
### Imports
```
import cudf
import numpy as np
import pandas as pd
```
### Sample DataFrame
```
# pandas
pandas_df = pd.DataFrame(
[
(39, 6.88, np.datetime64('2020-10-08T12:12:01'), np.timedelta64(14378,'s'), 'C', 'D', 'data'
, 'RAPIDS.ai is a suite of open-source libraries that allow you to run your end to end data science and analytics pipelines on GPUs.')
, (11, 4.21, None, None , 'A', 'D', 'cuDF'
, 'cuDF is a Python GPU DataFrame (built on the Apache Arrow columnar memory format)')
, (31, 4.71, np.datetime64('2020-10-10T09:26:43'), np.timedelta64(12909,'s'), 'U', 'D', 'memory'
, 'cuDF allows for loading, joining, aggregating, filtering, and otherwise manipulating tabular data using a DataFrame style API.')
, (40, 0.93, np.datetime64('2020-10-11T17:10:00'), np.timedelta64(10466,'s'), 'P', 'B', 'tabular'
, '''If your workflow is fast enough on a single GPU or your data comfortably fits in memory on
a single GPU, you would want to use cuDF.''')
, (33, 9.26, np.datetime64('2020-10-15T10:58:02'), np.timedelta64(35558,'s'), 'O', 'D', 'parallel'
, '''If you want to distribute your workflow across multiple GPUs or have more data than you can fit
in memory on a single GPU you would want to use Dask-cuDF''')
, (42, 4.21, np.datetime64('2020-10-01T10:02:23'), np.timedelta64(20480,'s'), 'U', 'C', 'GPUs'
, 'BlazingSQL provides a high-performance distributed SQL engine in Python')
, (36, 3.01, np.datetime64('2020-09-30T14:36:26'), np.timedelta64(24409,'s'), 'T', 'D', None
, 'BlazingSQL is built on the RAPIDS GPU data science ecosystem')
, (38, 6.44, np.datetime64('2020-10-10T08:34:36'), np.timedelta64(90171,'s'), 'X', 'B', 'csv'
, 'BlazingSQL lets you ETL raw data directly into GPU memory as a GPU DataFrame (GDF)')
, (17, 5.28, np.datetime64('2020-10-09T08:34:40'), np.timedelta64(30532,'s'), 'P', 'D', 'dataframes'
, 'Dask is a flexible library for parallel computing in Python')
, (10, 8.28, np.datetime64('2020-10-03T03:31:21'), np.timedelta64(23552,'s'), 'W', 'B', 'python'
, None)
]
, columns = ['num', 'float', 'datetime', 'timedelta', 'char', 'category', 'word', 'string']
)
pandas_df['category'] = pandas_df['category'].astype('category')
#cudf
cudf_df = cudf.DataFrame(
[
(39, 6.88, np.datetime64('2020-10-08T12:12:01'), np.timedelta64(14378,'s'), 'C', 'D', 'data'
, 'RAPIDS.ai is a suite of open-source libraries that allow you to run your end to end data science and analytics pipelines on GPUs.')
, (11, 4.21, None, None , 'A', 'D', 'cuDF'
, 'cuDF is a Python GPU DataFrame (built on the Apache Arrow columnar memory format)')
, (31, 4.71, np.datetime64('2020-10-10T09:26:43'), np.timedelta64(12909,'s'), 'U', 'D', 'memory'
, 'cuDF allows for loading, joining, aggregating, filtering, and otherwise manipulating tabular data using a DataFrame style API.')
, (40, 0.93, np.datetime64('2020-10-11T17:10:00'), np.timedelta64(10466,'s'), 'P', 'B', 'tabular'
, '''If your workflow is fast enough on a single GPU or your data comfortably fits in memory on
a single GPU, you would want to use cuDF.''')
, (33, 9.26, np.datetime64('2020-10-15T10:58:02'), np.timedelta64(35558,'s'), 'O', 'D', 'parallel'
, '''If you want to distribute your workflow across multiple GPUs or have more data than you can fit
in memory on a single GPU you would want to use Dask-cuDF''')
, (42, 4.21, np.datetime64('2020-10-01T10:02:23'), np.timedelta64(20480,'s'), 'U', 'C', 'GPUs'
, 'BlazingSQL provides a high-performance distributed SQL engine in Python')
, (36, 3.01, np.datetime64('2020-09-30T14:36:26'), np.timedelta64(24409,'s'), 'T', 'D', None
, 'BlazingSQL is built on the RAPIDS GPU data science ecosystem')
, (38, 6.44, np.datetime64('2020-10-10T08:34:36'), np.timedelta64(90171,'s'), 'X', 'B', 'csv'
, 'BlazingSQL lets you ETL raw data directly into GPU memory as a GPU DataFrame (GDF)')
, (17, 5.28, np.datetime64('2020-10-09T08:34:40'), np.timedelta64(30532,'s'), 'P', 'D', 'dataframes'
, 'Dask is a flexible library for parallel computing in Python')
, (10, 8.28, np.datetime64('2020-10-03T03:31:21'), np.timedelta64(23552,'s'), 'W', 'B', 'python'
, None)
]
, columns = ['num', 'float', 'datetime', 'timedelta', 'char', 'category', 'word', 'string']
)
cudf_df['category'] = cudf_df['category'].astype('category')
```
---
# Properties
---
## <span style="color:blue">DataFrame</span>
#### cudf.core.dataframe.DataFrame.at()
```
# pandas
pandas_df.at[2, 'string']
# cudf
cudf_df.at[2, 'string']
```
#### cudf.core.dataframe.DataFrame.columns()
```
# pandas
pandas_df.columns
#cudf
cudf_df.columns
```
#### cudf.core.dataframe.DataFrame.dtypes()
```
# pandas
pandas_df.dtypes
# cudf
cudf_df.dtypes
```
#### cudf.core.dataframe.DataFrame.iat()
```
# pandas
pandas_df.iat[2, 7]
# cudf
cudf_df.iat[2, 7]
```
#### cudf.core.dataframe.DataFrame.iloc()
```
# pandas
pandas_df.iloc[3]
# cudf
cudf_df.iloc[3]
pandas_df.iloc[3:5]
# pandas
pandas_df.iloc[2, 7]
# cudf
cudf_df.iloc[2, 7]
# pandas
pandas_df.iloc[2:5, 7]
# cudf
cudf_df.iloc[2:5, 7]
# pandas
pandas_df.iloc[2:5, [4,5,7]]
# cudf
cudf_df.iloc[2:5, [4,5,7]]
# pandas
pandas_df.iloc[[1,2,7], [4,5,6]]
# cudf
cudf_df.iloc[[1,2,7], [4,5,6]]
```
#### cudf.core.dataframe.DataFrame.index()
```
# pandas
pandas_df.index
# cudf
cudf_df.index
```
#### cudf.core.dataframe.DataFrame.loc()
```
# pandas
pandas_df.loc[3]
# cudf
cudf_df.loc[3]
# pandas
pandas_df.loc[3:6]
# cudf
cudf_df.loc[3:6]
# pandas
pandas_df.loc[2, 'string']
# cudf
cudf_df.loc[2, 'string']
# pandas
pandas_df.loc[3:6, ['string', 'float']]
# cudf
cudf_df.loc[3:6, ['string', 'float']]
```
#### cudf.core.dataframe.DataFrame.ndim()
```
# pandas
pandas_df.ndim
# cudf
cudf_df.ndim
```
#### cudf.core.dataframe.DataFrame.shape()
```
# pandas
pandas_df.shape
# cudf
cudf_df.shape
```
#### cudf.core.dataframe.DataFrame.size()
```
# pandas
pandas_df.size
# cudf
cudf_df.size
```
#### cudf.core.dataframe.DataFrame.T()
```
# pandas
pandas_df[['num']].T
# cudf
cudf_df[['num']].T
```
#### cudf.core.dataframe.DataFrame.values()
```
# pandas
pandas_df[['num', 'float']].values
# cudf
cudf_df[['num', 'float']].values
```
| github_jupyter |
# Housing prices and Machine Learning notebook with C# <img src ="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/207px-Jupyter_logo.svg.png" width="80px" alt="dotnet bot in space" align ="right"><img src ="https://user-images.githubusercontent.com/2546640/56708992-deee8780-66ec-11e9-9991-eb85abb1d10a.png" width="80px" alt="dotnet bot in space" align ="right">
### Add NuGet package references
```
#r "nuget:Microsoft.ML,1.4.0"
#r "nuget:Microsoft.ML.AutoML,0.16.0"
#r "nuget:Microsoft.Data.Analysis,0.1.0"
using Microsoft.Data.Analysis;
using XPlot.Plotly;
using Microsoft.AspNetCore.Html;
Formatter<DataFrame>.Register((df, writer) =>
{
var headers = new List<IHtmlContent>();
headers.Add(th(i("index")));
headers.AddRange(df.Columns.Select(c => (IHtmlContent) th(c.Name)));
var rows = new List<List<IHtmlContent>>();
var take = 20;
for (var i = 0; i < Math.Min(take, df.RowCount); i++)
{
var cells = new List<IHtmlContent>();
cells.Add(td(i));
foreach (var obj in df[i])
{
cells.Add(td(obj));
}
rows.Add(cells);
}
var t = table(
thead(
headers),
tbody(
rows.Select(
r => tr(r))));
writer.Write(t);
}, "text/html");
using System.IO;
using System.Net.Http;
string housingPath = "housing.csv";
if (!File.Exists(housingPath))
{
var contents = new HttpClient()
.GetStringAsync("https://raw.githubusercontent.com/ageron/handson-ml2/master/datasets/housing/housing.csv").Result;
File.WriteAllText("housing.csv", contents);
}
var housingData = DataFrame.LoadCsv(housingPath);
housingData
housingData.Description()
Chart.Plot(
new Graph.Histogram()
{
x = housingData["median_house_value"],
nbinsx = 20
}
)
var chart = Chart.Plot(
new Graph.Scattergl()
{
x = housingData["longitude"],
y = housingData["latitude"],
mode = "markers",
marker = new Graph.Marker()
{
color = housingData["median_house_value"],
colorscale = "Jet"
}
}
);
chart.Width = 600;
chart.Height = 600;
display(chart);
static T[] Shuffle<T>(T[] array)
{
Random rand = new Random();
for (int i = 0; i < array.Length; i++)
{
int r = i + rand.Next(array.Length - i);
T temp = array[r];
array[r] = array[i];
array[i] = temp;
}
return array;
}
int[] randomIndices = Shuffle(Enumerable.Range(0, (int)housingData.RowCount).ToArray());
int testSize = (int)(housingData.RowCount * .1);
int[] trainRows = randomIndices[testSize..];
int[] testRows = randomIndices[..testSize];
DataFrame housing_train = housingData[trainRows];
DataFrame housing_test = housingData[testRows];
display(housing_train.RowCount);
display(housing_test.RowCount);
using Microsoft.ML;
using Microsoft.ML.Data;
using Microsoft.ML.AutoML;
#!time
var mlContext = new MLContext();
var experiment = mlContext.Auto().CreateRegressionExperiment(maxExperimentTimeInSeconds: 15);
var result = experiment.Execute(housing_train, labelColumnName:"median_house_value");
var scatters = result.RunDetails.Where(d => d.ValidationMetrics != null).GroupBy(
r => r.TrainerName,
(name, details) => new Graph.Scattergl()
{
name = name,
x = details.Select(r => r.RuntimeInSeconds),
y = details.Select(r => r.ValidationMetrics.MeanAbsoluteError),
mode = "markers",
marker = new Graph.Marker() { size = 12 }
});
var chart = Chart.Plot(scatters);
chart.WithXTitle("Training Time");
chart.WithYTitle("Error");
display(chart);
Console.WriteLine($"Best Trainer:{result.BestRun.TrainerName}");
var testResults = result.BestRun.Model.Transform(housing_test);
var trueValues = testResults.GetColumn<float>("median_house_value");
var predictedValues = testResults.GetColumn<float>("Score");
var predictedVsTrue = new Graph.Scattergl()
{
x = trueValues,
y = predictedValues,
mode = "markers",
};
var maximumValue = Math.Max(trueValues.Max(), predictedValues.Max());
var perfectLine = new Graph.Scattergl()
{
x = new[] {0, maximumValue},
y = new[] {0, maximumValue},
mode = "lines",
};
var chart = Chart.Plot(new[] {predictedVsTrue, perfectLine });
chart.WithXTitle("True Values");
chart.WithYTitle("Predicted Values");
chart.WithLegend(false);
chart.Width = 600;
chart.Height = 600;
display(chart);
```
| github_jupyter |
```
from functools import partial
import itertools
import json
from pathlib import Path
import re
import sys
sys.path.append("../src")
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
import statsmodels.formula.api as smf
from tqdm import tqdm, tqdm_notebook
%matplotlib inline
sns.set(style="whitegrid", context="paper", font_scale=3.5, rc={"lines.linewidth": 2.5})
from IPython.display import set_matplotlib_formats
set_matplotlib_formats('png')
#set_matplotlib_formats('svg')
%load_ext autoreload
%autoreload 2
import util
```
## Data preparation
```
output_path = Path("../output")
bert_encoding_path = output_path / "encodings"
sprobe_results_path = output_path / "structural-probe"
checkpoints = [util.get_encoding_ckpt_id(dir_entry) for dir_entry in bert_encoding_path.iterdir()]
models = [model for model, _, _ in checkpoints]
baseline_model = "baseline"
if baseline_model not in models:
raise ValueError("Missing baseline model. This is necessary to compute performance deltas in the analysis of fine-tuning models. Stop.")
standard_models = [model for model in models if not model.startswith("LM_") and not model == baseline_model]
custom_models = [model for model in models if model.startswith("LM_") and not model == baseline_model]
runs = sorted(set(run for _, run, _ in checkpoints))
checkpoint_steps = sorted(set(step for _, _, step in checkpoints))
# Models which should appear in the final report figures
report_models = ["SQuAD", "QQP", "MNLI", "SST", "LM", "LM_scrambled", "LM_scrambled_para", "LM_pos", "glove"]
# Model subsets to render in different report figures
report_model_sets = [
("all", set(report_models)),
("standard", set(report_models) & set(standard_models)),
("custom", set(report_models) & set(custom_models)),
]
report_model_sets = [(name, model_set) for name, model_set in report_model_sets
if len(model_set) > 0]
RENDER_FINAL = True
figure_path = Path("../reports/figures")
figure_path.mkdir(exist_ok=True)
report_hues = dict(zip(sorted(report_models), sns.color_palette()))
```
## Collect results
```
eval_results = {}
for eval_dir in tqdm_notebook(list(sprobe_results_path.iterdir())):
if not eval_dir.is_dir(): continue
model, run, step = util.get_encoding_ckpt_id(eval_dir)
try:
uuas_file = list(eval_dir.glob("**/dev.uuas"))[0]
with uuas_file.open("r") as f:
uuas = float(f.read().strip())
except: continue
try:
spearman_file = list(eval_dir.glob("**/dev.spearmanr-*-mean"))[0]
with spearman_file.open("r") as f:
spearman = float(f.read().strip())
except: continue
eval_results[model, run, step] = pd.Series({"uuas": uuas, "spearman": spearman})
```
### Add non-BERT results
```
nonbert_models = []
# GloVe
# for glove_dir in tqdm_notebook(list(sprobe_glove_path.glob("*"))):
# if not glove_dir.is_dir(): continue
# model = glove_dir.name
# try:
# uuas_file = list(glove_dir.glob("**/dev.uuas"))[0]
# with uuas_file.open("r") as f:
# uuas = float(f.read().strip())
# except: continue
# try:
# spearman_file = list(glove_dir.glob("**/dev.spearmanr-*-mean"))[0]
# with spearman_file.open("r") as f:
# spearman = float(f.read().strip())
# except: continue
# nonbert_models.append(model)
# eval_results[model, 1, 250, 0] = pd.Series({"uuas": uuas, "spearman": spearman})
```
### Aggregate
```
eval_results = pd.DataFrame(pd.concat(eval_results, names=["model", "run", "step", "metric"]))
eval_results.tail(20)
# Only use spaCy results
nonbert_models_to_graph = [("spaCy-en_vectors_web_lg", "GloVe")]
nonbert_models_to_graph = [(name, label) for name, label in nonbert_models_to_graph if name in nonbert_models]
```
## Graph
```
graph_data = eval_results.reset_index()
graph_data = graph_data[~graph_data.model.isin(nonbert_models + [baseline_model])]
g = sns.FacetGrid(data=graph_data, col="metric", height=7, sharex=True, sharey=True)
g.map(sns.lineplot, "step", 0, "model")
for uuas_ax in g.axes[:, 0]:
for nonbert_model, label in nonbert_models_to_graph:
uuas_ax.axhline(eval_results.loc[nonbert_model, 1, 250, 0, "uuas"][0], linestyle='--', label=label)
for spearman_ax in g.axes[:, 1]:
for nonbert_model, label in nonbert_models_to_graph:
spearman_ax.axhline(eval_results.loc[nonbert_model, 1, 250, 0, "spearman"][0], linestyle='--', label=label)
g.add_legend()
g
g = sns.FacetGrid(data=graph_data, col="metric", row="model", height=7, sharex=True, sharey=True)
g.map(sns.lineplot, "step", 0).add_legend()
%matplotlib agg
if RENDER_FINAL:
dir = figure_path / "structural_probe"
dir.mkdir(exist_ok=True)
for metric, label in [("uuas", "UUAS"), ("spearman", "Spearman correlation")]:
fig = plt.figure(figsize=(15, 9))
ax = sns.lineplot(data=graph_data[(graph_data.metric == metric)], x="step", y=0,
hue="model", palette=report_hues)
for nonbert_model, nonbert_label in nonbert_models_to_graph:
ax.axhline(eval_results.loc[nonbert_model, 1, 0, metric][0],
linestyle='--', label=nonbert_label, linewidth=3)
plt.legend(loc="center left", bbox_to_anchor=(1, 0.5))
plt.xlim((0, checkpoint_steps[-1]))
plt.ylabel(label)
plt.xlabel("Training step")
plt.tight_layout()
plt.savefig(dir / ("%s.pdf" % metric))
plt.close()
%matplotlib inline
```
| github_jupyter |
# Exploratory Data Science (EDS) at scale with Dask
```
!pip install --upgrade "dask-cloudprovider[azure]" dask-lightgbm lightgbm
from azureml.core import Workspace
ws = Workspace.from_config()
ws
import git
from pathlib import Path
# get root of git repo
prefix = Path(git.Repo(".", search_parent_directories=True).working_tree_dir)
prefix
from azureml.core import Environment
from dask.distributed import Client
from dask_cloudprovider import AzureMLCluster
env = Environment.from_conda_specification(
"dask-tutorial", prefix.joinpath("environments", "dask.yml")
)
cluster = AzureMLCluster(
ws,
vm_size="STANDARD_DS5_V2",
environment_definition=env,
initial_node_count=60,
scheduler_idle_timeout=1200,
)
c = Client(cluster)
c
container_name = "isdweatherdatacontainer"
storage_options = {"account_name": "azureopendatastorage"}
from adlfs import AzureBlobFileSystem
fs = AzureBlobFileSystem(**storage_options)
fs
%%time
files = fs.glob(f"{container_name}/ISDWeather/year=*/month=*/*.parquet")
files = [f"az://{file}" for file in files]
len(files)
files[-5:]
%%time
import dask.dataframe as dd
npartitions = 256
engine = "pyarrow"
blocksize = "1GB"
ddf = dd.read_parquet(
files, storage_options=storage_options, engine=engine, blocksize=blocksize
).repartition(npartitions=npartitions)
ddf = ddf.set_index(
dd.to_datetime(ddf.datetime).dt.floor("d"), sorted=False
).persist()
ddf
%%time
len(ddf)
%%time
len(ddf)
%%time
gbs = round(ddf.memory_usage(index=True, deep=True).sum().compute() / 1e9, 2)
print(f"ddf is {gbs} GBs")
```
## EDA
```
%%time
ddf.describe().compute()
%%time
places = (
ddf.groupby(ddf.index)[["longitude", "latitude", "year"]].mean().compute()
)
import matplotlib.pyplot as plt
plt.figure(figsize=(16, 16))
plt.scatter(places.longitude, places.latitude, c=places.year)
plt.title("Lat/long")
plt.xlabel("Longitude")
plt.ylabel("Latitude")
plt.grid()
plt.colorbar()
plt.figure(figsize=(16, 16))
plt.scatter(places.longitude, places.latitude, c=places.year)
plt.title("Lat/long")
plt.xlabel("Longitude")
plt.ylabel("Latitude")
plt.xlim([-50, -30]) # zoom in
plt.ylim([35, 40]) # zoom in
plt.grid()
plt.colorbar()
%%time
means = ddf.groupby(ddf.index).mean().compute()
means.head()
from datetime import datetime
for col in list(means.columns):
fig = plt.figure(figsize=(16, 8))
# plt.style.use('dark_background')
means[col].plot(color="b")
plt.title("Average of {}".format(col))
plt.xlim([datetime(2008, 1, 1), datetime(2021, 1, 1)])
plt.grid()
```
## Process and persist
```
ddf["temperature"] = ddf["temperature"] * (9 / 5) + 32
ds = ws.get_default_datastore()
container_name = ds.container_name
storage_options = {
"account_name": ds.account_name,
"account_key": ds.account_key,
}
%%time
# ddf.to_csv("az://{container_name}/data/dask/isd", storage_options=storage_options)
```
## Prepare data and train LightGBM model
```
ddf = ddf.fillna(0)
cols = list(ddf.columns)
cols = [
col
for col in cols
if ddf.dtypes[col] != "object"
and col not in ["version", "datetime", "temperature"]
]
cols
X = ddf[cols].persist()
y = ddf.temperature.persist()
%%time
from dask_lightgbm import LGBMRegressor
params = {
"n_estimators": 64,
"num_iterations": 100,
"boosting_type": "gbdt",
"num_leaves": 31,
"max_depth": -1,
"learning_rate": 0.1,
"tree_learner": "data",
}
lgbm = LGBMRegressor(**params)
lgbm.fit(X, y)
%%time
y_pred = lgbm.predict(X)
%%time
rmse = ((((y.to_dask_array() - y_pred) ** 2).mean()) ** 0.5).compute()
print(f"Training RMSE: {round(rmse, 3)}")
```
## Close Cluster and Client
```
cluster.close()
c.close()
```
| github_jupyter |
# 第三讲 乘法和逆矩阵
## 矩阵乘法
矩阵乘法是指两个矩阵相乘得到一个新的矩阵,可表示为 $A * B = C$,如果矩阵 $A$ 为 $m * n$,矩阵 $B$ 为 $n * p$,则得到的矩阵 $C$ 为 $m * p$,注意两个矩阵可以相乘的条件是第一个矩阵的列数要等于第二个矩阵的行数。
计算矩阵乘法有多种方法,首先是**常规法**(Regular Way),逐个计算结果矩阵中的每一项 $C_{i,j} = \sum^{n}_{k} A_{i,k} * B_{k,j}$。
第二个方法是**整行相乘**(the Row Way),因为矩阵 $C$ 的行是矩阵 $B$ 行的线性组合。假设现有 $3 * 3$ 的矩阵 $A$ 乘以 $3 * 4$ 的矩阵 $B$,则计算过程为:$$\begin{bmatrix} a_{1,1} & a_{1,2} & a_{1,3} \\ b_{2,1} & b_{2,2} & b_{2,3} \\ c_{3,1} & c_{3,2} & c_{3,3} \end{bmatrix} \begin{bmatrix} row_1\\ row_2 \\ row_3 \end{bmatrix} = \begin{bmatrix} a_{1,1} * row_1 + a_{1,2} * row_2 + a_{1,3} * row_3 \\ a_{2,1} * row_1 + a_{2,2} * row_2 + a_{2,3} * row_3 \\ a_{3,1} * row_1 + a_{3,2} * row_2 + a_{3,3} * row_3 \end{bmatrix}$$
第三个方法是**整列相乘**(the Column Way),因为矩阵 $C$ 的列是矩阵 $A$ 列的线性组合。同样假设有 $3 * 3$ 的矩阵 $A$ 乘以 $3 * 4$ 的矩阵 $B$,则计算过程为:$$\begin{bmatrix} col_1 & col_2 & col_3\end{bmatrix}\begin{bmatrix} b_{1,1} & b_{1,2} & b_{1,3} & b_{1,4} \\ b_{2,1} & b_{2,2} & b_{2,3} & b_{2,4} \\ b_{3,1} & b_{3,2} & b_{3,3} & b_{3,4}\end{bmatrix} = \begin{bmatrix} col_1b_{1,1} & col_1b_{1,2} & col_1b_{1,3} & col_1b_{1,4} \\ + & + & + & + \\ col_2b_{2,1} & col_2b_{2,2} & col_2b_{2,3} & col_2b_{2,4}\\ + & + & + & + \\ col_3b_{3,1} & col_3b_{3,2} & col_3b_{3,3} & col_3b_{3,4}\end{bmatrix}$$
第四个方法为**列行相乘**(Columns Times Rows Way),一个 $3 * 1$ 的列乘以一个 $1 * 4$ 的行会得到一个 $3 * 4$ 的矩阵,如果把矩阵 $A$ 看成三个 $3 * 1$ 的列,将矩阵 $B$ 看成三个 $1 * 4$ 的行,则计算过程有:$$\begin{bmatrix} col_1 & col_2 & col_3 \end{bmatrix} \begin{bmatrix} row_1 \\ row_2 \\ row_3 \end{bmatrix} = col_1row_1 + col_2row_2 + col_3row_3$$
最后一个方法为**分块法**(Block Multiplication),将矩阵 $A$ 和矩阵 $B$ 分成若干的小相等的矩阵,然后按照常规法的规则相乘后相加:$$\begin{bmatrix} A_1 & A_2 \\ A_3 & A_4 \end{bmatrix} \begin{bmatrix} B_1 & B_2 \\ B_3 & B_4 \end{bmatrix} = \begin{bmatrix} A_1 * B_1 + A_2 * B_3 & A_1 * B_2 + A_2 * B_4\\ A_3 * B_1 + A_4 * B_3 & A_3 * B_2 + A_4 * B_4\end{bmatrix}$$
## 逆矩阵
**逆矩阵**(Inverses Matrix),给定一个 $n$ 阶方阵 $A$,如果存在一个矩阵 $B$ 使得 $A*B = B*A = I$,其中矩阵 $I$ 为一个 $n$ 阶单位矩阵,则称矩阵 $B$ 为矩阵 $A$ 的逆矩阵,记作 $A^{-1}$。注意,$n$ 阶**单位矩阵**(Identity Matrix)是一个 $n*n$ 且对角线元素全为 $1$,其它元素均为 $0$ 的方阵。我们一般考虑方阵的逆矩阵,但不是所有的方阵都有逆矩阵。对于有逆矩阵的方阵而言,左逆是和右逆相等的。对于有逆矩阵的非方阵而言,左逆和右逆是不相等的。
这里引入一个概念为奇异矩阵(Singular Matrix)和非奇异矩阵(Unsingular Matrix),奇异矩阵为**行列式**不为零的方阵(后面会讲到行列式的求法),对于非方阵而言不考虑矩阵的奇异性。而对于非奇异矩阵而言,是没有逆矩阵的。
考虑方阵没有逆矩阵的情况,如果存在一个向量 $x$ 使得 $Ax=0$,那么矩阵 $A$ 则没有逆矩阵。证明:$Ax = 0 \Rightarrow A^{-1} (Ax) = A^{-1} * 0 = 0 \Rightarrow Ix = 0$,但是任何矩阵乘以单位矩阵都等于其本身,即 $Ix = x$。
### 高斯-若尔当
高斯诺尔当(Gaussian Jordan)方法可以一次求解多个方程组,也可以来求解方阵的逆矩阵。首先我们考虑求解矩阵 $A = \begin{bmatrix} 1 & 3 \\ 2 & 7 \end{bmatrix}$ 的逆矩阵,设 $A^{-1} = \begin{bmatrix} a & c \\ b & d \end{bmatrix}$,则 $\begin{bmatrix} 1 & 3 \\ 2 & 7 \end{bmatrix} \begin{bmatrix} a & c \\ b & d \end{bmatrix} = \begin{bmatrix} 1 & 0 \\ 0 & 1 \end{bmatrix}$。这里我们可以将它看成是两个方程组:$\begin{bmatrix} 1 & 3 \\ 2 & 7 \end{bmatrix} \begin{bmatrix} a \\ b \end{bmatrix} = \begin{bmatrix} 1 \\ 0 \end{bmatrix}$ 和 $\begin{bmatrix} 1 & 3 \\ 2 & 7 \end{bmatrix} \begin{bmatrix} c \\ d \end{bmatrix} = \begin{bmatrix} 0 \\ 1 \end{bmatrix}$。
根据高斯诺尔当方法,我们将两个矩阵合并后进行先正向消元:$\left[\begin{array}{cc|cc}1&3&1&0\\2&7&0&1\end{array}\right] \xrightarrow{row_2-2row_1}\left[\begin{array}{cc|cc}1&3&1&0\\0&1&-2&1\end{array}\right]$,再进行逆向消元,即消除第一个方程中的第二个主元:$\left[\begin{array}{cc|cc}1&3&1&0\\0&1&-2&1\end{array}\right]\xrightarrow{row_1-3row_2}\left[\begin{array}{cc|cc}1&0&7&-3\\0&1&-2&1\end{array}\right]$。此时,合并矩阵左边的矩阵 $A$ 变成了单位矩阵,而右边的单位矩阵矩阵则变成了矩阵 $A$ 的逆,即 $A^{-1} = \begin{bmatrix}7&3\\1&-2\end{bmatrix}$。证明:$[ A\ |\ I] \Rightarrow [ A*A^{-1}\ |\ I*A^{-1}] \Rightarrow [ I\ |\ A^{-1}]$。
### 矩阵逆的性质
如果矩阵 $C$ 是矩阵 $A$ 和矩阵 $B$ 的乘积,那么矩阵 $C$ 的逆等于矩阵 $B$ 的逆乘以矩阵 $A$ 的逆:$(AB)^{-1} = B^{-1}A^{-1}$。证明,$A * B * B^{-1} * A^{-1} = A * A^{-1} = I$。
矩阵 $A$ 转置的逆等于矩阵 $A$ 的逆的转置:$(A^T)^{-1} = (A^{-1})^T$。证明,$I^T = (A * A^{-1})^T \Rightarrow (A^{-1})^TA^T = I$。
一个矩阵的逆的逆等于它本身:$A = (A^{-1})^{-1}$。
| github_jupyter |
<!--BOOK_INFORMATION-->
<img align="left" style="padding-right:10px;" src="fig/cover-small.jpg">
*This notebook contains an excerpt from the [Whirlwind Tour of Python](http://www.oreilly.com/programming/free/a-whirlwind-tour-of-python.csp) by Jake VanderPlas; the content is available [on GitHub](https://github.com/jakevdp/WhirlwindTourOfPython).*
*The text and code are released under the [CC0](https://github.com/jakevdp/WhirlwindTourOfPython/blob/master/LICENSE) license; see also the companion project, the [Python Data Science Handbook](https://github.com/jakevdp/PythonDataScienceHandbook).*
<!--NAVIGATION-->
< [List Comprehensions](11-List-Comprehensions.ipynb) | [Contents](Index.ipynb) | [Strings and Regular Expressions](13-Strings-and-Regular-Expressions.ipynb) >
# Generators
Here we'll take a deeper dive into Python generators, including *generator expressions* and *generator functions*.
## Generator Expressions
The difference between list comprehensions and generator expressions is sometimes confusing; here we'll quickly outline the differences between them:
### List comprehensions use square brackets, while generator expressions use parentheses
This is a representative list comprehension:
```
[n ** 2 for n in range(12)]
```
While this is a representative generator expression:
```
(n ** 2 for n in range(12))
```
Notice that printing the generator expression does not print the contents; one way to print the contents of a generator expression is to pass it to the ``list`` constructor:
```
G = (n ** 2 for n in range(10))
list(G)
```
Can you think of another possibility to print the element of a generator?
### A list is a collection of values, while a generator is a recipe for producing values
When you create a list, you are actually building a collection of values, and there is some memory cost associated with that.
When you create a generator, you are not building a collection of values, but a recipe for producing those values.
Both expose the same iterator interface, as we can see here:
```
L = [n ** 2 for n in range(12)]
for val in L:
print(val, end=' ')
G = (n ** 2 for n in range(12))
for val in G:
print(val, end=' ')
```
The difference is that a generator expression does not actually compute the values until they are needed.
This not only leads to memory efficiency, but to computational efficiency as well!
This also means that while the size of a list is limited by available memory, the size of a generator expression is unlimited!
An example of an infinite generator expression can be created using the ``count`` iterator defined in ``itertools``:
```
from itertools import count
count()
for i in count():
print(i, end=' ')
if i >= 10: break
```
The ``count`` iterator will go on happily counting forever until you tell it to stop; this makes it convenient to create generators that will also go on forever:
```
factors = [2, 3, 5, 7]
G = (i for i in count() if all(i % n > 0 for n in factors))
for val in G:
print(val, end=' ')
if val > 40: break
```
You might see what we're getting at here: if we were to expand the list of factors appropriately, what we would have the beginnings of is a prime number generator, using the Sieve of Eratosthenes algorithm. We'll explore this more momentarily.
### A list can be iterated multiple times; a generator expression is single-use
This is one of those potential gotchas of generator expressions.
With a list, we can straightforwardly do this:
```
L = [n ** 2 for n in range(12)]
for val in L:
print(val, end=' ')
print()
for val in L:
print(val, end=' ')
```
A generator expression, on the other hand, is used-up after one iteration:
```
G = (n ** 2 for n in range(12))
list(G)
list(G)
```
This can be very useful because it means iteration can be stopped and started:
```
G = (n**2 for n in range(12))
for n in G:
print(n, end=' ')
if n > 30: break
print("\ndoing something in between")
for n in G:
print(n, end=' ')
```
One place I've found this useful is when working with collections of data files on disk; it means that you can quite easily analyze them in batches, letting the generator keep track of which ones you have yet to see.
## Generator Functions: Using ``yield``
We saw in the previous section that list comprehensions are best used to create relatively simple lists, while using a normal ``for`` loop can be better in more complicated situations.
The same is true of generator expressions: we can make more complicated generators using *generator functions*, which make use of the ``yield`` statement.
Here we have two ways of constructing the same list:
```
L1 = [n ** 2 for n in range(12)]
L2 = []
for n in range(12):
L2.append(n ** 2)
print(L1)
print(L2)
```
Similarly, here we have two ways of constructing equivalent generators:
```
G1 = (n ** 2 for n in range(12))
def gen():
for n in range(12):
yield n ** 2
G2 = gen()
print(*G1)
print(*G2)
```
A generator function is a function that, rather than using ``return`` to return a value once, uses ``yield`` to yield a (potentially infinite) sequence of values.
Just as in generator expressions, the state of the generator is preserved between partial iterations, but if we want a fresh copy of the generator we can simply call the function again.
## Example: Prime Number Generator
Here I'll show my favorite example of a generator function: a function to generate an unbounded series of prime numbers.
A classic algorithm for this is the *Sieve of Eratosthenes*, which works something like this:
```
# Generate a list of candidates
L = [n for n in range(2, 40)]
print(L)
# Remove all multiples of the first value
L = [n for n in L if n == L[0] or n % L[0] > 0]
print(L)
# Remove all multiples of the second value
L = [n for n in L if n == L[1] or n % L[1] > 0]
print(L)
# Remove all multiples of the third value
L = [n for n in L if n == L[2] or n % L[2] > 0]
print(L)
```
If we repeat this procedure enough times on a large enough list, we can generate as many primes as we wish.
Let's encapsulate this logic in a generator function:
```
def gen_primes(N):
"""Generate primes up to N"""
primes = set()
for n in range(2, N):
if all(n % p > 0 for p in primes):
primes.add(n)
yield n
print(*gen_primes(100))
```
That's all there is to it!
While this is certainly not the most computationally efficient implementation of the Sieve of Eratosthenes, it illustrates how convenient the generator function syntax can be for building more complicated sequences.
<!--NAVIGATION-->
< [List Comprehensions](11-List-Comprehensions.ipynb) | [Contents](Index.ipynb) | [Strings and Regular Expressions](13-Strings-and-Regular-Expressions.ipynb) >
| github_jupyter |
**[Intermediate Machine Learning Home Page](https://www.kaggle.com/learn/intermediate-machine-learning)**
---
In this exercise, you will leverage what you've learned to tune a machine learning model with **cross-validation**.
# Setup
The questions below will give you feedback on your work. Run the following cell to set up the feedback system.
```
# set up code checking
import os
if not os.path.exists("../input/train.csv"):
os.symlink("../input/home-data-for-ml-course/train.csv", "../input/train.csv")
os.symlink("../input/home-data-for-ml-course/test.csv", "../input/test.csv")
from learntools.core import binder
binder.bind(globals())
from learntools.ml_intermediate.ex5 import *
print("Setup Complete")
```
You will work with the [Housing Prices Competition for Kaggle Learn Users](https://www.kaggle.com/c/home-data-for-ml-course) from the previous exercise.

Run the next code cell without changes to load the training and validation sets in `X_train`, `X_valid`, `y_train`, and `y_valid`. The test set is loaded in `X_test`.
For simplicity, we drop categorical variables.
```
from sklearn.model_selection import train_test_split
# load data
import pandas as pd
train_data = pd.read_csv('../input/train.csv', index_col='Id')
test_data = pd.read_csv('../input/test.csv', index_col='Id')
# remove rows with missing target, separate target from predictors
train_data.dropna(axis=0, subset=['SalePrice'], inplace=True)
y = train_data['SalePrice']
train_data.drop(['SalePrice'], axis=1, inplace=True)
# select numeric columns only
numeric_cols = [cname for cname in train_data.columns if train_data[cname].dtype in ['int64', 'float64']]
X = train_data[numeric_cols].copy()
X_test = test_data[numeric_cols].copy()
```
Use the next code cell to print the first several rows of the data.
```
X.head()
```
So far, you've learned how to build pipelines with scikit-learn. For instance, the pipeline below will use [`SimpleImputer()`](https://scikit-learn.org/stable/modules/generated/sklearn.impute.SimpleImputer.html) to replace missing values in the data, before using [`RandomForestRegressor()`](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.RandomForestRegressor.html) to train a random forest model to make predictions. We set the number of trees in the random forest model with the `n_estimators` parameter, and setting `random_state` ensures reproducibility.
```
from sklearn.pipeline import Pipeline
from sklearn.impute import SimpleImputer
from sklearn.ensemble import RandomForestRegressor
my_pipeline = Pipeline(steps=[
('preprocessor', SimpleImputer()),
('model', RandomForestRegressor(n_estimators=50, random_state=0))
])
```
You have also learned how to use pipelines in cross-validation. The code below uses the [`cross_val_score()`](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.cross_val_score.html) function to obtain the mean absolute error (MAE), averaged across five different folds. Recall we set the number of folds with the `cv` parameter.
```
from sklearn.model_selection import cross_val_score
# multiply by -1 since sklearn calculates *negative* MAE
scores = -1 * cross_val_score(my_pipeline, X, y,
cv=5,
scoring='neg_mean_absolute_error')
print("Average MAE score:", scores.mean())
```
<hr/>
# Step 1: Write a useful function
In this exercise, you'll use cross-validation to select parameters for a machine learning model.
Begin by writing a function `get_score()` that reports the average (over three cross-validation folds) MAE of a machine learning pipeline that uses:
- the data in `X` and `y` to create folds,
- `SimpleImputer()` (with all parameters left as default) to replace missing values, and
- `RandomForestRegressor()` (with `random_state=0`) to fit a random forest model.
The `n_estimators` parameter supplied to `get_score()` is used when setting the number of trees in the random forest model.
```
def get_score(n_estimators):
"""Return the average MAE over 3 CV folds of random forest model.
Keyword argument:
n_estimators -- the number of trees in the forest
"""
# define pipeline
my_pipeline = Pipeline(steps=[
('preprocessor', SimpleImputer()),
('model', RandomForestRegressor(n_estimators=n_estimators, random_state=0))
])
# evaluate model
scores = -1 * cross_val_score(my_pipeline, X, y,
cv=3,
scoring='neg_mean_absolute_error')
# return average of evalution scores
return scores.mean()
# check your answer
step_1.check()
# lines below will give you a hint or solution code
# step_1.hint()
# step_1.solution()
```
<hr/>
# Step 2: Test different parameter values
Now, you will use the function that you defined in Step 1 to evaluate the model performance corresponding to eight different values for the number of trees in the random forest: 50, 100, 150, ..., 300, 350, 400.
Store your results in a Python dictionary `results`, where `results[i]` is the average MAE returned by `get_scores(i)`.
```
results = {i: get_score(i) for i in range(50, 401, 50)}
# check your answer
step_2.check()
# lines below will give you a hint or solution code
# step_2.hint()
# step_2.solution()
```
<hr/>
# Step 3: Find the best parameter value
Use the next cell to visualize your results from Step 2. Run the code without changes.
```
import matplotlib.pyplot as plt
%matplotlib inline
plt.plot(results.keys(), results.values())
plt.show()
```
Given the results, which value for `n_estimators` seems best for the random forest model? Use your answer to set the value of `n_estimators_best`.
```
n_estimators_best = 200
# check your answer
step_3.check()
# lines below will give you a hint or solution code
# step_3.hint()
# step_3.solution()
```
In this exercise, you have explored one method for choosing appropriate parameters in a machine learning model.
If you'd like to learn more about [hyperparameter optimization](https://en.wikipedia.org/wiki/Hyperparameter_optimization), you're encouraged to start with **grid search**, which is a straightforward method for determining the best _combination_ of parameters for a machine learning model. Thankfully, scikit-learn also contains a built-in function [`GridSearchCV()`](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.GridSearchCV.html) that can make your grid search code very efficient!
# Keep going
Continue to learn about **[gradient boosting](https://www.kaggle.com/alexisbcook/xgboost)**, a powerful technique that achieves state-of-the-art results on a variety of datasets.
---
**[Intermediate Machine Learning Home Page](https://www.kaggle.com/learn/intermediate-machine-learning)**
*Have questions or comments? Visit the [Learn Discussion forum](https://www.kaggle.com/learn-forum) to chat with other Learners.*
| github_jupyter |
# How Big a Table can the Carpenter Build?
> You’re on a DIY kick and want to build a circular dining table which can be split in half so leaves can be added when entertaining guests. As luck would have it, on your last trip to the lumber yard, you came across the most pristine piece of exotic wood that would be perfect for the circular table top. Trouble is, the piece is rectangular. You are happy to have the leaves fashioned from one of the slightly-less-than-perfect pieces underneath it, but there’s still the issue of the main circle. You devise a plan: cut two congruent semicircles from the perfect 4-by-8-foot piece and reassemble them to form the circular top of your table. What is the radius of the largest possible circular table you can make? -- http://fivethirtyeight.com/features/how-big-a-table-can-the-carpenter-build/
Let's solve it!
```
%matplotlib inline
%config InlineBackend.figure_format='retina'
from __future__ import absolute_import, division, print_function
import matplotlib as mpl
from matplotlib import pyplot as plt
from matplotlib.pyplot import GridSpec
import seaborn as sns
import mpld3
import numpy as np
import pandas as pd
import os, sys
import warnings
from urllib.request import urlopen
from IPython.display import HTML
from ipywidgets import interactive, FloatSlider
warnings.filterwarnings('always')
plt.rcParams['figure.figsize'] = (8, 8)
def plot_circles(distance_from_side=2.5):
fig=plt.figure(1)
plt.axis([0,8,0,8])
plt.grid(b=None, )
ax=fig.add_subplot(1,1,1)
rect = plt.Rectangle((0, 2), 8, 4, alpha=0.4)
ax.add_patch(rect)
circ=plt.Circle((distance_from_side, 2.0), radius=distance_from_side,
lw=2.0, color='g', alpha=0.5, fill=False,)
ax.add_patch(circ)
circ2=plt.Circle((8 - distance_from_side, 6.0), radius=distance_from_side,
lw=2.0, color='b', alpha=0.5, fill=False,)
ax.add_patch(circ2)
ax.set_ylim(0, 8)
plt.scatter(distance_from_side, 2.0, c='g')
plt.scatter(8 - distance_from_side, 6.0, c='b')
ax.text(.5, 7, "Using long sides (2.5)", zorder=2, fontdict={'size':32})
fig.tight_layout()
plot_circles()
def plot_circles_diag(distance_from_center=1.33, radius=2.666):
slope = 0.5
intercept = 2.0
fig=plt.figure(1)
plt.axis([0,8,0,8])
plt.grid(b=None, )
ax=fig.add_subplot(1,1,1)
rect = plt.Rectangle((0, 2), 8, 4, alpha=0.4)
ax.add_patch(rect)
plt.plot([0, 8], [2, 6], c='k', lw=0.5)
center_1 = ((4 - distance_from_center), slope * (4 - distance_from_center) + intercept)
center_2 = ((4 + distance_from_center), slope * (4 + distance_from_center) + intercept)
circ=plt.Circle(center_1, radius=radius,
lw=2.0, color='g', alpha=0.5, fill=False,)
ax.add_patch(circ)
circ2=plt.Circle(center_2, radius=radius,
lw=2.0, color='b', alpha=0.5, fill=False,)
ax.add_patch(circ2)
ax.set_ylim(0, 8)
ax.set_title("Radius: " + str(round(radius, 5)))
fig.tight_layout()
plot_circles_diag()
distance_slider = FloatSlider(value=1.41, min=1.310, max=1.50, step=0.01)
radius_slider = FloatSlider(value=2.5, min=2.5, max=3.0, step=0.01)
interactive(plot_circles_diag,
radius=radius_slider,
distance_from_center=distance_slider,
)
# Proposed solution w/ radius 2.7054 -- I don't think that it works.
distance_slider = FloatSlider(value=1.41, min=1.310, max=1.50, step=0.01)
radius_slider = FloatSlider(value=2.7054, min=2.5, max=3.0, step=0.01)
interactive(plot_circles_diag,
radius=radius_slider,
distance_from_center=distance_slider,
)
```
| github_jupyter |
- https://gym.openai.com/
- https://openai.com/blog/universe/
- Pytorch参考, https://pytorch.org/tutorials/intermediate/reinforcement_q_learning.html
- https://github.com/Finspire13/pytorch-policy-gradient-example/blob/master/pg.py
- 一个博客, https://karpathy.github.io/2016/05/31/rl/
## 关于Gym的使用
```
#remove " > /dev/null 2>&1" to see what is going on under the hood
!pip install gym pyvirtualdisplay > /dev/null 2>&1
!apt-get install -y xvfb python-opengl ffmpeg > /dev/null 2>&1
!apt-get update > /dev/null 2>&1
!apt-get install cmake > /dev/null 2>&1
!pip install --upgrade setuptools 2>&1
!pip install ez_setup > /dev/null 2>&1
!pip install gym[atari] > /dev/null 2>&1
import gym
from gym import logger as gymlogger
from gym.wrappers import Monitor
gymlogger.set_level(40) #error only
import numpy as np
import random
import matplotlib
import matplotlib.pyplot as plt
%matplotlib inline
import math
import glob
import io
import base64
from IPython.display import HTML
from IPython import display as ipythondisplay
from pyvirtualdisplay import Display
display = Display(visible=0, size=(1400, 900))
display.start()
```
## Gym简单测试
```
import gym
env = gym.make('CartPole-v0')
env.reset()
for _ in range(1000):
env.render()
env.step(env.action_space.sample()) # take a random action
env.close()
```
### 关于 Action Space 与 Observation Space
```
# 打印 action space
print(env.action_space)
[env.action_space.sample() for i in range(10)]
# 打印 observation space
print(env.observation_space)
print(env.observation_space.high)
print(env.observation_space.low)
[env.observation_space.bounded_above, env.observation_space.bounded_below]
```
### 关于 step 返回的 obs, reward, done 与 info
```
env = gym.make('CartPole-v0')
obs0 = env.reset()
print("initial observation code:", obs0, type(obs0))
new_obs, reward, is_done, info = env.step(1) # taking action 1
print("new observation code:", new_obs, type(new_obs))
print("reward:", reward, type(reward))
print("is game over?:", is_done, type(is_done))
print("info:", info, type(info))
```
## 使用 Video 的方式可视化
```
"""
Utility functions to enable video recording of gym environment and displaying it
To enable video, just do "env = wrap_env(env)""
"""
def show_video():
mp4list = glob.glob('video/*.mp4')
if len(mp4list) > 0:
mp4 = mp4list[0]
video = io.open(mp4, 'r+b').read()
encoded = base64.b64encode(video)
ipythondisplay.display(HTML(data='''<video alt="test" autoplay
loop controls style="height: 400px;">
<source src="data:video/mp4;base64,{0}" type="video/mp4" />
</video>'''.format(encoded.decode('ascii'))))
else:
print("Could not find video")
def wrap_env(env):
env = Monitor(env, './video', force=True)
return env
env = wrap_env(gym.make("CartPole-v0"))
observation = env.reset()
while True:
env.render()
#your agent goes here
action = env.action_space.sample()
observation, reward, done, info = env.step(action)
if done:
break;
env.close()
show_video()
```
### 可视化例子
```
env = gym.make('CartPole-v0') # 初始化场景
env.reset() # 初始状态
env.render(mode='rgb_array')
t = 0
# 随便动一步(初始状态)
action = env.action_space.sample()
state, _, done, _ = env.step(action)
env.render(mode='rgb_array')
# 绘图
plt.figure()
plt.clf()
plt.title('Example extracted screen')
while True:
action = env.action_space.sample()
# 往后走一步
state, _, done, _ = env.step(action)
# 生成走后的场景
current_screen = env.render(mode='rgb_array') # 返回现在的图像, 用于可视化
# 绘制画面
plt.pause(0.7) # pause a bit so that plots are updated
ipythondisplay.clear_output(wait=True)
ipythondisplay.display(plt.gcf())
plt.title('Action: {}'.format(action))
plt.imshow(current_screen, interpolation='none')
t = t + 1
if done:
break
if t>100:
break
plt.show()
env.close()
print(t)
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.