text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
# Artificial Intelligence Nanodegree
## Convolutional Neural Networks
---
In this notebook, we train a CNN to classify images from the CIFAR-10 database.
### 1. Load CIFAR-10 Database
```
import keras
from keras.datasets import cifar10
# load the pre-shuffled train and test data
(x_train, y_train), (x_test, y_test) = cifar10.load_data()
```
### 2. Visualize the First 24 Training Images
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
fig = plt.figure(figsize=(20,5))
for i in range(36):
ax = fig.add_subplot(3, 12, i + 1, xticks=[], yticks=[])
ax.imshow(np.squeeze(x_train[i]))
```
### 3. Rescale the Images by Dividing Every Pixel in Every Image by 255
```
# rescale [0,255] --> [0,1]
x_train = x_train.astype('float32')/255
x_test = x_test.astype('float32')/255
```
### 4. Break Dataset into Training, Testing, and Validation Sets
```
from keras.utils import np_utils
# one-hot encode the labels
num_classes = len(np.unique(y_train))
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
# break training set into training and validation sets
(x_train, x_valid) = x_train[5000:], x_train[:5000]
(y_train, y_valid) = y_train[5000:], y_train[:5000]
# print shape of training set
print('x_train shape:', x_train.shape)
# print number of training, validation, and test images
print(x_train.shape[0], 'train samples')
print(x_test.shape[0], 'test samples')
print(x_valid.shape[0], 'validation samples')
```
### 5. Define the Model Architecture
```
from keras.models import Sequential
from keras.layers import Conv2D, MaxPooling2D, Flatten, Dense, Dropout
model = Sequential()
model.add(Conv2D(filters=16, kernel_size=2, padding='same', activation='relu',
input_shape=(32, 32, 3)))
model.add(MaxPooling2D(pool_size=2))
model.add(Conv2D(filters=32, kernel_size=2, padding='same', activation='relu'))
model.add(MaxPooling2D(pool_size=2))
model.add(Conv2D(filters=64, kernel_size=2, padding='same', activation='relu'))
model.add(MaxPooling2D(pool_size=2))
model.add(Dropout(0.3))
model.add(Flatten())
model.add(Dense(500, activation='relu'))
model.add(Dropout(0.4))
model.add(Dense(10, activation='softmax'))
model.summary()
```
### 6. Compile the Model
```
# compile the model
model.compile(loss='categorical_crossentropy', optimizer='rmsprop',
metrics=['accuracy'])
```
### 7. Train the Model
```
from keras.callbacks import ModelCheckpoint
# train the model
checkpointer = ModelCheckpoint(filepath='model.weights.best.hdf5', verbose=1,
save_best_only=True)
hist = model.fit(x_train, y_train, batch_size=32, epochs=100,
validation_data=(x_valid, y_valid), callbacks=[checkpointer],
verbose=2, shuffle=True)
```
### 8. Load the Model with the Best Validation Accuracy
```
# load the weights that yielded the best validation accuracy
model.load_weights('model.weights.best.hdf5')
```
### 9. Calculate Classification Accuracy on Test Set
```
# evaluate and print test accuracy
score = model.evaluate(x_test, y_test, verbose=0)
print('\n', 'Test accuracy:', score[1])
```
### 10. Visualize Some Predictions
This may give you some insight into why the network is misclassifying certain objects.
```
# get predictions on the test set
y_hat = model.predict(x_test)
# define text labels (source: https://www.cs.toronto.edu/~kriz/cifar.html)
cifar10_labels = ['airplane', 'automobile', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck']
# plot a random sample of test images, their predicted labels, and ground truth
fig = plt.figure(figsize=(20, 8))
for i, idx in enumerate(np.random.choice(x_test.shape[0], size=32, replace=False)):
ax = fig.add_subplot(4, 8, i + 1, xticks=[], yticks=[])
ax.imshow(np.squeeze(x_test[idx]))
pred_idx = np.argmax(y_hat[idx])
true_idx = np.argmax(y_test[idx])
ax.set_title("{} ({})".format(cifar10_labels[pred_idx], cifar10_labels[true_idx]),
color=("green" if pred_idx == true_idx else "red"))
```
| github_jupyter |
```
import datetime
lipidname = "PQP1"
tail = "CDDDC CCCC"
link = "G G"
head = "P1"
description = "; A phosphorylated phosphatidylinositol (PI) lipid \n; C16:0 palmitic acid , C20:3(5c,8c,11c) mead acid \n"
modeledOn="; This topology follows the standard Martini 2.0 lipid definitions and building block rules.\n; Reference(s): \n; S.J. Marrink, A.H. de Vries, A.E. Mark. Coarse grained model for semi-quantitative lipid simulations. JPC-B, 108:750-760, \n; 2004. doi:10.1021/jp036508g \n; S.J. Marrink, H.J. Risselada, S. Yefimov, D.P. Tieleman, A.H. de Vries. The MARTINI force field: coarse grained model for \n; biomolecular simulations. JPC-B, 111:7812-7824, 2007. doi:10.1021/jp071097f \n; T.A. Wassenaar, H.I. Ingolfsson, R.A. Bockmann, D.P. Tieleman, S.J. Marrink. Computational lipidomics with insane: a versatile \n; tool for generating custom membranes for molecular simulations. JCTC, 150410125128004, 2015. doi:10.1021/acs.jctc.5b00209\n; Created: "
now = datetime.datetime.now()
membrane="testmembrane"
insane="../insane+SF.py"
mdparams="../test.mdp"
martinipath="../martini.ff/"
ITPCatalogue="./epithelial.cat"
ITPMasterFile="martini_v2_epithelial.itp"
modeledOn+= now.strftime("%Y.%m.%d")+"\n"
# Cleaning up intermediate files from previous runs
!rm -f *#*
!rm -f *step*
!rm -f {membrane}*
import fileinput
import os.path
print("Create itp")
!python {martinipath}/lipid-martini-itp-v06.py -o {lipidname}.itp -alname {lipidname} -name {lipidname} -alhead '{head}' -allink '{link}' -altail '{tail}'
#update description and parameters
with fileinput.FileInput(lipidname+".itp", inplace=True) as file:
for line in file:
if line == "; This is a ...\n":
print(description, end='')
elif line == "; Was modeled on ...\n":
print(modeledOn, end='')
else:
print(line, end='')
#Add this ITP file to the catalogue file
if not os.path.exists(ITPCatalogue):
ITPCatalogueData = []
else:
with open(ITPCatalogue, 'r') as file :
ITPCatalogueData = file.read().splitlines()
ITPCatalogueData = [x for x in ITPCatalogueData if not x==lipidname+".itp"]
ITPCatalogueData.append(lipidname+".itp")
with open(ITPCatalogue, 'w') as file :
file.writelines("%s\n" % item for item in ITPCatalogueData)
#build ITPFile
with open(martinipath+ITPMasterFile, 'w') as masterfile:
for ITPfilename in ITPCatalogueData:
with open(ITPfilename, 'r') as ITPfile :
for line in ITPfile:
masterfile.write(line)
print("Done")
# build a simple membrane to visualize this species
!python2 {insane} -o {membrane}.gro -p {membrane}.top -d 0 -x 3 -y 3 -z 3 -sol PW -center -charge 0 -orient -u {lipidname}:1 -l {lipidname}:1 -itpPath {martinipath}
import os #Operating system specific commands
import re #Regular expression library
print("Test")
print("Grompp")
grompp = !gmx grompp -f {mdparams} -c {membrane}.gro -p {membrane}.top -o {membrane}.tpr
success=True
for line in grompp:
if re.search("ERROR", line):
success=False
if re.search("Fatal error", line):
success=False
#if not success:
print(line)
if success:
print("Run")
!export GMX_MAXCONSTRWARN=-1
!export GMX_SUPPRESS_DUMP=1
run = !gmx mdrun -v -deffnm {membrane}
summary=""
logfile = membrane+".log"
if not os.path.exists(logfile):
print("no log file")
print("== === ====")
for line in run:
print(line)
else:
try:
file = open(logfile, "r")
fe = False
for line in file:
if fe:
success=False
summary=line
elif re.search("^Steepest Descents.*converge", line):
success=True
summary=line
break
elif re.search("Fatal error", line):
fe = True
except IOError as exc:
sucess=False;
summary=exc;
if success:
print("Success")
else:
print(summary)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/Tessellate-Imaging/Monk_Object_Detection/blob/master/application_model_zoo/Example%20-%20STAIRS%20Action%20Recognition%20Dataset.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Table of contents
## Installation
## Training your own classifier
# Installation
- Run these commands
- git clone https://github.com/Tessellate-Imaging/Monk_Object_Detection.git
- cd Monk_Object_Detection/18_mmaction/installation
- Select the right file and run
- chmod +x install.sh && ./install.sh
```
! git clone https://github.com/Tessellate-Imaging/Monk_Object_Detection.git
! cd Monk_Object_Detection/18_mmaction/installation && chmod +x install.sh && ./install.sh
```
# Download and reformat Dataset
- Dataset Credits: https://actions.stair.center/
```
! git clone https://github.com/STAIR-Lab-CIT/STAIR-actions.git
! pip install tqdm
! mkdir videos
# Change line 47 in STAIR-actions/download.sh
# Change jq to .jq
! cd STAIR-actions && chmod +x download.sh && ./download.sh
! cd ..
import os
classes_list = sorted(os.listdir("STAIR-actions/STAIR_Actions_v1.1"));
f = open("classes.txt", 'w')
for i in range(len(classes_list)):
f.write(classes_list[i] + "\n");
f.close();
import os
from tqdm import tqdm
complete_list = [];
for i in tqdm(range(len(classes_list))):
folder_name = classes_list[i];
vid_label = str(classes_list.index(folder_name));
vid_list = os.listdir("STAIR-actions/STAIR_Actions_v1.1/" + folder_name);
for j in tqdm(range(len(vid_list))):
vid_name = vid_list[j];
wr = str(vid_name) + " " + vid_label;
os.system("cp STAIR-actions/STAIR_Actions_v1.1/" + folder_name + "/" + vid_name + " videos/")
complete_list.append(wr);
import random
random.shuffle(complete_list)
train_list = complete_list[:int(0.8*len(complete_list))];
val_list = complete_list[int(0.8*len(complete_list)):];
f = open("train.txt", 'w')
for i in range(len(train_list)):
f.write(train_list[i] + "\n");
f.close();
f = open("val.txt", 'w')
for i in range(len(val_list)):
f.write(val_list[i] + "\n");
f.close();
```
# Train your own network
```
import os
import sys
sys.path.append("Monk_Object_Detection/18_mmaction/lib");
from train_engine import Detector_Videos
gtf = Detector_Videos();
video_dir = 'videos/';
anno_file = 'train.txt';
classes_list_file = 'Dataset/classes.txt';
gtf.Train_Video_Dataset(video_dir,
anno_file,
classes_list_file);
video_dir = 'videos';
anno_file = 'val.txt';
gtf.Val_Video_Dataset(video_dir,
anno_file);
gtf.Dataset_Params(batch_size=4, num_workers=4)
gtf.List_Models();
gtf.Model_Params(model_name="tsm_r50", gpu_devices=[0])
gtf.Hyper_Params(lr=0.002, momentum=0.9, weight_decay=0.0001)
gtf.Training_Params(num_epochs=100, val_interval=10)
gtf.Train();
```
# Inference on trained network
```
import os
import sys
sys.path.append("Monk_Object_Detection/18_mmaction/lib");
from infer_engine import Infer_Videos
gtf = Infer_Videos();
gtf.Dataset_Params('classes.txt');
config_file = "work_dirs/config.py"
checkpoint_file = "work_dirs/latest.pth"
gtf.Model_Params(config_file, checkpoint_file, use_gpu=True)
import os
vid_list = os.listdir("videos")
video_path = "videos/" + vid_list[0]
results = gtf.Predict(video_path)
from IPython.display import Video
Video(video_path)
video_path = "videos/" + vid_list[1]
results = gtf.Predict(video_path)
from IPython.display import Video
Video(video_path)
```
| github_jupyter |
```
from __future__ import division
from __future__ import print_function
from __future__ import absolute_import
import os
import tempfile
import datetime
import pandas as pd
import numpy as np
import apache_beam as beam
import tensorflow as tf
import tensorflow_transform as tft
import tensorflow_transform.beam as tft_beam
from tensorflow_transform.tf_metadata import dataset_metadata
from tensorflow_transform.tf_metadata import dataset_schema
import tensorflow_transform.beam.impl as beam_impl
from utilities.transforms import MapAndFilterErrors
tf.__version__
PWD=os.getcwd()
BUCKET='going-tfx'
PROJECT='going-tfx'
EXTRA_PACKAGE='./utilities'
DATA_DIR="gs://{}/flight_data".format(BUCKET)
OUTPUT_DIR="gs://{}/output".format(BUCKET)
TMP_DIR="gs://{}/tmp".format(BUCKET)
ALL_COLUMNS = ['FL_DATE', 'FL_YEAR', 'FL_MONTH', 'FL_DOM', 'FL_DOW', 'UNIQUE_CARRIER', 'FL_NUM',
'ORIGIN_AIRPORT_SEQ_ID', 'DEST_AIRPORT_SEQ_ID', 'ORIGIN', 'DEST',
'CRS_DEP_TIME', 'DEP_DELAY', 'TAXI_OUT', 'WHEELS_OFF', 'WHEELS_ON',
'TAXI_IN', 'CRS_ARR_TIME', 'ARR_DELAY', 'CANCELLED',
'CANCELLATION_CODE', 'DIVERTED', 'DISTANCE']
DEFAULTS = [["-"], [], [], [], [], ["-"], ["-"], ["-"], ["-"], ["-"], ["-"], [], [], [], [], [], [], [], [], [], ['NONE'], [], []]
ORDERED_COLUMNS = ['FL_YEAR', 'FL_MONTH', 'FL_DOW', 'UNIQUE_CARRIER', 'ORIGIN', 'DEST', 'CRS_DEP_TIME', 'DEP_DELAY', 'CRS_ARR_TIME', 'ARR_DELAY']
SELECT = list(np.sort([ALL_COLUMNS.index(c) for c in ORDERED_COLUMNS]))
```
---
Get an impression of what's in the file
```
atlanta_june = tf.data.TextLineDataset(os.path.join(DATA_DIR, 'atl_june.csv'))
def decode_csv(row):
cols = tf.decode_csv(row, select_cols=SELECT, record_defaults=[DEFAULTS[i] for i in SELECT])
features = dict(zip([ALL_COLUMNS[i] for i in SELECT], cols))
return features
inp = atlanta_june.skip(1).map(decode_csv).batch(10)
input_raw = inp.make_one_shot_iterator().get_next()
with tf.Session() as sess:
b = sess.run(input_raw)
pd.DataFrame(b)
```
---
### Metadata and Schema
```
raw_data_schema = {
colname : dataset_schema.ColumnSchema(
tf.string, [], dataset_schema.FixedColumnRepresentation())
for colname in ['ORIGIN','UNIQUE_CARRIER','DEST']
}
raw_data_schema.update({
colname : dataset_schema.ColumnSchema(
tf.float32, [], dataset_schema.FixedColumnRepresentation())
for colname in ['DEP_DELAY','ARR_DELAY']
})
raw_data_schema.update({
colname : dataset_schema.ColumnSchema(
tf.int64, [], dataset_schema.FixedColumnRepresentation())
for colname in ['FL_YEAR','FL_MONTH','FL_DOW','CRS_DEP_TIME','CRS_ARR_TIME']
})
raw_data_metadata = dataset_metadata.DatasetMetadata(dataset_schema.Schema(raw_data_schema))
```
Another way of creating meta data is as follows
```
raw_data_metadata = dataset_metadata.DatasetMetadata(
dataset_schema.from_feature_spec({
'ORIGIN': tf.FixedLenFeature([], tf.string),
'FL_YEAR': tf.FixedLenFeature([], tf.int64),
'FL_MONTH': tf.FixedLenFeature([], tf.int64),
'FL_DOW': tf.FixedLenFeature([], tf.int64),
'UNIQUE_CARRIER': tf.FixedLenFeature([], tf.string),
'DEST': tf.FixedLenFeature([], tf.string),
'CRS_DEP_TIME': tf.FixedLenFeature([], tf.int64),
'CRS_ARR_TIME': tf.FixedLenFeature([], tf.int64),
'DEP_DELAY': tf.FixedLenFeature([], tf.float32),
'ARR_DELAY': tf.FixedLenFeature([], tf.float32)
}))
```
---
### The Preprocessing Pipeline
Here, we scale ```DEP_DELAY``` to the range $[0,1]$ and select only the interesting columns
```
def preprocessing_fn(inputs):
outputs = inputs.copy()
outputs = {k: outputs[k] for k in ORDERED_COLUMNS} # Projection: Select useful columns
outputs['DEP_DELAY'] = tft.scale_to_0_1(outputs['DEP_DELAY']) # Scale
return outputs
```
The CSV encoder and decoder
```
training_file = os.path.join(DATA_DIR, 'atl_june.csv') # 403'358 records
csv_decode = tft.coders.CsvCoder(ALL_COLUMNS, raw_data_metadata.schema).decode
csv_encode = tft.coders.CsvCoder(ORDERED_COLUMNS, raw_data_metadata.schema).encode
# Uncomment this to use a smaller test file
training_file = 'atl_june_11.csv' # 10 records only
```
A quick sanity check on for the error filter with a very small subset
```
with open('atl_june_11.csv') as f:
content = f.readlines()
(content
| MapAndFilterErrors(csv_decode)
| beam.Map(csv_encode))
```
Cleaning up any previously processed records...
```
!gsutil rm -f $OUTPUT_DIR/atl_june_transformed*
```
---
### The Pipeline
Runtime options:
```
test_mode = True
job_name = 'tft-tutorial-' + datetime.datetime.now().strftime('%y%m%d-%H%M%S')
tmp_dir = os.path.join(TMP_DIR, job_name)
options = {
'staging_location': os.path.join(tmp_dir, 'staging'),
'temp_location': tmp_dir,
'job_name': job_name,
'project': PROJECT,
'max_num_workers': 24,
'teardown_policy': 'TEARDOWN_ALWAYS',
'requirements_file': 'requirements.txt',
'setup_file': os.path.join(PWD, 'setup.py')
}
opts = beam.pipeline.PipelineOptions(flags=[], **options)
if test_mode:
RUNNER = 'DirectRunner'
else:
RUNNER = 'DataflowRunner'
print(RUNNER)
print(opts.get_all_options())
```
---
### Construct and run the pipeline
Be aware that this can take quite a while. You can watch your pipeline run on the dataflow console
```
tf.logging.set_verbosity(tf.logging.WARN)
with beam.Pipeline(RUNNER, options=opts) as pipeline:
with beam_impl.Context(temp_dir=tmp_dir):
# Decode the raw data from CSV file and filter outliers
raw_data = (
pipeline
| 'ReadData' >> beam.io.ReadFromText(training_file, skip_header_lines=1)
| 'Decode' >> MapAndFilterErrors(csv_decode)
| 'Filter_outliers' >> beam.Filter(lambda r: r['DEP_DELAY'] < 120.0)
)
# Analyse and transform - handle meta_data
raw_dataset = (raw_data, raw_data_metadata)
t_dataset, t_fn = (raw_dataset | beam_impl.AnalyzeAndTransformDataset(preprocessing_fn))
t_data, t_metadata = t_dataset
# Encode back to CSV file(s)
res = (t_data
| beam.Map(csv_encode)
| beam.io.WriteToText(file_path_prefix=os.path.join(OUTPUT_DIR, "atl_june_transformed")))
result = pipeline.run()
```
---
If you want to wait for the result, this would be the way to do it
```
# result.wait_until_finish()
```
---
The resulting CSV files
```
!gsutil ls -l $OUTPUT_DIR/atl_june_transformed*
!gsutil cat $OUTPUT_DIR/atl_june_transformed-00000-of-00001 | wc -l
!gsutil cat $OUTPUT_DIR/atl_june_transformed-00000-of-00001 | head -10
```
| github_jupyter |
# Amazon SageMaker Multi-Model Endpoints using Scikit Learn
With [Amazon SageMaker multi-model endpoints](https://docs.aws.amazon.com/sagemaker/latest/dg/multi-model-endpoints.html), customers can create an endpoint that seamlessly hosts up to thousands of models. These endpoints are well suited to use cases where any one of a large number of models, which can be served from a common inference container, needs to be invokable on-demand and where it is acceptable for infrequently invoked models to incur some additional latency. For applications which require consistently low inference latency, a traditional endpoint is still the best choice.
At a high level, Amazon SageMaker manages the loading and unloading of models for a multi-model endpoint, as they are needed. When an invocation request is made for a particular model, Amazon SageMaker routes the request to an instance assigned to that model, downloads the model artifacts from S3 onto that instance, and initiates loading of the model into the memory of the container. As soon as the loading is complete, Amazon SageMaker performs the requested invocation and returns the result. If the model is already loaded in memory on the selected instance, the downloading and loading steps are skipped and the invocation is performed immediately.
To demonstrate how multi-model endpoints are created and used, this notebook provides an example using a set of Scikit Learn models that each predict housing prices for a single location. This domain is used as a simple example to easily experiment with multi-model endpoints.
The Amazon SageMaker multi-model endpoint capability is designed to work across all machine learning frameworks and algorithms including those where you bring your own container.
### Contents
1. [Build and register a Scikit Learn container that can serve multiple models](#Build-and-register-a-Scikit-Learn-container-that-can-serve-multiple-models)
1. [Generate synthetic data for housing models](#Generate-synthetic-data-for-housing-models)
1. [Train multiple house value prediction models](#Train-multiple-house-value-prediction-models)
1. [Import models into hosting](#Import-models-into-hosting)
1. [Deploy model artifacts to be found by the endpoint](#Deploy-model-artifacts-to-be-found-by-the-endpoint)
1. [Create the Amazon SageMaker model entity](#Create-the-Amazon-SageMaker-model-entity)
1. [Create the multi-model endpoint](#Create-the-multi-model-endpoint)
1. [Exercise the multi-model endpoint](#Exercise-the-multi-model-endpoint)
1. [Dynamically deploy another model](#Dynamically-deploy-another-model)
1. [Invoke the newly deployed model](#Invoke-the-newly-deployed-model)
1. [Updating a model](#Updating-a-model)
1. [Clean up](#Clean-up)
## Build and register a Scikit Learn container that can serve multiple models
```
!pip install -qU awscli boto3 sagemaker
```
For the inference container to serve multiple models in a multi-model endpoint, it must implement [additional APIs](https://docs.aws.amazon.com/sagemaker/latest/dg/build-multi-model-build-container.html) in order to load, list, get, unload and invoke specific models.
The ['mme' branch of the SageMaker Scikit Learn Container repository](https://github.com/aws/sagemaker-scikit-learn-container/tree/mme) is an example implementation on how to adapt SageMaker's Scikit Learn framework container to use [Multi Model Server](https://github.com/awslabs/multi-model-server), a framework that provides an HTTP frontend that implements the additional container APIs required by multi-model endpoints, and also provides a pluggable backend handler for serving models using a custom framework, in this case the Scikit Learn framework.
Using this branch, below we will build a Scikit Learn container image that fulfills all of the multi-model endpoint container requirements, and then upload that image to Amazon Elastic Container Registry (ECR). Because uploading the image to ECR may create a new ECR repository, this notebook requires permissions in addition to the regular SageMakerFullAccess permissions. The easiest way to add these permissions is simply to add the managed policy AmazonEC2ContainerRegistryFullAccess to the role that you used to start your notebook instance. There's no need to restart your notebook instance when you do this, the new permissions will be available immediately.
```
ALGORITHM_NAME = 'multi-model-sklearn'
%%sh -s $ALGORITHM_NAME
algorithm_name=$1
account=$(aws sts get-caller-identity --query Account --output text)
# Get the region defined in the current configuration (default to us-west-2 if none defined)
region=$(aws configure get region)
ecr_image="${account}.dkr.ecr.${region}.amazonaws.com/${algorithm_name}:latest"
# If the repository doesn't exist in ECR, create it.
aws ecr describe-repositories --repository-names "${algorithm_name}" > /dev/null 2>&1
if [ $? -ne 0 ]
then
aws ecr create-repository --repository-name "${algorithm_name}" > /dev/null
fi
# Get the login command from ECR and execute it directly
$(aws ecr get-login --region ${region} --no-include-email --registry-ids ${account})
# Build the docker image locally with the image name and then push it to ECR
# with the full image name.
# First clear out any prior version of the cloned repo
rm -rf sagemaker-scikit-learn-container/
# Clone the sklearn container repo
git clone --single-branch --branch mme https://github.com/aws/sagemaker-scikit-learn-container.git
cd sagemaker-scikit-learn-container/
# Build the "base" container image that encompasses the installation of the
# scikit-learn framework and all of the dependencies needed.
docker build -q -t sklearn-base:0.20-2-cpu-py3 -f docker/0.20-2/base/Dockerfile.cpu --build-arg py_version=3 .
# Create the SageMaker Scikit-learn Container Python package.
python setup.py bdist_wheel --universal
# Build the "final" container image that encompasses the installation of the
# code that implements the SageMaker multi-model container requirements.
docker build -q -t ${algorithm_name} -f docker/0.20-2/final/Dockerfile.cpu .
docker tag ${algorithm_name} ${ecr_image}
docker push ${ecr_image}
```
## Generate synthetic data for housing models
```
import numpy as np
import pandas as pd
import json
import datetime
import time
from time import gmtime, strftime
import matplotlib.pyplot as plt
NUM_HOUSES_PER_LOCATION = 1000
LOCATIONS = ['NewYork_NY', 'LosAngeles_CA', 'Chicago_IL', 'Houston_TX', 'Dallas_TX',
'Phoenix_AZ', 'Philadelphia_PA', 'SanAntonio_TX', 'SanDiego_CA', 'SanFrancisco_CA']
PARALLEL_TRAINING_JOBS = 4 # len(LOCATIONS) if your account limits can handle it
MAX_YEAR = 2019
def gen_price(house):
_base_price = int(house['SQUARE_FEET'] * 150)
_price = int(_base_price + (10000 * house['NUM_BEDROOMS']) + \
(15000 * house['NUM_BATHROOMS']) + \
(15000 * house['LOT_ACRES']) + \
(15000 * house['GARAGE_SPACES']) - \
(5000 * (MAX_YEAR - house['YEAR_BUILT'])))
return _price
def gen_random_house():
_house = {'SQUARE_FEET': int(np.random.normal(3000, 750)),
'NUM_BEDROOMS': np.random.randint(2, 7),
'NUM_BATHROOMS': np.random.randint(2, 7) / 2,
'LOT_ACRES': round(np.random.normal(1.0, 0.25), 2),
'GARAGE_SPACES': np.random.randint(0, 4),
'YEAR_BUILT': min(MAX_YEAR, int(np.random.normal(1995, 10)))}
_price = gen_price(_house)
return [_price, _house['YEAR_BUILT'], _house['SQUARE_FEET'],
_house['NUM_BEDROOMS'], _house['NUM_BATHROOMS'],
_house['LOT_ACRES'], _house['GARAGE_SPACES']]
COLUMNS = ['PRICE', 'YEAR_BUILT', 'SQUARE_FEET', 'NUM_BEDROOMS',
'NUM_BATHROOMS', 'LOT_ACRES', 'GARAGE_SPACES']
def gen_houses(num_houses):
_house_list = []
for i in range(num_houses):
_house_list.append(gen_random_house())
_df = pd.DataFrame(_house_list,
columns=COLUMNS)
return _df
```
## Train multiple house value prediction models
```
import sagemaker
from sagemaker import get_execution_role
from sagemaker.predictor import csv_serializer
import boto3
sm_client = boto3.client(service_name='sagemaker')
runtime_sm_client = boto3.client(service_name='sagemaker-runtime')
s3 = boto3.resource('s3')
s3_client = boto3.client('s3')
sagemaker_session = sagemaker.Session()
role = get_execution_role()
ACCOUNT_ID = boto3.client('sts').get_caller_identity()['Account']
REGION = boto3.Session().region_name
BUCKET = sagemaker_session.default_bucket()
SCRIPT_FILENAME = 'script.py'
USER_CODE_ARTIFACTS = 'user_code.tar.gz'
MULTI_MODEL_SKLEARN_IMAGE = '{}.dkr.ecr.{}.amazonaws.com/{}:latest'.format(ACCOUNT_ID, REGION,
ALGORITHM_NAME)
DATA_PREFIX = 'DEMO_MME_SCIKIT'
HOUSING_MODEL_NAME = 'housing'
MULTI_MODEL_ARTIFACTS = 'multi_model_artifacts'
TRAIN_INSTANCE_TYPE = 'ml.m4.xlarge'
ENDPOINT_INSTANCE_TYPE = 'ml.m4.xlarge'
```
### Split a given dataset into train, validation, and test
```
from sklearn.model_selection import train_test_split
SEED = 7
SPLIT_RATIOS = [0.6, 0.3, 0.1]
def split_data(df):
# split data into train and test sets
seed = SEED
val_size = SPLIT_RATIOS[1]
test_size = SPLIT_RATIOS[2]
num_samples = df.shape[0]
X1 = df.values[:num_samples, 1:] # keep only the features, skip the target, all rows
Y1 = df.values[:num_samples, :1] # keep only the target, all rows
# Use split ratios to divide up into train/val/test
X_train, X_val, y_train, y_val = \
train_test_split(X1, Y1, test_size=(test_size + val_size), random_state=seed)
# Of the remaining non-training samples, give proper ratio to validation and to test
X_test, X_test, y_test, y_test = \
train_test_split(X_val, y_val, test_size=(test_size / (test_size + val_size)),
random_state=seed)
# reassemble the datasets with target in first column and features after that
_train = np.concatenate([y_train, X_train], axis=1)
_val = np.concatenate([y_val, X_val], axis=1)
_test = np.concatenate([y_test, X_test], axis=1)
return _train, _val, _test
```
### Launch a single training job for a given housing location
There is nothing specific to multi-model endpoints in terms of the models it will host. They are trained in the same way as all other SageMaker models. Here we are using the Scikit Learn estimator and not waiting for the job to complete.
```
%%writefile $SCRIPT_FILENAME
import argparse
import os
import glob
import numpy as np
import pandas as pd
from sklearn.ensemble import RandomForestRegressor
from sklearn.externals import joblib
# inference functions ---------------
def model_fn(model_dir):
print('loading model.joblib from: {}'.format(model_dir))
_loaded_model = joblib.load(os.path.join(model_dir, 'model.joblib'))
return _loaded_model
if __name__ =='__main__':
print('extracting arguments')
parser = argparse.ArgumentParser()
# hyperparameters sent by the client are passed as command-line arguments to the script.
# to simplify the demo we don't use all sklearn RandomForest hyperparameters
parser.add_argument('--n-estimators', type=int, default=10)
parser.add_argument('--min-samples-leaf', type=int, default=3)
# Data, model, and output directories
parser.add_argument('--model-dir', type=str, default=os.environ.get('SM_MODEL_DIR'))
parser.add_argument('--train', type=str, default=os.environ.get('SM_CHANNEL_TRAIN'))
parser.add_argument('--validation', type=str, default=os.environ.get('SM_CHANNEL_VALIDATION'))
parser.add_argument('--model-name', type=str)
args, _ = parser.parse_known_args()
print('reading data')
print('model_name: {}'.format(args.model_name))
train_file = os.path.join(args.train, args.model_name + '_train.csv')
train_df = pd.read_csv(train_file)
val_file = os.path.join(args.validation, args.model_name + '_val.csv')
test_df = pd.read_csv(os.path.join(val_file))
print('building training and testing datasets')
X_train = train_df[train_df.columns[1:train_df.shape[1]]]
X_test = test_df[test_df.columns[1:test_df.shape[1]]]
y_train = train_df[train_df.columns[0]]
y_test = test_df[test_df.columns[0]]
# train
print('training model')
model = RandomForestRegressor(
n_estimators=args.n_estimators,
min_samples_leaf=args.min_samples_leaf,
n_jobs=-1)
model.fit(X_train, y_train)
# print abs error
print('validating model')
abs_err = np.abs(model.predict(X_test) - y_test)
# print couple perf metrics
for q in [10, 50, 90]:
print('AE-at-' + str(q) + 'th-percentile: '
+ str(np.percentile(a=abs_err, q=q)))
# persist model
path = os.path.join(args.model_dir, 'model.joblib')
joblib.dump(model, path)
print('model persisted at ' + path)
# can test the model locally
# ! python script.py --n-estimators 100 \
# --min-samples-leaf 2 \
# --model-dir ./ \
# --model-name 'NewYork_NY' \
# --train ./data/NewYork_NY/train/ \
# --validation ./data/NewYork_NY/val/
# from sklearn.externals import joblib
# regr = joblib.load('./model.joblib')
# _start_time = time.time()
# regr.predict([[0.0, 0.0, 0.0, 0.0, 0.0, 0.0]])
# _duration = time.time() - _start_time
# print('took {:,d} ms'.format(int(_duration * 1000)))
from sagemaker.sklearn.estimator import SKLearn
def launch_training_job(location):
# clear out old versions of the data
_s3_bucket = s3.Bucket(BUCKET)
_full_input_prefix = '{}/model_prep/{}'.format(DATA_PREFIX, location)
_s3_bucket.objects.filter(Prefix=_full_input_prefix + '/').delete()
# upload the entire set of data for all three channels
_local_folder = 'data/{}'.format(location)
_inputs = sagemaker_session.upload_data(path=_local_folder,
key_prefix=_full_input_prefix)
print('Training data uploaded: {}'.format(_inputs))
_job = 'mme-{}'.format(location.replace('_', '-'))
_full_output_prefix = '{}/model_artifacts/{}'.format(DATA_PREFIX,
location)
_s3_output_path = 's3://{}/{}'.format(BUCKET, _full_output_prefix)
_estimator = SKLearn(
entry_point=SCRIPT_FILENAME, role=role,
train_instance_count=1, train_instance_type=TRAIN_INSTANCE_TYPE,
framework_version='0.20.0',
output_path=_s3_output_path,
base_job_name=_job,
metric_definitions=[
{'Name' : 'median-AE',
'Regex': 'AE-at-50th-percentile: ([0-9.]+).*$'}],
hyperparameters = {'n-estimators' : 100,
'min-samples-leaf': 3,
'model-name' : location})
DISTRIBUTION_MODE = 'FullyReplicated'
_train_input = sagemaker.s3_input(s3_data=_inputs+'/train',
distribution=DISTRIBUTION_MODE, content_type='csv')
_val_input = sagemaker.s3_input(s3_data=_inputs+'/val',
distribution=DISTRIBUTION_MODE, content_type='csv')
_remote_inputs = {'train': _train_input, 'validation': _val_input}
_estimator.fit(_remote_inputs, wait=False)
return _estimator.latest_training_job.name
```
### Kick off a model training job for each housing location
```
def save_data_locally(location, train, val, test):
_header = ','.join(COLUMNS)
os.makedirs('data/{}/train'.format(location))
np.savetxt( 'data/{0}/train/{0}_train.csv'.format(location), train, delimiter=',', fmt='%.2f')
os.makedirs('data/{}/val'.format(location))
np.savetxt( 'data/{0}/val/{0}_val.csv'.format(location), val, delimiter=',', fmt='%.2f')
os.makedirs('data/{}/test'.format(location))
np.savetxt( 'data/{0}/test/{0}_test.csv'.format(location), test, delimiter=',', fmt='%.2f')
import shutil
import os
training_jobs = []
shutil.rmtree('data', ignore_errors=True)
for loc in LOCATIONS[:PARALLEL_TRAINING_JOBS]:
_houses = gen_houses(NUM_HOUSES_PER_LOCATION)
_train, _val, _test = split_data(_houses)
save_data_locally(loc, _train, _val, _test)
_job = launch_training_job(loc)
training_jobs.append(_job)
print('{} training jobs launched: {}'.format(len(training_jobs), training_jobs))
```
### Wait for all model training to finish
```
def wait_for_training_job_to_complete(job_name):
print('Waiting for job {} to complete...'.format(job_name))
_resp = sm_client.describe_training_job(TrainingJobName=job_name)
_status = _resp['TrainingJobStatus']
while _status=='InProgress':
time.sleep(60)
_resp = sm_client.describe_training_job(TrainingJobName=job_name)
_status = _resp['TrainingJobStatus']
if _status == 'InProgress':
print('{} job status: {}'.format(job_name, _status))
print('DONE. Status for {} is {}\n'.format(job_name, _status))
# wait for the jobs to finish
for j in training_jobs:
wait_for_training_job_to_complete(j)
```
## Import models into hosting
When creating the Model entity for a multi-model endpoint, the `ModelDataUrl` of the ContainerDefinition is treated as an S3 prefix for the model artifacts that will be loaded on-demand. The rest of the S3 path will be specified in the `InvokeEndpoint` request. Remember to close the location with a trailing slash.
The `Mode` of container is specified as `MultiModel` to signify that the container will host multiple models.
### Deploy model artifacts to be found by the endpoint
As described above, the multi-model endpoint is configured to find its model artifacts in a specific location in S3. For each trained model, we make a copy of its model artifacts into that location.
In our example, we are storing all the models within a single folder. The implementation of multi-model endpoints is flexible enough to permit an arbitrary folder structure. For a set of housing models for example, you could have a top level folder for each region, and the model artifacts would be copied to those regional folders. The target model referenced when invoking such a model would include the folder path. For example, `northeast/Boston_MA.tar.gz`.
```
import re
def parse_model_artifacts(model_data_url):
# extract the s3 key from the full url to the model artifacts
_s3_key = model_data_url.split('s3://{}/'.format(BUCKET))[1]
# get the part of the key that identifies the model within the model artifacts folder
_model_name_plus = _s3_key[_s3_key.find('model_artifacts') + len('model_artifacts') + 1:]
# finally, get the unique model name (e.g., "NewYork_NY")
_model_name = re.findall('^(.*?)/', _model_name_plus)[0]
return _s3_key, _model_name
# make a copy of the model artifacts from the original output of the training job to the place in
# s3 where the multi model endpoint will dynamically load individual models
def deploy_artifacts_to_mme(job_name):
_resp = sm_client.describe_training_job(TrainingJobName=job_name)
_source_s3_key, _model_name = parse_model_artifacts(_resp['ModelArtifacts']['S3ModelArtifacts'])
_copy_source = {'Bucket': BUCKET, 'Key': _source_s3_key}
_key = '{}/{}/{}.tar.gz'.format(DATA_PREFIX, MULTI_MODEL_ARTIFACTS, _model_name)
print('Copying {} model\n from: {}\n to: {}...'.format(_model_name, _source_s3_key, _key))
s3_client.copy_object(Bucket=BUCKET, CopySource=_copy_source, Key=_key)
return _key
```
Note that we are purposely *not* copying the first model. This will be copied later in the notebook to demonstrate how to dynamically add new models to an already running endpoint.
```
# First, clear out old versions of the model artifacts from previous runs of this notebook
s3 = boto3.resource('s3')
s3_bucket = s3.Bucket(BUCKET)
full_input_prefix = '{}/multi_model_artifacts'.format(DATA_PREFIX)
print('Removing old model artifacts from {}'.format(full_input_prefix))
filter_resp = s3_bucket.objects.filter(Prefix=full_input_prefix + '/').delete()
# copy every model except the first one
for job in training_jobs[1:]:
deploy_artifacts_to_mme(job)
```
### Create the Amazon SageMaker model entity
Here we use `boto3` to create the model entity. Instead of describing a single model, it will indicate the use of multi-model semantics and will identify the source location of all specific model artifacts.
```
# When using multi-model endpoints with the Scikit Learn container, we need to provide an entry point for
# inference that will at least load the saved model. This function uploads a model artifact containing such a
# script. This tar.gz file will be fed to the SageMaker multi-model creation and pointed to by the
# SAGEMAKER_SUBMIT_DIRECTORY environment variable.
def upload_inference_code(script_file_name, prefix):
_tmp_folder = 'inference-code'
if not os.path.exists(_tmp_folder):
os.makedirs(_tmp_folder)
!tar -czvf $_tmp_folder/$USER_CODE_ARTIFACTS $script_file_name > /dev/null
_loc = sagemaker_session.upload_data(_tmp_folder,
key_prefix='{}/{}'.format(prefix, _tmp_folder))
return _loc + '/' + USER_CODE_ARTIFACTS
def create_multi_model_entity(multi_model_name, role):
# establish the place in S3 from which the endpoint will pull individual models
_model_url = 's3://{}/{}/{}/'.format(BUCKET, DATA_PREFIX, MULTI_MODEL_ARTIFACTS)
_container = {
'Image': MULTI_MODEL_SKLEARN_IMAGE,
'ModelDataUrl': _model_url,
'Mode': 'MultiModel',
'Environment': {
'SAGEMAKER_PROGRAM' : SCRIPT_FILENAME,
'SAGEMAKER_SUBMIT_DIRECTORY' : upload_inference_code(SCRIPT_FILENAME, DATA_PREFIX)
}
}
create_model_response = sm_client.create_model(
ModelName = multi_model_name,
ExecutionRoleArn = role,
Containers = [_container])
return _model_url
multi_model_name = '{}-{}'.format(HOUSING_MODEL_NAME, strftime('%Y-%m-%d-%H-%M-%S', gmtime()))
model_url = create_multi_model_entity(multi_model_name, role)
print('Multi model name: {}'.format(multi_model_name))
print('Here are the models that the endpoint has at its disposal:')
!aws s3 ls --human-readable --summarize $model_url
```
### Create the multi-model endpoint
There is nothing special about the SageMaker endpoint config for a multi-model endpoint. You need to consider the appropriate instance type and number of instances for the projected prediction workload. The number and size of the individual models will drive memory requirements.
Once the endpoint config is in place, the endpoint creation is straightforward.
```
endpoint_config_name = multi_model_name
print('Endpoint config name: ' + endpoint_config_name)
create_endpoint_config_response = sm_client.create_endpoint_config(
EndpointConfigName = endpoint_config_name,
ProductionVariants=[{
'InstanceType': ENDPOINT_INSTANCE_TYPE,
'InitialInstanceCount': 1,
'InitialVariantWeight': 1,
'ModelName' : multi_model_name,
'VariantName' : 'AllTraffic'}])
endpoint_name = multi_model_name
print('Endpoint name: ' + endpoint_name)
create_endpoint_response = sm_client.create_endpoint(
EndpointName=endpoint_name,
EndpointConfigName=endpoint_config_name)
print('Endpoint Arn: ' + create_endpoint_response['EndpointArn'])
print('Waiting for {} endpoint to be in service...'.format(endpoint_name))
waiter = sm_client.get_waiter('endpoint_in_service')
waiter.wait(EndpointName=endpoint_name)
```
## Exercise the multi-model endpoint
### Invoke multiple individual models hosted behind a single endpoint
Here we iterate through a set of housing predictions, choosing the specific location-based housing model at random. Notice the cold start price paid for the first invocation of any given model. Subsequent invocations of the same model take advantage of the model already being loaded into memory.
```
def predict_one_house_value(features, model_name):
print('Using model {} to predict price of this house: {}'.format(model_name,
features))
_float_features = [float(i) for i in features]
_body = ','.join(map(str, _float_features)) + '\n'
_start_time = time.time()
_response = runtime_sm_client.invoke_endpoint(
EndpointName=endpoint_name,
ContentType='text/csv',
TargetModel=model_name,
Body=_body)
_predicted_value = json.loads(_response['Body'].read())[0]
_duration = time.time() - _start_time
print('${:,.2f}, took {:,d} ms\n'.format(_predicted_value, int(_duration * 1000)))
# iterate through invocations with random inputs against a random model showing results and latency
for i in range(10):
model_name = LOCATIONS[np.random.randint(1, len(LOCATIONS[:PARALLEL_TRAINING_JOBS]))]
full_model_name = '{}.tar.gz'.format(model_name)
predict_one_house_value(gen_random_house()[1:], full_model_name)
```
### Dynamically deploy another model
Here we demonstrate the power of dynamic loading of new models. We purposely did not copy the first model when deploying models earlier. Now we deploy an additional model and can immediately invoke it through the multi-model endpoint. As with the earlier models, the first invocation to the new model takes longer, as the endpoint takes time to download the model and load it into memory.
```
# add another model to the endpoint and exercise it
deploy_artifacts_to_mme(training_jobs[0])
```
### Invoke the newly deployed model
Exercise the newly deployed model without the need for any endpoint update or restart.
```
print('Here are the models that the endpoint has at its disposal:')
!aws s3 ls $model_url
model_name = LOCATIONS[0]
full_model_name = '{}.tar.gz'.format(model_name)
for i in range(5):
features = gen_random_house()
predict_one_house_value(gen_random_house()[1:], full_model_name)
```
### Updating a model
To update a model, you would follow the same approach as above and add it as a new model. For example, if you have retrained the `NewYork_NY.tar.gz` model and wanted to start invoking it, you would upload the updated model artifacts behind the S3 prefix with a new name such as `NewYork_NY_v2.tar.gz`, and then change the `TargetModel` field to invoke `NewYork_NY_v2.tar.gz` instead of `NewYork_NY.tar.gz`. You do not want to overwrite the model artifacts in Amazon S3, because the old version of the model might still be loaded in the containers or on the storage volume of the instances on the endpoint. Invocations to the new model could then invoke the old version of the model.
Alternatively, you could stop the endpoint and re-deploy a fresh set of models.
## Clean up
Here, to be sure we are not billed for endpoints we are no longer using, we clean up.
```
# shut down the endpoint
sm_client.delete_endpoint(EndpointName=endpoint_name)
# and the endpoint config
sm_client.delete_endpoint_config(EndpointConfigName=endpoint_config_name)
# delete model too
sm_client.delete_model(ModelName=multi_model_name)
```
| github_jupyter |
```
%tensorflow_version 2.x
import tensorflow as tf
# layers
from tensorflow.keras.layers import Input, Dense, Flatten, GRU, LSTM, SimpleRNN, GlobalMaxPooling1D
# models
from tensorflow.keras.models import Sequential, Model
# optimizers
from tensorflow.keras.optimizers import Adam, SGD
# additional imports
import numpy as np # linear algebra
import pandas as pd # data manipulation
import matplotlib.pyplot as plt # data visualisation
### build the dataset
# This is a nonlinear and long-distance dataset
# (Actually, we'll test long-distance vs. short-distance patterns)
# Start with a small T and increase it later
T = 10
D = 1
X = []
Y = []
# We are making a time series signal and
# there is a pattern within this time series signal that leads
# to the classification result. If this pattern is at the end of
# the signal then it doesn't require the RNN to have a long-term memory
# if the pattern is at the beginning of the signal then RNN must
# successfully remember what it saw all the way to the end of the signal
# so the longer the signal is, the longer RNN has to remember.
# generating simple XOR pattern
def get_label(x, i1, i2, i3):
# x = sequence
if x[i1] < 0 and x[i2] < 0 and x[i3] < 0:
return 1
if x[i1] < 0 and x[i2] > 0 and x[i3] > 0:
return 1
if x[i1] > 0 and x[i2] < 0 and x[i3] > 0:
return 1
if x[i1] > 0 and x[i2] > 0 and x[i3] < 0:
return 1
return 0
for t in range(5000):
x = np.random.randn(T)
X.append(x)
y = get_label(x, -1, -2, -3) # short distance
# y = get_label(x, 1, 2, 3) # long distance
Y.append(y)
X = np.array(X)
Y = np.array(Y)
N = len(X)
def get_data(T, L = 5000,data_type = 'long'):
X = []
Y = []
for t in range(L):
x = np.random.randn(T)
X.append(x)
if data_type == 'short':
y = get_label(x, -1, -2, -3) # short distance
if data_type == 'long':
y = get_label(x, 1, 2, 3) # long distance
Y.append(y)
X = np.array(X)
Y = np.array(Y)
N = len(X)
return X,Y,N
# Try a linear model first
linear_model = Sequential([
Input(shape = (T,)),
# it is a classification problem now,
# so we need to map regression results to 0 or 1
Dense(1, activation = 'sigmoid'),
])
linear_model.compile(optimizer = Adam(lr = 0.01),
loss = 'binary_crossentropy',
metrics = ['accuracy'])
r_linear_model = linear_model.fit(X, Y,
validation_split = .5,
epochs = 100)
# plot the loss and accuracy
# loss
plt.plot(r_linear_model.history['loss'], label = 'loss')
plt.plot(r_linear_model.history['val_loss'], label = 'val_loss')
plt.legend()
plt.show()
# accuracy
plt.plot(r_linear_model.history['accuracy'], label = 'accuracy')
plt.plot(r_linear_model.history['val_accuracy'], label = 'val_accuracy')
plt.legend()
plt.show()
# Now let's try RNN
processed_X = np.expand_dims(X, -1) # Make it NxTxD
# SimpleRNN
simplernn = Sequential([
Input(shape = (T,D)),
SimpleRNN(10),
Dense(1, activation = 'sigmoid')
])
# LSTM
lstm = Sequential([
Input(shape = (T,D)),
LSTM(10),
Dense(1, activation = 'sigmoid')
])
# GRU
gru = Sequential([
Input(shape = (T,D)),
GRU(10),
Dense(1, activation = 'sigmoid')
])
simplernn.compile(loss = 'binary_crossentropy',
optimizer = Adam(lr=.01),
metrics = ['accuracy'])
lstm.compile(loss = 'binary_crossentropy',
optimizer = Adam(lr=.01),
metrics = ['accuracy'])
gru.compile(loss = 'binary_crossentropy',
optimizer = Adam(lr=.01),
metrics = ['accuracy'])
r_simplernn = simplernn.fit(processed_X,Y, epochs = 200, validation_split = .5 )
r_lstm = lstm.fit(processed_X,Y, epochs = 200, validation_split = .5 )
r_gru = gru.fit(processed_X,Y, epochs = 200, validation_split = .5 )
# plot the loss and accuracy
# loss
plt.plot(r_simplernn.history['loss'], label = 'loss')
plt.plot(r_simplernn.history['val_loss'], label = 'val_loss')
plt.title('SimpleRNN Loss')
plt.legend()
plt.show()
# accuracy
plt.plot(r_simplernn.history['accuracy'], label = 'accuracy')
plt.plot(r_simplernn.history['val_accuracy'], label = 'val_accuracy')
plt.title('SimpleRNN Accuracy')
plt.legend()
plt.show()
# loss
plt.plot(r_lstm.history['loss'], label = 'loss')
plt.plot(r_lstm.history['val_loss'], label = 'val_loss')
plt.title('LSTM Loss')
plt.legend()
plt.show()
# accuracy
plt.plot(r_lstm.history['accuracy'], label = 'accuracy')
plt.plot(r_lstm.history['val_accuracy'], label = 'val_accuracy')
plt.title('LSTM Accuracy')
plt.legend()
plt.show()
# loss
plt.plot(r_gru.history['loss'], label = 'loss')
plt.plot(r_gru.history['val_loss'], label = 'val_loss')
plt.title('GRU Loss')
plt.legend()
plt.show()
# accuracy
plt.plot(r_gru.history['accuracy'], label = 'accuracy')
plt.plot(r_gru.history['val_accuracy'], label = 'val_accuracy')
plt.title('GRU Accuracy')
plt.legend()
plt.show()
# Now change to the long distance problem
T = 10
D = 1
X = []
Y = []
for t in range(5000):
x = np.random.randn(T)
X.append(x)
y = get_label(x, 1, 2, 3) # long distance
Y.append(y)
X = np.array(X)
Y = np.array(Y)
N = len(X)
# Lets try every model again
processed_X = np.expand_dims(X, -1) # Make it NxTxD
# SimpleRNN
simplernn = Sequential([
Input(shape = (T,D)),
SimpleRNN(10),
Dense(1, activation = 'sigmoid')
])
# LSTM
lstm = Sequential([
Input(shape = (T,D)),
LSTM(10),
Dense(1, activation = 'sigmoid')
])
# GRU
gru = Sequential([
Input(shape = (T,D)),
GRU(10),
Dense(1, activation = 'sigmoid')
])
simplernn.compile(loss = 'binary_crossentropy',
optimizer = Adam(lr=.01),
metrics = ['accuracy'])
lstm.compile(loss = 'binary_crossentropy',
optimizer = Adam(lr=.01),
metrics = ['accuracy'])
gru.compile(loss = 'binary_crossentropy',
optimizer = Adam(lr=.01),
metrics = ['accuracy'])
r_simplernn = simplernn.fit(processed_X,Y, epochs = 200, validation_split = .5 )
r_lstm = lstm.fit(processed_X,Y, epochs = 200, validation_split = .5 )
r_gru = gru.fit(processed_X,Y, epochs = 200, validation_split = .5 )
# plot the loss and accuracy
# loss
plt.plot(r_simplernn.history['loss'], label = 'loss')
plt.plot(r_simplernn.history['val_loss'], label = 'val_loss')
plt.title('SimpleRNN Loss')
plt.legend()
plt.show()
# accuracy
plt.plot(r_simplernn.history['accuracy'], label = 'accuracy')
plt.plot(r_simplernn.history['val_accuracy'], label = 'val_accuracy')
plt.title('SimpleRNN Accuracy')
plt.legend()
plt.show()
# loss
plt.plot(r_lstm.history['loss'], label = 'loss')
plt.plot(r_lstm.history['val_loss'], label = 'val_loss')
plt.title('LSTM Loss')
plt.legend()
plt.show()
# accuracy
plt.plot(r_lstm.history['accuracy'], label = 'accuracy')
plt.plot(r_lstm.history['val_accuracy'], label = 'val_accuracy')
plt.title('LSTM Accuracy')
plt.legend()
plt.show()
# loss
plt.plot(r_gru.history['loss'], label = 'loss')
plt.plot(r_gru.history['val_loss'], label = 'val_loss')
plt.title('GRU Loss')
plt.legend()
plt.show()
# accuracy
plt.plot(r_gru.history['accuracy'], label = 'accuracy')
plt.plot(r_gru.history['val_accuracy'], label = 'val_accuracy')
plt.title('GRU Accuracy')
plt.legend()
plt.show()
# Now let's try with harder dataset
# To make the data difficult, I'll extend sequence length T to 20
# (T = 20)
X, Y, N = get_data(T = 20, data_type = 'long')
processed_X = np.expand_dims(X, -1) # make it NxTxD
# Test all the models again
# SimpleRNN
simplernn = Sequential([
Input(shape = (20,D)), # Now T = 20
SimpleRNN(10),
Dense(1, activation = 'sigmoid')
])
# LSTM
lstm = Sequential([
Input(shape = (20,D)),
LSTM(10),
Dense(1, activation = 'sigmoid')
])
# GRU
gru = Sequential([
Input(shape = (20,D)),
GRU(10),
Dense(1, activation = 'sigmoid')
])
simplernn.compile(loss = 'binary_crossentropy',
optimizer = Adam(lr=.01),
metrics = ['accuracy'])
lstm.compile(loss = 'binary_crossentropy',
optimizer = Adam(lr=.01),
metrics = ['accuracy'])
gru.compile(loss = 'binary_crossentropy',
optimizer = Adam(lr=.01),
metrics = ['accuracy'])
r_simplernn = simplernn.fit(processed_X,Y, epochs = 200, validation_split = .5 )
r_lstm = lstm.fit(processed_X,Y, epochs = 200, validation_split = .5 )
r_gru = gru.fit(processed_X,Y, epochs = 200, validation_split = .5 )
# plot the loss and accuracy
# loss
plt.plot(r_simplernn.history['loss'], label = 'loss')
plt.plot(r_simplernn.history['val_loss'], label = 'val_loss')
plt.title('SimpleRNN Loss')
plt.legend()
plt.show()
# accuracy
plt.plot(r_simplernn.history['accuracy'], label = 'accuracy')
plt.plot(r_simplernn.history['val_accuracy'], label = 'val_accuracy')
plt.title('SimpleRNN Accuracy')
plt.legend()
plt.show()
# loss
plt.plot(r_lstm.history['loss'], label = 'loss')
plt.plot(r_lstm.history['val_loss'], label = 'val_loss')
plt.title('LSTM Loss')
plt.legend()
plt.show()
# accuracy
plt.plot(r_lstm.history['accuracy'], label = 'accuracy')
plt.plot(r_lstm.history['val_accuracy'], label = 'val_accuracy')
plt.title('LSTM Accuracy')
plt.legend()
plt.show()
# loss
plt.plot(r_gru.history['loss'], label = 'loss')
plt.plot(r_gru.history['val_loss'], label = 'val_loss')
plt.title('GRU Loss')
plt.legend()
plt.show()
# accuracy
plt.plot(r_gru.history['accuracy'], label = 'accuracy')
plt.plot(r_gru.history['val_accuracy'], label = 'val_accuracy')
plt.title('GRU Accuracy')
plt.legend()
plt.show()
# Now try LSTM with global max pooling
lstm_gmp2d = Sequential([
Input(shape = (20,D)),
LSTM(5, return_sequences=True),
GlobalMaxPooling1D(),
Dense(1, activation = 'sigmoid')
])
lstm_gmp2d.compile(optimizer = Adam(lr=0.01),
loss = 'binary_crossentropy',
metrics = ['accuracy'])
# Train the RNN
r = lstm_gmp2d.fit(processed_X, Y, validation_split = .5, epochs = 150)
# plot the loss and the accuracy
plt.plot(r.history['loss'], label = 'loss')
plt.plot(r.history['val_loss'], label = 'val_loss')
plt.legend()
plt.show()
# accuracy
plt.plot(r.history['accuracy'], label = 'accuracy')
plt.plot(r.history['val_accuracy'], label = 'val_accuracy')
plt.legend()
plt.show()
# In the end, we can say that LSTM and GRU does a better job
# on long sequences and long distances
# also I used max pooling on LSTM in the last example
# Remember that LSTM outputs last h(t)s,
# If we get every h(t) sequences, we may select only the maximum
# efficient ones. By using max pooling, we select maximum efficient
# h(t) from the sequence. So in the end we pass the best h(t)s we have
# to the Dense layer.
# LSTM with Max Pooling gives the almost same result as others with
# less epochs, so this approach is more powerful than previous ones
```
| github_jupyter |
# Linear Regression and Regularization
Loosely following Chapter 10 of Python Machine Learning 3rd Edition, Raschka
>Disclaimer: Regression is a huge field. It is impossible to cover it all in one class (or even two).
<img src='files/diagrams/ols.png' style='width: 500px'>
[Image Source](https://scikit-learn.org/stable/auto_examples/linear_model/plot_ols.html)
>For a sense of the depth and potential complexity of regression models, see [Mostly Harmless Econometrics](https://www.mostlyharmlesseconometrics.com)
# Ordinary Least Squares (OLS) Linear Regression
>[All models are wrong, but some are useful.](https://en.wikipedia.org/wiki/All_models_are_wrong)
<br><br>George Box
Linear regression is one of the most popular, widely used, and foundational concepts in statistics, econometrics, and machine learning.
Boils down having a numeric target value ($y$) we want to either predict or understand the variance drivers. We use data ($X$) we think impacts our target to understand the underlying **linear** relationship.
### Big Assumption:
- The regression function $E(Y|X)$ is **linear** in the inputs $X_{1},\dots,X_{p}$
- Transformations can be applied to satisfy that requirment.
Typically will see it expressed as $y = \beta X$, or formally:
$$
f(X)=\beta_{0}+\sum{X{j}\beta_{j}}
$$
- Linear model assumes the function is linear or reasonably linear.
- The true $\beta_{j}$'s are unknown parameters (coefficients/weights).
- The features must be able to be represented within a non-null numeric matrix.
### Goal - Minimize the mean-squared error. Why?
Residuals will be positive and negative, need to penalize negative and positive equally.
Sum of errors: $\epsilon_{1} + \epsilon_{2} + \dots + \epsilon_{n}$
RSS: $\epsilon_{1}^2 + \epsilon_{2}^2 + \dots + \epsilon_{n}^2$
MSE: $\frac{1}{N}\sum{\epsilon_{i}^2}$
Most statistical programs solve for the $\beta$ values using plain old linear alegbra, in what is called the closed-form:
$\hat\beta = (X^TX)^{-1}X^{T}y$
### Closed Form Derivation
$$RSS(\beta)=\sum{(y_{i}-f(x_{i}))^2}$$
$$=\sum(y_{i}-\beta_{0}-\sum{x_{ij}\beta{j}})^2$$
$(x_i,y_i)$ should be independent from other $(x_i,y_i)$'s
- Time series models violate this without special treatment
We are seeking a $f(X)$ that minimizes the sum of squared residuals from $Y$:
<img src='files/diagrams/esl-3-1.png' style='width: 500px'>
[Image source: Elements of Statistical Learning, Figure 3.1](https://www.statlearning.com)
$$
RSS(\beta)=(y-X\beta)^T(y-X\beta)
$$
#### Differentiating:
$$
\frac{dRSS}{d\beta}=-2X^T(y-X\beta)
$$
#### And again:
$$
\frac{d^2RSS}{d\beta d \beta^T}=2X^TX
$$
#### Setting the first derivative to zero:
$$
X^T(y-X\beta)=0
$$
#### And we get:
$$
\hat{\beta}=(X^TX)^{-1}X^Ty
$$
#### And our predicted values are:
$$
\hat{y}=X\hat{\beta}
$$
#### And relates to $y$ by:
$$
y = \hat{y} + \epsilon =X\hat{\beta}+\epsilon
$$
> Unique solution means we can derive with pure linear algebra, i.e., no need to use convergence algorithms.
## Slope and Intercept
### Remember in its simple form: $$y=mx+b$$
<img src='files/diagrams/slope.png' style="height: 200px;width: 200px;">
[Image source](https://en.wikipedia.org/wiki/Linear_regression#/media/File:Linear_least_squares_example2.png)
> Since we need an estimate for the intercept, we'll need to manually add the constant. Not all implementations do this automatically, e.g., statsmodels.
- Intercept: where the line go through the y-axis.
- Slope: for a 1-unit change in x, y will increase by $\beta$
# Example - Credit Data
[Data from Elements of Statistical Learning](https://www.statlearning.com)
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
credit = pd.read_csv('data/islr-credit.csv')
credit = credit.iloc[:, 1:]
credit.head()
```
## Find Function so $Rating=f(Limit)$
We'll need to convert the pandas objects to numpy arrays.
```
credit.plot.scatter(x='Limit', y='Rating')
plt.show()
```
## Regression using closed-form:
```
X = np.array(credit['Limit']).reshape(-1,1)
y = np.array(credit['Rating']).reshape(-1,1)
X.shape, y.shape
```
And since we are going to implement a version, we'll need to manually add the constant for the intercept. Why?
$y=\beta_{0}(1)+\beta_{i}x_{i}$
```
from numpy.linalg import inv
'''
- Manually adding the constant
- Sometimes this is done via the API (check the docs)
'''
const = np.ones(shape=y.shape)
mat = np.concatenate( (const, X), axis=1)
# first 5 examples
mat[:5,:]
```
## Betas
We have a feature matrix that has 2 columns, so we'll get estimate for the constant ($\beta_{0}$) and the credit limit ($\beta_{1}$).
### Calculate the coefficient estimates
Recall $\hat\beta = (X^TX)^{-1}X^{T}y$
```
betas = inv(mat.transpose().dot(mat)).dot(mat.transpose()).dot(y)
b0, b1 = betas
print(f'Beta 0: {np.round(b0[0],3)}')
print(f'Beta 1: {np.round(b1[0],3)}')
```
### Predict $\hat{y}$ and plot the fit
$$
\begin{equation}
\hat{y}=\hat{\beta}X=
\hat{\beta_{0}}
\begin{pmatrix}
1 \\
\dots \\
1
\end{pmatrix}+
\hat{\beta_{1}}
\begin{pmatrix}
3606 \\
\dots \\
5524
\end{pmatrix}
\end{equation}
$$
```
yhat = mat.dot(betas)
plt.plot(X, y, 'bo')
plt.plot(X, yhat, 'r')
plt.xlabel('Credit Limit')
plt.ylabel('Credit Rating')
plt.title('$Rating=f(Limit)$', loc='left')
plt.show()
```
# Quantifying fit with metrics
Common metrics:
### $R^2$ [Wikipedia](https://en.wikipedia.org/wiki/Coefficient_of_determination)
$$1 - \frac{\sum (\hat {y}-y)^{2}}{\sum ({\bar y}-y)^{2}}$$
### Mean squared error (MSE) [Wikipedia](https://en.wikipedia.org/wiki/Mean_squared_error)
$$\frac{\sum (\hat {y}-y)^{2}}{n}$$
### Mean Absolute Error (MAE) [Wikipedia](https://en.wikipedia.org/wiki/Mean_absolute_error)
$$1/n\sum |\hat {y}-y|$$
### Root mean squared error (RMSE) [Wikipedia](https://en.wikipedia.org/wiki/Root_mean_square)
$$\sqrt \frac{\sum (\hat {y}-y)^{2}}{n}$$
#### Notes:
- $r^2$ expresses the percent of variance explained (bound between 0% and 100%).
- RMSE expresses the variance in unit terms.
- MSE/MAE are heavily influenced by outliers.
- Usually RMSE is chosen for optimizing since it's an unbiased estimator.
- If there are a lot of outliers, MAE may be a better choice.
### Further reading:
[$r^2$ vs. RMSE](https://www.statology.org/rmse-vs-r-squared/)
# Intrepretability
A nice property of linear regression is the relatively simple intrepretations that can be drawn. The implementation in statsmodels includes all the output needed to evaluate and interpret model output.
[statsmodels OLS regression](https://www.statsmodels.org/stable/regression.html)
```
# betas we calculated:
betas.reshape(1,-1)[0]
```
statsmodels output:
```
import statsmodels.api as smf
simpleModel = smf.OLS(y, mat).fit()
print(simpleModel.summary())
```
## Results Discussion:
- Overall $r^2$ of 99% - very strong linear relationship as we saw in the initial plot we created.
- You can ignore the t-statistic on the constant - it isn't meaningful.
- The t-statistic for the credit limit (x1) is very high (252), with a [p-value](https://en.wikipedia.org/wiki/P-value) of essentially 0 - recall a p-value is the probably that the effect we are seeing is by random chance.
- We can conclude there is a very strong linear relationship.
### Further reading:
[Rethinking p-values](https://www.vox.com/2016/3/15/11225162/p-value-simple-definition-hacking)
[Econometrics](https://en.wikipedia.org/wiki/Econometrics)
# A Note on p-Hacking
<img src='files/diagrams/bad-coef.png' style="width: 500px;">
>Taken from "Having Too Little or Too Much Time Is Linked to Lower Subjective Well-Being", Sharif et al.
# scikit-learn's LinearRegression
[scikit-learn's LinearRegression](https://scikit-learn.org/stable/modules/linear_model.html#ordinary-least-squares)
This will probably be the implementation you'd want to use for comparing various regression models for prediction problems since the API will be similar.
scikit-learn is geared more towards prediction and won't have some of the friendly output that statsmodels has - if you are going for an intrepretation, you may be better off using statsmodels.
```
from sklearn.linear_model import LinearRegression
# we added the intercept manually, so turn that option off
skOLS = LinearRegression(fit_intercept=False).fit(mat,y)
skOLS.coef_
```
#### And they all provide the same coefficient/model estimates - assuming the same set-up.
```
print(betas.reshape(1,-1)[0])
print(simpleModel.params)
print(skOLS.coef_[0])
```
# Weakness - Outlier Sensitivity
There aren't any outliers in the data, I'm going to introduce one.
```
matCopy = mat.copy()
matCopy[4, 1] = matCopy[4, 1] + 150000
outModel = smf.OLS(y, matCopy).fit()
print(outModel.summary())
yhat_out = outModel.predict(matCopy)
plt.plot(matCopy[:,1], y, 'bo')
plt.plot(matCopy[:,1], yhat_out, 'r')
plt.xlabel('Credit Limit')
plt.ylabel('Actual / Expected Credit Rating')
plt.show()
```
## Why the sensitivity?
Recall we are minimizing the sum of squared residuals. That really big outlier is going to a lot of influence.
## How to combat?
- [RANdom SAmple Consensus (RANSAC)](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.RANSACRegressor.html)
- Replace or remove the outliers.
## RANSAC
- Select random samples.
- Tests non-sample points and creates a inlier list (opposite of outlier).
- Refits models with all inliers.
- Evaluates error.
- Stops or repeats until iterations/threshold met.
- **Not guaranteed to get same answer each time* - why?**
> [More details](https://scikit-learn.org/stable/modules/linear_model.html#ransac-regression)
<img src='files/diagrams/ransac.png'>
[Image Source](https://scikit-learn.org/stable/auto_examples/linear_model/plot_ransac.html)
```
from sklearn.linear_model import RANSACRegressor
ransac = RANSACRegressor().fit(matCopy, y)
yhat_ransac = ransac.predict(matCopy)
plt.plot(matCopy[:,1], y, 'bo')
plt.plot(matCopy[:,1], yhat_ransac, 'r')
plt.xlabel('Credit Limit')
plt.ylabel('Actual / Expected Credit Rating')
plt.show()
```
# Strength: Robust to Overfitting
>Simple is better than complex.
### No Overfitting
<img src='files/diagrams/reg.png' style="width: 400px;">
### Severe Overfitting
<img src='files/diagrams/poly.png' style="width: 400px;">
# Multiple Regression
Instead of an $mx1$ input matrix, we'll have $mxn$.
### $y = w_{0} + w_{1}x_{1} + \dots + w_{m}x_{m} = \sum{w_{i}x_{i}}=w^{T}x$
Coefficients still reference the effect on $y$ of a 1-unit change to a $x$ - all else held constant.
Example with the California housing dataset:
## California Housing
```
import pandas as pd
import numpy as np
from sklearn.datasets import fetch_california_housing
from matplotlib import pyplot as plt
import warnings
%matplotlib inline
with warnings.catch_warnings():
warnings.filterwarnings("ignore")
X, y = fetch_california_housing(return_X_y=True, as_frame=True)
x0, x1 = X.shape
print(f'Rows: {x0:,}\nFeatures: {x1}')
california_df = pd.concat([X,y], axis=1)
california_df.head()
```
## Run the regression
Need to add a constant/intercept term manually as statsmodels doesn't handle this automatically.
```
california_df['const'] = 1
featureNames = [x for x in california_df.columns if x != 'MedHouseVal']
featureNames = ['const'] + list(featureNames)
print(featureNames)
```
> Double and triple check that you properly separated your target variable from your features!
```
import statsmodels.api as smf
statsModelsCoefs = smf.OLS(california_df['MedHouseVal'], california_df[featureNames]).fit()
print(statsModelsCoefs.summary())
```
## Check Residuals
```
yhat = statsModelsCoefs.predict(california_df[featureNames])
resid = y - yhat
plt.figure(figsize=(5,5))
plt.plot(y, yhat, 'ro')
plt.xlabel('Actual')
plt.ylabel('Predicted')
plt.title('Actual vs. Predicted')
plt.show()
plt.figure(figsize=(5,5))
plt.hist(resid)
plt.title('Residual Distribution')
plt.show()
```
### Patterns in residuals are not ideal. You'll want these to look like normally distributed white noise (ideally). We might be able to make those residuals behave with feature transformations and other techniques we'll talk about later.
>In certain cases, it may help to log-transform your target variable, which will compress some of the variance.
[Log-linear models](https://en.wikipedia.org/wiki/Log-linear_model)
```
import statsmodels.api as smf
logMV = np.log(california_df['MedHouseVal'])
statsModelsCoefs = smf.OLS(logMV, california_df[featureNames]).fit()
print(statsModelsCoefs.summary())
logyhat = statsModelsCoefs.predict(california_df[featureNames])
logresid = logMV - logyhat
plt.figure(figsize=(5,5))
plt.plot(np.exp(logMV), np.exp(logyhat), 'ro', alpha=0.1)
plt.xlabel('Actual')
plt.ylabel('Predicted')
plt.title('Actual vs. Predicted')
plt.show()
```
> We'll go over models later that can catch the tail of this data better.
# What if we had categorical variables?
## Role of [Dummy Variables](https://en.wikipedia.org/wiki/Dummy_variable_(statistics))
A way to incorporate categorical data into modeling, since models require numerical matrices. Essentially acts to change the intercept.
### Simple Example
```
dummy = pd.DataFrame([[2,4,1],
[3,6,1],
[4,8,1],
[6,12,1],
[7,14,1],
[2,6,0],
[4,10,0],
[6,14,0],
[7,16,0],
[3,8,0]], columns=['hgt', 'wgt', 'label'])
dummy
```
### Persistent differences between these lines. Track parallel to one another.
```
class1 = dummy.query('label==1')
class2 = dummy.query('label==0')
plt.figure(figsize=(6,6))
plt.plot(class1['hgt'], class1['wgt'], 'bo')
plt.plot(class2['hgt'], class2['wgt'], 'ro')
plt.legend(['Label = 1', 'Label = 0'])
plt.show()
```
### Let's look at the means for the data by group
```
dummy.groupby('label')['wgt'].mean().diff()[1]
```
### Compare running a model with and without a dummy variable
```
from sklearn.linear_model import LinearRegression
Xa = np.array(dummy['hgt']).reshape(-1,1)
Xb = np.array(dummy[['hgt','label']])
y = np.array(dummy['wgt']).reshape(-1,1)
bothOLS = LinearRegression().fit(Xa,y)
yhat_both = bothOLS.predict(Xa)
sepOLS = LinearRegression().fit(Xb, y)
yhat_sep = sepOLS.predict(Xb)
print('No Dummy:\n')
print(f' Intercept: {np.round(bothOLS.intercept_[0],2)}')
print(f' Slope: {np.round(bothOLS.coef_[0][0],2)}\n')
print('w/ Dummy:\n')
print(f' Intercept: {np.round(sepOLS.intercept_[0], 2)}')
print(f' Slope: {np.round(sepOLS.coef_[0][0],2)}')
print(f' Dummy: {np.round(sepOLS.coef_[0][1], 0)}')
```
### The dummy captures the mean difference! Otherwise the slope is identical.
If the dummy was present:
$$
y=1.0 + (2)(x_1) + (2)(1)
$$
If the dummy was not present:
$$
y=2.0 + (2)(x_1) + (2)(0)
$$
# Incorporating Categorical Variables into a Model w/ Pipelines
[Example from "Hands-on Machine Learning with Scikit-Learn, Keras & Tensorflow"](https://github.com/ageron/handson-ml2)
This is an expanded, rawer form of the california housing data that is available in scikit-learn.
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
housing = pd.read_csv('data/housing.csv')
housing.info()
```
## Goal - Predict Median House Value
#### Things we need to consider:
- Ocean Proximity is categorical.
- Missing values in Total Bedrooms.
- Data of significantly different scales.
```
housing.median_house_value.hist()
plt.title('Distribution of Median Home Values')
plt.show()
housing.hist(bins=50, figsize=(8,8))
plt.show()
```
Interesting items of note:
- Wide variances in scales.
- Median house value truncated at $500,000
- Outliers are present
```
housing.groupby('ocean_proximity')['median_house_value'].median().plot.barh()
plt.xlabel('Median of Median Home Value')
plt.ylabel('')
plt.title('Price differences by Ocean Proximity')
plt.show()
```
Inland homes have significantly lower prices than homes closer to the water.
```
housing.plot(kind='scatter', x='longitude', y='latitude', alpha=0.4, s=housing['population']/100,
label='population', figsize=(10,7), c='median_house_value', cmap=plt.get_cmap('jet'),
colorbar=True)
plt.legend('')
plt.show()
```
Higher values are largely clustered around the coast.
[Example from "Hands-on Machine Learning with Scikit-Learn, Keras & TensorFlow](https://github.com/ageron/handson-ml2)
# Missing Values
#### Options:
- Drop rows (can be bad, what if incoming data has missing values?).
- Drop columns (could be bad, what if there value in that feature?).
- Fill in the missing values (ding ding).
- If categorical, might want to add a dummy to indicate it was missing (ding ding).
- Best strategy will be situationally dependent. This can be treated as a hyperparameter - no perfect answer.
#### Strategies:
- Simple inputers with median, median, mode, random values.
- Estimate the missing value with another machine learning model (increasing overal complexity).
>Not all strategies will work for all data types, so you may need to split it up, e.g., one method for the numerical variables and another for the categorical variables.
```
housing.isna().sum()
```
### Filling with median
```
from sklearn.impute import SimpleImputer
example_imputer = SimpleImputer(strategy='median')
example_imputer.fit_transform(np.array(housing.total_bedrooms).reshape(-1,1))
example_imputer = pd.Series(example_imputer)
example_imputer.isna().sum()
```
### If you wanted to add an indictor for the missing value
> Probably more useful for categorical variables
```
from sklearn.impute import SimpleImputer
example_imputer = SimpleImputer(strategy='median', add_indicator=True)
example_imputer.fit_transform(np.array(housing.total_bedrooms).reshape(-1,1))
```
### Note on One-Hot Encoding
From "Hands-on Machine Learning with Scikit-Learn, Keras & TensorFlow":
> If a categorical attribute has a large number of possible categories, then one-hot encoding will result in a large number of input features. This may slow down training and degrade performance.
#### Possible alternatives in that situation:
- Recode to a numerical feature, e.g., distance to ocean.
- Only use the most frequent $N$ categories.
- Convert to embeddings.
#### A risk for us:
May not have any islands in the training data, what would happen if we encountered that in our test/evaluation data?
```
housing.ocean_proximity.value_counts()
```
### Islands are really rare
- Adding a dummy for this won't do much - it'll basically be zero.
- Replace to nearest category?
```
from sklearn.preprocessing import OneHotEncoder
example_ohe = OneHotEncoder()
example_ohe = example_ohe.fit_transform(np.array(housing['ocean_proximity']).reshape(-1,1))
example_ohe
```
> Sparse matrix returns lists of coordinates in the matrix with a non-zero. It's a more efficient structure:
```
print(example_ohe[:5,])
```
>Can be converted back to a dense format:
```
example_ohe.toarray()[:5,:]
```
### Should we use this for modeling?
>In statistics, multicollinearity (also collinearity) is a phenomenon in which one predictor variable in a multiple regression model can be linearly predicted from the others with a substantial degree of accuracy. In this situation, the coefficient estimates of the multiple regression may change erratically in response to small changes in the model or the data. Multicollinearity does not reduce the predictive power or reliability of the model as a whole, at least within the sample data set; it only affects calculations regarding individual predictors. That is, a multivariate regression model with collinear predictors can indicate how well the entire bundle of predictors predicts the outcome variable, but it may not give valid results about any individual predictor, or about which predictors are redundant with respect to others.
<br><br>[Wikipedia](https://en.wikipedia.org/wiki/Multicollinearity)
### Could argue that this could be represented as an ordinal variable (island > near ocean > near bay > ...)
See [OrdinalEncoder for as example](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.OrdinalEncoder.html)
You could try this using both methods to see if one is better.
# A Brief Rant on LabelEncoder
Should not be used on your feature set! This is commonly done on Kaggle (incorrectly!). Do not use this for your feature processing!
[LabelEncoder Documentation](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.LabelEncoder.html?highlight=labelencoder)
# Scaling Numerical Variables
Don't skip this - most models don't perform well when variables are on different scales.
```
housing.select_dtypes(['float','integer']).describe().round(0)
```
## Two main methods:
> Only fit these to the training data, no leaking information from the test set!
### Min-max scaling
- Simple
- Bound between 0 and 1 - a lot of algorithms like that, especially neural networks
- scikit-learn gives you some additional flexibility in terms of the range
- Very susceptible to outliers
$$
x_{scaled} = \frac{x - x_{min}}{x_{max}-x_{min}}
$$
### Standardization
- Little more involved.
- More robust to outliers.
- No specific range boundary.
$$
x_{scaled} = \frac{x - \hat{x}}{\sigma_{x}}
$$
Since we are doing regression and don't have a scaling boundary requirement and there are probably outliers, we'll use standardization.
```
from sklearn.preprocessing import StandardScaler
import matplotlib.pyplot as plt
example_SS = StandardScaler()
example_SS = example_SS.fit_transform(np.array(housing.total_rooms).reshape(-1,1))
plt.hist(housing.total_rooms, bins=100)
plt.title('Total Rooms', loc='left')
plt.show()
plt.hist(example_SS, bins=100)
plt.title('Total Rooms - Standardized', loc='left')
plt.show()
```
# Training/Test Splits
>Put the test data aside and never look at it again.
- All of the feature transformations and model training should be on the training data.
- In production, you wouldn't exactly know what the incoming data would look like ahead of time.
- If you use the test data to inform **any** of the feature transformations or modeling, then you are letting that test data leak into the training data and that will (may) bias your evaluations. This is called **leakage** or **data snooping** - both are not good.
### Simpliest form is splitting your data into two parts:
- Training: will base feature transforms and modeling on this.
- Test: evaluate the models on this data.
>There are more robust methods that we'll talk about later. You can think of this simple splitting as a quick and dirty way to evaluate performance, but it isn't a methodology you'd want to use to estimate what your performance is truly likely to be.
### Split off the features and the target variable
```
y = housing.median_house_value
features = ['housing_median_age', 'total_rooms', 'total_bedrooms',
'population', 'households', 'median_income', 'ocean_proximity', 'longitude', 'latitude'
]
X = housing[features]
X.info()
```
### Split into training and test sets
>80/20 split is pretty standard, but not universal. For very large datasets, I've heard of 99/1 splits.
```
from sklearn.model_selection import train_test_split
X_training, X_test, y_training, y_test = train_test_split(X, y, test_size=0.20)
print(f'Training samples: {X_training.shape[0]:,}')
print(f'Test samples: {X_test.shape[0]:,}')
```
>Remember, the test data is only for evaluation.
```
X_training.info()
```
# Pipelines
- Fill missing values.
- Create dummies for categorical.
- Standardize numerical variables.
- Fit the model.
### Pipelines can be made of collections of pipelines
```
from sklearn.pipeline import Pipeline
from sklearn.compose import ColumnTransformer
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import OneHotEncoder
from sklearn.preprocessing import StandardScaler
cat_vars = ['ocean_proximity']
num_vars = ['housing_median_age', 'total_rooms', 'total_bedrooms', 'population',
'households', 'median_income', 'longitude', 'latitude']
num_pipeline = Pipeline([('impute_missing', SimpleImputer(strategy='median')),
('standardize_num', StandardScaler())
])
cat_pipeline = Pipeline([('impute_missing_cats', SimpleImputer(strategy='most_frequent')),
('create_dummies_cats', OneHotEncoder(handle_unknown='ignore', drop='first'))])
processing_pipeline = ColumnTransformer(transformers=[('proc_numeric', num_pipeline, num_vars),
('create_dummies', cat_pipeline, cat_vars)])
print(processing_pipeline)
from sklearn.linear_model import LinearRegression
modeling_pipeline = Pipeline([('data_processing', processing_pipeline), ('lm', LinearRegression())])
modeling_pipeline.fit(X_training, y_training)
```
# Evaluating the model
>Really evaluatign the entire preprocessing process and the model itself.
```
housing_predictions = modeling_pipeline.predict(X_test)
```
### Get the mean squared error, root mean squared error, and $R^2$
```
from sklearn.metrics import mean_squared_error
mse = mean_squared_error(y_test, housing_predictions)
mse
rmse = np.sqrt(mse)
rmse
from sklearn.metrics import r2_score
r2 = r2_score(y_test, housing_predictions)
r2
```
### Plot the test and predictions
```
import matplotlib.pyplot as plt
plt.plot(y_test, housing_predictions, 'ro')
plt.xlabel('Actual')
plt.ylabel('Predicted')
plt.show()
```
# Observations
- If this was a perfect model, the residuals would be 0 for all actual/predicted values.
- Residuals should look like white noise across all values - seeing some patterns.
- Some information is leaking into the residuals that the model is capturing.
- Could be a feature we don't have access to.
- Could be noise in the data.
- Could be the underlying relationships are linear.
- Insert any number of additional explanations.
There may be some overfitting - the training data fits better than the test data. We can explore other models to see if they are able to reduce the overfitting.
# Bias-variance Tradeoff
<img src='files/diagrams/bullseye.png' style="width: 600px;">
[Image source](http://scott.fortmann-roe.com/docs/BiasVariance.html)
>At its root, dealing with bias and variance is really about dealing with **over- and under-fitting**. Bias is reduced and variance is increased in relation to model complexity. As more and more parameters are added to a model, the complexity of the model rises and variance becomes our primary concern while bias steadily falls
<br><br>Understanding the Bias-Variance Tradeoff, Fortmann-Roe
From Raschka (paraphrased):
>Variance measures the consistency (or variability) of the model prediction. If we retrain the model on different subsets of the training get and observe difference results, we say it is subject to high variance.
>Bias measures how far off the predictions are from the correct values. This will be error that isn't due to differences in the training datasets.
[Bias and variance from Raschka's Evaluation Lecture Notes](https://sebastianraschka.com/pdf/lecture-notes/stat479fs18/08_eval-intro_notes.pdf)
<img src='files/diagrams/high-bias.png' style="width: 600px;">
<img src='files/diagrams/high-variance.png' style="width: 600px;">
# Simple Usually Triumphs Over the Complex
It's a balancing act though. You'll need a minimum level of complexity to capture the relationships in the data.
<img src='files/diagrams/biasvariance.png'>
[Image source](http://scott.fortmann-roe.com/docs/BiasVariance.html)
## Potential Options
- Include less features.
- Shrinkage methods.
- Data over models.
# Low Variance, Forward, and Backward Selection
## [Low Variance](https://scikit-learn.org/stable/modules/feature_selection.html)
>You can automatically omit features with zero-variance (i.e., constants).
```
from sklearn.feature_selection import VarianceThreshold
X = np.array([[0, 0, 1], [0, 1, 0], [1, 0, 0], [0, 1, 1], [0, 1, 0], [0, 1, 1]])
print(X)
sel = VarianceThreshold(threshold=(.8 * (1 - .8)))
sel.fit_transform(X)
```
## [Forward or Backward Selection](https://scikit-learn.org/stable/modules/generated/sklearn.feature_selection.SequentialFeatureSelector.html#sklearn.feature_selection.SequentialFeatureSelector)
From scikit-learn:
>Forward-SFS is a greedy procedure that iteratively finds the best new feature to add to the set of selected features. Concretely, we initially start with zero feature and find the one feature that maximizes a cross-validated score when an estimator is trained on this single feature. Once that first feature is selected, we repeat the procedure by adding a new feature to the set of selected features. The procedure stops when the desired number of selected features is reached, as determined by the n_features_to_select parameter.
>Backward-SFS follows the same idea but works in the opposite direction: instead of starting with no feature and greedily adding features, we start with all the features and greedily remove features from the set. The direction parameter controls whether forward or backward SFS is used.
<br><br>In general, forward and backward selection do not yield equivalent results. Also, one may be much faster than the other depending on the requested number of selected features: if we have 10 features and ask for 7 selected features, forward selection would need to perform 7 iterations while backward selection would only need to perform 3.
>SFS differs from RFE and SelectFromModel in that it does not require the underlying model to expose a coef_ or feature_importances_ attribute. It may however be slower considering that more models need to be evaluated, compared to the other approaches. For example in backward selection, the iteration going from m features to m - 1 features using k-fold cross-validation requires fitting m * k models, while RFE would require only a single fit, and SelectFromModel always just does a single fit and requires no iterations.
[See a *Comparative Study of Techniques for Large-Scale Feature Selection* for more discussion.](https://citeseerx.ist.psu.edu/viewdoc/download?doi=10.1.1.24.4369&rep=rep1&type=pdf)
### Basic Idea
- Start with all the features.
- Determine the feature provided the least added benefit.
- Remove the above feature.
- Continue until reach the desired number of features or hit a threshold.
### Rational
> Automatically select the most relevant subset of features.
> Really only necessary if your models don't support regularization.
> Can help with the *Curse of Dimensionality* since it'll generally select a more parsimonious model with dense features.
From Machine Learning with Python, SBS (backward) showed a model with 3 features would have achieved 100% accuracy on the validation data set. See pages 137-139.
<img src='files/diagrams/04_08.png' style='width: 300px;'>
[Image source - Raschka's GitHub; Python for Machine Learning 3rd Edition, Figure 4.8](https://github.com/rasbt/python-machine-learning-book-3rd-edition/blob/master/ch04/images/04_08.png)
- I've never used these in practice - there are other ways to guard against overfitting and selecting a more parsimonious model.
- Can add a lot of computational overhead.
[See an example from scikit-learn for code example.](https://scikit-learn.org/stable/auto_examples/feature_selection/plot_select_from_model_diabetes.html#sphx-glr-auto-examples-feature-selection-plot-select-from-model-diabetes-py)
# Regularization
>Helps solve overfitting (high variance) - having too many parameters (i.e., too complex).
>Also helps with multicolinearity (which we saw) and filtering out noise.
### Types of regularization:
- $L1$ (lasso - least absolute shrinkage and selection operator): penalizing the sum of $\lvert \beta_{j} \rvert$
- $L2$ (ridge): penalizing the sum of the $\beta_{j}$'s
# Shrinkage with Ridge Regression ($l_2$ Regularization)
## We can modify our loss function to penalize complexity
Shrinking parameter values to penalize for increased complexity (i.e., parameters with meaningful weights).
>Shrinks, but does not eliminate. Coefficients will be non-zero. Will that help when there any many many features?
## Reduces the influence of parameters that carry less weight
Recall $\hat\beta = (X^TX)^{-1}X^{T}y$
Goal: $\frac{1}{N}\sum{\epsilon_{i}^2}$
#### We can force parameters to shrink towards zero, effectively reducing their influence by adding a penality to the loss function:
$argmin\frac{1}{N}\sum{\epsilon_{i}^2} + \sum{\lambda\beta_{j}^2}$
- What happens with $\lambda=0$?
New closed form: $\hat\beta = (X^TX+\lambda{I})^{-1}X^{T}y$
> Ridge has a unique solution, similar to OLS. You can modify the closed form implementation from last class as an independent proof. scikit-learn's implementation uses various optimization methods to solve the loss optimization problem, so it won't be 100% comparable to OLS with $\alpha=0$, it'll be close though.
## Introduces $\lambda$ - our first (official) hyperparameter!
$\lambda$ controls the amount of the penality, it is bounded between 0 and $\infty$.
>When $\lambda=0$, ridge will provide the same coefficients as OLS, since the $\lambda{I}$ will become zero.
```
from sklearn.linear_model import Ridge
modeling_pipeline_ridge = Pipeline([('data_processing', processing_pipeline), ('ridge', Ridge(alpha=0))])
modeling_pipeline_ridge.fit(X_training, y_training)
modeling_pipeline_ridge['ridge'].coef_
```
> Compared to the below from the original model:
```
modeling_pipeline['lm'].coef_
```
# Now to evaluate different $\lambda$ values
>We'll need to evaluate a gradient of $\lambda$ values to determine the best one to use.
```
from collections import defaultdict
alphas = [0, 1, 2, 5, 10, 50]
ridge_results = defaultdict(dict)
for alph in alphas:
modeling_pipeline_ridge = Pipeline([('data_processing', processing_pipeline), ('ridge', Ridge(alpha=alph))])
modeling_pipeline_ridge.fit(X_training, y_training)
ridge_results['coefficients'][alph] = modeling_pipeline_ridge['ridge'].coef_
ridge_results['training score'][alph] = modeling_pipeline_ridge.score(X_training, y_training)
ridge_results['test score'][alph] = modeling_pipeline_ridge.score(X_test, y_test)
print('Done')
coefficients_ridge = pd.DataFrame.from_dict(ridge_results['coefficients'])
coefficients_ridge = coefficients_ridge.reset_index()
coefficients_ridge = coefficients_ridge.rename(columns={'index':'coefficient_nbr'})
coefficients_ridge = coefficients_ridge.melt(id_vars='coefficient_nbr', var_name='alpha', value_name='coefficient')
(
coefficients_ridge.pivot_table(index='alpha', columns='coefficient_nbr', values='coefficient')
.plot(figsize=(8,4),legend=False)
)
plt.title('Ridge Coefficients', loc='left')
plt.xlabel('Alpha (Regularization Amount)')
plt.ylabel('Coefficient')
plt.show()
```
### Changes in $R^2$
```
ridge_training_r2 = pd.Series(ridge_results['training score'])
ridge_test_r2 = pd.Series(ridge_results['test score'])
ridge_training_r2.plot()
ridge_test_r2.plot()
plt.title('$R^2$ for Ridge Regression')
plt.legend(['Training','Test'])
plt.xlabel('Alpha (Regularization Level)')
plt.ylabel('Percent of Variance Explained')
plt.ylim(0.4, 1)
plt.show()
```
# Another Option is Lasso - Requires Gradient Descent (GD)
OLS and Ridge regression have unique solutions (even though scikit-learn uses optimization). In order to talk about some of the other variants, we need to talk about an optimization technique called gradient descent.
Gradient descent is an optimization technique that allows us to "learn" what the coefficients should be by iteration and continuous improvment. Traditional statisticans don't love this technique since it's not too far away from guessing a bunch of times until you can't guess much better.
<img src='files/diagrams/gs.png' style="width: 300px;">
[Image source](https://www.google.com/url?sa=i&url=https%3A%2F%2Fwww.datasciencecentral.com%2Fprofiles%2Fblogs%2Falternatives-to-the-gradient-descent-algorithm&psig=AOvVaw1ki8gWYTrWRy-NKpu7RFgo&ust=1631036623607000&source=images&cd=vfe&ved=0CAsQjRxqFwoTCJiSq6nz6vICFQAAAAAdAAAAABAD)
Gradient descent essentially is looking for the local minimums of loss functions that are differentiable.
While least-squares does not a closed-form solution, you can also approximate it using gradient descent. Gradient descent will reappear with other algorithms.
GD requires a loss function, which for OLS regression is the sum of squared errors:
$$J(w)=\frac{1}{2}\sum(y^{(i)} - \hat{y}^{(i)})^2$$
This also be used in logistic regression and neural networks.
## Updating weights
All of the weights are set simultaneously.
1. Initialize weights to 0 or small random numbers.
2. For each training example, $x^{i}$:
a. Compute the output value, $\hat{y}$.
b. Update the weights.
```
def gradientDescent(x, y, theta, alpha, m, numIterations):
thetaHistory = list()
xTrans = x.transpose()
costList = list()
for i in range(0, numIterations):
# data x feature weights = y_hat
hypothesis = np.dot(x, theta)
# how far we are off
loss = hypothesis - y
# mse
cost = np.sum(loss ** 2) / (2 * m)
costList.append(cost)
# avg gradient per example
gradient = np.dot(xTrans, loss) / m
# update
theta = theta - alpha * gradient
thetaHistory.append(theta)
return thetaHistory, costList
```
### Create training data
```
data_pipeline = Pipeline([('data_processing', processing_pipeline)])
data_pipeline.fit(X_training)
gs_training_data = data_pipeline.fit_transform(X_training)
```
### Run the model
```
import datetime
start_ts = datetime.datetime.now()
betaHistory, costList = gradientDescent(gs_training_data,y_training,
theta=np.zeros(gs_training_data.shape[1]),
alpha=0.01,
m=gs_training_data.shape[0], numIterations=5000)
end_ts = datetime.datetime.now()
print(f'Completed in {end_ts-start_ts}')
plt.plot(costList)
plt.title(f'Final cost: {costList[-1]:,.2f}', loc='left')
plt.show()
```
## Show changes in $\beta$ throughout the iterations
```
from collections import defaultdict
thetas = defaultdict(list)
for i in range(len(betaHistory)):
for j in range(len(betaHistory[i])):
thetas[j].append(betaHistory[i][j])
thetasD = pd.DataFrame.from_dict(thetas)
thetasD.plot(legend=False)
plt.title('Beta Estimates')
plt.ylabel('Coefficient')
plt.xlabel('Iteration')
plt.show()
```
### Predictions
```
gs_betas = betaHistory[4999]
gs_predictions = np.dot(gs_training_data, gs_betas)
plt.plot(y_training, gs_predictions, 'bo', alpha=0.4)
plt.xlabel('Actual')
plt.ylabel('Predicted')
plt.title('Gradient Descent Regression Fit on Training Data')
plt.show()
```
# Where things can go wrong:
- The learning rate (alpha generally) is really important. If you pick a rate that is too large, you may hop over the minimum and the models will either be very poor or never converge.
- Other thresholds may take some trial and error.
- You can get stuck at a local minima and never find the local maxima.
- You need to know when to stop. It'll keep adjusting coefficients until it reaches an iteration limit or a derivative threshold. Too long and overfitting could occur.
# Lasso Regression ($l_1$ Regularization)
No closed form solution - will need to use optimization techniques regardless.
$$
J(w)_{lasso}=\sum{(y^{(i)}-\hat{y}^{(i)})^2+\lambda \lvert \lvert w \rvert \rvert_1}
$$
$$
L1:\lambda \lvert \lvert w \rvert \rvert_1 = \lambda \sum{\lvert w_j \rvert}
$$
Could modify the gradient descent with the above, but we'll let sci-kit learn handle it.
## Differences from Ridge
<img src='files/diagrams/ridge-v-lasso.png'>
[Introduction to Statistical Learning, Figure 3.11](https://www.statlearning.com)
>Estimation picture for the lasso (left) and ridge regression (right). Shown are contours of the error and constraint functions. The solid blue areas are the constraint regions $\lvert \beta_1 \rvert \leq t$ and $\beta_{1}^2 + \beta_{2}^2 \leq t^2$, respectivity, while the red ellipses are the contours of the least squares error function.
#### Explanation from Raschka (Python Machine Learning 3rd Edition, Chapter 4, pages 129-131):
> We can think of regularization as adding a penalty term to the cost function to encourage smaller weighs; in other words, we penalize large weights. Thus, by increasing the regularization strength via the regularization parameter, we shrink the weights toward zero and decrease the dependence of our model on the training data.
> The shaded regions of represent the regularization "budget" - the combination of the weights cannot exceed those limits. As the regularization term increases, so does the area of that shaded region.
[See Elements of Statistical Learning Section 3.4 for a more thorough discussion.](https://web.stanford.edu/~hastie/ElemStatLearn/)
### TL:DR - Lasso can create sparse models, Ridge cannot.
Ridge will have non-zero estimates for its $\beta$ values, and lasso can result in some $\beta$ values equal to zero (i.e., sparse).
- Lasso should provide better protection to overfitting than Ridge and OLS.
- Can also be a technique by itself for feature selection.
```
from sklearn.linear_model import Lasso
from collections import defaultdict
alphas = [1, 2, 5, 10, 50]
lasso_results = defaultdict(dict)
for alph in alphas:
modeling_pipeline_lasso = Pipeline([('data_processing', processing_pipeline), ('lasso', Lasso(alpha=alph))])
modeling_pipeline_lasso.fit(X_training, y_training)
lasso_results['coefficients'][alph] = modeling_pipeline_lasso['lasso'].coef_
lasso_results['training score'][alph] = modeling_pipeline_lasso.score(X_training, y_training)
lasso_results['test score'][alph] = modeling_pipeline_lasso.score(X_test, y_test)
coefficients_lasso = pd.DataFrame.from_dict(lasso_results['coefficients'])
coefficients_lasso = coefficients_lasso.reset_index()
coefficients_lasso = coefficients_lasso.rename(columns={'index':'coefficient_nbr'})
coefficients_lasso = coefficients_lasso.melt(id_vars='coefficient_nbr', var_name='alpha', value_name='coefficient')
coefficients_lasso.pivot_table(index='alpha', columns='coefficient_nbr', values='coefficient').plot(figsize=(8,4))
plt.title('Lasso Coefficients', loc='left')
plt.xlabel('Alpha (Regularization Amount)')
plt.ylabel('Coefficient')
plt.legend('')
plt.show()
coefficients_ridge.query('coefficient_nbr == 9')
coefficients_lasso.query('coefficient_nbr == 9')
```
> Lasso shrunk to $0$ and ridge did not!
# Reading
[Spurious correlations (High $r^2$ useless relationships)](https://www.tylervigen.com/spurious-correlations)
<br>[Mostly Harmless Econometrics](https://www.mostlyharmlesseconometrics.com)
<br>[Additional examples from statsmodels](https://www.statsmodels.org/stable/examples/notebooks/generated/ols.html)
<br> [Common pitfalls](https://scikit-learn.org/stable/auto_examples/inspection/plot_linear_model_coefficient_interpretation.html)
<br>[Andrew Ng on Regularization](https://www.youtube.com/watch?v=NyG-7nRpsW8&list=PLkDaE6sCZn6Hn0vK8co82zjQtt3T2Nkqc&index=6)
<br>[Great notes from Goker Erdogan](https://github.com/gokererdogan/JaverianaMLCourse/blob/master/Lectures/05.pdf).
<br>[Raschka on Regularization Performance](https://sebastianraschka.com/faq/docs/regularized-logistic-regression-performance.html)
<br>[Raschka's Linear Regression Notes](http://rasbt.github.io/mlxtend/user_guide/regressor/LinearRegression/)
From Raschka's post:
>Regularization does NOT improve the performance on the data set that the algorithm used to learn the model parameters (feature weights). However, it can improve the generalization performance, i.e., the performance on new, unseen data, which is exactly what we want.
| github_jupyter |
<h1>Script-mode Custom Training Container</h1>
This notebook demonstrates how to build and use a custom Docker container for training with Amazon SageMaker that leverages on the <strong>Script Mode</strong> execution that is implemented by the sagemaker-training-toolkit library. Reference documentation is available at https://github.com/aws/sagemaker-training-toolkit
We start by defining some variables like the current execution role, the ECR repository that we are going to use for pushing the custom Docker container and a default Amazon S3 bucket to be used by Amazon SageMaker.
```
import boto3
import sagemaker
from sagemaker import get_execution_role
ecr_namespace = 'sagemaker-training-containers/'
prefix = 'script-mode-container'
ecr_repository_name = ecr_namespace + prefix
role = get_execution_role()
account_id = role.split(':')[4]
region = boto3.Session().region_name
sagemaker_session = sagemaker.session.Session()
bucket = sagemaker_session.default_bucket()
print(account_id)
print(region)
print(role)
print(bucket)
```
Let's take a look at the Dockerfile which defines the statements for building our script-mode custom training container:
```
! pygmentize ../docker/Dockerfile
```
At high-level the Dockerfile specifies the following operations for building this container:
<ul>
<li>Start from Ubuntu 16.04</li>
<li>Define some variables to be used at build time to install Python 3</li>
<li>Some handful libraries are installed with apt-get</li>
<li>We then install Python 3 and create a symbolic link</li>
<li>We install some Python libraries like numpy, pandas, ScikitLearn, etc.</li>
<li>We set e few environment variables, including PYTHONUNBUFFERED which is used to avoid buffering Python standard output (useful for logging)</li>
<li>We install the <strong>sagemaker-training-toolkit</strong> library</li>
<li>Finally, we copy all contents in <strong>code/</strong> (which is where our training code is) under <strong>/opt/ml/code/</strong> which is the path where sagemaker-training-toolkit expects to find training code</li>
</ul>
<h3>Build and push the container</h3>
We are now ready to build this container and push it to Amazon ECR. This task is executed using a shell script stored in the ../script/ folder. Let's take a look at this script and then execute it.
```
! pygmentize ../scripts/build_and_push.sh
```
<h3>--------------------------------------------------------------------------------------------------------------------</h3>
The script builds the Docker container, then creates the repository if it does not exist, and finally pushes the container to the ECR repository. The build task requires a few minutes to be executed the first time, then Docker caches build outputs to be reused for the subsequent build operations.
```
%%capture
! ../scripts/build_and_push.sh $account_id $region $ecr_repository_name
```
<h3>Training with Amazon SageMaker</h3>
Once we have correctly pushed our container to Amazon ECR, we are ready to start training with Amazon SageMaker, which requires the ECR path to the Docker container used for training as parameter for starting a training job.
```
container_image_uri = '{0}.dkr.ecr.{1}.amazonaws.com/{2}:latest'.format(account_id, region, ecr_repository_name)
print(container_image_uri)
```
Given the purpose of this example is explaining how to build custom script-mode containers, we are not going to train a real model. The script that will be executed does not define a specific training logic; it just outputs the configurations injected by SageMaker and implements a dummy training loop. Training data is also dummy. Let's analyze the script first:
```
! pygmentize ../docker/code/train.py
```
You can realize that the training code has been implemented as a standard Python script, that will be invoked by the sagemaker-training-toolkit library passing hyperparameters as arguments. This way of invoking training script is indeed called <strong>Script Mode</strong> for Amazon SageMaker containers.
Now, we upload some dummy data to Amazon S3, in order to define our S3-based training channels.
```
! echo "val1, val2, val3" > dummy.csv
print(sagemaker_session.upload_data('dummy.csv', bucket, prefix + '/train'))
print(sagemaker_session.upload_data('dummy.csv', bucket, prefix + '/val'))
! rm dummy.csv
```
Finally, we can execute the training job by calling the fit() method of the generic Estimator object defined in the Amazon SageMaker Python SDK (https://github.com/aws/sagemaker-python-sdk/blob/master/src/sagemaker/estimator.py). This corresponds to calling the CreateTrainingJob() API (https://docs.aws.amazon.com/sagemaker/latest/dg/API_CreateTrainingJob.html).
```
import sagemaker
import json
# JSON encode hyperparameters
def json_encode_hyperparameters(hyperparameters):
return {str(k): json.dumps(v) for (k, v) in hyperparameters.items()}
hyperparameters = json_encode_hyperparameters({
"hp1": "value1",
"hp2": 300,
"hp3": 0.001})
est = sagemaker.estimator.Estimator(container_image_uri,
role,
train_instance_count=1,
train_instance_type='local', # we use local mode
#train_instance_type='ml.m5.xlarge',
base_job_name=prefix,
hyperparameters=hyperparameters)
train_config = sagemaker.session.s3_input('s3://{0}/{1}/train/'.format(bucket, prefix), content_type='text/csv')
val_config = sagemaker.session.s3_input('s3://{0}/{1}/val/'.format(bucket, prefix), content_type='text/csv')
est.fit({'train': train_config, 'validation': val_config })
```
| github_jupyter |
# Tutorial Part 15: Synthetic Feasibility
Synthetic feasibility is a problem when running large scale enumerations. Ofen molecules that are enumerated are very difficult to make and thus not worth inspection even if their other chemical properties are good in silico. This tutorial goes through how to train the [ScScore](https://pubs.acs.org/doi/abs/10.1021/acs.jcim.7b00622) model [1].
The idea of the model is to train on pairs of molecules where one molecule is "more complex" than the other. The neural network then can make scores which attempt to keep this pairwise ordering of molecules. The final result is a model which can give a relative complexity of a molecule.
The paper trains on every reaction in reaxys, declaring products more complex than reactions. Since this training set is prohibitively expensive we will instead train on arbitrary molecules declaring one more complex if it's SMILES string is longer. In the real world you can use whatever measure of complexity makes sense for the project.
In this tutorial, we'll use the Tox21 dataset to train our simple synthetic feasibility model.
## Colab
This tutorial and the rest in this sequence are designed to be done in Google colab. If you'd like to open this notebook in colab, you can use the following link.
[](https://colab.research.google.com/github/deepchem/deepchem/blob/master/examples/tutorials/15_Synthetic_Feasibility_Scoring.ipynb)
## Setup
We recommend you run this tutorial on Google colab. You'll need to run the following commands to set up your colab environment to run the notebook.
```
%tensorflow_version 1.x
!curl -Lo deepchem_installer.py https://raw.githubusercontent.com/deepchem/deepchem/master/scripts/colab_install.py
import deepchem_installer
%time deepchem_installer.install(version='2.3.0')
import deepchem as dc
# Lets get some molecules to play with
from deepchem.molnet.load_function import tox21_datasets
tasks, datasets, transformers = tox21_datasets.load_tox21(featurizer='Raw', split=None, reload=False)
molecules = datasets[0].X
```
# Make The Datasets
Because ScScore is trained on relative complexities we have our `X` tensor in our dataset has 3 dimensions `(sample_id, molecule_id, features)`. the 1st dimension molecule_id is in \[0, 1\], because a sample is a pair of molecules. The label is 1 if the zeroth molecule is more complex than the first molecule. The function `create_dataset` we introduce below pulls random pairs of smiles strings out of a given list and ranks them according to this complexity measure.
In the real world you could use purchase cost, or number of reaction steps required as your complexity score.
```
from rdkit import Chem
import random
from deepchem.feat import CircularFingerprint
import deepchem as dc
import numpy as np
def create_dataset(fingerprints, smiles_lens, ds_size=100000):
"""
m1: list of np.Array
fingerprints for molecules
m2: list of int
length of a molecules SMILES string
returns:
dc.data.Dataset for input into ScScore Model
Dataset.X
shape is (sample_id, molecule_id, features)
Dataset.y
shape is (sample_id,)
values is 1 if the 0th index molecule is more complex
0 if the 1st index molecule is more complex
"""
X, y = [], []
all_data = list(zip(fingerprints, smiles_lens))
while len(y) < ds_size:
i1 = random.randrange(0, len(smiles_lens))
i2 = random.randrange(0, len(smiles_lens))
m1 = all_data[i1]
m2 = all_data[i2]
if m1[1] == m2[1]:
continue
if m1[1] > m2[1]:
y.append(1.0)
else:
y.append(0.0)
X.append([m1[0], m2[0]])
return dc.data.NumpyDataset(np.array(X), np.expand_dims(np.array(y), axis=1))
```
With our complexity ranker in place we can now construct our dataset. Let's start by loading the molecules in the Tox21 dataset into memory. We split the dataset at this stage to ensure that the training and test set have non-overlapping sets of molecules.
```
# Lets split our dataset into a train set and a test set
molecule_ds = dc.data.NumpyDataset(np.array(molecules))
splitter = dc.splits.RandomSplitter()
train_mols, test_mols = splitter.train_test_split(molecule_ds)
```
We'll featurize all our molecules with the ECFP fingerprint with chirality (matching the source paper), and will then construct our pairwise dataset using the code from above.
```
# In the paper they used 1024 bit fingerprints with chirality
n_features=1024
featurizer = dc.feat.CircularFingerprint(size=n_features, radius=2, chiral=True)
train_features = featurizer.featurize(train_mols.X)
train_smileslen = [len(Chem.MolToSmiles(x)) for x in train_mols.X]
train_dataset = create_dataset(train_features, train_smileslen)
```
Now that we have our dataset created, let's train a `ScScoreModel` on this dataset.
```
from deepchem.models import ScScoreModel
# Now to create the model and train it
model = ScScoreModel(n_features=n_features)
model.fit(train_dataset, nb_epoch=20)
```
# Model Performance
Lets evaluate how well the model does on our holdout molecules. The SaScores should track the length of SMILES strings from never before seen molecules.
```
import matplotlib.pyplot as plt
%matplotlib inline
mol_scores = model.predict_mols(test_mols.X)
smiles_lengths = [len(Chem.MolToSmiles(x)) for x in test_mols.X]
```
Let's now plot the length of the smiles string of the molecule against the SaScore using matplotlib.
```
plt.figure(figsize=(20,16))
plt.scatter(smiles_lengths, mol_scores)
plt.xlim(0,80)
plt.xlabel("SMILES length")
plt.ylabel("ScScore")
plt.show()
```
As we can see the model generally tracks SMILES length. It has good enrichment between 8 and 30 characters and gets both small and large SMILES strings extremes dead on.
Now you can train your own models on more meaningful metrics than SMILES length!
# Congratulations! Time to join the Community!
Congratulations on completing this tutorial notebook! If you enjoyed working through the tutorial, and want to continue working with DeepChem, we encourage you to finish the rest of the tutorials in this series. You can also help the DeepChem community in the following ways:
## Star DeepChem on [GitHub](https://github.com/deepchem/deepchem)
This helps build awareness of the DeepChem project and the tools for open source drug discovery that we're trying to build.
## Join the DeepChem Gitter
The DeepChem [Gitter](https://gitter.im/deepchem/Lobby) hosts a number of scientists, developers, and enthusiasts interested in deep learning for the life sciences. Join the conversation!
# Bibliography:
[1] https://pubs.acs.org/doi/abs/10.1021/acs.jcim.7b00622
| github_jupyter |
# Lecture 3 Exercise Solutions
## Exercise 1: Data Types and Operators
**Task 1.1:** Try how capitalization affects string comparison, e.g., compare "datascience" to "Datascience".
```
# Two strings are not considered equal if they have different capitalization.
"datascience" == "Datascience"
# I can make the strings lowercase to compare them, though.
# That works for ASCII, but there are Unicode cases where that breaks.
"dAtAscience".lower() == "Datascience".lower()
```
**Task 1.2:** Try to compare floats using the `==` operator defined as expressions of integers, e.g., whether 1/3 is equal to 2/6. Does that work?
```
# Comparing floats works sometimes, but you shouldn't rely on it ever!
1/5 == 2/10
a = 0.2 + 0.2 + 0.2
b = 0.6
a == b
a = 0.3
b = 0.1 + 0.1 + 0.1
a == b
a = 0.2
b = 0.1 + 0.1
a == b
a = 7/10
b = 14/20
a == b
a = 0.2 + 0.2 + 0.2 + 0.2
b = 0.8
a == b
```
**Task 1.3:** Write an expression that compares the "floor" value of a float to an integer, e.g., compare the floor of 1/3 to 0. There are two ways to calculate a floor value: using `int()` and using `math.floor()`. Are they equal? What is the data type of the returned values?
```
a = 0.1 + 0.1 + 0.1
type(a)
a
b
b = int(a)
0 == b
type(b)
import math
b = math.floor(a)
0 == b
type(b)
```
Both operations return an int and therefore can be safely compared.
## Exercise 3: Functions and If
Write a function that takes two integers. If either of the numbers can be divided by the other without a remainder, print the result of the division. If none of the numbers can divide the other one, print an error message.
```
def areFactors(x, y):
print("-------")
print("x:", x, "y:", y)
factorExists = False
if (not x % y):
print(x,"/",y,"=",x/y)
factorExists = True
if (not y % x):
print(y,"/",x,"=",y/x)
factorExists = True
if (not factorExists):
print("These numbers are not fators of each other")
areFactors(2, 4)
areFactors(4, 4)
areFactors(4, 2)
areFactors(6, 3)
areFactors(41263, 123789)
areFactors(7, 9)
```
## Exercise 4: Lists
```
# state of other notebook
beatles = ["Paul", "John", "George", "Ringo"]
zeppelin = ["Jimmy", "Robert", "John", "John"]
bands = [beatles, zeppelin]
stones = ["Mick", "Keith", "Charlie", "Ronnie"]
original_members = stones[0:3]
original_members
bands.append(original_members)
bands
```
## Exercise 5.1: While
```
sum = 0
count = 1
while (count <= 100):
sum += count
count += 1
sum
```
## Exercise 5.2: For
### 5.2.1: Array with even numbers
```
even_numbers = []
for i in range(51):
if (i == 0):
continue
elif (i%2 == 0):
even_numbers.append(i)
even_numbers
```
### 5.2.2: Foreach band member, add instrument
```
beatles = ["Paul", "John", "George", "Ringo"]
instruments = ["Bass", "Rythm Guitar", "Lead Guitar", "Drums"]
count = 0
for member in beatles:
print (member + ": " + instruments[count])
count +=1
```
## Exercise 6: Recursion
Write a recursive function that calculates the factorial of a number.
```
def factorial(n):
if(n>0):
return n * factorial(n-1)
else:
return 1
factorial(6)
```
## Exercise 7: List Comprehension
Write a list comprehension function that creates an array with the length of the words in the following sentence:
```
sentence = "the quick brown fox jumps over the lazy dog"
word_list = sentence.split()
word_list
[len(word) for word in word_list]
```
| github_jupyter |
```
import numpy as np
import numpy.fft as fft
import matplotlib.pyplot as plt
from IPython import display
from PIL import Image
import cv2
import os
import datetime
import matlab.engine
%matplotlib inline
def no_shift(plot_im=True):
delta = np.zeros((5,5))
delta[2][2] = 1
fft_mag = np.abs(fft.fft2(delta))
fft_arg = np.angle(fft.fft2(delta))
if plot_im:
fig, ax = plt.subplots(nrows=1, ncols=3)
fig.tight_layout()
ax[0].imshow(delta, cmap='gray')
ax[0].set_title('Delta function in \n real space')
ax[1].imshow(fft_mag,vmin=-3,vmax=3,cmap='gray')
ax[1].set_title('Magnitude of FT of \n a delta function')
ax[2].imshow(fft_arg,vmin=-3,vmax=3,cmap='gray')
ax[2].set_title('Phase of FT of \n delta function')
def shift(plot_im=True):
delta = np.zeros((5,5))
delta[2][2] = 1
delta_shifted = fft.ifftshift(delta)
fft_mag = np.abs(fft.fft2(delta_shifted))
fft_arg = np.angle(fft.fft2(delta_shifted))
if plot_im:
fig2, ax2 = plt.subplots(nrows=1, ncols=3)
fig2.tight_layout()
ax2[0].imshow(delta_shifted, cmap='gray')
ax2[0].set_title('Delta function shifted in \n real space')
ax2[1].imshow(fft_mag,vmin=-3,vmax=3,cmap='gray')
ax2[1].set_title('Magnitude of FT of a \n shifted delta function')
ax2[2].imshow(fft_arg,vmin=-3,vmax=3,cmap='gray')
ax2[2].set_title('Phase of FT of a \n shifted delta function')
def initMatrices(h):
pixel_start = (np.max(h) + np.min(h))/2
x = np.ones(h.shape)*pixel_start
init_shape = h.shape
print("This is the Init_shape: "+str(init_shape))
padded_shape = [nextPow2(2*n - 1) for n in init_shape]
print("This is the padded_shape: "+str(padded_shape))
starti = (padded_shape[0]- init_shape[0])//2
endi = starti + init_shape[0]
startj = (padded_shape[1]//2) - (init_shape[1]//2)
endj = startj + init_shape[1]
hpad = np.zeros(padded_shape)
hpad[starti:endi, startj:endj] = h
H = fft.fft2(fft.ifftshift(hpad), norm="ortho") #this calcualtes takes fourier transform of psf within padding
Hadj = np.conj(H)
def crop(X):
return X[starti:endi, startj:endj]
def pad(v):
vpad = np.zeros(padded_shape).astype(np.complex64)
vpad[starti:endi, startj:endj] = v
return vpad
utils = [crop, pad]
v = np.real(pad(x))
print("This is the v: "+str(v.shape))
pause()
return H, Hadj, v, utils
def nextPow2(n):
return int(2**np.ceil(np.log2(n)))
def grad(Hadj, H, vk, b, crop, pad):
Av = calcA(H, vk, crop)
diff = Av - b
norm2 = np.linalg.norm(diff)
print(norm2)
return np.real(calcAHerm(Hadj, diff, pad))
def calcA(H, vk, crop):
Vk = fft.fft2(fft.ifftshift(vk))
return crop(fft.fftshift(fft.ifft2(H*Vk)))
def calcAHerm(Hadj, diff, pad):
xpad = pad(diff)
X = fft.fft2(fft.ifftshift(xpad))
return fft.fftshift(fft.ifft2(Hadj*X))
def grad_descent(h, b, iters=200):
H, Hadj, v, utils = initMatrices(h)
crop = utils[0]
pad = utils[1]
alpha = np.real(1.8/(np.max(Hadj * H)))
iterations = 0
def non_neg(xi):
xi = np.maximum(xi,0)
return xi
#proj = lambda x:x #Do no projection
proj = non_neg #Enforce nonnegativity at every gradient step. Comment out as needed.
l2_error = np.zeros(iters)
parent_var = [H, Hadj, b, crop, pad, alpha, proj]
vk = v
mt = 0
#### uncomment for FISTA update ################
tk = 1
xk = v
#### uncomment for ADAM update ################
# vk = v
# #alpha = 0.01
# beta1 = 0.9
# beta2 = 0.999
# epsilon = 1e-06
# mt = 0
# t = 0 #time step
# vt = np.array([0])
# vth = np.array([0]) #np.zeros(padded_shape)
# amsgrad = False #set to true or false
for iterations in range(iters):
# uncomment for ADAM update
# vk, beta1, beta2, mt, vt, epsilon, t = adam_update(vk,beta1,beta2,mt,vt,vth,epsilon,parent_var,amsgrad,t)
vk, tk, xk = fista_update(vk, tk, xk, parent_var)
if iterations % 10 == 0:
image = proj(crop(vk))
f = plt.figure(1)
plt.imshow(image, cmap='gray')
plt.title('Reconstruction after iteration {}'.format(iterations))
display.display(f)
display.clear_output(wait=True)
return proj(crop(vk))
def adam_update(vk, beta1, beta2, mt, vt, vth, epsilon, parent_var, amsgrad, t):
H, Hadj, b, crop, pad, alpha, proj = parent_var
t = t+4
learn_rate = alpha * (np.sqrt(1-beta2**t)/(1-beta1**t))
gradient = grad(Hadj, H, vk, b, crop, pad)
mt1 = beta1*mt + (1-beta1)*gradient
vt1 = beta2*vt + (1-beta2)*gradient*gradient
#perform gradient update
if (amsgrad == True):
#print(type(vt))
#print(type(vth))
#pause()
vth1 = np.maximum(vth,vt1)
vk -= learn_rate*mt1/(np.sqrt(vth1)+epsilon)
vth = vth1
else:
vk -= learn_rate*mt1/(np.sqrt(vt1)+epsilon)
#vk -= learn_rate*mt1/(np.sqrt(vt1)+epsilon)
vk = proj(vk)
#t = t+1
vt = vt1
mt = mt1
return vk, beta1, beta2, mt, vt, epsilon, t
def fista_update(vk, tk, xk, parent_var):
H, Hadj, b, crop, pad, alpha, proj = parent_var
x_k1 = xk
gradient = grad(Hadj, H, vk, b, crop, pad)
vk -= alpha*gradient
xk = proj(vk)
t_k1 = (1+np.sqrt(1+4*tk**2))/2
vk = xk+(tk-1)/t_k1*(xk - x_k1)
tk = t_k1
#adaptive restart momentum check
'''grad_test = np.max(gradient*(xk-x_k1))
if (np.mean(xk) > np.mean(x_k1)): #grad_test > 0
print("Im in restart loop")
vk = xk
xk = vk
tk =1
else:
t_k1 = (1+np.sqrt(1+4*tk**2))/2
vk = xk+(tk-1)/t_k1*(xk - x_k1)
tk = t_k1'''
return vk, tk, xk
def loaddata(psf,data,dark=None,factor=8,show_im=True):
if type(data) == str:
with Image.open(psf) as psf_:
psf = np.array(psf_, dtype='float32')/255
with Image.open(data) as data_:
data = np.array(data_, dtype='float32')/255
if dark is not None:
bg = np.mean(dark)
else:
bg = np.mean(psf[5:15,5:15])
psf -= bg
data -= bg
"""Resize to a more manageable size to do reconstruction on.
Because resizing is downsampling, it is subject to aliasing
(artifacts produced by the periodic nature of sampling). Demosaicing is an attempt
to account for/reduce the aliasing caused. In this application, we do the simplest
possible demosaicing algorithm: smoothing/blurring the image with a box filter"""
psf_internal = internal_resize(psf.copy(),factor)
data_internal = internal_resize(data.copy(),factor)
del psf, data
psf = psf_internal
data = data_internal
"""Now we normalize the images so they have the same total power. Technically not a
necessary step, but the optimal hyperparameters are a function of the total power in
the PSF (among other things), so it makes sense to standardize it"""
psf /= np.linalg.norm(psf.ravel())
data /= np.linalg.norm(data.ravel())
if show_im:
fig1 = plt.figure()
plt.imshow(psf, cmap='gray')
plt.title('PSF')
display.display(fig1)
fig2 = plt.figure()
plt.imshow(data, cmap='gray')
plt.title('Raw data')
display.display(fig2)
return psf, data
def internal_resize(img, factor):
#factor = int(np.log2(factor))
width = int(img.shape[0]/factor)
height = int(img.shape[1]/factor)
out_dim = (height,width)
img_out = cv2.resize(img,out_dim, interpolation = cv2.INTER_LANCZOS4)
print(img_out.shape)
return img_out
class diffuseCAM(object):
def __init__(self,cam_idx=0,window_name='diffuseCAM',**kwargs):
self.window_name = window_name
self.sess = str(datetime.datetime.now()).replace(':','-')
self.cam_idx=cam_idx
super(diffuseCAM,self).__init__(**kwargs)
def preview(self):
self.start()
if self.cam.isOpened(): # try to get the first frame
rval, frame = self.cam.read()
else:
rval = False
while rval:
cv2.imshow(self.window_name, frame)
rval, frame = self.cam.read()
keypress = cv2.waitKey(20)
if keypress == 27: # exit on ESC
break
elif keypress == ord('q'):
frame = self.BGR2RGB(frame)
plt.figure()
plt.hist(frame.ravel())
# self.imsave(frame,'unsorted')
self.close()
def close(self):
cv2.destroyWindow(self.window_name)
self.cam.release()
def start(self):
cv2.namedWindow(self.window_name)
self.cam = cv2.VideoCapture(self.cam_idx)
if not self.cam.isOpened():
raise RuntimeError('Could not start camera')
def capture(self,folder='unsorted'):
self.start()
_, frame = self.cam.read()
self.close()
frame = self.BGR2RGB(frame)
self.imsave(frame,folder)
return frame
def BGR2RGB(self,frame):
frame = [frame[:,:,2],frame[:,:,1],frame[:,:,0]]
return np.asarray(np.swapaxes(np.swapaxes(frame,0,2),0,1),dtype=np.float32)/255
def calibrate_dark(self,num_frames=10):
self.start()
_, img = self.cam.read()
img = self.BGR2RGB(img)
for idx in range(num_frames-1):
_, frame = self.cam.read()
frame = self.BGR2RGB(frame)
img += frame
self.dark = img/num_frames
self.close()
def calibrate_psf(self,num_frames=5):
self.start()
_, img = self.cam.read()
img = self.BGR2RGB(img)
for idx in range(num_frames-1):
_, frame = self.cam.read()
frame = self.BGR2RGB(frame)
img += frame
self.psf = img/num_frames
self.imsave(frame,'psf')
self.close()
def imsave(self,img,folder):
time_now = self.sess = str(datetime.datetime.now()).replace(':','-')
plt.imsave('../data/%s/dc_%s.png'%(folder,time_now),img)
dC = diffuseCAM(1)
def pause():
programPause = input("Press the <ENTER> key to continue...")
eng = matlab.engine.start_matlab()
def rgb2gray(img,eng=None):
if eng is not None:
img = eng.rgb2gray(matlab.double(np.ndarray.tolist(img)))
else:
img = 0.2989 * img[:,:,0] + 0.5870 * img[:,:,1] + 0.1140 * img[:,:,2]
return img
dC.calibrate_dark()
plt.imshow(dC.dark)
dC.calibrate_psf()
plt.imshow(dC.psf)
img = dC.capture()
plt.imshow(img)
dC.preview()
psf = '../data/psf_sample.tif'
data = '../data/rawdata_hand_sample.tif'
iters = 200
psf, data = loaddata(psf,data,factor=8)
final_im = grad_descent(psf, data,iters)
plt.imshow(final_im, cmap='gray')
plt.title('Final reconstruction after {} iterations'.format(iters))
display.display()
#plt.savefig('final_img.pdf')
iters = 200
psf, data = loaddata(rgb2gray(dC.psf),
rgb2gray(dC.capture('raw_data')),
# rgb2gray(dC.dark),
factor=1)
final_im = grad_descent(psf,data,iters)
plt.imshow(final_im, cmap='gray')
dC.imsave(final_im,'reconstructed')
plt.title('Final reconstruction after {} iterations'.format(iters))
display.display()
#plt.savefig('final_img.pdf')
plt.imshow(rgb2gray(dC.psf))
```
| github_jupyter |
```
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
# You can write up to 20GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
df = pd.read_csv('/kaggle/input/titanic/train.csv')
```
### Visualization
```
df.head()
df.describe()
df.isnull().sum()
df.loc[df['Sex'] == 'male', 'Sex'] = 0
df.loc[df['Sex'] == 'female', 'Sex'] = 1
df.head()
#setdata = set(df['Name'].values.tolist())
#print(setdata)
df['Age'] = df['Age'].fillna(df['Age'].notnull().mean())
df.head()
df.isnull().sum()
```
scale = df[['Sex','Age','Survived']].to_numpy()
print(scale.shape)
```
titles = set()
for names in df['Name']:
titles.add(names.split(",")[1].split(".")[0].strip(" "))
print(titles)
Map = {'Mrs': 1, 'Miss': 2, 'Major': 3, 'Lady': 4, 'Sir': 5, 'Ms': 6, 'Mr': 7, 'Mlle': 8, 'Don': 9, 'Mme': 10, 'Capt': 11, 'Master': 12, 'the Countess': 13, 'Jonkheer': 14, 'Col': 15, 'Rev': 16, 'Dr': 17}
def func(row):
row['Name'] = row['Name'].split(",")[1].split(".")[0].strip(" ")
return Map.get(row['Name'])
df.loc[df['Embarked'] == 'C', 'Embarked'] = 0
df.loc[df['Embarked'] == 'Q', 'Embarked'] = 1
df.loc[df['Embarked'] == 'S', 'Embarked'] = 2
df['Embarked'] = df['Embarked'].fillna(df['Embarked'].notnull().mean())
df['Title'] = df.apply(func,axis=1)
df.head()
scale = df[['PassengerId','Pclass','Sex','Age','SibSp','Parch','Fare','Embarked','Title','Survived']].to_numpy()
print(scale.shape)
import matplotlib.pyplot as plt
df = df[['Age','Sex','Survived']]
X = df.iloc[:, :-1]
y = df.iloc[:, -1]
# filter out the applicants that got admitted
survival = df.loc[y == 1]
survival.head()
# filter out the applicants that din't get admission
not_survival = df.loc[y == 0]
# plots
plt.scatter(survival.iloc[:, 0], survival.iloc[:, 1], s=10, label='survival')
plt.scatter(not_survival.iloc[:, 0], not_survival.iloc[:, 1], s=10, label='not_survival')
plt.legend()
plt.show()
```
### Scale Data
```
t = int(0.8 * scale.shape[0])
train = scale[:t]
test = scale[t:]
print(train.shape, test.shape)
from sklearn import preprocessing
min_max_scaler = preprocessing.MinMaxScaler()
X_train = min_max_scaler.fit_transform(train[:,:-1])
X_test = min_max_scaler.transform(test[:,:-1])
print(X_train.shape, X_test.shape)
y_train = train[:,-1]
y_test = test[:,-1]
X_train = X_train
y_train = y_train.astype('int')
X_test = X_test
y_test = y_test.astype('int')
print(X_train)
```
### Training
```
%time
from sklearn.linear_model import LogisticRegression
model = LogisticRegression(random_state=0)
model.fit(X_train, y_train)
```
### Predict
```
model.predict(X_test)
```
### Accuracy
```
print("Train set: ", model.score(X_train, y_train))
print("Test set: ", model.score(X_test, y_test))
```
### Accuracy with Age, Sex:

### Accuracy with PassengerId, Pclass, Sex, Age, Fare:

### Submit
```
csv = pd.read_csv('/kaggle/input/titanic/test.csv')
csv.head()
csv.isnull().sum()
csv.loc[csv['Sex'] == 'male', 'Sex'] = 0
csv.loc[csv['Sex'] == 'female', 'Sex'] = 1
csv['Age'] = csv['Age'].fillna(csv['Age'].notnull().mean())
csv['Fare'] = csv['Fare'].fillna(csv['Fare'].notnull().mean())
Map = {'Mrs': 1, 'Miss': 2, 'Major': 3, 'Lady': 4, 'Sir': 5, 'Ms': 6, 'Mr': 7, 'Mlle': 8, 'Don': 9, 'Mme': 10, 'Capt': 11, 'Master': 12, 'the Countess': 13, 'Jonkheer': 14, 'Col': 15, 'Rev': 16, 'Dr': 17}
def func(row):
row['Name'] = row['Name'].split(",")[1].split(".")[0].strip(" ")
return Map.get(row['Name'])
csv.loc[csv['Embarked'] == 'C', 'Embarked'] = 0
csv.loc[csv['Embarked'] == 'Q', 'Embarked'] = 1
csv.loc[csv['Embarked'] == 'S', 'Embarked'] = 2
csv['Embarked'] = csv['Embarked'].fillna(csv['Embarked'].notnull().mean())
csv['Title'] = csv.apply(func,axis=1)
csv['Title'] = csv['Title'].fillna(csv['Title'].notnull().mean())
df.head()
X_test = min_max_scaler.transform(csv[['PassengerId','Pclass','Sex','Age','SibSp','Parch','Fare','Embarked','Title']].to_numpy())
csv.isnull().sum()
print(np.any(np.isnan(X_test)), np.all(np.isfinite(X_test)))
np.argwhere(np.isnan(X_test))
y_pred = model.predict(X_test)
print(y_pred)
output = pd.DataFrame({'PassengerId': csv.PassengerId, 'Survived': y_pred.astype(int)})
output.to_csv('submission.csv', index=False)
```
| github_jupyter |
```
%%HTML
<style>
.CodeMirror {
width: 100vw;
}
.container {
width: 79% !important;
}
.rendered_html {
font-size:0.8em;
}
.rendered_html table, .rendered_html th, .rendered_html tr, .rendered_html td {
font-size: 100%;
}
</style>
```
# Performance
http://docs.micropython.org/en/latest/pyboard/reference/constrained.html
[Writing fast and efficient MicroPython (Youtube)](https://www.youtube.com/watch?v=hHec4qL00x0&t=1261s) - Damien George






# Example: blinking leds
``` python
led = machine.Pin('LED_BLUE')
N = 200000
for i in range(N):
led.on()
led.off()
```
57.93 kblinks/sec
## Reasonable optimization (182.39 kblinks/sec)

## Hardcore optimizations
### @micropython.viper
- directly writing GPIO registers
- 12890 kblinks/sec
### @micropython.asm_thumb
- directly writing GPIO registers - in assembler..
- 27359 kblinks/sec
- 500 times faster than initial code

Compiler can emit machine code
> ahead-of-time (AOT) compilation is the act of compiling a higher-level programming language such as C or C++, or an intermediate representation such as Java bytecode or .NET Framework Common Intermediate Language (CIL) code, into a native (system-dependent) machine code so that the resulting binary file can execute natively.
## Flash
> For reasons connected with the physical architecture of the flash memory part of this capacity may be inaccessible as a filesystem. In such cases this space may be employed by incorporating user modules into a firmware build which is then flashed to the device.
### custom build

**scripts/**: raw Python code stored in the board's flash memory.
**modules/**: using cross-compiler to store python modules as bytecode
### frozen modules (.mpy)
Frozen modules store the Python source with the firmware.
### frozen bytecode
Frozen bytecode uses the cross compiler to convert the source to bytecode which is then stored with the firmware.
**Scripts/**: This saves you from having to copy that code onto the board's filesystem, but doesn't save a lot of memory or processing time.
### Steps:
1. Clone the MicroPython repository.
- Acquire the (platform specific) toolchain to build the firmware.
- Build the cross compiler.
- Place the modules to be frozen in a specified directory (dependent on whether the module is to be frozen as source or as bytecode).
- Build the firmware. A specific command may be required to build frozen code of either type - see the platform documentation.
- Flash the firmware to the device.
# RAM
- compilation phase
- execution phase
## Compilation Phase
#### compiling during runtime requires overhead RAM
- import module -> compiled to bytecode -> RAM
- instantiated objects -> RAM
- The compiler itself requires RAM
* **Limit module imports**
* __Avoid global objects in imported modules__
> In general it is best to avoid code which runs on import; a better approach is to have initialisation code which is run by the application after all modules have been imported.
When a module is imported, MicroPython compiles the code to bytecode which is then executed by the MicroPython virtual machine (VM). The bytecode is stored in RAM. The compiler itself requires RAM, but this becomes available for use when the compilation has completed.
### Frozen modules
### Frozen bytecode
on most platforms this saves even more RAM as the bytecode is run directly from flash rather than being stored in RAM.
## Exection phase
## Constants
```
from micropython import const
ROWS = const(33)
_COLS = const(0x10)
a = ROWS
b = _COLS
```
- compiler substitutes identifier with numeric value in bytecode
- this avoids a dictionary lookup at runtime.
- anything that evaluates to integer compile time
In both instances where the constant is assigned to a variable the compiler will avoid coding a lookup to the name of the constant by substituting its literal value. This saves bytecode and hence RAM. However the ROWS value will occupy at least two machine words, one each for the key and value in the globals dictionary. The presence in the dictionary is necessary because another module might import or use it. This RAM can be saved by prepending the name with an underscore as in _COLS: this symbol is not visible outside the module so will not occupy RAM.
The argument to const() may be anything which, at compile time, evaluates to an integer e.g. 0x100 or 1 << 8. It can even include other const symbols that have already been defined, e.g. 1 << BIT.
## Constant data structures
- Store data as bytes in frozen bytecode.
- Use ustruct to convert between bytes and python built-in types
- strings
- numeric data
The compiler ‘knows’ that bytes objects are immutable and ensures that the objects remain in flash memory rather than being copied to RAM.
Where there is a substantial volume of constant data and the platform supports execution from Flash, RAM may be saved as follows. The data should be located in Python modules and frozen as bytecode. The data must be defined as bytes objects. The compiler ‘knows’ that bytes objects are immutable and ensures that the objects remain in flash memory rather than being copied to RAM. The ustruct module can assist in converting between bytes types and other Python built-in types.
When considering the implications of frozen bytecode, note that in Python strings, floats, bytes, integers and complex numbers are immutable. Accordingly these will be frozen into flash. Thus, in the line
`mystring = "The quick brown fox"`
the actual string “The quick brown fox” will reside in flash. At runtime a reference to the string is assigned to the variable mystring. The reference occupies a single machine word. In principle a long integer could be used to store constant data:
`bar = 0xDEADBEEF0000DEADBEEF`
As in the string example, at runtime a reference to the arbitrarily large integer is assigned to the variable bar. That reference occupies a single machine word.
### String concatenation
```
var1 = "foo" "bar"
var2 = """\
foo\
bar"""
```
Creates at compile time
`var = "foo" + "bar"`
Creates "foo", "bar" and var at runtime
> The best way to create dynamic strings is by means of the string format() method:
`var = "Temperature {:5.2f} Pressure {:06d}\n".format(temp, press)`
## Buffers
When accessing devices such as instances of UART, I2C and SPI interfaces, using pre-allocated buffers avoids the creation of needless objects. Consider these two loops:
```
while True:
var = spi.read(100)
# process data
buf = bytearray(100)
while True:
spi.readinto(buf)
# process data in buf
```
The first creates a buffer on each pass whereas the second re-uses a pre-allocated buffer; this is both faster and more efficient in terms of memory fragmentation.
## Heap
## Heap fragmentation
<img width="500px" src="https://upload.wikimedia.org/wikipedia/commons/4/4a/External_Fragmentation.svg" align="right" alt="Alt text that describes the graphic" title="https://commons.wikimedia.org/wiki/File:External_Fragmentation.svg">
- <font color="red">Minimise</font> the repeated creation and destruction of objects!
- instantiate large buffers/objects early
- periodic use of `gc.collect()` (few ms)
> The discourse on this is somewhat involved. For a ‘quick fix’ issue the following periodically:
```
gc.collect()
gc.threshold( (gc.mem_free() // 10) + gc.mem_alloc())
```
This will provoke a GC when more than 10% of the currently free heap becomes occupied.
## Commands
```
import gc
import micropython
gc.collect()
micropython.mem_info()
print('-----------------------------')
print('Initial free: {} allocated: {}'.format(gc.mem_free(), gc.mem_alloc()))
def func():
a = bytearray(10000)
gc.collect()
print('Func definition: {} allocated: {}'.format(gc.mem_free(), gc.mem_alloc()))
func()
print('Func run free: {} allocated: {}'.format(gc.mem_free(), gc.mem_alloc()))
gc.collect()
print('Garbage collect free: {} allocated: {}'.format(gc.mem_free(), gc.mem_alloc()))
print('-----------------------------')
micropython.mem_info(1)
```
- gc.collect() Force a garbage collection.
- micropython.mem_info() Print a summary of RAM utilisation.
- gc.mem_free() Return the free heap size in bytes.
- gc.mem_alloc() Return the number of bytes currently allocated.
- micropython.mem_info(1) Print a table of heap utilisation (detailed below).
Error messages can be strange
```
with open("file.txt") as f:
f.write("hello world")
```
## Performance
- Locally scope your variables
- Avoid floating Point arithmetric
- Use allocate buffers (instead of appending list)
Some MicroPython ports allocate floating point numbers on heap. Some other ports may lack dedicated floating-point coprocessor, and perform arithmetic operations on them in “software” at considerably lower speed than on integers.
- Caching object references
```
class Tank(object):
def __init__(self):
self.targets = bytearray(100)
def identify_target(self, obj_display):
targets_ref = self.targets
fb = obj_display.framebuffer
(...)
```
| github_jupyter |
```
%matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
```
## Fetch test data
```
from geneview.utils import load_dataset
admixture_output_fn = load_dataset("admixture_output.Q")
population_group_fn = load_dataset("admixture_population.info")
```
## The output data of admixture
```
admixture_q = pd.read_table(admixture_output_fn, sep=" ", header=None)
admixture_q
```
## Sample information for admixture looks like:
```
# each row for one sample.
sample_info = pd.read_table(population_group_fn, sep="\t", header=None, names=["Group"])
sample_info
```
## A simple examples
```
from geneview.utils import load_dataset
from geneview import admixtureplot
f, ax = plt.subplots(1, 1, figsize=(14, 2), facecolor="w", constrained_layout=True, dpi=300)
admixtureplot(data=admixture_output_fn, population_info=population_group_fn, ax=ax)
admixtureplot(data=admixture_output_fn,
population_info=population_group_fn,
linewidth=2.0,
shuffle_popsample_kws={"frac": 0.2})
# define the order for population to plot
pop_group_1kg = ["KHV", "CDX", "CHS", "CHB", "JPT", "BEB", "STU", "ITU", "GIH", "PJL", "FIN",
"CEU", "GBR", "IBS", "TSI", "PEL", "PUR", "MXL", "CLM", "ASW", "ACB", "GWD",
"MSL", "YRI", "ESN", "LWK"]
f, ax = plt.subplots(1, 1, figsize=(14, 2), facecolor="w", constrained_layout=True, dpi=300)
admixtureplot(data=admixture_output_fn,
population_info=population_group_fn,
group_order=pop_group_1kg,
shuffle_popsample_kws={"n": 50},
ylabel_kws={"rotation": 0, "ha": "right"},
ax=ax)
df = pd.read_table(admixture_output_fn, sep=" ", header=None)
sample_info = pd.read_table(population_group_fn, sep="\t", header=None, names=["Group"])
popset = set(sample_info["Group"])
data = {}
for g in popset:
g_data = df[sample_info["Group"]==g].copy()
# Sub sampling: keep less than 140 samples for each group
data[g] = g_data.sample(n=140, random_state=100) if len(g_data)>140 else g_data
### Plot
params = {"legend.fontsize": 14,
"axes.labelsize": 16,
"xtick.labelsize": 14,
"ytick.labelsize": 14}
plt.rcParams.update(params)
# define a plot
f, ax = plt.subplots(1, 1, figsize=(14, 2), facecolor="w", constrained_layout=True, dpi=300)
_ = admixtureplot(data=data,
group_order=pop_group_1kg,
palette="Set1",
xticklabel_kws={"rotation": 45},
ylabel="K=11",
ylabel_kws={"rotation": 0, "ha": "right"},
ax=ax)
```
| github_jupyter |
## Convolutional Neural Network (CNN)
- CNN is basically two dimensional configuration of neural networks
- The input of CNN are image (3 N by N if it color image and N by N if its black and white image)
- The weights are also two dimensional array
<img src="rgb_image.png" width="300" height="300">
### The weights in CNN
- The weights in CNN are called:
- Kernel
or
- Filter matrix
<img src="kernel_image.png" width="400" height="400">
### Stride
- To define a CNN, we should specify the horizontal and vertical movement steps
- what is the output size with stride = 1 and stride =2?
<img src="stride_1.png" width="400" height="400">
<img src="stride_2.png" width="400" height="400">
- output_size = (input_size - filter_size)/stride + 1
- Stride visualization: http://deeplearning.stanford.edu/wiki/index.php/Feature_extraction_using_convolution
```
model.add(Conv2D(32, kernel_size=(3, 3), strides=(1, 1), activation='relu', input_shape=input_shape))
```
### Max Pooling vs Average Pooling
- Max pooling: take the maximum element from each window of a certain size
<img src="maxpooling.png" width="400" height="400">
### Faltten Layer
- After feature extraction that is done by multiple Convolutional layers, we use flatten layer to add MLP after convolutional layers in order to do classification task
- This one is simple--it's just Keras's verison of numpy.reshape. This reshapes n-dimensional arrays to a vector. This is necessary when moving from Conv2D layers, which expect 2-dimensional arrays as inputs, to Dense layers, which expect 1-dimension vectors as inputs. As a concrete example, a Flatten layer given a 28 x 28 array as input would output a vector of the shape (784, 1)
### Visualize the whole NN: CNN + MLP
<img src="CNN.png" width="600" height="600">
## Activity: Obtain the number of parameters for the following CNN
- By default, the strides = (1, 1)
```
from __future__ import print_function
import keras
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
from keras.layers import Conv2D, MaxPooling2D
model = Sequential()
model.add(Conv2D(32, kernel_size=(3, 3),
activation='relu',
input_shape=(28, 28, 1)))
model.add(Conv2D(64, (3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
# model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
# model.add(Dropout(0.5))
model.add(Dense(10, activation='softmax'))
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
print(model.summary())
```
### Solution
- output_size = (28 - 3)/1 + 1 = 26
- output_size = (26 - 3)/1 + 1 = 24
- The parameters for the first Conv2D = 32 x 9 + 32 = 320
- The parameters for the second Conv2D = 64 x 32 x 9 + 64 = 18496
- The shape for flatten is: 12 x 12 x 64 = 9216
- The parameters for dense_1 = 9216 x 128 + 128 = 1179776
- The parameters for dense_2 = 128 x 10 + 10 = 1290
## Data Preparation for CNN
- Suppose we want to feed a 4 by 4 image to a CNN network, how we should reshape the data?
```
from keras.layers import Input, Dense, Conv2D, MaxPooling2D, UpSampling2D
from keras.models import Model
from keras import backend as K
import numpy as np
input_img = Input(shape=(4, 4, 1)) # adapt this if using `channels_first` image data format
x = Conv2D(2, (2, 2), activation='relu')(input_img)
y = Conv2D(3, (2, 2), activation='relu')(x)
model = Model(input_img, y)
# cnv_ml_1 = Model(input_img, x)
data = np.array([[5, 12, 1, 8], [2, 10, 3, 6], [4, 7, 9, 1], [5, 7, 5, 6]])
data = data.reshape(4, 4, 1)
print(data[0])
data = np.expand_dims(data, axis=0)
# data = data.reshape(4, 4, 1)
print(data[0])
print(model.predict(data))
print('M :')
print(model.predict(data).reshape(3, 2, 2))
print(model.summary())
from keras.layers import Input, Dense, Conv2D, MaxPooling2D, UpSampling2D
from keras.models import Model
from keras import backend as K
import numpy as np
input_img = Input(shape=(4, 4, 1)) # adapt this if using `channels_first` image data format
x = Conv2D(2, (2, 2), activation='relu')(input_img)
y = Conv2D(3, (2, 2), activation='relu')(x)
model = Model(input_img, y)
# cnv_ml_1 = Model(input_img, x)
data = np.array([[5, 12, 1, 8], [2, 10, 3, 6], [4, 7, 9, 1], [5, 7, 5, 6]])
data = data.reshape(1, 4, 4, 1)
print(model.predict(data))
print('M :')
print(model.predict(data).reshape(3, 2, 2))
print(model.summary())
from keras.datasets import mnist
img_rows, img_cols = 28, 28
# the data, split between train and test sets
(x_train, y_train), (x_test, y_test) = mnist.load_data()
if K.image_data_format() == 'channels_first':
x_train = x_train.reshape(x_train.shape[0], 1, img_rows, img_cols)
x_test = x_test.reshape(x_test.shape[0], 1, img_rows, img_cols)
input_shape = (1, img_rows, img_cols)
else:
x_train = x_train.reshape(x_train.shape[0], img_rows, img_cols, 1)
x_test = x_test.reshape(x_test.shape[0], img_rows, img_cols, 1)
input_shape = (img_rows, img_cols, 1)
print(x_train[0].shape)
print(x_train[1].shape)
```
| github_jupyter |
# Finding a Tangent Disk
We are going to show how to use inversive geometry to find a disk that is internally tangent to the unit disk and externally tangent to two given example disks.
```
# Run this first: import necessary libraries
# Geometrical objects
from koebe.geometries.orientedProjective2 import PointOP2, DiskOP2
from koebe.geometries.euclidean2 import CircleE2, PointE2
from koebe.geometries.extendedComplex import ExtendedComplex, Mobius
# Viewers used
from koebe.graphics.euclidean2viewer import UnitScaleE2Sketch, makeStyle
# Other imports
import math
# Next, create the unit disk and two example disks that are tangent to one another and
# internally tangent to the unit disk.
unitDisk = DiskOP2(1/math.sqrt(2), 0, 0, -1/math.sqrt(2))
exampleDisk1 = DiskOP2.fromCenterAndRadius(PointOP2(-0.5, 0), 0.5)
exampleDisk2 = DiskOP2.fromCenterAndRadius(PointOP2( 0.5, 0), 0.5)
# Now, get the three points of tangency of the three disks.
p1 = unitDisk.intersectWithDiskOP2(exampleDisk1)[0]
p2 = exampleDisk1.intersectWithDiskOP2(exampleDisk2)[0]
p3 = exampleDisk2.intersectWithDiskOP2(unitDisk)[0]
# Create a viewer and draw the two disks
viewer = UnitScaleE2Sketch()
viewer.addAll(
[unitDisk, exampleDisk1, exampleDisk2, p1, p2, p3]
)
viewer.show()
```
Notice the three blue points that are the points of tangency between our circles given in left to right order as $p_1$, $p_2$, and $p_3$.
```
# Convert the points to ExtendedComplex numbers (the ExtendedComplex numbers use homogeneous coordinates
# to represent both the complex numbers and a projective line at infinity).
z1, z2, z3 = [p.toExtendedComplex() for p in (p1, p2, p3)]
# Compute the Möbius transformation taking z1 -> 0, z2 -> 1, and z3 -> infinity.
# Conceptually, this makes the unit disk the vertical line through the origin,
# exampleDisk1 a disk with radius 0.5 centered at (0.5, 0) and exampleDisk2 the
# vertical line through (1, 0).
M = Mobius.transformToZeroOneInfinity(z1, z2, z3)
# Under the transformation above, we are now just looking for a circle that is tangent vertical lines
# through (0, 0) and (1, 0) and the circle centered at (0.5, 0) with radius 0.5 This is now very easy
# and the three points of tangency of that circle are w1, w2, w3 given below.
w1 = ExtendedComplex(complex(0.0, 1.0))
w2 = ExtendedComplex(complex(0.5, 0.5))
w3 = ExtendedComplex(complex(1.0, 1.0))
# Apply the inverse Möbius transformation to the three points of tangency to get them in our
# original picture.
x1, x2, x3 = M.inverse.apply(w1, w2, w3)
# Finally, convert them to Point objects.
q1, q2, q3 = [x.toPointOP2() for x in (x1, x2, x3)]
# We can now view the three points q1, q2, q3 and the tangent disk (styled in red):
viewer = UnitScaleE2Sketch()
viewer.addAll([
unitDisk,
exampleDisk1,
exampleDisk2,
p1,
p2,
p3,
(q1, makeStyle(fill="#f00")),
(q2, makeStyle(fill="#f00")),
(q3, makeStyle(fill="#f00")),
(DiskOP2.fromPointOP2(q1, q2, q3), makeStyle(stroke="#f00"))
])
viewer.show()
```
Now let's make this a function! Note there are actually two solutions, so we may as well return both.
```
def mutuallyTangent(p1: PointOP2, p2: PointOP2, p3: PointOP2) -> (PointOP2, PointOP2, PointOP2):
"""Finds a circle C mutually tangent to three mutually circles C1, C2, C3 given their three points of tangency.
Args:
p1: The point of tangency between C1 and C2
p2: The point of tangency between C2 and C3
p3: The point of tangency between C3 and C1
Returns:
D1, D2, D3 the points of tangency between C and C1, C2, and C3.
"""
# Convert the points to ExtendedComplex numbers (the ExtendedComplex numbers use homogeneous coordinates
# to represent both the complex numbers and a projective line at infinity).
z1, z2, z3 = [p.toExtendedComplex() for p in (p1, p2, p3)]
# Compute the Möbius transformation taking z1 -> 0, z2 -> 1, and z3 -> infinity.
# Conceptually, this makes the unit disk the vertical line through the origin,
# exampleDisk1 a disk with radius 0.5 centered at (0.5, 0) and exampleDisk2 the
# vertical line through (1, 0).
M = Mobius.transformToZeroOneInfinity(z1, z2, z3)
# Solution 1:
# Under the transformation above, we are now just looking for a circle that is tangent vertical lines
# through (0, 0) and (1, 0) and the circle centered at (0.5, 0) with radius 0.5 This is now very easy
# and the three points of tangency of that circle are w1, w2, w3 given below.
w1a = ExtendedComplex(complex(0.0, 1.0))
w2a = ExtendedComplex(complex(0.5, 0.5))
w3a = ExtendedComplex(complex(1.0, 1.0))
# Apply the inverse Möbius transformation to the three points of tangency to get them in our
# original picture.
x1a, x2a, x3a = M.inverse.apply(w1a, w2a, w3a)
# Solution 2:
# Under the transformation above, we are now just looking for a circle that is tangent vertical lines
# through (0, 0) and (1, 0) and the circle centered at (0.5, 0) with radius 0.5 This is now very easy
# and the three points of tangency of that circle are w1, w2, w3 given below.
w1b = ExtendedComplex(complex(0.0, -1.0))
w2b = ExtendedComplex(complex(0.5, -0.5))
w3b = ExtendedComplex(complex(1.0, -1.0))
# Apply the inverse Möbius transformation to the three points of tangency to get them in our
# original picture.
x1b, x2b, x3b = M.inverse.apply(w1b, w2b, w3b)
# Finally, convert them to Point objects.
q1a, q2a, q3a, q1b, q2b, q3b = [x.toPointOP2() for x in (x1a, x2a, x3a, x1b, x2b, x3b)]
return (q1a, q2a, q3a), (q1b, q2b, q3b)
sols0 = mutuallyTangent(q2, p2, q3)[0]
sols1 = mutuallyTangent(q3, p3, q1)[0]
sols2 = mutuallyTangent(q1, p1, q2)[0]
# We can now view the three points q1, q2, q3 and the tangent disk (styled in red):
viewer = UnitScaleE2Sketch(width=400,height=400)
viewer.addAll([
unitDisk,
exampleDisk1,
exampleDisk2,
p1,
p2,
p3,
(q1, makeStyle(fill="#f00")),
(q2, makeStyle(fill="#f00")),
(q3, makeStyle(fill="#f00")),
(DiskOP2.fromPointOP2(q1, q2, q3), makeStyle(stroke="#f00")),
(sols0[0], makeStyle(fill="#0f0")),
(sols0[1], makeStyle(fill="#0f0")),
(sols0[2], makeStyle(fill="#0f0")),
(DiskOP2.fromPointOP2(*sols0), makeStyle(stroke="#0f0")),
(sols1[0], makeStyle(fill="#0f0")),
(sols1[1], makeStyle(fill="#0f0")),
(sols1[2], makeStyle(fill="#0f0")),
(DiskOP2.fromPointOP2(*sols1), makeStyle(stroke="#0f0")),
(sols2[0], makeStyle(fill="#0f0")),
(sols2[1], makeStyle(fill="#0f0")),
(sols2[2], makeStyle(fill="#0f0")),
(DiskOP2.fromPointOP2(*sols2), makeStyle(stroke="#0f0"))
])
viewer.show()
```
| github_jupyter |
```
import tkinter as tk
import numpy as np
from tkinter import Scale,ttk,Button,Text
from PIL import Image, ImageTk
import os
imageList=[]
folder=os.getcwd()+"\\images"
for filename in os.listdir(folder):
img=np.asarray(Image.open(os.path.join(folder,filename)))
if len(img.shape)<3: # Gif or Greyscale
continue
if img.shape[2]>3:
img=img[:,:,:3]
if img is not None:
imageList.append(img)
print(len(imageList))
if not os.path.exists(os.getcwd()+"\\heads"):
os.mkdir(os.getcwd()+"\\heads")
def square_update(length):
global currentIndex,canvas,source,photoImg,origin
length=int(length)
if origin[0]+length>=len(source):
origin[0]=len(source)-1-length
if origin[1]+length>=len(source[0]):
origin[1]=len(source[1])-1-length
toDraw=source.copy()
red=[255,0,0]
for i in range(origin[0],origin[0]+length):
if i>=len(source):
i=len(source)-1
toDraw[i][origin[1]]=red
x=origin[1]+length
if x>=len(source[0]):
x=len(source[0])-1
toDraw[i][x]=red
for j in range(origin[1],origin[1]+length):
if j>=len(source[0]):
j=len(source[0])-1
toDraw[origin[0]][j]=red
x=origin[0]+length
if x>=len(source):
x=len(source)-1
toDraw[x][j]=red
image=Image.fromarray(toDraw)
image=image.resize((canvas_width,canvas_height),resample=0)
photoImg=ImageTk.PhotoImage(image,master=canvas)
canvas.create_image(0,0,image=photoImg,anchor="nw")
def cropCurrent():
global source,origin,file_count,lengthSlider,text
length=lengthSlider.get()
sub_image=source[origin[0]:origin[0]+length,origin[1]:origin[1]+length]
image=Image.fromarray(sub_image)
image.save("heads\\"+str(file_count)+"_head.jpg")
print("Cropped image saved at","heads\\"+str(file_count)+"_head.jpg")
file_count+=1
text.configure(state='normal')
text.delete('1.0',tk.END)
text.insert(tk.END,"Image Number: "+str(currentIndex)+"\nFiles Saved: "+str(file_count))
text.configure(state='disabled')
def Up(event):
global pixel_per_move
pixel_per_move=int(pixel_per_move)
if origin[0]-pixel_per_move>=0:
origin[0]=origin[0]-pixel_per_move
else:
origin[0]=0
square_update(lengthSlider.get())
def Down(event):
global pixel_per_move
pixel_per_move=int(pixel_per_move)
if origin[0]+pixel_per_move<len(source)-1:
origin[0]=origin[0]+pixel_per_move
else:
origin[0]=len(source)-1
square_update(lengthSlider.get())
def Left(event):
global pixel_per_move
pixel_per_move=int(pixel_per_move)
if origin[1]-pixel_per_move>0:
origin[1]=origin[1]-pixel_per_move
else:
origin[1]=0
square_update(lengthSlider.get())
def Right(event):
global pixel_per_move
pixel_per_move=int(pixel_per_move)
if origin[1]+pixel_per_move<len(source[0])-1:
origin[1]=origin[1]+pixel_per_move
else:
origin[1]=len(source[0])-1
square_update(lengthSlider.get())
def loadImg():
global currentIndex,canvas,lengthSlider,source,photoImg,canvas_height,origin,text,file_count
if currentIndex==len(imageList):
print("Last image, program ends here.")
root.destroy()
return
canvas.delete('all')
source=imageList[currentIndex]
currentIndex+=1
text.configure(state='normal')
text.delete('1.0',tk.END)
text.insert(tk.END,"Image Number: "+str(currentIndex)+"\nFiles Saved: "+str(file_count))
text.configure(state='disabled')
image=Image.fromarray(source)
size=image.size
max_length=min(size)-1
length=max_length
canvas_height=int(size[1]*(canvas_width/size[0]))
origin=[0,0]
image=image.resize((canvas_width,canvas_height),resample=0)
canvas.config(height=canvas_height)
photoImg=ImageTk.PhotoImage(image,master=canvas)
canvas.create_image(0,0,image=photoImg,anchor="nw")
lengthSlider.config(to=max_length)
lengthSlider.set(max_length/2)
def loadPrevious():
global currentIndex,text
if currentIndex>1:
currentIndex-=2
loadImg()
def movement_update(value):
global pixel_per_move
pixel_per_move=value
file_count=0
origin=[0,0]
length=None
canvas_width=800
pixel_per_move=1
canvas_height=None
root=tk.Tk()
canvas=tk.Canvas(root,width=canvas_width,height=canvas_width,borderwidth=0,highlightthickness=0)
canvas.pack()
crop=Button(root,text="Crop",command=cropCurrent)
crop.pack(side="bottom")
moveOn=Button(root,text="Next",command=loadImg)
moveOn.pack(side="bottom")
moveBack=Button(root,text="Previous",command=loadPrevious)
moveBack.pack(side="bottom")
movementSlider=None
movementSlider=Scale(root,variable=pixel_per_move,orient="vertical",from_=1,to=5, command=lambda value=movementSlider:movement_update(value))
movementSlider.set(1)
movementSlider.pack(side="right")
def scroll_movement(event):
lengthSlider.set(lengthSlider.get()+2*(event.delta/120))
square_update(lengthSlider.get())
root.bind("<MouseWheel>", scroll_movement)
lengthSlider=None
lengthSlider=Scale(root,variable=length,orient="horizontal",from_=1,command=lambda value=lengthSlider:square_update(value),length=canvas_width*0.6)
lengthSlider.set(int(lengthSlider.cget("to")/2))
lengthSlider.pack(side="bottom")
text=Text(root,height=2,state="disabled")
text.configure(state='normal')
text.insert(tk.END,"Image Number: 0\nFiles Saved:0")
text.configure(state='disabled')
text.pack()
def arrowPrevious(event):
loadPrevious()
def arrowLoad(event):
loadImg()
root.bind("<Left>", arrowPrevious)
root.bind("<Right>", arrowLoad)
root.bind("w", Up)
root.bind("s", Down)
root.bind("a", Left)
root.bind("d", Right)
def clickCrop(event):
cropCurrent()
canvas.bind("<Double-Button-1>",clickCrop)
currentIndex=0
loadImg()
square_update(lengthSlider.get())
root.after(1, lambda: root.focus_force())
root.mainloop()
```
| github_jupyter |
```
import itertools as it
import sys
import os
import math
#if len(sys.argv) != 3:
# print("Usage: python3 " + sys.argv[0] + " N mode" )
# print("N is the max N-body contribution you want")
# sys.exit(1)
#nbmaxg = int(sys.argv[1])
#mode = int(sys.argv[2])
mode = 1
atlistg = [3,3,3,3,3]
opt_mon_eng = [-185.06839054,-185.06839054,-185.06839054,-185.06839054,-185.06839054]
#atlist = [3,3]
#opt_mon_en = [-186.561257471,-186.561257471]
nbmaxg = 5
nb = len(atlistg)
def get_mb(nbmax,atlist,opt_mon_en):
# Getting data we will need
nb = len(atlist)
nm = []
monsN = range(1,nb + 1)
combs = []
for i in range(1,nb+1):
combs.append(list(it.combinations(monsN,i)))
nm.append(len(combs[i-1]))
# Store energies
energies = []
for i in range(nbmax):
foldname = str(i + 1) + "b"
nbEn = []
for j in range(nm[i]):
fold = foldname + "/" + str(j+1)
ff = open(fold + "/input.energy", 'r')
nbEn.append(float(ff.readline().split()[0]))
energies.append(nbEn)
# Get 1b
enb = []
e = []
for i in range(nm[0]):
e.append(energies[0][i] - opt_mon_en[i])
enb.append(e)
# Get the many-body contributions (>1B)
# Loop overall the nb contributions
for nbx in range(1,nbmax):
n = nbx + 1
# Get the nbx-yh body contribution for each fragment
e = []
for frag in combs[nbx]:
sumE = []
e.append(0.0)
x = len(e) - 1
# Loop over all the nth subfragments of the cluster
for k in range(1,len(frag) + 1):
sumE.append(0.0)
icomb = list(it.combinations(frag,k))
for l2 in range(len(icomb)):
for l in range(1,len(combs[k - 1]) + 1):
filename = str(k) + "b/" + str(l) + "/input.xyz"
ff = open(filename)
ff.readline()
refcomb = ff.readline().split()
ff.close()
refcomb = [int(i) for i in refcomb]
if sorted(refcomb) == sorted(list(icomb[l2])):
sumE[k-1] += energies[k-1][l-1]
break
Nmax = len(frag)
for m2 in range(n):
m = m2 + 1
one = pow(-1,Nmax - m )
a = math.factorial(Nmax - m )
b = math.factorial(k - m)
c = math.factorial(Nmax - k)
g = one * a / b / c
e[x ] += g * sumE[m2]
enb.append(e)
return enb
if mode == 1 or mode == 2:
enb = get_mb(nbmaxg,atlistg,opt_mon_eng)
# Printing results
be = 0.0
for i in range(len(enb)):
enb[i] = [k * 627.503 for k in enb[i]]
be += sum(enb[i])
print(str(i+1) + "b = " + str(sum(enb[i])))
if nbmaxg == nb:
print("Interaction Energy = " + str(be - sum(enb[0])))
print("Binding Energy = " + str(be))
for i in range(len(enb)):
print(" ")
print("Individual " + str(i+1) + "-body contributions:")
for j in range(len(enb[i])):
print("Fragment " + str(j+1) + ": " + str(enb[i][j]) )
elif mode == 3:
nb = len(atlistg)
nm = []
monsN = range(1,nb + 1)
comb = []
enb = []
for i in range(1,nb+1):
comb.append(list(it.combinations(monsN,i)))
nm.append(len(comb[i-1]))
for i in range(nbmaxg):
enb.append([])
foldname = str(i + 1) + "b"
for j in range(len(comb[i])):
atl = []
monen = []
for k in range(len(comb[i][j])):
atl.append(atlistg[comb[i][j][k] - 1])
monen.append(opt_mon_eng[comb[i][j][k]] - 1)
os.chdir(foldname + "/" + str(j+1))
enbx = get_mb(i+1,atl,monen)
enb[i].append(enbx[i][0])
os.chdir("../../")
# Printing results
be = 0.0
for i in range(len(enb)):
enb[i] = [k * 627.503 for k in enb[i]]
be += sum(enb[i])
print(str(i+1) + "b = " + str(sum(enb[i])))
if nbmaxg == nb:
print("Interaction Energy = " + str(be - sum(enb[0])))
print("Binding Energy = " + str(be))
for i in range(len(enb)):
print(" ")
print("Individual " + str(i+1) + "-body contributions:")
for j in range(len(enb[i])):
print("Fragment " + str(j+1) + ": " + str(enb[i][j]) )
```
| github_jupyter |
<h1><center> Προσπαρασκευή 1ης εργαστηριακής άσκησης:</center></h1>
<h2><center> Μέρος πρώτο: Ορθογράφος</center><h2>
### ΕΚΤΕΛΕΣΗ
### Βήμα 1: Κατασκευή corpus
__α)__ Κατεβάζουμε λοιπόν από το project Gutenberg το βιβλίο __War of the Worlds__ σε plain txt μορφή και το αποθηκεύουμε με το όνομα __War.txt__ (με το ! στην αρχή μιάς εντολής δηλώνουμε ότι ακολουθεί ένα shell command).
```
! wget -c http://www.gutenberg.org/files/36/36-0.txt -O War.txt
```
__β)__ Προφανώς μπορούμε να ενώσουμε περισσότερα βιβλία για την κατασκευή ενός μεγαλύτερου corpus. Τα πλεονεκτήματα αυτής της τεχνικής (εκτός από την αύξηση του μεγέθους των δεδομένων) είναι:
1. Τα δεδομένα μας θα προέρχονται από πολλές διαφορετικές πηγές με αποτέλεσμα το γλωσσικό μοντέλο μας να γενικεύσει περισσότερο και να μην κάνει overfit στα δεδομένα μας.
2. Όταν έχουμε ένα μεγάλο corpus τα στατιστικά στοιχεία που θα υπολογίσουμε πάνω σε αυτό θα είναι ακριβέστερα και ακόμα θα είμαστε σε θέση να κρίνουμε ποιό στατιστικό στοιχείο είναι σημαντικότερο για το μοντέλο μας.
### Βήμα 2: Προεπεξεργασία corpus
Έχουμε λοιπόν το __corpus__ που θα χρησιμοποιήσουμε στον ίδιο φάκελο με το notebook μας. Θα ορίσουμε τώρα ορισμένες χρήσιμες συναρτήσεις για την προεπεξεργασία του.
__α)__ Η συνάρτηση __identity_preprocess__ διαβάζει ένα string και απλά γυρνάει τον εαυτό του.
```
def identity_preprocess(string_var):
return string_var
```
__β)__ Η συνάρτηση __read_path__ δέχεται σαν όρισμα το path του αρχείου που θέλουμε να διαβάσουμε και μία συνάρτηση preprocess και διαβάζει το αρχείο γραμμή προς γραμμή σε μία λίστα, καλώντας την preprocess σε κάθε γραμμή. Χρησιμοποιούμε σαν default όρισμα για την preprocess την identity_preprocess που ορίστηκε παραπάνω.
```
def read_path(file_path, preprocess = identity_preprocess):
#initilize the list of processed lines
processed_lines = []
#open file to read mode
with open(file_path, "r") as f:
for line in f:
#omit spaces
if not line.isspace():
processed_lines.extend(preprocess(line))
return processed_lines
```
__γ)__ Η συνάρτηση __tokenize__ δέχεται σαν όρισμα ένα string s και:
1. καλεί την strip() και lower() πάνω στο s,
2. αφαιρεί όλα τα σημεία στίξης / σύμβολα / αριθμούς, αφήνωντας μόνο αλφαριθμητικούς χαρακτήρες,
3. αντικαθιστά τα newlines με κένα,
4. κάνει split() τις λέξεις στα κενά.
```
import string
def tokenize(s):
#Remove possible spaces from the start or the end of the string and
#turn all letters lowercase.
s = s.strip().lower()
#Remove all punctuations, symbols and numbers from the string leaving
#only lowercase alphabetical letters.
s = "".join((char for char in s if char not in string.punctuation and not char.isdigit()))
#Replace new line characters with spaces
s = s.replace('\n',' ')
#Split the string in every space resulting in a list of tokens
res = s.split(" ")
return res
```
__δ)__ Μπορούμε αντί να κατασκευάσουμε τον δικό μας tokenizer να χρησιμοποιήσουμε έναν έτοιμο από την βιβλιοθήκη nltk που διαθέτει πόλλους διαφορετικούς tokenizer. Παρακάτω ορίζουμε έναν από αυτούς τον οποίο θα συγκρίνουμε αργότερα με τον δικό μας.
```
from nltk.tokenize import WordPunctTokenizer
def nltk_tokenizer(s):
words = WordPunctTokenizer().tokenize(text = s)
words = [word.lower() for word in words if word.isalpha() and not word.isdigit()]
return words
```
Στη συνέχεια μπορούμε να συγκρίνουμε τους δύο tokenizers δίνοντας τους ως είσοδο ένα string με διάφορα είδη χαρακτήρων, συμβόλων και αριθμών. Παρατηρούμε ότι και οι δύο ανταποκρίνονται πολύ καλά δίνοντάς μας το επιθυμητό αποτέλεσμα, απλά ο δικός μας αποθηκεύει και κάποια κενά ως token το οποίο μπορεί να διορθωθεί κατά την δημιουργία του λεξικού.
```
test_string = "hi my !@( name 32 is ... Panos"
print("My tokenizer :" + " ", end="")
print(tokenize(test_string))
print("Nltk tokenizer :" + " ", end="")
print(nltk_tokenizer(test_string))
```
### Βήμα 3: Κατασκευή λεξικού και αλφαβήτου
__α)__ Δημιουργούμε μία λίστα η οποία θα περιέχει όλα τα μοναδικά tokens που βρίσκονται στο corpus που θα χρησιμοποιήσουμε, σύμφωνα με τον tokenizer του Βήματος 2. Ουσιαστικά θα καλέσουμε την συνάρτηση read_path με όρισμα τον tokenizer που θέλουμε να εφαρμοστεί σε κάθε γραμμή του αρχείου και στη συνέχεια θα πάρουμε μόνο τις μοναδικές λέξεις για να σχηματίσουμε το λεξικό μας.
```
def get_tokens(file_path):
tokens = read_path(file_path, tokenize)
distinct_tokens = list(dict.fromkeys(tokens))
return distinct_tokens
tokens_dict = get_tokens("War.txt")
```
__β)__ Δημιουργούμε μία λίστα που περιέχει το αλφάβητο του corpus.
```
def get_alphabet(tokens):
alphabet = []
for token in tokens:
alphabet.extend(list(token))
alphabet = list(dict.fromkeys(alphabet))
return alphabet
alphabet = get_alphabet(tokens_dict)
```
### Βήμα 4: Δημιουργία συμβόλων εισόδου-εξόδου
Η συνάρτηση __alphabet _to_int__ δέχεται ως όρισμα το αλφάβητο που έχουμε ορίσει από το corpus, αντιστοιχίζει κάθε χαρακτήρα σε ένα αύξοντα ακέραιο index και γράφει το αποτέλεσμα σε ένα αρχείο chars.syms με αυτή τη μορφή http://www.openfst.org/twiki/pub/FST/FstExamples/ascii.syms. Σε όλα τα επόμενα βήματα θα χρησιμοποιήσουμε το αρχείο chars.syms για τα σύμβολα εισόδου και εξόδου των FSTs.
```
def alphabet_to_int(alphabet):
# open file
f = open("chars.syms", "w")
# match epsilon to 0
f.write("EPS" + 7*" " + str(0) + '\n')
num = 21
for character in alphabet:
# match every other character to an increasing index
f.write(character + 7*" " + str(num) + '\n')
num += 1
f.close()
alphabet_to_int(alphabet)
```
### Βήμα 5: Κατασκευή μετατροπέων FST
__α)__ Για να κατασκευάσουμε τον μετατροπέα σε αυτό το βήμα αλλά και τον αποδοχέα στο επόμενο θα χρειαστούμε μία συνάρτηση format_arc η οποία θα διαμορφώνει μία γραμμή από το αρχείο περιγραφής του κάθε FST. Συγκεκριμένα δέχεται ως όρισμα τα __src__, __dest__, __ilabel__, __olabel__ και το __weight__ (με default τιμή το 0) και τα επιστρέφει στην κατάλληλη μορφή όπως αναφέρεται και εδώ http://www.openfst.org/twiki/bin/view/FST/FstQuickTour#CreatingFsts/.
```
def format_arc(src, dest, ilabel, olabel, weight=0):
return (str(src) + " " + str(dest) + " " + str(ilabel) + " " + str(olabel) + " " + str(weight))
```
Τώρα λοιπόν θα διαμορφώσουμε το αρχείο περιγραφής του μετατροπέα μας, ο οποίος θα έχει μία κατάσταση και θα υλοποιεί την απόσταση Levenshtein αντιστοιχίζοντας:
1. Κάθε χαρακτήρα στον εαυτό του με βάρος 0 (no edit).
2. Κάθε χαρακτήρα στο EPS με βάρος 1 (deletion).
3. Το EPS σε κάθε χαρακτήρα με βάρος 1 (insertion).
4. Κάθε χαρακτήρα σε κάθε άλλο χαρακτήρα με βάρος 1 (substitution).
Η περιγραφή του μετατροπέα αποθηκεύεται στο αρχείο __transducer.fst__.
```
# get alphabet of the corpus
alphabet = get_alphabet(get_tokens("War.txt"))
# open file to write mode
f = open("transducer.fst", "w")
for letter in alphabet:
# no edit
f.write(format_arc(0, 0, letter, letter) + "\n")
# deletion
f.write(format_arc(0, 0, letter, "EPS", 1) + "\n")
# insertion
f.write(format_arc(0, 0, "EPS", letter, 1) + "\n")
for i in range(len(alphabet)):
for j in range(len(alphabet)):
if i != j:
# substitution
f.write(format_arc(0, 0, alphabet[i], alphabet[j], 1) + "\n")
# make initial state also final state
f.write("0")
# close file
f.close()
```
Τρέχοντας λοιπόν το παρακάτω shell command κάνουμε compile τον μετατροπέα μας. Το binary αρχείο που προκύπτει με όνομα __transducer.bin.fst__ είναι αυτό που θα χρησιμοποιήσουμε στις επόμενες λειτουργίες.
```
! ./compile_transducer.sh
```
Είναι προφανές ότι ο μετατροπέας μας αυτή την στιγμή σε μία λέξη εισόδου αν πάρουμε το shortest path απλά θα επιστρέψει την ίδια την λέξη αυτούσια χωρίς καμμία αλλαγή. Και αυτό γιατί για κάθε σύμβολο του αλφαβήτου έχουμε μετάβαση στον εαυτό του με κόστος 0. Ο μετατροπέας μας θα αποκτήσει νόημα όταν τον συνθέσουμε με τον αποδοχέα του επόμενου ερωτήματος.
__β)__ Αυτός είναι ένας αρκετά αφελής τρόπος για τον υπολογισμό των βαρών για κάθε edit. Αν τώρα είχαμε στη διάθεση μας ό,τι δεδομένα θέλουμε αυτό που θα κάναμε είναι ότι θα υπολογίζαμε τα βάρη με βάση το πόσο συχνά γίνεται αυτό το λάθος. Πιο συγκεκριμένα, θα υπολογίζαμε για κάθε σύμβολο του αλφαβήτου την πιθανότητα κάποιος να το διαγράψει, να το προσθέσει ή να το αντικαταστήσει με κάποιο άλλο. Στη συνέχεια, θα μετατρέπαμε αυτές τις πιθανότητες σε κόστη παίρνοντας τον αρνητικό λογάριθμο και θα είχαμε τα τελικά βάρη μας για κάθε σύμβολο στο deletion και το insertion και για κάθε δυάδα συμβόλων στο substitution.
### Βήμα 6: Κατασκευή αποδοχέα λεξικού
__α)__ Σε αυτό το βήμα θέλουμε να κατασκευάσουμε έναν αποδοχέα με μία αρχική κατάσταση που αποδέχεται κάθε κατάσταση του λεξικού μας. Τα βάρη όλων των ακμών θα είναι 0. Αντίστοιχα με πριν γράφουμε την περιγραφή του αποδοχέα σε ένα αρχείο __acceptor.fst__.
```
# get tokens of the corpus (our acceptor should accept only these words)
tokens = get_tokens("War.txt")
# open file to write mode
f = open("acceptor.fst", "w")
s = 1
for token in tokens:
letters = list(token)
for i in range(0, len(letters)):
if i == 0:
# For each token make state 1 its first state
f.write(format_arc(1, s+1, letters[i], letters[i]) + "\n")
else:
f.write(format_arc(s, s+1, letters[i], letters[i]) + "\n")
s += 1
if i == len(letters) - 1:
# When reaching the end of a token go to final state 0 though an ε-transition
f.write(format_arc(s, 0, "EPS", "EPS") + "\n")
# make state 0 final state
f.write("0")
# close the file
f.close()
```
Στη συνέχεια το fst μας γίνεται compile στο αρχείο __acceptor.bin.fst__.
```
! ./compile_acceptor.sh
```
__β)__ Στη συνέχεια καλούμε τις συναρτήσεις fstrmepsilon, fstdeterminize και fstminimize για να βελτιστοποιήσουμε το μοντέλο μας. Παρακάτω αναφέρεται η λειτουργία της κάθεμίας:
1. __fstrmepsilon__: Μετατρέπει το FST σε ένα ισοδύναμο που να μην περιέχει ε-μεταβάσεις
2. __fstdeterminize__: Μετατρέπει το FST σε ένα ισοδύναμο ντετερμινιστικό. Το τελικό FST, δηλαδή, θά έχει για κάθε κατάσταση και για κάθε σύμβολο εισόδου μία και μόνο μετάβαση.
2. __fstminimize__: Ελαχιστοποιεί το FST ώστε να έχει τον ελάχιστο αριθμό καταστάσεων.
```
! fstrmepsilon acceptor.bin.fst acceptor.bin.fst
! fstdeterminize acceptor.bin.fst acceptor.bin.fst
! fstminimize acceptor.bin.fst acceptor.bin.fst
```
### Βήμα 7: Κατασκευή ορθογράφου
Στο βήμα αυτό θα συνθέσουμε τον Levenshtein transducer με τον αποδοχέα του ερωτήματος 6α παράγοντας τον min edit distance spell checker. Αρχικά θα ταξινομήσουμε τις εξόδους του transducer και τις εισόδους του αποδοχέα με την συνάρτηση __fstarcsort__.
```
! fstarcsort --sort_type=olabel transducer.bin.fst transducer_sorted.fst
! fstarcsort --sort_type=ilabel acceptor.bin.fst acceptor_sorted.fst
```
Στη συνέχεια συνθέτουμε τον transducer με τον αποδοχέα με την συνάρτηση fstcompose αποθηκεύοντας τον min edit distance spell checker στο αρχείο __spell_checker.fst__.
```
! fstcompose transducer_sorted.fst acceptor_sorted.fst spell_checker.fst
```
Η συμπεριφορά του μετατροπέα που έχουμε υλοποιήσει διαφέρει ανάλογα με τα βάρη που θα βάλουμε σε κάθε edit. Συγκεκριμένα:
1. Στην περίπτωση που τα edits είναι ισοβαρή έχουμε ένα αφελή transducer που απλά διορθώνει τις λέξεις χωρίς να λαμβάνει υπόψιν του κάποια γλωσσική πληροφορία, με μόνο κριτήριο να κάνει τις ελάχιστες δυνατές μετατροπές στην λέξη εισόδου.
2. Στην περίπτωση τώρα που τα edits δεν είναι ισοβαρή μπορούμε να υλοποιήσουμε έναν πιο ισχυρό trandsucer. Συγκεκριμένα, μπορούμε να ορίσουμε με βάση κάποια γνωστά μοντέλα την πιθανότητα ένα σύμβολο να λείπει, να έχει προστεθεί λανθασμένα ή να έχει αντικατασταθεί από κάποιο άλλο σύμβολο και να μετατρέψουμε αυτές τις πιθανότητες σε κόστη τα οποία θα μεταφραστούν στη συνέχεια ως βάρη στα αντίστοιχα edits. Έτσι, η λέξη θα διορθώνεται όχι με βάση τις ελάχιστες δυνατές μετατροπές αλλά με το ποιά λέξη είναι πιο πίθανο να ήθελε να γράψει αυτός που την έγραψε λάθος.
__β)__ Στην περίπτωση, λοιπόν, που η είσοδος στον min edit spell checker μας είναι η λέξη __cit__ θα προσπαθήσει με τον ελάχιστο αριθμό που τις τρεις λειτουργίες που του επιτρέπονται (insert, delete, substitute) να την μετατρέψει σε μία λέξη που υπάρχει στο λεξικό μας. Επειδή, βέβαια, το λεξικό μας προέρχεται απλά από ένα βιβλίο είναι λογικό πολλές δημοφιλής λέξεις να λείπουν. Στην περίπτωση μας έχουμε τις εξής πιθανές προβλέψεις:
1. Με μία αλλαγή μόνο:
* __cat__
* __sit__
* __fit__
* __cut__
* __cite__
2. Σε περίπτωση τώρα που καμία από τις παραπάνω λέξεις δεν βρίσκεται στο λεξικό μας ο ορθογράφος θα κοιτάξει στις λέξεις που προκύπτουν από δύο αλλαγές στις λέξεις κ.ο.κ.
### Βήμα 8: Αξιολόγηση ορθογράφου
__α)__ Για να κάνουμε το evaluation του ορθογράφου κατεβάζουμε το παρακάτω σύνολο δεδομένων:
```
! wget https://raw.githubusercontent.com/georgepar/python-lab/master/spell_checker_test_set
```
__β)__ Από το παραπάνω test set λοιπόν θα επιλεξουμε 20 λέξεις στην τύχη και με βάση τον αλγόριθμο των ελάχιστων μονοπατιών στο γράφο του μετατροπέα του βήματος 7 θα την διορθώσουμε και θα εκτιμήσουμε την διόρθωση.
Δημιουργούμε αρχικά μία συνάρτηση __predict__ η οποία δέχεται μία λέξη που πρέπει να διορθωθεί και γράφει σε ένα αρχείο __pred_word.fst__ την περιγραφή ενός FST το οποίο αποδέχεται την συγκεκριμένη λέξη. Το FST αυτό θα το κάνουμε στη συνέχεια compose με τον ορθογράφο μας για να πάρουμε το τελικό αποτέλεσμα.
```
def predict(word):
s= 1
letters = list(word)
# open file to write mode
f = open("pred_word.fst", "w")
for i in range(0, len(letters)):
# for each letter of the word make a transition with zero weight
f.write(format_arc(s, s+1, letters[i], letters[i], 0) + '\n')
s += 1
if i == len(letters) - 1:
# when reaching the end the word make a ε-transition to the final state 0
f.write(format_arc(s, 0, "EPS", "EPS", 0) + '\n')
# final state
f.write("0")
# close the file
f.close()
```
Στη συνέχεια, επιλέγουμε από το test set 20 τυχαίες λέξεις. Συγκεκριμένα, επιλέγουμε 20 τυχαίες σειρές και από κάθε σειρά επιλέγουμε τυχαία μία λανθασμένη λέξη. Βάζουμε ένα συγκεκριμένο seed έτσι ώστε να προκύπτουν οι ίδιες λέξεις κάθε φορά και να γίνει η εκτίμηση αργότερα.
```
import random
random.seed(1)
test_words = []
for _ in range(20):
random_lines = random.choice(open('spell_checker_test_set').readlines())
test_words.append(random.choice(random_lines.strip('\n').split()[1:]))
```
Τέλος, για κάθε λέξη καλούμε την συνάρτηση predict και στη συνέχεια τρέχουμε στο bash (με το σύμβολο !) το αρχείο predict.sh δίνοντας ως όρισμα τον ορθογράφο μας.
```
for word in test_words:
print(word + ":" + " ",end='')
predict(word)
! ./predict.sh spell_checker.fst
print('\n')
```
##### Σχολιασμός αποτελεσμάτων:
Παρατηρούμε ότι ο ορθογράφος λειτουργεί με μικρή ακρίβεια. Υπάρχουν συγκεκριμένα λάθη σε κάποιες λέξεις που οφείλονται στο γεγονός ότι τα βάρη στον μετατροπέα είναι όλα 1 και ότι το corpus μας αποτελείται μόνο από ένα βιβλίο.
1. __chosing__ το οποίο διορθώνεται σε chasing αλλά θα μπορούσε να διορθωθεί σε choosing το οποίο έχει ίδιο αριθμό αλλαγών με το chasing αλλά θα μπορύσαμε να πούμε ότι όταν δύο γράμματα είναι συνεχόμενα είναι πολύ πιθανό να παραλειφθεί το ένα από τα δύο. Άρα το λάθος αυτό οφείλεται στά ίδια βάρη.
2. __aranging__, __awfall__, __defenition__, __compair__, __transfred__, __adress__, __juise__ που θα μπορούσαν να γίνουν με λιγότερα edits σε arranging,awful, definition, compare, transfered, address, juice αντίστοιχα. Τα λάθη αυτά οφείλονται στο ότι οι συγκεκριμένες λέξεις δεν υπήρχας στο corpus μας.
### Βήμα 9: Εξαγωγή αναπαραστάσεων word2vec
__α)__ Αρχικά θα διαβάσουμε το κείμενο σε μία λίστα από tokenized προτάσεις χρησιμοποιώντας τον κώδικα του βήματος 2γ. Για να χωρίσουμε το corpus σε προτάσεις χρησιμοποιούμε την βιβλιοθήκη __nltk__.
```
import nltk
# We split the corpus in a list of tokenized sentences.
file_path = "War.txt"
tokenized_sentences = []
with open(file_path, "r") as f:
text = f.read()
sentences = nltk.sent_tokenize(text)
tokenized_sentences = [tokenize(sentence) for sentence in sentences]
```
__β)__ Θα χρησιμοποιήσουμε την κλάση Word2Vec του gensim για να εκπαιδεύσουμε 100-διάστατα word2vec embeddings με βάση τις προτάσεις του βήματος 9α. Θα χρησιμοποιήσουμε window = 5 και 1000 εποχές.
```
from gensim.models import Word2Vec
# Initialize word2vec. Context is taken as the 2 previous and 2 next words
model = Word2Vec(tokenized_sentences, window=5, size=100, workers=4)
# Train the model for 1000 epochs
model.train(tokenized_sentences, total_examples=len(sentences), epochs=1000)
# get ordered vocabulary list
voc = model.wv.index2word
# get vector size
dim = model.vector_size
import numpy as np
# Convert to numpy 2d array (n_vocab x vector_size)
def to_embeddings_Matrix(model):
embedding_matrix = np.zeros((len(model.wv.vocab), model.vector_size))
for i in range(len(model.wv.vocab)):
embedding_matrix[i] = model.wv[model.wv.index2word[i]]
return embedding_matrix, model.wv.index2word
```
__γ)__ Θα επιλέξουμε 10 τυχαίες λέξεις από το λεξικό μας και θα βρούμε τις σημασιολογικά κοντινότερές τους.
```
import random
selected_words = random.sample(voc, 10)
for word in selected_words:
# get most similar words
sim = model.wv.most_similar(word, topn=5)
print('"' + word + '"' + " is similar with the following words:")
for s in sim:
print('"' + s[0] + '"' + " with similarity " + str(s[1]))
print()
```
Θα επιλέξουμε τώρα μεγαλύτερο παράθυρο κρατώντας ίδιο τον αριθμό των εποχών.
```
# Initialize word2vec. Context is taken as the 2 previous and 2 next words
model = Word2Vec(tokenized_sentences, window=20, size=100, workers=4)
# Train the model for 1000 epochs
model.train(tokenized_sentences, total_examples=len(sentences), epochs=1000)
# get ordered vocabulary list
voc = model.wv.index2word
# get vector size
dim = model.vector_size
import random
selected_words = random.sample(voc, 10)
for word in selected_words:
# get most similar words
sim = model.wv.most_similar(word, topn=5)
print('"' + word + '"' + " is similar with the following words:")
for s in sim:
print('"' + s[0] + '"' + " with similarity " + str(s[1]))
print()
```
Αντίθετα τώρα θα αυξήσουμε τον αριθμό των εποχών κρατώντας το μέγεθος του παραθύρου σταθερό.
```
# Initialize word2vec. Context is taken as the 2 previous and 2 next words
model = Word2Vec(tokenized_sentences, window=5, size=100, workers=4)
# Train the model for 1000 epochs
model.train(tokenized_sentences, total_examples=len(sentences), epochs=3000)
# get ordered vocabulary list
voc = model.wv.index2word
# get vector size
dim = model.vector_size
import random
selected_words = random.sample(voc, 10)
for word in selected_words:
# get most similar words
sim = model.wv.most_similar(word, topn=5)
print('"' + word + '"' + " is similar with the following words:")
for s in sim:
print('"' + s[0] + '"' + " with similarity " + str(s[1]))
print()
```
#### Σχολιασμός Αποτελεσμάτων
Τα αποτελέσματα δεν είναι τόσο ποιοτικά όσο περιμέναμε. Παρατηρούμε ότι σε κάποιες περιπτώσεις πετυχαίνει 1-2 λέξεις που όντως είναι σημασιολογικά κοντά με την επιλεγόμενη λέξη. Ακόμα και όταν αυξήσαμε το μέγεθος του παραθύρου ή τον αριθμό των εποχών η κατάσταση παρέμεινε η ίδια.Κατά την γνώμη μου αυτό συμβαίνει διότι έχουμε ένα πολύ μικρό corpus που δεν βοηθάει το μοντέλο ώστε να καταλάβει ποιες λέξεις είναι πραγματικά κοντά σημασιολογικά.
| github_jupyter |
# Pong with dqn
## Step 1: Import the libraries
```
import time
import gym
import random
import torch
import numpy as np
from collections import deque
import matplotlib.pyplot as plt
import math
import sys
sys.path.append('../../')
from algos.agents.dqn_agent import DQNAgent
from algos.models.dqn_cnn import DQNCnn
from algos.preprocessing.stack_frame import preprocess_frame, stack_frame
```
## Step 2: Create our environment
Initialize the environment in the code cell below.
```
env = gym.make('SpaceInvaders-v0')
env.seed(0)
# if gpu is to be used
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
print("Device: ", device)
```
## Step 3: Viewing our Enviroment
```
print("The size of frame is: ", env.observation_space.shape)
print("No. of Actions: ", env.action_space.n)
env.reset()
plt.figure()
plt.imshow(env.reset())
plt.title('Original Frame')
plt.show()
```
### Execute the code cell below to play Pong with a random policy.
```
def random_play():
score = 0
env.reset()
while True:
env.render()
action = env.action_space.sample()
state, reward, done, _ = env.step(action)
score += reward
if done:
env.close()
print("Your Score at end of game is: ", score)
break
random_play()
```
## Step 4:Preprocessing Frame
```
env.reset()
plt.figure()
plt.imshow(preprocess_frame(env.reset(), (8, -12, -12, 4), 84), cmap="gray")
plt.title('Pre Processed image')
plt.show()
```
## Step 5: Stacking Frame
```
def stack_frames(frames, state, is_new=False):
frame = preprocess_frame(state, (8, -12, -12, 4), 84)
frames = stack_frame(frames, frame, is_new)
return frames
```
## Step 6: Creating our Agent
```
INPUT_SHAPE = (4, 84, 84)
ACTION_SIZE = env.action_space.n
SEED = 0
GAMMA = 0.99 # discount factor
BUFFER_SIZE = 100000 # replay buffer size
BATCH_SIZE = 64 # Update batch size
LR = 0.0001 # learning rate
TAU = 1e-3 # for soft update of target parameters
UPDATE_EVERY = 1 # how often to update the network
UPDATE_TARGET = 10000 # After which thershold replay to be started
EPS_START = 0.99 # starting value of epsilon
EPS_END = 0.01 # Ending value of epsilon
EPS_DECAY = 100 # Rate by which epsilon to be decayed
agent = DQNAgent(INPUT_SHAPE, ACTION_SIZE, SEED, device, BUFFER_SIZE, BATCH_SIZE, GAMMA, LR, TAU, UPDATE_EVERY, UPDATE_TARGET, DQNCnn)
```
## Step 7: Watching untrained agent play
```
# watch an untrained agent
state = stack_frames(None, env.reset(), True)
for j in range(200):
env.render()
action = agent.act(state)
next_state, reward, done, _ = env.step(action)
state = stack_frames(state, next_state, False)
if done:
break
env.close()
```
## Step 8: Loading Agent
Uncomment line to load a pretrained agent
```
start_epoch = 0
scores = []
scores_window = deque(maxlen=20)
```
## Step 9: Train the Agent with DQN
```
epsilon_by_epsiode = lambda frame_idx: EPS_END + (EPS_START - EPS_END) * math.exp(-1. * frame_idx /EPS_DECAY)
plt.plot([epsilon_by_epsiode(i) for i in range(1000)])
def train(n_episodes=1000):
"""
Params
======
n_episodes (int): maximum number of training episodes
"""
for i_episode in range(start_epoch + 1, n_episodes+1):
state = stack_frames(None, env.reset(), True)
score = 0
eps = epsilon_by_epsiode(i_episode)
while True:
action = agent.act(state, eps)
next_state, reward, done, info = env.step(action)
score += reward
next_state = stack_frames(state, next_state, False)
agent.step(state, action, reward, next_state, done)
state = next_state
if done:
break
scores_window.append(score) # save most recent score
scores.append(score) # save most recent score
print('\rEpisode {}\tAverage Score: {:.2f}'.format(i_episode, np.mean(scores_window)), end="")
if i_episode % 100 == 0:
print('\rEpisode {}\tAverage Score: {:.2f}'.format(i_episode, np.mean(scores_window)))
fig = plt.figure()
ax = fig.add_subplot(111)
plt.plot(np.arange(len(scores)), scores)
plt.ylabel('Score')
plt.xlabel('Episode #')
plt.show()
return scores
scores = train(1000)
```
## Step 10: Watch a Smart Agent!
```
score = 0
state = stack_frames(None, env.reset(), True)
while True:
env.render()
action = agent.act(state)
next_state, reward, done, _ = env.step(action)
score += reward
state = stack_frames(state, next_state, False)
if done:
print("You Final score is:", score)
break
env.close()
```
| github_jupyter |
Example based on https://machinelearningmastery.com/binary-classification-tutorial-with-the-keras-deep-learning-library/
Data: https://archive.ics.uci.edu/ml/datasets/Connectionist+Bench+(Sonar,+Mines+vs.+Rocks)
activity_regularizer: https://machinelearningmastery.com/how-to-reduce-generalization-error-in-deep-neural-networks-with-activity-regularization-in-keras/
http://proceedings.mlr.press/v15/glorot11a.html
https://keras.io/layers/writing-your-own-keras-layers/
```
import numpy
import pandas
from tensorflow.keras.layers import Dense
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import Pipeline
import wvpy.util
from YConditionalRegularizedModel import YConditionalRegularizedModel
from eval_model import eval_model
# load dataset
dataframe = pandas.read_csv("sonar.csv", header=None)
dataframe.shape
dataframe
# deal with outcome column, which is the last column
last_col = dataframe.shape[1] - 1
Y = numpy.asarray(dataframe[last_col])
# encode class values as integers
encoder = LabelEncoder()
encoder.fit(Y)
encoded_Y = encoder.transform(Y)
Y = None
encoded_Y
X = dataframe.values[:, 0:last_col] # clean copy, no outcome columns
dataframe = None
X.shape
def example_model_steps():
return [
60, # input vector width
Dense(60, activation='relu'),
Dense(1, activation='sigmoid'),
]
batch_size = 100
estimators = list()
estimators.append(('standardize', StandardScaler()))
estimators.append(('mlp',
YConditionalRegularizedModel(
example_model_steps, alpha=0.01,
epochs=100, batch_size=batch_size, verbose=0)))
pipeline = Pipeline(estimators)
preds = pipeline.fit_transform(X, encoded_Y)
wvpy.util.dual_density_plot(probs=preds[:,1], istrue=encoded_Y, title="Smoothed In-Sample Performance")
preds_out, results = eval_model(model=pipeline, X=X, y=encoded_Y)
print("Smoothed: %.2f%% (%.2f%%)" % (results.mean()*100, results.std()*100))
wvpy.util.dual_density_plot(
probs=preds_out['prediction'], istrue=encoded_Y,
title="Smoothed Out-Of-Sample Performance")
wvpy.util.plot_roc(
prediction=preds_out['prediction'], istrue=encoded_Y,
title="Smoothed Out-Of-Sample Performance")
encoded_Y_shuffled = encoded_Y.copy()
numpy.random.shuffle(encoded_Y_shuffled)
preds_shuffled = pipeline.fit_transform(X, encoded_Y_shuffled)
wvpy.util.dual_density_plot(
probs=preds_shuffled[:,1], istrue=encoded_Y_shuffled,
title="Shuffled Smoothed In-Sample Performance")
preds_out_shuffled, results_shuffled = eval_model(model=pipeline, X=X, y=encoded_Y_shuffled)
print("Shuffled Smoothed: %.2f%% (%.2f%%)" % (results_shuffled.mean()*100, results_shuffled.std()*100))
wvpy.util.dual_density_plot(
probs=preds_out_shuffled['prediction'], istrue=encoded_Y_shuffled,
title="Shuffled Smoothed Out-Of-Sample Performance")
pf = pandas.DataFrame({
'prediction': preds_out['prediction'],
'truth': encoded_Y
})
estimators0 = list()
estimators0.append(('standardize', StandardScaler()))
estimators0.append(('mlp',
YConditionalRegularizedModel(
example_model_steps, alpha=0,
epochs=100, batch_size=batch_size, verbose=0)))
pipeline0 = Pipeline(estimators0)
preds0 = pipeline0.fit_transform(X, encoded_Y)
wvpy.util.dual_density_plot(
probs=preds0[:,1], istrue=encoded_Y,
title="Not Smoothed In-Sample Performance")
preds_out0, results0 = eval_model(model=pipeline0, X=X, y=encoded_Y)
print("Not smoothed: %.2f%% (%.2f%%)" % (results0.mean()*100, results0.std()*100))
wvpy.util.dual_density_plot(
probs=preds_out0['prediction'], istrue=encoded_Y,
title="Not Smoothed Out-Of-Sample Performance")
preds0s = pipeline0.fit_transform(X, encoded_Y_shuffled)
wvpy.util.dual_density_plot(
probs=preds0s[:,1], istrue=encoded_Y_shuffled,
title="Not Smoothed Shuffled In-Sample Performance")
preds_out0s, results0s = eval_model(model=pipeline0, X=X, y=encoded_Y_shuffled)
print("Not smoothed shuffle : %.2f%% (%.2f%%)" % (results0s.mean()*100, results0s.std()*100))
wvpy.util.dual_density_plot(
probs=preds_out0s['prediction'], istrue=encoded_Y_shuffled,
title="Not Smoothed Shuffled Out-Of-Sample Performance")
```
| github_jupyter |
# Preamble
```
from flair.models import SequenceTagger
from flair.data import Sentence, Token
from nltk import word_tokenize
import stanfordnlp
import nltk
import json
import spacy
import re
PATH_WIKIPEDIA_EXTRACTION = "../../data/causality-graphs/extraction/"
PATH_WIKIPEDIA_EXTRACTION += "wikipedia/wikipedia-extraction.tsv-backup"
PATH_FLAIR_FOLDER = "../../data/flair-models/lists/"
PATH_NLTK_RESOURCES = "../../data/external/nltk/"
PATH_STANFORD_RESOURCES = "../../data/external/stanfordnlp/"
PATH_OUTPUT_GRAPH = "../../data/causality-graphs/spotting/wikipedia/list-graph.json"
nltk.download('punkt', PATH_NLTK_RESOURCES)
nltk.data.path.append(PATH_NLTK_RESOURCES)
stanfordnlp.download('en', PATH_STANFORD_RESOURCES)
stanford_nlp = stanfordnlp.Pipeline(processors='tokenize,pos',
models_dir=PATH_STANFORD_RESOURCES,
treebank='en_ewt',
use_gpu=True)
stanford_nlp_pretokenized = stanfordnlp.Pipeline(processors='tokenize,pos',
models_dir=PATH_STANFORD_RESOURCES,
treebank='en_ewt',
tokenize_pretokenized=True,
use_gpu=True)
# Spacy version: 2.1.8
# Model version: 2.1.0
# pip install https://github.com/explosion/spacy-models/releases/download/en_core_web_sm-2.1.0/en_core_web_sm-2.1.0.tar.gz
spacy_nlp = spacy.load('en_core_web_sm')
print(spacy.__version__)
print(spacy_nlp.meta['version'])
```
# Loading Lists
```
list_set = []
for line in open(PATH_WIKIPEDIA_EXTRACTION, encoding="utf-8"):
parts = line.strip().split('\t')
if parts[0] != 'wikipedia_list':
continue
assert len(parts) == 9
list_data = {
"list": json.loads(parts[8]),
"type": "wikipedia_list",
"payload": {
"wikipedia_page_id": parts[1],
"wikipedia_page_title": parts[2],
"wikipedia_revision_id": parts[3],
"wikipedia_revision_timestamp": parts[4],
"list_toc_parent_title": parts[5][1:-1],
"list_toc_section_heading": parts[6][1:-1],
"list_toc_section_level": parts[7]
}
}
list_set.append(list_data)
```
# Preprocessing
```
def is_valid_article(title):
forbidden_title_parts = ['Wikipedia:', 'Template:', 'File:',
'Portal:', 'Category:', 'Draft:',
'List of', 'disambiguation']
contains_forbidden_title_part = False
for forbidden_title_part in forbidden_title_parts:
if forbidden_title_part in title:
contains_forbidden_title_part = True
break
return not contains_forbidden_title_part
def is_in_causal_section(section_title, section_level):
allowed_sections = ['Cause', 'Causes', 'Risk factor',
'Risk factors' 'Symptom', 'Symptoms',
'Signs and symptoms']
return section_title in allowed_sections and section_level == '2'
list_data_preprocessed = []
for list_data in list_set:
if not is_valid_article(list_data['payload']['wikipedia_page_title']):
continue
list_toc_section_heading = list_data['payload']['list_toc_section_heading']
list_toc_section_level = list_data['payload']['list_toc_section_level']
if not is_in_causal_section(list_toc_section_heading,
list_toc_section_level):
continue
list_data_preprocessed.append(list_data)
def get_pos(sentence):
tags = []
for token in sentence.tokens:
for word in token.words:
tags.append((word.text, word.pos))
return tags
def pos_tagging(doc):
tags = []
for sentence in doc.sentences:
for pos in get_pos(sentence):
tags.append(pos)
return tags
def is_verb(pos_tag):
'''
List of verb-POS-Tags:
VB - Verb, base form
VBD - Verb, past tense
VBG - Verb, gerund or present participle
VBN - Verb, past participle
VBP - Verb, non-3rd person singular present
VBZ - Verb, 3rd person singular present
'''
return pos_tag in ['VB', 'VBD', 'VBG', 'VBN', 'VBP', 'VBZ']
def preprocess(string):
# split newlines
newline_split = [elem.strip()
for elem in string.split('\n')
if elem.strip() != '']
# remove brackets
for i in range(len(newline_split)):
newline_split[i] = re.sub(r"\s?\([^)]*\)\s?",
"",
newline_split[i]).strip()
'''
- remove 'See ...'
- remove paragraphs
- (multiple sentences; contains more than a sentence -> is paragraph)
- remove whole sentences (phrases with verb)
'''
result = []
for string in newline_split:
# Remove 'See ...'
if re.match(r"^See\s.*", string) is not None:
continue
# Remove leading list headings like "Initially: ... Later: ..."
# heading at beginning
string = re.sub(r"^\w+:\s?", "", string).strip()
# heading within the string
string = re.sub(r"\w+:\s?", "", string).strip()
if string is None or string == '':
continue
doc = stanford_nlp(string)
# Remove paragraphs
if len(doc.sentences) != 1:
continue
# Remove whole sentences (with verb)
contains_verb = False
pos_tags = pos_tagging(doc)
for i in range(len(pos_tags)):
if is_verb(pos_tags[i][1]):
contains_verb = True
break
if contains_verb:
continue
result.append(string)
return result
for list_data in list_data_preprocessed:
list_data['list'] = preprocess(list_data['list'])
list_data_for_spotting = []
for list_data in list_data_preprocessed:
for data in list_data['list']:
result = {'list': data,
"sources": [{'type': list_data['type'],
'payload': list_data['payload']}]}
list_data_for_spotting.append(result)
```
# POS-Tagging
```
def get_offset_of_tags_in_sentence(sentence, tags):
# go from left to right and determine tag offsets
offsets = []
total_offset = 0
for tag in tags:
label = tag[0]
local_offset = sentence.find(label)
offset = total_offset + local_offset
offsets.append(offset)
# prepare for next iteration
sentence = sentence[local_offset + len(label):]
total_offset = offset + len(label)
return offsets
def get_pos_tags_of_sentence(sentence):
tags = []
for token in sentence.tokens:
for word in token.words:
tags.append((word.text, word.pos))
return tags
def calculate_pos_tags_for_string(doc):
tags = []
for sentence in doc.sentences:
sentence_pos = []
for pos in get_pos_tags_of_sentence(sentence):
sentence_pos.append(pos)
tags.append(sentence_pos)
return tags
def pos_tagging(strings):
batch = '\n\n'.join([' '.join(word_tokenize(string))
for string in strings])
doc = stanford_nlp_pretokenized(batch)
tags = calculate_pos_tags_for_string(doc)
assert len(tags) == len(strings)
result = []
for i in range(len(strings)):
offsets = get_offset_of_tags_in_sentence(strings[i], tags[i])
result.append([(tags[i][x][0], tags[i][x][1], str(offsets[x]))
for x in range(len(tags[i]))])
return result
strings = []
for sample in list_data_for_spotting:
strings.append(sample['list'])
tags = pos_tagging(strings)
for i in range(len(list_data_for_spotting)):
sample = list_data_for_spotting[i]
sample['list:POS'] = tags[i]
```
# List-Spotter: Prediction
```
def get_index(chunk):
return [int(chunk.tokens[0].get_tag('idx').value),
int(chunk.tokens[-1].get_tag('idx').value)
+ len(chunk.tokens[-1].text)]
classifier = SequenceTagger.load(PATH_FLAIR_FOLDER + 'final-model.pt')
for sample in list_data_for_spotting:
sentence = Sentence(use_tokenizer=False)
for pos in sample['list:POS']:
token = Token(pos[0])
token.add_tag('POS', pos[1])
token.add_tag('idx', pos[2])
sentence.add_token(token)
classifier.predict(sentence)
chunks = [get_index(chunk) for chunk in sentence.get_spans('chunk_BIO')]
extraction = []
for chunk in chunks:
extraction.append(sample['list'][chunk[0]:chunk[1]])
sample['extraction'] = extraction
```
# Post-processing
```
def spacy_tagging(sentence):
doc = spacy_nlp(sentence)
tagging = []
for token in doc:
tagging.append((token.text, token.tag_, token.idx))
return tagging
def post_process_value(value):
if value is None:
return value
tags = spacy_tagging(value)
left = 0
right = len(tags)-1
punctuation = ['.', ',', ';', '(', ')', '``', "''"]
cutoff = ['CC', 'DT', 'PRP', 'PRP$'] + punctuation
for tag in tags:
if tag[1] in cutoff:
left += 1
else:
break
for tag in reversed(tags):
if tag[1] in cutoff:
right -= 1
else:
break
try:
return value[tags[left][2]:tags[right][2] + len(tags[right][0])]
except:
return value
def get_relation(section_heading, parent_title, value):
if section_heading in ['Symptom', 'Symptoms', 'Signs and symptoms']:
return (parent_title, value)
if section_heading in ['Cause', 'Causes']:
return (value, parent_title)
if section_heading in ['Risk factor', 'Risk factors']:
return (value, parent_title)
raise Exception("Not handled.")
list_graph = []
for sample in list_data_for_spotting:
for value in sample['extraction']:
value = post_process_value(value)
if len(value) == 0:
continue
source = sample['sources'][0]['payload']
relation = get_relation(
source['list_toc_section_heading'],
source['list_toc_parent_title'],
value)
causal_relation = {'causal_relation': {
'cause': {'concept': relation[0]},
'effect': {'concept': relation[1]},
}, 'sources': sample['sources']}
list_graph.append(causal_relation)
```
# Save List-Graph
```
jsonarray = json.dumps(list_graph)
file_list_graph = open(PATH_OUTPUT_GRAPH, "w+")
file_list_graph.write(jsonarray)
file_list_graph.close()
```
| github_jupyter |
## 4-2. トロッター分解を用いた量子シミュレーション
この節では量子系のダイナミクスのシミュレーションする手法の一つである、トロッター分解に基づいたダイナミクスシミュレーションを紹介する。この手法はダイナミクスのシミュレーションとしてはある意味最も愚直な手法であり、NISQデバイスでも一定の規模の問題までは動作する。
ここでは、1次元の横磁場イジングモデルという物理系の量子ダイナミクスをシミュレートしてみる。
### トロッター分解とは
[(リー・)トロッター分解](https://ja.wikipedia.org/wiki/リー・トロッター積公式)とは、正方行列$A,B$の和の指数関数を、それぞれの指数関数の積に近似する公式である:
$$
e^{\delta (A+B)} = e^{\delta A} \cdot e^{\delta B} + O\left( \delta^2 \right).
$$
ここで$O$は2.3節コラムで紹介したオーダー記法であり、近似の大体の精度を表す。($A,B$が行列なので、$e^{A+B}\neq e^A\cdot e^B$ であることに注意)
### トロッター分解を用いた量子シミュレーションの仕組み
前節で学んだように、量子状態の時間発展はシュレディンガー方程式$i\frac{\partial}{\partial t}|\psi(t)\rangle = H|\psi(t)\rangle$に従う($\hbar=1$とした)。特に、ハミルトニアン$H$が時間に依存しない場合、時間$t$が経った時の状態は$|\psi(t)\rangle = e^{-iHt}|\psi(0)\rangle$となる($|\psi(0)\rangle$は初期状態)。つまり、量子系のダイナミクスのシミュレーションでは$e^{-iHt}$という演算子が計算できれば良いのだが、$n$量子ビット系だと$H$は$2^n$次元となって非常に大きな行列になる。ゆえに古典コンピュータでこの指数関数を計算するのは難しいし、量子コンピュータでもあらゆる$2^n$次元の行列$H$の指数関数を計算するのは難しいと考えられている。
しかし、ハミルトニアンが**特定の構造**を持つ場合、量子コンピュータは$e^{-iHt}$を効率よく計算できる。その際に用いるのがトロッター分解である。
具体例で考えてみよう。$n$量子ビットからなるイジングモデル(後述)$H = \sum_{i=1}^{n-1} Z_i Z_{i+1}$ を考える。この$H$は$2^n$次元の行列であるが、トロッター分解を用いると$e^{-iH t}$を
$$
e^{-iHt} = e^{-i\left(\sum_{i=1}^{n-1} Z_i Z_{i+1}\right) t} = \left(e^{-i(Z_1 Z_2 + Z_2 Z_3 \cdots )\frac{t}{M}}\right)^M
\approx \left(e^{-i(Z_1 Z_2)\frac{t}{M}}\cdot e^{-i(Z_2 Z_3)\frac{t}{M}}\cdot\cdots \right)^M \tag{1}
$$
のように分解できる。$M$は$e^{-iHt}$の作用を分割する総数で、近似の精度$O\left((t/M)^2\right)$が十分小さくなるように選んでおく。
ここで最後の式に注目すると、小さい行列(2量子ビットゲート、4次元)である $e^{-i(Z_i Z_{i+1})\frac{t}{M}}$の$nM$個の積でかけていることが分かる。つまり、**指数的に大きな行列の作用を、小さな行列(2量子ビットゲート)の多項式個の作用の積で書き表すことができた**のである。
このように、ハミルトニアンが少数の項の和で書ける場合には、トロッター分解を用いて量子コンピュータで高速に計算を行うことができるのだ。
※ ちなみに、物理・量子化学の分野で興味のあるハミルトニアンは、大抵は効率的なトロッター分解ができる形をしている。
### 量子ダイナミクスの実装(1):イジングモデル
量子シミュレーションを実装する具体的な系として、イジングモデルというものを考えてみよう。
(物理学に詳しくない読者は、とりあえず定義だけ受け入れて計算を追うので問題ない)
物質が磁場に反応する性質を磁性というが、ミクロな世界では電子や原子核が非常に小さな磁性を持ち、**スピン**と呼ばれている。大雑把には、スピンとは小さな磁石のことであると思って良い。私たちが日常目にする磁石は、物質中の電子(原子核)のスピンが示す小さな磁性が綺麗に揃った結果、巨大な磁性が現れたものなのである。
イジングモデルは、そんな物質中のスピンを振る舞いを記述するモデルで、磁石の本質を取り出したモデルとして考案された。定義は以下の通りである。
$$
H = J \sum_{i=1}^{n} Z_i Z_{i+1}
$$
ここで$J$はモデルのパラメータ、$n$はスピンを持つ粒子の数、$Z_n$は$n$番目の粒子のスピンの$z$軸方向の成分を表す。前節で学んだように、物理量はオブザーバブルというエルミート演算子で表されるので、スピンが示す磁化(磁力の強さ)も演算子で表される。空間は三次元だから、独立な磁化の成分として$x,y,z$軸の三方向の磁化が考えられ、それぞれ対応する演算子は$X,Y,Z$である。(パウリ演算子$X,Y,Z$の名前の由来はここから来ている)
ざっくりいうと、$n$番目の粒子=量子ビットであり、$Z|0\rangle=|0\rangle$, $Z|1\rangle= -|1\rangle$だから、$|0\rangle$はスピンが$+z$方向を向いている状態、$|1\rangle$はスピンが$-z$方向を向いている状態である。
このモデルを構成する$Z_i Z_{i+1}$は隣り合う量子ビット間の相互作用を表している。$J>0$の時、 $(Z_i,Z_{i+1})=(1,1),(-1,-1)$ で $Z_i Z_{i+1}=1$、$(Z_i,Z_{i+1})=(1,-1),(-1,1)$ で $Z_i Z_{i+1}=-1$ だから、エネルギーが低いのはスピンが互い違いになる場合$|010101\cdots\rangle$である。逆に$J<0$ではスピンが揃った場合 $|000\cdots\rangle, |111\cdots\rangle$ がエネルギーが低く安定になり、系は巨大な磁化を持った磁石として振る舞う。
さて、実はこのモデルそのものは量子コンピュータを使わなくても簡単に解けてしまうが、まずはこのモデルでトロッター分解の基本を実装してみよう。$e^{-iHt}$のトロッター分解は(1)式で導いているので、必要になるのは$e^{-i \delta Z_i Z_{i+1}}$というゲートをどのように実装するかである。これは次のようにすれば良い(計算して確かめてみてほしい):
$$
e^{-i \delta Z_i Z_{i+1}} = \operatorname{CNOT}_{i,i+1} \cdot e^{-i\delta Z_{i+1}} \cdot \operatorname{CNOT}_{i,i+1}.
$$
今回は**z方向の全磁化**
$$
\left\langle\psi\left|\sum_{i=1}^{n} Z_i\right|\psi\right\rangle
$$
という物理量に着目して、その時間変化を追いかける。
**初期状態はスピンが**$+z$**方向に揃った**$|000\cdots\rangle$とし、便宜上、系が周期的になっているとして$Z_{n+1}$ は $Z_1$ と同一視する。
```
## Google Colaboratoryの場合・Qulacsがインストールされていないlocal環境の場合のみ実行してください
!pip install qulacs
#必要なライブラリをインポート
from qulacs import QuantumState,QuantumCircuit, Observable, PauliOperator
from qulacs.gate import X,Z,RX,RY,RZ,CNOT,merge,DenseMatrix,add
from qulacs.state import inner_product
import matplotlib.pyplot as plt
import numpy as np
%matplotlib inline
# 今回は粒子6個の系を想定する。
nqubits = 6
# ダイナミクスをシミュレーションする時間
t = 1.0
# トロッター分解の分割数
M = 100
# 時間の刻み幅
delta = t/M
## 全磁化に対応するオブザーバブルを準備しておく
magnetization_obs = Observable(nqubits)
for i in range(nqubits):
magnetization_obs.add_operator(PauliOperator("Z "+str(i), 1.0/nqubits))
## 初期状態は|000000>
state = QuantumState(nqubits)
state.set_zero_state()
# トロッター分解の1回分、e^{iZ_1Z_2*delta}*e^{iZ_2Z_3*delta}*...e^{iZ_nZ_1*delta} を量子ゲートに変換
circuit_trotter_Ising = QuantumCircuit(nqubits)
for i in range(nqubits):
circuit_trotter_Ising.add_CNOT_gate(i,(i+1)%(nqubits))
circuit_trotter_Ising.add_RZ_gate((i+1)%nqubits,2*delta) ## RZ(a)=exp(i*a/2*Z)に注意
circuit_trotter_Ising.add_CNOT_gate(i,(i+1)%(nqubits))
## 時間と磁化を記録するリスト
x = [i*delta for i in range(M+1)]
y = []
#t=0の時の全磁化のみ先に計算
y.append( magnetization_obs.get_expectation_value(state) )
#t=0以降の全磁化を計算
for i in range(M):
# delta=t/Mだけ時間発展
circuit_trotter_Ising.update_quantum_state(state)
# 磁化を計算して記録
y.append(magnetization_obs.get_expectation_value(state))
#グラフの描画
plt.xlabel("time")
plt.ylabel("Value of magnetization")
plt.title("Dynamics of Ising model")
plt.plot(x, y, "-")
plt.show()
```
この結果を見ると分かるように、$z$方向の全磁化は一定である(縦軸のスケールにも注意)。
実は、イジングモデルでは$z$方向の相互作用しかないので、$z$方向の全磁化も保存してしまうのだ。これでは面白くないので、次は$x$軸方向に磁場を加えた横磁場イジングモデルの時間発展を見てみよう。
### 量子ダイナミクスの実装(2):横磁場イジングモデル
イジングモデルに、X軸方向の一様な磁場をかけた横磁場イジングモデルを考えよう。
$$
H = \sum_{i=1}^n Z_i Z_{i+1} + h\sum_{i=1}^n X_i
$$
この$h$は横磁場の強さを表す係数で、$X_i$は$i$番目の粒子のx方向の磁化を表すパウリ演算子(オブザーバブル)である。
このモデルでトロッター分解を行うと、先ほどのイジングモデルに加えて$e^{-ih X_1 \frac{t}{M}} \cdots e^{-ih X_n \frac{t}{M}}$という項が加わることになる。
では実際にこのモデルでの全磁化の動きを見てみよう。
```
# 今回は粒子6個の系を想定する。
nqubits = 6
# ダイナミクスをシミュレーションする時間
t = 3.0
# トロッター分解の分割数
M = 100
# 時間の刻み幅
delta = t/M
## 横磁場の強さ
h = 3.
## 全磁化に対応するオブザーバブルを準備しておく
magnetization_obs = Observable(nqubits)
for i in range(nqubits):
magnetization_obs.add_operator(PauliOperator("Z "+str(i), 1.0/nqubits))
## 初期状態は|000000>
state = QuantumState(nqubits)
state.set_zero_state()
# トロッター分解の1回分、
# e^{iZ_1Z_2*delta}*e^{iZ_2Z_3*delta}*...e^{iZ_nZ_1*delta} * e^{i X_1*delta}*...*e^{i X_n*delta} を量子ゲートに変換
circuit_trotter_transIsing = QuantumCircuit(nqubits)
for i in range(nqubits):
circuit_trotter_transIsing.add_CNOT_gate(i,(i+1)%(nqubits))
circuit_trotter_transIsing.add_RZ_gate((i+1)%nqubits,2*delta) ## RZ(a)=exp(i*a/2*Z)に注意
circuit_trotter_transIsing.add_CNOT_gate(i,(i+1)%(nqubits))
circuit_trotter_transIsing.add_RX_gate(i, 2*delta*h) ## RX(a)=exp(i*a/2*X)に注意
## 時間と磁化を記録するリスト
x = [i*delta for i in range(M+1)]
y = []
#t=0の時の全磁化のみ先に計算
y.append( magnetization_obs.get_expectation_value(state) )
#t=0以降の全磁化を計算
for i in range(M):
# delta=t/Mだけ時間発展
circuit_trotter_transIsing.update_quantum_state(state)
# 磁化を計算して記録
y.append(magnetization_obs.get_expectation_value(state))
#グラフの描画
plt.xlabel("time")
plt.ylabel("Value of magnetization")
plt.title("Dynamics of transverse Ising model")
plt.plot(x, y, "-")
plt.show()
```
$h=0$のイジングモデルの場合は全磁化の値が一定だったのに対し、横磁場$h=3$を入れた横磁場イジングモデルの場合は全磁化の値がその後0近くまで下がっている点で大きく違う。
これは横磁場の影響でz方向に揃っていたスピンが乱れて乱雑になり、結果として全磁化の値が変化していると考えれば良い。興味のある読者は$h$を強くして計算してみると、磁化の減少するスピードが速くなることが分かるであろう。
### 量子ダイナミクスの実装(3):厳密解との比較
トロッター分解には誤差がある。上で計算したダイナミクスがどれほどの精度のなのか、$e^{-iHt}$を直接計算した厳密なダイナミクスと比べてみよう。
```
# 今回は粒子6個の系を想定する。
nqubits = 6
# ダイナミクスをシミュレーションする時間
t = 3.0
# トロッター分解の分割数
M = 100
# 時間の刻み幅
delta = t/M
## 横磁場の強さ
h = 3.
## 全磁化に対応するオブザーバブル.
magnetization_obs = Observable(nqubits)
for i in range(nqubits):
magnetization_obs.add_operator(PauliOperator("Z "+str(i), 1.0))
## 初期状態は|000000>
state_trotter = QuantumState(nqubits)
state_trotter.set_zero_state()
state_exact = QuantumState(nqubits)
state_exact.set_zero_state()
# トロッター分解の1回分、
# e^{iZ_1Z_2*delta}*e^{iZ_2Z_3*delta}*...e^{iZ_nZ_1*delta} * e^{i X_1*delta}*...*e^{i X_n*delta} を量子ゲートに変換
circuit_trotter_transIsing = QuantumCircuit(nqubits)
for i in range(nqubits):
circuit_trotter_transIsing.add_CNOT_gate(i,(i+1)%(nqubits))
circuit_trotter_transIsing.add_RZ_gate((i+1)%nqubits,2*delta) ## RZ(a)=exp(i*a/2*Z)に注意
circuit_trotter_transIsing.add_CNOT_gate(i,(i+1)%(nqubits))
circuit_trotter_transIsing.add_RX_gate(i, 2*delta*h) ## RX(a)=exp(i*a/2*X)に注意
# e^{-iHt}を直接対角化する。Hの行列表現を得るために、gateを生成してそのmatrixを取得する
zz_matrix = np.array([[1,0,0,0],[0,-1,0,0],[0,0,-1,0],[0,0,0,1]]) ## Z_i*Z_{i+1}の行列表示
hx_matrix = h*np.array( [ [0,1], [1,0] ] )
zz = DenseMatrix([0,1], zz_matrix) ## 0~1間の相互作用
hx = DenseMatrix(0, hx_matrix) ## 0サイトへの横磁場
## qulacs.gate.addを用いて、1以降のサイトの相互作用と横磁場を足していく
for i in range(1, nqubits):
zz = add(zz, DenseMatrix([i,(i+1)%nqubits], zz_matrix))
hx = add(hx, DenseMatrix(i, hx_matrix) )
## 最終的なハミルトニアン
ham = add(zz, hx)
matrix = ham.get_matrix() #行列の取得
eigenvalue, P = np.linalg.eigh(np.array(matrix)) #取得した行列の固有値、固有ベクトルを取得
## e^{-i*H*delta}を行列として作る
e_iHdelta = np.diag(np.exp(-1.0j*eigenvalue*delta))
e_iHdelta = np.dot(P, np.dot(e_iHdelta, P.T))
## 回路に変換
circuit_exact_transIsing = QuantumCircuit(nqubits)
circuit_exact_transIsing.add_dense_matrix_gate( np.arange(nqubits), e_iHdelta)
## 時間と磁化を記録するリスト
x = [i*delta for i in range(M+1)]
y_trotter = []
y_exact = []
#t=0の時の全磁化のみ先に計算
y_trotter.append( magnetization_obs.get_expectation_value(state_trotter) )
y_exact.append( magnetization_obs.get_expectation_value(state_exact) )
#t=0以降の全磁化を計算
for i in range(M):
# delta=t/Mだけ時間発展
circuit_trotter_transIsing.update_quantum_state(state_trotter)
circuit_exact_transIsing.update_quantum_state(state_exact)
# 磁化を計算して記録
y_trotter.append( magnetization_obs.get_expectation_value(state_trotter) )
y_exact.append( magnetization_obs.get_expectation_value(state_exact) )
#グラフの描画
plt.xlabel("time")
plt.ylabel("Value of magnetization")
plt.title("Dynamics of transverse Ising model")
plt.plot(x, y_trotter, "-", label="Trotter")
plt.plot(x, y_exact, "-", label="exact")
plt.legend()
plt.show()
```
この範囲では、どうやらほぼ一致しているようだ。誤差を見てみよう。
```
#グラフの描画
plt.xlabel("time")
plt.ylabel("Value of magnetization")
plt.title("Error")
plt.plot(x, np.abs(np.array(y_trotter) - np.array(y_exact)), "-", label="Trotter")
plt.legend()
plt.show()
```
興味のある読者は、分割数$M$を荒くしたり、時間$t$を大きくしたり色々と試してみてほしい。
| github_jupyter |
```
import findspark
findspark.init()
import pyspark
import pyspark.sql
from pyspark import SparkConf
from pyspark import SparkContext
conf = SparkConf()#
conf.set("spark.python.profile", "true")
#conf = sc.getConf()
conf.set( "spark.driver.memory", "4g" )
conf.set( "spark.jars","/Users/crisliu/agile/lib/mongo-hadoop-spark-2.0.2.jar,\
/Users/crisliu/agile/lib/mongo-java-driver-3.4.0.jar,\
/Users/crisliu/agile/lib/elasticsearch-spark-20_2.10-5.0.0-alpha5.jar,\
/Users/crisliu/agile/lib/snappy-java-1.1.2.6.jar,\
/Users/crisliu/agile/lib/lzo-hadoop-1.0.5.jar")
#/Users/crisliu/agile/lib/elasticsearch-hadoop-5.0.0-alpha5.jar"
#conf.set("spark.executor.extraClassPath","~")
sc = SparkContext('local', 'test', conf=conf)
#sc = pyspark.SparkContext()
APP_NAME = "my_script.py"
spark = pyspark.sql.SparkSession(sc).builder.appName(APP_NAME).getOrCreate()
base_path = '/Users/crisliu/agile/'
#
# {
# "ArrDelay":5.0,"CRSArrTime":"2015-12-31T03:20:00.000-08:00","CRSDepTime":"2015-12-31T03:05:00.000-08:00",
# "Carrier":"WN","DayOfMonth":31,"DayOfWeek":4,"DayOfYear":365,"DepDelay":14.0,"Dest":"SAN","Distance":368.0,
# "FlightDate":"2015-12-30T16:00:00.000-08:00","FlightNum":"6109","Origin":"TUS"
# }
#
from pyspark.sql.types import StringType, IntegerType, FloatType, DoubleType, DateType, TimestampType
from pyspark.sql.types import StructType, StructField
from pyspark.sql.functions import udf
schema = StructType([
StructField("ArrDelay", DoubleType(), True), # "ArrDelay":5.0
StructField("CRSArrTime", TimestampType(), True), # "CRSArrTime":"2015-12-31T03:20:00.000-08:00"
StructField("CRSDepTime", TimestampType(), True), # "CRSDepTime":"2015-12-31T03:05:00.000-08:00"
StructField("Carrier", StringType(), True), # "Carrier":"WN"
StructField("DayOfMonth", IntegerType(), True), # "DayOfMonth":31
StructField("DayOfWeek", IntegerType(), True), # "DayOfWeek":4
StructField("DayOfYear", IntegerType(), True), # "DayOfYear":365
StructField("DepDelay", DoubleType(), True), # "DepDelay":14.0
StructField("Dest", StringType(), True), # "Dest":"SAN"
StructField("Distance", DoubleType(), True), # "Distance":368.0
StructField("FlightDate", DateType(), True), # "FlightDate":"2015-12-30T16:00:00.000-08:00"
StructField("FlightNum", StringType(), True), # "FlightNum":"6109"
StructField("Origin", StringType(), True), # "Origin":"TUS"
])
input_path = "{}/Agile_Data_Code_2/data/simple_flight_delay_features.jsonl.bz2".format(
base_path
)
features = spark.read.json(input_path, schema=schema)
features.first()
#
# Check for nulls in features before using Spark ML
#
null_counts = [(column, features.where(features[column].isNull()).count()) for column in features.columns]
cols_with_nulls = filter(lambda x: x[1] > 0, null_counts)
print(list(cols_with_nulls))
#
# Add a Route variable to replace FlightNum
#
from pyspark.sql.functions import lit, concat
features_with_route = features.withColumn(
'Route',
concat(
features.Origin,
lit('-'),
features.Dest
)
)
features_with_route.show(6)
#
# Use pysmark.ml.feature.Bucketizer to bucketize ArrDelay into on-time, slightly late, very late (0, 1, 2)
#
from pyspark.ml.feature import Bucketizer
# Setup the Bucketizer
splits = [-float("inf"), -15.0, 0, 30.0, float("inf")]
arrival_bucketizer = Bucketizer(
splits=splits,
inputCol="ArrDelay",
outputCol="ArrDelayBucket"
)
# Apply the bucketizer
ml_bucketized_features = arrival_bucketizer.transform(features_with_route)
ml_bucketized_features.select("ArrDelay", "ArrDelayBucket").show()
ml_bucketized_features
#
# Extract features tools in with pyspark.ml.feature
#
from pyspark.ml.feature import StringIndexer, VectorAssembler
# Turn category fields into indexes
for column in ["Carrier", "Origin", "Dest", "Route"]:
print(column)
string_indexer = StringIndexer(
inputCol=column,
outputCol=column + "_index"
)
string_indexer_model = string_indexer.fit(ml_bucketized_features)
ml_bucketized_features = string_indexer_model.transform(ml_bucketized_features)
# Drop the original column
ml_bucketized_features = ml_bucketized_features.drop(column)
# Save the pipeline model
string_indexer_output_path = "{}/models/string_indexer_model_{}.bin".format(
base_path,
column
)
string_indexer_model.write().overwrite().save(string_indexer_output_path)
# Combine continuous, numeric fields with indexes of nominal ones
# ...into one feature vector
numeric_columns = [
"DepDelay", "Distance",
"DayOfMonth", "DayOfWeek",
"DayOfYear"]
index_columns = ["Carrier_index", "Origin_index",
"Dest_index", "Route_index"]
vector_assembler = VectorAssembler(
inputCols=numeric_columns + index_columns,
outputCol="Features_vec"
)
final_vectorized_features = vector_assembler.transform(ml_bucketized_features)
# Drop the index columns
for column in index_columns:
final_vectorized_features = final_vectorized_features.drop(column)
# Inspect the finalized features
final_vectorized_features.show()
# Instantiate and fit random forest classifier on all the data
from pyspark.ml.classification import RandomForestClassifier
rfc = RandomForestClassifier(
featuresCol="Features_vec",
labelCol="ArrDelayBucket",
predictionCol="Prediction",
maxBins=4657,
maxMemoryInMB=1024
)
model = rfc.fit(final_vectorized_features)
# Evaluate model using test data
predictions = model.transform(final_vectorized_features)
from pyspark.ml.evaluation import MulticlassClassificationEvaluator
evaluator = MulticlassClassificationEvaluator(
predictionCol="Prediction",
labelCol="ArrDelayBucket",
metricName="accuracy"
)
accuracy = evaluator.evaluate(predictions)
print("Accuracy = {}".format(accuracy))
# Check the distribution of predictions
predictions.groupBy("Prediction").count().show()
# Check a sample
predictions.sample(False, 0.001, 18).orderBy("CRSDepTime").show(6)
# Save the pipeline model
string_indexer_output_path = "{}/models/string_indexer_model_{}.bin".format(
base_path,
column
)
string_indexer_model.write().overwrite().save(string_indexer_output_path)
# Save the numeric vector assembler
vector_assembler_path = "{}/models/numeric_vector_assembler.bin".format(base_path)
vector_assembler.write().overwrite().save(vector_assembler_path)
# Save the bucketizer
arrival_bucketizer_path = "{}/models/arrival_bucketizer_2.0.bin".format(base_path)
arrival_bucketizer.write().overwrite().save(arrival_bucketizer_path)
# Save the new model over the old one
model_output_path = "{}/models/spark_random_forest_classifier.flight_delays.5.0.bin".format(base_path)
model.write().overwrite().save(model_output_path)
```
| github_jupyter |
Data from EPS fig to TIKZ converter
```
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
data_1colon2 = """(600.8320,1527.5200) -- (600.8320,1525.0200) --
(601.6680,1524.1900) -- (601.6680,1521.6900) -- (602.5000,1520.8500) --
(602.5000,1516.6900) -- (603.3320,1515.8500) -- (603.3320,1513.3500) --
(604.1680,1511.6900) -- (604.1680,1508.3500) -- (605.0000,1506.6900) --
(605.0000,1504.1900) -- (605.8360,1503.3500) -- (605.8360,1499.1900) --
(606.6680,1498.3500) -- (606.6680,1495.8500) -- (607.5000,1495.0200) --
(607.5000,1490.8500) -- (608.3360,1490.0200) -- (608.3360,1485.8500) --
(609.1680,1485.0200) -- (609.1680,1482.5200) -- (610.0000,1480.8500) --
(610.0000,1477.5200) -- (610.8360,1475.8500) -- (610.8360,1473.3500) --
(611.6680,1472.5200) -- (611.6680,1468.3500) -- (612.5000,1467.5200) --
(612.5000,1465.0200) -- (613.3360,1464.1800) -- (613.3360,1460.0200) --
(614.1680,1459.1800) -- (614.1680,1456.6800) -- (615.0000,1455.0200) --
(615.0000,1451.6800) -- (615.8360,1450.0200) -- (615.8360,1447.5200) --
(616.6680,1446.6800) -- (616.6680,1442.5200) -- (617.5000,1441.6800) --
(617.5000,1439.1800) -- (618.3360,1437.5200) -- (618.3360,1434.1800) --
(619.1680,1433.3500) -- (619.1680,1430.8500) -- (620.0000,1429.1800) --
(620.0000,1425.8500) -- (620.8360,1424.1800) -- (620.8360,1420.8500) --
(621.6680,1419.1800) -- (621.6680,1416.6800) -- (622.5000,1415.8500) --
(622.5000,1411.6800) -- (623.3360,1410.8500) -- (623.3360,1408.3500) --
(624.1680,1407.5200) -- (624.1680,1403.3500) -- (625.0000,1402.5200) --
(625.0000,1400.0200) -- (625.8360,1398.3500) -- (625.8360,1395.0200) --
(626.6680,1393.3500) -- (626.6680,1390.8500) -- (627.5000,1390.0200) --
(627.5000,1385.8500) -- (628.3360,1385.0200) -- (628.3360,1382.5200) --
(629.1680,1380.8500) -- (629.1680,1377.5200) -- (630.0000,1376.6800) --
(630.0000,1374.1800) -- (630.8360,1372.5200) -- (630.8360,1369.1800) --
(631.6680,1367.5200) -- (631.6680,1365.0200) -- (632.5000,1364.1800) --
(632.5000,1360.0200) -- (633.3360,1359.1800) -- (633.3360,1356.6800) --
(634.1680,1355.0200) -- (634.1680,1351.6800) -- (635.0000,1350.0200) --
(635.0000,1348.3500) -- (635.8360,1346.6800) -- (635.8360,1343.3500) --
(636.6680,1341.6800) -- (636.6680,1339.1800) -- (637.5000,1338.3500) --
(637.5000,1334.1800) -- (638.3360,1333.3500) -- (638.3360,1330.8500) --
(639.1680,1329.1800) -- (639.1680,1325.8500) -- (640.0000,1324.1800) --
(640.0000,1321.6800) -- (640.8360,1320.8500) -- (640.8360,1318.3500) --
(641.6680,1317.5200) -- (641.6680,1313.3500) -- (642.5000,1312.5200) --
(642.5000,1310.0200) -- (643.3360,1308.3500) -- (643.3360,1305.0200) --
(644.1680,1303.3500) -- (644.1680,1300.8500) -- (645.0000,1300.0200) --
(645.0000,1295.8500) -- (645.8360,1295.0200) -- (645.8360,1292.5200) --
(646.6680,1290.8500) -- (646.6680,1287.5200) -- (647.5000,1286.6800) --
(647.5000,1284.1800) -- (648.3360,1282.5200) -- (648.3360,1279.1800) --
(649.1680,1277.5200) -- (649.1680,1275.0200) --
(650.0000,1274.1800) -- (650.0000,1270.0200) --
(650.8320,1269.1900) -- (650.8320,1266.6900) -- (651.6680,1265.0200) --
(651.6680,1261.6900) -- (652.5000,1260.8500) -- (652.5000,1258.3500) --
(653.3320,1256.6900) -- (653.3320,1253.3500) -- (654.1680,1251.6900) --
(654.1680,1249.1900) -- (655.0000,1248.3500) -- (655.0000,1245.8500) --
(655.8320,1244.1900) -- (655.8320,1240.8500) -- (656.6680,1239.1900) --
(656.6680,1236.6900) -- (657.5000,1235.8500) -- (657.5000,1231.6900) --
(658.3360,1230.8500) -- (658.3360,1228.3500) -- (659.1680,1227.5200) --
(659.1680,1223.3500) -- (660.0000,1222.5200) -- (660.0000,1220.0200) --
(660.8360,1218.3500) -- (660.8360,1215.0200) -- (661.6680,1213.3500) --
(661.6680,1210.8500) -- (662.5000,1210.0200) -- (662.5000,1207.5200) --
(663.3360,1205.8500) -- (663.3360,1202.5200) -- (664.1680,1201.6900) --
(664.1680,1199.1900) -- (665.0000,1197.5200) -- (665.0000,1194.1900) --
(665.8360,1192.5200) -- (665.8360,1190.0200) -- (666.6680,1189.1900) --
(666.6680,1185.0200) -- (667.5000,1184.1900) -- (667.5000,1181.6900) --
(668.3360,1180.0200) -- (668.3360,1176.6900) -- (669.1680,1175.0200) --
(669.1680,1173.3500) -- (670.0000,1171.6900) -- (670.0000,1169.1900) --
(670.8360,1168.3500) -- (670.8360,1164.1900) -- (671.6680,1163.3500) --
(671.6680,1160.8500) -- (672.5000,1159.1900) -- (672.5000,1155.8500) --
(673.3360,1154.1900) -- (673.3360,1151.6900) -- (674.1680,1150.8500) --
(674.1680,1146.6900) -- (675.0000,1145.8500) -- (675.0000,1143.3500) --
(675.8360,1142.5200) -- (675.8360,1140.0200) -- (676.6680,1138.3500) --
(676.6680,1135.0200) -- (677.5000,1133.3500) -- (677.5000,1130.8500) --
(678.3360,1130.0200) -- (678.3360,1125.8500) -- (679.1680,1125.0200) --
(679.1680,1122.5200) -- (680.0000,1120.8500) -- (680.0000,1118.3500) --
(680.8360,1117.5200) -- (680.8360,1114.1900) -- (681.6680,1112.5200) --
(681.6680,1110.0200) -- (682.5000,1109.1900) -- (682.5000,1105.0200) --
(683.3360,1104.1900) -- (683.3360,1101.6900) -- (684.1680,1100.0200) --
(684.1680,1097.5200) -- (685.0000,1096.6900) -- (685.0000,1092.5200) --
(685.8360,1091.6900) -- (685.8360,1089.1900) -- (686.6680,1087.5200) --
(686.6680,1084.1900) -- (687.5000,1083.3500) -- (687.5000,1080.8500) --
(688.3360,1079.1900) -- (688.3360,1076.6900) -- (689.1680,1075.8500) --
(689.1680,1071.6900) -- (690.0000,1070.8500) -- (690.0000,1068.3500) --
(690.8360,1066.6900) -- (690.8360,1063.3500) -- (691.6680,1061.6900) --
(691.6680,1059.1900) -- (692.5000,1058.3500) -- (692.5000,1055.8500) --
(693.3360,1055.0200) -- (693.3360,1050.8500) -- (694.1680,1050.0200) --
(694.1680,1047.5200) -- (695.0000,1045.8500) -- (695.0000,1042.5200) --
(695.8360,1040.8500) -- (695.8360,1038.3500) -- (696.6680,1037.5200) --
(696.6680,1035.0200) -- (697.5000,1033.3500) -- (697.5000,1030.0200) --
(698.3360,1028.3500) -- (698.3360,1026.6900) -- (699.1680,1025.0200) --
(699.1680,1022.5200) -- (700.0000,1021.6900) -- (700.0000,1017.5200) --
(700.8360,1016.6900) -- (700.8360,1014.1900) -- (701.6680,1012.5200) --
(701.6680,1009.1900) -- (702.5000,1007.5200) -- (702.5000,1005.0200) --
(703.3360,1004.1900) -- (703.3360,1001.6900) -- (704.1680,1000.0200) --
(704.1680,996.6880) -- (705.0000,995.8520) -- (705.0000,993.3520) --
(705.8360,991.6880) -- (705.8360,989.1880) -- (706.6680,988.3520) --
(706.6680,984.1880) -- (707.5000,983.3520) -- (707.5000,980.8520) --
(708.3360,979.1880) -- (708.3360,976.6880) -- (709.1680,975.8520) --
(709.1680,971.6880) -- (710.0000,970.8520) -- (710.0000,968.3520) --
(710.8360,967.5200) -- (710.8360,963.3520) -- (711.6680,962.5200) --
(711.6680,960.0200) -- (712.5000,958.3520) -- (712.5000,955.8520) --
(713.3360,955.0200) -- (713.3360,950.8520) -- (714.1680,950.0200) --
(714.1680,947.5200) -- (715.0000,945.8520) -- (715.0000,943.3520) --
(715.8360,942.5200) -- (715.8360,939.1880) -- (716.6680,937.5200) --
(716.6680,935.0200) -- (717.5000,934.1880) -- (717.5000,931.6880) --
(718.3360,930.0200) -- (718.3360,926.6880) -- (719.1680,925.0200) --
(719.1680,922.5200) -- (720.0000,921.6880) -- (720.0000,919.1880) --
(720.8360,917.5200) -- (720.8360,914.1880) -- (721.6680,912.5200) --
(721.6680,910.0200) -- (722.5040,909.1880) -- (722.5040,906.6880) --
(723.3360,905.8520) -- (723.3320,901.6880) --
(724.1680,900.8520) -- (724.1680,898.3520) -- (725.0000,896.6880) --
(725.0000,894.1880) -- (725.8360,893.3520) -- (725.8360,889.1880) --
(726.6680,888.3520) -- (726.6680,885.8520) -- (727.5000,884.1880) --
(727.5000,881.6880) -- (728.3360,880.8520) -- (728.3360,877.5200) --
(729.1680,875.8520) -- (729.1680,873.3520) -- (730.0000,872.5200) --
(730.0000,870.0200) -- (730.8360,868.3520) -- (730.8360,865.0200) --
(731.6680,863.3520) -- (731.6680,860.8520) -- (732.5000,860.0200) --
(732.5000,857.5200) -- (733.3360,855.8520) -- (733.3360,852.5200) --
(734.1680,851.6880) -- (734.1680,849.1880) -- (735.0000,847.5200) --
(735.0000,845.0200) -- (735.8360,844.1880) -- (735.8360,841.6880) --
(736.6680,840.0200) -- (736.6680,836.6880) -- (737.5000,835.0200) --
(737.5000,832.5200) -- (738.3360,831.6880) -- (738.3360,829.1880) --
(739.1680,827.5200) -- (739.1680,824.1880) -- (740.0000,822.5200) --
(740.0000,820.8520) -- (740.8360,819.1880) -- (740.8360,816.6880) --
(741.6680,815.8520) -- (741.6680,811.6880) -- (742.5000,810.8520) --
(742.5000,808.3520) -- (743.3360,806.6880) -- (743.3360,804.1880) --
(744.1680,803.3520) -- (744.1680,800.8520) -- (745.0000,799.1880) --
(745.0000,795.8520) -- (745.8360,794.1880) -- (745.8360,792.5200) --
(746.6680,790.8520) -- (746.6680,788.3520) -- (747.5000,787.5200) --
(747.5000,783.3520) -- (748.3360,782.5200) -- (748.3360,780.0200) --
(749.1680,778.3520) -- (749.1680,775.8520) -- (750.0000,775.0200) --
(750.0000,772.5200) -- (750.8360,770.8520) -- (750.8360,767.5200) --
(751.6680,765.8520) -- (751.6680,763.3520) -- (752.5000,762.5200) --
(752.5000,760.0200) -- (753.3360,759.1880) -- (753.3360,755.0200) --
(754.1680,754.1880) -- (754.1680,751.6880) -- (755.0000,750.0200) --
(755.0000,747.5200) -- (755.8360,746.6880) -- (755.8360,744.1880) --
(756.6680,742.5200) -- (756.6680,739.1880) -- (757.5000,737.5200) --
(757.5000,735.0200) -- (758.3360,734.1880) -- (758.3360,731.6880) --
(759.1680,730.8520) -- (759.1680,726.6880) -- (760.0000,725.8520) --
(760.0000,723.3520) -- (760.8360,721.6880) -- (760.8360,719.1880) --
(761.6680,718.3520) -- (761.6680,715.8520) -- (762.5000,714.1880) --
(762.5000,710.8520) -- (763.3360,709.1880) -- (763.3360,706.6880) --
(764.1680,705.8520) -- (764.1680,703.3520) -- (765.0000,702.5200) --
(765.0000,700.0200) -- (765.8360,698.3520) -- (765.8360,695.0200) --
(766.6680,693.3520) -- (766.6680,690.8520) -- (767.5000,690.0200) --
(767.5000,687.5200) -- (768.3360,685.8520) -- (768.3360,683.3520) --
(769.1680,682.5200) -- (769.1680,678.3520) -- (770.0000,677.5200) --
(770.0000,675.0200) -- (770.8360,674.1880) -- (770.8360,671.6880) --
(771.6680,670.0200) -- (771.6680,667.5200) -- (772.5000,666.6880) --
(772.5000,662.5200) -- (773.3360,661.6880) -- (773.3360,659.1880) --
(774.1680,657.5200) -- (774.1680,655.0200) -- (775.0000,654.1880) --
(775.0000,651.6880) -- (775.8360,650.0200) -- (775.8360,646.6880) --
(776.6680,645.8520) -- (776.6680,643.3520) -- (777.5000,641.6880) --
(777.5000,639.1880) -- (778.3360,638.3520) -- (778.3360,635.8520) --
(779.1680,634.1880) -- (779.1680,630.8520) -- (780.0000,629.1880) --
(780.0000,626.6880) -- (780.8360,625.8520) -- (780.8360,623.3520) --
(781.6680,621.6880) -- (781.6680,619.1880) -- (782.5000,618.3520) --
(782.5000,615.8520) -- (783.3360,615.0200) -- (783.3360,610.8520) --
(784.1680,610.0200) -- (784.1680,607.5200) -- (785.0000,605.8520) --
(785.0000,603.3550) -- (785.8360,602.5200) -- (785.8360,600.0200) --
(786.6680,598.3550) -- (786.6680,595.0200) -- (787.5000,593.3550) --
(787.5000,590.8550) -- (788.3360,590.0200) -- (788.3360,587.5200) --
(789.1680,586.6880) -- (789.1680,584.1910) -- (790.0040,582.5200) --
(790.0040,580.0200) -- (790.8360,579.1910) -- (790.8360,575.0200) --
(791.6680,574.1910) -- (791.6680,571.6910) -- (792.5040,570.0200) --
(792.5040,567.5200) -- (793.3360,566.6910) -- (793.3360,564.1910) --
(794.1680,562.5200) -- (794.1680,559.1910) -- (795.0040,558.3520) --
(795.0040,555.8520) -- (795.8360,554.1910) -- (795.8360,551.6910) --
(796.6680,550.8520) -- (796.6680,548.3520) -- (797.5040,546.6910) --
(797.5040,544.1910) -- (798.3360,543.3520) -- (798.3360,539.1910) --
(799.1680,538.3520) -- (799.1680,536.6910) --
(799.1680,535.8520) -- (800.0000,534.1910) -- (800.0000,531.6910) --
(800.8320,530.8520) -- (800.8320,528.3520) -- (801.6680,527.5200) --
(801.6680,525.0200) -- (802.5000,523.3520) -- (802.5000,520.0200) --
(803.3320,518.3520) -- (803.3320,515.8520) -- (804.1680,515.0200) --
(804.1680,512.5200) -- (805.0000,510.8520) -- (805.0000,508.3520) --
(805.8320,507.5200) -- (805.8320,505.0200) -- (806.6680,503.3520) --
(806.6680,500.0200) -- (807.5000,499.1910) -- (807.5000,496.6910) --
(808.3360,495.0200) -- (808.3360,492.5200) -- (809.1680,491.6910) --
(809.1680,489.1910) -- (810.0000,487.5200) -- (810.0000,485.0200) --
(810.8360,484.1910) -- (810.8360,481.6910) -- (811.6680,480.0200) --
(811.6680,476.6910) -- (812.5000,475.0200) -- (812.5000,472.5200) --
(813.3360,471.6910) -- (813.3360,469.1910) -- (814.1680,468.3520) --
(814.1680,465.8520) -- (815.0000,464.1910) -- (815.0000,461.6910) --
(815.8360,460.8520) -- (815.8360,458.3520) -- (816.6680,456.6910) --
(816.6680,453.3520) -- (817.5000,451.6910) -- (817.5000,449.1910) --
(818.3360,448.3520) -- (818.3360,445.8520) -- (819.1680,444.1910) --
(819.1680,441.6910) -- (820.0000,440.8520) -- (820.0000,438.3520) --
(820.8360,437.5200) -- (820.8360,435.0200) -- (821.6680,433.3520) --
(821.6680,430.0200) -- (822.5000,428.3520) -- (822.5000,425.8520) --
(823.3360,425.0200) -- (823.3360,422.5200) -- (824.1680,420.8520) --
(824.1680,418.3520) -- (825.0000,417.5200) -- (825.0000,415.0200) --
(825.8360,413.3520) -- (825.8360,411.6910) -- (826.6680,410.0200) --
(826.6680,406.6910) -- (827.5000,405.0200) -- (827.5000,402.5200) --
(828.3360,401.6910) -- (828.3360,399.1910) -- (829.1680,397.5200) --
(829.1680,395.0200) -- (830.0000,394.1910) -- (830.0000,391.6910) --
(830.8360,390.0200) -- (830.8360,387.5200) -- (831.6680,386.6910) --
(831.6680,384.1910) -- (832.5000,383.3520) -- (832.5000,379.1910) --
(833.3360,378.3520) -- (833.3360,375.8520) -- (834.1680,374.1910) --
(834.1680,371.6910) -- (835.0000,370.8520) -- (835.0000,368.3520) --
(835.8360,366.6910) -- (835.8360,364.1910) -- (836.6680,363.3520) --
(836.6680,360.8520) -- (837.5000,359.1910) -- (837.5000,356.6910) --
(838.3360,355.8520) -- (838.3360,352.5200) -- (839.1680,350.8520) --
(839.1680,348.3520) -- (840.0000,347.5200) -- (840.0000,345.0200) --
(840.8360,343.3520) -- (840.8360,340.8520) -- (841.6680,340.0200) --
(841.6680,337.5200) -- (842.5000,335.8520) -- (842.5000,333.3520) --
(843.3360,332.5200) -- (843.3360,330.0200) -- (844.1680,328.3520) --
(844.1680,325.0200) -- (845.0000,324.1910) -- (845.0000,321.6910) --
(845.8360,320.0200) -- (845.8360,317.5200) -- (846.6680,316.6910) --
(846.6680,314.1910) -- (847.5000,312.5200) -- (847.5000,310.0200) --
(848.3360,309.1910) -- (848.3360,306.6910) -- (849.1680,305.0200) --
(849.1680,302.5200) -- (850.0000,301.6910) -- (850.0000,299.1910) --
(850.8360,297.5200) -- (850.8360,296.6910) -- (851.6680,296.6910) --
(851.6680,300.0200) -- (852.5000,301.6910) -- (852.5000,304.1910) --
(853.3360,305.0200) -- (853.3360,307.5200) -- (854.1680,309.1910) --
(854.1680,311.6910) -- (855.0000,312.5200) -- (855.0000,315.0200) --
(855.8360,316.6910) -- (855.8360,319.1910) -- (856.6680,320.0200) --
(856.6680,322.5200) -- (857.5000,324.1910) -- (857.5000,325.8520) --
(858.3360,327.5200) -- (858.3360,330.0200) -- (859.1680,330.8520) --
(859.1680,335.0200) -- (860.0000,335.8520) -- (860.0000,338.3520) --
(860.8360,340.0200) -- (860.8360,342.5200) -- (861.6680,343.3520) --
(861.6680,345.8520) -- (862.5000,347.5200) -- (862.5000,350.0200) --
(863.3360,350.8520) -- (863.3360,353.3520) -- (864.1680,354.1910) --
(864.1680,356.6910) -- (865.0000,358.3520) -- (865.0000,360.8520) --
(865.8360,361.6910) -- (865.8360,364.1910) -- (866.6680,365.8520) --
(866.6680,368.3520) -- (867.5000,369.1910) -- (867.5000,371.6910) --
(868.3360,373.3520) -- (868.3360,376.6910) -- (869.1680,378.3520) --
(869.1680,380.8520) -- (870.0000,381.6910) -- (870.0000,384.1910) --
(870.8360,385.0200) -- (870.8360,387.5200) -- (871.6680,389.1910) --
(871.6680,391.6910) -- (872.5000,392.5200) -- (872.5000,395.0200) --
(873.3360,396.6910) -- (873.3360,399.1910) -- (874.1680,400.0200) --
(874.1680,402.5200) -- (875.0040,404.1910) -- (875.0040,406.6910) --
(875.8360,407.5200) -- (875.8360,410.0200) -- (876.6680,411.6910) --
(876.6680,413.3520) -- (877.5040,415.0200) -- (877.5040,417.5200) --
(878.3360,418.3520) -- (878.3360,420.8520) -- (879.1680,422.5200) --
(879.1680,423.3520) -- (879.1680,425.0200) --
(880.0000,425.8520) -- (880.0000,430.0200) -- (880.8320,430.8520) --
(880.8320,433.3520) -- (881.6680,435.0200) -- (881.6680,437.5200) --
(882.5000,438.3520) -- (882.5000,440.8520) -- (883.3360,441.6910) --
(883.3360,444.1910) -- (884.1680,445.8520) -- (884.1680,448.3520) --
(885.0000,449.1910) -- (885.0000,451.6910) -- (885.8360,453.3520) --
(885.8360,455.8520) -- (886.6680,456.6910) -- (886.6680,459.1910) --
(887.5000,460.8520) -- (887.5000,463.3520) -- (888.3360,464.1910) --
(888.3360,466.6910) -- (889.1680,468.3520) -- (889.1680,470.8520) --
(890.0000,471.6910) -- (890.0000,474.1910) -- (890.8360,475.0200) --
(890.8360,477.5200) -- (891.6680,479.1910) -- (891.6680,481.6910) --
(892.5000,482.5200) -- (892.5000,485.0200) -- (893.3360,486.6910) --
(893.3360,489.1910) -- (894.1680,490.0200) -- (894.1680,492.5200) --
(895.0000,494.1910) -- (895.0000,497.5200) -- (895.8360,499.1910) --
(895.8360,500.8520) -- (896.6680,502.5200) -- (896.6680,505.0200) --
(897.5000,505.8520) -- (897.5000,508.3520) -- (898.3360,510.0200) --
(898.3360,512.5200) -- (899.1680,513.3520) -- (899.1680,515.8520) --
(900.0000,517.5200) -- (900.0000,520.0200) -- (900.8360,520.8520) --
(900.8360,523.3520) -- (901.6680,525.0200) -- (901.6680,527.5200) --
(902.5000,528.3520) -- (902.5000,530.8520) -- (903.3360,531.6910) --
(903.3360,534.1910) -- (904.1680,535.8520) -- (904.1680,538.3520) --
(905.0000,539.1910) -- (905.0000,541.6910) -- (905.8360,543.3520) --
(905.8360,545.8520) -- (906.6680,546.6910) -- (906.6680,549.1910) --
(907.5000,550.8520) -- (907.5000,553.3520) -- (908.3360,554.1910) --
(908.3360,556.6910) -- (909.1680,558.3520) -- (909.1680,560.0200) --
(910.0000,561.6910) -- (910.0000,564.1910) -- (910.8360,565.0200) --
(910.8360,567.5200) -- (911.6680,569.1910) -- (911.6680,571.6910) --
(912.5000,572.5200) -- (912.5000,575.0200) -- (913.3360,576.6910) --
(913.3360,579.1910) -- (914.1680,580.0200) -- (914.1680,582.5200) --
(915.0000,584.1910) -- (915.0000,586.6880) -- (915.8360,587.5200) --
(915.8360,590.0200) -- (916.6680,590.8520) -- (916.6680,593.3520) --
(917.5000,595.0200) -- (917.5000,597.5200) -- (918.3360,598.3520) --
(918.3360,600.8520) -- (919.1680,602.5200) -- (919.1680,605.0200) --
(920.0000,605.8520) -- (920.0000,608.3520) -- (920.8360,610.0200) --
(920.8360,612.5200) -- (921.6680,613.3520) -- (921.6680,615.8520) --
(922.5000,617.5200) -- (922.5000,619.1880) -- (923.3360,620.8520) --
(923.3360,623.3520) -- (924.1680,624.1880) -- (924.1680,626.6880) --
(925.0000,628.3520) -- (925.0000,630.8520) -- (925.8360,631.6880) --
(925.8360,634.1880) -- (926.6680,635.8520) -- (926.6680,638.3520) --
(927.5000,639.1880) -- (927.5000,641.6880) -- (928.3360,643.3520) --
(928.3360,645.8520) -- (929.1680,646.6880) -- (929.1680,649.1880) --
(930.0000,650.0200) -- (930.0000,652.5200) -- (930.8360,654.1880) --
(930.8360,656.6880) -- (931.6680,657.5200) -- (931.6680,660.0200) --
(932.5000,661.6880) -- (932.5000,664.1880) -- (933.3360,665.0200) --
(933.3360,667.5200) -- (934.1680,669.1880) -- (934.1680,671.6880) --
(935.0000,672.5200) -- (935.0000,675.0200) -- (935.8360,675.8520) --
(935.8360,678.3520) -- (936.6680,680.0200) -- (936.6680,682.5200) --
(937.5000,683.3520) -- (937.5000,685.8520) -- (938.3360,687.5200) --
(938.3360,690.0200) -- (939.1680,690.8520) -- (939.1680,693.3520) --
(940.0000,695.0200) -- (940.0000,697.5200) -- (940.8360,698.3520) --
(940.8360,700.8520) -- (941.6680,702.5200) -- (941.6680,705.0200) --
(942.5000,705.8520) -- (942.5000,708.3520) -- (943.3360,709.1880) --
(943.3360,711.6880) -- (944.1680,713.3520) -- (944.1680,715.8520) --
(945.0000,716.6880) -- (945.0000,719.1880) -- (945.8360,720.8520) --
(945.8360,723.3520) -- (946.6680,724.1880) -- (946.6680,725.8520) --
(947.5000,726.6880) -- (947.5000,729.1880) -- (948.3360,730.8520) --
(948.3360,733.3520) -- (949.1680,734.1880) -- (949.1680,736.6880) --
(950.0000,737.5200) -- (950.0000,740.0200) -- (950.8360,741.6880) --
(950.8360,744.1880) -- (951.6680,745.0200) -- (951.6680,747.5200) --
(952.5040,749.1880) -- (952.5040,751.6880) -- (953.3360,752.5200) --
(953.3360,755.0200) -- (954.1680,756.6880) -- (954.1680,759.1880) --
(955.0040,760.0200) -- (955.0040,762.5200) -- (955.8360,763.3520) --
(955.8360,765.8520) -- (956.6680,767.5200) -- (956.6680,770.0200) --
(957.5040,770.8520) -- (957.5040,773.3520) -- (958.3360,775.0200) --
(958.3360,777.5200) -- (959.1680,778.3520) -- (959.1680,780.8520) --
(960.0040,782.5200) -- (960.0040,785.0200) -- (960.8360,785.8520) --
(960.8360,788.3520) -- (961.6680,790.0200) --
(961.6680,792.5200) -- (962.5000,793.3520) --
(962.5000,794.1880) -- (963.3320,795.8520) -- (963.3320,798.3520) --
(964.1680,799.1880) -- (964.1680,801.6880) -- (965.0000,803.3520) --
(965.0000,805.8520) -- (965.8360,806.6880) -- (965.8360,809.1880) --
(966.6680,810.8520) -- (966.6680,813.3520) -- (967.5000,814.1880) --
(967.5000,816.6880) -- (968.3360,818.3520) -- (968.3360,820.8520) --
(969.1680,821.6880) -- (969.1680,824.1880) -- (970.0000,825.0200) --
(970.0000,827.5200) -- (970.8360,829.1880) -- (970.8360,831.6880) --
(971.6680,832.5200) -- (971.6680,835.0200) -- (972.5000,836.6880) --
(972.5000,839.1880) -- (973.3360,840.0200) -- (973.3360,841.6880) --
(974.1680,842.5200) -- (974.1680,845.0200) -- (975.0000,846.6880) --
(975.0000,849.1880) -- (975.8360,850.0200) -- (975.8360,852.5200) --
(976.6680,853.3520) -- (976.6680,855.8520) -- (977.5000,857.5200) --
(977.5000,860.0200) -- (978.3360,860.8520) -- (978.3360,863.3520) --
(979.1680,865.0200) -- (979.1680,867.5200) -- (980.0000,868.3520) --
(980.0000,870.8520) -- (980.8360,872.5200) -- (980.8360,875.0200) --
(981.6680,875.8520) -- (981.6680,878.3520) -- (982.5000,880.0200) --
(982.5000,880.8520) -- (983.3360,881.6880) -- (983.3360,884.1880) --
(984.1680,885.8520) -- (984.1680,888.3520) -- (985.0000,889.1880) --
(985.0000,891.6880) -- (985.8360,893.3520) -- (985.8360,895.8520) --
(986.6680,896.6880) -- (986.6680,899.1880) -- (987.5000,900.8520) --
(987.5000,903.3520) -- (988.3360,904.1880) -- (988.3360,906.6880) --
(989.1680,908.3520) -- (989.1680,910.0200) -- (990.0000,911.6880) --
(990.0000,914.1880) -- (990.8360,915.0200) -- (990.8360,916.6880) --
(991.6680,917.5200) -- (991.6680,920.0200) -- (992.5000,921.6880) --
(992.5000,924.1880) -- (993.3360,925.0200) -- (993.3360,927.5200) --
(994.1680,929.1880) -- (994.1680,931.6880) -- (995.0000,932.5200) --
(995.0000,935.0200) -- (995.8360,936.6880) -- (995.8360,939.1880) --
(996.6680,940.0200) -- (996.6680,942.5200) -- (997.5000,943.3520) --
(997.5000,945.8520) -- (998.3360,947.5200) -- (998.3360,948.3520) --
(999.1680,950.0200) -- (999.1680,952.5200) -- (1000.0000,953.3520) --
(1000.0000,955.8520) -- (1000.8300,957.5200) -- (1000.8300,960.0200) --
(1001.6700,960.8520) -- (1001.6700,963.3520) -- (1002.5000,965.0200) --
(1002.5000,967.5200) -- (1003.3300,968.3520) -- (1003.3300,970.8520) --
(1004.1700,971.6880) -- (1004.1700,974.1880) -- (1005.0000,975.8520) --
(1005.0000,976.6880) -- (1005.8300,978.3520) -- (1005.8300,980.8520) --
(1006.6700,981.6880) -- (1006.6700,984.1880) -- (1007.5000,985.8520) --
(1007.5000,988.3520) -- (1008.3300,989.1880) -- (1008.3300,991.6880) --
(1009.1700,993.3520) -- (1009.1700,995.8520) -- (1010.0000,996.6880) --
(1010.0000,999.1880) -- (1010.8300,1000.0200) -- (1010.8300,1001.6900) --
(1011.6700,1002.5200) -- (1011.6700,1005.0200) -- (1012.5000,1006.6900) --
(1012.5000,1009.1900) -- (1013.3300,1010.0200) -- (1013.3300,1012.5200) --
(1014.1700,1014.1900) -- (1014.1700,1016.6900) -- (1015.0000,1017.5200) --
(1015.0000,1020.0200) -- (1015.8300,1021.6900) -- (1015.8300,1024.1900) --
(1016.6700,1025.0200) -- (1016.6700,1026.6900) -- (1017.5000,1027.5200) --
(1017.5000,1030.0200) -- (1018.3300,1030.8500) -- (1018.3300,1033.3500) --
(1019.1700,1035.0200) -- (1019.1700,1037.5200) -- (1020.0000,1038.3500) --
(1020.0000,1040.8500) -- (1020.8300,1042.5200) -- (1020.8300,1045.0200) --
(1021.6700,1045.8500) -- (1021.6700,1047.5200) -- (1022.5000,1048.3500) --
(1022.5000,1050.8500) -- (1023.3300,1052.5200) -- (1023.3300,1055.0200) --
(1024.1700,1055.8500) -- (1024.1700,1058.3500) -- (1025.0000,1059.1900) --
(1025.0000,1061.6900) -- (1025.8300,1063.3500) -- (1025.8300,1065.8500) --
(1026.6700,1066.6900) -- (1026.6700,1069.1900) -- (1027.5000,1070.8500) --
(1027.5000,1071.6900) -- (1028.3300,1073.3500) -- (1028.3300,1075.8500) --
(1029.1700,1076.6900) -- (1029.1700,1079.1900) -- (1030.0000,1080.8500) --
(1030.0000,1083.3500) -- (1030.8300,1084.1900) -- (1030.8300,1086.6900) --
(1031.6700,1087.5200) -- (1031.6700,1089.1900) -- (1032.5000,1090.0200) --
(1032.5000,1092.5200) -- (1033.3300,1094.1900) -- (1033.3300,1096.6900) --
(1034.1700,1097.5200) -- (1034.1700,1100.0200) -- (1035.0000,1101.6900) --
(1035.0000,1104.1900) -- (1035.8300,1105.0200) -- (1035.8300,1107.5200) --
(1036.6700,1109.1900) -- (1036.6700,1110.0200) -- (1037.5000,1111.6900) --
(1037.5000,1114.1900) -- (1038.3300,1115.0200) -- (1038.3300,1117.5200) --
(1039.1700,1118.3500) -- (1039.1700,1120.8500) -- (1040.0000,1122.5200) --
(1040.0000,1125.0200) -- (1040.8300,1125.8500) -- (1040.8300,1128.3500) --
(1041.6700,1130.0200) -- (1041.6700,1130.8500) -- (1042.5000,1132.5200) --
(1042.5000,1135.0200) -- (1043.3300,1135.8500) -- (1043.3300,1138.3500) --
(1044.1700,1140.0200) -- (1044.1700,1142.5200) -- (1045.0000,1143.3500) --
(1045.0000,1145.8500) -- (1045.8400,1146.6900) -- (1045.8400,1148.3500) --
(1046.6700,1149.1900) -- (1046.6700,1151.6900) -- (1047.5000,1153.3500) --
(1047.5000,1155.8500) -- (1048.3400,1156.6900) -- (1048.3400,1159.1900) --
(1049.1700,1160.8500) -- (1049.1700,1163.3500) --
(1050.0000,1164.1900) -- (1050.0000,1165.8500) -- (1050.8300,1166.6900) --
(1050.8300,1169.1900) -- (1051.6700,1170.8500) -- (1051.6700,1173.3500) --
(1052.5000,1174.1900) -- (1052.5000,1176.6900) -- (1053.3300,1177.5200) --
(1053.3300,1180.0200) -- (1054.1700,1181.6900) -- (1054.1700,1182.5200) --
(1055.0000,1184.1900) -- (1055.0000,1186.6900) -- (1055.8300,1187.5200) --
(1055.8300,1190.0200) -- (1056.6700,1191.6900) -- (1056.6700,1194.1900) --
(1057.5000,1195.0200) -- (1057.5000,1197.5200) -- (1058.3300,1199.1900) --
(1058.3300,1200.0200) -- (1059.1700,1201.6900) -- (1059.1700,1203.3500) --
(1060.0000,1205.0200) -- (1060.0000,1207.5200) -- (1060.8300,1208.3500) --
(1060.8300,1210.8500) -- (1061.6700,1212.5200) -- (1061.6700,1213.3600) --
(1062.5000,1215.0200) -- (1062.5000,1217.5200) -- (1063.3300,1218.3600) --
(1063.3300,1220.8600) -- (1064.1700,1222.5200) -- (1064.1700,1225.0200) --
(1065.0000,1225.8600) -- (1065.0000,1228.3600) -- (1065.8300,1230.0200) --
(1065.8300,1230.8600) -- (1066.6700,1231.6900) -- (1066.6700,1234.1900) --
(1067.5000,1235.8600) -- (1067.5000,1238.3600) -- (1068.3300,1239.1900) --
(1068.3300,1241.6900) -- (1069.1700,1243.3600) -- (1069.1700,1244.1900) --
(1070.0000,1245.8600) -- (1070.0000,1248.3600) -- (1070.8300,1249.1900) --
(1070.8300,1251.6900) -- (1071.6700,1253.3600) -- (1071.6700,1255.8600) --
(1072.5000,1256.6900) -- (1072.5000,1259.1900) -- (1073.3300,1260.8600) --
(1073.3300,1261.6900) -- (1074.1700,1262.5200) -- (1074.1700,1265.0200) --
(1075.0000,1266.6900) -- (1075.0000,1269.1900) -- (1075.8300,1270.0200) --
(1075.8300,1272.5200) -- (1076.6700,1274.1900) -- (1076.6700,1275.0200) --
(1077.5000,1276.6900) -- (1077.5000,1279.1900) -- (1078.3300,1280.0200) --
(1078.3300,1282.5200) -- (1079.1700,1284.1900) -- (1079.1700,1286.6900) --
(1080.0000,1287.5200) -- (1080.0000,1289.1900) -- (1080.8300,1290.0200) --
(1080.8300,1292.5200) -- (1081.6700,1293.3600) -- (1081.6700,1295.8600) --
(1082.5000,1297.5200) -- (1082.5000,1300.0200) -- (1083.3300,1300.8600) --
(1083.3300,1302.5200) -- (1084.1700,1303.3600) -- (1084.1700,1305.8600) --
(1085.0000,1307.5200) -- (1085.0000,1310.0200) -- (1085.8300,1310.8600) --
(1085.8300,1313.3600) -- (1086.6700,1315.0200) -- (1086.6700,1315.8600) --
(1087.5000,1317.5200) -- (1087.5000,1320.0200) -- (1088.3300,1320.8600) --
(1088.3300,1323.3600) -- (1089.1700,1324.1900) -- (1089.1700,1326.6900) --
(1090.0000,1328.3600) -- (1090.0000,1329.1900) -- (1090.8300,1330.8600) --
(1090.8300,1333.3600) -- (1091.6700,1334.1900) -- (1091.6700,1336.6900) --
(1092.5000,1338.3600) -- (1092.5000,1340.8600) -- (1093.3300,1341.6900) --
(1093.3300,1343.3600) -- (1094.1700,1344.1900) -- (1094.1700,1346.6900) --
(1095.0000,1348.3600) -- (1095.0000,1350.0200) -- (1095.8300,1351.6900) --
(1095.8300,1354.1900) -- (1096.6700,1355.0200) -- (1096.6700,1356.6900) --
(1097.5000,1357.5200) -- (1097.5000,1360.0200) -- (1098.3300,1361.6900) --
(1098.3300,1364.1900) -- (1099.1700,1365.0200) -- (1099.1700,1366.6900) --
(1100.0000,1367.5200) -- (1100.0000,1370.0200) -- (1100.8300,1371.6900) --
(1100.8300,1374.1900) -- (1101.6700,1375.0200) -- (1101.6700,1377.5200) --
(1102.5000,1378.3600) -- (1102.5000,1380.0200) -- (1103.3300,1380.8600) --
(1103.3300,1383.3600) -- (1104.1700,1385.0200) -- (1104.1700,1387.5200) --
(1105.0000,1388.3600) -- (1105.0000,1390.8600) -- (1105.8300,1392.5200) --
(1105.8300,1393.3600) -- (1106.6700,1395.0200) -- (1106.6700,1397.5200) --
(1107.5000,1398.3600) -- (1107.5000,1400.8600) -- (1108.3300,1402.5200) --
(1108.3300,1403.3600) -- (1109.1700,1405.0200) -- (1109.1700,1407.5200) --
(1110.0000,1408.3600) -- (1110.0000,1410.8600) -- (1110.8300,1411.6900) --
(1110.8300,1414.1900) -- (1111.6700,1415.8600) -- (1111.6700,1416.6900) --
(1112.5000,1418.3600) -- (1112.5000,1420.8600) -- (1113.3300,1421.6900) --
(1113.3300,1424.1900) -- (1114.1700,1425.8600) -- (1114.1700,1426.6900) --
(1115.0000,1428.3600) -- (1115.0000,1430.8600) -- (1115.8300,1431.6900) --
(1115.8300,1434.1900) -- (1116.6700,1435.8600) -- (1116.6700,1436.6900) --
(1117.5000,1437.5200) -- (1117.5000,1440.0200) -- (1118.3300,1441.6900) --
(1118.3300,1444.1900) -- (1119.1700,1445.0200) -- (1119.1700,1447.5200) --
(1120.0000,1449.1900) -- (1120.0000,1450.0200) -- (1120.8300,1451.6900) --
(1120.8300,1454.1900) -- (1121.6700,1455.0200) -- (1121.6700,1457.5200) --
(1122.5000,1459.1900) -- (1122.5000,1460.0200) -- (1123.3300,1461.6900) --
(1123.3300,1464.1900) -- (1124.1700,1465.0200) -- (1124.1700,1467.5200) --
(1125.0000,1468.3600) -- (1125.0000,1470.0200) -- (1125.8300,1470.8600) --
(1125.8300,1473.3600) -- (1126.6700,1475.0200) -- (1126.6700,1477.5200) --
(1127.5000,1478.3600) -- (1127.5000,1480.8600) -- (1128.3300,1482.5200) --
(1128.3300,1483.3600) -- (1129.1700,1485.0200) -- (1129.1700,1487.5200) --
(1130.0000,1488.3600) -- (1130.0000,1490.8600) -- (1130.8400,1492.5200) --
(1130.8400,1493.3600) -- (1131.6700,1495.0200) -- (1131.6700,1496.6900) --
(1132.5000,1498.3600) -- (1132.5000,1500.8600) -- (1133.3400,1501.6900) --
(1133.3400,1503.3600) -- (1134.1700,1504.1900) -- (1134.1700,1506.6900) --
(1135.0000,1508.3600) -- (1135.0000,1510.8600) -- (1135.8400,1511.6900) --
(1135.8400,1513.3600) -- (1136.6700,1514.1900) -- (1136.6700,1516.6900) --
(1137.5000,1518.3600) -- (1137.5000,1520.8600) -- (1138.3400,1521.6900) --
(1138.3400,1523.3600) -- (1139.1700,1524.1900) -- (1139.1700,1526.6900) --
(1140.0000,1527.5200) -- (600.8360,1527.5200)"""
data_2colon3 = """
(1177.5000,1527.5200) -- (1177.5000,1526.6900)
-- (1178.3300,1525.0200) -- (1178.3300,1524.1900) -- (1179.1700,1523.3500) --
(1179.1700,1521.6900) -- (1180.0000,1520.8500) -- (1180.0000,1519.1900) --
(1180.8300,1518.3500) -- (1181.6700,1516.6900) -- (1181.6700,1515.8500) --
(1182.5000,1514.1900) -- (1182.5000,1513.3500) -- (1183.3300,1511.6900) --
(1183.3300,1510.8500) -- (1184.1700,1509.1900) -- (1184.1700,1508.3500) --
(1185.0000,1506.6900) -- (1185.0000,1505.8500) -- (1185.8300,1504.1900) --
(1185.8300,1503.3500) -- (1186.6700,1501.6900) -- (1186.6700,1500.8500) --
(1187.5000,1499.1900) -- (1187.5000,1498.3500) -- (1188.3300,1496.6900) --
(1189.1700,1495.8500) -- (1189.1700,1495.0200) -- (1190.0000,1493.3500) --
(1190.0000,1492.5200) -- (1190.8300,1490.8500) -- (1190.8300,1490.0200) --
(1191.6700,1488.3500) -- (1191.6700,1487.5200) -- (1192.5000,1485.8500) --
(1192.5000,1485.0200) -- (1193.3300,1483.3500) -- (1193.3300,1482.5200) --
(1194.1700,1480.8500) -- (1194.1700,1480.0200) -- (1195.0000,1478.3500) --
(1195.0000,1477.5200) -- (1195.8300,1475.8500) -- (1195.8300,1475.0200) --
(1196.6700,1473.3500) -- (1196.6700,1472.5200) -- (1197.5000,1470.8500) --
(1197.5000,1470.0200) -- (1198.3300,1468.3500) -- (1198.3300,1467.5200) --
(1199.1700,1465.8500) -- (1199.1700,1465.0200) -- (1200.0000,1464.1800) --
(1200.0000,1462.5200) -- (1200.8300,1461.6800) -- (1200.8300,1460.0200) --
(1201.6700,1459.1800) -- (1201.6700,1457.5200) -- (1202.5000,1456.6800) --
(1202.5000,1455.0200) -- (1203.3300,1454.1800) -- (1203.3300,1452.5200) --
(1204.1700,1451.6800) -- (1204.1700,1450.0200) -- (1205.0000,1449.1800) --
(1205.0000,1447.5200) -- (1205.8300,1446.6800) -- (1205.8300,1445.0200) --
(1206.6700,1444.1800) -- (1206.6700,1442.5200) -- (1207.5000,1441.6800) --
(1207.5000,1440.0200) -- (1208.3300,1439.1800) -- (1208.3300,1437.5200) --
(1209.1700,1436.6800) -- (1209.1700,1435.8500) -- (1210.0000,1434.1800) --
(1210.0000,1433.3500) -- (1210.8300,1431.6800) -- (1210.8300,1430.8500) --
(1211.6700,1429.1800) -- (1211.6700,1428.3500) -- (1212.5000,1426.6800) --
(1212.5000,1425.8500) -- (1213.3300,1424.1800) -- (1213.3300,1423.3500) --
(1214.1700,1421.6800) -- (1214.1700,1420.8500) -- (1215.0000,1419.1800) --
(1215.0000,1418.3500) -- (1215.8300,1416.6800) -- (1215.8300,1415.8500) --
(1216.6700,1414.1800) -- (1216.6700,1413.3500) -- (1217.5000,1411.6800) --
(1217.5000,1409.1800) -- (1218.3300,1408.3500) -- (1218.3300,1407.5200) --
(1219.1700,1405.8500) -- (1219.1700,1405.0200) -- (1220.0000,1403.3500) --
(1220.0000,1402.5200) -- (1220.8300,1400.8500) -- (1220.8300,1400.0200) --
(1221.6700,1398.3500) -- (1221.6700,1397.5200) -- (1222.5000,1395.8500) --
(1222.5000,1395.0200) -- (1223.3300,1393.3500) -- (1223.3300,1392.5200) --
(1224.1700,1390.8500) -- (1224.1700,1388.3500) -- (1225.0000,1387.5200) --
(1225.0000,1385.8500) -- (1225.8300,1385.0200) -- (1225.8300,1383.3500) --
(1226.6700,1382.5200) -- (1226.6700,1380.8500) -- (1227.5000,1380.0200) --
(1227.5000,1378.3500) -- (1228.3300,1377.5200) -- (1228.3300,1376.6800) --
(1229.1700,1375.0200) -- (1229.1700,1372.5200) -- (1230.0000,1371.6800) --
(1230.0000,1370.0200) -- (1230.8300,1369.1800) -- (1230.8300,1367.5200) --
(1231.6700,1366.6800) -- (1231.6700,1365.0200) -- (1232.5000,1364.1800) --
(1232.5000,1362.5200) -- (1233.3300,1361.6800) -- (1233.3300,1359.1800) --
(1234.1700,1357.5200) -- (1234.1700,1356.6800) -- (1235.0000,1355.0200) --
(1235.0000,1354.1800) -- (1235.8300,1352.5200) -- (1235.8300,1351.6800) --
(1236.6700,1350.0200) -- (1236.6700,1348.3500) -- (1237.5000,1346.6800) --
(1237.5000,1345.8500) -- (1238.3300,1344.1800) -- (1238.3300,1343.3500) --
(1239.1700,1341.6800) -- (1239.1700,1340.8500) -- (1240.0000,1339.1800) --
(1240.0000,1336.6800) -- (1240.8300,1335.8500) -- (1240.8300,1334.1800) --
(1241.6700,1333.3500) -- (1241.6700,1331.6800) -- (1242.5000,1330.8500) --
(1242.5000,1329.1800) -- (1243.3300,1328.3500) -- (1243.3300,1325.8500) --
(1244.1700,1324.1800) -- (1244.1700,1323.3500) -- (1245.0000,1321.6800) --
(1245.0000,1320.8500) -- (1245.8300,1320.0200) -- (1245.8300,1317.5200) --
(1246.6700,1315.8500) -- (1246.6700,1315.0200) -- (1247.5000,1313.3500) --
(1247.5000,1312.5200) -- (1248.3300,1310.8500) -- (1248.3300,1308.3500) --
(1249.1700,1307.5200) -- (1249.1700,1305.8500) -- (1250.0000,1305.0200) --
(1250.0000,1303.3500) -- (1250.8300,1302.5200) -- (1250.8300,1300.0200) --
(1251.6700,1298.3500) -- (1251.6700,1297.5200) -- (1252.5000,1295.8500) --
(1252.5000,1293.3500) -- (1253.3300,1292.5200) -- (1253.3300,1290.8500) --
(1254.1700,1290.0200) -- (1254.1700,1289.1800) -- (1255.0000,1287.5200) --
(1255.0000,1285.0200) -- (1255.8300,1284.1800) -- (1255.8300,1282.5200) --
(1256.6700,1281.6800) -- (1256.6700,1279.1800) -- (1257.5000,1277.5200) --
(1257.5000,1276.6800) -- (1258.3300,1275.0200) -- (1258.3300,1272.5200) --
(1259.1700,1271.6800) -- (1259.1700,1270.0200) -- (1260.0000,1269.1800) --
(1260.0000,1267.5200) -- (1260.8400,1266.6800) -- (1260.8400,1264.1800) --
(1261.6700,1262.5200) -- (1261.6700,1261.6800) -- (1262.5000,1260.8500) --
(1262.5000,1258.3500) -- (1263.3400,1256.6800) -- (1263.3400,1255.8500) --
(1264.1700,1254.1800) -- (1264.1700,1251.6800) -- (1265.0000,1250.8500) --
(1265.0000,1249.1800) -- (1265.8400,1248.3500) -- (1265.8400,1245.8500) --
(1266.6700,1244.1800) -- (1266.6700,1241.6800) -- (1267.5000,1240.8500) --
(1267.5000,1239.1800) -- (1268.3400,1238.3500) -- (1268.3400,1235.8500) --
(1269.1700,1234.1800) -- (1269.1700,1233.3500) -- (1270.0000,1231.6800) --
(1270.0000,1230.0200) -- (1270.8400,1228.3500) -- (1270.8400,1227.5200) --
(1271.6700,1225.8500) -- (1271.6700,1223.3500) -- (1272.5000,1222.5200) --
(1272.5000,1220.0200) -- (1273.3400,1218.3500) -- (1273.3400,1217.5200) --
(1274.1700,1215.8500) -- (1274.1700,1213.3500) -- (1275.0000,1212.5200) --
(1275.0000,1210.8500) -- (1275.8400,1210.0200) -- (1275.8400,1207.5200) --
(1276.6700,1205.8500) -- (1276.6700,1203.3500) -- (1277.5000,1202.5200) --
(1277.5000,1201.6800) -- (1278.3400,1200.0200) -- (1278.3400,1197.5200) --
(1279.1700,1196.6800) -- (1279.1700,1194.1800) -- (1280.0000,1192.5200) --
(1280.0000,1191.6800) -- (1280.8400,1190.0200) -- (1280.8400,1187.5200) --
(1281.6700,1186.6800) -- (1281.6700,1184.1800) -- (1282.5000,1182.5200) --
(1282.5000,1180.0200) -- (1283.3400,1179.1800) -- (1283.3400,1177.5200) --
(1284.1700,1176.6800) -- (1284.1700,1174.1800) -- (1285.0000,1173.3500) --
(1285.0000,1170.8500) -- (1285.8400,1169.1800) -- (1285.8400,1166.6800) --
(1286.6700,1165.8500) -- (1286.6700,1164.1800) -- (1287.5000,1163.3500) --
(1287.5000,1160.8500) -- (1288.3400,1159.1800) -- (1288.3400,1156.6800) --
(1289.1700,1155.8500) -- (1289.1700,1153.3500) -- (1290.0000,1151.6800) --
(1290.0000,1149.1800) -- (1290.8400,1148.3500) -- (1290.8400,1145.8500) --
(1291.6700,1144.1800) -- (1291.6700,1143.3500) -- (1292.5000,1142.5200) --
(1292.5000,1140.0200) -- (1293.3400,1138.3500) -- (1293.3400,1135.8500) --
(1294.1700,1135.0200) -- (1294.1700,1132.5200) -- (1295.0000,1130.8500) --
(1295.0000,1128.3500) -- (1295.8400,1127.5200) -- (1295.8400,1125.0200) --
(1296.6700,1123.3500) -- (1296.6700,1120.8500) -- (1297.5000,1120.0200) --
(1297.5000,1117.5200) -- (1298.3400,1115.8500) -- (1298.3400,1114.1800) --
(1299.1700,1112.5200) -- (1299.1700,1110.0200) -- (1300.0000,1109.1800) --
(1300.0000,1106.6800) -- (1300.8400,1105.0200) -- (1300.8400,1102.5200) --
(1301.6700,1101.6800) -- (1301.6700,1099.1800) -- (1302.5000,1097.5200) --
(1302.5000,1095.0200) -- (1303.3400,1094.1800) -- (1303.3400,1091.6800) --
(1304.1700,1090.0200) -- (1304.1700,1087.5200) -- (1305.0000,1086.6800) --
(1305.0000,1084.1800) -- (1305.8400,1083.3500) -- (1305.8400,1080.8500) --
(1306.6700,1079.1800) -- (1306.6700,1076.6800) -- (1307.5000,1075.8500) --
(1307.5000,1073.3500) -- (1308.3400,1071.6800) -- (1308.3400,1069.1800) --
(1309.1700,1068.3500) -- (1309.1700,1064.1800) -- (1310.0000,1063.3500) --
(1310.0000,1060.8500) -- (1310.8400,1059.1800) -- (1310.8400,1056.6800) --
(1311.6700,1055.8500) -- (1311.6700,1053.3500) -- (1312.5000,1052.5200) --
(1312.5000,1050.0200) -- (1313.3400,1048.3500) -- (1313.3400,1045.0200) --
(1314.1700,1043.3500) -- (1314.1700,1040.8500) -- (1315.0000,1040.0200) --
(1315.0000,1037.5200) -- (1315.8400,1035.8500) -- (1315.8400,1033.3500) --
(1316.6700,1032.5200) -- (1316.6700,1028.3500) -- (1317.5000,1027.5200) --
(1317.5000,1025.0200) -- (1318.3400,1024.1800) -- (1318.3400,1021.6800) --
(1319.1700,1020.0200) -- (1319.1700,1016.6800) -- (1320.0000,1015.0200) --
(1320.0000,1012.5200) -- (1320.8400,1011.6800) -- (1320.8400,1007.5200) --
(1321.6700,1006.6800) -- (1321.6700,1004.1800) -- (1322.5000,1002.5200) --
(1322.5000,999.1840) -- (1323.3400,997.5160) -- (1323.3400,995.8480) --
(1324.1700,994.1840) -- (1324.1700,990.8480) -- (1325.0000,989.1840) --
(1325.0000,986.6840) -- (1325.8400,985.8480) -- (1325.8400,981.6840) --
(1326.6700,980.8480) -- (1326.6700,978.3520) -- (1327.5000,976.6840) --
(1327.5000,973.3520) -- (1328.3400,971.6840) -- (1328.3400,969.1840) --
(1329.1700,968.3520) -- (1329.1700,965.0160) -- (1330.0000,963.3520) --
(1330.0000,960.0160) -- (1330.8400,958.3520) -- (1330.8400,955.8520) --
(1331.6700,955.0160) -- (1331.6700,950.8520) -- (1332.5000,950.0160) --
(1332.5000,945.8520) -- (1333.3400,945.0160) -- (1333.3400,940.8520) --
(1334.1700,940.0160) -- (1334.1700,937.5160) -- (1335.0000,936.6840) --
(1335.0000,932.5160) -- (1335.8400,931.6840) -- (1335.8400,927.5160) --
(1336.6700,926.6840) -- (1336.6700,922.5160) -- (1337.5000,921.6840) --
(1337.5000,917.5160) -- (1338.3400,916.6840) -- (1338.3400,912.5160) --
(1339.1700,911.6840) -- (1339.1700,908.3520) -- (1340.0000,906.6840) --
(1340.0000,903.3520) -- (1340.8400,901.6840) -- (1340.8400,898.3520) --
(1341.6700,896.6840) -- (1341.6700,893.3520) -- (1342.5000,891.6840) --
(1342.5000,888.3520) -- (1343.3400,886.6840) -- (1343.3400,883.3520) --
(1344.1700,881.6840) -- (1344.1700,878.3520) -- (1345.0000,877.5160) --
(1345.0000,873.3520) -- (1345.8400,872.5160) -- (1345.8400,867.5160) --
(1346.6700,865.8520) -- (1346.6700,862.5160) -- (1347.5000,860.8520) --
(1347.5000,857.5160) -- (1348.3400,855.8520) -- (1348.3400,851.6840) --
(1349.1700,850.0160) -- (1349.1700,846.6840) -- (1350.0000,845.0160) --
(1350.0000,841.6840) -- (1350.8400,840.0160) -- (1350.8400,835.0160) --
(1351.6700,834.1840) -- (1351.6700,830.0160) -- (1352.5000,829.1840) --
(1352.5000,824.1840) -- (1353.3400,822.5160) -- (1353.3400,818.3520) --
(1354.1700,816.6840) -- (1354.1700,813.3520) -- (1355.0000,811.6840) --
(1355.0000,806.6840) -- (1355.8400,805.8520) -- (1355.8400,800.8520) --
(1356.6700,799.1840) -- (1356.6700,794.1840) -- (1357.5000,793.3520) --
(1357.5000,788.3520) -- (1358.3400,787.5160) -- (1358.3400,782.5160) --
(1359.1700,780.8520) -- (1359.1700,775.8520) -- (1360.0000,775.0160) --
(1360.0000,770.0160) -- (1360.8400,768.3520) -- (1360.8400,763.3520) --
(1361.6700,762.5160) -- (1361.6700,757.5160) -- (1362.5000,756.6840) --
(1362.5000,751.6840) -- (1363.3400,750.0160) -- (1363.3400,744.1840) --
(1364.1700,742.5160) -- (1364.1700,737.5160) -- (1365.0000,736.6840) --
(1365.0000,730.8520) -- (1365.8400,729.1840) -- (1365.8400,724.1840) --
(1366.6700,723.3520) -- (1366.6700,716.6840) -- (1367.5000,715.8520) --
(1367.5000,709.1840) -- (1368.3400,708.3520) -- (1368.3400,702.5160) --
(1369.1700,700.8520) -- (1369.1700,695.0160) -- (1370.0000,693.3520) --
(1370.0000,687.5160) -- (1370.8400,685.8520) -- (1370.8400,680.0160) --
(1371.6700,678.3520) -- (1371.6700,672.5160) -- (1372.5000,671.6840) --
(1372.5000,664.1840) -- (1373.3400,662.5160) -- (1373.3400,655.0160) --
(1374.1700,654.1840) -- (1374.1700,647.5160) -- (1375.0000,646.6840) --
(1375.0000,639.1840) -- (1375.8400,638.3520) -- (1375.8400,630.8520) --
(1376.6700,629.1840) -- (1376.6700,620.8520) -- (1377.5000,619.1840) --
(1377.5000,612.5160) -- (1378.3400,610.8520) -- (1378.3400,602.5160) --
(1379.1700,600.8520) -- (1379.1700,592.5160) -- (1380.0000,590.8520) --
(1380.0000,582.5200) -- (1380.8400,581.6800) -- (1380.8400,571.6800) --
(1381.6700,570.0200) -- (1381.6700,560.0200) -- (1382.5000,559.1800) --
(1382.5000,549.1800) -- (1383.3400,548.3520) -- (1383.3400,536.6800) --
(1384.1700,535.8520) -- (1384.1700,525.0200) -- (1385.0000,523.3520) --
(1385.0000,510.8520) -- (1385.8400,510.0200) -- (1385.8400,496.6800) --
(1386.6700,495.0200) -- (1386.6700,481.6800) -- (1387.5000,480.0200) --
(1387.5000,464.1800) -- (1388.3400,463.3520) -- (1388.3400,445.8520) --
(1389.1700,444.1800) -- (1389.1700,425.0200) -- (1390.0000,423.3520) --
(1390.0000,399.1800) -- (1390.8400,397.5200) -- (1390.8400,364.1800) --
(1391.6700,363.3520) -- (1391.6700,296.6800) -- (1391.6700,328.3520) --
(1392.5000,330.0200) -- (1392.5000,379.1800) -- (1393.3400,380.8520) --
(1393.3400,409.1800) -- (1394.1700,410.0200) -- (1394.1700,432.5200) --
(1395.0000,433.3520) -- (1395.0000,451.6800) -- (1395.8400,453.3520) --
(1395.8400,470.8520) -- (1396.6700,471.6800) -- (1396.6700,486.6800) --
(1397.5000,487.5200) -- (1397.5000,500.8520) -- (1398.3400,502.5200) --
(1398.3400,515.0200) -- (1399.1700,515.8520) -- (1399.1700,528.3520) --
(1400.0000,529.1800) -- (1400.0000,540.8520) -- (1400.8400,541.6800) --
(1400.8400,551.6800) -- (1401.6700,553.3520) -- (1401.6700,562.5200) --
(1402.5000,564.1800) -- (1402.5000,574.1800) -- (1403.3400,575.0200) --
(1403.3400,584.1800) -- (1404.1700,585.0200) -- (1404.1700,593.3520) --
(1405.0000,595.0160) -- (1405.0000,603.3520) -- (1405.8400,605.0160) --
(1405.8400,613.3520) -- (1406.6700,615.0160) -- (1406.6700,621.6840) --
(1407.5000,623.3520) -- (1407.5000,630.8520) -- (1408.3400,631.6840) --
(1408.3400,639.1840) -- (1409.1700,640.8520) -- (1409.1700,647.5160) --
(1410.0000,649.1840) -- (1410.0000,656.6840) -- (1410.8400,657.5160) --
(1410.8400,664.1840) -- (1411.6700,665.0160) -- (1411.6700,671.6840) --
(1412.5000,672.5160) -- (1412.5000,680.0160) -- (1413.3400,680.8520) --
(1413.3400,687.5160) -- (1414.1700,688.3520) -- (1414.1700,695.0160) --
(1415.0000,695.8520) -- (1415.0000,702.5160) -- (1415.8400,703.3520) --
(1415.8400,708.3520) -- (1416.6700,709.1840) -- (1416.6700,715.8520) --
(1417.5000,716.6840) -- (1417.5000,723.3520) -- (1418.3400,724.1840) --
(1418.3400,729.1840) -- (1419.1700,730.8520) -- (1419.1700,735.0160) --
(1420.0000,736.6840) -- (1420.0000,742.5160) -- (1420.8400,744.1840) --
(1420.8400,749.1840) -- (1421.6700,750.0160) -- (1421.6700,755.0160) --
(1422.5000,756.6840) -- (1422.5000,761.6840) -- (1423.3400,762.5160) --
(1423.3400,767.5160) -- (1424.1700,768.3520) -- (1424.1700,773.3520) --
(1425.0000,775.0160) -- (1425.0000,780.0160) -- (1425.8400,780.8520) --
(1425.8400,785.8520) -- (1426.6700,787.5160) -- (1426.6700,792.5160) --
(1427.5100,793.3520) -- (1427.5100,796.6840) -- (1428.3400,798.3520) --
(1428.3400,803.3520) -- (1429.1700,804.1840) -- (1429.1700,808.3520) --
(1430.0100,809.1840) -- (1430.0100,814.1840) -- (1430.8400,815.8520) --
(1430.8400,819.1840) -- (1431.6700,820.8520) -- (1431.6700,825.0160) --
(1432.5100,826.6840) -- (1432.5100,830.0160) -- (1433.3400,831.6840) --
(1433.3400,836.6840) -- (1434.1700,837.5160) -- (1434.1700,841.6840) --
(1435.0100,842.5160) -- (1435.0100,846.6840) -- (1435.8400,847.5160) --
(1435.8400,851.6840) -- (1436.6700,852.5160) -- (1436.6700,855.8520) --
(1437.5100,857.5160) -- (1437.5100,862.5160) -- (1438.3400,863.3520) --
(1438.3400,867.5160) -- (1439.1700,868.3520) -- (1439.1700,872.5160) --
(1440.0100,873.3520) -- (1440.0100,877.5160) -- (1440.8400,878.3520) --
(1440.8400,881.6840) -- (1441.6700,883.3520) -- (1441.6700,886.6840) --
(1442.5100,888.3520) -- (1442.5100,890.8520) -- (1443.3400,891.6840) --
(1443.3400,895.8520) -- (1444.1700,896.6840) -- (1444.1700,900.8520) --
(1445.0100,901.6840) -- (1445.0100,905.8520) -- (1445.8400,906.6840) --
(1445.8400,910.0160) -- (1446.6700,911.6840) -- (1446.6700,914.1840) --
(1447.5100,915.0160) -- (1447.5100,919.1840) -- (1448.3400,920.0160) --
(1448.3400,924.1840) -- (1449.1700,925.0160) -- (1449.1700,927.5160) --
(1450.0100,929.1840) -- (1450.0100,932.5160) -- (1450.8400,934.1840) --
(1450.8400,937.5160) -- (1451.6700,939.1840) -- (1451.6700,940.8520) --
(1452.5100,942.5160) -- (1452.5100,945.8520) -- (1453.3400,947.5160) --
(1453.3400,950.0160) -- (1454.1700,950.8520) -- (1454.1700,955.0160) --
(1455.0100,955.8520) -- (1455.0100,958.3520) -- (1455.8400,960.0160) --
(1455.8400,962.5160) -- (1456.6700,963.3520) -- (1456.6700,967.5160) --
(1457.5100,968.3520) -- (1457.5100,970.8520) -- (1458.3400,971.6840) --
(1458.3400,975.8520) -- (1459.1700,976.6840) -- (1459.1700,979.1840) --
(1460.0100,980.8480) -- (1460.0100,983.3480) -- (1460.8400,984.1840) --
(1460.8400,988.3480) -- (1461.6700,989.1840) -- (1461.6700,991.6840) --
(1462.5100,993.3480) -- (1462.5100,995.8480) -- (1463.3400,996.6840) --
(1463.3400,999.1840) -- (1464.1700,1000.0200) -- (1464.1700,1002.5200) --
(1465.0100,1004.1800) -- (1465.0100,1007.5200) -- (1465.8400,1009.1800) --
(1465.8400,1011.6800) -- (1466.6700,1012.5200) -- (1466.6700,1015.0200) --
(1467.5100,1016.6800) -- (1467.5100,1019.1800) -- (1468.3400,1020.0200) --
(1468.3400,1022.5200) -- (1469.1700,1024.1800) -- (1469.1700,1026.6800) --
(1470.0100,1027.5200) -- (1470.0100,1030.0200) -- (1470.8400,1030.8500) --
(1470.8400,1033.3500) -- (1471.6700,1035.0200) -- (1471.6700,1037.5200) --
(1472.5100,1038.3500) -- (1472.5100,1040.8500) -- (1473.3400,1042.5200) --
(1473.3400,1045.0200) -- (1474.1700,1045.8500) -- (1474.1700,1048.3500) --
(1475.0100,1050.0200) -- (1475.0100,1052.5200) -- (1475.8400,1053.3500) --
(1475.8400,1055.8500) -- (1476.6700,1056.6800) -- (1476.6700,1059.1800) --
(1477.5100,1060.8500) -- (1477.5100,1063.3500) -- (1478.3400,1064.1800) --
(1478.3400,1066.6800) -- (1479.1700,1068.3500) -- (1479.1700,1070.8500) --
(1480.0100,1071.6800) -- (1480.0100,1074.1800) -- (1480.8400,1075.8500) --
(1480.8400,1078.3500) -- (1481.6700,1079.1800) -- (1481.6700,1080.8500) --
(1482.5100,1081.6800) -- (1482.5100,1084.1800) -- (1483.3400,1085.8500) --
(1483.3400,1087.5200) -- (1484.1700,1089.1800) -- (1484.1700,1091.6800) --
(1485.0100,1092.5200) -- (1485.0100,1095.0200) -- (1485.8400,1096.6800) --
(1485.8400,1097.5200) -- (1486.6700,1099.1800) -- (1486.6700,1101.6800) --
(1487.5100,1102.5200) -- (1487.5100,1105.0200) -- (1488.3400,1106.6800) --
(1488.3400,1107.5200) -- (1489.1700,1109.1800) -- (1489.1700,1111.6800) --
(1490.0100,1112.5200) -- (1490.0100,1115.0200) -- (1490.8400,1115.8500) --
(1490.8400,1117.5200) -- (1491.6700,1118.3500) -- (1491.6700,1120.8500) --
(1492.5100,1122.5200) -- (1492.5100,1125.0200) -- (1493.3400,1125.8500) --
(1493.3400,1127.5200) -- (1494.1700,1128.3500) -- (1494.1700,1130.8500) --
(1495.0100,1132.5200) -- (1495.0100,1135.0200) -- (1495.8400,1135.8500) --
(1495.8400,1137.5200) -- (1496.6700,1138.3500) -- (1496.6700,1140.8500) --
(1497.5100,1142.5200) -- (1497.5100,1143.3500) -- (1498.3400,1144.1800) --
(1498.3400,1146.6800) -- (1499.1700,1148.3500) -- (1499.1700,1149.1800) --
(1500.0100,1150.8500) -- (1500.0100,1153.3500) -- (1500.8400,1154.1800) --
(1500.8400,1156.6800) -- (1501.6700,1158.3500) -- (1501.6700,1159.1800) --
(1502.5100,1160.8500) -- (1502.5100,1163.3500) -- (1503.3400,1164.1800) --
(1503.3400,1165.8500) -- (1504.1700,1166.6800) -- (1504.1700,1168.3500) --
(1505.0100,1169.1800) -- (1505.0100,1171.6800) -- (1505.8400,1173.3500) --
(1505.8400,1174.1800) -- (1506.6700,1175.0200) -- (1506.6700,1177.5200) --
(1507.5100,1179.1800) -- (1507.5100,1180.0200) -- (1508.3400,1181.6800) --
(1508.3400,1184.1800) -- (1509.1700,1185.0200) -- (1509.1700,1186.6800) --
(1510.0100,1187.5200) -- (1510.0100,1189.1800) -- (1510.8400,1190.0200) --
(1510.8400,1192.5200) -- (1511.6700,1194.1800) -- (1511.6700,1195.0200) --
(1512.5100,1196.6800) -- (1512.5100,1199.1800) -- (1513.3400,1200.0200) --
(1513.3400,1201.6800) -- (1514.1700,1202.5200) -- (1514.1700,1203.3500) --
(1515.0100,1205.0200) -- (1515.0100,1207.5200) -- (1515.8400,1208.3500) --
(1515.8400,1210.0200) -- (1516.6700,1210.8500) -- (1516.6700,1212.5200) --
(1517.5100,1213.3500) -- (1517.5100,1215.8500) -- (1518.3400,1217.5200) --
(1518.3400,1218.3500) -- (1519.1700,1220.0200) -- (1519.1700,1220.8500) --
(1520.0100,1222.5200) -- (1520.0100,1223.3500) -- (1520.8400,1225.0200) --
(1520.8400,1227.5200) -- (1521.6700,1228.3500) -- (1521.6700,1230.0200) --
(1522.5100,1230.8500) -- (1522.5100,1231.6800) -- (1523.3400,1233.3500) --
(1523.3400,1234.1800) -- (1524.1700,1235.8500) -- (1524.1700,1238.3500) --
(1525.0100,1239.1800) -- (1525.0100,1240.8500) -- (1525.8400,1241.6800) --
(1525.8400,1243.3500) -- (1526.6700,1244.1800) -- (1526.6700,1245.8500) --
(1527.5100,1246.6800) -- (1527.5100,1249.1800) -- (1528.3400,1250.8500) --
(1528.3400,1251.6800) -- (1529.1700,1253.3500) -- (1529.1700,1254.1800) --
(1530.0100,1255.8500) -- (1530.0100,1256.6800) -- (1530.8400,1258.3500) --
(1530.8400,1259.1800) -- (1531.6700,1260.8500) -- (1531.6700,1261.6800) --
(1532.5100,1262.5200) -- (1532.5100,1265.0200) -- (1533.3400,1266.6800) --
(1533.3400,1267.5200) -- (1534.1700,1269.1800) -- (1534.1700,1270.0200) --
(1535.0100,1271.6800) -- (1535.0100,1272.5200) -- (1535.8400,1274.1800) --
(1535.8400,1275.0200) -- (1536.6700,1276.6800) -- (1536.6700,1277.5200) --
(1537.5100,1279.1800) -- (1537.5100,1280.0200) -- (1538.3400,1281.6800) --
(1538.3400,1282.5200) -- (1539.1700,1284.1800) -- (1539.1700,1285.0200) --
(1540.0100,1286.6800) -- (1540.0100,1287.5200) -- (1540.8400,1289.1800) --
(1540.8400,1290.0200) -- (1541.6700,1290.8500) -- (1541.6700,1293.3500) --
(1542.5100,1295.0200) -- (1542.5100,1295.8500) -- (1543.3400,1297.5200) --
(1543.3400,1298.3500) -- (1544.1700,1300.0200) -- (1544.1700,1300.8500) --
(1545.0100,1302.5200) -- (1545.0100,1303.3500) -- (1545.8400,1305.0200) --
(1545.8400,1305.8500) -- (1546.6700,1307.5200) -- (1546.6700,1308.3500) --
(1547.5100,1310.0200) -- (1547.5100,1310.8500) -- (1548.3400,1312.5200) --
(1548.3400,1313.3500) -- (1549.1700,1315.0200) -- (1549.1700,1315.8500) --
(1550.0100,1317.5200) -- (1550.0100,1318.3500) -- (1550.8400,1320.0200) --
(1550.8400,1320.8500) -- (1551.6700,1321.6800) -- (1551.6700,1323.3500) --
(1552.5100,1324.1800) -- (1552.5100,1325.8500) -- (1553.3400,1326.6800) --
(1553.3400,1328.3500) -- (1554.1700,1329.1800) -- (1554.1700,1330.8500) --
(1555.0100,1331.6800) -- (1555.8400,1333.3500) -- (1555.8400,1334.1800) --
(1556.6700,1335.8500) -- (1556.6700,1336.6800) -- (1557.5100,1338.3500) --
(1557.5100,1339.1800) -- (1558.3400,1340.8500) -- (1558.3400,1341.6800) --
(1559.1700,1343.3500) -- (1559.1700,1344.1800) -- (1560.0100,1345.8500) --
(1560.0100,1346.6800) -- (1560.8400,1348.3500) -- (1560.8400,1349.1800) --
(1561.6700,1350.0200) -- (1561.6700,1351.6800) -- (1562.5100,1352.5200) --
(1562.5100,1354.1800) -- (1563.3400,1355.0200) -- (1564.1700,1356.6800) --
(1564.1700,1357.5200) -- (1565.0100,1359.1800) -- (1565.0100,1360.0200) --
(1565.8400,1361.6800) -- (1565.8400,1362.5200) -- (1566.6700,1364.1800) --
(1566.6700,1365.0200) -- (1567.5100,1366.6800) -- (1567.5100,1367.5200) --
(1568.3400,1369.1800) -- (1568.3400,1370.0200) -- (1569.1700,1371.6800) --
(1570.0100,1372.5200) -- (1570.0100,1374.1800) -- (1570.8400,1375.0200) --
(1570.8400,1376.6800) -- (1571.6700,1377.5200) -- (1571.6700,1378.3500) --
(1572.5100,1380.0200) -- (1572.5100,1380.8500) -- (1573.3400,1382.5200) --
(1574.1700,1383.3500) -- (1574.1700,1385.0200) -- (1575.0100,1385.8500) --
(1575.0100,1387.5200) -- (1575.8400,1388.3500) -- (1575.8400,1390.0200) --
(1576.6700,1390.8500) -- (1576.6700,1392.5200) -- (1577.5100,1393.3500) --
(1578.3400,1395.0200) -- (1578.3400,1395.8500) -- (1579.1700,1397.5200) --
(1579.1700,1398.3500) -- (1580.0100,1400.0200) -- (1580.0100,1400.8500) --
(1580.8400,1402.5200) -- (1581.6700,1403.3500) -- (1581.6700,1405.0200) --
(1582.5100,1405.8500) -- (1582.5100,1407.5200) -- (1583.3400,1408.3500) --
(1583.3400,1409.1800) -- (1584.1700,1410.8500) -- (1585.0100,1411.6800) --
(1585.0100,1413.3500) -- (1585.8400,1414.1800) -- (1585.8400,1415.8500) --
(1586.6700,1416.6800) -- (1586.6700,1418.3500) -- (1587.5100,1419.1800) --
(1588.3400,1420.8500) -- (1588.3400,1421.6800) -- (1589.1700,1423.3500) --
(1589.1700,1424.1800) -- (1590.0100,1425.8500) -- (1590.8400,1426.6800) --
(1590.8400,1428.3500) -- (1591.6700,1429.1800) -- (1591.6700,1430.8500) --
(1592.5100,1431.6800) -- (1593.3400,1433.3500) -- (1593.3400,1434.1800) --
(1594.1700,1435.8500) -- (1594.1700,1436.6800) -- (1595.0100,1437.5200) --
(1595.8400,1439.1800) -- (1595.8400,1440.0200) -- (1596.6700,1441.6800) --
(1596.6700,1442.5200) -- (1597.5100,1444.1800) -- (1598.3400,1445.0200) --
(1598.3400,1446.6800) -- (1599.1700,1447.5200) -- (1599.1700,1449.1800) --
(1600.0100,1450.0200) -- (1600.8400,1451.6800) -- (1600.8400,1452.5200) --
(1601.6700,1454.1800) -- (1602.5100,1455.0200) -- (1602.5100,1456.6800) --
(1603.3400,1457.5200) -- (1603.3400,1459.1800) -- (1604.1700,1460.0200) --
(1605.0100,1461.6800) -- (1605.0100,1462.5200) -- (1605.8400,1464.1800) --
(1606.6700,1465.0200) -- (1606.6700,1465.8500) -- (1607.5100,1467.5200) --
(1607.5100,1468.3500) -- (1608.3400,1470.0200) -- (1609.1800,1470.8500) --
(1609.1800,1472.5200) -- (1610.0100,1473.3500) -- (1610.8400,1475.0200) --
(1610.8400,1475.8500) -- (1611.6800,1477.5200) -- (1612.5100,1478.3500) --
(1612.5100,1480.0200) -- (1613.3400,1480.8500) -- (1613.3400,1482.5200) --
(1614.1800,1483.3500) -- (1615.0100,1485.0200) -- (1615.0100,1485.8500) --
(1615.8400,1487.5200) -- (1616.6800,1488.3500) -- (1616.6800,1490.0200) --
(1617.5100,1490.8500) -- (1618.3400,1492.5200) -- (1618.3400,1493.3500) --
(1619.1800,1495.0200) -- (1620.0100,1495.8500) -- (1620.0100,1496.6900) --
(1620.8400,1498.3500) -- (1621.6800,1499.1900) -- (1621.6800,1500.8500) --
(1622.5100,1501.6900) -- (1623.3400,1503.3500) -- (1623.3400,1504.1900) --
(1624.1800,1505.8500) -- (1625.0100,1506.6900) -- (1625.0100,1508.3500) --
(1625.8400,1509.1900) -- (1626.6800,1510.8500) -- (1626.6800,1511.6900) --
(1627.5100,1513.3500) -- (1628.3400,1514.1900) -- (1628.3400,1515.8500) --
(1629.1800,1516.6900) -- (1630.0100,1518.3500) -- (1630.0100,1519.1900) --
(1630.8400,1520.8500) -- (1631.6800,1521.6900) -- (1631.6800,1523.3500) --
(1632.5100,1524.1900) -- (1633.3400,1525.0200) -- (1633.3400,1526.6900) --
(1634.1800,1527.5200) -- (1177.5000,1527.5200)
"""
data_1colon1 = """(1795.0000,1527.5200) -- (1795.0000,1526.6900) --
(1795.8300,1525.0200) -- (1795.8300,1524.1900) -- (1796.6700,1523.3500) --
(1796.6700,1521.6900) -- (1797.5000,1520.8500) -- (1797.5000,1519.1900) --
(1798.3300,1518.3500) -- (1798.3300,1516.6900) -- (1799.1700,1515.8500) --
(1799.1700,1513.3500) -- (1800.0000,1511.6900) -- (1800.0000,1510.8500) --
(1800.8300,1509.1900) -- (1800.8300,1508.3500) -- (1801.6700,1506.6900) --
(1801.6700,1505.8500) -- (1802.5000,1504.1900) -- (1802.5000,1501.6900) --
(1803.3300,1500.8500) -- (1803.3300,1499.1900) -- (1804.1700,1498.3500) --
(1804.1700,1496.6900) -- (1805.0000,1495.8500) -- (1805.0000,1495.0200) --
(1805.8300,1493.3500) -- (1805.8300,1492.5200) -- (1806.6700,1490.8500) --
(1806.6700,1488.3500) -- (1807.5000,1487.5200) -- (1807.5000,1485.8500) --
(1808.3300,1485.0200) -- (1808.3300,1483.3500) -- (1809.1700,1482.5200) --
(1809.1700,1480.8500) -- (1810.0000,1480.0200) -- (1810.0000,1478.3500) --
(1810.8300,1477.5200) -- (1810.8300,1475.0200) -- (1811.6700,1473.3500) --
(1811.6700,1472.5200) -- (1812.5000,1470.8500) -- (1812.5000,1470.0200) --
(1813.3300,1468.3500) -- (1813.3300,1467.5200) -- (1814.1700,1465.8500) --
(1814.1700,1465.0200) -- (1815.0000,1464.1800) -- (1815.0000,1461.6800) --
(1815.8300,1460.0200) -- (1815.8300,1459.1800) -- (1816.6700,1457.5200) --
(1816.6700,1456.6800) -- (1817.5000,1455.0200) -- (1817.5000,1454.1800) --
(1818.3300,1452.5200) -- (1818.3300,1451.6800) -- (1819.1700,1450.0200) --
(1819.1700,1449.1800) -- (1820.0000,1447.5200) -- (1820.0000,1445.0200) --
(1820.8300,1444.1800) -- (1820.8300,1442.5200) -- (1821.6700,1441.6800) --
(1821.6700,1440.0200) -- (1822.5000,1439.1800) -- (1822.5000,1437.5200) --
(1823.3300,1436.6800) -- (1823.3300,1435.8500) -- (1824.1700,1434.1800) --
(1824.1700,1433.3500) -- (1825.0000,1431.6800) -- (1825.0000,1429.1800) --
(1825.8300,1428.3500) -- (1825.8300,1426.6800) -- (1826.6700,1425.8500) --
(1826.6700,1424.1800) -- (1827.5000,1423.3500) -- (1827.5000,1421.6800) --
(1828.3300,1420.8500) -- (1828.3300,1419.1800) -- (1829.1700,1418.3500) --
(1829.1700,1416.6800) -- (1830.0000,1415.8500) -- (1830.0000,1413.3500) --
(1830.8300,1411.6800) -- (1830.8300,1410.8500) -- (1831.6700,1409.1800) --
(1831.6700,1408.3500) -- (1832.5000,1407.5200) -- (1832.5000,1405.8500) --
(1833.3300,1405.0200) -- (1833.3300,1403.3500) -- (1834.1700,1402.5200) --
(1834.1700,1400.8500) -- (1835.0000,1400.0200) -- (1835.0000,1398.3500) --
(1835.8300,1397.5200) -- (1835.8300,1395.0200) -- (1836.6700,1393.3500) --
(1836.6700,1392.5200) -- (1837.5000,1390.8500) -- (1837.5000,1390.0200) --
(1838.3300,1388.3500) -- (1838.3300,1387.5200) -- (1839.1700,1385.8500) --
(1839.1700,1385.0200) -- (1840.0000,1383.3500) -- (1840.0000,1382.5200) --
(1840.8300,1380.8500) -- (1840.8300,1380.0200) -- (1841.6700,1378.3500) --
(1841.6700,1376.6800) -- (1842.5000,1375.0200) -- (1842.5000,1374.1800) --
(1843.3300,1372.5200) -- (1843.3300,1371.6800) -- (1844.1700,1370.0200) --
(1844.1700,1369.1800) -- (1845.0000,1367.5200) -- (1845.0000,1366.6800) --
(1845.8300,1365.0200) -- (1845.8300,1364.1800) -- (1846.6700,1362.5200) --
(1846.6700,1361.6800) -- (1847.5000,1360.0200) -- (1847.5000,1359.1800) --
(1848.3300,1357.5200) -- (1848.3300,1355.0200) -- (1849.1700,1354.1800) --
(1849.1700,1352.5200) -- (1850.0000,1351.6800) -- (1850.0000,1350.0200) --
(1850.8300,1349.1800) -- (1850.8300,1348.3500) -- (1851.6700,1346.6800) --
(1851.6700,1345.8500) -- (1852.5000,1344.1800) -- (1852.5000,1343.3500) --
(1853.3300,1341.6800) -- (1853.3300,1340.8500) -- (1854.1700,1339.1800) --
(1854.1700,1338.3500) -- (1855.0000,1336.6800) -- (1855.0000,1335.8500) --
(1855.8300,1334.1800) -- (1855.8300,1333.3500) -- (1856.6700,1331.6800) --
(1856.6700,1329.1800) -- (1857.5000,1328.3500) -- (1857.5000,1326.6800) --
(1858.3300,1325.8500) -- (1858.3300,1324.1800) -- (1859.1700,1323.3500) --
(1859.1700,1321.6800) -- (1860.0000,1320.8500) -- (1860.0000,1320.0200) --
(1860.8300,1318.3500) -- (1860.8300,1317.5200) -- (1861.6700,1315.8500) --
(1861.6700,1315.0200) -- (1862.5000,1313.3500) -- (1862.5000,1312.5200) --
(1863.3300,1310.8500) -- (1863.3300,1310.0200) -- (1864.1700,1308.3500) --
(1864.1700,1307.5200) -- (1865.0000,1305.8500) -- (1865.0000,1305.0200) --
(1865.8300,1303.3500) -- (1865.8300,1302.5200) -- (1866.6700,1300.8500) --
(1866.6700,1300.0200) -- (1867.5000,1298.3500) -- (1867.5000,1295.8500) --
(1868.3300,1295.0200) -- (1868.3300,1293.3500) -- (1869.1700,1292.5200) --
(1869.1700,1290.8500) -- (1870.0000,1290.0200) -- (1870.0000,1289.1800) --
(1870.8300,1287.5200) -- (1870.8300,1286.6800) -- (1871.6700,1285.0200) --
(1871.6700,1284.1800) -- (1872.5000,1282.5200) -- (1872.5000,1281.6800) --
(1873.3300,1280.0200) -- (1873.3300,1279.1800) -- (1874.1700,1277.5200) --
(1874.1700,1276.6800) -- (1875.0000,1275.0200) --
(1875.0000,1274.1800) -- (1875.8300,1272.5200) --
(1875.8300,1271.6900) -- (1876.6700,1270.0200) -- (1876.6700,1269.1900) --
(1877.5000,1267.5200) -- (1877.5000,1266.6900) -- (1878.3300,1265.0200) --
(1878.3300,1264.1900) -- (1879.1700,1262.5200) -- (1879.1700,1261.6900) --
(1880.0000,1260.8500) -- (1880.0000,1259.1900) -- (1880.8300,1258.3500) --
(1880.8300,1256.6900) -- (1881.6700,1255.8500) -- (1881.6700,1254.1900) --
(1882.5000,1253.3500) -- (1882.5000,1251.6900) -- (1883.3300,1250.8500) --
(1883.3300,1249.1900) -- (1884.1700,1248.3500) -- (1884.1700,1246.6900) --
(1885.0000,1245.8500) -- (1885.0000,1244.1900) -- (1885.8300,1243.3500) --
(1885.8300,1241.6900) -- (1886.6700,1240.8500) -- (1886.6700,1239.1900) --
(1887.5000,1238.3500) -- (1887.5000,1236.6900) -- (1888.3300,1235.8500) --
(1888.3300,1234.1900) -- (1889.1700,1233.3500) -- (1889.1700,1231.6900) --
(1890.0000,1230.8500) -- (1890.0000,1230.0200) -- (1890.8300,1228.3500) --
(1890.8300,1227.5200) -- (1891.6700,1225.8500) -- (1891.6700,1225.0200) --
(1892.5000,1223.3500) -- (1892.5000,1222.5200) -- (1893.3300,1220.8500) --
(1893.3300,1220.0200) -- (1894.1700,1218.3500) -- (1894.1700,1217.5200) --
(1895.0000,1215.8500) -- (1895.0000,1215.0200) -- (1895.8300,1213.3500) --
(1895.8300,1212.5200) -- (1896.6700,1210.8500) -- (1896.6700,1210.0200) --
(1897.5000,1208.3500) -- (1897.5000,1207.5200) -- (1898.3300,1205.8500) --
(1898.3300,1205.0200) -- (1899.1700,1203.3500) -- (1899.1700,1202.5200) --
(1900.0000,1201.6900) -- (1900.0000,1200.0200) -- (1900.8300,1199.1900) --
(1900.8300,1197.5200) -- (1901.6700,1196.6900) -- (1901.6700,1195.0200) --
(1902.5000,1194.1900) -- (1902.5000,1192.5200) -- (1903.3300,1191.6900) --
(1903.3300,1190.0200) -- (1904.1700,1189.1900) -- (1904.1700,1187.5200) --
(1905.0000,1186.6900) -- (1905.0000,1185.0200) -- (1905.8300,1184.1900) --
(1905.8300,1182.5200) -- (1906.6700,1181.6900) -- (1906.6700,1180.0200) --
(1907.5000,1179.1900) -- (1907.5000,1177.5200) -- (1908.3300,1176.6900) --
(1908.3300,1175.0200) -- (1909.1700,1174.1900) -- (1909.1700,1173.3500) --
(1910.0000,1171.6900) -- (1910.0000,1170.8500) -- (1910.8300,1169.1900) --
(1910.8300,1168.3500) -- (1911.6700,1166.6900) -- (1911.6700,1165.8500) --
(1912.5000,1164.1900) -- (1912.5000,1163.3500) -- (1913.3300,1161.6900) --
(1913.3300,1160.8500) -- (1914.1700,1159.1900) -- (1914.1700,1158.3500) --
(1915.0000,1156.6900) -- (1915.8300,1155.8500) -- (1915.8300,1154.1900) --
(1916.6700,1153.3500) -- (1916.6700,1151.6900) -- (1917.5000,1150.8500) --
(1917.5000,1149.1900) -- (1918.3300,1148.3500) -- (1918.3300,1146.6900) --
(1919.1700,1145.8500) -- (1919.1700,1144.1900) -- (1920.0000,1143.3500) --
(1920.0000,1142.5200) -- (1920.8300,1140.8500) -- (1920.8300,1140.0200) --
(1921.6700,1138.3500) -- (1921.6700,1137.5200) -- (1922.5000,1135.8500) --
(1922.5000,1135.0200) -- (1923.3300,1133.3500) -- (1923.3300,1132.5200) --
(1924.1700,1130.8500) -- (1924.1700,1130.0200) -- (1925.0000,1128.3500) --
(1925.8300,1127.5200) -- (1925.8300,1125.8500) -- (1926.6700,1125.0200) --
(1926.6700,1123.3500) -- (1927.5000,1122.5200) -- (1927.5000,1120.8500) --
(1928.3300,1120.0200) -- (1928.3300,1118.3500) -- (1929.1700,1117.5200) --
(1929.1700,1115.8500) -- (1930.0000,1115.0200) -- (1930.0000,1114.1900) --
(1930.8300,1112.5200) -- (1930.8300,1111.6900) -- (1931.6700,1110.0200) --
(1932.5000,1109.1900) -- (1932.5000,1107.5200) -- (1933.3300,1106.6900) --
(1933.3300,1105.0200) -- (1934.1700,1104.1900) -- (1934.1700,1102.5200) --
(1935.0000,1101.6900) -- (1935.0000,1100.0200) -- (1935.8300,1099.1900) --
(1935.8300,1097.5200) -- (1936.6700,1096.6900) -- (1936.6700,1095.0200) --
(1937.5000,1094.1900) -- (1938.3300,1092.5200) -- (1938.3300,1091.6900) --
(1939.1700,1090.0200) -- (1939.1700,1089.1900) -- (1940.0000,1087.5200) --
(1940.0000,1086.6900) -- (1940.8300,1085.8500) -- (1940.8300,1084.1900) --
(1941.6700,1083.3500) -- (1941.6700,1081.6900) -- (1942.5000,1080.8500) --
(1942.5000,1079.1900) -- (1943.3300,1078.3500) -- (1944.1700,1076.6900) --
(1944.1700,1075.8500) -- (1945.0000,1074.1900) -- (1945.0000,1073.3500) --
(1945.8300,1071.6900) -- (1945.8300,1070.8500) -- (1946.6700,1069.1900) --
(1946.6700,1068.3500) -- (1947.5000,1066.6900) -- (1948.3300,1065.8500) --
(1948.3300,1064.1900) -- (1949.1700,1063.3500) -- (1949.1700,1061.6900) --
(1950.0000,1060.8500) -- (1950.0000,1059.1900) -- (1950.8300,1058.3500) --
(1950.8300,1056.6900) -- (1951.6700,1055.8500) -- (1952.5000,1055.0200) --
(1952.5000,1053.3500) -- (1953.3300,1052.5200) -- (1953.3300,1050.8500) --
(1954.1700,1050.0200) -- (1954.1700,1048.3500) -- (1955.0000,1047.5200) --
(1955.0000,1045.8500) -- (1955.8300,1045.0200) -- (1956.6700,1043.3500) --
(1956.6700,1042.5200) -- (1957.5000,1040.8500) -- (1957.5000,1040.0200) --
(1958.3300,1038.3500) -- (1958.3300,1037.5200) -- (1959.1700,1035.8500) --
(1959.1700,1035.0200) -- (1960.0000,1033.3500) -- (1960.8300,1032.5200) --
(1960.8300,1030.8500) -- (1961.6700,1030.0200) -- (1961.6700,1028.3500) --
(1962.5000,1027.5200) -- (1962.5000,1026.6900) -- (1963.3300,1025.0200) --
(1964.1700,1024.1900) -- (1964.1700,1022.5200) -- (1965.0000,1021.6900) --
(1965.0000,1020.0200) -- (1965.8300,1019.1900) -- (1965.8300,1017.5200) --
(1966.6700,1016.6900) -- (1967.5000,1015.0200) -- (1967.5000,1014.1900) --
(1968.3300,1012.5200) -- (1968.3300,1011.6900) -- (1969.1700,1010.0200) --
(1970.0000,1009.1900) -- (1970.0000,1007.5200) -- (1970.8300,1006.6900) --
(1970.8300,1005.0200) -- (1971.6700,1004.1900) -- (1971.6700,1002.5200) --
(1972.5000,1001.6900) -- (1973.3300,1000.0200) -- (1973.3300,999.1880) --
(1974.1700,997.5200) -- (1974.1700,996.6880) -- (1975.0000,995.8520) --
(1975.8300,994.1880) -- (1975.8300,993.3520) -- (1976.6700,991.6880) --
(1976.6700,990.8520) -- (1977.5000,989.1880) -- (1978.3300,988.3520) --
(1978.3300,986.6880) -- (1979.1700,985.8520) -- (1979.1700,984.1880) --
(1980.0000,983.3520) -- (1980.8300,981.6880) -- (1980.8300,980.8520) --
(1981.6700,979.1880) -- (1981.6700,978.3520) -- (1982.5000,976.6880) --
(1983.3300,975.8520) -- (1983.3300,974.1880) -- (1984.1700,973.3520) --
(1984.1700,971.6880) -- (1985.0000,970.8520) -- (1985.8300,969.1880) --
(1985.8300,968.3520) -- (1986.6700,967.5200) -- (1986.6700,965.8520) --
(1987.5000,965.0200) -- (1988.3300,963.3520) -- (1988.3300,962.5200) --
(1989.1700,960.8520) -- (1989.1700,960.0200) -- (1990.0000,958.3520) --
(1990.8300,957.5200) -- (1990.8300,955.8520) -- (1991.6700,955.0200) --
(1992.5000,953.3520) -- (1992.5000,952.5200) -- (1993.3300,950.8520) --
(1993.3300,950.0200) -- (1994.1700,948.3520) -- (1994.1700,947.5200) --
(1995.0000,945.8520) -- (1995.0000,945.0200) -- (1995.8300,943.3520) --
(1995.8300,940.8520) -- (1996.6700,940.0200) -- (1996.6700,939.1880) --
(1997.5000,937.5200) -- (1997.5000,936.6880) -- (1998.3300,935.0200) --
(1998.3300,932.5200) -- (1999.1700,931.6880) -- (1999.1700,930.0200) --
(2000.0000,929.1880) -- (2000.0000,926.6880) -- (2000.8300,925.0200) --
(2000.8300,924.1880) -- (2001.6700,922.5200) -- (2001.6700,920.0200) --
(2002.5000,919.1880) -- (2002.5000,916.6880) -- (2003.3300,915.0200) --
(2003.3300,914.1880) -- (2004.1700,912.5200) -- (2004.1700,910.0200) --
(2005.0000,909.1880) -- (2005.0000,908.3520) -- (2005.8300,906.6880) --
(2005.8300,905.8520) -- (2005.8300,904.1880) --
(2006.6700,903.3520) -- (2006.6700,901.6880) -- (2007.5000,900.8520) --
(2007.5000,898.3520) -- (2008.3300,896.6880) -- (2008.3300,895.8520) --
(2009.1700,894.1880) -- (2009.1700,891.6880) -- (2010.0000,890.8520) --
(2010.0000,888.3520) -- (2010.8300,886.6880) -- (2010.8300,885.8520) --
(2011.6700,884.1880) -- (2011.6700,881.6880) -- (2012.5000,880.8520) --
(2012.5000,880.0200) -- (2013.3300,878.3520) -- (2013.3300,875.8520) --
(2014.1700,875.0200) -- (2014.1700,872.5200) -- (2015.0000,870.8520) --
(2015.0000,870.0200) -- (2015.8300,868.3520) -- (2015.8300,865.8520) --
(2016.6700,865.0200) -- (2016.6700,862.5200) -- (2017.5000,860.8520) --
(2017.5000,860.0200) -- (2018.3300,858.3520) -- (2018.3300,855.8520) --
(2019.1700,855.0200) -- (2019.1700,852.5200) -- (2020.0000,851.6880) --
(2020.0000,850.0200) -- (2020.8300,849.1880) -- (2020.8300,846.6880) --
(2021.6700,845.0200) -- (2021.6700,842.5200) -- (2022.5000,841.6880) --
(2022.5000,840.0200) -- (2023.3300,839.1880) -- (2023.3300,836.6880) --
(2024.1700,835.0200) -- (2024.1700,832.5200) -- (2025.0000,831.6880) --
(2025.0000,830.0200) -- (2025.8300,829.1880) -- (2025.8300,826.6880) --
(2026.6700,825.0200) -- (2026.6700,822.5200) -- (2027.5000,821.6880) --
(2027.5000,819.1880) -- (2028.3300,818.3520) -- (2028.3300,816.6880) --
(2029.1700,815.8520) -- (2029.1700,813.3520) -- (2030.0000,811.6880) --
(2030.0000,809.1880) -- (2030.8300,808.3520) -- (2030.8300,806.6880) --
(2031.6700,805.8520) -- (2031.6700,803.3520) -- (2032.5000,801.6880) --
(2032.5000,799.1880) -- (2033.3300,798.3520) -- (2033.3300,795.8520) --
(2034.1700,794.1880) -- (2034.1700,793.3520) -- (2035.0000,792.5200) --
(2035.0000,790.0200) -- (2035.8300,788.3520) -- (2035.8300,785.8520) --
(2036.6700,785.0200) -- (2036.6700,782.5200) -- (2037.5000,780.8520) --
(2037.5000,780.0200) -- (2038.3300,778.3520) -- (2038.3300,775.8520) --
(2039.1700,775.0200) -- (2039.1700,772.5200) -- (2040.0000,770.8520) --
(2040.0000,768.3520) -- (2040.8300,767.5200) -- (2040.8300,765.8520) --
(2041.6700,765.0200) -- (2041.6700,762.5200) -- (2042.5000,761.6880) --
(2042.5000,759.1880) -- (2043.3300,757.5200) -- (2043.3300,755.0200) --
(2044.1700,754.1880) -- (2044.1700,752.5200) -- (2045.0000,751.6880) --
(2045.0000,749.1880) -- (2045.8300,747.5200) -- (2045.8300,745.0200) --
(2046.6700,744.1880) -- (2046.6700,741.6880) -- (2047.5000,740.0200) --
(2047.5000,739.1880) -- (2048.3300,737.5200) -- (2048.3300,735.0200) --
(2049.1700,734.1880) -- (2049.1700,731.6880) -- (2050.0000,730.8520) --
(2050.0000,728.3520) -- (2050.8300,726.6880) -- (2050.8300,724.1880) --
(2051.6700,723.3520) -- (2051.6700,721.6880) -- (2052.5000,720.8520) --
(2052.5000,718.3520) -- (2053.3300,716.6880) -- (2053.3300,714.1880) --
(2054.1700,713.3520) -- (2054.1700,710.8520) -- (2055.0000,709.1880) --
(2055.0000,708.3520) -- (2055.8300,706.6880) -- (2055.8300,705.0200) --
(2056.6700,703.3520) -- (2056.6700,700.8520) -- (2057.5000,700.0200) --
(2057.5000,697.5200) -- (2058.3300,695.8520) -- (2058.3300,693.3520) --
(2059.1700,692.5200) -- (2059.1700,690.8520) -- (2060.0000,690.0200) --
(2060.0000,687.5200) -- (2060.8300,685.8520) -- (2060.8300,683.3520) --
(2061.6700,682.5200) -- (2061.6700,680.0200) -- (2062.5000,678.3520) --
(2062.5000,675.8520) -- (2063.3300,675.0200) -- (2063.3300,674.1880) --
(2064.1700,672.5200) -- (2064.1700,670.0200) -- (2065.0000,669.1880) --
(2065.0000,666.6880) -- (2065.8300,665.0200) -- (2065.8300,662.5200) --
(2066.6700,661.6880) -- (2066.6700,659.1880) -- (2067.5000,657.5200) --
(2067.5000,655.0200) -- (2068.3300,654.1880) -- (2068.3300,652.5200) --
(2069.1700,651.6880) -- (2069.1700,649.1880) -- (2070.0000,647.5200) --
(2070.0000,645.8520) -- (2070.8300,644.1880) -- (2070.8300,641.6880) --
(2071.6700,640.8520) -- (2071.6700,638.3520) -- (2072.5000,636.6880) --
(2072.5000,635.8520) -- (2073.3300,634.1880) -- (2073.3300,631.6880) --
(2074.1700,630.8520) -- (2074.1700,628.3520) -- (2075.0000,626.6880) --
(2075.0000,624.1880) -- (2075.8300,623.3520) -- (2075.8300,620.8520) --
(2076.6700,619.1880) -- (2076.6700,618.3520) -- (2077.5000,617.5200) --
(2077.5000,615.0200) -- (2078.3300,613.3520) -- (2078.3300,610.8520) --
(2079.1700,610.0200) -- (2079.1700,607.5200) -- (2080.0000,605.8520) --
(2080.0000,603.3550) -- (2080.8300,602.5200) -- (2080.8300,600.0200) --
(2081.6700,598.3550) -- (2081.6700,597.5200) -- (2082.5000,595.8550) --
(2082.5000,593.3550) -- (2083.3300,592.5200) -- (2083.3300,590.0200) --
(2084.1700,588.3550) -- (2084.1700,586.6880) -- (2085.0000,585.0200) --
(2085.0000,582.5200) -- (2085.8300,581.6910) -- (2085.8300,579.1910) --
(2086.6700,577.5200) -- (2086.6700,576.6910) -- (2087.5000,575.0200) --
(2087.5000,572.5200) -- (2088.3300,571.6910) -- (2088.3300,569.1910) --
(2089.1700,567.5200) -- (2089.1700,565.0200) -- (2090.0000,564.1910) --
(2090.0000,561.6910) -- (2090.8300,560.0200) -- (2090.8300,558.3520) --
(2091.6700,556.6910) -- (2091.6700,555.8520) -- (2092.5000,554.1910) --
(2092.5000,551.6910) -- (2093.3300,550.8520) -- (2093.3300,548.3520) --
(2094.1700,546.6910) -- (2094.1700,544.1910) -- (2095.0000,543.3520) --
(2095.0000,540.8520) -- (2095.8300,539.1910) -- (2095.8300,538.3520) --
(2096.6700,536.6910) -- (2096.6700,534.1910) --
(2097.5000,533.3520) -- (2097.5000,530.8520) -- (2098.3300,529.1910) --
(2098.3300,527.5200) -- (2099.1700,525.8520) -- (2099.1700,523.3520) --
(2100.0000,522.5200) -- (2100.0000,520.0200) -- (2100.8300,518.3520) --
(2100.8300,517.5200) -- (2101.6700,515.8520) -- (2101.6700,513.3520) --
(2102.5000,512.5200) -- (2102.5000,510.0200) -- (2103.3300,508.3520) --
(2103.3300,505.8520) -- (2104.1700,505.0200) -- (2104.1700,502.5200) --
(2105.0000,500.8520) -- (2105.0000,499.1910) -- (2105.8300,497.5200) --
(2105.8300,496.6910) -- (2106.6700,495.0200) -- (2106.6700,492.5200) --
(2107.5000,491.6910) -- (2107.5000,489.1910) -- (2108.3300,487.5200) --
(2108.3300,485.0200) -- (2109.1700,484.1910) -- (2109.1700,481.6910) --
(2110.0000,480.0200) -- (2110.0000,477.5200) -- (2110.8300,476.6910) --
(2110.8300,475.0200) -- (2111.6700,474.1910) -- (2111.6700,471.6910) --
(2112.5000,470.8520) -- (2112.5000,468.3520) -- (2113.3300,466.6910) --
(2113.3300,464.1910) -- (2114.1700,463.3520) -- (2114.1700,460.8520) --
(2115.0000,459.1910) -- (2115.0000,456.6910) -- (2115.8300,455.8520) --
(2115.8300,454.1910) -- (2116.6700,453.3520) -- (2116.6700,450.8520) --
(2117.5000,449.1910) -- (2117.5000,446.6910) -- (2118.3300,445.8520) --
(2118.3300,443.3520) -- (2119.1700,441.6910) -- (2119.1700,440.0200) --
(2120.0000,438.3520) -- (2120.0000,437.5200) -- (2120.8300,435.8520) --
(2120.8300,433.3520) -- (2121.6700,432.5200) -- (2121.6700,430.0200) --
(2122.5000,428.3520) -- (2122.5000,425.8520) -- (2123.3300,425.0200) --
(2123.3300,422.5200) -- (2124.1700,420.8520) -- (2124.1700,418.3520) --
(2125.0000,417.5200) -- (2125.0000,415.8520) -- (2125.8300,415.0200) --
(2125.8300,412.5200) -- (2126.6700,411.6910) -- (2126.6700,409.1910) --
(2127.5000,407.5200) -- (2127.5000,405.0200) -- (2128.3300,404.1910) --
(2128.3300,401.6910) -- (2129.1700,400.0200) -- (2129.1700,399.1910) --
(2130.0000,397.5200) -- (2130.0000,395.0200) -- (2130.8300,394.1910) --
(2130.8300,391.6910) -- (2131.6700,390.0200) -- (2131.6700,387.5200) --
(2132.5000,386.6910) -- (2132.5000,384.1910) -- (2133.3300,383.3520) --
(2133.3300,381.6910) -- (2134.1700,380.8520) -- (2134.1700,378.3520) --
(2135.0000,376.6910) -- (2135.0000,374.1910) -- (2135.8300,373.3520) --
(2135.8300,370.8520) -- (2136.6700,369.1910) -- (2136.6700,366.6910) --
(2137.5000,365.8520) -- (2137.5000,363.3520) -- (2138.3300,361.6910) --
(2138.3300,360.8520) -- (2139.1700,359.1910) -- (2139.1700,356.6910) --
(2140.0000,355.8520) -- (2140.0000,353.3520) -- (2140.8300,352.5200) --
(2140.8300,350.0200) -- (2141.6700,348.3520) -- (2141.6700,345.8520) --
(2142.5000,345.0200) -- (2142.5000,343.3520) -- (2143.3300,342.5200) --
(2143.3300,340.0200) -- (2144.1700,338.3520) -- (2144.1700,335.8520) --
(2145.0000,335.0200) -- (2145.0000,332.5200) -- (2145.8300,330.8520) --
(2145.8300,328.3520) -- (2146.6700,327.5200) -- (2146.6700,325.8520) --
(2147.5000,325.0200) -- (2147.5000,322.5200) -- (2148.3300,321.6910) --
(2148.3300,319.1910) -- (2149.1700,317.5200) -- (2149.1700,315.0200) --
(2150.0000,314.1910) -- (2150.0000,311.6910) -- (2150.8300,310.0200) --
(2150.8300,309.1910) -- (2151.6700,307.5200) -- (2151.6700,305.0200) --
(2152.5000,304.1910) -- (2152.5000,301.6910) -- (2153.3300,300.0200) --
(2153.3300,297.5200) -- (2154.1700,296.6910) -- (2155.0000,296.6910) --
(2155.0000,299.1910) -- (2155.8300,300.0200) -- (2155.8300,302.5200) --
(2156.6700,304.1910) -- (2156.6700,306.6910) -- (2157.5000,307.5200) --
(2157.5000,310.0200) -- (2158.3300,311.6910) -- (2158.3300,312.5200) --
(2159.1700,314.1910) -- (2159.1700,316.6910) -- (2160.0000,317.5200) --
(2160.0000,320.0200) -- (2160.8300,321.6910) -- (2160.8300,324.1910) --
(2161.6700,325.0200) -- (2161.6700,325.8520) -- (2162.5000,327.5200) --
(2162.5000,330.0200) -- (2163.3300,330.8520) -- (2163.3300,333.3520) --
(2164.1700,335.0200) -- (2164.1700,337.5200) -- (2165.0000,338.3520) --
(2165.0000,340.8520) -- (2165.8300,342.5200) -- (2165.8300,343.3520) --
(2166.6700,345.0200) -- (2166.6700,347.5200) -- (2167.5000,348.3520) --
(2167.5000,350.8520) -- (2168.3300,352.5200) -- (2168.3300,354.1910) --
(2169.1700,355.8520) -- (2169.1700,356.6910) -- (2170.0000,358.3520) --
(2170.0000,360.8520) -- (2170.8300,361.6910) -- (2170.8300,364.1910) --
(2171.6700,365.8520) -- (2171.6700,368.3520) -- (2172.5000,369.1910) --
(2172.5000,370.8520) -- (2173.3300,371.6910) -- (2173.3300,374.1910) --
(2174.1700,375.8520) -- (2174.1700,378.3520) -- (2175.0000,379.1910) --
(2175.0000,381.6910) -- (2175.8300,383.3520) -- (2175.8300,384.1910) --
(2176.6700,385.0200) -- (2176.6700,387.5200) -- (2177.5000,389.1910) --
(2177.5000,391.6910) -- (2178.3300,392.5200) -- (2178.3300,395.0200) --
(2179.1700,396.6910) -- (2179.1700,397.5200) -- (2180.0000,399.1910) --
(2180.0000,401.6910) -- (2180.8300,402.5200) -- (2180.8300,405.0200) --
(2181.6700,406.6910) -- (2181.6700,409.1910) -- (2182.5000,410.0200) --
(2182.5000,411.6910) -- (2183.3300,412.5200) -- (2183.3300,415.0200) --
(2184.1700,415.8520) -- (2184.1700,418.3520) -- (2185.0000,420.0200) --
(2185.0000,420.8520) -- (2185.8300,422.5200) --
(2185.8300,423.3520) -- (2185.8300,425.0200) --
(2186.6700,425.8520) -- (2186.6700,428.3520) -- (2187.5000,430.0200) --
(2187.5000,432.5200) -- (2188.3300,433.3520) -- (2188.3300,435.0200) --
(2189.1700,435.8520) -- (2189.1700,438.3520) -- (2190.0000,440.0200) --
(2190.0000,441.6910) -- (2190.8300,443.3520) -- (2190.8300,445.8520) --
(2191.6700,446.6910) -- (2191.6700,448.3520) -- (2192.5000,449.1910) --
(2192.5000,451.6910) -- (2193.3300,453.3520) -- (2193.3300,455.8520) --
(2194.1700,456.6910) -- (2194.1700,458.3520) -- (2195.0000,459.1910) --
(2195.0000,461.6910) -- (2195.8300,463.3520) -- (2195.8300,465.8520) --
(2196.6700,466.6910) -- (2196.6700,468.3520) -- (2197.5000,469.1910) --
(2197.5000,471.6910) -- (2198.3300,472.5200) -- (2198.3300,475.0200) --
(2199.1700,476.6910) -- (2199.1700,479.1910) -- (2200.0000,480.0200) --
(2200.0000,481.6910) -- (2200.8300,482.5200) -- (2200.8300,485.0200) --
(2201.6700,486.6910) -- (2201.6700,489.1910) -- (2202.5000,490.0200) --
(2202.5000,491.6910) -- (2203.3300,492.5200) -- (2203.3300,495.0200) --
(2204.1700,496.6910) -- (2204.1700,499.1910) -- (2205.0000,500.0200) --
(2205.0000,500.8520) -- (2205.8300,502.5200) -- (2205.8300,505.0200) --
(2206.6700,505.8520) -- (2206.6700,508.3520) -- (2207.5000,510.0200) --
(2207.5000,510.8520) -- (2208.3300,512.5200) -- (2208.3300,515.0200) --
(2209.1700,515.8520) -- (2209.1700,518.3520) -- (2210.0000,520.0200) --
(2210.0000,520.8520) -- (2210.8300,522.5200) -- (2210.8300,525.0200) --
(2211.6700,525.8520) -- (2211.6700,528.3520) -- (2212.5000,529.1910) --
(2212.5000,530.8520) -- (2213.3300,531.6910) -- (2213.3300,534.1910) --
(2214.1700,535.8520) -- (2214.1700,538.3520) -- (2215.0000,539.1910) --
(2215.0000,540.8520) -- (2215.8300,541.6910) -- (2215.8300,544.1910) --
(2216.6700,545.8520) -- (2216.6700,548.3520) -- (2217.5000,549.1910) --
(2217.5000,550.8520) -- (2218.3300,551.6910) -- (2218.3300,554.1910) --
(2219.1700,555.8520) -- (2219.1700,558.3520) -- (2220.0000,559.1910) --
(2220.0000,560.0200) -- (2220.8300,561.6910) -- (2220.8300,564.1910) --
(2221.6700,565.0200) -- (2221.6700,567.5200) -- (2222.5000,569.1910) --
(2222.5000,570.0200) -- (2223.3300,571.6910) -- (2223.3300,574.1910) --
(2224.1700,575.0200) -- (2224.1700,576.6910) -- (2225.0000,577.5200) --
(2225.0000,580.0200) -- (2225.8300,581.6910) -- (2225.8300,584.1910) --
(2226.6700,585.0200) -- (2226.6700,586.6880) -- (2227.5000,587.5200) --
(2227.5000,590.0200) -- (2228.3300,590.8520) -- (2228.3300,593.3520) --
(2229.1700,595.0200) -- (2229.1700,595.8520) -- (2230.0000,597.5200) --
(2230.0000,600.0200) -- (2230.8300,600.8520) -- (2230.8300,602.5200) --
(2231.6700,603.3520) -- (2231.6700,605.8520) -- (2232.5000,607.5200) --
(2232.5000,610.0200) -- (2233.3300,610.8520) -- (2233.3300,612.5200) --
(2234.1700,613.3520) -- (2234.1700,615.8520) -- (2235.0000,617.5200) --
(2235.0000,618.3520) -- (2235.8300,619.1880) -- (2235.8300,621.6880) --
(2236.6700,623.3520) -- (2236.6700,625.8520) -- (2237.5000,626.6880) --
(2237.5000,628.3520) -- (2238.3300,629.1880) -- (2238.3300,631.6880) --
(2239.1700,633.3520) -- (2239.1700,634.1880) -- (2240.0000,635.8520) --
(2240.0000,638.3520) -- (2240.8300,639.1880) -- (2240.8300,640.8520) --
(2241.6700,641.6880) -- (2241.6700,644.1880) -- (2242.5000,645.8520) --
(2242.5000,647.5200) -- (2243.3300,649.1880) -- (2243.3300,650.0200) --
(2244.1700,651.6880) -- (2244.1700,654.1880) -- (2245.0000,655.0200) --
(2245.0000,656.6880) -- (2245.8300,657.5200) -- (2245.8300,660.0200) --
(2246.6700,661.6880) -- (2246.6700,662.5200) -- (2247.5000,664.1880) --
(2247.5000,666.6880) -- (2248.3300,667.5200) -- (2248.3300,669.1880) --
(2249.1700,670.0200) -- (2249.1700,672.5200) -- (2250.0000,674.1880) --
(2250.0000,675.8520) -- (2250.8300,677.5200) -- (2250.8300,678.3520) --
(2251.6700,680.0200) -- (2251.6700,682.5200) -- (2252.5000,683.3520) --
(2252.5000,685.0200) -- (2253.3300,685.8520) -- (2253.3300,688.3520) --
(2254.1700,690.0200) -- (2254.1700,690.8520) -- (2255.0000,692.5200) --
(2255.0000,695.0200) -- (2255.8300,695.8520) -- (2255.8300,697.5200) --
(2256.6700,698.3520) -- (2256.6700,700.8520) -- (2257.5000,702.5200) --
(2257.5000,703.3520) -- (2258.3300,705.0200) -- (2258.3300,706.6880) --
(2259.1700,708.3520) -- (2259.1700,709.1880) -- (2260.0000,710.8520) --
(2260.0000,713.3520) -- (2260.8300,714.1880) -- (2260.8300,715.8520) --
(2261.6700,716.6880) -- (2261.6700,719.1880) -- (2262.5000,720.8520) --
(2262.5000,721.6880) -- (2263.3300,723.3520) -- (2263.3300,725.8520) --
(2264.1700,726.6880) -- (2264.1700,728.3520) -- (2265.0000,729.1880) --
(2265.0000,731.6880) -- (2265.8300,733.3520) -- (2265.8300,734.1880) --
(2266.6700,735.0200) -- (2266.6700,737.5200) -- (2267.5000,739.1880) --
(2267.5000,740.0200) -- (2268.3300,741.6880) -- (2268.3300,744.1880) --
(2269.1700,745.0200) -- (2269.1700,746.6880) -- (2270.0000,747.5200) --
(2270.0000,749.1880) -- (2270.8300,750.0200) -- (2270.8300,752.5200) --
(2271.6700,754.1880) -- (2271.6700,755.0200) -- (2272.5000,756.6880) --
(2272.5000,759.1880) -- (2273.3300,760.0200) -- (2273.3300,761.6880) --
(2274.1700,762.5200) -- (2274.1700,765.0200) -- (2275.0000,765.8520) --
(2275.0000,767.5200) -- (2275.8300,768.3520) -- (2275.8300,770.8520) --
(2276.6700,772.5200) -- (2276.6700,773.3520) -- (2277.5000,775.0200) --
(2277.5000,775.8520) -- (2278.3300,777.5200) -- (2278.3300,780.0200) --
(2279.1700,780.8520) -- (2279.1700,782.5200) -- (2280.0000,783.3520) --
(2280.0000,785.8520) -- (2280.8300,787.5200) -- (2280.8300,788.3520) --
(2281.6700,790.0200) -- (2281.6700,790.8520) --
(2282.5000,792.5200) -- (2282.5000,794.1880) --
(2283.3300,795.8520) -- (2283.3300,796.6880) -- (2284.1700,798.3520) --
(2284.1700,800.8520) -- (2285.0000,801.6880) -- (2285.0000,803.3520) --
(2285.8300,804.1880) -- (2285.8300,805.8520) -- (2286.6700,806.6880) --
(2286.6700,809.1880) -- (2287.5000,810.8520) -- (2287.5000,811.6880) --
(2288.3300,813.3520) -- (2288.3300,815.8520) -- (2289.1700,816.6880) --
(2289.1700,818.3520) -- (2290.0000,819.1880) -- (2290.0000,820.8520) --
(2290.8300,821.6880) -- (2290.8300,824.1880) -- (2291.6700,825.0200) --
(2291.6700,826.6880) -- (2292.5000,827.5200) -- (2292.5000,829.1880) --
(2293.3300,830.0200) -- (2293.3300,832.5200) -- (2294.1700,834.1880) --
(2294.1700,835.0200) -- (2295.0000,836.6880) -- (2295.0000,837.5200) --
(2295.8300,839.1880) -- (2295.8300,841.6880) -- (2296.6700,842.5200) --
(2296.6700,844.1880) -- (2297.5000,845.0200) -- (2297.5000,846.6880) --
(2298.3300,847.5200) -- (2298.3300,849.1880) -- (2299.1700,850.0200) --
(2299.1700,852.5200) -- (2300.0000,853.3520) -- (2300.0000,855.0200) --
(2300.8300,855.8520) -- (2300.8300,857.5200) -- (2301.6700,858.3520) --
(2301.6700,860.8520) -- (2302.5000,862.5200) -- (2302.5000,863.3520) --
(2303.3300,865.0200) -- (2303.3300,865.8520) -- (2304.1700,867.5200) --
(2304.1700,868.3520) -- (2305.0000,870.0200) -- (2305.0000,872.5200) --
(2305.8300,873.3520) -- (2305.8300,875.0200) -- (2306.6700,875.8520) --
(2306.6700,877.5200) -- (2307.5000,878.3520) -- (2307.5000,880.0200) --
(2308.3300,880.8520) -- (2308.3300,883.3520) -- (2309.1700,884.1880) --
(2309.1700,885.8520) -- (2310.0000,886.6880) -- (2310.0000,888.3520) --
(2310.8300,889.1880) -- (2310.8300,890.8520) -- (2311.6700,891.6880) --
(2311.6700,894.1880) -- (2312.5000,895.8520) -- (2312.5000,896.6880) --
(2313.3300,898.3520) -- (2313.3300,899.1880) -- (2314.1700,900.8520) --
(2314.1700,901.6880) -- (2315.0000,903.3520) -- (2315.0000,904.1880) --
(2315.8300,905.8520) -- (2315.8300,908.3520) -- (2316.6700,909.1880) --
(2316.6700,910.0200) -- (2317.5000,911.6880) -- (2317.5000,912.5200) --
(2318.3300,914.1880) -- (2318.3300,915.0200) -- (2319.1700,916.6880) --
(2319.1700,917.5200) -- (2320.0000,919.1880) -- (2320.0000,920.0200) --
(2320.8300,921.6880) -- (2320.8300,922.5200) -- (2321.6700,924.1880) --
(2321.6700,925.0200) -- (2322.5000,926.6880) -- (2322.5000,929.1880) --
(2323.3300,930.0200) -- (2323.3300,931.6880) -- (2324.1700,932.5200) --
(2324.1700,934.1880) -- (2325.0000,935.0200) -- (2325.0000,936.6880) --
(2325.8300,937.5200) -- (2325.8300,939.1880) -- (2326.6700,940.0200) --
(2326.6700,940.8520) -- (2327.5000,942.5200) -- (2327.5000,943.3520) --
(2328.3300,945.0200) -- (2328.3300,945.8520) -- (2329.1700,947.5200) --
(2329.1700,948.3520) -- (2330.0000,950.0200) -- (2330.8300,950.8520) --
(2330.8300,952.5200) -- (2331.6700,953.3520) -- (2332.5000,955.0200) --
(2333.3300,955.8520) -- (2333.3300,957.5200) -- (2334.1700,958.3520) --
(2335.0000,960.0200) -- (2335.8300,960.8520) -- (2335.8300,962.5200) --
(2336.6700,963.3520) -- (2337.5000,965.0200) -- (2338.3300,965.8520) --
(2338.3300,967.5200) -- (2339.1700,968.3520) -- (2340.0000,969.1880) --
(2340.0000,970.8520) -- (2340.8300,971.6880) -- (2341.6700,973.3520) --
(2342.5000,974.1880) -- (2342.5000,975.8520) -- (2343.3300,976.6880) --
(2344.1700,978.3520) -- (2345.0000,979.1880) -- (2345.0000,980.8520) --
(2345.8300,981.6880) -- (2346.6700,983.3520) -- (2347.5000,984.1880) --
(2347.5000,985.8520) -- (2348.3300,986.6880) -- (2349.1700,988.3520) --
(2349.1700,989.1880) -- (2350.0000,990.8520) -- (2350.8300,991.6880) --
(2351.6700,993.3520) -- (2351.6700,994.1880) -- (2352.5000,995.8520) --
(2353.3300,996.6880) -- (2353.3300,997.5200) -- (2354.1700,999.1880) --
(2355.0000,1000.0200) -- (2355.8300,1001.6900) -- (2355.8300,1002.5200) --
(2356.6700,1004.1900) -- (2357.5000,1005.0200) -- (2358.3300,1006.6900) --
(2358.3300,1007.5200) -- (2359.1700,1009.1900) -- (2360.0000,1010.0200) --
(2360.0000,1011.6900) -- (2360.8300,1012.5200) -- (2361.6700,1014.1900) --
(2362.5000,1015.0200) -- (2362.5000,1016.6900) -- (2363.3300,1017.5200) --
(2364.1700,1019.1900) -- (2364.1700,1020.0200) -- (2365.0000,1021.6900) --
(2365.8300,1022.5200) -- (2365.8300,1024.1900) -- (2366.6700,1025.0200) --
(2367.5000,1026.6900) -- (2368.3300,1027.5200) -- (2368.3300,1028.3500) --
(2369.1700,1030.0200) -- (2370.0000,1030.8500) -- (2370.0000,1032.5200) --
(2370.8300,1033.3500) -- (2371.6700,1035.0200) -- (2372.5000,1035.8500) --
(2372.5000,1037.5200) -- (2373.3300,1038.3500) -- (2374.1700,1040.0200) --
(2374.1700,1040.8500) -- (2375.0000,1042.5200) -- (2375.8300,1043.3500) --
(2375.8300,1045.0200) -- (2376.6700,1045.8500) -- (2377.5000,1047.5200) --
(2378.3300,1048.3500) -- (2378.3300,1050.0200) -- (2379.1700,1050.8500) --
(2380.0000,1052.5200) -- (2380.0000,1053.3500) -- (2380.8300,1055.0200) --
(2381.6700,1055.8500) -- (2381.6700,1056.6900) -- (2382.5000,1058.3500) --
(2383.3300,1059.1900) -- (2384.1700,1060.8500) -- (2384.1700,1061.6900) --
(2385.0000,1063.3500) -- (2385.8300,1064.1900) -- (2385.8300,1065.8500) --
(2386.6700,1066.6900) -- (2387.5000,1068.3500) -- (2387.5000,1069.1900) --
(2388.3300,1070.8500) -- (2389.1700,1071.6900) -- (2390.0000,1073.3500) --
(2390.0000,1074.1900) -- (2390.8300,1075.8500) -- (2391.6700,1076.6900) --
(2391.6700,1078.3500) -- (2392.5000,1079.1900) -- (2393.3300,1080.8500) --
(2393.3300,1081.6900) -- (2394.1700,1083.3500) -- (2395.0000,1084.1900) --
(2395.0000,1085.8500) -- (2395.8300,1086.6900) -- (2396.6700,1087.5200) --
(2397.5000,1089.1900) -- (2397.5000,1090.0200) -- (2398.3300,1091.6900) --
(2399.1700,1092.5200) -- (2399.1700,1094.1900) -- (2400.0000,1095.0200) --
(2400.8300,1096.6900) -- (2400.8300,1097.5200) -- (2401.6700,1099.1900) --
(2402.5000,1100.0200) -- (2402.5000,1101.6900) -- (2403.3300,1102.5200) --
(2404.1700,1104.1900) -- (2405.0000,1105.0200) -- (2405.0000,1106.6900) --
(2405.8300,1107.5200) -- (2406.6700,1109.1900) -- (2406.6700,1110.0200) --
(2407.5000,1111.6900) -- (2408.3300,1112.5200) -- (2408.3300,1114.1900) --
(2409.1700,1115.0200) -- (2410.0000,1115.8500) -- (2410.0000,1117.5200) --
(2410.8300,1118.3500) -- (2411.6700,1120.0200) -- (2411.6700,1120.8500) --
(2412.5000,1122.5200) -- (2413.3300,1123.3500) -- (2414.1700,1125.0200) --
(2414.1700,1125.8500) -- (2415.0000,1127.5200) -- (2415.8300,1128.3500) --
(2415.8300,1130.0200) -- (2416.6700,1130.8500) -- (2417.5000,1132.5200) --
(2417.5000,1133.3500) -- (2418.3300,1135.0200) -- (2419.1700,1135.8500) --
(2419.1700,1137.5200) -- (2420.0000,1138.3500) -- (2420.8300,1140.0200) --
(2420.8300,1140.8500) -- (2421.6700,1142.5200) -- (2422.5000,1143.3500) --
(2422.5000,1144.1900) -- (2423.3300,1145.8500) -- (2424.1700,1146.6900) --
(2424.1700,1148.3500) -- (2425.0000,1149.1900) -- (2425.8300,1150.8500) --
(2426.6700,1151.6900) -- (2426.6700,1153.3500) -- (2427.5000,1154.1900) --
(2428.3300,1155.8500) -- (2428.3300,1156.6900) -- (2429.1700,1158.3500) --
(2430.0000,1159.1900) -- (2430.0000,1160.8500) --
(2430.8300,1161.6900) -- (2431.6700,1163.3500) -- (2431.6700,1164.1900) --
(2432.5000,1165.8500) -- (2433.3300,1166.6900) -- (2433.3300,1168.3500) --
(2434.1700,1169.1900) -- (2435.0000,1170.8500) -- (2435.0000,1171.6900) --
(2435.8300,1173.3500) -- (2436.6700,1174.1900) -- (2436.6700,1175.0200) --
(2437.5000,1176.6900) -- (2438.3300,1177.5200) -- (2439.1700,1179.1900) --
(2439.1700,1180.0200) -- (2440.0000,1181.6900) -- (2440.8300,1182.5200) --
(2440.8300,1184.1900) -- (2441.6700,1185.0200) -- (2442.5000,1186.6900) --
(2442.5000,1187.5200) -- (2443.3300,1189.1900) -- (2444.1700,1190.0200) --
(2444.1700,1191.6900) -- (2445.0000,1192.5200) -- (2445.8300,1194.1900) --
(2445.8300,1195.0200) -- (2446.6700,1196.6900) -- (2447.5000,1197.5200) --
(2447.5000,1199.1900) -- (2448.3300,1200.0200) -- (2449.1700,1201.6900) --
(2449.1700,1202.5200) -- (2450.0000,1203.3500) -- (2450.8300,1205.0200) --
(2450.8300,1205.8500) -- (2451.6700,1207.5200) -- (2452.5000,1208.3500) --
(2452.5000,1210.0200) -- (2453.3300,1210.8500) -- (2454.1700,1212.5200) --
(2454.1700,1213.3600) -- (2455.0000,1215.0200) -- (2455.8300,1215.8500) --
(2456.6700,1217.5200) -- (2456.6700,1218.3600) -- (2457.5000,1220.0200) --
(2458.3300,1220.8600) -- (2458.3300,1222.5200) -- (2459.1700,1223.3600) --
(2460.0000,1225.0200) -- (2460.0000,1225.8600) -- (2460.8300,1227.5200) --
(2461.6700,1228.3600) -- (2461.6700,1230.0200) -- (2462.5000,1230.8600) --
(2463.3300,1231.6900) -- (2463.3300,1233.3600) -- (2464.1700,1234.1900) --
(2465.0000,1235.8600) -- (2465.0000,1236.6900) -- (2465.8300,1238.3600) --
(2466.6700,1239.1900) -- (2466.6700,1240.8600) -- (2467.5000,1241.6900) --
(2468.3300,1243.3600) -- (2468.3300,1244.1900) -- (2469.1700,1245.8600) --
(2470.0000,1246.6900) -- (2470.0000,1248.3600) -- (2470.8300,1249.1900) --
(2471.6700,1250.8600) -- (2471.6700,1251.6900) -- (2472.5000,1253.3600) --
(2473.3300,1254.1900) -- (2473.3300,1255.8600) -- (2474.1700,1256.6900) --
(2475.0000,1258.3600) -- (2475.8300,1259.1900) -- (2475.8300,1260.8600) --
(2476.6700,1261.6900) -- (2477.5000,1262.5200) -- (2477.5000,1264.1900) --
(2478.3300,1265.0200) -- (2479.1700,1266.6900) -- (2479.1700,1267.5200) --
(2480.0000,1269.1900) -- (2480.8300,1270.0200) -- (2480.8300,1271.6900) --
(2481.6700,1272.5200) -- (2482.5000,1274.1900) -- (2482.5000,1275.0200) --
(2483.3300,1276.6900) -- (2484.1700,1277.5200) -- (2484.1700,1279.1900) --
(2485.0000,1280.0200) -- (2485.8300,1281.6900) -- (2485.8300,1282.5200) --
(2486.6700,1284.1900) -- (2487.5000,1285.0200) -- (2487.5000,1286.6900) --
(2488.3300,1287.5200) -- (2489.1700,1289.1900) -- (2489.1700,1290.0200) --
(2490.0000,1290.8600) -- (2490.8300,1292.5200) -- (2490.8300,1293.3600) --
(2491.6700,1295.0200) -- (2492.5000,1295.8600) -- (2492.5000,1297.5200) --
(2493.3300,1298.3600) -- (2494.1700,1300.0200) -- (2494.1700,1300.8600) --
(2495.0000,1302.5200) -- (2495.8300,1303.3600) -- (2495.8300,1305.0200) --
(2496.6700,1305.8600) -- (2497.5000,1307.5200) -- (2498.3300,1308.3600) --
(2498.3300,1310.0200) -- (2499.1700,1310.8600) -- (2500.0000,1312.5200) --
(2500.0000,1313.3600) -- (2500.8300,1315.0200) -- (2501.6700,1315.8600) --
(2501.6700,1317.5200) -- (2502.5000,1318.3600) -- (2503.3300,1320.0200) --
(2503.3300,1320.8600) -- (2504.1700,1321.6900) -- (2505.0000,1323.3600) --
(2505.0000,1324.1900) -- (2505.8300,1325.8600) -- (2506.6700,1326.6900) --
(2506.6700,1328.3600) -- (2507.5000,1329.1900) -- (2508.3300,1330.8600) --
(2508.3300,1331.6900) -- (2509.1700,1333.3600) -- (2510.0000,1334.1900) --
(2510.0000,1335.8600) -- (2510.8300,1336.6900) -- (2511.6700,1338.3600) --
(2511.6700,1339.1900) -- (2512.5000,1340.8600) -- (2513.3400,1341.6900) --
(2513.3400,1343.3600) -- (2514.1700,1344.1900) -- (2515.0000,1345.8600) --
(2515.0000,1346.6900) -- (2515.8400,1348.3600) -- (2516.6700,1349.1900) --
(2516.6700,1350.0200) -- (2517.5000,1351.6900) -- (2518.3400,1352.5200) --
(2519.1700,1354.1900) -- (2519.1700,1355.0200) -- (2520.0000,1356.6900) --
(2520.8400,1357.5200) -- (2520.8400,1359.1900) -- (2521.6700,1360.0200) --
(2522.5000,1361.6900) -- (2522.5000,1362.5200) -- (2523.3400,1364.1900) --
(2524.1700,1365.0200) -- (2524.1700,1366.6900) -- (2525.0000,1367.5200) --
(2525.8400,1369.1900) -- (2525.8400,1370.0200) -- (2526.6700,1371.6900) --
(2527.5000,1372.5200) -- (2527.5000,1374.1900) -- (2528.3400,1375.0200) --
(2529.1700,1376.6900) -- (2529.1700,1377.5200) -- (2530.0000,1378.3600) --
(2530.8400,1380.0200) -- (2530.8400,1380.8600) -- (2531.6700,1382.5200) --
(2532.5000,1383.3600) -- (2532.5000,1385.0200) -- (2533.3400,1385.8600) --
(2534.1700,1387.5200) -- (2534.1700,1388.3600) -- (2535.0000,1390.0200) --
(2535.8400,1390.8600) -- (2536.6700,1392.5200) -- (2536.6700,1393.3600) --
(2537.5000,1395.0200) -- (2538.3400,1395.8600) -- (2538.3400,1397.5200) --
(2539.1700,1398.3600) -- (2540.0000,1400.0200) -- (2540.0000,1400.8600) --
(2540.8400,1402.5200) -- (2541.6700,1403.3600) -- (2541.6700,1405.0200) --
(2542.5000,1405.8600) -- (2543.3400,1407.5200) -- (2543.3400,1408.3600) --
(2544.1700,1409.1900) -- (2545.0000,1410.8600) -- (2545.0000,1411.6900) --
(2545.8400,1413.3600) -- (2546.6700,1414.1900) -- (2546.6700,1415.8600) --
(2547.5000,1416.6900) -- (2548.3400,1418.3600) -- (2548.3400,1419.1900) --
(2549.1700,1420.8600) -- (2550.0000,1421.6900) -- (2550.0000,1423.3600) --
(2550.8400,1424.1900) -- (2551.6700,1425.8600) -- (2552.5000,1426.6900) --
(2552.5000,1428.3600) -- (2553.3400,1429.1900) -- (2554.1700,1430.8600) --
(2554.1700,1431.6900) -- (2555.0000,1433.3600) -- (2555.8400,1434.1900) --
(2555.8400,1435.8600) -- (2556.6700,1436.6900) -- (2557.5000,1437.5200) --
(2557.5000,1439.1900) -- (2558.3400,1440.0200) -- (2559.1700,1441.6900) --
(2559.1700,1442.5200) -- (2560.0000,1444.1900) -- (2560.8400,1445.0200) --
(2560.8400,1446.6900) -- (2561.6700,1447.5200) -- (2562.5000,1449.1900) --
(2562.5000,1450.0200) -- (2563.3400,1451.6900) -- (2564.1700,1452.5200) --
(2564.1700,1454.1900) -- (2565.0000,1455.0200) -- (2565.8400,1456.6900) --
(2566.6700,1457.5200) -- (2566.6700,1459.1900) -- (2567.5000,1460.0200) --
(2568.3400,1461.6900) -- (2568.3400,1462.5200) -- (2569.1700,1464.1900) --
(2570.0000,1465.0200) -- (2570.0000,1465.8600) -- (2570.8400,1467.5200) --
(2571.6700,1468.3600) -- (2571.6700,1470.0200) -- (2572.5000,1470.8600) --
(2573.3400,1472.5200) -- (2573.3400,1473.3600) -- (2574.1700,1475.0200) --
(2575.0000,1475.8600) -- (2575.0000,1477.5200) -- (2575.8400,1478.3600) --
(2576.6700,1480.0200) -- (2577.5000,1480.8600) -- (2577.5000,1482.5200) --
(2578.3400,1483.3600) -- (2579.1700,1485.0200) -- (2579.1700,1485.8600) --
(2580.0000,1487.5200) -- (2580.8400,1488.3600) -- (2580.8400,1490.0200) --
(2581.6700,1490.8600) -- (2582.5000,1492.5200) -- (2582.5000,1493.3600) --
(2583.3400,1495.0200) -- (2584.1700,1495.8600) -- (2584.1700,1496.6900) --
(2585.0000,1498.3600) -- (2585.8400,1499.1900) -- (2585.8400,1500.8600) --
(2586.6700,1501.6900) -- (2587.5000,1503.3600) -- (2588.3400,1504.1900) --
(2588.3400,1505.8600) -- (2589.1700,1506.6900) -- (2590.0000,1508.3600) --
(2590.0000,1509.1900) -- (2590.8400,1510.8600) -- (2591.6700,1511.6900) --
(2591.6700,1513.3600) -- (2592.5000,1514.1900) -- (2593.3400,1515.8600) --
(2593.3400,1516.6900) -- (2594.1700,1518.3600) -- (2595.0000,1519.1900) --
(2595.0000,1520.8600) -- (2595.8400,1521.6900) -- (2596.6700,1523.3600) --
(2597.5000,1524.1900) -- (2597.5000,1525.0200) -- (2598.3400,1526.6900) --
(2599.1700,1527.5200) -- (1795.0000,1527.5200)"""
data_2colon1 = """(3138.3300,1527.5200) -- (3139.1700,1526.6900) --
(3140.0000,1525.0200) -- (3140.8300,1524.1900) -- (3141.6700,1523.3500) --
(3141.6700,1521.6900) -- (3142.5000,1520.8500) -- (3143.3300,1519.1900) --
(3144.1700,1518.3500) -- (3145.0000,1516.6900) -- (3145.8300,1515.8500) --
(3145.8300,1514.1900) -- (3146.6700,1513.3500) -- (3147.5000,1511.6900) --
(3148.3300,1510.8500) -- (3149.1700,1509.1900) -- (3150.0000,1508.3500) --
(3150.0000,1506.6900) -- (3150.8300,1505.8500) -- (3151.6700,1504.1900) --
(3152.5000,1503.3500) -- (3153.3300,1501.6900) -- (3154.1700,1500.8500) --
(3154.1700,1499.1900) -- (3155.0000,1498.3500) -- (3155.8300,1496.6900) --
(3156.6700,1495.8500) -- (3157.5000,1495.0200) -- (3158.3300,1493.3500) --
(3158.3300,1492.5200) -- (3159.1700,1490.8500) -- (3160.0000,1490.0200) --
(3160.8300,1488.3500) -- (3161.6700,1487.5200) -- (3162.5000,1485.8500) --
(3162.5000,1485.0200) -- (3163.3300,1483.3500) -- (3164.1700,1482.5200) --
(3165.0000,1480.8500) -- (3165.8300,1480.0200) -- (3166.6700,1478.3500) --
(3166.6700,1477.5200) -- (3167.5000,1475.8500) -- (3168.3300,1475.0200) --
(3169.1700,1473.3500) -- (3170.0000,1472.5200) -- (3170.0000,1470.8500) --
(3170.8300,1470.0200) -- (3171.6700,1468.3500) -- (3172.5000,1467.5200) --
(3173.3300,1465.8500) -- (3174.1700,1465.0200) -- (3174.1700,1464.1800) --
(3175.0000,1462.5200) -- (3175.8300,1461.6800) -- (3176.6700,1460.0200) --
(3177.5000,1459.1800) -- (3178.3300,1457.5200) -- (3178.3300,1456.6800) --
(3179.1700,1455.0200) -- (3180.0000,1454.1800) -- (3180.8300,1452.5200) --
(3181.6700,1451.6800) -- (3181.6700,1450.0200) -- (3182.5000,1449.1800) --
(3183.3300,1447.5200) -- (3184.1700,1446.6800) -- (3185.0000,1445.0200) --
(3185.8300,1444.1800) -- (3185.8300,1442.5200) -- (3186.6700,1441.6800) --
(3187.5000,1440.0200) -- (3188.3300,1439.1800) -- (3189.1700,1437.5200) --
(3190.0000,1436.6800) -- (3190.0000,1435.8500) -- (3190.8300,1434.1800) --
(3191.6700,1433.3500) -- (3192.5000,1431.6800) -- (3193.3300,1430.8500) --
(3193.3300,1429.1800) -- (3194.1700,1428.3500) -- (3195.0000,1426.6800) --
(3195.8300,1425.8500) -- (3196.6700,1424.1800) -- (3197.5000,1423.3500) --
(3197.5000,1421.6800) -- (3198.3300,1420.8500) -- (3199.1700,1419.1800) --
(3200.0000,1418.3500) -- (3200.8300,1416.6800) -- (3200.8300,1415.8500) --
(3201.6700,1414.1800) -- (3202.5000,1413.3500) -- (3203.3300,1411.6800) --
(3204.1700,1410.8500) -- (3205.0000,1409.1800) -- (3205.0000,1408.3500) --
(3205.8300,1407.5200) -- (3206.6700,1405.8500) -- (3207.5000,1405.0200) --
(3208.3300,1403.3500) -- (3209.1700,1402.5200) -- (3209.1700,1400.8500) --
(3210.0000,1400.0200) -- (3210.8300,1398.3500) -- (3211.6700,1397.5200) --
(3212.5000,1395.8500) -- (3212.5000,1395.0200) -- (3213.3300,1393.3500) --
(3214.1700,1392.5200) -- (3215.0000,1390.8500) -- (3215.8300,1390.0200) --
(3216.6700,1388.3500) -- (3216.6700,1387.5200) -- (3217.5000,1385.8500) --
(3218.3300,1385.0200) -- (3219.1700,1383.3500) -- (3220.0000,1382.5200) --
(3220.0000,1380.8500) -- (3220.8300,1380.0200) -- (3221.6700,1378.3500) --
(3222.5000,1377.5200) -- (3223.3300,1376.6800) -- (3224.1700,1375.0200) --
(3224.1700,1374.1800) -- (3225.0000,1372.5200) -- (3225.8300,1371.6800) --
(3226.6700,1370.0200) -- (3227.5000,1369.1800) -- (3227.5000,1367.5200) --
(3228.3300,1366.6800) -- (3229.1700,1365.0200) -- (3230.0000,1364.1800) --
(3230.8300,1362.5200) -- (3231.6700,1361.6800) -- (3231.6700,1360.0200) --
(3232.5000,1359.1800) -- (3233.3300,1357.5200) -- (3234.1700,1356.6800) --
(3235.0000,1355.0200) -- (3235.0000,1354.1800) -- (3235.8300,1352.5200) --
(3236.6700,1351.6800) -- (3237.5000,1350.0200) -- (3238.3300,1349.1800) --
(3239.1700,1348.3500) -- (3239.1700,1346.6800) -- (3240.0000,1345.8500) --
(3240.8400,1344.1800) -- (3241.6700,1343.3500) -- (3242.5000,1341.6800) --
(3243.3400,1340.8500) -- (3243.3400,1339.1800) -- (3244.1700,1338.3500) --
(3245.0000,1336.6800) -- (3245.8400,1335.8500) -- (3246.6700,1334.1800) --
(3246.6700,1333.3500) -- (3247.5000,1331.6800) -- (3248.3400,1330.8500) --
(3249.1700,1329.1800) -- (3250.0000,1328.3500) -- (3250.8400,1326.6800) --
(3250.8400,1325.8500) -- (3251.6700,1324.1800) -- (3252.5000,1323.3500) --
(3253.3400,1321.6800) -- (3254.1700,1320.8500) -- (3255.0000,1320.0200) --
(3255.0000,1318.3500) -- (3255.8400,1317.5200) -- (3256.6700,1315.8500) --
(3257.5000,1315.0200) -- (3258.3400,1313.3500) -- (3259.1700,1312.5200) --
(3259.1700,1310.8500) -- (3260.0000,1310.0200) -- (3260.8400,1308.3500) --
(3261.6700,1307.5200) -- (3262.5000,1305.8500) -- (3262.5000,1305.0200) --
(3263.3400,1303.3500) -- (3264.1700,1302.5200) -- (3265.0000,1300.8500) --
(3265.8400,1300.0200) -- (3266.6700,1298.3500) -- (3266.6700,1297.5200) --
(3267.5000,1295.8500) -- (3268.3400,1295.0200) -- (3269.1700,1293.3500) --
(3270.0000,1292.5200) -- (3270.8400,1290.8500) -- (3270.8400,1290.0200) --
(3271.6700,1289.1800) -- (3272.5000,1287.5200) -- (3273.3400,1286.6800) --
(3274.1700,1285.0200) -- (3275.0000,1284.1800) -- (3275.8400,1282.5200) --
(3275.8400,1281.6800) -- (3276.6700,1280.0200) -- (3277.5000,1279.1800) --
(3278.3400,1277.5200) -- (3279.1700,1276.6800) -- (3280.0000,1275.0200) --
(3280.0000,1274.1800) -- (3280.8300,1272.5200) --
(3281.6700,1271.6900) -- (3282.5000,1270.0200) -- (3283.3300,1269.1900) --
(3284.1700,1267.5200) -- (3284.1700,1266.6900) -- (3285.0000,1265.0200) --
(3285.8300,1264.1900) -- (3286.6700,1262.5200) -- (3287.5000,1261.6900) --
(3288.3300,1260.8500) -- (3289.1700,1259.1900) -- (3289.1700,1258.3500) --
(3290.0000,1256.6900) -- (3290.8300,1255.8500) -- (3291.6700,1254.1900) --
(3292.5000,1253.3500) -- (3293.3300,1251.6900) -- (3294.1700,1250.8500) --
(3294.1700,1249.1900) -- (3295.0000,1248.3500) -- (3295.8300,1246.6900) --
(3296.6700,1245.8500) -- (3297.5000,1244.1900) -- (3297.5000,1243.3500) --
(3298.3300,1241.6900) -- (3299.1700,1240.8500) -- (3299.1700,1239.1900) --
(3300.0000,1238.3500) -- (3300.0000,1236.6900) -- (3300.8300,1235.8500) --
(3300.8300,1234.1900) -- (3301.6700,1233.3500) -- (3301.6700,1231.6900) --
(3302.5000,1230.8500) -- (3303.3300,1230.0200) -- (3303.3300,1228.3500) --
(3304.1700,1227.5200) -- (3304.1700,1225.8500) -- (3305.0000,1225.0200) --
(3305.0000,1223.3500) -- (3305.8300,1222.5200) -- (3305.8300,1220.8500) --
(3306.6700,1220.0200) -- (3306.6700,1218.3500) -- (3307.5000,1217.5200) --
(3307.5000,1215.8500) -- (3308.3300,1215.0200) -- (3308.3300,1213.3500) --
(3309.1700,1212.5200) -- (3309.1700,1210.8500) -- (3310.0000,1210.0200) --
(3310.8300,1208.3500) -- (3310.8300,1207.5200) -- (3311.6700,1205.8500) --
(3311.6700,1205.0200) -- (3312.5000,1203.3500) -- (3312.5000,1202.5200) --
(3313.3300,1201.6900) -- (3313.3300,1200.0200) -- (3314.1700,1199.1900) --
(3314.1700,1197.5200) -- (3315.0000,1196.6900) -- (3315.0000,1195.0200) --
(3315.8300,1194.1900) -- (3315.8300,1192.5200) -- (3316.6700,1191.6900) --
(3316.6700,1190.0200) -- (3317.5000,1189.1900) -- (3317.5000,1187.5200) --
(3318.3300,1186.6900) -- (3318.3300,1185.0200) -- (3319.1700,1184.1900) --
(3319.1700,1181.6900) -- (3320.0000,1180.0200) -- (3320.0000,1179.1900) --
(3320.8300,1177.5200) -- (3320.8300,1176.6900) -- (3321.6700,1175.0200) --
(3321.6700,1174.1900) -- (3322.5000,1173.3500) -- (3322.5000,1171.6900) --
(3323.3300,1170.8500) -- (3323.3300,1169.1900) -- (3324.1700,1168.3500) --
(3324.1700,1166.6900) -- (3325.0000,1165.8500) -- (3325.0000,1164.1900) --
(3325.8300,1163.3500) -- (3325.8300,1161.6900) -- (3326.6700,1160.8500) --
(3326.6700,1159.1900) -- (3327.5000,1158.3500) -- (3327.5000,1155.8500) --
(3328.3300,1154.1900) -- (3328.3300,1153.3500) -- (3329.1700,1151.6900) --
(3329.1700,1150.8500) -- (3330.0000,1149.1900) -- (3330.0000,1148.3500) --
(3330.8300,1146.6900) -- (3330.8300,1145.8500) -- (3331.6700,1144.1900) --
(3331.6700,1143.3500) -- (3332.5000,1142.5200) -- (3332.5000,1140.0200) --
(3333.3300,1138.3500) -- (3333.3300,1137.5200) -- (3334.1700,1135.8500) --
(3334.1700,1135.0200) -- (3335.0000,1133.3500) -- (3335.0000,1132.5200) --
(3335.8300,1130.8500) -- (3335.8300,1128.3500) -- (3336.6700,1127.5200) --
(3336.6700,1125.8500) -- (3337.5000,1125.0200) -- (3337.5000,1123.3500) --
(3338.3300,1122.5200) -- (3338.3300,1120.8500) -- (3339.1700,1120.0200) --
(3339.1700,1117.5200) -- (3340.0000,1115.8500) -- (3340.0000,1115.0200) --
(3340.8300,1114.1900) -- (3340.8300,1112.5200) -- (3341.6700,1111.6900) --
(3341.6700,1109.1900) -- (3342.5000,1107.5200) -- (3342.5000,1106.6900) --
(3343.3300,1105.0200) -- (3343.3300,1104.1900) -- (3344.1700,1102.5200) --
(3344.1700,1100.0200) -- (3345.0000,1099.1900) -- (3345.0000,1097.5200) --
(3345.8300,1096.6900) -- (3345.8300,1095.0200) -- (3346.6700,1094.1900) --
(3346.6700,1091.6900) -- (3347.5000,1090.0200) -- (3347.5000,1089.1900) --
(3348.3300,1087.5200) -- (3348.3300,1085.8500) -- (3349.1700,1084.1900) --
(3349.1700,1083.3500) -- (3350.0000,1081.6900) -- (3350.0000,1079.1900) --
(3350.8300,1078.3500) -- (3350.8300,1076.6900) -- (3351.6700,1075.8500) --
(3351.6700,1074.1900) -- (3352.5000,1073.3500) -- (3352.5000,1070.8500) --
(3353.3300,1069.1900) -- (3353.3300,1068.3500) -- (3354.1700,1066.6900) --
(3354.1700,1064.1900) -- (3355.0000,1063.3500) -- (3355.0000,1061.6900) --
(3355.8300,1060.8500) -- (3355.8300,1058.3500) -- (3356.6700,1056.6900) --
(3356.6700,1055.8500) -- (3357.5000,1055.0200) -- (3357.5000,1052.5200) --
(3358.3300,1050.8500) -- (3358.3300,1050.0200) -- (3359.1700,1048.3500) --
(3359.1700,1045.8500) -- (3360.0000,1045.0200) -- (3360.0000,1042.5200) --
(3360.8300,1040.8500) -- (3360.8300,1040.0200) -- (3361.6700,1038.3500) --
(3361.6700,1035.8500) -- (3362.5000,1035.0200) -- (3362.5000,1033.3500) --
(3363.3300,1032.5200) -- (3363.3300,1030.0200) -- (3364.1700,1028.3500) --
(3364.1700,1026.6900) -- (3365.0000,1025.0200) -- (3365.0000,1024.1900) --
(3365.8300,1022.5200) -- (3365.8300,1020.0200) -- (3366.6700,1019.1900) --
(3366.6700,1016.6900) -- (3367.5000,1015.0200) -- (3367.5000,1014.1900) --
(3368.3300,1012.5200) -- (3368.3300,1010.0200) -- (3369.1700,1009.1900) --
(3369.1700,1006.6900) -- (3370.0000,1005.0200) -- (3370.0000,1004.1900) --
(3370.8300,1002.5200) -- (3370.8300,1000.0200) -- (3371.6700,999.1880) --
(3371.6700,996.6880) -- (3372.5000,995.8520) -- (3372.5000,993.3520) --
(3373.3300,991.6880) -- (3373.3300,989.1880) -- (3374.1700,988.3520) --
(3374.1700,986.6880) -- (3375.0000,985.8520) -- (3375.0000,983.3520) --
(3375.8400,981.6880) -- (3375.8400,979.1880) -- (3376.6700,978.3520) --
(3376.6700,975.8520) -- (3377.5000,974.1880) -- (3377.5000,971.6880) --
(3378.3400,970.8520) -- (3378.3400,968.3520) -- (3379.1700,967.5200) --
(3379.1700,965.8520) -- (3380.0000,965.0200) -- (3380.0000,962.5200) --
(3380.8400,960.8520) -- (3380.8400,958.3520) -- (3381.6700,957.5200) --
(3381.6700,955.0200) -- (3382.5000,953.3520) -- (3382.5000,950.8520) --
(3383.3400,950.0200) -- (3383.3400,947.5200) -- (3384.1700,945.8520) --
(3384.1700,943.3520) -- (3385.0000,942.5200) -- (3385.0000,940.0200) --
(3385.8400,939.1880) -- (3385.8400,936.6880) -- (3386.6700,935.0200) --
(3386.6700,932.5200) -- (3387.5000,931.6880) -- (3387.5000,929.1880) --
(3388.3400,927.5200) -- (3388.3400,925.0200) -- (3389.1700,924.1880) --
(3389.1700,920.0200) -- (3390.0000,919.1880) -- (3390.0000,916.6880) --
(3390.8400,915.0200) -- (3390.8400,912.5200) -- (3391.6700,911.6880) --
(3391.6700,909.1880) -- (3392.5000,908.3520) --
(3392.5000,905.8520) -- (3393.3300,904.1880) --
(3393.3300,901.6880) -- (3394.1700,900.8520) -- (3394.1700,896.6880) --
(3395.0000,895.8520) -- (3395.0000,893.3520) -- (3395.8300,891.6880) --
(3395.8300,889.1880) -- (3396.6700,888.3520) -- (3396.6700,885.8520) --
(3397.5000,884.1880) -- (3397.5000,880.8520) -- (3398.3300,880.0200) --
(3398.3300,877.5200) -- (3399.1700,875.8520) -- (3399.1700,873.3520) --
(3400.0000,872.5200) -- (3400.0000,868.3520) -- (3400.8300,867.5200) --
(3400.8300,865.0200) -- (3401.6700,863.3520) -- (3401.6700,860.0200) --
(3402.5000,858.3520) -- (3402.5000,855.8520) -- (3403.3300,855.0200) --
(3403.3300,851.6880) -- (3404.1700,850.0200) -- (3404.1700,847.5200) --
(3405.0000,846.6880) -- (3405.0000,842.5200) -- (3405.8300,841.6880) --
(3405.8300,839.1880) -- (3406.6700,837.5200) -- (3406.6700,834.1880) --
(3407.5000,832.5200) -- (3407.5000,829.1880) -- (3408.3300,827.5200) --
(3408.3300,825.0200) -- (3409.1700,824.1880) -- (3409.1700,820.8520) --
(3410.0000,819.1880) -- (3410.0000,815.8520) -- (3410.8300,814.1880) --
(3410.8300,811.6880) -- (3411.6700,810.8520) -- (3411.6700,806.6880) --
(3412.5000,805.8520) -- (3412.5000,801.6880) -- (3413.3300,800.8520) --
(3413.3300,796.6880) -- (3414.1700,795.8520) -- (3414.1700,792.5200) --
(3415.0000,790.8520) -- (3415.0000,787.5200) -- (3415.8300,785.8520) --
(3415.8300,782.5200) -- (3416.6700,780.8520) -- (3416.6700,777.5200) --
(3417.5000,775.8520) -- (3417.5000,772.5200) -- (3418.3300,770.8520) --
(3418.3300,767.5200) -- (3419.1700,765.8520) -- (3419.1700,762.5200) --
(3420.0000,761.6880) -- (3420.0000,757.5200) -- (3420.8300,756.6880) --
(3420.8300,751.6880) -- (3421.6700,750.0200) -- (3421.6700,746.6880) --
(3422.5000,745.0200) -- (3422.5000,741.6880) -- (3423.3300,740.0200) --
(3423.3300,735.0200) -- (3424.1700,734.1880) -- (3424.1700,730.8520) --
(3425.0000,729.1880) -- (3425.0000,725.8520) -- (3425.8300,724.1880) --
(3425.8300,719.1880) -- (3426.6700,718.3520) -- (3426.6700,713.3520) --
(3427.5000,711.6880) -- (3427.5000,708.3520) -- (3428.3300,706.6880) --
(3428.3300,702.5200) -- (3429.1700,700.8520) -- (3429.1700,695.8520) --
(3430.0000,695.0200) -- (3430.0000,690.0200) -- (3430.8300,688.3520) --
(3430.8300,683.3520) -- (3431.6700,682.5200) -- (3431.6700,677.5200) --
(3432.5000,675.8520) -- (3432.5000,671.6880) -- (3433.3300,670.0200) --
(3433.3300,665.0200) -- (3434.1700,664.1880) -- (3434.1700,659.1880) --
(3435.0000,657.5200) -- (3435.0000,651.6880) -- (3435.8300,650.0200) --
(3435.8300,645.8520) -- (3436.6700,644.1880) -- (3436.6700,638.3520) --
(3437.5000,636.6880) -- (3437.5000,630.8520) -- (3438.3300,629.1880) --
(3438.3300,623.3520) -- (3439.1700,621.6880) -- (3439.1700,617.5200) --
(3440.0000,615.8520) -- (3440.0000,608.3550) -- (3440.8300,607.5200) --
(3440.8300,600.8550) -- (3441.6700,600.0200) -- (3441.6700,593.3550) --
(3442.5000,592.5200) -- (3442.5000,585.0200) -- (3443.3300,584.1910) --
(3443.3300,576.6910) -- (3444.1700,575.0200) -- (3444.1700,567.5200) --
(3445.0000,566.6910) -- (3445.0000,559.1910) -- (3445.8300,558.3520) --
(3445.8300,550.8520) -- (3446.6700,549.1910) -- (3446.6700,540.8520) --
(3447.5000,539.1910) -- (3447.5000,536.6910) --
(3447.5000,530.8520) -- (3448.3300,529.1910) -- (3448.3300,520.8520) --
(3449.1700,520.0200) -- (3449.1700,510.0200) -- (3450.0000,508.3520) --
(3450.0000,499.1910) -- (3450.8300,497.5200) -- (3450.8300,486.6910) --
(3451.6700,485.0200) -- (3451.6700,472.5200) -- (3452.5000,471.6910) --
(3452.5000,459.1910) -- (3453.3300,458.3520) -- (3453.3300,444.1910) --
(3454.1700,443.3520) -- (3454.1700,427.5200) -- (3455.0000,425.8520) --
(3455.0000,407.5200) -- (3455.8300,406.6910) -- (3455.8300,384.1910) --
(3456.6700,383.3520) -- (3456.6700,350.8520) -- (3457.5000,350.0200) --
(3457.5000,296.6910) -- (3457.5000,337.5200) -- (3458.3300,338.3520) --
(3458.3300,375.8520) -- (3459.1700,376.6910) -- (3459.1700,400.0200) --
(3460.0000,401.6910) -- (3460.0000,420.8520) -- (3460.8300,422.5200) --
(3460.8300,423.3520) -- (3460.8300,438.3520) --
(3461.6700,440.0200) -- (3461.6700,454.1910) -- (3462.5000,455.8520) --
(3462.5000,468.3520) -- (3463.3300,469.1910) -- (3463.3300,481.6910) --
(3464.1700,482.5200) -- (3464.1700,494.1910) -- (3465.0000,495.0200) --
(3465.0000,505.0200) -- (3465.8300,505.8520) -- (3465.8300,515.8520) --
(3466.6700,517.5200) -- (3466.6700,525.8520) -- (3467.5000,527.5200) --
(3467.5000,536.6910) -- (3468.3300,538.3520) -- (3468.3300,545.8520) --
(3469.1700,546.6910) -- (3469.1700,555.8520) -- (3470.0000,556.6910) --
(3470.0000,564.1910) -- (3470.8300,565.0200) -- (3470.8300,572.5200) --
(3471.6700,574.1910) -- (3471.6700,581.6910) -- (3472.5000,582.5200) --
(3472.5000,588.3520) -- (3473.3300,590.0200) -- (3473.3300,597.5200) --
(3474.1700,598.3520) -- (3474.1700,605.0200) -- (3475.0000,605.8520) --
(3475.0000,612.5200) -- (3475.8300,613.3520) -- (3475.8300,619.1880) --
(3476.6700,620.8520) -- (3476.6700,626.6880) -- (3477.5000,628.3520) --
(3477.5000,633.3520) -- (3478.3300,634.1880) -- (3478.3300,640.8520) --
(3479.1700,641.6880) -- (3479.1700,646.6880) -- (3480.0000,647.5200) --
(3480.0000,652.5200) -- (3480.8300,654.1880) -- (3480.8300,660.0200) --
(3481.6700,661.6880) -- (3481.6700,666.6880) -- (3482.5000,667.5200) --
(3482.5000,672.5200) -- (3483.3300,674.1880) -- (3483.3300,678.3520) --
(3484.1700,680.0200) -- (3484.1700,685.0200) -- (3485.0000,685.8520) --
(3485.0000,690.0200) -- (3485.8300,690.8520) -- (3485.8300,695.8520) --
(3486.6700,697.5200) -- (3486.6700,702.5200) -- (3487.5000,703.3520) --
(3487.5000,706.6880) -- (3488.3300,708.3520) -- (3488.3300,713.3520) --
(3489.1700,714.1880) -- (3489.1700,718.3520) -- (3490.0000,719.1880) --
(3490.0000,724.1880) -- (3490.8300,725.8520) -- (3490.8300,729.1880) --
(3491.6700,730.8520) -- (3491.6700,734.1880) -- (3492.5000,735.0200) --
(3492.5000,740.0200) -- (3493.3300,741.6880) -- (3493.3300,745.0200) --
(3494.1700,746.6880) -- (3494.1700,750.0200) -- (3495.0000,751.6880) --
(3495.0000,755.0200) -- (3495.8300,756.6880) -- (3495.8300,760.0200) --
(3496.6700,761.6880) -- (3496.6700,765.0200) -- (3497.5000,765.8520) --
(3497.5000,770.0200) -- (3498.3300,770.8520) -- (3498.3300,775.0200) --
(3499.1700,775.8520) -- (3499.1700,780.0200) -- (3500.0000,780.8520) --
(3500.0000,785.0200) -- (3500.8300,785.8520) -- (3500.8300,788.3520) --
(3501.6700,790.0200) -- (3501.6700,792.5200) --
(3501.6700,793.3520) -- (3502.5000,794.1880) -- (3502.5000,798.3520) --
(3503.3300,799.1880) -- (3503.3300,803.3520) -- (3504.1700,804.1880) --
(3504.1700,806.6880) -- (3505.0000,808.3520) -- (3505.0000,811.6880) --
(3505.8300,813.3520) -- (3505.8300,815.8520) -- (3506.6700,816.6880) --
(3506.6700,820.8520) -- (3507.5000,821.6880) -- (3507.5000,824.1880) --
(3508.3300,825.0200) -- (3508.3300,829.1880) -- (3509.1700,830.0200) --
(3509.1700,832.5200) -- (3510.0000,834.1880) -- (3510.0000,837.5200) --
(3510.8300,839.1880) -- (3510.8300,841.6880) -- (3511.6700,842.5200) --
(3511.6700,845.0200) -- (3512.5000,846.6880) -- (3512.5000,850.0200) --
(3513.3300,851.6880) -- (3513.3300,853.3520) -- (3514.1700,855.0200) --
(3514.1700,857.5200) -- (3515.0000,858.3520) -- (3515.0000,862.5200) --
(3515.8300,863.3520) -- (3515.8300,865.8520) -- (3516.6700,867.5200) --
(3516.6700,870.0200) -- (3517.5000,870.8520) -- (3517.5000,873.3520) --
(3518.3300,875.0200) -- (3518.3300,877.5200) -- (3519.1700,878.3520) --
(3519.1700,881.6880) -- (3520.0000,883.3520) -- (3520.0000,885.8520) --
(3520.8300,886.6880) -- (3520.8300,889.1880) -- (3521.6700,890.8520) --
(3521.6700,893.3520) -- (3522.5000,894.1880) -- (3522.5000,896.6880) --
(3523.3300,898.3520) -- (3523.3300,900.8520) -- (3524.1700,901.6880) --
(3524.1700,904.1880) -- (3525.0000,905.8520) -- (3525.0000,908.3520) --
(3525.8300,909.1880) -- (3525.8300,911.6880) -- (3526.6700,912.5200) --
(3526.6700,915.0200) -- (3527.5000,916.6880) -- (3527.5000,919.1880) --
(3528.3300,920.0200) -- (3528.3300,921.6880) -- (3529.1700,922.5200) --
(3529.1700,925.0200) -- (3530.0000,926.6880) -- (3530.0000,929.1880) --
(3530.8300,930.0200) -- (3530.8300,932.5200) -- (3531.6700,934.1880) --
(3531.6700,936.6880) -- (3532.5000,937.5200) -- (3532.5000,940.0200) --
(3533.3300,940.8520) -- (3533.3300,942.5200) -- (3534.1700,943.3520) --
(3534.1700,945.8520) -- (3535.0000,947.5200) -- (3535.0000,950.0200) --
(3535.8300,950.8520) -- (3535.8300,953.3520) -- (3536.6700,955.0200) --
(3536.6700,955.8520) -- (3537.5000,957.5200) -- (3537.5000,960.0200) --
(3538.3300,960.8520) -- (3538.3300,963.3520) -- (3539.1700,965.0200) --
(3539.1700,965.8520) -- (3540.0000,967.5200) -- (3540.0000,969.1880) --
(3540.8300,970.8520) -- (3540.8300,973.3520) -- (3541.6700,974.1880) --
(3541.6700,975.8520) -- (3542.5000,976.6880) -- (3542.5000,979.1880) --
(3543.3300,980.8520) -- (3543.3300,981.6880) -- (3544.1700,983.3520) --
(3544.1700,985.8520) -- (3545.0000,986.6880) -- (3545.0000,989.1880) --
(3545.8300,990.8520) -- (3545.8300,991.6880) -- (3546.6700,993.3520) --
(3546.6700,995.8520) -- (3547.5000,996.6880) -- (3547.5000,997.5200) --
(3548.3300,999.1880) -- (3548.3300,1001.6900) -- (3549.1700,1002.5200) --
(3549.1700,1004.1900) -- (3550.0000,1005.0200) -- (3550.0000,1007.5200) --
(3550.8300,1009.1900) -- (3550.8300,1010.0200) -- (3551.6700,1011.6900) --
(3551.6700,1014.1900) -- (3552.5000,1015.0200) -- (3552.5000,1016.6900) --
(3553.3300,1017.5200) -- (3553.3300,1019.1900) -- (3554.1700,1020.0200) --
(3554.1700,1022.5200) -- (3555.0000,1024.1900) -- (3555.0000,1025.0200) --
(3555.8300,1026.6900) -- (3555.8300,1028.3500) -- (3556.6700,1030.0200) --
(3556.6700,1030.8500) -- (3557.5000,1032.5200) -- (3557.5000,1033.3500) --
(3558.3300,1035.0200) -- (3558.3300,1037.5200) -- (3559.1700,1038.3500) --
(3559.1700,1040.0200) -- (3560.0000,1040.8500) -- (3560.0000,1042.5200) --
(3560.8300,1043.3500) -- (3560.8300,1045.8500) -- (3561.6700,1047.5200) --
(3561.6700,1048.3500) -- (3562.5000,1050.0200) -- (3562.5000,1050.8500) --
(3563.3300,1052.5200) -- (3563.3300,1053.3500) -- (3564.1700,1055.0200) --
(3564.1700,1056.6900) -- (3565.0000,1058.3500) -- (3565.0000,1059.1900) --
(3565.8300,1060.8500) -- (3565.8300,1061.6900) -- (3566.6700,1063.3500) --
(3566.6700,1064.1900) -- (3567.5000,1065.8500) -- (3567.5000,1068.3500) --
(3568.3300,1069.1900) -- (3568.3300,1070.8500) -- (3569.1700,1071.6900) --
(3569.1700,1073.3500) -- (3570.0000,1074.1900) -- (3570.0000,1075.8500) --
(3570.8300,1076.6900) -- (3570.8300,1078.3500) -- (3571.6700,1079.1900) --
(3571.6700,1081.6900) -- (3572.5000,1083.3500) -- (3572.5000,1084.1900) --
(3573.3300,1085.8500) -- (3573.3300,1086.6900) -- (3574.1700,1087.5200) --
(3574.1700,1089.1900) -- (3575.0000,1090.0200) -- (3575.0000,1091.6900) --
(3575.8300,1092.5200) -- (3575.8300,1094.1900) -- (3576.6700,1095.0200) --
(3576.6700,1096.6900) -- (3577.5000,1097.5200) -- (3577.5000,1099.1900) --
(3578.3300,1100.0200) -- (3578.3300,1101.6900) -- (3579.1700,1102.5200) --
(3579.1700,1105.0200) -- (3580.0000,1106.6900) -- (3580.0000,1107.5200) --
(3580.8300,1109.1900) -- (3580.8300,1110.0200) -- (3581.6700,1111.6900) --
(3581.6700,1112.5200) -- (3582.5000,1114.1900) -- (3582.5000,1115.0200) --
(3583.3300,1115.8500) -- (3583.3300,1117.5200) -- (3584.1700,1118.3500) --
(3584.1700,1120.0200) -- (3585.0000,1120.8500) -- (3585.0000,1122.5200) --
(3585.8300,1123.3500) -- (3585.8300,1125.0200) -- (3586.6700,1125.8500) --
(3586.6700,1127.5200) -- (3587.5000,1128.3500) -- (3587.5000,1130.0200) --
(3588.3300,1130.8500) -- (3588.3300,1132.5200) -- (3589.1700,1133.3500) --
(3589.1700,1135.0200) -- (3590.0000,1135.8500) -- (3590.0000,1137.5200) --
(3590.8300,1138.3500) -- (3590.8300,1140.0200) -- (3591.6700,1140.8500) --
(3592.5000,1142.5200) -- (3592.5000,1143.3500) -- (3593.3300,1144.1900) --
(3593.3300,1145.8500) -- (3594.1700,1146.6900) -- (3594.1700,1148.3500) --
(3595.0000,1149.1900) -- (3595.0000,1150.8500) -- (3595.8300,1151.6900) --
(3595.8300,1153.3500) -- (3596.6700,1154.1900) -- (3596.6700,1155.8500) --
(3597.5000,1156.6900) -- (3597.5000,1158.3500) -- (3598.3300,1159.1900) --
(3598.3300,1160.8500) -- (3599.1700,1161.6900) --
(3600.0000,1163.3500) -- (3600.0000,1164.1900) -- (3600.8300,1165.8500) --
(3600.8300,1166.6900) -- (3601.6700,1168.3500) -- (3601.6700,1169.1900) --
(3602.5000,1170.8500) -- (3602.5000,1171.6900) -- (3603.3300,1173.3500) --
(3604.1700,1174.1900) -- (3604.1700,1175.0200) -- (3605.0000,1176.6900) --
(3605.0000,1177.5200) -- (3605.8300,1179.1900) -- (3605.8300,1180.0200) --
(3606.6700,1181.6900) -- (3606.6700,1182.5200) -- (3607.5000,1184.1900) --
(3608.3300,1185.0200) -- (3608.3300,1186.6900) -- (3609.1700,1187.5200) --
(3609.1700,1189.1900) -- (3610.0000,1190.0200) -- (3610.0000,1191.6900) --
(3610.8300,1192.5200) -- (3611.6700,1194.1900) -- (3611.6700,1195.0200) --
(3612.5000,1196.6900) -- (3612.5000,1197.5200) -- (3613.3300,1199.1900) --
(3614.1700,1200.0200) -- (3614.1700,1201.6900) -- (3615.0000,1202.5200) --
(3615.0000,1203.3500) -- (3615.8300,1205.0200) -- (3616.6700,1205.8500) --
(3616.6700,1207.5200) -- (3617.5000,1208.3500) -- (3617.5000,1210.0200) --
(3618.3300,1210.8500) -- (3619.1700,1212.5200) -- (3619.1700,1213.3600) --
(3620.0000,1215.0200) -- (3620.0000,1215.8500) -- (3620.8300,1217.5200) --
(3621.6700,1218.3600) -- (3621.6700,1220.0200) -- (3622.5000,1220.8600) --
(3623.3300,1222.5200) -- (3623.3300,1223.3600) -- (3624.1700,1225.0200) --
(3624.1700,1225.8600) -- (3625.0000,1227.5200) -- (3625.8300,1228.3600) --
(3625.8300,1230.0200) -- (3626.6700,1230.8600) -- (3627.5000,1231.6900) --
(3627.5000,1233.3600) -- (3628.3300,1234.1900) -- (3629.1700,1235.8600) --
(3629.1700,1236.6900) -- (3630.0000,1238.3600) -- (3630.8300,1239.1900) --
(3630.8300,1240.8600) -- (3631.6700,1241.6900) -- (3632.5000,1243.3600) --
(3632.5000,1244.1900) -- (3633.3300,1245.8600) -- (3634.1700,1246.6900) --
(3635.0000,1248.3600) -- (3635.8300,1249.1900) -- (3636.6700,1250.8600) --
(3638.3300,1251.6900) -- (3639.1700,1253.3600) -- (3640.0000,1254.1900) --
(3640.8300,1255.8600) -- (3641.6700,1256.6900) -- (3642.5000,1258.3600) --
(3643.3300,1259.1900) -- (3644.1700,1260.8600) -- (3645.0000,1261.6900) --
(3645.8300,1262.5200) -- (3646.6700,1264.1900) -- (3647.5000,1265.0200) --
(3648.3300,1266.6900) -- (3649.1700,1267.5200) -- (3650.0000,1269.1900) --
(3650.8300,1270.0200) -- (3651.6700,1271.6900) -- (3652.5000,1272.5200) --
(3653.3300,1274.1900) -- (3654.1700,1275.0200) -- (3655.0000,1276.6900) --
(3655.8300,1277.5200) -- (3656.6700,1279.1900) -- (3657.5000,1280.0200) --
(3658.3300,1281.6900) -- (3659.1700,1282.5200) -- (3660.0000,1284.1900) --
(3660.8300,1285.0200) -- (3661.6700,1286.6900) -- (3662.5000,1287.5200) --
(3663.3300,1289.1900) -- (3664.1700,1290.0200) -- (3665.0000,1290.8600) --
(3665.8300,1292.5200) -- (3666.6700,1293.3600) -- (3668.3300,1295.0200) --
(3669.1700,1295.8600) -- (3670.0000,1297.5200) -- (3670.8300,1298.3600) --
(3671.6700,1300.0200) -- (3672.5000,1300.8600) -- (3673.3300,1302.5200) --
(3674.1700,1303.3600) -- (3675.0000,1305.0200) -- (3675.8300,1305.8600) --
(3676.6700,1307.5200) -- (3677.5000,1308.3600) -- (3678.3300,1310.0200) --
(3679.1700,1310.8600) -- (3680.0000,1312.5200) -- (3680.8300,1313.3600) --
(3681.6700,1315.0200) -- (3682.5000,1315.8600) -- (3683.3300,1317.5200) --
(3684.1700,1318.3600) -- (3685.0000,1320.0200) -- (3685.8300,1320.8600) --
(3686.6700,1321.6900) -- (3687.5000,1323.3600) -- (3688.3300,1324.1900) --
(3689.1700,1325.8600) -- (3690.0000,1326.6900) -- (3690.8300,1328.3600) --
(3692.5000,1329.1900) -- (3693.3400,1330.8600) -- (3694.1700,1331.6900) --
(3695.0000,1333.3600) -- (3695.8400,1334.1900) -- (3696.6700,1335.8600) --
(3697.5000,1336.6900) -- (3698.3400,1338.3600) -- (3699.1700,1339.1900) --
(3700.0000,1340.8600) -- (3700.8400,1341.6900) -- (3701.6700,1343.3600) --
(3702.5000,1344.1900) -- (3703.3400,1345.8600) -- (3704.1700,1346.6900) --
(3705.0000,1348.3600) -- (3705.8400,1349.1900) -- (3706.6700,1350.0200) --
(3707.5000,1351.6900) -- (3708.3400,1352.5200) -- (3710.0000,1354.1900) --
(3710.8400,1355.0200) -- (3711.6700,1356.6900) -- (3712.5000,1357.5200) --
(3713.3400,1359.1900) -- (3714.1700,1360.0200) -- (3715.0000,1361.6900) --
(3715.8400,1362.5200) -- (3716.6700,1364.1900) -- (3717.5000,1365.0200) --
(3718.3400,1366.6900) -- (3719.1700,1367.5200) -- (3720.0000,1369.1900) --
(3720.8400,1370.0200) -- (3721.6700,1371.6900) -- (3722.5000,1372.5200) --
(3724.1700,1374.1900) -- (3725.0000,1375.0200) -- (3725.8400,1376.6900) --
(3726.6700,1377.5200) -- (3727.5000,1378.3600) -- (3728.3400,1380.0200) --
(3729.1700,1380.8600) -- (3730.0000,1382.5200) -- (3730.8400,1383.3600) --
(3731.6700,1385.0200) -- (3732.5000,1385.8600) -- (3733.3400,1387.5200) --
(3734.1700,1388.3600) -- (3735.8400,1390.0200) -- (3736.6700,1390.8600) --
(3737.5000,1392.5200) -- (3738.3400,1393.3600) -- (3739.1700,1395.0200) --
(3740.0000,1395.8600) -- (3740.8400,1397.5200) -- (3741.6700,1398.3600) --
(3742.5000,1400.0200) -- (3743.3400,1400.8600) -- (3744.1700,1402.5200) --
(3745.0000,1403.3600) -- (3746.6700,1405.0200) -- (3747.5000,1405.8600) --
(3748.3400,1407.5200) -- (3749.1700,1408.3600) -- (3750.0000,1409.1900) --
(3750.8400,1410.8600) -- (3751.6700,1411.6900) -- (3752.5000,1413.3600) --
(3753.3400,1414.1900) -- (3754.1700,1415.8600) -- (3755.8400,1416.6900) --
(3756.6700,1418.3600) -- (3757.5000,1419.1900) -- (3758.3400,1420.8600) --
(3759.1700,1421.6900) -- (3760.0000,1423.3600) -- (3760.8400,1424.1900) --
(3761.6700,1425.8600) -- (3762.5000,1426.6900) -- (3763.3400,1428.3600) --
(3765.0000,1429.1900) -- (3765.8400,1430.8600) -- (3766.6700,1431.6900) --
(3767.5000,1433.3600) -- (3768.3400,1434.1900) -- (3769.1700,1435.8600) --
(3770.0000,1436.6900) -- (3770.8400,1437.5200) -- (3772.5000,1439.1900) --
(3773.3400,1440.0200) -- (3774.1700,1441.6900) -- (3775.0000,1442.5200) --
(3775.8400,1444.1900) -- (3776.6700,1445.0200) -- (3777.5000,1446.6900) --
(3778.3400,1447.5200) -- (3780.0000,1449.1900) -- (3780.8400,1450.0200) --
(3781.6700,1451.6900) -- (3782.5000,1452.5200) -- (3783.3400,1454.1900) --
(3784.1700,1455.0200) -- (3785.0000,1456.6900) -- (3785.8400,1457.5200) --
(3787.5000,1459.1900) -- (3788.3400,1460.0200) -- (3789.1700,1461.6900) --
(3790.0000,1462.5200) -- (3790.8400,1464.1900) -- (3791.6700,1465.0200) --
(3792.5000,1465.8600) -- (3794.1700,1467.5200) -- (3795.0000,1468.3600) --
(3795.8400,1470.0200) -- (3796.6700,1470.8600) -- (3797.5000,1472.5200) --
(3798.3400,1473.3600) -- (3800.0000,1475.0200) -- (3800.8400,1475.8600) --
(3801.6700,1477.5200) -- (3802.5000,1478.3600) -- (3803.3400,1480.0200) --
(3804.1700,1480.8600) -- (3805.8400,1482.5200) -- (3806.6700,1483.3600) --
(3807.5000,1485.0200) -- (3808.3400,1485.8600) -- (3809.1700,1487.5200) --
(3810.0000,1488.3600) -- (3811.6700,1490.0200) -- (3812.5000,1490.8600) --
(3813.3400,1492.5200) -- (3814.1700,1493.3600) -- (3815.0000,1495.0200) --
(3815.8400,1495.8600) -- (3817.5000,1496.6900) -- (3818.3400,1498.3600) --
(3819.1700,1499.1900) -- (3820.0000,1500.8600) -- (3820.8400,1501.6900) --
(3821.6700,1503.3600) -- (3823.3400,1504.1900) -- (3824.1700,1505.8600) --
(3825.0000,1506.6900) -- (3825.8400,1508.3600) -- (3826.6700,1509.1900) --
(3828.3400,1510.8600) -- (3829.1700,1511.6900) -- (3830.0000,1513.3600) --
(3830.8400,1514.1900) -- (3831.6700,1515.8600) -- (3833.3400,1516.6900) --
(3834.1700,1518.3600) -- (3835.0000,1519.1900) -- (3835.8400,1520.8600) --
(3836.6700,1521.6900) -- (3838.3400,1523.3600) -- (3839.1700,1524.1900) --
(3840.0000,1525.0200) -- (3840.8400,1526.6900) -- (3841.6700,1527.5200) --
(3138.3400,1527.5200)"""
# data = data_1colon2
# tongue_pos = 1.0/2.0
# data = data_2colon3
# tongue_pos = 2.0/3.0
# data = data_1colon1
# tongue_pos = 1.0/1.0
# data = data_2colon1
# tongue_pos = 2.0/1.0
f, r = [], []
for d in data.split('--'):
# print d
x, y = d.strip().replace('(','').replace(')','').split(',')
# print float(x), float(y)
f.append(float(x))
r.append(float(y))
f = np.array(f)
r = np.array(r)
min_f, max_f = np.min(f), np.max(f)
min_r, max_r = np.min(r), np.max(r)
tongue_f_index = np.argmin(r)
tongue_f = f[tongue_f_index]
factor = np.log(tongue_pos) / tongue_f
# plt.figure(figsize=(12,5))
plt.semilogx(np.exp(f*factor), r)
plt.semilogx([tongue_pos, tongue_pos], [0, max_r], 'r:')
plt.xlim(np.exp(factor*min_f), np.exp(factor*max_f))
```
| github_jupyter |
# Training a model with Sklearn
In this example we will train a model in Ray Air using a Sklearn classifier.
Let's start with installing our dependencies:
```
!pip install -qU "ray[tune]" sklearn
```
Then we need some imports:
```
import argparse
import math
from typing import Tuple
import pandas as pd
import ray
from ray.data.dataset import Dataset
from ray.ml.batch_predictor import BatchPredictor
from ray.ml.predictors.integrations.sklearn import SklearnPredictor
from ray.ml.preprocessors import Chain, OrdinalEncoder, StandardScaler
from ray.ml.result import Result
from ray.ml.train.integrations.sklearn import SklearnTrainer
from sklearn.datasets import load_breast_cancer
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
try:
from cuml.ensemble import RandomForestClassifier as cuMLRandomForestClassifier
except ImportError:
cuMLRandomForestClassifier = None
```
Next we define a function to load our train, validation, and test datasets.
```
def prepare_data() -> Tuple[Dataset, Dataset, Dataset]:
data_raw = load_breast_cancer()
dataset_df = pd.DataFrame(data_raw["data"], columns=data_raw["feature_names"])
dataset_df["target"] = data_raw["target"]
# add a random categorical column
num_samples = len(dataset_df)
dataset_df["categorical_column"] = pd.Series(
(["A", "B"] * math.ceil(num_samples / 2))[:num_samples]
)
train_df, test_df = train_test_split(dataset_df, test_size=0.3)
train_dataset = ray.data.from_pandas(train_df)
valid_dataset = ray.data.from_pandas(test_df)
test_dataset = ray.data.from_pandas(test_df.drop("target", axis=1))
return train_dataset, valid_dataset, test_dataset
```
The following function will create a Sklearn trainer, train it, and return the result.
```
def train_sklearn(num_cpus: int, use_gpu: bool = False) -> Result:
if use_gpu and not cuMLRandomForestClassifier:
raise RuntimeError("cuML must be installed for GPU enabled sklearn estimators.")
train_dataset, valid_dataset, _ = prepare_data()
# Scale some random columns
columns_to_scale = ["mean radius", "mean texture"]
preprocessor = Chain(
OrdinalEncoder(["categorical_column"]), StandardScaler(columns=columns_to_scale)
)
if use_gpu:
trainer_resources = {"CPU": 1, "GPU": 1}
estimator = cuMLRandomForestClassifier()
else:
trainer_resources = {"CPU": num_cpus}
estimator = RandomForestClassifier()
trainer = SklearnTrainer(
estimator=estimator,
label_column="target",
datasets={"train": train_dataset, "valid": valid_dataset},
preprocessor=preprocessor,
cv=5,
scaling_config={
"trainer_resources": trainer_resources,
},
)
result = trainer.fit()
print(result.metrics)
return result
```
Once we have the result, we can do batch inference on the obtained model. Let's define a utility function for this.
```
def predict_sklearn(result: Result, use_gpu: bool = False):
_, _, test_dataset = prepare_data()
batch_predictor = BatchPredictor.from_checkpoint(
result.checkpoint, SklearnPredictor
)
predicted_labels = (
batch_predictor.predict(
test_dataset,
num_gpus_per_worker=int(use_gpu),
)
.map_batches(lambda df: (df > 0.5).astype(int), batch_format="pandas")
.to_pandas(limit=float("inf"))
)
print(f"PREDICTED LABELS\n{predicted_labels}")
```
Now we can run the training:
```
result = train_sklearn(num_cpus=2, use_gpu=False)
```
And perform inference on the obtained model:
```
predict_sklearn(result, use_gpu=False)
```
| github_jupyter |
```
import Augmentor
import keras
from keras.preprocessing.image import image, img_to_array
from keras.preprocessing.image import ImageDataGenerator
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
from keras.layers import Conv2D, MaxPooling2D
import numpy as np
import matplotlib.pyplot as plt
from keras.layers import Input, Dense, Conv2D, MaxPooling2D, UpSampling2D
from keras.models import Model
from keras import backend as K
p = Augmentor.Pipeline("data/train/")
#p.status()
p.rotate90(probability=0.05)
p.rotate270(probability=0.05)
p.flip_left_right(probability=0.08)
p.flip_top_bottom(probability=0.03)
p.crop_random(probability=1, percentage_area=0.95)
p.resize(probability=1.0, width=128, height=128)
p.greyscale(1.0)
p.sample(100)
batch_size = 128
g = p.keras_generator(batch_size=batch_size)
images, labels = next(g)
plt.imshow(images[0].reshape(128, 128), cmap="Spectral");
input_img = Input(shape=(128, 128, 1)) # adapt this if using `channels_first` image data format
x = Conv2D(32, (3, 3), activation='relu', padding='same')(input_img)
x = MaxPooling2D((2, 2), padding='same')(x)
x = Conv2D(32, (3, 3), activation='relu', padding='same')(x)
encoded = MaxPooling2D((2, 2), padding='same')(x)
# at this point the representation is (7, 7, 32)
x = Conv2D(32, (3, 3), activation='relu', padding='same')(encoded)
x = UpSampling2D((2, 2))(x)
x = Conv2D(32, (3, 3), activation='relu', padding='same')(x)
x = UpSampling2D((2, 2))(x)
decoded = Conv2D(1, (3, 3), activation='sigmoid', padding='same')(x)
autoencoder = Model(input_img, decoded)
autoencoder.compile(optimizer='adadelta', loss='binary_crossentropy')
x_train =images
noise_factor = 0.5
x_train_noisy = x_train + noise_factor * np.random.normal(loc=0.0, scale=1.0, size=x_train.shape)
x_train_noisy = np.clip(x_train_noisy, 0., 1.)
autoencoder.fit(x_train_noisy, x_train, epochs=500, batch_size=128, shuffle=True,
validation_split=0.33,verbose=1)
n = 8
plt.figure(figsize=(20, 2))
for i in range(n):
ax = plt.subplot(1, n, i+1)
plt.imshow(x_train_noisy[i].reshape(128, 128))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
plt.show()
import warnings
warnings.filterwarnings("ignore")
decoded_imgs = autoencoder.predict(x_train)
n =8
plt.figure(figsize=(20, 4))
for i in range(n):
# display original
ax = plt.subplot(3, n, i+1)
plt.imshow(x_train[i].reshape(128, 128))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
# display reconstruction
ax = plt.subplot(2, n, i + n+1)
plt.imshow(decoded_imgs[i].reshape(128, 128))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
plt.show()
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Hub Authors.
Licensed under the Apache License, Version 2.0 (the "License");
Created by @[Adrish Dey](https://github.com/captain-pool) for [Google Summer of Code](https://summerofcode.withgoogle.com/) 2019
```
# Copyright 2019 The TensorFlow Hub Authors. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
```
# 使用 ESRGAN 进行图像超解析
<table class="tfo-notebook-buttons" align="left">
<td> <a target="_blank" href="https://tensorflow.google.cn/hub/tutorials/image_enhancing"><img src="https://tensorflow.google.cn/images/tf_logo_32px.png">View on TensorFlow.org</a> </td>
<td> <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/zh-cn/hub/tutorials/image_enhancing.ipynb"><img src="https://tensorflow.google.cn/images/colab_logo_32px.png">Run in Google Colab</a> </td>
<td> <a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/zh-cn/hub/tutorials/image_enhancing.ipynb"><img src="https://tensorflow.google.cn/images/GitHub-Mark-32px.png">View on GitHub</a> </td>
<td> <a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/zh-cn/hub/tutorials/image_enhancing.ipynb"><img src="https://tensorflow.google.cn/images/download_logo_32px.png">Download notebook</a> </td>
<td><a href="https://tfhub.dev/captain-pool/esrgan-tf2/1"><img src="https://tensorflow.google.cn/images/hub_logo_32px.png">查看 TF Hub 模型</a></td>
</table>
此 Colab 演示了在 Enhanced Super Resolution Generative Adversarial Network(*由 Xintao Wang 等人撰写*)[[论文](https://arxiv.org/pdf/1809.00219.pdf)] [[代码](https://github.com/captain-pool/GSOC/)] 中如何使用 TensorFlow Hub 模块
进行图像增强。*(最好使用双三次降采样的图像。)*
模型在 DIV2K 数据集(双三次降采样的图像)中大小为 128 x 128 的图像块上进行了训练。
**准备环境**
```
import os
import time
from PIL import Image
import numpy as np
import tensorflow as tf
import tensorflow_hub as hub
import matplotlib.pyplot as plt
os.environ["TFHUB_DOWNLOAD_PROGRESS"] = "True"
!wget "https://user-images.githubusercontent.com/12981474/40157448-eff91f06-5953-11e8-9a37-f6b5693fa03f.png" -O original.png
# Declaring Constants
IMAGE_PATH = "original.png"
SAVED_MODEL_PATH = "https://tfhub.dev/captain-pool/esrgan-tf2/1"
```
**定义辅助函数**
```
def preprocess_image(image_path):
""" Loads image from path and preprocesses to make it model ready
Args:
image_path: Path to the image file
"""
hr_image = tf.image.decode_image(tf.io.read_file(image_path))
# If PNG, remove the alpha channel. The model only supports
# images with 3 color channels.
if hr_image.shape[-1] == 4:
hr_image = hr_image[...,:-1]
hr_size = (tf.convert_to_tensor(hr_image.shape[:-1]) // 4) * 4
hr_image = tf.image.crop_to_bounding_box(hr_image, 0, 0, hr_size[0], hr_size[1])
hr_image = tf.cast(hr_image, tf.float32)
return tf.expand_dims(hr_image, 0)
def save_image(image, filename):
"""
Saves unscaled Tensor Images.
Args:
image: 3D image tensor. [height, width, channels]
filename: Name of the file to save.
"""
if not isinstance(image, Image.Image):
image = tf.clip_by_value(image, 0, 255)
image = Image.fromarray(tf.cast(image, tf.uint8).numpy())
image.save("%s.jpg" % filename)
print("Saved as %s.jpg" % filename)
%matplotlib inline
def plot_image(image, title=""):
"""
Plots images from image tensors.
Args:
image: 3D image tensor. [height, width, channels].
title: Title to display in the plot.
"""
image = np.asarray(image)
image = tf.clip_by_value(image, 0, 255)
image = Image.fromarray(tf.cast(image, tf.uint8).numpy())
plt.imshow(image)
plt.axis("off")
plt.title(title)
```
#### 对从路径加载的图像执行超解析
```
hr_image = preprocess_image(IMAGE_PATH)
# Plotting Original Resolution image
plot_image(tf.squeeze(hr_image), title="Original Image")
save_image(tf.squeeze(hr_image), filename="Original Image")
model = hub.load(SAVED_MODEL_PATH)
start = time.time()
fake_image = model(hr_image)
fake_image = tf.squeeze(fake_image)
print("Time Taken: %f" % (time.time() - start))
# Plotting Super Resolution Image
plot_image(tf.squeeze(fake_image), title="Super Resolution")
save_image(tf.squeeze(fake_image), filename="Super Resolution")
```
### 评估模型的性能
```
!wget "https://lh4.googleusercontent.com/-Anmw5df4gj0/AAAAAAAAAAI/AAAAAAAAAAc/6HxU8XFLnQE/photo.jpg64" -O test.jpg
IMAGE_PATH = "test.jpg"
# Defining helper functions
def downscale_image(image):
"""
Scales down images using bicubic downsampling.
Args:
image: 3D or 4D tensor of preprocessed image
"""
image_size = []
if len(image.shape) == 3:
image_size = [image.shape[1], image.shape[0]]
else:
raise ValueError("Dimension mismatch. Can work only on single image.")
image = tf.squeeze(
tf.cast(
tf.clip_by_value(image, 0, 255), tf.uint8))
lr_image = np.asarray(
Image.fromarray(image.numpy())
.resize([image_size[0] // 4, image_size[1] // 4],
Image.BICUBIC))
lr_image = tf.expand_dims(lr_image, 0)
lr_image = tf.cast(lr_image, tf.float32)
return lr_image
hr_image = preprocess_image(IMAGE_PATH)
lr_image = downscale_image(tf.squeeze(hr_image))
# Plotting Low Resolution Image
plot_image(tf.squeeze(lr_image), title="Low Resolution")
model = hub.load(SAVED_MODEL_PATH)
start = time.time()
fake_image = model(lr_image)
fake_image = tf.squeeze(fake_image)
print("Time Taken: %f" % (time.time() - start))
plot_image(tf.squeeze(fake_image), title="Super Resolution")
# Calculating PSNR wrt Original Image
psnr = tf.image.psnr(
tf.clip_by_value(fake_image, 0, 255),
tf.clip_by_value(hr_image, 0, 255), max_val=255)
print("PSNR Achieved: %f" % psnr)
```
**并排比较输出大小。**
```
plt.rcParams['figure.figsize'] = [15, 10]
fig, axes = plt.subplots(1, 3)
fig.tight_layout()
plt.subplot(131)
plot_image(tf.squeeze(hr_image), title="Original")
plt.subplot(132)
fig.tight_layout()
plot_image(tf.squeeze(lr_image), "x4 Bicubic")
plt.subplot(133)
fig.tight_layout()
plot_image(tf.squeeze(fake_image), "Super Resolution")
plt.savefig("ESRGAN_DIV2K.jpg", bbox_inches="tight")
print("PSNR: %f" % psnr)
```
| github_jupyter |
# AdaGrad
Presented during ML reading group, 2019-11-5.
Author: Ioana Plajer, ioana.plajer@unitbv.ro
Reviewed: Lucian Sasu
```
#%matplotlib notebook
%matplotlib inline
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
print(f'Numpy version: {np.__version__}')
```
# AdaGrad
The [AdaGrad paper](http://www.jmlr.org/papers/volume12/duchi11a/duchi11a.pdf) comes with the idea of using different learning rates for each feature. Hence, instead of:
$$w_{t+1} = w_{t} - \eta \nabla J_{w}(w_t)$$
AdaGrad comes with:
$$w_{t+1}^{(j)} = w_{t}^{(j)} - \frac{\eta}{\sqrt{\varepsilon + \sum_{\tau=1}^{t}{(g_{\tau}^{(j)})^2}}} \nabla J_{w}(w_t^{(j)})$$
where $g_{\tau}$ is the gradient of error function at iteration $\tau$, $g_{\tau}^{(j)}$ is the partial derivative of the
error function in direction of the $j$ - th feature, at iteration $\tau$, $m$ - is the number of features, i.e.
$$g_{\tau}^{(j)} = \nabla J_{w}(w_\tau^{(j)})$$
AdaGrad specifies the update as:
$$w_{t+1} = w_{t} - \frac{\eta}{\sqrt{\varepsilon I + diag(G_t)}} \nabla J_{w}(w_t)$$
where:
* $\eta$ is the initial learning rate (hyperparameter)
* $n$ is the number of items in (mini)batch
* $G_t = \sum\limits_{\tau=1}^t \mathbf{g}_\tau \mathbf{g}_\tau^T$
* $diag(A)$ is the diagonal form of the square matrix $A$
* $\varepsilon > 0$ is used to avoid division by 0
* $I$ is the unit matrix of size $m$
* $G_t^{(j,j)} = \sum\limits_{\tau = 1}^{t}{(g_\tau^{(j)})^2}$ is the sum of the squared partial derivatives in direction
of the $j$ - th feature from the first iteration up to the current iteration
In a more detailed form, the update of the weights through AdaGrad is done by:
$$\left[\begin{array}{c}
w_{t+1}^{(1)}\\
w_{t+1}^{(2)}\\
\vdots\\
w_{t+1}^{(m)}
\end{array}\right] = \left[\begin{array}{c}
w_{t}^{(1)}\\
w_{t}^{(2)}\\
\vdots\\
w_{t}^{(m)}\end{array}\right] - \eta\left(\left[\begin{array}{cccc} \varepsilon & 0 & \ldots & 0\\
0 & \varepsilon & \ldots & 0\\
\vdots & \vdots & \ddots & \vdots \\
0 & 0 & \ldots & 0\end{array}\right]+
\left[\begin{array}{cccc}
G_{t}^{(1,1)} & 0 & \ldots & 0\\
0 & G_{t}^{(2,2)} & \ldots & 0\\
\vdots & \vdots & \ddots & \vdots\\
0 & 0 & \ldots & G_{t}^{(m,m)}\end{array}\right]\right)^{-1/2}
\left[\begin{array}{c}
g_t^{(1)}\\
g_t^{(2)}\\
\vdots\\
g_t^{(m)}\end{array}\right]$$
which simplifies to:
$$\left[\begin{array}{c}
w_{t+1}^{(1)}\\
w_{t+1}^{(2)}\\
\vdots\\
w_{t+1}^{(m)}
\end{array}\right] = \left[\begin{array}{c}
w_{t}^{(1)}\\
w_{t}^{(2)}\\
\vdots\\
w_{t}^{(m)}\end{array}\right] - \left[\begin{array}{c}
\frac{\eta}{\sqrt{\varepsilon+G_{t}^{(1,1)}}}g_t^{(1)}\\
\frac{\eta}{\sqrt{\varepsilon+G_{t}^{(2,2)}}}g_t^{(2)}\\
\vdots\\
\frac{\eta}{\sqrt{\varepsilon+G_{t}^{(m,m)}}}g_t^{(m)}
\end{array}\right]$$
## Generate data
```
from scipy.sparse import random #to generate sparse data
np.random.seed(10) # for reproducibility
m_data = 100
n_data = 4 #number of features of the data
_scales = np.array([1,10, 10,1]) # play with these...
_parameters = np.array([3, 0.5, 1, 7])
def gen_data(m, n, scales, parameters, add_noise=True):
# Adagrad is designed especially for sparse data.
# produce: X, a 2d tensor with m lines and n columns
# and X[:, k] uniformly distributed in [-scale_k, scale_k] with the first and the last column containing sparse data
#(approx 75% of the elements are 0)
#
# To generate a sparse data matrix with m rows and n columns
# and random values use S = random(m, n, density=0.25).A, where density = density of the data. S will be the
# resulting matrix
# more information at https://docs.scipy.org/doc/scipy/reference/generated/scipy.sparse.random.html
#
# To obtain X - generate a random matrix with X[:, k] uniformly distributed in [-scale_k, scale_k]
# set X[:, 0] and X[:, -1] to 0 and add matrix S with the sparse data.
#
# let y be X@parameters.T + epsilon, with epsilon ~ N(0, 1); y is a vector with m elements
# parameters - the ideal weights, used to produce output values y
#
return X, y
X, y = gen_data(m_data, n_data, _scales, _parameters)
```
## Define error function, gradient, inference
```
def model_estimate(X, w):
'''Computes the linear regression estimation on the dataset X, using coefficients w
:param X: a 2d tensor with m_data lines and n_data columns
:param w: a 1d tensor with n_data coefficients (no intercept)
:return: a 1d tensor with m_data elements y_hat = w @X.T
'''
return y_hat
def J(X, y, w):
"""Computes the mean squared error of model. See the picture from last week's sheet.
:param X: input values, of shape m_data x n_data
:param y: ground truth, column vector with m_data values
:param w: column with n_data coeffieicnts for the linear form
:return: a scalar value >= 0
:use the same formula as in the exercise from last week
"""
return err
def gradient(X, y, w):
'''Commputes the gradients to be used for gradient descent.
:param X: 2d tensor with training data
:param y: 1d tensor with y.shape[0] == W.shape[0]
:param w: 1d tensor with current values of the coefficients
:return: gradients to be used for gradient descent.
:use the same formula as in the exercise from last week
'''
return grad## implement
#The function from last week for comparison
def gd_no_momentum(X, y, w_init, eta=1e-1, thresh = 0.001):
'''Iterates with gradient descent algorithm
:param X: 2d tensor with data
:param y: 1d tensor, ground truth
:param w_init: 1d tensor with the X.shape[1] initial coefficients
:param eta: the learning rate hyperparameter
:param thresh: the threshold for the gradient norm (to stop iterations)
:return: the list of succesive errors and the found w* vector
'''
w = w_init
w_err=[]
while True:
grad = gradient(X, y, w)
err = J(X, y, w)
w_err.append(err)
w = w - eta * grad
if np.linalg.norm(grad) < thresh:
break;
return w_err, w
w_init = np.array([0, 0, 0, 0])
errors, w_best = gd_no_momentum(X, y, w_init, 0.0001)
print(f'How many iterations were made: {len(errors)}')
w_best
fig, axes = plt.subplots()
axes.plot(list(range(len(errors))), errors)
axes.set_xlabel('Epochs')
axes.set_ylabel('Error')
axes.set_title('Optimization without momentum')
```
## Momentum algorithm
```
#The function from last week for comparison
def gd_with_momentum(X, y, w_init, eta=1e-1, gamma = 0.9, thresh = 0.001):
"""Applies gradient descent with momentum coefficient
:params: as in gd_no_momentum
:param gamma: momentum coefficient
:param thresh: the threshold for the gradient norm (to stop iterations)
:return: the list of succesive errors and the found w* vector
"""
w = w_init
w_err=[]
delta = np.zeros_like(w)
while True:
grad = gradient(X, y, w)
err = J(X, y, w)
w_err.append(err)
w_new = w + gamma * delta - eta * grad
delta = w_new - w
w = w_new
if np.linalg.norm(grad) < thresh :
break;
return w_err, w
w_init = np.array([0, 0, 0, 0])
errors_momentum, w_best = gd_with_momentum(X, y, w_init,0.0001, 0.9)
print(f'How many iterations were made: {len(errors_momentum)}')
w_best
fig, axes = plt.subplots()
axes.plot(list(range(len(errors_momentum))), errors_momentum)
axes.set_xlabel('Epochs')
axes.set_ylabel('Error')
axes.set_title('Optimization with momentum')
```
## Apply AdaGrad and report resulting $\eta$'s
```
def ada_grad(X, y, w_init, eta_init=1e-1, eps = 0.001,thresh = 0.001):
'''Iterates with gradient descent algorithm
:param X: 2d tensor with data
:param y: 1d tensor, ground truth
:param w_init: 1d tensor with the X.shape[1] initial coefficients
:param eps: the epsilon value from the AdaGrad formula
:param thresh: the threshold for the gradient norm (to stop iterations)
:return: the list of succesive errors w_err, the found w - the estimated feature vector
:and rates the learning rates after the final iteration
'''
return w_err, w, rates
w_init = np.array([0,0,0,0])
adaGerr, w_ada_best, rates = ada_grad(X, y, w_init)
print(f'How many iterations were made: {len(adaGerr)}')
w_ada_best
fig, axes = plt.subplots()
axes.plot(list(range(len(adaGerr))),adaGerr)
axes.set_xlabel('Epochs')
axes.set_ylabel('Error')
axes.set_title('Optimization with AdaGrad')
```
| github_jupyter |
# Meraki Python SDK Demo: Uplink Preference Backup
*This notebook demonstrates using the Meraki Python SDK to back up Internet (WAN) and VPN traffic uplink preferences, as well as custom performance classes, to a single Excel file. If you have hundreds of WAN/VPN uplink preferences, they can be a challenge to manipulate. This demo seeks to prove how using the Meraki API and Python SDK can substantially streamline such complex deployments.*
If an admin has any Internet traffic or VPN traffic uplink preferences, or custom performance classes, this tool will download them to an Excel file. This is a more advanced demo, intended for intermediate to advanced Python programmers, but has been documented thoroughly with the intention that even a determined Python beginner can understand the concepts involved.
---
>NB: Throughout this notebook, we will print values for demonstration purposes. In a production Python script, the coder would likely remove these print statements to clean up the console output.
In this first cell, we import the required `meraki` and `os` modules, and open the Dashboard API connection using the SDK. We also import `openpyxl` for working with Excel files, and `netaddr` for working with IP addresses.
```
# Install the relevant modules. If you are using a local editor (e.g. VS Code, rather than Colab) you can run these commands, without the preceding %, via a terminal. NB: Run `pip install meraki==` to find the latest version of the Meraki SDK. Uncomment these lines to run them in Google Colab.
#%pip install meraki
#%pip install openpyxl
# If you are using Google Colab, please ensure you have set up your environment variables as linked above, then delete the two lines of ''' to activate the following code:
'''
%pip install colab-env -qU
import colab_env
'''
# The Meraki SDK
import meraki
# The built-in OS module, to read environment variables
import os
# We're also going to import Python's built-in JSON module, but only to make the console output pretty. In production, you wouldn't need any of the printing calls at all, nor this import!
import json
# The openpyxl module, to manipulate Excel files
import openpyxl
# The datetime module, to generate timestamps
import datetime
# Treat your API key like a password. Store it in your environment variables as 'MERAKI_DASHBOARD_API_KEY' and let the SDK call it for you.
# Or, call it manually after importing Python's os module:
# API_KEY = os.getenv('MERAKI_DASHBOARD_API_KEY')
# Initialize the Dashboard connection.
dashboard = meraki.DashboardAPI(suppress_logging=True)
# We'll also create a few reusable strings for later interactivity.
string_constants = dict()
string_constants['CONFIRM'] = 'OK, are you sure you want to do this? This script does not have an "undo" feature.'
string_constants['CANCEL'] = 'OK. Operation canceled.'
string_constants['WORKING'] = 'Working...'
string_constants['COMPLETE'] = 'Operation complete.'
string_constants['NETWORK_SELECTED'] = 'Network selected.'
string_constants['NO_VALID_OPTIONS'] = 'There are no valid options. Please try again with an API key that has access to the appropriate resources.'
# Some of the parameters we'll work with are optional. This string defines what value will be put into a cell corresponding with a parameter that is not set on that rule.
string_constants['NOT_APPLICABLE'] = 'N/A'
# This script is interactive; user choices and data will be stored here.
user_choices = dict()
user_data = dict()
```
A basic pretty print formatter, `printj()`. It will make reading JSON on the console easier, but won't be necessary in production scripts.
```
def printj(ugly_json_object):
# The json.dumps() method converts a JSON object into human-friendly formatted text
pretty_json_string = json.dumps(ugly_json_object, indent = 2, sort_keys = False)
return print(pretty_json_string)
```
## Introducing a Python class
To streamline user interaction in a re-usable way, we'll create a class called UserChoice. Think of classes like a superset of functions, where you can store related functions and variables. Later, we'll create an instance of this class to prompt the user for input, and validate that input.
It may look complex, but it will streamline our code later, and is a great example of code-reuse in Python. For more information on classes, [click here](https://docs.python.org/3/tutorial/classes.html). Alternatively, you could reduce the script's interactivity and have it read required settings from a settings file, as in [this example](https://github.com/meraki/dashboard-api-python/tree/master/examples/merakiApplianceVlanToL3SwitchInterfaceMigrator).
```
class UserChoice:
'A re-usable CLI option prompt.'
def __init__(self, options_list=[], subject_of_choice='available options', single_option_noun='option', id_parameter='id', name_parameter='name', action_verb='choose', no_valid_options_message=string_constants['NO_VALID_OPTIONS']):
self.options_list = options_list # options_list is a list of dictionaries containing attributes id_parameter and name_parameter
self.subject_of_choice = subject_of_choice # subject_of_choice is a string that names the subject of the user's choice. It is typically a plural noun.
self.single_option_noun = single_option_noun # single_option_noun is a string that is a singular noun corresponding to the subject_of_choice
self.id_parameter = id_parameter # id_parameter is a string that represents the name of the sub-parameter that serves as the ID value for the option in options_list. It should be a unique value for usability.
self.name_parameter = name_parameter # name_paraemter is a string that represents the name of the sub-parameter that serves as the name value for the option in options_list. It does not need to be unique.
self.action_verb = action_verb # action_verb is a string that represents the verb of the user's action. For example, to "choose"
self.no_valid_options_message = no_valid_options_message # no_valid_options_message is a string that represents an error message if options_list is empty
# Confirm there are options in the list
if len(self.options_list):
print(f'We found {len(self.options_list)} {self.subject_of_choice}:')
# Label each option and show the user their choices.
option_index = 0
for option in self.options_list:
print(f"{option_index}. {option[self.id_parameter]} with name {option[self.name_parameter]}")
option_index+=1
print(f'Which {self.single_option_noun} would you like to {self.action_verb}?')
self.active_option = int(input(f'Choose 0-{option_index-1}:'))
# Ask until the user provides valid input.
while self.active_option not in list(range(option_index)):
print(f'{self.active_option} is not a valid choice. Which {self.single_option_noun} would you like to {self.action_verb}?')
self.active_option = int(input(f'Choose 0-{option_index-1}:'))
print(f'Your {self.single_option_noun} is {self.options_list[self.active_option][self.name_parameter]}.')
# Assign the class id and name vars to the chosen item's
self.id = self.options_list[self.active_option][self.id_parameter]
self.name = self.options_list[self.active_option][self.name_parameter]
```
## Pulling organization and network IDs
Most API calls require passing values for the organization ID and/or the network ID. Remember that `UserChoice` class we created earlier? We'll call that and supply parameters defining what the user can choose. Notice how, having defined the class earlier, we can re-use it with only a single declaration.
```
# getOrganizations will return all orgs to which the supplied API key has access
user_choices['all_organizations'] = dashboard.organizations.getOrganizations()
# Prompt the user to pick an organization.
user_choices['organization'] = UserChoice(
options_list=user_choices['all_organizations'],
subject_of_choice='organizations',
single_option_noun='organization'
)
```
## Identify networks with MX appliances, and prompt the user to choose one
We want to:
> Download any existing uplink selection preferences and custom performance classes.
> Optionally, upload a replacement list of preferences.
We can only run this on networks that have appliance devices, so we will find networks where `productTypes` contains `appliance`. Then we'll ask the user to pick one, then pull the uplink selection rules from it.
Then let's ask the user which network they'd like to use.
```
# Make a list of all networks in the org
user_choices['all_networks'] = dashboard.organizations.getOrganizationNetworks(organizationId=user_choices['organization'].id)
# Find the networks with appliances
user_choices['networks_with_appliances'] = [network for network in user_choices['all_networks'] if 'appliance' in network['productTypes']]
# If any are found, let the user choose a network. Otherwise, let the user know that none were found. The logic for this class is defined in a cell above.
user_choices['network'] = UserChoice(
options_list = user_choices['networks_with_appliances'],
subject_of_choice = 'networks with appliances',
single_option_noun = 'network'
)
```
## Pulling uplink preferences for the network
Let's pull the uplink preferences via the API using the SDK's `getNetworkApplianceTrafficShapingUplinkSelection` and `getNetworkApplianceTrafficShapingCustomPerformanceClasses` methods. The associated endpoints will return all of the relevant information for the chosen network. We'll create an Excel file of it later.
Let's define this operation as a function so we can re-use it later.
```
def getNetworkClassesAndPrefs(*, networkId):
# Get the uplink preferences (WAN and VPN)
uplink_prefs = dashboard.appliance.getNetworkApplianceTrafficShapingUplinkSelection(networkId=networkId)
# Get the custom performance classes
custom_performance_classes = dashboard.appliance.getNetworkApplianceTrafficShapingCustomPerformanceClasses(networkId=networkId)
# Create a dict network_classes_and_prefs with both that we can return and call later
network_classes_and_prefs = {
'uplinkPrefs': uplink_prefs,
'customPerformanceClasses': custom_performance_classes
}
return(network_classes_and_prefs)
user_data['current_classes_and_prefs'] = getNetworkClassesAndPrefs(networkId = user_choices['network'].id)
```
The above returns all uplink preferences (e.g. Internet traffic and VPN traffic). We can review the output of each individually if we'd like to see the raw data:
```
# We can review the wanTrafficUplinkPreferences in plain text.
printj(user_data['current_classes_and_prefs']['uplinkPrefs']['wanTrafficUplinkPreferences'])
# We can also review the vpnTrafficUplinkPreferences in plain text.
printj(user_data['current_classes_and_prefs']['uplinkPrefs']['vpnTrafficUplinkPreferences'])
# We can also review the customPerformanceClasses in plain text.
printj(user_data['current_classes_and_prefs']['customPerformanceClasses'])
```
## Let's make a backup
Before we start modifying any of these rules or classes, it'd be good to make a backup, don't you think? We'll use `openpyxl` to create a new Excel workbook with two worksheets: one for Internet traffic preferences and the other for VPN traffic preferences. If you don't care to use Excel, you could instead create a CSV or potentially any other type of file you like, using an appropriate Python module. Consider a self-paced excercise where you save these values into some other file format.
We'll start with the creation of the workbook. We'll make this a function for easy calling later.
```
def create_workbook():
# Create a workbook with the appropriate column headers and/or worksheets
# First we'll create the workbook, then we'll design the worksheets. Just like in Excel, by default, the workbook has one worksheet, titled 'Sheet'.
new_workbook = openpyxl.Workbook()
# We'll specify that the active worksheet is our wan_prefs_worksheet
wan_prefs_worksheet = new_workbook.active
# Let's rename the worksheet from 'Sheet' to something more descriptive
wan_prefs_worksheet.title = 'wanUplinkPreferences'
# Name the columns for the wan_prefs_worksheet. Think of this like a single-line CSV:
wan_title_row_headers = (
'Protocol',
'Source',
'Src port',
'Destination',
'Dst port',
'Preferred uplink'
)
# Add that title row to the worksheet
wan_prefs_worksheet.append(wan_title_row_headers)
# Let's make the title row bold for easier reading
for row in wan_prefs_worksheet.iter_rows():
for cell in row:
cell.font = openpyxl.styles.Font(bold=True)
# Now let's do the same for the VPN uplink preferences, and custom performance classes.
# First, create a separate worksheet for the VPN uplink preferences
vpn_prefs_worksheet = new_workbook.create_sheet(title='vpnUplinkPreferences')
custom_performance_classes_worksheet = new_workbook.create_sheet(title='customPerformanceClasses')
# Name the columns for the vpn_prefs_worksheet and custom_performance_classes_worksheet.
vpn_title_row_headers = (
'Type',
'Protocol or App ID',
'Source or App Name',
'Src port',
'Destination',
'Dst port',
'Preferred uplink',
'Failover criterion',
'Performance class type',
'Performance class name',
'Performance class ID'
)
classes_title_row_headers = (
'ID',
'Name',
'Max Latency',
'Max Jitter',
'Max Loss Percentage'
)
# Add the title rows to the appropriate worksheets
vpn_prefs_worksheet.append(vpn_title_row_headers)
custom_performance_classes_worksheet.append(classes_title_row_headers)
# Let's make those title rows bold, too
for row in vpn_prefs_worksheet.iter_rows():
for cell in row:
cell.font = openpyxl.styles.Font(bold=True)
for row in custom_performance_classes_worksheet.iter_rows():
for cell in row:
cell.font = openpyxl.styles.Font(bold=True)
print(f'Created formatted workbook.')
return(new_workbook)
```
### Function to add custom performance classes to a workbook
VPN uplink prefs might reference performance classes. Accordingly, we need to back those up as well.
```
def add_custom_performance_classes_to_workbook(workbook):
# We'll specify that the active worksheet is our custom_performance_classes worksheet
custom_performance_classes_worksheet = workbook['customPerformanceClasses']
# We'll also count the number of classes to help the user know that it's working.
performance_class_count = 0
# Let's add all the vpnTrafficUplinkPreferences to the VPN worksheet
for performance_class in user_data['current_classes_and_prefs']['customPerformanceClasses']:
performance_class_id = performance_class['customPerformanceClassId']
performance_class_name = performance_class['name']
performance_class_max_latency = performance_class['maxLatency']
performance_class_max_jitter = performance_class['maxJitter']
performance_class_max_loss_percentage = performance_class['maxLossPercentage']
# We assemble the parameters into a tuple that will represent a row.
performance_class_row = (
performance_class_id,
performance_class_name,
performance_class_max_latency,
performance_class_max_jitter,
performance_class_max_loss_percentage
)
# We then add that row to the worksheet.
custom_performance_classes_worksheet.append(performance_class_row)
# increase the rule_count
performance_class_count += 1
print(f'Added {performance_class_count} performance classes to customPerformanceClasses.')
return(workbook)
```
### Function to add WAN preferences to a workbook
Transposing the settings into a two-dimensional table doesn't require anything too fancy. We simply iterate through the rules in the `uplinkPrefs['wanTrafficUplinkPreferences']` list, and pull out the relevant information for each key-value pair. We'll also make this a function so we can call it later.
```
def add_wan_prefs_to_workbook(workbook):
# We'll specify that the active worksheet is our wan_prefs_worksheet
wan_prefs_worksheet = workbook['wanUplinkPreferences']
# We'll also count the number of rules to help the user know that it's working.
rule_count = 0
# Let's add all the wanTrafficUplinkPreferences
for rule in user_data['current_classes_and_prefs']['uplinkPrefs']['wanTrafficUplinkPreferences']:
rule_preferred_uplink = rule['preferredUplink']
rule_protocol = rule['trafficFilters'][0]['value']['protocol']
rule_source = rule['trafficFilters'][0]['value']['source']['cidr']
# An 'any' value in the protocol removes the need for either source or destination port numbers to be defined, so the API doesn't specify port numbers in the output if 'any' is the protocol. However, we don't want to leave those cells blank, so we will fill them in accordingly.
if rule_protocol == 'any':
rule_src_port = 'any'
rule_dst_port = 'any'
else:
rule_src_port = rule['trafficFilters'][0]['value']['source']['port']
rule_dst_port = rule['trafficFilters'][0]['value']['destination']['port']
rule_destination = rule['trafficFilters'][0]['value']['destination']['cidr']
# We assemble the parameters into a tuple that will represent a row.
rule_row = (
rule_protocol,
rule_source,
rule_src_port,
rule_destination,
rule_dst_port,
rule_preferred_uplink
)
# We then add that row to the worksheet.
wan_prefs_worksheet.append(rule_row)
# increase the rule_count
rule_count += 1
print(f'Added {rule_count} rules to wanUplinkPreferences.')
return(workbook)
```
### Function to add VPN preferences to a workbook
We'll do almost the exact same for the VPN traffic uplink preferences, but since there are more parameters available here, we employ a bit more `if`/`else` logic to handle the varied key structures.
```
def add_vpn_prefs_to_workbook(workbook):
# We'll specify that the active worksheet is our wan_prefs_worksheet
vpn_prefs_worksheet = workbook['vpnUplinkPreferences']
# We'll also count the number of rules to help the user know that it's working.
rule_count = 0
# Let's add all the vpnTrafficUplinkPreferences to the VPN worksheet
for rule in user_data['current_classes_and_prefs']['uplinkPrefs']['vpnTrafficUplinkPreferences']:
rule_preferred_uplink = rule['preferredUplink']
rule_type = rule['trafficFilters'][0]['type']
# Application rules have different parameters. This checks, and assigns values accordingly.
if 'application' in rule_type:
rule_protocol = rule['trafficFilters'][0]['value']['id']
rule_source = rule['trafficFilters'][0]['value']['name']
rule_src_port = string_constants['NOT_APPLICABLE']
rule_destination = string_constants['NOT_APPLICABLE']
rule_dst_port = string_constants['NOT_APPLICABLE']
else:
rule_protocol = rule['trafficFilters'][0]['value']['protocol']
rule_source = rule['trafficFilters'][0]['value']['source']['cidr']
rule_destination = rule['trafficFilters'][0]['value']['destination']['cidr']
# An 'any' or 'icmp' value in the protocol removes the need for either source or destination port numbers to be defined, so the API doesn't specify port numbers in the output if 'any' or 'icmp' is the protocol. However, we don't want to leave those cells blank, so we will fill them in accordingly.
if rule_protocol == 'any':
rule_src_port = 'any'
rule_dst_port = 'any'
elif rule_protocol == 'icmp':
rule_src_port = string_constants['NOT_APPLICABLE']
rule_dst_port = string_constants['NOT_APPLICABLE']
else:
rule_src_port = rule['trafficFilters'][0]['value']['source']['port']
rule_dst_port = rule['trafficFilters'][0]['value']['destination']['port']
# A failover criterion and performance class are optional parameters, so we cannot assume they are there. We'll first check to see if they exist before pulling their values.
# Check if the rule has failOverCriterion set
if rule.get('failOverCriterion', False):
rule_failover_criterion = rule['failOverCriterion']
else: # No failOverCriterion set
rule_failover_criterion = string_constants['NOT_APPLICABLE']
# Check if the rule has performanceClass set
if rule.get('performanceClass', False):
rule_performance_class_type = rule['performanceClass']['type']
# If the performance class is set, and is 'builtin', then we use 'builtinPerformanceClassName'. If it's 'custom', then we use 'customPerformanceClassId' to identify it.
if rule_performance_class_type == 'builtin':
rule_performance_class_id = string_constants['NOT_APPLICABLE']
rule_performance_class_name = rule['performanceClass']['builtinPerformanceClassName']
else:
rule_performance_class_id = rule['performanceClass']['customPerformanceClassId']
# search user_data['current_classes_and_prefs']['customPerformanceClasses'] for the class that has that ID, then pull the corresponding name
for performance_class in user_data['current_classes_and_prefs']['customPerformanceClasses']:
if rule_performance_class_id == performance_class['customPerformanceClassId']:
rule_performance_class_name = performance_class['name']
# Else, there's no performanceClass set, so we'll set these values accordingly.
else:
rule_performance_class_type = string_constants['NOT_APPLICABLE']
rule_performance_class_name = string_constants['NOT_APPLICABLE']
rule_performance_class_id = string_constants['NOT_APPLICABLE']
# We assemble the parameters into a tuple that will represent a row.
rule_row = (
rule_type,
rule_protocol,
rule_source,
rule_src_port,
rule_destination,
rule_dst_port,
rule_preferred_uplink,
rule_failover_criterion,
rule_performance_class_type,
rule_performance_class_name,
rule_performance_class_id
)
# We then add that row to the worksheet.
vpn_prefs_worksheet.append(rule_row)
# increase the rule_count
rule_count += 1
print(f'Added {rule_count} rules to wanUplinkPreferences.')
return(workbook)
```
### Function to save a workbook
This function takes a single workbook as an argument, and saves it.
```
# Function that saves a workbook
def save_workbook(workbook):
# Finally, we save the worksheet.
# Let's give it a name with a date/time stamp
downloaded_rules_workbook_filename = f'downloaded_rules_workbook_{datetime.datetime.now()}.xlsx'.replace(':','')
workbook.save(downloaded_rules_workbook_filename)
print(f'Saved {downloaded_rules_workbook_filename}.')
```
### Build and save the workbook
Now that we've defined the functions that will build the workbook object in Python and save it to a file, we need to run them.
```
uplink_prefs_workbook = create_workbook()
add_custom_performance_classes_to_workbook(uplink_prefs_workbook)
add_wan_prefs_to_workbook(uplink_prefs_workbook)
add_vpn_prefs_to_workbook(uplink_prefs_workbook)
save_workbook(uplink_prefs_workbook)
```
# Final thoughts
Hopefully this was a useful deep dive into Python programming and interacting with the Meraki SDK and Excel workbooks. We tackled a problem that is tough to solve in the Dashboard GUI and showed how it can be done very quickly via API and the Python SDK.
But what if we want to RESTORE that backup? Well, that's the next challenge! [Return to the notebooks folder](https://github.com/meraki/dashboard-api-python/tree/master/notebooks) for an example notebook that can restore such a backup to the Dashboard.
Here we used Excel workbooks, but you can imagine that there are all types of data structures that might be used instead of Excel workbooks, e.g. CSVs, plain text, YAML, XML, LibreOffice files or others, and with the right code you can use those instead of Excel.
## Further learning
[Meraki Interactive API Docs](https://developer.cisco.com/meraki/api-v1/#!overview): The official (and interactive!) Meraki API and SDK documentation repository on DevNet.
| github_jupyter |
### plot the results from the hyper parameter optimization
```
import matplotlib.pyplot as plt
import numpy as np
import os
classifier = 'dt'
#classifier = 'bdt'
classifier = 'rf'
#classifier = 'mlp'
outfile = "/Users/manuelmeyer/Python/TESdata/data-04202020/" \
"dv1p7-all-expflare/dvdtthr10_vthr10_steploNone_stepupNone/ml-results/"
results = np.load(os.path.join(outfile, f'results_{classifier:s}.npy'),
allow_pickle=True).flat[0]
```
### plot the parameter profiles
```
scores = list(results['learning_curve'].keys())
print (scores)
plt.figure(figsize=(4*3, 4))
ax = []
for i, score in enumerate(scores):
color = 'g' if score == 'AUC' else 'k'
for j, par in enumerate(results['profile']['mean_test'][score].keys()):
if not i:
ax.append(plt.subplot(1,len(results['profile']['mean_test'][score].keys()), j+1))
print (par)
if par == 'hidden_layer_sizes':
x = np.unique([np.sum(results['gs_cv'][f'param_{par}'].data[i]) for i in \
range(results['gs_cv'][f'param_{par}'].data.size)])
else:
x = np.unique(results['gs_cv'][f'param_{par}'].data).astype(np.float)
for t in ['test', 'train']:
ax[j].plot(x, results['profile'][f'mean_{t:s}'][score][par],
color=color,
ls='-' if t == 'test' else '--',
label=score + " " + t
)
if t == 'test':
ax[j].fill_between(x, results['profile'][f'mean_{t:s}'][score][par] - \
0.5 * results['profile'][f'std_{t:s}'][score][par],
y2=results['profile'][f'mean_{t:s}'][score][par] + \
0.5 * results['profile'][f'std_{t:s}'][score][par],
color=color,
alpha=0.3)
if not i:
ax[j].set_xlabel(par)
else:
ax[j].legend()
ax[j].set_ylim(0.973,1.001)
ax[j].grid()
if j:
ax[j].tick_params(labelleft=False)
plt.subplots_adjust(wspace = 0.05)
plt.savefig(f"{classifier}_profiles.png", dpi=150)
```
### plot the parameter profiles
### plot the learning curve
```
for score, val in results['learning_curve'].items():
if not score == 'Accuracy':
continue
train_sizes, train_scores, valid_scores = val
plt.plot(train_sizes, train_scores.mean(axis=1),
marker='o',
label=score + " Train",
ls='--',
color='g' if score == 'AUC' else 'k'
)
plt.fill_between(train_sizes,
train_scores.mean(axis=1) - np.sqrt(train_scores.var()),
y2=train_scores.mean(axis=1) + np.sqrt(train_scores.var()),
alpha=0.3,
color='g' if score == 'AUC' else 'k',
zorder=-1
)
plt.plot(train_sizes, valid_scores.mean(axis=1),
marker='o',
label=score + " valid", ls='-',
color= 'g' if score == 'AUC' else 'k',
)
plt.fill_between(train_sizes,
valid_scores.mean(axis=1) - np.sqrt(valid_scores.var()),
y2=valid_scores.mean(axis=1) + np.sqrt(valid_scores.var()),
alpha=0.3,
color='g' if score == 'AUC' else 'k',
zorder=-1
)
plt.legend(title=classifier)
plt.grid()
plt.xlabel("Sample Size")
plt.ylabel("Score")
plt.savefig(f"{classifier}_learning_curve.png", dpi=150)
```
### print the performance on the validation set:
```
for score, val in results['score_validation'].items():
print(f"score {score:s} on validation sample is {val:.5f} for {classifier:s}")
```
Compute expected background rate with observation time and number of fitted pulses in extrinsic sample:
```
T_obs = 66407.566110
n_tot_bkg_pulses = 2966
bkg_rate = (1. - results['score_validation']['Accuracy']) * n_tot_bkg_pulses / T_obs
print (f"Expected bkg rate of {classifier:s}: {bkg_rate:.5e} Hz")
```
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
from tensorflow import keras
def plot_series(time, series, format="-", start=0, end=None, label=None):
plt.plot(time[start:end], series[start:end], format, label=label)
plt.xlabel("Time")
plt.ylabel("Value")
if label:
plt.legend(fontsize=14)
plt.grid(True)
def trend(time, slope=0):
return slope * time
time = np.arange(4 * 365 + 1)
baseline = 10
series = trend(time, 0.1)
plt.figure(figsize=(10, 6))
plot_series(time, series)
plt.show()
def seasonal_pattern(season_time):
"""Just an arbitrary pattern, you can change it if you wish"""
return np.where(season_time < 0.4,
np.cos(season_time * 2 * np.pi),
1 / np.exp(3 * season_time))
def seasonality(time, period, amplitude=1, phase=0):
"""Repeats the same pattern at each period"""
season_time = ((time + phase) % period) / period
return amplitude * seasonal_pattern(season_time)
baseline = 10
amplitude = 40
series = seasonality(time, period=365, amplitude=amplitude)
plt.figure(figsize=(10, 6))
plot_series(time, series)
plt.show()
slope = 0.05
series = baseline + trend(time, slope) + seasonality(time, period=365, amplitude=amplitude)
plt.figure(figsize=(10, 6))
plot_series(time, series)
plt.show()
def white_noise(time, noise_level=1, seed=None):
rnd = np.random.RandomState(seed)
return rnd.randn(len(time)) * noise_level
noise_level = 5
noise = white_noise(time, noise_level, seed=42)
plt.figure(figsize=(10, 6))
plot_series(time, noise)
plt.show()
series += noise
plt.figure(figsize=(10, 6))
plot_series(time, series)
plt.show()
split_time = 1000
time_train = time[:split_time]
x_train = series[:split_time]
time_valid = time[split_time:]
x_valid = series[split_time:]
def autocorrelation(time, amplitude, seed=None):
rnd = np.random.RandomState(seed)
φ1 = 0.5
φ2 = -0.1
ar = rnd.randn(len(time) + 50)
ar[:50] = 100
for step in range(50, len(time) + 50):
ar[step] += φ1 * ar[step - 50]
ar[step] += φ2 * ar[step - 33]
return ar[50:] * amplitude
def autocorrelation(time, amplitude, seed=None):
rnd = np.random.RandomState(seed)
φ = 0.8
ar = rnd.randn(len(time) + 1)
for step in range(1, len(time) + 1):
ar[step] += φ * ar[step - 1]
return ar[1:] * amplitude
series = autocorrelation(time, 10, seed=42)
plot_series(time[:200], series[:200])
plt.show()
series = autocorrelation(time, 10, seed=42) + trend(time, 2)
plot_series(time[:200], series[:200])
plt.show()
series = autocorrelation(time, 10, seed=42) + seasonality(time, period=50, amplitude=150) + trend(time, 2)
plot_series(time[:200], series[:200])
plt.show()
series = autocorrelation(time, 10, seed=42) + seasonality(time, period=50, amplitude=150) + trend(time, 2)
series2 = autocorrelation(time, 5, seed=42) + seasonality(time, period=50, amplitude=2) + trend(time, -1) + 550
series[200:] = series2[200:]
#series += noise(time, 30)
plot_series(time[:300], series[:300])
plt.show()
def impulses(time, num_impulses, amplitude=1, seed=None):
rnd = np.random.RandomState(seed)
impulse_indices = rnd.randint(len(time), size=10)
series = np.zeros(len(time))
for index in impulse_indices:
series[index] += rnd.rand() * amplitude
return series
series = impulses(time, 10, seed=42)
plot_series(time, series)
plt.show()
def autocorrelation(source, φs):
ar = source.copy()
max_lag = len(φs)
for step, value in enumerate(source):
for lag, φ in φs.items():
if step - lag > 0:
ar[step] += φ * ar[step - lag]
return ar
signal = impulses(time, 10, seed=42)
series = autocorrelation(signal, {1: 0.99})
plot_series(time, series)
plt.plot(time, signal, "k-")
plt.show()
signal = impulses(time, 10, seed=42)
series = autocorrelation(signal, {1: 0.70, 50: 0.2})
plot_series(time, series)
plt.plot(time, signal, "k-")
plt.show()
series_diff1 = series[1:] - series[:-1]
plot_series(time[1:], series_diff1)
from pandas.plotting import autocorrelation_plot
autocorrelation_plot(series)
from statsmodels.tsa.arima_model import ARIMA
model = ARIMA(series, order=(5, 1, 0))
model_fit = model.fit(disp=0)
print(model_fit.summary())
import pandas as pd
df = pd.read_csv("sunspots.csv", parse_dates=["Date"], index_col="Date")
series = df["Monthly Mean Total Sunspot Number"].asfreq("1M")
series.head()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/dlmacedo/starter-academic/blob/master/4The_ultimate_guide_to_Encoder_Decoder_Models_4_4.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
%%capture
!pip install -qq git+https://github.com/huggingface/transformers.git
```
# **The ultimate guide to Encoder Decoder Models 4/4**
The *transformer-based* encoder-decoder model was introduced by Vaswani et al. in the famous [Attention is all you need paper](https://arxiv.org/abs/1706.03762) and is today the *de-facto* standard encoder-decoder architecture in natural language processing (NLP).
Recently, there has been a lot of research on different *pre-training* objectives for transformer-based encoder-decoder models, *e.g.* T5, Bart, Pegasus, ProphetNet, Marge, *etc*..., but the model architecture has stayed largely the same.
The goal of the blog post is to give an **in-detail** explanation of **how** the transformer-based encoder-decoder architecture models *sequence-to-sequence* problems. We will focus on the mathematical model defined by the architecture and how the model can be used in inference. Along the way, we will give some background on sequence-to-sequence models in NLP and break down the *transformer-based* encoder-decoder architecture into its **encoder** and **decoder** part. We provide many illustrations and establish the link
between the theory of *transformer-based* encoder-decoder models and their practical usage in 🤗Transformers for inference.
Note that this blog post does *not* explain how such models can be trained - this will be the topic of a future blog post.
Transformer-based encoder-decoder models are the result of years of research on *representation learning* and *model architectures*.
This notebook provides a short summary of the history of neural encoder-decoder models. For more context, the reader is advised to read this awesome [blog post](https://ruder.io/a-review-of-the-recent-history-of-nlp/) by Sebastion Ruder. Additionally, a basic understanding of the *self-attention architecture* is recommended.
The following blog post by Jay Alammar serves as a good refresher on the original Transformer model [here](http://jalammar.github.io/illustrated-transformer/).
At the time of writing this notebook, 🤗Transformers comprises the encoder-decoder models *T5*, *Bart*, *MarianMT*, and *Pegasus*, which are summarized in the docs under [model summaries](https://huggingface.co/transformers/model_summary.html#sequence-to-sequence-models).
The notebook is divided into four parts:
- **Background** - *A short history of neural encoder-decoder models is given with a focus on on RNN-based models.* - [click here](https://colab.research.google.com/drive/18ZBlS4tSqSeTzZAVFxfpNDb_SrZfAOMf?usp=sharing)
- **Encoder-Decoder** - *The transformer-based encoder-decoder model is presented and it is explained how the model is used for inference.* - [click here](https://colab.research.google.com/drive/1XpKHijllH11nAEdPcQvkpYHCVnQikm9G?usp=sharing)
- **Encoder** - *The encoder part of the model is explained in detail.* - [click here](https://colab.research.google.com/drive/1HJhnWMFizEKKWEAb-k7QDBv4c03hXbCR?usp=sharing)
- **Decoder** - *The decoder part of the model is explained in detail.*
Each part builds upon the previous part, but can also be read on its own.
## **Decoder**
As mentioned in the *Encoder-Decoder* section, the *transformer-based* decoder defines the conditional probability distribution of a target sequence given the contextualized encoding sequence:
$$ p_{\theta_{dec}}(\mathbf{Y}_{1: m} | \mathbf{\overline{X}}_{1:n}), $$
which by Bayes' rule can be decomposed into a product of conditional distributions of the next target vector given the contextualized encoding sequence and all previous target vectors:
$$ p_{\theta_{dec}}(\mathbf{Y}_{1:m} | \mathbf{\overline{X}}_{1:n}) = \prod_{i=1}^{m} p_{\theta_{dec}}(\mathbf{y}_i | \mathbf{Y}_{0: i-1}, \mathbf{\overline{X}}_{1:n}). $$
Let's first understand how the transformer-based decoder defines a probability distribution. The transformer-based decoder is a stack of *decoder blocks* followed by a dense layer, the "LM head".
The stack of decoder blocks maps the contextualized encoding sequence $\mathbf{\overline{X}}_{1:n}$ and a target vector sequence prepended by the $\text{BOS}$ vector and cut to the last target vector, *i.e.* $\mathbf{Y}_{0:i-1}$, to an encoded sequence of target vectors $\mathbf{\overline{Y}}_{0: i-1}$. Then, the "LM head" maps the encoded sequence of target vectors $\mathbf{\overline{Y}}_{0: i-1}$ to a sequence of logit vectors $\mathbf{L}_{1:n} = \mathbf{l}_1, \ldots, \mathbf{l}_n$, whereas the dimensionality of each logit vector $\mathbf{l}_i$ corresponds to the size of the vocabulary. This way, for each $i \in \{1, \ldots, n\}$ a probability distribution over the whole vocabulary can be obtained by applying a softmax operation on $\mathbf{l}_i$. These distributions define the conditional distribution:
$$p_{\theta_{dec}}(\mathbf{y}_i | \mathbf{Y}_{0: i-1}, \mathbf{\overline{X}}_{1:n}), \forall i \in \{1, \ldots, n\},$$
respectively. The "LM head" is often tied to the transpose of the word embedding matrix, *i.e.* $\mathbf{W}_{\text{emb}}^{\intercal} = \left[\mathbf{y}^1, \ldots, \mathbf{y}^{\text{vocab}}\right]^{\intercal}$ ${}^1$. Intuitively this means that for all $i \in \{0, \ldots, n - 1\}$ the "LM Head" layer compares the encoded output vector $\mathbf{\overline{y}}_i$ to all word embeddings in the vocabulary $\mathbf{y}^1, \ldots, \mathbf{y}^{\text{vocab}}$ so that the logit vector $\mathbf{l}_{i+1}$ represents the similarity scores between the encoded output vector and each word embedding. The softmax operation simply transformers the similarity scores to a probability distribution. For each $i \in \{1, \ldots, n\}$, the following equations hold:
$$ p_{\theta_{dec}}(\mathbf{y} | \mathbf{\overline{X}}_{1:n}, \mathbf{Y}_{0:i-1})$$
$$ = \text{Softmax}(f_{\theta_{\text{dec}}}(\mathbf{\overline{X}}_{1:n}, \mathbf{Y}_{0:i-1}))$$
$$ = \text{Softmax}(\mathbf{W}_{\text{emb}}^{\intercal} \mathbf{\overline{y}}_{i-1})$$
$$ = \text{Softmax}(\mathbf{l}_i). $$
Putting it all together, in order to model the conditional distribution of a target vector sequence $\mathbf{Y}_{1: m}$, the target vectors $\mathbf{Y}_{1:m-1}$ prepended by the special $\text{BOS}$ vector, *i.e.* $\mathbf{y}_0$, are first mapped together with the contextualized encoding sequence $\mathbf{\overline{X}}_{1:n}$ to the logit vector sequence $\mathbf{L}_{1:m}$.
Consequently, each logit target vector $\mathbf{l}_i$ is transformed into a conditional probability distribution of the target vector $\mathbf{y}_i$ using the softmax operation. Finally, the conditional probabilities of all target vectors $\mathbf{y}_1, \ldots, \mathbf{y}_m$ multiplied together to yield the conditional probability of the complete target vector sequence:
$$ p_{\theta_{dec}}(\mathbf{Y}_{1:m} | \mathbf{\overline{X}}_{1:n}) = \prod_{i=1}^{m} p_{\theta_{dec}}(\mathbf{y}_i | \mathbf{Y}_{0: i-1}, \mathbf{\overline{X}}_{1:n}).$$
In contrast to transformer-based encoders, in transformer-based decoders, the encoded output vector $\mathbf{\overline{y}}_i$ should be a good representation of the *next* target vector $\mathbf{y}_{i+1}$ and not of the input vector itself. Additionally, the encoded output vector $\mathbf{\overline{y}}_i$ should be conditioned on all contextualized encoding sequence $\mathbf{\overline{X}}_{1:n}$. To meet these requirements each decoder block consists of a **uni-directional** self-attention layer, followed by a **cross-attention** layer and two feed-forward layers ${}^2$.
The uni-directional self-attention layer puts each of its input vectors $\mathbf{y'}_j$ only into relation with all previous input vectors $\mathbf{y'}_i, \text{ with } i \le q$ for all $j \in \{1, \ldots, n\}$ to model the probability distribution of the next target vectors.
The cross-attention layer puts each of its input vectors $\mathbf{y''}_j$ into relation with all contextualized encoding vectors $\mathbf{\overline{X}}_{1:n}$ to condition the probability distribution of the next target vectors on the input of the encoder as well.
Alright, let's visualize the *transformer-based* decoder for our English to German translation example.
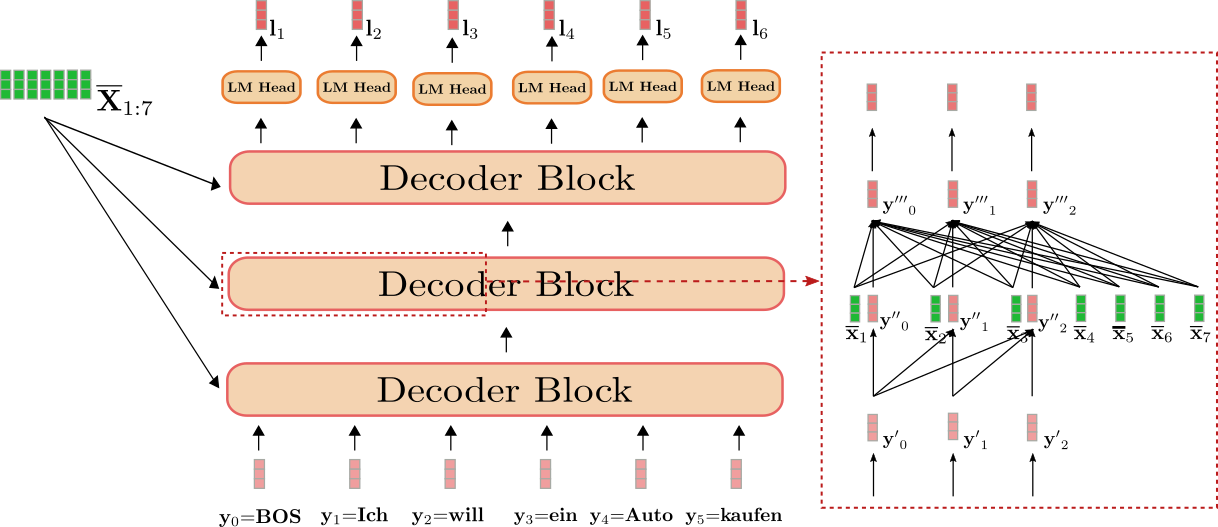
We can see that the decoder maps the input $\mathbf{Y}_{0:5}$ "BOS", "Ich", "will", "ein", "Auto", "kaufen" (shown in light red) together with the contextualized sequence of "I", "want", "to", "buy", "a", "car", "EOS", *i.e.* $\mathbf{\overline{X}}_{1:7}$ (shown in dark green) to the logit vectors $\mathbf{L}_{1:6}$ (shown in dark red).
Applying a softmax operation on each $\mathbf{l}_1, \mathbf{l}_2, \ldots, \mathbf{l}_5$ can thus define the conditional probability distributions:
$$ p_{\theta_{dec}}(\mathbf{y} | \text{BOS}, \mathbf{\overline{X}}_{1:7}), $$
$$ p_{\theta_{dec}}(\mathbf{y} | \text{BOS Ich}, \mathbf{\overline{X}}_{1:7}), $$
$$ \ldots, $$
$$ p_{\theta_{dec}}(\mathbf{y} | \text{BOS Ich will ein Auto kaufen}, \mathbf{\overline{X}}_{1:7}). $$
The overall conditional probability of:
$$ p_{\theta_{dec}}(\text{Ich will ein Auto kaufen EOS} | \mathbf{\overline{X}}_{1:n})$$
can therefore be computed as the following product:
$$ p_{\theta_{dec}}(\text{Ich} | \text{BOS}, \mathbf{\overline{X}}_{1:7}) \times \ldots \times p_{\theta_{dec}}(\text{EOS} | \text{BOS Ich will ein Auto kaufen}, \mathbf{\overline{X}}_{1:7}). $$
The red box on the right shows a decoder block for the first three target vectors $\mathbf{y}_0, \mathbf{y}_1, \mathbf{y}_2$. In the lower part, the uni-directional self-attention mechanism is illustrated and in the middle, the cross-attention mechanism is illustrated. Let's first focus on uni-directional self-attention.
As in bi-directional self-attention, in uni-directional self-attention, the query vectors $\mathbf{q}_0, \ldots, \mathbf{q}_{m-1}$ (shown in purple below), key vectors $\mathbf{k}_0, \ldots, \mathbf{k}_{m-1}$ (shown in orange below), and value vectors $\mathbf{v}_0, \ldots, \mathbf{v}_{m-1}$ (shown in blue below) are projected from their respective input vectors $\mathbf{y'}_0, \ldots, \mathbf{y}_{m-1}$ (shown in light red below). However, in uni-directional self-attention, each query vector $\mathbf{q}_i$ is compared *only* to its respective key vector and all previous ones, namely $\mathbf{k}_0, \ldots, \mathbf{k}_i$ to yield the respective *attention weights*.
This prevents an output vector $\mathbf{y''}_j$ (shown in dark red below) to include any information about the following input vector $\mathbf{y}_i, \text{ with } i > 1$ for all $j \in \{0, \ldots, m - 1 \}$. As is the case in bi-directional self-attention, the attention weights are then multiplied by their respective value vectors and summed together.
We can summarize uni-directional self-attention as follows:
$$\mathbf{y''}_i = \mathbf{V}_{0: i} \textbf{Softmax}(\mathbf{K}_{0: i}^\intercal \mathbf{q}_i) + \mathbf{y'}_i. $$
Note that the index range of the key and value vectors is $0:i$ instead of $0: m-1$ which would be the range of the key vectors in bi-directional self-attention.
Let's illustrate uni-directional self-attention for the input vector $\mathbf{y'}_1$ for our example above.
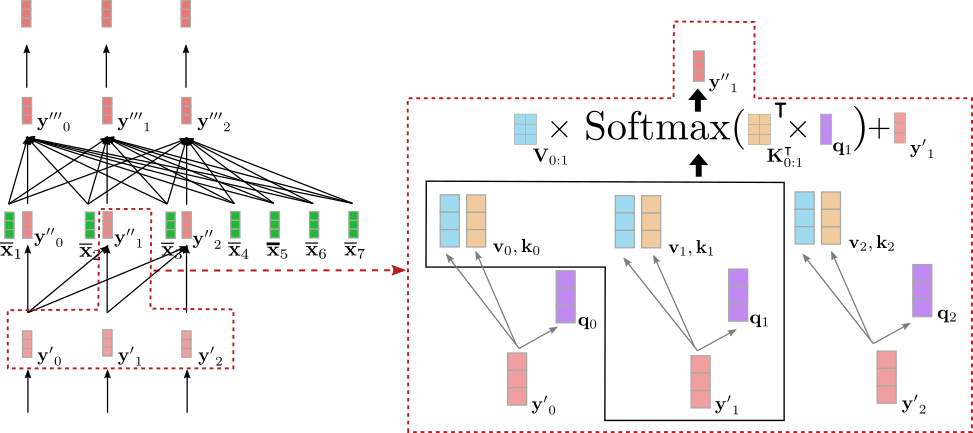
As can be seen $\mathbf{y''}_1$ only depends on $\mathbf{y'}_0$ and $\mathbf{y'}_1$. Therefore, we put the vector representation of the word "Ich", *i.e.* $\mathbf{y'}_1$ only into relation with itself and the "BOS" target vector, *i.e.* $\mathbf{y'}_0$, but **not** with the vector representation of the word "will", *i.e.* $\mathbf{y'}_2$.
So why is it important that we use uni-directional self-attention in the decoder instead of bi-directional self-attention?
As stated above, a transformer-based decoder defines a mapping from a sequence of input vector $\mathbf{Y}_{0: m-1}$ to the logits corresponding to the **next** decoder input vectors, namely $\mathbf{L}_{1:m}$.
In our example, this means, *e.g.* that the input vector $\mathbf{y}_1$ = "Ich" is mapped to the logit vector $\mathbf{l}_2$, which is then used to predict the input vector $\mathbf{y}_2$. Thus, if $\mathbf{y'}_1$ would have access to the following input vectors $\mathbf{Y'}_{2:5}$, the decoder would simply copy the vector representation of "will", *i.e.* $\mathbf{y'}_2$, to be its output $\mathbf{y''}_1$. This would be forwarded to the last layer so that the encoded output vector $\mathbf{\overline{y}}_1$ would essentially just correspond to the vector representation $\mathbf{y}_2$.
This is obviously disadvantageous as the transformer-based decoder would never learn to predict the next word given all previous words, but just copy the target vector $\mathbf{y}_i$ through the network to $\mathbf{\overline{y}}_{i-1}$ for all $i \in \{1, \ldots, m \}$. In order to define a conditional distribution of the next target vector, the distribution cannot be conditioned on the next target vector itself. It does not make much sense to predict $\mathbf{y}_i$ from $p(\mathbf{y} | \mathbf{Y}_{0:i}, \mathbf{\overline{X}})$ because the distribution is conditioned on the target vector it is supposed to model.
The uni-directional self-attention architecture, therefore, allows us to define a *causal* probability distribution, which is necessary to effectively model a conditional distribution of the next target vector.
Great! Now we can move to the layer that connects the encoder and decoder - the *cross-attention* mechanism!
The cross-attention layer takes two vector sequences as inputs: the outputs of the uni-directional self-attention layer, *i.e.* $\mathbf{Y''}_{0: m-1}$ and the contextualized encoding vectors $\mathbf{\overline{X}}_{1:n}$.
As in the self-attention layer, the query vectors $\mathbf{q}_0, \ldots, \mathbf{q}_{m-1}$ are projections of the output vectors of the previous layer, *i.e.* $\mathbf{Y''}_{0: m-1}$. However, the key and value vectors $\mathbf{k}_0, \ldots, \mathbf{k}_{m-1}$ and $\mathbf{v}_0, \ldots, \mathbf{v}_{m-1}$ are projections of the contextualized encoding vectors $\mathbf{\overline{X}}_{1:n}$. Having defined key, value, and query vectors, a query vector $\mathbf{q}_i$ is then compared to *all* key vectors and the corresponding score is used to weight the respective value vectors, just as is the case for *bi-directional* self-attention to give the output vector $\mathbf{y'''}_i$ for all $i \in \{0, \ldots, m-1\}. Cross-attention can be summarized as follows:
$$
\mathbf{y'''}_i = \mathbf{V}_{1:n} \textbf{Softmax}(\mathbf{K}_{1: n}^\intercal \mathbf{q}_i) + \mathbf{y''}_i.
$$
Note that the index range of the key and value vectors is $1:n$ corresponding to the number of contextualized encoding vectors.
Let's visualize the cross-attention mechanism Let's for the input vector $\mathbf{y''}_1$ for our example above.
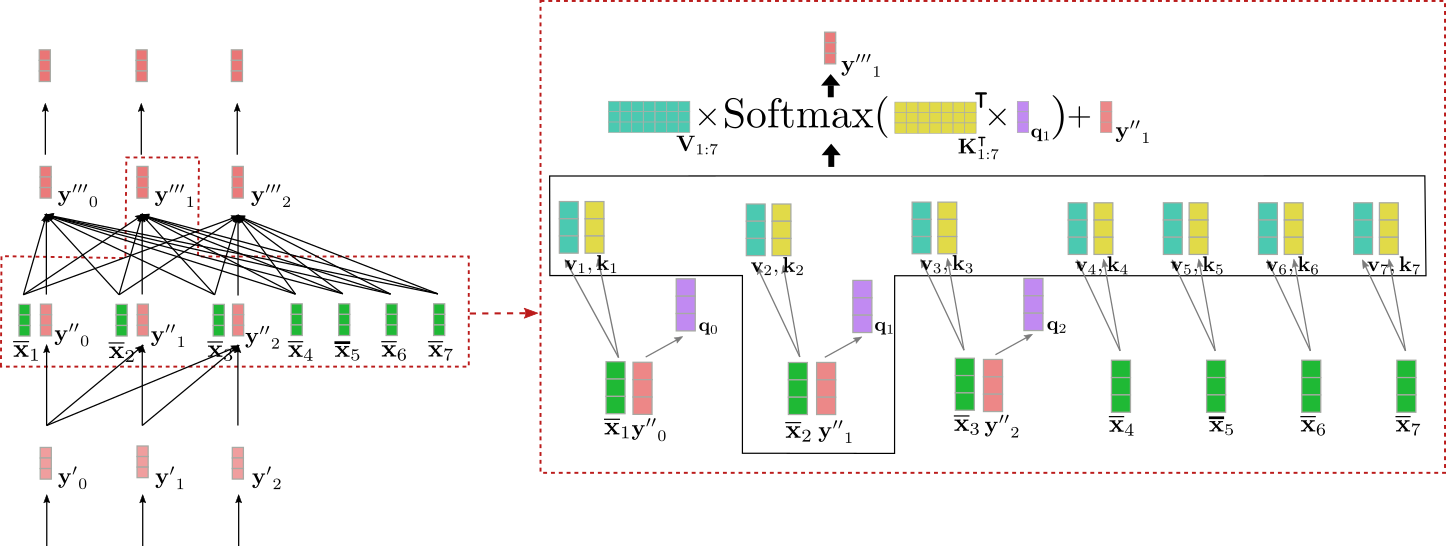
We can see that the query vector $\mathbf{q}_1$ (shown in purple) is derived from $\mathbf{y''}_1$ and therefore relies on a vector representation of the word "Ich". The query vector $\mathbf{q}_1$ (shown in red) is then compared to the key vectors $\mathbf{k}_1, \ldots, \mathbf{k}_7$ (shown in yellow) corresponding to the contextual encoding representation of all encoder input vectors $\mathbf{X}_{1:n}$ = "I want to buy a car EOS". This puts the vector representation of "Ich" into direct relation with all encoder input vectors. Finally, the attention weights are multiplied by the value vectors $\mathbf{v}_1, \ldots, \mathbf{v}_7$ (shown in turquoise) to yield in addition to the input vector $\mathbf{y''}_1$ the output vector $\mathbf{y'''}_1$ (shown in dark red).
So intuitively, what happens here exactly?
Each output vector $\mathbf{y'}_i$ is a weighted sum of all value projections of the encoder inputs $\mathbf{v}_{1}, \ldots, \mathbf{v}_7$ plus the input vector itself $\mathbf{y}_i$ (*c.f.* illustrated formula above). The key mechanism to understand is the following: Depending on how similar a query projection of the *input decoder vector* $\mathbf{q}_i$ is to a key projection of the *encoder input vector* $\mathbf{k}_j$, the more important is the value projection of the encoder input vector $\mathbf{v}_j$. In loose terms this means, the more "related" a decoder input representation is to an encoder input representation, the more does the input representation influence the decoder output representation.
Cool! Now we can see how this architecture nicely conditions each output vector $\mathbf{y'''}_i$ on the interaction between the encoder input vectors $\mathbf{\overline{X}}_{1:n}$ and the input vector $\mathbf{y''}_i$.
Another important observation at this point is that the architecture is completely independent of the number $n$ of contextualized encoding vectors $\mathbf{\overline{X}}_{1:n}$ on which the output vector $\mathbf{y'''}_i$ is conditioned on. All projection matrices $\mathbf{W}^{\text{cross}}_{k}$ and $\mathbf{W}^{\text{cross}}_{v}$ to derive the key vectors $\mathbf{k}_1, \ldots, \mathbf{k}_n$ and the value vectors $\mathbf{v}_1, \ldots, \mathbf{v}_n$ respectively are shared across all positions $1, \ldots, n$ and all value vectors $\mathbf{v}_1, \ldots, \mathbf{v}_n$ are summed together to a single weighted averaged vector.
Now it becomes obvious as well, why the transformer-based decoder does not suffer from the long-range dependency problem, the RNN-based decoder suffers from. Because each decoder logit vector is *directly* dependent on every single encoded output vector, the number of mathematical operations to compare the first encoded output vector and the last decoder logit vector amounts essentially to just one.
To conclude, the uni-directional self-attention layer is responsible for conditioning each output vector on all previous decoder input vectors and the current input vector and the cross-attention layer is responsible to further condition each output vector on all encoded input vectors.
To verify our theoretical understanding, let's continue our code example from the encoder section above.
---
${}^1$ The word embedding matrix $\mathbf{W}_{\text{emb}}$ gives each input word a unique *context-independent* vector representation. This matrix is often fixed as the "LM Head" layer. However, the "LM Head" layer can very well consist of a completely independent "encoded vector-to-logit" weight mapping.
${}^2$ Again, an in-detail explanation of the role the feed-forward layers play in transformer-based models is out-of-scope for this notebook. It is argued in [Yun et. al, (2017)](https://arxiv.org/pdf/1912.10077.pdf) that feed-forward layers are crucial to map each contextual vector $\mathbf{x'}_i$ individually to the desired output space, which the *self-attention* layer does not manage to do on its own. It should be noted here, that each output token $\mathbf{x'}$ is processed by the same feed-forward layer. For more detail, the reader is advised to read the paper.
```
from transformers import MarianMTModel, MarianTokenizer
import torch
tokenizer = MarianTokenizer.from_pretrained("Helsinki-NLP/opus-mt-en-de")
model = MarianMTModel.from_pretrained("Helsinki-NLP/opus-mt-en-de")
embeddings = model.get_input_embeddings()
# get encoded input vectors
input_ids = tokenizer("I want to buy a car", return_tensors="pt").input_ids
encoded_output_vectors = model.base_model.encoder(input_ids, return_dict=True).last_hidden_state
# create ids of encoded input vectors
decoder_input_ids = tokenizer("<pad> Ich will ein", return_tensors="pt", add_special_tokens=False).input_ids
# pass decoder input_ids and encoded input vectors to decoder
decoder_output_vectors = model.base_model.decoder(decoder_input_ids, encoded_output_vectors, None, None, None, return_dict=True).last_hidden_state
# derive embeddings by multiplying decoder outputs with embedding weights
lm_logits = torch.nn.functional.linear(decoder_output_vectors, embeddings.weight, bias=model.final_logits_bias)
# change the decoder input slightly
decoder_input_ids_perturbed = tokenizer("</s> Ich will das", return_tensors="pt").input_ids
decoder_output_vectors_perturbed = model.base_model.decoder(decoder_input_ids, encoded_output_vectors, None, None, None, return_dict=True).last_hidden_state
lm_logits_perturbed = torch.nn.functional.linear(decoder_output_vectors_perturbed, embeddings.weight, bias=model.final_logits_bias)
# compare shape and encoding of first vector
print(f"Shape of decoder input vectors {embeddings(decoder_input_ids).shape}. Shape of decoder logits {lm_logits.shape}")
# compare values of word embedding of "I" for input_ids and perturbed input_ids
print("Is encoding for `Ich` equal to its perturbed version?: ", torch.allclose(lm_logits[0, 0], lm_logits_perturbed[0, 0], atol=1e-3))
```
We compare the output shape of the decoder input word embeddings, *i.e.* `embeddings(decoder_input_ids)` (corresponds to $\mathbf{Y}_{0: 4}$, here `<pad>` corresponds to BOS and "Ich will das" is tokenized to 4 tokens) with the dimensionality of the `lm_logits`(corresponds to $\mathbf{L}_{1:5}$).
Also, we have passed the word sequence "<pad> Ich will das" and a slightly perturbated version "<pad> Ich will das" together with the `encoder_output_vectors` through the encoder to check if the second `lm_logit`, corresponding to "Ich", differs when only the last word is changed in the input sequence ("ein" -> "das").
As expected the output shapes of the decoder input word embeddings and lm_logits, *i.e.* the dimensionality of $\mathbf{Y}_{0: 4}$ and $\mathbf{L}_{1:5}$ are different in the last dimension. While the sequence length is the same (=5), the dimensionality of a decoder input word embedding corresponds to `model.config.hidden_size`, whereas the dimensionality of a `lm_logit` corresponds to the vocabulary size `model.config.vocab_size`, as explained above.
Second, it can be noted that the values of the encoded output vector of $\mathbf{l}_1 = \text{"Ich"}$ are the same when the last word is changed from "ein" to "das". This however should not come as a surprise if one has understood uni-directional self-attention.
On a final side-note, *auto-regressive* models, such as GPT2, have the same architecture as *transformer-based* decoder models **if** one removes the cross-attention layer because stand-alone auto-regressive models are not conditioned on any encoder outputs.
So auto-regressive models are essentially the same as *auto-encoding* models but replace bi-directional attention with uni-directional attention. These models can also be pre-trained on massive open-domain text data to show impressive performances on natural language generation (NLG) tasks. In [Radford et al. (2019)](https://cdn.openai.com/better-language-models/language_models_are_unsupervised_multitask_learners.pdf), the authors show that a pre-trained GPT2 model can achieve SOTA or close to SOTA results on a variety of NLG tasks without much fine-tuning. All *auto-regressive* models of 🤗Transformers can be found [here](https://huggingface.co/transformers/model_summary.html#autoregressive-models).
Alright, that's it! Now, you should have gotten a good understanding of *transformer-based* encoder-decoder models and how to use them with the 🤗Transformers library.
Thanks a lot to Victor Sanh, Sasha Rush, Sam Shleifer, Oliver Åstrand,
Ted Moskovitz and Kristian Kyvik for giving valuable feedback.
| github_jupyter |
```
import numpy as np
from scipy.optimize import minimize
# ##############################################################################
# LoadData takes the file location for the yacht_hydrodynamics.data and returns
# the data set partitioned into a training set and a test set.
# the X matrix, deal with the month and day strings.
# Do not change this function!
# ##############################################################################
def loadData(df):
data = np.loadtxt(df)
Xraw = data[:,:-1]
# The regression task is to predict the residuary resistance per unit weight of displacement
yraw = (data[:,-1])[:, None]
X = (Xraw-Xraw.mean(axis=0))/np.std(Xraw, axis=0)
y = (yraw-yraw.mean(axis=0))/np.std(yraw, axis=0)
ind = range(X.shape[0])
test_ind = ind[0::4] # take every fourth observation for the test set
train_ind = list(set(ind)-set(test_ind))
X_test = X[test_ind]
X_train = X[train_ind]
y_test = y[test_ind]
y_train = y[train_ind]
return X_train, y_train, X_test, y_test
# ##############################################################################
# Returns a single sample from a multivariate Gaussian with mean and cov.
# ##############################################################################
def multivariateGaussianDraw(mean, cov):
#n = mean.shape[0]
#sample = np.zeros((mean.shape[0], 1)) # This is only a placeholder
# Task 2:
# TODO: Implement a draw from a multivariate Gaussian here
# K = np.linalg.cholesky(cov + 1e-15*np.eye(n))
# sample = np.dot(L, np.random.normal(size=(n,1)))
sample = np.random.multivariate_normal(mean, cov, 1)
# Return drawn sample
return sample
# ##############################################################################
# RadialBasisFunction for the kernel function
# k(x,x') = s2_f*exp(-norm(x,x')^2/(2l^2)). If s2_n is provided, then s2_n is
# added to the elements along the main diagonal, and the kernel function is for
# the distribution of y,y* not f, f*.
# ##############################################################################
class RadialBasisFunction():
def __init__(self, params):
self.ln_sigma_f = params[0]
self.ln_length_scale = params[1]
self.ln_sigma_n = params[2]
self.sigma2_f = np.exp(2*self.ln_sigma_f)
self.sigma2_n = np.exp(2*self.ln_sigma_n)
self.length_scale = np.exp(self.ln_length_scale)
def setParams(self, params):
self.ln_sigma_f = params[0]
self.ln_length_scale = params[1]
self.ln_sigma_n = params[2]
self.sigma2_f = np.exp(2*self.ln_sigma_f)
self.sigma2_n = np.exp(2*self.ln_sigma_n)
self.length_scale = np.exp(self.ln_length_scale)
def getParams(self):
return np.array([self.ln_sigma_f, self.ln_length_scale, self.ln_sigma_n])
def getParamsExp(self):
return np.array([self.sigma2_f, self.length_scale, self.sigma2_n])
# ##########################################################################
# covMatrix computes the covariance matrix for the provided matrix X using
# the RBF. If two matrices are provided, for a training set and a test set,
# then covMatrix computes the covariance matrix between all inputs in the
# training and test set.
# ##########################################################################
def covMatrix(self, X, Xa=None):
if Xa is not None:
X_aug = np.zeros((X.shape[0]+Xa.shape[0], X.shape[1]))
X_aug[:X.shape[0], :X.shape[1]] = X
X_aug[X.shape[0]:, :X.shape[1]] = Xa
X=X_aug
n = X.shape[0]
covMat = np.zeros((n,n))
# Task 1:
# TODO: Implement the covariance matrix here
for i in range(n):
for j in range(n):
sqdist = np.exp(-0.5 * (1/(self.length_scale)**2)* (np.linalg.norm(X[i,:]-X[j,:]))**2)
covMat[i,j] = self.sigma2_f * sqdist
# If additive Gaussian noise is provided, this adds the sigma2_n along
# the main diagonal. So the covariance matrix will be for [y y*]. If
# you want [y f*], simply subtract the noise from the lower right
# quadrant.
if self.sigma2_n is not None:
covMat += self.sigma2_n*np.identity(n)
# Return computed covariance matrix
return covMat
# ln_sigma_f = np.log(1.27)
# ln_length_scale = np.log(1)
# ln_sigma_n = np.log(0.3)
# basis = RadialBasisFunction([ln_sigma_f,ln_length_scale,ln_sigma_n])
# X = np.array([-1.50,-1.00,-0.7,-0.40, -0.25, 0.00]).reshape(-1,1)
#np.linalg.det(basis.covMatrix(X))
#X_train, y_train, X_test, y_test = loadData('yacht_hydrodynamics.data')
#basis = RadialBasisFunction([ 1.10442998 ,17.38230593 ,-10.10133779])
#np.linalg.det(basis.covMatrix(X_train))
class GaussianProcessRegression():
def __init__(self, X, y, k):
self.X = X
self.n = X.shape[0]
self.y = y
self.k = k
self.K = self.KMat(self.X)
self.L = np.linalg.cholesky(self.K)
# ##########################################################################
# Recomputes the covariance matrix and the inverse covariance
# matrix when new hyperparameters are provided.
# ##########################################################################
def KMat(self, X, params=None):
if params is not None:
self.k.setParams(params)
K = self.k.covMatrix(X)
self.K = K
self.L = np.linalg.cholesky(self.K)
return K
# ##########################################################################
# Computes the posterior mean of the Gaussian process regression and the
# covariance for a set of test points.
# NOTE: This should return predictions using the 'clean' (not noisy) covariance
# ##########################################################################
def predict(self, Xa):
mean_fa = np.zeros((Xa.shape[0], 1))
mean_fa = np.array(mean_fa.flatten())
cov_fa = np.zeros((Xa.shape[0], Xa.shape[0]))
# Task 3:
# TODO: compute the mean and covariance of the prediction
#beginning of my solution: ------->
na = Xa.shape[0]
Ktotal = self.k.covMatrix(self.X, Xa)
# Covariance between training sample points (without Gaussian noise)
Kxx = Ktotal[0:self.n,0:self.n] # + 1 * np.eye(10) if there is Gaussian noise
# Covariance between training and test points
Kxs = Ktotal[self.n:self.n+na, 0:self.n]
# Covariance between test points
Kss = Ktotal[self.n:self.n+na,self.n:self.n+na]
# The mean of the GP fit (note that @ is matrix multiplcation: A @ B is equivalent to np.matmul(A,B))
mean = Kxs @ np.linalg.inv(Kxx) @ self.y
for i in range(mean.shape[0]):
mean_fa[i] = (mean[i])[0]
# The covariance matrix of the GP fit
cov_fa = Kss - Kxs @ np.linalg.inv(Kxx) @ Kxs.T
# Return the mean and covariance
return mean_fa, cov_fa
# ##########################################################################
# Return negative log marginal likelihood of training set. Needs to be
# negative since the optimiser only minimises.
# ##########################################################################
def logMarginalLikelihood(self, params=None):
if params is not None:
self.KMat(self.X, params)
mll = 0
# Task 4:
# TODO: Calculate the log marginal likelihood ( mll ) of self.y
mll = 0.5* (self.y).T @ np.linalg.solve((self.L).T, np.linalg.solve(self.L, self.y)) + np.log(np.sum((self.L).diagonal())) + 0.5 * self.n * np.log(2 * np.pi)
# Return mll
return mll
# ##########################################################################
# Computes the gradients of the negative log marginal likelihood wrt each
# hyperparameter.
# ##########################################################################
def gradLogMarginalLikelihood(self, params=None):
if params is not None:
K = self.KMat(self.X, params)
grad_ln_sigma_f = grad_ln_length_scale = grad_ln_sigma_n = 0
# Task 5:
# TODO: calculate the gradients of the negative log marginal likelihood
# wrt. the hyperparameters
beta_ln_length_scale = np.zeros((self.n, self.n))
beta_ln_sigma_f = np.zeros((self.n, self.n))
beta_ln_sigma_n = np.zeros((self.n, self.n))
param = self.k.getParamsExp()
alpha = np.linalg.solve(self.K, self.y)
#inv = np.linalg.solve((self.L).T, np.linalg.solve(self.L, np.eye(self.n,self.n)))
inv = np.linalg.inv(self.K)
#L = self.L
for i in range(self.n):
for j in range(self.n):
beta_ln_length_scale[i,j] = (1/(param[1]**2)) * ((np.linalg.norm(self.X[i]-self.X[j]))**2) * param[0] * np.exp(-0.5*(np.linalg.norm(self.X[i]-self.X[j])**2)/(param[1]**2))
beta_ln_sigma_n[i,j] = 2 * param[2] if (i==j) else 0
beta_ln_sigma_f[i,j] = 2 * param[0] * (np.exp(-0.5*(np.linalg.norm(self.X[i]-self.X[j])**2)/(param[1]**2)))
#beta_ln_sigma_n =beta_ln_sigma_n @
grad_ln_sigma_f = -0.5 * np.trace((alpha @ alpha.T - inv) @ beta_ln_sigma_f)
grad_ln_length_scale = -0.5 * np.trace((alpha @ alpha.T - inv) @ beta_ln_length_scale)
grad_ln_sigma_n = -0.5 * np.trace((alpha @ alpha.T - inv) @ beta_ln_sigma_n)
# Combine gradients
gradients = np.array([grad_ln_sigma_f, grad_ln_length_scale, grad_ln_sigma_n])
# Return the gradients
return gradients
# ##########################################################################
# Computes the mean squared error between two input vectors.
# ##########################################################################
def mse(self, ya, fbar):
mse = 0
# Task 7:
# TODO: Implement the MSE between ya and fbar
mse = np.sum((ya[i,:] - fbar[i,:])**2)/(ya.shape[0])
# Return mse
return mse
# ##########################################################################
# Computes the mean standardised log loss.
# ##########################################################################
def msll(self, ya, fbar, cov):
msll = 0
# Task 7:
# TODO: Implement MSLL of the prediction fbar, cov given the target ya
return msll
# ##########################################################################
# Minimises the negative log marginal likelihood on the training set to find
# the optimal hyperparameters using BFGS.
# ##########################################################################
def optimize(self, params, disp=True):
#params = np.array([0.5*np.log(1.0), np.log(1.0), 0.5*np.log(0.5)])
res = minimize(self.logMarginalLikelihood, params, method ='BFGS', jac = self.gradLogMarginalLikelihood, options = {'disp':disp})
return res.x
if __name__ == '__main__':
np.random.seed(42)
##########################
# You can put your tests here - marking
# will be based on importing this code and calling
# specific functions with custom input.
##########################
X_train, y_train, X_test, y_test = loadData('yacht_hydrodynamics.data')
params = [0.5*np.log(1.0), np.log(0.1), 0.5*np.log(0.5)]
basis = RadialBasisFunction(params)
basis.sigma2_f
GPR = GaussianProcessRegression(X_train, y_train,basis)
basis.covMatrix(X_train)
(basis.covMatrix(X_train, X_test)).shape
(GPR.KMat(X_train)).shape
mean, cov = GPR.predict(X_test)
mean.shape
cov.shape
GPR.logMarginalLikelihood()
GPR.gradLogMarginalLikelihood()
sample = multivariateGaussianDraw(mean, cov)
mean = [0, 0]
cov = [[1, 0], [0, 100]]
import matplotlib.pyplot as plt
x, y = np.random.multivariate_normal(mean, cov, 5000).T
plt.plot(x, y, 'x')
plt.axis('equal')
plt.show()
sigma2_f = np.exp(2*0.5*np.log(1.0))
length_scale= np.exp(np.log(0.1))
sigma2_n = np.exp(2*0.5*np.log(0.5))
def covariance(X,length_scale,sigma2_n,sigma2_f):
n = X.shape[0]
covMat = np.zeros((n,n))
for i in range(n):
for j in range(n):
sqdist = np.exp(-0.5 * (1/(length_scale)**2)* (np.linalg.norm(X[i,:]-X[j,:]))**2)
covMat[i,j] = sigma2_f * sqdist
covMat += sigma2_n*np.identity(n)
return covMat
def mll(X,y,length_scale,sigma2_n,sigma2_f):
n = X.shape[0]
K = covariance(X,length_scale,sigma2_n,sigma2_f)
L = np.linalg.cholesky(K)
mll = 0.5* y.T @ np.linalg.solve(L.T, np.linalg.solve(L, y)) + np.log(np.sum(L.diagonal())) + 0.5 * n * np.log(2 * np.pi)
return mll
# def gradient_checker():
# pass
covariance(X_train,length_scale,sigma2_n,sigma2_f).shape
h = 1e-6
Gsigmaf_h = mll(X_train,y_train,length_scale,sigma2_n,sigma2_f + h)
Gsigmaf = mll(X_train,y_train,length_scale,sigma2_n,sigma2_f)
(Gsigmaf_h - Gsigmaf)/h
mll(X_train,y_train,length_scale,sigma2_n,sigma2_f)
Gsigman_h = mll(X_train,y_train,length_scale,sigma2_n+h,sigma2_f)
Gsigman = mll(X_train,y_train,length_scale,sigma2_n,sigma2_f)
(Gsigman_h - Gsigman)/h
Gl_h = mll(X_train,y_train,length_scale + h,sigma2_n,sigma2_f)
Gl = mll(X_train,y_train,length_scale,sigma2_n,sigma2_f)
(Gl_h - Gl)/h
from scipy.optimize import check_grad
check_grad(GPR.logMarginalLikelihood(), GPR.gradLogMarginalLikelihood(), params)
```
| github_jupyter |
### Additional Uses
Remember what I said in the last lecture about some common patterns we can implement with context managers:
* Open - Close
* Change - Reset
* Start - Stop
The open file context manager is an example of the **Open - Close** pattern. But we have other ones as well.
#### Decimal Contexts
Decimals have a context which can be used to define many things, such as precision, rounding mechanism, etc.
By default, Decimals have a "global" context - i.e. one that will apply to any Decimal object by default:
```
import decimal
decimal.getcontext()
```
If we create a decimal object, then it will use those settings.
We can certainly change the properties of that global context:
```
decimal.getcontext().prec=14
decimal.getcontext()
```
And now the default (global) context has a precision set to 14.
Let's reset it back to 28:
```
decimal.getcontext().prec = 28
```
Suppose now that we just want to temporarily change something in the context - we would have to do something like this:
```
old_prec = decimal.getcontext().prec
decimal.getcontext().prec = 4
print(decimal.Decimal(1) / decimal.Decimal(3))
decimal.getcontext().prec = old_prec
print(decimal.Decimal(1) / decimal.Decimal(3))
```
Of course, this is kind of a pain to have to store the current value, set it to something new, and then remember to set it back to it's original value.
How about writing a context manager to do that seamlessly for us:
```
class precision:
def __init__(self, prec):
self.prec = prec
self.current_prec = decimal.getcontext().prec
def __enter__(self):
decimal.getcontext().prec = self.prec
def __exit__(self, exc_type, exc_value, exc_traceback):
decimal.getcontext().prec = self.current_prec
return False
```
Now we can do this:
```
with precision(3):
print(decimal.Decimal(1) / decimal.Decimal(3))
print(decimal.Decimal(1) / decimal.Decimal(3))
```
And as you can see, the precision was set back to it's original value once the context was exited.
In fact, the decimal class already has a context manager, and it's way better than ours, because we can set not only the precision, but anything else we want:
```
with decimal.localcontext() as ctx:
ctx.prec = 3
print(decimal.Decimal(1) / decimal.Decimal(3))
print(decimal.Decimal(1) / decimal.Decimal(3))
```
So this is an example of using a context manager for a **Change - Reset** type of situation.
#### Timing a with block
Here's another example of a **Start - Stop** type of context manager.
We'll create a context manager to time the code inside the `with` block:
```
from time import perf_counter, sleep
class Timer:
def __init__(self):
self.elapsed = 0
def __enter__(self):
self.start = perf_counter()
return self
def __exit__(self, exc_type, exc_value, exc_traceback):
self.stop = perf_counter()
self.elapsed = self.stop - self.start
return False
```
You'll note that this time we are returning the context manager itself from the `__enter__` statement. This will allow us to look at the `elapsed` property of the context manager once the `with` statement has finished running.
```
with Timer() as timer:
sleep(1)
print(timer.elapsed)
```
#### Redirecting stdout
Here we are going to temporarily redirect `stdout` to a file instead fo the console:
```
import sys
class OutToFile:
def __init__(self, fname):
self._fname = fname
self._current_stdout = sys.stdout
def __enter__(self):
self._file = open(self._fname, 'w')
sys.stdout = self._file
def __exit__(self, exc_type, exc_value, exc_tb):
sys.stdout = self._current_stdout
if self._file:
self._file.close()
return False
with OutToFile('test.txt'):
print('Line 1')
print('Line 2')
```
As you can see, no output happened on the console... Instead the output went to the file we specified. And our print statements will now output to the console again:
```
print('back to console output')
with open('test.txt') as f:
print(f.readlines())
```
#### HTML Tags
In this example, we're going to basically use a context manager to inject opening and closing html tags as we print to the console (of course we could redirect our prints somewhere else as we just saw!):
```
class Tag:
def __init__(self, tag):
self._tag = tag
def __enter__(self):
print(f'<{self._tag}>', end='')
def __exit__(self, exc_type, exc_value, exc_tb):
print(f'</{self._tag}>', end='')
return False
with Tag('p'):
print('some ', end='')
with Tag('b'):
print('bold', end='')
print(' text', end='')
```
#### Re-entrant Context Managers
We can also write context managers that can be re-entered in the sense that we can call `__enter__` and `__exit__` more than once on the **same** context manager.
These methods are called when a `with` statement is used, so we'll need to be able to get our hands on the context manager object itself - but that's easy, we just return `self` from the `__enter__` method.
Let's write a ListMaker context manager to do see how this works.
```
class ListMaker:
def __init__(self, title, prefix='- ', indent=3):
self._title = title
self._prefix = prefix
self._indent = indent
self._current_indent = 0
print(title)
def __enter__(self):
self._current_indent += self._indent
return self
def __exit__(self, exc_type, exc_value, exc_tb):
self._current_indent -= self._indent
return False
def print(self, arg):
s = ' ' * self._current_indent + self._prefix + str(arg)
print(s)
```
Because `__enter__` is returning `self`, the instance of the context manager, we can call `with` on that context manager and it will automatically call the `__enter__` and `__exit__` methods. Each time we run `__enter__` we increase the indentation, each time we run `__exit__` we decrease the indentation.
Our `print` method then takes that into account when it prints the requested string argument.
```
with ListMaker('Items') as lm:
lm.print('Item 1')
with lm:
lm.print('item 1a')
lm.print('item 1b')
lm.print('Item 2')
with lm:
lm.print('item 2a')
lm.print('item 2b')
```
Of course, we can easily redirect the output to a file instead, using the context manager we wrote earlier:
```
with OutToFile('my_list.txt'):
with ListMaker('Items') as lm:
lm.print('Item 1')
with lm:
lm.print('item 1a')
lm.print('item 1b')
lm.print('Item 2')
with lm:
lm.print('item 2a')
lm.print('item 2b')
with open('my_list.txt') as f:
for row in f:
print(row, end='')
```
| github_jupyter |
<h1>Sustainable Energy Transitions</h1>
<div>A project by <a href="http://www.ssgouridis.org" target="_blank" >Sgouris Sgouridis</a> and <a href="http://www.csaladen.es" target="_blank" >Dénes Csala</a> at <a href="http://www.masdar.ac.ae" target="_blank">Masdar Institute of Science and Technology</a></div>
<h2><br>Resource distribution notebook</h2>
This workbook will guide you through the mining, processing, formatting and saving the renewable resource distribution input data for the <a target="_blank" href="http://set.csaladen.es/set2.html">SET 2.0</a> model.
<p>This is document has been created using <a target="_blank" href="http://ipython.org/">IPython Notebook</a> in the <a target="_blank" href="https://store.continuum.io/cshop/anaconda/">Anaconda</a> distribution and it can be edited and run in <i>active</i> mode by clicking download in top right corner of this page. The code is partitioned into sections, called <i>cells</i>. When you are using this workbook in <i>active</i> mode, double-click on a cell to edit it and then run using <code>Ctrl + Enter</code>. Hitting <code>Shift + Enter</code> runs the code and steps into the next cell, while <code>Alt + Enter</code> runs the code and adds a new, empty cell. If you are running this notebook on a presonal computer, you will need a machine with at least <i>1GB</i> of memory (<i>2GB</i> recommended) and a processor of <i>1GHz</i>.
<h2>Data</h2>
<p>The outputs of this work are the calibration values for $\omega$, $\theta$ and $j$.
</p>
<h2>Processing</h2>
<p>The data processing has been done using the <a target="_blank" href="https://store.continuum.io/cshop/anaconda/">Anaconda</a> distribution of <a target="_blank" href="https://www.python.org/">Python 2.7</a> using the <i>IPython Notebook</i> editor. If you have not generated your own data, please download <a href="http://set.csaladen.es/data/results/index.html" target="_blank">all</a> results files, and run this <i>Iptyhon</i> notebook in the same folder. Then the data is loaded into a <a target="_blank" href="http://pandas.pydata.org/">pandas</a> dataframe, which represents the backbone of the data analysis. Numerical processing is done with <a target="_blank" href="http://www.numpy.org/">NumPy</a> and for plotting we use <a target="_blank" href="http://matplotlib.org/">matplotlib</a>. Please make sure you have all of these compoents set up in your <i>Python</i> installation.
</p>
<h1>Code</h1>
```
import matplotlib.pyplot as plt, pandas as pd, numpy as np, json, copy
%matplotlib inline
plt.style.use('ggplot')
```
## Country and region name converters
```
#country name converters
#EIA->pop
clist1={'North America':'Northern America',
'United States':'United States of America',
'Central & South America':'Latin America and the Caribbean',
'Bahamas, The':'Bahamas',
'Saint Vincent/Grenadines':'Saint Vincent and the Grenadines',
'Venezuela':'Venezuela (Bolivarian Republic of)',
'Macedonia':'The former Yugoslav Republic of Macedonia',
'Moldova':'Republic of Moldova',
'Russia':'Russian Federation',
'Iran':'Iran (Islamic Republic of)',
'Palestinian Territories':'State of Palestine',
'Syria':'Syrian Arab Republic',
'Yemen':'Yemen ',
'Congo (Brazzaville)':'Congo',
'Congo (Kinshasa)':'Democratic Republic of the Congo',
'Cote dIvoire (IvoryCoast)':"C\xc3\xb4te d'Ivoire",
'Gambia, The':'Gambia',
'Libya':'Libyan Arab Jamahiriya',
'Reunion':'R\xc3\xa9union',
'Somalia':'Somalia ',
'Sudan and South Sudan':'Sudan',
'Tanzania':'United Republic of Tanzania',
'Brunei':'Brunei Darussalam',
'Burma (Myanmar)':'Myanmar',
'Hong Kong':'China, Hong Kong Special Administrative Region',
'Korea, North':"Democratic People's Republic of Korea",
'Korea, South':'Republic of Korea',
'Laos':"Lao People's Democratic Republic",
'Macau':'China, Macao Special Administrative Region',
'Timor-Leste (East Timor)':'Timor-Leste',
'Virgin Islands, U.S.':'United States Virgin Islands',
'Vietnam':'Viet Nam'}
#BP->pop
clist2={u' European Union #':u'Europe',
u'Rep. of Congo (Brazzaville)':u'Congo (Brazzaville)',
'Republic of Ireland':'Ireland',
'China Hong Kong SAR':'China, Hong Kong Special Administrative Region',
u'Total Africa':u'Africa',
u'Total North America':u'Northern America',
u'Total S. & Cent. America':'Latin America and the Caribbean',
u'Total World':u'World',
u'Total World ':u'World',
'South Korea':'Republic of Korea',
u'Trinidad & Tobago':u'Trinidad and Tobago',
u'US':u'United States of America'}
#WD->pop
clist3={u"Cote d'Ivoire":"C\xc3\xb4te d'Ivoire",
u'Congo, Rep.':u'Congo (Brazzaville)',
u'Caribbean small states':'Carribean',
u'East Asia & Pacific (all income levels)':'Eastern Asia',
u'Egypt, Arab Rep.':'Egypt',
u'European Union':u'Europe',
u'Hong Kong SAR, China':u'China, Hong Kong Special Administrative Region',
u'Iran, Islamic Rep.':u'Iran (Islamic Republic of)',
u'Kyrgyz Republic':u'Kyrgyzstan',
u'Korea, Rep.':u'Republic of Korea',
u'Latin America & Caribbean (all income levels)':'Latin America and the Caribbean',
u'Macedonia, FYR':u'The former Yugoslav Republic of Macedonia',
u'Korea, Dem. Rep.':u"Democratic People's Republic of Korea",
u'South Asia':u'Southern Asia',
u'Sub-Saharan Africa (all income levels)':u'Sub-Saharan Africa',
u'Slovak Republic':u'Slovakia',
u'Venezuela, RB':u'Venezuela (Bolivarian Republic of)',
u'Yemen, Rep.':u'Yemen ',
u'Congo, Dem. Rep.':u'Democratic Republic of the Congo'}
#COMTRADE->pop
clist4={u"Bosnia Herzegovina":"Bosnia and Herzegovina",
u'Central African Rep.':u'Central African Republic',
u'China, Hong Kong SAR':u'China, Hong Kong Special Administrative Region',
u'China, Macao SAR':u'China, Macao Special Administrative Region',
u'Czech Rep.':u'Czech Republic',
u"Dem. People's Rep. of Korea":"Democratic People's Republic of Korea",
u'Dem. Rep. of the Congo':"Democratic Republic of the Congo",
u'Dominican Rep.':u'Dominican Republic',
u'Fmr Arab Rep. of Yemen':u'Yemen ',
u'Fmr Ethiopia':u'Ethiopia',
u'Fmr Fed. Rep. of Germany':u'Germany',
u'Fmr Panama, excl.Canal Zone':u'Panama',
u'Fmr Rep. of Vietnam':u'Viet Nam',
u"Lao People's Dem. Rep.":u"Lao People's Democratic Republic",
u'Occ. Palestinian Terr.':u'State of Palestine',
u'Rep. of Korea':u'Republic of Korea',
u'Rep. of Moldova':u'Republic of Moldova',
u'Serbia and Montenegro':u'Serbia',
u'US Virgin Isds':u'United States Virgin Islands',
u'Solomon Isds':u'Solomon Islands',
u'United Rep. of Tanzania':u'United Republic of Tanzania',
u'TFYR of Macedonia':u'The former Yugoslav Republic of Macedonia',
u'USA':u'United States of America',
u'USA (before 1981)':u'United States of America',
}
#Jacobson->pop
clist5={u"Korea, Democratic People's Republic of":"Democratic People's Republic of Korea",
u'All countries':u'World',
u"Cote d'Ivoire":"C\xc3\xb4te d'Ivoire",
u'Iran, Islamic Republic of':u'Iran (Islamic Republic of)',
u'Macedonia, Former Yugoslav Republic of':u'The former Yugoslav Republic of Macedonia',
u'Congo, Democratic Republic of':u"Democratic Republic of the Congo",
u'Korea, Republic of':u'Republic of Korea',
u'Tanzania, United Republic of':u'United Republic of Tanzania',
u'Moldova, Republic of':u'Republic of Moldova',
u'Hong Kong, China':u'China, Hong Kong Special Administrative Region',
u'All countries.1':"World"
}
#NREL solar->pop
clist6={u"Antigua & Barbuda":u'Antigua and Barbuda',
u"Bosnia & Herzegovina":u"Bosnia and Herzegovina",
u"Brunei":u'Brunei Darussalam',
u"Cote d'Ivoire":"C\xc3\xb4te d'Ivoire",
u"Iran":u'Iran (Islamic Republic of)',
u"Laos":u"Lao People's Democratic Republic",
u"Libya":'Libyan Arab Jamahiriya',
u"Moldova":u'Republic of Moldova',
u"North Korea":"Democratic People's Republic of Korea",
u"Reunion":'R\xc3\xa9union',
u'Sao Tome & Principe':u'Sao Tome and Principe',
u'Solomon Is.':u'Solomon Islands',
u'St. Lucia':u'Saint Lucia',
u'St. Vincent & the Grenadines':u'Saint Vincent and the Grenadines',
u'The Bahamas':u'Bahamas',
u'The Gambia':u'Gambia',
u'Virgin Is.':u'United States Virgin Islands',
u'West Bank':u'State of Palestine'
}
#NREL wind->pop
clist7={u"Antigua & Barbuda":u'Antigua and Barbuda',
u"Bosnia & Herzegovina":u"Bosnia and Herzegovina",
u'Occupied Palestinian Territory':u'State of Palestine',
u'China Macao SAR':u'China, Macao Special Administrative Region',
#"C\xc3\xb4te d'Ivoire":"C\xc3\xb4te d'Ivoire",
u'East Timor':u'Timor-Leste',
u'TFYR Macedonia':u'The former Yugoslav Republic of Macedonia',
u'IAM-country Total':u'World'
}
#country entroids->pop
clist8={u'Burma':'Myanmar',
u"Cote d'Ivoire":"C\xc3\xb4te d'Ivoire",
u'Republic of the Congo':u'Congo (Brazzaville)',
u'Reunion':'R\xc3\xa9union'
}
def cnc(country):
if country in clist1: return clist1[country]
elif country in clist2: return clist2[country]
elif country in clist3: return clist3[country]
elif country in clist4: return clist4[country]
elif country in clist5: return clist5[country]
elif country in clist6: return clist6[country]
elif country in clist7: return clist7[country]
elif country in clist8: return clist8[country]
else: return country
```
# Population
Consult the notebook entitled *pop.ipynb* for the details of mining the data from the UN statistics division online database. Due to being the reference database for country names cell, the cell below needs to be run first, before any other databases.
```
#use local path
#pop_path='https://dl.dropboxusercontent.com/u/531697/datarepo/netset/db/
pop_path='E:/Dropbox/Public/datarepo/netset/db/'
#population data
pop=pd.read_csv(pop_path+'pop.csv').set_index(['Country','Year']).unstack(level=1)
pop.head(3)
```
# Define database parsers and units
```
#initialize data and constants
data={}
countries={i for i in pop.index}
years={int(i[1]) for i in pop.columns}
dbs={'bp','eia'}
datatypes={'prod','cons','emi','res'}
allfuels=['oil','coal','gas','nuclear','biofuels','hydro','geo_other','solar','wind']
fossils=['oil','coal','gas']+['nrg','nrg_sum']
transp=1 #transparency
#colorlist=np.array([[166,86,40,transp*255],[153,153,153,transp*255],[152,78,163,transp*255],
# [228,26,28,transp*255],[247,129,191,transp*255],[55,126,184,transp*255],
# [82,56,65,transp*255],[255,255,51,transp*255],[77,175,74,transp*255]])/255.0
colorlist=np.array([[131,13,9,transp*255],[85,20,52,transp*255],[217,20,14,transp*255],
[213,9,98,transp*255],[64,185,85,transp*255],[202,200,46,transp*255],
[106,23,9,transp*255],[251,212,31,transp*255],[112,133,16,transp*255]])/255.0
gcolors={allfuels[i]:colorlist[i] for i in range(len(allfuels))}
def reset(what='all',datatype='all'):
global data
if what=='all':
#reset all values of database
fuels=allfuels+['nrg','nrg_sum']
data={i:{int(k[1]):{'energy':{j:{k:{l:np.NaN for l in dbs} for k in datatypes}\
for j in fuels},'population':long(pop.loc[i][k])*1000,\
'consumer_efficiency':0.5,\
'cumulative_emissions':0}\
for k in pop.columns}\
#we use population as the default database for country names
for i in pop.index}
else:
countries=data.keys()
for i in countries:
for j in years:
if datatype=='all':
data[i][j]['energy'][what]={k:{l:np.NaN for l in dbs} for k in datatypes}
else:
data[i][j]['energy'][what][datatype]={l:np.NaN for l in dbs}
reset()
kbpd_to_TWh=365.25*0.001628200 #unit conversion from thousand barrels of oil per day to TWh per year
Gboe_to_TWh=1628.2 #unit conversion from thousand million barrels of oil to TWh
EJ_to_TWh=277.77 #unit conversion from exa Joule to TWh
bcf_to_TWh=0.2931 #unit conversion from billion cubic feet of natural gas to TWh
tcf_to_TWh=bcf_to_TWh*1000.0 #unit conversion from trillion cubic feet of natural gas to TWh
qbtu_to_TWh=293.297222 #unit conversion from quadrillion British thermal units to TWh
mtoe_to_TWh=11.63 #unit conversion million metric tons of oil equivalent to TWh
kgge_to_gm3=1.49 #unit conversion from kilogram of natural gas to cubic meter, based on CH4
mtlnge_to_TWh=14.45 #unit conversion million metric tons of gas (LNG) equivalent to TWh
cm_to_cf=35.3 #unit conversion from million cubic meters to million cubic feet
tcm_to_TWh=tcf_to_TWh*cm_to_cf #unit conversion from trillion cubic meters of natural gas to TWh
kgge_to_TWh=kgge_to_gm3*tcf_to_TWh*cm_to_cf*1e-18 #unit conversion from kilogram of natural gas to TWh
#mtge_to_TWh=kgge_to_gm3*tcf_to_TWh*cm_to_cf*1e-9 #unit conversion from kilogram of natural gas to TWh
mtge_to_GJ=53.6
mtge_to_TWh=mtge_to_GJ*1e-9*EJ_to_TWh
t_to_st=1.10231 #unit conversion from metric ton to short ton
tcoe_to_toe=0.7 #unit conversion from metric tons of coal equivalent to metric tons of oil equivalent
mtcoe_to_TWh=tcoe_to_toe*mtoe_to_TWh #unit conversion million metric tons of coal equivalent to TWh
#mtcoe_to_TWh=8.141
mstcoe_to_TWh=mtcoe_to_TWh*t_to_st #unit conversion million metric short tons of coal equivalent to TWh
c_to_co2=44.0/12 #unit conversion from C to CO2 mass
carbon_budget=840*c_to_co2 #840 GtC as per http://www.ipcc.ch/report/ar5/wg1/
path = 2 #set database path
#online path = 0
#backup path = 1
#local path = 2
```
## Read, process & convert renewable resource maps
CSP
```
solar=pd.read_excel(pop_path+'maps/nrelsolar.xlsx',sheetname='DNI')
#http://en.openei.org/doe-opendata/dataset/19316a50-e55d-45e2-b692-efcebfe16c95/resource/3e72f32a-7de1-4e5d-a25a-76928769625f/download/solarresourceenergy.xlsx
solar_res={}
solar_cf={}
for i in solar.T.iteritems():
country=cnc(i[1][0])
if i[0]<10: #record capacity factors
solar_cf[i[1][9]]=i[1][10]
if country in data:
if country not in solar_res:solar_res[country]={}
clas=i[1][1]
cap=i[1][5]/1000.0 #workbook doest not import commas correctly, fix
solar_res[country][clas]=cap
plt.bar(solar_res['United States of America'].keys(),solar_res['United States of America'].values())
plt.ylabel('avail energy [Twh/year]')
plt.title('DNI-based solar thermal power\nwith 6 hrs of storage\n 31MW/km$^2$ plant power density')
plt.xlabel('solar resource class (~ 1/cost)')
```
PV
```
solar=pd.read_excel(pop_path+'maps/nrelsolar.xlsx',sheetname='Tilt=Lat')
#http://en.openei.org/doe-opendata/dataset/19316a50-e55d-45e2-b692-efcebfe16c95/resource/3e72f32a-7de1-4e5d-a25a-76928769625f/download/solarresourceenergy.xlsx
solar_res2={}
for i in solar.T.iteritems():
country=cnc(i[1][0])
if country in data:
if country not in solar_res2:solar_res2[country]={}
clas=i[1][1]
cap=i[1][6]
solar_res2[country][clas]=cap
plt.bar(solar_res2['United States of America'].keys(),solar_res2['United States of America'].values())
plt.ylabel('avail energy [Twh/year]')
plt.title('GHI-based solar PV power\nwith 10% efficiency\n 1.5% productive land')
plt.xlabel('solar resource class (~ 1/cost)')
plt.bar(solar_res2['United Arab Emirates'].keys(),solar_res2['United Arab Emirates'].values())
plt.ylabel('avail energy [Twh/year]')
plt.title('GHI-based solar PV power\nwith 10% efficiency\n 1.5% productive land')
plt.xlabel('solar resource class (~ 1/cost)')
```
convert to capacities
```
def interpolate(d,years,gfit=2,depth=1,polyorder=1,override=False,ends=False):
#d=helper
#years=[2015]
#gfit=1
#depth=extrapolatedict[fuel]
#polyorder=1
#override=True
#ends=True
#if True:
#depth * length of interpolation substrings will be taken to the left and right
#for example for {1971:5,1972:6,1973:7,1974:5} interpolating it over 1969-1990
#for the section 1960-1970 (2 elements) the values from 1972,1973,1974 (3 elements) will be taken with depth 1.5
#for the section 1974-1990 (15 elements) all values (4 elements) will be taken to extrapolate
#override to extend interpolation to edges, i.e. extrapolate
if (gfit>2):
print 'interpolate takes only 1 (polynomial) or 2 (exponential) as 3rd argument [default=2]'
return
mydict={}
if d!={}:
missing_points=[[]]
onbeginning=False
onend=False
for year in years:
if year not in d.keys():
missing_points[-1].append(year)
else:
missing_points.append([])
for m in missing_points:
if m:
fit=gfit
#if only one point, set min extrapolation depth to 2
if (len(m)==1): depth=max(depth,2)
#check if it is ends of the interval,
if ((m[-1]<np.sort(d.keys())[0])|(m[0]>np.sort(d.keys())[-1])):
#if not set to override then extrapolate mean only
if not override:
fit=0
if fit==0: #take average
y = {k: d[k] for k in set(d.keys()).intersection(range(int(max(min(years),min(m)-int(3))),\
int(min(max(years),max(m)+int(3))+1)))}
#returned empty, on beginning
if y=={}:
if m[-1]<np.sort(d.keys())[0]:y={np.sort(d.keys())[0]:d[np.sort(d.keys())[0]]}
elif m[0]>np.sort(d.keys())[-1]:y={np.sort(d.keys())[-1]:d[np.sort(d.keys())[-1]]}
for i in range(len(m)):
mydict[m[i]]=np.mean(y.values())
elif fit==1:
#intersector
y = {k: d[k] for k in set(d.keys()).intersection(range(int(max(min(years),\
min(m)-int(depth*len(m)))),int(min(max(years),max(m)+int(depth*len(m)))+1)))}
#returned empty
if y=={}:
if m[-1]<np.sort(d.keys())[0]:y={np.sort(d.keys())[0]:d[np.sort(d.keys())[0]]}
elif m[0]>np.sort(d.keys())[-1]:y={np.sort(d.keys())[-1]:d[np.sort(d.keys())[-1]]}
w = np.polyfit(y.keys(),y.values(),polyorder) # obtaining regression parameters
if (polyorder==1):
intersector=w[0]*np.array(m)+w[1]
else:
intersector=w[0]*np.array(m)*np.array(m)+w[1]*np.array(m)+w[2]
for i in range(len(m)):
mydict[m[i]]=max(0,intersector[i])
else:
#exponential intersector
y = {k: d[k] for k in set(d.keys()).intersection(range(int(max(min(years),\
min(m)-int(depth*len(m)))),int(min(max(years),max(m)+int(depth*len(m)))+1)))}
#returned empty
if y=={}:
if m[-1]<np.sort(d.keys())[0]:y={np.sort(d.keys())[0]:d[np.sort(d.keys())[0]]}
elif m[0]>np.sort(d.keys())[-1]:y={np.sort(d.keys())[-1]:d[np.sort(d.keys())[-1]]}
w = np.polyfit(y.keys(),np.log(y.values()),1) # obtaining log regression parameters (exp fitting)
intersector=np.exp(w[1])*np.exp(w[0]*np.array(m))
for i in range(len(m)):
mydict[m[i]]=max(0,intersector[i])
#record ends adjustment beginning and end
if ends:
if (m[-1]<np.sort(d.keys())[0]):
onbeginning=True
beginning=m[-1]
if (m[0]>np.sort(d.keys())[-1]):
onend=True
end=m[0]
#finish ends adjustment
if ends:
if onbeginning:
#calculate adjustment scaler
if (mydict[beginning]==0): scaler=0
elif (beginning+1 in d): scaler=d[beginning+1]*1.0/mydict[beginning]
else: scaler=d[np.sort(d.keys())[0]]*1.0/mydict[beginning]
#readjust data
for year in mydict:
if (year<=beginning):
mydict[year]*=scaler
if onend:
#calculate adjustment scaler
if (mydict[end]==0): scaler=0
elif (end-1 in d): scaler=d[end-1]*1.0/mydict[end]
else: scaler=d[np.sort(d.keys())[-1]]*1.0/mydict[end]
#readjust data
for year in mydict:
if (year>=end):
mydict[year]*=scaler
#return interpolated points
return mydict
#extrapolate solar classes
helper=interpolate(solar_cf,range(0,15),1,1,2,True,True)
solar_cf.update(helper)
solar_cap={}
for i in solar_res:
if i not in solar_cap:solar_cap[i]={}
for j in solar_res[i]:
solar_cap[i][j]=solar_res[i][j]*solar_cf[j]/8760
solar_cap2={}
for i in solar_res2:
if i not in solar_cap2:solar_cap2[i]={}
for j in solar_res2[i]:
solar_cap2[i][j]=solar_res2[i][j]*solar_cf[j]/8760
```
wind
```
wind=pd.read_excel(pop_path+'maps/nrelwind.xlsx',sheetname='Onshore Power',skiprows=2)
#http://en.openei.org/doe-opendata/dataset/c186913f-6684-4455-a2f2-f26e152a9b35/resource/4dc4a6fd-3a63-47df-bcbe-e9c83b83b38e/download/nrelcfddawindsc20130603.xlsx
wind_cap={}
for i in wind.T.iteritems():
country=i[1][0]
if country in data:
for k in range(1,10):
if country not in wind_cap:wind_cap[country]={}
clas=k
cap=i[1][k] + i[1][k+10]# + i[1][k+20] #last one too far 0-50,50-100,100-5000
wind_cap[country][clas]=cap/1000.0 #workbook reported values in GW, we use TW
plt.bar(wind_cap['Argentina'].keys(),wind_cap['Denmark'].values())
plt.ylabel('avail cap [TW]')
plt.title('onshore wind resources\nwith 90m hub height\n 5MW/km$^2$ power density')
plt.xlabel('wind resource class (~ 1/cost)')
wind=pd.read_excel(pop_path+'maps/nrelwind.xlsx',sheetname='Onshore Energy',skiprows=2)
#http://en.openei.org/doe-opendata/dataset/c186913f-6684-4455-a2f2-f26e152a9b35/resource/4dc4a6fd-3a63-47df-bcbe-e9c83b83b38e/download/nrelcfddawindsc20130603.xlsx
wind_res={}
for i in wind.T.iteritems():
country=i[1][0]
if country in data:
for k in range(1,10):
if country not in wind_res:wind_res[country]={}
clas=k
cap=i[1][k] + i[1][k+10]# + i[1][k+20] #last one too far 0-50,50-100,100-5000
wind_res[country][clas]=cap*1000.0 #workbook reported values in PWh, we use TWh
plt.bar(wind_res['Argentina'].keys(),wind_res['Denmark'].values())
plt.ylabel('avail energy [TWh/year]')
plt.title('onshore wind resources\nwith 90m hub height\n 5MW/km$^2$ power density')
plt.xlabel('wind resource class (~ 1/cost)')
wind=pd.read_excel(pop_path+'maps/nrelwind.xlsx',sheetname='Offshore Energy',skiprows=3)
#http://en.openei.org/doe-opendata/dataset/c186913f-6684-4455-a2f2-f26e152a9b35/resource/4dc4a6fd-3a63-47df-bcbe-e9c83b83b38e/download/nrelcfddawindsc20130603.xlsx
wind_res2={}
country=''
for i in wind.T.iteritems():
if str(i[1][0])!='nan':
country=str(i[1][0])
water=str(i[1][1])
if water in {'shallow','transitional'}: #skip deep for now
if country in data:
for k in range(2,11):
if country not in wind_res2:wind_res2[country]={}
clas=k-1
cap=i[1][k] + i[1][k+10]# + i[1][k+20] #last one too far 0-50,50-100,100-5000
if clas not in wind_res2[country]:wind_res2[country][clas]=0
wind_res2[country][clas]+=cap*1000.0
plt.bar(wind_res2['Denmark'].keys(),wind_res2['Denmark'].values())
plt.ylabel('avail energy [TWh/year]')
plt.title('offshore wind resources\nwith 90m hub height\n 5MW/km$^2$ power density')
plt.xlabel('wind resource class (~ 1/cost)')
wind=pd.read_excel(pop_path+'maps/nrelwind.xlsx',sheetname='Offshore Power',skiprows=3)
#http://en.openei.org/doe-opendata/dataset/c186913f-6684-4455-a2f2-f26e152a9b35/resource/4dc4a6fd-3a63-47df-bcbe-e9c83b83b38e/download/nrelcfddawindsc20130603.xlsx
wind_cap2={}
country=''
for i in wind.T.iteritems():
if str(i[1][0])!='nan':
country=str(i[1][0])
water=str(i[1][1])
if water in {'shallow','transitional'}: #skip deep for now
if country in data:
for k in range(2,11):
if country not in wind_cap2:wind_cap2[country]={}
clas=k-1
cap=i[1][k] + i[1][k+10]# + i[1][k+20] #last one too far 0-50,50-100,100-5000
if clas not in wind_cap2[country]:wind_cap2[country][clas]=0
wind_cap2[country][clas]+=cap/1000.0
plt.bar(wind_cap2['Denmark'].keys(),wind_cap2['Denmark'].values())
plt.ylabel('avail cap [TW]')
plt.title('offshore wind resources\nwith 90m hub height\n 5MW/km$^2$ power density')
plt.xlabel('wind resource class (~ 1/cost)')
```
## Save resource data
```
res={}
for i in solar_res:
if i not in res: res[i]={"wind":{"cap":{},"res":{}},"pv":{"cap":{},"res":{}},"csp":{"cap":{},"res":{}}}
res[i]["csp"]["res"]=solar_res[i]
res[i]["csp"]["cap"]=solar_cap[i]
res[i]["pv"]["res"]=solar_res2[i]
res[i]["pv"]["cap"]=solar_cap2[i]
for i in wind_res: #onshore
if i not in res: res[i]={"wind":{"cap":{},"res":{}},"pv":{"cap":{},"res":{}},"csp":{"cap":{},"res":{}}}
res[i]["wind"]["res"]=wind_res[i]
res[i]["wind"]["cap"]=wind_cap[i]
for i in wind_res2: #offshore
if i not in res: res[i]={"wind":{"cap":{},"res":{}},"pv":{"cap":{},"res":{}},"csp":{"cap":{},"res":{}}}
for j in wind_res2[i]:
if j not in res[i]["wind"]["res"]:res[i]["wind"]["res"][j]=0
res[i]["wind"]["res"][j]+=wind_res2[i][j]
for j in wind_cap2[i]:
if j not in res[i]["wind"]["cap"]:res[i]["wind"]["cap"][j]=0
res[i]["wind"]["cap"][j]+=wind_cap2[i][j]
plt.bar(res['Norway']['wind']['cap'].keys(),res['Norway']['wind']['cap'].values())
plt.ylabel('avail cap [TW]')
plt.title('total wind resources\nwith 90m hub height\n 5MW/km$^2$ power density')
plt.xlabel('wind resource class (~ 1/cost)')
#save data
file('data/resources/res.json','w').write(json.dumps(res))
```
## Potentials
Estimate remaining potentials
```
#biomass
#http://www.sciencedirect.com/science/article/pii/S0961953414005340
#Table 5, they all come on one tier
#nuclear
#the chance for developing nuclear is essentially zero, e.g. infinite cost
#hydro
#http://www.intpow.com/index.php?id=487&download=1
```
create EROEI - resource class functions
```
#load savedata if exists
predata=json.loads(file('data/resources/res.json','r').read())
res={}
for c in predata:
res[c]={}
for f in predata[c]:
res[c][f]={}
for r in predata[c][f]:
res[c][f][r]={}
for year in predata[c][f][r]:
res[c][f][r][int(year)]=predata[c][f][r][year]
predata={}
#check
res['Afghanistan']['csp']['res']
```
resource normalization
```
#pv classes = 4-14
#wind classes = 1-9
#csp classes = 1-10
def histo(x,y,b): #one elment histogram rebin interpolator (for wind 9->10 classes)
if b>len(x):
if b==10 and len(x)==9:
yy=np.histogram(x,weights=y,bins=b)[0]
#return yy #turn off
toadd=((yy[3]+yy[5])/2.0)*0.7 #multipl. to adjust for recurrency .88 approx.
shares=np.array(yy)/np.sum(yy) #redistribute based on resource size
for s in range(10):
if s not in [4]:
shares[s]*=(abs(1.0/(4-s))) #scale based on dist from number
shares=shares/np.sum(shares)
for s in range(10):
if s not in [4]:
yy[s]-=toadd*shares[s]
yy[4]+=toadd*shares[s]
return yy
else: print "error, this is a custom function..."
else:
return np.histogram(x,weights=y,bins=b)[0]
lim={"pv":[4,14],"wind":[1,9],"csp":[1,10]}
newres={}
for c in res:
newres[c]={}
for r in res[c]:
newres[c][r]={}
for f in res[c][r]:
newres[c][r][f]={}
helper=dict(res[c][r][f])
x=[]
y=[]
for k in range(0,15):
if k<lim[r][0] or k>lim[r][1]: #pop items outside class limits - these are small errors
if k in helper:helper.pop(k);
elif k not in helper:helper[k]=0 #else insert it if is not there already
for i in range(len(sorted(helper.keys()))):
x.append(float(sorted(helper.keys())[i]))
y.append(float(helper[sorted(helper.keys())[i]]))
#plt.figure()
#plt.bar(x,y,alpha=0.6)
yy=histo(x,y,10)
xx=range(1,11)
#plt.bar(xx,yy,alpha=0.6,color='red')
for i in xx:
newres[c][r][f][i]=yy[i-1]
res=newres
#save data
file('data/resources/res_exp.json','w').write(json.dumps(res))
#load res savedata if exists
predata=json.loads(file('data/resources/res_exp.json','r').read())
res={}
for c in predata:
res[c]={}
for f in predata[c]:
res[c][f]={}
for r in predata[c][f]:
res[c][f][r]={}
for year in predata[c][f][r]:
res[c][f][r][int(year)]=predata[c][f][r][year]
predata={}
#load energy savedata (from output of calculator.ipynb)
predata=json.loads(file('data/historical/savedata.json','r').read())
data={}
for c in predata:
data[c]={}
for year in predata[c]:
data[c][int(year)]=predata[c][year]
predata={}
```
exponential distribution
```
#class-based distribution function
pwr=3
s=sum([i**pwr for i in range(10)])*1.0
classconverter={i+1:i**pwr/s for i in range(10)}
plt.plot(classconverter.keys(),np.array(classconverter.values())*100)#,color='#dd1c77',lw=2,alpha=0.9, align='center')
plt.xlabel('Resource Quality [decile]')
plt.ylabel('Percentage allocated [%]')
plt.xlim(0.5,10.5)
plt.show()
l=4
k=np.array([i**3 for i in range(l)])
plt.plot(np.arange(l)/3.0,k*100.0/sum(k),'magenta',lw=2,label=u'$f(x)=x^3/\sum_{classes}{x^3}$')
print k*100.0/sum(k),k*6.0/sum(k)
plt.ylabel("allocated share [%]")
plt.xlabel("resource quality distribution [1 - best, 0 - worst]\nexample with 3 classes")
plt.title("Function for distributing in-country\n generation into resource classes")
plt.legend(loc=2)
plt.savefig('res3exp.png',bbox_inches = 'tight', pad_inches = 0.1, dpi=150)
plt.show()
l=7
k=np.array([i**3 for i in range(l)])
plt.plot(np.arange(l)/6.0,k*100.0/sum(k),'magenta',lw=2,label=u'$f(x)=x^3/\sum_{classes}{x^3}$')
print k*100.0/sum(k),k*5.0/sum(k)
plt.ylabel("allocated share [%]")
plt.xlabel("resource quality distribution [1 - best, 0 - worst]\nexample with 6 classes")
plt.title("Function for distributing in-country\n generation into resource classes")
plt.legend(loc=2)
plt.savefig('res6exp.png',bbox_inches = 'tight', pad_inches = 0.1, dpi=150)
plt.show()
def classconvert(a,b,pwr=3):#a classed resource in 10 classes, return exponential distribution, with fill control
classvector=[]
for i in range(1,11):
if a[i]>0:
if not np.isnan(a[i]):
classvector.append(i)
s=sum([i**pwr for i in classvector])*1.0
z={}
for i in range(1,11):
if i in classvector:
z[i]=i**pwr/s
else: z[i]=0
w={}
tofill=b
for i in range(10,0,-1):
if a[i]>0:
if tofill*z[i]>a[i]: #used to be tofill>a[i]:
w[i]=a[i]/b
tofill-=a[i]
z.pop(i);
else: break
q=sum(w.values())
for i in range(1,11):
if i not in w:
w[i]=max(0,z[i]/sum(z.values())*(1.0-q))
return w
#pv classes = 4-14
#wind classes = 1-9
#csp classes = 1-10
groei={}#resources
groei3={}#built
sd={"pv":[],"wind":[],"csp":[]}
nru={"pv":[],"wind":[],"csp":[]}
sd2={"pv":[],"wind":[],"csp":[]}
rr={"pv":"solar","wind":"wind","csp":"csp"}
for c in res:
croei={}
#if True:
try:
for r in res[c]:
if r not in groei: groei[r]={}
if r not in groei3: groei3[r]={}
#calculate exponential dsitribution in-country
dynamicres={} #record initial sum of res - necessary for exhaustion mechanism
for cl in range(10,0,-1): #start from top, step down
if not np.isnan(res[c][r]['res'][cl]):
if cl not in groei[r]: groei[r][cl]=0
if cl not in croei: croei[cl]=0
v=res[c][r]['res'][cl]
groei[r][cl]=np.nansum([groei[r][cl],v])
croei[cl]=np.nansum([croei[cl],v])
#exponential distribution across resource classes for existing production
#s=res[c][r]['res'][cl]*1.0/sum(res[c][r]['res'].values()) #old, even distribution
if c in data:
for year in data[c]:
if rr[r] in data[c][year]['energy']:
if year<2015:
if cl not in groei3[r]: groei3[r][cl]=0
s=classconvert(res[c][r]['res'],data[c][year]['energy'][rr[r]]['cons']['navg3'])[cl]
v=s*data[c][year]['energy'][rr[r]]['cons']['navg3']
if not np.isnan(v):
if v>0.01:
sd2[r].append({"country":c,"class":cl,"value":v,"year":year})
if year==2014:
groei3[r][cl]=np.nansum([groei3[r][cl],v])
sd[r].append({"country":c,"class":cl,"value":res[c][r]['res'][cl],"year":2014})
except: print 'ERROR',c
aroei={}
titles=['Photovoltaic','Solar Thermal','Wind']
t2=['\n10% overall efficiency\ncovering 1.5% of productive land in each country',
'\ncalculated with 6 hours of storage\n31MW/km$^2$ power density',
'\nonshore + offshore, calculated for 90m hub height\n 5MW/km$^2$ power density']
t2=['','','']
t3=['Global Horizontal Irradiance [kWh / sqm / day]','Direct Normal Irradiance [kWh / sqm / day]','NREL Wind Power Class']
t4=[[0.5,10.5],[0.5,10.5],[0.5,10.5]]
t5=[[0,70000],[0,250000],[0,100000]]
t6=[[0,70],[0,10],[0,300]]
t7=[[0,50000],[0,50000],[0,2000]]
locs=[2,2,1]
cvtr={"pv":0.5,"wind":1,"csp":0.64}
cvtr2={"pv":0.25,"wind":0,"csp":1.87}
for j,r in enumerate(groei):
#if True:
x=[]
y=[]
for i in range(len(sorted(groei[r].keys()))):
if float(groei[r][sorted(groei[r].keys())[i]])>0.1:
x.append(float(sorted(groei[r].keys())[i]))
y.append(float(groei[r][sorted(groei[r].keys())[i]]))
aroei[r]=np.average(x,weights=y)
fig=plt.figure(figsize=(7,5))
plt.bar(x,y,color='#dd1c77',lw=2,alpha=0.7, align='center')
plt.title('Global '+titles[j]+' Energy Resource'+t2[j]+' Utilization',fontsize=14,y=1.05)
plt.ylabel('Available energy [TWh / year]')
plt.xlabel('Normalized Resource Quality Class [1-worst to 10-best]')
plt.ylim(t5[j])
plt.tick_params(axis='y', colors='#dd1c77')
z=np.average(x,weights=y)
print r,'weighted average class of resources',z
z2=z*1.0
plt.axvline(z,color='#dd1c77',lw=2,ls='--')
#new_tick_locations = [2,8]+[np.round(z,1)]
new_tick_locations = [np.round(z,1)]
x=[]
y=[]
for i in range(len(sorted(groei3[r].keys()))):
x.append(float(sorted(groei[r].keys())[i]))
y.append(float(groei3[r][sorted(groei3[r].keys())[i]]))
ax=plt.twinx()
plt.plot([-10],[-10],color='#dd1c77',lw=2,label='Total Energy Resource')
plt.plot([-10],[-10],color='royalBlue',lw=2,label='Generation in 2014')
ax.bar(x,y,color='royalBlue',lw=2,alpha=0.7, align='center')
plt.plot([-10],[-10],color='grey',lw=2,ls='',label=' ')
plt.plot([-10],[-10],color='grey',lw=2,ls='--',label='weighted average')
ax.tick_params(axis='y', colors='royalBlue')
z=np.average(x,weights=y)
print r,'weighted average class of generation',z
print r,'normalized resource utilization',z/z2
plt.axvline(z,color='royalBlue',lw=2,ls='--')
plt.ylim(t6[j])
plt.xlim(t4[j])
plt.legend(loc=locs[j],framealpha=0)
ax2 = fig.add_axes((0.16,-0.01,0.705,0.0))
ax2.yaxis.set_visible(False)
ax2.set_xlabel('Normalized Resource Utilization')
ax2.set_xlim(1,10)
new_tick_locations += [np.round(z,1)]
new_ticks = np.round(np.array(new_tick_locations)/new_tick_locations[0],2)
nru[r]=new_tick_locations
ax2.set_xticks(new_tick_locations)
ax2.set_xticklabels(new_ticks)
#wind class http://rredc.nrel.gov/wind/pubs/atlas/appendix_A.html
#NREL Renewable Resource Data Center
plt.savefig(str(r)+'3.png',bbox_inches = 'tight', pad_inches = 0.1, dpi=150)
plt.savefig('plots/'+str(r)+'.png',bbox_inches = 'tight', pad_inches = 0.1, dpi=150)
plt.show()
```
save for viz
```
#save
file(pop_path+'../universal/res.json','w').write(json.dumps({"res":sd,"inst":sd2}))
import zipfile
try:
import zlib
compression = zipfile.ZIP_DEFLATED
except:
compression = zipfile.ZIP_STORED
zf = zipfile.ZipFile(pop_path+'../universal/res.zip', mode='w')
zf.write(pop_path+'../universal/res.json','res.json',compress_type=compression)
zf.close()
```
save for later
```
file('data/resources/res_redist.json','w').write(json.dumps(newres))
```
shares
```
#normalized resource utilization
nru
#shares
w=[0.827]
p=[0.157]
c=[0.016]
#Shares
W=[0.827]
P=[0.157]
C=[0.016]
#thetas - normalized resource utilization
wt=[1.09] #wind
wp=[0.68] #pv
wc=[1.18] #csp
wt=[nru['wind'][1]/nru['wind'][0]] #wind
wp=[nru['pv'][1]/nru['pv'][0]] #pv
wc=[nru['csp'][1]/nru['csp'][0]] #csp
#js
jt=[1] #wind
jp=[1.77] #pv
jc=[1.59] #csp
javg=(nru['wind'][0]+nru['csp'][0]+nru['pv'][0])/3.0
jt=[nru['wind'][0]/javg] #wind
jp=[nru['pv'][0]/javg] #pv
jc=[nru['csp'][0]/javg] #csp
tgt=1.57 #target share wind
tgp=1 #target share pv
tgc=1.48 #target share csp
s=0.35
z=0.5
f=2
x=[2014]
for i in range(2015,2101):
k=i-2014
x.append(i)
wt.append(wt[0]+(tgt-wt[0])*(i-2014)/(2101-2015))
wp.append(wp[0]+(tgp-wp[0])*(i-2014)/(2101-2015))
wc.append(wc[0]+(tgc-wc[0])*(i-2014)/(2101-2015))
w.append(w[k-1]+((1.0/3)*(1/wt[k-1])**z*jt[0]**s)**f)
p.append(p[k-1]+((1.0/3)*(1/wp[k-1])**z*jp[0]**s)**f)
c.append(c[k-1]+((1.0/3)*(1/wc[k-1])**z*jc[0]**s)**f)
jt.append(jt[0])
jp.append(jp[0])
jc.append(jc[0])
W.append(W[k-1]+w[k]/(w[k]+p[k]+c[k]))
P.append(P[k-1]+p[k]/(w[k]+p[k]+c[k]))
C.append(C[k-1]+c[k]/(w[k]+p[k]+c[k]))
fig,ax=plt.subplots(1,5,figsize=(17,4))
ax[0].plot(x,wt)
ax[0].plot(x,wp)
ax[0].plot(x,wc)
ax[0].set_title('Utilization')
ax[1].plot(x,w,label='wind')
ax[1].plot(x,p,label='pv')
ax[1].plot(x,c,label='csp')
ax[1].legend(loc=2)
ax[1].set_title('New additions')
ax[3].plot(x,W)
ax[3].plot(x,P)
ax[3].plot(x,C)
ax[3].set_title('Cumulative added')
ax[2].plot(x,[w[k]/(w[k]+p[k]+c[k]) for k in range(len(w))])
ax[2].plot(x,[p[k]/(w[k]+p[k]+c[k]) for k in range(len(w))])
ax[2].plot(x,[c[k]/(w[k]+p[k]+c[k]) for k in range(len(w))])
ax[2].set_title('Share in new additions')
ax[4].plot(x,[W[k]/W[k] for k in range(len(w))])
ax[4].plot(x,[P[k]/W[k] for k in range(len(w))])
ax[4].plot(x,[C[k]/W[k] for k in range(len(w))])
ax[4].set_title('Share in cu. relative to wind')
plt.show()
for year in [86]:
print x[year],'omega wind',W[year]*1.0/(W[year]+P[year]+C[year]),
print 'omega pv',P[year]*1.0/(W[year]+P[year]+C[year]),
print 'omega csp',C[year]*1.0/(W[year]+P[year]+C[year])
for year in [6,26,46,66,86]:
print x[year],'omega/wind wind',W[year]*1.0/(W[year]),
print 'omega/wind pv',P[year]*1.0/(W[year]),
print 'omega/wind csp',C[year]*1.0/(W[year])
eroei={"pv":12,"wind":22,"csp":12}
xk={"pv":tgp,"wind":tgt,"csp":tgc}
rtau={}
for r in eroei:
rtau[r]=eroei[r]*1.0/(nru[r][1]/nru[r][0])
print r,': theta',nru[r][1]/nru[r][0],'xk',xk[r],'rtau',rtau[r],'avg class',nru[r][0]
```
<p><br><p>We would like to express our gratitude to all of the developers of the libraries used and especially to the affiliates of <i>EIA, BP and World Bank</i> - and for this particular workbook <i>NREL</i>, for their great database and openly accesible data. The data manipulation algorithms are open sourced and freely reproducible under an <a href="http://opensource.org/licenses/MIT" target="_blank">MIT license</a>.</p>
<br>
<p><a href="http://www.csaladen.es" target="_blank">Dénes Csala</a> | 2016</p>
| github_jupyter |
```
import datarobot as dr
from datarobot import Project, Deployment
import pandas as pd
from pandas.io.json import json_normalize
import numpy as np
import plotly.express as px
import seaborn as sns
import matplotlib.pyplot as plt
import datetime as dt
from datetime import datetime
import dateutil.parser
import os
import re
from importlib import reload
import random
import math
import numpy.ma as ma
import timesynth as ts
import matplotlib.pyplot as plt
from matplotlib.pyplot import figure
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
# Set Pandas configuration to show all columns
pd.set_option('display.max_columns', None)
pd.set_option('display.max_colwidth', -1)
dr.Client(config_path='../drconfig.yaml');
df = pd.read_csv(""" Enter Code """)
df.head(5)
df.plot(x='Date', y='Sales', figsize=(20, 8));
```
## Months
```
MONTHS = ''
def months(df):
global MONTHS
MIN_DATE = df['Date'].min()
MAX_DATE = df['Date'].max()
MONTHS = str(int((MAX_DATE - MIN_DATE).days / 30))
print('Min Date: ', MIN_DATE)
print('Max Date: ', MAX_DATE)
print('Months: ', MONTHS)
months(df)
```
## Create TS Settings
```
DATE = """ Enter Code """
TARGET = """ Enter Code """
PROJECT_NAME = 'Lab_2'
VERSION = '1'
MODE = 'Q'
FDWS = """ Enter Code """
FDS = """ Enter Code """
BASE = 'L2_Base_V:'
PREFIX = BASE + VERSION + '_Mnths:' + MONTHS + '_Mode:' + MODE
DATASET_FILENAME = 'Months_' + MONTHS
MAX_WAIT = 14400
READ_TIMEOUT = 14400
HOLDOUT_START_DATE = None
VALIDATION_DURATION = None
HOLDOUT_DURATION = None
NUMBER_BACKTESTS = None
GAP_DURATION = None
FEATURE_SETTINGS = []
CAL_ID = None
print(FEATURE_SETTINGS)
print(CAL_ID)
```
## Create Project
### Function to create projects
```
def create_dr_project(df, project_name, fw_start=None, fw_end=None, fdw_start=None, fdw_end=None):
###############################
# Create Datetime Specification
###############################
# SERIES_COL = [SERIES]
time_partition = dr.DatetimePartitioningSpecification(
datetime_partition_column = DATE,
forecast_window_start = fw_start,
forecast_window_end = fw_end,
feature_derivation_window_start = fdw_start,
feature_derivation_window_end = fdw_end,
holdout_start_date = HOLDOUT_START_DATE ,
validation_duration = VALIDATION_DURATION,
holdout_duration = HOLDOUT_DURATION,
gap_duration = GAP_DURATION,
number_of_backtests = NUMBER_BACKTESTS,
feature_settings = FEATURE_SETTINGS,
use_time_series = True,
calendar_id = CAL_ID
)
################
# Create Project
################
project = dr.Project.create(
project_name = project_name,
sourcedata = df,
max_wait = MAX_WAIT,
read_timeout = READ_TIMEOUT,
dataset_filename = DATASET_FILENAME
)
print("Post-Project MB: ", (df.memory_usage(index=True).sum()/1024/1024).round(2))
print("Post-Project Records: {:,}".format(len(df)))
print(f'Project {project_name} Created...')
#################
# Start Autopilot
#################
project.set_target(
target = TARGET,
metric = None,
mode = dr.AUTOPILOT_MODE.QUICK , # dr.AUTOPILOT_MODE.FULL_AUTO,
#advanced_options = opts,
worker_count = -1,
partitioning_method = time_partition,
max_wait = MAX_WAIT
)
return project
```
## Model Factory
```
projects = [] # Keep List of all projects
```
### Function to iterate through various FDWs & FDs
```
def model_factory(df, FDWS, FDS):
PREFIX = BASE + str(VERSION) + '_Mnths:' + MONTHS + '_Mode:' + MODE
for fdw in FDWS:
for fd in FDS:
fd_start = fd[0]
fd_end = fd[1]
fdw_start = fdw[0]
fdw_end = fdw[1]
# Name project
project_name = f"{PREFIX}_FDW:{fdw_start}-{fdw_end}_FD:{fd_start}-{fd_end}"
print(project_name)
data = df.copy()
# Create project
project = create_dr_project(data, project_name,
fw_start=fd_start, fw_end=fd_end,
fdw_start=fdw_start, fdw_end=fdw_end)
projects.append(project)
```
## Default Model
```
model_factory(df, FDWS, FDS)
```
## Forecast Distance Breakout
```
# Update Version
VERSION = 2
FDS = """ Enter Code """
model_factory(df, FDWS, FDS)
```
# Pull Results
## Get Project Names in a List
```
projects = dr.Project.list(search_params={'project_name': BASE})
projects
```
## Get Project Names and PIDs in a List
```
lst = []
for p in projects:
r = ((p, p.id))
lst.append(r)
lst
```
## Unlock Holdouts
```
for i in lst:
project = Project.get(i[1])
project.unlock_holdout()
```
## Compute Backtests for Blenders
```
for i in lst:
project = Project.get(i[1])
lb = project.get_datetime_models()
for model in lb:
if 'Blender' in model.model_type:
try:
print(project.project_name, model)
dr.DatetimeModel.score_backtests(model)
print(f'Computing backtests for model {model.id} in Project {project.project_name}')
except dr.errors.ClientError:
pass
print(f'All available backtests have been submitted for scoring for project {project.project_name}')
print(' ')
else:
None
```
## Compute All Backtests for Top Models in Backtest 1 and Holdout groups
```
OPTIMIZE_GROUP = ['validation', 'holdout']
PROJECT_METRIC = project.metric
METRICS = list(set([PROJECT_METRIC, 'MASE', 'RMSE']))
for p in lst :
for met in METRICS:
for o in OPTIMIZE_GROUP:
project = Project.get(p[1])
lb = project.get_datetime_models()
best_models = sorted(
[model for model in lb if model.metrics[met][o]],
key=lambda m: m.metrics[met][o],
)[0:3]
for mod in best_models:
if mod.metrics[met]["backtesting"] == None:
try:
print(project.project_name, mod)
dr.DatetimeModel.score_backtests(mod)
print(f'Computing backtests for model {mod.model_type} in Project {project.project_name}')
except dr.errors.ClientError:
pass
print(f'All available backtests have been submitted for scoring for project {project.project_name}')
print(' ')
else:
print(project.project_name)
print(f'{mod.model_type} All Backtests Already Computed')
print(' ')
```
## Get Project and Model Scores
```
OPTIMIZATION_PERIOD = 'backtesting' # BackTest 1: validation All Backtest: backtesting Holdout: holdout
```
### Get scores in a DataFrame
```
models = []
scores = pd.DataFrame()
for p in lst:
project = Project.get(p[1])
lb = project.get_datetime_models()
best_model = sorted(
[model for model in lb if model.metrics[project.metric][OPTIMIZATION_PERIOD]],
key=lambda m: m.metrics[project.metric][OPTIMIZATION_PERIOD],
)[0]
backtest_scores = pd.DataFrame(
[
{
'Project_Name': project.project_name,
'Project_ID': project.id,
'Model_ID': best_model.id,
'Model_Type': best_model.model_type,
'Featurelist': best_model.featurelist_name,
'Optimization_Metric': project.metric,
'Scores': best_model.metrics,
}
]
)
scores = scores.append(backtest_scores, sort=False).reset_index(drop=True)
print(f'Scores for all {len(projects)} projects have been computed')
print('')
scores = scores.join(json_normalize(scores["Scores"].tolist())).drop(labels=['Scores'], axis=1)
# Drop Empty Columns
scores = scores[scores.columns.drop(list(scores.filter(regex='crossValidation$')))]
# Rename Columns
scores.columns = scores.columns.str.replace(".backtesting", "_All_BT")
scores.columns = scores.columns.str.replace(".holdout", "_Holdout")
scores.columns = scores.columns.str.replace(".validation", "_BT_1")
scores.columns = scores.columns.str.replace(' ', '_')
scores = scores[scores.columns.drop(list(scores.filter(regex='_All_BTScores$')))]
scores.head(2)
```
### Put column names into variables for easy reference
```
METRICS = scores.filter(regex='MASE|RMSE').columns.to_list()
PROJECT = ['Project_Name', 'Project_ID', 'Model_ID', 'Model_Type', 'Featurelist']
COLS = PROJECT + METRICS
scores[COLS]
```
### Sort dataframe by score column
```
scores['FDW_Start'] = scores['Project_Name'].str.extract(r'FDW:(-\d{1,2})')
scores['FDW_End'] = scores['Project_Name'].str.extract(r'FDW:-\d{1,2}-(\d{1,2})_')
scores['FD_Start'] = scores['Project_Name'].str.extract(r'FD:(\d{1,2})')
scores['FD_End'] = scores['Project_Name'].str.extract(r'FD:\d{1,2}-(\d{1,2})')
scores['Months'] = scores['Project_Name'].str.extract(r'_Mnths:(\d{1,2})_')
scores.rename(columns={'All_Backtests_Poisson Deviance':'All_Backtests_Poisson_Deviance',
'Backtest_1_Poisson Deviance':'Backtest_1_Poisson_Deviance',
'Holdout_Poisson Deviance':'Holdout_Poisson_Deviance',
'Holdout_Tweedie Deviance':'Holdout_Tweedie_Deviance',
'All_Backtests_Tweedie Deviance':'All_Backtests_Tweedie_Deviance',
'Backtest_1_Tweedie Deviance':'Backtest_1_Tweedie_Deviance',
'Holdout_Tweedie Deviance':'Holdout_Tweedie_Deviance'}, inplace=True)
META = ['FDW_Start', 'FDW_End', 'FD_Start', 'FD_End', 'Months']
MORE = PROJECT + META + METRICS
scores[MORE].sort_values(by=[""" Enter Code """], ascending=True)
```
## Get Best Model
```
hrmse = scores.loc[scores[""" Enter Code """].notnull()]
# Take the Single Best model
# hrmse_best = pd.DataFrame(hrmse.loc[hrmse.MASE_All_BT.idxmin()]).transpose()
# Take the Best model by Project Name
hrmse_best = hrmse.loc[hrmse.groupby('Project_Name').MASE_All_BT.idxmin()]
best_models = pd.DataFrame(hrmse_best)
best_models
```
## Generate Predictions
### Select the project and model from the dataframe
```
RECORD = 0
# Verify correct project and model
PID = best_models['Project_ID'].values[RECORD]
MID = best_models['Model_ID'].values[RECORD]
project = dr.Project.get(PID)
model = dr.Model.get(PID, MID)
print(project, model)
PID = best_models['Project_ID'].values[RECORD]
MID = best_models['Model_ID'].values[RECORD]
project = dr.Project.get(PID)
model = dr.Model.get(PID, MID)
dataset = project.upload_dataset(""" Enter Code """)
pred_job = model.request_predictions(dataset_id = dataset.id)
preds = pred_job.get_result_when_complete()
preds.head(5)
```
## Generate Predictions with 90% Prediction Interval
```
PID = best_models['Project_ID'].values[RECORD]
MID = best_models['Model_ID'].values[RECORD]
project = dr.Project.get(PID)
model = dr.Model.get(PID, MID)
dataset = project.upload_dataset(""" Enter Code """)
pred_job = model.request_predictions(dataset_id = dataset.id,
""" Enter Code """)
preds = pred_job.get_result_when_complete()
preds.head(5)
```
| github_jupyter |
Importing the dataset and libraries.
```
import pandas as pd
solicitations=pd.read_csv("Data\sedec_solicitacoes.csv",delimiter =';')
```
Renaming the column processo_numero to ID to match the other datasets, and while doing that dropping processo_origem.
```
solicitations = solicitations.rename(columns={'processo_numero':'ID'})
solicitations.drop(columns='processo_origem', axis=1, inplace=True)
```
Taking a look at the dataset.
```
solicitations.info()
solicitations.head()
```
In the solicitacao_descricao column there is information about if a sliding took place. Additionally the processo_situacao column says if the process is considered complete or not. So we are going to isolate the keywords deslizamento and completo and transform into 0/1 values.
```
solicitations['processo_situacao'] = solicitations['processo_situacao'].astype('str')
solicitations['solicitacao_descricao'] = solicitations['solicitacao_descricao'].astype('str')
solicitations['solicitacao_descricao'].str.lower()
solicitations['solicitacao_descricao']=[1 if 'deslizamento' in review else 0 for review in solicitations['solicitacao_descricao']]
solicitations['processo_situacao']=[1 if sit == 'completo' else 0 for sit in solicitations['processo_situacao']]
solicitations
```
Dropping the columns that have no relevant information.
```
solicitations.drop(columns='solicitacao_hora', axis=1, inplace=True)
solicitations.drop(columns='solicitacao_plantao', axis=1, inplace=True)
solicitations.drop(columns='solicitacao_localidade', axis=1, inplace=True)
solicitations.drop(columns='rpa_nome', axis=1, inplace=True)
solicitations.drop(columns='rpa_codigo', axis=1, inplace=True)
solicitations.drop(columns='solicitacao_regional', axis=1, inplace=True)
solicitations.drop(columns='solicitacao_microrregiao', axis=1, inplace=True)
solicitations.drop(columns='solicitacao_endereco', axis=1, inplace=True)
solicitations.drop(columns='solicitacao_roteiro', axis=1, inplace=True)
solicitations.drop(columns='processo_solicitacao', axis=1, inplace=True)
solicitations.head()
solicitations.info()
```
Renaming the columns for better understading.
```
solicitations = solicitations.rename(columns={'ano':'year','mes':'month','solicitacao_data':'day',
'solicitacao_descricao':'description','solicitacao_bairro':'neighbourhood',
'processo_situacao':'situation'})
solicitations.head()
```
Dropping the duplicate values.
```
solicitations = solicitations.sort_values('description', ascending=False).drop_duplicates('ID').sort_index()
solicitations.info()
```
Isolating the month and day values in the respective columns.
```
aux=[]
for i in range (0,len(solicitations['month'])):
aux = solicitations.loc[solicitations.index[i],'month']
solicitations.loc[solicitations.index[i],'month'] = aux[:2]
solicitations.loc[solicitations.index[i],'month'] = int(solicitations.loc[solicitations.index[i],'month'])
solicitations['month'][:10]
aux=[]
for i in range (0,len(solicitations['day'])):
aux = solicitations.loc[solicitations.index[i],'day']
solicitations.loc[solicitations.index[i],'day'] = aux[-2:]
solicitations['day'][:10]
solicitations.head()
solicitations['day'] = solicitations['day'].astype(int)
solicitations.head()
```
There is still one column with no relevant information. We are keeping the neighbourhood column because there are no null values in this one, so we can cross-reference that with the latitude/longitude values and complete the data about the locale.
```
solicitations = solicitations.drop(columns='processo_tipo',axis=1)
solicitations.head()
```
Taking a look at the unique values and information of the dataset.
```
solicitations['description'].value_counts()
solicitations['situation'].value_counts()
solicitations['neighbourhood'].value_counts()
locations = solicitations.groupby(['neighbourhood']).size().reset_index().rename(columns={0:'count'})
locations[70:90]
number_locations = locations['count'].value_counts()
number_locations
len(number_locations)
```
Converting the date information to a pandas time series.
```
import datetime
solicitations['Date'] = pd.to_datetime(solicitations[['year', 'month', 'day']])
solicitations.head()
```
Droping the unnecessary date columns.
```
solicitations = solicitations.drop(columns=['year','month','day'],axis=1)
solicitations.head()
```
The analysis ends in december 2019 so we need to remove the first row.
```
solicitations = solicitations.drop([0])
solicitations.head()
```
Transforming the neighbourhood data into numerical values, we will just transform it into a sequential integer list. Additionally there are some rows which do not have information about the location ('Não informado') so we need to remove those rows.
```
locations= locations[locations['neighbourhood']!='Não informado']
locations['list']=range(1,120)
locations = locations.drop(columns='count',axis=1)
locations.head()
solicitations['location']= 0
solicitations = solicitations[solicitations['neighbourhood']!='Não informado']
solicitations.head()
solicitations.info()
for n in range (0,len(solicitations)):
b= locations.loc[locations['neighbourhood']==solicitations.neighbourhood.iloc[n]]
solicitations.location.iloc[n]= b.list.iloc[0]
solicitations.head()
```
Dropping the neighbourhood column since there is no need for it anymore.
```
solicitations = solicitations.drop(columns='neighbourhood',axis=1)
solicitations.head()
solicitations.to_csv(path_or_buf='solicitations_prepared.csv')
```
| github_jupyter |
# functorial-learning
```
from discopy import Tensor
import jax.numpy as np; import numpy; np.random = numpy.random; np.random.seed(42)
Tensor.np = np # this line is necessary when using jax
```
## 0. Generate a language
We first build the pregroup diagrams for a simple subject-verb-object language.
```
from discopy import Ty, Word, Cup, Id
nouns = ["Alice", "Bob", "Charlie", "Diane", "Eve", "Fred", "George", "Isabel"]
verbs = ["loves", "hates", "kills"]
s, n = Ty('s'), Ty('n')
tv = n.r @ s @ n.l
vocab = [Word(noun, n) for noun in nouns] + [Word(verb, tv) for verb in verbs]
grammar = Cup(n, n.r) @ Id(s) @ Cup(n.l, n)
sentences = [
"{} {} {}".format(subj, verb, obj)
for subj in nouns for verb in verbs for obj in nouns]
print("{}*{}*{} = {} sentences:\n{}\n".format(
len(nouns), len(verbs), len(nouns), len(sentences), "\n".join(sentences[:3] + ["..."] + sentences[-3:])))
diagrams = {
sentence: Word(subj, n) @ Word(verb, tv) @ Word(obj, n) >> grammar
for sentence in sentences for subj, verb, obj in [sentence.split(' ')]}
diagrams[sentences[1]].draw()
print("Pregroup diagram for '{}'".format(sentences[1]))
```
## 1. Build a random dataset
We then generate a toy dataset with scalars attached to each diagram.
In order to do so, we apply a random functor from Pregroup to Tensor to each diagram in the corpus.
A `TensorFunctor` is defined by two mappings `ob` from types to dimension (i.e. hyper-parameters) and `ar` from words to matrices (i.e. parameters).
```
from discopy import TensorFunctor
# N.B. defining ob[tv] is not necessary, indeed F(tv) == F(n) * F(s) * F(n)
ob = {n: 2, s: 1, tv: 4}
ar0 = {word: np.random.randn(ob[word.cod]) for word in vocab}
F0 = TensorFunctor(ob, ar0)
def sigmoid(x):
return 1 / (1 + np.exp(-x))
dataset = {sentence: sigmoid(F0(diagram).array) for sentence, diagram in diagrams.items()}
y_true = [float(y) for y in dataset.values()]
```
Let's have a look at what this random dataset looks like:
```
from matplotlib import pyplot as plt
plt.hist(y_true, bins=100)
plt.show()
from random import shuffle
delta = 0.1
true_sentences = [sentence for sentence in dataset if dataset[sentence] > 0.5 + delta]
shuffle(true_sentences)
print("{} true sentences:\n{}\n".format(
len(true_sentences), "\n".join(true_sentences[:3] + ['...'] + true_sentences[-3:])))
print("Does Alice love Bob?\n{}".format(
"Yes" if dataset["Alice loves Bob"] > 0.5 + delta else "No"))
```
## 2. Learn the functor from data
We split the dataset into training and testing.
We forget `F0` and initialise some new random functor.
We define the mean squared loss on testing and training set and compile them just-in-time.
We use automatic differentiation and vanilla gradient descent to learn the parameters for the functor.
```
from sklearn.model_selection import train_test_split
training, testing = train_test_split(sentences, test_size=0.25, random_state=42)
def loss(ar, sample=training):
F = TensorFunctor(ob, ar)
y_pred = [sigmoid(F(diagrams[sentence]).array) for sentence in sample]
y_true = [dataset[sentence] for sentence in sample]
return np.mean((np.array(y_true) - np.array(y_pred)) ** 2)
ar = {word: np.random.randn(ob[word.cod]) for word in vocab}
print("Initial training loss: {}\nInitial testing loss: {}".format(loss(ar), loss(ar, testing)))
from time import time
from jax import jit, grad
training_loss = jit(loss)
testing_loss = jit(lambda ar: loss(ar, testing))
gradient = jit(grad(training_loss))
start = time()
print("{1:.0f} seconds to just-in-time compile the gradient:\n{0}".format("\n".join(
"{}: {}".format(word, dx) for word, dx in gradient(ar).items()), time() - start))
learning_rate = 1
epochs = 10
iterations = 100
training_losses = []
testing_losses = []
for i in range(epochs):
start = time()
for _ in range(iterations):
g = gradient(ar)
for word in vocab:
ar[word] = ar[word] - g[word] * learning_rate
training_losses.append(float(training_loss(ar)))
testing_losses.append(float(testing_loss(ar)))
print("Epoch {} ({:.3f} seconds)".format(i + 1, time() - start))
F = TensorFunctor(ob, ar)
y_pred = [float(sigmoid(F(diagrams[sentence]).array)) for sentence in sentences]
plt.plot(training_losses)
plt.xlabel("n_iterations")
plt.ylabel("training_loss")
plt.show()
plt.plot(testing_losses)
plt.xlabel("n_iterations")
plt.ylabel("testing_loss")
plt.show()
```
## 3. Evaluate the learnt functor on some task
Here we show the results for a binary classification task, i.e. yes-no question answering:
```
from sklearn.metrics import classification_report
print(classification_report(np.array(y_true) > .5 + delta, np.array(y_pred) > .5 + delta))
print("Does Alice love Bob?\n{}".format(
"Yes" if y_pred[sentences.index("Alice loves Bob")] > 0.5 + delta else "No"))
```
| github_jupyter |
# Convolutional Neural Networks
---
In this notebook, we train a **CNN** to classify images from the CIFAR-10 database.
The images in this database are small color images that fall into one of ten classes; some example images are pictured below.
<img src='https://github.com/udacity/deep-learning-v2-pytorch/blob/master/convolutional-neural-networks/cifar-cnn/notebook_ims/cifar_data.png?raw=1' width=70% height=70% />
### Test for [CUDA](http://pytorch.org/docs/stable/cuda.html)
Since these are larger (32x32x3) images, it may prove useful to speed up your training time by using a GPU. CUDA is a parallel computing platform and CUDA Tensors are the same as typical Tensors, only they utilize GPU's for computation.
```
import torch
import numpy as np
# check if CUDA is available
train_on_gpu = torch.cuda.is_available()
if not train_on_gpu:
print('CUDA is not available. Training on CPU ...')
else:
print('CUDA is available! Training on GPU ...')
```
---
## Load the [Data](http://pytorch.org/docs/stable/torchvision/datasets.html)
Downloading may take a minute. We load in the training and test data, split the training data into a training and validation set, then create DataLoaders for each of these sets of data.
```
from torchvision import datasets
import torchvision.transforms as transforms
from torch.utils.data.sampler import SubsetRandomSampler
# number of subprocesses to use for data loading
num_workers = 0
# how many samples per batch to load
batch_size = 20
# percentage of training set to use as validation
valid_size = 0.2
# convert data to a normalized torch.FloatTensor
transform = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))
])
# choose the training and test datasets
train_data = datasets.CIFAR10('data', train=True,
download=True, transform=transform)
test_data = datasets.CIFAR10('data', train=False,
download=True, transform=transform)
# obtain training indices that will be used for validation
num_train = len(train_data)
indices = list(range(num_train))
np.random.shuffle(indices)
split = int(np.floor(valid_size * num_train))
train_idx, valid_idx = indices[split:], indices[:split]
# define samplers for obtaining training and validation batches
train_sampler = SubsetRandomSampler(train_idx)
valid_sampler = SubsetRandomSampler(valid_idx)
# prepare data loaders (combine dataset and sampler)
train_loader = torch.utils.data.DataLoader(train_data, batch_size=batch_size,
sampler=train_sampler, num_workers=num_workers)
valid_loader = torch.utils.data.DataLoader(train_data, batch_size=batch_size,
sampler=valid_sampler, num_workers=num_workers)
test_loader = torch.utils.data.DataLoader(test_data, batch_size=batch_size,
num_workers=num_workers)
# specify the image classes
classes = ['airplane', 'automobile', 'bird', 'cat', 'deer',
'dog', 'frog', 'horse', 'ship', 'truck']
```
### Visualize a Batch of Training Data
```
import matplotlib.pyplot as plt
%matplotlib inline
# helper function to un-normalize and display an image
def imshow(img):
img = img / 2 + 0.5 # unnormalize
plt.imshow(np.transpose(img, (1, 2, 0))) # convert from Tensor image
# obtain one batch of training images
dataiter = iter(train_loader)
images, labels = dataiter.next()
images = images.numpy() # convert images to numpy for display
# plot the images in the batch, along with the corresponding labels
fig = plt.figure(figsize=(25, 4))
# display 20 images
for idx in np.arange(20):
ax = fig.add_subplot(2, 20/2, idx+1, xticks=[], yticks=[])
imshow(images[idx])
ax.set_title(classes[labels[idx]])
```
### View an Image in More Detail
Here, we look at the normalized red, green, and blue (RGB) color channels as three separate, grayscale intensity images.
```
rgb_img = np.squeeze(images[3])
channels = ['red channel', 'green channel', 'blue channel']
fig = plt.figure(figsize = (20, 20))
for idx in np.arange(rgb_img.shape[0]):
ax = fig.add_subplot(1, 3, idx + 1)
img = rgb_img[idx]
ax.imshow(img, cmap='gray')
ax.set_title(channels[idx])
width, height = img.shape
thresh = img.max()/2.5
for x in range(width):
for y in range(height):
val = round(img[x][y],2) if img[x][y] !=0 else 0
ax.annotate(str(val), xy=(y,x),
horizontalalignment='center',
verticalalignment='center', size=8,
color='white' if img[x][y]<thresh else 'black')
```
---
## Define the Network [Architecture](http://pytorch.org/docs/stable/nn.html)
This time, you'll define a CNN architecture. Instead of an MLP, which used linear, fully-connected layers, you'll use the following:
* [Convolutional layers](https://pytorch.org/docs/stable/nn.html#conv2d), which can be thought of as stack of filtered images.
* [Maxpooling layers](https://pytorch.org/docs/stable/nn.html#maxpool2d), which reduce the x-y size of an input, keeping only the most _active_ pixels from the previous layer.
* The usual Linear + Dropout layers to avoid overfitting and produce a 10-dim output.
A network with 2 convolutional layers is shown in the image below and in the code, and you've been given starter code with one convolutional and one maxpooling layer.
<img src='https://github.com/udacity/deep-learning-v2-pytorch/blob/master/convolutional-neural-networks/cifar-cnn/notebook_ims/2_layer_conv.png?raw=1' height=50% width=50% />
#### TODO: Define a model with multiple convolutional layers, and define the feedforward network behavior.
The more convolutional layers you include, the more complex patterns in color and shape a model can detect. It's suggested that your final model include 2 or 3 convolutional layers as well as linear layers + dropout in between to avoid overfitting.
It's good practice to look at existing research and implementations of related models as a starting point for defining your own models. You may find it useful to look at [this PyTorch classification example](https://github.com/pytorch/tutorials/blob/master/beginner_source/blitz/cifar10_tutorial.py) or [this, more complex Keras example](https://github.com/keras-team/keras/blob/master/examples/cifar10_cnn.py) to help decide on a final structure.
#### Output volume for a convolutional layer
To compute the output size of a given convolutional layer we can perform the following calculation (taken from [Stanford's cs231n course](http://cs231n.github.io/convolutional-networks/#layers)):
> We can compute the spatial size of the output volume as a function of the input volume size (W), the kernel/filter size (F), the stride with which they are applied (S), and the amount of zero padding used (P) on the border. The correct formula for calculating how many neurons define the output_W is given by `(W−F+2P)/S+1`.
For example for a 7x7 input and a 3x3 filter with stride 1 and pad 0 we would get a 5x5 output. With stride 2 we would get a 3x3 output.
```
import torch.nn as nn
import torch.nn.functional as F
# define the CNN architecture
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(3, 6, 5)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(6, 16, 5)
self.fc1 = nn.Linear(16 * 5 * 5, 120)
self.fc2 = nn.Linear(120, 84)
self.fc3 = nn.Linear(84, 10)
def forward(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
x = x.view(-1, 16 * 5 * 5)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.fc3(x)
return x
# create a complete CNN
model = Net()
print(model)
# move tensors to GPU if CUDA is available
if train_on_gpu:
model.cuda()
```
### Specify [Loss Function](http://pytorch.org/docs/stable/nn.html#loss-functions) and [Optimizer](http://pytorch.org/docs/stable/optim.html)
Decide on a loss and optimization function that is best suited for this classification task. The linked code examples from above, may be a good starting point; [this PyTorch classification example](https://github.com/pytorch/tutorials/blob/master/beginner_source/blitz/cifar10_tutorial.py) or [this, more complex Keras example](https://github.com/keras-team/keras/blob/master/examples/cifar10_cnn.py). Pay close attention to the value for **learning rate** as this value determines how your model converges to a small error.
#### TODO: Define the loss and optimizer and see how these choices change the loss over time.
```
import torch.optim as optim
# specify loss function
criterion = nn.CrossEntropyLoss()
# specify optimizer
optimizer = optim.Adam(model.parameters(), lr=0.001)
```
---
## Train the Network
Remember to look at how the training and validation loss decreases over time; if the validation loss ever increases it indicates possible overfitting.
```
# number of epochs to train the model
n_epochs = 8 # you may increase this number to train a final model
valid_loss_min = np.Inf # track change in validation loss
for epoch in range(1, n_epochs+1):
# keep track of training and validation loss
train_loss = 0.0
valid_loss = 0.0
###################
# train the model #
###################
model.train()
for data, target in train_loader:
# move tensors to GPU if CUDA is available
if train_on_gpu:
data, target = data.cuda(), target.cuda()
# clear the gradients of all optimized variables
optimizer.zero_grad()
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the batch loss
loss = criterion(output, target)
# backward pass: compute gradient of the loss with respect to model parameters
loss.backward()
# perform a single optimization step (parameter update)
optimizer.step()
# update training loss
train_loss += loss.item()*data.size(0)
######################
# validate the model #
######################
model.eval()
for data, target in valid_loader:
# move tensors to GPU if CUDA is available
if train_on_gpu:
data, target = data.cuda(), target.cuda()
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the batch loss
loss = criterion(output, target)
# update average validation loss
valid_loss += loss.item()*data.size(0)
# calculate average losses
train_loss = train_loss/len(train_loader.dataset)
valid_loss = valid_loss/len(valid_loader.dataset)
# print training/validation statistics
print('Epoch: {} \tTraining Loss: {:.6f} \tValidation Loss: {:.6f}'.format(
epoch, train_loss, valid_loss))
# save model if validation loss has decreased
if valid_loss <= valid_loss_min:
print('Validation loss decreased ({:.6f} --> {:.6f}). Saving model ...'.format(
valid_loss_min,
valid_loss))
torch.save(model.state_dict(), 'model_cifar.pt')
valid_loss_min = valid_loss
```
### Load the Model with the Lowest Validation Loss
```
model.load_state_dict(torch.load('model_cifar.pt'))
```
---
## Test the Trained Network
Test your trained model on previously unseen data! A "good" result will be a CNN that gets around 70% (or more, try your best!) accuracy on these test images.
```
# track test loss
test_loss = 0.0
class_correct = list(0. for i in range(10))
class_total = list(0. for i in range(10))
model.eval()
# iterate over test data
for data, target in test_loader:
# move tensors to GPU if CUDA is available
if train_on_gpu:
data, target = data.cuda(), target.cuda()
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the batch loss
loss = criterion(output, target)
# update test loss
test_loss += loss.item()*data.size(0)
# convert output probabilities to predicted class
_, pred = torch.max(output, 1)
# compare predictions to true label
correct_tensor = pred.eq(target.data.view_as(pred))
correct = np.squeeze(correct_tensor.numpy()) if not train_on_gpu else np.squeeze(correct_tensor.cpu().numpy())
# calculate test accuracy for each object class
for i in range(batch_size):
label = target.data[i]
class_correct[label] += correct[i].item()
class_total[label] += 1
# average test loss
test_loss = test_loss/len(test_loader.dataset)
print('Test Loss: {:.6f}\n'.format(test_loss))
for i in range(10):
if class_total[i] > 0:
print('Test Accuracy of %5s: %2d%% (%2d/%2d)' % (
classes[i], 100 * class_correct[i] / class_total[i],
np.sum(class_correct[i]), np.sum(class_total[i])))
else:
print('Test Accuracy of %5s: N/A (no training examples)' % (classes[i]))
print('\nTest Accuracy (Overall): %2d%% (%2d/%2d)' % (
100. * np.sum(class_correct) / np.sum(class_total),
np.sum(class_correct), np.sum(class_total)))
```
### Question: What are your model's weaknesses and how might they be improved?
The model is too simple to capture the features from the images. It needs more complexity to capture them. Dropout and data augmentation will also be helpful.
### Visualize Sample Test Results
```
# obtain one batch of test images
dataiter = iter(test_loader)
images, labels = dataiter.next()
images.numpy()
# move model inputs to cuda, if GPU available
if train_on_gpu:
images = images.cuda()
# get sample outputs
output = model(images)
# convert output probabilities to predicted class
_, preds_tensor = torch.max(output, 1)
preds = np.squeeze(preds_tensor.cpu().numpy())
# plot the images in the batch, along with predicted and true labels
fig = plt.figure(figsize=(25, 4))
for idx in np.arange(20):
ax = fig.add_subplot(2, 20/2, idx+1, xticks=[], yticks=[])
images = images.cpu()
imshow(images[idx])
ax.set_title("{} ({})".format(classes[preds[idx]], classes[labels[idx]]),
color=("green" if preds[idx]==labels[idx].item() else "red"))
```
| github_jupyter |
Create a file called ref_trajectory.py. Copy the functions from week3 that enabled you to generate a trajectory given a sequence of straight/ turn commands
```
import cv2
import numpy as np
from PIL import Image
import matplotlib.pyplot as plt
#from ref_trajectory import generate_trajectory as traj
v = 1
dt = 0.1
num_st_pts = int(v/dt)
num_pts = 50
def cubic_spiral(theta_i, theta_f, n=10):
x = np.linspace(0, 1, num=n)
#-2*x**3 + 3*x**2
return (theta_f-theta_i)*(-2*x**3 + 3*x**2) + theta_i
def straight(dist, curr_pose, n=num_st_pts):
# the straight-line may be along x or y axis
#curr_theta will determine the orientation
x0, y0, t0 = curr_pose
xf, yf = x0 + dist*np.cos(t0), y0 + dist*np.sin(t0)
x = (xf - x0) * np.linspace(0, 1, n) + x0
y = (yf - y0) * np.linspace(0, 1, n) + y0
return x, y, t0*np.ones_like(x)
def turn(change, curr_pose, n=num_pts):
# adjust scaling constant for desired turn radius
x0, y0, t0 = curr_pose
theta = cubic_spiral(t0, t0 + np.deg2rad(change), n)
x= x0 + np.cumsum(v*np.cos(theta)*dt)
y= y0 + np.cumsum(v*np.sin(theta)*dt)
return x, y, theta
def generate_trajectory(route, init_pose = (0, 0,np.pi/2)):
curr_pose = init_pose
func = {'straight': straight, 'turn': turn}
x, y, t = np.array([]), np.array([]),np.array([])
for manoeuvre, command in route:
px, py, pt = func[manoeuvre](command, curr_pose)
curr_pose = px[-1],py[-1],pt[-1]
x = np.concatenate([x, px])
y = np.concatenate([y, py])
t = np.concatenate([t, pt])
return np.vstack([x, y, t])
traj = generate_trajectory
```
We will look at 2 ways to make a grid
(a) Load an existing circuit (Use MS paint or anything else)
(b) Make a custom grid
Remember grid resolution is the key parameter to go from grid co-ordinates to world co-ordinates. What is the value of 1 pixel?
```
grid_res = 0.05
image = Image.open('./circuit.png').convert('LA') # open the test circuit
grid_data = np.asarray(image)[:,:,0]/255
#find the shape
print(grid_data.shape)
grid_shape = np.array(grid_data.shape)
print(grid_shape)
#find the real-world extents of the grid: opposite vertices of the rectangle
x1, y1 = 0, 0
#use shape and grid_res to calculate the top-right vertex
x2, y2 = grid_shape*grid_res
print(x2, y2)
#display the image using imshow()
#pay attention to the keywords origin, cmap and extent
#Default origin is top-left, but we should use 'lower'
#how should grid_data be transformed
plt.figure()
#complete
plt.imshow(grid_data, cmap = plt.cm.gray, extent=[x1,x2,y1,y2])
start_pose = np.array([12, 4, np.pi/2])
plt.plot(start_pose[0], start_pose[1],'r+')
plt.show()
#custom grid
#choose resolution and dimensions
grid_res = 0.05
grid_span = 50 # square circuit dimensions in m
#calculate grid_shape from grid
# dimensions have to be integers
grid_shape = (np.array([grid_span]*2)/grid_res).astype('int')
#Initialize
grid_data = np.zeros(grid_shape)
#Create rectangular obstacles in world co-ordinates
#xmin, xmax, ymin, ymax
obstacles = np.array([[25, 26, 10, 40],
[2, 8, 16, 20]])
for obs in obstacles:
# calculate obstacles extent in pixel coords
xmin, xmax, ymin, ymax = (obs/grid_res).astype('int')
# mark them as occupied
grid_data[xmin:xmax ,ymin:ymax ] = 1.0
#calculate the extents
x1, y1 = 0, 0
x2, y2 = grid_span, grid_span
#imshow() from prior cell
plt.figure()
plt.imshow(1-grid_data.T, origin='lower', cmap=plt.cm.gray, extent=[x1,x2,y1,y2] )
```
Plot the reference trajectory on the grid environment
```
#pick a suitable starting pose
start_pose = np.array([2, 6, np.pi/2])
#we will use same route as before
route = [("straight", 5),("turn", -90),("straight", 6),("turn", 90)]
#calculate ref_path from traj()
# we want it to be numpy array of dimensions (N, 3)
ref_path = traj(route, start_pose).T
#plot the grid and
plt.imshow(1-grid_data.T, origin='lower', cmap=plt.cm.gray, extent=[x1,x2,y1,y2])
plt.plot(ref_path[:,0], ref_path[:,1])
plt.plot(start_pose[0], start_pose[1],'r+')
```
Let us now use circle approximation to perform collision checking. We will use w=0.8, l=1.2m as mule dimensions. Calculate the 4 vertices of the mule
Circle parameters are offset and radii
You would have to plot the mule rectangle and 3 circles
```
w=0.8
l=1.2
mule_extents = np.array([[-w/2,-l/2],
[w/2, -l/2],
[w/2, l/2],
[-w/2, l/2],
[-w/2,-l/2]])
r = 0.5
l = 0.4
circles = [(0, 0, r), (0, l, r), (0, -l, r)]
plt.figure()
#plot rectangle or just the 4 vertices
for vertex in mule_extents:
p, q = vertex
plt.plot(p, q, 'b.')
for v1, v2 in zip(mule_extents[:-1],mule_extents[1:]):
p1, q1 = v1
p2, q2 = v2
plt.plot((p1, p2), (q1,q2), 'k-')
ax = plt.gca()
for x,y,rad in circles:
ax.add_patch(plt.Circle((x,y), rad, fill=False))
```
Write a function to find if there is a collision given the grid and the local path
1. Rotate the 3 circle vertices along the local path
2. Check if $(x-x_c)^2 + (y-y_c)^2 < r$ and grid[x, y] = 1.0
3. Return True or False
You can assume circles variable (approximation of the vehicle dimensions) is assumed as a global variable
What do you think is a simple unit test?
```
grid_res = 0.05
def circle_collision_check(grid, local_traj):
xmax, ymax = grid.shape
all_x = np.arange(xmax)
all_y = np.arange(ymax)
X, Y = np.meshgrid(all_x, all_y)
for xl, yl, tl in local_traj:
rot = np.array([[np.sin(tl), -np.cos(tl)],[np.cos(tl), np.sin(tl)]])
for xc, yc, rc in circles:
xc_rot, yc_rot = rot @ np.array([xc, yc]) + np.array([xl, yl])
print((np.array([xc, yc]) + np.array([xl, yl])).shape)
xc_pix, yc_pix = int(xc_rot/grid_res), int(yc_rot/ grid_res)
rc_pix = (rc/ grid_res)
inside_circle = ((X-xc_pix)**2 +(Y-yc_pix)**2 - rc_pix**2 < 0)
occupied_pt = grid[X, Y] == 1
if np.sum(np.multiply( inside_circle, occupied_pt)):
return True
return False
sush = np.array([[1, 2, 3]])
print(np.shape(sush))
local_traj = np.array([[0, 0.25, np.pi/2], [0, 1.0, np.pi/2], [0, 1.5, np.pi/2]])
grid = np.zeros([50, 50])
grid[0:2,30:32] = 1.0
print(circle_collision_check(grid, local_traj))
plt.imshow(1 - grid.T, origin='lower',cmap = plt.cm.gray, extent = [0, 2.5, 0, 2.5])
```
Let us move onto the more complicated swath collision checking
Write a function to trace the footprint of the vehicle along the local path.
Input: grid_size, local path
Output: footprint
Start with an empty grid of similar size
1. Rotate the 4 vertices of the mule rectangle along the local path
2. Collect all the vertices and calculate the convex hull. Remember to typecast them as float32 as cv2..convexHull() expects that
3. Use fillConvexPoly() to fill up the hull
```
def swath_footprint(local_path,grid_size):
#initialize image
img = np.zeros(grid_size)
vertices = []
for xl,yl,tl in local_path:
rot = np.array([[np.sin(tl), -np.cos(tl)],[np.cos(tl), np.sin(tl)]])
for vertex in mule_extents:
rot_vertex = rot @ vertex + np.array([xl, yl])
vertices.append(rot_vertex)
vertices = np.array(vertices,dtype='float32')
hull_vertices = cv2.convexHull(vertices)
#change hull vertices to grid coords and typecast as int32
hull_vertices = (hull_vertices/grid_res).astype('int32')
cv2.fillConvexPoly(img,hull_vertices,1)
return img.T == 1
s, e = 10, 20
footprint = swath_footprint(ref_path[s:e], grid_data.shape)
plt.imshow(1 - footprint.T, origin='lower', cmap=plt.cm.gray, extent=[x1,x2,y1,y2])
plt.plot(ref_path[:,0], ref_path[:,1],'b-')
plt.plot(ref_path[s:e,0], ref_path[s:e,1],'r-')
plt.xlim(0, 10)
plt.ylim(0, 20)
```
Once you know the footprint, it should be easy to see if the footprint coincides with any occupied cells in the original grid
Try out some examples to unit test your code for both circle checking and swath checking
In practice, you can write this as a CollisionChecking class where the mule dimensions are read from a config file and initialized. Global variables are best avoided in production code
```
```
| github_jupyter |
# Welcome to fastai
```
from fastai import *
from fastai.vision import *
from fastai.gen_doc.nbdoc import *
from fastai.core import *
from fastai.basic_train import *
```
The fastai library simplifies training fast and accurate neural nets using modern best practices. It's based on research in to deep learning best practices undertaken at [fast.ai](http://www.fast.ai), including "out of the box" support for [`vision`](/vision.html#vision), [`text`](/text.html#text), [`tabular`](/tabular.html#tabular), and [`collab`](/collab.html#collab) (collaborative filtering) models. If you're looking for the source code, head over to the [fastai repo](https://github.com/fastai/fastai) on GitHub. For brief examples, see the [examples](https://github.com/fastai/fastai/tree/master/examples) folder; detailed examples are provided in the full documentation (see the sidebar). For example, here's how to train an MNIST model using [resnet18](https://arxiv.org/abs/1512.03385) (from the [vision example](https://github.com/fastai/fastai/blob/master/examples/vision.ipynb)):
```
path = untar_data(URLs.MNIST_SAMPLE)
data = ImageDataBunch.from_folder(path)
learn = create_cnn(data, models.resnet18, metrics=accuracy)
learn.fit(1)
jekyll_note("""This documentation is all built from notebooks;
that means that you can try any of the code you see in any notebook yourself!
You'll find the notebooks in the <a href="https://github.com/fastai/fastai/tree/master/docs_src">docs_src</a> folder of the
<a href="https://github.com/fastai/fastai">fastai</a> repo. For instance,
<a href="https://nbviewer.jupyter.org/github/fastai/fastai/blob/master/docs_src/index.ipynb">here</a>
is the notebook source of what you're reading now.""")
```
## Installation and updating
To install or update fastai, we recommend `conda`:
```
conda install -c pytorch -c fastai fastai pytorch-nightly cuda92
```
For troubleshooting, and alternative installations (including pip and CPU-only options) see the [fastai readme](https://github.com/fastai/fastai/blob/master/README.md).
## Reading the docs
To get started quickly, click *Applications* on the sidebar, and then choose the application you're interested in. That will take you to a walk-through of training a model of that type. You can then either explore the various links from there, or dive more deeply into the various fastai modules.
We've provided below a quick summary of the key modules in this library. For details on each one, use the sidebar to find the module you're interested in. Each module includes an overview and example of how to use it, along with documentation for every class, function, and method. API documentation looks, for example, like this:
### An example function
```
show_doc(rotate)
```
---
Types for each parameter, and the return type, are displayed following standard Python [type hint syntax](https://www.python.org/dev/peps/pep-0484/). Sometimes for compound types we use [type variables](/fastai_typing.html). Types that are defined by fastai or Pytorch link directly to more information about that type; try clicking *Image* in the function above for an example. The docstring for the symbol is shown immediately after the signature, along with a link to the source code for the symbol in GitHub. After the basic signature and docstring you'll find examples and additional details (not shown in this example). As you'll see at the top of the page, all symbols documented like this also appear in the table of contents.
For inherited classes and some types of decorated function, the base class or decorator type will also be shown at the end of the signature, delimited by `::`. For `vision.transforms`, the random number generator used for data augmentation is shown instead of the type, for randomly generated parameters.
## Module structure
### Imports
fastai is designed to support both interactive computing as well as traditional software development. For interactive computing, where convenience and speed of experimentation is a priority, data scientists often prefer to grab all the symbols they need, with `import *`. Therefore, fastai is designed to support this approach, without compromising on maintainability and understanding.
In order to do so, the module dependencies are carefully managed (see next section), with each exporting a carefully chosen set of symbols when using `import *`. In general, for interactive computing, you'll want to import from both `fastai`, and from one of the *applications*, such as:
```
from fastai import *
from fastai.vision import *
```
That will give you all the standard external modules you'll need, in their customary namespaces (e.g. `pandas as pd`, `numpy as np`, `matplotlib.pyplot as plt`), plus the core fastai libraries. In addition, the main classes and functions for your application ([`fastai.vision`](/vision.html#vision), in this case), e.g. creating a [`DataBunch`](/basic_data.html#DataBunch) from an image folder and training a convolutional neural network (with [`create_cnn`](/vision.learner.html#create_cnn)), are also imported. Alternatively, use `from fastai import vision` to import the vision
If you wish to see where a symbol is imported from, either just type the symbol name (in a REPL such as Jupyter Notebook or IPython), or (in most editors) wave your mouse over the symbol to see the definition. For instance:
```
Learner
```
### Dependencies
At the base of everything are the two modules [`core`](/core.html#core) and [`torch_core`](/torch_core.html#torch_core) (we're not including the `fastai.` prefix when naming modules in these docs). They define the basic functions we use in the library; [`core`](/core.html#core) only relies on general modules, whereas [`torch_core`](/torch_core.html#torch_core) requires pytorch. Most type-hinting shortcuts are defined there too (at least the one that don't depend on fastai classes defined later). Nearly all modules below import [`torch_core`](/torch_core.html#torch_core).
Then, there are three modules directly on top of [`torch_core`](/torch_core.html#torch_core):
- [`data`](/vision.data.html#vision.data), which contains the class that will take a [`Dataset`](https://pytorch.org/docs/stable/data.html#torch.utils.data.Dataset) or pytorch [`DataLoader`](https://pytorch.org/docs/stable/data.html#torch.utils.data.DataLoader) to wrap it in a [`DeviceDataLoader`](/basic_data.html#DeviceDataLoader) (a class that sits on top of a [`DataLoader`](https://pytorch.org/docs/stable/data.html#torch.utils.data.DataLoader) and is in charge of putting the data on the right device as well as applying transforms such as normalization) and regroup then in a [`DataBunch`](/basic_data.html#DataBunch).
- [`layers`](/layers.html#layers), which contains basic functions to define custom layers or groups of layers
- [`metrics`](/metrics.html#metrics), which contains all the metrics
This takes care of the basics, then we regroup a model with some data in a [`Learner`](/basic_train.html#Learner) object to take care of training. More specifically:
- [`callback`](/callback.html#callback) (depends on [`data`](/vision.data.html#vision.data)) defines the basis of callbacks and the [`CallbackHandler`](/callback.html#CallbackHandler). Those are functions that will be called every step of the way of the training loop and can allow us to customize what is happening there;
- [`basic_train`](/basic_train.html#basic_train) (depends on [`callback`](/callback.html#callback)) defines [`Learner`](/basic_train.html#Learner) and [`Recorder`](/basic_train.html#Recorder) (which is a callback that records training stats) and has the training loop;
- [`callbacks`](/callbacks.html#callbacks) (depends on [`basic_train`](/basic_train.html#basic_train)) is a submodule defining various callbacks, such as for mixed precision training or 1cycle annealing;
- `learn` (depends on [`callbacks`](/callbacks.html#callbacks)) defines helper functions to invoke the callbacks more easily.
From [`data`](/vision.data.html#vision.data) we can split on one of the four main *applications*, which each has their own module: [`vision`](/vision.html#vision), [`text`](/text.html#text) [`collab`](/collab.html#collab), or [`tabular`](/tabular.html#tabular). Each of those submodules is built in the same way with:
- a submodule named <code>transform</code> that handles the transformations of our data (data augmentation for computer vision, numericalizing and tokenizing for text and preprocessing for tabular)
- a submodule named <code>data</code> that contains the class that will create datasets specific to this application and the helper functions to create [`DataBunch`](/basic_data.html#DataBunch) objects.
- a submodule named <code>models</code> that contains the models specific to this application.
- optionally, a submodule named <code>learn</code> that will contain [`Learner`](/basic_train.html#Learner) speficic to the application.
Here is a graph of the key module dependencies:

| github_jupyter |
```
from pathlib import Path
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import pylab as pl
from matplotlib.lines import Line2D
from shapely.geometry import Point
import geopandas
import seaborn as sns
from scipy import stats
from imblearn import under_sampling, over_sampling
from sklearn import preprocessing
from sklearn.model_selection import train_test_split
from sklearn.neural_network import MLPClassifier
from sklearn.metrics import classification_report
from sklearn.metrics import log_loss
from sklearn.metrics import plot_confusion_matrix
from sklearn.metrics import confusion_matrix
from sklearn.metrics import roc_auc_score
from sklearn.metrics import pairwise
from sklearn.model_selection import cross_val_score
from scipy.spatial.distance import squareform
from scipy.spatial.distance import pdist
from sklearn.cluster import KMeans
from sklearn.decomposition import PCA
%matplotlib inline
from matplotlib import rcParams
rcParams.update({'figure.autolayout': True})
#plt.style.use('dark_background')
pd.set_option('display.max_columns',500)
sns.set()
import random
SEED = 111
random.seed(SEED)
np.random.seed(SEED)
p = Path.cwd()
data_path = p.parent.parent / 'data' / 'Master Project Data'
nfirs_path = data_path / 'NFIRS Fire Incident Data.csv'
# List the columns you want to download from the NFIRS csv
cols_to_use = ['state','fdid','inc_date','oth_inj','oth_death','prop_loss',
'cont_loss','tot_loss','geoid']
# Specify particular data type for geoid column
col_dtypes = {'geoid':str}
# Read in NFIRS dataframe
nfirs = pd.read_csv(nfirs_path,
dtype = col_dtypes,
usecols = cols_to_use,
encoding='latin-1')
# Convert inc_date column values to python datetime type
nfirs['inc_date'] = pd.to_datetime(nfirs['inc_date'], infer_datetime_format=True)
#Read in ACS dataframe
ACS_path = data_path / 'ACS 5YR Block Group Data.csv'
ACS = pd.read_csv(ACS_path,
dtype = {'GEOID':'object'},
index_col = 1)
#Read in Lives Saved & House Visits dataframes
lives_saved_path = data_path / 'ARC Lives Saved Data.csv'
lives_saved = pd.read_csv(lives_saved_path)
arc_visits_path = data_path / 'ARC Response Data.csv'
arc_visits = pd.read_csv(arc_visits_path)
#NFIRS Munging
# Ensure correct calculation of tot_loss column
nfirs['tot_loss'] = nfirs['prop_loss'] + nfirs['cont_loss']
# Create mask for new severe fire variable
sev_fire_mask = (nfirs['oth_death'] > 0) | (nfirs['oth_inj'] > 0) | (nfirs['tot_loss'] >= 10000)
# By default assigns values of severe fire column as not severe
nfirs['severe_fire'] = 'not_sev_fire'
# Applies filter to severe fire column to label the severe fire instances correctly
nfirs.loc[sev_fire_mask,'severe_fire'] = 'sev_fire'
sev_nfirs = nfirs.loc[sev_fire_mask, :].copy()
# create function for additional nfirs munging
def nfirs_munging(nfirs):
""" apply nfirs munginng
:param nfirs: pandas dataframe
:return: pandas dataframe
"""
# Create new NFIRS variables based on specified thresholds of existing variables in dataframe
nfirs['had_inj'] = np.where(nfirs['oth_inj']>0,'had_inj','no_inj')
nfirs['had_death'] = np.where(nfirs['oth_death']>0,'had_death','no_death')
nfirs['10k_loss'] = np.where(nfirs['tot_loss']>=10000,'had_10k_loss','no_10k_loss')
# Extract just the numeric portion of the geoid
nfirs['geoid'] = nfirs['geoid'].str.strip('#_')
# Add a year column to be used to groupby in addition to geoid
nfirs['year'] = nfirs['inc_date'].dt.year.astype('str')
nfirs.set_index('geoid',inplace = True)
return nfirs
nfirs = nfirs_munging(nfirs)
print('nfirs', nfirs.shape)
sev_nfirs = nfirs_munging(sev_nfirs)
print('severe nfirs', sev_nfirs.shape)
# ACS Munging
# Ensures GEOID variable is in the correct format and sets it as the dataframe index
ACS['GEOID'] = ACS['GEOID'].str[2:]
ACS.set_index(['GEOID'],inplace = True)
# Captures name properies of GEOIDs for later use before filtering dataframe to be numeric features only
Names = ACS[['county_name', 'state_name']]
# Removes extraneous features (i.e. non-numeric) in the dataframe
if 'Unnamed: 0' in ACS.columns:
ACS.drop('Unnamed: 0','columns',inplace= True)
if 'NAME' in ACS.columns:
ACS.drop('NAME','columns',inplace= True)
if 'inc_pcincome' in ACS.columns:
ACS.drop('inc_pcincome','columns',inplace= True)
# Creates vector of total populations for each census block to be used to normalize total fires per year variable
tot_pop = ACS[['tot_population']]
# Drop all total count columns in ACS and keeps all percentage columns
cols = ACS.columns.to_list()
for col in cols:
if col.find('tot') != -1 :
ACS.drop(col,'columns', inplace = True)
# Integer indexing for all rows, but gets rid of county_name, state_name, and in_poverty
ACS = ACS.iloc[:,3:]
# Remove missing values from dataframe
ACS.replace([np.inf, -np.inf], np.nan,inplace = True)
ACS.dropna(inplace = True)
print('ACS', ACS.shape, 'top_pop', tot_pop.shape)
## Adjust total fires per year by the population counts
def fires_munging(nfirs):
# Creates dataframe that shows the number of fires in each census block each year
fires = pd.crosstab(nfirs.index,nfirs['year'])
fires.index.rename('GEOID',inplace = True)
# Grab total population values pulled from ACS dataframe and assign to each census block in NFIRS dataframe
fires = fires.merge(tot_pop, how = 'left', left_index = True, right_index = True)
# Remove resulting NaN/infinity values following merge
fires.replace([np.inf, -np.inf], np.nan,inplace = True)
fires.dropna(inplace = True)
# drop rows with no population count
fires = fires[fires['tot_population'] != 0 ]
# population adjustment
fires.loc[:,'2009':'2016'] = fires.loc[:,'2009':'2016'].div(fires['tot_population'], axis = 'index') * 1000
fires = fires.loc[:,:'2016']
# view fires by year across geoids; displays additional information regarding # of fires in higher percentile categories
fires.describe(percentiles=[.75, .85, .9 ,.95, .99])
# define variables to indicate census blocks in the top 10% percent of fire risk scores
top10 = fires > fires.quantile(.9)
return fires, top10
fires, top10 = fires_munging(nfirs)
print(nfirs.shape, fires.shape, top10.shape)
sev_fires, sev_top10 = fires_munging(sev_nfirs)
print(sev_nfirs.shape, sev_fires.shape, sev_top10.shape)
# function to create histogram of with # of fires on x-axis and # of census blocks on y-axis
def plotFires(df):
figsize = (20, 16)
cols = 4
rows = 2
f, axs = plt.subplots(cols,rows,figsize= figsize)
cases = df.columns.to_list()
for case in enumerate(cases):
ax = plt.subplot(cols,rows,case[0]+1)
ax.set_title('All Fires {}'.format(str(case[1])) )
plt.hist(df[case[1]],bins=[0,1,2,3,4,5,6,7,8,9,10,11,12,13,15,20,40,80,100])
# Find correlated features in ACS dataset and identify the highly correlated relationships
# Create ACS correlation matrix
corr = ACS.corr()
# Generate a mask for the upper triangle
mask = np.triu(np.ones_like(corr, dtype=np.bool))
# Set up the matplotlib figure
f, ax = plt.subplots(figsize=(11, 9))
# Generate a custom diverging colormap
cmap = sns.diverging_palette(220, 10, as_cmap=True)
#Draw the heatmap with the mask and correct aspect ratio
sns.heatmap(corr, mask=mask, cmap=cmap, vmax=.3, center=0,
square=True, linewidths=.5, cbar_kws={"shrink": .5})
# Filtering out lower/upper triangular duplicates
corr_high = corr[abs(corr) > 0.7].stack().reset_index()
corr_high = corr_high[corr_high['level_0'].astype(str)!=corr_high['level_1'].astype(str)]
corr_high['ordered-cols'] = corr_high.apply(lambda x: '-'.join(sorted([x['level_0'],x['level_1']])),axis=1)
corr_high = corr_high.drop_duplicates(['ordered-cols'])
corr_high.drop(['ordered-cols'], axis=1, inplace=True)
corr_high.columns = ['Pair Var 1', 'Pair Var 2', 'Corr Value']
# Display highly correlated pairs
print(corr_high.sort_values(by=['Corr Value'], ascending=False))
# From highly correlated pairs, remove one of the Pair Vars from the ACS dataset except for the 'mort' variables
ACS = ACS.drop(['house_pct_vacant', 'did_not_work_past_12_mo', 'house_pct_non_family', 'house_pct_rent_occupied',
'race_pct_nonwhite', 'race_pct_nonwhitenh', 'house_pct_incomplete_plumb',
'house_pct_incomplete_kitchen', 'race_pct_whitenh'], axis=1)
# Based on feature importance experiments, select features with consistence importance across annual predictions
ACS = ACS[['house_yr_pct_earlier_1939', 'house_pct_occupied', 'house_pct_family_married', 'race_pct_black',
'worked_past_12_mo', 'heat_pct_fueloil_kerosene', 'educ_bachelors', 'house_pct_live_alone',
'educ_some_col_no_grad', 'house_pct_ownd_occupied', 'house_w_home_equity_loan', 'house_val_175K_200K',
'house_val_200K_250K']]
print (ACS.head())
# Function to upsample or downsample our dataframe features if we have unbalanced classes
def resample_df(X,y,upsample=True,seed = SEED):
from sklearn.utils import resample
# check which of our two classes is overly represented
if np.mean(y) > .5:
major,minor = 1,0
else:
major,minor = 0, 1
# Add Class feature to dataframe equal to our existing dependent variable
X['Class'] = y
df_major = X[X.Class == major ]
df_minor = X[X.Class == minor ]
if upsample:
df_minor_resampled = resample(df_minor,
replace = True,
n_samples = df_major.shape[0],
random_state = seed)
combined = pd.concat([df_major,df_minor_resampled])
# Debug
#print('minor class {}, major class {}'.format(df_minor_resampled.shape[0],
#df_major.shape[0]))
else: # downsample
df_major_resampled = resample(df_major,
replace = False,
n_samples = df_minor.shape[0],
random_state = seed)
combined = pd.concat([df_major_resampled,df_minor])
#print('minor class {}, major class {}'.format(df_minor.shape[0],
#df_major_resampled.shape[0]))
y_out = combined['Class']
X_out = combined.drop('Class', axis =1)
return X_out , y_out
# Function to train model that predicts whether each census block is in the top 10% percent of fire risk scores
def train_model(top10,fires, ACS = pd.DataFrame(), nyears = 4, modeltype='LogisticRegression', seed = SEED):
from scipy.stats import zscore
# Define model types & parameters
if modeltype =='LogisticRegression':
from sklearn.linear_model import LogisticRegression
model = LogisticRegression(warm_start=True,
class_weight = 'balanced',
max_iter = 1000)
elif modeltype =='BalBagged':
from imblearn.ensemble import BalancedBaggingClassifier
from sklearn.tree import DecisionTreeClassifier
model = BalancedBaggingClassifier(base_estimator=DecisionTreeClassifier(),
n_estimators=80, sampling_strategy='auto',
random_state=0)
elif modeltype =='BalRF':
from imblearn.ensemble import BalancedRandomForestClassifier
model = BalancedRandomForestClassifier(n_estimators=80, sampling_strategy='auto',
max_depth=10, random_state=0,
max_features=None, min_samples_leaf=40)
elif modeltype =='Bagged':
from sklearn.ensemble import BaggingClassifier
from sklearn.tree import DecisionTreeClassifier
model = BaggingClassifier(base_estimator=DecisionTreeClassifier(),
n_estimators=40,
random_state=0)
elif modeltype =='RF':
from imblearn.ensemble import BalancedRandomForestClassifier
model = BalancedRandomForestClassifier(n_estimators=60,
warm_start = False,
max_depth = 10,
random_state = 0)
# Create framework to predict whether a given census block has a fire risk score in the 90th percentile
# based on the specific number of previous years' data
years = top10.columns
start_pointer = 0
end_pointer = nyears-1
y_pointer = nyears
while y_pointer < len(years):
X_start, X_end = start_pointer, end_pointer
X = fires.iloc[:,X_start:X_end].copy()
L = X.shape[1]
X.columns = ['year-{}'.format(L - year) for year in range(L)]
sm = np.sum(X, axis = 1 )
mu = np.mean(X, axis = 1)
mx = np.max(X, axis =1)
X['Sum'] = sm
X['Mean'] = mu
X['Max'] = mx
y = top10.iloc[:,y_pointer]
# merge in ACS Data into X unless NFIRS-Only model
if not ACS.empty:
X=X[['Sum','Mean','Max']] # drop all other NFIRS columns that have low feature importance scores
X = X.merge(ACS, how ='left',left_index = True, right_index = True)
X = X.dropna()
y = y.filter(X.index)
# Create 80/20 training/testing set split
X_train, X_test, y_train, y_test = train_test_split(X, y,test_size = .2 )
# Perform resampling if data classes are unbalanced
X_train, y_train = resample_df(X_train,y_train)
# Perform cross-validation
#scaler = preprocessing.StandardScaler().fit(X)
#scaler.transform(X)
#print ('Cross Val Score:')
#print(cross_val_score(model, X, y))
# Standardize features by removing the mean and scaling to unit variance
scaler = preprocessing.StandardScaler().fit(X_train)
scaler.transform(X_train)
scaler.transform(X_test)
# Fit model to training set
print('Predicting {}:'.format(years[y_pointer]) )
model = model.fit(X_train,y_train)
# Calculate training set performance
train_prediction_probs = model.predict_proba(X_train)
train_predictions = model.predict(X_train)
print (confusion_matrix(y_train, train_predictions))
print (roc_auc_score(y_train, train_prediction_probs[:,1]))
# Calculate test set performance
test_prediction_probs = model.predict_proba(X_test)
test_predictions = model.predict(X_test)
print (confusion_matrix(y_test, test_predictions))
print (roc_auc_score(y_test, test_prediction_probs[:,1]))
print (classification_report(y_test,test_predictions))
print (log_loss(y_test,test_predictions))
#Calculate feature importance for each model
if modeltype=="LogisticRegression":
feature_importance = {}
for coef, feat in zip(abs(model.coef_[0]),X_test.columns.tolist()):
feature_importance[feat] = coef
print("Feature ranking:")
print (feature_importance)
else:
if modeltype=="RF" or modeltype=="BalRF":
importances = model.feature_importances_
elif modeltype=="Bagged":
importances = np.mean([model.estimators_[i].feature_importances_ for i
in range(len(model.estimators_))], axis=0)
elif modeltype=="BalBagged":
importances = np.mean([model.estimators_[i].steps[1][1].feature_importances_ for i
in range(len(model.estimators_))], axis=0)
indices = np.argsort(importances)[::-1]
print("Feature ranking:")
for f in range(len(X_test.columns)):
print("%d. %s (%f)" % (f + 1, X_test.columns[indices[f]], importances[indices[f]]))
# Increment sliding prediction window
#start_pointer += 1
end_pointer += 1
y_pointer += 1
return model,X_test,y_test,X
# Train NFIRS Only Model and output prediction performance metrics for each year
# mdl,X_test,y_test=train_model(top10.loc[:,'2009':'2016'],fires,nyears = 5, modeltype='RF', resample = True)
# Train NFIRS + ACS Model and output prediction performance metrics for each year
mdl,X_test,y_test, X =train_model(top10.loc[:,'2009':'2016'],fires,ACS = ACS,nyears=5, modeltype='BalRF')
# Function to visualize relative feature importance for trained Logistic Regression models
def plot_LR_feat_importance(model):
# Calculate and store relative importance of model features
feature_importance = {}
for coef, feat in zip(abs(model.coef_[0]),X_test.columns.tolist()):
feature_importance[feat] = coef
data_to_plot = pd.DataFrame.from_dict(features,orient ='index').reset_index()
data_to_plot.columns = ['Variables','Importance Coefficients']
data_to_plot = data_to_plot.sort_values('Importance Coefficients', ascending=False )
data_to_plot = data_to_plot[0:10]
plt.figure(figsize=(16, 8))
sns.barplot(
x='Variables', y='Importance Coefficients',
data=data_to_plot, palette='Blues_r')
plt.xticks(
range(len(data_to_plot)),
data_to_plot['Variables'], rotation='45', size=10)
plt.xlabel('Variables', fontsize=30)
plt.ylabel('Coeff', fontsize=30)
plt.tight_layout()
plt.show()
# Display list of relative feature importance for RF models
def rf_feat_importance(model, num_feats_to_display):
importances = model.feature_importances_
indices = np.argsort(importances)[::-1]
# Print the feature ranking
print("Feature ranking:")
for f in range(num_feats_to_display):
print("%d. %s (%f)" % (f + 1, X_test.columns[indices[f]], importances[indices[f]]))
# Function to visualize example tree from random forest (only functional if model max_depth<=5)
def plot_rf_tree(model):
estimator = model.estimators_[1]
from sklearn import tree
fig, axes = plt.subplots(nrows = 1,ncols = 1,figsize = (4,4), dpi=800)
tree.plot_tree(estimator,
feature_names = X_test.columns,
class_names=y_test.index.name,
filled = True);
fig.savefig('rf_individualtree.png')
def process_data(X,y, test_size=0.2, seed=SEED):
from scipy.stats import zscore
X = np.log(X +1)
y = np.log(y+1)
print (X.shape)
# rename features and add new features
X.columns = ['year-{}'.format(year[0]+1) for year in enumerate(X.columns)]
X.columns
X['Sum'] = np.sum(X, axis = 1 )
X['mean']= np.mean(X, axis = 1)
X['median'] = np.median(x, axis = 1)
X = zscore(X.astype(float), axis=0)
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=test_size, random_state=seed)
return X_train, X_test, y_train, y_test
def plot_boundaries(X,y,model,n):
from itertools import combinations
n_elms = max(y.shape[0],1000) # only plot 1000 elements from each class
n_cols = np.ceil(np.sqrt(n))
n_rows = np.ceil(n / n_cols)
names = X.columns
combos = [c for c in combinations(range(X.shape[1]), 2) ]
W = model.coef_[0]
b = model.intercept_
for c in enumerate(combos[:n]):
x_idx = c[1][0]
y_idx = c[1][1]
plot_num = c[0]
print('subplot{} {}: {}'.format(n_rows,n_cols,c[0]))
plt.subplot(n_rows,n_cols,c[0]+1)
plt.scatter(X.iloc[:n_elms, x_idx],
X.iloc[:n_elms, y_idx],
c=y[:n_elms], edgecolors='k',
alpha = .3, cmap=plt.cm.binary)
ax = plt.gca()
x_values = np.array(ax.get_xlim())
y_values = -(b + np.dot(W[x_idx],x_values)) / W[y_idx]
plt.plot(x_values, y_values )
plt.xticks(())
plt.xlabel(names[x_idx])
plt.yticks(())
plt.ylabel(names[y_idx])
plt.show()
#plot_boundaries(X_test,y_test,mdl,16)
x = np.random.multivariate_normal([0,0,0,0],np.identity(4),1000)
x[0:500,2] += 5
x[0:500,3] -= 5
x[:500,1] += -1
x[:500,0] -= 10
y = np.zeros([1000,])
y[:500] = 1
from sklearn.linear_model import LogisticRegression
tmdl = LogisticRegression().fit(x,y)
x =pd.DataFrame(data = x,columns={'0','1','2','3'})
plot_boundaries(x,y,tmdl,12)
from itertools import combinations
c= [c for c in combinations(range(5),2)]
for a in enumerate(c[:5]):
print(a[0],a[1], a[1][0],a[1][1])
def fit_clf_model(data, model='LogisticRegression', seed=0, verbose=True, variables=None):
_allowed_models = ["LogisticRegression", "SVM", "Tree", "RandomForest"]
assert model in _allowed_models, "Invalid model name entered. Allowed options: %s" % _allowed_models
X_train, X_test, y_train, y_test = process_data(data, variables)
if verbose:
print('-' * 90)
print('[PROGRESS] Training %s classifier...' % model)
if model == 'LogisticRegression':
from sklearn.linear_model import LogisticRegression
clf = LogisticRegression(
solver='lbfgs', max_iter=1000,
random_state=seed,
).fit(X_train, y_train)
elif model == 'SVM':
from sklearn.svm import SVC
clf = SVC(
gamma='auto',
kernel='linear',
random_state=seed,
).fit(X_train, y_train)
elif model == 'Tree':
from sklearn import tree
clf = tree.DecisionTreeClassifier(
random_state=seed,
).fit(X_train,y_train)
elif model == 'RandomForest':
from sklearn.ensemble import RandomForestClassifier
clf = RandomForestClassifier(
n_estimators=100,
random_state=seed,
).fit(X_train,y_train)
else:
raise(NotImplemented)
if verbose:
print('[PROGRESS] ...done!')
msg = "\n[INFO] Train accuracy: {0:.1f} %, Test accuracy: {1:.1f} %".format(
clf.score(X_train, y_train) * 100, clf.score(X_test, y_test) * 100)
print(msg)
print('-' * 90)
print('\n')
return clf
# Train severe model
sev_mdl, sev_X_test, sev_y_test, sev_X =train_model(sev_top10.loc[:,'2009':'2016'],sev_fires,ACS = ACS,nyears=5, modeltype='BalRF')
# build a dataframe with both feature importances
def feature_impt_df(model, X_test, col_name):
importances = model.feature_importances_
features = pd.DataFrame(importances, columns=[col_name])
features['feature_name'] = X_test.columns
features = features[['feature_name', col_name]]
return features
all_features = feature_impt_df(mdl, X_test, 'all_fires')
sev_features = feature_impt_df(sev_mdl, sev_X_test, 'severe_fires')
feature_comp = pd.merge(all_features, sev_features)
feature_comp.set_index('feature_name', inplace=True)
feature_comp.to_csv('all_vs_severe_feature_importances.csv')
feature_comp.sort_values('all_fires', inplace=True)
feature_comp.plot(kind='barh', title='Feature Importances All vs Severe Fires', figsize=(12,8))
plt.savefig('feature_imp_comp.png', replace=True)
# temp = feature_comp.copy()
# temp.loc['Max', 'all_fires'] = temp.loc['Max', 'severe_fires']
# temp.plot(kind='barh', title='Feature Importances All vs Severe Fires (trimmed)', figsize=(12,8))
# plt.savefig('feature_imp_comp_trimmed.png', replace=True)
# compare predictions
all_comp = pd.DataFrame(y_test)
all_comp.rename({'2016': 'true_all_fires'}, axis='columns', inplace=True)
all_yhat = mdl.predict(X_test)
all_comp['pred_all_fires'] = all_yhat
display(all_comp.shape)
sev_comp = pd.DataFrame(sev_y_test)
sev_comp.rename({'2016': 'true_sev_fires'}, axis='columns', inplace=True)
sev_yhat = sev_mdl.predict(sev_X_test)
sev_comp['pred_sev_fires'] = sev_yhat
display(sev_comp.shape)
pred_comp = pd.merge(all_comp, sev_comp, left_index=True, right_index=True)
pred_comp.to_csv('all_vs_severe_predictions.csv')
display(pred_comp.shape)
pred_all = pred_comp.groupby(['true_all_fires', 'true_sev_fires', 'pred_all_fires']).count().reset_index()
pred_all.rename({'pred_sev_fires': 'pred_all'}, axis='columns', inplace=True)
pred_all = pred_all.set_index(['true_all_fires', 'true_sev_fires', 'pred_all_fires']).unstack()
pred_all.reset_index(inplace=True)
pred_all.columns = ['true_all_fires', 'true_sev_fires', 'pred_all_false', 'pred_all_true']
pred_sev = pred_comp.groupby(['true_all_fires', 'true_sev_fires', 'pred_sev_fires']).count().reset_index()
pred_sev.rename({'pred_all_fires': 'pred_sev'}, axis='columns', inplace=True)
pred_sev = pred_sev.set_index(['true_all_fires', 'true_sev_fires', 'pred_sev_fires']).unstack()
pred_sev.reset_index(inplace=True)
pred_sev.columns = ['true_all_fires', 'true_sev_fires', 'pred_sev_false', 'pred_sev_true']
pred = pd.merge(pred_all, pred_sev)
pred['total'] = pred[['pred_all_false', 'pred_all_true']].sum(axis=1)
pred = pred[['true_all_fires', 'true_sev_fires', 'total', 'pred_all_false', 'pred_all_true', 'pred_sev_false', 'pred_sev_true']]
pred.to_csv('all_vs_severe_predictions_counts.csv', index=False)
display(pred)
all_yhat = mdl.predict(X)
sev_yhat = sev_mdl.predict(X)
comp = pd.DataFrame(all_yhat)
comp['severe'] = sev_yhat
comp.columns = ['all', 'severe']
comp['dummy'] = 1
comp['count'] = comp.groupby(['all', 'severe']).transform('count')['dummy']
comp.drop_duplicates(inplace=True)
comp.drop('dummy', axis=1, inplace=True)
comp.sort_values('all', inplace=True)
display(comp)
```
| github_jupyter |
```
from SimPEG import Mesh, Maps, Utils
import numpy as np
from matplotlib.colors import LogNorm
from simpegem1d import (
GlobalEM1DProblem, GlobalEM1DSurveyFD, get_vertical_discretization
)
from pymatsolver import PardisoSolver
%pylab inline
import SimPEG
import h5py
import tarfile
import os
import shutil
def download_and_unzip_data(
url = "https://storage.googleapis.com/simpeg/bookpurnong/bookpurnong_inversion.tar.gz"
):
"""
Download the data from the storage bucket, unzip the tar file, return
the directory where the data are
"""
# download the data
downloads = Utils.download(url)
# directory where the downloaded files are
directory = downloads.split(".")[0]
# unzip the tarfile
tar = tarfile.open(downloads, "r")
tar.extractall()
tar.close()
return downloads, directory
# download the data
downloads, directory = download_and_unzip_data()
# Load resolve data
resolve = h5py.File(
os.path.sep.join([directory, "booky_resolve.hdf5"]),
"r"
)
river_path = resolve["river_path"].value # River path
nskip = 1
nSounding = resolve["data"][::nskip, :].shape[0] # the # of soundings
# Bird height from surface
b_height_resolve = (resolve["src_elevation"])[::nskip]
# fetch the frequencies we are considering
cpi_inds = [0, 2, 6, 8, 10] # Indices for HCP in-phase
cpq_inds = [1, 3, 7, 9, 11] # Indices for HCP quadrature
frequency_cp = resolve["frequency_cp"].value
xy = (resolve["xy"].value)[::nskip, :]
line = resolve['line'][::nskip]
data_cpi = resolve["data"][::nskip, cpi_inds].astype(float)
data_cpq = resolve["data"][::nskip, cpq_inds].astype(float)
plt.plot(river_path[:,0], river_path[:,1], 'k-')
plt.plot(xy[:,0], xy[:,1], 'k.', ms=1)
```
# Seting up a 2D mesh and model
```
from scipy.constants import mu_0
frequency = np.array([382, 1822, 7970, 35920, 130100], dtype=float)
hz = get_vertical_discretization(frequency, sigma_background=1./50)
# survey parameters
rxOffset = 7.86 # tx-rx separation
bp = -mu_0/(4*np.pi*rxOffset**3) # primary magnetic field
import scipy as sp
tri = sp.spatial.Delaunay(xy)
# Split the triangulation into connections
edges = np.r_[tri.simplices[:,:2],tri.simplices[:,1:],tri.simplices[:,[0,2]]]
# Sort and keep uniques
edges = np.sort(edges, axis=1)
edges = np.unique(edges[np.argsort(edges[:,0]),:], axis=0)
# Create 2D operator, dimensionless for now
nN = edges.shape[0]
nStn = xy.shape[0]
stn, count = np.unique(edges[:,0], return_counts = True)
col = []
row = []
dm = []
avg = []
for ii in range(nN):
row += [ii]*2
col += [edges[ii,0], edges[ii,1]]
scale = count[stn==edges[ii,0]][0]
dm += [-1., 1.]
avg += [0.5, 0.5]
D = sp.sparse.csr_matrix((dm, (row, col)), shape=(nN, nStn))
A = sp.sparse.csr_matrix((avg, (row, col)), shape=(nN, nStn))
# Kron vertically for nCz
Grad = sp.sparse.kron(D, Utils.speye(hz.size))
Avg = sp.sparse.kron(A, Utils.speye(hz.size))
figsize(10, 10)
plt.triplot(xy[:,0], xy[:,1], tri.simplices)
plt.plot(xy[:,0], xy[:,1], '.')
plt.show()
n_sounding = xy.shape[0]
b_height_resolve = resolve["src_elevation"].value[::nskip]
topo_resolve = resolve["ground_elevation"].value[::nskip]
x = xy[:,0]
y = xy[:,1]
z = topo_resolve + b_height_resolve
rx_locations = np.c_[x, y, z]
src_locations = np.c_[x, y, z]
topo = np.c_[x, y, topo_resolve].astype(float)
mapping = Maps.ExpMap(nP=int(n_sounding*hz.size))
survey = GlobalEM1DSurveyFD(
rx_locations = rx_locations,
src_locations = src_locations,
frequency = frequency,
offset = np.ones_like(frequency) * 7.86,
src_type = "VMD",
rx_type = "Hz",
field_type = 'secondary',
topo = topo
)
prob = GlobalEM1DProblem(
[], sigmaMap=mapping, hz=hz, parallel=True, n_cpu=2,
Solver=PardisoSolver
)
prob.pair(survey)
m0 = np.ones(mapping.nP) * np.log(1./50)
%%time
pred = survey.dpred(m0)
DOBS = np.empty((prob.n_sounding, 2, survey.n_frequency))
DPRED = pred.reshape((prob.n_sounding, 2, survey.n_frequency))
for i_freq in range(frequency_cp.size):
DOBS[:,0,i_freq] = data_cpi[:, i_freq]
DOBS[:,1,i_freq] = data_cpq[:, i_freq]
DOBS *= bp * 1e-6
uniq_line = np.unique(line)
i_line = 0
ind_line = line == uniq_line[i_line]
figsize(10, 5)
for i_freq in range(survey.n_frequency):
plt.semilogy(xy[ind_line,0], -DOBS[ind_line,0,i_freq], 'k')
plt.semilogy(xy[ind_line,0], -DPRED[ind_line,0,i_freq], 'k.')
for i_freq in range(survey.n_frequency):
plt.semilogy(xy[ind_line,0], -DOBS[ind_line,1,i_freq], 'r')
plt.semilogy(xy[ind_line,0], -DPRED[ind_line,1,i_freq], 'r.')
fig = plt.figure(figsize=(10, 5))
plt.plot(topo[ind_line,0], src_locations[ind_line,2]-topo[ind_line,2])
from SimPEG import (
Regularization, Directives, Inversion, InvProblem, Optimization, DataMisfit, Utils
)
hx = np.ones(n_sounding)
mesh_reg = Mesh.TensorMesh([hz, hx])
# Now we can create the regularization using the 2D mesh
reg = Regularization.Sparse(mesh_reg, mapping=Maps.IdentityMap(nP=mesh_reg.nC))
# Override the gradient operator in X with the one created above
reg.regmesh._cellDiffyStencil = Grad
# Do the same for the averaging operator
reg.regmesh._aveCC2Fy = Avg
DOBS.flatten().shape
dobs = DOBS.flatten()
%%time
std = 0.1
floor = abs(20 * bp * 1e-6) # floor of 20ppm
uncert = std*abs(dobs)+floor
np.random.seed(1)
survey.dobs = dobs
m0 = np.ones(mesh_reg.nC) * np.log(1./100.)
dmisfit = DataMisfit.l2_DataMisfit(survey)
dmisfit.W = 1./uncert
regmap = Maps.IdentityMap(mesh_reg)
# mapping is required ... for IRLS
reg = Regularization.Sparse(
mesh_reg, mapping=regmap,
alpha_s=1.,
alpha_x = 1.,
alpha_y = 10.,
)
p = 0
qx, qz = 1., 1.
reg.norms = np.c_[p, qx, qz, 0.]
# reg.norms = np.c_[p, qx, qz, 0.]
IRLS = Directives.Update_IRLS(maxIRLSiter=10, minGNiter=1)
opt = Optimization.InexactGaussNewton(maxIter = 20)
invProb = InvProblem.BaseInvProblem(dmisfit, reg, opt)
beta = Directives.BetaSchedule(coolingFactor=2, coolingRate=1)
betaest = Directives.BetaEstimate_ByEig(beta0_ratio=1.)
target = Directives.TargetMisfit()
# inv = Inversion.BaseInversion(invProb, directiveList=[beta,betaest,target])
inv = Inversion.BaseInversion(invProb, directiveList=[IRLS,betaest])
prob.counter = opt.counter = Utils.Counter()
opt.LSshorten = 0.5
opt.remember('xc')
mopt = inv.run(m0)
reg[0]
reg[1]
reg[2]
out = hist(np.log10(mapping*mopt), bins=100)
# out = hist(np.log10(mapping*invProb.l2model), bins=100)
m0
sigma = mapping * mopt
# sigma_l2 = mapping * invProb.l2model
PRED = invProb.dpred.reshape((prob.n_sounding, 2, survey.n_frequency))
i_line = 4
ind_line = line == uniq_line[i_line]
figsize(10, 5)
for i_freq in range(survey.n_frequency):
plt.semilogy(xy[ind_line,0], -DOBS[ind_line,0,i_freq], 'k')
plt.semilogy(xy[ind_line,0], -PRED[ind_line,0,i_freq], 'k.')
for i_freq in range(survey.n_frequency):
plt.semilogy(xy[ind_line,0], -DOBS[ind_line,1,i_freq], 'r')
plt.semilogy(xy[ind_line,0], -PRED[ind_line,1,i_freq], 'r.')
```
# Final 3D mesh output
```
from simpegem1d import set_mesh_1d
mesh_1d = set_mesh_1d(hz)
depth = -mesh_1d.gridN[:-1]
xyz = np.empty((hz.size, xy.shape[0], 3), order='F')
for i_xy in range(xy.shape[0]):
z = - mesh_1d.vectorCCx + topo[i_xy, 2]
x = np.ones_like(z) * xy[i_xy,0]
y = np.ones_like(z) * xy[i_xy,1]
xyz[:, i_xy, :] = np.c_[x, y, z]
Sigma = sigma.reshape((hz.size, n_sounding), order='F')
# Sigma_l2 = sigma_l2.reshape((hz.size, n_sounding), order='F')
xmin, xmax = xy[:,0].min(), xy[:,0].max()
ymin, ymax = xy[:,1].min(), xy[:,1].max()
zmin, zmax = topo[:,2].min() + depth.min(), topo[:,2].max()
zmin
iz = 2
i_line = 11
ind_line = line == uniq_line[i_line]
xz = xyz[:, ind_line, :][:,:,[0,2]].reshape((int(hz.size*ind_line.sum()), 2), order='F')
print (depth[iz])
temp = sigma.reshape((mesh_reg.vnC), order='F')[iz,:]
fig = plt.figure(figsize=(8, 8))
out = Utils.plot2Ddata(xy, temp, scale='log', contourOpts={'cmap':'viridis'}, ncontour=10)
plt.plot(xy[ind_line,0], xy[ind_line,1], 'k.')
plt.colorbar(out[0])
fig = plt.figure()
sigma_xz = Utils.mkvc(Sigma[:,ind_line])
Utils.plot2Ddata(xz, sigma_xz, scale='log', dataloc=False)
plt.fill_between(topo[ind_line, 0], topo[ind_line, 2], topo[ind_line, 1]*0. + zmax, color='w')
plt.ylim(zmax-100, zmax)
plt.gca().set_aspect(10)
# iz = 4
# i_line = 10
# ind_line = line == uniq_line[i_line]
# xz = xyz[:, ind_line, :][:,:,[0,2]].reshape((int(hz.size*ind_line.sum()), 2), order='F')
# print (depth[iz])
# temp = sigma_l2.reshape((mesh_reg.vnC), order='F')[iz,:]
# fig = plt.figure(figsize=(8, 8))
# out = Utils.plot2Ddata(xy, temp, scale='log', contourOpts={'cmap':'viridis'}, ncontour=10)
# plt.plot(xy[ind_line,0], xy[ind_line,1], 'k.')
# plt.colorbar(out[0])
# fig = plt.figure()
# sigma_xz = Utils.mkvc(Sigma_l2[:,ind_line])
# Utils.plot2Ddata(xz, sigma_xz, scale='log', dataloc=False)
# plt.fill_between(topo[ind_line, 0], topo[ind_line, 2], topo[ind_line, 1]*0. + zmax, color='w')
# plt.ylim(zmax-100, zmax)
# plt.gca().set_aspect(10)
print (reg[0][0], reg[0][1](mopt))
print (reg[0][0], reg[1][1](mopt))
print (reg[0][0], reg[2][1](mopt))
# # Use Kdtree
# xyz = xyz.reshape((hz.size*xy.shape[0], 3), order='F')
# from scipy.spatial import cKDTree
# tree = cKDTree(xyz)
# xmin, xmax = xyz[:,0].min(), xyz[:,0].max()
# ymin, ymax = xyz[:,1].min(), xyz[:,1].max()
# zmin, zmax = xyz[:,2].min(), xyz[:,2].max()
# print (xmin, xmax)
# print (ymin, ymax)
# print (zmin, zmax)
# dx = 40.
# dy = 40.
# dz = hz.min()
# nx = int(np.floor((xmax-xmin)/dx))
# ny = int(np.floor((ymax-ymin)/dy))
# nz = int(np.floor((zmax-zmin)/dz))
# x0 = [xmin-dx/2., ymin-dy/2., zmin-dz/2.]
# x = np.linspace(xmin-dx/2., xmax+dx/2., nx)
# y = np.linspace(ymin-dy/2., ymax+dy/2., ny)
# z = np.linspace(zmin-dz/2., zmax+dz/2., nz)
# mesh = Mesh.TensorMesh([np.diff(x),np.diff(y), np.diff(z)], x0=x0)
# d, inds = tree.query(mesh.gridCC, k = 10)
# w = 1.0 / (d+10.)**2
# sigma_out = np.sum(w * sigma[inds], axis=1) / np.sum(w, axis=1)
```
| github_jupyter |
# Example 4: Open Seismic Usage Template
In this example, we will walk you through how to use your files. This example should be treated as a template for you to get up and running with Open Seismic.
### Sections
4.1 **Defining the JSON Configuration**<br/>
4.2 **Organization of Files**<br/>
4.3 **Using Open Seismic's Docker Image** <br/>
## Imports
Make sure to run `pip install -r requirements.txt` and build/pull the docker image for Open Seismic.
**TODO:**
1. Make a copy of `template_dir` in `examples/assets/`. Put your files in the appropriate locations and edit the config.json to reflect correct namings.
2. Edit the `local_dir` variable and the `name_of_config` variable in the next cell with the path to the edited `template_dir` directory and the name of the configuration JSON file, respectively.
3. Run the rest of the notebook.
```
import os
from pathlib import PurePath
# PurePath is used so notebook is OS independent.
# You can specify a path such as path/to/vol by doing PurePath('path', 'to', 'vol').
# local_dir must be an absolute path.
# We have provided a way to get the absolute path, but this might not meet your needs.
cwd_path = PurePath(os.getcwd())
local_dir = cwd_path / PurePath('assets', 'template_dir_copy') # TODO
name_of_config = 'template_config.json' # TODO
```
Here are some handy imports.
```
import json
import os
import numpy as np
import matplotlib.pyplot as plt
def list_files(startpath):
for root, dirs, files in os.walk(startpath):
level = root.replace(startpath, '').count(os.sep)
indent = ' ' * 4 * (level)
print('{}{}/'.format(indent, os.path.basename(root)))
subindent = ' ' * 4 * (level + 1)
for f in files:
print('{}{}'.format(subindent, f))
open_seismic_docker = 'open_seismic'
os_path = PurePath(os.getcwd()).parent
models_path = os_path.joinpath('models')
models_mount = f' -v {models_path}:/core/python/models/'
```
## Section 4.1: Defining the JSON Configuration
Here is a peek at your JSON configuration. Make sure this lines up with your expectations.
```
json_path = local_dir.joinpath(name_of_config)
with open(str(json_path), 'rb') as f:
json_config = json.load(f)
print(json.dumps(json_config, indent=4))
json_path
```
## Section 4.2: Organization of Files
Here is the recommended file structure:
```
my_local_dir/
config.json
my_data_folder\
...
data_file_i
...
my_optimization_folder\
converter_script.sh
converter_script_helper.py
my_scripts_folder\
model.py
preprocessor.py
postprocessor.py
modelname.xml
modelname.bin
modelname.mapping
```
Here is a peek at your `local_dir` file structure. Make sure that they follow the same structure.
```
list_files(str(local_dir))
```
## Section 4.3: Using Open Seismic's Docker Image
Run the cells below to use Open Seismic!
```
run_dir = local_dir.joinpath('runs')
os_input = f'/core/mnt/{name_of_config}'
mounts = f'-v {local_dir}:/core/mnt/ -v {run_dir}:/core/runs {models_mount}'
os_cmd = f"docker run {mounts} {open_seismic_docker} ./run.sh -c {os_input}"
os_cmd
! {os_cmd}
```
Congratulations! You have completed the example series.
| github_jupyter |
```
# Dependencies
from splinter import Browser
from webdriver_manager.chrome import ChromeDriverManager
from bs4 import BeautifulSoup
import requests
import numpy as np
# Setup splinter
executable_path = {'executable_path': ChromeDriverManager().install()}
browser = Browser('chrome', **executable_path, headless=False)
from selenium import webdriver
from selenium.webdriver.chrome.options import Options
# from selenium import webdriver
# from selenium.webdriver.support.ui import WebDriverWait
# from selenium.webdriver.support import expected_conditions as EC
# from selenium.webdriver.common.by import By
# from selenium.webdriver.common.action_chains import ActionChains
# # driver = webdriver.Chrome()
# # driver.get('https://www.soleretriever.com/raffles')
# elem = WebDriverWait(driver, 20).until(EC.presence_of_element_located((By.XPATH, '/html/body/div[6]/div/div/div[2]/form/div[6]/button')))
# action = ctionChains(driver)
# action.move_to_element(elem).move_by_offset(250, 0).click().perform()
# PATH = "C:\Program Files (x86)\chromedriver.exe"
# CHROME_DRIVER_PATH = "C:\Program Files (x86)\chromedriver.exe"
# BIN_LOC = "C:\Program Files\BraveSoftware\Brave-Browser\Application\\brave.exe"
# options = Options()
# options.binary_location = BIN_LOC
# driver = webdriver.Chrome(options=options, executable_path=CHROME_DRIVER_PATH)
# # Retrieve page with the requests module
# response = requests.get(url)
# # Create BeautifulSoup object; parse with 'html.parser'
# soup = BeautifulSoup(response.text, 'html.parser')
CHROME_DRIVER_PATH = "C:\Program Files (x86)\chromedriver.exe"
chrome_options = webdriver.ChromeOptions();
chrome_options.add_experimental_option("excludeSwitches", ['enable-automation']);
driver = webdriver.Chrome(options=chrome_options, executable_path=CHROME_DRIVER_PATH);
# URL of page to be scraped
url = 'http://house.speakingsame.com/p.php?q=Perth%2C+WA'
# browser.visit(url)
driver.get(url)
# URL of page to be scraped
url = 'http://house.speakingsame.com/p.php?q=Perth%2C+WA'
browser.visit(url)
driver.title
# desired_cap = {
# 'browser': 'chrome',
# 'browser_version': 'latest',
# 'os': 'Windows',
# 'os_version': '10',
# 'build': 'Python Sample Build',
# 'name': 'Pop-ups testing'
# }
# desired_cap["chromeOptions"] = {}
# desired_cap["chromeOptions"]["excludeSwitches"] = ["disable-popup-blocking"]
# driver = webdriver.Remote(
# command_executor='http://house.speakingsame.com/p.php?q=Perth&p=0&s=1&st=&type=&count=300®ion=Perth&lat=0&lng=0&sta=wa&htype=&agent=0&minprice=0&maxprice=0&minbed=0&maxbed=0&minland=0&maxland=0',
# desired_capabilities=desired_cap
# )
l = driver.find_elements_by_class_name("addr")
for i in l:
print("address:"+ i.text)
driver.find_element_by_partial_link_text('Next').click()
addresses = []
properties_info = []
pages = np.arange(0, 2, 1)
for page in pages:
page_url = "http://house.speakingsame.com/p.php?q=Perth&p="+ str(page) +"&s=1&st=&type=&count=300®ion=Perth&lat=0&lng=0&sta=wa&htype=&agent=0&minprice=0&maxprice=0&minbed=0&maxbed=0&minland=0&maxland=0"
driver.get(page_url)
# html = driver.html
# soup = BeautifulSoup(html, 'html.parser')
# Scrape the address
addr_list = driver.find_elements_by_class_name("addr")
for address in addr_list:
addresses.append(address.text)
# Scrape the rest of property information
# tables = driver.find_elements_by_tag_name('table', style="font-size:13px") #tables list (except 0,1 index) contians list of properties in the page
# tables = driver.find_element(By.XPATH,"//table[@style='font-size:13px']")
tables = driver.find_elements_by_xpath("//table[@style='font-size:13px']")
# tds = driver.find_element_by_xpath("//table[@style='font-size:13px']/tbody/tr/td").text
for i in range(2,12):
try:
info = []
trs = tables[i].find_elements_by_xpath('tr')
for tr in trs:
td = tr.find_element_by_xpath('td')
info.append(td.text)
except:
info.append('')
properties_info.append(info)
# #once all data scraped from 1st page, moving onto the next
# next_page_url = 'http://house.speakingsame.com/p.php?q=Perth&p='+str(page+1)+'&s=1&st=&type=&count=300®ion=Perth&lat=0&lng=0&sta=wa&htype=&agent=0&minprice=0&maxprice=0&minbed=0&maxbed=0&minland=0&maxland=0'
# driver.find_element_by_link_text(next_page_url).click()
next_click = driver.find_element_by_partial_link_text('Next')
driver.execute_script("arguments[0].click();", next_click)
properties_info
len(tds)
tds
len(properties_info)
print(tables.text)
tables[3].find_elements_by_xpath("//tr")
for i in tds:
print(i.text)
len(addresses)
Properties = []
for i in range(0, len(properties_info)):
Property_dict = {}
Property_dict['address'] = addresses[i]
for j in range (0, len(properties_info[i])):
try:
if properties_info[i][j].split(' ')[0] == "Sold":
Property_dict['price'] = properties_info[i][j].split(' ')[1]
Property_dict['sold date'] = properties_info[i][j].split('in ')[1]
if properties_info[i][j].split(' ')[0] in ["Apartment:", "House:", "Townhouse:"]:
Property_dict['type'] = properties_info[i][j].split(' ')[0].split(':')[0]
Property_dict['bedrooms'] = properties_info[i][j].split(' ')[1]
try:
Property_dict['bathrooms'] = properties_info[i][j].split(' ')[3]
except:
Property_dict['bathrooms'] = ''
try:
Property_dict['car space'] = properties_info[i][j].split(' ')[5]
except:
Property_dict['car space'] = ''
if properties_info[i][j].split(' ')[0] == "Agent:":
Property_dict['agent'] = properties_info[i][j].split(': ')[1]
if properties_info[i][j].split(' ')[0] == "Rent":
Property_dict['rent'] = properties_info[i][j].split(' ')[1]
Property_dict['rent date'] = properties_info[i][j].split('in ')[1]
if properties_info[i][j].split(' ')[1] == "size:":
if properties_info[i][j].split(' ')[0] == "Land":
Property_dict['Land size'] = properties_info[i][j].split(' ')[2]
try:
if properties_info[i][j].split(' ')[5] == "Building":
Property_dict['Building size'] = properties_info[i][j].split(' ')[7]
except:
continue
elif properties_info[i][j].split(' ')[0] == "Building":
Property_dict['Building size'] = properties_info[i][j].split(' ')[2]
except:
continue
Properties.append(Property_dict)
Properties
len(Properties)
# URL of page to be scraped
url = 'http://house.speakingsame.com/p.php?q=Perth%2C+WA'
browser.visit(url)
# # Retrieve page with the requests module
# response = requests.get(url)
# # Create BeautifulSoup object; parse with 'html.parser'
# soup = BeautifulSoup(response.text, 'html.parser')
addresses = []
properties_info = []
pages = np.arange(0, 2, 1)
for page in pages:
page_url = "http://house.speakingsame.com/p.php?q=Perth&p="+ str(page) +"&s=1&st=&type=&count=300®ion=Perth&lat=0&lng=0&sta=wa&htype=&agent=0&minprice=0&maxprice=0&minbed=0&maxbed=0&minland=0&maxland=0"
html = browser.html
soup = BeautifulSoup(html, 'html.parser')
addr_spans = soup.find_all('span', class_='addr')
addresses = []
for span in addr_spans:
addresses.append(span.a.text)
addresses
tables = soup.find_all('table', style="font-size:13px") #tables list (except 0,1 index) contians list of properties in the page
for i in range(2,12):
try:
info = []
trs = tables[i].find_all('tr')
for tr in trs:
info.append(tr.td.text)
except:
info.append('')
properties_info.append(info)
#once all data scraped from 1st page, moving onto the next
next_page_url = 'http://house.speakingsame.com/p.php?q=Perth&p='+str(page+1)+'&s=1&st=&type=&count=300®ion=Perth&lat=0&lng=0&sta=wa&htype=&agent=0&minprice=0&maxprice=0&minbed=0&maxbed=0&minland=0&maxland=0'
driver.find_element_by_link_text(next_page_url).click()
properties_info
tables = soup.find_all('table', style="font-size:13px") #tables list (except 0,1 index) contians list of properties in the page
page_info = []
for i in range(2,12):
try:
info = []
trs = tables[i].find_all('tr')
for tr in trs:
info.append(tr.td.text)
except:
info.append('')
page_info.append(info)
page_info
Properties = []
for i in range(0,10):
Property_dict = {}
Property_dict['address'] = addresses[i]
for j in range (0, len(page_info[i])):
try:
if page_info[i][j].split(' ')[0] == "Sold":
Property_dict['price'] = page_info[i][j].split(' ')[1]
Property_dict['sold date'] = page_info[i][j].split('in ')[1]
if page_info[i][j].split(' ')[0] in ["Apartment:", "House:", "Townhouse:"]:
Property_dict['type'] = page_info[i][j].split(' ')[0].split(':')[0]
Property_dict['bedrooms'] = page_info[i][j].split(' ')[1]
try:
Property_dict['bathrooms'] = page_info[i][j].split(' ')[3]
except:
Property_dict['bathrooms'] = ''
try:
Property_dict['car space'] = page_info[i][j].split(' ')[5]
except:
Property_dict['car space'] = ''
if page_info[i][j].split(' ')[0] == "Agent:":
Property_dict['agent'] = page_info[i][j].split(': ')[1]
if page_info[i][j].split(' ')[0] == "Rent":
Property_dict['rent'] = page_info[i][j].split(' ')[1]
Property_dict['rent date'] = page_info[i][j].split('in ')[1]
if page_info[i][j].split(' ')[1] == "size":
if page_info[i][j].split(' ')[0] == "Land":
Property_dict['Land size'] = page_info[i][j].split(' ')[2]
try:
if page_info[i][j].split(' ')[5] == "Building":
Property_dict['Building size'] = page_info[i][j].split(' ')[7]
except:
continue
elif page_info[i][j].split(' ')[0] == "Building":
Property_dict['Building size'] = page_info[i][j].split(' ')[2]
except:
continue
Properties.append(Property_dict)
Properties
```
| github_jupyter |
# Maxpooling Layer
In this notebook, we add and visualize the output of a maxpooling layer in a CNN.
A convolutional layer + activation function, followed by a pooling layer, and a linear layer (to create a desired output size) make up the basic layers of a CNN.
<img src='notebook_ims/CNN_all_layers.png' height=50% width=50% />
### Import the image
```
import cv2
import matplotlib.pyplot as plt
%matplotlib inline
# TODO: Feel free to try out your own images here by changing img_path
# to a file path to another image on your computer!
img_path = 'data/udacity_sdc.png'
# load color image
bgr_img = cv2.imread(img_path)
# convert to grayscale
gray_img = cv2.cvtColor(bgr_img, cv2.COLOR_BGR2GRAY)
# normalize, rescale entries to lie in [0,1]
gray_img = gray_img.astype("float32")/255
# plot image
plt.imshow(gray_img, cmap='gray')
plt.show()
```
### Define and visualize the filters
```
import numpy as np
## TODO: Feel free to modify the numbers here, to try out another filter!
filter_vals = np.array([[-1, -1, 1, 1], [-1, -1, 1, 1], [-1, -1, 1, 1], [-1, -1, 1, 1]])
print('Filter shape: ', filter_vals.shape)
# Defining four different filters,
# all of which are linear combinations of the `filter_vals` defined above
# define four filters
filter_1 = filter_vals
filter_2 = -filter_1
filter_3 = filter_1.T
filter_4 = -filter_3
filters = np.array([filter_1, filter_2, filter_3, filter_4])
# For an example, print out the values of filter 1
print('Filter 1: \n', filter_1)
```
### Define convolutional and pooling layers
You've seen how to define a convolutional layer, next is a:
* Pooling layer
In the next cell, we initialize a convolutional layer so that it contains all the created filters. Then add a maxpooling layer, [documented here](http://pytorch.org/docs/stable/_modules/torch/nn/modules/pooling.html), with a kernel size of (2x2) so you can see that the image resolution has been reduced after this step!
A maxpooling layer reduces the x-y size of an input and only keeps the most *active* pixel values. Below is an example of a 2x2 pooling kernel, with a stride of 2, applied to a small patch of grayscale pixel values; reducing the x-y size of the patch by a factor of 2. Only the maximum pixel values in 2x2 remain in the new, pooled output.
<img src='notebook_ims/maxpooling_ex.png' height=50% width=50% />
```
import torch
import torch.nn as nn
import torch.nn.functional as F
# define a neural network with a convolutional layer with four filters
# AND a pooling layer of size (2, 2)
class Net(nn.Module):
def __init__(self, weight):
super(Net, self).__init__()
# initializes the weights of the convolutional layer to be the weights of the 4 defined filters
k_height, k_width = weight.shape[2:]
# assumes there are 4 grayscale filters
self.conv = nn.Conv2d(1, 4, kernel_size=(k_height, k_width), bias=False)
self.conv.weight = torch.nn.Parameter(weight)
# define a pooling layer
self.pool = nn.MaxPool2d(2, 2)
def forward(self, x):
# calculates the output of a convolutional layer
# pre- and post-activation
conv_x = self.conv(x)
activated_x = F.relu(conv_x)
# applies pooling layer
pooled_x = self.pool(activated_x)
# returns all layers
return conv_x, activated_x, pooled_x
# instantiate the model and set the weights
weight = torch.from_numpy(filters).unsqueeze(1).type(torch.FloatTensor)
model = Net(weight)
# print out the layer in the network
print(model)
```
### Visualize the output of each filter
First, we'll define a helper function, `viz_layer` that takes in a specific layer and number of filters (optional argument), and displays the output of that layer once an image has been passed through.
```
# helper function for visualizing the output of a given layer
# default number of filters is 4
def viz_layer(layer, n_filters= 4):
fig = plt.figure(figsize=(20, 20))
for i in range(n_filters):
ax = fig.add_subplot(1, n_filters, i+1)
# grab layer outputs
ax.imshow(np.squeeze(layer[0,i].data.numpy()), cmap='gray')
ax.set_title('Output %s' % str(i+1))
```
Let's look at the output of a convolutional layer after a ReLu activation function is applied.
#### ReLu activation
A ReLu function turns all negative pixel values in 0's (black). See the equation pictured below for input pixel values, `x`.
<img src='notebook_ims/relu_ex.png' height=50% width=50% />
```
# plot original image
plt.imshow(gray_img, cmap='gray')
# visualize all filters
fig = plt.figure(figsize=(12, 6))
fig.subplots_adjust(left=0, right=1.5, bottom=0.8, top=1, hspace=0.05, wspace=0.05)
for i in range(4):
ax = fig.add_subplot(1, 4, i+1, xticks=[], yticks=[])
ax.imshow(filters[i], cmap='gray')
ax.set_title('Filter %s' % str(i+1))
# convert the image into an input Tensor
gray_img_tensor = torch.from_numpy(gray_img).unsqueeze(0).unsqueeze(1)
# get all the layers
conv_layer, activated_layer, pooled_layer = model(gray_img_tensor)
# visualize the output of the activated conv layer
viz_layer(activated_layer)
```
### Visualize the output of the pooling layer
Then, take a look at the output of a pooling layer. The pooling layer takes as input the feature maps pictured above and reduces the dimensionality of those maps, by some pooling factor, by constructing a new, smaller image of only the maximum (brightest) values in a given kernel area.
Take a look at the values on the x, y axes to see how the image has changed size.
```
# visualize the output of the pooling layer
viz_layer(pooled_layer)
```
| github_jupyter |
ADS Sample Notebook.
Copyright (c) 2021 Oracle, Inc. All rights reserved. Licensed under the Universal Permissive License v 1.0 as shown at https://oss.oracle.com/licenses/upl.
***
# <font color=red>Working with an ADSDataset Object</font>
<p style="margin-left:10%; margin-right:10%;">by the <font color=teal> Oracle Cloud Infrastructure Data Science Team </font></p>
***
## Overview of this Notebook
One of the most important elements of any data science project is the data itself. This notebook demonstrates how to work with the `ADSDataset` class.
---
## Prerequisites:
- Experience with the topic: Novice
- Professional experience: None
---
## Business Uses:
The `ADSDataset` class is a flexible container for working with data. It is backed by `dask` and allows distributed operations on the data. This greatly improves the ability of the data scientist to work on large scale problems.
---
## Objectives:
To demonstrate the following methods features of the `ADSDataset` class:
- <a href='#columnbasedops'>Column Based Operations</a>
- <a href='#rename_columns'>Renaming Columns</a>
- <a href='#feature_types'>Feature Type</a>
- <a href='#assign_column'>Assigning Values to a Column</a>
- <a href='#astype'>Altering a Column's Data Type</a>
- <a href='#merge'>Merging ADSDataset Objects</a>
- <a href='#target'>Target</a>
- <a href='#correlation'>Correlation</a>
- <a href='#corr'>Correlation Calculation</a>
- <a href='#showcorr'>Correlation Visualization</a>
- <a href='#reference'>References</a>
***
<font color=gray>Datasets are provided as a convenience. Datasets are considered Third Party Content and are not considered Materials
under your agreement with Oracle applicable to the Services.
The `wine` dataset license is available [here](https://github.com/scikit-learn/scikit-learn/blob/master/COPYING)
</font>
***
```
import ads
import logging
import pandas as pd
import numpy as np
import warnings
from ads.dataset.dataset_browser import DatasetBrowser
from ads.dataset.factory import DatasetFactory
from os import path
warnings.filterwarnings('ignore')
logging.basicConfig(format='%(levelname)s:%(message)s', level=logging.ERROR)
ads.set_documentation_mode(False)
```
<a id='columnbasedops'></a>
# Column Based Operations
The sections below demonstrate the common operations that a data scientist will perform. For the purposes of the example, a small `ADSDataset` will be created with some basic student information.
```
data = [["ID", "Name", "GPA"],
[1, "Bob", 3.7],
[2, "Sam", 4.3],
[3, "Erin", 2.6]]
df = pd.DataFrame(data[1:], columns=data[0])
ds = DatasetFactory.open(df)
ds.head()
```
<a id='rename_columns'></a>
## Renaming Columns
The `ADSDataset` has the method `rename_columns` which is used to rename the columns. It takes a JSON object where the key is the old column name and the value is the new column name. If the column name does not exist, the key is ignored. Note, the column names are processed in the order listed in the JSON. Thus, it is possible to swap column names if the order of updating does not conflict with any column name.
In this example, the column `ID` will be changed to `StudentID` and `Name` will be changed to `GivenName`.
```
rename = ds.rename_columns({'ID': "StudentID", 'Name': 'GivenName'})
rename.columns
```
<a id='feature_types'></a>
## Feature Type
Each column in an `ADSDataset` object has a feature type. This determines how the information will be used in modeling. The `feature_types` attribute provides information on them. Common feature types are ordinal, categorical and continuous.
```
for key in ds.feature_types:
print('{} is {}'.format(key, ds.feature_types[key]['type']))
```
Calling `feature_types` returns a dictionary with information about the feature. This would include its type, percentage of missing values, the low-level data type and summary statistics. The summary statistics vary for each feature type. For example, continuous features report measures of centrality (mode, median, mean), kurtosis, skewness, variance, standard deviation, count, the percentage of outliers, observed range (minimum and maximum) and quantiles (25, 50 and 75th).
```
ds.feature_types['GPA']
```
<a id='assign_column'></a>
## Assigning Values to a Column
The `assign_column` will add a new column to an `ADSDataset` object or update the values of an existing column. The method accepts the column name and then a list of values.
```
surname = ds.assign_column("Surname", ['Smith', 'Jones', 'Allan'])
surname
```
It is possible to update the values in a column using a lamba function. In this example, the student IDs will be increased by 1000.
```
new_id = ds.assign_column("ID", lambda x: int(x+1000))
new_id
```
When values are updated, the `feature_types` information is automatically updated.
```
new_id.feature_types['ID']
```
<a id='astype'></a>
## Altering a Column's Data Type
The student ID column is an ordinal data type. The `astype` method allows this to be changed. It takes a JSON object where the key is the column name and the value is the new column type. If the column name does not exist, an error is created.
In this example, the column `ID` will be changed to `categorical`
```
new_type = ds.astype({'ID': 'categorical'})
new_type.feature_types['ID']['type']
```
<a id='merge'></a>
## Merging ADSDataset Objects
Two `ADSDataset`s can be merged with the `merge` method. In this example an `ADSDataset` will be created with two columns that represent the student's surname and if the are on the Honors list or not.
```
data = [["Surname", "Honors"],
['Smith', True],
['Jones', True],
['Allan', False]]
df = pd.DataFrame(data[1:], columns=data[0])
honors = DatasetFactory.open(df)
honors.head()
```
The two datasets can be merged with:
```
honors_list = ds.merge(honors)
honors_list
```
<a id='target'></a>
<a id='target'></a>
# Target
The target in a dataset is the value that is to be predicted. The target can be set with the `target` method and it accepts the name of a column. By setting the target, the class of the object will be changed to an object that is customized for the properties of the target. In the example below the target will be set to the `Honors` column. Since `Honors` is binary-valued the `ADSDataset` will be converted to a `BinaryClassificationDataset` object.
ADS will attempt to determine which class is the positive class for the `BinaryClassificationDataset` object. However, this can be sent manually with the `set_positive_class()` method. The parameter is the value of the positive class.
```
honors_target = honors_list.set_target("Honors")
type(honors_target)
```
The `target.show_in_notebook()` will generate a plot of the target column. The type of plot that is generated is specific to the data type. Generally, this is a bar plot for categorical and ordinal data. A histogram is used for continuous data.
```
honors_target.target.show_in_notebook()
```
The `target.show_in_notebook` method also takes a list of columns in the `feature_names` parameter. It will generate one plot for each listed feature against the target column.
```
honors_target.target.show_in_notebook(feature_names=["GPA", "Name"])
```
The `Honors` target resulted in a `BinaryClassificationDataset` because it had true and false values and any prediction that would be done on that type of data would be a binary classification problem. However, changing the target to the `GPA` column will result in a `RegressionDataset` because `GPA` is a continuous variable.
```
type(honors_list.set_target("GPA"))
```
Changing the target to `Name` will result in a `MultiClassClassificationDataset` object because the data is categorical, like the `Honors` column, but there are more than two categories. Therefore, the prediction problem would be a multiclass classification problem.
```
type(honors_list.set_target("Name"))
```
<a id='correlation'></a>
# Correlation
The `corr()` methods uses the following techniques to calculate the correlation based on the data types:
- Continuous-continuous: `Pearson` method [(link)](https://en.wikipedia.org/wiki/Pearson_correlation_coefficient). The correlations range from -1 to 1.
- Categorical-categorical: `Cramer's V`[(link)](https://en.wikipedia.org/wiki/Cram%C3%A9r%27s_V). The correlations range from 0 to 1.
- Continuous-categorical: `Correlation Ratio`[(link)](https://en.wikipedia.org/wiki/Correlation_ratio). The correlations range from 0 to 1.
**Note:** `Continuous` features consist of `continuous` and `ordinal` type.
`Categorical` features consist of `categorical` and `zipcode` type.
```
ds = DatasetFactory.open(
path.join("/", "opt", "notebooks", "ads-examples", "oracle_data", "orcl_attrition.csv"),
target="Attrition").set_positive_class('Yes')
```
To check the `Datatype`, we can use `summary` function.
```
ds.summary()
```
<a id='corr'></a>
## Correlation Calculation
- `correlation_methods`: Methods to calculate the correlation and it defaults to 'pearson'.
- `frac`: The portion of the data used to calculate the correlation. It must be greater than 0 and less than or equal to 1. Defaults to 1.
- `nan_threshold`: Drop a column from the analysis if the proportion of values that are NaN exceed this threshold. Valid values are 0 to 1. Default is 0.8.
- `force_recompute`: Force a recompute of the cached correlation matrices. Default is False.
**Note**: For Cramer's V, a bias correction is applied, but not the Yate's correction for continuity([link](https://en.wikipedia.org/wiki/Yates%27s_correction_for_continuity)).
```
cts_vs_cts = ds.corr()
```
Can select single `correlation_methods` from 'pearson', 'cramers v', and 'correlation ratio', parse in as a string
```
cat_vs_cat = ds.corr(correlation_methods='cramers v')
```
Or choose multiple ones among 'pearson', 'cramers v', and 'correlation ratio', parse in as a list
```
cat_vs_cat, cat_vs_cts = ds.corr(correlation_methods=['cramers v', 'correlation ratio'])
```
Use 'all' in `correlation_methods` equivalent to ['pearson', 'cramers v', 'correlation ratio']
```
cts_vs_cts, cat_vs_cat, cat_vs_cts = ds.corr(correlation_methods='all')
```
Use Pearson's Correlation between continuous features
```
cts_vs_cts
```
Use Cramer's V correlations between categorical features
```
cat_vs_cat
```
Use Correlation Ratio Correlation between categorical and continuous features
```
cat_vs_cts
```
<a id='showcorr'></a>
## Correlation Visualization
The `show_corr()` method creates a heatmap or barplot for the pairwise correlations. The parameter `correlation_methods` assigns the methods the correlation methods to use. By default, this is the 'pearson' correlation. You can select one or more from 'pearson', 'cramers v', and 'correlation ratio'. Or set it to 'all' to show all correlation charts. The usage is the same as in the `corr()` method.
The `correlation_threshold` method applies a filter to the correlation matrices and only exhibits the pairs whose correlation values are greater than or equal to the `correlation_threshold`.
The correlation results are cached as they are expensive to compute. If the parameters`nan_threshold`, `frac` or `correlation_target` are changed, the parameter `force_recompute` need to be set to be `True`. Otherwise, it will output the old result.
When the `correlation_target` parameter is not `None`, the `plot_type` can be `'bar'` or `'heatmap'`. However, when `plot_type` is set to be `'bar'`, `correlation_target` also has to be set.
The following cell will show all the correlation charts.
```
ds.show_corr(plot_type='heatmap', correlation_methods='all')
```
<a id='reference'></a>
# 4. References
- <a href="https://docs.cloud.oracle.com/en-us/iaas/tools/ads-sdk/latest/index.html">Oracle ADS</a>
| github_jupyter |
<a href="https://www.kaggle.com/code/ritvik1909/named-entity-recognition-attention" target="_blank"><img align="left" alt="Kaggle" title="Open in Kaggle" src="https://kaggle.com/static/images/open-in-kaggle.svg"></a>
# Named Entity Recognition (NER)
NER is an information extraction technique to identify and classify named entities in text. These entities can be pre-defined and generic like location names, organizations, time and etc, or they can be very specific like the example with the resume.
The goal of a named entity recognition (NER) system is to identify all textual mentions of the named entities. This can be broken down into two sub-tasks: identifying the boundaries of the NE, and identifying its type.
Named entity recognition is a task that is well-suited to the type of classifier-based approach. In particular, a tagger can be built that labels each word in a sentence using the IOB format, where chunks are labelled by their appropriate type.
The IOB Tagging system contains tags of the form:
* B - {CHUNK_TYPE} – for the word in the Beginning chunk
* I - {CHUNK_TYPE} – for words Inside the chunk
* O – Outside any chunk
## Approaches to NER
* **Classical Approaches:** mostly rule-based.
* **Machine Learning Approaches:** there are two main methods in this category:
* Treat the problem as a multi-class classification where named entities are our labels so we can apply different classification algorithms. The problem here is that identifying and labeling named entities require thorough understanding of the context of a sentence and sequence of the word labels in it, which this method ignores that.
* Conditional Random Field (CRF) model. It is a probabilistic graphical model that can be used to model sequential data such as labels of words in a sentence. The CRF model is able to capture the features of the current and previous labels in a sequence but it cannot understand the context of the forward labels; this shortcoming plus the extra feature engineering involved with training a CRF model, makes it less appealing to be adapted by the industry.
* **Deep Learning Approaches:** Bidirectional RNNs, Transformers
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from datasets import load_dataset
plt.style.use('fivethirtyeight')
```
# Getting the Dataset and EDA
We will be working on S800 Corpus, which is a novel abstract-based manually annotated corpus. S800 comprises 800 PubMed abstracts in which organism mentions were identified and mapped to the corresponding NCBI Taxonomy identifiers.
It is available on Hugging Face Datasets Hub
```
dataset = load_dataset('species_800')
train_df = pd.DataFrame(dataset['train']).explode(['tokens', 'ner_tags']).dropna()
valid_df = pd.DataFrame(dataset['validation']).explode(['tokens', 'ner_tags']).dropna()
test_df = pd.DataFrame(dataset['test']).explode(['tokens', 'ner_tags']).dropna()
train_df.head()
```
Here ner_tag 1 indicates B
and ner_tag 2 indicates I
while ner_tag 0 indicates O
for simplicity we will combine B and I
```
train_df['ner_tags'] = train_df['ner_tags'].apply(lambda x: 1 if x > 0 else 0)
valid_df['ner_tags'] = valid_df['ner_tags'].apply(lambda x: 1 if x > 0 else 0)
test_df['ner_tags'] = test_df['ner_tags'].apply(lambda x: 1 if x > 0 else 0)
```
**Create list of list of tuples to differentiate each sentence from each other**
```
class SentenceGetter(object):
def __init__(self, dataset, word_col, tag_col, sent_id_col):
self.n_sent = 1
self.dataset = dataset
self.empty = False
agg_func = lambda s: [
(w, t) for w,t in zip(s[word_col].values.tolist(), s[tag_col].values.tolist())
]
self.grouped = self.dataset.groupby(sent_id_col).apply(agg_func)
self.sentences = [s for s in self.grouped]
def get_next(self):
try:
s = self.grouped["Sentence: {}".format(self.n_sent)]
self.n_sent += 1
return s
except:
return None
train_getter = SentenceGetter(dataset=train_df, word_col='tokens', tag_col='ner_tags', sent_id_col='id')
valid_getter = SentenceGetter(dataset=valid_df, word_col='tokens', tag_col='ner_tags', sent_id_col='id')
test_getter = SentenceGetter(dataset=test_df, word_col='tokens', tag_col='ner_tags', sent_id_col='id')
train_sentences = train_getter.sentences
valid_sentences = valid_getter.sentences
test_sentences = test_getter.sentences
print('Sample Sentence')
print(train_sentences[0])
fig, ax = plt.subplots(figsize=(20, 6))
ax.hist([len(s) for s in train_sentences], bins=50)
ax.set_title('Number of words in each Sentence')
maxlen = max([len(s) for s in train_sentences])
print('Number of Sentences:', len(train_sentences))
print ('Maximum sequence length:', maxlen)
words = list(set(train_df["tokens"].values))
words.append("ENDPAD")
words.append("UNK")
n_words = len(words)
print('Number of unique words:', n_words)
```
**Converting words to numbers and numbers to words**
```
word2idx = {w: i for i, w in enumerate(words)}
idx2word = {i: w for i, w in enumerate(words)}
```
# Modelling
```
import tensorflow as tf
from tensorflow import keras
import tensorflow.keras.layers as L
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing import sequence
from tensorflow.keras.callbacks import EarlyStopping, CSVLogger, ReduceLROnPlateau
from tensorflow.keras.utils import plot_model, to_categorical
```
**Lets first tokenize the data**
```
X_train = [[word2idx.get(w[0], len(word2idx)-1) for w in s] for s in train_sentences]
X_train = sequence.pad_sequences(maxlen=maxlen, sequences=X_train, padding="post",value=n_words - 2)
y_train = [[[w[1]] for w in s] for s in train_sentences]
y_train = sequence.pad_sequences(maxlen=maxlen, sequences=y_train, padding="post", value=0)
y_train = np.array([to_categorical(i, num_classes=2) for i in y_train])
print('X shape', X_train.shape, 'y shape', y_train.shape)
X_valid = [[word2idx.get(w[0], len(word2idx)-1) for w in s] for s in valid_sentences]
X_valid = sequence.pad_sequences(maxlen=maxlen, sequences=X_valid, padding="post",value=n_words - 2)
y_valid = [[[w[1]] for w in s] for s in valid_sentences]
y_valid = sequence.pad_sequences(maxlen=maxlen, sequences=y_valid, padding="post", value=0)
y_valid = np.array([to_categorical(i, num_classes=2) for i in y_valid])
print('X shape', X_valid.shape, 'y shape', y_valid.shape)
X_test = [[word2idx.get(w[0], len(word2idx)-1) for w in s] for s in test_sentences]
X_test = sequence.pad_sequences(maxlen=maxlen, sequences=X_test, padding="post",value=n_words - 2)
y_test = [[[w[1]] for w in s] for s in test_sentences]
y_test = sequence.pad_sequences(maxlen=maxlen, sequences=y_test, padding="post", value=0)
y_test = np.array([to_categorical(i, num_classes=2) for i in y_test])
print('X shape', X_test.shape, 'y shape', y_test.shape)
```
**Training Parameters**
```
class config():
VOCAB = n_words
MAX_LEN = maxlen
N_OUPUT = 2
EMBEDDING_VECTOR_LENGTH = 50
DENSE_DIM = 32
NUM_HEADS = 2
OUTPUT_ACTIVATION = 'softmax'
LOSS = 'categorical_crossentropy'
OPTIMIZER = 'adam'
METRICS = ['accuracy']
MAX_EPOCHS = 100
```
**Lets define a standard Transformer Encoder Block**
```
class TransformerEncoder(L.Layer):
def __init__(self, embed_dim, dense_dim, num_heads, **kwargs):
super().__init__(**kwargs)
self.embed_dim = embed_dim
self.dense_dim = dense_dim
self.num_heads = num_heads
self.attention = L.MultiHeadAttention(num_heads=num_heads, key_dim=embed_dim)
self.dense_proj = keras.Sequential([L.Dense(dense_dim, activation='relu'), L.Dense(embed_dim)])
self.layernorm1 = L.LayerNormalization()
self.layernorm2 = L.LayerNormalization()
def call(self, inputs, mask=None):
if mask is not None:
mask = mask[: tf.newaxis, :]
attention_output = self.attention(inputs, inputs, attention_mask=mask)
proj_input = self.layernorm1(inputs + attention_output)
proj_output = self.dense_proj(proj_input)
return self.layernorm2(proj_input + proj_output)
def get_config(self):
config = super().get_confog()
config.update({
"embed_dim": self.embed_dim,
"num_heads": self.num_heads,
"dense_dim": self.dense_dim
})
return config
```
**Some standard Callbacks**
```
es = EarlyStopping(monitor='val_loss', mode='min', patience=10, restore_best_weights=True)
rlp = ReduceLROnPlateau(monitor='loss', patience=3)
csv_logger = CSVLogger('training_log.csv')
```
**Lets define our model**
A token classification is pretty simple and similar to that of sequence classification, ie there is only one change, since we need predictions for each input tokken we do not use the Global Pooling Layer, therefore the architechture looks something like:
* Input Layer
* Embeddings
* Transformer Encoder Block
* Dropout (optional)
* Classification Layer (where n_units = number of classes, ie 2 in out case)
```
inputs = keras.Input(shape=(None, ), dtype="int64")
x = L.Embedding(config.VOCAB, config.EMBEDDING_VECTOR_LENGTH)(inputs)
x = TransformerEncoder(config.EMBEDDING_VECTOR_LENGTH, config.DENSE_DIM, config.NUM_HEADS)(x)
x = L.Dropout(0.5)(x)
outputs = L.Dense(config.N_OUPUT, activation=config.OUTPUT_ACTIVATION)(x)
model = keras.Model(inputs, outputs)
model.compile(loss=config.LOSS, optimizer=config.OPTIMIZER, metrics=config.METRICS)
model.summary()
plot_model(model, show_shapes=True)
history = model.fit(X_train, y_train, validation_data=(X_valid, y_valid),
callbacks=[es, rlp, csv_logger], epochs=config.MAX_EPOCHS
)
fig, ax = plt.subplots(2, 1, figsize=(20, 8))
df = pd.DataFrame(history.history)
df[['accuracy', 'val_accuracy']].plot(ax=ax[0])
df[['loss', 'val_loss']].plot(ax=ax[1])
ax[0].set_title('Model Accuracy', fontsize=12)
ax[1].set_title('Model Loss', fontsize=12)
fig.suptitle('Model Metrics', fontsize=18);
```
# Evaluation
```
i = np.random.randint(0, X_test.shape[0])
p = model.predict(np.array([X_test[i]]))
p = np.argmax(p, axis=-1)
y_true = np.argmax(y_test, axis=-1)[i]
print(f"{'Word':15}{'True':5}\t{'Pred'}")
print("-"*30)
for (w, t, pred) in zip(X_test[i], y_true, p[0]):
print(f"{words[w]:15}{t}\t{pred}")
if words[w] == 'ENDPAD':
break
```
| github_jupyter |
# Introduction to Probability and Statistics!
In this lab we will introduce you to some basic probability and statistics concepts. You may have heard of many of the terms we plan to go over today but if not, do not fret. Hopefully by the end of this session you will all leave here feeling like stats masters!
<br> <br> <br>
By the end of this section, hopefully you will be able to answer the following questions with confidence:
<br>
<br>
1) What is the mean, standard deviation, and variance of a collection of data?
<br>
2) How do these concepts help me interpret and better understand the data I'm given?
# 1.1 Mean, Variance, and Standard Deviation
TODO: data8 Lab 5 may be helpful here! Just got over the basic intuition of what the mean and std. dev. are, maybe with some plots. Tie in the connection to variance and how they together help to distinguish rare and expected values.
First off, you all may have heard of the mean value of a set of data, but if not, the mean is simply the average. Suppose you're given the following values:
$$ 5,3,7,4,2 $$
What is their average? We can compute this by hand:
$$ \frac{1}{5} \left( 5 + 3 + 7 + 4 + 2 \right) = \frac{21}{5} = 4.2$$
This is fine while we have a small set of values, but what if we heard thousands of values, or even hundreds of thousands? This is where computers can be very handy.
<br> <br>
Before we get into the coding part of the lesson, let's look at these values on a number line and get some intuition for what the mean value represents.
```
import matplotlib.pyplot as plt
plt.style.use('seaborn-whitegrid')
import numpy as np
fig = plt.figure()
ax = plt.axes()
x = np.array((5,3,7,4,2))
ax.scatter(x,np.zeros(len(x)), s=75, label = 'Values')
ax.scatter(4.2,0, s=75, label = 'Mean')
ax.legend()
```
Now let's generate some random numbers and calculate their means. I know this may seem tedious but the code will be useful later!
```
import random as rnd
for x in range(10):
print rnd.randint(1,101) # rnd.randint(start,end) returns an integer value between values start and end
for x in range(10):
print rnd.random() # rnd.random() returns a pseudo-random value between 0.0 and 1.0
# Fill in the following code to compute the mean of a set of randomly generated numbers
num_values = 2 # change to reflect number of values
current_total = 0.0 # variable to keep track of current total
for x in range(num_values):
val = rnd.random() # generate random number (either integer or decimal between 0 and 1) and store it in the variable named val
current_total = current_total + val # keep a running sum of the values
print val
print "The average value is", current_total/num_values
# If we'd like to plot our values and their corresponding means we would need to store the generated values.
# Rewrite the code above to store the values as they are generated. Instead of keeping a running sum, use
# numpy's "sum" function to calculate the mean after the loop!
# Code goes here
```
Using the provided code above, can you plot the generated values with their mean?
```
# Plot code goes here!
```
Now that we've got a better understanding for the mean, what about the standard deviation and variance?
<br> <br>
Intuitively, the standard deviation is a measure of spread of the data.
<br>
Suppose we have a 3-ft ruler that we'd like to measure a friend's height with. Let's also assume that the person of interest is taller than 3 feet and WE ONLY HAVE ONE RULER. Therefore, we need to be creative as to how we measure our friend. We could use a 3-ft long string as a placeholder and place the ruler tip-to-tail to measure the remaining distance. As you could imagine, our measurements would be slightly off. However, it's reasonable to think that if we took many measurements, that the mean measurement would be pretty close to the true height! In this case, the standard deviation would be a measure of the "spread" of all our (potentially) inaccurate measurements around the average (or mean) height.
<br> <br>
Using the standard deviation, we can get an idea for the interval for which a majority of our measurements lie. If we plotted our data on a number line as before and included the mean, we would see that 68% of our measurements fall within one standard deviation (on either side) of the mean!
<br> <br>
Finally, the variance is simply the square of the standard deviation!
$$ std \ dev = \sqrt{var} $$
```
fig = plt.figure()
ax = plt.axes()
x = np.array((5,3,7,4,2))
ax.scatter(x,np.zeros(len(x)), s=75, label = 'Values')
ax.scatter(4.2,0, s=75, label = 'Mean')
std = np.std(x)
x_ticks = np.append([1,3,4,5,7], [4.2-std,4.2+std])
plt.axvline(x=4.2-std,color='m')
plt.axvline(x=4.2+std,color='m')
ax.legend()
ax.set_xticks(x_ticks)
fig.suptitle('One Standard Deviation',fontsize=30)
fig = plt.figure()
ax = plt.axes()
x = np.array((5,3,7,4,2))
ax.scatter(x,np.zeros(len(x)), s=75, label = 'Values')
ax.scatter(4.2,0, s=75, label = 'Mean')
std = np.std(x)
x_ticks = np.append([1,3,4,5,7], [4.2-2*std,4.2+2*std])
plt.axvline(x=4.2-2*std,color='m')
plt.axvline(x=4.2+2*std,color='m')
ax.legend()
ax.set_xticks(x_ticks)
fig.suptitle('Two Standard Deviations',fontsize=30)
fig = plt.figure()
ax = plt.axes()
x = np.array((5,3,7,4,2))
ax.scatter(x,np.zeros(len(x)), s=75, label = 'Values')
ax.scatter(4.2,0, s=75, label = 'Mean')
std = np.std(x)
x_ticks = np.append([1,3,4,5,7], [4.2-3*std,4.2+3*std])
plt.axvline(x=4.2-3*std,color='m')
plt.axvline(x=4.2+3*std,color='m')
ax.legend()
ax.set_xticks(x_ticks)
fig.suptitle('Three Standard Deviations',fontsize=30)
from IPython.display import Image
Image(url = 'https://amp.businessinsider.com/images/546e68776bb3f74f68b7d0ba-750-544.jpg')
# Let's simulate this on our own!
num_draws = 10 # enter number of measurements to take
true_height, std_dev = 160, 1 # play around with changing the standard deviation. Do your results make sense?
height_msr = np.random.normal(true_height, std_dev,num_draws)
fig = plt.figure()
ax = plt.axes()
plt.hist(height_msr)
x_ticks = np.append(ax.get_xticks(), true_height)
plt.axvline(x=true_height,color='m')
ax.legend()
ax.set_xticks(x_ticks)
fig.suptitle('Simulated Height Measurements',fontsize=30)
plt.ylabel('Number of Measurements',fontsize = 15)
plt.xlabel('Measured Height (cm)',fontsize = 15)
```
# 1.2 Probability
Khan Academy says it the best: <br>
"Probability is simply how likely something is to happen. <br><br>
Whenever we're unsure about the outsome of an event, we can talk about the probabilities of certain outcomes - how likely they are. The analysis of events governed by probability is called statistics."
# 1.2.1 Flipping a Coin
The best example for understanding probability is flipping a coin.
There are two possible outcomes - heads or tails.
What's the probability of the coin landing on heads? We can find out using the equation $$P(H) = ? $$
You may already know the likelihood is 50% (assuming a fair coin). But how do we work that out?
$$ P(Event) = \frac{\text{number of ways it can happen}}{\text{total number of outcomes}} $$
In the case of flipping a coin, $P(H) = \frac{1}{2}$. <br><br>
Let's test this by simulation!
```
# In this cell we will write a block of code to simulate flipping a coin and plotting the results.
p = 0.5 # pick probability of getting heads
n = 10# pick number of trials
outcome = [] # empty container to store the results
for i in range(n):
result = rnd.random()# generate a number between 0 and 1 (Hint: we used a built-in function before that can help us with this)
if result <= p:
outcome.append(1)
else:
outcome.append(0)
fig = plt.figure()
plt.hist(outcome)
fig.suptitle('Flipping a Coin, Heads = 1, Tails = 0',fontsize=20)
```
# 1.2.2 Drawing cards from a Deck
```
Image(url='http://www.milefoot.com/math/discrete/counting/images/cards.png')
```
The primary deck of 52 playing cards in use today includes thirteen ranks of each of the four Franch suits, diamonds (♦), spades (♠), hearts (♥) and clubs (♣), with reversible Rouennais "court" or face cards. <br>
Each suit includes an ace, a king, a queen, a jack, and ranks two through ten, with each card depicting that many symbols of its suit.
<br>
Two (sometimes one or four) Jokers are included in commercial decks but many games require them to be removed before play. For simplicity, we will also assume there the Jokers of our deck have been removed.
<br>
Going back to our definition of probability
$$ P(Event) = \frac{\text{number of ways it can happen}}{\text{total number of outcomes}}, $$
what is the probability of drawing an ace?
<br> <br>
$$ P(ace) = \frac{\text{number of aces}}{\text{total playing cards}} = \frac{4}{52} = \frac{1}{13}. $$
Similarly, we could compute the probability of NOT drawing an ace
$$ P(not \ ace) = \frac{\text{number of non-ace cards}}{\text{total playing cards}} = \frac{48}{52} = \frac{12}{13}. $$
$$ P(not \ ace) = \frac{52}{52} - \frac{\text{number of aces}}{\text{total playing cards}} = \frac{48}{52} = \frac{12}{13}. $$
<br> <br>
Notice that $P(ace) + P(not \ ace) = 1$. That is, the probability of either drawing an ace from a deck of cards or not drawing an ace is 100%!
```
# The code below to takes a random draw from a deck of cards
# Define the special face cards, suits and color
faces = {11: "Jack", 12: "Queen", 13: "King", 1: "Ace"}
suits = {1: "Spades", 2: "Hearts", 3: "Diamonds", 4: "Clubs"}
colors = {"Spades" : "Black", "Clubs" : "Black", "Diamonds" : "Red", "Hearts" : "Red"}
# Draw a random integer value. Card_num can be any of the 13 cards to choose from and suits picks a random suit
card_num = rnd.randint(1,13)
suit_num = rnd.randint(1,4)
# Get the card face, suit and corresponding color
card_face = faces.get(card_num,card_num)
card_suit = suits[suit_num]
card_color = colors.get(card_suit)
print card_face, card_suit, card_color
num_draws = 10 # Can change number of draws but slow (on personal computer)
i = 0
cards = []
while i < num_draws:
# Draw a random integer value. Card_num can be any of the 13 cards to choose from and suits picks a random suit
card_num = rnd.randint(1,13)
suit_num = rnd.randint(1,4)
# Get the card face, suit and corresponding color
card_face = faces.get(card_num,card_num)
card_suit = suits[suit_num]
cards.append([card_face,card_suit,1])
i = i+1
# remove duplicates (assuming we are using only one deck)
''' for j in range(i-1):
if cards[j][0] == cards[i-1][0]:
if cards[j][1] == cards[i-1][1]:
cards.remove(cards[i-1])
i = i-1
'''
# Note: Won't remove duplicates so that we see probability of drawing a specific card
# Instead, count number of occurences
for j in range(i-1):
if cards[j][0] == cards[i-1][0]:
if cards[j][1] == cards[i-1][1]:
cards[j][2] = cards[j][2] + 1
cards.remove(cards[i-1])
i = i-1
print cards
```
| github_jupyter |
# Exploring RESTful APIs inside Notebooks
## Presentation at [PyData Berlin Meetup, 2018-10-17](https://www.meetup.com/PyData-Berlin/events/255361308/)
This notebook contains examples for how to explore RESTful APIs inside a [Jupyter](https://jupyter.org) notebook environment. It is somewhat inspired by [Postman](https://www.getpostman.com) and aims at providing more flexibility for customising the UI when entering input and rendering output.
## Ways of Working RESTfully…
Curl and friends
```
! curl https://xkcd.com/552/info.0.json
! curl -s https://xkcd.com/552/info.0.json | jq .
! http https://xkcd.com/552/info.0.json
! curl -s https://xkcd.com/552/info.0.json | jq .img | sed s/\"//g
! curl -s -k $(curl -s 'https://xkcd.com/552/info.0.json' | jq .img | sed s/\"//g) --output xkcd.png
! open xkcd.png
```
The following few examples require an HTTP API token, see https://wit.ai/docs/http/.
```
import os
WIT_TOKEN = os.getenv('WIT_TOKEN')
! curl -s -H 'Authorization: Bearer {WIT_TOKEN}' \
'https://api.wit.ai/message?q=silly+nonsense' | jq .
! curl -s -H 'Authorization: Bearer {WIT_TOKEN}' \
'https://api.wit.ai/message?'\
'q="how+is+the+traffic+around+office+in+chicago"' | jq .
! curl 'https://gist.githubusercontent.com/'\
'deeplook/71e9ded257cfc2d8e5e9/raw/f0cfbab5f266fcb8056e8aea046f1f222346b76b/2013.geojson'
```
Requests
```
import requests
requests.get('http://python.org').headers
```
Postman
<img src="images/postman.png" alt="Postman UI" style="width: 90%;"/>
## Enter ipyrest
```
from ipyrest import Api
Api('http://www.apple.com')
```
## Input Arguments & Parameters, etc.
This requires API access tokens as explained on https://developer.here.com/documentation.
```
import os
from ipyrest import Api
url = 'https://1.{maptype}.maps.api.here.com/' \
'maptile/2.1/{tiletype}/newest/{scheme}/{zoom}/{xtile}/{ytile}/{size}/{format}'
args = dict(
maptype='traffic',
tiletype='traffictile',
scheme='normal.day',
zoom='11',
xtile='525',
ytile='761',
size='256',
format='png8',
)
params = dict(
app_id=os.getenv('HEREMAPS_APP_ID'),
app_code=os.getenv('HEREMAPS_APP_CODE'),
ppi='320',
)
Api(url, args=args, params=params)
```
## GeoJSON Output
```
! curl 'https://gist.githubusercontent.com/deeplook/'\
'71e9ded257cfc2d8e5e9/raw/f0cfbab5f266fcb8056e8aea046f1f222346b76b/2013.geojson'
from ipyrest import Api
url = 'https://gist.githubusercontent.com/' \
'deeplook/71e9ded257cfc2d8e5e9/raw/f0cfbab5f266fcb8056e8aea046f1f222346b76b/2013.geojson'
def post(resp):
"Post-process response content-type since gists seem to use text/plain."
resp.headers['Content-Type'] = 'application/vnd.geo+json'
Api(url, post_process_resp=post)
```
## Simple custom rendering view
```
from ipywidgets import Textarea, Layout
from ipyrest import Api
from ipyrest.responseviews import ResponseView
class HelloWorldView(ResponseView):
name = 'HelloWorld'
mimetype_pats = ['text/html']
def render(self, resp):
layout = Layout(width='100%', height='100px')
return Textarea(value='Hello World!', layout=layout)
url = 'https://python.org'
Api(url, additional_views=[HelloWorldView])
```
## Advanced rendering view
This example requires API access tokens as explained on https://developer.here.com/documentation.
```
import os
from ipyleaflet import Map, Marker, Polyline
from ipyrest import Api
from ipyrest.responseviews import ResponseView, zoom_for_bbox
class HereIsolinesView(ResponseView):
"""
A view for the isolines from the HERE Routing API, see
https://developer.here.com/documentation/routing/topics/request-isoline.html.
"""
name = 'HereIsolines'
mimetype_pats = ['application/json']
def render(self, resp):
obj = resp.json()
center = obj['response']['center']
lat, lon = center['latitude'], center['longitude']
m = Map(center=(lat, lon))
m += Marker(location=(lat, lon))
mins, maxs = [], []
for isoline in obj['response']['isoline']:
shape = isoline['component'][0]['shape']
path = [tuple(map(float, pos.split(','))) for pos in shape]
m += Polyline(locations=path, color='red', weight=2, fill=True)
mins.append(min(path))
maxs.append(max(path))
m.zoom = zoom_for_bbox(*min(mins), *max(maxs))
self.data = m
return m
url = 'https://isoline.route.api.here.com' \
'/routing/7.2/calculateisoline.json'
lat, lon = 52.5, 13.4
params = dict(
app_id=os.getenv('HEREMAPS_APP_ID'),
app_code=os.getenv('HEREMAPS_APP_CODE'),
start=f'geo!{lat},{lon}',
mode='fastest;car;traffic:disabled',
rangetype='time', # time/distance
range='300,600', # seconds/meters
resolution='20', # meters
#departure='now', # 2018-07-04T17:00:00+02
)
Api(url, params=params, additional_views=[HereIsolinesView])
```
## 3D Output (Experimental)
This might have issues on JupyterLab, but a [classic notebook](http://localhost:8888/notebooks/pysdk/docs/postman/postmanbox-meetup.ipynb) is fine.
```
from ipyrest import Api
def post(resp):
"Post-proess response content-type since gists seem to have text/plain."
resp.headers['Content-Type'] = 'application/vnd.3d+txt'
url = 'https://gist.githubusercontent.com/deeplook/4568232f2ca9388942aab9830ceeb21f'\
'/raw/782da3be33080ff7c7d2bd25b7d96b6bb455d570/sample_xyz_1000.txt'
Api(url, post_process_resp=post)
```
As an aside, see a crude examples of ipyvolume and pptk using Lidar data in another notebook, [x_point_clouds.ipynb](x_point_clouds.ipynb).
## XKCD Variants
```
from ipyrest import Api
url = 'https://xkcd.com/552/info.0.json'
Api(url)
import requests
from ipyrest import Api
url = requests.get('https://xkcd.com/552/info.0.json').json()['img']
Api(url)
import requests
from ipywidgets import Image
from ipyrest import Api
from ipyrest.responseviews import ResponseView, builtin_view_classes
class XKCDView(ResponseView):
"Api rendering view for XKCD comics taken from XKCD JSON API."
name = 'XKCD'
mimetype_pats = ['application/json']
def render(self, resp):
return Image(value=requests.get(resp.json()['img']).content)
url = 'https://xkcd.com/552/info.0.json'
Api(url, views=builtin_view_classes + [XKCDView])
```
### What's the Latest Comic, BTW?
```
url = 'https://xkcd.com/info.0.json'
Api(url, views=builtin_view_classes + [XKCDView], click_send=True)
```
Compare with [xkcd.com](https://xkcd.com)...
## Dynamic Views
These examples require an API access token as explained on https://docs.gitlab.com/ce/user/profile/personal_access_tokens.html.
```
import os
TOKEN = os.environ['GITLAB_TOKEN']
server = 'https://gitlab.com/api/v4'
from ipyrest import Api, ResponseView
Api(f'{server}/snippets', headers={'PRIVATE-TOKEN': TOKEN},
cassette_path='snippets.yaml',
timeout=20)
from ipywidgets import HBox, VBox, Text, Button, Layout
from ipyrest import Api
from ipyrest.responseviews import ResponseView
class DynamicSnippetView(ResponseView):
"ResponseView showing snippet IDs with some decent 'UI'."
name = 'DynamicSnippetView'
mimetype_pats = ['application/json']
def render(self, resp):
return VBox([
HBox([Text(str(snippet['id'])),
Text(snippet['title']),
Button(description='Delete (dummy)')]
)
for snippet in resp.json()])
Api(f'{server}/snippets', headers={'PRIVATE-TOKEN': TOKEN},
cassette_path='snippets.yaml',
additional_views=[DynamicSnippetView], timeout=20)
```
## Skipped for Brevity
- Protobuf example
- Caching
- Timeouts
- HTTP methods other than GET
- Accessing data between response views
- Ipywidgets UI **executable w/o browser**
- Ipywidgets UI **testable via pytest**
- etc.
## Test-Suite
```
# Run local server for testing the API in a different process locally.
import multiprocessing
import subprocess
import requests
def start_test_server():
subprocess.check_call(['python', 'tests/api_server.py'])
url = 'http://localhost:5000/'
if requests.get(url).status_code >= 400:
print('Starting test server...')
multiprocessing.Process(target=start_test_server).start()
else:
print('Test server is already running.')
! pytest -s -v ../tests
```
## Ipyrest Under the Hood
### Architecture & Documentation
**"It's in a flux!"**
### Dependencies
- **requests**
- **ipywidgets**
- **timeout_decorator**
- **typing**
- ipyleaflet
- ipyvolume
- pandas
- pytest
- ...
### To-do-to-do-to-do
- make *true* widget
- version
- package
- docker
- binder
- swagger?
## Wrap-Up
- http://github.com/deeplook/ipyrest
- Ipyrest is an _emerging_ tool for _exploring_ APIs inside _notebooks_.
- Best used for _testing_ new APIs _quickly_ and _interactively_.
- It _might_ be also useful for providing *executable API examples* online.
- It is **#WIP #Alpha #Prototype**!
- But already useful!
- Try it!
- Contribute!
- provide ResponseViews for your use case
- provide examples using other APIs
- push for missing ipywidgets issues like [this on styling](https://github.com/jupyter-widgets/ipywidgets/issues/2206)
## Q & A & T
- Questions?
- Answers!
- Thank You!
| github_jupyter |
**Chapter 03**
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
%matplotlib inline
```
# 8 Auto Data set (Linear Regression)
```
auto_file_path = '../data/Auto'
autos = pd.read_table(auto_file_path,sep='\s+')
autos.head()
# clearn the data
autos=autos.replace('?',np.NAN).dropna()
autos['horsepower']=autos['horsepower'].astype('float')
autos.plot.scatter(x='horsepower', y='mpg')
g = sns.jointplot('horsepower','mpg',data=autos,kind='reg', xlim=(25,225))
from IPython.display import HTML, display
import statsmodels.api as sm
from statsmodels.formula.api import ols
mpg_model = ols("mpg ~ horsepower", autos).fit()
mpg_model_summary=mpg_model.summary()
# convert our table to HTML and add colors to headers for explanatory purposes
HTML(
mpg_model_summary\
.as_html()\
.replace(' Adj. R-squared: ', ' Adj. R-squared: ')\
.replace('coef', 'coef')\
.replace('std err', 'std err')\
.replace('P>|t|', 'P>|t|')\
.replace('[95.0% Conf. Int.]', '[95.0% Conf. Int.]')
)
# confidence interval
from statsmodels.sandbox.regression.predstd import wls_prediction_std
_, confidence_interval_lower, confidence_interval_upper = wls_prediction_std(mpg_model)
x = autos['horsepower']
y = autos['mpg']
fig, ax = plt.subplots(figsize=(10,7))
ax.plot(x, confidence_interval_lower,'g--')
ax.plot(x, confidence_interval_upper,'g--')
ax.scatter(x,y)
sns.lmplot(x='horsepower', y='mpg',data=autos)
sns.residplot(x='horsepower', y='mpg',data=autos)
```
# 2 Auto Data set (Multiple Linear Regression)
```
from pandas.tools.plotting import scatter_matrix
# mpg cylinders displacement horsepower weight acceleration
fig, ax = plt.subplots(figsize=(15, 15))
df_auto = autos[['mpg','cylinders','displacement','horsepower','weight','acceleration','year','origin']]
scatter_matrix(df_auto, alpha=0.5,diagonal='kde', ax=ax);
df_auto.corr()
mpg_multi_model = ols('mpg ~ cylinders + displacement + horsepower + weight + acceleration + year + origin',
data=df_auto).fit()
mpg_multi_model_summary = mpg_multi_model.summary()
HTML(
mpg_multi_model_summary\
.as_html()\
.replace(' Adj. R-squared: ', ' Adj. R-squared: ')\
.replace('coef', 'coef')\
.replace('std err', 'std err')\
.replace('P>|t|', 'P>|t|')\
.replace('[95.0% Conf. Int.]', '[95.0% Conf. Int.]')
)
```
# 10 Carseats Data set
```
carseats_file_name = '../data/Carseats.csv'
carseats = pd.read_csv(carseats_file_name, index_col=0)
carseats.head()
carseats_subset = carseats[['Sales','Price','Urban','US']]
carseats_subset=carseats_subset.replace(['Yes','No'],[1,-1])
sales_multi_model = ols('Sales ~ Price + Urban + US', data=carseats_subset).fit()
sales_multi_model_summary = sales_multi_model.summary()
HTML(
sales_multi_model_summary\
.as_html()\
.replace(' Adj. R-squared: ', ' Adj. R-squared: ')\
.replace('coef', 'coef')\
.replace('std err', 'std err')\
.replace('P>|t|', 'P>|t|')\
.replace('[95.0% Conf. Int.]', '[95.0% Conf. Int.]')
)
```
$$
y_i = \beta_0+\beta_1\times price + \beta_2 \times urban + \beta_3\times us+\epsilon_i =
\begin{cases}
\beta_0+\beta_1\times price + \beta_2 + \beta_3 + \epsilon_i & i \text{th carseat is Urban is Yes and US is Yes} \\
\beta_0+\beta_1\times price - \beta_2 + \beta_3 + \epsilon_i & i \text{th carseat is Urban is No and US is Yes} \\
\beta_0+\beta_1\times price + \beta_2 - \beta_3 + \epsilon_i & i \text{th carseat is Urban is Yes and US is No} \\
\beta_0+\beta_1\times price - \beta_2 - \beta_3 + \epsilon_i & i \text{th carseat is Urban is No and US is No} \\
\end{cases}
$$
The coefficients' null hypotheis of Intercept, Price and US can be rejected.
```
sales_multi_model = ols('Sales ~ Price + US', data=carseats_subset).fit()
sales_multi_model_summary = sales_multi_model.summary()
HTML(
sales_multi_model_summary\
.as_html()\
.replace(' Adj. R-squared: ', ' Adj. R-squared: ')\
.replace('coef', 'coef')\
.replace('std err', 'std err')\
.replace('P>|t|', 'P>|t|')\
.replace('[95.0% Conf. Int.]', '[95.0% Conf. Int.]')
)
from statsmodels.graphics.regressionplots import plot_leverage_resid2
plot_leverage_resid2(sales_multi_model);
fig, ax = plt.subplots(figsize=(10,10))
fig=sm.graphics.influence_plot(sales_multi_model, ax=ax, criterion="cooks")
```
# 11
```
np.random.seed(1)
x = np.random.normal(0,1,100)
y = 2*x + np.random.normal(0, 1, 100)
```
## 11(a)
```
df = pd.DataFrame({'x':x,'y':y})
df_y_x_model = ols('y ~ x + 0', data=df).fit()
df_y_x_model_summary = df_y_x_model.summary()
HTML(
df_y_x_model_summary\
.as_html()\
.replace(' Adj. R-squared: ', ' Adj. R-squared: ')\
.replace('coef', 'coef')\
.replace('std err', 'std err')\
.replace('P>|t|', 'P>|t|')\
.replace('[95.0% Conf. Int.]', '[95.0% Conf. Int.]')
)
```
## 11(b)
```
df_x_y_model = ols('x ~ y + 0', data=df).fit()
df_x_y_model_summary = df_x_y_model.summary()
HTML(
df_x_y_model_summary\
.as_html()\
.replace(' Adj. R-squared: ', ' Adj. R-squared: ')\
.replace('coef', 'coef')\
.replace('std err', 'std err')\
.replace('P>|t|', 'P>|t|')\
.replace('[95.0% Conf. Int.]', '[95.0% Conf. Int.]')
)
```
## 11(c)
The $y=2x+\epsilon$ can be written $x=0.5(y-\epsilon)$
## 11(d)
We draw $x$ from the Gaussian normal distribution, so the $\bar{x}=0$, and the $\bar{y}=0$. $\hat{\beta}=\frac{\sum_{i=1}^n(x_i-\bar{x})(y_i-\bar{y})}{\sum_{i=1}^{n}(x_i-\bar{x})^2}=\frac{\sum_{i=1}^nx_iy_i}{\sum_{i=1}^{n}x_i^2}$
Here is the Proof:
$$
t=\frac{\hat{\beta}}{\text{SE}(\hat{\beta})}=\frac{\sum x_iy_i}{\sum x_i^2}\sqrt{\frac{(n-1)\sum x_i^2}{\sum (y_i-x_i\hat{\beta})^2}}
$$
$$
=\frac{\sqrt{n-1}\sum x_iy_i}{\sqrt{\sum x_i^2\sum (y_i-x_i\hat{\beta})^2}} = \frac{\sqrt{n-1}\sum x_iy_i}{\sqrt{\sum x_i^2\sum (y_i^2-2\hat{\beta}x_iy_i+x_i^2\hat{\beta}^2)}}
$$
$$
=\frac{\sqrt{n-1}\sum x_iy_i}{\sqrt{\sum x_i^2 \sum y_i^2 - \sum x_i^2\hat{\beta}(2\sum x_iy_i - \hat{\beta}\sum x_i^2)}} = \frac{\sqrt{n-1}\sum x_iy_i}{\sqrt{\sum x_i^2 \sum y_i^2 - \sum x_iy_i(2\sum x_iy_i - \sum x_iy_i)}}
$$
$$
t=\frac{\sqrt{n-1}\sum x_iy_i}{\sqrt{\sum x_i^2 \sum y_i^2-(\sum x_iy_i)^2}}
$$
## 11(f)
```
df_y_x_intercept_model = ols('y ~ x', data=df).fit()
df_y_x_intercept_model_summary = df_y_x_intercept_model.summary()
HTML(
df_y_x_intercept_model_summary\
.as_html()\
.replace(' Adj. R-squared: ', ' Adj. R-squared: ')\
.replace('coef', 'coef')\
.replace('std err', 'std err')\
.replace('P>|t|', 'P>|t|')\
.replace('[95.0% Conf. Int.]', '[95.0% Conf. Int.]')
)
df_x_y_intercept_model = ols('x ~ y', data=df).fit()
df_x_y_intercept_model_summary = df_x_y_intercept_model.summary()
HTML(
df_x_y_intercept_model_summary\
.as_html()\
.replace(' Adj. R-squared: ', ' Adj. R-squared: ')\
.replace('coef', 'coef')\
.replace('std err', 'std err')\
.replace('P>|t|', 'P>|t|')\
.replace('[95.0% Conf. Int.]', '[95.0% Conf. Int.]')
)
```
# 13
## 13(a)
```
np.random.seed(1)
x = np.random.normal(0,1,100)
```
## 13(b)
```
eps = np.random.normal(0,0.25,100)
```
## 13(c)
```
y = -1.0 + 0.5*x+eps
df = pd.DataFrame({'x':x,'y':y})
```
## 13(d)
```
df.plot.scatter(x='x',y='y');
lm_model = ols('y ~ x', data=df).fit()
lm_model_summary = lm_model.summary()
HTML(
lm_model_summary\
.as_html()\
.replace(' Adj. R-squared: ', ' Adj. R-squared: ')\
.replace('coef', 'coef')\
.replace('std err', 'std err')\
.replace('P>|t|', 'P>|t|')\
.replace('[95.0% Conf. Int.]', '[95.0% Conf. Int.]')
)
```
## 11(e)
```
X = np.linspace(-2,2,100)
y_pred = X*0.5239 + (-0.9632)
y_actu = X*0.5 + (-1.0)
plt.plot(X, y_pred, 'r', label='Predict')
plt.plot(X, y_actu, 'b', label='Actual')
plt.scatter(x,y)
plt.legend()
plt.show()
```
## 13(g)
```
df['x2'] = x**2
lm_quadratic_model = ols('y ~ x + x2', data= df).fit()
lm_quadratic_model_summary = lm_quadratic_model.summary()
HTML(
lm_quadratic_model_summary\
.as_html()\
.replace(' Adj. R-squared: ', ' Adj. R-squared: ')\
.replace('coef', 'coef')\
.replace('std err', 'std err')\
.replace('P>|t|', 'P>|t|')\
.replace('[95.0% Conf. Int.]', '[95.0% Conf. Int.]')
)
```
As the $x^2$'s $p-$value is big enough to reject this relationship
# 14
## 14(a)
```
np.random.seed(1)
x1 = np.random.uniform(0,1,100)
x2 = 0.5*x1+ np.random.normal(100)/10
y = 2 + 2*x1 + 0.3*x2+np.random.normal(100)
```
$$
y=2 + 2\times x_1 + 0.3\times x_2
$$
## 14(b)
```
df = pd.DataFrame({'x1':x1,'x2':x2,'y':y})
df[['x1','x2']].corr()
df[['x1','x2']].plot.scatter(x='x1',y='x2');
```
## 14(c)
```
lm_y_x1_x2_model = ols('y ~ x1 + x2', data=df).fit()
lm_y_x1_x2_model_summary = lm_y_x1_x2_model.summary()
HTML(
lm_y_x1_x2_model_summary\
.as_html()\
.replace(' Adj. R-squared: ', ' Adj. R-squared: ')\
.replace('coef', 'coef')\
.replace('std err', 'std err')\
.replace('P>|t|', 'P>|t|')\
.replace('[95.0% Conf. Int.]', '[95.0% Conf. Int.]')
)
```
From above tables, we get the $\hat{\beta_0}=1.577,\hat{\beta_1}=-2.9255,\hat{\beta_2}=10.15009$ which are quite various from the $\beta_0,\beta_1,\beta_2$
## 14(d)
```
lm_y_x1_model = ols('y ~ x1', data=df).fit()
lm_y_x1_model_summary = lm_y_x1_model.summary()
HTML(
lm_y_x1_model_summary\
.as_html()\
.replace(' Adj. R-squared: ', ' Adj. R-squared: ')\
.replace('coef', 'coef')\
.replace('std err', 'std err')\
.replace('P>|t|', 'P>|t|')\
.replace('[95.0% Conf. Int.]', '[95.0% Conf. Int.]')
)
```
## 14(e)
```
lm_y_x2_model = ols('y ~ x2', data=df).fit()
lm_y_x2_model_summary = lm_y_x2_model.summary()
HTML(
lm_y_x2_model_summary\
.as_html()\
.replace(' Adj. R-squared: ', ' Adj. R-squared: ')\
.replace('coef', 'coef')\
.replace('std err', 'std err')\
.replace('P>|t|', 'P>|t|')\
.replace('[95.0% Conf. Int.]', '[95.0% Conf. Int.]')
)
```
## 14(f)
Because the $x_1$ and $x_2$ have colineartiy, it is hard to distinguish their effects.
```
x1 = np.hstack((x1,[0.6]))
x2 = np.hstack([x2,[0.8]])
y = np.hstack([y, 6])
df = pd.DataFrame({'x1':x1,'x2':x2,'y':y})
lm_y_x1_x2_model = ols('y ~ x1 + x2',data=df).fit()
lm_y_x1_x2_model_summary = lm_y_x1_x2_model.summary()
HTML(
lm_y_x1_x2_model_summary\
.as_html()\
.replace(' Adj. R-squared: ', ' Adj. R-squared: ')\
.replace('coef', 'coef')\
.replace('std err', 'std err')\
.replace('P>|t|', 'P>|t|')\
.replace('[95.0% Conf. Int.]', '[95.0% Conf. Int.]')
)
from statsmodels.graphics.regressionplots import plot_leverage_resid2
plot_leverage_resid2(lm_y_x1_x2_model);
```
# 15
```
boston_file_name = '../data/Boston.csv'
bostons = pd.read_csv(boston_file_name, index_col=0)
bostons.head()
```
## 15(a)
```
print(ols('crim ~ zn',data=bostons).fit().summary())
print(ols('crim ~ indus',data=bostons).fit().summary())
print(ols('crim ~ chas',data=bostons).fit().summary())
print(ols('crim ~ nox',data=bostons).fit().summary())
print(ols('crim ~ rm',data=bostons).fit().summary())
print(ols('crim ~ age',data=bostons).fit().summary())
print(ols('crim ~ dis',data=bostons).fit().summary())
print(ols('crim ~ rad',data=bostons).fit().summary())
print(ols('crim ~ tax',data=bostons).fit().summary())
print(ols('crim ~ ptratio',data=bostons).fit().summary())
print(ols('crim ~ black',data=bostons).fit().summary())
print(ols('crim ~ lstat',data=bostons).fit().summary())
print(ols('crim ~ medv',data=bostons).fit().summary())
```
## 15(b)
```
print(ols('crim ~ zn + indus + chas + nox + rm + age + dis + rad + tax + ptratio + black + lstat + medv',
data=bostons).fit().summary())
```
| github_jupyter |
# Error and Exception Handling
```
5/0
print("hellow")
a =int(input("Enter a number"))
b =int(input("Enter a number"))
try:
c = a/b
print(c)
except ZeroDivisionError:
print("Exception is caught ZeroDivisionError")
print('Hello Error')
a =int(input("Enter a number"))
b =int(input("Enter a number"))
c = a/b
print(c)
print('Hello World')
a = [1,2,3]
try:
print(a[8])
except IndexError:
print('Yeah! I Have handled that error')
a =int(input("Enter a number"))
b =int(input("Enter a number"))
d = []
try:
c = a/b
print(c)
print(d[2])
except ZeroDivisionError:
print('Exception is caught ZeroDivisionError')
print('Hello World')
a =int(input("Enter a number"))
b =int(input("Enter a number"))
try:
c = a/b
except ZeroDivisionError:
print("Handled ZeroDivisionError")
else:
print("Else " +str(c))
a =int(input("Enter a number"))
b =int(input("Enter a number"))
d = []
try:
c = a/b
e = d[5]
print(e)
except ZeroDivisionError:
print("Handled ZeroDivisionError")
else:
print(c)
print("First")
a =int(input("Enter a number"))
b =int(input("Enter a number"))
d = []
try:
c = a/b
# e = d[5]
except ZeroDivisionError:
print("Handled ZeroDivisionError")
else:
print(c)
print("last")
a =int(input("Enter a number"))
b =int(input("Enter a number"))
d = []
try:
c = a/b
e = d[5]
except ZeroDivisionError: #Type Z and press tab to get list of Exception
print("Exception is caught ZeroDivisionError")
except IndexError: #Type I and press tab to get list of Exception
print("Exception is caught is list index out of range")
else:
print(c)
print(e)
d = []
try:
a =int(input("Enter a number"))
b =int(input("Enter a number"))
c = a/b
e = d[5]
except: # here we did not explain the type of exception. Genral exception will
# be used. means it will catch every kind of exception
print("koi si bhi Exception is caught")
else:
print(c)
# a =int(input("Enter a number"))
# b =int(input("Enter a number"))
d = []
try:
'''built in message print karna hy exception ka to {except Exception as e:}
is me jistrah ki acception aegi wo uska message khud he utha k laega or print karega
is code ko khud se run kare or exception raise kar k dekhen'''
a =int(input("Enter a number"))
b =int(input("Enter a number"))
c = a/b
ee = d[5]
except Exception as e:
print("Handled Error == " +str(e))
else:
print(c)
print('HelloWorld')
a =int(input("Enter a number"))
b =int(input("Enter a number"))
d = []
try:
c = a/b
e = d[5]
"""yahan ek humne apni message likha hy or ek built in use kia hy"""
except ZeroDivisionError:
print("Handel ZeroDivisionError")
except Exception as e:
print("Handle Error == " +str(e))
else:
print(c)
a =int(input("Enter a number"))
b =int(input("Enter a number"))
d = []
try:
c = a/b
e = d[5]
except ZeroDivisionError:
print("Handel ZeroDivisionError")
else:
print(c)
finally:
print("Finally will always run")
```
```
class Student():
def __init__(self,name,age):
if age > 80 or age <16:
raise Exception("Age can not be greater then 80 and less then 16")
self.name = name
self.age = age
age = int(input("Enter students age: "))
st = Student("ALi",age)
print(st.age)
```
```
class Student():
def __init__(self,name,age):
if age > 80:
raise Exception("Age can not be greater then 80")
self.name = name
self.age = age
'''ab humen try except laga diya hy code crash nahi hoga or izzat batch jaegi hahahahah'''
try:
age = int(input("Enter students age: "))
st = Student("Hello",age) # send different ages and see the result
except Exception as e:
print("Exception "+str(e))
else :
print(st.age)
```
```
class Student():
def __init__(self,name,age):
if age > 80 or age <16:
raise StudentAgeException("Age can not be greater then 80 and less then 16")
# we used here own class of exceptio
self.name = name
self.age = age
'''All the exception raised are derived from a class Exception by default
We can make our own class of exception and can derived(inherit) it from parent
Exception class like this:'''
class StudentAgeException(Exception):
pass
age = int(input("Enter students age: "))
st = Student("Hello",age)
# abhi it is creating exception that is not handled
print(st.age)
####Note###
'''exception raise karane k liye humne Exception python ki general wali use nahiiiiiii ki hy
abhi code crash hogaya hy kyu k humne abhi exception catch nahiiiiiiiiiiiiiiiiii ki hy'''
```
```
class Student():
def __init__(self,name,age):
if age > 80 or age <16:
raise StudentAgeException("Age can not be greater then 80 and less then 16")
self.name = name
self.age = age
class StudentAgeException(Exception):
pass
try:
age = int(input("Enter students age: "))
st = Student("Hello",age)
except StudentAgeException as e:
print("StudentAgeException == "+str(e))
else :
print(st.age)
```
| github_jupyter |
# Preprocessing and Feature Engineering
One of the biggest challenges and perhaps most time consuming step in any advanced analytics project is preprocessing. In order to obtain a meaningful result from a machine learning algorithm a thorough preprocessing is required. It’s not that it’s particularly complicated programming, but rather that it requires deep knowledge of the data you are working with and an understanding of what your model needs in order to successfully leverage this data.
Feature engineering is the process of using domain knowledge of the data to create features that make machine learning algorithms work. Feature engineering is fundamental to the application of machine learning, and is both difficult and expensive.
## Formatting Models According to Your Use Case
To preprocess data for Spark’s different advanced analytics tools, you must consider your end objective. The following list walks through the requirements for input data structure for each advanced analytics task in MLlib:
* In the case of most **classification** and **regression** algorithms, you want to get your data into a column of type Double to represent the label and a column of type Vector (either dense or sparse) to represent the features.
* In the case of **recommendation**, you want to get your data into a column of users, a column of items (say movies or books), and a column of ratings.
* In the case of **unsupervised learning**, a column of type Vector (either dense or sparse) is needed to represent the features.
Before we proceed, we’re going to read in several different sample datasets, each of which has different properties we will manipulate:
```
# the following line gets the bucket name attached to our cluster
bucket = spark._jsc.hadoopConfiguration().get("fs.gs.system.bucket")
# specifying the path to our bucket where the data is located (no need to edit this path anymore)
data = "gs://" + bucket + "/notebooks/jupyter/data/"
print(data)
sales = spark.read.format("csv")\
.option("header", "true")\
.option("inferSchema", "true")\
.load(data + "retail-data/by-day/*.csv")\
.coalesce(5)\
.where("Description IS NOT NULL")
fakeIntDF = spark.read.parquet(data + "simple-ml-integers")
simpleDF = spark.read.json(data + "simple-ml")
scaleDF = spark.read.parquet(data + "simple-ml-scaling")
```
In addition to this realistic *sales* data, we’re going to use several simple synthetic datasets as well. *FakeIntDF*, *simpleDF*, and *scaleDF* all have very few rows. This will give you the ability to focus on the exact data manipulation we are performing instead of the various inconsistencies of any particular dataset. Because we’re going to be accessing the sales data a number of times, we’re going to cache it so we can read it efficiently from memory as opposed to reading it from disk every time we need it. Let’s also check out the first several rows of data in order to better understand what’s in the dataset:
```
sales.cache()
sales.show(3)
print("sales datasets consists of {} rows.".format(sales.count()))
```
**Note:** It is important to note that we filtered out null values here. MLlib does not always play nicely with null values at this point in time. This is a frequent cause for problems and errors and a great first step when you are debugging. Improvements are also made with every Spark release to improve algorithm handling of null values.
## Transformers
The best way to get your data in the desired format is through **transformers**. Transformers are functions that accept a DataFrame as an argument and return a new DataFrame as a response. This notebook will focus on what transformers are relevant for particular use cases rather than attempting to enumerate every possible transformer.
Spark provides a number of transformers that can be found under *pyspark.ml.feature*. For more information on transformers you can visit [Extracting, transforming and selecting features doccumentation page](http://spark.apache.org/docs/latest/ml-features.html).
Transformers are functions that convert raw data in some way. This might be to create a new interaction variable (from two other variables), to normalize a column, or to simply turn it into a Double to be input into a model. Transformers are primarily used in preprocessing or feature generation. RFormula that we saw in the previous lecture is an example of a transformer.
Spark’s transformer only includes a transform method. This is because it will not change based on the input data. Figure below is a simple illustration. On the left is an input DataFrame with the column to be manipulated. On the right is the input DataFrame with a new column representing the output transformation.
<img src="https://github.com/soltaniehha/Big-Data-Analytics-for-Business/blob/master/figs/10-01-A-Spark-transformer.png?raw=true" width="800" align="center"/>
The **`Tokenizer`** is an example of a transformer. It tokenizes a string (converts the input string to lowercase and then splits it by white spaces) and has nothing to learn from our data; it simply applies a function. Here’s a small code snippet showing how a tokenizer is built to accept the input column, how it transforms the data, and then the output from that transformation:
```
from pyspark.ml.feature import Tokenizer
tokenizer = Tokenizer(inputCol="Description", outputCol="words")
tokenized = tokenizer.transform(sales).select("Description", "words")
tokenized.show(5, False)
```
We can simply create a "udf" function and apply it on our newly created column to count the number of words:
```
from pyspark.sql import functions as F
countTokens = F.udf(lambda words: len(words))
tokenized.select("Description", "words")\
.withColumn("tokens", countTokens(F.col("words"))).show(5, False)
```
## Estimators for Preprocessing
Another set of tools for preprocessing are estimators. An estimator is necessary when a transformation you would like to perform must be initialized with data or information about the input column (often derived by doing a pass over the input column itself). For example, if you wanted to scale the values in our column to have mean zero and unit variance, you would need to perform a pass over the entire data in order to calculate the values you would use to normalize the data to mean zero and unit variance. In effect, an estimator can be a transformer configured according to your particular input data. In simplest terms, you can either blindly apply a transformation (a “regular” transformer type) or perform a transformation based on your data (an estimator type). Figure below is a simple illustration of an estimator fitting to a particular input dataset, generating a transformer that is then applied to the input dataset to append a new column (of the transformed data).
<img src="https://github.com/soltaniehha/Big-Data-Analytics-for-Business/blob/master/figs/10-01-A-Spark-estimator.png?raw=true" width="800" align="center"/>
An example of this type of estimator is the **`StandardScaler`**, which scales your input column according to the range of values in that column to have a zero mean and a variance of 1 in each dimension. For that reason it must first perform a pass over the data to create the transformer. Here’s a sample code snippet showing the entire process, as well as the output:
```
scaleDF.show(2)
from pyspark.ml.feature import StandardScaler
ss = StandardScaler(inputCol="features", outputCol="scaledFeatures")
FittedSs = ss.fit(scaleDF)
print("mean:", FittedSs.mean)
print("std:", FittedSs.std)
FittedSs.transform(scaleDF).show(5, False)
```
**Note:** that we have used the same DF to fit and to transform our data, but in reality one could use a different DF to obtain the parameters needed to scale (e.g. on the training set) and then apply it to other DFs (test, validation).
### Transformer Properties
All transformers require you to specify, at a minimum, the inputCol and the outputCol, which represent the column name of the input and output, respectively. There are some defaults (you can find these in the documentation), but it is a best practice to manually specify them yourself for clarity. In addition to input and output columns, all transformers have different parameters that you can tune (whenever we mention a parameter in this chapter you must set it with a set() method). In Python, we also have another method to set these values with keyword arguments to the object’s constructor. Estimators require you to fit the transformer to your particular dataset and then call transform on the resulting object.
## High-Level Transformers
High-level transformers, such as the RFormula we saw in the previous lecture, allow you to concisely specify a number of transformations in one. These operate at a “high level”, and allow you to avoid doing data manipulations or transformations one by one. In general, **you should try to use the highest level transformers you can, in order to minimize the risk of error and help you focus on the business problem instead of the smaller details of implementation**. While this is not always possible, it’s a good objective.
### RFormula
The **`RFormula`** is the easiest transfomer to use when you have “conventionally” formatted data. Spark borrows this transformer from the R language to make it simple to declaratively specify a set of transformations for your data. With this transformer, values can be either numerical or categorical and you do not need to extract values from strings or manipulate them in any way. The RFormula will automatically handle categorical inputs (specified as strings) by performing something called one-hot encoding. In brief, one-hot encoding converts a set of values into a set of binary columns specifying whether or not the data point has each particular value (we’ll discuss one-hot encoding in more depth later in the notebook). With the RFormula, numeric columns will be cast to Double but will not be one-hot encoded. If the label column is of type String, it will be first transformed to Double with StringIndexer.
The RFormula allows you to specify your transformations in declarative syntax. It is simple to use once you understand the syntax. Currently, RFormula supports a limited subset of the R operators that in practice work quite well for simple transformations. The basic operators are:
~
Separate target and terms
+
Concat terms; “+ 0” means removing the intercept (this means that the y-intercept of the line that we will fit will be 0)
-
Remove a term; “- 1” means removing the intercept (this means that the y-intercept of the line that we will fit will be 0—yes, this does the same thing as “+ 0”
:
Interaction (multiplication for numeric values, or binarized categorical values)
.
All columns except the target/dependent variable
RFormula also uses default columns of label and features to label, you guessed it, the label and the set of features that it outputs (for supervised machine learning). The models covered later on in this chapter by default require those column names, making it easy to pass the resulting transformed DataFrame into a model for training.
Let’s use RFormula in an example. In this case, we want to use all available variables (the .) and then specify an interaction between value1 and color and value2 and color as additional features to generate:
```
from pyspark.ml.feature import RFormula
supervised = RFormula(formula="lab ~ . + color:value1 + color:value2")
supervised.fit(simpleDF).transform(simpleDF).show(2, False)
```
### SQL Transformers
A **`SQLTransformer`** allows you to leverage Spark’s vast library of SQL-related manipulations just as you would a MLlib transformation. Any SELECT statement you can use in SQL is a valid transformation. The only thing you need to change is that instead of using the table name, you should just use the keyword \__THIS__. You might want to use SQLTransformer if you want to formally codify some DataFrame manipulation as a preprocessing step, or try different SQL expressions for features during hyperparameter tuning. Also note that the output of this transformation will be appended as a column to the output DataFrame.
You might want to use an SQLTransformer in order to represent all of your manipulations on the very rawest form of your data so you can version different variations of manipulations as transformers. This gives you the benefit of building and testing varying pipelines, all by simply swapping out transformers. The following is a basic example of using SQLTransformer:
```
from pyspark.ml.feature import SQLTransformer
basicTransformation = SQLTransformer()\
.setStatement("""
SELECT CustomerID, sum(Quantity) total_quantity, count(*) number_of_transactions
FROM __THIS__
GROUP BY CustomerID
""")
basicTransformation.transform(sales).show(5)
```
### VectorAssembler
The **`VectorAssembler`** is a tool you’ll use in nearly every single pipeline you generate. It helps concatenate all your features into one big vector you can then pass into an estimator. It’s used typically in the last step of a machine learning pipeline and takes as input a number of columns of Boolean, Double, or Vector. This is particularly helpful if you’re going to perform a number of manipulations using a variety of transformers and need to gather all of those results together.
Its output is similar to an RFormula transformation without the label column and any particular transformation based on a formula.
The output from the following code snippet will make it clear how this works:
```
from pyspark.ml.feature import VectorAssembler
va = VectorAssembler().setInputCols(["int1", "int2", "int3"])
va.transform(fakeIntDF).show()
```
## Working with Continuous Features
Continuous features are just values on the number line, from positive infinity to negative infinity. There are two common transformers for continuous features. First, you can convert continuous features into categorical features via a process called bucketing, or you can scale and normalize your features according to several different requirements. These transformers will only work on Double types, so make sure you’ve turned any other numerical values to Double:
```
contDF = spark.range(20).selectExpr("cast(id as double)")
contDF.show(2)
```
### Bucketing
The most straightforward approach to *bucketing* or *binning* is using the **`Bucketizer`**. This will split a given continuous feature into the buckets of your designation. You specify how buckets should be created via an array or list of Double values. This is useful because you may want to simplify the features in your dataset or simplify their representations for interpretation later on. For example, imagine you have a column that represents a patient’s BMI and you would like to predict some value based on this information. In some cases, it might be simpler to create three buckets of “overweight,” “average,” and “underweight.”
To specify the bucket, set its borders. For example, setting splits to 5.0, 10.0, 250.0 on our contDF will actually fail because we don’t cover all possible input ranges. When specifying your bucket points, the values you pass into splits must satisfy three requirements:
* The minimum value in your splits array must be less than the minimum value in your DataFrame.
* The maximum value in your splits array must be greater than the maximum value in your DataFrame.
* You need to specify at a minimum three values in the splits array, which creates two buckets.
To cover all possible ranges we can use `float("inf")` and `float("-inf")`.
In order to handle null or NaN values, we must specify the `handleInvalid` parameter as a certain value. We can either keep those values (keep), error or null, or skip those rows. Here’s an example of using bucketing:
```
from pyspark.ml.feature import Bucketizer
bucketBorders = [-1.0, 5.0, 10.0, 250.0, 600.0]
bucketer = Bucketizer(inputCol="id", outputCol="id_bucket", splits = bucketBorders)
bucketer.transform(contDF).show()
```
In addition to splitting based on hardcoded values, another option is to split based on percentiles in our data. This is done with **`QuantileDiscretizer`**, which will bucket the values into user-specified buckets with the splits being determined by approximate quantiles values. For instance, the 90th quantile is the point in your data at which 90% of the data is below that value. You can control how finely the buckets should be split by setting the relative error for the approximate quantiles calculation using `setRelativeError`. Spark does this is by allowing you to specify the number of buckets you would like out of the data and it will split up your data accordingly. The following is an example:
```
from pyspark.ml.feature import QuantileDiscretizer
bucketer = QuantileDiscretizer(numBuckets=5, inputCol="id", outputCol="id_bucket")
fittedBucketer = bucketer.fit(contDF)
fittedBucketer.transform(contDF).show()
```
**NOTE:** If the user chooses to keep NaN values, they will be handled specially and placed into their own bucket, for example, if 4 buckets are used, then non-NaN data will be put into buckets[0-3], but NaNs will be counted in a special bucket[4].
### Scaling and Normalization
We saw how we can use bucketing to create groups out of continuous variables. Another common task is to scale and normalize continuous data. While not always necessary, doing so is usually a best practice. You might want to do this when your data contains a number of columns based on different scales. For instance, say we have a DataFrame with two columns: weight (in ounces) and height (in feet). If you don’t scale or normalize, the algorithm will be less sensitive to variations in height because height values in feet are much lower than weight values in ounces. That’s an example where you should scale your data.
An example of normalization might involve transforming the data so that each point’s value is a representation of its distance from the mean of that column. Using the same example from before, we might want to know how far a given individual’s height is from the mean height. Many algorithms assume that their input data is normalized.
As you might imagine, there are a multitude of algorithms we can apply to our data to scale or normalize it. Enumerating them all is unnecessary here because they are covered in many other texts and machine learning libraries. If you’re unfamiliar with the concept in detail, check out any of the books referenced in the previous lecture. Just keep in mind the fundamental goal—we want our data on the same scale so that values can easily be compared to one another in a sensible way.
**`StandardScaler`**
The `StandardScaler` standardizes a set of features to have zero mean and a standard deviation of 1. The flag `withStd` will scale the data to unit standard deviation while the flag `withMean` (false by default) will center the data prior to scaling it.
WARNING: Centering can be very expensive on sparse vectors because it generally turns them into dense vectors, so be careful before centering your data.
Here’s an example of using a `StandardScaler`:
```
from pyspark.ml.feature import StandardScaler
sScaler = StandardScaler(inputCol="features", withMean=True)
sScaler.fit(scaleDF).transform(scaleDF).show(5, False)
```
**`MINMAXSCALER`**
The `MinMaxScaler` will scale the values in a vector (component wise) to the proportional values on a scale from a given min value to a max value. If you specify the minimum value to be 0 and the maximum value to be 1, then all the values will fall in between 0 and 1:
```
from pyspark.ml.feature import MinMaxScaler
minMax = MinMaxScaler(min=5, max=10, inputCol="features", outputCol="features_MinMaxScaler")
fittedminMax = minMax.fit(scaleDF)
fittedminMax.transform(scaleDF).show()
```
**`MAXABSSCALER`**
The max absolute scaler (`MaxAbsScaler`) scales the data by dividing each value by the maximum absolute value in this feature. All values therefore end up between −1 and 1. This transformer does not shift or center the data at all in the process:
```
from pyspark.ml.feature import MaxAbsScaler
maScaler = MaxAbsScaler(inputCol="features", outputCol="features_MaxAbsScaler")
fittedmaScaler = maScaler.fit(scaleDF)
fittedmaScaler.transform(scaleDF).show(5, False)
```
## Working with Categorical Features
The most common task for categorical features is indexing. Indexing converts a categorical variable in a column to a numerical one that you can plug into machine learning algorithms. While this is conceptually simple, there are some catches that are important to keep in mind so that Spark can do this in a stable and repeatable manner.
In general, we recommend re-indexing every categorical variable when pre-processing just for consistency’s sake. This can be helpful in maintaining your models over the long run as your encoding practices may change over time.
**`StringIndexer`**
The simplest way to index is via the StringIndexer, which maps strings to different numerical IDs. Spark’s StringIndexer also creates metadata attached to the DataFrame that specify what inputs correspond to what outputs. This allows us later to get inputs back from their respective index values:
```
from pyspark.ml.feature import StringIndexer
lblIndxr = StringIndexer(inputCol="lab", outputCol="labelInd")
idxRes = lblIndxr.fit(simpleDF).transform(simpleDF)
idxRes.select("lab", "labelInd").show(5)
```
The default behavior is `stringOrderType='frequencyDesc'`, meaning that the most frequent label gets index 0.
We can also apply `StringIndexer` to columns that are not strings, in which case, they will be converted to strings before being indexed:
```
valIndexer = StringIndexer(inputCol="value1", outputCol="value1Ind")
valIndexer.fit(simpleDF).transform(simpleDF).select("value1", "value1Ind").show(10)
```
Keep in mind that the StringIndexer is an estimator that must be fit on the input data. This means it must see all inputs to select a mapping of inputs to IDs. If you train a StringIndexer on inputs “a,” “b,” and “c” and then go to use it against input “d,” it will throw an error by default. Another option is to skip the entire row if the input value was not a value seen during training. Going along with the previous example, an input value of “d” would cause that row to be skipped entirely. We can set this option before or after training the indexer or pipeline. More options may be added to this feature in the future but as of Spark 2.2, you can only skip or throw an error on invalid inputs.
```
valIndexer.setHandleInvalid("skip")
```
Or while definition:
```
valIndexer = StringIndexer(inputCol="value1", outputCol="value1Ind", handleInvalid="skip")
```
### Converting Indexed Values Back to Text
When inspecting your machine learning results, you’re likely going to want to map back to the original values. Since MLlib classification models make predictions using the indexed values, this conversion is useful for converting model predictions (indices) back to the original categories. We can do this with `IndexToString`. You’ll notice that we do not have to input our value to the String key; Spark’s MLlib maintains this metadata for you. You can optionally specify the outputs.
```
from pyspark.ml.feature import IndexToString
labelReverse = IndexToString(inputCol="labelInd")
labelReverse.transform(idxRes).show(5)
```
**`One-Hot Encoding`**
Indexing categorical variables is only half of the story. One-hot encoding is an extremely common data transformation performed after indexing categorical variables. This is because indexing does not always represent our categorical variables in the correct way for downstream models to process. For instance, when we index our “color” column, you will notice that some colors have a higher value (or index number) than others (in our case, blue is 1 and green is 2).
This is incorrect because it gives the mathematical appearance that the input to the machine learning algorithm seems to specify that green > blue, which makes no sense in the case of the current categories. To avoid this, we use `OneHotEncoder`, which will convert each distinct value to a Boolean flag (1 or 0) as a component in a vector. When we encode the color value, then we can see these are no longer ordered, making them easier for downstream models (e.g., a linear model) to process:
```
from pyspark.ml.feature import OneHotEncoder, StringIndexer
lblIndxr = StringIndexer(inputCol="color", outputCol="colorInd")
colorLab = lblIndxr.fit(simpleDF).transform(simpleDF.select("color"))
ohe = OneHotEncoder(inputCol="colorInd", outputCol="one-hot-enc-color").fit(colorLab)
ohe.transform(colorLab).show(5)
```
### Text Data Transformers
Text is always tricky input because it often requires lots of manipulation to map to a format that a machine learning model will be able to use effectively. There are generally two kinds of texts you’ll see: free-form text and string categorical variables. We already discussed categorical variables in the previous section. At the beginning of this notebook we also saw an example of free-form text and were able to tokenize it. Text is a very interesting and broad discussion and beyond the scope of this course. For more text transformations I refer you to chapter 25 of the textbook and Spark's documentation.
### Further Reading
For more information on transformers visit [Extracting, transforming and selecting features doccumentation page](http://spark.apache.org/docs/latest/ml-features.html).
| github_jupyter |
```
import matplotlib.pyplot as plt
import numpy as np
import os
%matplotlib inline
```
Data loading
--
Graphs for individual subjects are adjcacency matrices organized as follows (n_subjects,nrow,ncol) where nrow,ncol are 65 regions.
Several connectivity measures were calculated : we will use one measure named "tangent"
```
allscales = ['scale007','scale036','scale064','scale122','scale444']
curscale = allscales[1]
# Loading data
curfile = "../mpilmbb/fconn_158subjects_%s_regions.npz" % curscale
# Current ATLAS
roi_maps = '../mpilmbb/basc_2mm_%s.nii.gz' % curscale
X = np.load(curfile)['tangent']
print('X shape (n_subjects,n_regions,n_regions):')
print(X.shape)
```
Here is the final list of subject ids. Note that this is the order that is used in the first axis of the X array
```
subjfiles = np.load(curfile)['subjectids']
subjids = [cur[60:70] for cur in subjfiles]
print(subjids)
```
Visualizing the brain parcellation
--
We learnt a brain parcellation using the resting state data from all subjects of the dataset.
Let's load the file that described the final regions of interest and visualize the 65 regions.
```
from nilearn.plotting import plot_prob_atlas,plot_roi
from nilearn.regions import connected_label_regions
labels_maps = connected_label_regions(roi_maps)
plot_roi(labels_maps,cut_coords=8,display_mode='x')
```
This summarizes all regions over the whole brain. Let's rather plot them one by one.
As this will generate a lot of figures, we create a directory (in the current directory) and directly save the images there.
```
nrois = np.unique(labels_maps.get_data())[-1]
print("Atlas has %d rois " % nrois)
import os
from nilearn.image import math_img
if not(os.path.isdir("roi_images_%s" % curscale)):
os.makedirs("roi_images_%s" % curscale)
from nilearn.image import iter_img
from nilearn.plotting import plot_roi,plot_stat_map
for i in range(1,94):
curimg = math_img('img == %d' % i ,img=labels_maps)
plot_roi(curimg,title="Region %d" % i)
plt.savefig("roi_images_basc/roi_%d.png" % i)
plt.close()
```
Let's extract the center of each region, we will need them for further visualization
```
from nilearn.plotting import find_xyz_cut_coords
roi_maps_4D = []
for i in range(1,nrois+1):
curimg = math_img('img == %d' % i ,img=labels_maps)
roi_maps_4D.append(curimg)
coords_connectome = [find_xyz_cut_coords(img) for img in iter_img(roi_maps_4D)]
coords_connectome = np.stack(coords_connectome)
coords_connectome.shape # Each row is a 3D coordinate of the center of the corresponding region.
```
Now we can visualize connectomes.
We choose a bunch of them randomly.
Note that the choice of threshold will highly influence visualization.
```
from nilearn.plotting import plot_matrix
n_subj_to_plot = 3
for i in np.random.choice(len(subjids),n_subj_to_plot):
plot_matrix(X[i],title='%s' % subjids[i])
plt.show()
from nilearn.plotting import plot_connectome
n_subj_to_plot = 3
for i in np.random.choice(len(subjids),n_subj_to_plot):
curA = k_nearest_neighbor_graph(X[i],2)
plot_connectome(curA,coords_connectome,edge_threshold='99%',title='%s' % subjids[i])
plt.show()
def k_nearest_neighbor_graph(graph, k):
ng = np.zeros(graph.shape)
for i in range(graph.shape[0]):
a = graph[i, :]
ind = np.argpartition(a, -(k+1))[-(k+1):]
for j in range(graph.shape[0]):
if j in ind and j != i:
ng[i,j] = graph[i,j]
for i in range(graph.shape[0]):
for j in range(graph.shape[0]):
ng[i,j] = max(ng[i,j], ng[j,i])
return ng
```
| github_jupyter |
# GLM: Mini-batch ADVI on hierarchical regression model
Unlike Gaussian mixture models, (hierarchical) regression models have independent variables. These variables affect the likelihood function, but are not random variables. When using mini-batch, we should take care of that.
```
%matplotlib inline
%env THEANO_FLAGS=device=cpu, floatX=float32, warn_float64=ignore
import theano
import matplotlib.pyplot as plt
import numpy as np
import pymc3 as pm
import pandas as pd
data = pd.read_csv(pm.get_data('radon.csv'))
county_names = data.county.unique()
county_idx = data['county_code'].values
n_counties = len(data.county.unique())
total_size = len(data)
```
Here, 'log_radon_t' is a dependent variable, while 'floor_t' and 'county_idx_t' determine independent variable.
```
import theano.tensor as tt
log_radon_t = pm.Minibatch(data.log_radon.values, 100)
floor_t = pm.Minibatch(data.floor.values, 100)
county_idx_t = pm.Minibatch(data.county_code.values, 100)
with pm.Model() as hierarchical_model:
# Hyperpriors for group nodes
mu_a = pm.Normal('mu_alpha', mu=0., sigma=100**2)
sigma_a = pm.Uniform('sigma_alpha', lower=0, upper=100)
mu_b = pm.Normal('mu_beta', mu=0., sigma=100**2)
sigma_b = pm.Uniform('sigma_beta', lower=0, upper=100)
```
Intercept for each county, distributed around group mean mu_a. Above we just set mu and sd to a fixed value while here we plug in a common group distribution for all a and b (which are vectors of length n_counties).
```
with hierarchical_model:
a = pm.Normal('alpha', mu=mu_a, sigma=sigma_a, shape=n_counties)
# Intercept for each county, distributed around group mean mu_a
b = pm.Normal('beta', mu=mu_b, sigma=sigma_b, shape=n_counties)
```
Model prediction of radon level `a[county_idx]` translates to `a[0, 0, 0, 1, 1, ...]`, we thus link multiple household measures of a county to its coefficients.
```
with hierarchical_model:
radon_est = a[county_idx_t] + b[county_idx_t] * floor_t
```
Finally, we specify the likelihood:
```
with hierarchical_model:
# Model error
eps = pm.Uniform('eps', lower=0, upper=100)
# Data likelihood
radon_like = pm.Normal('radon_like', mu=radon_est, sigma=eps, observed=log_radon_t, total_size=len(data))
```
Random variable 'radon_like', associated with 'log_radon_t', should be given to the function for ADVI to denote that as observations in the likelihood term.
On the other hand, 'minibatches' should include the three variables above.
Then, run ADVI with mini-batch.
```
with hierarchical_model:
approx = pm.fit(100000, callbacks=[pm.callbacks.CheckParametersConvergence(tolerance=1e-4)])
```
Check the trace of ELBO and compare the result with MCMC.
```
import matplotlib.pyplot as plt
plt.plot(approx.hist);
# Inference button (TM)!
with pm.Model():
mu_a = pm.Normal('mu_alpha', mu=0., sigma=100**2)
sigma_a = pm.Uniform('sigma_alpha', lower=0, upper=100)
mu_b = pm.Normal('mu_beta', mu=0., sigma=100**2)
sigma_b = pm.Uniform('sigma_beta', lower=0, upper=100)
a = pm.Normal('alpha', mu=mu_a, sigma=sigma_a, shape=n_counties)
b = pm.Normal('beta', mu=mu_b, sigma=sigma_b, shape=n_counties)
# Model error
eps = pm.Uniform('eps', lower=0, upper=100)
radon_est = a[county_idx] + b[county_idx] * data.floor.values
radon_like = pm.Normal(
'radon_like', mu=radon_est, sigma=eps, observed=data.log_radon.values)
# essentially, this is what init='advi' does
step = pm.NUTS(scaling=approx.cov.eval(), is_cov=True)
hierarchical_trace = pm.sample(2000, step, start=approx.sample()[0], progressbar=True)
means = approx.bij.rmap(approx.mean.eval())
sds = approx.bij.rmap(approx.std.eval())
from scipy import stats
import seaborn as sns
varnames = means.keys()
fig, axs = plt.subplots(nrows=len(varnames), figsize=(12, 18))
for var, ax in zip(varnames, axs):
mu_arr = means[var]
sigma_arr = sds[var]
ax.set_title(var)
for i, (mu, sigma) in enumerate(zip(mu_arr.flatten(), sigma_arr.flatten())):
sd3 = (-4*sigma + mu, 4*sigma + mu)
x = np.linspace(sd3[0], sd3[1], 300)
y = stats.norm(mu, sigma).pdf(x)
ax.plot(x, y)
if hierarchical_trace[var].ndim > 1:
t = hierarchical_trace[var][i]
else:
t = hierarchical_trace[var]
sns.distplot(t, kde=False, norm_hist=True, ax=ax)
fig.tight_layout()
```
| github_jupyter |
[](https://colab.research.google.com/github/cadCAD-org/demos/blob/master/tutorials/robots_and_marbles/robot-marbles-part-7/robot-marbles-part-7.ipynb)
# cadCAD Tutorials: The Robot and the Marbles, part 7
In parts [1](../robot-marbles-part-1/robot-marbles-part-1.ipynb) and [2](../robot-marbles-part-2/robot-marbles-part-2.ipynb) we introduced the 'language' in which a system must be described in order for it to be interpretable by cadCAD and some of the basic concepts of the library:
* State Variables
* Timestep
* State Update Functions
* Partial State Update Blocks
* Simulation Configuration Parameters
* Policies
In [part 3](../robot-marbles-part-3/robot-marbles-part-3.ipynb) we covered how to describe the presence of asynchronous subsystems within the system being modeled in cadCAD. And [part 4](../robot-marbles-part-4/robot-marbles-part-4.ipynb) introduced Monte Carlo simulations.
In [part 6](../robot-marbles-part-4/robot-marbles-part-6.ipynb) we conducted A/B tests using different system initial conditions to see the differences.
In this notebook, we'll cover cadCAD's support for parameters and parameter sweeping, a useful feature to produce multiple configurations with different parameters. As in Part 6, let's start by copying the base configuration with which we ended Part 4. Here's the description of that system:
__The robot and the marbles__
* Picture a box (`box_A`) with ten marbles in it; an empty box (`box_B`) next to the first one; and __two__ robot arms capable of taking a marble from any one of the boxes and dropping it into the other one.
* The robots are programmed to take one marble at a time from the box containing the largest number of marbles and drop it in the other box. They repeat that process until the boxes contain an equal number of marbles.
* The robots act __asynchronously__ and __non-deterministically__; at every timestep each robot acts with a probability P: 50% for robot 1 and 33.33% for robot 2.
We have added a new parameter in the `robot_arm` function called `capacity`. This is the amount of marbles a robot can move from one box to the other each time.
```
%%capture
# Only run this cell if you need to install the libraries
# If running in google colab, this is needed.
!pip install cadcad matplotlib pandas numpy
# Import dependancies
# Data processing and plotting libraries
import pandas as pd
import numpy as np
from random import random as rand
import matplotlib.pyplot as plt
from pprint import pprint
# cadCAD specific libraries
from cadCAD.configuration.utils import config_sim
from cadCAD.configuration import Experiment
from cadCAD.engine import ExecutionContext, Executor,ExecutionMode
from cadCAD import configs
def p_robot_arm(params, substep, state_history, previous_state, capacity=1):
# Parameters & variables
box_a = previous_state['box_A']
box_b = previous_state['box_B']
# Logic
if box_b > box_a:
b_to_a = capacity
elif box_b < box_a:
b_to_a = -capacity
else:
b_to_a = 0
# Output
return({'add_to_A': b_to_a, 'add_to_B': -b_to_a})
# We specify each of the robots logic in a Policy Function
robots_probabilities = [0.5,1/3] # Robot 1 acts with a 50% probability; Robot 2, 33.33%
def robot_arm_1(params, substep, state_history, previous_state):
_robotId = 1
if rand()<robots_probabilities[_robotId-1]: # draw a random number between 0 and 1; if it's smaller than the robot's parameter, it acts
return p_robot_arm(params, substep, state_history, previous_state)
else:
return({'add_to_A': 0, 'add_to_B': 0})
def robot_arm_2(params, substep, state_history, previous_state):
_robotId = 2
if rand()<robots_probabilities[_robotId-1]: # draw a random number between 0 and 1; if it's smaller than the robot's parameter, it acts
return p_robot_arm(params, substep, state_history, previous_state)
else:
return({'add_to_A': 0, 'add_to_B': 0})
def s_box_A(params, substep, state_history, previous_state, policy_input):
# Parameters & variables
box_A_current = previous_state['box_A']
box_A_change = policy_input['add_to_A']
# Logic
box_A_new = box_A_current + box_A_change
# Output
return ('box_A', box_A_new)
def s_box_B(params, substep, state_history, previous_state, policy_input):
# Parameters & variables
box_B_current = previous_state['box_B']
box_B_change = policy_input['add_to_B']
# Logic
box_B_new = box_B_current + box_B_change
# Output
return ('box_B', box_B_new)
partial_state_update_blocks = [
{
'policies': {
'robot_arm_1': robot_arm_1,
'robot_arm_2': robot_arm_2
},
'variables': { # The following state variables will be updated simultaneously
'box_A': s_box_A,
'box_B': s_box_B
}
}
]
initial_state = {
'box_A': 10, # box_A starts out with 10 marbles in it
'box_B': 0 # box_B starts out empty
}
del configs[:]
MONTE_CARLO_RUNS = 30
SIMULATION_TIMESTEPS = 10
sim_config = config_sim(
{
'N': MONTE_CARLO_RUNS,
'T': range(SIMULATION_TIMESTEPS),
#'M': {} # This will be explained in later tutorials
}
)
experiment = Experiment()
experiment.append_configs(
sim_configs=sim_config,
initial_state=initial_state,
partial_state_update_blocks=partial_state_update_blocks,
)
exec_context = ExecutionContext()
run = Executor(exec_context=exec_context, configs=configs)
(system_events, tensor_field, sessions) = run.execute()
df = pd.DataFrame(system_events)
df.set_index(['simulation', 'run', 'timestep', 'substep'])
ax = None
for i in range(0,MONTE_CARLO_RUNS):
ax = df[df['run']==i+1].plot(x='timestep', y=['box_A','box_B'], marker='o', markersize=12,
markeredgewidth=4, alpha=0.2, markerfacecolor='black',
linewidth=5, figsize=(12,8), title="Marbles in each box as a function of time",
ylabel='Number of Marbles', grid=True, fillstyle='none',
xticks=list(df['timestep'].drop_duplicates()), legend=None,
yticks=list(range(1+(df['box_A']+df['box_B']).max())), ax=ax);
```
## Parameters
What would happen if we "upgrade" the capacity of one of our robot arm to being able to move two marbles at once?
In order to test that, let's first redefine robot arms policies to get their capacity from the `params` parameter. That first `params` parameter is present in policy and state functions, and it makes reference to the dictionary of parameters of the system.
```
robots_probabilities = [0.5,1/3] # Robot 1 acts with a 50% probability; Robot 2, 33.33%
def robot_arm_1(params, substep, state_history, previous_state):
_robotId = 1
capacity = params['capacity'][f'robot_{_robotId}']
if rand()<robots_probabilities[_robotId-1]: # draw a random number between 0 and 1; if it's smaller than the robot's parameter, it acts
return p_robot_arm(params, substep, state_history, previous_state, capacity)
else:
return({'add_to_A': 0, 'add_to_B': 0})
def robot_arm_2(params, substep, state_history, previous_state):
_robotId = 2
capacity = params['capacity'][f'robot_{_robotId}']
if rand()<robots_probabilities[_robotId-1]: # draw a random number between 0 and 1; if it's smaller than the robot's parameter, it acts
return p_robot_arm(params, substep, state_history, previous_state, capacity)
else:
return({'add_to_A': 0, 'add_to_B': 0})
partial_state_update_blocks = [
{
'policies': {
'robot_arm_1': robot_arm_1,
'robot_arm_2': robot_arm_2
},
'variables': { # The following state variables will be updated simultaneously
'box_A': s_box_A,
'box_B': s_box_B
}
}
]
```
You may remember in previous tutorials we left the `M` key as an empty object and promised to use the parameters dictionary in further articles. This is the time. We set each arm's `capacity` here. Let's set the first robot arm capacity to 2 meanwhile we keep the second robot arm's capacity to 1.
Settings of general simulation parameters, unrelated to the system itself
* `T` is a range with the number of discrete units of time the simulation will run for;
* `N` is the number of times the simulation will be run (Monte Carlo runs)
* `M` is a dictionary of parameter keys
```
del configs[:]
MONTE_CARLO_RUNS = 30
SIMULATION_TIMESTEPS = 10
sim_config = config_sim(
{
'N': MONTE_CARLO_RUNS,
'T': range(SIMULATION_TIMESTEPS),
'M': {
'capacity': [
{'robot_1': 2, 'robot_2': 1 }
]
}
}
)
experiment.append_configs(
sim_configs=sim_config,
initial_state=initial_state,
partial_state_update_blocks=partial_state_update_blocks,
)
```
When using params, we recommend to use the multi_proc execution mode. Let's see what is the effect of having an "upgraded" arm over the system.
```
exec_context = ExecutionContext()
run = Executor(exec_context=exec_context, configs=configs)
(system_events, tensor_field, sessions) = run.execute()
df = pd.DataFrame(system_events)
ax = None
for i in range(0,MONTE_CARLO_RUNS):
ax = df[df['run']==i+1].plot(x='timestep', y=['box_A','box_B'], marker='o', markersize=12,
markeredgewidth=4, alpha=0.2, markerfacecolor='black',
linewidth=5, figsize=(12,8), title="Marbles in each box as a function of time",
ylabel='Number of Marbles', grid=True, fillstyle='none',
xticks=list(df['timestep'].drop_duplicates()), legend=None,
yticks=list(range(1+(df['box_A']+df['box_B']).max())), ax=ax);
```
## Parameter sweeping
In order to simulate more than one versions of the same system, we can create many `Configuration` objects to pass to the `Executor` instead of just one. For example, suppose we wanted to test the system under three different arm capacitiy configurations.
Just like we did before, we package those initial conditions along with the partial state update blocks and the simulation parameters into `Configuration` objects. This time using the `append_configs` util for convenience. The configurations are stored in the `configs` list.
```
del configs[:]
MONTE_CARLO_RUNS = 30
SIMULATION_TIMESTEPS = 10
sim_config = config_sim(
{
'N': MONTE_CARLO_RUNS,
'T': range(SIMULATION_TIMESTEPS),
'M': {
'capacity' : [
{ 'robot_1': 1, 'robot_2': 1 }, # Each robot has capacity 1
{ 'robot_1': 2, 'robot_2': 1 }, # Arm 1 has been "upgraded"
{ 'robot_1': 2, 'robot_2': 2 } # Both arms have been "upgraded"
]
}
}
)
experiment.append_configs(
sim_configs=sim_config,
initial_state=initial_state,
partial_state_update_blocks=partial_state_update_blocks,
)
```
And now we can execute the simulation of those three different versions of the system in parallel.
```
execution_mode = ExecutionMode()
exec_context = ExecutionContext(execution_mode.multi_mode)
run = Executor(exec_context=exec_context, configs=configs)
```
And we are ready to execute the simulation. The `execute()` method will return a list of tuples - the first element of those tuples correspond to the datapoints of each one of the versions of the system being simulated.
```
(system_events, tensor_field, sessions) = run.execute()
df = pd.DataFrame(system_events)
for subset in df.subset.unique():
ax = None
for i in range(0,MONTE_CARLO_RUNS):
ax = df[(df['subset']==subset) & (df['run']==i+1)].plot(x='timestep', y=['box_A','box_B'], marker='o', markersize=12,
markeredgewidth=4, alpha=0.2, markerfacecolor='black',
linewidth=5, figsize=(12,8), title="Marbles in each box as a function of time",
ylabel='Number of Marbles', grid=True, fillstyle='none',
xticks=list(df['timestep'].drop_duplicates()), legend=None,
yticks=list(range(1+(df[(df['subset']==subset)]['box_A']+df[(df['subset']==subset)]['box_B']).max())), ax=ax);
```
Can you see the differences between the graphs? You probably can see setting higher capacities in robot arms make the the marbles in the boxes converge faster but have a loosened variation, meanwhile lower capacities make them converge slower but tighter.
As stated, parameters can be used to parametrize the states and policies, and we can run parallel simulations testing different parameters generating different configurations with `config_sim`.
| github_jupyter |
```
%pylab inline
%load_ext autoreload
%autoreload 2
import os
os.chdir("../")
from interpolate import get_latent_vector, slerp
from deepflow.utils import load_generator
from deepflow.plotting_utils import update_matplotlib_config, colorbar
update_matplotlib_config()
import torch
from sklearn.manifold import MDS
from sklearn.decomposition import PCA
import xarray as xr
import tifffile
import matplotlib.colors as colors
import matplotlib.cm as cmx
from mpl_toolkits.axes_grid1.inset_locator import zoomed_inset_axes
from mpl_toolkits.axes_grid1.inset_locator import mark_inset
from mpl_toolkits.axes_grid1 import make_axes_locatable
from matplotlib.ticker import FuncFormatter, MaxNLocator
plot=False
```
## Creating an interpolation animation between the three obtained MAP maxima
```
generator = load_generator("./checkpoints/generator_facies_multichannel_4_6790.pth")
z1_file = "./results/interpolations/run_1/iteration_233.nc"
z2_file = "./results/interpolations/run_4/iteration_253.nc"
z3_file = "./results/interpolations/run_5/iteration_475.nc"
z1 = get_latent_vector(z1_file)
z2 = get_latent_vector(z2_file)
z3 = get_latent_vector(z3_file)
vals = np.linspace(0, 1, 101)
print(vals)
z_int = [torch.from_numpy(slerp(val, z1, z2)).view(1, 50, 1, 2) for val in vals]+ \
[torch.from_numpy(slerp(val, z2, z3)).view(1, 50, 1, 2) for val in vals] + \
[torch.from_numpy(slerp(val, z3, z1)).view(1, 50, 1, 2) for val in vals]
print(len(z_int))
ks = []
for i, z in enumerate(z_int):
k, poro, x = generator(z)
ks.append(k.detach().numpy())
tifffile.imsave("./results/animations/interpolated_1_4_5.tiff", np.array([k[0, 0].T for k in ks]).astype(np.float32))
```
## Plotting the resulting flow, well, and prior loss for interpolated latent vectors
```
def get_interpolations(folder, N=10):
curves = []
poros, perms = [], []
misfits = []
zs = []
for i in range(0, N):
try:
ds = xr.open_dataset(folder+'/iteration_'+str(i)+'.nc')
qo = ds['state_variables'][dict(state_variable=2, well=1)]*(-60*60*24)
qw = ds['state_variables'][dict(state_variable=1, well=1)]*(-60*60*24)
p = ds['state_variables'][dict(state_variable=0, well=0)]/1e5
poros.append(ds['material_properties'][0].values)
perms.append(ds['material_properties'][1].values)
curves.append([qo, qw, p])
zs.append(ds['latent_variables'].values[1])
misfits.append([ds['misfit_value'].values])
ds.close()
except FileNotFoundError:
print(i, " not found ")
return np.array(curves), np.array(poros), np.array(perms), np.array(misfits), np.array(zs)
folders = ['./results/interpolations/interpolation_1_4/', './results/interpolations/interpolation_4_5/', './results/interpolations/interpolation_5_1/']
interpolated_data = [get_interpolations(folder, N=101) for folder in folders]
total_loss = np.concatenate([data[3][:, 0, 0, [0, 1, 3]].sum(1) for data in interpolated_data], 0)
accuracies = np.concatenate([data[3][:, 0, 0, 2] for data in interpolated_data], 0)
flow_loss = np.concatenate([data[3][:, 0, 0, 0] for data in interpolated_data], 0)
well_loss = np.concatenate([data[3][:, 0, 0, 1] for data in interpolated_data], 0)
prior_loss = np.concatenate([data[3][:, 0, 0, 3] for data in interpolated_data], 0)
zs = np.concatenate([data[4] for data in interpolated_data], 0)
N_interpolations = len(zs)
def make_patch_spines_invisible(ax):
ax.set_frame_on(True)
ax.patch.set_visible(False)
for sp in ax.spines.values():
sp.set_visible(False)
def plot_interpolation_losses(host, run_names, flow_loss, well_loss, prior_loss, ranges=[[1.2, 1.4], [1e-1, 1e6], [1e-1, 1e6]]):
fig = host.figure
labels = ['']*303
labels[0] = run_names[0]
labels[100] = run_names[1]
labels[201] = run_names[2]
labels[302] = run_names[0]
f = 16
def format_fn(tick_val, tick_pos):
if int(tick_val) in [0, 100, 201, 302]:
return labels[int(tick_val)]
else:
return ''
host.xaxis.set_ticks([-25, 0, 100, 201, 302, 325])
host.xaxis.set_major_formatter(FuncFormatter(format_fn))
par1 = host.twinx()
par2 = host.twinx()
par2.spines["right"].set_position(("axes", 1.1))
make_patch_spines_invisible(par2)
par2.spines["right"].set_visible(True)
p1, = host.plot(flow_loss, color="black", linestyle="-.", label="Flow Loss")#, c=colors, s=1)
host.set_ylabel("Flow Loss", fontsize=f)
p2, = par1.plot(well_loss, color="red", linestyle="-", label="Well Loss")
par1.set_ylabel("Well Loss", fontsize=f)
p3, = par2.plot(prior_loss, color="blue", linestyle="--", label="Prior Loss")
par2.set_ylabel("Prior Loss", fontsize=f)
host.set_yscale("log")
par1.set_yscale("log")
host.set_xlabel("Interpolations", fontsize=16)
par2.set_ylim(*ranges[0])
host.set_ylim(*ranges[1])
par1.set_ylim(*ranges[2])
host.yaxis.label.set_color(p1.get_color())
par1.yaxis.label.set_color(p1.get_color())
tkw = dict(size=4, width=1.05)
host.tick_params(axis='y', colors=p1.get_color(), **tkw)
par1.tick_params(axis='y', colors=p1.get_color(), **tkw)
host.tick_params(axis='x', **tkw)
host.scatter([0, 100, 201, 302], flow_loss[[0, 100, 201, 302]], marker="o", s=50, color="black")
par1.scatter([0, 100, 201, 302], well_loss[[0, 100, 201, 302]], marker="o", s=50, color="black")
par2.scatter([0, 100, 201, 302], prior_loss[[0, 100, 201, 302]], marker="o", s=50, color="black")
lines = [p1, p2, p3]
host.legend(lines, [l.get_label() for l in lines], fontsize=14, loc=3)
return None
def plot_interpolation_total_loss(host, total_loss, accuracies):
fig = host.figure
labels = ['']*303
labels[0] = run_names[0]
labels[100] = run_names[1]
labels[201] = run_names[2]
labels[302] = run_names[0]
f = 16
def format_fn(tick_val, tick_pos):
if int(tick_val) in [0, 100, 201, 302]:
return labels[int(tick_val)]
else:
return ''
host.xaxis.set_ticks([-25, 0, 100, 201, 302, 325])
host.xaxis.set_major_formatter(FuncFormatter(format_fn))
par1 = host.twinx()
p1, = host.plot(total_loss, color="black", linestyle="-.", label="Total Loss")#, c=colors, s=1)
host.set_ylabel("Total Loss", fontsize=f)
p2, = par1.plot(accuracies, color="red", linestyle="-", label="Well Acc.")
par1.set_ylabel("Well Accuracy", fontsize=f)
host.set_yscale("log")
host.set_xlabel("Interpolations", fontsize=16)
par1.set_ylim(0.8, 1.01)
host.set_ylim(1e-1, 1e6)
host.yaxis.label.set_color(p1.get_color())
par1.yaxis.label.set_color(p1.get_color())
tkw = dict(size=4, width=1.05)
host.tick_params(axis='y', colors=p1.get_color(), **tkw)
par1.tick_params(axis='y', colors=p1.get_color(), **tkw)
host.tick_params(axis='x', **tkw)
host.scatter([0, 100, 201, 302], total_loss[[0, 100, 201, 302]], marker="o", s=50, color="black")
par1.scatter([0, 100, 201, 302], accuracies[[0, 100, 201, 302]], marker="o", s=50, color="black")
lines = [p1, p2]#, p3]
host.legend(lines, [l.get_label() for l in lines], fontsize=14, loc=3)
return None
run_names = ['Run 1', 'Run 4', 'Run 5']
fig, axarr = plt.subplots(figsize=(12, 6))
fig.subplots_adjust(right=0.9)
plot_interpolation_losses(axarr, run_names, flow_loss[:303], well_loss[:303], prior_loss[:303],
[[1.23, 1.42], [1e-1, 1e6], [1e-3, 1e3]])
if plot:
plt.savefig("./results/figures/interpolation_losses_1_4_5_1.png", dpi=300, bbox_inches="tight")
fig, axarr = plt.subplots(figsize=(12, 6))
fig.subplots_adjust(right=0.9)
plot_interpolation_total_loss(axarr, total_loss[0:303], accuracies[0:303])
if plot:
plt.savefig("./results/figures/interpolation_total_loss_1_4_5_1.png", dpi=300, bbox_inches="tight")
```
| github_jupyter |
```
# Taken from https://github.com/xiongyihui/tdoa/blob/master/gcc_phat.py
"""
Estimate time delay using GCC-PHAT
Copyright (c) 2017 Yihui Xiong
Licensed under the Apache License, Version 2.0 (the "License");
you may not use this file except in compliance with the License.
You may obtain a copy of the License at
http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software
distributed under the License is distributed on an "AS IS" BASIS,
WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
See the License for the specific language governing permissions and
limitations under the License.
"""
import numpy as np
def gcc_phat(sig, refsig, fs=1, max_tau=None, interp=16):
'''
This function computes the offset between the signal sig and the reference signal refsig
using the Generalized Cross Correlation - Phase Transform (GCC-PHAT)method.
'''
# make sure the length for the FFT is larger or equal than len(sig) + len(refsig)
n = sig.shape[0] + refsig.shape[0]
# Generalized Cross Correlation Phase Transform
SIG = np.fft.rfft(sig, n=n)
REFSIG = np.fft.rfft(refsig, n=n)
R = SIG * np.conj(REFSIG)
cc = np.fft.irfft(R / np.abs(R), n=(interp * n))
max_shift = int(interp * n / 2)
if max_tau:
max_shift = np.minimum(int(interp * fs * max_tau), max_shift)
cc = np.concatenate((cc[-max_shift:], cc[:max_shift+1]))
# find max cross correlation index
shift = np.argmax(np.abs(cc)) - max_shift
tau = shift / float(interp * fs)
return tau, cc
def main():
refsig = np.linspace(1, 10, 10)
for i in range(0, 10):
sig = np.concatenate((np.linspace(0, 0, i), refsig, np.linspace(0, 0, 10 - i)))
offset, _ = gcc_phat(sig, refsig)
print(offset)
if __name__ == "__main__":
main()
Fs = 8000
dt = 1/Fs #0.125e-3
f1 = 100
tdelay = 0.625e-03 # try different values
t3 = np.linspace(0.0,1.0,Fs)
x3 = np.cos(2*np.pi*f1*t3);
x4 = np.cos(2*np.pi*f1*(t3-tdelay));
gcc_phat(x4,x3)
import numpy as np
import statistics
from scipy.io import wavfile
import matplotlib.pyplot as plt
from IPython.display import Audio
from scipy.signal import kaiserord, lfilter, firwin, freqz
samplerate, data = wavfile.read("Q69905.wav",'rb')
samples = np.arange(20000,30000)
fir_filter = firwin(123,0.75)
channel_1 = lfilter(fir_filter,1, data[samples,0])
channel_2 = lfilter(fir_filter,1, data[samples,1])
channel_3 = lfilter(fir_filter,1, data[samples,2])
channel_4 = lfilter(fir_filter,1, data[samples,3])
noise_1 = np.random.normal(0,1000,len(channel_1))
noise_2 = np.random.normal(0,1000,len(channel_2))
noise_3 = np.random.normal(0,1000,len(channel_3))
noise_4 = np.random.normal(0,1000,len(channel_4))
print([statistics.mean(data[:,0]), statistics.mean(data[:,1]), statistics.mean(data[:,2]), statistics.mean(data[:,3])])
# delay, gcc = gcc_phat(data[samples,0].astype(float)+10, data[samples,2].astype(float)+12, interp=1)
delay, gcc = gcc_phat(channel_1 + noise_1, channel_3 + noise_3, interp=1)
delay_no_noise, gcc_no_noise = gcc_phat(channel_1 , channel_3 , interp=1)
plt.figure(figsize=(20,30))
plt.subplot(4, 1, 1)
plt.plot(data[samples,0])
plt.subplot(4, 1, 2)
plt.plot(np.arange(-10,10),gcc[9990:10010],'.') # [9950:10050]
plt.subplot(4, 1, 3)
plt.plot(np.arange(-10,10),gcc_no_noise[9990:10010],'.') # [9950:10050]
plt.subplot(4, 1, 4)
lags,c, line, b = plt.xcorr(channel_1,channel_3)
plt.plot(lags,c,color='r')
print('GCC-PHAT: ' + str(delay))
print('XCORR: ' + str(lags[np.argmax(c)]))
Audio(channel_1 + noise_1, rate=44100)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/HSE-LAMBDA/DeepGenerativeModels/blob/master/seminars/seminar-9/2.alpha-gan.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
Some code is taken from [github](https://github.com/victor-shepardson/alpha-GAN)
```
use_gpu = True
data_dir = './data/'
import sys
sys.path.append('../')
COLAB = True # Change this variable if you are running the notebook on Colab
if COLAB:
!git clone https://github.com/HSE-LAMBDA/DeepGenerativeModels.git
%cd DeepGenerativeModels/seminars/seminar-9/
%load_ext autoreload
%autoreload 1
%aimport alphagan
!pip install psutil
from collections import defaultdict
from psutil import cpu_count
import numpy as np
import pandas as pd
import torch
from torch import nn
from torch.nn import init, Parameter
import torch.nn.functional as F
from torch.autograd import Variable
from torch.utils.data import DataLoader
from torchvision import datasets, models, transforms
from torchvision.utils import make_grid
import matplotlib.pyplot as plt
%matplotlib inline
# get a image data set into a tensor, scale to (-1, 1), discarding labels
def torchvision_dataset(dset_fn, train=True):
dset = dset_fn(
data_dir,
train=train,
transform=transforms.Compose([
transforms.ToTensor()
]),
target_transform=None,
download=True)
return torch.stack(list(zip(*dset))[0])*2-1
cifar = torchvision_dataset(datasets.CIFAR10, train=True)
cifar_test = torchvision_dataset(datasets.CIFAR10, train=False)
print(cifar.size())
batch_size = 64
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
num_workers = cpu_count() if use_gpu else 0
X_train = DataLoader(cifar, batch_size=batch_size, shuffle=True,
num_workers=num_workers, pin_memory=use_gpu)
X_test = DataLoader(cifar_test, batch_size=batch_size, shuffle=False,
num_workers=num_workers, pin_memory=use_gpu)
class ChannelsToLinear(nn.Linear):
"""Flatten a Variable to 2d and apply Linear layer"""
def forward(self, x):
b = x.size(0)
return super().forward(x.view(b,-1))
class LinearToChannels2d(nn.Linear):
"""Reshape 2d Variable to 4d after Linear layer"""
def __init__(self, m, n, w=1, h=None, **kw):
h = h or w
super().__init__(m, n*w*h, **kw)
self.w = w
self.h = h
def forward(self, x):
b = x.size(0)
return super().forward(x).view(b, -1, self.w, self.h)
class ResBlock(nn.Module):
"""Simple ResNet block"""
def __init__(self, c,
activation=nn.LeakyReLU, norm=nn.BatchNorm2d,
init_gain=1, groups=1):
super().__init__()
self.a1 = activation()
self.a2 = activation()
self.norm1 = norm and norm(c)
self.norm2 = norm and norm(c)
to_init = []
self.conv1 = nn.Conv2d(
c, c, 3, 1, 1, bias=bool(norm), groups=groups)
to_init.append(self.conv1.weight)
self.conv2 = nn.Conv2d(
c, c, 3, 1, 1, bias=bool(norm), groups=groups)
to_init.append(self.conv2.weight)
# if using grouping, add a 1x1 convolution to each conv layer
if groups!=1:
self.conv1 = nn.Sequential(
self.conv1, nn.Conv2d(c,c,1,bias=bool(norm)))
self.conv2 = nn.Sequential(
self.conv2, nn.Conv2d(c,c,1,bias=bool(norm)))
to_init.extend([self.conv1[1].weight, self.conv2[1].weight])
# init
for w in to_init:
init.xavier_normal(w, init_gain)
def forward(self, x):
y = self.conv1(x)
if self.norm1:
y = self.norm1(y)
y = self.a1(y)
y = self.conv2(y)
if self.norm2:
y = self.norm2(y)
return self.a2(x+y)
latent_dim = 128
# encoder network
h = 128
resample = nn.AvgPool2d
norm = nn.BatchNorm2d#None
a, g = nn.ReLU, init.calculate_gain('relu')
groups = 1#h//8
E = nn.Sequential(
nn.Conv2d(3,h,5,1,2), resample(2), a(),
ResBlock(h, activation=a, norm=norm, init_gain=g, groups=groups), resample(2),
ResBlock(h, activation=a, norm=norm, init_gain=g, groups=groups), resample(2),
ResBlock(h, activation=a, norm=norm, init_gain=g, groups=groups),
ChannelsToLinear(h*16, latent_dim)
)
for layer in (0,8):
init.xavier_normal(E[layer].weight, g)
t = Variable(torch.randn(batch_size,3,32,32))
assert E(t).size() == (batch_size,latent_dim)
# generator network
h = 128
norm = nn.BatchNorm2d#None
a, g = nn.ReLU, init.calculate_gain('relu')
groups = 1#h//8
resample = lambda x: nn.Upsample(scale_factor=x)
G = nn.Sequential(
LinearToChannels2d(latent_dim,h,4,4), a(),
ResBlock(h, activation=a, norm=norm, init_gain=g, groups=groups), resample(2),
ResBlock(h, activation=a, norm=norm, init_gain=g, groups=groups), resample(2),
ResBlock(h, activation=a, norm=norm, init_gain=g, groups=groups), resample(2),
ResBlock(h, activation=a, norm=norm, init_gain=g, groups=groups),
nn.Conv2d(h, 3, 1), nn.Tanh()
)
for layer in (0,9):
init.xavier_normal(G[layer].weight, g)
t = Variable(torch.randn(batch_size,latent_dim))
assert G(t).size() == (batch_size,3,32,32)
# discriminator network
h = 128
resample = nn.AvgPool2d
norm = nn.BatchNorm2d
a, g = lambda: nn.LeakyReLU(.2), init.calculate_gain('leaky_relu', .2)
groups = 1
D = nn.Sequential(
nn.Conv2d(3,h,5,1,2), resample(2), a(),
ResBlock(h, activation=a, norm=norm, init_gain=g, groups=groups), resample(2),
ResBlock(h, activation=a, norm=norm, init_gain=g, groups=groups), resample(2),
ResBlock(h, activation=a, norm=norm, init_gain=g, groups=groups),
ChannelsToLinear(h*16, 1), nn.Sigmoid()
)
for layer in (0,8):
init.xavier_normal(D[layer].weight, g)
t = Variable(torch.randn(batch_size,3,32,32))
assert D(t).size() == (batch_size,1)
# code discriminator network
# no batch norm in the code discriminator, it causes trouble
h = 700
a, g = lambda: nn.LeakyReLU(.2), init.calculate_gain('leaky_relu', .2)
C = nn.Sequential(
nn.Linear(latent_dim, h), a(),
nn.Linear(h, h), a(),
nn.Linear(h, 1), nn.Sigmoid(),
)
for i,layer in enumerate(C):
if i%2==0:
init.xavier_normal(layer.weight, g)
t = Variable(torch.randn(batch_size,latent_dim))
assert C(t).size() == (batch_size,1)
model = alphagan.AlphaGAN(E, G, D, C, latent_dim, lambd=40, z_lambd=0).to(device)
# model.load_state_dict(torch.load('../models/cifar10_dim_{}_lambda_{}_zlambd_{}_epochs_{}.torch'.format(
# latent_dim, model.lambd, model.z_lambd, 8
# )))
diag = []
def log_fn(d):
d = pd.DataFrame(d)
diag.append(d)
print(d)
def checkpoint_fn(model, epoch):
path = '../models/cifar10_dim_{}_lambda_{}_zlambd_{}_epochs_{}.torch'.format(
model.latent_dim, model.lambd, model.z_lambd, epoch
)
torch.save(model.state_dict(), path)
model.fit(
X_train, X_test,
n_iter=(2,1,1), n_epochs=10,
log_fn=log_fn, log_every=1,
checkpoint_fn=checkpoint_fn, checkpoint_every=2
)
%matplotlib inline
fig, ax = plt.subplots(1,1,figsize=(14,8))
diagnostic = pd.concat([pd.DataFrame(d.stack(), columns=[i]).T for i,d in enumerate(diag)])
cols = list('rgbcmy')
colors = defaultdict(lambda: cols.pop())
for c in diagnostic:
component, dataset = c
kw = {}
if dataset=='valid':
kw['label'] = component
else:
kw['ls'] = '--'
ax.plot(diagnostic[c].values, c=colors[component], **kw)
ax.legend(bbox_to_anchor=(1, 0.7))
model.eval()
pass
# samples
z, x = model(128, mode='sample')
fig, ax = plt.subplots(1,1,figsize=(16,12))
ax.imshow(make_grid(
x.data, nrow=16, range=(-1,1), normalize=True
).cpu().numpy().transpose(1,2,0), interpolation='nearest')
pass
fig, ax = plt.subplots(1,1,figsize=(16,4))
# training reconstructions
x = cifar[:16]
z, x_rec = model(x)
ax.imshow(make_grid(
torch.cat((x, x_rec.cpu().data)), nrow=16, range=(-1,1), normalize=True
).cpu().numpy().transpose(1,2,0), interpolation='nearest')
pass
fig, ax = plt.subplots(1,1,figsize=(16,4))
# test reconstructions
x = cifar_test[:16]
z, x_rec = model(x)
ax.imshow(make_grid(
torch.cat((x, x_rec.cpu().data)), nrow=16, range=(-1,1), normalize=True
).cpu().numpy().transpose(1,2,0), interpolation='nearest')
pass
```
### pairwise distances give a sense of how encoded z are distributed compared the prior:
```
fig, ax = plt.subplots(1,1,figsize=(16,5))
n = 64
import scipy.spatial
train_z = model(cifar[:n], mode='encode').data.cpu().numpy()
test_z = model(cifar_test[:n], mode='encode').data.cpu().numpy()
sampled_z = model.sample_prior(n).data.cpu().numpy()
train_cdist = scipy.spatial.distance.cdist(train_z,train_z)
test_cdist = scipy.spatial.distance.cdist(test_z,test_z)
sampled_cdist = scipy.spatial.distance.cdist(sampled_z,sampled_z)
cax = ax.imshow(np.concatenate((train_cdist, test_cdist, sampled_cdist),1))
fig.colorbar(cax)
ax.set_xlabel('train, test, sampled z pairwise distances')
pass
```
### interpolation
```
def slerp(z0, z1, t):
"""Spherical linear interpolation over last dimension:
z0.shape = z1.shape = (m,...n, d) and t.shape = (q,) -> (q, m,...n, d)
Project to unit sphere and linearly interpolate magnitudes.
"""
m0, m1 = (np.linalg.norm(z, 2, -1) for z in (z0, z1))
p0, p1 = z0/np.expand_dims(m0,-1), z1/np.expand_dims(m1,-1)
omega = np.arccos((p0*p1).sum(-1))
while t.ndim<=omega.ndim:
t = np.expand_dims(t,-1)
sin_omega = np.sin(omega)
t0 = np.sin((1-t)*omega)/sin_omega
t1 = np.sin(t*omega)/sin_omega
lim = abs(omega)<1e-15
t1[lim] = t[lim]
t0[lim] = (1-t)[lim]
t0, t1 = np.expand_dims(t0, -1), np.expand_dims(t1, -1)
m = np.expand_dims((1-t)*m0 + t*m1, -1)
return m*(t0*p0 + t1*p1)
def slerp4(z, t):
# z (4,n) and t (m,) -> z' (m, m, n)
return slerp(
slerp(z[0], z[1], t),
slerp(z[2], z[3], t),
t)
# torch.manual_seed(2)
n = 7
fig,ax = plt.subplots(2,3,figsize=(17,11))
for ax in ax.flatten():
x = cifar_test[torch.LongTensor(np.random.randint(len(cifar_test),size=4))]
z = model(x, mode='encode')
z = slerp4(z.data.cpu().numpy(), np.linspace(0,1,n)).reshape(n*n,-1)
x_rec = model(z, mode='generate')
ax.imshow(make_grid(
x_rec.cpu().data, nrow=n, range=(-1,1), normalize=True
).numpy().transpose(1,2,0), interpolation='nearest')
```
| github_jupyter |
```
!pip install -q ../input/segmentation-models-wheels/efficientnet_pytorch-0.6.3-py3-none-any.whl
!pip install -q ../input/segmentation-models-wheels/pretrainedmodels-0.7.4-py3-none-any.whl
!pip install -q ../input/segmentation-models-wheels/timm-0.3.2-py3-none-any.whl
!pip install -q ../input/segmentation-models-wheels/segmentation_models_pytorch-0.1.3-py3-none-any.whl
import wandb
import os
import cv2
import time
import warnings
import random
import numpy as np
import pandas as pd
from tqdm import tqdm_notebook as tqdm
from torch.optim.lr_scheduler import ReduceLROnPlateau
from sklearn.model_selection import KFold
import torch
import torch.nn as nn
from torch.nn import functional as F
import torch.optim as optim
import torch.backends.cudnn as cudnn
from torch.utils.data import DataLoader, Dataset, sampler
from matplotlib import pyplot as plt
import albumentations as A
from albumentations.pytorch import ToTensorV2
from PIL import Image
from sklearn.metrics import accuracy_score, f1_score
import torch
import collections.abc as container_abcs
torch._six.container_abcs = container_abcs
import segmentation_models_pytorch as smp
warnings.filterwarnings("ignore")
def fix_all_seeds(seed):
np.random.seed(seed)
random.seed(seed)
os.environ['PYTHONHASHSEED'] = str(seed)
torch.manual_seed(seed)
torch.cuda.manual_seed(seed)
torch.cuda.manual_seed_all(seed)
fix_all_seeds(2021)
# from wandb.keras import WandbCallback
try:
from kaggle_secrets import UserSecretsClient
user_secrets = UserSecretsClient()
api_key = user_secrets.get_secret("WANDB")
wandb.login(key=api_key)
anonymous = None
except:
anonymous = "must"
print('To use your W&B account,\nGo to Add-ons -> Secrets and provide your W&B access token. Use the Label name as WANDB. \nGet your W&B access token from here: https://wandb.ai/authorize')
class config:
seed = 101
debug = False # set debug=False for Full Training
exp_name = 'Unet-effnetb2-512x512-aug2'
model_name = 'Unet'
backbone0 = 'efficientnet-b2'
backbone1 = 'efficientnet-b7'
backbone2 = 'efficientnet-b6'
backbone3 = 'efficientnet-b5'
batch_size = 8
train_bs = 24
valid_bs = 48
IMAGE_RESIZE = (512, 512)
epochs = 2
lr = 5e-3
scheduler = 'CosineAnnealingLR'
min_lr = 1e-6
T_max = int(100*6*1.8)
T_0 = 25
warmup_epochs = 0
wd = 1e-6
n_accumulate = 32//train_bs
# n_fold = 5
num_classes = 1
DEVICE = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
competition = 'sartorius'
_wandb_kernel = 'somusan'
TEST_PATH = "../input/sartorius-cell-instance-segmentation/test"
model = smp.Unet(config.backbone0, encoder_weights="imagenet", classes=1,activation=None)
model1 = smp.Unet(config.backbone1, encoder_weights="imagenet", classes=1,activation=None)
model2 = smp.Unet(config.backbone2, encoder_weights="imagenet", classes=1,activation=None)
model3 = smp.Unet(config.backbone3, encoder_weights="imagenet", classes=1,activation=None)
best_model0 = wandb.restore('model0.pth', run_path="somusan/kaggle_sartorius_unet_ensemble/dnhtjlrc")
model.load_state_dict(torch.load(best_model0.name))
best_model1 = wandb.restore('model1.pth', run_path="somusan/kaggle_sartorius_unet_ensemble/dnhtjlrc")
model1.load_state_dict(torch.load(best_model1.name))
best_model2 = wandb.restore('model2.pth', run_path="somusan/kaggle_sartorius_unet_ensemble/dnhtjlrc")
model2.load_state_dict(torch.load(best_model2.name))
best_model3 = wandb.restore('model3.pth', run_path="somusan/kaggle_sartorius_unet_ensemble/dnhtjlrc")
model3.load_state_dict(torch.load(best_model3.name))
class TestCellDataset(Dataset):
def __init__(self):
self.test_path = config.TEST_PATH
self.image_ids = [f[:-4]for f in os.listdir(self.test_path)]
self.num_samples = len(self.image_ids)
self.transform = A.Compose([A.Resize(config.IMAGE_RESIZE[0], config.IMAGE_RESIZE[1]), ToTensorV2()])
def __getitem__(self, idx):
image_id = self.image_ids[idx]
path = os.path.join(self.test_path, image_id + ".png")
image = cv2.imread(path)
image = self.transform(image=image)['image']
return {'image': image, 'id': image_id}
def __len__(self):
return self.num_samples
ds_test = TestCellDataset()
dl_test = DataLoader(ds_test, batch_size=1, shuffle=False, num_workers=4, pin_memory=True)
def post_process(probability, threshold=0.5, min_size=300):
mask = cv2.threshold(probability, threshold, 1, cv2.THRESH_BINARY)[1]
num_component, component = cv2.connectedComponents(mask.astype(np.uint8))
predictions = []
for c in range(1, num_component):
p = (component == c)
if p.sum() > min_size:
a_prediction = np.zeros((520, 704), np.float32)
a_prediction[p] = 1
predictions.append(a_prediction)
return predictions
def rle_encoding(x):
dots = np.where(x.flatten() == 1)[0]
run_lengths = []
prev = -2
for b in dots:
if (b>prev+1): run_lengths.extend((b + 1, 0))
run_lengths[-1] += 1
prev = b
return ' '.join(map(str, run_lengths))
def get_preds(model):
model.eval()
submission = []
for i, batch in enumerate(tqdm(dl_test)):
preds = torch.sigmoid(model(batch['image'].float()))
preds = preds.detach().cpu().numpy()[:, 0, :, :] # (batch_size, 1, size, size) -> (batch_size, size, size)
for image_id, probability_mask in zip(batch['id'], preds):
probability_mask = cv2.resize(probability_mask, dsize=(704, 520), interpolation=cv2.INTER_LINEAR)
submission.append(probability_mask)
return submission
plt.imshow(submission[0]>0.5)
import numpy as np
import cv2 as cv
from matplotlib import pyplot as plt
# img = cv.imread('opencv_logo.png')
plt.figure(figsize=(20,20))
img = submission[0] > 0.5
# kernel = np.ones((5,5),np.float32)/25
# dst = cv.filter2D(submission[0],-1,kernel)
dst = cv2.medianBlur(img, 5)
plt.subplot(121),plt.imshow(img),plt.title('Original')
plt.xticks([]), plt.yticks([])
plt.subplot(122),plt.imshow(dst),plt.title('Averaging')
plt.xticks([]), plt.yticks([])
plt.show()
model.eval()
submission = []
for i, batch in enumerate(tqdm(dl_test)):
preds = torch.sigmoid(model(batch['image'].cuda()))
preds = preds.detach().cpu().numpy()[:, 0, :, :] # (batch_size, 1, size, size) -> (batch_size, size, size)
for image_id, probability_mask in zip(batch['id'], preds):
try:
#if probability_mask.shape != IMAGE_RESIZE:
# probability_mask = cv2.resize(probability_mask, dsize=IMAGE_RESIZE, interpolation=cv2.INTER_LINEAR)
probability_mask = cv2.resize(probability_mask, dsize=(704, 520), interpolation=cv2.INTER_LINEAR)
predictions = post_process(probability_mask)
for prediction in predictions:
#plt.imshow(prediction)
#plt.show()
try:
submission.append((image_id, rle_encoding(prediction)))
except:
print("Error in RL encoding")
except Exception as e:
print(f"Exception for img: {image_id}: {e}")
# Fill images with no predictions
image_ids = [image_id for image_id, preds in submission]
if image_id not in image_ids:
submission.append((image_id, ""))
df_submission = pd.DataFrame(submission, columns=['id', 'predicted'])
df_submission.to_csv('submission.csv', index=False)
if df_submission['predicted'].apply(check_is_run_length).mean() != 1:
print("Check run lenght failed")
create_empty_submission()
# from keras.models import load_model
#Set compile=False as we are not loading it for training, only for prediction.
# model1 = load_model('saved_models/res34_backbone_50epochs.hdf5', compile=False)
# model2 = load_model('saved_models/inceptionv3_backbone_50epochs.hdf5', compile=False)
# model3 = load_model('saved_models/vgg19_backbone_50epochs.hdf5', compile=False)
#Weighted average ensemble
# models = [model1, model2, model3]
#preds = [model.predict(X_test) for model in models]
# pred1 = model1.predict(X_test1)
# pred2 = model2.predict(X_test2)
# pred3 = model3.predict(X_test3)
pred0 = get_preds(model)
pred1 = get_preds(model1)
pred2 = get_preds(model2)
pred3 = get_preds(model3)
preds=np.array([pred0[0], pred1[0], pred2[0], pred3[0]])
#preds=np.array(preds)
weights = [0.3, 0.5, 0.3,0.4]
#Use tensordot to sum the products of all elements over specified axes.
weighted_preds = np.tensordot(preds, weights, axes=((0),(0)))
weighted_ensemble_prediction = np.argmax(weighted_preds, axis=3)
y_pred1_argmax=np.argmax(pred1, axis=3)
y_pred2_argmax=np.argmax(pred2, axis=3)
y_pred3_argmax=np.argmax(pred3, axis=3)
#Using built in keras function
n_classes = 1
IOU1 = MeanIoU(num_classes=n_classes)
IOU2 = MeanIoU(num_classes=n_classes)
IOU3 = MeanIoU(num_classes=n_classes)
IOU_weighted = MeanIoU(num_classes=n_classes)
IOU1.update_state(y_test[:,:,:,0], y_pred1_argmax)
IOU2.update_state(y_test[:,:,:,0], y_pred2_argmax)
IOU3.update_state(y_test[:,:,:,0], y_pred3_argmax)
IOU_weighted.update_state(y_test[:,:,:,0], weighted_ensemble_prediction)
print('IOU Score for model1 = ', IOU1.result().numpy())
print('IOU Score for model2 = ', IOU2.result().numpy())
print('IOU Score for model3 = ', IOU3.result().numpy())
print('IOU Score for weighted average ensemble = ', IOU_weighted.result().numpy())
```
| github_jupyter |
<a href="https://colab.research.google.com/github/trendinafrica/python_workshop/blob/main/06_TReND_Python_Practical2_all_mixed.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>

# TReND Python Course: Day 2, Practical 2
# Topic: Data Types, Operators, Loops & Data Structures
__Content creators:__ Cameron Smith & Brian Mboya
# Introduction
In this course we will practice what we have learned in previous courses on Data Types (Theory 1), Operators (Theory 2), Data Structures (Theory 3) and Operators (Theory 4). The idea is to train our understanding of the theory concepts. This tutorial provides you with problem sets and your goal is to formulate a solution to it. Note that in most cases different solutions can solve the problem. The problems are divided into levels: Think of levels as difficulty.
- Level 1: [tricky](#easy)
- Level 2: [trickier](#medium)
- Level 3: [trickiest](#difficult)
Note: Do whatever can be done in the time you have. If not all, then the rest can be done during free time.
---
# Let's practise!
```
# @title The code hidden here is for testing your code, do not modify it please :).
def test_results(your_input, expected_input):
try:
assert your_input == expected_input
except AssertionError as e:
print("\x1b[6;30;91m", "TEST FAILED - You are almost there. Give it another try :)" + "\x1b[0m")
if type(your_input) == type(expected_input):
raise AssertionError(f'\x1b[6;30;91m {your_input} != {expected_input} \x1b[0m')
elif type(your_input) != type(expected_input):
raise AssertionError(f'\x1b[6;30;91m {type(your_input)} != {type(expected_input)} \x1b[0m')
else:
raise e
else:
print('\x1b[6;30;92m' + 'TEST PASSED! - Congratulations, you found a solution! Good Job!' + '\x1b[0m')
```
---
<a name="easy"></a>
### **Level 1: tricky**
### 1.1 Lists
- Given a list `nums`, print the length of the list.
```
nums = [73, 23, 45, 54, 29, 49, 2, -13, 48, 82, 49, 284, 204, 1337] # list to use
# Write your code here!
your_answer = ...
# @title ##### Run this. Have you found a solution?
# Tests
test_results(your_answer, 14)
# @title Double click here for the solution
# solution to be added
your_answer = len(nums)
```
- Given `nums`, print the value of the last index.
```
# Write your code here!
your_answer = ...
# @title ##### Run this. Have you found a solution?
# Tests
test_results(your_answer, 1337)
# @title Double click here for the solution
# solution to be added
your_answer = nums[-1]
```
- Given `nums`, select the last five elements of the list.
```
# Write your code here!
your_answer = ...
# @title ##### Run this. Have you found a solution?
# Tests
test_results(your_answer, [82, 49, 284, 204, 1337])
# @title Double click here for the solution
# solution to be added
your_answer = nums[-5:]
```
- Given `nums`, find the sum of all elements in the list using a `for loop`. Compare your result with the output found using the `sum`; python in-built function.
```
# Write your code here!
# Using for loop
total_using_forloop = 0
# Using the sum in-built function
total_using_sum = ...
# @title ##### Run this. Have you found a solution?
# Test 1
test_results(total_using_forloop, 2266)
# Test 2
test_results(total_using_sum == total_using_forloop, True)
# @title Double click here for the solution
# solution to be added
# Using for loop
total_using_forloop = 0
for num in nums:
total_using_forloop += num
# Using the sum in-built function
total_using_sum = sum(nums)
```
- Given a list of integers `odds_evens` (below), loop through the list and using `append`, add all the even numbers in the list `evens` and all odds in list `odds`. Then print the result. Note: To check if a number is even in python, you do, `if number % 2 == 0:` otherwise it's odd
```
odds_evens = [2, 4, 5, 6, 92, 54, 55, 53, 12, 32, 13, 95, 82, 77, 45, 65, 66, 22, 23, 15, 91, 100, 35, 88, 6, 8, 9, 11, 7]
evens = []
odds = []
# Write your code here
# @title ##### Run this. Have you found a solution?
# Test even
test_results(evens, [2, 4, 6, 92, 54, 12, 32, 82, 66, 22, 100, 88, 6, 8])
# Test odd
test_results(odds, [5, 55, 53, 13, 95, 77, 45, 65, 23, 15, 91, 35, 9, 11, 7])
# @title Double click here for the solution
# solution to be added
# Write your code here!
for num in odds_evens:
if num & 1: # or num % 2 != 0
odds.append(num)
else:
evens.append(num)
```
- Using the above results, sort the resulting lists; `evens` and `odds`
```
# Write your code here!
# @title ##### Run this. Have you found a solution?
# Test even sorted
test_results(evens, [2, 4, 6, 6, 8, 12, 22, 32, 54, 66, 82, 88, 92, 100])
# Test odd sorted
test_results(odds, [5, 7, 9, 11, 13, 15, 23, 35, 45, 53, 55, 65, 77, 91, 95])
# @title Double click here for the solution
# solution to be added
evens.sort()
odds.sort()
```
- Concatenate `evens` and `odds` above and assert that your result is equally the same as the original list `odds_evens`. Note; you need to sort the list before asserting. Also, you can use both methods learned to concatenate the lists; not a requirement
```
# Write your code here!
new_nums = ...
# @title ##### Run this. Have you found a solution?
# Test
test_results(new_nums == odds_evens, True)
# @title Double click here for the solution
# solution to be added
new_odds_evens = sorted(evens + odds)
```
- Alice is attending a private party but she's not sure if her name is on the invited guests list.
```
# @title ##### Run this. The list is hidden here. Don't peek!
names = ['Olivia', 'Emma', 'Ava', 'Charlotte', 'Sophia', 'Amelia', 'Isabella', 'Mia', 'Evelyn', 'Harper', 'Camila', 'Gianna', 'Abigail', 'Luna', 'Ella', 'Elizabeth', 'Sofia', 'Emily', 'Avery', 'Mila', 'Scarlett', 'Eleanor', 'Madison', 'Layla', 'Penelope', 'Aria', 'Chloe', 'Grace', 'Alice', 'Ellie', 'Nora', 'Hazel', 'Zoey', 'Riley', 'Victoria', 'Lily', 'Aurora', 'Violet', 'Nova', 'Hannah', 'Emilia', 'Zoe', 'Stella', 'Everly', 'Isla', 'Leah', 'Lillian', 'Addison', 'Willow', 'Lucy', 'Paisley']
```
- Alice doesn't like embarrassment and does not want to walk up to the gate not knowing, if her name is on the list or not. Alices hands you a list of names of the guests called `names`. Help her find out if she's on the list or not. Print the results.
```
my_name = 'Alice'
# Write your code here!
name_is_on_list = ... # is the name on the list (True) or not (False)?
# @title ##### Run this to know: Did you get it right?
# Tests
test_results(name_is_on_list, True)
# @title Double click here for the solution
# solution to be added
name_is_on_list = 'Alice' in names
```
### 1.2 Dictionaries
- Given an (unkonw) dictionary `persons`, which contatins the name of a person and their age. Perform the following operations.
```
# @title ##### Run this. The unkown dictionary is hidden here. Don't peek!
persons = {'Alice': 22, 'Bob': 26, 'Yoda': 123, 'Xii': 1, 'V': 28, 'Watson': 12}
```
- Given the dictionary `persons`, print a list of all the keys of the dictionary.
```
# Write your code here!
keys = ...
# @title ##### Run this. Have you found a solution?
# Tests
test_results(keys, ['Alice', 'Bob', 'Yoda', 'Xii', 'V', 'Watson'])
# @title Double click here for the solution
# solution to be added
keys = list(persons.keys())
```
- Given the dictionary `persons`, print a list of all values.
```
# Write your code here!
values = ...
# @title ##### Run this. Have you found a solution?
# Test
test_results(values, [22, 26, 123, 1, 28, 12])
# @title Double click here for the solution
# solution to be added
values = list(persons.values())
```
- Given the dictionary `persons`, print key-value pairs or items.
```
# Write your code here!
```
- Given the dictionary `persons`, return the age of the person mentioned below,
if the person exists in the dictionary. If it does not exist, add the person with age 999 to the dictionary and print the new dict. Check, if you have successfully added the person. Add your own name and age to the dictionary.
```
person = 'Yoda'
# Write your code here!
age = ...
# @title ##### Run this. Have you found a solution?
# Test
test_results(age, 123)
# @title Double click here for the solution
# solution to be added
age = persons[person]
person2 = 'Brian'
# Check if person exists
person_exits = ...
# If person_exits is false, write your code here to add the person to the dictionary persons. Remember age is 999 if they aren't included!
...
# @title ##### Run this. Have you found a solution?
# Test new persons
test_results(persons, {'Alice': 22, 'Bob': 26, 'Yoda': 123, 'Xii': 1, 'V': 28, 'Watson': 12, 'Brian': 999})
# @title Double click here for the solution
# solution to be added
# Check if person exists
person_exits = person2 in persons
# If person_exits is false, write your code here to add the person to the dictionary persons
if not person_exits:
persons[person2] = 999
```
- Add a new key called `'location'` with the value `'New Zealand'` to the dictionary `my_friend`. Print the new dictionary.
Be sure to add it on a new line. Do not add the value in the first line where the dictionary is defined. Remember, capitalization matters!
```
my_friend = {'name': 'Fluffball', 'occupation': 'Cat'} # don't change anything here
## Add your code here!
# @title ##### Run this. Have you found a solution?
test_results(my_friend, {'name': 'Fluffball', 'occupation': 'Cat', 'location': 'New Zealand'})
# @title Double click here for the solution
# solution to be added
my_friend['location'] = 'New Zealand'
```
### 1.3 Loops 'n Lists
- Given the top secret hidden list `top_secret`, use a for-loop to print each element in the given list on a new line.
```
# @title ##### Run this. Top secret hidden list. Don't peek!
top_secret = [1,2,'woah',4,5,6]
## Add your code here!
```
- Use a `while`-Loop to add one to the variable `counter` until it is greater than zero. Print the new value of `counter`.
```
counter = -10
## Add your code here!
# @title ##### Run this. Have you found a solution?
test_results(counter, 1)
# @title Double click here for the solution
# solution to be added
while counter < 1:
counter = counter + 1
```
- Insert the string `'owl'` into the list `animals_seen` at the first position using the `.insert()` function. Print the new list. You can read a short paragraph on the `insert()` function [here](https://www.programiz.com/python-programming/methods/list/insert).
```
animals_seen = ['cow', 'emu', 'crane']
# Add your code here!
# @title ##### Run this. Have you found a solution?
test_results(animals_seen, ['owl', 'cow', 'emu', 'crane'])
# @title Double click here for the solution
# solution to be added
animals_seen.insert(0, 'owl')
```
### 1.4 [SELF-STUDY] Tuples
- Create two lists `my_fav_food` and `my_fav_friends`. Make a tuple containing both lists. Then print the tuple.
```
## TODO: create lists and tuple
my_food_friends_tuple = ...
## TODO print
# @title ##### Run this. Have you found a solution?
test_results([type(my_food_friends_tuple), type(my_food_friends_tuple[0]), type(my_food_friends_tuple[1])], [tuple, list, list])
# @title Double click here for the solution
# solution to be added
my_food_friends_tuple = (['Pizza'], ['Pizza is also my favorite friend'])
```
---
<a name="medium"></a>
### **Level 2: trickier**
### 2.1 Lists
- Given the list of numbers `num`, perform the following operations.
```
# Given this list of numbers
nums = [73, 23, 45, 54, 29, 49, 2, -13, 48, 82, 49, 284, 204, 1337] # list to use
# Operation 1: Change the value at index 7 to 147 in copy_nums. Do not change original nums.
copy_nums = nums[::] # this is how copying a list is done in python. There other ways too, find link below
# Write your code here!
```
- [Copying a list in python](https://www.programiz.com/python-programming/methods/list/copy)
```
# @title ##### Run this. Have you found a solution?
# Tests
test_results(copy_nums, [73, 23, 45, 54, 29, 49, 2, 147, 48, 82, 49, 284, 204, 1337])
# @title Double click here for the solution
# solution to be added
copy_nums[7] = 147
# Operation 2: Take the value at index 2 and set it equal to 4 using the copy_nums1. Do not change original nums.
copy_nums1 = nums[::]
# Write your code here!
# @title ##### Run this. Have you found a solution?
# Tests
test_results(copy_nums1, [73, 23, 4, 54, 29, 49, 2, -13, 48, 82, 49, 284, 204, 1337])
# @title Double click here for the solution
# solution to be added
copy_nums1[2] = 4
# Operation 3: Remove the first and last element from the list copy_nums2
# Note: do not use the original list nums defined above, make a copy.
copy_nums2 = nums[::]
# Write your code here!
# @title ##### Run this. Have you found a solution?
# Tests
test_results(copy_nums2, [23, 45, 54, 29, 49, 2, -13, 48, 82, 49, 284, 204])
# @title Double click here for the solution
# solution to be added
copy_nums2 = copy_nums2[1:-1]
```
- Given a sorted list of distinct integers; `sorted_nums` and a target value, the following function returns the index if the target is found. If not, it returns the index where it would be if it were inserted in order. Complete the function by writing your for loop where asked.
For example:
```
Input: sorted_nums = [1, 3, 4, 5, 6, 8]
target = 2
Output: 1
input: sorted_nums = [1, 3, 4, 5, 6, 8]
target = 5
Output: 3
```
```
sorted_nums = [2, 3, 5, 6, 9, 10, 11, 14, 16, 18, 20, 23, 25, 28, 32, 44, 54, 67, 77, 88, 90, 95, 100, 117]
target1 = 28
target2 = 22
target3 = 200
target4 = 1
def find_index(target):
found_index = -1
end = None # we set this when there is no such value in the list, this hold the next index
if target < sorted_nums[0]:
return 0
# Write your code here!
...
if found_index < 0 and end:
return end - 1
elif found_index < 0:
return len(sorted_nums)
return found_index
# target1
result1 = find_index(target1)
# target2
result2 = find_index(target2)
# target3
result3 = find_index(target3)
# target4
result4 = find_index(target4)
# @title ##### Run this. Have you found a solution?
# Test target1
test_results(result1, 13)
# Test target2
test_results(result2, 10)
# Test target3
test_results(result3, 24)
# Test target4
test_results(result4, 0)
# @title Double click here for the solution
# solution to be added
def find_index(target):
found_index = -1
end = None
if target < sorted_nums[0]:
return 0
# Write your code here!
for index, num in enumerate(sorted_nums):
if num == target:
found_index = index
break
elif num > target:
end = index
break
if found_index < 0 and end:
return end - 1
elif found_index < 0:
return len(sorted_nums)
return found_index
```
### 2.2 Dictionaries
- Given a dict of passwords, print password owner and the password, if valid. A valid password is one that does not have repeating characters.
```
Input: passwords = {1: 'ahsgdfk', 2: 'sbgri', 3: 'jjshhgs', 4: 'shggryyas'}
Output: 1: ahsgdfk
2: sbgri
```
```
passwords = {'A': 'aseddc', 'B': 'cdeesaq', 'C': 'lkkdjur', 'D': 'kksgrruee', 'E': 'jueysh', 'F': 'grteusi', 'G': 'uyttee', 'H': 'hfgtry', 'I': 'lkbyww', 'J': 'gfbtwsj'}
# Write your code here!
# @title Double click here for the solution
# solution to be added
for owner, password in passwords.items():
dupes_removed = set(password)
valid = len(dupes_removed) == len(password)
if valid:
print(f'{owner}: {password}')
```
- Using the previous code, loop through all passwords. Instead of printing the valid passwords, create a string formatted as `'Key: Value'`. Add this string to a set called `result`.
```
# Write your code here!
# @title ##### Run this. Have you found a solution?
# Test
test_results(owners, {'J: gfbtwsj', 'F: grteusi', 'H: hfgtry', 'E: jueysh'})
# @title Double click here for the solution
# solution to be added
for owner, password in passwords.items():
dupes_removed = set(password)
valid = len(dupes_removed) == len(password)
if valid:
owners.add(f'{owner}: {password}')
```
### 2.3 Loops
- Write a nested for-loop that combines all letters in the variable given below. Set the variable `answer` equal to that string.
```
prob_six_list = [['g','o'], ['o','d'], [' '], ['j','o','b','!']]
answer = ''
# Add your code here!
# @title ##### Run this. Have you found a solution?
test_results(answer, 'good job!')
# @title Double click here for the solution
# solution to be added
for single_list in prob_six_list:
for element in single_list:
answer = answer + element
```
- Complete a program that keeps asking an interger input from the user and print the sum of all the user unput so far until the user input `-1`
```
# Insert your code here
result = 0
while True:
number = int(input("Enter an integer: "))
if #insert your code here:
break
result += # insert your code here
print(result)
# @title Double click here for the solution
result = 0
while True:
number = int(input("Enter an integer: "))
if number == -1 :
break
result += number
print(result)
```
---
<a name="difficult"></a>
### **Level 3: trickiest**
### 3.1 Lists
- You are given a list of lists of scores, the outer list represents the students and the inner list represents student scores for each subject the student undertake. Find the student with the highest score and return the score.
For example;
Given;
```
Input: students = [[23, 32, 41, 25], [43, 55, 45, 57], [12, 32, 43, 55]]
Output: 200
Explanation:
student 1: 23 + 32 + 41 + 25 = 121
student 2: 43 + 55 + 45 + 57 = 200
student 3: 12 + 32 + 43 + 55 = 142
You can use sum python in-built function, sum([23, 32, 41, 25])
The highest is student 2 with 200 score
```
```
students = [[23, 32, 41, 25], [43, 55, 45, 57], [12, 32, 43, 55], [21, 23, 34, 59], [32, 23, 14, 52], [34, 55, 54, 75], [93, 77, 65, 67]]
# Write your code here!
highest_score = ...
...
# @title ##### Run this. Have you found a solution?
# Tests
test_results(highest_score, 302)
# @title Double click here for the solution
# solution to be added
for student in students:
total = sum(student)
if total > highest_score:
highest_score = total
```
### 3.2 Lists & Dictionaries
- Given a list of integers, create a dictionary, call it `seen`, of key-value representing the amount of time a number is seen in the list. Print the dictionary.
For example:
```
Input: numbers = [1, 2, 2, 7, 7, 7, 5, 4, 4, 8, 8]
Output: {1: 1, 2: 2, 7: 3, 5: 1, 4: 2, 8: 2}
```
```
numbers = [1, 2, 2, 7, 7, 7, 5, 4, 4, 8, 8]
# Write your code here
seen = {}
...
# @title ##### Run this. Have you found a solution?
# Test
test_results(seen, {1: 1, 2: 2, 7: 3, 5: 1, 4: 2, 8: 2})
# @title Double click here for the solution
# solution to be added
seen = {}
for num in numbers:
if num in seen:
seen[num] += 1
else:
seen[num] = 1
```
### 3.3 Loops & Dictionaries
- Write a loop which goes through all keys in the dictionary below. If the key is an integer, pass without doing anything. Otherwise, sum the list corresponding to that key and append that sum to the variable answer.
```
prob_seven_list = {'a': [2,4,2], 1: [44,2], 2: [1,2,3], 'b': [0,0,10], 'c': [7]}
answer = []
## Add your code here!
# @title ##### Double click here for a hint!
# `type()` tells you the type of a variable. `pass` tells the loop to do nothing.*
# @title ##### Run this. Have you found a solution?
test_results(answer, [8, 10, 7])
# @title Double click here for the solution
for key, value in prob_seven_list.items():
if type(k) == int:
pass
else:
total = sum(value)
answer.append(total)
```
### 3.4 Operators
- Use relational operators, for loop, and if statements to complete the following problem.
- Given a list of dictionaries that contain students' names as keys and values as the students' list of performance in several subjects, write a program to print a student's name followed by the student's performance score in each subject. See the example below.
```
Operations:
- if score is below 30, print E
- if score is greater than or equals to 30 and less than 50, print D
- if score is greater than or equals to 50 and less than or equals to 70, print C
- if score is greater than 70 and less than 80, print B
- Otherwise, print A
Input: students = [{'py': [23, 45, 56, 67, 89, 90], 'js': [32, 88, 77, 98, 89, 56]}]
Output:
py
23 is E
45 is D
56 is C
67 is C
89 is A
90 is A
js
32 is D
88 is A
77 is B
98 is A
89 is A
56 is C
```
```
csstudents = [('py', [23, 45, 56, 67, 89, 90]), ('js', [32, 88, 77, 98, 89, 56]), ('cpp', [70, 60, 90, 80, 78, 85]), ('lisp', [90, 95, 77, 86, 97, 67, 65])]
# Write your code here!
# @title Double click here for the solution
# solution to be added
for student in csstudents:
name, scores = student
print(name)
print()
for score in scores:
if score < 30:
print(f'{score} is E')
elif score >= 30 and score < 50:
print(f'{score} is D')
elif score >= 50 and score <= 70:
print(f'{score} is C')
elif score > 70 and score < 80:
print(f'{score} is B')
else:
print(f'{score} is A')
```
- Check for parity; parity refers to evenness or oddness of a number
- Given a list of numbers or integers, print the number followed by `is even` if the number is even. Else, print the number followed by `is odd`.
```
numbers = [2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20]
# Write your code here!
# @title Double click here for the solution
# solution to be added
for number in numbers:
if number % 2 == 0:
print(f'{number} is even')
else:
print(f'{number} is odd')
```
---
#Congrats! That's it for this tutorial - and for this course.
## Thank you so much for being part of this adventure - keep up the good work! Hope to welcome you soon again!
---
*This notebook was created by Cameron Smith and Brian Mboya for the [TReND](https://trendinafrica.org/) Python course 2021 and is published under [MIT license](https://choosealicense.com/licenses/mit/).*
| github_jupyter |
# Optimization
- [Least squares](#Least-squares)
- [Gradient descent](#Gradient-descent)
- [Constraint optimization](#Constraint-optimization)
- [Global optimization](#Global-optimization)
Start with describing the problem, like here
- https://scaron.info/blog/linear-programming-in-python-with-cvxopt.html
- https://scaron.info/blog/linear-programming-in-python-with-pulp.html
- http://benalexkeen.com/linear-programming-with-python-and-pulp/
- https://stackoverflow.com/questions/10697995/linear-programming-in-python
- http://isaacslavitt.com/2014/06/20/linear-optimization-in-python/
nonlinear opt course:
- https://github.com/maeehart/TIES483
- https://indico.io/blog/visualizing-with-t-sne/
integer
https://github.com/avivace/dna-recombination/blob/master/script/gurobi_tests.py
## Intro
Biological research uses optimization when performing many types of machine learning, or when it interfaces with engineering. A particular example is metabolic engineering. As a topic in itself, optimization is extremely complex and useful, so much so that it touches to the core of mathematics and computing.
An optimization problem complexity is dependent on several factors, such as:
- Do you intend a local or a global optimization?
- Is the function linear or nonlinear?
- Is the function convex or not?
- Can a gradient be computed?
- Can the Hessian matrix be computed?
- Do we perform optimization under constraints?
- Are those constraints integers?
- Is there a single objective or several?
Scipy does not cover all solvers efficiently but there are several Python packages specialized for certain classes of optimization problems. In general though, the heavier optimization tasks are solved with dedicated programs, many of whom have language bindings for Python.
## Least squares
In practical terms, the most basic application of optimization is computing the local vs global minima of functions. We will exemplify this with the method of least squares. This method is being used to fit the parameters of a function by performing an error minimization.
**Problem context** Having a set of $m$ data points, $(x_1, y_1), (x_2, y_2),\dots,(x_m, y_m)$ and a curve (model function) $y=f(x, \boldsymbol \beta)$ that in addition to the variable $x$ also depends on $n$ parameters, $\boldsymbol \beta = (\beta_1, \beta_2, \dots, \beta_n)$ with $m\ge n$.
It is desired to find the vector $\boldsymbol \beta$ of parameters such that the curve fits best the given data in the least squares sense, that is, the sum of squares of the residuals is minimized:
$$ min \sum_{i=1}^{m}(y_i - f(x_i, \boldsymbol \beta))^2$$
Let us use a similar exercise as the basic linear regression performet in the statistics chapter, but fit a curve instead. That is to say, we are now performing a very basic form of nonlinear regression. While not strictly statistics related, this exercise can be useful for example if we want to decide how a probability distribution fits our data. We will use the least-square again, through the optimization module of scipy.
```
%matplotlib inline
import numpy as np
import pylab as plt
from scipy import optimize
nsamp = 30
x = np.linspace(0,1,nsamp)
"""
y = -0.5*x**2 + 7*sin(x)
This is what we try to fit against. Suppose we know our function is generated
by this law and want to find the (-0.5, 7) parameters. Alternatively we might
not know anything about this dataset but just want to fit this curve to it.
"""
# define the normal function
f = lambda p, x: p[0]*x*x + p[1]*np.sin(x)
testp = (-0.5, 7)
print("True(unknown) parameter value:", testp)
y = f(testp,x)
yr = y + .5*np.random.normal(size=nsamp) # adding a small noise
# define the residual function
e = lambda p, x, y: (abs((f(p,x)-y))).sum()
p0 = (5, 20) # initial parameter value
print("Initial parameter value:", p0)
# uses the standard least squares algorithm
p_est1 = optimize.least_squares(e, p0, args=(x, yr))
print("Parameters estimated with least squares:",p_est1.x)
y_est1 = f(p_est1.x, x)
plt.plot(x,y_est1,'r-', x,yr,'o', x,y,'b-')
plt.show()
# uses a simplex algorithm
p_est2 = optimize.fmin(e, p0, args=(x,yr))
print("Parameters estimated with the simplex algorithm:",p_est2)
y_est2 = f(p_est2, x)
plt.plot(x,y_est2,'r-', x,yr,'o', x,y,'b-')
plt.show()
```
Exercises:
- Use a different nonlinear function.
- Define a normal Python function f() instead!
- Improve the LS fit by using nonstandart loss functions (soft_l1, cauchy)
- Improve the LS fit by using different methods {‘dogbox’, ‘lm’}
## Gradient descent
Note that least squares is not an optimization method per se, it is a method to frame linear regression in the terms of an optimization problem. Gradient descent is the basis optimizatin method for most of modern machine learning, and any processor software today is judged by its speed to compute gradient descent. On GPUs, it sits as the foundation for Deep Learning and Reinforcement Learning.
The method is making an iterative walk in the direction of the local gradient, until the step size becomes :
$$\mathbf{a}_{n+1} = \mathbf{a}_n-\gamma\nabla F(\mathbf{a}_n)$$
Here is the naive algorithm, adapted from Wikipedia:
```
%matplotlib inline
import numpy as np
import pylab as plt
cur_x = 6 # The algorithm starts at x=6
gamma = 0.01 # step size multiplier
precision = 0.00001
previous_step_size = 1/precision; # some large value
f = lambda x: x**4 - 3 * x**3 + 2
df = lambda x: 4 * x**3 - 9 * x**2
x = np.linspace(-4,cur_x,100)
while previous_step_size > precision:
prev_x = cur_x
cur_x += -gamma * df(prev_x)
previous_step_size = abs(cur_x - prev_x)
print("The local minimum occurs at %f" % cur_x)
plt.plot(x,f(x),'b-')
```
The naive implementations suffer from many downsides, from speed to oscilating accross a valey. Another typical fault of the naive approach is assuming that the functions to be optimized are smooth, with expected variation for small variations in their parameters. Here is an example of how to compute the gradient descent with scipy, for the rosenbrock function, a function known to be ill-conditioned.
But before, here is a [practical advice from Scipy](https://www.scipy-lectures.org/advanced/mathematical_optimization/index.html):
- Gradient not known:
> In general, prefer BFGS or L-BFGS, even if you have to approximate numerically gradients. These are also the default if you omit the parameter method - depending if the problem has constraints or bounds. On well-conditioned problems, Powell and Nelder-Mead, both gradient-free methods, work well in high dimension, but they collapse for ill-conditioned problems.
- With knowledge of the gradient:
> BFGS or L-BFGS. Computational overhead of BFGS is larger than that L-BFGS, itself larger than that of conjugate gradient. On the other side, BFGS usually needs less function evaluations than CG. Thus conjugate gradient method is better than BFGS at optimizing computationally cheap functions.
With the Hessian:
- If you can compute the Hessian, prefer the Newton method (Newton-CG or TCG).
- If you have noisy measurements: Use Nelder-Mead or Powell.
```
import numpy as np
import scipy.optimize as optimize
def f(x): # The rosenbrock function
return .5*(1 - x[0])**2 + (x[1] - x[0]**2)**2
def fprime(x):
return np.array((-2*.5*(1 - x[0]) - 4*x[0]*(x[1] - x[0]**2), 2*(x[1] - x[0]**2)))
print(optimize.fmin_ncg(f, [2, 2], fprime=fprime))
def hessian(x): # Computed with sympy
return np.array(((1 - 4*x[1] + 12*x[0]**2, -4*x[0]), (-4*x[0], 2)))
print(optimize.fmin_ncg(f, [2, 2], fprime=fprime, fhess=hessian))
%matplotlib inline
from matplotlib import cm
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
fig = plt.figure()
ax = fig.gca(projection='3d')
X = np.arange(-1, 1, 0.005)
Y = np.arange(-1, 1, 0.005)
X, Y = np.meshgrid(X, Y)
Z = .5*(1 - X)**2 + (Y - X**2)**2
surf = ax.plot_surface(X, Y, Z, rstride=1, cstride=1, cmap=cm.coolwarm,
linewidth=0, antialiased=False)
#ax.set_zlim(-1000.01, 1000.01)
fig.colorbar(surf, shrink=0.5, aspect=5)
plt.show()
```
## Constraint optimization
Problem definition:
$$
\begin{array}{rcll}
\min &~& f(\mathbf{x}) & \\
\mathrm{subject~to} &~& g_i(\mathbf{x}) = c_i &\text{for } i=1,\ldots,n \quad \text{Equality constraints} \\
&~& h_j(\mathbf{x}) \geqq d_j &\text{for } j=1,\ldots,m \quad \text{Inequality constraints}
\end{array}
$$
Let's take the particular case when the objective function and the constraints linear, such as in the cannonical form:
$$\begin{align}
& \text{maximize} && \mathbf{c}^\mathrm{T} \mathbf{x}\\
& \text{subject to} && A \mathbf{x} \leq \mathbf{b} \\
& \text{and} && \mathbf{x} \ge \mathbf{0}
\end{align}$$
Scipy has methods for optimizing functions under constraints, including linear programming. Aditionally many (linear or nonlinear) constraint optimization problems can be turned into full optimization problems using Lagrange multipliers. We will learn how to run linear problems with PuLP.
```
"""
maximize: 4x + 3y
x > 0
y >= 2
x + 2y <= 25
2x - 4y <= 8
-2x + y <= -5
"""
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
x = np.linspace(0, 20, 2000)
y1 = (x*0) + 2
y2 = (25-x)/2.0
y3 = (2*x-8)/4.0
y4 = 2 * x -5
# Make plot
plt.plot(x, y1, label=r'$y\geq2$')
plt.plot(x, y2, label=r'$2y\leq25-x$')
plt.plot(x, y3, label=r'$4y\geq 2x - 8$')
plt.plot(x, y4, label=r'$y\leq 2x-5$')
plt.xlim((0, 16))
plt.ylim((0, 11))
plt.xlabel(r'$x$')
plt.ylabel(r'$y$')
# Fill feasible region
y5 = np.minimum(y2, y4)
y6 = np.maximum(y1, y3)
plt.fill_between(x, y5, y6, where=y5>y6, color='grey', alpha=0.5)
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
```
The solution to an optimization problem with linear constraints lies somewhere on the gray area, and it is not necessarily unique. However for linear optimization functions the solution is on one of the vertices. Now let us frame it with PuLP:
```
conda install -c conda-forge pulp
```
```
import pulp
my_lp_problem = pulp.LpProblem("My LP Problem", pulp.LpMaximize)
x = pulp.LpVariable('x', lowBound=0, cat='Continuous')
y = pulp.LpVariable('y', lowBound=2, cat='Continuous')
# Objective function
my_lp_problem += 4 * x + 3 * y, "Z"
# Constraints
my_lp_problem += 2 * y <= 25 - x
my_lp_problem += 4 * y >= 2 * x - 8
my_lp_problem += y <= 2 * x - 5
my_lp_problem
my_lp_problem.solve()
print(pulp.LpStatus[my_lp_problem.status])
for variable in my_lp_problem.variables():
print("{} = {}".format(variable.name, variable.varValue))
print(pulp.value(my_lp_problem.objective))
```
Further complications arise when some of the constraints need to be integers, in which case the problem becomes known as Mixed Integer Liniar Programming and it computationally more expensive. Yet such problems are quite frequent, for example in metabolic engineering, where you need to deal machines operating on discrete intervals, or when studying protein folding or DNA recombination. In such cases one can also install python packages that deal with the specific problem, such as [cobrapy](https://cobrapy.readthedocs.io/en/latest/) for metabolic engineering.
Further reading:
- For LP with PuLP, I recommend this tutorial, which also uses some real life problems, and has a github link for the notebooks: [http://benalexkeen.com/linear-programming-with-python-and-pulp/](http://benalexkeen.com/linear-programming-with-python-and-pulp/)
- For a great list of LP solvers check [https://stackoverflow.com/questions/26305704/python-mixed-integer-linear-programming](https://stackoverflow.com/questions/26305704/python-mixed-integer-linear-programming)
## Global optimization
The most computation efficient optimization methods can only find local optima, while using different heuristics when needing to access global optima. In the class of global optimization algorithms a few methods to mention are:
- Grid search: These methods belong to the class of brute force or greedy searches and check for solutions in multidimensional sollution spaces. Typicall employed when having to find optimal parameter combinations for machine learning problems (hyperparametrization)
- Branch and Bound: This method, belonging to the more general class called dynamical programming, uses an optimal rooted tree, thus breaking the problem into smaller local optima problems. It is used in LP/MILP for example, or by sequence alignment programs such as BLAST.
- Monte Carlo: This method belongs to the stochastic optimization class, that instead of looking for an exact fit is using random sampling and bayesian statistics. These methods are expected to gain more traction with the evolution of computing power.
- Heuristics: Many methods in this class are nature inspired, such as genetic programming, ant colony optimization etc.
```
import sys
sys.version
sys.path
```
| github_jupyter |
# Batch Normalization
One way to make deep networks easier to train is to use more sophisticated optimization procedures such as SGD+momentum, RMSProp, or Adam. Another strategy is to change the architecture of the network to make it easier to train.
One idea along these lines is batch normalization which was proposed by [3] in 2015.
The idea is relatively straightforward. Machine learning methods tend to work better when their input data consists of uncorrelated features with zero mean and unit variance. When training a neural network, we can preprocess the data before feeding it to the network to explicitly decorrelate its features; this will ensure that the first layer of the network sees data that follows a nice distribution. However, even if we preprocess the input data, the activations at deeper layers of the network will likely no longer be decorrelated and will no longer have zero mean or unit variance since they are output from earlier layers in the network. Even worse, during the training process the distribution of features at each layer of the network will shift as the weights of each layer are updated.
The authors of [3] hypothesize that the shifting distribution of features inside deep neural networks may make training deep networks more difficult. To overcome this problem, [3] proposes to insert batch normalization layers into the network. At training time, a batch normalization layer uses a minibatch of data to estimate the mean and standard deviation of each feature. These estimated means and standard deviations are then used to center and normalize the features of the minibatch. A running average of these means and standard deviations is kept during training, and at test time these running averages are used to center and normalize features.
It is possible that this normalization strategy could reduce the representational power of the network, since it may sometimes be optimal for certain layers to have features that are not zero-mean or unit variance. To this end, the batch normalization layer includes learnable shift and scale parameters for each feature dimension.
[3] [Sergey Ioffe and Christian Szegedy, "Batch Normalization: Accelerating Deep Network Training by Reducing
Internal Covariate Shift", ICML 2015.](https://arxiv.org/abs/1502.03167)
```
# As usual, a bit of setup
import time
import numpy as np
import matplotlib.pyplot as plt
from cs682.classifiers.fc_net import *
from cs682.data_utils import get_CIFAR10_data
from cs682.gradient_check import eval_numerical_gradient, eval_numerical_gradient_array
from cs682.solver import Solver
%matplotlib inline
plt.rcParams['figure.figsize'] = (10.0, 8.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
# for auto-reloading external modules
# see http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
%load_ext autoreload
%autoreload 2
def rel_error(x, y):
""" returns relative error """
return np.max(np.abs(x - y) / (np.maximum(1e-8, np.abs(x) + np.abs(y))))
def print_mean_std(x,axis=0):
print(' means: ', x.mean(axis=axis))
print(' stds: ', x.std(axis=axis))
print()
# Load the (preprocessed) CIFAR10 data.
data = get_CIFAR10_data()
for k, v in data.items():
print('%s: ' % k, v.shape)
```
## Batch normalization: forward
In the file `cs682/layers.py`, implement the batch normalization forward pass in the function `batchnorm_forward`. Once you have done so, run the following to test your implementation.
Referencing the paper linked to above would be helpful!
```
# Check the training-time forward pass by checking means and variances
# of features both before and after batch normalization
# Simulate the forward pass for a two-layer network
np.random.seed(231)
N, D1, D2, D3 = 200, 50, 60, 3
X = np.random.randn(N, D1)
W1 = np.random.randn(D1, D2)
W2 = np.random.randn(D2, D3)
a = np.maximum(0, X.dot(W1)).dot(W2)
print('Before batch normalization:')
print_mean_std(a,axis=0)
gamma = np.ones((D3,))
beta = np.zeros((D3,))
# Means should be close to zero and stds close to one
print('After batch normalization (gamma=1, beta=0)')
a_norm, _ = batchnorm_forward(a, gamma, beta, {'mode': 'train'})
print_mean_std(a_norm,axis=0)
gamma = np.asarray([1.0, 2.0, 3.0])
beta = np.asarray([11.0, 12.0, 13.0])
# Now means should be close to beta and stds close to gamma
print('After batch normalization (gamma=', gamma, ', beta=', beta, ')')
a_norm, _ = batchnorm_forward(a, gamma, beta, {'mode': 'train'})
print_mean_std(a_norm,axis=0)
# Check the test-time forward pass by running the training-time
# forward pass many times to warm up the running averages, and then
# checking the means and variances of activations after a test-time
# forward pass.
np.random.seed(231)
N, D1, D2, D3 = 200, 50, 60, 3
W1 = np.random.randn(D1, D2)
W2 = np.random.randn(D2, D3)
bn_param = {'mode': 'train'}
gamma = np.ones(D3)
beta = np.zeros(D3)
for t in range(50):
X = np.random.randn(N, D1)
a = np.maximum(0, X.dot(W1)).dot(W2)
batchnorm_forward(a, gamma, beta, bn_param)
bn_param['mode'] = 'test'
X = np.random.randn(N, D1)
a = np.maximum(0, X.dot(W1)).dot(W2)
a_norm, _ = batchnorm_forward(a, gamma, beta, bn_param)
# Means should be close to zero and stds close to one, but will be
# noisier than training-time forward passes.
print('After batch normalization (test-time):')
print_mean_std(a_norm,axis=0)
```
## Batch normalization: backward
Now implement the backward pass for batch normalization in the function `batchnorm_backward`.
To derive the backward pass you should write out the computation graph for batch normalization and backprop through each of the intermediate nodes. Some intermediates may have multiple outgoing branches; make sure to sum gradients across these branches in the backward pass.
Once you have finished, run the following to numerically check your backward pass.
```
# Gradient check batchnorm backward pass
np.random.seed(231)
N, D = 4, 5
x = 5 * np.random.randn(N, D) + 12
gamma = np.random.randn(D)
beta = np.random.randn(D)
dout = np.random.randn(N, D)
bn_param = {'mode': 'train'}
fx = lambda x: batchnorm_forward(x, gamma, beta, bn_param)[0]
fg = lambda a: batchnorm_forward(x, a, beta, bn_param)[0]
fb = lambda b: batchnorm_forward(x, gamma, b, bn_param)[0]
dx_num = eval_numerical_gradient_array(fx, x, dout)
da_num = eval_numerical_gradient_array(fg, gamma.copy(), dout)
db_num = eval_numerical_gradient_array(fb, beta.copy(), dout)
_, cache = batchnorm_forward(x, gamma, beta, bn_param)
dx, dgamma, dbeta = batchnorm_backward(dout, cache)
#You should expect to see relative errors between 1e-13 and 1e-8
print('dx error: ', rel_error(dx_num, dx))
print('dgamma error: ', rel_error(da_num, dgamma))
print('dbeta error: ', rel_error(db_num, dbeta))
```
## Batch normalization: alternative backward
In class we talked about two different implementations for the sigmoid backward pass. One strategy is to write out a computation graph composed of simple operations and backprop through all intermediate values. Another strategy is to work out the derivatives on paper. For example, you can derive a very simple formula for the sigmoid function's backward pass by simplifying gradients on paper.
Surprisingly, it turns out that you can do a similar simplification for the batch normalization backward pass too.
Given a set of inputs $X=\begin{bmatrix}x_1\\x_2\\...\\x_N\end{bmatrix}$,
we first calculate the mean $\mu=\frac{1}{N}\sum_{k=1}^N x_k$ and variance $v=\frac{1}{N}\sum_{k=1}^N (x_k-\mu)^2.$
With $\mu$ and $v$ calculated, we can calculate the standard deviation $\sigma=\sqrt{v+\epsilon}$ and normalized data $Y$ with $y_i=\frac{x_i-\mu}{\sigma}.$
The meat of our problem is to get $\frac{\partial L}{\partial X}$ from the upstream gradient $\frac{\partial L}{\partial Y}.$ It might be challenging to directly reason about the gradients over $X$ and $Y$ - try reasoning about it in terms of $x_i$ and $y_i$ first.
You will need to come up with the derivations for $\frac{\partial L}{\partial x_i}$, by relying on the Chain Rule to first calculate the intermediate $\frac{\partial \mu}{\partial x_i}, \frac{\partial v}{\partial x_i}, \frac{\partial \sigma}{\partial x_i},$ then assemble these pieces to calculate $\frac{\partial y_i}{\partial x_i}$. You should make sure each of the intermediary steps are all as simple as possible.
After doing so, implement the simplified batch normalization backward pass in the function `batchnorm_backward_alt` and compare the two implementations by running the following. Your two implementations should compute nearly identical results, but the alternative implementation should be a bit faster.
```
np.random.seed(231)
N, D = 100, 500
x = 5 * np.random.randn(N, D) + 12
gamma = np.random.randn(D)
beta = np.random.randn(D)
dout = np.random.randn(N, D)
bn_param = {'mode': 'train'}
out, cache = batchnorm_forward(x, gamma, beta, bn_param)
t1 = time.time()
dx1, dgamma1, dbeta1 = batchnorm_backward(dout, cache)
t2 = time.time()
dx2, dgamma2, dbeta2 = batchnorm_backward_alt(dout, cache)
t3 = time.time()
print('dx difference: ', rel_error(dx1, dx2))
print('dgamma difference: ', rel_error(dgamma1, dgamma2))
print('dbeta difference: ', rel_error(dbeta1, dbeta2))
print('speedup: %.2fx' % ((t2 - t1) / (t3 - t2)))
```
## Fully Connected Nets with Batch Normalization
Now that you have a working implementation for batch normalization, go back to your `FullyConnectedNet` in the file `cs682/classifiers/fc_net.py`. Modify your implementation to add batch normalization.
Concretely, when the `normalization` flag is set to `"batchnorm"` in the constructor, you should insert a batch normalization layer before each ReLU nonlinearity. The outputs from the last layer of the network should not be normalized. Once you are done, run the following to gradient-check your implementation.
HINT: You might find it useful to define an additional helper layer similar to those in the file `cs682/layer_utils.py`. If you decide to do so, do it in the file `cs682/classifiers/fc_net.py`.
```
np.random.seed(231)
N, D, H1, H2, C = 2, 15, 20, 30, 10
X = np.random.randn(N, D)
y = np.random.randint(C, size=(N,))
# You should expect losses between 1e-4~1e-10 for W,
# losses between 1e-08~1e-10 for b,
# and losses between 1e-08~1e-09 for beta and gammas.
for reg in [0, 3.14]:
print('Running check with reg = ', reg)
model = FullyConnectedNet([H1, H2], input_dim=D, num_classes=C,
reg=reg, weight_scale=5e-2, dtype=np.float64,
normalization='batchnorm')
loss, grads = model.loss(X, y)
print('Initial loss: ', loss)
for name in sorted(grads):
f = lambda _: model.loss(X, y)[0]
grad_num = eval_numerical_gradient(f, model.params[name], verbose=False, h=1e-5)
print('%s relative error: %.2e' % (name, rel_error(grad_num, grads[name])))
if reg == 0: print()
```
# Batchnorm for deep networks
Run the following to train a six-layer network on a subset of 1000 training examples both with and without batch normalization.
```
np.random.seed(231)
# Try training a very deep net with batchnorm
hidden_dims = [100, 100, 100, 100, 100]
num_train = 1000
small_data = {
'X_train': data['X_train'][:num_train],
'y_train': data['y_train'][:num_train],
'X_val': data['X_val'],
'y_val': data['y_val'],
}
weight_scale = 2e-2
bn_model = FullyConnectedNet(hidden_dims, weight_scale=weight_scale, normalization='batchnorm')
model = FullyConnectedNet(hidden_dims, weight_scale=weight_scale, normalization=None)
bn_solver = Solver(bn_model, small_data,
num_epochs=10, batch_size=50,
update_rule='adam',
optim_config={
'learning_rate': 1e-3,
},
verbose=True,print_every=20)
bn_solver.train()
solver = Solver(model, small_data,
num_epochs=10, batch_size=50,
update_rule='adam',
optim_config={
'learning_rate': 1e-3,
},
verbose=True, print_every=20)
solver.train()
```
Run the following to visualize the results from two networks trained above. You should find that using batch normalization helps the network to converge much faster.
```
def plot_training_history(title, label, baseline, bn_solvers, plot_fn, bl_marker='.', bn_marker='.', labels=None):
"""utility function for plotting training history"""
plt.title(title)
plt.xlabel(label)
bn_plots = [plot_fn(bn_solver) for bn_solver in bn_solvers]
bl_plot = plot_fn(baseline)
num_bn = len(bn_plots)
for i in range(num_bn):
label='with_norm'
if labels is not None:
label += str(labels[i])
plt.plot(bn_plots[i], bn_marker, label=label)
label='baseline'
if labels is not None:
label += str(labels[0])
plt.plot(bl_plot, bl_marker, label=label)
plt.legend(loc='lower center', ncol=num_bn+1)
plt.subplot(3, 1, 1)
plot_training_history('Training loss','Iteration', solver, [bn_solver], \
lambda x: x.loss_history, bl_marker='o', bn_marker='o')
plt.subplot(3, 1, 2)
plot_training_history('Training accuracy','Epoch', solver, [bn_solver], \
lambda x: x.train_acc_history, bl_marker='-o', bn_marker='-o')
plt.subplot(3, 1, 3)
plot_training_history('Validation accuracy','Epoch', solver, [bn_solver], \
lambda x: x.val_acc_history, bl_marker='-o', bn_marker='-o')
plt.gcf().set_size_inches(15, 15)
plt.show()
```
# Batch normalization and initialization
We will now run a small experiment to study the interaction of batch normalization and weight initialization.
The first cell will train 8-layer networks both with and without batch normalization using different scales for weight initialization. The second layer will plot training accuracy, validation set accuracy, and training loss as a function of the weight initialization scale.
```
np.random.seed(231)
# Try training a very deep net with batchnorm
hidden_dims = [50, 50, 50, 50, 50, 50, 50]
num_train = 1000
small_data = {
'X_train': data['X_train'][:num_train],
'y_train': data['y_train'][:num_train],
'X_val': data['X_val'],
'y_val': data['y_val'],
}
bn_solvers_ws = {}
solvers_ws = {}
weight_scales = np.logspace(-4, 0, num=20)
for i, weight_scale in enumerate(weight_scales):
print('Running weight scale %d / %d' % (i + 1, len(weight_scales)))
bn_model = FullyConnectedNet(hidden_dims, weight_scale=weight_scale, normalization='batchnorm')
model = FullyConnectedNet(hidden_dims, weight_scale=weight_scale, normalization=None)
bn_solver = Solver(bn_model, small_data,
num_epochs=10, batch_size=50,
update_rule='adam',
optim_config={
'learning_rate': 1e-3,
},
verbose=False, print_every=200)
bn_solver.train()
bn_solvers_ws[weight_scale] = bn_solver
solver = Solver(model, small_data,
num_epochs=10, batch_size=50,
update_rule='adam',
optim_config={
'learning_rate': 1e-3,
},
verbose=False, print_every=200)
solver.train()
solvers_ws[weight_scale] = solver
# Plot results of weight scale experiment
best_train_accs, bn_best_train_accs = [], []
best_val_accs, bn_best_val_accs = [], []
final_train_loss, bn_final_train_loss = [], []
for ws in weight_scales:
best_train_accs.append(max(solvers_ws[ws].train_acc_history))
bn_best_train_accs.append(max(bn_solvers_ws[ws].train_acc_history))
best_val_accs.append(max(solvers_ws[ws].val_acc_history))
bn_best_val_accs.append(max(bn_solvers_ws[ws].val_acc_history))
final_train_loss.append(np.mean(solvers_ws[ws].loss_history[-100:]))
bn_final_train_loss.append(np.mean(bn_solvers_ws[ws].loss_history[-100:]))
plt.subplot(3, 1, 1)
plt.title('Best val accuracy vs weight initialization scale')
plt.xlabel('Weight initialization scale')
plt.ylabel('Best val accuracy')
plt.semilogx(weight_scales, best_val_accs, '-o', label='baseline')
plt.semilogx(weight_scales, bn_best_val_accs, '-o', label='batchnorm')
plt.legend(ncol=2, loc='lower right')
plt.subplot(3, 1, 2)
plt.title('Best train accuracy vs weight initialization scale')
plt.xlabel('Weight initialization scale')
plt.ylabel('Best training accuracy')
plt.semilogx(weight_scales, best_train_accs, '-o', label='baseline')
plt.semilogx(weight_scales, bn_best_train_accs, '-o', label='batchnorm')
plt.legend()
plt.subplot(3, 1, 3)
plt.title('Final training loss vs weight initialization scale')
plt.xlabel('Weight initialization scale')
plt.ylabel('Final training loss')
plt.semilogx(weight_scales, final_train_loss, '-o', label='baseline')
plt.semilogx(weight_scales, bn_final_train_loss, '-o', label='batchnorm')
plt.legend()
plt.gca().set_ylim(1.0, 3.5)
plt.gcf().set_size_inches(15, 15)
plt.show()
```
## Inline Question 1:
Describe the results of this experiment. How does the scale of weight initialization affect models with/without batch normalization differently, and why?
## Answer:
When without batch normalization, in deeper networks, it is easier to have the weights blow up.
But when with batch normalization, each layer’s inputs are normalized so the weights would stay more stable.
# Batch normalization and batch size
We will now run a small experiment to study the interaction of batch normalization and batch size.
The first cell will train 6-layer networks both with and without batch normalization using different batch sizes. The second layer will plot training accuracy and validation set accuracy over time.
```
def run_batchsize_experiments(normalization_mode):
np.random.seed(231)
# Try training a very deep net with batchnorm
hidden_dims = [100, 100, 100, 100, 100]
num_train = 1000
small_data = {
'X_train': data['X_train'][:num_train],
'y_train': data['y_train'][:num_train],
'X_val': data['X_val'],
'y_val': data['y_val'],
}
n_epochs=10
weight_scale = 2e-2
batch_sizes = [5,10,50]
lr = 10**(-3.5)
solver_bsize = batch_sizes[0]
print('No normalization: batch size = ',solver_bsize)
model = FullyConnectedNet(hidden_dims, weight_scale=weight_scale, normalization=None)
solver = Solver(model, small_data,
num_epochs=n_epochs, batch_size=solver_bsize,
update_rule='adam',
optim_config={
'learning_rate': lr,
},
verbose=False)
solver.train()
bn_solvers = []
for i in range(len(batch_sizes)):
b_size=batch_sizes[i]
print('Normalization: batch size = ',b_size)
bn_model = FullyConnectedNet(hidden_dims, weight_scale=weight_scale, normalization=normalization_mode)
bn_solver = Solver(bn_model, small_data,
num_epochs=n_epochs, batch_size=b_size,
update_rule='adam',
optim_config={
'learning_rate': lr,
},
verbose=False)
bn_solver.train()
bn_solvers.append(bn_solver)
return bn_solvers, solver, batch_sizes
batch_sizes = [5,10,50]
bn_solvers_bsize, solver_bsize, batch_sizes = run_batchsize_experiments('batchnorm')
plt.subplot(2, 1, 1)
plot_training_history('Training accuracy (Batch Normalization)','Epoch', solver_bsize, bn_solvers_bsize, \
lambda x: x.train_acc_history, bl_marker='-^', bn_marker='-o', labels=batch_sizes)
plt.subplot(2, 1, 2)
plot_training_history('Validation accuracy (Batch Normalization)','Epoch', solver_bsize, bn_solvers_bsize, \
lambda x: x.val_acc_history, bl_marker='-^', bn_marker='-o', labels=batch_sizes)
plt.gcf().set_size_inches(15, 10)
plt.show()
```
## Inline Question 2:
Describe the results of this experiment. What does this imply about the relationship between batch normalization and batch size? Why is this relationship observed?
## Answer:
Our experiment shows that, with larger batch size, the accuracy would be higher with batchnorm, this is because when using batchnorm, if the batch size is too small, we are getting a worse representation for the data.
# Layer Normalization
Batch normalization has proved to be effective in making networks easier to train, but the dependency on batch size makes it less useful in complex networks which have a cap on the input batch size due to hardware limitations.
Several alternatives to batch normalization have been proposed to mitigate this problem; one such technique is Layer Normalization [4]. Instead of normalizing over the batch, we normalize over the features. In other words, when using Layer Normalization, each feature vector corresponding to a single datapoint is normalized based on the sum of all terms within that feature vector.
[4] [Ba, Jimmy Lei, Jamie Ryan Kiros, and Geoffrey E. Hinton. "Layer Normalization." stat 1050 (2016): 21.](https://arxiv.org/pdf/1607.06450.pdf)
## Inline Question 3:
Which of these data preprocessing steps is analogous to batch normalization, and which is analogous to layer normalization?
1. Scaling each image in the dataset, so that the RGB channels for each row of pixels within an image sums up to 1.
2. Scaling each image in the dataset, so that the RGB channels for all pixels within an image sums up to 1.
3. Subtracting the mean image of the dataset from each image in the dataset.
4. Setting all RGB values to either 0 or 1 depending on a given threshold.
## Answer:
Batch normalization: 3
Layer normalization: 2
# Layer Normalization: Implementation
Now you'll implement layer normalization. This step should be relatively straightforward, as conceptually the implementation is almost identical to that of batch normalization. One significant difference though is that for layer normalization, we do not keep track of the moving moments, and the testing phase is identical to the training phase, where the mean and variance are directly calculated per datapoint.
Here's what you need to do:
* In `cs682/layers.py`, implement the forward pass for layer normalization in the function `layernorm_backward`.
Run the cell below to check your results.
* In `cs682/layers.py`, implement the backward pass for layer normalization in the function `layernorm_backward`.
Run the second cell below to check your results.
* Modify `cs682/classifiers/fc_net.py` to add layer normalization to the `FullyConnectedNet`. When the `normalization` flag is set to `"layernorm"` in the constructor, you should insert a layer normalization layer before each ReLU nonlinearity.
Run the third cell below to run the batch size experiment on layer normalization.
```
# Check the training-time forward pass by checking means and variances
# of features both before and after layer normalization
# Simulate the forward pass for a two-layer network
np.random.seed(231)
N, D1, D2, D3 =4, 50, 60, 3
X = np.random.randn(N, D1)
W1 = np.random.randn(D1, D2)
W2 = np.random.randn(D2, D3)
a = np.maximum(0, X.dot(W1)).dot(W2)
print('Before layer normalization:')
print_mean_std(a,axis=1)
gamma = np.ones(D3)
beta = np.zeros(D3)
# Means should be close to zero and stds close to one
print('After layer normalization (gamma=1, beta=0)')
a_norm, _ = layernorm_forward(a, gamma, beta, {'mode': 'train'})
print_mean_std(a_norm,axis=1)
gamma = np.asarray([3.0,3.0,3.0])
beta = np.asarray([5.0,5.0,5.0])
# Now means should be close to beta and stds close to gamma
print('After layer normalization (gamma=', gamma, ', beta=', beta, ')')
a_norm, _ = layernorm_forward(a, gamma, beta, {'mode': 'train'})
print_mean_std(a_norm,axis=1)
# Gradient check batchnorm backward pass
np.random.seed(231)
N, D = 4, 5
x = 5 * np.random.randn(N, D) + 12
gamma = np.random.randn(D)
beta = np.random.randn(D)
dout = np.random.randn(N, D)
ln_param = {}
fx = lambda x: layernorm_forward(x, gamma, beta, ln_param)[0]
fg = lambda a: layernorm_forward(x, a, beta, ln_param)[0]
fb = lambda b: layernorm_forward(x, gamma, b, ln_param)[0]
dx_num = eval_numerical_gradient_array(fx, x, dout)
da_num = eval_numerical_gradient_array(fg, gamma.copy(), dout)
db_num = eval_numerical_gradient_array(fb, beta.copy(), dout)
_, cache = layernorm_forward(x, gamma, beta, ln_param)
dx, dgamma, dbeta = layernorm_backward(dout, cache)
#You should expect to see relative errors between 1e-12 and 1e-8
print('dx error: ', rel_error(dx_num, dx))
print('dgamma error: ', rel_error(da_num, dgamma))
print('dbeta error: ', rel_error(db_num, dbeta))
```
# Layer Normalization and batch size
We will now run the previous batch size experiment with layer normalization instead of batch normalization. Compared to the previous experiment, you should see a markedly smaller influence of batch size on the training history!
```
ln_solvers_bsize, solver_bsize, batch_sizes = run_batchsize_experiments('layernorm')
plt.subplot(2, 1, 1)
plot_training_history('Training accuracy (Layer Normalization)','Epoch', solver_bsize, ln_solvers_bsize, \
lambda x: x.train_acc_history, bl_marker='-^', bn_marker='-o', labels=batch_sizes)
plt.subplot(2, 1, 2)
plot_training_history('Validation accuracy (Layer Normalization)','Epoch', solver_bsize, ln_solvers_bsize, \
lambda x: x.val_acc_history, bl_marker='-^', bn_marker='-o', labels=batch_sizes)
plt.gcf().set_size_inches(15, 10)
plt.show()
```
## Inline Question 4:
When is layer normalization likely to not work well, and why?
1. Using it in a very deep network
2. Having a very small dimension of features
3. Having a high regularization term
## Answer:
Having a very small dimension of features, this is the same as when using a batchnorm and having a very small batch size, you are getting a worse representation of the data.
| github_jupyter |
```
import pandas as pd
import numpy as np
import regex as re
import nltk
nltk.download('wordnet')
nltk.download('punkt')
nltk.download('stopwords')
from nltk.corpus import stopwords
import tensorflow as tf
from tensorflow.python.keras.preprocessing import sequence
from tensorflow.python.keras.preprocessing import text
from sklearn.model_selection import train_test_split
from keras.models import Sequential
from keras.layers import Dense, Embedding, LSTM,Bidirectional, GlobalMaxPooling1D,Dropout,Flatten,Conv1D,MaxPooling1D
from keras.models import load_model
from keras.callbacks import ModelCheckpoint
data = pd.read_csv('version-4.csv',index_col=0)
data.reset_index(drop=True,inplace = True)
for i,j in enumerate(data['labels']):
try:
j = int(j)
except ValueError as e:
print(f'error on {i} line')
data.drop(labels=[2478],inplace= True)
data['labels'].astype(int);
data.dropna(inplace = True)
cleaned_text = []
for text in data['texts']:
text = " ".join(word for word in text.split() if not word.isdigit())
cleaned_text.append(text)
data['cleaned_text'] = cleaned_text
vocab = {}
for text in data['cleaned_text']:
sen = text.split()
for word in sen:
try:
vocab[word] += 1
except KeyError:
vocab[word] = 1
vocab = dict(sorted(vocab.items(), key=lambda item: item[1]))
rare_words = []
for key,value in vocab.items():
if value<=10:
rare_words.append(key)
stopwords_en = set(stopwords.words('english'))
cleaner_text = []
for text in data['cleaned_text']:
text = " ".join([word for word in text.split() if len(word)>2 and word not in rare_words])
cleaner_text.append(text)
data['final_text'] = cleaner_text
vocab = {}
for text in data['final_text']:
sen = text.split()
for word in sen:
try:
vocab[word] += 1
except KeyError:
vocab[word] = 1
vocab = dict(sorted(vocab.items(), key=lambda item: item[1]))
vocab_list = list(vocab.items())
vocab_size = len(vocab)
x = data['final_text'].values
y = data['labels'].values
X_train,X_test,y_train, y_test = train_test_split(x,y, test_size = 0.2, shuffle = True)
y_train = y_train.astype(int)
y_test = y_test.astype(int)
embeddings_index = dict()
f = open('glove.twitter.27B.200d.txt')
for line in f:
values = line.split()
word = values[0]
try:
coefs = np.asarray(values[1:], dtype='float32')
except ValueError:
pass
embeddings_index[word] = coefs
f.close()
print('Loaded %s word vectors.' % len(embeddings_index))
from tensorflow.python.keras.preprocessing import sequence
from tensorflow.python.keras.preprocessing import text
tokenizer = text.Tokenizer()
tokenizer.fit_on_texts(X_train)
X_train = tokenizer.texts_to_sequences(X_train)
X_test = tokenizer.texts_to_sequences(X_test)
X_train = sequence.pad_sequences(X_train,maxlen=200)
X_test = sequence.pad_sequences(X_test,maxlen=200)
tokens = len(tokenizer.word_index) + 2
embedding_matrix = np.zeros((tokens, 200))
count = 0
unknown = []
for word, i in tokenizer.word_index.items():
embedding_vector = embeddings_index.get(word)
if embedding_vector is not None:
try:
embedding_matrix[i] = embedding_vector
except ValueError:
unknown.append(word)
count += 1
else:
unknown.append(word)
count += 1
print(1-(count/vocab_size))
model = Sequential()
model.add(Embedding(tokens,200,weights = [embedding_matrix],input_length = embedding_matrix.shape[1]))
model.add(LSTM(100))
model.add(Dropout(0.2))
model.add(Flatten())
model.add(Dense(1,activation='sigmoid'))
model.compile(optimizer='adam',loss = 'binary_crossentropy',metrics=['accuracy'])
print(model.summary())
model.fit(X_train,y_train,epochs=30)
loss,accuracy = model.evaluate(X_train,y_train)
print(f'acc: {accuracy}')
predictions = model.predict(X_test)
predictions = np.round(predictions)
from sklearn.metrics import accuracy_score, confusion_matrix
score = accuracy_score(y_test,predictions)
cm = confusion_matrix(y_test,predictions)
print("score: {} cm: {}".format(score,cm))
```
| github_jupyter |
# Skip-gram Word2Vec
In this notebook, I'll lead you through using PyTorch to implement the [Word2Vec algorithm](https://en.wikipedia.org/wiki/Word2vec) using the skip-gram architecture. By implementing this, you'll learn about embedding words for use in natural language processing. This will come in handy when dealing with things like machine translation.
## Readings
Here are the resources I used to build this notebook. I suggest reading these either beforehand or while you're working on this material.
* A really good [conceptual overview](http://mccormickml.com/2016/04/19/word2vec-tutorial-the-skip-gram-model/) of Word2Vec from Chris McCormick
* [First Word2Vec paper](https://arxiv.org/pdf/1301.3781.pdf) from Mikolov et al.
* [Neural Information Processing Systems, paper](http://papers.nips.cc/paper/5021-distributed-representations-of-words-and-phrases-and-their-compositionality.pdf) with improvements for Word2Vec also from Mikolov et al.
---
## Word embeddings
When you're dealing with words in text, you end up with tens of thousands of word classes to analyze; one for each word in a vocabulary. Trying to one-hot encode these words is massively inefficient because most values in a one-hot vector will be set to zero. So, the matrix multiplication that happens in between a one-hot input vector and a first, hidden layer will result in mostly zero-valued hidden outputs.
<img src='assets/one_hot_encoding.png' width=50%>
To solve this problem and greatly increase the efficiency of our networks, we use what are called **embeddings**. Embeddings are just a fully connected layer like you've seen before. We call this layer the embedding layer and the weights are embedding weights. We skip the multiplication into the embedding layer by instead directly grabbing the hidden layer values from the weight matrix. We can do this because the multiplication of a one-hot encoded vector with a matrix returns the row of the matrix corresponding the index of the "on" input unit.
<img src='assets/lookup_matrix.png' width=50%>
Instead of doing the matrix multiplication, we use the weight matrix as a lookup table. We encode the words as integers, for example "heart" is encoded as 958, "mind" as 18094. Then to get hidden layer values for "heart", you just take the 958th row of the embedding matrix. This process is called an **embedding lookup** and the number of hidden units is the **embedding dimension**.
<img src='assets/tokenize_lookup.png' width=50%>
There is nothing magical going on here. The embedding lookup table is just a weight matrix. The embedding layer is just a hidden layer. The lookup is just a shortcut for the matrix multiplication. The lookup table is trained just like any weight matrix.
Embeddings aren't only used for words of course. You can use them for any model where you have a massive number of classes. A particular type of model called **Word2Vec** uses the embedding layer to find vector representations of words that contain semantic meaning.
---
## Word2Vec
The Word2Vec algorithm finds much more efficient representations by finding vectors that represent the words. These vectors also contain semantic information about the words.
<img src="assets/context_drink.png" width=40%>
Words that show up in similar **contexts**, such as "coffee", "tea", and "water" will have vectors near each other. Different words will be further away from one another, and relationships can be represented by distance in vector space.
<img src="assets/vector_distance.png" width=40%>
There are two architectures for implementing Word2Vec:
>* CBOW (Continuous Bag-Of-Words) and
* Skip-gram
<img src="assets/word2vec_architectures.png" width=60%>
In this implementation, we'll be using the **skip-gram architecture** because it performs better than CBOW. Here, we pass in a word and try to predict the words surrounding it in the text. In this way, we can train the network to learn representations for words that show up in similar contexts.
---
## Loading Data
Next, we'll ask you to load in data and place it in the `data` directory
1. Load the [text8 dataset](https://s3.amazonaws.com/video.udacity-data.com/topher/2018/October/5bbe6499_text8/text8.zip); a file of cleaned up *Wikipedia article text* from Matt Mahoney.
2. Place that data in the `data` folder in the home directory.
3. Then you can extract it and delete the archive, zip file to save storage space.
After following these steps, you should have one file in your data directory: `data/text8`.
```
# read in the extracted text file
with open('data/text8') as f:
text = f.read()
# print out the first 100 characters
print(text[:100])
type(text)
```
## Pre-processing
Here I'm fixing up the text to make training easier. This comes from the `utils.py` file. The `preprocess` function does a few things:
>* It converts any punctuation into tokens, so a period is changed to ` <PERIOD> `. In this data set, there aren't any periods, but it will help in other NLP problems.
* It removes all words that show up five or *fewer* times in the dataset. This will greatly reduce issues due to noise in the data and improve the quality of the vector representations.
* It returns a list of words in the text.
This may take a few seconds to run, since our text file is quite large. If you want to write your own functions for this stuff, go for it!
```
import utils
# get list of words
words = utils.preprocess(text)
print(words[:30])
# print some stats about this word data
print("Total words in text: {}".format(len(words)))
print("Unique words: {}".format(len(set(words)))) # `set` removes any duplicate words
```
### Dictionaries
Next, I'm creating two dictionaries to convert words to integers and back again (integers to words). This is again done with a function in the `utils.py` file. `create_lookup_tables` takes in a list of words in a text and returns two dictionaries.
>* The integers are assigned in descending frequency order, so the most frequent word ("the") is given the integer 0 and the next most frequent is 1, and so on.
Once we have our dictionaries, the words are converted to integers and stored in the list `int_words`.
```
vocab_to_int, int_to_vocab = utils.create_lookup_tables(words)
int_words = [vocab_to_int[word] for word in words]
print(int_words[:30])
```
## Subsampling
Words that show up often such as "the", "of", and "for" don't provide much context to the nearby words. If we discard some of them, we can remove some of the noise from our data and in return get faster training and better representations. This process is called subsampling by Mikolov. For each word $w_i$ in the training set, we'll discard it with probability given by
$$ P(w_i) = 1 - \sqrt{\frac{t}{f(w_i)}} $$
where $t$ is a threshold parameter and $f(w_i)$ is the frequency of word $w_i$ in the total dataset.
$$ P(0) = 1 - \sqrt{\frac{1*10^{-5}}{1*10^6/16*10^6}} = 0.98735 $$
I'm going to leave this up to you as an exercise. Check out my solution to see how I did it.
> **Exercise:** Implement subsampling for the words in `int_words`. That is, go through `int_words` and discard each word given the probablility $P(w_i)$ shown above. Note that $P(w_i)$ is the probability that a word is discarded. Assign the subsampled data to `train_words`.
QUIZ
```
1 - ( np.sqrt(0.0001/(700/1000000)) )
from collections import Counter
import random
import numpy as np
threshold = 1e-5
word_counts = Counter(int_words)
print(list(word_counts.items())[0]) # dictionary of int_words, how many times they appear
# discard some frequent words, according to the subsampling equation
# create a new list of words for training
train_words = None
#print(train_words[:30])
len(word_counts)
total_count = len(word_counts)
#see formula above
freq = {word: count/total_count for word, count in word_counts.items()}
p_drop = {word: 1- np.sqrt(threshold / freq[word]) for word in word_counts}
train_words = [word for word in int_words if random.random() < (1 - p_drop[word])]
print(train_words[:30])
```
## Making batches
Now that our data is in good shape, we need to get it into the proper form to pass it into our network. With the skip-gram architecture, for each word in the text, we want to define a surrounding _context_ and grab all the words in a window around that word, with size $C$.
From [Mikolov et al.](https://arxiv.org/pdf/1301.3781.pdf):
"Since the more distant words are usually less related to the current word than those close to it, we give less weight to the distant words by sampling less from those words in our training examples... If we choose $C = 5$, for each training word we will select randomly a number $R$ in range $[ 1: C ]$, and then use $R$ words from history and $R$ words from the future of the current word as correct labels."
> **Exercise:** Implement a function `get_target` that receives a list of words, an index, and a window size, then returns a list of words in the window around the index. Make sure to use the algorithm described above, where you chose a random number of words to from the window.
Say, we have an input and we're interested in the idx=2 token, `741`:
```
[5233, 58, 741, 10571, 27349, 0, 15067, 58112, 3580, 58, 10712]
```
For `R=2`, `get_target` should return a list of four values:
```
[5233, 58, 10571, 27349]
```
```
def get_target(words, idx, window_size=5):
''' Get a list of words in a window around an index. '''
# TODO: implement this function
R = np.random.randint(1, window_size+1) # range
start = idx - R if (idx-R) > 0 else 0
stop = idx + R
target_words = words[start:idx] + words[idx+1:stop+1]
return list(target_words)
# test your code!
# run this cell multiple times to check for random window selection
int_text = [i for i in range(10)]
print('Input: ', int_text)
idx=5 # word index of interest
target = get_target(int_text, idx=idx, window_size=5)
print('Target: ', target) # you should get some indices around the idx
```
### Generating Batches
Here's a generator function that returns batches of input and target data for our model, using the `get_target` function from above. The idea is that it grabs `batch_size` words from a words list. Then for each of those batches, it gets the target words in a window.
```
def get_batches(words, batch_size, window_size=5):
''' Create a generator of word batches as a tuple (inputs, targets) '''
n_batches = len(words)//batch_size
# only full batches
words = words[:n_batches*batch_size]
for idx in range(0, len(words), batch_size):
x, y = [], []
batch = words[idx:idx+batch_size]
for ii in range(len(batch)):
batch_x = batch[ii]
batch_y = get_target(batch, ii, window_size)
y.extend(batch_y)
x.extend([batch_x]*len(batch_y))
yield x, y
int_text = [i for i in range(20)]
x,y = next(get_batches(int_text, batch_size=4, window_size=5))
print('x\n', x)
print('y\n', y)
```
## Building the graph
Below is an approximate diagram of the general structure of our network.
<img src="assets/skip_gram_net_arch.png" width=60%>
>* The input words are passed in as batches of input word tokens.
* This will go into a hidden layer of linear units (our embedding layer).
* Then, finally into a softmax output layer.
We'll use the softmax layer to make a prediction about the context words by sampling, as usual.
The idea here is to train the embedding layer weight matrix to find efficient representations for our words. We can discard the softmax layer because we don't really care about making predictions with this network. We just want the embedding matrix so we can use it in _other_ networks we build using this dataset.
---
## Validation
Here, I'm creating a function that will help us observe our model as it learns. We're going to choose a few common words and few uncommon words. Then, we'll print out the closest words to them using the cosine similarity:
$$
\mathrm{similarity} = \cos(\theta) = \frac{\vec{a} \cdot \vec{b}}{|\vec{a}||\vec{b}|}
$$
We can encode the validation words as vectors $\vec{a}$ using the embedding table, then calculate the similarity with each word vector $\vec{b}$ in the embedding table. With the similarities, we can print out the validation words and words in our embedding table semantically similar to those words. It's a nice way to check that our embedding table is grouping together words with similar semantic meanings.
```
def cosine_similarity(embedding, valid_size=16, valid_window=100, device='cpu'):
""" Returns the cosine similarity of validation words with words in the embedding matrix.
Here, embedding should be a PyTorch embedding module.
"""
# Here we're calculating the cosine similarity between some random words and
# our embedding vectors. With the similarities, we can look at what words are
# close to our random words.
# sim = (a . b) / |a||b|
embed_vectors = embedding.weight
# magnitude of embedding vectors, |b|
magnitudes = embed_vectors.pow(2).sum(dim=1).sqrt().unsqueeze(0)
# pick N words from our ranges (0,window) and (1000,1000+window). lower id implies more frequent
valid_examples = np.array(random.sample(range(valid_window), valid_size//2))
valid_examples = np.append(valid_examples,
random.sample(range(1000,1000+valid_window), valid_size//2))
valid_examples = torch.LongTensor(valid_examples).to(device)
valid_vectors = embedding(valid_examples)
similarities = torch.mm(valid_vectors, embed_vectors.t())/magnitudes
return valid_examples, similarities
```
## SkipGram model
Define and train the SkipGram model.
> You'll need to define an [embedding layer](https://pytorch.org/docs/stable/nn.html#embedding) and a final, softmax output layer.
An Embedding layer takes in a number of inputs, importantly:
* **num_embeddings** – the size of the dictionary of embeddings, or how many rows you'll want in the embedding weight matrix
* **embedding_dim** – the size of each embedding vector; the embedding dimension
```
import torch
from torch import nn
import torch.optim as optim
class SkipGram(nn.Module):
def __init__(self, n_vocab, n_embed):
super().__init__()
self.embed = nn.Embedding(n_vocab, n_embed)
self.output = nn.Linear(n_embed, n_vocab)
self.log_Softmax = nn.LogSoftmax(dim=1)
# complete this SkipGram model
def forward(self, x):
# define the forward behavior
x = self.embed(x)
scores = self.output(x)
log_ps = self.log_Softmax(scores)
return log_ps
```
### Training
Below is our training loop, and I recommend that you train on GPU, if available.
**Note that, because we applied a softmax function to our model output, we are using NLLLoss** as opposed to cross entropy. This is because Softmax in combination with NLLLoss = CrossEntropy loss .
```
# check if GPU is available
device = 'cuda' if torch.cuda.is_available() else 'cpu'
embedding_dim=300 # you can change, if you want
model = SkipGram(len(vocab_to_int), embedding_dim).to(device)
criterion = nn.NLLLoss()
optimizer = optim.Adam(model.parameters(), lr=0.003)
print_every = 500
steps = 0
# i am going to trained for only 2 epochs due to limited GPU access, but we can improve with more epochs definitely in this case
epochs = 2
# train for some number of epochs
for e in range(epochs):
# get input and target batches
for inputs, targets in get_batches(train_words, 512):
steps += 1
inputs, targets = torch.LongTensor(inputs), torch.LongTensor(targets)
inputs, targets = inputs.to(device), targets.to(device)
log_ps = model(inputs)
loss = criterion(log_ps, targets)
optimizer.zero_grad()
loss.backward()
optimizer.step()
if steps % print_every == 0:
# getting examples and similarities
valid_examples, valid_similarities = cosine_similarity(model.embed, device=device)
_, closest_idxs = valid_similarities.topk(6) # topk highest similarities
valid_examples, closest_idxs = valid_examples.to('cpu'), closest_idxs.to('cpu')
for ii, valid_idx in enumerate(valid_examples):
closest_words = [int_to_vocab[idx.item()] for idx in closest_idxs[ii]][1:]
print(int_to_vocab[valid_idx.item()] + " | " + ', '.join(closest_words))
print("...")
```
## Visualizing the word vectors
Below we'll use T-SNE to visualize how our high-dimensional word vectors cluster together. T-SNE is used to project these vectors into two dimensions while preserving local stucture. Check out [this post from Christopher Olah](http://colah.github.io/posts/2014-10-Visualizing-MNIST/) to learn more about T-SNE and other ways to visualize high-dimensional data.
```
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import matplotlib.pyplot as plt
from sklearn.manifold import TSNE
# getting embeddings from the embedding layer of our model, by name
embeddings = model.embed.weight.to('cpu').data.numpy()
viz_words = 600
tsne = TSNE()
embed_tsne = tsne.fit_transform(embeddings[:viz_words, :])
fig, ax = plt.subplots(figsize=(16, 16))
for idx in range(viz_words):
plt.scatter(*embed_tsne[idx, :], color='steelblue')
plt.annotate(int_to_vocab[idx], (embed_tsne[idx, 0], embed_tsne[idx, 1]), alpha=0.7)
```
NOTE: I HAVE ONLY TRAINED IT FOR 2 EPOCHS DUE TO GPU LIMITED ACCESS
| github_jupyter |
# Predicting sentiment from product reviews
The goal of this first notebook is to explore logistic regression and feature engineering with existing GraphLab functions.
In this notebook you will use product review data from Amazon.com to predict whether the sentiments about a product (from its reviews) are positive or negative.
* Use SFrames to do some feature engineering
* Train a logistic regression model to predict the sentiment of product reviews.
* Inspect the weights (coefficients) of a trained logistic regression model.
* Make a prediction (both class and probability) of sentiment for a new product review.
* Given the logistic regression weights, predictors and ground truth labels, write a function to compute the **accuracy** of the model.
* Inspect the coefficients of the logistic regression model and interpret their meanings.
* Compare multiple logistic regression models.
Let's get started!
## Fire up GraphLab Create
Make sure you have the latest version of GraphLab Create.
```
from __future__ import division
import graphlab
import math
import string
```
# Data preparation
We will use a dataset consisting of baby product reviews on Amazon.com.
```
products = graphlab.SFrame('amazon_baby.gl/')
```
Now, let us see a preview of what the dataset looks like.
```
products
```
## Build the word count vector for each review
Let us explore a specific example of a baby product.
```
products[269]
```
Now, we will perform 2 simple data transformations:
1. Remove punctuation using [Python's built-in](https://docs.python.org/2/library/string.html) string functionality.
2. Transform the reviews into word-counts.
**Aside**. In this notebook, we remove all punctuations for the sake of simplicity. A smarter approach to punctuations would preserve phrases such as "I'd", "would've", "hadn't" and so forth. See [this page](https://www.cis.upenn.edu/~treebank/tokenization.html) for an example of smart handling of punctuations.
```
def remove_punctuation(text):
return text.translate(None, string.punctuation)
review_without_punctuation = products['review'].apply(remove_punctuation)
products['word_count'] = graphlab.text_analytics.count_words(review_without_punctuation)
```
Now, let us explore what the sample example above looks like after these 2 transformations. Here, each entry in the **word_count** column is a dictionary where the key is the word and the value is a count of the number of times the word occurs.
```
products[269]['word_count']
```
## Extract sentiments
We will **ignore** all reviews with *rating = 3*, since they tend to have a neutral sentiment.
```
products = products[products['rating'] != 3]
len(products)
```
Now, we will assign reviews with a rating of 4 or higher to be *positive* reviews, while the ones with rating of 2 or lower are *negative*. For the sentiment column, we use +1 for the positive class label and -1 for the negative class label.
```
products['sentiment'] = products['rating'].apply(lambda rating : +1 if rating > 3 else -1)
products
```
Now, we can see that the dataset contains an extra column called **sentiment** which is either positive (+1) or negative (-1).
## Split data into training and test sets
Let's perform a train/test split with 80% of the data in the training set and 20% of the data in the test set. We use `seed=1` so that everyone gets the same result.
```
train_data, test_data = products.random_split(.8, seed=1)
print len(train_data)
print len(test_data)
```
# Train a sentiment classifier with logistic regression
We will now use logistic regression to create a sentiment classifier on the training data. This model will use the column **word_count** as a feature and the column **sentiment** as the target. We will use `validation_set=None` to obtain same results as everyone else.
**Note:** This line may take 1-2 minutes.
```
sentiment_model = graphlab.logistic_classifier.create(train_data,
target = 'sentiment',
features=['word_count'],
validation_set=None)
sentiment_model
```
**Aside**. You may get a warning to the effect of "Terminated due to numerical difficulties --- this model may not be ideal". It means that the quality metric (to be covered in Module 3) failed to improve in the last iteration of the run. The difficulty arises as the sentiment model puts too much weight on extremely rare words. A way to rectify this is to apply regularization, to be covered in Module 4. Regularization lessens the effect of extremely rare words. For the purpose of this assignment, however, please proceed with the model above.
Now that we have fitted the model, we can extract the weights (coefficients) as an SFrame as follows:
```
weights = sentiment_model.coefficients
weights.column_names()
```
There are a total of `121713` coefficients in the model. Recall from the lecture that positive weights $w_j$ correspond to weights that cause positive sentiment, while negative weights correspond to negative sentiment.
Fill in the following block of code to calculate how many *weights* are positive ( >= 0). (**Hint**: The `'value'` column in SFrame *weights* must be positive ( >= 0)).
```
num_positive_weights = (weights['value'] >= 0).sum()
num_negative_weights = (weights['value'] < 0).sum()
print "Number of positive weights: %s " % num_positive_weights
print "Number of negative weights: %s " % num_negative_weights
```
**Quiz Question:** How many weights are >= 0?
## Making predictions with logistic regression
Now that a model is trained, we can make predictions on the **test data**. In this section, we will explore this in the context of 3 examples in the test dataset. We refer to this set of 3 examples as the **sample_test_data**.
```
sample_test_data = test_data[10:13]
print sample_test_data['rating']
sample_test_data
```
Let's dig deeper into the first row of the **sample_test_data**. Here's the full review:
```
sample_test_data[0]['review']
```
That review seems pretty positive.
Now, let's see what the next row of the **sample_test_data** looks like. As we could guess from the sentiment (-1), the review is quite negative.
```
sample_test_data[1]['review']
```
We will now make a **class** prediction for the **sample_test_data**. The `sentiment_model` should predict **+1** if the sentiment is positive and **-1** if the sentiment is negative. Recall from the lecture that the **score** (sometimes called **margin**) for the logistic regression model is defined as:
$$
\mbox{score}_i = \mathbf{w}^T h(\mathbf{x}_i)
$$
where $h(\mathbf{x}_i)$ represents the features for example $i$. We will write some code to obtain the **scores** using GraphLab Create. For each row, the **score** (or margin) is a number in the range **[-inf, inf]**.
```
scores = sentiment_model.predict(sample_test_data, output_type='margin')
print scores
```
### Predicting sentiment
These scores can be used to make class predictions as follows:
$$
\hat{y} =
\left\{
\begin{array}{ll}
+1 & \mathbf{w}^T h(\mathbf{x}_i) > 0 \\
-1 & \mathbf{w}^T h(\mathbf{x}_i) \leq 0 \\
\end{array}
\right.
$$
Using scores, write code to calculate $\hat{y}$, the class predictions:
```
def calc_prediction (scores) :
predictions = []
for score in scores:
if score > 0 :
predict = 1
else:
predict = -1
predictions.append(predict)
return predictions
calc_prediction(scores)
```
Run the following code to verify that the class predictions obtained by your calculations are the same as that obtained from GraphLab Create.
```
print "Class predictions according to GraphLab Create:"
print sentiment_model.predict(sample_test_data)
```
**Checkpoint**: Make sure your class predictions match with the one obtained from GraphLab Create.
### Probability predictions
Recall from the lectures that we can also calculate the probability predictions from the scores using:
$$
P(y_i = +1 | \mathbf{x}_i,\mathbf{w}) = \frac{1}{1 + \exp(-\mathbf{w}^T h(\mathbf{x}_i))}.
$$
Using the variable **scores** calculated previously, write code to calculate the probability that a sentiment is positive using the above formula. For each row, the probabilities should be a number in the range **[0, 1]**.
```
import numpy as np
props = 1/(1+np.exp(-scores))
print props
```
**Checkpoint**: Make sure your probability predictions match the ones obtained from GraphLab Create.
```
print "Class predictions according to GraphLab Create:"
print sentiment_model.predict(sample_test_data, output_type='probability')
```
** Quiz Question:** Of the three data points in **sample_test_data**, which one (first, second, or third) has the **lowest probability** of being classified as a positive review?
# Find the most positive (and negative) review
We now turn to examining the full test dataset, **test_data**, and use GraphLab Create to form predictions on all of the test data points for faster performance.
Using the `sentiment_model`, find the 20 reviews in the entire **test_data** with the **highest probability** of being classified as a **positive review**. We refer to these as the "most positive reviews."
To calculate these top-20 reviews, use the following steps:
1. Make probability predictions on **test_data** using the `sentiment_model`. (**Hint:** When you call `.predict` to make predictions on the test data, use option `output_type='probability'` to output the probability rather than just the most likely class.)
2. Sort the data according to those predictions and pick the top 20. (**Hint:** You can use the `.topk` method on an SFrame to find the top k rows sorted according to the value of a specified column.)
```
test_data['probability'] = sentiment_model.predict(test_data, output_type='probability')
test_data['name','probability'].topk('probability', k=20).print_rows(num_rows = 20)
```
**Quiz Question**: Which of the following products are represented in the 20 most positive reviews? [multiple choice]
Now, let us repeat this exercise to find the "most negative reviews." Use the prediction probabilities to find the 20 reviews in the **test_data** with the **lowest probability** of being classified as a **positive review**. Repeat the same steps above but make sure you **sort in the opposite order**.
```
test_data['name','probability'].topk('probability', k=20, reverse= True).print_rows(num_rows = 20)
```
**Quiz Question**: Which of the following products are represented in the 20 most negative reviews? [multiple choice]
## Compute accuracy of the classifier
We will now evaluate the accuracy of the trained classifier. Recall that the accuracy is given by
$$
\mbox{accuracy} = \frac{\mbox{# correctly classified examples}}{\mbox{# total examples}}
$$
This can be computed as follows:
* **Step 1:** Use the trained model to compute class predictions (**Hint:** Use the `predict` method)
* **Step 2:** Count the number of data points when the predicted class labels match the ground truth labels (called `true_labels` below).
* **Step 3:** Divide the total number of correct predictions by the total number of data points in the dataset.
Complete the function below to compute the classification accuracy:
```
def get_classification_accuracy(model, data, true_labels):
# First get the predictions
## YOUR CODE HERE
predictions = model.predict(data)
# Compute the number of correctly classified examples
## YOUR CODE HERE
num_correct = sum(predictions == true_labels)
# Then compute accuracy by dividing num_correct by total number of examples
## YOUR CODE HERE
accuracy = num_correct / len(data)
return accuracy
```
Now, let's compute the classification accuracy of the **sentiment_model** on the **test_data**.
```
get_classification_accuracy(sentiment_model, test_data, test_data['sentiment'])
```
**Quiz Question**: What is the accuracy of the **sentiment_model** on the **test_data**? Round your answer to 2 decimal places (e.g. 0.76).
**Quiz Question**: Does a higher accuracy value on the **training_data** always imply that the classifier is better?
## Learn another classifier with fewer words
There were a lot of words in the model we trained above. We will now train a simpler logistic regression model using only a subset of words that occur in the reviews. For this assignment, we selected a 20 words to work with. These are:
```
significant_words = ['love', 'great', 'easy', 'old', 'little', 'perfect', 'loves',
'well', 'able', 'car', 'broke', 'less', 'even', 'waste', 'disappointed',
'work', 'product', 'money', 'would', 'return']
len(significant_words)
```
For each review, we will use the **word_count** column and trim out all words that are **not** in the **significant_words** list above. We will use the [SArray dictionary trim by keys functionality]( https://dato.com/products/create/docs/generated/graphlab.SArray.dict_trim_by_keys.html). Note that we are performing this on both the training and test set.
```
train_data['word_count_subset'] = train_data['word_count'].dict_trim_by_keys(significant_words, exclude=False)
test_data['word_count_subset'] = test_data['word_count'].dict_trim_by_keys(significant_words, exclude=False)
```
Let's see what the first example of the dataset looks like:
```
train_data[0]['review']
```
The **word_count** column had been working with before looks like the following:
```
print train_data[0]['word_count']
```
Since we are only working with a subset of these words, the column **word_count_subset** is a subset of the above dictionary. In this example, only 2 `significant words` are present in this review.
```
print train_data[0]['word_count_subset']
```
## Train a logistic regression model on a subset of data
We will now build a classifier with **word_count_subset** as the feature and **sentiment** as the target.
```
simple_model = graphlab.logistic_classifier.create(train_data,
target = 'sentiment',
features=['word_count_subset'],
validation_set=None)
simple_model
```
We can compute the classification accuracy using the `get_classification_accuracy` function you implemented earlier.
```
get_classification_accuracy(simple_model, test_data, test_data['sentiment'])
```
Now, we will inspect the weights (coefficients) of the **simple_model**:
```
simple_model.coefficients
```
Let's sort the coefficients (in descending order) by the **value** to obtain the coefficients with the most positive effect on the sentiment.
```
simple_model.coefficients.sort('value', ascending=False).print_rows(num_rows=21)
```
**Quiz Question**: Consider the coefficients of **simple_model**. There should be 21 of them, an intercept term + one for each word in **significant_words**. How many of the 20 coefficients (corresponding to the 20 **significant_words** and *excluding the intercept term*) are positive for the `simple_model`?
```
simple_weights = simple_model.coefficients
positive_significant_words = simple_weights[(simple_weights['value'] > 0) & (simple_weights['name'] == "word_count_subset")]['index']
print len(positive_significant_words)
print positive_significant_words
```
**Quiz Question**: Are the positive words in the **simple_model** (let us call them `positive_significant_words`) also positive words in the **sentiment_model**?
```
weights.filter_by(positive_significant_words, 'index')
```
# Comparing models
We will now compare the accuracy of the **sentiment_model** and the **simple_model** using the `get_classification_accuracy` method you implemented above.
First, compute the classification accuracy of the **sentiment_model** on the **train_data**:
```
get_classification_accuracy(sentiment_model, train_data, train_data['sentiment'])
```
Now, compute the classification accuracy of the **simple_model** on the **train_data**:
```
get_classification_accuracy(simple_model, train_data, train_data['sentiment'])
```
**Quiz Question**: Which model (**sentiment_model** or **simple_model**) has higher accuracy on the TRAINING set?
Now, we will repeat this exercise on the **test_data**. Start by computing the classification accuracy of the **sentiment_model** on the **test_data**:
```
get_classification_accuracy(sentiment_model, test_data, test_data['sentiment'])
```
Next, we will compute the classification accuracy of the **simple_model** on the **test_data**:
```
get_classification_accuracy(simple_model, test_data, test_data['sentiment'])
```
**Quiz Question**: Which model (**sentiment_model** or **simple_model**) has higher accuracy on the TEST set?
## Baseline: Majority class prediction
It is quite common to use the **majority class classifier** as the a baseline (or reference) model for comparison with your classifier model. The majority classifier model predicts the majority class for all data points. At the very least, you should healthily beat the majority class classifier, otherwise, the model is (usually) pointless.
What is the majority class in the **train_data**?
```
num_positive = (train_data['sentiment'] == +1).sum()
num_negative = (train_data['sentiment'] == -1).sum()
print num_positive
print num_negative
```
Now compute the accuracy of the majority class classifier on **test_data**.
**Quiz Question**: Enter the accuracy of the majority class classifier model on the **test_data**. Round your answer to two decimal places (e.g. 0.76).
```
print (test_data['sentiment'] == +1).sum()
print (test_data['sentiment'] == -1).sum()
print (test_data['sentiment'] == +1).sum()/len(test_data['sentiment'])
```
**Quiz Question**: Is the **sentiment_model** definitely better than the majority class classifier (the baseline)?
| github_jupyter |
```
# from google.colab import files
# uploaded = files.upload()
import numpy as np
import matplotlib.pyplot as plt
import h5py
from PIL import Image
from scipy import ndimage
import os
import io
from keras.applications.vgg16 import VGG16
from keras.applications.vgg16 import preprocess_input
from keras.preprocessing import image
from keras.preprocessing.image import ImageDataGenerator
from keras.models import Model, load_model
from tensorflow.keras.callbacks import EarlyStopping
from tensorflow.keras.layers import Dense, Dropout, Flatten, Conv2D, MaxPooling2D,Input
from tensorflow.keras.utils import to_categorical
from tensorflow.keras.models import Sequential
from tensorflow.keras import optimizers
train_dataset = h5py.File('train_catvnoncat.h5' , 'r')
train_features = np.array(train_dataset["train_set_x"][:])
train_labels = np.array(train_dataset["train_set_y"][:])
test_dataset = h5py.File('test_catvnoncat.h5', "r")
test_features = np.array(test_dataset["test_set_x"][:])
test_labels = np.array(test_dataset["test_set_y"][:])
classes = ['not-cat' , 'cat']
plt.imshow(train_features[2])
train_labels = to_categorical(train_labels,2)
test_labels = to_categorical(test_labels,2)
print(train_features.shape)
print(train_labels.shape)
```
DEFINING GENERATOR
```
datagen = ImageDataGenerator(preprocessing_function=preprocess_input,
rotation_range=40,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,
fill_mode='nearest')
```
TRAINING SET GENERATOR
```
train_generator = datagen.flow(train_features, train_labels)
```
VALIDATION SET GENERATOR
```
test_generator = datagen.flow(test_features, test_labels)
```
GETTING VGG
```
vgg_model=VGG16(include_top=False, weights='imagenet', input_shape=(64,64,3))
model = Sequential()
for layer in vgg_model.layers[:-1]:
model.add(layer)
for layer in model.layers:
layer.trainable = False
model.add(Flatten())
model.add(Dense(units=2,activation='softmax'))
#model.summary()
```
MODEL COMPILING
```
opt = optimizers.Adam(learning_rate=0.01)
model.compile(optimizer = opt , loss='categorical_crossentropy' , metrics = ['accuracy'] )
print(train_labels.shape)
print(test_labels.shape)
```
OUTPUT WITH DATA AUGMENTATION
```
model.fit(train_generator,epochs=12,verbose=1, steps_per_epoch=5 , validation_data=test_generator)
model.save('data_augmented.h5')
```
OUTPUT WHEN RAN WITHOUT DATA AUGMENTATION
```
#---------- THIS PROCESS REDUCES ACCURACY TO 60 % --------------
# train_features = train_features / 255
# test_features = test_features / 255
train_features=preprocess_input(train_features)
test_features=preprocess_input(test_features)
model.fit(train_features,train_labels,epochs=4,verbose=1,batch_size=10, validation_data=(test_features,test_labels) , shuffle=True)
model.save('regular.h5')
```
LOADING THE MODEL
```
from tensorflow.keras.preprocessing.image import load_img,img_to_array
from tensorflow.keras.applications.vgg16 import preprocess_input
from tensorflow.keras.models import Model, load_model
import cv2
import numpy as np
# IMAGE PREPROCESSING
def preprocess_image(name):
# img = cv2.imread(name)
img = load_img(name, target_size=(64,64))
res = img_to_array(img)
# res = cv2.resize(img, dsize=(64, 64), interpolation=cv2.INTER_CUBIC)
preprocessed_image = preprocess_input(img)
samples = np.expand_dims(preprocessed_image, 0)
return samples
image = preprocess_image('cat.png')
image2 = preprocess_image('not_cat.png')
reconstructed_model = load_model("data_augmented.h5")
prediction1 = reconstructed_model.predict(image)
prediction2 = reconstructed_model.predict(image2)
print(prediction1,prediction2)
```
| github_jupyter |
# SOP060 - Uninstall kubernetes module
## Steps
### Common functions
Define helper functions used in this notebook.
```
# Define `run` function for transient fault handling, hyperlinked suggestions, and scrolling updates on Windows
import sys
import os
import re
import platform
import shlex
import shutil
import datetime
from subprocess import Popen, PIPE
from IPython.display import Markdown
retry_hints = {} # Output in stderr known to be transient, therefore automatically retry
error_hints = {} # Output in stderr where a known SOP/TSG exists which will be HINTed for further help
install_hint = {} # The SOP to help install the executable if it cannot be found
def run(cmd, return_output=False, no_output=False, retry_count=0, base64_decode=False, return_as_json=False):
"""Run shell command, stream stdout, print stderr and optionally return output
NOTES:
1. Commands that need this kind of ' quoting on Windows e.g.:
kubectl get nodes -o jsonpath={.items[?(@.metadata.annotations.pv-candidate=='data-pool')].metadata.name}
Need to actually pass in as '"':
kubectl get nodes -o jsonpath={.items[?(@.metadata.annotations.pv-candidate=='"'data-pool'"')].metadata.name}
The ' quote approach, although correct when pasting into Windows cmd, will hang at the line:
`iter(p.stdout.readline, b'')`
The shlex.split call does the right thing for each platform, just use the '"' pattern for a '
"""
MAX_RETRIES = 5
output = ""
retry = False
# When running `azdata sql query` on Windows, replace any \n in """ strings, with " ", otherwise we see:
#
# ('HY090', '[HY090] [Microsoft][ODBC Driver Manager] Invalid string or buffer length (0) (SQLExecDirectW)')
#
if platform.system() == "Windows" and cmd.startswith("azdata sql query"):
cmd = cmd.replace("\n", " ")
# shlex.split is required on bash and for Windows paths with spaces
#
cmd_actual = shlex.split(cmd)
# Store this (i.e. kubectl, python etc.) to support binary context aware error_hints and retries
#
user_provided_exe_name = cmd_actual[0].lower()
# When running python, use the python in the ADS sandbox ({sys.executable})
#
if cmd.startswith("python "):
cmd_actual[0] = cmd_actual[0].replace("python", sys.executable)
# On Mac, when ADS is not launched from terminal, LC_ALL may not be set, which causes pip installs to fail
# with:
#
# UnicodeDecodeError: 'ascii' codec can't decode byte 0xc5 in position 4969: ordinal not in range(128)
#
# Setting it to a default value of "en_US.UTF-8" enables pip install to complete
#
if platform.system() == "Darwin" and "LC_ALL" not in os.environ:
os.environ["LC_ALL"] = "en_US.UTF-8"
# When running `kubectl`, if AZDATA_OPENSHIFT is set, use `oc`
#
if cmd.startswith("kubectl ") and "AZDATA_OPENSHIFT" in os.environ:
cmd_actual[0] = cmd_actual[0].replace("kubectl", "oc")
# To aid supportability, determine which binary file will actually be executed on the machine
#
which_binary = None
# Special case for CURL on Windows. The version of CURL in Windows System32 does not work to
# get JWT tokens, it returns "(56) Failure when receiving data from the peer". If another instance
# of CURL exists on the machine use that one. (Unfortunately the curl.exe in System32 is almost
# always the first curl.exe in the path, and it can't be uninstalled from System32, so here we
# look for the 2nd installation of CURL in the path)
if platform.system() == "Windows" and cmd.startswith("curl "):
path = os.getenv('PATH')
for p in path.split(os.path.pathsep):
p = os.path.join(p, "curl.exe")
if os.path.exists(p) and os.access(p, os.X_OK):
if p.lower().find("system32") == -1:
cmd_actual[0] = p
which_binary = p
break
# Find the path based location (shutil.which) of the executable that will be run (and display it to aid supportability), this
# seems to be required for .msi installs of azdata.cmd/az.cmd. (otherwise Popen returns FileNotFound)
#
# NOTE: Bash needs cmd to be the list of the space separated values hence shlex.split.
#
if which_binary == None:
which_binary = shutil.which(cmd_actual[0])
# Display an install HINT, so the user can click on a SOP to install the missing binary
#
if which_binary == None:
print(f"The path used to search for '{cmd_actual[0]}' was:")
print(sys.path)
if user_provided_exe_name in install_hint and install_hint[user_provided_exe_name] is not None:
display(Markdown(f'HINT: Use [{install_hint[user_provided_exe_name][0]}]({install_hint[user_provided_exe_name][1]}) to resolve this issue.'))
raise FileNotFoundError(f"Executable '{cmd_actual[0]}' not found in path (where/which)")
else:
cmd_actual[0] = which_binary
start_time = datetime.datetime.now().replace(microsecond=0)
print(f"START: {cmd} @ {start_time} ({datetime.datetime.utcnow().replace(microsecond=0)} UTC)")
print(f" using: {which_binary} ({platform.system()} {platform.release()} on {platform.machine()})")
print(f" cwd: {os.getcwd()}")
# Command-line tools such as CURL and AZDATA HDFS commands output
# scrolling progress bars, which causes Jupyter to hang forever, to
# workaround this, use no_output=True
#
# Work around a infinite hang when a notebook generates a non-zero return code, break out, and do not wait
#
wait = True
try:
if no_output:
p = Popen(cmd_actual)
else:
p = Popen(cmd_actual, stdout=PIPE, stderr=PIPE, bufsize=1)
with p.stdout:
for line in iter(p.stdout.readline, b''):
line = line.decode()
if return_output:
output = output + line
else:
if cmd.startswith("azdata notebook run"): # Hyperlink the .ipynb file
regex = re.compile(' "(.*)"\: "(.*)"')
match = regex.match(line)
if match:
if match.group(1).find("HTML") != -1:
display(Markdown(f' - "{match.group(1)}": "{match.group(2)}"'))
else:
display(Markdown(f' - "{match.group(1)}": "[{match.group(2)}]({match.group(2)})"'))
wait = False
break # otherwise infinite hang, have not worked out why yet.
else:
print(line, end='')
if wait:
p.wait()
except FileNotFoundError as e:
if install_hint is not None:
display(Markdown(f'HINT: Use {install_hint} to resolve this issue.'))
raise FileNotFoundError(f"Executable '{cmd_actual[0]}' not found in path (where/which)") from e
exit_code_workaround = 0 # WORKAROUND: azdata hangs on exception from notebook on p.wait()
if not no_output:
for line in iter(p.stderr.readline, b''):
try:
line_decoded = line.decode()
except UnicodeDecodeError:
# NOTE: Sometimes we get characters back that cannot be decoded(), e.g.
#
# \xa0
#
# For example see this in the response from `az group create`:
#
# ERROR: Get Token request returned http error: 400 and server
# response: {"error":"invalid_grant",# "error_description":"AADSTS700082:
# The refresh token has expired due to inactivity.\xa0The token was
# issued on 2018-10-25T23:35:11.9832872Z
#
# which generates the exception:
#
# UnicodeDecodeError: 'utf-8' codec can't decode byte 0xa0 in position 179: invalid start byte
#
print("WARNING: Unable to decode stderr line, printing raw bytes:")
print(line)
line_decoded = ""
pass
else:
# azdata emits a single empty line to stderr when doing an hdfs cp, don't
# print this empty "ERR:" as it confuses.
#
if line_decoded == "":
continue
print(f"STDERR: {line_decoded}", end='')
if line_decoded.startswith("An exception has occurred") or line_decoded.startswith("ERROR: An error occurred while executing the following cell"):
exit_code_workaround = 1
# inject HINTs to next TSG/SOP based on output in stderr
#
if user_provided_exe_name in error_hints:
for error_hint in error_hints[user_provided_exe_name]:
if line_decoded.find(error_hint[0]) != -1:
display(Markdown(f'HINT: Use [{error_hint[1]}]({error_hint[2]}) to resolve this issue.'))
# Verify if a transient error, if so automatically retry (recursive)
#
if user_provided_exe_name in retry_hints:
for retry_hint in retry_hints[user_provided_exe_name]:
if line_decoded.find(retry_hint) != -1:
if retry_count < MAX_RETRIES:
print(f"RETRY: {retry_count} (due to: {retry_hint})")
retry_count = retry_count + 1
output = run(cmd, return_output=return_output, retry_count=retry_count)
if return_output:
if base64_decode:
import base64
return base64.b64decode(output).decode('utf-8')
else:
return output
elapsed = datetime.datetime.now().replace(microsecond=0) - start_time
# WORKAROUND: We avoid infinite hang above in the `azdata notebook run` failure case, by inferring success (from stdout output), so
# don't wait here, if success known above
#
if wait:
if p.returncode != 0:
raise SystemExit(f'Shell command:\n\n\t{cmd} ({elapsed}s elapsed)\n\nreturned non-zero exit code: {str(p.returncode)}.\n')
else:
if exit_code_workaround !=0 :
raise SystemExit(f'Shell command:\n\n\t{cmd} ({elapsed}s elapsed)\n\nreturned non-zero exit code: {str(exit_code_workaround)}.\n')
print(f'\nSUCCESS: {elapsed}s elapsed.\n')
if return_output:
if base64_decode:
import base64
return base64.b64decode(output).decode('utf-8')
else:
return output
# Hints for tool retry (on transient fault), known errors and install guide
#
retry_hints = {'azdata': ['Endpoint sql-server-master does not exist', 'Endpoint livy does not exist', 'Failed to get state for cluster', 'Endpoint webhdfs does not exist', 'Adaptive Server is unavailable or does not exist', 'Error: Address already in use', 'Login timeout expired (0) (SQLDriverConnect)', 'SSPI Provider: No Kerberos credentials available', ], 'kubectl': ['A connection attempt failed because the connected party did not properly respond after a period of time, or established connection failed because connected host has failed to respond', ], 'python': [ ], }
error_hints = {'azdata': [['Please run \'azdata login\' to first authenticate', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['The token is expired', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['Reason: Unauthorized', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['Max retries exceeded with url: /api/v1/bdc/endpoints', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['Look at the controller logs for more details', 'TSG027 - Observe cluster deployment', '../diagnose/tsg027-observe-bdc-create.ipynb'], ['provided port is already allocated', 'TSG062 - Get tail of all previous container logs for pods in BDC namespace', '../log-files/tsg062-tail-bdc-previous-container-logs.ipynb'], ['Create cluster failed since the existing namespace', 'SOP061 - Delete a big data cluster', '../install/sop061-delete-bdc.ipynb'], ['Failed to complete kube config setup', 'TSG067 - Failed to complete kube config setup', '../repair/tsg067-failed-to-complete-kube-config-setup.ipynb'], ['Data source name not found and no default driver specified', 'SOP069 - Install ODBC for SQL Server', '../install/sop069-install-odbc-driver-for-sql-server.ipynb'], ['Can\'t open lib \'ODBC Driver 17 for SQL Server', 'SOP069 - Install ODBC for SQL Server', '../install/sop069-install-odbc-driver-for-sql-server.ipynb'], ['Control plane upgrade failed. Failed to upgrade controller.', 'TSG108 - View the controller upgrade config map', '../diagnose/tsg108-controller-failed-to-upgrade.ipynb'], ['NameError: name \'azdata_login_secret_name\' is not defined', 'SOP013 - Create secret for azdata login (inside cluster)', '../common/sop013-create-secret-for-azdata-login.ipynb'], ['ERROR: No credentials were supplied, or the credentials were unavailable or inaccessible.', 'TSG124 - \'No credentials were supplied\' error from azdata login', '../repair/tsg124-no-credentials-were-supplied.ipynb'], ['Please accept the license terms to use this product through', 'TSG126 - azdata fails with \'accept the license terms to use this product\'', '../repair/tsg126-accept-license-terms.ipynb'], ], 'kubectl': [['no such host', 'TSG010 - Get configuration contexts', '../monitor-k8s/tsg010-get-kubernetes-contexts.ipynb'], ['No connection could be made because the target machine actively refused it', 'TSG056 - Kubectl fails with No connection could be made because the target machine actively refused it', '../repair/tsg056-kubectl-no-connection-could-be-made.ipynb'], ], 'python': [['Library not loaded: /usr/local/opt/unixodbc', 'SOP012 - Install unixodbc for Mac', '../install/sop012-brew-install-odbc-for-sql-server.ipynb'], ['WARNING: You are using pip version', 'SOP040 - Upgrade pip in ADS Python sandbox', '../install/sop040-upgrade-pip.ipynb'], ], }
install_hint = {'azdata': [ 'SOP063 - Install azdata CLI (using package manager)', '../install/sop063-packman-install-azdata.ipynb' ], 'kubectl': [ 'SOP036 - Install kubectl command line interface', '../install/sop036-install-kubectl.ipynb' ], }
print('Common functions defined successfully.')
```
### Pip uninstall the kubernetes module
```
import sys
run(f'python -m pip uninstall kubernetes -y')
```
### Pip list installed modules
```
run(f'python -m pip list')
print("Notebook execution is complete.")
```
| github_jupyter |
# Importance of decision tree hyperparameters on generalization
In this notebook, we will illustrate the importance of some key
hyperparameters on the decision tree; we will demonstrate their effects on
the classification and regression problems we saw previously.
First, we will load the classification and regression datasets.
```
import pandas as pd
data_clf_columns = ["Culmen Length (mm)", "Culmen Depth (mm)"]
target_clf_column = "Species"
data_clf = pd.read_csv("../datasets/penguins_classification.csv")
data_reg_columns = ["Flipper Length (mm)"]
target_reg_column = "Body Mass (g)"
data_reg = pd.read_csv("../datasets/penguins_regression.csv")
```
<div class="admonition note alert alert-info">
<p class="first admonition-title" style="font-weight: bold;">Note</p>
<p class="last">If you want a deeper overview regarding this dataset, you can refer to the
Appendix - Datasets description section at the end of this MOOC.</p>
</div>
## Create helper functions
We will create some helper functions to plot the data samples as well as the
decision boundary for classification and the regression line for regression.
```
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from helpers.plotting import DecisionBoundaryDisplay
def fit_and_plot_classification(model, data, feature_names, target_names):
model.fit(data[feature_names], data[target_names])
if data[target_names].nunique() == 2:
palette = ["tab:red", "tab:blue"]
else:
palette = ["tab:red", "tab:blue", "black"]
DecisionBoundaryDisplay.from_estimator(
model, data[feature_names], response_method="predict",
cmap="RdBu", alpha=0.5
)
sns.scatterplot(data=data, x=feature_names[0], y=feature_names[1],
hue=target_names, palette=palette)
plt.legend(bbox_to_anchor=(1.05, 1), loc='upper left')
def fit_and_plot_regression(model, data, feature_names, target_names):
model.fit(data[feature_names], data[target_names])
data_test = pd.DataFrame(
np.arange(data.iloc[:, 0].min(), data.iloc[:, 0].max()),
columns=data[feature_names].columns,
)
target_predicted = model.predict(data_test)
sns.scatterplot(
x=data.iloc[:, 0], y=data[target_names], color="black", alpha=0.5)
plt.plot(data_test.iloc[:, 0], target_predicted, linewidth=4)
```
## Effect of the `max_depth` parameter
The hyperparameter `max_depth` controls the overall complexity of a decision
tree. This hyperparameter allows to get a trade-off between an under-fitted
and over-fitted decision tree. Let's build a shallow tree and then a deeper
tree, for both classification and regression, to understand the impact of the
parameter.
We can first set the `max_depth` parameter value to a very low value.
```
from sklearn.tree import DecisionTreeClassifier, DecisionTreeRegressor
max_depth = 2
tree_clf = DecisionTreeClassifier(max_depth=max_depth)
tree_reg = DecisionTreeRegressor(max_depth=max_depth)
fit_and_plot_classification(
tree_clf, data_clf, data_clf_columns, target_clf_column)
_ = plt.title(f"Shallow classification tree with max-depth of {max_depth}")
fit_and_plot_regression(
tree_reg, data_reg, data_reg_columns, target_reg_column)
_ = plt.title(f"Shallow regression tree with max-depth of {max_depth}")
```
Now, let's increase the `max_depth` parameter value to check the difference
by observing the decision function.
```
max_depth = 30
tree_clf = DecisionTreeClassifier(max_depth=max_depth)
tree_reg = DecisionTreeRegressor(max_depth=max_depth)
fit_and_plot_classification(
tree_clf, data_clf, data_clf_columns, target_clf_column)
_ = plt.title(f"Deep classification tree with max-depth of {max_depth}")
fit_and_plot_regression(
tree_reg, data_reg, data_reg_columns, target_reg_column)
_ = plt.title(f"Deep regression tree with max-depth of {max_depth}")
```
For both classification and regression setting, we observe that
increasing the depth will make the tree model more expressive. However, a
tree that is too deep will overfit the training data, creating partitions
which are only correct for "outliers" (noisy samples). The `max_depth` is one
of the hyperparameters that one should optimize via cross-validation and
grid-search.
```
from sklearn.model_selection import GridSearchCV
param_grid = {"max_depth": np.arange(2, 10, 1)}
tree_clf = GridSearchCV(DecisionTreeClassifier(), param_grid=param_grid)
tree_reg = GridSearchCV(DecisionTreeRegressor(), param_grid=param_grid)
fit_and_plot_classification(
tree_clf, data_clf, data_clf_columns, target_clf_column)
_ = plt.title(f"Optimal depth found via CV: "
f"{tree_clf.best_params_['max_depth']}")
fit_and_plot_regression(
tree_reg, data_reg, data_reg_columns, target_reg_column)
_ = plt.title(f"Optimal depth found via CV: "
f"{tree_reg.best_params_['max_depth']}")
```
With this example, we see that there is not a single value that is optimal
for any dataset. Thus, this parameter is required to be optimized for each
application.
## Other hyperparameters in decision trees
The `max_depth` hyperparameter controls the overall complexity of the tree.
This parameter is adequate under the assumption that a tree is built
symmetrically. However, there is no guarantee that a tree will be symmetrical.
Indeed, optimal generalization performance could be reached by growing some of
the branches deeper than some others.
We will built a dataset where we will illustrate this asymmetry. We will
generate a dataset composed of 2 subsets: one subset where a clear separation
should be found by the tree and another subset where samples from both
classes will be mixed. It implies that a decision tree will need more splits
to classify properly samples from the second subset than from the first
subset.
```
from sklearn.datasets import make_blobs
data_clf_columns = ["Feature #0", "Feature #1"]
target_clf_column = "Class"
# Blobs that will be interlaced
X_1, y_1 = make_blobs(
n_samples=300, centers=[[0, 0], [-1, -1]], random_state=0)
# Blobs that will be easily separated
X_2, y_2 = make_blobs(
n_samples=300, centers=[[3, 6], [7, 0]], random_state=0)
X = np.concatenate([X_1, X_2], axis=0)
y = np.concatenate([y_1, y_2])
data_clf = np.concatenate([X, y[:, np.newaxis]], axis=1)
data_clf = pd.DataFrame(
data_clf, columns=data_clf_columns + [target_clf_column])
data_clf[target_clf_column] = data_clf[target_clf_column].astype(np.int32)
sns.scatterplot(data=data_clf, x=data_clf_columns[0], y=data_clf_columns[1],
hue=target_clf_column, palette=["tab:red", "tab:blue"])
_ = plt.title("Synthetic dataset")
```
We will first train a shallow decision tree with `max_depth=2`. We would
expect this depth to be enough to separate the blobs that are easy to
separate.
```
max_depth = 2
tree_clf = DecisionTreeClassifier(max_depth=max_depth)
fit_and_plot_classification(
tree_clf, data_clf, data_clf_columns, target_clf_column)
_ = plt.title(f"Decision tree with max-depth of {max_depth}")
```
As expected, we see that the blue blob on the right and the red blob on the
top are easily separated. However, more splits will be required to better
split the blob were both blue and red data points are mixed.
Indeed, we see that red blob on the top and the blue blob on the right of
the plot are perfectly separated. However, the tree is still making mistakes
in the area where the blobs are mixed together. Let's check the tree
representation.
```
from sklearn.tree import plot_tree
_, ax = plt.subplots(figsize=(10, 10))
_ = plot_tree(tree_clf, ax=ax, feature_names=data_clf_columns)
```
We see that the right branch achieves perfect classification. Now, we
increase the depth to check how the tree will grow.
```
max_depth = 6
tree_clf = DecisionTreeClassifier(max_depth=max_depth)
fit_and_plot_classification(
tree_clf, data_clf, data_clf_columns, target_clf_column)
_ = plt.title(f"Decision tree with max-depth of {max_depth}")
_, ax = plt.subplots(figsize=(11, 7))
_ = plot_tree(tree_clf, ax=ax, feature_names=data_clf_columns)
```
As expected, the left branch of the tree continue to grow while no further
splits were done on the right branch. Fixing the `max_depth` parameter would
cut the tree horizontally at a specific level, whether or not it would
be more beneficial that a branch continue growing.
The hyperparameters `min_samples_leaf`, `min_samples_split`,
`max_leaf_nodes`, or `min_impurity_decrease` allows growing asymmetric trees
and apply a constraint at the leaves or nodes level. We will check the effect
of `min_samples_leaf`.
```
min_samples_leaf = 60
tree_clf = DecisionTreeClassifier(min_samples_leaf=min_samples_leaf)
fit_and_plot_classification(
tree_clf, data_clf, data_clf_columns, target_clf_column)
_ = plt.title(
f"Decision tree with leaf having at least {min_samples_leaf} samples")
_, ax = plt.subplots(figsize=(10, 7))
_ = plot_tree(tree_clf, ax=ax, feature_names=data_clf_columns)
```
This hyperparameter allows to have leaves with a minimum number of samples
and no further splits will be searched otherwise. Therefore, these
hyperparameters could be an alternative to fix the `max_depth`
hyperparameter.
| github_jupyter |
```
import sys
sys.path.insert(0,'..')
from src.evaluation.gnn_evaluation_module import train_and_get_embeddings
from src.models.multi_layered_model import MonoModel
from torch_geometric.nn import GCNConv
from src.data.data_loader import GraphDataset
def eval_original(dataset_name, directionality='undirected',
train_examples=20, val_examples=30):
isDirected = (directionality != 'undirected')
isReversed = (directionality == 'reversed')
dataset = GraphDataset(f'../data/tmp/{dataset_name}{("_" + directionality) if isDirected else ""}', dataset_name,
f'../data/graphs/processed/{dataset_name}/{dataset_name}.cites',
f'../data/graphs/processed/{dataset_name}/{dataset_name}.content',
directed=isDirected, reverse=isReversed)
emb = train_and_get_embeddings(dataset, channels=[96], modelType=GCNConv, architecture=MonoModel,
lr=0.01, wd=0.01, heads=1, dropout=0.8, attention_dropout=None,
epochs=200,
train_examples=train_examples, val_examples=val_examples,
split_seed=0, init_seed=0,
test_score=False, actual_predictions=False, add_complete_edges=False)
return emb, dataset[0].y
def eval_modcm(dataset_name, directionality='undirected',
train_examples=20, val_examples=30, modcm_inits=1):
isDirected = (directionality != 'undirected')
isReversed = (directionality == 'reversed')
for i in range(modcm_inits):
dataset = GraphDataset(f'../data/tmp/{dataset_name}{("_" + directionality) if isDirected else ""}-modcm{i}', dataset_name,
f'../data/graphs/modcm/{dataset_name}/{dataset_name}_modcm_{i}.cites',
f'../data/graphs/processed/{dataset_name}/{dataset_name}.content',
directed=isDirected, reverse=isReversed)
emb = train_and_get_embeddings(dataset, channels=[96], modelType=GCNConv, architecture=MonoModel,
lr=0.01, wd=0.01, heads=1, dropout=0.8, attention_dropout=None,
epochs=200,
train_examples=train_examples, val_examples=val_examples,
split_seed=0, init_seed=0,
test_score=False, actual_predictions=False, add_complete_edges=False)
return emb, dataset[0].y
x_original,y_original = eval_original('cora')
x_lcm,y_lcm = eval_modcm('cora')
x_original = x_original.detach().cpu().numpy()
y_original = y_original.detach().cpu().numpy()
x_lcm = x_lcm.detach().cpu().numpy()
y_lcm = y_lcm.detach().cpu().numpy()
import umap
```
| github_jupyter |
[](https://lab.mlpack.org/v2/gh/mlpack/examples/master?urlpath=lab%2Ftree%2Fcifar10_transformation_with_pca%2Fcifar-10-pca-py.ipynb)
```
# @file cifar-10-pca-py.ipynb
#
# A simple example usage of Principal Component Analysis (PCA)
# applied to the CIFAR-10 dataset.
#
# https://www.cs.toronto.edu/~kriz/cifar.html
import mlpack
import pandas as pd
import numpy as np
import glob
import os
from matplotlib import pyplot as plt
import matplotlib.image as mpimg
%matplotlib inline
# Load the CIFAR10 data from an online URL.
!wget -O - https://lab.mlpack.org/data/cifar10-images.tar.gz | tar -xz
# Create a list with all train and test images.
train_images_path = glob.glob('cifar10-images/train/**/*.png', recursive=True)
test_images_path = glob.glob('cifar10-images/test/**/*.png', recursive=True)
# Extract the labels from the image path.
train_labels = []
for img in train_images_path:
train_labels.append(img.split('/')[2])
test_labels = []
for img in test_images_path:
test_labels.append(img.split('/')[2])
# Print the image shape of the first image from the testing data.
img = mpimg.imread(test_images_path[0])
img.shape
# Display images from the training and testing data.
plt.figure(figsize=[20,20])
# Display image number 0 from the training data.
plt.subplot(141)
plt.imshow(mpimg.imread(train_images_path[0]))
plt.title("Label: " + train_labels[0]);
# Display image number 3000 from the training data.
plt.subplot(142)
plt.imshow(mpimg.imread(train_images_path[5000]))
plt.title("Label: " + train_labels[5000]);
# Display image number 2000 from the testing data.
plt.subplot(143)
plt.imshow(mpimg.imread(test_images_path[2000]))
plt.title("Label: " + test_labels[2000]);
# Display image number 7000 from the testing data.
plt.subplot(144)
plt.imshow(mpimg.imread(test_images_path[7000]))
plt.title("Label: " + test_labels[7000]);
# Create Dataframe from the testing images, this can take some time.
features = ['pixel' + str(i) for i in range(img.shape[0] * img.shape[1] * img.shape[2])]
samples = {}
for c, im in enumerate(test_images_path):
if c % 1000 == 0:
print('Loaded images: %d' % (c))
im = mpimg.imread(im)
im = im.reshape(-1, img.shape[0] * img.shape[1] * img.shape[2])
samples[str(c)] = im.tolist()[0]
cifar = pd.DataFrame.from_dict(samples, orient='index')
# Print the first 10 rows of the data.
cifar.head(10)
# Perform Principal Components Analysis using the randomized method.
# Other decomposition methods are 'exact', 'randomized-block-krylov', 'quic'.
#
# For more information checkout https://www.mlpack.org/doc/stable/python_documentation.html#pca
# or uncomment the line below.
# mlpack.pca?
data = mlpack.pca(decomposition_method='randomized', scale=True, input=cifar, new_dimensionality=2)
# Plot the transformed input.
plt.figure();
plt.figure(figsize=(10, 10))
plt.xticks(fontsize=10)
plt.yticks(fontsize=10)
plt.xlabel('Principal Component - 1', fontsize=20)
plt.ylabel('Principal Component - 2', fontsize=20)
plt.title('Projection of CIFAR-10 dataset onto first two principal components', fontsize=20)
targets = ['Airplane', 'Automobile', 'Bird', 'Cat', 'Deer', 'Dog', 'Frog', 'Horse', 'Ship', 'Truck']
colors = ['#DB579E', '#C957DB', '#7957DB', '#5784DB', '#57D3DB', '#57DB94', '#69DB57', '#B9DB57', '#DBAE57', '#DB5F57']
test_labels = np.array(test_labels)
for target, color in zip(targets, colors):
indices = np.where(test_labels == target)
plt.scatter(data['output'][indices,[0]], data['output'][indices,[1]], c=color, s=50)
plt.legend(targets, prop={'size': 15});
```
We can observe some structure in the plot above, samples belonging to the same class are close to each other;
images that are different like a Truck and a Horse are further away. We can also observe that the
first two principal components aren't sufficient for separating the classes.
| github_jupyter |
# Crystal1Johnson Tweets
@Crystal1Johnson's Twitter account was swept up in Twitter's suspension of Russian disinformation campaigns. This notebook explores what tweets of hers are in the Internet Archive's Wayback Machine using the [wayback](https://wayback.readthedocs.io/en/stable/) Python library that the kind folks at [EDGI](https://envirodatagov.org/) have created.
```
import wayback
```
First lets see what days her profile page has been archived on: https://twitter.com/Crystal1Johnson.
```
wb = wayback.WaybackClient()
for result in wb.search('https://twitter.com/Crystal1Johnson'):
print(result.timestamp, result.view_url)
```
You can see that there was some sporadic coverage and then a ton of archive requests on 2016-12-01. Something must have happened in Twitter then for so many requests.
Lets write the days that her profile page was archived, and the Wayback URL to view it for Kevin.
```
import csv
out = csv.writer(open('data/crystal1johnson-profile.csv', 'w'))
out.writerow(['archive time', 'wayback url'])
for result in wb.search('https://twitter.com/Crystal1Johnson'):
out.writerow([str(result.timestamp), result.view_url])
```
Now lets look to see what individual tweets have been archived. Since you can't edit tweets we are really only interested in unique URLs that match the pattern https://twitter.com/Crystal1Johnson/status/{id} Also it's worth pointing out that Twitter screen names are cast insenstive, so https://twitter.com/Crystal1Johnson/status/123 is the same as https://twitter.com/crystal1johnson/status/123
```
import re
from urllib.parse import urlparse
tweets = set()
archive_urls = {}
for result in wb.search('https://twitter.com/Crystal1Johnson', matchType='prefix'):
url = urlparse(result.url)
if re.match(r'/crystal1johnson/status/\d+$', url.path, re.IGNORECASE):
tweet_url = '{0.scheme}://{0.netloc}{0.path}'.format(url).lower()
tweets.add(tweet_url)
archive_urls[tweet_url] = result.view_url
for url in seen_tweets:
print('{0} archived at {1}'.format(url, archive_urls[url]))
```
It might be interesting to see the number of times the tweet was archived, since this can be a proxy for user attention on the tweet. So lets do it again using a counter.
```
from collections import Counter
tweet_counter = Counter()
archived_urls = {}
for result in wb.search('https://twitter.com/Crystal1Johnson', matchType='prefix'):
url = urlparse(result.url)
if re.match(r'/crystal1johnson/status/\d+$', url.path, re.IGNORECASE):
tweet_url = '{0.scheme}://{0.netloc}{0.path}'.format(url).lower()
tweet_counter.update([tweet_url])
archived_urls[tweet_url] = result.view_url
for url, count in tweet_counter.most_common():
print(url, count)
```
Now I'm going to write these out to a CSV to give to Kevin :-)
```
import csv
out = csv.writer(open('data/crystal1johnson.csv', 'w'))
out.writerow(['tweet url', 'count', 'archive url'])
for url, count in tweet_counter.most_common():
out.writerow([url, count, archived_urls[url]])
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
df = pd.read_csv('../csv/SBER_101001_201001_1d.csv',
delimiter=',',
parse_dates=[[0,1]],
index_col=0,
names=['Date','Time','Open', 'High', 'Low', 'Close', 'Volume'],
header=0,
encoding='utf-8'
)
df['Close'].plot(figsize=(10,7))
```
# Подготовка данных
```
df['Buy'] = (df['Close']-df['Open'])/df['Open']
df['Sell'] = (df['Open']-df['Close'])/df['Open']
```
# Покупки после каждого закрытия вниз
```
df['Buy_1_Day'] = np.where(df['Buy'].shift(1)<0, df['Buy'], np.nan)
df['Equity_Buy_1_Day'] = df['Buy_1_Day'].cumsum().apply(np.exp)
```
# Продажа после каждого закрытия вверх
```
df['Sell_1_Day'] = np.where(df['Sell'].shift(1)<0, df['Sell'], np.nan)
df['Equity_Sell_1_Day'] = df['Sell_1_Day'].cumsum().apply(np.exp)
```
# Покупка и продажа вместе
```
df['Buysell'] = np.where(df['Buy'].shift(1)<0, df['Buy'], 0)
df['Buysell'] = np.where(df['Buy'].shift(1)>0, df['Sell'], 0)
df['Equity_Buysell_1_Day'] = df['Buysell'].cumsum().apply(np.exp)
```
# Покупка после двух подряд закрытий вниз
```
def calc_n_days(values, n):
if (values[:n] < 0).all():
return values[-1]
else:
return np.nan
df['Buy_2_Days'] = df['Buy'].rolling(3).apply(calc_n_days, args=(2,))
df['Equity_Buy_2_Days'] = df['Buy_2_Days'].cumsum().apply(np.exp)
```
# Продажа после двух подряд закрытий вверх
```
df['Sell_2_Days'] = df['Sell'].rolling(3).apply(calc_n_days, args=(2,))
df['Equity_Sell_2_Days'] = df['Sell_2_Days'].cumsum().apply(np.exp)
```
# Продажа и покупка вместе после двух дней подряд
```
def calc_n_days_buysell(values, n):
if (values[:n] < 0).all():
return values[-1]
elif (values[:n] > 0).all():
return -1*values[-1]
else:
return np.nan
df['Buysell_2_Days'] = df['Buysell'].rolling(3).apply(calc_n_days_buysell, args=(2,))
df['Equity_Buysell_2_Days'] = df['Buysell_2_Days'].cumsum().apply(np.exp)
```
# Покупка после трёх подряд закрытий вниз
```
df['Buy_3_Days'] = df['Buy'].rolling(4).apply(calc_n_days, args=(3,))
df['Equity_Buy_3_Days'] = df['Buy_3_Days'].cumsum().apply(np.exp)
```
# Продажа после трёх подряд закрытий вверх
```
df['Sell_3_Days'] = df['Sell'].rolling(4).apply(calc_n_days, args=(3,))
df['Equity_Sell_3_Days'] = df['Sell_3_Days'].cumsum().apply(np.exp)
```
# Покупка и продажа вместе после трёх дней
```
df['Buysell_3_Days'] = df['Buysell'].rolling(4).apply(calc_n_days_buysell, args=(3,))
df['Equity_Buysell_3_Days'] = df['Buysell_3_Days'].cumsum().apply(np.exp)
```
# Графики
```
df[[
'Equity_Buy_1_Day',
'Equity_Buy_2_Days',
'Equity_Buy_3_Days'
]].interpolate(method='index').plot(figsize=(10,7))
df[[
'Equity_Sell_1_Day',
'Equity_Sell_2_Days',
'Equity_Sell_3_Days'
]].interpolate(method='index').plot(figsize=(10,7))
df[[
'Equity_Buysell_1_Day',
'Equity_Buysell_2_Days',
'Equity_Buysell_3_Days'
]].interpolate(method='index').plot(figsize=(10,7))
```
| github_jupyter |
# Data Science Ex 12 - Clustering (Density-based Methods)
12.05.2021, Lukas Kretschmar (lukas.kretschmar@ost.ch)
## Let's have some Fun with Density-based Clustering approaches!
In this exercise, we are going to have a look at density-based clustering.
Further, we have a look at possibilities to scale and normalize data.
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
sns.set()
```
## Introduction
Before we go to the density-based clustering approach, we want to introduce some preprocessing steps for a data scientist.
- Scaling numerical data
- Normalizing numerical data
- Encoding categorical data
These knowledge is important since we need to prepare the data for clustering.
For one, it could be a good idea to reduce the dimensions (number of features) of our data and therefore improve the runtime of our algorithms.
Or we can visualize the data easier.
Further, since clustering needs to calculate the distance between points, the values should be in the same range.
Otherwise, some features will dominate over others.
## Preprocessing
### Preprocessing Data
When building clusters, it's essential that the values are in a comparable range.
Otherwise, calculating distances will include biases (higher values have a larger impact - e.g. one column is in `km`, another in `mm`).
Therefore, we need to know some techniques to scale our data.
#### Numerical Data
References: https://scikit-learn.org/stable/modules/preprocessing.html & https://scikit-learn.org/stable/auto_examples/preprocessing/plot_all_scaling.html
We will have a look at the follwing possibilities (there are many more):
- Normalizer: Normalizes a row to unit norm (the sum of all values is `1`, the values are relative to each other).
- MinMaxScaler: Transforms features into a defined scale.
- RobustScaler: Scales features but mitigates outliers.
- StandardScaler: Scales features to unit variance.
```
from sklearn.preprocessing import Normalizer, MinMaxScaler, RobustScaler, StandardScaler
rng = np.random.RandomState(42)
data = pd.DataFrame(rng.randn(100000) + 5, columns=["Values"])
data
data.hist(bins=100)
```
The sample data has a normal distribution around a mean of 5.
Now, let's have a look what effects the scalers and normalizers make to this data.
##### Normalizer
Reference: https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.Normalizer.html
The `Normalizer()` scales the values in a way that the sum is `1`.
We already saw this method in action in the last exercise - when we called `normalize()`.
`Normalize()` is just the class implementing the method and can be used in `Pipelines` or when a class instead of a method is needed.
```
norm = Normalizer()
data_norm = norm.fit_transform(data.T) # Since the algorithm works on a row, we have to transform the data
data_norm = pd.DataFrame(data_norm.T)
fig, ax = plt.subplots(1,2,figsize=(20,5))
data_norm.hist(bins=100, ax=ax[0])
(data_norm[0]
.sort_values() # Sorting all values in ascending order
.reset_index(drop=True) # Removing index
.apply(lambda v: v**2) # Squaring values
.cumsum() # Taking the cumulative sum
.plot(ax=ax[1])) # Plotting the line
ax[0].set(title="Histogram of Normalizer (l2, default)")
ax[1].set(title="Cumulative sum of normalized data")
```
We had to square each value since the default behavior of `Normalize()` uses squares when normalizing.
If this behavior is not needed, but just the values relative to each other, we can set value of the `norm` parameter.
```
norm = Normalizer(norm="l1")
data_norm = norm.fit_transform(data.T)
data_norm = pd.DataFrame(data_norm.T)
fig, ax = plt.subplots(1,2,figsize=(20,5))
data_norm.hist(bins=100, ax=ax[0])
(data_norm[0]
.sort_values() # Sorting all values in ascending order
.reset_index(drop=True) # Removing index
.cumsum() # Taking the cumulative sum
.plot(ax=ax[1])) # Plotting the line
ax[0].set(title="Histogram of Normalizer (l1)")
ax[1].set(title="Cumulative sum of normalized data")
```
As you can see, the values changed but the sum is still `1`.
##### MinMaxScaler
Reference: https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.MinMaxScaler.html
The `MinMaxScaler` transforms the data so it is in a given range.
The default range is `(0,1)`.
```
minmax = MinMaxScaler()
data_minmax = minmax.fit_transform(data)
data_minmax = pd.DataFrame(data_minmax)
print(f"Min: {data_minmax.min()[0]}")
print(f"Max: {data_minmax.max()[0]}")
fig, ax = plt.subplots(figsize=(10,5))
data_minmax.hist(bins=100, ax=ax)
ax.set(title="Histogram of MinMaxScaler")
```
As you can see, all the values are now scaled to a range from `0` to `1`.
We can change this by providing an argument for parameter `feature_range`.
```
minmax = MinMaxScaler(feature_range=(2,4))
data_minmax = minmax.fit_transform(data)
data_minmax = pd.DataFrame(data_minmax)
print(f"Min: {data_minmax.min()[0]}")
print(f"Max: {data_minmax.max()[0]}")
fig, ax = plt.subplots(figsize=(10,5))
data_minmax.hist(bins=100, ax=ax)
ax.set(title="Histogram of MinMaxScaler (2,4)")
```
##### RobustScaler
Reference: https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.RobustScaler.html
The `RobustScaler` transforms the data based on a given quantile range (default is 1st quartile (25%) to 3rd quartile (75%)).
With this approach we try to remove the impact of outliers.
```
robust = RobustScaler()
data_robust = robust.fit_transform(data)
data_robust = pd.DataFrame(data_robust)
fig, ax = plt.subplots(1,2,figsize=(20, 5))
(data - 5).hist(bins=100, ax=ax[0])
data_robust.hist(bins=100, ax=ax[1])
ax[0].set(title="Original Data (shifted to 0)")
ax[1].set(title="Data with RobustScaler")
```
As you can see, the scaled data has a smaller range of values.
##### StandardScaler
Reference: https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.StandardScaler.html
The `StandardScaler` removes the mean and transforms the data using the variance.
```
std = StandardScaler()
data_std = std.fit_transform(data)
data_std = pd.DataFrame(data_std)
fig, ax = plt.subplots(1,2,figsize=(20,5))
data.hist(bins=100, ax=ax[0])
data_std.hist(bins=100, ax=ax[1])
ax[0].set(title="Original Data")
ax[1].set(title="Data with StandardScaler")
rng = np.random.RandomState(42)
data_exp = pd.DataFrame(rng.uniform(5,10,size=100000), columns=["Values"])
data_exp_std = StandardScaler().fit_transform(data_exp)
data_exp_std = pd.DataFrame(data_exp_std)
fig, ax = plt.subplots(1,2,figsize=(20,5))
data_exp.hist(bins=100, ax=ax[0])
data_exp_std.hist(bins=100, ax=ax[1])
ax[0].set(title="Original Data (Uniform Distribution)")
ax[1].set(title="Data with StandardScaler")
```
As we can see, the scaled data has now a mean of `0` but the distribution hasn't changed.
#### Categorial Data
When working with categorial data, we usually need to transform it into numbers.
We have already seen one approach with the `pd.get_dummies()` method.
Here, we will introduce another method that accomplishes the same.
```
jobs = ["Engineer", "Accountant", "Manager", "Professor", "Student"]
people = pd.DataFrame({"Name" : ["Johnny", "Jenny", "Jake"], "Job":["Engineer", "Manager", "Student"]})
people
```
##### OneHotEncoder
Reference: https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.OneHotEncoder.html
```
from sklearn.preprocessing import OneHotEncoder
enc = OneHotEncoder(sparse=False) # sparse=False will get us an array as result and not a sparse array object
data_cat = enc.fit_transform(people["Job"].to_numpy().reshape(-1,1))
data_cat = pd.DataFrame(data_cat, columns=["Is_" + str(c) for c in enc.categories_[0]])
data_cat
```
As we can see, the result is a dataset with multiple columns containing a `1` if a category was present in a row.
Or `0` to indicate the absence of this category.
We can also provide a complete list of all possible values.
And then for every possibility a column is provided.
```
enc = OneHotEncoder(categories=[jobs], sparse=False)
data_cat = enc.fit_transform(people["Job"].to_numpy().reshape(-1,1))
data_cat = pd.DataFrame(data_cat, columns=["Is_" + str(c) for c in enc.categories_[0]])
data_cat
```
*Please note:* Compared with the examples above, we had to transform the values first.
Throwing a whole dataset (as shown in the examples above) at a scaler works well.
But when we just want to use the scaler for one column, we have to reshape the values first.
The column `Job` looks like this:
```
people["Job"]
```
When we just take the numpy array, we have an array of all values.
But the values would be treated as one row.
```
people["Job"].to_numpy()
```
Calling `reshape(-1,1)` switches the row to a column representation.
```
people["Job"].to_numpy().reshape(-1,1)
```
And with this kind of input, the scalers can work.
As an alternative, we could have also created a `DataFrame` with the one `Series`.
##### Distances with Categories
We've learned when working with categorial values, we have to use specific ways to calculate the distances.
Using methods from the [`DistanceMetric` class](https://scikit-learn.org/stable/modules/generated/sklearn.neighbors.DistanceMetric.html) or own implementations, we can use some clustering algorithms also with categorial data.
But we won't got into details in this exercise.
It's primarily an FIY.
### DENCLUE
Unfortunatelly, there is no implementation of DENCLUE provided by any package in Anaconda or other python packages.
Therefore, we had to find another way to get a DENCLUE algorithm up and running.
Luckily for us, there is an [open-source implementation](https://github.com/mgarrett57/DENCLUE) available which we are going to use.
**Disclaimer:** While testing the implementation, I ran into several issues with the code.
The algorithm was implemented in April 2017, and since then, a module and methods the algorithm uses were changed.
Thus, I had to fix the implementation so it runs with the current version of the `networkx` module.
It works now, but I cannot guarantee that no other problems will occur.
If you want to see how it's implemented, feel free to check out the [code in denclue.py](./denclue.py) yourself.
```
from denclue import DENCLUE
```
And now we can simply use the clustering algorithm by calling it.
```
DENCLUE()
```
The algorithm offers some hyperparameters as well (all are optional):
- **h**: Hill-climbing parameter (you can steer the size of the neighborhood)
- **eps**: Convergence threshold for density (you can stop the hill-climbing at a certain level)
- **min_density**: Threshold to consider a cluster and not noise
- **metric**: Distance metric used
Compared to the algorithms from sklearn, the interface for this algortihm is limited.
There is only a `fit()` method that we can use and the cluster assignments are stored in the `label_` property and we get information of all clusters by calling `clust_info_`.
Let's start with some data.
```
data = pd.read_csv("./Demo_3Cluster_Noise.csv")
data.head(5)
fig, ax = plt.subplots(figsize=(10,10))
data.plot.scatter("x", "y", c="b", ax=ax)
```
You can see, that we have 3 clusters in here, but there are points that aren't that close to an obvious cluster.
So let's see what the DENCLUE algorithm can do with such data.
*Note:* The execution might take a while.
```
model = DENCLUE()
model.fit(data.to_numpy()) # Unfortunately, the algorithm cannot handle DataFrams - so we need to provide an array.
```
By calling `labels_` we get the cluster numbers assigned to each point.
```
model.labels_
```
And with `clust_info_` we get some more insights how the clusters are set together.
```
model.clust_info_
```
So we see, that the algorithm found a total of 7 clusters in the given data.
But we see also, that 4 of the 7 cluster only contain 1 or 2 points.
Before we head into plotting, let's introduce a helper method.
This method creates a new dataset containing the centroids of each cluster.
And we can filter the clusters by specifying a `min_density`.
```
def get_centroids(model, min_density=0.0):
centroids = pd.DataFrame(columns=["x", "y", "density"])
for i in range(len(model.clust_info_)):
clust = model.clust_info_[i]
if(clust["density"] < min_density):
continue
centroid = pd.DataFrame([clust["centroid"]], columns=["x", "y"])
centroid["density"] = clust["density"]
centroids = centroids.append(centroid, ignore_index=True)
return centroids
centroids = get_centroids(model)
fig, ax = plt.subplots(figsize=(10,10))
data.plot.scatter("x", "y", ax=ax, c=model.labels_, cmap="rainbow", colorbar=False)
centroids.plot.scatter("x", "y", ax=ax, c="k", s=200, alpha=.5)
```
By calling the `set_minimum_density` we can change the number of clusters found in the data.
Those clusters not fulfilling the density requirement, count as outliers.
```
model.set_minimum_density(0.01)
centroids = get_centroids(model, min_density=0.01)
fig, ax = plt.subplots(figsize=(10,10))
data.plot.scatter("x", "y", ax=ax, c=model.labels_, cmap="rainbow", colorbar=False)
centroids.plot.scatter("x", "y", ax=ax, c="k", s=200, alpha=.5)
```
If we are honest, although some outliers were detected, the clusters still have some points that are quite far away from the center and could also be counted as outliers.
To reduce the cluster size, we can limit the boundaries of a cluster.
```
model_lim = DENCLUE(h=.5, min_density=0.01)
model_lim.fit(data.to_numpy())
len(model_lim.clust_info_)
centroids_lim = get_centroids(model_lim, min_density=0.01)
fig, ax = plt.subplots(figsize=(10,10))
data.plot.scatter("x", "y", ax=ax, c=model_lim.labels_, cmap="rainbow", colorbar=False)
centroids.plot.scatter("x", "y", ax=ax, c="k", s=200, alpha=.5)
```
The clusters detected are now smaller.
And the one on the bottom left even got split into two clusters.
## Exercises
### Ex01 - Preprocessing
In this exercise, you are going to use the scalers and normalizer introduced above.
First, load the data from **Ex12_01_Data.csv**.
What you have here are some specs on cars.
And you will now scale this data.
But before you start, create a new empty dataset with just the name and year since you won't scale these columns.
First, use a `MinMaxScaler` with a range of `(-1,1)` to scale the `mpg`.
Assign the results to the new dataset you've created above.
Next, the `cylinders`.
Use a `MinMaxScaler` again, but this time with a range of `(-2,2)`.
Now, use a `StandardScaler` for the `horsepower`.
Do the same for the `acceleration`.
In the last step, use a `RobustScaler` for `displacement` and `weight`.
And you are finished.
You've successfully scaled some features into more comparable ranges.
#### Solution
```
# %load ./Ex12_01_Sol.py
```
### Ex02 - Simple DENCLUE
In this exercise, you are going to use the DENCLUE algorithm in it's simplest form.
To begin, load **Ex12_02_Data.csv**.
Plot the clusters.
Use the value in the `label` column for coloring.
Run the `DENCLUE` algorithm for the dataset (use only columns `x` & `y`).
How many cluster were found?
Get the centroids of these clusters.
*Hint:* You may use the method defined in the introduction.
But feel free to code it yourself.
Plot the data.
Use the clusters assigned by the DENCLUE algorithm for coloring.
Plot the centroids as well.
We didn't limit the cluster density.
So, we will do it now to get only our 4 expected clusters.
Use the `set_minimum_density` method and use a reasonable value for the density so only the 4 clusters remain.
Plot the data again, with the centroids.
Congratulations!
You have used DENCLUE successfully.
#### Solutions
```
# %load ./Ex12_02_Sol.py
```
### Ex03 - More DENCLUE
Let's use DENCLUE again.
Load **Ex12_03_Data.csv**.
Plot the data so you have an idea with what you are dealing.
Use the DENCLUE algorithm again.
But this time specify from the beginning a `min_density` of `0.01`.
And set `h=.75`.
Get the centroids for the clusters.
How many cluster were found? - Only count those with a density equal or greater than specified above.
Plot the data with the found clusters.
Also plot the centroids.
So you see, clusters were found, but not the number as expected.
You know from the original data that we expect 6 clusters.
Try to find good values for the parameters to get close to these 6 clusters.
This exercise has no right or wrong answer.
It shows how hard it can be to find good parameters.
A good starting point is the following model:
```python
DENCLUE(h=.11, eps=.005, min_density=0.1)
```
But there is space to improve.
#### Solution
```
# %load ./Ex12_03_Sol.py
```
### Ex04 - Airbnb Clustering
In this exercise, you are going to find clusters of Airbnb listings in Zurich.
In the dataset **Ex12_04_Data.csv** you'll find the raw data from Zurich.
Plot these listings in a scatter plot using `longitude` and `latitude` and use the `availability_365` for coloring.
Select all listings that are *Entire home/apt* (`room_type`) and are at least availbale *300* days a year (`availability_365`).
How many listings would that be?
**Note:** During the rest of the exercise, we will just use this reduced dataset.
Working with the full dataset would result in a quite long execution of the *DENCLUE* algorithm.
And you want to finish this exercise, eventually.
Create and run the *DENCLUE* algorithm with `h=0.003`.
This code will take some time to run.
Set the `minimum_density` to *300*.
Get the centroids of the model.
Plot the data again, but with
- Outliers should be grayed out (visible as outliers)
- Clusters
- Centroids
#### Solution
```
# %load ./Ex12_04_Sol.py
```
| github_jupyter |
# Tools - NumPy
*NumPy is the fundamental library for scientific computing with Python. NumPy is centered around a powerful N-dimensional array object, and it also contains useful linear algebra, Fourier transform, and random number functions.*
## Creating arrays
Now let's import `numpy`. Most people import it as `np`:
```
import numpy as np
```
## `np.zeros`
The `zeros` function creates an array containing any number of zeros:
```
np.zeros(5)
```
It's just as easy to create a 2D array (ie. a matrix) by providing a tuple with the desired number of rows and columns. For example, here's a 3x4 matrix:
```
np.zeros((3,4))
```
## Some vocabulary
* In NumPy, each dimension is called an **axis**.
* The number of axes is called the **rank**.
* For example, the above 3x4 matrix is an array of rank 2 (it is 2-dimensional).
* The first axis has length 3, the second has length 4.
* An array's list of axis lengths is called the **shape** of the array.
* For example, the above matrix's shape is `(3, 4)`.
* The rank is equal to the shape's length.
* The **size** of an array is the total number of elements, which is the product of all axis lengths (eg. 3*4=12)
```
a = np.zeros((3,4))
a
a.shape
a.ndim # equal to len(a.shape)
a.size
```
## N-dimensional arrays
You can also create an N-dimensional array of arbitrary rank. For example, here's a 3D array (rank=3), with shape `(2,3,4)`:
```
np.zeros((2,3,4))
```
## Array type
NumPy arrays have the type `ndarray`s:
```
type(np.zeros((3,4)))
```
## `np.ones`
Many other NumPy functions create `ndarrays`.
Here's a 3x4 matrix full of ones:
```
np.ones((3,4))
```
## `np.full`
Creates an array of the given shape initialized with the given value. Here's a 3x4 matrix full of `π`.
```
np.full((3,4), np.pi)
```
## `np.empty`
An uninitialized 2x3 array (its content is not predictable, as it is whatever is in memory at that point):
```
np.empty((2,3))
```
## np.array
Of course you can initialize an `ndarray` using a regular python array. Just call the `array` function:
```
np.array([[1,2,3,4], [10, 20, 30, 40]])
```
## `np.arange`
You can create an `ndarray` using NumPy's `arange` function, which is similar to python's built-in `range` function:
```
np.arange(1, 5)
```
It also works with floats:
```
np.arange(1.0, 5.0)
```
Of course you can provide a step parameter:
```
np.arange(1, 5, 0.5)
```
However, when dealing with floats, the exact number of elements in the array is not always predictible. For example, consider this:
```
print(np.arange(0, 5/3, 1/3)) # depending on floating point errors, the max value is 4/3 or 5/3.
print(np.arange(0, 5/3, 0.333333333))
print(np.arange(0, 5/3, 0.333333334))
```
## `np.linspace`
For this reason, it is generally preferable to use the `linspace` function instead of `arange` when working with floats. The `linspace` function returns an array containing a specific number of points evenly distributed between two values (note that the maximum value is *included*, contrary to `arange`):
```
print(np.linspace(0, 5/3, 6))
```
## `np.rand` and `np.randn`
A number of functions are available in NumPy's `random` module to create `ndarray`s initialized with random values.
For example, here is a 3x4 matrix initialized with random floats between 0 and 1 (uniform distribution):
```
np.random.rand(3,4)
```
Here's a 3x4 matrix containing random floats sampled from a univariate [normal distribution](https://en.wikipedia.org/wiki/Normal_distribution) (Gaussian distribution) of mean 0 and variance 1:
```
np.random.randn(3,4)
```
To give you a feel of what these distributions look like, let's use matplotlib (see the [matplotlib tutorial](tools_matplotlib.ipynb) for more details):
```
%matplotlib inline
import matplotlib.pyplot as plt
plt.hist(np.random.rand(100000), density=True, bins=100, histtype="step", color="blue", label="rand")
plt.hist(np.random.randn(100000), density=True, bins=100, histtype="step", color="red", label="randn")
plt.axis([-2.5, 2.5, 0, 1.1])
plt.legend(loc = "upper left")
plt.title("Random distributions")
plt.xlabel("Value")
plt.ylabel("Density")
plt.show()
```
## np.fromfunction
You can also initialize an `ndarray` using a function:
```
def my_function(z, y, x):
return x * y + z
np.fromfunction(my_function, (3, 2, 10))
```
NumPy first creates three `ndarrays` (one per dimension), each of shape `(2, 10)`. Each array has values equal to the coordinate along a specific axis. For example, all elements in the `z` array are equal to their z-coordinate:
[[[ 0. 0. 0. 0. 0. 0. 0. 0. 0. 0.]
[ 0. 0. 0. 0. 0. 0. 0. 0. 0. 0.]]
[[ 1. 1. 1. 1. 1. 1. 1. 1. 1. 1.]
[ 1. 1. 1. 1. 1. 1. 1. 1. 1. 1.]]
[[ 2. 2. 2. 2. 2. 2. 2. 2. 2. 2.]
[ 2. 2. 2. 2. 2. 2. 2. 2. 2. 2.]]]
So the terms x, y and z in the expression `x * y + z` above are in fact `ndarray`s (we will discuss arithmetic operations on arrays below). The point is that the function `my_function` is only called *once*, instead of once per element. This makes initialization very efficient.
## Array data
## `dtype`
NumPy's `ndarray`s are also efficient in part because all their elements must have the same type (usually numbers).
You can check what the data type is by looking at the `dtype` attribute:
```
c = np.arange(1, 5)
print(c.dtype, c)
c = np.arange(1.0, 5.0)
print(c.dtype, c)
```
Instead of letting NumPy guess what data type to use, you can set it explicitly when creating an array by setting the `dtype` parameter:
```
d = np.arange(1, 5, dtype=np.complex64)
print(d.dtype, d)
```
Available data types include `int8`, `int16`, `int32`, `int64`, `uint8`|`16`|`32`|`64`, `float16`|`32`|`64` and `complex64`|`128`. Check out [the documentation](http://docs.scipy.org/doc/numpy-1.10.1/user/basics.types.html) for the full list.
## `itemsize`
The `itemsize` attribute returns the size (in bytes) of each item:
```
e = np.arange(1, 5, dtype=np.complex64)
e.itemsize
```
## `data` buffer
An array's data is actually stored in memory as a flat (one dimensional) byte buffer. It is available *via* the `data` attribute (you will rarely need it, though).
```
f = np.array([[1,2],[1000, 2000]], dtype=np.int32)
f.data
```
In python 2, `f.data` is a buffer. In python 3, it is a memoryview.
```
if (hasattr(f.data, "tobytes")):
data_bytes = f.data.tobytes() # python 3
else:
data_bytes = memoryview(f.data).tobytes() # python 2
data_bytes
```
Several `ndarrays` can share the same data buffer, meaning that modifying one will also modify the others. We will see an example in a minute.
## Reshaping an array
## In place
Changing the shape of an `ndarray` is as simple as setting its `shape` attribute. However, the array's size must remain the same.
```
g = np.arange(24)
print(g)
print("Rank:", g.ndim)
g.shape = (6, 4)
print(g)
print("Rank:", g.ndim)
g.shape = (2, 3, 4)
print(g)
print("Rank:", g.ndim)
```
## `reshape`
The `reshape` function returns a new `ndarray` object pointing at the *same* data. This means that modifying one array will also modify the other.
```
g2 = g.reshape(4,6)
print(g2)
print("Rank:", g2.ndim)
```
Set item at row 1, col 2 to 999 (more about indexing below).
```
g2[1, 2] = 999
g2
```
The corresponding element in `g` has been modified.
```
g
```
## `ravel`
Finally, the `ravel` function returns a new one-dimensional `ndarray` that also points to the same data:
```
g.ravel()
```
## Arithmetic operations
All the usual arithmetic operators (`+`, `-`, `*`, `/`, `//`, `**`, etc.) can be used with `ndarray`s. They apply *elementwise*:
```
a = np.array([14, 23, 32, 41])
b = np.array([5, 4, 3, 2])
print("a + b =", a + b)
print("a - b =", a - b)
print("a * b =", a * b)
print("a / b =", a / b)
print("a // b =", a // b)
print("a % b =", a % b)
print("a ** b =", a ** b)
```
Note that the multiplication is *not* a matrix multiplication. We will discuss matrix operations below.
The arrays must have the same shape. If they do not, NumPy will apply the *broadcasting rules*.
## Broadcasting
In general, when NumPy expects arrays of the same shape but finds that this is not the case, it applies the so-called *broadcasting* rules:
## First rule
*If the arrays do not have the same rank, then a 1 will be prepended to the smaller ranking arrays until their ranks match.*
```
h = np.arange(5).reshape(1, 1, 5)
h
```
Now let's try to add a 1D array of shape `(5,)` to this 3D array of shape `(1,1,5)`. Applying the first rule of broadcasting!
```
h + [10, 20, 30, 40, 50] # same as: h + [[[10, 20, 30, 40, 50]]]
```
## Second rule
*Arrays with a 1 along a particular dimension act as if they had the size of the array with the largest shape along that dimension. The value of the array element is repeated along that dimension.*
```
k = np.arange(6).reshape(2, 3)
k
```
Let's try to add a 2D array of shape `(2,1)` to this 2D `ndarray` of shape `(2, 3)`. NumPy will apply the second rule of broadcasting:
```
k + [[100], [200]] # same as: k + [[100, 100, 100], [200, 200, 200]]
```
Combining rules 1 & 2, we can do this:
```
k + [100, 200, 300] # after rule 1: [[100, 200, 300]], and after rule 2: [[100, 200, 300], [100, 200, 300]]
```
And also, very simply:
```
k + 1000 # same as: k + [[1000, 1000, 1000], [1000, 1000, 1000]]
```
## Third rule
*After rules 1 & 2, the sizes of all arrays must match.*
```
try:
k + [33, 44]
except ValueError as e:
print(e)
```
Broadcasting rules are used in many NumPy operations, not just arithmetic operations, as we will see below.
For more details about broadcasting, check out [the documentation](https://docs.scipy.org/doc/numpy-dev/user/basics.broadcasting.html).
## Upcasting
When trying to combine arrays with different `dtype`s, NumPy will *upcast* to a type capable of handling all possible values (regardless of what the *actual* values are).
```
k1 = np.arange(0, 5, dtype=np.uint8)
print(k1.dtype, k1)
k2 = k1 + np.array([5, 6, 7, 8, 9], dtype=np.int8)
print(k2.dtype, k2)
```
Note that `int16` is required to represent all *possible* `int8` and `uint8` values (from -128 to 255), even though in this case a uint8 would have sufficed.
```
k3 = k1 + 1.5
print(k3.dtype, k3)
```
## Conditional operators
The conditional operators also apply elementwise:
```
m = np.array([20, -5, 30, 40])
m < [15, 16, 35, 36]
```
And using broadcasting:
```
m < 25 # equivalent to m < [25, 25, 25, 25]
```
This is most useful in conjunction with boolean indexing (discussed below).
```
m[m < 25]
```
##Mathematical and statistical functions
Many mathematical and statistical functions are available for `ndarray`s.
## `ndarray` methods
Some functions are simply `ndarray` methods, for example:
```
a = np.array([[-2.5, 3.1, 7], [10, 11, 12]])
print(a)
print("mean =", a.mean())
```
Note that this computes the mean of all elements in the `ndarray`, regardless of its shape.
Here are a few more useful `ndarray` methods:
```
for func in (a.min, a.max, a.sum, a.prod, a.std, a.var):
print(func.__name__, "=", func())
```
These functions accept an optional argument `axis` which lets you ask for the operation to be performed on elements along the given axis. For example:
```
c=np.arange(24).reshape(2,3,4)
c
c.sum(axis=0) # sum across matrices
c.sum(axis=1) # sum across rows
```
You can also sum over multiple axes:
```
c.sum(axis=(0,2)) # sum across matrices and columns
0+1+2+3 + 12+13+14+15, 4+5+6+7 + 16+17+18+19, 8+9+10+11 + 20+21+22+23
```
## Universal functions
NumPy also provides fast elementwise functions called *universal functions*, or **ufunc**. They are vectorized wrappers of simple functions. For example `square` returns a new `ndarray` which is a copy of the original `ndarray` except that each element is squared:
```
a = np.array([[-2.5, 3.1, 7], [10, 11, 12]])
np.square(a)
```
Here are a few more useful unary ufuncs:
```
print("Original ndarray")
print(a)
for func in (np.abs, np.sqrt, np.exp, np.log, np.sign, np.ceil, np.modf, np.isnan, np.cos):
print("\n", func.__name__)
print(func(a))
```
## Binary ufuncs
There are also many binary ufuncs, that apply elementwise on two `ndarray`s. Broadcasting rules are applied if the arrays do not have the same shape:
```
a = np.array([1, -2, 3, 4])
b = np.array([2, 8, -1, 7])
np.add(a, b) # equivalent to a + b
np.greater(a, b) # equivalent to a > b
np.maximum(a, b)
np.copysign(a, b)
```
## Array indexing
## One-dimensional arrays
One-dimensional NumPy arrays can be accessed more or less like regular python arrays:
```
a = np.array([1, 5, 3, 19, 13, 7, 3])
a[3]
a[2:5]
a[2:-1]
a[:2]
a[2::2]
a[::-1]
```
Of course, you can modify elements:
```
a[3]=999
a
```
You can also modify an `ndarray` slice:
```
a[2:5] = [997, 998, 999]
a
```
## Differences with regular python arrays
Contrary to regular python arrays, if you assign a single value to an `ndarray` slice, it is copied across the whole slice, thanks to broadcasting rules discussed above.
```
a[2:5] = -1
a
```
Also, you cannot grow or shrink `ndarray`s this way:
```
try:
a[2:5] = [1,2,3,4,5,6] # too long
except ValueError as e:
print(e)
```
You cannot delete elements either:
```
try:
del a[2:5]
except ValueError as e:
print(e)
```
Last but not least, `ndarray` **slices are actually *views*** on the same data buffer. This means that if you create a slice and modify it, you are actually going to modify the original `ndarray` as well!
```
a_slice = a[2:6]
a_slice[1] = 1000
a # the original array was modified!
a[3] = 2000
a_slice # similarly, modifying the original array modifies the slice!
```
If you want a copy of the data, you need to use the `copy` method:
```
another_slice = a[2:6].copy()
another_slice[1] = 3000
a # the original array is untouched
a[3] = 4000
another_slice # similary, modifying the original array does not affect the slice copy
```
## Multi-dimensional arrays
Multi-dimensional arrays can be accessed in a similar way by providing an index or slice for each axis, separated by commas:
```
b = np.arange(48).reshape(4, 12)
b
b[1, 2] # row 1, col 2
b[1, :] # row 1, all columns
b[:, 1] # all rows, column 1
```
**Caution**: note the subtle difference between these two expressions:
```
b[1, :]
b[1:2, :]
```
The first expression returns row 1 as a 1D array of shape `(12,)`, while the second returns that same row as a 2D array of shape `(1, 12)`.
## Fancy indexing
You may also specify a list of indices that you are interested in. This is referred to as *fancy indexing*.
```
b[(0,2), 2:5] # rows 0 and 2, columns 2 to 4 (5-1)
b[:, (-1, 2, -1)] # all rows, columns -1 (last), 2 and -1 (again, and in this order)
```
If you provide multiple index arrays, you get a 1D `ndarray` containing the values of the elements at the specified coordinates.
```
b[(-1, 2, -1, 2), (5, 9, 1, 9)] # returns a 1D array with b[-1, 5], b[2, 9], b[-1, 1] and b[2, 9] (again)
```
## Higher dimensions
Everything works just as well with higher dimensional arrays, but it's useful to look at a few examples:
```
c = b.reshape(4,2,6)
c
c[2, 1, 4] # matrix 2, row 1, col 4
c[2, :, 3] # matrix 2, all rows, col 3
```
If you omit coordinates for some axes, then all elements in these axes are returned:
```
c[2, 1] # Return matrix 2, row 1, all columns. This is equivalent to c[2, 1, :]
```
## Ellipsis (`...`)
You may also write an ellipsis (`...`) to ask that all non-specified axes be entirely included.
```
c[2, ...] # matrix 2, all rows, all columns. This is equivalent to c[2, :, :]
c[2, 1, ...] # matrix 2, row 1, all columns. This is equivalent to c[2, 1, :]
c[2, ..., 3] # matrix 2, all rows, column 3. This is equivalent to c[2, :, 3]
c[..., 3] # all matrices, all rows, column 3. This is equivalent to c[:, :, 3]
```
## Boolean indexing
You can also provide an `ndarray` of boolean values on one axis to specify the indices that you want to access.
```
b = np.arange(48).reshape(4, 12)
b
rows_on = np.array([True, False, True, False])
b[rows_on, :] # Rows 0 and 2, all columns. Equivalent to b[(0, 2), :]
cols_on = np.array([False, True, False] * 4)
b[:, cols_on] # All rows, columns 1, 4, 7 and 10
```
## `np.ix_`
You cannot use boolean indexing this way on multiple axes, but you can work around this by using the `ix_` function:
```
b[np.ix_(rows_on, cols_on)]
np.ix_(rows_on, cols_on)
```
If you use a boolean array that has the same shape as the `ndarray`, then you get in return a 1D array containing all the values that have `True` at their coordinate. This is generally used along with conditional operators:
```
b[b % 3 == 1]
```
## Iterating
Iterating over `ndarray`s is very similar to iterating over regular python arrays. Note that iterating over multidimensional arrays is done with respect to the first axis.
```
c = np.arange(24).reshape(2, 3, 4) # A 3D array (composed of two 3x4 matrices)
c
for m in c:
print("Item:")
print(m)
for i in range(len(c)): # Note that len(c) == c.shape[0]
print("Item:")
print(c[i])
```
If you want to iterate on *all* elements in the `ndarray`, simply iterate over the `flat` attribute:
```
for i in c.flat:
print("Item:", i)
```
##Stacking arrays
It is often useful to stack together different arrays. NumPy offers several functions to do just that. Let's start by creating a few arrays.
```
q1 = np.full((3,4), 1.0)
q1
q2 = np.full((4,4), 2.0)
q2
q3 = np.full((3,4), 3.0)
q3
```
## `vstack`
Now let's stack them vertically using `vstack`:
```
q4 = np.vstack((q1, q2, q3))
q4
q4.shape
```
This was possible because q1, q2 and q3 all have the same shape (except for the vertical axis, but that's ok since we are stacking on that axis).
## `hstack`
We can also stack arrays horizontally using `hstack`:
```
q5 = np.hstack((q1, q3))
q5
q5.shape
```
This is possible because q1 and q3 both have 3 rows. But since q2 has 4 rows, it cannot be stacked horizontally with q1 and q3:
```
try:
q5 = np.hstack((q1, q2, q3))
except ValueError as e:
print(e)
```
## `concatenate`
The `concatenate` function stacks arrays along any given existing axis.
```
q7 = np.concatenate((q1, q2, q3), axis=0) # Equivalent to vstack
q7
q7.shape
```
As you might guess, `hstack` is equivalent to calling `concatenate` with `axis=1`.
## `stack`
The `stack` function stacks arrays along a new axis. All arrays have to have the same shape.
```
q8 = np.stack((q1, q3))
q8
q8.shape
```
##Splitting arrays
Splitting is the opposite of stacking. For example, let's use the `vsplit` function to split a matrix vertically.
First let's create a 6x4 matrix:
```
r = np.arange(24).reshape(6,4)
r
```
Now let's split it in three equal parts, vertically:
```
r1, r2, r3 = np.vsplit(r, 3)
r1
r2
r3
```
There is also a `split` function which splits an array along any given axis. Calling `vsplit` is equivalent to calling `split` with `axis=0`. There is also an `hsplit` function, equivalent to calling `split` with `axis=1`:
```
r4, r5 = np.hsplit(r, 2)
r4
r5
```
##Transposing arrays
The `transpose` method creates a new view on an `ndarray`'s data, with axes permuted in the given order.
For example, let's create a 3D array:
```
t = np.arange(24).reshape(4,2,3)
t
```
Now let's create an `ndarray` such that the axes `0, 1, 2` (depth, height, width) are re-ordered to `1, 2, 0` (depth→width, height→depth, width→height):
```
t1 = t.transpose((1,2,0))
t1
t1.shape
```
By default, `transpose` reverses the order of the dimensions:
```
t2 = t.transpose() # equivalent to t.transpose((2, 1, 0))
t2
t2.shape
```
NumPy provides a convenience function `swapaxes` to swap two axes. For example, let's create a new view of `t` with depth and height swapped:
```
t3 = t.swapaxes(0,1) # equivalent to t.transpose((1, 0, 2))
t3
t3.shape
```
## Linear algebra
NumPy 2D arrays can be used to represent matrices efficiently in python. We will just quickly go through some of the main matrix operations available. For more details about Linear Algebra, vectors and matrics, go through the [Linear Algebra tutorial](math_linear_algebra.ipynb).
## Matrix transpose
The `T` attribute is equivalent to calling `transpose()` when the rank is ≥2:
```
m1 = np.arange(10).reshape(2,5)
m1
m1.T
```
The `T` attribute has no effect on rank 0 (empty) or rank 1 arrays:
```
m2 = np.arange(5)
m2
m2.T
```
We can get the desired transposition by first reshaping the 1D array to a single-row matrix (2D):
```
m2r = m2.reshape(1,5)
m2r
m2r.T
```
## Matrix multiplication
Let's create two matrices and execute a [matrix multiplication](https://en.wikipedia.org/wiki/Matrix_multiplication) using the `dot()` method.
```
n1 = np.arange(10).reshape(2, 5)
n1
n2 = np.arange(15).reshape(5,3)
n2
n1.dot(n2)
```
**Caution**: as mentionned previously, `n1*n2` is *not* a matric multiplication, it is an elementwise product (also called a [Hadamard product](https://en.wikipedia.org/wiki/Hadamard_product_(matrices))).
## Matrix inverse and pseudo-inverse
Many of the linear algebra functions are available in the `numpy.linalg` module, in particular the `inv` function to compute a square matrix's inverse:
```
import numpy.linalg as linalg
m3 = np.array([[1,2,3],[5,7,11],[21,29,31]])
m3
linalg.inv(m3)
```
You can also compute the [pseudoinverse](https://en.wikipedia.org/wiki/Moore%E2%80%93Penrose_pseudoinverse) using `pinv`:
```
linalg.pinv(m3)
```
## Identity matrix
The product of a matrix by its inverse returns the identiy matrix (with small floating point errors):
```
m3.dot(linalg.inv(m3))
```
You can create an identity matrix of size NxN by calling `eye`:
```
np.eye(3)
```
## QR decomposition
The `qr` function computes the [QR decomposition](https://en.wikipedia.org/wiki/QR_decomposition) of a matrix:
```
q, r = linalg.qr(m3)
q
r
q.dot(r) # q.r equals m3
```
## Determinant
The `det` function computes the [matrix determinant](https://en.wikipedia.org/wiki/Determinant):
```
linalg.det(m3) # Computes the matrix determinant
```
## Eigenvalues and eigenvectors
The `eig` function computes the [eigenvalues and eigenvectors](https://en.wikipedia.org/wiki/Eigenvalues_and_eigenvectors) of a square matrix:
```
eigenvalues, eigenvectors = linalg.eig(m3)
eigenvalues # λ
eigenvectors # v
m3.dot(eigenvectors) - eigenvalues * eigenvectors # m3.v - λ*v = 0
```
## Singular Value Decomposition
The `svd` function takes a matrix and returns its [singular value decomposition](https://en.wikipedia.org/wiki/Singular_value_decomposition):
```
m4 = np.array([[1,0,0,0,2], [0,0,3,0,0], [0,0,0,0,0], [0,2,0,0,0]])
m4
U, S_diag, V = linalg.svd(m4)
U
S_diag
```
The `svd` function just returns the values in the diagonal of Σ, but we want the full Σ matrix, so let's create it:
```
S = np.zeros((4, 5))
S[np.diag_indices(4)] = S_diag
S # Σ
V
U.dot(S).dot(V) # U.Σ.V == m4
```
## Diagonal and trace
```
np.diag(m3) # the values in the diagonal of m3 (top left to bottom right)
np.trace(m3) # equivalent to np.diag(m3).sum()
```
## Solving a system of linear scalar equations
The `solve` function solves a system of linear scalar equations, such as:
* $2x + 6y = 6$
* $5x + 3y = -9$
```
coeffs = np.array([[2, 6], [5, 3]])
depvars = np.array([6, -9])
solution = linalg.solve(coeffs, depvars)
solution
```
Let's check the solution:
```
coeffs.dot(solution), depvars # yep, it's the same
```
Looks good! Another way to check the solution:
```
np.allclose(coeffs.dot(solution), depvars)
```
## Vectorization
Instead of executing operations on individual array items, one at a time, your code is much more efficient if you try to stick to array operations. This is called *vectorization*. This way, you can benefit from NumPy's many optimizations.
For example, let's say we want to generate a 768x1024 array based on the formula $sin(xy/40.5)$. A **bad** option would be to do the math in python using nested loops:
```
import math
data = np.empty((768, 1024))
for y in range(768):
for x in range(1024):
data[y, x] = math.sin(x*y/40.5) # BAD! Very inefficient.
```
Sure, this works, but it's terribly inefficient since the loops are taking place in pure python. Let's vectorize this algorithm. First, we will use NumPy's `meshgrid` function which generates coordinate matrices from coordinate vectors.
```
x_coords = np.arange(0, 1024) # [0, 1, 2, ..., 1023]
y_coords = np.arange(0, 768) # [0, 1, 2, ..., 767]
X, Y = np.meshgrid(x_coords, y_coords)
X
Y
```
As you can see, both `X` and `Y` are 768x1024 arrays, and all values in `X` correspond to the horizontal coordinate, while all values in `Y` correspond to the the vertical coordinate.
Now we can simply compute the result using array operations:
```
data = np.sin(X*Y/40.5)
```
Now we can plot this data using matplotlib's `imshow` function (see the [matplotlib tutorial](tools_matplotlib.ipynb)).
```
import matplotlib.pyplot as plt
import matplotlib.cm as cm
fig = plt.figure(1, figsize=(7, 6))
plt.imshow(data, cmap=cm.hot, interpolation="bicubic")
plt.show()
```
## Saving and loading
NumPy makes it easy to save and load `ndarray`s in binary or text format.
## Binary `.npy` format
Let's create a random array and save it.
```
a = np.random.rand(2,3)
a
np.save("my_array", a)
```
Done! Since the file name contains no file extension was provided, NumPy automatically added `.npy`. Let's take a peek at the file content:
```
with open("my_array.npy", "rb") as f:
content = f.read()
content
```
To load this file into a NumPy array, simply call `load`:
```
a_loaded = np.load("my_array.npy")
a_loaded
```
## Text format
Let's try saving the array in text format:
```
np.savetxt("my_array.csv", a)
```
Now let's look at the file content:
```
with open("my_array.csv", "rt") as f:
print(f.read())
```
This is a CSV file with tabs as delimiters. You can set a different delimiter:
```
np.savetxt("my_array.csv", a, delimiter=",")
```
To load this file, just use `loadtxt`:
```
a_loaded = np.loadtxt("my_array.csv", delimiter=",")
a_loaded
```
## Zipped `.npz` format
It is also possible to save multiple arrays in one zipped file:
```
b = np.arange(24, dtype=np.uint8).reshape(2, 3, 4)
b
np.savez("my_arrays", my_a=a, my_b=b)
```
Again, let's take a peek at the file content. Note that the `.npz` file extension was automatically added.
```
with open("my_arrays.npz", "rb") as f:
content = f.read()
repr(content)[:180] + "[...]"
```
You then load this file like so:
```
my_arrays = np.load("my_arrays.npz")
my_arrays
```
This is a dict-like object which loads the arrays lazily:
```
my_arrays.keys()
my_arrays["my_a"]
```
## What next?
Now you know all the fundamentals of NumPy, but there are many more options available. The best way to learn more is to experiment with NumPy, and go through the excellent [reference documentation](http://docs.scipy.org/doc/numpy/reference/index.html) to find more functions and features you may be interested in.
| github_jupyter |
```
import pandas as pd
import glob
import matplotlib.pyplot as plt
import datetime, sys
import numpy as np
%matplotlib inline
def parse(t):
string_ = str(t)
try:
return datetime.date(int(string_[:4]), int(string_[4:6]), int(string_[6:]))
except:
return datetime.date(1900,1,1)
def readAllFiles():
allFiles = glob.iglob("data/atp_rankings_" + "*.csv")
ranks = pd.DataFrame()
list_ = list()
for filen in allFiles:
df = pd.read_csv(filen,
index_col=None,
header=None,
parse_dates=[0],
date_parser=lambda t:parse(t),
names = ["ranking_date", "ranking", "player_id", "ranking_points"])
list_.append(df)
ranks = pd.concat(list_)
return ranks
def readPlayers():
return pd.read_csv("data/atp_players.csv",
index_col=None,
header=None,
parse_dates=[4],
names = ["player_id", "first_name", "last_name", "hand", "birth_date", "country_code"],
date_parser=lambda t:parse(t))
# merge rankings with players but keep only top 100
ranks = readAllFiles()
ranks = ranks[(ranks['ranking']<100)]
ranks['player_id'] = ranks['player_id'].apply(lambda row: int(row))
players = readPlayers()
players = ranks.merge(players,right_on="player_id",left_on="player_id")
players.head()
# we are interested in ranking date, country code and counting the number of players.
# To keep the data cleaner get only countries with at least 4 players in top 100
aggregate = players[["ranking_date", "country_code"]].groupby(["ranking_date","country_code"]).size().to_frame()
aggregate = pd.DataFrame(aggregate.to_records()) # transform the series to df
aggregate_least_4 = aggregate[(aggregate["0"] > 3)]
aggregate_least_10 = aggregate[(aggregate["0"] > 9)]
# countries with at least 10 players in top 10 at a given time.
# we can see that the US has dominated until 1997, when Spain started to take the lead.
# Since 2000, Spain and France had the most players in top 100.
pivoted = pd.pivot_table(aggregate_least_10, values='0', columns='country_code', index="ranking_date")
pivoted.plot(figsize=(20,10))
# Let's see that figure again without the outlier USA
aggregate_no_usa = aggregate[(aggregate["country_code"] != "USA") & (aggregate["0"] > 9)]
pivoted = pd.pivot_table(aggregate_no_usa, values='0', columns='country_code', index="ranking_date")
pivoted.plot(figsize=(20,10))
# after 2000 Spain and France dominate with an average of 15 players in top 100
aggregate_after_2000 = aggregate[(aggregate["ranking_date"] > "2000-01-01" ) & (aggregate["0"] > 9)]
pivoted = pd.pivot_table(aggregate_after_2000, values='0', columns='country_code', index="ranking_date")
pivoted.plot(figsize=(20,10))
# let's see how it fairs by continent
import json
with open('iso3.json') as data_file:
countries = json.load(data_file)
with open('continent.json') as data_file:
continents = json.load(data_file)
# we can see that North America dominated in the 80s after which Europe took lead.
# Now North America is on par with South America and Asia.
ioc_countries = {}
for country in countries:
if country["ioc"]:
ioc_countries[country["ioc"]] = country["alpha2"]
ioc_countries["YUG"] = "MK"
aggregate_continents = aggregate.copy()
ioc_countries
aggregate_continents
aggregate_continents["country_code"] = aggregate_continents["country_code"].apply(lambda row: continents[ioc_countries[row]])
aggregate_continents = aggregate_continents[["ranking_date", "country_code", "0"]].groupby(["ranking_date", "country_code"]).sum()
aggregate_continents = pd.DataFrame(aggregate_continents.to_records()) # transform the series to df
pivoted = pd.pivot_table(aggregate_continents, values='0', columns='country_code', index="ranking_date")
pivoted.plot(figsize=(20,10))
```
| github_jupyter |
# Multithreading
```
import sys
import queue
import threading
sys.path.append("../")
from IoTPy.concurrency.multithread import iThread
from IoTPy.core.stream import Stream
def test_multithread_1():
from IoTPy.agent_types.sink import sink_element, stream_to_queue
# The thread target that reads output queues and puts the results
# in a list.
def output_thread_target(q_out, output):
while True:
try:
w = q_out.get()
if w == '_finished': break
else: output.append(w)
except:
time.sleep(0.0001)
# Declare output queue q of the IoTPy thread.
x_q = queue.Queue()
# Create threads to read output queues of IoTPy thread.
x_output = []
output_thread = threading.Thread(
target=output_thread_target,
args=(x_q, x_output))
# Declare streams
x = Stream('x')
# Declare agents
stream_to_queue(x, x_q)
# Create thread: This is the IoTPy thread. This thread
# reads a single input stream x and puts results of its
# computations in queue, x_q.
ithread = iThread(in_streams=[x], output_queues=[x_q])
# Start threads.
ithread.start()
output_thread.start()
# Put data into streams.
# in_stream_name is the name of the input stream which is
# extended.
test_data = list(range(5))
ithread.extend(in_stream_name='x', list_of_elements=test_data)
# Indicate stream 'x' will get no further input.
ithread.terminate(in_stream_name='x')
# Join threads
ithread.thread.join()
output_thread.join()
# Check output
assert x_output == test_data
print (x_output)
test_multithread_1()
def test_multithread_2():
from IoTPy.agent_types.sink import sink_element, stream_to_queue
from IoTPy.agent_types.merge import zip_map, zip_stream
from IoTPy.agent_types.op import map_element
# The thread target that reads output queues and puts the results
# in a list.
def output_thread_target(q_out, output):
while True:
try:
w = q_out.get()
if w == '_finished': break
else: output.append(w)
except:
time.sleep(0.0001)
# Declare output queues
z_q = queue.Queue()
b_q = queue.Queue()
# Create threads to read output queues of IoTPy thread.
z_output = []
z_output_thread = threading.Thread(
target=output_thread_target,
args=(z_q, z_output))
b_output = []
b_output_thread = threading.Thread(
target=output_thread_target,
args=(b_q, b_output))
# Declare streams
x = Stream('x')
y = Stream('y')
z = Stream('z')
a = Stream('a')
b = Stream('b')
# Declare agents
zip_stream(in_streams=[x, y], out_stream=z)
stream_to_queue(z, z_q)
def g(v): return 2* v
map_element(func=g, in_stream=a, out_stream=b)
stream_to_queue(b, b_q)
# Create threads
ithread_1 = iThread(in_streams=[x,y], output_queues=[z_q])
ithread_2 = iThread(in_streams=[a], output_queues=[b_q])
# Start threads.
ithread_1.start()
ithread_2.start()
z_output_thread.start()
b_output_thread.start()
# Put data into streams.
x_data = list(range(5))
y_data = list(range(100, 105))
a_data = list(range(1000, 1008))
ithread_1.extend(in_stream_name='x', list_of_elements=x_data)
ithread_1.extend(in_stream_name='y', list_of_elements=y_data)
ithread_2.extend(in_stream_name='a', list_of_elements=a_data)
# Indicate stream is finished
ithread_1.terminate('x')
ithread_1.terminate('y')
ithread_2.terminate('a')
# Join threads
ithread_1.join()
ithread_2.join()
z_output_thread.join()
b_output_thread.join()
# Check output
assert z_output == list(zip(x_data, y_data))
assert b_output == [g(v) for v in a_data]
print (z_output)
test_multithread_2()
```
| github_jupyter |
# SGDRegressor with Power Transformer
This Code template is for regression analysis using the SGDRegressor where feature transformation is done using PowerTransformer.
### Required Packages
```
import warnings
import numpy as np
import pandas as pd
import seaborn as se
import matplotlib.pyplot as plt
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import PowerTransformer
from sklearn.model_selection import train_test_split
from sklearn.metrics import r2_score, mean_absolute_error, mean_squared_error
from sklearn.linear_model import SGDRegressor
warnings.filterwarnings('ignore')
```
### Initialization
Filepath of CSV file
```
#filepath
file_path= ""
```
List of features which are required for model training.
```
#x_values
features=[]
```
Target feature for prediction.
```
#y_value
target=''
```
### Data Fetching
Pandas is an open-source, BSD-licensed library providing high-performance, easy-to-use data manipulation and data analysis tools.
We will use panda's library to read the CSV file using its storage path.And we use the head function to display the initial row or entry.
```
df=pd.read_csv(file_path)
df.head()
```
### Feature Selections
It is the process of reducing the number of input variables when developing a predictive model. Used to reduce the number of input variables to both reduce the computational cost of modelling and, in some cases, to improve the performance of the model.
We will assign all the required input features to X and target/outcome to Y.
```
X=df[features]
Y=df[target]
```
### Data Preprocessing
Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
```
def NullClearner(df):
if(isinstance(df, pd.Series) and (df.dtype in ["float64","int64"])):
df.fillna(df.mean(),inplace=True)
return df
elif(isinstance(df, pd.Series)):
df.fillna(df.mode()[0],inplace=True)
return df
else:return df
def EncodeX(df):
return pd.get_dummies(df)
```
Calling preprocessing functions on the feature and target set.
```
x=X.columns.to_list()
for i in x:
X[i]=NullClearner(X[i])
X=EncodeX(X)
Y=NullClearner(Y)
X.head()
```
#### Correlation Map
In order to check the correlation between the features, we will plot a correlation matrix. It is effective in summarizing a large amount of data where the goal is to see patterns.
```
f,ax = plt.subplots(figsize=(18, 18))
matrix = np.triu(X.corr())
se.heatmap(X.corr(), annot=True, linewidths=.5, fmt= '.1f',ax=ax, mask=matrix)
plt.show()
```
### Data Splitting
The train-test split is a procedure for evaluating the performance of an algorithm. The procedure involves taking a dataset and dividing it into two subsets. The first subset is utilized to fit/train the model. The second subset is used for prediction. The main motive is to estimate the performance of the model on new data.
```
x_train,x_test,y_train,y_test=train_test_split(X,Y,test_size=0.2,random_state=123)
```
### Model
Stochastic Gradient Descent (SGD) is a simple yet very efficient approach to fitting linear classifiers and regressors under convex loss functions such as (linear) Support Vector Machines and Logistic Regression. SGD is merely an optimization technique and does not correspond to a specific family of machine learning models. It is only a way to train a model. Often, an instance of SGDClassifier or SGDRegressor will have an equivalent estimator in the scikit-learn API, potentially using a different optimization technique.
For example, using SGDRegressor(loss='squared_loss', penalty='l2') and Ridge solve the same optimization problem, via different means.
#### Model Tuning Parameters
> - **loss** -> The loss function to be used. The possible values are ‘squared_loss’, ‘huber’, ‘epsilon_insensitive’, or ‘squared_epsilon_insensitive’
> - **penalty** -> The penalty (aka regularization term) to be used. Defaults to ‘l2’ which is the standard regularizer for linear SVM models. ‘l1’ and ‘elasticnet’ might bring sparsity to the model (feature selection) not achievable with ‘l2’.
> - **alpha** -> Constant that multiplies the regularization term. The higher the value, the stronger the regularization. Also used to compute the learning rate when set to learning_rate is set to ‘optimal’.
> - **l1_ratio** -> The Elastic Net mixing parameter, with 0 <= l1_ratio <= 1. l1_ratio=0 corresponds to L2 penalty, l1_ratio=1 to L1. Only used if penalty is ‘elasticnet’.
> - **tol** -> The stopping criterion
> - **learning_rate** -> The learning rate schedule,possible values {'optimal','constant','invscaling','adaptive'}
> - **eta0** -> The initial learning rate for the ‘constant’, ‘invscaling’ or ‘adaptive’ schedules.
> - **power_t** -> The exponent for inverse scaling learning rate.
> - **epsilon** -> Epsilon in the epsilon-insensitive loss functions; only if loss is ‘huber’, ‘epsilon_insensitive’, or ‘squared_epsilon_insensitive’.
### Power Transformer
Power transforms are a family of parametric, monotonic transformations that are applied to make data more Gaussian-like. This is useful for modeling issues related to heteroscedasticity (non-constant variance), or other situations where normality is desired.
Currently, PowerTransformer supports the Box-Cox transform and the Yeo-Johnson transform. The optimal parameter for stabilizing variance and minimizing skewness is estimated through maximum likelihood.
Apply a power transform featurewise to make data more Gaussian-like.
```
model=make_pipeline(PowerTransformer(), SGDRegressor(random_state=123))
model.fit(x_train,y_train)
```
#### Model Accuracy
We will use the trained model to make a prediction on the test set.Then use the predicted value for measuring the accuracy of our model.
score: The score function returns the coefficient of determination R2 of the prediction.
```
print("Accuracy score {:.2f} %\n".format(model.score(x_test,y_test)*100))
```
> **r2_score**: The **r2_score** function computes the percentage variablility explained by our model, either the fraction or the count of correct predictions.
> **mae**: The **mean abosolute error** function calculates the amount of total error(absolute average distance between the real data and the predicted data) by our model.
> **mse**: The **mean squared error** function squares the error(penalizes the model for large errors) by our model.
```
y_pred=model.predict(x_test)
print("R2 Score: {:.2f} %".format(r2_score(y_test,y_pred)*100))
print("Mean Absolute Error {:.2f}".format(mean_absolute_error(y_test,y_pred)))
print("Mean Squared Error {:.2f}".format(mean_squared_error(y_test,y_pred)))
```
#### Prediction Plot
First, we make use of a plot to plot the actual observations, with x_train on the x-axis and y_train on the y-axis.
For the regression line, we will use x_train on the x-axis and then the predictions of the x_train observations on the y-axis.
```
n=len(x_test) if len(x_test)<20 else 20
plt.figure(figsize=(14,10))
plt.plot(range(n),y_test[0:n], color = "green")
plt.plot(range(n),model.predict(x_test[0:n]), color = "red")
plt.legend(["Actual","prediction"])
plt.title("Predicted vs True Value")
plt.xlabel("Record number")
plt.ylabel(target)
plt.show()
```
#### Creator: Ayush Gupta , Github: [Profile](https://github.com/guptayush179)
| github_jupyter |
# Import Libraries
```
import requests#Connect to endpoints
import time #time script
import sys
import csv
import json
import pandas as pd
import spotipy#authentication
import spotipy.util as util#authentication
from spotipy.oauth2 import SpotifyClientCredentials#authentication
```
# declaring credentials
```
cid = 'f61689bf814a44dab11d7e63d18dc38a'
secret = '7d2c54296b204a20b26b1c7eb5a93601'
redirect_uri = 'http://google.com/'
username = '31tabw4566sgjjakrw5afmpspdv4'
```
# authorization flow
```
scope = 'user-library-read'
token = util.prompt_for_user_token(username, scope, client_id=cid, client_secret=secret, redirect_uri=redirect_uri)
if token:
sp = spotipy.Spotify(auth=token)
else:
print("Can't get token for", username)
```
# Global list for Tracks
```
#Move tracks list to first line of Extracting Liked Tracks Cell
#Use this global list in the get_tracks function
tracks = []
```
# Retrieving Like Tracks
```
#Add all_parameter arguement to get_user_tracks
def get_user_tracks(nexturl,show=True,all_parameters=False):
#tracks = [] #list tracks
try:
resp = requests.get(url=nexturl,
headers={'Authorization': 'Bearer ' + token})
resp.raise_for_status()
except requests.exceptions.HTTPError as err:
print(err)
response = resp.json()
track_limit = (response['limit'])-1
for x in range(track_limit):
try:
track_uri = response['items'][x]['track']['uri']
track_name = response['items'][x]['track']['name']
if all_parameters:
#Add artist_uri
#Add artist_name
#Add album_uri
#Add album_name
#To find path go to this link and fill in with data sample
#and paste the json into https://jsonformatter.org/json-editor
#append to dictionary add all variables under if condition
#tracks.append({'track_uri':track_uri,'track_name':track_name})
else:
#Change note for consistency
#for track in pmtracks:
tracks.append({'track_uri':track_uri,'track_name':track_name})
if show:
print(track_uri, track_name)
except IndexError as error:
continue
try:
if (nexturl is not None):
get_user_tracks(response['next'])
else:
return
except:
return tracks
```
# Extracting Liked Tracks
```
#Rename pmtracks
#Add all_parameters as boolean value as a parameter to get_user_tracks
start = time.time()
#output tracks
pmtracks = get_user_tracks('https://api.spotify.com/v1/me/tracks?limit=50', show=True,all_parameters=True)
stop = time.time()
duration = (stop - start)/60
#should be NoneType
type(pmtracks)
str(duration)+' minutes'
#If a dataframe doesn't show make sure there is an append to tracks in the get_tracks function
#If your tracks are way larger than the number next to offeset in the links rereun the global track cell and rerun
df = pd.DataFrame(tracks)
df
#Should be found in the Data folder in the repo
df.to_csv('Data/user_track_uri.csv',sep='|',index=False)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/mnassar/segfault/blob/main/SegFault_Segmentation_examples.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Segmentation Fault: A cheap defense against adversarial machine learning
## Table III, IV and V
## Authors: Mohamed Nassar, Doha Al Bared
### Department of Computer Science
### AUB
```
!wget --no-check-certificate 'https://docs.google.com/uc?export=download&id=1H4KEE0Vp8DFZOe_QfcxqOxEVnpun-uka' -O CIFAR10model.h5
# load the cifar classifier
from tensorflow.keras.models import load_model
pretrained_model = load_model('CIFAR10model.h5')
pretrained_model.trainable = False
pretrained_model.summary()
```
# CIFAR 10
```
#get dataset: cifar10
import tensorflow_datasets as tfds
from keras.datasets import cifar10
(ds_train, ds_test), ds_info = tfds.load(
'cifar10',
split=['train', 'test'],
shuffle_files=True,
as_supervised=True,
with_info=True,
)
print("-------------")
print (ds_info)
import tensorflow.compat.v2 as tf
tf.enable_v2_behavior()
# normalize images
# these are the numbers used during training the model
mean = 120.70748
std = 64.150024
bound_min = (0-mean)/std
bound_max = (255-mean)/std
class_names = ['airplane', 'automobile', 'bird', 'cat', 'deer',
'dog', 'frog', 'horse', 'ship', 'truck']
def normalize_img(image, label):
"""Normalizes images: `uint8` -> `float32`."""
# return tf.cast(image, tf.float32) / 255., tf.one_hot(label, 10)
return (tf.cast(image, tf.float32) - mean) / std, tf.one_hot(label, 10)
ds_train = ds_train.map(
normalize_img, num_parallel_calls=tf.data.experimental.AUTOTUNE)
ds_train = ds_train.cache()
ds_train = ds_train.shuffle(ds_info.splits['train'].num_examples)
ds_train = ds_train.batch(128)
ds_train = ds_train.prefetch(tf.data.experimental.AUTOTUNE)
ds_test = ds_test.map(
normalize_img, num_parallel_calls=tf.data.experimental.AUTOTUNE)
ds_test = ds_test.batch(128)
ds_test = ds_test.cache()
ds_test = ds_test.prefetch(tf.data.experimental.AUTOTUNE)
iter_ds_train = iter(ds_train)
import matplotlib.pyplot as plt
import numpy as np
images, labels = next(iter_ds_train)
preds = pretrained_model.predict(images)
img = images[0]
label = labels[0]
pred = preds[0]
img = img.numpy()
plt.figure(figsize = (1,1))
plt.axis('off')
plt.imshow((img * std + mean).astype(np.uint8))
plt.show()
print (class_names[np.argmax(label)], class_names[np.argmax(pred)])
```
# Felzenszwalb
```
from skimage.segmentation import felzenszwalb, slic, quickshift
from skimage.segmentation import mark_boundaries
from skimage.util import img_as_float
images, labels = next(iter_ds_train)
preds = pretrained_model.predict(images)
img = images[0]
label = labels[0]
pred = preds[0]
img = img.numpy()
for s in [1,10,100, 1000]:
segImg = felzenszwalb(img, scale=s)
print("Felzenszwalb's number of segments: %d" % len(np.unique(segImg)))
plt.figure(figsize = (1,1))
plt.axis('off')
plt.imshow(mark_boundaries((img * std + mean).astype(np.uint8), segImg))
plt.show()
print (class_names[np.argmax(label)], class_names[np.argmax(pred)])
```
# QuickShift
```
d={}
images, labels = next(iter_ds_train)
preds = pretrained_model.predict(images)
img = images[0]
label = labels[0]
pred = preds[0]
img = img.numpy()
d[class_names[np.argmax(label)]] = {}
for s in [0,1,2]:
for m in [10, 20]:
segImg = quickshift(img, sigma=s, max_dist=m)
print("Quick Shift's number of segments: %d" % len(np.unique(segImg)))
d[class_names[np.argmax(label)]][(s,m)] =len(np.unique(segImg))
plt.figure(figsize = (1,1))
plt.axis('off')
plt.imshow(mark_boundaries((img * std + mean).astype(np.uint8), segImg))
plt.show()
print (class_names[np.argmax(label)], class_names[np.argmax(pred)])
import pprint
pp = pprint.PrettyPrinter(indent=4)
pp.pprint(d)
```
# SLIC
```
images, labels = next(iter_ds_train)
preds = pretrained_model.predict(images)
img = images[0]
label = labels[0]
pred = preds[0]
img = img.numpy()
for n in [1, 32, 64, 128]:
segImg = slic(img, n_segments=n)
print("SLIC number of segments: %d" % len(np.unique(segImg)))
plt.figure(figsize = (1,1))
plt.axis('off')
plt.imshow(mark_boundaries((img * std + mean).astype(np.uint8), segImg))
plt.show()
print (class_names[np.argmax(label)], class_names[np.argmax(pred)])
```
| github_jupyter |
```
!pip install --user qctrl
!pip install --user qctrl-visualizer qctrl-open-controls
import matplotlib.pyplot as plt
import numpy as np
from qctrlvisualizer import get_qctrl_style, plot_controls
from qctrl import Qctrl
qctrl = Qctrl()
```
# The Challenge
```
control_count = 5
segment_count = 256
duration = 30.0
shot_count = 32
controls = []
for k in range(control_count):
# Create a random string of complex numbers for each controls.
real_part = np.random.random(size=[segment_count])
imag_part = np.random.random(size=[segment_count])
values = 0.15 * k * (real_part + 1j * imag_part)
controls.append({"duration": duration, "values": values})
# Plot the last control as an example.
plot_controls(
figure=plt.figure(),
controls={
"$\Omega$": [
{"duration": duration / segment_count, "value": value} for value in values
]
},
)
experiment_results = qctrl.functions.calculate_qchack_measurements(
controls=controls,
shot_count=shot_count,
)
```
# Band-Limited Pulses
```
sigma_x = np.array([[0, 1], [1, 0]])
sigma_z = np.array([[1, 0], [0, -1]])
sigma_y = np.array([[0, -1j], [1j, 0]])
ideal_not_gate = np.array([[0, 1], [1, 0]])
ideal_h_gate = (1 / np.sqrt(2)) * np.array([[1, -1j], [-1j, 1]])
# Define physical constraints
alpha_max = 2 * np.pi * 8.5e6 # Hz
sinc_cutoff_frequency = 1 * np.pi * 48e6 # Hz
nu = 2 * np.pi * 6e6 # Hz
segment_count = 50
duration = 400e-9 # s
experiment_results = qctrl.functions.calculate_qchack_measurements(
controls=controls,
shot_count=shot_count,
)
def get_optimized_matrix(target_operator,repetitions=1,shots=1024):
# 5. SPAM error confusion matrix
confusion_matrix = np.array([[0.99, 0.01], [0.02, 0.98]])
with qctrl.create_graph() as graph:
# Create initial alpha_1(t) signal
alpha_1_values = qctrl.operations.bounded_optimization_variable(
count=segment_count,
lower_bound=-alpha_max,
upper_bound=alpha_max,
)
alpha_1 = qctrl.operations.pwc_signal(
values=alpha_1_values,
duration=duration,
)
# Create filtered signal
alpha_1_filtered = qctrl.operations.convolve_pwc(
alpha_1,
qctrl.operations.sinc_integral_function(sinc_cutoff_frequency),
)
# Discretize to obtain smoothed alpha_1(t) signal
alpha_1 = qctrl.operations.discretize_stf(
stf=alpha_1_filtered,
duration=duration,
segments_count=segment_count,
name="alpha_1",
)
# Similarly, create alpha_2(t) signal
alpha_2_values = qctrl.operations.bounded_optimization_variable(
count=segment_count,
lower_bound=-alpha_max,
upper_bound=alpha_max,
)
alpha_2 = qctrl.operations.pwc_signal(
values=alpha_2_values,
duration=duration,
)
# Create filtered signal
alpha_2_filtered = qctrl.operations.convolve_pwc(
alpha_2,
qctrl.operations.sinc_integral_function(sinc_cutoff_frequency),
)
# Discretize to obtain smoothed alpha_2(t) signal
alpha_2 = qctrl.operations.discretize_stf(
stf=alpha_2_filtered,
duration=duration,
segments_count=segment_count,
name="alpha_2",
)
# Create drive term
drive = qctrl.operations.pwc_operator_hermitian_part(
qctrl.operations.pwc_operator(signal=alpha_1, operator=sigma_x / 2)
)
# Create clock shift term
shift = qctrl.operations.pwc_operator(signal=alpha_2, operator=sigma_z / 2)
# Create dephasing noise term
dephasing = qctrl.operations.constant_pwc_operator(
duration=duration, operator=sigma_z / duration
)
# Create target
target_operator = qctrl.operations.target(operator=target_operator)
#Construct Hamiltonian.
hamiltonian = qctrl.operations.pwc_sum(
[
drive,
shift,
dephasing,
]
)
# Solve Schrodinger's equation and get total unitary at the end
unitary = qctrl.operations.time_evolution_operators_pwc(
hamiltonian=hamiltonian,
sample_times=np.array([duration]),
)[-1]
unitary.name = "unitary"
# Repeat final unitary
repeated_unitary = np.eye(2)
for _ in range(repetitions):
repeated_unitary = repeated_unitary @ unitary
repeated_unitary.name = "repeated_unitary"
# Initial state
initial_state = np.array([[1], [0]])
# Calculate final state.
state = repeated_unitary @ initial_state
# Calculate final populations.
populations = qctrl.operations.abs(state[:, 0]) ** 2
# Normalize populations
norm = qctrl.operations.sum(populations)
populations = populations / norm
populations.name = "populations"
# Create infidelity
infidelity = qctrl.operations.infidelity_pwc(
hamiltonian=hamiltonian,
target_operator=target_operator,
noise_operators=[dephasing],
name="infidelity",
)
# Run the optimization
optimization_result = qctrl.functions.calculate_optimization(
cost_node_name="infidelity",
output_node_names=["alpha_1","alpha_2","unitary", "repeated_unitary", "populations"],
graph=graph,
)
print("Optimized cost:\t", optimization_result.cost)
# Plot the optimized controls
plot_controls(
plt.figure(),
controls={
"$\\alpha_1$": optimization_result.output["alpha_1"],
"$\\alpha_2$": optimization_result.output["alpha_2"],
},
)
plt.show()
#Extract Outputs
unitary = optimization_result.output["unitary"]["value"]
populations = optimization_result.output["populations"]["value"]
# Sample projective measurements.
true_measurements = np.random.choice(2, size=shots, p=populations)
measurements = np.array(
[np.random.choice(2, p=confusion_matrix[m]) for m in true_measurements]
)
results = {"unitary": unitary, "measurements": measurements}
return results
error_norm = (
lambda operate_a, operator_b: 1
- np.abs(np.trace((operate_a.conj().T @ operator_b)) / 2) ** 2
)
def estimate_probability_of_one(measurements):
size = len(measurements)
probability = np.mean(measurements)
standard_error = np.std(measurements) / np.sqrt(size)
return (probability, standard_error)
not_op_matrix = get_optimized_matrix(ideal_not_gate,repetitions=100)
h_op_matrix = get_optimized_matrix(ideal_h_gate,repetitions=100)
realised_not_gate = not_op_matrix["unitary"]
ideal_not_gate = np.array([[0, -1j], [-1j, 0]])
not_error = error_norm(realised_not_gate, ideal_not_gate)
realised_h_gate = h_op_matrix["unitary"]
ideal_h_gate = (1 / np.sqrt(2)) * np.array([[1, -1j], [-1j, 1]])
h_error = error_norm(realised_h_gate, ideal_h_gate)
not_measurements = not_op_matrix["measurements"]
h_measurements = h_op_matrix["measurements"]
not_probability, not_standard_error = estimate_probability_of_one(not_measurements)
h_probability, h_standard_error = estimate_probability_of_one(h_measurements)
print("Realised NOT Gate:")
print(realised_not_gate)
print("Ideal NOT Gate:")
print(ideal_not_gate)
print("NOT Gate Error:" + str(not_error))
print("NOT estimated probability of getting 1:" + str(not_probability))
print("NOT estimate standard error:" + str(not_standard_error) + "\n")
print("Realised H Gate:")
print(realised_h_gate)
print("Ideal H Gate:")
print(ideal_h_gate)
print("H Gate Error:" + str(h_error))
print("H estimated probability of getting 1:" + str(h_probability))
print("H estimate standard error:" + str(h_standard_error))
```
| github_jupyter |
# Practical Deep Learning for Coders, v3
# Lesson7_resnet_mnist
# MNIST CNN
```
%reload_ext autoreload
%autoreload 2
%matplotlib inline
from fastai.vision import *
```
### Data 数据
```
path = untar_data(URLs.MNIST)
path.ls()
il = ImageList.from_folder(path, convert_mode='L')
il.items[0]
defaults.cmap='binary'
il
il[0].show()
sd = il.split_by_folder(train='training', valid='testing')
sd
(path/'training').ls()
ll = sd.label_from_folder()
ll
x,y = ll.train[0]
x.show()
print(y,x.shape)
tfms = ([*rand_pad(padding=3, size=28, mode='zeros')], [])
ll = ll.transform(tfms)
bs = 128
# not using imagenet_stats because not using pretrained model
data = ll.databunch(bs=bs).normalize()
x,y = data.train_ds[0]
x.show()
print(y)
def _plot(i,j,ax): data.train_ds[0][0].show(ax, cmap='gray')
plot_multi(_plot, 3, 3, figsize=(8,8))
xb,yb = data.one_batch()
xb.shape,yb.shape
data.show_batch(rows=3, figsize=(5,5))
```
### Basic CNN with batchnorm 批量正规化(归一化)的基础版CNN模型
```
def conv(ni,nf): return nn.Conv2d(ni, nf, kernel_size=3, stride=2, padding=1)
model = nn.Sequential(
conv(1, 8), # 14
nn.BatchNorm2d(8),
nn.ReLU(),
conv(8, 16), # 7
nn.BatchNorm2d(16),
nn.ReLU(),
conv(16, 32), # 4
nn.BatchNorm2d(32),
nn.ReLU(),
conv(32, 16), # 2
nn.BatchNorm2d(16),
nn.ReLU(),
conv(16, 10), # 1
nn.BatchNorm2d(10),
Flatten() # remove (1,1) grid
)
learn = Learner(data, model, loss_func = nn.CrossEntropyLoss(), metrics=accuracy)
print(learn.summary())
xb = xb.cuda()
model(xb).shape
learn.lr_find(end_lr=100)
learn.recorder.plot()
learn.fit_one_cycle(3, max_lr=0.1)
```
### Refactor 重构
```
def conv2(ni,nf): return conv_layer(ni,nf,stride=2)
model = nn.Sequential(
conv2(1, 8), # 14
conv2(8, 16), # 7
conv2(16, 32), # 4
conv2(32, 16), # 2
conv2(16, 10), # 1
Flatten() # remove (1,1) grid
)
learn = Learner(data, model, loss_func = nn.CrossEntropyLoss(), metrics=accuracy)
learn.fit_one_cycle(10, max_lr=0.1)
```
### Resnet-ish
```
class ResBlock(nn.Module):
def __init__(self, nf):
super().__init__()
self.conv1 = conv_layer(nf,nf)
self.conv2 = conv_layer(nf,nf)
def forward(self, x): return x + self.conv2(self.conv1(x))
help(res_block)
model = nn.Sequential(
conv2(1, 8),
res_block(8),
conv2(8, 16),
res_block(16),
conv2(16, 32),
res_block(32),
conv2(32, 16),
res_block(16),
conv2(16, 10),
Flatten()
)
def conv_and_res(ni,nf): return nn.Sequential(conv2(ni, nf), res_block(nf))
model = nn.Sequential(
conv_and_res(1, 8),
conv_and_res(8, 16),
conv_and_res(16, 32),
conv_and_res(32, 16),
conv2(16, 10),
Flatten()
)
learn = Learner(data, model, loss_func = nn.CrossEntropyLoss(), metrics=accuracy)
learn.lr_find(end_lr=100)
learn.recorder.plot()
learn.fit_one_cycle(12, max_lr=0.05)
print(learn.summary())
```
| github_jupyter |
```
#cell-width control
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:80% !important; }</style>"))
%%javascript
IPython.OutputArea.prototype._should_scroll = function(lines) {
return false;
}
#packages
import numpy
import tensorflow as tf
from tensorflow.core.example import example_pb2
#utils
import os
import random
import pickle
import struct
import time
from noise import *
#keras
import keras
from keras.preprocessing import text, sequence
from keras.preprocessing.text import Tokenizer
from keras.models import Model
from keras.models import load_model
from keras.layers import Dense, Dropout, Activation, Concatenate, Dot, Embedding, LSTM, Conv1D, MaxPooling1D, Input, Lambda
numpy.random.seed(47)
####TRAIN####
filename = "/home/oala/Documents/MT/data/datasets/finished_files/test.bin"
#noise_candidates_path = '/home/oala/Documents/MT/noising/4-beam-PGC-noise-on-train/pretrained_model_tf1.2.1/decode_train_400maxenc_4beam_35mindec_120maxdec_ckpt-238410/decoded/'
#filename = "/home/donald/documents/MT/data/data-essentials-mini/finished_files/val.bin"
nc_dist = (0.5,0.5)
corr_sample = False
separate = False
fractions = {"switch-pairs":0.25,"sentence-switch-entire-bank":0.25,\
"sentence-switch-same-text-bank":0.25,"word-switch-entire-bank":0.25}
#read in clean data
print('Reading clean data...')
text_summ_pairs = []
with open(filename, 'r') as data:
text = data.readline()
summ = data.readline()
while summ:
if len(text) > 2 and len(summ) > 2:
text_summ_pairs.append([text[0:-1], summ[0:-1]])
text = data.readline()
summ = data.readline()
clean_2d = numpy.array(text_summ_pairs, dtype=object)
print('...done!')
#preprocess clean data, i.e. remove <s> and </s>
print('Preprocess clean data, i.e. remove <s> and </s>...')
for i in range(clean_2d.shape[0]):
clean_2d[i,1] = clean_2d[i,1].replace('<s> ', '')
clean_2d[i,1] = clean_2d[i,1].replace(' </s>', '')
print('...done!')
partition = {}
partition['test'] = []
labels = {}
id_counter = 1
#get targets
N_clean = clean_2d.shape[0]
#read in candidate noise points
orig_data_2d = clean_2d
###get all sentences and all words###
N = orig_data_2d.shape[0]
sentences = []
summ_sentences = []
text_sentences = []
text_words = []
summ_words = []
all_words = []
removes = []
sent_detector = nltk.data.load('tokenizers/punkt/english.pickle')
for i in range(N):
#sentences += sent_detector.tokenize(orig_data_2d[i,0].strip())
#sentences += sent_detector.tokenize(orig_data_2d[i,1].strip())
text_list = sent_detector.tokenize(orig_data_2d[i,0].strip())
summ_list = sent_detector.tokenize(orig_data_2d[i,1].strip())
text_words.append(orig_data_2d[i,0].split(" "))
summ_words.append(orig_data_2d[i,1].split(" "))
removes.append(summ_list[-1])
summ_list = summ_list[0:-1]
new_summ_list = []
if len(summ_list) > 0:
for j in range(len(summ_list)):
if j == 0:
new_summ_list.append(summ_list[j]+" </s>")
else:
new_summ_list.append(summ_list[j][5:]+" </s>")
else:
new_summ_list.append(orig_data_2d[i,1])
new_text_list = []
for sentence in text_list:
new_text_list.append("<s> "+sentence+" </s> ")
sentences += new_text_list + new_summ_list
summ_sentences.append(new_summ_list)
text_sentences.append(new_text_list)
all_words = summ_words + text_words
all_words = set([item for sublist in all_words for item in sublist])
all_words.discard('<s>')
all_words.discard('</s>')
all_words = list(all_words)
N_clean = orig_data_2d.shape[0]
clean_ratio, noise_ratio = nc_dist
N_noise = int((N_clean - N_clean*clean_ratio)/clean_ratio)
###noise: switch pairs###
rel_fact = fractions["switch-pairs"]
N = orig_data_2d.shape[0]
N_noise1 = int(N_noise*rel_fact)
#get regular indices and randomized indices
if type(rel_fact) == int:
indices_in_order = numpy.array(list(range(N))*rel_fact)
if type(rel_fact) == float:
start = int(rel_fact//1)
indices_in_order = list(range(N))*start
on_top = N_noise1 - len(indices_in_order)
#relative_index = int((N*(rel_fact%1))/100)
indices_in_order += list(range(N))[0:on_top]
random_indices = numpy.random.choice(list(range(N)), size=N_noise1, replace=True)
#quick fix noise indices that would create signal data
#TODO: make this noise fixing truly random
#print(len(indices_in_order))
#print(random_indices.shape)
#print(orig_data_2d.shape)
#print(noise_data1_2d.shape)
random_indices[indices_in_order == random_indices] += 1
random_indices[random_indices > N] -= 2
#print noise 1 to file
for i in range(N_noise1):
store_string = orig_data_2d[indices_in_order[i],0] + '\n'+orig_data_2d[random_indices[i],1].replace('<s> ', '').replace(' </s>', '')
id_name = 'id-'+str(id_counter)
with open('/media/oala/4TB/experimental-data/evaluation-data/test-onlynoise/pseudorandom-dist/only-noise/'+id_name, 'w') as file:
file.write(store_string)
partition['test'] += [id_name]
labels[id_name] = 0
id_counter += 1
###noise: sentence switch (entire bank)###
rel_fact = fractions["sentence-switch-entire-bank"]
N = orig_data_2d.shape[0]
N_noise2 = int(N_noise*rel_fact)
#initialize noise 2d array
#noise_data2_2d = numpy.ndarray((N_noise2,2), dtype=object)
#get total number of sentences
num_sentences = len(sentences)
#track avg number of changes across all changes
avg_num_changes = 0
for i in range(N_noise2): #iterate through the number of noisy points we want to create
textID = numpy.random.randint(0,N)
#summID = numpy.random.randint(0,N)
summID = textID
len_summ = len(summ_sentences[summID])
if len_summ == 0:
print('summID: ', summID)
print('textID: ', textID)
print(orig_data_2d[summID,1])
print(summ_sentences[summID])
#pick number of changes
num_changes = numpy.random.randint(1,len_summ+1)
avg_num_changes += num_changes/N_noise2
#pick positions for changes
pos_changes = numpy.random.randint(0,len_summ, size=num_changes)
old_summ = summ_sentences[summID][:]
#new_summ = ""
for j in pos_changes:
old_summ[j] = sentences[numpy.random.randint(0,num_sentences)]
new_summ = " ".join(old_summ)
if textID == summID:
if old_summ == new_summ:
print('ouch')
store_string = orig_data_2d[textID,0] + '\n'+ new_summ.replace('<s> ', '').replace(' </s>', '')
id_name = 'id-'+str(id_counter)
with open('/media/oala/4TB/experimental-data/evaluation-data/test-onlynoise/pseudorandom-dist/only-noise/'+id_name, 'w') as file:
file.write(store_string)
partition['test'] += [id_name]
labels[id_name] = 0
id_counter += 1
###noise: sentence switch (same text bank)###
rel_fact = fractions["sentence-switch-same-text-bank"]
N = orig_data_2d.shape[0]
N_noise3 = int(N_noise*rel_fact)
#initialize noise 2d array
#noise_data3_2d = numpy.ndarray((N_noise3,2), dtype=object)
#track avg number of changes across all changes
avg_num_changes = 0
for i in range(N_noise3): #iterate through the number of noisy points we want to create
textID = numpy.random.randint(0,N)
#summID = numpy.random.randint(0,N)
summID = textID
len_summ = len(summ_sentences[summID])
#get total number of sentences in the corresponding text
num_sentences = len(text_sentences[textID])
if len_summ == 0:
print('summID: ', summID)
print('textID: ', textID)
print(orig_data_2d[summID,1])
print(summ_sentences[summID])
#pick number of changes
num_changes = numpy.random.randint(1,len_summ+1)
avg_num_changes += num_changes/N_noise3
#pick positions for changes
pos_changes = numpy.random.randint(0,len_summ, size=num_changes)
old_summ = summ_sentences[summID][:]
#new_summ = ""
for j in pos_changes:
old_summ[j] = text_sentences[textID][numpy.random.randint(0,num_sentences)]
new_summ = " ".join(old_summ)
if textID == summID:
if old_summ == new_summ:
print('ouch')
store_string = orig_data_2d[textID,0] + '\n'+ new_summ.replace('<s> ', '').replace(' </s>', '')
id_name = 'id-'+str(id_counter)
with open('/media/oala/4TB/experimental-data/evaluation-data/test-onlynoise/pseudorandom-dist/only-noise/'+id_name, 'w') as file:
file.write(store_string)
partition['test'] += [id_name]
labels[id_name] = 0
id_counter += 1
###noise: word switch (entire bank)###
rel_fact = fractions["word-switch-entire-bank"]
N = orig_data_2d.shape[0]
N_noise4 = int(N_noise*rel_fact)
#initialize noise 2d array
#noise_data4_2d = numpy.ndarray((N_noise4,2), dtype=object)
#get total number of sentences
num_sentences = len(sentences)
num_words = len(all_words)
#track avg number of changes across all changes
avg_num_changes = 0
for i in range(N_noise4): #iterate through the number of noisy points we want to create
textID = numpy.random.randint(0,N)
#summID = numpy.random.randint(0,N)
summID = textID
len_summ = len(summ_words[summID])
if len_summ == 0:
print('summID: ', summID)
print('textID: ', textID)
print(orig_data_2d[summID,1])
print(summ_sentences[summID])
#pick number of changes
num_changes = numpy.random.randint(1,len_summ+1)
avg_num_changes += num_changes/N_noise4
#pick positions for changes
pos_changes = numpy.random.randint(0,len_summ, size=num_changes)
old_summ = summ_words[summID][:]
#new_summ = ""
for j in pos_changes:
if old_summ[j] == '<s>' or old_summ[j] == '</s>':
pass
else:
old_summ[j] = all_words[numpy.random.randint(0,num_words)]
new_summ = " ".join(old_summ)
if textID == summID:
if old_summ == new_summ:
print('ouch')
store_string = orig_data_2d[textID,0] + '\n'+ new_summ.replace('<s> ', '').replace(' </s>', '')
id_name = 'id-'+str(id_counter)
with open('/media/oala/4TB/experimental-data/evaluation-data/test-onlynoise/pseudorandom-dist/only-noise/'+id_name, 'w') as file:
file.write(store_string)
partition['test'] += [id_name]
labels[id_name] = 0
id_counter += 1
print(len(partition['test']))
count = 0
for key in partition['test']:
count += labels[key]
print(count/len(partition['test']))
#store label dict
print('store label dict...')
with open('/media/oala/4TB/experimental-data/evaluation-data/test-onlynoise/pseudorandom-dist/only-noise/'+'labels.pickle', 'wb') as handle:
pickle.dump(labels, handle, protocol=pickle.HIGHEST_PROTOCOL)
print('...done!')
#store partition dict
print('partition dict...')
with open('/media/oala/4TB/experimental-data/evaluation-data/test-onlynoise/pseudorandom-dist/only-noise/'+'partition.pickle', 'wb') as handle:
pickle.dump(partition, handle, protocol=pickle.HIGHEST_PROTOCOL)
print('...done!')
```
| github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#Goal" data-toc-modified-id="Goal-1"><span class="toc-item-num">1 </span>Goal</a></span></li><li><span><a href="#Var" data-toc-modified-id="Var-2"><span class="toc-item-num">2 </span>Var</a></span></li><li><span><a href="#Init" data-toc-modified-id="Init-3"><span class="toc-item-num">3 </span>Init</a></span></li><li><span><a href="#Checking-overlap-with-train" data-toc-modified-id="Checking-overlap-with-train-4"><span class="toc-item-num">4 </span>Checking overlap with train</a></span></li><li><span><a href="#DM-run" data-toc-modified-id="DM-run-5"><span class="toc-item-num">5 </span>DM run</a></span></li><li><span><a href="#Communites" data-toc-modified-id="Communites-6"><span class="toc-item-num">6 </span>Communites</a></span><ul class="toc-item"><li><span><a href="#n100_r5" data-toc-modified-id="n100_r5-6.1"><span class="toc-item-num">6.1 </span>n100_r5</a></span></li><li><span><a href="#n100_r25" data-toc-modified-id="n100_r25-6.2"><span class="toc-item-num">6.2 </span>n100_r25</a></span></li></ul></li><li><span><a href="#Estimated-coverage" data-toc-modified-id="Estimated-coverage-7"><span class="toc-item-num">7 </span>Estimated coverage</a></span></li><li><span><a href="#Feature-tables" data-toc-modified-id="Feature-tables-8"><span class="toc-item-num">8 </span>Feature tables</a></span><ul class="toc-item"><li><span><a href="#n100_r5" data-toc-modified-id="n100_r5-8.1"><span class="toc-item-num">8.1 </span>n100_r5</a></span></li><li><span><a href="#n100_r25" data-toc-modified-id="n100_r25-8.2"><span class="toc-item-num">8.2 </span>n100_r25</a></span><ul class="toc-item"><li><span><a href="#Number-of-contigs" data-toc-modified-id="Number-of-contigs-8.2.1"><span class="toc-item-num">8.2.1 </span>Number of contigs</a></span></li><li><span><a href="#Misassembly-types" data-toc-modified-id="Misassembly-types-8.2.2"><span class="toc-item-num">8.2.2 </span>Misassembly types</a></span></li></ul></li></ul></li><li><span><a href="#sessionInfo" data-toc-modified-id="sessionInfo-9"><span class="toc-item-num">9 </span>sessionInfo</a></span></li></ul></div>
# Goal
* evaluate the results of the test dataset generation
# Var
```
# location of all training dataset runs
work_dir = '/ebio/abt3_projects/databases_no-backup/DeepMAsED/test_runs/'
# 100 ref genomes, 3 metagenomes
n100_r5_dir = file.path(work_dir, 'n100_r5')
# 100 ref genomes, 25 metagenomes (due to error with one, used 5 more)
n100_r25_dir = file.path(work_dir, 'n100_r25')
# input genome table
test_n100_ref_file = '/ebio/abt3_projects/databases_no-backup/DeepMAsED/GTDB_ref_genomes/DeepMAsED_GTDB_genome-refs_test_n100.tsv'
train_n1000_ref_file = '/ebio/abt3_projects/databases_no-backup/DeepMAsED/GTDB_ref_genomes/DeepMAsED_GTDB_genome-refs_train_n1000.tsv'
```
# Init
```
library(dplyr)
library(tidyr)
library(ggplot2)
library(data.table)
source('/ebio/abt3_projects/software/dev/DeepMAsED/bin/misc_r_functions/init.R')
```
# Checking overlap with train
* making sure train/test genomes don't overlap
# DM run
```
# pipeline run config files
cat_file(file.path(n100_r5_dir, 'config.yaml'))
# pipeline run config files
cat_file(file.path(n100_r25_dir, 'config.yaml'))
```
# Communites
* Simulated abundances of each ref genome
## n100_r5
```
comm_files = list.files(file.path(n100_r5_dir, 'MGSIM'), 'comm_wAbund.txt', full.names=TRUE, recursive=TRUE)
comm_files
comms = list()
for(F in comm_files){
df = read.delim(F, sep='\t')
df$Rep = basename(dirname(F))
comms[[F]] = df
}
comms = do.call(rbind, comms)
rownames(comms) = 1:nrow(comms)
comms %>% dfhead
comms$Perc_rel_abund %>% summary %>% print
p = comms %>%
group_by(Taxon) %>%
mutate(mean_Rank = mean(Rank)) %>%
ungroup() %>%
mutate(Taxon = reorder(Taxon, mean_Rank)) %>%
ggplot(aes(Taxon, Perc_rel_abund, color=Rep)) +
geom_point(size=0.5, alpha=0.4) +
scale_y_log10() +
labs(y='% abundance') +
theme_bw() +
theme(
axis.text.x = element_blank()
)
dims(10,3)
plot(p)
p = comms %>%
mutate(Perc_rel_abund = ifelse(Perc_rel_abund == 0, 1e-5, Perc_rel_abund)) %>%
group_by(Taxon) %>%
summarize(mean_perc_abund = mean(Perc_rel_abund),
sd_perc_abund = sd(Perc_rel_abund)) %>%
ungroup() %>%
mutate(neg_sd_perc_abund = mean_perc_abund - sd_perc_abund,
pos_sd_perc_abund = mean_perc_abund + sd_perc_abund,
neg_sd_perc_abund = ifelse(neg_sd_perc_abund <= 0, 1e-5, neg_sd_perc_abund)) %>%
mutate(Taxon = Taxon %>% reorder(-mean_perc_abund)) %>%
ggplot(aes(Taxon, mean_perc_abund)) +
geom_linerange(aes(ymin=neg_sd_perc_abund, ymax=pos_sd_perc_abund),
size=0.3, alpha=0.3) +
geom_point(size=0.5, alpha=0.4, color='red') +
labs(y='% abundance') +
theme_bw() +
theme(
axis.text.x = element_blank(),
panel.grid.major.x = element_blank(),
panel.grid.major.y = element_blank(),
panel.grid.minor.x = element_blank(),
panel.grid.minor.y = element_blank()
)
dims(10,2.5)
plot(p)
dims(10,2.5)
plot(p + scale_y_log10())
```
## n100_r25
```
comm_files = list.files(file.path(n100_r25_dir, 'MGSIM'), 'comm_wAbund.txt', full.names=TRUE, recursive=TRUE)
comm_files %>% length %>% print
comm_files %>% head
comms = list()
for(F in comm_files){
df = read.delim(F, sep='\t')
df$Rep = basename(dirname(F))
comms[[F]] = df
}
comms = do.call(rbind, comms)
rownames(comms) = 1:nrow(comms)
comms %>% dfhead
p = comms %>%
mutate(Perc_rel_abund = ifelse(Perc_rel_abund == 0, 1e-5, Perc_rel_abund)) %>%
group_by(Taxon) %>%
summarize(mean_perc_abund = mean(Perc_rel_abund),
sd_perc_abund = sd(Perc_rel_abund)) %>%
ungroup() %>%
mutate(neg_sd_perc_abund = mean_perc_abund - sd_perc_abund,
pos_sd_perc_abund = mean_perc_abund + sd_perc_abund,
neg_sd_perc_abund = ifelse(neg_sd_perc_abund <= 0, 1e-5, neg_sd_perc_abund)) %>%
mutate(Taxon = Taxon %>% reorder(-mean_perc_abund)) %>%
ggplot(aes(Taxon, mean_perc_abund)) +
geom_linerange(aes(ymin=neg_sd_perc_abund, ymax=pos_sd_perc_abund),
size=0.3, alpha=0.3) +
geom_point(size=0.5, alpha=0.4, color='red') +
labs(y='% abundance') +
theme_bw() +
theme(
axis.text.x = element_blank(),
panel.grid.major.x = element_blank(),
panel.grid.major.y = element_blank(),
panel.grid.minor.x = element_blank(),
panel.grid.minor.y = element_blank()
)
dims(10,2.5)
plot(p)
dims(10,2.5)
plot(p + scale_y_log10())
```
# Estimated coverage
* Coverage estimated with nonpareil
```
F = file.path(n100_r5_dir, 'coverage', 'nonpareil', 'all_summary.txt')
cov = read.delim(F, sep='\t')
cov %>% summary %>% print
F = file.path(n100_r25_dir, 'coverage', 'nonpareil', 'all_summary.txt')
cov = read.delim(F, sep='\t')
cov %>% summary %>% print
```
# Feature tables
* feature tables for ML model training/testing
## n100_r5
```
feat_files = list.files(file.path(n100_r5_dir, 'map'), 'features.tsv.gz', full.names=TRUE, recursive=TRUE)
feat_files %>% length %>% print
feat_files %>% head
feats = list()
for(F in feat_files){
cmd = glue::glue('gunzip -c {F}', F=F)
df = fread(cmd, sep='\t') %>%
distinct(contig, assembler, Extensive_misassembly)
df$Rep = basename(dirname(dirname(F)))
feats[[F]] = df
}
feats = do.call(rbind, feats)
rownames(feats) = 1:nrow(feats)
feats %>% dfhead
p = feats %>%
mutate(Extensive_misassembly = ifelse(Extensive_misassembly == '', 'None',
Extensive_misassembly)) %>%
group_by(Extensive_misassembly, assembler, Rep) %>%
summarize(n = n()) %>%
ungroup() %>%
ggplot(aes(Extensive_misassembly, n, color=assembler)) +
geom_boxplot() +
scale_y_log10() +
labs(x='metaQUAST extensive mis-assembly', y='Count') +
coord_flip() +
theme_bw() +
theme(
axis.text.x = element_text(angle=45, hjust=1)
)
dims(7,4)
plot(p)
```
## n100_r25
```
feat_files = list.files(file.path(n100_r25_dir, 'map'), 'features.tsv.gz', full.names=TRUE, recursive=TRUE)
feat_files %>% length %>% print
feat_files %>% head
feats = list()
for(F in feat_files){
cmd = glue::glue('gunzip -c {F}', F=F)
df = fread(cmd, sep='\t') %>%
distinct(contig, assembler, Extensive_misassembly)
df$Rep = basename(dirname(dirname(F)))
feats[[F]] = df
}
feats = do.call(rbind, feats)
rownames(feats) = 1:nrow(feats)
feats %>% dfhead
```
### Number of contigs
```
feats_s = feats %>%
group_by(assembler, Rep) %>%
summarize(n_contigs = n_distinct(contig)) %>%
ungroup
feats_s$n_contigs %>% summary
```
### Misassembly types
```
p = feats %>%
mutate(Extensive_misassembly = ifelse(Extensive_misassembly == '', 'None',
Extensive_misassembly)) %>%
group_by(Extensive_misassembly, assembler, Rep) %>%
summarize(n = n()) %>%
ungroup() %>%
ggplot(aes(Extensive_misassembly, n, color=assembler)) +
geom_boxplot() +
scale_y_log10() +
labs(x='metaQUAST extensive mis-assembly', y='Count') +
coord_flip() +
theme_bw() +
theme(
axis.text.x = element_text(angle=45, hjust=1)
)
dims(7,4)
plot(p)
```
# sessionInfo
```
sessionInfo()
```
| github_jupyter |
<i>Copyright (c) Microsoft Corporation. All rights reserved.</i>
<i>Licensed under the MIT License.</i>
# DKN : Deep Knowledge-Aware Network for News Recommendation
DKN \[1\] is a deep learning model which incorporates information from knowledge graph for better news recommendation. Specifically, DKN uses TransX \[2\] method for knowledge graph representaion learning, then applies a CNN framework, named KCNN, to combine entity embedding with word embedding and generate a final embedding vector for a news article. CTR prediction is made via an attention-based neural scorer.
## Properties of DKN:
- DKN is a content-based deep model for CTR prediction rather than traditional ID-based collaborative filtering.
- It makes use of knowledge entities and common sense in news content via joint learning from semantic-level and knnowledge-level representations of news articles.
- DKN uses an attention module to dynamically calculate a user's aggregated historical representaition.
## Data format:
One simple example: <br>
`0 CandidateNews:1050,5238,5165,6996,1922,7058,7263,0,0,0 entity:0,0,0,0,2562,2562,2562,0,0,0 clickedNews0:7680,3978,1451,409,4681,0,0,0,0,0 entity0:0,0,0,395,0,0,0,0,0,0 clickedNews1:3698,2301,1055,6481,6242,7203,0,0,0,0 entity1:0,0,1904,1904,1904,0,0,0,0,0 ...`
<br>
In general, each line in data file represents one instance, the format is like: <br>
`[label] [CandidateNews:w1,w2,w3,...] [entity:e1,e2,e3,...] [clickedNews0:w1,w2,w3,...] [entity0:e1,e2,e3,...] ...` <br>
It contains several parts seperated by space, i.e. label part, CandidateNews part, ClickedHistory part. CandidateNews part describes the target news article we are going to score in this instance, it is represented by (aligned) title words and title entities. To take a quick example, a news title may be : `Trump to deliver State of the Union address next week` , then the title words value may be `CandidateNews:34,45,334,23,12,987,3456,111,456,432` and the title entitie value may be: `entity:45,0,0,0,0,0,0,0,0,0`. Only the first value of entity vector is non-zero due to the word `Trump`. <br>
clickedNewsk and entityk describe the k-th news article the user ever clicked and the format is the same as candidate news. Words and entities are aligned in news title. We use a fixed length to describe an article, if the title is less than the fixed length, just pad it with zeros.
## Global settings and imports
```
import sys
sys.path.append("../../")
from reco_utils.recommender.deeprec.deeprec_utils import *
from reco_utils.recommender.deeprec.models.dkn import *
from reco_utils.recommender.deeprec.IO.dkn_iterator import *
import papermill as pm
```
## Download and load data
```
data_path = '../../tests/resources/deeprec/dkn'
yaml_file = os.path.join(data_path, r'dkn.yaml')
train_file = os.path.join(data_path, r'final_train_with_entity.txt')
valid_file = os.path.join(data_path, r'final_test_with_entity.txt')
wordEmb_file = os.path.join(data_path, r'word_embeddings_100.npy')
entityEmb_file = os.path.join(data_path, r'TransE_entity2vec_100.npy')
if not os.path.exists(yaml_file):
download_deeprec_resources(r'https://recodatasets.blob.core.windows.net/deeprec/', data_path, 'dknresources.zip')
```
## Create hyper-parameters
```
epoch=10
hparams = prepare_hparams(yaml_file, wordEmb_file=wordEmb_file, entityEmb_file=entityEmb_file, epochs=epoch)
print(hparams)
input_creator = DKNTextIterator
```
## Train the DKN model
```
model = DKN(hparams, input_creator)
print(model.run_eval(valid_file))
model.fit(train_file, valid_file)
```
Now we can test again the performance on valid set:
```
res = model.run_eval(valid_file)
print(res)
pm.record("res", res)
```
## Reference
\[1\] Wang, Hongwei, et al. "DKN: Deep Knowledge-Aware Network for News Recommendation." Proceedings of the 2018 World Wide Web Conference on World Wide Web. International World Wide Web Conferences Steering Committee, 2018.<br>
\[2\] Knowledge Graph Embeddings including TransE, TransH, TransR and PTransE. https://github.com/thunlp/KB2E
| github_jupyter |
<h1> Structured data prediction using Cloud ML Engine </h1>
This notebook illustrates:
<ol>
<li> Exploring a BigQuery dataset using Datalab
<li> Creating datasets for Machine Learning using Dataflow
<li> Creating a model using the high-level Estimator API
<li> Training on Cloud ML Engine
<li> Deploying model
<li> Predicting with model
</ol>
```
# change these to try this notebook out
BUCKET = 'cloud-training-demos-ml'
PROJECT = 'cloud-training-demos'
REGION = 'us-central1'
import os
os.environ['BUCKET'] = BUCKET
os.environ['PROJECT'] = PROJECT
os.environ['REGION'] = REGION
%%bash
if ! gsutil ls | grep -q gs://${BUCKET}/; then
gsutil mb -l ${REGION} gs://${BUCKET}
fi
```
<h2> Explore data </h2>
The data is natality data (record of births in the US). My goal is to predict the baby's weight given a number of factors about the pregnancy and the baby's mother. Later, we will want to split the data into training and eval datasets. The hash of the year-month will be used for that.
```
query="""
SELECT
weight_pounds,
is_male,
mother_age,
mother_race,
plurality,
gestation_weeks,
mother_married,
ever_born,
cigarette_use,
alcohol_use,
FARM_FINGERPRINT(CONCAT(CAST(YEAR AS STRING), CAST(month AS STRING))) AS hashmonth
FROM
publicdata.samples.natality
WHERE year > 2000
"""
import google.datalab.bigquery as bq
df = bq.Query(query + " LIMIT 100").execute().result().to_dataframe()
df.head()
```
Let's write a query to find the unique values for each of the columns and the count of those values.
```
def get_distinct_values(column_name):
sql = """
SELECT
{0},
COUNT(1) AS num_babies,
AVG(weight_pounds) AS avg_wt
FROM
publicdata.samples.natality
WHERE
year > 2000
GROUP BY
{0}
""".format(column_name)
return bq.Query(sql).execute().result().to_dataframe()
df = get_distinct_values('mother_race')
df.plot(x='mother_race', logy='num_babies', kind='bar');
df.plot(x='mother_race', y='avg_wt', kind='bar');
df = get_distinct_values('is_male')
df.plot(x='is_male', logy='num_babies', kind='bar');
df.plot(x='is_male', y='avg_wt', kind='bar');
df = get_distinct_values('mother_age')
df = df.sort_values('mother_age')
df.plot(x='mother_age', y='num_babies');
df.plot(x='mother_age', y='avg_wt');
df = get_distinct_values('plurality')
df = df.sort_values('plurality')
df.plot(x='plurality', logy='num_babies', kind='bar');
df.plot(x='plurality', y='avg_wt', kind='bar');
df = get_distinct_values('gestation_weeks')
df = df.sort_values('gestation_weeks')
df.plot(x='gestation_weeks', logy='num_babies', kind='bar');
df.plot(x='gestation_weeks', y='avg_wt', kind='bar');
df = get_distinct_values('mother_married')
df.plot(x='mother_married', logy='num_babies', kind='bar');
df.plot(x='mother_married', y='avg_wt', kind='bar');
df = get_distinct_values('ever_born')
df = df.sort_values('ever_born')
df.plot(x='ever_born', logy='num_babies', kind='bar');
df.plot(x='ever_born', y='avg_wt', kind='bar');
df = get_distinct_values('cigarette_use')
df.plot(x='cigarette_use', logy='num_babies', kind='bar');
df.plot(x='cigarette_use', y='avg_wt', kind='bar');
df = get_distinct_values('alcohol_use')
df.plot(x='alcohol_use', logy='num_babies', kind='bar');
df.plot(x='alcohol_use', y='avg_wt', kind='bar');
```
Other than the ever_born (the total number of babies born to this mother), the factors all seem to play a part in the baby's weight. Male babies are heavier on average than female babies. The mother's age and race play a part (age much more than race -- teenage and middle-aged moms tend to have lower-weight babies). Twins, triplets, etc. are lower weight than single births. Preemies weigh in lower as do babies born to single moms. Moms who use alcohol or cigarettes have babies that weigh lower on average.
<p>
In the rest of this notebook, we will use machine learning to combine all of these factors to come up with a prediction of a baby's weight.
<h2> Create ML dataset using Dataflow </h2>
<p>
Let's use Cloud Dataflow to read in the BigQuery data and write it out as CSV files.
Instead of using Beam/Dataflow, I had two other options:
<ol>
<li> Read from BigQuery directly using TensorFlow. However, using CSV files gives us the advantage of shuffling during read. This is important for distributed training because some workers might be slower than others, and shuffling the data helps prevent the same data from being assigned to the slow workers.
<li> Use the BigQuery console (http://bigquery.cloud.google.com) to run a Query and save the result as a CSV file. For larger datasets, you may have to select the option to "allow large results" and save the result into a CSV file on Google Cloud Storage. However, in this case, I want to do some preprocessing (on the "race" column). If I didn't need preprocessing, I could have used the web console. Also, I prefer to script it out rather than run queries on the user interface, so I am using Cloud Dataflow for the preprocessing.
</ol>
<p>
Note that after you launch this, the notebook will appear to be hung. Go to the GCP webconsole to the Dataflow section and monitor the running job. It took about <b>30 minutes</b> for me.
```
import apache_beam as beam
import datetime
def to_csv(rowdict):
# pull columns from BQ and create a line
import hashlib
import copy
CSV_COLUMNS = 'weight_pounds,is_male,mother_age,mother_race,plurality,gestation_weeks,mother_married,cigarette_use,alcohol_use'.split(',')
# modify opaque numeric race code into human-readable data
races = dict(zip([1,2,3,4,5,6,7,18,28,39,48],
['White', 'Black', 'American Indian', 'Chinese',
'Japanese', 'Hawaiian', 'Filipino',
'Asian Indian', 'Korean', 'Samaon', 'Vietnamese']))
result = copy.deepcopy(rowdict)
if 'mother_race' in rowdict and rowdict['mother_race'] in races:
result['mother_race'] = races[rowdict['mother_race']]
else:
result['mother_race'] = 'Unknown'
data = ','.join([str(result[k]) if k in result else 'None' for k in CSV_COLUMNS])
key = hashlib.sha224(data).hexdigest() # hash the columns to form a key
return str('{},{}'.format(data, key))
def preprocess(in_test_mode):
job_name = 'preprocess-babyweight-features' + '-' + datetime.datetime.now().strftime('%y%m%d-%H%M%S')
if in_test_mode:
print 'Launching local job ... hang on'
OUTPUT_DIR = './preproc'
else:
print 'Launching Dataflow job {} ... hang on'.format(job_name)
OUTPUT_DIR = 'gs://{0}/babyweight/preproc/'.format(BUCKET)
options = {
'staging_location': os.path.join(OUTPUT_DIR, 'tmp', 'staging'),
'temp_location': os.path.join(OUTPUT_DIR, 'tmp'),
'job_name': job_name,
'project': PROJECT,
'teardown_policy': 'TEARDOWN_ALWAYS',
'no_save_main_session': True
}
opts = beam.pipeline.PipelineOptions(flags=[], **options)
if in_test_mode:
RUNNER = 'DirectRunner'
else:
RUNNER = 'DataflowRunner'
p = beam.Pipeline(RUNNER, options=opts)
query = """
SELECT
weight_pounds,
is_male,
mother_age,
mother_race,
plurality,
gestation_weeks,
mother_married,
ever_born,
cigarette_use,
alcohol_use,
FARM_FINGERPRINT(CONCAT(CAST(YEAR AS STRING), CAST(month AS STRING))) AS hashmonth
FROM
publicdata.samples.natality
WHERE year > 2000
AND weight_pounds > 0
AND mother_age > 0
AND plurality > 0
AND gestation_weeks > 0
AND month > 0
"""
if in_test_mode:
query = query + ' LIMIT 100'
for step in ['train', 'eval']:
if step == 'train':
selquery = 'SELECT * FROM ({}) WHERE MOD(ABS(hashmonth),4) < 3'.format(query)
else:
selquery = 'SELECT * FROM ({}) WHERE MOD(ABS(hashmonth),4) = 3'.format(query)
(p
| '{}_read'.format(step) >> beam.io.Read(beam.io.BigQuerySource(query=selquery, use_standard_sql=True))
| '{}_csv'.format(step) >> beam.Map(to_csv)
| '{}_out'.format(step) >> beam.io.Write(beam.io.WriteToText(os.path.join(OUTPUT_DIR, '{}.csv'.format(step))))
)
job = p.run()
preprocess(in_test_mode=False)
```
<img src="dataflow.png" width="500"/>
```
%bash
gsutil ls gs://${BUCKET}/babyweight/preproc/*-00000*
```
<h2> Create TensorFlow model using TensorFlow's Estimator API </h2>
<p>
First, write an input_fn to read the data.
```
import shutil
import tensorflow as tf
import tensorflow.contrib.learn as tflearn
import tensorflow.contrib.layers as tflayers
from tensorflow.contrib.learn.python.learn import learn_runner
import tensorflow.contrib.metrics as metrics
CSV_COLUMNS = 'weight_pounds,is_male,mother_age,mother_race,plurality,gestation_weeks,mother_married,cigarette_use,alcohol_use,key'.split(',')
LABEL_COLUMN = 'weight_pounds'
KEY_COLUMN = 'key'
DEFAULTS = [[0.0], ['null'], [0.0], ['null'], [0.0], [0.0], ['null'], ['null'], ['null'], ['nokey']]
TRAIN_STEPS = 1000
def read_dataset(prefix, pattern, batch_size=512):
# use prefix to create filename
filename = 'gs://{}/babyweight/preproc/{}*{}*'.format(BUCKET, prefix, pattern)
if prefix == 'train':
mode = tf.contrib.learn.ModeKeys.TRAIN
else:
mode = tf.contrib.learn.ModeKeys.EVAL
# the actual input function passed to TensorFlow
def _input_fn():
# could be a path to one file or a file pattern.
input_file_names = tf.train.match_filenames_once(filename)
filename_queue = tf.train.string_input_producer(
input_file_names, shuffle=True)
# read CSV
reader = tf.TextLineReader()
_, value = reader.read_up_to(filename_queue, num_records=batch_size)
value_column = tf.expand_dims(value, -1)
columns = tf.decode_csv(value_column, record_defaults=DEFAULTS)
features = dict(zip(CSV_COLUMNS, columns))
features.pop(KEY_COLUMN)
label = features.pop(LABEL_COLUMN)
return features, label
return _input_fn
```
Next, define the feature columns
```
def get_wide_deep():
# define column types
races = ['White', 'Black', 'American Indian', 'Chinese',
'Japanese', 'Hawaiian', 'Filipino', 'Unknown',
'Asian Indian', 'Korean', 'Samaon', 'Vietnamese']
is_male,mother_age,mother_race,plurality,gestation_weeks,mother_married,cigarette_use,alcohol_use = \
[ \
tflayers.sparse_column_with_keys('is_male', keys=['True', 'False']),
tflayers.real_valued_column('mother_age'),
tflayers.sparse_column_with_keys('mother_race', keys=races),
tflayers.real_valued_column('plurality'),
tflayers.real_valued_column('gestation_weeks'),
tflayers.sparse_column_with_keys('mother_married', keys=['True', 'False']),
tflayers.sparse_column_with_keys('cigarette_use', keys=['True', 'False', 'None']),
tflayers.sparse_column_with_keys('alcohol_use', keys=['True', 'False', 'None'])
]
# which columns are wide (sparse, linear relationship to output) and which are deep (complex relationship to output?)
wide = [is_male, mother_race, plurality, mother_married, cigarette_use, alcohol_use]
deep = [\
mother_age,
gestation_weeks,
tflayers.embedding_column(mother_race, 3)
]
return wide, deep
```
To predict with the TensorFlow model, we also need a serving input function. We will want all the inputs from our user.
```
def serving_input_fn():
feature_placeholders = {
'is_male': tf.placeholder(tf.string, [None]),
'mother_age': tf.placeholder(tf.float32, [None]),
'mother_race': tf.placeholder(tf.string, [None]),
'plurality': tf.placeholder(tf.float32, [None]),
'gestation_weeks': tf.placeholder(tf.float32, [None]),
'mother_married': tf.placeholder(tf.string, [None]),
'cigarette_use': tf.placeholder(tf.string, [None]),
'alcohol_use': tf.placeholder(tf.string, [None])
}
features = {
key: tf.expand_dims(tensor, -1)
for key, tensor in feature_placeholders.items()
}
return tflearn.utils.input_fn_utils.InputFnOps(
features,
None,
feature_placeholders)
```
Finally, train!
```
from tensorflow.contrib.learn.python.learn.utils import saved_model_export_utils
pattern = "00001-of-" # process only one of the shards, for testing purposes
def experiment_fn(output_dir):
wide, deep = get_wide_deep()
return tflearn.Experiment(
tflearn.DNNLinearCombinedRegressor(model_dir=output_dir,
linear_feature_columns=wide,
dnn_feature_columns=deep,
dnn_hidden_units=[64, 32]),
train_input_fn=read_dataset('train', pattern),
eval_input_fn=read_dataset('eval', pattern),
eval_metrics={
'rmse': tflearn.MetricSpec(
metric_fn=metrics.streaming_root_mean_squared_error
)
},
export_strategies=[saved_model_export_utils.make_export_strategy(
serving_input_fn,
default_output_alternative_key=None,
exports_to_keep=1
)],
train_steps=TRAIN_STEPS
)
shutil.rmtree('babyweight_trained', ignore_errors=True) # start fresh each time
learn_runner.run(experiment_fn, 'babyweight_trained')
```
Now that we have the TensorFlow code working on a subset of the data (in the code above, I was reading only the 00001-of-x file), we can package the TensorFlow code up as a Python module and train it on Cloud ML Engine.
<p>
<h2> Train on Cloud ML Engine </h2>
<p>
Training on Cloud ML Engine requires:
<ol>
<li> Making the code a Python package
<li> Using gcloud to submit the training code to Cloud ML Engine
</ol>
<p>
The code in model.py is the same as in the above cells. I just moved it to a file so that I could package it up as a module.
(explore the <a href="babyweight/trainer">directory structure</a>).
```
%bash
grep "^def" babyweight/trainer/model.py
```
After moving the code to a package, make sure it works standalone. (Note the --pattern and --train_steps lines so that I am not trying to boil the ocean on my laptop). Even then, this takes about <b>a minute</b> in which you won't see any output ...
```
%bash
echo "bucket=${BUCKET}"
rm -rf babyweight_trained
export PYTHONPATH=${PYTHONPATH}:${PWD}/babyweight
python -m trainer.task \
--bucket=${BUCKET} \
--output_dir=babyweight_trained \
--job-dir=./tmp \
--pattern="00001-of-" --train_steps=1000
```
Once the code works in standalone mode, you can run it on Cloud ML Engine. Because this is on the entire dataset, it will take a while. The training run took about <b> an hour </b> for me. You can monitor the job from the GCP console in the Cloud Machine Learning Engine section.
```
%bash
OUTDIR=gs://${BUCKET}/babyweight/trained_model
JOBNAME=babyweight_$(date -u +%y%m%d_%H%M%S)
echo $OUTDIR $REGION $JOBNAME
#gsutil -m rm -rf $OUTDIR
gcloud ml-engine jobs submit training $JOBNAME \
--region=$REGION \
--module-name=trainer.task \
--package-path=$(pwd)/babyweight/trainer \
--job-dir=$OUTDIR \
--staging-bucket=gs://$BUCKET \
--scale-tier=STANDARD_1 \
-- \
--bucket=${BUCKET} \
--output_dir=${OUTDIR} \
--train_steps=2000000
```
Training finished with a RMSE of 1 lb. Obviously, this is our first model. We could probably add in some features, discretize the mother's age, and do some hyper-parameter tuning to get to a lower RMSE. I'll leave that to you. If you create a better model, I'd love to hear about it -- please do write a short blog post about what you did, and tweet it at me -- @lak_gcp.
```
from google.datalab.ml import TensorBoard
TensorBoard().start('gs://{}/babyweight/trained_model'.format(BUCKET))
for pid in TensorBoard.list()['pid']:
TensorBoard().stop(pid)
print 'Stopped TensorBoard with pid {}'.format(pid)
```
<table width="70%">
<tr><td><img src="weights.png"/></td><td><img src="rmse.png" /></tr>
</table>
<h2> Deploy trained model </h2>
<p>
Deploying the trained model to act as a REST web service is a simple gcloud call.
```
%bash
gsutil ls gs://${BUCKET}/babyweight/trained_model/export/Servo/
%bash
MODEL_NAME="babyweight"
MODEL_VERSION="v1"
MODEL_LOCATION=$(gsutil ls gs://${BUCKET}/babyweight/trained_model/export/Servo/ | tail -1)
echo "Deleting and deploying $MODEL_NAME $MODEL_VERSION from $MODEL_LOCATION ... this will take a few minutes"
#gcloud ml-engine versions delete ${MODEL_VERSION} --model ${MODEL_NAME}
#gcloud ml-engine models delete ${MODEL_NAME}
gcloud ml-engine models create ${MODEL_NAME} --regions $REGION
gcloud ml-engine versions create ${MODEL_VERSION} --model ${MODEL_NAME} --origin ${MODEL_LOCATION}
```
<h2> Use model to predict </h2>
<p>
Send a JSON request to the endpoint of the service to make it predict a baby's weight ... I am going to try out how well the model would have predicted the weights of our two kids and a couple of variations while we are at it ...
```
from googleapiclient import discovery
from oauth2client.client import GoogleCredentials
import json
credentials = GoogleCredentials.get_application_default()
api = discovery.build('ml', 'v1', credentials=credentials)
request_data = {'instances':
[
{
'is_male': 'True',
'mother_age': 26.0,
'mother_race': 'Asian Indian',
'plurality': 1.0,
'gestation_weeks': 39,
'mother_married': 'True',
'cigarette_use': 'False',
'alcohol_use': 'False'
},
{
'is_male': 'False',
'mother_age': 29.0,
'mother_race': 'Asian Indian',
'plurality': 1.0,
'gestation_weeks': 38,
'mother_married': 'True',
'cigarette_use': 'False',
'alcohol_use': 'False'
},
{
'is_male': 'True',
'mother_age': 26.0,
'mother_race': 'White',
'plurality': 1.0,
'gestation_weeks': 39,
'mother_married': 'True',
'cigarette_use': 'False',
'alcohol_use': 'False'
},
{
'is_male': 'True',
'mother_age': 26.0,
'mother_race': 'White',
'plurality': 2.0,
'gestation_weeks': 37,
'mother_married': 'True',
'cigarette_use': 'False',
'alcohol_use': 'False'
}
]
}
parent = 'projects/%s/models/%s/versions/%s' % (PROJECT, 'babyweight', 'v1')
response = api.projects().predict(body=request_data, name=parent).execute()
print "response={0}".format(response)
```
According to the model, our son would have clocked in at 7.3 lbs and our daughter at 6.8 lbs.
<p>
The weights are off by about 0.5 lbs. Pretty cool!
Copyright 2017 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
| github_jupyter |
### Importing necessary modules
```
import pandas as pd
import numpy as np
import os
import numpy as np
import requests
import json
#Reading the data from csv and storing the df in empty dictionary
years = [2015,2016,2017,2018,2019]
df={}
for year in years:
url = f"../Resources/{year}.csv"
pd.set_option("display.max_rows",800)
df[year] = pd.read_csv(url)
```
### Cleaning the df over the period 2016-2019
```
#Unpacking the df
df_2015,df_2016,df_2017,df_2018,df_2019 = df[2015],df[2016],df[2017],df[2018],df[2019]
# Checking for null values in the df
for year in years:
print(f"""{year} \n{df[year].isna().sum()}
----------""")
# Checked for null values - replaced it with with zero.
df_2018.fillna(0,inplace = True)
df_2018.isna().sum()
# replacing spaces with underscore
for year in years:
df[year].columns = [x.replace(" ","_") for x in df[year].columns.values]
print(f"""{year} \n{df[year].columns}
----------""")
#Removing non-contributing columns of the df
col = ['Lower_Confidence_Interval','Upper_Confidence_Interval','Region','Whisker.high','Whisker.low','Standard_Error']
for year in years:
for value in df[year].columns:
if value in col:
del df[year][value]
df[2019]
#Checking the missing columns of the df
for year in years:
print(f"""{year} \n{df[year].shape}
----------""")
#Calculating the missing dystopian columnn values for 2018 and 2019 and Renaming the columns over that period
for year in years[-2:]:
df[year]['Dystopia_Residual'] = df[year].loc[:,'Score']-df[year].loc[:,'GDP_per_capita':].sum(axis=1)
df[year].rename(columns={
'Score':'Happiness_Score',
'GDP_per_capita':'Economy_(GDP_per_Capita)',
'Social_support':'Family',
'Healthy_life_expectancy':'Health_(Life_Expectancy)',
'Freedom_to_make_life_choices':'Freedom',
'Perceptions_of_corruption':'Trust_(Government_Corruption)',
'Overall_rank':'Happiness_Rank',
'Country_or_region':'Country'
},inplace=True)
#Renaming 2017 df column names
df[2017].rename(columns={
'Happiness.Score':'Happiness_Score',
'Economy..GDP.per.Capita.':'Economy_(GDP_per_Capita)',
'Health..Life.Expectancy.':'Health_(Life_Expectancy)',
'Trust..Government.Corruption.':'Trust_(Government_Corruption)',
'Happiness.Rank':'Happiness_Rank',
'Dystopia.Residual':'Dystopia_Residual'
},inplace=True)
#Showing all the columns of the df are equal size
for year in years:
print(f"""{year} \n{df[year].shape}
----------""")
#Finding any duplicates in df
for year in years:
print(f"""{year} \n{df[year]['Country'].nunique()}
----------""")
#Checking if the right data type in data frames
for year in years:
print(f"""\n{year} \n{df[year].info()}
----------\n""")
for year in years:
print(f"""\n{year} \n{df[year].columns}
----------\n""")
#Restructurin the column names for uniformity in df
#Writing the cleaned df into csv
for year in years:
df[year] = df[year][['Country', 'Happiness_Rank', 'Happiness_Score',
'Economy_(GDP_per_Capita)', 'Family', 'Health_(Life_Expectancy)',
'Freedom', 'Trust_(Government_Corruption)', 'Generosity',
'Dystopia_Residual']]
df[year].to_csv(f"../Output/{year}.csv",index=False)
for year in years:
df[year]['year'] = [year for x in range(len(df[year].Country))]
df_list = [df[x] for x in range(2015,2020)]
concated_df = pd.concat(df_list)
concated_df
concated_df.sort_values('Happiness_Rank',inplace=True)
concated_df.reset_index(drop=True,inplace=True)
concated_df.to_csv(f"../Output/concated_df.csv",index=False)
concated_df.head(50)
for year in years:
print(f"{year} Total Countries: {df[year]['Country'].count()} Last Rank is: {df[year]['Happiness_Rank'].max()}")
# Merging all data frames into one data frame on Country column
merge_df = df_2015.merge(df_2016, on=['Country'], how='outer', suffixes=['_2015','_2016'])
merge_df = merge_df.merge(df_2017, on=['Country'], how='outer')
merge_df = merge_df.merge(df_2018, on=['Country'], how='outer', suffixes=['_2017','_2018'])
merge_df = merge_df.merge(df_2019, on=['Country'], how='outer')
# Adding suffix to 2019 columns
merge_df = merge_df.rename(columns={"Happiness_Rank": "Happiness_Rank_2019",
"Happiness_Score": "Happiness_Score_2019",
"Economy_(GDP_per_Capita)": "Economy_(GDP_per_Capita)_2019",
"Family": "Family_2019",
"Health_(Life_Expectancy)": "Health_(Life_Expectancy)_2019",
"Trust_(Government_Corruption)": "Trust_(Government Corruption)_2019",
"Freedom": "Freedom_2019",
"Generosity": "Generosity_2019",
"Dystopia_Residual": "Dystopia_Residual_2019"
})
merge_df.info()
# Checking for countries that has records in 2015, but not in 2016
df_2015[~df_2015.Country.isin(df_2016.Country)]
# Checking for countries that has records in 2016, but not in 2015
df_2016[~df_2016.Country.isin(df_2015.Country)]
# Checking for countries that has records in later years but not in 2015
merge_df[~merge_df.Country.isin(df_2015.Country)]
# Replacing null value found in merged dataframe with 0
merge_df.fillna(0,inplace = True)
merge_df.isna().sum()
# Checking the country names which are in Merged Data frame, but missing in any of the years data frames
merge_df["Country"].loc[(~merge_df["Country"].isin(df_2015["Country"])) | \
(~merge_df["Country"].isin(df_2016["Country"])) | \
(~merge_df["Country"].isin(df_2017["Country"])) | \
(~merge_df["Country"].isin(df_2018["Country"])) | \
(~merge_df["Country"].isin(df_2019["Country"]))
].sort_values()
# Making the country names matching for those refering to the same country
merge_df["Country"].loc[merge_df.Country == "North Cyprus"] = "Northern Cyprus"
merge_df["Country"].loc[merge_df.Country == "Hong Kong S.A.R., China"] = "Hong Kong"
merge_df["Country"].loc[merge_df.Country == "Taiwan Province of China"] = "Taiwan"
merge_df["Country"].loc[merge_df.Country == "Macedonia"] = "North Macedonia"
merge_df["Country"].loc[merge_df.Country == "Trinidad & Tobago"] = "Trinidad and Tobago"
merge_df["Country"].loc[merge_df.Country == "Somaliland region"] = "Somaliland Region"
# Checking the country names which were not present in any of the years data frames
country_check_list = merge_df["Country"].loc[(~merge_df["Country"].isin(df_2015["Country"])) | \
(~merge_df["Country"].isin(df_2016["Country"])) | \
(~merge_df["Country"].isin(df_2017["Country"])) | \
(~merge_df["Country"].isin(df_2018["Country"])) | \
(~merge_df["Country"].isin(df_2019["Country"]))
].sort_values().tolist()
country_check_list = set(country_check_list)
country_check_list
# Displaying the list of Countries missing data in some year
merge_df.loc[merge_df["Country"].isin(country_check_list)]
# Creating a list of country names duplicated
dup_countries = merge_df['Country'].loc[merge_df['Country'].duplicated()].tolist()
dup_countries
# Merging duplicate country names row wise by getting sum of each column values
# Loop through each duplicated country names in the list
for country in dup_countries:
# Making a new data frame having only the duplicated country names
joined_rows = merge_df.loc[merge_df.Country == country]
# Adding a row to the new data frame with the sum of each columns
joined_rows.loc[country,:] = joined_rows.sum(axis=0)
# Correcting the Country column value
joined_rows['Country'] = country
# Removing those rows from the merged data frame
merge_df.drop(merge_df[merge_df.Country == country].index, inplace=True)
# Concatenating the last row added(sum) to the original merged data frame
merge_df = pd.concat([merge_df, joined_rows.tail(1)])
# Reseting the merged data frame's index
merge_df.reset_index(drop=True, inplace=True)
# Displaying the countries data which were missing in any of the year's data frame - after cleaning
merge_df.loc[merge_df["Country"].isin(country_check_list)]
# Replacing 0 values with NAN
cols = merge_df.columns
merge_df[cols] = merge_df[cols].replace({'0':np.nan, 0:np.nan})
merge_df.sort_values
# Writing the cleaned merged data frame to a csv file
merge_df.to_csv("../Output/happiness_merged.csv", index=False)
```
### Getting Region data using API
```
# Loading data from merged csv
df = pd.read_csv("../Output/happiness_merged.csv")
# URL for GET requests to retrieve Region data of Countries from Rest Countries API
base_url = "https://restcountries.eu/rest/v2/name/"
# Defining a function to fetch each Contries Region with API
def fetchRegion(countries):
country_list = []
region_list = []
not_found = []
for country in countries:
try:
# Appending country name to the base url
url = f"{base_url}{country}"
# Perform a get request for each country
response = requests.get(url)
# Storing the JSON response within a variable
data = response.json()
# Adding the region and country name to lists
region_list.append(data[0]['region'])
country_list.append(country)
except:
not_found.append(country)
# function return all the list of values
return country_list, region_list, not_found
# Creating a list of all countries from each years data
countries = df['Country']
# Calling out the function to fetch regions of countries in countries list
country_list, region_list, not_found = fetchRegion(countries)
# Creating a data frame with the list values returned from the function
df_region = pd.DataFrame({"Country": country_list, "Region": region_list})
print(f"Countries Not Found: {not_found}")
df_region
# Getting region from nearby location or other name
df_not_found = pd.DataFrame({'Country': not_found,
'Try_Name':['Korea', 'Palestine',
'Congo', 'Congo',
'Somalia', 'Cyprus',
'Macedonia']
})
df_not_found
# Creating a new list of countries to fecth region which didn't get results yet
countries = df_not_found["Try_Name"].tolist()
# Calling out the function to fetch regions of countries in the new list
country_list, region_list, not_found = fetchRegion(countries)
# Creating a data frame with the list values returned from the function
df_found = pd.DataFrame({"Try_Name": country_list, "Region": region_list})
# Checking if any Country's Region is still missing
print(f"Number of Countries Not Found: {len(not_found)}")
df_found
# Merging The 2 data frames to eliminate Try_Name in between
merge_df = df_not_found.merge(df_found, on='Try_Name', how= 'left')
merge_df.drop(['Try_Name'], axis=1, inplace= True)
merge_df
# Concatenating the data frames with Country and Region found in 2 function calls
df_region = pd.concat([df_region, merge_df]).reset_index(drop=True)
df_region
# Merging region with the happiness data frame
df = df_region.merge(df, on="Country", how="left")
# Writing the data back to the merge csv file
df.to_csv('../Output/happiness_merged.csv', index=False)
```
### Suicide data cleaning
```
# Reading all the needed csv data into dataframes
df = pd.read_csv("../Output/happiness_merged.csv")
df_suicide = pd.read_csv("../Resources/suicide_death_rate.csv", encoding='utf-8')
df.head()
# Selecting the needed Columns and dropping the unnecessary from the happiness data frame
df = df.loc[:,['Country',
'Region',
'Happiness_Score_2015',
'Happiness_Score_2016',
'Happiness_Score_2017',
'Happiness_Score_2018',
'Happiness_Score_2019']]
# Renaming the Yearwise Column names
df = df.rename(columns={'Happiness_Score_2015': '2015',
'Happiness_Score_2016': '2016',
'Happiness_Score_2017': '2017',
'Happiness_Score_2018': '2018',
'Happiness_Score_2019': '2019'})
df.info()
# Selecting the needed Columns and dropping the unnecessary from the suicide data frame
df_suicide = df_suicide.loc[:,['location','year','val']]
# Renaming the coulumns to match with Happiness data frame
df_suicide = df_suicide.rename(columns={"location": "Country", "year": "Year", "val": "Suicide_Rate"})
# Changing the type of values in Year column to string to match with column names in happiness data frame
df_suicide.Year = df_suicide.Year.astype('str')
df_suicide.head()
# Checking for Country name mismatches in both data frame
# Listing Country nemes in Happiness but not found in Suicide data frame
mismatch = df[~df.Country.isin(df_suicide.Country)].Country
mismatch
# Checking for Country names in Suicide dataframe which contains, Country names in Happiness dataframe as part of it
temp_df = pd.DataFrame()
for country in mismatch:
temp_df2 = df_suicide.loc[df_suicide['Country'].str.contains(country)]
temp_df = pd.concat([temp_df, temp_df2])
temp_df
# Changing the Country names in Suicide data frame to match with Happiness dataframe
df_suicide["Country"].loc[df_suicide.Country == "United States of America"] = "United States"
df_suicide["Country"].loc[df_suicide.Country == "Venezuela (Bolivarian Republic of)"] = "Venezuela"
df_suicide["Country"].loc[df_suicide.Country == "Bolivia (Plurinational State of)"] = "Bolivia"
df_suicide["Country"].loc[df_suicide.Country == "Republic of Moldova"] = "Moldova"
df_suicide["Country"].loc[df_suicide.Country == "Russian Federation"] = "Russia"
df_suicide["Country"].loc[df_suicide.Country == "Iran (Islamic Republic of)"] = "Iran"
df_suicide["Country"].loc[df_suicide.Country == "United Republic of Tanzania"] = "Tanzania"
df_suicide["Country"].loc[df_suicide.Country == "Syrian Arab Republic"] = "Syria"
df_suicide["Country"].loc[df_suicide.Country == "Taiwan (Province of China)"] = "Taiwan"
# Again checking for mismatch, Listing Country nemes in Happiness but not found in Suicide data frame
mismatch = df[~df.Country.isin(df_suicide.Country)].Country
mismatch
# Checking for Country names in Suicide dataframe which starts with first 4 characters of,
# Country names in Happiness dataframe as part of it
temp_df = pd.DataFrame()
for country in mismatch:
temp_df2 = df_suicide.loc[df_suicide['Country'].str.startswith(country[:4])]
temp_df = pd.concat([temp_df, temp_df2])
temp_df
# Changing the Country names in Suicide data frame to match with Happiness dataframe
df_suicide["Country"].loc[df_suicide.Country == "Czechia"] = "Czech Republic"
df_suicide["Country"].loc[df_suicide.Country == "Viet Nam"] = "Vietnam"
df_suicide["Country"].loc[df_suicide.Country == "Republic of Korea"] = "South Korea"
df_suicide["Country"].loc[df_suicide.Country == "Palestine"] = "Palestinian Territories"
df_suicide["Country"].loc[df_suicide.Country == "Democratic Republic of the Congo"] = "Congo (Kinshasa)"
df_suicide["Country"].loc[df_suicide.Country == "Congo"] = "Congo (Brazzaville)"
df_suicide["Country"].loc[df_suicide.Country == "Lao People's Democratic Republic"] = "Laos"
df_suicide["Country"].loc[df_suicide.Country == "Eswatini"] = "Swaziland"
df_suicide["Country"].loc[df_suicide.Country == "Côte d'Ivoire"] = "Ivory Coast"
# Again checking for mismatch, Listing Country nemes in Happiness but not found in Suicide data frame
mismatch = df[~df.Country.isin(df_suicide.Country)].Country
# The mismatch is due to missing data.
mismatch
# Melting the happiness dataframe to make a new column 'Year'
# Listing the values of Year column
value_vars = ['2015', '2016', '2017', '2018', '2019']
# Melting keeping Country and Region same and Creating Year column, assigning year values and corresponding happiness score
df = pd.melt(df, id_vars=['Country','Region'], value_vars=value_vars,
var_name='Year', value_name='Happiness_Score', col_level=None)
df.head()
# Merging Suicide rates to the happiness dataframe
df = df.merge(df_suicide, left_on=["Country","Year"], right_on=["Country","Year"], how="left")
# Dropping rows with happiness score as NAN
df = df[df['Happiness_Score'].notna()]
# Writing the data to a file to easy access in analysis
df.to_csv("../Output/suicide_concat.csv", index = False)
df.head()
```
| github_jupyter |
# import the packages
```
# Add the path of the PyOTIC Software to the system path
import sys
sys.path.append('../..')
sys.path.append('../../../PyOTIC/pyoti/')
#Load investigator module
import pyoti
pyoti.info()
import pyotc
```
### create experiment and set data path
```
# Import os to easily join names of filepaths
import os
# Choose the path, were the experiment should be created (or opened from)
#
# datadir: The path to where the experiment (and the data) are located
# datafile: The name of the file that contains the data. Here it is only used to generate dbfile.
# The data is loaded further down upon creation of a Record.
# dbfile: The name of the database file the experiment is saved to (or loaded from).
datadir = '../exampleData/height_calibration_time_series/'
datafile = 'B01.bin'
# For the name of the experiment, exchange the extension '.bin' with '.fs'
dbfile = os.path.join(datadir, datafile.replace('.bin', '.fs'))
# Create the experiment (this will implicitly create/open the dbfile in datadir)
# and assign it to the variable 'experiment'
experiment = pyoti.open_experiment(dbfile)
cfgfile = '../../../PyOTIC/pyoti/pyoti/etc/record/ASWAD.cfg'
record = experiment.create_record(filename=datafile, # data file to read
cfgfile=cfgfile, # setup specific configuration file
directory=datadir) # data directory
experiment._grs.displayrate = 50
experiment.add_view('psd1', 'alpha', traces=['psdXYZ', 'positionXYZ'])
```
### determine amount of data (time) needed
```
N_avg = 80
ex_freq = 32.0
# posible minimum frequencies
print([ex_freq/ndiv for ndiv in range(1, 6)])
print('choose the minimum frequency:')
min_freq = float(input())
t_psd = 1/min_freq
print('Time for one psd: {0:1.3f} s'.format(t_psd))
t_total = N_avg * t_psd
print('Time needed for measurement: {0:1.3f} s'.format(t_total))
from pyotc import gen_psdm_from_region
region = experiment.view('psd1')
T_msr = 5.0
T_delay = 0.5
N_avg = N_avg
es = pyotc.ExpSetting(temp=29.2, radius=0.43,
temp_err=0.001, radius_err=0.43e-3,
temp_unit='degC', radius_unit='um',
height=-1 * region.get_data('positionZ').mean(),
height_unit='um',
material='ps', medium='water')
# traces to calculate psds of
psd_traces = ['psdX', 'psdY', 'psdZ']
position_unit = 'um'
pm = gen_psdm_from_region(region, T_msr, N_avg, T_delay=1.0, psd_traces=psd_traces, exp_setting=es,
active_calibration=True, ex_freq=ex_freq)
pm.plot_psds()
pf = pyotc.PSDFit(pm)
pf.setup_fit(model='hydro')
bounds = {'psdX': (40, 15e3), 'psdY': (40, 15e3), 'psdZ': (200, 4e3)}
pf.fit_psds(dynamic_bounds=True, bounds=bounds, plot_fits=True)
pf.write_results_to_file(directory=pm.directory, fname=pm.paramfile)
```
| github_jupyter |
## 1.3 Concatenate and visualize
We have downloaded 175 netcdf files, that we will compile into one netcdf file here. Also we will plot the modeled and observed precipitation.
### Import packages
```
##This is so variables get printed within jupyter
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
##import packages
import os
import xarray as xr
import numpy as np
import matplotlib.pyplot as plt
import cartopy
import cartopy.crs as ccrs
##We want the working directory to be the UNSEEN-open directory
pwd = os.getcwd() ##current working directory is UNSEEN-open/Notebooks/1.Download
pwd #print the present working directory
os.chdir(pwd+'/../../') # Change the working directory to UNSEEN-open
os.getcwd() #print the working directory
```
## Concatenate
Here we will create an netcdf file that contains the dimensions lat, lon, number (25 ensembles), time (35 years) and leadtime (5 initialization months). We loop over lead times, and open all 35 years of the lead time and then concatenate those leadtimes.
```
init_months = np.append(np.arange(9,13),1) ## Initialization months 9-12,1 (Sep-Jan)
SEAS5_ld1= xr.open_mfdataset('../UK_example/SEAS5/' + '*01.nc', combine='by_coords') # Load the first lead time
SEAS5= SEAS5_ld1 # Create the xarray dataset to concatenate over
for init_month in init_months[0:-1][::-1]: ## Remove the first that we already have and invert so it is lead time 1-5
print(init_month)
SEAS5_ld = xr.open_mfdataset('../UK_example/SEAS5/' + '*' + "%.2i" % init_month + '.nc', combine='by_coords') # Load the first lead time to concatenate over
SEAS5=xr.concat([SEAS5,SEAS5_ld],dim='leadtime')
SEAS5 = SEAS5.assign_coords(leadtime = np.arange(5) + 1) # assign leadtime coordinates
SEAS5['tprate'] = SEAS5['tprate'] * 1000 * 3600 * 24 ## From m/s to mm/d
SEAS5['tprate'].attrs = {'long_name': 'rainfall',
'units': 'mm/day',
'standard_name': 'thickness_of_rainfall_amount'}
SEAS5.load()
```
## Visualize
Here I will compare SEAS5 to the EOBS historical precipitation. I tried to download EOBS through CDS, but the Product was temporally disabled for maintenance purposes. Here, I use EOBS v17 (insted of v20) through KNMI opendap. It runs from 1950 - 2017. Additionally, I download the recent EOBS data (2019 and 2020) [here](https://surfobs.climate.copernicus.eu/dataaccess/access_eobs_months.php).
I will select February monthly mean precipitation to compare to SEAS5.
```
EOBS = xr.open_dataset('http://opendap.knmi.nl/knmi/thredds/dodsC/e-obs_0.50regular/rr_0.50deg_reg_v17.0.nc')
# Concatenated_file.groupby("time.month").mean(dim=['start','ensemble',"time"])
EOBS = EOBS.sel(time=EOBS['time.month'] == 2) ## Select only February
EOBS = EOBS.groupby("time.year").mean('time') ## And calculate the monthly mean precipitation
EOBS
EOBS = xr.open_dataset('http://opendap.knmi.nl/knmi/thredds/dodsC/e-obs_0.50regular/rr_0.50deg_reg_v17.0.nc')
# Concatenated_file.groupby("time.month").mean(dim=['start','ensemble',"time"])
EOBS = EOBS.resample(time='1m').mean()
EOBS
EOBS = EOBS.sel(time=EOBS['time.month'] == 2) ## Select only February
EOBS
EOBS['rr'].attrs = {'long_name': 'rainfall',
'units': 'mm/day',
'standard_name': 'thickness_of_rainfall_amount'}
EOBS['rr'].mean('time').plot()
EOBS2020 = xr.open_dataset('../UK_example/EOBS/rr_0.25deg_day_2020_grid_ensmean.nc')
EOBS2020 = EOBS2020.resample(time='1m').mean()
EOBS2020 = EOBS2020.sel(time=EOBS2020['time.month'] == 2) ## Select only February
EOBS2020['rr'].sel(latitude=slice(48,60),longitude=slice(-10,5)).plot()
fig, axs = plt.subplots(1, 3,subplot_kw={'projection': ccrs.OSGB()})
SEAS5['tprate'].mean(dim = ['time','leadtime','number']).plot(transform=ccrs.PlateCarree(), vmin = 0, vmax = 8, cmap=plt.cm.Blues, ax=axs[0]) #,cmap=plt.cm.Blues,
EOBS['rr'].mean('time').plot(transform=ccrs.PlateCarree(), vmin = 0, vmax = 8, cmap=plt.cm.Blues, ax=axs[1])
EOBS2020['rr'].plot(transform=ccrs.PlateCarree(), vmin = 0, vmax = 8, cmap=plt.cm.Blues, ax=axs[2])
for ax in axs.flat:
ax.coastlines(resolution='110m')
axs[0].set_title('SEAS5')
axs[1].set_title('EOBS')
axs[2].set_title('EOBS 2020')
EOBS2020
EOBS
(EOBS2020['rr']-EOBS['rr'].mean('time'))
# /EOBS['rr'].std('time')
fig, axs = plt.subplots(1, 2,subplot_kw={'projection': ccrs.OSGB()})
(SEAS5['tprate'].mean(dim = ['time','leadtime','number'])
.sel(latitude=slice(),longitude=slice())
.plot(transform=ccrs.PlateCarree(), vmin = 0, vmax = 8, cmap=plt.cm.Blues, ax=axs[0])) #,cmap=plt.cm.Blues,
EOBS['rr'].mean('time').plot(transform=ccrs.PlateCarree(), vmin = 0, vmax = 8, cmap=plt.cm.Blues, ax=axs[1])
for ax in axs.flat:
ax.coastlines(resolution='110m')
axs[0].set_title('SEAS5')
axs[1].set_title('EOBS')
# np.arange('1950', '2017', dtype='datetime64[Y]')
EOBS['rr'].mean(dim = ['latitude','longitude']).plot(x='time')
Quantiles = SEAS5['tprate'].mean(dim = ['latitude','longitude']).quantile([2.5/100, 0.5, 97.5/100], dim=['number','leadtime'])
Quantiles.plot.line(x='time')
# SEAS5['tprate'].mean(dim = ['latitude','longitude','number','leadtime']).plot.line(x = 'time')
Quantiles = SEAS5['tprate'].mean(dim = ['latitude','longitude']).quantile([2.5/100, 0.5, 97.5/100], dim=['number','leadtime'])
Quantiles.plot.line(x='time')
# plt.plot(Quantiles.sel(quantile=0.5),label = 'SEAS5 Median')
# plt.fill_between(Quantiles.sel(quantile=0.025), Quantiles.sel(quantile=0.975), color='green', alpha=0.2,label = '97.5% percentile')
SEAS5['tprate'].mean(dim = ['latitude','longitude','number']).plot.line(x = 'time')
Quantiles = Tapajos.assign_coords(time=np.tile(time_year,400)).groupby("time.month").quantile([2.5/100, 0.5, 97.5/100, 99/100, 0.999, 0.9999], dim="time")
plt.plot(np.arange(1,13), Quantiles.sel(quantile=0.5),label = 'Median of 2000 years')
plt.fill_between(np.arange(1,13), Quantiles.sel(quantile=0.025), Quantiles.sel(quantile=0.975), color='green', alpha=0.2,label = '97.5% percentile')
```
| github_jupyter |
# Sky with the Moon Up
* https://desi.lbl.gov/trac/wiki/CommissioningCommissioningPlanning/commishdata#NightSky
* https://portal.nersc.gov/project/desi/collab/nightwatch/kpno/20191112/exposures.html
* http://desi-www.kpno.noao.edu:8090/nightsum/nightsum-2019-11-12/nightsum.html
**Data**
* /global/projecta/projectdirs/desi/spectro/redux/daily/exposures/20191112
* /global/projecta/projectdirs/desi/spectro/redux/daily/exposures/20191112
* Exposures 27337-27396
**Data Model**
* https://desidatamodel.readthedocs.io/en/latest/DESI_SPECTRO_REDUX/SPECPROD/exposures/NIGHT/EXPID/sky-CAMERA-EXPID.html
John Moustakas
```
import os, sys, glob, pdb
import numpy as np
import fitsio
from astropy.table import Table, vstack
import astropy.units as u
from astropy.coordinates import SkyCoord
import astropy.time
import astropy.coordinates
# hack!
from astropy.utils.iers import conf
from astropy.utils import iers
conf.auto_max_age = None
iers.Conf.iers_auto_url.set('ftp://cddis.gsfc.nasa.gov/pub/products/iers/finals2000A.all')
loc = astropy.coordinates.EarthLocation.of_site('Kitt Peak')
import desispec.io
import desimodel.io
import desimodel.io
# https://astroplan.readthedocs.io/en/v0.1/_modules/astroplan/moon.html
import ephem
def moon_phase_angle(time, location):
sun = astropy.coordinates.get_sun(time).transform_to(astropy.coordinates.AltAz(
location=location, obstime=time))
moon = astropy.coordinates.get_moon(time, location)
elongation = sun.separation(moon)
return np.arctan2(sun.distance*np.sin(elongation),
moon.distance - sun.distance*np.cos(elongation))
def moon_illumination(time, location):
i = moon_phase_angle(time, location)
k = (1 + np.cos(i))/2.0
return k.value
```
import speclite
import specsim.atmosphere
import specsim.simulator
```
import matplotlib.pyplot as plt
import seaborn as sns
sns.set(context='talk', style='ticks', font_scale=1.1)
%matplotlib inline
allcam = ('b', 'r', 'z')
allspec = ['{}'.format(nn) for nn in np.arange(10)]
topdir = os.getenv('DESI_ROOT')+'/spectro'
```
#### Initialize access to the online database.
https://github.com/desihub/desicmx/blob/master/analysis/gfa/DESI-Online-Database-Tutorial.ipynb
```
dbfile = os.getenv('HOME')+'/software/desi-telemetry-db.yaml'
if not os.path.exists(dbfile):
import getpass
pw = getpass.getpass(prompt='Enter database password: ')
with open(dbfile, 'w') as f:
print('host: db.replicator.dev-cattle.stable.spin.nersc.org', file=f)
print('dbname: desi_dev', file=f)
print('port: 60042', file=f)
print('user: desi_reader', file=f)
print(f'password: {pw}', file=f)
print('Created {}'.format(dbfile))
from desietcimg.db import DB, Exposures, NightTelemetry
db = DB(config_name=dbfile)
ExpInfo = Exposures(db)
print(ExpInfo.what)
TcsInfo = NightTelemetry(db, 'tcs_info')
print(TcsInfo(20191112).columns)
#dir(exp(27339))
#ss = desispec.io.read_sky('/global/project/projectdirs/desi/spectro/redux/daily/exposures/20191112/00027345/sky-r3-00027345.fits')
#ss.header[:40]
#tcsinfo = tcs('20191112')
#ii = np.argmin(np.abs(tcsinfo.mjd.values-58800.22016993))
#tcsinfo.iloc[ii]
fp = desimodel.io.load_fiberpos()
tt = desimodel.io.load_throughput('r')
tt.extinction(tt._wave)
tt = desimodel.io.load_throughput('r')
plt.plot(tt._wave, tt.thru(tt._wave))
plt.plot(tt._wave, tt.atmospheric_throughput(tt._wave))
plt.plot(tt._wave, tt.fiberinput_throughput(tt._wave))
dd = '/global/project/projectdirs/desi/spectro/redux/daily/exposures/20191112/00027345'
for cam in ('b', 'r', 'z'):
tt = desimodel.io.load_throughput(cam)
sr = desispec.io.read_frame(dd+'/frame-{}3-00027345.fits'.format(cam))
ss = desispec.io.read_sky(dd+'/sky-{}3-00027345.fits'.format(cam))
plt.plot(ss.wave, ss.flux[0, :] / tt.thru(ss.wave))
plt.ylim(0, 20000)
sr.fibermap
```
#### Troll the reduced data looking for processed sky exposures.
```
def gather_skyspec_info():
allnights = glob.glob(topdir+'/redux/daily/exposures/20191112')
#print(allnights)
info = []
for fullnight in sorted(allnights[:5]):
night = os.path.basename(fullnight)
tcs = TcsInfo(night)
allexpids = glob.glob(fullnight+'/????????')
#print(allexpids)
for datadir in sorted(allexpids):
expid = os.path.basename(datadir)
#exptime = ExpInfo(expid, 'exptime')
for cam in ('r'): # allcam:
for spec in '3': # allspec:
skyfile = '{}/sky-{}{}-{}.fits'.format(datadir, cam, spec, expid)
#print(skyfile)
if os.path.isfile(skyfile):
#print('Reading {}'.format(skyfile))
sky = desispec.io.read_sky(skyfile)
pdb.set_trace()
# gather metadata for this exposure and pack into an astropy Table
mjdobs = sky.header['MJD-OBS']
indx = np.argmin(np.abs(tcs['mjd'].values-mjdobs))
tel = tcs.iloc[indx]
info1 = Table()
info1['NIGHT'] = [night]
info1['EXPID'] = [expid]
info1['CAMERA'] = [cam]
info1['SPECTROGRAPH'] = [spec] # should get from the header...
info1['MJD-OBS'] = [mjdobs] # mid-exposure?
info1['EXPTIME'] = [sky.header['EXPTIME']]
info1['DATE-OBS'] = [sky.header['DATE-OBS']] # mid-exposure?
info1['DATE-UT'] = tel.date_ut # start of exposure?
info1['MJD'] = tel.mjd
info1['SKYRA'] = tel.sky_ra * u.degree
info1['SKYDEC'] = tel.sky_dec * u.degree
#info1['TARGRA'] = tel.target_ra
#info1['TARGDEC'] = tel.target_dec
info1['AIRMASS'] = tel.airmass.astype('f4')
info1['TEL_MOONRA'] = tel.moon_ra * u.degree
info1['TEL_MOONDEC'] = tel.moon_dec * u.degree
info.append(info1)
#pdb.set_trace()
info = vstack(info)
time = astropy.time.Time(info['MJD-OBS'], format='mjd')
info['MOONPHASE'] = np.array([moon_illumination(astropy.time.Time(tt.to_datetime(), scale='utc'), loc)
for tt in time]).astype('f4')
moonpos = astropy.coordinates.get_moon(time, loc)
info['MOONRA'], info['MOONDEC'] = moonpos.ra, moonpos.dec
# compute the moon separation in degrees
csky = SkyCoord(ra=info['SKYRA'], dec=info['SKYDEC'], frame='icrs')
cmoon = SkyCoord(ra=info['MOONRA'], dec=info['MOONDEC'], frame='icrs')
info['MOONSEP'] = np.array(csky.separation(cmoon).value).astype('f4') # [degree]
# get the zenith angle of the moon
moon_altaz = moonpos.transform_to(astropy.coordinates.AltAz(obstime=time, location=loc))
moon_az = moon_altaz.az.value * u.degree
info['MOONAZ'] = moon_altaz.az
info['MOONALT'] = moon_altaz.alt
info['MOONZD'] = 90.0 * u.degree - moon_altaz.alt
return info
%time info = gather_skyspec_info()
info
stop
```
#### Previous code below here
```
gfadir = '/project/projectdirs/desi/users/ameisner/GFA/reduced/v0001'
```
#### Load the GFA filter curve.
```
def load_gfa_filter():
if False:
rfilt = speclite.filters.load_filter('decam2014-r')
else:
if False:
filtfile = '/global/homes/a/ameisner/ci_throughput/etc/gfa_filter_transmission_DESI-1297.dat'
filtwave, filtresp, _, _ = np.loadtxt(filtfile, unpack=True)
else:
filt = Table.read('/global/homes/a/ameisner/ci_throughput/etc/gfa_throughput-airmass_1.00.fits')
filtwave, filtresp = filt['LAMBDA_NM'], filt['THROUGHPUT']
filtresp[filtresp < 0] = 0
filtresp[0] = 0
filtresp[-1] = 0
srt = np.argsort(filtwave)
filtwave, filtresp = filtwave[srt] * 10, filtresp[srt]
rfilt = speclite.filters.FilterResponse(wavelength=filtwave * u.Angstrom,
response=filtresp,
meta=dict(group_name='gfa', band_name='r'))
return rfilt
rfilt = load_gfa_filter()
plt.plot(rfilt.wavelength, rfilt.response)
```
#### Specify the night and range of GFA and spectrograph exposure IDs.
```
night = 20191112
expid_start, expid_end = 27337, 27396
expids = np.arange(expid_end - expid_start + 1) + expid_start
for col in ('airmass', 'skyra', 'skydec', 'moonangl', 'moonra', 'moondec'):
print(col, ExpInfo(27339, col))
```
#### Read A. Meisner's reductions to get the sky background in the GFAs vs moon separation.
```
def read_gfa(night, expid, raw=False):
sexpid = '{:08d}'.format(expid)
rawfname = topdir+'/data/{0}/{1}/gfa-{1}.fits.fz'.format(night, sexpid)
fname = gfadir+'/{0}/{1}/gfa-{1}_ccds.fits'.format(night, sexpid)
if os.path.isfile(fname):
# get the target-moon separation
hdr = fitsio.read_header(rawfname, ext=1)
#print(hdr)
ra, dec, moonra, moondec = hdr['SKYRA'], hdr['SKYDEC'], hdr['MOONRA'], hdr['MOONDEC']
csky = SkyCoord(ra=ra*u.degree, dec=dec*u.degree, frame='icrs')
cmoon = SkyCoord(ra=moonra*u.degree, dec=moondec*u.degree, frame='icrs')
moonsep = np.array(csky.separation(cmoon).value).astype('f4') # [degree]
#print('Reading {}'.format(fname))
data = Table.read(fname)
#print(data.colnames)
# pack into a table
out = Table()
out['sky_mag_ab'] = [np.mean(data['sky_mag_ab'])]
out['sky_mag_ab_err'] = [np.std(data['sky_mag_ab']) / np.sqrt(len(data))]
out['moonsep'] = [moonsep]
#out['airmass'] = [hdr['AIRMASS']]
out = out[np.argsort(out['moonsep'])[::-1]]
return out
else:
return None
#data = read_gfa(night, 27390)
#data
def gfa_sky(night):
out = []
for expid in expids:
dd = read_gfa(night, expid)
if dd is not None:
out.append(dd)
return vstack(out)
#Table(fitsio.read(os.getenv('DESI_ROOT')+'/users/ameisner/GFA/files/skymags-prelim.fits')).colnames
Table(fitsio.read(os.getenv('DESI_ROOT')+'/users/ameisner/GFA/files/skymags-prelim.with_airmass.fits')).colnames
```
Read Aaron's updated GFA analysis outputs.
```
def gfa_sky_updated():
#gfafile = os.getenv('DESI_ROOT')+'/users/ameisner/GFA/files/skymags-prelim.fits'
gfafile = os.getenv('DESI_ROOT')+'/users/ameisner/GFA/files/skymags-prelim.with_airmass.fits'
cat = Table(fitsio.read(gfafile, lower=True))
cat = cat[(cat['expid'] >= expid_start) * (cat['expid'] <= expid_end)]
ra, dec, moonra, moondec = cat['skyra'], cat['skydec'], cat['ra_moon_deg'], cat['dec_moon_deg']
#print(moonra, moondec)
csky = SkyCoord(ra=ra*u.degree, dec=dec*u.degree, frame='icrs')
cmoon = SkyCoord(ra=moonra*u.degree, dec=moondec*u.degree, frame='icrs')
moonsep = np.array(csky.separation(cmoon).value) # [degree]
out = Table()
out['camera'] = cat['extname']
out['expid'] = cat['expid']
out['airmass'] = cat['airmass']
out['ra'] = cat['skyra']
out['dec'] = cat['skydec']
out['moon_phase'] = cat['moon_phase']
out['sky_mag_ab'] = cat['skymag_median_top_camera'].astype('f4')
#out['sky_mag_ab_err'] = [np.std(data['sky_mag_ab']) / np.sqrt(len(data))]
out['moonsep'] = moonsep.astype('f4')
out = out[np.argsort(out['moonsep'])[::-1]]
return out
if False:
gfa = gfa_sky(night)
else:
gfa = gfa_sky_updated()
gfa
fig, ax = plt.subplots(figsize=(8, 6))
for cam, mark in zip(set(gfa['camera']), ('s', 'o')):
ww = gfa['camera'] == cam
ax.scatter(gfa['moonsep'][ww], gfa['sky_mag_ab'][ww],
label=cam, marker=mark, s=80, alpha=0.7)
#ax.invert_yaxis()
#ax.set_ylim(17.5, 14.5)
ax.set_xlabel('Target-Moon Separation (degree)')
ax.set_ylabel('Sky Brightness (AB mag)')
ax.legend()
```
#### Read the spectroscopic reductions to get the sky spectra vs moon separation.
```
skymodel = specsim.simulator.Simulator('desi').atmosphere
skymodel.airmass = 1.1
skymodel.moon.moon_phase = 0.05
skymodel.moon.moon_zenith = 40 * u.deg
for moonsep in (45, 10):
skymodel.moon.separation_angle = moonsep * u.deg
skymodelflux = (skymodel.surface_brightness * np.pi * (0.75 * u.arcsec) ** 2).to(
u.erg / (u.Angstrom * u.cm ** 2 * u.s))
plt.plot(skymodel._wavelength, skymodelflux.value)
def read_spec(night, expid):
sexpid = '{:08d}'.format(expid)
datadir = topdir+'/redux/daily/exposures/{}/{}/'.format(night, sexpid)
if os.path.isdir(datadir):
#fr = desispec.io.read_frame('{}/frame-r3-{}.fits'.format(datadir, sexpid))
sp = desispec.io.read_frame('{}/sframe-r3-{}.fits'.format(datadir, sexpid))
sky = desispec.io.read_sky('{}/sky-r3-{}.fits'.format(datadir, sexpid))
return sp, sky
#return fr, sp, sky
else:
return None, None
#sp, sky = read_spec(night, 27339)
def spec_sky():
import astropy.time
import astropy.coordinates
# hack!
from astropy.utils.iers import conf
from astropy.utils import iers
conf.auto_max_age = None
iers.Conf.iers_auto_url.set('ftp://cddis.gsfc.nasa.gov/pub/products/iers/finals2000A.all')
moonphase = 0.99 # from elog
infastropy.coordinates.EarthLocation.of_site('Kitt Peak')
# skymodel
skymodel = specsim.simulator.Simulator('desi').atmosphere
_rfilt = speclite.filters.FilterSequence([rfilt])
rand = np.random.RandomState(seed=1)
allmoonsep, meansky, stdsky, skymodelmag = [], [], [], []
specsky, specwave = [], []
for expid in expids:
sp, sky = read_spec(night, expid)
if sky is not None:
# blarg! there's no metadata in the headers (or in the database), so
# choose the RA, Dec, and airmass from the nearest GFA exposure.
this = np.argmin(np.abs(gfa['expid'] - expid))
ra, dec = gfa['ra'][this], gfa['dec'][this]
moonphase, airmass = gfa['moon_phase'][this], gfa['airmass'][this],
# get the object-moon separation!!
#moonsep.append(sky.header['PROGRAM'][:3])
hdr = sky.header
date, obstime = hdr['DATE-OBS'], hdr['TIME-OBS']
mjd, exptime = hdr['MJD-OBS'], hdr['EXPTIME']
time = astropy.time.Time(mjd, format='mjd')
#time = astropy.time.Time(date, format='isot', scale='utc')
moonpos = astropy.coordinates.get_moon(time, loc)
moonra, moondec = moonpos.ra, moonpos.dec
csky = SkyCoord(ra=ra*u.degree, dec=dec*u.degree, frame='icrs')
cmoon = SkyCoord(ra=moonra, dec=moondec, frame='icrs')
moonsep = csky.separation(cmoon)
allmoonsep.append(moonsep.value)
moon_altaz = moonpos.transform_to(astropy.coordinates.AltAz(obstime=time, location=loc))
moon_az = moon_altaz.az.value * u.degree
moon_zenith = (90. - moon_altaz.alt.value) * u.degree
# get the model sky brightness
skymodel.airmass = airmass
skymodel.moon.moon_phase = 1 - moonphase
skymodel.moon.moon_zenith = moon_zenith
skymodel.moon.separation_angle = moonsep
print(skymodel.airmass, skymodel.moon.moon_phase, skymodel.moon.moon_zenith.value,
skymodel.moon.separation_angle.value)
skymodelflux = (skymodel.surface_brightness * np.pi * (0.75 * u.arcsec) ** 2).to(
u.erg / (u.Angstrom * u.cm ** 2 * u.s))
pad_skymodelflux, pad_skymodelwave = _rfilt.pad_spectrum(
skymodelflux.value, skymodel._wavelength)
#plt.plot(pad_skymodelwave, pad_skymodelflux)
#import pdb ; pdb.set_trace()
skymodel_abmags = _rfilt.get_ab_magnitudes(pad_skymodelflux, pad_skymodelwave)[rfilt.name]
skymodelmag.append(skymodel_abmags)
# now the data: convolve the spectrum with the r-band filter curve
keep = sp.fibermap['OBJTYPE'] == 'SKY'
padflux, padwave = _rfilt.pad_spectrum(sky.flux[keep, :], sky.wave, method='edge')
abmags = _rfilt.get_ab_magnitudes(padflux, padwave)[rfilt.name]
meansky.append(np.mean(abmags))
stdsky.append(np.std(abmags) / np.sqrt(len(abmags)))
# get the median spectra for this moon separation
specwave.append(sky.wave)
specsky.append(np.percentile(sky.flux[keep, :], axis=0, q=50))
#q25sky = np.percentile(sky.flux[keep, :], axis=0, q=25)
#q75sky = np.percentile(sky.flux[keep, :], axis=0, q=75)
#these = rand.choice(keep, size=20, replace=False)
#[plt.plot(sky.wave, sky.flux[ii, :], alpha=1.0) for ii in these]
#plt.plot(sky.wave, medsky, alpha=0.7, color='k')
out = Table()
out['moonsep'] = np.hstack(allmoonsep).astype('f4')
out['meansky'] = np.hstack(meansky).astype('f4')
out['stdsky'] = np.hstack(stdsky).astype('f4')
out['specwave'] = specwave
out['specsky'] = specsky
out['skymodel'] = np.hstack(skymodelmag).astype('f4')
out = out[np.argsort(out['moonsep'])[::-1]]
return out
spec = spec_sky()
spec
fig, ax = plt.subplots(figsize=(8, 6))
ref = spec['moonsep'].argmax()
#print(ref, spec['specsky'][ref])
for ss in spec:
#print(ss['specwave'], ss['specsky'] / spec['specsky'][ref])
ax.plot(ss['specwave'], ss['specsky'],# / spec['specsky'][0],
label=r'{:.0f}$^{{\circ}}$'.format(ss['moonsep']))
ax.plot(rfilt.wavelength, rfilt.response / np.max(rfilt.response) * np.median(spec['specsky'][-1]),
color='k', ls='--', lw=2)
ax.set_xlim(5500, 7800)
#ax.set_yscale('log')
ax.set_xlabel('Wavelength ($\AA$)')
ax.set_ylabel('Sky Spectra (counts / $\AA$)')
ax.legend(ncol=5, fontsize=12)
#ax.set_title('Night {}'.format(night))
fig, ax = plt.subplots(figsize=(8, 6))
for cam, mark in zip(set(gfa['camera']), ('s', 'o')):
ww = gfa['camera'] == cam
ax.plot(gfa['moonsep'][ww], gfa['sky_mag_ab'][ww], '{}-'.format(mark),
alpha=0.8, markersize=10, label='GFA-{}'.format(cam))
ax.plot(spec['moonsep'], spec['skymodel'], 'o--', color='k', label='Model Sky')
ax.invert_xaxis()
#ax.set_ylim(17.5, 14.5)
ax.set_xlabel('Target-Moon Separation (degree)')
ax.set_ylabel('$r$-band Sky Brightness (AB mag)')
ax.legend()
fig, ax = plt.subplots(figsize=(10, 8))
specref = spec['meansky'][spec['moonsep'].argmax()]
for cam, mark in zip(set(gfa['camera']), ('s', 'o')):
ww = gfa['camera'] == cam
gfaref = gfa['sky_mag_ab'][gfa['moonsep'].argmax()]
ax.plot(gfa['moonsep'][ww], gfa['sky_mag_ab'][ww] - gfaref, '{}-'.format(mark),
alpha=0.8, markersize=10,
label='GFA-{}'.format(cam))#, marker=mark, s=80)
#ax.scatter(gfa['moonsep'], gfa['sky_mag_ab'] - gfaref, label='GFAs')
ax.plot(spec['moonsep'], spec['meansky'] - specref, '^--',
alpha=0.8, markersize=13, label='Spectra')
modelref = spec['skymodel'][spec['moonsep'].argmax()]
ax.plot(spec['moonsep'], spec['skymodel'] - modelref, 'p-.',
alpha=0.8, markersize=13, label='Sky Model')
ax.invert_xaxis()
#ax.set_ylim(17.5, 14.5)
ax.set_xlabel('Target-Moon Separation (degree)')
ax.set_ylabel('Relative $r$-band Sky Brightness (AB mag)')
ax.legend(loc='lower left')
#ax.set_title('Night {}'.format(night))
```
#### Playing around
```
import specsim.simulator
desi = specsim.simulator.Simulator('desi', num_fibers=1)
desi.instrument.fiberloss_method = 'table'
desi.simulate()
plt.plot(desi.atmosphere._wavelength.value, desi.atmosphere.surface_brightness.value)
plt.xlim(5500, 8500)
from astropy.coordinates import EarthLocation
loc = EarthLocation.of_site('Kitt Peak')
loc
time = astropy.time.Time(t['DATE'], format='jd')
moon_position = astropy.coordinates.get_moon(time, loc)
moon_ra = moon_position.ra.value
moon_dec = moon_position.dec.value
moon_position_altaz = moon_position.transform_to(astropy.coordinates.AltAz(obstime=time, location=location))
moon_alt = moon_position_altaz.alt.value
moon_az = moon_position_altaz.az.value
EarthLocation.from_geodetic(lat='-30d10m10.78s', lon='-70d48m23.49s', height=2241.4*u.m)
```
| github_jupyter |
```
# Copyright 2020 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Vertex client library: Custom training image classification model for online prediction
<table align="left">
<td>
<a href="https://colab.research.google.com/github/GoogleCloudPlatform/vertex-ai-samples/blob/master/notebooks/community/gapic/custom/showcase_custom_image_classification_online.ipynb">
<img src="https://cloud.google.com/ml-engine/images/colab-logo-32px.png" alt="Colab logo"> Run in Colab
</a>
</td>
<td>
<a href="https://github.com/GoogleCloudPlatform/vertex-ai-samples/blob/master/notebooks/community/gapic/custom/showcase_custom_image_classification_online.ipynb">
<img src="https://cloud.google.com/ml-engine/images/github-logo-32px.png" alt="GitHub logo">
View on GitHub
</a>
</td>
</table>
<br/><br/><br/>
## Overview
This tutorial demonstrates how to use the Vertex client library for Python to train and deploy a custom image classification model for online prediction.
### Dataset
The dataset used for this tutorial is the [CIFAR10 dataset](https://www.tensorflow.org/datasets/catalog/cifar10) from [TensorFlow Datasets](https://www.tensorflow.org/datasets/catalog/overview). The version of the dataset you will use is built into TensorFlow. The trained model predicts which type of class an image is from ten classes: airplane, automobile, bird, cat, deer, dog, frog, horse, ship, truck.
### Objective
In this tutorial, you create a custom model from a Python script in a Google prebuilt Docker container using the Vertex client library, and then do a prediction on the deployed model by sending data. You can alternatively create custom models using `gcloud` command-line tool or online using Google Cloud Console.
The steps performed include:
- Create a Vertex custom job for training a model.
- Train a TensorFlow model.
- Retrieve and load the model artifacts.
- View the model evaluation.
- Upload the model as a Vertex `Model` resource.
- Deploy the `Model` resource to a serving `Endpoint` resource.
- Make a prediction.
- Undeploy the `Model` resource.
### Costs
This tutorial uses billable components of Google Cloud (GCP):
* Vertex AI
* Cloud Storage
Learn about [Vertex AI
pricing](https://cloud.google.com/vertex-ai/pricing) and [Cloud Storage
pricing](https://cloud.google.com/storage/pricing), and use the [Pricing
Calculator](https://cloud.google.com/products/calculator/)
to generate a cost estimate based on your projected usage.
## Installation
Install the latest version of Vertex client library.
```
import os
import sys
# Google Cloud Notebook
if os.path.exists("/opt/deeplearning/metadata/env_version"):
USER_FLAG = "--user"
else:
USER_FLAG = ""
! pip3 install -U google-cloud-aiplatform $USER_FLAG
```
Install the latest GA version of *google-cloud-storage* library as well.
```
! pip3 install -U google-cloud-storage $USER_FLAG
```
### Restart the kernel
Once you've installed the Vertex client library and Google *cloud-storage*, you need to restart the notebook kernel so it can find the packages.
```
if not os.getenv("IS_TESTING"):
# Automatically restart kernel after installs
import IPython
app = IPython.Application.instance()
app.kernel.do_shutdown(True)
```
## Before you begin
### GPU runtime
*Make sure you're running this notebook in a GPU runtime if you have that option. In Colab, select* **Runtime > Change Runtime Type > GPU**
### Set up your Google Cloud project
**The following steps are required, regardless of your notebook environment.**
1. [Select or create a Google Cloud project](https://console.cloud.google.com/cloud-resource-manager). When you first create an account, you get a $300 free credit towards your compute/storage costs.
2. [Make sure that billing is enabled for your project.](https://cloud.google.com/billing/docs/how-to/modify-project)
3. [Enable the Vertex APIs and Compute Engine APIs.](https://console.cloud.google.com/flows/enableapi?apiid=ml.googleapis.com,compute_component)
4. [The Google Cloud SDK](https://cloud.google.com/sdk) is already installed in Google Cloud Notebook.
5. Enter your project ID in the cell below. Then run the cell to make sure the
Cloud SDK uses the right project for all the commands in this notebook.
**Note**: Jupyter runs lines prefixed with `!` as shell commands, and it interpolates Python variables prefixed with `$` into these commands.
```
PROJECT_ID = "[your-project-id]" # @param {type:"string"}
if PROJECT_ID == "" or PROJECT_ID is None or PROJECT_ID == "[your-project-id]":
# Get your GCP project id from gcloud
shell_output = !gcloud config list --format 'value(core.project)' 2>/dev/null
PROJECT_ID = shell_output[0]
print("Project ID:", PROJECT_ID)
! gcloud config set project $PROJECT_ID
```
#### Region
You can also change the `REGION` variable, which is used for operations
throughout the rest of this notebook. Below are regions supported for Vertex. We recommend that you choose the region closest to you.
- Americas: `us-central1`
- Europe: `europe-west4`
- Asia Pacific: `asia-east1`
You may not use a multi-regional bucket for training with Vertex. Not all regions provide support for all Vertex services. For the latest support per region, see the [Vertex locations documentation](https://cloud.google.com/vertex-ai/docs/general/locations)
```
REGION = "us-central1" # @param {type: "string"}
```
#### Timestamp
If you are in a live tutorial session, you might be using a shared test account or project. To avoid name collisions between users on resources created, you create a timestamp for each instance session, and append onto the name of resources which will be created in this tutorial.
```
from datetime import datetime
TIMESTAMP = datetime.now().strftime("%Y%m%d%H%M%S")
```
### Authenticate your Google Cloud account
**If you are using Google Cloud Notebook**, your environment is already authenticated. Skip this step.
**If you are using Colab**, run the cell below and follow the instructions when prompted to authenticate your account via oAuth.
**Otherwise**, follow these steps:
In the Cloud Console, go to the [Create service account key](https://console.cloud.google.com/apis/credentials/serviceaccountkey) page.
**Click Create service account**.
In the **Service account name** field, enter a name, and click **Create**.
In the **Grant this service account access to project** section, click the Role drop-down list. Type "Vertex" into the filter box, and select **Vertex Administrator**. Type "Storage Object Admin" into the filter box, and select **Storage Object Admin**.
Click Create. A JSON file that contains your key downloads to your local environment.
Enter the path to your service account key as the GOOGLE_APPLICATION_CREDENTIALS variable in the cell below and run the cell.
```
# If you are running this notebook in Colab, run this cell and follow the
# instructions to authenticate your GCP account. This provides access to your
# Cloud Storage bucket and lets you submit training jobs and prediction
# requests.
# If on Google Cloud Notebook, then don't execute this code
if not os.path.exists("/opt/deeplearning/metadata/env_version"):
if "google.colab" in sys.modules:
from google.colab import auth as google_auth
google_auth.authenticate_user()
# If you are running this notebook locally, replace the string below with the
# path to your service account key and run this cell to authenticate your GCP
# account.
elif not os.getenv("IS_TESTING"):
%env GOOGLE_APPLICATION_CREDENTIALS ''
```
### Create a Cloud Storage bucket
**The following steps are required, regardless of your notebook environment.**
When you submit a custom training job using the Vertex client library, you upload a Python package
containing your training code to a Cloud Storage bucket. Vertex runs
the code from this package. In this tutorial, Vertex also saves the
trained model that results from your job in the same bucket. You can then
create an `Endpoint` resource based on this output in order to serve
online predictions.
Set the name of your Cloud Storage bucket below. Bucket names must be globally unique across all Google Cloud projects, including those outside of your organization.
```
BUCKET_NAME = "gs://[your-bucket-name]" # @param {type:"string"}
if BUCKET_NAME == "" or BUCKET_NAME is None or BUCKET_NAME == "gs://[your-bucket-name]":
BUCKET_NAME = "gs://" + PROJECT_ID + "aip-" + TIMESTAMP
```
**Only if your bucket doesn't already exist**: Run the following cell to create your Cloud Storage bucket.
```
! gsutil mb -l $REGION $BUCKET_NAME
```
Finally, validate access to your Cloud Storage bucket by examining its contents:
```
! gsutil ls -al $BUCKET_NAME
```
### Set up variables
Next, set up some variables used throughout the tutorial.
### Import libraries and define constants
#### Import Vertex client library
Import the Vertex client library into our Python environment.
```
import time
from google.cloud.aiplatform import gapic as aip
from google.protobuf import json_format
from google.protobuf.json_format import MessageToJson, ParseDict
from google.protobuf.struct_pb2 import Struct, Value
```
#### Vertex constants
Setup up the following constants for Vertex:
- `API_ENDPOINT`: The Vertex API service endpoint for dataset, model, job, pipeline and endpoint services.
- `PARENT`: The Vertex location root path for dataset, model, job, pipeline and endpoint resources.
```
# API service endpoint
API_ENDPOINT = "{}-aiplatform.googleapis.com".format(REGION)
# Vertex location root path for your dataset, model and endpoint resources
PARENT = "projects/" + PROJECT_ID + "/locations/" + REGION
```
#### Hardware Accelerators
Set the hardware accelerators (e.g., GPU), if any, for training and prediction.
Set the variables `TRAIN_GPU/TRAIN_NGPU` and `DEPLOY_GPU/DEPLOY_NGPU` to use a container image supporting a GPU and the number of GPUs allocated to the virtual machine (VM) instance. For example, to use a GPU container image with 4 Nvidia Telsa K80 GPUs allocated to each VM, you would specify:
(aip.AcceleratorType.NVIDIA_TESLA_K80, 4)
For GPU, available accelerators include:
- aip.AcceleratorType.NVIDIA_TESLA_K80
- aip.AcceleratorType.NVIDIA_TESLA_P100
- aip.AcceleratorType.NVIDIA_TESLA_P4
- aip.AcceleratorType.NVIDIA_TESLA_T4
- aip.AcceleratorType.NVIDIA_TESLA_V100
Otherwise specify `(None, None)` to use a container image to run on a CPU.
*Note*: TF releases before 2.3 for GPU support will fail to load the custom model in this tutorial. It is a known issue and fixed in TF 2.3 -- which is caused by static graph ops that are generated in the serving function. If you encounter this issue on your own custom models, use a container image for TF 2.3 with GPU support.
```
if os.getenv("IS_TESTING_TRAIN_GPU"):
TRAIN_GPU, TRAIN_NGPU = (
aip.AcceleratorType.NVIDIA_TESLA_K80,
int(os.getenv("IS_TESTING_TRAIN_GPU")),
)
else:
TRAIN_GPU, TRAIN_NGPU = (aip.AcceleratorType.NVIDIA_TESLA_K80, 1)
if os.getenv("IS_TESTING_DEPOLY_GPU"):
DEPLOY_GPU, DEPLOY_NGPU = (
aip.AcceleratorType.NVIDIA_TESLA_K80,
int(os.getenv("IS_TESTING_DEPOLY_GPU")),
)
else:
DEPLOY_GPU, DEPLOY_NGPU = (None, None)
```
#### Container (Docker) image
Next, we will set the Docker container images for training and prediction
- TensorFlow 1.15
- `gcr.io/cloud-aiplatform/training/tf-cpu.1-15:latest`
- `gcr.io/cloud-aiplatform/training/tf-gpu.1-15:latest`
- TensorFlow 2.1
- `gcr.io/cloud-aiplatform/training/tf-cpu.2-1:latest`
- `gcr.io/cloud-aiplatform/training/tf-gpu.2-1:latest`
- TensorFlow 2.2
- `gcr.io/cloud-aiplatform/training/tf-cpu.2-2:latest`
- `gcr.io/cloud-aiplatform/training/tf-gpu.2-2:latest`
- TensorFlow 2.3
- `gcr.io/cloud-aiplatform/training/tf-cpu.2-3:latest`
- `gcr.io/cloud-aiplatform/training/tf-gpu.2-3:latest`
- TensorFlow 2.4
- `gcr.io/cloud-aiplatform/training/tf-cpu.2-4:latest`
- `gcr.io/cloud-aiplatform/training/tf-gpu.2-4:latest`
- XGBoost
- `gcr.io/cloud-aiplatform/training/xgboost-cpu.1-1`
- Scikit-learn
- `gcr.io/cloud-aiplatform/training/scikit-learn-cpu.0-23:latest`
- Pytorch
- `gcr.io/cloud-aiplatform/training/pytorch-cpu.1-4:latest`
- `gcr.io/cloud-aiplatform/training/pytorch-cpu.1-5:latest`
- `gcr.io/cloud-aiplatform/training/pytorch-cpu.1-6:latest`
- `gcr.io/cloud-aiplatform/training/pytorch-cpu.1-7:latest`
For the latest list, see [Pre-built containers for training](https://cloud.google.com/vertex-ai/docs/training/pre-built-containers).
- TensorFlow 1.15
- `gcr.io/cloud-aiplatform/prediction/tf-cpu.1-15:latest`
- `gcr.io/cloud-aiplatform/prediction/tf-gpu.1-15:latest`
- TensorFlow 2.1
- `gcr.io/cloud-aiplatform/prediction/tf2-cpu.2-1:latest`
- `gcr.io/cloud-aiplatform/prediction/tf2-gpu.2-1:latest`
- TensorFlow 2.2
- `gcr.io/cloud-aiplatform/prediction/tf2-cpu.2-2:latest`
- `gcr.io/cloud-aiplatform/prediction/tf2-gpu.2-2:latest`
- TensorFlow 2.3
- `gcr.io/cloud-aiplatform/prediction/tf2-cpu.2-3:latest`
- `gcr.io/cloud-aiplatform/prediction/tf2-gpu.2-3:latest`
- XGBoost
- `gcr.io/cloud-aiplatform/prediction/xgboost-cpu.1-2:latest`
- `gcr.io/cloud-aiplatform/prediction/xgboost-cpu.1-1:latest`
- `gcr.io/cloud-aiplatform/prediction/xgboost-cpu.0-90:latest`
- `gcr.io/cloud-aiplatform/prediction/xgboost-cpu.0-82:latest`
- Scikit-learn
- `gcr.io/cloud-aiplatform/prediction/sklearn-cpu.0-23:latest`
- `gcr.io/cloud-aiplatform/prediction/sklearn-cpu.0-22:latest`
- `gcr.io/cloud-aiplatform/prediction/sklearn-cpu.0-20:latest`
For the latest list, see [Pre-built containers for prediction](https://cloud.google.com/vertex-ai/docs/predictions/pre-built-containers)
```
if os.getenv("IS_TESTING_TF"):
TF = os.getenv("IS_TESTING_TF")
else:
TF = "2-1"
if TF[0] == "2":
if TRAIN_GPU:
TRAIN_VERSION = "tf-gpu.{}".format(TF)
else:
TRAIN_VERSION = "tf-cpu.{}".format(TF)
if DEPLOY_GPU:
DEPLOY_VERSION = "tf2-gpu.{}".format(TF)
else:
DEPLOY_VERSION = "tf2-cpu.{}".format(TF)
else:
if TRAIN_GPU:
TRAIN_VERSION = "tf-gpu.{}".format(TF)
else:
TRAIN_VERSION = "tf-cpu.{}".format(TF)
if DEPLOY_GPU:
DEPLOY_VERSION = "tf-gpu.{}".format(TF)
else:
DEPLOY_VERSION = "tf-cpu.{}".format(TF)
TRAIN_IMAGE = "gcr.io/cloud-aiplatform/training/{}:latest".format(TRAIN_VERSION)
DEPLOY_IMAGE = "gcr.io/cloud-aiplatform/prediction/{}:latest".format(DEPLOY_VERSION)
print("Training:", TRAIN_IMAGE, TRAIN_GPU, TRAIN_NGPU)
print("Deployment:", DEPLOY_IMAGE, DEPLOY_GPU, DEPLOY_NGPU)
```
#### Machine Type
Next, set the machine type to use for training and prediction.
- Set the variables `TRAIN_COMPUTE` and `DEPLOY_COMPUTE` to configure the compute resources for the VMs you will use for for training and prediction.
- `machine type`
- `n1-standard`: 3.75GB of memory per vCPU.
- `n1-highmem`: 6.5GB of memory per vCPU
- `n1-highcpu`: 0.9 GB of memory per vCPU
- `vCPUs`: number of \[2, 4, 8, 16, 32, 64, 96 \]
*Note: The following is not supported for training:*
- `standard`: 2 vCPUs
- `highcpu`: 2, 4 and 8 vCPUs
*Note: You may also use n2 and e2 machine types for training and deployment, but they do not support GPUs*.
```
if os.getenv("IS_TESTING_TRAIN_MACHINE"):
MACHINE_TYPE = os.getenv("IS_TESTING_TRAIN_MACHINE")
else:
MACHINE_TYPE = "n1-standard"
VCPU = "4"
TRAIN_COMPUTE = MACHINE_TYPE + "-" + VCPU
print("Train machine type", TRAIN_COMPUTE)
if os.getenv("IS_TESTING_DEPLOY_MACHINE"):
MACHINE_TYPE = os.getenv("IS_TESTING_DEPLOY_MACHINE")
else:
MACHINE_TYPE = "n1-standard"
VCPU = "4"
DEPLOY_COMPUTE = MACHINE_TYPE + "-" + VCPU
print("Deploy machine type", DEPLOY_COMPUTE)
```
# Tutorial
Now you are ready to start creating your own custom model and training for CIFAR10.
## Set up clients
The Vertex client library works as a client/server model. On your side (the Python script) you will create a client that sends requests and receives responses from the Vertex server.
You will use different clients in this tutorial for different steps in the workflow. So set them all up upfront.
- Model Service for `Model` resources.
- Endpoint Service for deployment.
- Job Service for batch jobs and custom training.
- Prediction Service for serving.
```
# client options same for all services
client_options = {"api_endpoint": API_ENDPOINT}
def create_job_client():
client = aip.JobServiceClient(client_options=client_options)
return client
def create_model_client():
client = aip.ModelServiceClient(client_options=client_options)
return client
def create_endpoint_client():
client = aip.EndpointServiceClient(client_options=client_options)
return client
def create_prediction_client():
client = aip.PredictionServiceClient(client_options=client_options)
return client
clients = {}
clients["job"] = create_job_client()
clients["model"] = create_model_client()
clients["endpoint"] = create_endpoint_client()
clients["prediction"] = create_prediction_client()
for client in clients.items():
print(client)
```
## Train a model
There are two ways you can train a custom model using a container image:
- **Use a Google Cloud prebuilt container**. If you use a prebuilt container, you will additionally specify a Python package to install into the container image. This Python package contains your code for training a custom model.
- **Use your own custom container image**. If you use your own container, the container needs to contain your code for training a custom model.
## Prepare your custom job specification
Now that your clients are ready, your first step is to create a Job Specification for your custom training job. The job specification will consist of the following:
- `worker_pool_spec` : The specification of the type of machine(s) you will use for training and how many (single or distributed)
- `python_package_spec` : The specification of the Python package to be installed with the pre-built container.
### Prepare your machine specification
Now define the machine specification for your custom training job. This tells Vertex what type of machine instance to provision for the training.
- `machine_type`: The type of GCP instance to provision -- e.g., n1-standard-8.
- `accelerator_type`: The type, if any, of hardware accelerator. In this tutorial if you previously set the variable `TRAIN_GPU != None`, you are using a GPU; otherwise you will use a CPU.
- `accelerator_count`: The number of accelerators.
```
if TRAIN_GPU:
machine_spec = {
"machine_type": TRAIN_COMPUTE,
"accelerator_type": TRAIN_GPU,
"accelerator_count": TRAIN_NGPU,
}
else:
machine_spec = {"machine_type": TRAIN_COMPUTE, "accelerator_count": 0}
```
### Prepare your disk specification
(optional) Now define the disk specification for your custom training job. This tells Vertex what type and size of disk to provision in each machine instance for the training.
- `boot_disk_type`: Either SSD or Standard. SSD is faster, and Standard is less expensive. Defaults to SSD.
- `boot_disk_size_gb`: Size of disk in GB.
```
DISK_TYPE = "pd-ssd" # [ pd-ssd, pd-standard]
DISK_SIZE = 200 # GB
disk_spec = {"boot_disk_type": DISK_TYPE, "boot_disk_size_gb": DISK_SIZE}
```
### Define the worker pool specification
Next, you define the worker pool specification for your custom training job. The worker pool specification will consist of the following:
- `replica_count`: The number of instances to provision of this machine type.
- `machine_spec`: The hardware specification.
- `disk_spec` : (optional) The disk storage specification.
- `python_package`: The Python training package to install on the VM instance(s) and which Python module to invoke, along with command line arguments for the Python module.
Let's dive deeper now into the python package specification:
-`executor_image_spec`: This is the docker image which is configured for your custom training job.
-`package_uris`: This is a list of the locations (URIs) of your python training packages to install on the provisioned instance. The locations need to be in a Cloud Storage bucket. These can be either individual python files or a zip (archive) of an entire package. In the later case, the job service will unzip (unarchive) the contents into the docker image.
-`python_module`: The Python module (script) to invoke for running the custom training job. In this example, you will be invoking `trainer.task.py` -- note that it was not neccessary to append the `.py` suffix.
-`args`: The command line arguments to pass to the corresponding Pythom module. In this example, you will be setting:
- `"--model-dir=" + MODEL_DIR` : The Cloud Storage location where to store the model artifacts. There are two ways to tell the training script where to save the model artifacts:
- direct: You pass the Cloud Storage location as a command line argument to your training script (set variable `DIRECT = True`), or
- indirect: The service passes the Cloud Storage location as the environment variable `AIP_MODEL_DIR` to your training script (set variable `DIRECT = False`). In this case, you tell the service the model artifact location in the job specification.
- `"--epochs=" + EPOCHS`: The number of epochs for training.
- `"--steps=" + STEPS`: The number of steps (batches) per epoch.
- `"--distribute=" + TRAIN_STRATEGY"` : The training distribution strategy to use for single or distributed training.
- `"single"`: single device.
- `"mirror"`: all GPU devices on a single compute instance.
- `"multi"`: all GPU devices on all compute instances.
```
JOB_NAME = "custom_job_" + TIMESTAMP
MODEL_DIR = "{}/{}".format(BUCKET_NAME, JOB_NAME)
if not TRAIN_NGPU or TRAIN_NGPU < 2:
TRAIN_STRATEGY = "single"
else:
TRAIN_STRATEGY = "mirror"
EPOCHS = 20
STEPS = 100
DIRECT = True
if DIRECT:
CMDARGS = [
"--model-dir=" + MODEL_DIR,
"--epochs=" + str(EPOCHS),
"--steps=" + str(STEPS),
"--distribute=" + TRAIN_STRATEGY,
]
else:
CMDARGS = [
"--epochs=" + str(EPOCHS),
"--steps=" + str(STEPS),
"--distribute=" + TRAIN_STRATEGY,
]
worker_pool_spec = [
{
"replica_count": 1,
"machine_spec": machine_spec,
"disk_spec": disk_spec,
"python_package_spec": {
"executor_image_uri": TRAIN_IMAGE,
"package_uris": [BUCKET_NAME + "/trainer_cifar10.tar.gz"],
"python_module": "trainer.task",
"args": CMDARGS,
},
}
]
```
### Assemble a job specification
Now assemble the complete description for the custom job specification:
- `display_name`: The human readable name you assign to this custom job.
- `job_spec`: The specification for the custom job.
- `worker_pool_specs`: The specification for the machine VM instances.
- `base_output_directory`: This tells the service the Cloud Storage location where to save the model artifacts (when variable `DIRECT = False`). The service will then pass the location to the training script as the environment variable `AIP_MODEL_DIR`, and the path will be of the form:
<output_uri_prefix>/model
```
if DIRECT:
job_spec = {"worker_pool_specs": worker_pool_spec}
else:
job_spec = {
"worker_pool_specs": worker_pool_spec,
"base_output_directory": {"output_uri_prefix": MODEL_DIR},
}
custom_job = {"display_name": JOB_NAME, "job_spec": job_spec}
```
### Examine the training package
#### Package layout
Before you start the training, you will look at how a Python package is assembled for a custom training job. When unarchived, the package contains the following directory/file layout.
- PKG-INFO
- README.md
- setup.cfg
- setup.py
- trainer
- \_\_init\_\_.py
- task.py
The files `setup.cfg` and `setup.py` are the instructions for installing the package into the operating environment of the Docker image.
The file `trainer/task.py` is the Python script for executing the custom training job. *Note*, when we referred to it in the worker pool specification, we replace the directory slash with a dot (`trainer.task`) and dropped the file suffix (`.py`).
#### Package Assembly
In the following cells, you will assemble the training package.
```
# Make folder for Python training script
! rm -rf custom
! mkdir custom
# Add package information
! touch custom/README.md
setup_cfg = "[egg_info]\n\ntag_build =\n\ntag_date = 0"
! echo "$setup_cfg" > custom/setup.cfg
setup_py = "import setuptools\n\nsetuptools.setup(\n\n install_requires=[\n\n 'tensorflow_datasets==1.3.0',\n\n ],\n\n packages=setuptools.find_packages())"
! echo "$setup_py" > custom/setup.py
pkg_info = "Metadata-Version: 1.0\n\nName: CIFAR10 image classification\n\nVersion: 0.0.0\n\nSummary: Demostration training script\n\nHome-page: www.google.com\n\nAuthor: Google\n\nAuthor-email: aferlitsch@google.com\n\nLicense: Public\n\nDescription: Demo\n\nPlatform: Vertex"
! echo "$pkg_info" > custom/PKG-INFO
# Make the training subfolder
! mkdir custom/trainer
! touch custom/trainer/__init__.py
```
#### Task.py contents
In the next cell, you write the contents of the training script task.py. We won't go into detail, it's just there for you to browse. In summary:
- Get the directory where to save the model artifacts from the command line (`--model_dir`), and if not specified, then from the environment variable `AIP_MODEL_DIR`.
- Loads CIFAR10 dataset from TF Datasets (tfds).
- Builds a model using TF.Keras model API.
- Compiles the model (`compile()`).
- Sets a training distribution strategy according to the argument `args.distribute`.
- Trains the model (`fit()`) with epochs and steps according to the arguments `args.epochs` and `args.steps`
- Saves the trained model (`save(args.model_dir)`) to the specified model directory.
```
%%writefile custom/trainer/task.py
# Single, Mirror and Multi-Machine Distributed Training for CIFAR-10
import tensorflow_datasets as tfds
import tensorflow as tf
from tensorflow.python.client import device_lib
import argparse
import os
import sys
tfds.disable_progress_bar()
parser = argparse.ArgumentParser()
parser.add_argument('--model-dir', dest='model_dir',
default=os.getenv("AIP_MODEL_DIR"), type=str, help='Model dir.')
parser.add_argument('--lr', dest='lr',
default=0.01, type=float,
help='Learning rate.')
parser.add_argument('--epochs', dest='epochs',
default=10, type=int,
help='Number of epochs.')
parser.add_argument('--steps', dest='steps',
default=200, type=int,
help='Number of steps per epoch.')
parser.add_argument('--distribute', dest='distribute', type=str, default='single',
help='distributed training strategy')
args = parser.parse_args()
print('Python Version = {}'.format(sys.version))
print('TensorFlow Version = {}'.format(tf.__version__))
print('TF_CONFIG = {}'.format(os.environ.get('TF_CONFIG', 'Not found')))
print('DEVICES', device_lib.list_local_devices())
# Single Machine, single compute device
if args.distribute == 'single':
if tf.test.is_gpu_available():
strategy = tf.distribute.OneDeviceStrategy(device="/gpu:0")
else:
strategy = tf.distribute.OneDeviceStrategy(device="/cpu:0")
# Single Machine, multiple compute device
elif args.distribute == 'mirror':
strategy = tf.distribute.MirroredStrategy()
# Multiple Machine, multiple compute device
elif args.distribute == 'multi':
strategy = tf.distribute.experimental.MultiWorkerMirroredStrategy()
# Multi-worker configuration
print('num_replicas_in_sync = {}'.format(strategy.num_replicas_in_sync))
# Preparing dataset
BUFFER_SIZE = 10000
BATCH_SIZE = 64
def make_datasets_unbatched():
# Scaling CIFAR10 data from (0, 255] to (0., 1.]
def scale(image, label):
image = tf.cast(image, tf.float32)
image /= 255.0
return image, label
datasets, info = tfds.load(name='cifar10',
with_info=True,
as_supervised=True)
return datasets['train'].map(scale).cache().shuffle(BUFFER_SIZE).repeat()
# Build the Keras model
def build_and_compile_cnn_model():
model = tf.keras.Sequential([
tf.keras.layers.Conv2D(32, 3, activation='relu', input_shape=(32, 32, 3)),
tf.keras.layers.MaxPooling2D(),
tf.keras.layers.Conv2D(32, 3, activation='relu'),
tf.keras.layers.MaxPooling2D(),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(10, activation='softmax')
])
model.compile(
loss=tf.keras.losses.sparse_categorical_crossentropy,
optimizer=tf.keras.optimizers.SGD(learning_rate=args.lr),
metrics=['accuracy'])
return model
# Train the model
NUM_WORKERS = strategy.num_replicas_in_sync
# Here the batch size scales up by number of workers since
# `tf.data.Dataset.batch` expects the global batch size.
GLOBAL_BATCH_SIZE = BATCH_SIZE * NUM_WORKERS
train_dataset = make_datasets_unbatched().batch(GLOBAL_BATCH_SIZE)
with strategy.scope():
# Creation of dataset, and model building/compiling need to be within
# `strategy.scope()`.
model = build_and_compile_cnn_model()
model.fit(x=train_dataset, epochs=args.epochs, steps_per_epoch=args.steps)
model.save(args.model_dir)
```
#### Store training script on your Cloud Storage bucket
Next, you package the training folder into a compressed tar ball, and then store it in your Cloud Storage bucket.
```
! rm -f custom.tar custom.tar.gz
! tar cvf custom.tar custom
! gzip custom.tar
! gsutil cp custom.tar.gz $BUCKET_NAME/trainer_cifar10.tar.gz
```
### Train the model
Now start the training of your custom training job on Vertex. Use this helper function `create_custom_job`, which takes the following parameter:
-`custom_job`: The specification for the custom job.
The helper function calls job client service's `create_custom_job` method, with the following parameters:
-`parent`: The Vertex location path to `Dataset`, `Model` and `Endpoint` resources.
-`custom_job`: The specification for the custom job.
You will display a handful of the fields returned in `response` object, with the two that are of most interest are:
`response.name`: The Vertex fully qualified identifier assigned to this custom training job. You save this identifier for using in subsequent steps.
`response.state`: The current state of the custom training job.
```
def create_custom_job(custom_job):
response = clients["job"].create_custom_job(parent=PARENT, custom_job=custom_job)
print("name:", response.name)
print("display_name:", response.display_name)
print("state:", response.state)
print("create_time:", response.create_time)
print("update_time:", response.update_time)
return response
response = create_custom_job(custom_job)
```
Now get the unique identifier for the custom job you created.
```
# The full unique ID for the custom job
job_id = response.name
# The short numeric ID for the custom job
job_short_id = job_id.split("/")[-1]
print(job_id)
```
### Get information on a custom job
Next, use this helper function `get_custom_job`, which takes the following parameter:
- `name`: The Vertex fully qualified identifier for the custom job.
The helper function calls the job client service's`get_custom_job` method, with the following parameter:
- `name`: The Vertex fully qualified identifier for the custom job.
If you recall, you got the Vertex fully qualified identifier for the custom job in the `response.name` field when you called the `create_custom_job` method, and saved the identifier in the variable `job_id`.
```
def get_custom_job(name, silent=False):
response = clients["job"].get_custom_job(name=name)
if silent:
return response
print("name:", response.name)
print("display_name:", response.display_name)
print("state:", response.state)
print("create_time:", response.create_time)
print("update_time:", response.update_time)
return response
response = get_custom_job(job_id)
```
# Deployment
Training the above model may take upwards of 20 minutes time.
Once your model is done training, you can calculate the actual time it took to train the model by subtracting `end_time` from `start_time`. For your model, we will need to know the location of the saved model, which the Python script saved in your local Cloud Storage bucket at `MODEL_DIR + '/saved_model.pb'`.
```
while True:
response = get_custom_job(job_id, True)
if response.state != aip.JobState.JOB_STATE_SUCCEEDED:
print("Training job has not completed:", response.state)
model_path_to_deploy = None
if response.state == aip.JobState.JOB_STATE_FAILED:
break
else:
if not DIRECT:
MODEL_DIR = MODEL_DIR + "/model"
model_path_to_deploy = MODEL_DIR
print("Training Time:", response.update_time - response.create_time)
break
time.sleep(60)
print("model_to_deploy:", model_path_to_deploy)
```
## Load the saved model
Your model is stored in a TensorFlow SavedModel format in a Cloud Storage bucket. Now load it from the Cloud Storage bucket, and then you can do some things, like evaluate the model, and do a prediction.
To load, you use the TF.Keras `model.load_model()` method passing it the Cloud Storage path where the model is saved -- specified by `MODEL_DIR`.
```
import tensorflow as tf
model = tf.keras.models.load_model(MODEL_DIR)
```
## Evaluate the model
Now find out how good the model is.
### Load evaluation data
You will load the CIFAR10 test (holdout) data from `tf.keras.datasets`, using the method `load_data()`. This will return the dataset as a tuple of two elements. The first element is the training data and the second is the test data. Each element is also a tuple of two elements: the image data, and the corresponding labels.
You don't need the training data, and hence why we loaded it as `(_, _)`.
Before you can run the data through evaluation, you need to preprocess it:
x_test:
1. Normalize (rescaling) the pixel data by dividing each pixel by 255. This will replace each single byte integer pixel with a 32-bit floating point number between 0 and 1.
y_test:<br/>
2. The labels are currently scalar (sparse). If you look back at the `compile()` step in the `trainer/task.py` script, you will find that it was compiled for sparse labels. So we don't need to do anything more.
```
import numpy as np
from tensorflow.keras.datasets import cifar10
(_, _), (x_test, y_test) = cifar10.load_data()
x_test = (x_test / 255.0).astype(np.float32)
print(x_test.shape, y_test.shape)
```
### Perform the model evaluation
Now evaluate how well the model in the custom job did.
```
model.evaluate(x_test, y_test)
```
## Upload the model for serving
Next, you will upload your TF.Keras model from the custom job to Vertex `Model` service, which will create a Vertex `Model` resource for your custom model. During upload, you need to define a serving function to convert data to the format your model expects. If you send encoded data to Vertex, your serving function ensures that the data is decoded on the model server before it is passed as input to your model.
### How does the serving function work
When you send a request to an online prediction server, the request is received by a HTTP server. The HTTP server extracts the prediction request from the HTTP request content body. The extracted prediction request is forwarded to the serving function. For Google pre-built prediction containers, the request content is passed to the serving function as a `tf.string`.
The serving function consists of two parts:
- `preprocessing function`:
- Converts the input (`tf.string`) to the input shape and data type of the underlying model (dynamic graph).
- Performs the same preprocessing of the data that was done during training the underlying model -- e.g., normalizing, scaling, etc.
- `post-processing function`:
- Converts the model output to format expected by the receiving application -- e.q., compresses the output.
- Packages the output for the the receiving application -- e.g., add headings, make JSON object, etc.
Both the preprocessing and post-processing functions are converted to static graphs which are fused to the model. The output from the underlying model is passed to the post-processing function. The post-processing function passes the converted/packaged output back to the HTTP server. The HTTP server returns the output as the HTTP response content.
One consideration you need to consider when building serving functions for TF.Keras models is that they run as static graphs. That means, you cannot use TF graph operations that require a dynamic graph. If you do, you will get an error during the compile of the serving function which will indicate that you are using an EagerTensor which is not supported.
### Serving function for image data
To pass images to the prediction service, you encode the compressed (e.g., JPEG) image bytes into base 64 -- which makes the content safe from modification while transmitting binary data over the network. Since this deployed model expects input data as raw (uncompressed) bytes, you need to ensure that the base 64 encoded data gets converted back to raw bytes before it is passed as input to the deployed model.
To resolve this, define a serving function (`serving_fn`) and attach it to the model as a preprocessing step. Add a `@tf.function` decorator so the serving function is fused to the underlying model (instead of upstream on a CPU).
When you send a prediction or explanation request, the content of the request is base 64 decoded into a Tensorflow string (`tf.string`), which is passed to the serving function (`serving_fn`). The serving function preprocesses the `tf.string` into raw (uncompressed) numpy bytes (`preprocess_fn`) to match the input requirements of the model:
- `io.decode_jpeg`- Decompresses the JPG image which is returned as a Tensorflow tensor with three channels (RGB).
- `image.convert_image_dtype` - Changes integer pixel values to float 32.
- `image.resize` - Resizes the image to match the input shape for the model.
- `resized / 255.0` - Rescales (normalization) the pixel data between 0 and 1.
At this point, the data can be passed to the model (`m_call`).
```
CONCRETE_INPUT = "numpy_inputs"
def _preprocess(bytes_input):
decoded = tf.io.decode_jpeg(bytes_input, channels=3)
decoded = tf.image.convert_image_dtype(decoded, tf.float32)
resized = tf.image.resize(decoded, size=(32, 32))
rescale = tf.cast(resized / 255.0, tf.float32)
return rescale
@tf.function(input_signature=[tf.TensorSpec([None], tf.string)])
def preprocess_fn(bytes_inputs):
decoded_images = tf.map_fn(
_preprocess, bytes_inputs, dtype=tf.float32, back_prop=False
)
return {
CONCRETE_INPUT: decoded_images
} # User needs to make sure the key matches model's input
@tf.function(input_signature=[tf.TensorSpec([None], tf.string)])
def serving_fn(bytes_inputs):
images = preprocess_fn(bytes_inputs)
prob = m_call(**images)
return prob
m_call = tf.function(model.call).get_concrete_function(
[tf.TensorSpec(shape=[None, 32, 32, 3], dtype=tf.float32, name=CONCRETE_INPUT)]
)
tf.saved_model.save(
model, model_path_to_deploy, signatures={"serving_default": serving_fn}
)
```
## Get the serving function signature
You can get the signatures of your model's input and output layers by reloading the model into memory, and querying it for the signatures corresponding to each layer.
For your purpose, you need the signature of the serving function. Why? Well, when we send our data for prediction as a HTTP request packet, the image data is base64 encoded, and our TF.Keras model takes numpy input. Your serving function will do the conversion from base64 to a numpy array.
When making a prediction request, you need to route the request to the serving function instead of the model, so you need to know the input layer name of the serving function -- which you will use later when you make a prediction request.
```
loaded = tf.saved_model.load(model_path_to_deploy)
serving_input = list(
loaded.signatures["serving_default"].structured_input_signature[1].keys()
)[0]
print("Serving function input:", serving_input)
```
### Upload the model
Use this helper function `upload_model` to upload your model, stored in SavedModel format, up to the `Model` service, which will instantiate a Vertex `Model` resource instance for your model. Once you've done that, you can use the `Model` resource instance in the same way as any other Vertex `Model` resource instance, such as deploying to an `Endpoint` resource for serving predictions.
The helper function takes the following parameters:
- `display_name`: A human readable name for the `Endpoint` service.
- `image_uri`: The container image for the model deployment.
- `model_uri`: The Cloud Storage path to our SavedModel artificat. For this tutorial, this is the Cloud Storage location where the `trainer/task.py` saved the model artifacts, which we specified in the variable `MODEL_DIR`.
The helper function calls the `Model` client service's method `upload_model`, which takes the following parameters:
- `parent`: The Vertex location root path for `Dataset`, `Model` and `Endpoint` resources.
- `model`: The specification for the Vertex `Model` resource instance.
Let's now dive deeper into the Vertex model specification `model`. This is a dictionary object that consists of the following fields:
- `display_name`: A human readable name for the `Model` resource.
- `metadata_schema_uri`: Since your model was built without an Vertex `Dataset` resource, you will leave this blank (`''`).
- `artificat_uri`: The Cloud Storage path where the model is stored in SavedModel format.
- `container_spec`: This is the specification for the Docker container that will be installed on the `Endpoint` resource, from which the `Model` resource will serve predictions. Use the variable you set earlier `DEPLOY_GPU != None` to use a GPU; otherwise only a CPU is allocated.
Uploading a model into a Vertex Model resource returns a long running operation, since it may take a few moments. You call response.result(), which is a synchronous call and will return when the Vertex Model resource is ready.
The helper function returns the Vertex fully qualified identifier for the corresponding Vertex Model instance upload_model_response.model. You will save the identifier for subsequent steps in the variable model_to_deploy_id.
```
IMAGE_URI = DEPLOY_IMAGE
def upload_model(display_name, image_uri, model_uri):
model = {
"display_name": display_name,
"metadata_schema_uri": "",
"artifact_uri": model_uri,
"container_spec": {
"image_uri": image_uri,
"command": [],
"args": [],
"env": [{"name": "env_name", "value": "env_value"}],
"ports": [{"container_port": 8080}],
"predict_route": "",
"health_route": "",
},
}
response = clients["model"].upload_model(parent=PARENT, model=model)
print("Long running operation:", response.operation.name)
upload_model_response = response.result(timeout=180)
print("upload_model_response")
print(" model:", upload_model_response.model)
return upload_model_response.model
model_to_deploy_id = upload_model(
"cifar10-" + TIMESTAMP, IMAGE_URI, model_path_to_deploy
)
```
### Get `Model` resource information
Now let's get the model information for just your model. Use this helper function `get_model`, with the following parameter:
- `name`: The Vertex unique identifier for the `Model` resource.
This helper function calls the Vertex `Model` client service's method `get_model`, with the following parameter:
- `name`: The Vertex unique identifier for the `Model` resource.
```
def get_model(name):
response = clients["model"].get_model(name=name)
print(response)
get_model(model_to_deploy_id)
```
## Deploy the `Model` resource
Now deploy the trained Vertex custom `Model` resource. This requires two steps:
1. Create an `Endpoint` resource for deploying the `Model` resource to.
2. Deploy the `Model` resource to the `Endpoint` resource.
### Create an `Endpoint` resource
Use this helper function `create_endpoint` to create an endpoint to deploy the model to for serving predictions, with the following parameter:
- `display_name`: A human readable name for the `Endpoint` resource.
The helper function uses the endpoint client service's `create_endpoint` method, which takes the following parameter:
- `display_name`: A human readable name for the `Endpoint` resource.
Creating an `Endpoint` resource returns a long running operation, since it may take a few moments to provision the `Endpoint` resource for serving. You call `response.result()`, which is a synchronous call and will return when the Endpoint resource is ready. The helper function returns the Vertex fully qualified identifier for the `Endpoint` resource: `response.name`.
```
ENDPOINT_NAME = "cifar10_endpoint-" + TIMESTAMP
def create_endpoint(display_name):
endpoint = {"display_name": display_name}
response = clients["endpoint"].create_endpoint(parent=PARENT, endpoint=endpoint)
print("Long running operation:", response.operation.name)
result = response.result(timeout=300)
print("result")
print(" name:", result.name)
print(" display_name:", result.display_name)
print(" description:", result.description)
print(" labels:", result.labels)
print(" create_time:", result.create_time)
print(" update_time:", result.update_time)
return result
result = create_endpoint(ENDPOINT_NAME)
```
Now get the unique identifier for the `Endpoint` resource you created.
```
# The full unique ID for the endpoint
endpoint_id = result.name
# The short numeric ID for the endpoint
endpoint_short_id = endpoint_id.split("/")[-1]
print(endpoint_id)
```
### Compute instance scaling
You have several choices on scaling the compute instances for handling your online prediction requests:
- Single Instance: The online prediction requests are processed on a single compute instance.
- Set the minimum (`MIN_NODES`) and maximum (`MAX_NODES`) number of compute instances to one.
- Manual Scaling: The online prediction requests are split across a fixed number of compute instances that you manually specified.
- Set the minimum (`MIN_NODES`) and maximum (`MAX_NODES`) number of compute instances to the same number of nodes. When a model is first deployed to the instance, the fixed number of compute instances are provisioned and online prediction requests are evenly distributed across them.
- Auto Scaling: The online prediction requests are split across a scaleable number of compute instances.
- Set the minimum (`MIN_NODES`) number of compute instances to provision when a model is first deployed and to de-provision, and set the maximum (`MAX_NODES) number of compute instances to provision, depending on load conditions.
The minimum number of compute instances corresponds to the field `min_replica_count` and the maximum number of compute instances corresponds to the field `max_replica_count`, in your subsequent deployment request.
```
MIN_NODES = 1
MAX_NODES = 1
```
### Deploy `Model` resource to the `Endpoint` resource
Use this helper function `deploy_model` to deploy the `Model` resource to the `Endpoint` resource you created for serving predictions, with the following parameters:
- `model`: The Vertex fully qualified model identifier of the model to upload (deploy) from the training pipeline.
- `deploy_model_display_name`: A human readable name for the deployed model.
- `endpoint`: The Vertex fully qualified endpoint identifier to deploy the model to.
The helper function calls the `Endpoint` client service's method `deploy_model`, which takes the following parameters:
- `endpoint`: The Vertex fully qualified `Endpoint` resource identifier to deploy the `Model` resource to.
- `deployed_model`: The requirements specification for deploying the model.
- `traffic_split`: Percent of traffic at the endpoint that goes to this model, which is specified as a dictionary of one or more key/value pairs.
- If only one model, then specify as **{ "0": 100 }**, where "0" refers to this model being uploaded and 100 means 100% of the traffic.
- If there are existing models on the endpoint, for which the traffic will be split, then use `model_id` to specify as **{ "0": percent, model_id: percent, ... }**, where `model_id` is the model id of an existing model to the deployed endpoint. The percents must add up to 100.
Let's now dive deeper into the `deployed_model` parameter. This parameter is specified as a Python dictionary with the minimum required fields:
- `model`: The Vertex fully qualified model identifier of the (upload) model to deploy.
- `display_name`: A human readable name for the deployed model.
- `disable_container_logging`: This disables logging of container events, such as execution failures (default is container logging is enabled). Container logging is typically enabled when debugging the deployment and then disabled when deployed for production.
- `dedicated_resources`: This refers to how many compute instances (replicas) that are scaled for serving prediction requests.
- `machine_spec`: The compute instance to provision. Use the variable you set earlier `DEPLOY_GPU != None` to use a GPU; otherwise only a CPU is allocated.
- `min_replica_count`: The number of compute instances to initially provision, which you set earlier as the variable `MIN_NODES`.
- `max_replica_count`: The maximum number of compute instances to scale to, which you set earlier as the variable `MAX_NODES`.
#### Traffic Split
Let's now dive deeper into the `traffic_split` parameter. This parameter is specified as a Python dictionary. This might at first be a tad bit confusing. Let me explain, you can deploy more than one instance of your model to an endpoint, and then set how much (percent) goes to each instance.
Why would you do that? Perhaps you already have a previous version deployed in production -- let's call that v1. You got better model evaluation on v2, but you don't know for certain that it is really better until you deploy to production. So in the case of traffic split, you might want to deploy v2 to the same endpoint as v1, but it only get's say 10% of the traffic. That way, you can monitor how well it does without disrupting the majority of users -- until you make a final decision.
#### Response
The method returns a long running operation `response`. We will wait sychronously for the operation to complete by calling the `response.result()`, which will block until the model is deployed. If this is the first time a model is deployed to the endpoint, it may take a few additional minutes to complete provisioning of resources.
```
DEPLOYED_NAME = "cifar10_deployed-" + TIMESTAMP
def deploy_model(
model, deployed_model_display_name, endpoint, traffic_split={"0": 100}
):
if DEPLOY_GPU:
machine_spec = {
"machine_type": DEPLOY_COMPUTE,
"accelerator_type": DEPLOY_GPU,
"accelerator_count": DEPLOY_NGPU,
}
else:
machine_spec = {
"machine_type": DEPLOY_COMPUTE,
"accelerator_count": 0,
}
deployed_model = {
"model": model,
"display_name": deployed_model_display_name,
"dedicated_resources": {
"min_replica_count": MIN_NODES,
"max_replica_count": MAX_NODES,
"machine_spec": machine_spec,
},
"disable_container_logging": False,
}
response = clients["endpoint"].deploy_model(
endpoint=endpoint, deployed_model=deployed_model, traffic_split=traffic_split
)
print("Long running operation:", response.operation.name)
result = response.result()
print("result")
deployed_model = result.deployed_model
print(" deployed_model")
print(" id:", deployed_model.id)
print(" model:", deployed_model.model)
print(" display_name:", deployed_model.display_name)
print(" create_time:", deployed_model.create_time)
return deployed_model.id
deployed_model_id = deploy_model(model_to_deploy_id, DEPLOYED_NAME, endpoint_id)
```
## Make a online prediction request
Now do a online prediction to your deployed model.
### Get test item
You will use an example out of the test (holdout) portion of the dataset as a test item.
```
test_image = x_test[0]
test_label = y_test[0]
print(test_image.shape)
```
### Prepare the request content
You are going to send the CIFAR10 image as compressed JPG image, instead of the raw uncompressed bytes:
- `cv2.imwrite`: Use openCV to write the uncompressed image to disk as a compressed JPEG image.
- Denormalize the image data from \[0,1) range back to [0,255).
- Convert the 32-bit floating point values to 8-bit unsigned integers.
- `tf.io.read_file`: Read the compressed JPG images back into memory as raw bytes.
- `base64.b64encode`: Encode the raw bytes into a base 64 encoded string.
```
import base64
import cv2
cv2.imwrite("tmp.jpg", (test_image * 255).astype(np.uint8))
bytes = tf.io.read_file("tmp.jpg")
b64str = base64.b64encode(bytes.numpy()).decode("utf-8")
```
### Send the prediction request
Ok, now you have a test image. Use this helper function `predict_image`, which takes the following parameters:
- `image`: The test image data as a numpy array.
- `endpoint`: The Vertex fully qualified identifier for the `Endpoint` resource where the `Model` resource was deployed to.
- `parameters_dict`: Additional parameters for serving.
This function calls the prediction client service `predict` method with the following parameters:
- `endpoint`: The Vertex fully qualified identifier for the `Endpoint` resource where the `Model` resource was deployed to.
- `instances`: A list of instances (encoded images) to predict.
- `parameters`: Additional parameters for serving.
To pass the image data to the prediction service, in the previous step you encoded the bytes into base64 -- which makes the content safe from modification when transmitting binary data over the network. You need to tell the serving binary where your model is deployed to, that the content has been base64 encoded, so it will decode it on the other end in the serving binary.
Each instance in the prediction request is a dictionary entry of the form:
{serving_input: {'b64': content}}
- `input_name`: the name of the input layer of the underlying model.
- `'b64'`: A key that indicates the content is base64 encoded.
- `content`: The compressed JPG image bytes as a base64 encoded string.
Since the `predict()` service can take multiple images (instances), you will send your single image as a list of one image. As a final step, you package the instances list into Google's protobuf format -- which is what we pass to the `predict()` service.
The `response` object returns a list, where each element in the list corresponds to the corresponding image in the request. You will see in the output for each prediction:
- `predictions`: Confidence level for the prediction, between 0 and 1, for each of the classes.
```
def predict_image(image, endpoint, parameters_dict):
# The format of each instance should conform to the deployed model's prediction input schema.
instances_list = [{serving_input: {"b64": image}}]
instances = [json_format.ParseDict(s, Value()) for s in instances_list]
response = clients["prediction"].predict(
endpoint=endpoint, instances=instances, parameters=parameters_dict
)
print("response")
print(" deployed_model_id:", response.deployed_model_id)
predictions = response.predictions
print("predictions")
for prediction in predictions:
print(" prediction:", prediction)
predict_image(b64str, endpoint_id, None)
```
## Undeploy the `Model` resource
Now undeploy your `Model` resource from the serving `Endpoint` resoure. Use this helper function `undeploy_model`, which takes the following parameters:
- `deployed_model_id`: The model deployment identifier returned by the endpoint service when the `Model` resource was deployed to.
- `endpoint`: The Vertex fully qualified identifier for the `Endpoint` resource where the `Model` is deployed to.
This function calls the endpoint client service's method `undeploy_model`, with the following parameters:
- `deployed_model_id`: The model deployment identifier returned by the endpoint service when the `Model` resource was deployed.
- `endpoint`: The Vertex fully qualified identifier for the `Endpoint` resource where the `Model` resource is deployed.
- `traffic_split`: How to split traffic among the remaining deployed models on the `Endpoint` resource.
Since this is the only deployed model on the `Endpoint` resource, you simply can leave `traffic_split` empty by setting it to {}.
```
def undeploy_model(deployed_model_id, endpoint):
response = clients["endpoint"].undeploy_model(
endpoint=endpoint, deployed_model_id=deployed_model_id, traffic_split={}
)
print(response)
undeploy_model(deployed_model_id, endpoint_id)
```
# Cleaning up
To clean up all GCP resources used in this project, you can [delete the GCP
project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial.
Otherwise, you can delete the individual resources you created in this tutorial:
- Dataset
- Pipeline
- Model
- Endpoint
- Batch Job
- Custom Job
- Hyperparameter Tuning Job
- Cloud Storage Bucket
```
delete_dataset = True
delete_pipeline = True
delete_model = True
delete_endpoint = True
delete_batchjob = True
delete_customjob = True
delete_hptjob = True
delete_bucket = True
# Delete the dataset using the Vertex fully qualified identifier for the dataset
try:
if delete_dataset and "dataset_id" in globals():
clients["dataset"].delete_dataset(name=dataset_id)
except Exception as e:
print(e)
# Delete the training pipeline using the Vertex fully qualified identifier for the pipeline
try:
if delete_pipeline and "pipeline_id" in globals():
clients["pipeline"].delete_training_pipeline(name=pipeline_id)
except Exception as e:
print(e)
# Delete the model using the Vertex fully qualified identifier for the model
try:
if delete_model and "model_to_deploy_id" in globals():
clients["model"].delete_model(name=model_to_deploy_id)
except Exception as e:
print(e)
# Delete the endpoint using the Vertex fully qualified identifier for the endpoint
try:
if delete_endpoint and "endpoint_id" in globals():
clients["endpoint"].delete_endpoint(name=endpoint_id)
except Exception as e:
print(e)
# Delete the batch job using the Vertex fully qualified identifier for the batch job
try:
if delete_batchjob and "batch_job_id" in globals():
clients["job"].delete_batch_prediction_job(name=batch_job_id)
except Exception as e:
print(e)
# Delete the custom job using the Vertex fully qualified identifier for the custom job
try:
if delete_customjob and "job_id" in globals():
clients["job"].delete_custom_job(name=job_id)
except Exception as e:
print(e)
# Delete the hyperparameter tuning job using the Vertex fully qualified identifier for the hyperparameter tuning job
try:
if delete_hptjob and "hpt_job_id" in globals():
clients["job"].delete_hyperparameter_tuning_job(name=hpt_job_id)
except Exception as e:
print(e)
if delete_bucket and "BUCKET_NAME" in globals():
! gsutil rm -r $BUCKET_NAME
```
| github_jupyter |
```
import platform
print(platform.architecture())
!python --version
pwd
# 네이버에서 검색어 입력받아 검색 한 후 블로그 메뉴를 선택하고
# 오른쪽에 있는 검색옵션 버튼을 눌러서
# 정렬 방식과 기간을 입력하기
#Step 0. 필요한 모듈과 라이브러리를 로딩하고 검색어를 입력 받습니다.
import sys
import os
import pandas as pd
import numpy as np
import math
from bs4 import BeautifulSoup
import requests
import urllib.request as req
from selenium import webdriver
from selenium.webdriver.common.keys import Keys
import time
import tqdm
from tqdm.notebook import tqdm
query_txt = '성심당여행대전'
start_date= "20190101"
end_date= "20210501"
os.getenv('HOME')
webdriver.__version__
#Step 1. 크롬 웹브라우저 실행
path = os.getenv('HOME')+ '/chromedriver'
driver = webdriver.Chrome(path)
# 사이트 주소는 네이버
driver.get('https://blog.naver.com/')
time.sleep(1)
#Step 2. 네이버 검색창에 "검색어" 검색
element = driver.find_element_by_name("sectionBlogQuery").send_keys(query_txt)
# element.submit()
time.sleep(2)
driver.find_element_by_class_name("button.button_blog").click() # 관련도순 xpath
# element.find_element_by_css_selector("#header > div.header_common > div > div.area_search > form > fieldset > a.button.button_blog").click() # 관련도순 xpath
# element.clear()
# element.send_keys(query_txt) # query_txt는 위에서 입력한 '이재용'
# element.submit()
#Step 1. 크롬 웹브라우저 실행
path = os.getenv('HOME')+ '/chromedriver'
driver = webdriver.Chrome(path)
# 사이트 주소는 네이버
driver.get('https://blog.naver.com/')
time.sleep(0.1)
#Step 2. 네이버 검색창에 "검색어" 검색
element = driver.find_element_by_name("sectionBlogQuery").send_keys(query_txt)
# element.submit()
time.sleep(0.1)
driver.find_element_by_class_name("button.button_blog").click() # 관련도순 xpath
time.sleep(0.1)
#Step 4. 오른쪽의 검색 옵션 버튼 클릭
driver.find_element_by_class_name("present_selected").click()
#Step 6. 날짜 입력
driver.find_element_by_id("search_start_date").send_keys(start_date)
driver.find_element_by_id("search_end_date").send_keys(end_date)
time.sleep(0.1)
driver.find_element_by_id("periodSearch").click()
time.sleep(0.1)
searched_post_num = driver.find_element_by_class_name('search_number').text
print(searched_post_num)
url_list = []
title_list = []
total_page = 600
# total_page = math.ceil(int(searched_post_num.replace(',', '').strip('건')) / 7)
print('total_page :', total_page)
for i in tqdm(range(0, total_page)): # 페이지 번호
url = f'https://section.blog.naver.com/Search/Post.naver?pageNo={i}&rangeType=sim&orderBy=recentdate&startDate={start_date}&endDate={end_date}&keyword={query_txt}'
driver.get(url)
response = requests.get(url)
soup = BeautifulSoup(response.text, 'html.parser')
print(soup)
# time.sleep(0.5)
# area = soup.findAll('div', {'class' : 'list_search_post'}) #.find_all('a', {'class' : 'url'})
# print(area)
# URL 크롤링 시작
# titles = "a.sh_blog_title._sp_each_url._sp_each_title" # #content
# # article_raw = driver.find_elements_by_class_name('area_list_search')
# # article_raw = driver.find_elements_by_css_selector('#content > section > div.area_list_search > div:nth-child(1)')
# article_raw = driver.find_elements_by_xpath(f'//*[@id="content"]/section/div[2]/div[{i}]')
# print(article_raw)
# # url 크롤링 시작 # 7개
# for article in article_raw:
# url = article.get_attribute('href')
# print(url)
# url_list.append(url)
# # 제목 크롤링 시작
# for article in article_raw:
# title = article.get_attribute('title')
# title_list.append(title)
# print(title)
# print('url갯수: ', len(url_list))
# print('url갯수: ', len(title_list))
# df = pd.DataFrame({'url':url_list, 'title':title_list})
# # 저장하기
# df.to_csv("./blog_url.csv")
li = [2, 3, 4, 4, 5, 6, 7, 8]
len(li)
for i in range(0, 8, 2):
print(i)
new = []
for i in range(0, len(li)-1, 2):
new.append([li[i], li[i+1]])
new
article_raw = driver.find_elements_by_xpath('//*[@id="content"]/section/div[2]/div[1]')
# article_raw.get_attribute('href')
for i in article_raw:
print(i.get_attribute('href'))
//*[@id="content"]/section/div[2]
//*[@id="content"]/section/div[2]
//*[@id="content"]/section/div[2]
//*[@id="content"]/section/div[2]/div[1]
//*[@id="content"]/section/div[2]/div[2]
//*[@id="content"]/section/div[2]/div[3]
...
//*[@id="content"]/section/div[2]/div[7]
```
1 page = 7 posts
72 page search
sample = https://section.blog.naver.com/Search/Post.naver?pageNo=1&rangeType=PERIOD&orderBy=sim&startDate=2019-01-01&endDate=2021-05-01&keyword=%EC%84%B1%EC%8B%AC%EB%8B%B9%EC%97%AC%ED%96%89%EB%8C%80%EC%A0%84
```
## 제목 눌러서 블로그 페이지 열기
driver.find_element_by_class_name('title').click()
time.sleep(1)
type(searched_post_num), searched_post_num
import re
re.sub('^[0-9]', '', searched_post_num)
searched_post_num
searched_post_num.replace(',', '').replace('건', '')
total_page = math.ceil(int(searched_post_num.replace(',', '').strip('건')) / 7)
total_page
```
| github_jupyter |
```
#Importing packages
import time
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
from tensorflow.keras.layers import Dense
from tensorflow.keras.datasets import mnist
from tensorflow.keras.models import Sequential
from tensorflow.keras.utils import to_categorical
#MNIST Dataset has 70000 instances of hand written digits from 0-9, each image size is 28*28
(x_train, y_train), (x_test, y_test) = mnist.load_data()
#Get the data ready for training
#Convert to Float
x_train, x_test = np.array(x_train, np.float32), np.array(x_test, np.float32)
#Flatten images to 1-D vector of 784 features (28*28).
x_train, x_test = x_train.reshape([-1, 784]), x_test.reshape([-1, 784])
#One hot encoding of labels
y_train = to_categorical(y_train, 10)
y_test = to_categorical(y_test, 10)
#Normalize images value from [0, 255] to [0, 1].
x_train = x_train / 255.
x_test = x_test / 255.
def build_model(activation, initializer):
model = Sequential()
model.add(Dense(512, input_dim = 784, activation = activation, kernel_initializer = initializer))
model.add(Dense(256, activation = activation, kernel_initializer = initializer))
model.add(Dense(128, activation = activation, kernel_initializer = initializer))
model.add(Dense(64, activation = activation, kernel_initializer = initializer))
model.add(Dense(10, activation = 'softmax', kernel_initializer = initializer))
model.compile(loss = 'binary_crossentropy', optimizer = tf.keras.optimizers.Adam(), metrics = ['acc'])
return model
random_normal_model = build_model(activation = 'relu', initializer = tf.keras.initializers.RandomNormal(mean = 0.0, stddev = 0.01))
random_normal_hist = random_normal_model.fit(x_train, y_train, epochs = 15, validation_data = (x_test, y_test), batch_size = 128)
random_normal_model1 = build_model(activation = 'relu', initializer = tf.keras.initializers.RandomNormal(mean = 0.0, stddev = 0.1))
random_normal_hist1 = random_normal_model1.fit(x_train, y_train, epochs = 15, validation_data = (x_test, y_test), batch_size = 128)
random_normal_model2 = build_model(activation = 'relu', initializer = tf.keras.initializers.RandomNormal(mean = 0.0, stddev = 1.0))
random_normal_hist2 = random_normal_model2.fit(x_train, y_train, epochs = 15, validation_data = (x_test, y_test), batch_size = 128)
random_uniform_model = build_model(activation = 'relu', initializer = tf.keras.initializers.RandomUniform(minval = -1, maxval = 1))
random_uniform_hist = random_uniform_model.fit(x_train, y_train, epochs = 15, validation_data = (x_test, y_test), batch_size = 128)
random_uniform_model1 = build_model(activation = 'relu', initializer = tf.keras.initializers.RandomUniform(minval = 0, maxval = 1))
random_uniform_hist1 = random_uniform_model1.fit(x_train, y_train, epochs = 15, validation_data = (x_test, y_test), batch_size = 128)
ones_model = build_model(activation = 'relu', initializer = tf.keras.initializers.Ones())
ones_hist = ones_model.fit(x_train, y_train, epochs = 15, validation_data = (x_test, y_test), batch_size = 128)
zeros_model = build_model(activation = 'relu', initializer = tf.keras.initializers.Zeros())
zeros_hist = zeros_model.fit(x_train, y_train, epochs = 15, validation_data = (x_test, y_test), batch_size = 128)
glorot_normal_model = build_model(activation = 'relu', initializer = tf.keras.initializers.GlorotNormal())
glorot_normal_hist = model.fit(x_train, y_train, epochs = 15, validation_data = (x_test, y_test), batch_size = 128)
glorot_uniform_model1 = build_model(activation = 'relu', initializer = tf.keras.initializers.GlorotUniform())
glorot_uniform_hist1 = model.fit(x_train, y_train, epochs = 15, validation_data = (x_test, y_test), batch_size = 128)
he_normal_model = build_model(activation = 'relu', initializer = tf.keras.initializers.HeNormal())
he_normal_hist = model.fit(x_train, y_train, epochs = 15, validation_data = (x_test, y_test), batch_size = 128)
he_uniform_model = build_model(activation = 'relu', initializer = tf.keras.initializers.HeUniform())
he_uniform_hist = model.fit(x_train, y_train, epochs = 15, validation_data = (x_test, y_test), batch_size = 128)
lecun_uniform_model = build_model(activation = 'relu', initializer = tf.keras.initializers.LecunUniform())
lecun_uniform_hist = model.fit(x_train, y_train, epochs = 15, validation_data = (x_test, y_test), batch_size = 128)
lecun_normal_model = build_model(activation = 'relu', initializer = tf.keras.initializers.LecunNormal())
lecun_normal_hist = model.fit(x_train, y_train, epochs = 15, validation_data = (x_test, y_test), batch_size = 128)
fig, ax = plt.subplots(5, 2, figsize = (12, 28))
ax[0, 0].plot(random_normal_hist.history['acc'])
ax[0, 0].plot(random_normal_hist.history['val_acc'])
ax[0, 0].set_ylabel('Accuracy')
ax[0, 0].set_xlabel('Epochs')
ax[0, 0].legend(['Train', 'Test'], loc='upper left')
ax[0, 0].set_title('Random Normal - StdDev of 0.05')
ax[0, 1].plot(random_normal_hist.history['loss'])
ax[0, 1].plot(random_normal_hist.history['val_loss'])
ax[0, 1].set_ylabel('Loss')
ax[0, 1].set_xlabel('Epoch')
ax[0, 1].legend(['Train', 'Test'], loc='upper left')
ax[0, 1].set_title('Random Normal - StdDev of 0.05')
ax[1, 0].plot(random_normal_hist1.history['acc'])
ax[1, 0].plot(random_normal_hist1.history['val_acc'])
ax[1, 0].set_ylabel('Accuracy')
ax[1, 0].set_xlabel('Epochs')
ax[1, 0].legend(['Train', 'Test'], loc='upper left')
ax[1, 0].set_title('Random Normal - StdDev of 0.1')
ax[1, 1].plot(random_normal_hist1.history['loss'])
ax[1, 1].plot(random_normal_hist1.history['val_loss'])
ax[1, 1].set_ylabel('Loss')
ax[1, 1].set_xlabel('Epoch')
ax[1, 1].legend(['Train', 'Test'], loc='upper left')
ax[1, 1].set_title('Random Normal - StdDev of 0.1')
ax[2, 0].plot(random_normal_hist2.history['acc'])
ax[2, 0].plot(random_normal_hist2.history['val_acc'])
ax[2, 0].set_ylabel('Accuracy')
ax[2, 0].set_xlabel('Epochs')
ax[2, 0].legend(['Train', 'Test'], loc='upper left')
ax[2, 0].set_title('Random Normal - StdDev of 1.0')
ax[2, 1].plot(random_normal_hist2.history['loss'])
ax[2, 1].plot(random_normal_hist2.history['val_loss'])
ax[2, 1].set_ylabel('Loss')
ax[2, 1].set_xlabel('Epoch')
ax[2, 1].legend(['Train', 'Test'], loc='upper left')
ax[2, 1].set_title('Random Normal - StdDev of 1.0')
ax[3, 0].plot(random_uniform_hist.history['acc'])
ax[3, 0].plot(random_uniform_hist.history['val_acc'])
ax[3, 0].set_ylabel('Accuracy')
ax[3, 0].set_xlabel('Epochs')
ax[3, 0].legend(['Train', 'Test'], loc='upper left')
ax[3, 0].set_title('Random Uniform - [-1, 1]')
ax[3, 1].plot(random_uniform_hist.history['loss'])
ax[3, 1].plot(random_uniform_hist.history['val_loss'])
ax[3, 1].set_ylabel('Loss')
ax[3, 1].set_xlabel('Epoch')
ax[3, 1].legend(['Train', 'Test'], loc='upper left')
ax[3, 1].set_title('Random Uniform - [-1, 1]')
ax[4, 0].plot(random_uniform_hist1.history['acc'])
ax[4, 0].plot(random_uniform_hist1.history['val_acc'])
ax[4, 0].set_ylabel('Accuracy')
ax[4, 0].set_xlabel('Epochs')
ax[4, 0].legend(['Train', 'Test'], loc='upper left')
ax[4, 0].set_title('Random Uniform - [0, 1]')
ax[4, 1].plot(random_uniform_hist1.history['loss'])
ax[4, 1].plot(random_uniform_hist1.history['val_loss'])
ax[4, 1].set_ylabel('Loss')
ax[4, 1].set_xlabel('Epoch')
ax[4, 1].legend(['Train', 'Test'], loc='upper left')
ax[4, 1].set_title('Random Uniform - [0, 1]')
plt.show()
fig, ax = plt.subplots(2, 2, figsize = (12, 10))
ax[0, 0].plot(ones_hist.history['acc'])
ax[0, 0].plot(ones_hist.history['val_acc'])
ax[0, 0].set_ylabel('Accuracy')
ax[0, 0].set_xlabel('Epochs')
ax[0, 0].legend(['Train', 'Test'], loc='upper left')
ax[0, 0].set_title('Ones')
ax[0, 1].plot(ones_hist.history['loss'])
ax[0, 1].plot(ones_hist.history['val_loss'])
ax[0, 1].set_ylabel('Loss')
ax[0, 1].set_xlabel('Epoch')
ax[0, 1].legend(['Train', 'Test'], loc='upper left')
ax[0, 1].set_title('Ones')
ax[1, 0].plot(zeros_hist.history['acc'])
ax[1, 0].plot(zeros_hist.history['val_acc'])
ax[1, 0].set_ylabel('Accuracy')
ax[1, 0].set_xlabel('Epochs')
ax[1, 0].legend(['Train', 'Test'], loc='upper left')
ax[1, 0].set_title('Zeros')
ax[1, 1].plot(zeros_hist.history['loss'])
ax[1, 1].plot(zeros_hist.history['val_loss'])
ax[1, 1].set_ylabel('Loss')
ax[1, 1].set_xlabel('Epoch')
ax[1, 1].legend(['Train', 'Test'], loc='upper left')
ax[1, 1].set_title('Zeros')
plt.show()
```
## Given below is a method to manually initialize and have complete control (If necessary and for custom weight init as required)
```
#Iterate over the layers of a given model
for layer in model.layers:
if isinstance(layer, tf.keras.layers.Dense):
#Initialize weights in the range of [-y, y], where y = 1/√n (n is no. of neurons in the next layer)
shape = (layer.weights[0].shape[0], layer.weights[0].shape[1])
y = 1.0/np.sqrt(shape[0])
rule_weights = np.random.uniform(-y, y, shape)
#Weights
layer.weights[0] = rule_weights
#Bias
layer.weights[1] = 0
```
## You can visualize layer weight init distribution in a graphical way also, using Tensorboard - Head over to the notebook [Tensorboard Demo]()
## In the above given weight init techniques, feel free to change and run the respective init techniques, make slight changes to the matplotlib cell to accomodate your interested weight init graph for visualization
## You can combine various weight init in a single graph also, for a visualization and comparision
| github_jupyter |
<a href="https://colab.research.google.com/github/yukinaga/lecture_pytorch/blob/master/lecture2/exercise.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# 演習
第2講の演習です。
PyTorchを使ってモデルを構築し、最適化アルゴリズムを設定しましょう。
## データを訓練用とテスト用に分割
```
import torch
from sklearn import datasets
from sklearn.model_selection import train_test_split
digits_data = datasets.load_digits()
digit_images = digits_data.data
labels = digits_data.target
x_train, x_test, t_train, t_test = train_test_split(digit_images, labels) # 25%がテスト用
# Tensorに変換
x_train = torch.tensor(x_train, dtype=torch.float32)
t_train = torch.tensor(t_train, dtype=torch.int64)
x_test = torch.tensor(x_test, dtype=torch.float32)
t_test = torch.tensor(t_test, dtype=torch.int64)
```
## モデルの構築
`nn`モジュールの`Sequential`クラスを使い、`print(net)`で以下のように表示されるモデルを構築しましょう。
```
Sequential(
(0): Linear(in_features=64, out_features=128, bias=True)
(1): ReLU()
(2): Linear(in_features=128, out_features=64, bias=True)
(3): ReLU()
(4): Linear(in_features=64, out_features=10, bias=True)
)
```
```
from torch import nn
net = nn.Sequential(
# ------- ここからコードを記述 -------
# ------- ここまで -------
)
print(net)
```
## 学習
モデルを訓練します。
最適化アルゴリズムの設定をしましょう。
最適化アルゴリズムは、以下のページから好きなものを選択してください。
https://pytorch.org/docs/stable/optim.html
```
from torch import optim
# 交差エントロピー誤差関数
loss_fnc = nn.CrossEntropyLoss()
# 最適化アルゴリズム
optimizer = # ここにコードを記述
# 損失のログ
record_loss_train = []
record_loss_test = []
# 1000エポック学習
for i in range(1000):
# 勾配を0に
optimizer.zero_grad()
# 順伝播
y_train = net(x_train)
y_test = net(x_test)
# 誤差を求める
loss_train = loss_fnc(y_train, t_train)
loss_test = loss_fnc(y_test, t_test)
record_loss_train.append(loss_train.item())
record_loss_test.append(loss_test.item())
# 逆伝播(勾配を求める)
loss_train.backward()
# パラメータの更新
optimizer.step()
if i%100 == 0:
print("Epoch:", i, "Loss_Train:", loss_train.item(), "Loss_Test:", loss_test.item())
```
## 誤差の推移
```
import matplotlib.pyplot as plt
plt.plot(range(len(record_loss_train)), record_loss_train, label="Train")
plt.plot(range(len(record_loss_test)), record_loss_test, label="Test")
plt.legend()
plt.xlabel("Epochs")
plt.ylabel("Error")
plt.show()
```
## 正解率
```
y_test = net(x_test)
count = (y_test.argmax(1) == t_test).sum().item()
print("正解率:", str(count/len(y_test)*100) + "%")
```
# 解答例
以下は、どうしても手がかりがないときのみ参考にしましょう。
```
from torch import nn
net = nn.Sequential(
# ------- ここからコードを記述 -------
nn.Linear(64, 128), # 全結合層
nn.ReLU(), # ReLU
nn.Linear(128, 64),
nn.ReLU(),
nn.Linear(64, 10)
# ------- ここまで -------
)
print(net)
from torch import optim
# 交差エントロピー誤差関数
loss_fnc = nn.CrossEntropyLoss()
# 最適化アルゴリズム
optimizer = optim.Adam(net.parameters()) # ここにコードを記述
# 損失のログ
record_loss_train = []
record_loss_test = []
# 1000エポック学習
for i in range(1000):
# 勾配を0に
optimizer.zero_grad()
# 順伝播
y_train = net(x_train)
y_test = net(x_test)
# 誤差を求める
loss_train = loss_fnc(y_train, t_train)
loss_test = loss_fnc(y_test, t_test)
record_loss_train.append(loss_train.item())
record_loss_test.append(loss_test.item())
# 逆伝播(勾配を求める)
loss_train.backward()
# パラメータの更新
optimizer.step()
if i%100 == 0:
print("Epoch:", i, "Loss_Train:", loss_train.item(), "Loss_Test:", loss_test.item())
```
| github_jupyter |
```
import numpy as np, requests, pandas as pd, zipfile, StringIO,json
df15=pd.read_csv('https://www.worldenergy.org/csv/index-table/2015').set_index('Country name')
df14=pd.read_csv('https://www.worldenergy.org/csv/index-table/2014').set_index('Country name')
df13=pd.read_csv('https://www.worldenergy.org/csv/index-table/2013').set_index('Country name')
df12=pd.read_csv('https://www.worldenergy.org/csv/index-table/2012').set_index('Country name')
df11=pd.read_csv('https://www.worldenergy.org/csv/index-table/2011').set_index('Country name')
df10=pd.read_csv('https://www.worldenergy.org/csv/index-table/2010').set_index('Country name')
df10.columns
df15.loc['Luxembourg']
#hdi:e3a
#edu:energy equity
#gni:energy affordability
#le:env susti
#idhi:contextual performance
#edu:societal strength
#gni:economic strength
#le:political strength
pop=pd.read_csv('pop.csv').set_index(['Country','Year'])
#country name converter
def cnc(c):
if c==u'Democratic Republic of the Congo':return 'Congo (Dem. Rep.)'
elif c==u'Ivory Coast':return "C\xc3\xb4te d'Ivoire"
return c
def cnc2(c):
if c==u'Ivory Coast':return "C\xc3\xb4te d'Ivoire"
elif c=="Syria":return 'Syrian Arab Republic'
elif c=="Moldova":return 'Republic of Moldova'
elif c=="South Korea":return 'Republic of Korea'
elif c=="Hong Kong":return 'China, Hong Kong Special Administrative Region'
elif c=="Vietnam":return 'Viet Nam'
elif c=="Tanzania":return 'United Republic of Tanzania'
elif c=="Iran":return 'Iran (Islamic Republic of)'
elif c=="United States":return 'United States of America'
elif c=="Russia":return 'Russian Federation'
elif c=="Libya":return 'Libyan Arab Jamahiriya'
elif c=="Venezuela":return 'Venezuela (Bolivarian Republic of)'
elif c=="Yemen":return 'Yemen '
elif c=="Macedonia":return 'The former Yugoslav Republic of Macedonia'
return c
#load dictionary preformatted for viz type
data3=json.loads(file('data3.json','r').read())
data={}
for d in data3:
if d not in data: data[d]={}
if 'e3' not in data[d]:data[d]['e3']={}
if 'cp' not in data[d]:data[d]['cp']={}
if 'name' not in data[d]:data[d]['name']=data3[d]['name']
if 'code' not in data[d]:data[d]['code']=data3[d]['code']
if 'population' not in data[d]:data[d]['population']={}
for y in range(2010,2017):
if repr(y) not in data[d]['e3']:data[d]['e3'][repr(y)]={u'edu': u'NaN', u'gni': u'NaN', u'le': u'NaN'}
if repr(y) not in data[d]['cp']:data[d]['cp'][repr(y)]={u'edu': u'NaN', u'gni': u'NaN', u'le': u'NaN'}
try:
data[d]['population'][repr(y)]=str(pop.loc[cnc2(data3[d]['name'])].loc[y]['Value']*1000)
#print "success population",y
except: pass
try:
data[d]['e3']['2010']['gni']=str(np.round(df10.loc[cnc(data3[d]['name'])]['Social equity score 2010']/10.0,3))
data[d]['e3']['2010']['edu']=str(np.round(df10.loc[cnc(data3[d]['name'])]['Energy security score 2010']/10.0,3))
data[d]['e3']['2010']['le']=str(np.round(df10.loc[cnc(data3[d]['name'])]['Environmental impact mitigation score 2010']/10.0,3))
data[d]['cp']['2010']['edu']=str(np.round(df10.loc[cnc(data3[d]['name'])]['Societal strength score 2010']/10.0,3))
data[d]['cp']['2010']['gni']=str(np.round(df10.loc[cnc(data3[d]['name'])]['Economic strength score 2010']/10.0,3))
data[d]['cp']['2010']['le']=str(np.round(df10.loc[cnc(data3[d]['name'])]['Political strength score 2010']/10.0,3))
#print "success 2010"
except:pass
try:
data[d]['e3']['2011']['gni']=str(np.round(df11.loc[cnc(data3[d]['name'])]['Social equity score 2011']/10.0,3))
data[d]['e3']['2011']['edu']=str(np.round(df11.loc[cnc(data3[d]['name'])]['Energy security score 2011']/10.0,3))
data[d]['e3']['2011']['le']=str(np.round(df11.loc[cnc(data3[d]['name'])]['Environmental impact mitigation score 2011']/10.0,3))
data[d]['cp']['2011']['edu']=str(np.round(df11.loc[cnc(data3[d]['name'])]['Societal strength score 2011']/10.0,3))
data[d]['cp']['2011']['gni']=str(np.round(df11.loc[cnc(data3[d]['name'])]['Economic strength score 2011']/10.0,3))
data[d]['cp']['2011']['le']=str(np.round(df11.loc[cnc(data3[d]['name'])]['Political strength score 2011']/10.0,3))
#print "success 2011"
except:pass
try:
data[d]['e3']['2012']['gni']=str(np.round(df12.loc[cnc(data3[d]['name'])]['Social equity score 2012']/10.0,3))
data[d]['e3']['2012']['edu']=str(np.round(df12.loc[cnc(data3[d]['name'])]['Energy security score 2012']/10.0,3))
data[d]['e3']['2012']['le']=str(np.round(df12.loc[cnc(data3[d]['name'])]['Environmental impact mitigation score 2012']/10.0,3))
data[d]['cp']['2012']['edu']=str(np.round(df12.loc[cnc(data3[d]['name'])]['Societal strength score 2012']/10.0,3))
data[d]['cp']['2012']['gni']=str(np.round(df12.loc[cnc(data3[d]['name'])]['Economic strength score 2012']/10.0,3))
data[d]['cp']['2012']['le']=str(np.round(df12.loc[cnc(data3[d]['name'])]['Political strength score 2012']/10.0,3))
#print "success 2012"
except:pass
try:
data[d]['e3']['2013']['gni']=str(np.round(df13.loc[cnc(data3[d]['name'])]['Social equity score 2013']/10.0,3))
data[d]['e3']['2013']['edu']=str(np.round(df13.loc[cnc(data3[d]['name'])]['Energy security score 2013']/10.0,3))
data[d]['e3']['2013']['le']=str(np.round(df13.loc[cnc(data3[d]['name'])]['Environmental impact mitigation score 2013']/10.0,3))
data[d]['cp']['2013']['edu']=str(np.round(df13.loc[cnc(data3[d]['name'])]['Societal strength score 2013']/10.0,3))
data[d]['cp']['2013']['gni']=str(np.round(df13.loc[cnc(data3[d]['name'])]['Economic strength score 2013']/10.0,3))
data[d]['cp']['2013']['le']=str(np.round(df13.loc[cnc(data3[d]['name'])]['Political strength score 2013']/10.0,3))
#print "success 2013"
except:pass
try:
data[d]['e3']['2014']['gni']=str(np.round(df14.loc[cnc(data3[d]['name'])]['Social equity score 2014']/10.0,3))
data[d]['e3']['2014']['edu']=str(np.round(df14.loc[cnc(data3[d]['name'])]['Energy security score 2014']/10.0,3))
data[d]['e3']['2014']['le']=str(np.round(df14.loc[cnc(data3[d]['name'])]['Environmental impact mitigation score 2014']/10.0,3))
data[d]['cp']['2014']['edu']=str(np.round(df14.loc[cnc(data3[d]['name'])]['Societal strength score 2014']/10.0,3))
data[d]['cp']['2014']['gni']=str(np.round(df14.loc[cnc(data3[d]['name'])]['Economic strength score 2014']/10.0,3))
data[d]['cp']['2014']['le']=str(np.round(df14.loc[cnc(data3[d]['name'])]['Political strength score 2014']/10.0,3))
#print "success 2014"
except:pass
try:
data[d]['e3']['2015']['gni']=str(np.round(df15.loc[cnc(data3[d]['name'])]['Social equity score 2015']/10.0,3))
data[d]['e3']['2015']['edu']=str(np.round(df15.loc[cnc(data3[d]['name'])]['Energy security score 2015']/10.0,3))
data[d]['e3']['2015']['le']=str(np.round(df15.loc[cnc(data3[d]['name'])]['Environmental impact mitigation score 2015']/10.0,3))
data[d]['cp']['2015']['edu']=str(np.round(df15.loc[cnc(data3[d]['name'])]['Societal strength score 2015']/10.0,3))
data[d]['cp']['2015']['gni']=str(np.round(df15.loc[cnc(data3[d]['name'])]['Economic strength score 2015']/10.0,3))
data[d]['cp']['2015']['le']=str(np.round(df15.loc[cnc(data3[d]['name'])]['Political strength score 2015']/10.0,3))
#print "success 2015"
except:pass
file('data.json','w').write(json.dumps(data))
```
| github_jupyter |
### <font color = "darkblue">Updates to Assignment</font>
#### If you were working on the older version:
* Please click on the "Coursera" icon in the top right to open up the folder directory.
* Navigate to the folder: Week 3/ Planar data classification with one hidden layer. You can see your prior work in version 6b: "Planar data classification with one hidden layer v6b.ipynb"
#### List of bug fixes and enhancements
* Clarifies that the classifier will learn to classify regions as either red or blue.
* compute_cost function fixes np.squeeze by casting it as a float.
* compute_cost instructions clarify the purpose of np.squeeze.
* compute_cost clarifies that "parameters" parameter is not needed, but is kept in the function definition until the auto-grader is also updated.
* nn_model removes extraction of parameter values, as the entire parameter dictionary is passed to the invoked functions.
# Planar data classification with one hidden layer
Welcome to your week 3 programming assignment. It's time to build your first neural network, which will have a hidden layer. You will see a big difference between this model and the one you implemented using logistic regression.
**You will learn how to:**
- Implement a 2-class classification neural network with a single hidden layer
- Use units with a non-linear activation function, such as tanh
- Compute the cross entropy loss
- Implement forward and backward propagation
## 1 - Packages ##
Let's first import all the packages that you will need during this assignment.
- [numpy](https://www.numpy.org/) is the fundamental package for scientific computing with Python.
- [sklearn](http://scikit-learn.org/stable/) provides simple and efficient tools for data mining and data analysis.
- [matplotlib](http://matplotlib.org) is a library for plotting graphs in Python.
- testCases provides some test examples to assess the correctness of your functions
- planar_utils provide various useful functions used in this assignment
```
# Package imports
import numpy as np
import matplotlib.pyplot as plt
from testCases_v2 import *
import sklearn
import sklearn.datasets
import sklearn.linear_model
from planar_utils import plot_decision_boundary, sigmoid, load_planar_dataset, load_extra_datasets
%matplotlib inline
np.random.seed(1) # set a seed so that the results are consistent
```
## 2 - Dataset ##
First, let's get the dataset you will work on. The following code will load a "flower" 2-class dataset into variables `X` and `Y`.
```
X, Y = load_planar_dataset()
```
Visualize the dataset using matplotlib. The data looks like a "flower" with some red (label y=0) and some blue (y=1) points. Your goal is to build a model to fit this data. In other words, we want the classifier to define regions as either red or blue.
```
# Visualize the data:
plt.scatter(X[0, :], X[1, :], c=Y, s=40, cmap=plt.cm.Spectral);
```
You have:
- a numpy-array (matrix) X that contains your features (x1, x2)
- a numpy-array (vector) Y that contains your labels (red:0, blue:1).
Lets first get a better sense of what our data is like.
**Exercise**: How many training examples do you have? In addition, what is the `shape` of the variables `X` and `Y`?
**Hint**: How do you get the shape of a numpy array? [(help)](https://docs.scipy.org/doc/numpy/reference/generated/numpy.ndarray.shape.html)
```
### START CODE HERE ### (≈ 3 lines of code)
shape_X = X.shape
shape_Y = Y.shape
m = (X.size)/shape_X[0] # training set size
### END CODE HERE ###
print ('The shape of X is: ' + str(shape_X))
print ('The shape of Y is: ' + str(shape_Y))
print ('I have m = %d training examples!' % (m))
```
**Expected Output**:
<table style="width:20%">
<tr>
<td>**shape of X**</td>
<td> (2, 400) </td>
</tr>
<tr>
<td>**shape of Y**</td>
<td>(1, 400) </td>
</tr>
<tr>
<td>**m**</td>
<td> 400 </td>
</tr>
</table>
## 3 - Simple Logistic Regression
Before building a full neural network, lets first see how logistic regression performs on this problem. You can use sklearn's built-in functions to do that. Run the code below to train a logistic regression classifier on the dataset.
```
# Train the logistic regression classifier
clf = sklearn.linear_model.LogisticRegressionCV();
clf.fit(X.T, Y.T);
```
You can now plot the decision boundary of these models. Run the code below.
```
# Plot the decision boundary for logistic regression
plot_decision_boundary(lambda x: clf.predict(x), X, Y)
plt.title("Logistic Regression")
# Print accuracy
LR_predictions = clf.predict(X.T)
print ('Accuracy of logistic regression: %d ' % float((np.dot(Y,LR_predictions) + np.dot(1-Y,1-LR_predictions))/float(Y.size)*100) +
'% ' + "(percentage of correctly labelled datapoints)")
```
**Expected Output**:
<table style="width:20%">
<tr>
<td>**Accuracy**</td>
<td> 47% </td>
</tr>
</table>
**Interpretation**: The dataset is not linearly separable, so logistic regression doesn't perform well. Hopefully a neural network will do better. Let's try this now!
## 4 - Neural Network model
Logistic regression did not work well on the "flower dataset". You are going to train a Neural Network with a single hidden layer.
**Here is our model**:
<img src="images/classification_kiank.png" style="width:600px;height:300px;">
**Mathematically**:
For one example $x^{(i)}$:
$$z^{[1] (i)} = W^{[1]} x^{(i)} + b^{[1]}\tag{1}$$
$$a^{[1] (i)} = \tanh(z^{[1] (i)})\tag{2}$$
$$z^{[2] (i)} = W^{[2]} a^{[1] (i)} + b^{[2]}\tag{3}$$
$$\hat{y}^{(i)} = a^{[2] (i)} = \sigma(z^{ [2] (i)})\tag{4}$$
$$y^{(i)}_{prediction} = \begin{cases} 1 & \mbox{if } a^{[2](i)} > 0.5 \\ 0 & \mbox{otherwise } \end{cases}\tag{5}$$
Given the predictions on all the examples, you can also compute the cost $J$ as follows:
$$J = - \frac{1}{m} \sum\limits_{i = 0}^{m} \large\left(\small y^{(i)}\log\left(a^{[2] (i)}\right) + (1-y^{(i)})\log\left(1- a^{[2] (i)}\right) \large \right) \small \tag{6}$$
**Reminder**: The general methodology to build a Neural Network is to:
1. Define the neural network structure ( # of input units, # of hidden units, etc).
2. Initialize the model's parameters
3. Loop:
- Implement forward propagation
- Compute loss
- Implement backward propagation to get the gradients
- Update parameters (gradient descent)
You often build helper functions to compute steps 1-3 and then merge them into one function we call `nn_model()`. Once you've built `nn_model()` and learnt the right parameters, you can make predictions on new data.
### 4.1 - Defining the neural network structure ####
**Exercise**: Define three variables:
- n_x: the size of the input layer
- n_h: the size of the hidden layer (set this to 4)
- n_y: the size of the output layer
**Hint**: Use shapes of X and Y to find n_x and n_y. Also, hard code the hidden layer size to be 4.
```
# GRADED FUNCTION: layer_sizes
def layer_sizes(X, Y):
"""
Arguments:
X -- input dataset of shape (input size, number of examples)
Y -- labels of shape (output size, number of examples)
Returns:
n_x -- the size of the input layer
n_h -- the size of the hidden layer
n_y -- the size of the output layer
"""
### START CODE HERE ### (≈ 3 lines of code)
n_x = X.shape[0] # size of input layer
n_h = 4
n_y = Y.shape[0] # size of output layer
### END CODE HERE ###
return (n_x, n_h, n_y)
X_assess, Y_assess = layer_sizes_test_case()
(n_x, n_h, n_y) = layer_sizes(X_assess, Y_assess)
print("The size of the input layer is: n_x = " + str(n_x))
print("The size of the hidden layer is: n_h = " + str(n_h))
print("The size of the output layer is: n_y = " + str(n_y))
```
**Expected Output** (these are not the sizes you will use for your network, they are just used to assess the function you've just coded).
<table style="width:20%">
<tr>
<td>**n_x**</td>
<td> 5 </td>
</tr>
<tr>
<td>**n_h**</td>
<td> 4 </td>
</tr>
<tr>
<td>**n_y**</td>
<td> 2 </td>
</tr>
</table>
### 4.2 - Initialize the model's parameters ####
**Exercise**: Implement the function `initialize_parameters()`.
**Instructions**:
- Make sure your parameters' sizes are right. Refer to the neural network figure above if needed.
- You will initialize the weights matrices with random values.
- Use: `np.random.randn(a,b) * 0.01` to randomly initialize a matrix of shape (a,b).
- You will initialize the bias vectors as zeros.
- Use: `np.zeros((a,b))` to initialize a matrix of shape (a,b) with zeros.
```
# GRADED FUNCTION: initialize_parameters
def initialize_parameters(n_x, n_h, n_y):
"""
Argument:
n_x -- size of the input layer
n_h -- size of the hidden layer
n_y -- size of the output layer
Returns:
params -- python dictionary containing your parameters:
W1 -- weight matrix of shape (n_h, n_x)
b1 -- bias vector of shape (n_h, 1)
W2 -- weight matrix of shape (n_y, n_h)
b2 -- bias vector of shape (n_y, 1)
"""
np.random.seed(2) # we set up a seed so that your output matches ours although the initialization is random.
### START CODE HERE ### (≈ 4 lines of code)
W1 = np.random.randn(n_h,n_x) * 0.01
b1 = np.zeros((n_h,1))
W2 = np.random.randn(n_y,n_h) * 0.01
b2 = np.zeros((n_y,1))
### END CODE HERE ###
assert (W1.shape == (n_h, n_x))
assert (b1.shape == (n_h, 1))
assert (W2.shape == (n_y, n_h))
assert (b2.shape == (n_y, 1))
parameters = {"W1": W1,
"b1": b1,
"W2": W2,
"b2": b2}
return parameters
n_x, n_h, n_y = initialize_parameters_test_case()
parameters = initialize_parameters(n_x, n_h, n_y)
print("W1 = " + str(parameters["W1"]))
print("b1 = " + str(parameters["b1"]))
print("W2 = " + str(parameters["W2"]))
print("b2 = " + str(parameters["b2"]))
```
**Expected Output**:
<table style="width:90%">
<tr>
<td>**W1**</td>
<td> [[-0.00416758 -0.00056267]
[-0.02136196 0.01640271]
[-0.01793436 -0.00841747]
[ 0.00502881 -0.01245288]] </td>
</tr>
<tr>
<td>**b1**</td>
<td> [[ 0.]
[ 0.]
[ 0.]
[ 0.]] </td>
</tr>
<tr>
<td>**W2**</td>
<td> [[-0.01057952 -0.00909008 0.00551454 0.02292208]]</td>
</tr>
<tr>
<td>**b2**</td>
<td> [[ 0.]] </td>
</tr>
</table>
### 4.3 - The Loop ####
**Question**: Implement `forward_propagation()`.
**Instructions**:
- Look above at the mathematical representation of your classifier.
- You can use the function `sigmoid()`. It is built-in (imported) in the notebook.
- You can use the function `np.tanh()`. It is part of the numpy library.
- The steps you have to implement are:
1. Retrieve each parameter from the dictionary "parameters" (which is the output of `initialize_parameters()`) by using `parameters[".."]`.
2. Implement Forward Propagation. Compute $Z^{[1]}, A^{[1]}, Z^{[2]}$ and $A^{[2]}$ (the vector of all your predictions on all the examples in the training set).
- Values needed in the backpropagation are stored in "`cache`". The `cache` will be given as an input to the backpropagation function.
```
# GRADED FUNCTION: forward_propagation
def forward_propagation(X, parameters):
"""
Argument:
X -- input data of size (n_x, m)
parameters -- python dictionary containing your parameters (output of initialization function)
Returns:
A2 -- The sigmoid output of the second activation
cache -- a dictionary containing "Z1", "A1", "Z2" and "A2"
"""
# Retrieve each parameter from the dictionary "parameters"
### START CODE HERE ### (≈ 4 lines of code)
W1 = parameters["W1"]
b1 = parameters["b1"]
W2 = parameters["W2"]
b2 = parameters["b2"]
### END CODE HERE ###
# Implement Forward Propagation to calculate A2 (probabilities)
### START CODE HERE ### (≈ 4 lines of code)
Z1 = np.dot(W1,X) + b1
A1 = np.tanh(Z1)
Z2 = np.dot(W2,A1) + b2
A2 = sigmoid(Z2)
### END CODE HERE ###
assert(A2.shape == (1, X.shape[1]))
# Values needed in the backpropagation are stored in "cache". This will be given as an input to the backpropagation
cache = {"Z1": Z1,
"A1": A1,
"Z2": Z2,
"A2": A2}
return A2, cache
X_assess, parameters = forward_propagation_test_case()
A2, cache = forward_propagation(X_assess, parameters)
# Note: we use the mean here just to make sure that your output matches ours.
print(np.mean(cache['Z1']) ,np.mean(cache['A1']),np.mean(cache['Z2']),np.mean(cache['A2']))
```
**Expected Output**:
<table style="width:50%">
<tr>
<td> 0.262818640198 0.091999045227 -1.30766601287 0.212877681719 </td>
</tr>
</table>
Now that you have computed $A^{[2]}$ (in the Python variable "`A2`"), which contains $a^{[2](i)}$ for every example, you can compute the cost function as follows:
$$J = - \frac{1}{m} \sum\limits_{i = 1}^{m} \large{(} \small y^{(i)}\log\left(a^{[2] (i)}\right) + (1-y^{(i)})\log\left(1- a^{[2] (i)}\right) \large{)} \small\tag{13}$$
**Exercise**: Implement `compute_cost()` to compute the value of the cost $J$.
**Instructions**:
- There are many ways to implement the cross-entropy loss. To help you, we give you how we would have implemented
$- \sum\limits_{i=0}^{m} y^{(i)}\log(a^{[2](i)})$:
```python
logprobs = np.multiply(np.log(A2),Y)
cost = - np.sum(logprobs) # no need to use a for loop!
```
(you can use either `np.multiply()` and then `np.sum()` or directly `np.dot()`).
Note that if you use `np.multiply` followed by `np.sum` the end result will be a type `float`, whereas if you use `np.dot`, the result will be a 2D numpy array. We can use `np.squeeze()` to remove redundant dimensions (in the case of single float, this will be reduced to a zero-dimension array). We can cast the array as a type `float` using `float()`.
```
# GRADED FUNCTION: compute_cost
def compute_cost(A2, Y, parameters):
"""
Computes the cross-entropy cost given in equation (13)
Arguments:
A2 -- The sigmoid output of the second activation, of shape (1, number of examples)
Y -- "true" labels vector of shape (1, number of examples)
parameters -- python dictionary containing your parameters W1, b1, W2 and b2
[Note that the parameters argument is not used in this function,
but the auto-grader currently expects this parameter.
Future version of this notebook will fix both the notebook
and the auto-grader so that `parameters` is not needed.
For now, please include `parameters` in the function signature,
and also when invoking this function.]
Returns:
cost -- cross-entropy cost given equation (13)
"""
m = Y.shape[1] # number of example
# Compute the cross-entropy cost
### START CODE HERE ### (≈ 2 lines of code)
logprobs = logprobs = np.multiply(Y ,np.log(A2)) + np.multiply((1-Y), np.log(1-A2))
cost = (-1/m) * np.sum(logprobs)
### END CODE HERE ###
cost = float(np.squeeze(cost)) # makes sure cost is the dimension we expect.
# E.g., turns [[17]] into 17
assert(isinstance(cost, float))
return cost
A2, Y_assess, parameters = compute_cost_test_case()
print("cost = " + str(compute_cost(A2, Y_assess, parameters)))
```
**Expected Output**:
<table style="width:20%">
<tr>
<td>**cost**</td>
<td> 0.693058761... </td>
</tr>
</table>
Using the cache computed during forward propagation, you can now implement backward propagation.
**Question**: Implement the function `backward_propagation()`.
**Instructions**:
Backpropagation is usually the hardest (most mathematical) part in deep learning. To help you, here again is the slide from the lecture on backpropagation. You'll want to use the six equations on the right of this slide, since you are building a vectorized implementation.
<img src="images/grad_summary.png" style="width:600px;height:300px;">
<!--
$\frac{\partial \mathcal{J} }{ \partial z_{2}^{(i)} } = \frac{1}{m} (a^{[2](i)} - y^{(i)})$
$\frac{\partial \mathcal{J} }{ \partial W_2 } = \frac{\partial \mathcal{J} }{ \partial z_{2}^{(i)} } a^{[1] (i) T} $
$\frac{\partial \mathcal{J} }{ \partial b_2 } = \sum_i{\frac{\partial \mathcal{J} }{ \partial z_{2}^{(i)}}}$
$\frac{\partial \mathcal{J} }{ \partial z_{1}^{(i)} } = W_2^T \frac{\partial \mathcal{J} }{ \partial z_{2}^{(i)} } * ( 1 - a^{[1] (i) 2}) $
$\frac{\partial \mathcal{J} }{ \partial W_1 } = \frac{\partial \mathcal{J} }{ \partial z_{1}^{(i)} } X^T $
$\frac{\partial \mathcal{J} _i }{ \partial b_1 } = \sum_i{\frac{\partial \mathcal{J} }{ \partial z_{1}^{(i)}}}$
- Note that $*$ denotes elementwise multiplication.
- The notation you will use is common in deep learning coding:
- dW1 = $\frac{\partial \mathcal{J} }{ \partial W_1 }$
- db1 = $\frac{\partial \mathcal{J} }{ \partial b_1 }$
- dW2 = $\frac{\partial \mathcal{J} }{ \partial W_2 }$
- db2 = $\frac{\partial \mathcal{J} }{ \partial b_2 }$
!-->
- Tips:
- To compute dZ1 you'll need to compute $g^{[1]'}(Z^{[1]})$. Since $g^{[1]}(.)$ is the tanh activation function, if $a = g^{[1]}(z)$ then $g^{[1]'}(z) = 1-a^2$. So you can compute
$g^{[1]'}(Z^{[1]})$ using `(1 - np.power(A1, 2))`.
```
# GRADED FUNCTION: backward_propagation
def backward_propagation(parameters, cache, X, Y):
"""
Implement the backward propagation using the instructions above.
Arguments:
parameters -- python dictionary containing our parameters
cache -- a dictionary containing "Z1", "A1", "Z2" and "A2".
X -- input data of shape (2, number of examples)
Y -- "true" labels vector of shape (1, number of examples)
Returns:
grads -- python dictionary containing your gradients with respect to different parameters
"""
m = X.shape[1]
# First, retrieve W1 and W2 from the dictionary "parameters".
### START CODE HERE ### (≈ 2 lines of code)
W1 = parameters["W1"]
b1 = parameters["b1"]
W2 = parameters["W2"]
b2 = parameters["b2"]
### END CODE HERE ###
# Retrieve also A1 and A2 from dictionary "cache".
### START CODE HERE ### (≈ 2 lines of code)
A1 = cache["A1"]
A2 = cache["A2"]
Z1 = cache["Z1"]
Z2 = cache["Z2"]
### END CODE HERE ###
# Backward propagation: calculate dW1, db1, dW2, db2.
### START CODE HERE ### (≈ 6 lines of code, corresponding to 6 equations on slide above)
dZ2 = A2 - Y
dW2 = (1/m) * np.dot(dZ2,A1.T)
db2 = (1/m) *(np.sum(dZ2,axis=1,keepdims=True))
dZ1 = np.dot(W2.T,dZ2) * (1 - np.power(A1,2))
dW1 = (1/m) *(np.dot(dZ1,X.T))
db1 = (1/m) *(np.sum(dZ1, axis=1, keepdims=True))
### END CODE HERE ###
grads = {"dW1": dW1,
"db1": db1,
"dW2": dW2,
"db2": db2}
return grads
parameters, cache, X_assess, Y_assess = backward_propagation_test_case()
grads = backward_propagation(parameters, cache, X_assess, Y_assess)
print ("dW1 = "+ str(grads["dW1"]))
print ("db1 = "+ str(grads["db1"]))
print ("dW2 = "+ str(grads["dW2"]))
print ("db2 = "+ str(grads["db2"]))
```
**Expected output**:
<table style="width:80%">
<tr>
<td>**dW1**</td>
<td> [[ 0.00301023 -0.00747267]
[ 0.00257968 -0.00641288]
[-0.00156892 0.003893 ]
[-0.00652037 0.01618243]] </td>
</tr>
<tr>
<td>**db1**</td>
<td> [[ 0.00176201]
[ 0.00150995]
[-0.00091736]
[-0.00381422]] </td>
</tr>
<tr>
<td>**dW2**</td>
<td> [[ 0.00078841 0.01765429 -0.00084166 -0.01022527]] </td>
</tr>
<tr>
<td>**db2**</td>
<td> [[-0.16655712]] </td>
</tr>
</table>
**Question**: Implement the update rule. Use gradient descent. You have to use (dW1, db1, dW2, db2) in order to update (W1, b1, W2, b2).
**General gradient descent rule**: $ \theta = \theta - \alpha \frac{\partial J }{ \partial \theta }$ where $\alpha$ is the learning rate and $\theta$ represents a parameter.
**Illustration**: The gradient descent algorithm with a good learning rate (converging) and a bad learning rate (diverging). Images courtesy of Adam Harley.
<img src="images/sgd.gif" style="width:400;height:400;"> <img src="images/sgd_bad.gif" style="width:400;height:400;">
```
# GRADED FUNCTION: update_parameters
def update_parameters(parameters, grads, learning_rate):
"""
Updates parameters using the gradient descent update rule given above
Arguments:
parameters -- python dictionary containing your parameters
grads -- python dictionary containing your gradients
Returns:
parameters -- python dictionary containing your updated parameters
"""
# Retrieve each parameter from the dictionary "parameters"
### START CODE HERE ### (≈ 4 lines of code)
W1 = parameters["W1"]
b1 = parameters["b1"]
W2 = parameters["W2"]
b2 = parameters["b2"]
### END CODE HERE ###
# Retrieve each gradient from the dictionary "grads"
### START CODE HERE ### (≈ 4 lines of code)
dW1 = grads["dW1"]
db1 = grads["db1"]
dW2 = grads["dW2"]
db2 = grads["db2"]
## END CODE HERE ###
# Update rule for each parameter
### START CODE HERE ### (≈ 4 lines of code)
W1 = W1 - learning_rate * dW1
b1 = b1 - learning_rate * db1
W2 = W2 - learning_rate * dW2
b2 = b2 - learning_rate * db2
### END CODE HERE ###
parameters = {"W1": W1,
"b1": b1,
"W2": W2,
"b2": b2}
return parameters
parameters, grads = update_parameters_test_case()
parameters = update_parameters(parameters, grads,1.2)
print("W1 = " + str(parameters["W1"]))
print("b1 = " + str(parameters["b1"]))
print("W2 = " + str(parameters["W2"]))
print("b2 = " + str(parameters["b2"]))
```
**Expected Output**:
<table style="width:80%">
<tr>
<td>**W1**</td>
<td> [[-0.00643025 0.01936718]
[-0.02410458 0.03978052]
[-0.01653973 -0.02096177]
[ 0.01046864 -0.05990141]]</td>
</tr>
<tr>
<td>**b1**</td>
<td> [[ -1.02420756e-06]
[ 1.27373948e-05]
[ 8.32996807e-07]
[ -3.20136836e-06]]</td>
</tr>
<tr>
<td>**W2**</td>
<td> [[-0.01041081 -0.04463285 0.01758031 0.04747113]] </td>
</tr>
<tr>
<td>**b2**</td>
<td> [[ 0.00010457]] </td>
</tr>
</table>
### 4.4 - Integrate parts 4.1, 4.2 and 4.3 in nn_model() ####
**Question**: Build your neural network model in `nn_model()`.
**Instructions**: The neural network model has to use the previous functions in the right order.
```
# NN_model
def nn_model(X, Y, n_h, learning_rate, num_iterations = 10000, print_cost=False):
n_x = layer_sizes(X, Y)[0]
n_y = layer_sizes(X, Y)[2]
# Initialize parameters
parameters = initialize_parameters(n_x, n_h, n_y)
W1 = parameters["W1"]
b1 = parameters["b1"]
W2 = parameters["W2"]
b2 = parameters["b2"]
# Loop (gradient descent)
for i in range(0, num_iterations):
# Forward propagation. Inputs: "X, parameters". Outputs: "A2, cache"
A2, cache = forward_propagation(X, parameters)
# Cost function. Inputs: "A2, Y, parameters". Outputs: "cost"
cost = compute_cost(A2, Y, parameters)
# Backpropagation. Inputs: "parameters, cache, X, Y". Outputs: "grads"
grads = backward_propagation(parameters, cache, X, Y)
# Update rule for each parameter
parameters = update_parameters(parameters, grads, learning_rate)
# If print_cost=True, Print the cost every 1000 iterations
if print_cost and i % 1000 == 0:
print ("Cost after iteration %i: %f" %(i, cost))
# Returns parameters learnt by the model. They can then be used to predict output
return parameters
X_assess, Y_assess = nn_model_test_case()
parameters = nn_model(X_assess, Y_assess, 4, 1.02,num_iterations=10000, print_cost=True)
print("W1 = " + str(parameters["W1"]))
print("b1 = " + str(parameters["b1"]))
print("W2 = " + str(parameters["W2"]))
print("b2 = " + str(parameters["b2"]))
```
**Expected Output**:
<table style="width:90%">
<tr>
<td>
**cost after iteration 0**
</td>
<td>
0.692739
</td>
</tr>
<tr>
<td>
<center> $\vdots$ </center>
</td>
<td>
<center> $\vdots$ </center>
</td>
</tr>
<tr>
<td>**W1**</td>
<td> [[-0.65848169 1.21866811]
[-0.76204273 1.39377573]
[ 0.5792005 -1.10397703]
[ 0.76773391 -1.41477129]]</td>
</tr>
<tr>
<td>**b1**</td>
<td> [[ 0.287592 ]
[ 0.3511264 ]
[-0.2431246 ]
[-0.35772805]] </td>
</tr>
<tr>
<td>**W2**</td>
<td> [[-2.45566237 -3.27042274 2.00784958 3.36773273]] </td>
</tr>
<tr>
<td>**b2**</td>
<td> [[ 0.20459656]] </td>
</tr>
</table>
### 4.5 Predictions
**Question**: Use your model to predict by building predict().
Use forward propagation to predict results.
**Reminder**: predictions = $y_{prediction} = \mathbb 1 \text{{activation > 0.5}} = \begin{cases}
1 & \text{if}\ activation > 0.5 \\
0 & \text{otherwise}
\end{cases}$
As an example, if you would like to set the entries of a matrix X to 0 and 1 based on a threshold you would do: ```X_new = (X > threshold)```
```
# GRADED FUNCTION: predict
def predict(parameters, X):
"""
Using the learned parameters, predicts a class for each example in X
Arguments:
parameters -- python dictionary containing your parameters
X -- input data of size (n_x, m)
Returns
predictions -- vector of predictions of our model (red: 0 / blue: 1)
"""
# Computes probabilities using forward propagation, and classifies to 0/1 using 0.5 as the threshold.
### START CODE HERE ### (≈ 2 lines of code)
A2, cache = forward_propagation(X, parameters)
predictions = (A2 > 0.5)
### END CODE HERE ###
return predictions
parameters, X_assess = predict_test_case()
predictions = predict(parameters, X_assess)
print("predictions mean = " + str(np.mean(predictions)))
```
**Expected Output**:
<table style="width:40%">
<tr>
<td>**predictions mean**</td>
<td> 0.666666666667 </td>
</tr>
</table>
It is time to run the model and see how it performs on a planar dataset. Run the following code to test your model with a single hidden layer of $n_h$ hidden units.
```
# Build a model with a n_h-dimensional hidden layer
parameters = nn_model(X, Y, 4, 1.2 , num_iterations = 10000, print_cost=True)
# Plot the decision boundary
plot_decision_boundary(lambda x: predict(parameters, x.T), X, Y)
plt.title("Decision Boundary for hidden layer size " + str(4))
```
**Expected Output**:
<table style="width:40%">
<tr>
<td>**Cost after iteration 9000**</td>
<td> 0.218607 </td>
</tr>
</table>
```
# Print accuracy
predictions = predict(parameters, X)
print ('Accuracy: %d' % float((np.dot(Y,predictions.T) + np.dot(1-Y,1-predictions.T))/float(Y.size)*100) + '%')
```
**Expected Output**:
<table style="width:15%">
<tr>
<td>**Accuracy**</td>
<td> 90% </td>
</tr>
</table>
Accuracy is really high compared to Logistic Regression. The model has learnt the leaf patterns of the flower! Neural networks are able to learn even highly non-linear decision boundaries, unlike logistic regression.
Now, let's try out several hidden layer sizes.
### 4.6 - Tuning hidden layer size (optional/ungraded exercise) ###
Run the following code. It may take 1-2 minutes. You will observe different behaviors of the model for various hidden layer sizes.
```
# This may take about 2 minutes to run
plt.figure(figsize=(16, 32))
hidden_layer_sizes = [1, 2, 3, 4, 5, 20, 50]
for i, n_h in enumerate(hidden_layer_sizes):
plt.subplot(5, 2, i+1)
plt.title('Hidden Layer of size %d' % n_h)
parameters = nn_model(X, Y, n_h,1.2, num_iterations = 5000)
plot_decision_boundary(lambda x: predict(parameters, x.T), X, Y)
predictions = predict(parameters, X)
accuracy = float((np.dot(Y,predictions.T) + np.dot(1-Y,1-predictions.T))/float(Y.size)*100)
print ("Accuracy for {} hidden units: {} %".format(n_h, accuracy))
```
**Interpretation**:
- The larger models (with more hidden units) are able to fit the training set better, until eventually the largest models overfit the data.
- The best hidden layer size seems to be around n_h = 5. Indeed, a value around here seems to fits the data well without also incurring noticeable overfitting.
- You will also learn later about regularization, which lets you use very large models (such as n_h = 50) without much overfitting.
**Optional questions**:
**Note**: Remember to submit the assignment by clicking the blue "Submit Assignment" button at the upper-right.
Some optional/ungraded questions that you can explore if you wish:
- What happens when you change the tanh activation for a sigmoid activation or a ReLU activation?
- Play with the learning_rate. What happens?
- What if we change the dataset? (See part 5 below!)
<font color='blue'>
**You've learnt to:**
- Build a complete neural network with a hidden layer
- Make a good use of a non-linear unit
- Implemented forward propagation and backpropagation, and trained a neural network
- See the impact of varying the hidden layer size, including overfitting.
Nice work!
## 5) Performance on other datasets
If you want, you can rerun the whole notebook (minus the dataset part) for each of the following datasets.
```
# Datasets
noisy_circles, noisy_moons, blobs, gaussian_quantiles, no_structure = load_extra_datasets()
datasets = {"noisy_circles": noisy_circles,
"noisy_moons": noisy_moons,
"blobs": blobs,
"gaussian_quantiles": gaussian_quantiles}
### START CODE HERE ### (choose your dataset)
dataset = "noisy_moons"
### END CODE HERE ###
X, Y = datasets[dataset]
X, Y = X.T, Y.reshape(1, Y.shape[0])
# make blobs binary
if dataset == "blobs":
Y = Y%2
# Visualize the data
plt.scatter(X[0, :], X[1, :], c=Y, s=40, cmap=plt.cm.Spectral);
```
Congrats on finishing this Programming Assignment!
Reference:
- http://scs.ryerson.ca/~aharley/neural-networks/
- http://cs231n.github.io/neural-networks-case-study/
| github_jupyter |
# Reto 01
> Consiste en encontrar la carpeta de descargas del usuario (independientemente de sus personalizaciones, tiene que funcionar si la tiene en otro lado o es un usuario cualquiera) y listar solo los archivos de la carpeta.
Sacado de https://j2logo.com/python/listar-directorio-en-python/
Para listar o recorrer un directorio:
```
import os
contenido = os.listdir('/Users/noave/Downloads')
contenido
```
El primer problema es que me muestra también todas las carpetas. El segundo, que aún no sé cómo encontrar un directorio con un comodín. Al parecer, en Windows requiere acceder al registro, pero he leído que hay otro método.
Para mostrar los archivos de tipo jpg de una carpeta:
```
import os
ejemplo_dir = '/Users/noave/Downloads'
contenido = os.listdir(ejemplo_dir)
imagenes = []
for fichero in contenido:
if os.path.isfile(os.path.join(ejemplo_dir, fichero)) and fichero.endswith('.jpg'):
imagenes.append(fichero)
print(imagenes)
```
Con scandir() puedo listar los ficheros de un directorio. Es la opción recomendada.
```
import os
ejemplo_dir = '/Users/noave/Downloads'
with os.scandir(ejemplo_dir) as ficheros:
for fichero in ficheros:
print(fichero.name)
```
Y también está iterdir() del módulo pathlib, también recomendado y que permite filtrar con is_file() o is_dir().
```
import pathlib
ejemplo_dir = '/Users/noave/Downloads'
directorio = pathlib.Path(ejemplo_dir)
for fichero in directorio.iterdir():
print(fichero.name)
```
Para recorrer solo los ficheros de un directorio, dos métodos con lo que hemos visto.
```
import os
ejemplo_dir = '/Users/noave/Downloads'
with os.scandir(ejemplo_dir) as ficheros:
ficheros = [fichero.name for fichero in ficheros if fichero.is_file()]
print(ficheros)
import pathlib
ejemplo_dir = '/Users/noave/Downloads/'
directorio = pathlib.Path(ejemplo_dir)
ficheros = [fichero.name for fichero in directorio.iterdir() if fichero.is_file()]
for i in ficheros:
print(i)
#Y si solo queremos las carpetas:
import os
ejemplo_dir = '/Users/noave/Downloads/'
with os.scandir(ejemplo_dir) as ficheros:
subdirectorios = [fichero.name for fichero in ficheros if fichero.is_dir()]
print(subdirectorios)
```
Pues ya tenemos un ganador. Ahora solo nos falta descubrir la manera de que Python encuentre la carpeta de descargas, esté donde esté.
```
DIRECTORIO_BASE_2 = os.getcwd()
print(DIRECTORIO_BASE_2)
```
https://stackoverflow.com/questions/35851281/python-finding-the-users-downloads-folder
Alguien dio esta posible solución, pero como toca registro, no me atrevo a meterla tal cual:
```import os
def get_download_path():
"""Returns the default downloads path for linux or windows"""
if os.name == 'nt':
import winreg
sub_key = r'SOFTWARE\Microsoft\Windows\CurrentVersion\Explorer\Shell Folders'
downloads_guid = '{374DE290-123F-4565-9164-39C4925E467B}'
with winreg.OpenKey(winreg.HKEY_CURRENT_USER, sub_key) as key:
location = winreg.QueryValueEx(key, downloads_guid)[0]
return location
else:
return os.path.join(os.path.expanduser('~'), 'downloads')
```
Este me inspira más confianza, aunque de novato no tiene nada:
```
import os
if os.name == 'nt':
import ctypes
from ctypes import windll, wintypes
from uuid import UUID
#ctypes GUID copied from MSDN sample code
class GUID(ctypes.Structure):
_fields_ = [
("Data1", wintypes.DWORD),
("Data2", wintypes.WORD),
("Data3", wintypes.WORD),
("Data4", wintypes.BYTE * 8)
]
def __init__(self, uuidstr):
uuid = UUID(uuidstr)
ctypes.Structure.__init__(self)
self.Data1, self.Data2, self.Data3, \
self.Data4[0], self.Data4[1], rest = uuid.fields
for i in range(2, 8):
self.Data4[i] = rest>>(8-i-1)*8 & 0xff
SHGetKnownFolderPath = windll.shell32.SHGetKnownFolderPath
SHGetKnownFolderPath.argtypes = [
ctypes.POINTER(GUID), wintypes.DWORD,
wintypes.HANDLE, ctypes.POINTER(ctypes.c_wchar_p)
]
def _get_known_folder_path(uuidstr):
pathptr = ctypes.c_wchar_p()
guid = GUID(uuidstr)
if SHGetKnownFolderPath(ctypes.byref(guid), 0, 0, ctypes.byref(pathptr)):
raise ctypes.WinError()
return pathptr.value
FOLDERID_Download = '{374DE290-123F-4565-9164-39C4925E467B}'
def get_download_folder():
return _get_known_folder_path(FOLDERID_Download)
else:
def get_download_folder():
home = os.path.expanduser("~")
return os.path.join(home, "Downloads")
print(get_download_folder())
```
```
def get_download_folder():
home = os.path.expanduser("~")
return os.path.join(home, "Downloads")
print(get_download_folder())
```
Vale, me ha funcionado. Supongo que si moviese la carpeta a otra parte, dejaría de funcionar (en realidad es la ruta por defecto home+downloads). Pero ya es algo.
Ahora solo me falta… Hacer que imprima los archivos de ese directorio.
```
import pathlib
def get_download_folder():
home = os.path.expanduser("~")
return os.path.join(home, "Downloads")
print(get_download_folder())
ejemplo_dir = get_download_folder()
directorio = pathlib.Path(ejemplo_dir)
ficheros = [fichero.name for fichero in directorio.iterdir() if fichero.is_file()]
for i in ficheros:
print(i)
```
Reto conseguido. ¿Funcionará con otros usuarios?
```
import os
def get_download_folder():
home = os.path.expanduser("~")
return os.path.join(home, "Downloads")
print(get_download_folder())
ejemplo_dir = get_download_folder()
with os.scandir(ejemplo_dir) as ficheros:
ficheros = [fichero.name for fichero in ficheros if fichero.is_file()]
for i in ficheros:
print(i)
```
---
# Reto 2
> En este caso, y partiendo de la solución anterior, ahora hay que trabajar con archivos. El objetivo es que listes todos los archivos JPEG que se encuentran en el directorio de descargas del usuario.
> Tienes que tener en cuenta, que imágenes JPEG, son los .jpg o .jpeg o incluso JPEG por citarte algunas. Pero además en el listado se cumplirán las siguientes condiciones,
> - Si en el nombre del archivo aparece un número, el nombre del achivo apararecerá en mayúsculas, mientras que si no tiene ningún número aparecerá en minúsculas.
> - Además en ese listado, los que sean pares, tendrán una flecha delante, del tipo =>.
> Como ves se trata de un reto sencillo, y que nos permitirá profundizar en bucles, módulos y otros temas similares.
Este reto tiene una parte que ya está solucionada más arriba, en mis notas del reto 1. Es la parte de los archivos en JPG. Para extender las variaciones dentro de la extensión (jpeg, JPEG, jpg, etc.), tal vez pueda aplicar expresiones regulares. No sé si algo como jp*g funcionará, imagino que tendría que importar el módulo y ver cuál es la forma correcta. Y, de paso, tal vez tenga una forma de evitar el case sensitive sin recurrir a .lower().
También habrá que hacer un bucle for para cada carácter dentro de cada archivo, un for i in str, para determinar si hay números. Aquí imagino que sí necesitaré expresiones regulares, ya que al ser str no puedo preguntar si hay int. Necesitaría filtrar con [0-9]. Si la respuesta es True: upper(str).
Por último, hay varias formas de hacer algo con los elementos pares o impares. Se puede hacer con bucles while tanto con for. Por lo general, se determinan los pares cuando su index % 2 = 0. O bien podemos establecer un range(start, end, 2), pero no me acuerdo de si puedo usar el valor len(archivos) para determinar el end del range.
Empecemos.
Antes de andar jugueteando con mis directorios, voy a simular con una lista.
```
# Encontrar archivos cuya extensión coincida con .jpg y variantes:
import re
archivos = ['imagen01.jpg', 'oso_trospido.JPG', "Karaoke.mp3", 'Ashjkdaslkl.JPEG', 'AKAKAKAKA9.jpeg', 'final_image.jpg']
patron = re.compile(r"\.jp[e]?g", re.IGNORECASE)
prueba = "prueba.JPEG"
print(patron.search(prueba)) # No sé por qué, pero .match() me da siempre None.
print(prueba, "\n---------------")
for i in archivos:
print(patron.search(i))
if patron.search(i):
print(i)
# Determinar si los archivos llevan números.
# Si es True, ponerlo todo en mayúsculas.
import re
archivos = ['imagen01.jpg', 'oso_trospido.JPG',
'Ashjkdaslkl.JPEG', 'AKAKAKAKA9.jpeg', 'final_image.jpg']
patron = re.compile(r"[0-9]", re.IGNORECASE)
prueba = "imagen5.jpg"
print(patron.search(prueba))
print(prueba, "\n---------------")
for i in archivos:
# print(patron.search(i))
if patron.search(i):
print(i.upper())
else:
print(i)
# Si es par, añadir una flecha.
archivos = ['imagen01.jpg', 'oso_trospido.JPG',
'Ashjkdaslkl.JPEG', 'AKAKAKAKA9.jpeg', 'final_image.jpg']
pos=0
for i in archivos:
pos+=1
if pos%2==0:
print("=> ", i)
else:
print(i)
```
Ya tendríamos prácticamente todo por separado. Vamos a ponerlo todo junto.
```
import os
import re
def get_download_folder():
home = os.path.expanduser("~")
return os.path.join(home, "Downloads")
print(get_download_folder())
patronJPG = re.compile(r"\.jp[e]?g", re.IGNORECASE)
patronNum = re.compile(r"[0-9]", re.IGNORECASE)
descargasDir = get_download_folder()
contenido = os.listdir(descargasDir)
imagenes = []
for fichero in contenido:
if os.path.isfile(os.path.join(descargasDir, fichero)) and patronJPG.search(fichero):
imagenes.append(fichero)
pos = 0
for i in imagenes:
pos += 1
if pos % 2 == 0:
i = "=> " + i
if patronNum.search(i):
print(i.upper())
else:
print(i.lower())
```
¡Reto número 2 conseguido!
---
`[Actualización 220307]`
Al hacer Pull Request, Github me da el siguiente mensaje a través de [pep8speaks](https://github.com/pep8speaks):
> Hello @NoaVelasco! Thanks for opening this PR. We checked the lines you've touched for PEP 8 issues, and found:
- In the file reto-01/NoaVelasco/reto01.py:
Line 3:1: E302 expected 2 blank lines, found 1
Line 4:9: E117 over-indented
Line 6:1: W293 blank line contains whitespace
Line 7:1: E305 expected 2 blank lines after class or function definition, found 1
Line 7:29: W291 trailing whitespace
Line 13:13: W292 no newline at end of file
- In the file reto-02/NoaVelasco/reto02.py:
Line 19:80: E501 line too long (89 > 79 characters)
Así que he hecho algunos cambios para dejar el código lo mejor posible.
| github_jupyter |
```
# default_exp mailer
# hide
_FNAME='mailer'
import unittest
from unittest import mock
from nbdev.export import notebook2script
import os
TESTCASE = unittest.TestCase()
_nbpath = os.path.join(_dh[0], _FNAME+'.ipynb')
notebook2script(_nbpath)
#export
import os
import logging
from sendgrid import SendGridAPIClient
from sendgrid.helpers.mail import Mail
from sewing import is_main, start_log
logger = logging.getLogger()
DEFAULT_MAIL_API = None
def set_default_mail_api(mail_api=None):
global DEFAULT_MAIL_API
if mail_api is None:
SENDGRID_API_KEY = os.environ.get("SENDGRID_KEY")
mail_api = SendGridAPIClient(SENDGRID_API_KEY)
mail_api.api_name='sendgrid'
DEFAULT_MAIL_API=mail_api
return mail_api
FROM_DEFAULT = 'will@endao.network'
def send(to_emails, subject, html_content, from_email=FROM_DEFAULT, mail_api=DEFAULT_MAIL_API):
logger.debug('Email request received {}'.format(
{'to': to_emails,
'subject': subject,
'body': html_content,
'from': from_email
}))
if mail_api is None:
mail_api = set_default_mail_api()
message = Mail(from_email = from_email,
to_emails = to_emails,
subject = subject,
html_content = html_content)
email_resp = mail_api.send(message)
er = email_resp
logger.debug('Email sent with response {}'.format({
'status_code': er.status_code,
'headers': dict(er.headers),
'body': er.body}))
return email_resp
def send_test_email(to_emails=None, from_email=FROM_DEFAULT, subject='Test email', content_file=None):
if not content_file:
html_content='This was an automated email.'
else:
with open(content_file, 'r') as f:
html_content = f.read()
return send(to_emails=to_emails, from_email=from_email, subject=subject, html_content=html_content)
if is_main(globals()):
start_log()
import argparse
parser = argparse.ArgumentParser("Send test email")
parser.add_argument('--to', type=str, help="Send email to")
parser.add_argument('--sender', type=str, help='From email address', default=FROM_DEFAULT)
parser.add_argument('--content', type=str, help="Filename of email body", default=None)
parser.add_argument('--subject', type=str, help="Email subject", default='Test email')
args = parser.parse_args()
send_test_email(to_emails=[args.to], from_email=args.sender, subject=args.subject, content_file=args.content)
```
| github_jupyter |
# Simulating Language 5, Simple Innate Signalling (lab) (some answers)
The code below builds up to a function, ```ca_monte```, which measures and returns the level of communicative accuracy between a production system and a reception system. Signalling systems are stored as lists of lists of association weights. This list of lists structure can be thought of as a matrix with meanings on the rows and signals on the columns (for production matrices) or signals on the rows and meanings on the columns (for reception matrices).
You should go through this notebook cell by cell, running each cell by using SHIFT+ENTER.
First, we import the random library and the usual plotting stuff.
```
import random
%matplotlib inline
import matplotlib.pyplot as plt
from IPython.display import set_matplotlib_formats
set_matplotlib_formats('svg', 'pdf')
```
Now here's the bulk of the code, in three functions, ```wta``` (for "winner take all"), ```communicate```, and ```ca_monte```.
```
def wta(items):
maxweight = max(items)
candidates = []
for i in range(len(items)):
if items[i] == maxweight:
candidates.append(i)
return random.choice(candidates)
def communicate(speaker_system, hearer_system, meaning):
speaker_signal = wta(speaker_system[meaning])
hearer_meaning = wta(hearer_system[speaker_signal])
if meaning == hearer_meaning:
return 1
else:
return 0
def ca_monte(speaker_system, hearer_system, trials):
total = 0.
accumulator = []
for n in range(trials):
total += communicate(speaker_system, hearer_system,
random.randrange(len(speaker_system)))
accumulator.append(total / (n + 1))
return accumulator
```
We'll try and understand what this code does and why in a minute, but for now let's try getting it to do something. Let's say we had three meanings, and three signals. Let's imagine a speaker who used signal 0 for meaning 0, signal 1 for meaning 1, and both signal 1 and 2 for meaning 2, and a hearer who understood meaning 0 for signal 0, meaning 1 or 2 for signal 1, and meaning 2 for signal 2. What do we expect the communicative accuracy to be for these two individuals?
Try and figure this out in your head first (it might help to write out the matrices on a piece of paper).
Now, let's simulate it! Enter the following code in the next cell (note I've formatted the list of lists nicely so you can read them as matrices, but these can also be entered all on one line if you want):
```python
speaker = [[1, 0, 0],
[0, 1, 0],
[0, 1, 1]]
hearer = [[1, 0, 0],
[0, 1, 1],
[0, 0, 1]]
ca_monte(speaker, hearer, 100)
```
This simulates 100 interactions between the speaker and hearer and returns a list consisting of the average communicative accuracy so far at each interaction (so the values in this list become more and more accurate reflections of the communicative accuracy as you go through the list).
```
speaker = [[1, 0, 0],
[0, 1, 0],
[0, 1, 1]]
hearer = [[1, 0, 0],
[0, 1, 1],
[0, 0, 1]]
ca_monte(speaker, hearer, 100)
```
So, this has given us our first simulation results! We can look at these as a big list, but it might be better to plot it as a graph. Try the following, for example:
```python
data = ca_monte(speaker, hearer, 100)
plt.plot(data)
plt.xlabel('trials')
plt.ylabel('communicative accuracy')
plt.ylim([0,1])
plt.grid()
```
```
data = ca_monte(speaker, hearer, 100)
plt.plot(data)
plt.xlabel('trials')
plt.ylabel('communicative accuracy')
plt.ylim([0,1])
plt.grid()
```
So, this plots a single simulation run on a graph. More often, we want to plot lots of runs on a graph. We can do this by making a loop, and gathering a lot of simulation runs in a list. (This will actually be a list of lists now...)
For example, we could do the following to get ten simulation runs:
```python
lots_of_data = []
for n in range(10):
data = ca_monte(speaker, hearer, 100)
lots_of_data.append(data)
print(lots_of_data)
```
```
lots_of_data = []
for n in range(10):
data = ca_monte(speaker, hearer, 100)
lots_of_data.append(data)
print(lots_of_data)
```
Hmmm... I'm not sure we want to spend much time looking at screens full of numbers! Let's plot a graph!
The following command produces a nicer visualisation of our data, showing all the graphs with transparent lines so we can see the aggregate behaviour of multiple simulations. See if you can figure out how this bit of code works. _Hint: in graphics "alpha" is a technical term for transparency._
```python
for data in lots_of_data:
plt.plot(data, color='orange', alpha=0.5)
plt.xlabel('trials')
plt.ylabel('communicative accuracy')
plt.ylim([0,1])
plt.grid()
```
```
for data in lots_of_data:
plt.plot(data, color='orange', alpha=0.5)
plt.xlabel('trials')
plt.ylabel('communicative accuracy')
plt.ylim([0,1])
plt.grid()
```
Another useful type of plot is a histogram showing a distribution of values. Let's say we wanted to know what the end result of a lot of simulation runs looked like. We need to do a little bit of work to pull out the last item of each run of a simulation and put it in a list. This code will do that:
```python
end_result = []
for n in range(10):
data = ca_monte(speaker, hearer, 100)
last_item = data[len(data) - 1]
end_result.append(last_item)
```
Now we can plot the distribution using another handy plotting command, `hist`:
```python
plt.hist(end_result)
plt.xlabel('communicative accuracy')
plt.xlim([0, 1])
```
```
end_result = []
for n in range(10):
data = ca_monte(speaker, hearer, 100)
last_item = data[len(data) - 1]
end_result.append(last_item)
plt.hist(end_result)
plt.xlabel('communicative accuracy')
plt.xlim([0, 1])
```
OK, now we've had a bit of fun, let's understand what's going on!
Try and understand what each function does. Look at the main function `ca_monte` first, then the function it calls (`communicate`), and so on until you have inspected each function separately. Can you see why the program has been divided into functions in the way it has?
If you're not sure what each function does, you can have a look at the separate (very!) detailed walkthrough notebook I've provided on the Learn page. Work through that first and then come back here.
When you are satisfied that you understand roughly how the code works, answer the following questions. 1-3 should be completed by everyone. **4 and 5 are optional!** Only attempt them if you are happy you have completed 1-3.
1. How many trials should there be in the Monte Carlo simulation to work out communicative accuracy? *Hint: answer this question empirically by plotting the results and comparing to what the "real" answer should be for various numbers of trials.*
```
end_results = []
for trials in [10, 100, 1000, 10000]:
end_result = []
for n in range(50):
data = ca_monte(speaker, hearer, trials)
last_item = data[len(data) - 1]
end_result.append(last_item)
end_results.append(end_result)
fig, axs = plt.subplots(nrows=4, ncols=1, sharex=True)
for i in range(4):
axs[i].hist(end_results[i], range=(0,1), bins=50,
label=['10 trials', '100 trial', '1000 trials', '10000 trials'][i])
axs[i].legend()
axs[3].set_xlabel('communicative accuracy')
```
2. How do synonymy and homonymy affect communicative accuracy? Create production and reception systems with different degrees of homonymy and synonymy to explore this. *Note: you don’t have to start from the production and reception systems given above, and you don’t have to restrict yourself to systems with 3 meanings and 3 signals.*
3. What alternatives to "winner-take-all" might there be in the model of production/ reception? What difference might this make? Would they be more or less realistic, or powerful as a model? *Hint: how might you interpret weights as probabilities?*
4. **[optional]** How could you model production and reception using a single underlying matrix, rather than separate production and reception matrices? Is this kind of model better or worse than a model where we use separate matrices?
5. **[optional]** How would you go about calculating communicative accuracy exactly, i.e. rather than via Monte Carlo techniques?
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.