text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
```
#r "nuget:Newtonsoft.Json, 12.0.2"
#r "nuget:CsvHelper, 26.1.0"
#r "nuget:DataBoss.DataPackage, 0.0.93"
using DataBoss.Data;
using DataBoss.DataPackage;
```
_visualization functions, you can safely skip the following cell_
```
using System.Data;
using Microsoft.DotNet.Interactive.Formatting;
using static Microsoft.DotNet.Interactive.Formatting.PocketViewTags;
static void Dump(DataPackage dp, int? limit = null) {
var rs = dp.Resources
.Where(x => x.Format == "csv")
.Select(x => (Heading: th(x.Name), Result: td(ReadAsHtmlTable(x.Read(), limit: limit))))
.ToList();
display(table(
thead(rs.Select(x => x.Heading)),
tbody(rs.Select(x => x.Result))
));
}
static object Dump(DataBoss.DataPackage.TabularDataResource xs, int? limit = null) => Dump(xs.Read(), limit: limit);
static object Dump(IDataReader xs, int? limit = null) => display(ReadAsHtmlTable(xs, limit: limit));
static object ReadAsHtmlTable(IDataReader xs, int? limit = null) {
try {
limit ??= int.MaxValue;
var rows = new List<object>();
for(var i = 0;xs.Read() && i < limit; ++i)
rows.Add(Enumerable.Range(0, xs.FieldCount).Select(x => td(xs.GetValue(x))).ToList());
return table(
thead(Enumerable.Range(0, xs.FieldCount).Select(x => th[style:"font-weight:bold"](xs.GetName(x)))),
tbody(rows.Select(x => tr(x))));
} finally {
xs.Dispose();
}
}
```
# Defining a simple resource
```
var dp = new DataPackage();
dp.AddResource(xs => xs.WithName("numbers").WithData(Enumerable.Range(0, 2).Select(x => new { Value = x })));
Dump(dp);
```
**DataPackage.Load** supports directory paths containing a datapackage.json, zip files and http.
```
var countries = DataPackage.Load(@"https://datahub.io/core/country-list/r/country-list_zip.zip");
Dump(countries.GetResource("data_csv"), limit: 10);
```
# Resource (DataReader) Transformation
```
var c2 = new DataPackage();
c2.AddResource(countries.GetResource("data_csv"));
c2.UpdateResource("data_csv", xs => xs
.WithName("countries") //resource can be renamed.
.Transform(x =>
{
var id = 0;
x.Transform("Code", (string value) => value.ToLower()); //typed transform
x.Add(0, "Id", r => ++id); //columns can be added at any existing ordinal
x.Transform("Name", (string value) => value.ToUpper());
x.Add("NameLen", r => r["Name"].ToString().Length); //record based
x.Add("Source", r => $"{r.Source["Name"]} ({r.Source["Code"]})"); //from non transformed source
})
);
Dump(c2, limit: 10);
```
# Creating Resrouces Incrementally
```
var n = 0;
var numbers = Enumerable.Range(0, 3).Select(x => new { Value = ++n });
var myNumbers = new DataPackage();
void AddOrAppend<T>(DataPackage dp, string name, IEnumerable<T> rows) {
dp.AddOrUpdateResource(name, xs => xs.WithData(rows), xs => xs.WithData(() => xs.Read().Concat(rows)));
}
AddOrAppend(myNumbers, "numbers", numbers.ToList());
Dump(myNumbers);
AddOrAppend(myNumbers, "numbers", numbers.ToList());
Dump(myNumbers);
```
| github_jupyter |
# Azure ML Inference Pipeline for HIFIS-RNN-MLP
This notebook defines an Azure machine learning pipeline for batch inference and submits the pipeline as an experiment to be run on an Azure virtual machine. It then publishes the pipeline in the workspace and schedules it to be run at a regular interval.
```
# Import statements
import azureml.core
from azureml.core import Experiment
from azureml.core import Workspace, Datastore
from azureml.data.data_reference import DataReference
from azureml.pipeline.core import PipelineData
from azureml.pipeline.core import Pipeline, PipelineRun
from azureml.pipeline.core.schedule import ScheduleRecurrence, Schedule
from azureml.pipeline.steps import PythonScriptStep, EstimatorStep
from azureml.train.dnn import TensorFlow
from azureml.train.estimator import Estimator
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.core.compute_target import ComputeTargetException
from azureml.core.environment import Environment
from azureml.core.runconfig import RunConfiguration
import shutil
```
### Register the workspace and configure its Python environment.
```
# Get reference to the workspace
ws = Workspace.from_config("./ws_config.json")
# Set workspace's environment
env = Environment.from_pip_requirements(name = "HIFIS_env", file_path = "./../requirements.txt")
env.python.conda_dependencies.add_pip_package("azureml-core")
env.python.conda_dependencies.add_pip_package("sendgrid")
env.register(workspace=ws)
runconfig = RunConfiguration(conda_dependencies=env.python.conda_dependencies)
print(env.python.conda_dependencies.serialize_to_string())
# Move AML ignore file to root folder
aml_ignore_path = shutil.copy('./.amlignore', './../.amlignore')
```
### Create references to persistent and intermediate data
Create DataReference objects that point to our raw data on the blob. Configure a PipelineData object to point to preprocessed data stored on the blob. Pipeline data is intermediate, meaning that it is produced by a step and will be fed as input to a subsequent step.
```
# Get the blob datastores associated with this workspace
hifis_blob_ds = Datastore(ws, name='hifisrnnmlp_ds')
raw_data_blob_ds = Datastore(ws, name='hifis_raw_ds')
# Create data references to folders on the blobs
raw_data_dr = DataReference(
datastore=raw_data_blob_ds,
data_reference_name="raw_data",
path_on_datastore="hifis/")
inference_dr = DataReference(
datastore=hifis_blob_ds,
data_reference_name="inference",
path_on_datastore="inference/")
# Set up intermediate pipeline data to store preprocessed data and serialized objects needed for inference (e.g. explainer)
preprocess_pd = PipelineData(
"preprocessed_output",
datastore=hifis_blob_ds,
output_name="preprocessed_output",
output_mode="mount")
```
### Compute Target
Specify and configure the compute target for this workspace. If a compute cluster by the name we specified does not exist, create a new compute cluster.
```
# Define some constants
CT_NAME = "ds3v2-infer" # Name of our compute cluster
VM_SIZE = "STANDARD_DS3_V2" # Specify the Azure VM for execution of our pipeline
MIN_NODES = 0 # Min number of compute nodes in cluster
MAX_NODES = 3 # Max number of compute nodes in cluster
# Set up the compute target for this experiment
try:
compute_target = AmlCompute(ws, CT_NAME)
print("Found existing compute target.")
except ComputeTargetException:
print("Creating new compute target")
provisioning_config = AmlCompute.provisioning_configuration(vm_size=VM_SIZE, min_nodes=MIN_NODES, max_nodes=MAX_NODES)
compute_target = ComputeTarget.create(ws, CT_NAME, provisioning_config) # Create the compute cluster
# Wait for cluster to be provisioned
compute_target.wait_for_completion(show_output=True, min_node_count=None, timeout_in_minutes=20)
print("Azure Machine Learning Compute attached")
print("Compute targets: ", ws.compute_targets)
compute_target = ws.compute_targets[CT_NAME]
```
### Define pipeline and submit experiment.
Define the steps of an Azure machine learning pipeline. Create an Azure Experiment that will run our pipeline. Submit the experiment to the execution environment.
```
# Define preprocessing step in the ML pipeline
step1 = PythonScriptStep(name="preprocess_step",
script_name="azure/preprocess_step/preprocess_step.py",
arguments=["--rawdatadir", raw_data_dr, "--inferencedir", inference_dr, "--preprocessedoutputdir",
preprocess_pd],
inputs=[raw_data_dr, inference_dr],
outputs=[preprocess_pd],
compute_target=compute_target,
source_directory="./../",
runconfig=runconfig,
params={"PIPELINE": "inference"},
allow_reuse=False)
# Define batch inference step in the ML pipeline
step2 = PythonScriptStep(name="inference_step",
script_name="azure/inference_step/inference_step.py",
arguments=["--preprocessedoutputdir", preprocess_pd, "--inferencedir", inference_dr],
inputs=[preprocess_pd, inference_dr],
outputs=[],
compute_target=compute_target,
source_directory="./../",
runconfig=runconfig,
allow_reuse=False)
# Construct the ML pipeline from the steps
steps = [step1, step2]
inference_pipeline = Pipeline(workspace=ws, steps=steps)
inference_pipeline.validate()
# Define a new experiment and submit a new pipeline run to the compute target.
experiment = Experiment(workspace=ws, name='BatchInferenceExperiment_v1')
inference_run = experiment.submit(inference_pipeline, regenerate_outputs=False)
print("Pipeline is submitted for execution")
# Move AML ignore file back to original folder
aml_ignore_path = shutil.move(aml_ignore_path, './.amlignore')
```
### Publish the pipeline
Wait for the pipeline run to finish. Then publish the pipeline. The pipeline will be visible as an endpoint in the Pipelines tab in the workspace on Azure Machine Learning studio. Delete the training compute cluster to prevent further cost.
```
# Wait for the pipeline to finish running.
inference_run.wait_for_completion()
# Publish the pipeline.
published_pipeline = inference_run.publish_pipeline(
name="HIFIS-RNN-MLP Inference Pipeline",
description="Azure ML Pipeline that runs model inference on all clients in HIFIS database and computes LIME explanations",
version="1.0")
```
### Schedule the pipeline
Schedule the published pipeline to run weekly.
```
recurrence = ScheduleRecurrence(frequency="Week", interval=1)
pipeline_id = "inference-pipeline"
recurring_schedule = Schedule.create(ws, name="WeeklyInferenceSchedule",
description="Scheduled weekly batch inference",
pipeline_id=published_pipeline.id,
experiment_name=experiment.name,
recurrence=recurrence)
```
### Disable scheduled pipelines
Disable all scheduled pipelines
```
schedules = Schedule.list(ws)
for schedule in schedules:
print(schedule)
schedule.disable()
```
| github_jupyter |
# Gems of the Python Standard Library
### Alex Zharichenko
# Data Types
## enum
---
#### Support for enumerations
```
from enum import Enum, IntEnum
class Color(IntEnum):
RED = 1
GREEN = 2
BLUE = 3
Color.RED
Color.RED == 1
# list(Color)
from enum import Flag, auto
class Color(Flag):
RED = auto()
BLUE = auto()
GREEN = auto()
WHITE = RED | BLUE | GREEN
list(Color)
```
## bisect
```
from bisect import bisect
def grade(score, breakpoints=[60, 70, 80, 90], grades='FDCBA'):
i = bisect(breakpoints, score)
return grades[i]
[grade(score) for score in [33, 99, 77, 70, 89, 90, 100]]
```
# Text Processing Services
## difflib
---
### Helpers for computing deltas
```
import difflib
s1 = ['bacon\n', 'eggs\n', 'ham\n', 'guido\n']
s2 = ['python\n', 'eggy\n', 'hamster\n', 'guido\n']
print(''.join(difflib.context_diff(s1, s2)))
print(''.join(difflib.unified_diff(s1, s2)))
print(''.join(difflib.ndiff(s1, s2)))
diff = list(difflib.ndiff(s1, s2))
print(''.join(difflib.restore(diff, 1)))
print(''.join(difflib.restore(diff, 2)))
difflib.get_close_matches('appel', ['ape', 'apple', 'peach', 'puppy'])
```
# File and Directory Access
## pathlib
---
### Object-oriented filesystem paths
```
from pathlib import Path
Path('.')
Path(r'C:/Users/Alex/Projects/gems-of-python-stdlib')
Path('.').absolute()
output_dir = Path() / 'output'
output_dir
print(output_dir.exists())
output_dir.mkdir()
print(output_dir.exists())
print(output_dir.is_dir())
some_file = output_dir / "test.txt"
with some_file.open("w") as f:
f.write("hello world")
some_file.is_file()
list(output_dir.glob("*.txt"))
```
# Concurrent Execution
## sched
---
### Event scheduler
```
import sched
import time
s = sched.scheduler()
def do_action(action="default"):
print("do_action", action, time.time())
s.enter(5, 1, do_action)
s.enter(10, 2, do_action, argument=("do_nothing",))
s.enter(10, 1, do_action, kwargs={"action": "do_something"})
s.queue
print(time.time())
s.run()
print(time.time())
```
# Internet Protocols and Support
## webbrowser
---
### Convenient Web-browser controller
```
import webbrowser
url = 'http://docs.python.org/'
webbrowser.open(url)
webbrowser.open_new(url)
webbrowser.open_new_tab(url)
webbrowser.get()
```
## UUID
---
### UUID objects according to RFC 4122¶
```
import uuid
# Version 1: date-time and host id
uuid.uuid1()
# Version 3 & 5: namespace, v3 using md5, v5 using sha1
uuid.uuid3(namespace=uuid.NAMESPACE_DNS, name='python.org')
uuid.uuid5(namespace=uuid.NAMESPACE_DNS, name='python.org')
# Version 4: Random
uuid.uuid4()
id = uuid.uuid4()
print(str(id))
print(id.int)
```
| github_jupyter |
---
title: Toggling code cells in Jupyter HTML Outputs
summary: "Or, How I Wasted an Afternoon."
date: 2020-01-11
source: jupyter
---
When writing a blog post in Twitter, I found there was no easy way to
reproduce the rather lovely **code folding** effect you get in
([Rmarkdown documents](https://bookdown.org/yihui/rmarkdown/html-document.html#code-folding)),
so I went and made one myself, illustrated here.
Actually, I've written a few of them, as the code changes slightly depending on where you want to achieve code folding.
First, here's some toggleable code.
```
for i in range(9, -1, -1):
if i > 0:
print('.' * (i) + ('%i' % i) + '.' * (9-i))
else:
print('>Lift Off!')
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
plt.hist(np.random.normal(0, 1, 100));
```
## Jupyter Magic
To activate toggling directly in a live Notebook using
the [%%JavaScript magic](https://jupyter-notebook.readthedocs.io/en/stable/examples/Notebook/JavaScript%20Notebook%20Extensions.html), just nclude the cell below in your notebook
If the notebook is [truster](https://jupyter-notebook.readthedocs.io/en/stable/security.html),
it will automatically run when you load the page.
```
%%javascript
function toggler(){
if(window.already_toggling){
// Don't add multiple buttons.
return 0
}
let btn = $('.input').append('<button>Toggle Code</button>')
.children('button');
btn.on('click', function(e){
let tgt = e.currentTarget;
$(tgt).parent().children('.inner_cell').toggle()
})
window.already_toggling = true;
}
// Since javascript cells are executed as soon as we load
// the notebook (if it's trusted), and this cell might be at the
// top of the notebook (so is executed first), we need to
// allow time for all of the other code cells to load before
// running. Let's give it 5 seconds.
setTimeout(toggler, 5000);
```
Notebooks that have toggling enabled will keep this feature
when you convert them to HTML pages using `jupyter nbconvert`.
I believe that this code could be packaged to create a jupyter extension, but I don't know how, and don't have much interest right now.
## NBConvert Templates
It should be possible to create a
[custom template](https://nbconvert.readthedocs.io/en/latest/customizing.html)
for `jupyter nbconvert` that injects the same code,
or something very like it.
Unfortunately, the documentation on how to do this is not straightforward.
## Hugo
This site is generated using the [Academic](https://github.com/gcushen/hugo-academic) theme for
[Hugo](https://gohugo.io/).
To post Jupyter notebooks, I convert them to `.md` files using the command
`jupyter nbconvert index.ipynb --to markdown --NbConvertApp.output_files_dir=.`,
and following the instructions [here](https://github.com/gcushen/hugo-academic/blob/master/exampleSite/content/post/jupyter/index.md).
I've added the following rule to the site header template for my version of the theme,
located in `/path/to/my_page/themes/academic/layouts/partials/site_head.html`,
just before the `</head>` tag.
```js
{{ if eq $.Params.source "jupyter"}}
<script
src="https://code.jquery.com/jquery-3.4.1.min.js"
crossorigin="anonymous"></script>
<script type="text/javascript" async
src="https://cdnjs.cloudflare.com/ajax/libs/mathjax/2.7.5/MathJax.js?config=TeX-MML-AM_CHTML">
</script>
<script type="text/x-mathjax-config">
MathJax.Hub.Config({
tex2jax: {
inlineMath: [['$','$'], ['\\(','\\)']],
displayMath: [['$$','$$'], ['\[','\]']],
processEscapes: true,
processEnvironments: true,
skipTags: ['script', 'noscript', 'style', 'textarea', 'pre'],
TeX: { equationNumbers: { autoNumber: "AMS" },
extensions: ["AMSmath.js", "AMSsymbols.js"] }
}
});
</script>
<script>
function toggler(){
let btn = $('.language-python').parent()
.prepend('<button>Toggle Code</button>')
.children('button');
btn.on('click', function(e){
let tgt = e.currentTarget;
$(tgt).parent().children('code').toggle()
})
}
$(window).on('load', toggler)
</script>
{{ end }}
```
I then add `source: jupyter` to the metadata of all Jupyter posts.
This means that this code is added to these posts, and only these posts.
Something very similar should work for other static site generators such as Pelican.
# Toggle All
Finally, it would be simple to add a Show/Hide All button at the top of the page, but I haven't got around to doing so yet. Soon, maybe.
| github_jupyter |
# KNN CLASSIFIER
KNN can be used for both classification and regression predictive problems. However, it is more widely used in classification problems in the industry. To evaluate any technique we generally look at 3 important aspects:
1. Ease to interpret output
2. Calculation time
3. Predictive Power
```
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from sklearn import datasets
from sklearn.decomposition import PCA
# import some data to play with
iris = datasets.load_iris()
X = iris.data[:, :2] # we only take the first two features.
y = iris.target
x_min, x_max = X[:, 0].min() - .5, X[:, 0].max() + .5
y_min, y_max = X[:, 1].min() - .5, X[:, 1].max() + .5
plt.figure(2, figsize=(8, 6))
plt.clf()
# Plot the training points
plt.scatter(X[:, 0], X[:, 1], c=y, cmap=plt.cm.Set1,
edgecolor='k')
plt.xlabel('Sepal length')
plt.ylabel('Sepal width')
plt.xlim(x_min, x_max)
plt.ylim(y_min, y_max)
plt.xticks(())
plt.yticks(())
# To getter a better understanding of interaction of the dimensions
# plot the first three PCA dimensions
fig = plt.figure(1, figsize=(8, 6))
ax = Axes3D(fig, elev=-150, azim=110)
X_reduced = PCA(n_components=3).fit_transform(iris.data)
ax.scatter(X_reduced[:, 0], X_reduced[:, 1], X_reduced[:, 2], c=y,
cmap=plt.cm.Set1, edgecolor='k', s=40)
ax.set_title("First three PCA directions")
ax.set_xlabel("1st eigenvector")
ax.w_xaxis.set_ticklabels([])
ax.set_ylabel("2nd eigenvector")
ax.w_yaxis.set_ticklabels([])
ax.set_zlabel("3rd eigenvector")
ax.w_zaxis.set_ticklabels([])
plt.show()
from matplotlib.colors import ListedColormap
import matplotlib.pyplot as plt
def plot_decision_regions(X, y, classifier, test_idx=None, resolution=0.02):
# setup marker generator and color map
markers = ('s', 'x', 'o', '^', 'v')
colors = ('red', 'blue', 'lightgreen', 'gray', 'cyan')
cmap = ListedColormap(colors[:len(np.unique(y))])
# plot the decision surface
x1_min, x1_max = X[:, 0].min() - 1, X[:, 0].max() + 1
x2_min, x2_max = X[:, 1].min() - 1, X[:, 1].max() + 1
xx1, xx2 = np.meshgrid(np.arange(x1_min, x1_max, resolution),
np.arange(x2_min, x2_max, resolution))
Z = classifier.predict(np.array([xx1.ravel(), xx2.ravel()]).T)
Z = Z.reshape(xx1.shape)
plt.contourf(xx1, xx2, Z, alpha=0.3, cmap=cmap)
plt.xlim(xx1.min(), xx1.max())
plt.ylim(xx2.min(), xx2.max())
for idx, cl in enumerate(np.unique(y)):
plt.scatter(x=X[y == cl, 0],
y=X[y == cl, 1],
alpha=0.8,
c=colors[idx],
marker=markers[idx],
label=cl,
edgecolor='black')
# highlight test samples
if test_idx:
# plot all samples
X_test, y_test = X[test_idx, :], y[test_idx]
plt.scatter(X_test[:, 0],
X_test[:, 1],
c='',
edgecolor='black',
alpha=1.0,
linewidth=1,
marker='o',
s=100,
label='test set')
from sklearn.neighbors import KNeighborsClassifier as KNN
import pandas as pd
import numpy as np
for x in range(1,8):
model=KNN(n_neighbors=x)
model.fit(X,y)
plot_decision_regions(X,y,model)
print("number of nearest neighbours")
print(x)
plt.show()
```
| github_jupyter |
# Run a Gene Ontology Enrichment Analysis (GOEA)
We use data from a 2014 Nature paper:
[Computational analysis of cell-to-cell heterogeneity
in single-cell RNA-sequencing data reveals hidden
subpopulations of cells
](http://www.nature.com/nbt/journal/v33/n2/full/nbt.3102.html#methods)
Note: you must have the Python package, **xlrd**, installed to run this example.
Note: To create plots, you must have:
* Python packages: **pyparsing**, **pydot**
* [Graphviz](http://www.graphviz.org/) loaded and your PATH environmental variable pointing to the Graphviz bin directory.
## 1. Download Ontologies and Associations
### 1a. Download Ontologies, if necessary
```
# Get http://geneontology.org/ontology/go-basic.obo
from goatools.base import download_go_basic_obo
obo_fname = download_go_basic_obo()
```
### 1b. Download Associations, if necessary
```
# Get ftp://ftp.ncbi.nlm.nih.gov/gene/DATA/gene2go.gz
from goatools.base import download_ncbi_associations
gene2go = download_ncbi_associations()
```
## 2. Load Ontologies, Associations and Background gene set
### 2a. Load Ontologies
```
from goatools.obo_parser import GODag
obodag = GODag("go-basic.obo")
```
### 2b. Load Associations
```
from __future__ import print_function
from goatools.associations import read_ncbi_gene2go
geneid2gos_mouse = read_ncbi_gene2go("gene2go", taxids=[10090])
print("{N:,} annotated mouse genes".format(N=len(geneid2gos_mouse)))
```
### 2c. Load Background gene set
In this example, the background is all mouse protein-codinge genes
```
from goatools.test_data.genes_NCBI_10090_ProteinCoding import GeneID2nt as GeneID2nt_mus
```
## 3. Initialize a GOEA object
The GOEA object holds the Ontologies, Associations, and background.
Numerous studies can then be run withough needing to re-load the above items.
In this case, we only run one GOEA.
```
from goatools.go_enrichment import GOEnrichmentStudy
goeaobj = GOEnrichmentStudy(
GeneID2nt_mus.keys(), # List of mouse protein-coding genes
geneid2gos_mouse, # geneid/GO associations
obodag, # Ontologies
propagate_counts = False,
alpha = 0.05, # default significance cut-off
methods = ['fdr_bh']) # defult multipletest correction method
```
## 4. Read study genes
~400 genes from the Nature paper supplemental table 4
```
# Data will be stored in this variable
import os
geneid2symbol = {}
# Get xlsx filename where data is stored
ROOT = os.path.dirname(os.getcwd()) # go up 1 level from current working directory
din_xlsx = os.path.join(ROOT, "tests/data/nbt_3102/nbt.3102-S4_GeneIDs.xlsx")
# Read data
if os.path.isfile(din_xlsx):
import xlrd
book = xlrd.open_workbook(din_xlsx)
pg = book.sheet_by_index(0)
for r in range(pg.nrows):
symbol, geneid, pval = [pg.cell_value(r, c) for c in range(pg.ncols)]
if geneid:
geneid2symbol[int(geneid)] = symbol
```
## 5. Run Gene Ontology Enrichment Analysis (GOEA)
You may choose to keep all results or just the significant results. In this example, we choose to keep only the significant results.
```
# 'p_' means "pvalue". 'fdr_bh' is the multipletest method we are currently using.
geneids_study = geneid2symbol.keys()
goea_results_all = goeaobj.run_study(geneids_study)
goea_results_sig = [r for r in goea_results_all if r.p_fdr_bh < 0.05]
```
## 6. Write results to an Excel file and to a text file
```
goeaobj.wr_xlsx("nbt3102.xlsx", goea_results_sig)
goeaobj.wr_txt("nbt3102.txt", goea_results_sig)
```
## 7. Plot all significant GO terms
Plotting all significant GO terms produces a messy spaghetti plot. Such a plot can be useful sometimes because you can open it and zoom and scroll around. But sometimes it is just too messy to be of use.
The **"{NS}"** in **"nbt3102_{NS}.png"** indicates that you will see three plots, one for "biological_process"(BP), "molecular_function"(MF), and "cellular_component"(CC)
```
from goatools.godag_plot import plot_gos, plot_results, plot_goid2goobj
plot_results("nbt3102_{NS}.png", goea_results_sig)
```
### 7a. These plots are likely to messy
The *Cellular Component* plot is the smallest plot...

### 7b. So make a smaller sub-plot
This plot contains GOEA results:
* GO terms colored by P-value:
* pval < 0.005 (light red)
* pval < 0.01 (light orange)
* pval < 0.05 (yellow)
* pval > 0.05 (grey) Study terms that are not statistically significant
* GO terms with study gene counts printed. e.g., "32 genes"
```
# Plot subset starting from these significant GO terms
goid_subset = [
'GO:0003723', # MF D04 RNA binding (32 genes)
'GO:0044822', # MF D05 poly(A) RNA binding (86 genes)
'GO:0003729', # MF D06 mRNA binding (11 genes)
'GO:0019843', # MF D05 rRNA binding (6 genes)
'GO:0003746', # MF D06 translation elongation factor activity (5 genes)
]
plot_gos("nbt3102_MF_RNA_genecnt.png",
goid_subset, # Source GO ids
obodag,
goea_results=goea_results_all) # Use pvals for coloring
```

### 7c. Add study gene Symbols to plot
e.g., *11 genes: Calr, Eef1a1, Pabpc1*
```
plot_gos("nbt3102_MF_RNA_Symbols.png",
goid_subset, # Source GO ids
obodag,
goea_results=goea_results_all, # use pvals for coloring
# We can further configure the plot...
id2symbol=geneid2symbol, # Print study gene Symbols, not Entrez GeneIDs
study_items=6, # Only only 6 gene Symbols max on GO terms
items_p_line=3, # Print 3 genes per line
)
```

Copyright (C) 2016, DV Klopfenstein, H Tang. All rights reserved.
| github_jupyter |
# Flax SST-2 Example
<a href="https://colab.research.google.com/github/google/flax/blob/master/examples/sst2/sst2.ipynb" ><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
Demonstration notebook for
https://github.com/google/flax/tree/master/examples/sst2
**Before you start:** Select Runtime -> Change runtime type -> GPU.
The **Flax Notebook Workflow**:
1. Run the entire notebook end-to-end and check out the outputs.
- This will open Python files in the right-hand editor!
- You'll be able to interactively explore metrics in TensorBoard.
2. Change `config` and train for different hyperparameters. Check out the
updated TensorBoard plots.
3. Update the code in `train.py`. Thanks to `%autoreload`, any changes you
make in the file will automatically appear in the notebook. Some ideas to
get you started:
- Change the model.
- Log some per-batch metrics during training.
- Add new hyperparameters to `configs/default.py` and use them in
`train.py`.
4. At any time, feel free to paste code from `train.py` into the notebook
and modify it directly there!
## Setup
```
example_directory = 'examples/sst2'
editor_relpaths = ('configs/default.py', 'train.py', 'models.py')
# (If you run this code in Jupyter[lab], then you're already in the
# example directory and nothing needs to be done.)
#@markdown **Fetch newest Flax, copy example code**
#@markdown
#@markdown **If you select no** below, then the files will be stored on the
#@markdown *ephemeral* Colab VM. **After some time of inactivity, this VM will
#@markdown be restarted an any changes are lost**.
#@markdown
#@markdown **If you select yes** below, then you will be asked for your
#@markdown credentials to mount your personal Google Drive. In this case, all
#@markdown changes you make will be *persisted*, and even if you re-run the
#@markdown Colab later on, the files will still be the same (you can of course
#@markdown remove directories inside your Drive's `flax/` root if you want to
#@markdown manually revert these files).
if 'google.colab' in str(get_ipython()):
import os
os.chdir('/content')
# Download Flax repo from Github.
if not os.path.isdir('flaxrepo'):
!git clone --depth=1 https://github.com/google/flax flaxrepo
# Copy example files & change directory.
mount_gdrive = 'no' #@param ['yes', 'no']
if mount_gdrive == 'yes':
DISCLAIMER = 'Note: Editing in your Google Drive, changes will persist.'
from google.colab import drive
drive.mount('/content/gdrive')
example_root_path = f'/content/gdrive/My Drive/flax/{example_directory}'
else:
DISCLAIMER = 'WARNING: Editing in VM - changes lost after reboot!!'
example_root_path = f'/content/{example_directory}'
from IPython import display
display.display(display.HTML(
f'<h1 style="color:red;" class="blink">{DISCLAIMER}</h1>'))
if not os.path.isdir(example_root_path):
os.makedirs(example_root_path)
!cp -r flaxrepo/$example_directory/* "$example_root_path"
os.chdir(example_root_path)
from google.colab import files
for relpath in editor_relpaths:
s = open(f'{example_root_path}/{relpath}').read()
open(f'{example_root_path}/{relpath}', 'w').write(
f'## {DISCLAIMER}\n' + '#' * (len(DISCLAIMER) + 3) + '\n\n' + s)
files.view(f'{example_root_path}/{relpath}')
# Note: In Colab, above cell changed the working directory.
!pwd
# Install SST-2 dependencies.
!pip install -q -r requirements.txt
```
## Imports / Helpers
```
# If you want to use TPU instead of GPU, you need to run this to make it work.
try:
import jax.tools.colab_tpu
jax.tools.colab_tpu.setup_tpu()
except KeyError:
print('\n### NO TPU CONNECTED - USING CPU or GPU ###\n')
import os
os.environ['XLA_FLAGS'] = '--xla_force_host_platform_device_count=8'
jax.devices()
from absl import logging
import flax
import jax.numpy as jnp
import numpy as np
import tensorflow as tf
import tensorflow_datasets as tfds
import time
logging.set_verbosity(logging.INFO)
# Make sure the GPU is for JAX, not for TF.
tf.config.experimental.set_visible_devices([], 'GPU')
# Local imports from current directory - auto reload.
# Any changes you make to train.py will appear automatically.
%load_ext autoreload
%autoreload 2
import train
import models
import vocabulary
import input_pipeline
from configs import default as config_lib
config = config_lib.get_config()
```
## Dataset
```
# Get datasets.
# If you get an error you need to install tensorflow_datasets from Github.
train_dataset = input_pipeline.TextDataset(split='train')
eval_dataset = input_pipeline.TextDataset(split='validation')
```
## Training
```
# Get a live update during training - use the "refresh" button!
# (In Jupyter[lab] start "tensorboard" in the local directory instead.)
if 'google.colab' in str(get_ipython()):
%load_ext tensorboard
%tensorboard --logdir=.
config.num_epochs = 10
model_name = 'bilstm'
start_time = time.time()
optimizer = train.train_and_evaluate(config, workdir=f'./models/{model_name}')
logging.info('Walltime: %f s', time.time() - start_time)
if 'google.colab' in str(get_ipython()):
#@markdown You can upload the training results directly to https://tensorboard.dev
#@markdown
#@markdown Note that everbody with the link will be able to see the data.
upload_data = 'yes' #@param ['yes', 'no']
if upload_data == 'yes':
!tensorboard dev upload --one_shot --logdir ./models --name 'Flax examples/mnist'
```
| github_jupyter |
# Character level language model - Dinosaurus land
Welcome to Dinosaurus Island! 65 million years ago, dinosaurs existed, and in this assignment they are back. You are in charge of a special task. Leading biology researchers are creating new breeds of dinosaurs and bringing them to life on earth, and your job is to give names to these dinosaurs. If a dinosaur does not like its name, it might go beserk, so choose wisely!
<table>
<td>
<img src="images/dino.jpg" style="width:250;height:300px;">
</td>
</table>
Luckily you have learned some deep learning and you will use it to save the day. Your assistant has collected a list of all the dinosaur names they could find, and compiled them into this [dataset](dinos.txt). (Feel free to take a look by clicking the previous link.) To create new dinosaur names, you will build a character level language model to generate new names. Your algorithm will learn the different name patterns, and randomly generate new names. Hopefully this algorithm will keep you and your team safe from the dinosaurs' wrath!
By completing this assignment you will learn:
- How to store text data for processing using an RNN
- How to synthesize data, by sampling predictions at each time step and passing it to the next RNN-cell unit
- How to build a character-level text generation recurrent neural network
- Why clipping the gradients is important
We will begin by loading in some functions that we have provided for you in `rnn_utils`. Specifically, you have access to functions such as `rnn_forward` and `rnn_backward` which are equivalent to those you've implemented in the previous assignment.
```
import numpy as np
from utils import *
import random
```
## 1 - Problem Statement
### 1.1 - Dataset and Preprocessing
Run the following cell to read the dataset of dinosaur names, create a list of unique characters (such as a-z), and compute the dataset and vocabulary size.
```
data = open('dinos.txt', 'r').read()
data= data.lower()
chars = list(set(data))
data_size, vocab_size = len(data), len(chars)
print('There are %d total characters and %d unique characters in your data.' % (data_size, vocab_size))
```
The characters are a-z (26 characters) plus the "\n" (or newline character), which in this assignment plays a role similar to the `<EOS>` (or "End of sentence") token we had discussed in lecture, only here it indicates the end of the dinosaur name rather than the end of a sentence. In the cell below, we create a python dictionary (i.e., a hash table) to map each character to an index from 0-26. We also create a second python dictionary that maps each index back to the corresponding character character. This will help you figure out what index corresponds to what character in the probability distribution output of the softmax layer. Below, `char_to_ix` and `ix_to_char` are the python dictionaries.
```
char_to_ix = { ch:i for i,ch in enumerate(sorted(chars)) }
ix_to_char = { i:ch for i,ch in enumerate(sorted(chars)) }
print(ix_to_char)
```
### 1.2 - Overview of the model
Your model will have the following structure:
- Initialize parameters
- Run the optimization loop
- Forward propagation to compute the loss function
- Backward propagation to compute the gradients with respect to the loss function
- Clip the gradients to avoid exploding gradients
- Using the gradients, update your parameter with the gradient descent update rule.
- Return the learned parameters
<img src="images/rnn.png" style="width:450;height:300px;">
<caption><center> **Figure 1**: Recurrent Neural Network, similar to what you had built in the previous notebook "Building a RNN - Step by Step". </center></caption>
At each time-step, the RNN tries to predict what is the next character given the previous characters. The dataset $X = (x^{\langle 1 \rangle}, x^{\langle 2 \rangle}, ..., x^{\langle T_x \rangle})$ is a list of characters in the training set, while $Y = (y^{\langle 1 \rangle}, y^{\langle 2 \rangle}, ..., y^{\langle T_x \rangle})$ is such that at every time-step $t$, we have $y^{\langle t \rangle} = x^{\langle t+1 \rangle}$.
## 2 - Building blocks of the model
In this part, you will build two important blocks of the overall model:
- Gradient clipping: to avoid exploding gradients
- Sampling: a technique used to generate characters
You will then apply these two functions to build the model.
### 2.1 - Clipping the gradients in the optimization loop
In this section you will implement the `clip` function that you will call inside of your optimization loop. Recall that your overall loop structure usually consists of a forward pass, a cost computation, a backward pass, and a parameter update. Before updating the parameters, you will perform gradient clipping when needed to make sure that your gradients are not "exploding," meaning taking on overly large values.
In the exercise below, you will implement a function `clip` that takes in a dictionary of gradients and returns a clipped version of gradients if needed. There are different ways to clip gradients; we will use a simple element-wise clipping procedure, in which every element of the gradient vector is clipped to lie between some range [-N, N]. More generally, you will provide a `maxValue` (say 10). In this example, if any component of the gradient vector is greater than 10, it would be set to 10; and if any component of the gradient vector is less than -10, it would be set to -10. If it is between -10 and 10, it is left alone.
<img src="images/clip.png" style="width:400;height:150px;">
<caption><center> **Figure 2**: Visualization of gradient descent with and without gradient clipping, in a case where the network is running into slight "exploding gradient" problems. </center></caption>
**Exercise**: Implement the function below to return the clipped gradients of your dictionary `gradients`. Your function takes in a maximum threshold and returns the clipped versions of your gradients. You can check out this [hint](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.clip.html) for examples of how to clip in numpy. You will need to use the argument `out = ...`.
```
### GRADED FUNCTION: clip
def clip(gradients, maxValue):
'''
Clips the gradients' values between minimum and maximum.
Arguments:
gradients -- a dictionary containing the gradients "dWaa", "dWax", "dWya", "db", "dby"
maxValue -- everything above this number is set to this number, and everything less than -maxValue is set to -maxValue
Returns:
gradients -- a dictionary with the clipped gradients.
'''
dWaa, dWax, dWya, db, dby = gradients['dWaa'], gradients['dWax'], gradients['dWya'], gradients['db'], gradients['dby']
### START CODE HERE ###
# clip to mitigate exploding gradients, loop over [dWax, dWaa, dWya, db, dby]. (≈2 lines)
for gradient in [dWax, dWaa, dWya, db, dby]:
gradient = np.clip(gradient, -maxValue, maxValue, out = gradient)
### END CODE HERE ###
gradients = {"dWaa": dWaa, "dWax": dWax, "dWya": dWya, "db": db, "dby": dby}
return gradients
np.random.seed(3)
dWax = np.random.randn(5,3)*10
dWaa = np.random.randn(5,5)*10
dWya = np.random.randn(2,5)*10
db = np.random.randn(5,1)*10
dby = np.random.randn(2,1)*10
gradients = {"dWax": dWax, "dWaa": dWaa, "dWya": dWya, "db": db, "dby": dby}
gradients = clip(gradients, 10)
print("gradients[\"dWaa\"][1][2] =", gradients["dWaa"][1][2])
print("gradients[\"dWax\"][3][1] =", gradients["dWax"][3][1])
print("gradients[\"dWya\"][1][2] =", gradients["dWya"][1][2])
print("gradients[\"db\"][4] =", gradients["db"][4])
print("gradients[\"dby\"][1] =", gradients["dby"][1])
```
** Expected output:**
<table>
<tr>
<td>
**gradients["dWaa"][1][2] **
</td>
<td>
10.0
</td>
</tr>
<tr>
<td>
**gradients["dWax"][3][1]**
</td>
<td>
-10.0
</td>
</td>
</tr>
<tr>
<td>
**gradients["dWya"][1][2]**
</td>
<td>
0.29713815361
</td>
</tr>
<tr>
<td>
**gradients["db"][4]**
</td>
<td>
[ 10.]
</td>
</tr>
<tr>
<td>
**gradients["dby"][1]**
</td>
<td>
[ 8.45833407]
</td>
</tr>
</table>
### 2.2 - Sampling
Now assume that your model is trained. You would like to generate new text (characters). The process of generation is explained in the picture below:
<img src="images/dinos3.png" style="width:500;height:300px;">
<caption><center> **Figure 3**: In this picture, we assume the model is already trained. We pass in $x^{\langle 1\rangle} = \vec{0}$ at the first time step, and have the network then sample one character at a time. </center></caption>
**Exercise**: Implement the `sample` function below to sample characters. You need to carry out 4 steps:
- **Step 1**: Pass the network the first "dummy" input $x^{\langle 1 \rangle} = \vec{0}$ (the vector of zeros). This is the default input before we've generated any characters. We also set $a^{\langle 0 \rangle} = \vec{0}$
- **Step 2**: Run one step of forward propagation to get $a^{\langle 1 \rangle}$ and $\hat{y}^{\langle 1 \rangle}$. Here are the equations:
$$ a^{\langle t+1 \rangle} = \tanh(W_{ax} x^{\langle t \rangle } + W_{aa} a^{\langle t \rangle } + b)\tag{1}$$
$$ z^{\langle t + 1 \rangle } = W_{ya} a^{\langle t + 1 \rangle } + b_y \tag{2}$$
$$ \hat{y}^{\langle t+1 \rangle } = softmax(z^{\langle t + 1 \rangle })\tag{3}$$
Note that $\hat{y}^{\langle t+1 \rangle }$ is a (softmax) probability vector (its entries are between 0 and 1 and sum to 1). $\hat{y}^{\langle t+1 \rangle}_i$ represents the probability that the character indexed by "i" is the next character. We have provided a `softmax()` function that you can use.
- **Step 3**: Carry out sampling: Pick the next character's index according to the probability distribution specified by $\hat{y}^{\langle t+1 \rangle }$. This means that if $\hat{y}^{\langle t+1 \rangle }_i = 0.16$, you will pick the index "i" with 16% probability. To implement it, you can use [`np.random.choice`](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.random.choice.html).
Here is an example of how to use `np.random.choice()`:
```python
np.random.seed(0)
p = np.array([0.1, 0.0, 0.7, 0.2])
index = np.random.choice([0, 1, 2, 3], p = p.ravel())
```
This means that you will pick the `index` according to the distribution:
$P(index = 0) = 0.1, P(index = 1) = 0.0, P(index = 2) = 0.7, P(index = 3) = 0.2$.
- **Step 4**: The last step to implement in `sample()` is to overwrite the variable `x`, which currently stores $x^{\langle t \rangle }$, with the value of $x^{\langle t + 1 \rangle }$. You will represent $x^{\langle t + 1 \rangle }$ by creating a one-hot vector corresponding to the character you've chosen as your prediction. You will then forward propagate $x^{\langle t + 1 \rangle }$ in Step 1 and keep repeating the process until you get a "\n" character, indicating you've reached the end of the dinosaur name.
```
# GRADED FUNCTION: sample
def sample(parameters, char_to_ix, seed):
"""
Sample a sequence of characters according to a sequence of probability distributions output of the RNN
Arguments:
parameters -- python dictionary containing the parameters Waa, Wax, Wya, by, and b.
char_to_ix -- python dictionary mapping each character to an index.
seed -- used for grading purposes. Do not worry about it.
Returns:
indices -- a list of length n containing the indices of the sampled characters.
"""
# Retrieve parameters and relevant shapes from "parameters" dictionary
Waa, Wax, Wya, by, b = parameters['Waa'], parameters['Wax'], parameters['Wya'], parameters['by'], parameters['b']
vocab_size = by.shape[0]
n_a = Waa.shape[1]
### START CODE HERE ###
# Step 1: Create the one-hot vector x for the first character (initializing the sequence generation). (≈1 line)
x = np.zeros((vocab_size,1))
# Step 1': Initialize a_prev as zeros (≈1 line)
a_prev = np.zeros((n_a,1))
# Create an empty list of indices, this is the list which will contain the list of indices of the characters to generate (≈1 line)
indices = []
# Idx is a flag to detect a newline character, we initialize it to -1
idx = -1
# Loop over time-steps t. At each time-step, sample a character from a probability distribution and append
# its index to "indices". We'll stop if we reach 50 characters (which should be very unlikely with a well
# trained model), which helps debugging and prevents entering an infinite loop.
counter = 0
newline_character = char_to_ix['\n']
while (idx != newline_character and counter != 50):
# Step 2: Forward propagate x using the equations (1), (2) and (3)
a = np.tanh(np.dot(Wax,x) + np.dot(Waa,a_prev) + b)
z = np.dot(Wya,a) + by
y = softmax(z)
# for grading purposes
np.random.seed(counter+seed)
# Step 3: Sample the index of a character within the vocabulary from the probability distribution y
idx = np.random.choice(list(range(vocab_size)), p = y.ravel())
# Append the index to "indices"
indices.append(idx)
# Step 4: Overwrite the input character as the one corresponding to the sampled index.
x = np.zeros((vocab_size, 1))
x[idx] = 1
# Update "a_prev" to be "a"
a_prev = a
# for grading purposes
seed += 1
counter +=1
### END CODE HERE ###
if (counter == 50):
indices.append(char_to_ix['\n'])
return indices
np.random.seed(2)
_, n_a = 20, 100
Wax, Waa, Wya = np.random.randn(n_a, vocab_size), np.random.randn(n_a, n_a), np.random.randn(vocab_size, n_a)
b, by = np.random.randn(n_a, 1), np.random.randn(vocab_size, 1)
parameters = {"Wax": Wax, "Waa": Waa, "Wya": Wya, "b": b, "by": by}
indices = sample(parameters, char_to_ix, 0)
print("Sampling:")
print("list of sampled indices:", indices)
print("list of sampled characters:", [ix_to_char[i] for i in indices])
```
** Expected output:**
<table>
<tr>
<td>
**list of sampled indices:**
</td>
<td>
[12, 17, 24, 14, 13, 9, 10, 22, 24, 6, 13, 11, 12, 6, 21, 15, 21, 14, 3, 2, 1, 21, 18, 24, <br>
7, 25, 6, 25, 18, 10, 16, 2, 3, 8, 15, 12, 11, 7, 1, 12, 10, 2, 7, 7, 11, 5, 6, 12, 25, 0, 0]
</td>
</tr><tr>
<td>
**list of sampled characters:**
</td>
<td>
['l', 'q', 'x', 'n', 'm', 'i', 'j', 'v', 'x', 'f', 'm', 'k', 'l', 'f', 'u', 'o', <br>
'u', 'n', 'c', 'b', 'a', 'u', 'r', 'x', 'g', 'y', 'f', 'y', 'r', 'j', 'p', 'b', 'c', 'h', 'o', <br>
'l', 'k', 'g', 'a', 'l', 'j', 'b', 'g', 'g', 'k', 'e', 'f', 'l', 'y', '\n', '\n']
</td>
</tr>
</table>
## 3 - Building the language model
It is time to build the character-level language model for text generation.
### 3.1 - Gradient descent
In this section you will implement a function performing one step of stochastic gradient descent (with clipped gradients). You will go through the training examples one at a time, so the optimization algorithm will be stochastic gradient descent. As a reminder, here are the steps of a common optimization loop for an RNN:
- Forward propagate through the RNN to compute the loss
- Backward propagate through time to compute the gradients of the loss with respect to the parameters
- Clip the gradients if necessary
- Update your parameters using gradient descent
**Exercise**: Implement this optimization process (one step of stochastic gradient descent).
We provide you with the following functions:
```python
def rnn_forward(X, Y, a_prev, parameters):
""" Performs the forward propagation through the RNN and computes the cross-entropy loss.
It returns the loss' value as well as a "cache" storing values to be used in the backpropagation."""
....
return loss, cache
def rnn_backward(X, Y, parameters, cache):
""" Performs the backward propagation through time to compute the gradients of the loss with respect
to the parameters. It returns also all the hidden states."""
...
return gradients, a
def update_parameters(parameters, gradients, learning_rate):
""" Updates parameters using the Gradient Descent Update Rule."""
...
return parameters
```
```
# GRADED FUNCTION: optimize
def optimize(X, Y, a_prev, parameters, learning_rate = 0.01):
"""
Execute one step of the optimization to train the model.
Arguments:
X -- list of integers, where each integer is a number that maps to a character in the vocabulary.
Y -- list of integers, exactly the same as X but shifted one index to the left.
a_prev -- previous hidden state.
parameters -- python dictionary containing:
Wax -- Weight matrix multiplying the input, numpy array of shape (n_a, n_x)
Waa -- Weight matrix multiplying the hidden state, numpy array of shape (n_a, n_a)
Wya -- Weight matrix relating the hidden-state to the output, numpy array of shape (n_y, n_a)
b -- Bias, numpy array of shape (n_a, 1)
by -- Bias relating the hidden-state to the output, numpy array of shape (n_y, 1)
learning_rate -- learning rate for the model.
Returns:
loss -- value of the loss function (cross-entropy)
gradients -- python dictionary containing:
dWax -- Gradients of input-to-hidden weights, of shape (n_a, n_x)
dWaa -- Gradients of hidden-to-hidden weights, of shape (n_a, n_a)
dWya -- Gradients of hidden-to-output weights, of shape (n_y, n_a)
db -- Gradients of bias vector, of shape (n_a, 1)
dby -- Gradients of output bias vector, of shape (n_y, 1)
a[len(X)-1] -- the last hidden state, of shape (n_a, 1)
"""
### START CODE HERE ###
# Forward propagate through time (≈1 line)
loss, cache = rnn_forward(X,Y,a_prev,parameters)
# Backpropagate through time (≈1 line)
gradients, a = rnn_backward(X,Y,parameters, cache)
# Clip your gradients between -5 (min) and 5 (max) (≈1 line)
gradients = clip(gradients,5)
# Update parameters (≈1 line)
parameters = update_parameters(parameters,gradients,learning_rate)
### END CODE HERE ###
return loss, gradients, a[len(X)-1]
np.random.seed(1)
vocab_size, n_a = 27, 100
a_prev = np.random.randn(n_a, 1)
Wax, Waa, Wya = np.random.randn(n_a, vocab_size), np.random.randn(n_a, n_a), np.random.randn(vocab_size, n_a)
b, by = np.random.randn(n_a, 1), np.random.randn(vocab_size, 1)
parameters = {"Wax": Wax, "Waa": Waa, "Wya": Wya, "b": b, "by": by}
X = [12,3,5,11,22,3]
Y = [4,14,11,22,25, 26]
loss, gradients, a_last = optimize(X, Y, a_prev, parameters, learning_rate = 0.01)
print("Loss =", loss)
print("gradients[\"dWaa\"][1][2] =", gradients["dWaa"][1][2])
print("np.argmax(gradients[\"dWax\"]) =", np.argmax(gradients["dWax"]))
print("gradients[\"dWya\"][1][2] =", gradients["dWya"][1][2])
print("gradients[\"db\"][4] =", gradients["db"][4])
print("gradients[\"dby\"][1] =", gradients["dby"][1])
print("a_last[4] =", a_last[4])
```
** Expected output:**
<table>
<tr>
<td>
**Loss **
</td>
<td>
126.503975722
</td>
</tr>
<tr>
<td>
**gradients["dWaa"][1][2]**
</td>
<td>
0.194709315347
</td>
<tr>
<td>
**np.argmax(gradients["dWax"])**
</td>
<td> 93
</td>
</tr>
<tr>
<td>
**gradients["dWya"][1][2]**
</td>
<td> -0.007773876032
</td>
</tr>
<tr>
<td>
**gradients["db"][4]**
</td>
<td> [-0.06809825]
</td>
</tr>
<tr>
<td>
**gradients["dby"][1]**
</td>
<td>[ 0.01538192]
</td>
</tr>
<tr>
<td>
**a_last[4]**
</td>
<td> [-1.]
</td>
</tr>
</table>
### 3.2 - Training the model
Given the dataset of dinosaur names, we use each line of the dataset (one name) as one training example. Every 100 steps of stochastic gradient descent, you will sample 10 randomly chosen names to see how the algorithm is doing. Remember to shuffle the dataset, so that stochastic gradient descent visits the examples in random order.
**Exercise**: Follow the instructions and implement `model()`. When `examples[index]` contains one dinosaur name (string), to create an example (X, Y), you can use this:
```python
index = j % len(examples)
X = [None] + [char_to_ix[ch] for ch in examples[index]]
Y = X[1:] + [char_to_ix["\n"]]
```
Note that we use: `index= j % len(examples)`, where `j = 1....num_iterations`, to make sure that `examples[index]` is always a valid statement (`index` is smaller than `len(examples)`).
The first entry of `X` being `None` will be interpreted by `rnn_forward()` as setting $x^{\langle 0 \rangle} = \vec{0}$. Further, this ensures that `Y` is equal to `X` but shifted one step to the left, and with an additional "\n" appended to signify the end of the dinosaur name.
```
# GRADED FUNCTION: model
def model(data, ix_to_char, char_to_ix, num_iterations = 35000, n_a = 50, dino_names = 7, vocab_size = 27):
"""
Trains the model and generates dinosaur names.
Arguments:
data -- text corpus
ix_to_char -- dictionary that maps the index to a character
char_to_ix -- dictionary that maps a character to an index
num_iterations -- number of iterations to train the model for
n_a -- number of units of the RNN cell
dino_names -- number of dinosaur names you want to sample at each iteration.
vocab_size -- number of unique characters found in the text, size of the vocabulary
Returns:
parameters -- learned parameters
"""
# Retrieve n_x and n_y from vocab_size
n_x, n_y = vocab_size, vocab_size
# Initialize parameters
parameters = initialize_parameters(n_a, n_x, n_y)
# Initialize loss (this is required because we want to smooth our loss, don't worry about it)
loss = get_initial_loss(vocab_size, dino_names)
# Build list of all dinosaur names (training examples).
with open("dinos.txt") as f:
examples = f.readlines()
examples = [x.lower().strip() for x in examples]
# Shuffle list of all dinosaur names
np.random.seed(0)
np.random.shuffle(examples)
# Initialize the hidden state of your LSTM
a_prev = np.zeros((n_a, 1))
# Optimization loop
for j in range(num_iterations):
### START CODE HERE ###
# Use the hint above to define one training example (X,Y) (≈ 2 lines)
index = j % len(examples)
X = [None] + [char_to_ix[ch] for ch in examples[index]]
Y = X[1:] + [char_to_ix['\n']]
# Perform one optimization step: Forward-prop -> Backward-prop -> Clip -> Update parameters
# Choose a learning rate of 0.01
curr_loss, gradients, a_prev = optimize(X,Y,a_prev,parameters)
### END CODE HERE ###
# Use a latency trick to keep the loss smooth. It happens here to accelerate the training.
loss = smooth(loss, curr_loss)
# Every 2000 Iteration, generate "n" characters thanks to sample() to check if the model is learning properly
if j % 2000 == 0:
print('Iteration: %d, Loss: %f' % (j, loss) + '\n')
# The number of dinosaur names to print
seed = 0
for name in range(dino_names):
# Sample indices and print them
sampled_indices = sample(parameters, char_to_ix, seed)
print_sample(sampled_indices, ix_to_char)
seed += 1 # To get the same result for grading purposed, increment the seed by one.
print('\n')
return parameters
```
Run the following cell, you should observe your model outputting random-looking characters at the first iteration. After a few thousand iterations, your model should learn to generate reasonable-looking names.
```
parameters = model(data, ix_to_char, char_to_ix)
```
## Conclusion
You can see that your algorithm has started to generate plausible dinosaur names towards the end of the training. At first, it was generating random characters, but towards the end you could see dinosaur names with cool endings. Feel free to run the algorithm even longer and play with hyperparameters to see if you can get even better results. Our implemetation generated some really cool names like `maconucon`, `marloralus` and `macingsersaurus`. Your model hopefully also learned that dinosaur names tend to end in `saurus`, `don`, `aura`, `tor`, etc.
If your model generates some non-cool names, don't blame the model entirely--not all actual dinosaur names sound cool. (For example, `dromaeosauroides` is an actual dinosaur name and is in the training set.) But this model should give you a set of candidates from which you can pick the coolest!
This assignment had used a relatively small dataset, so that you could train an RNN quickly on a CPU. Training a model of the english language requires a much bigger dataset, and usually needs much more computation, and could run for many hours on GPUs. We ran our dinosaur name for quite some time, and so far our favoriate name is the great, undefeatable, and fierce: Mangosaurus!
<img src="images/mangosaurus.jpeg" style="width:250;height:300px;">
## 4 - Writing like Shakespeare
The rest of this notebook is optional and is not graded, but we hope you'll do it anyway since it's quite fun and informative.
A similar (but more complicated) task is to generate Shakespeare poems. Instead of learning from a dataset of Dinosaur names you can use a collection of Shakespearian poems. Using LSTM cells, you can learn longer term dependencies that span many characters in the text--e.g., where a character appearing somewhere a sequence can influence what should be a different character much much later in ths sequence. These long term dependencies were less important with dinosaur names, since the names were quite short.
<img src="images/shakespeare.jpg" style="width:500;height:400px;">
<caption><center> Let's become poets! </center></caption>
We have implemented a Shakespeare poem generator with Keras. Run the following cell to load the required packages and models. This may take a few minutes.
```
from __future__ import print_function
from keras.callbacks import LambdaCallback
from keras.models import Model, load_model, Sequential
from keras.layers import Dense, Activation, Dropout, Input, Masking
from keras.layers import LSTM
from keras.utils.data_utils import get_file
from keras.preprocessing.sequence import pad_sequences
from shakespeare_utils import *
import sys
import io
```
To save you some time, we have already trained a model for ~1000 epochs on a collection of Shakespearian poems called [*"The Sonnets"*](shakespeare.txt).
Let's train the model for one more epoch. When it finishes training for an epoch---this will also take a few minutes---you can run `generate_output`, which will prompt asking you for an input (`<`40 characters). The poem will start with your sentence, and our RNN-Shakespeare will complete the rest of the poem for you! For example, try "Forsooth this maketh no sense " (don't enter the quotation marks). Depending on whether you include the space at the end, your results might also differ--try it both ways, and try other inputs as well.
```
print_callback = LambdaCallback(on_epoch_end=on_epoch_end)
model.fit(x, y, batch_size=128, epochs=1, callbacks=[print_callback])
# Run this cell to try with different inputs without having to re-train the model
generate_output()
```
The RNN-Shakespeare model is very similar to the one you have built for dinosaur names. The only major differences are:
- LSTMs instead of the basic RNN to capture longer-range dependencies
- The model is a deeper, stacked LSTM model (2 layer)
- Using Keras instead of python to simplify the code
If you want to learn more, you can also check out the Keras Team's text generation implementation on GitHub: https://github.com/keras-team/keras/blob/master/examples/lstm_text_generation.py.
Congratulations on finishing this notebook!
**References**:
- This exercise took inspiration from Andrej Karpathy's implementation: https://gist.github.com/karpathy/d4dee566867f8291f086. To learn more about text generation, also check out Karpathy's [blog post](http://karpathy.github.io/2015/05/21/rnn-effectiveness/).
- For the Shakespearian poem generator, our implementation was based on the implementation of an LSTM text generator by the Keras team: https://github.com/keras-team/keras/blob/master/examples/lstm_text_generation.py
| github_jupyter |
### *IPCC SR15 scenario assessment*
<img style="float: right; height: 80px; padding-left: 20px;" src="../_static/IIASA_logo.png">
<img style="float: right; height: 80px;" src="../_static/IAMC_logo.jpg">
# Descriptive statistics of the primary energy development
This notebook computes indicators and diagnostics of the primary-energy timeseries by fuel
for **Table 2.6** in the IPCC's _"Special Report on Global Warming of 1.5°C"_.
The scenario data used in this analysis can be accessed and downloaded at [https://data.ene.iiasa.ac.at/iamc-1.5c-explorer](https://data.ene.iiasa.ac.at/iamc-1.5c-explorer).
## Load `pyam` package and other dependencies
```
import pandas as pd
import numpy as np
import warnings
import io
import itertools
import yaml
import math
import matplotlib.pyplot as plt
plt.style.use('style_sr15.mplstyle')
%matplotlib inline
import pyam
```
## Import scenario data, categorization and specifications files
The metadata file must be generated from the notebook `sr15_2.0_categories_indicators` included in this repository.
If the snapshot file has been updated, make sure that you rerun the categorization notebook.
The last cell of this section loads and assigns a number of auxiliary lists as defined in the categorization notebook.
```
sr1p5 = pyam.IamDataFrame(data='../data/iamc15_scenario_data_world_r1.1.xlsx')
sr1p5.load_metadata('sr15_metadata_indicators.xlsx')
with open("sr15_specs.yaml", 'r') as stream:
specs = yaml.load(stream, Loader=yaml.FullLoader)
rc = pyam.run_control()
for item in specs.pop('run_control').items():
rc.update({item[0]: item[1]})
cats = specs.pop('cats')
cats_15 = specs.pop('cats_15')
cats_15_no_lo = specs.pop('cats_15_no_lo')
marker= specs.pop('marker')
```
## Downselect scenario ensemble to categories of interest for this assessment
```
years = [2020, 2030, 2050]
df = sr1p5.filter(category=cats_15, year=years)
```
## Initialize a `pyam.Statistics` instance
```
stats = pyam.Statistics(df=df,
filters=[('all 1.5', {}),
('no & lo os 1.5', {'category': cats_15_no_lo}),
('hi os 1.5', {'category': ['1.5C high overshoot']})
], rows=True)
header='Primary energy supply (EJ)'
header_share='Share in primary energy (%)'
header_growth='Growth (factor)'
statistics_settings = dict(
header=header,
header_share=header_share,
header_growth= header_growth,
growth_year=2050,
base_year=2020
)
def add_statistics(data, base, row, growth_year, base_year,
header, header_share, header_growth):
stats.add(data, header=header, row=row)
if base is not None:
stats.add(data / base * 100, header=header_share, row=row)
stats.add(data[growth_year] / data[base_year] - 1,
header=header_growth, row=row,
subheader='{}-{}'.format(base_year, growth_year))
```
## Extract primary energy timeseries data and add summary statistics
```
pe = df.filter(variable='Primary Energy').timeseries()
pe.index = pe.index.droplevel([2, 3, 4])
add_statistics(pe, None, 'total primary', **statistics_settings)
```
## Compute share of renewables by various types in primary energy
Only use scenarios for this indicator that report both biomass and the aggregate non-biomass timeseries - otherwise, the share would be distorted.
### All renewables (biomass and non-biomass)
```
df_pe_res = df.filter()
df_pe_res.require_variable('Primary Energy|Non-Biomass Renewables', exclude_on_fail=True)
df_pe_res.require_variable('Primary Energy|Biomass', exclude_on_fail=True)
df_pe_res.filter(exclude=False, inplace=True)
res = (
df_pe_res.filter(variable=['Primary Energy|Biomass', 'Primary Energy|Non-Biomass Renewables'])
.timeseries()
.groupby(['model', 'scenario']).sum()
)
add_statistics(res, pe, 'renewables', **statistics_settings)
```
### Biomass
```
res_bio = (
df.filter(variable=['Primary Energy|Biomass'])
.timeseries()
.groupby(['model', 'scenario']).sum()
)
add_statistics(res_bio, pe, 'biomass', **statistics_settings)
```
### Non-biomass renewables
```
res_non_bio = (
df.filter(variable=['Primary Energy|Non-Biomass Renewables'])
.timeseries()
.groupby(['model', 'scenario']).sum()
)
add_statistics(res_non_bio, pe, 'non-biomass', **statistics_settings)
```
### Renewable energy from wind and solar
As above, verify that scenarios report values for both 'Wind' and 'Solar'
```
df_win_sol = df.filter()
df_win_sol.require_variable('Primary Energy|Wind', exclude_on_fail=True)
df_win_sol.require_variable('Primary Energy|Solar', exclude_on_fail=True)
df_win_sol.filter(exclude=False, inplace=True)
win_sol = (
df_win_sol.filter(variable=['Primary Energy|Wind',
'Primary Energy|Solar'])
.timeseries()
.groupby(['model', 'scenario']).sum()
)
add_statistics(win_sol, pe, 'wind & solar', **statistics_settings)
```
## Compute share of nuclear in primary energy
```
nuc = (
df.filter(variable=['Primary Energy|Nuclear'])
.timeseries()
.groupby(['model', 'scenario']).sum()
)
add_statistics(nuc, pe, 'nuclear', **statistics_settings)
```
## Compute share of fossil in primary energy
```
fossil = (
df.filter(variable=['Primary Energy|Coal', 'Primary Energy|Gas', 'Primary Energy|Oil'])
.timeseries()
.groupby(['model', 'scenario']).sum()
)
add_statistics(fossil, pe, 'fossil', **statistics_settings)
coal = (
df.filter(variable=['Primary Energy|Coal'])
.timeseries()
.groupby(['model', 'scenario']).sum()
)
add_statistics(coal, pe, 'coal', **statistics_settings)
gas = (
df.filter(variable=['Primary Energy|Gas'])
.timeseries()
.groupby(['model', 'scenario']).sum()
)
add_statistics(gas, pe, 'gas', **statistics_settings)
oil = (
df.filter(variable=['Primary Energy|Oil'])
.timeseries()
.groupby(['model', 'scenario']).sum()
)
add_statistics(oil, pe, 'oil', **statistics_settings)
```
## Display and export summary statistics for all 1.5C scenarios to `xlsx`
```
summary = (
stats.summarize(center='median', fullrange=True)
.reindex(columns=['count', header, header_share, header_growth], level=0)
)
summary
summary.to_excel('output/table_2.6_primary_energy_supply.xlsx')
```
| github_jupyter |
# Caffe2 Basic Concepts - Operators & Nets
In this tutorial we will go through a set of Caffe2 basics: the basic concepts including how operators and nets are being written.
First, let's import caffe2. `core` and `workspace` are usually the two that you need most. If you want to manipulate protocol buffers generated by caffe2, you probably also want to import `caffe2_pb2` from `caffe2.proto`.
```
# We'll also import a few standard python libraries
from matplotlib import pyplot
import numpy as np
import time
# These are the droids you are looking for.
from caffe2.python import core, workspace
from caffe2.proto import caffe2_pb2
```
You might see a warning saying that caffe2 does not have GPU support. That means you are running a CPU-only build. Don't be alarmed - anything CPU is still runnable without problem.
## Workspaces
Let's cover workspaces first, where all the data reside.
If you are familiar with Matlab, workspace consists of blobs you create and store in memory. For now, consider a blob to be a N-dimensional Tensor similar to numpy's ndarray, but is contiguous. Down the road, we will show you that a blob is actually a typed pointer that can store any type of C++ objects, but Tensor is the most common type stored in a blob. Let's show what the interface looks like.
`Blobs()` prints out all existing blobs in the workspace.
`HasBlob()` queries if a blob exists in the workspace. For now, we don't have anything yet.
```
print("Current blobs in the workspace: {}".format(workspace.Blobs()))
print("Workspace has blob 'X'? {}".format(workspace.HasBlob("X")))
```
We can feed blobs into the workspace using `FeedBlob()`.
```
X = np.random.randn(2, 3).astype(np.float32)
print("Generated X from numpy:\n{}".format(X))
workspace.FeedBlob("X", X)
```
Now, let's take a look what blobs there are in the workspace.
```
print("Current blobs in the workspace: {}".format(workspace.Blobs()))
print("Workspace has blob 'X'? {}".format(workspace.HasBlob("X")))
print("Fetched X:\n{}".format(workspace.FetchBlob("X")))
```
Let's verify that the arrays are equal.
```
np.testing.assert_array_equal(X, workspace.FetchBlob("X"))
```
Also, if you are trying to access a blob that does not exist, an error will be thrown:
```
try:
workspace.FetchBlob("invincible_pink_unicorn")
except RuntimeError as err:
print(err)
```
One thing that you might not use immediately: you can have multiple workspaces in Python using different names, and switch between them. Blobs in different workspaces are separate from each other. You can query the current workspace using `CurrentWorkspace`. Let's try switching the workspace by name (gutentag) and creating a new one if it doesn't exist.
```
print("Current workspace: {}".format(workspace.CurrentWorkspace()))
print("Current blobs in the workspace: {}".format(workspace.Blobs()))
# Switch the workspace. The second argument "True" means creating
# the workspace if it is missing.
workspace.SwitchWorkspace("gutentag", True)
# Let's print the current workspace. Note that there is nothing in the
# workspace yet.
print("Current workspace: {}".format(workspace.CurrentWorkspace()))
print("Current blobs in the workspace: {}".format(workspace.Blobs()))
```
Let's switch back to the default workspace.
```
workspace.SwitchWorkspace("default")
print("Current workspace: {}".format(workspace.CurrentWorkspace()))
print("Current blobs in the workspace: {}".format(workspace.Blobs()))
```
Finally, `ResetWorkspace()` clears anything that is in the current workspace.
```
workspace.ResetWorkspace()
```
## Operators
Operators in Caffe2 are kind of like functions. From the C++ side, they all derive from a common interface, and are registered by type, so that we can call different operators during runtime. The interface of operators is defined in `caffe2/proto/caffe2.proto`. Basically, it takes in a bunch of inputs, and produces a bunch of outputs.
Remember, when we say "create an operator" in Caffe2 Python, nothing gets run yet. All it does is to create the protocol buffere that specifies what the operator should be. At a later time it will be sent to the C++ backend for execution. If you are not familiar with protobuf, it is a json-like serialization tool for structured data. Find more about protocol buffers [here](https://developers.google.com/protocol-buffers/).
Let's see an actual example.
```
# Create an operator.
op = core.CreateOperator(
"Relu", # The type of operator that we want to run
["X"], # A list of input blobs by their names
["Y"], # A list of output blobs by their names
)
# and we are done!
```
As we mentioned, the created op is actually a protobuf object. Let's show the content.
```
print("Type of the created op is: {}".format(type(op)))
print("Content:\n")
print(str(op))
```
OK, let's run the operator. We first feed in the input X to the workspace.
Then the simplest way to run an operator is to do `workspace.RunOperatorOnce(operator)`
```
workspace.FeedBlob("X", np.random.randn(2, 3).astype(np.float32))
workspace.RunOperatorOnce(op)
```
After execution, let's see if the operator is doing the right thing, which is our neural network's activation function ([Relu](https://en.wikipedia.org/wiki/Rectifier_(neural_networks))) in this case.
```
print("Current blobs in the workspace: {}\n".format(workspace.Blobs()))
print("X:\n{}\n".format(workspace.FetchBlob("X")))
print("Y:\n{}\n".format(workspace.FetchBlob("Y")))
print("Expected:\n{}\n".format(np.maximum(workspace.FetchBlob("X"), 0)))
```
This is working if you're expected output matches your Y output in this example.
Operators also take optional arguments if needed. They are specified as key-value pairs. Let's take a look at one simple example, which is to create a tensor and fills it with Gaussian random variables.
```
op = core.CreateOperator(
"GaussianFill",
[], # GaussianFill does not need any parameters.
["Z"],
shape=[100, 100], # shape argument as a list of ints.
mean=1.0, # mean as a single float
std=1.0, # std as a single float
)
print("Content of op:\n")
print(str(op))
```
Let's run it and see if things are as intended.
```
workspace.RunOperatorOnce(op)
temp = workspace.FetchBlob("Z")
pyplot.hist(temp.flatten(), bins=50)
pyplot.title("Distribution of Z")
```
If you see a bell shaped curve then it worked!
## Nets
Nets are essentially computation graphs. We keep the name `Net` for backward consistency (and also to pay tribute to neural nets). A Net is composed of multiple operators just like a program written as a sequence of commands. Let's take a look.
When we talk about nets, we will also talk about BlobReference, which is an object that wraps around a string so we can do easy chaining of operators.
Let's create a network that is essentially the equivalent of the following python math:
```
X = np.random.randn(2, 3)
W = np.random.randn(5, 3)
b = np.ones(5)
Y = X * W^T + b
```
We'll show the progres step by step. Caffe2's `core.Net` is a wrapper class around a NetDef protocol buffer.
When creating a network, its underlying protocol buffer is essentially empty other than the network name. Let's create the net and then show the proto content.
```
net = core.Net("my_first_net")
print("Current network proto:\n\n{}".format(net.Proto()))
```
Let's create a blob called X, and use GaussianFill to fill it with some random data.
```
X = net.GaussianFill([], ["X"], mean=0.0, std=1.0, shape=[2, 3], run_once=0)
print("New network proto:\n\n{}".format(net.Proto()))
```
You might have boserved a few differences from the earlier `core.CreateOperator` call. Basically, when we have a net, you can direct create an operator *and* add it to the net at the same time using Python tricks: essentially, if you call `net.SomeOp` where SomeOp is a registered type string of an operator, this essentially gets translated to
```
op = core.CreateOperator("SomeOp", ...)
net.Proto().op.append(op)
```
Also, you might be wondering what X is. X is a `BlobReference` which basically records two things:
- what its name is. You can access the name by str(X)
- which net it gets created from. It is recorded by an internal variable `_from_net`, but most likely
you won't need that.
Let's verify it. Also, remember, we are not actually running anything yet, so X contains nothing but a symbol. Don't expect to get any numerical values out of it right now :)
```
print("Type of X is: {}".format(type(X)))
print("The blob name is: {}".format(str(X)))
```
Let's continue to create W and b.
```
W = net.GaussianFill([], ["W"], mean=0.0, std=1.0, shape=[5, 3], run_once=0)
b = net.ConstantFill([], ["b"], shape=[5,], value=1.0, run_once=0)
```
Now, one simple code sugar: since the BlobReference objects know what net it is generated from, in addition to creating operators from net, you can also create operators from BlobReferences. Let's create the FC operator in this way.
```
Y = X.FC([W, b], ["Y"])
```
Under the hood, `X.FC(...)` simply delegates to `net.FC` by inserting `X` as the first input of the corresponding operator, so what we did above is equivalent to
```
Y = net.FC([X, W, b], ["Y"])
```
Let's take a look at the current network.
```
print("Current network proto:\n\n{}".format(net.Proto()))
```
Too verbose huh? Let's try to visualize it as a graph. Caffe2 ships with a very minimal graph visualization tool for this purpose. Let's show that in ipython.
```
from caffe2.python import net_drawer
from IPython import display
graph = net_drawer.GetPydotGraph(net, rankdir="LR")
display.Image(graph.create_png(), width=800)
```
So we have defined a `Net`, but nothing gets executed yet. Remember that the net above is essentially a protobuf that holds the definition of the network. When we actually want to run the network, what happens under the hood is:
- Instantiate a C++ net object from the protobuf;
- Call the instantiated net's Run() function.
Before we do anything, we should clear any earlier workspace variables with `ResetWorkspace()`.
Then there are two ways to run a net from Python. We will do the first option in the example below.
1. Using `workspace.RunNetOnce()`, which instantiates, runs and immediately destructs the network.
2. A little bit more complex and involves two steps:
(a) call `workspace.CreateNet()` to create the C++ net object owned by the workspace, and
(b) use `workspace.RunNet()` by passing the name of the network to it.
```
workspace.ResetWorkspace()
print("Current blobs in the workspace: {}".format(workspace.Blobs()))
workspace.RunNetOnce(net)
print("Blobs in the workspace after execution: {}".format(workspace.Blobs()))
# Let's dump the contents of the blobs
for name in workspace.Blobs():
print("{}:\n{}".format(name, workspace.FetchBlob(name)))
```
Now let's try the second way to create the net, and run it. First clear the variables with `ResetWorkspace()`, create the net with the workspace's net object you created earlier `CreateNet(net_object)`, and then run the net by name with `RunNet(net_name)`.
```
workspace.ResetWorkspace()
print("Current blobs in the workspace: {}".format(workspace.Blobs()))
workspace.CreateNet(net)
workspace.RunNet(net.Proto().name)
print("Blobs in the workspace after execution: {}".format(workspace.Blobs()))
for name in workspace.Blobs():
print("{}:\n{}".format(name, workspace.FetchBlob(name)))
```
There are a few differences between `RunNetOnce` and `RunNet`, but probably the main difference is the computation time overhead. Since `RunNetOnce` involves serializing the protobuf to pass between Python and C and instantiating the network, it may take longer to run. Let's see in this case what the overhead is.
```
# It seems that %timeit magic does not work well with
# C++ extensions so we'll basically do for loops
start = time.time()
for i in range(1000):
workspace.RunNetOnce(net)
end = time.time()
print('Run time per RunNetOnce: {}'.format((end - start) / 1000))
start = time.time()
for i in range(1000):
workspace.RunNet(net.Proto().name)
end = time.time()
print('Run time per RunNet: {}'.format((end - start) / 1000))
```
OK, so above are a few key components if you would like to use Caffe2 from the python side. We are going to add more to the tutorial as we find more needs. For now, kindly check out the rest of the tutorials!
| github_jupyter |
#Introduction
In this notebook we will use a clustering algorithm to analyze our data (i.e. YouTube comments of a single video).
This will help us extract topics of discussion.
We use the embeddings generated in Assignment 4 as input.
(This notebook will not run without first running the assignment 4 Notebook, as it relies on the data in the folder 'output/')
Each of our comments has been assigned a vector that encodes information about its meaning.
The closer two vectors are, the more similar the meaning.
Each vector is of 512 Dimensions.
Before we can cluster our data we need to reduce the embeddings' dimensionality to overcome the curse of dimensionality.
We use the UMAP ALgorithm for this.
After that we use the KMedoids Algorithm to partition the embedding space and generate our clusters this way.
We need to define the number of clusters we want to have.
To find the optimal number of clusters, we use a simple optimization scheme.
Once the clusters are created, we visualize them.
To do this we reduce the dimensionality of the embeddings again to two dimensions.
Then we render a scatterplot of our data.
Furthermore we want to analyze and interpret our clusters.
To do this, we:
- print some statistics about each of the clusters
- print cluster's medoid (the central sample)
- print the cluster(s) we want to analyze further
Check to see if jupyter lab uses the correct python interpreter with '!which python'.
It should be something like '/opt/anaconda3/envs/[environment name]/bin/python' (on Mac).
If not, try this: https://github.com/jupyter/notebook/issues/3146#issuecomment-352718675
```
!which python
```
# Install dependencies:
```
install_packages = False
if install_packages:
!conda install -c conda-forge umap-learn -y
!conda install -c conda-forge scikit-learn-extra -y
```
# Imports
```
#imports
import pandas as pd
import numpy as np
import os
import time
import matplotlib.pyplot as plt
import umap
from sklearn_extra.cluster import KMedoids
import seaborn as sns
#from sklearn.cluster import AgglomerativeClustering, DBSCAN, KMeans, OPTICS
from sklearn.metrics import silhouette_samples, silhouette_score, pairwise_distances
```
# Functions to Save and load manually
```
# Save and load your data after clustering
def save_results():
data.to_pickle(output_path+'data_clustered'+'.pkl')
def load_results():
data = pd.read_pickle(output_path+'data_clustered'+'.pkl')
```
# Set pandas print options
This will improve readability of printed pandas dataframe.
```
pd.set_option('display.max_rows', None)
pd.set_option('display.max_columns', None)
pd.set_option('display.width', None)
pd.set_option('display.max_colwidth', None)
```
## Set global Parameters
Set your parameters here:
output_path: Files generated in this notebook will be saved here.
model_type: Define which model was used to produce the embeddings. (Check the name of the .npy-file containing the embeddings)
```
output_path = "./output/"
model_type = 'Transformer' #@param ['DAN','Transformer','Transformer_Multilingual']
```
# Load Data
Load the preprocessed data as a pandas dataframe.
And load the embeddings as a numpy ndarray (a matrix in our case).
```
data = pd.read_pickle(output_path+'data_preprocessed'+'.pkl')
labels_default = np.zeros(len(data.index))-1
data['label_manual'] = labels_default
embeddings = np.load(output_path+'/embeddings'+model_type+'.npy', mmap_mode=None, allow_pickle=False, fix_imports=True, encoding='ASCII')
```
# Dimensionality reduction with UMAP
We reduce the number of dimensions of our embeddings to make possibly present clusters more pronounced.
The number of dimensions (num_dimensions) depends on the number of samples
```
# Set the number of dimensions to reduce to
num_dimensions =100
reducer_clustering = umap.UMAP(n_neighbors=50,
n_components=num_dimensions,
metric='cosine',
#n_epochs=200,
learning_rate=.5,
init='spectral',
min_dist=0,
#spread=5.0,
#set_op_mix_ratio=1.0,
#local_connectivity=1.0,
#negative_sample_rate=5,
#transform_queue_size=4.0,
force_approximation_algorithm=True,
unique=True)
embeddings_umap = reducer_clustering.fit_transform(embeddings)
```
# Optimize the Number of Clusters
```
#optimize number of clusters
optimize_number_of_clusters = True#@param {type:'boolean'}
min_clusters= 5
max_clusters=55
step=5
if optimize_number_of_clusters:
rows_list = []
inertias = []
n_clusters = []
silouette_scores = []
init_param = 'k-medoids++' #@param ['random', 'heuristic', 'k-medoids++']
random_state_param=134 #@param {type:'number'}
for i in range(min_clusters,max_clusters, step):
temp_clustering = KMedoids(n_clusters=i, metric='euclidean', init=init_param, max_iter=300, random_state=random_state_param).fit(embeddings_umap)
silhouette_avg = silhouette_score(embeddings_umap, temp_clustering.labels_)
print("n_clusters:",i, "silhouette_avg:",silhouette_avg)
silhouette_dict = {'number of clusters': i, 'silhouette average': silhouette_avg}
rows_list.append(silhouette_dict)
results = pd.DataFrame(rows_list)
sns.lineplot(x = 'number of clusters', y = 'silhouette average',data = results)
```
# Clustering with KMedoids
```
number_of_clusters = 20
init_param = 'k-medoids++' #@param ['random', 'heuristic', 'k-medoids++']
clustering_model = KMedoids(n_clusters=number_of_clusters,
metric='cosine',
init=init_param,
max_iter=150,
random_state=None).fit(embeddings_umap)
clustering_model
labels = clustering_model.labels_
data["label_kmedoids"] = labels
print("cluster","members", data["label_kmedoids"].value_counts().sort_values())
clustering_model.inertia_
medoids_indices = clustering_model.medoid_indices_
#calculate distances
distances = np.diag(pairwise_distances(X = clustering_model.cluster_centers_[labels], Y = embeddings_umap[:], metric='cosine'))
data["distance_kmedoids"] = distances
```
# Dimensionality Reduction for Visualization
```
num_dimensions =2
reducer_visualization = umap.UMAP(n_neighbors=50,
n_components=num_dimensions,
metric='cosine',
output_metric='euclidean',
#n_epochs=200,
learning_rate=.5,
init='spectral',
min_dist=.1,
spread=5.0,
set_op_mix_ratio=1.0,
local_connectivity=1.0,
negative_sample_rate=5,
transform_queue_size=4.0,
force_approximation_algorithm=True,
unique=True)
embeddings_umap_2d = reducer_visualization.fit_transform(embeddings)
```
# Visualize clustering results
```
#@markdown Set the color palette used for visualizing different clusters
palette_param = "Spectral" #@param ['Accent','cubehelix', "tab10", 'Paired', "Spectral"]
#@markdown Set opacity of data points (1 = opaque, 0 = invisible)
alpha_param = 1 #@param {type:"slider", min:0, max:1, step:0.01}
sns.relplot(x = embeddings_umap_2d[:, 0], y = embeddings_umap_2d[:, 1], hue = data['label_kmedoids'], palette = palette_param,alpha = alpha_param,height = 10)
```
## Highlight one cluster
```
## Choose a cluster to higlight:
cluster_num = 15
cluster_num_2 = 14
data['highlight'] = np.zeros(len(data.index))
data.loc[data['label_kmedoids'] == cluster_num, 'highlight'] = 1
data.loc[data['label_kmedoids'] == cluster_num_2, 'highlight'] = 2
sns.relplot(x = embeddings_umap_2d[:, 0], y = embeddings_umap_2d[:, 1], hue = data['highlight'], palette = "Accent",alpha = 0.8,height = 10)
```
# Print Medoids and cluster statistics
```
# print the medoids
data.iloc[medoids_indices]
# print statistics for each cluster
data['label_kmedoids'].value_counts().sort_values()
for k,g in data.groupby(by = 'label_kmedoids'):
print(g.iloc[0]['label_kmedoids'],"number of samples: ",len(g.index),"mean distance from center: ", 100*np.mean(g['distance_kmedoids']), "Proportion of replies:", 100*np.sum(g['isReply'])/len(g.index))
```
# Print Cluster
Print the comments within a cluster. Comments are sorted by their distance from the cluster medoid
```
# Choose a cluster to print
cluster_number = 14
# Choose the number of samples to print
number_of_samples_to_print = 1000
data['label_kmedoids'] = data['label_kmedoids'].astype('category')
cluster = data[data['label_kmedoids']==cluster_number]
if cluster["text"].count()<=number_of_samples_to_print:
number_of_samples_to_print = cluster["text"].count()
cluster = cluster.sort_values(by='distance_kmedoids')
print("Number of samples in the cluster:", cluster["text"].count())
print("Average Distance from cluster center:", np.mean(cluster['distance_kmedoids']))
cluster['text']
```
# Assign Cluster labels manually
cluster_number: which cluster would you like to assign labels to?
min_distance: the minimum distance from the cluster medoid be for a data point to still get the specified label
max_distance: the maximum distance from the cluster medoid be for a data point to still get the specified label
label_manual: your label
```
#which cluster would you like to assign labels to?
cluster_number = 15
#your label
label_manual = 'rationality'
#the minimum distance from the cluster medoid be for a data point to still get the specified label
min_distance = 0
#the maximum distance from the cluster medoid be for a data point to still get the specified label
max_distance = 1000
# 2. Filter data by cluster label and specified label to filtered data
data.loc[(data['label_kmedoids']==cluster_number) & (data['distance_kmedoids'] <= max_distance) & (data['distance_kmedoids'] >= min_distance), 'label_manual'] = label_manual
data[data['label_kmedoids']==cluster_number].sort_values(by='distance_kmedoids')
```
| github_jupyter |
## Classifying Iris Notebook
Please make sure you have **matplotlib** installed in the compute context you choose as kernel.
- For local or remote **Docker kernels**, please ensure **notebook** and **matplotlib** are listed in your **conda_dependencies.yml** file under **aml_config** folder.
```
name: project_environment
dependencies:
- python=3.5.2
- scikit-learn
- pip:
- notebook
- matplotlib
```
```
%matplotlib inline
%azureml history off
import pickle
import sys
import os
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from sklearn import decomposition
from sklearn import datasets
from sklearn import preprocessing
from sklearn.datasets import load_iris
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import precision_recall_curve
from azureml.logging import get_azureml_logger
from azureml.dataprep import package
logger = get_azureml_logger()
print ('Python version: {}'.format(sys.version))
print ()
%azureml history on
#load Iris dataset from a DataPrep package
iris = package.run('iris.dprep', dataflow_idx=0, spark=False)
# load features and labels
X, Y = iris[['Sepal Length', 'Sepal Width', 'Petal Length', 'Petal Width']].values, iris['Species'].values
# tag this cell to measure duration
logger.log("Cell","Load Data")
logger.log("Rows",iris.shape[0])
print ('Iris dataset shape: {}'.format(iris.shape))
logger.log("Cell", "Training")
# change regularization rate and you will likely get a different accuracy.
reg = 0.01
print("Regularization rate is {}".format(reg))
logger.log('Regularization Rate', reg)
# train a logistic regression model
clf = LogisticRegression(C=1/reg).fit(X, Y)
print (clf)
# Log curves for label value 'Iris-versicolor'
y_scores = clf.predict_proba(X)
precision, recall, thresholds = precision_recall_curve(Y, y_scores[:,1],pos_label='Iris-versicolor')
logger.log("Precision",precision)
logger.log("Recall",recall)
logger.log("Thresholds",thresholds)
accuracy = clf.score(X, Y)
logger.log('Accuracy', accuracy)
print ("Accuracy is {}".format(accuracy))
logger.log("Cell", "Scoring")
# predict a new sample
X_new = [[3.0, 3.6, 1.3, 0.25]]
print ('New sample: {}'.format(X_new))
pred = clf.predict(X_new)
logger.log('Prediction', pred.tolist())
print('Predicted class is {}'.format(pred))
%azureml history off
# Plot Iris data in 3D
centers = [[1, 1], [-1, -1], [1, -1]]
fig = plt.figure(1, figsize=(8, 6))
plt.clf()
ax = Axes3D(fig, rect=[0, 0, .95, 1], elev=48, azim=134)
plt.cla()
# decompose 4 feature columns into 3 components for 3D plotting
pca = decomposition.PCA(n_components=3)
pca.fit(X)
X = pca.transform(X)
le = preprocessing.LabelEncoder()
le.fit(Y)
Y = le.transform(Y)
for name, label in [('Setosa', 0), ('Versicolour', 1), ('Virginica', 2)]:
ax.text3D(X[Y == label, 0].mean(),
X[Y == label, 1].mean() + 1.5,
X[Y == label, 2].mean(), name,
horizontalalignment='center',
bbox=dict(alpha=.5, edgecolor='w', facecolor='w'))
# Reorder the labels to have colors matching the cluster results
y = np.choose(Y, [1, 2, 0]).astype(np.float)
ax.scatter(X[:, 0], X[:, 1], X[:, 2], c=Y, cmap=plt.cm.spectral)
ax.w_xaxis.set_ticklabels([])
ax.w_yaxis.set_ticklabels([])
ax.w_zaxis.set_ticklabels([])
plt.show()
```
| github_jupyter |
```
# Copyright 2022 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Get started with Vertex AI TensorBoard
In machine learning, to improve something you often need to be able to measure it. TensorBoard is a tool for providing the measurements and visualizations needed during the machine learning workflow. It enables tracking experiment metrics like loss and accuracy, visualizing the model graph, projecting embeddings to a lower dimensional space, and much more.
Vertex AI TensorBoard is the managed enterprise-ready version of the open source TensorBoard project.
This quickstart will show how to quickly get started with Vertex AI TensorBoard.
```
# Installing the Vertex AI TensorBoard uploader
!python -m pip install google-cloud-aiplatform['tensorboard'] --upgrade
import datetime
import tensorflow as tf
# Clear any logs from previous runs
!rm -rf ./logs/
project_id = "YOUR PROJECT ID"
region = "YOUR REGION"
content_name = "vertex-ai-tb-get-started"
```
Using the [MNIST](https://en.wikipedia.org/wiki/MNIST_database) dataset as the example, normalize the data and write a function that creates a simple Keras model for classifying the images into 10 classes.
```
mnist = tf.keras.datasets.mnist
(x_train, y_train), (x_test, y_test) = mnist.load_data()
x_train, x_test = x_train / 255.0, x_test / 255.0
def create_model():
return tf.keras.models.Sequential(
[
tf.keras.layers.Flatten(input_shape=(28, 28)),
tf.keras.layers.Dense(512, activation="relu"),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(10, activation="softmax"),
]
)
```
## Using TensorBoard with Keras Model.fit()
When training with Keras's [Model.fit()](https://www.tensorflow.org/api_docs/python/tf/keras/models/Model#fit), adding the `tf.keras.callbacks.TensorBoard` callback ensures that logs are created and stored. Additionally, enable histogram computation every epoch with `histogram_freq=1` (this is off by default)
Place the logs in a timestamped subdirectory to allow easy selection of different training runs.
```
model = create_model()
model.compile(
optimizer="adam", loss="sparse_categorical_crossentropy", metrics=["accuracy"]
)
log_dir = "logs/fit/" + datetime.datetime.now().strftime("%Y%m%d-%H%M%S")
tensorboard_callback = tf.keras.callbacks.TensorBoard(log_dir=log_dir, histogram_freq=1)
model.fit(
x=x_train,
y=y_train,
epochs=5,
validation_data=(x_test, y_test),
callbacks=[tensorboard_callback],
)
from google.cloud import aiplatform
aiplatform.init(project=project_id, location=region)
tensorboard = aiplatform.Tensorboard.create(display_name=content_name)
TENSORBOARD_RESOURCE_NAME = tensorboard.resource_name
!tb-gcp-uploader --one_shot=True --tensorboard_resource_name $TENSORBOARD_RESOURCE_NAME -logdir="logs/fit" --experiment_name="test-experiment1"
```
<!-- <img class="tfo-display-only-on-site" src="https://github.com/tensorflow/tensorboard/blob/master/docs/images/quickstart_model_fit.png?raw=1"/> -->
A brief overview of the dashboards shown (tabs in top navigation bar):
* The **Scalars** dashboard shows how the loss and metrics change with every epoch. You can use it to also track training speed, learning rate, and other scalar values.
* The **Graphs** dashboard helps you visualize your model. In this case, the Keras graph of layers is shown which can help you ensure it is built correctly.
* The **Distributions** and **Histograms** dashboards show the distribution of a Tensor over time. This can be useful to visualize weights and biases and verify that they are changing in an expected way.
Additional TensorBoard plugins are automatically enabled when you log other types of data. For example, the Keras TensorBoard callback lets you log images and embeddings as well. You can see what other plugins are available in TensorBoard by clicking on the "inactive" dropdown towards the top right.
## Using Vertex AI TensorBoard with other methods
When training with methods such as [`tf.GradientTape()`](https://www.tensorflow.org/api_docs/python/tf/GradientTape), use `tf.summary` to log the required information.
Use the same dataset as above, but convert it to `tf.data.Dataset` to take advantage of batching capabilities:
```
train_dataset = tf.data.Dataset.from_tensor_slices((x_train, y_train))
test_dataset = tf.data.Dataset.from_tensor_slices((x_test, y_test))
train_dataset = train_dataset.shuffle(60000).batch(64)
test_dataset = test_dataset.batch(64)
```
The training code follows the [advanced quickstart](https://www.tensorflow.org/tutorials/quickstart/advanced) tutorial, but shows how to log metrics to TensorBoard. Choose loss and optimizer:
```
loss_object = tf.keras.losses.SparseCategoricalCrossentropy()
optimizer = tf.keras.optimizers.Adam()
```
Create stateful metrics that can be used to accumulate values during training and logged at any point:
```
# Define our metrics
train_loss = tf.keras.metrics.Mean("train_loss", dtype=tf.float32)
train_accuracy = tf.keras.metrics.SparseCategoricalAccuracy("train_accuracy")
test_loss = tf.keras.metrics.Mean("test_loss", dtype=tf.float32)
test_accuracy = tf.keras.metrics.SparseCategoricalAccuracy("test_accuracy")
```
Define the training and test functions:
```
def train_step(model, optimizer, x_train, y_train):
with tf.GradientTape() as tape:
predictions = model(x_train, training=True)
loss = loss_object(y_train, predictions)
grads = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(grads, model.trainable_variables))
train_loss(loss)
train_accuracy(y_train, predictions)
def test_step(model, x_test, y_test):
predictions = model(x_test)
loss = loss_object(y_test, predictions)
test_loss(loss)
test_accuracy(y_test, predictions)
```
Set up summary writers to write the summaries to disk in a different logs directory:
```
current_time = datetime.datetime.now().strftime("%Y%m%d-%H%M%S")
train_log_dir = "logs/gradient_tape/" + current_time + "/train"
test_log_dir = "logs/gradient_tape/" + current_time + "/test"
train_summary_writer = tf.summary.create_file_writer(train_log_dir)
test_summary_writer = tf.summary.create_file_writer(test_log_dir)
```
Start training. Use `tf.summary.scalar()` to log metrics (loss and accuracy) during training/testing within the scope of the summary writers to write the summaries to disk. You have control over which metrics to log and how often to do it. Other `tf.summary` functions enable logging other types of data.
```
model = create_model() # reset our model
EPOCHS = 5
for epoch in range(EPOCHS):
for (x_train, y_train) in train_dataset:
train_step(model, optimizer, x_train, y_train)
with train_summary_writer.as_default():
tf.summary.scalar("loss", train_loss.result(), step=epoch)
tf.summary.scalar("accuracy", train_accuracy.result(), step=epoch)
for (x_test, y_test) in test_dataset:
test_step(model, x_test, y_test)
with test_summary_writer.as_default():
tf.summary.scalar("loss", test_loss.result(), step=epoch)
tf.summary.scalar("accuracy", test_accuracy.result(), step=epoch)
template = "Epoch {}, Loss: {}, Accuracy: {}, Test Loss: {}, Test Accuracy: {}"
print(
template.format(
epoch + 1,
train_loss.result(),
train_accuracy.result() * 100,
test_loss.result(),
test_accuracy.result() * 100,
)
)
# Reset metrics every epoch
train_loss.reset_states()
test_loss.reset_states()
train_accuracy.reset_states()
test_accuracy.reset_states()
```
Upload the TensorBoard logs again, this time pointing it at the new log directory and uploading to a different experiment (test-experiment2).
We could have also monitored training while it progresses by continuously uploading the TensorBoard logs, by omitting the "one_shot" flag.
```
!tb-gcp-uploader --one_shot=True --tensorboard_resource_name $TENSORBOARD_RESOURCE_NAME --logdir="logs/gradient_tape" --experiment_name="test-experiment2"
```
<!-- <img class="tfo-display-only-on-site" src="https://github.com/tensorflow/tensorboard/blob/master/docs/images/quickstart_gradient_tape.png?raw=1"/> -->
That's it! You have now seen how to use Vertex AI TensorBoard both through the Keras callback and through `tf.summary` for more custom scenarios.
| github_jupyter |
[](https://colab.research.google.com/github/eirasf/GCED-AA2/blob/main/lab7/lab7-parte2.ipynb)
# Práctica 7: Residual neural networks
### Pre-requisitos. Instalar paquetes
Para la segunda parte de este Laboratorio 7 necesitaremos TensorFlow y TensorFlow-Datasets. Además, como habitualmente, fijaremos la semilla aleatoria para asegurar la reproducibilidad de los experimentos.
```
import tensorflow as tf
import tensorflow_datasets as tfds
#Fijamos la semilla para poder reproducir los resultados
import os
import numpy as np
import random
seed=1234
os.environ['PYTHONHASHSEED']=str(seed)
tf.random.set_seed(seed)
np.random.seed(seed)
random.seed(seed)
```
Además, cargamos también APIs que vamos a emplear para que el código quede más legible
```
#API de Keras, modelo Sequential y la capa Dense
from tensorflow import keras
from keras.models import Sequential
from keras.layers import Dense
#Para mostrar gráficas
from matplotlib import pyplot
```
### Carga del conjunto de datos
De nuevo, seguimos empleando el conjunto *german_credit_numeric* ya empleado en los laboratorios anteriores.
```
# TODO: Carga el conjunto german_credit como ds_train
# Indica además un tamaño de batch de 128 y que se repita indefinidamente
ds_train = ...
```
## Visualización del desvanecimiento del gradiente
En este apartado visualizaremos los tamaños de los gradientes, como hicimos en la primera parte. Para ello mantenemos la declaración de `GradientLoggingSequentialModel`.
```
class GradientLoggingSequentialModel(tf.keras.models.Sequential):
def __init__(self, **kwargs):
super().__init__(**kwargs)
# En la inicialización instanciamos una nueva variable en la que
# registraremos el historial de tamaños de gradientes de cada capa
self.gradient_history = {}
def compile(self, **kwargs):
result = super().compile(**kwargs)
# Una vez sabemos la arquitectura, podemos inicializar la historia
# de gradientes de cada capa a una lista vacía.
for l in self.layers:
self.gradient_history[l.name] = []
return result
def _save_gradients(self, gradients):
# A cada paso de entrenamiento llamaremos a esta función para que
# registre los gradientes.
# En la lista gradients se encuentran los gradientes de las distintas
# capas por orden. Cada capa l tendrá un número de gradientes que
# concidirá con l.trainable_variables.
# Teniendo esto en cuenta, recorremos los gradientes, calculamos su
# tamaño y guardamos la media de tamaños de cada capa en el histórico
i = 0
for layer in self.layers:
gradient_sizes = []
for lw in layer.trainable_variables:
g_size = np.linalg.norm(gradients[i].numpy())
gradient_sizes.append(g_size)
i += 1
mean_gradient_size = np.mean(gradient_sizes)
self.gradient_history[layer.name].append(mean_gradient_size)
def train_step(self, data):
# Haremos un paso de entrenamiento personalizado basado en
# https://www.tensorflow.org/guide/keras/customizing_what_happens_in_fit#a_first_simple_example
# Dejaremos el ejemplo tal cual, añadiendo tan solo la llamada a
# _save_gradients una vez que disponemos de los gradientes
# Unpack the data. Its structure depends on your model and
# on what you pass to `fit()`.
x, y = data
with tf.GradientTape() as tape:
y_pred = self(x, training=True) # Forward pass
# Compute the loss value
# (the loss function is configured in `compile()`)
loss = self.compiled_loss(y, y_pred, regularization_losses=self.losses)
# Compute gradients
trainable_vars = self.trainable_variables
gradients = tape.gradient(loss, trainable_vars)
# Update weights
self.optimizer.apply_gradients(zip(gradients, trainable_vars))
# Update metrics (includes the metric that tracks the loss)
self.compiled_metrics.update_state(y, y_pred)
# Llamada añadida para grabar los gradientes.
self._save_gradients(gradients)
# Return a dict mapping metric names to current value
return {m.name: m.result() for m in self.metrics}
```
### Creación del bloque con conexión residual
En una red neuronal *feed-forward* convencional, la salida de cada capa se utiliza como entrada de las capa siguiente.
<img src="./img/resnet.png" alt="Dos capas en una red FF" width="400"/>
En contraste, en una ResNet se introducen bloques que incluyen conexiones residuales con el objetivo de favorecer la propagación de los gradientes.
<img src="./img/resnet-2.png" alt="Dos capas en una red FF con conexión residual" width="400"/>
A pesar de que las ResNet se suelen utilizar con redes convolucionales, en este Laboratorio utilizaremos una red *feed-forward*.
Para poder utilizar este tipo de bloques en nuestra arquitectura definiremos un nuevo tipo de modelo denominado `DoubleDenseWithSkipModel` que consistirá en lo representado en la imagen:
- Una primera capa Dense, cuya salida sirve de entrada a...
- Una segunda capa Dense lineal (sin activación), cuya salida sirve de entrada a...
- A una operación suma que la añade a la entrada original a la primera capa. La salida de esta suma servirá de entrada a...
- Una función de activación, cuya salida será la salida del bloque.
Utilizaremos la función sigmoide como función de activación en ambos casos. La entrada y la salida del bloque deberán tener la misma dimensión para que se pueda realizar la suma. Por simplicidad, mantendremos la salida de la primera capa con la misma dimensión.
Nuestra nueva clase heredará de `Model`, por lo que deberá implementar los métodos `build` y `call`. En celdas posteriores añadiremos estos bloques a un modelo `Sequential` como si fuesen capas.
```
class DoubleDenseWithSkipModel(tf.keras.models.Model):
def __init__(self, **kwargs):
super(DoubleDenseWithSkipModel, self).__init__(**kwargs)
# TODO: Completa el método build
def build(self, input_shape):
...
# TODO: Completa el método skip
def call(self, x):
...
```
## Creamos un modelo *GradientLoggingSequentialModel* utilizando las nuevas capas
Creamos un modelo *GradientLoggingSequentialModel* para ajustar a los datos de entrada siguiendo las especificaciones dadas. Deberá incluir (además de las capas de entrada y salida) una capa de 10 unidades con activación sigmoide y 10 de los nuevos bloques.
```
# TODO - Define en model una red GradientLoggingSequentialModel
# Pon una capa densa con 10 unidades con activación sigmoide y 10 capas DoubleDenseWithSkipModel
model = ...
#Construimos el modelo y mostramos
model.build()
print(model.summary())
```
### Entrenamiento del modelo
Como en la primera parte, vamos a establecer la función de pérdida (entropía cruzada binaria), el optimizador (SGD con LR $10^{-3}$) y la métrica que nos servirá para evaluar el rendimiento del modelo entrenado (área bajo la curva).
```
#TODO - Compila el modelo. Utiliza la opción run_eagerly=True para que se puedan registrar los gradientes a cada paso
model.compile(...)
```
Entrenamos el modelo usando model.fit
```
#TODO - entrenar el modelo utilizando 8 steps por epoch. Con 10 epochs nos valdrá para comprobar el desvanecimiento de gradientes.
model.fit(...)
```
Una vez hayas encontrado un valor de learning rate que consiga una convergencia rápida, guarda el history de la pérdida en la variable history_sgd para poder hacer comparativas.
```
pyplot.figure(figsize=(14, 6), dpi=80)
pyplot.boxplot(model.gradient_history.values())
pyplot.yscale('log')
pyplot.xticks(ticks=range(1,len(model.gradient_history)+1), labels=model.gradient_history.keys())
pyplot.show()
```
## Comparativa
- Contrasta los resultados con los obtenidos en la primera parte
- Cambia las activaciones del bloque y de la primera capa oculta de la red a ReLU y observa la diferencia.
- Alarga el entrenamiento y prueba distintos optimizadores para intentar que el modelo entrene correct
| github_jupyter |
A workbook illustrating how to use the gp_sandbox functions such as simulating data (random, clustered, candence), sampling the data, and testing multiple input values for parameters of kernels.
```
%matplotlib notebook
import gp_sandbox as cgp
import pandas as pd
import george
import numpy as np
import matplotlib.pyplot as plt
from ipywidgets import interact
```
# Simulated Data
```
def simulated_data_comparison(gamma, period, amp):
x_pos, py, gp1 = cgp.create_data(gamma, period, amp)
def _do_work(n = 10, m = 0, o = 0):
x, y, yerr = cgp.sample_data(x_pos, py, 1, n, m, o)
gp2, ln_likelihood = cgp.fit_data(x,y,yerr,gamma,period,gp1)
gp3, ln_likelihood_opt, result = cgp.optimize(y, gp2, ln_likelihood, print_results = True)
#ln_prior = cgp.log_prior(gamma, period, amp)
#ln_post = cgp.log_post(ln_prior, ln_likelihood_opt)
#print("Initial likelihood: " + str(ln_likelihood))
#print("Optimized likelihood: " + str(ln_likelihood_opt))
#print("Log prior: " + str(ln_prior))
#print("Log post: " + str(ln_post))
cgp.plotting(x,y,yerr,py,x_pos,gp2)
#cgp.plotting(x,y,yerr,py,x_pos,gp3)
return _do_work
#n will only work if m is set equal to 0
vary_nm = simulated_data_comparison(10,2,2)
interact(vary_nm, n=(0, 100,5), m = (0, 10), o = (0,5), continuous_update=False)
```
# Imported Data
## Asteriod 1291
```
#import file into a pandas dataframe
df = pd.read_csv('../data/1291_lc_49627_to_49787.txt', delimiter=' ',header=None, names=['x','y'], dtype={'x':float, 'y':float})
pre_x = (df.x)
pre_y = df.y
cap = 1000
#sample only from the first 2000 points for x and y (out of 400,000ish...)
x, y, yerr = cgp.sample_data(pre_x[0:cap], pre_y[0:cap], 1 , 0, 0, 1)
#plot the two original points
plt.figure(figsize=(5, 3.75))
plt.plot(pre_x[0:cap], pre_y[0:cap], '.', alpha=0.5,label="Original")
plt.plot(x,y, '.', alpha=0.5, label="Sample")
#guess the kernel numbers
kernel = 5*george.kernels.ExpSine2Kernel(gamma=1, log_period=-1.5) #optimal gamma was ~15
gp = george.GP(kernel)
gp.compute(x,yerr)
pred, pred_var = gp.predict(y, x, return_var=True)
#plot the kernel-fitted guess
plt.fill_between(x, pred - np.sqrt(pred_var), pred + np.sqrt(pred_var), color="green", alpha=0.4)
plt.plot(x, pred, lw=1.5, alpha=0.7, label="Before")
#all the x-values visually present we want to map onto
x_short = np.linspace(pre_x[0], pre_x[cap], 1000)
#optimize the fit
gp2, ln_like2, result = cgp.optimize(y,gp,gp.lnlikelihood, print_results=True)
pred, pred_var = gp2.predict(y, x_short, return_var=True)
#print the optimized fit
plt.fill_between(x_short, pred - np.sqrt(pred_var), pred + np.sqrt(pred_var), color="red", alpha=0.4)
plt.plot(x_short, pred, "red", lw=1.5, alpha=0.5, label="After")
plt.xlim([pre_x[0], pre_x[cap]])
plt.legend()
```
## Asteriod 3200
```
#import file into a pandas dataframe
df = pd.read_csv('../data/3200_lc_49627_to_49787.txt', delimiter=' ',header=None, names=['x','y'], dtype={'x':float, 'y':float})
pre_x = df.x
pre_y = df.y
cap = 5000
#sample only from the first 2000 points for x and y (out of 400,000ish...)
x, y, yerr = cgp.sample_data(pre_x[0:cap], pre_y[0:cap], 0.2 , 0, 0, 2)
#plot the two original points
plt.figure(figsize=(5, 3.75))
plt.plot(pre_x[0:cap], pre_y[0:cap], '-', alpha=0.5, label="Original")
plt.plot(x,y, 'ko', ms = 3, alpha=0.5, label="Sample")
#guess the kernel numbers
kernel = 0.5*george.kernels.ExpSine2Kernel(gamma=1, log_period=-1.8) #optimal gamma was ~15
gp = george.GP(kernel)
gp.compute(x,yerr)
pred, pred_var = gp.predict(y, x, return_var=True)
#plot the kernel-fitted guess
#plt.fill_between(x, pred - np.sqrt(pred_var), pred + np.sqrt(pred_var), color="green", alpha=0.4)
#plt.plot(x, pred, lw=1.5, alpha=0.7, label="Before")
#all the x-values visually present we want to map onto
x_short = np.linspace(pre_x[0], pre_x[cap], 1000)
#optimize the fit
gp2, ln_like2, result = cgp.optimize(y,gp,gp.lnlikelihood, print_results=True)
pred, pred_var = gp2.predict(y, x_short, return_var=True)
#print the optimized fit
plt.fill_between(x_short, pred - np.sqrt(pred_var), pred + np.sqrt(pred_var), color="red", alpha=0.4)
plt.plot(x_short, pred, "red", lw=1.5, alpha=0.5, label="After")
plt.xlim([pre_x[0], pre_x[cap]])
plt.legend()
gp.get_parameter_names()
```
# Parameter Study
```
parameter = np.zeros(len(fun_values))
function_value = np.zeros(len(fun_values))
def fun_generator(amp, gamma, log_period):
#guess the kernel numbers
kernel = amp*george.kernels.ExpSine2Kernel(gamma=gamma, log_period=log_period) #optimal gamma was ~15
gp = george.GP(kernel)
gp.compute(x,yerr)
return gp.lnlikelihood(y)
#pred, pred_var = gp.predict(y, x, return_var=True)
#optimize the fit
#gp2, ln_like2, fun = cgp.optimize(y,gp,gp.lnlikelihood, print_results=False)
#return fun
#import file into a pandas dataframe
df = pd.read_csv('3200_lc_49627_to_49787.txt', delimiter=' ',header=None, names=['x','y'], dtype={'x':float, 'y':float})
pre_x = df.x
pre_y = df.y
#sample only from the first 2000 points for x and y (out of 400,000ish...)
x, y, yerr = cgp.sample_data(pre_x[0:2000], pre_y[0:2000], 0.2, int(0.1*len(pre_x[0:2000])), 0, 0)
```
### Period
```
periods = np.zeros((12,2))
for i in np.arange(len(periods)):
periods[i][0] = 2*(i+1)
periods[i][1] = np.log(2.0*(i+1)/24)
fun_values = np.zeros((len(periods),2))
for i in np.arange(len(periods)):
fun_values[i][0] = periods[i][0]
#fun_values[i][1] = periods[i][1]
#print(periods[i][1])
fun_values[i][1] = fun_generator(1,1,periods[i][1])
print(fun_values)
```
### Gamma
```
gamma = np.zeros(12)
for i in np.arange(len(gamma)):
gamma[i] = i+1
fun_values = np.zeros((len(gamma),2))
for i in np.arange(len(gamma)):
fun_values[i][0] = gamma[i]
fun_values[i][1] = fun_generator(1,gamma[i], -1.09861229e+00)
print(fun_values)
```
### Amplitude
```
amp = np.zeros(12)
for i in np.arange(len(amp)):
amp[i] = i+1
fun_values = np.zeros((len(amp),2))
for i in np.arange(len(amp)):
fun_values[i][0] = amp[i]
fun_values[i][1] = fun_generator(amp[i], 1, -1.09861229e+00)
print(fun_values)
```
### Plotting
```
for i in np.arange(len(fun_values)):
parameter[i] = fun_values[i][0]
function_value[i] = fun_values[i][1]
plt.figure()
plt.plot(parameter, function_value, 'o')
plt.ylim([-500,500])
```
## To Do
add more data and sample more sparsely
test sampling with m parameter
make yerr an input (1%)
look at emcee documentation
make parameter graphs
definition - gamma: length scale of variation
### parameter graphs
* amp, gamma, period = 5-10 each
* P = (1-24 hrs)
* 3 separate plots of parameter vs fun output (get from looping)
### cadence
1 pt every 10 minutes of 8 hrs per night
2/4 nights per month (consecutive for 6/2 months)
```
cgp.cadence_set(pre_y, 5)
```
| github_jupyter |
```
# CART
import numpy as np
import random
class CART(object):
def __init__(self, x, y, prune=False):
self.x = x
self.y = y
# print('build cart...')
self.branches = self.buildTree(x,y,prune)
def sign(self,v):
if v < 0:
return -1
else:
return 1
def buildTree(self,x,y,prune=False):
branches = []
if prune: #只有一层分支
branch, left, right = self.branchStump(x,y)
branches.append(branch)
if sum(left[1]) >= 0: #表示对left数据集的y值求和
branches.append([1])
else:
branches.append([-1])
if sum(right[1]) >= 0:
branches.append([1])
else:
branches.append([-1])
return branches
else: #fully grown
if abs(sum(y)) == len(y): #二叉决策树终止条件:当前各个分支下包含的所有样本yn都是同类的
branches.append(y[0])
return branches
else:
branch, left, right = self.branchStump(x,y) #二叉决策树前序排列
branches.append(branch)
branches.append(self.buildTree(left[0],left[1]))
branches.append(self.buildTree(right[0],right[1]))
return branches
def branchStump(self, x, y): #运用desicion stump方法对x和y找到分类出子树的最佳方法!
# print('13. branch tree once...')
dimensions = len(x[0]) #这里dimensions为2,相当于特征数
best_s = True
best_theta = 0
best_dim = 0
best_gini = 100
for dim in range(dimensions):
thetas = np.sort(x[:,dim]) #1/ 每次只选出某一个特征dim用来做decision stump,并且依据该特征对数据集sort。
#2/ 对每个特征,不知道把数据集分到哪一个子分支依据的界限theta是多少,又要每次选出一个theta实验
ss = [True,False] #3/ 不知道x的该特征是正增益还是负增益。所以s为正遍历一遍,为负遍历一遍
for i,theta in enumerate(thetas):
if i > 0:
theta = (theta+thetas[i-1])/2 #取每个分割段的中间作为theta
else:
theta = theta/2 #考虑i=0的特殊情况
for s in ss:
gini = self.computeDivideGini(x,y,s,theta,dim)
if gini < best_gini:
best_gini = gini
best_s = s
best_theta = theta
best_dim = dim
branch = [best_s,best_theta,best_dim] #结果:得到了最适合的s,theta和dim!!!,最适合的依据是基尼系数最小
left,right = self.divideData(x,y,best_s,best_theta,best_dim) #得到了最适合的左右子树
return branch,left,right
def hFunc(self, x, s, theta): #比较x和theta的大小 #s表示方向,可以为true,可以为false
if s:
return self.sign(x-theta) #表示正方向
else:
return -self.sign(x-theta)
def divideData(self,x,y,s,theta,dim): #依据theta这个分割线来具体划分数据集成为两个子数据集
left_x = []
left_y = []
right_x = []
right_y = []
for i in range(len(y)):
if self.hFunc(x[i][dim],s,theta) == -1: #表示在s方向上x[i][dim]大于theta
left_x.append(x[i])
left_y.append(y[i])
else:
right_x.append(x[i])
right_y.append(y[i])
# print('left data:',len(left_y), 'right data:',len(right_y))
# 注意:下面这种列表list的拼接方法,如a=[np.array(left_x),np.array(left_y)]表示得到了一个长度为2的数组a,
# a[0]表示left_x,a[1]表示left_y,则a[0][0][0]表示lef_x的第1个元素的第1个特征值
return [np.array(left_x),np.array(left_y)],[np.array(right_x),np.array(right_y)]
def computeDivideGini(self,x,y,s,theta,dim): #依据theta这个分割线得到对应的两个子数据集
left,right = self.divideData(x,y,s,theta,dim)
left_gini = self.computeGini(left[0],left[1])
right_gini = self.computeGini(right[0],right[1])
return len(left[1])*left_gini+len(right[1])*right_gini
def computeGini(self,x,y): #计算基尼指数
sums = sum(y)
lens = len(y)
if lens == 0:
return 0
pos_num = (sums+lens)/2 #标签为正的y个数
neg_num = lens-pos_num #标签为负的y个数
return 1-(pos_num/lens)*(pos_num/lens)-(neg_num/lens)*(neg_num/lens)
def fit(self,x,branch): #这里x表示单个数据
if len(branch) == 3: #父亲+左子树+右子树的结构
dim = branch[0][2] #dim为父亲的最佳dim
y = self.hFunc(x[dim],branch[0][0],branch[0][1])
if y == -1:
return self.fit(x,branch[1])
else:
return self.fit(x,branch[2])
else: #当最后长度不为3了,表示已经到叶子了,则取出该叶子的值
return branch[0]
def predict(self,x):
res = []
for i in range(len(x)):
res.append(self.fit(x[i],self.branches))
return np.array(res)
# RandomForest
class RandomForest(object):
"""docstring for RandomForest""" # T表示有多少棵树
def __init__(self, x, y, T, prune=False):
self.x = x
self.y = y
self.T = T
self.trees = self.buildRF(x,y,T,prune) #所有的树构成的列表
def boostrap(self,x,y,N): #boostrap:对N个数重新排序,创建新数据集
indexs = [random.randint(0,N) for _ in range(N)]
return x[indexs], y[indexs]
def buildRF(self,x,y,T,prune):
trees = []
for i in range(T):
tx, ty = self.boostrap(x,y,len(y)-1)
trees.append(CART(tx,ty,prune))
return trees
def fit(self,x): #这里的x是一个数据
res = []
for i in range(self.T):
res.append(self.trees[i].fit(x,self.trees[i].branches))
if sum(res) >= 0: #所有数的fit的平均
return 1
else:
return -1
def predict(self,x):
res = []
for i in range(len(x)):
res.append(self.fit(x[i]))
return np.array(res)
if __name__ == '__main__':
print('load data...')
train_data=np.loadtxt('train_data.txt')
test_data=np.loadtxt('test_data.txt')
train_x=train_data[:,0:2]
train_y=train_data[:,2]
test_x=test_data[:,0:2]
test_y=test_data[:,2]
cart = CART(train_x,train_y)
train_predict = cart.predict(train_x)
E_in = 1-sum(train_predict==train_y)/len(train_y)
print('14. E_in:',E_in)
predicted = cart.predict(test_x)
print('15. E_out:',1-sum(predicted==test_y)/len(test_y))
times = 10
E_in = 0
E_out = 0
E_g_in = 0
T = 300
for i in range(times): #随机10次,做10个随机森林,其结果会随着boostrap随机生成的训练集不同而不同
rf = RandomForest(train_x,train_y,T)
for tree in rf.trees:
train_predict = tree.predict(train_x)
g_in = 1-sum(train_predict==train_y)/len(train_y)
E_g_in += g_in #这是对300棵树*10==3000棵树分别做预测的训练误差的和
train_predict = rf.predict(train_x)
e_in = 1-sum(train_predict==train_y)/len(train_y)
E_in += e_in #这是随机森林(1个随机森林(300棵树)*10)的训练误差
predicted = rf.predict(test_x)
e_out = 1-sum(predicted==test_y)/len(test_y)
E_out += e_out #这是随机森林(1个随机森林(300棵树)*10)的验证误差
print('random forest:',i,'e_in:',e_in,'e_out:',e_out)
print('16. E_g_in:', E_g_in/T/times)
print('17. E_in:', E_in/times)
print('18. E_out:', E_out/times)
E_in = 0
E_out = 0
for i in range(times):
rf = RandomForest(train_x,train_y,T,True)
train_predict = rf.predict(train_x)
e_in = 1-sum(train_predict==train_y)/len(train_y)
E_in += e_in
predicted = rf.predict(test_x)
e_out = 1-sum(predicted==test_y)/len(test_y)
E_out += e_out
print('pruned random forest:',i,'e_in:',e_in,'e_out:',e_out)
print('19. E_in:', E_in/times)
print('20. E_out:', E_out/times)
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import glob
import os
import matplotlib.pyplot as plt
from scipy import signal
import sys
def GetRMSE(x2, y2, x1, y1):
from scipy.spatial.distance import pdist
er = []
for idx in range(len(x2)):
X = np.asarray([[x1[idx], y1[idx]],
[x2[idx], y2[idx]]])
temp_er = pdist(X,metric = 'euclidean')
er.append(temp_er[0])
er = np.asarray(er)
return(er)
# Do this for all the raw data..
# [### check list for video annotation - mannually or automated
#### If frames are manually annotated, then read them directly and add to next folder.
#### If automated via Deeplabcut, filter them based on filtering parameters.]
# parameters used to filter data
cutoff = 24
interpol_order = 3 # order for polynomial interpolation
# win_gauss = signal.gaussian(10,3) # gaussian kernal for smoothening interpolated data
# parameters for the savitzky-golay filter
savgol_win = 11
savgol_polyorder = 3
# window_length=7, polyorder=2,
# ROLLING_WINDOW = [11, 21, 31][0]
```
### remove x y points based on threshold ( of rmse distance from previous frame )
```
circ_parameters_path = glob.glob('./dataFolders/Output/Centroids_v3/circle_parameters.csv')
circ_parameters = pd.read_csv(circ_parameters_path[0])
direc = r"./dataFolders/Output/Proboscis/RawTracks/"
visitnum = ['FirstVisit/', 'LastVisit/']
for visit in visitnum:
outpath = os.path.join('./dataFolders/Output/Proboscis/FilteredTracks/', visit)
print(outpath)
if not os.path.exists(outpath):
try:
os.mkdir(outpath)
except OSError:
print ("Creation of the directory %s failed" % outpath)
# def removelist(visit):
# if visit == 'FirstVisit/':
# newlist = ['c-1_m2', 'c-2_m2']
# elif visit == 'Later7thVisit/':
# newlist = ['c-10_m2_visit_6', 'c-1_m1_visit_6']
# elif visit == 'Later20thVisit/':
# newlist = ['c-1_m1_visit_19', 'c-2_m2_visit_19']
# return(newlist)
for visit in visitnum:
path = os.path.join(direc, visit)
trackslist = glob.glob(path + '*.csv')
# # remove problem cases
# n_remove = removelist(visit)
# newlist = []
# for n in trackslist:
# m_temp = os.path.basename(n)[:-4]
# a = m_temp.split('_')[0]
# b = m_temp.split('_')[1]
# m = a+'_'+b
# m in n_remove
# if m not in n_remove:
# newlist.append(n)
newlist = trackslist
outpath = os.path.join('./dataFolders/Output/Proboscis/FilteredTracks/', visit)
outpathfig = os.path.join('./dataFolders/Output/Proboscis/FilteredTracks/Figures', visit)
for data in newlist:
# data = trackslist[0]
name = os.path.basename(data)[:-4]
print('working on ' + name)
file = pd.read_csv(data)
x = file.x.values
y = file.y.values
p = file.likelihood
x_notinView = x <=5
y_notinView = y <=5
x[x_notinView & y_notinView]=np.nan
y[x_notinView & y_notinView]=np.nan
# add filter for DLC likelihood
med = file['likelihood'].rolling(11).median()
x[med < 0.6] = np.nan
y[med < 0.6] = np.nan
if x.size == 0 or y.size == 0:
print(name + 'has emtpy x y tracks')
continue
mothname = [n for n in circ_parameters.name if n + '_' in data][0]
circ_x = circ_parameters.loc[circ_parameters.name == mothname, 'circ_x'].values
circ_y = circ_parameters.loc[circ_parameters.name == mothname, 'circ_y'].values
circ_radii = circ_parameters.loc[circ_parameters.name == mothname, 'circ_radii'].values
# get rmse values for subsequent frames
rmse = GetRMSE(x[1:], y[1:], x[:-1], y[:-1])
filtered_x = np.copy(x[1:])
filtered_y = np.copy(y[1:])
filtered_x[(rmse > cutoff) | (rmse == np.nan)] = np.nan
filtered_y[(rmse > cutoff) | (rmse == np.nan)] = np.nan
filtered_r = np.linalg.norm([filtered_x - circ_x, filtered_y - circ_y], axis = 0)
filtered_r = filtered_r/circ_radii
filt_trajectory = pd.DataFrame([filtered_x, filtered_y, filtered_r]).T
filt_trajectory.columns = ['x', 'y', 'r']
t = (pd.Series(filtered_x).rolling(30).median(center=True))
t_reverse = t[::-1]
t_reverse
s = np.argmax([ ~np.isnan(t) for t in t_reverse ] )
trim = len(t)-s
# Apply filters
trajectory = filt_trajectory.copy()
trajectory = trajectory.loc[0:trim, :]
print(trajectory.shape)
for colname in trajectory.columns:
# print(colname)
trajectory.loc[:, colname] = signal.medfilt(trajectory.loc[:, colname], kernel_size=11)
trajectory.loc[:, colname] = trajectory.loc[:, colname].interpolate(method = 'polynomial'
,order = 3, limit = 40)
nans = trajectory.loc[:,colname].isnull()
trajectory.loc[:,colname] = trajectory.loc[:,colname].interpolate(method = 'pad')
trajectory.loc[:, colname] = signal.savgol_filter(trajectory.loc[:, colname],
window_length=savgol_win,
polyorder=savgol_polyorder,
axis=0)
trajectory.loc[nans, colname]= np.nan
trajectory_r = np.linalg.norm([trajectory.loc[:,'x'].values - circ_x, trajectory.loc[:,'y'].values - circ_y], axis = 0)
trajectory['r'] = trajectory_r/circ_radii
trajectory['x_centered'] = trajectory.x - circ_x
trajectory['y_centered'] = trajectory.y - circ_y
# fig = plt.figure()
axes = pd.concat([filt_trajectory, trajectory], axis = 1).plot(subplots = True, figsize = (15,8))
fig = plt.gcf()
# .get_figure()
fig.savefig(outpathfig + mothname + '_' + visit[:-1] + '.pdf')
plt.close()
trajectory.to_csv(outpath + mothname + '_' + visit[:-1] + '.csv')
```
| github_jupyter |
Imports
-------
```
%matplotlib inline
import numpy as np
from sklearn.pipeline import Pipeline
from msmbuilder.example_datasets import FsPeptide
from msmbuilder.featurizer import DihedralFeaturizer
from msmbuilder.preprocessing import RobustScaler
from msmbuilder.decomposition import tICA
from msmbuilder.cluster import KMeans
from msmbuilder.msm import MarkovStateModel
from msmbuilder.tpt import net_fluxes, paths
import mdtraj as md
from matplotlib.colors import rgb2hex
from nglview import MDTrajTrajectory, NGLWidget
import msmexplorer as msme
from mdentropy.metrics import DihedralMutualInformation
rs = np.random.RandomState(42)
```
Load Trajectories
-----------------
```
trajectories = FsPeptide(verbose=False).get().trajectories
```
Build Markov Model
------------------
```
pipeline = Pipeline([
('dihedrals', DihedralFeaturizer()),
('scaler', RobustScaler()),
('tica', tICA(n_components=2, lag_time=10)),
('kmeans', KMeans(n_clusters=12, random_state=rs)),
('msm', MarkovStateModel(lag_time=1))
])
msm_assignments = pipeline.fit_transform(trajectories)
msm = pipeline.get_params()['msm']
```
Identify Top Folding Pathway
----------------------------
```
sources, sinks = [msm.populations_.argmin()], [msm.populations_.argmax()]
net_flux = net_fluxes(sources, sinks, msm)
paths, _ = paths(sources, sinks, net_flux, num_paths=0)
samples = msm.draw_samples(msm_assignments, n_samples=1000, random_state=rs)
xyz = []
for state in paths[0]:
for traj_id, frame in samples[state]:
xyz.append(trajectories[traj_id][frame].xyz)
pathway = md.Trajectory(np.concatenate(xyz, axis=0), trajectories[0].topology)
pathway.superpose(pathway[0])
```
Calculate Mutual information
----------------------------
```
dmutinf = DihedralMutualInformation(n_bins=3, method='knn', normed=True)
M = dmutinf.partial_transform(pathway)
M -= M.diagonal() * np.eye(*M.shape)
labels = [str(res.index) for res in trajectories[0].topology.residues
if res.name not in ['ACE', 'NME']]
ax = msme.plot_chord(M, threshold=.5, labels=labels,)
from nglview import MDTrajTrajectory, NGLWidget
t = MDTrajTrajectory(pathway)
view = NGLWidget(t)
view
scores = np.real(np.linalg.eig(M)[1][0])
scores -= scores.min()
scores /= scores.max()
cmap = msme.utils.make_colormap(['rawdenim', 'lightgrey', 'pomegranate'])
reslist = [str(res.index) for res in pathway.topology.residues][1:-1]
view.clear()
view.clear_representations()
view.add_cartoon('protein', color='white')
for i, color in enumerate(cmap(scores)):
view.add_representation('ball+stick', reslist[i], color=rgb2hex(color))
view.camera = 'orthographic'
```
| github_jupyter |
```
import math
import pandas as pd
import matplotlib.pyplot as plt
from scipy import stats
import seaborn as sns
from pylab import rcParams
from sklearn.model_selection import train_test_split
from sklearn import preprocessing
from sklearn.metrics import mean_squared_error
import numpy as np
import time
from numpy import arange, sin, pi, random
%matplotlib inline
sns.set(style='whitegrid', palette='muted', font_scale=1.5)
rcParams['figure.figsize'] = 14, 8
RANDOM_SEED = 42
LABELS = ["Normal", "Anomaly"]
from zoo.pipeline.api.keras.layers import Dense, Dropout, Activation, Input
from zoo.pipeline.api.keras.models import Sequential, Model
from bigdl.optim.optimizer import *
```
## Read data from csv
```
df = pd.read_csv("data/CPU_example_new_1.csv")
df.shape
df.head()
```
## Basic sanity check of data and normalization
```
# check for null data
df.isnull().values.any()
# Standard scaling : mean 0, stddev 1
scaler = preprocessing.MinMaxScaler(feature_range=(0, 1))
df['CPU'] = preprocessing.StandardScaler().fit_transform(df['CPU'].values.reshape(-1, 1))
df.head()
# let's explore the distribution of input data
count_classes = pd.value_counts(df['Class'], sort = False)
print(count_classes.count)
count_classes.plot(kind = 'bar', rot=0)
plt.title("Transaction class distribution")
plt.xticks(range(2), LABELS)
plt.xlabel("Class")
plt.ylabel("Frequency");
# Just checking the relative counts
anomaly = df[df.Class == 1]
normal = df[df.Class == 0]
anomaly.shape
normal.shape
anomaly.CPU.describe()
normal.CPU.describe()
```
## Any correlation between time and CPU metrics ?
```
f, (ax1, ax2) = plt.subplots(2, 1, sharex=True)
f.suptitle('Time vs CPU by class')
ax1.scatter(anomaly.Time, anomaly.CPU)
ax1.set_title('Anomaly')
ax2.scatter(normal.Time, normal.CPU)
ax2.set_title('Normal')
plt.xlabel('Time (in Seconds)')
plt.ylabel('CPU')
plt.show()
```
## Feature Re-engineering
```
indexed_df = df.set_index(["Time"])
df = indexed_df.rolling(10).mean() ## mean over rolling window of size 10
df.loc[df['Class'] > 0.5, 'Class'] = 1.0
df.loc[df['Class'] <= 0.5, 'Class'] = 0.0
df = df[np.isfinite(df['CPU'])] ## remove NaNs
df.head()
```
## Split into training and test data
```
X_train, X_test = train_test_split(df.values[:, 1:2], test_size=0.2, random_state=RANDOM_SEED)
Y_train, Y_test = train_test_split(df.values[:, -1], test_size=0.2, random_state=RANDOM_SEED)
X_train.shape
X_test.shape
Y_train.shape
Y_test.shape
class_0 = list(filter(lambda x: x == 0.0, Y_test))
class_1 = list(filter(lambda x: x == 1.0, Y_test))
print(len(class_0), len(class_1))
```
# Build the Model
```
inputs = Input(shape = (X_train.shape[1],))
activation = Activation('relu')(inputs)
hidden1 = Dense(16, activation = 'relu')(activation)
hd1 = Dropout(0.2)(hidden1)
hidden2 = Dense(32, activation = 'relu')(hidden1)
hd2 = Dropout(0.2)(hidden2)
hidden3 = Dense(8, activation = 'relu')(hidden2)
hd3 = Dropout(0.6)(hidden3)
outputs = Dense(output_dim = 1, activation = 'sigmoid')(hd3)
model = Model(inputs, outputs)
# Build the model
# model = Sequential()
# 1st Fully connected layer + DropOut
# model.add(Dense(16, input_shape=(X_train.shape[1],), activation = 'relu'))
# model.add(Dropout(0.2))
# 2nd Fully connected layer + DropOut
# model.add(Dense(32, activation = 'relu'))
# model.add(Dropout(0.2))
# 3rd Fully connected layer + DropOut
# model.add(Dense(32, activation = 'relu'))
# model.add(Dropout(0.6))
# Output a Sigmoid since we need probability (normal / anomaly)
# model.add(Dense(output_dim=1, activation = 'sigmoid'))
# model.summary()
```
# Train
```
log_dir = 'dglogdir'
app_name = 'anomaly-cpu'
tf_model_dump_file_name = 'anomaly-tfdump'
model.save_graph_topology(log_dir + '/' + tf_model_dump_file_name)
model.compile(optimizer=Adam(learningrate = 1e-4),
loss = 'binary_crossentropy',
metrics = ['accuracy'])
model.set_tensorboard(log_dir, app_name)
history = model.fit(X_train, Y_train,
nb_epoch = 8,
batch_size = 256)
train_summary = TrainSummary(log_dir = log_dir, app_name = app_name)
# loss = np.array(train_summary.read_scalar('Loss'))
# df_loss = pd.DataFrame({'Loss' :np.array(loss)})
# df_loss.describe
# plt.plot(loss)
# plt.plot(history.history['val_loss'])
# plt.title('model train vs validation loss')
# plt.ylabel('loss')
# plt.xlabel('epoch')
# plt.legend(['train', 'validation'], loc='upper right')
# plt.show()
```
# Now predict on test set
```
predictions = model.predict(X_test)
first10 = predictions.take(10)
list(map(lambda x: x[0], first10))
p = predictions.collect()
## find classes from probabilities
classes = list(map(lambda x: 1.0 if x[0] > 0.5 else 0.0, p))
## check diff between predicted classes and real classes
df_prediction = pd.DataFrame({'Real Class' :np.array(Y_test)})
df_prediction['Prediction'] = classes
df_prediction[ df_prediction['Real Class'] != df_prediction['Prediction'] ]
scores = model.evaluate(X_test, Y_test)
list(map(lambda x: (x.result, x.total_num, x.method), scores))
```
# Load another test dataset drawn from same distribution
```
df = pd.read_csv("data/CPU_example_test.csv")
df.head()
df.isnull().values.any()
scaler = preprocessing.MinMaxScaler(feature_range=(0, 1))
df['CPU'] = preprocessing.StandardScaler().fit_transform(df['CPU'].values.reshape(-1, 1))
df.head()
indexed_df = df.set_index(["Time"])
df = indexed_df.rolling(10).mean()
df.loc[df['Class'] > 0.5, 'Class'] = 1.0
df.loc[df['Class'] <= 0.5, 'Class'] = 0.0
df = df[np.isfinite(df['CPU'])] ## remove NaNs
df.head()
X_test = df.values[:, 1:2]
Y_test = df.values[:, -1]
scores = model.evaluate(X_test, Y_test)
results = list(map(lambda x: (x.result, x.total_num, x.method), scores))
print(results)
```
# Another test set
```
df = pd.read_csv("data/CPU_example_test_1.csv")
df.isnull().values.any()
scaler = preprocessing.MinMaxScaler(feature_range=(0, 1))
df['CPU'] = preprocessing.StandardScaler().fit_transform(df['CPU'].values.reshape(-1, 1))
df.head()
indexed_df = df.set_index(["Time"])
df = indexed_df.rolling(10).mean()
df.loc[df['Class'] > 0.5, 'Class'] = 1.0
df.loc[df['Class'] <= 0.5, 'Class'] = 0.0
df = df[np.isfinite(df['CPU'])] ## remove NaNs
df.head()
X_test = df.values[:, 1:2]
Y_test = df.values[:, -1]
scores = model.evaluate(X_test, Y_test)
results = list(map(lambda x: (x.result, x.total_num, x.method), scores))
print(results)
```
| github_jupyter |
# Part 2: Deploy a model trained using SageMaker distributed data parallel
Use this notebook after you have completed **Part 1: Distributed data parallel MNIST training with PyTorch and SageMaker's distributed data parallel library** in the notebook pytorch_smdataparallel_mnist_demo.ipynb. To deploy the model you previously trained, you need to create a Sagemaker Endpoint. This is a hosted prediction service that you can use to perform inference.
## Finding the model
This notebook uses a stored model if it exists. If you recently ran a training example that use the `%store%` magic, it will be restored in the next cell.
Otherwise, you can pass the URI to the model file (a .tar.gz file) in the `model_data` variable.
To find the location of model files in the [SageMaker console](https://console.aws.amazon.com/sagemaker/home), do the following:
1. Go to the SageMaker console: https://console.aws.amazon.com/sagemaker/home.
1. Select **Training** in the left navigation pane and then Select **Training jobs**.
1. Find your recent training job and choose it.
1. In the **Output** section, you should see an S3 URI under **S3 model artifact**. Copy this S3 URI.
1. Uncomment the `model_data` line in the next cell that manually sets the model's URI and replace the placeholder value with that S3 URI.
```
# Retrieve a saved model from a previous notebook run's stored variable
%store -r model_data
# If no model was found, set it manually here.
# model_data = 's3://sagemaker-us-west-2-XXX/pytorch-smdataparallel-mnist-2020-10-16-17-15-16-419/output/model.tar.gz'
print("Using this model: {}".format(model_data))
```
## Create a model object
You define the model object by using the SageMaker Python SDK's `PyTorchModel` and pass in the model from the `estimator` and the `entry_point`. The endpoint's entry point for inference is defined by `model_fn` as seen in the following code block that prints out `inference.py`. The function loads the model and sets it to use a GPU, if available.
```
!pygmentize code/inference.py
import sagemaker
role = sagemaker.get_execution_role()
from sagemaker.pytorch import PyTorchModel
model = PyTorchModel(
model_data=model_data,
source_dir="code",
entry_point="inference.py",
role=role,
framework_version="1.6.0",
py_version="py3",
)
```
### Deploy the model on an endpoint
You create a `predictor` by using the `model.deploy` function. You can optionally change both the instance count and instance type.
```
predictor = model.deploy(initial_instance_count=1, instance_type="ml.m4.xlarge")
```
## Test the model
You can test the depolyed model using samples from the test set.
```
# Download the test set
import torchvision
from torchvision import datasets, transforms
from torch.utils.data import DataLoader
from packaging.version import Version
# Set the source to download MNIST data from
TORCHVISION_VERSION = "0.9.1"
if Version(torchvision.__version__) < Version(TORCHVISION_VERSION):
# Set path to data source and include checksum key to make sure data isn't corrupted
datasets.MNIST.resources = [
(
"https://sagemaker-sample-files.s3.amazonaws.com/datasets/image/MNIST/train-images-idx3-ubyte.gz",
"f68b3c2dcbeaaa9fbdd348bbdeb94873",
),
(
"https://sagemaker-sample-files.s3.amazonaws.com/datasets/image/MNIST/train-labels-idx1-ubyte.gz",
"d53e105ee54ea40749a09fcbcd1e9432",
),
(
"https://sagemaker-sample-files.s3.amazonaws.com/datasets/image/MNIST/t10k-images-idx3-ubyte.gz",
"9fb629c4189551a2d022fa330f9573f3",
),
(
"https://sagemaker-sample-files.s3.amazonaws.com/datasets/image/MNIST/t10k-labels-idx1-ubyte.gz",
"ec29112dd5afa0611ce80d1b7f02629c",
),
]
else:
# Set path to data source
datasets.MNIST.mirrors = [
"https://sagemaker-sample-files.s3.amazonaws.com/datasets/image/MNIST/"
]
test_set = datasets.MNIST(
"data",
download=True,
train=False,
transform=transforms.Compose(
[transforms.ToTensor(), transforms.Normalize((0.1307,), (0.3081,))]
),
)
# Randomly sample 16 images from the test set
test_loader = DataLoader(test_set, shuffle=True, batch_size=16)
test_images, _ = iter(test_loader).next()
# inspect the images
import torchvision
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
def imshow(img):
img = img.numpy()
img = np.transpose(img, (1, 2, 0))
plt.imshow(img)
return
# unnormalize the test images for displaying
unnorm_images = (test_images * 0.3081) + 0.1307
print("Sampled test images: ")
imshow(torchvision.utils.make_grid(unnorm_images))
# Send the sampled images to endpoint for inference
outputs = predictor.predict(test_images.numpy())
predicted = np.argmax(outputs, axis=1)
print("Predictions: ")
print(predicted.tolist())
```
## Cleanup
If you don't intend on trying out inference or to do anything else with the endpoint, you should delete it.
```
predictor.delete_endpoint()
```
| github_jupyter |
# TransformerScheduler Class
## How to use it
**Functions defined to be applied on rows of Pandas dataframes** can be wrapped in a TransformerScheduler object in order to be applied on a full Pandas dataframe in a specific order.
TransformerScheduler objects :
- are compatible with the scikit-learn API (they have fit and transform methods).
- can be integrated into an execution Pipeline.
- allows the wrapped functions to be applied in multiprocessing.
A TransformerScheduler object is initialized with a functions_scheduler argument. The functions_scheduler argument is a **list of tuples** containing information about the desired pre-processing functions. Each tuple describe an individual function and should contain the following elements:
1. A function
2. A tuple with the function’s arguments (if no arguments are required, use None or an empty tuple)
3. A column(s) name list returned by the function (if no arguments are required, use None or an empty list)
#### Defining a TransformerScheduler object
```python
from melusine.utils.transformer_scheduler import TransformerScheduler
melusine_transformer = TransformerScheduler(
functions_scheduler=[
(my_function_1, (argument1, argument2), ['return_col_A']),
(my_function_2, None, ['return_col_B', 'return_col_C'])
(my_function_3, (argument1, ), None),
mode='apply_by_multiprocessing',
n_jobs=4)
])
```
#### Parameters
The other parameters of the TransformerScheduler class are:
- **mode** (optional): Define mode to apply function along a row axis (axis=1) If set to ‘apply_by_multiprocessing’, it uses multiprocessing tool to parallelize computation. Possible values are ‘apply’ (default) and ‘apply_by_multiprocessing’
- **n_jobs** (optional): Number of cores used for computation. Default value, 1. Possible values are integers ranging from 1 (default) to the number of cores available for computation
#### Applying the TransformerScheduler object
```python
df = melusine_transformer.fit_transform(df)
```
#### Chaining transformers in a scikit-learn pipeline
Once all the desired functions and transformers have been defined, transformers can be chained in a Scikit-Learn Pipeline. The code below describes the definition of a pipeline:
```python
from sklearn.pipeline import Pipeline
pipeline = Pipeline([
('TransformerName1', TransformerObject1),
('TransformerName2', TransformerObject2),
('TransformerName3', TransformerObject3),
])
df = pipeline.fit_tranform(df)
```
## Example
```
import pandas as pd
df_example = pd.DataFrame({'col': [1,2,3,4,5]})
df_example
```
Let's define three **functions that are applied on rows of dataframes** :
```
def fonction_example_1(row):
return row['col']+1
def fonction_example_x(row,x):
return row['col']+x
def fonction_example_x_y(row,x,y):
return row['col']+x, row['col']+y
from melusine.utils.transformer_scheduler import TransformerScheduler
transformer_example = TransformerScheduler(
functions_scheduler=[
(fonction_example_1, None, ['col_1']),
(fonction_example_x, (2,), ['col_x']),
(fonction_example_x_y, (3,4), ['col_x2', 'col_y'])
],
mode='apply_by_multiprocessing',
n_jobs=4
)
df_example = transformer_example.fit_transform(df_example)
df_example
```
| github_jupyter |
# ETL & Visualization Project
## UV Exposure & Melanoma Rates Correlation in United States
### Extract: UV Exposure and Melanoma Data (csv)
```
# Dependencies and Setup
%matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import json
import pandas as pd
from sqlalchemy import create_engine
# from scipy.stats import linregress
# from scipy import stats
# import pingouin as pg # Install pingouin stats package (pip install pingouin)
# import seaborn as sns # Install seaborn data visualization library (pip install seaborn)
# from scipy.stats import pearsonr
yr_list= [2005, 2006, 2007, 2008, 2009, 2010, 2011, 2012, 2013,
2014, 2015]
# Hide warning messages in notebook
import warnings
warnings.filterwarnings('ignore')
# File to Load
CDI_data_to_load = "CDI_data.csv"
# Read the Population Health Data
CDI_data_pd = pd.read_csv(CDI_data_to_load, encoding="utf-8").fillna(0)
# Display the data table for preview
CDI_data_pd
# Extracting cancer data
topic_sorted_df = CDI_data_pd.groupby('Topic')
topic_sorted_df
cancer_df = topic_sorted_df.get_group('Cancer')
cancer_df
cancer_df = cancer_df.sort_values('LocationDesc')
cancer_df[[]]
new_cancer_df = cancer_df[['LocationAbbr','LocationDesc','Topic',
'Question','DataValueType','DataValue']].copy()
new_cancer_df
# Cancer Incidence Values for Continental United States
incidence_df = new_cancer_df.loc[new_cancer_df['Question'] == 'Invasive melanoma, incidence']
incidence_df
incidence_df = incidence_df.loc[incidence_df['DataValueType'] == 'Average Annual Number']
incidence_df.set_index('LocationAbbr', inplace=True)
# incidence_df.to_csv('incidence.csv')
incidence_df.drop(["AK", "HI", "US"], inplace = True)
incidence_df
# Cancer Mortality Values for Continental United States
mortality_df = new_cancer_df.loc[new_cancer_df['Question'] == 'Melanoma, mortality']
mortality_df
mortality_df = mortality_df.loc[mortality_df['DataValueType'] == 'Average Annual Number']
mortality_df.set_index('LocationAbbr', inplace=True)
#mortality_df.to_csv('mortality.csv')
mortality_df.drop(["AK", "HI", "US"], inplace = True)
mortality_df
# 2nd File to Load
UV_data_to_load = "UV_data.csv"
UV_data_df = pd.read_csv(UV_data_to_load, encoding="utf-8").fillna(0)
# Read the Population Health Data
UV_data_df = pd.read_csv(UV_data_to_load)
# # Display the data table for preview
UV_data_df = UV_data_df.groupby("STATENAME", as_index=False)["UV_Wh/square_meter"].mean()
UV_data_df.set_index('STATENAME', inplace=True)
UV_data_df.to_csv('UV_data_post.csv')
UV_data_df
```
### Load: Database (MongoDB)
```
# Dependencies
import pymongo
import pandas as pd
# Initialize PyMongo to work with MongoDBs
conn = 'mongodb://localhost:27017'
client = pymongo.MongoClient(conn)
```
#### Upload Clean Data to Database
#### 1. Melanoma Incidence Data
```
# Define database and collection
db = client.uv_melanoma_db
collection = db.melanoma_incidence
# Convert the data frame of melanoma incidence data to dictionary
incidence_dict = incidence_df.to_dict("records")
incidence_dict
# Upload melanoma incidence data to MongoDB
for incidence_data in range(len(incidence_dict)):
collection.insert_one(incidence_dict[incidence_data])
# Display the MongoDB records created above
melanoma_incidence_records = db.melanoma_incidence.find()
for melanoma_incidence_record in melanoma_incidence_records:
print(melanoma_incidence_record)
```
#### 2. Melanoma Mortality Data
```
# Define database and collection
db = client.uv_melanoma_db
collection = db.melanoma_mortality
# Convert the data frame of melanoma mortality data to dictionary
mortality_dict = mortality_df.to_dict("records")
mortality_dict
# Upload melanoma mortality data to MongoDB
for mortality_data in range(len(mortality_dict)):
collection.insert_one(mortality_dict[mortality_data])
# Display the MongoDB records created above
melanoma_mortality_records = db.melanoma_mortality.find()
for melanoma_mortality_record in melanoma_mortality_records:
print(melanoma_mortality_record)
```
#### 3. UV Exposure Data
```
# Define database and collection
db = client.uv_melanoma_db
collection = db.uv
# Convert the data frame of UV exposure data to dictionary
UV_dict = UV_data_df.to_dict("records")
UV_dict
# Upload UV exposure data to MongoDB
for UV_data in range(len(UV_dict)):
collection.insert_one(UV_dict[UV_data])
# Display the MongoDB records created above
UV_records = db.uv.find()
for UV_record in UV_records:
print(UV_record)
```
PLAN:
- Dropdown for each states
- Map showing the UV exposure and layers for incidence and mortality
```
# CLEANING WITH PANDAS - DONE
# MONGODB - DONE
# FLASK APP
# VISUALIZATIONS (JS)
# WEB DEPLOYMENT
%load_ext sql
DB_ENDPOINT = "localhost"
DB = 'melanoma_db'
DB_USER = 'postgres'
DB_PASSWORD = [REDACTED]
DB_PORT = '5432'
# postgresql://username:password@host:port/database
conn_string = "postgresql://{}:{}@{}:{}/{}" \
.format(DB_USER, DB_PASSWORD, DB_ENDPOINT, DB_PORT, DB)
print(conn_string)
%sql $conn_string
rds_connection_string = "postgres:password@localhost:5432/melanoma_db"
engine = create_engine(f'postgresql://{rds_connection_string}')
engine.table_names()
pd.read_sql_query('select * from uv', con=engine)
```
| github_jupyter |
# Train you first forecasting model
So you picked the forecasting project. Good for you! Forecasting is an important skill, and it's helpful to learn about the techchnologies out there that can make your life easier. Or worse, depending on how you look at it. ;)
### 1. Download the Kaggle forecasting data
Use the notebook example for this.
### 2. Access your data
```
import pandas as pd
import json
import sagemaker
import boto3
df = pd.read_csv('train.csv')
df.shape
df.head()
```
Each row here is a new point in time, and each column is an energy station. That means that each COLUMN is a unique time series data set. We are going to train our first model on a single column. Then, you can extend it by adding more columns.
```
target = df['ACME']
target.shape
```
Great! We have 5113 obervations. That's well over the 300-limit on DeepAR.
### 3. Create Train and Test Sets
Now, we'll build 2 datasets. One for training, another for testing. Both need to be written to json files, then copied over to S3.
```
def get_split(df, freq='D', split_type = 'train', cols_to_use = ['ACME']):
rt_set = []
# use 70% for training
if split_type == 'train':
lower_bound = 0
upper_bound = round(df.shape[0] * .7)
# use 15% for validation
elif split_type == 'validation':
lower_bound = round(df.shape[0] * .7)
upper_bound = round(df.shape[0] * .85)
# use 15% for test
elif split_type == 'test':
lower_bound = round(df.shape[0] * .85)
upper_bound = df.shape[0]
# loop through columns you want to use
for h in list(df):
if h in cols_to_use:
target_column = df[h].values.tolist()[lower_bound:upper_bound]
date_str = str(df.iloc[0]['Date'])
year = date_str[0:4]
month = date_str[4:6]
date = date_str[7:]
start_dataset = pd.Timestamp("{}-{}-{} 00:00:00".format(year, month, date, freq=freq))
# create a new json object for each column
json_obj = {'start': str(start_dataset),
'target':target_column}
rt_set.append(json_obj)
return rt_set
train_set = get_split(df, freq)
test_set = get_split(df, freq, split_type = 'test')
def write_dicts_to_file(path, data):
with open(path, 'wb') as fp:
for d in data:
fp.write(json.dumps(d).encode("utf-8"))
# fp.write("\n".encode('utf-8'))
write_dicts_to_file('train.json', train_set)
write_dicts_to_file('test.json', test_set)
!aws s3 cp train.json s3://forecasting-do-not-delete/train/train.json
!aws s3 cp test.json s3://forecasting-do-not-delete/test/test.json
```
### 4. Run a SageMaker Training Job
Ok! If everything worked, we should be able to train a model in SageMaker straight away.
```
sess = sagemaker.Session()
region = sess.boto_region_name
image = sagemaker.amazon.amazon_estimator.get_image_uri(region, "forecasting-deepar", "latest")
role = sagemaker.get_execution_role()
estimator = sagemaker.estimator.Estimator(
sagemaker_session=sess,
image_name=image,
role=role,
train_instance_count=1,
train_instance_type='ml.c4.2xlarge',
base_job_name='deepar-electricity-demo',
output_path='s3://forecasting-do-not-delete/output'
)
hyperparameters = {
# frequency interval is once per day
"time_freq": 'D',
"epochs": "400",
"early_stopping_patience": "40",
"mini_batch_size": "64",
"learning_rate": "5E-4",
# let's use the last 30 days for context
"context_length": str(30),
# let's forecast for 30 days
"prediction_length": str(30)
}
estimator.set_hyperparameters(**hyperparameters)
data_channels = {
"train": "s3://forecasting-do-not-delete/train/train.json",
"test": "s3://forecasting-do-not-delete/test/test.json"
}
estimator.fit(inputs=data_channels, wait=True)
```
### 5. Run Inference
If you made it this far, congratulations! None of this is easy. For your next steps, please open up the example notebook under the SageMakerExamples:
- SageMakerExamples/Introduction To Amazon Algorithms/DeepAR-Electricity.
That will walk you through both how to add more timeseries to your model, and how to get inference results out of it.
### 6. Extend Your Solution
Now you're getting forecasts, how will you extend your solution? How good are your forecasts? What about getting forecasts for the other stations? Is your model cognizant of the weather?
Spend your remaining time growing your modeling solution to leverage additional datasets. Then, think through how you'd set this up to run in production.
| github_jupyter |
# How GPflow relates to TensorFlow: tips & tricks
GPflow is built on top of TensorFlow, so it is useful to have some understanding of how TensorFlow works. In particular, TensorFlow's two-stage concept of first building a static compute graph and then executing it for specific input values can cause problems. This notebook aims to help with the most common issues.
```
import gpflow
import tensorflow as tf
import numpy as np
from gpflow import settings
```
## 1. Computation time increases when I create GPflow objects
The following example shows a typical situation when computation time increases proportionally to the number of GPflow objects that are created.
```
for n in range(2, 4):
kernel = gpflow.kernels.RBF(input_dim=1) # This is a gpflow object with tf.Variables inside
x = np.random.randn(n, 1) # gpflow expects rank-2 input matrices, even for D=1
kxx = kernel.K(x) # This is a tensor!
```
**Remember, we operate on a TensorFlow graph!**
Every time we create (build and compile) a new GPflow object, we continue to add more tensors to the graph and change only the reference to them, despite overwriting (in this case) the kernel variable.
So, unnecessary expansion of the graph slows down your computation!
The following examples show how to fix the issue (imagine running this code snippet in `ipython` repeatedly):
```
for n in range(2, 4):
gpflow.reset_default_graph_and_session()
kernel = gpflow.kernels.RBF(1)
x = np.random.randn(n, 1)
kxx = kernel.K(x)
```
Here we were simply resetting the default graph and session using GPflow's `reset_default_graph_and_session()` function. In the next example we explicitly build new `tf.Graph()` and `tf.Session()` objects:
```
for n in range(2, 4):
with tf.Graph().as_default() as graph:
with tf.Session(graph=graph).as_default():
kernel = gpflow.kernels.RBF(1)
x = np.random.randn(n, 1)
kxx = kernel.K(x)
```
In the [Custom mean functions](tailor/external-mean-function.ipynb) notebook we show a real-world example of this idea.
## 2. I want to reuse a model on different data
```
np.random.seed(1)
x = np.random.randn(2, 1)
y = np.random.randn(2, 1)
kernel = gpflow.kernels.RBF(1)
model = gpflow.models.GPR(x, y, kernel)
print(model.compute_log_likelihood())
x_new = np.random.randn(100, 1)
y_new = np.random.randn(100, 1)
```
We can compute the log-likelihood of the model on different data. Note that we didn't change the original model!
```
x_tensor = model.X.parameter_tensor
y_tensor = model.Y.parameter_tensor
model.compute_log_likelihood(feed_dict={x_tensor: x_new, y_tensor: y_new}) # we can still probe the model with the old data
```
We can do the same by permanently updating the values of the dataholders.
```
model.X = x_new
model.Y = y_new
model.compute_log_likelihood()
```
## 3. I want to use external TensorFlow tensors and pass them to a GPflow model
You can pass TensorFlow tensors for any non-trainable parameters of the GPflow objects like DataHolders.
```
np.random.seed(1)
kernel = gpflow.kernels.RBF(1)
likelihood = gpflow.likelihoods.Gaussian()
x_tensor = tf.random_normal((100, 1), dtype=settings.float_type)
y_tensor = tf.random_normal((100, 1), dtype=settings.float_type)
z = np.random.randn(10, 1)
model = gpflow.models.SVGP(x_tensor, y_tensor, kern=kernel, likelihood=likelihood, Z=z)
model.compute_log_likelihood()
```
You can also use TensorFlow variables for trainable objects:
```
z = tf.Variable(np.random.randn(10, 1))
model = gpflow.models.SVGP(x_tensor, y_tensor, kern=kernel, likelihood=likelihood, Z=z)
```
However, in this case you have to initialise them manually, before interacting with a model:
```
session = gpflow.get_default_session()
session.run(z.initializer)
model.compute_log_likelihood()
```
## 4. I want to share parameters between GPflow objects
Sometimes we want to impose a hard-coded structure on the model (for example, if we have a multi-output model where some output dimensions share the same kernel and others don't). Unfortunately we cannot do this after the kernel object is compiled. We have to do it at build time and then manually compile the object.
```
with gpflow.decors.defer_build():
kernels = [gpflow.kernels.RBF(1) for _ in range(3)]
mo_kernels = gpflow.multioutput.kernels.SeparateMixedMok(kernels, W=np.random.randn(3, 4))
mo_kernels.kernels[0].lengthscales = mo_kernels.kernels[1].lengthscales
mo_kernels.compile()
assert mo_kernels.kernels[0].lengthscales is mo_kernels.kernels[1].lengthscales
```
## 5. Optimising my model repeatedly slows down the computation time
The following is an example of bad practice:
```
x = np.random.randn(100, 1)
y = np.random.randn(100, 1)
model = gpflow.models.GPR(x, y, kernel)
optimizer = gpflow.training.AdamOptimizer()
optimizer.minimize(model, maxiter=2)
# Do something with the model
optimizer.minimize(model, maxiter=2)
```
The `minimize()` call creates a bunch of optimisation tensors. Calling `minimize()` again causes the same issue discussed under issue (1).
The correct way of optimising your model without polluting your graph is as follows:
```
kernel = gpflow.kernels.RBF(1)
x = np.random.randn(100, 1)
y = np.random.randn(100, 1)
model = gpflow.models.GPR(x, y, kernel)
optimizer = gpflow.training.AdamOptimizer()
optimizer_tensor = optimizer.make_optimize_tensor(model)
session = gpflow.get_default_session()
for _ in range(2):
session.run(optimizer_tensor)
```
Don't forget to **anchor** your model to the session after optimisation. Then you can continue working with your model.<br/>
```
model.anchor(session)
```
Now, if you need to optimise it again, you can reuse the same optimiser tensor.
```
for _ in range(2):
session.run(optimizer_tensor)
model.anchor(session)
```
## 6. When I try to read parameter values, I'm getting stale values
```
np.random.seed(1)
x = np.random.randn(100, 1)
y = np.random.randn(100, 1)
kernel = gpflow.kernels.RBF(1)
model = gpflow.models.GPR(x, y, kernel)
optimizer = gpflow.training.AdamOptimizer()
optimizer_tensor = optimizer.make_optimize_tensor(model)
```
The initial value before optimisation is:
```
model.kern.lengthscales.value
```
Let's call one step of the optimisation and check the new value of the parameter.
```
gpflow.get_default_session().run(optimizer_tensor)
model.kern.lengthscales.value
```
After optimisation you would expect that the parameters were updated, but they weren't. The trick is that the `value` property returns a cached NumPy value of a parameter.
You can get the value of the optimised parameter by using the `read_value()` method, specifying the correct `session`.
```
model.kern.lengthscales.read_value(session)
```
Alternatively, you can `anchor(session)` your model to the session after the optimisation step. The `anchor()` updates the parameters' cache.
**NOTE:** The `anchor(session)` method is significantly more time-consuming than `read_value(session)`. Do not call it too often unless you need to.
```
model.anchor(session)
model.kern.lengthscales.value
```
## 7. I want to save and load a GPflow model
```
kernel = gpflow.kernels.RBF(1)
x = np.random.randn(100, 1)
y = np.random.randn(100, 1)
model = gpflow.models.GPR(x, y, kernel)
from pathlib import Path
filename = "/tmp/gpr.gpflow"
path = Path(filename)
if path.exists():
path.unlink()
saver = gpflow.saver.Saver()
saver.save(filename, model)
```
You can load the model into a different graph:
```
with tf.Graph().as_default() as graph, tf.Session().as_default():
model_copy = saver.load(filename)
```
Alternatively, you can load the model into the same session:
```
ctx_for_loading = gpflow.saver.SaverContext(autocompile=False)
model_copy = saver.load(filename, context=ctx_for_loading)
model_copy.clear()
model_copy.compile()
```
The difference between the former approach and the latter lies in the TensorFlow name scopes which are used for naming variables. The former approach replicates the instance of the TensorFlow objects (which already exist in the original graph), so we need to load the model into a new graph.
The latter approach uses different name scopes for the variables so that you can dump the model in the same graph.
| github_jupyter |
```
import itertools
import json
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
import numpy as np
import os
import seaborn as sns
import sqlite3
cc = itertools.cycle(sns.color_palette('colorblind'))
color_map = {
"simple_cnn": next(cc),
"resnet": next(cc),
"resnet_pretrained": next(cc),
"resnet_pretrained_embeddings": next(cc),
"resnet_no_pool": next(cc),
"unseen": next(cc),
"geo_mean_seen": next(cc),
"geo_mean_unseen": next(cc)
}
model_name_map = {
"simple_cnn": "2-Layer CNN",
"resnet": "ResNet-18",
"resnet_pretrained": "ResNet-18 (CIFAR-10 Pretrained)",
"resnet_pretrained_embeddings": "ResNet-18 (CIFAR-10 Embeddings)",
"resnet_no_pool": "ResNet-18 (Avg. Pooling Layer Removed)",
}
def get_timestamps(input_name, model_name, output_name, experiment_name):
conn = sqlite3.connect('../results/results_lookup.db')
c = conn.cursor()
timestamps = []
for row in c.execute('''
SELECT * FROM experiments
WHERE input=?
AND output=?
AND model=?
AND experiment=?
AND timestamp>'2020-01-21_00:00:00_000000'
''', (input_name, output_name, model_name, experiment_name)):
timestamps.append(row[0])
# print(str(len(timestamps)) + " experiment entries found")
return timestamps[-30:]
def get_results(timestamps):
train_results, test_results = [], []
for trial_timestamp in timestamps:
try:
with open('../results/' + trial_timestamp + "/train.json") as train_fp:
train_results.append(json.load(train_fp))
with open('../results/' + trial_timestamp + "/test.json") as test_fp:
test_results.append(json.load(test_fp))
except:
pass
print(len(test_results))
return train_results, test_results
def get_keep_pcts(test_results):
return sorted(list(test_results[0].keys()))[::-1]
def get_label_names(test_results, keep_pcts):
if "class_3_name" not in test_results[0][keep_pcts[0]][0]:
return test_results[0][keep_pcts[0]][0]["class_1_name"].capitalize(), \
test_results[0][keep_pcts[0]][0]["class_2_name"].capitalize()
else:
return test_results[0][keep_pcts[0]][0]["class_1_name"].capitalize(), \
test_results[0][keep_pcts[0]][0]["class_2_name"].capitalize(), \
test_results[0][keep_pcts[0]][0]["class_3_name"].capitalize()
def get_left_out_accs(test_results, keep_pcts):
left_out_final_num_accs_raw = np.array([[trial_test_results[keep_pct][-1]["left_out_num_acc"] for keep_pct in keep_pcts] for trial_test_results in test_results])
left_out_final_num_accs = np.mean(left_out_final_num_accs_raw, axis=0)
left_out_final_num_accs_std = np.std(left_out_final_num_accs_raw, axis=0)
left_out_final_col_accs_raw = np.array([[trial_test_results[keep_pct][-1]["left_out_col_acc"] for keep_pct in keep_pcts] for trial_test_results in test_results])
left_out_final_col_accs = np.mean(left_out_final_col_accs_raw, axis=0)
left_out_final_col_accs_std = np.std(left_out_final_col_accs_raw, axis=0)
left_out_final_loc_accs_raw = np.array([[trial_test_results[keep_pct][-1]["left_out_loc_acc"] for keep_pct in keep_pcts] for trial_test_results in test_results])
left_out_final_loc_accs = np.mean(left_out_final_loc_accs_raw, axis=0)
left_out_final_loc_accs_std = np.std(left_out_final_loc_accs_raw, axis=0)
return (left_out_final_num_accs, left_out_final_num_accs_std), \
(left_out_final_col_accs, left_out_final_col_accs_std), \
(left_out_final_loc_accs, left_out_final_loc_accs_std)
def get_non_left_out_accs(test_results, keep_pcts):
non_left_out_final_num_accs_raw = np.array([[trial_test_results[keep_pct][-1]["non_left_out_num_acc"] for keep_pct in keep_pcts] for trial_test_results in test_results])
non_left_out_final_num_accs = np.mean(non_left_out_final_num_accs_raw, axis=0)
non_left_out_final_num_accs_std = np.std(non_left_out_final_num_accs_raw, axis=0)
non_left_out_final_col_accs_raw = np.array([[trial_test_results[keep_pct][-1]["non_left_out_col_acc"] for keep_pct in keep_pcts] for trial_test_results in test_results])
non_left_out_final_col_accs = np.mean(non_left_out_final_col_accs_raw, axis=0)
non_left_out_final_col_accs_std = np.std(non_left_out_final_col_accs_raw, axis=0)
non_left_out_final_loc_accs_raw = np.array([[trial_test_results[keep_pct][-1]["non_left_out_loc_acc"] for keep_pct in keep_pcts] for trial_test_results in test_results])
non_left_out_final_loc_accs = np.mean(non_left_out_final_loc_accs_raw, axis=0)
non_left_out_final_loc_accs_std = np.std(non_left_out_final_loc_accs_raw, axis=0)
return (non_left_out_final_num_accs, non_left_out_final_num_accs_std), \
(non_left_out_final_col_accs, non_left_out_final_col_accs_std), \
(non_left_out_final_loc_accs, non_left_out_final_loc_accs_std)
def get_directory(input_name, model_name, output_name, experiment_name):
directory = "../plots/{}/{}/{}/{}/".format(experiment_name, output_name, input_name, model_name)
if not os.path.exists(directory):
os.makedirs(directory)
return directory
def plot(keep_pcts, label_names, left_out_accs, non_left_out_accs, directory, model_name):
sns.set()
sns.set_style('ticks')
sns.set_context("poster")
for i in range(len(label_names)):
label_name = label_names[i]
left_out_final_accs, left_out_final_accs_std = left_out_accs[i]
non_left_out_final_accs, non_left_out_final_accs_std = non_left_out_accs[i]
fig, ax = plt.subplots(figsize=(8, 6))
ax.errorbar(keep_pcts[:0:-1], non_left_out_final_accs[:0:-1],
yerr=non_left_out_final_accs_std[:0:-1], zorder=10,
fmt='-s', clip_on=False, linewidth=4, ls='--', color=color_map[model_name])
ax.errorbar(keep_pcts[:0:-1], left_out_final_accs[:0:-1],
yerr=left_out_final_accs_std[:0:-1], zorder=10,
fmt='-o', clip_on=False, linewidth=4, color=color_map[model_name])
ax.set_ylim(0, 1)
ax.set_xlabel('Keep %')
ax.set_ylabel('Test Accuracy')
ax.set_title((r'Final Hold-Out $\bf {}$ Classification' + '\nAccuracy v. Keep %').format(label_name), fontsize=25, y=1.08)
ax.legend(["Seen Combinations", "Unseen Combinations"], loc='best', fontsize=20, framealpha=1,)
# Set minor tick locations.
ax.yaxis.set_minor_locator(ticker.MultipleLocator(0.2))
ax.xaxis.set_minor_locator(ticker.MultipleLocator(1))
# Set grid to use minor tick locations.
ax.grid(which = 'both')
plt.subplots_adjust(top=0.8, bottom=0.2, left=0.17)
plt.xticks(keep_pcts[:0:-2], labels=[str(i + 1) + "/" + str(len(keep_pcts)) for i in range(0, len(keep_pcts), 2)])
plt.yticks([x / 10. for x in range(0, 11, 2)])
for item in ([ax.xaxis.label, ax.yaxis.label] +
ax.get_xticklabels() + ax.get_yticklabels()):
item.set_fontsize(20)
fig.savefig(directory + "hold_out_{}_acc.pdf".format(label_name.lower()), dpi=500)
plt.show()
fig.clf()
def plot_geo_mean(keep_pcts, test_results, directory, model_name):
sns.set()
sns.set_style('ticks')
sns.set_context("poster")
left_out_final_num_accs_raw = np.array([[trial_test_results[keep_pct][-1]["left_out_num_acc"] for keep_pct in keep_pcts] for trial_test_results in test_results])
left_out_final_col_accs_raw = np.array([[trial_test_results[keep_pct][-1]["left_out_col_acc"] for keep_pct in keep_pcts] for trial_test_results in test_results])
left_out_final_loc_accs_raw = np.array([[trial_test_results[keep_pct][-1]["left_out_loc_acc"] for keep_pct in keep_pcts] for trial_test_results in test_results])
non_left_out_final_num_accs_raw = np.array([[trial_test_results[keep_pct][-1]["non_left_out_num_acc"] for keep_pct in keep_pcts] for trial_test_results in test_results])
non_left_out_final_col_accs_raw = np.array([[trial_test_results[keep_pct][-1]["non_left_out_col_acc"] for keep_pct in keep_pcts] for trial_test_results in test_results])
non_left_out_final_loc_accs_raw = np.array([[trial_test_results[keep_pct][-1]["non_left_out_loc_acc"] for keep_pct in keep_pcts] for trial_test_results in test_results])
left_out_geo_mean_acc_raw = np.sqrt(left_out_final_num_accs_raw * left_out_final_col_accs_raw * left_out_final_loc_accs_raw)
left_out_geo_mean_acc = np.mean(left_out_geo_mean_acc_raw, axis=0)
left_out_geo_mean_acc_std = np.std(left_out_geo_mean_acc_raw, axis=0)
non_left_out_geo_mean_acc_raw = np.sqrt(non_left_out_final_num_accs_raw * non_left_out_final_col_accs_raw * non_left_out_final_loc_accs_raw)
non_left_out_geo_mean_acc = np.mean(non_left_out_geo_mean_acc_raw, axis=0)
non_left_out_geo_mean_acc_std = np.std(non_left_out_geo_mean_acc_raw, axis=0)
fig, ax = plt.subplots(figsize=(8, 6))
ax.errorbar(keep_pcts[:0:-1], non_left_out_geo_mean_acc[:0:-1],
yerr=non_left_out_geo_mean_acc_std[:0:-1], zorder=10,
fmt='-s', clip_on=False, linewidth=4, ls='--', color=color_map[model_name])
ax.errorbar(keep_pcts[:0:-1], left_out_geo_mean_acc[:0:-1],
yerr=left_out_geo_mean_acc_std[:0:-1], zorder=10,
fmt='-o', clip_on=False, linewidth=4, color=color_map[model_name])
ax.set_ylim(0, 1)
ax.set_xlabel('Keep %')
ax.set_ylabel('Test Accuracy')
ax.set_title(r'$\bf{Geometric}$ $\bf{Mean}$ of' + '\nTest Accuracy v. Keep %', fontsize=25, y=1.08)
ax.legend(["Seen Combinations", "Unseen Combinations"], loc='best', fontsize=20, framealpha=1,)
# Set minor tick locations.
ax.yaxis.set_minor_locator(ticker.MultipleLocator(0.2))
ax.xaxis.set_minor_locator(ticker.MultipleLocator(1))
# Set grid to use minor tick locations.
ax.grid(which = 'both')
plt.xticks(keep_pcts[:0:-2], labels=[str(i + 1) + "/" + str(len(keep_pcts)) for i in range(0, len(keep_pcts), 2)])
# plt.yticks([y / 4. for y in range(0, 5)])
plt.yticks([x / 10. for x in range(0, 11, 2)])
for item in ([ax.xaxis.label, ax.yaxis.label] +
ax.get_xticklabels() + ax.get_yticklabels()):
item.set_fontsize(20)
plt.subplots_adjust(top=0.8, bottom=0.2, left=0.17)
fig.savefig(directory + "geo_mean_acc.pdf", dpi=500)
plt.show()
fig.clf()
def run_all(input_name, model_name, output_name, experiment_name):
timestamps = get_timestamps(input_name, model_name, output_name, experiment_name)
_, test_results = get_results(timestamps)
keep_pcts = get_keep_pcts(test_results)
label_names = get_label_names(test_results, keep_pcts)
left_out_accs = get_left_out_accs(test_results, keep_pcts)
non_left_out_accs = get_non_left_out_accs(test_results, keep_pcts)
directory = get_directory(input_name, model_name, output_name, experiment_name)
plot(keep_pcts, label_names, left_out_accs, non_left_out_accs, directory, model_name)
plot_geo_mean(keep_pcts, test_results, directory, model_name)
input_name = "left_out_varied_location_mnist"
model_name = "resnet_no_pool"
output_name = "keep_pct_readout_dump"
experiment_name = "keep_pct_readout_9_class_weighted_save"
for input_name, experiment_name in [("left_out_colored_location_mnist", "keep_pct_readout_3_task")]:
for model_name in [\
# "simple_cnn",
"resnet",
# "resnet_pretrained",
# "resnet_pretrained_embeddings",
# "resnet_no_pool",
]:
print("Task:", input_name, experiment_name)
print("Architecture:", model_name)
run_all(input_name, model_name, output_name, experiment_name)
def setup_fig(n, ylabel, title):
# plt.clf()
fig, ax = plt.subplots(figsize=(8, 6))
ax.set_ylim(0, 1)
ax.set_xlabel('Keep %')
ax.set_ylabel(ylabel)
ax.set_title(title, fontsize=25, y=1.08)
# Set minor tick locations.
ax.yaxis.set_minor_locator(ticker.MultipleLocator(0.2))
# ax.xaxis.set_minor_locator(ticker.MultipleLocator(1/9))
ax.xaxis.set_minor_locator(ticker.MultipleLocator(1./n))
# Set grid to use minor tick locations.
ax.grid(which = 'both')
ax.set_yticks([x / 10. for x in range(0, 11, 2)])
for item in ([ax.xaxis.label, ax.yaxis.label] +
ax.get_xticklabels() + ax.get_yticklabels()):
item.set_fontsize(20)
return fig, ax
model_names = \
[
# "simple_cnn",
"resnet",
# "resnet_pretrained",
# "resnet_pretrained_embeddings",
# "resnet_no_pool"
]
task_to_labels = {
"left_out_colored_mnist": ("Shape", "Color"),
"left_out_varied_location_mnist": ("Shape", "Position")
}
keep_pct_map = {
"left_out_colored_mnist": ['1', '0.9', '0.8', '0.7', '0.6', '0.5', '0.4', '0.3', '0.2', '0.1'],
"left_out_varied_location_mnist": ['1.0', '0.8888888888888888', '0.7777777777777778', '0.6666666666666666', '0.5555555555555556', '0.4444444444444444', '0.3333333333333333', '0.2222222222222222', '0.1111111111111111']
}
model_to_name = {"simple_cnn" : "2-Layer CNN",
"resnet": "ResNet-18",
"resnet_pretrained": "ResNet-18 (CIFAR10 Pretrained)",
"resnet_pretrained_embeddings": "ResNet-18 (CIFAR10 Embeddings)",
"resnet_no_pool": "ResNet-18 (No Avg. Pooling)"
}
def combined_plot():
# for input_name, experiment_name, n in [("left_out_colored_mnist", "keep_pct_readout_default_v2_weighted_save", 10),
# ("left_out_varied_location_mnist", "keep_pct_readout_9_class_v2_weighted_save", 9)]:
for input_name, experiment_name in [("left_out_colored_mnist", "keep_pct_readout_color_only"),
("left_out_colored_mnist", "keep_pct_readout_shape_only"),
("left_out_varied_location_mnist", "keep_pct_readout_loc_only"),
("left_out_varied_location_mnist", "keep_pct_readout_shape_only")]:
shape_fig, shape_ax = setup_fig(n, "Test Accuracy", (r'Final Hold-Out $\bf {}$ Classification' + '\nAccuracy v. Keep %').format(task_to_labels[input_name][0]))
color_fig, color_ax = setup_fig(n, "Test Accuracy", (r'Final Hold-Out $\bf {}$ Classification' + '\nAccuracy v. Keep %').format(task_to_labels[input_name][1]))
geo_m_fig, geo_m_ax = setup_fig(n, "Geometric Mean", r'$\bf{Geometric}$ $\bf{Mean}$ of' + '\nTest Accuracy v. Keep %')
for ax in [shape_ax, color_ax, geo_m_ax]:
ax.set_xticks(np.array(keep_pct_map[input_name][:0:-2], dtype=float), minor=False)
ax.set_xticklabels(np.array([str(i + 1) + "/" + str(len(keep_pct_map[input_name])) for i in range(0, len(keep_pct_map[input_name]), 2)]), minor=False)
for model_name in model_names:
# break
timestamps = get_timestamps(input_name, model_name, output_name, experiment_name)
_, test_results = get_results(timestamps)
keep_pcts = get_keep_pcts(test_results)
directory = get_directory(input_name, model_name, output_name, experiment_name)
left_out_final_num_accs_raw = np.array([[trial_test_results[keep_pct][-1]["left_out_num_acc"] for keep_pct in keep_pcts] for trial_test_results in test_results])
left_out_final_num_accs = np.mean(left_out_final_num_accs_raw, axis=0)
left_out_final_num_accs_std = np.std(left_out_final_num_accs_raw, axis=0)
left_out_final_col_accs_raw = np.array([[trial_test_results[keep_pct][-1]["left_out_col_acc"] for keep_pct in keep_pcts] for trial_test_results in test_results])
left_out_final_col_accs = np.mean(left_out_final_col_accs_raw, axis=0)
left_out_final_col_accs_std = np.std(left_out_final_col_accs_raw, axis=0)
left_out_geo_mean_acc_raw = np.sqrt(left_out_final_num_accs_raw * left_out_final_col_accs_raw)
left_out_geo_mean_acc = np.mean(left_out_geo_mean_acc_raw, axis=0)
left_out_geo_mean_acc_std = np.std(left_out_geo_mean_acc_raw, axis=0)
shape_ax.errorbar([float(x) for x in keep_pcts[:0:-1]], left_out_final_num_accs[:0:-1],
yerr=left_out_final_num_accs_std[:0:-1], zorder=10,
fmt='-o', clip_on=False, linewidth=4, color=color_map[model_name],
label=model_to_name[model_name])
color_ax.errorbar([float(x) for x in keep_pcts[:0:-1]], left_out_final_col_accs[:0:-1],
yerr=left_out_final_col_accs_std[:0:-1], zorder=10,
fmt='-o', clip_on=False, linewidth=4, color=color_map[model_name],
label=model_to_name[model_name])
location_ax.errorbar([float(x) for x in keep_pcts[:0:-1]], left_out_final_loc_accs[:0:-1],
yerr=left_out_final_loc_accs_std[:0:-1], zorder=10,
fmt='-o', clip_on=False, linewidth=4, color=color_map[model_name],
label=model_to_name[model_name])
geo_m_ax.errorbar([float(x) for x in keep_pcts[:0:-1]], left_out_geo_mean_acc[:0:-1],
yerr=left_out_geo_mean_acc_std[:0:-1], zorder=10,
fmt='-o', clip_on=False, linewidth=4, color=color_map[model_name],
label=model_to_name[model_name])
# shape_ax.legend(fontsize=16)
shape_fig.subplots_adjust(top=0.8, bottom=0.2, left=0.17)
shape_fig.savefig("shape-" + input_name + ".pdf")
shape_fig.clf()
# color_ax.legend(fontsize=16)
color_fig.subplots_adjust(top=0.8, bottom=0.2, left=0.17)
color_fig.savefig("color-" + input_name + ".pdf")
color_fig.clf()
# geo_m_ax.legend(fontsize=16, framealpha=1, loc='upper center', bbox_to_anchor=(0.5, -0.05),)
geo_m_fig.subplots_adjust(top=0.8, bottom=0.2, left=0.17)
geo_m_fig.savefig("geo_m-" + input_name + ".pdf")
geo_m_fig.clf()
plt.show()
combined_plot()
# def combined_plot():
# for input_name, experiment_name, n in [("left_out_varied_location_mnist", "keep_pct_readout_9_class_weighted_save", 9)]:
# shape_fig, shape_ax = setup_fig(n, "Test Accuracy", (r'Final Hold-Out $\bf {}$ Classification' + '\nAccuracy v. Keep %').format(task_to_labels[input_name][0]))
# color_fig, color_ax = setup_fig(n, "Test Accuracy", (r'Final Hold-Out $\bf {}$ Classification' + '\nAccuracy v. Keep %').format(task_to_labels[input_name][1]))
# geo_m_fig, geo_m_ax = setup_fig(n, "Geometric Mean", r'$\bf{Geometric}$ $\bf{Mean}$ of' + '\nTest Accuracy v. Keep %')
# for ax in [shape_ax, color_ax, geo_m_ax]:
# ax.set_xticks(np.array(keep_pct_map[input_name][:0:-2], dtype=float), minor=False)
# ax.set_xticklabels(np.array([str(i + 1) + "/" + str(len(keep_pct_map[input_name])) for i in range(0, len(keep_pct_map[input_name]), 2)]), minor=False)
# for model_name in model_names:
# # break
# timestamps = get_timestamps(input_name, model_name, output_name, experiment_name)
# _, test_results = get_results(timestamps)
# keep_pcts = get_keep_pcts(test_results)
# directory = get_directory(input_name, model_name, output_name, experiment_name)
# left_out_final_num_accs_raw = np.array([[trial_test_results[keep_pct][-1]["left_out_num_acc"] for keep_pct in keep_pcts] for trial_test_results in test_results])
# left_out_final_num_accs = np.mean(left_out_final_num_accs_raw, axis=0)
# left_out_final_num_accs_std = np.std(left_out_final_num_accs_raw, axis=0)
# left_out_final_col_accs_raw = np.array([[trial_test_results[keep_pct][-1]["left_out_col_acc"] for keep_pct in keep_pcts] for trial_test_results in test_results])
# left_out_final_col_accs = np.mean(left_out_final_col_accs_raw, axis=0)
# left_out_final_col_accs_std = np.std(left_out_final_col_accs_raw, axis=0)
# left_out_geo_mean_acc_raw = np.sqrt(left_out_final_num_accs_raw * left_out_final_col_accs_raw)
# left_out_geo_mean_acc = np.mean(left_out_geo_mean_acc_raw, axis=0)
# left_out_geo_mean_acc_std = np.std(left_out_geo_mean_acc_raw, axis=0)
# print(keep_pcts[:0:-1])
# shape_ax.errorbar([float(x) for x in keep_pcts[:0:-1]], left_out_final_num_accs[:0:-1],
# yerr=left_out_final_num_accs_std[:0:-1], zorder=10,
# fmt='-o', clip_on=False, linewidth=4, color=color_map[model_name])
# color_ax.errorbar([float(x) for x in keep_pcts[:0:-1]], left_out_final_col_accs[:0:-1],
# yerr=left_out_final_col_accs_std[:0:-1], zorder=10,
# fmt='-o', clip_on=False, linewidth=4, color=color_map[model_name])
# geo_m_ax.errorbar([float(x) for x in keep_pcts[:0:-1]], left_out_geo_mean_acc[:0:-1],
# yerr=left_out_geo_mean_acc_std[:0:-1], zorder=10,
# fmt='-o', clip_on=False, linewidth=4, color=color_map[model_name])
# plt.show()
# shape_fig.clf()
# color_fig.clf()
# geo_m_fig.clf()
# combined_plot()
```
| github_jupyter |
# Compound Charts
Altair provides a concise API for creating multi-panel and layered charts, and we'll mention three of them explicitly here:
- Layering
- Horizontal Concatenation
- Vertical Concatenation
- Repeat Charts
We'll explore those briefly here.
```
import altair as alt
# Altair plots render by default in JupyterLab and nteract
# Uncomment/run this line to enable Altair in the classic notebook (not in JupyterLab)
# alt.renderers.enable('notebook')
# Uncomment/run this line to enable Altair in Colab
# alt.renderers.enable('colab')
```
## Layering
Layering lets you put layer multiple marks on a single Chart. One common example is creating a plot with both points and lines representing the same data.
Let's use the ``stocks`` data for this example:
```
from vega_datasets import data
stocks = data.stocks()
stocks.head()
```
Here is a simple line plot for the stocks data:
```
alt.Chart(stocks).mark_line().encode(
x='date:T',
y='price:Q',
color='symbol:N'
)
```
and here is the same plot with a ``circle`` mark:
```
alt.Chart(stocks).mark_circle().encode(
x='date:T',
y='price:Q',
color='symbol:N'
)
```
We can layer these two plots together using a ``+`` operator:
```
lines = alt.Chart(stocks).mark_line().encode(
x='date:T',
y='price:Q',
color='symbol:N'
)
points = alt.Chart(stocks).mark_circle().encode(
x='date:T',
y='price:Q',
color='symbol:N'
)
lines + points
```
This ``+`` is just a shortcut to the ``alt.layer()`` function, which does the same thing:
```
alt.layer(lines, points)
```
One pattern we'll use often is to create a base chart with the common elements, and add together two copies with just a single change:
```
base = alt.Chart(stocks).encode(
x='date:T',
y='price:Q',
color='symbol:N'
)
base.mark_line() + base.mark_circle()
```
## Horizontal Concatenation
Just as we can layer charts on top of each other, we can concatenate horizontally using ``alt.hconcat``, or equivalently the ``|`` operator:
```
base.mark_line() | base.mark_circle()
alt.hconcat(base.mark_line(),
base.mark_circle())
```
This can be most useful for creating multi-panel views; for example, here is the iris dataset:
```
iris = data.iris()
iris.head()
base = alt.Chart(iris).mark_point().encode(
x='petalWidth',
y='petalLength',
color='species'
)
base | base.encode(x='sepalWidth')
```
## Vertical Concatenation
Vertical concatenation looks a lot like horizontal concatenation, but using either the ``alt.hconcat()`` function, or the ``&`` operator:
```
base & base.encode(y='sepalWidth')
```
## Repeat Chart
Because it is such a common pattern to horizontally and vertically concatenate charts while changing one encoding, Altair offers a shortcut for this, using the ``repeat()`` operator.
```
import altair as alt
from vega_datasets import data
iris = data.iris()
fields = ['petalLength', 'petalWidth', 'sepalLength', 'sepalWidth']
alt.Chart(iris).mark_point().encode(
alt.X(alt.repeat("column"), type='quantitative'),
alt.Y(alt.repeat("row"), type='quantitative'),
color='species'
).properties(
width=200,
height=200
).repeat(
row=fields,
column=fields[::-1]
).interactive()
```
This repeat API is still not as streamlined as it could be, but we'll be working on that.
| github_jupyter |
```
import reltest
import reltest.util as util
from reltest.mctest import MCTestPSI, MCTestCorr
from reltest.mmd import MMD_U, med_heuristic
from reltest import kernel
from helper import download_to, summary, fid
import numpy as np
import os
import logging
import pandas as pd
logging.getLogger().setLevel(logging.INFO)
%load_ext autoreload
%autoreload 2
```
#### Download Dataset (Default = ```False```)
To download dataset change ```download``` to ```True```.
```
download = False
feature_url = "http://ftp.tuebingen.mpg.de/pub/is/wittawat/kmod_share/problems/celeba/inception_features/"
feature_dir = os.getcwd() + "/celeba_features"
celeba_classes = ['gen_smile', 'gen_nonsmile', 'ref_smile', 'ref_nonsmile']
mix_ratio = [1,1,0,0]
inception_dim = 2048
## Normalised ratio
celeb_class_ratio = dict(zip(celeba_classes, mix_ratio/np.sum(mix_ratio)))
if download:
os.mkdir(feature_dir)
for celeba_class in celeba_classes:
filename= '{}.npy'.format(celeba_class)
npy_path = os.path.join(feature_dir, filename)
url_path = os.path.join(feature_url, filename)
download_to(url_path,npy_path)
```
### Problem Setting
```
class CelebASampler():
def __init__(self, model_classes_mix, ref_classes_mix):
celeba_features = []
for celeba_class in celeba_classes:
filename= '{}.npy'.format(celeba_class)
npy_path = os.path.join(feature_dir, filename)
celeba_features.append(np.load(npy_path))
self.celeba_features = dict(zip(celeba_classes, celeba_features))
self.model_classes_mix = model_classes_mix
self.ref_classes_mix = ref_classes_mix
def sample(self, n_samples, seed):
## DISJOINT SET
model_features = {}
ref_samples = []
with util.NumpySeedContext(seed=seed):
## FOR EACH CELEBA CLASS
for key, features in self.celeba_features.items():
# CALCULATE HOW MUCH SHOULD BE IN THE REFERENCE POOL
n_ref_samples = int(np.round(self.ref_classes_mix[key] * n_samples))
random_features = np.random.permutation(features)
## FOR THE CANDIDATE MODELS
model_features[key] = random_features[n_ref_samples:]
## FOR THE REFERENCE
ref_samples.append(random_features[:n_ref_samples])
## samples for models
model_samples = []
for j,class_ratios in enumerate(model_classes_mix):
model_class_samples = []
for i, data_class in enumerate(class_ratios.keys()):
n_class_samples = int(np.round(class_ratios[data_class] * n_samples))
seed_class = i*n_samples+seed*j
with util.NumpySeedContext(seed=seed_class):
indices = np.random.choice(model_features[data_class].shape[0], n_class_samples)
model_class_samples.append(model_features[data_class][indices])
class_samples = dict(zip(class_ratios.keys(),model_class_samples))
model_class_stack = np.vstack(list(class_samples.values()))
model_samples.append(model_class_stack)
assert model_class_stack.shape[0] == n_samples, "Sample size mismatch: {0} instead of {1}".format(samples.shape[0],n)
with util.NumpySeedContext(seed=seed+5):
ref_samples = np.random.permutation(np.vstack(ref_samples))
model_samples = [np.random.permutation(samples) for samples in model_samples]
assert ref_samples.shape[0] == n_samples, "Sample size mismatch: {0} instead of {1}".format(samples.shape[0],n)
return model_samples, ref_samples
model_mixtures = [
[0.50,0.50,0.0,0.0], # Model 5
[0.0,0.0,0.60,0.40], # Model 2
[0.0,0.0,0.40,0.60], # Model 3
[0.0,0.0,0.51,0.49], # Model 4
[0.0,0.0,0.52,0.48], # Model 1
]
ref_mixtures = [[0,0,0.5,0.5]]
def class_mix_ratio(classes, mixtures):
return [dict(zip(classes, mixture/np.sum(mixture))) for mixture in mixtures]
model_classes_mix = class_mix_ratio(celeba_classes, model_mixtures)
ref_classes_mix = class_mix_ratio(celeba_classes, ref_mixtures)[0]
n_samples = 2000
n_models = len(model_mixtures)
setting = {'n': n_models,
'dim':inception_dim}
celeba = CelebASampler(model_classes_mix,ref_classes_mix)
n_trials = 100
psi_res = []
cor_res = []
fid_scores = []
for seed in range(n_trials):
## Sample from model
model_samples, ref_samples = celeba.sample(n_samples, seed)
psiTest = MCTestPSI(ref_samples)
corTest = MCTestCorr(ref_samples)
## Perform Test
bandwidth = med_heuristic(model_samples, ref_samples,
subsample=1000)
mmd_u = MMD_U(kernel.KIMQ())
psi_res.append(psiTest.perform_tests(model_samples,mmd_u))
cor_res.append(corTest.perform_tests(model_samples, mmd_u, split=0.5, density=False, correction=0))
## Calculate FID
fid_score = []
for j, model_samples in enumerate(model_samples):
fid_score.append(fid(model_samples,ref_samples))
fid_scores.append(fid_score)
n_models = len(model_mixtures)
print(summary(psi_res,n_models))
print(summary(cor_res,n_models))
fid_results = { 'ind_sel':np.zeros(n_models),
'av_fid': np.zeros(n_models)}
for trial in fid_scores:
l_fids=[model1 for model1 in trial]
ind_sel = np.argmin(l_fids)
fid_results['ind_sel'][ind_sel] += 1/len(fid_scores)
fid_results['av_fid'] =fid_results['av_fid']+ np.array(l_fids)/len(fid_scores)
fids=[]
for trial in fid_scores:
fids.append([model1 for model1 in trial])
print(np.mean(np.array(fids),axis=0))
print(np.median(np.array(fids),axis=0))
dict_res = {
'model_mix':model_mixtures,
'test_psi_sel':summary(psi_res,n_models)['av_sel'],
'test_psi_rej':summary(psi_res,n_models)['av_rej'],
'test_cor_sel':summary(cor_res,n_models)['av_sel'],
'test_cor_rej':summary(cor_res,n_models)['av_rej'],
'fid_results_sel':fid_results['ind_sel'],
'fid_results_av':fid_results['av_fid'],
'fid_results_std':np.std(np.array(fids),axis=0),
}
pd.DataFrame.from_dict(dict_res)
```
| github_jupyter |
# Training Gradient Boosting Models on Large Datasets with CatBoost
[](https://colab.research.google.com/github/catboost/tutorials/blob/master/events/2019_10_30_odsc_west.ipynb)
In this tutorial we will use dataset Amazon Employee Access Challenge from [Kaggle](https://www.kaggle.com) competition for our experiments. Data can be downloaded [here](https://www.kaggle.com/c/amazon-employee-access-challenge/data).
## Libraries installation
```
#!pip install --user --upgrade catboost
#!pip install --user --upgrade ipywidgets
#!pip install shap
#!pip install sklearn
#!jupyter nbextension enable --py widgetsnbextension
import os
import pandas as pd
import numpy as np
np.set_printoptions(precision=4)
import catboost
print(catboost.__version__)
```
## Reading the data
```
from catboost.datasets import amazon
# If you have "URLError: SSL: CERTIFICATE_VERIFY_FAILED" uncomment next two lines:
# import ssl
# ssl._create_default_https_context = ssl._create_unverified_context
# If you have another error:
# Download datasets from bit.ly/2MSENXn and uncomment next three lines:
# def amazon():
# return (pd.read_csv('train.csv', sep=',', header='infer'),
# pd.read_csv('test.csv', sep=',', header='infer'))
(train_df, test_df) = amazon()
train_df.head()
```
## Preparing the data
Label values extraction
```
y = train_df.ACTION
X = train_df.drop('ACTION', axis=1)
```
Categorical features declaration
```
cat_features = list(range(0, X.shape[1]))
print(cat_features)
cat_features = train_df.columns[1:].tolist()
cat_features
```
Looking on label balance in dataset
```
print('Labels: {}'.format(set(y)))
print('Zero count = {}, One count = {}'.format(len(y) - sum(y), sum(y)))
```
Ways to create Pool class. If you have a big dataset it is effective (in terms of time) to load data from file, instead of pandas Dataframe.
```
dataset_dir = './amazon'
if not os.path.exists(dataset_dir):
os.makedirs(dataset_dir)
train_df.to_csv(
os.path.join(dataset_dir, 'train.csv'),
index=False, sep=',', header=True
)
test_df.to_csv(
os.path.join(dataset_dir, 'test.csv'),
index=False, sep=',', header=True
)
!head -3 amazon/train.csv
from catboost.utils import create_cd
feature_names = dict(list(enumerate(train_df.keys()[1:])))
create_cd(
label=0,
cat_features=list(range(1, train_df.shape[1])),
feature_names=feature_names,
output_path=os.path.join(dataset_dir, 'train.cd')
)
!cat amazon/train.cd
from catboost import Pool
pool1 = Pool(data=X, label=y, cat_features=cat_features)
pool2 = Pool(
data=os.path.join(dataset_dir, 'train.csv'),
delimiter=',',
column_description=os.path.join(dataset_dir, 'train.cd'),
has_header=True
)
print('Dataset shape: {}\n'.format(pool1.shape))
```
## Dataset Quantization
Features quantization. It is effective to quantize features single time before several trainings.
```
pool1.quantize(
border_count=254,
feature_border_type='GreedyLogSum',
#input_borders='.../path/to/borders',
per_float_feature_quantization=['0:border_count=1024']
)
```
## Split your data into train and validation
```
from sklearn.model_selection import train_test_split
data = train_test_split(X, y, train_size=0.8, random_state=0)
X_train, X_validation, y_train, y_validation = data
train_pool = Pool(
data=X_train,
label=y_train,
cat_features=cat_features
)
validation_pool = Pool(
data=X_validation,
label=y_validation,
cat_features=cat_features
)
```
## Training
```
from catboost import CatBoostClassifier
model = CatBoostClassifier(
iterations=5,
learning_rate=0.1,
)
model.fit(train_pool, eval_set=validation_pool, verbose=False)
print('Model is fitted: {}'.format(model.is_fitted()))
print('Model params:\n{}'.format(model.get_params()))
```
## Stdout of the training
```
model = CatBoostClassifier(
iterations=15,
# verbose=5,
)
model.fit(train_pool, eval_set=validation_pool);
```
## Metrics calculation and graph plotting
```
model = CatBoostClassifier(
iterations=200,
learning_rate=0.1,
custom_loss=['AUC', 'Accuracy']
)
model.fit(
train_pool,
eval_set=validation_pool,
verbose=False,
plot=True
);
```
## Model comparison
```
model1 = CatBoostClassifier(
learning_rate=0.7,
iterations=100,
train_dir='learing_rate_0.7'
)
model2 = CatBoostClassifier(
learning_rate=0.01,
iterations=100,
train_dir='learing_rate_0.01'
)
model1.fit(train_pool, eval_set=validation_pool, verbose=False)
model2.fit(train_pool, eval_set=validation_pool, verbose=False);
from catboost import MetricVisualizer
MetricVisualizer(['learing_rate_0.7', 'learing_rate_0.01']).start()
```
## Best iteration
```
model = CatBoostClassifier(
iterations=100,
learning_rate=0.5,
# use_best_model=False
)
model.fit(
train_pool,
eval_set=validation_pool,
verbose=False,
plot=True
);
print('Tree count: ' + str(model.tree_count_))
```
## Overfitting Detector
```
model_with_early_stop = CatBoostClassifier(
iterations=200,
learning_rate=0.5,
early_stopping_rounds=20
)
model_with_early_stop.fit(
train_pool,
eval_set=validation_pool,
verbose=False,
plot=True
);
print(model_with_early_stop.tree_count_)
```
### Overfitting Detector with eval metric
```
model_with_early_stop = CatBoostClassifier(
eval_metric='AUC',
iterations=200,
learning_rate=0.5,
early_stopping_rounds=20
)
model_with_early_stop.fit(
train_pool,
eval_set=validation_pool,
verbose=False,
plot=True
);
print(model_with_early_stop.tree_count_)
```
## Cross-validation
```
from catboost import cv
params = {
'loss_function': 'Logloss',
'iterations': 80,
'custom_loss': 'AUC',
'learning_rate': 0.5,
}
cv_data = cv(
params = params,
pool = train_pool,
fold_count=5,
shuffle=True,
partition_random_seed=0,
plot=True,
stratified=False,
verbose=False
)
cv_data.head(10)
best_value = cv_data['test-Logloss-mean'].min()
best_iter = cv_data['test-Logloss-mean'].values.argmin()
print('Best validation Logloss score: {:.4f}±{:.4f} on step {}'.format(
best_value,
cv_data['test-Logloss-std'][best_iter],
best_iter)
)
cv_data = cv(
params = params,
pool = train_pool,
fold_count=10,
shuffle=True,
partition_random_seed=0,
plot=True,
stratified=True,
verbose=False
)
```
## Grid Search
```
model = CatBoostClassifier(iterations=10, eval_metric='AUC')
grid = {'learning_rate': [0.001, 0.01, 0.1, 1.0, 10.0], 'depth': [4, 5, 6]}
result = model.grid_search(grid, train_pool)
print('Best parameters: {}\n'.format(result['params']))
msg = 'Mean AUC value on validation set per each iteration:\n{}'
print(msg.format(np.round(result['cv_results']['test-AUC-mean'], 4)))
model.get_params()
model.predict(validation_pool)
model.grid_search(grid, train_pool, plot=True, verbose=False);
```
More about parameter tuning you can find in [tutorial](https://github.com/catboost/catboost/blob/master/catboost/tutorials/hyperparameters_tuning/hyperparameters_tuning.ipynb).
## Model predictions
```
model = CatBoostClassifier(iterations=200, learning_rate=0.03)
model.fit(
train_pool,
verbose=False,
plot=True
);
```
If you have a big dataset, it is effective to make predictions for batch of objects, not single one.
```
print(model.predict_proba(X_validation))
raw_pred = model.predict(
X_validation,
prediction_type='RawFormulaVal'
)
print(raw_pred)
def sigmoid(x):
return 1 / (1 + np.exp(-x))
probabilities = sigmoid(raw_pred)
print(probabilities)
print(model.predict(X_validation))
```
## Select decision boundary

```
import matplotlib.pyplot as plt
from catboost.utils import get_roc_curve
from catboost.utils import get_fpr_curve
from catboost.utils import get_fnr_curve
curve = get_roc_curve(model, validation_pool)
(fpr, tpr, thresholds) = curve
(thresholds, fpr) = get_fpr_curve(curve=curve)
(thresholds, fnr) = get_fnr_curve(curve=curve)
plt.figure(figsize=(16, 8))
style = {'alpha':0.5, 'lw':2}
plt.plot(thresholds, fpr, color='blue', label='FPR', **style)
plt.plot(thresholds, fnr, color='green', label='FNR', **style)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xticks(fontsize=16)
plt.yticks(fontsize=16)
plt.grid(True)
plt.xlabel('Threshold', fontsize=16)
plt.ylabel('Error Rate', fontsize=16)
plt.title('FPR-FNR curves', fontsize=20)
plt.legend(loc="lower left", fontsize=16);
from catboost.utils import select_threshold
print(select_threshold(model, validation_pool, FNR=0.01))
print(select_threshold(model, validation_pool, FPR=0.01))
```
## Metric evaluation on a new dataset
```
metrics = model.eval_metrics(
data=validation_pool,
metrics=['Logloss','AUC'],
plot=True
)
print('AUC values:\n{}'.format(np.array(metrics['AUC'])))
```
## Feature importances
### Prediction values change
```
model.get_feature_importance(prettified=True)
```
### Loss function change
```
model.get_feature_importance(
validation_pool,
'LossFunctionChange',
prettified=True
)
```
### Shap values
```
shap_values = model.get_feature_importance(
train_pool,
'ShapValues'
)
expected_value = shap_values[0,-1]
shap_values = shap_values[:,:-1]
print(shap_values.shape)
import shap
shap.initjs()
shap.force_plot(
expected_value,
shap_values[2,:],
feature_names=train_pool.get_feature_names()
)
shap.force_plot(
expected_value,
shap_values[0,:],
feature_names=train_pool.get_feature_names()
)
shap.summary_plot(shap_values, X_train)
```
More information about shap value usage you can find in [tutorial](https://github.com/catboost/catboost/blob/master/catboost/tutorials/model_analysis/shap_values_tutorial.ipynb).
## Tree Visualization
```
model = CatBoostClassifier(iterations=2, depth=1)
features = [
[1, 2, 3],
[4, 5, 6],
[7, 8, 9]
]
labels = [1, 0, 1]
model.fit(features, labels, verbose=False);
# This cell doesn't work without graphviz package
# You can install it by link https://graphviz.gitlab.io/download/
# Installation can take a lot of time. You can do it at home.
from IPython.display import display
display(model.plot_tree(0))
display(model.plot_tree(1))
x = [0, 3, 2]
raw_pred = model.predict(x, prediction_type='RawFormulaVal')
print(raw_pred)
```
## Snapshotting
```
# !rm 'catboost_info/snapshot.bkp'
model = CatBoostClassifier(
iterations=200,
save_snapshot=True,
snapshot_file='snapshot.bkp',
snapshot_interval=1
)
model.fit(train_pool, eval_set=validation_pool, verbose=10);
```
## Saving the model
```
model = CatBoostClassifier(iterations=10)
model.fit(train_pool, eval_set=validation_pool, verbose=False)
model.save_model('catboost_model.bin')
model.save_model('catboost_model.json', format='json')
model.load_model('catboost_model.bin')
print(model.get_params())
print(model.learning_rate_)
```
## Hyperparameter tunning
```
tunned_model = CatBoostClassifier(
iterations=1000,
learning_rate=0.03,
# Quality
depth=6,
l2_leaf_reg=3,
random_strength=1,
one_hot_max_size=2,
# Final Model Size
max_ctr_complexity=1,
model_size_reg=50,
ctr_leaf_count_limit=100,
)
tunned_model.fit(
X_train, y_train,
cat_features=cat_features,
verbose=False,
eval_set=(X_validation, y_validation),
plot=True
);
```
| github_jupyter |
```
import cv2
import matplotlib.pyplot as plt
import numpy as np
import random
import os
from ssd_model import SSD300, SSD512
from ssd_utils import PriorUtil
from ssd_data import InputGenerator
from ssd_data import preprocess
from ssd_training import SSDLoss
from utils.model import load_weights
from data_voc import GTUtility
gt_util = GTUtility('data/VOC2007/')
gt_util_train, gt_util_val = gt_util.split(gt_util, split=0.8)
experiment = 'ssd_voc'
num_classes = gt_util.num_classes
#model = SSD300(num_classes=num_classes)
model = SSD512(num_classes=num_classes)
image_size = model.image_size
_, inputs, images, data = gt_util_train.sample_random_batch(batch_size=16, input_size=image_size)
test_idx = 5
test_img = images[test_idx]
test_gt = data[test_idx]
plt.imshow(test_img)
gt_util.plot_gt(test_gt, show_labels=True)
plt.show()
prior_util = PriorUtil(model)
egt = prior_util.encode(test_gt)
idxs = np.where(np.logical_not(egt[:,4]))[0]
egt[idxs][:,:5]
#x = prior_util.encode(test_gt)
#y = prior_util.decode(x)
self = prior_util
from utils.vis import to_rec
def plot_assignment(self, map_idx):
ax = plt.gca()
im = plt.gci()
image_height, image_width = image_size = im.get_size()
# ground truth
boxes = self.gt_boxes
boxes_x = (boxes[:,0] + boxes[:,2]) / 2. * image_height
boxes_y = (boxes[:,1] + boxes[:,3]) / 2. * image_width
for box in boxes:
xy_rec = to_rec(box[:4], image_size)
ax.add_patch(plt.Polygon(xy_rec, fill=False, edgecolor='b', linewidth=2))
plt.plot(boxes_x, boxes_y, 'bo', markersize=8)
# prior boxes
for idx, box_idx in self.match_indices.items():
if idx >= self.map_offsets[map_idx] and idx < self.map_offsets[map_idx+1]:
x, y = self.priors_xy[idx]
w, h = self.priors_wh[idx]
plt.plot(x, y, 'ro', markersize=4)
plt.plot([x, boxes_x[box_idx]], [y, boxes_y[box_idx]], '-r', linewidth=1)
ax.add_patch(plt.Rectangle((x-w/2, y-h/2), w+1, h+1,
fill=False, edgecolor='y', linewidth=2))
from ssd_utils import iou
def encode(self, gt_data, overlap_threshold=0.5, debug=False):
# calculation is done with normalized sizes
gt_boxes = self.gt_boxes = np.copy(gt_data[:,:4]) # normalized xmin, ymin, xmax, ymax
gt_labels = self.gt_labels = np.copy(gt_data[:,4:]) # one_hot classes including background
num_priors = self.priors.shape[0]
num_classes = gt_labels.shape[1]
# TODO: empty ground truth
if gt_data.shape[0] == 0:
print('gt_data', type(gt_data), gt_data.shape)
gt_iou = np.array([iou(b, self.priors_norm) for b in gt_boxes]).T
# assigne gt to priors
max_idxs = np.argmax(gt_iou, axis=1)
max_val = gt_iou[np.arange(num_priors), max_idxs]
prior_mask = max_val > overlap_threshold
match_indices = max_idxs[prior_mask]
self.match_indices = dict(zip(list(np.argwhere(prior_mask)[:,0]), list(match_indices)))
# prior labels
confidence = np.zeros((num_priors, num_classes))
confidence[:,0] = 1
confidence[prior_mask] = gt_labels[match_indices]
# compute local offsets from ground truth boxes
gt_xy = (gt_boxes[:,2:4] + gt_boxes[:,0:2]) / 2.
gt_wh = gt_boxes[:,2:4] - gt_boxes[:,0:2]
gt_xy = gt_xy[match_indices]
gt_wh = gt_wh[match_indices]
priors_xy = self.priors_xy[prior_mask] / self.image_size
priors_wh = self.priors_wh[prior_mask] / self.image_size
offsets = np.zeros((num_priors, 4))
offsets[prior_mask, 0:2] = (gt_xy - priors_xy) / priors_wh
offsets[prior_mask, 2:4] = np.log(gt_wh / priors_wh)
offsets[prior_mask, :] /= self.priors[prior_mask,-4:] # variances
return np.concatenate([offsets, confidence], axis=1)
def decode(self, model_output, confidence_threshold=0.01, keep_top_k=200):
# calculation is done with normalized sizes
offsets = model_output[:,:4]
confidence = model_output[:,4:]
num_priors = offsets.shape[0]
num_classes = confidence.shape[1]
priors_xy = self.priors_xy / self.image_size
priors_wh = self.priors_wh / self.image_size
# compute bounding boxes from local offsets
boxes = np.empty((num_priors, 4))
offsets *= self.priors[:,-4:] # variances
boxes_xy = priors_xy + offsets[:,0:2] * priors_wh
boxes_wh = priors_wh * np.exp(offsets[:,2:4])
boxes[:,0:2] = boxes_xy - boxes_wh / 2. # xmin, ymin
boxes[:,2:4] = boxes_xy + boxes_wh / 2. # xmax, ymax
boxes = np.clip(boxes, 0.0, 1.0)
prior_mask = confidence > confidence_threshold
# TODO: number of confident boxes, compute bounding boxes only for those?
#print(np.sum(np.any(prior_mask[:,1:], axis=1)))
# do non maximum suppression
results = []
for c in range(1, num_classes):
mask = prior_mask[:,c]
boxes_to_process = boxes[mask]
if len(boxes_to_process) > 0:
confs_to_process = confidence[mask, c]
feed_dict = {
self.boxes: boxes_to_process,
self.scores: confs_to_process
}
idx = self.sess.run(self.nms, feed_dict=feed_dict)
good_boxes = boxes_to_process[idx]
good_confs = confs_to_process[idx][:, None]
labels = np.ones((len(idx),1)) * c
c_pred = np.concatenate((good_boxes, good_confs, labels), axis=1)
results.extend(c_pred)
results = np.array(results)
if len(results) > 0:
order = np.argsort(results[:, 1])[::-1]
results = results[order]
results = results[:keep_top_k]
self.results = results
return results
x = encode(self, test_gt)
y = decode(self, x)
#for idx in range(len(prior_util.prior_maps)):
for idx in [1,2,3,4]:
m = prior_util.prior_maps[idx]
plt.figure(figsize=[10]*2)
plt.imshow(test_img)
m.plot_locations()
#m.plot_boxes([0, 10, 100])
#gt_util.plot_gt(test_gt)
plot_assignment(self, idx)
prior_util.plot_results(y, show_labels=False)
plt.show()
enc = prior_util.encode(test_gt)
prior_util.decode(enc)[:,:4]
test_gt[:,:4]
# plot ground truth
for i in range(4):
plt.figure(figsize=[8]*2)
plt.imshow(images[i])
gt_util.plot_gt(data[i])
plt.show()
# plot prior boxes
for i, m in enumerate(prior_util.prior_maps):
plt.figure(figsize=[8]*2)
#plt.imshow(images[7])
plt.imshow(images[8])
m.plot_locations()
#m.plot_boxes([0, 10, 100])
m.plot_boxes([0])
plt.axis('off')
#plt.savefig('plots/ssd_priorboxes_%i.pgf' % (i), bbox_inches='tight')
#print(m.map_size)
plt.show()
for l in model.layers:
try:
ks = l.weights[0].shape
except:
ks = ""
print("%30s %16s %24s %20s" % (l.name, l.__class__.__name__, l.output_shape, ks))
model.summary()
batch_size = 24
gen = InputGenerator(gt_util_train, prior_util, batch_size, model.image_size,
augmentation=False,
vflip_prob=0.0, do_crop=False)
g = gen.generate()
%%time
for j in range(3000):
i, o = next(g)
print(j, len(i))
if len(i) != batch_size:
print('FOOOOOOOOOOO')
import cProfile
g = gen.generate()
p = cProfile.Profile()
p.enable()
i, o = next(g)
p.disable()
p.print_stats(sort='cumulative')
```
| github_jupyter |
```
import numpy as np
import pandas as pd
from keras.layers import Dense, LSTM
import matplotlib.pyplot as plt
import tensorflow as tf
data_train = pd.read_csv('austin_weather_new.csv')
data_train.head()
len(data_train)
data_train.shape[0]
len(data_train.index)
data_train.dtypes
data_train['TempHighF'] = data_train.TempHighF.astype(float)
data_train['TempAvgF'] = data_train.TempAvgF.astype(float)
data_train['TempLowF'] = data_train.TempLowF.astype(float)
data_train['DewPointHighF'] = data_train.DewPointHighF.astype(float)
data_train['DewPointAvgF'] = data_train.DewPointAvgF.astype(float)
data_train['DewPointLowF'] = data_train.DewPointLowF.astype(float)
data_train['HumidityHighPercent'] = data_train.HumidityHighPercent.astype(float)
data_train['HumidityAvgPercent'] = data_train.HumidityAvgPercent.astype(float)
data_train['HumidityLowPercent'] = data_train.HumidityLowPercent.astype(float)
data_train.dtypes
data_train.isnull().sum()
dates = data_train['Date'].values
temp = data_train['TempAvgF'].values
dew = data_train['DewPointAvgF'].values
hum = data_train['HumidityAvgPercent'].values
plt.figure(figsize=(8,5))
plt.plot(dates, temp)
plt.title('Rataan Suhu',
fontsize=20);
plt.figure(figsize=(8,5))
plt.plot(dates, dew)
plt.title('Rataan Dew',
fontsize=20);
plt.figure(figsize=(8,5))
plt.plot(dates, hum)
plt.title('Rataan Lembab',
fontsize=20);
def windowed_dataset(series, window_size, batch_size, shuffle_buffer):
series = tf.expand_dims(series, axis=-1)
ds = tf.data.Dataset.from_tensor_slices(series)
ds = ds.window(window_size + 1, shift=1, drop_remainder=True)
ds = ds.flat_map(lambda w: w.batch(window_size + 1))
ds = ds.shuffle(shuffle_buffer)
ds = ds.map(lambda w: (w[:-1], w[1:]))
return ds.batch(batch_size).prefetch(1)
```
**Model sequential dan Learning Rate pada Optimizer**
```
train_set = windowed_dataset(temp, window_size=60, batch_size=100, shuffle_buffer=1000)
model = tf.keras.models.Sequential([
tf.keras.layers.LSTM(60, return_sequences=True),
tf.keras.layers.LSTM(60),
tf.keras.layers.Dense(30, activation="relu"),
tf.keras.layers.Dense(10, activation="relu"),
tf.keras.layers.Dense(1),
])
```
**Training MAE**
```
optimizer = tf.keras.optimizers.SGD(lr=1.0000e-04, momentum=0.9)
model.compile(loss=tf.keras.losses.Huber(),
optimizer = optimizer,
metrics=["mae"])
history = model.fit(train_set,epochs=100)
```
| github_jupyter |
# 23) Intro to machine learning (ML)
*This notebook contains excerpts from the [Python Data Science Handbook](http://shop.oreilly.com/product/0636920034919.do) by Jake VanderPlas; the original content is available [on GitHub](https://github.com/jakevdp/PythonDataScienceHandbook).*
## What Is Machine Learning?
- Not magic; requires knowledge of which methods are appropriate when
- One definition: *building models of data*
- The "learning" is from tuning parameters to fit training set data
- The results are models that predict and understand aspects of newly observed data
### Categories of Machine Learning
- *Supervised learning* involves somehow modeling the relationship between measured features of data and some label associated with the data; once this model is determined, it can be used to apply labels to new, unknown data.
- This is further subdivided into *classification* tasks and *regression* tasks: in classification, the labels are discrete categories, while in regression, the labels are continuous quantities.
- *Unsupervised learning* involves modeling the features of a dataset without reference to any label, and is often described as "letting the dataset speak for itself." These models include tasks such as *clustering* and *dimensionality reduction.*
- Clustering algorithms identify distinct groups of data, while dimensionality reduction algorithms search for more succinct representations of the data.
- *Semi-supervised learning* methods, which falls somewhere between supervised learning and unsupervised learning. Semi-supervised learning methods are often useful when only incomplete labels are available.

from [Ferguson, AL, "Machine learning and data science in soft materials engineering," **2018** *J. Phys.: Condens. Matter* **30** 043002](https://doi.org/10.1088/1361-648X/aa98bd)
### Qualitative Examples of Machine Learning Applications
To make these ideas more concrete, let's take a look at a few very simple examples of a machine learning task.
These examples are meant to give an intuitive, non-quantitative overview of the types of machine learning tasks we will be looking at in this chapter.
FYI: you can find the Python source that generates the following figures in the [Appendix: Figure Code](https://github.com/jakevdp/PythonDataScienceHandbook/blob/master/notebooks/06.00-Figure-Code.ipynb).
#### Classification: Predicting discrete labels
We will first take a look at a simple *classification* task, in which you are given a set of labeled points and want to use these to classify some unlabeled points.
Imagine that we have the data shown in this figure:

Here we have two-dimensional data: that is, we have two *features* for each point, represented by the *(x,y)* positions of the points on the plane.
In addition, we have one of two *class labels* for each point, here represented by the colors of the points.
From these features and labels, we would like to create a model that will let us decide whether a new point should be labeled "blue" or "red."
There are a number of possible models for such a classification task, but here we will use an extremely simple one. We will make the assumption that the two groups can be separated by drawing a straight line through the plane between them, such that points on each side of the line fall in the same group.
**Here the *model* is a quantitative version of the statement "a straight line separates the classes", while the *model parameters* are the particular numbers describing the location and orientation of that line for our data.**
The optimal values for these model parameters are learned from the data (this is the "learning" in machine learning), which is often called *training the model*.
The following figure shows a visual representation of what the trained model looks like for this data:

Now that this model has been trained, it can be generalized to new, unlabeled data.
In other words, we can take a new set of data, draw this model line through it, and assign labels to the new points based on this model.
This stage is usually called *prediction*. See the following figure:

This is the basic idea of a classification task in machine learning, where **"classification" indicates that the data has discrete class labels.**
At first glance this may look fairly trivial: it would be relatively easy to simply look at this data and draw such a discriminatory line to accomplish this classification.
A benefit of the machine learning approach, however, is that it can generalize to much larger datasets in many more dimensions.
For example, this is similar to the task of automated spam detection for email; in this case, we might use the following features and labels:
- *feature 1*, *feature 2*, etc. $\to$ normalized counts of important words or phrases ("Viagra", "Nigerian prince", etc.)
- *label* $\to$ "spam" or "not spam"
For the training set, these labels might be determined by individual inspection of a small representative sample of emails; for the remaining emails, the label would be determined using the model.
For a suitably trained classification algorithm with enough well-constructed features (typically thousands or millions of words or phrases), this type of approach can be very effective.
### Regression: Predicting continuous labels
In contrast with the discrete labels of a classification algorithm, we will next look at a simple *regression* task in which the labels are continuous quantities.
Consider the data shown in the following figure, which consists of a set of points each with a continuous label:

As with the classification example, we have two-dimensional data: that is, there are two features describing each data point.
The color of each point represents the continuous label for that point.
There are a number of possible regression models we might use for this type of data, but here we will use a simple linear regression to predict the points.
This simple linear regression model assumes that if we treat the label as a third spatial dimension, we can fit a plane to the data.
This is a higher-level generalization of the well-known problem of fitting a line to data with two coordinates.
We can visualize this setup as shown in the following figure:

Notice that the *feature 1-feature 2* plane here is the same as in the two-dimensional plot from before; in this case, however, we have represented the labels by both color and three-dimensional axis position.
From this view, it seems reasonable that fitting a plane through this three-dimensional data would allow us to predict the expected label for any set of input parameters.
Returning to the two-dimensional projection, when we fit such a plane we get the result shown in the following figure:

This plane of fit gives us what we need to predict labels for new points.
Visually, we find the results shown in the following figure:

As with the classification example, this may seem rather trivial in a low number of dimensions.
But the power of these methods is that they can be straightforwardly applied and evaluated in the case of data with many, many features.
For example, this is similar to the task of computing the distance to galaxies observed through a telescope—in this case, we might use the following features and labels:
- *feature 1*, *feature 2*, etc. $\to$ brightness of each galaxy at one of several wave lengths or colors
- *label* $\to$ distance or redshift of the galaxy
The distances for a small number of these galaxies might be determined through an independent set of (typically more expensive) observations.
Distances to remaining galaxies could then be estimated using a suitable regression model, without the need to employ the more expensive observation across the entire set.
In astronomy circles, this is known as the "photometric redshift" problem.
### Clustering: Inferring labels on unlabeled data
The classification and regression illustrations we just looked at are examples of supervised learning algorithms, in which we are trying to build a model that will predict labels for new data.
Unsupervised learning involves models that describe data without reference to any known labels.
One common case of unsupervised learning is "clustering," in which data is automatically assigned to some number of discrete groups.
For example, we might have some two-dimensional data like that shown in the following figure:

By eye, it is clear that each of these points is part of a distinct group.
Given this input, a clustering model will use the intrinsic structure of the data to determine which points are related.
Using the very fast and intuitive *k*-means algorithm (which we'll return to), we find the clusters shown in the following figure:

*k*-means fits a model consisting of *k* cluster centers; the optimal centers are assumed to be those that minimize the distance of each point from its assigned center.
Again, this might seem like a trivial exercise in two dimensions, but as our data becomes larger and more complex, such clustering algorithms can be employed to extract useful information from the dataset.
### Dimensionality reduction: Inferring structure of unlabeled data
Dimensionality reduction is another example of an unsupervised algorithm, in which labels or other information are inferred from the structure of the dataset itself.
Dimensionality reduction is a bit more abstract than the examples we looked at before, but generally it seeks to pull out some low-dimensional representation of data that in some way preserves relevant qualities of the full dataset.
As an example of this, consider the data shown in the following figure:

Visually, it is clear that there is some structure in this data: it is drawn from a one-dimensional line that is arranged in a spiral within this two-dimensional space.
In a sense, you could say that this data is "intrinsically" only one dimensional, though this one-dimensional data is embedded in higher-dimensional space.
A suitable dimensionality reduction model in this case would be sensitive to this nonlinear embedded structure, and be able to pull out this lower-dimensionality representation.
The following figure shows a visualization of the results of the Isomap algorithm, a manifold learning algorithm that does exactly this:

Notice that the colors (which represent the extracted one-dimensional latent variable) change uniformly along the spiral, which indicates that the algorithm did in fact detect the structure we saw by eye.
As with the previous examples, the power of dimensionality reduction algorithms becomes clearer in higher-dimensional cases.
For example, we might wish to visualize important relationships within a dataset that has 100 or 1,000 features.
Visualizing 1,000-dimensional data is a challenge, and one way we can make this more manageable is to use a dimensionality reduction technique to reduce the data to two or three dimensions.
## Summary
Here we have seen a few simple examples of some of the basic types of machine learning approaches.
Needless to say, there are a number of important practical details that we have glossed over, but I hope this section was enough to give you a basic idea of what types of problems machine learning approaches can solve.
In short, we saw the following:
- *Supervised learning*: Models that can predict labels based on labeled training data
- *Classification*: Models that predict labels as two or more discrete categories
- *Regression*: Models that predict continuous labels
- *Unsupervised learning*: Models that identify structure in unlabeled data
- *Clustering*: Models that detect and identify distinct groups in the data
- *Dimensionality reduction*: Models that detect and identify lower-dimensional structure in higher-dimensional data
In the following sections we will go into much greater depth within these categories, and see some more interesting examples of where these concepts can be useful.
All of the figures in the preceding discussion are generated based on actual machine learning computations; the code behind them can be found in [Appendix: Figure Code](06.00-Figure-Code.ipynb).
| github_jupyter |
# VQE on Aer simulator with noise
This notebook demonstrates using the [Qiskit Aer](https://qiskit.org/documentation/the_elements.html#aer) `aer_simulator` to run a simulation with noise, based on a given noise model. This can be useful to investigate behavior under different noise conditions. Aer not only allows you to define your own custom noise model, but also allows a noise model to be easily created based on the properties of a real quantum device. The latter is what this notebook will demonstrate since the goal is to show VQE with noise and not the more complex task of how to build custom noise models.
The `QuantumInstance` module provides a solution to mitigate the measurement error when running on a noisy simulation or a real quantum device. This allows any algorithm using it to automatically have measurement noise mitigation applied.
Further information on Qiskit Aer noise model can be found in the online [Qiskit Aer documentation](https://qiskit.org/documentation/apidoc/aer_noise.html), also there is tutorial for [building noise models](../simulators/3_building_noise_models.ipynb).
```
import numpy as np
import pylab
from qiskit import Aer
from qiskit.utils import QuantumInstance, algorithm_globals
from qiskit.algorithms import VQE, NumPyMinimumEigensolver
from qiskit.algorithms.optimizers import SPSA
from qiskit.circuit.library import TwoLocal
from qiskit.opflow import I, X, Z
```
Noisy simulation will be demonstrated here with VQE, finding the minimum (ground state) energy of an Hamiltonian, but the technique applies to any quantum algorithm from Qiskit.
So for VQE we need a qubit operator as input. Here, once again, we will take a set of paulis that were originally computed by Qiskit Nature, for an H2 molecule, so we can quickly create an Operator.
```
H2_op = (-1.052373245772859 * I ^ I) + \
(0.39793742484318045 * I ^ Z) + \
(-0.39793742484318045 * Z ^ I) + \
(-0.01128010425623538 * Z ^ Z) + \
(0.18093119978423156 * X ^ X)
print(f'Number of qubits: {H2_op.num_qubits}')
```
As the above problem is still easily tractable classically we can use NumPyMinimumEigensolver to compute a reference value so we can compare later the results.
```
npme = NumPyMinimumEigensolver()
result = npme.compute_minimum_eigenvalue(operator=H2_op)
ref_value = result.eigenvalue.real
print(f'Reference value: {ref_value:.5f}')
```
## Performance *without* noise
First we will run on the simulator without adding noise to see the result. I have created the backend and QuantumInstance, which holds the backend as well as various other run time configuration, which are defaulted here, so it easy to compare when we get to the next section where noise is added. There is no attempt to mitigate noise or anything in this notebook so the latter setup and running of VQE is identical.
```
seed = 170
iterations = 125
algorithm_globals.random_seed = seed
backend = Aer.get_backend('aer_simulator')
qi = QuantumInstance(backend=backend, seed_simulator=seed, seed_transpiler=seed)
counts = []
values = []
def store_intermediate_result(eval_count, parameters, mean, std):
counts.append(eval_count)
values.append(mean)
ansatz = TwoLocal(rotation_blocks='ry', entanglement_blocks='cz')
spsa = SPSA(maxiter=iterations)
vqe = VQE(ansatz, optimizer=spsa, callback=store_intermediate_result, quantum_instance=qi)
result = vqe.compute_minimum_eigenvalue(operator=H2_op)
print(f'VQE on Aer qasm simulator (no noise): {result.eigenvalue.real:.5f}')
print(f'Delta from reference energy value is {(result.eigenvalue.real - ref_value):.5f}')
```
We captured the energy values above during the convergence so we can see what went on in the graph below.
```
pylab.rcParams['figure.figsize'] = (12, 4)
pylab.plot(counts, values)
pylab.xlabel('Eval count')
pylab.ylabel('Energy')
pylab.title('Convergence with no noise')
```
## Performance *with* noise
Now we will add noise. Here we will create a noise model for Aer from an actual device. You can create custom noise models with Aer but that goes beyond the scope of this notebook. Links to further information on Aer noise model, for those that may be interested in doing this, were given above.
First we need to get an actual device backend and from its `configuration` and `properties` we can setup a coupling map and a noise model to match the device. While we could leave the simulator with the default all to all map, this shows how to set the coupling map too. Note: We can also use this coupling map as the entanglement map for the variational form if we choose.
Note: simulation with noise will take longer than without noise.
Terra Mock Backends:
We will use real noise data for an IBM Quantum device using the date stored in Qiskit Terra. Specifically, in this tutorial, the device is ibmq_vigo.
```
import os
from qiskit.providers.aer import QasmSimulator
from qiskit.providers.aer.noise import NoiseModel
from qiskit.test.mock import FakeVigo
device_backend = FakeVigo()
backend = Aer.get_backend('aer_simulator')
counts1 = []
values1 = []
noise_model = None
device = QasmSimulator.from_backend(device_backend)
coupling_map = device.configuration().coupling_map
noise_model = NoiseModel.from_backend(device)
basis_gates = noise_model.basis_gates
print(noise_model)
print()
algorithm_globals.random_seed = seed
qi = QuantumInstance(backend=backend, seed_simulator=seed, seed_transpiler=seed,
coupling_map=coupling_map, noise_model=noise_model,)
def store_intermediate_result1(eval_count, parameters, mean, std):
counts1.append(eval_count)
values1.append(mean)
var_form = TwoLocal(rotation_blocks='ry', entanglement_blocks='cz')
spsa = SPSA(maxiter=iterations)
vqe = VQE(ansatz, optimizer=spsa, callback=store_intermediate_result1, quantum_instance=qi)
result1 = vqe.compute_minimum_eigenvalue(operator=H2_op)
print(f'VQE on Aer qasm simulator (with noise): {result1.eigenvalue.real:.5f}')
print(f'Delta from reference energy value is {(result1.eigenvalue.real - ref_value):.5f}')
if counts1 or values1:
pylab.rcParams['figure.figsize'] = (12, 4)
pylab.plot(counts1, values1)
pylab.xlabel('Eval count')
pylab.ylabel('Energy')
pylab.title('Convergence with noise')
```
## Performance *with* noise and measurement error mitigation
Now we will add method for measurement error mitigation, which increases the fidelity of measurement. Here we choose `CompleteMeasFitter` to mitigate the measurement error. The calibration matrix will be auto-refresh every 30 minute (default value).
Note: simulation with noise will take longer than without noise.
```
from qiskit.utils.mitigation import CompleteMeasFitter
counts2 = []
values2 = []
if noise_model is not None:
algorithm_globals.random_seed = seed
qi = QuantumInstance(backend=backend, seed_simulator=seed, seed_transpiler=seed,
coupling_map=coupling_map, noise_model=noise_model,
measurement_error_mitigation_cls=CompleteMeasFitter,
cals_matrix_refresh_period=30)
def store_intermediate_result2(eval_count, parameters, mean, std):
counts2.append(eval_count)
values2.append(mean)
ansatz = TwoLocal(rotation_blocks='ry', entanglement_blocks='cz')
spsa = SPSA(maxiter=iterations)
vqe = VQE(ansatz, optimizer=spsa, callback=store_intermediate_result2, quantum_instance=qi)
result2 = vqe.compute_minimum_eigenvalue(operator=H2_op)
print(f'VQE on Aer qasm simulator (with noise and measurement error mitigation): {result2.eigenvalue.real:.5f}')
print(f'Delta from reference energy value is {(result2.eigenvalue.real - ref_value):.5f}')
if counts2 or values2:
pylab.rcParams['figure.figsize'] = (12, 4)
pylab.plot(counts2, values2)
pylab.xlabel('Eval count')
pylab.ylabel('Energy')
pylab.title('Convergence with noise, measurement error mitigation enabled')
```
Lets bring the results together here for a summary.
We produced a reference value using a classical algorithm and then proceeded to run VQE on a qasm simulator. While the simulation is ideal (no noise) there is so called shot-noise due to sampling - increasing the number of shots reduces this as more and more samples are taken, but shots was left at the default of 1024 and we see a small effect in the outcome.
Then we added noise using a model taken off a real device and can see the result is affected. Finally we added measurement noise mitigation which adjusts the results in an attempt to alleviate the affect of noise in the classical equipment measuring the qubits.
```
print(f'Reference value: {ref_value:.5f}')
print(f'VQE on Aer qasm simulator (no noise): {result.eigenvalue.real:.5f}')
print(f'VQE on Aer qasm simulator (with noise): {result1.eigenvalue.real:.5f}')
print(f'VQE on Aer qasm simulator (with noise and measurement error mitigation): {result2.eigenvalue.real:.5f}')
import qiskit.tools.jupyter
%qiskit_version_table
%qiskit_copyright
```
| github_jupyter |
```
from pathlib import Path
from src.notebookcodesnippets.hideInputs import hide_toggle
hide_toggle()
from IPython.display import display, Markdown
with open(Path.cwd().parent.parent.parent / "src" / "notebookcodesnippets" / "supervised" / "intro.md", 'r') as fh:
content = fh.read()
display(Markdown(content))
hide_toggle()
title = "Insightful"
InsertApproach = "03-insightful-approach"
numberOfPrincipalComponents = 176
```
# Imports and reading of data
Most of the code used in this notebook is found in the src library.
```
# Optional: Load the "autoreload" extension so that code can change
%load_ext autoreload
#OPTIONAL: Always reload modules so that as you change code in src, it gets loaded
%autoreload 2
hide_toggle()
import sys
sys.path.insert(0, "../")
data_dir = Path.cwd().parent.parent.parent / "data"
print("Current data directory {}".format(data_dir))
models_dir = Path.cwd().parent.parent.parent / "models"
# src
from src.models import train_model, predict_model
from src.features import build_features
from src.visualization import visualize
#Standard libraries
import numpy as np
import pandas as pd
import pickle
#Models
from sklearn.ensemble import RandomForestClassifier, GradientBoostingClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
# CV
from sklearn.model_selection import RepeatedStratifiedKFold
#visualizations
import plotly.graph_objs as go
from tqdm import tqdm
# setting random seed for reproducibility
random_state=1
hide_toggle()
data = pd.read_pickle(data_dir / "processed" / "processedData.pkl")
trainingData = pd.read_pickle(data_dir / InsertApproach / "processed" / "trainingData.pkl")
trainingTarget= pd.read_pickle(data_dir / InsertApproach / "processed" / "trainingTarget.pkl")
testSet = pd.read_pickle(data_dir / InsertApproach / "processed" / "testSet.pkl")
trainingData
testSet
from IPython.display import display, Markdown
with open(Path.cwd().parent.parent.parent / "src" / "notebookcodesnippets" / "supervised" / "algorithms.md", 'r') as fh:
content = fh.read()
display(Markdown(content))
hide_toggle()
InsertAlgorithms = [LogisticRegression (random_state = random_state, max_iter=200),
DecisionTreeClassifier (random_state = random_state, max_features = "auto"),
RandomForestClassifier (random_state = random_state, max_features = "auto", max_depth=6),\
GradientBoostingClassifier(random_state = random_state, max_features = "auto")]
InsertAbbreviations = ["LOG", "DT", "RF", "GB"]
InsertprettyNames = ["Logistic regression", "Decision Tree", "Random Forest", "Gradient Boost"]
from IPython.display import display, Markdown
with open(Path.cwd().parent.parent.parent / "src" / "notebookcodesnippets" / "supervised" / "optimalparameters.md", 'r') as fh:
content = fh.read()
display(Markdown(content))
hide_toggle()
# Optional to include any sampling method. Default is none.
includeSampleMethods = [""]#, "under", "over", "both"]
numberRuns = 5
numberSplits = 5
rskfold = RepeatedStratifiedKFold(n_splits=numberSplits, n_repeats=numberRuns, random_state=random_state)
ModelsBestParams = pd.Series({}, dtype="string")
Abbreviations = []
prettyNames = []
Algorithms = []
for i, algorithm in tqdm(enumerate(InsertAlgorithms)):
for method in includeSampleMethods:
print("Finding best params for: {}".format(InsertAbbreviations[i] + " " + method))
bestEstimator, ModelsBestParams[InsertAbbreviations[i] + " " + method] = train_model.applyGridSearch(
X = trainingData.drop(["material_id", "full_formula"], axis=1),
y = trainingTarget.values.reshape(-1,),
model = algorithm,
cv = rskfold,
numPC = numberOfPrincipalComponents,
sampleMethod = method,
searchPC = True )
Abbreviations.append(InsertAbbreviations[i] + " " + method)
prettyNames.append(InsertAbbreviations[i] + " " + method)
Algorithms.append(bestEstimator)
from IPython.display import display, Markdown
with open(Path.cwd().parent.parent.parent / "src" / "notebookcodesnippets" / "supervised" / "visualoptimalparameters.md", 'r') as fh:
content = fh.read()
display(Markdown(content))
hide_toggle()
visualize.principalComponentsVSscores(X = trainingData.drop(["material_id", "full_formula"], axis=1),
ModelsBestParams = ModelsBestParams,
prettyNames = prettyNames,
numPC = numberOfPrincipalComponents,
approach = InsertApproach)
visualize.gridsearchVSscores(X = trainingData.drop(["material_id", "full_formula"], axis=1),
ModelsBestParams = ModelsBestParams,
prettyNames = prettyNames,
approach = InsertApproach)
from scikitplot.metrics import plot_calibration_curve
#plot_calibration_curve()
ModelsBestParams["RF "].best_estimator_
from IPython.display import display, Markdown
with open(Path.cwd().parent.parent.parent / "src" / "notebookcodesnippets" / "supervised" / "roc-auc.md", 'r') as fh:
content = fh.read()
display(Markdown(content))
hide_toggle()
SupervisedModels = pd.Series({}, dtype="string")
for i, algorithm in enumerate(Algorithms):
print("Current training algorithm: {}".format(prettyNames[i]))
SupervisedModels[Abbreviations[i]] = (
visualize.evaluatePrecisionRecallMetrics(classifier = algorithm,
X = trainingData.drop(["material_id", "full_formula"], axis=1),
y = trainingTarget.values.reshape(-1,),
k = numberSplits,
n = numberRuns,
cv = rskfold,
featureImportance = True,
title = prettyNames[i],
numPC = numberOfPrincipalComponents,
approach = InsertApproach )
)
from IPython.display import display, Markdown
with open(Path.cwd().parent.parent.parent / "src" / "notebookcodesnippets" / "supervised" / "visualize-cv.md", 'r') as fh:
content = fh.read()
display(Markdown(content))
hide_toggle()
visualize.plot_accuracy(SupervisedModels, prettyNames,
prettyNames = prettyNames,
numPC = numberOfPrincipalComponents,
approach = InsertApproach)
from IPython.display import display, Markdown
with open(Path.cwd().parent.parent.parent / "src" / "notebookcodesnippets" / "supervised" / "visualizerelevant.md", 'r') as fh:
content = fh.read()
display(Markdown(content))
hide_toggle()
visualize.plot_important_features(SupervisedModels,
X = trainingData.drop(["material_id", "full_formula"], axis=1),
k = numberSplits,
n = numberRuns,
prettyNames = prettyNames,
numPC = numberOfPrincipalComponents,
approach = InsertApproach,
numFeat = 15)
from IPython.display import display, Markdown
with open(Path.cwd().parent.parent.parent / "src" / "notebookcodesnippets" / "supervised" / "falselypredicted.md", 'r') as fh:
content = fh.read()
display(Markdown(content))
hide_toggle()
visualize.plot_confusion_metrics(SupervisedModels, prettyNames, trainingData,
k = numberSplits,
n = numberRuns,
prettyNames = prettyNames,
numPC = numberOfPrincipalComponents,
approach = InsertApproach)
visualize.confusion_matrixQT(SupervisedModels, trainingTarget,
prettyNames = prettyNames,
numPC = numberOfPrincipalComponents,
approach = InsertApproach)
from IPython.display import display, Markdown
with open(Path.cwd().parent.parent.parent / "src" / "notebookcodesnippets" / "supervised" / "predictions.md", 'r') as fh:
content = fh.read()
display(Markdown(content))
hide_toggle()
Summary = pd.DataFrame({}, dtype="string")
Summary["material_id"] = testSet["material_id"]
Summary["full_formula"] = testSet["full_formula"]
Summary["pretty_formula"] = testSet["pretty_formula"]
PredictedCandidates = pd.Series({}, dtype="string")
#threshold = numberSplits*numberRuns/2 #50% when equal
trainSet = trainingData.drop(["material_id", "full_formula"], axis=1)
testData = testSet.drop(["pretty_formula", "candidate", "full_formula", "material_id"], axis=1)
fittedAlgorithms = []
for i, algorithm in tqdm(enumerate(Algorithms)):
# Train the final model with the best estimators
fittedAlgorithm = train_model.fitAlgorithm(algorithm,
trainingData = trainSet,#trainSet[trainSet.columns[SupervisedModels[Abbreviations[i]]["importantKeys"]>threshold]],\
trainingTarget = trainingTarget.values.reshape(-1,),)
fittedAlgorithms.append(fittedAlgorithm)
# Predict the final model on unseen data
PredictedCandidates[Abbreviations[i]],\
PredictedCandidates[Abbreviations[i]+"Prob"] = predict_model.runPredictions(fittedAlgorithm,\
testData = testData)#[testData.columns[SupervisedModels[Abbreviations[i]]["importantKeys"]>threshold]])
for abbreviation in Abbreviations:
Summary[abbreviation] = PredictedCandidates[abbreviation]
Summary[abbreviation + "Prob"] = PredictedCandidates[abbreviation + "Prob"]
print("{} predict the number of candidates as: {}".format(abbreviation, int(np.sum(PredictedCandidates[abbreviation]))))
Summary
from IPython.display import display, Markdown
with open(Path.cwd().parent.parent.parent / "src" / "notebookcodesnippets" / "supervised" / "save.md", 'r') as fh:
content = fh.read()
display(Markdown(content))
hide_toggle()
for i, fitted_algorithm in tqdm(enumerate(fittedAlgorithms)):
file_path = Path(models_dir / InsertApproach / "trained-models" / Path("PCA-" + str(numberOfPrincipalComponents) + "-" + prettyNames[i] + ".pkl"))
with file_path.open("wb") as fp:
pickle.dump(fitted_algorithm, fp)
file_path = Path(models_dir / InsertApproach / "summary" / Path("PCA-" + str(numberOfPrincipalComponents) + "-" + "summary.pkl"))
with file_path.open("wb") as fp:
pickle.dump(Summary, fp)
```
| github_jupyter |
# Notebook for ingesting ENEM data
This notebook aims to ingest ENEM microdata stored on the [INEP](http://portal.inep.gov.br/web/guest/microdados) website, where it basically consists of the following steps:
1. Prepare the environment (install unzips, create folders, define variables and import python libraries)
2. Create Python functions that will extract the data from the unzipped file, convert and upload it to the S3 bucket
3. Download the files from the INEP website according to the list defined in the **step 1** variable
4. Perform the transformation functions defined in **step 2** in the downloaded .zip files
5. Create the database in AWS Glue, then create and run the crawler that will register the tables in the data catalog
After completing this notebook, you can proceed with the data transformation, exploration and consumption processes in Athena.
*Note: it is a prerequisite for running this notebook to create the [CloudFormation stack](https://github.com/aws-samples/aws-edu-exam-analytics/blob/main/templates/EduLabCFN.yaml) as specified in the aws-samples repository.*
---
### 1.1- *Download and binary installation for rar uncompress (one of the downloaded files contains a .rar file)*
```
%%bash
wget https://www.rarlab.com/rar/rarlinux-x64-5.9.1.tar.gz
tar -xvzf rarlinux-x64-5.9.1.tar.gz
rm rarlinux-x64-5.9.1.tar.gz
```
### 1.2- *Download and install wget library in Python*
```
!pip install wget
```
### 1.3- *Import Python libraries that will be used by code in next cells*
```
import zipfile
import wget
import fnmatch
import os
import gzip
import boto3
import botocore
import sys
import shutil
from zipfile import ZipFile
from pprint import pprint
```
### 1.4- *Variables definition and folders criation to store zip packages downloaded and microdata files in csv format*
```
#define variáveis e cria diretorios de trabalho
zipdir = 'zips'
outdir = 'microdados'
list_arq=[
'microdados_enem2012'
,'microdados_enem2013'
,'microdados_enem2014'
,'microdados_enem2015'
,'microdados_enem2016'
,'microdados_enem2017'
,'microdados_enem2018'
,'microdados_enem_2019']
#troque o nome do bucket para o criado no stack do Cloudformation
bucket='edu-bucket-999'
folder='data'
print(list_arq)
os.mkdir(zipdir)
os.mkdir(outdir)
```
### 2- *Function blocks to provide wget download status, uncompress, transform, and upload data to S3 bucket*
```
#bloco de funções
#status do download wget
def bar_custom(current, total, width=80):
progress_message = "Downloading: %d%% [%d / %d] bytes" % (current / total * 100, current, total)
sys.stdout.write("\r" + progress_message)
#converte pra utf8 e comprime
def convert_compress_file(csvfile):
year=csvfile[-8:-4]
filenameout="microdados/MICRODADOS_ENEM_"+year+".csv.gz"
print("Convertendo "+csvfile+" para utf-8 e compactando para "+filenameout+"...")
#converte, compacta e remove aspas se for o caso
with open(csvfile,encoding='cp1252') as filein,gzip.open(filenameout,'wt',encoding='utf8') as fileout:
for line in filein:
fileout.write(line.replace('"', ''))
os.remove(csvfile)
return filenameout
#carrega dados no bucket
def upload_s3(upfile,bucket,folder):
year=upfile[-11:-7]
s3 = boto3.resource('s3')
data = open(upfile, "rb")
key = folder + '/enem_microdados_' + year + '/' + os.path.basename(upfile)
print("Carregando "+key+" para o bucket "+bucket)
s3.Bucket(bucket).put_object(Key=key, Body=data)
# transformação e upload do arquivo csv
def microdados_transform(microfile):
pattern1="*/DADOS_*.csv"
pattern2="*/[Mm][Ii][Cc][Rr][Oo]*.csv"
unrarcmd="/home/ec2-user/SageMaker/rar/unrar e "
with ZipFile(microfile, 'r') as zipObj:
listOfiles = zipObj.namelist()
#Se for arquivo rar
if fnmatch.filter(listOfiles, '*.rar'):
rarfile=fnmatch.filter(listOfiles, '*.rar')[0]
print("Arquivo rar "+rarfile)
zipObj.extractall()
unrarlb="/home/ec2-user/SageMaker/rar/unrar lb "+rarfile+" | grep MICRO | grep csv"
extractfile=os.popen(unrarlb).readline().rstrip("\r\n")
#extractfile=os.path.basename(result)
print("Extraindo arquivo "+extractfile)
#print(unrarcmd+rarfile+" "+extractfile)
os.system(unrarcmd+rarfile+" "+extractfile)
print("Movendo arquivo para pasta microdados")
finalfile='microdados/'+os.path.basename(extractfile)
os.rename(os.path.basename(extractfile),finalfile)
os.remove(rarfile)
else:
for extractfile in fnmatch.filter(listOfiles, '*.csv'):
if fnmatch.fnmatch(extractfile,pattern1) or fnmatch.fnmatch(extractfile,pattern2):
print("Arquivo zip "+microfile)
print("Extraindo arquivo "+extractfile)
zipObj.extract(extractfile)
print("Movendo arquivo para pasta microdados")
finalfile='microdados/'+os.path.basename(extractfile)
os.rename(extractfile,finalfile)
basepath=extractfile.split("/")[0]
print("Removendo "+ basepath)
shutil.rmtree(basepath)
return finalfile
```
### 3- *Download of microdata file packages from INEP site according to file list in variable previously defined*
```
#Download dos arquivos (com base em list_arq)
for item in list_arq:
year=item[-4:]
if os.path.isfile('zips/'+item+'.zip'):
print("arquivo "+item+".zip já existe")
else:
print("carregando arquivo "+item+"...")
url='http://download.inep.gov.br/microdados/'+item+'.zip'
wget.download(url,bar=bar_custom, out='zips')
print(" ok")
#os.system('wget '+url)
#break
print("fim dos downloads")
```
### 4- *Execute in loop for each downloaded .zip file the uncompress, convertion and upload functions*
```
#loop completo
bucket='edu-bucket-999'
for filename in sorted(os.listdir('zips')):
print(">>Processando zips/"+filename)
#extraindo arquivo csv
result_tr=microdados_transform('zips/'+filename)
#convertendo e comprimindo
result_conv=convert_compress_file(result_tr)
#enviando para o bucket s3
upload_s3(result_conv,bucket,folder)
```
### 5.1- *Define database name variable, and initialize boto3 object to execute AWS Glue related tasks*
```
#Change the database name if desired
dbname = 'db_education'
#Change the region name if necessary
region='us-east-1'
glue_client = boto3.client('glue', region_name=region)
```
### 5.2- *Creates Glue Crawler for S3 bucket where the microdata files were uploaded*
*Obs: Update the **rolename** variable with IAM Role created in CloudFormation stack*
```
#Criando Crawler
#Mude para o arn do Role criado no Cloudformation - coloque sua account ID e troque o nome se for o caso
rolename='arn:aws:iam::99999999999:role/AWSGlueServiceRole-Educ'
response = glue_client.create_crawler(
Name='enem-crawler',
Role=rolename,
DatabaseName=dbname,
Description='Crawler para bucket da educação',
Targets={
'S3Targets': [
{
'Path': 's3://'+bucket+'/'+folder,
'Exclusions': [
]
},
]
},
SchemaChangePolicy={
'UpdateBehavior': 'UPDATE_IN_DATABASE',
'DeleteBehavior': 'DELETE_FROM_DATABASE'
}
)
pprint(response)
```
### 5.4- *Run Crawler previously defined*
```
response = glue_client.start_crawler(Name='enem-crawler')
pprint(response)
```
---
### After executing steps above, it's available in Glue Data Catalog the microdata tables, that can be used in Amazon Athena and other tools/services.
| github_jupyter |
# Representing Data and Engineering Features
In the last chapter, we built our very first supervised learning models and applied them to
some classic datasets, such as the **Iris** and the **Boston** datasets. However, in the real world,
data rarely comes in a neat `<n_samples x n_features>` **feature matrix** that is part of a
pre-packaged database. Instead, it is our own responsibility to find a way to represent the
data in a meaningful way. The process of finding the best way to represent our data is
known as **feature engineering**, and it is one of the main tasks of data scientists and machine
learning practitioners trying to solve real-world problems.
I know you would rather jump right to the end and build the deepest neural network
mankind has ever seen. But, trust me, this stuff is important! Representing our data in the
right way can have a much greater influence on the performance of our supervised model
than the exact parameters we choose. And we get to invent our own features, too.
In this chapter, we will therefore go over some common feature engineering tasks.
Specifically, we want to answer the following questions:
- What are some common preprocessing techniques that everyone uses but nobody talks about?
- How do we represent categorical variables, such as the names of products, of colors, or of fruits?
- How would we even go about representing text?
- What is the best way to encode images, and what do SIFT and SURF stand for?
## Outline
- [Preprocessing Data](04.01-Preprocessing-Data.ipynb)
- [Reducing the Dimensionality of the Data](04.02-Reducing-the-Dimensionality-of-the-Data.ipynb)
- [Representing Categorical Variables](04.03-Representing-Categorical-Variables.ipynb)
- [Representing Text Features](04.04-Represening-Text-Features.ipynb)
- [Representing Images](04.05-Representing-Images.ipynb)
## Feature engineering
Feature engineering comes in two stages:
- **Feature selection**: This is the process of identifying important attributes (or
features) in the data. Possible features of an image might be the location of edges,
corners, or ridges. In this chapter, we will look at some of the more advanced
feature descriptors that OpenCV provides, such as the Scale-Invariant Feature
Transform (SIFT) and Speeded Up Robust Features (SURF).
- **Feature extraction**: This is the actual process of transforming the raw data into
the desired feature space used to feed a machine learning algorithm as illustrated
in the figure above. An example would be the Harris operator, which allows you
to extract corners (that is, a selected feature) in an image.
## Preprocessing data
The preprocessing pipeline:
- *Data formatting*: The data may not be in a format that is suitable for you to work
with. For example, the data might be provided in a proprietary file format, which
your favorite machine learning algorithm does not understand.
- *Data cleaning*: The data may contain invalid or missing entries, which need to be
cleaned up or removed.
- *Data sampling*: The data may be far too numerous for your specific purpose,
forcing you to sample the data in a smart way.
## Transforming data
Once the data has been preprocessed, you are ready for the actual feature engineering: to transform the
preprocessed data to fit your specific machine learning algorithm. This step usually involves one or more of
three possible procedures:
- *Scaling*: Machine learning algorithms often require the data to be within a
common range, such as to have zero mean and unit variance. Scaling is the
process of bringing all features (which might have different physical units) into a
common range of values.
- *Decomposition*: Datasets often have many more features than you could possibly
process. Feature decomposition is the process of compressing data into a smaller
number of highly informative data components.
- *Aggregation*: There may be features that can be aggregated into a single feature
that would be more meaningful to the problem you are trying to solve. For
example, a database might contain the date and time each user logged into a webbased
system. Depending on the task, this data might be better represented by
simply counting the number of logins per user.
| github_jupyter |
## Training Notebook
This notebook illustrates training of a simple model to classify digits using the MNIST dataset. This code is used to train the model included with the templates. This is meant to be a starter model to show you how to set up Serverless applications to do inferences. For deeper understanding of how to train a good model for MNIST, we recommend literature from the [MNIST website](http://yann.lecun.com/exdb/mnist/). The dataset is made available under a [Creative Commons Attribution-Share Alike 3.0](https://creativecommons.org/licenses/by-sa/3.0/) license.
```
# We'll use scikit-learn to load the dataset
! pip install -q scikit-learn==0.23.2
# Load the mnist dataset
from sklearn.datasets import fetch_openml
from sklearn.model_selection import train_test_split
X, y = fetch_openml('mnist_784', return_X_y=True)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=10000)
# Next, let's add code for deskewing images (we will use this to improve accuracy)
# This code comes from https://fsix.github.io/mnist/Deskewing.html
from scipy.ndimage import interpolation
def moments(image):
c0, c1 = np.mgrid[:image.shape[0], :image.shape[1]]
img_sum = np.sum(image)
m0 = np.sum(c0 * image) / img_sum
m1 = np.sum(c1 * image) / img_sum
m00 = np.sum((c0-m0)**2 * image) / img_sum
m11 = np.sum((c1-m1)**2 * image) / img_sum
m01 = np.sum((c0-m0) * (c1-m1) * image) / img_sum
mu_vector = np.array([m0,m1])
covariance_matrix = np.array([[m00, m01],[m01, m11]])
return mu_vector, covariance_matrix
def deskew(image):
c, v = moments(image)
alpha = v[0,1] / v[0,0]
affine = np.array([[1,0], [alpha,1]])
ocenter = np.array(image.shape) / 2.0
offset = c - np.dot(affine, ocenter)
return interpolation.affine_transform(image, affine, offset=offset)
def deskew_images(images):
output_images = []
for image in images:
output_images.append(deskew(image.reshape(28, 28)).flatten())
return np.array(output_images)
```
## XGBoost Model Training
For this example, we will train a simple XGBoost classifier using scikit-learn to classify the MNIST digits. We will then freeze the model using `save_model()`. This is same as the starter model file included with the SAM templates.
```
! pip -q install xgboost==1.1.1
import sklearn
import xgboost as xgb
import numpy as np
from sklearn.metrics import accuracy_score
print('Using versions: sklearn', sklearn.__version__, 'xgboost', xgb.__version__)
# Remove the "tree_method" and "gpu_id" params if you don't have CUDA-compatible GPU set up
xgb_model = xgb.XGBClassifier(n_jobs=-1, tree_method='gpu_hist', gpu_id=0)
xgb_model.fit(X_train, y_train, eval_metric='error')
accuracy = accuracy_score(y_test, xgb_model.predict(np.array(X_test)))
print('Accuracy without deskewing:', accuracy)
# Next, we will try with deskewing to see if accuracy improves
# Remove the "tree_method" and "gpu_id" params if you don't have CUDA-compatible GPU set up
xgb_model = xgb.XGBClassifier(n_jobs=-1, tree_method='gpu_hist', gpu_id=0)
xgb_model.fit(deskew_images(X_train), y_train, eval_metric='error')
accuracy = accuracy_score(y_test, xgb_model.predict(deskew_images(X_test)))
print('Accuracy with deskewing:', accuracy)
# Save model to the disk
xgb_model.save_model('xgb_digit_classifier.model')
```
| github_jupyter |
# Interpretation of the data
This notebook produces the tables shown in the paper. It is assumed that the scripts in `scripts/` have been run and the results have been written to `results` directory.
### Dependencies
```
import pandas as pd
import numpy as np
import glob
def create_table_of_results(directory):
files = [filename for filename in glob.glob("results/" + directory + "/*.csv")]
tables = [pd.read_csv(f, header=None).T for f in files]
for t in tables:
t.columns = list([t.iloc[0]])
t.drop(0, inplace=True)
all_results = pd.concat(tables)
all_results.columns = [y[0] for y in all_results.columns]
return all_results
def get_sig_figs(x, sig=1):
return round(x, -int(np.log10(x))+sig)
```
## Results for PIR with one plaintext payload
```
# Reading data
pir_data = create_table_of_results('pir-one-plaintext')
pir_data["db_size"] = pir_data["num_keywords"] * (2**pir_data["log_poly_mod_degree"]) * 20 / 8
```
### Folklore with N=8192
```
selected_data = pir_data[
(
(pir_data['eq_type'] == 0) &
(pir_data['log_poly_mod_degree'] == 13) &
(pir_data['num_threads'] == 1)
)
]
selected_data = selected_data.reset_index().drop(columns=["index"])
selected_data = selected_data.astype('int64')
averaged_data = selected_data.groupby(["num_keywords"], as_index=True).mean()
averaged_data = averaged_data.astype('int64')
for col in ["db_size", "time_query", "time_expansion","time_sel_vec", "time_inner_prod", "time_server_latency"]:
averaged_data[col] = averaged_data[col].apply(lambda x: "{:1g}".format(get_sig_figs(x/1000000)))
res = averaged_data.reset_index(level=0)[["num_keywords", "db_size", "encoding_size", "time_expansion", "time_sel_vec", "time_inner_prod", "time_server_latency"]]
res.astype('str').apply(lambda x: x + " &")
```
### Folklore with N=16384
```
selected_data = pir_data[
(
(pir_data['eq_type'] == 0) &
(pir_data['log_poly_mod_degree'] == 14) &
(pir_data['num_threads'] == 1)
)
]
selected_data = selected_data.reset_index().drop(columns=["index"])
selected_data = selected_data.astype('int64')
averaged_data = selected_data.groupby(["num_keywords"], as_index=True).mean()
averaged_data = averaged_data.astype('int64')
for col in ["db_size", "time_query", "time_expansion", "time_sel_vec", "time_inner_prod", "time_server_latency"]:
averaged_data[col] = averaged_data[col].apply(lambda x: "{:1g}".format(get_sig_figs(x/1000000)))
res = averaged_data.reset_index(level=0)[["num_keywords", "db_size", "encoding_size", "time_expansion", "time_sel_vec", "time_inner_prod", "time_server_latency"]]
res.astype('str').apply(lambda x: x + " &")
```
### Constant-weight k=2 Single Thread
```
selected_data = pir_data[
(
(pir_data['hamming_weight'] == 2) &
(pir_data['eq_type'] == 1) &
(pir_data['num_threads'] == 1)
)
]
selected_data = selected_data.reset_index().drop(columns=["index"])
selected_data = selected_data.astype('int64')
averaged_data = selected_data.groupby(["hamming_weight", "num_keywords"], as_index=True).mean()
averaged_data = averaged_data.astype('int64')
for col in ["time_query", "time_expansion",]:
averaged_data[col] = averaged_data[col].apply(lambda x: "{:.1g}".format(x/1000000))
for col in ["db_size", "time_sel_vec", "time_inner_prod", "time_server_latency"]:
averaged_data[col] = averaged_data[col].apply(lambda x: "{:1g}".format(get_sig_figs(x/1000000)))
averaged_data.index = averaged_data.index.droplevel(level=0)
res = averaged_data[["db_size", "encoding_size", "time_expansion", "time_sel_vec", "time_inner_prod", "time_server_latency"]]
res.astype('str').apply(lambda x: x + " &")
```
### Constant-weight k=2 Parallel
```
selected_data = pir_data[
(
(pir_data['hamming_weight'] == 2) &
(pir_data['eq_type'] == 1) &
(pir_data['num_threads'] == 64)
)
]
selected_data = selected_data.reset_index().drop(columns=["index"])
selected_data = selected_data.astype('int64')
averaged_data = selected_data.groupby(["hamming_weight", "num_keywords"], as_index=True).mean()
averaged_data = averaged_data.astype('int64')
for col in ["db_size", "time_query", "time_expansion","time_sel_vec", "time_inner_prod", "time_server_latency"]:
averaged_data[col] = averaged_data[col].apply(lambda x: "{:1g}".format(get_sig_figs(x/1000000)))
res = averaged_data.reset_index(level=0)[["db_size", "encoding_size", "time_expansion", "time_sel_vec", "time_inner_prod", "time_server_latency"]]
res.astype('str').apply(lambda x: x + " &")
```
### Unary Single Thread
```
selected_data = pir_data[
(
(pir_data['hamming_weight'] == 1) &
(pir_data['eq_type'] == 1) &
(pir_data['num_threads'] == 1)
)
]
selected_data = selected_data.reset_index().drop(columns=["index"])
selected_data = selected_data.astype('int64')
averaged_data = selected_data.groupby(["hamming_weight", "num_keywords"], as_index=True).mean()
averaged_data = averaged_data.astype('int64')
for col in ["time_query", "time_expansion",]:
averaged_data[col] = averaged_data[col].apply(lambda x: "{:1g}".format(get_sig_figs(x/1000000)))
for col in ["db_size", "time_sel_vec", "time_inner_prod", "time_server_latency"]:
averaged_data[col] = averaged_data[col].apply(lambda x: "{:1g}".format(get_sig_figs(x/1000000)))
res = averaged_data.reset_index(level=0)[["db_size", "encoding_size", "time_expansion", "time_sel_vec", "time_inner_prod", "time_server_latency"]]
res.astype('str').apply(lambda x: x + " &")
```
## Results for Databases with Large Payloads (large databases)
```
# Reading data
large_pir_data = create_table_of_results("pir-large-payload")
results = []
for h, kw in [(2, 16), (3, 32), (4, 48)]:
for nk in [1000, 10000]:
data = large_pir_data[(large_pir_data['num_keywords'] == nk) & (large_pir_data['hamming_weight']==h) & (large_pir_data["valid_response"] == 1)]
data = data.reset_index().drop(columns=["index"])
data = data.astype('int64')
data["db_size"] = (data["num_keywords"] * (2**data["log_poly_mod_degree"]) * data["num_output_ciphers"] * 20 / 8000000000).apply(lambda x : get_sig_figs(x))
data["item_size"] = ((2**data["log_poly_mod_degree"]) * data["num_output_ciphers"] * 20 / 8000000).apply(lambda x : get_sig_figs(x))
data = data[data.db_size >= 0.5]
data= data.groupby(["num_output_ciphers"], as_index=True).mean()
for col in ["time_server_latency", "time_expansion", "time_sel_vec", "time_inner_prod",]:
data[col] = data[col].apply(lambda x: "{:1g}".format(get_sig_figs(x/1000000,2)))
data["keyword_bitlength"] = kw
res = data[["keyword_bitlength", "num_keywords", "db_size", "item_size", "time_server_latency",]]
res = res.sort_values(by=["db_size"])
res.reset_index(drop=True, inplace=True)
res.set_index('keyword_bitlength', inplace=True)
res=res.astype('str').apply(lambda x: x + " &")
results += [res]
# hamming_weight = 2
# keyword bitlength = 16
# number of rows in database = 1000
results[0]
# hamming_weight = 2
# keyword bitlength = 16
# number of rows in database = 10000
results[1]
# hamming_weight = 3
# keyword bitlength = 32
# number of rows in database = 1000
results[2]
# hamming_weight = 3
# keyword bitlength = 32
# number of rows in database = 10000
results[3]
# hamming_weight = 4
# keyword bitlength = 48
# number of rows in database = 1000
results[4]
# hamming_weight = 4
# keyword bitlength = 48
# number of rows in database = 10000
results[5]
```
| github_jupyter |
```
```
## Neural Networks: Network, Layers & Neurons
Different architectures & neurons are possible to be used. In one way it's a very advanced LEGO, you just need to understand what each LEGO part is built of and how it interacts with everything else and how the LEGO works as a system all together.
A Layer in a Neural Network is built from multiple Neurons being stacked in width.
I'll start from Neurons and build it upwards to first understand the single LEGO piece and then the bigger picture.
### Backpropagation
Backpropagation is the key in neural network, it is used to calculate the _gradient_ that is required by the "update" function in the neuron where it recalculates its weights. Backpropagation is short for "the backward propagation of errors".
In simple terms, we push backward what we learned by our prediction and how well it went compared to the answer and we update our network automatically to create a better guess next time.
```
# PyTorch
# Fully connected neural network with one hidden layer
class NeuralNet(nn.Module):
def __init__(self, input_size, hidden_size, num_classes):
super(NeuralNet, self).__init__()
self.fc1 = nn.Linear(input_size, hidden_size)
self.relu = nn.ReLU()
self.fc2 = nn.Linear(hidden_size, num_classes)
def forward(self, x):
out = self.fc1(x)
out = self.relu(out)
out = self.fc2(out)
return out
# Init model
model = NeuralNet(input_size, hidden_size, num_classes).to(device)
# Loss and optimizer
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.Adam(model.parameters(), lr=learning_rate)
# Train the model
total_step = len(train_loader)
for epoch in range(num_epochs):
for i, (images, labels) in enumerate(train_loader):
# Move tensors to the configured device
images = images.reshape(-1, 28*28).to(device)
labels = labels.to(device)
# Forward pass
outputs = model(images)
loss = criterion(outputs, labels)
# Backward and optimize
optimizer.zero_grad()
loss.backward()
optimizer.step()
if (i+1) % 100 == 0:
print ('Epoch [{}/{}], Step [{}/{}], Loss: {:.4f}'
.format(epoch+1, num_epochs, i+1, total_step, loss.item()))
# Test the model
# In test phase, we don't need to compute gradients (for memory efficiency)
with torch.no_grad():
correct = 0
total = 0
for images, labels in test_loader:
images = images.reshape(-1, 28*28).to(device)
labels = labels.to(device)
outputs = model(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: {} %'.format(100 * correct / total))
# Save the model checkpoint
torch.save(model.state_dict(), 'model.ckpt')
# Keras
# create model
model = Sequential()
model.add(Dense(12, input_dim=8, activation='relu'))
model.add(Dense(1, activation='sigmoid'))
# Compile model
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
# Fit the model
model.fit(X, Y, epochs=150, batch_size=10)
# evaluate the model
scores = model.evaluate(X, Y)
print("\n%s: %.2f%%" % (model.metrics_names[1], scores[1]*100))
# calculate predictions
predictions = model.predict(X)
```
### Neurons
There exist an crazy amont of neurons that can be used and I'll try to quickly go through the most common ones.
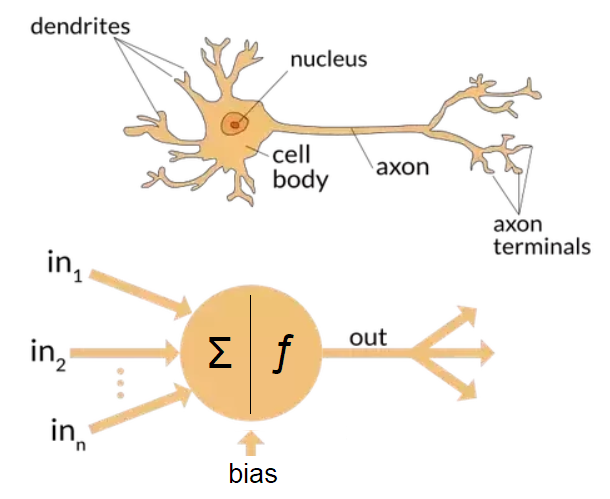
#### 1. Dense / Linear
Dense layers are the most simple. They're basically just an matrix (dot) multiplication. Each neuron receives input from all the neurons in the earlier layer, hence densely connected.
In Keras it's implemented as the following: `output = activation(dot(input, kernel) + bias)`
We also include a bias vector as well as the weight vector on this. The activation is selected when creating the layer.
#### 2. Dropout
Dropout is a way to combat overfitting. We randomly drop a node completely from the network, this is done by supplying a float inbetween 0 and 1.
There is another dropout called **Spatial Dropout**. If we think about CNNs this would rather remove the whole feature map (convolution) rather than individual pixels. So basically we drop an entire featuremap instead of an individual feature.

#### 3. Pooling
Pooling is used to reduce dimensions, by doing this we also make the network more independent in form of input / pattern transition / changes. It goes very well hand in hand with CNNs (which will be explained soon). We have a few different types of pooling such as Maxpool, Avgpool and Global XPool.
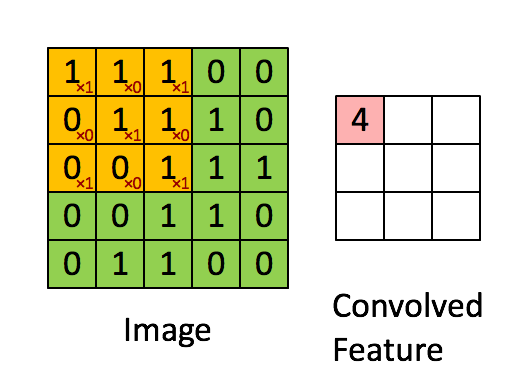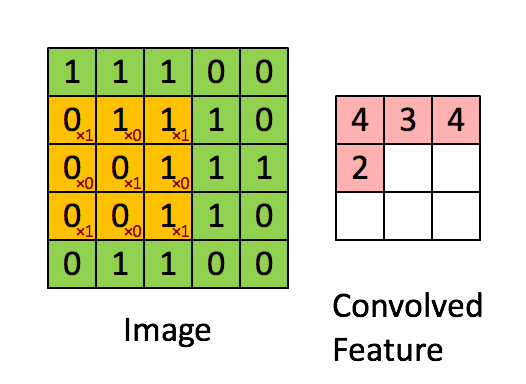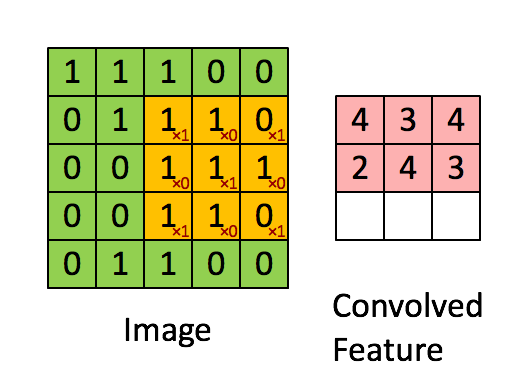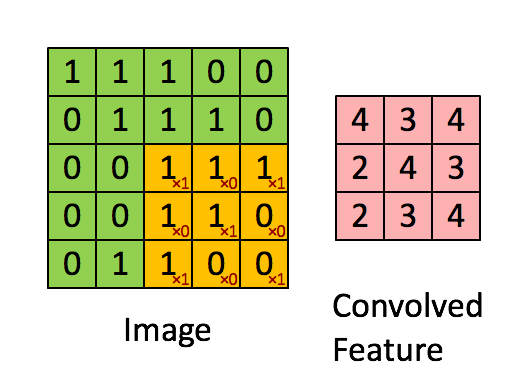
#### 4. Recurrent: LSTM & GRU
The "two" neurons that exist:
LSTM - Long Short Term Memory.
GRU - Gated Recurrent Unit
ELI5: We add a memory to the gate. This makes the network context aware which is great for natural language. We can now remember things over a sequence
More in depth we have three outputs of an LSTM and two out of an GRU. We have the cell-state (c), hidden state (h) & for LSTM we also have output (o). Important to add on this is that outside of memory we also have the possibility to forget.
LSTM & GRU solves the problem of vanishing gradient where when we backpropagation.
As they're very complex I won't go through the teory but just the basics.
I might extend this.
### Layers/Network
All from above are layers too.
#### 1. Convolutional
One can see Convolutional Neural Networks (CNNs) as a type of filter. We apply matrix multiplication to find some kind of feature. One example of this is in images one such matrix multiplication could be to find all edges in an image.
In CNNs, as with other Neural Networks, this is trained automatically by backpropagation.
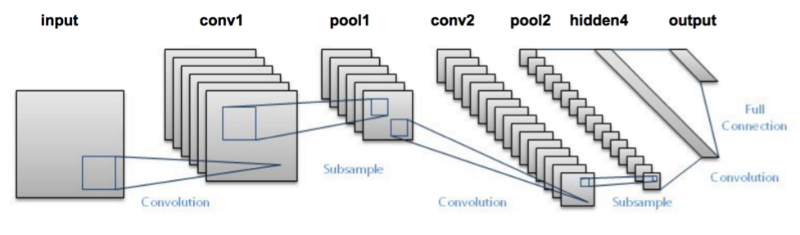
#### 2. Embedding
The embedding layer does exactly what the name suggests, embedds the feature. In our case we always talk about Word Embeddings, and I'm not sure there is any other use-case of embedding layers.
How we did it:
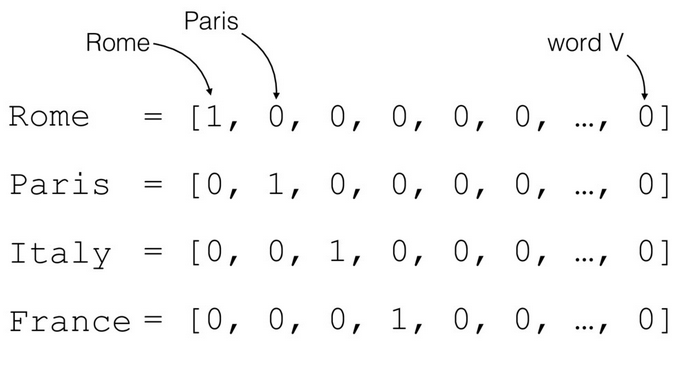

How Embedding does it (dense representation):

#### 3. Recurrent: LSTM & GRU
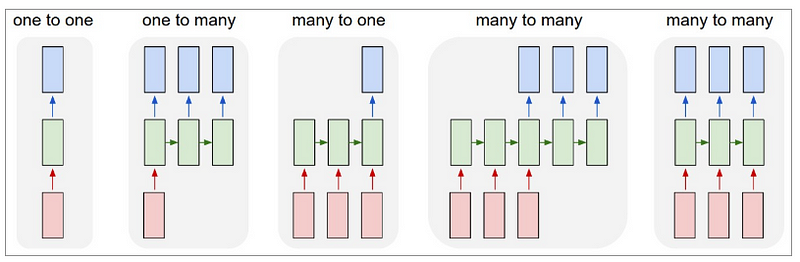
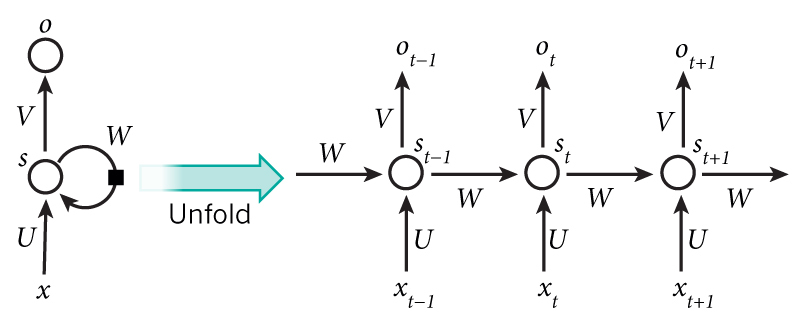
#### 4. Seq2Seq
Sequence 2 sequence is an special version RNNs.
#### Bonus: Capsule, Autoencoding & Attention
Let's **CODE**!
```
```
| github_jupyter |
# Introduction to DSP with PYNQ
# 02: DSP & PYNQ
> In the previous notebook, we used SciPy to analyse an audio recording of 2 birds and used filtering to isolate one of them. In this notebook the same techniques will be used but, this time, we'll be moving the software FFT and FIR functions over to FPGA hardware and controlling them using PYNQ as if they were software!
## Reusing Code
As this is a whole, new notebook we will need to: load in our audio file again, create a way to get our data into the pandas format for Plotly, and set up our plotting template. To keep things simple we can just reuse code from the previous lab rather than writing it all again.
```
from scipy.io import wavfile
import numpy as np
fs, aud_in = wavfile.read("assets/birds.wav")
import plotly_express as px
import pandas as pd
# Derive a custom plotting template from `plotly_dark`
import plotly.io as pio
new_template = pio.templates['plotly_white']
new_template.update(dict(layout = dict(
width = 800,
autosize = False,
legend = dict(x=1.1),
paper_bgcolor = 'rgb(0,0,0,0)',
plot_bgcolor = 'rgb(0,0,0,0)',
)))
# Register new template as the default
pio.templates['light_plot'] = new_template
pio.templates.default = 'light_plot'
def to_freq_dataframe(samples, fs):
"""Create a pandas dataframe from an ndarray frequency domain samples"""
sample_freqs = np.linspace(0, fs, len(samples))
return pd.DataFrame(dict(
amplitude = samples[0:int(len(samples)/2)],
freq = sample_freqs[0:int(len(samples)/2)]
))
```
## Moving to Hardware
The hardware design consists of a LogiCore FIR Compiler (with 99 reprogrammable weights), and a LogiCore FFT (with a fixed length of $2^{14}$). These are then connected, via AXI4-Stream, to the standard AXI DMAs that allow shared access to PS memory. The IPI diagram below shows how the filter and FFT IPs are both within their own hierarchy, this is for two reasons. One is to keep the top diagram simple and easy to follow, the other it makes referencing the IPs in PYNQ a little simpler.
<img src="assets/dsp_pynq_top.png" width="1200"/>
Of course, to get a better idea of how the hardware is set up, it's best to show you what is inside one of these hierarchies - let's look at the filter. You'll notice straight away that there's not a whole lot in here, just the FIR and a few DMAs for transferring our audio and configuration data. With PYNQ, this is all you need to be able to send data to and from your IP using AXI4-Stream!
<img src="assets/dsp_pynq_filt.png" width="900"/>
To start working with our design, we need to first download our bitstream onto the FPGA. PYNQ makes things simple by giving us the Overlay class...
```
from pynq import Overlay
ol = Overlay('assets/dsp_pynq.bit')
```
As easy as that! PYNQ also makes use of the accompanying Hardware Handoff (HWH) file to create a dictionary of the IP in the design. This helps with identifying any IP for which drivers can be automatically assigned, such as the DMAs. You can view the keys for IP dictionary by running the command in the cell below.
```
list(ol.ip_dict.keys())
```
Another great feature of PYNQ is having quick access to the entire register map of any IP in your design. As an example, let's have a look at our "fft_data" DMA. You can refer to [PG021](https://www.xilinx.com/support/documentation/ip_documentation/axi_dma/v7_1/pg021_axi_dma.pdf#page=12) to check these results.
```
ol.fft.fft_data.register_map
```
Now that we have a better idea of how the hardware looks, let's start off with the FFT and check our signal in the frequency domain...
## Hardware FFT
First off we need to create the DMA objects for the FFT IP. There are two associated DMAs here, one for data, and the other for configuration.
```
fft_data = ol.fft.fft_data
fft_config = ol.fft.fft_config
```
The IP will be set up for a forward FFT with a selected scaling schedule. We do this by sending a single, 16-bit packet to the FFT AXI4-Stream config port. This involves a few steps:
- First we create our config packet (in this case it's easier to show in binary)
- Next we create a contiguous memory buffer using allocate()
- Then we fill our buffer with the config packet
- Finally we transfer our packet to the DMA
To learn more about the FFT configuration, you can refer to [PG109](https://www.xilinx.com/support/documentation/ip_documentation/xfft/v9_1/pg109-xfft.pdf#page=16). And you can learn more about the DMA class in the [PYNQ documentation](https://pynq.readthedocs.io/en/v2.1/pynq_libraries/dma.html).
```
from pynq import allocate
def get_config_value(forwards, scaling_sched):
val = 0
for scaling in scaling_sched: # [14:1] = scaling schedule
val = (val << 2) + scaling
return (val << 1) + int(forwards) # [0] = direction
config_value = get_config_value(True, [1, 1, 2, 2, 2, 2, 2])
fft_buffer_config = allocate(shape=(1,),dtype=np.int16)
fft_buffer_config[0] = config_value
fft_config.sendchannel.transfer(fft_buffer_config)
fft_config.sendchannel.wait()
```
### Using the FFT
The LogiCore FFT IP data port expects a complex number with 16-bit components (32-bits in total) with the real part in the lower 2-bytes. It returns an equivalent complex 32-bit output as well.
As our input values are real only, we can just convert the signal to 32-bit values, ensuring the upper 2-bytes are 0.
```
# Imag[31:16] Real[15:0] --- imag all zeros
aud_hw = np.asarray(aud_in,np.int32)
```
Now all we need to do is set up our FFT data buffers and transfer the data. Our FFT is set up in *Non Real Time* throttle scheme, so for every frame of data we transmit, we need to read a frame of data out.
This would be simple if our signal was an exact multiple of a frame length (in our case 16384 samples), but unfortunately that rarely happens in the real world. To counteract this mismatch in length we need to append zeros to our signal up to the next frame length.
We can do this all within one function as shown in the next cell. You will recognise some of the syntax from the DMA transfer when we sent the configuration packet. The only difference here is that we also expect a packet *back* from the DMA this time as well, so we need to set up an output buffer.
```
def fft_hw(signal, NFFT):
# calculate how many NFFT frames are needed to iterate through entire signal
max_iters = np.int16(np.ceil(len(signal)/NFFT))
# calculate amount of zeros to add to make up to NFFT multiple
zeros = np.int16(np.ceil(len(signal)/NFFT))*NFFT - len(signal)
# increase length to multiple of NFFT
signal = np.int32(np.append(signal, np.zeros(zeros)))
fft_in_buffer = allocate(shape=(NFFT,),dtype=np.int32)
fft_out_buffer = allocate(shape=(NFFT*2,),dtype=np.int16)
fft_out = np.zeros(len(fft_out_buffer))
for i in range(0,max_iters):
np.copyto(fft_in_buffer,signal[NFFT*i:(NFFT*(i+1))])
fft_data.sendchannel.transfer(fft_in_buffer)
fft_data.recvchannel.transfer(fft_out_buffer)
fft_data.sendchannel.wait()
fft_data.recvchannel.wait()
fft_out = fft_out + np.array(fft_out_buffer)
fft_out_buffer.close()
fft_in_buffer.close()
return fft_out
```
It's important to note here that the function we just created now works in the same way as the SciPy equivalent that we used in the last lab (i.e. it takes the same data and the same arguments) - but this time the FFT is in hardware. This really shows the power of PYNQ: that you can so easily switch between hardware and software and never feel like you've moved between either!
With that said, let's apply the hardware FFT to our audio file...
```
NFFT = 2**14
# only perform FFT over small subset of data
aud_fft = fft_hw(aud_hw[np.int16(fs*0.3):np.int16(fs*0.718)],NFFT)
```
You may have noticed in the function definition that the output buffer is comprised of 16-bit integers while the input buffer has 32-bit integers. You may also have noticed that the length of the output array is double that of the input. Why are we doing this? Well, this is an intentional exploitation of a NumPy feature, where our 32-bit value will be reshaped into two 16-bit values. We use this to make it easier for ourselves to combine the complex output values together, seen in the cell below...
```
# make complex number x[n] + j*x[n+1]
aud_fft_c = np.int16(aud_fft[0::2])+1j*np.int16(aud_fft[1::2])
```
We then take the magnitude of our complex values and plot the results.
```
aud_fft_abs = np.abs(aud_fft_c)
# Plot FFT
px.line(
to_freq_dataframe(aud_fft_abs, fs),
x='freq', y='amplitude',
labels = dict(amplitude='Amplitude', freq='Freq (Hz)'),
template='light_plot'
)
```
Perfect! Now let's move onto filtering the signal with our hardware FIR...
## Hardware FIR Filter
The LogiCore FIR Compiler gives the user the ability to load and reload filter coefficients, on-the-fly, over AXI4-Stream. In this section we use this functionality to filter our audio data in hardware as well.
### Configuring the FIR
Similar to our FFT, we first have to set up the DMAs associated with the FIRs. There are 3 DMAs here, one for data and two for configuration.
```
dma_data = ol.filter.fir_data
dma_config = ol.filter.fir_config
dma_reload = ol.filter.fir_reload
```
One of the greatest benefits of using PYNQ is that it encourages us to mix our software and hardware in ways rarely implemented before. Remember the coefficients we designed with SciPy's `firwin` function in the previous notebook? We can use those to program the FIR in hardware!
The FIR Compiler is set up to accept 16-bit integers, so we will first need to convert them from their original type of *float*.
```
hpf_coeffs = np.load('assets/hpf_coeffs.npy')
hpf_coeffs_hw = np.int16(hpf_coeffs/np.max(abs(hpf_coeffs)) * 2**15 - 1)
```
and because our filter is symmetrical we need only to send half the weights and the FIR compiler will infer the rest...
```
hpf_coeffs_hw = hpf_coeffs_hw[0:int(len(hpf_coeffs_hw)/2)+1] # 1/2 + 1 weights
```
Now we can load the new coefficients to the FIR IP over AXI4-Stream using the same DMA transfer routine we used when configuring the FFT, albeit with an extra step. This *reload/config* transfer is explained in more detail in [PG149](https://www.xilinx.com/support/documentation/ip_documentation/fir_compiler/v7_2/pg149-fir-compiler.pdf#page=16).
```
# Create DMA buffer for coefs
fir_buffer_reload = allocate(shape=(len(hpf_coeffs_hw),),dtype=np.int16)
# Copy coefs to buffer
for i in range(len(hpf_coeffs_hw)):
fir_buffer_reload[i] = hpf_coeffs_hw[i]
# Transfer coefficients to FIR
dma_reload.sendchannel.transfer(fir_buffer_reload)
dma_reload.sendchannel.wait()
# Send an empty 8-bit packet to FIR config port to complete reload
fir_buffer_config = allocate(shape=(1,),dtype=np.int8)
fir_buffer_config[0] = 0
dma_config.sendchannel.transfer(fir_buffer_config)
dma_config.sendchannel.wait()
# Close the buffers
fir_buffer_reload.close()
fir_buffer_config.close()
```
### Using the FIR
Now we can try filtering the signal using our weights from SciPy. First we need to convert our `aud_hw` signal back to a 16-bit integer then, similarly to how we transferred data to and from the FFT, can do the same for our filter.
>You'll notice that, as with the FFT, the output buffer is again a different type from the input. In this case the 32-bit output is to take into account bit growth during the filtering process.
```
aud_hw = np.int16(aud_hw)
# Create DMA buffer
fir_in_buffer = allocate(shape=(len(aud_hw),),dtype=np.int16)
fir_out_buffer = allocate(shape=(len(aud_hw),),dtype=np.int32)
# Copy aud_hw to buffer
for i in range(len(aud_hw)):
fir_in_buffer[i] = aud_hw[i]
# Transfer
dma_data.sendchannel.transfer(fir_in_buffer)
dma_data.recvchannel.transfer(fir_out_buffer)
dma_data.sendchannel.wait()
dma_data.recvchannel.wait()
```
Then all we need to do is send the result from our FIR to our hardware FFT function, and then plot it!
```
# increase the dynamic range and send to FFT
aud_hpf = np.int16(fir_out_buffer/np.max(abs(fir_out_buffer)) * 2**15 - 1)
# only perform FFT over small subset of data
hpf_fft = fft_hw(aud_hpf[np.int16(fs*0.3):np.int16(fs*0.718)],NFFT)
# make complex number x[n] + j*x[n+1]
hpf_fft_c = np.int16(hpf_fft[0::2])+1j*np.int16(hpf_fft[1::2])
# Plot FFT
px.line(
to_freq_dataframe(np.abs(hpf_fft_c), fs),
x='freq', y='amplitude',
labels = dict(amplitude='Amplitude', freq='Freq (Hz)'),
template='light_plot'
)
```
We can see from the new FFT plot that the lower frequencies have been removed by our hardware filter! We can also check the output by converting the signal back to audio.
```
from IPython.display import Audio
scaled = np.int16(fir_out_buffer/np.max(abs(fir_out_buffer)) * 2**15 - 1)
wavfile.write('assets/hpf_hw.wav', fs, scaled)
Audio('assets/hpf_hw.wav')
```
Perfect! If you have time, try out the bandpass filter on your own and plot the results.
## Summary
Let's recap what we've covered in the second notebook:
* Reusing code between notebooks (including filter coefficients)
* Getting to know features unique to PYNQ
+ Viewing the IP dictionary and register maps
+ Transferring data between PS and PL using the DMA class
+ On-the-fly IP reconfiguration
* Creating Python functions for FPGA hardware
| github_jupyter |
```
#all_slow
#export
from fastai2.basics import *
from fastai2.vision.all import *
#default_exp vision.gan
#default_cls_lvl 3
#hide
from nbdev.showdoc import *
```
# GAN
> Basic support for [Generative Adversial Networks](https://arxiv.org/abs/1406.2661)
GAN stands for [Generative Adversarial Nets](https://arxiv.org/pdf/1406.2661.pdf) and were invented by Ian Goodfellow. The concept is that we train two models at the same time: a generator and a critic. The generator will try to make new images similar to the ones in a dataset, and the critic will try to classify real images from the ones the generator does. The generator returns images, the critic a single number (usually a probability, 0. for fake images and 1. for real ones).
We train them against each other in the sense that at each step (more or less), we:
1. Freeze the generator and train the critic for one step by:
- getting one batch of true images (let's call that `real`)
- generating one batch of fake images (let's call that `fake`)
- have the critic evaluate each batch and compute a loss function from that; the important part is that it rewards positively the detection of real images and penalizes the fake ones
- update the weights of the critic with the gradients of this loss
2. Freeze the critic and train the generator for one step by:
- generating one batch of fake images
- evaluate the critic on it
- return a loss that rewards posisitively the critic thinking those are real images
- update the weights of the generator with the gradients of this loss
> Note: The fastai library provides support for training GANs through the GANTrainer, but doesn't include more than basic models.
## Wrapping the modules
```
#export
class GANModule(Module):
"Wrapper around a `generator` and a `critic` to create a GAN."
def __init__(self, generator=None, critic=None, gen_mode=False):
if generator is not None: self.generator=generator
if critic is not None: self.critic =critic
store_attr(self, 'gen_mode')
def forward(self, *args):
return self.generator(*args) if self.gen_mode else self.critic(*args)
def switch(self, gen_mode=None):
"Put the module in generator mode if `gen_mode`, in critic mode otherwise."
self.gen_mode = (not self.gen_mode) if gen_mode is None else gen_mode
```
This is just a shell to contain the two models. When called, it will either delegate the input to the `generator` or the `critic` depending of the value of `gen_mode`.
```
show_doc(GANModule.switch)
```
By default (leaving `gen_mode` to `None`), this will put the module in the other mode (critic mode if it was in generator mode and vice versa).
```
#export
@delegates(ConvLayer.__init__)
def basic_critic(in_size, n_channels, n_features=64, n_extra_layers=0, norm_type=NormType.Batch, **kwargs):
"A basic critic for images `n_channels` x `in_size` x `in_size`."
layers = [ConvLayer(n_channels, n_features, 4, 2, 1, norm_type=None, **kwargs)]
cur_size, cur_ftrs = in_size//2, n_features
layers += [ConvLayer(cur_ftrs, cur_ftrs, 3, 1, norm_type=norm_type, **kwargs) for _ in range(n_extra_layers)]
while cur_size > 4:
layers.append(ConvLayer(cur_ftrs, cur_ftrs*2, 4, 2, 1, norm_type=norm_type, **kwargs))
cur_ftrs *= 2 ; cur_size //= 2
init = kwargs.get('init', nn.init.kaiming_normal_)
layers += [init_default(nn.Conv2d(cur_ftrs, 1, 4, padding=0), init), Flatten()]
return nn.Sequential(*layers)
#export
class AddChannels(Module):
"Add `n_dim` channels at the end of the input."
def __init__(self, n_dim): self.n_dim=n_dim
def forward(self, x): return x.view(*(list(x.shape)+[1]*self.n_dim))
#export
@delegates(ConvLayer.__init__)
def basic_generator(out_size, n_channels, in_sz=100, n_features=64, n_extra_layers=0, **kwargs):
"A basic generator from `in_sz` to images `n_channels` x `out_size` x `out_size`."
cur_size, cur_ftrs = 4, n_features//2
while cur_size < out_size: cur_size *= 2; cur_ftrs *= 2
layers = [AddChannels(2), ConvLayer(in_sz, cur_ftrs, 4, 1, transpose=True, **kwargs)]
cur_size = 4
while cur_size < out_size // 2:
layers.append(ConvLayer(cur_ftrs, cur_ftrs//2, 4, 2, 1, transpose=True, **kwargs))
cur_ftrs //= 2; cur_size *= 2
layers += [ConvLayer(cur_ftrs, cur_ftrs, 3, 1, 1, transpose=True, **kwargs) for _ in range(n_extra_layers)]
layers += [nn.ConvTranspose2d(cur_ftrs, n_channels, 4, 2, 1, bias=False), nn.Tanh()]
return nn.Sequential(*layers)
critic = basic_critic(64, 3)
generator = basic_generator(64, 3)
tst = GANModule(critic=critic, generator=generator)
real = torch.randn(2, 3, 64, 64)
real_p = tst(real)
test_eq(real_p.shape, [2,1])
tst.switch() #tst is now in generator mode
noise = torch.randn(2, 100)
fake = tst(noise)
test_eq(fake.shape, real.shape)
tst.switch() #tst is back in critic mode
fake_p = tst(fake)
test_eq(fake_p.shape, [2,1])
#export
_conv_args = dict(act_cls = partial(nn.LeakyReLU, negative_slope=0.2), norm_type=NormType.Spectral)
def _conv(ni, nf, ks=3, stride=1, self_attention=False, **kwargs):
if self_attention: kwargs['xtra'] = SelfAttention(nf)
return ConvLayer(ni, nf, ks=ks, stride=stride, **_conv_args, **kwargs)
#export
@delegates(ConvLayer)
def DenseResBlock(nf, norm_type=NormType.Batch, **kwargs):
"Resnet block of `nf` features. `conv_kwargs` are passed to `conv_layer`."
return SequentialEx(ConvLayer(nf, nf, norm_type=norm_type, **kwargs),
ConvLayer(nf, nf, norm_type=norm_type, **kwargs),
MergeLayer(dense=True))
#export
def gan_critic(n_channels=3, nf=128, n_blocks=3, p=0.15):
"Critic to train a `GAN`."
layers = [
_conv(n_channels, nf, ks=4, stride=2),
nn.Dropout2d(p/2),
DenseResBlock(nf, **_conv_args)]
nf *= 2 # after dense block
for i in range(n_blocks):
layers += [
nn.Dropout2d(p),
_conv(nf, nf*2, ks=4, stride=2, self_attention=(i==0))]
nf *= 2
layers += [
ConvLayer(nf, 1, ks=4, bias=False, padding=0, norm_type=NormType.Spectral, act_cls=None),
Flatten()]
return nn.Sequential(*layers)
#export
class GANLoss(GANModule):
"Wrapper around `crit_loss_func` and `gen_loss_func`"
def __init__(self, gen_loss_func, crit_loss_func, gan_model):
super().__init__()
store_attr(self, 'gen_loss_func,crit_loss_func,gan_model')
def generator(self, output, target):
"Evaluate the `output` with the critic then uses `self.gen_loss_func`"
fake_pred = self.gan_model.critic(output)
self.gen_loss = self.gen_loss_func(fake_pred, output, target)
return self.gen_loss
def critic(self, real_pred, input):
"Create some `fake_pred` with the generator from `input` and compare them to `real_pred` in `self.crit_loss_func`."
fake = self.gan_model.generator(input).requires_grad_(False)
fake_pred = self.gan_model.critic(fake)
self.crit_loss = self.crit_loss_func(real_pred, fake_pred)
return self.crit_loss
```
In generator mode, this loss function expects the `output` of the generator and some `target` (a batch of real images). It will evaluate if the generator successfully fooled the critic using `gen_loss_func`. This loss function has the following signature
```
def gen_loss_func(fake_pred, output, target):
```
to be able to combine the output of the critic on `output` (which the first argument `fake_pred`) with `output` and `target` (if you want to mix the GAN loss with other losses for instance).
In critic mode, this loss function expects the `real_pred` given by the critic and some `input` (the noise fed to the generator). It will evaluate the critic using `crit_loss_func`. This loss function has the following signature
```
def crit_loss_func(real_pred, fake_pred):
```
where `real_pred` is the output of the critic on a batch of real images and `fake_pred` is generated from the noise using the generator.
```
#export
class AdaptiveLoss(Module):
"Expand the `target` to match the `output` size before applying `crit`."
def __init__(self, crit): self.crit = crit
def forward(self, output, target):
return self.crit(output, target[:,None].expand_as(output).float())
#export
def accuracy_thresh_expand(y_pred, y_true, thresh=0.5, sigmoid=True):
"Compute accuracy after expanding `y_true` to the size of `y_pred`."
if sigmoid: y_pred = y_pred.sigmoid()
return ((y_pred>thresh).byte()==y_true[:,None].expand_as(y_pred).byte()).float().mean()
```
## Callbacks for GAN training
```
#export
def set_freeze_model(m, rg):
for p in m.parameters(): p.requires_grad_(rg)
#export
class GANTrainer(Callback):
"Handles GAN Training."
run_after = TrainEvalCallback
def __init__(self, switch_eval=False, clip=None, beta=0.98, gen_first=False, show_img=True):
store_attr(self, 'switch_eval,clip,gen_first,show_img')
self.gen_loss,self.crit_loss = AvgSmoothLoss(beta=beta),AvgSmoothLoss(beta=beta)
def _set_trainable(self):
train_model = self.generator if self.gen_mode else self.critic
loss_model = self.generator if not self.gen_mode else self.critic
set_freeze_model(train_model, True)
set_freeze_model(loss_model, False)
if self.switch_eval:
train_model.train()
loss_model.eval()
def before_fit(self):
"Initialize smootheners."
self.generator,self.critic = self.model.generator,self.model.critic
self.gen_mode = self.gen_first
self.switch(self.gen_mode)
self.crit_losses,self.gen_losses = [],[]
self.gen_loss.reset() ; self.crit_loss.reset()
#self.recorder.no_val=True
#self.recorder.add_metric_names(['gen_loss', 'disc_loss'])
#self.imgs,self.titles = [],[]
def before_validate(self):
"Switch in generator mode for showing results."
self.switch(gen_mode=True)
def before_batch(self):
"Clamp the weights with `self.clip` if it's not None, set the correct input/target."
if self.training and self.clip is not None:
for p in self.critic.parameters(): p.data.clamp_(-self.clip, self.clip)
if not self.gen_mode:
(self.learn.xb,self.learn.yb) = (self.yb,self.xb)
def after_batch(self):
"Record `last_loss` in the proper list."
if not self.training: return
if self.gen_mode:
self.gen_loss.accumulate(self.learn)
self.gen_losses.append(self.gen_loss.value)
self.last_gen = to_detach(self.pred)
else:
self.crit_loss.accumulate(self.learn)
self.crit_losses.append(self.crit_loss.value)
def before_epoch(self):
"Put the critic or the generator back to eval if necessary."
self.switch(self.gen_mode)
#def after_epoch(self):
# "Show a sample image."
# if not hasattr(self, 'last_gen') or not self.show_img: return
# data = self.learn.data
# img = self.last_gen[0]
# norm = getattr(data,'norm',False)
# if norm and norm.keywords.get('do_y',False): img = data.denorm(img)
# img = data.train_ds.y.reconstruct(img)
# self.imgs.append(img)
# self.titles.append(f'Epoch {epoch}')
# pbar.show_imgs(self.imgs, self.titles)
# return add_metrics(last_metrics, [getattr(self.smoothenerG,'smooth',None),getattr(self.smoothenerC,'smooth',None)])
def switch(self, gen_mode=None):
"Switch the model and loss function, if `gen_mode` is provided, in the desired mode."
self.gen_mode = (not self.gen_mode) if gen_mode is None else gen_mode
self._set_trainable()
self.model.switch(gen_mode)
self.loss_func.switch(gen_mode)
```
> Warning: The GANTrainer is useless on its own, you need to complete it with one of the following switchers
```
#export
class FixedGANSwitcher(Callback):
"Switcher to do `n_crit` iterations of the critic then `n_gen` iterations of the generator."
run_after = GANTrainer
def __init__(self, n_crit=1, n_gen=1): store_attr(self, 'n_crit,n_gen')
def before_train(self): self.n_c,self.n_g = 0,0
def after_batch(self):
"Switch the model if necessary."
if not self.training: return
if self.learn.gan_trainer.gen_mode:
self.n_g += 1
n_iter,n_in,n_out = self.n_gen,self.n_c,self.n_g
else:
self.n_c += 1
n_iter,n_in,n_out = self.n_crit,self.n_g,self.n_c
target = n_iter if isinstance(n_iter, int) else n_iter(n_in)
if target == n_out:
self.learn.gan_trainer.switch()
self.n_c,self.n_g = 0,0
#export
class AdaptiveGANSwitcher(Callback):
"Switcher that goes back to generator/critic when the loss goes below `gen_thresh`/`crit_thresh`."
run_after = GANTrainer
def __init__(self, gen_thresh=None, critic_thresh=None):
store_attr(self, 'gen_thresh,critic_thresh')
def after_batch(self):
"Switch the model if necessary."
if not self.training: return
if self.gan_trainer.gen_mode:
if self.gen_thresh is None or self.loss < self.gen_thresh: self.gan_trainer.switch()
else:
if self.critic_thresh is None or self.loss < self.critic_thresh: self.gan_trainer.switch()
#export
class GANDiscriminativeLR(Callback):
"`Callback` that handles multiplying the learning rate by `mult_lr` for the critic."
run_after = GANTrainer
def __init__(self, mult_lr=5.): self.mult_lr = mult_lr
def before_batch(self):
"Multiply the current lr if necessary."
if not self.learn.gan_trainer.gen_mode and self.training:
self.learn.opt.set_hyper('lr', self.learn.opt.hypers[0]['lr']*self.mult_lr)
def after_batch(self):
"Put the LR back to its value if necessary."
if not self.learn.gan_trainer.gen_mode: self.learn.opt.set_hyper('lr', self.learn.opt.hypers[0]['lr']/self.mult_lr)
```
## GAN data
```
#export
class InvisibleTensor(TensorBase):
def show(self, ctx=None, **kwargs): return ctx
#export
def generate_noise(fn, size=100): return cast(torch.randn(size), InvisibleTensor)
#export
@typedispatch
def show_batch(x:InvisibleTensor, y:TensorImage, samples, ctxs=None, max_n=10, nrows=None, ncols=None, figsize=None, **kwargs):
if ctxs is None: ctxs = get_grid(min(len(samples), max_n), nrows=nrows, ncols=ncols, figsize=figsize)
ctxs = show_batch[object](x, y, samples, ctxs=ctxs, max_n=max_n, **kwargs)
return ctxs
#export
@typedispatch
def show_results(x:InvisibleTensor, y:TensorImage, samples, outs, ctxs=None, max_n=10, nrows=None, ncols=None, figsize=None, **kwargs):
if ctxs is None: ctxs = get_grid(min(len(samples), max_n), nrows=nrows, ncols=ncols, add_vert=1, figsize=figsize)
ctxs = [b.show(ctx=c, **kwargs) for b,c,_ in zip(outs.itemgot(0),ctxs,range(max_n))]
return ctxs
bs = 128
size = 64
dblock = DataBlock(blocks = (TransformBlock, ImageBlock),
get_x = generate_noise,
get_items = get_image_files,
splitter = IndexSplitter([]),
item_tfms=Resize(size, method=ResizeMethod.Crop),
batch_tfms = Normalize.from_stats(torch.tensor([0.5,0.5,0.5]), torch.tensor([0.5,0.5,0.5])))
path = untar_data(URLs.LSUN_BEDROOMS)
dls = dblock.dataloaders(path, path=path, bs=bs)
dls.show_batch(max_n=16)
```
## GAN Learner
```
#export
def gan_loss_from_func(loss_gen, loss_crit, weights_gen=None):
"Define loss functions for a GAN from `loss_gen` and `loss_crit`."
def _loss_G(fake_pred, output, target, weights_gen=weights_gen):
ones = fake_pred.new_ones(fake_pred.shape[0])
weights_gen = ifnone(weights_gen, (1.,1.))
return weights_gen[0] * loss_crit(fake_pred, ones) + weights_gen[1] * loss_gen(output, target)
def _loss_C(real_pred, fake_pred):
ones = real_pred.new_ones (real_pred.shape[0])
zeros = fake_pred.new_zeros(fake_pred.shape[0])
return (loss_crit(real_pred, ones) + loss_crit(fake_pred, zeros)) / 2
return _loss_G, _loss_C
#export
def _tk_mean(fake_pred, output, target): return fake_pred.mean()
def _tk_diff(real_pred, fake_pred): return real_pred.mean() - fake_pred.mean()
#export
@delegates()
class GANLearner(Learner):
"A `Learner` suitable for GANs."
def __init__(self, dls, generator, critic, gen_loss_func, crit_loss_func, switcher=None, gen_first=False,
switch_eval=True, show_img=True, clip=None, cbs=None, metrics=None, **kwargs):
gan = GANModule(generator, critic)
loss_func = GANLoss(gen_loss_func, crit_loss_func, gan)
if switcher is None: switcher = FixedGANSwitcher(n_crit=5, n_gen=1)
trainer = GANTrainer(clip=clip, switch_eval=switch_eval, gen_first=gen_first, show_img=show_img)
cbs = L(cbs) + L(trainer, switcher)
metrics = L(metrics) + L(*LossMetrics('gen_loss,crit_loss'))
super().__init__(dls, gan, loss_func=loss_func, cbs=cbs, metrics=metrics, **kwargs)
@classmethod
def from_learners(cls, gen_learn, crit_learn, switcher=None, weights_gen=None, **kwargs):
"Create a GAN from `learn_gen` and `learn_crit`."
losses = gan_loss_from_func(gen_learn.loss_func, crit_learn.loss_func, weights_gen=weights_gen)
return cls(gen_learn.dls, gen_learn.model, crit_learn.model, *losses, switcher=switcher, **kwargs)
@classmethod
def wgan(cls, dls, generator, critic, switcher=None, clip=0.01, switch_eval=False, **kwargs):
"Create a WGAN from `data`, `generator` and `critic`."
return cls(dls, generator, critic, _tk_mean, _tk_diff, switcher=switcher, clip=clip, switch_eval=switch_eval, **kwargs)
GANLearner.from_learners = delegates(to=GANLearner.__init__)(GANLearner.from_learners)
GANLearner.wgan = delegates(to=GANLearner.__init__)(GANLearner.wgan)
from fastai2.callback.all import *
generator = basic_generator(64, n_channels=3, n_extra_layers=1)
critic = basic_critic (64, n_channels=3, n_extra_layers=1, act_cls=partial(nn.LeakyReLU, negative_slope=0.2))
learn = GANLearner.wgan(dls, generator, critic, opt_func = RMSProp)
learn.recorder.train_metrics=True
learn.recorder.valid_metrics=False
learn.fit(1, 2e-4, wd=0.)
learn.show_results(max_n=9, ds_idx=0)
```
## Export -
```
#hide
from nbdev.export import notebook2script
notebook2script()
```
| github_jupyter |
```
%reload_ext autoreload
%autoreload 2
%matplotlib inline
# To support both python 2 and python 3
from __future__ import division, print_function, unicode_literals
# Common imports
import numpy as np
import os
import pandas as pd
import featuretools as ft
import matplotlib as mpl
import matplotlib.pyplot as plt
import re
import datetime
from datetime import date
from pathlib import Path
# to make this notebook's output stable across runs
np.random.seed(42)
mpl.rc('axes', labelsize=14)
mpl.rc('xtick', labelsize=12)
mpl.rc('ytick', labelsize=12)
# Ignore useless warnings (see SciPy issue #5998)
import warnings
warnings.filterwarnings(action="ignore", message="^internal gelsd")
from IPython.display import display
pd.options.display.max_columns = 50
pd.options.display.html.table_schema = True
```
## Functions
```
def add_datepart(df, fldname, drop=True, time=False):
"""Helper function that adds columns relevant to a date."""
fld = df[fldname]
fld_dtype = fld.dtype
if isinstance(fld_dtype, pd.core.dtypes.dtypes.DatetimeTZDtype):
fld_dtype = np.datetime64
if not np.issubdtype(fld_dtype, np.datetime64):
df[fldname] = fld = pd.to_datetime(fld, infer_datetime_format=True)
targ_pre = re.sub('[Dd]ate$', '', fldname)
attr = ['Year', 'Month', 'Week', 'Day', 'Dayofweek', 'Dayofyear',
'Is_month_end', 'Is_month_start', 'Is_quarter_end', 'Is_quarter_start', 'Is_year_end', 'Is_year_start']
if time: attr = attr + ['Hour', 'Minute', 'Second']
for n in attr: df[targ_pre + n] = getattr(fld.dt, n.lower())
df[targ_pre + 'Elapsed'] = fld.astype(np.int64) // 10 ** 9
if drop: df.drop(fldname, axis=1, inplace=True)
```
## Constants
```
TOTAL_NUMBERS = 10000
```
## Load Data
```
PATH = Path("datasets/lotto")
DATASET = PATH/'data_all.csv'
DATASET_PROCESSED = PATH/'data_processed.csv'
# df will store the original dataset
dataset = pd.read_csv(DATASET, parse_dates=['DrawDate'], dtype={'PrizeType': str})
dataset.columns
dataset.info()
dataset.dtypes
dataset.nunique()
len(dataset)
```
## Train Test Split
```
from sklearn.model_selection import train_test_split
train_df, test_df = train_test_split(dataset, test_size=0.20, shuffle=False)
combined_df = [train_df, test_df]
display(len(train_df))
display(len(test_df))
```
## Feature generation
#### Days since last exact match
```
df = dataset.copy()
df['DaysSinceLastExactMatch'] = 0
result = df.loc[df.LuckyNo == 1234]
display(len(result), result)
datetime.datetime.strptime('1997-07-12', '%Y-%m-%d').date() - datetime.datetime.strptime('1992-05-28', '%Y-%m-%d').date()
# df_temp['DaysSinceLastExactMatch'] = (df_temp.DrawDate - df_temp.loc[df_temp.LuckyNo == 9999].DrawDate.shift(1)).dt.days
# df_temp.loc[df_temp.LuckyNo == 9999].head(3)
matched_dates = df.loc[df.LuckyNo == 1234].DrawDate.values
days = np.diff(matched_dates).astype('timedelta64[D]')
days = np.insert(days, 0,0)
display(matched_dates, days)
len(days)
def get_elapsed_days(data, no, fld, func):
"""Get elapsed between the draw dates"""
df_filtered = data[func(data.LuckyNo, no)]
days = np.absolute(np.diff(df_filtered.DrawDate.values).astype('timedelta64[D]')).astype('int64')
days = np.insert(days, 0,0)
for idx, val in zip(df_filtered.index, days):
data.at[idx, fld] = val
def exact_match(lucky_no, no):
return lucky_no == no
df.sort_values(by=['DrawDate'], ascending=True, inplace=True)
for no in range(0, TOTAL_NUMBERS):
if no % 1000 == 0:
print('Processing %s of 10000' % str(no))
get_elapsed_days(df, no, 'DaysSinceLastExactMatch', exact_match)
display(df[df.LuckyNo==1234])
```
#### Days Until Next Exact Match
```
df['DaysUntilNextExactMatch'] = 0
df.tail(20)
df.sort_values(by=['DrawDate'], ascending=False, inplace=True)
display(df[df.LuckyNo==1234])
for no in range(0, TOTAL_NUMBERS):
if no % 1000 == 0:
print('Processing %s of 10000' % str(no))
get_elapsed_days(df, no, 'DaysUntilNextExactMatch', exact_match)
display(df[df.LuckyNo==1234])
```
#### Days Since Last Any Match
```
def pad(val):
return str(val).zfill(4)
from itertools import permutations
def get_permutations(no):
no_list = []
for p in list(set(permutations(pad(no)))):
no_list.append(int(''.join(p)))
return no_list
def any_match(lucky_no, no):
return lucky_no.isin(get_permutations(no))
df['DaysSinceLastAnyMatch'] = 0
df.sort_values(by=['DrawDate'], ascending=True, inplace=True)
display(df[df.LuckyNo==1234])
get_elapsed_days(df, 1234, 'DaysSinceLastAnyMatch', any_match)
display(df[df.LuckyNo.isin(get_permutations(1234))].head(10))
for no in range(0, TOTAL_NUMBERS):
if no % 1000 == 0:
print('Processing %s of 10000' % str(no))
get_elapsed_days(df, no, 'DaysSinceLastAnyMatch', any_match)
display(df[df.LuckyNo.isin(get_permutations(123))].head(20))
```
#### Days Until Next Any Match
```
df['DaysUntilNextAnyMatch'] = 0
df.sort_values(by=['DrawDate'], ascending=False, inplace=True)
display(df[df.LuckyNo.isin(get_permutations(123))].head(10))
for no in range(0, TOTAL_NUMBERS):
if no % 1000 == 0:
print('Processing %s of 10000' % str(no))
get_elapsed_days(df, no, 'DaysUntilNextAnyMatch', any_match)
display(df[df.LuckyNo.isin(get_permutations(123))].head(10))
```
#### Extract digits by position
```
df['LuckyNo_str'] = df['LuckyNo'].apply(str).apply(pad)
df['1st_digit'] = df['LuckyNo_str'].str[0:1].apply(int)
df['2nd_digit'] = df['LuckyNo_str'].str[1:2].apply(int)
df['3rd_digit'] = df['LuckyNo_str'].str[2:3].apply(int)
df['4th_digit'] = df['LuckyNo_str'].str[3:4].apply(int)
df.drop(columns=['LuckyNo_str'], axis=1, inplace=True)
display(df[df.LuckyNo.isin(get_permutations(123))].head(10))
df.info()
```
#### Add Date Parts
```
columns_to_drop = ['DrawIs_month_end','DrawIs_month_start', 'DrawIs_quarter_end', 'DrawIs_quarter_start', 'DrawIs_year_end', 'DrawIs_year_start' ,'DrawElapsed']
add_datepart(df, 'DrawDate', drop=False)
df.drop(columns=columns_to_drop, axis=1, inplace=True)
display(df[df.LuckyNo.isin(get_permutations(123))].head(10))
```
#### Calculate Combinations Per X Draws
```
df.sort_values(by=['DrawDate', 'PrizeType'], ascending=False, inplace=True)
df.head(23)
# Derive number of combinations for the past X draw
X = 3
total_numbers = 23 * X
print(total_numbers)
def get_combinations(df, dates):
total_distinct_combinations = 0
total_random_combinations = 0
combination_list = {}
for no in df[df.DrawDate.isin(dates)].LuckyNo:
found = False
for k, v in combination_list.items():
perms = get_permutations(k)
if no in perms:
found = True
combination_list[k] = combination_list.get(k) + 1
if not found:
combination_list[no] = 1
return combination_list
# get_combinations(df,['2019-06-09', '2019-06-08'])
# Past 10 draws
result = get_combinations(df,df.DrawDate[0:total_numbers].unique())
print(total_numbers, len(result), result)
```
#### Cumulative Probability Exact Match
```
df['CumProbaExactMatch'] = 0
df.sort_values(by=['DrawDate', 'PrizeType'], ascending=True, inplace=True)
df[df.LuckyNo.isin(get_permutations(2646))].tail(10)
df[df.LuckyNo.isin([2646])].tail(10)
def get_cum_proba_match(df, numbers, field):
"""Cumulative probability for the numbers"""
count = 0
for index, row in df[df.LuckyNo.isin(numbers)].iterrows():
count = count + 1
# print(index)
all_numbers = len(df.iloc[0:index]) + 1
proba = round((count / TOTAL_NUMBERS) * 100,5)
# print(index, row['DrawDate'], all_numbers, proba)
# print(row)
df[field].iloc[index] = proba
for no in range(0, TOTAL_NUMBERS):
if no % 1000 == 0:
print('Processing %s of 10000' % str(no))
get_cum_proba_match(df, [no], 'CumProbaExactMatch')
df.tail(10)
```
#### Cumulative Probability Any Match
```
df['CumProbaAnyMatch'] = 0
df.sort_values(by=['DrawDate', 'PrizeType'], ascending=True, inplace=True)
df[df.LuckyNo.isin(get_permutations(2646))].tail(10)
no_list = []
for no in range(0, TOTAL_NUMBERS):
if no % 1000 == 0:
print('Processing %s of 10000' % str(no))
if no not in no_list:
nos = get_permutations(no)
# breakpoint()
no_list.extend(nos)
get_cum_proba_match(df, nos , 'CumProbaAnyMatch')
df.tail(10)
df[df.LuckyNo.isin(get_permutations(2976))].head(33)
df.dtypes
```
#### Digits Pairings
##### Create the columns
```
columns = ['DrawDate', '1st_digit', '2nd_digit', '3rd_digit', '4th_digit' ]
new_columns = []
import itertools
digits = list(range(0,10))
pairings = list(itertools.combinations(digits,2))
for p in pairings:
new_columns.append("_".join(str(i) for i in p))
new_columns.extend(["0_0", "1_1", "2_2", "3_3", "4_4", "5_5", "6_6", "7_7", "8_8", "9_9"])
print(new_columns)
```
##### Derive the pairings
```
def get_pairings(df, no):
pairs = list(set(itertools.combinations(no, 2)))
cols = []
for p in pairs:
cols.append("_".join(d for d in sorted(p)))
return cols
for c in new_columns:
df[c] = 0
stats = {}
count = 0
for draw_date in df.DrawDate.unique():
if count % 1000 == 0:
print(f"Working on {draw_date}")
count = count + 1
for index, row in df[df.DrawDate == draw_date].iterrows():
pairings = get_pairings(df, pad(row.LuckyNo))
for p in pairings:
if p in df.columns:
# df.at[index, p] = df.at[index, p] + 1
if p in stats:
stats[p] = stats[p] + 1
else:
stats[p] = 1
else:
print("combination does not exists")
for k in stats:
df[k][df.DrawDate == draw_date] = stats[k]
# for index, row in df[df.DrawDate == '1992-05-06 00:00:00'].iterrows():
# pairings = get_pairings(df, pad(row.LuckyNo))
# for p in pairings:
# if p in df.columns:
# if p in stats:
# stats[p] = stats[p] + 1
# else:
# stats[p] = 1
# else:
# print("combination does not exists")
# for k in stats:
# df[k][df.DrawDate == '1992-05-06 00:00:00'] = stats[k]
# df[df.DrawDate == '1992-05-06 00:00:00'][["DrawDate","LuckyNo"] + new_columns]
df[["DrawDate","LuckyNo"] + new_columns].head(66)
# Total 55 pairing combinations
len(new_columns)
```
#### Digits Frequencies
```
columns = ['DrawDate', '1st_digit', '2nd_digit', '3rd_digit', '4th_digit']
df[columns].head(23)
def get_digit_count(df, fld, digit, draw_date)->int:
"""Get digit count for a particular position"""
return len(df[ (df[fld]== digit) & (df.DrawDate == draw_date)])
# Derive new columns
prefix = 'pos_{}_{}_freq'
new_columns=[prefix.format(pos, digit) for pos in range(1, 5) for digit in range(0,10)]
print(new_columns)
def get_draw_digit_freq(df, draw_date, stats):
"""Get digits freqs per draw"""
first_freq = second_freq = third_freq = forth_freq = 0
for digit in range(0, 10):
stats[prefix.format(1, digit)] = stats[prefix.format(1, digit)] + get_digit_count(df, "1st_digit", digit, draw_date)
stats[prefix.format(2, digit)] = stats[prefix.format(1, digit)] + get_digit_count(df, "2nd_digit", digit, draw_date)
stats[prefix.format(3, digit)] = stats[prefix.format(1, digit)] + get_digit_count(df, "3rd_digit", digit, draw_date)
stats[prefix.format(4, digit)] = stats[prefix.format(1, digit)] + get_digit_count(df, "4th_digit", digit, draw_date)
stats = {}
for c in new_columns:
df[c] = 0
stats[c] = 0
# for pos in range(1,5):
# for digit in range(0, 10):
# stats[prefix.format(pos, digit)] = 0
# print(stats)
# draw_date = '1992-05-06 00:00:00'
#get_draw_digit_freq(df, draw_date, stats)
#print(stats)
# display(df[df.DrawDate == draw_date][["LuckyNo"] + new_columns])
count = 0
for draw_date in df.DrawDate.unique():
if count % 500 == 0:
print(f"Working on {draw_date}")
count = count + 1
get_draw_digit_freq(df, draw_date, stats)
for k in stats:
df[k][df.DrawDate == draw_date] = stats[k]
display(df.head(46))
len(df.columns)
```
#### Save to file
```
df.to_csv(DATASET_PROCESSED, index=False, header=True)
```
#### Averaging gaps between dates
#### Visualisation
#### Digits Rolling Mean
```
# df['1st_digit_rolling_mean'] = df['1st_digit'].rolling(window=23).mean()
# df['2st_digit_rolling_mean'] = df['2st_digit'].rolling(window=23).mean()
# df['3st_digit_rolling_mean'] = df['3st_digit'].rolling(window=23).mean()
# df['4st_digit_rolling_mean'] = df['4st_digit'].rolling(window=23).mean()
# df['total_digits_rolling_mean'] = (df['1st_digit_rolling_mean'] + df['2st_digit_rolling_mean'] + df['3st_digit_rolling_mean'] + df['4st_digit_rolling_mean']) / 4
# df.head(46)
```
| github_jupyter |
```
%load_ext lab_black
# from __future__ import print_function
import bs4 as bs # import BeautifulSoup as bs
import re
import requests
import pandas as pd
import json
from datetime import date
import os
import random
import time
# from IPython.display import display
from six.moves.urllib import parse
def news_parser(news_url):
"""parse news from HTML to a readable article
"""
print("Parsing news at\n %s" % news_url)
news_id = re.search(pattern="\d{7}", string=news_url).group(0)
news_title = (news_url.split("/"))[-1]
news_page = requests.get(news_url)
news_page.encoding = "utf-8"
news = bs.BeautifulSoup(news_page.text, "lxml")
article = [""] # pseudo initialization, in the case some news cannot be parsed
# step1: extract all HTML strings containing 'APOLLO_STATE'
APOLLO_string = []
scripts = news.find_all("script")
for script in scripts:
sString = script.string
if sString != None and sString.find("APOLLO_STATE") > 0:
APOLLO_string.append(sString)
# step2: extract contents from {}
if APOLLO_string != []:
regBrackets = r"\{(.*?)\}"
matches = re.finditer(regBrackets, APOLLO_string[0], re.MULTILINE | re.DOTALL)
bracket_content = []
# bracket_content is the content between {}
for match in matches:
bracket_content.append(match.group(1))
# step3: match sentence
article_end_index = bracket_content.index(
'"type":"reading-100-percent-completion-tracker"'
)
sentence = []
# sentence[] contains the sentence behind "type":"text","data":, and that's the article sentence
sen_code = '(?<="type":"p","children":\[{"type":"text","data":).*$'
for content in bracket_content[0 : article_end_index + 1]:
sen = re.findall(sen_code, content)
if not sen == [] and not sen == ['"\\n"']: # clean blank lists and \\n
sentence += sen
# step4: clean sentences and combine them into an article
for i in range(len(sentence)):
sentence[i] = sentence[i].strip('"')
article = ""
for s in sentence:
article += s
# change: new sentence has a space ahead
article += " "
article = [article]
# record date and build a dataframe
news_time = news.find("time")
if news_time == None:
news_time = ["cannot parse"]
else:
news_time = news_time["datetime"]
now = date.today()
news_df = pd.DataFrame(
data={
"news_id": news_id,
"title": news_title,
"article": article,
"release_time": news_time,
"collecting_date": now,
"URL": news_url,
}
)
time.sleep(random.random() * 5)
return news_df
def comments_parser(comments_url):
"""parse comments and return a dataframe
"""
print("Parsing comments at\n %s" % comments_url)
comments_page = requests.get(comments_url)
comments_page.encoding = "utf-8"
comments = bs.BeautifulSoup(comments_page.text, "lxml")
words = comments.find_all(class_="card-content-action")
# the very raw comments, every "card-content-action" is a person's comment block.
user_name = []
sentence = []
comment_time = []
for word in words: # every word is a comment block
user_name.append(word.find(attrs={"class": "comment-author-name"}).text)
sentence.append(
word.find(attrs={"class": "comment-content"})
.text.strip()
.replace("\n", " ")
)
comment_time.append((word.find("time"))["datetime"])
sentence_cleaned = []
for s in sentence:
if "@******" in s:
sentence_cleaned.append((s.split("@******"))[1].strip())
else:
sentence_cleaned.append(s)
news_id = comments_url[-7:]
now = date.today()
comments_df = pd.DataFrame(
data={
"user_name": user_name,
"comment_raw": sentence,
"comment_cleaned": sentence_cleaned,
"date": comment_time,
"news_id": news_id,
"is_reply": True,
"collecting_date": now,
}
)
for index, row in comments_df.iterrows():
if "@******" in row["comment_raw"]:
comments_df.at[index, "is_reply"] = True
else:
comments_df.at[index, "is_reply"] = False
time.sleep(random.random() * 3)
return comments_df
```
## Collect all URLs of the articles
**Method**
Since SCMP uses infinite roll to load old articles, I catch the URL of every request and find its pattern. Some short pages use 0,20,40,60 as their identifier, while other pages may use timestamps. For those timestamps, I collect them manually.
I've also tried to download rolled down pages as mHTML, but bs.find_all("a", href=True) or bs.find_all(class_=) doesn't work as in online pages.
```
url_list = []
```
https://www.scmp.com/coronavirus/greater-china
```
for i in range(4):
url = "https://apigw.scmp.com/content-delivery/v1?operationName=QueueById&variables=%7B%22itemLimit%22%3A20%2C%22offset%22%3A{offset}%2C%22name%22%3A%22section_top_505356%22%7D&extensions=%7B%22persistedQuery%22%3A%7B%22version%22%3A1%2C%22sha256Hash%22%3A%223f30ddb476061fe29f0dba291c35330b61a58fb0aaf08096d985f6b81e107840%22%7D%7D".format(
offset=i * 20
)
headers = {"apikey": "MyYvyg8M9RTaevVlcIRhN5yRIqqVssNY"}
text = requests.get(url=url, headers=headers).text
result = json.loads(text)["data"]["queue"]["items"]
for item in result:
url_list.append(item["urlAlias"])
# print(item["urlAlias"]) # item["headline"],
```
https://www.scmp.com/coronavirus/asia
```
# 0-60,step=20
for i in range(4):
url = "https://apigw.scmp.com/content-delivery/v1?operationName=QueueById&variables=%7B%22itemLimit%22%3A20%2C%22offset%22%3A{offset}%2C%22name%22%3A%22section_top_505326%22%7D&extensions=%7B%22persistedQuery%22%3A%7B%22version%22%3A1%2C%22sha256Hash%22%3A%223f30ddb476061fe29f0dba291c35330b61a58fb0aaf08096d985f6b81e107840%22%7D%7D".format(
offset=i * 20
)
headers = {"apikey": "MyYvyg8M9RTaevVlcIRhN5yRIqqVssNY"}
text = requests.get(url=url, headers=headers).text
result = json.loads(text)["data"]["queue"]["items"]
for item in result:
url_list.append(item["urlAlias"])
# print(item["urlAlias"]) # item["headline"],
```
https://www.scmp.com/coronavirus/europe
```
# 0-60,step=20
for i in range(4):
url = "https://apigw.scmp.com/content-delivery/v1?operationName=QueueById&variables=%7B%22itemLimit%22%3A20%2C%22offset%22%3A{offset}%2C%22name%22%3A%22section_top_505325%22%7D&extensions=%7B%22persistedQuery%22%3A%7B%22version%22%3A1%2C%22sha256Hash%22%3A%223f30ddb476061fe29f0dba291c35330b61a58fb0aaf08096d985f6b81e107840%22%7D%7D".format(
offset=i * 20
)
headers = {"apikey": "MyYvyg8M9RTaevVlcIRhN5yRIqqVssNY"}
text = requests.get(url=url, headers=headers).text
result = json.loads(text)["data"]["queue"]["items"]
for item in result:
url_list.append(item["urlAlias"])
# print(item["urlAlias"]) # item["headline"],
```
https://www.scmp.com/coronavirus/us-canada
```
# 0-60,step=20
for i in range(4):
url = "https://apigw.scmp.com/content-delivery/v1?operationName=QueueById&variables=%7B%22itemLimit%22%3A20%2C%22offset%22%3A{offset}%2C%22name%22%3A%22section_top_505354%22%7D&extensions=%7B%22persistedQuery%22%3A%7B%22version%22%3A1%2C%22sha256Hash%22%3A%223f30ddb476061fe29f0dba291c35330b61a58fb0aaf08096d985f6b81e107840%22%7D%7D".format(
offset=i * 20
)
headers = {"apikey": "MyYvyg8M9RTaevVlcIRhN5yRIqqVssNY"}
text = requests.get(url=url, headers=headers).text
result = json.loads(text)["data"]["queue"]["items"]
for item in result:
url_list.append(item["urlAlias"])
# print(item["urlAlias"]) # item["headline"],
```
https://www.scmp.com/coronavirus/health-medicine
```
# 0-60,step=20
for i in range(4):
url = "https://apigw.scmp.com/content-delivery/v1?operationName=QueueById&variables=%7B%22itemLimit%22%3A20%2C%22offset%22%3A{offset}%2C%22name%22%3A%22section_top_505355%22%7D&extensions=%7B%22persistedQuery%22%3A%7B%22version%22%3A1%2C%22sha256Hash%22%3A%223f30ddb476061fe29f0dba291c35330b61a58fb0aaf08096d985f6b81e107840%22%7D%7D".format(
offset=i * 20
)
headers = {"apikey": "MyYvyg8M9RTaevVlcIRhN5yRIqqVssNY"}
text = requests.get(url=url, headers=headers).text
result = json.loads(text)["data"]["queue"]["items"]
for item in result:
url_list.append(item["urlAlias"])
# print(item["urlAlias"]) # item["headline"],
```
https://www.scmp.com/coronavirus/economic-impact
```
# 0-60,step=20
for i in range(4):
url = "https://apigw.scmp.com/content-delivery/v1?operationName=QueueById&variables=%7B%22itemLimit%22%3A20%2C%22offset%22%3A{offset}%2C%22name%22%3A%22section_top_505328%22%7D&extensions=%7B%22persistedQuery%22%3A%7B%22version%22%3A1%2C%22sha256Hash%22%3A%223f30ddb476061fe29f0dba291c35330b61a58fb0aaf08096d985f6b81e107840%22%7D%7D".format(
offset=i * 20
)
headers = {"apikey": "MyYvyg8M9RTaevVlcIRhN5yRIqqVssNY"}
text = requests.get(url=url, headers=headers).text
result = json.loads(text)["data"]["queue"]["items"]
for item in result:
url_list.append(item["urlAlias"])
# print(item["urlAlias"]) # item["headline"],
```
https://www.scmp.com/coronavirus/analysis-opinion
```
# 0-60,step=20
for i in range(4):
url = "https://apigw.scmp.com/content-delivery/v1?operationName=QueueById&variables=%7B%22itemLimit%22%3A20%2C%22offset%22%3A{offset}%2C%22name%22%3A%22section_top_505327%22%7D&extensions=%7B%22persistedQuery%22%3A%7B%22version%22%3A1%2C%22sha256Hash%22%3A%223f30ddb476061fe29f0dba291c35330b61a58fb0aaf08096d985f6b81e107840%22%7D%7D".format(
offset=i * 20
)
headers = {"apikey": "MyYvyg8M9RTaevVlcIRhN5yRIqqVssNY"}
text = requests.get(url=url, headers=headers).text
result = json.loads(text)["data"]["queue"]["items"]
for item in result:
url_list.append(item["urlAlias"])
# print(item["urlAlias"]) # item["headline"],
```
https://www.scmp.com/topics/coronavirus-china
```
timestamp_list_COVCHINA = [
"1587436199000",
"1587117610000",
"1586913303000",
"1586610083000",
"1586361914000",
"1585954821000",
"1585735387000",
"1585528208000",
"1585229419000",
"1584874812000",
"1584581236000",
"1581608419000",
"1589540408000",
"1589274011000",
"1588939218000",
"1588673798000",
"1588345217000",
"1588063310000",
"1587770106000",
"1587558821000",
"1587295804000",
"1587036700000",
"1586842439000",
"1586527218000",
"1586317395000",
"1585896342000",
"1585400413000",
"1585132217000",
"1584768613000",
"1584493217000",
"1579925153000", # back to Mar 19, the earliest
]
timestamp_list_COVCHINA.sort()
timestamp_list_COVCHINA
for timestamp in timestamp_list_COVCHINA:
url = (
"https://apigw.scmp.com/content-delivery/v1?operationName=gettopicbyentityuuid&variables=%7B%22latestContentsLimit%22%3A30%2C%22latestOpinionsLimit%22%3A30%2C%22entityUuid%22%3A%22c7985da3-2f6d-4540-a1c2-875c2d7881b6%22%2C%22articleTypeId%22%3A%22012d7708-2959-4b2b-9031-23e3d025a08d%22%2C%22applicationIds%22%3A%5B%222695b2c9-96ef-4fe4-96f8-ba20d0a020b3%22%5D%2C%22after%22%3A%22"
+ timestamp
+ "%22%7D&extensions=%7B%22persistedQuery%22%3A%7B%22version%22%3A1%2C%22sha256Hash%22%3A%22a78c49ca1280c93d31533f172e1a837d7e8aebf1215195364cb7fd85c76d2e0e%22%7D%7D"
)
headers = {"apikey": "MyYvyg8M9RTaevVlcIRhN5yRIqqVssNY"}
r = requests.get(url=url, headers=headers)
j = r.json()["data"]["topic"]["latestContentsWithCursor"]["items"]
for item in j:
url_list.append(item["urlAlias"])
# print(item["urlAlias"])
```
## Append new urls and save them in a csv
```
if not os.path.isfile("news_url.csv"):
# creat the news url list for the first time
print("New file created.")
news_url_list = []
for item in url_list:
news_url_list.append("https://www.scmp.com" + item)
news_url_list = list(dict.fromkeys(news_url_list)) # rid duplicates
news_url_df = pd.DataFrame(data={"news_url": news_url_list})
news_url_df.to_csv(
"../data/news_url.csv", encoding="utf-8-sig", header="column_names", index=False
)
else:
# there is already a news url list file
print("File already exist.")
news_url_df = pd.read_csv("news_url.csv")
news_url_list = list(news_url_df["news_url"])
for item in url_list:
news_url_list.append("https://www.scmp.com" + item)
news_url_list = list(dict.fromkeys(news_url_list)) # rid duplicates
news_url_df = pd.DataFrame(data={"news_url": news_url_list})
news_url_df.to_csv(
"../data/news_url.csv",
encoding="utf-8-sig",
mode="w",
header="column_names",
index=False,
)
```
## Test
```
# read news url list from csv
news_url_df = pd.read_csv("../data/news_url.csv")
news_url_list = list(news_url_df["news_url"].unique())
print("We have collected %d news URLs in all." % len(list(news_url_df["news_url"])))
```
After 386 loops, the connection breaks.
```
counter = 0
# news_url_list is the list of all news URLs
for url in news_url_list[1174:]:
re_try = re.search(pattern="\d{7}", string=url)
if re_try != None:
news_id = re_try.group(0)
news_df = news_parser(url) # parse news article
comments_id = news_id
comments_url = "https://www.scmp.com/scmp_comments/popup/" + comments_id
comments_df = comments_parser(comments_url) # parse comments
if not news_df["article"][0] == [""]:
news_df.to_csv(
"../data/news_new.csv",
encoding="utf-8-sig",
header=False,
index=False,
mode="a",
)
if not comments_df["user_name"][0] == None:
comments_df.to_csv(
"../data/comments_new.csv",
encoding="utf-8-sig",
header=False,
index=False,
mode="a",
)
counter += 1
print("Loop %d finished." % counter)
```
| github_jupyter |
# Training and Deploying A Deep Learning Model in Keras MobileNet V2 and Heroku: A Step-by-Step Tutorial
**This is part of my blog [here](https://medium.com/@malnakli/tf-serving-keras-mobilenetv2-632b8d92983c)**
## Prepare data for training
|Label|Description|
|- | - |
| 0 | T-shirt/top |
| 1 | Trouser |
| 2 | Pullover |
| 3 | Dress |
| 4 | Coat |
| 5 | Sandal |
| 6 | Shirt |
| 7 | Sneaker |
| 8 | Bag |
| 9 | Ankle boot |
```
from keras.datasets.fashion_mnist import load_data
# Load the fashion-mnist train data and test data
(x_train, y_train), (x_test, y_test) = load_data()
print("x_train shape:", x_train.shape, "y_train shape:", y_train.shape)
```
## Helper function to display imags
```
import matplotlib.pyplot as plt
import numpy as np
%matplotlib inline
def show_images(images):
"""
images : numpy arrays
"""
n_images = len(images)
titles = ['(%d)' % i for i in range(1, n_images + 1)]
num = 5
iter_num = np.ceil(n_images / num).astype(int)
for i in range(iter_num):
fig = plt.figure()
sub_images = images[i * num:num * (i + 1)]
sub_titles = titles[i * num:num * (i + 1)]
for n, (image, title) in enumerate(zip(sub_images, sub_titles)):
a = fig.add_subplot(1, np.ceil(len(sub_images)), n + 1)
if image.ndim == 2:
plt.gray()
a.set_title(title, fontsize=15)
plt.imshow(image)
```
### Show samples of data
```
show_images(x_train[:10])
```
### Data normalization
Normalize the data dimensions so that they are of approximately the same scale.In general, normalization makes very deep NN easier to train, special in Convolutional and Recurrent neural network.
Here is a nice explanation [video](https://www.coursera.org/lecture/deep-neural-network/normalizing-activations-in-a-network-4ptp2) and an [article](https://medium.com/@darrenyaoyao.huang/why-we-need-normalization-in-deep-learning-from-batch-normalization-to-group-normalization-d06ea0e59c17)
```
norm_x_train = x_train.astype('float32') / 255
norm_x_test = x_test.astype('float32') / 255
# dsiplay images
show_images(norm_x_train[:10])
```
### Convert labels (y_train and y_test) to one hot encoding
A one hot encoding is a representation of categorical variables as binary vectors.
[Here is the full explanation](https://machinelearningmastery.com/how-to-one-hot-encode-sequence-data-in-python/) If you would like to have a deep understanding.
```
from keras.utils import to_categorical
encoded_y_train = to_categorical(y_train, num_classes=10, dtype='float32')
encoded_y_test = to_categorical(y_test, num_classes=10, dtype='float32')
```
### Resize images & convert to 3 channel (RGB)
[MobileNet V2](https://keras.io/applications/#mobilenetv2) model accepts one of the following formats: (96, 96), (128, 128), (160, 160),(192, 192), or (224, 224). In addition, the image has to be 3 channel (RGB) format. Therefore, We need to resize & convert our images. from (28 X 28) to (96 X 96 X 3).
```
from skimage.transform import resize
target_size = 96
def preprocess_image(x):
# Resize the image to have the shape of (96,96)
x = resize(x, (target_size, target_size),
mode='constant',
anti_aliasing=False)
# convert to 3 channel (RGB)
x = np.stack((x,)*3, axis=-1)
# Make sure it is a float32, here is why
# https://www.quora.com/When-should-I-use-tf-float32-vs-tf-float64-in-TensorFlow
return x.astype(np.float32)
```
Running the previous code in all our data data, it may eat up a lot of memory resources; therefore, we are going to use generator.
[Python Generator](https://www.programiz.com/python-programming/generator) is a function that returns an object (iterator) which we can iterate over (one value at a time).
```
from sklearn.utils import shuffle
def load_data_generator(x, y, batch_size=64):
num_samples = x.shape[0]
while 1: # Loop forever so the generator never terminates
try:
shuffle(x)
for i in range(0, num_samples, batch_size):
x_data = [preprocess_image(im) for im in x[i:i+batch_size]]
y_data = y[i:i + batch_size]
# convert to numpy array since this what keras required
yield shuffle(np.array(x_data), np.array(y_data))
except Exception as err:
print(err)
```
## Train a Deep Learning model
```
from keras.applications.mobilenetv2 import MobileNetV2
from keras.layers import Dense, Input, Dropout
from keras.models import Model
def build_model( ):
input_tensor = Input(shape=(target_size, target_size, 3))
base_model = MobileNetV2(
include_top=False,
weights='imagenet',
input_tensor=input_tensor,
input_shape=(target_size, target_size, 3),
pooling='avg')
for layer in base_model.layers:
layer.trainable = True # trainable has to be false in order to freeze the layers
op = Dense(256, activation='relu')(base_model.output)
op = Dropout(.25)(op)
##
# softmax: calculates a probability for every possible class.
#
# activation='softmax': return the highest probability;
# for example, if 'Coat' is the highest probability then the result would be
# something like [0,0,0,0,1,0,0,0,0,0] with 1 in index 5 indicate 'Coat' in our case.
##
output_tensor = Dense(10, activation='softmax')(op)
model = Model(inputs=input_tensor, outputs=output_tensor)
return model
from keras.optimizers import Adam
model = build_model()
model.compile(optimizer=Adam(),
loss='categorical_crossentropy',
metrics=['categorical_accuracy'])
train_generator = load_data_generator(norm_x_train, encoded_y_train, batch_size=64)
model.fit_generator(
generator=train_generator,
steps_per_epoch=5,
verbose=1,
epochs=2)
```
### Test
```
test_generator = load_data_generator(norm_x_test, encoded_y_test, batch_size=64)
model.evaluate_generator(generator=test_generator,
steps=900,
verbose=1)
```
## Save the model
Make sure you save the model, because we are going to use in next part
```
model_name = "tf_serving_keras_mobilenetv2"
model.save(f"models/{model_name}.h5")
```
## Part 2 (if you follow the blog)
## Makes Model ready to tensorflow serving
### Tensorflow serving
[TensorFlow Serving](https://www.tensorflow.org/serving/overview) is a flexible, high-performance serving system for machine learning models, designed for **production** environments.
```
from keras.models import load_model
model = load_model(f"models/{model_name}.h5")
```
### Build & Save model to be tensorflow serving ready
```
import os
import tensorflow as tf
import keras
# Import the libraries needed for saving models
# Note that in some other tutorials these are framed as coming from tensorflow_serving_api which is no longer correct
from tensorflow.python.saved_model import builder as saved_model_builder
from tensorflow.python.saved_model import tag_constants, signature_constants, signature_def_utils_impl
# images will be the input key name
# scores will be the out key name
prediction_signature = tf.saved_model.signature_def_utils.predict_signature_def(
{
"images": model.input
}, {
"scores": model.output
})
# export_path is a directory in which the model will be created
export_path = os.path.join(
tf.compat.as_bytes('models/export/{}'.format(model_name)),
tf.compat.as_bytes('1'))
builder = saved_model_builder.SavedModelBuilder(export_path)
sess = keras.backend.get_session()
# Add the meta_graph and the variables to the builder
builder.add_meta_graph_and_variables(
sess, [tag_constants.SERVING],
signature_def_map={
'prediction': prediction_signature,
})
# save the graph
builder.save()
```
## Setting up Heroku & Docker
- [Install docker for macOS and windows](https://www.docker.com/products/docker-desktop)
- A little more work for Ubuntu users but still straightforward [Install docker for Ubuntu](https://docs.docker.com/install/linux/docker-ce/ubuntu/#upgrade-docker-ce)
- [Signup to Heroku](https://signup.heroku.com/)
- [Install heroku-cli](https://devcenter.heroku.com/articles/heroku-cli#download-and-install)
### After you have installed docker and heroku-cli.
Run the following to make sure docker & heroku have been installed correctly
```
> docker ps
CONTAINER ID IMAGE COMMAND CREATED STATUS PORTS NAMES
> heroku --version
heroku/7.18.3 darwin-x64 node-v10.12.0
# make sure you have logged in to your heroku account
heroku login
# Output should have:
Logged in as xxxxx@xxx.xx
```
## Deploy the model to Heroku
#### Download tensorflow serving image from `hub.docker.com`
Because tensorflow serving docker image was not optimized for heroku.
I have created a dockerfile that following heroku instructure. cleck [here](https://github.com/malnakli/ML/tf_serving_keras_mobilenetv2/Dockerfile) to look at.
Also I have pushed the a docker image that ready to deploy to heroku,
which already have the trained model.
`docker pull malnakli/ml:tf-serving-heroku-1.11`
**To build you own**
Run the following:
make sure you ar in the right folder
`cd ML/tf_serving_keras_mobilenetv2`
Build docker image:
`docker build -t tf-serving-heroku-1.11 .`
Once the image build. you can run it locally if you would like,
otherwise go to deploy section.
`docker run -p 8501:8501 -e PORT=8501 -t tf-serving-heroku-1.11`
If you see the following the last output, then it works.
```
... tensorflow_serving/model_servers/server.cc:301] Exporting HTTP/REST API at:localhost:8501 ...
```
### Deploy
##### Log in to Container Registry:
`heroku container:login`
##### Create a heroku app
`heroku create ${YOUR_APP_NAME}`
#### Push the docker image to heroku
`heroku container:push web -a ${YOUR_APP_NAME}`
`heroku container:release web -a ${YOUR_APP_NAME}`
## Call the model
#### The loacl url
> `http://localhost:8501//v1/models/tf_serving_keras_mobilenetv2/versions/1:predict`
#### The Heroku url
> http://https://tf-serving-keras-mobilenetv2.herokuapp.com//v1/models/tf_serving_keras_mobilenetv2/versions/1:predict
JSON data that is sent to TensorFlow Model Server:
```
{
"signature_name":'prediction',
"instances": [{"images":image.tolist()}]
}
```
And you can see the full documentation about RESTful API in [Here](https://www.tensorflow.org/serving/api_rest).
```
from urllib import request
from PIL import Image
image_url = "https://cdn.shopify.com/s/files/1/2029/4253/products/Damb_Back_2a3cc4cc-06c2-488e-8918-2e7a1cde3dfc_530x@2x.jpg"
image_path = f"tmp/{image_url.split('/')[-1]}"
# download image
with request.urlopen(url=image_url, timeout=10) as response:
data = response.read()
with open(image_path, 'wb') as f:
f.write(data)
# convert image to grayscale.
image = Image.open(image_path).convert('L')
# resize the image to 28 28 to make sure it is similar to our dataset
image.thumbnail((28,28))
image = preprocess_image(np.array(image))
print(image.shape)
show_images([image])
import requests
import json
"""
NOTE:
change https://tf-serving-keras-mobilenetv2.herokuapp.com to your url or
if you ran the docker locally, then replace with http://localhost:8501
"""
url = "https://tf-serving-keras-mobilenetv2.herokuapp.com"
full_url = f"{url}/v1/models/tf_serving_keras_mobilenetv2/versions/1:predict"
data = {"signature_name":"prediction",
"instances":[{"images":image.tolist()}]}
data = json.dumps(data)
import sys
labels = ['T-shirt/top' ,'Trouser','Pullover','Dress','Coat','Sandal','Shirt','Sneaker','Bag', 'Ankle boot']
try:
response = requests.post(full_url,data=data)
response = response.json()
highest_index = np.argmax(response['predictions'])
print(labels[highest_index])
except:
print(sys.exc_info()[0])
```
**If you have any suggest or question you can create an issue in this repo**
# References:
Here are some articles that it convert similar aspect cover in this notebook.
- [Fashion-MNIST with tf.Keras](https://medium.com/tensorflow/hello-deep-learning-fashion-mnist-with-keras-50fcff8cd74a)
- [A Comprehensive guide to Fine-tuning Deep Learning Models in Keras ](https://flyyufelix.github.io/2016/10/03/fine-tuning-in-keras-part1.html)
- [Transfer Learning and Fine Tuning: Let's discuss](https://www.linkedin.com/pulse/transfer-learning-fine-tuning-lets-discuss-arun-das/)
- [Serving Image-Based Deep Learning Models with TensorFlow-Serving’s RESTful API](https://medium.com/@tmlabonte/serving-image-based-deep-learning-models-with-tensorflow-servings-restful-api-d365c16a7dc4)
- [How to Setup Tensorflow Serving For Production](https://medium.com/@brianalois/how-to-setup-tensorflow-serving-for-production-3cc2abf7efa)
- [How to Run Dockerized Apps on Heroku… and it’s pretty sweet](https://medium.com/travis-on-docker/how-to-run-dockerized-apps-on-heroku-and-its-pretty-great-76e07e610e22)
| github_jupyter |
# Convergence Acceleration
This notebook shows crude implementations of some convergence acceleration techniques. We are testing on a very simple case: Jacobi iterations for the solution of an SPD linear system.
```
import numpy as np
from numpy import linalg as LA
import itertools
from collections import OrderedDict
from bokeh.io import push_notebook, show, output_notebook
from bokeh.layouts import row
from bokeh.plotting import figure
from bokeh.palettes import Dark2_5 as palette
output_notebook()
```
## Jacobi iterations
The next function performs standad Jacobi iterations to solve a linear system with matrix $\mathbf{A}$ and right-hand side vector $\mathbf{b}$.
Let us split the stiffness matrix into its diagonal and off-diagonal components:
\begin{equation}
\mathbf{A} = \mathbf{O} + \mathbf{D}
\end{equation}
which leads to:
\begin{equation}
\mathbf{A}\mathbf{x} = (\mathbf{O} + \mathbf{D})\mathbf{x} = \mathbf{b},
\end{equation}
then moving the vector $\mathbf{O}\mathbf{x}$ to the right-hand side gives us the, formally trivial, diagonal system:
\begin{equation}
\mathbf{D}\mathbf{x} = \mathbf{b} - \mathbf{O}\mathbf{x},
\end{equation}
whose solution yields the form of the fixed-point iterations we seek:
\begin{equation}
\mathbf{x}^{[n+1]} = \mathbf{D}^{-1}(\mathbf{b} - \mathbf{O}\mathbf{x}^{[n]})
\end{equation}
```
def apply_jacobi(A, b, x):
"""
Compute next solution of linear system by applying a Jacobi step.
Parameters
----------
A
Stiffness matrix
b
RHS vector
x
Current solution
Returns
-------
Next iteratate in Jacobi fixed-point iteration sequence
"""
D = np.diag(A)
O = A - np.diagflat(D)
return (b - np.einsum('ij,j->i', O, x)) / D
def jacobi(A, b, rtol=1.0e-8, max_it=50, x_0=None, print_report=False):
"""
Solve linear system of equations with Jacobi iterations.
Parameters
----------
A
Stiffness matrix
b
RHS vector
rtol
Tolerance for solution norm
max_it
Maximum number of iterations
x_0
Initial guess
print_report
Whether to print iteration report inside the function
Returns
-------
x
Linear system solution vector
n_it
Number of iterations performed
report
Iteration number, residual norm sequence
"""
x = np.zeros_like(b) if x_0 is None else x_0
it = 0
report = []
if print_report:
print('Iteration # Residual norm')
while it < max_it:
# Compute residual
r = apply_jacobi(A, b, x) - x
rnorm = LA.norm(r)
# Report
report.append(rnorm)
if print_report:
print(' {:4d} {:.5E}'.format(it, rnorm))
# Check convergence
if rnorm < rtol:
break
# Update solution vector
x = apply_jacobi(A, b, x)
it += 1
else:
raise RuntimeError(
'Maximum number of iterations ({0:d}) exceeded, but residual norm {1:.5E} still greater than threshold {2:.5E}'.
format(max_it, rnorm, rtol))
return x, it, report
```
## Conjugate gradient and conjugate residual
Assuming that the stiffness matrix is SPD, the conjugate gradient (CG) method would be a more robust choice of iterative method than Jacobi iterations. If the stiffness matrix is only known to be Hermitian, the conjugate residual (CR) method can be used instead.
```
def cg(A, b, rtol=1.0e-8, max_it=25, x_0=None, print_report=False):
"""
Solve linear system of equations with conjugate gradient.
Parameters
----------
A
Stiffness matrix
b
RHS vector
rtol
Tolerance for solution norm
max_it
Maximum number of iterations
x_0
Initial guess
print_report
Whether to print iteration report inside the function
Returns
-------
x
Linear system solution vector
n_it
Number of iterations performed
report
Iteration number, residual norm sequence
"""
x = np.zeros_like(b) if x_0 is None else x_0
it = 0
report = []
r = b - np.einsum('ij,j->i', A, x)
p = r
if print_report:
print('Iteration # Residual norm')
while it < max_it:
Ap = np.einsum('ij,j->i', A, p)
rtr = np.einsum('i,i', r, r)
alpha = rtr / np.einsum('i,i', p, Ap)
x = x + alpha * p
r = r - alpha * Ap
rnorm = LA.norm(r)
# Report
report.append(rnorm)
if print_report:
print(' {:4d} {:.5E}'.format(it, rnorm))
# Check convergence
if rnorm < rtol:
break
beta = np.einsum('i,i', r, r) / rtr
p = r + beta * p
it += 1
else:
raise RuntimeError(
'Maximum number of iterations ({0:d}) exceeded, but residual norm {1:.5E} still greater than threshold {2:.5E}'.
format(max_it, rnorm, rtol))
return x, it, report
def cr(A, b, rtol=1.0e-8, max_it=25, x_0=None, print_report=False):
"""
Solve linear system of equations with conjugate residual.
Parameters
----------
A
Stiffness matrix
b
RHS vector
rtol
Tolerance for solution norm
max_it
Maximum number of iterations
x_0
Initial guess
print_report
Whether to print iteration report inside the function
Returns
-------
x
Linear system solution vector
n_it
Number of iterations performed
report
Iteration number, residual norm sequence
"""
x = np.zeros_like(b) if x_0 is None else x_0
it = 0
report = []
r = b - np.einsum('ij,j->i', A, x)
p = r
if print_report:
print('Iteration # Residual norm')
while it < max_it:
Ap = np.einsum('ij,j->i', A, p)
rtAr = np.einsum('i,ij,j', r, A, r)
alpha = rtAr / np.einsum('i,i', Ap, Ap)
x = x + alpha * p
r = r - alpha * Ap
rnorm = LA.norm(r)
# Report
report.append(rnorm)
if print_report:
print(' {:4d} {:.5E}'.format(it, rnorm))
# Check convergence
if rnorm < rtol:
break
beta = np.einsum('i,ij,j', r, A, r) / rtAr
p = r + beta * p
it += 1
else:
raise RuntimeError(
'Maximum number of iterations ({0:d}) exceeded, but residual norm {1:.5E} still greater than threshold {2:.5E}'.
format(max_it, rnorm, rtol))
return x, it, report
```
## Kaczmarz iterations
\begin{equation}
\mathbf{x}^{[k+1]} = \mathbf{x}^{[k]} + \frac{b_i - \mathbf{A}_{i}\cdot\mathbf{x}^{[k]}}{||\mathbf{A}_{i}||^{2}}\mathbf{A}_{i}^{t},
\end{equation}
with $i = k \mod m$.
```
def kaczmarz(A, b, rtol=1.0e-8, max_it=2005, x_0=None, print_report=False):
x = np.zeros_like(b) if x_0 is None else x_0
it = 0
report = []
r = b - np.einsum('ij,j->i', A, x)
# Number of rows
m = A.shape[0]
if print_report:
print('Iteration # Residual norm')
while it < max_it:
# Kaczmarz sweeps
i = it%m
alpha = (b[i] - np.einsum('i,i->', A[i,:], x)) / np.einsum('i,i->', A[i,:], A[i,:])
x = x + alpha * A[i,:].T
r = b - np.einsum('ij,j->i', A, x)
rnorm = LA.norm(r)
# Report
report.append(rnorm)
if print_report:
print(' {:4d} {:.5E}'.format(it, rnorm))
# Check convergence
if rnorm < rtol:
break
it +=1
else:
raise RuntimeError(
'Maximum number of iterations ({0:d}) exceeded, but residual norm {1:.5E} still greater than threshold {2:.5E}'.
format(max_it, rnorm, rtol))
return x, it, report
```
## DIIS acceleration
Direct inversion in the iterative subpace (DIIS) is an acceleration method ubiquitous in quantum chemistry. Given a queue of iterates $\lbrace \mathbf{x}^{[i]} \rbrace_{i=0}^{h}$, where $h$ is the maximum size of the queue, the next iterate in the sequence is obtained as a linear combination of the iterates in the queue:
\begin{equation}
\tilde{\mathbf{x}} := \sum_{i=0}^{h} c_{i} \mathbf{x}_{i}
\end{equation}
with the coefficients subject to the normalization condition:
\begin{equation}
\mathbf{1} \cdot \mathbf{c} = \sum_{i=0}^{h} c_{i} = 1.
\end{equation}
The optimal coefficients are found by minimizing the squared norm of the extrapolated iterate, under the normalization constraint. This leads to a linear system, sometimes known as _Pulay's equations_:
\begin{equation}
\begin{pmatrix}
\mathbf{x}_{0}\cdot\mathbf{x}_{0} & \cdots & \mathbf{x}_{0}\cdot\mathbf{x}_{n} & -1 \\
\vdots & \cdots & \vdots \\
\mathbf{x}_{n}\cdot\mathbf{x}_{0} & \cdots & \mathbf{x}_{n}\cdot\mathbf{x}_{n} & -1 \\
-1 & \cdots & -1 & 0 \\
\end{pmatrix}
\begin{pmatrix}
c_{0} \\
\vdots \\
c_{n} \\
\lambda \\
\end{pmatrix}
=
\begin{pmatrix}
0 \\
\vdots \\
0 \\
-1 \\
\end{pmatrix}
\end{equation}
```
def jacobi_diis(A, b, rtol=1.0e-8, max_it=25, max_hist=8, x_0=None, print_report=False):
"""
Solve linear system of equations with Jacobi iterations and DIIS acceleration.
Parameters
----------
A
Stiffness matrix
b
RHS vector
rtol
Tolerance for solution norm
max_it
Maximum number of iterations
max_hist
Maximum size of DIIS queue
x_0
Initial guess
print_report
Whether to print iteration report inside the function
Returns
-------
x
Linear system solution vector
n_it
Number of iterations performed
report
Iteration number, residual norm sequence
"""
x = np.zeros_like(b) if x_0 is None else x_0
# Lists of iterates and residuals
xs = []
rs = []
report = []
it = 0
if print_report:
print('Iteration # Residual norm')
while it < max_it:
# Compute residual
r = apply_jacobi(A, b, x) - x
rnorm = LA.norm(r)
# Collect DIIS history
xs.append(x)
rs.append(r)
# Report
report.append(rnorm)
if print_report:
print(' {:4d} {:.5E}'.format(it, rnorm))
# Check convergence
if rnorm < rtol:
break
if it >= 2:
# Prune DIIS history
diis_hist = len(xs)
if diis_hist > max_hist:
xs.pop(0)
rs.pop(0)
diis_hist -= 1
# Build error matrix B
B = np.empty((diis_hist + 1, diis_hist + 1))
B[-1, :] = -1
B[:, -1] = -1
B[-1, -1] = 0
for i, ri in enumerate(rs):
for j, rj in enumerate(rs):
if j > i: continue
val = np.einsum('i,i', ri, rj)
B[i, j] = val
B[j, i] = val
# Normalize B
B[:-1, :-1] /= np.abs(B[:-1, :-1]).max()
# Build rhs vector
rhs = np.zeros(diis_hist + 1)
rhs[-1] = -1
# Solve Pulay equations
cs = LA.solve(B, rhs)
# Calculate new solution as linear
# combination of previous solutions
x = np.zeros_like(x)
for i, c in enumerate(cs[:-1]):
x += c * xs[i]
# Update solution vector
x = apply_jacobi(A, b, x)
it += 1
else:
raise RuntimeError(
'Maximum number of iterations ({0:d}) exceeded, but residual norm {1:.5E} still greater than threshold {2:.5E}'.
format(max_it, rnorm, rtol))
return x, it, report
```
## KAIN acceleration
The Krylov accelerated inexact Newton (KAIN) method is another example of convergence acceleration algorithm similar to DIIS.
Given the nonlinear equation $\mathbf{\omega}(\mathbf{x}) = 0$, with $\mathbf{\omega}: \mathbb{R}^{N} \rightarrow \mathbb{R}^{N}$,
the Newton method formulates the solution as iterative updates $\mathbf{x}^{[n+1]} = \mathbf{x}^{[n]} + \mathbf{\delta}^{[n]}$ where $\mathbf{\delta}^{[n]}$ is of the form:
\begin{equation}
\mathbf{A}^{[n]}\mathbf{\delta}^{[n]} = -\mathbf{\omega}^{[n]},
\end{equation}
with $\mathbf{A}^{[n]}$ the Jacobian computed at the $n$-th iteration, _i.e._ from the $\mathbf{x}^{[n]}$ iterate.
We assume that we have generated a queue of iterates $\lbrace \mathbf{x}^{[i]}\rbrace_{i=0}^{h}$ and corresponding queue of function iterates $\lbrace \mathbf{\omega}^{[i]} \equiv \mathbf{\omega}(\mathbf{x}^{[i]})\rbrace_{i=0}^{h}$. As before, here $h$ is the maximum size of the queues.
Similarly to DIIS we seek to construct the update _without_ ever forming the full Jacobian. However, at variance with DIIS, we would like to include information from _outside_ the iterative subspace in our update.
An approximation of the Jacobian _within the subspace of iterates differences_ can be built as:
\begin{equation}
\omega^{[i]} = \omega(\mathbf{x}^{[i]} + \mathbf{x}^{[h]} - \mathbf{x}^{[h]})
\simeq
\omega^{[h]} + \mathbf{A}^{[h]}(\mathbf{x}^{[i]} - \mathbf{x}^{[h]})
\end{equation}
such that _within the subspace_ $\mathcal{D} = \lbrace \mathbf{y}^{[k]} | \mathbf{y}^{[k]} = \mathbf{x}^{[i]} - \mathbf{x}^{[h]},\, \forall k = 0,\ldots, h-1 \rbrace$:
\begin{equation}
\mathbf{A}^{[h]}(\mathbf{x}^{[i]} - \mathbf{x}^{[h]}) \simeq \omega^{[i]} - \omega^{[h]}.
\end{equation}
We introduce the projector into the $\mathcal{D}$ subspace $\hat{P}$ and its complement $\hat{Q}$ and we partition the Newton equation accordingly:
\begin{equation}
\mathbf{A}^{[h]}\hat{P}\delta^{[h]} + \mathbf{D}^{[h]}\hat{Q}\delta^{[h]} = - \omega^{[h]},
\end{equation}
where $\mathbf{D}^{[h]}$ is a _readily invertible approximation_ to the full Jacobian _outside_ the the $\mathcal{D}$ subspace.
Now, the update _within_ the subspace is a clearly a linear combination of differences of previous iterates:
\begin{equation}
\hat{P}\delta^{[h]} = \sum_{i=0}^{h-1} c_{i}(\mathbf{x}^{[i]} - \mathbf{x}^{[h]})
\end{equation}
with coefficients yet to be determined. The update _outside_ the subspace is then:
\begin{equation}
\hat{Q}\delta^{[h]} = - (\mathbf{D}^{[h]})^{-1}\left( \mathbf{A}^{[h]}\hat{P}\mathbf{\delta}^{[h]} + \omega^{[h]}\right).
\end{equation}
Projection onto the the $\mathcal{D}$ subspace and insertion of the equation for $\hat{P}\delta^{[h]}$ yields the subspace equations for the mixing coefficients $\mathbf{c}$:
\begin{equation}
\sum_{j=0}^{h-1}
\langle \mathbf{x}^{[i]} - \mathbf{x}^{[h]} | (\mathbf{D}^{[h]})^{-1}(\omega^{[j]} - \omega^{[h]}) \rangle c_{j}
= - \langle \mathbf{x}^{[i]} - \mathbf{x}^{[h]} | (\mathbf{D}^{[h]})^{-1}\omega^{[h]} \rangle
\end{equation}
The final form for the inexact Newton update is then:
\begin{equation}
\delta^{[h]} = \sum_{i=0}^{h-1} c_{i}(\mathbf{x}^{[i]} - \mathbf{x}^{[h]})
- (\mathbf{D}^{[h]})^{-1}\left(\sum_{i=0}^{h-1} c_{i}(\omega^{[i]} - \omega^{[h]}) + \omega^{[h]} \right)
\end{equation}
### The $\mathbf{Q}$ matrix
The KAIN subspace equations can be manipulated a bit further:
\begin{equation}
\sum_{j=0}^{h-1}
(Q^{[h]}_{ij} - Q^{[h]}_{ih} - Q^{[h]}_{hj} + Q^{[h]}_{hh})c_{j}
=
- Q^{[h]}_{ih} + Q^{[h]}_{hh}
\end{equation}
with the $\mathbf{Q}$ matrix being:
\begin{equation}
Q^{[h]}_{ij} = \langle \mathbf{x}^{[i]} | (\mathbf{D}^{[h]})^{-1}\omega^{[j]} \rangle
\end{equation}
Some further manipulation yields:
\begin{equation}
\sum_{j=0}^{h-1}
(Q^{[h]}_{ij} - Q^{[h]}_{hj})c_{j} + (Q^{[h]}_{hh} - Q^{[h]}_{ih})\left(\sum_{j=0}^{h-1} c_{j} \right)
=
- Q^{[h]}_{ih} + Q^{[h]}_{hh}.
\end{equation}
We now impose the normalization condition $\sum_{j=0}^{h} c_{j} = 1$ such that the subspace equation becomes:
\begin{equation}
\sum_{j=0}^{h}
(Q^{[h]}_{ij} - Q^{[h]}_{hj})c_{j}
=
0,
\end{equation}
while the update now reads:
\begin{equation}
\tilde{\mathbf{x}} = \sum_{j=0}^{h} c_{j}\mathbf{x}^{[j]} - (\mathbf{D}^{[h]})^{-1}\left(\sum_{j=0}^{h} c_{j}\omega^{[j]}\right)
\end{equation}
```
def jacobi_kain(A, b, rtol=1.0e-8, max_it=25, max_hist=8, x_0=None, print_report=False):
"""
Solve linear system of equations with Jacobi iterations and KAIN acceleration.
Parameters
----------
A
Stiffness matrix
b
RHS vector
rtol
Tolerance for solution norm
max_it
Maximum number of iterations
max_hist
Maximum size of KAIN queue
x_0
Initial guess
print_report
Whether to print iteration report inside the function
Returns
-------
x
Linear system solution vector
n_it
Number of iterations performed
report
Iteration number, residual norm sequence
"""
x = np.zeros_like(b) if x_0 is None else x_0
# Lists of iterates and residuals
xs = []
rs = []
report = []
it = 0
if print_report:
print('Iteration # Residual norm')
while it < max_it:
# Compute residual
r = apply_jacobi(A, b, x) - x
rnorm = LA.norm(r)
# Collect DIIS history
xs.append(x)
rs.append(r)
# Report
report.append(rnorm)
if print_report:
print(' {:4d} {:.5E}'.format(it, rnorm))
# Check convergence
if rnorm < rtol:
break
if it >= 2:
# Prune KAIN history
kain_hist = len(xs)
if kain_hist > max_hist:
xs.pop(0)
rs.pop(0)
kain_hist -= 1
# Build Q matrix
Q = np.empty((kain_hist, kain_hist))
for i, x in enumerate(xs):
for j, r in enumerate(rs):
Q[i, j] = np.einsum('i,i', x, r)
# Build subspace matrix B and vector rhs
B = np.empty((kain_hist - 1, kain_hist - 1))
rhs = np.zeros(kain_hist - 1)
for i in range(kain_hist - 1):
rhs[i] = -Q[i, -1] + Q[-1, -1]
for j in range(kain_hist - 1):
B[i, j] = Q[i, j] - Q[i, -1] - Q[-1, j] + Q[-1, -1]
# Solve KAIN equations
cs = LA.solve(B, rhs)
cs = np.append(cs, 1.0 - np.sum(cs))
# Calculate new solution as linear
# combination of previous solutions
x = np.zeros_like(x)
for i, c in enumerate(cs):
x += c * xs[i] - c * rs[i]
# Update solution vector
x = apply_jacobi(A, b, x)
it += 1
else:
raise RuntimeError(
'Maximum number of iterations ({0:d}) exceeded, but residual norm {1:.5E} still greater than threshold {2:.5E}'.
format(max_it, rnorm, rtol))
return x, it, report
```
## CROP acceleration
CHECK: CROP should only need 3 previous iterates at most for linear problems.
```
def jacobi_crop(A, b, rtol=1.0e-8, max_it=25, max_hist=8, x_0=None):
"""
Solve linear system of equations with Jacobi iterations and CROP acceleration.
Parameters
----------
A
Stiffness matrix
b
RHS vector
rtol
Tolerance for solution norm
max_it
Maximum number of iterations
max_hist
Maximum size of CROP queue
x_0
Initial guess
Returns
-------
x
Linear system solution vector
"""
pass
```
## Regularized nonlinear acceleration
See also: https://github.com/windows7lover/RegularizedNonlinearAcceleration
```
def jacobi_rna(A, b, rtol=1.0e-8, max_it=25, max_hist=8, x_0=None, reg=0.1, print_report=False):
"""
Solve linear system of equations with Jacobi iterations and RNA.
Parameters
----------
A
Stiffness matrix
b
RHS vector
rtol
Tolerance for solution norm
max_it
Maximum number of iterations
max_hist
Maximum size of RNA queue
x_0
Initial guess
reg
Regularization parameter
print_report
Whether to print iteration report inside the function
Returns
-------
x
Linear system solution vector
n_it
Number of iterations performed
report
Iteration number, residual norm sequence
"""
x = np.zeros_like(b) if x_0 is None else x_0
# Lists of iterates
xs = []
report = []
it = 0
if print_report:
print('Iteration # Residual norm')
while it < max_it:
# Compute residual
r = apply_jacobi(A, b, x) - x
rnorm = LA.norm(r)
# Collect history
xs.append(x)
# Report
report.append(rnorm)
if print_report:
print(' {:4d} {:.5E}'.format(it, rnorm))
# Check convergence
if rnorm < rtol:
break
if it >= 2:
# Prune history
hist = len(xs)
if hist > max_hist:
xs.pop(0)
hist -= 1
# Build R matrix of residuals
X = np.asarray(xs)
R = np.diff(X, axis=0)
# Build R^{t}R and normalize it
RR = np.einsum('ik,jk->ij', R, R)
print('RR is ', RR)
my_RR = RR = np.dot(np.transpose(R),R)
print('my_RR is ', my_RR)
RR /= np.sqrt(np.einsum('ii', RR))
# Solve RNA equations
zs = LA.solve(RR + reg*np.eye(RR.shape[0]), np.ones(RR.shape[0]))
# Recover mixing weights
cs = zs / np.sum(zs)
# Calculate new solution as linear
# combination of previous solutions
x = np.zeros_like(b)
for i, c in enumerate(cs[:-1]):
x += c * xs[i]
# Update solution vector
x = apply_jacobi(A, b, x)
# Update iteration counter
it += 1
else:
raise RuntimeError(
'Maximum number of iterations ({0:d}) exceeded, but residual norm {1:.5E} still greater than threshold {2:.5E}'.
format(max_it, rnorm, rtol))
return x, it, report
def relative_error_to_reference(x, x_ref):
return LA.norm(x - x_ref) / LA.norm(x_ref)
print('Experiments with linear solvers')
dim = 100
M = np.random.randn(dim, dim)
# Make sure our matrix is SPD
A = 0.5 * (M + M.transpose())
A = A * A.transpose()
A += dim * np.eye(dim)
b = np.random.rand(dim)
x_ref = LA.solve(A, b)
methods = OrderedDict({
'Jacobi' : jacobi,
'Jacobi-DIIS' : jacobi_diis,
'Jacobi-KAIN' : jacobi_kain,
#'Jacobi-RNA' : jacobi_rna, # NOT working
'Kaczmarz' : kaczmarz,
'Conjugate gradient' : cg,
'Conjugate residual': cr
})
reports = OrderedDict()
for k, v in methods.items():
print('@ {:s} algorithm'.format(k))
x, it, report = v(A, b)
rel_err = relative_error_to_reference(x, x_ref)
print('{:s} converged in {:d} iterations with relative error to reference {:.5E}\n'.format(k, it, rel_err))
reports.update({k: report})
print('--- Final report ---')
# TODO generate table
#header = ' '.join(reports.keys())
colors = itertools.cycle(palette)
p = figure(plot_width=800, plot_height=400, title='Convergence', y_axis_type='log')
for method, report, color in zip(reports.keys(), reports.values(), colors):
p.line(np.arange(len(reports['Kaczmarz'])), report, line_width=2, legend=method, color=color)
show(p)
```
| github_jupyter |
<a href="https://qworld.net" target="_blank" align="left"><img src="../qworld/images/header.jpg" align="left"></a>
$ \newcommand{\bra}[1]{\langle #1|} $
$ \newcommand{\ket}[1]{|#1\rangle} $
$ \newcommand{\braket}[2]{\langle #1|#2\rangle} $
$ \newcommand{\dot}[2]{ #1 \cdot #2} $
$ \newcommand{\biginner}[2]{\left\langle #1,#2\right\rangle} $
$ \newcommand{\mymatrix}[2]{\left( \begin{array}{#1} #2\end{array} \right)} $
$ \newcommand{\myvector}[1]{\mymatrix{c}{#1}} $
$ \newcommand{\myrvector}[1]{\mymatrix{r}{#1}} $
$ \newcommand{\mypar}[1]{\left( #1 \right)} $
$ \newcommand{\mybigpar}[1]{ \Big( #1 \Big)} $
$ \newcommand{\sqrttwo}{\frac{1}{\sqrt{2}}} $
$ \newcommand{\dsqrttwo}{\dfrac{1}{\sqrt{2}}} $
$ \newcommand{\onehalf}{\frac{1}{2}} $
$ \newcommand{\donehalf}{\dfrac{1}{2}} $
$ \newcommand{\hadamard}{ \mymatrix{rr}{ \sqrttwo & \sqrttwo \\ \sqrttwo & -\sqrttwo }} $
$ \newcommand{\vzero}{\myvector{1\\0}} $
$ \newcommand{\vone}{\myvector{0\\1}} $
$ \newcommand{\stateplus}{\myvector{ \sqrttwo \\ \sqrttwo } } $
$ \newcommand{\stateminus}{ \myrvector{ \sqrttwo \\ -\sqrttwo } } $
$ \newcommand{\myarray}[2]{ \begin{array}{#1}#2\end{array}} $
$ \newcommand{\X}{ \mymatrix{cc}{0 & 1 \\ 1 & 0} } $
$ \newcommand{\I}{ \mymatrix{rr}{1 & 0 \\ 0 & 1} } $
$ \newcommand{\Z}{ \mymatrix{rr}{1 & 0 \\ 0 & -1} } $
$ \newcommand{\Htwo}{ \mymatrix{rrrr}{ \frac{1}{2} & \frac{1}{2} & \frac{1}{2} & \frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & \frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} & \frac{1}{2} } } $
$ \newcommand{\CNOT}{ \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0} } $
$ \newcommand{\norm}[1]{ \left\lVert #1 \right\rVert } $
$ \newcommand{\pstate}[1]{ \lceil \mspace{-1mu} #1 \mspace{-1.5mu} \rfloor } $
$ \newcommand{\greenbit}[1] {\mathbf{{\color{green}#1}}} $
$ \newcommand{\bluebit}[1] {\mathbf{{\color{blue}#1}}} $
$ \newcommand{\redbit}[1] {\mathbf{{\color{red}#1}}} $
$ \newcommand{\brownbit}[1] {\mathbf{{\color{brown}#1}}} $
$ \newcommand{\blackbit}[1] {\mathbf{{\color{black}#1}}} $
<font style="font-size:28px;" align="left"><b>Basics of Python: Lists </b></font>
<br>
_prepared by Abuzer Yakaryilmaz_
<br><br>
We review using Lists in Python here.
Run each cell and check the results.
A list (or array) is a collection of objects (variables) separated by comma.
The order is important, and we can access each element in the list with its index starting from 0.
```
# here is a list holding all even numbers between 10 and 20
L = [10, 12, 14, 16, 18, 20]
# let's print the list
print(L)
# let's print each element by using its index but in reverse order
print(L[5],L[4],L[3],L[2],L[1],L[0])
# let's print the length (size) of list
print(len(L))
# let's print each element and its index in the list
# we use a for-loop, and the number of iteration is determined by the length of the list
L = [10, 12, 14, 16, 18, 20]
for i in range(len(L)):
print(L[i],"is the element in our list with the index",i)
# let's replace each number in the above list with its double value
# L = [10, 12, 14, 16, 18, 20]
# print the list before doubling operation
print("the list before doubling operation is",L)
for i in range(len(L)):
current_element=L[i] # get the value of the i-th element
L[i] = 2 * current_element # update the value of the i-th element
# let's shorten the code as
#L[i] = 2 * L[i]
# or
#L[i] *= 2
# print the list after doubling operation
print("the list after doubling operation is",L)
# after each execution of this cell, the latest values will be doubled
# so the values in the list will be exponentially increased
# let's define two lists
L1 = [1,2,3,4]
L2 = [-5,-6,-7,-8]
# two lists can be concatenated
# the result is a new list
print("the concatenation of L1 and L2 is",L1+L2)
# the order of terms is important
print("the concatenation of L2 and L1 is",L2+L1) # this is a different list than L1+L2
# we can add a new element to a list, which increases its length/size by 1
L = [10, 12, 14, 16, 18, 20]
print(L,"the current length is",len(L))
# we add two values by showing two different methods
# L.append(value) directly adds the value as a new element to the list
L.append(-4)
# we can also use concatenation operator +
L = L + [-8] # here [-8] is a list having a single element
print(L,"the new length is",len(L))
# a list can be multiplied with an integer
L = [1,2]
# we can consider the multiplication of L by an integer as a repeated "summation" (concatenation) of L by itself
# L * 1 is the list itself
# L * 2 is L + L (the concatenation of L with itself)
# L * 3 is L + L + L (the concatenation of L with itself twice)
# L * m is L + ... + L (the concatenation of L with itself m-1 times)
# L * 0 is the empty list
# L * i is the same as i * L
# let's print the different cases
for i in range(6):
print(i,"* L is",i*L)
# this operation can be useful when initializing a list with the same value(s)
# let's create a list of prime numbers less than 100
# here is a function that determines whether a given number is prime or not
def prime(number):
if number < 2: return False
if number == 2: return True
if number % 2 == 0: return False
for i in range(3,number,2):
if number % i == 0: return False
return True
# end of a function
# let's start with an empty list
L=[]
# what can the length of this list be?
print("my initial length is",len(L))
for i in range(2,100):
if prime(i):
L.append(i)
# alternative methods:
#L = L + [i]
#L += [i]
# print the final list
print(L)
print("my final length is",len(L))
```
For a given integer $n \geq 0$, $ S(0) = 0 $, $ S(1)=1 $, and $ S(n) = 1 + 2 + \cdots + n $.
We define list $ L(n) $ such that the element with index $n$ holds $ S(n) $.
In other words, the elements of $ L(n) $ are $ [ S(0)~~S(1)~~S(2)~~\cdots~~S(n) ] $.
Let's build the list $ L(20) $.
```
# let's define the list with S(0)
L = [0]
# let's iteratively define n and S
# initial values
n = 0
S = 0
# the number of iterations
N = 20
while n <= N: # we iterate all values from 1 to 20
n = n + 1
S = S + n
L.append(S)
# print the final list
print(L)
```
<h3> Task 1 </h3>
Fibonacci sequence starts with $ 1 $ and $ 1 $. Then, each next element is equal to the summation of the previous two elements:
$$
1, 1, 2 , 3 , 5, 8, 13, 21, 34, 55, \ldots
$$
Find the first 30 elements of the Fibonacci sequence, store them in a list, and then print the list.
You can verify the first 10 elements of your result with the above list.
```
#
# your solution is here
#
F = [1,1]
```
<a href="Python20_Basics_Lists_Solutions.ipynb#task1">click for our solution</a>
<h3> Lists of different objects </h3>
A list can have any type of values.
```
# the following list stores certain information about Asja
# name, surname, age, profession, partner(s) if any, kid(s) if any, the updated date of list
ASJA = ['Asja','Karaindrou',31,'composer',[],['Cemile','Aitmatov'],"March 07, 2021"]
print(ASJA)
# Remark that an element of a list can be another list as well.
```
<h3> Task 2 </h3>
Define a list $ N $ with 11 elements such that $ N[i] $ is another list with four elements such that $ [i, i^2, i^3, i^2+i^3] $.
The index $ i $ should be between $ 0 $ and $ 10 $.
```
#
# your solution is here
#
```
<a href="Python20_Basics_Lists_Solutions.ipynb#task2">click for our solution</a>
<h3> Dictionaries </h3>
The outcomes of a quantum program (circuit) will be stored in a dictionary.
Therefore, we very shortly mention about the dictionary data type.
A dictionary is a set of paired elements.
Each pair is composed by a key and its value, and any value can be accessed by its key.
```
# let's define a dictionary pairing a person with her/his age
ages = {
'Asja':32,
'Balvis':28,
'Fyodor':43
}
# let print all keys
for person in ages:
print(person)
# let's print the values
for person in ages:
print(ages[person])
```
| github_jupyter |
```
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
# You can write up to 20GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
train=pd.read_csv("/kaggle/input/jobathon-analytics-vidhya/train.csv")
test=pd.read_csv("/kaggle/input/jobathon-analytics-vidhya/test.csv")
s=pd.read_csv("/kaggle/input/jobathon-analytics-vidhya/sample_submission.csv")
train
train[train['Response']==1]
df=train.append(test,ignore_index=True)
df
df.isnull().sum()
for k in df.columns:
print(k,df[k].nunique())
# df['upper_age_bin'] = pd.qcut(df['Upper_Age'],q=10,labels=['upp_age'+str(x) for x in range(10)])
# df['lower_age_bin'] = pd.qcut(df['Lower_Age'],q=10,labels=['low_age'+str(x) for x in range(10)])
# df['Reco_Policy_Premium_bin'] = pd.cut(df['Reco_Policy_Premium'],bins=5,labels=['premium_'+str(x) for x in range(5)])
# df['Health Indicator']=df.groupby('age')['Health Indicator'].transform(
# lambda grp: grp.fillna(grp.mode().iloc[0])
# )
df['Health_Indicator_null'] = df['Health Indicator'].apply(lambda x: 1 if x!=x else 0)
df['Holding_Policy_Duration_null'] = df['Holding_Policy_Duration'].apply(lambda x: 1 if x!=x else 0)
df['Health Indicator']=df['Health Indicator'].fillna('X10')
df['age_dist']=(df['Upper_Age']-df['Lower_Age'])/df['Lower_Age']
df['age']=(df['Upper_Age']-df['Lower_Age'])
# df['Health Indicator']=df.groupby('age')['Health Indicator'].transform(
# lambda grp: grp.fillna(grp.mode().iloc[0])
# )
# df['Holding_Policy_Type']=df.groupby('age')['Holding_Policy_Type'].transform(
# lambda grp: grp.fillna(grp.mode().iloc[0])
# )
# df['Holding_Policy_Duration']=df.groupby('age')['Holding_Policy_Duration'].transform(
# lambda grp: grp.fillna(grp.mode().iloc[0])
# )
df['Holding_Policy_Type']=df['Holding_Policy_Type'].fillna('5.0')
df['Holding_Policy_Duration']=df['Holding_Policy_Duration'].fillna('5.0')
dum=['Accomodation_Type','Reco_Insurance_Type','Is_Spouse','Holding_Policy_Type']
df.isnull().sum()
df.columns
df.head()
agg_data={
'Reco_Policy_Premium':['sum','max','min','mean'],
'age_dist':['sum','max','min','mean'],
'Upper_Age':['mean','max'],
'Lower_Age':['mean','min'],
'Health Indicator':['size','nunique'],
'Region_Code':['size','nunique'],
'Holding_Policy_Duration':['size','nunique'],
'City_Code':['size','nunique'],
}
df1 = df.groupby(['Reco_Policy_Cat']).agg(agg_data)
df1.columns=['A_' + '_'.join(col).strip() for col in df1.columns.values]
df1.reset_index(inplace=True)
df1.head()
df=df.merge(df1,on='Reco_Policy_Cat',how='left')
agg_data={
'Reco_Policy_Premium':['sum','max','min','mean'],
'age_dist':['sum','max','min','mean'],
'Upper_Age':['mean','max'],
'Lower_Age':['mean','min'],
'Health Indicator':['size','nunique'],
'Holding_Policy_Duration':['size','nunique'],
'City_Code':['size','nunique'],
}
df1 = df.groupby(['Region_Code','Reco_Policy_Cat']).agg(agg_data)
df1.columns=['B_' + '_'.join(col).strip() for col in df1.columns.values]
df1.reset_index(inplace=True)
df1.head()
df=df.merge(df1,on=['Region_Code','Reco_Policy_Cat'],how='left')
agg_data={
'Reco_Policy_Premium':['sum','max','min','mean'],
'age_dist':['sum','max','min','mean'],
'Health Indicator':['size','nunique'],
'Upper_Age':['mean','max'],
'Holding_Policy_Duration':['size','nunique'],
'City_Code':['size','nunique'],
'Region_Code':['size','nunique'],
'Lower_Age':['mean','min'],
}
df1 = df.groupby(['Region_Code','Reco_Insurance_Type']).agg(agg_data)
df1.columns=['C_' + '_'.join(col).strip() for col in df1.columns.values]
df1.reset_index(inplace=True)
df1.head()
df=df.merge(df1,on=['Region_Code','Reco_Insurance_Type'],how='left')
agg_data={
'Reco_Policy_Premium':['sum','max','min','mean'],
'age_dist':['sum','max','min','mean'],
'Lower_Age':['mean','min'],
'Health Indicator':['size','nunique'],
'Holding_Policy_Duration':['size','nunique'],
'City_Code':['size','nunique'],
'Region_Code':['size','nunique'],
'Reco_Insurance_Type':['size','nunique'],
}
df1 = df.groupby(['Reco_Policy_Cat','Reco_Insurance_Type','Upper_Age']).agg(agg_data)
df1.columns=['D_' + '_'.join(col).strip() for col in df1.columns.values]
df1.reset_index(inplace=True)
df1.head()
df=df.merge(df1,on=['Reco_Policy_Cat','Reco_Insurance_Type','Upper_Age'],how='left')
df['F1'] = (df['Health Indicator'].astype(str)+' '+df['Holding_Policy_Duration'].astype(str)).astype('category')
df['F2'] = (df['Health Indicator'].astype(str)+' '+df['Holding_Policy_Type'].astype(str)).astype('category')
df['F3'] = (df['Health Indicator'].astype(str)+' '+df['Reco_Policy_Cat'].astype(str)).astype('category')
df['F4'] = (df['Health Indicator'].astype(str)+' '+df['Region_Code'].astype(str)).astype('category')
df['F5'] = (df['Holding_Policy_Duration'].astype(str)+' '+df['Holding_Policy_Type'].astype(str)).astype('category')
df['F6'] = (df['Holding_Policy_Duration'].astype(str)+' '+df['Reco_Policy_Cat'].astype(str)).astype('category')
df['F7'] = (df['Holding_Policy_Duration'].astype(str)+' '+df['Region_Code'].astype(str)).astype('category')
df['F8'] = (df['Holding_Policy_Type'].astype(str)+' '+df['Reco_Policy_Cat'].astype(str)).astype('category')
df['F9'] = (df['Holding_Policy_Type'].astype(str)+' '+df['Region_Code'].astype(str)).astype('category')
df['F10'] = (df['Region_Code'].astype(str)+' '+df['Reco_Policy_Cat'].astype(str)).astype('category')
df=pd.get_dummies(df,columns=dum,drop_first=True)
cat=['City_Code','Region_Code','Health Indicator','Holding_Policy_Duration'
,'Reco_Policy_Cat'
]
for k in cat:
df[k]=df[k].astype('category')
to_drop=[]
corr_matrix = df.drop(['ID','Response'],axis=1).corr()
# print(corr_matrix)
upper = corr_matrix.where(np.triu(np.ones(corr_matrix.shape), k=1).astype(np.bool))
upper
to_drop = [column for column in upper.columns if any(upper[column] > 0.95)]
print(to_drop)
df.drop(to_drop,inplace=True,axis=1)
# agg_data={
# 'Reco_Policy_Premium':['sum','max','min','mean'],
# 'age_dist':['sum','max','min','mean'],
# 'Upper_Age':['mean','max'],
# 'Lower_Age':['mean','min'],
# 'Region_Code':['size','nunique'],
# }
# ,
# df1 = df.groupby(['City_Code','Health Indicator', 'Holding_Policy_Duration', 'Reco_Policy_Cat']).agg(agg_data)
# df1.columns=['B_' + '_'.join(col).strip() for col in df1.columns.values]
# df1.reset_index(inplace=True)
# df1.head()
# df=df.merge(df1,on=['City_Code','Health Indicator', 'Holding_Policy_Duration', 'Reco_Policy_Cat'],how='left')
dftrain=df[df['Response'].isnull()!=True]
dftest=df[df['Response'].isnull()==True]
dftest.head()
X,y=dftrain.drop(['ID','Response'],axis=1),dftrain['Response']
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score,f1_score
X_train,X_val,y_train,y_val = train_test_split(X,y,test_size=0.25,random_state = 1994,stratify=y)
from catboost import CatBoostClassifier,Pool, cv
from lightgbm import LGBMClassifier,LGBMRegressor
from sklearn.model_selection import StratifiedKFold,train_test_split
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import accuracy_score,confusion_matrix,roc_auc_score,f1_score,mean_squared_error
import seaborn as sns
%matplotlib inline
import matplotlib.pyplot as plt
from sklearn.metrics import f1_score
def evaluate_exRmse_lgb(truth, predictions):
ex = np.exp(-np.sqrt(mean_squared_error(truth,predictions)))
return ('EXRMSE', ex, True)
m=LGBMClassifier(n_estimators=5000,random_state=1994,learning_rate=0.05,colsample_bytree=0.5,reg_alpha=4
)
# m=RidgeCV(cv=4)
m.fit(X_train,y_train,eval_set=[(X_train,y_train),(X_val, y_val)],eval_metric='AUC', early_stopping_rounds=200,verbose=200)
p=m.predict_proba(X_val)[:,-1]
print(roc_auc_score(y_val,p))
# import catboost
# class ModelOptimizer:
# best_score = None
# opt = None
# def __init__(self, model, X_train, y_train, categorical_columns_indices=None, n_fold=3, seed=1994, early_stopping_rounds=30, is_stratified=True, is_shuffle=True):
# self.model = model
# self.X_train = X_train
# self.y_train = y_train
# self.categorical_columns_indices = categorical_columns_indices
# self.n_fold = n_fold
# self.seed = seed
# self.early_stopping_rounds = early_stopping_rounds
# self.is_stratified = is_stratified
# self.is_shuffle = is_shuffle
# def update_model(self, **kwargs):
# for k, v in kwargs.items():
# setattr(self.model, k, v)
# def evaluate_model(self):
# pass
# def optimize(self, param_space, max_evals=10, n_random_starts=2):
# start_time = time.time()
# @use_named_args(param_space)
# def _minimize(**params):
# self.model.set_params(**params)
# return self.evaluate_model()
# opt = gp_minimize(_minimize, param_space, n_calls=max_evals, n_random_starts=n_random_starts, random_state=2405, n_jobs=-1)
# best_values = opt.x
# optimal_values = dict(zip([param.name for param in param_space], best_values))
# best_score = opt.fun
# self.best_score = best_score
# self.opt = opt
# print('optimal_parameters: {}\noptimal score: {}\noptimization time: {}'.format(optimal_values, best_score, time.time() - start_time))
# print('updating model with optimal values')
# self.update_model(**optimal_values)
# plot_convergence(opt)
# return optimal_values
# class CatboostOptimizer(ModelOptimizer):
# def evaluate_model(self):
# validation_scores = catboost.cv(
# catboost.Pool(self.X_train,
# self.y_train,
# cat_features=self.categorical_columns_indices),
# self.model.get_params(),
# nfold=self.n_fold,
# stratified=self.is_stratified,
# seed=self.seed,
# early_stopping_rounds=self.early_stopping_rounds,
# shuffle=self.is_shuffle,
# verbose=100,
# plot=False)
# self.scores = validation_scores
# test_scores = validation_scores.iloc[:, 2]
# best_metric = test_scores.max()
# return 1 - best_metric
# from skopt import gp_minimize
# from skopt.space import Real, Integer
# from skopt.utils import use_named_args
# from skopt.plots import plot_convergence
# import time
# cb = catboost.CatBoostClassifier(n_estimators=4000, # use large n_estimators deliberately to make use of the early stopping
# loss_function='Logloss',
# eval_metric='AUC',
# boosting_type='Ordered', # use permutations
# random_seed=1994,
# use_best_model=True,random_strength=200,bagging_temperature=0.5,l2_leaf_reg=4)
# cb_optimizer = CatboostOptimizer(cb, X_train, y_train,categorical_columns_indices=categorical_features_indices)
# params_space = [Real(0.01, 0.8, name='learning_rate'),]
# cb_optimal_values = cb_optimizer.optimize(params_space)
# 45034503462176484
train['Response'].value_counts()/train.shape[0]
from catboost import CatBoostRegressor,CatBoostClassifier
categorical_features_indices = np.where(X_train.dtypes =='category')[0]
categorical_features_indices
m2=CatBoostClassifier(n_estimators=3000,random_state=1994,
eval_metric='AUC',learning_rate=0.05,random_strength=200,
bagging_temperature=0.6,
l2_leaf_reg=4,
# class_weights={0:0.239947,1:0.760053},
# min_data_in_leaf=5,
# auto_class_weights='Balanced',
one_hot_max_size=10,
task_type='GPU')
m2.fit(X_train,y_train,eval_set=[(X_val, y_val)], early_stopping_rounds=100,verbose=200,cat_features=categorical_features_indices)
p=m2.predict_proba(X_val)[:,-1]
print(roc_auc_score(y_val,p))
from catboost import CatBoostRegressor,CatBoostClassifier
categorical_features_indices = np.where(X_train.dtypes =='category')[0]
categorical_features_indices
m3=CatBoostClassifier(n_estimators=3000,random_state=1994,eval_metric='AUC',learning_rate=0.05,random_strength=200,bagging_temperature=0.6,l2_leaf_reg=4,one_hot_max_size=10)
m3.fit(X_train,y_train,eval_set=[(X_train,y_train),(X_val, y_val)], early_stopping_rounds=100,verbose=200,cat_features=categorical_features_indices)
p3=m3.predict_proba(X_val)[:,-1]
print(roc_auc_score(y_val,p3))
# print(roc_auc_score(y_val,(p3+p)/2))
import matplotlib.pyplot as plt
import warnings
import seaborn as sns
sns.set_style('darkgrid')
warnings.filterwarnings('ignore')
%matplotlib inline
feature_imp = pd.DataFrame(sorted(zip(m2.feature_importances_, X.columns), reverse=True)[:200],
columns=['Value','Feature'])
plt.figure(figsize=(12,8))
sns.barplot(x="Value", y="Feature", data=feature_imp.sort_values(by="Value", ascending=False))
plt.title('LightGBM Features')
plt.tight_layout()
plt.show()
feat_cols = feature_imp['Feature'][:55].values
feat_cols
Xtest=dftest.drop(['ID','Response'],axis=1)
Xtest[feat_cols]
categorical_features_indices = np.where(X[feat_cols].dtypes =='category')[0]
categorical_features_indices
err=[]
y_pred_tot=[]
from sklearn.model_selection import KFold,StratifiedKFold,GroupKFold,GroupShuffleSplit
fold=StratifiedKFold(n_splits=10,shuffle=True,random_state=1994)
for train_index, test_index in fold.split(X[feat_cols],y):
X_train, X_test = X[feat_cols].iloc[train_index], X[feat_cols].iloc[test_index]
y_train, y_test = y[train_index], y[test_index]
m=CatBoostClassifier(n_estimators=3000,random_state=1994,
eval_metric='AUC',learning_rate=0.05,random_strength=200,
bagging_temperature=0.6,
l2_leaf_reg=4,
# class_weights={0:0.239947,1:0.760053},
# min_data_in_leaf=5,
# auto_class_weights='Balanced',
one_hot_max_size=10,
task_type='GPU')
m.fit(X_train,y_train,eval_set=[(X_test, y_test)], early_stopping_rounds=100,verbose=200,cat_features=categorical_features_indices)
preds=m.predict_proba(X_test,ntree_end=m.best_iteration_)[:,-1]
print("err: ",roc_auc_score(y_test,preds))
err.append(roc_auc_score(y_test,preds))
p = m.predict_proba(Xtest[feat_cols],ntree_end=m.best_iteration_)[:,-1]
y_pred_tot.append(p)
err2=[]
y_pred_tot2=[]
from sklearn.model_selection import KFold,StratifiedKFold,GroupKFold,GroupShuffleSplit
fold=StratifiedKFold(n_splits=10,shuffle=True,random_state=1994)
for train_index, test_index in fold.split(X[feat_cols],y):
X_train, X_test = X[feat_cols].iloc[train_index], X[feat_cols].iloc[test_index]
y_train, y_test = y[train_index], y[test_index]
m=CatBoostClassifier(n_estimators=3000,random_state=1994,eval_metric='AUC',learning_rate=0.05,random_strength=200,bagging_temperature=0.6,l2_leaf_reg=4,one_hot_max_size=10)
m.fit(X_train,y_train,eval_set=[(X_test, y_test)], early_stopping_rounds=100,verbose=200,cat_features=categorical_features_indices)
preds=m.predict_proba(X_test,ntree_end=m.best_iteration_)[:,-1]
print("err: ",roc_auc_score(y_test,preds))
err2.append(roc_auc_score(y_test,preds))
p = m.predict_proba(Xtest[feat_cols],ntree_end=m.best_iteration_)[:,-1]
y_pred_tot2.append(p)
err1=[]
y_pred_tot1=[]
from sklearn.model_selection import KFold,StratifiedKFold,GroupKFold,GroupShuffleSplit
fold=StratifiedKFold(n_splits=10,shuffle=True,random_state=1994)
for train_index, test_index in fold.split(X[feat_cols],y):
X_train, X_test = X[feat_cols].iloc[train_index], X[feat_cols].iloc[test_index]
y_train, y_test = y[train_index], y[test_index]
m=LGBMClassifier(n_estimators=5000,random_state=1994,learning_rate=0.05,colsample_bytree=0.5,reg_alpha=4)
m.fit(X_train,y_train,eval_set=[(X_train,y_train),(X_test, y_test)],eval_metric='AUC', early_stopping_rounds=50,verbose=200)
preds=m.predict_proba(X_test,num_iteration=m.best_iteration_)[:,-1]
print("err: ",roc_auc_score(y_test,preds))
err1.append(roc_auc_score(y_test,preds))
p = m.predict_proba(Xtest[feat_cols],num_iteration=m.best_iteration_)[:,-1]
y_pred_tot1.append(p)
# np.mean(y_pred_tot1,0)
np.mean(err),np.mean(err1),np.mean(err2)
# dftest['Response']=np.mean(y_pred_tot1,0)
# dftest[['ID','Response']].to_csv('AV-job-lgb-sub3.csv',index=False)
dftest['Response']=np.mean(y_pred_tot2,0)
dftest[['ID','Response']].to_csv('AV-job-cb-sub9.csv',index=False)
dftest['Response']=np.mean(y_pred_tot1,0)
dftest[['ID','Response']].to_csv('AV-job-lgb-sub9.csv',index=False)
dftest['Response']=np.mean(y_pred_tot,0)*0.1+np.mean(y_pred_tot1,0)*0.05+np.mean(y_pred_tot2,0)*0.85
dftest[['ID','Response']].to_csv('AV-job-stack-sub9.csv',index=False)
```
| github_jupyter |
```
#r "nuget: Mermaid.InteractiveExtension"
#!mermaid
%%{init: { 'logLevel': 'debug', 'theme': 'base', 'gitGraph': {'showBranches': false}} }%%
gitGraph
commit
branch hotfix
checkout hotfix
commit
branch develop
checkout develop
commit id:"ash" tag:"abc"
branch featureB
checkout featureB
commit type:HIGHLIGHT
checkout main
checkout hotfix
commit type:NORMAL
checkout develop
commit type:REVERSE
checkout featureB
commit
checkout main
merge hotfix
checkout featureB
commit
checkout develop
branch featureA
commit
checkout develop
merge hotfix
checkout featureA
commit
checkout featureB
commit
checkout develop
merge featureA
branch release
checkout release
commit
checkout main
commit
checkout release
merge main
checkout develop
merge release
#!mermaid
stateDiagram-v2
[*] --> Active
state Active {
[*] --> NumLockOff
NumLockOff --> NumLockOn : EvNumLockPressed
NumLockOn --> NumLockOff : EvNumLockPressed
--
[*] --> CapsLockOff
CapsLockOff --> CapsLockOn : EvCapsLockPressed
CapsLockOn --> CapsLockOff : EvCapsLockPressed
--
[*] --> ScrollLockOff
ScrollLockOff --> ScrollLockOn : EvScrollLockPressed
ScrollLockOn --> ScrollLockOff : EvScrollLockPressed
}
#!mermaid
stateDiagram-v2
state fork_state <<fork>>
[*] --> fork_state
fork_state --> State2
fork_state --> State3
state join_state <<join>>
State2 --> join_state
State3 --> join_state
join_state --> State4
State4 --> [*]
#!mermaid
graph TB
sq[Square shape] --> ci((Circle shape))
subgraph A
od>Odd shape]-- Two line<br/>edge comment --> ro
di{Diamond with <br/> line break} -.-> ro(Rounded<br>square<br>shape)
di==>ro2(Rounded square shape)
end
%% Notice that no text in shape are added here instead that is appended further down
e --> od3>Really long text with linebreak<br>in an Odd shape]
%% Comments after double percent signs
e((Inner / circle<br>and some odd <br>special characters)) --> f(,.?!+-*ز)
cyr[Cyrillic]-->cyr2((Circle shape Начало));
classDef green fill:#9f6,stroke:#333,stroke-width:2px;
classDef orange fill:#f96,stroke:#333,stroke-width:4px;
class sq,e green
class di orange
```
# UML from .NET types
Use the extension method `ExploreWithUmlClassDiagram()` to see the uml diagram of a .NET type
```
using Microsoft.DotNet.Interactive;
typeof(Kernel).ExploreWithUmlClassDiagram().Display()
```
| github_jupyter |
# K-Nearest Neighbor Regressor
This Code template is for the regression analysis using a simple KNeighborsRegressor based on the K-Nearest Neighbors algorithm.
### Required Packages
```
import warnings
import numpy as np
import pandas as pd
import seaborn as se
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.metrics import r2_score, mean_absolute_error, mean_squared_error
from sklearn.neighbors import KNeighborsRegressor
warnings.filterwarnings('ignore')
```
### Initialization
Filepath of CSV file
```
#filepath
file_path= ""
```
List of features which are required for model training .
```
#x_values
features=[]
```
Target feature for prediction.
```
#y_value
target=''
```
### Data Fetching
Pandas is an open-source, BSD-licensed library providing high-performance, easy-to-use data manipulation and data analysis tools.
We will use panda's library to read the CSV file using its storage path.And we use the head function to display the initial row or entry.
```
df=pd.read_csv(file_path)
df.head()
```
### Feature Selections
It is the process of reducing the number of input variables when developing a predictive model. Used to reduce the number of input variables to both reduce the computational cost of modelling and, in some cases, to improve the performance of the model.
We will assign all the required input features to X and target/outcome to Y.
```
X=df[features]
Y=df[target]
```
### Data Preprocessing
Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
```
def NullClearner(df):
if(isinstance(df, pd.Series) and (df.dtype in ["float64","int64"])):
df.fillna(df.mean(),inplace=True)
return df
elif(isinstance(df, pd.Series)):
df.fillna(df.mode()[0],inplace=True)
return df
else:return df
def EncodeX(df):
return pd.get_dummies(df)
```
Calling preprocessing functions on the feature and target set.
```
x=X.columns.to_list()
for i in x:
X[i]=NullClearner(X[i])
X=EncodeX(X)
Y=NullClearner(Y)
X.head()
```
#### Correlation Map
In order to check the correlation between the features, we will plot a correlation matrix. It is effective in summarizing a large amount of data where the goal is to see patterns.
```
f,ax = plt.subplots(figsize=(18, 18))
matrix = np.triu(X.corr())
se.heatmap(X.corr(), annot=True, linewidths=.5, fmt= '.1f',ax=ax, mask=matrix)
plt.show()
```
### Data Splitting
The train-test split is a procedure for evaluating the performance of an algorithm. The procedure involves taking a dataset and dividing it into two subsets. The first subset is utilized to fit/train the model. The second subset is used for prediction. The main motive is to estimate the performance of the model on new data.
```
x_train,x_test,y_train,y_test=train_test_split(X,Y,test_size=0.2,random_state=123)
```
### Model
KNN is one of the easiest Machine Learning algorithms based on Supervised Machine Learning technique. The algorithm stores all the available data and classifies a new data point based on the similarity. It assumes the similarity between the new data and data and put the new case into the category that is most similar to the available categories.KNN algorithm at the training phase just stores the dataset and when it gets new data, then it classifies that data into a category that is much similar to the available data.
#### Model Tuning Parameters
> - n_neighbors -> Number of neighbors to use by default for kneighbors queries.
> - weights -> weight function used in prediction. {**uniform,distance**}
> - algorithm -> Algorithm used to compute the nearest neighbors. {**‘auto’, ‘ball_tree’, ‘kd_tree’, ‘brute’**}
> - p -> Power parameter for the Minkowski metric. When p = 1, this is equivalent to using manhattan_distance (l1), and euclidean_distance (l2) for p = 2. For arbitrary p, minkowski_distance (l_p) is used.
> - leaf_size -> Leaf size passed to BallTree or KDTree. This can affect the speed of the construction and query, as well as the memory required to store the tree. The optimal value depends on the nature of the problem.
```
model=KNeighborsRegressor(n_jobs=-1)
model.fit(x_train,y_train)
```
#### Model Accuracy
We will use the trained model to make a prediction on the test set.Then use the predicted value for measuring the accuracy of our model.
```
print("Accuracy score {:.2f} %\n".format(model.score(x_test,y_test)*100))
```
> **r2_score**: The **r2_score** function computes the percentage variablility explained by our model, either the fraction or the count of correct predictions.
> **mae**: The **mean abosolute error** function calculates the amount of total error(absolute average distance between the real data and the predicted data) by our model.
> **mse**: The **mean squared error** function squares the error(penalizes the model for large errors) by our model.
```
y_pred=model.predict(x_test)
print("R2 Score: {:.2f} %".format(r2_score(y_test,y_pred)*100))
print("Mean Absolute Error {:.2f}".format(mean_absolute_error(y_test,y_pred)))
print("Mean Squared Error {:.2f}".format(mean_squared_error(y_test,y_pred)))
```
#### Prediction Plot
First, we make use of a plot to plot the actual observations, with x_train on the x-axis and y_train on the y-axis.
For the regression line, we will use x_train on the x-axis and then the predictions of the x_train observations on the y-axis.
```
plt.figure(figsize=(14,10))
plt.plot(range(20),y_test[0:20], color = "green")
plt.plot(range(20),model.predict(x_test[0:20]), color = "red")
plt.legend(["Actual","prediction"])
plt.title("Predicted vs True Value")
plt.xlabel("Record number")
plt.ylabel(target)
plt.show()
```
#### Creator: Thilakraj Devadiga , Github: [Profile](https://github.com/Thilakraj1998)
| github_jupyter |
# TensorFlow Dataset API
**Learning Objectives**
1. Learn how use tf.data to read data from memory
1. Learn how to use tf.data in a training loop
1. Learn how use tf.data to read data from disk
1. Learn how to write production input pipelines with feature engineering (batching, shuffling, etc.)
In this notebook, we will start by refactoring the linear regression we implemented in the previous lab so that is takes its data from a`tf.data.Dataset`, and we will learn how to implement **stochastic gradient descent** with it. In this case, the original dataset will be synthetic and read by the `tf.data` API directly from memory.
In a second part, we will learn how to load a dataset with the `tf.data` API when the dataset resides on disk.
```
!sudo chown -R jupyter:jupyter /home/jupyter/training-data-analyst
# Ensure the right version of Tensorflow is installed.
!pip freeze | grep tensorflow==2.1 || pip install tensorflow==2.1
import json
import math
import os
from pprint import pprint
import numpy as np
import tensorflow as tf
print(tf.version.VERSION)
```
## Loading data from memory
### Creating the dataset
Let's consider the synthetic dataset of the previous section:
```
N_POINTS = 10
X = tf.constant(range(N_POINTS), dtype=tf.float32)
Y = 2 * X + 10
```
We begin with implementing a function that takes as input
- our $X$ and $Y$ vectors of synthetic data generated by the linear function $y= 2x + 10$
- the number of passes over the dataset we want to train on (`epochs`)
- the size of the batches the dataset (`batch_size`)
and returns a `tf.data.Dataset`:
**Remark:** Note that the last batch may not contain the exact number of elements you specified because the dataset was exhausted.
If you want batches with the exact same number of elements per batch, we will have to discard the last batch by
setting:
```python
dataset = dataset.batch(batch_size, drop_remainder=True)
```
We will do that here.
```
# TODO 1
def create_dataset(X, Y, epochs, batch_size):
dataset = tf.data.Dataset.from_tensor_slices((X, Y))
dataset = dataset.repeat(epochs).batch(batch_size, drop_remainder=True)
return dataset
```
Let's test our function by iterating twice over our dataset in batches of 3 datapoints:
```
BATCH_SIZE = 3
EPOCH = 2
dataset = create_dataset(X, Y, epochs=1, batch_size=3)
for i, (x, y) in enumerate(dataset):
print("x:", x.numpy(), "y:", y.numpy())
assert len(x) == BATCH_SIZE
assert len(y) == BATCH_SIZE
assert EPOCH
```
### Loss function and gradients
The loss function and the function that computes the gradients are the same as before:
```
def loss_mse(X, Y, w0, w1):
Y_hat = w0 * X + w1
errors = (Y_hat - Y)**2
return tf.reduce_mean(errors)
def compute_gradients(X, Y, w0, w1):
with tf.GradientTape() as tape:
loss = loss_mse(X, Y, w0, w1)
return tape.gradient(loss, [w0, w1])
```
### Training loop
The main difference now is that now, in the traning loop, we will iterate directly on the `tf.data.Dataset` generated by our `create_dataset` function.
We will configure the dataset so that it iterates 250 times over our synthetic dataset in batches of 2.
```
# TODO 2
EPOCHS = 250
BATCH_SIZE = 2
LEARNING_RATE = .02
MSG = "STEP {step} - loss: {loss}, w0: {w0}, w1: {w1}\n"
w0 = tf.Variable(0.0)
w1 = tf.Variable(0.0)
dataset = create_dataset(X, Y, epochs=EPOCHS, batch_size=BATCH_SIZE)
for step, (X_batch, Y_batch) in enumerate(dataset):
dw0, dw1 = compute_gradients(X_batch, Y_batch, w0, w1)
w0.assign_sub(dw0 * LEARNING_RATE)
w1.assign_sub(dw1 * LEARNING_RATE)
if step % 100 == 0:
loss = loss_mse(X_batch, Y_batch, w0, w1)
print(MSG.format(step=step, loss=loss, w0=w0.numpy(), w1=w1.numpy()))
assert loss < 0.0001
assert abs(w0 - 2) < 0.001
assert abs(w1 - 10) < 0.001
```
## Loading data from disk
### Locating the CSV files
We will start with the **taxifare dataset** CSV files that we wrote out in a previous lab.
The taxifare datast files been saved into `../data`.
Check that it is the case in the cell below, and, if not, regenerate the taxifare
dataset by running the provious lab notebook:
```
!ls -l ../data/taxi*.csv
```
### Use tf.data to read the CSV files
The `tf.data` API can easily read csv files using the helper function
[tf.data.experimental.make_csv_dataset](https://www.tensorflow.org/api_docs/python/tf/data/experimental/make_csv_dataset)
If you have TFRecords (which is recommended), you may use
[tf.data.experimental.make_batched_features_dataset](https://www.tensorflow.org/api_docs/python/tf/data/experimental/make_batched_features_dataset)
The first step is to define
- the feature names into a list `CSV_COLUMNS`
- their default values into a list `DEFAULTS`
```
CSV_COLUMNS = [
'fare_amount',
'pickup_datetime',
'pickup_longitude',
'pickup_latitude',
'dropoff_longitude',
'dropoff_latitude',
'passenger_count',
'key'
]
LABEL_COLUMN = 'fare_amount'
DEFAULTS = [[0.0], ['na'], [0.0], [0.0], [0.0], [0.0], [0.0], ['na']]
```
Let's now wrap the call to `make_csv_dataset` into its own function that will take only the file pattern (i.e. glob) where the dataset files are to be located:
```
# TODO 3
def create_dataset(pattern):
return tf.data.experimental.make_csv_dataset(
pattern, 1, CSV_COLUMNS, DEFAULTS)
tempds = create_dataset('../data/taxi-train*')
print(tempds)
```
Note that this is a prefetched dataset, where each element is an `OrderedDict` whose keys are the feature names and whose values are tensors of shape `(1,)` (i.e. vectors).
Let's iterate over the two first element of this dataset using `dataset.take(2)` and let's convert them ordinary Python dictionary with numpy array as values for more readability:
```
for data in tempds.take(2):
pprint({k: v.numpy() for k, v in data.items()})
print("\n")
```
### Transforming the features
What we really need is a dictionary of features + a label. So, we have to do two things to the above dictionary:
1. Remove the unwanted column "key"
1. Keep the label separate from the features
Let's first implement a funciton that takes as input a row (represented as an `OrderedDict` in our `tf.data.Dataset` as above) and then returns a tuple with two elements:
* The first element beeing the same `OrderedDict` with the label dropped
* The second element beeing the label itself (`fare_amount`)
Note that we will need to also remove the `key` and `pickup_datetime` column, which we won't use.
```
UNWANTED_COLS = ['pickup_datetime', 'key']
# TODO 4a
def features_and_labels(row_data):
label = row_data.pop(LABEL_COLUMN)
features = row_data
for unwanted_col in UNWANTED_COLS:
features.pop(unwanted_col)
return features, label
```
Let's iterate over 2 examples from our `tempds` dataset and apply our `feature_and_labels`
function to each of the examples to make sure it's working:
```
for row_data in tempds.take(2):
features, label = features_and_labels(row_data)
pprint(features)
print(label, "\n")
assert UNWANTED_COLS[0] not in features.keys()
assert UNWANTED_COLS[1] not in features.keys()
assert label.shape == [1]
```
### Batching
Let's now refactor our `create_dataset` function so that it takes an additional argument `batch_size` and batch the data correspondingly. We will also use the `features_and_labels` function we implemented in order for our dataset to produce tuples of features and labels.
```
# TODO 4b
def create_dataset(pattern, batch_size):
dataset = tf.data.experimental.make_csv_dataset(
pattern, batch_size, CSV_COLUMNS, DEFAULTS)
return dataset.map(features_and_labels)
```
Let's test that our batches are of the right size:
```
BATCH_SIZE = 2
tempds = create_dataset('../data/taxi-train*', batch_size=2)
for X_batch, Y_batch in tempds.take(2):
pprint({k: v.numpy() for k, v in X_batch.items()})
print(Y_batch.numpy(), "\n")
assert len(Y_batch) == BATCH_SIZE
```
### Shuffling
When training a deep learning model in batches over multiple workers, it is helpful if we shuffle the data. That way, different workers will be working on different parts of the input file at the same time, and so averaging gradients across workers will help. Also, during training, we will need to read the data indefinitely.
Let's refactor our `create_dataset` function so that it shuffles the data, when the dataset is used for training.
We will introduce a additional argument `mode` to our function to allow the function body to distinguish the case
when it needs to shuffle the data (`mode == "train"`) from when it shouldn't (`mode == "eval"`).
Also, before returning we will want to prefetch 1 data point ahead of time (`dataset.prefetch(1)`) to speedup training:
```
# TODO 4c
def create_dataset(pattern, batch_size=1, mode="eval"):
dataset = tf.data.experimental.make_csv_dataset(
pattern, batch_size, CSV_COLUMNS, DEFAULTS)
dataset = dataset.map(features_and_labels).cache()
if mode == "train":
dataset = dataset.shuffle(1000).repeat()
# take advantage of multi-threading; 1=AUTOTUNE
dataset = dataset.prefetch(1)
return dataset
```
Let's check that our function work well in both modes:
```
tempds = create_dataset('../data/taxi-train*', 2, "train")
print(list(tempds.take(1)))
tempds = create_dataset('../data/taxi-valid*', 2, "eval")
print(list(tempds.take(1)))
```
In the next notebook, we will build the model using this input pipeline.
Copyright 2019 Google Inc.
Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
| github_jupyter |
<h1>CS4618: Artificial Intelligence I</h1>
<h1>Introduction</h1>
<h2>
Derek Bridge<br />
School of Computer Science and Information Technology<br />
University College Cork
</h2>
<h1>What is intelligence?</h1>
<ul>
<li>Some people define it in terms of outward behaviours, e.g. the <i>Turing Test</i>.</li>
<li>Some people define it in terms of various skills, e.g. reasoning, planning, learning, …
or playing chess, composing poetry, …
<ul>
<li>In particular, these people often concentrate on human skills (chess, language,...),
resulting in a very human-centric definition.
</li>
</ul>
</li>
<li>Here are three better definitions:
<ul>
<li>Nils J. Nilsson: "…intelligence is that quality that enables an entity to
function appropriately and with foresight in its environment."
</li>
<li>My definition: "A system's degree of intelligence is defined in terms of its capacity to
act autonomously and rationally when faced with disorder, uncertainty, imprecision and
intractability."
</li>
<li>François Chollet: "The intelligence of a system is a measure of its skill-acquisition
efficiency over a scope of tasks, with respect to priors, experience, and generalization
difficulty."
</li>
</ul>
</li>
<li>No matter how we define it, intelligence lies on a multi-dimensional spectrum.</li>
</ul>
<figure style="text-align: center;">
<img src="images/other_intelligences.jpg" />
<figcaption>
Cartoon by <a href="https://twitter.com/FalseKnees">False Knees</a>
</figcaption>
</figure>
<h1>What is AI?</h1>
<ul>
<li>AI is the field that studies intelligence by trying to synthesize it.</li>
</ul>
<h1>Is AI even possible?</h1>
<table>
<tr>
<td style="border: 1px solid blue; vertical-align: top; text-align: left;">
<b>No</b>: there's a special and essential ingredient that can't be replicated, e.g. soul, spirit,
consciousness, free will, creativity, humour, …
<p>
Perhaps we can <b>simulate</b> intelligence:
</p>
<ul>
<li>Outwardly, systems may <em>behave as if</em> intelligent.</li>
<li>But the way they achieve this behaviour (the internal process) doesn't qualify as true
thinking.
</li>
</ul>
</td>
<td style="border: 1px solid blue; vertical-align: top;">
<b>Yes</b>, we can build <b>true human-like</b> intelligence.
</td>
<td style="border: 1px solid blue; vertical-align: top;">
<b>Yes</b>, we can build true intelligences but they won't necessarily be like us.<br />
AI = <b>alien intelligence</b>.
</td>
</tr>
</table>
<ul>
<li>Where do you sit in this table? Or, do you have a different view?</li>
</ul>
<h1>Why do we want to build intelligent systems?</h1>
<ul>
<li>The goal of most people who work in AI is to build smarter tools.</li>
<li>Some rough-and-ready definitions:
<ul>
<li>A <b>tool</b> is an object that a creature uses intentionally to transform another object
(possibly including the creature itself).
</li>
<li>A tool is only <b>useful</b> if it:
<ul>
<li><b>increases throughput</b>: more output for the same input or less input (better use of scarce
resources, including time); and/or
</li>
<li><b>reduces externalities</b>: fewer undesirable side-effects (e.g. less pollution, lower
risks to life and limb, etc.).
</li>
</ul>
</li>
</ul>
</li>
</ul>
<h2>Examples</h2>
<ul>
<li>So, based on the previous definitions, why do we want to build…
<table style="width: 100%;">
<tr>
<td>…self-driving vehicles?</td><td>…intelligent news filters?</td>
<td>…medical diagnosis tools?</td><td>…bomb disposal robots?</td>
</tr>
<tr>
<td><img src="images/car.jpg" /></td>
<td><img src="images/news.jpg" /></td>
<td><img src="images/medicine.jpg" /></td>
<td><img src="images/bomb.jpg" /></td>
</tr>
</table>
</li>
<li>All social media companies use AI. Why?
<div style="display: flex; border: 1px solid black;">
<img src="images/google.jpg" />
<img src="images/facebook.jpg" />
<img src="images/spotify.jpg" />
<img src="images/amazon.jpg" />
</div>
</li>
</ul>
<h1>Narrow AI versus Artificial General Intelligence (AGI)</h1>
<ul>
<li>Narrow AI:
<ul>
<li>We are bulding systems that are highly specialized, often for a single task in a single domain.
</li>
</ul>
<table>
<tr>
<td style="border-right-width: 0"><img style="width: 200px" src="images/togelius.jpg" /></td>
<td style="border-left-width: 0">
<a href="http://togelius.blogspot.ie/2017/07/some-advice-for-journalists-writing.html">
Some advice for journalists writing about AI:
</a><br />
"AI is a collection of methods … that can do something impressive, such as playing
a game or drawing pictures of cats. However, you can safely assume that the same system
cannot both play games and draw pictures of cats. … [Journalists can] make it seem
like there are machines with general intelligence out there. There are not."
</td>
</tr>
</table>
<ul>
<li>For narrow AI, we are having some amazing successes, e.g. <a href="https://www.nature.com/articles/d41586-020-00018-3">discovery of a new antibiotic</a>.
</li>
<li>But, equally, these narrow AI systems are surprisingly brittle.
</li>
</ul>
</li>
<li>AGI:
<ul>
<li>We are nowhere near close to achieving AGI.</li>
<li>In fact, we are not working on this!</li>
</ul>
</li>
</ul>
<h1>Risks</h1>
<ul>
<li>Journalists are exaggerating AI success stories (narrow AI) to imply that AGI is just around the corner.
<ul>
<li>And if AGI is just around the corner, this leads observers to warn about existential risks:
<table style="width: 100%;">
<tr>
<td style="border-right-width: 0">
<img style="width: 100px" src="images/musk.jpg" />
</td>
<td style="border-left-width: 0; border-right: 1px solid black;">
"…the most serious threat to the survival of the human race…"
</td>
<td style="border-right-width: 0">
<img style="width: 100px" src="images/hawking.jpg" />
</td>
<td style="border-left-width: 0">
"The development of full artificial intelligence could spell the end of the human
race…It would take off on its own, and re-design itself at an ever
increasing rate."
</td>
</tr>
</table>
</li>
<li>People who work within the field are more realistic:
<table style="width: 100%;">
<tr>
<td style="border-right-width: 0">
<img style="width: 100px" src="images/ng.jpg" />
</td>
<td style="border-left-width: 0">
"I don’t work on not turning AI evil today for the same reason I don't worry
about the problem of overpopulation on the planet Mars."
</td>
</tr>
</table>
</li>
</ul>
</li>
<li>The real risks:
<ul>
<li>The real risks include the following: lack of explainability; fragility; bias and data quality;
fakes; high energy costs; military uses; unemployment; surveillance by government agencies;
surveillance by big tech; …
</li>
<li>Let's briefly look at one: fragility. The problem is not that AI is too good.
The problem is that AI is not good enough.
<figure style="width: 100%; border: 1px solid black;" >
<img src="images/adv1.png" /><br />
<img src="images/adv2.png" />
</figure>
</li>
<li>People are more and more thinking about our responsibilities in this field, e.g.:
<a href="https://ec.europa.eu/futurium/en/ai-alliance-consultation">
Ethics Guidelines for Trustworthy AI
</a>
</li>
</ul>
</li>
</ul>
| github_jupyter |
# BOFdat usage example
```
import cobra
from BOFdat import step1,step2,step3
from BOFdat.util import update
from cobra.util.solver import linear_reaction_coefficients
```
## Generating stoichiometric coefficients for the biomass objective function of *E.coli* genome-scale model *i*ML1515
The weight percentage and abundance of each molecule in the cell may vary from an organism to another and vary between growth conditions for a given organism [1,2]. BOFdat allows to incorporate macromolecular cell composition obtained from literature or new experiments to generate new stoichiometric coefficients for your model's biomass objective function (BOF). Once weight percentages are obtained, OMIC data can be incorporated to buff the coefficients and fit to experimental reality.
### Steps
The following example will lead you through all the necessary steps for the generation of the BOF stoichiometric coefficients (BOFsc) for *E.coli* K12 MG1655 GEM *i*ML1515 [3].
1. Obtain the macromolecular composition of the organism
2. Obtain OMICs experimental data
3. Generate BOFsc
4. Generate NGAM and GAM
5. Update BOF (BOFdat!)
#### Sources
[1] Dennis P. Patrick and Bremmer Hans. (1974) Macromolecular composition during steady-state growth of *Escherichia coli* B/r. Journal of bacteriology
[2] Benjamin Volkmer and Matthias Heinemann. (2011) Condition-Dependent Cell Volume and Concentration of Escherichia coli to Facilitate Data Conversion for Systems Biology Modeling. PLoS One
[3] Jonathan M Monk, Colton J Lloyd, Elizabeth Brunk, Nathan Mih, Anand Sastry, Zachary King, Rikiya Takeuchi, Wataru Nomura, Zhen Zhang, Hirotada Mori, Adam M Feist and Bernhard O Palsson. (2017) *i*ML1515, a knowledgebase that computes Escherichia coli traits. Nat. Biotech.
### 1. Obtain the macromolecular compositon of the organism
*E.coli* has been characterized thoroughly in literature. The BOFsc used in *i*AF1260 [4] are the same in *i*ML1515 [3] and were obtained from Neidhart *et. al* [5].
**Note:** The package also provides the option to include the percentage of each type of RNA molecule in the cell (ribosomal, transfer and messenger). The default values are rRNA: 0.9, tRNA 0.05 and mRNA: 0.05.
#### Sources
[4] Adam M Feist, Christopher S Henry, Jennifer L Reed, Markus Krummenacker, Andrew R Joyce, Peter D Karp, Linda J Broadbelt, Vassily Hatzimanikatis and Bernhard Ø Palsson. (2007) A genome-scale metabolic reconstruction for Escherichia coli K-12 MG1655 that accounts for 1260 ORFs and thermodynamic information. Mol. Syst. Bio.
[5] Neidhardt FC, Ingraham JL, Schaechter M (1990) Physiology of the Bacterial Cell: a Molecular Approach. Sinauer Associates: Sunderland, Mass
```
#Set parameters based on dry weight composition
dna_weight_fraction = 0.031
rna_weight_fraction = 0.205
protein_weight_fraction = 0.55
lipid_weight_fraction = 0.1
metabolite_weight_fraction = 0.1
#Give the path to each file as function parameters
#Genome file in BioPython supported format (.faa, .fna) and GenBank file
#also in BioPython supported format (.gb, .gbff)
genome = 'Ecoli_DNA.fna'
genbank = 'Ecoli_K12_MG1655.gbff'
#OMICs data as a 2 column csv file, gene and abundance
transcriptomic = 'transcriptomic.csv'
proteomic = 'proteomic.csv'
#Lipidomic abundances and conversion to model identifier
lipidomic_abundances = 'lipidomic_abundances.csv'
lipidomic_conversion = 'lipidomic_conversion.csv'
#Metabolomic abundances and conversion to model identifier
metabolomic_abundances = 'metabolomic_abundances.csv'
metabolomic_conversion = 'metabolomic_conversion.csv'
#Growth data on different carbon sources, uptake and secretion rates
maintenance = 'maintenance.csv'
#The model for which the coefficients are generated
model = 'iML1515.json'
```
### 2. Obtain OMICs experimental data
Your genome should have a GenBank annotated file. This file should be provided in a BioPython supported format (.gb, .gbff).
Search in literature allowed to find multiple OMICs dataset for different macromolecules that can be used to generate stoichiometric coefficients [6,7,8]. The data should be converted into a 2 column csv file.
The genome file should be provided in a standard BioPython supported format (.faa or .fna) and is used to calculate the abundance of each base in the genome.
Transcriptomic and proteomic files are 2 column csv files where the first column is the **gene identifier ** and the second column is the relative abundance of each of these genes in the cell.
Unlike DNA, RNA and proteins that are standard amongst every known life form, the lipid and metabolites in different organisms may vary. Hence a conversion file is required. This first column of this file is the original name of the compound and the second is the target identifier that this compound should have in your model. The first column of the abundance file gives the compound identifier in the model and the second column gives the abundance of that compound in the OMIC dataset.
#### Sources
[6] Sang Woo Seo, Donghyuk Kim, Haythem Latif, Edward J. O’Brien, Richard Szubin & Bernhard O. Palsson. (2014) Deciphering Fur transcriptional regulatory network highlights its complex role beyond iron metabolism in Escherichia coli. Nat. Comm.
[7] Alexander Schmidt, Karl Kochanowski, Silke Vedelaar, Erik Ahrné, Benjamin Volkmer, Luciano Callipo, Kèvin Knoops, Manuel Bauer, Ruedi Aebersold and Matthias Heinemann. (2016) The quantitative and condition-dependent *Escherichia coli* proteome. Nat. Biotech.
[8] Kian-Kai Cheng, Baek-Seok Lee, Takeshi Masuda, Takuro Ito, Kazutaka Ikeda, Akiyoshi Hirayama, Lingli Deng, Jiyang Dong, Kazuyuki Shimizu, Tomoyoshi Soga, Masaru Tomita, Bernhard O. Palsson and Martin Robert. (2014) Global metabolic network reorganization by adaptive mutations allows fast growth of Escherichia coli on glycerol. Nat Comm.
### 3. STEP 1: Generate BOFsc for macromolecules and generate maintenance costs
BOFdat operates with a single get_coefficient function for each macromolecule used. Input the parameters determined above as function parameters. Each function outputs a dictionary of metabolite and stoichiometric coefficients. The dictionary can be used to update the BOF (Step 5).
```
dna_coefficients = step1.generate_dna_coefficients(genome,model,DNA_WEIGHT_FRACTION=dna_weight_fraction)
dna_coefficients
rna_coefficients = step1.generate_rna_coefficients(genbank,model,transcriptomic,RNA_WEIGHT_FRACTION=rna_weight_fraction)
rna_coefficients
protein_coefficients = step1.generate_protein_coefficients(genbank,model,proteomic,PROTEIN_WEIGHT_FRACTION=protein_weight_fraction)
protein_coefficients
lipid_coefficients = step1.generate_lipid_coefficients(lipidomic_abundances,lipidomic_conversion,model,LIPID_WEIGHT_FRACTION=lipid_weight_fraction)
lipid_coefficients
```
### 4. Generate GAM and NGAM
Growth-associated maintenance (GAM) is the ATP cost related to growth. This includes the polymerization cost of each macromolecule. This cost is unaccounted for in the BOF because the model synthesizes the building blocks of each macromolecule in sufficient quantity to reflect the cell composition but not the cost of assembling those building blocks together. The GAM can be calculated experimentally by growing the bacteria on different sources of carbon at different starting concentrations. The carbon source should be the sole source of carbon in the media and its concentration should be measured after a given time. These remaining concentrations along with the excretion products are used by the package to constrain the model and calculate the ATP cost of growth.
The file format is shown in the image below:
The carbon sources are in the first column. The second column entails
Non growth-associated maintenance (NGAM) is the ATP cost of sustaining life without providing and
```
maintenance_cost = step1.generate_maintenance_costs(maintenance,model)
json_model = cobra.io.load_json_model('iML1515.json')
update.make_new_BOF(json_model,
dna_coefficients,
rna_coefficients,
protein_coefficients,
lipid_coefficients,maintenance=maintenance_cost)
#Model now has a step1 biomass
json_model.slim_optimize()
```
### 5- Update BOF (BOFdat!)
All the dictionaries have been generated. Now it would be fun to start playing with the model. Actually BOFdat allows you to use the generated dictionaries to update and buff your BOF experimental data. Just buff that!
```
#Let's load the model we want to update
ecoli = cobra.io.load_json_model(model)
#Let's see what the BOF looks like for E.coli iML1515
from cobra.util.solver import linear_reaction_coefficients
biomass = linear_reaction_coefficients(ecoli).keys()[0]
model_metab_id = [m.id[:-2] for m in ecoli.metabolites]
model_metab_id
bd.dna.update_biomass_coefficients(dna_coefficients,ecoli)
bd.rna.update_biomass_coefficients(rna_coefficients,ecoli)
bd.protein.update_biomass_coefficients(protein_coefficients,ecoli)
bd.lipid.update_biomass_coefficients(lipid_coefficients,ecoli)
bd.metabolite.update_biomass_coefficients(metab_coefficients,ecoli)
linear_reaction_coefficients(ecoli).keys()[0]
```
# That's it!
Your BOFsc have been updated in your model using BOFdat.
| github_jupyter |
```
from pynq.overlays.base import BaseOverlay
base = BaseOverlay('base.bit')
base.init_rf_clks()
#!/usr/bin/env python
"""
"""
import redis
import re
import numpy as np
import json
import time
board_name = "bu_rfsoc"
max_len = 10000000
r = redis.Redis(host='155.41.106.70', port=6379, db=0, password='Qsh4r9.VlMj5_HrvY#0f36')
def set_transmitter_channel(channel, enable, gain, frequency):
channel.control.enable = enable
channel.control.gain = gain
channel.dac_block.MixerSettings['Freq'] = frequency
def board_requests_handler(msg):
board_channel=1
if board_channel == 1:
set_transmitter_channel(base.radio.transmitter.channel[1], True, 0.8, 750)
number_samples=32768
print(f"GOT REQUEST: {msg}")
r.srem("active_command_boards", board_name)
channel = msg['channel'].decode()
# get id for request from channel name
req_id = re.findall(r'\d+', channel)[0]
r.delete(f"board-responses:{board_name}:{req_id}:metadata")
r.delete(f"board-responses:{board_name}:{req_id}:real")
r.delete(f"board-responses:{board_name}:{req_id}:imag")
r.delete(f"board-responses:{board_name}:{req_id}:complete")
req_params = json.loads(msg['data'])
# duration
duration = req_params['duration']
fc=base.radio.transmitter.channel[board_channel].dac_block.MixerSettings['Freq']*1e6
fs=base.radio.transmitter.channel[board_channel].dac_block.BlockStatus['SamplingFreq']*1e9
r.set(f"board-responses:{board_name}:{req_id}:complete", 'False')
cdata =[]
count = 0
b = base.radio.receiver.channel[board_channel]
#TODO: Find a better way to get a large amount of data from the board!!!!
start = time.time()
end = start + duration
try:
while time.time() < end:
cdata.append(b.transfer(number_samples))
count += 1
print(f"count: {count}")
for d in cdata:
xreal=d.real
xreal_json = json.dumps(xreal.tolist())
r.xadd(f"board-responses:{board_name}:{req_id}:real", {'data': xreal_json}, maxlen=max_len)
ximag=d.imag
ximag_json = json.dumps(ximag.tolist())
r.xadd(f"board-responses:{board_name}:{req_id}:imag", {'data': ximag_json}, maxlen=max_len)
finally:
print("ENTER BOARD_REQUEST_HANDLER FINALLY")
r.set(f"board-responses:{board_name}:{req_id}:complete", 'True')
# TODO: A lot of this metadata was taken from a random DigitalRF file.
# It should be updated to match the actual board.
metadata = {
'sfreq': fs,
'cfreq': fc,
'number_samples': number_samples,
'start_time': start,
'sample_rate_numerator' : int(fs),
'sample_rate_denominator' : 1,
'dtype_str' : "f8", # short int
'sub_cadence_secs' : (
3600 # Number of seconds of data in a subdirectory - typically MUCH larger
),
'file_cadence_seconds' : 1, # Each file will have up to 400 ms of data
'compression_level' : 1, # low level of compression
'checksum' : False, # no checksum
'is_complex' : True,# complex values
'is_continuous' : True,
'num_subchannels' : 1, # only one subchannel
'marching_periods' : False, # no marching periods when writing
'uuid' : None,# The digitalRF library will generate a UUID for us
'processing' : {
"channelizer_filter_taps" : [],
"decimation" : 2,
"interpolation" : 1,
"resampling_filter_taps" : [],
"scaling" : 1.0,
},
'receiver': {
"antenna" : "ADC2",
"bandwidth" : 100000000.0,
"center_freq" : fs / 4,
"clock_rate" : 125000000.0,
"clock_source" : 'external',
"dc_offset" : False,
"description" : 'BU RFSoC',
"gain" : 50.0,
"id" : 'BU_RFSoC1',
"info" : { },
"iq_balance" : '',
"lo_export" : '',
"lo_offset" : 0.0,
"lo_source" : '',
"otw_format" : '',
"stream_args" : '',
"subdev" : '',
"time_source" : 'external',
},
"uuid_str" : 'a8012bf59eeb49d6a71fbfdcddf1efbb',
}
print(f"sending metadata: {metadata}")
r.xadd(f"board-responses:{board_name}:{req_id}:metadata", {'data': json.dumps(metadata)})
r.sadd("active_command_boards", board_name)
def run_stream():
"""Set ups a redis connection and updates data to the stream."""
p = r.pubsub(ignore_subscribe_messages=True)
p.psubscribe(**{f'board-requests:{board_name}:*': board_requests_handler})
r.set(f'board-request-id:{board_name}', 0)
r.sadd("active_command_boards", board_name)
print("Waiting...")
try:
while True:
p.get_message()
time.sleep(.01)
finally:
print("ENTER FINALLY")
r.srem("active_command_boards", board_name)
if __name__ == '__main__':
run_stream()
```
| github_jupyter |
Use logistic regression to A/B test engineered features
```
import os
import logging
dir_path = os.path.realpath('..')
```
## Import data
```
import numpy as np
import pandas as pd
path = 'data/raw/train.csv'
full_path = os.path.join(dir_path, path)
df = pd.read_csv(full_path, header=0, index_col=0)
print("Dataset has {} rows, {} columns.".format(*df.shape))
# fill NaN with string "unknown"
df.fillna('unknown',inplace=True)
```
## Feature engineering
```
# Uppercase count
df['processed'] = df['comment_text'].str.split()
df['uppercase_count'] = df['processed'].apply(lambda x: sum(1 for t in x if t.isupper() and len(t)>2))
df = df.drop(['processed'], axis=1)
df.head()
```
## Pre-processing
```
from sklearn.model_selection import train_test_split
seed = 42
np.random.seed(seed)
test_size = 0.2
target = ['toxic', 'severe_toxic', 'obscene', 'threat', 'insult', 'identity_hate']
corpus = 'comment_text'
X = df.drop(target, axis=1)
y = df[target]
Xtrain, Xtest, ytrain, ytest = train_test_split(X, y, test_size=test_size, random_state=seed)
%%time
from sklearn.feature_extraction.text import TfidfVectorizer
vectorizer = TfidfVectorizer( max_features = 20000, ngram_range = ( 1, 3 ), sublinear_tf = True )
Xtrain_TFIDF = vectorizer.fit_transform(Xtrain[corpus])
Xtest_TFIDF = vectorizer.transform(Xtest[corpus])
```
## Model train - benchmark
```
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import log_loss
from sklearn.multiclass import OneVsRestClassifier
%%time
# Fit model
param_grid = {'estimator__C': [1, 10, 100] }
# Fit model
clf = OneVsRestClassifier(LogisticRegression(), n_jobs=-1)
clf_cv = GridSearchCV(clf, param_grid, cv=3, verbose=1)
clf_cv.fit(Xtrain_TFIDF, ytrain)
# Evaluate
y_pred = clf_cv.predict_proba(Xtest_TFIDF)
hold_out_preds = pd.DataFrame(y_pred, index=ytest.index, columns=target)
losses = []
for label in target:
loss = log_loss(ytest[label], hold_out_preds[label])
losses.append(loss)
print("{} log loss is {} .".format(label, loss))
print("Combined log loss: {} .".format(np.mean(losses)))
print(clf_cv.best_params_)
```
## Model train - with uppercase count
```
from scipy import sparse
Xtrain_upper = sparse.hstack((Xtrain_TFIDF, sparse.csr_matrix(Xtrain['uppercase_count'].values).T))
Xtest_upper = sparse.hstack((Xtest_TFIDF, sparse.csr_matrix(Xtest['uppercase_count'].values).T))
%%time
#Tune
param_grid = {'estimator__C': [0.1, 1, 10, 100] }
# Fit model
clf = OneVsRestClassifier(LogisticRegression(), n_jobs=-1)
clf_cv = GridSearchCV(clf, param_grid, cv=3, verbose=1)
clf_cv.fit(Xtrain_upper, ytrain)
# Evaluate
y_pred = clf_cv.predict_proba(Xtest_upper)
hold_out_preds = pd.DataFrame(y_pred, index=ytest.index, columns=target)
losses = []
for label in target:
loss = log_loss(ytest[label], hold_out_preds[label])
losses.append(loss)
print("{} log loss is {} .".format(label, loss))
print("Combined log loss: {} .".format(np.mean(losses)))
print(clf_cv.best_params_)
```
| github_jupyter |
**Plan for today**
Now we will start pulling things together.
- reading data
- storing it in a data structure
- sorting things stored in data structures
But before we start:
Over the next weeks we will be doing more pulling things together, review, etc.
To decide what we will be talking about, I need your help!
Anonymous mailbag tool:
www.slido.com
use event code:
cs1px
Let me know what you want to go over in more detail!
**We're going to use a modified subset of the firefighter game as an example today.**
We have a network that describes connections between nodes (which could be people/places/animals/whatever), and a fire is going to start somewhere and the spread over the network. We get to protect some vertices and they stay protected forever. This is a whole research area! We're going to do a very simplified task related to this game:
- we have decided that we want to defend in order of priority by number of connections
- so we want to sort our nodes by the number of connections they have, and give back that sorted list.
We want to write several functions:
1. Something to read in connections from a file (with exception-catching) and store the connections each node has in a data structure.
2. Something to tell us in any pair of nodes which should be higher-priority.
3. Using that, we want to sort the nodes by their number of connections.
First, let's look at an example file of connections:
```
1 2
2 3
2 4
4 1
4 6
6 5
```
Questions we need to answer:
1. How are we going to store these data?
2. How are we going to read the data?
One way we can store the data: as a dictionary of lists,
where the keys are vertices, and the values are lists of the vertices it is connected to
```
connectionsDict = {}
connectionsDict['1'] = ['2', '4']
connectionsDict['2'] = ['1', '3', '4']
connectionsDict['3'] = ['2']
connectionsDict['4'] = ['2', '1', '6']
connectionsDict['5'] = ['6']
connectionsDict['6'] = ['4', '5']
print(connectionsDict)
```
Now I want to read from a file:
```
def readConnections(filename):
connDict = {}
try:
with open(filename,"r") as f:
line = f.readline()
while line != "":
split = line.split()
if len(split) >= 2:
firstNode = split[0]
secondNode = split[1]
if firstNode not in connDict:
connDict[firstNode] = []
if secondNode not in connDict:
connDict[secondNode] = []
connDict[firstNode].append(secondNode)
connDict[secondNode].append(firstNode)
line = f.readline()
except Exception as e:
print('Something has gone wrong reading file ' + filename)
print (str(e))
finally:
return connDict
connDict = readConnections('edges.txt')
print(connDict)
```
Say I want to sort the vertices by the number of connections they have.
One way to do this is to define a special lessThan function
```
def lessThan(vertex1, vertex2, connDict):
numConnectionsV1 = len(connDict[vertex1])
numConnectionsV2 = len(connDict[vertex2])
return numConnectionsV1 < numConnectionsV2
lessThan('5', '3', connDict)
```
Now we can use this in a sorting algorithm:
```
def swap(myList, i, j):
tmp = myList[i]
myList[i] = myList[j]
myList[j] = tmp
def findMinimumAfterIncluding(myList, afterInd, infoDict):
minSoFar = afterInd
for i in range(afterInd, len(myList)):
if lessThan(myList[i], myList[minSoFar], infoDict):
minSoFar = i
return minSoFar
def selectionSort(myList, infoDict):
for i in range(len(myList)):
smallest = findMinimumAfterIncluding(myList, i, infoDict)
swap(myList, smallest, i)
listOfVertices = list(connDict.keys())
selectionSort(listOfVertices, connDict)
print(listOfVertices)
```
| github_jupyter |
```
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
from os import listdir
from os.path import join
from matplotlib.colors import LogNorm
path = "/Users/dgagne/data/cam_mp_files_run2_csv/"
mp_files = sorted(listdir(path))
mp_data_list = []
for mp_file in mp_files[:10]:
print(mp_file)
mp_data_list.append(pd.read_csv(join(path, mp_file), index_col="Index"))
mp_data = pd.concat(mp_data_list, ignore_index=True)
del mp_data_list[:]
del mp_data_list
mp_data["lambda_c"] =(np.pi * 1000.0 * mp_data["NC_TAU_in"] / (
mp_data["RHO_CLUBB_lev"] * mp_data["QC_TAU_in"])) ** 0.25
mp_data["lambda_r"] =(np.pi * 1000.0 * mp_data["NR_TAU_in"] / (
mp_data["RHO_CLUBB_lev"] * mp_data["QR_TAU_in"])) ** 0.25
mp_data.columns
plt.hist(mp_data["nrtend_MG2"], bins=100)
plt.gca().set_yscale("log")
np.count_nonzero((mp_data["nrtend_MG2"] > 0).values)
plt.hist(mp_data["lambda_c"], bins=100)
plt.gca().set_yscale("log")
np.count_nonzero(mp_data["QR_TAU_in"] == 0)
diameters = np.logspace(-4, -1, 50)
def size_distribution(diameters, l, n):
return n * np.exp(-l * diameters)
size_dist_out = size_distribution(diameters, mp_data.loc[::5000, ["lambda_c"]].values,
mp_data.loc[0::5000, ["NC_TAU_in"]].values)
plt.semilogx(diameters, np.sum(size_dist_out > 0, axis=0))
plt.semilogx(diameters, np.log10(size_dist_out[200]))
for size_dist_set in size_dist_out:
plt.semilogx(diameters, size_dist_set)
plt.gca().set_yscale("log")
plt.ylim(1e-10, 1e10)
plt.hist(np.log10(mp_data["lambda_c"][~np.isinf(mp_data["lambda_c"])]), bins=100)
plt.gca().set_yscale("log")
plt.hist(np.log(mp_data["NC_TAU_in"]))
plt.hist(np.log(mp_data["lambda_c"]), bins=100)
plt.gca().set_yscale("log")
np.count_nonzero(mp_data["qctend_MG2"] == 0)
mp_data.loc[mp_data["qctend_MG2"] == 0,
["QC_TAU_in", "NC_TAU_in", "QR_TAU_in", "NR_TAU_in", "RHO_CLUBB_lev"]].max()
mp_data.loc[(mp_data["qctend_MG2"] < 0) & (mp_data["QC_TAU_in"] < 1e-10),
["QC_TAU_in", "NC_TAU_in", "QR_TAU_in", "NR_TAU_in"]].max()
plt.hist2d(np.log10(mp_data.loc[mp_data["qctend_MG2"]==0, "QC_TAU_in"]),
np.log10(mp_data.loc[mp_data["qctend_MG2"]==0, "NC_TAU_in"]),
bins=100, cmin=1, norm=LogNorm())
fig, ax = plt.subplots(1,1,figsize=(8, 8))
plt.hist2d(np.log10(mp_data.loc[mp_data["QC_TAU_in"] > 1e-18, "QC_TAU_in"]),
np.log10(mp_data.loc[mp_data["QC_TAU_in"] > 1e-18,"NC_TAU_in"]),
bins=100, cmin=1, norm=LogNorm(), cmap="bone")
ax.set_facecolor("k")
plt.xlabel("QC")
plt.ylabel("NC")
plt.savefig("qc_nc_enterprise.png", dpi=200, bbox_inches="tight")
mean_tau = mp_data[["QC_TAU_in", "NC_TAU_in", "qrtend_TAU"]].groupby(["QC_TAU_in", "NC_TAU_in"]).mean()
mean_ta.stack()
np.log10(mp_data.loc[:,"NC_TAU_in"]).min()
mp_data
np.count_nonzero((mp_data["QC_TAU_in"] > 1e-6) & (mp_data["NC_TAU_in"] > 1e3)) / mp_data.shape[0]
mp_data.loc[mp_data["qctend_TAU"] < -1e-8,"NC_TAU_in"].min()
mp_data["qctend_TAU"].max()
mp_data["qrtend_TAU"].min()
plt.hist(np.log10(mp_data["NR_TAU_in"][mp_data["NR_TAU_in"] > 0]), bins=100)
plt.gca().set_yscale("log")
np.count_nonzero((mp_data["NC_TAU_in"] > 1e-12) & (mp_data["NR_TAU_in"] > 1e-12))
plt.hist(mp_data.loc[(mp_data["NC_TAU_in"] == 1e-12) & (mp_data["NR_TAU_in"]== 1e-12), "pressure"],
bins=20)
plt.hist(mp_data.loc[(mp_data["NC_TAU_in"] == 1e-12) & (mp_data["NR_TAU_in"]== 1e-12), "qrtend_MG2"],
bins=20)
plt.gca().set_yscale("log")
np.count_nonzero((mp_data["QC_TAU_in"] > 0) & (mp_data["QR_TAU_in"] == 0))
mp_data.loc[mp_data["QR_TAU_in"] == 0, ["lev"]].max()
plt.hist(mp_data["lambda_c"], bins=100)
plt.gca().set_yscale("log")
mp_data["lambda_c"].idxmax()
mp_data.loc[971060]
plt.hist2d(mp_data["qrtend_MG2"], mp_data["qrtend_TAU"], norm=LogNorm())
idx = np.where((mp_data["nrtend_MG2"] > 0) & (mp_data["nrtend_TAU"] > 0))[0]
plt.hist2d(np.log10(mp_data.loc[idx, "nrtend_MG2"]), np.log10(mp_data.loc[idx, "nrtend_TAU"]), norm=LogNorm(), bins=100)
np.count_nonzero((mp_data["NR_TAU_in"] + mp_data["nrtend_TAU"] * 1800) < 0)
(mp_data["nctend_MG2"] + mp_data["NC_TAU_in"]).min()
(mp_data["NR_TAU_in"] + mp_data["nrtend_TAU"] * 1800).min()
(mp_data["QC_TAU_in"] + mp_data["qctend_TAU"]).max()
mp_data.loc[(mp_data["NC_TAU_in"] + mp_data["nctend_TAU"] * 1800) < -1e-10, "nctend_TAU"]
plt.hist((mp_data["NC_TAU_in"] + mp_data["nctend_TAU"] * 1800), bins=100)
plt.gca().set_yscale("log")
```
| github_jupyter |
# Multi-Layer Perceptron, MNIST
---
In this notebook, we will train an MLP to classify images from the [MNIST database](http://yann.lecun.com/exdb/mnist/) hand-written digit database.
The process will be broken down into the following steps:
>1. Load and visualize the data
2. Define a neural network
3. Train the model
4. Evaluate the performance of our trained model on a test dataset!
Before we begin, we have to import the necessary libraries for working with data and PyTorch.
```
# import libraries
import torch
import numpy as np
```
---
## Load and Visualize the [Data](http://pytorch.org/docs/stable/torchvision/datasets.html)
Downloading may take a few moments, and you should see your progress as the data is loading. You may also choose to change the `batch_size` if you want to load more data at a time.
This cell will create DataLoaders for each of our datasets.
```
from torchvision import datasets
import torchvision.transforms as transforms
from torch.utils.data.sampler import SubsetRandomSampler
# number of subprocesses to use for data loading
num_workers = 0
# how many samples per batch to load
batch_size = 20
# percentage of training set to use as validation
valid_size = 0.2
# convert data to torch.FloatTensor
transform = transforms.ToTensor()
# choose the training and test datasets
train_data = datasets.MNIST(root='data', train=True,
download=True, transform=transform)
test_data = datasets.MNIST(root='data', train=False,
download=True, transform=transform)
# obtain training indices that will be used for validation
num_train = len(train_data)
indices = list(range(num_train))
np.random.shuffle(indices)
split = int(np.floor(valid_size * num_train))
train_idx, valid_idx = indices[split:], indices[:split]
# define samplers for obtaining training and validation batches
train_sampler = SubsetRandomSampler(train_idx)
valid_sampler = SubsetRandomSampler(valid_idx)
# prepare data loaders
train_loader = torch.utils.data.DataLoader(train_data, batch_size=batch_size,
sampler=train_sampler, num_workers=num_workers)
valid_loader = torch.utils.data.DataLoader(train_data, batch_size=batch_size,
sampler=valid_sampler, num_workers=num_workers)
test_loader = torch.utils.data.DataLoader(test_data, batch_size=batch_size,
num_workers=num_workers)
```
### Visualize a Batch of Training Data
The first step in a classification task is to take a look at the data, make sure it is loaded in correctly, then make any initial observations about patterns in that data.
```
import matplotlib.pyplot as plt
%matplotlib inline
# obtain one batch of training images
dataiter = iter(train_loader)
images, labels = dataiter.next()
images = images.numpy()
# plot the images in the batch, along with the corresponding labels
fig = plt.figure(figsize=(25, 4))
for idx in np.arange(20):
ax = fig.add_subplot(2, 20/2, idx+1, xticks=[], yticks=[])
ax.imshow(np.squeeze(images[idx]), cmap='gray')
# print out the correct label for each image
# .item() gets the value contained in a Tensor
ax.set_title(str(labels[idx].item()))
```
### View an Image in More Detail
```
img = np.squeeze(images[1])
fig = plt.figure(figsize = (12,12))
ax = fig.add_subplot(111)
ax.imshow(img, cmap='gray')
width, height = img.shape
thresh = img.max()/2.5
for x in range(width):
for y in range(height):
val = round(img[x][y],2) if img[x][y] !=0 else 0
ax.annotate(str(val), xy=(y,x),
horizontalalignment='center',
verticalalignment='center',
color='white' if img[x][y]<thresh else 'black')
```
---
## Define the Network [Architecture](http://pytorch.org/docs/stable/nn.html)
The architecture will be responsible for seeing as input a 784-dim Tensor of pixel values for each image, and producing a Tensor of length 10 (our number of classes) that indicates the class scores for an input image. This particular example uses two hidden layers and dropout to avoid overfitting.
```
import torch.nn as nn
import torch.nn.functional as F
# define the NN architecture
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
# number of hidden nodes in each layer (512)
hidden_1 = 512
hidden_2 = 512
# linear layer (784 -> hidden_1)
self.fc1 = nn.Linear(28 * 28, hidden_1)
# linear layer (n_hidden -> hidden_2)
self.fc2 = nn.Linear(hidden_1, hidden_2)
# linear layer (n_hidden -> 10)
self.fc3 = nn.Linear(hidden_2, 10)
# dropout layer (p=0.2)
# dropout prevents overfitting of data
self.dropout = nn.Dropout(0.2)
def forward(self, x):
# flatten image input
x = x.view(-1, 28 * 28)
# add hidden layer, with relu activation function
x = F.relu(self.fc1(x))
# add dropout layer
x = self.dropout(x)
# add hidden layer, with relu activation function
x = F.relu(self.fc2(x))
# add dropout layer
x = self.dropout(x)
# add output layer
x = self.fc3(x)
return x
# initialize the NN
model = Net()
print(model)
```
### Specify [Loss Function](http://pytorch.org/docs/stable/nn.html#loss-functions) and [Optimizer](http://pytorch.org/docs/stable/optim.html)
It's recommended that you use cross-entropy loss for classification. If you look at the documentation (linked above), you can see that PyTorch's cross entropy function applies a softmax funtion to the output layer *and* then calculates the log loss.
```
# specify loss function (categorical cross-entropy)
criterion = nn.CrossEntropyLoss()
# specify optimizer (stochastic gradient descent) and learning rate = 0.01
optimizer = torch.optim.SGD(model.parameters(), lr=0.01)
```
---
## Train the Network
The steps for training/learning from a batch of data are described in the comments below:
1. Clear the gradients of all optimized variables
2. Forward pass: compute predicted outputs by passing inputs to the model
3. Calculate the loss
4. Backward pass: compute gradient of the loss with respect to model parameters
5. Perform a single optimization step (parameter update)
6. Update average training loss
The following loop trains for 50 epochs; take a look at how the values for the training loss decrease over time. We want it to decrease while also avoiding overfitting the training data.
```
# number of epochs to train the model
n_epochs = 50
# initialize tracker for minimum validation loss
valid_loss_min = np.Inf # set initial "min" to infinity
for epoch in range(n_epochs):
# monitor training loss
train_loss = 0.0
valid_loss = 0.0
###################
# train the model #
###################
model.train() # prep model for training
for data, target in train_loader:
# clear the gradients of all optimized variables
optimizer.zero_grad()
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the loss
loss = criterion(output, target)
# backward pass: compute gradient of the loss with respect to model parameters
loss.backward()
# perform a single optimization step (parameter update)
optimizer.step()
# update running training loss
train_loss += loss.item()*data.size(0)
######################
# validate the model #
######################
model.eval() # prep model for evaluation
for data, target in valid_loader:
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the loss
loss = criterion(output, target)
# update running validation loss
valid_loss += loss.item()*data.size(0)
# print training/validation statistics
# calculate average loss over an epoch
train_loss = train_loss/len(train_loader.sampler)
valid_loss = valid_loss/len(valid_loader.sampler)
print('Epoch: {} \tTraining Loss: {:.6f} \tValidation Loss: {:.6f}'.format(
epoch+1,
train_loss,
valid_loss
))
# save model if validation loss has decreased
if valid_loss <= valid_loss_min:
print('Validation loss decreased ({:.6f} --> {:.6f}). Saving model ...'.format(
valid_loss_min,
valid_loss))
torch.save(model.state_dict(), 'model.pt')
valid_loss_min = valid_loss
```
### Load the Model with the Lowest Validation Loss
```
model.load_state_dict(torch.load('model.pt'))
```
---
## Test the Trained Network
Finally, we test our best model on previously unseen **test data** and evaluate it's performance. Testing on unseen data is a good way to check that our model generalizes well. It may also be useful to be granular in this analysis and take a look at how this model performs on each class as well as looking at its overall loss and accuracy.
```
# initialize lists to monitor test loss and accuracy
test_loss = 0.0
class_correct = list(0. for i in range(10))
class_total = list(0. for i in range(10))
model.eval() # prep model for evaluation
for data, target in test_loader:
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the loss
loss = criterion(output, target)
# update test loss
test_loss += loss.item()*data.size(0)
# convert output probabilities to predicted class
_, pred = torch.max(output, 1)
# compare predictions to true label
correct = np.squeeze(pred.eq(target.data.view_as(pred)))
# calculate test accuracy for each object class
for i in range(len(target)):
label = target.data[i]
class_correct[label] += correct[i].item()
class_total[label] += 1
# calculate and print avg test loss
test_loss = test_loss/len(test_loader.sampler)
print('Test Loss: {:.6f}\n'.format(test_loss))
for i in range(10):
if class_total[i] > 0:
print('Test Accuracy of %5s: %2d%% (%2d/%2d)' % (
str(i), 100 * class_correct[i] / class_total[i],
np.sum(class_correct[i]), np.sum(class_total[i])))
else:
print('Test Accuracy of %5s: N/A (no training examples)' % (classes[i]))
print('\nTest Accuracy (Overall): %2d%% (%2d/%2d)' % (
100. * np.sum(class_correct) / np.sum(class_total),
np.sum(class_correct), np.sum(class_total)))
```
### Visualize Sample Test Results
This cell displays test images and their labels in this format: `predicted (ground-truth)`. The text will be green for accurately classified examples and red for incorrect predictions.
```
# obtain one batch of test images
dataiter = iter(test_loader)
images, labels = dataiter.next()
# get sample outputs
output = model(images)
# convert output probabilities to predicted class
_, preds = torch.max(output, 1)
# prep images for display
images = images.numpy()
# plot the images in the batch, along with predicted and true labels
fig = plt.figure(figsize=(25, 4))
for idx in np.arange(20):
ax = fig.add_subplot(2, 20/2, idx+1, xticks=[], yticks=[])
ax.imshow(np.squeeze(images[idx]), cmap='gray')
ax.set_title("{} ({})".format(str(preds[idx].item()), str(labels[idx].item())),
color=("green" if preds[idx]==labels[idx] else "red"))
```
| github_jupyter |
```
# %load_ext autoreload
# %autoreload 2
import syft as sy
import numpy as np
import pandas as pd
domain = sy.login(
email="info@openmined.org",
password="changethis",
port=8081,
)
# Let's create an empty dataset
domain.create_dataset(
name="Empty Dataset",
description="Test delete",
)
# We can see that there are not existing assets attached to the dataset
domain.datasets
# Let's grab the dataset pointer
dataset_pointer = domain.datasets[0]
dataset_pointer
# Let's add a random asset to the dataset
dataset_pointer.add(name="random-asset", value=np.random.rand(100, 8))
# Let's check the dataset pointer
dataset_pointer
# Great !!! we have an asset attached to the dataset
# Let's try an asset with the same name again
dataset_pointer.add(name="random-asset", value=np.random.rand(50, 4))
# Great!!! So, we cannot add asset with the same name.
# Let's try and delete the asset
dataset_pointer.delete(name="random-asset")
# The dataset is deleted.
# let's check the dataset_pointer again
dataset_pointer
# We don't have any assets attached, which is what is expected.
# If we delete an asset that does not exists, we will get an error
dataset_pointer.delete("key-not-exists")
# Great, now that we can add/delete assets, we can upload the dataset
# in batches, let's try to do that.
```
Let's load some data... into memory
```
# Load the first 100 rows
ca_data = pd.read_csv("../../trade_demo/datasets/ca - feb 2021.csv")[:100]
# We will transfer data in batches of 10
start, end = 0, len(ca_data)
batch_size = 10
idx = 0
while(start < end):
idx += 1
batch = ca_data[start:start+batch_size] # Select a batch of images
name = f"ca_batch_{idx+1}" # Asset key name
dataset_pointer.add(name=name, value=batch) # Add asset to the dataset pointer
start += batch_size
print(f"Batches successfully uploaded: {idx}/{end//batch_size}")
# Great !!! we uploaded the whole data into batches of 10.
# Let's check the dataset pointer
dataset_pointer
# Let's check the dataset in the domain
domain.datasets
# Now, lastly, accessing the assets as an iterator
# We want to access the assets of this dataset as an iterator
dataset_ptr = domain.datasets[0] # Selected the MedNIST dataset
# We want an iterator and exclude certain assets.
data_iterator = dataset_ptr.iter(exclude=["ca_batch_2", "ca_batch_3", "ca_batch_4"])
cnt = 0
for d in data_iterator:
cnt += 1
print(d)
print(f"total assets in the iterator: {cnt}")
```
Since we excluded three assets, so we had pointers to the remaining seven assets.
```
# Finally, the delete dataset functionality is also fixed,
# So, let's delete the whole dataset
domain.datasets
# Deleting the dataset
del domain.datasets[0]
# There are no datasets in the domain
domain.datasets
```
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
from sklearn import datasets
iris = datasets.load_iris()
train_x, classes = iris.data, iris.target
# whiten features
train_x -= np.mean(train_x, axis=0)
train_x /= np.std(train_x, axis=0) + 1e-8
# convert target to one hot vectors
train_y = np.zeros((150, 3))
train_y[np.arange(150), classes] = 1
import keras
import keras.layers as layers
import keras.backend as K
from model import LogGaussMF
def train_model():
model = keras.Sequential([
layers.Dense(4, activation="relu", input_shape=(4,)),
LogGaussMF(3),
layers.Lambda(lambda x: K.sum(x, axis=-1)),
layers.Lambda(lambda x: K.exp(x - K.max(x, axis=-1, keepdims=True))),
layers.Dense(3),
layers.Activation("softmax")
])
model.compile(
optimizer=keras.optimizers.adam(lr=0.03),
loss="binary_crossentropy",
metrics=[keras.metrics.categorical_accuracy])
# train the model
history = model.fit(
x=train_x,
y=train_y,
batch_size=16,
epochs=200,
verbose=0,
shuffle=True)
print(model.evaluate(train_x, train_y))
# plot the loss and accuracy
fig, axes = plt.subplots(1, 2, figsize=(8, 4), squeeze=True)
axes[0].set_title("Loss")
axes[0].plot(history.history["loss"], c="b")
axes[1].set_title("Accuracy")
axes[1].plot(history.history["categorical_accuracy"], c="b")
return model
model = train_model()
def find_representatives(model):
pred_y = model.predict(train_x)
medoid_idxs = []
for i in range(3):
medoid_idxs.append(np.argmax(pred_y[:,i]))
return train_x[medoid_idxs]
medoids = find_representatives(model)
print(medoids)
import innvestigate
from keras.models import Model
def analyze_representative(model, representative, top_k=10):
assert(isinstance(model.layers[-5], LogGaussMF))
assert(isinstance(model.layers[-4], layers.Lambda))
assert(isinstance(model.layers[-3], layers.Lambda))
assert(isinstance(model.layers[-2], layers.Dense))
# outputs needed for analyzing the network
features_tensor = model.layers[-6].output
memberships_tensor = model.layers[-5].output
firing_strength_tensor = model.layers[-3].output
logits_tensor = model.layers[-2].output
# new model with the outputs from above
analysis_model = Model(model.inputs, [
features_tensor,
memberships_tensor,
firing_strength_tensor,
logits_tensor,
])
# compute the values of those tensors on the representative
features, memberships, firing_strength, logits = analysis_model.predict(
representative.reshape(1, 4))
# find and output the max logit
logit_val, logit_idx = np.max(logits), np.argmax(logits)
print("Logits: {}".format(logits))
print("Max Logit")
print("Value = {}, Index = {}\n".format(logit_val, logit_idx))
# find the rule that contributed most to the logit
# this is computed as firing strength multiplied by the weights
W = analysis_model.layers[-1].get_weights()[0]
R = W[:,logit_idx] * firing_strength
rule_val, rule_idx = np.max(R), np.argmax(R)
print("Firing Strength: {}".format(firing_strength))
print("Weights: {}".format(W[:,logit_idx]))
print("Most important rule")
print("Value = {}, Index = {}\n".format(rule_val, rule_idx))
# the maximum membership value does not appear to be a good indicator of the
# most important membership function. a large number of rules seem to share
# the maximum possible value of the membership function, 1.
mu, beta = analysis_model.layers[-4].get_weights()
R = np.exp(memberships[:,rule_idx,:]) * features
membership_val = np.max(R)
membership_idx = np.argmax(R)
print("Features: {}".format(features))
print("Memberships: {}".format(np.exp(memberships[:,rule_idx,:])))
print("Most important MF")
print("Value = {}, Index = {}\n".format(membership_val, membership_idx))
model_wo_softmax = Model(model.inputs, [model.layers[-6].output])
analyzer = innvestigate.create_analyzer(
"guided_backprop",
model_wo_softmax,
neuron_selection_mode="index",
allow_lambda_layers=True)
analysis = analyzer.analyze(
representative.reshape(1, 4),
membership_idx).squeeze()
print("Input Space Relevance")
print(analysis)
print("\n")
for medoid in medoids:
analyze_representative(model, medoid, top_k=1)
```
| github_jupyter |
<div class="alert alert-info">
*Objective of this sheet*
</div>
* concatenating
* merging
* reshaping (pivoting)
```
import pandas as pd
# 1 Concatenating : the pandas.concat( ) function concatenates the objects along an axis.
NZ_population = pd.DataFrame({
'city': ['Auckland','Wellington','Hamilton'],
'population' : [1534700,412500,235900],
'density': [1413,929,269]
})
NZ_population
AUS_population = pd.DataFrame({
'city': ['Sydney','Melbourne','Perth'],
'population' : [5029768,4725316,2022044],
'density': [1900 ,1500,1000 ]
})
AUS_population
# for concatenation we have to just pass the dataframe objects, here we have two, but even can be more then that
con = pd.concat([NZ_population, AUS_population])
con
# if we will have a look at the above index, we can see above the index are taken what were originally,
# now we will make it a continous index continous index
df = pd.concat([NZ_population, AUS_population], ignore_index = True)
df
# now to retrive a particular data from the dataframe, e.g I want to retreve Australia data
dif = pd.concat([NZ_population, AUS_population], keys=['NZ','AUS'])
dif
# now it has created an additional index for a subset of a datframe
# it will be helpful to retrieve a subset of a dataframe
dif.loc['AUS']
# so to get the orignal dataframe back it is a good choice
# If we just want to concatenate column wise,then
NZ_population = pd.DataFrame({
'city': ['Auckland','Wellington','Hamilton'],
'population' : [1534700,412500,235900]})
NZ_population
AUS_population = pd.DataFrame({
'city': ['Sydney','Melbourne','Perth'],
'population' : [5029768,4725316,2022044] })
AUS_population
# this is how we can concate columns wise
df1 = pd.concat([NZ_population,AUS_population], axis = 1)
df1
# we can also join the dataframe with a series, now down temperature_df is my dataframe
AUS_population
# now we are creating a series
s = pd.Series([1900 ,1500,1000 ], name = 'density')
s
#so down we have added series to the dataframe
pd.concat([AUS_population, s], axis = 1)
#2 MERGE DataFrame : the pandas.merge( ) function connects the rows in a DataFrame based on keys it implements join operations.
AUS_population = pd.DataFrame({
'city': ['Sydney','Melbourne','Perth'],
'population' : [5029768,4725316,2022044]})
AUS_population
AUS_density = pd.DataFrame({
'city': ['Sydney','Melbourne','Perth','Adelaide'],
'density': [1900 ,1500,1000,1300 ]})
AUS_density
# now we want to join these two dataframes by merge function
data = pd.merge(AUS_population,AUS_density ,right_on = 'city', left_on = 'city')
data
# it is better then concat, because it is looking the values in the cities and then merging
# inner join is by defauklt, we can also use outer, left or right as well
# in the same way here we have used the left join
data = pd.merge(AUS_population,AUS_density ,on = 'city', how = 'left')
data
# if we will write here the right join then Adelade will come.
# in the same way here we have used the outer join, here we are setting the data first.
NZ_population = pd.DataFrame({
'city': ['Auckland','Wellington','Hamilton'],
'population' : [1534700,412500,235900],
'density': [1413,929,269]
})
NZ_population
AUS_population = pd.DataFrame({
'city': ['Sydney','Melbourne','Perth'],
'population' : [5029768,4725316,2022044],
'density': [1900 ,1500,1000 ]
})
AUS_population
#df3 = pd.merge(NZ_population,AUS_population,on='city', how = 'outer')
## here suffixes will come handy, it will distinguish between the two columns
df3 = pd.merge(NZ_population,AUS_population,on='city', how = 'outer', suffixes=('_left','_right'))
df3
#3 Pivot tables
df =pd.read_excel('pivo.xlsx')
df
# pivot functions arguments
df2 = df.pivot_table(index='date', columns='city', values = 'humidity')
df2
# we can see three things written above i.e. index, columns and values:
# index : what I want on 'X' axis or as rows?
# columns: what do I want to present in the columns?
# values: what data I want to present?
# mean is the default function that comes!
# In pivot method: the columns shold have most number of unique values.
# condensing the dataframe by aligning the common values.
# Why pivot table? Pivot table is used to summarize and aggregate data inside dataframe
df =pd.read_excel('pitab.xlsx')
df
# we have aggregated the input dataframe into meaningful output: we can use any number of functions
df.pivot_table(index='city', columns = 'date', aggfunc='mean')
# aggfunc can be anything like mean,sum, median etc
# seing another example of using the pivot in Python.
import seaborn as sns
# loading the dataset for the using purpose. This entire data is related to the TITANIC ship.
titanic = sns.load_dataset('titanic')
# How many people survived the attack?
titanic.pivot_table(values = 'survived', index='sex', columns='class', aggfunc = 'mean')
# MULTILEVEL PIVOT TABLES : cut here means bining
# binning : Binning or discretization is the process of transforming numerical variables into categorical counterparts.
# An example is to bin values for Age into categories such as 20-39, 40-59, and 60-79. Numerical variables are usually
#discretized in the modeling methods
age = pd.cut(titanic['age'], [0,18,80])
titanic.pivot_table(values = 'survived', index = ['sex', age], columns = ['class'])
```
### Exercise
##### Q1 Please rerun the cells and try to understand the logic.
##### Q2 Please tell the difference between pivot and the groupby. When which technique should be used?
###### Q3 What is the default join of the concatenation?
| github_jupyter |
# Dynamics 365 Business Central Trouble Shooting Guide (TSG) - Extensions
This notebook contains Kusto queries that can help getting to the root cause of an issue with extensions for one or more environments.
NB! Some of the signal used in this notebook is only available in newer versions of Business Central, so check the version of your environment if some sections do not return any data. The signal documentation states in which version a given signal was introduced.
## 1\. Get setup: Load up Python libraries and connect to Application Insights
First you need to set the notebook Kernel to Python3, load the KQLmagic module (did you install it?) and connect to your Application Insights resource (get appid and appkey from the API access page in the Application Insights portal)
```
# load the KQLmagic module
%reload_ext Kqlmagic
# Connect to the Application Insights API
%kql appinsights://appid='<add app id from the Application Insights portal>';appkey='<add API key from the Application Insights portal>'
```
## 2\. Define filters
This workbook is designed for troubleshooting extensions. Please provide values for aadTenantId, environmentName, and extensionId (or use a config file).
You can also specify limits to the period of time that the analysis should include.
```
# Add values for AAD tenant id, environment name, and extension id.
# It is possible to leave one or more values blank (if you want to analyze across all values of the parameter)
# You can either use configuration file (INI file format) or set filters directly.
# If you specify a config file, then variables set here takes precedence over manually set filter variables
# config file name and directory (full path)
configFile = "c:/tmp/notebook.ini"
# Add AAD tenant id and environment name here
aadTenantId = "MyaaDtenantId"
environmentName = "MyEnvironmentName"
extensionId = "MyExtensionId"
# date filters for the analysis
# use YYYY-MM-DD format for the dates (ISO 8601)
startDate = "2020-11-20"
endDate = "2020-11-24"
# Do not edit this code section
import configparser
config = configparser.ConfigParser()
config.read(configFile)
if bool(config.defaults()):
if config.has_option('DEFAULT', 'aadTenantId'):
aadTenantId = config['DEFAULT']['aadTenantId']
if config.has_option('DEFAULT', 'environmentName'):
environmentName = config['DEFAULT']['environmentName']
if config.has_option('DEFAULT', 'extensionId'):
extensionId = config['DEFAULT']['extensionId']
if config.has_option('DEFAULT', 'startDate'):
startDate = config['DEFAULT']['startDate']
if config.has_option('DEFAULT', 'endDate'):
endDate = config['DEFAULT']['endDate']
print("Using these parameters for the analysis:")
print("----------------------------------------")
print("aadTenantId " + aadTenantId)
print("environmentName " + environmentName)
print("extensionId " + extensionId)
print("startDate " + startDate)
print("endDate " + endDate)
```
# Analyze extension events
Now you can run Kusto queries to look for possible root causes for issues about extensions.
Either click **Run All** above to run all sections, or scroll down to the type of analysis you want to do and manually run queries
## Extension event overview
Event telemetry docs:
* https://docs.microsoft.com/en-us/dynamics365/business-central/dev-itpro/administration/telemetry-extension-lifecycle-trace
* https://docs.microsoft.com/en-us/dynamics365/business-central/dev-itpro/administration/telemetry-extension-update-trace
KQL samples: https://github.com/microsoft/BCTech/blob/master/samples/AppInsights/KQL/RawData/ExtensionLifecycle.kql
```
%%kql
//
// extension event types stats
//
let _aadTenantId = aadTenantId;
let _environmentName = environmentName;
let _extensionId = extensionId;
let _startDate = startDate;
let _endDate = endDate;
traces
| where 1==1
and timestamp >= todatetime(_startDate)
and timestamp <= todatetime(_endDate) + totimespan(24h) - totimespan(1ms)
and (_aadTenantId == '' or customDimensions.aadTenantId == _aadTenantId)
and (_environmentName == '' or customDimensions.environmentName == _environmentName )
and (_extensionId == '' or customDimensions.extensionId == _extensionId)
and customDimensions.eventId in ('RT0010', 'LC0010', 'LC0011', 'LC0012', 'LC0013', 'LC0014', 'LC0015', 'LC0016', 'LC0017', 'LC0018', 'LC0019', 'LC020', 'LC021', 'LC022', 'LC023')
| extend aadTenantId=tostring( customDimensions.aadTenantId)
, environmentName=tostring( customDimensions.environmentName )
, extensionId=tostring( customDimensions.extensionId )
, eventId=tostring(customDimensions.eventId)
| extend eventMessageShort= strcat( case(
eventId=='RT0010', 'Update failed (upgrade code)'
, eventId=='LC0011', 'Install failed'
, eventId=='LC0012', 'Synch succeeded'
, eventId=='LC0013', 'Synch failed'
, eventId=='LC0014', 'Publish succeeded'
, eventId=='LC0015', 'Publish failed'
, eventId=='LC0016', 'Un-install succeeded'
, eventId=='LC0017', 'Un-install failed'
, eventId=='LC0018', 'Un-publish succeeded'
, eventId=='LC0019', 'Un-publish failed'
, eventId=='LC0020', 'Compilation succeeded'
, eventId=='LC0021', 'Compilation failed'
, eventId=='LC0022', 'Update succeeded'
, eventId=='LC0023', 'Update failed (other)'
, 'Unknown message'
), " (", eventId, ')' )
| summarize count=count() by eventType=eventMessageShort
| order by eventType
| render barchart with (title='Extension lifecycle event overview', legend=hidden)
%%kql
//
// top 100 extension events
//
let _aadTenantId = aadTenantId;
let _environmentName = environmentName;
let _extensionId = extensionId;
let _startDate = startDate;
let _endDate = endDate;
traces
| where 1==1
and timestamp >= todatetime(_startDate)
and timestamp <= todatetime(_endDate) + totimespan(24h) - totimespan(1ms)
and (_aadTenantId == '' or customDimensions.aadTenantId == _aadTenantId)
and (_environmentName == '' or customDimensions.environmentName == _environmentName )
and (_extensionId == '' or customDimensions.extensionId == _extensionId)
and customDimensions.eventId in ('RT0010', 'LC0010', 'LC0011', 'LC0012', 'LC0013', 'LC0014', 'LC0015', 'LC0016', 'LC0017', 'LC0018', 'LC0019', 'LC020', 'LC021', 'LC022', 'LC023')
| extend aadTenantId=tostring( customDimensions.aadTenantId)
, environmentName=tostring( customDimensions.environmentName )
, extensionId=tostring( customDimensions.extensionId )
, extensionName=tostring( customDimensions.extensionName )
, eventId=tostring(customDimensions.eventId)
| extend eventMessageShort= strcat( case(
eventId=='RT0010', 'Update failed (upgrade code)'
, eventId=='LC0011', 'Install failed'
, eventId=='LC0012', 'Synch succeeded'
, eventId=='LC0013', 'Synch failed'
, eventId=='LC0014', 'Publish succeeded'
, eventId=='LC0015', 'Publish failed'
, eventId=='LC0016', 'Un-install succeeded'
, eventId=='LC0017', 'Un-install failed'
, eventId=='LC0018', 'Un-publish succeeded'
, eventId=='LC0019', 'Un-publish failed'
, eventId=='LC0020', 'Compilation succeeded'
, eventId=='LC0021', 'Compilation failed'
, eventId=='LC0022', 'Update succeeded'
, eventId=='LC0023', 'Update failed (other)'
, 'Unknown message'
), " (", eventId, ')' )
| project timestamp, eventMessageShort, extensionName, aadTenantId, environmentName, extensionId
| order by aadTenantId, environmentName, extensionId, timestamp asc
| limit 100
```
## Extension failures
Event telemetry docs:
* https://docs.microsoft.com/en-us/dynamics365/business-central/dev-itpro/administration/telemetry-extension-lifecycle-trace
* https://docs.microsoft.com/en-us/dynamics365/business-central/dev-itpro/administration/telemetry-extension-update-trace
```
%%kql
//
// extension event failure overview
//
let _aadTenantId = aadTenantId;
let _environmentName = environmentName;
let _extensionId = extensionId;
let _startDate = startDate;
let _endDate = endDate;
traces
| where 1==1
and timestamp >= todatetime(_startDate)
and timestamp <= todatetime(_endDate) + totimespan(24h) - totimespan(1ms)
and (_aadTenantId == '' or customDimensions.aadTenantId == _aadTenantId)
and (_environmentName == '' or customDimensions.environmentName == _environmentName )
and (_extensionId == '' or customDimensions.extensionId == _extensionId)
and customDimensions.eventId in ('RT0010', 'LC0011', 'LC0013', 'LC0015', 'LC0017', 'LC0019', 'LC021', 'LC023')
| extend aadTenantId=tostring( customDimensions.aadTenantId)
, environmentName=tostring( customDimensions.environmentName )
, extensionId=tostring( customDimensions.extensionId )
, eventId=tostring(customDimensions.eventId)
| extend eventMessageShort= strcat( case(
eventId=='RT0010', 'Update failed (upgrade code)'
, eventId=='LC0011', 'Install failed'
, eventId=='LC0013', 'Synch failed'
, eventId=='LC0015', 'Publish failed'
, eventId=='LC0017', 'Un-install failed'
, eventId=='LC0019', 'Un-publish failed'
, eventId=='LC0021', 'Compilation failed'
, eventId=='LC0023', 'Update failed (other)'
, 'Unknown message'
), " (", eventId, ')' )
| summarize count=count() by eventType=eventMessageShort
| order by eventType
| render barchart with (title='Failure type overview', xtitle="", legend=hidden)
%%kql
//
// top 100 latest extension event failure details
//
let _aadTenantId = aadTenantId;
let _environmentName = environmentName;
let _extensionId = extensionId;
let _startDate = startDate;
let _endDate = endDate;
traces
| where 1==1
and timestamp >= todatetime(_startDate)
and timestamp <= todatetime(_endDate) + totimespan(24h) - totimespan(1ms)
and (_aadTenantId == '' or customDimensions.aadTenantId == _aadTenantId)
and (_environmentName == '' or customDimensions.environmentName == _environmentName )
and (_extensionId == '' or customDimensions.extensionId == _extensionId)
and customDimensions.eventId in ('RT0010', 'LC0011', 'LC0013', 'LC0015', 'LC0017', 'LC0019', 'LC021', 'LC023')
| extend aadTenantId=tostring( customDimensions.aadTenantId)
, environmentName=tostring( customDimensions.environmentName )
, extensionId=tostring( customDimensions.extensionId )
, eventId=tostring(customDimensions.eventId)
, extensionName=tostring(customDimensions.extensionName)
| extend eventMessageShort= strcat( case(
eventId=='RT0010', 'Update failed (upgrade code)'
, eventId=='LC0011', 'Install failed'
, eventId=='LC0013', 'Synch failed'
, eventId=='LC0015', 'Publish failed'
, eventId=='LC0017', 'Un-install failed'
, eventId=='LC0019', 'Un-publish failed'
, eventId=='LC0021', 'Compilation failed'
, eventId=='LC0023', 'Update failed (other)'
, 'Unknown message'
), " (", eventId, ')' )
| project timestamp, extensionName, eventType=eventMessageShort
, version=customDimensions.extensionVersion
, failureReason=customDimensions.failureReason
, aadTenantId, environmentName, extensionId
| order by timestamp desc
| limit 100
%%kql
//
// top 20 latest update failures (due to upgrade code)
//
let _aadTenantId = aadTenantId;
let _environmentName = environmentName;
let _extensionId = extensionId;
let _startDate = startDate;
let _endDate = endDate;
traces
| where 1==1
and timestamp >= todatetime(_startDate)
and timestamp <= todatetime(_endDate) + totimespan(24h) - totimespan(1ms)
and (_aadTenantId == '' or customDimensions.aadTenantId == _aadTenantId)
and (_environmentName == '' or customDimensions.environmentName == _environmentName )
and (_extensionId == '' or customDimensions.extensionId == _extensionId)
and customDimensions.eventId == 'RT0010'
| extend aadTenantId=tostring( customDimensions.aadTenantId)
, environmentName=tostring( customDimensions.environmentName )
, extensionId=tostring( customDimensions.extensionId )
, eventId=tostring(customDimensions.eventId)
, extensionName=tostring(customDimensions.extensionName)
| project timestamp, extensionName
, version=customDimensions.extensionVersion
, targetedVersion =customDimensions.extensionTargetedVersion
, failureType =customDimensions.failureType
, alStackTrace =customDimensions.alStackTrace
, companyName = customDimensions.companyName
, extensionPublisher = customDimensions.extensionPublisher
, aadTenantId, environmentName, extensionId
| order by timestamp desc
| limit 20
%%kql
//
// top 20 latest synch failures
//
let _aadTenantId = aadTenantId;
let _environmentName = environmentName;
let _extensionId = extensionId;
let _startDate = startDate;
let _endDate = endDate;
traces
| where 1==1
and timestamp >= todatetime(_startDate)
and timestamp <= todatetime(_endDate) + totimespan(24h) - totimespan(1ms)
and (_aadTenantId == '' or customDimensions.aadTenantId == _aadTenantId)
and (_environmentName == '' or customDimensions.environmentName == _environmentName )
and (_extensionId == '' or customDimensions.extensionId == _extensionId)
and customDimensions.eventId == 'LC0013'
| extend aadTenantId=tostring( customDimensions.aadTenantId)
, environmentName=tostring( customDimensions.environmentName )
, extensionId=tostring( customDimensions.extensionId )
, eventId=tostring(customDimensions.eventId)
, extensionName=tostring(customDimensions.extensionName)
| project timestamp, extensionName
, version=customDimensions.extensionVersion
, failureReason=customDimensions.failureReason
, publishedAs = customDimensions.extensionPublishedAs
, extensionPublisher = customDimensions.extensionPublisher
, extensionScope = customDimensions.extensionScope
, extensionSynchronizationMode = customDimensions.extensionSynchronizationMode
, aadTenantId, environmentName, extensionId
| order by timestamp desc
| limit 20
%%kql
//
// top 20 latest compilation failures
//
let _aadTenantId = aadTenantId;
let _environmentName = environmentName;
let _extensionId = extensionId;
let _startDate = startDate;
let _endDate = endDate;
traces
| where 1==1
and timestamp >= todatetime(_startDate)
and timestamp <= todatetime(_endDate) + totimespan(24h) - totimespan(1ms)
and (_aadTenantId == '' or customDimensions.aadTenantId == _aadTenantId)
and (_environmentName == '' or customDimensions.environmentName == _environmentName )
and (_extensionId == '' or customDimensions.extensionId == _extensionId)
and customDimensions.eventId == 'LC0021'
| extend aadTenantId=tostring( customDimensions.aadTenantId)
, environmentName=tostring( customDimensions.environmentName )
, extensionId=tostring( customDimensions.extensionId )
, eventId=tostring(customDimensions.eventId)
, extensionName=tostring(customDimensions.extensionName)
| project timestamp, extensionName
, version=customDimensions.extensionVersion
, failureReason=customDimensions.failureReason
, compilationResult = customDimensions.extensionCompilationResult
, compilationDependencyList = customDimensions.extensionCompilationDependencyList
, publisher = customDimensions.extensionPublisher
, publishedAs = customDimensions.extensionPublishedAs
, extensionScope = customDimensions.extensionScope
, aadTenantId, environmentName, extensionId
| order by timestamp desc
| limit 20
```
| github_jupyter |
# Solve a Generalized Assignment Problem using Lagrangian relaxation
This tutorial includes data and information that you need to set up decision optimization engines and build mathematical programming models to solve a Generalized Assignment Problem using Lagrangian relaxation.
When you finish this tutorial, you'll have a foundational knowledge of _Prescriptive Analytics_.
>This notebook is part of the [Prescriptive Analytics for Python](https://rawgit.com/IBMDecisionOptimization/docplex-doc/master/docs/index.html)
>You will need a valid subscription to Decision Optimization on Cloud ([here](https://developer.ibm.com/docloud)) or a local installation of CPLEX Optimizers.
Some familiarity with Python is recommended. This notebook runs on Python 2.
## Table of contents
* [Describe the business problem](#describe-problem)
* [How Decision Optimization can help](#do-help)
* [Use Decision Optimization to create and solve the model](#do-model-create-solve)
* [Summary](#summary)<br>
## Describe the business problem
This notebook illustrates how to solve an optimization model using Lagrangian relaxation technics.
It solves a generalized assignment problem (GAP), as defined by Wolsey, using this relaxation technic.
The main aim is to show multiple optimization through modifications of different models existing in a single environment, not to show how to solve a GAP problem.
In the field of Mathematical Programming, this technic consists in the approximation of a difficult constrained problem by a simpler problem: you remove difficult constraints by integrating them in the objective function, penalizing it if the constraint is not respected.
The method penalizes violations of inequality constraints using a Lagrange multiplier, which imposes a cost on violations. These added costs are used instead of the strict inequality constraints in the optimization. In practice, this relaxed problem can often be solved more easily than the original problem.
For more information, see the following Wikipedia articles: [Generalized assignment problem](https://en.wikipedia.org/wiki/Generalized_assignment_problem) and [Lagrangian relaxation](https://en.wikipedia.org/wiki/Lagrangian_relaxation).
This notebook first solves the standard problem (which is not important here), then shows how to reformulate it to meet the Lagrangian Relaxation features.
## How decision optimization can help
Prescriptive analytics (decision optimization) technology recommends actions that are based on desired outcomes. It considers specific scenarios, resources, and knowledge of past and current events. With this insight, your organization can make better decisions and have greater control over business outcomes.
Prescriptive analytics is the next step on the path to insight-based actions. It creates value through synergy with predictive analytics, which analyzes data to predict future outcomes. Prescriptive analytics takes that insight to the next level by suggesting the optimal way to handle a future situation. Organizations that act fast in dynamic conditions and make superior decisions in uncertain environments gain a strong competitive advantage.
With prescriptive analytics, you can:
* Automate the complex decisions and trade-offs to better manage your limited resources.
* Take advantage of a future opportunity or mitigate a future risk.
* Proactively update recommendations based on changing events.
* Meet operational goals, increase customer loyalty, prevent threats and fraud, and optimize business processes.
## Use Decision Optimization
Perform the following steps to create and solve the model.
1. [Download the library](#Step-1:-Download-the-library)<br>
2. [Set up the engines](#Step-2:-Set-up-the-prescriptive-engine)<br>
3. [Model the Data](#Step-3:-Model-the-data)<br>
4. [Prepare the data](#Step-4:-Prepare-the-data)<br>
5. [Set up the prescriptive model](#Step-5:-Set-up-the-prescriptive-model)<br>
5.1 [Define the decision variables](#Define-the-decision-variables)<br>
5.2 [Express the business constraints](#Express-the-business-constraints)<br>
5.3 [Express the objective](#Express-the-objective)<br>
5.4 [Solve with the Decision Optimization solve service](#Solve-with-the-Decision-Optimization-solve-service)<br>
6. [Investigate the solution and run an example analysis](#Step-6:-Investigate-the-solution-and-then-run-an-)<br>
### 1. Download the library
Run the following code to install the Decision Optimization CPLEX Modeling library. The *DOcplex* library contains two modeling packages, mathematical programming (docplex.mp) package and constraint programming (docplex.cp) package.
In the following code, `real.prefix` is used to ensure that the script is not running inside a virtual environment.
```
import sys
try:
import docplex.mp
except:
if hasattr(sys, 'real_prefix'):
!pip install docplex
else:
!pip install --user docplex
```
### 2. Set up the prescriptive engine
To solve DOcplex models, you need access to the DOcplexcloud solve service.
* Subscribe to the [Decision Optimization on Cloud](https://developer.ibm.com/docloud) (DOcplexcloud) service.
* Get the service base URL and personal API key.
* Enter the URL and API key in the cell below, enclosed in quotation marks (""), and run the cell.
```
url = None
key = None
```
### 3. Model the data
In this scenario, the data is simple. It is delivered as 3 input arrays: A, B, and C. The data does not need changing or refactoring.
```
B = [15, 15, 15]
C = [
[ 6, 10, 1],
[12, 12, 5],
[15, 4, 3],
[10, 3, 9],
[8, 9, 5]
]
A = [
[ 5, 7, 2],
[14, 8, 7],
[10, 6, 12],
[ 8, 4, 15],
[ 6, 12, 5]
]
```
### 4. Set up the prescriptive model
Start with viewing the environment information. This information should be updated when you run the notebook.
```
from docplex.mp.environment import Environment
env = Environment()
env.print_information()
```
We will firt create an optimization problem, composed of 2 basic constraints blocks, then we will resolve it using Lagrangian Relaxation on 1 of the constraints block.
#### 4.1 Create the DOcplex model
The model contains the business constraints and the objective.
```
from docplex.mp.model import Model
mdl = Model("GAP per Wolsey")
```
#### 4.2 Define the decision variables
```
print("#As={}, #Bs={}, #Cs={}".format(len(A), len(B), len(C)))
number_of_cs = len(C)
# variables
x_vars = [mdl.binary_var_list(c, name=None) for c in C]
```
#### 4.3 Define the business constraints
```
# constraints
cts = mdl.add_constraints(mdl.sum(xv) <= 1 for xv in x_vars)
mdl.add_constraints(mdl.sum(x_vars[ii][j] * A[ii][j] for ii in range(number_of_cs)) <= bs for j, bs in enumerate(B))
# objective
total_profit = mdl.sum(mdl.scal_prod(x_i, c_i) for c_i, x_i in zip(C, x_vars))
mdl.maximize(total_profit)
mdl.print_information()
```
#### 4.4. Solve the model
Use the Decision Optimization on Cloud solve service to solve the model or a local engine.
```
s = mdl.solve(url=url, key=key)
assert s is not None
obj = s.objective_value
print("* GAP with no relaxation run OK, best objective is: {:g}".format(obj))
```
#### 4.5. Solve the model with Lagrangian Relaxation method
Let's consider for the demonstration of the Lagrangian Relaxation that this model was hard to solve for CPLEX.
We will approximate this problem by doing an iterative model, where the objective is modified at each iteration.
(Wait a few seconds for the solution, due to a time limit parameter.)
We first remove the culprit constraints from the model
```
for ct in cts:
mdl.remove_constraint(ct)
#p_vars are the penalties attached to violating the constraints
p_vars = mdl.continuous_var_list(C, name='p') # new for relaxation
# new version of the approximated constraint where we apply the penalties
mdl.add_constraints(mdl.sum(xv) == 1 - pv for xv, pv in zip(x_vars, p_vars))
;
#Define the maximum number of iterations
max_iters = 10
number_of_cs = len(C)
c_range = range(number_of_cs)
# Langrangian relaxation loop
eps = 1e-6
loop_count = 0
best = 0
initial_multiplier = 1
multipliers = [initial_multiplier] * len(C)
# Objective function
# I'd write the key perfromance indicator (kpi) as
# total_profit = mdl.sum(mdl.sum(x_vars[task][worker] * C[task][worker]) for task, worker in zip(tasks, workers))
total_profit = mdl.sum(mdl.scal_prod(x_i, c_i) for c_i, x_i in zip(C, x_vars))
mdl.add_kpi(total_profit, "Total profit")
print("starting the loop")
while loop_count <= max_iters:
loop_count += 1
# Rebuilt at each loop iteration
total_penalty = mdl.scal_prod(p_vars, multipliers)
mdl.maximize(total_profit + total_penalty)
s = mdl.solve(url=url, key=key)
if not s:
print("*** solve fails, stopping at iteration: %d" % loop_count)
break
best = s.objective_value
penalties = [pv.solution_value for pv in p_vars]
print('%d> new lagrangian iteration:\n\t obj=%g, m=%s, p=%s' % (loop_count, best, str(multipliers), str(penalties)))
do_stop = True
justifier = 0
for k in c_range:
penalized_violation = penalties[k] * multipliers[k]
if penalized_violation >= eps:
do_stop = False
justifier = penalized_violation
break
if do_stop:
print("* Lagrangian relaxation succeeds, best={:g}, penalty={:g}, #iterations={}"
.format(best, total_penalty.solution_value, loop_count))
break
else:
# Update multipliers and start the loop again.
scale_factor = 1.0 / float(loop_count)
multipliers = [max(multipliers[i] - scale_factor * penalties[i], 0.) for i in c_range]
print('{0}> -- loop continues, m={1!s}, justifier={2:g}'.format(loop_count, multipliers, justifier))
print(best)
```
### 6. Investigate the solution and run an example analysis
You can see that with this relaxation method applied to this simple model, we find the same solution to the problem.
## Summary
You learned how to set up and use IBM Decision Optimization CPLEX Modeling for Python to formulate a Constraint Programming model and solve it with IBM Decision Optimization on Cloud.
## References
* [CPLEX Modeling for Python documentation](https://rawgit.com/IBMDecisionOptimization/docplex-doc/master/docs/index.html)
* [Decision Optimization on Cloud](https://developer.ibm.com/docloud/)
* [Decision Optimization documentation](https://datascience.ibm.com/docs/content/DO/DOinDSX.html)
* For help with DOcplex, or to report a defect, go [here](https://developer.ibm.com/answers/smartspace/docloud).
* Contact us at dofeedback@wwpdl.vnet.ibm.com
<hr>
Copyright © IBM Corp. 2017. Released as licensed Sample Materials.
| github_jupyter |
# 02 Continuing the Introduction
For a simpler introduction to `pycox` see the [01_introduction.ipynb](https://nbviewer.jupyter.org/github/havakv/pycox/blob/master/examples/01_introduction.ipynb) instead.
In this notebook we will show some more functionality of the `pycox` package in addition to more functionality of the `torchtuples` package.
We will continue with the `LogisticHazard` method for simplicity.
We will in the following:
- use SUPPORT as an example dataset,
- use entity embeddings for categorical variables,
- use the [AdamWR optimizer](https://arxiv.org/pdf/1711.05101.pdf) with cyclical learning rates,
- use the scheme proposed by [Smith 2017](https://arxiv.org/pdf/1506.01186.pdf) to find a suitable learning rate.
```
import numpy as np
import matplotlib.pyplot as plt
from sklearn.preprocessing import StandardScaler
from sklearn_pandas import DataFrameMapper
import torch
import torchtuples as tt
from pycox.datasets import support
from pycox.preprocessing.feature_transforms import OrderedCategoricalLong
from pycox.models import LogisticHazard
from pycox.evaluation import EvalSurv
np.random.seed(123456)
_ = torch.manual_seed(123456)
```
## Dataset
We load the SUPPORT data set and split in train, test and validation.
```
df_train = support.read_df()
df_test = df_train.sample(frac=0.2)
df_train = df_train.drop(df_test.index)
df_val = df_train.sample(frac=0.2)
df_train = df_train.drop(df_val.index)
df_train.head()
```
## Feature transforms
We have 14 covariates, in addition to the durations and event indicators.
We will standardize the 8 numerical covariates, and leave the 3 binary variables unaltered.
We will use entity embedding for the 3 categorical variables `x2`, `x3`, and `x6`.
Hence, they are transformed to `int64` integers representing the categories. The category 0 is reserved for `None` and very small categories that are set to `None`.
We use the `OrderedCategoricalLong` transform to achieve this.
```
cols_standardize = ['x0', 'x7', 'x8', 'x9', 'x10', 'x11', 'x12', 'x13']
cols_leave = ['x1', 'x4', 'x5']
cols_categorical = ['x2', 'x3', 'x6']
standardize = [([col], StandardScaler()) for col in cols_standardize]
leave = [(col, None) for col in cols_leave]
categorical = [(col, OrderedCategoricalLong()) for col in cols_categorical]
x_mapper_float = DataFrameMapper(standardize + leave)
x_mapper_long = DataFrameMapper(categorical) # we need a separate mapper to ensure the data type 'int64'
x_fit_transform = lambda df: tt.tuplefy(x_mapper_float.fit_transform(df), x_mapper_long.fit_transform(df))
x_transform = lambda df: tt.tuplefy(x_mapper_float.transform(df), x_mapper_long.transform(df))
x_train = x_fit_transform(df_train)
x_val = x_transform(df_val)
x_test = x_transform(df_test)
```
In the `x_fit_transform` and `x_transform` we have wrapped the results with `tt.tuplefy`. The result is a `TupleTree` which equivalent to a regular `tuple`, but with some added functionality that makes it easier to investigate the data.
From the code below we see that `x_train` is a `tuple` with two arrays representing the transformed numerical and categorical covariates.
```
x_train.types()
x_train.shapes()
x_train.dtypes()
```
## Label transforms
`LogisticHazard` is a discrete-time method, meaning it requires discretization of the event times to be applied to continuous-time data.
We let `num_durations` define the size of the discretization grid, but we will now let the **quantiles** of the estimated event-time distribution define the grid, as explained in [this paper](https://arxiv.org/abs/1910.06724).
```
num_durations = 20
scheme = 'quantiles'
labtrans = LogisticHazard.label_transform(num_durations, scheme)
get_target = lambda df: (df['duration'].values, df['event'].values)
y_train = labtrans.fit_transform(*get_target(df_train))
y_val = labtrans.transform(*get_target(df_val))
durations_test, events_test = get_target(df_test)
```
Note that the discretization grid is far from equidistant. The idea behind the quantile discretization is that the grid is finer where there are many events and coarser where there are few.
```
labtrans.cuts
```
We can visualize the grid together with the Kaplan-Meier estaimtes to see this clearly.
```
from pycox.utils import kaplan_meier
plt.vlines(labtrans.cuts, 0, 1, colors='gray', linestyles="--", label='Discretization Grid')
kaplan_meier(*get_target(df_train)).plot(label='Kaplan-Meier')
plt.ylabel('S(t)')
plt.legend()
_ = plt.xlabel('Time')
```
## Investigating the data
Next we collect the training and validation data with `tt.tuplefy` in a nested tuple to make it simpler to inspect them
```
train = tt.tuplefy(x_train, y_train)
val = tt.tuplefy(x_val, y_val)
train.types()
train.shapes()
train.dtypes()
```
We can now alternatively transform the data to torch tensors with `to_tensor`. This is not useful for this notebook, but can be very handy for development.
```
train_tensor = train.to_tensor()
train_tensor.types()
train_tensor.shapes()
train_tensor.dtypes()
del train_tensor
```
## Neural net
We want our network to take two input arguments, one for the numerical covariates and one for the categorical covariates such that we can apply entity embedding.
The `tt.practical.MixedInputMLP` does exactly this for us. If first applies entity embeddings to the categorical covariates and then concatenate the embeddings with the numerical covariates.
First we need to define the embedding sizes. Here we will let the embedding dimensions be half the size of the number of categories.
This means that each category is represented by a vector that is half the size of the number of categories.
```
num_embeddings = x_train[1].max(0) + 1
embedding_dims = num_embeddings // 2
num_embeddings, embedding_dims
```
We then define a net with four hidden layers, each of size 32, and include batch normalization and dropout between each layer.
```
in_features = x_train[0].shape[1]
out_features = labtrans.out_features
num_nodes = [32, 32, 32, 32]
batch_norm = True
dropout = 0.2
net = tt.practical.MixedInputMLP(in_features, num_embeddings, embedding_dims,
num_nodes, out_features, batch_norm, dropout)
net
```
## Fitting model
We want to use the cyclic [AdamWR optimizer](https://arxiv.org/abs/1711.05101) where we multiply the learning rate with 0.8 and double then cycle length after every cycle.
Also, we add [decoupled weight decay](https://arxiv.org/abs/1711.05101) for regularization.
```
optimizer = tt.optim.AdamWR(decoupled_weight_decay=0.01, cycle_eta_multiplier=0.8,
cycle_multiplier=2)
model = LogisticHazard(net, optimizer, duration_index=labtrans.cuts)
```
We can use `lr_finder` to find a suitable initial learning rate
with the scheme proposed by [Smith 2017](https://arxiv.org/pdf/1506.01186.pdf).
See [this post](https://towardsdatascience.com/finding-good-learning-rate-and-the-one-cycle-policy-7159fe1db5d6) for an explanation.
The `tolerance` argument just defines the largest loss allowed before terminating the procedure. Is serves mostly a visual purpose.
```
batch_size = 256
lrfind = model.lr_finder(x_train, y_train, batch_size, tolerance=50)
_ = lrfind.plot()
```
We can see that this sets the optimizer learning rate in our model to the same value as that of `get_best_lr`.
```
model.optimizer.param_groups[0]['lr']
lrfind.get_best_lr()
```
However, we have found that `get_best_lr` sometimes gives a little high learning rate, so we instead set it to
```
model.optimizer.set_lr(0.02)
```
For early stopping, we will use `EarlyStoppingCycle` which work in the same manner as `EarlyStopping` but will stop at the end of **the cycle** if the current best model was not obtained in the current cycle.
```
epochs = 512
callbacks = [tt.callbacks.EarlyStoppingCycle()]
verbose = False # set to True if you want printout
%%time
log = model.fit(x_train, y_train, batch_size, epochs, callbacks, verbose,
val_data=val)
_ = log.to_pandas().iloc[1:].plot()
```
We can now plot the learning rates used through the training with following piece of code
```
lrs = model.optimizer.lr_scheduler.to_pandas() * model.optimizer.param_groups[0]['initial_lr']
lrs.plot()
plt.grid(linestyle='--')
```
## Evaluation
The `LogisticHazard` method has two implemented interpolation schemes: the constant density interpolation (default) and constant hazard interpolation. See [this paper](https://arxiv.org/abs/1910.06724) for details.
```
surv_cdi = model.interpolate(100).predict_surv_df(x_test)
surv_chi = model.interpolate(100, 'const_hazard').predict_surv_df(x_test)
ev_cdi = EvalSurv(surv_cdi, durations_test, events_test, censor_surv='km')
ev_chi = EvalSurv(surv_chi, durations_test, events_test, censor_surv='km')
ev_cdi.concordance_td(), ev_chi.concordance_td()
time_grid = np.linspace(durations_test.min(), durations_test.max(), 100)
ev_cdi.integrated_brier_score(time_grid), ev_chi.integrated_brier_score(time_grid)
ev_cdi.brier_score(time_grid).rename('CDI').plot()
ev_chi.brier_score(time_grid).rename('CHI').plot()
plt.legend()
plt.ylabel('Brier score')
_ = plt.xlabel('Time')
```
We see from the figures that, in this case, the constant hazard interpolated estimates are not as good as the constant density interpolated estimates.
## Investigate what's going on
The instabilities at the end of the plot above is a consequence for our discretization scheme.
From `labtrans` we can get the last two discretization points
```
labtrans.cuts[-2:]
```
Now, because the censoring times in this interval are rounded down while events times are rounded up, we get an unnatural proportion of events at the final time point.
```
data = train.iloc[train[1][0] == train[1][0].max()]
data[1][1]
```
We see the almost all training individuals here have an event!
```
data[1][1].mean()
```
While the true event proportion of individuals that survival past 1500 is almost zero
```
df_train.loc[lambda x: x['duration'] > 1500]['event'].mean()
```
This is one of the dangers with discretization, and one should take caution. The simple solution would be to ensure that there are more discretization point in this interval, or simply not evaluate past the time 1348.
If we take a look at individuals in the test set that are censored some time after 1500, we see that the survival estimates are not very appropriate.
```
test = tt.tuplefy(x_test, (durations_test, events_test))
data = test.iloc[(durations_test > 1500) & (events_test == 0)]
n = data[0][0].shape[0]
idx = np.random.choice(n, 6)
fig, axs = plt.subplots(2, 3, figsize=(12, 6), sharex=True, sharey=True)
for i, ax in zip(idx, axs.flat):
x, (t, _) = data.iloc[[i]]
surv = model.predict_surv_df(x)
surv[0].rename('Survival estimate').plot(ax=ax)
ax.vlines(t, 0, 1, colors='red', linestyles="--",
label='censoring time')
ax.grid(linestyle='--')
ax.legend()
ax.set_ylabel('S(t | x)')
_ = ax.set_xlabel('Time')
```
| github_jupyter |
## Part 1: Import, Load Data.
* ### Import
```
# import standard libraries
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import pylab as pl
from scipy import stats
# import models and metrics
from sklearn import metrics, linear_model, model_selection
from sklearn.metrics import r2_score, mean_squared_error, mean_squared_log_error, mean_absolute_error
from sklearn.model_selection import cross_val_score
from sklearn.linear_model import LinearRegression
from sklearn.ensemble import GradientBoostingRegressor
```
* ### Load Data
```
# read data from '.csv' files
train = pd.read_csv("train.csv")
test = pd.read_csv("test.csv")
# identify target
target = train['Profit']
```
## Part 2: Exploratory Data Analysis.
* ### Info
```
# print the full summary of the Train dataset
train.info()
# print the full summary of the Test dataset
test.info()
```
* ### Head
```
# preview of the first 5 lines of the loaded Train data
train.head()
# preview of the first 5 lines of the loaded Test data
test.head()
```
* ### Observation of target variable
```
# target variable
train['Profit'].describe()
# visualisation of 'Profit' distribution
sns.distplot(train['Profit'], color='g')
# set 'ID' to index
train = train.set_index('ID')
test = test.set_index('ID')
```
* ### Numerical and Categorical features
#### List of Numerical and Categorical features
```
# check for Numerical and Categorical features in Train
numerical_feats_train = train.dtypes[train.dtypes != 'object'].index
print ('Quantity of Numerical features: ', len(numerical_feats_train))
print ()
print (train[numerical_feats_train].columns)
print ()
categorical_feats_train = train.dtypes[train.dtypes == 'object'].index
print ('Quantity of Categorical features: ', len(categorical_feats_train))
print ()
print (train[categorical_feats_train].columns)
```
* ### Missing values
#### List of data features with missing values
```
# check the Train features with missing values
nan_columns = [i for i in train.columns if train[i].isnull().any()]
print(train.isnull().sum())
print()
print("There are " + str(len(nan_columns)) +" columns with NAN values for 50 rows.")
nan_columns
# check the Test features with missing values
nan_columns = [i for i in test.columns if test[i].isnull().any()]
print(test.isnull().sum())
print()
print("There are " + str(len(nan_columns)) +" columns with NAN values for 50 rows.")
nan_columns
```
#### Filling missing values
Fields where NAN values have meaning.
Explaining in further depth:
* 'R&D Spend': Numerical - replacement of NAN by 'mean';
* 'Administration': Numerical - replacement of NAN by 'mean';
* 'Marketing Spend': Numerical - replacement of NAN by 'mean';
* 'State': Categorical - replacement of NAN by 'None';
* 'Category': Categorical - replacement of NAN by 'None'.
```
# Numerical NAN columns to fill in Train and Test datasets
nan_columns_fill = [
'R&D Spend',
'Administration',
'Marketing Spend'
]
# replace 'NAN' with 'mean' in these columns
train.fillna(train.mean(), inplace = True)
test.fillna(test.mean(), inplace = True)
# Categorical NAN columns to fill in Train and Test datasets
na_columns_fill = [
'State',
'Category'
]
# replace 'NAN' with 'None' in these columns
for col in na_columns_fill:
train[col].fillna('None', inplace=True)
test[col].fillna('None', inplace=True)
# check is there any mising values left in Train
train.isnull().sum().sum()
# check is there any mising values left in Test
test.isnull().sum().sum()
```
#### Visualisation of Numerical features (regplot)
```
nr_rows = 2
nr_cols = 2
fig, axs = plt.subplots(nr_rows, nr_cols, figsize=(nr_cols*3.5,nr_rows*3))
num_feats = list(numerical_feats_train)
not_plot = ['Id', 'Profit']
plot_num_feats = [c for c in list(numerical_feats_train) if c not in not_plot]
for r in range(0,nr_rows):
for c in range(0,nr_cols):
i = r*nr_cols + c
if i < len(plot_num_feats):
sns.regplot(train[plot_num_feats[i]], train['Profit'], ax = axs[r][c], color = "#5081ac" )
stp = stats.pearsonr(train[plot_num_feats[i]], train['Profit'])
str_title = "r = " + "{0:.2f}".format(stp[0]) + " " "p = " + "{0:.2f}".format(stp[1])
axs[r][c].set_title(str_title, fontsize=11)
plt.tight_layout()
plt.show()
# profit split in State level
sns.barplot(x = 'State', y = 'Profit', data = train, palette = "Blues_d")
# profit split in State level
sns.barplot(x = 'Category', y = 'Profit', data = train, palette = "Blues_d")
plt.xticks(rotation=90)
```
| github_jupyter |
Copyright (c) Microsoft Corporation. All rights reserved.
Licensed under the MIT License.

# Register TensorFlow SavedModel and deploy as webservice
Following this notebook, you will:
- Learn how to register a TF SavedModel in your Azure Machine Learning Workspace.
- Deploy your model as a web service in an Azure Container Instance.
## Prerequisites
If you are using an Azure Machine Learning Notebook VM, you are all set. Otherwise, make sure you go through the [configuration notebook](../../../configuration.ipynb) to install the Azure Machine Learning Python SDK and create a workspace.
```
import azureml.core
# Check core SDK version number.
print('SDK version:', azureml.core.VERSION)
```
## Initialize workspace
Create a [Workspace](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.workspace%28class%29?view=azure-ml-py) object from your persisted configuration.
```
from azureml.core import Workspace
ws = Workspace.from_config()
print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep='\n')
```
### Download the Model
Download and extract the model from https://amlsamplenotebooksdata.blob.core.windows.net/data/flowers_model.tar.gz to "models" directory
```
import os
import tarfile
import urllib.request
# create directory for model
model_dir = 'models'
if not os.path.isdir(model_dir):
os.mkdir(model_dir)
url="https://amlsamplenotebooksdata.blob.core.windows.net/data/flowers_model.tar.gz"
response = urllib.request.urlretrieve(url, model_dir + "/flowers_model.tar.gz")
tar = tarfile.open(model_dir + "/flowers_model.tar.gz", "r:gz")
tar.extractall(model_dir)
```
## Register model
Register a file or folder as a model by calling [Model.register()](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.model.model?view=azure-ml-py#register-workspace--model-path--model-name--tags-none--properties-none--description-none--datasets-none--model-framework-none--model-framework-version-none--child-paths-none-). For this example, we have provided a TensorFlow SavedModel (`flowers_model` in the notebook's directory).
In addition to the content of the model file itself, your registered model will also store model metadata -- model description, tags, and framework information -- that will be useful when managing and deploying models in your workspace. Using tags, for instance, you can categorize your models and apply filters when listing models in your workspace. Also, marking this model with the scikit-learn framework will simplify deploying it as a web service, as we'll see later.
```
from azureml.core import Model
model = Model.register(workspace=ws,
model_name='flowers', # Name of the registered model in your workspace.
model_path= model_dir + '/flowers_model', # Local Tensorflow SavedModel folder to upload and register as a model.
model_framework=Model.Framework.TENSORFLOW, # Framework used to create the model.
model_framework_version='1.14.0', # Version of Tensorflow used to create the model.
description='Flowers model')
print('Name:', model.name)
```
## Deploy model
Deploy your model as a web service using [Model.deploy()](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.model.model?view=azure-ml-py#deploy-workspace--name--models--inference-config--deployment-config-none--deployment-target-none-). Web services take one or more models, load them in an environment, and run them on one of several supported deployment targets.
For this example, we will deploy your TensorFlow SavedModel to an Azure Container Instance (ACI).
### Use a default environment (for supported models)
The Azure Machine Learning service provides a default environment for supported model frameworks, including TensorFlow, based on the metadata you provided when registering your model. This is the easiest way to deploy your model.
**Note**: This step can take several minutes.
```
from azureml.core import Webservice
from azureml.exceptions import WebserviceException
service_name = 'tensorflow-flower-service'
# Remove any existing service under the same name.
try:
Webservice(ws, service_name).delete()
except WebserviceException:
pass
service = Model.deploy(ws, service_name, [model])
service.wait_for_deployment(show_output=True)
```
After your model is deployed, perform a call to the web service.
```
import requests
headers = {'Content-Type': 'application/json'}
if service.auth_enabled:
headers['Authorization'] = 'Bearer '+ service.get_keys()[0]
elif service.token_auth_enabled:
headers['Authorization'] = 'Bearer '+ service.get_token()[0]
scoring_uri = service.scoring_uri # If you have a SavedModel with classify and regress,
# you can change the scoring_uri from 'uri:predict' to 'uri:classify' or 'uri:regress'.
print(scoring_uri)
with open('tensorflow-flower-predict-input.json', 'rb') as data_file:
response = requests.post(
scoring_uri, data=data_file, headers=headers)
print(response.status_code)
print(response.elapsed)
print(response.json())
```
When you are finished testing your service, clean up the deployment.
```
service.delete()
```
| github_jupyter |

# Fundamentos de Ciência de Dados
---
[](https://zenodo.org/badge/latestdoi/335308405)
---
# PPGI/UFRJ 2020.3
## Prof Sergio Serra e Jorge Zavaleta
---
# Módulo 3 - Gráficos Simples
## Matplolib
> **Matplotlib** é uma biblioteca Python especializada em de gráficos bidimensionais (incluindo gráficos 3D), muito conhecida nos círculos científicos e de engenharia (http://matplolib.org).
> ### Recursos de Matplotlib:
> * Extrema simplicidade em seu uso
> * Desenvolvimento gradual e visualização interativa de dados
> * Expressões e texto em LaTeX
> * Maior controle sobre os elementos gráficos
> * Exporte para vários formatos, como PNG, PDF, SVG e EPS
### Importando a biblioteca
```
# importando as bibliotecas
import matplotlib as mpl # usada ocasionalmente 3D
import matplotlib.pyplot as plt # importar a biblioteca matplotlib
import numpy as np # importar numpy
import pandas as pd # importat pandas
plt.style.use('classic') # estilo da biblioteca
%matplotlib inline
#%config InlineBackend.figure_format='svg'
# exemplo - grafico
x = np.linspace(-6, 3, 100)
y1 = x**4 + 5*x**3 + 5*x
y2 = 4*x**3 + 15*x**2 + 5
y3 = 12*x**2 + 30*x
#
fig, ax = plt.subplots()
ax.plot(x, y1, color="blue", label="y(x)")
ax.plot(x, y2, color="red", label="y'(x)")
ax.plot(x, y3, color="green", label="y”(x)")
ax.set_xlabel("x")
ax.set_ylabel("y")
ax.set_title('Polinomios')
ax.legend();
# ipython notebook
x1 = np.linspace(-1,4,50)
y1 = 3*(np.power(x1,4)) - 16*(np.power(x1,3))+18*(np.power(x1,2))
# para gravar a figura
fig = plt.figure(figsize=(7,4))
plt.plot(x1,y1,'-')
plt.plot(x1,y1+6,'r--'); # ;->show()
fig.savefig('imagens/figura1.png')
# verificando se a imagem foi gerada
from IPython.display import Image
Image('imagens/figura1.png')
#fig = plt.figure(figsize=(8, 2.5), facecolor="#f1f1f1")
fig = plt.figure(figsize=(8, 2.5))
# axes coordinates as fractions of the canvas width and height
left, bottom, width, height = 0.1, 0.1, 0.8, 0.8
# valores de x, y
x = np.linspace(-2, 2, 1000)
y1 = np.cos(40 * x)
y2 = np.exp(-x**2)
# figure
ax = fig.add_axes((left, bottom, width, height),facecolor="#fff0ed")
ax.plot(x, y1 * y2)
ax.plot(x, y2, 'g')
ax.plot(x, -y2, 'g')
ax.set_xlabel("x")
ax.set_ylabel("y")
# save figure
#fig.savefig("imagens/figura2.png", dpi=100, facecolor="#e1e1e1")
# verificando se a imagem foi gerada
from IPython.display import Image
Image('imagens/figura2.png')
```
### Tipos de gráficos suportados por Matplotlib
```
# tipos de formatos de graficos suportados
fig.canvas.get_supported_filetypes()
```
### Usando Varias Interfaces - Subplots
```
# exemplo de subplots
plt.figure() # criar a figura
# criando a painel 1
plt.subplot(3, 1, 1) # (linha, coluna, numero do panel)
plt.plot(x, y1,'r')
# criando o painel 2
plt.subplot(3, 1, 2)
plt.plot(x, y1+6,'b');
# criando o painel 3
plt.subplot(3, 1, 3)
plt.plot(x, y1-6,'g');
```
> **Interfaces Orientadas a Objetos**
```
# intercaes orientadas como objetos
# grid de subplots
fig, ax = plt.subplots(2)
ax[0].plot(x, y1,'r')
ax[1].plot(x, y1-6);
# subplot vertical
x1 = np.linspace(-2, 2, 100)
y1 = x1**3+ x1*2 + 4
fig, (ax1, ax2) = plt.subplots(1, 2)
fig.suptitle('Subplot Vertical')
ax1.plot(x1, y1,'.r')
ax2.plot(x1, y1-6);
#fig.savefig('imagens/fx.png')
# grid de subplots
#fig,ax = plt.subplots(2,2,figsize=(8,8))
fig,ax = plt.subplots(2,2,figsize=(8,8),constrained_layout=True)
ax[0][0].plot(x1,y1,'r')
ax[0][0].set_title('$f(x)=3x^4-16x^3+18x^2$')
ax[0][0].set_ylabel('y')
ax[0][0].set_xlabel('x')
#
ax[0][1].plot(x1,y1-6,'g')
ax[0][1].set_title('$f(x)=3x^4-16x^3+18x^2-6$')
#
ax[1][0].plot(x1,y1+6,'orange')
ax[1][0].set_title('$f(x)=3x^4-16x^3+18x^2+6$')
#
ax[1][1].plot(x1,y1+10,'blue')
ax[1][1].set_title('$f(x)=3x^4-16x^3+18x^2+10$')
#
#plt.suptitle('$f(x)=3x^4-16x^3+18x^2$', verticalalignment='bottom', fontsize=14)
plt.show()
#fig.savefig('imagens/subpots.png')
```
> **Graficar linhas simples**
```
# grid de linhas
plt.style.use('seaborn-whitegrid')
fig = plt.figure()
ax = plt.axes()
plt.plot(x1,y1,'-');
```
> ### Cores e Estilos de Linhas
```
# graficos de diversas cores
plt.plot(x1, y1, color='blue') # specify color by name
plt.plot(x1, y1-5, color='g') # short color code (rgbcmyk)
plt.plot(x1, y1-10, color='0.75') # Grayscale between 0 and 1
plt.plot(x1, y1-15, color='#FFDD44') # Hex code (RRGGBB from 00 to FF)
plt.plot(x1, y1+5, color=(1.0,0.2,0.3)) # RGB tuple, values 0 and 1
plt.plot(x1, y1+10, color='chartreuse');
# Graficos e estilos de linhas
plt.plot(x1, x1 + 0, linestyle='solid')
plt.plot(x1, x1 + 2, linestyle='dashed')
plt.plot(x1, x1 + 4, linestyle='dashdot')
plt.plot(x1, x1 + 6, linestyle='dotted');
# usando codigos
plt.plot(x1, x1 + 8, linestyle='-') # solid
plt.plot(x1, x1 + 10, linestyle='--') # dashed
plt.plot(x1, x1 + 12, linestyle='-.') # dashdot
plt.plot(x1, x1 + 14, linestyle=':'); # dotted
```
> ### Ajustando os limites do eixos
```
# Ajustando limites de eixos
plt.plot(x1, y1)
plt.xlim(-1.5, 4.5)
plt.ylim(-35, 45);
# Ajustando eixos
plt.plot(x1, y1,'r')
# [xmin,xmax, ymin,ymax]
plt.axis([-1, 5, -30, 40]);
```
>### Melhorando os gráficos - subplot
```
# exemplo de subplots - problema
# gerando os valores
x = np.linspace(-6, 3, 100)
y1 = x**4 + 5*x**3 + 5*x
# plot
fig = plt.figure() # criar a figura
# criando a painel 1
plt.subplot(3, 1, 1) # (linha, coluna, numero do panel)
plt.plot(x, y1,'r')
plt.xlabel('eixo x')
plt.ylabel('y')
plt.title('Três subplots')
# criando o painel 2
plt.subplot(3, 1, 2)
plt.plot(x, y1+6,'b')
plt.xlabel('x2')
# criando o painel 3
plt.subplot(3, 1, 3)
plt.plot(x, y1-6,'g')
plt.xlabel('x3')
#
# melhorar o plot
#fig.tight_layout() # melhorar
plt.show()
# grid de subplots
# valores
x1 = np.linspace(-1,4,50)
y1 = 3*(np.power(x1,4)) - 16*(np.power(x1,3))+18*(np.power(x1,2))
#
fig,ax = plt.subplots(2,2,figsize=(8,8),constrained_layout=False) # True
ax[0][0].plot(x1,y1,'r')
ax[0][0].set_title('$f(x)=3x^4-16x^3+18x^2$')
ax[0][0].set_ylabel('y')
ax[0][0].set_xlabel('Polinomio em x')
#
ax[0][1].plot(x1,y1-6,'g')
ax[0][1].set_title('$f(x)=3x^4-16x^3+18x^2-6$')
#
ax[1][0].plot(x1,y1+6,'orange')
ax[1][0].set_title('$f(x)=3x^4-16x^3+18x^2+6$')
#
ax[1][1].plot(x1,y1+10,'blue')
ax[1][1].set_title('$f(x)=3x^4-16x^3+18x^2+10$')
#
plt.suptitle('$f(x)=3x^4-16x^3+18x^2$', verticalalignment='bottom', fontsize=14)
# melhorar
#plt.subplots_adjust(left=0.125,bottom=0.1, right=0.9, top=0.9, wspace=0.2, hspace=0.35) #
#plt.show()
#fig.savefig('imagens/subpots.png')
```
> ### Gráficos 3D
```
# matplotlib-exemplo-30
# define uma função
def f(x, y):
return np.sin(x) ** 10 + np.cos(10 + y * x) * np.cos(x)
# gerando os eixos 3D
#from mpl_toolkits import mplot3d
#
fig = plt.figure(figsize=(10,7))
ax = plt.axes(projection="3d")
plt.show()
```
>### Curva 3D
```
# curva 3D
x = np.linspace(0, 5, 50)
y = np.linspace(0, 5, 50)
#X, Y = np.meshgrid(x, y)
Z = f(x, y)
#
fig = plt.figure(figsize=(10,7))
ax = plt.axes(projection='3d')
ax.set_xlabel('seno-coseno')
#
ax.plot3D(x, y, Z, 'red')
plt.show()
```
>### Superficie 3D
```
# superficie 3D
x = np.linspace(0, 5, 50)
y = np.linspace(0, 5, 50)
X, Y = np.meshgrid(x, y)
Z = f(X, Y)
#
fig = plt.figure(figsize=(10,7))
ax = plt.axes(projection='3d')
ax.set_xlabel('Imagem 3D')
#
ax.plot_surface(X, Y, Z, cmap=plt.cm.jet, rstride=1, cstride=1, linewidth=0)
#ax.plot_surface(X, Y, Z, cmap='RdGy', rstride=1, cstride=1, linewidth=0)
plt.show()
```
>### Contorno
```
# matplotlib-exemplo-31
# gerando os valores usando meshgrid
x = np.linspace(0, 5, 50)
y = np.linspace(0, 5, 50)
X, Y = np.meshgrid(x, y)
Z = f(X, Y)
plt.contour(X, Y, Z, colors='blue'); #grafico 3-D
# matplotlib-exemplo-32
# usando contornos
plt.contour(X, Y, Z, 20, cmap='RdGy');
# matplotlib-exemplo-32
# Graficando barra de cores
plt.contourf(X, Y, Z, 20, cmap='RdGy')
plt.colorbar();
```
> ### Curva 3D e Pontos discretos
```
# linha 3D + scatter
x = np.linspace(0, 5, 50)
y = np.linspace(0, 5, 50)
#X, Y = np.meshgrid(x, y)
Z = f(x, y)
#
fig = plt.figure(figsize=(10,7))
ax = plt.axes(projection='3d')
ax.set_xlabel('seno-coseno')
#
ax.plot3D(x, y, Z, 'red')
# scatter
x = x + 0.1 * np.random.randn(50)
y = y + 0.1 * np.random.randn(50)
zdata = f(x,y)
ax.scatter3D(x, y, zdata, c=zdata, cmap='Greens')
ax.set_xlabel('seno')
plt.show()
```
## Histogramas
```
# matplotlib-exemplo-33
# graficando um histograma
%matplotlib inline
plt.style.use('seaborn-white')
data = np.random.randn(900)
plt.hist(data);
# matplotlib-exemplo-34
# Histograma customizado
plt.hist(data, bins=30, density=True, alpha=0.5,histtype='stepfilled', color='steelblue',edgecolor='none');
# matplotlib-exemplo-35
# histogramas sobrepostos
x1 = np.random.normal(0, 0.8, 1000)
x2 = np.random.normal(-2, 1, 1000)
x3 = np.random.normal(3, 2, 1000)
kwargs = dict(histtype='stepfilled', alpha=0.8, density=True, bins=40)
plt.hist(x1, **kwargs)
plt.hist(x2, **kwargs)
plt.hist(x3, **kwargs);
```
## Customizando as Legendas
```
# matplotlib-exemplo-36
# Personalizando as legendas
%matplotlib inline
plt.style.use('classic')
x2 = np.linspace(0, 10, 1000)
fig, ax2 = plt.subplots()
ax2.plot(x2, np.sin(x2), '-b', label='Seno()')
ax2.plot(x2, np.cos(2*x2), '--r', label='Coseno()')
ax2.axis('equal')
leg = ax2.legend()
plt.show()
# matplotlib-exemplo-37
# mudando a legenda de posição e colunas =2
ax2.legend(loc='upper left', frameon=False, ncol=2)
fig
# matplotlib-exemplo-38
#
plt.style.use('seaborn-white')
x3 = np.arange(10)
y3 = (20, 35, 30, 35, 27, 40, 45, 41, 39, 30)
plt.bar(x3,y3)
plt.show() # Try commenting this an run
plt.scatter(x3,y3) # scatter plot
plt.show()
```
>### Exemplo IRIS
```
# imagem iris
# creditos Sebastian Raschka: https://sebastianraschka.com/Articles/2015_pca_in_3_steps.html
from IPython.display import Image
Image('imagens/iris.png')
# matplotlib-exemplo-39
# Graficando um dataset conhecido com pandas
df = pd.read_csv('data/iris.csv')
df.hist()# Histograma
print('\n')
df.plot(); # grafico linear
```
>### Gráfico de Barras
```
# matplotlib-exemplo-40
# Histogramas de cores
pre = np.array([19, 6, 11, 9])
post = np.array([15, 11, 9, 8])
labels=['Pesquisa '+x for x in list('ABCD')]
fig, ax = plt.subplots(figsize=(8, 3.5))
width = 0.4 # largura da barra
xlocs = np.arange(len(pre))
ax.bar(xlocs-width, pre, width, color='green', label='Verde')
ax.bar(xlocs, post, width, color='#1f10ed', label='Azul')
# labels, grids, titulos e grvar a figura
ax.set_xticks(ticks=range(len(pre)))
ax.set_xticklabels(labels)
ax.yaxis.grid(True)
ax.legend(loc='best')
ax.set_ylabel('Quantidade')
fig.suptitle('Barras Simples')
fig.tight_layout(pad=1)
#fig.savefig('data/barra.png', dpi=125)
```
>### Gráfico de PIE
```
# matplotlib-exemplo-41
# grafico de pizza
datax = np.array([15,8,4])
labels = ['Engenharia', 'Matemática', 'Física']
explode = (0, 0.1, 0)
colrs=['cyan', 'tan', 'wheat']
# graficos
fig, ax = plt.subplots(figsize=(8, 3.5))
ax.pie(datax, explode=explode, labels=labels, autopct='%1.1f%%',startangle=270, colors=colrs)
ax.axis('equal') # mantenha um circulo
fig.suptitle("Pizza de Cursos");
#fig.savefig('data/pizza.png', dpi=125)
# matplotlib-exemplo-42
# grafico de pizza usando opção 'shadow'
xp = np.array([1,2,3,4,5,6,7,8,9,0])
label = ['a','b','c','d','e','f','g','h','i','j']
explode = [0.2, 0.1, 0.5, 0, 0, 0.3, 0.3, 0.2, 0.1,0]
# grafico
fig, axx = plt.subplots(figsize=(6, 6))
axx.pie(xp, labels=label, explode = explode, shadow=True, autopct ='%2.2f%%')
plt.title('Gráfico de Pizza')
plt.show()
#fig.savefig('data/pizzashadow.png', dpi=125)
# matplotlib-exemplo-43
# graficando coordenadas polares
r = np.arange(0, 10.0, 0.1)
theta = 2* np.pi * r
plt.polar(theta, r, color ='r')
plt.show()
```
>### Decorando os Gráficos com Textos, Setas e Anotações
```
# matplotlib-exemplo-44
# decorando: texto, setas e anotações
sx = np.arange(0, 2*np.pi, .01)
sy = np.sin(sx)
plt.plot(sx, sy, color = 'r')
plt.text(0.1, -0.04, '$sin(0) = 0$')
plt.text(1.5, 0.9, '$sin(90) = 1$')
plt.text(2.0, 0, '$sin(180) = 0$')
plt.text(4.5, -0.9, '$sin(270) = -1$')
plt.text(6.0, 0.0, '$sin(360) = 1$')
#plt.annotate('$sin(theta)=0$', xy=(3, 0.1), xytext=(5, 0.7),arrowprops=dict(facecolor='blue', shrink=0.05))
plt.title('Insertando texto e anotações no Gráfico')
plt.xlabel('$theta$')
plt.ylabel('$sy = sin( theta)$')
plt.show()
```
> ### Gráficos Usando os datasets do PROUNI
```
# Dataset prouni 2020
prouni = pd.read_csv('data/cursos-prouni2020.csv',delimiter=',')
prouni.head()
# valores dos eixos
print(len(prouni))
x = range(0,len(prouni),1)
print(len(x))
y = prouni['bolsa_integral_cotas']
y1 = prouni['nota_integral_ampla']
len(y1)
plt.plot(x,y,'r')
#plt.plot(x,y1,'+b')
plt.ylabel('Bolsa Integral')
plt.xlabel('Prouni 2020')
plt.title('Exame PROUNI 20/02/2021')
plt.show()
```
## Seaborn
> **Seaborn** é uma biblioteca de visualização de dados Python baseada em matplotlib e fornece uma interface de alto nível para desenhar gráficos estatísticos atraentes e informativos. Maiores informações em --> [Seaborn]('https://seaborn.pydata.org/index.html')
>> Importar a biblioteca:```import seaborn as sns```
```
# importar a bibliteca
import seaborn as sns
#plt.style.use('seaborn-whitegrid')
sns.set_theme() # tema
# dados
tips = sns.load_dataset("tips")
tips
# Create a visualization
sns.relplot(data=tips, x="total_bill", y="tip", col="time",
hue="smoker", style="smoker", size="size");
# dados prouni
prouni.head()
dados_rj = prouni[prouni['uf_busca']=='RJ']
dados_rj.head()
#print(len(dados_rj))
# visualização do prouni
sns.relplot(data=dados_rj, x="bolsa_integral_cotas", y="nota_integral_ampla", col="turno",
hue="nome", style="nome", size="turno");
```
### API em visualizações
```
# exmplo
dots = sns.load_dataset("dots") # dados embutidos no seaborn
# cria o gráfico
sns.relplot(data=dots, kind="line", x="time", y="firing_rate", col="align", hue="choice", size="coherence", style="choice",
facet_kws=dict(sharex=False),);
```
### Estimativa estatística e barras de erro
```
# Exemplo de estimativa
fmri = sns.load_dataset("fmri") # dados embutidos no seaborn
# cria o gráfico
sns.relplot(data=fmri, kind="line", x="timepoint", y="signal", col="region",
hue="event", style="event",);
# dados prouni = RJ
sns.relplot(data=dados_rj, kind="line", x="nota_integral_ampla", y="bolsa_integral_cotas",
col="turno", hue='cidade_busca',style ="cidade_busca",);
```
#### Usando o modelo de regressão linear - lmplot()
```
# exemplo: modelo de regressão linear
sns.lmplot(data=tips, x="total_bill", y="tip", col="time", hue="smoker");
# dados prouni = RJ
sns.lmplot(data=dados_rj, x="nota_integral_ampla", y="bolsa_integral_cotas", col="turno", hue='cidade_busca');
```
### Resumos informativos de distribuição
> As análises estatísticas requerem conhecimento sobre a distribuição de variáveis no conjunto de dados.
```
sns.displot(data=tips, x="total_bill", col="time", kde=True);
sns.displot(data=dados_rj, x="nota_integral_cotas", col="turno", kde=True);
```
### Gráficos especializados para dados categóricos
```
# exemplo seaborn
sns.catplot(data=tips, kind="swarm", x="day", y="total_bill", hue="smoker");
# exemplo dados prouni
sns.catplot(data=dados_rj, kind="swarm", x="grau", y="nota_integral_cotas", hue="turno",);
# exemplo dados prouni
sns.stripplot(data=dados_rj, x="grau", y="nota_integral_cotas", hue="turno",);
# Exemplo dados prouni
sns.stripplot(data=dados_rj, x="grau", y="nota_integral_cotas", hue="turno",jitter=0.5);
# exemplo embutido no seaborn
sns.catplot(data=tips, kind="bar", x="day", y="total_bill", hue="smoker");
# Exemplo dados prouni
sns.catplot(data=dados_rj, kind='bar', x="grau", y="nota_integral_cotas", hue="turno");
```
### Visualizações compostas de conjuntos de dados multivariados
```
# exemplo do seaborn
penguins = sns.load_dataset("penguins")
sns.jointplot(data=penguins, x="flipper_length_mm", y="bill_length_mm", hue="species")
# Exemplo dados prouni
sns.jointplot(data=dados_rj, x="bolsa_integral_cotas", y="nota_integral_cotas", hue="turno");
# dados prouni
sns.pairplot(data=dados_rj, hue="grau");
```
### Classes e funções para fazer gráficos complexos
> Estas ferramentas funcionam combinando funções de plotagem no nível dos eixos com objetos que gerenciam o layout da figura, ligando a estrutura de um conjunto de dados a uma grade de eixos.
```
# exemplo
g = sns.PairGrid(dados_rj, hue="grau", corner=True)
g.map_lower(sns.kdeplot, hue=None, levels=5, color=".2")
g.map_lower(sns.scatterplot, marker="+")
g.map_diag(sns.histplot, element="step", linewidth=0, kde=True)
g.add_legend(frameon=True)
g.legend.set_bbox_to_anchor((.61, .6))
```
---
#### Fudamentos para Ciência Dados © Copyright 2021, Sergio Serra & Jorge Zavaleta
| github_jupyter |
```
#load watermark
%load_ext watermark
%watermark -a 'Gopala KR' -u -d -v -p watermark,numpy,pandas,matplotlib,nltk,sklearn,tensorflow,theano,mxnet,chainer,seaborn,keras,tflearn,bokeh,gensim
import keras
keras.__version__
from keras import backend as K
K.clear_session()
```
# Generating images
This notebook contains the second code sample found in Chapter 8, Section 4 of [Deep Learning with Python](https://www.manning.com/books/deep-learning-with-python?a_aid=keras&a_bid=76564dff). Note that the original text features far more content, in particular further explanations and figures: in this notebook, you will only find source code and related comments.
---
## Variational autoencoders
Variational autoencoders, simultaneously discovered by Kingma & Welling in December 2013, and Rezende, Mohamed & Wierstra in January 2014,
are a kind of generative model that is especially appropriate for the task of image editing via concept vectors. They are a modern take on
autoencoders -- a type of network that aims to "encode" an input to a low-dimensional latent space then "decode" it back -- that mixes ideas
from deep learning with Bayesian inference.
A classical image autoencoder takes an image, maps it to a latent vector space via an "encoder" module, then decode it back to an output
with the same dimensions as the original image, via a "decoder" module. It is then trained by using as target data the _same images_ as the
input images, meaning that the autoencoder learns to reconstruct the original inputs. By imposing various constraints on the "code", i.e.
the output of the encoder, one can get the autoencoder to learn more or less interesting latent representations of the data. Most
commonly, one would constraint the code to be very low-dimensional and sparse (i.e. mostly zeros), in which case the encoder acts as a way
to compress the input data into fewer bits of information.
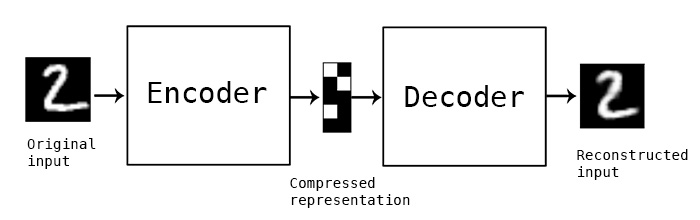
In practice, such classical autoencoders don't lead to particularly useful or well-structured latent spaces. They're not particularly good
at compression, either. For these reasons, they have largely fallen out of fashion over the past years. Variational autoencoders, however,
augment autoencoders with a little bit of statistical magic that forces them to learn continuous, highly structured latent spaces. They
have turned out to be a very powerful tool for image generation.
A VAE, instead of compressing its input image into a fixed "code" in the latent space, turns the image into the parameters of a statistical
distribution: a mean and a variance. Essentially, this means that we are assuming that the input image has been generated by a statistical
process, and that the randomness of this process should be taken into accounting during encoding and decoding. The VAE then uses the mean
and variance parameters to randomly sample one element of the distribution, and decodes that element back to the original input. The
stochasticity of this process improves robustness and forces the latent space to encode meaningful representations everywhere, i.e. every
point sampled in the latent will be decoded to a valid output.
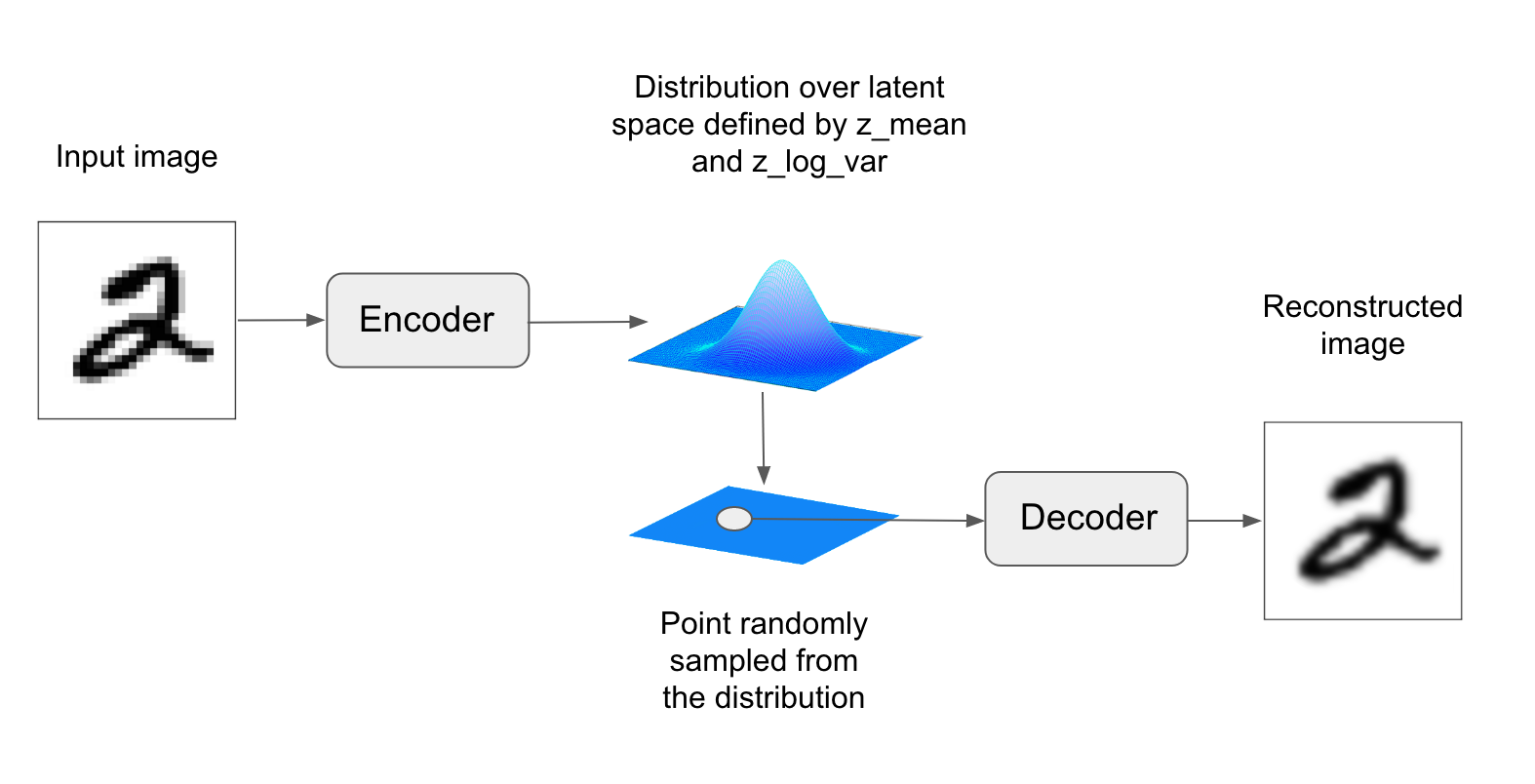
In technical terms, here is how a variational autoencoder works. First, an encoder module turns the input samples `input_img` into two
parameters in a latent space of representations, which we will note `z_mean` and `z_log_variance`. Then, we randomly sample a point `z`
from the latent normal distribution that is assumed to generate the input image, via `z = z_mean + exp(z_log_variance) * epsilon`, where
epsilon is a random tensor of small values. Finally, a decoder module will map this point in the latent space back to the original input
image. Because `epsilon` is random, the process ensures that every point that is close to the latent location where we encoded `input_img`
(`z-mean`) can be decoded to something similar to `input_img`, thus forcing the latent space to be continuously meaningful. Any two close
points in the latent space will decode to highly similar images. Continuity, combined with the low dimensionality of the latent space,
forces every direction in the latent space to encode a meaningful axis of variation of the data, making the latent space very structured
and thus highly suitable to manipulation via concept vectors.
The parameters of a VAE are trained via two loss functions: first, a reconstruction loss that forces the decoded samples to match the
initial inputs, and a regularization loss, which helps in learning well-formed latent spaces and reducing overfitting to the training data.
Let's quickly go over a Keras implementation of a VAE. Schematically, it looks like this:
```
# Encode the input into a mean and variance parameter
#z_mean, z_log_variance = encoder(input_img)
# Draw a latent point using a small random epsilon
#z = z_mean + exp(z_log_variance) * epsilon
# Then decode z back to an image
#reconstructed_img = decoder(z)
# Instantiate a model
#model = Model(input_img, reconstructed_img)
# Then train the model using 2 losses:
# a reconstruction loss and a regularization loss
```
Here is the encoder network we will use: a very simple convnet which maps the input image `x` to two vectors, `z_mean` and `z_log_variance`.
```
import keras
from keras import layers
from keras import backend as K
from keras.models import Model
import numpy as np
img_shape = (28, 28, 1)
batch_size = 16
latent_dim = 2 # Dimensionality of the latent space: a plane
input_img = keras.Input(shape=img_shape)
x = layers.Conv2D(32, 3,
padding='same', activation='relu')(input_img)
x = layers.Conv2D(64, 3,
padding='same', activation='relu',
strides=(2, 2))(x)
x = layers.Conv2D(64, 3,
padding='same', activation='relu')(x)
x = layers.Conv2D(64, 3,
padding='same', activation='relu')(x)
shape_before_flattening = K.int_shape(x)
x = layers.Flatten()(x)
x = layers.Dense(32, activation='relu')(x)
z_mean = layers.Dense(latent_dim)(x)
z_log_var = layers.Dense(latent_dim)(x)
```
Here is the code for using `z_mean` and `z_log_var`, the parameters of the statistical distribution assumed to have produced `input_img`, to
generate a latent space point `z`. Here, we wrap some arbitrary code (built on top of Keras backend primitives) into a `Lambda` layer. In
Keras, everything needs to be a layer, so code that isn't part of a built-in layer should be wrapped in a `Lambda` (or else, in a custom
layer).
```
def sampling(args):
z_mean, z_log_var = args
epsilon = K.random_normal(shape=(K.shape(z_mean)[0], latent_dim),
mean=0., stddev=1.)
return z_mean + K.exp(z_log_var) * epsilon
z = layers.Lambda(sampling)([z_mean, z_log_var])
```
This is the decoder implementation: we reshape the vector `z` to the dimensions of an image, then we use a few convolution layers to obtain a final
image output that has the same dimensions as the original `input_img`.
```
# This is the input where we will feed `z`.
decoder_input = layers.Input(K.int_shape(z)[1:])
# Upsample to the correct number of units
x = layers.Dense(np.prod(shape_before_flattening[1:]),
activation='relu')(decoder_input)
# Reshape into an image of the same shape as before our last `Flatten` layer
x = layers.Reshape(shape_before_flattening[1:])(x)
# We then apply then reverse operation to the initial
# stack of convolution layers: a `Conv2DTranspose` layers
# with corresponding parameters.
x = layers.Conv2DTranspose(32, 3,
padding='same', activation='relu',
strides=(2, 2))(x)
x = layers.Conv2D(1, 3,
padding='same', activation='sigmoid')(x)
# We end up with a feature map of the same size as the original input.
# This is our decoder model.
decoder = Model(decoder_input, x)
# We then apply it to `z` to recover the decoded `z`.
z_decoded = decoder(z)
```
The dual loss of a VAE doesn't fit the traditional expectation of a sample-wise function of the form `loss(input, target)`. Thus, we set up
the loss by writing a custom layer with internally leverages the built-in `add_loss` layer method to create an arbitrary loss.
```
class CustomVariationalLayer(keras.layers.Layer):
def vae_loss(self, x, z_decoded):
x = K.flatten(x)
z_decoded = K.flatten(z_decoded)
xent_loss = keras.metrics.binary_crossentropy(x, z_decoded)
kl_loss = -5e-4 * K.mean(
1 + z_log_var - K.square(z_mean) - K.exp(z_log_var), axis=-1)
return K.mean(xent_loss + kl_loss)
def call(self, inputs):
x = inputs[0]
z_decoded = inputs[1]
loss = self.vae_loss(x, z_decoded)
self.add_loss(loss, inputs=inputs)
# We don't use this output.
return x
# We call our custom layer on the input and the decoded output,
# to obtain the final model output.
y = CustomVariationalLayer()([input_img, z_decoded])
```
Finally, we instantiate and train the model. Since the loss has been taken care of in our custom layer, we don't specify an external loss
at compile time (`loss=None`), which in turns means that we won't pass target data during training (as you can see we only pass `x_train`
to the model in `fit`).
```
from keras.datasets import mnist
vae = Model(input_img, y)
vae.compile(optimizer='rmsprop', loss=None)
vae.summary()
# Train the VAE on MNIST digits
(x_train, _), (x_test, y_test) = mnist.load_data()
x_train = x_train.astype('float32') / 255.
x_train = x_train.reshape(x_train.shape + (1,))
x_test = x_test.astype('float32') / 255.
x_test = x_test.reshape(x_test.shape + (1,))
vae.fit(x=x_train, y=None,
shuffle=True,
epochs=2,
batch_size=batch_size,
validation_data=(x_test, None))
```
Once such a model is trained -- e.g. on MNIST, in our case -- we can use the `decoder` network to turn arbitrary latent space vectors into
images:
```
import matplotlib.pyplot as plt
from scipy.stats import norm
# Display a 2D manifold of the digits
n = 15 # figure with 15x15 digits
digit_size = 28
figure = np.zeros((digit_size * n, digit_size * n))
# Linearly spaced coordinates on the unit square were transformed
# through the inverse CDF (ppf) of the Gaussian
# to produce values of the latent variables z,
# since the prior of the latent space is Gaussian
grid_x = norm.ppf(np.linspace(0.05, 0.95, n))
grid_y = norm.ppf(np.linspace(0.05, 0.95, n))
for i, yi in enumerate(grid_x):
for j, xi in enumerate(grid_y):
z_sample = np.array([[xi, yi]])
z_sample = np.tile(z_sample, batch_size).reshape(batch_size, 2)
x_decoded = decoder.predict(z_sample, batch_size=batch_size)
digit = x_decoded[0].reshape(digit_size, digit_size)
figure[i * digit_size: (i + 1) * digit_size,
j * digit_size: (j + 1) * digit_size] = digit
plt.figure(figsize=(10, 10))
plt.imshow(figure, cmap='Greys_r')
plt.show()
```
The grid of sampled digits shows a completely continuous distribution of the different digit classes, with one digit morphing into another
as you follow a path through latent space. Specific directions in this space have a meaning, e.g. there is a direction for "four-ness",
"one-ness", etc.
| github_jupyter |
# Training Neural Networks
The network we built in the previous part isn't so smart, it doesn't know anything about our handwritten digits. Neural networks with non-linear activations work like universal function approximators. There is some function that maps your input to the output. For example, images of handwritten digits to class probabilities. The power of neural networks is that we can train them to approximate this function, and basically any function given enough data and compute time.
<img src="assets/function_approx.png" width=500px>
At first the network is naive, it doesn't know the function mapping the inputs to the outputs. We train the network by showing it examples of real data, then adjusting the network parameters such that it approximates this function.
To find these parameters, we need to know how poorly the network is predicting the real outputs. For this we calculate a **loss function** (also called the cost), a measure of our prediction error. For example, the mean squared loss is often used in regression and binary classification problems
$$
\large \ell = \frac{1}{2n}\sum_i^n{\left(y_i - \hat{y}_i\right)^2}
$$
where $n$ is the number of training examples, $y_i$ are the true labels, and $\hat{y}_i$ are the predicted labels.
By minimizing this loss with respect to the network parameters, we can find configurations where the loss is at a minimum and the network is able to predict the correct labels with high accuracy. We find this minimum using a process called **gradient descent**. The gradient is the slope of the loss function and points in the direction of fastest change. To get to the minimum in the least amount of time, we then want to follow the gradient (downwards). You can think of this like descending a mountain by following the steepest slope to the base.
<img src='assets/gradient_descent.png' width=350px>
## Backpropagation
For single layer networks, gradient descent is straightforward to implement. However, it's more complicated for deeper, multilayer neural networks like the one we've built. Complicated enough that it took about 30 years before researchers figured out how to train multilayer networks.
Training multilayer networks is done through **backpropagation** which is really just an application of the chain rule from calculus. It's easiest to understand if we convert a two layer network into a graph representation.
<img src='assets/backprop_diagram.png' width=550px>
In the forward pass through the network, our data and operations go from bottom to top here. We pass the input $x$ through a linear transformation $L_1$ with weights $W_1$ and biases $b_1$. The output then goes through the sigmoid operation $S$ and another linear transformation $L_2$. Finally we calculate the loss $\ell$. We use the loss as a measure of how bad the network's predictions are. The goal then is to adjust the weights and biases to minimize the loss.
To train the weights with gradient descent, we propagate the gradient of the loss backwards through the network. Each operation has some gradient between the inputs and outputs. As we send the gradients backwards, we multiply the incoming gradient with the gradient for the operation. Mathematically, this is really just calculating the gradient of the loss with respect to the weights using the chain rule.
$$
\large \frac{\partial \ell}{\partial W_1} = \frac{\partial L_1}{\partial W_1} \frac{\partial S}{\partial L_1} \frac{\partial L_2}{\partial S} \frac{\partial \ell}{\partial L_2}
$$
**Note:** I'm glossing over a few details here that require some knowledge of vector calculus, but they aren't necessary to understand what's going on.
We update our weights using this gradient with some learning rate $\alpha$.
$$
\large W^\prime_1 = W_1 - \alpha \frac{\partial \ell}{\partial W_1}
$$
The learning rate $\alpha$ is set such that the weight update steps are small enough that the iterative method settles in a minimum.
## Losses in PyTorch
Let's start by seeing how we calculate the loss with PyTorch. Through the `nn` module, PyTorch provides losses such as the cross-entropy loss (`nn.CrossEntropyLoss`). You'll usually see the loss assigned to `criterion`. As noted in the last part, with a classification problem such as MNIST, we're using the softmax function to predict class probabilities. With a softmax output, you want to use cross-entropy as the loss. To actually calculate the loss, you first define the criterion then pass in the output of your network and the correct labels.
Something really important to note here. Looking at [the documentation for `nn.CrossEntropyLoss`](https://pytorch.org/docs/stable/nn.html#torch.nn.CrossEntropyLoss),
> This criterion combines `nn.LogSoftmax()` and `nn.NLLLoss()` in one single class.
>
> The input is expected to contain scores for each class.
This means we need to pass in the raw output of our network into the loss, not the output of the softmax function. This raw output is usually called the *logits* or *scores*. We use the logits because softmax gives you probabilities which will often be very close to zero or one but floating-point numbers can't accurately represent values near zero or one ([read more here](https://docs.python.org/3/tutorial/floatingpoint.html)). It's usually best to avoid doing calculations with probabilities, typically we use log-probabilities.
```
import torch
from torch import nn
import torch.nn.functional as F
from torchvision import datasets, transforms
# Define a transform to normalize the data
transform = transforms.Compose([transforms.ToTensor(),
transforms.Normalize((0.5,), (0.5,)),
])
# Download and load the training data
trainset = datasets.MNIST('~/.pytorch/MNIST_data/', download=True, train=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=64, shuffle=True)
```
### Note
If you haven't seen `nn.Sequential` yet, please finish the end of the Part 2 notebook.
```
# Build a feed-forward network
model = nn.Sequential(nn.Linear(784, 128),
nn.ReLU(),
nn.Linear(128, 64),
nn.ReLU(),
nn.Linear(64, 10))
# Define the loss
criterion = nn.CrossEntropyLoss()
# Get our data
images, labels = next(iter(trainloader))
# Flatten images
images = images.view(images.shape[0], -1)
# Forward pass, get our logits
logits = model(images)
# Calculate the loss with the logits and the labels
loss = criterion(logits, labels)
print(loss)
```
In my experience it's more convenient to build the model with a log-softmax output using `nn.LogSoftmax` or `F.log_softmax` ([documentation](https://pytorch.org/docs/stable/nn.html#torch.nn.LogSoftmax)). Then you can get the actual probabilities by taking the exponential `torch.exp(output)`. With a log-softmax output, you want to use the negative log likelihood loss, `nn.NLLLoss` ([documentation](https://pytorch.org/docs/stable/nn.html#torch.nn.NLLLoss)).
>**Exercise:** Build a model that returns the log-softmax as the output and calculate the loss using the negative log likelihood loss. Note that for `nn.LogSoftmax` and `F.log_softmax` you'll need to set the `dim` keyword argument appropriately. `dim=0` calculates softmax across the rows, so each column sums to 1, while `dim=1` calculates across the columns so each row sums to 1. Think about what you want the output to be and choose `dim` appropriately.
```
# TODO: Build a feed-forward network
model = nn.Sequential(nn.Linear(784, 128),
nn.ReLU(),
nn.Linear(128, 64),
nn.ReLU(),
nn.Linear(64, 10),
nn.LogSoftmax(dim=1))
# TODO: Define the loss
criterion = nn.NLLLoss()
### Run this to check your work
# Get our data
images, labels = next(iter(trainloader))
# Flatten images
images = images.view(images.shape[0], -1)
# Forward pass, get our logits
logits = model(images)
# Calculate the loss with the logits and the labels
loss = criterion(logits, labels)
print(loss)
```
## Autograd
Now that we know how to calculate a loss, how do we use it to perform backpropagation? Torch provides a module, `autograd`, for automatically calculating the gradients of tensors. We can use it to calculate the gradients of all our parameters with respect to the loss. Autograd works by keeping track of operations performed on tensors, then going backwards through those operations, calculating gradients along the way. To make sure PyTorch keeps track of operations on a tensor and calculates the gradients, you need to set `requires_grad = True` on a tensor. You can do this at creation with the `requires_grad` keyword, or at any time with `x.requires_grad_(True)`.
You can turn off gradients for a block of code with the `torch.no_grad()` content:
```python
x = torch.zeros(1, requires_grad=True)
>>> with torch.no_grad():
... y = x * 2
>>> y.requires_grad
False
```
Also, you can turn on or off gradients altogether with `torch.set_grad_enabled(True|False)`.
The gradients are computed with respect to some variable `z` with `z.backward()`. This does a backward pass through the operations that created `z`.
```
x = torch.randn(2,2, requires_grad=True)
print(x)
y = x**2
print(y)
```
Below we can see the operation that created `y`, a power operation `PowBackward0`.
```
## grad_fn shows the function that generated this variable
print(y.grad_fn)
```
The autograd module keeps track of these operations and knows how to calculate the gradient for each one. In this way, it's able to calculate the gradients for a chain of operations, with respect to any one tensor. Let's reduce the tensor `y` to a scalar value, the mean.
```
z = y.mean()
print(z)
```
You can check the gradients for `x` and `y` but they are empty currently.
```
print(x.grad)
```
To calculate the gradients, you need to run the `.backward` method on a Variable, `z` for example. This will calculate the gradient for `z` with respect to `x`
$$
\frac{\partial z}{\partial x} = \frac{\partial}{\partial x}\left[\frac{1}{n}\sum_i^n x_i^2\right] = \frac{x}{2}
$$
```
z.backward()
print(x.grad)
print(x/2)
```
These gradients calculations are particularly useful for neural networks. For training we need the gradients of the cost with respect to the weights. With PyTorch, we run data forward through the network to calculate the loss, then, go backwards to calculate the gradients with respect to the loss. Once we have the gradients we can make a gradient descent step.
## Loss and Autograd together
When we create a network with PyTorch, all of the parameters are initialized with `requires_grad = True`. This means that when we calculate the loss and call `loss.backward()`, the gradients for the parameters are calculated. These gradients are used to update the weights with gradient descent. Below you can see an example of calculating the gradients using a backwards pass.
```
# Build a feed-forward network
model = nn.Sequential(nn.Linear(784, 128),
nn.ReLU(),
nn.Linear(128, 64),
nn.ReLU(),
nn.Linear(64, 10),
nn.LogSoftmax(dim=1))
criterion = nn.NLLLoss()
images, labels = next(iter(trainloader))
images = images.view(images.shape[0], -1)
logits = model(images)
loss = criterion(logits, labels)
print('Before backward pass: \n', model[0].weight.grad)
loss.backward()
print('After backward pass: \n', model[0].weight.grad)
```
## Training the network!
There's one last piece we need to start training, an optimizer that we'll use to update the weights with the gradients. We get these from PyTorch's [`optim` package](https://pytorch.org/docs/stable/optim.html). For example we can use stochastic gradient descent with `optim.SGD`. You can see how to define an optimizer below.
```
from torch import optim
# Optimizers require the parameters to optimize and a learning rate
optimizer = optim.SGD(model.parameters(), lr=0.01)
```
Now we know how to use all the individual parts so it's time to see how they work together. Let's consider just one learning step before looping through all the data. The general process with PyTorch:
* Make a forward pass through the network
* Use the network output to calculate the loss
* Perform a backward pass through the network with `loss.backward()` to calculate the gradients
* Take a step with the optimizer to update the weights
Below I'll go through one training step and print out the weights and gradients so you can see how it changes. Note that I have a line of code `optimizer.zero_grad()`. When you do multiple backwards passes with the same parameters, the gradients are accumulated. This means that you need to zero the gradients on each training pass or you'll retain gradients from previous training batches.
```
print('Initial weights - ', model[0].weight)
images, labels = next(iter(trainloader))
images.resize_(64, 784)
# Clear the gradients, do this because gradients are accumulated
optimizer.zero_grad()
# Forward pass, then backward pass, then update weights
output = model(images)
loss = criterion(output, labels)
loss.backward()
print('Gradient -', model[0].weight.grad)
# Take an update step and few the new weights
optimizer.step()
print('Updated weights - ', model[0].weight)
```
### Training for real
Now we'll put this algorithm into a loop so we can go through all the images. Some nomenclature, one pass through the entire dataset is called an *epoch*. So here we're going to loop through `trainloader` to get our training batches. For each batch, we'll doing a training pass where we calculate the loss, do a backwards pass, and update the weights.
>**Exercise:** Implement the training pass for our network. If you implemented it correctly, you should see the training loss drop with each epoch.
```
## Your solution here
model = nn.Sequential(nn.Linear(784, 128),
nn.ReLU(),
nn.Linear(128, 64),
nn.ReLU(),
nn.Linear(64, 10),
nn.LogSoftmax(dim=1))
criterion = nn.NLLLoss()
optimizer = optim.SGD(model.parameters(), lr=0.003)
epochs = 5
for e in range(epochs):
running_loss = 0
for images, labels in trainloader:
# Flatten MNIST images into a 784 long vector
images = images.view(images.shape[0], -1)
# TODO: Training pass
optimizer.zero_grad()
output = model.forward(images)
loss = criterion(output, labels)
loss.backward()
running_loss += loss.item()
optimizer.step()
else:
print(f"Training loss: {running_loss/len(trainloader)}")
```
With the network trained, we can check out it's predictions.
```
%matplotlib inline
import helper
images, labels = next(iter(trainloader))
img = images[0].view(1, 784)
# Turn off gradients to speed up this part
with torch.no_grad():
logps = model(img)
# Output of the network are log-probabilities, need to take exponential for probabilities
ps = torch.exp(logps)
helper.view_classify(img.view(1, 28, 28), ps)
```
Now our network is brilliant. It can accurately predict the digits in our images. Next up you'll write the code for training a neural network on a more complex dataset.
| github_jupyter |
# Expecting the unexpected
To handle errors properly deserves a chapter on its own in any programming book. Python gives us many ways do deal with errors fatal and otherwise: try, except, assert, if ...
Using these mechanisms in a naive way may lead to code that is littered with safety `if` statements and `try`-`except` blocks, just because we need to account for errors at every level in a program.
In this tutorial we'll see how we can use exceptions in a more effective way. As an added bonus we learn how to use exceptions in a manner that is compatible with the Noodles programming model. Let's try something dangerous! We'll compute the reciprocal of a list of numbers. To see what is happening, the function `something_dangerous` contains a print statement.
```
import sys
def something_dangerous(x):
print("computing reciprocal of", x)
return 1 / x
try:
for x in [2, 1, 0, -1]:
print("1/{} = {}".format(x, something_dangerous(x)))
except ArithmeticError as error:
print("Something went terribly wrong:", error)
```
This shows how exceptions are raised and caught, but this approach is somewhat limited. Suppose now, that we weren't expecting this expected unexpected behaviour and we wanted to compute everything before displaying our results.
```
input_list = [2, 1, 0, -1]
reciprocals = [something_dangerous(item)
for item in input_list]
print("The reciprocal of", input_list, "is", reciprocals)
```
Ooops! Let's fix that.
```
try:
reciprocals = [something_dangerous(item)
for item in input_list]
except ArithmeticError as error:
print("Something went terribly wrong:", error)
else:
print("The reciprocal of\n\t", input_list,
"\nis\n\t", reciprocals)
```
That's also not what we want. We wasted all this time computing nice reciprocals of numbers, only to find all of our results being thrown away because of one stupid zero in the input list. We can fix this.
```
import math
def something_safe(x):
try:
return something_dangerous(x)
except ArithmeticError as error:
return math.nan
reciprocals = [something_safe(item)
for item in input_list]
print("The reciprocal of\n\t", input_list,
"\nis\n\t", reciprocals)
```
That's better! We skipped right over the error and continued to more interesting results. So how are we going to make this solution more generic? Subsequent functions may not know how to handle that little `nan` in our list.
```
square_roots = [math.sqrt(item) for item in reciprocals]
```
Hmmmpf. There we go again.
```
def safe_sqrt(x):
try:
return math.sqrt(x)
except ValueError as error:
return math.nan
[safe_sqrt(item) for item in reciprocals]
```
This seems Ok, but there are two problems here. For one, it feels like we're doing too much work! We have a repeating code pattern here. That's always a moment to go back and consider making parts of our code more generic. At the same time, this is when we need some more advanced Python concepts to get us out of trouble. We're going to define a function in a function!
```
def secure_function(dangerous_function):
def something_safe(x):
"""A safer version of something dangerous."""
try:
return dangerous_function(x)
except (ArithmeticError, ValueError):
return math.nan
return something_safe
```
Consider what happens here. The function `secure_function` takes a function `something_dangerous` as an argument and returns a new function `something_safe`. This new function executes `something_dangerous` within a `try`-`except` block to deal with the possibility of failure. Let's see how this works.
```
safe_sqrt = secure_function(math.sqrt)
print("⎷2 =", safe_sqrt(2))
print("⎷-1 =", safe_sqrt(-1))
print()
help(safe_sqrt)
```
Ok, so that works! However, the documentation of `safe_sqrt` is not yet very useful. There is a nice library routine that may help us here: `functools.wraps`; this utility function sets the correct name and doc-string to our new function.
```
import functools
def secure_function(dangerous_function):
"""Create a function that doesn't raise ValueErrors."""
@functools.wraps(dangerous_function)
def something_safe(x):
"""A safer version of something dangerous."""
try:
return dangerous_function(x)
except (ArithmeticError, ValueError):
return math.nan
return something_safe
safe_sqrt = secure_function(math.sqrt)
help(safe_sqrt)
```
Now it is very easy to also rewrite our function computing the reciprocals safely:
```
something_safe = secure_function(something_dangerous)
[safe_sqrt(something_safe(item)) for item in input_list]
```
There is a second problem to this approach, which is a bit more subtle. How do we know where the error occured? We got two values of `nan` and are desperate to find out what went wrong. We'll need a little class to capture all aspects of failure.
```
class Fail:
"""Keep track of failures."""
def __init__(self, exception, trace):
self.exception = exception
self.trace = trace
def extend_trace(self, f):
"""Grow a stack trace."""
self.trace.append(f)
return self
def __str__(self):
return "Fail in " + " -> ".join(
f.__name__ for f in reversed(self.trace)) \
+ ":\n\t" + type(self.exception).__name__ \
+ ": " + str(self.exception)
```
We will adapt our earlier design for `secure_function`. If the given argument is a `Fail`, we don't even attempt to run the next function. In stead, we extend the trace of the failure, so that we can see what happened later on.
```
def secure_function(dangerous_function):
"""Create a function that doesn't raise ValueErrors."""
@functools.wraps(dangerous_function)
def something_safe(x):
"""A safer version of something dangerous."""
if isinstance(x, Fail):
return x.extend_trace(dangerous_function)
try:
return dangerous_function(x)
except Exception as error:
return Fail(error, [dangerous_function])
return something_safe
```
Now we can rewrite our little program entirely from scratch:
```
@secure_function
def reciprocal(x):
return 1 / x
@secure_function
def square_root(x):
return math.sqrt(x)
reciprocals = map(reciprocal, input_list)
square_roots = map(square_root, reciprocals)
for x, result in zip(input_list, square_roots):
print("sqrt( 1 /", x, ") =", result)
```
See how we retain a trace of the functions that were involved in creating the failed state, even though the execution of that produced those values is entirely decoupled. This is exactly what we need to trace errors in Noodles.
## Handling errors in Noodles
Noodles has the functionality of `secure_function` build in by the name of `maybe`. The following code implements the above example in terms of `noodles.maybe`:
```
import noodles
import math
from noodles.tutorial import display_workflows
@noodles.maybe
def reciprocal(x):
return 1 / x
@noodles.maybe
def square_root(x):
return math.sqrt(x)
results = [square_root(reciprocal(x)) for x in [2, 1, 0, -1]]
for result in results:
print(str(result))
```
The `maybe` decorator works well together with `schedule`. The following workflow is full of errors!
```
@noodles.schedule
@noodles.maybe
def add(a, b):
return a + b
workflow = add(noodles.schedule(reciprocal)(0),
noodles.schedule(square_root)(-1))
display_workflows(arithmetic=workflow, prefix='errors')
```
Both the reciprocal and the square root functions will fail. Noodles is smart enough to report on both errors.`
```
result = noodles.run_single(workflow)
print(result)
```
## Example: parallel stat
Let's do an example that works with external processes. The UNIX command `stat` gives the status of a file
```
!stat -t -c '%A %10s %n' /dev/null
```
If a file does note exist, `stat` returns an error-code of 1.
```
!stat -t -c '%A %10s %n' does-not-exist
```
We can wrap the execution of the `stat` command in a helper function.
```
from subprocess import run, PIPE, CalledProcessError
@noodles.schedule
@noodles.maybe
def stat_file(filename):
p = run(['stat', '-t', '-c', '%A %10s %n', filename],
check=True, stdout=PIPE, stderr=PIPE)
return p.stdout.decode().strip()
```
The `run` function runs the given command and returns a `CompletedProcess` object. The `check=True` argument enables checking for return value of the child process. If the return value is any other then 0, a `CalledProcessError` is raised. Because we decorated our function with `noodles.maybe`, such an error will be caught and a `Fail` object will be returned.
```
files = ['/dev/null', 'does-not-exist', '/home', '/usr/bin/python3']
workflow = noodles.gather_all(stat_file(f) for f in files)
display_workflows(stat=workflow, prefix='errors')
```
We can now run this workflow and print the output in a table.
```
result = noodles.run_parallel(workflow, n_threads=4)
for file, stat in zip(files, result):
print('stat {:18} -> {}'.format(
file, stat if not noodles.failed(stat)
else 'failed: ' + stat.exception.stderr.decode().strip()))
```
| github_jupyter |
```
import numpy as np
import keras
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
import os
import pandas as pd
from keras.utils import to_categorical
import matplotlib.pyplot as plt
df_train = pd.read_csv('train.txt', header =None, sep =';', names = ['Input','Sentiment'], encoding='utf-8')
df_test = pd.read_csv('test.txt', header = None, sep =';', names = ['Input','Sentiment'],encoding='utf-8')
df_val=pd.read_csv('val.txt',header=None,sep=';',names=['Input','Sentiment'],encoding='utf-8')
df_train.Sentiment.value_counts()
X=df_train['Input']
lst=[]
for i in X:
lst.append(len(i))
len1=pd.DataFrame(lst)
len1.describe()
cts=[]
for i in range(7,301):
ct=0
for k in lst:
if k==i:
ct+=1
cts.append(ct)
plt.bar(range(7,301),cts)
plt.show()
tokenizer=Tokenizer(15212,lower=True,oov_token='UNK')
tokenizer.fit_on_texts(X)
len(tokenizer.word_index)
X_train=tokenizer.texts_to_sequences(X)
X_train_pad=pad_sequences(X_train,maxlen=80,padding='post')
df_train['Sentiment']=df_train.Sentiment.replace({'joy':0,'anger':1,'love':2,'sadness':3,'fear':4,'surprise':5})
Y_train=df_train['Sentiment'].values
Y_train_f=to_categorical(Y_train)
Y_train_f[:6]
X_val=df_val['Input']
Y_val=df_val.Sentiment.replace({'joy':0,'anger':1,'love':2,'sadness':3,'fear':4,'surprise':5})
X_val_f=tokenizer.texts_to_sequences(X_val)
X_val_pad=pad_sequences(X_val_f,maxlen=80,padding='post')
Y_val_f=to_categorical(Y_val)
Y_val_f[:6]
from keras.models import Sequential
from keras.layers import LSTM,Bidirectional,Dense,Embedding,Dropout
model=Sequential()
model.add(Embedding(15212,64,input_length=80))
model.add(Dropout(0.6))
model.add(Bidirectional(LSTM(80,return_sequences=True)))
model.add(Bidirectional(LSTM(160)))
model.add(Dense(6,activation='softmax'))
print(model.summary())
model.compile(optimizer='adam',loss='categorical_crossentropy',metrics=['accuracy'])
hist=model.fit(X_train_pad,Y_train_f,epochs=12,validation_data=(X_val_pad,Y_val_f))
plt.plot(hist.history['accuracy'],c='b',label='train')
plt.plot(hist.history['val_accuracy'],c='r',label='validation')
plt.legend(loc='lower right')
plt.show()
plt.plot(hist.history['loss'],c='orange',label='train')
plt.plot(hist.history['val_loss'],c='g',label='validation')
plt.legend(loc='upper right')
plt.show()
X_test=df_test['Input']
Y_test=df_test.Sentiment.replace({'joy':0,'anger':1,'love':2,'sadness':3,'fear':4,'surprise':5})
X_test_f=tokenizer.texts_to_sequences(X_test)
X_test_pad=pad_sequences(X_test_f,maxlen=80,padding='post')
Y_test_f=to_categorical(Y_test)
X_test_pad.shape
Y_test_f[:7]
model.evaluate(X_test_pad,Y_test_f)
def get_key(value):
dictionary={'joy':0,'anger':1,'love':2,'sadness':3,'fear':4,'surprise':5}
for key,val in dictionary.items():
if (val==value):
return key
def predict(sentence):
sentence_lst=[]
sentence_lst.append(sentence)
sentence_seq=tokenizer.texts_to_sequences(sentence_lst)
sentence_padded=pad_sequences(sentence_seq,maxlen=80,padding='post')
ans=get_key(model.predict_classes(sentence_padded))
print("The emotion predicted is",ans)
predict(str(input('Enter a sentence : ')))
predict(str(input('Enter a sentence : ')))
```
| github_jupyter |
As the name suggests, Time Series is a collection of data points collected at constant time intervals. These are analyzed to determine the long term trend so as to forecast the future or perform some other form of analysis. But what makes a TS different from say a regular regression problem? There are 2 things:
* It is time dependent. So the basic assumption of a linear regression model that the observations are independent doesn’t hold in this case.
* Along with an increasing or decreasing trend, most TS have some form of seasonality trends, i.e. variations specific to a particular time frame. For example, if you see the sales of a woolen jacket over time, you will invariably find higher sales in winter seasons.
* Because of the inherent properties of a TS, there are various steps involved in analyzing it. These are discussed in detail below. Lets start by loading a TS object in Python. We’ll be using the popular AirPassengers data set.
```
import pandas as pd
import numpy as np
import matplotlib.pylab as plt
%matplotlib inline
from matplotlib.pylab import rcParams
rcParams['figure.figsize'] = 15, 6
data = pd.read_csv('AirPassengers.csv')
print data.head()
print '\n Data Types:'
print data.dtypes
print data.shape
dateparse = lambda dates: pd.datetime.strptime(dates, '%Y-%m')
data = pd.read_csv('AirPassengers.csv', parse_dates=True, index_col='Month',date_parser=dateparse)
print data.head()
data.index
data.shape
ts = data['#Passengers']
ts.head(10)
from datetime import datetime
ts[datetime(1949,1,1)]
ts['1959']
```
#### How to Check Stationarity of a Time Series?
A TS is said to be stationary if its statistical properties such as mean, variance remain constant over time. But why is it important? Most of the TS models work on the assumption that the TS is stationary. Intuitively, we can sat that if a TS has a particular behaviour over time, there is a very high probability that it will follow the same in the future. Also, the theories related to stationary series are more mature and easier to implement as compared to non-stationary series.
Stationarity is defined using very strict criterion. However, for practical purposes we can assume the series to be stationary if it has constant statistical properties over time, ie. the following:
1. constant mean
2. constant variance
3. an autocovariance that does not depend on time.
```
plt.plot(ts)
```
we can check stationarity using the following:
1. Plotting Rolling Statistics: We can plot the moving average or moving variance and see if it varies with time. By moving average/variance I mean that at any instant ‘t’, we’ll take the average/variance of the last year, i.e. last 12 months. But again this is more of a visual technique.
2. Dickey-Fuller Test: This is one of the statistical tests for checking stationarity. Here the null hypothesis is that the TS is non-stationary. The test results comprise of a Test Statistic and some Critical Values for difference confidence levels. If the ‘Test Statistic’ is less than the ‘Critical Value’, we can reject the null hypothesis and say that the series is stationary.
```
from statsmodels.tsa.stattools import adfuller ##dickey fuller test implementation
def test_stationarity(timeseries):
#Determing rolling statistics
rolmean = pd.rolling_mean(timeseries, window=12)
rolstd = pd.rolling_std(timeseries, window=12)
#Plot rolling statistics:
orig = plt.plot(timeseries, color='blue',label='Original')
mean = plt.plot(rolmean, color='red', label='Rolling Mean')
std = plt.plot(rolstd, color='black', label = 'Rolling Std')
plt.legend(loc='best')
plt.title('Rolling Mean & Standard Deviation')
plt.show(block=False)
#Perform Dickey-Fuller test:
print 'Results of Dickey-Fuller Test:'
dftest = adfuller(timeseries, autolag='AIC')
dfoutput = pd.Series(dftest[0:4], index=['Test Statistic','p-value','#Lags Used','Number of Observations Used'])
for key,value in dftest[4].items():
dfoutput['Critical Value (%s)'%key] = value
print dfoutput
test_stationarity(ts)
```
Though stationarity assumption is taken in many TS models, almost none of practical time series are stationary. So statisticians have figured out ways to make series stationary, which we’ll discuss now. Actually, its almost impossible to make a series perfectly stationary, but we try to take it as close as possible.
Lets understand what is making a TS non-stationary. There are 2 major reasons behind non-stationaruty of a TS:
1. Trend – varying mean over time. For eg, in this case we saw that on average, the number of passengers was growing over time.
2. Seasonality – variations at specific time-frames. eg people might have a tendency to buy cars in a particular month because of pay increment or festivals.
The underlying principle is to model or estimate the trend and seasonality in the series and remove those from the series to get a stationary series. Then statistical forecasting techniques can be implemented on this series. The final step would be to convert the forecasted values into the original scale by applying trend and seasonality constraints back.
#### Estimating & Eliminating Trend
One of the first tricks to reduce trend can be transformation. For example, in this case we can clearly see that the there is a significant positive trend. So we can apply transformation which penalize higher values more than smaller values. These can be taking a log, square root, cube root, etc. Lets take a log transform here for simplicity:
```
ts_log = np.log(ts)
plt.plot(ts_log)
```
In this simpler case, it is easy to see a forward trend in the data. But its not very intuitive in presence of noise. So we can use some techniques to estimate or model this trend and then remove it from the series. There can be many ways of doing it and some of most commonly used are:
1. Aggregation – taking average for a time period like monthly/weekly averages
2. Smoothing – taking rolling averages
3. Polynomial Fitting – fit a regression model
Lets try smoothing:
Smoothing refers to taking rolling estimates, i.e. considering the past few instances. There are can be various ways but we will discuss two of those here.
#### Moving average
In this approach, we take average of ‘k’ consecutive values depending on the frequency of time series. Here we can take the average over the past 1 year, i.e. last 12 values. Pandas has specific functions defined for determining rolling statistics.
```
moving_avg = pd.rolling_mean(ts_log,12)
plt.plot(ts_log)
plt.plot(moving_avg, color='red')
##The red line shows the rolling mean.
##Lets subtract this from the original series. Note that since we are taking
##average of last 12 values, rolling mean is not defined for first 11 values. This can be observed as:
ts_log_moving_avg_diff = ts_log - moving_avg
ts_log_moving_avg_diff.head(12)
##Notice the first 11 being Nan. Lets drop these NaN values and check the plots to test stationarity.
ts_log_moving_avg_diff.dropna(inplace=True)
test_stationarity(ts_log_moving_avg_diff)
```
This looks like a much better series. The rolling values appear to be varying slightly but there is no specific trend. Also, the test statistic is smaller than the 5% critical values so we can say with 95% confidence that this is a stationary series.
However, a drawback in this particular approach is that the time-period has to be strictly defined. In this case we can take yearly averages but in complex situations like forecasting a stock price, its difficult to come up with a number. So we take a ‘weighted moving average’ where more recent values are given a higher weight. There can be many technique for assigning weights. A popular one is exponentially weighted moving average where weights are assigned to all the previous values with a decay factor. This can be implemented in Pandas as:
```
expwighted_avg = pd.ewma(ts_log, halflife=12)
plt.plot(ts_log)
plt.plot(expwighted_avg, color='red')
##Note that here the parameter ‘halflife’ is used to define the amount of exponential decay.
##This is just an assumption here and would depend largely on the business domain.
##Other parameters like span and center of mass can also be used to define decay .
##Now, let’s remove this from series and check stationarity:
ts_log_ewma_diff = ts_log - expwighted_avg
test_stationarity(ts_log_ewma_diff)
```
#### Eliminating Trend and Seasonality
The simple trend reduction techniques discussed before don’t work in all cases, particularly the ones with high seasonality. Lets discuss two ways of removing trend and seasonality:
1. Differencing – taking the differece with a particular time lag
2. Decomposition – modeling both trend and seasonality and removing them from the model.
#### Differencing
One of the most common methods of dealing with both trend and seasonality is differencing. In this technique, we take the difference of the observation at a particular instant with that at the previous instant. This mostly works well in improving stationarity. First order differencing can be done in Pandas as:
```
ts_log_diff = ts_log - ts_log.shift()
plt.plot(ts_log_diff)
ts_log_diff.dropna(inplace=True)
test_stationarity(ts_log_diff)
```
We can see that the mean and std variations have small variations with time. Also, the Dickey-Fuller test statistic is less than the 10% critical value, thus the TS is stationary with 90% confidence. We can also take second or third order differences which might get even better results in certain applications.
#### Decomposing
In this approach, both trend and seasonality are modeled separately and the remaining part of the series is returned.
```
from statsmodels.tsa.seasonal import seasonal_decompose
decomposition = seasonal_decompose(ts_log)
trend = decomposition.trend
seasonal = decomposition.seasonal
residual = decomposition.resid
plt.subplot(411)
plt.plot(ts_log, label='Original')
plt.legend(loc='best')
plt.subplot(412)
plt.plot(trend, label='Trend')
plt.legend(loc='best')
plt.subplot(413)
plt.plot(seasonal,label='Seasonality')
plt.legend(loc='best')
plt.subplot(414)
plt.plot(residual, label='Residuals')
plt.legend(loc='best')
plt.tight_layout()
##Lets check stationarity of residuals:
ts_log_decompose = residual
ts_log_decompose.dropna(inplace=True)
test_stationarity(ts_log_decompose)
```
### Forecasting a time series:
We saw different techniques and all of them worked reasonably well for making the TS stationary. Lets make model on the TS after differencing as it is a very popular technique. Also, its relatively easier to add noise and seasonality back into predicted residuals in this case. Having performed the trend and seasonality estimation techniques, there can be two situations:
1. A strictly stationary series with no dependence among the values. This is the easy case wherein we can model the residuals as white noise. But this is very rare.
2. A series with significant dependence among values. In this case we need to use some statistical models like ARIMA to forecast the data.
ARIMA stands for Auto-Regressive Integrated Moving Averages. The ARIMA forecasting for a stationary time series is nothing but a linear (like a linear regression) equation. The predictors depend on the parameters (p,d,q) of the ARIMA model:
1. Number of AR (Auto-Regressive) terms (p): AR terms are just lags of dependent variable. For instance if p is 5, the predictors for x(t) will be x(t-1)….x(t-5).
2. Number of MA (Moving Average) terms (q): MA terms are lagged forecast errors in prediction equation. For instance if q is 5, the predictors for x(t) will be e(t-1)….e(t-5) where e(i) is the difference between the moving average at ith instant and actual value.
3. Number of Differences (d): These are the number of nonseasonal differences, i.e. in this case we took the first order difference. So either we can pass that variable and put d=0 or pass the original variable and put d=1. Both will generate same results.
An important concern here is how to determine the value of ‘p’ and ‘q’. We use two plots to determine these numbers. Lets discuss them first.
1. Autocorrelation Function (ACF): It is a measure of the correlation between the the TS with a lagged version of itself. For instance at lag 5, ACF would compare series at time instant ‘t1’…’t2’ with series at instant ‘t1-5’…’t2-5’ (t1-5 and t2 being end points).
2. Partial Autocorrelation Function (PACF): This measures the correlation between the TS with a lagged version of itself but after eliminating the variations already explained by the intervening comparisons. Eg at lag 5, it will check the correlation but remove the effects already explained by lags 1 to 4.
The ACF and PACF plots for the TS after differencing can be plotted as:
(for more understanding on ARMA and ACF and PACF https://www.analyticsvidhya.com/blog/2015/12/complete-tutorial-time-series-modeling/)
How to determine if it is AR or MA .
Plot ACF and PACF , if it cuts below threshhold in PACF then it is AR model.
If it cuts below threshold in ACF it is MA model
```
#ACF and PACF plots:
from statsmodels.tsa.stattools import acf, pacf
lag_acf = acf(ts_log_diff, nlags=20)
lag_pacf = pacf(ts_log_diff, nlags=20, method='ols')
#Plot ACF:
plt.subplot(121)
plt.plot(lag_acf)
plt.axhline(y=0,linestyle='--',color='gray')
plt.axhline(y=-1.96/np.sqrt(len(ts_log_diff)),linestyle='--',color='gray')
plt.axhline(y=1.96/np.sqrt(len(ts_log_diff)),linestyle='--',color='gray')
plt.title('Autocorrelation Function')
#Plot PACF:
plt.subplot(122)
plt.plot(lag_pacf)
plt.axhline(y=0,linestyle='--',color='gray')
plt.axhline(y=-1.96/np.sqrt(len(ts_log_diff)),linestyle='--',color='gray')
plt.axhline(y=1.96/np.sqrt(len(ts_log_diff)),linestyle='--',color='gray')
plt.title('Partial Autocorrelation Function')
plt.tight_layout()
```
In this plot, the two dotted lines on either sides of 0 are the confidence interevals. These can be used to determine the ‘p’ and ‘q’ values as:
* p – The lag value where the PACF chart crosses the upper confidence interval for the first time. If you notice closely, in this case p=2.
* q – The lag value where the ACF chart crosses the upper confidence interval for the first time. If you notice closely, in this case q=2.
Now, lets make 3 different ARIMA models considering individual as well as combined effects. We will also print the RSS for each. Please note that here RSS is for the values of residuals and not actual series.
We need to load the ARIMA model first:
```
from statsmodels.tsa.arima_model import ARIMA
##AR model
model = ARIMA(ts_log, order=(2, 1, 0))
results_AR = model.fit(disp=-1)
plt.plot(ts_log_diff)
plt.plot(results_AR.fittedvalues, color='red')
plt.title('RSS: %.4f'% sum((results_AR.fittedvalues-ts_log_diff)**2))
##MA model
model = ARIMA(ts_log, order=(0, 1, 2))
results_MA = model.fit(disp=-1)
plt.plot(ts_log_diff)
plt.plot(results_MA.fittedvalues, color='red')
plt.title('RSS: %.4f'% sum((results_MA.fittedvalues-ts_log_diff)**2))
## combined model
model = ARIMA(ts_log, order=(2, 1, 2))
results_ARIMA = model.fit(disp=-1)
plt.plot(ts_log_diff)
plt.plot(results_ARIMA.fittedvalues, color='red')
plt.title('RSS: %.4f'% sum((results_ARIMA.fittedvalues-ts_log_diff)**2))
predictions_ARIMA_diff = pd.Series(results_ARIMA.fittedvalues, copy=True)
print predictions_ARIMA_diff.head()
```
Notice that these start from ‘1949-02-01’ and not the first month. Why? This is because we took a lag by 1 and first element doesn’t have anything before it to subtract from. The way to convert the differencing to log scale is to add these differences consecutively to the base number. An easy way to do it is to first determine the cumulative sum at index and then add it to the base number. The cumulative sum can be found as:
```
predictions_ARIMA_diff_cumsum = predictions_ARIMA_diff.cumsum()
print predictions_ARIMA_diff_cumsum.head()
```
Next we’ve to add them to base number. For this lets create a series with all values as base number and add the differences to it. This can be done as:
```
predictions_ARIMA_log = pd.Series(ts_log.ix[0], index=ts_log.index)
predictions_ARIMA_log = predictions_ARIMA_log.add(predictions_ARIMA_diff_cumsum,fill_value=0)
predictions_ARIMA_log.head()
```
Here the first element is base number itself and from thereon the values cumulatively added. Last step is to take the exponent and compare with the original series.
```
predictions_ARIMA = np.exp(predictions_ARIMA_log)
plt.plot(ts)
plt.plot(predictions_ARIMA)
plt.title('RMSE: %.4f'% np.sqrt(sum((predictions_ARIMA-ts)**2)/len(ts)))
```
| github_jupyter |
# Computational and Numerical Methods
## Group 16
### Set 11 (08-10-2018): Nonlinear Systems and the Newton Method
#### Vidhin Parmar 201601003
#### Parth Shah 201601086
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
import scipy
import scipy.linalg as la
from sympy import symbols, diff
from sympy import *
import mpmath
```
$x^2+4y^2-9=0$
$18y−14x^2+ 45 = 0$
```
th = np.arange(0, 2*np.pi + np.pi/50, np.pi/50)
plt.plot(3*np.cos(th), 1.5*np.sin(th), label="x^2+y^2=9")
t = np.arange(-5, 5 + 0.01, 0.01)
plt.plot((9/14)*t, (-2.5 + ((9/28)*(t**2))), label="18y - 14x^2 +45 =0")
plt.legend(loc="best")
plt.grid(True)
plt.show()
x,y = symbols('x y')
f = lambda a,b: a**2 + 4*b**2 - 9
g = lambda a,b: -14*a**2 + 18*b + 45
fx = lambdify((x,y), diff(x**2 + 4*y**2 - 9, x), "numpy")
fy = lambdify((x,y), diff(x**2 + 4*y**2 - 9, y), "numpy")
gx = lambdify((x,y), diff(-14*x**2 + 18*y + 45, x), "numpy")
gy = lambdify((x,y), diff(-14*x**2 + 18*y + 45, y), "numpy")
x0, y0 = 1, 1
for i in range(25):
mat1 = np.array([[fx(x0,y0), fy(x0,y0)], [gx(x0,y0), gy(x0,y0)]], dtype=np.float32)
mat2 = np.array([[-f(x0,y0)], [-g(x0,y0)]], dtype=np.float32)
ans = la.solve(mat1, mat2)
x0, y0 = x0+ans[0,0], y0+ans[1,0]
"Root1 : ", x0, y0
x,y = symbols('x y')
f = lambda a,b: a**2 + 4*b**2 - 9
g = lambda a,b: -14*a**2 + 18*b + 45
fx = lambdify((x,y), diff(x**2 + 4*y**2 - 9, x), "numpy")
fy = lambdify((x,y), diff(x**2 + 4*y**2 - 9, y), "numpy")
gx = lambdify((x,y), diff(-14*x**2 + 18*y + 45, x), "numpy")
gy = lambdify((x,y), diff(-14*x**2 + 18*y + 45, y), "numpy")
x0, y0 = -1, 1
for i in range(25):
mat1 = np.array([[fx(x0,y0), fy(x0,y0)], [gx(x0,y0), gy(x0,y0)]], dtype=np.float32)
mat2 = np.array([[-f(x0,y0)], [-g(x0,y0)]], dtype=np.float32)
ans = la.solve(mat1, mat2)
x0, y0 = x0+ans[0,0], y0+ans[1,0]
"Root2 : ", x0, y0
x,y = symbols('x y')
f = lambda a,b: a**2 + 4*b**2 - 9
g = lambda a,b: -14*a**2 + 18*b + 45
fx = lambdify((x,y), diff(x**2 + 4*y**2 - 9, x), "numpy")
fy = lambdify((x,y), diff(x**2 + 4*y**2 - 9, y), "numpy")
gx = lambdify((x,y), diff(-14*x**2 + 18*y + 45, x), "numpy")
gy = lambdify((x,y), diff(-14*x**2 + 18*y + 45, y), "numpy")
x0, y0 = -1, -1
for i in range(25):
mat1 = np.array([[fx(x0,y0), fy(x0,y0)], [gx(x0,y0), gy(x0,y0)]], dtype=np.float32)
mat2 = np.array([[-f(x0,y0)], [-g(x0,y0)]], dtype=np.float32)
ans = la.solve(mat1, mat2)
x0, y0 = x0+ans[0,0], y0+ans[1,0]
"Root3 : ", x0, y0
x,y = symbols('x y')
f = lambda a,b: a**2 + 4*b**2 - 9
g = lambda a,b: -14*a**2 + 18*b + 45
fx = lambdify((x,y), diff(x**2 + 4*y**2 - 9, x), "numpy")
fy = lambdify((x,y), diff(x**2 + 4*y**2 - 9, y), "numpy")
gx = lambdify((x,y), diff(-14*x**2 + 18*y + 45, x), "numpy")
gy = lambdify((x,y), diff(-14*x**2 + 18*y + 45, y), "numpy")
x0, y0 = 1, -1
for i in range(25):
mat1 = np.array([[fx(x0,y0), fy(x0,y0)], [gx(x0,y0), gy(x0,y0)]], dtype=np.float32)
mat2 = np.array([[-f(x0,y0)], [-g(x0,y0)]], dtype=np.float32)
ans = la.solve(mat1, mat2)
x0, y0 = x0+ans[0,0], y0+ans[1,0]
"Root4 : ", x0, y0
```
| github_jupyter |
# Some Random SymPy Stuff
```
import sympy as sp
sp.init_printing()
# for older SymPy versions:
#%load_ext sympy.interactive.ipythonprinting
# for even older IPython/SymPy versions:
#%load_ext sympyprinting
x, y = sp.symbols("x y")
(sp.pi + x) ** 2
expr = ((x + y)**2 * (x + 1))
expr
sp.expand(expr)
sp.diff(sp.cos(x**2)**2 / (1 + x), x)
x = sp.symbols('x', positive=True)
x > 0
sp.sqrt(x ** 2)
sp.sqrt(y ** 2)
sp.exp(sp.I * sp.pi)
sp.oo
a, b, c = sp.symbols("a, b, c")
sp.expand(sp.sin(a + b))
sp.expand(sp.sin(a + b), trig=True)
sp.simplify(sp.sin(x)**2 + sp.cos(x)**2)
sp.factor(x**3 + 6 * x**2 + 11*x + 6)
sp.simplify(sp.cos(x) / sp.sin(x))
expr1 = 1 / ((a + 1) * (a + 2))
expr1
expr1.is_rational_function()
sp.apart(expr1)
z = sp.symbols('z')
f = sp.sin(x * y) + sp.cos(y * z)
f
sp.integrate(f, x)
sp.integrate(f, y)
sp.integrate(sp.exp(-x**2), x)
sp.integrate(sp.exp(-x**2), (x, -sp.oo, sp.oo))
sp.integrate(x + sp.sinh(x), x)
sp.diff(_, x)
sp.limit(sp.sin(sp.pi * x) / x, x, 0)
4 * sp.atan(1)
n = sp.symbols('n')
expr = (-1)**n / (2 * n + 1)
expr
sp.summation(expr, (n, 0, sp.oo))
sp.series(sp.atan(x), x, n=13)
expr = 1 / (x + y)
expr
expr.series(x)
sp.series(sp.exp(x), x)
sp.summation(1 / sp.factorial(n), (n, 0, sp.oo))
expr = (1 + 1/n) ** n
expr
sp.limit(expr, n, sp.oo)
sp.pi.evalf(100)
sp.solve(x**4 - x**2 - 1, x)
sp.solve([x + y - 1, x - y - 1], [x,y])
sp.solve([x + y - a, x - y - c], [x,y])
sp.symbols('m:q')
sp.symbols('r:5')
m = sp.symbols('m')
M = sp.Matrix([[1, m], [3, 4]])
M
M.inv()
A = sp.Matrix(sp.symbols("m:2:2")).reshape(2, 2)
A
A[1, 0]
b = sp.Matrix(sp.symbols("b:2"))
b
A**2
A * b
A.det()
A.inv()
```
from https://github.com/sympy/sympy/wiki/Matrices-eigenvalues
```
M = sp.Matrix(4, 4, lambda i, j: i * j + 1)
M
M.eigenvals()
M = sp.Matrix(4, 4, lambda i, j: i * j + 2)
M
M.eigenvals()
M = sp.Matrix(4, 4, lambda i, j: i * j + 3)
M
M.eigenvals()
x = sp.symbols('x')
M = sp.Matrix(4, 4, lambda i, j: i * j + x)
M
M.eigenvals()
```
from https://groups.google.com/forum/#!topic/sympy/mISLFQcEUIM
```
f = sp.Piecewise((0, x <= -1), (sp.Rational(1, 2), x < 1), (0, True))
f
#sp.integrate(f.subs(x, x - y) * f(x), y)
sp.integrate(sp.Heaviside(x + 1) / 2 - sp.Heaviside(x - 1) / 2, (x, -10, -5))
q = sp.Piecewise((0, x < 0), (x, x < 1), (0, True))
q
q.args
```
from https://groups.google.com/forum/#!topic/sympy/tt8IA3y2sfQ
```
f = sp.Lambda(x, sp.exp(-x**2))
f
conv = sp.Integral(f(x-y)*f(y), (y, -sp.oo, sp.oo))
conv
conv.subs(x, 0)
l = sp.lambdify(x, sp.Integral(f(x), (x, -sp.oo, sp.oo)))
#l(3) # doesn't work!
l
l = sp.lambdify(x, sp.Integral(f(x), (x, -sp.oo, sp.oo)), "sympy")
l
l(x)
```
from http://www.mare.ee/indrek/misc/convolution.pdf
```
# Convolute two "one-piece" functions. Arguments F and G
# are tuples in form (h(x) , a_h, b_h), where h(x) is
# the function and [a_h, b_h) is the range where the functions
# are non-zero.
def convolute_onepiece(x, F, G):
f, a_f, b_f = F
g, a_g, b_g = G
f = sp.S(f)
g = sp.S(g)
# make sure ranges are in order, swap values if necessary
if b_f - a_f > b_g - a_g:
f, a_f, b_f, g, a_g, b_g = g, a_g, b_g, f, a_f, b_f
y = sp.Dummy('y')
i = sp.integrate(f.subs(x, y) * g.subs(x, x-y), y)
return [
(i.subs(y, x-a_g) - i.subs(y, a_f), a_f+a_g, b_f+a_g),
(i.subs(y, b_f) - i.subs(y, a_f), b_f+a_g, a_f+b_g),
(i.subs(y, b_f) - i.subs(y, x-b_g), a_f+b_g, b_f+b_g) ]
# Two "flat" functions, uniform centered PDFs
F = 0.5, -1, 1
G = 0.05, -10, 10
convolute_onepiece(sp.abc.x, F, G)
```
from https://github.com/fperez/sympy/blob/master/sympy/solvers/tests/test_pde.py
```
from sympy import Derivative as D
x, y, z, t = sp.symbols("x y z t")
c = sp.symbols("c", real=True)
Phi = sp.Function('Phi')
F, R, T, X, Y, Z, u = map(sp.Function, 'FRTXYZu')
r, theta, z = sp.symbols('r theta z')
eq = sp.Eq(D(F(x, y, z), x) + D(F(x, y, z), y) + D(F(x, y, z), z))
eq
sp.pde_separate_mul(eq, F(x, y, z), [Y(y), u(x, z)])
sp.pde_separate_mul(eq, F(x, y, z), [X(x), Y(y), Z(z)])
# wave equation
wave = sp.Eq(D(u(x, t), t, t), c**2*D(u(x, t), x, x))
wave
sp.pde_separate_mul(wave, u(x, t), [X(x), T(t)])
# Laplace equation in cylindrical coords
eq = sp.Eq(1/r * D(Phi(r, theta, z), r) + D(Phi(r, theta, z), r, 2) +
1/r**2 * D(Phi(r, theta, z), theta, 2) + D(Phi(r, theta, z), z, 2))
eq
# Separate z
sp.pde_separate_mul(eq, Phi(r, theta, z), [Z(z), u(theta, r)])
# Lets use the result to create a new equation...
eq = sp.Eq(_[1], c)
eq
# ...and separate theta...
sp.pde_separate_mul(eq, u(theta, r), [T(theta), R(r)])
# ...or r...
sp.pde_separate_mul(eq, u(theta, r), [R(r), T(theta)])
```
<p xmlns:dct="http://purl.org/dc/terms/">
<a rel="license"
href="http://creativecommons.org/publicdomain/zero/1.0/">
<img src="http://i.creativecommons.org/p/zero/1.0/88x31.png" style="border-style: none;" alt="CC0" />
</a>
<br />
To the extent possible under law,
<span rel="dct:publisher" resource="[_:publisher]">the person who associated CC0</span>
with this work has waived all copyright and related or neighboring
rights to this work.
</p>
| github_jupyter |
```
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import classification_report,confusion_matrix
import numpy as np
import plotly.offline as py
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import StandardScaler
py.init_notebook_mode(connected=True)
import warnings
warnings.simplefilter('ignore')
train = pd.read_csv("Train_v2.csv")
test = pd.read_csv("Test_v2.csv")
test1 = pd.read_csv('Test_v2.csv')
train.info()
train.isnull().sum()
test.isnull().sum()
train.country.value_counts()
train.head()
def preprocess_data(data) :
dic1 = {"Yes" : 1 , "No" : 0}
dic2 = {"Rural" : 0 , "Urban" : 1}
data["location_type"]=data["location_type"].map(dic2)
data["cellphone_access"]=data["cellphone_access"].map(dic1)
one_hot = pd.get_dummies(data["gender_of_respondent"])
data = data.drop("gender_of_respondent",axis=1)
data = data.join(one_hot)
one_hot = pd.get_dummies(data["country"])
data = data.drop("country",axis=1)
data = data.join(one_hot)
le = LabelEncoder()
data['relationship_with_head']=le.fit_transform(data['relationship_with_head'])
data['marital_status']=le.fit_transform(data["marital_status"])
data['education_level']=le.fit_transform(data["education_level"])
data['job_type']=le.fit_transform(data['job_type'])
data = data.drop('uniqueid',axis=1)
data.loc[ data['age_of_respondent'] <= 16, 'age_of_respondent'] = 0
data.loc[(data['age_of_respondent'] > 16) & (data['age_of_respondent'] <= 32), 'age_of_respondent'] = 1
data.loc[(data['age_of_respondent'] > 32) & (data['age_of_respondent'] <= 48), 'age_of_respondent'] = 2
data.loc[(data['age_of_respondent'] > 48) & (data['age_of_respondent'] <= 64), 'age_of_respondent'] = 3
data.loc[ data['age_of_respondent'] > 64, 'age_of_respondent'] = 4
return data
def preprocess_data2(data) :
dic1 = {"Yes" : 1 , "No" : 0}
dic2 = {"Rural" : 0 , "Urban" : 1}
data["location_type"]=data["location_type"].map(dic2)
data["cellphone_access"]=data["cellphone_access"].map(dic1)
one_hot = pd.get_dummies(data["gender_of_respondent"])
data = data.drop("gender_of_respondent",axis=1)
data = data.join(one_hot)
one_hot = pd.get_dummies(data["country"])
data = data.drop("country",axis=1)
data = data.join(one_hot)
one_hot = pd.get_dummies(data["relationship_with_head"])
data = data.drop("relationship_with_head",axis=1)
data = data.join(one_hot)
one_hot = pd.get_dummies(data["marital_status"])
data = data.drop("marital_status",axis=1)
data = data.join(one_hot)
one_hot = pd.get_dummies(data["education_level"])
data = data.drop("education_level",axis=1)
data = data.join(one_hot)
one_hot = pd.get_dummies(data["job_type"])
data = data.drop("job_type",axis=1)
data = data.join(one_hot)
data = data.drop('uniqueid',axis=1)
data.loc[ data['age_of_respondent'] <= 16, 'age_of_respondent'] = 0
data.loc[(data['age_of_respondent'] > 16) & (data['age_of_respondent'] <= 32), 'age_of_respondent'] = 1
data.loc[(data['age_of_respondent'] > 32) & (data['age_of_respondent'] <= 48), 'age_of_respondent'] = 2
data.loc[(data['age_of_respondent'] > 48) & (data['age_of_respondent'] <= 64), 'age_of_respondent'] = 3
data.loc[ data['age_of_respondent'] > 64, 'age_of_respondent'] = 4
return data
def feature_engineering(data) :
data['Has_Education'] = data["education_level"].apply(
lambda x: 1 if x=='Primary education' or x=='Secondary education' or x=='Tertiary education' else 0)
return data
def feature_engineering2 (data) :
data['works_for_gov'] = data["job_type"].apply(
lambda x: 1 if x=='Formally employed Government' or x=='Government Dependent' else 0)
return data
def cleaning_marital_status(data) :
train_wrong=data.loc[(data["household_size"]==1) & (data['marital_status']=='Married/Living together') ]
data= pd.concat([data,train_wrong,train_wrong]).drop_duplicates(keep=False)
return data
train_cleaned = feature_engineering(train)
train_cleaned = feature_engineering2(train_cleaned)
#train_cleaned = cleaning_marital_status(train_cleaned)
train_cleaned = preprocess_data(train_cleaned)
train_cleaned.head()
train_cleaned2 = preprocess_data2(train)
train_cleaned2["bank_account"]=train_cleaned2["bank_account"].map(dic1)
train_cleaned2.head()
dic1 = {"Yes" : 1 , "No" : 0}
train_cleaned["bank_account"]=train_cleaned["bank_account"].map(dic1)
#test_cleaned = feature_engineering(test)
#test_cleaned = feature_engineering2(test_cleaned)
#test_cleaned = cleaning_marital_status(test_cleaned)
test_cleaned = preprocess_data(test)
test_cleaned.head()
test_cleaned2 = preprocess_data2(test)
test_cleaned2.head()
```
# Manually
```
dic3 = {"Kenya" : 3 , "Rwanda" : 2 , "Tanzania" : 1 , "Uganda" : 0 }
dic4 = {"Primary education" : 5, "Secondary education" : 4 ,"Tertiary education" : 3,"Vocational/Specialised training" :2,"No formal education" :1 ,"Other/Dont know/RTA" : 0}
dic5 = {"Married/Living together" : 4,"Single/Never Married" :3,"Widowed":2 ,"Divorced/Seperated" :1 ,"Dont know" : 0}
dic6 = {"Head of Household":5,"Spouse":4,"Child":3,"Other relative":2,"Parent":1,"Other non-relatives":0}
dic7 = {"Self employed":9,"Farming and Fishing":8,"Formally employed Private":7,"Informally employed":6,"Formally employed Government":5,"Remittance Dependent":4,"Other Income":3,"Government Dependent":2,"Dont Know/Refuse to answer":1,"No Income":0}
train['country']=train['country'].map(dic3)
train['education_level']=train['education_level'].map(dic4)
train['marital_status']=train['marital_status'].map(dic5)
train['relationship_with_head'] = train['relationship_with_head'].map(dic6)
train['job_type']=train['job_type'].map(dic7)
test['country']=test['country'].map(dic3)
test['education_level']=test['education_level'].map(dic4)
test['marital_status']=test['marital_status'].map(dic5)
test['relationship_with_head'] = test['relationship_with_head'].map(dic6)
test['job_type']=test['job_type'].map(dic7)
def plot_correlation_map( df ):
corr = df.corr()
s , ax = plt.subplots( figsize =( 20 , 20 ) )
cmap = sns.diverging_palette( 175, 20 , as_cmap = True )
s = sns.heatmap(
corr,
cmap = cmap,
square=True,
cbar_kws={ 'shrink' : .9 },
ax=ax,
annot = True,
annot_kws = { 'fontsize' : 10 }
)
plot_correlation_map(train_cleaned)
sorted(train_cleaned.corr()["bank_account"].iteritems(), key=lambda x:abs(x[1]), reverse=True)[1:9]
X= train_cleaned[["year","location_type","cellphone_access","household_size","age_of_respondent","relationship_with_head","marital_status","education_level","job_type","Female","Male","Kenya","Tanzania","Rwanda","Uganda","Has_Education","works_for_gov"]]
y=train_cleaned["bank_account"]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.30, random_state=30)
X_train, X_test, y_train, y_test = train_test_split(
train_cleaned2.drop(columns=["bank_account"]),
train_cleaned2["bank_account"],
test_size=0.2,
random_state=20)
from sklearn.linear_model import LogisticRegression
logreg = LogisticRegression()
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train)
X_test_scaled = scaler.transform(X_test)
logreg.fit(X_train_scaled, y_train)
y_pred = logreg.predict(X_test_scaled)
from sklearn.metrics import classification_report
print(classification_report(y_test, y_pred))
print('Accuracy = {:.2f}'.format(logreg.score(X_test_scaled, y_test)))
def plot_confusion_matrix(cm,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix')
print(cm)
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
import itertools
cnfe = confusion_matrix(y_test,y_pred)
plt.figure()
plot_confusion_matrix(cnfe,
title='Confusion matrix')
plt.show()
test_scaled = scaler.transform(test_cleaned2)
predictions = logreg.predict(test_scaled)
data_predictions = pd.DataFrame(predictions,columns=['bank_account'])
data_predictions.insert(0,'unique_id',test1['uniqueid'])
data_predictions["unique_id"]=data_predictions["unique_id"]+' x '+test1['country']
data_predictions.to_csv("Mysubmission1.csv",index=False)
from sklearn import model_selection
from sklearn.model_selection import cross_val_score
kfold = model_selection.KFold(n_splits=10, random_state=5)
modelCV = LogisticRegression()
scoring = 'accuracy'
results = model_selection.cross_val_score(modelCV, X_train, y_train, cv=kfold, scoring=scoring)
print("10-fold cross validation average accuracy: %.3f" % (results.mean()))
from sklearn.neighbors import KNeighborsClassifier
from sklearn.metrics import accuracy_score
knn = KNeighborsClassifier(n_neighbors=17)
knn.fit(X_train, y_train)
y_pred = knn.predict(X_test)
print("Accuracy = ",accuracy_score(y_pred,y_test))
from sklearn.neighbors import KNeighborsClassifier
from sklearn.metrics import accuracy_score
scores=[]
k_range= range(1,30)
for i in k_range :
knn = KNeighborsClassifier(n_neighbors=i)
knn.fit(X_train, y_train)
y_pred = knn.predict(X_test)
scores.append(accuracy_score(y_pred,y_test))
plt.plot(k_range,scores)
rfc = RandomForestClassifier(n_estimators=350)
rfc = rfc.fit(X_train, y_train)
rfc_pred = rfc.predict(X_test)
from sklearn.metrics import accuracy_score
from sklearn.metrics import classification_report,confusion_matrix
print(classification_report(y_test,rfc_pred))
print("accuracy = ",accuracy_score(rfc_pred,y_test))
for name, score in zip(X_train, rfc.feature_importances_):
print(name, score)
rfc_predic = rfc.predict(test)
data_predictions1 = pd.DataFrame(rfc_predic,columns=['bank_account'])
data_predictions1.insert(0,'unique_id',test1['uniqueid'])
data_predictions1["unique_id"]=data_predictions1["unique_id"]+' x '+test1['country']
data_predictions1.to_csv("Mysubmission2.csv",index=False)
from sklearn import tree
dtc = tree.DecisionTreeClassifier(max_depth=3)
dtc = dtc.fit(X_train,y_train)
from sklearn.metrics import accuracy_score
dtc_pred = dtc.predict(X_test)
from sklearn.metrics import classification_report,confusion_matrix
print(classification_report(y_test,dtc_pred))
print("Accuracy = ",accuracy_score(dtc_pred,y_test))
predictions1 = dtc.predict(test)
data_predictions1 = pd.DataFrame(predictions1,columns=['bank_account'])
data_predictions1.insert(0,'unique_id',test1['uniqueid'])
data_predictions1["unique_id"]=data_predictions1["unique_id"]+' x '+test1['country']
data_predictions1.to_csv("Mysubmission2.csv",index=False)
```
# XGBOOST
```
import xgboost as xgb
XGB= xgb.XGBClassifier()
XGB.fit(X_train_scaled,y_train)
pred1=XGB.predict(X_test_scaled)
print ('Accuracy ={:.2f}'.format(XGB.score(X_test_scaled,y_test)))
xgb_pred = XGB.predict(test_scaled)
xgb_pred = pd.DataFrame(xgb_pred,columns=['bank_account'])
xgb_pred.insert(0,'unique_id',test1['uniqueid'])
xgb_pred["unique_id"]=xgb_pred["unique_id"]+' x '+test1['country']
xgb_pred.to_csv("MysubmissionXGB.csv",index=False)
from sklearn.ensemble import GradientBoostingClassifier
learning_rates = [0.05, 0.1,0.15,0.2, 0.25,0.3,0.35,0.4,0.45, 0.5,0.55,0.6,0.65,0.7, 0.75,0.8,0.85,0.9,0.95, 1,1.05,1.1,1.15,1.2,1.25]
for learning_rate in learning_rates:
gb = GradientBoostingClassifier(n_estimators=20, learning_rate = learning_rate, max_features=2, max_depth = 2, random_state = 0)
gb.fit(X_train, y_train)
print("Learning rate: ", learning_rate)
print("Accuracy score (training): {0:.3f}".format(gb.score(X_train, y_train)))
print("Accuracy score (validation): {0:.3f}".format(gb.score(X_test, y_test)))
print()
```
# ADA Boost
```
from sklearn.ensemble import AdaBoostClassifier
ada = AdaBoostClassifier(n_estimators=500)
ada = ada.fit(X_train_scaled,y_train)
ada_pred = ada.predict(X_test_scaled)
ada.score(X_test_scaled,y_test)
from sklearn.ensemble import AdaBoostClassifier
ada = AdaBoostClassifier(n_estimators=500)
ada = ada.fit(X,y)
ada_pred = ada.predict(test)
ada_predictions1 = pd.DataFrame(ada_pred,columns=['bank_account'])
ada_predictions1.insert(0,'unique_id',test1['uniqueid'])
ada_predictions1["unique_id"]=ada_predictions1["unique_id"]+' x '+test1['country']
ada_predictions1.to_csv("Mysubmission3.csv",index=False)
from sklearn.ensemble import BaggingClassifier
bagging = BaggingClassifier(logreg)
bagging = bagging.fit(X_train,y_train)
bagging.score(X_test,y_test)
```
# MLP
```
from sklearn.neural_network import MLPClassifier
mlp = MLPClassifier()
mlp = mlp.fit(X_train,y_train)
mlp_pred = mlp.predict(X_test)
mlp.score(X_test,y_test)
```
# EXTRA TREES CLASSIFIER
```
from sklearn.ensemble import ExtraTreesClassifier
etc = ExtraTreesClassifier(n_estimators=200)
etc = etc.fit(X_train,y_train)
etc.score(X_test,y_test)
```
# LDA
```
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis as LDA
lda = LDA(n_components=1)
X_train = lda.fit_transform(X_train, y_train)
X_test = lda.transform(X_test)
test.columns
test = lda.transform(test)
```
# PCA
```
features=["country","year","location_type","cellphone_access","household_size","age_of_respondent","relationship_with_head","marital_status","education_level","job_type","Female","Male"]
x = train.loc[:,features].values
y = train.loc[:,['bank_account']].values
x = scaler.fit_transform(x)
from sklearn.decomposition import PCA
pca = PCA(n_components=12)
principalComponents = pca.fit_transform(x)
principalDf = pd.DataFrame(data = principalComponents)
finalDf = pd.concat([principalDf,train['bank_account']],axis= 1)
x = test.loc[:,features].values
x= scaler.fit_transform(x)
x_pca = pca.fit_transform(x)
X_train, X_test, y_train, y_test = train_test_split(principalDf, train['bank_account'], test_size=0.30, random_state=30)
from sklearn.ensemble import AdaBoostClassifier
ada = AdaBoostClassifier(n_estimators=100)
ada = ada.fit(X_train,y_train)
ada_pred = ada.predict(X_test)
ada.score(X_test,y_test)
cnfe = confusion_matrix(y_test,y_pred)
plt.figure()
plot_confusion_matrix(cnfe,
title='Confusion matrix')
plt.show()
ada_pca = ada.predict(x_pca)
ada_pred = ada.predict(test)
ada_predictions1 = pd.DataFrame(ada_pred,columns=['bank_account'])
ada_predictions1.insert(0,'unique_id',test1['uniqueid'])
ada_predictions1["unique_id"]=ada_predictions1["unique_id"]+' x '+test1['country']
ada_predictions1.to_csv("Mysubmission4.csv",index=False)
sns.boxplot(data=train)
plt.xticks(rotation=-90)
```
# SMOTE
```
from imblearn.over_sampling import SMOTE
sm = SMOTE(random_state=2)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.30, random_state=30)
X_res, y_res = sm.fit_sample(X_train, y_train.ravel())
import numpy as np
from sklearn.model_selection import GridSearchCV
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import confusion_matrix, precision_recall_curve, auc, roc_auc_score, roc_curve, recall_score, classification_report
parameters = {
'C': np.linspace(1, 10, 10)
}
lr = LogisticRegression()
clf = GridSearchCV(lr, parameters, cv=5, verbose=5, n_jobs=3)
clf.fit(X_res, y_res.ravel())
clf.score(X_test,y_test)
from sklearn.ensemble import AdaBoostClassifier
ada = AdaBoostClassifier(n_estimators=100)
ada = ada.fit(X_res,y_res)
ada_pred = ada.predict(X_test)
ada.score(X_test,y_test)
ada_pred = ada.predict(test)
ada_predictions1 = pd.DataFrame(ada_pred,columns=['bank_account'])
ada_predictions1.insert(0,'unique_id',test1['uniqueid'])
ada_predictions1["unique_id"]=ada_predictions1["unique_id"]+' x '+test1['country']
ada_predictions1.to_csv("Mysubmission4.csv",index=False)
from sklearn.metrics import roc_curve
from sklearn.metrics import roc_auc_score
from matplotlib import pyplot
plt.figure(figsize=(15,10))
probs = logreg.predict_proba(X_test)
probs1 = ada.predict_proba(X_test)
probs2 = rfc.predict_proba(X_test)
probs3 = dtc.predict_proba(X_test)
probs = probs[:, 1]
probs1 = probs1[:, 1]
probs2 = probs2[:, 1]
probs3 = probs3[:,1]
auc = roc_auc_score(y_test, probs)
auc1 = roc_auc_score(y_test, probs1)
auc2 = roc_auc_score(y_test, probs2)
auc3 = roc_auc_score(y_test,probs3)
print('AUC of LR: %.3f' % auc)
print('AUC of ADA: %.3f' % auc1)
print('AUC of RFC: %.3f' % auc2)
print('AUC of DTC: %.3f' % auc3)
fpr, tpr, thresholds = roc_curve(y_test, probs)
fpr1, tpr1, thresholds1 = roc_curve(y_test, probs1)
fpr2, tpr2, thresholds2 = roc_curve(y_test, probs2)
fpr3, tpr3, thresholds3 = roc_curve(y_test, probs3)
pyplot.plot([0, 1], [0, 1], linestyle='--')
pyplot.plot(fpr, tpr, marker='.')
pyplot.plot(fpr1, tpr1, marker='.')
pyplot.plot(fpr2, tpr2, marker='.')
pyplot.plot(fpr3, tpr3, marker='.')
pyplot.show()
```
# Keras
```
import tensorflow as tf
from tensorflow.keras import layers
model = tf.keras.models.Sequential([
tf.keras.layers.Flatten(input_shape=(38,)),
tf.keras.layers.Dense(128, activation="relu"),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(32, activation="relu"),
tf.keras.layers.Dropout(0.1),
tf.keras.layers.Dense(1, activation="sigmoid")
])
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train)
X_test_scaled = scaler.transform(X_test)
model.compile(optimizer=tf.keras.optimizers.Adam(lr=0.00005),
loss='binary_crossentropy',
metrics=['accuracy'])
history = model.fit(X_train_scaled,
y_train,
epochs=500,
batch_size=4096,
validation_data=(X_test_scaled, y_test),
verbose=1)
model.evaluate(X_test_scaled, y_test)
test_scaled = scaler.transform(test_cleaned2.values)
tf_predictions = model.predict(test_scaled)
tf_predictions = pd.DataFrame(tf_predictions,columns=['bank_account'])
tf_predictions.insert(0,'unique_id',test1['uniqueid'])
tf_predictions["unique_id"]=tf_predictions["unique_id"]+' x '+test1['country']
tf_predictions.to_csv("MysubmissionTF.csv",index=False)
from keras.models import Sequential
from keras.layers import Dense, Activation
import numpy as np
model = Sequential()
model.add(Dense(input_dim=15, output_dim=12))
model.add(Dense(input_dim=12, output_dim=12))
model.add(Dense(input_dim=12, output_dim=2))
model.add(Activation("softmax"))
model.compile(loss='sparse_categorical_crossentropy', optimizer='sgd', metrics=['accuracy'])
scaler = StandardScaler()
model.fit(scaler.fit_transform(X_train.values), y_train,epochs=100)
y_prediction = model.predict_classes(scaler.transform(X_test.values))
print ("\n\naccuracy" , np.sum(y_prediction == y_test) / float(len(y_test)))
from sklearn import model_selection
from sklearn.linear_model import LogisticRegression
from sklearn.neighbors import KNeighborsClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.ensemble import RandomForestClassifier
from mlxtend.classifier import StackingClassifier
from sklearn.ensemble import AdaBoostClassifier
import numpy as np
import warnings
warnings.simplefilter('ignore')
clf1 = KNeighborsClassifier(n_neighbors=17)
from sklearn.neural_network import MLPClassifier
mlp = MLPClassifier()
from sklearn.ensemble import ExtraTreesClassifier
etc = ExtraTreesClassifier(n_estimators=100)
lr = LogisticRegression()
sclf = StackingClassifier(classifiers=[mlp,lr,etc , clf1],
meta_classifier=lr)
print('3-fold cross validation:\n')
for clf, label in zip([mlp, lr,etc,clf1, sclf],
['mlp',
'lr','etc','knn',
'StackingClassifier']):
scores = model_selection.cross_val_score(clf, X, y,
cv=3, scoring='accuracy')
print("Accuracy: %0.2f (+/- %0.2f) [%s]"
% (scores.mean(), scores.std(), label))
def Stacking(model,train,y,test,n_fold):
folds=StratifiedKFold(n_splits=n_fold,random_state=1)
test_pred=np.empty((test.shape[0],1),float)
train_pred=np.empty((0,1),float)
for train_indices,val_indices in folds.split(train,y.values):
x_train,x_val=train.iloc[train_indices],train.iloc[val_indices]
y_train,y_val=y.iloc[train_indices],y.iloc[val_indices]
model.fit(X=x_train,y=y_train)
train_pred=np.append(train_pred,model.predict(x_val))
test_pred=np.append(test_pred,model.predict(test))
return test_pred.reshape(-1,1),train_pred
from sklearn.model_selection import StratifiedKFold
from sklearn import tree
model1 = tree.DecisionTreeClassifier(random_state=1)
test_pred1 ,train_pred1=Stacking(model=model1,n_fold=10, train=X_train,test=X_test,y=y_train)
train_pred1=pd.DataFrame(train_pred1)
test_pred1=pd.DataFrame(test_pred1)
model2 = KNeighborsClassifier()
test_pred2 ,train_pred2=Stacking(model=model2,n_fold=10,train=X_train,test=X_test,y=y_train)
train_pred2=pd.DataFrame(train_pred2)
test_pred2=pd.DataFrame(test_pred2)
df = pd.concat([train_pred1, train_pred2], axis=1)
df_test = pd.concat([test_pred1, test_pred2], axis=1)
model = LogisticRegression(random_state=1)
model.fit(df,y_train)
#model.score(df_test, y_test)
print(df_test.shape)
print(y_test.shape)
```
# Stacking that actually works
```
from sklearn.ensemble import (RandomForestClassifier, AdaBoostClassifier,
GradientBoostingClassifier, ExtraTreesClassifier)
from sklearn.svm import SVC
from sklearn.model_selection import KFold
# Some useful parameters which will come in handy later on
ntrain = train_cleaned2.shape[0]
ntest = test_cleaned2.shape[0]
SEED = 0 # for reproducibility
NFOLDS = 5 # set folds for out-of-fold prediction
kf = KFold( n_splits= NFOLDS, random_state=SEED)
# Class to extend the Sklearn classifier
class SklearnHelper(object):
def __init__(self, clf, seed=0, params=None):
params['random_state'] = seed
self.clf = clf(**params)
def train(self, x_train, y_train):
self.clf.fit(x_train, y_train)
def predict(self, x):
return self.clf.predict(x)
def fit(self,x,y):
return self.clf.fit(x,y)
def feature_importances(self,x,y):
print(self.clf.fit(x,y).feature_importances_)
from xgboost.sklearn import XGBRegressor
from xgboost.sklearn import XGBClassifier
from catboost import CatBoostRegressor
from catboost import CatBoostClassifier
from lightgbm import LGBMRegressor
from lightgbm import LGBMClassifier
def get_oof(clf, x_train, y_train, x_test):
oof_train = np.zeros((ntrain,))
oof_test = np.zeros((ntest,))
oof_test_skf = np.empty((NFOLDS, ntest))
for i, (train_index, test_index) in enumerate(kf.split(train_cleaned2)):
x_tr = x_train[train_index]
y_tr = y_train[train_index]
x_te = x_train[test_index]
clf.train(x_tr, y_tr)
oof_train[test_index] = clf.predict(x_te)
oof_test_skf[i, :] = clf.predict(x_test)
oof_test[:] = oof_test_skf.mean(axis=0)
return oof_train.reshape(-1, 1), oof_test.reshape(-1, 1)
# Put in our parameters for said classifiers
# Random Forest parameters
rf_params = {
'n_jobs': -1,
'n_estimators': 500,
'warm_start': True,
#'max_features': 0.2,
'max_depth': 6,
'min_samples_leaf': 2,
'max_features' : 'sqrt',
'verbose': 0
}
# Extra Trees Parameters
et_params = {
'n_jobs': -1,
'n_estimators':500,
#'max_features': 0.5,
'max_depth': 8,
'min_samples_leaf': 2,
'verbose': 0
}
# AdaBoost parameters
ada_params = {
'n_estimators': 500,
'learning_rate' : 0.75
}
# Gradient Boosting parameters
gb_params = {
'n_estimators': 500,
#'max_features': 0.2,
'max_depth': 5,
'min_samples_leaf': 2,
'verbose': 0
}
# Support Vector Classifier parameters
svc_params = {
'kernel' : 'linear',
'C' : 0.025
}
xgb_params = {
'seed': 0,
'colsample_bytree': 0.7,
'silent': 1,
'subsample': 0.7,
'learning_rate': 0.075,
'objective': 'binary:logistic',
'max_depth': 4,
'num_parallel_tree': 1,
'min_child_weight': 1,
'nrounds': 200
}
catboost_params = {
'iterations': 200,
'learning_rate': 0.5,
'depth': 3,
'l2_leaf_reg': 40,
'bootstrap_type': 'Bernoulli',
'subsample': 0.7,
'scale_pos_weight': 5,
'eval_metric': 'AUC',
'od_type': 'Iter',
'allow_writing_files': False
}
lightgbm_params = {
'n_estimators':200,
'learning_rate':0.1,
'num_leaves':123,
'colsample_bytree':0.8,
'subsample':0.9,
'max_depth':15,
'reg_alpha':0.1,
'reg_lambda':0.1,
'min_split_gain':0.01,
'min_child_weight':2
}
# Create 5 objects that represent our 4 models
rf = SklearnHelper(clf=RandomForestClassifier, seed=SEED, params=rf_params)
et = SklearnHelper(clf=ExtraTreesClassifier, seed=SEED, params=et_params)
ada = SklearnHelper(clf=AdaBoostClassifier, seed=SEED, params=ada_params)
gb = SklearnHelper(clf=GradientBoostingClassifier, seed=SEED, params=gb_params)
svc = SklearnHelper(clf=SVC, seed=SEED, params=svc_params)
catboost = SklearnHelper(clf=CatBoostClassifier, seed=SEED, params=catboost_params)
xgb = SklearnHelper(clf=XGBClassifier, seed=SEED, params=xgb_params)
lgbm = SklearnHelper(clf=LGBMClassifier, seed=SEED, params=lightgbm_params)
y_train = train_cleaned2['bank_account'].ravel()
train_cleaned2 = train_cleaned2.drop(['bank_account'], axis=1)
x_train = train_cleaned2.values # Creates an array of the train data
x_test = test_cleaned2.values # Creats an array of the test data
# Create our OOF train and test predictions. These base results will be used as new features
et_oof_train, et_oof_test = get_oof(et, x_train, y_train, x_test) # Extra Trees
rf_oof_train, rf_oof_test = get_oof(rf,x_train, y_train, x_test) # Random Forest
ada_oof_train, ada_oof_test = get_oof(ada, x_train, y_train, x_test) # AdaBoost
gb_oof_train, gb_oof_test = get_oof(gb,x_train, y_train, x_test) # Gradient Boost
svc_oof_train, svc_oof_test = get_oof(svc,x_train, y_train, x_test) # Support Vector Classifier
print("Training is complete")
# Create our OOF train and test predictions. These base results will be used as new features
lgbm_oof_train, lgbm_oof_test = get_oof(lgbm, x_train, y_train, x_test) # Extra Trees
xgb_oof_train, xgb_oof_test = get_oof(xgb,x_train, y_train, x_test) # Random Forest
cat_oof_train, cat_oof_test = get_oof(catboost, x_train, y_train, x_test) # AdaBoost
print("Training is complete")
x_train.shape
rf_feature = rf.feature_importances(x_train,y_train)
et_feature = et.feature_importances(x_train, y_train)
ada_feature = ada.feature_importances(x_train, y_train)
gb_feature = gb.feature_importances(x_train,y_train)
rf_features = [0.02205724 ,0.03262432, 0.1064319 , 0.01796495 ,0.02636287, 0.03023319
,0.01720656 ,0.42861682 ,0.12952454 ,0.01675376 ,0.06027649, 0.0216796,
0.01794121 ,0.02149129, 0.02180122 ,0.02557075, 0.00346328]
print(len(rf_features))
et_features = [0.02929168, 0.07039986 ,0.13213236, 0.008584 , 0.01627604, 0.01950861,
0.01259213 ,0.30534888, 0.07075466, 0.02989719 ,0.12130776 ,0.02191149,
0.02456045 ,0.07707529, 0.01941821 ,0.03262196 ,0.00831943]
print(len(et_features))
ada_features =[0.004 ,0.008, 0.01 , 0.076 ,0.024, 0.1 , 0.016, 0.418, 0.306 ,0.002, 0.002, 0.
,0.002 ,0.002 ,0.002 ,0.006 ,0.022]
print(len(ada_features))
gb_features = [0.01230186 ,0.04541501 ,0.05320673 ,0.05736801, 0.0601205 , 0.03931794
, 0.02873651, 0.38957657 ,0.15897751 ,0.01012928 ,0.02283663, 0.0141792,
0.0138869 ,0.04215608, 0.01158523 ,0.03136426 ,0.00884177]
print(len(gb_features))
cols = train_cleaned.columns.values
# Create a dataframe with features
feature_dataframe = pd.DataFrame( {'features': cols,
'Random Forest feature importances': rf_features,
'Extra Trees feature importances': et_features,
'AdaBoost feature importances': ada_features,
'Gradient Boost feature importances': gb_features
})
feature_dataframe['mean'] = feature_dataframe.mean(axis= 1)
feature_dataframe
y = feature_dataframe['mean'].values
x = feature_dataframe['features'].values
import plotly.graph_objs as go
data = [go.Bar(
x= x,
y= y,
width = 0.5,
marker=dict(
color = feature_dataframe['mean'].values,
colorscale='Portland',
showscale=True,
reversescale = False
),
opacity=0.6
)]
layout= go.Layout(
autosize= True,
title= 'Barplots of Mean Feature Importance',
hovermode= 'closest',
# xaxis= dict(
# title= 'Pop',
# ticklen= 5,
# zeroline= False,
# gridwidth= 2,
# ),
yaxis=dict(
title= 'Feature Importance',
ticklen= 5,
gridwidth= 2
),
showlegend= False
)
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='bar-direct-labels')
base_predictions_train = pd.DataFrame( {'RandomForest': rf_oof_train.ravel(),
'ExtraTrees': et_oof_train.ravel(),
'AdaBoost': ada_oof_train.ravel(),
'GradientBoost': gb_oof_train.ravel()
})
base_predictions_train.head()
data = [
go.Heatmap(
z= base_predictions_train.astype(float).corr().values ,
x=base_predictions_train.columns.values,
y= base_predictions_train.columns.values,
colorscale='Viridis',
showscale=True,
reversescale = True
)
]
py.iplot(data, filename='labelled-heatmap')
x_train = np.concatenate(( xgb_oof_train, lgbm_oof_train, cat_oof_train), axis=1)
x_test = np.concatenate(( xgb_oof_test, lgbm_oof_test, cat_oof_test), axis=1)
gbm = XGBClassifier(
#learning_rate = 0.02,
n_estimators= 2000,
max_depth= 4,
min_child_weight= 2,
#gamma=1,
gamma=0.9,
subsample=0.8,
colsample_bytree=0.8,
objective= 'binary:logistic',
nthread= -1,
scale_pos_weight=1).fit(x_train, y_train)
predictions = gbm.predict(x_test)
predictions
stacking_model_pred = pd.DataFrame(predictions,columns=['bank_account'])
stacking_model_pred.insert(0,'unique_id',test1['uniqueid'])
stacking_model_pred["unique_id"]=stacking_model_pred["unique_id"]+' x '+test1['country']
stacking_model_pred.to_csv("MysubmissionX.csv",index=False)
# add new feature educated : yes or no
# add new feature has income : yes or no
# change age into categories
train = pd.read_csv("Train_v2.csv")
train['Has_Education'] = train["education_level"].apply(
lambda x: 1 if x=='Primary education' or x=='Secondary education' or x=='Tertiary education' else 0)
train.head()
train['job_type'].value_counts()
train_cleaned = cleaning_marital_status(train)
train_cleaned = preprocess_data(train_cleaned)
train_cleaned.head()
dic1 = {"Yes" : 1,"No" : 0}
train_cleaned['bank_account']=train_cleaned['bank_account'].map(dic1)
train_cleaned = feature_engineering(train_cleaned)
```
| github_jupyter |
STAT 453: Deep Learning (Spring 2020)
Instructor: Sebastian Raschka (sraschka@wisc.edu)
Course website: http://pages.stat.wisc.edu/~sraschka/teaching/stat453-ss2020/
GitHub repository: https://github.com/rasbt/stat453-deep-learning-ss20
```
%load_ext watermark
%watermark -a 'Sebastian Raschka' -v -p torch
```
- Runs on CPU or GPU (if available)
# Generative Adversarial Networks (GAN)AN)
A GAN with mode collapse.
## Imports
```
import time
import numpy as np
import torch
import torch.nn.functional as F
from torchvision import datasets
from torchvision import transforms
import torch.nn as nn
from torch.utils.data import DataLoader
if torch.cuda.is_available():
torch.backends.cudnn.deterministic = True
```
## Settings and Dataset
```
##########################
### SETTINGS
##########################
# Device
device = torch.device("cuda:1" if torch.cuda.is_available() else "cpu")
# Hyperparameters
random_seed = 0
generator_learning_rate = 0.001
discriminator_learning_rate = 0.001
num_epochs = 100
BATCH_SIZE = 128
LATENT_DIM = 100
IMG_SHAPE = (1, 28, 28)
IMG_SIZE = 1
for x in IMG_SHAPE:
IMG_SIZE *= x
##########################
### MNIST DATASET
##########################
# Note transforms.ToTensor() scales input images
# to 0-1 range
train_dataset = datasets.MNIST(root='data',
train=True,
transform=transforms.ToTensor(),
download=True)
test_dataset = datasets.MNIST(root='data',
train=False,
transform=transforms.ToTensor())
train_loader = DataLoader(dataset=train_dataset,
batch_size=BATCH_SIZE,
num_workers=4,
shuffle=True)
test_loader = DataLoader(dataset=test_dataset,
batch_size=BATCH_SIZE,
num_workers=4,
shuffle=False)
# Checking the dataset
for images, labels in train_loader:
print('Image batch dimensions:', images.shape)
print('Image label dimensions:', labels.shape)
break
```
## Model
```
##########################
### MODEL
##########################
class Flatten(nn.Module):
def forward(self, input):
return input.view(input.size(0), -1)
class GAN(torch.nn.Module):
def __init__(self):
super(GAN, self).__init__()
self.generator = nn.Sequential(
nn.Linear(LATENT_DIM, 128),
nn.LeakyReLU(inplace=True),
nn.Dropout(p=0.5),
nn.Linear(128, IMG_SIZE),
nn.Tanh()
)
self.discriminator = nn.Sequential(
nn.Conv2d(in_channels=1, out_channels=8, padding=1, kernel_size=(3, 3)),
nn.LeakyReLU(inplace=True),
nn.Conv2d(in_channels=8, out_channels=8, padding=1, stride=2, kernel_size=(3, 3)),
nn.LeakyReLU(inplace=True),
nn.Conv2d(in_channels=8, out_channels=16, padding=1, kernel_size=(3, 3)),
nn.LeakyReLU(inplace=True),
nn.Conv2d(in_channels=16, out_channels=16, padding=1, stride=2, kernel_size=(3, 3)),
nn.LeakyReLU(inplace=True),
nn.Conv2d(in_channels=16, out_channels=32, padding=1, kernel_size=(3, 3)),
nn.LeakyReLU(inplace=True),
nn.Conv2d(in_channels=32, out_channels=32, padding=1, stride=2, kernel_size=(3, 3)),
nn.LeakyReLU(inplace=True),
nn.AdaptiveAvgPool2d(1),
Flatten(),
nn.Linear(32, 16),
nn.LeakyReLU(inplace=True),
nn.Linear(16, 1),
nn.Sigmoid()
)
def generator_forward(self, z):
img = self.generator(z)
return img
def discriminator_forward(self, img):
pred = model.discriminator(img)
return pred.view(-1)
torch.manual_seed(random_seed)
#del model
model = GAN()
model = model.to(device)
print(model)
################################
## FOR DEBUGGING
"""
outputs= []
def hook(module, input, output):
outputs.append(output)
for i, layer in enumerate(model.discriminator):
if isinstance(layer, torch.nn.modules.conv.Conv2d):
model.discriminator[i].register_forward_hook(hook)
"""
optim_gener = torch.optim.Adam(model.generator.parameters(), lr=generator_learning_rate)
optim_discr = torch.optim.Adam(model.discriminator.parameters(), lr=discriminator_learning_rate)
```
## Training
```
start_time = time.time()
discr_costs = []
gener_costs = []
for epoch in range(num_epochs):
model = model.train()
for batch_idx, (features, targets) in enumerate(train_loader):
# Normalize images to [-1, 1] range
features = (features - 0.5)*2.
features = features.view(-1, IMG_SIZE).to(device)
targets = targets.to(device)
valid = torch.ones(targets.size(0)).float().to(device)
fake = torch.zeros(targets.size(0)).float().to(device)
### FORWARD AND BACK PROP
# --------------------------
# Train Generator
# --------------------------
# Make new images
z = torch.zeros((targets.size(0), LATENT_DIM)).uniform_(-1.0, 1.0).to(device)
generated_features = model.generator_forward(z)
# Loss for fooling the discriminator
discr_pred = model.discriminator_forward(generated_features.view(targets.size(0), 1, 28, 28))
gener_loss = F.binary_cross_entropy(discr_pred, valid)
optim_gener.zero_grad()
gener_loss.backward()
optim_gener.step()
# --------------------------
# Train Discriminator
# --------------------------
discr_pred_real = model.discriminator_forward(features.view(targets.size(0), 1, 28, 28))
real_loss = F.binary_cross_entropy(discr_pred_real, valid)
discr_pred_fake = model.discriminator_forward(generated_features.view(targets.size(0), 1, 28, 28).detach())
fake_loss = F.binary_cross_entropy(discr_pred_fake, fake)
discr_loss = 0.5*(real_loss + fake_loss)
optim_discr.zero_grad()
discr_loss.backward()
optim_discr.step()
discr_costs.append(discr_loss)
gener_costs.append(gener_loss)
### LOGGING
if not batch_idx % 100:
print ('Epoch: %03d/%03d | Batch %03d/%03d | Gen/Dis Loss: %.4f/%.4f'
%(epoch+1, num_epochs, batch_idx,
len(train_loader), gener_loss, discr_loss))
print('Time elapsed: %.2f min' % ((time.time() - start_time)/60))
print('Total Training Time: %.2f min' % ((time.time() - start_time)/60))
### For Debugging
#for i in outputs:
# print(i.size())
```
## Evaluation
```
%matplotlib inline
import matplotlib.pyplot as plt
plt.plot(range(len(gener_costs)), gener_costs, label='generator loss')
plt.plot(range(len(discr_costs)), discr_costs, label='discriminator loss')
plt.legend()
plt.show()
##########################
### VISUALIZATION
##########################
model.eval()
# Make new images
z = torch.zeros((5, LATENT_DIM)).uniform_(-1.0, 1.0).to(device)
generated_features = model.generator_forward(z)
imgs = generated_features.view(-1, 28, 28)
fig, axes = plt.subplots(nrows=1, ncols=5, figsize=(20, 2.5))
for i, ax in enumerate(axes):
axes[i].imshow(imgs[i].to(torch.device('cpu')).detach(), cmap='binary')
##########################
### VISUALIZATION
##########################
model.eval()
# Make new images
z = torch.zeros((5, LATENT_DIM)).uniform_(-1.0, 1.0).to(device)
generated_features = model.generator_forward(z)
imgs = generated_features.view(-1, 28, 28)
fig, axes = plt.subplots(nrows=1, ncols=5, figsize=(20, 2.5))
for i, ax in enumerate(axes):
axes[i].imshow(imgs[i].to(torch.device('cpu')).detach(), cmap='binary')
##########################
### VISUALIZATION
##########################
model.eval()
# Make new images
z = torch.zeros((10, LATENT_DIM)).uniform_(-1.0, 1.0).to(device)
generated_features = model.generator_forward(z)
imgs = generated_features.view(-1, 28, 28)
fig, axes = plt.subplots(nrows=1, ncols=10, figsize=(20, 2.5))
for i, ax in enumerate(axes):
axes[i].imshow(imgs[i].to(torch.device('cpu')).detach(), cmap='binary')
```
| github_jupyter |
# An introduction to functions
Functions are the building blocks of most programs. Without functions, we would often have to repeat ourselves.
They have the following pattern:
```python
def my_function(x):
"""Docstring."""
y = operations on x
return y
```
The general idea is that you can organize your code using functions. Instead of just being a long series of instructions, your code can use re-usable blocks, jumping around from block to block.
When you 'call' a function, by using its name and passing it any arguments (input) it needs, it returns its output to exactly the place it was called from.
For example, instead of writing:
```python
x = (x_in_f - 32) * 5/9
y = (y_in_f - 32) * 5/9
z = (z_in_f - 32) * 5/9
```
It is more readable and easier to maintain if we do this:
```python
def f_to_c(temp):
return (temp - 32) * 5/9
x = f_to_c(x_in_f)
y = f_to_c(y_in_f)
z = f_to_c(z_in_f)
```
Better yet:
```python
temps = [x_in_f, y_in_f, z_in_f]
x, y, z = [f_to_c(t) for t in temps]
```
## A simple function
We'll start by defining a very simple function, implementing the acoustic impedance equation:
$$ Z = \rho V_\mathrm{P} $$
```
def impedance(rho, vp):
"""Compute acoustic impedance from density and velocity."""
z = rho * vp
return z
impedance(2300, 2500)
```
Note that we do not have access to the variables inside the function.
```
z
```
Similarly, if the variable `z` already exists outside the function, it is unaffected by the function:
```
z = 'Not even a number.'
impedance(rho=2400, vp=2100)
print(z)
```
<div class="alert alert-success">
<h3>Exercise</h3>
Add your function to a new text file called `utils.py`, and save it in the current directory.
</div>
```
# The current directory is whatever shows up when you run this cell.
%pwd
```
## Default values
We can set default values for some or all of the things we pass to the function — its 'arguments'.
```
def quad(x, a=1, b=1, c=0):
"""
Returns the quadratic function of x,
a.x^2 + b.x + c
where by default
a = b = 1 and c = 0.
"""
return a*x**2 + b*x + c
quad(10, c=4)
```
Now we can call the function with only the required argument, `x`, and let the others fall back on their default values.
```
quad(10)
```
We can change one or more of these arguments when we call the function:
```
quad(10, a=3, c=1) # b is 1 by default.
```
The argument `x` in this example is a 'positional argument', while `a`, `b`, and `c` are called 'keyword arguments'. They are sometimes also called 'optional arguments' because they can be omitted from the function call.
<div class="alert alert-success">
<h3>Exercise</h3>
Go to the [`Practice_functions.ipynb`](Practice_functions.ipynb) notebook.
</div>
----
# Intro to Python students stop here
----
## `args` and `kwargs`
This is a more advanced concept, but it's good to know about. We can write functions that take an arbitrary number of arguments, as well as arguments you give specific names to:
```
def add(*args):
print(args)
return sum(args)
add(2, 3, 4, 5, 6, 7)
```
We can mix this with an arbitrary number of positional and keyword arguments:
```
def foo(x, y, *args):
"""
Print these things.
"""
print(x, y)
print(args)
return
foo(2, 'this', 'that', 45)
```
The unnamed `args` are stored in a `tuple`, and this is what was printed out by `print(args)`.
You can pass keyword arguments in the same way, with a slightly different syntax:
```
def bar(x, y=1, **kwargs):
print(x, y)
print(kwargs)
return
```
This time the unspecified `kwargs` are stored in a `dict`:
```
bar(2, 'this', that='that', other=45)
```
You can create a `dict` of containing the keywords and values outside the function and pass it in. The `**kwargs` syntax unpacks the element of the dictionary for use inside the function:
```
func_params = dict(param1 = 'alpha', param2 = 'beta', param3 = 99.0)
bar(2, **func_params)
```
## Passing functions, and unnamed functions
Functions have names, just like other objects in Python. We can pass them around, just like other objects. Sometimes we might want to use a function as input to another function.
For example, the `sorted` function sorts sequences:
```
x = list('I Love Geology')
sorted(x)
```
Notice how Python sorts the upper case characters before the lower case ones. Supposing we'd like to ignore the case of the letters when sorting? Then we can define a function that returns the thing we'd like to sort on:
```
def ignore_case(x):
return x.lower()
sorted(x, key=ignore_case)
```
Sometimes in a situation like this you don't actually want to go to the trouble of defining a function, especially such a short one. Python has a 'function-lite' option — the `lambda` — which we define but don't name. You can think of it as a one-shot, throwaway, or temporary function:
```
sorted(x, key=lambda x: x.lower())
```
The syntax is a little weird though, and some people think that using a lot of `lambda` functions makes for hard-to-read code.
<div class="alert alert-success">
<h3>Exercise</h3>
I have a list of rocks, can you sort them by the last letter in each name?
Here's the desired result: `['sandstone', 'chalk', 'basalt']`
</div>
```
# Add your code here. HINT: define a function.
rocks = ['basalt', 'chalk', 'sandstone']
# Add your code here. HINT: define a function.
sorted(y, key=last_char)
```
<div class="alert alert-success">
<h3>Exercise</h3>
Can you sort this list of files in numerical order?
Here's the desired result: `['file01.txt', 'file2.txt', 'file03.txt', 'file11.txt', 'file12.txt']`
</div>
```
files = ['file2.txt', 'file03.txt', 'file11.txt', 'file12.txt', 'file01.txt']
# Your code goes here
```
## A note about documentation
There are two main kinds of documentation:
- Docs that you write to help people understand how to use your tool.
- Comments in the code to help other coders understand how the tool works.
In general, you should try to write code that does not need copious documentation. Using descriptive variable and function names, keeping functions simple, and writing function docstrings all help.
Docstrings are the least that is required of you — these will become available to users of your code, as shown in the examples that follow. First, here's how to write a docstring:
```
def ignore_case(x):
"""
This is a docstring. It's special.
Args:
x (str). A string to send to lowercase.
Returns:
str. The string in lowercase.
"""
# This is just a normal comment.
return x.lower() # So is this.
```
Now let's look at how another person might read this information. The easiest way to get it is to call `help()` on the function:
```
help(ignore_case)
```
You could also inspect the docstring directly; Python stores it on an attribute of the function called `__doc__`:
```
print(ignore_case.__doc__)
```
In Jupyter Notebooks and in the IPython interpreter you can also type the name of the function with a `?`
```
ignore_case?
```
## A note about testing
Testing is an important step in writing correct code. Indeed, programmers have a saying:
> Untested code is broken code.
In our experience, the process of writing tests often reveals bugs, and almost always results in better code.
A quick and easy way to get started with testing is the built0in `doctest` module. Simply add one or more examples yo the docstring of the function:
```
def ignore_case(x):
"""
This is a docstring. It's special.
Args:
x (str). A string to send to lowercase.
Returns:
str. The string in lowercase.
Example:
>>> ignore_case('Geology.')
'geology.'
"""
# This is just a normal comment.
return x.lower() # So is this.
```
Then we can test the function like this:
```
import doctest
doctest.testmod()
```
Our test passed! `doctest` looked for the line that looks like interactive Python input (with the `>>>` at the start), ran it, and compared its output to the line in my example. Since the lines matched, the test passed. You can add as many tests as you like to the docstring.
See [the `docstring` documentation](https://docs.python.org/3.6/library/doctest.html) for more details, or check out the notebook [Introduction to testing](Intro_to_testing.ipynb) to go deeper.
## A note about type hints
New in Python 3. Essentially a type of documentation. [Read about them.](https://docs.python.org/3/library/typing.html) [Read PEP484](https://www.python.org/dev/peps/pep-0484/).
You can check the internal consistency of types using [mypy](http://mypy-lang.org/index.html).
Python is **strongly typed** — you cannot add an `int` to a `str`. For example, `2 + '3'` throws a `TypeError`, whereas in JavaScript, which is weakly typed, it returns `'23'`.
But Python is **dynamically typed**, so I can do `x = 5` and then, later, `x = 'Hello'` — the type of `x` is dynamic, and depends only on the data I point it to. Similarly, I can pass ints, floats or strings into a function that multiplies things:
```
def double(n):
return 2 * n
double('this')
```
As you might imagine, sometimes this kind of flexibility can be the cause of bugs.
The basic idea of type hints is to bridge the gap between dynamic typing (Python's usual mode, so to speak), and static typing (a popular feature of some other languages, such as Java or C).
You can annotate a variable assignment with the expected type of the variable, for example:
```
n: float = 3.14159
```
There's a similar signature for annotating functions, with some special syntax for annotating the return variable too:
```
def double(n: float) -> float:
return 2 * n
double(2.1)
```
These are just annotations, however, there is no actual type checking. You can still do whatever you want.
```
double('that')
```
You can, however, check the internal consistency of types using [mypy](http://mypy-lang.org/index.html).
The `typing` module helps make hybrid types, new types, etc.
```
from typing import List
def scale(scalar: float, vector: List[float]) -> List[float]:
return [scalar * num for num in vector]
new_vector = scale(2.0, [1.0, -4.2, 5.4])
new_vector
```
None of this changes the actual type of the variables:
```
type(new_vector)
```
<hr />
<div>
<img src="https://avatars1.githubusercontent.com/u/1692321?s=50"><p style="text-align:center">© Agile Geoscience 2018</p>
</div>
| github_jupyter |
```
# Erasmus+ ICCT project (2018-1-SI01-KA203-047081)
# Toggle cell visibility
from IPython.display import HTML
tag = HTML('''<script>
code_show=true;
function code_toggle() {
if (code_show){
$('div.input').hide()
} else {
$('div.input').show()
}
code_show = !code_show
}
$( document ).ready(code_toggle);
</script>
Toggle cell visibility <a href="javascript:code_toggle()">here</a>.''')
display(tag)
# Hide the code completely
# from IPython.display import HTML
# tag = HTML('''<style>
# div.input {
# display:none;
# }
# </style>''')
# display(tag)
```
## Prehod iz sistema diferencialnih enačb v model prostora stanj
### Modeliranje sistema masa-vzmet-dušilka
Ta interaktivni primer omogoča opazovanje vpliva parametrov sistema na naravni (nevsiljeni oz. lastni) in vsiljeni odziv sistema masa-vzmet-dušilka; sistem je podrobneje obravnavan v spremljajoči knjigi.
<img src="Images\mass-spring-damper.PNG" alt="drawing" width="300">
Obravnavani sistem lahko popišemo z naslednjo enačbo:
$$m\ddot{x}=-kx-c\dot{x}+F(t),$$
kjer $x$ označuje lego mase vzdolž njene prostostne stopnje, $m$ njeno dejansko maso, $k$ koeficient vzmeti (iz tega sledi, da je $kx$ sila, s katero vzmet deluje na maso), $c$ koeficient dušenja (sledi, da $c\dot{x}$ sila, s katero dušilka deluje na maso, in $F(t)$ zunanjo silo, s katero delujemo na maso (ta sila predstavlja vhod sistema).
Z določitvijo vektorja stanj $\textbf{x}=[x_1, x_2]^T$, kjer sta $x_1=x$ in $x_2=\dot{x}$, in vhoda $u(t)=F(t)$, je možno opisati odziv sistema z naslednjima enačbama:
\begin{cases}
\dot{x_2}=-\frac{k}{m}x_1-\frac{c}{m}x_2+u(t) \\
\dot{x_1}=x_2
\end{cases}
(upoštevamo, da je $\dot{x_2}=\ddot{x}$). Enačbi nato zapišemo v matrični obliki
$$
\begin{bmatrix}
\dot{x_1} \\
\dot{x_2}
\end{bmatrix}=\underbrace{\begin{bmatrix}
0 && 1 \\
-\frac{k}{m} && -\frac{c}{m}
\end{bmatrix}}_{A}\begin{bmatrix}
x_1 \\
x_2
\end{bmatrix}+\underbrace{\begin{bmatrix}
0 \\
\frac{1}{m}
\end{bmatrix}}_{B}u.
$$
### Kako upravljati s tem interaktivnim primerom?
- Razišči različne odzive sistema na različne začetne pogoje in vhode v sistem.
- Opazuj, kako vrednost koeficienta dušenja $c$ vpliva na prisotnost oscilacij lege mase.
Poizkusi nastaviti parametre sistema tako, 1) da bosta pomik in hitrost mase oscilirali v neskončnost (da bo sistem nedušen) in 2) da bo sistem čim hitreje dosegel želeno končno stanje brez oscilacij.
Poizkusi nastaviti parametre sistema $m$, $k$ in $c$ ter izbrati vhodno funkcijo tako, da se bo amplituda oscilacij povečevala s časom. Ali je možno doseči tak odziv sistema?
```
#Preparatory Cell
%matplotlib notebook
import control
import numpy
from IPython.display import display, Markdown
import ipywidgets as widgets
import matplotlib.pyplot as plt
from matplotlib import animation
%matplotlib inline
#print a matrix latex-like
def bmatrix(a):
"""Returns a LaTeX bmatrix - by Damir Arbula (ICCT project)
:a: numpy array
:returns: LaTeX bmatrix as a string
"""
if len(a.shape) > 2:
raise ValueError('bmatrix can at most display two dimensions')
lines = str(a).replace('[', '').replace(']', '').splitlines()
rv = [r'\begin{bmatrix}']
rv += [' ' + ' & '.join(l.split()) + r'\\' for l in lines]
rv += [r'\end{bmatrix}']
return '\n'.join(rv)
# Display formatted matrix:
def vmatrix(a):
if len(a.shape) > 2:
raise ValueError('bmatrix can at most display two dimensions')
lines = str(a).replace('[', '').replace(']', '').splitlines()
rv = [r'\begin{vmatrix}']
rv += [' ' + ' & '.join(l.split()) + r'\\' for l in lines]
rv += [r'\end{vmatrix}']
return '\n'.join(rv)
#matrixWidget is a matrix looking widget built with a VBox of HBox(es) that returns a numPy array as value !
class matrixWidget(widgets.VBox):
def updateM(self,change):
for irow in range(0,self.n):
for icol in range(0,self.m):
self.M_[irow,icol] = self.children[irow].children[icol].value
#print(self.M_[irow,icol])
self.value = self.M_
def dummychangecallback(self,change):
pass
def __init__(self,n,m):
self.n = n
self.m = m
self.M_ = numpy.matrix(numpy.zeros((self.n,self.m)))
self.value = self.M_
widgets.VBox.__init__(self,
children = [
widgets.HBox(children =
[widgets.FloatText(value=0.0, layout=widgets.Layout(width='90px')) for i in range(m)]
)
for j in range(n)
])
#fill in widgets and tell interact to call updateM each time a children changes value
for irow in range(0,self.n):
for icol in range(0,self.m):
self.children[irow].children[icol].value = self.M_[irow,icol]
self.children[irow].children[icol].observe(self.updateM, names='value')
#value = Unicode('example@example.com', help="The email value.").tag(sync=True)
self.observe(self.updateM, names='value', type= 'All')
def setM(self, newM):
#disable callbacks, change values, and reenable
self.unobserve(self.updateM, names='value', type= 'All')
for irow in range(0,self.n):
for icol in range(0,self.m):
self.children[irow].children[icol].unobserve(self.updateM, names='value')
self.M_ = newM
self.value = self.M_
for irow in range(0,self.n):
for icol in range(0,self.m):
self.children[irow].children[icol].value = self.M_[irow,icol]
for irow in range(0,self.n):
for icol in range(0,self.m):
self.children[irow].children[icol].observe(self.updateM, names='value')
self.observe(self.updateM, names='value', type= 'All')
#self.children[irow].children[icol].observe(self.updateM, names='value')
#overlaod class for state space systems that DO NOT remove "useless" states (what "professor" of automatic control would do this?)
class sss(control.StateSpace):
def __init__(self,*args):
#call base class init constructor
control.StateSpace.__init__(self,*args)
#disable function below in base class
def _remove_useless_states(self):
pass
#define matrixes
C = numpy.matrix([[1,0],[0,1]])
D = numpy.matrix([[0],[0]])
X0 = matrixWidget(2,1)
m = widgets.FloatSlider(
value=5,
min=0.1,
max=10.0,
step=0.1,
description='m [kg]:',
disabled=False,
continuous_update=False,
orientation='horizontal',
readout=True,
readout_format='.1f',
)
k = widgets.FloatSlider(
value=1,
min=0,
max=10.0,
step=0.1,
description='k [N/m]:',
disabled=False,
continuous_update=False,
orientation='horizontal',
readout=True,
readout_format='.1f',
)
c = widgets.FloatSlider(
value=0.5,
min=0,
max=10.0,
step=0.1,
description='c [Ns/m]:',
disabled=False,
continuous_update=False,
orientation='horizontal',
readout=True,
readout_format='.1f',
)
u = widgets.FloatSlider(
value=1,
min=0,
max=10.0,
step=0.1,
description='',
disabled=False,
continuous_update=False,
orientation='horizontal',
readout=True,
readout_format='.1f',
)
omega = widgets.FloatSlider(
value=5,
min=0,
max=10.0,
step=0.1,
description='',
disabled=False,
continuous_update=False,
orientation='horizontal',
readout=True,
readout_format='.1f',
)
def main_callback(X0, m, k, c, u, selu, omega):#m, k, c, u, selu, DW
a = numpy.matrix([[0,1],[-k/m,-c/m]])
b = numpy.matrix([[0],[1/m]])
eig = numpy.linalg.eig(a)
sys = sss(a,b,C,D)
if min(numpy.real(abs(eig[0]))) != 0:
T = numpy.linspace(0,100/min(numpy.real(abs(eig[0]))),1000)
else:
if max(numpy.real(abs(eig[0]))) != 0:
T = numpy.linspace(0,100/max(numpy.real(abs(eig[0]))),1000)
else:
T = numpy.linspace(0,1000,1000)
if selu == 'impulzna funkcija': #selu
U = [0 for t in range(0,len(T))]
U[0] = u
y = control.forced_response(sys,T,U,X0)
if selu == 'koračna funkcija':
U = [u for t in range(0,len(T))]
y = control.forced_response(sys,T,U,X0)
if selu == 'sinusoidna funkcija':
U = u*numpy.sin(omega*T)
y = control.forced_response(sys,T,U,X0)
fig=plt.figure(num=1,figsize=[15, 4])
fig.add_subplot(121)
plt.plot(T,y[1][0])
plt.grid()
plt.xlabel('čas [s]')
plt.ylabel('pomik [m]')
fig.add_subplot(122)
plt.plot(T,y[1][1])
plt.grid()
plt.xlabel('čas [s]')
plt.ylabel('hitrost [m/s]')
#display(Markdown('The A matrix is: $%s$ and the eigenvalues are: $%s$' % (bmatrix(a),eig[0])))
#create dummy widget
DW = widgets.FloatText(layout=widgets.Layout(width='0px', height='0px'))
#create button widget
START = widgets.Button(
description='Test',
disabled=False,
button_style='', # 'success', 'info', 'warning', 'danger' or ''
tooltip='Test',
icon='check'
)
def on_start_button_clicked(b):
#This is a workaround to have intreactive_output call the callback:
# force the value of the dummy widget to change
if DW.value> 0 :
DW.value = -1
else:
DW.value = 1
pass
START.on_click(on_start_button_clicked)
#define type of ipout
SELECT = widgets.Dropdown(
options=['impulzna funkcija', 'koračna funkcija', 'sinusoidna funkcija'],
value='impulzna funkcija',
description='',
disabled=False
)
#create a graphic structure to hold all widgets
alltogether = widgets.VBox([widgets.HBox([widgets.VBox([m,
k,
c]),
widgets.HBox([widgets.VBox([widgets.Label('Izberi vhodno funkcijo:',border=3),
widgets.Label('u [N]:',border=3),
widgets.Label('omega [rad/s]:',border=3)]),
widgets.VBox([SELECT,u,omega])])]),
widgets.HBox([widgets.Label('Začetni pogoj X0:',border=3),X0])])
out = widgets.interactive_output(main_callback,{'X0':X0, 'm': m, 'k': k, 'c': c, 'u': u, 'selu': SELECT, 'omega':omega})
#out.layout.height = '300px'
display(out,alltogether)
```
| github_jupyter |
```
# Support Vector Machine
# Importing the libraries
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# Importing the datasets
datasets = pd.read_csv('Social_Network_Ads.csv')
X = datasets.iloc[:, [2,3]].values
Y = datasets.iloc[:, 4].values
datasets.head()
# Splitting the dataset into the Training set and Test set
from sklearn.model_selection import train_test_split
X_Train, X_Test, Y_Train, Y_Test = train_test_split(X, Y, test_size = 0.25, random_state = 0)
# Feature Scaling
from sklearn.preprocessing import StandardScaler
sc_X = StandardScaler()
X_Train = sc_X.fit_transform(X_Train)
X_Test = sc_X.transform(X_Test)
# Fitting the classifier into the Training set
from sklearn.svm import SVC
classifier = SVC(kernel = 'linear', random_state = 0)
classifier.fit(X_Train, Y_Train)
# Predicting the test set results
Y_Pred = classifier.predict(X_Test)
# Making the Confusion Matrix
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(Y_Test, Y_Pred)
# Visualising the Training set results
from matplotlib.colors import ListedColormap
X_Set, Y_Set = X_Train, Y_Train
X1, X2 = np.meshgrid(np.arange(start = X_Set[:, 0].min() - 1, stop = X_Set[:, 0].max() + 1, step = 0.01),
np.arange(start = X_Set[:, 1].min() - 1, stop = X_Set[:, 1].max() + 1, step = 0.01))
plt.contourf(X1, X2, classifier.predict(np.array([X1.ravel(), X2.ravel()]).T).reshape(X1.shape),
alpha = 0.75, cmap = ListedColormap(('red', 'green')))
plt.xlim(X1.min(), X1.max())
plt.ylim(X2.min(), X2.max())
for i, j in enumerate(np.unique(Y_Set)):
plt.scatter(X_Set[Y_Set == j, 0], X_Set[Y_Set == j, 1],
c = ListedColormap(('red', 'green'))(i), label = j)
plt.title('Support Vector Machine (Training set)')
plt.xlabel('Age')
plt.ylabel('Estimated Salary')
plt.legend()
plt.show()
# Visualising the Test set results
from matplotlib.colors import ListedColormap
X_Set, Y_Set = X_Test, Y_Test
X1, X2 = np.meshgrid(np.arange(start = X_Set[:, 0].min() - 1, stop = X_Set[:, 0].max() + 1, step = 0.01),
np.arange(start = X_Set[:, 1].min() - 1, stop = X_Set[:, 1].max() + 1, step = 0.01))
plt.contourf(X1, X2, classifier.predict(np.array([X1.ravel(), X2.ravel()]).T).reshape(X1.shape),
alpha = 0.75, cmap = ListedColormap(('red', 'green')))
plt.xlim(X1.min(), X1.max())
plt.ylim(X2.min(), X2.max())
for i, j in enumerate(np.unique(Y_Set)):
plt.scatter(X_Set[Y_Set == j, 0], X_Set[Y_Set == j, 1],
c = ListedColormap(('red', 'green'))(i), label = j)
plt.title('Support Vector Machine (Test set)')
plt.xlabel('Age')
plt.ylabel('Estimated Salary')
plt.legend()
plt.show()
```
| github_jupyter |
```
import mdtraj as md
import numpy as np
import matplotlib.pyplot as plot
import subprocess
import os
BCCID= ['md1/']
mydir='/net/jam-amaro-shared/bccgc4/Clustering/Gromos/clustering/'
workingdir=mydir
filedir='/net/jam-amaro-shared/bccgc4/Strided_Traj/'
j='joined_traj_md'
traj1=md.load([filedir+j+'1.nc', filedir+j+'2.nc', filedir+j+'3.nc', filedir+j+'4.nc', filedir+j+'5.nc'], top=filedir+'protein.h5')
print (traj1)
#alignment by alpha carbons
alphacarbon_indices=traj1.topology.select('name CA')
print(alphacarbon_indices)
traj1.superpose(traj1, 0, alphacarbon_indices)
#proteinv selection (trajectory.pdb)
#protein1selection=traj1.topology.select('protein')
#print(protein1selection)
#protein1=traj1.atom_slice(protein1selection)
#print(protein1)
#print(protein1.topology)
#protein1.save_pdb(workingdir+'trajectory.pdb')
traj1.save_pdb(workingdir+'trajectory.pdb')
print("YAY")
###trajectory.pdb formatting
get_ipython().system('cat '+mydir+'trajectory.pdb | grep -v CRYST1 > '+mydir+'temp.pdb')
get_ipython().system('mv -f '+mydir+'temp.pdb '+mydir+'trajectory.pdb')
#get_ipython().system('perl -pi -e "s/END/ENDMDL/g" trajectory.pdb')
ref_frame=traj1[0]
print(ref_frame)
ref_frame.save_pdb(workingdir+'ref_frame.pdb')
get_ipython().system(" cat "+mydir+"ref_frame.pdb | awk '{print $6}' | sort -n | uniq > "+mydir+"resid_ref_frame.dat")
print("Woooh")
# resid_activesite.dat formatting
f=open(mydir+'resid_ref_frame.dat','r')
lines=f.readlines()
f.close()
f=open(mydir+'resid_ref_frame.dat','w')
for line in lines:
if (line [0:1]!='\n'):
if "," not in line:
f.write(line)
f.close()
print("Duck Tales")
get_ipython().system('cat '+mydir+'resid_ref_frame.dat | awk \'{print "cat /net/jam-amaro-shared/bccgc4/Clustering/Gromos/first_frame.pdb | awk STARTif ($6==" $1 ") print $0 END" }\' | sed "s/START/\'{/g" | sed "s/END/}\'/g" | csh > '+mydir+'ref_frame_correct$| csh > '+mydir+'ref_frame_correct_residues.pdb')
get_ipython().system('cat '+mydir+'ref_frame_correct_residues.pdb | awk \'{printf $2 " "}\' > '+mydir+'ref_frame_atoms_indices.dat')
get_ipython().system('cat '+mydir+'ref_frame_correct_residues.pdb | grep " CA " | awk \'{ if ( NR%15 == 0){ {printf "%4i", $2} {printf "\\n"} } else {printf "%4i ", $2} }\' > '+mydir+'ref_frame.ndx')
get_ipython().system('cat /net/jam-amaro-shared/bccgc4/Clustering/Gromos/first_frame.pdb | grep " CA " | awk \'{ if ( NR%15 == 0){ {printf "%4i", $2} {printf "\\n"} } else {printf "%4i ", $2} }\' > '+mydir+'alpha_carbons_indices.ndx')
get_ipython().system('cat '+mydir+'alpha_carbons_indices.ndx '+mydir+'ref_frame.ndx > '+mydir+'selections.ndx')
print("Woooh")
f=open(mydir+'ref_frame.ndx','r+')
lines=f.readlines()
f.seek(0)
f.write('[ ref_frame_CA ] \n')
for line in lines:
f.write(line)
f.close()
print("David tenat's in this one")
f=open(mydir+'alpha_carbons_indices.ndx','r+')
lines=f.readlines()
f.seek(0)
f.write('[ C-alpha ] \n')
for line in lines:
f.write(line)
f.close()
print("And Lin Manuel Miranda")
get_ipython().system('cat '+mydir+'alpha_carbons_indices.ndx '+mydir+'ref_frame.ndx > '+mydir+'selections.ndx')
```
When you finish the above step, get the first frame of each trajectory and name it first_frame.pdb in the working directory.
| github_jupyter |
```
# # from google.colab import drive
# drive.mount('/content/drive')
# from google.colab import drive
# drive.mount('/content/drive')
!pwd
path = '/kaggle/working/run_'
# path=''
import torch.nn as nn
import torch.nn.functional as F
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import torch
import torchvision
import torchvision.transforms as transforms
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
from matplotlib import pyplot as plt
import copy
# from sparsemax import Sparsemax
import torch.optim as optim
# Ignore warnings
import warnings
warnings.filterwarnings("ignore")
# n_seed = 0
#k = 0
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark= False
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
trainset = torchvision.datasets.CIFAR10(root='./data', train=True, download=True, transform=transform)
testset = torchvision.datasets.CIFAR10(root='./data', train=False, download=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=10, shuffle=False)
testloader = torch.utils.data.DataLoader(testset, batch_size=10, shuffle=False)
classes = ('plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
foreground_classes = {'plane', 'car', 'bird'}
#foreground_classes = {'bird', 'cat', 'deer'}
background_classes = {'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck'}
#background_classes = {'plane', 'car', 'dog', 'frog', 'horse','ship', 'truck'}
fg1,fg2,fg3 = 0,1,2
dataiter = iter(trainloader)
background_data=[]
background_label=[]
foreground_data=[]
foreground_label=[]
batch_size=10
for i in range(5000):
images, labels = dataiter.next()
for j in range(batch_size):
if(classes[labels[j]] in background_classes):
img = images[j].tolist()
background_data.append(img)
background_label.append(labels[j])
else:
img = images[j].tolist()
foreground_data.append(img)
foreground_label.append(labels[j])
foreground_data = torch.tensor(foreground_data)
foreground_label = torch.tensor(foreground_label)
background_data = torch.tensor(background_data)
background_label = torch.tensor(background_label)
def create_mosaic_img(bg_idx,fg_idx,fg):
"""
bg_idx : list of indexes of background_data[] to be used as background images in mosaic
fg_idx : index of image to be used as foreground image from foreground data
fg : at what position/index foreground image has to be stored out of 0-8
"""
image_list=[]
j=0
for i in range(9):
if i != fg:
image_list.append(background_data[bg_idx[j]])#.type("torch.DoubleTensor"))
j+=1
else:
image_list.append(foreground_data[fg_idx])#.type("torch.DoubleTensor"))
label = foreground_label[fg_idx]- fg1 # minus fg1 because our fore ground classes are fg1,fg2,fg3 but we have to store it as 0,1,2
#image_list = np.concatenate(image_list ,axis=0)
image_list = torch.stack(image_list)
return image_list,label
desired_num = 30000
mosaic_list_of_images =[] # list of mosaic images, each mosaic image is saved as list of 9 images
fore_idx =[] # list of indexes at which foreground image is present in a mosaic image i.e from 0 to 9
mosaic_label=[] # label of mosaic image = foreground class present in that mosaic
for i in range(desired_num):
np.random.seed(i)
bg_idx = np.random.randint(0,35000,8)
fg_idx = np.random.randint(0,15000)
fg = np.random.randint(0,9)
fore_idx.append(fg)
image_list,label = create_mosaic_img(bg_idx,fg_idx,fg)
mosaic_list_of_images.append(image_list)
mosaic_label.append(label)
plt.imshow(torch.transpose(mosaic_list_of_images[0][1],dim0= 0,dim1 = 2))
class MosaicDataset(Dataset):
"""MosaicDataset dataset."""
def __init__(self, mosaic_list_of_images, mosaic_label, fore_idx):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.mosaic = mosaic_list_of_images
self.label = mosaic_label
self.fore_idx = fore_idx
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.mosaic[idx] , self.label[idx], self.fore_idx[idx]
batch = 250
msd = MosaicDataset(mosaic_list_of_images, mosaic_label , fore_idx)
train_loader = DataLoader( msd,batch_size= batch ,shuffle=True)
class Focus(nn.Module):
def __init__(self):
super(Focus, self).__init__()
self.conv1 = nn.Conv2d(in_channels=3, out_channels=32, kernel_size=3, padding=0,bias=False)
self.conv2 = nn.Conv2d(in_channels=32, out_channels=64, kernel_size=3, padding=0,bias=False)
self.conv3 = nn.Conv2d(in_channels=64, out_channels=64, kernel_size=3, padding=0,bias=False)
self.conv4 = nn.Conv2d(in_channels=64, out_channels=128, kernel_size=3, padding=0,bias=False)
self.conv5 = nn.Conv2d(in_channels=128, out_channels=256, kernel_size=3, padding=0,bias=False)
self.conv6 = nn.Conv2d(in_channels=256, out_channels=256, kernel_size=3, padding=1,bias=False)
self.pool = nn.MaxPool2d(kernel_size=2, stride=2)
self.batch_norm1 = nn.BatchNorm2d(32,track_running_stats=False)
self.batch_norm2 = nn.BatchNorm2d(64,track_running_stats=False)
self.batch_norm3 = nn.BatchNorm2d(256,track_running_stats=False)
self.dropout1 = nn.Dropout2d(p=0.05)
self.dropout2 = nn.Dropout2d(p=0.1)
self.fc1 = nn.Linear(256,64,bias=False)
self.fc2 = nn.Linear(64, 32,bias=False)
self.fc3 = nn.Linear(32, 10,bias=False)
self.fc4 = nn.Linear(10, 1,bias=False)
torch.nn.init.xavier_normal_(self.conv1.weight)
torch.nn.init.xavier_normal_(self.conv2.weight)
torch.nn.init.xavier_normal_(self.conv3.weight)
torch.nn.init.xavier_normal_(self.conv4.weight)
torch.nn.init.xavier_normal_(self.conv5.weight)
torch.nn.init.xavier_normal_(self.conv6.weight)
torch.nn.init.xavier_normal_(self.fc1.weight)
torch.nn.init.xavier_normal_(self.fc2.weight)
torch.nn.init.xavier_normal_(self.fc3.weight)
torch.nn.init.xavier_normal_(self.fc4.weight)
def forward(self,z): #y is avg image #z batch of list of 9 images
y = torch.zeros([batch,256, 3,3], dtype=torch.float64)
x = torch.zeros([batch,9],dtype=torch.float64)
ftr = torch.zeros([batch,9,256,3,3])
y = y.to("cuda")
x = x.to("cuda")
ftr = ftr.to("cuda")
for i in range(9):
out,ftrs = self.helper(z[:,i])
#print(out.shape)
x[:,i] = out
ftr[:,i] = ftrs
log_x = F.log_softmax(x,dim=1) # log_alpha
x = F.softmax(x,dim=1)
for i in range(9):
x1 = x[:,i]
y = y + torch.mul(x1[:,None,None,None],ftr[:,i])
return x, y, log_x, #alpha, log_alpha, avg_data
def helper(self, x):
#x1 = x
#x1 =x
x = self.conv1(x)
x = F.relu(self.batch_norm1(x))
x = (F.relu(self.conv2(x)))
x = self.pool(x)
x = self.conv3(x)
x = F.relu(self.batch_norm2(x))
x = (F.relu(self.conv4(x)))
x = self.pool(x)
x = self.dropout1(x)
x = self.conv5(x)
x = F.relu(self.batch_norm3(x))
x = self.conv6(x)
x1 = F.tanh(x)
x = F.relu(x)
x = self.pool(x)
x = x.view(x.size(0), -1)
x = self.dropout2(x)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.dropout2(x)
x = F.relu(self.fc3(x))
x = self.fc4(x)
x = x[:,0]
# print(x.shape)
return x,x1
class Classification(nn.Module):
def __init__(self):
super(Classification, self).__init__()
self.conv1 = nn.Conv2d(in_channels=256, out_channels=128, kernel_size=3, padding=1)
self.conv2 = nn.Conv2d(in_channels=128, out_channels=128, kernel_size=3, padding=1)
self.conv3 = nn.Conv2d(in_channels=128, out_channels=256, kernel_size=3, padding=1)
self.conv4 = nn.Conv2d(in_channels=256, out_channels=256, kernel_size=3, padding=1)
self.conv5 = nn.Conv2d(in_channels=256, out_channels=512, kernel_size=3, padding=1)
self.conv6 = nn.Conv2d(in_channels=512, out_channels=512, kernel_size=3, padding=1)
self.pool = nn.MaxPool2d(kernel_size=2, stride=2,padding=1)
self.batch_norm1 = nn.BatchNorm2d(128,track_running_stats=False)
self.batch_norm2 = nn.BatchNorm2d(256,track_running_stats=False)
self.batch_norm3 = nn.BatchNorm2d(512,track_running_stats=False)
self.dropout1 = nn.Dropout2d(p=0.05)
self.dropout2 = nn.Dropout2d(p=0.1)
self.global_average_pooling = nn.AvgPool2d(kernel_size=2)
self.fc1 = nn.Linear(512,128)
# self.fc2 = nn.Linear(128, 64)
# self.fc3 = nn.Linear(64, 10)
self.fc2 = nn.Linear(128, 3)
torch.nn.init.xavier_normal_(self.conv1.weight)
torch.nn.init.xavier_normal_(self.conv2.weight)
torch.nn.init.xavier_normal_(self.conv3.weight)
torch.nn.init.xavier_normal_(self.conv4.weight)
torch.nn.init.xavier_normal_(self.conv5.weight)
torch.nn.init.xavier_normal_(self.conv6.weight)
torch.nn.init.zeros_(self.conv1.bias)
torch.nn.init.zeros_(self.conv2.bias)
torch.nn.init.zeros_(self.conv3.bias)
torch.nn.init.zeros_(self.conv4.bias)
torch.nn.init.zeros_(self.conv5.bias)
torch.nn.init.zeros_(self.conv6.bias)
torch.nn.init.xavier_normal_(self.fc1.weight)
torch.nn.init.xavier_normal_(self.fc2.weight)
torch.nn.init.zeros_(self.fc1.bias)
torch.nn.init.zeros_(self.fc2.bias)
def forward(self, x):
x = self.conv1(x)
x = F.relu(self.batch_norm1(x))
x = (F.relu(self.conv2(x)))
x = self.pool(x)
x = self.conv3(x)
x = F.relu(self.batch_norm2(x))
x = (F.relu(self.conv4(x)))
x = self.pool(x)
x = self.dropout1(x)
x = self.conv5(x)
x = F.relu(self.batch_norm3(x))
x = (F.relu(self.conv6(x)))
x = self.pool(x)
#print(x.shape)
x = self.global_average_pooling(x)
x = x.squeeze()
#x = x.view(x.size(0), -1)
#print(x.shape)
x = self.dropout2(x)
x = F.relu(self.fc1(x))
#x = F.relu(self.fc2(x))
#x = self.dropout2(x)
#x = F.relu(self.fc3(x))
x = self.fc2(x)
return x
test_images =[] #list of mosaic images, each mosaic image is saved as laist of 9 images
fore_idx_test =[] #list of indexes at which foreground image is present in a mosaic image
test_label=[] # label of mosaic image = foreground class present in that mosaic
for i in range(10000):
np.random.seed(i+30000)
bg_idx = np.random.randint(0,35000,8)
fg_idx = np.random.randint(0,15000)
fg = np.random.randint(0,9)
fore_idx_test.append(fg)
image_list,label = create_mosaic_img(bg_idx,fg_idx,fg)
test_images.append(image_list)
test_label.append(label)
test_data = MosaicDataset(test_images,test_label,fore_idx_test)
test_loader = DataLoader( test_data,batch_size= batch ,shuffle=False)
criterion = nn.CrossEntropyLoss()
def my_cross_entropy(x, y,alpha,log_alpha,k):
# log_prob = -1.0 * F.log_softmax(x, 1)
# loss = log_prob.gather(1, y.unsqueeze(1))
# loss = loss.mean()
loss = criterion(x,y)
#alpha = torch.clamp(alpha,min=1e-10)
b = -1.0* alpha * log_alpha
b = torch.mean(torch.sum(b,dim=1))
closs = loss
entropy = b
loss = (1-k)*loss + ((k)*b)
return loss,closs,entropy
def calculate_attn_loss(dataloader,what,where,k):
what.eval()
where.eval()
r_loss = 0
alphas = []
lbls = []
pred = []
fidices = []
with torch.no_grad():
for i, data in enumerate(dataloader, 0):
inputs, labels,fidx = data
lbls.append(labels)
fidices.append(fidx)
inputs = inputs.double()
inputs, labels = inputs.to("cuda"),labels.to("cuda")
alpha, avg_images,log_alphas = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
pred.append(predicted.cpu().numpy())
alphas.append(alpha.cpu().numpy())
loss,_,_ = my_cross_entropy(outputs, labels, alpha, log_alphas, k)
r_loss += loss.item()
alphas = np.concatenate(alphas,axis=0)
pred = np.concatenate(pred,axis=0)
lbls = np.concatenate(lbls,axis=0)
fidices = np.concatenate(fidices,axis=0)
#print(alphas.shape,pred.shape,lbls.shape,fidices.shape)
# value>0.01
sparsity_val = np.sum(np.sum(alphas>0.01,axis=1))
# simplex distance
argmax_index = np.argmax(alphas,axis=1)
simplex_pt = np.zeros(alphas.shape)
simplex_pt[np.arange(argmax_index.size),argmax_index] = 1
shortest_distance_simplex = np.sum(np.sqrt(np.sum((alphas-simplex_pt)**2,axis=1)))
# entropy
entropy = np.sum((-alphas*np.log2(alphas)).sum(axis=1))
analysis = analyse_data(alphas,lbls,pred,fidices)
return analysis,[sparsity_val,shortest_distance_simplex ,entropy]
def analyse_data(alphas,lbls,predicted,f_idx):
'''
analysis data is created here
'''
batch = len(predicted)
amth,alth,ftpt,ffpt,ftpf,ffpf = 0,0,0,0,0,0
for j in range (batch):
focus = np.argmax(alphas[j])
if(alphas[j][focus] >= 0.5):
amth +=1
else:
alth +=1
if(focus == f_idx[j] and predicted[j] == lbls[j]):
ftpt += 1
elif(focus != f_idx[j] and predicted[j] == lbls[j]):
ffpt +=1
elif(focus == f_idx[j] and predicted[j] != lbls[j]):
ftpf +=1
elif(focus != f_idx[j] and predicted[j] != lbls[j]):
ffpf +=1
return [ftpt,ffpt,ftpf,ffpf,amth,alth]
n_seed =[0, 1, 2]
lr = [0.0005, 0.001]
k = 0 # k 0.001 0.003, 0.005
Analysis_ = {}
Train_Loss_ = []
for n_seed_ in n_seed:
for lr_ in lr:
analyse_data_train = []
analyse_data_test = []
sparsty_train = []
sparsty_test = []
tr_loss = []
print("initializing models using seed",n_seed_)
torch.manual_seed(n_seed_)
focus_net = Focus().double()
focus_net = focus_net.to("cuda")
torch.manual_seed(n_seed_)
classify = Classification().double()
classify = classify.to("cuda")
criterion = nn.CrossEntropyLoss()
print("using lr",lr_)
optimizer_focus = optim.Adam(focus_net.parameters(), lr=lr_)#, momentum=0.9)
optimizer_classify = optim.Adam(classify.parameters(), lr=lr_)#, momentum=0.9)
analysis_data_train, sparsity_value_train =calculate_attn_loss(train_loader,classify,focus_net,k)
analysis_data_test, sparsity_value_test =calculate_attn_loss(test_loader,classify,focus_net,k)
analyse_data_train.append(analysis_data_train)
analyse_data_test.append(analysis_data_test)
sparsty_train.append(sparsity_value_train)
sparsty_test.append(sparsity_value_test)
nos_epochs = 60
focus_net.train()
classify.train()
for epoch in range(nos_epochs): # loop over the dataset multiple time
focus_net.train()
classify.train()
epoch_loss = []
cnt=0
running_loss = 0
iteration = desired_num // batch
for i, data in enumerate(train_loader):
inputs , labels , fore_idx = data
inputs = inputs.double()
inputs, labels = inputs.to("cuda"), labels.to("cuda")
# zero the parameter gradients
optimizer_focus.zero_grad()
optimizer_classify.zero_grad()
alphas, avg_images,log_alphas = focus_net(inputs)
outputs = classify(avg_images)
# outputs, alphas, avg_images = classify(inputs)
_, predicted = torch.max(outputs.data, 1)
loss,_,_ = my_cross_entropy(outputs, labels,alphas,log_alphas,k)
loss.backward()
optimizer_focus.step()
optimizer_classify.step()
running_loss += loss.item()
mini = 60
if cnt % mini == mini-1: # print every 60 mini-batches
print('[%d, %5d] loss: %.3f' %(epoch + 1, cnt + 1, running_loss / mini))
epoch_loss.append(running_loss/mini)
running_loss = 0.0
cnt=cnt+1
tr_loss.append(np.mean(epoch_loss))
if epoch % 1 == 0:
analysis_data_train, sparsity_value_train = calculate_attn_loss(train_loader,classify,focus_net,k)
analysis_data_test, sparsity_value_test = calculate_attn_loss(test_loader,classify,focus_net,k)
analyse_data_train.append(analysis_data_train)
analyse_data_test.append(analysis_data_test)
sparsty_train.append(sparsity_value_train)
sparsty_test.append(sparsity_value_test)
if(np.mean(epoch_loss) <= 0.05):
break
print('Finished Training')
print("train FTPT Analysis and sparsity values",analysis_data_train,sparsity_value_train)
print("test FTPT Analysis and sparsity values",analysis_data_test,sparsity_value_test)
torch.save(focus_net.state_dict(),path+"seed_"+str(n_seed_)+"lr_"+str(lr_)+"weights_focus.pt")
torch.save(classify.state_dict(),path+"seed_"+str(n_seed_)+"lr_"+str(lr_)+"weights_classify.pt")
Analysis_["train_seed_"+str(n_seed_)+"_lr_"+str(lr_)] = np.array(analyse_data_train)
Analysis_["test_seed_"+str(n_seed_)+"_lr_"+str(lr_)] = np.array(analyse_data_test)
Analysis_["train_sparsity_"+str(n_seed_)+"_lr_"+str(lr_)] = np.array(sparsty_train)
Analysis_["test_sparsity_"+str(n_seed_)+"_lr_"+str(lr_)] = np.array(sparsty_test)
Train_Loss_.append(tr_loss)
np.save("analysis.npy",Analysis_)
np.save("training_loss.npy",Train_Loss_)
Analysis_
for i in range(len(Train_Loss_)):
plt.figure(figsize=(6,5))
plt.plot(np.arange(1,len(Train_Loss_[i])+1),Train_Loss_[i])
plt.xlabel("epochs", fontsize=14, fontweight = 'bold')
plt.ylabel("Loss", fontsize=14, fontweight = 'bold')
plt.xticks(np.arange(1,epoch+2))
plt.title("Train Loss")
# #plt.grid()
plt.show()
for seed_ in n_seed:
for lr_ in lr:
data = Analysis_['train_seed_'+ str(seed_)+"_lr_" +str(lr_)]
plt.figure(figsize=(6,5))
plt.plot(np.arange(0,len(data)),data[:,0]/300, label ="FTPT ")
plt.plot(np.arange(0,len(data)),data[:,1]/300, label ="FFPT ")
plt.plot(np.arange(0,len(data)),data[:,2]/300, label ="FTPF ")
plt.plot(np.arange(0,len(data)),data[:,3]/300, label ="FFPF ")
plt.title("On Training set " + 'seed ' + str(seed_)+" lr " +str(lr_))
#plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs", fontsize=14, fontweight = 'bold')
plt.ylabel("percentage train data", fontsize=14, fontweight = 'bold')
# plt.xlabel("epochs")
# plt.ylabel("training data")
plt.legend()
plt.savefig(path + 'seed ' + str(seed_)+" lr " +str(lr_)+ "_train.png",bbox_inches="tight")
plt.savefig(path + 'seed ' + str(seed_)+" lr " +str(lr_)+ "_train.pdf",bbox_inches="tight")
plt.grid()
plt.show()
for seed_ in n_seed:
for lr_ in lr:
data = Analysis_['test_seed_'+ str(seed_)+"_lr_" +str(lr_)]
plt.figure(figsize=(6,5))
plt.plot(np.arange(0,len(data)),data[:,0]/100, label ="FTPT ")
plt.plot(np.arange(0,len(data)),data[:,1]/100, label ="FFPT ")
plt.plot(np.arange(0,len(data)),data[:,2]/100, label ="FTPF ")
plt.plot(np.arange(0,len(data)),data[:,3]/100, label ="FFPF ")
plt.title("On Testing set " + 'seed ' + str(seed_)+" lr " +str(lr_))
#plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs", fontsize=14, fontweight = 'bold')
plt.ylabel("percentage test data", fontsize=14, fontweight = 'bold')
# plt.xlabel("epochs")
# plt.ylabel("training data")
plt.legend()
plt.savefig(path + 'seed ' + str(seed_)+" lr " +str(lr_)+"_test.png",bbox_inches="tight")
plt.savefig(path + 'seed ' + str(seed_)+" lr " +str(lr_)+ "_test.pdf",bbox_inches="tight")
plt.grid()
plt.show()
```
| github_jupyter |
# VTK tools
Pygslib use VTK:
- as data format and data converting tool
- to plot in 3D
- as a library with some basic computational geometry functions, for example to know if a point is inside a surface
Some of the functions in VTK were obtained or modified from Adamos Kyriakou at https://pyscience.wordpress.com/
```
import pygslib
import numpy as np
```
## Functions in vtktools
```
help(pygslib.vtktools)
```
## Load a cube defined in an stl file and plot it
STL is a popular mesh format included an many non-commercial and commercial software, example: Paraview, Datamine Studio, etc.
```
#load the cube
mycube=pygslib.vtktools.loadSTL('../datasets/stl/cube.stl')
# see the information about this data... Note that it is an vtkPolyData
print mycube
# Create a VTK render containing a surface (mycube)
renderer = pygslib.vtktools.polydata2renderer(mycube, color=(1,0,0), opacity=0.50, background=(1,1,1))
# Now we plot the render
pygslib.vtktools.vtk_show(renderer, camera_position=(-20,20,20), camera_focalpoint=(0,0,0))
```
## Ray casting to find intersections of a lines with the cube
This is basically how we plan to find points inside solid and to define blocks inside solid
```
# we have a line, for example a block model row
# defined by two points or an infinite line passing trough a dillhole sample
pSource = [-50.0, 0.0, 0.0]
pTarget = [50.0, 0.0, 0.0]
# now we want to see how this looks like
pygslib.vtktools.addLine(renderer,pSource, pTarget, color=(0, 1, 0))
pygslib.vtktools.vtk_show(renderer) # the camera position was already defined
# now we find the point coordinates of the intersections
intersect, points, pointsVTK= pygslib.vtktools.vtk_raycasting(mycube, pSource, pTarget)
print "the line intersects? ", intersect==1
print "the line is over the surface?", intersect==-1
# list of coordinates of the points intersecting
print points
#Now we plot the intersecting points
# To do this we add the points to the renderer
for p in points:
pygslib.vtktools.addPoint(renderer, p, radius=0.5, color=(0.0, 0.0, 1.0))
pygslib.vtktools.vtk_show(renderer)
```
### Test line on surface
```
# we have a line, for example a block model row
# defined by two points or an infinite line passing trough a dillhole sample
pSource = [-50.0, 5.01, 0]
pTarget = [50.0, 5.01, 0]
# now we find the point coordinates of the intersections
intersect, points, pointsVTK= pygslib.vtktools.vtk_raycasting(mycube, pSource, pTarget)
print "the line intersects? ", intersect==1
print "the line is over the surface?", intersect==-1
# list of coordinates of the points intersecting
print points
# now we want to see how this looks like
pygslib.vtktools.addLine(renderer,pSource, pTarget, color=(0, 1, 0))
for p in points:
pygslib.vtktools.addPoint(renderer, p, radius=0.5, color=(0.0, 0.0, 1.0))
pygslib.vtktools.vtk_show(renderer) # the camera position was already defined
# note that there is a tolerance of about 0.01
```
# Finding points
```
#using same cube but generation arbitrary random points
x = np.random.uniform(-10,10,150)
y = np.random.uniform(-10,10,150)
z = np.random.uniform(-10,10,150)
```
## Find points inside a solid
```
# selecting all inside the solid
# This two methods are equivelent but test=4 also works with open surfaces
inside,p=pygslib.vtktools.pointquering(mycube, azm=0, dip=0, x=x, y=y, z=z, test=1)
inside1,p=pygslib.vtktools.pointquering(mycube, azm=0, dip=0, x=x, y=y, z=z, test=4)
err=inside==inside1
#print inside, tuple(p)
print x[~err]
print y[~err]
print z[~err]
# here we prepare to plot the solid, the x,y,z indicator and we also
# plot the line (direction) used to ray trace
# convert the data in the STL file into a renderer and then we plot it
renderer = pygslib.vtktools.polydata2renderer(mycube, color=(1,0,0), opacity=0.70, background=(1,1,1))
# add indicator (r->x, g->y, b->z)
pygslib.vtktools.addLine(renderer,[-10,-10,-10], [-7,-10,-10], color=(1, 0, 0))
pygslib.vtktools.addLine(renderer,[-10,-10,-10], [-10,-7,-10], color=(0, 1, 0))
pygslib.vtktools.addLine(renderer,[-10,-10,-10], [-10,-10,-7], color=(0, 0, 1))
# add ray to see where we are pointing
pygslib.vtktools.addLine(renderer, (0.,0.,0.), tuple(p), color=(0, 0, 0))
# here we plot the points selected and non-selected in different color and size
# add the points selected
for i in range(len(inside)):
p=[x[i],y[i],z[i]]
if inside[i]!=0:
#inside
pygslib.vtktools.addPoint(renderer, p, radius=0.5, color=(0.0, 0.0, 1.0))
else:
pygslib.vtktools.addPoint(renderer, p, radius=0.2, color=(0.0, 1.0, 0.0))
#lets rotate a bit this
pygslib.vtktools.vtk_show(renderer, camera_position=(0,0,50), camera_focalpoint=(0,0,0))
```
# Find points over a surface
```
# selecting all over a solid (test = 2)
inside,p=pygslib.vtktools.pointquering(mycube, azm=0, dip=0, x=x, y=y, z=z, test=2)
# here we prepare to plot the solid, the x,y,z indicator and we also
# plot the line (direction) used to ray trace
# convert the data in the STL file into a renderer and then we plot it
renderer = pygslib.vtktools.polydata2renderer(mycube, color=(1,0,0), opacity=0.70, background=(1,1,1))
# add indicator (r->x, g->y, b->z)
pygslib.vtktools.addLine(renderer,[-10,-10,-10], [-7,-10,-10], color=(1, 0, 0))
pygslib.vtktools.addLine(renderer,[-10,-10,-10], [-10,-7,-10], color=(0, 1, 0))
pygslib.vtktools.addLine(renderer,[-10,-10,-10], [-10,-10,-7], color=(0, 0, 1))
# add ray to see where we are pointing
pygslib.vtktools.addLine(renderer, (0.,0.,0.), tuple(-p), color=(0, 0, 0))
# here we plot the points selected and non-selected in different color and size
# add the points selected
for i in range(len(inside)):
p=[x[i],y[i],z[i]]
if inside[i]!=0:
#inside
pygslib.vtktools.addPoint(renderer, p, radius=0.5, color=(0.0, 0.0, 1.0))
else:
pygslib.vtktools.addPoint(renderer, p, radius=0.2, color=(0.0, 1.0, 0.0))
#lets rotate a bit this
pygslib.vtktools.vtk_show(renderer, camera_position=(0,0,50), camera_focalpoint=(0,0,0))
```
# Find points below a surface
```
# selecting all over a solid (test = 2)
inside,p=pygslib.vtktools.pointquering(mycube, azm=0, dip=0, x=x, y=y, z=z, test=3)
# here we prepare to plot the solid, the x,y,z indicator and we also
# plot the line (direction) used to ray trace
# convert the data in the STL file into a renderer and then we plot it
renderer = pygslib.vtktools.polydata2renderer(mycube, color=(1,0,0), opacity=0.70, background=(1,1,1))
# add indicator (r->x, g->y, b->z)
pygslib.vtktools.addLine(renderer,[-10,-10,-10], [-7,-10,-10], color=(1, 0, 0))
pygslib.vtktools.addLine(renderer,[-10,-10,-10], [-10,-7,-10], color=(0, 1, 0))
pygslib.vtktools.addLine(renderer,[-10,-10,-10], [-10,-10,-7], color=(0, 0, 1))
# add ray to see where we are pointing
pygslib.vtktools.addLine(renderer, (0.,0.,0.), tuple(p), color=(0, 0, 0))
# here we plot the points selected and non-selected in different color and size
# add the points selected
for i in range(len(inside)):
p=[x[i],y[i],z[i]]
if inside[i]!=0:
#inside
pygslib.vtktools.addPoint(renderer, p, radius=0.5, color=(0.0, 0.0, 1.0))
else:
pygslib.vtktools.addPoint(renderer, p, radius=0.2, color=(0.0, 1.0, 0.0))
#lets rotate a bit this
pygslib.vtktools.vtk_show(renderer, camera_position=(0,0,50), camera_focalpoint=(0,0,0))
```
## Export points to a VTK file
```
data = {'inside': inside}
pygslib.vtktools.points2vtkfile('points', x,y,z, data)
```
The results can be ploted in an external viewer, for example mayavi or paraview:
<img src="figures/Fig_paraview.png">
| github_jupyter |
# AutoGluon Tabular with SageMaker
[AutoGluon](https://github.com/awslabs/autogluon) automates machine learning tasks enabling you to easily achieve strong predictive performance in your applications. With just a few lines of code, you can train and deploy high-accuracy deep learning models on tabular, image, and text data.
This notebook shows how to use AutoGluon-Tabular with Amazon SageMaker by creating custom containers.
## Prerequisites
If using a SageMaker hosted notebook, select kernel `conda_mxnet_p36`.
```
import subprocess
# Make sure docker compose is set up properly for local mode
subprocess.run("./setup.sh", shell=True)
# For Studio
subprocess.run("apt-get update -y", shell=True)
subprocess.run("apt install unzip", shell=True)
import os
import sys
import boto3
import sagemaker
from time import sleep
from collections import Counter
import numpy as np
import pandas as pd
from sagemaker import get_execution_role, local, Model, utils, s3
from sagemaker.estimator import Estimator
from sagemaker.predictor import Predictor
from sagemaker.serializers import CSVSerializer
from sagemaker.deserializers import StringDeserializer
from sklearn.metrics import accuracy_score, classification_report
from IPython.core.display import display, HTML
from IPython.core.interactiveshell import InteractiveShell
# Print settings
InteractiveShell.ast_node_interactivity = "all"
pd.set_option('display.max_columns', 500)
pd.set_option('display.max_rows', 10)
# Account/s3 setup
session = sagemaker.Session()
local_session = local.LocalSession()
bucket = session.default_bucket()
prefix = 'sagemaker/autogluon-tabular'
region = session.boto_region_name
role = get_execution_role()
client = session.boto_session.client(
"sts", region_name=region, endpoint_url=utils.sts_regional_endpoint(region)
)
account = client.get_caller_identity()['Account']
registry_uri_training = sagemaker.image_uris.retrieve('mxnet', region, version= '1.7.0', py_version='py3', instance_type='ml.m5.2xlarge', image_scope='training')
registry_uri_inference = sagemaker.image_uris.retrieve('mxnet', region, version= '1.7.0', py_version='py3', instance_type='ml.m5.2xlarge', image_scope='inference')
ecr_uri_prefix = account +'.'+'.'.join(registry_uri_training.split('/')[0].split('.')[1:])
```
### Build docker images
Build the training/inference image and push to ECR
```
training_algorithm_name = 'autogluon-sagemaker-training'
inference_algorithm_name = 'autogluon-sagemaker-inference'
```
First, you may want to remove existing docker images to make a room to build autogluon containers.
```
subprocess.run("docker system prune -af", shell=True)
subprocess.run(f"/bin/bash ./container-training/build_push_training.sh {account} {region} {training_algorithm_name} {ecr_uri_prefix} {registry_uri_training.split('/')[0].split('.')[0]} {registry_uri_training}", shell=True)
subprocess.run("docker system prune -af", shell=True)
subprocess.run(f"/bin/bash ./container-inference/build_push_inference.sh {account} {region} {inference_algorithm_name} {ecr_uri_prefix} {registry_uri_training.split('/')[0].split('.')[0]} {registry_uri_inference}", shell=True)
subprocess.run("docker system prune -af", shell=True)
```
### Alternative way of building docker images using sm-docker
The new Amazon SageMaker Studio Image Build convenience package allows data scientists and developers to easily build custom container images from your Studio notebooks via a new CLI.
Newly built Docker images are tagged and pushed to Amazon ECR.
To use the CLI, you need to ensure the Amazon SageMaker execution role used by your Studio notebook environment (or another AWS Identity and Access Management (IAM) role, if you prefer) has the required permissions to interact with the resources used by the CLI, including access to CodeBuild and Amazon ECR. Your role should have a trust policy with CodeBuild.
You also need to make sure the appropriate permissions are included in your role to run the build in CodeBuild, create a repository in Amazon ECR, and push images to that repository.
See also: https://aws.amazon.com/blogs/machine-learning/using-the-amazon-sagemaker-studio-image-build-cli-to-build-container-images-from-your-studio-notebooks/
```
#subprocess.run("pip install sagemaker-studio-image-build", shell=True)
'''
training_repo_name = training_algorithm_name + ':latest'
training_repo_name
!sm-docker build . --repository {training_repo_name} \
--file ./container-training/Dockerfile.training --build-arg REGISTRY_URI={registry_uri_training}
inference_repo_name = inference_algorithm_name + ':latest'
inference_repo_name
!sm-docker build . --repository {inference_repo_name} \
--file ./container-inference/Dockerfile.inference --build-arg REGISTRY_URI={registry_uri_inference}
'''
```
### Get the data
In this example we'll use the direct-marketing dataset to build a binary classification model that predicts whether customers will accept or decline a marketing offer.
First we'll download the data and split it into train and test sets. AutoGluon does not require a separate validation set (it uses bagged k-fold cross-validation).
```
# Download and unzip the data
subprocess.run(f"aws s3 cp --region {region} s3://sagemaker-sample-data-{region}/autopilot/direct_marketing/bank-additional.zip .", shell=True)
subprocess.run("unzip -qq -o bank-additional.zip", shell=True)
subprocess.run("rm bank-additional.zip", shell=True)
local_data_path = './bank-additional/bank-additional-full.csv'
data = pd.read_csv(local_data_path)
# Split train/test data
train = data.sample(frac=0.7, random_state=42)
test = data.drop(train.index)
# Split test X/y
label = 'y'
y_test = test[label]
X_test = test.drop(columns=[label])
```
##### Check the data
```
train.head(3)
train.shape
test.head(3)
test.shape
X_test.head(3)
X_test.shape
```
Upload the data to s3
```
train_file = 'train.csv'
train.to_csv(train_file,index=False)
train_s3_path = session.upload_data(train_file, key_prefix='{}/data'.format(prefix))
test_file = 'test.csv'
test.to_csv(test_file,index=False)
test_s3_path = session.upload_data(test_file, key_prefix='{}/data'.format(prefix))
X_test_file = 'X_test.csv'
X_test.to_csv(X_test_file,index=False)
X_test_s3_path = session.upload_data(X_test_file, key_prefix='{}/data'.format(prefix))
```
## Hyperparameter Selection
The minimum required settings for training is just a target label, `init_args['label']`.
Additional optional hyperparameters can be passed to the `autogluon.tabular.TabularPredictor.fit` function via `fit_args`.
Below shows a more in depth example of AutoGluon-Tabular hyperparameters from the example [Predicting Columns in a Table - In Depth](https://auto.gluon.ai/stable/tutorials/tabular_prediction/tabular-indepth.html). Please see [fit parameters](https://auto.gluon.ai/stable/_modules/autogluon/tabular/predictor/predictor.html#TabularPredictor) for further information. Note that in order for hyperparameter ranges to work in SageMaker, values passed to the `fit_args['hyperparameters']` must be represented as strings.
```python
nn_options = {
'num_epochs': "10",
'learning_rate': "ag.space.Real(1e-4, 1e-2, default=5e-4, log=True)",
'activation': "ag.space.Categorical('relu', 'softrelu', 'tanh')",
'layers': "ag.space.Categorical([100],[1000],[200,100],[300,200,100])",
'dropout_prob': "ag.space.Real(0.0, 0.5, default=0.1)"
}
gbm_options = {
'num_boost_round': "100",
'num_leaves': "ag.space.Int(lower=26, upper=66, default=36)"
}
model_hps = {'NN': nn_options, 'GBM': gbm_options}
init_args = {
'eval_metric' : 'roc_auc'
'label': 'y'
}
fit_args = {
'presets': ['best_quality', 'optimize_for_deployment'],
'time_limits': 60*10,
'hyperparameters': model_hps,
'hyperparameter_tune': True,
'search_strategy': 'skopt'
}
hyperparameters = {
'fit_args': fit_args,
'feature_importance': True
}
```
**Note:** Your hyperparameter choices may affect the size of the model package, which could result in additional time taken to upload your model and complete training. Including `'optimize_for_deployment'` in the list of `fit_args['presets']` is recommended to greatly reduce upload times.
<br>
```
# Define required label and optional additional parameters
init_args = {
'label': 'y'
}
# Define additional parameters
fit_args = {
# Adding 'best_quality' to presets list will result in better performance (but longer runtime)
'presets': ['optimize_for_deployment'],
}
# Pass fit_args to SageMaker estimator hyperparameters
hyperparameters = {
'init_args': init_args,
'fit_args': fit_args,
'feature_importance': True
}
tags = [{
'Key' : 'AlgorithmName',
'Value' : 'AutoGluon-Tabular'
}]
```
## Train
For local training set `train_instance_type` to `local` .
For non-local training the recommended instance type is `ml.m5.2xlarge`.
**Note:** Depending on how many underlying models are trained, `train_volume_size` may need to be increased so that they all fit on disk.
```
%%time
instance_type = 'ml.m5.2xlarge'
#instance_type = 'local'
ecr_image = f'{ecr_uri_prefix}/{training_algorithm_name}:latest'
estimator = Estimator(image_uri=ecr_image,
role=role,
instance_count=1,
instance_type=instance_type,
hyperparameters=hyperparameters,
volume_size=100,
tags=tags)
# Set inputs. Test data is optional, but requires a label column.
inputs = {'training': train_s3_path, 'testing': test_s3_path}
estimator.fit(inputs)
```
### Review the performance of the trained model
```
from utils.ag_utils import launch_viewer
launch_viewer(is_debug=False)
```
### Create Model
```
# Create predictor object
class AutoGluonTabularPredictor(Predictor):
def __init__(self, *args, **kwargs):
super().__init__(*args,
serializer=CSVSerializer(),
deserializer=StringDeserializer(), **kwargs)
ecr_image = f'{ecr_uri_prefix}/{inference_algorithm_name}:latest'
if instance_type == 'local':
model = estimator.create_model(image_uri=ecr_image, role=role)
else:
#model_uri = os.path.join(estimator.output_path, estimator._current_job_name, "output", "model.tar.gz")
model_uri = estimator.model_data
model = Model(ecr_image, model_data=model_uri, role=role, sagemaker_session=session, predictor_cls=AutoGluonTabularPredictor)
```
### Batch Transform
For local mode, either `s3://<bucket>/<prefix>/output/` or `file:///<absolute_local_path>` can be used as outputs.
By including the label column in the test data, you can also evaluate prediction performance (In this case, passing `test_s3_path` instead of `X_test_s3_path`).
```
output_path = f's3://{bucket}/{prefix}/output/'
# output_path = f'file://{os.getcwd()}'
transformer = model.transformer(instance_count=1,
instance_type=instance_type,
strategy='MultiRecord',
max_payload=6,
max_concurrent_transforms=1,
output_path=output_path)
transformer.transform(test_s3_path, content_type='text/csv', split_type='Line')
transformer.wait()
```
### Endpoint
##### Deploy remote or local endpoint
```
instance_type = 'ml.m5.2xlarge'
#instance_type = 'local'
predictor = model.deploy(initial_instance_count=1,
instance_type=instance_type)
```
##### Attach to endpoint (or reattach if kernel was restarted)
```
# Select standard or local session based on instance_type
if instance_type == 'local':
sess = local_session
else:
sess = session
# Attach to endpoint
predictor = AutoGluonTabularPredictor(predictor.endpoint_name, sagemaker_session=sess)
```
##### Predict on unlabeled test data
```
results = predictor.predict(X_test.to_csv(index=False)).splitlines()
# Check output
threshold = 0.5
y_results = np.array(['yes' if float(i.split(",")[1]) > threshold else 'no' for i in results])
print(Counter(y_results))
```
##### Predict on data that includes label column
Prediction performance metrics will be printed to endpoint logs.
```
results = predictor.predict(test.to_csv(index=False)).splitlines()
# Check output
threshold = 0.5
y_results = np.array(['yes' if float(i.split(",")[1]) > threshold else 'no' for i in results])
print(Counter(y_results))
```
##### Check that classification performance metrics match evaluation printed to endpoint logs as expected
```
threshold = 0.5
y_results = np.array(['yes' if float(i.split(",")[1]) > threshold else 'no' for i in results])
print("accuracy: {}".format(accuracy_score(y_true=y_test, y_pred=y_results)))
print(classification_report(y_true=y_test, y_pred=y_results, digits=6))
```
##### Clean up endpoint
```
predictor.delete_endpoint()
```
## Explainability with Amazon SageMaker Clarify
There are growing business needs and legislative regulations that require explainations of why a model made a certain decision. SHAP (SHapley Additive exPlanations) is an approach to explain the output of machine learning models. SHAP values represent a feature's contribution to a change in the model output. SageMaker Clarify uses SHAP to explain the contribution that each input feature makes to the final decision.
##### Set parameters for SHAP calculation
```
seed = 0
num_rows = 500
#Write a csv file used by SageMaker Clarify
test_explainavility_file = 'test_explainavility.csv'
train.head(num_rows).to_csv(test_explainavility_file, index=False, header=False)
test_explainavility_s3_path = session.upload_data(test_explainavility_file, key_prefix='{}/data'.format(prefix))
```
##### Specify computing resources
```
from sagemaker import clarify
model_name = estimator.latest_training_job.job_name
container_def = model.prepare_container_def()
session.create_model(model_name,
role,
container_def)
clarify_processor = clarify.SageMakerClarifyProcessor(role=role,
instance_count=1,
instance_type='ml.c4.xlarge',
sagemaker_session=session)
model_config = clarify.ModelConfig(model_name=model_name,
instance_type='ml.c5.xlarge',
instance_count=1,
accept_type='text/csv')
```
##### Run a SageMaker Clarify job
```
shap_config = clarify.SHAPConfig(baseline=X_test.sample(15, random_state=seed).values.tolist(),
num_samples=100,
agg_method='mean_abs')
explainability_output_path = 's3://{}/{}/{}/clarify-explainability'.format(bucket, prefix, model_name)
explainability_data_config = clarify.DataConfig(s3_data_input_path=test_explainavility_s3_path,
s3_output_path=explainability_output_path,
label='y',
headers=train.columns.to_list(),
dataset_type='text/csv')
predictions_config = clarify.ModelPredictedLabelConfig(probability_threshold=0.5)
clarify_processor.run_explainability(data_config=explainability_data_config,
model_config=model_config,
explainability_config=shap_config)
```
##### View the Explainability Report
You can view the explainability report in Studio under the experiments tab. If you're not a Studio user yet, as with the Bias Report, you can access this report at the following S3 bucket.
```
subprocess.run(f"aws s3 cp {explainability_output_path} . --recursive", shell=True)
```
Global explanatory methods allow understanding the model and its feature contributions in aggregate over multiple datapoints. Here we show an aggregate bar plot that plots the mean absolute SHAP value for each feature.
```
subprocess.run(f"{sys.executable} -m pip install shap", shell=True)
```
##### Compute global shap values out of out.csv
```
shap_values_ = pd.read_csv('explanations_shap/out.csv')
shap_values_.abs().mean().to_dict()
num_features = len(train.head(num_rows).drop(['y'], axis = 1).columns)
import shap
shap_values = [shap_values_.to_numpy()[:,:num_features], shap_values_.to_numpy()[:,num_features:]]
shap.summary_plot(shap_values,
plot_type='bar',
feature_names=train.head(num_rows).drop(['y'], axis = 1).columns.tolist())
```
The detailed summary plot below can provide more context over the above bar chart. It tells which features are most important and, in addition, their range of effects over the dataset. The color allows us to match how changes in the value of a feature effect the change in prediction. The 'red' indicates higher value of the feature and 'blue' indicates lower (normalized over the features).
```
shap.summary_plot(shap_values_[shap_values_.columns[20:]].to_numpy(),
train.head(num_rows).drop(['y'], axis = 1))
```
| github_jupyter |
# Hybrid audit examples
```
from __future__ import division, print_function
%matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
import math
import numpy as np
import numpy.random
import scipy as sp
import scipy.stats
from ballot_comparison import findNmin_ballot_comparison_rates
from hypergeometric import trihypergeometric_optim, simulate_ballot_polling_power
from fishers_combination import simulate_fisher_combined_audit, calculate_lambda_range
```
# Example 1 - medium sized election, close race
There are two strata. One contains every CVR county and the other contains every no-CVR county.
There were 110,000 ballots cast in the election, 100,000 in the CVR stratum and 10,000 in the no-CVR stratum.
In the CVR stratum, there were 45,500 votes reported for A, 49,500 votes for candidate B, and 5,000 invalid ballots.
In the no-CVR stratum, there were 7,500 votes reported for A, 1,500 votes for B, and 1000 invalid ballots.
A won overall, with 53,000 votes to B's 51,000, but not in the CVR stratum.
The reported vote margin between A and B is 2,000 votes, a "diluted margin" of $2,000/110,000 = 1.8\%$.
Candidate | Stratum 1 | Stratum 2 | total
---|---|---|---
A | 45,500 | 7,500 | 53,000
B | 49,500 | 1,500 | 51,000
Ballots | 100,000 | 10,000 | 110,000
Diluted margin | -4% | 60% | 1.8%
The reported outcome of the election is correct if, for every $\lambda$, either the overstatement of the margin in the CVR stratum is less than $2000\lambda$ votes or the overstatement of the margin in the no-CVR stratum is less than $2000(1-\lambda)$ votes.
We want to limit the risk of certifying an incorrect outcome to at most $\alpha=10\%$.
# Using Fisher's method to combine the audits
```
alpha = 0.1
N_w1 = 45500
N_l1 = 49500
N_w2 = 7500
N_l2 = 1500
N1 = 100000
N2 = 10000
margin = (N_w1 + N_w2 - N_l1 - N_l2)
calculate_lambda_range(N_w1, N_l1, N1, N_w2, N_l2, N2)
```
By plotting the Fisher's combined $p$-value along a grid, we determined that the Fisher's combined $p$-value is maximized somewhere on $[0, 1]$. Below, we restrict the search to that region.
```
np.random.seed(20180514)
n1 = 750
n2 = 500
power = simulate_fisher_combined_audit(N_w1, N_l1, N1, N_w2, N_l2, N2, n1, n2, alpha,
reps=10000, feasible_lambda_range=(-7.0, 3.0))
print("In 10000 simulations with a CVR stratum sample size of 700 ballots and \
\n no-CVR stratum sample size of 500 ballots, the rate of stopping the audit is ", \
power)
```
# What if we could do a ballot-comparison audit for the entire contest?
With current technology, this isn't possible. We'll use a risk limit of 10% to be consistent with the example above.
```
# Assuming that the audit will find no errors
findNmin_ballot_comparison_rates(alpha=alpha, gamma=1.03905,
r1=0, s1=0, r2=0, s2=0,
reported_margin=margin, N=N1+N2,
null_lambda=1)
# Assuming that the audit will find 1-vote overstatements at rate 0.1%
findNmin_ballot_comparison_rates(alpha=alpha, gamma=1.03905,
r1=0.001, s1=0, r2=0, s2=0,
reported_margin=margin, N=N1+N2,
null_lambda=1)
```
# Instead, what if we did an inefficient approach?
In Section 2.3 of the paper, we suggest a simple-but-pessimistic approach: sample uniformly from all counties as if one were performing a ballot-level comparison audit everywhere, but to treat any ballot selected from a legacy county as a two-vote overstatement.
In this example, $10,000/1,100,000 \approx 9\%$ of ballots come from the no-CVR stratum. We find that we'd proceed to a full hand count.
```
# Assuming that the audit will find no errors
findNmin_ballot_comparison_rates(alpha=alpha, gamma=1.03905,
r1=0, s1=0, r2=N1/(N1+N2), s2=0,
reported_margin=margin, N=N1+N2, null_lambda=1)
```
If, instead, the margin were a bit larger (in this example, let's say 10,000 votes) and the no-CVR counties made up only 1.2% of total ballots, things would be more favorable.
```
# Assuming that the audit will find no errors
findNmin_ballot_comparison_rates(alpha=alpha, gamma=1.03905,
r1=0, s1=0, r2=0.012, s2=0,
reported_margin=10000, N=N1+N2, null_lambda=1)
```
# Example 2 - large election, large margin
There are two strata. One contains every CVR county and the other contains every no-CVR county.
There were 2 million ballots cast in the election, 1.9 million in the CVR stratum and 100,000 in the no-CVR stratum.
In the CVR stratum, the diluted margin was $21\%$: there were 1,102,000 votes reported for A, 703,000 votes reported for candidate B, and 76,000 invalid ballots.
In the no-CVR stratum, the diluted margin was $-10\%$: there were 42,500 votes reported for A, 52,500 votes for B, and 5,000 invalid ballots.
A won overall, with 1,144,500 votes to B's 755,500, but not in the CVR stratum.
The reported vote margin between A and B is 389,000 votes, a "diluted margin" of $389,000/2,000,000 = 19.45\%$.
Candidate | Stratum 1 | Stratum 2 | total
---|---|---|---
A | 1,102,000 | 42,500 | 1,144,500
B | 703,000 | 52,500 | 755,500
Ballots | 1,900,000 | 100,000 | 2,000,000
Diluted margin | 21% | -10% | 19.45%
We want to limit the risk of certifying an incorrect outcome to at most $\alpha=5\%$.
# Using Fisher's method to combine the audits
```
alpha = 0.05
N1 = 1900000
N2 = 100000
N_w1 = 1102000
N_l1 = 703000
N_w2 = 42500
N_l2= 52500
margin = (N_w1 + N_w2 - N_l1 - N_l2)
calculate_lambda_range(N_w1, N_l1, N1, N_w2, N_l2, N2)
```
By plotting the Fisher's combined $p$-value along a grid, we determined that the Fisher's combined $p$-value is maximized somewhere on $[0.5, 1.5]$. Below, we restrict the search to that region.
```
np.random.seed(20180514)
n1 = 50
n2 = 25
power = simulate_fisher_combined_audit(N_w1, N_l1, N1, N_w2, N_l2, N2, n1, n2, alpha,
reps=10000, feasible_lambda_range = (0.5, 2))
print("In 10,000 simulations with a CVR stratum sample size of 50 ballots \n \
and no-CVR stratum sample size of 25 ballots, the rate of stopping the audit is ", \
power)
```
# What if we could do a ballot-comparison audit for the entire contest?
With current technology, this isn't possible.
```
# Assuming that the audit will find no errors
findNmin_ballot_comparison_rates(alpha=alpha, gamma=1.03905,
r1=0, s1=0, r2=0, s2=0,
reported_margin=margin, N=N1+N2, null_lambda=1)
# Assuming that the audit will find 1-vote overstatements at rate 0.1%
findNmin_ballot_comparison_rates(alpha=alpha, gamma=1.03905,
r1=0.001, s1=0, r2=0, s2=0,
reported_margin=margin, N=N1+N2, null_lambda=1)
```
# Instead, what if we did an inefficient approach?
In Section 2.3 of the paper, we suggest a simple-but-pessimistic approach: sample uniformly from all counties as if one were performing a ballot-level comparison audit everywhere, but to treat any ballot selected from a legacy county as a two-vote overstatement.
In this example, $100,000/2,000,000 = 5\%$ of ballots come from the no-CVR stratum. That is large enough
that if we treat all ballots sampled from the no-CVR stratum as 2-vote overstatemnts, the audit would be expected to require a full hand count.
```
# Assuming that the audit will find no errors
findNmin_ballot_comparison_rates(alpha=alpha, gamma=1.03905,
r1=0, s1=0, r2=N1/(N1+N2), s2=0,
reported_margin=margin, N=N1+N2, null_lambda=1)
```
| github_jupyter |
# A Primer on Bayesian Methods for Multilevel Modeling
Hierarchical or multilevel modeling is a generalization of regression modeling.
*Multilevel models* are regression models in which the constituent model parameters are given **probability models**. This implies that model parameters are allowed to **vary by group**.
Observational units are often naturally **clustered**. Clustering induces dependence between observations, despite random sampling of clusters and random sampling within clusters.
A *hierarchical model* is a particular multilevel model where parameters are nested within one another.
Some multilevel structures are not hierarchical.
* e.g. "country" and "year" are not nested, but may represent separate, but overlapping, clusters of parameters
We will motivate this topic using an environmental epidemiology example.
### Example: Radon contamination (Gelman and Hill 2006)
Let's revisit the radon contamination example from the previous section. For hierarchical modeling, we will use more of the data; we will focus on modeling radon levels in Minnesota. The EPA did the radon study in 80,000 houses. There were two important predictors:
* measurement in basement or first floor (radon higher in basements)
* county uranium level (positive correlation with radon levels)
The hierarchy in this example is households within county.
### Data organization
First, we import the data from a local file, and extract Minnesota's data.
```
%matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns; sns.set_context('notebook')
import warnings
warnings.simplefilter("ignore")
# Import radon data
radon_data = pd.read_csv('../data/radon.csv', index_col=0)
RANDOM_SEED = 20090425
counties = radon_data.county.unique()
n_counties = counties.shape[0]
county = radon_data.county_code.values
log_radon = radon_data.log_radon.values
floor_measure = radon_data.floor.values
log_uranium = np.log(radon_data.Uppm.values)
county_lookup = dict(zip(counties, np.arange(n_counties)))
```
Distribution of radon levels in MN (log scale):
```
radon_data.activity.apply(lambda x: np.log(x+0.1)).hist(bins=25, grid=False);
```
## Conventional approaches
The two conventional alternatives to modeling radon exposure represent the two extremes of the bias-variance tradeoff:
***Complete pooling***:
Treat all counties the same, and estimate a single radon level.
$$y_i = \alpha + \beta x_i + \epsilon_i$$
***No pooling***:
Model radon in each county independently.
$$y_i = \alpha_{j[i]} + \beta x_i + \epsilon_i$$
where $j = 1,\ldots,85$
The errors $\epsilon_i$ may represent measurement error, temporal within-house variation, or variation among houses.
Here are the point estimates of the slope and intercept for the complete pooling model:
```
from pymc3 import Model, sample, Normal, HalfCauchy, Uniform
floor = radon_data.floor.values
log_radon = radon_data.log_radon.values
with Model() as pooled_model:
β = Normal('β', 0, sigma=10, shape=2)
σ = HalfCauchy('σ', 5)
θ = β[0] + β[1]*floor
y = Normal('y', θ, sigma=σ, observed=log_radon)
```
Before sampling, it is useful to conduct **prior predictive checks** to ensure that our priors are appropriate, and do not unduly constrain inference.
```
from pymc3 import sample_prior_predictive
with pooled_model:
prior_checks = sample_prior_predictive(samples=1000)
plt.plot(
[0, 1],
[prior_checks["β"][:, 0], prior_checks["β"][:, 1]],
"ok",
alpha=0.2)
plt.xlabel("Floor measurement location")
plt.xticks([0,1], ["Basement", "Floor"])
plt.ylabel("Mean log radon level");
```
These look fine--they allow for any radon values that we would reasonably expect to see in the data.
```
with pooled_model:
pooled_trace = sample(1000, tune=1000, cores=2)
b0, m0 = pooled_trace['β'].mean(axis=0)
plt.scatter(radon_data.floor, np.log(radon_data.activity+0.1))
xvals = np.linspace(-0.2, 1.2)
plt.plot(xvals, m0*xvals+b0, 'r--');
```
Estimates of county radon levels for the unpooled model:
```
with Model() as unpooled_model:
β0 = Normal('β0', 0, sigma=10, shape=n_counties)
β1 = Normal('β1', 0, sigma=10)
σ = HalfCauchy('σ', 5)
θ = β0[county] + β1*floor
y = Normal('y', θ, sigma=σ, observed=log_radon)
with unpooled_model:
unpooled_trace = sample(1000, tune=1000, cores=2, random_seed=RANDOM_SEED)
from arviz import plot_forest
(ax,) = plot_forest(unpooled_trace, var_names=['β0'])
ax.set_yticklabels([""]);
unpooled_estimates = pd.Series(unpooled_trace['β0'].mean(axis=0), index=counties)
unpooled_se = pd.Series(unpooled_trace['β0'].std(axis=0), index=counties)
```
We can plot the ordered estimates to identify counties with high radon levels:
```
order = unpooled_estimates.sort_values().index
plt.scatter(range(len(unpooled_estimates)), unpooled_estimates[order])
for i, m, se in zip(range(len(unpooled_estimates)), unpooled_estimates[order], unpooled_se[order]):
plt.plot([i,i], [m-se, m+se], 'b-')
plt.xlim(-1,86); plt.ylim(-1,4)
plt.ylabel('Radon estimate');plt.xlabel('Ordered county');
```
Here are visual comparisons between the pooled and unpooled estimates for a subset of counties representing a range of sample sizes.
```
sample_counties = ('LAC QUI PARLE', 'AITKIN', 'KOOCHICHING',
'DOUGLAS', 'CLAY', 'STEARNS', 'RAMSEY', 'ST LOUIS')
fig, axes = plt.subplots(2, 4, figsize=(12, 6), sharey=True, sharex=True)
axes = axes.ravel()
m = unpooled_trace['β1'].mean()
for i,c in enumerate(sample_counties):
y = radon_data.log_radon[radon_data.county==c]
x = radon_data.floor[radon_data.county==c]
axes[i].scatter(x + np.random.randn(len(x))*0.01, y, alpha=0.4)
# No pooling model
b = unpooled_estimates[c]
# Plot both models and data
xvals = np.linspace(-0.2, 1.2)
axes[i].plot(xvals, m*xvals+b)
axes[i].plot(xvals, m0*xvals+b0, 'r--')
axes[i].set_xticks([0,1])
axes[i].set_xticklabels(['basement', 'floor'])
axes[i].set_ylim(-1, 3)
axes[i].set_title(c)
if not i%2:
axes[i].set_ylabel('log radon level')
```
Neither of these models are satisfactory:
* if we are trying to identify high-radon counties, pooling is useless
* we do not trust extreme unpooled estimates produced by models using few observations
## Multilevel and hierarchical models
When we pool our data, we imply that they are sampled from the same model. This ignores any variation among sampling units (other than sampling variance):

When we analyze data unpooled, we imply that they are sampled independently from separate models. At the opposite extreme from the pooled case, this approach claims that differences between sampling units are to large to combine them:

In a hierarchical model, parameters are viewed as a sample from a population distribution of parameters. Thus, we view them as being neither entirely different or exactly the same. This is ***parital pooling***.

We can use PyMC to easily specify multilevel models, and fit them using Markov chain Monte Carlo.
## Partial pooling model
The simplest partial pooling model for the household radon dataset is one which simply estimates radon levels, without any predictors at any level. A partial pooling model represents a compromise between the pooled and unpooled extremes, approximately a weighted average (based on sample size) of the unpooled county estimates and the pooled estimates.
$$\hat{\alpha} \approx \frac{(n_j/\sigma_y^2)\bar{y}_j + (1/\sigma_{\alpha}^2)\bar{y}}{(n_j/\sigma_y^2) + (1/\sigma_{\alpha}^2)}$$
Estimates for counties with smaller sample sizes will shrink towards the state-wide average.
Estimates for counties with larger sample sizes will be closer to the unpooled county estimates.
```
with Model() as partial_pooling:
# Priors
μ_a = Normal('μ_a', mu=0., sigma=10)
σ_a = HalfCauchy('σ_a', 5)
# Random intercepts
a = Normal('a', mu=μ_a, sigma=σ_a, shape=n_counties)
# Model error
σ_y = HalfCauchy('σ_y',5)
# Expected value
y_hat = a[county]
# Data likelihood
y_like = Normal('y_like', mu=y_hat, sigma=σ_y, observed=log_radon)
with partial_pooling:
partial_pooling_trace = sample(1000, tune=1000, cores=2, random_seed=RANDOM_SEED)
sample_trace = partial_pooling_trace['a']
fig, axes = plt.subplots(1, 2, figsize=(14,6), sharex=True, sharey=True)
n_samples, n_counties = sample_trace.shape
jitter = np.random.normal(scale=0.1, size=n_counties)
n_county = radon_data.groupby('county')['county_code'].count()
unpooled_means = radon_data.groupby('county')['log_radon'].mean()
unpooled_sd = radon_data.groupby('county')['log_radon'].std()
unpooled = pd.DataFrame({'n':n_county, 'm':unpooled_means, 'sd':unpooled_sd})
unpooled['se'] = unpooled.sd/np.sqrt(unpooled.n)
axes[0].plot(unpooled.n + jitter, unpooled.m, 'b.')
for j, row in zip(jitter, unpooled.iterrows()):
name, dat = row
axes[0].plot([dat.n+j,dat.n+j], [dat.m-dat.se, dat.m+dat.se], 'b-')
axes[0].set_xscale('log')
axes[0].hlines(sample_trace.mean(), 0.9, 100, linestyles='--')
n_samples, n_counties = sample_trace.shape
means = sample_trace.mean(axis=0)
sd = sample_trace.std(axis=0)
axes[1].scatter(n_county.values + jitter, means)
axes[1].set_xscale('log')
axes[1].set_xlim(1,100)
axes[1].set_ylim(0, 3)
axes[1].hlines(sample_trace.mean(), 0.9, 100, linestyles='--')
for j,n,m,s in zip(jitter, n_county.values, means, sd):
axes[1].plot([n+j]*2, [m-s, m+s], 'b-');
```
Notice the difference between the unpooled and partially-pooled estimates, particularly at smaller sample sizes. The former are both more extreme and more imprecise.
## Varying intercept model
This model allows intercepts to vary across county, according to a random effect.
$$y_i = \alpha_{j[i]} + \beta x_{i} + \epsilon_i$$
where
$$\epsilon_i \sim N(0, \sigma_y^2)$$
and the intercept random effect:
$$\alpha_{j[i]} \sim N(\mu_{\alpha}, \sigma_{\alpha}^2)$$
As with the the “no-pooling” model, we set a separate intercept for each county, but rather than fitting separate least squares regression models for each county, multilevel modeling **shares strength** among counties, allowing for more reasonable inference in counties with little data.
```
with Model() as varying_intercept:
# Priors
μ_a = Normal('μ_a', mu=0., sigma=10)
σ_a = HalfCauchy('σ_a', 5)
# Random intercepts
a = Normal('a', mu=μ_a, sigma=σ_a, shape=n_counties)
# Common slope
b = Normal('b', mu=0., sigma=10)
# Model error
sd_y = HalfCauchy('sd_y', 5)
# Expected value
y_hat = a[county] + b * floor_measure
# Data likelihood
y_like = Normal('y_like', mu=y_hat, sigma=sd_y, observed=log_radon)
with varying_intercept:
varying_intercept_trace = sample(1000, tune=1000, cores=2, random_seed=RANDOM_SEED)
(ax,) = plot_forest(varying_intercept_trace, var_names=['a'])
ax.set_yticklabels([""]);
from arviz import plot_posterior
plot_posterior(varying_intercept_trace, var_names=['σ_a', 'b']);
```
The estimate for the `floor` coefficient is approximately -0.66, which can be interpreted as houses without basements having about half ($\exp(-0.66) = 0.52$) the radon levels of those with basements, after accounting for county.
```
from pymc3 import summary
summary(varying_intercept_trace, var_names=['b'])
xvals = np.arange(2)
bp = varying_intercept_trace[a].mean(axis=0)
mp = varying_intercept_trace[b].mean()
for bi in bp:
plt.plot(xvals, mp*xvals + bi, 'bo-', alpha=0.4)
plt.xlim(-0.1,1.1);
```
It is easy to show that the partial pooling model provides more objectively reasonable estimates than either the pooled or unpooled models, at least for counties with small sample sizes.
```
fig, axes = plt.subplots(2, 4, figsize=(12, 6), sharey=True, sharex=True)
axes = axes.ravel()
for i,c in enumerate(sample_counties):
# Plot county data
y = radon_data.log_radon[radon_data.county==c]
x = radon_data.floor[radon_data.county==c]
axes[i].scatter(x + np.random.randn(len(x))*0.01, y, alpha=0.4)
# No pooling model
b = unpooled_estimates[c]
m = unpooled_trace['β1'].mean()
xvals = np.linspace(-0.2, 1.2)
# Unpooled estimate
axes[i].plot(xvals, m*xvals+b)
# Pooled estimate
axes[i].plot(xvals, m0*xvals+b0, 'r--')
# Partial pooling esimate
axes[i].plot(xvals, mp*xvals+bp[county_lookup[c]], 'k:')
axes[i].set_xticks([0,1])
axes[i].set_xticklabels(['basement', 'floor'])
axes[i].set_ylim(-1, 3)
axes[i].set_title(c)
if not i%2:
axes[i].set_ylabel('log radon level');
```
## Varying slope model
Alternatively, we can posit a model that allows the counties to vary according to how the location of measurement (basement or floor) influences the radon reading.
$$y_i = \alpha + \beta_{j[i]} x_{i} + \epsilon_i$$
```
with Model() as varying_slope:
# Priors
μ_b = Normal('μ_b', mu=0., sigma=10)
σ_b = HalfCauchy('σ_b', 5)
# Common intercepts
a = Normal('a', mu=0., sigma=10)
# Random slopes
b = Normal('b', mu=μ_b, sigma=σ_b, shape=n_counties)
# Model error
σ_y = HalfCauchy('σ_y',5)
# Expected value
y_hat = a + b[county] * floor_measure
# Data likelihood
y_like = Normal('y_like', mu=y_hat, sigma=σ_y, observed=log_radon)
with varying_slope:
varying_slope_trace = sample(1000, tune=1000, cores=2, random_seed=RANDOM_SEED)
(ax,) = plot_forest(varying_slope_trace, var_names=['b'])
ax.set_yticklabels([""]);
xvals = np.arange(2)
b = varying_slope_trace['a'].mean()
m = varying_slope_trace['b'].mean(axis=0)
for mi in m:
plt.plot(xvals, mi*xvals + b, 'bo-', alpha=0.4)
plt.xlim(-0.2, 1.2);
```
## Exercise: Varying intercept and slope model
The most general model allows both the intercept and slope to vary by county:
$$y_i = \alpha_{j[i]} + \beta_{j[i]} x_{i} + \epsilon_i$$
Combine these two models to create a version with both slope and intercept varying.
```
with Model() as varying_intercept_slope:
# Write your model here
with varying_intercept_slope:
varying_intercept_slope_trace = sample(1000, tune=1000, cores=2, random_seed=RANDOM_SEED)
```
Forest plot of slopes and intercepts:
```
plot_forest(varying_intercept_slope_trace, var_names=['a','b']);
xvals = np.arange(2)
b = varying_intercept_slope_trace['a'].mean(axis=0)
m = varying_intercept_slope_trace['b'].mean(axis=0)
for bi,mi in zip(b,m):
plt.plot(xvals, mi*xvals + bi, 'bo-', alpha=0.4)
plt.xlim(-0.1, 1.1);
```
## Adding group-level predictors
A primary strength of multilevel models is the ability to handle predictors on multiple levels simultaneously. If we consider the varying-intercepts model above:
$$y_i = \alpha_{j[i]} + \beta x_{i} + \epsilon_i$$
we may, instead of a simple random effect to describe variation in the expected radon value, specify another regression model with a county-level covariate. Here, we use the county uranium reading $u_j$, which is thought to be related to radon levels:
$$\alpha_j = \gamma_0 + \gamma_1 u_j + \zeta_j$$
$$\zeta_j \sim N(0, \sigma_{\alpha}^2)$$
Thus, we are now incorporating a house-level predictor (floor or basement) as well as a county-level predictor (uranium).
Note that the model has both indicator variables for each county, plus a county-level covariate. In classical regression, this would result in collinearity. In a multilevel model, the partial pooling of the intercepts towards the expected value of the group-level linear model avoids this.
Group-level predictors also serve to reduce group-level variation $\sigma_{\alpha}$. An important implication of this is that the group-level estimate induces stronger pooling.
```
from pymc3 import Deterministic
with Model() as hierarchical_intercept:
# Priors
σ_a = HalfCauchy('σ_a', 5)
# County uranium model
γ_0 = Normal('γ_0', mu=0., sigma=10)
γ_1 = Normal('γ_1', mu=0., sigma=10)
# Uranium model for intercept
μ_a = γ_0 + γ_1*log_uranium
# County variation not explained by uranium
ϵ_a = Normal('ϵ_a', mu=0, sigma=1, shape=n_counties)
a = Deterministic('a', μ_a + σ_a*ϵ_a[county])
# Common slope
b = Normal('b', mu=0., sigma=10)
# Model error
σ_y = Uniform('σ_y', lower=0, upper=100)
# Expected value
y_hat = a + b * floor_measure
# Data likelihood
y_like = Normal('y_like', mu=y_hat, sigma=σ_y, observed=log_radon)
with hierarchical_intercept:
hierarchical_intercept_trace = sample(1000, tune=1000, cores=2, random_seed=RANDOM_SEED)
a_means = hierarchical_intercept_trace['a'].mean(axis=0)
plt.scatter(log_uranium, a_means)
g0 = hierarchical_intercept_trace['γ_0'].mean()
g1 = hierarchical_intercept_trace['γ_1'].mean()
xvals = np.linspace(-1, 0.8)
plt.plot(xvals, g0+g1*xvals, 'k--')
plt.xlim(-1, 0.8)
a_se = hierarchical_intercept_trace['a'].std(axis=0)
for ui, m, se in zip(log_uranium, a_means, a_se):
plt.plot([ui,ui], [m-se, m+se], 'b-')
plt.xlabel('County-level uranium'); plt.ylabel('Intercept estimate');
```
The standard errors on the intercepts are narrower than for the partial-pooling model without a county-level covariate.
### Correlations among levels
In some instances, having predictors at multiple levels can reveal correlation between individual-level variables and group residuals. We can account for this by including the average of the individual predictors as a covariate in the model for the group intercept.
$$\alpha_j = \gamma_0 + \gamma_1 u_j + \gamma_2 \bar{x} + \zeta_j$$
These are broadly referred to as ***contextual effects***.
```
# Create new variable for mean of floor across counties
xbar = radon_data.groupby('county')['floor'].mean().rename(county_lookup).values
with Model() as contextual_effect:
# Priors
σ_a = HalfCauchy('σ_a', 5)
# County uranium model for slope
γ = Normal('γ', mu=0., sigma=10, shape=3)
# Uranium model for intercept
μ_a = Deterministic('μ_a', γ[0] + γ[1]*log_uranium + γ[2]*xbar[county])
# County variation not explained by uranium
ϵ_a = Normal('ϵ_a', mu=0, sigma=1, shape=n_counties)
a = Deterministic('a', μ_a + σ_a*ϵ_a[county])
# Common slope
b = Normal('b', mu=0., sigma=10)
# Model error
σ_y = Uniform('σ_y', lower=0, upper=100)
# Expected value
y_hat = a + b * floor_measure
# Data likelihood
y_like = Normal('y_like', mu=y_hat, sigma=σ_y, observed=log_radon)
with contextual_effect:
contextual_effect_trace = sample(1000, tune=1000, cores=2, random_seed=RANDOM_SEED)
plot_forest(contextual_effect_trace, var_names=['γ']);
summary(contextual_effect_trace, var_names=['γ'])
```
So, we might infer from this that counties with higher proportions of houses without basements tend to have higher baseline levels of radon. Perhaps this is related to the soil type, which in turn might influence what type of structures are built.
### Prediction
Gelman (2006) used cross-validation tests to check the prediction error of the unpooled, pooled, and partially-pooled models
**root mean squared cross-validation prediction errors**:
* unpooled = 0.86
* pooled = 0.84
* multilevel = 0.79
There are two types of prediction that can be made in a multilevel model:
1. a new individual within an existing group
2. a new individual within a new group
For example, if we wanted to make a prediction for a new house with no basement in St. Louis county, we just need to sample from the radon model with the appropriate intercept.
```
county_lookup['ST LOUIS']
```
That is,
$$\tilde{y}_i \sim N(\alpha_{69} + \beta (x_i=1), \sigma_y^2)$$
This is a matter of adding a single additional line in PyMC:
```
with Model() as contextual_pred:
# Priors
σ_a = HalfCauchy('σ_a', 5)
# County uranium model for slope
γ = Normal('γ', mu=0., sigma=10, shape=3)
# Uranium model for intercept
μ_a = Deterministic('μ_a', γ[0] + γ[1]*log_uranium + γ[2]*xbar[county])
# County variation not explained by uranium
ϵ_a = Normal('ϵ_a', mu=0, sigma=1, shape=n_counties)
a = Deterministic('a', μ_a + σ_a*ϵ_a[county])
# Common slope
b = Normal('b', mu=0., sigma=10)
# Model error
σ_y = Uniform('σ_y', lower=0, upper=100)
# Expected value
y_hat = a + b * floor_measure
# Data likelihood
y_like = Normal('y_like', mu=y_hat, sigma=σ_y, observed=log_radon)
# St Louis county prediction
stl_pred = Normal('stl_pred', mu=a[69] + b, sigma=σ_y)
with contextual_pred:
contextual_pred_trace = sample(2000, tune=1000, cores=2, random_seed=RANDOM_SEED)
plot_posterior(contextual_pred_trace, var_names=['stl_pred']);
```
## Exercise
How would we make a prediction from a new county (*e.g.* one not included in this dataset)?
```
# Write your answer here
```
## Benefits of Multilevel Models
- Accounting for natural hierarchical structure of observational data
- Estimation of coefficients for (under-represented) groups
- Incorporating individual- and group-level information when estimating group-level coefficients
- Allowing for variation among individual-level coefficients across groups
---
## References
Gelman, A., & Hill, J. (2006). Data Analysis Using Regression and Multilevel/Hierarchical Models (1st ed.). Cambridge University Press.
Betancourt, M. J., & Girolami, M. (2013). Hamiltonian Monte Carlo for Hierarchical Models.
Gelman, A. (2006). Multilevel (Hierarchical) modeling: what it can and cannot do. Technometrics, 48(3), 432–435.
| github_jupyter |
## Lateral displacements
In this notebook we explore a few different things.
- We can compute mixing length scales as variance divided by gradient. This is a nice metric when we are thinking about how a gradient gets mixed.
- We can also potentially compute a subduction length scale as the nearest mean upwelling region that had similar properties. Maybe this can be done using spice.
Combining these length scales, with a time scale can help.
Timescale can also be computed in multiple ways:
- Either using some estimate of EKE along with a length scale
- Or getting a ventilation timescale using the AOU.
```
import numpy as np
import xarray as xr
import gsw
import matplotlib.pyplot as plt
data_dir = '/Users/dhruvbalwada/OneDrive/sogos_data/data/'
# Climatology data
Tclim = xr.open_dataset(data_dir + "raw/climatology/RG_ArgoClim_Temperature_2017.nc", decode_times=False)
Sclim = xr.open_dataset(data_dir + "raw/climatology/RG_ArgoClim_Salinity_2017.nc", decode_times=False)
Climextra = xr.open_mfdataset(data_dir+ 'raw/climatology/RG_ArgoClim_201*', decode_times=False)
RG_clim = xr.merge([Tclim, Sclim, Climextra])
# Calendar type was missing, and giving errors in decoding time
RG_clim.TIME.attrs['calendar'] = '360_day'
RG_clim = xr.decode_cf(RG_clim);
## Add density and other things
SA = xr.apply_ufunc(gsw.SA_from_SP, RG_clim.ARGO_SALINITY_MEAN+RG_clim.ARGO_SALINITY_ANOMALY, RG_clim.PRESSURE ,
RG_clim.LONGITUDE, RG_clim.LATITUDE,
dask='parallelized', output_dtypes=[float,]).rename('SA')
CT = xr.apply_ufunc(gsw.CT_from_t, SA, RG_clim.ARGO_TEMPERATURE_MEAN+RG_clim.ARGO_TEMPERATURE_ANOMALY, RG_clim.PRESSURE,
dask='parallelized', output_dtypes=[float,]).rename('CT')
SIGMA0 = xr.apply_ufunc(gsw.sigma0, SA, CT, dask='parallelized', output_dtypes=[float,]).rename('SIGMA0')
RG_clim = xr.merge([RG_clim, SA, CT, SIGMA0])
CT_region = RG_clim.CT.sel(LONGITUDE=30, method='nearest').sel(
LATITUDE=slice(-65,-30), PRESSURE=slice(0,1200)
).groupby('TIME.month').mean().load()
SA_region = RG_clim.SA.sel(LONGITUDE=30, method='nearest').sel(
LATITUDE=slice(-65,-30), PRESSURE=slice(0,1200)
).groupby('TIME.month').mean().load()
rho_region = RG_clim.SIGMA0.sel(LONGITUDE=30, method='nearest').sel(
LATITUDE=slice(-65,-30), PRESSURE=slice(0,1200)
).groupby('TIME.month').mean().load()
Clim_region = xr.merge([CT_region, SA_region, rho_region])
Clim_region
Clim_region.CT.isel(month=0).plot.contourf(levels=19, vmin=-17)
Clim_region.SIGMA0.isel(month=0).plot.contour()
plt.gca().invert_yaxis()
```
| github_jupyter |
# Wiki Networking - Extended Example
### Introduction
Network graphs consisting of nodes and edges and be used to visualize the relationships between people. The results can inform a viewer of groups of relationships. This notebook has examples for all functions available and a variety of text mining examples. It demonstrates how to crawl Wiki articles and represent the links between these articles as a graph.
### `import` statements
First, we will import a few necessary packages, including `wikinetworking` from this repository. It includes functions for crawling and text mining Wiki data as well as graphing the result of the crawl.
```
import wikinetworking as wn
import networkx as nx
from pyquery import PyQuery
%matplotlib inline
print "OK"
```
### `filter_links`
This function accepts a PyQuery object and returns a list of Wikipedia article links from the article's main body text. It will not return links that are redirects or links that are anchors within other pages. Optionally, you can specify a DOM selector for the type of element containing links you wish to retrieve.
In this example, we create a PyQuery object for the [Iron Man](https://en.wikipedia.org/wiki/Iron_Man) article on Wikipedia, retrieve its links with `filter_links` and output the number of links we retrieved.
```
iron_man_page = PyQuery(url="https://en.wikipedia.org/wiki/Iron_man")
iron_man_links = wn.filter_links(iron_man_page)
print len(iron_man_links), "links retrieved"
```
In this example, we retrieve links from a [list of Hip Hop musicians](https://en.wikipedia.org/wiki/List_of_hip_hop_musicians). Notice that each link is contained inside of a `li` list item HTML tag.
```
hip_hop_page = PyQuery(url="https://en.wikipedia.org/wiki/List_of_hip_hop_musicians")
hip_hop_links = wn.filter_links(hip_hop_page, "li")
print len(hip_hop_links), "links retrieved"
```
In this example, we do the same thing for [Harry Potter Characters](https://en.wikipedia.org/wiki/List_of_Harry_Potter_characters). Notice that many links are omitted - these links are redirects or links to sections within larger articles.
```
harry_potter_page = PyQuery(url="https://en.wikipedia.org/wiki/List_of_Harry_Potter_characters")
harry_potter_links = wn.filter_links(harry_potter_page, "li")
print len(harry_potter_links), "links retrieved"
```
In this example, we are retrieving a list of [Marvel Comics Characters beginning with the letter D](https://en.wikipedia.org/wiki/List_of_Marvel_Comics_characters:_D). In this case, we specify the DOM selector `.hatnote`. This is because we only want links for Marvel characters that have a dedicated _*Main article*_. The _*Main article*_ elements all have the `.hatnote` CSS class, so it is an easy way of selecting these articles. In this example, we simply print the contents of the list.
```
marvel_d_page = PyQuery(url="https://en.wikipedia.org/wiki/List_of_Marvel_Comics_characters:_D")
marvel_d_links = wn.filter_links(marvel_d_page, ".hatnote")
print marvel_d_links
```
### `retrieve_multipage`
This function retrieves data from a list of URLs. This is useful for when data on a Wikipedia category is broken up into multiple individual pages.
In this example, we are retrieving lists from all subpages from the [List of Marvel Comics Characters](https://en.wikipedia.org/wiki/List_of_Marvel_Comics_characters). The URL pattern for these articles is `https://en.wikipedia.org/wiki/List_of_Marvel_Comics_characters:_` followed by a section name.
```
sections = [letter for letter in 'ABCDEFGHIJKLMNOPQRSTUVWXYZ']
sections.append('0-9')
urls = ["https://en.wikipedia.org/wiki/List_of_Marvel_Comics_characters:_" + section for section in sections]
print "URLs:", urls
character_links = wn.retrieve_multipage(urls, ".hatnote", True)
print len(character_links), "links retrieved"
```
### `write_list` and `read_list`
These are convenience functions for writing and reading list data.
In this example, we write all of the Marvel character links we retrieved to a text file, and verify that we can read the data from the file.
```
wn.write_list(character_links, "all_marvel_chars.txt")
print len(wn.read_list("all_marvel_chars.txt")), "read"
```
### `intersection`
Returns a list of elements that appear in both provided lists
The `intersection` function is useful for cross referencing links from one Wikipedia page with another. For example, the [List of Hip Hop Musicians](https://en.wikipedia.org/wiki/List_of_hip_hop_musicians) contains many musicians, but we may not have the time or resources to crawl every single artist. The [BET Hip Hop Awards page](https://en.wikipedia.org/wiki/BET_Hip_Hop_Awards) has links to artists that have won an award and may be significant for our search, but it also contains links to songs and other media that may not be articles about individual Hip Hop artists. In addition, each Wiki page will have links to completely unrelated articles. By taking the `intersection` of both, we get a list containing links of only Hip Hop arists that have won a BET Hip Hop Award.
```
hip_hop_page = PyQuery(url="https://en.wikipedia.org/wiki/List_of_hip_hop_musicians")
hip_hop_links = wn.filter_links(hip_hop_page, "li")
print len(hip_hop_links), "links retrieved from List of Hip Hop Musicians"
bet_hip_hop_awards_page = PyQuery(url="https://en.wikipedia.org/wiki/BET_Hip_Hop_Awards")
bet_hip_hop_awards_links = wn.filter_links(bet_hip_hop_awards_page, "li")
print len(bet_hip_hop_awards_links), "links retrieved from BET Hip Hop Awards"
award_winner_links = wn.intersection(hip_hop_links, bet_hip_hop_awards_links)
print "BET Hip Hop Award winners:", award_winner_links
```
### `crawl`
This crawls articles iteratively, with a two second delay between each page retrieval. It requires a starting URL fragment and an `accept` list of URLs the crawler should follow. It returns a dictionary structure with each URL fragment as a key and crawl data (`links`, `title` and `depth`) as a value.
In this example, we start a crawl at the article for [Jadakiss](https://en.wikipedia.org/wiki/Jadakiss). As the `accept` list of URLs that the crawler is allowed to follow, we will use the `award_winner_links` list from earlier. This isolates the crawl to only [Artist winners of BET Hip Hop Awards](https://en.wikipedia.org/wiki/BET_Hip_Hop_Awards). Therefore, our social network will be a network built of BET Hip Hop Award Winners and stored in the `bet_winner_crawl` dictionary.
```
bet_winner_crawl = wn.crawl("/wiki/Jadakiss", accept=award_winner_links)
print bet_winner_crawl
```
### `directed_graph` and `undirected_graph`
These functions flatten the raw crawl data into something a little more usable. It returns a dictionary with only article title names as the key, and corresponding urls and a dictionary of edges and weights to each article. The `directed_graph` function allows each article to retain a separate edge representing back links, while the `undirected_graph` creates one edge from each pair of article where the weight is a sum of the links between each article. For example, `directed_graph` might produce:
```
{
'Iron Man' : {
'url' : '/wiki/Iron_Man',
'edges' : {
'Captain America' : 11
}
},
'Captain America' : {
'url' : '/wiki/Captain_America',
'edges' : {
'Iron Man' : 8
}
}
}
```
...while the same data passed to `undirected_graph` might produce:
```
{
'Iron Man' : {
'url' : '/wiki/Iron_Man',
'edges' : {
'Captain America' : 19
}
},
'Captain America' : {
'url' : /wiki/Captain_America',
'edges' : {
}
}
}
```
```
bet_directed_graph_data = wn.directed_graph(bet_winner_crawl)
print bet_directed_graph_data
bet_undirected_graph_data = wn.undirected_graph(bet_winner_crawl)
print bet_undirected_graph_data
```
### `save_dict` and `load_dict`
These functions save and load Python dictionaries. This is convenient when you want to save crawl data.
```
wn.save_dict(bet_undirected_graph_data, "bet_network.json")
print wn.load_dict("bet_network.json")
```
### `create_graph`
Now that we have a dictionary representing an undirected graph, we need to turn it into a `networkx.graph` object which can be drawn. Optionally, we can pre-generate a node layout and save it.
```
# Just in case we don't want to re-run the crawl, we will load the data directly
import wikinetworking as wn
import networkx as nx
%matplotlib inline
bet_undirected_graph_data = wn.load_dict("bet_network.json")
# Now create the graph
graph = wn.create_graph(bet_undirected_graph_data)
layout = nx.spring_layout(graph)
```
### `make_interactive_graph`
The `networkx.graph` object can be rendered as an interactive, clickable graph.
```
graph_html = wn.make_interactive_graph(graph, pos=layout)
```
You can save the resulting HTML of the interactive graph to a file, if you wish to load it outside of this notebook.
```
with open("bet_network.html", "w") as f:
f.write(graph_html)
f.close()
```
### `save_big_graph`
If you would like to save a very high resolution version of this graph for use on a display system like [Texas Advanced Computing Center's Stallion](https://www.tacc.utexas.edu/vislab/stallion), you can use the `save_big_graph` function. The default output size at 600dpi will be 4800x3600 pixels. Warning: This function can take a little while to run.
```
wn.save_big_graph(graph, pos=layout)
```
| github_jupyter |
# XGBoost model for prediction of avalanche events
```
# import of standard Python libraries for data analysis
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
# installation of XGBoost
!pip install xgboost
# import needed objects from Scikit learn library for machine learning
from numpy import loadtxt
from xgboost import XGBClassifier
from sklearn.metrics import confusion_matrix, classification_report
from sklearn import preprocessing
from sklearn import metrics
from sklearn.metrics import f1_score, accuracy_score
from sklearn.model_selection import train_test_split, cross_val_score
from numpy import mean
from sklearn.model_selection import RepeatedStratifiedKFold
# I would like to see all rows and columns of dataframes
pd.set_option('display.max_columns', None)
pd.set_option('display.max_rows', None)
```
## Reading dataset and final preprocessing
```
# reading dataset
df = pd.read_csv("final_data.csv")
df.head()
# viewing all variables to decide which are not useful
df.columns
# removing variables not needed for the model
df_clean = df.drop(columns=["massif_num","lon","lat","aval_type", "acccidental_risk_index",
'snow_thickness_1D', 'snow_thickness_3D', 'snow_thickness_5D',
'snow_water_1D', 'snow_water_3D', 'snow_water_5D', 'risk_index',
'thickness_of_wet_snow_top_of_snowpack','thickness_of_frozen_snow_top_of_snowpack',
'surface_air_pressure_mean', 'rainfall_rate', 'drainage', 'runoff',
'liquid_water_in_soil', 'frozen_water_in_soil', 'elevation','snow_melting_rate', 'aval_accident'])
# viewing if data type of the variables is suitable for XGBoost
# there are 2 string categorical variables that needs to be transformed: day and massif name
df_clean.info()
df_clean.shape
# for transformation of massif names I will use OneHotEncoding with pd.get_dummies method
# selecting of values for dummies
df_clean.massif_name.unique()
massifs = ('Chablais', 'Aravis', 'Mont-Blanc', 'Bauges', 'Beaufortin',
'Hte-tarent', 'Chartreuse', 'Belledonne', 'Maurienne', 'Vanoise',
'Hte-maurie', 'Gdes-rouss', 'Thabor', 'Vercors', 'Oisans',
'Pelvoux', 'Queyras', 'Devoluy', 'Champsaur', 'Parpaillon',
'Ubaye', 'Ht_Var-Ver', 'Mercantour')
# creating initial dataframe
df_massifs = pd.DataFrame(massifs, columns=['massif_name'])
# generate binary values using get_dummies
dum_df = pd.get_dummies(df_massifs, columns=["massif_name"], prefix="massif")
# merge initial dataframe with dummies
df_massifs = df_massifs.join(dum_df)
# merge final datatset with dataframe with dummies
df_clean = df_clean.merge(df_massifs, how="left", on="massif_name")
# checking dataset with dummy variables
df_clean.head()
# getting rid of redundant variable
df_clean = df_clean.drop(columns=['massif_name'])
# second categorical variable to transform is day
# slicing is used to get years and months from day variable
# and after we need to transform years and months to integers
df_clean["year"] = (df_clean.day.str[:4]).astype(int)
df_clean["month"] = (df_clean.day.str[5:7]).astype(int)
df_clean.head()
# getting rid of redundant variable
df_clean = df_clean.drop(columns=['day', 'year'])
# checking dataset with new time variables
df_clean.head()
# verifying there are no null values in dataset
(df_clean.apply(lambda x: x.isnull().sum())).sum()
# checking one last time my dataset before creating RF models
df_clean.info()
```
## Problem with very imbalanced dataset
- metrics like Accuracy will be very high, but that does not provide any significant insight, because of imbalanced dataset with majority of No avalanche events
- in classification report metrics **Precision will have lesser importance for my analysis than Recall**, because I want to reduce number of False Negatives rather then number of False Positives
- **For performance of XGBoost model, I will look mainly on weighted F1-score** metrics because it is the most suitable for this kind of analysis. F1-score is in fact weighted average of the precision and recall.
```
# Percentage of cases with and without avalanches showing imbalanced dataset
round((df_clean.aval_event.value_counts()/540818)*100, 2)
# getting number of positives and negatives in order to calculate scale_pos_weight
total_negative = df_clean.aval_event.value_counts()[0]
total_positive = df_clean.aval_event.value_counts()[1]
f"Number of total positives is {total_positive} and for total negatives it is {total_negative}"
# scale_pos_weight for class weights
scale_pos_weight=total_negative/total_positive
scale_pos_weight
# labels are the values we want to predict
labels_ev = np.array(df_clean['aval_event'])
# removing the labels from the features
features_ev = df_clean.drop(columns=['aval_event'])
# saving feature names for later use
feature_list_ev = list(features_ev.columns)
# converting to numpy array
features_ev = np.array(features_ev)
# splitting dataset into train and test
train_features_ev, test_features_ev, train_labels_ev, test_labels_ev = train_test_split(features_ev, labels_ev, test_size = 0.33, random_state = 42)
# displaying sizes of train/test features and labels
print('Training Features Shape:', train_features_ev.shape)
print('Training Labels Shape:', train_labels_ev.shape)
print('Testing Features Shape:', test_features_ev.shape)
print('Testing Labels Shape:', test_labels_ev.shape)
# defining and fitting model
xgb = XGBClassifier(scale_pos_weight=scale_pos_weight)
xgb.fit(train_features_ev, train_labels_ev)
# making prediction
label_pred_ev = xgb.predict(test_features_ev)
%%time
# defining evaluation procedure
cv = RepeatedStratifiedKFold(n_splits=10, n_repeats=3, random_state=1)
# evaluating model
scores = cross_val_score(xgb, features_ev, labels_ev, scoring='roc_auc', cv=cv, n_jobs=-1)
# summary of performance
print('Mean ROC AUC: %.5f' % mean(scores))
print("Accuracy:",round(metrics.accuracy_score(test_labels_ev, label_pred_ev),5))
print("F1_score weighted:", round(f1_score(test_labels_ev, label_pred_ev, average='weighted'),5))
# displaying confusion matrix
sns.set(font_scale=1.75)
f, ax = plt.subplots(figsize=(5, 5))
cf_matrix_basic = confusion_matrix(test_labels_ev, label_pred_ev)
sns.heatmap(cf_matrix_basic, annot=True, ax=ax, cmap="YlGnBu", fmt=".0f", linewidths=.5)
plt.title("Confusion matrix for avalanches")
plt.xlabel('Predicted')
plt.ylabel('Real');
# displaying report of performance of model
print(classification_report(test_labels_ev, label_pred_ev, digits=3))
```
## Results without summer months
```
# creating dataframe without summer months
summer_season = [6, 7, 8]
df_winter = df_clean[~df_clean["month"].isin(summer_season)]
df_winter.head()
# labels are the values we want to predict
labels_win = np.array(df_winter['aval_event'])
# removing the labels from the features
features_win = df_winter.drop(columns=['aval_event'])
# saving feature names for later use
feature_list_win = list(features_win.columns)
# converting to numpy array
features_win = np.array(features_win)
# splitting dataset into train and test
train_features_win, test_features_win, train_labels_win, test_labels_win = train_test_split(features_win, labels_win, test_size = 0.33, random_state = 42)
# displaying sizes of train/test features and labels
print('Training Features Shape:', train_features_win.shape)
print('Training Labels Shape:', train_labels_win.shape)
print('Testing Features Shape:', test_features_win.shape)
print('Testing Labels Shape:', test_labels_win.shape)
# getting number of positives and negatives in order to calculate scale_pos_weight
total_negative_win = df_winter.aval_event.value_counts()[0]
total_positive_win = df_winter.aval_event.value_counts()[1]
f"Number of total positives is {total_positive_win} and for total negatives it is {total_negative_win}"
# scale_pos_weight for class weights
scale_pos_weight_win=total_negative_win/total_positive_win
scale_pos_weight_win
%%time
# defining and fitting model
xgb_win = XGBClassifier(scale_pos_weight=scale_pos_weight_win)
xgb_win.fit(train_features_win, train_labels_win)
# making prediction
label_pred_win = xgb_win.predict(test_features_win)
%%time
# evaluate model
scores = cross_val_score(xgb_win, features_win, labels_win, scoring='roc_auc', cv=cv, n_jobs=-1)
# summary of performance
print('Mean ROC AUC: %.5f' % mean(scores))
print("Accuracy:",round(metrics.accuracy_score(test_labels_win, label_pred_win),5))
print("F1_score weighted:", round(f1_score(test_labels_win, label_pred_win, average='weighted'),5))
cf_matrix_win = confusion_matrix(test_labels_win, label_pred_win)
f, ax = plt.subplots(figsize=(5, 5))
sns.heatmap(cf_matrix_win, annot=True, ax=ax, cmap="YlGnBu", fmt=".0f", linewidths=.5)
plt.title("Confusion matrix for avalanches without summer months")
plt.xlabel('Predicted')
plt.ylabel('Real');
# displaying report of performance of model
print(classification_report(test_labels_win, label_pred_win, digits=3))
```
## Results for altitude from 1500 to 3600 metres
```
df.elevation.value_counts()
df_elevation = df.drop(columns=["massif_num","lon","lat","aval_type", "acccidental_risk_index",
'snow_thickness_1D', 'snow_thickness_3D', 'snow_thickness_5D',
'snow_water_1D', 'snow_water_3D', 'snow_water_5D', 'risk_index',
'thickness_of_wet_snow_top_of_snowpack','thickness_of_frozen_snow_top_of_snowpack',
'surface_air_pressure_mean', 'rainfall_rate', 'drainage', 'runoff',
'liquid_water_in_soil', 'frozen_water_in_soil', 'snow_melting_rate', 'aval_accident'])
# creating initial dataframe
df_massifs = pd.DataFrame(massifs, columns=['massif_name'])
# generate binary values using get_dummies
dum_df = pd.get_dummies(df_massifs, columns=["massif_name"], prefix="massif")
# merge initial dataframe with dummies
df_massifs = df_massifs.join(dum_df)
# merge final datatset with dataframe with dummies
df_elevation = df_elevation.merge(df_massifs, how="left", on="massif_name")
# getting rid of redundant variable
df_elevation = df_elevation.drop(columns=['massif_name'])
# creating variable for months
df_elevation["month"] = (df_elevation.day.str[5:7]).astype(int)
df_elevation = df_elevation.drop(columns=['day'])
# verifying there are no null values in dataset
(df_elevation.apply(lambda x: x.isnull().sum())).sum()
# reducing altitudes where avalanches rarely occur
df_elevation = df_elevation[(df_elevation["elevation"]>1499) & (df_elevation["elevation"]< 3601)]
# last check of dataframe before using RF model
df_elevation.info()
# labels are the values we want to predict
labels_el = np.array(df_elevation['aval_event'])
# removing the labels from the features
features_el = df_elevation.drop(columns=['aval_event'])
# saving feature names for later use
feature_list_el = list(features_el.columns)
# converting to numpy array
features_el = np.array(features_el)
# splitting dataset into train and test
train_features_el, test_features_el, train_labels_el, test_labels_el = train_test_split(features_el, labels_el, test_size = 0.33, random_state = 42)
# displaying sizes of train/test features and labels
print('Training Features Shape:', train_features_el.shape)
print('Training Labels Shape:', train_labels_el.shape)
print('Testing Features Shape:', test_features_el.shape)
print('Testing Labels Shape:', test_labels_el.shape)
# getting number of positives and negatives in order to calculate scale_pos_weight
total_negative_el = df_elevation.aval_event.value_counts()[0]
total_positive_el = df_elevation.aval_event.value_counts()[1]
f"Number of total positives is {total_positive_el} and for total negatives it is {total_negative_el}"
# scale_pos_weight for class weights
scale_pos_weight_el =total_negative_el/total_positive_el
scale_pos_weight_el
%%time
# defining and fitting model
xgb_el = XGBClassifier(scale_pos_weight=scale_pos_weight_el)
xgb_el.fit(train_features_el, train_labels_el)
# making prediction
label_pred_el = xgb_el.predict(test_features_el)
%%time
# evaluate model
scores = cross_val_score(xgb_el, features_el, labels_el, scoring='roc_auc', cv=cv, n_jobs=-1)
# summary of performance
print('Mean ROC AUC: %.5f' % mean(scores))
print("Accuracy:",round(metrics.accuracy_score(test_labels_el, label_pred_el),5))
print("F1_score weighted:", round(f1_score(test_labels_el, label_pred_el, average='weighted'),5))
cf_matrix_el = confusion_matrix(test_labels_el, label_pred_el)
f, ax = plt.subplots(figsize=(5, 5))
sns.heatmap(cf_matrix_el, annot=True, ax=ax, cmap="YlGnBu", fmt=".0f", linewidths=.5)
plt.title("Confusion matrix for avalanches for altitude 1500 - 3600 m.")
plt.xlabel('Predicted')
plt.ylabel('Real');
# displaying report of performance of model
print(classification_report(test_labels_el, label_pred_el, digits=3))
```
## Results for limited altitude and without summer
```
# creating new dataframe for altitude from 1500 to 3600 and with exclusion of summer months
df_elev_win = df_elevation[~df_elevation["month"].isin(summer_season)]
# labels are the values we want to predict
labels_el_w = np.array(df_elev_win['aval_event'])
# removing the labels from the features
features_el_w = df_elev_win.drop(columns=['aval_event'])
# saving feature names for later use
feature_list_el_w = list(features_el_w.columns)
# converting to numpy array
features_el_w = np.array(features_el_w)
# splitting dataset into train and test
train_features_el_w, test_features_el_w, train_labels_el_w, test_labels_el_w = train_test_split(features_el_w, labels_el_w, test_size = 0.33, random_state = 42)
# displaying sizes of train/test features and labels
print('Training Features Shape:', train_features_el_w.shape)
print('Training Labels Shape:', train_labels_el_w.shape)
print('Testing Features Shape:', test_features_el_w.shape)
print('Testing Labels Shape:', test_labels_el_w.shape)
# getting number of positives and negatives in order to calculate scale_pos_weight
total_negative_el_w = df_elev_win.aval_event.value_counts()[0]
total_positive_el_w = df_elev_win.aval_event.value_counts()[1]
f"Number of total positives is {total_positive_el_w} and for total negatives it is {total_negative_el_w}"
# scale_pos_weight for class weights
scale_pos_weight_el_w =total_negative_el_w/total_positive_el_w
scale_pos_weight_el_w
%%time
# defining and fitting model
xgb_el_w = XGBClassifier(scale_pos_weight=scale_pos_weight_el_w)
xgb_el_w.fit(train_features_el_w, train_labels_el_w)
# making prediction
label_pred_el_w = xgb_el_w.predict(test_features_el_w)
%%time
# evaluate model
scores = cross_val_score(xgb_el_w, features_el_w, labels_el_w, scoring='roc_auc', cv=cv, n_jobs=-1)
# summary of performance
print('Mean ROC AUC: %.5f' % mean(scores))
print("Accuracy:",round(metrics.accuracy_score(test_labels_el_w, label_pred_el_w),5))
print("F1_score weighted:", round(f1_score(test_labels_el_w, label_pred_el_w, average='weighted'),5))
cf_matrix_el_w = confusion_matrix(test_labels_el_w, label_pred_el_w)
f, ax = plt.subplots(figsize=(5, 5))
sns.heatmap(cf_matrix_el_w, annot=True, ax=ax, cmap="YlGnBu", fmt=".0f", linewidths=.5)
plt.title("Confusion matrix for avalanches for altitude 1500 - 3600 m. without summer")
plt.xlabel('Predicted')
plt.ylabel('Real');
# displaying report of performance of model
print(classification_report(test_labels_el_w, label_pred_el_w, digits=3))
```
## Results for French Alps (without massifs)
```
df_clean.columns
# removing variables with massifs
df_no_massifs = df_clean[['temp_soil_0.005_m', 'temp_soil_0.08_m', 'whiteness_albedo',
'net_radiation', 'surface_temperature', 'surface_snow_amount',
'thickness_of_snowfall', 'snow_thickness_7D', 'snow_water_7D',
'penetration_ram_resistance', 'near_surface_humidity_mean',
'relative_humidity_mean', 'freezing_level_altitude_mean',
'rain_snow_transition_altitude_mean', 'air_temp_max', 'wind_speed_max',
'snowfall_rate_max', 'nebulosity_max', 'air_temp_min', 'aval_event', 'month']]
# checking new dataframe
df_no_massifs.info()
# labels are the values we want to predict
labels_alps = np.array(df_no_massifs['aval_event'])
# removing the labels from the features
features_alps = df_no_massifs.drop(columns=['aval_event'])
# saving feature names for later use
feature_list_alps = list(features_alps.columns)
# converting to numpy array
features_alps = np.array(features_alps)
# splitting dataset into train and test
train_features_alps, test_features_alps, train_labels_alps, test_labels_alps = train_test_split(features_alps, labels_alps, test_size = 0.33, random_state = 42)
# displaying sizes of train/test features and labels
print('Training Features Shape:', train_features_alps.shape)
print('Training Labels Shape:', train_labels_alps.shape)
print('Testing Features Shape:', test_features_alps.shape)
print('Testing Labels Shape:', test_labels_alps.shape)
# getting number of positives and negatives in order to calculate scale_pos_weight
total_negative_alps = df_no_massifs.aval_event.value_counts()[0]
total_positive_alps = df_no_massifs.aval_event.value_counts()[1]
f"Number of total positives is {total_positive_alps} and for total negatives it is {total_negative_alps}"
# scale_pos_weight for class weights
scale_pos_weight_alps =total_negative_alps/total_positive_alps
scale_pos_weight_alps
%%time
# defining and fitting model
xgb_alps = XGBClassifier(scale_pos_weight=scale_pos_weight_alps)
xgb_alps.fit(train_features_alps, train_labels_alps)
# making prediction
label_pred_alps = xgb_alps.predict(test_features_alps)
%%time
# evaluate model
scores = cross_val_score(xgb_alps, features_alps, labels_alps, scoring='roc_auc', cv=cv, n_jobs=-1)
# summary of performance
print('Mean ROC AUC: %.5f' % mean(scores))
print("Accuracy:",round(metrics.accuracy_score(test_labels_alps, label_pred_alps),5))
print("F1_score weighted:", round(f1_score(test_labels_alps, label_pred_alps, average='weighted'),5))
cf_matrix_alps = confusion_matrix(test_labels_alps, label_pred_alps)
f, ax = plt.subplots(figsize=(5, 5))
sns.heatmap(cf_matrix_alps, annot=True, ax=ax, cmap="YlGnBu", fmt=".0f", linewidths=.5)
plt.title("Confusion matrix for avalanches in French Alps")
plt.xlabel('Predicted')
plt.ylabel('Real');
# displaying report of performance of model
print(classification_report(test_labels_alps, label_pred_alps, digits=3))
```
## Results for French Alps without summer
```
df_alps_win = df_no_massifs[~df_no_massifs["month"].isin(summer_season)]
# labels are the values we want to predict
labels_alps_win = np.array(df_alps_win['aval_event'])
# removing the labels from the features
features_alps_win = df_alps_win.drop(columns=['aval_event'])
# saving feature names for later use
feature_list_alps_win = list(features_alps_win.columns)
# converting to numpy array
features_alps_win = np.array(features_alps_win)
# splitting dataset into train and test
train_features_alps_win, test_features_alps_win, train_labels_alps_win, test_labels_alps_win = train_test_split(features_alps_win, labels_alps_win, test_size = 0.33, random_state = 42)
# displaying sizes of train/test features and labels
print('Training Features Shape:', train_features_alps_win.shape)
print('Training Labels Shape:', train_labels_alps_win.shape)
print('Testing Features Shape:', test_features_alps_win.shape)
print('Testing Labels Shape:', test_labels_alps_win.shape)
# getting number of positives and negatives in order to calculate scale_pos_weight
total_negative_alps_win = df_alps_win.aval_event.value_counts()[0]
total_positive_alps_win = df_alps_win.aval_event.value_counts()[1]
f"Number of total positives is {total_positive_alps_win} and for total negatives it is {total_negative_alps_win}"
# scale_pos_weight for class weights
scale_pos_weight_alps_win =total_negative_alps_win/total_positive_alps_win
scale_pos_weight_alps_win
%%time
# defining and fitting model
xgb_alps_win = XGBClassifier(scale_pos_weight=scale_pos_weight_alps_win)
xgb_alps_win.fit(train_features_alps_win, train_labels_alps_win)
# making prediction
label_pred_alps_win = xgb_alps_win.predict(test_features_alps_win)
%%time
# evaluate model
scores = cross_val_score(xgb_alps_win, features_alps_win, labels_alps_win, scoring='roc_auc', cv=cv, n_jobs=-1)
# summary of performance
print('Mean ROC AUC: %.5f' % mean(scores))
print("Accuracy:",round(metrics.accuracy_score(test_labels_alps_win, label_pred_alps_win),5))
print("F1_score weighted:", round(f1_score(test_labels_alps_win, label_pred_alps_win, average='weighted'),5))
cf_matrix_alps_win = confusion_matrix(test_labels_alps_win, label_pred_alps_win)
f, ax = plt.subplots(figsize=(5, 5))
sns.heatmap(cf_matrix_alps_win, annot=True, ax=ax, cmap="YlGnBu", fmt=".0f", linewidths=.5)
plt.title("Confusion matrix for avalanches in French Alps without summer")
plt.xlabel('Predicted')
plt.ylabel('Real');
# displaying report of performance of model
print(classification_report(test_labels_alps_win, label_pred_alps_win, digits=3))
```
## Summary of XGBoost results:
Recall for days with avalanche (in order to reduce number of false negatives) was my primary metrics to evaluate performance of avalanche prediction, with weighted F1 score following closely behind. **I used 5 variants of feature selection in order to improve recall. Original sample without feature selection resulted in recall 0.887 for days with avalanches and weighted F1-score 0.99443.** Precision and recall for days without avalanches were in all 6 variants 1 or very close to 1, therefore this metrics won't be discussed further.
- **Feature selection**: I used 5 different changes to features:
A) sample without summer months
B) sample with limited altitude (not more than 3600 metres)
C) sample without massif division.
Also I checked combinations of A+B and A+C.
The worst results were from option C) and A+C). F1-score and recall for avalanche days has dropped. On the other hand, **the best recall for avalanche days had option A) (0.916), therefore this would be my preferred XGBoost model.** Even though other options B) and B+A) had slightly better F1-score (0.995 instead of 0.994 for option A) still I prefer A) option. Lower F1-score was consequence of slightly lower recall for days without any avalanche, but this metrics has lesser importance.
All the models generated similar F1-scores 0.992–0.995. **Sample without summer months had 0.916 recall for avalanche days, therefore would be the most prefered option**.
-**Comparision with Random Forest model**:
Previously, I tried Random Forest ML model, which had slightly better weighted F1-scores (0.997-0.998), but recall for days with avalanche was significatly worse, only 0.58 at best, and therefore XGBoost machine learning models are better option than Random Forest.
**Conclusion: I would choose XGBoost on sample without summer months, because it has the best recall for avalanche days 0.916 and acceptable F1 score.** When comparing with results from Random Forest, XGBoost proved to be better choice, because of significant improvement in recall for avavlanche days, even though overall F1-score was little bit lower for all XGBoost models. But this was because of lower metrics like recall for days without avalanche or precision for avalanche days, which are not that important.
| github_jupyter |
<img src="../img/saturn_logo.png" width="300" />
We don't need to run all of Notebook 5 again, we'll just call `setup2.py` in the next chunk to get ourselves back to the right state. This also includes the reindexing work from Notebook 5, and a couple of visualization functions that we'll talk about later.
***
**Note: This notebook assumes you have an S3 bucket where you can store your model performance statistics.**
If you don't have access to an S3 bucket, but would still like to train your model and review results, please visit [Notebook 6b](06b-transfer-training-local.ipynb) and [Notebook 7](07-learning-results.ipynb) to see detailed examples of how you can do that.
***
## Connect to Cluster
```
%run -i ../tools/setup2.py
display(HTML(gpu_links))
import torch
from tensorboardX import SummaryWriter
from torch import nn, optim
from torch.nn.parallel import DistributedDataParallel as DDP
from torchvision import datasets, transforms, models
from torch.utils.data import DataLoader
from torch.utils.data.sampler import SubsetRandomSampler
import torch.distributed as dist
client
```
We're ready to do some learning!
## Model Parameters
Aside from the Special Elements noted below, we can write this section essentially the same way we write any other PyTorch training loop.
* Cross Entropy Loss for our loss function
* SGD (Stochastic Gradient Descent) for our optimizer
We have two stages in this process, as well - training and evaluation. We run the training set completely using batches of 100 before we move to the evaluation step, where we run the eval set completely also using batches of 100.
Most of the training workflow function shown will be very familiar for users of PyTorch. However, there are a couple of elements that are different.
### 1. Tensorboard Writer
We're using Tensorboard to monitor the model's performance, so we'll create a SummaryWriter object in our training function, and use that to write out statistics and sample image classifications.
### 2. Model to GPU Resources
```
device = torch.device(0)
net = models.resnet50(pretrained=True)
model = net.to(device)
```
We need to make sure our model is assigned to a GPU resource- here we do it one time before the training loops begin. We will also assign each image and its label to a GPU resource within the training and evaluation loops.
### 3. DDP Wrapper
```
model = DDP(model)
```
And finally, we need to enable the DistributedDataParallel framework. To do this, we are using the `DDP()` wrapper around the model, which is short for the PyTorch function `torch.nn.parallel.DistributedDataParallel`. There is a lot to know about this, but for our purposes the important thing is to understand that this allows the model training to run in parallel on our cluster. https://pytorch.org/docs/stable/notes/ddp.html
> **Discussing DDP**
It may be interesting for you to know what DDP is really doing under the hood: for a detailed discussion and more tips about this same workflow, you can visit our blog to read more! [https://www.saturncloud.io/s/combining-dask-and-pytorch-for-better-faster-transfer-learning/](https://www.saturncloud.io/s/combining-dask-and-pytorch-for-better-faster-transfer-learning/)
***
# Training time!
Our whole training process is going to be contained in one function, here named `run_transfer_learning`.
## Modeling Functions
Setting these pretty basic steps into a function just helps us ensure perfect parity between our train and evaluation steps.
```
def iterate_model(inputs, labels, model, device):
# Pass items to GPU
inputs = inputs.to(device)
labels = labels.to(device)
# Run model iteration
outputs = model(inputs)
# Format results
_, preds = torch.max(outputs, 1)
perct = [torch.nn.functional.softmax(el, dim=0)[i].item() for i, el in zip(preds, outputs)]
return inputs, labels, outputs, preds, perct
def run_transfer_learning(bucket, prefix, train_pct, batch_size,
n_epochs, base_lr, imagenetclasses,
n_workers = 1, subset = False):
'''Load basic Resnet50, run transfer learning over given epochs.
Uses dataset from the path given as the pool from which to take the
training and evaluation samples.'''
worker_rank = int(dist.get_rank())
# Set results writer
writer = SummaryWriter(f's3://pytorchtraining/pytorch_bigbatch/learning_worker{worker_rank}')
executor = ThreadPoolExecutor(max_workers=64)
# --------- Format model and params --------- #
device = torch.device("cuda")
net = models.resnet50(pretrained=True) # True means we start with the imagenet version
model = net.to(device)
model = DDP(model)
criterion = nn.CrossEntropyLoss().cuda()
optimizer = optim.SGD(model.parameters(), lr=base_lr, momentum=0.9)
# --------- Retrieve data for training and eval --------- #
whole_dataset = prepro_batches(bucket, prefix)
new_class_to_idx = {x: int(replace_label(x, imagenetclasses)[1]) for x in whole_dataset.classes}
whole_dataset.class_to_idx = new_class_to_idx
train, val = get_splits_parallel(train_pct, whole_dataset, batch_size=batch_size, subset = subset, workers = n_workers)
dataloaders = {'train' : train, 'val': val}
# --------- Start iterations --------- #
count = 0
t_count = 0
for epoch in range(n_epochs):
agg_loss = []
agg_loss_t = []
agg_cor = []
agg_cor_t = []
# --------- Training section --------- #
model.train() # Set model to training mode
for inputs, labels in dataloaders["train"]:
dt = datetime.datetime.now().isoformat()
inputs, labels, outputs, preds, perct = iterate_model(inputs, labels, model, device)
loss = criterion(outputs, labels)
correct = (preds == labels).sum().item()
# zero the parameter gradients
optimizer.zero_grad()
loss.backward()
optimizer.step()
count += 1
# Track statistics
for param_group in optimizer.param_groups:
current_lr = param_group['lr']
agg_loss.append(loss.item())
agg_cor.append(correct)
if ((count % 25) == 0):
future = executor.submit(
writer.add_hparams(
hparam_dict = {'lr': current_lr, 'bsize': batch_size, 'worker':worker_rank},
metric_dict = {'correct': correct,'loss': loss.item()},
name = 'train-iter',
global_step=count)
)
# Save a matplotlib figure showing a small sample of actual preds for spot check
# Functions used here are in setup2.py
if ((count % 50) == 0):
future = executor.submit(
writer.add_figure(
'predictions vs. actuals, training',
plot_classes_preds(model, inputs, labels, preds, perct, imagenetclasses),
global_step=count
)
)
# --------- Evaluation section --------- #
with torch.no_grad():
model.eval() # Set model to evaluation mode
for inputs_t, labels_t in dataloaders["val"]:
dt = datetime.datetime.now().isoformat()
inputs_t, labels_t, outputs_t, pred_t, perct_t = iterate_model(inputs_t, labels_t, model, device)
loss_t = criterion(outputs_t, labels_t)
correct_t = (pred_t == labels_t).sum().item()
t_count += 1
# Track statistics
for param_group in optimizer.param_groups:
current_lr = param_group['lr']
agg_loss_t.append(loss_t.item())
agg_cor_t.append(correct_t)
if ((t_count % 25) == 0):
future = executor.submit(
writer.add_hparams(
hparam_dict = {'lr': current_lr, 'bsize': batch_size, 'worker':worker_rank},
metric_dict = {'correct': correct_t,'loss': loss_t.item()},
name = 'eval-iter',
global_step=t_count)
)
future = executor.submit(
writer.add_hparams(
hparam_dict = {'lr': current_lr, 'bsize': batch_size, 'worker':worker_rank},
metric_dict = {'correct': np.mean(agg_cor),'loss': np.mean(agg_loss),
'last_correct': correct,'last_loss': loss.item()},
name = 'train',
global_step=epoch)
)
future = executor.submit(
writer.add_hparams(
hparam_dict = {'lr': current_lr, 'bsize': batch_size, 'worker':worker_rank},
metric_dict = {'correct': np.mean(agg_cor_t),'loss': np.mean(agg_loss_t),
'last_correct': correct_t,'last_loss': loss_t.item()},
name = 'eval',
global_step=epoch)
)
pickle.dump(model.state_dict(), s3.open(f"pytorchtraining/pytorch_bigbatch/model_epoch{epoch}_iter{count}_{dt}.pkl",'wb'))
```
######
Now we've done all the hard work, and just need to run our function! Using `dispatch.run` from `dask-pytorch-ddp`, we pass in the transfer learning function so that it gets distributed correctly across our cluster. This creates futures and starts computing them.
```
import math
import numpy as np
import multiprocessing as mp
import datetime
import json
import pickle
from concurrent.futures import ThreadPoolExecutor
num_workers = 64
s3 = s3fs.S3FileSystem()
with s3.open('s3://saturn-public-data/dogs/imagenet1000_clsidx_to_labels.txt') as f:
imagenetclasses = [line.strip() for line in f.readlines()]
client.restart() # Clears memory on cluster- optional but recommended.
startparams = {'n_epochs': 6,
'batch_size': 100,
'train_pct': .8,
'base_lr': 0.01,
'imagenetclasses':imagenetclasses,
'subset': True,
'n_workers': 3} #only necessary if you select subset
```
## Kick Off Job
### Send Tasks to Workers
We talked in Notebook 2 about how we distribute tasks to the workers in our cluster, and now you get to see it firsthand. Inside the `dispatch.run()` function in `dask-pytorch-ddp`, we are actually using the `client.submit()` method to pass tasks to our workers, and collecting these as futures in a list. We can prove this by looking at the results, here named "futures", where we can see they are in fact all pending futures, one for each of the workers in our cluster.
> *Why don't we use `.map()` in this function?*
> Recall that `.map` allows the Cluster to decide where the tasks are completed - it has the ability to choose which worker is assigned any task. That means that we don't have the control we need to ensure that we have one and only one job per GPU. This could be a problem for our methodology because of the use of DDP.
> Instead we use `.submit` and manually assign it to the workers by number. This way, each worker is attacking the same problem - our transfer learning problem - and pursuing a solution simultaneously. We'll have one and only one job per worker.
```
%%time
futures = dispatch.run(client, run_transfer_learning, bucket = "saturn-public-data", prefix = "dogs/Images", **startparams)
futures
futures
futures[0].result()
```
<img src="https://media.giphy.com/media/VFDeGtRSHswfe/giphy.gif" alt="parallel" style="width: 200px;"/>
Now we let our workers run for awhile. This step will take time, so you may not be able to see the full results during our workshop. See the dashboards to view the GPUs efforts as the job runs.
***
If you don't have access to an S3 bucket, but would still like to do model performance review, please visit [Notebook 6b](06b-transfer-training-local.ipynb) and [Notebook 7](07-learning-results.ipynb) to see detailed examples of how you can do that.
***
## Optional: Launch Tensorboard
### If you save files to S3
Open a terminal on your local machine, run `tensorboard --logdir=s3://[NAMEOFBUCKET]/runs`. Ensure that your AWS creds are in your bash profile/environment.
#### Example of creds you should have
export AWS_SECRET_ACCESS_KEY=`your secret key`
export AWS_ACCESS_KEY_ID=`your access key id`
export S3_REGION=us-east-2 `substitute your region`
export S3_ENDPOINT=https://s3.us-east-2.amazonaws.com `match to your region`
### If you save files locally
When you are ready to start viewing the board, run this at the terminal inside Jupyter Labs:
`tensorboard --logdir=runs`
Then, in a terminal on your local machine, run:
`ssh -L 6006:localhost:6006 -i ~/.ssh/PATHTOPRIVATEKEY SSHURLFORJUPYTER`
You'll find the private key path on your local machine, and the SSH URL on the project page for this project. You can change the local port (the first 6006) if you like.
At this stage, you'll likely not have any data, but the board will update itself every thirty seconds.
| github_jupyter |
# Supervised Fine-Tuning Best Artworks of All Time
Same as the base experiment, except that it was applied to extracted patches of paintings. Sample-level classification follows the `averaging` voting strategy.
Code: [github:lucasdavid/experiments/.../supervised/fine-tuning/best-artworks-of-all-time](https://github.com/lucasdavid/experiments/blob/main/notebooks/supervised/fine-tuning/best-artworks-of-all-time/best-artworks-of-all-time.ipynb)
Dataset: https://www.kaggle.com/ikarus777/best-artworks-of-all-time
Docker image: `tensorflow/tensorflow:latest-gpu-jupyter`
```
from time import time
import tensorflow as tf
class RC:
AUTOTUNE = tf.data.experimental.AUTOTUNE
seed = 21392
class DC:
path = '/tf/datasets/best-artworks-of-all-time'
images = path + '/images/patches'
info = path + '/artists.csv'
batch_size = 32
image_size = (299, 299)
channels = 3
input_shape = (batch_size, *image_size, channels)
buffer_size = 100000
class TC:
epochs = 200
learning_rate = .001
validation_split = .3
reduce_lr_on_plateau_pacience = 20
reduce_lr_on_plateau_factor = .5
early_stopping_patience = 50
splits = [f'train[{validation_split}:]', f'train[:{validation_split}]', 'test']
augment = True
epochs_fine_tuning = 0
learning_rate_fine_tuning = .0005
fine_tuning_layers = .2 # 20%
class LogConfig:
tensorboard = (f'/tf/logs/d:baoat-patches '
f'e:{TC.epochs} fte:{TC.epochs_fine_tuning} b:{DC.batch_size} '
f'v:{TC.validation_split} m:inceptionv3 aug:{TC.augment}'
f'/{int(time())}')
class Config:
run = RC
data = DC
training = TC
log = LogConfig
```
## Setup
```
import os
import pathlib
from math import ceil
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from tensorflow.keras import Model, Sequential, Input
from tensorflow.keras.layers import (Conv2D, Dense, Dropout, BatchNormalization,
Activation, Lambda)
def plot(y, titles=None, rows=1, i0=0):
for i, image in enumerate(y):
if image is None:
plt.subplot(rows, ceil(len(y) / rows), i0+i+1)
plt.axis('off')
continue
t = titles[i] if titles else None
plt.subplot(rows, ceil(len(y) / rows), i0+i+1, title=t)
plt.imshow(image)
plt.axis('off')
sns.set()
```
## Dataset
```
class Data:
info = pd.read_csv(Config.data.info)
data_dir = pathlib.Path(Config.data.images)
class_names = np.array(sorted([item.name
for item in data_dir.glob('*')
if item.name != "LICENSE.txt"]))
dataset_args = dict(
label_mode='int',
image_size=Config.data.image_size, batch_size=Config.data.batch_size,
validation_split=Config.training.validation_split,
seed=Config.run.seed)
train_ds = tf.keras.preprocessing.image_dataset_from_directory(
data_dir,
subset='training',
**dataset_args)
val_ds = tf.keras.preprocessing.image_dataset_from_directory(
data_dir,
subset='validation',
**dataset_args)
```
## Augmentation Policy
```
batchwise_augmentation = Sequential([
tf.keras.layers.experimental.preprocessing.RandomZoom((-.3, .3)),
tf.keras.layers.experimental.preprocessing.RandomFlip(),
tf.keras.layers.experimental.preprocessing.RandomRotation(0.2),
], name='batch_aug')
def augment_fn(image, label):
image = batchwise_augmentation(image)
image = tf.clip_by_value(image, 0, 255)
return image, label
def prepare(ds, augment=False):
ds = ds.cache()
if augment: ds = ds.map(augment_fn, num_parallel_calls=Config.run.AUTOTUNE)
return ds.prefetch(buffer_size=Config.run.AUTOTUNE)
train_ds = prepare(Data.train_ds, augment=Config.training.augment)
val_ds = prepare(Data.val_ds)
# test_ds = prepare(Data.test_ds)
for x, y in train_ds:
print('Shapes:', x.shape, 'and', y.shape)
print("Labels: ", y.numpy())
plt.figure(figsize=(16, 9))
plot(x.numpy().astype(int), rows=4)
plt.tight_layout()
break
```
## Model Definition
```
from tensorflow.keras.applications import inception_resnet_v2
encoder = inception_resnet_v2.InceptionResNetV2(include_top=False, pooling='avg',
input_shape=Config.data.input_shape[1:])
# encoder = Model(encoder.input, encoder.get_layer('block_9_add').output)
def encoder_pre(x):
return Lambda(inception_resnet_v2.preprocess_input, name='pre_incresnet')(x)
from tensorflow.keras.layers import GlobalAveragePooling2D
def dense_block(x, units, activation='relu', name=None):
y = Dense(units, name=f'{name}_fc', use_bias=False)(x)
y = BatchNormalization(name=f'{name}_bn')(y)
y = Activation(activation, name=f'{name}_relu')(y)
return y
def discriminator():
y = x = Input(shape=Config.data.input_shape[1:], name='inputs')
y = encoder_pre(y)
y = encoder(y)
y = Dense(len(Data.class_names), name='predictions')(y)
return tf.keras.Model(x, y, name='author_disc')
disc = discriminator()
disc.summary()
disc.get_layer('inception_resnet_v2').trainable = False
from tensorflow.keras import losses, metrics, optimizers
disc.compile(
optimizer=optimizers.Adam(lr=Config.training.learning_rate),
loss=losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=[
metrics.SparseCategoricalAccuracy(),
metrics.SparseTopKCategoricalAccuracy()
]
)
```
## Training
### Initial Training for Final Classification Layer
The final layer --- currently containing random values --- must be first adjusted to match the the encoder's layers' current state.
```
from tensorflow.keras import callbacks
cs = [
callbacks.TerminateOnNaN(),
callbacks.ModelCheckpoint(Config.log.tensorboard + '/weights.h5',
save_best_only=True,
save_weights_only=True,
verbose=1),
callbacks.ReduceLROnPlateau(patience=Config.training.reduce_lr_on_plateau_pacience,
factor=Config.training.reduce_lr_on_plateau_factor,
verbose=1),
callbacks.EarlyStopping(patience=Config.training.early_stopping_patience, verbose=1),
callbacks.TensorBoard(Config.log.tensorboard, write_graph=False)
]
try:
disc.fit(
train_ds,
validation_data=val_ds,
epochs=Config.training.epochs,
initial_epoch=0,
callbacks=cs);
except KeyboardInterrupt:
print('stopped')
disc.load_weights(Config.log.tensorboard + '/weights.h5')
disc.get_layer('inception_resnet_v2').trainable = True
disc.save_weights(Config.log.tensorboard + '/weights.h5')
```
### Fine-Tuning All Layers
```
if Config.training.epochs_fine_tuning:
_enc = disc.get_layer('inception_resnet_v2')
ft_layer_ix = int((1-Config.training.fine_tuning_layers)*len(_enc.layers))
for ix, l in enumerate(_enc.layers):
l.trainable = ix >= ft_layer_ix
try: disc.fit(
train_ds,
validation_data=val_ds,
initial_epoch=disc.history.epoch[-1] + 1,
epochs=len(disc.history.epoch) + Config.training.epochs_fine_tuning,
callbacks=cs);
except KeyboardInterrupt: print('stopped')
if Config.training.epochs_fine_tuning:
disc.load_weights(Config.log.tensorboard + '/weights.h5')
for ix, l in enumerate(_enc.layers):
l.trainable = True
_enc.trainable = True
disc.save_weights(Config.log.tensorboard + '/weights.h5')
```
## Testing
```
disc.load_weights(Config.log.tensorboard + '/weights.h5')
from sklearn import metrics as skmetrics
def labels_and_predictions(model, ds):
labels, predictions = [], []
for x, y in ds:
p = model(x).numpy()
p = p.argmax(axis=1)
labels.append(y.numpy())
predictions.append(p)
labels, predictions = np.concatenate(labels), np.concatenate(predictions)
labels, predictions = Data.class_names[labels], Data.class_names[predictions]
return labels, predictions
def evaluate(model, ds):
labels, predictions = labels_and_predictions(model, ds)
print('balanced acc:', skmetrics.balanced_accuracy_score(labels, predictions))
print('accuracy :', skmetrics.accuracy_score(labels, predictions))
print('Classification report:')
print(skmetrics.classification_report(labels, predictions))
```
#### Training Report
```
evaluate(disc, train_ds)
```
#### Validation Report
```
evaluate(disc, val_ds)
```
#### Test Report
```
test_ds = val_ds
evaluate(disc, test_ds)
labels, predictions = labels_and_predictions(disc, test_ds)
cm = skmetrics.confusion_matrix(labels, predictions)
cm = cm / cm.sum(axis=1, keepdims=True)
sorted_by_most_accurate = cm.diagonal().argsort()[::-1]
cm = cm[sorted_by_most_accurate][:, sorted_by_most_accurate]
plt.figure(figsize=(12, 12))
with sns.axes_style("white"):
sns.heatmap(cm, cmap='RdPu', annot=False, cbar=False,
yticklabels=Data.class_names[sorted_by_most_accurate], xticklabels=False);
def plot_predictions(model, ds, take=1):
figs, titles = [], []
plt.figure(figsize=(16, 16))
for ix, (x, y) in enumerate(ds.take(take)):
p = model.predict(x)
p = tf.nn.softmax(p).numpy()
figs.append(x.numpy().astype(int))
titles.append([f'label: {a}\npredicted: {b}\nproba:{c:.0%}'
for a, b, c in zip(Data.class_names[y],
Data.class_names[p.argmax(axis=-1)],
p.max(axis=-1))])
plot(np.concatenate(figs),
titles=sum(titles, []),
rows=6)
plt.tight_layout()
plot_predictions(disc, train_ds)
```
| github_jupyter |
```
# Direct Python to plot all figures inline (i.e., not in a separate window)
%matplotlib inline
# Load libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from scipy import stats
# Division of two integers in Python 2.7 does not return a floating point result. The default is to round down
# to the nearest integer. The following piece of code changes the default.
from __future__ import division
# Directory where the csv data file is located
workdir = '../../data/raw_data/'
data_message = pd.read_csv('../../data/raw_data/AAPL_05222012_0930_1300_message.tar.gz',compression='gzip')
data_lob = pd.read_csv('../../data/raw_data/AAPL_05222012_0930_1300_LOB_2.tar.gz',compression='gzip')
#drop redundant time
col_names=data_lob.columns
delete_list=[i for i in col_names if 'UPDATE_TIME' in i]
for i in delete_list:
data_lob=data_lob.drop(i,1)
#functions for renaming
def rename(txt):
txt=txt[16:].split('..')[0]
index=0
ask_bid=''
p_v=''
if txt[-2].isdigit():
index=txt[-2:]
else:
index=txt[-1]
if txt[:3]=="BID":
ask_bid='bid'
else:
ask_bid='ask'
if txt[4:9]=="PRICE":
p_v='P'
else:
p_v='V'
return('_'.join([p_v,index,ask_bid]))
#rename columns
col_names=data_lob.columns
new_col_names=[]
new_col_names.append('index')
new_col_names.append('Time')
for i in col_names[2:]:
new_col_names.append(rename(i))
len(new_col_names)
data_lob.columns=new_col_names
#feature: bid-ask spreads and mid price
for i in list(range(1, 11)):
bid_ask_col_name='_'.join(['spreads',str(i)])
p_i_ask='_'.join(['P',str(i),'ask'])
p_i_bid='_'.join(['P',str(i),'bid'])
data_lob[bid_ask_col_name]=data_lob[p_i_ask]-data_lob[p_i_bid]
mid_price_col_name = '_'.join(['mid_price',str(i)])
data_lob[mid_price_col_name]=(data_lob[p_i_ask]+data_lob[p_i_bid])/2
data_lob.head()
```
# Analyze $\Delta t$ and generate good graphs!
```
#compute frequency of midprice stationary
diff=data_lob['mid_price_1']
bid = data_lob['P_1_bid']
ask = data_lob['P_1_ask']
# print('t', 'up', 'stationary','down', 'up', 'stationary','down')
upper_step = 100
for t in list(range(1, upper_step, 5)):
print(t)
# label from mid-price
delta_midprice = np.array(diff[t:]) - np.array(diff[:(-1*t)])
dummy_up = delta_midprice > 0
dummy_down = delta_midprice < 0
dummy_stationary = delta_midprice == 0
freq_stationary = sum(dummy_stationary) / len(dummy_stationary)
freq_up = sum(dummy_up) / len(dummy_up)
freq_down = sum(dummy_down) / len(dummy_down)
if t == 1:
delta_t = np.array([t,])
up_MP = np.array([freq_up,])
stationary_MP = np.array([freq_stationary,])
down_MP = np.array([freq_down,])
else:
delta_t = np.append(delta_t, t)
up_MP = np.append(up_MP, freq_up)
stationary_MP = np.append(stationary_MP, freq_stationary)
down_MP = np.append(down_MP, freq_down)
df_MP = pd.DataFrame({'delta_t' : delta_t,
'1' : up_MP ,
'0' : stationary_MP ,
'-1': down_MP})
upper_step_sc = 2000
for t in range(1, upper_step_sc, 100):
print(t)
# label from spread crossing
up = np.array(bid[t:]) - np.array(ask[:(-1*t)]) > 0
down = np.array(ask[t:]) - np.array(bid[:(-1*t)]) < 0
stationary = np.logical_not(np.logical_or(up, down))
freq_stationary_crossing = sum(stationary) / len(stationary)
freq_up_crossing = sum(up) / len(up)
freq_down_crossing = sum(down) / len(down)
if t == 1:
delta_t = np.array([t,])
up_SC = np.array([freq_up_crossing,])
stationary_SC = np.array([freq_stationary_crossing,])
down_SC = np.array([freq_down_crossing,])
else:
delta_t = np.append(delta_t, t)
up_SC = np.append(up_SC, freq_up_crossing)
stationary_SC = np.append(stationary_SC, freq_stationary_crossing)
down_SC = np.append(down_SC, freq_down_crossing)
df_SC = pd.DataFrame({'delta_t' : delta_t,
'1' : up_SC ,
'0' : stationary_SC ,
'-1': down_SC})
print(df_MP)
print(df_SC)
fig_MP = plt.figure()
plt.plot(df_MP['delta_t'], df_MP['1'], 'r--',
df_MP['delta_t'], df_MP['0'], 'b--',
df_MP['delta_t'], df_MP['-1'], 'g--')
plt.legend(['Up', 'Stationary', 'Down'])
plt.xlabel('delta t')
plt.ylabel('Frequency')
plt.title('Proportion of labels: Midprice')
plt.show()
fig_SC = plt.figure()
plt.plot(df_SC['delta_t'], df_SC['1'], 'r--',
df_SC['delta_t'], df_SC['0'], 'b--',
df_SC['delta_t'], df_SC['-1'], 'g--')
plt.legend(['Up', 'Stationary', 'Down'])
plt.xlabel('delta t')
plt.ylabel('Frequency')
plt.title('Proportion of labels: Bid-ask spread crossing')
plt.show()
fig_MP.savefig('../report/delta_t_MP.png')
fig_SC.savefig('../report/delta_t_SC.png')
```
# A test for profit calculator
```
##### Old version #####
def profit_calculator(data, delta_t = 30, simple = True):
"""Calculate the profit of trading strategy based on precisely the prediction of the model
Parameters
----------
data : a data frame with "predicted" "P_1_bid" "P_1_ask"
delta_t : time gap between
simple : a dummy, True, means we make transection decisions only every delta_t period. False, means we track the current
hand every period, only if we don't have anything at hand, we make transactions
Returns
-------
profit : a numeric, the net profit at the end
profit_series : a np.array, time series tracking net profit at each point of time
"""
if simple == True:
data_effective = data.loc[np.arange(len(data)) % delta_t == 0]
bid = data_effective['P_1_bid']
ask = data_effective['P_1_ask']
trade_decision = data_effective['predicted'][:-1]
buy_profit = np.array(bid[1:]) - np.array(ask[:-1])
profit = sum(trade_decision * buy_profit)
return profit
else:
# print(data_effective)
# print(data['P_1_bid'])
# print(data['P_1_ask'])
# print(data['P_1_bid'][delta_t:])
# print(data['P_1_ask'][:(-1 * delta_t)])
# print(np.array(data['P_1_bid'][delta_t:]))
# print(np.array(data['P_1_ask'][:(-1 * delta_t)]))
buy_profit = np.array(data['P_1_bid'][delta_t:]) - np.array(data['P_1_ask'][:(-1 * delta_t)])
trade_decision_draft = data['predicted'][:(-1 * delta_t)]
T = len(buy_profit)
#print(T)
#print(buy_profit)
#print(trade_decision_draft)
current_state = [0] * T
trade_decision = [0] * T
#print(current_state[T])
for i in range(T):
if current_state[i] == 1:
trade_decision[i] = 0
else:
trade_decision[i] = trade_decision_draft[i]
if i < T-1:
#print(i)
current_state[i+1] = int(sum(trade_decision[max(0, i - delta_t):i]) != 0)
profit = sum(trade_decision * buy_profit)
return profit
d = {'P_1_bid' : pd.Series([1., 2., 3., 5., 1., 2., 3., 5.]),
'P_1_ask' : pd.Series([1., 2., 3., 4., 1., 2., 3., 4.]),
'predicted' : pd.Series([0, 1, -1, 1, 1, 1, -1, 1])}
df = pd.DataFrame(d)
print(df)
print(profit_calculator(df, delta_t = 1, simple = True))
print(profit_calculator(df, delta_t = 1, simple = False))
df.iloc[:-1,:]
##### New version #####
def profit_calculator(data, delta_t = 30, simple = False):
"""Calculate the profit of trading strategy based on precisely the prediction of the model
Parameters
----------
data : a data frame with "predicted" "P_1_bid" "P_1_ask"
delta_t : time gap between
simple : a dummy, True, means we make transection decisions only every delta_t period. False, means we track the current
hand every period, only if we don't have anything at hand, we make new transactions
Returns
-------
profit : a numeric, the net profit at the end
"""
if simple == True:
data_effective = data.loc[np.arange(len(data)) % delta_t == 0]
bid = data_effective['P_1_bid']
ask = data_effective['P_1_ask']
trade_decision = data_effective['predicted'][:-1]
buy_profit = np.array(bid[1:]) - np.array(ask[:-1])
sell_profit = np.array(bid[:-1]) - np.array(ask[1:])
profit = sum((np.array(trade_decision) > 0) * buy_profit + (np.array(trade_decision) < 0) * sell_profit)
print(np.array(trade_decision) > 0, np.array(trade_decision) < 0, buy_profit, sell_profit)
print((np.array(trade_decision) > 0) * buy_profit)
return profit
else:
buy_profit = np.array(data['P_1_bid'][delta_t:]) - np.array(data['P_1_ask'][:(-1 * delta_t)])
sell_profit = np.array(data['P_1_bid'][:(-1 * delta_t)]) - np.array(data['P_1_ask'][delta_t:])
trade_decision_draft = data['predicted'][:(-1 * delta_t)]
T = len(buy_profit)
current_state = [0] * T
trade_decision = [0] * T
profit = 0
for i in range(T):
if current_state[i] == 1:
trade_decision[i] = 0
else:
trade_decision[i] = trade_decision_draft[i]
if trade_decision[i] == 1:
profit += buy_profit[i]
elif trade_decision[i] == -1:
profit += sell_profit[i]
if i < T-1:
current_state[i+1] = int(sum(trade_decision[max(0, i - delta_t):i]) != 0)
profit = sum(trade_decision * buy_profit)
return profit
```
| github_jupyter |
Processing Copy and Paste from THES Rankings
========================
Simple processing of the text files that result from copying and pasting the tables from the THES rankings. There are two tabs in the table, one gives rankings alongside some demographic information. The other other gives the elements that make up the score.
Each tab was copy-pasted into a text file (THES_2016-17_ranks.txt and THES_2016-17_scores.txt) which have some idiosyncratic formatting. The goal here is to bring them into a single csv file that has been normalised to a standardised set of countries and ultimately to GRID IDs.
```
import os.path
input_data_folder = 'data/input/'
thes = []
filepath = os.path.join(input_data_folder, 'THES_2016-17_ranks.txt' )
with open(filepath, encoding = 'utf-8-sig') as f:
lines = f.readlines()
for n in range(0, len(lines), 3):
uni = {}
uni['rank'], uni['name'] = lines[n].split(' ')
#uni['name'] = uni['name'].rstrip('\n')
uni['country'] = lines[n+1]
uni['student_fte'], uni['students_per_staff'], \
uni['foreign_students_%'], uni['gender_ratio'] = lines[n+2].split(' ')
for k in uni.keys():
uni[k] = uni[k].rstrip('\n').rstrip('%')
if uni['foreign_students_%'] != 'n/a':
uni['foreign_students_%'] = int(uni['foreign_students_%'])
uni['student_fte'] = int(''.join(uni['student_fte'].split(',')))
uni['students_per_staff'] = float(uni['students_per_staff'])
if uni['gender_ratio'] != 'n/a':
uni['female_%'] = int(uni['gender_ratio'].split(':')[0].rstrip(' '))
thes.append(uni)
```
Check the length and the that the first and last entry are correct
```
len(thes)
thes[0]
thes[1102]
```
Now to load in the other file and load the other variables. Some of the overall scores are given as ranges so will leave those as strings for the moment.
```
filepath = os.path.join(input_data_folder, 'THES_2016-17_scores.txt')
with open(filepath, encoding = 'utf-8-sig') as f:
lines = f.readlines()
for n in range(0, len(lines), 3):
uni = {}
rank, name = lines[n].split(' ')
uni['overall_score'], uni['teaching_score'], uni['research_score'], \
uni['citations'], uni['industry_income'], uni['international_outlook'] = lines[n+2].split(' ')
for k in uni.keys():
try:
uni[k] = float(uni[k])
except ValueError:
print(n, name, k, uni[k])
thes_line = thes[int(n/3)]
try:
assert rank == thes_line['rank']
assert name.rstrip('\n') == thes_line['name']
except AssertionError:
print(uni['rank'], thes_line['rank'])
print(uni['name'], thes_line['name'])
thes_line.update(uni)
```
Cleaning up the rankings
-----------------------
Some cleanup is required to make use of the ranking properly. Best done by setting up a new attribute using the equal rank for both/all of those where that is the case and the lower rank where a range is given.
```
for u in thes:
u['int_rank'] = u['rank'].lstrip('=')
if '–' in u['rank']:
u['int_rank'] = u['int_rank'].split('–')[1]
if u['int_rank'] == '1001+':
u['int_rank'] = 1103
u['int_rank'] = int(u['int_rank'])
unique_rankings = set()
for uni in thes:
unique_rankings.add(uni['int_rank'])
unique_rankings = list(unique_rankings)
unique_rankings.sort()
unique_rankings
unique_countries = set()
for uni in thes:
unique_countries.add(uni['country'])
unique_countries = list(unique_countries)
unique_countries.sort()
country_counts = []
for country in unique_countries:
country_counts.append((country, len([u for u in thes if u['country'] == country])))
country_counts
```
Attempting to analyse by region
-----------------------------
We load the countries.json file from https://github.com/mledoze/countries and aim to normalise by country and determine a sub-region for each university for analysis. The subregions are the roughly right level of granularity for this analysis.
```
import json
with open(os.path.join(input_data_folder,'countries.json')) as f:
countries = json.load(f)
country_lookup = {}
for i, country in enumerate(countries):
country_lookup[country['name']['common']] = i
country_lookup[country['name']['official']] = i
```
Identify those cases where we don't have a matching country to the standard set
```
for uni in thes:
try:
country_lookup[uni['country']]
except KeyError:
print(uni['name'], uni['country'])
```
These two universities show countries that are not in our normalised data set. One appears to be due to a differing spelling of Macau and the other is a political issue as to the status of Northern Cyprus. The spelling is easily fixed. For the purpose of this analysis we will treat Northern and Southern Cyprus as the one country.
```
country_lookup.get('Macau')
country_lookup['Macao'] = 137
country_lookup.get('Cyprus')
country_lookup['Northern Cyprus'] = 58
for uni in thes:
i = country_lookup[uni['country']]
uni['country_code'] = countries[i].get('cca2')
uni['region'] = countries[i].get('region')
uni['subregion'] = countries[i].get('subregion')
```
Creating a set of groups based on the ranking and creating a new heading for these. Moderately arbitrary grouping into 0-10, 11-50, 51-100, 101-200, 201-500, 1000-
```
for uni in thes:
if uni['int_rank'] <=15:
uni['rank_group'] = '0001-15'
if uni['int_rank'] <=50 and uni['int_rank'] > 15:
uni['rank_group'] = '0016-50'
if uni['int_rank'] <=100 and uni['int_rank'] > 50:
uni['rank_group'] = '0051-100'
if uni['int_rank'] <=200 and uni['int_rank'] > 100:
uni['rank_group'] = '0101-200'
if uni['int_rank'] <=500 and uni['int_rank'] > 200:
uni['rank_group'] = '0201-500'
if uni['int_rank'] <=1000 and uni['int_rank'] > 500:
uni['rank_group'] = '0501-1000'
if uni['int_rank'] <=2000 and uni['int_rank'] > 1000:
uni['rank_group'] = '1000+'
```
Will write out to a csv file for further manipulation and later normalisation to GRID ID's for each university, as well as to allow for sampling.
```
import csv
output_data_folder = 'data/output/'
with open(os.path.join(output_data_folder,'thes_2016_ranking.csv'), 'w', encoding = 'utf-8') as f:
writer = csv.DictWriter(f, fieldnames=[f for f in iter(thes[0])])
writer.writeheader()
writer.writerows(thes)
```
| github_jupyter |
# LSTM Model to detect Heel Strike and Toe-off events
In this notebook, we will predict whether each sample is a heel strike or toe-off based on the accelerometer data. To do so, we train an LSTM (Long-Short Term Memory) model on 70% of the data and test it on the remaining 30%
The LSTM architecture has
- a lookback window of 3 timesteps
- 6 features (X,Y,Z acceleration of left foot and right foot)
- 44 hidden nodes
The first step to select the Subject Id, activity and event of interest
The subsequent steps to tune and test the model are as follows:
1. Subset the data based on subject, activity, and event
2. Oversampling and adding in composite accelerations
3. Remove noise from the data
4. Re-scale each of the features to range (0,5)
5. Split the data into training and testing sets
6. Since we are using an LSTM architecture, we will implement a look-back function that will introduce historical data determined by the window size
7. Build and compile the Neural Network
8. Fit the model
9. Save the predictions and actual value to evaluate the model. The model is evaluated in terms of
- F1 score
- Percentage of true positives
- Mean of the absolute difference in time between true positives and corresponding GT events
# Note: before you run the code
You can adjust the subject Id, activity and event in the next cell.
Some parts are specific to indoors/outdoors, so you will need to comment the part if needed, and uncomment it it's if not needed.
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
```
## 1. Subset the data based on subject, activity and event
```
####### Select Subject Id #######
# Key in a number from 1 to 20.
# 1 to 11: indoors
# 12 to 20: outdoors
SUBJECT_ID = '20'
###### Select Activity ######
# Key in one of the options below:
## For indoors:
# treadmill_walk
# treadmill_run
# treadmill_walknrun
# treadmill_all (treadmill_walk + treadmill_walknrun + treadmill_slope_walk)
# indoor_walk
# indoor_run
# indoor_walknrun
## For outdoors:
# outdoor_walk
# outdoor_run
# outdoor_walknrun
ACTIVITY = 'outdoor_walknrun'
###### Select Event ######
# LF_HS
# LF_TO
# RF_HS
# RF_TO
EVENT = 'RF_TO'
# read in the data
DATA_PATH = './Combined Data_csv format/'
df = pd.read_csv(DATA_PATH + 'Sub_'+ SUBJECT_ID + '.csv', header = 0)
df = df.drop(df.columns[0], axis=1)
df.head()
```
Since there is no indicator variable for treadmill_all for subjects 1-11, let's create another column for this. The value of this data is 1 if either treadmill_walknrun is 1 or treadmill_slope_walk is 1
```
###############################################
######## Specific to indoors only #############
###############################################
# for SUBJECT_ID 1 to 11, uncomment if otherwise
# def label_treadmill_all (row):
# if row['treadmill_walknrun'] == 1:
# return 1
# if row['treadmill_slope_walk'] == 1:
# return 1
# return 0
# df['treadmill_all']=df.apply (lambda row: label_treadmill_all(row),axis=1)
# def label_indoor_run (row):
# if row['indoor_walknrun'] == 1 and row['indoor_walk'] == 0:
# return 1
# return 0
# df['indoor_run']=df.apply (lambda row: label_indoor_run(row),axis=1)
# def label_treadmill_run (row):
# if row['treadmill_walknrun'] == 1 and row['treadmill_walk'] == 0:
# return 1
# return 0
# df['treadmill_run']=df.apply (lambda row: label_indoor_run(row),axis=1)
# df.head()
# ###############################################
# ######## Specific to outdoors only ############
# ###############################################
# for SUBJECT_ID 12 to 20, uncomment if otherwise
def label_outdoor_run (row):
if row['outdoor_walknrun'] == 1 and row['outdoor_walk'] == 0:
return 1
return 0
df['outdoor_run']=df.apply (lambda row: label_outdoor_run(row),axis=1)
df.head()
# data munging: change the indicator variable +/- 2 timesteps away from the true value to '1'
def oversample(event):
global df
temp_list = []
for i in range(len(df)):
if i>=2 and i+3<=len(df):
if df[event][i-1]==1 or df[event][i]==1 or df[event][i+1]==1:
# if df[event][i-2]==1 or df[event][i-1]==1 or df[event][i]==1 or df[event][i+1]==1 or df[event][i+2]==1:
temp_list.append(1)
else:
temp_list.append(0)
else:
temp_list.append(0)
# return temp_list
df[event+'_2']=temp_list
oversample(EVENT)
# Subset out the data by activity of interest
k1=df[df[ACTIVITY]==1]
k1.head()
# Add in the composite acceleration: sqrt(accX^2+accY^2+accZ^2)
k1['acc_LF'] = np.sqrt(k1['accX_LF']*k1['accX_LF']+k1['accY_LF']*k1['accY_LF']+k1['accZ_LF']*k1['accZ_LF'])
k1['acc_RF'] = np.sqrt(k1['accX_RF']*k1['accX_RF']+k1['accY_RF']*k1['accY_RF']+k1['accZ_RF']*k1['accZ_RF'])
k1['acc_LF_xy'] = np.sqrt(k1['accX_LF']*k1['accX_LF']+k1['accY_LF']*k1['accY_LF'])
k1['acc_RF_xy'] = np.sqrt(k1['accX_RF']*k1['accX_RF']+k1['accY_RF']*k1['accY_RF'])
k1['acc_LF_yz'] = np.sqrt(k1['accY_LF']*k1['accY_LF']+k1['accZ_LF']*k1['accZ_LF'])
k1['acc_RF_yz'] = np.sqrt(k1['accY_RF']*k1['accY_RF']+k1['accZ_RF']*k1['accZ_RF'])
k1['acc_LF_xz'] = np.sqrt(k1['accX_LF']*k1['accX_LF']+k1['accZ_LF']*k1['accZ_LF'])
k1['acc_RF_xz'] = np.sqrt(k1['accX_RF']*k1['accX_RF']+k1['accZ_RF']*k1['accZ_RF'])
# Select the columns you want -- the first few accelerometer data and event of interest
k2 = k1[['accX_LF','accY_LF','accZ_LF','accX_RF','accY_RF','accZ_RF','acc_LF','acc_RF','acc_LF_xy','acc_RF_xy','acc_LF_yz','acc_RF_yz','acc_LF_xz','acc_RF_xz', EVENT, EVENT+'_2']]
k2.head()
```
## 2. Remove noise from the data
```
k2['accX_LF_median']=k2['accX_LF'].rolling(window=3).mean()
k2['accY_LF_median']=k2['accY_LF'].rolling(window=3).mean()
k2['accZ_LF_median']=k2['accZ_LF'].rolling(window=3).mean()
k2['acc_LF_median']=k2['acc_LF'].rolling(window=3).mean()
k2['acc_LF_median_xy']=k2['acc_LF_xy'].rolling(window=3).mean()
k2['acc_LF_median_yz']=k2['acc_LF_yz'].rolling(window=3).mean()
k2['acc_LF_median_xz']=k2['acc_LF_xz'].rolling(window=3).mean()
k2['accX_RF_median']=k2['accX_RF'].rolling(window=3).mean()
k2['accY_RF_median']=k2['accY_RF'].rolling(window=3).mean()
k2['accZ_RF_median']=k2['accZ_RF'].rolling(window=3).mean()
k2['acc_RF_median']=k2['acc_RF'].rolling(window=3).mean()
k2['acc_RF_median_xy']=k2['acc_RF_xy'].rolling(window=3).mean()
k2['acc_RF_median_yz']=k2['acc_RF_yz'].rolling(window=3).mean()
k2['acc_RF_median_xz']=k2['acc_RF_xz'].rolling(window=3).mean()
k2[:10]
# Visualise
k2[['accX_LF','accX_LF_median']][:500].plot()
k2[['accY_LF','accY_LF_median']][:500].plot()
k2[['accZ_LF','accZ_LF_median']][:500].plot()
k2[['acc_LF','acc_LF_median']][:500].plot()
k2[['acc_LF_xy','acc_LF_median_xy']][:500].plot()
k2[['acc_LF_yz','acc_LF_median_yz']][:500].plot()
k2[['acc_LF_xz','acc_LF_median_xz']][:500].plot()
k2[['accX_RF','accX_RF_median']][:500].plot()
k2[['accY_RF','accY_RF_median']][:500].plot()
k2[['accZ_RF','accZ_RF_median']][:500].plot()
k2[['acc_RF','acc_RF_median']][:500].plot()
k2[['acc_RF_xy','acc_RF_median_xy']][:500].plot()
k2[['acc_RF_yz','acc_RF_median_yz']][:500].plot()
k2[['acc_RF_xz','acc_RF_median_xz']][:500].plot()
# Select the columns you want -- the rolling median accelerometer data and event of interest
# Remove the first two rows which have no accelerometer data
k3 = k2[['accX_LF_median','accY_LF_median','accZ_LF_median','acc_LF_median','acc_LF_median_xy','acc_LF_median_yz','acc_LF_median_xz','accX_RF_median','accY_RF_median','accZ_RF_median','acc_RF_median','acc_RF_median_xy','acc_RF_median_yz', 'acc_RF_median_xz', EVENT, EVENT+'_2']]
k3 = k3.iloc[2:]
k3.head()
```
## 3. Re-scale each of the features to range (0,5)
"LSTMs are sensitive to the scale of the input data, specifically when the sigmoid (default) or tanh activation functions are used. It can be a good practice to rescale the data, also called normalizing. We can easily normalize the dataset using the MinMaxScaler preprocessing class from the scikit-learn library."
```
dataset = k3.values
from sklearn.preprocessing import MinMaxScaler
# normalize the dataset
scaler = MinMaxScaler(feature_range=(0, 5))
dataset = scaler.fit_transform(dataset)
```
## 4. Split the data into training and test sets
```
# split into train and test sets
train_size = int(len(dataset) * 0.7)
test_size = len(dataset) - train_size
train, test = dataset[0:train_size,:], dataset[train_size:len(dataset),:]
print(len(train), len(test))
print(train)
# Example of first 500 and last 500 samples in training dataset
fig = plt.figure(figsize=(15,4))
ax = fig.add_subplot(1,2,1)
ax.plot(train[:500])
ax = fig.add_subplot(1,2,2)
ax.plot(test[-500:])
```
## 5. Implement a look-back function that will introduce historical data determined by the window size
Now we can define a function to create a new dataset, as described above.
The function takes two arguments: the dataset, which is a NumPy array that we want to convert into a dataset, and the look_back, which is the number of previous time steps to use as input variables to predict the next time period — in this case defaulted to 1
There are three sets of data we want in the training and testing datasets
1. X - these are the features, inputs to the LSTM model.
2. Y - these are the oversampled HS/TOs which we use for our training.
3. Z - these are the ground truth HS and TOs for evaluation.
```
# convert an array of values into a dataset matrix
def create_dataset(dataset, look_back=1):
dataX, dataY, dataZ = [], [], []
for i in range(len(dataset)-look_back-1):
a = dataset[i:(i+look_back), 0:14]
dataX.append(a)
dataY.append(dataset[i + look_back, 15])
dataZ.append(dataset[i + look_back, 14])
return numpy.array(dataX), numpy.array(dataY), numpy.array(dataZ)
import numpy
look_back=3 # 3 timesteps
trainX, trainY, trainZ = create_dataset(train, look_back)
testX, testY, testZ = create_dataset(test, look_back)
trainX[0:5]
trainY[0:500]
trainZ[0:500]
```
## 6. Build and compile the Neural Network
input to be [samples, time steps, features]
```
from keras.models import Sequential
from keras.layers import LSTM, Dense, Dropout
model = Sequential()
# Recurrent layer
model.add(LSTM(44, return_sequences=True, dropout=0.1, recurrent_dropout=0.1))
# Fully connected layer
model.add(Dense(44, activation='relu'))
# Recurrent layer
model.add(LSTM(44, return_sequences=False, dropout=0.1, recurrent_dropout=0.1))
# Fully connected layer
model.add(Dense(44, activation='relu'))
# Dropout for regularization
model.add(Dropout(0.5))
# Output layer
model.add(Dense(1, activation='sigmoid'))
# Compile the model
model.compile(
optimizer='adam', loss='binary_crossentropy', metrics=['accuracy'])
```
## 7. Fit the model
Further split the training dataset into 2/3 train and 1/3 validation.
Epoch size is 20
Batch size is 20
```
from keras_tqdm import TQDMNotebookCallback
from keras.callbacks import EarlyStopping
es= EarlyStopping(monitor='val_loss', min_delta=0,patience=5,verbose=0, mode='auto')
# Fit the model
history = model.fit(trainX, trainY, validation_split = 0.33, epochs=50, batch_size=20, verbose=0, callbacks=[TQDMNotebookCallback(),es])
from matplotlib import pyplot as plt
# List all data in history
print(history.history.keys())
# summarize history for accuracy
plt.plot(history.history['acc'])
plt.plot(history.history['val_acc'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train','validation'])
plt.show()
# summarize history for loss
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train','validation'])
plt.show()
model.summary()
```
## 8. Save the predictions and actual value to evaluate the model
The predictions are floating point values and we recode them as 1 if greater than 0.5 and 0 otherwise.
```
testY_predict = model.predict(testX)
print(testY_predict)
print(len(testZ), len(testY_predict))
predicted = []
actual = []
for i in range(len(testZ)):
#print(int(testY_predict[i]>0.5), int(testZ[i]>0.5))
predicted.append(int(testY_predict[i]>0.5))
actual.append(int(testZ[i]>0.5))
PREDICTIONS_DF = pd.DataFrame(list(zip(predicted, actual)),
columns=['predicted','actual'])
PREDICTIONS_DF.head()
# write dataframe to .csv
SAVE_PATH = './Predicted Data_model_csv format/'
PREDICTIONS_DF.to_csv(SAVE_PATH+'Sub'+SUBJECT_ID+'_'+ ACTIVITY + '_'+ EVENT + '.csv', encoding='utf-8')
```
| github_jupyter |
# Doc2Vec
```
import collections
import glob
from itertools import chain
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
%matplotlib inline
np.random.seed(1)
files = glob.glob('*.txt')
words = []
for f in files:
file = open(f)
words.append(file.read())
file.close()
words = list(chain.from_iterable(words))
words = ''.join(words)[:-1]
sentences = words.split('\n')
len(sentences)
vocabulary_size = 40000
def build_dataset(sentences):
words = ''.join(sentences).split()
count = [['UNK', -1]]
count.extend(collections.Counter(words).most_common(vocabulary_size - 1))
dictionary = dict()
for word, _ in count:
dictionary[word] = len(dictionary)
unk_count = 0
sent_data = []
for sentence in sentences:
data = []
for word in sentence.split():
if word in dictionary:
index = dictionary[word]
else:
index = 0 # dictionary['UNK']
unk_count = unk_count + 1
data.append(index)
sent_data.append(data)
count[0][1] = unk_count
reverse_dictionary = dict(zip(dictionary.values(), dictionary.keys()))
return sent_data, count, dictionary, reverse_dictionary
data, count, dictionary, reverse_dictionary = build_dataset(sentences)
print('Most common words (+UNK)', count[:5])
print('Sample data', data[:2])
# del words # Hint to reduce memory.
```
## Tensorflow Model
```
skip_window = 3
instances = 0
# Pad sentence with skip_windows
for i in range(len(data)):
data[i] = [vocabulary_size]*skip_window+data[i]+[vocabulary_size]*skip_window
# Check how many training samples that we get
for sentence in data:
instances += len(sentence)-2*skip_window
print(instances)
sent_len = np.array([len(d) for d in data])
plt.hist(sent_len,100)
plt.show()
context = np.zeros((instances,skip_window*2+1),dtype=np.int32)
labels = np.zeros((instances,1),dtype=np.int32)
doc = np.zeros((instances,1),dtype=np.int32)
k = 0
for doc_id, sentence in enumerate(data):
for i in range(skip_window, len(sentence)-skip_window):
# buffer = sentence[i-skip_window:i+skip_window+1]
# labels[k] = sentence[i]
# del buffer[skip_window]
# context[k] = buffer
# doc[k] = doc_id
# k += 1
context[k] = sentence[i-skip_window:i+skip_window+1] # Get surrounding words
labels[k] = sentence[i] # Get target variable
doc[k] = doc_id
k += 1
context = np.delete(context,skip_window,1) # delete the middle word
shuffle_idx = np.random.permutation(k)
labels = labels[shuffle_idx]
doc = doc[shuffle_idx]
context = context[shuffle_idx]
batch_size = 256
context_window = 2*skip_window
embedding_size = 50 # Dimension of the embedding vector.
softmax_width = embedding_size # +embedding_size2+embedding_size3
num_sampled = 5 # Number of negative examples to sample.
sum_ids = np.repeat(np.arange(batch_size),context_window)
len_docs = len(data)
graph = tf.Graph()
with graph.as_default(): # , tf.device('/cpu:0')
# Input data.
train_word_dataset = tf.placeholder(tf.int32, shape=[batch_size*context_window])
train_doc_dataset = tf.placeholder(tf.int32, shape=[batch_size])
train_labels = tf.placeholder(tf.int32, shape=[batch_size, 1])
segment_ids = tf.constant(sum_ids, dtype=tf.int32)
word_embeddings = tf.Variable(tf.random_uniform([vocabulary_size,embedding_size],-1.0,1.0))
word_embeddings = tf.concat([word_embeddings,tf.zeros((1,embedding_size))],0)
doc_embeddings = tf.Variable(tf.random_uniform([len_docs,embedding_size],-1.0,1.0))
softmax_weights = tf.Variable(tf.truncated_normal([vocabulary_size, softmax_width],
stddev=1.0 / np.sqrt(embedding_size)))
softmax_biases = tf.Variable(tf.zeros([vocabulary_size]))
# Model.
# Look up embeddings for inputs.
embed_words = tf.segment_mean(tf.nn.embedding_lookup(word_embeddings, train_word_dataset),segment_ids)
embed_docs = tf.nn.embedding_lookup(doc_embeddings, train_doc_dataset)
embed = (embed_words+embed_docs)/2.0#+embed_hash+embed_users
# Compute the softmax loss, using a sample of the negative labels each time.
loss = tf.reduce_mean(tf.nn.nce_loss(softmax_weights, softmax_biases, train_labels,
embed, num_sampled, vocabulary_size))
# Optimizer.
optimizer = tf.train.AdagradOptimizer(0.5).minimize(loss)
norm = tf.sqrt(tf.reduce_sum(tf.square(doc_embeddings), 1, keep_dims=True))
normalized_doc_embeddings = doc_embeddings / norm
############################
# Chunk the data to be passed into the tensorflow Model
###########################
data_idx = 0
def generate_batch(batch_size):
global data_idx
if data_idx+batch_size<instances:
batch_labels = labels[data_idx:data_idx+batch_size]
batch_doc_data = doc[data_idx:data_idx+batch_size]
batch_word_data = context[data_idx:data_idx+batch_size]
data_idx += batch_size
else:
overlay = batch_size - (instances-data_idx)
batch_labels = np.vstack([labels[data_idx:instances],labels[:overlay]])
batch_doc_data = np.vstack([doc[data_idx:instances],doc[:overlay]])
batch_word_data = np.vstack([context[data_idx:instances],context[:overlay]])
data_idx = overlay
batch_word_data = np.reshape(batch_word_data,(-1,1))
return batch_labels, batch_word_data, batch_doc_data
num_steps = 1000001
step_delta = int(num_steps/20)
with tf.Session(graph=graph) as session:
tf.global_variables_initializer().run()
print('Initialized')
average_loss = 0
for step in range(num_steps):
batch_labels, batch_word_data, batch_doc_data\
= generate_batch(batch_size)
feed_dict = {train_word_dataset : np.squeeze(batch_word_data),
train_doc_dataset : np.squeeze(batch_doc_data),
train_labels : batch_labels}
_, l = session.run([optimizer, loss], feed_dict=feed_dict)
average_loss += l
if step % step_delta == 0:
if step > 0:
average_loss = average_loss / step_delta
# The average loss is an estimate of the loss over the last 2000 batches.
print('Average loss at step %d: %f' % (step, average_loss))
average_loss = 0
# Get the weights to save for later
# final_doc_embeddings = normalized_doc_embeddings.eval()
final_word_embeddings = word_embeddings.eval()
final_word_embeddings_out = softmax_weights.eval()
final_doc_embeddings = normalized_doc_embeddings.eval()
# rand_doc = np.random.randint(len_docs)
dist = final_doc_embeddings.dot(final_doc_embeddings[rand_doc][:,None])
closest_doc = np.argsort(dist,axis=0)[-4:][::-1]
furthest_doc = np.argsort(dist,axis=0)[0][::-1]
for idx in closest_doc:
print(dist[idx][0][0])
print(dist[furthest_doc][0][0])
plt.hist(dist,100)
plt.show()
sentences[rand_doc]
sentences[closest_doc[1][0]]
sentences[closest_doc[2][0]]
sentences[closest_doc[3][0]]
sentences[furthest_doc[0]]
import matplotlib.pyplot as plt
%matplotlib inline
plt.hist(dist,100)
plt.show()
```
## Gensim Model
```
import gensim
from gensim.models import Doc2Vec
from multiprocessing import cpu_count
cpus = cpu_count()
def read_corpus():
for i,sentence in enumerate(words.split('\n')):
yield gensim.models.doc2vec.TaggedDocument(gensim.utils.simple_preprocess(sentence), [i])
train_corpus = list(read_corpus())
model = Doc2Vec(dm=1, dm_concat=0, size=embedding_size, window=skip_window,
negative=5,hs=0, min_count=5, workers=cpus, iter=2)
model.build_vocab(train_corpus)
%%time
model.train(train_corpus)
closest_doc2 = model.docvecs.most_similar([model.docvecs[rand_doc]],topn=4)
for _, sim in closest_doc2:
print(sim)
sentences[rand_doc]
sentences[closest_doc2[1][0]]
sentences[closest_doc2[2][0]]
sentences[closest_doc2[3][0]]
norm_vec = np.array([vec for vec in model.docvecs])
norm_vec = norm_vec/np.sqrt(np.sum(np.square(norm_vec),axis=1,keepdims=True))
norm_vec[rand_doc].dot(norm_vec[closest_doc2[1][0]])
```
| github_jupyter |
<a href="https://colab.research.google.com/github/yohanesnuwara/geostatistics/blob/main/project_notebooks/EDA_LWD_openhole_volve_well_15_9_F_15.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
from scipy.interpolate import interp1d
plt.style.use("classic")
!pip install lasio
import lasio
import missingno as msno
import plotly.express as px
!git clone https://github.com/yohanesnuwara/geostatistics
```
## LWD data
Copying script from the previous notebook, "EDA_realtime_volve_well_15_9_F-15.ipynb". There are two files, merged into one.
```
# Realtime data
LAS_file1 = "/content/geostatistics/data/volve-realtime/well_15_9_F-15/WL_RAW_CAL-DEN-ELEM-GR-NEU-REMP_MWD_1~LF1~FRM1.las"
LAS_file2 = "/content/geostatistics/data/volve-realtime/well_15_9_F-15/WL_RAW_CAL-DEN-ELEM-GR-NEU-REMP_MWD_2~LF1~FRM1.las"
mwd1 = lasio.read(LAS_file1)
mwd2 = lasio.read(LAS_file2)
# Convert LAS to dataframe
mwd1_df = mwd1.df().reset_index()
mwd2_df = mwd2.df().reset_index()
# Selected LWD logs. Special here is Bit size (BS_ARC)
mwd_lognames = ["TDEP", "GRMA", "BCAV", "WCLA", "WCAR", "WQFM",
"RHOB", "BPHI", "PEF", "BS_ARC"]
# Select subset dataframe
mwd1_df = mwd1_df[mwd_lognames]
mwd2_df = mwd2_df[mwd_lognames]
# Merging dataframes
mwd_df = pd.concat([mwd1_df, mwd2_df], axis=0)
# Converting ft to m
mwd_df["TDEP"] = mwd_df["TDEP"] / 3.281
# Converting NPHI in % to v/v
mwd_df["BPHI"] = mwd_df["BPHI"] / 100
mwd_df.head()
mwd_df.describe()
```
## Openhole data
```
# Open hole data
LAS_file3 = "/content/geostatistics/data/volve-openhole/well_15_9_F-15/WLC_PETRO_COMPUTED_INPUT_1~LF1~FRM1.las"
oh = lasio.read(LAS_file3)
oh_df = oh.df().reset_index()
# Converting ft to m
oh_df["DEPTH"] = oh_df["DEPTH"] / 3.281
oh_df.head()
oh_df.describe()
```
## Merge Openhole to LWD data by interpolation
I have shown interpolation method in the previous notebook. I'll now use Scipy cubic interpolation that's found to be more accurate than Numpy linear interpolation.
```
def interpolate(df_data, df_new, x_data, y_data, x_new):
xd, yd = df_data[x_data], df_data[y_data]
f = interp1d(xd, yd)
xn = df_new[x_new]
return f(xn)
lognames_data = ["CALI", "DT", "GR", "NPHI", "PEF", "RD", "RHOB"]
lognames_new = ["CALI_OH", "DT_OH", "GR_OH", "NPHI_OH", "PEF_OH", "RD_OH", "RHOB_OH"]
new_logs = []
for i in lognames_data:
_ = interpolate(oh_df, mwd_df, x_data="DEPTH", y_data=i, x_new="TDEP")
new_logs.append(_)
# Merge the result to LWD dataframe
for i in range(len(lognames_new)):
mwd_df[lognames_new[i]] = new_logs[i]
mwd_df.head()
msno.matrix(mwd_df)
plt.show()
```
## Scatter plot between openhole and LWD
I make a scatter plot below to compare measurements between openhole and LWD. As we see below, only GR fits perfectly on the 1:1 line.
```
# LWD
lwd_rec_name = ["GRMA", "RHOB", "PEF", "BPHI"]
lwd_recs = []
for i in lwd_rec_name:
_ = mwd_df[i].values
lwd_recs.append(_)
# Openhole
oh_rec_name = ["GR_OH", "RHOB_OH", "PEF_OH", "NPHI_OH"]
oh_recs = []
for i in oh_rec_name:
_ = mwd_df[i].values
oh_recs.append(_)
# Scatter plot
plt.figure(figsize=(8,8))
xlabels = ["GR Openhole", "RHOB Openhole", "PEF Openhole", "NPHI Openhole"]
ylabels = ["GR LWD", "RHOB LWD", "PEF LWD", "NPHI LWD"]
for i in range(len(lwd_rec_name)):
plt.subplot(2,2,i+1)
plt.scatter(lwd_recs[i], oh_recs[i])
# Plot 1:1 line
x = [0, max(oh_recs[i])]
plt.plot(x, x, '--', color='red')
plt.xlabel(xlabels[i])
plt.ylabel(ylabels[i])
plt.tight_layout(1.3)
plt.show()
```
## Adding formation tops and facies
## Percent clay, quartz, and carbonates
Actually there are not only WCLA (clay), WCAR (carb), & WQFM (qtz), but also siderite, pyrite, anhydrite, etc. So, not all WCLA+WCAR+WQFM sums up to 1.
To assume a composition of ONLY clay, carb, & qtz, I need to make correction, e.g. for clay: %WCLA / (%WCLA + %WCAR + %WQFM) * 100%
```
# Correction for WCLA, WCAR, WQM to sum up 1
xsum = mwd_df["WCLA"] + mwd_df["WCAR"] + mwd_df["WQFM"]
mwd_df["XCLA"] = mwd_df["WCLA"] / xsum
mwd_df["XCAR"] = mwd_df["WCAR"] / xsum
mwd_df["XQFM"] = mwd_df["WQFM"] / xsum
# Ternary plot
fig = px.scatter_ternary(mwd_df, a="XCLA", b="XCAR", c="XQFM", color="TDEP")
fig.show()
```
| github_jupyter |
# Clustering via $k$-means
We previously studied the classification problem using the logistic regression algorithm. Since we had labels for each data point, we may regard the problem as one of _supervised learning_. However, in many applications, the data have no labels but we wish to discover possible labels (or other hidden patterns or structures). This problem is one of _unsupervised learning_. How can we approach such problems?
**Clustering** is one class of unsupervised learning methods. In this lab, we'll consider the following form of the clustering task. Suppose you are given
- a set of observations, $X \equiv \{\hat{x}_i \,|\, 0 \leq i < n\}$, and
- a target number of _clusters_, $k$.
Your goal is to partition the points into $k$ subsets, $C_0,\dots, C_{k-1} \subseteq X$, which are
- disjoint, i.e., $i \neq j \implies C_i \cap C_j = \emptyset$;
- but also complete, i.e., $C_0 \cup C_1 \cup \cdots \cup C_{k-1} = X$.
Intuitively, each cluster should reflect some "sensible" grouping. Thus, we need to specify what constitutes such a grouping.
## Setup: Dataset
The following cell will download the data you'll need for this lab. Run it now.
```
import requests
import os
import hashlib
import io
def on_vocareum():
return os.path.exists('.voc')
def download(file, local_dir="", url_base=None, checksum=None):
local_file = "{}{}".format(local_dir, file)
if not os.path.exists(local_file):
if url_base is None:
url_base = "https://cse6040.gatech.edu/datasets/"
url = "{}{}".format(url_base, file)
print("Downloading: {} ...".format(url))
r = requests.get(url)
with open(local_file, 'wb') as f:
f.write(r.content)
if checksum is not None:
with io.open(local_file, 'rb') as f:
body = f.read()
body_checksum = hashlib.md5(body).hexdigest()
assert body_checksum == checksum, \
"Downloaded file '{}' has incorrect checksum: '{}' instead of '{}'".format(local_file,
body_checksum,
checksum)
print("'{}' is ready!".format(file))
if on_vocareum():
URL_BASE = "https://cse6040.gatech.edu/datasets/kmeans/"
DATA_PATH = "../resource/asnlib/publicdata/"
else:
URL_BASE = "https://github.com/cse6040/labs-fa17/raw/master/datasets/kmeans/"
DATA_PATH = ""
datasets = {'logreg_points_train.csv': '9d1e42f49a719da43113678732491c6d',
'centers_initial_testing.npy': '8884b4af540c1d5119e6e8980da43f04',
'compute_d2_soln.npy': '980fe348b6cba23cb81ddf703494fb4c',
'y_test3.npy': 'df322037ea9c523564a5018ea0a70fbf',
'centers_test3_soln.npy': '0c594b28e512a532a2ef4201535868b5',
'assign_cluster_labels_S.npy': '37e464f2b79dc1d59f5ec31eaefe4161',
'assign_cluster_labels_soln.npy': 'fc0e084ac000f30948946d097ed85ebc'}
for filename, checksum in datasets.items():
download(filename, local_dir=DATA_PATH, url_base=URL_BASE, checksum=checksum)
print("\n(All data appears to be ready.)")
```
## The $k$-means clustering criterion
Here is one way to measure the quality of a set of clusters. For each cluster $C$, consider its center $\mu$ and measure the distance $\|x-\mu\|$ of each observation $x \in C$ to the center. Add these up for all points in the cluster; call this sum is the _within-cluster sum-of-squares (WCSS)_. Then, set as our goal to choose clusters that minimize the total WCSS over _all_ clusters.
More formally, given a clustering $C = \{C_0, C_1, \ldots, C_{k-1}\}$, let
$$
\mathrm{WCSS}(C) \equiv \sum_{i=0}^{k-1} \sum_{x\in C_i} \|x - \mu_i\|^2,
$$
where $\mu_i$ is the center of $C_i$. This center may be computed simply as the mean of all points in $C_i$, i.e.,
$$
\mu_i \equiv \dfrac{1}{|C_i|} \sum_{x \in C_i} x.
$$
Then, our objective is to find the "best" clustering, $C_*$, which is the one that has a minimum WCSS.
$$
C_* = \arg\min_C \mathrm{WCSS}(C).
$$
## The standard $k$-means algorithm (Lloyd's algorithm)
Finding the global optimum is [NP-hard](https://en.wikipedia.org/wiki/NP-hardness), which is computer science mumbo jumbo for "we don't know whether there is an algorithm to calculate the exact answer in fewer steps than exponential in the size of the input." Nevertheless, there is an iterative method, Lloyd’s algorithm, that can quickly converge to a _local_ (as opposed to _global_) minimum. The procedure alternates between two operations: _assignment_ and _update_.
**Step 1: Assignment.** Given a fixed set of $k$ centers, assign each point to the nearest center:
$$
C_i = \{\hat{x}: \| \hat{x} - \mu_i \| \le \| \hat{x} - \mu_j \|, 1 \le j \le k \}.
$$
**Step 2: Update.** Recompute the $k$ centers ("centroids") by averaging all the data points belonging to each cluster, i.e., taking their mean:
$$
\mu_i = \dfrac{1}{|C_i|} \sum_{\hat{x} \in C_i} \hat{x}
$$

> Figure adapted from: http://stanford.edu/~cpiech/cs221/img/kmeansViz.png
In the code that follows, it will be convenient to use our usual "data matrix" convention, that is, each row of a data matrix $X$ is one of $m$ observations and each column (coordinate) is one of $d$ predictors. However, we will _not_ need a dummy column of ones since we are not fitting a function.
$$
X
\equiv \left(\begin{array}{c} \hat{x}_0^T \\ \vdots \\ \hat{x}_{m}^T \end{array}\right)
= \left(\begin{array}{ccc} x_0 & \cdots & x_{d-1} \end{array}\right).
$$
```
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
%matplotlib inline
import matplotlib as mpl
mpl.rc("savefig", dpi=100) # Adjust for higher-resolution figures
```
We will use the following data set which some of you may have seen previously.
```
df = pd.read_csv('{}logreg_points_train.csv'.format(DATA_PATH))
df.head()
# Helper functions from Logistic Regression Lesson
def make_scatter_plot(df, x="x_1", y="x_2", hue="label",
palette={0: "red", 1: "olive"},
size=5,
centers=None):
sns.lmplot(x=x, y=y, hue=hue, data=df, palette=palette,
fit_reg=False)
if centers is not None:
plt.scatter(centers[:,0], centers[:,1],
marker=u'*', s=500,
c=[palette[0], palette[1]])
def mark_matches(a, b, exact=False):
"""
Given two Numpy arrays of {0, 1} labels, returns a new boolean
array indicating at which locations the input arrays have the
same label (i.e., the corresponding entry is True).
This function can consider "inexact" matches. That is, if `exact`
is False, then the function will assume the {0, 1} labels may be
regarded as the same up to a swapping of the labels. This feature
allows
a == [0, 0, 1, 1, 0, 1, 1]
b == [1, 1, 0, 0, 1, 0, 0]
to be regarded as equal. (That is, use `exact=False` when you
only care about "relative" labeling.)
"""
assert a.shape == b.shape
a_int = a.astype(dtype=int)
b_int = b.astype(dtype=int)
all_axes = tuple(range(len(a.shape)))
assert ((a_int == 0) | (a_int == 1)).all()
assert ((b_int == 0) | (b_int == 1)).all()
exact_matches = (a_int == b_int)
if exact:
return exact_matches
assert exact == False
num_exact_matches = np.sum(exact_matches)
if (2*num_exact_matches) >= np.prod (a.shape):
return exact_matches
return exact_matches == False # Invert
def count_matches(a, b, exact=False):
"""
Given two sets of {0, 1} labels, returns the number of mismatches.
This function can consider "inexact" matches. That is, if `exact`
is False, then the function will assume the {0, 1} labels may be
regarded as similar up to a swapping of the labels. This feature
allows
a == [0, 0, 1, 1, 0, 1, 1]
b == [1, 1, 0, 0, 1, 0, 0]
to be regarded as equal. (That is, use `exact=False` when you
only care about "relative" labeling.)
"""
matches = mark_matches(a, b, exact=exact)
return np.sum(matches)
make_scatter_plot(df)
```
Let's extract the data points as a data matrix, `points`, and the labels as a vector, `labels`. Note that the k-means algorithm you will implement should **not** reference `labels` -- that's the solution we will try to predict given only the point coordinates (`points`) and target number of clusters (`k`).
```
points = df[['x_1', 'x_2']].values
labels = df['label'].values
n, d = points.shape
k = 2
```
Note that the labels should _not_ be used in the $k$-means algorithm. We use them here only as ground truth for later verification.
### How to start? Initializing the $k$ centers
To start the algorithm, you need an initial guess. Let's randomly choose $k$ observations from the data.
**Exercise 1** (2 points). Complete the following function, `init_centers(X, k)`, so that it randomly selects $k$ of the given observations to serve as centers. It should return a Numpy array of size `k`-by-`d`, where `d` is the number of columns of `X`.
```
def init_centers(X, k):
"""
Randomly samples k observations from X as centers.
Returns these centers as a (k x d) numpy array.
"""
from numpy.random import choice
samples = choice(len(X), size=k, replace=False)
return X[samples, :]
# Test cell: `init_centers_test`
centers_initial = init_centers(points, k)
print("Initial centers:\n", centers_initial)
assert type(centers_initial) is np.ndarray, "Your function should return a Numpy array instead of a {}".format(type(centers_initial))
assert centers_initial.shape == (k, d), "Returned centers do not have the right shape ({} x {})".format(k, d)
assert (sum(centers_initial[0, :] == points) == [1, 1]).all(), "The centers must come from the input."
assert (sum(centers_initial[1, :] == points) == [1, 1]).all(), "The centers must come from the input."
print("\n(Passed!)")
```
### Computing the distances
**Exercise 2** (3 points). Implement a function that computes a distance matrix, $S = (s_{ij})$ such that $s_{ij} = d_{ij}^2$ is the _squared_ distance from point $\hat{x}_i$ to center $\mu_j$. It should return a Numpy matrix `S[:m, :k]`.
```
def compute_d2(X, centers):
m = len(X)
k = len(centers)
S = np.empty((m, k))
return np.linalg.norm(X[:, np.newaxis, :] - centers, ord=2, axis=2) ** 2
# Test cell: `compute_d2_test`
centers_initial_testing = np.load("{}centers_initial_testing.npy".format(DATA_PATH))
compute_d2_soln = np.load("{}compute_d2_soln.npy".format(DATA_PATH))
S = compute_d2 (points, centers_initial_testing)
assert (np.linalg.norm (S - compute_d2_soln, axis=1) <= (20.0 * np.finfo(float).eps)).all ()
print("\n(Passed!)")
```
**Exercise 3** (2 points). Write a function that uses the (squared) distance matrix to assign a "cluster label" to each point.
That is, consider the $m \times k$ squared distance matrix $S$. For each point $i$, if $s_{i,j}$ is the minimum squared distance for point $i$, then the index $j$ is $i$'s cluster label. In other words, your function should return a (column) vector $y$ of length $m$ such that
$$
y_i = \underset{j \in \{0, \ldots, k-1\}}{\operatorname{argmin}} s_{ij}.
$$
> Hint: Judicious use of Numpy's [`argmin()`](https://docs.scipy.org/doc/numpy/reference/generated/numpy.argmin.html) makes for a nice one-line solution.
```
def assign_cluster_labels(S):
return np.argmin(S, axis=1)
# Cluster labels: 0 1
S_test1 = np.array([[0.3, 0.2], # --> cluster 1
[0.1, 0.5], # --> cluster 0
[0.4, 0.2]]) # --> cluster 1
y_test1 = assign_cluster_labels(S_test1)
print("You found:", y_test1)
assert (y_test1 == np.array([1, 0, 1])).all()
# Test cell: `assign_cluster_labels_test`
S_test2 = np.load("{}assign_cluster_labels_S.npy".format(DATA_PATH))
y_test2_soln = np.load("{}assign_cluster_labels_soln.npy".format(DATA_PATH))
y_test2 = assign_cluster_labels(S_test2)
assert (y_test2 == y_test2_soln).all()
print("\n(Passed!)")
```
**Exercise 4** (2 points). Given a clustering (i.e., a set of points and assignment of labels), compute the center of each cluster.
```
def update_centers(X, y):
# X[:m, :d] == m points, each of dimension d
# y[:m] == cluster labels
m, d = X.shape
k = max(y) + 1
assert m == len(y)
assert (min(y) >= 0)
centers = np.empty((k, d))
for j in range(k):
# Compute the new center of cluster j,
# i.e., centers[j, :d].
centers[j, :d] = np.mean(X[y == j, :], axis=0)
return centers
# Test cell: `update_centers_test`
y_test3 = np.load("{}y_test3.npy".format(DATA_PATH))
centers_test3_soln = np.load("{}centers_test3_soln.npy".format(DATA_PATH))
centers_test3 = update_centers(points, y_test3)
delta_test3 = np.abs(centers_test3 - centers_test3_soln)
assert (delta_test3 <= 2.0*len(centers_test3_soln)*np.finfo(float).eps).all()
print("\n(Passed!)")
```
**Exercise 5** (2 points). Given the squared distances, return the within-cluster sum of squares.
In particular, your function should have the signature,
```python
def WCSS(S):
...
```
where `S` is an array of distances as might be computed from Exercise 2.
For example, suppose `S` is defined as follows:
```python
S = np.array([[0.3, 0.2],
[0.1, 0.5],
[0.4, 0.2]])
```
Then `WCSS(S) == 0.2 + 0.1 + 0.2 == 0.5.`
> _Hint_: See [numpy.amin](https://docs.scipy.org/doc/numpy/reference/generated/numpy.amin.html#numpy.amin).
```
def WCSS(S):
return np.sum(np.amin(S, axis=1))
# Quick test:
print("S ==\n", S_test1)
WCSS_test1 = WCSS(S_test1)
print("\nWCSS(S) ==", WCSS(S_test1))
# Test cell: `WCSS_test`
assert np.abs(WCSS_test1 - 0.5) <= 3.0*np.finfo(float).eps, "WCSS(S_test1) should be close to 0.5, not {}".format(WCSS_test1)
print("\n(Passed!)")
```
Lastly, here is a function to check whether the centers have "moved," given two instances of the center values. It accounts for the fact that the order of centers may have changed.
```
def has_converged(old_centers, centers):
return set([tuple(x) for x in old_centers]) == set([tuple(x) for x in centers])
```
**Exercise 6** (3 points). Put all of the preceding building blocks together to implement Lloyd's $k$-means algorithm.
```
def kmeans(X, k,
starting_centers=None,
max_steps=np.inf):
if starting_centers is None:
centers = init_centers(X, k)
else:
centers = starting_centers
converged = False
labels = np.zeros(len(X))
i = 1
while (not converged) and (i <= max_steps):
old_centers = centers
S = compute_d2(X, centers)
labels = assign_cluster_labels(S)
centers = update_centers(X, labels)
converged = has_converged(old_centers, centers)
print ("iteration", i, "WCSS = ", WCSS (S))
i += 1
return labels
clustering = kmeans(points, k, starting_centers=points[[0, 187], :])
```
Let's visualize the results.
```
# Test cell: `kmeans_test`
df['clustering'] = clustering
centers = update_centers(points, clustering)
make_scatter_plot(df, hue='clustering', centers=centers)
n_matches = count_matches(df['label'], df['clustering'])
print(n_matches,
"matches out of",
len(df), "possible",
"(~ {:.1f}%)".format(100.0 * n_matches / len(df)))
assert n_matches >= 320
```
**Applying k-means to an image.** In this section of the notebook, you will apply k-means to an image, for the purpose of doing a "stylized recoloring" of it. (You can view this example as a primitive form of [artistic style transfer](http://genekogan.com/works/style-transfer/), which state-of-the-art methods today [accomplish using neural networks](https://medium.com/artists-and-machine-intelligence/neural-artistic-style-transfer-a-comprehensive-look-f54d8649c199).)
In particlar, let's take an input image and cluster pixels based on the similarity of their colors. Maybe it can become the basis of your own [Instagram filter](https://blog.hubspot.com/marketing/instagram-filters)!
```
from PIL import Image
from matplotlib.pyplot import imshow
%matplotlib inline
def read_img(path):
"""
Read image and store it as an array, given the image path.
Returns the 3 dimensional image array.
"""
img = Image.open(path)
img_arr = np.array(img, dtype='int32')
img.close()
return img_arr
def display_image(arr):
"""
display the image
input : 3 dimensional array
"""
arr = arr.astype(dtype='uint8')
img = Image.fromarray(arr, 'RGB')
imshow(np.asarray(img))
img_arr = read_img("../resource/asnlib/publicdata/football.bmp")
display_image(img_arr)
print("Shape of the matrix obtained by reading the image")
print(img_arr.shape)
```
Note that the image is stored as a "3-D" matrix. It is important to understand how matrices help to store a image. Each pixel corresponds to a intensity value for Red, Green and Blue. If you note the properties of the image, its resolution is 620 x 412. The image width is 620 pixels and height is 412 pixels, and each pixel has three values - **R**, **G**, **B**. This makes it a 412 x 620 x 3 matrix.
**Exercise 7** (1 point). Write some code to *reshape* the matrix into "img_reshaped" by transforming "img_arr" from a "3-D" matrix to a flattened "2-D" matrix which has 3 columns corresponding to the RGB values for each pixel. In this form, the flattened matrix must contain all pixels and their corresponding RGB intensity values. Remember in the previous modules we had discussed a C type indexing style and a Fortran type indexing style. In this problem, refer to the C type indexing style. The numpy reshape function may be of help here.
```
r, c, l = img_arr.shape
img_reshaped = np.reshape(img_arr, (r*c, l), order="C")
# Test cell - 'reshape_test'
r, c, l = img_arr.shape
# The reshaped image is a flattened '2-dimensional' matrix
assert len(img_reshaped.shape) == 2
r_reshaped, c_reshaped = img_reshaped.shape
assert r * c * l == r_reshaped * c_reshaped
assert c_reshaped == 3
print("Passed")
```
**Exercise 8** (1 point). Now use the k-means function that you wrote above to divide the image in **3** clusters. The result would be a vector named labels, which assigns the label to each pixel.
```
labels = kmeans(img_reshaped, 3)
# Test cell - 'labels'
assert len(labels) == r_reshaped
assert set(labels) == {0, 1, 2}
print("\nPassed!")
```
**Exercise 9** (2 points). Write code to calculate the mean of each cluster and store it in a dictionary, named centers, as label:array(cluster_center). For 3 clusters, the dictionary should have three keys as the labels and their corresponding cluster centers as values, i.e. {0:array(center0), 1: array(center1), 2:array(center2)}.
```
ind = np.column_stack((img_reshaped, labels))
centers = {}
for i in set(labels):
c = ind[ind[:,3] == i].mean(axis=0)
centers[i] = c[:3]
print("Free points here! But you need to implement the above section correctly for you to see what we want you to see later.")
print("\nPassed!")
```
Below, we have written code to generate a matrix "img_clustered" of the same dimensions as img_reshaped, where each pixel is replaced by the cluster center to which it belongs.
```
img_clustered = np.array([centers[i] for i in labels])
```
Let us display the clustered image and see how kmeans works on the image.
```
r, c, l = img_arr.shape
img_disp = np.reshape(img_clustered, (r, c, l), order="C")
display_image(img_disp)
```
You can visually inspect the original image and the clustered image to get a sense of what kmeans is doing here. You can also try to vary the number of clusters to see how the output image changes
## Built-in $k$-means
The preceding exercises walked you through how to implement $k$-means, but as you might have imagined, there are existing implementations as well! The following shows you how to use Scipy's implementation, which should yield similar results. If you are asked to use $k$-means in a future lab (or exam!), you can use this one.
```
from scipy.cluster import vq
# `distortion` below is the similar to WCSS.
# It is called distortion in the Scipy documentation
# since clustering can be used in compression.
k = 2
centers_vq, distortion_vq = vq.kmeans(points, k)
# vq return the clustering (assignment of group for each point)
# based on the centers obtained by the kmeans function.
# _ here means ignore the second return value
clustering_vq, _ = vq.vq(points, centers_vq)
print("Centers:\n", centers_vq)
print("\nCompare with your method:\n", centers, "\n")
print("Distortion (WCSS):", distortion_vq)
df['clustering_vq'] = clustering_vq
make_scatter_plot(df, hue='clustering_vq', centers=centers_vq)
n_matches_vq = count_matches(df['label'], df['clustering_vq'])
print(n_matches_vq,
"matches out of",
len(df), "possible",
"(~ {:.1f}%)".format(100.0 * n_matches_vq / len(df)))
```
**Fin!** That marks the end of this notebook. Don't forget to submit it!
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
import matplotlib
%matplotlib inline
num_imgs = 50000
img_size = 16
min_rect_size = 3
max_rect_size = 8
num_objects = 2
bboxes = np.zeros((num_imgs, num_objects, 4))
imgs = np.zeros((num_imgs, img_size, img_size))
shapes = np.zeros((num_imgs, num_objects, 1))
for i_img in range(num_imgs):
for i_object in range(num_objects):
if np.random.choice([True, False]):
width, height = np.random.randint(min_rect_size, max_rect_size, size=2)
x = np.random.randint(0, img_size - width)
y = np.random.randint(0, img_size - height)
imgs[i_img, x:x+width, y:y+height] = 1.
bboxes[i_img, i_object] = [x, y, width, height]
shapes[i_img, i_object] = [0]
else:
size = np.random.randint(min_rect_size, max_rect_size)
x, y = np.random.randint(0, img_size - size, size=2)
mask = np.tril_indices(size)
imgs[i_img, x + mask[0], y + mask[1]] = 1.
bboxes[i_img, i_object] = [x, y, size, size]
shapes[i_img, i_object] = [1]
imgs.shape, bboxes.shape
i = 0
# TODO: Why does the array have to be transposed?
plt.imshow(imgs[i].T, cmap='Greys', interpolation='none', origin='lower', extent=[0, img_size, 0, img_size])
for bbox, shape in zip(bboxes[i], shapes[i]):
plt.gca().add_patch(matplotlib.patches.Rectangle((bbox[0], bbox[1]), bbox[2], bbox[3], ec='r' if shape[0] == 0 else 'y', fc='none'))
X = (imgs.reshape(num_imgs, -1) - np.mean(imgs)) / np.std(imgs)
X.shape, np.mean(X), np.std(X)
# TODO: We use binary classification here - for multiple classes, convert classes to one-hot vectors.
y = np.concatenate([bboxes / img_size, shapes], axis=-1).reshape(num_imgs, -1)
y.shape
i = int(0.8 * num_imgs)
train_X = X[:i]
test_X = X[i:]
train_y = y[:i]
test_y = y[i:]
test_imgs = imgs[i:]
test_bboxes = bboxes[i:]
from keras.models import Sequential
from keras.layers import Dense, Activation, Dropout
from keras.optimizers import SGD
model = Sequential([
Dense(256, input_dim=X.shape[-1]),
Activation('relu'),
Dropout(0.4),
Dense(y.shape[-1])
])
model.compile('adadelta', 'mse')
# Flip bboxes during training.
# Note: The validation loss is always quite big here because we don't flip the bounding boxes for the validation data.
def IOU(bbox1, bbox2):
'''Calculate overlap between two bounding boxes [x, y, w, h] as the area of intersection over the area of unity'''
x1, y1, w1, h1 = bbox1[0], bbox1[1], bbox1[2], bbox1[3] # TODO: Check if its more performant if tensor elements are accessed directly below.
x2, y2, w2, h2 = bbox2[0], bbox2[1], bbox2[2], bbox2[3]
w_I = min(x1 + w1, x2 + w2) - max(x1, x2)
h_I = min(y1 + h1, y2 + h2) - max(y1, y2)
w_I = max(w_I, 0) # set w_I and h_I zero if there is no intersection
h_I = max(h_I, 0)
I = w_I * h_I
U = w1 * h1 + w2 * h2 - I
return I / U
def dist(bbox1, bbox2):
return np.sqrt(np.sum(np.square(bbox1[:2] - bbox2[:2])))
num_epochs = 50
flipped_train_y = np.array(train_y)
flipped = np.zeros((len(flipped_train_y), num_epochs))
ious = np.zeros((len(flipped_train_y), num_epochs))
dists = np.zeros((len(flipped_train_y), num_epochs))
for epoch in range(num_epochs):
print 'Epoch', epoch
model.fit(train_X, flipped_train_y, nb_epoch=1, validation_data=(test_X, test_y), verbose=2)
pred_y = model.predict(train_X)
for i, (pred_bboxes, exp_bboxes) in enumerate(zip(pred_y, flipped_train_y)):
flipped_exp_bboxes = np.concatenate([exp_bboxes[4:], exp_bboxes[:4]])
mse = np.mean(np.square(pred_bboxes - exp_bboxes))
mse_flipped = np.mean(np.square(pred_bboxes - flipped_exp_bboxes))
iou = IOU(pred_bboxes[:4], exp_bboxes[:4]) + IOU(pred_bboxes[4:], exp_bboxes[4:])
iou_flipped = IOU(pred_bboxes[:4], flipped_exp_bboxes[:4]) + IOU(pred_bboxes[4:], flipped_exp_bboxes[4:])
dist = IOU(pred_bboxes[:4], exp_bboxes[:4]) + IOU(pred_bboxes[4:], exp_bboxes[4:])
dist_flipped = IOU(pred_bboxes[:4], flipped_exp_bboxes[:4]) + IOU(pred_bboxes[4:], flipped_exp_bboxes[4:])
if mse_flipped < mse: # using iou or dist here leads to similar results
flipped_train_y[i] = flipped_exp_bboxes
flipped[i, epoch] = 1
ious[i, epoch] = iou_flipped / 2.
dists[i, epoch] = dist_flipped / 2.
else:
ious[i, epoch] = iou / 2.
dists[i, epoch] = dist / 2.
print 'Flipped {} training samples ({} %)'.format(np.sum(flipped[:, epoch]), np.mean(flipped[:, epoch]) * 100.)
print 'Mean IOU: {}'.format(np.mean(ious[:, epoch]))
print 'Mean dist: {}'.format(np.mean(dists[:, epoch]))
print
plt.pcolor(flipped[:1000], cmap='Greys')
plt.xlabel('Epoch')
plt.ylabel('Training sample')
plt.plot(np.mean(ious, axis=0), label='Mean IOU') # between predicted and assigned true bboxes
plt.plot(np.mean(dists, axis=0), label='Mean distance') # relative to image size
plt.legend()
plt.ylim(0, 1)
pred_y = model.predict(test_X)
pred_y = pred_y.reshape(len(pred_y), num_objects, -1)
pred_bboxes = pred_y[..., :4] * img_size
pred_shapes = pred_y[..., 4:5]
pred_bboxes.shape, pred_shapes.shape
plt.figure(figsize=(16, 8))
for i_subplot in range(1, 5):
plt.subplot(1, 4, i_subplot)
i = np.random.randint(len(test_X))
plt.imshow(test_imgs[i].T, cmap='Greys', interpolation='none', origin='lower', extent=[0, img_size, 0, img_size])
for pred_bbox, exp_bbox, pred_shape in zip(pred_bboxes[i], test_bboxes[i], pred_shapes[i]):
plt.gca().add_patch(matplotlib.patches.Rectangle((pred_bbox[0], pred_bbox[1]), pred_bbox[2], pred_bbox[3], ec='r' if pred_shape[0] <= 0.5 else 'y', fc='none'))
# TODO: Calculate max IOU with all expected bounding boxes.
# plt.annotate('IOU: {:.2f}'.format(IOU(pred_bbox, exp_bbox)), (pred_bbox[0], pred_bbox[1]+pred_bbox[3]+0.4), color='r')
# plt.savefig('plots/bw-two-rectangles-or-triangles4.png', dpi=300)
```
| github_jupyter |
# Introduction
We need to convert the file type and reproject our Snow Water Equivalent data into the format accepted by Google Earth Engine.
Google Earth Engine accepts .tifs in WGS 83 (EPSG: 4326).
```
# Imports
from glob import glob
import matplotlib.pyplot as plt
import numpy as np
import os
from os.path import exists, join
import pandas as pd
from PIL import Image
import re
import subprocess
import urllib
import webbrowser
import xarray as xr
# Silence warnings
import warnings
warnings.simplefilter(action='ignore', category=FutureWarning)
```
# EASE Grids Map Projection to WGS84
The GlobSnow data product is produced by the European Space Agency as a HDF4 file. Plotting the SWE from the original file using xarray shows a Northern Hemisphere, Lambert Azimuthal projection.
```
# Take a look at one of the .hdf files
f = "../data/01/GlobSnow_SWE_L3B_monthly_201601_v2.0.hdf"
data = xr.open_dataset(f)
data.swe_average.plot()
```
We would like to reproject this to WGS84. Unfortunately, if we look at the xarray dataset we see there are no latitude, longitude data variables. The metadata concerning the projection is missing. Without this projection information, GDAL does not know how to map the pixel coordinates to latitude and longitude.
```
print(data)
```
### Programmatically obtain projection data
The parameters of the grid can be found at the [National Snow & Ice Data Center](https://nsidc.org/ease/ease-grid-projection-gt). We wrote a class to obtain the projection data from the data provided on the NSIDC website. The EPSG and map coordinates of pixel corners must be manually provided to the GDAL translate utility.
```
class EASE_Parameters(object):
def __init__(self, version=1, hemisphere='Northern'):
'''
Retrieve CRS information from URL.
:param version: The version number (1 or 2) of the grid
:param hemisphere: The hemisphere of interest (Northern or Southern)
'''
# Set the grid name using the supplied version number
self.version = version
grid_name = {
1: 'EASE-Grid',
2: 'EASE-Grid 2.0'
}
# Validate user input
try:
self.grid = grid_name[self.version]
except KeyError:
raise UserWarning('You must choose between version 1 and version 2.')
# Validate the hemisphere input
self.hemisphere = hemisphere
if self.hemisphere.lower() not in ['northern', 'southern']:
raise UserWarning('You must chose between the Northern and Southern hemisphere.')
self.url = "https://nsidc.org/ease/ease-grid-projection-gt"
self._table_number = -2
@property
def table_number(self):
return self._table_number
@table_number.setter
def table_number(self, n):
self._table_number = n
@property
def table(self):
'''
Return the dataframe for the grid and resolution of interest.
Rather than pass resolution as a parameter, select the table number from
'''
tables = pd.read_html(self.url)
num_tables = len(tables) -1
if abs(self._table_number) > num_tables:
raise UserWarning(f'There are only {num_tables} tables, please select a valid table number.')
df = tables[self._table_number]
# The Grid Name column name varies over tables, use column number
return df[(df[df.columns[0]]==self.grid) & (df['Projection'].str.contains(self.hemisphere, case=False))]
@property
def epsg(self):
'''
Return the EPSG code corresponding to the CRS.
'''
proj = self.table['Projection'].item()
pattern = re.compile('EPSG: \d{4}')
epsg = re.search(pattern, proj)[0]
return epsg.replace(' ', '')
@property
def num_cols(self):
'''
Return the number of columns in the grid.
'''
return self.table['Number of Columns'].item()
@property
def num_rows(self):
'''
Return the number of rows in teh grid.
'''
return self.table['Number of Rows'].item()
@property
def grid_size(self):
'''
Grid cell area is reported as a string {number} {unit} x {number} {unit}.
Parse the string and take the first number as the size
'''
return float(self.table['Grid Cell Area'].item().split(' ')[0].replace(',',''))
# TODO: Debug key error
@property
def ulx(self):
try:
return self.table['x-axis map coordinate of the outer edge of the upper-left pixel'].item()
except KeyError:
return self.table[self.table.columns[-2]].item()
@property
def uly(self):
try:
return self.table['y-axis map coordinate of the outer edge of the upper-left pixel'].item()
except KeyError:
return self.table[self.table.columns[-1]].item()
@property
def lrx(self):
'''
Return the lower right x coord by multiplying the grid size by the number of columns.
'''
return self.ulx + self.grid_size*self.num_cols
@property
def lry(self):
'''
Return the lower right x coord by multiplying the grid size by the number of rows.
'''
return self.uly - self.grid_size*self.num_rows
```
### Convert HDF file to TIF file
When GDAL is given the projection information, it can be used to convert to a GeoTiff file.
```
def download_globsnow(directory):
'''
Download all available data.
'''
for year in range(1997, 2017):
for month in range(1, 13):
mon = "{0:0=2d}".format(month)
filename = f"GlobSnow_SWE_L3B_monthly_{year}{mon}_v2.0.hdf"
folder = join(directory, mon)
if not exists(folder):
os.makedirs(folder)
if not exists(join(folder, filename)):
url = f"http://www.globsnow.info/swe/archive_v2.0/{year}/L3B_monthly_SWE_HDF/{filename}"
try:
urllib.request.urlretrieve(url, join(folder, filename))
except:
continue
# File conversion function
def translate_globsnow(directory, month):
'''
Add meta data to the raw file and convert to gdal virtual format.
'''
# Obtain CRS information
param = EASE_Parameters()
epsg = param.epsg
ulx, uly, lrx, lry = param.ulx, param.uly, param.lrx, param.lry
# Find files that match this pattern
pattern = re.compile("(GlobSnow_SWE_L3B_monthly_(\d{4})(\d{2})_v2.0.hdf)")
folder = join(directory, month)
for f in glob(join(folder, '*.hdf')):
match = re.search(pattern, f)
if match is not None:
# Create output filename from input filename
input_file, year, month = match.groups()
output_file = f"GlobSnow_SWE_Average_{year}_{month}.vrt"
bash_cmd = (
f"gdal_translate -of VRT -a_nodata -1 "
f"-a_srs {epsg} -a_ullr {ulx} {uly} {lrx} {lry} "
f"HDF4_SDS:UNKNOWN:\"{input_file}\":0 {output_file}"
)
subprocess.Popen(bash_cmd, cwd=folder, shell=True, executable='/bin/bash')
def convert_globsnow(directory, month):
'''
Convert .vrt to a .tif file.
'''
pattern = re.compile("GlobSnow_SWE_Average_\d{4}_\d{2}.vrt")
folder = join(directory, month)
for f in glob(join(folder, '*.vrt')):
match = re.search(pattern, f)
if match is not None:
input_file = match[0]
output_file = input_file.replace('.vrt', '.tif')
bash_cmd = (
f"gdalwarp -of GTiff "
f"-t_srs EPSG:4326 -r cubic "
f"{input_file} {output_file}"
)
subprocess.Popen(bash_cmd, cwd=folder, shell=True, executable='/bin/bash')
def plot(filename):
'''
Plot image.
'''
im = Image.open(filename)
plt.imshow(im)
```
Replotting the data we can see it has been reprojected.
```
# Run code
# download_globsnow("../data")
# for month in ["{0:0=2d}".format(m) for m in range(1,13)]:
# translate_globsnow("../data", month)
# convert_globsnow("../data", month)
plot("../data/01/GlobSnow_SWE_Average_2016_01.tif")
```
### Google Earth Engine
Import tif to Google Earth Engine and include the data as a parameter in classification. [Open GEE Code Editor]('https://code.earthengine.google.com/0eb3b00f6829f6e73f53dea2fe325761')
```
url = 'https://code.earthengine.google.com/0eb3b00f6829f6e73f53dea2fe325761'
webbrowser.open_new(url)
```
| github_jupyter |
# Basic Spark SQL Usage
### Example of using Spark SQL with Stroom DataFrame
#### Prerequisites
This notebook is designed to work with a `stroom-full-test` Stroom stack intalled on `localhost`.
You must set the environmental variable `STROOM_API_KEY` to the API token associated with a suitably privileged Stroom user account before starting the Jupyter notebook server process.
#### Use Java 8
It is necessary to start `pyspark` from a Java 8 shell. Failure to do so will result in errors, including quite mysterious ones relating to missing Hive classes.
#### Setup
Import standard utility classes/functions, including JSON handling XSLT.
```
from pyspark.sql.types import *
from pyspark.sql.functions import from_json, col
from IPython.display import display
from pyspark.sql import SparkSession
```
#### Create a schema using XPaths
N.B. XPath @* is used to extract both StreamId and EventId from the Event, and placed into a single field.
This field has unique values, handy for working with SQL.
```
mySchema = StructType([StructField("user", StringType(), True,
metadata={"get": "EventSource/User/Id"}),
StructField("operation", StringType(), True,
metadata={"get": "EventDetail/TypeId"}),
StructField("eventid", StringType(), False,
metadata={"get": "@*"})])
stroomDf = spark.read.format('stroom.spark.datasource.StroomDataSource').load(
token=os.environ['STROOM_API_KEY'],host='localhost',protocol='http',
uri='api/stroom-index/v2',
index='57a35b9a-083c-4a93-a813-fc3ddfe1ff44', pipeline='26ed1000-255e-4182-b69b-00266be891ee',
schema=mySchema).select('eventid','user','operation','idxUserId')
display(stroomDf.limit(5).toPandas().head())
```
#### Using Spark SQL
In order to start actually writing SQL queries, it is necessary to create a temporary view onto the
Stroom DataFrame created above.
Results are returned as DataFrames themselves, making further operations possible.
```
stroomDf.createOrReplaceTempView("userops")
sqlDf = spark.sql("select * from userops where user = 'admin' and operation='StroomIndexQueryResourceImpl.search'")
display(sqlDf.limit(5).toPandas().head())
sqlDf2 = spark.sql("select user,operation, count (eventid) as events from userops \
where idxUserId != 'admin' group by user, operation \
order by events desc")
display(sqlDf2.toPandas())
```
| github_jupyter |
# Import and format custom datasets for SCHOLAR
```
import os, sys, scipy, json
from scipy import sparse
import codecs
import numpy as np
import pandas as pd
import file_handling as fh
```
# Semi-synthetic data BOOKING
## load data and save in SCHOLAR format
```
if sys.platform == "darwin":
#raw_data_path = "/Users/maximilianahrens/OneDrive - Nexus365/00_datasets/booking/booking_btr/"
output_dir = raw_data_path + "scholar/"
else:
raw_data_path = "/nfs/home/maxa/data/semisynth_btr/"
output_dir = raw_data_path + "scholar/"
print(raw_data_path, "\n",output_dir)
```
## vocab
```
vocab_df = pd.read_csv(raw_data_path +'preprocessed/booking_synth_vocab.csv', header = 0)
vocab_df
vocab = list(vocab_df["x1"])
len(vocab)
fh.write_to_json(vocab, output_dir + "train.vocab.json", indent=2, sort_keys=True)
```
## dtm
```
x_bow_raw = pd.read_csv(raw_data_path + "preprocessed/booking_synth_dtm.csv", header = 0).values
x_bow = np.matrix(x_bow_raw)
x_bow.shape
# insample
sparse_Xtr = sparse.coo_matrix(x_bow).tocsr()
fh.save_sparse(sparse_Xtr, os.path.join(raw_data_path, "scholar", "train.npz"))
```
## doc IDs
```
#length = 50000
#train_ids = ["train_" + str(x) for x in list(range(length))]
# insample
train_ids = ["train_" + str(x) for x in list(range(x_bow.shape[0]))]
fh.write_to_json(train_ids, output_dir + "train.ids.json", indent=2, sort_keys=True)
semisynth_data = pd.read_csv(raw_data_path + "booking_semisynth_sample.csv",header = 0)
semisynth_data.shape
semisynth_data.head(3)
```
## labels
```
train_y = semisynth_data.synth_y
train_y.index = train_ids
train_y.to_csv(output_dir + "train.target.csv")
train_y
```
## covariates
```
train_covars = semisynth_data[["Leisure","av_score"]].astype("float32") # choose which features to include
train_covars.index = train_ids
train_covars.to_csv(output_dir + "train.covars.csv")
train_covars.shape
train_covars
```
# For train-test split
```
output_dir_tt = "/nfs/home/maxa/data/semisynth_btr/scholar_split/"
```
## vocab
```
vocab_df = pd.read_csv(raw_data_path +'preprocessed/booking_synth_vocab.csv', header = 0)
vocab_df
vocab = list(vocab_df["x1"])
len(vocab)
fh.write_to_json(vocab, output_dir_tt + "train.vocab.json", indent=2, sort_keys=True)
```
## dtm
```
x_bow.shape
cut = int(0.8*x_bow.shape[0])
cut
x_bow_train = x_bow[:cut]
x_bow_test = x_bow[cut:]
x_bow_train.shape, x_bow_test.shape
# train
sparse_Xtr = sparse.coo_matrix(x_bow_train).tocsr()
fh.save_sparse(sparse_Xtr, os.path.join(raw_data_path, "scholar_split", "train.npz"))
# test
sparse_Xte = sparse.coo_matrix(x_bow_test).tocsr()
fh.save_sparse(sparse_Xte, os.path.join(raw_data_path, "scholar_split", "test.npz"))
```
## doc IDs
```
train_ids = ["train_" + str(x) for x in list(range(x_bow_train.shape[0]))]
test_ids = ["test_" + str(x) for x in list(range(x_bow_test.shape[0]))]
fh.write_to_json(train_ids, output_dir_tt + "train.ids.json", indent=2, sort_keys=True)
fh.write_to_json(test_ids, output_dir_tt + "test.ids.json", indent=2, sort_keys=True)
```
## labels
```
semisynth_data = pd.read_csv(raw_data_path + "booking_semisynth_sample.csv",header = 0)
semisynth_data.shape
semisynth_data.head(2)
data_y = semisynth_data.synth_y
train_y = data_y[:cut]
test_y = data_y[cut:]
train_y.index = train_ids
train_y.to_csv(output_dir_tt + "train.target.csv")
train_y
test_y.index = test_ids
test_y.to_csv(output_dir_tt + "test.target.csv")
test_y
```
## covariates
```
data_covars = semisynth_data[["Leisure","av_score"]].astype("float32") # choose which features to include
train_covars = data_covars[:cut]
test_covars = data_covars[cut:]
train_covars.index = train_ids
train_covars.to_csv(output_dir_tt + "train.covars.csv")
test_covars.index = test_ids
test_covars.to_csv(output_dir_tt + "test.covars.csv")
train_covars.shape, test_covars.shape
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Dogs vs Cats Image Classification With Image Augmentation
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/examples/blob/master/courses/udacity_intro_to_tensorflow_for_deep_learning/l05c02_dogs_vs_cats_with_augmentation.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/examples/blob/master/courses/udacity_intro_to_tensorflow_for_deep_learning/l05c02_dogs_vs_cats_with_augmentation.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
</table>
In this tutorial, we will discuss how to classify images into pictures of cats or pictures of dogs. We'll build an image classifier using `tf.keras.Sequential` model and load data using `tf.keras.preprocessing.image.ImageDataGenerator`.
## Specific concepts that will be covered:
In the process, we will build practical experience and develop intuition around the following concepts
* Building _data input pipelines_ using the `tf.keras.preprocessing.image.ImageDataGenerator` class — How can we efficiently work with data on disk to interface with our model?
* _Overfitting_ - what is it, how to identify it, and how can we prevent it?
* _Data Augmentation_ and _Dropout_ - Key techniques to fight overfitting in computer vision tasks that we will incorporate into our data pipeline and image classifier model.
## We will follow the general machine learning workflow:
1. Examine and understand data
2. Build an input pipeline
3. Build our model
4. Train our model
5. Test our model
6. Improve our model/Repeat the process
<hr>
**Before you begin**
Before running the code in this notebook, reset the runtime by going to **Runtime -> Reset all runtimes** in the menu above. If you have been working through several notebooks, this will help you avoid reaching Colab's memory limits.
# Importing packages
Let's start by importing required packages:
* os — to read files and directory structure
* numpy — for some matrix math outside of TensorFlow
* matplotlib.pyplot — to plot the graph and display images in our training and validation data
```
from __future__ import absolute_import, division, print_function, unicode_literals
try:
# Use the %tensorflow_version magic if in colab.
%tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
from tensorflow.keras.preprocessing.image import ImageDataGenerator
import os
import numpy as np
import matplotlib.pyplot as plt
```
# Data Loading
To build our image classifier, we begin by downloading the dataset. The dataset we are using is a filtered version of <a href="https://www.kaggle.com/c/dogs-vs-cats/data" target="_blank">Dogs vs. Cats</a> dataset from Kaggle (ultimately, this dataset is provided by Microsoft Research).
In previous Colabs, we've used <a href="https://www.tensorflow.org/datasets" target="_blank">TensorFlow Datasets</a>, which is a very easy and convenient way to use datasets. In this Colab however, we will make use of the class `tf.keras.preprocessing.image.ImageDataGenerator` which will read data from disk. We therefore need to directly download *Dogs vs. Cats* from a URL and unzip it to the Colab filesystem.
```
_URL = 'https://storage.googleapis.com/mledu-datasets/cats_and_dogs_filtered.zip'
zip_dir = tf.keras.utils.get_file('cats_and_dogs_filterted.zip', origin=_URL, extract=True)
```
The dataset we have downloaded has following directory structure.
<pre style="font-size: 10.0pt; font-family: Arial; line-height: 2; letter-spacing: 1.0pt;" >
<b>cats_and_dogs_filtered</b>
|__ <b>train</b>
|______ <b>cats</b>: [cat.0.jpg, cat.1.jpg, cat.2.jpg ....]
|______ <b>dogs</b>: [dog.0.jpg, dog.1.jpg, dog.2.jpg ...]
|__ <b>validation</b>
|______ <b>cats</b>: [cat.2000.jpg, cat.2001.jpg, cat.2002.jpg ....]
|______ <b>dogs</b>: [dog.2000.jpg, dog.2001.jpg, dog.2002.jpg ...]
</pre>
We'll now assign variables with the proper file path for the training and validation sets.
```
base_dir = os.path.join(os.path.dirname(zip_dir), 'cats_and_dogs_filtered')
train_dir = os.path.join(base_dir, 'train')
validation_dir = os.path.join(base_dir, 'validation')
train_cats_dir = os.path.join(train_dir, 'cats') # directory with our training cat pictures
train_dogs_dir = os.path.join(train_dir, 'dogs') # directory with our training dog pictures
validation_cats_dir = os.path.join(validation_dir, 'cats') # directory with our validation cat pictures
validation_dogs_dir = os.path.join(validation_dir, 'dogs') # directory with our validation dog pictures
```
### Understanding our data
Let's look at how many cats and dogs images we have in our training and validation directory
```
num_cats_tr = len(os.listdir(train_cats_dir))
num_dogs_tr = len(os.listdir(train_dogs_dir))
num_cats_val = len(os.listdir(validation_cats_dir))
num_dogs_val = len(os.listdir(validation_dogs_dir))
total_train = num_cats_tr + num_dogs_tr
total_val = num_cats_val + num_dogs_val
print('total training cat images:', num_cats_tr)
print('total training dog images:', num_dogs_tr)
print('total validation cat images:', num_cats_val)
print('total validation dog images:', num_dogs_val)
print("--")
print("Total training images:", total_train)
print("Total validation images:", total_val)
```
# Setting Model Parameters
For convenience, let us set up variables that will be used later while pre-processing our dataset and training our network.
```
BATCH_SIZE = 100
IMG_SHAPE = 150 # Our training data consists of images with width of 150 pixels and height of 150 pixels
```
After defining our generators for training and validation images, **flow_from_directory** method will load images from the disk and will apply rescaling and will resize them into required dimensions using single line of code.
# Data Augmentation
Overfitting often occurs when we have a small number of training examples. One way to fix this problem is to augment our dataset so that it has sufficient number and variety of training examples. Data augmentation takes the approach of generating more training data from existing training samples, by augmenting the samples through random transformations that yield believable-looking images. The goal is that at training time, your model will never see the exact same picture twice. This exposes the model to more aspects of the data, allowing it to generalize better.
In **tf.keras** we can implement this using the same **ImageDataGenerator** class we used before. We can simply pass different transformations we would want to our dataset as a form of arguments and it will take care of applying it to the dataset during our training process.
To start off, let's define a function that can display an image, so we can see the type of augmentation that has been performed. Then, we'll look at specific augmentations that we'll use during training.
```
# This function will plot images in the form of a grid with 1 row and 5 columns where images are placed in each column.
def plotImages(images_arr):
fig, axes = plt.subplots(1, 5, figsize=(20,20))
axes = axes.flatten()
for img, ax in zip(images_arr, axes):
ax.imshow(img)
plt.tight_layout()
plt.show()
```
### Flipping the image horizontally
We can begin by randomly applying horizontal flip augmentation to our dataset and seeing how individual images will look after the transformation. This is achieved by passing `horizontal_flip=True` as an argument to the `ImageDataGenerator` class.
```
image_gen = ImageDataGenerator(rescale=1./255, horizontal_flip=True)
train_data_gen = image_gen.flow_from_directory(batch_size=BATCH_SIZE,
directory=train_dir,
shuffle=True,
target_size=(IMG_SHAPE,IMG_SHAPE))
```
To see the transformation in action, let's take one sample image from our training set and repeat it five times. The augmentation will be randomly applied (or not) to each repetition.
```
augmented_images = [train_data_gen[0][0][0] for i in range(5)]
plotImages(augmented_images)
```
### Rotating the image
The rotation augmentation will randomly rotate the image up to a specified number of degrees. Here, we'll set it to 45.
```
image_gen = ImageDataGenerator(rescale=1./255, rotation_range=45)
train_data_gen = image_gen.flow_from_directory(batch_size=BATCH_SIZE,
directory=train_dir,
shuffle=True,
target_size=(IMG_SHAPE, IMG_SHAPE))
```
To see the transformation in action, let's once again take a sample image from our training set and repeat it. The augmentation will be randomly applied (or not) to each repetition.
```
augmented_images = [train_data_gen[0][0][0] for i in range(5)]
plotImages(augmented_images)
```
### Applying Zoom
We can also apply Zoom augmentation to our dataset, zooming images up to 50% randomly.
```
image_gen = ImageDataGenerator(rescale=1./255, zoom_range=0.5)
train_data_gen = image_gen.flow_from_directory(batch_size=BATCH_SIZE,
directory=train_dir,
shuffle=True,
target_size=(IMG_SHAPE, IMG_SHAPE))
```
One more time, take a sample image from our training set and repeat it. The augmentation will be randomly applied (or not) to each repetition.
```
augmented_images = [train_data_gen[0][0][0] for i in range(5)]
plotImages(augmented_images)
```
### Putting it all together
We can apply all these augmentations, and even others, with just one line of code, by passing the augmentations as arguments with proper values.
Here, we have applied rescale, rotation of 45 degrees, width shift, height shift, horizontal flip, and zoom augmentation to our training images.
```
image_gen_train = ImageDataGenerator(
rescale=1./255,
rotation_range=40,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,
fill_mode='nearest')
train_data_gen = image_gen_train.flow_from_directory(batch_size=BATCH_SIZE,
directory=train_dir,
shuffle=True,
target_size=(IMG_SHAPE,IMG_SHAPE),
class_mode='binary')
```
Let's visualize how a single image would look like five different times, when we pass these augmentations randomly to our dataset.
```
augmented_images = [train_data_gen[0][0][0] for i in range(5)]
plotImages(augmented_images)
```
### Creating Validation Data generator
Generally, we only apply data augmentation to our training examples, since the original images should be representative of what our model needs to manage. So, in this case we are only rescaling our validation images and converting them into batches using ImageDataGenerator.
```
image_gen_val = ImageDataGenerator(rescale=1./255)
val_data_gen = image_gen_val.flow_from_directory(batch_size=BATCH_SIZE,
directory=validation_dir,
target_size=(IMG_SHAPE, IMG_SHAPE),
class_mode='binary')
```
# Model Creation
## Define the model
The model consists of four convolution blocks with a max pool layer in each of them.
Before the final Dense layers, we're also applying a Dropout probability of 0.5. It means that 50% of the values coming into the Dropout layer will be set to zero. This helps to prevent overfitting.
Then we have a fully connected layer with 512 units, with a `relu` activation function. The model will output class probabilities for two classes — dogs and cats — using `softmax`.
```
model = tf.keras.models.Sequential([
tf.keras.layers.Conv2D(32, (3,3), activation='relu', input_shape=(150, 150, 3)),
tf.keras.layers.MaxPooling2D(2, 2),
tf.keras.layers.Conv2D(64, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
tf.keras.layers.Conv2D(128, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
tf.keras.layers.Conv2D(128, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
tf.keras.layers.Dropout(0.5),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(512, activation='relu'),
tf.keras.layers.Dense(2)
])
```
### Compiling the model
As usual, we will use the `adam` optimizer. Since we output a softmax categorization, we'll use `sparse_categorical_crossentropy` as the loss function. We would also like to look at training and validation accuracy on each epoch as we train our network, so we are passing in the metrics argument.
```
model.compile(optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
```
### Model Summary
Let's look at all the layers of our network using **summary** method.
```
model.summary()
```
### Train the model
It's time we train our network.
Since our batches are coming from a generator (`ImageDataGenerator`), we'll use `fit_generator` instead of `fit`.
```
epochs=100
history = model.fit_generator(
train_data_gen,
steps_per_epoch=int(np.ceil(total_train / float(BATCH_SIZE))),
epochs=epochs,
validation_data=val_data_gen,
validation_steps=int(np.ceil(total_val / float(BATCH_SIZE)))
)
```
### Visualizing results of the training
We'll now visualize the results we get after training our network.
```
acc = history.history['accuracy']
val_acc = history.history['val_accuracy']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs_range = range(epochs)
plt.figure(figsize=(8, 8))
plt.subplot(1, 2, 1)
plt.plot(epochs_range, acc, label='Training Accuracy')
plt.plot(epochs_range, val_acc, label='Validation Accuracy')
plt.legend(loc='lower right')
plt.title('Training and Validation Accuracy')
plt.subplot(1, 2, 2)
plt.plot(epochs_range, loss, label='Training Loss')
plt.plot(epochs_range, val_loss, label='Validation Loss')
plt.legend(loc='upper right')
plt.title('Training and Validation Loss')
plt.show()
```
| github_jupyter |
Isolation Forest algorithm works for both univariate and multivariate datasets, and uses unsupervised learning
This algo is ideal for detecting anomalies when you have multiple input features because of its ability to handle multivariate data. However, we are mostly using univariate data (just our univariate timeseries metrics), but we also exercise the multivariate capabilities when we perform Principle Component Analysis on the whole dataset to see when the repo as a whole is in an anomolous state.
In our case, our input features will be various different metrics provided by our own api.
The goal of this algo is to determine when a particular metric or the whole repo is in an anomalous state. This could mean, for example, that we are detecting a spike in issues opened.
BELOW IS EXTRA INFO THAT EXPLAINS MORE ABOUT THE ALGORITHM IF YOU ARE INTERESTED:
It works similar to Decision trees algorithm, where we start with a root node and keep on partitioning the space. In Isolation forest we partition randomly, unlike Decision trees where the partition is based on information gain.
Partitions are created by randomly selecting a feature and then randomly creating a split value between the maximum and the minimum value of the feature. We keep on creating the partitions until we isolate all the points(in most cases we also set a limit on number of partitions/height of the tree). The anomaly will be detected in smaller number of partitions than a normal point
So clearly the path length indicates whether a point is a normal or an anomalous point. (Path length - of a point x is measured by the number of edges x traverses an Isolation tree from the root node until the traversal is terminated at an external node)
Isolation forest is an ensemble method. So we create multiple Isolation trees(generally 100 trees will suffice) and we take the average of all the path lengths. This average path length will then decide whether a point is anomalous or not.
```
import pandas as pd
import requests
import datetime
import json
# Declare method for collecting data
def collect_data():
# this part will be in config or provided by broker in implementation
training_days = 365
repo_id = 21000
# Declare the endpoints we want to discover insights for
endpoints = [{'issues-new': 'issues'}, {'code-changes': 'commit_count'}, {'code-changes-lines': 'added'},
{'reviews': 'pull_requests'}, {'contributors-new': 'new_contributors'}]
base_url = 'http://localhost:5002/api/unstable/repo-groups/9999/repos/{}/'.format(repo_id)
# Dataframe to hold all endpoint results
# Subtract configurable amount of time
begin_date = datetime.datetime.now().replace(hour=0, minute=0, second=0, microsecond=0) - datetime.timedelta(days=training_days)
# begin_date = begin_date.strftime('%Y-%m-%d')
index = pd.date_range(begin_date, periods=training_days, freq='D')
df = pd.DataFrame(index=index)#, columns=[key for d in endpoints for key in d.keys()])
# Hit and discover insights for every endpoint we care about
for metric_meta in endpoints:
for endpoint, field in metric_meta.items():
# Hit endpoint
url = base_url + endpoint
print("Hitting endpoint: " + url + "\n")
# try:
data = requests.get(url=url).json()
# except:
# data = json.loads(json.dumps(requests.get(url=url).text))
if len(data) == 0:
print("Endpoint with url: {} returned an empty response. Moving on to next endpoint.\n".format(url))
continue
if 'date' not in data[0]:
print("Endpoint {} is not a timeseries, moving to next endpoint.".format(endpoint))
continue
metric_df = pd.DataFrame.from_records(data)
metric_df.index = pd.to_datetime(metric_df['date'], utc=True).dt.date
# print(metric_df)
# df.add(metric_df[field])
# df = pd.concat(df, metric_df[field])
df = df.join(metric_df[field]).fillna(0)
# print(df)
return df
df = collect_data()
df
```
Visualize all our metrics before doing anomaly detection
```
df.plot(y=['issues'], kind='line')
df.plot(y=['commit_count'], kind='line')
df.plot(y=['pull_requests'], kind='line')
df.plot(y=['added'], kind='line')
df.plot(y=['new_contributors'], kind='line')
```
Isolation forest tries to separate each point in the data. In case of 2D data it randomly creates a line and tries to single out a point. Here an anomalous point could be separated in a few steps while normal points which are closer could take significantly more steps to be segregated.
I am using sklearn’s Isolation Forest here as it is a small dataset with 1 year of data, worth noting h2o’s isolation forest is a potentially more scalable possibility on high volume datasets, if the amount of data starts to impact sklearn's performace
Contamination is an important parameter here. It is the assumption of the percentage of outlier points in the data. Through testing I found 0.041 to be a fitting value for our use case (which will give us around 15 anomalies / year per metric).
THIS CAN AND SHOULD BE TWEAKED THOUGH:
15-30 anomalies a year makes sense for rails/rails but not so much for smaller repos where there is only 15 commits / year, we don't want EVERY action to be considered anomalous. A table of updated contamination values could help control something like this, provide flexibility for different repos, and allow a possibility to make this algorithm's implementation semi-supervised (maybe with questions in the slack bot: Would you consider this anomaly that you were just notified about a TRUE anomaly? [yes / no]) if yes -> do nothing, if no -> lower the contamination value for this repo slightly. Or another question: did any events occur in your repository that YOU would consider anomalous that our slack bot did not notify you about? if yes -> increase contamination parameter
```
from sklearn.ensemble import IsolationForest
# specify the 4 metric names that we will model
to_model_columns = df.columns[0:5]
model = IsolationForest(n_estimators=100, max_samples='auto', contamination=float(.041), \
max_features=1.0, bootstrap=False, n_jobs=-1, random_state=32, verbose=0)
model.fit(df[to_model_columns])
import numpy as np
pred = model.predict(df[to_model_columns])
df['anomaly'] = pred
outliers = df.loc[df['anomaly'] == -1]
outlier_index = list(outliers.index)
# Find the number of anomalies and normal points here points classified -1 are anomalous
print(df['anomaly'].value_counts())
boolean_anomalies = df['anomaly'].where(df['anomaly'] == -1, False).where(df['anomaly'] != -1, True).values
boolean_anomalies = boolean_anomalies.astype(np.bool_)
```
Now we have 4 metrics on which we have classified anomalies based on isolation forest. We will try to visualize the results and check if the classification makes sense.
We will now normalize and fit the metrics to a PCA to reduce the number of dimensions and then plot them in 3D highlighting the anomalies.
This chart below is very conceptual as it is plotting anomalies for the repo as a whole, and our repo is then reduced to 3 defining fields (which do NOT map to any specific metric, but are rather entirely "new" parameters that define the whole repo) which are the parameters that make up the x, y, and x axes.
Anomalies are red X's and normal points are indicated with green points in the plot.
```
import matplotlib.pyplot as plt
from sklearn.decomposition import PCA
from sklearn.preprocessing import StandardScaler
from mpl_toolkits.mplot3d import Axes3D
pca = PCA(n_components=3) # Reduce to k=3 dimensions
scaler = StandardScaler()
# normalize the metrics
X = scaler.fit_transform(df[to_model_columns])
X_reduce = pca.fit_transform(X)
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
ax.set_zlabel("x_composite_3") # Plot the compressed data points
ax.scatter(X_reduce[:, 0], X_reduce[:, 1], zs=X_reduce[:, 2], s=4, lw=1, label="inliers",c="green")# Plot x's for the ground truth outliers
ax.scatter(X_reduce[boolean_anomalies, 0],X_reduce[boolean_anomalies, 1], X_reduce[boolean_anomalies, 2],
lw=2, s=60, marker="x", c="red", label="outliers")
ax.legend()
plt.show()
```
Now as 2D
Anomalies are highlighted as red edges and normal points are indicated with green points in the plot.
Here the contamination parameter plays a great factor.
Another conceptual chart that helps see anomalies across the whole repo as well, but also helps in the parameter-tuning process, specifically for the contamination parameter.
Our idea here is to capture all the anomalous point in the system. So its better to identify few points which might be normal as anomalous (false positives), but not to miss out catching an anomaly (true negative). (So I have specified 4.1% as contamination which varies based on use case and repo)
```
pca = PCA(n_components=2)
scaler = StandardScaler()
# normalize the metrics
X = scaler.fit_transform(df[to_model_columns])
X_reduce = pca.fit_transform(X)
# pca.fit(df[to_model_columns])
# res = pd.DataFrame(pca.transform(df[to_model_columns]))
# Z = np.array(res)
plt.title("IsolationForest")
# plt.contourf(Z, cmap=plt.cm.Blues_r)
# b1 = plt.scatter(res[0], res[1], c='green', s=20, label="normal points")
# b1 = plt.scatter(res.iloc[boolean_anomalies,0], res.iloc[boolean_anomalies,1], c='green', s=20, edgecolor="red", label="predicted outliers")
b1 = plt.scatter(X_reduce[:, 0], X_reduce[:, 1], c='green', s=20, label="normal points")
b1 = plt.scatter(X_reduce[boolean_anomalies, 0], X_reduce[boolean_anomalies, 1], c='green', s=20, edgecolor="red", label="predicted outliers")
plt.legend(loc="upper right")
plt.show()
```
Now we have figured the anomalous behavior at a use case level, but to be actionable on the anomaly its important to identify and provide information on which specific metrics are anomalous individually.
The anomalies identified by the algorithm should make sense when viewed visually (sudden dip/peaks) by the business user to act upon it. So creating a good visualization is equally important in this process.
The following function creates actuals plot on a time series with anomaly points highlighted on it. Also a table which provides actual data, the change and conditional formatting based on anomalies.
```
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot
import chart_studio.plotly as py
from matplotlib import pyplot
import plotly.graph_objs as go
init_notebook_mode(connected=True)
def plot_anomaly(df,field_name):
df.date = pd.to_datetime(df['date_col'].astype(str))#, format="%Y%m%d")
dates = df.date
# Identify the anomaly points and create a array of its values for plot
bool_array = (abs(df['anomaly']) > 0)
metric_value = df[field_name][-len(bool_array):]
anomaly_points = bool_array * metric_value
anomaly_points[anomaly_points == 0] = np.nan
med_bool_array = (abs(test_df['anomaly_class']) == 1)
med_metric_value = df[field_name][-len(med_bool_array):]
med_anomaly_points = med_bool_array * med_metric_value
med_anomaly_points[med_anomaly_points == 0] = np.nan
# A dictionary for conditional format table based on anomaly
color_map = {0: "rgba(228, 222, 249, 0.65)", 1: "yellow", 2: "red"} #rgba(228, 222, 249, 0.65)
# Table which includes Date, Actuals, Change occured from previous point
table = go.Table(
domain = dict(x = [0, 1], y = [0, 0.3]),
columnwidth = [1, 2],
# columnorder=[0, 1, 2,],
header = dict(
height=20,
values=[['<b>Date</b>'], ['<b>Metric Value </b>'], ['<b>% Change </b>'],],
font=dict(color=['rgb(45, 45, 45)'] * 5, size=14),
fill=dict(color='#d562be')
),
cells = dict(
values = [df.round(3)[k].tolist() for k in ['date_col', field_name, 'percentage_change']],
line = dict(color='#506784'),
align = ['center'] * 5,
font = dict(color=['rgb(40, 40, 40)'] * 5, size=12),
# format = [None] + [",.4f"] + [',.4f'],
# suffix = [None] * 4,
suffix = [None] + [''] + [''] + ['%'] + [''],
height = 27,
fill = dict(color = [test_df['anomaly_class'].map(color_map)],)#map based on anomaly level from dictionary
)
)
#Plot the actuals points
metric = go.Scatter(
name = 'Metric Value',
x = dates,
y = df[field_name],
xaxis = 'x1',
yaxis = 'y1',
mode = 'lines',
marker = dict(
size = 12,
line = dict(width=1),
color = "blue")
) #Highlight the anomaly points
anomalies_map = go.Scatter(
name = "Anomaly",
showlegend = True,
x = dates,
y = anomaly_points,
mode = 'markers',
xaxis = 'x1',
yaxis = 'y1',
marker = dict(
color = "red",
size = 11,
line = dict(
color = "red",
width = 2)
)
)
med_anomalies_map = go.Scatter(
name = "Light Anomaly",
showlegend = True,
x = dates,
y = med_anomaly_points,
mode = 'markers',
xaxis = 'x1',
yaxis = 'y1',
marker = dict(
color = "yellow",
size = 11,
line = dict(
color = "yellow",
width = 2)
)
)
axis = dict(
showline = True,
zeroline = False,
showgrid = True,
mirror = True,
ticklen = 4,
gridcolor = '#ffffff',
tickfont = dict(size=10)
)
layout = dict(
width = 1000,
height = 865,
autosize = False,
title = field_name,
margin = dict(t = 75),
showlegend = True,
xaxis1 = dict(axis, **dict(domain=[0, 1], anchor='y1', showticklabels=True)),
yaxis1 = dict(axis, **dict(domain=[2 * 0.21 + 0.20, 1], anchor='x1', hoverformat='.2f'))
)
fig = go.Figure(data=[table, anomalies_map, med_anomalies_map, metric], layout=layout)
iplot(fig)
pyplot.show()
```
A helper function to find percentage change,classify anomaly based on severity.
The predict function classifies the data as anomalies based on the results from decision function on crossing a threshold. Say if the business needs to find the next level of anomalies which might have an impact, this could be used to identify those points.
The top 12 quantiles are identified anomalies(high severity), based on decision function here we identify the 12–24 quantile points and classify them as low severity anomalies.
```
def classify_anomalies(df,field_name):
# df[field_name] = metric_name
df = df.sort_values(by='date_col', ascending=False)
# Shift metric values by one date to find the percentage chage between current and previous data point
df['shift'] = df[field_name].shift(-1)
df['percentage_change'] = ((df[field_name] - df['shift']) / df[field_name]) * 100
# Categorise anomalies as 0 - no anomaly, 1 - low anomaly , 2 - high anomaly
df['anomaly'].loc[df['anomaly'] == 1] = 0
df['anomaly'].loc[df['anomaly'] == -1] = 2
df['anomaly_class'] = df['anomaly']
max_anomaly_score = df['score'].loc[df['anomaly_class'] == 2].max()
medium_percentile = df['score'].quantile(0.24)
df['anomaly_class'].loc[(df['score'] > max_anomaly_score) & (df['score'] <= medium_percentile)] = 1
return df
```
Identify anomalies for individual metrics and plot the results.
X axis — date Y axis — Metric values and anomaly points.
The plots allow us to see the sudden spikes and dips in the metric's value and project the anomalous points onto the plot.
The tables below each plot allow us to examine the raw data and see potential holes or broken pipelines in the data that would need to be fixed in our preprocessing steps. They also let us see which anomalies are low-level (yellow) or high level (red)
```
import warnings
warnings.filterwarnings('ignore')
for i, field_name in enumerate(to_model_columns):
model.fit(df.iloc[:,i:i+1])
pred = model.predict(df.iloc[:,i:i+1])
test_df = pd.DataFrame()
test_df['date_col'] = df.index#df['date']
test_df.index = df.index
# Find decision function to find the score and classify anomalies
test_df['score'] = model.decision_function(df.iloc[:,i:i+1])
test_df[field_name] = df.iloc[:,i:i+1]
test_df['anomaly'] = pred
# Get the indexes of outliers in order to compare the metrics with use case anomalies if required
outliers = test_df.loc[test_df['anomaly'] == -1]
outlier_index = list(outliers.index)
test_df = classify_anomalies(test_df,field_name)
plot_anomaly(test_df,field_name)
```
Things to do next:
- map PCA results over time (ie show WHEN the repo AS A WHOLE is in an anomalous state)
- appeal to a "detection period" (ie still train over whole year but only show anomalies in past x days)
- other ideas I can't remember right now
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.