text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
<a href="https://colab.research.google.com/github/satyajitghana/TSAI-DeepNLP-END2.0/blob/main/08_TorchText/pytorch-seq2seq-modern/4_Packed_Padded_Sequences%2C_Masking%2C_Inference_and_BLEU.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# 4 - Packed Padded Sequences, Masking, Inference and BLEU
## Introduction
In this notebook we will be adding a few improvements - packed padded sequences and masking - to the model from the previous notebook. Packed padded sequences are used to tell our RNN to skip over padding tokens in our encoder. Masking explicitly forces the model to ignore certain values, such as attention over padded elements. Both of these techniques are commonly used in NLP.
We will also look at how to use our model for inference, by giving it a sentence, seeing what it translates it as and seeing where exactly it pays attention to when translating each word.
Finally, we'll use the BLEU metric to measure the quality of our translations.
## Preparing Data
First, we'll import all the modules as before, with the addition of the `matplotlib` modules used for viewing the attention.
```
! pip install spacy==3.0.6 --quiet
```
You might need to restart the Runtime after installing the spacy models
```
! python -m spacy download en_core_web_sm --quiet
! python -m spacy download de_core_news_sm --quiet
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
# from torchtext.legacy.datasets import Multi30k
# from torchtext.legacy.data import Field, BucketIterator
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
import spacy
import numpy as np
import random
import math
import time
from typing import *
```
Next, we'll set the random seed for reproducability.
```
SEED = 1234
random.seed(SEED)
np.random.seed(SEED)
torch.manual_seed(SEED)
torch.cuda.manual_seed(SEED)
torch.backends.cudnn.deterministic = True
```
As before, we'll import spaCy and define the German and English tokenizers.
```
from torchtext.data.utils import get_tokenizer
from torchtext.vocab import build_vocab_from_iterator
from torchtext.datasets import Multi30k
SRC_LANGUAGE = 'de'
TGT_LANGUAGE = 'en'
# Place-holders
token_transform = {}
vocab_transform = {}
# Create source and target language tokenizer. Make sure to install the dependencies.
# the 'language' should be a full qualified name, since shortcuts like `de` and `en` are deprecated in spaCy 3.0+
token_transform[SRC_LANGUAGE] = get_tokenizer('spacy', language='de_core_news_sm')
token_transform[TGT_LANGUAGE] = get_tokenizer('spacy', language='en_core_web_sm')
# Training, Validation and Test data Iterator
train_iter, val_iter, test_iter = Multi30k(split=('train', 'valid', 'test'), language_pair=(SRC_LANGUAGE, TGT_LANGUAGE))
train_list, val_list, test_list = list(train_iter), list(val_iter), list(test_iter)
train_list[0]
```
Build the vocabulary.
```
# helper function to yield list of tokens
def yield_tokens(data_iter: Iterable, language: str) -> List[str]:
language_index = {SRC_LANGUAGE: 0, TGT_LANGUAGE: 1}
for data_sample in data_iter:
yield token_transform[language](data_sample[language_index[language]])
# Define special symbols and indices
UNK_IDX, PAD_IDX, BOS_IDX, EOS_IDX = 0, 1, 2, 3
# Make sure the tokens are in order of their indices to properly insert them in vocab
special_symbols = ['<unk>', '<pad>', '<bos>', '<eos>']
for ln in [SRC_LANGUAGE, TGT_LANGUAGE]:
# Create torchtext's Vocab object
vocab_transform[ln] = build_vocab_from_iterator(yield_tokens(train_list, ln),
min_freq=1,
specials=special_symbols,
special_first=True)
# Set UNK_IDX as the default index. This index is returned when the token is not found.
# If not set, it throws RuntimeError when the queried token is not found in the Vocabulary.
for ln in [SRC_LANGUAGE, TGT_LANGUAGE]:
vocab_transform[ln].set_default_index(UNK_IDX)
```
Define the device.
```
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
```
Create the iterators.
```
from torch.nn.utils.rnn import pad_sequence
# helper function to club together sequential operations
def sequential_transforms(*transforms):
def func(txt_input):
for transform in transforms:
txt_input = transform(txt_input)
return txt_input
return func
# function to add BOS/EOS and create tensor for input sequence indices
def tensor_transform(token_ids: List[int]):
return torch.cat((torch.tensor([BOS_IDX]),
torch.tensor(token_ids),
torch.tensor([EOS_IDX])))
# src and tgt language text transforms to convert raw strings into tensors indices
text_transform = {}
for ln in [SRC_LANGUAGE, TGT_LANGUAGE]:
text_transform[ln] = sequential_transforms(token_transform[ln], #Tokenization
vocab_transform[ln], #Numericalization
tensor_transform) # Add BOS/EOS and create tensor
# function to collate data samples into batch tesors
def collate_fn(batch):
src_batch, src_len, tgt_batch = [], [], []
for src_sample, tgt_sample in batch:
src_batch.append(text_transform[SRC_LANGUAGE](src_sample.rstrip("\n")))
src_len.append(len(src_batch[-1]))
tgt_batch.append(text_transform[TGT_LANGUAGE](tgt_sample.rstrip("\n")))
src_batch = pad_sequence(src_batch, padding_value=PAD_IDX)
tgt_batch = pad_sequence(tgt_batch, padding_value=PAD_IDX)
return src_batch, torch.LongTensor(src_len), tgt_batch
from torch.utils.data import DataLoader
BATCH_SIZE = 128
train_dataloader = DataLoader(train_list, batch_size=BATCH_SIZE, collate_fn=collate_fn)
val_dataloader = DataLoader(val_list, batch_size=BATCH_SIZE, collate_fn=collate_fn)
test_dataloader = DataLoader(test_list, batch_size=BATCH_SIZE, collate_fn=collate_fn)
```
When using packed padded sequences, we need to tell PyTorch how long the actual (non-padded) sequences are. Luckily for us, TorchText's `Field` objects allow us to use the `include_lengths` argument, this will cause our `batch.src` to be a tuple. The first element of the tuple is the same as before, a batch of numericalized source sentence as a tensor, and the second element is the non-padded lengths of each source sentence within the batch.
We then load the data.
Next, we handle the iterators.
One quirk about packed padded sequences is that all elements in the batch need to be sorted by their non-padded lengths in descending order, i.e. the first sentence in the batch needs to be the longest. We use two arguments of the iterator to handle this, `sort_within_batch` which tells the iterator that the contents of the batch need to be sorted, and `sort_key` a function which tells the iterator how to sort the elements in the batch. Here, we sort by the length of the `src` sentence.
## Building the Model
### Encoder
Next up, we define the encoder.
The changes here all within the `forward` method. It now accepts the lengths of the source sentences as well as the sentences themselves.
After the source sentence (padded automatically within the iterator) has been embedded, we can then use `pack_padded_sequence` on it with the lengths of the sentences. Note that the tensor containing the lengths of the sequences must be a CPU tensor as of the latest version of PyTorch, which we explicitly do so with `to('cpu')`. `packed_embedded` will then be our packed padded sequence. This can be then fed to our RNN as normal which will return `packed_outputs`, a packed tensor containing all of the hidden states from the sequence, and `hidden` which is simply the final hidden state from our sequence. `hidden` is a standard tensor and not packed in any way, the only difference is that as the input was a packed sequence, this tensor is from the final **non-padded element** in the sequence.
We then unpack our `packed_outputs` using `pad_packed_sequence` which returns the `outputs` and the lengths of each, which we don't need.
The first dimension of `outputs` is the padded sequence lengths however due to using a packed padded sequence the values of tensors when a padding token was the input will be all zeros.
```
class Encoder(nn.Module):
def __init__(self, input_dim, emb_dim, enc_hid_dim, dec_hid_dim, dropout):
super().__init__()
self.embedding = nn.Embedding(input_dim, emb_dim)
self.rnn = nn.GRU(emb_dim, enc_hid_dim, bidirectional = True)
self.fc = nn.Linear(enc_hid_dim * 2, dec_hid_dim)
self.dropout = nn.Dropout(dropout)
def forward(self, src, src_len):
#src = [src len, batch size]
#src_len = [batch size]
embedded = self.dropout(self.embedding(src))
#embedded = [src len, batch size, emb dim]
#need to explicitly put lengths on cpu!
packed_embedded = nn.utils.rnn.pack_padded_sequence(embedded, src_len.to('cpu'), enforce_sorted=False)
packed_outputs, hidden = self.rnn(packed_embedded)
#packed_outputs is a packed sequence containing all hidden states
#hidden is now from the final non-padded element in the batch
outputs, _ = nn.utils.rnn.pad_packed_sequence(packed_outputs)
#outputs is now a non-packed sequence, all hidden states obtained
# when the input is a pad token are all zeros
#outputs = [src len, batch size, hid dim * num directions]
#hidden = [n layers * num directions, batch size, hid dim]
#hidden is stacked [forward_1, backward_1, forward_2, backward_2, ...]
#outputs are always from the last layer
#hidden [-2, :, : ] is the last of the forwards RNN
#hidden [-1, :, : ] is the last of the backwards RNN
#initial decoder hidden is final hidden state of the forwards and backwards
# encoder RNNs fed through a linear layer
hidden = torch.tanh(self.fc(torch.cat((hidden[-2,:,:], hidden[-1,:,:]), dim = 1)))
#outputs = [src len, batch size, enc hid dim * 2]
#hidden = [batch size, dec hid dim]
return outputs, hidden
```
### Attention
The attention module is where we calculate the attention values over the source sentence.
Previously, we allowed this module to "pay attention" to padding tokens within the source sentence. However, using *masking*, we can force the attention to only be over non-padding elements.
The `forward` method now takes a `mask` input. This is a **[batch size, source sentence length]** tensor that is 1 when the source sentence token is not a padding token, and 0 when it is a padding token. For example, if the source sentence is: ["hello", "how", "are", "you", "?", `<pad>`, `<pad>`], then the mask would be [1, 1, 1, 1, 1, 0, 0].
We apply the mask after the attention has been calculated, but before it has been normalized by the `softmax` function. It is applied using `masked_fill`. This fills the tensor at each element where the first argument (`mask == 0`) is true, with the value given by the second argument (`-1e10`). In other words, it will take the un-normalized attention values, and change the attention values over padded elements to be `-1e10`. As these numbers will be miniscule compared to the other values they will become zero when passed through the `softmax` layer, ensuring no attention is payed to padding tokens in the source sentence.
```
class Attention(nn.Module):
def __init__(self, enc_hid_dim, dec_hid_dim):
super().__init__()
self.attn = nn.Linear((enc_hid_dim * 2) + dec_hid_dim, dec_hid_dim)
self.v = nn.Linear(dec_hid_dim, 1, bias = False)
def forward(self, hidden, encoder_outputs, mask):
#hidden = [batch size, dec hid dim]
#encoder_outputs = [src len, batch size, enc hid dim * 2]
batch_size = encoder_outputs.shape[1]
src_len = encoder_outputs.shape[0]
#repeat decoder hidden state src_len times
hidden = hidden.unsqueeze(1).repeat(1, src_len, 1)
encoder_outputs = encoder_outputs.permute(1, 0, 2)
#hidden = [batch size, src len, dec hid dim]
#encoder_outputs = [batch size, src len, enc hid dim * 2]
energy = torch.tanh(self.attn(torch.cat((hidden, encoder_outputs), dim = 2)))
#energy = [batch size, src len, dec hid dim]
attention = self.v(energy).squeeze(2)
#attention = [batch size, src len]
attention = attention.masked_fill(mask == 0, -1e10)
return F.softmax(attention, dim = 1)
```
### Decoder
The decoder only needs a few small changes. It needs to accept a mask over the source sentence and pass this to the attention module. As we want to view the values of attention during inference, we also return the attention tensor.
```
class Decoder(nn.Module):
def __init__(self, output_dim, emb_dim, enc_hid_dim, dec_hid_dim, dropout, attention):
super().__init__()
self.output_dim = output_dim
self.attention = attention
self.embedding = nn.Embedding(output_dim, emb_dim)
self.rnn = nn.GRU((enc_hid_dim * 2) + emb_dim, dec_hid_dim)
self.fc_out = nn.Linear((enc_hid_dim * 2) + dec_hid_dim + emb_dim, output_dim)
self.dropout = nn.Dropout(dropout)
def forward(self, input, hidden, encoder_outputs, mask):
#input = [batch size]
#hidden = [batch size, dec hid dim]
#encoder_outputs = [src len, batch size, enc hid dim * 2]
#mask = [batch size, src len]
input = input.unsqueeze(0)
#input = [1, batch size]
embedded = self.dropout(self.embedding(input))
#embedded = [1, batch size, emb dim]
a = self.attention(hidden, encoder_outputs, mask)
#a = [batch size, src len]
a = a.unsqueeze(1)
#a = [batch size, 1, src len]
encoder_outputs = encoder_outputs.permute(1, 0, 2)
#encoder_outputs = [batch size, src len, enc hid dim * 2]
weighted = torch.bmm(a, encoder_outputs)
#weighted = [batch size, 1, enc hid dim * 2]
weighted = weighted.permute(1, 0, 2)
#weighted = [1, batch size, enc hid dim * 2]
rnn_input = torch.cat((embedded, weighted), dim = 2)
#rnn_input = [1, batch size, (enc hid dim * 2) + emb dim]
output, hidden = self.rnn(rnn_input, hidden.unsqueeze(0))
#output = [seq len, batch size, dec hid dim * n directions]
#hidden = [n layers * n directions, batch size, dec hid dim]
#seq len, n layers and n directions will always be 1 in this decoder, therefore:
#output = [1, batch size, dec hid dim]
#hidden = [1, batch size, dec hid dim]
#this also means that output == hidden
assert (output == hidden).all()
embedded = embedded.squeeze(0)
output = output.squeeze(0)
weighted = weighted.squeeze(0)
prediction = self.fc_out(torch.cat((output, weighted, embedded), dim = 1))
#prediction = [batch size, output dim]
return prediction, hidden.squeeze(0), a.squeeze(1)
```
### Seq2Seq
The overarching seq2seq model also needs a few changes for packed padded sequences, masking and inference.
We need to tell it what the indexes are for the pad token and also pass the source sentence lengths as input to the `forward` method.
We use the pad token index to create the masks, by creating a mask tensor that is 1 wherever the source sentence is not equal to the pad token. This is all done within the `create_mask` function.
The sequence lengths as needed to pass to the encoder to use packed padded sequences.
The attention at each time-step is stored in the `attentions`
```
class Seq2Seq(nn.Module):
def __init__(self, encoder, decoder, src_pad_idx, device):
super().__init__()
self.encoder = encoder
self.decoder = decoder
self.src_pad_idx = src_pad_idx
self.device = device
def create_mask(self, src):
mask = (src != self.src_pad_idx).permute(1, 0)
return mask
def forward(self, src, src_len, trg, teacher_forcing_ratio = 0.5):
#src = [src len, batch size]
#src_len = [batch size]
#trg = [trg len, batch size]
#teacher_forcing_ratio is probability to use teacher forcing
#e.g. if teacher_forcing_ratio is 0.75 we use teacher forcing 75% of the time
batch_size = src.shape[1]
trg_len = trg.shape[0]
trg_vocab_size = self.decoder.output_dim
#tensor to store decoder outputs
outputs = torch.zeros(trg_len, batch_size, trg_vocab_size).to(self.device)
#encoder_outputs is all hidden states of the input sequence, back and forwards
#hidden is the final forward and backward hidden states, passed through a linear layer
encoder_outputs, hidden = self.encoder(src, src_len)
#first input to the decoder is the <sos> tokens
input = trg[0,:]
mask = self.create_mask(src)
#mask = [batch size, src len]
for t in range(1, trg_len):
#insert input token embedding, previous hidden state, all encoder hidden states
# and mask
#receive output tensor (predictions) and new hidden state
output, hidden, _ = self.decoder(input, hidden, encoder_outputs, mask)
#place predictions in a tensor holding predictions for each token
outputs[t] = output
#decide if we are going to use teacher forcing or not
teacher_force = random.random() < teacher_forcing_ratio
#get the highest predicted token from our predictions
top1 = output.argmax(1)
#if teacher forcing, use actual next token as next input
#if not, use predicted token
input = trg[t] if teacher_force else top1
return outputs
```
## Training the Seq2Seq Model
Next up, initializing the model and placing it on the GPU.
```
INPUT_DIM = len(vocab_transform[SRC_LANGUAGE])
OUTPUT_DIM = len(vocab_transform[TGT_LANGUAGE])
ENC_EMB_DIM = 256
DEC_EMB_DIM = 256
ENC_HID_DIM = 512
DEC_HID_DIM = 512
ENC_DROPOUT = 0.5
DEC_DROPOUT = 0.5
SRC_PAD_IDX = PAD_IDX
attn = Attention(ENC_HID_DIM, DEC_HID_DIM)
enc = Encoder(INPUT_DIM, ENC_EMB_DIM, ENC_HID_DIM, DEC_HID_DIM, ENC_DROPOUT)
dec = Decoder(OUTPUT_DIM, DEC_EMB_DIM, ENC_HID_DIM, DEC_HID_DIM, DEC_DROPOUT, attn)
model = Seq2Seq(enc, dec, SRC_PAD_IDX, device).to(device)
```
Then, we initialize the model parameters.
```
def init_weights(m):
for name, param in m.named_parameters():
if 'weight' in name:
nn.init.normal_(param.data, mean=0, std=0.01)
else:
nn.init.constant_(param.data, 0)
model.apply(init_weights)
```
We'll print out the number of trainable parameters in the model, noticing that it has the exact same amount of parameters as the model without these improvements.
```
def count_parameters(model):
return sum(p.numel() for p in model.parameters() if p.requires_grad)
print(f'The model has {count_parameters(model):,} trainable parameters')
```
Then we define our optimizer and criterion.
The `ignore_index` for the criterion needs to be the index of the pad token for the target language, not the source language.
```
optimizer = optim.Adam(model.parameters())
TRG_PAD_IDX = PAD_IDX
criterion = nn.CrossEntropyLoss(ignore_index = TRG_PAD_IDX)
```
Next, we'll define our training and evaluation loops.
As we are using `include_lengths = True` for our source field, `batch.src` is now a tuple with the first element being the numericalized tensor representing the sentence and the second element being the lengths of each sentence within the batch.
Our model also returns the attention vectors over the batch of source source sentences for each decoding time-step. We won't use these during the training/evaluation, but we will later for inference.
```
def train(model, iterator, optimizer, criterion, clip):
model.train()
epoch_loss = 0
for i, batch in enumerate(iterator):
src, src_len, trg = batch
src, src_len, trg = src.to(device), src_len.to(device), trg.to(device)
optimizer.zero_grad()
output = model(src, src_len, trg)
#trg = [trg len, batch size]
#output = [trg len, batch size, output dim]
output_dim = output.shape[-1]
output = output[1:].view(-1, output_dim)
trg = trg[1:].view(-1)
#trg = [(trg len - 1) * batch size]
#output = [(trg len - 1) * batch size, output dim]
loss = criterion(output, trg)
loss.backward()
torch.nn.utils.clip_grad_norm_(model.parameters(), clip)
optimizer.step()
epoch_loss += loss.item()
return epoch_loss / len(iterator)
def evaluate(model, iterator, criterion):
model.eval()
epoch_loss = 0
with torch.no_grad():
for i, batch in enumerate(iterator):
src, src_len, trg = batch
src, src_len, trg = src.to(device), src_len.to(device), trg.to(device)
output = model(src, src_len, trg, 0) #turn off teacher forcing
#trg = [trg len, batch size]
#output = [trg len, batch size, output dim]
output_dim = output.shape[-1]
output = output[1:].view(-1, output_dim)
trg = trg[1:].view(-1)
#trg = [(trg len - 1) * batch size]
#output = [(trg len - 1) * batch size, output dim]
loss = criterion(output, trg)
epoch_loss += loss.item()
return epoch_loss / len(iterator)
```
Then, we'll define a useful function for timing how long epochs take.
```
def epoch_time(start_time, end_time):
elapsed_time = end_time - start_time
elapsed_mins = int(elapsed_time / 60)
elapsed_secs = int(elapsed_time - (elapsed_mins * 60))
return elapsed_mins, elapsed_secs
```
The penultimate step is to train our model. Notice how it takes almost half the time as our model without the improvements added in this notebook.
```
N_EPOCHS = 10
CLIP = 1
best_valid_loss = float('inf')
for epoch in range(N_EPOCHS):
start_time = time.time()
train_loss = train(model, train_dataloader, optimizer, criterion, CLIP)
valid_loss = evaluate(model, val_dataloader, criterion)
end_time = time.time()
epoch_mins, epoch_secs = epoch_time(start_time, end_time)
if valid_loss < best_valid_loss:
best_valid_loss = valid_loss
torch.save(model.state_dict(), 'tut4-model.pt')
print(f'Epoch: {epoch+1:02} | Time: {epoch_mins}m {epoch_secs}s')
print(f'\tTrain Loss: {train_loss:.3f} | Train PPL: {math.exp(train_loss):7.3f}')
print(f'\t Val. Loss: {valid_loss:.3f} | Val. PPL: {math.exp(valid_loss):7.3f}')
```
Finally, we load the parameters from our best validation loss and get our results on the test set.
We get the improved test perplexity whilst almost being twice as fast!
```
model.load_state_dict(torch.load('tut4-model.pt'))
test_loss = evaluate(model, test_dataloader, criterion)
print(f'| Test Loss: {test_loss:.3f} | Test PPL: {math.exp(test_loss):7.3f} |')
```
## Inference
Now we can use our trained model to generate translations.
**Note:** these translations will be poor compared to examples shown in paper as they use hidden dimension sizes of 1000 and train for 4 days! They have been cherry picked in order to show off what attention should look like on a sufficiently sized model.
Our `translate_sentence` will do the following:
- ensure our model is in evaluation mode, which it should always be for inference
- tokenize the source sentence if it has not been tokenized (is a string)
- numericalize the source sentence
- convert it to a tensor and add a batch dimension
- get the length of the source sentence and convert to a tensor
- feed the source sentence into the encoder
- create the mask for the source sentence
- create a list to hold the output sentence, initialized with an `<sos>` token
- create a tensor to hold the attention values
- while we have not hit a maximum length
- get the input tensor, which should be either `<sos>` or the last predicted token
- feed the input, all encoder outputs, hidden state and mask into the decoder
- store attention values
- get the predicted next token
- add prediction to current output sentence prediction
- break if the prediction was an `<eos>` token
- convert the output sentence from indexes to tokens
- return the output sentence (with the `<sos>` token removed) and the attention values over the sequence
```
def translate_sentence(sentence, vocabs, init_token, eos_token, model, device, max_len = 50):
model.eval()
if isinstance(sentence, str):
nlp = spacy.load('de_core_news_sm')
tokens = [token.text.lower() for token in nlp(sentence)]
else:
tokens = [token.lower() for token in sentence]
tokens = [init_token] + tokens + [eos_token]
src_indexes = [vocabs['de'][token] for token in tokens]
src_tensor = torch.LongTensor(src_indexes).unsqueeze(1).to(device)
src_len = torch.LongTensor([len(src_indexes)])
with torch.no_grad():
encoder_outputs, hidden = model.encoder(src_tensor, src_len)
mask = model.create_mask(src_tensor)
trg_indexes = [vocabs['en'][init_token]]
attentions = torch.zeros(max_len, 1, len(src_indexes)).to(device)
for i in range(max_len):
trg_tensor = torch.LongTensor([trg_indexes[-1]]).to(device)
with torch.no_grad():
output, hidden, attention = model.decoder(trg_tensor, hidden, encoder_outputs, mask)
attentions[i] = attention
pred_token = output.argmax(1).item()
trg_indexes.append(pred_token)
if pred_token == vocabs['en'][eos_token]:
break
trg_tokens = [vocabs['en'].vocab.get_itos()[i] for i in trg_indexes]
return trg_tokens[1:], attentions[:len(trg_tokens)-1]
```
Next, we'll make a function that displays the model's attention over the source sentence for each target token generated.
```
def display_attention(sentence, translation, attention):
fig = plt.figure(figsize=(10,10))
ax = fig.add_subplot(111)
attention = attention.squeeze(1).cpu().detach().numpy()
cax = ax.matshow(attention, cmap='bone')
ax.tick_params(labelsize=15)
if isinstance(sentence, str):
nlp = spacy.load('de_core_news_sm')
x_ticks = [''] + ['<sos>'] + [t.text.lower() for t in nlp(sentence)] + ['<eos>']
else:
x_ticks = [''] + ['<sos>'] + [t.lower() for t in sentence] + ['<eos>']
y_ticks = [''] + translation
ax.set_xticklabels(x_ticks, rotation=45)
ax.set_yticklabels(y_ticks)
ax.xaxis.set_major_locator(ticker.MultipleLocator(1))
ax.yaxis.set_major_locator(ticker.MultipleLocator(1))
plt.show()
plt.close()
```
Now, we'll grab some translations from our dataset and see how well our model did. Note, we're going to cherry pick examples here so it gives us something interesting to look at, but feel free to change the `example_idx` value to look at different examples.
First, we'll get a source and target from our dataset.
```
example_idx = 12
src, trg = train_list[example_idx]
print(f'src = {src}')
print(f'trg = {trg}')
```
Then we'll use our `translate_sentence` function to get our predicted translation and attention. We show this graphically by having the source sentence on the x-axis and the predicted translation on the y-axis. The lighter the square at the intersection between two words, the more attention the model gave to that source word when translating that target word.
Below is an example the model attempted to translate, it gets the translation correct except changes *are fighting* to just *fighting*.
```
translation, attention = translate_sentence(src, vocab_transform, '<bos>', '<eos>', model, device)
print(f'predicted trg = {translation}')
display_attention(src, translation, attention)
```
Translations from the training set could simply be memorized by the model. So it's only fair we look at translations from the validation and testing set too.
Starting with the validation set, let's get an example.
```
example_idx = 14
src, trg = train_list[example_idx]
print(f'src = {src}')
print(f'trg = {trg}')
```
Then let's generate our translation and view the attention.
Here, we can see the translation is the same except for swapping *female* with *woman*.
```
translation, attention = translate_sentence(src, vocab_transform, '<bos>', '<eos>', model, device)
print(f'predicted trg = {translation}')
display_attention(src, translation, attention)
```
Finally, let's get an example from the test set.
```
example_idx = 18
src, trg = train_list[example_idx]
print(f'src = {src}')
print(f'trg = {trg}')
```
Again, it produces a slightly different translation than target, a more literal version of the source sentence. It swaps *mountain climbing* for *climbing a mountain*.
```
translation, attention = translate_sentence(src, vocab_transform, '<bos>', '<eos>', model, device)
print(f'predicted trg = {translation}')
display_attention(src, translation, attention)
```
## BLEU
Previously we have only cared about the loss/perplexity of the model. However there metrics that are specifically designed for measuring the quality of a translation - the most popular is *BLEU*. Without going into too much detail, BLEU looks at the overlap in the predicted and actual target sequences in terms of their n-grams. It will give us a number between 0 and 1 for each sequence, where 1 means there is perfect overlap, i.e. a perfect translation, although is usually shown between 0 and 100. BLEU was designed for multiple candidate translations per source sequence, however in this dataset we only have one candidate per source.
We define a `calculate_bleu` function which calculates the BLEU score over a provided TorchText dataset. This function creates a corpus of the actual and predicted translation for each source sentence and then calculates the BLEU score.
```
from torchtext.data.metrics import bleu_score
from tqdm.auto import tqdm
def calculate_bleu(data, vocabs, init_token, eos_token, model, device, max_len = 50):
nlp = spacy.load('en_core_web_sm')
trgs = []
pred_trgs = []
for datum in tqdm(data):
src, trg = datum
if isinstance(trg, str):
trg = [t.text.lower() for t in nlp(trg)]
pred_trg, _ = translate_sentence(src, vocabs, init_token, eos_token, model, device, max_len)
#cut off <eos> token
pred_trg = pred_trg[:-1]
pred_trgs.append(pred_trg)
trgs.append([trg])
return bleu_score(pred_trgs, trgs)
```
We get a BLEU of around 28. If we compare it to the paper that the attention model is attempting to replicate, they achieve a BLEU score of 26.75. This is similar to our score, however they are using a completely different dataset and their model size is much larger - 1000 hidden dimensions which takes 4 days to train! - so we cannot really compare against that either.
This number isn't really interpretable, we can't really say much about it. The most useful part of a BLEU score is that it can be used to compare different models on the same dataset, where the one with the **higher** BLEU score is "better".
```
bleu_score_this = calculate_bleu(test_list, vocab_transform, '<bos>', '<eos>', model, device)
print(f'BLEU score = {bleu_score_this*100:.2f}')
```
In the next tutorials we will be moving away from using recurrent neural networks and start looking at other ways to construct sequence-to-sequence models. Specifically, in the next tutorial we will be using convolutional neural networks.
| github_jupyter |
<a href="https://colab.research.google.com/github/SummerLife/EmbeddedSystem/blob/master/MachineLearning/gist/visualization_of_the_filters_of_VGG16.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
"""
#Visualization of the filters of VGG16, via gradient ascent in input space.
This script can run on CPU in a few minutes.
Results example: 
"""
from __future__ import print_function
import time
import numpy as np
from PIL import Image as pil_image
from keras.preprocessing.image import save_img
from keras import layers
from keras.applications import vgg16
from keras import backend as K
def normalize(x):
"""utility function to normalize a tensor.
# Arguments
x: An input tensor.
# Returns
The normalized input tensor.
"""
return x / (K.sqrt(K.mean(K.square(x))) + K.epsilon())
def deprocess_image(x):
"""utility function to convert a float array into a valid uint8 image.
# Arguments
x: A numpy-array representing the generated image.
# Returns
A processed numpy-array, which could be used in e.g. imshow.
"""
# normalize tensor: center on 0., ensure std is 0.25
x -= x.mean()
x /= (x.std() + K.epsilon())
x *= 0.25
# clip to [0, 1]
x += 0.5
x = np.clip(x, 0, 1)
# convert to RGB array
x *= 255
if K.image_data_format() == 'channels_first':
x = x.transpose((1, 2, 0))
x = np.clip(x, 0, 255).astype('uint8')
return x
def process_image(x, former):
"""utility function to convert a valid uint8 image back into a float array.
Reverses `deprocess_image`.
# Arguments
x: A numpy-array, which could be used in e.g. imshow.
former: The former numpy-array.
Need to determine the former mean and variance.
# Returns
A processed numpy-array representing the generated image.
"""
if K.image_data_format() == 'channels_first':
x = x.transpose((2, 0, 1))
return (x / 255 - 0.5) * 4 * former.std() + former.mean()
def visualize_layer(model,
layer_name,
step=1.,
epochs=15,
upscaling_steps=9,
upscaling_factor=1.2,
output_dim=(412, 412),
filter_range=(0, None)):
"""Visualizes the most relevant filters of one conv-layer in a certain model.
# Arguments
model: The model containing layer_name.
layer_name: The name of the layer to be visualized.
Has to be a part of model.
step: step size for gradient ascent.
epochs: Number of iterations for gradient ascent.
upscaling_steps: Number of upscaling steps.
Starting image is in this case (80, 80).
upscaling_factor: Factor to which to slowly upgrade
the image towards output_dim.
output_dim: [img_width, img_height] The output image dimensions.
filter_range: Tupel[lower, upper]
Determines the to be computed filter numbers.
If the second value is `None`,
the last filter will be inferred as the upper boundary.
"""
def _generate_filter_image(input_img,
layer_output,
filter_index):
"""Generates image for one particular filter.
# Arguments
input_img: The input-image Tensor.
layer_output: The output-image Tensor.
filter_index: The to be processed filter number.
Assumed to be valid.
#Returns
Either None if no image could be generated.
or a tuple of the image (array) itself and the last loss.
"""
s_time = time.time()
# we build a loss function that maximizes the activation
# of the nth filter of the layer considered
if K.image_data_format() == 'channels_first':
loss = K.mean(layer_output[:, filter_index, :, :])
else:
loss = K.mean(layer_output[:, :, :, filter_index])
# we compute the gradient of the input picture wrt this loss
grads = K.gradients(loss, input_img)[0]
# normalization trick: we normalize the gradient
grads = normalize(grads)
# this function returns the loss and grads given the input picture
iterate = K.function([input_img], [loss, grads])
# we start from a gray image with some random noise
intermediate_dim = tuple(
int(x / (upscaling_factor ** upscaling_steps)) for x in output_dim)
if K.image_data_format() == 'channels_first':
input_img_data = np.random.random(
(1, 3, intermediate_dim[0], intermediate_dim[1]))
else:
input_img_data = np.random.random(
(1, intermediate_dim[0], intermediate_dim[1], 3))
input_img_data = (input_img_data - 0.5) * 20 + 128
# Slowly upscaling towards the original size prevents
# a dominating high-frequency of the to visualized structure
# as it would occur if we directly compute the 412d-image.
# Behaves as a better starting point for each following dimension
# and therefore avoids poor local minima
for up in reversed(range(upscaling_steps)):
# we run gradient ascent for e.g. 20 steps
for _ in range(epochs):
loss_value, grads_value = iterate([input_img_data])
input_img_data += grads_value * step
# some filters get stuck to 0, we can skip them
if loss_value <= K.epsilon():
return None
# Calculate upscaled dimension
intermediate_dim = tuple(
int(x / (upscaling_factor ** up)) for x in output_dim)
# Upscale
img = deprocess_image(input_img_data[0])
img = np.array(pil_image.fromarray(img).resize(intermediate_dim,
pil_image.BICUBIC))
input_img_data = np.expand_dims(
process_image(img, input_img_data[0]), 0)
# decode the resulting input image
img = deprocess_image(input_img_data[0])
e_time = time.time()
print('Costs of filter {:3}: {:5.0f} ( {:4.2f}s )'.format(filter_index,
loss_value,
e_time - s_time))
return img, loss_value
def _draw_filters(filters, n=None):
"""Draw the best filters in a nxn grid.
# Arguments
filters: A List of generated images and their corresponding losses
for each processed filter.
n: dimension of the grid.
If none, the largest possible square will be used
"""
if n is None:
n = int(np.floor(np.sqrt(len(filters))))
# the filters that have the highest loss are assumed to be better-looking.
# we will only keep the top n*n filters.
filters.sort(key=lambda x: x[1], reverse=True)
filters = filters[:n * n]
# build a black picture with enough space for
# e.g. our 8 x 8 filters of size 412 x 412, with a 5px margin in between
MARGIN = 5
width = n * output_dim[0] + (n - 1) * MARGIN
height = n * output_dim[1] + (n - 1) * MARGIN
stitched_filters = np.zeros((width, height, 3), dtype='uint8')
# fill the picture with our saved filters
for i in range(n):
for j in range(n):
img, _ = filters[i * n + j]
width_margin = (output_dim[0] + MARGIN) * i
height_margin = (output_dim[1] + MARGIN) * j
stitched_filters[
width_margin: width_margin + output_dim[0],
height_margin: height_margin + output_dim[1], :] = img
# save the result to disk
save_img('vgg_{0:}_{1:}x{1:}.png'.format(layer_name, n), stitched_filters)
# this is the placeholder for the input images
assert len(model.inputs) == 1
input_img = model.inputs[0]
# get the symbolic outputs of each "key" layer (we gave them unique names).
layer_dict = dict([(layer.name, layer) for layer in model.layers[1:]])
output_layer = layer_dict[layer_name]
assert isinstance(output_layer, layers.Conv2D)
# Compute to be processed filter range
filter_lower = filter_range[0]
filter_upper = (filter_range[1]
if filter_range[1] is not None
else len(output_layer.get_weights()[1]))
assert(filter_lower >= 0
and filter_upper <= len(output_layer.get_weights()[1])
and filter_upper > filter_lower)
print('Compute filters {:} to {:}'.format(filter_lower, filter_upper))
# iterate through each filter and generate its corresponding image
processed_filters = []
for f in range(filter_lower, filter_upper):
img_loss = _generate_filter_image(input_img, output_layer.output, f)
if img_loss is not None:
processed_filters.append(img_loss)
print('{} filter processed.'.format(len(processed_filters)))
# Finally draw and store the best filters to disk
_draw_filters(processed_filters)
if __name__ == '__main__':
# the name of the layer we want to visualize
# (see model definition at keras/applications/vgg16.py)
LAYER_NAME = 'block5_conv1'
# build the VGG16 network with ImageNet weights
vgg = vgg16.VGG16(weights='imagenet', include_top=False)
print('Model loaded.')
vgg.summary()
# example function call
# visualize_layer(vgg, LAYER_NAME)
visualize_layer(vgg, "block1_conv1")
visualize_layer(vgg, "block2_conv1")
```
| github_jupyter |
# Neural Machine Translation
Welcome to your first programming assignment for this week!
* You will build a Neural Machine Translation (NMT) model to translate human-readable dates ("25th of June, 2009") into machine-readable dates ("2009-06-25").
* You will do this using an attention model, one of the most sophisticated sequence-to-sequence models.
This notebook was produced together with NVIDIA's Deep Learning Institute.
## <font color='darkblue'>Updates</font>
#### If you were working on the notebook before this update...
* The current notebook is version "4a".
* You can find your original work saved in the notebook with the previous version name ("v4")
* To view the file directory, go to the menu "File->Open", and this will open a new tab that shows the file directory.
#### List of updates
* Clarified names of variables to be consistent with the lectures and consistent within the assignment
- pre-attention bi-directional LSTM: the first LSTM that processes the input data.
- 'a': the hidden state of the pre-attention LSTM.
- post-attention LSTM: the LSTM that outputs the translation.
- 's': the hidden state of the post-attention LSTM.
- energies "e". The output of the dense function that takes "a" and "s" as inputs.
- All references to "output activation" are updated to "hidden state".
- "post-activation" sequence model is updated to "post-attention sequence model".
- 3.1: "Getting the activations from the Network" renamed to "Getting the attention weights from the network."
- Appropriate mentions of "activation" replaced "attention weights."
- Sequence of alphas corrected to be a sequence of "a" hidden states.
* one_step_attention:
- Provides sample code for each Keras layer, to show how to call the functions.
- Reminds students to provide the list of hidden states in a specific order, in order to pause the autograder.
* model
- Provides sample code for each Keras layer, to show how to call the functions.
- Added a troubleshooting note about handling errors.
- Fixed typo: outputs should be of length 10 and not 11.
* define optimizer and compile model
- Provides sample code for each Keras layer, to show how to call the functions.
* Spelling, grammar and wording corrections.
Let's load all the packages you will need for this assignment.
```
from keras.layers import Bidirectional, Concatenate, Permute, Dot, Input, LSTM, Multiply
from keras.layers import RepeatVector, Dense, Activation, Lambda
from keras.optimizers import Adam
from keras.utils import to_categorical
from keras.models import load_model, Model
import keras.backend as K
import numpy as np
from faker import Faker
import random
from tqdm import tqdm
from babel.dates import format_date
from nmt_utils import *
import matplotlib.pyplot as plt
%matplotlib inline
```
## 1 - Translating human readable dates into machine readable dates
* The model you will build here could be used to translate from one language to another, such as translating from English to Hindi.
* However, language translation requires massive datasets and usually takes days of training on GPUs.
* To give you a place to experiment with these models without using massive datasets, we will perform a simpler "date translation" task.
* The network will input a date written in a variety of possible formats (*e.g. "the 29th of August 1958", "03/30/1968", "24 JUNE 1987"*)
* The network will translate them into standardized, machine readable dates (*e.g. "1958-08-29", "1968-03-30", "1987-06-24"*).
* We will have the network learn to output dates in the common machine-readable format YYYY-MM-DD.
<!--
Take a look at [nmt_utils.py](./nmt_utils.py) to see all the formatting. Count and figure out how the formats work, you will need this knowledge later. !-->
### 1.1 - Dataset
We will train the model on a dataset of 10,000 human readable dates and their equivalent, standardized, machine readable dates. Let's run the following cells to load the dataset and print some examples.
```
m = 10000
dataset, human_vocab, machine_vocab, inv_machine_vocab = load_dataset(m)
dataset[:10]
```
You've loaded:
- `dataset`: a list of tuples of (human readable date, machine readable date).
- `human_vocab`: a python dictionary mapping all characters used in the human readable dates to an integer-valued index.
- `machine_vocab`: a python dictionary mapping all characters used in machine readable dates to an integer-valued index.
- **Note**: These indices are not necessarily consistent with `human_vocab`.
- `inv_machine_vocab`: the inverse dictionary of `machine_vocab`, mapping from indices back to characters.
Let's preprocess the data and map the raw text data into the index values.
- We will set Tx=30
- We assume Tx is the maximum length of the human readable date.
- If we get a longer input, we would have to truncate it.
- We will set Ty=10
- "YYYY-MM-DD" is 10 characters long.
```
Tx = 30
Ty = 10
X, Y, Xoh, Yoh = preprocess_data(dataset, human_vocab, machine_vocab, Tx, Ty)
print("X.shape:", X.shape)
print("Y.shape:", Y.shape)
print("Xoh.shape:", Xoh.shape)
print("Yoh.shape:", Yoh.shape)
```
You now have:
- `X`: a processed version of the human readable dates in the training set.
- Each character in X is replaced by an index (integer) mapped to the character using `human_vocab`.
- Each date is padded to ensure a length of $T_x$ using a special character (< pad >).
- `X.shape = (m, Tx)` where m is the number of training examples in a batch.
- `Y`: a processed version of the machine readable dates in the training set.
- Each character is replaced by the index (integer) it is mapped to in `machine_vocab`.
- `Y.shape = (m, Ty)`.
- `Xoh`: one-hot version of `X`
- Each index in `X` is converted to the one-hot representation (if the index is 2, the one-hot version has the index position 2 set to 1, and the remaining positions are 0.
- `Xoh.shape = (m, Tx, len(human_vocab))`
- `Yoh`: one-hot version of `Y`
- Each index in `Y` is converted to the one-hot representation.
- `Yoh.shape = (m, Tx, len(machine_vocab))`.
- `len(machine_vocab) = 11` since there are 10 numeric digits (0 to 9) and the `-` symbol.
* Let's also look at some examples of preprocessed training examples.
* Feel free to play with `index` in the cell below to navigate the dataset and see how source/target dates are preprocessed.
```
index = 0
print("Source date:", dataset[index][0])
print("Target date:", dataset[index][1])
print()
print("Source after preprocessing (indices):", X[index])
print("Target after preprocessing (indices):", Y[index])
print()
print("Source after preprocessing (one-hot):", Xoh[index])
print("Target after preprocessing (one-hot):", Yoh[index])
```
## 2 - Neural machine translation with attention
* If you had to translate a book's paragraph from French to English, you would not read the whole paragraph, then close the book and translate.
* Even during the translation process, you would read/re-read and focus on the parts of the French paragraph corresponding to the parts of the English you are writing down.
* The attention mechanism tells a Neural Machine Translation model where it should pay attention to at any step.
### 2.1 - Attention mechanism
In this part, you will implement the attention mechanism presented in the lecture videos.
* Here is a figure to remind you how the model works.
* The diagram on the left shows the attention model.
* The diagram on the right shows what one "attention" step does to calculate the attention variables $\alpha^{\langle t, t' \rangle}$.
* The attention variables $\alpha^{\langle t, t' \rangle}$ are used to compute the context variable $context^{\langle t \rangle}$ for each timestep in the output ($t=1, \ldots, T_y$).
<table>
<td>
<img src="images/attn_model.png" style="width:500;height:500px;"> <br>
</td>
<td>
<img src="images/attn_mechanism.png" style="width:500;height:500px;"> <br>
</td>
</table>
<caption><center> **Figure 1**: Neural machine translation with attention</center></caption>
Here are some properties of the model that you may notice:
#### Pre-attention and Post-attention LSTMs on both sides of the attention mechanism
- There are two separate LSTMs in this model (see diagram on the left): pre-attention and post-attention LSTMs.
- *Pre-attention* Bi-LSTM is the one at the bottom of the picture is a Bi-directional LSTM and comes *before* the attention mechanism.
- The attention mechanism is shown in the middle of the left-hand diagram.
- The pre-attention Bi-LSTM goes through $T_x$ time steps
- *Post-attention* LSTM: at the top of the diagram comes *after* the attention mechanism.
- The post-attention LSTM goes through $T_y$ time steps.
- The post-attention LSTM passes the hidden state $s^{\langle t \rangle}$ and cell state $c^{\langle t \rangle}$ from one time step to the next.
#### An LSTM has both a hidden state and cell state
* In the lecture videos, we were using only a basic RNN for the post-attention sequence model
* This means that the state captured by the RNN was outputting only the hidden state $s^{\langle t\rangle}$.
* In this assignment, we are using an LSTM instead of a basic RNN.
* So the LSTM has both the hidden state $s^{\langle t\rangle}$ and the cell state $c^{\langle t\rangle}$.
#### Each time step does not use predictions from the previous time step
* Unlike previous text generation examples earlier in the course, in this model, the post-attention LSTM at time $t$ does not take the previous time step's prediction $y^{\langle t-1 \rangle}$ as input.
* The post-attention LSTM at time 't' only takes the hidden state $s^{\langle t\rangle}$ and cell state $c^{\langle t\rangle}$ as input.
* We have designed the model this way because unlike language generation (where adjacent characters are highly correlated) there isn't as strong a dependency between the previous character and the next character in a YYYY-MM-DD date.
#### Concatenation of hidden states from the forward and backward pre-attention LSTMs
- $\overrightarrow{a}^{\langle t \rangle}$: hidden state of the forward-direction, pre-attention LSTM.
- $\overleftarrow{a}^{\langle t \rangle}$: hidden state of the backward-direction, pre-attention LSTM.
- $a^{\langle t \rangle} = [\overrightarrow{a}^{\langle t \rangle}, \overleftarrow{a}^{\langle t \rangle}]$: the concatenation of the activations of both the forward-direction $\overrightarrow{a}^{\langle t \rangle}$ and backward-directions $\overleftarrow{a}^{\langle t \rangle}$ of the pre-attention Bi-LSTM.
#### Computing "energies" $e^{\langle t, t' \rangle}$ as a function of $s^{\langle t-1 \rangle}$ and $a^{\langle t' \rangle}$
- Recall in the lesson videos "Attention Model", at time 6:45 to 8:16, the definition of "e" as a function of $s^{\langle t-1 \rangle}$ and $a^{\langle t \rangle}$.
- "e" is called the "energies" variable.
- $s^{\langle t-1 \rangle}$ is the hidden state of the post-attention LSTM
- $a^{\langle t' \rangle}$ is the hidden state of the pre-attention LSTM.
- $s^{\langle t-1 \rangle}$ and $a^{\langle t \rangle}$ are fed into a simple neural network, which learns the function to output $e^{\langle t, t' \rangle}$.
- $e^{\langle t, t' \rangle}$ is then used when computing the attention $a^{\langle t, t' \rangle}$ that $y^{\langle t \rangle}$ should pay to $a^{\langle t' \rangle}$.
- The diagram on the right of figure 1 uses a `RepeatVector` node to copy $s^{\langle t-1 \rangle}$'s value $T_x$ times.
- Then it uses `Concatenation` to concatenate $s^{\langle t-1 \rangle}$ and $a^{\langle t \rangle}$.
- The concatenation of $s^{\langle t-1 \rangle}$ and $a^{\langle t \rangle}$ is fed into a "Dense" layer, which computes $e^{\langle t, t' \rangle}$.
- $e^{\langle t, t' \rangle}$ is then passed through a softmax to compute $\alpha^{\langle t, t' \rangle}$.
- Note that the diagram doesn't explicitly show variable $e^{\langle t, t' \rangle}$, but $e^{\langle t, t' \rangle}$ is above the Dense layer and below the Softmax layer in the diagram in the right half of figure 1.
- We'll explain how to use `RepeatVector` and `Concatenation` in Keras below.
### Implementation Details
Let's implement this neural translator. You will start by implementing two functions: `one_step_attention()` and `model()`.
#### one_step_attention
* The inputs to the one_step_attention at time step $t$ are:
- $[a^{<1>},a^{<2>}, ..., a^{<T_x>}]$: all hidden states of the pre-attention Bi-LSTM.
- $s^{<t-1>}$: the previous hidden state of the post-attention LSTM
* one_step_attention computes:
- $[\alpha^{<t,1>},\alpha^{<t,2>}, ..., \alpha^{<t,T_x>}]$: the attention weights
- $context^{ \langle t \rangle }$: the context vector:
$$context^{<t>} = \sum_{t' = 1}^{T_x} \alpha^{<t,t'>}a^{<t'>}\tag{1}$$
##### Clarifying 'context' and 'c'
- In the lecture videos, the context was denoted $c^{\langle t \rangle}$
- In the assignment, we are calling the context $context^{\langle t \rangle}$.
- This is to avoid confusion with the post-attention LSTM's internal memory cell variable, which is also denoted $c^{\langle t \rangle}$.
#### Implement `one_step_attention`
**Exercise**: Implement `one_step_attention()`.
* The function `model()` will call the layers in `one_step_attention()` $T_y$ using a for-loop.
* It is important that all $T_y$ copies have the same weights.
* It should not reinitialize the weights every time.
* In other words, all $T_y$ steps should have shared weights.
* Here's how you can implement layers with shareable weights in Keras:
1. Define the layer objects in a variable scope that is outside of the `one_step_attention` function. For example, defining the objects as global variables would work.
- Note that defining these variables inside the scope of the function `model` would technically work, since `model` will then call the `one_step_attention` function. For the purposes of making grading and troubleshooting easier, we are defining these as global variables. Note that the automatic grader will expect these to be global variables as well.
2. Call these objects when propagating the input.
* We have defined the layers you need as global variables.
* Please run the following cells to create them.
* Please note that the automatic grader expects these global variables with the given variable names. For grading purposes, please do not rename the global variables.
* Please check the Keras documentation to learn more about these layers. The layers are functions. Below are examples of how to call these functions.
* [RepeatVector()](https://keras.io/layers/core/#repeatvector)
```Python
var_repeated = repeat_layer(var1)
```
* [Concatenate()](https://keras.io/layers/merge/#concatenate)
```Python
concatenated_vars = concatenate_layer([var1,var2,var3])
```
* [Dense()](https://keras.io/layers/core/#dense)
```Python
var_out = dense_layer(var_in)
```
* [Activation()](https://keras.io/layers/core/#activation)
```Python
activation = activation_layer(var_in)
```
* [Dot()](https://keras.io/layers/merge/#dot)
```Python
dot_product = dot_layer([var1,var2])
```
```
# Defined shared layers as global variables
repeator = RepeatVector(Tx)
concatenator = Concatenate(axis=-1)
densor1 = Dense(10, activation = "tanh")
densor2 = Dense(1, activation = "relu")
activator = Activation(softmax, name='attention_weights') # We are using a custom softmax(axis = 1) loaded in this notebook
dotor = Dot(axes = 1)
# GRADED FUNCTION: one_step_attention
def one_step_attention(a, s_prev):
"""
Performs one step of attention: Outputs a context vector computed as a dot product of the attention weights
"alphas" and the hidden states "a" of the Bi-LSTM.
Arguments:
a -- hidden state output of the Bi-LSTM, numpy-array of shape (m, Tx, 2*n_a)
s_prev -- previous hidden state of the (post-attention) LSTM, numpy-array of shape (m, n_s)
Returns:
context -- context vector, input of the next (post-attention) LSTM cell
"""
### START CODE HERE ###
# Use repeator to repeat s_prev to be of shape (m, Tx, n_s) so that you can concatenate it with all hidden states "a" (≈ 1 line)
s_prev =repeator(s_prev)
# Use concatenator to concatenate a and s_prev on the last axis (≈ 1 line)
# For grading purposes, please list 'a' first and 's_prev' second, in this order.
concat = concatenator([a,s_prev])
# Use densor1 to propagate concat through a small fully-connected neural network to compute the "intermediate energies" variable e. (≈1 lines)
e = densor1(concat)
# Use densor2 to propagate e through a small fully-connected neural network to compute the "energies" variable energies. (≈1 lines)
energies = densor2(e)
# Use "activator" on "energies" to compute the attention weights "alphas" (≈ 1 line)
alphas = activator(energies)
# Use dotor together with "alphas" and "a" to compute the context vector to be given to the next (post-attention) LSTM-cell (≈ 1 line)
context = dotor([alphas,a])
### END CODE HERE ###
return context
```
You will be able to check the expected output of `one_step_attention()` after you've coded the `model()` function.
#### model
* `model` first runs the input through a Bi-LSTM to get $[a^{<1>},a^{<2>}, ..., a^{<T_x>}]$.
* Then, `model` calls `one_step_attention()` $T_y$ times using a `for` loop. At each iteration of this loop:
- It gives the computed context vector $context^{<t>}$ to the post-attention LSTM.
- It runs the output of the post-attention LSTM through a dense layer with softmax activation.
- The softmax generates a prediction $\hat{y}^{<t>}$.
**Exercise**: Implement `model()` as explained in figure 1 and the text above. Again, we have defined global layers that will share weights to be used in `model()`.
```
n_a = 32 # number of units for the pre-attention, bi-directional LSTM's hidden state 'a'
n_s = 64 # number of units for the post-attention LSTM's hidden state "s"
# Please note, this is the post attention LSTM cell.
# For the purposes of passing the automatic grader
# please do not modify this global variable. This will be corrected once the automatic grader is also updated.
post_activation_LSTM_cell = LSTM(n_s, return_state = True) # post-attention LSTM
output_layer = Dense(len(machine_vocab), activation=softmax)
```
Now you can use these layers $T_y$ times in a `for` loop to generate the outputs, and their parameters will not be reinitialized. You will have to carry out the following steps:
1. Propagate the input `X` into a bi-directional LSTM.
* [Bidirectional](https://keras.io/layers/wrappers/#bidirectional)
* [LSTM](https://keras.io/layers/recurrent/#lstm)
* Remember that we want the LSTM to return a full sequence instead of just the last hidden state.
Sample code:
```Python
sequence_of_hidden_states = Bidirectional(LSTM(units=..., return_sequences=...))(the_input_X)
```
2. Iterate for $t = 0, \cdots, T_y-1$:
1. Call `one_step_attention()`, passing in the sequence of hidden states $[a^{\langle 1 \rangle},a^{\langle 2 \rangle}, ..., a^{ \langle T_x \rangle}]$ from the pre-attention bi-directional LSTM, and the previous hidden state $s^{<t-1>}$ from the post-attention LSTM to calculate the context vector $context^{<t>}$.
2. Give $context^{<t>}$ to the post-attention LSTM cell.
- Remember to pass in the previous hidden-state $s^{\langle t-1\rangle}$ and cell-states $c^{\langle t-1\rangle}$ of this LSTM
* This outputs the new hidden state $s^{<t>}$ and the new cell state $c^{<t>}$.
Sample code:
```Python
next_hidden_state, _ , next_cell_state =
post_activation_LSTM_cell(inputs=..., initial_state=[prev_hidden_state, prev_cell_state])
```
Please note that the layer is actually the "post attention LSTM cell". For the purposes of passing the automatic grader, please do not modify the naming of this global variable. This will be fixed when we deploy updates to the automatic grader.
3. Apply a dense, softmax layer to $s^{<t>}$, get the output.
Sample code:
```Python
output = output_layer(inputs=...)
```
4. Save the output by adding it to the list of outputs.
3. Create your Keras model instance.
* It should have three inputs:
* `X`, the one-hot encoded inputs to the model, of shape ($T_{x}, humanVocabSize)$
* $s^{\langle 0 \rangle}$, the initial hidden state of the post-attention LSTM
* $c^{\langle 0 \rangle}$), the initial cell state of the post-attention LSTM
* The output is the list of outputs.
Sample code
```Python
model = Model(inputs=[...,...,...], outputs=...)
```
```
# GRADED FUNCTION: model
def model(Tx, Ty, n_a, n_s, human_vocab_size, machine_vocab_size):
"""
Arguments:
Tx -- length of the input sequence
Ty -- length of the output sequence
n_a -- hidden state size of the Bi-LSTM
n_s -- hidden state size of the post-attention LSTM
human_vocab_size -- size of the python dictionary "human_vocab"
machine_vocab_size -- size of the python dictionary "machine_vocab"
Returns:
model -- Keras model instance
"""
# Define the inputs of your model with a shape (Tx,)
# Define s0 (initial hidden state) and c0 (initial cell state)
# for the decoder LSTM with shape (n_s,)
X = Input(shape=(Tx, human_vocab_size))
s0 = Input(shape=(n_s,), name='s0')
c0 = Input(shape=(n_s,), name='c0')
s = s0
c = c0
# Initialize empty list of outputs
outputs = []
### START CODE HERE ###
# Step 1: Define your pre-attention Bi-LSTM. (≈ 1 line)
a = Bidirectional(LSTM(n_a,return_sequences=True))(X)
# Step 2: Iterate for Ty steps
for t in range(Ty):
# Step 2.A: Perform one step of the attention mechanism to get back the context vector at step t (≈ 1 line)
context = one_step_attention(a,s)
# Step 2.B: Apply the post-attention LSTM cell to the "context" vector.
# Don't forget to pass: initial_state = [hidden state, cell state] (≈ 1 line)
s, _, c = post_activation_LSTM_cell(context,initial_state=[s,c])
# Step 2.C: Apply Dense layer to the hidden state output of the post-attention LSTM (≈ 1 line)
out = output_layer(s)
# Step 2.D: Append "out" to the "outputs" list (≈ 1 line)
outputs.append(out)
# Step 3: Create model instance taking three inputs and returning the list of outputs. (≈ 1 line)
model = Model(inputs=[X,s0,c0],outputs=outputs)
### END CODE HERE ###
return model
```
Run the following cell to create your model.
```
model = model(Tx, Ty, n_a, n_s, len(human_vocab), len(machine_vocab))
```
#### Troubleshooting Note
* If you are getting repeated errors after an initially incorrect implementation of "model", but believe that you have corrected the error, you may still see error messages when building your model.
* A solution is to save and restart your kernel (or shutdown then restart your notebook), and re-run the cells.
Let's get a summary of the model to check if it matches the expected output.
```
model.summary()
```
**Expected Output**:
Here is the summary you should see
<table>
<tr>
<td>
**Total params:**
</td>
<td>
52,960
</td>
</tr>
<tr>
<td>
**Trainable params:**
</td>
<td>
52,960
</td>
</tr>
<tr>
<td>
**Non-trainable params:**
</td>
<td>
0
</td>
</tr>
<tr>
<td>
**bidirectional_1's output shape **
</td>
<td>
(None, 30, 64)
</td>
</tr>
<tr>
<td>
**repeat_vector_1's output shape **
</td>
<td>
(None, 30, 64)
</td>
</tr>
<tr>
<td>
**concatenate_1's output shape **
</td>
<td>
(None, 30, 128)
</td>
</tr>
<tr>
<td>
**attention_weights's output shape **
</td>
<td>
(None, 30, 1)
</td>
</tr>
<tr>
<td>
**dot_1's output shape **
</td>
<td>
(None, 1, 64)
</td>
</tr>
<tr>
<td>
**dense_3's output shape **
</td>
<td>
(None, 11)
</td>
</tr>
</table>
#### Compile the model
* After creating your model in Keras, you need to compile it and define the loss function, optimizer and metrics you want to use.
* Loss function: 'categorical_crossentropy'.
* Optimizer: [Adam](https://keras.io/optimizers/#adam) [optimizer](https://keras.io/optimizers/#usage-of-optimizers)
- learning rate = 0.005
- $\beta_1 = 0.9$
- $\beta_2 = 0.999$
- decay = 0.01
* metric: 'accuracy'
Sample code
```Python
optimizer = Adam(lr=..., beta_1=..., beta_2=..., decay=...)
model.compile(optimizer=..., loss=..., metrics=[...])
```
```
### START CODE HERE ### (≈2 lines)
opt = Adam(lr=0.005, beta_1=0.9, beta_2=0.999, decay=0.01)
model.compile(optimizer=opt,metrics=['accuracy'],loss='categorical_crossentropy')
### END CODE HERE ###
```
#### Define inputs and outputs, and fit the model
The last step is to define all your inputs and outputs to fit the model:
- You have input X of shape $(m = 10000, T_x = 30)$ containing the training examples.
- You need to create `s0` and `c0` to initialize your `post_attention_LSTM_cell` with zeros.
- Given the `model()` you coded, you need the "outputs" to be a list of 10 elements of shape (m, T_y).
- The list `outputs[i][0], ..., outputs[i][Ty]` represents the true labels (characters) corresponding to the $i^{th}$ training example (`X[i]`).
- `outputs[i][j]` is the true label of the $j^{th}$ character in the $i^{th}$ training example.
```
s0 = np.zeros((m, n_s))
c0 = np.zeros((m, n_s))
outputs = list(Yoh.swapaxes(0,1))
```
Let's now fit the model and run it for one epoch.
```
model.fit([Xoh, s0, c0], outputs, epochs=1, batch_size=100)
```
While training you can see the loss as well as the accuracy on each of the 10 positions of the output. The table below gives you an example of what the accuracies could be if the batch had 2 examples:
<img src="images/table.png" style="width:700;height:200px;"> <br>
<caption><center>Thus, `dense_2_acc_8: 0.89` means that you are predicting the 7th character of the output correctly 89% of the time in the current batch of data. </center></caption>
We have run this model for longer, and saved the weights. Run the next cell to load our weights. (By training a model for several minutes, you should be able to obtain a model of similar accuracy, but loading our model will save you time.)
```
model.load_weights('models/model.h5')
```
You can now see the results on new examples.
```
EXAMPLES = ['3 May 1979', '5 April 09', '21th of August 2016', 'Tue 10 Jul 2007', 'Saturday May 9 2018', 'March 3 2001', 'March 3rd 2001', '1 March 2001']
for example in EXAMPLES:
source = string_to_int(example, Tx, human_vocab)
source = np.array(list(map(lambda x: to_categorical(x, num_classes=len(human_vocab)), source))).swapaxes(0,1)
prediction = model.predict([source, s0, c0])
prediction = np.argmax(prediction, axis = -1)
output = [inv_machine_vocab[int(i)] for i in prediction]
print("source:", example)
print("output:", ''.join(output),"\n")
```
You can also change these examples to test with your own examples. The next part will give you a better sense of what the attention mechanism is doing--i.e., what part of the input the network is paying attention to when generating a particular output character.
## 3 - Visualizing Attention (Optional / Ungraded)
Since the problem has a fixed output length of 10, it is also possible to carry out this task using 10 different softmax units to generate the 10 characters of the output. But one advantage of the attention model is that each part of the output (such as the month) knows it needs to depend only on a small part of the input (the characters in the input giving the month). We can visualize what each part of the output is looking at which part of the input.
Consider the task of translating "Saturday 9 May 2018" to "2018-05-09". If we visualize the computed $\alpha^{\langle t, t' \rangle}$ we get this:
<img src="images/date_attention.png" style="width:600;height:300px;"> <br>
<caption><center> **Figure 8**: Full Attention Map</center></caption>
Notice how the output ignores the "Saturday" portion of the input. None of the output timesteps are paying much attention to that portion of the input. We also see that 9 has been translated as 09 and May has been correctly translated into 05, with the output paying attention to the parts of the input it needs to to make the translation. The year mostly requires it to pay attention to the input's "18" in order to generate "2018."
### 3.1 - Getting the attention weights from the network
Lets now visualize the attention values in your network. We'll propagate an example through the network, then visualize the values of $\alpha^{\langle t, t' \rangle}$.
To figure out where the attention values are located, let's start by printing a summary of the model .
```
model.summary()
```
Navigate through the output of `model.summary()` above. You can see that the layer named `attention_weights` outputs the `alphas` of shape (m, 30, 1) before `dot_2` computes the context vector for every time step $t = 0, \ldots, T_y-1$. Let's get the attention weights from this layer.
The function `attention_map()` pulls out the attention values from your model and plots them.
```
attention_map = plot_attention_map(model, human_vocab, inv_machine_vocab, "Tuesday 09 Oct 1993", num = 7, n_s = 64);
```
On the generated plot you can observe the values of the attention weights for each character of the predicted output. Examine this plot and check that the places where the network is paying attention makes sense to you.
In the date translation application, you will observe that most of the time attention helps predict the year, and doesn't have much impact on predicting the day or month.
### Congratulations!
You have come to the end of this assignment
## Here's what you should remember
- Machine translation models can be used to map from one sequence to another. They are useful not just for translating human languages (like French->English) but also for tasks like date format translation.
- An attention mechanism allows a network to focus on the most relevant parts of the input when producing a specific part of the output.
- A network using an attention mechanism can translate from inputs of length $T_x$ to outputs of length $T_y$, where $T_x$ and $T_y$ can be different.
- You can visualize attention weights $\alpha^{\langle t,t' \rangle}$ to see what the network is paying attention to while generating each output.
Congratulations on finishing this assignment! You are now able to implement an attention model and use it to learn complex mappings from one sequence to another.
| github_jupyter |
##### Copyright 2020 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Calculate gradients
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/quantum/tutorials/gradients"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/quantum/blob/master/docs/tutorials/gradients.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/quantum/blob/master/docs/tutorials/gradients.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/quantum/docs/tutorials/gradients.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
This tutorial explores gradient calculation algorithms for the expectation values of quantum circuits.
Calculating the gradient of the expectation value of a certain observable in a quantum circuit is an involved process. Expectation values of observables do not have the luxury of having analytic gradient formulas that are always easy to write down—unlike traditional machine learning transformations such as matrix multiplication or vector addition that have analytic gradient formulas which are easy to write down. As a result, there are different quantum gradient calculation methods that come in handy for different scenarios. This tutorial compares and contrasts two different differentiation schemes.
## Setup
```
try:
%tensorflow_version 2.x
except Exception:
pass
```
Install TensorFlow Quantum:
```
!pip install tensorflow-quantum
```
Now import TensorFlow and the module dependencies:
```
import tensorflow as tf
import tensorflow_quantum as tfq
import cirq
import sympy
import numpy as np
# visualization tools
%matplotlib inline
import matplotlib.pyplot as plt
from cirq.contrib.svg import SVGCircuit
```
## 1. Preliminary
Let's make the notion of gradient calculation for quantum circuits a little more concrete. Suppose you have a parameterized circuit like this one:
```
qubit = cirq.GridQubit(0, 0)
my_circuit = cirq.Circuit(cirq.Y(qubit)**sympy.Symbol('alpha'))
SVGCircuit(my_circuit)
```
Along with an observable:
```
pauli_x = cirq.X(qubit)
pauli_x
```
Looking at this operator you know that $⟨Y(\alpha)| X | Y(\alpha)⟩ = \sin(\pi \alpha)$
```
def my_expectation(op, alpha):
"""Compute ⟨Y(alpha)| `op` | Y(alpha)⟩"""
params = {'alpha': alpha}
sim = cirq.Simulator()
final_state = sim.simulate(my_circuit, params).final_state
return op.expectation_from_wavefunction(final_state, {qubit: 0}).real
my_alpha = 0.3
print("Expectation=", my_expectation(pauli_x, my_alpha))
print("Sin Formula=", np.sin(np.pi * my_alpha))
```
and if you define $f_{1}(\alpha) = ⟨Y(\alpha)| X | Y(\alpha)⟩$ then $f_{1}^{'}(\alpha) = \pi \cos(\pi \alpha)$. Let's check this:
```
def my_grad(obs, alpha, eps=0.01):
grad = 0
f_x = my_expectation(obs, alpha)
f_x_prime = my_expectation(obs, alpha + eps)
return ((f_x_prime - f_x) / eps).real
print('Finite difference:', my_grad(pauli_x, my_alpha))
print('Cosine formula: ', np.pi * np.cos(np.pi * my_alpha))
```
## 2. The need for a differentiator
With larger circuits, you won't always be so lucky to have a formula that precisely calculates the gradients of a given quantum circuit. In the event that a simple formula isn't enough to calculate the gradient, the `tfq.differentiators.Differentiator` class allows you to define algorithms for computing the gradients of your circuits. For instance you can recreate the above example in TensorFlow Quantum (TFQ) with:
```
expectation_calculation = tfq.layers.Expectation(
differentiator=tfq.differentiators.ForwardDifference(grid_spacing=0.01))
expectation_calculation(my_circuit,
operators=pauli_x,
symbol_names=['alpha'],
symbol_values=[[my_alpha]])
```
However, if you switch to estimating expectation based on sampling (what would happen on a true device) the values can change a little bit. This means you now have an imperfect estimate:
```
sampled_expectation_calculation = tfq.layers.SampledExpectation(
differentiator=tfq.differentiators.ForwardDifference(grid_spacing=0.01))
sampled_expectation_calculation(my_circuit,
operators=pauli_x,
repetitions=500,
symbol_names=['alpha'],
symbol_values=[[my_alpha]])
```
This can quickly compound into a serious accuracy problem when it comes to gradients:
```
# Make input_points = [batch_size, 1] array.
input_points = np.linspace(0, 5, 200)[:, np.newaxis].astype(np.float32)
exact_outputs = expectation_calculation(my_circuit,
operators=pauli_x,
symbol_names=['alpha'],
symbol_values=input_points)
imperfect_outputs = sampled_expectation_calculation(my_circuit,
operators=pauli_x,
repetitions=500,
symbol_names=['alpha'],
symbol_values=input_points)
plt.title('Forward Pass Values')
plt.xlabel('$x$')
plt.ylabel('$f(x)$')
plt.plot(input_points, exact_outputs, label='Analytic')
plt.plot(input_points, imperfect_outputs, label='Sampled')
plt.legend()
# Gradients are a much different story.
values_tensor = tf.convert_to_tensor(input_points)
with tf.GradientTape() as g:
g.watch(values_tensor)
exact_outputs = expectation_calculation(my_circuit,
operators=pauli_x,
symbol_names=['alpha'],
symbol_values=values_tensor)
analytic_finite_diff_gradients = g.gradient(exact_outputs, values_tensor)
with tf.GradientTape() as g:
g.watch(values_tensor)
imperfect_outputs = sampled_expectation_calculation(
my_circuit,
operators=pauli_x,
repetitions=500,
symbol_names=['alpha'],
symbol_values=values_tensor)
sampled_finite_diff_gradients = g.gradient(imperfect_outputs, values_tensor)
plt.title('Gradient Values')
plt.xlabel('$x$')
plt.ylabel('$f^{\'}(x)$')
plt.plot(input_points, analytic_finite_diff_gradients, label='Analytic')
plt.plot(input_points, sampled_finite_diff_gradients, label='Sampled')
plt.legend()
```
Here you can see that although the finite difference formula is fast to compute the gradients themselves in the analytical case, when it came to the sampling based methods it was far too noisy. More careful techniques must be used to ensure a good gradient can be calculated. Next you will look at a much slower technique that wouldn't be as well suited for analytical expectation gradient calculations, but does perform much better in the real-world sample based case:
```
# A smarter differentiation scheme.
gradient_safe_sampled_expectation = tfq.layers.SampledExpectation(
differentiator=tfq.differentiators.ParameterShift())
with tf.GradientTape() as g:
g.watch(values_tensor)
imperfect_outputs = gradient_safe_sampled_expectation(
my_circuit,
operators=pauli_x,
repetitions=500,
symbol_names=['alpha'],
symbol_values=values_tensor)
sampled_param_shift_gradients = g.gradient(imperfect_outputs, values_tensor)
plt.title('Gradient Values')
plt.xlabel('$x$')
plt.ylabel('$f^{\'}(x)$')
plt.plot(input_points, analytic_finite_diff_gradients, label='Analytic')
plt.plot(input_points, sampled_param_shift_gradients, label='Sampled')
plt.legend()
```
From the above you can see that certain differentiators are best used for particular research scenarios. In general, the slower sample-based methods that are robust to device noise, etc., are great differentiators when testing or implementing algorithms in a more "real world" setting. Faster methods like finite difference are great for analytical calculations and you want higher throughput, but aren't yet concerned with the device viability of your algorithm.
## 3. Multiple observables
Let's introduce a second observable and see how TensorFlow Quantum supports multiple observables for a single circuit.
```
pauli_z = cirq.Z(qubit)
pauli_z
```
If this observable is used with the same circuit as before, then you have $f_{2}(\alpha) = ⟨Y(\alpha)| Z | Y(\alpha)⟩ = \cos(\pi \alpha)$ and $f_{2}^{'}(\alpha) = -\pi \sin(\pi \alpha)$. Perform a quick check:
```
test_value = 0.
print('Finite difference:', my_grad(pauli_z, test_value))
print('Sin formula: ', -np.pi * np.sin(np.pi * test_value))
```
It's a match (close enough).
Now if you define $g(\alpha) = f_{1}(\alpha) + f_{2}(\alpha)$ then $g'(\alpha) = f_{1}^{'}(\alpha) + f^{'}_{2}(\alpha)$. Defining more than one observable in TensorFlow Quantum to use along with a circuit is equivalent to adding on more terms to $g$.
This means that the gradient of a particular symbol in a circuit is equal to the sum of the gradients with regards to each observable for that symbol applied to that circuit. This is compatible with TensorFlow gradient taking and backpropagation (where you give the sum of the gradients over all observables as the gradient for a particular symbol).
```
sum_of_outputs = tfq.layers.Expectation(
differentiator=tfq.differentiators.ForwardDifference(grid_spacing=0.01))
sum_of_outputs(my_circuit,
operators=[pauli_x, pauli_z],
symbol_names=['alpha'],
symbol_values=[[test_value]])
```
Here you see the first entry is the expectation w.r.t Pauli X, and the second is the expectation w.r.t Pauli Z. Now when you take the gradient:
```
test_value_tensor = tf.convert_to_tensor([[test_value]])
with tf.GradientTape() as g:
g.watch(test_value_tensor)
outputs = sum_of_outputs(my_circuit,
operators=[pauli_x, pauli_z],
symbol_names=['alpha'],
symbol_values=test_value_tensor)
sum_of_gradients = g.gradient(outputs, test_value_tensor)
print(my_grad(pauli_x, test_value) + my_grad(pauli_z, test_value))
print(sum_of_gradients.numpy())
```
Here you have verified that the sum of the gradients for each observable is indeed the gradient of $\alpha$. This behavior is supported by all TensorFlow Quantum differentiators and plays a crucial role in the compatibility with the rest of TensorFlow.
## 4. Advanced usage
Here you will learn how to define your own custom differentiation routines for quantum circuits.
All differentiators that exist inside of TensorFlow Quantum subclass `tfq.differentiators.Differentiator`. A differentiator must implement `differentiate_analytic` and `differentiate_sampled`.
The following uses TensorFlow Quantum constructs to implement the closed form solution from the first part of this tutorial.
```
class MyDifferentiator(tfq.differentiators.Differentiator):
"""A Toy differentiator for <Y^alpha | X |Y^alpha>."""
def __init__(self):
pass
@tf.function
def _compute_gradient(self, symbol_values):
"""Compute the gradient based on symbol_values."""
# f(x) = sin(pi * x)
# f'(x) = pi * cos(pi * x)
return tf.cast(tf.cos(symbol_values * np.pi) * np.pi, tf.float32)
@tf.function
def differentiate_analytic(self, programs, symbol_names, symbol_values,
pauli_sums, forward_pass_vals, grad):
"""Specify how to differentiate a circuit with analytical expectation.
This is called at graph runtime by TensorFlow. `differentiate_analytic`
should calculate the gradient of a batch of circuits and return it
formatted as indicated below. See
`tfq.differentiators.ForwardDifference` for an example.
Args:
programs: `tf.Tensor` of strings with shape [batch_size] containing
the string representations of the circuits to be executed.
symbol_names: `tf.Tensor` of strings with shape [n_params], which
is used to specify the order in which the values in
`symbol_values` should be placed inside of the circuits in
`programs`.
symbol_values: `tf.Tensor` of real numbers with shape
[batch_size, n_params] specifying parameter values to resolve
into the circuits specified by programs, following the ordering
dictated by `symbol_names`.
pauli_sums: `tf.Tensor` of strings with shape [batch_size, n_ops]
containing the string representation of the operators that will
be used on all of the circuits in the expectation calculations.
forward_pass_vals: `tf.Tensor` of real numbers with shape
[batch_size, n_ops] containing the output of the forward pass
through the op you are differentiating.
grad: `tf.Tensor` of real numbers with shape [batch_size, n_ops]
representing the gradient backpropagated to the output of the
op you are differentiating through.
Returns:
A `tf.Tensor` with the same shape as `symbol_values` representing
the gradient backpropagated to the `symbol_values` input of the op
you are differentiating through.
"""
# Computing gradients just based off of symbol_values.
return self._compute_gradient(symbol_values) * grad
@tf.function
def differentiate_sampled(self, programs, symbol_names, symbol_values,
pauli_sums, num_samples, forward_pass_vals, grad):
"""Specify how to differentiate a circuit with sampled expectation.
This is called at graph runtime by TensorFlow. `differentiate_sampled`
should calculate the gradient of a batch of circuits and return it
formatted as indicated below. See
`tfq.differentiators.ForwardDifference` for an example.
Args:
programs: `tf.Tensor` of strings with shape [batch_size] containing
the string representations of the circuits to be executed.
symbol_names: `tf.Tensor` of strings with shape [n_params], which
is used to specify the order in which the values in
`symbol_values` should be placed inside of the circuits in
`programs`.
symbol_values: `tf.Tensor` of real numbers with shape
[batch_size, n_params] specifying parameter values to resolve
into the circuits specified by programs, following the ordering
dictated by `symbol_names`.
pauli_sums: `tf.Tensor` of strings with shape [batch_size, n_ops]
containing the string representation of the operators that will
be used on all of the circuits in the expectation calculations.
num_samples: `tf.Tensor` of positive integers representing the
number of samples per term in each term of pauli_sums used
during the forward pass.
forward_pass_vals: `tf.Tensor` of real numbers with shape
[batch_size, n_ops] containing the output of the forward pass
through the op you are differentiating.
grad: `tf.Tensor` of real numbers with shape [batch_size, n_ops]
representing the gradient backpropagated to the output of the
op you are differentiating through.
Returns:
A `tf.Tensor` with the same shape as `symbol_values` representing
the gradient backpropagated to the `symbol_values` input of the op
you are differentiating through.
"""
return self._compute_gradient(symbol_values) * grad
```
This new differentiator can now be used with existing `tfq.layer` objects:
```
custom_dif = MyDifferentiator()
custom_grad_expectation = tfq.layers.Expectation(differentiator=custom_dif)
# Now let's get the gradients with finite diff.
with tf.GradientTape() as g:
g.watch(values_tensor)
exact_outputs = expectation_calculation(my_circuit,
operators=[pauli_x],
symbol_names=['alpha'],
symbol_values=values_tensor)
analytic_finite_diff_gradients = g.gradient(exact_outputs, values_tensor)
# Now let's get the gradients with custom diff.
with tf.GradientTape() as g:
g.watch(values_tensor)
my_outputs = custom_grad_expectation(my_circuit,
operators=[pauli_x],
symbol_names=['alpha'],
symbol_values=values_tensor)
my_gradients = g.gradient(my_outputs, values_tensor)
plt.subplot(1, 2, 1)
plt.title('Exact Gradient')
plt.plot(input_points, analytic_finite_diff_gradients.numpy())
plt.xlabel('x')
plt.ylabel('f(x)')
plt.subplot(1, 2, 2)
plt.title('My Gradient')
plt.plot(input_points, my_gradients.numpy())
plt.xlabel('x')
```
This new differentiator can now be used to generate differentiable ops.
Key Point: A differentiator that has been previously attached to an op must be refreshed before attaching to a new op, because a differentiator may only be attached to one op at a time.
```
# Create a noisy sample based expectation op.
expectation_sampled = tfq.get_sampled_expectation_op(
cirq.DensityMatrixSimulator(noise=cirq.depolarize(0.01)))
# Make it differentiable with your differentiator:
# Remember to refresh the differentiator before attaching the new op
custom_dif.refresh()
differentiable_op = custom_dif.generate_differentiable_op(
sampled_op=expectation_sampled)
# Prep op inputs.
circuit_tensor = tfq.convert_to_tensor([my_circuit])
op_tensor = tfq.convert_to_tensor([[pauli_x]])
single_value = tf.convert_to_tensor([[my_alpha]])
num_samples_tensor = tf.convert_to_tensor([[1000]])
with tf.GradientTape() as g:
g.watch(single_value)
forward_output = differentiable_op(circuit_tensor, ['alpha'], single_value,
op_tensor, num_samples_tensor)
my_gradients = g.gradient(forward_output, single_value)
print('---TFQ---')
print('Foward: ', forward_output.numpy())
print('Gradient:', my_gradients.numpy())
print('---Original---')
print('Forward: ', my_expectation(pauli_x, my_alpha))
print('Gradient:', my_grad(pauli_x, my_alpha))
```
Success: Now you can use all the differentiators that TensorFlow Quantum has to offer—and define your own.
| github_jupyter |
```
import os
os.environ['CUDA_VISIBLE_DEVICES'] = '2'
import numpy as np
import tensorflow as tf
import json
with open('train-test.json') as fopen:
dataset = json.load(fopen)
with open('dictionary.json') as fopen:
dictionary = json.load(fopen)
train_X = dataset['train_X']
train_Y = dataset['train_Y']
test_X = dataset['test_X']
test_Y = dataset['test_Y']
dictionary.keys()
dictionary_from = dictionary['from']['dictionary']
rev_dictionary_from = dictionary['from']['rev_dictionary']
dictionary_to = dictionary['to']['dictionary']
rev_dictionary_to = dictionary['to']['rev_dictionary']
GO = dictionary_from['GO']
PAD = dictionary_from['PAD']
EOS = dictionary_from['EOS']
UNK = dictionary_from['UNK']
for i in range(len(train_X)):
train_X[i] += ' EOS'
train_X[0]
for i in range(len(test_X)):
test_X[i] += ' EOS'
test_X[0]
def pad_second_dim(x, desired_size):
padding = tf.tile([[[0.0]]], tf.stack([tf.shape(x)[0], desired_size - tf.shape(x)[1], tf.shape(x)[2]], 0))
return tf.concat([x, padding], 1)
class Translator:
def __init__(self, size_layer, num_layers, embedded_size,
from_dict_size, to_dict_size, learning_rate,
beam_width=5, force_teaching_ratio=0.5):
def lstm_cell(size, reuse=False):
return tf.nn.rnn_cell.LSTMCell(size, initializer=tf.orthogonal_initializer(),reuse=reuse)
self.X = tf.placeholder(tf.int32, [None, None])
self.Y = tf.placeholder(tf.int32, [None, None])
self.X_seq_len = tf.count_nonzero(self.X, 1, dtype=tf.int32)
self.Y_seq_len = tf.count_nonzero(self.Y, 1, dtype=tf.int32)
batch_size = tf.shape(self.X)[0]
encoder_embeddings = tf.Variable(tf.random_uniform([from_dict_size, embedded_size], -1, 1))
decoder_embeddings = tf.Variable(tf.random_uniform([to_dict_size, embedded_size], -1, 1))
self.encoder_out = tf.nn.embedding_lookup(encoder_embeddings, self.X)
for n in range(num_layers):
(out_fw, out_bw), (state_fw, state_bw) = tf.nn.bidirectional_dynamic_rnn(
cell_fw = lstm_cell(size_layer // 2),
cell_bw = lstm_cell(size_layer // 2),
inputs = self.encoder_out,
sequence_length = self.X_seq_len,
dtype = tf.float32,
scope = 'bidirectional_rnn_%d'%(n))
self.encoder_out = tf.concat((out_fw, out_bw), 2)
bi_state_c = tf.concat((state_fw.c, state_bw.c), -1)
bi_state_h = tf.concat((state_fw.h, state_bw.h), -1)
bi_lstm_state = tf.nn.rnn_cell.LSTMStateTuple(c=bi_state_c, h=bi_state_h)
encoder_state = tuple([bi_lstm_state] * num_layers)
with tf.variable_scope('decode'):
attention_mechanism = tf.contrib.seq2seq.BahdanauMonotonicAttention(
num_units = size_layer,
memory = self.encoder_out,
memory_sequence_length = self.X_seq_len)
decoder_cell = tf.contrib.seq2seq.AttentionWrapper(
cell = tf.nn.rnn_cell.MultiRNNCell([lstm_cell(size_layer) for _ in range(num_layers)]),
attention_mechanism = attention_mechanism,
attention_layer_size = size_layer)
main = tf.strided_slice(self.Y, [0, 0], [batch_size, -1], [1, 1])
decoder_input = tf.concat([tf.fill([batch_size, 1], GO), main], 1)
training_helper = tf.contrib.seq2seq.ScheduledEmbeddingTrainingHelper(
inputs = tf.nn.embedding_lookup(decoder_embeddings, decoder_input),
sequence_length = self.Y_seq_len,
embedding = decoder_embeddings,
sampling_probability = 1 - force_teaching_ratio,
time_major = False)
training_decoder = tf.contrib.seq2seq.BasicDecoder(
cell = decoder_cell,
helper = training_helper,
initial_state = decoder_cell.zero_state(batch_size, tf.float32).clone(cell_state=encoder_state),
output_layer = tf.layers.Dense(to_dict_size))
training_decoder_output, _, _ = tf.contrib.seq2seq.dynamic_decode(
decoder = training_decoder,
impute_finished = True,
maximum_iterations = tf.reduce_max(self.Y_seq_len))
self.training_logits = training_decoder_output.rnn_output
with tf.variable_scope('decode', reuse=True):
encoder_out_tiled = tf.contrib.seq2seq.tile_batch(self.encoder_out, beam_width)
encoder_state_tiled = tf.contrib.seq2seq.tile_batch(encoder_state, beam_width)
X_seq_len_tiled = tf.contrib.seq2seq.tile_batch(self.X_seq_len, beam_width)
attention_mechanism = tf.contrib.seq2seq.BahdanauMonotonicAttention(
num_units = size_layer,
memory = encoder_out_tiled,
memory_sequence_length = X_seq_len_tiled)
decoder_cell = tf.contrib.seq2seq.AttentionWrapper(
cell = tf.nn.rnn_cell.MultiRNNCell([lstm_cell(size_layer, reuse=True) for _ in range(num_layers)]),
attention_mechanism = attention_mechanism,
attention_layer_size = size_layer)
predicting_decoder = tf.contrib.seq2seq.BeamSearchDecoder(
cell = decoder_cell,
embedding = decoder_embeddings,
start_tokens = tf.tile(tf.constant([GO], dtype=tf.int32), [batch_size]),
end_token = EOS,
initial_state = decoder_cell.zero_state(batch_size * beam_width, tf.float32).clone(cell_state = encoder_state_tiled),
beam_width = beam_width,
output_layer = tf.layers.Dense(to_dict_size, _reuse=True),
length_penalty_weight = 0.0)
predicting_decoder_output, _, _ = tf.contrib.seq2seq.dynamic_decode(
decoder = predicting_decoder,
impute_finished = False,
maximum_iterations = 2 * tf.reduce_max(self.X_seq_len))
self.predicting_ids = predicting_decoder_output.predicted_ids[:, :, 0]
masks = tf.sequence_mask(self.Y_seq_len, tf.reduce_max(self.Y_seq_len), dtype=tf.float32)
self.cost = tf.contrib.seq2seq.sequence_loss(logits = self.training_logits,
targets = self.Y,
weights = masks)
self.optimizer = tf.train.AdamOptimizer(learning_rate).minimize(self.cost)
y_t = tf.argmax(self.training_logits,axis=2)
y_t = tf.cast(y_t, tf.int32)
self.prediction = tf.boolean_mask(y_t, masks)
mask_label = tf.boolean_mask(self.Y, masks)
correct_pred = tf.equal(self.prediction, mask_label)
correct_index = tf.cast(correct_pred, tf.float32)
self.accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
size_layer = 256
num_layers = 2
embedded_size = 256
learning_rate = 1e-3
batch_size = 128
epoch = 20
tf.reset_default_graph()
sess = tf.InteractiveSession()
model = Translator(size_layer, num_layers, embedded_size, len(dictionary_from),
len(dictionary_to), learning_rate)
sess.run(tf.global_variables_initializer())
def str_idx(corpus, dic):
X = []
for i in corpus:
ints = []
for k in i.split():
ints.append(dic.get(k,UNK))
X.append(ints)
return X
def pad_sentence_batch(sentence_batch, pad_int):
padded_seqs = []
seq_lens = []
max_sentence_len = max([len(sentence) for sentence in sentence_batch])
for sentence in sentence_batch:
padded_seqs.append(sentence + [pad_int] * (max_sentence_len - len(sentence)))
seq_lens.append(len(sentence))
return padded_seqs, seq_lens
train_X = str_idx(train_X, dictionary_from)
test_X = str_idx(test_X, dictionary_from)
train_Y = str_idx(train_Y, dictionary_to)
test_Y = str_idx(test_Y, dictionary_to)
import tqdm
for e in range(epoch):
pbar = tqdm.tqdm(
range(0, len(train_X), batch_size), desc = 'minibatch loop')
train_loss, train_acc, test_loss, test_acc = [], [], [], []
for i in pbar:
index = min(i + batch_size, len(train_X))
maxlen = max([len(s) for s in train_X[i : index] + train_Y[i : index]])
batch_x, seq_x = pad_sentence_batch(train_X[i : index], PAD)
batch_y, seq_y = pad_sentence_batch(train_Y[i : index], PAD)
feed = {model.X: batch_x,
model.Y: batch_y}
accuracy, loss, _ = sess.run([model.accuracy,model.cost,model.optimizer],
feed_dict = feed)
train_loss.append(loss)
train_acc.append(accuracy)
pbar.set_postfix(cost = loss, accuracy = accuracy)
pbar = tqdm.tqdm(
range(0, len(test_X), batch_size), desc = 'minibatch loop')
for i in pbar:
index = min(i + batch_size, len(test_X))
batch_x, seq_x = pad_sentence_batch(test_X[i : index], PAD)
batch_y, seq_y = pad_sentence_batch(test_Y[i : index], PAD)
feed = {model.X: batch_x,
model.Y: batch_y,}
accuracy, loss = sess.run([model.accuracy,model.cost],
feed_dict = feed)
test_loss.append(loss)
test_acc.append(accuracy)
pbar.set_postfix(cost = loss, accuracy = accuracy)
print('epoch %d, training avg loss %f, training avg acc %f'%(e+1,
np.mean(train_loss),np.mean(train_acc)))
print('epoch %d, testing avg loss %f, testing avg acc %f'%(e+1,
np.mean(test_loss),np.mean(test_acc)))
rev_dictionary_to = {int(k): v for k, v in rev_dictionary_to.items()}
test_size = 20
batch_x, seq_x = pad_sentence_batch(test_X[: test_size], PAD)
batch_y, seq_y = pad_sentence_batch(test_Y[: test_size], PAD)
feed = {model.X: batch_x}
logits = sess.run(model.predicting_ids, feed_dict = feed)
logits.shape
rejected = ['PAD', 'EOS', 'UNK', 'GO']
for i in range(test_size):
predict = [rev_dictionary_to[i] for i in logits[i] if rev_dictionary_to[i] not in rejected]
actual = [rev_dictionary_to[i] for i in batch_y[i] if rev_dictionary_to[i] not in rejected]
print(i, 'predict:', ' '.join(predict))
print(i, 'actual:', ' '.join(actual))
print()
```
| github_jupyter |
```
import matplotlib
matplotlib.use('nbagg')
import matplotlib.animation as anm
import matplotlib.pyplot as plt
import math
import matplotlib.patches as patches
import numpy as np
class World:
def __init__(self, debug=False):
self.objects = []
self.debug = debug
def append(self,obj): # オブジェクトを登録するための関数
self.objects.append(obj)
def draw(self):
fig = plt.figure(figsize=(4,4)) # 8x8 inchの図を準備
ax = fig.add_subplot(111) # サブプロットを準備
ax.set_aspect('equal') # 縦横比を座標の値と一致させる
ax.set_xlim(-5,5) # X軸を-5m x 5mの範囲で描画
ax.set_ylim(-5,5) # Y軸も同様に
ax.set_xlabel("X",fontsize=10) # X軸にラベルを表示
ax.set_ylabel("Y",fontsize=10) # 同じくY軸に
elems = []
if self.debug:
for i in range(1000): self.one_step(i, elems, ax)
else:
self.ani = anm.FuncAnimation(fig, self.one_step, fargs=(elems, ax), frames=100, interval=1000, repeat=False)
plt.show()
def one_step(self, i, elems, ax): ### fig:one_step_add_one_step
while elems: elems.pop().remove()
elems.append(ax.text(-4.4, 4.5, "t = "+str(i), fontsize=10))
for obj in self.objects:
obj.draw(ax, elems)
if hasattr(obj, "one_step"): obj.one_step(1.0) # 追加
class IdealRobot: ### fig:rewrite_init_for_agent
def __init__(self, pose, agent=None, color="black"): # agentという引数を追加
self.pose = pose
self.r = 0.2
self.color = color
self.agent = agent # 追加
self.poses = [pose] # 軌跡の描画用。追加
def draw(self, ax, elems): ###idealrobot6draw
x, y, theta = self.pose #ここから15行目までは変えなくて良い
xn = x + self.r * math.cos(theta)
yn = y + self.r * math.sin(theta)
elems += ax.plot([x,xn], [y,yn], color=self.color)
c = patches.Circle(xy=(x, y), radius=self.r, fill=False, color=self.color)
elems.append(ax.add_patch(c))
self.poses.append(self.pose) #以下追加。軌跡の描画
elems += ax.plot([e[0] for e in self.poses], [e[1] for e in self.poses], linewidth=0.5, color="black")
@classmethod
def state_transition(cls, nu, omega, time, pose): ### fig:state_transition(20-35行目)
t0 = pose[2]
if math.fabs(omega) < 1e-10: #角速度がほぼゼロの場合とそうでない場合で場合分け
return pose + np.array( [nu*math.cos(t0),
nu*math.sin(t0),
omega ] ) * time
else:
return pose + np.array( [nu/omega*(math.sin(t0 + omega*time) - math.sin(t0)),
nu/omega*(-math.cos(t0 + omega*time) + math.cos(t0)),
omega*time ] )
def one_step(self, time_interval): ### fig:robot_one_step
if not self.agent: return
nu, omega = self.agent.decision()
self.pose = self.state_transition(nu, omega, time_interval, self.pose)
class Agent: ### fig:Agent
def __init__(self, nu, omega):
self.nu = nu
self.omega = omega
def decision(self, observation=None):
return self.nu, self.omega
world = World() ### fig:rewrite_robot_for_agent
straight = Agent(0.2, 0.0) # 0.2[m/s]で直進
circling = Agent(0.2, 10.0/180*math.pi) # 0.2[m/s], 10[deg/s](円を描く)
robot1 = IdealRobot( np.array([ 2, 3, math.pi/6]).T, straight )
robot2 = IdealRobot( np.array([-2, -1, math.pi/5*6]).T, circling, "red")
robot3 = IdealRobot( np.array([ 0, 0, 0]).T, color="blue") # エージェントを与えないロボット
world.append(robot1)
world.append(robot2)
world.append(robot3)
world.draw()
## 原点から0.1[m/s]で1[s]直進 ## ### fig:using_state_transition(セル6まで)
IdealRobot.state_transition(0.1, 0.0, 1.0, np.array([0,0,0]).T)
## 原点から0.1[m/s], 10[deg/s]で9[s]移動 ##
IdealRobot.state_transition(0.1, 10.0/180*math.pi, 9.0, np.array([0,0,0]).T)
## 原点から0.1[m/s], 10[deg/s]で18[s]移動 ##
IdealRobot.state_transition(0.1, 10.0/180*math.pi, 18.0, np.array([0,0,0]).T)
```
| github_jupyter |
# Activity: Logistic Functions
```
from cyllene import *
from sympy import solve,log
from f_special import logistic_function, logistic_plot_L, second_derivative
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from scipy.optimize import curve_fit
%matplotlib inline
from ipywidgets import interact, interactive
from IPython.display import Image
```
## The definition of a logistic function
A *logistic function* is a function of the form
$f(x) = \dfrac{L}{1+C \, e^{-kx}},$
where $L, C, k$ are *parameters*.
Logistic functions *initially* exhibit growth that is similar to *exponential*, but after a while the growth begins to slow down as a saturation property takes hold of the growth process.
```
f = function('1/(1+10*exp(-x))')
graph(f(x))
```
### The parameter $L$
Let us take a look at the paramater $L$. Use the slider below to change the value of $L$.
What effect does this have on the graph of a logistic function?
```
interactive_plot = interactive(logistic_plot_L, L=(1, 8, 0.5))
output = interactive_plot.children[-1]
output.layout.height = '300px'
interactive_plot
```
We see that $L$ determines the **height of the graph**. It is the **limiting value** of the logistic function. The values of $f$ will approach $L$ as $x$ goes to infinity. The parameter $L$ is also called the **carrying capacity**.
## The inflection point
Let us again consider a simple logistic function:
```
f = function('8/(1+10*exp(-2*x))')
```
To find possible inflection points, we need to compute the second derivative:
```
f2 = second_derivative(f(x))
f2
```
Try to find the zeros for this expression.
You can check your answer by running the code below:
```
p = solve(f2,x)[0]
solve(f2,x)[0]
```
Now plug the solution into the original function:
```
f(p)
```
Repeat this with different values for the parameters $L, C, k$. Do you observe a relation between $L$ and the location of the inflection point?
### **FACT**: The inflection point of a logistic function occurs where $f(x) = L/2$.
```
Image(filename='logistic_function.png', width=600)
```
---
## Logistic modeling of COVID-19 data
In the following cell, we read the current Coronavirus data from CSSE at Johns Hopkins University. We extract a time series for the US, giving us for each day since January 22 the total number of infected persons.
```
url = "https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_confirmed_global.csv"
df = pd.read_csv(url)
# country = df[df["Country/Region"] == "Italy"]
# country = df[df["Country/Region"] == "Sweden"]
# country = df[df["Country/Region"] == "Germany"]
country = df[df["Country/Region"] == "US"]
interval_len = 200
x_data = list(range(interval_len))
y_data = list(country.iloc[int(0),int(4):int(interval_len+4)])
```
We run a logistic regression for our data and the logistic model.
```
fit = curve_fit(logistic_function, x_data, y_data)
k = round(fit[0][0],3)
C = round(fit[0][1],3)
L = int(fit[0][2])
print("We estimated the following parameters:")
print("k = ", k)
print("C = ", C)
print("L = ", L)
```
### Question: What does the parameter $L$ reflect in this case?
#### Plotting the data
```
plt.figure(figsize=(10,8))
plt.scatter(list(range(interval_len)),y_data,label="Real data",color="red")
# Predicted logistic curve
t = np.arange(0, interval_len, 1)
s = L/(1+C*np.exp(-k*t))
plt.plot(t,s)
plt.xlabel("Days since January 22")
plt.ylabel("Total number of infected people")
plt.show()
```
| github_jupyter |
# Files
[Python File I/O
](https://www.programiz.com/python-programming/file-operation)
## Open Files
```
f = open('file.txt')
```
### Python File Modes
| Mode | Description |
|---|---|
| 'r' | Open a file for reading. (default) |
| 'w' | Open a file for writing. Creates a new file if it does not exist or truncates the file if it exists. |
| 'x' | Open a file for exclusive creation. If the file already exists, the operation fails. |
| 'a' | Open for appending at the end of the file without truncating it. Creates a new file if it does not exist. |
| 't' | Open in text mode. (default) |
| 'b' | Open in binary mode. |
| '+' | Open a file for updating (reading and writing) |
```
f = open("test.txt") # equivalent to 'r' or 'rt'
f = open("test.txt", 'w') # write in text mode
f = open("python.png",'r+b') # read and write in binary mode
```
### File Encodings
Unlike other languages, the character 'a' does not imply the number 97 until it is encoded using ASCII (or other equivalent encodings). The default encoding is platform dependent.
* In Windows it is 'cp1252'
* In Linux it is 'utf-8'
You cannot rely on the default encoding or else our code will behave differently in different platforms. It is highly recommended to specify the encoding type when working with text files.
```
f = open("file.txt", mode='r', encoding='utf-8')
```
## Closing Files
```
f = open("file.txt", encoding='utf-8')
# perform file operations
f.close()
```
In the last example exceptions could crash your program. Exception handlig is a topic in itself, but here we'll simply wrap the code in a try...catch block. Now the file handle will be closed regardless of whether or not there was an exception. In the example above, an exeption might result in the file remaining open (and potentially locked) from other applications in the event of an exception.
```
try:
f = open("file.txt",encoding = 'utf-8')
# perform file operations
finally:
f.close()
```
## Using with
The best way to ensure that you close your files when the code block terminates is by using the with statement. When using the with statement you do not have to explicitly call the file's close() method.
```
with open('file.txt') as f:
text = f.read()
print(text)
# f.close() # not necessary
```
## Writing Files
```
with open("file.txt", 'w', encoding='utf-8') as f:
print(type(f))
f.write("hello class\n" )
f.write("python is fun\n\n")
f.write("let's get started\n")
```
## Reading Files
```
f = open("file.txt", 'r', encoding='utf-8')
print('1:', f.read(3)) # read the 1st 3 characters
print('2:', f.read(3)) # read the next 3 characters
print('3:', f.read()) # read until the end
print('4:', f.read()) # return's an empty string
f.tell() # get the current file position
f.seek(0) # bring file cursor to initial position
f.read() # read the entire file
f.seek(0) # reset the cursor
# read line by line
for line in f:
print(line, end='')
f.seek(0) # reset the cursor
# read line by line
f.readline()
f.readline()
f.readline()
line = f.readline()
print(type(line), line)
f.readline()
f.seek(0) # reset the cursor
# read all lines at once
f.readlines()
```
## Some Notes on Printing
```
print('abcd')
print('efgh')
print('abcd', end='')
print('efgh')
text = ' abcd '
print(text, end='')
print(text)
text = ' abcd '
text = text.strip()
print(text, end='')
print(text)
print?
print('abc', 'def', 'xyz', sep=',')
```
| github_jupyter |
# Example 1e: Spin-bath model (very strong coupling)
### Introduction
The HEOM method solves the dynamics and steady state of a system and its environment, the latter of which is encoded in a set of auxiliary density matrices.
In this example we show the evolution of a single two-level system in contact with a single Bosonic environment. The properties of the system are encoded in Hamiltonian, and a coupling operator which describes how it is coupled to the environment.
The Bosonic environment is implicitly assumed to obey a particular Hamiltonian (see paper), the parameters of which are encoded in the spectral density, and subsequently the free-bath correlation functions.
In the example below we show how to model the overdamped Drude-Lorentz Spectral Density, commonly used with the HEOM. We show how to do the Matsubara and Pade analytical decompositions, as well as how to fit the latter with a finite set of approximate exponentials.
### Drude-Lorentz spectral density
The Drude-Lorentz spectral density is:
$$J(\omega)=\omega \frac{2\lambda\gamma}{{\gamma}^2 + \omega^2}$$
where $\lambda$ scales the coupling strength, and $\gamma$ is the cut-off frequency.
With the HEOM we must use an exponential decomposition:
\begin{equation*}
C(t)=\sum_{k=0}^{k=\infty} c_k e^{-\nu_k t}
\end{equation*}
The Matsubara decomposition of the Drude-Lorentz spectral density is given by:
\begin{equation*}
\nu_k = \begin{cases}
\gamma & k = 0\\
{2 \pi k} / {\beta \hbar} & k \geq 1\\
\end{cases}
\end{equation*}
\begin{equation*}
c_k = \begin{cases}
\lambda \gamma (\cot(\beta \gamma / 2) - i) / \hbar & k = 0\\
4 \lambda \gamma \nu_k / \{(nu_k^2 - \gamma^2)\beta \hbar^2 \} & k \geq 1\\
\end{cases}
\end{equation*}
```
%pylab inline
from qutip import *
from qutip.ipynbtools import HTMLProgressBar
%load_ext autoreload
%autoreload 2
from bofin.heom import BosonicHEOMSolver
# Defining the system Hamiltonian
eps = .0 # Energy of the 2-level system.
Del = .2 # Tunnelling term
Hsys = 0.5 * eps * sigmaz() + 0.5 * Del* sigmax()
# System-bath coupling (Drude-Lorentz spectral density)
Q = sigmaz() # coupling operator
#tlist = np.linspace(0, 600, 600)
tlist = np.linspace(0, pi/Del, 600)
#Bath properties: (see Q. Shi, L. Chen, G. Nan, R.-X. Xu, and Y. Yan, The Journal of chemical physics 130, 084105 (2009))
gamma = 1. # cut off frequency
lam = 2.5 # coupling strength
T = 1. # in units where Boltzmann factor is 1
beta = 1./T
#HEOM parameters
Nk = 1 # number of exponentials in approximation of the the spectral density
NC = 13 # cut off parameter for the bath
###############################################
# Plot the spectral density
def cot(x):
return 1./np.tan(x)
wlist = np.linspace(0, 5, 1000)
pref = 1.
J = [w * 2 * lam * gamma / ((gamma**2 + w**2)) for w in wlist]
# Plot the results
fig, axes = plt.subplots(1, 1, sharex=True, figsize=(8,8))
axes.plot(wlist, J, 'r', linewidth=2)
axes.set_xlabel(r'$\omega$', fontsize=28)
axes.set_ylabel(r'J', fontsize=28)
#Define correlation functions with Matsubara Decomposition with Nk Matsubara terms
def c(t):
c_temp =[]
c_temp.append(pref * lam * gamma * (-1.0j + cot(gamma / (2 * T))) * np.exp(-gamma * t))
for k in range(1,15000):
vk = 2 * np.pi * k * T
c_temp.append((pref * 4 * lam * gamma * T * vk / (vk**2 - gamma**2)) * np.exp(- vk * t) )
return c_temp
ckAR = [pref * lam * gamma * (cot(gamma / (2 * T))) + 0.j]
ckAR.extend([(pref * 4 * lam * gamma * T * 2 * np.pi * k * T / (( 2 * np.pi * k * T)**2 - gamma**2))+0.j for k in range(1,Nk+1)])
vkAR = [gamma+0.j]
vkAR.extend([2 * np.pi * k * T + 0.j for k in range(1,Nk+1)])
ckAI = [pref * lam * gamma * (-1.0) + 0.j]
vkAI = [gamma+0.j]
print(ckAR)
print(ckAI)
print(vkAR)
print(vkAI)
def Jw(w):
Jt = w * 2 * lam * gamma/ ((gamma**2 + w**2))
return Jt
#Collate large number of matsubara terms for a comparison
anamax = 15000
def c(t,mats):
c_temp =[]
c_temp.append(pref * lam * gamma * (-1.0j + cot(gamma / (2 * T))) * np.exp(-gamma * t))
for k in range(1,mats):
vk = 2 * np.pi * k * T
c_temp.append((pref * 4 * lam * gamma * T * vk / (vk**2 - gamma**2)) * np.exp(- vk * t) )
return c_temp
# Reals parts
corrRana = [np.real(sum(c(t,anamax))) for t in tlist]
# Imaginary parts
corrIana = [np.imag(sum(c(t,1))) for t in tlist] #only 1 imaginary term, no need to sum again
cppL = [sum(c(t,Nk)) for t in tlist]
#Construct RHS with specifief Nk Matsubara terms
NR = len(ckAR)
NI = len(ckAI)
Q2 = [Q for kk in range(NR+NI)]
print(Q2)
options = Options(nsteps=15000, store_states=True, rtol=1e-14, atol=1e-14)
HEOMMats = BosonicHEOMSolver(Hsys, Q2, ckAR, ckAI, vkAR, vkAI, NC, options=options)
#Solve ODE for time steps in tlist
# Initial state of the system.
rho0 = basis(2,0) * basis(2,0).dag()
resultMats = HEOMMats.run(rho0, tlist)
# Define some operators with which we will measure the system
# 1,1 element of density matrix - corresonding to groundstate
P11p=basis(2,0) * basis(2,0).dag()
P22p=basis(2,1) * basis(2,1).dag()
# 1,2 element of density matrix - corresonding to coherence
P12p=basis(2,0) * basis(2,1).dag()
# Calculate expectation values in the bases
P11exp = expect(resultMats.states, P11p)
P22exp = expect(resultMats.states, P22p)
P12exp = expect(resultMats.states, P12p)
#do version with tanimura terminator
op = -2*spre(Q)*spost(Q.dag()) + spre(Q.dag()*Q) + spost(Q.dag()*Q)
approx_factr = ((2 * lam / (beta * gamma)) - 1j*lam)
approx_factr -= lam * gamma * (-1.0j + cot(gamma / (2 * T)))/gamma
for k in range(1,Nk+1):
vk = 2 * np.pi * k * T
#c_temp.append((pref * 4 * lam * gamma * T * vk / (vk**2 - gamma**2)) * np.exp(- vk * t) )
approx_factr -= ((pref * 4 * lam * gamma * T * vk / (vk**2 - gamma**2))/ vk)
#approx_factr -= (c[k] / nu[k])
L_bnd = -approx_factr*op
Ltot = -1.0j*(spre(Hsys)-spost(Hsys)) + L_bnd
Ltot = liouvillian(Hsys) + L_bnd
NR = len(ckAR)
NI = len(ckAI)
Q2 = [Q for kk in range(NR+NI)]
print(Q2)
options = Options(nsteps=15000, store_states=True, rtol=1e-14, atol=1e-14)
HEOMMatsT = BosonicHEOMSolver(Ltot, Q2, ckAR, ckAI, vkAR, vkAI, NC, options=options)
# Initial state of the system.
rho0 = basis(2,0) * basis(2,0).dag()
resultMatsT = HEOMMatsT.run(rho0, tlist)
# Define some operators with which we will measure the system
# 1,1 element of density matrix - corresonding to groundstate
P11p=basis(2,0) * basis(2,0).dag()
P22p=basis(2,1) * basis(2,1).dag()
# 1,2 element of density matrix - corresonding to coherence
P12p=basis(2,0) * basis(2,1).dag()
# Calculate expectation values in the bases
P11expT = expect(resultMatsT.states, P11p)
P22expT = expect(resultMatsT.states, P22p)
P12expT = expect(resultMatsT.states, P12p)
# Plot the results
fig, axes = plt.subplots(1, 1, sharex=True, figsize=(8,8))
axes.plot(tlist, np.real(P11exp), 'b', linewidth=2, label="P11")
axes.plot(tlist, np.real(P11expT), 'b--', linewidth=2, label="P11")
axes.set_xlabel(r't', fontsize=28)
axes.legend(loc=0, fontsize=12)
#as a comparison, this is pade
lmax = 1
def deltafun(j,k):
if j==k:
return 1.
else:
return 0.
Alpha =np.zeros((2*lmax,2*lmax))
for j in range(2*lmax):
for k in range(2*lmax):
#Alpha[j][k] = (deltafun(j,k+1)+deltafun(j,k-1))/sqrt((2*(j+1)-1)*(2*(k+1)-1)) #fermi
Alpha[j][k] = (deltafun(j,k+1)+deltafun(j,k-1))/sqrt((2*(j+1)+1)*(2*(k+1)+1)) #bose
eigvalsA=eigvalsh(Alpha)
eps = []
for val in eigvalsA[0:lmax]:
#print(-2/val)
eps.append(-2/val)
AlphaP =np.zeros((2*lmax-1,2*lmax-1))
for j in range(2*lmax-1):
for k in range(2*lmax-1):
#AlphaP[j][k] = (deltafun(j,k+1)+deltafun(j,k-1))/sqrt((2*(j+1)+1)*(2*(k+1)+1)) #fermi
AlphaP[j][k] = (deltafun(j,k+1)+deltafun(j,k-1))/sqrt((2*(j+1)+3)*(2*(k+1)+3)) #Bos: This is +3 because +1 (bose) + 2*(+1)(from bm+1)
eigvalsAP=eigvalsh(AlphaP)
chi = []
for val in eigvalsAP[0:lmax-1]:
#print(-2/val)
chi.append(-2/val)
#eta_list=[0.5*lmax*(2*(lmax + 1) - 1)*(
# np.prod([chi[k]**2 - eps[j]**2 for k in range(lmax - 1)])/
# np.prod([eps[k]**2 - eps[j]**2 +deltafun(j,k) for k in range(lmax)]))
# for j in range(lmax)]
eta_list = []
prefactor = 0.5*lmax*(2*(lmax + 1) + 1)
for j in range(lmax):
term = prefactor
for k1 in range(lmax - 1):
term *= (chi[k1]**2 - eps[j]**2)/(eps[k1]**2 - eps[j]**2 + deltafun(j,k1))
#term2 = 1
for k2 in range(lmax-1,lmax):
term /= (eps[k2]**2 - eps[j]**2 + deltafun(j,k2))
#print(term2)
eta_list.append(term)
#kappa = [0,1,1,1.56,14.44]
kappa = [0]+eta_list
print(kappa)
#epsilon = [0,1*pi,1*3*pi,1.06*5*pi,2.11*7*pi]
#print([0,1*pi,1*3*pi,1.06*5*pi,2.11*7*pi])
epsilon = [0]+eps
beta = 1/T
def f_approx(x):
f = 0.5
for l in range(1,lmax+1):
f= f - 2*kappa[l]*x/(x**2+epsilon[l]**2)
return f
def f(x):
kB=1.
return 1/(1-exp(-x)) #this is n(w)+1 btw! (for bosons)
def C(tlist):
eta_list = []
gamma_list =[]
#l = 0
#eta_0 = 0.5*lam*gamma*f(1.0j*beta*gamma)
eta_0 =lam*gamma*(1.0/np.tan(gamma*beta/2.0) - 1.0j)
gamma_0 = gamma
eta_list.append(eta_0)
gamma_list.append(gamma_0)
if lmax>0:
for l in range(1,lmax+1):
eta_list.append((kappa[l]/beta)*4*lam*gamma*(epsilon[l]/beta)/((epsilon[l]**2/beta**2)-gamma**2))
gamma_list.append(epsilon[l]/beta)
c_tot = []
for t in tlist:
c_tot.append(sum([eta_list[l]*exp(-gamma_list[l]*t) for l in range(lmax+1)]))
return c_tot, eta_list, gamma_list
cppLP,etapLP,gampLP = C(tlist)
fig, ax1 = plt.subplots(figsize=(12, 7))
#print(gam_list)
ax1.plot(tlist,real(cppLP), color="b", linewidth=3, label= r"real pade 2")
#ax1.plot(tlist,imag(cppL), color="r", linewidth=3, label= r"imag alt")
ax1.plot(tlist,corrRana, "r--", linewidth=3, label= r"real mats 15000")
ax1.plot(tlist,real(cppL), "g--", linewidth=3, label= r"real mats 2")
#ax1.plot(tlist,corrIana, "r--", linewidth=3, label= r"imag ana")
ax1.set_xlabel("t")
ax1.set_ylabel(r"$C$")
ax1.legend()
fig, ax1 = plt.subplots(figsize=(12, 7))
ax1.plot(tlist,real(cppLP)-corrRana, color="b", linewidth=3, label= r"pade error")
ax1.plot(tlist,real(cppL)-corrRana, "r--", linewidth=3, label= r"mats error")
ax1.set_xlabel("t")
ax1.set_ylabel(r"$C$")
ax1.legend()
ckAR = [real(eta) +0j for eta in etapLP]
ckAI = [imag(etapLP[0]) + 0j]
vkAR = [gam +0j for gam in gampLP]
vkAI = [gampLP[0] + 0j]
vkAR
NR = len(ckAR)
NI = len(ckAI)
Q2 = [Q for kk in range(NR+NI)]
print(Q2)
options = Options(nsteps=15000, store_states=True, rtol=1e-14, atol=1e-14)
HEOMPade = BosonicHEOMSolver(Hsys, Q2, ckAR, ckAI, vkAR, vkAI, NC, options=options)
# Initial state of the system.
rho0 = basis(2,0) * basis(2,0).dag()
# Times to record state
#tlist = np.linspace(0, 40, 600)
resultPade = HEOMPade.run(rho0, tlist)
# Define some operators with which we will measure the system
# 1,1 element of density matrix - corresonding to groundstate
P11p=basis(2,0) * basis(2,0).dag()
P22p=basis(2,1) * basis(2,1).dag()
# 1,2 element of density matrix - corresonding to coherence
P12p=basis(2,0) * basis(2,1).dag()
# Calculate expectation values in the bases
P11expP = expect(resultPade.states, P11p)
P22expP = expect(resultPade.states, P22p)
P12expP = expect(resultPade.states, P12p)
# Plot the results
fig, axes = plt.subplots(1, 1, sharex=True, figsize=(8,8))
#axes.plot(tlist, np.real(P11exp)+ np.real(P22exp), 'b', linewidth=2, label="P11")
axes.plot(tlist, np.real(P11exp), 'b', linewidth=2, label="P11 mats")
axes.plot(tlist, np.real(P11expT), 'r', linewidth=2, label="P11 mats++term")
axes.plot(tlist, np.real(P11expP), 'b--', linewidth=2, label="P11 pade")
#axes.plot(tlist, np.real(P12exp), 'r', linewidth=2, label="P12 mats")
#axes.plot(tlist, np.real(P12expP), 'r--', linewidth=2, label="P12 pade")
axes.set_xlabel(r't', fontsize=28)
axes.legend(loc=0, fontsize=12)
```
### Do fitting of Matsubara expansion
```
tlist2= linspace(0,6,10000)
lmaxmats = 15000
def c(t,anamax):
c_temp = (pref * lam * gamma * (-1.0j + cot(gamma / (2 * T))) * np.exp(-gamma * t))
for k in range(1, anamax):
vk = 2 * np.pi * k * T
c_temp += ((pref * 4 * lam * gamma * T * vk / (vk**2 - gamma**2)) * np.exp(- vk * t) )
return c_temp
# Reals parts
corrRana = [np.real(c(t,lmaxmats)) for t in tlist2]
# Imaginary parts
corrIana = [np.imag((pref * lam * gamma * (-1.0j + cot(gamma / (2 * T))) * np.exp(-gamma * t))) for t in tlist2]
from scipy.optimize import curve_fit
def wrapper_fit_func(x, N, *args):
a, b = list(args[0][:N]), list(args[0][N:2*N])
# print("debug")
return fit_func(x, a, b, N)
# actual fitting function
def fit_func(x, a, b, N):
tot = 0
for i in range(N):
# print(i)
tot += a[i]*np.exp(b[i]*x)
return tot
def fitter(ans, tlist, k):
# the actual computing of fit
popt = []
pcov = []
# tries to fit for k exponents
for i in range(k):
params_0 = [0]*(2*(i+1))
upper_a = abs(max(ans, key = abs))*10
#sets initial guess
guess = []
aguess = [ans[0]]*(i+1)#[max(ans)]*(i+1)
bguess = [0]*(i+1)
guess.extend(aguess)
guess.extend(bguess)
# sets bounds
# a's = anything , b's negative
# sets lower bound
b_lower = []
alower = [-upper_a]*(i+1)
blower = [-np.inf]*(i+1)
b_lower.extend(alower)
b_lower.extend(blower)
# sets higher bound
b_higher = []
ahigher = [upper_a]*(i+1)
bhigher = [0]*(i+1)
b_higher.extend(ahigher)
b_higher.extend(bhigher)
param_bounds = (b_lower, b_higher)
p1, p2 = curve_fit(lambda x, *params_0: wrapper_fit_func(x, i+1, \
params_0), tlist, ans, p0=guess, sigma=[0.01 for t in tlist2], bounds = param_bounds, maxfev = 1e8)
popt.append(p1)
pcov.append(p2)
print(i+1)
return popt
# print(popt)
# function that evaluates values with fitted params at
# given inputs
def checker(tlist, vals):
y = []
for i in tlist:
# print(i)
y.append(wrapper_fit_func(i, int(len(vals)/2), vals))
return y
k = 3
popt1 = fitter(corrRana, tlist2, k)
for i in range(k):
y = checker(tlist2, popt1[i])
plt.plot(tlist2, corrRana, tlist2, y)
plt.show()
k1 = 1
popt2 = fitter(corrIana, tlist2, k1)
for i in range(k1):
y = checker(tlist2, popt2[i])
plt.plot(tlist2, corrIana, tlist2, y)
plt.show()
ckAR1 = list(popt1[k-1])[:len(list(popt1[k-1]))//2]
ckAR = [x+0j for x in ckAR1]
ckAI1 = list(popt2[k1-1])[:len(list(popt2[k1-1]))//2]
#minus?wtfiforgotagain
ckAI = [x+0j for x in ckAI1]
# vkAR, vkAI
vkAR1 = list(popt1[k-1])[len(list(popt1[k-1]))//2:]
vkAR = [-x+0j for x in vkAR1]
vkAI1 = list(popt2[k1-1])[len(list(popt2[k1-1]))//2:]
vkAI = [-x+0j for x in vkAI1]
# NC, NR, NI
# print(np.array(H).shape, np.array(Q).shape)
NC = 13
NR = len(ckAR)
NI = len(ckAI)
options = Options(nsteps=15000, store_states=True, rtol=1e-14, atol=1e-14)
#overwrite imaginary fit with analytical value jsut in case
ckAI = [pref * lam * gamma * (-1.0) + 0.j]
vkAI = [gamma+0.j]
print(ckAI)
print(vkAI)
NR = len(ckAR)
NI = len(ckAI)
Q2 = [Q for kk in range(NR+NI)]
print(Q2)
# Initial state of the system.
rho0 = basis(2,0) * basis(2,0).dag()
# Times to record state
#tlist = np.linspace(0, 40, 600)
options = Options(nsteps=1500, store_states=True, rtol=1e-12, atol=1e-12)
HEOMFit = BosonicHEOMSolver(Hsys, Q2, ckAR, ckAI, vkAR, vkAI, NC, ode_use_mkl = True,options=options)
start = time.time()
resultFitMKL = HEOMFit.run(rho0, tlist)
end = time.time()
print(end - start)
# Initial state of the system.
rho0 = basis(2,0) * basis(2,0).dag()
# Times to record state
#tlist = np.linspace(0, 40, 600)
options = Options(nsteps=1500, store_states=True, rtol=1e-12, atol=1e-12)
HEOMFit = BosonicHEOMSolver(Hsys, Q2, ckAR, ckAI, vkAR, vkAI, NC, ode_use_mkl = False,options=options)
start = time.time()
resultFit = HEOMFit.run(rho0, tlist)
end = time.time()
print(end - start)
#do version with tanimura terminator
op = -2*spre(Q)*spost(Q.dag()) + spre(Q.dag()*Q) + spost(Q.dag()*Q)
approx_factr = ((2 * lam / (beta * gamma)) - 1j*lam)
approx_factr -= lam * gamma * (-1.0j + cot(gamma / (2 * T)))/gamma
for k in range(1,lmaxmats):
vk = 2 * np.pi * k * T
#c_temp.append((pref * 4 * lam * gamma * T * vk / (vk**2 - gamma**2)) * np.exp(- vk * t) )
approx_factr -= ((pref * 4 * lam * gamma * T * vk / (vk**2 - gamma**2))/ vk)
#approx_factr -= (c[k] / nu[k])
L_bnd = -approx_factr*op
print(approx_factr)
Ltot = -1.0j*(spre(Hsys)-spost(Hsys)) + L_bnd
Ltot = liouvillian(Hsys) + L_bnd
NR = len(ckAR)
NI = len(ckAI)
Q2 = [Q for kk in range(NR+NI)]
print(Q2)
options = Options(nsteps=1500, store_states=True, rtol=1e-12, atol=1e-12, method="bdf")
HEOMFitT = BosonicHEOMSolver(Ltot, Q2, ckAR, ckAI, vkAR, vkAI, NC, options=options)
# Initial state of the system.
rho0 = basis(2,0) * basis(2,0).dag()
# Times to record state
#tlist = np.linspace(0, 40, 600)
resultFitT = HEOMFitT.run(rho0, tlist)
# Define some operators with which we will measure the system
# 1,1 element of density matrix - corresonding to groundstate
P11p=basis(2,0) * basis(2,0).dag()
P22p=basis(2,1) * basis(2,1).dag()
# 1,2 element of density matrix - corresonding to coherence
P12p=basis(2,0) * basis(2,1).dag()
# Calculate expectation values in the bases
P11expFT = expect(resultFitT.states, P11p)
P22expFT = expect(resultFitT.states, P22p)
P12expFT = expect(resultFitT.states, P12p)
qsave(resultMats, 'data/resultMatsOD')
qsave(resultMatsT, 'data/resultMatsTOD')
qsave(resultPade, 'data/resultPadeOD')
qsave(resultFit, 'data/resultFitOD')
##qsave(resultFitT, 'resultFitTOD')
resultFit = qload('data/resultFitOD')
print(Hsys.eigenstates())
energies, states = Hsys.eigenstates()
rhoss = (states[0]*states[0].dag()*exp(-beta*energies[0]) + states[1]*states[1].dag()*exp(-beta*energies[1]))
rhoss = rhoss/rhoss.norm()
P11 = expect(rhoss,P11p)
P12 = expect(rhoss,P12p)
DL = " 2*pi* 2.0 * {lam} / (pi * {gamma} * {beta}) if (w==0) else 2*pi*(2.0*{lam}*{gamma} *w /(pi*(w**2+{gamma}**2))) * ((1/(exp((w) * {beta})-1))+1)".format(gamma=gamma, beta = beta, lam = lam)
optionsODE = Options(nsteps=15000, store_states=True,rtol=1e-12,atol=1e-12)
outputBR = brmesolve(Hsys, rho0, tlist, a_ops=[[sigmaz(),DL]], options = optionsODE)
# Calculate expectation values in the bases
P11BR = expect(outputBR.states, P11p)
P22BR = expect(outputBR.states, P22p)
P12BR = expect(outputBR.states, P12p)
matplotlib.rcParams['figure.figsize'] = (7, 5)
matplotlib.rcParams['axes.titlesize'] = 25
matplotlib.rcParams['axes.labelsize'] = 30
matplotlib.rcParams['xtick.labelsize'] = 28
matplotlib.rcParams['ytick.labelsize'] = 28
matplotlib.rcParams['legend.fontsize'] = 28
matplotlib.rcParams['axes.grid'] = False
matplotlib.rcParams['savefig.bbox'] = 'tight'
matplotlib.rcParams['lines.markersize'] = 5
matplotlib.rcParams['font.family'] = 'STIXgeneral'
matplotlib.rcParams['mathtext.fontset'] = 'stix'
matplotlib.rcParams["font.serif"] = "STIX"
matplotlib.rcParams['text.usetex']=False
# Define some operators with which we will measure the system
# 1,1 element of density matrix - corresonding to groundstate
P11p=basis(2,0) * basis(2,0).dag()
P22p=basis(2,1) * basis(2,1).dag()
# 1,2 element of density matrix - corresonding to coherence
P12p=basis(2,0) * basis(2,1).dag()
# Calculate expectation values in the bases
P11expF = expect(resultFit.states, P11p)
P22expF = expect(resultFit.states, P22p)
P12expF = expect(resultFit.states, P12p)
# Plot the results
fig, axes = plt.subplots(1, 1, sharex=True, figsize=(12,7))
plt.yticks([0.99,1.0],[0.99,1])
axes.plot(tlist, np.real(P11exp), 'b', linewidth=2, label=r"Matsubara $N_k=1$")
axes.plot(tlist, np.real(P11expT), 'g--', linewidth=3, label=r"Matsubara $N_k=1$ & terminator")
axes.plot(tlist, np.real(P11expP), 'y-.', linewidth=2, label=r"Padé $N_k=1$")
#axes.plot(tlist, np.real(P11BR), 'y-.', linewidth=2, label="Bloch Redfield")
axes.plot(tlist, np.real(P11expF), 'r',dashes=[3,2], linewidth=2, label=r"Fit $N_f = 3$, $N_k=15 \times 10^3$")
axes.locator_params(axis='y', nbins=6)
axes.locator_params(axis='x', nbins=6)
axes.set_ylabel(r'$\rho_{11}$',fontsize=30)
axes.set_xlabel(r'$t\;\gamma$',fontsize=30)
axes.set_xlim(tlist[0],tlist[-1])
axes.set_ylim(0.98405,1.0005)
axes.legend(loc=0)
fig.savefig("figures/fig2.pdf")
from qutip.ipynbtools import version_table
version_table()
```
| github_jupyter |
```
# Install TensorFlow
# !pip install -q tensorflow-gpu==2.0.0-rc0
try:
%tensorflow_version 2.x # Colab only.
except Exception:
pass
import tensorflow as tf
print(tf.__version__)
# More imports
from tensorflow.keras.layers import Input, Dense, Flatten
from tensorflow.keras.applications.vgg16 import VGG16 as PretrainedModel, \
preprocess_input
from tensorflow.keras.models import Model
from tensorflow.keras.optimizers import SGD, Adam
from tensorflow.keras.preprocessing import image
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from glob import glob
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import sys, os
# Data from: https://mmspg.epfl.ch/downloads/food-image-datasets/
# !wget --passive-ftp --prefer-family=ipv4 --ftp-user FoodImage@grebvm2.epfl.ch \
# --ftp-password Cahc1moo -nc ftp://tremplin.epfl.ch/Food-5K.zip
!wget -nc https://lazyprogrammer.me/course_files/Food-5K.zip
!unzip -qq -o Food-5K.zip
!ls
!mv Food-5K/* .
!ls training
# look at an image for fun
plt.imshow(image.load_img('training/0_808.jpg'))
plt.show()
# Food images start with 1, non-food images start with 0
plt.imshow(image.load_img('training/1_616.jpg'))
plt.show()
!mkdir data
# Make directories to store the data Keras-style
!mkdir data/train
!mkdir data/test
!mkdir data/train/nonfood
!mkdir data/train/food
!mkdir data/test/nonfood
!mkdir data/test/food
# Move the images
# Note: we will consider 'training' to be the train set
# 'validation' folder will be the test set
# ignore the 'evaluation' set
!mv training/0*.jpg data/train/nonfood
!mv training/1*.jpg data/train/food
!mv validation/0*.jpg data/test/nonfood
!mv validation/1*.jpg data/test/food
train_path = 'data/train'
valid_path = 'data/test'
# These images are pretty big and of different sizes
# Let's load them all in as the same (smaller) size
IMAGE_SIZE = [200, 200]
# useful for getting number of files
image_files = glob(train_path + '/*/*.jpg')
valid_image_files = glob(valid_path + '/*/*.jpg')
# useful for getting number of classes
folders = glob(train_path + '/*')
folders
# look at an image for fun
plt.imshow(image.load_img(np.random.choice(image_files)))
plt.show()
ptm = PretrainedModel(
input_shape=IMAGE_SIZE + [3],
weights='imagenet',
include_top=False)
# map the data into feature vectors
x = Flatten()(ptm.output)
# create a model object
model = Model(inputs=ptm.input, outputs=x)
# view the structure of the model
model.summary()
# create an instance of ImageDataGenerator
gen = ImageDataGenerator(preprocessing_function=preprocess_input)
batch_size = 128
# create generators
train_generator = gen.flow_from_directory(
train_path,
target_size=IMAGE_SIZE,
batch_size=batch_size,
class_mode='binary',
)
valid_generator = gen.flow_from_directory(
valid_path,
target_size=IMAGE_SIZE,
batch_size=batch_size,
class_mode='binary',
)
Ntrain = len(image_files)
Nvalid = len(valid_image_files)
# Figure out the output size
feat = model.predict(np.random.random([1] + IMAGE_SIZE + [3]))
D = feat.shape[1]
X_train = np.zeros((Ntrain, D))
Y_train = np.zeros(Ntrain)
X_valid = np.zeros((Nvalid, D))
Y_valid = np.zeros(Nvalid)
# populate X_train and Y_train
i = 0
for x, y in train_generator:
# get features
features = model.predict(x)
# size of the batch (may not always be batch_size)
sz = len(y)
# assign to X_train and Ytrain
X_train[i:i + sz] = features
Y_train[i:i + sz] = y
# increment i
i += sz
print(i)
if i >= Ntrain:
print('breaking now')
break
print(i)
# populate X_valid and Y_valid
i = 0
for x, y in valid_generator:
# get features
features = model.predict(x)
# size of the batch (may not always be batch_size)
sz = len(y)
# assign to X_train and Ytrain
X_valid[i:i + sz] = features
Y_valid[i:i + sz] = y
# increment i
i += sz
if i >= Nvalid:
print('breaking now')
break
print(i)
X_train.max(), X_train.min()
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
X_train2 = scaler.fit_transform(X_train)
X_valid2 = scaler.transform(X_valid)
# Try the built-in logistic regression
from sklearn.linear_model import LogisticRegression
logr = LogisticRegression()
logr.fit(X_train2, Y_train)
print(logr.score(X_train2, Y_train))
print(logr.score(X_valid2, Y_valid))
# Do logistic regression in Tensorflow
i = Input(shape=(D,))
x = Dense(1, activation='sigmoid')(i)
linearmodel = Model(i, x)
linearmodel.compile(
loss='binary_crossentropy',
optimizer='adam',
metrics=['accuracy'])
# Can try both normalized and unnormalized data
r = linearmodel.fit(
X_train, Y_train,
batch_size=128,
epochs=10,
validation_data=(X_valid, Y_valid),
)
# loss
plt.plot(r.history['loss'], label='train loss')
plt.plot(r.history['val_loss'], label='val loss')
plt.legend()
plt.show()
# accuracies
plt.plot(r.history['accuracy'], label='train acc')
plt.plot(r.history['val_accuracy'], label='val acc')
plt.legend()
plt.show()
```
| github_jupyter |
```
# TO DO
# - Nothing for the moment
# PENALTY FUNCTIONS - SOME EXAMPLES
# multiplier is a positive number > 0 that determines the slope
# Linear Penalty Function
def linearPenalty(x, multiplier=1):
return x * multiplier
# Flipped/Inverse Linear Penalty Function
def invLinearPenalty(x, multiplier=1):
return -x * multiplier
# Linear for negative x and zero for positive x
def leftLinearPenalty(x, multiplier=1):
if(x < 0): return -x * multiplier
else: return 0
# Linear for positive x and zero for negative x
def rightLinearPenalty(x, multiplier=1):
if(x < 0): return 0
else: return x * multiplier
# V shape penalty
def VPenalty(x, multiplier=1):
if (x < 0): return -x * multiplier
else: return x
# Inverted V shape penalty
def invertedVPenalty(x, multiplier=1):
if (x < 0): return x * multiplier
else: return -x * multiplier
# Positive parabola penalty
def squaredPenalty(x, multiplier=1):
return (x**2) * multiplier
# Inverted parabola penalty
def invertedSquaredPenalty(x, multiplier=1):
return -(x**2) * multiplier
# Non-linear penalty
def nonLinearPenalty(x, multiplier=1):
return x + x**2 + x**3
penaltyFunctions = {linearPenalty: "Linear Penalty",
invLinearPenalty: "Inverse Linear Penalty",
leftLinearPenalty: "Left-Linear Penalty",
rightLinearPenalty: "Right-Linear Penalty",
VPenalty: "V Penalty",
invertedVPenalty: "Inverted-V Penalty",
squaredPenalty: "Squared Penalty",
invertedSquaredPenalty: "Inverted Squared Penalty",
nonLinearPenalty: "Non-Linear Penalty"
}
# Given a list of error values, plot the penalty function
# Error = Actual - Predicted value - This is always along a single dimension because the output is always a single
# column in a dataset.
import matplotlib.pyplot as plt
import matplotlib.cm as cm
%matplotlib inline
import seaborn as sns
# Plot the penalty function for a given list of error values and a given penalty function
def penaltyPlot(errorList, penaltyFunction):
# Set up the x-axis
num_points = 200
x = np.linspace(min(errorList), max(errorList), num_points)
fig, ax = plt.subplots(figsize=(6,4))
ax.set(xlabel='Predicted Value - Actual Value')
ax.set(ylabel='Penalty')
ax.axvline(x=0, color='black')
ax.axhline(y=0, color='black')
ax.set(title=penaltyFunctions[penaltyFunction])
ax.plot(x, list(map(penaltyFunction,x)))
# Load up the packages to investigate the data
import numpy as np
import pandas as pd
# Add a column of ones to the first column of a dataframe
# and turn it into a matrix
def df_addOnes(dataFrame):
vals = dataFrame.values
#add_ones_column = zip(np.ones(len(dataFrame)), vals)
#feature_matrix = np.matrix([val for val in add_ones_column])
feature_matrix = np.c_[np.ones(len(dataFrame)), vals]
return feature_matrix
# Making it easy to calculate the total penalty over the entire dataset
def penalty(df_features, df_output, paramater_value_list, penalty_function):
# df_features is a dataframe of the features (no column of ones added)
# df_output is a dataframe of the output column (target variable)
# parameter_value_list is a list of w0, w1, ..., wn+1 where n is the number of features
# i.e., the number of columns in df_features.
# Cost of being wrong calculated over the entire data set
# Will take X and add a first column of 1s to it to enable the matrix multiplication
# Therefore: X is an m x n matrix and theta is a n x 1 matrix
#### Turn the function inputs into matrices ####
# Get X and y into the right shapes for use in the penalty function
# Add a first column of ones to the feature matrix
# Add a column of 1s to X
feature_matrix = df_addOnes(df_features)
output_matrix = np.matrix(df_output.values)
parameter_matrix = np.matrix(paramater_value_list).T
#print(feature_matrix.shape, parameter_matrix.shape, output_matrix.shape)
# Difference between the predicted and the actual value
error = (feature_matrix * parameter_matrix) - output_matrix
#print(error.shape)
# penaltyPerOutput is an m x 1 matrix where each element is the penalty for
# the input and its associated output for a particular value of W
# Apply a penalty function to the errors from each row of the dataset
penaltyPerOutput = list(map(penalty_function,error))
# totalPenalty is the sum of the penalties of each row of the dataset
totalPenalty = np.sum(penaltyPerOutput)
# The penalty of getting it wrong is 1/2m of the totalPenalty (normalized penalty)
# m is the number of rows in df_features
totalPenaltyNorm = totalPenalty / (2 * len(df_features))
return totalPenaltyNorm
# Implement Gradient Descent
# **NOTE: ONLY for a squared penalty function**
def gradientDescent(df_features,
df_output,
init_params_list,
num_iterations=100,
learning_rate=0.0001,
penalty_function=squaredPenalty):
# df_features is a dataframe with the features
# df_ouptut is a dataframe of the output column
# init_params_list is the list of initial W values, e.g., [-1.0, 3.53]
# num_iterations is the number of steps taken by the algorithm as it descends the penalty surface
# learning_rate is the multiplier that determines step size (smaller = smaller step size)
# penalty_function is the penalty function applied to the machine learning problem
# NOTE: The formula for gradient descent we're implementing works only for the squaredPenalty function
# Get the inputs into matrix form so we can use matrix multiplication (more efficient)
feature_matrix = df_addOnes(df_features)
m = len(feature_matrix) # number of rows of data
output_matrix = np.matrix(df_output.values)
parameter_matrix = np.matrix(init_params_list).T
# This is the initial value of the parameters in matrix form
w = parameter_matrix
# Set up arrays to capture the running results
# Specify dtype=object because we're putting arrays into an array
#running_w = np.empty(num_iterations, dtype = object)
#running_w = np.array([[ 0.50182941],[-0.07935517]])
running_w = np.array(parameter_matrix)
# don't have to specify dtype for the other arrays because we're putting single values into the array
running_error = np.zeros(num_iterations)
running_normError = np.zeros(num_iterations)
running_penalty = np.zeros(num_iterations)
# Iterate over the dataset num_iterations times and adjust the values of the parameters each time
for i in range(num_iterations):
#print(w)
for j in range(len(parameter_matrix)):
error = ((feature_matrix * w) - output_matrix).T * np.matrix(feature_matrix[:,j]).T
normError = (learning_rate/m) * error
w[j] = w[j] - normError
#print(w[j])
# w, error, normError and penalty after each iteration
#running_w[i] = w
running_w = np.append(running_w, w, axis=0)
#print(i)
#print(w)
#print(running_w)
running_error[i] = np.sum((feature_matrix * w) - output_matrix.T)
running_normError[i] = (learning_rate/m) * running_error[i]
running_penalty[i] = penalty_function(running_error[i])
# w is the value of parameters afer num_iterations
#print(w)
# Get the running_w into the right form
# From https://jasonstitt.com/python-group-iterator-list-function
running_w = list(zip(*[iter(running_w)] * len(parameter_matrix)))
# error after num_iterations
final_error = np.sum((feature_matrix * w) - output_matrix.T)
# Penalty after num_iterations
final_penalty = penalty_function(final_error)
return w, final_penalty, running_w, running_penalty
```
| github_jupyter |
## Fixed-Type Arrays in Python
Python offers several different options for storing data in efficient, fixed-type data buffers.
The built-in ``array`` module (available since Python 3.3) can be used to create dense arrays of a uniform type:
```
import array
L = list(range(10))
A = array.array('i', L)
A
```
Here ``'i'`` is a type code indicating the contents are integers.
Much more useful, however, is the ``ndarray`` object of the NumPy package.
While Python's ``array`` object provides efficient storage of array-based data, NumPy adds to this efficient *operations* on that data.
We will explore these operations in later sections; here we'll demonstrate several ways of creating a NumPy array.
We'll start with the standard NumPy import, under the alias ``np``:
```
import numpy as np
```
## Creating Arrays from Python Lists
First, we can use ``np.array`` to create arrays from Python lists:
```
# integer array:
np.array([1, 4, 2, 5, 3])
```
Remember that unlike Python lists, NumPy is constrained to arrays that all contain the same type.
If types do not match, NumPy will upcast if possible (here, integers are up-cast to floating point):
```
np.array([3.14, 4, 2, 3])
```
If we want to explicitly set the data type of the resulting array, we can use the ``dtype`` keyword:
```
np.array([1, 2, 3, 4], dtype='float32')
```
Finally, unlike Python lists, NumPy arrays can explicitly be multi-dimensional; here's one way of initializing a multidimensional array using a list of lists:
```
# nested lists result in multi-dimensional arrays
np.array([range(i, i + 3) for i in [2, 4, 6]])
```
The inner lists are treated as rows of the resulting two-dimensional array.
## Creating Arrays from Scratch
Especially for larger arrays, it is more efficient to create arrays from scratch using routines built into NumPy.
Here are several examples:
```
# Create a length-10 integer array filled with zeros
np.zeros(10, dtype=int)
# Create a 3x5 floating-point array filled with ones
np.ones((3, 5), dtype=float)
# Create a 3x5 array filled with 3.14
np.full((3, 5), 3.14)
# Create an array filled with a linear sequence
# Starting at 0, ending at 20, stepping by 2
# (this is similar to the built-in range() function)
np.arange(0, 20, 2)
# Create an array of five values evenly spaced between 0 and 1
np.linspace(0, 1, 5)
# Create a 3x3 array of uniformly distributed
# random values between 0 and 1
np.random.random((3, 3))
# Create a 3x3 array of normally distributed random values
# with mean 0 and standard deviation 1
np.random.normal(0, 1, (3, 3))
# Create a 3x3 array of random integers in the interval [0, 10)
np.random.randint(0, 10, (3, 3))
# Create a 3x3 identity matrix
np.eye(3)
# Create an uninitialized array of three integers
# The values will be whatever happens to already exist at that memory location
np.empty(3)
```
## NumPy Standard Data Types
NumPy arrays contain values of a single type, so it is important to have detailed knowledge of those types and their limitations.
Because NumPy is built in C, the types will be familiar to users of C, Fortran, and other related languages.
The standard NumPy data types are listed in the following table.
Note that when constructing an array, they can be specified using a string:
```python
np.zeros(10, dtype='int16')
```
Or using the associated NumPy object:
```python
np.zeros(10, dtype=np.int16)
```
| Data type | Description |
|---------------|-------------|
| ``bool_`` | Boolean (True or False) stored as a byte |
| ``int_`` | Default integer type (same as C ``long``; normally either ``int64`` or ``int32``)|
| ``intc`` | Identical to C ``int`` (normally ``int32`` or ``int64``)|
| ``intp`` | Integer used for indexing (same as C ``ssize_t``; normally either ``int32`` or ``int64``)|
| ``int8`` | Byte (-128 to 127)|
| ``int16`` | Integer (-32768 to 32767)|
| ``int32`` | Integer (-2147483648 to 2147483647)|
| ``int64`` | Integer (-9223372036854775808 to 9223372036854775807)|
| ``uint8`` | Unsigned integer (0 to 255)|
| ``uint16`` | Unsigned integer (0 to 65535)|
| ``uint32`` | Unsigned integer (0 to 4294967295)|
| ``uint64`` | Unsigned integer (0 to 18446744073709551615)|
| ``float_`` | Shorthand for ``float64``.|
| ``float16`` | Half precision float: sign bit, 5 bits exponent, 10 bits mantissa|
| ``float32`` | Single precision float: sign bit, 8 bits exponent, 23 bits mantissa|
| ``float64`` | Double precision float: sign bit, 11 bits exponent, 52 bits mantissa|
| ``complex_`` | Shorthand for ``complex128``.|
| ``complex64`` | Complex number, represented by two 32-bit floats|
| ``complex128``| Complex number, represented by two 64-bit floats|
More advanced type specification is possible, such as specifying big or little endian numbers; for more information, refer to the [NumPy documentation](http://numpy.org/).
NumPy also supports compound data types, which will be covered in [Structured Data: NumPy's Structured Arrays](02.09-Structured-Data-NumPy.ipynb).
# The Basics of NumPy Arrays
Data manipulation in Python is nearly synonymous with NumPy array manipulation: even newer tools like Pandas are built around the NumPy array.
This section will present several examples of using NumPy array manipulation to access data and subarrays, and to split, reshape, and join the arrays.
While the types of operations shown here may seem a bit dry and pedantic, they comprise the building blocks of many other examples.
Get to know them well!
We'll cover a few categories of basic array manipulations here:
- *Attributes of arrays*: Determining the size, shape, memory consumption, and data types of arrays
- *Indexing of arrays*: Getting and setting the value of individual array elements
- *Slicing of arrays*: Getting and setting smaller subarrays within a larger array
- *Reshaping of arrays*: Changing the shape of a given array
- *Joining and splitting of arrays*: Combining multiple arrays into one, and splitting one array into many
## NumPy Array Attributes
First let's discuss some useful array attributes.
We'll start by defining three random arrays, a one-dimensional, two-dimensional, and three-dimensional array.
We'll use NumPy's random number generator, which we will *seed* with a set value in order to ensure that the same random arrays are generated each time this code is run:
```
import numpy as np
np.random.seed(0) # seed for reproducibility
x1 = np.random.randint(10, size=6) # One-dimensional array
x2 = np.random.randint(10, size=(3, 4)) # Two-dimensional array
x3 = np.random.randint(10, size=(3, 4, 5)) # Three-dimensional array
```
Each array has attributes ``ndim`` (the number of dimensions), ``shape`` (the size of each dimension), and ``size`` (the total size of the array):
```
print("x3 ndim: ", x3.ndim)
print("x3 shape:", x3.shape)
print("x3 size: ", x3.size)
```
Another useful attribute is the ``dtype``, the data type of the array:
```
print("dtype:", x3.dtype)
```
Other attributes include ``itemsize``, which lists the size (in bytes) of each array element, and ``nbytes``, which lists the total size (in bytes) of the array:
```
print("itemsize:", x3.itemsize, "bytes")
print("nbytes:", x3.nbytes, "bytes")
```
In general, we expect that ``nbytes`` is equal to ``itemsize`` times ``size``.
## Array Indexing: Accessing Single Elements
If you are familiar with Python's standard list indexing, indexing in NumPy will feel quite familiar.
In a one-dimensional array, the $i^{th}$ value (counting from zero) can be accessed by specifying the desired index in square brackets, just as with Python lists:
```
x1
x1[0]
x1[4]
```
To index from the end of the array, you can use negative indices:
```
x1[-1]
x1[-2]
```
In a multi-dimensional array, items can be accessed using a comma-separated tuple of indices:
```
x2
x2[0, 0]
x2[2, 0]
x2[2, -1]
```
Values can also be modified using any of the above index notation:
```
x2[0, 0] = 12
x2
```
Keep in mind that, unlike Python lists, NumPy arrays have a fixed type.
This means, for example, that if you attempt to insert a floating-point value to an integer array, the value will be silently truncated. Don't be caught unaware by this behavior!
```
x1[0] = 3.14159 # this will be truncated!
x1
```
## Array Slicing: Accessing Subarrays
Just as we can use square brackets to access individual array elements, we can also use them to access subarrays with the *slice* notation, marked by the colon (``:``) character.
The NumPy slicing syntax follows that of the standard Python list; to access a slice of an array ``x``, use this:
``` python
x[start:stop:step]
```
If any of these are unspecified, they default to the values ``start=0``, ``stop=``*``size of dimension``*, ``step=1``.
We'll take a look at accessing sub-arrays in one dimension and in multiple dimensions.
### One-dimensional subarrays
```
x = np.arange(10)
x
x[:5] # first five elements
x[5:] # elements after index 5
x[4:7] # middle sub-array
x[::2] # every other element
x[1::2] # every other element, starting at index 1
```
A potentially confusing case is when the ``step`` value is negative.
In this case, the defaults for ``start`` and ``stop`` are swapped.
This becomes a convenient way to reverse an array:
```
x[::-1] # all elements, reversed
x[5::-2] # reversed every other from index 5
```
### Multi-dimensional subarrays
Multi-dimensional slices work in the same way, with multiple slices separated by commas.
For example:
```
x2
x2[:2, :3] # two rows, three columns
x2[:3, ::2] # all rows, every other column
```
Finally, subarray dimensions can even be reversed together:
```
x2[::-1, ::-1]
```
#### Accessing array rows and columns
One commonly needed routine is accessing of single rows or columns of an array.
This can be done by combining indexing and slicing, using an empty slice marked by a single colon (``:``):
```
print(x2[:, 0]) # first column of x2
print(x2[0, :]) # first row of x2
```
In the case of row access, the empty slice can be omitted for a more compact syntax:
```
print(x2[0]) # equivalent to x2[0, :]
```
### Subarrays as no-copy views
One important–and extremely useful–thing to know about array slices is that they return *views* rather than *copies* of the array data.
This is one area in which NumPy array slicing differs from Python list slicing: in lists, slices will be copies.
Consider our two-dimensional array from before:
```
print(x2)
```
Let's extract a $2 \times 2$ subarray from this:
```
x2_sub = x2[:2, :2]
print(x2_sub)
```
Now if we modify this subarray, we'll see that the original array is changed! Observe:
```
x2_sub[0, 0] = 99
print(x2_sub)
print(x2)
```
This default behavior is actually quite useful: it means that when we work with large datasets, we can access and process pieces of these datasets without the need to copy the underlying data buffer.
### Creating copies of arrays
Despite the nice features of array views, it is sometimes useful to instead explicitly copy the data within an array or a subarray. This can be most easily done with the ``copy()`` method:
```
x2_sub_copy = x2[:2, :2].copy()
print(x2_sub_copy)
```
If we now modify this subarray, the original array is not touched:
```
x2_sub_copy[0, 0] = 42
print(x2_sub_copy)
print(x2)
```
## Reshaping of Arrays
Another useful type of operation is reshaping of arrays.
The most flexible way of doing this is with the ``reshape`` method.
For example, if you want to put the numbers 1 through 9 in a $3 \times 3$ grid, you can do the following:
```
grid = np.arange(1, 10).reshape((3, 3))
print(grid)
```
Note that for this to work, the size of the initial array must match the size of the reshaped array.
Where possible, the ``reshape`` method will use a no-copy view of the initial array, but with non-contiguous memory buffers this is not always the case.
Another common reshaping pattern is the conversion of a one-dimensional array into a two-dimensional row or column matrix.
This can be done with the ``reshape`` method, or more easily done by making use of the ``newaxis`` keyword within a slice operation:
```
x = np.array([1, 2, 3])
# row vector via reshape
x.reshape((1, 3))
# row vector via newaxis
x[np.newaxis, :]
# column vector via reshape
x.reshape((3, 1))
# column vector via newaxis
x[:, np.newaxis]
```
We will see this type of transformation often.
## Array Concatenation and Splitting
All of the preceding routines worked on single arrays. It's also possible to combine multiple arrays into one, and to conversely split a single array into multiple arrays. We'll take a look at those operations here.
### Concatenation of arrays
Concatenation, or joining of two arrays in NumPy, is primarily accomplished using the routines ``np.concatenate``, ``np.vstack``, and ``np.hstack``.
``np.concatenate`` takes a tuple or list of arrays as its first argument, as we can see here:
```
x = np.array([1, 2, 3])
y = np.array([3, 2, 1])
np.concatenate([x, y])
```
You can also concatenate more than two arrays at once:
```
z = [99, 99, 99]
print(np.concatenate([x, y, z]))
```
It can also be used for two-dimensional arrays:
```
grid = np.array([[1, 2, 3],
[4, 5, 6]])
# concatenate along the first axis
np.concatenate([grid, grid])
# concatenate along the second axis (zero-indexed)
np.concatenate([grid, grid], axis=1)
```
For working with arrays of mixed dimensions, it can be clearer to use the ``np.vstack`` (vertical stack) and ``np.hstack`` (horizontal stack) functions:
```
x = np.array([1, 2, 3])
grid = np.array([[9, 8, 7],
[6, 5, 4]])
# vertically stack the arrays
np.vstack([x, grid])
# horizontally stack the arrays
y = np.array([[99],
[99]])
np.hstack([grid, y])
```
Similary, ``np.dstack`` will stack arrays along the third axis.
### Splitting of arrays
The opposite of concatenation is splitting, which is implemented by the functions ``np.split``, ``np.hsplit``, and ``np.vsplit``. For each of these, we can pass a list of indices giving the split points:
```
x = [1, 2, 3, 99, 99, 3, 2, 1]
x1, x2, x3 = np.split(x, [3, 5])
print(x1, x2, x3)
```
Notice that _N_ split-points, leads to *N + 1* subarrays.
The related functions ``np.hsplit`` and ``np.vsplit`` are similar:
```
grid = np.arange(16).reshape((4, 4))
grid
upper, lower = np.vsplit(grid, [2])
print(upper)
print(lower)
left, right = np.hsplit(grid, [2])
print(left)
print(right)
```
Similarly, ``np.dsplit`` will split arrays along the third axis.
# Computation on NumPy Arrays: Universal Functions
Up until now, we have been discussing some of the basic nuts and bolts of NumPy; in the next few sections, we will dive into the reasons that NumPy is so important in the Python data science world.
Namely, it provides an easy and flexible interface to optimized computation with arrays of data.
Computation on NumPy arrays can be very fast, or it can be very slow.
The key to making it fast is to use *vectorized* operations, generally implemented through NumPy's *universal functions* (ufuncs).
This section motivates the need for NumPy's ufuncs, which can be used to make repeated calculations on array elements much more efficient.
It then introduces many of the most common and useful arithmetic ufuncs available in the NumPy package.
## The Slowness of Loops
Python's default implementation (known as CPython) does some operations very slowly.
This is in part due to the dynamic, interpreted nature of the language: the fact that types are flexible, so that sequences of operations cannot be compiled down to efficient machine code as in languages like C and Fortran.
Recently there have been various attempts to address this weakness: well-known examples are the [PyPy](http://pypy.org/) project, a just-in-time compiled implementation of Python; the [Cython](http://cython.org) project, which converts Python code to compilable C code; and the [Numba](http://numba.pydata.org/) project, which converts snippets of Python code to fast LLVM bytecode.
Each of these has its strengths and weaknesses, but it is safe to say that none of the three approaches has yet surpassed the reach and popularity of the standard CPython engine.
The relative sluggishness of Python generally manifests itself in situations where many small operations are being repeated – for instance looping over arrays to operate on each element.
For example, imagine we have an array of values and we'd like to compute the reciprocal of each.
A straightforward approach might look like this:
```
import numpy as np
np.random.seed(0)
def compute_reciprocals(values):
output = np.empty(len(values))
for i in range(len(values)):
output[i] = 1.0 / values[i]
return output
values = np.random.randint(1, 10, size=5)
compute_reciprocals(values)
```
This implementation probably feels fairly natural to someone from, say, a C or Java background.
But if we measure the execution time of this code for a large input, we see that this operation is very slow, perhaps surprisingly so!
We'll benchmark this with IPython's ``%timeit`` magic:
```
big_array = np.random.randint(1, 100, size=1000000)
%timeit compute_reciprocals(big_array)
```
It takes several seconds to compute these million operations and to store the result!
When even cell phones have processing speeds measured in Giga-FLOPS (i.e., billions of numerical operations per second), this seems almost absurdly slow.
It turns out that the bottleneck here is not the operations themselves, but the type-checking and function dispatches that CPython must do at each cycle of the loop.
Each time the reciprocal is computed, Python first examines the object's type and does a dynamic lookup of the correct function to use for that type.
If we were working in compiled code instead, this type specification would be known before the code executes and the result could be computed much more efficiently.
## Introducing UFuncs
For many types of operations, NumPy provides a convenient interface into just this kind of statically typed, compiled routine. This is known as a *vectorized* operation.
This can be accomplished by simply performing an operation on the array, which will then be applied to each element.
This vectorized approach is designed to push the loop into the compiled layer that underlies NumPy, leading to much faster execution.
Compare the results of the following two:
```
print(compute_reciprocals(values))
print(1.0 / values)
```
Looking at the execution time for our big array, we see that it completes orders of magnitude faster than the Python loop:
```
%timeit (1.0 / big_array)
```
Vectorized operations in NumPy are implemented via *ufuncs*, whose main purpose is to quickly execute repeated operations on values in NumPy arrays.
Ufuncs are extremely flexible – before we saw an operation between a scalar and an array, but we can also operate between two arrays:
```
np.arange(5) / np.arange(1, 6)
```
And ufunc operations are not limited to one-dimensional arrays–they can also act on multi-dimensional arrays as well:
```
x = np.arange(9).reshape((3, 3))
2 ** x
```
Computations using vectorization through ufuncs are nearly always more efficient than their counterpart implemented using Python loops, especially as the arrays grow in size.
Any time you see such a loop in a Python script, you should consider whether it can be replaced with a vectorized expression.
## Exploring NumPy's UFuncs
Ufuncs exist in two flavors: *unary ufuncs*, which operate on a single input, and *binary ufuncs*, which operate on two inputs.
We'll see examples of both these types of functions here.
### Array arithmetic
NumPy's ufuncs feel very natural to use because they make use of Python's native arithmetic operators.
The standard addition, subtraction, multiplication, and division can all be used:
```
x = np.arange(4)
print("x =", x)
print("x + 5 =", x + 5)
print("x - 5 =", x - 5)
print("x * 2 =", x * 2)
print("x / 2 =", x / 2)
print("x // 2 =", x // 2) # floor division
```
There is also a unary ufunc for negation, and a ``**`` operator for exponentiation, and a ``%`` operator for modulus:
```
print("-x = ", -x)
print("x ** 2 = ", x ** 2)
print("x % 2 = ", x % 2)
```
In addition, these can be strung together however you wish, and the standard order of operations is respected:
```
-(0.5*x + 1) ** 2
```
Each of these arithmetic operations are simply convenient wrappers around specific functions built into NumPy; for example, the ``+`` operator is a wrapper for the ``add`` function:
```
np.add(x, 2)
```
The following table lists the arithmetic operators implemented in NumPy:
| Operator | Equivalent ufunc | Description |
|---------------|---------------------|---------------------------------------|
|``+`` |``np.add`` |Addition (e.g., ``1 + 1 = 2``) |
|``-`` |``np.subtract`` |Subtraction (e.g., ``3 - 2 = 1``) |
|``-`` |``np.negative`` |Unary negation (e.g., ``-2``) |
|``*`` |``np.multiply`` |Multiplication (e.g., ``2 * 3 = 6``) |
|``/`` |``np.divide`` |Division (e.g., ``3 / 2 = 1.5``) |
|``//`` |``np.floor_divide`` |Floor division (e.g., ``3 // 2 = 1``) |
|``**`` |``np.power`` |Exponentiation (e.g., ``2 ** 3 = 8``) |
|``%`` |``np.mod`` |Modulus/remainder (e.g., ``9 % 4 = 1``)|
Additionally there are Boolean/bitwise operators; we will explore these in [Comparisons, Masks, and Boolean Logic](02.06-Boolean-Arrays-and-Masks.ipynb).
### Absolute value
Just as NumPy understands Python's built-in arithmetic operators, it also understands Python's built-in absolute value function:
```
x = np.array([-2, -1, 0, 1, 2])
abs(x)
```
The corresponding NumPy ufunc is ``np.absolute``, which is also available under the alias ``np.abs``:
```
np.absolute(x)
np.abs(x)
```
This ufunc can also handle complex data, in which the absolute value returns the magnitude:
```
x = np.array([3 - 4j, 4 - 3j, 2 + 0j, 0 + 1j])
np.abs(x)
```
### Trigonometric functions
NumPy provides a large number of useful ufuncs, and some of the most useful for the data scientist are the trigonometric functions.
We'll start by defining an array of angles:
```
theta = np.linspace(0, np.pi, 3)
```
Now we can compute some trigonometric functions on these values:
```
print("theta = ", theta)
print("sin(theta) = ", np.sin(theta))
print("cos(theta) = ", np.cos(theta))
print("tan(theta) = ", np.tan(theta))
```
The values are computed to within machine precision, which is why values that should be zero do not always hit exactly zero.
Inverse trigonometric functions are also available:
```
x = [-1, 0, 1]
print("x = ", x)
print("arcsin(x) = ", np.arcsin(x))
print("arccos(x) = ", np.arccos(x))
print("arctan(x) = ", np.arctan(x))
```
### Exponents and logarithms
Another common type of operation available in a NumPy ufunc are the exponentials:
```
x = [1, 2, 3]
print("x =", x)
print("e^x =", np.exp(x))
print("2^x =", np.exp2(x))
print("3^x =", np.power(3, x))
```
The inverse of the exponentials, the logarithms, are also available.
The basic ``np.log`` gives the natural logarithm; if you prefer to compute the base-2 logarithm or the base-10 logarithm, these are available as well:
```
x = [1, 2, 4, 10]
print("x =", x)
print("ln(x) =", np.log(x))
print("log2(x) =", np.log2(x))
print("log10(x) =", np.log10(x))
```
# Aggregations: Min, Max, and Everything In Between
Often when faced with a large amount of data, a first step is to compute summary statistics for the data in question.
Perhaps the most common summary statistics are the mean and standard deviation, which allow you to summarize the "typical" values in a dataset, but other aggregates are useful as well (the sum, product, median, minimum and maximum, quantiles, etc.).
NumPy has fast built-in aggregation functions for working on arrays; we'll discuss and demonstrate some of them here.
## Summing the Values in an Array
As a quick example, consider computing the sum of all values in an array.
Python itself can do this using the built-in ``sum`` function:
```
import numpy as np
L = np.random.random(100)
sum(L)
```
The syntax is quite similar to that of NumPy's ``sum`` function, and the result is the same in the simplest case:
```
np.sum(L)
```
However, because it executes the operation in compiled code, NumPy's version of the operation is computed much more quickly:
```
big_array = np.random.rand(1000000)
%timeit sum(big_array)
%timeit np.sum(big_array)
```
Be careful, though: the ``sum`` function and the ``np.sum`` function are not identical, which can sometimes lead to confusion!
In particular, their optional arguments have different meanings, and ``np.sum`` is aware of multiple array dimensions, as we will see in the following section.
## Minimum and Maximum
Similarly, Python has built-in ``min`` and ``max`` functions, used to find the minimum value and maximum value of any given array:
```
min(big_array), max(big_array)
```
NumPy's corresponding functions have similar syntax, and again operate much more quickly:
```
np.min(big_array), np.max(big_array)
%timeit min(big_array)
%timeit np.min(big_array)
```
For ``min``, ``max``, ``sum``, and several other NumPy aggregates, a shorter syntax is to use methods of the array object itself:
```
print(big_array.min(), big_array.max(), big_array.sum())
```
Whenever possible, make sure that you are using the NumPy version of these aggregates when operating on NumPy arrays!
### Multi dimensional aggregates
One common type of aggregation operation is an aggregate along a row or column.
Say you have some data stored in a two-dimensional array:
```
M = np.random.random((3, 4))
print(M)
```
By default, each NumPy aggregation function will return the aggregate over the entire array:
```
M.sum()
```
Aggregation functions take an additional argument specifying the *axis* along which the aggregate is computed. For example, we can find the minimum value within each column by specifying ``axis=0``:
```
M.min(axis=0)
```
The function returns four values, corresponding to the four columns of numbers.
Similarly, we can find the maximum value within each row:
```
M.max(axis=1)
```
The way the axis is specified here can be confusing to users coming from other languages.
The ``axis`` keyword specifies the *dimension of the array that will be collapsed*, rather than the dimension that will be returned.
So specifying ``axis=0`` means that the first axis will be collapsed: for two-dimensional arrays, this means that values within each column will be aggregated.
### Other aggregation functions
NumPy provides many other aggregation functions, but we won't discuss them in detail here.
Additionally, most aggregates have a ``NaN``-safe counterpart that computes the result while ignoring missing values, which are marked by the special IEEE floating-point ``NaN`` value (for a fuller discussion of missing data, see [Handling Missing Data](03.04-Missing-Values.ipynb)).
Some of these ``NaN``-safe functions were not added until NumPy 1.8, so they will not be available in older NumPy versions.
The following table provides a list of useful aggregation functions available in NumPy:
|Function Name | NaN-safe Version | Description |
|-------------------|---------------------|-----------------------------------------------|
| ``np.sum`` | ``np.nansum`` | Compute sum of elements |
| ``np.prod`` | ``np.nanprod`` | Compute product of elements |
| ``np.mean`` | ``np.nanmean`` | Compute mean of elements |
| ``np.std`` | ``np.nanstd`` | Compute standard deviation |
| ``np.var`` | ``np.nanvar`` | Compute variance |
| ``np.min`` | ``np.nanmin`` | Find minimum value |
| ``np.max`` | ``np.nanmax`` | Find maximum value |
| ``np.argmin`` | ``np.nanargmin`` | Find index of minimum value |
| ``np.argmax`` | ``np.nanargmax`` | Find index of maximum value |
| ``np.median`` | ``np.nanmedian`` | Compute median of elements |
| ``np.percentile`` | ``np.nanpercentile``| Compute rank-based statistics of elements |
| ``np.any`` | N/A | Evaluate whether any elements are true |
| ``np.all`` | N/A | Evaluate whether all elements are true |
We will see these aggregates often throughout the rest of the book.
## Comparison Operators as ufuncs
We introduced ufuncs, and focused in particular on arithmetic operators. We saw that using ``+``, ``-``, ``*``, ``/``, and others on arrays leads to element-wise operations.
NumPy also implements comparison operators such as ``<`` (less than) and ``>`` (greater than) as element-wise ufuncs.
The result of these comparison operators is always an array with a Boolean data type.
All six of the standard comparison operations are available:
```
x = np.array([1, 2, 3, 4, 5])
x < 3 # less than
x > 3 # greater than
x <= 3 # less than or equal
x >= 3 # greater than or equal
x != 3 # not equal
x == 3 # equal
```
It is also possible to do an element-wise comparison of two arrays, and to include compound expressions:
```
(2 * x) == (x ** 2)
```
As in the case of arithmetic operators, the comparison operators are implemented as ufuncs in NumPy; for example, when you write ``x < 3``, internally NumPy uses ``np.less(x, 3)``.
A summary of the comparison operators and their equivalent ufunc is shown here:
| Operator | Equivalent ufunc |-| Operator | Equivalent ufunc |
|---------------|---------------------|---|---------------|---------------------|
|``==`` |``np.equal`` | -|``!=`` |``np.not_equal`` |
|``<`` |``np.less`` | -|``<=`` |``np.less_equal`` |
|``>`` |``np.greater`` | -|``>=`` |``np.greater_equal`` |
Using comparison operators to subset an array:
```
x[x > 3]
```
## Fast Sorting in NumPy: ``np.sort`` and ``np.argsort``
Although Python has built-in ``sort`` and ``sorted`` functions to work with lists, we won't discuss them here because NumPy's ``np.sort`` function turns out to be much more efficient and useful for our purposes.
By default ``np.sort`` uses an $\mathcal{O}[N\log N]$, *quicksort* algorithm, though *mergesort* and *heapsort* are also available. For most applications, the default quicksort is more than sufficient.
To return a sorted version of the array without modifying the input, you can use ``np.sort``:
```
x = np.array([2, 1, 4, 3, 5])
np.sort(x)
```
If you prefer to sort the array in-place, you can instead use the ``sort`` method of arrays:
```
x.sort()
print(x)
```
A related function is ``argsort``, which instead returns the *indices* of the sorted elements:
```
x = np.array([2, 1, 4, 3, 5])
i = np.argsort(x)
print(i)
```
The first element of this result gives the index of the smallest element, the second value gives the index of the second smallest, and so on.
These indices can then be used (via fancy indexing) to construct the sorted array if desired:
```
x[i]
```
### Sorting along rows or columns
A useful feature of NumPy's sorting algorithms is the ability to sort along specific rows or columns of a multidimensional array using the ``axis`` argument. For example:
```
rand = np.random.RandomState(42)
X = rand.randint(0, 10, (4, 6))
print(X)
# sort each column of X
np.sort(X, axis=0)
# sort each row of X
np.sort(X, axis=1)
```
| github_jupyter |
<img src="images/dask_horizontal.svg" align="right" width="30%">
# Distributed, Advanced
## Distributed futures
```
from dask.distributed import Client
c = Client(n_workers=4)
c.cluster
```
In the previous chapter, we showed that executing a calculation (created using delayed) with the distributed executor is identical to any other executor. However, we now have access to additional functionality, and control over what data is held in memory.
To begin, the `futures` interface (derived from the built-in `concurrent.futures`) allows map-reduce like functionality. We can submit individual functions for evaluation with one set of inputs, or evaluated over a sequence of inputs with `submit()` and `map()`. Notice that the call returns immediately, giving one or more *futures*, whose status begins as "pending" and later becomes "finished". There is no blocking of the local Python session.
Here is the simplest example of `submit` in action:
```
def inc(x):
return x + 1
fut = c.submit(inc, 1)
fut
```
We can re-execute the following cell as often as we want as a way to poll the status of the future. This could of course be done in a loop, pausing for a short time on each iteration. We could continue with our work, or view a progressbar of work still going on, or force a wait until the future is ready.
In the meantime, the `status` dashboard (link above next to the Cluster widget) has gained a new element in the task stream, indicating that `inc()` has completed, and the progress section at the problem shows one task complete and held in memory.
```
fut
```
Possible alternatives you could investigate:
```python
from dask.distributed import wait, progress
progress(fut)
```
would show a progress bar in *this* notebook, rather than having to go to the dashboard. This progress bar is also asynchronous, and doesn't block the execution of other code in the meanwhile.
```python
wait(fut)
```
would block and force the notebook to wait until the computation pointed to by `fut` was done. However, note that the result of `inc()` is sitting in the cluster, it would take **no time** to execute the computation now, because Dask notices that we are asking for the result of a computation it already knows about. More on this later.
```
# grab the information back - this blocks if fut is not ready
c.gather(fut)
# equivalent action when only considering a single future
# fut.result()
```
Here we see an alternative way to execute work on the cluster: when you submit or map with the inputs as futures, the *computation moves to the data* rather than the other way around, and the client, in the local Python session, need never see the intermediate values. This is similar to building the graph using delayed, and indeed, delayed can be used in conjunction with futures. Here we use the delayed object `total` from before.
```
# Some trivial work that takes time
# repeated from the Distributed chapter.
from dask import delayed
import time
def inc(x):
time.sleep(5)
return x + 1
def dec(x):
time.sleep(3)
return x - 1
def add(x, y):
time.sleep(7)
return x + y
x = delayed(inc)(1)
y = delayed(dec)(2)
total = delayed(add)(x, y)
# notice the difference from total.compute()
# notice that this cell completes immediately
fut = c.compute(total)
fut
c.gather(fut) # waits until result is ready
```
### `Client.submit`
`submit` takes a function and arguments, pushes these to the cluster, returning a *Future* representing the result to be computed. The function is passed to a worker process for evaluation. Note that this cell returns immediately, while computation may still be ongoing on the cluster.
```
fut = c.submit(inc, 1)
fut
```
This looks a lot like doing `compute()`, above, except now we are passing the function and arguments directly to the cluster. To anyone used to `concurrent.futures`, this will look familiar. This new `fut` behaves the same way as the one above. Note that we have now over-written the previous definition of `fut`, which will get garbage-collected, and, as a result, that previous result is released by the cluster
### Exercise: Rebuild the above delayed computation using `Client.submit` instead
The arguments passed to `submit` can be futures from other submit operations or delayed objects. The former, in particular, demonstrated the concept of *moving the computation to the data* which is one of the most powerful elements of programming with Dask.
```
# Your code here
x = c.submit(inc, 1)
y = c.submit(dec, 2)
total = c.submit(add, x, y)
print(total) # This is still a future
c.gather(total) # This blocks until the computation has finished
```
Each futures represents a result held, or being evaluated by the cluster. Thus we can control caching of intermediate values - when a future is no longer referenced, its value is forgotten. In the solution, above, futures are held for each of the function calls. These results would not need to be re-evaluated if we chose to submit more work that needed them.
We can explicitly pass data from our local session into the cluster using `scatter()`, but usually better is to construct functions that do the loading of data within the workers themselves, so that there is no need to serialise and communicate the data. Most of the loading functions within Dask, sudh as `dd.read_csv`, work this way. Similarly, we normally don't want to `gather()` results that are too big in memory.
The [full API](http://distributed.readthedocs.io/en/latest/api.html) of the distributed scheduler gives details of interacting with the cluster, which remember, can be on your local machine or possibly on a massive computational resource.
The futures API offers a work submission style that can easily emulate the map/reduce paradigm (see `c.map()`) that may be familiar to many people. The intermediate results, represented by futures, can be passed to new tasks without having to bring the pull locally from the cluster, and new work can be assigned to work on the output of previous jobs that haven't even begun yet.
Generally, any Dask operation that is executed using `.compute()` can be submitted for asynchronous execution using `c.compute()` instead, and this applies to all collections. Here is an example with the calculation previously seen in the Bag chapter. We have replaced the `.compute()` method there with the distributed client version, so, again, we could continue to submit more work (perhaps based on the result of the calculation), or, in the next cell, follow the progress of the computation. A similar progress-bar appears in the monitoring UI page.
```
%run prep.py -d accounts
import dask.bag as db
import os
import json
filename = os.path.join('data', 'accounts.*.json.gz')
lines = db.read_text(filename)
js = lines.map(json.loads)
f = c.compute(js.filter(lambda record: record['name'] == 'Alice')
.pluck('transactions')
.flatten()
.pluck('amount')
.mean())
from dask.distributed import progress
# note that progress must be the last line of a cell
# in order to show up
progress(f)
# get result.
c.gather(f)
# release values by deleting the futures
del f, fut, x, y, total
```
### Persist
Considering which data should be loaded by the workers, as opposed to passed, and which intermediate values to persist in worker memory, will in many cases determine the computation efficiency of a process.
In the example here, we repeat a calculation from the Array chapter - notice that each call to `compute()` is roughly the same speed, because the loading of the data is included every time.
```
%run prep.py -d random
import h5py
import os
f = h5py.File(os.path.join('data', 'random.hdf5'), mode='r')
dset = f['/x']
import dask.array as da
x = da.from_array(dset, chunks=(1000000,))
%time x.sum().compute()
%time x.sum().compute()
```
If, instead, we persist the data to RAM up front (this takes a few seconds to complete - we could `wait()` on this process), then further computations will be much faster.
```
# changes x from a set of delayed prescriptions
# to a set of futures pointing to data in RAM
# See this on the UI dashboard.
x = c.persist(x)
%time x.sum().compute()
%time x.sum().compute()
```
Naturally, persisting every intermediate along the way is a bad idea, because this will tend to fill up all available RAM and make the whole system slow (or break!). The ideal persist point is often at the end of a set of data cleaning steps, when the data is in a form which will get queried often.
**Exercise**: how is the memory associated with `x` released, once we know we are done with it?
## Asynchronous computation
<img style="float: right;" src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/32/Rosenbrock_function.svg/450px-Rosenbrock_function.svg.png" height=200 width=200>
One benefit of using the futures API is that you can have dynamic computations that adjust as things progress. Here we implement a simple naive search by looping through results as they come in, and submit new points to compute as others are still running.
Watching the [diagnostics dashboard](../../9002/status) as this runs you can see computations are being concurrently run while more are being submitted. This flexibility can be useful for parallel algorithms that require some level of synchronization.
Lets perform a very simple minimization using dynamic programming. The function of interest is known as Rosenbrock:
```
# a simple function with interesting minima
import time
def rosenbrock(point):
"""Compute the rosenbrock function and return the point and result"""
time.sleep(0.1)
score = (1 - point[0])**2 + 2 * (point[1] - point[0]**2)**2
return point, score
```
Initial setup, including creating a graphical figure. We use Bokeh for this, which allows for dynamic update of the figure as results come in.
```
from bokeh.io import output_notebook, push_notebook
from bokeh.models.sources import ColumnDataSource
from bokeh.plotting import figure, show
import numpy as np
output_notebook()
# set up plot background
N = 500
x = np.linspace(-5, 5, N)
y = np.linspace(-5, 5, N)
xx, yy = np.meshgrid(x, y)
d = (1 - xx)**2 + 2 * (yy - xx**2)**2
d = np.log(d)
p = figure(x_range=(-5, 5), y_range=(-5, 5))
p.image(image=[d], x=-5, y=-5, dw=10, dh=10, palette="Spectral11");
```
We start off with a point at (0, 0), and randomly scatter test points around it. Each evaluation takes ~100ms, and as result come in, we test to see if we have a new best point, and choose random points around that new best point, as the search box shrinks.
We print the function value and current best location each time we have a new best value.
```
from dask.distributed import as_completed
from random import uniform
scale = 5 # Intial random perturbation scale
best_point = (0, 0) # Initial guess
best_score = float('inf') # Best score so far
startx = [uniform(-scale, scale) for _ in range(10)]
starty = [uniform(-scale, scale) for _ in range(10)]
# set up plot
source = ColumnDataSource({'x': startx, 'y': starty, 'c': ['grey'] * 10})
p.circle(source=source, x='x', y='y', color='c')
t = show(p, notebook_handle=True)
# initial 10 random points
futures = [c.submit(rosenbrock, (x, y)) for x, y in zip(startx, starty)]
iterator = as_completed(futures)
for res in iterator:
# take a completed point, is it an improvement?
point, score = res.result()
if score < best_score:
best_score, best_point = score, point
print(score, point)
x, y = best_point
newx, newy = (x + uniform(-scale, scale), y + uniform(-scale, scale))
# update plot
source.stream({'x': [newx], 'y': [newy], 'c': ['grey']}, rollover=20)
push_notebook(document=t)
# add new point, dynamically, to work on the cluster
new_point = c.submit(rosenbrock, (newx, newy))
iterator.add(new_point) # Start tracking new task as well
# Narrow search and consider stopping
scale *= 0.99
if scale < 0.001:
break
point
```
## Debugging
When something goes wrong in a distributed job, it is hard to figure out what the problem was and what to do about it. When a task raises an exception, the exception will show up when that result, or other result that depend upon it, is gathered.
Consider the following delayed calculation to be computed by the cluster. As usual, we get back a future, which the cluster is working on to compute (this happens very slowly for the trivial procedure).
```
@delayed
def ratio(a, b):
return a // b
ina = [5, 25, 30]
inb = [5, 5, 6]
out = delayed(sum)([ratio(a, b) for (a, b) in zip(ina, inb)])
f = c.compute(out)
f
```
We only get to know what happened when we gather the result (this is also true for `out.compute()`, except we could not have done other stuff in the meantime). For the first set of inputs, it works fine.
```
c.gather(f)
```
But if we introduce bad input, an exception is raised. The exception happens in `ratio`, but only comes to our attention when calculating the sum.
```
ina = [5, 25, 30]
inb = [5, 0, 6]
out = delayed(sum)([ratio(a, b) for (a, b) in zip(ina, inb)])
f = c.compute(out)
c.gather(f)
```
The display in this case makes the origin of the exception obvious, but this is not always the case. How should this be debugged, how would we go about finding out the exact conditions that caused the exception?
The first step, of course, is to write well-tested code which makes appropriate assertions about its input and clear warnings and error messages when something goes wrong. This applies to all code.
The most typical thing to do is to execute some portion of the computation in the local thread, so that we can run the Python debugger and query the state of things at the time that the exception happened. Obviously, this cannot be performed on the whole data-set when dealing with Big Data on a cluster, but a suitable sample will probably do even then.
```
import dask
with dask.config.set(scheduler="sync"):
# do NOT use c.compute(out) here - we specifically do not
# want the distributed scheduler
out.compute()
# uncomment to enter post-mortem debugger
# %debug
```
The trouble with this approach is that Dask is meant for the execution of large datasets/computations - you probably can't simply run the whole thing
in one local thread, else you wouldn't have used Dask in the first place. So the code above should only be used on a small part of the data that also exihibits the error.
Furthermore, the method will not work when you are dealing with futures (such as `f`, above, or after persisting) instead of delayed-based computations.
As an alternative, you can ask the scheduler to analyze your calculation and find the specific sub-task responsible for the error, and pull only it and its dependnecies locally for execution.
```
c.recreate_error_locally(f)
# uncomment to enter post-mortem debugger
# %debug
```
Finally, there are errors other than exceptions, when we need to look at the state of the scheduler/workers. In the standard "LocalCluster" we started, we
have direct access to these.
```
[(k, v.state) for k, v in c.cluster.scheduler.tasks.items() if v.exception is not None]
```
| github_jupyter |
# Db2 Jupyter Notebook Extensions Tutorial
Updated: 2019-10-03
The SQL code tutorials for Db2 rely on a Jupyter notebook extension, commonly refer to as a "magic" command. The beginning of all of the notebooks begin with the following command which will load the extension and allow the remainder of the notebook to use the %sql magic command.
<pre>
%run db2.ipynb
</pre>
The cell below will load the Db2 extension. Note that it will take a few seconds for the extension to load, so you should generally wait until the "Db2 Extensions Loaded" message is displayed in your notebook.
```
%run ../db2.ipynb
%run ../connection.ipynb
```
## Options
There are four options that can be set with the **`%sql`** command. These options are shown below with the default value shown in parenthesis.
- **`MAXROWS n (10)`** - The maximum number of rows that will be displayed before summary information is shown. If the answer set is less than this number of rows, it will be completely shown on the screen. If the answer set is larger than this amount, only the first 5 rows and last 5 rows of the answer set will be displayed. If you want to display a very large answer set, you may want to consider using the grid option `-g` to display the results in a scrollable table. If you really want to show all results then setting MAXROWS to -1 will return all output.
- **`MAXGRID n (5)`** - The maximum size of a grid display. When displaying a result set in a grid `-g`, the default size of the display window is 5 rows. You can set this to a larger size so that more rows are shown on the screen. Note that the minimum size always remains at 5 which means that if the system is unable to display your maximum row size it will reduce the table display until it fits.
- **`DISPLAY PANDAS | GRID (PANDAS)`** - Display the results as a PANDAS dataframe (default) or as a scrollable GRID
- **`RUNTIME n (1)`** - When using the timer option on a SQL statement, the statement will execute for **`n`** number of seconds. The result that is returned is the number of times the SQL statement executed rather than the execution time of the statement. The default value for runtime is one second, so if the SQL is very complex you will need to increase the run time.
- **`LIST`** - Display the current settings
To set an option use the following syntax:
```
%sql option option_name value option_name value ....
```
The following example sets all options:
```
%sql option maxrows 100 runtime 2 display grid maxgrid 10
```
The values will **not** be saved between Jupyter notebooks sessions. If you need to retrieve the current options values, use the LIST command as the only argument:
```
%sql option list
```
## Connections to Db2
Before any SQL commands can be issued, a connection needs to be made to the Db2 database that you will be using. The connection can be done manually (through the use of the CONNECT command), or automatically when the first `%sql` command is issued.
The Db2 magic command tracks whether or not a connection has occured in the past and saves this information between notebooks and sessions. When you start up a notebook and issue a command, the program will reconnect to the database using your credentials from the last session. In the event that you have not connected before, the system will prompt you for all the information it needs to connect. This information includes:
- Database name (SAMPLE)
- Hostname - localhost (enter an IP address if you need to connect to a remote server)
- PORT - 50000 (this is the default but it could be different)
- Userid - DB2INST1
- Password - No password is provided so you have to enter a value
- Maximum Rows - 10 lines of output are displayed when a result set is returned
There will be default values presented in the panels that you can accept, or enter your own values. All of the information will be stored in the directory that the notebooks are stored on. Once you have entered the information, the system will attempt to connect to the database for you and then you can run all of the SQL scripts. More details on the CONNECT syntax will be found in a section below.
If you have credentials available from Db2 on Cloud or DSX, place the contents of the credentials into a variable and then use the `CONNECT CREDENTIALS <var>` syntax to connect to the database.
```Python
db2blu = { "uid" : "xyz123456", ...}
%sql CONNECT CREDENTIALS db2blu
```
If the connection is successful using the credentials, the variable will be saved to disk so that you can connected from within another notebook using the same syntax.
The next statement will force a CONNECT to occur with the default values. If you have not connected before, it will prompt you for the information.
```
%sql CONNECT
```
## Line versus Cell Command
The Db2 extension is made up of one magic command that works either at the LINE level (`%sql`) or at the CELL level (`%%sql`). If you only want to execute a SQL command on one line in your script, use the `%sql` form of the command. If you want to run a larger block of SQL, then use the `%%sql` form. Note that when you use the `%%sql` form of the command, the entire contents of the cell is considered part of the command, so you cannot mix other commands in the cell.
The following is an example of a line command:
```
%sql VALUES 'HELLO THERE'
```
If you have SQL that requires multiple lines, of if you need to execute many lines of SQL, then you should
be using the CELL version of the `%sql` command. To start a block of SQL, start the cell with `%%sql` and do not place any SQL following the command. Subsequent lines can contain SQL code, with each SQL statement delimited with the semicolon (`;`). You can change the delimiter if required for procedures, etc... More details on this later.
```
%%sql
VALUES
1,
2,
3
```
If you are using a single statement then there is no need to use a delimiter. However, if you are combining a number of commands then you must use the semicolon.
```
%%sql
DROP TABLE STUFF;
CREATE TABLE STUFF (A INT);
INSERT INTO STUFF VALUES
1,2,3;
SELECT * FROM STUFF;
```
The script will generate messages and output as it executes. Each SQL statement that generates results will have a table displayed with the result set. If a command is executed, the results of the execution get listed as well. The script you just ran probably generated an error on the DROP table command.
## Options
Both forms of the `%sql` command have options that can be used to change the behavior of the code. For both forms of the command (`%sql`, `%%sql`), the options must be on the same line as the command:
<pre>
%sql -t ...
%%sql -t
</pre>
The only difference is that the `%sql` command can have SQL following the parameters, while the `%%sql` requires the SQL to be placed on subsequent lines.
There are a number of parameters that you can specify as part of the `%sql` statement.
* `-d` - Use alternative statement delimiter `@`
* `-t,-time` - Time the statement execution
* `-q,-quiet` - Suppress messages
* `-j` - JSON formatting of the first column
* `-json` - Retrieve the result set as a JSON record
* `-a,-all` - Show all output
* `-pb,-bar` - Bar chart of results
* `-pp,-pie` - Pie chart of results
* `-pl,-line` - Line chart of results
* `-sampledata` Load the database with the sample EMPLOYEE and DEPARTMENT tables
* `-r,-array` - Return the results into a variable (list of rows)
* `-e,-echo` - Echo macro substitution
* `-h,-help` - Display help information
* `-grid` - Display results in a scrollable grid
Multiple parameters are allowed on a command line. Each option should be separated by a space:
<pre>
%sql -a -j ...
</pre>
A `SELECT` statement will return the results as a dataframe and display the results as a table in the notebook. If you use the assignment statement, the dataframe will be placed into the variable and the results will not be displayed:
<pre>
r = %sql SELECT * FROM EMPLOYEE
</pre>
The sections below will explain the options in more detail.
## Delimiters
The default delimiter for all SQL statements is the semicolon. However, this becomes a problem when you try to create a trigger, function, or procedure that uses SQLPL (or PL/SQL). Use the `-d` option to turn the SQL delimiter into the at (`@`) sign and `-q` to suppress error messages. The semi-colon is then ignored as a delimiter.
For example, the following SQL will use the `@` sign as the delimiter.
```
%%sql -d -q
DROP TABLE STUFF
@
CREATE TABLE STUFF (A INT)
@
INSERT INTO STUFF VALUES
1,2,3
@
SELECT * FROM STUFF
@
```
The delimiter change will only take place for the statements following the `%%sql` command. Subsequent cells
in the notebook will still use the semicolon. You must use the `-d` option for every cell that needs to use the
semicolon in the script.
## Limiting Result Sets
The default number of rows displayed for any result set is 10. You have the option of changing this option when initially connecting to the database. If you want to override the number of rows display you can either update
the control variable, or use the -a option. The `-a` option will display all of the rows in the answer set. For instance, the following SQL will only show 10 rows even though we inserted 15 values:
```
%sql values 1,2,3,4,5,6,7,8,9,10,11,12,13,14,15
```
You will notice that the displayed result will split the visible rows to the first 5 rows and the last 5 rows.
Using the `-a` option will display all of the values.
```
%sql -a values 1,2,3,4,5,6,7,8,9,10,11,12,13,14,15
```
If you want a scrollable list, use the `-grid` option.
```
%sql -grid values 1,2,3,4,5,6,7,8,9,10,11,12,13,14,15
```
To change the default value of rows displayed, you can use the `%sql option maxrow` command to set the value to something else. A value of -1 means unlimited output. Note that `MAXROWS` will display all of the data for answer sets that are less than `MAXROWS` in size. For instance, if you set `MAXROWS` to 20, then any answer set less than or equal to 20 will be shown on the screen. Anything larger than this amount will be summarized with the first `MAXROWS/2` rows displayed followed by the last `MAXROWS/2` rows.
The following example will set the maximum rows to 8. Since our answer set is greater than 8, only the first 4 (8/2) rows will be shown, followed by the last 4.
```
%sql option maxrows 8
%sql values 1,2,3,4,5,6,7,8,9,10,11,12,13,14,15
```
For a grid display `-grid -g`, the `MAXGRID` setting will try to display the scrollable table with *at least* `MAXGRID` rows. The minimum display size of a table is 5 rows so if the table can't fit on the screen it will try to force at least 5 rows to be displayed. The size of the table display does not impact your ability to use the scrollbars to see the entire answer set.
```
%sql option maxrows 10
%sql values 1,2,3,4,5,6,7,8,9,10,11,12,13,14,15
```
A special note regarding the output from a `SELECT` statement. If the SQL statement is the last line of a block, the results will be displayed by default (unless you assigned the results to a variable). If the SQL is in the middle of a block of statements, the results will not be displayed.
## Quiet Mode
Every SQL statement will result in some output. You will either get an answer set (`SELECT`), or an indication if
the command worked. For instance, the following set of SQL will generate some error messages since the tables
will probably not exist:
```
%%sql
DROP TABLE TABLE_NOT_FOUND;
DROP TABLE TABLE_SPELLED_WRONG;
```
If you know that these errors may occur you can silence them with the -q option.
```
%%sql -q
DROP TABLE TABLE_NOT_FOUND;
DROP TABLE TABLE_SPELLED_WRONG;
```
SQL output will not be suppressed, so the following command will still show the results.
```
%%sql -q
DROP TABLE TABLE_NOT_FOUND;
DROP TABLE TABLE_SPELLED_WRONG;
VALUES 1,2,3;
```
## Variables in %sql Blocks
Python variables can be passed to a `%sql` line command, and to a `%%sql` block. For both forms of the `%sql` command you can pass variables by placing a colon in front of the variable name.
```python
%sql SELECT * FROM EMPLOYEE WHERE EMPNO = :empno
```
The following example illustrates the use of a variable in the SQL.
```
empno = '000010'
%sql SELECT * FROM EMPLOYEE WHERE EMPNO = :empno
```
You can doublecheck that the substitution took place by using the `-e` option which echos the SQL command after substitution.
```
%sql -echo SELECT * FROM EMPLOYEE WHERE EMPNO = :empno
```
Note that the variable `:empno` did not have quotes around it, although it is a string value. The `%sql` call will examine the contents of the variable and add quotes around strings so you do not have to supply them in the SQL command.
Variables can also be array types. Arrays are expanded into multiple values, each separated by commas. This is useful when building SQL `IN` lists. The following example searches for 3 employees based on their employee number.
```
empnos = ['000010','000020','000030']
%sql SELECT * FROM EMPLOYEE WHERE EMPNO IN (:empnos)
```
You can reference individual array items using this technique as well. If you wanted to search for only the first value in the `empnos` array, use `:empnos[0]` instead.
```
%sql SELECT * FROM EMPLOYEE WHERE EMPNO IN (:empnos[0])
```
One final type of variable substitution that is allowed is for dictionaries. Python dictionaries resemble JSON objects and can be used to insert JSON values into Db2. For instance, the following variable contains company information in a JSON structure.
```
customer = {
"name" : "Aced Hardware Stores",
"city" : "Rockwood",
"employees" : 14
}
```
Db2 has builtin functions for dealing with JSON objects. There is another Jupyter notebook which goes through this in detail. Rather than using those functions, the following code will create a Db2 table with a string column that will contain the contents of this JSON record.
```
%%sql
DROP TABLE SHOWJSON;
CREATE TABLE SHOWJSON (INJSON VARCHAR(256));
```
To insert the Dictionary (JSON Record) into this Db2 table, you only need to use the variable name as one of the fields being inserted.
```
%sql INSERT INTO SHOWJSON VALUES :customer
```
Selecting from this table will show that the data has been inserted as a string.
```
%sql select * from showjson
```
If you want to retrieve the data from a column that contains JSON records, you must use the `-j` flag to insert the contents back into a variable.
```
v = %sql -j SELECT * FROM SHOWJSON
```
The variable `v` now contains the original JSON record for you to use.
```
v
```
## SQL Character Strings
Character strings require special handling when dealing with Db2. The single quote character `'` is reserved for delimiting string constants, while the double quote `"` is used for naming columns that require special characters. You cannot use the double quote character to delimit strings that happen to contain the single quote character. What Db2 requires you do is placed two quotes in a row to have them interpreted as a single quote character. For instance, the next statement will select one employee from the table who has a quote in their last name: `O'CONNELL`.
```
%sql SELECT * FROM EMPLOYEE WHERE LASTNAME = 'O''CONNELL'
```
Python handles quotes differently! You can assign a string to a Python variable using single or double quotes. The following assignment statements are not identical!
```
lastname = "O'CONNELL"
print(lastname)
lastname = 'O''CONNELL'
print(lastname)
```
If you use the same syntax as Db2, Python will remove the quote in the string! It interprets this as two strings (O and CONNELL) being concatentated together. That probably isn't what you want! So the safest approach is to use double quotes around your string when you assign it to a variable. Then you can use the variable in the SQL statement as shown in the following example.
```
lastname = "O'CONNELL"
%sql -e SELECT * FROM EMPLOYEE WHERE LASTNAME = :lastname
```
Notice how the string constant was updated to contain two quotes when inserted into the SQL statement. This is done automatically by the `%sql` magic command, so there is no need to use the two single quotes when assigning a string to a variable. However, you must use the two single quotes when using constants in a SQL statement.
## Builtin Variables
There are 5 predefined variables defined in the program:
- database - The name of the database you are connected to
- uid - The userid that you connected with
- hostname = The IP address of the host system
- port - The port number of the host system
- max - The maximum number of rows to return in an answer set
Theses variables are all part of a structure called _settings. To retrieve a value, use the syntax:
```python
db = _settings['database']
```
There are also 3 variables that contain information from the last SQL statement that was executed.
- sqlcode - SQLCODE from the last statement executed
- sqlstate - SQLSTATE from the last statement executed
- sqlerror - Full error message returned on last statement executed
You can access these variables directly in your code. The following code segment illustrates the use of the SQLCODE variable.
```
empnos = ['000010','999999']
for empno in empnos:
ans1 = %sql -r SELECT SALARY FROM EMPLOYEE WHERE EMPNO = :empno
if (sqlcode != 0):
print("Employee "+ empno + " left the company!")
else:
print("Employee "+ empno + " salary is " + str(ans1[1][0]))
```
## Timing SQL Statements
Sometimes you want to see how the execution of a statement changes with the addition of indexes or other
optimization changes. The `-t` option will run the statement on the LINE or one SQL statement in the CELL for
exactly one second. The results will be displayed and optionally placed into a variable. The syntax of the
command is:
<pre>
sql_time = %sql -t SELECT * FROM EMPLOYEE
</pre>
For instance, the following SQL will time the VALUES clause.
```
%sql -t VALUES 1,2,3,4,5,6,7,8,9
```
When timing a statement, no output will be displayed. If your SQL statement takes longer than one second you
will need to modify the runtime options. You can use the `%sql option runtime` command to change the duration the statement runs.
```
%sql option runtime 5
%sql -t VALUES 1,2,3,4,5,6,7,8,9
%sql option runtime 1
```
## JSON Formatting
Db2 supports querying JSON that is stored in a column within a table. Standard output would just display the
JSON as a string. For instance, the following statement would just return a large string of output.
```
%%sql
VALUES
'{
"empno":"000010",
"firstnme":"CHRISTINE",
"midinit":"I",
"lastname":"HAAS",
"workdept":"A00",
"phoneno":[3978],
"hiredate":"01/01/1995",
"job":"PRES",
"edlevel":18,
"sex":"F",
"birthdate":"08/24/1963",
"pay" : {
"salary":152750.00,
"bonus":1000.00,
"comm":4220.00}
}'
```
Adding the -j option to the `%sql` (or `%%sql`) command will format the first column of a return set to better
display the structure of the document. Note that if your answer set has additional columns associated with it, they will not be displayed in this format.
```
%%sql -j
VALUES
'{
"empno":"000010",
"firstnme":"CHRISTINE",
"midinit":"I",
"lastname":"HAAS",
"workdept":"A00",
"phoneno":[3978],
"hiredate":"01/01/1995",
"job":"PRES",
"edlevel":18,
"sex":"F",
"birthdate":"08/24/1963",
"pay" : {
"salary":152750.00,
"bonus":1000.00,
"comm":4220.00}
}'
```
JSON fields can be inserted into Db2 columns using Python dictionaries. This makes the input and output of JSON fields much simpler. For instance, the following code will create a Python dictionary which is similar to a JSON record.
```
employee = {
"firstname" : "John",
"lastname" : "Williams",
"age" : 45
}
```
The field can be inserted into a character column (or BSON if you use the JSON functions) by doing a direct variable insert.
```
%%sql -q
DROP TABLE SHOWJSON;
CREATE TABLE SHOWJSON(JSONIN VARCHAR(128));
```
An insert would use a variable parameter (colon in front of the variable) instead of a character string.
```
%sql INSERT INTO SHOWJSON VALUES (:employee)
%sql SELECT * FROM SHOWJSON
```
An assignment statement to a variable will result in an equivalent Python dictionary type being created. Note that we must use the raw `-j` flag to make sure we only get the data and not a data frame.
```
x = %sql -j SELECT * FROM SHOWJSON
print("First Name is " + x[0]["firstname"] + " and the last name is " + x[0]['lastname'])
```
## Plotting
Sometimes it would be useful to display a result set as either a bar, pie, or line chart. The first one or two
columns of a result set need to contain the values need to plot the information.
The three possible plot options are:
* `-pb` - bar chart (x,y)
* `-pp` - pie chart (y)
* `-pl` - line chart (x,y)
The following data will be used to demonstrate the different charting options.
```
%sql values 1,2,3,4,5
```
Since the results only have one column, the pie, line, and bar charts will not have any labels associated with
them. The first example is a bar chart.
```
%sql -pb values 1,2,3,4,5
```
The same data as a pie chart.
```
%sql -pp values 1,2,3,4,5
```
And finally a line chart.
```
%sql -pl values 1,2,3,4,5
```
If you retrieve two columns of information, the first column is used for the labels (X axis or pie slices) and
the second column contains the data.
```
%sql -pb values ('A',1),('B',2),('C',3),('D',4),('E',5)
```
For a pie chart, the first column is used to label the slices, while the data comes from the second column.
```
%sql -pp values ('A',1),('B',2),('C',3),('D',4),('E',5)
```
Finally, for a line chart, the x contains the labels and the y values are used.
```
%sql -pl values ('A',1),('B',2),('C',3),('D',4),('E',5)
```
The following SQL will plot the number of employees per department.
```
%%sql -pb
SELECT WORKDEPT, COUNT(*)
FROM EMPLOYEE
GROUP BY WORKDEPT
```
## Sample Data
Many of the Db2 notebooks depend on two of the tables that are found in the `SAMPLE` database. Rather than
having to create the entire `SAMPLE` database, this option will create and populate the `EMPLOYEE` and
`DEPARTMENT` tables in your database. Note that if you already have these tables defined, they will not be dropped.
```
%sql -sampledata
```
## Result Sets
By default, any `%sql` block will return the contents of a result set as a table that is displayed in the notebook. The results are displayed using a feature of pandas dataframes. The following select statement demonstrates a simple result set.
```
%sql select * from employee fetch first 3 rows only
```
You can assign the result set directly to a variable.
```
x = %sql select * from employee fetch first 3 rows only
```
The variable x contains the dataframe that was produced by the `%sql` statement so you access the result set by using this variable or display the contents by just referring to it in a command line.
```
x
```
There is an additional way of capturing the data through the use of the `-r` flag.
<pre>
var = %sql -r select * from employee
</pre>
Rather than returning a dataframe result set, this option will produce a list of rows. Each row is a list itself. The column names are found in row zero (0) and the data rows start at 1. To access the first column of the first row, you would use var[1][0] to access it.
```
rows = %sql -r select * from employee fetch first 3 rows only
print(rows[1][0])
```
The number of rows in the result set can be determined by using the length function and subtracting one for the header row.
```
print(len(rows)-1)
```
If you want to iterate over all of the rows and columns, you could use the following Python syntax instead of
creating a for loop that goes from 0 to 41.
```
for row in rows:
line = ""
for col in row:
line = line + str(col) + ","
print(line)
```
If you don't want the header row, modify the first line to start at the first row instead of row zero.
```
for row in rows[1:]:
line = ""
for col in row:
line = line + str(col) + ","
print(line)
```
Since the data may be returned in different formats (like integers), you should use the str() function to convert the values to strings. Otherwise, the concatenation function used in the above example might fail. For instance, the 9th field is an education level. If you retrieve it as an individual value and try and concatenate a string to it, you get the following error.
```
try:
print("Education level="+rows[1][8])
except Exception as err:
print("Oops... Something went wrong!")
print(err)
```
You can fix this problem by adding the str function to convert the date.
```
print("Education Level="+str(rows[1][8]))
```
## Development SQL
The previous set of `%sql` and `%%sql` commands deals with SQL statements and commands that are run in an interactive manner. There is a class of SQL commands that are more suited to a development environment where code is iterated or requires changing input. The commands that are associated with this form of SQL are:
- AUTOCOMMIT
- COMMIT/ROLLBACK
- PREPARE
- EXECUTE
In addition, the `sqlcode`, `sqlstate` and `sqlerror` fields are populated after every statement so you can use these variables to test for errors.
Autocommit is the default manner in which SQL statements are executed. At the end of the successful completion of a statement, the results are commited to the database. There is no concept of a transaction where multiple DML/DDL statements are considered one transaction. The `AUTOCOMMIT` command allows you to turn autocommit `OFF` or `ON`. This means that the set of SQL commands run after the `AUTOCOMMIT OFF` command are executed are not commited to the database until a `COMMIT` or `ROLLBACK` command is issued.
`COMMIT (WORK)` will finalize all of the transactions (`COMMIT`) to the database and `ROLLBACK` will undo all of the changes. If you issue a `SELECT` statement during the execution of your block, the results will reflect all of your changes. If you `ROLLBACK` the transaction, the changes will be lost.
`PREPARE` is typically used in a situation where you want to repeatidly execute a SQL statement with different variables without incurring the SQL compilation overhead. For instance:
```
x = %sql PREPARE SELECT LASTNAME FROM EMPLOYEE WHERE EMPNO=?
for y in ['000010','000020','000030']:
%sql execute :x using :y
```
`EXECUTE` is used to execute a previously compiled statement.
## Db2 CONNECT Statement
As mentioned at the beginning of this notebook, connecting to Db2 is automatically done when you issue your first
`%sql` statement. Usually the program will prompt you with what options you want when connecting to a database. The other option is to use the CONNECT statement directly. The CONNECT statement is similar to the native Db2
CONNECT command, but includes some options that allow you to connect to databases that has not been
catalogued locally.
The CONNECT command has the following format:
<pre>
%sql CONNECT TO <database> USER <userid> USING <password | ?> HOST <ip address> PORT <port number>
</pre>
If you use a "?" for the password field, the system will prompt you for a password. This avoids typing the
password as clear text on the screen. If a connection is not successful, the system will print the error
message associated with the connect request.
If the connection is successful, the parameters are saved on your system and will be used the next time you
run a SQL statement, or when you issue the %sql CONNECT command with no parameters.
If you want to force the program to connect to a different database (with prompting), use the CONNECT RESET command. The next time you run a SQL statement, the program will prompt you for the the connection
and will force the program to reconnect the next time a SQL statement is executed.
#### Credits: IBM 2019, George Baklarz [baklarz@ca.ibm.com]
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Unicode strings
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/beta/tutorials/text/unicode"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/r2/tutorials/text/unicode.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/r2/tutorials/text/unicode.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/r2/tutorials/text/unicode.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
## Introduction
Models that process natural language often handle different languages with different character sets. *Unicode* is a standard encoding system that is used to represent character from almost all languages. Each character is encoded using a unique integer [code point](https://en.wikipedia.org/wiki/Code_point) between `0` and `0x10FFFF`. A *Unicode string* is a sequence of zero or more code points.
This tutorial shows how to represent Unicode strings in TensorFlow and manipulate them using Unicode equivalents of standard string ops. It separates Unicode strings into tokens based on script detection.
```
from __future__ import absolute_import, division, print_function, unicode_literals
!pip install tensorflow==2.0.0-beta1
import tensorflow as tf
```
## The `tf.string` data type
The basic TensorFlow `tf.string` `dtype` allows you to build tensors of byte strings.
Unicode strings are utf-8 encoded by default.
```
tf.constant(u"Thanks 😊")
```
A `tf.string` tensor can hold byte strings of varying lengths because the byte strings are treated as atomic units. The string length is not included in the tensor dimensions.
```
tf.constant([u"You're", u"welcome!"]).shape
```
Note: When using python to construct strings, the handling of unicode differs betweeen v2 and v3. In v2, unicode strings are indicated by the "u" prefix, as above. In v3, strings are unicode-encoded by default.
## Representing Unicode
There are two standard ways to represent a Unicode string in TensorFlow:
* `string` scalar — where the sequence of code points is encoded using a known [character encoding](https://en.wikipedia.org/wiki/Character_encoding).
* `int32` vector — where each position contains a single code point.
For example, the following three values all represent the Unicode string `"语言处理"` (which means "language processing" in Chinese):
```
# Unicode string, represented as a UTF-8 encoded string scalar.
text_utf8 = tf.constant(u"语言处理")
text_utf8
# Unicode string, represented as a UTF-16-BE encoded string scalar.
text_utf16be = tf.constant(u"语言处理".encode("UTF-16-BE"))
text_utf16be
# Unicode string, represented as a vector of Unicode code points.
text_chars = tf.constant([ord(char) for char in u"语言处理"])
text_chars
```
### Converting between representations
TensorFlow provides operations to convert between these different representations:
* `tf.strings.unicode_decode`: Converts an encoded string scalar to a vector of code points.
* `tf.strings.unicode_encode`: Converts a vector of code points to an encoded string scalar.
* `tf.strings.unicode_transcode`: Converts an encoded string scalar to a different encoding.
```
tf.strings.unicode_decode(text_utf8,
input_encoding='UTF-8')
tf.strings.unicode_encode(text_chars,
output_encoding='UTF-8')
tf.strings.unicode_transcode(text_utf8,
input_encoding='UTF8',
output_encoding='UTF-16-BE')
```
### Batch dimensions
When decoding multiple strings, the number of characters in each string may not be equal. The return result is a [`tf.RaggedTensor`](../../guide/ragged_tensors.ipynb), where the length of the innermost dimension varies depending on the number of characters in each string:
```
# A batch of Unicode strings, each represented as a UTF8-encoded string.
batch_utf8 = [s.encode('UTF-8') for s in
[u'hÃllo', u'What is the weather tomorrow', u'Göödnight', u'😊']]
batch_chars_ragged = tf.strings.unicode_decode(batch_utf8,
input_encoding='UTF-8')
for sentence_chars in batch_chars_ragged.to_list():
print(sentence_chars)
```
You can use this `tf.RaggedTensor` directly, or convert it to a dense `tf.Tensor` with padding or a `tf.SparseTensor` using the methods `tf.RaggedTensor.to_tensor` and `tf.RaggedTensor.to_sparse`.
```
batch_chars_padded = batch_chars_ragged.to_tensor(default_value=-1)
print(batch_chars_padded.numpy())
batch_chars_sparse = batch_chars_ragged.to_sparse()
```
When encoding multiple strings with the same lengths, a `tf.Tensor` may be used as input:
```
tf.strings.unicode_encode([[99, 97, 116], [100, 111, 103], [ 99, 111, 119]],
output_encoding='UTF-8')
```
When encoding multiple strings with varyling length, a `tf.RaggedTensor` should be used as input:
```
tf.strings.unicode_encode(batch_chars_ragged, output_encoding='UTF-8')
```
If you have a tensor with multiple strings in padded or sparse format, then convert it to a `tf.RaggedTensor` before calling `unicode_encode`:
```
tf.strings.unicode_encode(
tf.RaggedTensor.from_sparse(batch_chars_sparse),
output_encoding='UTF-8')
tf.strings.unicode_encode(
tf.RaggedTensor.from_tensor(batch_chars_padded, padding=-1),
output_encoding='UTF-8')
```
## Unicode operations
### Character length
The `tf.strings.length` operation has a parameter `unit`, which indicates how lengths should be computed. `unit` defaults to `"BYTE"`, but it can be set to other values, such as `"UTF8_CHAR"` or `"UTF16_CHAR"`, to determine the number of Unicode codepoints in each encoded `string`.
```
# Note that the final character takes up 4 bytes in UTF8.
thanks = u'Thanks 😊'.encode('UTF-8')
num_bytes = tf.strings.length(thanks).numpy()
num_chars = tf.strings.length(thanks, unit='UTF8_CHAR').numpy()
print('{} bytes; {} UTF-8 characters'.format(num_bytes, num_chars))
```
### Character substrings
Similarly, the `tf.strings.substr` operation accepts the "`unit`" parameter, and uses it to determine what kind of offsets the "`pos`" and "`len`" paremeters contain.
```
# default: unit='BYTE'. With len=1, we return a single byte.
tf.strings.substr(thanks, pos=7, len=1).numpy()
# Specifying unit='UTF8_CHAR', we return a single character, which in this case
# is 4 bytes.
print(tf.strings.substr(thanks, pos=7, len=1, unit='UTF8_CHAR').numpy())
```
### Split Unicode strings
The `tf.strings.unicode_split` operation splits unicode strings into substrings of individual characters:
```
tf.strings.unicode_split(thanks, 'UTF-8').numpy()
```
### Byte offsets for characters
To align the character tensor generated by `tf.strings.unicode_decode` with the original string, it's useful to know the offset for where each character begins. The method `tf.strings.unicode_decode_with_offsets` is similar to `unicode_decode`, except that it returns a second tensor containing the start offset of each character.
```
codepoints, offsets = tf.strings.unicode_decode_with_offsets(u"🎈🎉🎊", 'UTF-8')
for (codepoint, offset) in zip(codepoints.numpy(), offsets.numpy()):
print("At byte offset {}: codepoint {}".format(offset, codepoint))
```
## Unicode scripts
Each Unicode code point belongs to a single collection of codepoints known as a [script](https://en.wikipedia.org/wiki/Script_%28Unicode%29) . A character's script is helpful in determining which language the character might be in. For example, knowing that 'Б' is in Cyrillic script indicates that modern text containing that character is likely from a Slavic language such as Russian or Ukrainian.
TensorFlow provides the `tf.strings.unicode_script` operation to determine which script a given codepoint uses. The script codes are `int32` values corresponding to [International Components for
Unicode](http://site.icu-project.org/home) (ICU) [`UScriptCode`](http://icu-project.org/apiref/icu4c/uscript_8h.html) values.
```
uscript = tf.strings.unicode_script([33464, 1041]) # ['芸', 'Б']
print(uscript.numpy()) # [17, 8] == [USCRIPT_HAN, USCRIPT_CYRILLIC]
```
The `tf.strings.unicode_script` operation can also be applied to multidimensional `tf.Tensor`s or `tf.RaggedTensor`s of codepoints:
```
print(tf.strings.unicode_script(batch_chars_ragged))
```
## Example: Simple segmentation
Segmentation is the task of splitting text into word-like units. This is often easy when space characters are used to separate words, but some languages (like Chinese and Japanese) do not use spaces, and some languages (like German) contain long compounds that must be split in order to analyze their meaning. In web text, different languages and scripts are frequently mixed together, as in "NY株価" (New York Stock Exchange).
We can perform very rough segmentation (without implementing any ML models) by using changes in script to approximate word boundaries. This will work for strings like the "NY株価" example above. It will also work for most languages that use spaces, as the space characters of various scripts are all classified as USCRIPT_COMMON, a special script code that differs from that of any actual text.
```
# dtype: string; shape: [num_sentences]
#
# The sentences to process. Edit this line to try out different inputs!
sentence_texts = [u'Hello, world.', u'世界こんにちは']
```
First, we decode the sentences into character codepoints, and find the script identifeir for each character.
```
# dtype: int32; shape: [num_sentences, (num_chars_per_sentence)]
#
# sentence_char_codepoint[i, j] is the codepoint for the j'th character in
# the i'th sentence.
sentence_char_codepoint = tf.strings.unicode_decode(sentence_texts, 'UTF-8')
print(sentence_char_codepoint)
# dtype: int32; shape: [num_sentences, (num_chars_per_sentence)]
#
# sentence_char_scripts[i, j] is the unicode script of the j'th character in
# the i'th sentence.
sentence_char_script = tf.strings.unicode_script(sentence_char_codepoint)
print(sentence_char_script)
```
Next, we use those script identifiers to determine where word boundaries should be added. We add a word boundary at the beginning of each sentence, and for each character whose script differs from the previous character:
```
# dtype: bool; shape: [num_sentences, (num_chars_per_sentence)]
#
# sentence_char_starts_word[i, j] is True if the j'th character in the i'th
# sentence is the start of a word.
sentence_char_starts_word = tf.concat(
[tf.fill([sentence_char_script.nrows(), 1], True),
tf.not_equal(sentence_char_script[:, 1:], sentence_char_script[:, :-1])],
axis=1)
# dtype: int64; shape: [num_words]
#
# word_starts[i] is the index of the character that starts the i'th word (in
# the flattened list of characters from all sentences).
word_starts = tf.squeeze(tf.where(sentence_char_starts_word.values), axis=1)
print(word_starts)
```
We can then use those start offsets to build a `RaggedTensor` containing the list of words from all batches:
```
# dtype: int32; shape: [num_words, (num_chars_per_word)]
#
# word_char_codepoint[i, j] is the codepoint for the j'th character in the
# i'th word.
word_char_codepoint = tf.RaggedTensor.from_row_starts(
values=sentence_char_codepoint.values,
row_starts=word_starts)
print(word_char_codepoint)
```
And finally, we can segment the word codepoints `RaggedTensor` back into sentences:
```
# dtype: int64; shape: [num_sentences]
#
# sentence_num_words[i] is the number of words in the i'th sentence.
sentence_num_words = tf.reduce_sum(
tf.cast(sentence_char_starts_word, tf.int64),
axis=1)
# dtype: int32; shape: [num_sentences, (num_words_per_sentence), (num_chars_per_word)]
#
# sentence_word_char_codepoint[i, j, k] is the codepoint for the k'th character
# in the j'th word in the i'th sentence.
sentence_word_char_codepoint = tf.RaggedTensor.from_row_lengths(
values=word_char_codepoint,
row_lengths=sentence_num_words)
print(sentence_word_char_codepoint)
```
To make the final result easier to read, we can encode it back into UTF-8 strings:
```
tf.strings.unicode_encode(sentence_word_char_codepoint, 'UTF-8').to_list()
```
| github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#Objectives" data-toc-modified-id="Objectives-1"><span class="toc-item-num">1 </span>Objectives</a></span></li><li><span><a href="#Motivating--The-Bayesian-Classifier-🧐" data-toc-modified-id="Motivating--The-Bayesian-Classifier-🧐-2"><span class="toc-item-num">2 </span>Motivating The Bayesian Classifier 🧐</a></span><ul class="toc-item"><li><span><a href="#Naive-Bayes-Setup" data-toc-modified-id="Naive-Bayes-Setup-2.1"><span class="toc-item-num">2.1 </span>Naive Bayes Setup</a></span><ul class="toc-item"><li><span><a href="#What's-So-Great-About-This?" data-toc-modified-id="What's-So-Great-About-This?-2.1.1"><span class="toc-item-num">2.1.1 </span>What's So Great About This?</a></span></li></ul></li><li><span><a href="#The-Naive-Assumption" data-toc-modified-id="The-Naive-Assumption-2.2"><span class="toc-item-num">2.2 </span>The Naive Assumption</a></span></li><li><span><a href="#The-Formula" data-toc-modified-id="The-Formula-2.3"><span class="toc-item-num">2.3 </span>The Formula</a></span><ul class="toc-item"><li><span><a href="#What-Parts-Can-We-Find?" data-toc-modified-id="What-Parts-Can-We-Find?-2.3.1"><span class="toc-item-num">2.3.1 </span>What Parts Can We Find?</a></span></li></ul></li><li><span><a href="#Calculating-That-Our-Email-Is-Spam" data-toc-modified-id="Calculating-That-Our-Email-Is-Spam-2.4"><span class="toc-item-num">2.4 </span>Calculating That Our Email Is Spam</a></span></li><li><span><a href="#Extending-It-With-Multiple-Words" data-toc-modified-id="Extending-It-With-Multiple-Words-2.5"><span class="toc-item-num">2.5 </span>Extending It With Multiple Words</a></span></li></ul></li><li><span><a href="#Naive-Bayes-Modeling-Example" data-toc-modified-id="Naive-Bayes-Modeling-Example-3"><span class="toc-item-num">3 </span>Naive Bayes Modeling Example</a></span><ul class="toc-item"><li><span><a href="#Using-Bayes's-Theorem-for-Classification" data-toc-modified-id="Using-Bayes's-Theorem-for-Classification-3.1"><span class="toc-item-num">3.1 </span>Using Bayes's Theorem for Classification</a></span><ul class="toc-item"><li><span><a href="#Does-this-look-like-a-classification-problem?" data-toc-modified-id="Does-this-look-like-a-classification-problem?-3.1.1"><span class="toc-item-num">3.1.1 </span>Does this look like a classification problem?</a></span></li></ul></li><li><span><a href="#Elephant-Example" data-toc-modified-id="Elephant-Example-3.2"><span class="toc-item-num">3.2 </span>Elephant Example</a></span><ul class="toc-item"><li><span><a href="#Naive-Bayes-by-Hand" data-toc-modified-id="Naive-Bayes-by-Hand-3.2.1"><span class="toc-item-num">3.2.1 </span>Naive Bayes by Hand</a></span></li><li><span><a href="#Calculation-of-Likelihoods" data-toc-modified-id="Calculation-of-Likelihoods-3.2.2"><span class="toc-item-num">3.2.2 </span>Calculation of Likelihoods</a></span></li><li><span><a href="#Posteriors" data-toc-modified-id="Posteriors-3.2.3"><span class="toc-item-num">3.2.3 </span>Posteriors</a></span></li><li><span><a href="#More-Dimensions" data-toc-modified-id="More-Dimensions-3.2.4"><span class="toc-item-num">3.2.4 </span>More Dimensions</a></span><ul class="toc-item"><li><span><a href="#What's-"Naive"-about-This?" data-toc-modified-id="What's-"Naive"-about-This?-3.2.4.1"><span class="toc-item-num">3.2.4.1 </span>What's "Naive" about This?</a></span></li><li><span><a href="#Posteriors" data-toc-modified-id="Posteriors-3.2.4.2"><span class="toc-item-num">3.2.4.2 </span>Posteriors</a></span></li></ul></li><li><span><a href="#GaussianNB" data-toc-modified-id="GaussianNB-3.2.5"><span class="toc-item-num">3.2.5 </span><a href="https://scikit-learn.org/stable/modules/generated/sklearn.naive_bayes.GaussianNB.html" target="_blank"><code>GaussianNB</code></a></a></span></li></ul></li><li><span><a href="#Comma-Survey-Example" data-toc-modified-id="Comma-Survey-Example-3.3"><span class="toc-item-num">3.3 </span>Comma Survey Example</a></span><ul class="toc-item"><li><span><a href="#Calculating-Priors-and-Likelihoods" data-toc-modified-id="Calculating-Priors-and-Likelihoods-3.3.1"><span class="toc-item-num">3.3.1 </span>Calculating Priors and Likelihoods</a></span></li><li><span><a href="#Calculating-Posteriors" data-toc-modified-id="Calculating-Posteriors-3.3.2"><span class="toc-item-num">3.3.2 </span>Calculating Posteriors</a></span></li><li><span><a href="#Comparison-with-MultinomialNB" data-toc-modified-id="Comparison-with-MultinomialNB-3.3.3"><span class="toc-item-num">3.3.3 </span>Comparison with <a href="https://scikit-learn.org/stable/modules/generated/sklearn.naive_bayes.MultinomialNB.html" target="_blank"><code>MultinomialNB</code></a></a></span></li></ul></li></ul></li></ul></div>
```
import numpy as np
import pandas as pd
from scipy import stats
from sklearn.naive_bayes import MultinomialNB, GaussianNB
# There is also a BernoulliNB for a dataset with binary predictors
import seaborn as sns
from matplotlib import pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.metrics import plot_confusion_matrix
from sklearn.preprocessing import OneHotEncoder
%matplotlib inline
```
# Objectives
- Describe how Bayes's Theorem can be used to make predictions of a target
- Identify the appropriate variant of Naive Bayes models for a particular business problem
# Motivating The Bayesian Classifier 🧐
> Let's take a second to go through an example to get a feel for how Bayes' Theorem can help us with classification. Specifically about document classification
Spam, Spam, Spam, Spam, Spam...

> This is the classic example: detecting email spam!
**The Problem Setup**
> We get emails that can be either emails we care about (***ham*** 🐷) or emails we don't care about (***spam*** 🥫).
>
> We can probably look at the words in the email and get an idea of whether they are spam or not just by observing if they contain red-flag words 🚩
>
> We won't always be right, but if we see an email that uses word(s) that are more often associated with spam, then we can feel more confident as labeling that email as spam!
## Naive Bayes Setup
What we gotta do:
1. Look at spam and not spam (ham) emails
2. Identify words that suggest classification
3. Determine probability that words occur in each classification
4. Profit (classify new emails as "spam" or "ham")
### What's So Great About This?
- We can keep updating our belief based on the emails we detect
- Relatively simple
- Can expand to multiple words
## The Naive Assumption
$P(A,B) = P(A\cap B) = P(A)\ P(B)$ only if independent
In practice, makes sense & is usually pretty good assumption
## The Formula
Let's say the word that occurs is "cash":
$$ P(🥫 | "cash") = \frac{P("cash" | 🥫)P(🥫)}{P("cash")}$$
### What Parts Can We Find?
- $P("cash")$
* That's just the probability of finding the word "cash"! Frequency of the word!
- $P(🥫)$
* Well, we start with some data (_prior knowledge_). So frequency of the spam occurring!
- $P("cash" | 🥫)$
* How frequently "cash" is used in known spam emails. Count the frequency across all spam emails
## Calculating That Our Email Is Spam
```
# Let's just say 2% of all emails have the word "cash" in them
p_cash = 0.02
# We normally would measure this from our data, but we'll take
# it that 10% of all emails we collected were spam
p_spam = 0.10
# 12% of all spam emails have the word "cash"
p_cash_given_its_spam = 0.12
p_spam_given_cash = p_cash_given_its_spam * p_spam / p_cash
print(f'If the email has the word "cash" in it, there is a \
{p_spam_given_cash*100}% chance the email is spam')
```
> **Check it**: Does this make sense? <br>
> Suppose I had 250 total emails.
> - How many should I expect to have the word 'cash' in them? (Ans. 5)
> - How many should I expect to be spam? (Ans. 25)
> - How many *of the spam emails* should I expect to have the word 'cash' in them? (Ans. 3)
## Extending It With Multiple Words
> With more words, the more certain we can be if it is/isn't spam
Spam:
$$ P(🥫\ |"buy",\ "cash") \propto P("buy",\ "cash"|\ 🥫)\ P(🥫)$$
But because of independence:
$$ P("buy",\ "cash"|\ 🥫) = P("buy"|\ 🥫)\ P("cash"|\ 🥫)$$
Normalize by dividing!
$$
P(🥫\ |"buy",\ "cash") =
\frac
{P("buy"|\ 🥫)P("cash"|\ 🥫)\ P(🥫)}
{P("buy"|\ 🥫)P("cash"|\ 🥫)\ P(🥫) + P("buy"|\ 🐷)P("cash"|\ 🐷)\ P(🐷)}
$$
> **Note:** If we wanted to find the most probable class (especially useful for _multiclass_), we find the maximum numerator for the given criteria
# Naive Bayes Modeling Example
## Using Bayes's Theorem for Classification
Let's recall Bayes's Theorem:
$\large P(h|e) = \frac{P(h)P(e|h)}{P(e)}$
### Does this look like a classification problem?
- Suppose we have three competing hypotheses $\{h_1, h_2, h_3\}$ that would explain our evidence $e$.
- Then we could use Bayes's Theorem to calculate the posterior probabilities for each of these three:
- $P(h_1|e) = \frac{P(h_1)P(e|h_1)}{P(e)}$
- $P(h_2|e) = \frac{P(h_2)P(e|h_2)}{P(e)}$
- $P(h_3|e) = \frac{P(h_3)P(e|h_3)}{P(e)}$
- Suppose the evidence is a collection of elephant weights.
- Suppose each of the three hypotheses claims that the elephant whose measurements we have belongs to one of the three extant elephant species (*L. africana*, *L. cyclotis*, and *E. maximus*).
In that case the left-hand sides of these equations represent the probability that the elephant in question belongs to a given species.
If we think of the species as our target, then **this is just an ordinary classification problem**.
What about the right-hand sides of the equations? **These other probabilities we can calculate from our dataset.**
- The priors can simply be taken to be the percentages of the different classes in the dataset.
- What about the likelihoods?
- If the relevant features are **categorical**, we can simply count the numbers of each category in the dataset. For example, if the features are whether the elephant has tusks or not, then, to calculate the likelihoods, we'll just count the tusked and non-tuksed elephants per species.
- If the relevant features are **numerical**, we'll have to do something else. A good way of proceeding is to rely on (presumed) underlying distributions of the data. [Here](https://medium.com/analytics-vidhya/use-naive-bayes-algorithm-for-categorical-and-numerical-data-classification-935d90ab273f) is an example of using the normal distribution to calculate likelihoods. We'll follow this idea below for our elephant data.
## Elephant Example
Suppose we have a dataset that looks like this:
```
elephs = pd.read_csv('data/elephants.csv', usecols=['height (cm)',
'species'])
elephs.head()
plt.style.use('fivethirtyeight')
fig, ax = plt.subplots()
sns.kdeplot(data=elephs[elephs['species'] == 'maximus']['height (cm)'],
ax=ax, label='maximus')
sns.kdeplot(data=elephs[elephs['species'] == 'africana']['height (cm)'],
ax=ax, label='africana')
sns.kdeplot(data=elephs[elephs['species'] == 'cyclotis']['height (cm)'],
ax=ax, label='cyclotis')
plt.legend();
```
### Naive Bayes by Hand
Suppose we want to make prediction of species for some new elephant whose weight we've just recorded. We'll suppose the new elephant has:
```
new_ht = 263
```
What we want to calculate is the mean and standard deviation for height for each elephant species. We'll use these to calculate the relevant likelihoods.
So:
```
max_stats = elephs[elephs['species'] == 'maximus'].describe().loc[['mean', 'std'], :]
max_stats
cyc_stats = elephs[elephs['species'] == 'cyclotis'].describe().loc[['mean', 'std'], :]
cyc_stats
afr_stats = elephs[elephs['species'] == 'africana'].describe().loc[['mean', 'std'], :]
afr_stats
elephs['species'].value_counts()
```
### Calculation of Likelihoods
We'll use the PDFs of the normal distributions with the discovered means and standard deviations to calculate likelihoods:
```
stats.norm(loc=max_stats['height (cm)'][0],
scale=max_stats['height (cm)'][1]).pdf(263)
stats.norm(loc=cyc_stats['height (cm)'][0],
scale=cyc_stats['height (cm)'][1]).pdf(263)
stats.norm(loc=afr_stats['height (cm)'][0],
scale=afr_stats['height (cm)'][1]).pdf(263)
```
### Posteriors
What we have just calculated are (approximations of) the likelihoods, i.e.:
- $P(height=263 | species=maximus) = 2.04\%$
- $P(height=263 | species=cyclotis) = 1.50\%$
- $P(height=263 | species=africana) = 0.90\%$
(Notice that they do NOT sum to 1!) But what we'd really like to know are the posteriors. I.e. what are:
- $P(species=maximus | height=263)$?
- $P(species=cyclotis | height=263)$?
- $P(species=africana | height=263)$?
Since we have equal numbers of each species, every prior is equal to $\frac{1}{3}$. Thus we can calculate the probability of the evidence:
$P(height=263) = \frac{1}{3}(0.0204 + 0.0150 + 0.0090) = 0.0148$
And therefore calculate the posteriors using Bayes's Theorem:
- $P(species=maximus | height=263) = \frac{1}{3}\frac{0.0204}{0.0148} = 45.9\%$;
- $P(species=cyclotis | height=263) = \frac{1}{3}\frac{0.0150}{0.0148} = 33.8\%$;
- $P(species=africana | height=263) = \frac{1}{3}\frac{0.0090}{0.0148} = 20.3\%$.
Bayes's Theorem shows us that the largest posterior belongs to the *maximus* species. (Note also that, since the priors are all the same, the largest posterior will necessarily belong to the species with the largest likelihood!)
Therefore, the *maximus* species will be our prediction for an elephant of this height.
### More Dimensions
In fact, we also have elephant *weight* data available in addition to their heights. To accommodate multiple features we can make use of **multivariate normal** distributions.
#### What's "Naive" about This?
For multiple predictors, we make the simplifying assumption that **our predictors are probablistically independent**. This will often be unrealistic, but it simplifies our calculations a great deal.
```
elephants = pd.read_csv('data/elephants.csv',
usecols=['height (cm)', 'weight (lbs)', 'species'])
elephants.head()
maximus = elephants[elephants['species'] == 'maximus']
cyclotis = elephants[elephants['species'] == 'cyclotis']
africana = elephants[elephants['species'] == 'africana']
```
Suppose our new elephant with a height of 263 cm also has a weight of 7009 lbs.
```
likeli_max = stats.multivariate_normal(mean=maximus.mean(),
cov=maximus.cov()).pdf([263, 7009])
likeli_max
likeli_cyc = stats.multivariate_normal(mean=cyclotis.mean(),
cov=cyclotis.cov()).pdf([263, 7009])
likeli_cyc
likeli_afr = stats.multivariate_normal(mean=africana.mean(),
cov=africana.cov()).pdf([263, 7009])
likeli_afr
```
#### Posteriors
```
post_max = likeli_max / sum([likeli_max, likeli_cyc, likeli_afr])
post_cyc = likeli_cyc / sum([likeli_max, likeli_cyc, likeli_afr])
post_afr = likeli_afr / sum([likeli_max, likeli_cyc, likeli_afr])
print(post_max)
print(post_cyc)
print(post_afr)
```
### [`GaussianNB`](https://scikit-learn.org/stable/modules/generated/sklearn.naive_bayes.GaussianNB.html)
```
gnb = GaussianNB(priors=[1/3, 1/3, 1/3])
X = elephants.drop('species', axis=1)
y = elephants['species']
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
gnb.fit(X_train, y_train)
gnb.predict_proba(np.array([263, 7009]).reshape(1, -1))
gnb.score(X_test, y_test)
plot_confusion_matrix(gnb, X_test, y_test);
```
## Comma Survey Example
```
commas = pd.read_csv('data/comma-survey.csv')
commas.head()
```
The first question on the survey was about the Oxford comma.
```
commas.isna().sum().sum()
```
We'll go ahead and drop the NaNs:
```
commas = commas.dropna()
commas.shape
commas['In your opinion, which sentence is more gramatically correct?'].value_counts()
```
Personally, I like the Oxford comma, since it can help eliminate ambiguities, such as:
"This book is dedicated to my parents, Ayn Rand, and God" <br/> vs. <br/>
"This book is dedicated to my parents, Ayn Rand and God"
Let's see how a Naive Bayes model would make a prediction here. We'll think of the comma preference as our target.
```
commas['Age'].value_counts()
```
Suppose we want to make a prediction about Oxford comma usage for a new person who falls into the **45-60 age group**.
### Calculating Priors and Likelihoods
The following code makes a table of values that count up the number of survey respondents who fall into each of eight bins (the four age groups and the two answers to the first comma question).
```
table = np.zeros((2, 4))
for idx, value in enumerate(commas['Age'].value_counts().index):
table[0, idx] = len(commas[(commas['In your opinion, which sentence is '\
'more gramatically correct?'] ==\
'It\'s important for a person to be '\
'honest, kind, and loyal.') & (commas['Age'] == value)])
table[1, idx] = len(commas[(commas['In your opinion, which sentence is '\
'more gramatically correct?'] ==\
'It\'s important for a person to be '\
'honest, kind and loyal.') & (commas['Age'] == value)])
table
df = pd.DataFrame(table, columns=['Age45-60',
'Age>60',
'Age30-44',
'Age18-29'])
df
df['Oxford'] = [True, False]
df = df[['Age>60', 'Age45-60', 'Age30-44', 'Age18-29', 'Oxford']]
df
```
Since all we have is a single categorical feature here we can just read our likelihoods and priors right off of this table:
Likelihoods:
- Age45-60:
- P(Age45-60 | Oxford=True) = $\frac{123}{470} = 0.2617$;
- P(Age45-60 | Oxford=False) = $\frac{125}{355} = 0.3521$.
Priors:
- P(Oxford=True) = $\frac{470}{825} = 0.5697$;
- P(Oxford=False) = $\frac{355}{825} = 0.4303$.
### Calculating Posteriors
First we'll calculate the probability of the evidence:
$$\begin{align}
P(Age45-60) &= P(Age45-60 | Oxford=True) \cdot P(Oxford=True) \\
& \hspace{1cm} + P(Age45-60 | Oxford=False) \cdot P(Oxford=False)\\
&= 0.2617 \cdot 0.5697 + 0.3521 \cdot 0.4303 \\
&= 0.3006
\end{align}$$
```
(123+125)/825
```
Now use Bayes's Theorem to calculate the posteriors:
$$\begin{align}
P(Oxford=True | Age45-60) &= P(Oxford=True) \cdot P(Age45-60 | Oxford=True) / P(Age45-60) \\
&= 0.5697 \cdot 0.2617 / 0.3006 \\
&= 0.4960 \\
\\
P(Oxford=False | Age45-60) &= P(Oxford=False) \cdot P(Age45-60 | Oxford=False) / P(Age45-60) \\
&= 0.4303 \cdot 0.3521 / 0.3006 \\
&= 0.5040
\end{align}$$
Close! But our prediction for someone in the 45-60 age group will be that they **do not** favor the Oxford comma.
### Comparison with [`MultinomialNB`](https://scikit-learn.org/stable/modules/generated/sklearn.naive_bayes.MultinomialNB.html)
```
comma_model = MultinomialNB()
ohe = OneHotEncoder()
ohe.fit(commas['Age'].values.reshape(-1, 1))
X = ohe.transform(commas['Age'].values.reshape(-1, 1)).todense()
y = commas['In your opinion, which sentence is more gramatically correct?']
comma_model.fit(X, y)
comma_model.predict_proba(np.array([0, 0, 1, 0]).reshape(1, -1))
```
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#@title MIT License
#
# Copyright (c) 2017 François Chollet
#
# Permission is hereby granted, free of charge, to any person obtaining a
# copy of this software and associated documentation files (the "Software"),
# to deal in the Software without restriction, including without limitation
# the rights to use, copy, modify, merge, publish, distribute, sublicense,
# and/or sell copies of the Software, and to permit persons to whom the
# Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in
# all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL
# THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING
# FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER
# DEALINGS IN THE SOFTWARE.
```
# Регрессия: Спрогнозируй эффективность расхода топлива
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/tutorials/keras/regression"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />Смотрите на TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/ru/tutorials/keras/regression.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Запустите в Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/ru/tutorials/keras/regression.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />Изучайте код на GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/ru/tutorials/keras/regression.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Скачайте ноутбук</a>
</td>
</table>
Note: Вся информация в этом разделе переведена с помощью русскоговорящего Tensorflow сообщества на общественных началах. Поскольку этот перевод не является официальным, мы не гарантируем что он на 100% аккуратен и соответствует [официальной документации на английском языке](https://www.tensorflow.org/?hl=en). Если у вас есть предложение как исправить этот перевод, мы будем очень рады увидеть pull request в [tensorflow/docs](https://github.com/tensorflow/docs) репозиторий GitHub. Если вы хотите помочь сделать документацию по Tensorflow лучше (сделать сам перевод или проверить перевод подготовленный кем-то другим), напишите нам на [docs-ru@tensorflow.org list](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs-ru).
В задаче *регрессии*, мы хотим дать прогноз какого-либо непрерывного значения, например цену или вероятность. Сравните это с задачей *классификации*, где нужно выбрать конкретную категорию из ограниченного списка (например, есть ли на картинке яблоко или апельсин, распознайте какой фрукт на изображении).
Этот урок использует классический датасет [Auto MPG](https://archive.ics.uci.edu/ml/datasets/auto+mpg) и строит модель, предсказывающую эффективность расхода топлива автомобилей конца 70-х и начала 80-х. Чтобы сделать это, мы предоставим модели описания множества различных автомобилей того времени. Эти описания будут содержать такие параметры как количество цилиндров, лошадиных сил, объем двигателя и вес.
В этом примере используется tf.keras API, подробнее [смотри здесь](https://www.tensorflow.org/guide/keras).
```
# Установим библиотеку seaborn для построения парных графиков
!pip install seaborn
import pathlib
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
print(tf.__version__)
```
## Датасет Auto MPG
Датасет доступен в [репозитории машинного обучения UCI](https://archive.ics.uci.edu/ml/).
### Получите данные
Сперва загрузим датасет.
```
dataset_path = keras.utils.get_file("auto-mpg.data", "http://archive.ics.uci.edu/ml/machine-learning-databases/auto-mpg/auto-mpg.data")
dataset_path
```
Импортируем его при помощи библиотеки Pandas:
```
column_names = ['MPG','Cylinders','Displacement','Horsepower','Weight',
'Acceleration', 'Model Year', 'Origin']
raw_dataset = pd.read_csv(dataset_path, names=column_names,
na_values = "?", comment='\t',
sep=" ", skipinitialspace=True)
dataset = raw_dataset.copy()
dataset.tail()
```
### Подготовьте данные
Датасет содержит несколько неизвестных значений.
```
dataset.isna().sum()
```
Чтобы урок оставался простым, удалим эти строки.
```
dataset = dataset.dropna()
```
Столбец `"Origin"` на самом деле категорийный, а не числовой. Поэтому конвертируем его в one-hot
```
origin = dataset.pop('Origin')
dataset['USA'] = (origin == 1)*1.0
dataset['Europe'] = (origin == 2)*1.0
dataset['Japan'] = (origin == 3)*1.0
dataset.tail()
```
### Разделите данные на обучающую и тестовую выборки
Сейчас разделим датасет на обучающую и тестовую выборки.
Тестовую выборку будем использовать для итоговой оценки нашей модели
```
train_dataset = dataset.sample(frac=0.8,random_state=0)
test_dataset = dataset.drop(train_dataset.index)
```
### Проверьте данные
Посмотрите на совместное распределение нескольких пар колонок из тренировочного набора данных:
```
sns.pairplot(train_dataset[["MPG", "Cylinders", "Displacement", "Weight"]], diag_kind="kde")
```
Также посмотрите на общую статистику:
```
train_stats = train_dataset.describe()
train_stats.pop("MPG")
train_stats = train_stats.transpose()
train_stats
```
### Отделите признаки от меток
Отделите целевые значения или "метки" от признаков. Обучите модель для предсказания значений.
```
train_labels = train_dataset.pop('MPG')
test_labels = test_dataset.pop('MPG')
```
### Нормализуйте данные
Взгляните еще раз на блок train_stats приведенный выше. Обратите внимание на то, как отличаются диапазоны каждого из признаков.
Это хорошая практика - нормализовать признаки у которых различные масштабы и диапазон изменений. Хотя модель *может* сходиться и без нормализации признаков, обучение при этом усложняется и итоговая модель становится зависимой от выбранных единиц измерения входных данных..
Примечание. Мы намеренно генерируем эти статистические данные только из обучающей выборки, они же будут использоваться для нормализации тестовой выборки. Мы должны сделать это, чтобы тестовая выборка была из того распределения на которой обучалась модель.
```
def norm(x):
return (x - train_stats['mean']) / train_stats['std']
normed_train_data = norm(train_dataset)
normed_test_data = norm(test_dataset)
```
Для обучения модели мы будем использовать эти нормализованные данные.
Внимание: статистики использованные для нормализации входных данных (среднее и стандартное отклонение) должны быть применены к любым другим данным, которые используются в модели. Это же касается one-hot кодирования которое мы делали ранее. Преобразования необходимо применять как к тестовым данным, так и к данным с которыми модель используется в работе.
## Модель
### Постройте модель
Давайте построим нашу модель. Мы будем использовать `Sequential` (последовательную) модель с двумя полносвязными скрытыми слоями, а выходной слой будет возвращать одно непрерывное значение. Этапы построения модели мы опишем в функции build_model, так как позже мы создадим еще одну модель.
```
def build_model():
model = keras.Sequential([
layers.Dense(64, activation='relu', input_shape=[len(train_dataset.keys())]),
layers.Dense(64, activation='relu'),
layers.Dense(1)
])
optimizer = tf.keras.optimizers.RMSprop(0.001)
model.compile(loss='mse',
optimizer=optimizer,
metrics=['mae', 'mse'])
return model
model = build_model()
```
### Проверьте модель
Используйте метод `.summary` чтобы напечатать простое описание модели.
```
model.summary()
```
Сейчас попробуем нашу модель. Возьмем пакет из`10` примеров из обучающей выборки и вызовем `model.predict` на них.
```
example_batch = normed_train_data[:10]
example_result = model.predict(example_batch)
example_result
```
Похоже все работает правильно, модель показывает результат ожидаемой размерности и типа.
### Обучите модель
Обучите модель за 1000 эпох и запишите точность модели на тренировочных и проверочных данных в объекте `history`.
```
# Выведем прогресс обучения в виде точек после каждой завершенной эпохи
class PrintDot(keras.callbacks.Callback):
def on_epoch_end(self, epoch, logs):
if epoch % 100 == 0: print('')
print('.', end='')
EPOCHS = 1000
history = model.fit(
normed_train_data, train_labels,
epochs=EPOCHS, validation_split = 0.2, verbose=0,
callbacks=[PrintDot()])
```
Визуализируйте процесс обучения модели используя статистику содержащуюся в объекте `history`.
```
hist = pd.DataFrame(history.history)
hist['epoch'] = history.epoch
hist.tail()
def plot_history(history):
hist = pd.DataFrame(history.history)
hist['epoch'] = history.epoch
plt.figure()
plt.xlabel('Epoch')
plt.ylabel('Mean Abs Error [MPG]')
plt.plot(hist['epoch'], hist['mae'],
label='Train Error')
plt.plot(hist['epoch'], hist['val_mae'],
label = 'Val Error')
plt.ylim([0,5])
plt.legend()
plt.figure()
plt.xlabel('Epoch')
plt.ylabel('Mean Square Error [$MPG^2$]')
plt.plot(hist['epoch'], hist['mse'],
label='Train Error')
plt.plot(hist['epoch'], hist['val_mse'],
label = 'Val Error')
plt.ylim([0,20])
plt.legend()
plt.show()
plot_history(history)
```
Полученный график показывает, небольшое улучшение или даже деградацию ошибки валидации после примерно 100 эпох обучения. Давай обновим метод model.fit чтобы автоматически прекращать обучение когда ошибка валидации Val loss прекращает улучшаться. Для этого используем функцию *EarlyStopping callback* которая проверяет показатели обучения после каждой эпохи. Если после определенного количество эпох нет никаких улучшений, то функция автоматически остановит обучение.
Вы можете больше узнать про этот коллбек [здесь](https://www.tensorflow.org/versions/master/api_docs/python/tf/keras/callbacks/EarlyStopping).
```
model = build_model()
# Параметр patience определяет количество эпох, проверяемых на улучшение
early_stop = keras.callbacks.EarlyStopping(monitor='val_loss', patience=10)
history = model.fit(normed_train_data, train_labels, epochs=EPOCHS,
validation_split = 0.2, verbose=0, callbacks=[early_stop, PrintDot()])
plot_history(history)
```
График показывает что среднее значение ошибки на проверочных данных - около 2 галлонов на милю. Хорошо это или плохо? Решать тебе.
Давай посмотрим как наша модель справится на **тестовой** выборке, которую мы еще не использовали при обучении модели. Эта проверка покажет нам какого результата ожидать от модели, когда мы будем ее использовать в реальном мире
```
loss, mae, mse = model.evaluate(normed_test_data, test_labels, verbose=2)
print("Testing set Mean Abs Error: {:5.2f} MPG".format(mae))
```
### Сделайте прогноз
Наконец, спрогнозируйте значения миль-на-галлон используя данные из тестовой выборки:
```
test_predictions = model.predict(normed_test_data).flatten()
plt.scatter(test_labels, test_predictions)
plt.xlabel('True Values [MPG]')
plt.ylabel('Predictions [MPG]')
plt.axis('equal')
plt.axis('square')
plt.xlim([0,plt.xlim()[1]])
plt.ylim([0,plt.ylim()[1]])
_ = plt.plot([-100, 100], [-100, 100])
```
Вроде наша модель дает хорошие предсказания. Давайте посмотрим распределение ошибки.
```
error = test_predictions - test_labels
plt.hist(error, bins = 25)
plt.xlabel("Prediction Error [MPG]")
_ = plt.ylabel("Count")
```
Она не достаточно гауссова, но мы могли это предполагать потому что количество примеров очень мало.
## Заключение
Это руководство познакомило тебя с несколькими способами решения задач регрессии.
* Среднеквадратичная ошибка (MSE) это распространенная функция потерь используемая для задач регрессии (для классификации используются другие функции).
* Аналогично, показатели оценки модели для регрессии отличаются от используемых в классификации. Обычной метрикой для регрессии является средняя абсолютная ошибка (MAE).
* Когда значения числовых входных данных из разных диапазонов, каждый признак должен быть незавизимо масштабирован до одного и того же диапазона.
* Если данных для обучения немного, используй небольшую сеть из нескольких скрытых слоев. Это поможет избежать переобучения.
* Метод ранней остановки очень полезная техника для избежания переобучения.
| github_jupyter |
```
import re
import pandas as pd
import numpy as np
import pickle
import nltk
from sklearn.preprocessing import LabelEncoder
from nltk.corpus import stopwords
from nltk.tokenize import word_tokenize
from nltk.stem.porter import PorterStemmer
from sklearn.feature_extraction.text import TfidfVectorizer
```
## Forming the dataframe
```
df = pd.read_json('../src/utility/final.json')
df = pd.DataFrame(df.data.values.tolist())
df.head()
df = df.drop(['id', 'withheld'], axis=1)
df.head()
lb = LabelEncoder()
label = lb.fit_transform(df["label"])
lb.classes_
df.head(20)
df = df.drop(["label"], axis='columns')
df.head()
df["label"]=label
df.head()
```
## Preprocessing the data
```
def cleanTweet(text):
text = re.sub(r'@[A-Za-z0-9]+','',text) # Removing @mentions
text = re.sub(r'#','',text) # Removing the '#' symbol
text = re.sub(r'RT[\s]+','',text) # Removing RT
text = re.sub(r'https?:\/\/\S+','',text) # Removing hyperlinks
text = re.sub(r'[^a-zA-Z ]',' ', text) # Removing all the punctuations and numbers
text = text.lower()
return text
df['text'] = df['text'].apply(cleanTweet)
df.head()
# Removing the stop words and tokeinzing the sentences
stop_words = set(stopwords.words('english'))
def removeStopWords(text):
words = word_tokenize(text)
filtered_sentence = [w for w in words if not w in stop_words]
return filtered_sentence
tokenized_tweet = df['text'].apply(removeStopWords)
tokenized_tweet.head()
# Stemming
stemmer = PorterStemmer()
def stemTweet(text):
text = [stemmer.stem(word) for word in text]
return text
tokenized_tweet = tokenized_tweet.apply(stemTweet)
tokenized_tweet.head()
tweet=[]
for i in range(len(tokenized_tweet)):
s = tokenized_tweet[i]
sent = ' '.join([str(elem) for elem in s])
tweet.append(sent)
df['clean_tweet'] = tweet
df['clean_tweet'][1]
```
## Feature Extraction
```
vectorizer = TfidfVectorizer()
vector = vectorizer.fit_transform(df['clean_tweet'])
print(type(df['clean_tweet']))
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(vector, df['label'], random_state=42, test_size=0.25)
```
## Model Training
```
# from sklearn.ensemble import RandomForestClassifier
# model = RandomForestClassifier(bootstrap=True)
# model.fit(x_train, y_train)
# # model.score(x_test, y_test)
# from sklearn.metrics import f1_score, accuracy_score
# pred = model.predict(x_test)
# f1_score(y_test, pred, average=None)
# accuracy_score(y_test, pred)
from sklearn.linear_model import LogisticRegression
model = LogisticRegression(penalty = 'elasticnet', warm_start = True, max_iter = 1000, C=1.3, solver='saga', l1_ratio=0.9)
model.fit(x_train, y_train)
from sklearn.metrics import f1_score, accuracy_score
pred = model.predict(x_test)
f1_score(y_test, pred, average=None)
accuracy_score(y_test, pred)
```
## Exporting the trained model
```
fname = 'logistic_regression_model.sav'
pickle.dump(model, open(fname, 'wb'))
def cleanInput(text):
text = cleanTweet(text)
text = removeStopWords(text)
text = stemTweet(text)
text = ' '.join([str(elem) for elem in text])
text = [text]
return text
inp = 'I am a morning person'
inp = cleanInput(inp)
data = vectorizer.transform(inp)
data
model.predict(data)
```
| github_jupyter |
<center><h2>Python Script that charts bitcoin prices in US dollars</h2></center>
```
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib.dates import YearLocator, MonthLocator, WeekdayLocator, DayLocator, DateFormatter
import dateutil
from matplotlib.ticker import MultipleLocator
from datetime import datetime
from datetime import date
# Data metadata:
# CSV file where first column is string date in dd/mm/YYYY H:M:S format and second column is the bitcoin price in US dollars
# Pandas.read_csv has an awesome converters parameter that allows us to convert or modify a column's data on the fly!
# Therefore, we can read the first column as an actual Python datetime object instead of as a string.
# Because there is time info with the date, it causes the x-axis ticks to not correspond with the data.
# As a workaround, I stripped the time from the date values by invoking datetime.date()
bitcoin_data = pd.read_csv("http://blockchain.info/charts/market-price?showDataPoints=false×pan=&show_header=true&daysAverageString=1&scale=0&format=csv&address=",
index_col=0, names=['price'], converters={0: lambda x: datetime.date(datetime.strptime(x, '%d/%m/%Y %H:%M:%S'))})
```
#### Let's view just the last 5 rows of the bitcoin data:
```
bitcoin_data.tail()
```
#### Now, let's plot the data. Top chart will be all data available in monthly x-ticks intervals, bottom chart will be just last 2 calendar months of data, with daily x-ticks intervals.
```
fig, axes = plt.subplots(2, 1) # Make 2 rows and 1 column of plots (2 plots total)
# These are just time intervals we can use to format our X axis tick intervals
year = YearLocator()
month = MonthLocator(bymonth=range(1,13), bymonthday=1, interval=1)
week = WeekdayLocator(byweekday=dateutil.rrule.MO) # Every MOnday or WeekdayLocator(byweekday=0) works too
day = DayLocator(bymonthday=range(1,32), interval=1)
####################### Let's make the top plot ######################
# Use trial and error to get the size you want
axes[0] = fig.add_axes([0, 1.4, 1.5, 1]) # left, bottom, width, height
# Format the x-axis, set tick labels as Month 'Yr format, turn on grid for major ticks
monthsFmt = DateFormatter("%b '%y")
axes[0].xaxis.set_major_formatter(monthsFmt)
axes[0].xaxis.set_major_locator(month) # Make the x-axis tick interval to be monthly interval size
axes[0].xaxis.grid(which='major')
# Format the y-axis
y_major_ticks = MultipleLocator(200)
axes[0].yaxis.set_major_locator(y_major_ticks)
axes[0].yaxis.set_ticks_position('right') # Not sure why 'both' don't work
axes[0].yaxis.set_label_position('right')
axes[0].yaxis.grid(which='major')
# Plot the data...
axes[0].plot_date(bitcoin_data.index, bitcoin_data.values, 'r')
axes[0].set_title("Bitcoin Value in US$", weight="bold")
axes[0].set_xlabel("Month - Year")
axes[0].set_ylabel("US$")
plt.setp(axes[0].get_xticklabels(), rotation=-90)
############# Let's get last 2 calendar months' worth of bitcoin prices ###########
# But first, we must account for situation where current month is January
today = datetime.today()
this_month = today.month
if this_month==1:
last_year = today.year - 1 # if month is January, then year will be one less than current year
last_month = 12 # if month is January, then previous month is Dec (12)
else:
last_year = today.year # Otherwise, last year is really same as current year and
last_month = this_month - 1 # last month will equal current month minus 1
this_year = today.year
# Pandas' filtering by date ranges only work if the dates are of type datetime.datetime
# Unfortunately, the data parsed from the CSV file, was parsed as datetime.date
# So we will convert bitcoin_data.index to datetime.datetime:
bitcoin_data.index = bitcoin_data.index.to_datetime()
# Now I will take advantage of Pandas partial string indexing described below to get just 2 months of data:
# http://pandas.pydata.org/pandas-docs/stable/timeseries.html#partial-string-indexing
last2months = bitcoin_data[str(last_month)+'-'+str(last_year):str(this_month)+'-'+str(this_year)]
# Get current price and also the current date, convert it to a datetime.date, then make a string from it in mm/dd/YYYY format
current_price = last2months[-1:].values[0][0]
last_date = last2months[-1:].index
current_date_unformatted = date(last_date.year[0], last_date.month[0], last_date.day[0])
current_date = datetime.strftime(current_date_unformatted, "%m/%d/%Y")
####################### Let's make the bottom plot ######################
axes[1] = fig.add_axes([0, 0, 1.5, 1]) # left, bottom, width, height
# Format the x-axis tick labels as mm/dd/YYYY format, turn on grid for major ticks
dayFmt = DateFormatter("%b-%d-%Y")
axes[1].xaxis.set_major_locator(day) # Make x-axis tick interval to be daily interval size
axes[1].xaxis.set_major_formatter(dayFmt)
axes[1].xaxis.grid(which='major')
# Format the y-axis
y_major_ticks = MultipleLocator(20)
axes[1].yaxis.set_major_locator(y_major_ticks)
axes[1].yaxis.set_ticks_position('right') # Not sure why 'both' don't work
axes[1].yaxis.set_label_position('right')
axes[1].yaxis.grid(which='major')
# Plot the data...
axes[1].plot_date(last2months.index, last2months.values, 'r')
axes[1].set_title("Current Value@"+current_date+": "+str(current_price), weight="bold", size=10)
# MATPLOTLIB does not have a good way of creating a title with 2 rows of text with different font sizes
# Below is a workaround(http://matplotlib.org/users/text_intro.html), sort of "hackish" and requires trial and error (x,y)
axes[1].text(.22,1.07, "Last 2 Calendar Months of Bitcoin Values in US$", transform=axes[1].transAxes,
weight="bold", size=14)
axes[1].set_xlabel("Month - Day - Year")
axes[1].set_ylabel("US$")
plt.setp(axes[1].get_xticklabels(), rotation=-90, size=8)
plt.setp(axes[1].get_yticklabels(), size=9)
plt.show()
```
| github_jupyter |
```
%pylab inline
import csv
import os
import sklearn
from sklearn.metrics import mean_squared_error
import nltk
LEV1_GT_FILE = 'ADMIN_metadata.csv'
LEV2_GT_FILE = '2ADMIN_metadata.csv'
LEV3_GT_FILE = '3ADMIN_metadata.csv'
def read_labels(filename):
labels = {}
with open(filename, 'r') as f:
csvr = csv.reader(f, delimiter=',', quotechar='"')
for r in csvr:
if r[-1] == 'label' or r[-1] == 'pred_label' or r[-1] == 'predict':
# ignore header
continue
labels[r[0]] = r
return labels
def compare(gt_value, pred_value):
is_array = False
if gt_value.startswith('['):
# parse array
is_array = True
gt_value = [float(v) for v in gt_value[1:-1].split(',')]
else:
gt_value = [float(gt_value)]
if pred_value.startswith('['):
if not is_array:
# penalize
return 1000000
# parse array
try:
pred_value = [float(v) for v in pred_value[1:-1].split(',')]
except:
pred_value = []
if len(gt_value) != len(pred_value):
return 1000000
else:
if is_array:
# penalize
return 1000000
pred_value = [float(pred_value)]
# try:
rmse = mean_squared_error(gt_value, pred_value, squared=False)
# except:
# print(gt_value, pred_value)
return rmse
def is_float(value):
try:
float(value)
return True
except:
return False
def compare3(gt_value, pred_value):
error = 0
if pred_value == '':
return 1000000
if gt_value == 'Yes' or gt_value == 'No':
if gt_value != pred_value:
error += 1
elif is_float(gt_value):
error += mean_squared_error([float(gt_value)], [float(pred_value)], squared=False)
else:
# Levenstein edit distance
error += nltk.edit_distance(gt_value, pred_value)
return error
def compare_many(gt_values, pred_values, lev3=False):
errors = []
for i,v in enumerate(gt_values.keys()):
if not v in pred_values:
error = 1000000
else:
if lev3:
error = compare3(gt_values[v][-1], pred_values[v][-1])
else:
error = compare(gt_values[v][-1], pred_values[v][-1])
errors.append(error)
return np.mean(errors)
lev1_gt = read_labels(LEV1_GT_FILE)
lev2_gt = read_labels(LEV2_GT_FILE)
lev3_gt = read_labels(LEV3_GT_FILE)
LEV1_SUB_FILE = 'lev1_TEST_metadata.csv'
LEV2_SUB_FILE = 'lev2_TEST_metadata.csv'
LEV3_SUB_FILE = 'lev3_TEST_metadata.csv'
SUBMISSIONS_DIR = '/home/d/Projects/cqaw_submission/SUBMISSIONS/'
submissions = os.listdir(SUBMISSIONS_DIR)
for s in submissions:
print('-'*80)
print('SUBMISSION BY', s)
lev1_sub_file = os.path.join(SUBMISSIONS_DIR, s, LEV1_SUB_FILE)
lev2_sub_file = os.path.join(SUBMISSIONS_DIR, s, LEV2_SUB_FILE)
lev3_sub_file = os.path.join(SUBMISSIONS_DIR, s, LEV3_SUB_FILE)
if os.path.exists(lev1_sub_file):
lev1_sub_labels = read_labels(lev1_sub_file)
mean_rmse = compare_many(lev1_gt, lev1_sub_labels)
print('LEVEL 1 mean RMSE:', mean_rmse)
if os.path.exists(lev2_sub_file):
lev2_sub_labels = read_labels(lev2_sub_file)
mean_rmse = compare_many(lev2_gt, lev2_sub_labels)
print('LEVEL 2 mean RMSE:', mean_rmse)
if os.path.exists(lev3_sub_file):
lev3_sub_labels = read_labels(lev3_sub_file)
mean_err = compare_many(lev3_gt, lev3_sub_labels, lev3=True)
print('LEVEL 3 mean ERROR:', mean_err)
```
| github_jupyter |
# SQL Local Queries
###
```
import numpy as np
import pandas as pd
import wget
import psycopg2
import pymongo
import sqlite3
from sqlalchemy import create_engine
from bson.json_util import dumps, loads
import requests
import json
import os
import sys
import dotenv
# sql passwords file
os.chdir("C:\\Users\\dwagn\\git\\ds-6001\\mod7")
dotenv.load_dotenv()
sys.tracebacklimit = 0 # error tracebacks
# 4 separate tables in db
works = pd.read_csv("https://github.com/jkropko/DS-6001/raw/master/localdata/Works.csv")
characters = pd.read_csv("https://github.com/jkropko/DS-6001/raw/master/localdata/Characters.csv")
chapters = pd.read_csv("https://github.com/jkropko/DS-6001/raw/master/localdata/Chapters.csv")
paragraphs = pd.read_csv("https://github.com/jkropko/DS-6001/raw/master/localdata/Paragraphs.csv")
works.columns = works.columns.str.lower()
characters.columns = characters.columns.str.lower()
chapters.columns = chapters.columns.str.lower()
paragraphs.columns = paragraphs.columns.str.lower()
```
## PostgreSQL
```
# PostgreSQL
dotenv.load_dotenv()
pgpassword = os.getenv("pgpass")
dbserver = psycopg2.connect(
user='postgres',
password=pgpassword,
host="localhost"
)
dbserver.autocommit = True
cursor = dbserver.cursor()
try:
cursor.execute("CREATE DATABASE shake")
except:
cursor.execute("DROP DATABASE shake")
cursor.execute("CREATE DATABASE shake")
engine = create_engine("postgresql+psycopg2://{user}:{pw}@localhost/{db}"
.format(user='postgres',
pw=pgpassword,
db="teams"))
works.to_sql('works', con = engine, index=False, chunksize=1000, if_exists = 'replace')
characters.to_sql('characters', con = engine, index=False, chunksize=1000, if_exists = 'replace')
chapters.to_sql('chapters', con = engine, index=False, chunksize=1000, if_exists = 'replace')
paragraphs.to_sql('paragraphs', con = engine, index=False, chunksize=1000, if_exists = 'replace')
'''
Takes title, date, and totalwords from works table, sorts by totalwords in descending order,
creates new column (era) which classifies time periods from 1607,
then finally displays rows 7 through 11:
'''
myquery = """
SELECT title, date as year, totalwords,
CASE WHEN date < 1600 THEN 'early'
WHEN date BETWEEN 1600 AND 1607 THEN 'middle'
WHEN date > 1607 THEN 'late' ELSE NULL END AS era FROM works ORDER BY totalwords DESC LIMIT 5 OFFSET 6;
"""
pd.read_sql_query(myquery, con=engine)
'''
Reports average number of words in works by genre type, then displays genre type and
average wordcount within genre, then sorts by average in descending order
'''
myquery = """
SELECT genretype, AVG(totalwords) as avg_words
FROM works
GROUP BY genretype
ORDER BY avg_words DESC;
"""
pd.read_sql_query(myquery, con=engine)
'''
Generates table containig text of the longest speech in Hamlet
'''
myquery = """
SELECT plaintext, charid, wordcount
FROM paragraphs
WHERE charid = (SELECT charid
FROM characters
WHERE charname = 'Hamlet')
AND wordcount IS NOT null
ORDER BY wordcount DESC
LIMIT 1;
"""
pd.read_sql_query(myquery, con=engine)
'''
Lists all chapters that are set in a palace or castle
'''
myquery = """
SELECT works.title, works.workid, chapters.section AS act, chapters.chapter AS scene, chapters.description
FROM works, chapters
WHERE lower(chapters.description) LIKE '%%palace%%'
OR lower(chapters.description) LIKE '%%castle%%'
"""
pd.read_sql_query(myquery, con=engine)
'''
Example of joins to list columns from different tables, sorting by speech length and
restricting to those who have given at least 20 speeches
'''
myquery = """
SELECT
c.charname as character,
c.description as description,
w.title as work_title,
c.speechcount as speech_count,
AVG(p.wordcount) as avg_speech_length
FROM characters c
LEFT JOIN paragraphs p ON c.charid = p.charid
INNER JOIN works w ON c.works = w.workid
WHERE c.speechcount >= 20
GROUP BY c.charname, c.description, work_title, c.speechcount
ORDER BY avg_speech_length
"""
pd.read_sql_query(myquery, con=engine)
'''
Works that have no scenes in a castle or palace
'''
myquery = """
SELECT
DISTINCT w.title,
w.genretype,
w.date
FROM works w
INNER JOIN chapters c ON c.workid = w.workid
WHERE lower(c.description) NOT LIKE '%%palace%%'
AND lower(c.description) NOT LIKE '%%castle%%'
"""
pd.read_sql_query(myquery, con=engine).head() # cutting off at 5
```
## MongoDB
```
const = requests.get("https://github.com/jkropko/DS-6001/raw/master/localdata/const.json")
const_json = json.loads(const.text)
pd.DataFrame.from_records(const_json).head()
myclient = pymongo.MongoClient("mongodb://localhost/")
constdb = myclient["const"]
collist = constdb.list_collection_names()
if "constcollection" in collist:
constdb.constcollection.drop()
constcollection = constdb['constcollection']
constcollection.delete_many({}) # remove existing data from collection
allconst = constcollection.insert_many(const_json)
constcollection.count_documents({})
'''
Adopted after 1990 or less than 50% democracy score
'''
myquery = constcollection.find({'$or' :
[{'adopted' : {'$gt':1990},
'democracy' : {'$lt':0.5}}]
},
{'country':1,
'adopted':1,
'democracy':1,
'_id':0})
const_text = dumps(myquery)
const_records = loads(const_text)
const_df = pd.DataFrame.from_records(const_records)
const_df.head()
'''
Updating record's democracy score
'''
constcollection.update_one({'country' : 'Hungary'},
{'$set' : {'democracy': 0.4}})
myquery = constcollection.find({'country':'Hungary'})
const_text = dumps(myquery)
const_records = loads(const_text)
const_df = pd.DataFrame.from_records(const_records)
const_df.head()
'''
Searching for texts that contain specific phrase
'''
def read_mongo_query(col, query):
query_text = dumps(col.find(query))
query_rec = loads(query_text)
query_df = pd.DataFrame.from_records(query_rec)
return query_df
constcollection.create_index([('text', 'text')])
df = mongo_read_query(constcollection,
{'$text': {'$search':'\"freedom of speech\"',
'$caseSensitive': False}}) \
[['country', 'adopted', 'democracy']]
df.head()
'''
Searching for any of multiple terms
'''
cursor = constcollection.find(
{'$text': {'$search': 'freedom liberty legal justice rights',
'$caseSensitive': False}},
{'score': {'$meta': 'textScore'}})
cursor.sort([('score', {'$meta': 'textScore'})])
qtext = dumps(cursor)
qrec = loads(qtext)
df = pd.DataFrame.from_records(qrec)
df.head()
# end connections
dbserver.close()
myclient.close()
## Reads in files
# url = "https://github.com/nolanbconaway/pitchfork-data/raw/master/pitchfork.db"
# pfork = wget.download(url)
# pitchfork = sqlite3.connect(pfork)
# for t in ['artists','content','genres','labels','reviews','years']:
# datatable = pd.read_sql_query("SELECT * FROM {tab}".format(tab=t), pitchfork)
# datatable.to_csv("{tab}.csv".format(tab=t))
reviews = pd.read_csv("reviews.csv")
artists = pd.read_csv("artists.csv")
content = pd.read_csv("content.csv")
genres = pd.read_csv("genres.csv")
labels = pd.read_csv("labels.csv")
years = pd.read_csv("years.csv")
music_db = sqlite3.connect("music.db")
reviews.to_sql('reviews', music_db, index=False, chunksize=1000, if_exists='replace')
artists.to_sql('artists', music_db, index=False, chunksize=1000, if_exists='replace')
content.to_sql('content', music_db, index=False, chunksize=1000, if_exists='replace')
genres.to_sql('genres', music_db, index=False, chunksize=1000, if_exists='replace')
labels.to_sql('labels', music_db, index=False, chunksize=1000, if_exists='replace')
years.to_sql('years', music_db, index=False, chunksize=1000, if_exists='replace')
```
| github_jupyter |
<a href="https://colab.research.google.com/github/DingLi23/s2search/blob/pipelining/pipelining/exp-cscr/exp-cscr_cscr_shapley_value.ipynb" target="_blank"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
### Experiment Description
> This notebook is for experiment \<exp-cscr\> and data sample \<cscr\>.
### Initialization
```
%reload_ext autoreload
%autoreload 2
import numpy as np, sys, os
sys.path.insert(1, '../../')
from shapley_value import compute_shapley_value, feature_key_list
sv = compute_shapley_value('exp-cscr', 'cscr')
```
### Plotting
```
import matplotlib.pyplot as plt
import numpy as np
from s2search_score_pdp import pdp_based_importance
fig, axs = plt.subplots(nrows=1, ncols=2, figsize=(12, 5), dpi=200)
# generate some random test data
all_data = []
average_sv = []
sv_global_imp = []
for player_sv in [f'{player}_sv' for player in feature_key_list]:
all_data.append(sv[player_sv])
average_sv.append(pdp_based_importance(sv[player_sv]))
sv_global_imp.append(np.mean(np.abs(list(sv[player_sv]))))
# average_sv.append(np.std(sv[player_sv]))
# print(np.max(sv[player_sv]))
# plot violin plot
axs[0].violinplot(all_data,
showmeans=False,
showmedians=True)
axs[0].set_title('Violin plot')
# plot box plot
axs[1].boxplot(all_data,
showfliers=False,
showmeans=True,
)
axs[1].set_title('Box plot')
# adding horizontal grid lines
for ax in axs:
ax.yaxis.grid(True)
ax.set_xticks([y + 1 for y in range(len(all_data))],
labels=['title', 'abstract', 'venue', 'authors', 'year', 'n_citations'])
ax.set_xlabel('Features')
ax.set_ylabel('Shapley Value')
plt.show()
plt.rcdefaults()
fig, ax = plt.subplots(figsize=(12, 4), dpi=200)
# Example data
feature_names = ('title', 'abstract', 'venue', 'authors', 'year', 'n_citations')
y_pos = np.arange(len(feature_names))
# error = np.random.rand(len(feature_names))
# ax.xaxis.grid(True)
ax.barh(y_pos, average_sv, align='center', color='#008bfb')
ax.set_yticks(y_pos, labels=feature_names)
ax.invert_yaxis() # labels read top-to-bottom
ax.set_xlabel('PDP-based Feature Importance on Shapley Value')
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
_, xmax = plt.xlim()
plt.xlim(0, xmax + 1)
for i, v in enumerate(average_sv):
margin = 0.05
ax.text(v + margin if v > 0 else margin, i, str(round(v, 4)), color='black', ha='left', va='center')
plt.show()
plt.rcdefaults()
fig, ax = plt.subplots(figsize=(12, 4), dpi=200)
# Example data
feature_names = ('title', 'abstract', 'venue', 'authors', 'year', 'n_citations')
y_pos = np.arange(len(feature_names))
# error = np.random.rand(len(feature_names))
# ax.xaxis.grid(True)
ax.barh(y_pos, sv_global_imp, align='center', color='#008bfb')
ax.set_yticks(y_pos, labels=feature_names)
ax.invert_yaxis() # labels read top-to-bottom
ax.set_xlabel('SHAP Feature Importance')
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
_, xmax = plt.xlim()
plt.xlim(0, xmax + 1)
for i, v in enumerate(sv_global_imp):
margin = 0.05
ax.text(v + margin if v > 0 else margin, i, str(round(v, 4)), color='black', ha='left', va='center')
plt.show()
```
| github_jupyter |
## This notebook is create sample site along the street map
1. Prepare the street map
2. Set the sampling distance along the street
### 1. Clean the street map remove some types of streets
If you download the street map from open street map, you may need to remove some types of streets for your analyses, like highways, motor ways.
```
import fiona
import os,os.path
from shapely.geometry import shape,mapping
from shapely.ops import transform
from functools import partial
import pyproj
from fiona.crs import from_epsg
from shapely.geometry import LineString, Point # To create line geometries that can be used in a GeoDataFrame
import math
count = 0
# s = {'trunk_link','tertiary','motorway','motorway_link','steps', None, ' ','pedestrian','primary', 'primary_link','footway','tertiary_link', 'trunk','secondary','secondary_link','tertiary_link','bridleway','service'}
# s = {'trunk_link','tertiary','motorway','motorway_link','steps', ' ','pedestrian','primary', 'primary_link','footway','tertiary_link', 'trunk','secondary','secondary_link','tertiary_link','bridleway','service'}
s = {}
# set as your own filename
inshp = '../sample-spatialdata/CambridgeStreet_wgs84.shp'
outshp = '../sample-spatialdata/Cambridge20m.shp'
inshp = '/Users/xiaojiang/Dropbox (MIT)/ResearchProj/treepedia/cities-proj/chihuahua/red_vial_wgs.shp'
outshp = '/Users/xiaojiang/Dropbox (MIT)/ResearchProj/treepedia/cities-proj/chihuahua/street20m.shp'
# specify the distance
mini_dist = 20
# the temporaray file of the cleaned data
root = os.path.dirname(inshp)
basename = 'clean_' + os.path.basename(inshp)
temp_cleanedStreetmap = os.path.join(root,basename)
# if the tempfile exist then delete it
if os.path.exists(temp_cleanedStreetmap):
fiona.remove(temp_cleanedStreetmap, 'ESRI Shapefile')
print ('removed the existed tempfile')
# clean the original street maps by removing highways, if it the street map not from Open street data, users'd better to clean the data themselve
with fiona.open(inshp) as source, fiona.open(temp_cleanedStreetmap, 'w', driver=source.driver, crs=source.crs,schema=source.schema) as dest:
for feat in source:
try:
i = feat['properties']['highway'] # for the OSM street data
# i = feat['properties']['fclass'] # for the OSM tokyo street data
if i in s:
continue
except:
# if the street map is not osm, do nothing. You'd better to clean the street map, if you don't want to map the GVI for highways
key = list(dest.schema['properties'].keys())[0]
# key = dest.schema['properties'].keys()[0] # get the field of the input shapefile and duplicate the input feature
i = feat['properties'][key]
if i in s:
continue
# print feat
dest.write(feat)
schema = {
'geometry': 'Point',
'properties': {'id': 'int'},
}
```
### 2. Example of using Pyproj to do projection transform
```
import numpy as np
import pyproj
# Use the "new" numpy API for the random number generation (requires numpy >=1.17)
rng = np.random.default_rng(seed=1)
# Create arrays with random points
no_points = 5
longitudes = rng.uniform(low=-110, high=-100, size=no_points)
latitudes = rng.uniform(low=33, high=45, size=no_points)
print(" Longitude \t Latitude")
print(np.column_stack((longitudes, latitudes)))
print()
# Create the transformer object.
# You can create the object using either integers or "epsg" strings
transformer = pyproj.Transformer.from_crs(crs_from=4326, crs_to=3857)
# pyproj.Transformer.from_crs(crs_from="epsg:4326", crs_to="epsg:3857")
# Apply the transformation.
# Be careful! The order of the output here is Y/X
y1, x1 = transformer.transform(longitudes, latitudes)
# print(np.column_stack((x1, y1)))
# print()
# If you prefer to work with X/Y instead you need to set `always_xy=True` to the transformer object
transformer = pyproj.Transformer.from_crs(crs_from=4326, crs_to=3857, always_xy=True)
x2, y2 = transformer.transform(longitudes, latitudes)
print(np.column_stack((x2, y2)))
print()
```
### Create sample sites along the streets
Be careful of the pyproj version for the transform
```
from pyproj import Transformer
import math
# the transformer used to switch between the projections unit in degree to meter
transformer = Transformer.from_crs(4326, 3857, always_xy=True) #don't miss always_xy=True
transformer_back = Transformer.from_crs(3857, 4326, always_xy=True)
# Create point along the streets
with fiona.Env():
#with fiona.open(outshp, 'w', 'ESRI Shapefile', crs=source.crs, schema) as output:
with fiona.open(outshp, 'w', crs = from_epsg(4326), driver = 'ESRI Shapefile', schema = schema) as output:
i = 0
for line in fiona.open(temp_cleanedStreetmap):
i = i + 1
if i %1000 == 0: print(i)
# try:
# deal with MultiLineString and LineString
featureType = line['geometry']['type']
# for the LineString
if featureType == "LineString":
first = shape(line['geometry'])
length = first.length
# deal with different version of pyproj
if pyproj.__version__[0] != '2':
project = partial(pyproj.transform,pyproj.Proj(init='EPSG:4326'),pyproj.Proj(init='EPSG:3857')) #3857 is psudo WGS84 the unit is meter
line2 = transform(project, first)
print('using partial')
else:
# loop all vertices in the line and then reproj
line2_coord = []
for (lon, lat) in first.coords:
x, y = transformer.transform(lon, lat)
line2_coord.append((x, y))
line2 = LineString(line2_coord)
linestr = list(line2.coords)
dist = mini_dist
if math.isnan(line2.length): continue
# create sample points along lines and save to shapefile
for distance in range(0,int(line2.length), dist):
point = line2.interpolate(distance)
if pyproj.__version__[0]!='2':
project2 = partial(pyproj.transform,pyproj.Proj(init='EPSG:3857'),pyproj.Proj(init='EPSG:4326')) #3857 is psudo WGS84 the unit is meter
point = transform(project2, point)
else:
point = Point(transformer_back.transform(point.x, point.y))
output.write({'geometry':mapping(point),'properties': {'id':1}})
# for the MultiLineString, seperate these lines, then partition those lines
elif featureType == "MultiLineString":
multiline_geom = shape(line['geometry'])
print ('This is a multiline')
for singleLine in multiline_geom:
length = singleLine.length
if pyproj.__version__[0]!='2':
project = partial(pyproj.transform,pyproj.Proj(init='EPSG:4326'),pyproj.Proj(init='EPSG:3857')) #3857 is psudo WGS84 the unit is meter
line2 = transform(project, singleLine)
else:
# loop all vertices in the line and then reproj
line2_coord = []
for (lon, lat) in singleLine.coords:
x, y = transformer.transform(lon, lat)
line2_coord.append((x, y))
line2 = LineString(line2_coord)
linestr = list(line2.coords)
dist = mini_dist #set
if math.isnan(line2.length): continue
for distance in range(0,int(line2.length), dist):
point = line2.interpolate(distance)
if pyproj.__version__[0]!='2':
project2 = partial(pyproj.transform,pyproj.Proj(init='EPSG:3857'),pyproj.Proj(init='EPSG:4326')) #3857 is psudo WGS84 the unit is meter
point = transform(project2, point)
else:
point = Point(transformer_back.transform(point.x, point.y))
output.write({'geometry':mapping(point),'properties': {'id':1}})
else:
print('Else--------')
continue
# except:
# print ("You should make sure the input shapefile is WGS84")
# return
print("Process Complete")
# delete the temprary cleaned shapefile
# fiona.remove(temp_cleanedStreetmap, 'ESRI Shapefile')
```
| github_jupyter |
# User Defined Functions
User defined functions make for neater and more efficient programming.
We have already made use of several library functions in the math, scipy and numpy libraries.
```
import numpy as np
import scipy.constants as constants
print('Pi = ', constants.pi)
h = float(input("Enter the height of the tower (in metres): "))
t = float(input("Enter the time interval (in seconds): "))
s = constants.g*t**2/2
print("The height of the ball is",h-s,"meters")
```
Link to What's in Scipy.constants: https://docs.scipy.org/doc/scipy/reference/constants.html
## Library Functions in Maths
(and numpy)
```
x = 4**0.5
print(x)
x = np.sqrt(4)
print(x)
```
# User Defined Functions
Here we'll practice writing our own functions.
Functions start with
```python
def name(input):
```
and must end with a statement to return the value calculated
```python
return x
```
To run a function your code would look like this:
```python
import numpy as np
def name(input)
```
> FUNCTION CODE HERE
```python
return D
y=int(input("Enter y:"))
D = name(y)
print(D)
```
First - write a function to calculate n factorial. Reminder:
$n! = \pi^n_{k=1} k$
~
~
~
~
~
~
~
~
~
~
~
~
~
~
```
def factorial(n):
f = 1.0
for k in range(1,n+1):
f *= k
return f
print("This programme calculates n!")
n = int(input("Enter n:"))
a = factorial(10)
print("n! = ", a)
```
Finding distance to the origin in cylindrical co-ordinates:
```
from math import sqrt, cos, sin
def distance(r,theta,z):
x = r*cos(theta)
y = r*sin(theta)
d = sqrt(x**2+y**2+z**2)
return d
D = distance(2.0,0.1,1.5)
print(D)
```
# Another Example: Prime Factors and Prime Numbers
Reminder: prime factors are the numbers which divide another number exactly.
Factors of the integer n can be found by dividing by all integers from 2 up to n and checking to see which remainders are zero.
Remainder in python calculated using
```python
n % k
```
```
def factors(n):
factorlist=[]
k = 2
while k<=n:
while n%k==0:
factorlist.append(n)
n //= k
k += 1
return factorlist
list=factors(12)
print(list)
print(factors(17556))
print(factors(23))
```
The reason these are useful are for things like the below, where you want to make the same calculation many times. This finds all the prime numbers (only divided by 1 and themselves) from 2 to 100.
```
for n in range(2,100):
if len(factors(n))==1:
print(n)
```
# More Extended Example
Exercise 2.11 in Newman - binomial coefficients.
| github_jupyter |
# Classificação de músicas do Spotify
# Decision tree
Árvores de decisão são modelos estatísticos que utilizam um treinamento supervisionado para a classificação e previsão de dados. Uma árvore de decisão é utilizada para representar visualmente e explicitamente as decisões e a tomada de decisão. Apesar de ser amplamente utilizada em mineração de dados com algumas variações pode e é amplamente utilizada em aprendizado de máquina.
### Representação de uma árvore de decisão:
Uma arvore de decisão como representada na figura abaixo é representada por nós e ramos, onde cada nó representa um atributo a ser testado e cada ramo descendente representa um possível valor para tal atributo.

+ Dataset: https://www.kaggle.com/geomack/spotifyclassification
+ Info sobre as colunas: https://developer.spotify.com/web-api/get-audio-features/
+ https://graphviz.gitlab.io/download/
+ conda install -c anaconda graphviz
+ conda install -c conda-forge pydotplus
```
import graphviz
from matplotlib import pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
import pydotplus
import io
from scipy import misc
from sklearn import tree # pack age tree
from sklearn.metrics import accuracy_score # medir % acerto
from sklearn.model_selection import train_test_split # cortar dataset
from sklearn.tree import DecisionTreeClassifier, export_graphviz # arvore de decixao classificacao e graphviz para visualizar
```
*** Verificando os dados ***
```
df = pd.read_csv('data.csv')
df.head(3)
df.describe()
```
## Divisão treino e teste
### Divisão 70 - 30
```
train, test = train_test_split(df, test_size=round(len(df)*0.3))
print('Tamanho do set de treino: {},\nTamanho teste: {}'.format(len(train), len(test) ))
df.head()
```
O objetivo é dividir em grupos homogênioss com valor de 1 ou 0, dando uma serie de "caminhos" para determinar se o usuário gostou ou não da música.
```
# falar de overfitting comparando pagode funk e etc
# quanto maior o valor menor a arvore
tree = DecisionTreeClassifier(min_samples_split=100)
tree
features = ["danceability", "loudness", "valence", "energy", "instrumentalness", "acousticness", "key", "speechiness", "duration_ms"]
x_train = train[features]
y_train = train['target']
x_test = test[features]
y_test = test['target']
dct = tree.fit(x_train, y_train) # scikit fez uma decision tree
# visualizando
def showTree(tree, features, path):
file=io.StringIO()
export_graphviz(tree, out_file=file, feature_names=features)
pydotplus.graph_from_dot_data(file.getvalue()).write_png(path)
img = misc.imread(path)
plt.rcParams["figure.figsize"] = (20, 20)
plt.imshow(img)
#showTree(dct, features, 'minhaprimeiradct.png')
y_pred = tree.predict(x_test)
y_pred
score = accuracy_score(y_test, y_pred)*100
print('Score = {}'.format(score))
```
# Overfitting e Underfitting
*** Underfitting:*** Quando você treina seu algoritmo e testa ele no próprio conjunto de treino e percebe que ele ainda tem uma tacha de erro considerável e então testa ele no conjunto de teste e percebe que a taxa de erro é semelhante mas ainda alta.
Isso quer dizer que estamos diante de um caso de Underfitting o algoritmo tem um alto Bias e ainda podemos melhorar sua classificação, para isso deveremos mexer em alguns parâmetros do algoritmo.
Claro que em nem todos os casos ira ocorrer dessa forma depende da natureza do algoritmo.
*** Overfitting: *** Agora você treinou seu algoritmo e depois disso resolveu aplicá-lo em seu conjunto de treino e fica feliz quando percebe que ele teve uma taxa de erro de 00,35% por exemplo. Mas quando aplica no conjunto de teste percebe que ele tem um desempenho horrível.
# Random Forest Classifier
Florestas aleatórias ou florestas de decisão aleatórias são um método de aprendizado conjunto para classificação, regressão e outras tarefas que operam construindo uma multiplicidade de árvores de decisão no momento do treinamento e gerando a classe que é o modo das classes (classificação) ou predição de média ) das árvores individuais. Florestas de decisão aleatórias corrigem o hábito das árvores de decisão de overfitting em seu conjunto de treinamento.
```
from sklearn.ensemble import RandomForestClassifier
clf = RandomForestClassifier(n_estimators = 100)
clf.fit(x_train, y_train)
f_ypred = clf.predict(x_test)
score = accuracy_score(y_test, f_ypred) * 100
print('Score da decision tree: {}'.format(score))
```
# DESAFIO
https://www.kaggle.com/c/wine-quality-decision-tree
| github_jupyter |
# Solving the Diet Problem with *sasoptpy*
Diet problem, also known as *Stigler diet problem*, is one of the earliest optimization problems in the literature. [George J. Stigler](https://en.wikipedia.org/wiki/George_Stigler) originally posed the question of finding the cheapest diet, while satisfying the minimum nutritionial requirements [(Stigler, 1945)](www.jstor.org/stable/1231810).
This well-known problem can be solved with Linear Optimization easily. Since methodology was not developed in 1937, Stigler solved this problem using heuristics, albeit his solution was not the optimal (best) solution. He only missed the best solution by 24 cents per year.
Here, we will see how the problem can written in terms of mathematical equations and fed into SAS Viya Optimization solvers using the modeling capabilities of *sasoptpy* package.
## Organizing data
```
# Let us start with the nutritionial requrirements
# We use pandas package to represent tabular data
import pandas as pd
req = pd.DataFrame([
['calories', 3, 'kilocalories'],
['protein', 70, 'grams'],
['calcium', 0.8, 'grams'],
['iron', 12, 'milligrams'],
['vitamin_A', 5, 'thousand IU'],
['vitamin_B1', 1.8, 'milligrams'],
['vitamin_B2', 2.7, 'milligrams'],
['niacin', 18, 'milligrams'],
['vitamin_C', 75, 'milligrams']
], columns=['nutrient', 'allowance', 'unit']).set_index('nutrient')
# Print the table
from IPython.display import display, HTML
display(req)
```
The `diet.csv` file includes nutritional information from Stigler's 1945 paper. Each nutrient information is listed for $1 equivalent of the listed foods.
```
# Load the nutritional information from the file
nutr = pd.read_csv('diet.csv', index_col=0)
nutr.columns = req.index.tolist()
# Print first 10 rows for reference
display(nutr.head(10))
%matplotlib inline
# Show histogram of available values
import matplotlib.pyplot as plt
h = nutr.hist(figsize=(15,10))
display(h)
```
## Problem formulation
The diet problem can be written as a linear optimization formulation.
Denote $x_i$ as the servings of food item $i$ in our daily diet. The total cost of the diet is
$$
\sum_{i} c_i x_i
$$
where $c_i$ is the cost per serving of food item $i$.
Using the nutritional info, let us denote $n_{ij}$ as the amount of nutrition $j$ in food item $i$. Since we are trying to satisfy the minimum daily intake, we can write this constraint using the following mathematical expression:
$$
\sum_{i} n_{ij} x_i \geq m_j
$$
where $m_j$ is the minimum intake for nutrient $j$.
We can write the linear optimization model as follows
$$
\begin{array}{rrclcl}
\displaystyle \textrm{minimize}_{x} & \displaystyle {\sum_{i \in F} c_i x_i} \\
\textrm{subject to:} & \displaystyle \sum_{i \in F} n_{ij} x_i & \geq & m_j & & \forall j \in N \\
\end{array}
$$
## Modeling with Python
```
# We use sasoptpy to solve this problem
import sys
sys.path.insert(0, r'../../')
import sasoptpy as so
so.reset()
# We need swat package for connecting a running SAS Viya instance
from swat import CAS
# We will connect to the server
import os
host = os.environ['CASHOST']
port = os.environ['CASPORT']
session = CAS(host, port)
# Create a model object
m = so.Model(name='diet_problem', session=session)
# Define sets
F = nutr.index.tolist()
N = nutr.columns.tolist()
# Define variables
x = m.add_variables(F, name='intake', lb=0)
# Define objective
total_cost = m.set_objective(so.expr_sum(x[i] for i in F), sense=so.MIN, name='total_cost')
# Add the constraint
nutrient_con = m.add_constraints(
(so.expr_sum(nutr.at[i, j] * x[i] for i in F) >= req['allowance'][j] for j in N), name='nutrient_con'
)
# Solve the problem
sol = m.solve()
```
## Solution
The daily cost of our diet is 0.1086 USD (in 1939 dollars). It is equivalent to only 39.66 USD per year.
```
# Let us print the selected food items
for i in F:
if x[i].get_value() > 0:
print(i, round(x[i].get_value(), 4))
# Let us visualize the results
res = []
for i in F:
if x[i].get_value() > 0:
res.append([i, x[i].get_value(),])
resdf = pd.DataFrame(res, columns=['food', 'cost']).set_index(['food'])
resdf = resdf.join(nutr)
for i in N:
resdf['total_'+i] = resdf['cost'] * resdf[i]
resdf = resdf.drop([i], axis=1)
totaldf = resdf.append(resdf.sum().rename('Total'))
display(totaldf)
import matplotlib.pyplot as plt
fig, ax = plt.subplots(nrows=5, ncols=2, figsize=(14,20))
plt.subplots_adjust(wspace=0.6, hspace=0.4)
resdf.plot(kind='barh', subplots=True, ax=ax)
```
Navy beans and wheat flour dominate calories, protein, iron, vitamin B1, vitamin B2, and niacin intake.
| github_jupyter |
# Disambiguate sample editors
This notebook is concerned with identifying collected editorial board members in MAG. The process relies on the assumption that it is extremely unlikely that two scientists that shares the same name are affiliated with the same institution in any given years. Therefore, editors are disambiguated based on their first and last name (or first and middle initial and last name), affiliation, and year(s) of the affiliation.
```
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.metrics.pairwise import cosine_similarity
import pandas as pd
from fuzzywuzzy import fuzz
from unidecode import unidecode
import re
```
## Parse editor name
We allow first names with two letters to be interpreted in two ways. It is possible that "DJ Trump" is actually "D. J. Trump" but mistakened to be "Dj Trump" in the process of OCR, and it is also possible that "Dj" is actually someone's first name. Therefore, we allow both possiblities to exist in the first step of parsing.
In the later steps, only the correct interpretation of name can find a match within MAG, because it is extremely unlikely that "Dj Trump" and "D. J. Trump" are both affiliation with the same institution in the same year.
```
def parse_initial(name, mode=0):
'''
param: name - string
mode: 0 or 1, whether to treat length-2 first name as seperate initials
return: first, middle, last, first_initial, middle_initial, last_initial
'''
if mode != 0 and mode != 1:
raise("Error. Mode values must be 0 or 1")
l = name.split()
if len(l) == 0:
return '', '', '', '', '', ''
if len(l) == 1:
first, middle, last = '', '', l[0]
f_ini, m_ini, l_ini = '', '', last[0]
return first, middle, last, f_ini, m_ini, l_ini
# if the name is two parts, first name and last name
# assumes that if first or middle name is of length 1, it is an initial
elif len(l) == 2:
first, middle, last = l[0], '', l[1]
f_ini, m_ini, l_ini = first[0], '', last[0]
if len(first) == 1:
first = ''
elif len(first) == 2:
# if mode == 0, treat as first name
if mode == 1:
# treat as combination of two initials
f_ini, m_ini = first[0], first[1]
first = ''
return first, middle, last, f_ini, m_ini, l_ini
# if the name is three parts, first, middle, and last
elif len(l) == 3:
first, middle, last = l[0], l[1], l[2]
f_ini, m_ini, l_ini = first[0], middle[0], last[0]
if len(first) == 1:
first = ''
if len(middle) == 1:
middle = ''
return first, middle, last, f_ini, m_ini, l_ini
# if the name is more than three parts, first, middle, ..., last
elif len(l) > 3:
first, middle, last = l[0], ' '.join(l[1:-1]), l[-1]
f_ini, m_ini, l_ini = first[0], middle[0], last[0]
if len(first) == 1:
first = ''
if len(middle) == 1:
middle = ''
return first, middle, last, f_ini, m_ini, l_ini
all_editors = pd.read_csv("../data/SampleEditorsVol15.csv",sep='\t',
usecols=['issn','title','editorName','editorAff','Year'],
dtype={'issn':str,'Year':int,'issn':str})
print(all_editors.shape)
all_editors = all_editors.drop_duplicates()
print(all_editors.shape)
all_editors = all_editors.rename(columns={'editorName':'normalized_name'})
editors_0 = all_editors.assign(first = all_editors.normalized_name.apply(lambda x: parse_initial(x, 0)[0]),
middle = all_editors.normalized_name.apply(lambda x: parse_initial(x, 0)[1]),
last = all_editors.normalized_name.apply(lambda x: parse_initial(x, 0)[2]),
f_ini = all_editors.normalized_name.apply(lambda x: parse_initial(x, 0)[3]),
m_ini= all_editors.normalized_name.apply(lambda x: parse_initial(x, 0)[4]))
editors_1 = all_editors.assign(first = all_editors.normalized_name.apply(lambda x: parse_initial(x, 1)[0]),
middle = all_editors.normalized_name.apply(lambda x: parse_initial(x, 1)[1]),
last = all_editors.normalized_name.apply(lambda x: parse_initial(x, 1)[2]),
f_ini = all_editors.normalized_name.apply(lambda x: parse_initial(x, 1)[3]),
m_ini= all_editors.normalized_name.apply(lambda x: parse_initial(x, 1)[4]))
all_editors = editors_1.append(editors_0, ignore_index=True, sort=False)
all_editors.shape
all_editors = all_editors.dropna(subset=['editorAff'])
print(all_editors.shape)
all_editors = all_editors.drop_duplicates()
print(all_editors.shape)
all_editors.head()
```
## Remove country
This step is to remove as much noise as possible from intitution names.
Notice that Elsevier journals sometimes display affiliations as "institution name, city, country" but affiliations in MAG is much cleaner, i.e. without the city and country info. Therefore, to increase the accuracy when matching affiliations that we collected with the affiliations in MAG, we first remove all city, country, and country abbreviations from editors' affiliation. If an affiliation is within US or Canada, we also remove two-letter state abbreviations if found.
```
cities = pd.read_csv("../data/worldcities.csv", sep=",",
usecols=['city_ascii', 'country', 'iso2', 'iso3'],
dtype={'city_ascii':str, 'country':str, 'iso2':str, 'iso3':str}).drop_duplicates()
cities.shape
city_names = set([x.lower() for x in cities.city_ascii])
country_names = [x.lower() for x in cities.country]
country_names.extend([x.lower() for x in cities.iso2 if type(x) == str])
country_names.extend([x.lower() for x in cities.iso3])
country_names = set(country_names)
country_names.add('scotland')
country_names.add('uk')
us_states = [x.lower() for x in ["AL", "AK", "AZ", "AR", "CA", "CO", "CT", "DC", "DE", "FL", "GA",
"HI", "ID", "IL", "IN", "IA", "KS", "KY", "LA", "ME", "MD",
"MA", "MI", "MN", "MS", "MO", "MT", "NE", "NV", "NH", "NJ",
"NM", "NY", "NC", "ND", "OH", "OK", "OR", "PA", "RI", "SC",
"SD", "TN", "TX", "UT", "VT", "VA", "WA", "WV", "WI", "WY"] ]
canada_states = [x.lower() for x in ['AB', 'BC', 'MB', 'NB', 'NL', 'NT', 'NS', 'NU', 'ON', 'PE', 'QC', 'SK', 'YT'] ]
def clean_aff(x):
# throw away country and city names
# if in the States, identify state as well
l = str(x).split(',')
country = re.sub('[^0-9a-zA-Z ]+', '', l[-1]).strip()
if country in country_names:
l = l[:-1]
if len(l) > 0 and country in ['usa', 'us', 'united states', 'the united states']:
state = re.sub('[^0-9a-zA-Z]+', '', l[-1]).strip()
if state in us_states:
l = l[:-1]
if len(l) > 0 and country == 'canada':
state = re.sub('[^0-9a-zA-Z]+', '', l[-1]).strip()
if state in canada_states:
l = l[:-1]
if len(l) > 0:
city = re.sub('[^0-9a-zA-Z ]+', '', l[-1]).strip()
if city in city_names:
l = l[:-1]
return ' '.join(l).strip()
assert(clean_aff('university of california, ca, usa') == 'university of california')
assert(clean_aff('hong kong') == '')
assert(clean_aff('c a, united states') == '')
assert(clean_aff('peking university, beijing, china') == 'peking university')
all_editors = all_editors.assign(cleaned_aff = all_editors.editorAff.apply(clean_aff))
all_editors.head()
```
## Match editor names
Find potential matches who have the same name as editors.
```
def filterSameName(merged):
merged = merged[(merged.first_x == '') | (merged.first_y == '') | (merged.first_x == merged.first_y)]
merged = merged[(merged.middle_x =='') | (merged.middle_y == '') | (merged.middle_x == merged.middle_y)]
merged = merged[(merged.m_ini_x == '') | (merged.m_ini_y == '') | (merged.m_ini_x == merged.m_ini_y)]
merged = merged.drop(['first_x','first_y','middle_x','middle_y','last','f_ini','m_ini_x','m_ini_y'],axis=1)
return merged
%%time
authors = pd.read_csv("../data/mag/AuthorNames.csv", sep='\t', memory_map=True,
usecols=['NewAuthorId', 'first','middle','last', 'f_ini', 'm_ini'],
dtype={'NewAuthorId':int,'first':str,'last':str,'middle':str,'m_ini':str,'f_ini':str})
print(authors.shape)
all_editors = all_editors.fillna('')
authors = authors.fillna('')
%%time
merged = all_editors.merge(authors, on=['last', 'f_ini'])
print(merged.shape)
merged = filterSameName(merged)
print(merged.shape) # (195631, 7)
merged.head()
```
## Match editor affiliations
We use a combination of three measures of similarity to determine whether two institution names, when spelled differently, are representing the same affiliation.
`../data/manual_labeled_same.csv` and `../data/manual_labeled_different.csv` contains pairs of institutions that are manually labelled to be same or not. We use these manually labelled data as verification sets to determine the performance of chosen parameter. `../data/AllAffiliationSpellings.csv` contains all affiliations appeared as the affiliation of and editor (according to Elsevier) and all affiliations of a possible match (according to MAG).
```
def subfuzz(a, b):
if len(a) < len(b):
a, b = b, a
# len(a) >= len(b)
score = -1
lenb = len(b)
for start in range(len(a) - len(b)+1):
score = max(score, fuzz.ratio(a[start:start+lenb], b))
return score
assert(subfuzz('aaab', 'ab') == 100)
assert(subfuzz('ab', 'aaab') == 100)
assert(subfuzz('abcdabcabc', 'bcxy') == 50)
```
#### Get affiliation names
```
%%time
year_aff = pd.read_csv("../data/mag/AuthorAffYear.csv", sep='\t', memory_map=True,
usecols=['Year', 'NewAuthorId', 'AffiliationId'],
dtype={'Year':int, 'NewAuthorId':int, 'AffiliationId':int})
print(year_aff.shape)
affiliations = pd.read_csv("../data/mag/Affiliations.txt", sep="\t", memory_map=True,
names=['AffiliationId', "Rank", "NormalizedName", "DisplayName", "GridId",
"OfficialPage", "WikiPage", "PaperCount", "CitationCount", "Latitude",
"Longitude", 'CreatedDate'], usecols=['AffiliationId', 'NormalizedName'],
dtype={'AffiliationId': int, 'NormalizedName': str}).rename(
columns={'NormalizedName':'AffName'}).drop_duplicates()
print(affiliations.shape)
%%time
merged = merged.merge(year_aff, on=['NewAuthorId','Year'])
print(merged.shape)
merged = merged.merge(affiliations, on='AffiliationId')
print(merged.shape)
all_matched = merged.drop(['Year', 'AffiliationId'], axis=1).drop_duplicates() # all potential matches
all_matched = all_matched.rename(columns={'normalized_name':'EditorName'})
all_matched.shape
all_matched.head(3)
```
#### Get corpus
```
same = pd.read_csv("../data/manual_labeled_same.csv", sep="\t", index_col=0)
diff = pd.read_csv("../data/manual_labeled_different.csv", sep="\t", index_col=0)
allAff = pd.read_csv("../data/AllAffiliationSpellings.csv", sep="\t").fillna('')
same.shape, diff.shape, allAff.shape
allAff.head()
corpus = list(set(allAff.Affiliations).union(
set(same.affiliation)).union(set(same.affiliationMag)).union(
set(diff.affiliation)).union(set(diff.affiliationMag)))
# corpus = [re.sub('[^0-9a-zA-Z]+', ' ', x) for x in corpus]
len(corpus)
m = {}
for ind in range(len(corpus)):
m[ corpus[ind] ] = ind
%%time
vectorizer = TfidfVectorizer()
sparse_matrix = vectorizer.fit_transform(corpus)
doc_term_matrix = sparse_matrix.todense()
print(doc_term_matrix.shape)
```
#### Three outcomes and filter
```
%%time
all_matched = all_matched.assign(tfidf=all_matched.apply(lambda row:
cosine_similarity(doc_term_matrix[m[row['AffName']]], doc_term_matrix[m[row['cleaned_aff']]])[0][0], axis=1))
%%time
all_matched = all_matched.assign(fuz = all_matched.apply(
lambda row: fuzz.ratio(row['cleaned_aff'], row['AffName']), axis=1))
all_matched = all_matched.assign(subfuz = all_matched.apply(
lambda row: subfuzz(row['cleaned_aff'], row['AffName']), axis=1))
filtered = all_matched[(all_matched.tfidf>=33/100) & (all_matched.subfuz>=94) & (all_matched.subfuz>=19)]
filtered.shape # (1915, 11) # (1617, 11)
# only keep unique matches (each editor (issn, name) is uniquely matched to one author in MAG)
filtered = filtered.drop_duplicates(subset=['issn', 'EditorName', 'NewAuthorId'])
filtered = filtered.drop_duplicates(subset=['issn', 'EditorName'], keep=False)
filtered.shape
# find the start and end of editors
editors = filtered.merge(all_editors[['issn','normalized_name','Year']].drop_duplicates().rename(
columns={'normalized_name':'EditorName'}), on=['issn','EditorName']).groupby(
['issn','NewAuthorId']).agg({'Year':[min, max]}).reset_index()
editors.shape # 47
editors.columns = [f'{i} {j}'.strip() for i, j in editors.columns]
editors = editors.rename(columns={'Year min':'start_year', 'Year max':'end_year'})
editors.head()
```
Since we only parsed one editorial page, the start and end of editorial career is 2015 for all editors. Were the same procedure to be applied on the entired set of editorial pages, you can end up with the complete set of editors and their accurate start and end of editorial career.
From the set of editors that we have obtained accordingly, we sample 10 editors from the aforementioned journal, stored at `../data/SampleEditors.csv`, and use them to demonstrate susbsequent analysis.
| github_jupyter |
# Experiments with ConvNet Module for CpGNet
```
import keras
from keras.models import Sequential
from keras.layers import Dense, Activation, Dropout, Conv2D
from keras.layers import Embedding, GlobalAveragePooling1D, MaxPooling1D, MaxPooling2D,Flatten,Input,LeakyReLU
from keras.callbacks import EarlyStopping, ModelCheckpoint
from keras.models import Model
from CpG_Net import CpGNet
from CpG_Bin import Bin
import numpy as np
import cPickle as pickle
import matplotlib.pyplot as plt
import numpy as np
import matplotlib.pyplot as plt
from tqdm import tqdm
from sklearn.model_selection import train_test_split
from sklearn import preprocessing
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import AdaBoostClassifier
from sklearn.metrics import accuracy_score
from random import shuffle
from sklearn.metrics import roc_curve, auc
import random
from sklearn.metrics import roc_curve, auc
%load_ext autoreload
%autoreload 2
%matplotlib inline
CPG_DENSITY=6
data = pickle.load(open("HAMbins.p","rb"))
net = CpGNet(CPG_DENSITY)
```
Create complete bins
```
# Preliminary filter, useful to speed up computation
min_read_depth = 20
read_filtered_data = [bin_ for bin_ in data if bin_.matrix.shape[0] >= min_read_depth]
cpg_bins = [bin_ for bin_ in read_filtered_data if bin_.matrix.shape[1]==CPG_DENSITY]
shuffle(cpg_bins)
# Filters out missing data
cpg_bins_complete = net.filter_bad_reads(cpg_bins)
# secondary filter
cpg_bins_complete_depth = [bin_ for bin_ in cpg_bins_complete if bin_.matrix.shape[0] >= min_read_depth]
```
Create masks
```
masks = net.extract_masks(cpg_bins)
# apply masks
ready_bins = []
for Bin in cpg_bins_complete_depth:
truth_matrix = Bin.matrix
m_shape = truth_matrix.shape
if m_shape in masks:
if len(masks[m_shape]) > 0:
mask = random.choice(masks[m_shape])
observed = np.minimum(truth_matrix, mask)
Bin.tag2 = {"truth":truth_matrix, "observed":observed, "mask":mask}
ready_bins.append(Bin)
len(masks)
X,Y,Z = net.advanced_feature_collect(ready_bins)
```
Preprocessing
```
print X.shape
print Y.shape
print Z.shape
X[0]
max_depth = max([len(m) for m in X])
X_pad = np.zeros((len(X),max_depth, CPG_DENSITY))
temp = -5
for i,x in tqdm(enumerate(X)):
X_pad[i] = np.pad(x, ((0, max_depth-len(x)),(0,0)), "constant", constant_values=(temp))
# convert 1 to 3, 0 to -1, missing to 1, since we are 0 padding
X_pad[X_pad==1]=3
X_pad[X_pad==-1]=1
X_pad[X_pad==0]=-1
X_pad[X_pad==temp]=0
X[100]
X_pad[100]
X_exp = np.expand_dims(X_pad, axis=2) # add extra dimesion to make keras happy
X_exp=X_exp.reshape(len(X_exp),max_depth, CPG_DENSITY, 1)
Y_norm = preprocessing.scale(Y)
Z_exp = np.expand_dims(Z,-1)
#Y_exp = np.expand_dims(Y, axis=2) # add extra dimesion to make keras happy
print X_exp.shape
print Y_norm.shape
print Z.shape
Xf = X_exp.flatten()
Xfr = Xf.reshape(49229,498)
len(np.unique(X_exp,axis=0))
Xfr
np.save("npX",X_exp)
np.save("npY",Y_norm)
np.save("npZ",Z)
# Conv Module
convInput = Input(shape=(max_depth,CPG_DENSITY,1), dtype='float', name='input2')
filter_size = CPG_DENSITY
stride = filter_size
convLayer = Conv2D(32, kernel_size=(4,4), strides=2, padding="same",activation="linear")(convInput)
convLayer = LeakyReLU(alpha=.001)(convLayer)
convLayer = Conv2D(16, kernel_size=(2,2), strides=2, padding="same",activation="linear")(convLayer)
convLayer = LeakyReLU(alpha=.001)(convLayer)
convLayer = Conv2D(8, kernel_size=(2,2), strides=2, padding="same",activation="linear")(convLayer)
convLayer = LeakyReLU(alpha=.001)(convLayer)
#convLayer = MaxPooling2D()(convLayer)
convLayer = Flatten()(convLayer)
#convLayer = Flatten()(convInput)
#convLayer = Dense(1000, activation="relu")(convLayer)
# Numerical Module
numericalInput = Input(shape=(Y[0].size,), dtype='float', name='input1')
layer1 = Dense(1000, activation="linear")(numericalInput)
layer1 = LeakyReLU(alpha=.01)(layer1)
combined = Dropout(0.9)(combined)
layer1 = Dense(800, activation="linear")(layer1)
layer1 = LeakyReLU(alpha=.01)(layer1)
combined = Dropout(0.9)(combined)
layer1 = Dense(600, activation="linear")(layer1)
layer1 = LeakyReLU(alpha=.01)(layer1)
combined = Dropout(0.9)(combined)
layer1 = Dense(200, activation="linear")(layer1)
layer1 = LeakyReLU(alpha=.01)(layer1)
layer1 = Dense(100, activation="linear")(layer1)
layer1 = LeakyReLU(alpha=.01)(layer1)
layer1 = Dense(10, activation="linear")(layer1)
layer1 = LeakyReLU(alpha=.01)(layer1)
layer1 = Dense(3, activation="linear")(layer1)
layer1 = LeakyReLU(alpha=.01)(layer1)
layer1 = Dense(10, activation="linear")(layer1)
layer1 = LeakyReLU(alpha=.01)(layer1)
# Combined Module
combined = keras.layers.concatenate([convLayer, numericalInput])
combined = Dense(1000, activation="linear")(combined)
combined = LeakyReLU(alpha=.01)(combined)
combined = Dropout(0.9)(combined)
combined = Dense(800, activation="linear")(combined)
combined = LeakyReLU(alpha=.01)(combined)
combined = Dropout(0.9)(combined)
combined = Dense(400, activation="linear")(combined)
combined = LeakyReLU(alpha=.01)(combined)
combined = Dropout(0.9)(combined)
combined = Dense(1, activation="sigmoid")(combined)
model = Model(inputs=[convInput, numericalInput], outputs=[combined])
adam = keras.optimizers.Adam(lr=0.0001)
model.compile(optimizer="adam",loss = "binary_crossentropy",metrics=["acc"])
u, indices = np.unique(Xfr,axis=0, return_index=True)
X_u = X_exp[indices]
Y_u = Y_norm[indices]
Z_u = Z[indices]
history = model.fit([X_exp, Y_norm], [Z], epochs=100, validation_split=0.2, batch_size=16)
#history = model.fit([X_u, Y_u], [Z_u], epochs=100, validation_split=0.2, batch_size=16)
indices
Y[0]
# model = Sequential()
# model.add(Conv1D(16, 3, strides=3, activation='relu', input_shape=(seq_length,1)))
# model.add(Conv1D(5, 1, strides=1, activation='relu'))
# model.add(GlobalAveragePooling1D())
# model.add(Flatten())
# model.add(Dense(2, activation='sigmoid'))
# model.compile(loss='mse',
# optimizer='adam',
# metrics=['mse'])
# history = model.fit(X_exp, y, batch_size=1, epochs=1000, validation_split=0.0)
model.predict(X_exp)
y.shape
y_exp = y_exp.reshape(2,1,1)
y_exp
```
# Multiple input
Let's see if we can get a neural net to add two numbers together, x+y=z
We did it!
```
x = np.array([[[1,0,0],[0,1,1],[0,1,1]],[[0,0,0],[0,1,1],[0,1,1]]])
x_exp = np.expand_dims(x, axis=2)
y = np.array([1,0])
model = Sequential()
model.add(Conv2D(3, kernel_size=(2, 2), strides=(1, 1),
activation='relu',
padding ="same",
input_shape=x_exp[0].shape))
model.add(Flatten())
model.add(Dense(10, activation='relu'))
model.add(Dense(1, activation='sigmoid'))
model.compile(loss='binary_crossentropy',metrics=["acc"], optimizer="adam")
model.fit(x_exp, y, batch_size=32, epochs=1000)
model.predict(x_exp)
test = np.array([[1,1,2],[2,1,1]])
max_len = 5
test_pad = np.pad(test, ((0,max_len-len(test)),(0,0)), "constant", constant_values=(0))
test_pad
rf = RandomForestClassifier(n_estimators=100)
rf.fit(Y_norm[:10000], Z[:10000])
rf.score(Y_norm[:1000], Z[:1000])
rf.score(Y_norm[10000:11000], Z[10000:11000])
```
| github_jupyter |
## Introduction
## Preprocessing
Our data originates from plain text files in this format:
authorID [tab] review_text

Using some of our scripts, you can easily convert any document with the afore mentioned format
to be encoded in two different ways:
1. Our Custom Encoding
2. Doc2Vec Encoding
For our custom encoding, the script can be run as follows:
./customDoc2Vec.py [input filename] [output filename]
For the Doc2Vec encoding:
./spacyDoc2Vec.py [input filename] [output filename]
The format of the resulting encoding is as follows:
authorID [tab] word_vector
You can also reverse the sentences in a text corpus by running (it will output the same filename with 'r_' in front):
./reverseText.py [filename]
These methods above use pretrained word vector models; to create your own you can do the following:
./gensimDoc2Vec.py [input filename] [output filename] [model name]
## Setup
```
import keras
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
from IPython.display import display
from keras_tqdm import TQDMNotebookCallback
import copy
from keras import regularizers
from sklearn.utils import shuffle
```
## Reading Vector Data and One-Hot Creation
```
# choose file here
# options:
# 1. c_encoded.txt
# 2. c_encoded_r.txt
# 3. g_encoded.txt
# 4. g_encoded_r.txt
# 5. s_encoded.txt
# 6. s_encoded_r.txt
# Specifically for the demo:
# 1. demo_reviews.txt
# 2. demo_reversed.txt
filename = 's_encoded.txt'
data = [] # holds all data (i.e. encoded vectors)
unique = {} # holds unique author IDs here
index = 0 # used for determining which array inded will be 1 in the hot hot encoding
y_train = [] # one hots are in here
author_reviews_count = 0
authors = 3 # number of unique authors to read data from
with open(filename) as file:
for line in file:
author = line.split('\t')[0].rstrip('\n')
r = line.split('\t')[1].rstrip('\n')[1:-1].split(',')
vector = [float(x.strip(',')) for x in r if x != '']
if author not in unique:
if len(unique) == authors:
break
unique[author] = np.zeros(authors)
unique[author][index] = 1
index += 1
if len(vector) == 300:
data.append(vector)
y_train.append(unique[author])
```
## Seperate Training/Test Sets
```
x_tr = []
y_tr = []
x_te = []
y_te = []
cur = y_train[0]
count = 0
for i in range(len(data)):
if (y_train[i] == cur).all():
# training set
if count < 4:
x_tr.append(data[i])
y_tr.append(y_train[i])
# test set
else:
x_te.append(data[i])
y_te.append(y_train[i])
else:
cur = y_train[i]
count = 0
# training set
if count < 4:
x_tr.append(data[i])
y_tr.append(y_train[i])
# test set
else:
x_te.append(data[i])
y_te.append(y_train[i])
count += 1
```
## Oversampling Option
```
'''
x1 = copy.deepcopy(x_tr)
y1 = copy.deepcopy(y_tr)
for x in range(2):
for i in x1:
x_tr.append(i)
for i in y1:
y_tr.append(i)
'''
print(len(x_tr)) # x training set size
print(len(y_tr)) # y training set size
print(len(x_te)) # x testing set size
print(len(y_te)) # y testing set size
```
## Shuffle and Reshape
```
x_training,y_training = shuffle(x_tr,y_tr, random_state=0) # shuffle training data
# ========================================================
x_training = np.array(x_tr)
print(x_training.shape)
# convert the 2d to 3d representation
x_training = x_training.reshape(x_training.shape + (1,))
print(x_training.shape)
output_shape = x_training.shape[1]
input_shape = x_training.shape[2]
y_training = np.array(y_tr)
print('X_training shape: ' + str(x_training.shape))
# ========================================================
print(x_training.shape)
print(y_training.shape)
x_te = np.array(x_te)
# convert the 2d to 3d representation
x_te = x_te.reshape(x_te.shape + (1,))
print('X_testing shape: ' + str(x_te.shape))
print(x_te.shape)
y_te = np.array(y_te)
print(y_te.shape)
# ========================================================
```
## Network 1
```
model_n1 = keras.Sequential()
model_n1.add(keras.layers.Conv1D(300, kernel_size=(3),strides=(1),
activation='relu',
kernel_regularizer=regularizers.l2(0.01),
activity_regularizer=regularizers.l1(0.01),
input_shape=[output_shape,
input_shape]))
model_n1.add(keras.layers.MaxPooling1D())
model_n1.add(keras.layers.Flatten())
model_n1.add(keras.layers.Dense(300, activation='relu'))
model_n1.add(keras.layers.Dense(y_training.shape[1], activation='softmax'))
#sgd = keras.optimizers.SGD(lr=0.01, decay=1e-6, momentum=0.9, nesterov=True)
model_n1.compile(loss=keras.losses.categorical_crossentropy,
#optimizer = sgd,
optimizer=keras.optimizers.Adam(),
metrics=['accuracy'])
model_n1.summary()
```
## Network 2
```
model_n2 = keras.Sequential()
model_n2.add(keras.layers.Conv1D(300, kernel_size=(3),strides=(1),
activation='relu',
input_shape=[output_shape,
input_shape]))
model_n2.add(keras.layers.MaxPooling1D())
model_n2.add(keras.layers.Flatten())
model_n2.add(keras.layers.Dense(300, activation='relu'))
model_n2.add(keras.layers.Dense(y_training.shape[1], activation='softmax'))
#sgd = keras.optimizers.SGD(lr=0.01, decay=1e-6, momentum=0.9, nesterov=True)
model_n2.compile(loss=keras.losses.categorical_crossentropy,
#optimizer = sgd,
optimizer=keras.optimizers.Adam(),
metrics=['accuracy'])
model_n2.summary()
```
## Network 3
```
model_n3 = keras.Sequential()
model_n3.add(keras.layers.Dense(300,
activation='relu',
input_shape=[output_shape,input_shape]))
model_n3.add(keras.layers.Flatten())
model_n3.add(keras.layers.Dense(y_training.shape[1], activation='softmax'))
#sgd = keras.optimizers.SGD(lr=0.01, decay=1e-6, momentum=0.9, nesterov=True)
model_n3.compile(loss=keras.losses.categorical_crossentropy,
#optimizer = sgd,
optimizer=keras.optimizers.Adam(),
metrics=['accuracy'])
model_n3.summary()
```
## Training
```
# choose model here ...
current_model = model_n2
batch_size = 5
epochs = 5
history = current_model.fit(x_training, y_training,
batch_size=batch_size,
epochs=epochs,
verbose=0,
shuffle=True,
callbacks=[TQDMNotebookCallback()],
validation_split = 0.2)
#x_te = np.array(x_te)
#score = model.evaluate(x_te, np.array([y_te]), verbose=1)
#print('Test loss:', score[0])
#print('Test accuracy:', score[1])
```
## Testing
```
score = current_model.evaluate(x_te, y_te, verbose=1)
preds = current_model.predict_on_batch(x_training)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
```
## Graphing
```
score = current_model.evaluate(x_te, y_te, verbose=1)
preds = current_model.predict_on_batch(x_te)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
plt.subplot(211)
plt.plot(history.history['acc'])
plt.plot(history.history['val_acc'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
# summarize history for loss
plt.subplot(212)
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.tight_layout()
plt.show()
```
## Predictions
```
def get_auth(l):
#l = list(l)
for i in unique:
if (unique[i] == l).all():
return i
index = 0
incorrect = 0
correct = 0
for i in range(len(preds)):
m = max(preds[i])
index = list(preds[i]).index(m)
l = np.zeros(authors)
l[index] = 1
print(preds[i],index,y_te[i],'Guessed:',get_auth(l),'Actual:',get_auth(y_te[i]),end=' ')
if y_te[i][index] == 1:
print('Correct')
correct += 1
else:
print('Incorrect')
incorrect += 1
print('Correct:',correct,'Incorrect:',incorrect,'Out of:',(correct+incorrect))
```
| github_jupyter |
# Control Flow Graph
The code in this notebook helps with obtaining the control flow graph of python functions.
**Prerequisites**
* This notebook needs some understanding on advanced concepts in Python, notably
* classes
## Control Flow Graph
The class `PyCFG` allows one to obtain the control flow graph.
```Python
from ControlFlow import gen_cfg, to_graph
cfg = gen_cfg(inspect.getsource(my_function))
to_graph(cfg)
```
```
import bookutils
from bookutils import print_content
import ast
import re
from graphviz import Source, Digraph
```
### Registry
```
REGISTRY_IDX = 0
REGISTRY = {}
def get_registry_idx():
global REGISTRY_IDX
v = REGISTRY_IDX
REGISTRY_IDX += 1
return v
def reset_registry():
global REGISTRY_IDX
global REGISTRY
REGISTRY_IDX = 0
REGISTRY = {}
def register_node(node):
node.rid = get_registry_idx()
REGISTRY[node.rid] = node
def get_registry():
return dict(REGISTRY)
```
### CFGNode
We start with the `CFGNode` representing each node in the control flow graph.
\todo{Augmented and annotated assignments (`a += 1`), (`a:int = 1`)}.
```
class CFGNode(dict):
def __init__(self, parents=[], ast=None):
assert type(parents) is list
register_node(self)
self.parents = parents
self.ast_node = ast
self.update_children(parents) # requires self.rid
self.children = []
self.calls = []
def i(self):
return str(self.rid)
def update_children(self, parents):
for p in parents:
p.add_child(self)
def add_child(self, c):
if c not in self.children:
self.children.append(c)
def lineno(self):
return self.ast_node.lineno if hasattr(self.ast_node, 'lineno') else 0
def __str__(self):
return "id:%d line[%d] parents: %s : %s" % (
self.rid, self.lineno(), str([p.rid for p in self.parents]),
self.source())
def __repr__(self):
return str(self)
def __eq__(self, other):
return self.rid == other.rid
def __neq__(self, other):
return self.rid != other.rid
def set_parents(self, p):
self.parents = p
def add_parent(self, p):
if p not in self.parents:
self.parents.append(p)
def add_parents(self, ps):
for p in ps:
self.add_parent(p)
def add_calls(self, func):
self.calls.append(func)
def source(self):
return ast.unparse(self.ast_node).strip()
def to_json(self):
return {
'id': self.rid,
'parents': [p.rid for p in self.parents],
'children': [c.rid for c in self.children],
'calls': self.calls,
'at': self.lineno(),
'ast': self.source()
}
```
### PyCFG
Next, the `PyCFG` class which is responsible for parsing, and holding the graph.
```
class PyCFG:
def __init__(self):
self.founder = CFGNode(
parents=[], ast=ast.parse('start').body[0]) # sentinel
self.founder.ast_node.lineno = 0
self.functions = {}
self.functions_node = {}
class PyCFG(PyCFG):
def parse(self, src):
return ast.parse(src)
class PyCFG(PyCFG):
def walk(self, node, myparents):
fname = "on_%s" % node.__class__.__name__.lower()
if hasattr(self, fname):
fn = getattr(self, fname)
v = fn(node, myparents)
return v
else:
return myparents
class PyCFG(PyCFG):
def on_module(self, node, myparents):
"""
Module(stmt* body)
"""
# each time a statement is executed unconditionally, make a link from
# the result to next statement
p = myparents
for n in node.body:
p = self.walk(n, p)
return p
class PyCFG(PyCFG):
def on_augassign(self, node, myparents):
"""
AugAssign(expr target, operator op, expr value)
"""
p = [CFGNode(parents=myparents, ast=node)]
p = self.walk(node.value, p)
return p
class PyCFG(PyCFG):
def on_annassign(self, node, myparents):
"""
AnnAssign(expr target, expr annotation, expr? value, int simple)
"""
p = [CFGNode(parents=myparents, ast=node)]
p = self.walk(node.value, p)
return p
class PyCFG(PyCFG):
def on_assign(self, node, myparents):
"""
Assign(expr* targets, expr value)
"""
if len(node.targets) > 1:
raise NotImplemented('Parallel assignments')
p = [CFGNode(parents=myparents, ast=node)]
p = self.walk(node.value, p)
return p
class PyCFG(PyCFG):
def on_pass(self, node, myparents):
return [CFGNode(parents=myparents, ast=node)]
class PyCFG(PyCFG):
def on_break(self, node, myparents):
parent = myparents[0]
while not hasattr(parent, 'exit_nodes'):
# we have ordered parents
parent = parent.parents[0]
assert hasattr(parent, 'exit_nodes')
p = CFGNode(parents=myparents, ast=node)
# make the break one of the parents of label node.
parent.exit_nodes.append(p)
# break doesn't have immediate children
return []
class PyCFG(PyCFG):
def on_continue(self, node, myparents):
parent = myparents[0]
while not hasattr(parent, 'exit_nodes'):
# we have ordered parents
parent = parent.parents[0]
assert hasattr(parent, 'exit_nodes')
p = CFGNode(parents=myparents, ast=node)
# make continue one of the parents of the original test node.
parent.add_parent(p)
# return the parent because a continue is not the parent
# for the just next node
return []
class PyCFG(PyCFG):
def on_for(self, node, myparents):
# node.target in node.iter: node.body
# The For loop in python (no else) can be translated
# as follows:
#
# for a in iterator:
# mystatements
#
# __iv = iter(iterator)
# while __iv.__length_hint() > 0:
# a = next(__iv)
# mystatements
init_node = CFGNode(parents=myparents,
ast=ast.parse('__iv = iter(%s)' % ast.unparse(node.iter).strip()).body[0])
ast.copy_location(init_node.ast_node, node.iter)
_test_node = CFGNode(
parents=[init_node],
ast=ast.parse('_for: __iv.__length__hint__() > 0').body[0])
ast.copy_location(_test_node.ast_node, node)
# we attach the label node here so that break can find it.
_test_node.exit_nodes = []
test_node = self.walk(node.iter, [_test_node])
extract_node = CFGNode(parents=test_node,
ast=ast.parse('%s = next(__iv)' % ast.unparse(node.target).strip()).body[0])
ast.copy_location(extract_node.ast_node, node.iter)
# now we evaluate the body, one at a time.
p1 = [extract_node]
for n in node.body:
p1 = self.walk(n, p1)
# the test node is looped back at the end of processing.
_test_node.add_parents(p1)
return _test_node.exit_nodes + test_node
class PyCFG(PyCFG):
def on_while(self, node, myparents):
# For a while, the earliest parent is the node.test
_test_node = CFGNode(
parents=myparents,
ast=ast.parse(
'_while: %s' % ast.unparse(node.test).strip()).body[0])
ast.copy_location(_test_node.ast_node, node.test)
_test_node.exit_nodes = []
test_node = self.walk(node.test, [_test_node])
# we attach the label node here so that break can find it.
# now we evaluate the body, one at a time.
assert len(test_node) == 1
p1 = test_node
for n in node.body:
p1 = self.walk(n, p1)
# the test node is looped back at the end of processing.
_test_node.add_parents(p1)
# link label node back to the condition.
return _test_node.exit_nodes + test_node
class PyCFG(PyCFG):
def on_if(self, node, myparents):
_test_node = CFGNode(
parents=myparents,
ast=ast.parse(
'_if: %s' % ast.unparse(node.test).strip()).body[0])
ast.copy_location(_test_node.ast_node, node.test)
test_node = self.walk(node.test, [ _test_node])
assert len(test_node) == 1
g1 = test_node
for n in node.body:
g1 = self.walk(n, g1)
g2 = test_node
for n in node.orelse:
g2 = self.walk(n, g2)
return g1 + g2
class PyCFG(PyCFG):
def on_binop(self, node, myparents):
left = self.walk(node.left, myparents)
right = self.walk(node.right, left)
return right
class PyCFG(PyCFG):
def on_compare(self, node, myparents):
left = self.walk(node.left, myparents)
right = self.walk(node.comparators[0], left)
return right
class PyCFG(PyCFG):
def on_unaryop(self, node, myparents):
return self.walk(node.operand, myparents)
class PyCFG(PyCFG):
def on_call(self, node, myparents):
def get_func(node):
if type(node.func) is ast.Name:
mid = node.func.id
elif type(node.func) is ast.Attribute:
mid = node.func.attr
elif type(node.func) is ast.Call:
mid = get_func(node.func)
else:
raise Exception(str(type(node.func)))
return mid
#mid = node.func.value.id
p = myparents
for a in node.args:
p = self.walk(a, p)
mid = get_func(node)
myparents[0].add_calls(mid)
# these need to be unlinked later if our module actually defines these
# functions. Otherwsise we may leave them around.
# during a call, the direct child is not the next
# statement in text.
for c in p:
c.calllink = 0
return p
class PyCFG(PyCFG):
def on_expr(self, node, myparents):
p = [CFGNode(parents=myparents, ast=node)]
return self.walk(node.value, p)
class PyCFG(PyCFG):
def on_return(self, node, myparents):
if type(myparents) is tuple:
parent = myparents[0][0]
else:
parent = myparents[0]
val_node = self.walk(node.value, myparents)
# on return look back to the function definition.
while not hasattr(parent, 'return_nodes'):
parent = parent.parents[0]
assert hasattr(parent, 'return_nodes')
p = CFGNode(parents=val_node, ast=node)
# make the break one of the parents of label node.
parent.return_nodes.append(p)
# return doesnt have immediate children
return []
class PyCFG(PyCFG):
def on_functiondef(self, node, myparents):
# a function definition does not actually continue the thread of
# control flow
# name, args, body, decorator_list, returns
fname = node.name
args = node.args
returns = node.returns
enter_node = CFGNode(
parents=[],
ast=ast.parse('enter: %s(%s)' % (node.name, ', '.join(
[a.arg for a in node.args.args]))).body[0]) # sentinel
enter_node.calleelink = True
ast.copy_location(enter_node.ast_node, node)
exit_node = CFGNode(
parents=[],
ast=ast.parse('exit: %s(%s)' % (node.name, ', '.join(
[a.arg for a in node.args.args]))).body[0]) # sentinel
exit_node.fn_exit_node = True
ast.copy_location(exit_node.ast_node, node)
enter_node.return_nodes = [] # sentinel
p = [enter_node]
for n in node.body:
p = self.walk(n, p)
for n in p:
if n not in enter_node.return_nodes:
enter_node.return_nodes.append(n)
for n in enter_node.return_nodes:
exit_node.add_parent(n)
self.functions[fname] = [enter_node, exit_node]
self.functions_node[enter_node.lineno()] = fname
return myparents
class PyCFG(PyCFG):
def get_defining_function(self, node):
if node.lineno() in self.functions_node:
return self.functions_node[node.lineno()]
if not node.parents:
self.functions_node[node.lineno()] = ''
return ''
val = self.get_defining_function(node.parents[0])
self.functions_node[node.lineno()] = val
return val
class PyCFG(PyCFG):
def link_functions(self):
for nid, node in REGISTRY.items():
if node.calls:
for calls in node.calls:
if calls in self.functions:
enter, exit = self.functions[calls]
enter.add_parent(node)
if node.children:
# # until we link the functions up, the node
# # should only have succeeding node in text as
# # children.
# assert(len(node.children) == 1)
# passn = node.children[0]
# # We require a single pass statement after every
# # call (which means no complex expressions)
# assert(type(passn.ast_node) == ast.Pass)
# # unlink the call statement
assert node.calllink > -1
node.calllink += 1
for i in node.children:
i.add_parent(exit)
# passn.set_parents([exit])
# ast.copy_location(exit.ast_node, passn.ast_node)
# #for c in passn.children: c.add_parent(exit)
# #passn.ast_node = exit.ast_node
class PyCFG(PyCFG):
def update_functions(self):
for nid, node in REGISTRY.items():
_n = self.get_defining_function(node)
class PyCFG(PyCFG):
def update_children(self):
for nid, node in REGISTRY.items():
for p in node.parents:
p.add_child(node)
class PyCFG(PyCFG):
def gen_cfg(self, src):
"""
>>> i = PyCFG()
>>> i.walk("100")
5
"""
node = self.parse(src)
nodes = self.walk(node, [self.founder])
self.last_node = CFGNode(parents=nodes, ast=ast.parse('stop').body[0])
ast.copy_location(self.last_node.ast_node, self.founder.ast_node)
self.update_children()
self.update_functions()
self.link_functions()
```
### Supporting Functions
```
def compute_dominator(cfg, start=0, key='parents'):
dominator = {}
dominator[start] = {start}
all_nodes = set(cfg.keys())
rem_nodes = all_nodes - {start}
for n in rem_nodes:
dominator[n] = all_nodes
c = True
while c:
c = False
for n in rem_nodes:
pred_n = cfg[n][key]
doms = [dominator[p] for p in pred_n]
i = set.intersection(*doms) if doms else set()
v = {n} | i
if dominator[n] != v:
c = True
dominator[n] = v
return dominator
def compute_flow(pythonfile):
cfg, first, last = get_cfg(pythonfile)
return cfg, compute_dominator(
cfg, start=first), compute_dominator(
cfg, start=last, key='children')
def gen_cfg(fnsrc, remove_start_stop=True):
reset_registry()
cfg = PyCFG()
cfg.gen_cfg(fnsrc)
cache = dict(REGISTRY)
if remove_start_stop:
return {
k: cache[k]
for k in cache if cache[k].source() not in {'start', 'stop'}
}
else:
return cache
def get_cfg(src):
reset_registry()
cfg = PyCFG()
cfg.gen_cfg(src)
cache = dict(REGISTRY)
g = {}
for k, v in cache.items():
j = v.to_json()
at = j['at']
parents_at = [cache[p].to_json()['at'] for p in j['parents']]
children_at = [cache[c].to_json()['at'] for c in j['children']]
if at not in g:
g[at] = {'parents': set(), 'children': set()}
# remove dummy nodes
ps = set([p for p in parents_at if p != at])
cs = set([c for c in children_at if c != at])
g[at]['parents'] |= ps
g[at]['children'] |= cs
if v.calls:
g[at]['calls'] = v.calls
g[at]['function'] = cfg.functions_node[v.lineno()]
return (g, cfg.founder.ast_node.lineno, cfg.last_node.ast_node.lineno)
def to_graph(cache, arcs=[]):
graph = Digraph(comment='Control Flow Graph')
colors = {0: 'blue', 1: 'red'}
kind = {0: 'T', 1: 'F'}
cov_lines = set(i for i, j in arcs)
for nid, cnode in cache.items():
lineno = cnode.lineno()
shape, peripheries = 'oval', '1'
if isinstance(cnode.ast_node, ast.AnnAssign):
if cnode.ast_node.target.id in {'_if', '_for', '_while'}:
shape = 'diamond'
elif cnode.ast_node.target.id in {'enter', 'exit'}:
shape, peripheries = 'oval', '2'
else:
shape = 'rectangle'
graph.node(cnode.i(), "%d: %s" % (lineno, unhack(cnode.source())),
shape=shape, peripheries=peripheries)
for pn in cnode.parents:
plineno = pn.lineno()
if hasattr(pn, 'calllink') and pn.calllink > 0 and not hasattr(
cnode, 'calleelink'):
graph.edge(pn.i(), cnode.i(), style='dotted', weight=100)
continue
if arcs:
if (plineno, lineno) in arcs:
graph.edge(pn.i(), cnode.i(), color='green')
elif plineno == lineno and lineno in cov_lines:
graph.edge(pn.i(), cnode.i(), color='green')
# child is exit and parent is covered
elif hasattr(cnode, 'fn_exit_node') and plineno in cov_lines:
graph.edge(pn.i(), cnode.i(), color='green')
# parent is exit and one of its parents is covered.
elif hasattr(pn, 'fn_exit_node') and len(
set(n.lineno() for n in pn.parents) | cov_lines) > 0:
graph.edge(pn.i(), cnode.i(), color='green')
# child is a callee (has calleelink) and one of the parents is covered.
elif plineno in cov_lines and hasattr(cnode, 'calleelink'):
graph.edge(pn.i(), cnode.i(), color='green')
else:
graph.edge(pn.i(), cnode.i(), color='red')
else:
order = {c.i(): i for i, c in enumerate(pn.children)}
if len(order) < 2:
graph.edge(pn.i(), cnode.i())
else:
o = order[cnode.i()]
graph.edge(pn.i(), cnode.i(), color=colors[o], label=kind[o])
return graph
def unhack(v):
for i in ['if', 'while', 'for', 'elif']:
v = re.sub(r'^_%s:' % i, '%s:' % i, v)
return v
```
### Examples
#### check_triangle
```
def check_triangle(a, b, c):
if a == b:
if a == c:
if b == c:
return "Equilateral"
else:
return "Isosceles"
else:
return "Isosceles"
else:
if b != c:
if a == c:
return "Isosceles"
else:
return "Scalene"
else:
return "Isosceles"
import inspect
to_graph(gen_cfg(inspect.getsource(check_triangle)))
```
#### cgi_decode
Note that we do not yet support _augmented assignments_: i.e assignments such as `+=`
```
def cgi_decode(s):
hex_values = {
'0': 0, '1': 1, '2': 2, '3': 3, '4': 4,
'5': 5, '6': 6, '7': 7, '8': 8, '9': 9,
'a': 10, 'b': 11, 'c': 12, 'd': 13, 'e': 14, 'f': 15,
'A': 10, 'B': 11, 'C': 12, 'D': 13, 'E': 14, 'F': 15,
}
t = ""
i = 0
while i < len(s):
c = s[i]
if c == '+':
t += ' '
elif c == '%':
digit_high, digit_low = s[i + 1], s[i + 2]
i += 2
if digit_high in hex_values and digit_low in hex_values:
v = hex_values[digit_high] * 16 + hex_values[digit_low]
t += chr(v)
else:
raise ValueError("Invalid encoding")
else:
t += c
i += 1
return t
to_graph(gen_cfg(inspect.getsource(cgi_decode)))
```
#### gcd
```
def gcd(a, b):
if a<b:
c: int = a
a: int = b
b: int = c
while b != 0 :
c: int = a
a: int = b
b: int = c % b
return a
to_graph(gen_cfg(inspect.getsource(gcd)))
def compute_gcd(x, y):
if x > y:
small = y
else:
small = x
for i in range(1, small+1):
if((x % i == 0) and (y % i == 0)):
gcd = i
return gcd
to_graph(gen_cfg(inspect.getsource(compute_gcd)))
```
#### fib
Note that the *for-loop* requires additional massaging. While we show the labels correctly, the *comparison node* needs to be extracted. Hence, the representation is not accurate.
```
def fib(n,):
ls = [0, 1]
for i in range(n-2):
ls.append(ls[-1] + ls[-2])
return ls
to_graph(gen_cfg(inspect.getsource(fib)))
```
#### quad_solver
```
def quad_solver(a, b, c):
discriminant = b^2 - 4*a*c
r1, r2 = 0, 0
i1, i2 = 0, 0
if discriminant >= 0:
droot = math.sqrt(discriminant)
r1 = (-b + droot) / (2*a)
r2 = (-b - droot) / (2*a)
else:
droot = math.sqrt(-1 * discriminant)
droot_ = droot/(2*a)
r1, i1 = -b/(2*a), droot_
r2, i2 = -b/(2*a), -droot_
if i1 == 0 and i2 == 0:
return (r1, r2)
return ((r1,i1), (r2,i2))
to_graph(gen_cfg(inspect.getsource(quad_solver)))
```
## Call Graph
### Install: Pyan Static Call Graph Lifter
```
import os
import networkx as nx # type: ignore
```
### Call Graph Helpers
```
import shutil
PYAN = 'pyan3' if shutil.which('pyan3') is not None else 'pyan'
if shutil.which(PYAN) is None:
# If installed from pypi, pyan may still be missing
os.system('pip install "git+https://github.com/uds-se/pyan#egg=pyan"')
PYAN = 'pyan3' if shutil.which('pyan3') is not None else 'pyan'
assert shutil.which(PYAN) is not None
def construct_callgraph(code, name="callgraph"):
file_name = name + ".py"
with open(file_name, 'w') as f:
f.write(code)
cg_file = name + '.dot'
os.system(f'{PYAN} {file_name} --uses --defines --colored --grouped --annotated --dot > {cg_file}')
def callgraph(code, name="callgraph"):
if not os.path.isfile(name + '.dot'):
construct_callgraph(code, name)
return Source.from_file(name + '.dot')
def get_callgraph(code, name="callgraph"):
if not os.path.isfile(name + '.dot'):
construct_callgraph(code, name)
return nx.drawing.nx_pydot.read_dot(name + '.dot')
```
### Example: Maze
To provide a meaningful example where you can easily change the code complexity and target location, we generate the maze source code from the maze provided as string. This example is loosely based on an old [blog post](https://feliam.wordpress.com/2010/10/07/the-symbolic-maze/) on symbolic execution by Felipe Andres Manzano (Quick shout-out!).
You simply specify the maze as a string. Like so.
```
maze_string = """
+-+-----+
|X| |
| | --+ |
| | | |
| +-- | |
| |#|
+-----+-+
"""
```
Each character in `maze_string` represents a tile. For each tile, a tile-function is generated.
* If the current tile is "benign" (` `), the tile-function corresponding to the next input character (D, U, L, R) is called. Unexpected input characters are ignored. If no more input characters are left, it returns "VALID" and the current maze state.
* If the current tile is a "trap" (`+`,`|`,`-`), it returns "INVALID" and the current maze state.
* If the current tile is the "target" (`#`), it returns "SOLVED" and the current maze state.
The code is generated using the function `generate_maze_code`.
```
# ignore
def maze(s: str) -> str:
return "" # Will be overwritten by exec()
# ignore
def target_tile() -> str:
return ' ' # Will be overwritten by exec()
def generate_print_maze(maze_string):
return """
def print_maze(out, row, col):
output = out +"\\n"
c_row = 0
c_col = 0
for c in list(\"\"\"%s\"\"\"):
if c == '\\n':
c_row += 1
c_col = 0
output += "\\n"
else:
if c_row == row and c_col == col: output += "X"
elif c == "X": output += " "
else: output += c
c_col += 1
return output
""" % maze_string
def generate_trap_tile(row, col):
return """
def tile_%d_%d(input, index):
try: HTMLParser().feed(input)
except: pass
return print_maze("INVALID", %d, %d)
""" % (row, col, row, col)
def generate_good_tile(c, row, col):
code = """
def tile_%d_%d(input, index):
if (index == len(input)): return print_maze("VALID", %d, %d)
elif input[index] == 'L': return tile_%d_%d(input, index + 1)
elif input[index] == 'R': return tile_%d_%d(input, index + 1)
elif input[index] == 'U': return tile_%d_%d(input, index + 1)
elif input[index] == 'D': return tile_%d_%d(input, index + 1)
else : return tile_%d_%d(input, index + 1)
""" % (row, col, row, col,
row, col - 1,
row, col + 1,
row - 1, col,
row + 1, col,
row, col)
if c == "X":
code += """
def maze(input):
return tile_%d_%d(list(input), 0)
""" % (row, col)
return code
def generate_target_tile(row, col):
return """
def tile_%d_%d(input, index):
return print_maze("SOLVED", %d, %d)
def target_tile():
return "tile_%d_%d"
""" % (row, col, row, col, row, col)
def generate_maze_code(maze, name="maze"):
row = 0
col = 0
code = generate_print_maze(maze)
for c in list(maze):
if c == '\n':
row += 1
col = 0
else:
if c == "-" or c == "+" or c == "|":
code += generate_trap_tile(row, col)
elif c == " " or c == "X":
code += generate_good_tile(c, row, col)
elif c == "#":
code += generate_target_tile(row, col)
else:
print("Invalid maze! Try another one.")
col += 1
return code
```
Now you can generate the maze code for an arbitrary maze.
```
maze_code = generate_maze_code(maze_string)
print_content(maze_code, filename='.py')
exec(maze_code)
# Appending one more 'D', you have reached the target.
print(maze("DDDDRRRRUULLUURRRRDDD"))
```
This is the corresponding call graph.
```
callgraph(maze_code)
```
## Cleanup
We're done, so we clean up:
```
if os.path.exists('callgraph.dot'):
os.remove('callgraph.dot')
if os.path.exists('callgraph.py'):
os.remove('callgraph.py')
```
| github_jupyter |
# Rolling window on NumPy arrays without `for` loops
[Self-link](https://colab.research.google.com/drive/1Zru_-zzbtylgitbwxbi0eDBNhwr8qYl6)
**Disclaimer**: *author could be wrong!* If you see the error, please, write me at [foobar167@gmail.com](mailto:foobar167@gmail.com?subject=%20Rolling%20window%20error).
It is possible to implement a rolling window for NumPy arrays and images *without explicit cycles* in Python. As a result the speed of a such rolling window will be comparable to the speed of **C** programming language. Because NumPy library is implemented in the **C** programming language. It is in several thousand times faster than explicit Python `for` cycles.
00. [Introduction](#introduction)
00. [Rolling 1D window for ND array in Numpy](#1d)
00. [Rolling 2D window for ND array in Numpy](#2d)
00. [Rolling 3D window for ND array in Numpy](#3d)
00. [Rolling MD window for ND array, where M ≤ N](#md)
00. [Rolling MD window for ND array for any M and N](#md-extended)
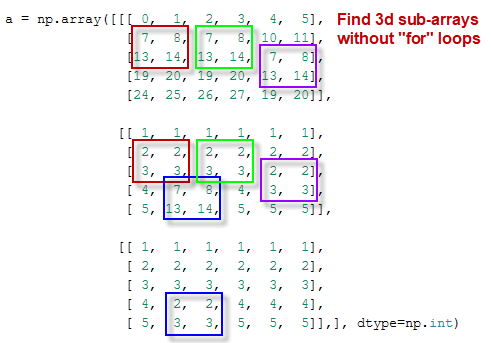
## <a name="introduction">Introduction</a>
This article is an extension of [my answer](https://stackoverflow.com/a/46237736/7550928) on the StackOverflow. My first experiments with the rolling window are [here](https://github.com/foobar167/junkyard/blob/master/rolling_window.py) and [here](https://github.com/foobar167/junkyard/blob/master/rolling_window_advanced.py).
**Practical implementation** of the rolling 2D window for 2D array in NumPy is in the `roll` function of the [`logic_tools.py`](https://github.com/foobar167/junkyard/blob/master/manual_image_annotation1/polygon/logic_tools.py) file of the [Manual image annotation with polygons](https://github.com/foobar167/junkyard/tree/master/manual_image_annotation1) project.
Basics rolling window technique for 1D array is already explained [here](https://stackoverflow.com/a/7100681/7550928), [here](https://rigtorp.se/2011/01/01/rolling-statistics-numpy.html) and [here](https://stackoverflow.com/questions/6811183/rolling-window-for-1d-arrays-in-numpy).
To understand the topic, you must know what [strides](https://stackoverflow.com/a/53099870/7550928) are.
```
# Import necessary libraries
import numpy as np
```
## <a name="1d" />1. Rolling 1D window for ND array in Numpy

```
# Rolling 1D window for ND array
def roll(a, # ND array
b, # rolling 1D window array
dx=1): # step size (horizontal)
shape = a.shape[:-1] + (int((a.shape[-1] - b.shape[-1]) / dx) + 1,) + b.shape
strides = a.strides[:-1] + (a.strides[-1] * dx,) + a.strides[-1:]
return np.lib.stride_tricks.as_strided(a, shape=shape, strides=strides)
```
There are 2 major things for newly created array: **shape** and **strides**. Shape of the new array is created via shape of the input ND array and shape of the rolling 1D window. While strides are created only via strides of the input ND array (without rolling window).
Step size of the rolling window **dx** is equal to 1, 2, 3, etc. and always horizontal for 1D rolling window.
*Shape* consists from 3 terms:
* `a.shape[:-1]` is the array shape remainder for ND arrays where `N > 1`. For `N == 1` array remainder will be equal to empty tuple `t == ()`, so it is not necessary for `N == 1`.
* `(int((a.shape[-1] - b.shape[-1]) / dx) + 1,)` is the number of steps of the rolling 1D window over last dimension of the ND array.
* `b.shape` is the shape of the rolling window.
*Strides* also consist from 3 terms:
* `a.strides[:-1]` is the array strides remainder for ND array where `N > 1`. For `N == 1` array remainder will be equal to empty tuple `t == ()`, so it is not necessary for `N == 1`.
* `(a.strides[-1] * dx,)` is the number of bytes between steps of the rolling window. For example, `int` array has 4 bytes stride between neighbour elements, so for step `dx == 2` it should be `4 * 2 = 8` bytes between steps of the rolling window.
* `(a.strides[-1],)` is the stride between neighbour elements of the input ND array. For example, for `int` it should be equal 4 bytes or tuple `(4,)`.
Function [`numpy.lib.stride_tricks.as_strided`](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.lib.stride_tricks.as_strided.html) creates a view into the array with the given shape and strides.
```
# Tests
def show_results(a, b, dx=1):
axis = a.ndim # number of dimensions
bool_array = np.all(roll(a, b, dx) == b, axis=axis)
counts = np.count_nonzero(bool_array)
coords = np.transpose(np.nonzero(bool_array)) * [1, dx]
print("Found {counts} elements with coordinates:\n{coords}".format(
counts=counts, coords=coords))
a = np.array([[ 0, 1, 2, 3, 4, 5],
[ 7, 8, 7, 8, 10, 11],
[13, 14, 13, 14, 7, 8],
[19, 20, 19, 20, 13, 14],
[24, 25, 26, 27, 19, 20]], dtype=np.int)
# Should find: 3 elements if dx == 1 or dx == 2
# 1 element if dx == 3
# 2 elements if dx == 4
b1 = np.array([7, 8], dtype=np.int)
show_results(a, b1)
show_results(a, b1, 2)
show_results(a, b1, 3)
show_results(a, b1, 4)
# Should find: 1 element if dx == 1
# 0 elements if dx == 2
b2 = np.array([8, 7], dtype=np.int)
print("----------")
show_results(a, b2)
show_results(a, b2, 2)
```
## <a name="2d" />2. Rolling 2D window for ND array in Numpy

Implementation examples of the rolling 2D window for 2D array are:
* find smaller subimage in the bigger image;
* do a convolution operation in the artificial neural network;
* apply filter of the artificial neural network or of the classical algorithm (Sobel, Gaussian or Blur filter) to the image.
In general it allows to make a *periodic operations* (comparison, convolution, subtraction, multiplication, some filter application, etc.) on a submatrix with some step size.
```
# Rolling 2D window for ND array
def roll(a, # ND array
b, # rolling 2D window array
dx=1, # horizontal step, abscissa, number of columns
dy=1): # vertical step, ordinate, number of rows
shape = a.shape[:-2] + \
((a.shape[-2] - b.shape[-2]) // dy + 1,) + \
((a.shape[-1] - b.shape[-1]) // dx + 1,) + \
b.shape # sausage-like shape with 2D cross-section
strides = a.strides[:-2] + \
(a.strides[-2] * dy,) + \
(a.strides[-1] * dx,) + \
a.strides[-2:]
return np.lib.stride_tricks.as_strided(a, shape=shape, strides=strides)
```
There are also shape and strides like in [section one](#1d), but for rolling 2D window.
*Shape* consists from 4 terms: 3 terms like in [section one](#1d) and the 4th term is the number of vertical steps `((a.shape[-2] - b.shape[-2]) // dy + 1,)` of the rolling 2D window.
Here we replaced:
```python
(int((a.shape[-1] - b.shape[-1]) / dx) + 1,)
```
with
```python
((a.shape[-1] - b.shape[-1]) // dx + 1,)
```
because these two expressions are equivalent.
*Strides* are also similar to [section one](#1d), but with additional stride `(a.strides[-2] * dy,)` for vertical step of the rolling 2D window.
```
# Tests
def show_results(a, b, dx=1, dy=1):
n = a.ndim # number of dimensions
# np.all over 2 dimensions of the rolling 2D window for 4D array
bool_array = np.all(roll(a, b, dx, dy) == b, axis=(n, n+1))
counts = np.count_nonzero(bool_array)
coords = np.transpose(np.nonzero(bool_array)) * [dy, dx]
print("Found {counts} elements with coordinates:\n{coords}".format(
counts=counts, coords=coords))
a = np.array([[ 0, 1, 2, 3, 4, 5],
[ 7, 8, 7, 8, 10, 11],
[13, 14, 13, 14, 7, 8],
[19, 20, 19, 20, 13, 14],
[24, 25, 26, 27, 19, 20]], dtype=np.int)
# Should find: 3 elements if dx == 1 or dx == 2
# 1 element if dx == 1 and dy == 2
# 2 elements if dx == 4
b1 = np.array([[ 7, 8],
[13, 14]], dtype=np.int)
show_results(a, b1)
show_results(a, b1, 2)
show_results(a, b1, 1, 2)
show_results(a, b1, 4)
# Should find: 1 element if dx == 1
# 0 elements if dx == 2
b2 = np.array([[ 8, 7],
[14, 13]], dtype=np.int)
print("----------")
show_results(a, b2)
show_results(a, b2, 2)
```
## <a name="3d" />3. Rolling 3D window for ND array in Numpy

You can see the pattern for 1D and 2D rolling window. It is difficult to understand the algorithm in higher dimensions, however it is possible to implement rolling 3D window for ND array in NumPy according to the pattern and then test it.
With this practical implementation you can apply voxels over 3D image.
```
# Rolling 3D window for ND array
def roll(a, # ND array
b, # rolling 2D window array
dx=1, # horizontal step, abscissa, number of columns
dy=1, # vertical step, ordinate, number of rows
dz=1): # transverse step, applicate, number of layers
shape = a.shape[:-3] + \
((a.shape[-3] - b.shape[-3]) // dz + 1,) + \
((a.shape[-2] - b.shape[-2]) // dy + 1,) + \
((a.shape[-1] - b.shape[-1]) // dx + 1,) + \
b.shape # multidimensional "sausage" with 3D cross-section
strides = a.strides[:-3] + \
(a.strides[-3] * dz,) + \
(a.strides[-2] * dy,) + \
(a.strides[-1] * dx,) + \
a.strides[-3:]
#print('shape =', shape, " strides =", strides) # for debugging
return np.lib.stride_tricks.as_strided(a, shape=shape, strides=strides)
# Tests
def show_results(a, b, dx=1, dy=1, dz=1):
n = a.ndim # number of dimensions == 3
# np.all over 3 dimensions of the rolling 3D window for 6D array
bool_array = np.all(roll(a, b, dx, dy, dz) == b, axis=(n, n+1, n+2))
counts = np.count_nonzero(bool_array)
coords = np.transpose(np.nonzero(bool_array)) * [dz, dy, dx]
print("Found {counts} elements with coordinates:\n{coords}".format(
counts=counts, coords=coords))
a = np.array([[[ 0, 1, 2, 3, 4, 5],
[ 7, 8, 7, 8, 10, 11],
[13, 14, 13, 14, 7, 8],
[19, 20, 19, 20, 13, 14],
[24, 25, 26, 27, 19, 20]],
[[ 1, 1, 1, 1, 1, 1],
[ 2, 2, 2, 2, 2, 2],
[ 3, 3, 3, 3, 2, 2],
[ 4, 7, 8, 4, 3, 3],
[ 5, 13, 14, 5, 5, 5]],
[[ 1, 1, 1, 1, 1, 1],
[ 2, 2, 2, 2, 2, 2],
[ 3, 3, 3, 3, 3, 3],
[ 4, 2, 2, 4, 4, 4],
[ 5, 3, 3, 5, 5, 5]],], dtype=np.int)
# Should find: 4 elements if dx == 1, dy == 1, dz == 1
# 3 elements if dx == 2, dy == 1, dz == 1
# 1 element if dx == 1, dy == 3, dz == 1
b1 = np.array([[[ 7, 8], [13, 14]],
[[ 2, 2], [ 3, 3]]], dtype=np.int)
show_results(a, b1)
show_results(a, b1, 2, 1, 1)
show_results(a, b1, 1, 3, 1)
# Should find: 1 element if dx == 1
# 0 elements if dx == 2
b2 = np.array([[[ 8, 7], [14, 13]],
[[ 2, 2], [ 3, 3]]], dtype=np.int)
print("----------")
show_results(a, b2)
show_results(a, b2, 2)
```
## <a name="md" />4. Rolling MD window for ND array, where M ≤ N
Generalize the `roll` function for rolling MD window over ND array, where M ≤ N.
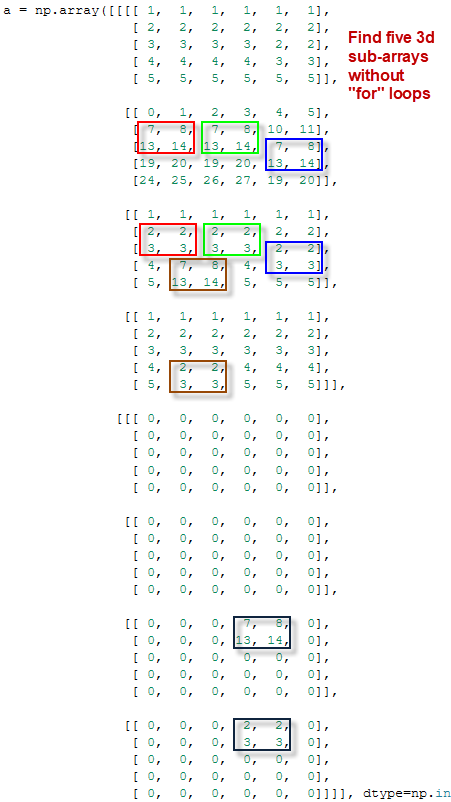
```
# Rolling MD window for ND array
def roll(a, # ND array
b, # rolling MD window array
d=None): # steps array
# Make several verifications
n = a.ndim # array dimensions
m = b.ndim # rolling window dimensions
if m > n: # check if M ≤ N
print("Error: rolling window dimensions is larger than the array dims")
return None
if d is None: # steps are equal to 1 by default
d = np.ones(m, dtype=np.uint32)
elif d.ndim != 1 and d.size != m:
print("Error: steps number must be equal to rolling window dimensions")
return None
elif not np.issubdtype(d.dtype, np.integer) or \
not (d > 0).all():
print("Error: steps must be integer and > 0")
return None
s = np.flip(d) # flip the 1D array of step sizes
sub = np.subtract(a.shape[-m:], b.shape[-m:])
steps = tuple(np.divide(sub, s).astype(np.uint32) + 1)
shape = a.shape[:-m] + steps + b.shape
section = tuple(np.multiply(a.strides[-m:], s))
strides = a.strides[:-m] + section + a.strides[-m:]
#print('shape =', shape, " strides =", strides) # for debugging
return np.lib.stride_tricks.as_strided(a, shape=shape, strides=strides)
```
The non-trivial (non-verifying) parts of the `roll` function are as follows:
* `steps = tuple(np.divide(sub, s).astype(np.uint32) + 1)` — calculate steps number of the rolling window in the multidimensional array.
* `section = tuple(np.multiply(a.strides[-m:], s))` — calculate strides section for multidimensional "sausage".
* Create that multidimensional "sausage" inserting this `section` into ND array: `strides = a.strides[:-m] + section + a.strides[-m:]`.
```
# Tests
def show_results(a, b, d=None):
n = a.ndim # array number of dimensions == N
m = b.ndim # rolling window dimensions == M
if d is None: # step sizes are equal to 1 by default
d = np.ones(m, dtype=np.uint32)
bool_array = roll(a, b, d) == b # get (N+M)D boolean array
# np.all over M dimensions of the rolling MD window for (N+M)D array
bool_array = np.all(bool_array, axis=tuple(range(n, n + m)))
counts = np.count_nonzero(bool_array)
# flip 1D array of step sizes and concatenate it with remaining dimensions
s = np.concatenate((np.ones(n-m, dtype=int), np.flip(d)))
coords = np.transpose(np.nonzero(bool_array)) * s
print("Found {counts} elements with coordinates:\n{coords}".format(
counts=counts, coords=coords))
a = np.array([[[[ 1, 1, 1, 1, 1, 1],
[ 2, 2, 2, 2, 2, 2],
[ 3, 3, 3, 3, 2, 2],
[ 4, 4, 4, 4, 3, 3],
[ 5, 5, 5, 5, 5, 5]],
[[ 0, 1, 2, 3, 4, 5],
[ 7, 8, 7, 8, 10, 11],
[13, 14, 13, 14, 7, 8],
[19, 20, 19, 20, 13, 14],
[24, 25, 26, 27, 19, 20]],
[[ 1, 1, 1, 1, 1, 1],
[ 2, 2, 2, 2, 2, 2],
[ 3, 3, 3, 3, 2, 2],
[ 4, 7, 8, 4, 3, 3],
[ 5, 13, 14, 5, 5, 5]],
[[ 1, 1, 1, 1, 1, 1],
[ 2, 2, 2, 2, 2, 2],
[ 3, 3, 3, 3, 3, 3],
[ 4, 2, 2, 4, 4, 4],
[ 5, 3, 3, 5, 5, 5]]],
[[[ 0, 0, 0, 0, 0, 0],
[ 0, 0, 0, 0, 0, 0],
[ 0, 0, 0, 0, 0, 0],
[ 0, 0, 0, 0, 0, 0],
[ 0, 0, 0, 0, 0, 0]],
[[ 0, 0, 0, 0, 0, 0],
[ 0, 0, 0, 0, 0, 0],
[ 0, 0, 0, 0, 0, 0],
[ 0, 0, 0, 0, 0, 0],
[ 0, 0, 0, 0, 0, 0]],
[[ 0, 0, 0, 7, 8, 0],
[ 0, 0, 0, 13, 14, 0],
[ 0, 0, 0, 0, 0, 0],
[ 0, 0, 0, 0, 0, 0],
[ 0, 0, 0, 0, 0, 0]],
[[ 0, 0, 0, 2, 2, 0],
[ 0, 0, 0, 3, 3, 0],
[ 0, 0, 0, 0, 0, 0],
[ 0, 0, 0, 0, 0, 0],
[ 0, 0, 0, 0, 0, 0]]]], dtype=np.int)
# Should find: 5 elements if dx == 1, dy == 1, dz == 1
# 3 elements if dx == 2, dy == 1, dz == 1
# 2 elements if dx == 1, dy == 3, dz == 1
# 2 elements if dx == 1, dy == 1, dz == 2
b1 = np.array([[[ 7, 8], [13, 14]],
[[ 2, 2], [ 3, 3]]], dtype=np.int)
show_results(a, b1)
show_results(a, b1, np.array([2, 1, 1]))
show_results(a, b1, np.array([1, 3, 1]))
show_results(a, b1, np.array([1, 1, 2]))
# Should find: 1 element if dx == 1
# 0 elements if dx == 2
b2 = np.array([[[ 8, 7], [14, 13]],
[[ 2, 2], [ 3, 3]]], dtype=np.int)
print("----------")
show_results(a, b2)
show_results(a, b2, np.array([2, 1, 1]))
```
The non-trivial parts of the `show_results` function are as follows:
* Get boolean array `bool_array` or **mask** of the rolling window searches. Then apply [`numpy.all`](https://docs.scipy.org/doc/numpy/reference/generated/numpy.all.html) to all `M` dimensions to test whether all array elements along a given dimension evaluate to `True`. Note that `bool_array` is (N+M)D array, but `np.all` applies `m` times for MD array of the rolling window:
```python
bool_array = roll(a, b, d) == b # get (N+M)D boolean array
# np.all over M dimensions of the rolling MD window for (N+M)D array
bool_array = np.all(bool_array, axis=tuple(range(n, n + m)))
```
* Another non-trivial part is for `M < N`. If `M < N` we should not only flip 1D array of step sizes, but to concatenate it with ones (step size is equal to 1) for remaining dimensions `N-M`. If `M == N` the remaining dimensions is 0, so in this case concatenation is not needed:
```python
# flip 1D array of step sizes and concatenate it with remaining dimensions
s = np.concatenate((np.ones(n-m, dtype=int), np.flip(d)))
```
## <a name="md-extended" />5. Rolling MD window for ND array for any M and N
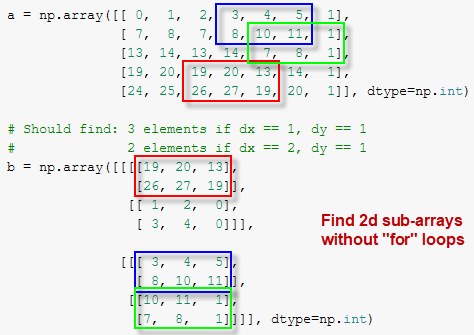

Can we roll MD window over ND arrays, where M > N? Actually, we can, but only part of the rolling window which **intersects** with ND array.
In other words lets find **common intersections** between MD and ND arrays.
Implement a rolling MD window for ND array for any M and N. For this we use the previous `roll` and `show_results` functions.
```
# Tests
def get_results(a, b, d=None): # the same as `show_results` function
n = a.ndim # array number of dimensions == N
m = b.ndim # rolling window dimensions == M
if d is None: # step sizes are equal to 1 by default
d = np.ones(m, dtype=np.uint32)
bool_array = roll(a, b, d) == b # get (N+M)D boolean array
# np.all over M dimensions of the rolling MD window for (N+M)D array
bool_array = np.all(bool_array, axis=tuple(range(n, n + m)))
counts = np.count_nonzero(bool_array)
# flip 1D array of step sizes and concatenate it with remaining dimensions
s = np.concatenate((np.ones(n-m, dtype=int), np.flip(d)))
coords = np.transpose(np.nonzero(bool_array)) * s
return (counts, coords)
def show_intersections(a, b, d=None):
d_tmp = d
n = a.ndim # array number of dimensions == N
m = b.ndim # rolling window dimensions == M
#
if d_tmp is None: # step sizes are equal to 1 by default
d_tmp = np.ones(m, dtype=np.uint32)
elif m > n and d_tmp.size == n: # for m > n case
# Concatenate d_tmp with remaining dimensions
d_tmp = np.concatenate((np.ones(m-n, dtype=int), d_tmp))
#
counts = 0
coords = None
if m <= n:
results = get_results(a, b, d_tmp) # return previous example
counts = results[0]
coords = results[1]
else: # if m > n
t = m - n # excessive dimensions
layers = np.prod(b.shape[:t]) # find number of layers
# Reshape MD array into (N+1)D array.
temp = b.reshape((layers,) + b.shape[t:])
# Get results for every layer in the intersection
for i in range(layers):
results = get_results(a, temp[i], d_tmp[t:])
counts += results[0]
if coords is None:
coords = results[1]
else:
coords = np.concatenate((coords, results[1]))
print("Found {counts} elements with coordinates:\n{coords}".format(
counts=counts, coords=coords))
a = np.array([[ 0, 1, 2, 3, 4, 5, 1],
[ 7, 8, 7, 8, 10, 11, 1],
[13, 14, 13, 14, 7, 8, 1],
[19, 20, 19, 20, 13, 14, 1],
[24, 25, 26, 27, 19, 20, 1]], dtype=np.int)
# Should find: 3 elements if dx == 1, dy == 1
# 2 elements if dx == 2, dy == 1
b = np.array([[[[19, 20, 13],
[26, 27, 19]],
[[ 1, 2, 0],
[ 3, 4, 0]]],
[[[ 3, 4, 5],
[ 8, 10, 11]],
[[10, 11, 1],
[7, 8, 1]]]], dtype=np.int)
show_intersections(a, b)
show_intersections(a, b, np.array([2, 1]))
# Should find: 1 element if dx == 1, dy == 1
print("----------")
b = np.array([[19, 20, 13],
[26, 27, 19]], dtype=np.int)
show_intersections(a, b)
```
`get_results` function is the same as `show_results` function without significant differences.
`show_intersections` function obtains intersections between arrays. If `M <= N` function `show_intersections` just returns `get_results` function. If `M > N` we should find intersection between `b` and `a` arrays. For this get excessive dimensions `t = m - n` between MD `b` array and ND `a` array. And find the number of layers in the intersection between `b` and `a`: `layers = np.prod(b.shape[:t])`.
Then reshape `b` array from MD array to (N+1)D array:
```python
# Reshape MD array into (N+1)D array.
temp = b.reshape((layers,) + b.shape[t:])
```
Finally we get results for every layer in the intersection:
```python
# Get results for every layer in the intersection
for i in range(layers):
results = get_results(a, temp[i], d_tmp[t:])
```
and aggregate counts and coordinates of all matches in the `counts` and `coords` variables:
```python
# Get results for every layer in the intersection
for i in range(layers):
results = get_results(a, temp[i], d_tmp[t:])
counts += results[0]
if coords is None:
coords = results[1]
else:
coords = np.concatenate((coords, results[1]))
```
```
```
| github_jupyter |
```
from __future__ import absolute_import, division, print_function
import torch
import torchvision.models as models
vgg19 = models.vgg19(pretrained=True)
import torch.nn as nn
import torchvision.transforms as transforms
import torch.nn.functional as F
from torch.autograd import Variable
from glob import glob
import os
import numpy as np
import pandas as pd
import json
import re
import matplotlib
from matplotlib import pylab, mlab, pyplot
%matplotlib inline
from IPython.core.pylabtools import figsize, getfigs
plt = pyplot
import seaborn as sns
sns.set_context('talk')
sns.set_style('white')
from PIL import Image
import base64
import sys
sys.path.insert(0, '../../python/')
from embeddings import *
import warnings
warnings.simplefilter("ignore")
!pip install argparse
import argparse
# directory & file hierarchy
proj_dir = os.path.abspath('../../..')
analysis_dir = os.getcwd()
results_dir = os.path.join(proj_dir,'results')
plot_dir = os.path.join(results_dir,'plots')
csv_dir = os.path.join(results_dir,'csv')
exp_dir = os.path.abspath(os.path.join(proj_dir,'experiments'))
sketch_dir = os.path.abspath(os.path.join(proj_dir,'sketches'))
#drawings_dir = os.path.join(sketch_dir,'run3_run4')
drawings_dir = os.path.join(os.path.join(sketch_dir, 'stroke_analysis'), 'png')
#h.save_sketches(D_run3_correct, complete_games, sketch_dir, 'run3_run4', iterationNum)
# retrieve sketch paths
def list_files(path, ext='png'):
result = [y for x in os.walk(path) for y in glob(os.path.join(x[0], '*.%s' % ext))]
return result
def check_invalid_sketch(filenames,invalids_path='drawings_to_exclude.txt'):
if not os.path.exists(invalids_path):
print('No file containing invalid paths at {}'.format(invalids_path))
invalids = []
else:
x = pd.read_csv(invalids_path, header=None)
x.columns = ['filenames']
invalids = list(x.filenames.values)
valids = []
basenames = [f.split('/')[-1] for f in filenames]
for i,f in enumerate(basenames):
if f not in invalids:
valids.append(filenames[i])
return valids
def make_dataframe(Labels):
Y = pd.DataFrame(Labels)
Y = Y.transpose()
Y.columns = ['label']
return Y
def normalize(X):
X = X - X.mean(0)
X = X / np.maximum(X.std(0), 1e-5)
return X
def preprocess_features(Features, Y):
_Y = Y.sort_values(['label'])
inds = np.array(_Y.index)
_Features = normalize(Features[inds])
_Y = _Y.reset_index(drop=True) # reset pandas dataframe index
return _Features, _Y
def save_features(Features, Y, layer_num, data_type,feat_path='./sketch_features'):
if not os.path.exists('./sketch_features'):
os.makedirs('./sketch_features')
layers = ['P1','P2','P3','P4','P5','FC6','FC7']
np.save(os.path.join(feat_path,'FEATURES_{}_{}_stroke_analysis.npy'.format(layers[int(layer_num)], data_type)), Features)
Y.to_csv(os.path.join(feat_path,'METADATA_{}_stroke_analysis.csv'.format(data_type)))
return layers[int(layer_num)]
parser = argparse.ArgumentParser()
parser.add_argument('--data', type=str, help='full path to images', default=drawings_dir)#'combined_final_png_drawings')
parser.add_argument('--layer_ind', help='fc6 = 5, fc7 = 6', default=5)
parser.add_argument('--data_type', help='"images" or "sketch"', default='sketch')
parser.add_argument('--spatial_avg', type=bool, help='collapse over spatial dimensions, preserving channel activation only if true', default=False)
parser.add_argument('--test', type=bool, help='testing only, do not save features', default=False)
parser.add_argument('--ext', type=str, help='image extension type (e.g., "png")', default="png")
args = parser.parse_args(args=[])
# args = parser.parse_args(argv[1:])
## get list of all sketch paths
image_paths = sorted(list_files(args.data,args.ext))
print('Length of image_paths before filtering: {}'.format(len(image_paths)))
## filter out invalid sketches
image_paths = check_invalid_sketch(image_paths)
print('Length of image_paths after filtering: {}'.format(len(image_paths)))
## extract features
layers = ['P1','P2','P3','P4','P5','FC6','FC7']
extractor = FeatureExtractor(image_paths,layer=args.layer_ind,data_type=args.data_type,spatial_avg=args.spatial_avg)
Features, Labels = extractor.extract_feature_matrix()
# # organize metadata into dataframe
# Y = make_dataframe(Labels)
Y = pd.DataFrame(Labels)
Y = Y.transpose()
Y.columns = ['0', '1', '2', '3', '4', '5', '6']
m = pd.DataFrame()
label_list = []
for i,d in Y.iterrows():
label = d['0'] + '_' + d['1'] + '_' + d['2'] + '_' + d['3'] + '_' + d['4'] + '_' + d['5'] + '_' + d['6']
label_list.append(label)
m['label'] = label_list
# if args.test==False:
# layer = save_features(_Features, _Y, args.layer_ind, args.data_type) # g,trialNum,target, repetition, iterationNum
_Features, _Y = preprocess_features(Features, m)
if args.test==False:
layer = save_features(_Features, _Y, args.layer_ind, args.data_type) # g,trialNum,target, repetition, iterationNum
#Y.to_csv('feature_matrix_raw.csv')
```
##### manually save it
```
feat_path = './sketch_features'
layer_num = args.layer_ind
data_type = 'sketch'
layers = ['P1','P2','P3','P4','P5','FC6','FC7']
out_path = os.path.join(feat_path,'FEATURES_{}_{}_stroke_analysis.npy'.format(layers[int(layer_num)], data_type))
np.save(out_path, _Features)
```
| github_jupyter |
```
import sys
sys.path.append(r'C:\Users\moallemie\EMAworkbench-master')
sys.path.append(r'C:\Users\moallemie\EM_analysis')
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from ema_workbench import load_results, ema_logging
sc = 2000 # Specify the number of scenarios where the convergence in the SA indices occured
n_exp = 2000 # Indicate the number of LHS experiments to run
t = 2100
nprocess = 25
```
## Load model, uncertainities; generate initial experiments
```
# Open Excel input data from the notebook directory before runnign the code in multi-processing.
def create_sets(outcome_var):
from Model_init import vensimModel
directory = 'C:/Users/moallemie/EM_analysis/Model/'
df_unc = pd.read_excel(directory+'ScenarioFramework.xlsx', sheet_name='Uncertainties')
# 0.75/1.25 multiplication is added to Min/Max for parameters with Reference values 0
#or min/max manually set in the spreadsheet
#df_unc['Min'] = df_unc['Min'] + df_unc['Reference'] * 0.75
#df_unc['Max'] = df_unc['Max'] + df_unc['Reference'] * 1.25
# From the Scenario Framework (all uncertainties), filter only those top 20 sensitive uncertainties under each outcome
sa_dir='C:/Users/moallemie/EM_analysis/Data/'
mu_df = pd.read_csv(sa_dir+"MorrisIndices_{}_sc{}_t{}_test.csv".format(outcome_var, sc, t))
mu_df.rename(columns={'Unnamed: 0': 'Uncertainty'}, inplace=True)
mu_df.sort_values(by=['mu_star'], ascending=False, inplace=True)
mu_df = mu_df.head(20)
mu_unc = mu_df['Uncertainty']
mu_unc_df = mu_unc.to_frame()
# Remove the rest of insensitive uncertainties from the Scenario Framework and update df_unc
keys = list(mu_unc_df.columns.values)
i1 = df_unc.set_index(keys).index
i2 = mu_unc_df.set_index(keys).index
df_unc2 = df_unc[i1.isin(i2)]
# Reorder the dataframe of mu index according to the model uncertainty.
# It will be used when we generate Sets 1 & 2
mu_df1 = mu_df[['Uncertainty', 'mu_star']]
mu_df1 = mu_df1.set_index('Uncertainty')
mu_df1 = mu_df1.reindex(index=df_unc2['Uncertainty'])
mu_df1 = mu_df1.reset_index()
vensimModel.uncertainties = [RealParameter(row['Uncertainty'], row['Min'], row['Max']) for index, row in df_unc2.iterrows()]
df_out = pd.read_excel(directory+'ScenarioFramework.xlsx', sheet_name='Outcomes')
vensimModel.outcomes = [TimeSeriesOutcome(out) for out in df_out['Outcome']]
from ema_workbench import MultiprocessingEvaluator
from ema_workbench.em_framework.evaluators import (MC, LHS, FAST, FF, PFF, SOBOL, MORRIS)
try:
with MultiprocessingEvaluator(vensimModel, n_processes=nprocess) as evaluator:
results = evaluator.perform_experiments(scenarios=n_exp, uncertainty_sampling=LHS) # The number of scenarios here is only for identifying the number of important parameters in SA results.
except (BrokenPipeError, IOError):
pass
experiments, outcomes = results
# Sort factors by importance
inds_mu = mu_df1['mu_star'].values
factors_sorted = np.argsort(inds_mu)[::-1]
# Set up DataFrame of default values to use for experiment
nsamples = len(experiments.index)
defaultvalues = df_unc2['Reference'].values
X_defaults = np.tile(defaultvalues,(nsamples, 1))
# Create Set 1 from experiments
exp_T = experiments.drop(['scenario', 'policy', 'model'], axis=1).T.reindex(df_unc2['Uncertainty'])
exp_ordered = exp_T.T
X_Set1 = exp_ordered.values
# Create initial Sets 2 and 3
X_Set2 = np.copy(X_defaults)
X_Set3 = np.copy(X_Set1)
return df_unc2, factors_sorted, X_defaults, X_Set1, X_Set2, X_Set3, outcomes, mu_df
```
## Where to draw the line between important and not important?
```
'''
Modifed from Waterprogramming blog by Antonia Hadgimichael: https://github.com/antonia-had/SA_verification
The idea is that we create 2 additiopnal Sets (current SA samples are Set 1).
We create a Set 2, using only the T most important factors from our Set 1 sample,
and fixing all other factors to their default values.
We also create a Set 3, now fixing the T most important factors to defaults
and using the sampled values of all other factors from Set 1.
If we classified our important and unimportant factors correctly,
then the correlation coefficient between the model outputs of Set 2 and Set 1 should approximate 1
(since we’re fixing all factors that don’t matter),
and the correlation coefficient between outputs from Set 3 and Set 1 should approximate 0
(since the factors we sampled are inconsequential to the output).
'''
# Define a function to convert your Set 2 and Set 3 into experiments structure in the EMA Workbench
def SA_experiments_to_scenarios(experiments, top_unc, model=None):
'''
"Slighlty modifed from the EMA Workbench"
This function transform a structured experiments array into a list
of Scenarios.
If model is provided, the uncertainties of the model are used.
Otherwise, it is assumed that all non-default columns are
uncertainties.
Parameters
----------
experiments : numpy structured array
a structured array containing experiments
model : ModelInstance, optional
Returns
-------
a list of Scenarios
'''
from ema_workbench import Scenario
# get the names of top 20 uncertainties
uncertainties = list(top_unc['Uncertainty'])
# make list of of tuples of tuples
cases = []
cache = set()
for i in range(experiments.shape[0]):
case = {}
case_tuple = []
for uncertainty in uncertainties:
entry = experiments[uncertainty][i]
case[uncertainty] = entry
case_tuple.append(entry)
case_tuple = tuple(case_tuple)
cases.append(case)
cache.add((case_tuple))
scenarios = [Scenario(**entry) for entry in cases]
return scenarios
```
# Run the models for the top n factors in Set 2 and Set 3 and generate correlation figures
def ranking_verification(outcome_var, df_unc2, factors_sorted, X_defaults, X_Set1, X_Set2, X_Set3, outcomes):
from Model_init import vensimModel
coefficient_S1_S2 = 0
coefficient_S1_S3 = 0.99
n = 0
for f in range(1, len(factors_sorted)+1):
ntopfactors = f
if (coefficient_S1_S2 <=0.95 or coefficient_S1_S3 >= 0.1):
for i in range(ntopfactors): #Loop through all important factors
X_Set2[:,factors_sorted[i]] = X_Set1[:,factors_sorted[i]] #Fix use samples for important
X_Set3[:,factors_sorted[i]] = X_defaults[:,factors_sorted[i]] #Fix important to defaults
X_Set2_exp = pd.DataFrame(data=X_Set2, columns=df_unc2['Uncertainty'].tolist())
X_Set3_exp = pd.DataFrame(data=X_Set3, columns=df_unc2['Uncertainty'].tolist())
scenarios_Set2 = SA_experiments_to_scenarios(X_Set2_exp, df_unc2, model=vensimModel)
scenarios_Set3 = SA_experiments_to_scenarios(X_Set3_exp, df_unc2, model=vensimModel)
vensimModel.outcomes = [TimeSeriesOutcome(outcome_var)]
with MultiprocessingEvaluator(vensimModel, n_processes=nprocess) as evaluator:
experiments_Set2, outcomes_Set2 = evaluator.perform_experiments(scenarios=scenarios_Set2)
experiments_Set3, outcomes_Set3 = evaluator.perform_experiments(scenarios=scenarios_Set3)
# Calculate coefficients of correlation
data_Set1 = outcomes[outcome_var][:,-1]
data_Set2 = outcomes_Set2[outcome_var][:,-1]
data_Set3 = outcomes_Set3[outcome_var][:,-1]
coefficient_S1_S2 = np.corrcoef(data_Set1,data_Set2)[0][1]
coefficient_S1_S3 = np.corrcoef(data_Set1,data_Set3)[0][1]
# Plot outputs and correlation
fig = plt.figure(figsize=(14,7))
ax1 = fig.add_subplot(1,2,1)
ax1.plot(data_Set1,data_Set1, color='#39566E')
ax1.scatter(data_Set1,data_Set2, color='#8DCCFC')
ax1.set_xlabel("Set 1",fontsize=14)
ax1.set_ylabel("Set 2",fontsize=14)
ax1.tick_params(axis='both', which='major', labelsize=10)
ax1.set_title('Set 1 vs Set 2 - ' + str(f) + ' top factors',fontsize=15)
ax1.text(0.05,0.95,'R= '+"{0:.3f}".format(coefficient_S1_S2),transform = ax1.transAxes,fontsize=16)
ax2 = fig.add_subplot(1,2,2)
ax2.plot(data_Set1,data_Set1, color='#39566E')
ax2.scatter(data_Set1,data_Set3, color='#FFE0D5')
ax2.set_xlabel("Set 1",fontsize=14)
ax2.set_ylabel("Set 3",fontsize=14)
ax2.tick_params(axis='both', which='major', labelsize=10)
ax2.set_title('Set 1 vs Set 3 - ' + str(f) + ' top factors',fontsize=15)
ax2.text(0.05,0.95,'R= '+"{0:.3f}".format(coefficient_S1_S3),transform = ax2.transAxes,fontsize=16)
plt.savefig('{}/{}_{}_topfactors.png'.format(r'C:/Users/moallemie/EM_analysis/Fig/sa_verification', outcome_var, str(f)),bbox_inches='tight')
plt.close()
n += 1
n_factor = n
return fig, n_factor
```
# Run the models for the top n factors in Set 2 and Set 3 and generate correlation figures
def ranking_verification(outcome_var, df_unc2, factors_sorted, X_defaults, X_Set1, X_Set2, X_Set3, outcomes):
from Model_init import vensimModel
coefficient_S1_S2 = 0
coefficient_S1_S3 = 0.99
pd_corr = pd.DataFrame(columns=['Outcome', 'Number_of_top_parameters', 'Correlation_Set1_Set3', 'Correlation_Set1_Set2'])
n = 0
for f in range(1, len(factors_sorted)+1):
ntopfactors = f
if (coefficient_S1_S2 <=0.99 or coefficient_S1_S3 >= 0.5):
for i in range(ntopfactors): #Loop through all important factors
X_Set2[:,factors_sorted[i]] = X_Set1[:,factors_sorted[i]] #Fix use samples for important
X_Set3[:,factors_sorted[i]] = X_defaults[:,factors_sorted[i]] #Fix important to defaults
X_Set2_exp = pd.DataFrame(data=X_Set2, columns=df_unc2['Uncertainty'].tolist())
X_Set3_exp = pd.DataFrame(data=X_Set3, columns=df_unc2['Uncertainty'].tolist())
scenarios_Set2 = SA_experiments_to_scenarios(X_Set2_exp, df_unc2, model=vensimModel)
scenarios_Set3 = SA_experiments_to_scenarios(X_Set3_exp, df_unc2, model=vensimModel)
vensimModel.outcomes = [TimeSeriesOutcome(outcome_var)]
try:
with MultiprocessingEvaluator(vensimModel, n_processes=nprocess) as evaluator:
experiments_Set2, outcomes_Set2 = evaluator.perform_experiments(scenarios=scenarios_Set2)
except (BrokenPipeError, IOError):
pass
try:
with MultiprocessingEvaluator(vensimModel, n_processes=nprocess) as evaluator:
experiments_Set3, outcomes_Set3 = evaluator.perform_experiments(scenarios=scenarios_Set3)
except (BrokenPipeError, IOError):
pass
# Calculate coefficients of correlation
data_Set1 = outcomes[outcome_var][:,-1]
data_Set2 = outcomes_Set2[outcome_var][:,-1]
data_Set3 = outcomes_Set3[outcome_var][:,-1]
coefficient_S1_S2 = np.corrcoef(data_Set1,data_Set2)[0][1]
coefficient_S1_S3 = np.corrcoef(data_Set1,data_Set3)[0][1]
pd_corr.loc[n] = [outcome_var, ntopfactors, coefficient_S1_S3, coefficient_S1_S2]
# Plot outputs and correlation
fig = plt.figure(figsize=(14,7))
ax1 = fig.add_subplot(1,2,1)
ax1.plot(data_Set1,data_Set1, color='#39566E')
ax1.scatter(data_Set1,data_Set2, color='#8DCCFC')
ax1.set_xlabel("Set 1",fontsize=14)
ax1.set_ylabel("Set 2",fontsize=14)
ax1.tick_params(axis='both', which='major', labelsize=10)
ax1.set_title('Set 1 vs Set 2 - ' + str(f) + ' top factors',fontsize=15)
ax1.text(0.05,0.95,'R= '+"{0:.3f}".format(coefficient_S1_S2),transform = ax1.transAxes,fontsize=16)
ax2 = fig.add_subplot(1,2,2)
ax2.plot(data_Set1,data_Set1, color='#39566E')
ax2.scatter(data_Set1,data_Set3, color='#FFE0D5')
ax2.set_xlabel("Set 1",fontsize=14)
ax2.set_ylabel("Set 3",fontsize=14)
ax2.tick_params(axis='both', which='major', labelsize=10)
ax2.set_title('Set 1 vs Set 3 - ' + str(f) + ' top factors',fontsize=15)
ax2.text(0.05,0.95,'R= '+"{0:.3f}".format(coefficient_S1_S3),transform = ax2.transAxes,fontsize=16)
plt.savefig('{}/{}_{}_topfactors.png'.format(r'C:/Users/moallemie/EM_analysis/Fig/sa_verification', outcome_var, str(f)),bbox_inches='tight')
plt.close()
n += 1
n_factor = n
return fig, n_factor, pd_corr
# define the ploting function
def plot_scores(outcome_var, n, sc, t):
mu_df2 = mu_df.set_index('Uncertainty')
mu_df2.sort_values(by=['mu_star'], ascending=True, inplace=True)
sns.set_style('white')
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=(3, 6))
ind = mu_df2.iloc[:,0]
err = mu_df2.iloc[:,1]
ind.plot.barh(xerr=err.values.T,ax=ax, color = ['#D7F0BC']*(20-n)+['#62C890']*n,
ecolor='dimgray', capsize=2, width=.9)
ax.set_ylabel('')
ax.legend().set_visible(False)
ax.set_xlabel('mu_star index', fontsize=12)
ylabels = ax.get_yticklabels()
ylabels = [item.get_text()[:-10] for item in ylabels]
ax.set_yticklabels(ylabels, fontsize=12)
title = outcome_var[:-10]
ax.set_title("{}".format(title), fontsize=12)
plt.suptitle("Target year: "+str(t), y=0.94, fontsize=12)
plt.rcParams["figure.figsize"] = [7.08,7.3]
plt.savefig('{}/Morris_ranking_sorted_{}_sc{}_t{}.png'.format(r'C:/Users/moallemie/EM_analysis/Fig/sa_verification', outcome_var, sc, t), dpi=300, bbox_inches='tight')
return fig
if __name__ == '__main__':
ema_logging.log_to_stderr(ema_logging.INFO)
from ema_workbench import (TimeSeriesOutcome,
perform_experiments,
RealParameter,
CategoricalParameter,
ema_logging,
save_results,
load_results)
from ema_workbench import MultiprocessingEvaluator
out_dir = 'C:/Users/moallemie/EM_analysis/Model/'
df_out = pd.read_excel(out_dir+'ScenarioFramework.xlsx', sheet_name='Outcomes')
outcome_vars = ['Total Population Indicator',
'Total Primary Education Graduates Indicator',
'Total Secondary Education Graduates Indicator',
'Total Tertiary Education Graduates Indicator',
'GWP per Capita Indicator',
'Energy Demand Indicator',
'Solar Energy Production Indicator',
'Wind Energy Production Indicator',
'Biomass Energy Production Indicator',
'Oil Production Indicator',
'Gas Production Indicator',
'Coal Production Indicator',
'Total Croplands Indicator',
'Forest Land Indicator',
'Pasture Land Indicator',
'Urban Industrial Land Indicator',
'Nonenergy Crops Production Indicator',
'Livestock Production Indicator',
'Total CO2 Emissions Indicator',
'CO2 Radiative Forcing Indicator']
import time
start = time.time()
top_factors = {}
pd_corr = pd.DataFrame(columns=['Outcome', 'Number_of_top_parameters', 'Correlation_Set1_Set3', 'Correlation_Set1_Set2'])
for out, outcome_var in enumerate(outcome_vars):
ema_logging.log_to_stderr(ema_logging.INFO)
df_unc2, factors_sorted, X_defaults, X_Set1, X_Set2, X_Set3, outcomes, mu_df = create_sets(outcome_var)
fig, n_factor, pd_corr_var = ranking_verification(outcome_var, df_unc2, factors_sorted, X_defaults, X_Set1, X_Set2, X_Set3, outcomes)
plot_scores(outcome_var, n_factor, sc, t)
plt.close()
top_factors[outcome_var] = n_factor
pd_corr = pd.concat([pd_corr, pd_corr_var])
import json
with open('C:/Users/moallemie/EM_analysis/Fig/sa_verification/note.txt', 'w') as file:
file.write(json.dumps(top_factors))
end = time.time()
print("took {} seconds".format(end-start))
with pd.option_context('display.max_rows', None, 'display.max_columns', None): # more options can be specified also
display(pd_corr)
```
| github_jupyter |
```
!pip install pycocotools
!cp ../input/torchdetection/* .
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import numpy as np
import pandas as pd
import math
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
import sys
# You can write up to 20GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
#my imports
import pydicom as dicom
import pickle
import matplotlib as plt
import matplotlib.patches as patches
import pandas as pd
import numpy as np
import PIL
from PIL import Image
import torch
import transforms as T
import torchvision.transforms as T2
import utils
import torchvision
import tqdm
import warnings
warnings.filterwarnings("ignore")
device = torch.device('cuda') if torch.cuda.is_available() else torch.device('cpu')
csv = pd.read_csv("../input/vinbigdata-1024-image-dataset/vinbigdata/train.csv")
def custom_normalize(img, mean, std, max_pixel_value=255.0):
mean = np.array(mean, dtype=np.float32)
mean *= max_pixel_value
std = np.array(std, dtype=np.float32)
std *= max_pixel_value
denominator = np.reciprocal(std, dtype=np.float32)
img = img.astype(np.float32)
img -= mean
img *= denominator
return img
class ChestXRayDataset(object):
def __init__ (self, path, tfms, csv, weights=None):
#defaults
self.path=path
self.tfms=tfms
self.csv = csv
self.weights = weights
#images
self.imgs = [x+".png" for x in csv['image_id'].tolist()]
def __getitem__ (self, idx):
img_path = os.path.join(self.path, 'train', self.imgs[idx])
img = Image.open(img_path)
# img = np.asarray(img)
# img/np.max(img)
boxes = []
labs = []
ic = []
dk = self.csv.loc[self.csv['image_id']==self.imgs[idx][:-4]]
area = []
for idx, row in dk.iterrows():
if (int(row['class_id'])!=14):
labs.append(int(row['class_id'])+1)
ic.append(0)
xmin = float(row['x_min'])
xmax = float(row['x_max'])
ymin = float(row['y_min'])
ymax = float(row['y_max'])
xmin*=1024
xmin/=row['width']
xmax*=1024
xmax/=row['width']
ymin*=1024
ymin/=row['height']
ymax*=1024
ymax/=row['height']
# if (img.shape[0]>img.shape[1]):
# rat = 800/img.shape[0]
# xmin = rat*xmin
# xmax = rat*xmax
# ymin = rat*ymin
# ymax = rat*ymax
# else:
# rat = 800/img.shape[1]
# xmin = rat*xmin
# xmax = rat*xmax
# ymin = rat*ymin
# ymax = rat*ymax
# xmin = floor(xmin)
# ymin = floor(ymin)
# xmax = floor(xmax)
# ymax = floor(ymax)
boxes.append([xmin, ymin, xmax, ymax])
area.append((xmax-xmin)*(ymax-ymin))
if (len(boxes)==0):
boxes = [[0, 0, 1, 1]]
boxes = torch.as_tensor(boxes, dtype=torch.float32)
labels = torch.zeros((1,), dtype=torch.int64)
area = torch.ones((1,), dtype=torch.float32)
iscrowd = torch.as_tensor([0],dtype=torch.int64)
else:
boxes = torch.as_tensor(boxes, dtype=torch.float32)
labels = torch.as_tensor(labs, dtype=torch.int64)
area = torch.as_tensor(area, dtype=torch.float32)
iscrowd = torch.as_tensor(ic, dtype=torch.int64)
image_id = torch.tensor([idx])
target = {}
target["boxes"] = boxes
target["labels"] = labels
target["image_id"] = image_id
target["area"] = area
target["iscrowd"] = iscrowd
if self.tfms is not None:
# img = custom_normalize(img, (0,0,0), (1,1,1))
# img = Image.fromarray(img)
img = self.tfms(img)
return img, target
def __len__(self):
return len(self.imgs)
def get_transform():
transform = []
transform.append(T2.ToTensor())
return T2.Compose(transform)
file_train = open(f"../input/private-model-save-dataset/train-set-v3 (1).pkl", "rb")
ds_train = pickle.load(file_train)
ds_train.head()
csv.head()
ds_test = pd.merge(csv,ds_train, indicator=True, how='outer').query('_merge=="left_only"').drop('_merge', axis=1)
ds_test[ds_test['image_id']=='afb6230703512afc370f236e8fe98806']
test_imgs = ds_test['image_id'].unique()
test_imgs
def bb_intersection_over_union(boxA, boxB):
# determine the (x, y)-coordinates of the intersection rectangle
xA = max(boxA[0], boxB[0])
yA = max(boxA[1], boxB[1])
xB = min(boxA[2], boxB[2])
yB = min(boxA[3], boxB[3])
# compute the area of intersection rectangle
interArea = abs(max((xB - xA, 0)) * max((yB - yA), 0))
if interArea == 0:
return 0
# compute the area of both the prediction and ground-truth
# rectangles
boxAArea = abs((boxA[2] - boxA[0]) * (boxA[3] - boxA[1]))
boxBArea = abs((boxB[2] - boxB[0]) * (boxB[3] - boxB[1]))
# compute the intersection over union by taking the intersection
# area and dividing it by the sum of prediction + ground-truth
# areas - the interesection area
iou = interArea / float(boxAArea + boxBArea - interArea)
# return the intersection over union value
return iou
for i in test_imgs:
df = ds_test.loc[ds_test['image_id']==i]
i = 0
while (i<len(df)):
j = i+1
while (j<len(df)):
if bb_intersection_over_union([df.iloc[i]['x_min'], df.iloc[i]['y_min'], df.iloc[i]['x_max'], df.iloc[i]['y_max']],
[df.iloc[j]['x_min'], df.iloc[j]['y_min'], df.iloc[j]['x_max'], df.iloc[j]['y_max']])>=0.4:
if (df.iloc[i]['class_id']==df.iloc[j]['class_id']):
try:
ds_test.drop(df.index[j], inplace=True)
except:
pass
j+=1
i+=1
# df.iloc[0]
def intersect(boxa, boxb):
minx = max(boxa[0], boxb[0])
miny = max(boxa[1], boxb[1])
maxx = min(boxa[2], boxb[2])
maxy = min(boxa[3], boxb[3])
return [minx, miny, maxx, maxy]
def box_area(box):
return (box[2]-box[0])*(box[3]-box[1])
def iou(box_a, box_b):
box_i = intersect(box_a, box_b)
box_i_area = box_area(box_i)
box_a_area = box_area(box_a)
box_b_area = box_area(box_b)
u_area = box_a_area+box_b_area-box_i_area
return (box_i_area/u_area, box_i_area, u_area)
def score_ap(model,dataloader,threshold=0.7):
model.eval()
tp_list = []
target_labels_list = []
scores_list = []
labels_list = []
iou_list = []
size = 0
print_threshold = 200
with torch.no_grad():
for index, (imgs,targets) in enumerate(dataloader):
imgs = list(img.to(device) for img in imgs)
predictions = model(imgs)
size += index
for i, prediction in enumerate(predictions):
img = imgs[i]
target = targets[i]
boxes = []
scores = []
labels = []
for i,p in enumerate(prediction["scores"]):
if p>threshold:
boxes.append(prediction["boxes"][i].cpu().numpy())
scores.append(prediction["scores"][i].cpu().item())
labels.append(prediction["labels"][i].cpu().item())
labels_np = np.array(labels)
scores_np = np.array(scores)
tp, labels, scores, iou = compute_tp(target["boxes"].cpu().numpy(),
target["labels"].cpu().numpy(),
boxes, labels, labels_np, scores_np, iou_threshold=0.5)
scores_list.extend(scores)
labels_list.extend(labels)
target_labels_list.extend(target["labels"])
tp_list.extend(tp)
iou_list.extend(iou)
p, r, ap, f1, unique_labels = ap_per_class(
np.asarray(tp_list),
np.asarray(scores_list),
np.asarray(labels_list),
np.asarray(target_labels_list))
m_ap = np.mean(ap)
return m_ap
def ap_per_class(tp, conf, pred_cls, target_cls):
""" Compute the average precision, given the recall and precision curves.
Source: https://github.com/rafaelpadilla/Object-Detection-Metrics.
# Arguments
tp: True positives (list).
conf: Objectness value from 0-1 (list).
pred_cls: Predicted object classes (list).
target_cls: True object classes (list).
# Returns
The average precision as computed in py-faster-rcnn.
"""
# Sort by objectness
i = np.argsort(-conf)
tp, conf, pred_cls = tp[i], conf[i], pred_cls[i]
# Find unique classes
unique_classes = np.unique(target_cls)
# Create Precision-Recall curve and compute AP for each class
ap, p, r = [], [], []
for c in tqdm.tqdm(unique_classes, desc="Computing AP"):
i = pred_cls == c
n_gt = (target_cls == c).sum() # Number of ground truth objects
n_p = i.sum() # Number of predicted objects
if n_p == 0 and n_gt == 0:
continue
elif n_p == 0 or n_gt == 0:
ap.append(0)
r.append(0)
p.append(0)
else:
# Accumulate FPs and TPs
fpc = (1 - tp[i]).cumsum()
tpc = (tp[i]).cumsum()
# Recall
recall_curve = tpc / (n_gt + 1e-16)
r.append(recall_curve[-1])
# Precision
precision_curve = tpc / (tpc + fpc)
p.append(precision_curve[-1])
# AP from recall-precision curve
ap.append(compute_ap(recall_curve, precision_curve))
# Compute F1 score (harmonic mean of precision and recall)
p, r, ap = np.array(p), np.array(r), np.array(ap)
f1 = 2 * p * r / (p + r + 1e-16)
return p, r, ap, f1, unique_classes.astype("int32")
def compute_ap(recall, precision):
""" Compute the average precision, given the recall and precision curves.
Code originally from https://github.com/rbgirshick/py-faster-rcnn.
# Arguments
recall: The recall curve (list).
precision: The precision curve (list).
# Returns
The average precision as computed in py-faster-rcnn.
"""
# correct AP calculation
# first append sentinel values at the end
mrec = np.concatenate(([0.0], recall, [1.0]))
mpre = np.concatenate(([0.0], precision, [0.0]))
# compute the precision envelope
for i in range(mpre.size - 1, 0, -1):
mpre[i - 1] = np.maximum(mpre[i - 1], mpre[i])
# to calculate area under PR curve, look for points
# where X axis (recall) changes value
i = np.where(mrec[1:] != mrec[:-1])[0]
# and sum (\Delta recall) * prec
ap = np.sum((mrec[i + 1] - mrec[i]) * mpre[i + 1])
return ap
def compute_tp(target_boxes, target_labels, pred_boxes, pred_labels, pred_cls, conf, iou_threshold=0.5, print_fp=False):
'''
calculate true positives, align pred labels and conf scores with true positives indexes, IoUs of bounding boxes
Args:
target_boxes: REQUIRED, array of target boxes
target_labels: REQUIRED, array of target labels
pred_boxes: REQUIRED, array of prediction boxes
pred_labels: REQUIRED, array of prediction labels
pred_cls: REQUIRED, array of predicted object classes
conf: REQUIRED, array of objectness value ranging from 0-1
iou_threshold: OPTIONAL, float, threshold of min iou, default - iou_threshold=0.5
Return value: array of true positives, new_pred_cls, new_conf, iou_return_arr
'''
avg_iou = 0
# all three will be set to length of target_labels
true_positives = np.zeros(len(target_labels))
new_pred_cls = np.full(len(target_labels), -1)
new_conf = np.zeros(len(target_labels))
false_positives = 0
fp_array = []
detected_boxes = []
iou_return_arr = np.zeros(len(target_labels))
for pred_i, pred_label in enumerate(pred_labels):
if pred_label not in target_labels:
fp_array.append(pred_label)
false_positives += 1
continue
pred_box = np.array([pred_boxes[pred_i][0],
pred_boxes[pred_i][1],
pred_boxes[pred_i][2],
pred_boxes[pred_i][3]])
iou_list=bbox_iou(np.expand_dims(pred_box,0), target_boxes) # calculate iou between prediction and target box
box_index = np.argmax(iou_list)
iou = iou_list[box_index]
if iou >= iou_threshold and box_index not in detected_boxes and pred_i < len(true_positives):
if pred_label != target_labels[box_index]:
print(f"pred_label: {pred_label}, box_index,{box_index}, target_labels:{target_labels}, detected_boxes: {detected_boxes}")
else:
true_positives[pred_i] = 1
new_pred_cls[pred_i] = pred_cls[pred_i]
new_conf[pred_i] = conf[pred_i]
iou_return_arr[pred_i] = iou
detected_boxes += [box_index]
if (print_fp):
print(fp_array)
return true_positives, new_pred_cls, new_conf, iou_return_arr
def bbox_iou(box1, box2):
"""
Returns the IoU of two bounding boxes
"""
# Get the coordinates of bounding boxes
b1_x1, b1_y1, b1_x2, b1_y2 = box1[:, 0], box1[:, 1], box1[:, 2], box1[:, 3]
b2_x1, b2_y1, b2_x2, b2_y2 = box2[:, 0], box2[:, 1], box2[:, 2], box2[:, 3]
# get the corrdinates of the intersection rectangle
inter_rect_x1 = np.maximum(b1_x1, b2_x1)
inter_rect_y1 = np.maximum(b1_y1, b2_y1)
inter_rect_x2 = np.minimum(b1_x2, b2_x2)
inter_rect_y2 = np.minimum(b1_y2, b2_y2)
# Intersection area
inter_area = np.clip(inter_rect_x2 - inter_rect_x1 + 1, 0, None) * np.clip(
inter_rect_y2 - inter_rect_y1 + 1, 0, None
)
# Union Area
b1_area = (b1_x2 - b1_x1 + 1) * (b1_y2 - b1_y1 + 1)
b2_area = (b2_x2 - b2_x1 + 1) * (b2_y2 - b2_y1 + 1)
iou = inter_area / (b1_area + b2_area - inter_area + 1e-16)
return iou
def main ():
model = torch.load("../input/private-model-save-dataset/model-save-5.pkl")
model.eval()
dataset_test = ChestXRayDataset('../input/vinbigdata-1024-image-dataset/vinbigdata', get_transform(), ds_test)
data_loader_test = torch.utils.data.DataLoader(
dataset_test, batch_size=1, shuffle=False, num_workers=4,
collate_fn=utils.collate_fn)
model.to(device)
m_ap = score_ap(model, data_loader_test, threshold=0.9)
print(m_ap)
main()
```
| github_jupyter |
```
#take shannon matlab & put into netcdf
from scipy.io import loadmat
import h5py
import xarray as xr
import numpy as np
#PLOTTING
import cartopy
from mpl_toolkits.axes_grid1.inset_locator import inset_axes
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
from matplotlib.colorbar import Colorbar
import matplotlib.ticker as mticker
import cartopy.crs as ccrs
import cartopy.feature as cfeature
from cartopy.mpl.gridliner import LONGITUDE_FORMATTER, LATITUDE_FORMATTER
#resampling to grid
from pyresample.geometry import AreaDefinition
from pyresample.geometry import GridDefinition
from pyresample import image, geometry, load_area, save_quicklook, SwathDefinition, area_def2basemap
from pyresample.kd_tree import resample_nearest
from pyresample.utils import check_and_wrap
from scipy import spatial
import glob
#where to find the data
adir_data= 'f:/data/project_data/fluxsat/orbit/'
adir_figs= 'f:/data/project_data/fluxsat/figures/'
# add land mask
#get bathymetry from ETOPO1
fname_topo = 'F:/data/topo/ETOPO1_Ice_g_gmt4.grd'
ds = xr.open_dataset(fname_topo)
ds_topo = ds.rename_dims({'x':'lon','y':'lat'}).rename({'x':'lon','y':'lat'})
tem = ds_topo.attrs
ds_topo = ds_topo.rename({'z':'etopo_depth'})
ds_topo.etopo_depth.attrs=tem
_, index = np.unique(ds_topo['lon'], return_index=True)
ds_topo = ds_topo.isel(lon=index)
_, index = np.unique(ds_topo['lat'], return_index=True)
ds_topo = ds_topo.isel(lat=index)
#get all filenames
filenames = glob.glob(adir_data+'FS*.mat')
filenames
#read in data from all 3 files, put into xarray dataset
#change UTCtime from days since 0000/1/1 to datetime64 object
#make utctime the coordinate
ds_sv=[]
for fname in filenames:
f = h5py.File(fname,mode='r') #open file
struArray = f['rad']
ilen = struArray['SClat']['data'].shape[0]
da = xr.DataArray(None, coords=[np.arange(ilen)], dims=['iob'])
ds = xr.Dataset({'test':xr.DataArray(None, coords=[np.arange(ilen)], dims=['iob'])})
for v in struArray:
data = struArray[v]['data']
da = xr.DataArray(data[:,0], coords=[np.arange(ilen)], dims=['iob'])
ds[v]=da
ds['UTCtime'] = np.datetime64('0000-01-01')+ds.UTCtime.astype('timedelta64[D]')+((ds.UTCtime-ds.UTCtime[0])*24*60*60*1e9).astype('timedelta64[ns]')
ds = ds.drop('test')
ds = ds.swap_dims({'iob':'UTCtime'}).reset_coords()
ds_sv.append(ds)
ds = xr.concat(ds_sv, dim='UTCtime')
#ds['UTCtime']=np.datetime64('0000-01-01')+ds.UTCtime.astype('int').data.astype('timedelta64[D]')
#create 1 day of data dataarray in xarray
dylist = ['2018-11-28','2018-11-29','2018-11-30']
da_sv = []
for dnum,dstr in enumerate(dylist):
dy = ds.sel(UTCtime=slice(dylist[dnum],dylist[dnum]))
for iorbit in range(16):
i1=iorbit*2777296
i2=(iorbit+1)*2777296
i2= [dy.blon.size-1 if i2>dy.blon.size else i2]
i2=i2[0]
dy2 = dy.sel(UTCtime=slice(dy.UTCtime[i1],dy.UTCtime[i2]))
x = dy2.blon.data
y = dy2.blat.data
z = dy2.blat.data*0+1
lons,lats,data = x,y,z
lons,lats = check_and_wrap(lons, lats)
swath_def = SwathDefinition(lons, lats)
grid_def_lons, grid_def_lats = np.arange(-180,180,0.1), np.arange(-90,90,0.1)
grid_mesh_lons,grid_mesh_lats = np.meshgrid(grid_def_lons,grid_def_lats)
grid_def = GridDefinition(lons=grid_mesh_lons,lats=grid_mesh_lats)
result1 = resample_nearest(swath_def, data, grid_def, radius_of_influence=20000, fill_value=None)
rr=np.expand_dims(result1,axis=2)
iorbit_num = iorbit+dnum*16
da = xr.DataArray(rr,name='sec',
coords={'lat':grid_def_lats,'lon':grid_def_lons,'orbit':[iorbit_num]},
dims=('lat','lon','orbit'))
da_sv.append(da)
da2 = xr.concat(da_sv, dim='orbit')
#ds = ds.drop({'SCinc','iob','SCalt','belev','bhorz','binc','scanang','SClat','SClon'})
fname_out = adir_data + '3dys_butterfly_obs.nc'
ds.to_netcdf(fname_out)
fname_out = adir_data + '3dys_butterfly_grid.nc'
da2.to_netcdf(fname_out)
#breakup
fname_out = adir_data + '3dys_butterfly_grid.nc'
ds = xr.open_dataset(fname_out)
fname_out = adir_data + '3dys_butterfly_gridA.nc'
ds.sec[:,:,0:16].to_netcdf(fname_out,encoding={"sec": {"dtype": "int8", "scale_factor": 0.1, "zlib": True, "complevel": 9}})
fname_out = adir_data + '3dys_butterfly_gridB.nc'
ds.sec[:,:,16:32].to_netcdf(fname_out,encoding={"sec": {"dtype": "int8", "scale_factor": 0.1, "zlib": True, "complevel": 9}})
fname_out = adir_data + '3dys_butterfly_gridC.nc'
ds.sec[:,:,32:].to_netcdf(fname_out,encoding={"sec": {"dtype": "int8", "scale_factor": 0.1, "zlib": True, "complevel": 9}})
tem = xr.open_dataset(fname_out)
tem.close()
tem
tem.sec[:,:,0].plot()
# CAC data
from scipy.io import loadmat
#load data
x = loadmat(adir_data + 'clayson_fluxes.mat')
#put into xarray dataarray
da_lo = xr.DataArray(x['lhf1deg'],
coords={'latlo':x['lat1deg'][0],'lonlo':x['lon1deg'][0]},
dims=['lonlo','latlo']).T
da_hi = xr.DataArray(x['lhfhires'],
coords={'lat':x['latitude'][:,0],'lon':x['longitude'][:,0]},
dims=['lon','lat']).T
#put into xarray dataset & write to netcdf file
ds = xr.Dataset({'lo':da_lo,'hi':da_hi})
ds.to_netcdf(adir_data + 'clayson_fluxes.nc')
da_hi.plot()
# shannons different resolution fluxes
#fname = adir_data + 'shannon_flux_res.mat'
# CAC data
from scipy.io import loadmat
#load data
x = loadmat(adir_data + 'shannon_flux_res.mat')
lat1 = x['XI'][1,:]
lon1 = x['YI'][:,1]
hi = x['hires']
lo = x['lores']
#put into xarray dataarray
da_lo = xr.DataArray(lo,
coords={'lat':lat1,'lon':lon1,'res':[0,1,2,3]},
dims=['lon','lat','res']).T
da_hi = xr.DataArray(hi,
coords={'lat':lat1,'lon':lon1,'res':[0,1,2]},
dims=['lon','lat','res']).T
#put into xarray dataset & write to netcdf file
ds = xr.Dataset({'lo':da_lo,'hi':da_hi})
ds.to_netcdf(adir_data + 'clayson_fluxes_hilo.nc')
ds.hi[0,:,:].sel(lon=slice(120,180),lat=slice(10,50)).plot(cmap='jet')
ds.hi[2,:,:].sel(lon=slice(120,180),lat=slice(10,50)).plot(cmap='jet')
```
| github_jupyter |
# Initialize the environment
```
from google.colab import drive
drive.mount('/content/drive/')
import os
os.chdir("/content/drive/MyDrive/Hate Speech Detection/ABSA-PyTorch/")
# fix the problem: ImportError: cannot import name 'SAVE_STATE_WARNING' from 'torch.optim.lr_scheduler' (/usr/local/lib/python3.7/dist-packages/torch/optim/lr_scheduler.py)
!pip install torch==1.4.0
!pip install -r requirements.txt
```
# Apply the model
```
# !python dependency_graph.py
```
## All other English based models BERT SPC
```
!python train-test.py --model_name bert_spc --dataset 2016-dataset --num_epoch 10 --device cuda:0 --polarities_dim 3 --valset_ratio 0.25 --pretrained_bert_name bert-large-cased --bert_dim 1024
!python train-test.py --model_name bert_spc --dataset 2016-dataset --num_epoch 10 --device cuda:0 --polarities_dim 3 --valset_ratio 0.25 --pretrained_bert_name bert-large-uncased --bert_dim 1024
!python train-test.py --model_name bert_spc --dataset 2016-dataset --num_epoch 10 --device cuda:0 --polarities_dim 3 --valset_ratio 0.25 --pretrained_bert_name bert-base-cased-finetuned-mrpc --bert_dim 768
!python train-test.py --model_name bert_spc --dataset 2016-dataset --num_epoch 10 --device cuda:0 --polarities_dim 3 --valset_ratio 0.25 --pretrained_bert_name bert-large-cased-whole-word-masking-finetuned-squad --bert_dim 1024
!python train-test.py --model_name bert_spc --dataset 2016-dataset --num_epoch 10 --device cuda:0 --polarities_dim 3 --valset_ratio 0.25 --pretrained_bert_name bert-large-uncased-whole-word-masking-finetuned-squad --bert_dim 1024
!python train-test.py --model_name bert_spc --dataset 2016-dataset --num_epoch 10 --device cuda:0 --polarities_dim 3 --valset_ratio 0.25 --pretrained_bert_name bert-large-uncased-whole-word-masking --bert_dim 1024
!python train-test.py --model_name bert_spc --dataset 2016-dataset --num_epoch 10 --device cuda:0 --polarities_dim 3 --valset_ratio 0.25 --pretrained_bert_name bert-large-cased-whole-word-masking --bert_dim 1024
```
## BERT uncased AEN
```
!python train-test.py --model_name aen_bert --dataset 2016-dataset --num_epoch 10 --device cuda:0 --polarities_dim 3 --valset_ratio 0.25 --pretrained_bert_name bert-large-uncased --bert_dim 1024
!python train-test.py --model_name aen_bert --dataset 2016-dataset --num_epoch 10 --device cuda:0 --polarities_dim 3 --valset_ratio 0.25 --pretrained_bert_name bert-large-uncased-whole-word-masking-finetuned-squad --bert_dim 1024
!python train-test.py --model_name aen_bert --dataset 2016-dataset --num_epoch 10 --device cuda:0 --polarities_dim 3 --valset_ratio 0.25 --pretrained_bert_name bert-large-uncased-whole-word-masking --bert_dim 1024
!python train-test.py --model_name aen_bert --dataset 2016-dataset --num_epoch 10 --device cuda:0 --polarities_dim 3 --valset_ratio 0.25 --pretrained_bert_name bert-base-cased-finetuned-mrpc --bert_dim 768
```
| github_jupyter |
[](http://rpi.analyticsdojo.com)
<center><h1>Introduction to Spark</h1></center>
<center><h3><a href = 'http://rpi.analyticsdojo.com'>rpi.analyticsdojo.com</a></h3></center>
Adopted from work by Steve Phelps:
https://github.com/phelps-sg/python-bigdata
This work is licensed under the Creative Commons Attribution 4.0 International license agreement.
### Reference
- [Spark Documentation](http://spark.apache.org/docs/latest/)
- [Spark Programming Guide](http://spark.apache.org/docs/latest/programming-guide.html)
- [DataBricks Login](https://community.cloud.databricks.com)
- [Pyspark](https://github.com/jupyter/docker-stacks)
- [Conda](
```conda install -c anaconda-cluster/label/dev spark
conda install -c conda-forge pyspark
```
### Overview
- History
- Data Structures
- Using Apache Spark with Python
## History
- Apache Spark was first released in 2014.
- It was originally developed by [Matei Zaharia](http://people.csail.mit.edu/matei) as a class project, and later a PhD dissertation, at University of California, Berkeley.
- In contrast to Hadoop, Apache Spark:
- is easy to install and configure.
- provides a much more natural *iterative* workflow
## Resilient Distributed Datasets (RDD)
- The fundamental abstraction of Apache Spark is a read-only, parallel, distributed, fault-tolerent collection called a resilient distributed datasets (RDD).
- When working with Apache Spark we iteratively apply functions to every elelement of these collections in parallel to produce *new* RDDs.
- For the most part, you can think/use RDDs like distributed dataframes.
## Resilient Distributed Datasets (RDD)
- Properties resilient distributed datasets (RDDs):
- The data is distributed across nodes in a cluster of computers.
- No data is lost if a single node fails.
- Data is typically stored in HBase tables, or HDFS files.
- The `map` and `reduce` functions can work in *parallel* across
different keys, or different elements of the collection.
- The underlying framework (e.g. Hadoop or Apache Spark) allocates data and processing to different nodes, without any intervention from the programmer.
## Word Count Example
- In this simple example, the input is a set of URLs, each record is a document. <br> <br> <br>
- **Problem: Compute how many times each word has occurred across data set.**
## Word Count: Map
The input to $\operatorname{map}$ is a mapping:
- Key: URL
- Value: Contents of document <br>
$\left< document1, to \; be \; or \; not \; to \; be \right>$
- In this example, our $\operatorname{map}$ function will process a given URL, and produces a mapping:
- So our original data-set will be transformed to:
$\left< to, 1 \right>$
$\left< be, 1 \right>$
$\left< or, 1 \right>$
$\left< not, 1 \right>$
$\left< to, 1 \right>$
$\left< be, 1 \right>$
## Word Count: Reduce
- The reduce operation groups values according to their key, and then performs areduce on each key.
- The collections are partitioned across different storage units, therefore.
- Map-Reduce will fold the data in such a way that it minimises data-copying across the cluster.
- Data in different partitions are reduced separately in parallel.
- The final result is a reduce of the reduced data in each partition.
- Therefore it is very important that our operator *is both commutative and associative*.
- In our case the function is the `+` operator
$\left< be, 2 \right>$
$\left< not, 1 \right>$
$\left< or, 1 \right>$
$\left< to, 2 \right>$
## Map-Reduce on a Cluster of Computers
- The code we have written so far will *not* allow us to exploit parallelism from multiple computers in a [cluster](https://en.wikipedia.org/wiki/Computer_cluster).
- Developing such a framework would be a very large software engineering project.
- There are existing frameworks we can use:
- [Apache Hadoop](https://hadoop.apache.org/)
- [Apache Spark](https://spark.apache.org/)
- This notebook covers Apache Spark.
## Apache Spark
- Apache Spark provides an object-oriented library for processing data on the cluster.
- It provides objects which represent resilient distributed datasets (RDDs).
- RDDs behave a bit like Python collections (e.g. lists).
- However:
- the underlying data is distributed across the nodes in the cluster, and
- the collections are *immutable*.
## Apache Spark and Map-Reduce
- We process the data by using higher-order functions to map RDDs onto *new* RDDs.
- Each instance of an RDD has at least two *methods* corresponding to the Map-Reduce workflow:
- `map`
- `reduceByKey`
- These methods work in the same way as the corresponding functions we defined earlier to work with the standard Python collections.
- There are also additional RDD methods in the Apache Spark API including ones for SQL.
## Word-count in Apache Spark
```
words = "to be or not to be".split()
words
```
### The `SparkContext` class
- When working with Apache Spark we invoke methods on an object which is an instance of the `pyspark.context.SparkContext` context.
- Typically, (such as when running on DataBricks) an instance of this object will be created automatically for you and assigned to the variable `sc`.
- The `parallelize` method in `SparkContext` can be used to turn any ordinary Python collection into an RDD;
```
#Don't Execute this on Databricks
#To be used if executing via docker
import pyspark
#sc = pyspark.SparkContext('local[*]')
words_rdd = sc.parallelize(words)
words_rdd
```
### Mapping an RDD
- Now when we invoke the `map` or `reduceByKey` methods on `my_rdd` we can set up a parallel processing computation across the cluster.
```
word_tuples_rdd = words_rdd.map(lambda x: (x, 1))
word_tuples_rdd
```
### Collecting the RDD
- Notice that we do not have a result yet.
- The computation is not performed until we request the final result to be *collected*.
- We do this by invoking the `collect()` method.
- Be careful with the `collect` method, as all data you are collecting must fit in memory.
- The `take` method is similar to `collect`, but only returns the first $n$ elements.
```
word_tuples_rdd.collect()
word_tuples_rdd.take(4)
```
### Reducing an RDD
- However, we require additional processing to reduce the data using the word key.
```
word_counts_rdd = word_tuples_rdd.reduceByKey(lambda x, y: x + y)
word_counts_rdd
```
- Now we request the final result:
```
word_counts = word_counts_rdd.collect()
word_counts
```
### Lazy evaluation
- It is only when we invoke `collect()` that the processing is performed on the cluster.
- Invoking `collect()` will cause both the `map` and `reduceByKey` operations to be performed.
- If the resulting collection is very large then this can be an expensive operation.
```
word_counts_rdd.take(2)
```
### Connecting MapReduce in Single Command
- Can string together `map` and `reduce` commands.
- Not executed until it is collected.
```
text = "to be or not to be".split()
rdd = sc.parallelize(text)
counts = rdd.map(lambda word: (word, 1)).reduceByKey(lambda x, y: x + y)
counts.collect()
```
## Additional RDD transformations
- Apache Spark offers many more methods for operating on collections of tuples over and above the standard Map-Reduce framework:
- Sorting: `sortByKey`, `sortBy`, `takeOrdered`
- Mapping: `flatMap`
- Filtering: `filter`
- Counting: `count`
- Set-theoretic: `intersection`, `union`
- Many others: [see the Transformations section of the programming guide](https://spark.apache.org/docs/latest/programming-guide.html#transformations)
## Creating an RDD from a text file
- In the previous example, we created an RDD from a Python collection.
- This is *not* typically how we would work with big data.
- More commonly we would create an RDD corresponding to data in an
HBase table, or an HDFS file.
- The following example creates an RDD from a text file on the native filesystem (ext4);
- With bigger data, you would use an HDFS file, but the principle is the same.
- Each element of the RDD corresponds to a single *line* of text.
```
genome = sc.textFile('../input/iris.csv')
```
## Calculating $\pi$ using Spark
- We can estimate an approximate value for $\pi$ using the following Monte-Carlo method:
1. Inscribe a circle in a square
2. Randomly generate points in the square
3. Determine the number of points in the square that are also in the circle
4. Let $r$ be the number of points in the circle divided by the number of points in the square, then $\pi \approx 4 r$.
- Note that the more points generated, the better the approximation
See [this tutorial](https://computing.llnl.gov/tutorials/parallel_comp/#ExamplesPI).
```
import numpy as np
def sample(p):
#here x,y are the x,y coordinate
x, y = np.random.random(), np.random.random()
#Because the circle is of
return 1 if x*x + y*y < 1 else 0
NUM_SAMPLES = 1000000
count = sc.parallelize(range(0, NUM_SAMPLES)).map(sample) \
.reduce(lambda a, b: a + b)
#Area = 4*PI*r
r = float(count) / float(NUM_SAMPLES)
r
print ("Pi is approximately %f" % (4.0 * r))
```
| github_jupyter |
## Face and Facial Keypoint detection
After you've trained a neural network to detect facial keypoints, you can then apply this network to *any* image that includes faces. The neural network expects a Tensor of a certain size as input and, so, to detect any face, you'll first have to do some pre-processing.
1. Detect all the faces in an image using a face detector (we'll be using a Haar Cascade detector in this notebook).
2. Pre-process those face images so that they are grayscale, and transformed to a Tensor of the input size that your net expects. This step will be similar to the `data_transform` you created and applied in Notebook 2, whose job was tp rescale, normalize, and turn any iimage into a Tensor to be accepted as input to your CNN.
3. Use your trained model to detect facial keypoints on the image.
---
In the next python cell we load in required libraries for this section of the project.
```
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
%matplotlib inline
```
#### Select an image
Select an image to perform facial keypoint detection on; you can select any image of faces in the `images/` directory.
```
import cv2
# load in color image for face detection
image = cv2.imread('images/obamas.jpg')
# switch red and blue color channels
# --> by default OpenCV assumes BLUE comes first, not RED as in many images
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
# plot the image
fig = plt.figure(figsize=(9,9))
plt.imshow(image)
```
## Detect all faces in an image
Next, you'll use one of OpenCV's pre-trained Haar Cascade classifiers, all of which can be found in the `detector_architectures/` directory, to find any faces in your selected image.
In the code below, we loop over each face in the original image and draw a red square on each face (in a copy of the original image, so as not to modify the original). You can even [add eye detections](https://docs.opencv.org/3.4.1/d7/d8b/tutorial_py_face_detection.html) as an *optional* exercise in using Haar detectors.
An example of face detection on a variety of images is shown below.
<img src='images/haar_cascade_ex.png' width=80% height=80%/>
```
# load in a haar cascade classifier for detecting frontal faces
face_cascade = cv2.CascadeClassifier('detector_architectures/haarcascade_frontalface_default.xml')
# run the detector
# the output here is an array of detections; the corners of each detection box
# if necessary, modify these parameters until you successfully identify every face in a given image
faces = face_cascade.detectMultiScale(image, 1.2, 2)
# make a copy of the original image to plot detections on
image_with_detections = image.copy()
# loop over the detected faces, mark the image where each face is found
for (x,y,w,h) in faces:
# draw a rectangle around each detected face
# you may also need to change the width of the rectangle drawn depending on image resolution
cv2.rectangle(image_with_detections,(x,y),(x+w,y+h),(255,0,0),3)
fig = plt.figure(figsize=(9,9))
plt.imshow(image_with_detections)
```
## Loading in a trained model
Once you have an image to work with (and, again, you can select any image of faces in the `images/` directory), the next step is to pre-process that image and feed it into your CNN facial keypoint detector.
First, load your best model by its filename.
```
import torch
from models import Net
n_outputs = 136 # Number of outputs of the CNN
input_shape=(224, 224)
net = Net(input_shape=input_shape, output_len=n_outputs, dropout_p=0.2, n_blocks=5)
## TODO: load the best saved model parameters (by your path name)
## You'll need to un-comment the line below and add the correct name for *your* saved model
net.load_state_dict(torch.load('saved_models/model_chpt.pt'))
## print out your net and prepare it for testing (uncomment the line below)
net.eval()
```
## Keypoint detection
Now, we'll loop over each detected face in an image (again!) only this time, you'll transform those faces in Tensors that your CNN can accept as input images.
### TODO: Transform each detected face into an input Tensor
You'll need to perform the following steps for each detected face:
1. Convert the face from RGB to grayscale
2. Normalize the grayscale image so that its color range falls in [0,1] instead of [0,255]
3. Rescale the detected face to be the expected square size for your CNN (224x224, suggested)
4. Reshape the numpy image into a torch image.
You may find it useful to consult to transformation code in `data_load.py` to help you perform these processing steps.
### TODO: Detect and display the predicted keypoints
After each face has been appropriately converted into an input Tensor for your network to see as input, you'll wrap that Tensor in a Variable() and can apply your `net` to each face. The ouput should be the predicted the facial keypoints. These keypoints will need to be "un-normalized" for display, and you may find it helpful to write a helper function like `show_keypoints`. You should end up with an image like the following with facial keypoints that closely match the facial features on each individual face:
<img src='images/michelle_detected.png' width=30% height=30%/>
```
# We use prediction on cpu here
image_copy = np.copy(image)
scale_roi = 2 # ROI scale factor. We need to upscale the detected ROI because the cascade detected ROI is too tight
# loop over the detected faces from your haar cascade
for (x, y, w, h) in faces:
# Select the region of interest that is the face in the image
w_new = w * scale_roi
h_new = h * scale_roi
dw = w_new - w
dh = h_new - h
x = max(int(x-dw/2), 0) # Avoid cropping region out of the frame
y = max(int(y-dh/2), 0)
x_max = min(image_copy.shape[1], int(x+w_new))
y_max = min(image_copy.shape[0], int(y+h_new))
roi = image_copy[y:y_max, x:x_max]
## TODO: Convert the face region from RGB to grayscale
roi = cv2.cvtColor(roi, cv2.COLOR_RGB2GRAY)
## TODO: Normalize the grayscale image so that its color range falls in [0,1] instead of [0,255]
roi = roi / 255.0
## TODO: Rescale the detected face to be the expected square size for your CNN (224x224, suggested)
roi = cv2.resize(roi, input_shape)
image_to_show = roi.copy()
## TODO: Reshape the numpy image shape (H x W x C) into a torch image shape (C x H x W)
roi = roi[:,:, np.newaxis].transpose((2, 0, 1))
## TODO: Make facial keypoint predictions using your loaded, trained network
## perform a forward pass to get the predicted facial keypoints
input_tensor = torch.from_numpy(roi.astype(np.float32))
input_tensor = torch.FloatTensor(input_tensor)
input_tensor = input_tensor.unsqueeze(0)
with torch.no_grad():
output_pts = net(input_tensor)
#output_pts = output_pts.view(output_pts.size()[0], 68, -1)
output_pts = output_pts.view(output_pts.size()[0], 68, -1)[0].data.cpu().numpy()
output_pts = output_pts * 50.0 + 100
#output_pts = output_pts.numpy()
## TODO: Display each detected face and the corresponding keypoints
plt.figure(dpi=150)
plt.imshow(image_to_show, cmap='gray')
plt.scatter(output_pts[:, 0], output_pts[:, 1], s=20, marker='.', c='m')
plt.show()
```
| github_jupyter |
# Tensorflow Timeline Analysis on Model Zoo Benchmark between Intel optimized and stock Tensorflow
This jupyter notebook will help you evaluate performance benefits from Intel-optimized Tensorflow on the level of Tensorflow operations via several pre-trained models from Intel Model Zoo. The notebook will show users a bar chart like the picture below for the Tensorflow operation level performance comparison. The red horizontal line represents the performance of Tensorflow operations from Stock Tensorflow, and the blue bars represent the speedup of Intel Tensorflow operations. The operations marked as "mkl-True" are accelerated by oneDNN a.k.a MKL-DNN, and users should be able to see a good speedup for those operations accelerated by oneDNN.
> NOTE : Users need to get Tensorflow timeline json files from other Jupyter notebooks like benchmark_perf_comparison
first to proceed this Jupyter notebook.
> NOTE: Users could also compare elapsed time of TF ops among any two different TF timeline files.
<img src="images\compared_tf_op_duration_ratio_bar.png" width="700">
The notebook will also show users two pie charts like the picture below for elapsed time percentage among different Tensorflow operations.
Users can easily find the Tensorflow operation hotspots in these pie charts among Stock and Intel Tensorflow.
<img src="images\compared_tf_op_duration_pie.png" width="700">
# Get Platform Information
```
# ignore all warning messages
import warnings
warnings.filterwarnings('ignore')
from profiling.profile_utils import PlatformUtils
plat_utils = PlatformUtils()
plat_utils.dump_platform_info()
```
# Section 1: TensorFlow Timeline Analysis
## Prerequisites
```
!pip install cxxfilt
%matplotlib inline
import matplotlib.pyplot as plt
import tensorflow as tf
import pandas as pd
pd.set_option('display.max_rows', 500)
pd.set_option('display.max_columns', 500)
pd.set_option('display.width', 1500)
```
## List out the Timeline folders
First, list out all Timeline folders from previous runs.
```
import os
filenames= os.listdir (".")
result = []
keyword = "Timeline"
for filename in filenames:
if os.path.isdir(os.path.join(os.path.abspath("."), filename)):
if filename.find(keyword) != -1:
result.append(filename)
result.sort()
index =0
for folder in result:
print(" %d : %s " %(index, folder))
index+=1
```
## Select a Timeline folder from previous runs
#### ACTION: Please select one Timeline folder and change FdIndex accordingly
```
# use the "FD_INDEX" environment variable value if it exists.
import os
env_fd_index=os.environ.get('FD_INDEX', '')
if env_fd_index != '':
FdIndex= int(env_fd_index)
else:
## USER INPUT
FdIndex= int(input('Input a index number of a folder: '))
```
List out all Timeline json files inside Timeline folder.
```
import os
TimelineFd = result[FdIndex]
print(TimelineFd)
datafiles = [TimelineFd +os.sep+ x for x in os.listdir(TimelineFd) if '.json' == x[-5:]]
print(datafiles)
if len(datafiles) is 0:
print("ERROR! No json file in the selected folder. Please select other folder.")
elif len(datafiles) is 1:
print("WARNING! There is only 1 json file in the selected folder. Please select other folder to proceed Section 1.2.")
```
> **Users can bypass below Section 1.1 and analyze performance among Stock and Intel TF by clicking the link : [Section 1_2](#section_1_2).**
<a id='section_1_1'></a>
## Section 1.1: Performance Analysis for one TF Timeline result
### Step 1: Pick one of the Timeline files
#### List out all the Timeline files first
```
index = 0
for file in datafiles:
print(" %d : %s " %(index, file))
index+=1
```
#### ACTION: Please select one timeline json file and change file_index accordingly
```
## USER INPUT
# use the "FILE_INDEX" environment variable value if it exists.
import os
env_file_index=os.environ.get('FILE_INDEX', '')
if env_file_index != '':
file_index= int(env_file_index)
else:
## USER INPUT
file_index= int(input('Input a index number of a file: '))
fn = datafiles[file_index]
tfile_prefix = fn.split('_')[0]
tfile_postfix = fn.strip(tfile_prefix)[1:]
fn
```
### Step 2: Parse timeline into pandas format
```
from profiling.profile_utils import TFTimelinePresenter
tfp = TFTimelinePresenter(True)
timeline_pd = tfp.postprocess_timeline(tfp.read_timeline(fn))
timeline_pd = timeline_pd[timeline_pd['ph'] == 'X']
```
### Step 3: Sum up the elapsed time of each TF operation
```
tfp.get_tf_ops_time(timeline_pd,fn,tfile_prefix)
```
### Step 4: Draw a bar chart for elapsed time of TF ops
```
filename= tfile_prefix +'_tf_op_duration_bar.png'
title_=tfile_prefix +'TF : op duration bar chart'
ax=tfp.summarize_barh(timeline_pd, 'arg_op', title=title_, topk=50, logx=True, figsize=(10,10))
tfp.show(ax,'bar')
```
### Step 5: Draw a pie chart for total time percentage of TF ops
```
filename= tfile_prefix +'_tf_op_duration_pie.png'
title_=tfile_prefix +'TF : op duration pie chart'
timeline_pd_known = timeline_pd[ ~timeline_pd['arg_op'].str.contains('unknown') ]
ax=tfp.summarize_pie(timeline_pd_known, 'arg_op', title=title_, topk=50, logx=True, figsize=(10,10))
tfp.show(ax,'pie')
ax.figure.savefig(filename,bbox_inches='tight')
```
<a id='section_1_2'></a>
## Section 1.2: Analyze TF Timeline results between Stock and Intel Tensorflow
> NOTE : Users could also compare elapsed time of TF ops among any two different TF timeline files.
### Speedup from oneDNN among different TF operations
### Step 1: Select one Intel and one Stock TF timeline files for analysis
> NOTE: Users could also pick any two different TF timeline files.
#### List out all timeline files in the selected folder
```
if len(datafiles) is 1:
print("ERROR! There is only 1 json file in the selected folder.")
print("Please select other Timeline folder from beginnning to proceed Section 1.2.")
for i in range(len(datafiles)):
print(" %d : %s " %(i, datafiles[i]))
```
#### ACTION: Please select one timeline file as a perfomance baseline and the other as a comparison target
put the related index for your selected timeline file.
In general, please put stock_timeline_xxxxx as the baseline.
```
# perfomance baseline
# use the "BASELINE_INDEX" environment variable value if it exists.
import os
env_baseline_index=os.environ.get('BASELINE_INDEX', '')
if env_baseline_index != '':
Baseline_Index= int(env_baseline_index)
else:
## USER INPUT
Baseline_Index= int(input('Input a index number of a Performance Baseline: '))
# comparison target
Comparison_Index = 0 if Baseline_Index else 1
```
#### List out two selected timeline files
```
selected_datafiles = []
selected_datafiles.append(datafiles[Baseline_Index])
selected_datafiles.append(datafiles[Comparison_Index])
print(selected_datafiles)
```
### Step 2: Parsing timeline results into CSV files
```
%matplotlib agg
from profiling.profile_utils import TFTimelinePresenter
csvfiles=[]
percentage_filename = ''
tfp = TFTimelinePresenter(True)
for fn in selected_datafiles:
if fn.find('/'):
fn_nofd=fn.split('/')[1]
else:
fn_nofd=fn
tfile_name= fn_nofd.split('.')[0]
tfile_prefix = fn_nofd.split('_')[0]
tfile_postfix = fn_nofd.strip(tfile_prefix)[1:]
csvpath = TimelineFd +os.sep+tfile_name+'.csv'
print(csvpath)
csvfiles.append(csvpath)
timeline_pd = tfp.postprocess_timeline(tfp.read_timeline(fn))
timeline_pd = timeline_pd[timeline_pd['ph'] == 'X']
sitems, percentage_filename = tfp.get_tf_ops_time(timeline_pd,fn,tfile_prefix)
```
#### The pie chart for elapsed time of oneDNN operations from Intel TF
```
%matplotlib inline
if percentage_filename != '':
print(percentage_filename)
tfp.plot_pie_chart(percentage_filename, 'mkl_percentage')
```
### Step 3: Pre-processing for the two CSV files
```
import os
import pandas as pd
csvarray = []
csvfilenames= []
for csvf in csvfiles:
print("read into pandas :",csvf)
a = pd.read_csv(csvf)
csvarray.append(a)
if csvf.find(os.sep) > 0:
csvfilenames.append(csvf.split(os.sep)[-1])
else:
csvfilenames.append(csvf)
a = csvarray[0]
b = csvarray[1]
# Find tags among CSV files
tags=[]
from profiling.profile_utils import PerfPresenter
perfp=PerfPresenter()
tag0, tag1 = perfp.get_diff_from_csv_filenames(csvfilenames[0][:-4],csvfilenames[1][:-4])
tags = [tag0, tag1]
print('tags : ',tags)
```
### Step 4: Merge two CSV files and caculate the speedup accordingly
#### Merge two csv files
```
import os
import pandas as pd
fdir='merged'
if not os.path.exists(fdir):
os.mkdir(fdir)
fpaths=[]
fpaths.append(fdir+os.sep+'merged.csv')
fpaths.append(fdir+os.sep+'diff_'+tags[0]+'.csv')
fpaths.append(fdir+os.sep+'diff_'+tags[1]+'.csv')
#merged=tfp.merge_two_csv_files(fpath,a,b)
merged=tfp.merge_two_csv_files_v2(fpaths, a, b, tags)
```
### Compare common operations among those two csv files
```
print("Compare common operations between ", tags)
merged_df = pd.read_csv(fpaths[0])
merged_df
```
#### The unique Tensorflow operations from the first csv/Timline file
```
%matplotlib inline
print("Operations are only in", tags[0], " run")
extra1 = pd.read_csv(fpaths[1])
extra1
```
#### The unique Tensorflow operations from the second csv/Timline file
```
print("Operations are only in", tags[1], " run")
extra2 = pd.read_csv(fpaths[2])
extra2
```
### Step 5: Draw a bar chart for elapsed time of common TF ops among stock TF and Intel TF
> NOTE: Users could also compare elapsed time of TF ops among any two different TF timeline files.
```
%matplotlib inline
print(fpaths[0])
tfp.plot_compare_bar_charts(fpaths[0], tags=tags)
tfp.plot_compare_ratio_bar_charts(fpaths[0], tags=['','oneDNN ops'], max_speedup=20)
```
### Step 6: Draw pie charts for elapsed time of TF ops among stock TF and Intel TF
> NOTE: Users could also compare elapsed time of TF ops among any two different TF timeline files.
We will have following pie charts in sequence:
1. the pie chart for elpased time of TF ops from stock TF or the first csv/Timeline file
2. the pie chart for elpased time of unique TF ops from stock TF or the first csv/Timeline file
3. the pie chart for elpased time of TF ops from Intel TF or the second csv/Timeline file
4. the pie chart for elpased time of unique TF ops from Intel TF or the second csv/Timeline file
5. the pie chart for elpased time of common TF ops among stock & Intel TF or two csv/Timeline files
#### The pie chart for elapsed time of TF ops from Stock TF or the first csv/Timline file
understand which TF operations spend most of time.
```
tfp.plot_pie_chart(csvfiles[0], tags[0])
```
#### The pie chart for elapsed time of unique TF operations from Stock TF or the first csv/Timline file
understand if there is any unique TF operation.
```
tfp.plot_pie_chart(fpaths[1], tags[0])
```
#### The pie chart for elapsed time of TF ops from Intel TF or the second csv/Timline file
understand which TF operations spend most of time.
```
tfp.plot_pie_chart(csvfiles[1], tags[1])
```
#### The pie chart for elapsed time of unique TF operations from Intel TF or the seond csv/Timline file
understand if there is any unique TF operation.
```
tfp.plot_pie_chart(fpaths[2], tags[1])
```
#### The pie chart for elapsed time of common TF ops among Stock & Intel TF or two csv/Timline files
understand top hotspots differences among Stock & Intel TF or two csv/Timeline files.
```
tfp.plot_compare_pie_charts(fpaths[0], tags=tags)
```
| github_jupyter |
# Introductory Exercises
These exercises are meant to supplement the lecture notes based on Jack VanderPlas's [excellent book](https://github.com/jakevdp/WhirlwindTourOfPython) as well as material from http://introtopython.org.
You are encouraged to modify your local version of this iPython notebook. Please add in any additional comments or advice you found helpful and definitely try to experiment with new code!
## Preamble
Before we begin, we need to run this code block to ensure the code is compatible between Python 2 and 3.
```
# Ensures compatibility between Python 2 and 3.
from __future__ import print_function, division
from __builtin__ import range
```
## Hello World
Let's quickly go through some basics of variable assignment.
```
print('Hello Python world!')
```
Try to get the following code to print the same message as the one above.
```
...
print(message)
```
## Strings
```
# Example of two strings.
my_string = "This is a double-quoted string."
your_string = 'This is a single-quoted string.'
# Example of types.
quote = "Josh once asked, 'Is this distinction useful?'"
followup = 'What do you think?'
```
We now want to manipulate these strings. A few exercises are below. Take as much time as you need; you are not expected to be able to complete them all.
```
# Combine the two lines (`quote`, `followup`) into one string with appropriate spacing.
combined_quote = ...
print(quote, followup) # printing two strings
print(combined_quote) # print the combined string
# Check whether the two strings are identical.
s1, s2 = "Test.", 'Test.'
same = ...
print(same)
# Split `my_string` into a separate string for each word.
collection_of_strings = ...
print(collection_of_strings)
# Check whether the *letter* `a` is in `quote`.
a_lettercheck = ...
print(a_lettercheck)
# Check whether the *word* `a` is in `quote`.
a_wordcheck = ...
print(a_wordcheck)
```
## Numbers
While you can define some variables explicitly, Python defines many of them implicitly and can change them when doing math. This mostly matters when working with integers and floats.
```
# Define two variables.
a, b, c = 3, 2, 5.
print('a = {0}'.format(a))
print('b =', b)
print('c', '=', c)
```
Let's see this in action below.
```
# Addition.
x = a + b
print(x, type(x))
# Subtraction.
x = c - a
print(x, type(x))
# Multiplication.
x = b * a
print(x, type(x))
# Division.
x = a / b
print(x, type(x))
# Exponentiation.
x = c ** b
print(x, type(x))
# ???
x = b // a
print(x, type(x))
x = c // a
print(x, type(x))
```
This type of distinction matters because roundoff errors can be an issue. These are a humongous pain but can be dealt with once you know what to watch out for.
```
f1, f2 = 0.3, 0.1 + 0.2
f1, f2, f1 == f2 # leaving variables for auto-output
```
Note that everything in Python is at some level an **object**. Our variables are no exception.
```
f1.as_integer_ratio(), f2.as_integer_ratio(), (0.5).as_integer_ratio()
```
## Math
Magnitudes are a pretty common unit in astronomy and are defined in terms of $m \equiv -2.5 \log(F)$ where $F$ is in units of flux density (energy/area/time/frequency).
Compute $m$ given $F = b(a-1)^2 + c(a-1) + 5$. Note that not all math operations are included in Python by default.
```
# Compute magnitude given flux.
F = ...
m = ...
print(m)
```
## Lists
We can keep track of many objects simultaneously by using **lists**.
```
dogs = ['border collie', 'australian cattle dog', 'labrador retriever']
```
Now let's print out an element of the list and capitalize it. We can figure out how to do this within the notebook by taking advantage of tab completion.
```
# Access element.
dog = dogs[0]
# Print result.
print(dog.)
```
See if you can figure out how to find where `'australian cattle dog'` is located using just standard Python operators and built-in list functions.
```
# Searching list.
dogs.index(...)
```
As before, see if you can figure out how to append items to the end of a list or insert them in the middle of a list. Append `'poodle'` to the end of the list, then insert `'chihuahua'` after `'border collie'` *without* entering the position by hand.
```
# Append.
# Insert.
print(dogs)
```
Now change border collie to `'maltese'` and remove `'labrador retriever'` from the list.
```
# Remove element.
dogs.remove(...)
dogs[0] = '...'
print(dogs)
```
Finally, a hugely useful concept in Python is **slicing** objects, where we can return only relevant portions of an object without modifying the relevant object itself.
```
# Slice an array
print(dogs[1:3])
print(dogs)
```
## Tuples
Like lists, **tuples** can keep track of many items at once. However, unlike lists, they're static and can't be changed after initialization.
```
date = (6, 11, 2018)
# Check tomorrow's date.
```
Why might we want to use tuples instead of lists?
## Dictionaries
**Dictionaries** are extremely flexible mappings of *keys* to values, and form the basis of much of Python's internal implementation. They are again another alternative to lists.
```
# Initialize dictionary.
numbers = {'one': 1, 'two': 2, 'three': 3}
# Access a value using its key.
print(numbers['two'])
# Add a new element.
numbers['ninety'] = 90
print(numbers)
```
Note that dictionaries do not maintain any sense of order for the input parameters. This is by design, since this lack of ordering allows dictionaries to be implemented very efficiently.
## Sets
**Sets** contain unordered collections of unique items only. It is really fast to perform bulk quick checks but can be hard to search. They are most useful for union, intersection, difference, symmetric difference math operations, and others. Python's sets have all of these operations built-in, via methods or operators.
```
primes = {2, 3, 5, 7}
odds = {1, 3, 5, 7, 9}
# Union: items appearing in either
print(primes.union(odds)) # equivalently with a method
# Intersction
{1, 2, 3, 5, 7, 9}
print(primes.intersection(odds)) # equivalently with a method
# difference: Items in primes but not in odds
print(primes.difference(odds)) # equivalently with a method
```
## Conditionals
Python's conditional statements are pretty straightforrward.
```
x = -15
if x == 0:
print(x, "is zero")
elif x > 0:
print(x, "is positive")
elif x < 0:
print(x, "is negative")
else:
print(x, "is unlike anything I've ever seen...")
```
## Loops
Python is able to work with any type of iterable, not just integers. This gives loops a lot more flexibility.
```
# Traditional loop.
for i in range(3):
print('I like ' + dogs[i] + 's.')
# "Pythonic" loop.
for dog in dogs:
print('I like ' + dog + 's.')
```
This flexibility makes loops in Python incredibly powerful. As an example, see if you can get the loop below to run using some combination of (1) `range`, (2) `zip`, and (3) `enumerate`.
```
for ... in ...:
print(counter, ':', dog)
```
Alternately, we can use while loops to get things done.
```
i = 0
while i < len(dogs):
print(i, ':', dogs[i])
i += 1
```
## Variable (Re-)Assignment
Beware (re-)assigning things without warning! Python by default only "protects" a certain subset of objects by automatically copying them.
```
# Example 1.
josh = quote # reassign
josh += " He got #rekt." # ???
print(quote)
print(josh)
# Example 2.
quotes = quote.split() # reassign
josh = quotes
josh += " He got #rekt.".split() # ???
print(quotes)
print(josh)
# Example 3.
quotes = quote.split() # reassign
josh = quotes[:]
josh += " He got #rekt.".split() # ???
print(quotes)
print(josh)
# Example 4.
d = b
d += 3.
print(b)
print(d)
```
## Good Coding Practices
Finally, now that our code is becoming slightly longer, let's take a step back and talk about good coding practices. Some of these are coded into Python itself.
```
import this
```
In addition, there's also the [PEP8 style guide](https://www.python.org/dev/peps/pep-0008/).
## Review
Starting [here](https://github.com/jakevdp/WhirlwindTourOfPython/blob/6f1daf714fe52a8dde6a288674ba46a7feed8816/02-Basic-Python-Syntax.ipynb).
## Exercises
Courtesy of Luke Kelley.
References:
- [pyformat.info](pyformat.info)
- [docs.python.org/3/library/string.html](https://docs.python.org/3/library/string.html#format-specification-mini-language)
Starting with the following variables:
```
title = "Banneker + Aztlan Initiative"
month = 6
day = 11
year = 2018
width = 30
```
**1) Write out the date in the following formats using the above variables:**
a) *Note that the month is "zero-padded" ('`06`', instead of just '`6`'):*
```
2018/06/11
```
b) *Note that only the last two digits of the year are shown:*
```
06/11/18
```
### 2) Draw an underline to match '`title`'
Print out the following string using the `title` variable and without manually typing more than a single '`=`' character:
```
Banneker + Aztlan Initiative
============================
```
*Hint: Not only can you add two strings together (i.e. `abc` + `def`), but you can also multiple a string by an integer - try it!*
### 3) Use a `for`-loop and `if`-statement to write:
Note: *there are `width` (i.e. 30) characters between each pair of "pipes": `|...|`*
```
|==============================|
|Banneker |
| Aztlan|
| Initiative |
| 2018/06/11 |
|==============================|
```
| github_jupyter |
# Benchmark results reporting
## Setup
### Prerequirements
This notebook requires a kernel running Python 3.5+.
You can skip this section if the kernel is already configured.
```
!pip install -r ./requirements.txt
#!pip install jupyter_contrib_nbextensions
#!jupyter contrib nbextension install --user
#!jupyter nbextension enable python-markdown/main
#!pip install jupyter_nbextensions_configurator
#!jupyter nbextensions_configurator enable --user
```
### Imports and selection of the results directory
```
import glob
import os
import sys
automlbenchmark_path = ".."
amlb_dir = os.path.realpath(os.path.expanduser(automlbenchmark_path))
for lib in [amlb_dir]:
sys.path.insert(0, lib)
import numpy as np
import pandas as pd
from amlb_report import draw_score_heatmap, draw_score_parallel_coord, draw_score_pointplot, draw_score_stripplot, draw_score_barplot\
, prepare_results, render_leaderboard, render_metadata, render_summary
from amlb_report.util import create_file, display
from amlb_report.visualizations.util import register_colormap, render_colormap, savefig
import amlb_report.config as config
```
## Results
#### Loading results, formatting and adding columns
- `result` is the raw result metric computed from predictions at the end the benchmark.
For classification problems, it is usually `auc` for binomial classification and `neg_logloss` for multinomial classification (higher is always better).
- `norm_result` is a normalization of `result` on a `[0, 1]` scale, with `{{normalization[0]}}` result as `0` and `{{normalization[1]}}` result as `1`.
- `imp_result` for imputed results. Given a task and a framework:
- if **all folds results are missing**, then no imputation occurs, and the result is `nan` for each fold.
- if **only some folds results are missing**, then the missing result can be imputed by setting `{{imputation='framework'}}` and use that framework to impute the result for this fold.
### Default config
```
#! avoid editing this cell: custom config should be applied in the next cell.
constraint = "1h8c"
results_dir = "."
output_dir = "."
tasks_sort_by = 'nrows'
results_group = ''
included_frameworks = []
excluded_frameworks = []
frameworks_sort_key = None
# frameworks_sort_key = lambda f: definitions[f]['key'] if 'key' in definitions[f] else f.lower()
frameworks_labels = None
# frameworks_labels = lambda l: definitions[l]['framework'].lower()
duplicates_handling = 'fail' # accepted values: 'fail', 'keep_first', 'keep_last', 'keep_none'
imputation = None
normalization = None
# normalization = (0, 'h2o', 'mean')
row_filter = None
# row_filter = lamdba r: r.fold == 0 #! r is a pd.Series
title_extra = ""
binary_result_label = 'AUC'
multiclass_result_label = 'neg. Log loss'
regression_result_label = 'neg. RMSE'
# register_colormap(config.colormap, ('colorblind', [1, 0, 2, 3, 4, 5]))
```
### Config and results definitions for current run
```
# this cell is an example showing how to use/customize this notebook depending on your results
config.nfolds = 1
results_dir = "../results"
output_dir = "./tmp"
duplicates_handling = 'keep_last'
normalization = (0, 'constantpredictor', 'mean') # normalizes results between 0 and constantpredictor
# row_filter = lambda r: ~r.task.isin(['kddcup09_appetency', 'colleges'])
definitions = dict(
constantpredictor=dict(
ref = True,
framework='constantpredictor_enc',
results=glob.glob(f"{results_dir}/constantpredictor*/scores/results.csv")
),
autogluon=dict(
framework='AutoGluon',
results=glob.glob(f"{results_dir}/autogluon*/scores/results.csv")
),
autosklearn=dict(
framework='autosklearn',
results=glob.glob(f"{results_dir}/autosklearn*/scores/results.csv")
),
h2oautoml=dict(
framework='H2OAutoML',
results=glob.glob(f"{results_dir}/h2oautoml*/scores/results.csv")
),
tpot=dict(
framework='TPOT',
results=glob.glob(f"{results_dir}/tpot*/scores/results.csv")
),
# rf=dict(
# framework='RandomForest',
# results=my_results_df[my_results_df['framework']=='RandomForest'] # example showing that we can also use a dataframe (or its subset)
# )
)
#definitions
```
## Load and prepare results
```
runs = {k:v for k, v in definitions.items()
if (k in included_frameworks if included_frameworks else True)
and k not in excluded_frameworks}
#runs
def results_as_df(results_dict, row_filter=None):
def apply_filter(res, filtr):
r = res.results
return r.loc[filtr(r)]
if row_filter is None:
row_filter = lambda r: True
return pd.concat([apply_filter(res, lambda r: (r.framework==name) & row_filter(r))
for name, res in results_dict.items()
if res is not None])
ref_results = {name: prepare_results(run['results'],
renamings={run['framework']: name},
exclusions=excluded_frameworks,
normalization=normalization,
duplicates_handling=duplicates_handling,
include_metadata=True
)
for name, run in runs.items() if runs[name].get('ref', False)}
all_ref_res = results_as_df(ref_results, row_filter)
runs_results = {name: prepare_results(run['results'],
renamings={run['framework']: name},
exclusions=excluded_frameworks,
imputation=imputation,
normalization=normalization,
ref_results=all_ref_res,
duplicates_handling=duplicates_handling
)
for name, run in runs.items() if name not in ref_results}
all_res = pd.concat([
all_ref_res,
results_as_df(runs_results, row_filter)
])
all_results = {**ref_results, **runs_results}
from functools import reduce
metadata = reduce(lambda l, r: {**r, **l},
[res.metadata
for res in list(ref_results.values())+list(runs_results.values())
if res is not None],
{})
# metadata = next(res for res in ref_results.values()).metadata
problem_types = pd.DataFrame(m.__dict__ for m in metadata.values())['type'].unique().tolist()
```
## Tasks lists
```
merged_res = pd.concat([r.done.reset_index() for r in all_results.values() if r is not None])
merged_res = merged_res[merged_res['id'].notna()]
merged_results = prepare_results(merged_res)
def render_tasks_by_state(state='done'):
# tasks = pd.concat([getattr(r, state).reset_index()
# .groupby(['task', 'framework'])['fold']
# .unique()
# for r in all_results.values()
# if r is not None])
tasks = (getattr(merged_results, state).reset_index()
.groupby(['task', 'framework'])['fold']
.unique())
display(tasks, pretty=True)
# display(tabulate(done, tablefmt='plain'))
render_metadata(metadata,
filename=create_file(output_dir, "datasets", results_group, "metadata.csv"))
```
### Completed tasks/folds
```
render_tasks_by_state('done')
```
### Missing or crashed/aborted tasks/folds
```
render_tasks_by_state('missing')
```
### Failing tasks/folds
```
render_tasks_by_state('failed')
failures = (merged_results.failed.groupby(['task', 'fold', 'framework'])['info']
.unique())
display(failures)
```
### Results anomalies
```
from amlb_report.analysis import list_outliers
display(list_outliers('result',
results=merged_results.results,
# results=merged_results.loc[merged_results.framework=='h2oautoml']
z_threshold=2.5,
))
```
## Data Reports
### Results summary
Averaging using arithmetic mean over fold `result`.
In following summaries, if not mentioned otherwise, and if results imputation was enabled, the means are computed over imputed results .
Given a task and a framework:
- if **all folds results are missing**, then no imputation occured, and the mean result is `nan`.
- if **only some folds results are missing**, then the amount of imputed results that contributed to the mean are displayed between parenthesis.
#### Number of models trained
When available, displays the average amount of models trained by the framework for each dataset.
This amount should be interpreted differently for each framework.
For example, with *RandomForest*, this amount corresponds to the number of trees.
```
models_summary = render_summary('models_count',
results=all_res)
models_summary.to_csv(create_file(output_dir, "tables", "models_summary.csv"))
```
### Resuls mean
```
res_summary = render_summary('result',
results=all_res)
res_summary.to_csv(create_file(output_dir, "tables", "results_summary.csv"))
if normalization:
norm_result_summary = render_summary('norm_result',
results=all_res)
norm_result_summary.to_csv(create_file(output_dir, "tables", "normalized_result_summary.csv"))
```
### Tasks leaderboard
```
benchmark_leaderboard = render_leaderboard('result',
results=all_res,
aggregate=True)
benchmark_leaderboard.to_csv(create_file(output_dir, "tables", "benchmark_leaderboard.csv"))
```
## Visualizations
```
render_colormap(config.colormap)
```
### Heatmaps
```
if 'binary' in problem_types:
fig = draw_score_heatmap('result',
results=all_res,
type_filter='binary',
metadata=metadata,
x_labels=frameworks_labels or True,
x_sort_by=frameworks_sort_key,
y_sort_by='nrows',
title=f"Results ({binary_result_label}) on {results_group} binary classification problems{title_extra}",
center=0.5
);
savefig(fig, create_file(output_dir, "visualizations", "binary_result_heat.png"))
if 'multiclass' in problem_types:
fig = draw_score_heatmap('result',
results=all_res,
type_filter='multiclass',
metadata=metadata,
x_labels=frameworks_labels or True,
x_sort_by=frameworks_sort_key,
y_sort_by='nrows',
title=f"Results ({multiclass_result_label}) on {results_group} multi-class classification problems{title_extra}",
center=0
);
savefig(fig, create_file(output_dir, "visualizations", "multiclass_result_heat.png"))
if 'regression' in problem_types:
fig = draw_score_heatmap('result',
results=all_res,
type_filter='regression',
metadata=metadata,
x_labels=frameworks_labels or True,
x_sort_by=frameworks_sort_key,
y_sort_by='nrows',
title=f"Results ({regression_result_label}) on {results_group} regression problems{title_extra}",
center=0
);
savefig(fig, create_file(output_dir, "visualizations", "regression_result_heat.png"))
```
### Bar plots
```
if 'binary' in problem_types:
fig = draw_score_barplot('result',
results=all_res,
type_filter='binary',
metadata=metadata,
x_sort_by=tasks_sort_by,
ylabel=binary_result_label,
ylim=dict(bottom=.5),
hue_sort_by=frameworks_sort_key,
ci=95,
title=f"Results ({binary_result_label}) on {results_group} binary classification problems{title_extra}",
legend_loc='lower center',
legend_labels=frameworks_labels,
);
savefig(fig, create_file(output_dir, "visualizations", "binary_result_barplot.png"))
if 'multiclass' in problem_types:
fig = draw_score_barplot('result',
results=all_res,
type_filter='multiclass',
metadata=metadata,
x_sort_by=tasks_sort_by,
ylabel=multiclass_result_label,
ylim=dict(top=0.1),
hue_sort_by=frameworks_sort_key,
ci=95,
title=f"Results ({multiclass_result_label}) on {results_group} multiclass classification problems{title_extra}",
legend_loc='lower center',
legend_labels=frameworks_labels,
);
savefig(fig, create_file(output_dir, "visualizations", "multiclass_result_barplot.png"))
if 'regression' in problem_types:
fig = draw_score_barplot('result',
results=all_res,
type_filter='regression',
metadata=metadata,
x_sort_by=tasks_sort_by,
yscale='symlog',
ylabel=regression_result_label,
ylim=dict(top=0.1),
hue_sort_by=frameworks_sort_key,
ci=95,
title=f"Results ({regression_result_label}) on {results_group} regression classification problems{title_extra}",
legend_loc='lower center',
legend_labels=frameworks_labels,
size=(8, 6),
);
savefig(fig, create_file(output_dir, "visualizations", "regression_result_barplot.png"))
```
### Point plots
```
if 'binary' in problem_types:
fig = draw_score_pointplot('result',
results=all_res,
type_filter='binary',
metadata=metadata,
x_sort_by=tasks_sort_by,
ylabel=binary_result_label,
ylim=dict(bottom=.5),
hue_sort_by=frameworks_sort_key,
join='none', marker='hline_xspaced', ci=95,
title=f"Results ({binary_result_label}) on {results_group} binary classification problems{title_extra}",
legend_loc='lower center',
legend_labels=frameworks_labels,
);
savefig(fig, create_file(output_dir, "visualizations", "binary_result_pointplot.png"))
if 'multiclass' in problem_types:
fig = draw_score_pointplot('result',
results=all_res,
type_filter='multiclass',
metadata=metadata,
x_sort_by=tasks_sort_by,
ylabel=multiclass_result_label,
hue_sort_by=frameworks_sort_key,
join='none', marker='hline_xspaced', ci=95,
title=f"Results ({multiclass_result_label}) on {results_group} multiclass classification problems{title_extra}",
legend_loc='lower center',
legend_labels=frameworks_labels,
);
savefig(fig, create_file(output_dir, "visualizations", "multiclass_result_pointplot.png"))
if 'regression' in problem_types:
fig = draw_score_pointplot('result',
results=all_res,
type_filter='regression',
metadata=metadata,
x_sort_by=tasks_sort_by,
ylabel=regression_result_label,
yscale='symlog',
ylim=dict(top=0.1),
hue_sort_by=frameworks_sort_key,
join='none', marker='hline_xspaced', ci=95,
title=f"Results ({regression_result_label}) on {results_group} regression classification problems{title_extra}",
legend_loc='lower center',
legend_labels=frameworks_labels,
size=(8, 6),
);
savefig(fig, create_file(output_dir, "visualizations", "regression_result_pointplot.png"))
```
### Strip plots
```
if 'binary' in problem_types:
fig = draw_score_stripplot('result',
results=all_res.sort_values(by=['framework']),
type_filter='binary',
metadata=metadata,
xlabel=binary_result_label,
y_sort_by=tasks_sort_by,
hue_sort_by=frameworks_sort_key,
title=f"Results ({binary_result_label}) on {results_group} binary classification problems{title_extra}",
legend_labels=frameworks_labels,
);
savefig(fig, create_file(output_dir, "visualizations", "binary_result_stripplot.png"))
if 'multiclass' in problem_types:
fig = draw_score_stripplot('result',
results=all_res.sort_values(by=['framework']),
type_filter='multiclass',
metadata=metadata,
xlabel=multiclass_result_label,
xscale='symlog',
y_sort_by=tasks_sort_by,
hue_sort_by=frameworks_sort_key,
title=f"Results ({multiclass_result_label}) on {results_group} multi-class classification problems{title_extra}",
legend_labels=frameworks_labels,
);
savefig(fig, create_file(output_dir, "visualizations", "multiclass_result_stripplot.png"))
if 'regression' in problem_types:
fig = draw_score_stripplot('result',
results=all_res,
type_filter='regression',
metadata=metadata,
xlabel=regression_result_label,
xscale='symlog',
y_sort_by=tasks_sort_by,
hue_sort_by=frameworks_sort_key,
title=f"Results ({regression_result_label}) on {results_group} regression problems{title_extra}",
legend_labels=frameworks_labels,
);
savefig(fig, create_file(output_dir, "visualizations", "regression_result_stripplot.png"))
```
### Normalized strip plots
```
if 'binary' in problem_types and normalization:
fig = draw_score_stripplot('norm_result',
results=all_res,
type_filter='binary',
metadata=metadata,
xlabel=f"rel. {binary_result_label}",
y_sort_by='nrows',
hue_sort_by=frameworks_sort_key,
title=f"Relative results ({binary_result_label}) on {results_group} binary classification problems{title_extra}",
legend_labels=frameworks_labels,
);
savefig(fig, create_file(output_dir, "visualizations", "binary_rel_result_stripplot.png"))
if 'multiclass' in problem_types and normalization:
fig = draw_score_stripplot('norm_result',
results=all_res,
type_filter='multiclass',
metadata=metadata,
xlabel=f"rel. {multiclass_result_label}",
xscale='symlog',
y_sort_by='nrows',
hue_sort_by=frameworks_sort_key,
title=f"Relative results ({multiclass_result_label}) on {results_group} multi-class classification problems{title_extra}",
legend_labels=frameworks_labels,
);
savefig(fig, create_file(output_dir, "visualizations", "multiclass_rel_result_stripplot.png"))
if 'regression' in problem_types and normalization:
fig = draw_score_stripplot('norm_result',
results=all_res,
type_filter='regression',
metadata=metadata,
xlabel=f"rel. {regression_result_label}",
y_sort_by='nrows',
hue_sort_by=frameworks_sort_key,
title=f"Relative results ({regression_result_label}) on {results_group} regression problems{title_extra}",
legend_labels=frameworks_labels,
);
savefig(fig, create_file(output_dir, "visualizations", "regression_rel_result_stripplot.png"))
```
## Playground
| github_jupyter |
# TF on GKE
This notebook shows how to run the [TensorFlow CIFAR10 sample](https://github.com/tensorflow/models/tree/master/tutorials/image/cifar10_estimator) on GKE using [TFJobs](https://github.com/kubeflow/tf-operator)
## Requirements
To run this notebook you must have the following installed
* gcloud
* kubectl
* helm
* kubernetes python client library
There is a Docker image based on Datalab suitable for running this notebook.
You can start that container as follows
```
docker run --name=gke-datalab -p "127.0.0.1:8081:8080" \
-v "${HOME}:/content/datalab/home" \
-v /var/run/docker.sock:/var/run/docker.sock -d -e "PROJECT_ID=" \
gcr.io/tf-on-k8s-dogfood/gke-datalab:v20171103-73616f0
```
* You need to map in docker if you want tobuild docker images inside the container.
* Alternatively, you can set "use_gcb" to true in order to build the images using Google Container Builder
Additionally the [py package](https://github.com/kubeflow/tf-operator/tree/master/py) must be a top level package importable as py
* If you cloned [kubeflow/tf-operator](https://github.com/kubeflow/tf-operator) and are running this notebook in place the path with be configured automatically
## Preliminaries
```
# Turn on autoreloading
%load_ext autoreload
%autoreload 2
```
import a bunch of modules and set some constants
```
from __future__ import print_function
import logging
import os
import sys
# Assumes we are running inside the cloned repo.
# Try to setup the path so we can import py as a top level package
ROOT_DIR = os.path.abspath(os.path.join("../.."))
if os.path.exists(os.path.join(ROOT_DIR, "py")):
if not ROOT_DIR in sys.path:
sys.path.append(ROOT_DIR)
import kubernetes
from kubernetes import client as k8s_client
from kubernetes import config as k8s_config
from kubernetes.client.rest import ApiException
from kubernetes.client.models.v1_label_selector import V1LabelSelector
import datetime
from googleapiclient import discovery
from googleapiclient import errors
from oauth2client.client import GoogleCredentials
from pprint import pprint
try:
from py import build_and_push_image
from py import util
except ImportError:
raise ImportError("Please ensure the py package in https://github.com/kubeflow/tf-operator is a top level package")
import StringIO
import subprocess
import urllib
import urllib2
import time
import yaml
logging.getLogger().setLevel(logging.INFO)
TF_JOB_GROUP = "kubeflow.org"
TF_JOB_VERSION = "v1alpha1"
TF_JOB_PLURAL = "tfjobs"
TF_JOB_KIND = "TFJob"
```
### Configure the notebook for your use
Change the constants defined below.
1. Change **project** to a project you have access to.
* GKE should be enabled for that project
1. Change **data_dir** and **job_dir**
* Use a GCS bucket that you have access to
* Ensure the service account on your GKE cluster can read/write to this GCS bucket
* Optional change the cluster name
```
project="cloud-ml-dev"
zone="us-east1-d"
cluster_name="gke-tf-example"
registry = "gcr.io/" + project
data_dir = "gs://cloud-ml-dev_jlewi/cifar10/data"
job_dirs = "gs://cloud-ml-dev_jlewi/cifar10/jobs"
gke = discovery.build("container", "v1")
namespace = "default"
# Whether to build containers using Google Container Builder.
# Set to false it will build by shelling out to docker build.
use_gcb = "false"
```
## GKE Cluster Setup
* The instructions below create a **CPU** cluster
* To create a GKE cluster with GPUs sign up for the [GKE GPU Alpha](https://goo.gl/forms/ef7eh2x00hV3hahx1)
* To use GPUs set accelerator and accelerator_count
* For a full list of cluster options see the [Cluster object](https://cloud.google.com/container-engine/reference/rest/v1/projects.zones.clusters#Cluster)
in the GKE API docs
To use an existing GKE cluster call **configure_kubectl** but not **create_cluster**
* The code below issues a GKE request to create the cluster by calling util.create_cluster
* util.create_cluster uses the GKE python client library
* After creating the cluster we call util.configure_kubectl
* This configures your machine to talk to the K8s master of the newly created cluster
```
reload(util)
machine_type = "n1-standard-8"
use_gpu = True
if use_gpu:
accelerator = "nvidia-tesla-k80"
accelerator_count = 1
else:
accelerator = None
accelerator_count = 0
cluster_request = {
"cluster": {
"name": cluster_name,
"description": "A GKE cluster for TF.",
"initialNodeCount": 1,
"nodeConfig": {
"machineType": machine_type,
"oauthScopes": [
"https://www.googleapis.com/auth/cloud-platform",
],
},
# TODO(jlewi): Stop pinning GKE version once 1.8 becomes the default.
"initialClusterVersion": "1.8.1-gke.1",
}
}
if bool(accelerator) != (accelerator_count > 0):
raise ValueError("If accelerator is set accelerator_count must be > 0")
if accelerator:
# TODO(jlewi): Stop enabling Alpha once GPUs make it out of Alpha
cluster_request["cluster"]["enableKubernetesAlpha"] = True
cluster_request["cluster"]["nodeConfig"]["accelerators"] = [
{
"acceleratorCount": accelerator_count,
"acceleratorType": accelerator,
},
]
util.create_cluster(gke, project, zone, cluster_request)
util.configure_kubectl(project, zone, cluster_name)
k8s_config.load_kube_config()
# Create an API client object to talk to the K8s master.
api_client = k8s_client.ApiClient()
```
### Install the Operator
* We need to deploy the [TFJob](https://github.com/kubeflow/tf-operator) custom resource on our K8s cluster
* TFJob is deployed using the [helm](https://github.com/kubernetes/helm) package manager so first we need to setup helm on our cluster
```
util.setup_cluster(api_client)
```
Now that helm is setup we can deploy the TFJob CRD
```
CHART="https://storage.googleapis.com/tf-on-k8s-dogfood-releases/latest/tf-job-operator-chart-latest.tgz"
util.run(["helm", "install", CHART, "-n", "tf-job", "--wait", "--replace", "--set", "rbac.install=true,cloud=gke"])
```
## Build Docker images
To run a TensorFlow program on K8s we need to package our code as Docker images.
The [Dockerfile](https://github.com/jlewi/k8s/blob/73616f09f335defc92f9b20225c272862e92e32b/examples/tensorflow-models/Dockerfile.template)
for this example starts with the published Docker images for TensorFlow ands
the code for our TensorFlow program
* In this example we are using the CIFAR10 example in the [TensorFlow's model zoo](https://github.com/tensorflow/models)
* So our Dockerfile just clones that repo
* Using TF's Docker images ensures we start with a reliable TF environment
We need to build separate Docker images for CPU and GPU versions of TensorFlow.
* **modes** controls whether we build images for CPU, GPU or both
* Our Dockerfile is a [Jinja2](http://jinja.pocoo.org/) template, so we can easily
build docker images based on different TensorFlow versions
The base images controls which version of TensorFlow we will use
* Change the base images if you want to use a different version.
```
reload(build_and_push_image)
if use_gpu:
modes = ["cpu", "gpu"]
else:
modes = ["cpu"]
image = os.path.join(registry, "tf-models")
dockerfile = os.path.join(ROOT_DIR, "examples", "tensorflow-models", "Dockerfile.template")
base_images = {
"cpu": "gcr.io/tensorflow/tensorflow:1.3.0",
"gpu": "gcr.io/tensorflow/tensorflow:1.3.0-gpu",
}
images = build_and_push_image.build_and_push(dockerfile, image, modes=modes, base_images=base_images)
```
## Create the CIFAR10 Datasets
We need to create the cifar10 TFRecord files by running [generate_cifar10_tfrecords.py](https://github.com/tensorflow/models/blob/master/tutorials/image/cifar10_estimator/generate_cifar10_tfrecords.py)
* We submit a K8s job to run this program
* You can skip this step if your data is already available in data_dir
```
batch_api = k8s_client.BatchV1Api(api_client)
job_name = "cifar10-data-"+ datetime.datetime.now().strftime("%y%m%d-%H%M%S")
body = {}
body['apiVersion'] = "batch/v1"
body['kind'] = "Job"
body['metadata'] = {}
body['metadata']['name'] = job_name
body['metadata']['namespace'] = namespace
# Note backoffLimit requires K8s >= 1.8
spec = """
backoffLimit: 4
template:
spec:
containers:
- name: cifar10
image: {image}
command: ["python", "/tensorflow_models/tutorials/image/cifar10_estimator/generate_cifar10_tfrecords.py", "--data-dir={data_dir}"]
restartPolicy: Never
""".format(data_dir=data_dir, image=images["cpu"])
spec_buffer = StringIO.StringIO(spec)
body['spec'] = yaml.load(spec_buffer)
try:
# Create a Resource
api_response = batch_api.create_namespaced_job(namespace, body)
print("Created job %s" % api_response.metadata.name)
except ApiException as e:
print(
"Exception when calling DefaultApi->apis_fqdn_v1_namespaces_namespace_resource_post: %s\n" %
e)
```
wait for the job to finish
```
while True:
results = batch_api.read_namespaced_job(job_name, namespace)
if results.status.succeeded >= 1 or results.status.failed >= 3:
break
print("Waiting for job %s ...." % results.metadata.name)
time.sleep(5)
if results.status.succeeded >= 1:
print("Job completed successfully")
else:
print("Job failed")
```
## Create a TFJob
To submit a TFJob, we define a TFJob spec and then create it in our cluster
```
crd_api = k8s_client.CustomObjectsApi(api_client)
namespace = "default"
job_name = "cifar10-"+ datetime.datetime.now().strftime("%y%m%d-%H%M%S")
job_dir = os.path.join(job_dirs, job_name)
num_steps = 10
body = {}
body['apiVersion'] = TF_JOB_GROUP + "/" + TF_JOB_VERSION
body['kind'] = TF_JOB_KIND
body['metadata'] = {}
body['metadata']['name'] = job_name
body['metadata']['namespace'] = namespace
master_image = images["cpu"]
if use_gpu:
master_image = images["gpu"]
spec = """
replicaSpecs:
- replicas: 1
tfReplicaType: MASTER
template:
spec:
containers:
- image: {master_image}
name: tensorflow
command:
- python
- /tensorflow_models/tutorials/image/cifar10_estimator/cifar10_main.py
- --data-dir={data_dir}
- --job-dir={job_dir}
- --train-steps={num_steps}
- --num-gpus={num_gpus}
restartPolicy: OnFailure
tfImage: {cpu_image}
tensorBoard:
logDir: {job_dir}
""".format(master_image=master_image, cpu_image=images["cpu"], data_dir=data_dir, job_dir=job_dir, num_steps=num_steps, num_gpus=accelerator_count)
spec_buffer = StringIO.StringIO(spec)
body['spec'] = yaml.load(spec_buffer)
if use_gpu:
body['spec']['replicaSpecs'][0]["template"]["spec"]["containers"][0]["resources"] = {
"limits": {
"nvidia.com/gpu": accelerator_count,
}
}
try:
# Create a Resource
api_response = crd_api.create_namespaced_custom_object(TF_JOB_GROUP, TF_JOB_VERSION, namespace, TF_JOB_PLURAL, body)
logging.info("Created job %s", api_response["metadata"]["name"])
except ApiException as e:
print(
"Exception when calling DefaultApi->apis_fqdn_v1_namespaces_namespace_resource_post: %s\n" %
e)
```
## Monitoring your job and waiting for it to finish
We can monitor the job a number of ways
* We can poll K8s to get the status of the TFJob
* We can check the TensorFlow logs
* These are available in StackDriver
* We can access TensorBoard if the TFJob was configured to launch TensorBoard
Running the code below will poll K8s for the TFJob status and also print out relevant links for TensorBoard and the StackDriver logs
To access TensorBoard you will need to run **kubectl proxy** to create a proxy connection to your K8s cluster
```
# Get pod logs
v1 = k8s_client.CoreV1Api(api_client)
k8s_config.load_kube_config()
api_client = k8s_client.ApiClient()
crd_api = k8s_client.CustomObjectsApi(api_client)
master_started = False
runtime_id = None
while True:
results = crd_api.get_namespaced_custom_object(TF_JOB_GROUP, TF_JOB_VERSION, namespace, TF_JOB_PLURAL, job_name)
if not runtime_id:
runtime_id = results["spec"]["RuntimeId"]
logging.info("Job has runtime id: %s", runtime_id)
tensorboard_url = "http://127.0.0.1:8001/api/v1/proxy/namespaces/{namespace}/services/tensorboard-{runtime_id}:80/".format(
namespace=namespace, runtime_id=runtime_id)
logging.info("Tensorboard will be available at job\n %s", tensorboard_url)
if not master_started:
# Get the master pod
# TODO(jlewi): V1LabelSelector doesn't seem to help
pods = v1.list_namespaced_pod(namespace=namespace, label_selector="runtime_id={0},job_type=MASTER".format(runtime_id))
# TODO(jlewi): We should probably handle the case where more than 1 pod gets started.
# TODO(jlewi): Once GKE logs pod labels we can just filter by labels to get all logs for a particular task
# and not have to identify the actual pod.
if pods.items:
pod = pods.items[0]
logging.info("master pod is %s", pod.metadata.name)
query={
'advancedFilter': 'resource.type="container"\nresource.labels.namespace_id="default"\nresource.labels.pod_id="{0}"'.format(pod.metadata.name),
'dateRangeStart': pod.metadata.creation_timestamp.isoformat(),
'expandAll': 'false',
'interval': 'NO_LIMIT',
'logName': 'projects/{0}/logs/tensorflow'.format(project),
'project': project,
}
logging.info("Logs will be available in stackdriver at\n"
"https://console.cloud.google.com/logs/viewer?" + urllib.urlencode(query))
master_started = True
if results["status"]["phase"] == "Done":
break
print("Job status {0}".format(results["status"]["phase"]))
time.sleep(5)
logging.info("Job %s", results["status"]["state"])
```
## Cleanup
* Delete the GKE cluster
```
util.delete_cluster(gke, cluster_name, project, zone)
```
## Appendix
```
from kubernetes.client.models.v1_label_selector import V1LabelSelector
import urllib2
# Get pod logs
k8s_config.load_kube_config()
api_client = k8s_client.ApiClient()
v1 = k8s_client.CoreV1Api(api_client)
runtime_id = results["spec"]["RuntimeId"]
# TODO(jlewi): V1LabelSelector doesn't seem to help
pods = v1.list_namespaced_pod(namespace=namespace, label_selector="runtime_id={0},job_type=MASTER".format(runtime_id))
pod = pods.items[0]
```
#### Read the Pod Logs From K8s
We can read pod logs directly from K8s and not depend on stackdriver
```
ret = v1.read_namespaced_pod_log(namespace=namespace, name=pod.metadata.name)
print(ret)
```
#### Fetch Logs from StackDriver Programmatically
* On GKE pod logs are stored in stackdriver
* These logs will stick around longer than pod logs
* Fetching from stackNote this tends to be a little slow()
```
from google.cloud import logging as gcp_logging
pod_filter = 'resource.type="container" AND resource.labels.pod_id="master-hrhh-0-wrh6g"'
client = gcp_logging.Client(project=project)
for entry in client.list_entries(filter_=pod_filter):
print(entry.payload.strip())
```
| github_jupyter |
```
import keras
keras.__version__
```
# Compreendendo as redes neurais recursivas
## A first recurrent layer in Keras
Para implementar uma Recursive Neural Network lançamos mão de um tipo de camada na Keras denominado `SimpleRNN`.
```
from keras.layers import SimpleRNN
```
Há apenas uma pequena diferença: `SimpleRNN` processa lotes de sequências, como todas as outras camadas Keras, e não apenas uma única sequência, como no nosso exemplo Numpy. Isso significa que ele recebe entradas da forma `(batch_size, timesteps, input_features)`, em vez de `(timesteps, input_features)`.
Como todas as camadas recorrentes no Keras, o `SimpleRNN` pode ser executado em dois modos diferentes: ele pode retornar as seqüências completas de saídas sucessivas para cada timestep (um tensor 3D de forma` (batch_size, timesteps, output_features) `) ou pode retornar apenas a última saída para cada sequência de entrada (um tensor 2D da forma `(batch_size, output_features)`). Esses dois modos são controlados pelo argumento do construtor `return_sequences`. Vamos dar uma olhada em um exemplo:
```
from keras.models import Sequential
from keras.layers import Embedding, SimpleRNN
model = Sequential()
model.add(Embedding(10000, 32))
model.add(SimpleRNN(32))
model.summary()
model = Sequential()
model.add(Embedding(10000, 32))
model.add(SimpleRNN(32, return_sequences=True))
model.summary()
```
Às vezes, é útil empilhar várias camadas recorrentes uma após a outra para aumentar o poder representacional de uma rede. Nessa configuração, é necessário obter todas as camadas intermediárias para retornar sequências completas:
```
model = Sequential()
model.add(Embedding(10000, 32))
model.add(SimpleRNN(32, return_sequences=True))
model.add(SimpleRNN(32, return_sequences=True))
model.add(SimpleRNN(32, return_sequences=True))
model.add(SimpleRNN(32)) # This last layer only returns the last outputs.
model.summary()
```
Agora vamos tentar usar esse modelo no problema de classificação de revisão de filmes do IMDB. Primeiro, vamos pré-processar os dados:
```
from keras.datasets import imdb
from keras.preprocessing import sequence
max_features = 10000 # number of words to consider as features
maxlen = 500 # cut texts after this number of words (among top max_features most common words)
batch_size = 32
print('Loading data...')
(input_train, y_train), (input_test, y_test) = imdb.load_data(num_words=max_features)
print(len(input_train), 'train sequences')
print(len(input_test), 'test sequences')
print('Pad sequences (samples x time)')
input_train = sequence.pad_sequences(input_train, maxlen=maxlen)
input_test = sequence.pad_sequences(input_test, maxlen=maxlen)
print('input_train shape:', input_train.shape)
print('input_test shape:', input_test.shape)
```
Vamos treinar uma rede recorrente simples usando uma camada `Embedding` e uma camada `SimpleRNN`:
```
from keras.layers import Dense
model = Sequential()
model.add(Embedding(max_features, 32))
model.add(SimpleRNN(32))
model.add(Dense(1, activation='sigmoid'))
model.compile(optimizer='rmsprop', loss='binary_crossentropy', metrics=['acc'])
history = model.fit(input_train, y_train,
epochs=10,
batch_size=128,
validation_split=0.2)
```
Vamos exibir a perda e a precisão do treinamento e da validação:
```
import matplotlib.pyplot as plt
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc))
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
```
Como lembrete, no capítulo 3, nossa primeira abordagem ingênua a esse mesmo conjunto de dados nos levou a 88% de precisão no teste. Infelizmente, nossa pequena rede recorrente não apresenta um desempenho muito bom em comparação com essa linha de base (apenas até 85% de precisão de validação). Parte do problema é que nossas entradas consideram apenas as primeiras 500 palavras e não as seqüências completas - portanto, nossa RNN tem acesso a menos informações do que nosso modelo de linha de base anterior. O restante do problema é simplesmente que o `SimpleRNN` não é muito bom no processamento de longas sequências como texto. Outros tipos de camadas recorrentes têm um desempenho muito melhor. Vamos dar uma olhada em algumas camadas mais avançadas.
## Um exemplo de LSTM em Keras
Agora vamos mudar para preocupações mais práticas: vamos configurar um modelo usando uma camada LSTM e treiná-lo nos dados do IMDB. Aqui está a rede, semelhante à do SimpleRNN que acabamos de apresentar. Especificamos apenas a dimensionalidade de saída da camada LSTM e deixamos todos os outros argumentos (existem muitos) para os padrões do Keras. Keras tem bons padrões, e as coisas quase sempre "simplesmente funcionam" sem você
tendo que gastar tempo ajustando parâmetros manualmente.
```
from keras.layers import LSTM
model = Sequential()
model.add(Embedding(max_features, 32))
model.add(LSTM(32))
model.add(Dense(1, activation='sigmoid'))
model.compile(optimizer='rmsprop',
loss='binary_crossentropy',
metrics=['acc'])
history = model.fit(input_train, y_train,
epochs=10,
batch_size=128,
validation_split=0.2)
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc))
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
```
| github_jupyter |
### Scrapy
- 파이썬 언어를 이용한 웹 데이터 수집 프레임워크
- 프레임워크 vs 라이브러리 또는 패키지
- 프레임워크 : 특정목적을 가진 기능의 코드가 미리 설정되어서 빈칸채우기 식으로 코드를 작성
- 패키지는 다른 사람이 작성해 놓은 코드를 가져다가 사용하는 방법
- scrapy
- `$pip install scrapy`
- tree
- `$sudo apt install tree`
서버에서 설치하기~~
scrapy는 자체적으로 강력한(?) xpath를 이용하기 때문에.
Beautifulsoup을 굳이 쓸 필요없다.-> BeautifulSoup 쓰는 이유는 css-selector 쓰기 위해서.
### 2. Scrapy Project
- scrapy 프로젝트 생성
- scrapy 구조
- gmarket 베스트 상품 링크 수집, 링크 안에 있는 상세 정보 수집
```
# 프로젝트 생성
!scrapy startproject crawler
!ls
!tree crawler
```
#### scrapy의 구조
- spiders
- 어떤 웹서비스를 어떻게 크롤링할것인지에 대한 코드를 작성(.py 파일로 작성)
- items.py
- 모델에 해당하는 코드, 저장하는 데이터의 자료구조를 설정
- piplines.py
- 스크래핑한 결과물을 items 형태로 구성하고 처리하는 방법에 대한 코드
- settings.py
- 스크래핑 할때의 설정값을 지정
- robots.txt : 따를지, 안따를지
#### gmarket 베스트 셀러 상품 수집
- 상품명, 상세페이지 URL, 원가, 판매가, 할인율
- xpath 확인
- items.py
- spider.py
- 크롤러 실행
### 1. xpath 확인
```
import requests
import pandas as pd
import scrapy
from scrapy.http import TextResponse
req = requests.get("http://corners.gmarket.co.kr/Bestsellers")
response = TextResponse(req.url, body=req.text, encoding="utf-8")
items = response.xpath('//*[@id="gBestWrap"]/div/div[3]/div[2]/ul/li')
len(items)
links = response.xpath(
'//*[@id="gBestWrap"]/div/div[3]/div[2]/ul/li/div[1]/a/@href').extract()
len(links)
links[0]
## 결과데이터가 무조건 list라서 [0] 써준다.
req = requests.get(links[0])
response = TextResponse(req.url, body=req.text, encoding="utf-8")
title = response.xpath('//*[@id="itemcase_basic"]/h1/text()')[0].extract()
s_price = response.xpath('//*[@id="itemcase_basic"]/p/span/strong/text()')[0].extract().replace(",", "")
o_price = response.xpath('//*[@id="itemcase_basic"]/p/span/span/text()')[0].extract().replace(",", "")
discount_rate = str(round((1 - int(s_price) / int(o_price))*100, 2)) + "%"
title, s_price, o_price, discount_rate
```
### 2. items.py 작성
- 크롤링할 데이터의 모델을 정해줌.
```
!cat crawler/crawler/items.py
%%writefile crawler/crawler/items.py
import scrapy
class CrawlerItem(scrapy.Item):
title = scrapy.Field()
s_price = scrapy.Field()
o_price = scrapy.Field()
discount_rate = scrapy.Field()
link = scrapy.Field()
```
### 3. spider.py 작성
실제로 작성하는건 요정도밖에 안된다~~
```
%%writefile crawler/crawler/items.py
import scrapy
from crawler.items import CrawlerItem
class Spider(scrapy.Spider):
name = "GmarketBestsellers"
allow_domain = ["gmarket.co.kr"]
start_urls = ["http://corners.gmarket.co.kr/BestSellers"]
def parse(self, response):
links = response.xpath("//*[@id="gBestWrap"]/div/div[3]/div[2]/ul/li")
for link in links[:10]: ##실행하는데 시간이 너무많이 걸려서 links 10개까지만 하자.
yield scrapy.Request(link, callback=self.page_content)
def page_content(self, response):
item = CrawlerItem()
item["title"] = response.xpath('//*[@id="itemcase_basic"]/h1/text()')[0].extract()
item["s_price"] = response.xpath('//*[@id="itemcase_basic"]/p/span/strong/text()')[0].extract().replace(",", "")
try:
item["o_price"] = response.xpath('//*[@id="itemcase_basic"]/p/span/span/text()')[0].extract().replace(",", "")
except:
item["o_price"] = response.xpath('//*[@id="itemcase_basic"]/p/span/span/text()')[0].extract().replace(",", "")
item["discount_rate"] = str(round((1 - int(s_price) / int(o_price))*100, 2)) + "%"
item["link"] = response.url
yield item
```
### 4. Scrapy 실행
```
!ls crawler
## scrapy.cfg 파일이 있는 디렉토리 안에서 다음의 커맨드를 실행해야한다
%%writefile run.sh
cd crawler # crawler 디렉토리로 이동
scrapy crawl GmarketBestsellers # shell script를 실행한다.
# +x : 파일에 대한 실행권한을 추가해준다
!chmod +x run.sh
!./run.sh
```
- 결과를 CSV로 저장
```
%%writefile run.sh
cd crawler
scrapy crawl GmarketBestsellers -o GmarketBestsellers.csv
! ./run.sh
!ls crawler/
## 정상적으로 크롤링되어 데이터가 저장되어 있는지 확인
import pandas as pd
path = !pwd
files = !ls crawler/
files
files = !ls crawler/ # crawler프로젝트 안의 파일을 불러옴.
files
"crawler/{}".format(files[0])
df = pd.read_csv("GmarketBestsellers.csv")
df.tail(2)
```
### 5. Pipelines 설정
- item 을 출력하기 전에 실행되는 코드를 정의
```
import requests
import json
class send_slack(object):
def __send_slack(self, msg):
WEBHOOK_URL = ""
payload = {
"channel": "",
"username": "",
"text": msg,
}
requests.post(WEBHOOK_URL, json.dumps(payload))
def process_item(self, item, spider):
keyword = "세트",
print("="*100)
print(item["title"], keyword)
print("="*100)
if keyword in item["title"]:
self.__send_slack("{},{},{}".format(
item["title"], item["s_price"], item["link"]))
return item #return된 item이 출력된당
##실행해서 Overwriting 해준당
```
- pipeline 설정 : settings.py
```
ITEM_PIPELINES = {
'crawler.pipelines.CrawlerPipeline': 300,
}
```
- 숫자 : 1~1000까지 가능. 파이프라인의 순서 정해줌. 숫자 낮을수록 더 일찍 실행된다.
```
# echo : 뒤에오는 문자열을 출력시켜주는 shell 커맨드
!echo "ITEM_PIPELINES = {" >> crawler/crawler/settings.py
!echo " 'crawler.pipelines.CrawlerPipeline': 300," >> crawler/crawler/settings.py
!echo "}" >> crawler/crawler/settings.py
!tail -n 3 cralwer/crawler/settings.py
#slack?? 으로 전송되는지 확인~~~
!./run.sh
```
| github_jupyter |
# Ranking
## Install packages
```
import sys
!{sys.executable} -m pip install -r requirements.txt
import cvxpy as cvx
import numpy as np
import pandas as pd
import time
import os
import quiz_helper
import matplotlib.pyplot as plt
%matplotlib inline
plt.style.use('ggplot')
plt.rcParams['figure.figsize'] = (14, 8)
```
### data bundle
```
import os
import quiz_helper
from zipline.data import bundles
os.environ['ZIPLINE_ROOT'] = os.path.join(os.getcwd(), '..', '..','data','module_4_quizzes_eod')
ingest_func = bundles.csvdir.csvdir_equities(['daily'], quiz_helper.EOD_BUNDLE_NAME)
bundles.register(quiz_helper.EOD_BUNDLE_NAME, ingest_func)
print('Data Registered')
```
### Build pipeline engine
```
from zipline.pipeline import Pipeline
from zipline.pipeline.factors import AverageDollarVolume
from zipline.utils.calendars import get_calendar
universe = AverageDollarVolume(window_length=120).top(500)
trading_calendar = get_calendar('NYSE')
bundle_data = bundles.load(quiz_helper.EOD_BUNDLE_NAME)
engine = quiz_helper.build_pipeline_engine(bundle_data, trading_calendar)
```
### View Data¶
With the pipeline engine built, let's get the stocks at the end of the period in the universe we're using. We'll use these tickers to generate the returns data for the our risk model.
```
universe_end_date = pd.Timestamp('2016-01-05', tz='UTC')
universe_tickers = engine\
.run_pipeline(
Pipeline(screen=universe),
universe_end_date,
universe_end_date)\
.index.get_level_values(1)\
.values.tolist()
universe_tickers
```
# Get Returns data
```
from zipline.data.data_portal import DataPortal
data_portal = DataPortal(
bundle_data.asset_finder,
trading_calendar=trading_calendar,
first_trading_day=bundle_data.equity_daily_bar_reader.first_trading_day,
equity_minute_reader=None,
equity_daily_reader=bundle_data.equity_daily_bar_reader,
adjustment_reader=bundle_data.adjustment_reader)
```
## Get pricing data helper function
```
def get_pricing(data_portal, trading_calendar, assets, start_date, end_date, field='close'):
end_dt = pd.Timestamp(end_date.strftime('%Y-%m-%d'), tz='UTC', offset='C')
start_dt = pd.Timestamp(start_date.strftime('%Y-%m-%d'), tz='UTC', offset='C')
end_loc = trading_calendar.closes.index.get_loc(end_dt)
start_loc = trading_calendar.closes.index.get_loc(start_dt)
return data_portal.get_history_window(
assets=assets,
end_dt=end_dt,
bar_count=end_loc - start_loc,
frequency='1d',
field=field,
data_frequency='daily')
```
## get pricing data into a dataframe
```
returns_df = \
get_pricing(
data_portal,
trading_calendar,
universe_tickers,
universe_end_date - pd.DateOffset(years=5),
universe_end_date)\
.pct_change()[1:].fillna(0) #convert prices into returns
returns_df
```
## Sector data helper function
We'll create an object for you, which defines a sector for each stock. The sectors are represented by integers. We inherit from the Classifier class. [Documentation for Classifier](https://www.quantopian.com/posts/pipeline-classifiers-are-here), and the [source code for Classifier](https://github.com/quantopian/zipline/blob/master/zipline/pipeline/classifiers/classifier.py)
```
from zipline.pipeline.classifiers import Classifier
from zipline.utils.numpy_utils import int64_dtype
class Sector(Classifier):
dtype = int64_dtype
window_length = 0
inputs = ()
missing_value = -1
def __init__(self):
self.data = np.load('../../data/project_4_sector/data.npy')
def _compute(self, arrays, dates, assets, mask):
return np.where(
mask,
self.data[assets],
self.missing_value,
)
sector = Sector()
```
## We'll use 2 years of data to calculate the factor
**Note:** Going back 2 years falls on a day when the market is closed. Pipeline package doesn't handle start or end dates that don't fall on days when the market is open. To fix this, we went back 2 extra days to fall on the next day when the market is open.
```
factor_start_date = universe_end_date - pd.DateOffset(years=2, days=2)
factor_start_date
```
## Explore the rank function
The Returns class inherits from zipline.pipeline.factors.factor.
[The documentation for rank is located here](https://www.zipline.io/appendix.html#zipline.pipeline.factors.Factor.rank), and is also pasted below:
```
rank(method='ordinal', ascending=True, mask=sentinel('NotSpecified'), groupby=sentinel('NotSpecified'))[source]
Construct a new Factor representing the sorted rank of each column within each row.
Parameters:
method (str, {'ordinal', 'min', 'max', 'dense', 'average'}) – The method used to assign ranks to tied elements. See scipy.stats.rankdata for a full description of the semantics for each ranking method. Default is ‘ordinal’.
ascending (bool, optional) – Whether to return sorted rank in ascending or descending order. Default is True.
mask (zipline.pipeline.Filter, optional) – A Filter representing assets to consider when computing ranks. If mask is supplied, ranks are computed ignoring any asset/date pairs for which mask produces a value of False.
groupby (zipline.pipeline.Classifier, optional) – A classifier defining partitions over which to perform ranking.
Returns:
ranks – A new factor that will compute the ranking of the data produced by self.
Return type:
zipline.pipeline.factors.Rank
```
## Quiz 1
Create a factor of one year returns, demeaned, and ranked
## Answer 1 here
```
from zipline.pipeline.factors import Returns
#TODO
# create a pipeline called p
p = Pipeline(screen=universe)
# create a factor of one year returns, deman by sector, then rank
factor = (
Returns(window_length=252, mask=universe).
demean(groupby=Sector()). #we use the custom Sector class that we reviewed earlier
rank()
)
# add the factor to the pipeline
p.add(factor, 'Momentum_1YR_demean_by_sector_ranked')
```
## visualize the pipeline
```
p.show_graph(format='png')
```
## run pipeline and view the factor data
```
df = engine.run_pipeline(p, factor_start_date, universe_end_date)
df.head()
```
## Quiz 2
What do you notice about the factor values?
## Answer 2 here:
The factor values are now integers from 1 to N, where N is the number of stocks in the universe.
## Solution notebook
[The solution notebook is here.](rank_solution.ipynb)
| github_jupyter |
The objective of this short pipeline is to design a recurrent neural network (RNN) with the Long Short Term Memory (LSTM) Model to predict future market closing prices for the Dow Jones Industrial Average (DJIA) through a closed technical analysis approach. It attempts to draw market insights by leveraging underlying relationships between price, information and volume for the index.
<h2> Load Requirements </h2>
```
import pandas as pd
import numpy as np
import scipy as sp
import pickle
import gc
import requests
import urllib.request
import psutil
import sklearn.ensemble
import sklearn.preprocessing
import sklearn.metrics
import matplotlib.pyplot as plt
import seaborn as sns
import tensorflow as tf
import keras
from keras.callbacks import EarlyStopping, ModelCheckpoint
from keras.wrappers.scikit_learn import KerasRegressor
from keras.models import Sequential, load_model
from keras.layers import Dense, Dropout, LSTM, RNN
djiadata = pd.read_csv('https://www.alphavantage.co/query?function=TIME_SERIES_DAILY&symbol=DJIA&apikey='+
'1D9IC2FZ6E6ABYLY&datatype=csv&outputsize=full')
with open('djia_historic_data.csv', 'wb') as f:
f.write(djiadata.to_csv(index=False).encode())
djiadata.info()
```
<h2> Prepare Data </h2>
<b> STEP 1: Clean Dataset </b>
A fine balance must be established between training a network on excessively historic data such that it loses predictive power on current trends, vis-a-vis limited short-term market information. Therefore, it was decided to train the network on data from 2008-2017, test it on 2018-19, and run future predictions. In addition, we are attempting a 'many-to-one' network, effectively trying to predict only day-end closing prices for the index.
```
djiadata = djiadata.iloc[::-1].reset_index(drop=True)
djiadata.drop(['open', 'high', 'low', 'volume'], axis=1, inplace=True)
djiadata.timestamp = djiadata.timestamp.apply(lambda s: int(''.join(s.split('-'))))
djiadata.drop(djiadata[djiadata.timestamp < 20080000].index.values, axis=0, inplace=True)
djiadata.reset_index(drop=True, inplace=True)
```
<b> STEP 2: Add Indicator Features </b>
In order to effectively uncover and dissect market trends, additional indicators and oscillators are added to DJIA data obtained above. The choice of indicators was difficult, mostly to only include tools that are known from constricted present data, and thus moving averages (simple and exponential), bollinger bands, simplified aaron oscillator and change momentum oscillators are used. We engineer said features on input dataset, and proceed to model development.
```
#19-Day SMA
djiadata['sma19'] = np.cumsum(djiadata['close'])
djiadata.sma19 = (djiadata.sma19 - djiadata.sma19.shift(19))/19
djiadata.loc[djiadata.index.values==18, 'sma19'] = np.mean(djiadata.close[:19])
#19-Day (Double-Weighted) EMA
djiadata['ema19'] = np.nan
djiadata.loc[:19, 'ema19'] = djiadata.loc[:19, 'sma19']
for i in range(19, len(djiadata)):
djiadata.loc[i,'ema19']=(djiadata.loc[i,'close']-djiadata.loc[i-1,'ema19'])*0.1 + djiadata.loc[i-1,'ema19']
#19-Day Double-STD Bollinger Bands
lstcls = djiadata['close'].astype('str')
for i in range(1, 19):
lstcls += ' ' + djiadata['close'].astype('str').shift(i)
stddevval = lstcls.apply(lambda s: s if str(s)=='nan' else np.std(np.vectorize(lambda g: float(g))(s.split())))
djiadata['upperbol'] = djiadata['sma19'] + 2 * stddevval
djiadata['lowerbol'] = djiadata['sma19'] - 2 * stddevval
#Simplified Aaron Oscillator
#lstcls is currently at 19-day
for i in range(19,25):
lstcls += ' ' + djiadata['close'].astype('str').shift(i)
aarminmax = lstcls.apply(lambda s: s if str(s)=='nan' else np.vectorize(lambda g: float(g))(s.split()))
aarminmax = aarminmax.apply(lambda s: s if str(s)=='nan' else 100*(25-np.argmax(s))/25 - 100*(25-np.argmin(s))/25)
djiadata['aaronval'] = aarminmax
#25-Period Chande Momentum Oscillator
djiadata['highc'] = 0
djiadata['lowc'] = 0
djiadata.loc[djiadata.close < djiadata.close.shift(1), 'lowc'] = djiadata['close']
djiadata.loc[djiadata.close > djiadata.close.shift(1), 'highc'] = djiadata['close']
djiadata['sumhighc'] = np.cumsum(djiadata['highc'])
djiadata['sumhighc'] = djiadata['sumhighc'] - djiadata['sumhighc'].shift(25)
djiadata['sumlowc'] = np.cumsum(djiadata['lowc'])
djiadata['sumlowc'] = djiadata['sumlowc'] - djiadata['sumlowc'].shift(25)
djiadata.loc[djiadata.index==24, ['sumhighc','sumlowc']] = [np.sum(djiadata.highc[:25]), np.sum(djiadata.lowc[:25])]
djiadata['chande'] = (djiadata['sumhighc']-djiadata['sumlowc'])/(djiadata['sumhighc']+djiadata['sumlowc']) * 100
djiadata.drop(['highc', 'lowc', 'sumhighc', 'sumlowc'], axis=1, inplace=True)
djiadata = djiadata.dropna(axis=0)
djiadata.reset_index(drop=True, inplace=True)
```
<b> STEP 3: Data Scaling </b>
We must scale all data down to a specified feature range so that it is significantly easier for the model to learn underlying data trends. In interest of the same, it was decided to min-max scale each feature to [0,1]. This choice was primarily driven by the lack of outliers in the data that nullify the robustness advantage of standardization, along with the simplicity of scaling back and forth based on minima and maxima.
```
sc = sklearn.preprocessing.MinMaxScaler(feature_range=(0,1), copy=True)
cols = djiadata.columns.values[1:]
djiadata[cols] = pd.DataFrame(sc.fit_transform(djiadata[cols]), columns=cols)
with open('scaler.pkl', 'wb') as f:
pickle.dump(sc, f)
```
<b> STEP 3: Design Input for Recurrent Neural Network
For our LSTM-based Recurrent Neural Network, we decide to have 50 time-stamps at a time, allowing the model to predict the 51st closing price. The rationale behind choosing a 50-time frame was to balance between short-term trends and long-term market indicators, whilst ensuring that all 19- and 25-day oscillators were adequately represented in each model training sequence. Thus, each time-stamp is a 7-element numpy array containing the current closing price, 19-day simple and exponential moving averages, upper and lower bollinger bands, as well as a simplified aaron and chande momentum oscillator, in said order. We thus convert our dataset into independent and response variable sets.
```
time_stamp_num = 50
Y = djiadata.close[time_stamp_num:].values
rawdata = djiadata.values[:, 1:]
X = np.zeros((len(djiadata)-time_stamp_num, time_stamp_num, 7), dtype='float')
for i in range(len(djiadata)-time_stamp_num):
X[i, :, :] = rawdata[i:i+time_stamp_num]
X.shape, Y.shape
```
<h2> Design Recurrent Neural Network </h2>
<b> STEP 1: Separate Train and Test Sets </b>
```
#2008-2018 is the training set, and 2019 YTD is the test set
#Define the number of entries in the test set
sum(djiadata.timestamp > 20180000)
testsize = sum(djiadata.timestamp > 20180000)
trainx = X[:-testsize, :, :]
testx = X[-testsize:, :, :]
trainy = Y[:-testsize]
testy = Y[-testsize:]
trainx.shape, testx.shape, trainy.shape, testy.shape
```
<b> STEP 2: Build and Train Network </b>
When designing the model's architecture, LSTM cells were chosen over simple RNN cells available through Tensorflow in order to accomodate the model's ability to learn long-term dependencies in the time-series data. Multiple layers of LSTM cells were decided to be stacked for greater memory retention. The exact architecture planned was as follows:
1) First Layer of LSTM cells, returns sequence to next
2) Second layer of LSTM cells, returns sequence to next
3) Third layer of LSTM cells, returns last-value for time-stamps
4) Dense (fully-connected) layer returning the actual 51st period closing price
```
def mdlbdr():
model = Sequential()
model.add(LSTM(units=50, return_sequences=True, input_shape=(50, 7)))
model.add(LSTM(units=50, return_sequences=True))
model.add(LSTM(units=50))
model.add(Dense(units=25, activation='relu'))
model.add(Dense(units=1))
model.compile(optimizer='adam', loss='mean_squared_error')
return model
chk = ModelCheckpoint('./djiarnn2.h5', monitor='loss', save_best_only=True, period=10)
callbacklist = [chk]
djmdl = KerasRegressor(build_fn=mdlbdr, epochs=100, batch_size=30, verbose=True, callbacks=callbacklist,
validation_data=(testx, testy))
djmdl.fit(trainx, trainy)
djmdl.model.save('./djiarnn2.h5')
```
<b> STEP 3: Run Test Data </b>
```
Ypred = djmdl.predict(testx)
tmp = np.concatenate([Ypred.reshape(-1,1), np.zeros((len(Ypred), 6))], axis=1)
Ypredorig = sc.inverse_transform(tmp)[:, 0]
tmpt = np.concatenate([testy.reshape(-1,1), np.zeros((len(testy), 6))], axis=1)
testyorig = sc.inverse_transform(tmpt)[:, 0]
sklearn.metrics.mean_absolute_error(testyorig, Ypredorig)
fig = plt.figure(figsize=(10,7))
ax = fig.add_subplot(111)
plt.plot(Ypredorig)
plt.plot(testyorig)
plt.title('DJIA Predictions on Test Data (2018 + 2019 YTD)')
plt.xlabel('Time (Oldest -> Newest)')
plt.ylabel('DJIA Price')
plt.show()
fig = plt.figure(figsize=(10,7))
ax = fig.add_subplot(111)
plt.plot(Ypredorig)
plt.plot(testyorig)
plt.title('DJIA Predictions on Test Data (2018 + 2019 YTD)')
plt.xlabel('Time (Oldest -> Newest)')
plt.ylabel('DJIA Price')
plt.show()
```
<b> STEP 4: Build September Future-Prediction Algorithm </b>
```
djiadata
```
| github_jupyter |
# Grunnleggende Python
## Skrive ut i terminalen
Når du programmerer i Python brukes funksjonen `print()` til å skrive ut i terminalen. Det som skal printes må legges inni parentesen.
```
print(2)
```
Du kan skrive ut flere ting i terminalen ved å skille med komma.
```
print(0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10)
```
Hvis du skal printe tekst, må du ramme inn teksten med fnutter ("), slik som i eksempelet under.
```
print("This message will be printed")
```
## Variabler
I Python kan man deklarere variabler som vist i kodesnutten under.
Merk at det er venstresiden som settes til å være lik høyresiden. I motsetning til matte, er det derfor "lov" til å f.eks sette variabler lik seg selv pluss noe, slik som vist under.
Merk at `b` ikke endrer verdi når du endrer `a` *etter* å ha deklarert `b`.
**Husk at Python leser filen fra øverste linje og nedover.** Derfor MÅ man deklarer `a` før `b`i kodesnutten under.
```
a = 2
print("1: a =", a)
b = a + 1
print("2: b =", b)
a = a + 2
print("3: a =", a)
print("4: b =", b)
```
### Variabelnavn
Python dømmer ikke særlig på hvilke variabelnavn du bruker. Kriteriene til et variabelnavn er
- Variabelnavn kan inneholde alle bokstaver, både store og små, som finnes i det **engelske alfabetet**. Æ, ø og å fungerer i Python 3, men konvensjon er å ikke bruke disse.
- Variabelnavn kan inneholde **understrek**
- Variabelnavn kan inneholde tall, men **et tall kan ikke være første tegn i navnet**
- Ellers er ingen andre tegn lov i variabelnavn
Disse reglene er illustrert under, med et fryktelig langt variabelnavn:
```
VARIABLE_NAME_with_1_2_3 = 0
```
Som sagt, Python dømmer ikke særlig på variabelnavn. Men du bør likevel lage hensiktsmessige variabelnavn for å øke lesbarheten til koden din. Variabelnavn bør ikke være kjempelang, men likevel være beskrivende nok til at andre (eller deg i fremtiden) skal kunne forstå hva variabelen er.
Når variabelen er deklarert, kan den brukes!
```
pi = 3.14
print("pi = ", pi)
```
## Python som kalkulator
Slik gjør du enkel matte i Python:
$\Large{\cdot}$ $a - b$ i Python : `a - b`
$\Large{\cdot}$ $a \times b$ i Python : `a*b`
$\Large{\cdot}$ $\frac{a}{b}$ i Python : `a/b`
$\Large{\cdot}$ $a^b$ i Python : `a**b`
$\Large{\cdot}$ $3.2 \times 10^7$ i Python : `3.2e7`
Pass på at du brukes parenteser når du gjør utregningene dine! Se nøye på eksemplene under.
```
a = 1
b = 2
c = 3
print("2 + 3/3 = ", (b + c/c))
print("(2 + 3)/3 = ", ((b + c)/c))
print("3/2 + 1 = ", (c/b + a))
print("3/(2 + 1) = ", (c/(b + a)))
```
## Datatyper
Variabler kan være mye mer enn bare heltall. Noen av de enkleste typene vil bli gjennomgått. Du kan bruke `type()` for å sjekke hvilken datatype en variabel er. Eksempler på dette vil bli vist.
### Integers (Heltall)
Heltall i Python heter *integer*, men betegnes som `int`.
Heltall i Python 2 og Python 3 oppfører seg litt annerledes. I Python 2 vil heltall gi heltallsdivisjon, som er vanlig i programmeringsspråk. Dette betyr at når du deler to heltall, så vil svaret også bli et heltall. Altså blir $3/2 = 1$. I Python 3 derimot, vil to heltall kunne gi et desimall tall, slik at $3/2 = 1,5$.
```
print(type(2))
```
### Float (desimaltall)
Desimaltall i Python heter `floating point number`, men betegnes som `float`.
```
print(type(1.2))
```
### String (tekst)
Tekst i Python heter *String*, men betegnes som `str`.
Lurte du på hvorfor du trengte fnutter for å printe ut tekst? Hvis ikke dette gjøres vil Python tro at verdien til en variabel med det navnet skal printes. Fnuttene spesifiserer at det *ikke* er snakk om et variabelnavn (eller andre typer, men det ignoreres foreløpig), men en `String`.
```
print(type("spam"))
```
Under er det vist feilmeldingen som oppstår av at en Pythonista ikke har spesifisert at en `String` skulle printes. Det gjelder å lære seg at feilmeldinger er en Pythonista sin beste venn. Her sier feilmeldingen tydelig at `spam is not defined`.
```
print(spam)
```
### <span style="color:green"> Test deg selv </span>
Kan du endre på variablene under slik at alle tre variabler inneholder verdien 2, men er av forskjellige datatyper?
```
int_2 = ?
print(type(int_2))
float_2 = ?
print(type(float_2))
str_2 = ?
print(type(str_2))
```
## Import av bibliotek
En modul er en samling ferdig skrevne funksjoner. Et eksempel er `math` som inneholder både variabler og funksjoner. Ved å skrive `import <module_name>`, hentes modulen slik at innholdet kan brukes i koden din. For å bruke funksjonene fra modulen, må man presisere at de er fra modulen. Dette gjøres ved å skrive `<module_name>.<function_name>`, som vist under.
```
import math
a = math.exp(2)
```
Hvis man ikke skal bruke altfor mange funksjoner fra en modul, vil det være kortere å spesifisere i `import`-linjen hva som skal brukes. Dette gjøres ved å skrive `from <module_name> import <function_name_1>, <function_name_2>`. Hvis du henter mer enn én funksjon fra modulen, skiller du dem med komma. Under ligger et eksempel.
```
from math import pi, sin
b = sin(0.5*pi)
```
## Kommentering av kode
For å kommentere koden kan du enten bruke `#` hvis kommentaren kun går over én linje, eller `'''` før og og etter kommentaren om den går over flere linjer.
```
# This is how to make a comment on a single line
a = 13 # Can be used on the right side of your code
''' This is how to
comment over
several
lines '''
```
## Formatering
For å inkludere variabler i en String, brukes %-tegnet. %-tegnet symboliserer at det skal komme inn en variabel, som er gitt etter Stringen. Etterfulgt av %-tegnet kommer et tegn som viser hvilken type variabel som skal plasseres der. **En tabell med format-symboler finner du på side 10 i læreboken.** Etter Stringen er avsluttet kommer et nytt %-tegn etterfulgt av variablene som skal inn. Hvis det er flere variabler, separeres disse med komma.
Et eksempel:
%f symboliserer at det kommer en `float` (tall med desimaler). Med denne kan du bestemme både hvor mye plass variabelen skal oppta, og hvor mange desimaler som skal tas med. Dette gjøres ved å skrive
`" %<total_space>.<decimals>f " % variable`
I eksempelet under er det skrevet %<span style="color:red">.4</span><span style="color:blue">f</span> for å presisere at det er en <span style="color:blue">float</span> og at det tallet skal inkludere <span style="color:red">fire desimaler</span>. Den neste er tilsvarende, %.2f presiserer `float` med to desimaler. **Merk at rekkefølgen på variablene som er satt inn bak bestemmer rekkefølgen de vil vises i teksten.**
Under er det printet ut 1/3 uten å formatere. Ser du forskjellen?
```
a_third = 1./3
a_fourth = 1./4
print("1/3 = %.4f and 1/4 = %.2f" %(a_third, a_fourth))
print("1/3 = ", a_third)
```
## Kjøreeksempel
Et kjøreeksempel er en kopi av det som skrives ut i terminalvinduet når du kjører koden. Copy-paste nederest i .py-filen før du leverer.
```
print("Hello world")
'''
eirillsh$ python hello_world.py
Hello world
'''
```
| github_jupyter |
# Indonesian Text Classification
This is a text classification of indonesian corpus using several different technique such as Naive Bayes, SVM, Random Forest, Convolition Neural Network (CNN), LSTM or GRU. An indonesian [pre-trained word vectors](https://fasttext.cc/docs/en/pretrained-vectors.html) from FastText has ben also used in our Neural Network models.
We use [Word Bahasa Indonesia Corpus and Parallel English Translation](https://www.panl10n.net/english/outputs/Indonesia/BPPT/0902/BPPTIndToEngCorpusHalfM.zip) dataset from PAN Localization.
It contains 500,000 words from various online sources translated into English.
For our text classification, we use only the indonesian part.
The corpus has 4 classes:
- 0: Economy
- 1: International
- 2: Science
- 3: Sport
Originally each class is in separate file, we combine, randomize and split it to train and test file with 90:10.
```
!pip install xgboost
from sklearn import model_selection, preprocessing
from sklearn import linear_model, naive_bayes, metrics, svm
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer
from sklearn import decomposition, ensemble
import tensorflow as tf
#import pandas as pd, xgboost, numpy, textblob, string
import pandas as pd, xgboost, numpy, string
from tensorflow.keras.preprocessing import text, sequence
from tensorflow.keras import layers, models, optimizers
from keras.preprocessing.sequence import pad_sequences
from pathlib import Path
import os
import numpy as np
#import ntlk
# Parameters
#LMDATA = Path('/content/drive/My Drive/lmdata')
LMDATA = Path('lmdata/id/dataset')
params = {'batch_size': 1024,
'n_classes': 2,
'max_len': 100,
'n_words': 50000,
'shuffle': True}
try:
TPU_WORKER = 'grpc://' + os.environ['COLAB_TPU_ADDR']
strategy=tf.contrib.tpu.TPUDistributionStrategy(
tf.contrib.cluster_resolver.TPUClusterResolver(TPU_WORKER))
except KeyError:
TPU_WORKER = None
np.random.seed(seed=10)
# Load the Drive helper and mount
from google.colab import drive
# This will prompt for authorization.
drive.mount('/content/drive')
!pwd
print(LMDATA)
!ls -lh "$LMDATA"
!ls -lh "$LMDATA/BPPTIndToEngCorpus"
train_df = pd.read_csv(LMDATA/'BPPTIndToEngCorpus/bppt_panl_train.csv')
train_df.columns = ['label', 'text']
test_df = pd.read_csv(LMDATA/'BPPTIndToEngCorpus/bppt_panl_test.csv')
test_df.columns = ['label', 'text']
print(train_df.head())
#print(train_df['label'][:10].values)
#print(train_df['text'][:10].values)
!set |grep -i tpu|grep -v grep
# split the dataset into training and validation datasets
train_x, valid_x, train_y, valid_y = model_selection.train_test_split(train_df['text'], train_df['label'])
# label encode the target variable
encoder = preprocessing.LabelEncoder()
train_y = encoder.fit_transform(train_y)
valid_y = encoder.fit_transform(valid_y)
print(train_x[:5])
print(train_y[:5])
print(valid_x[:5])
print(valid_y[:5])
# create a count vectorizer object
count_vect = CountVectorizer(analyzer='word', token_pattern=r'\w{1,}')
count_vect.fit(train_df['text'])
# transform the training and validation data using count vectorizer object
xtrain_count = count_vect.transform(train_x)
xvalid_count = count_vect.transform(valid_x)
# word level tf-idf
tfidf_vect = TfidfVectorizer(analyzer='word', token_pattern=r'\w{1,}',
max_features=5000)
tfidf_vect.fit(train_df['text'])
xtrain_tfidf = tfidf_vect.transform(train_x)
xvalid_tfidf = tfidf_vect.transform(valid_x)
# ngram level tf-idf
tfidf_vect_ngram = TfidfVectorizer(analyzer='word', token_pattern=r'\w{1,}',
ngram_range=(2,3), max_features=5000)
tfidf_vect_ngram.fit(train_df['text'])
xtrain_tfidf_ngram = tfidf_vect_ngram.transform(train_x)
xvalid_tfidf_ngram = tfidf_vect_ngram.transform(valid_x)
# characters level tf-idf
tfidf_vect_ngram_chars = TfidfVectorizer(analyzer='char', token_pattern=r'\w{1,}',
ngram_range=(2,3), max_features=5000)
tfidf_vect_ngram_chars.fit(train_df['text'])
xtrain_tfidf_ngram_chars = tfidf_vect_ngram_chars.transform(train_x)
xvalid_tfidf_ngram_chars = tfidf_vect_ngram_chars.transform(valid_x)
# load the pre-trained word-embedding vectors
max_words = params['n_words']
embeddings_index = {}
for i, line in enumerate(open(LMDATA/'wiki.id.300K.vec')):
if i%50000 == 0:
print(i)
values = line.split()
try:
embeddings_index[" ".join(values[0:-300])] = numpy.asarray(values[-300:], dtype='float32')
except ValueError:
print("Values: {}: {}".format(i, values))
# create a tokenizer
token = text.Tokenizer()
token.fit_on_texts(train_df['text'])
word_index = token.word_index
# convert text to sequence of tokens and pad them to ensure equal length vectors
train_seq_x = sequence.pad_sequences(token.texts_to_sequences(train_x),
maxlen=params['max_len'])
valid_seq_x = sequence.pad_sequences(token.texts_to_sequences(valid_x),
maxlen=params['max_len'])
# create token-embedding mapping
embedding_matrix = numpy.zeros((len(word_index) + 1, 300))
for word, i in word_index.items():
if i>=max_words:
break
embedding_vector = embeddings_index.get(word)
if embedding_vector is not None:
embedding_matrix[i] = embedding_vector
print(LMDATA)
```
## Konventional Machine Learning
```
def train_model(classifier, feature_vector_train, label, feature_vector_valid,
is_neural_net=False, epochs=1):
# fit the training dataset on the classifier
if is_neural_net:
classifier.fit(feature_vector_train, label, epochs=epochs)
else:
classifier.fit(feature_vector_train, label)
# predict the labels on validation dataset
predictions = classifier.predict(feature_vector_valid)
if is_neural_net:
predictions = [int(round(p[0])) for p in predictions]
#predictions = predictions.argmax(axis=-1)
print(" predictions:", predictions[:20])
print("ground truth:", valid_y[:20])
return metrics.accuracy_score(predictions, valid_y)
```
### Naive Bayes
```
# Naive Bayes on Count Vectors
accuracy = train_model(naive_bayes.MultinomialNB(), xtrain_count, train_y, xvalid_count)
print("NB, Count Vectors: ", accuracy)
# Naive Bayes on Word Level TF IDF Vectors
accuracy = train_model(naive_bayes.MultinomialNB(), xtrain_tfidf, train_y, xvalid_tfidf)
print("NB, WordLevel TF-IDF: ", accuracy)
# Naive Bayes on Ngram Level TF IDF Vectors
accuracy = train_model(naive_bayes.MultinomialNB(), xtrain_tfidf_ngram, train_y, xvalid_tfidf_ngram)
print("NB, N-Gram Vectors: ", accuracy)
# Naive Bayes on Character Level TF IDF Vectors
accuracy = train_model(naive_bayes.MultinomialNB(), xtrain_tfidf_ngram_chars, train_y, xvalid_tfidf_ngram_chars)
print("NB, CharLevel Vectors: ", accuracy)
```
### Linear Classifier
```
# Linear Classifier on Count Vectors
accuracy = train_model(linear_model.LogisticRegression(), xtrain_count, train_y, xvalid_count)
print("LR, Count Vectors: ", accuracy)
# Linear Classifier on Word Level TF IDF Vectors
accuracy = train_model(linear_model.LogisticRegression(), xtrain_tfidf, train_y, xvalid_tfidf)
print("LR, WordLevel TF-IDF: ", accuracy)
# Linear Classifier on Ngram Level TF IDF Vectors
accuracy = train_model(linear_model.LogisticRegression(), xtrain_tfidf_ngram, train_y, xvalid_tfidf_ngram)
print( "LR, N-Gram Vectors: ", accuracy)
# Linear Classifier on Character Level TF IDF Vectors
accuracy = train_model(linear_model.LogisticRegression(), xtrain_tfidf_ngram_chars, train_y, xvalid_tfidf_ngram_chars)
print("LR, CharLevel Vectors: ", accuracy)
```
### SVM
```
# SVM on Ngram Level TF IDF Vectors
# It seems there is something wrong here, since the predictions are always 0
accuracy = train_model(svm.SVC(), xtrain_tfidf_ngram, train_y, xvalid_tfidf_ngram)
print("SVM, N-Gram Vectors: ", accuracy)
```
### Random Forest
```
# RF on Count Vectors
accuracy = train_model(ensemble.RandomForestClassifier(), xtrain_count, train_y, xvalid_count)
print("RF, Count Vectors: ", accuracy)
# RF on Word Level TF IDF Vectors
accuracy = train_model(ensemble.RandomForestClassifier(), xtrain_tfidf, train_y, xvalid_tfidf)
print("RF, WordLevel TF-IDF: ", accuracy)
```
### Extreme Gradient Boosting
```
# Extreme Gradient Boosting on Count Vectors
accuracy = train_model(xgboost.XGBClassifier(), xtrain_count.tocsc(), train_y, xvalid_count.tocsc())
print("Xgb, Count Vectors: ", accuracy)
# Extreme Gradient Boosting on Word Level TF IDF Vectors
accuracy = train_model(xgboost.XGBClassifier(), xtrain_tfidf.tocsc(), train_y, xvalid_tfidf.tocsc())
print("Xgb, WordLevel TF-IDF: ", accuracy)
# Extreme Gradient Boosting on Character Level TF IDF Vectors
accuracy = train_model(xgboost.XGBClassifier(), xtrain_tfidf_ngram_chars.tocsc(), train_y, xvalid_tfidf_ngram_chars.tocsc())
print("Xgb, CharLevel Vectors: ", accuracy)
```
## Neural Network
```
from tensorflow.keras.utils import Sequence, to_categorical
def tokenize(texts, n_words=1000):
tokenizer = Tokenizer(num_words=n_words)
tokenizer.fit_on_texts(texts)
return tokenizer
class DataGenerator(Sequence):
'Generates data for Keras'
def __init__(self, texts, labels, tokenizer, batch_size=32, max_len=100,
n_classes=2, n_words=1000, shuffle=True):
'Initialization'
self.max_len = max_len
self.batch_size = batch_size
self.texts = texts
self.labels = labels
self.n_classes = n_classes
self.shuffle = shuffle
self.tokenizer = tokenizer
self.steps_per_epoch = int(np.floor(self.texts.size / self.batch_size))
self.on_epoch_end()
def __len__(self):
'Denotes the number of batches per epoch'
return self.steps_per_epoch
def __getitem__(self, index):
'Generate one batch of data'
# Generate indexes of the batch
indexes = self.indexes[index*self.batch_size:(index+1)*self.batch_size]
texts = np.array([self.texts[k] for k in indexes])
sequences = self.tokenizer.texts_to_sequences(texts)
X = pad_sequences(sequences, maxlen=self.max_len)
y = np.array([to_categorical(self.labels[k], 4) for k in indexes])
return X, y
def on_epoch_end(self):
'Updates indexes after each epoch'
self.indexes = np.arange(self.texts.size)
if self.shuffle == True:
np.random.shuffle(self.indexes)
# Create data generator
training_generator = DataGenerator(train_x.values, train_y, token, **params)
valid_generator = DataGenerator(valid_x.values, valid_y, token, **params)
def tpu_wrapper(func):
if TPU_WORKER is not None:
tpu_model = tf.contrib.tpu.keras_to_tpu_model(func, strategy)
return tpu_model
else:
return func
# This model doesn't work with current dataset
def create_simple_model(input_length=100):
# create input layer
input_layer = layers.Input((input_length, ), sparse=True)
# create hidden layer
hidden_layer = layers.Dense(100, activation="relu", name="D1")(input_layer)
# create output layer
output_layer = layers.Dense(4, activation="softmax")(hidden_layer)
classifier = models.Model(inputs = input_layer, outputs = output_layer)
classifier.compile(optimizer=optimizers.Adam(), loss='categorical_crossentropy')
return classifier
#classifier = create_simple_model(xtrain_tfidf_ngram.shape[1])
#accuracy = train_model(classifier, xtrain_tfidf_ngram, train_y, xvalid_tfidf_ngram,
# is_neural_net=True, epochs=1)
#print("NN, Ngram Level TF IDF Vectors", accuracy)
%%time
#NN, Ngram Level TF IDF Vectors
tf.keras.backend.clear_session()
classifier = create_simple_model(input_length=params['max_len'])
classifier = tpu_wrapper(classifier)
classifier.fit_generator(
generator=training_generator,
validation_data=valid_generator,
#use_multiprocessing=True,
#workers=6,
epochs=20
)
classifier.save_weights(str(LMDATA/'bard.h5'), overwrite=True)
xtrain_tfidf_ngram.shape
"""
def to_tpu(func):
def wrapper(*args, **kwargs):
print("TPU Wrapper start")
print(func)
if TPU_WORKER is not None:
tpu_model = tf.contrib.tpu.keras_to_tpu_model(func, strategy)
print("TPU exist")
return tpu_model(*args, **kwargs)
else:
print("TPU not exist")
return func(*args, **kwargs)
print("TO_TPU")
return wrapper
"""
```
### CNN
```
def create_cnn(input_length=100):
# Add an Input Layer
input_layer = layers.Input((input_length, ))
# Add the word embedding Layer
embedding_layer = layers.Embedding(len(word_index) + 1, 300,
weights=[embedding_matrix],
trainable=False)(input_layer)
embedding_layer = layers.SpatialDropout1D(0.3)(embedding_layer)
# Add the convolutional Layer
conv_layer = layers.Convolution1D(100, 3, activation="relu")(embedding_layer)
# Add the pooling Layer
pooling_layer = layers.GlobalMaxPool1D()(conv_layer)
# Add the output Layers
output_layer1 = layers.Dense(100, activation="relu")(pooling_layer)
output_layer1 = layers.Dropout(0.3)(output_layer1)
output_layer2 = layers.Dense(4, activation="softmax")(output_layer1)
# Compile the model
model = models.Model(inputs=input_layer, outputs=output_layer2)
model.compile(optimizer=optimizers.Adam(),
loss='categorical_crossentropy',
metrics=['accuracy'])
return model
%%time
tf.keras.backend.clear_session()
classifier = create_cnn(input_length=params['max_len'])
classifier = tpu_wrapper(classifier)
classifier.fit_generator(
generator=training_generator,
validation_data=valid_generator,
#use_multiprocessing=True,
#workers=6,
epochs=20
)
classifier.save_weights(str(LMDATA/'bard.h5'), overwrite=True)
classifier.save_weights(str(LMDATA/'cnn.h5'), overwrite=True)
# read test dataset
test_x = test_df['text'].values
sequences = token.texts_to_sequences(test_x)
test_x_seq = pad_sequences(sequences, maxlen=params['max_len'])
#print(valid_x_seq[:5])
# predict the labels on test dataset
classifier = create_cnn(input_length=params['max_len'])
classifier.load_weights(str(LMDATA/'bard.h5'))
# TPU can be enabled here if we need
# classifier = tpu_wrapper(classifier)
labels = np.array([list(to_categorical(label, 4).astype(int)) for label in test_df['label'].values])
score = classifier.evaluate(test_x_seq, labels, verbose=1)
print('Loss for final step: {}, accuracy: {}'.format(score[0], score[1]))
"""
# Just for testing
labels = [list(to_categorical(label, 4).astype(int)) for label in test_df['label'].values]
#print(labels[:5])
print(np.array(labels)[:5])
print(test_df['label'].values)
print(to_categorical(3, 4))
"""
prediction = classifier.predict(test_x_seq, verbose=1)
#print(prediction)
print('Predictions for final step: {}'.format(np.argmax(prediction, axis=1)[:10]))
print(test_x[:10])
!free
#print('Accuracy ', score[1])
"""
# First, run the seed forward to prime the state of the model.
#prediction_model.reset_states()
strategy = tf.contrib.tpu.TPUDistributionStrategy(
tf.contrib.cluster_resolver.TPUClusterResolver(TPU_WORKER))
prediction_model = tf.contrib.tpu.keras_to_tpu_model(
prediction_model, strategy=strategy)
predictions = prediction_model.predict(valid_x_seq)
print("predictions", predictions[:20])
predictions = tpu_model.predict(valid_x_seq)
predictions = [int(round(p[0])) for p in predictions]
#predictions = predictions.argmax(axis=-1)
print("predictions", predictions[:20])
print("valid_y", valid_y[:20])
return metrics.accuracy_score(predictions, valid_y)
"""
print(prediction)
print(np.argmax(prediction, axis=1))
```
### Kim Yoon’s CNN
```
# The following model is similar to
# Kim Yoon’s Convolutional Neural Networks for Sentence Classification
# (https://arxiv.org/abs/1408.5882)
#
# Model Hyperparameters
embedding_dim = 300
filter_sizes = (3, 5, 7)
num_filters = 100
dropout_prob = (0.5, 0.5)
hidden_dims = 50
def create_cnn_kimyoon(input_length=100):
# Add an Input Layer
input_layer = layers.Input((input_length, ))
# Add the word embedding Layer
embedding_layer = layers.Embedding(len(word_index) + 1, embedding_dim,
weights=[embedding_matrix],
trainable=False)(input_layer)
embedding_layer = layers.SpatialDropout1D(dropout_prob[0])(embedding_layer)
conv_array = []
for sz in filter_sizes:
conv = layers.Convolution1D(filters=num_filters,
kernel_size=sz,
padding="valid",
activation="relu",
strides=1)(embedding_layer)
conv = layers.MaxPooling1D(pool_size=2)(conv)
conv = layers.Flatten()(conv)
conv_array.append(conv)
layer = layers.Concatenate()(conv_array) if len(conv_array) > 1 else conv_array[0]
layer = layers.Dropout(dropout_prob[1])(layer)
layer = layers.Dense(hidden_dims, activation="relu")(layer)
output_layer = layers.Dense(4, activation="softmax")(layer)
# Compile the model
model = models.Model(inputs=input_layer, outputs=output_layer)
model.compile(optimizer=optimizers.Adam(),
loss='categorical_crossentropy',
metrics=['accuracy'])
return model
%%time
tf.keras.backend.clear_session()
classifier = create_cnn_kimyoon(input_length=params['max_len'])
classifier = tpu_wrapper(classifier)
classifier.fit_generator(
generator=training_generator,
validation_data=valid_generator,
#use_multiprocessing=True,
#workers=6,
epochs=20
)
classifier.save_weights(str(LMDATA/'cnn_kimyoon.h5'), overwrite=True)
# predict the labels on test dataset
classifier = create_cnn_kimyoon(input_length=params['max_len'])
classifier.load_weights(str(LMDATA/'cnn_kimyoon.h5'))
# TPU can be enabled here if we need
# classifier = tpu_wrapper(classifier)
labels = np.array([list(to_categorical(label, 4).astype(int)) for label in test_df['label'].values])
score = classifier.evaluate(test_x_seq, labels, verbose=1)
print('Loss for final step: {}, accuracy: {}'.format(score[0], score[1]))
```
### RNN-LSTM
```
def create_rnn_lstm(input_length=100):
# Add an Input Layer
input_layer = layers.Input((input_length, ))
# Add the word embedding Layer
embedding_layer = layers.Embedding(len(word_index) + 1, 300, weights=[embedding_matrix], trainable=False)(input_layer)
embedding_layer = layers.SpatialDropout1D(0.2)(embedding_layer)
# Add the LSTM Layer
lstm_layer = layers.LSTM(100)(embedding_layer)
#lstm_layer = layers.LSTM(100)(lstm_layer)
#lstm_layer = layers.LSTM(100, return_sequences=True)(lstm_layer)
#lstm_layer = layers.TimeDistributed(layers.Dense(100))(lstm_layer)
# Add the output Layers
output_layer1 = layers.Dense(100, activation="relu")(lstm_layer)
output_layer1 = layers.Dropout(0.2)(output_layer1)
output_layer2 = layers.Dense(4, activation="softmax")(output_layer1)
# Compile the model
model = models.Model(inputs=input_layer, outputs=output_layer2)
model.compile(optimizer=optimizers.Adam(), loss='categorical_crossentropy',
metrics=['accuracy'])
return model
%%time
# RNN-LSTM, Word Embeddings
# Speed comparison between CPU vs TPU
# First we test CPU with 2 epochs
tf.keras.backend.clear_session()
classifier = create_rnn_lstm(input_length=params['max_len'])
#classifier = tf.contrib.tpu.keras_to_tpu_model(classifier, strategy=strategy)
classifier.fit_generator(
generator=training_generator,
validation_data=valid_generator,
#use_multiprocessing=True,
#workers=6,
epochs=2
)
%%time
# RNN-LSTM, Word Embeddings
# We test now the TPU
# The result is:
# CPU: 65s/epoch
# TPU: 3.15s/epoch
tf.keras.backend.clear_session()
classifier = create_rnn_lstm(input_length=params['max_len'])
classifier = tpu_wrapper(classifier)
classifier.fit_generator(
generator=training_generator,
validation_data=valid_generator,
#use_multiprocessing=True,
#workers=6,
epochs=20
)
classifier.save_weights(str(LMDATA/'rnn_lstm.h5'), overwrite=True)
# predict the labels on test dataset
classifier = create_rnn_lstm(input_length=params['max_len'])
classifier.load_weights(str(LMDATA/'rnn_lstm.h5'))
# TPU can be enabled here if we need
# classifier = tpu_wrapper(classifier)
labels = np.array([list(to_categorical(label, 4).astype(int)) for label in test_df['label'].values])
score = classifier.evaluate(test_x_seq, labels, verbose=1)
print('Loss for final step: {}, accuracy: {}'.format(score[0], score[1]))
```
### RNN-GRU
```
def create_rnn_gru(input_length=100):
# Add an Input Layer
input_layer = layers.Input((input_length, ))
# Add the word embedding Layer
embedding_layer = layers.Embedding(len(word_index) + 1, 300, weights=[embedding_matrix], trainable=False)(input_layer)
embedding_layer = layers.SpatialDropout1D(0.3)(embedding_layer)
# Add the GRU Layer
lstm_layer = layers.GRU(100)(embedding_layer)
# Add the output Layers
output_layer1 = layers.Dense(50, activation="relu")(lstm_layer)
output_layer1 = layers.Dropout(0.25)(output_layer1)
output_layer2 = layers.Dense(4, activation="softmax")(output_layer1)
# Compile the model
model = models.Model(inputs=input_layer, outputs=output_layer2)
model.compile(optimizer=optimizers.Adam(), loss='categorical_crossentropy',
metrics=['accuracy'])
return model
%%time
# RNN-GRU, Word Embeddings
tf.keras.backend.clear_session()
classifier = create_rnn_gru(input_length=params['max_len'])
classifier = tpu_wrapper(classifier)
classifier.fit_generator(
generator=training_generator,
validation_data=valid_generator,
#use_multiprocessing=True,
#workers=6,
epochs=20
)
classifier.save_weights(str(LMDATA/'rnn_gru.h5'), overwrite=True)
# predict the labels on test dataset
classifier = create_rnn_gru(input_length=params['max_len'])
classifier.load_weights(str(LMDATA/'rnn_gru.h5'))
# TPU can be enabled here if we need
# classifier = tpu_wrapper(classifier)
labels = np.array([list(to_categorical(label, 4).astype(int)) for label in test_df['label'].values])
score = classifier.evaluate(test_x_seq, labels, verbose=1)
print('Loss for final step: {}, accuracy: {}'.format(score[0], score[1]))
```
### Biderectional RNN
```
# RNN-Bidirectional, Word Embeddings
# It doesn't work with TPU, but it works on CPU/GPU
def create_bidirectional_rnn(input_length=100):
# Add an Input Layer
input_layer = layers.Input((input_length, ))
# Add the word embedding Layer
embedding_layer = layers.Embedding(len(word_index) + 1, 300, weights=[embedding_matrix], trainable=False)(input_layer)
embedding_layer = layers.SpatialDropout1D(0.3)(embedding_layer)
# Add the LSTM Layer
lstm_layer = layers.Bidirectional(layers.GRU(100))(embedding_layer)
# Add the output Layers
output_layer1 = layers.Dense(50, activation="relu")(lstm_layer)
output_layer1 = layers.Dropout(0.25)(output_layer1)
output_layer2 = layers.Dense(4, activation="softmax")(output_layer1)
# Compile the model
model = models.Model(inputs=input_layer, outputs=output_layer2)
model.compile(optimizer=optimizers.Adam(), loss='categorical_crossentropy',
metrics=['accuracy'])
return model
%%time
# RNN-Bidirectional, Word Embeddings
# It doesn't work with TPU, but it works on CPU/GPU
tf.keras.backend.clear_session()
classifier = create_bidirectional_rnn(input_length=params['max_len'])
#classifier = tpu_wrapper(classifier)
classifier.fit_generator(
generator=training_generator,
validation_data=valid_generator,
#use_multiprocessing=True,
#workers=6,
epochs=20
)
classifier.save_weights(str(LMDATA/'rnn_bidirectional.h5'), overwrite=True)
# predict the labels on test dataset
classifier = create_bidirectional_rnn(input_length=params['max_len'])
classifier.load_weights(str(LMDATA/'rnn_bidirectional.h5'))
# TPU can be enabled here if we need
# classifier = tpu_wrapper(classifier)
labels = np.array([list(to_categorical(label, 4).astype(int)) for label in test_df['label'].values])
score = classifier.evaluate(test_x_seq, labels, verbose=1)
print('Loss for final step: {}, accuracy: {}'.format(score[0], score[1]))
```
### RCNN
```
def create_rcnn(input_length=100):
# Add an Input Layer
input_layer = layers.Input((input_length, ))
# Add the word embedding Layer
embedding_layer = layers.Embedding(len(word_index) + 1, 300, weights=[embedding_matrix], trainable=False)(input_layer)
embedding_layer = layers.SpatialDropout1D(0.3)(embedding_layer)
# Add the recurrent layer
rnn_layer = layers.Bidirectional(layers.GRU(50, return_sequences=True))(embedding_layer)
# Add the convolutional Layer
conv_layer = layers.Convolution1D(100, 3, activation="relu")(embedding_layer)
# Add the pooling Layer
pooling_layer = layers.GlobalMaxPool1D()(conv_layer)
# Add the output Layers
output_layer1 = layers.Dense(50, activation="relu")(pooling_layer)
output_layer1 = layers.Dropout(0.25)(output_layer1)
output_layer2 = layers.Dense(4, activation="sigmoid")(output_layer1)
# Compile the model
model = models.Model(inputs=input_layer, outputs=output_layer2)
model.compile(optimizer=optimizers.Adam(), loss='categorical_crossentropy',
metrics=['accuracy'])
return model
%%time
# RCNN, Word Embeddings
tf.keras.backend.clear_session()
classifier = create_rcnn(input_length=params['max_len'])
classifier = tpu_wrapper(classifier)
classifier.fit_generator(
generator=training_generator,
validation_data=valid_generator,
#use_multiprocessing=True,
#workers=6,
epochs=20
)
classifier.save_weights(str(LMDATA/'rcnn.h5'), overwrite=True)
# predict the labels on test dataset
classifier = create_rcnn(input_length=params['max_len'])
classifier.load_weights(str(LMDATA/'rcnn.h5'))
# TPU can be enabled here if we need
# classifier = tpu_wrapper(classifier)
labels = np.array([list(to_categorical(label, 4).astype(int)) for label in test_df['label'].values])
score = classifier.evaluate(test_x_seq, labels, verbose=1)
print('Loss for final step: {}, accuracy: {}'.format(score[0], score[1]))
```
| github_jupyter |
# Simulating noisy quantum circuits with Paddle Quantum
<em> Copyright (c) 2021 Institute for Quantum Computing, Baidu Inc. All Rights Reserved. </em>
## Introduction to quantum noises
In ideal models, we usually assume that quantum circuits are operating on a **closed physical system**. However, real quantum devices suffer from **incoherent noises** introduced by unwanted interactions between the system and the environment. This type of noise can significantly change the performance of quantum computation tasks and hence can hardly be ignored for near-term quantum devices. Consequently, designing robust quantum algorithms under the presence of noise is crucial for utilizing quantum computation in the real world. With the noise module of Paddle Quantum, we can now not only design and simulate quantum algorithms but also examine various noises' influence and further develop error mitigation schemes.
## Building noise models in Paddle Quantum
### Noise model and quantum channel
The evolution of a closed quantum system is always unitary. Mathematically, we can describe such a process as implementing a parameterized quantum circuit $U(\vec{\theta})$,
$$
\rho \longrightarrow U(\vec{\theta}) \rho U^\dagger(\vec{\theta}),
\tag{1}
$$
where $\rho$ is the initial quantum state, $\vec{\theta}$ is a vector containing all the parameters. The most intuitive type of noise one can think of is the error that appears in these parameters,
$$
\rho \longrightarrow U(\vec{\theta}+\vec{\epsilon}) \rho U^\dagger(\vec{\theta}+\vec{\epsilon}),
\tag{2}
$$
$\vec{\epsilon}$ can be a white noise sampled from Gaussian distributions. This kind of noise is a specific example of **coherent noises**. Coherent noise usually occurs due to device calibration errors or quantum control errors. We want to emphasize that one also uses unitary transformation $U(\vec{\epsilon})$ to describe coherent noises. In certain cases, coherent noises can be more damaging than incoherent noises [1].
Most of the time, the real problem lies on the evolution of an **open quantum system** that is non-unitary. Under this circumstance, we need a more general description beyond the unitary transformation to characterize incoherent noises, the language of **quantum channels**. To keep the discussion precise, we use *operator-sum representation* [2] to introduce a quantum channel as
$$
\mathcal{E}(\rho) = \sum_{k=0}^{m-1} E_k \rho E_k^{\dagger},
\tag{3}
$$
where $\{E_k\}$ are *Kraus* operators, and they satisfy the completeness condition $\sum_k E_k^\dagger E_k = I$. Mathematically, a quantum channel is completely positive and trace-preserving [2].
Under this representation, we can explicitly observe the results of implementing a quantum channel: Suppose we start with a pure state $\rho = |\psi\rangle\langle \psi|$, then we send it through a noisy quantum channel (e.g., $m = 2$ ). Eventually, we will get a mixed state $\mathcal{E}(\rho) = E_0 \rho E_0^\dagger + E_1 \rho E_1^\dagger$. Let's take the bit flip noise as an example:
$$
\mathcal{E}_{BF}(\rho) = (1 - p) I \rho I+ p X \rho X,
\tag{4}
$$
where $X,I$ are Pauli operators. The corresponding *Kraus* operators are:
$$
E_0 = \sqrt{1-p}
\begin{bmatrix}
1&0 \\
0&1
\end{bmatrix},
E_1 = \sqrt{p}
\begin{bmatrix}
0& 1 \\
1 &0
\end{bmatrix}.
\tag{5}
$$
The physical meaning of this quantum channel is there exist a probability $p$ that the state $|0\rangle$ will flip into $|1\rangle$, and vice versa. In Paddle Quantum, we can use this quantum channel by `UAnsatz.bit_flip(p, which_qubit)`, where `p` is the noise level.
**Note:** For a quantum channel, the Kraus operator representation is not necessarily unique [3].
### Implementation with Paddle Quantum
In this section, we will learn how to build a noise model in Paddle Quantum. First, we initialize a qubit to $|0\rangle$.
```
import paddle
from paddle_quantum.circuit import UAnsatz
# Define the number of qubits, here we use one single qubit
num_qubits = 1
# Initialize the quantum circuit
cir = UAnsatz(num_qubits)
# Initialize the qubit to |0><0|
init_state = cir.run_density_matrix()
# Mesure in the computational basis
cir.measure(plot=True)
```
Then, we add a bit flip channel with $p=0.1$, and measure the qubit after this channel.
**Note:** Noisy module in Paddle Quantum only supports density matrix operation mode.
```
# Noise level
p = 0.1
# Add the bit flip noisy channel
cir.bit_flip(p, 0)
# Execute the circuit
# Note: Noisy module in Paddle Quantum only supports density matrix operation mode
fin_state = cir.run_density_matrix()
# Measure in the computational basis
cir.measure(plot=True)
print('Quantum state after the bit flip quantum channel:\n', fin_state.numpy())
```
As we can see, the quantum state has been transformed to a mixed state $0.9 | 0 \rangle \langle 0 | + 0.1 | 1 \rangle \langle 1 |$ (with probability $p=0.1$ ) after the bit flip channel.
### Common quantum channels
Paddle Quantum supports many other common noisy channels.
- **Phase Flip Channel**
Similar to the bit-flip channel, the phase flip channel flips the phase of a qubit with probability $p$,
$$
\mathcal{E}_{PF}(\rho) = (1 - p) \rho + p Z \rho Z.
\tag{6}
$$
- **Bit-Phase Flip Channel**
$$
\mathcal{E}_{BPF}(\rho) = (1-p) \rho + p Y \rho Y.
\tag{7}
$$
- **Depolarizing Channel**
The quantum state will be in the maximally mixed state $I/2$ with probability $p$ or in the original state with probability $1-p$ after the single qubit depolarizing channel. The depolarizing channel can also be understood as applying Pauli noises symmetrically,
$$
\mathcal{E}_{D}(\rho) = (1 - p) \rho + \frac{p}{3}
\left( X \rho X+ Y \rho Y + Z \rho Z \right).
\tag{8}
$$
- **Pauli Channel**
The Pauli channel applies Pauli noises asymmetrically,
$$
\mathcal{E}_{Pauli}(\rho) = (1 - p_x - p_y - p_z) \rho + p_x X \rho X + p_y Y \rho Y + p_z Z \rho Z.
\tag{9}
$$
- **Amplitude Damping Channel**
The amplitude damping channel can be used to model the process of **energy dissipation**,
$$
\mathcal{E}_{AD}(\rho) = E_0 \rho E_0^\dagger + E_1 \rho E_1^\dagger,
\tag{10}
$$
where $\gamma$ is the damping factor,
$$
E_0 =
\begin{bmatrix}
1 & 0 \\ 0 & \sqrt{1 - \gamma}
\end{bmatrix},
E_1 =
\begin{bmatrix}
0 & \sqrt{\gamma} \\ 0 & 0
\end{bmatrix}.
\tag{11}
$$
- **Phase Damping Channel**
The phase damping channel describes the loss of **quantum information** without loss of energy,
$$
\mathcal{E}_{PD}(\rho) = E_0 \rho E_0^\dagger + E_1 \rho E_1^\dagger,
\tag{12}
$$
where $\gamma$ is the damping factor,
$$
E_0 =
\begin{bmatrix}
1 & 0 \\ 0 & \sqrt{1 - \gamma}
\end{bmatrix},
E_1 =
\begin{bmatrix}
0 & 0 \\ 0 & \sqrt{\gamma}
\end{bmatrix}.
\tag{13}
$$
- **Generalized Amplitude Damping Channel**
The generalized amplitude damping channel describes energy exchange between the system and the environment at **finite temperatures**. It is a common noise in superconducting quantum computations [4]. Interested readers can find more information here [API document](https://qml.baidu.com/api/paddle_quantum.circuit.uansatz.html).
**Note:** In Paddle Quantum, we can use these noisy channels through `UAnsatz.phase_flip()`, `UAnsatz.bit_phase_flip()`, `UAnsatz.depolarizing()`, `UAnsatz.pauli_channel()`, `UAnsatz.amplitude_damping()`, `UAnsatz.phase_damping()`, and `UAnsatz.generalized_amplitude_damping()`.
**Note:** One usually choose the amplitude damping channel and the phase damping channel to model noises since they describe the physical process in real quantum systems (modeling $T_1$ and $T_2$ process).
### Customized Channel
One can also use `UAnsatz.customized_channel()` in Paddle Quantum to add customized noisy channels. This is accomplished through user-defined Kraus operators. Here, we provide an example to reproduce the bit flip channel using customized_channel function:
```
import paddle
import numpy as np
from paddle_quantum.circuit import UAnsatz
# Noise level
p = 0.1
# We use customized Kraus operator to represent the bit flip channel
# Note that the data type of a Kraus operator is complex_128
a_0 = paddle.to_tensor(np.sqrt(1 - p) * np.array([[1, 0], [0, 1]], dtype='complex128'))
a_1 = paddle.to_tensor(np.sqrt(p) * np.array([[0, 1], [1, 0]], dtype='complex128'))
Kraus_ops = [a_0, a_1]
# Initialize the circuit
num_qubits = 1
cir = UAnsatz(num_qubits)
# Add customized channel, input is a list of Kraus operators
cir.customized_channel(Kraus_ops, 0)
# Execute the circuit
fin_state = cir.run_density_matrix()
# Compare the results
cir_1 = UAnsatz(num_qubits)
cir_1.bit_flip(p, 0)
fin_state_1 = cir_1.run_density_matrix()
print('quantum state after the customized channel:\n', fin_state.numpy())
print('\n quantum state after the bit flip channel:\n', fin_state_1.numpy())
print('\n are the two the same?', bool((fin_state - fin_state_1).abs().sum() < 1e-8))
```
## Discussion: Simulating noisy entanglement resources with Paddle Quantum
Many important quantum technologies require pre-shared entanglement resources, including quantum teleportation, state transformation, and distributed quantum computing. For instance, we want the allocated entanglement resources are in **maximally entangled states** under ideal circumstances. But in reality, noise always exists due to interactions between the system and the environment during preparation stage, transmission, and preservation. Here, we use the depolarized channel to simulate how a white noise could affect Bell states:
```
import paddle
from paddle import matmul, trace
from paddle_quantum.circuit import UAnsatz
from paddle_quantum.state import bell_state
# Noise level
p_trans = 0.1
p_store = 0.01
# Initialize the circuit
num_qubits = 2
cir = UAnsatz(num_qubits)
# The initial state is Bell state
init_state = paddle.to_tensor(bell_state(2))
# Apply the depolarized channel to each qubit, modeling the noise introduced by transmission
cir.depolarizing(p_trans, 0)
cir.depolarizing(p_trans, 1)
# Execute the circuit
status_mid = cir.run_density_matrix(init_state)
# Apply the amplitude damping channel to each qubit, modeling the noise introduced by storage
cir.amplitude_damping(p_store, 0)
cir.amplitude_damping(p_store, 1)
# Execute the circuit
status_fin = cir.run_density_matrix(status_mid)
fidelity_mid = paddle.real(trace(matmul(init_state, status_mid)))
fidelity_fin = paddle.real(trace(matmul(init_state, status_fin)))
print("Fidelity between the initial state and the Bell state", 1)
print("after transmission (depolarized channel), the fidelity between the entangled state and Bell state {:.5f}".format(fidelity_mid.numpy()[0]))
print("after preservation (amplitude damping channel), the fidelity between the entangled state and Bell state {:.5f}".format(fidelity_fin.numpy()[0]))
```
**Note:** Interested readers can check tutorials on the LOCCNet module of Paddle Quantum, where we discuss the concept of [entanglement distillation](../locc/EntanglementDistillation_LOCCNET_EN.ipynb).
## Application: Simulating noisy VQE with Paddle Quantum
Variational Quantum Eigensolver (VQE) [5] is designed to find the ground state energy of a given molecular Hamiltonian using variational quantum circuits. Interested readers can find more details from the previous tutorial [VQE](../quantum_simulation/VQE_EN.ipynb).
For illustration purposes, we use VQE to find the ground state energy for the following Hamiltonian:
$$
H = 0.4 \, Z \otimes I + 0.4 \, I \otimes Z + 0.2 \, X \otimes X.
\tag{14}
$$
Then, we add the amplitude damping channel and compare the performance of the noisy circuit and the noiseless circuit on this task:
```
import numpy as np
import paddle
from paddle_quantum.circuit import UAnsatz
from paddle_quantum.utils import pauli_str_to_matrix
# Construct Hamiltonian using Pauli string
H_info = [[0.4, 'z0'], [0.4, 'z1'], [0.2, 'x0,x1']]
# Convert the Pauli string to a matrix
H_matrix = pauli_str_to_matrix(H_info, num_qubits)
# Hyperparameters
num_qubits = 2
theta_size = 4
ITR = 100
LR = 0.4
SEED = 999
p = 0.1
class vqe_noisy(paddle.nn.Layer):
def __init__(self, shape, dtype='float64'):
super(vqe_noisy, self).__init__()
# Initialize a learnable parameter list with length theta_size, initial values are sampled from uniform distribution between [0, 2*pi]
self.theta = self.create_parameter(shape=shape,
default_initializer=paddle.nn.initializer.Uniform(low=0., high=2*np.pi),
dtype=dtype, is_bias=False)
# Define loss function and forward function
def forward(self):
# Initialize circuit
cir = UAnsatz(num_qubits)
# Add parameterized gates
cir.ry(self.theta[0], 0)
cir.ry(self.theta[1], 1)
cir.cnot([0, 1])
cir.ry(self.theta[2], 0)
cir.ry(self.theta[3], 1)
# Add amplitude damping channel
cir.amplitude_damping(p, 0)
cir.amplitude_damping(p, 1)
# Execute the circuit
cir.run_density_matrix()
# Expectation value of Hamiltonian
loss = cir.expecval(H_info)
return loss
# Construct a noiseless circuit
class vqe_noise_free(paddle.nn.Layer):
def __init__(self, shape, dtype='float64'):
super(vqe_noise_free, self).__init__()
self.theta = self.create_parameter(shape=shape,
default_initializer=paddle.nn.initializer.Uniform(low=0., high=2*np.pi),
dtype=dtype, is_bias=False)
def forward(self):
cir = UAnsatz(num_qubits)
cir.ry(self.theta[0], 0)
cir.ry(self.theta[1], 1)
cir.cnot([0, 1])
cir.ry(self.theta[2], 0)
cir.ry(self.theta[3], 1)
cir.run_density_matrix()
loss = cir.expecval(H_info)
return loss
# Train noisy VQE circuit
print('========== Training Noisy VQE ==========')
loss_list = []
parameter_list = []
# Define the dimension of parameters
vqe = vqe_noisy([theta_size])
# Generally, we use Adam optimizer to get a better convergence, you can change to SVG or RMS prop.
opt = paddle.optimizer.Adam(learning_rate = LR, parameters = vqe.parameters())
# Optimization iteration
for itr in range(ITR):
# Forward, to calculate loss function
loss = vqe()
# Backpropagate to minimize the loss function
loss.backward()
opt.minimize(loss)
opt.clear_grad()
# Record the learning curve
loss_list.append(loss.numpy()[0])
parameter_list.append(vqe.parameters()[0].numpy())
if itr % 10 == 0:
print('iter:', itr, ' loss: %.4f' % loss.numpy())
# Train the noiseless VQE in the same way
print('========== Training Noise Free VQE ==========')
loss_list_no_noise = []
parameter_list_no_noise = []
vqe_no_noise = vqe_noise_free([theta_size])
opt_no_noise = paddle.optimizer.Adam(learning_rate = LR, parameters = vqe_no_noise.parameters())
for itr in range(ITR):
loss = vqe_no_noise()
loss.backward()
opt_no_noise.minimize(loss)
opt_no_noise.clear_grad()
loss_list_no_noise.append(loss.numpy()[0])
parameter_list_no_noise.append(vqe_no_noise.parameters()[0].numpy())
if itr % 10 == 0:
print('iter:', itr, ' loss: %.4f' % loss.numpy())
print('\nGround state energy from noisy circuit: ', loss_list[-1], "Ha")
print('Ground state energy from noiseless circuit: ', loss_list_no_noise[-1], "Ha")
print('Actual ground state energy: ', np.linalg.eigh(H_matrix)[0][0], "Ha")
```
As we can see, noisy VQE behaves much worse than the noiseless version as expected and couldn't satisfy chemical accuracy $\varepsilon = 0.0016$ Ha.
## Conclusion
Noise is an unavoidable feature of quantum devices in the NISQ era. Therefore, designing robust quantum algorithms under the presence of noise and further developing error mitigation schemes are two important research directions. With the noise module in Paddle Quantum, we hope to provide a platform simulating real physical systems and help developing near-term quantum computation applications. Standing together with the research community, the noise module will help us explore what we can achieve with noisy devices, design more robust quantum algorithms, and eventually leads to trustworthy quantum solutions in areas including AI and quantum chemistry.
---
## References
[1] Iverson, J. K., & Preskill, J. Coherence in logical quantum channels. [New Journal of Physics, 22(7), 073066 (2020).](https://iopscience.iop.org/article/10.1088/1367-2630/ab8e5c)
[2] Nielsen, M. A. & Chuang, I. L. Quantum computation and quantum information. Cambridge university press (2010).
[3] Preskill, J. Quantum Information Lecture Notes. Chapter 3 (2018).
[4] Chirolli, L., & Burkard, G. Decoherence in solid-state qubits. [Advances in Physics, 57(3), 225-285 (2008).](https://www.tandfonline.com/doi/abs/10.1080/00018730802218067)
[5] Peruzzo, A. et al. A variational eigenvalue solver on a photonic quantum processor. [Nat. Commun. 5, 4213 (2014).](https://www.nature.com/articles/ncomms5213)
| github_jupyter |
```
from utils import *
import tensorflow as tf
from sklearn.cross_validation import train_test_split
import time
trainset = sklearn.datasets.load_files(container_path = 'data', encoding = 'UTF-8')
trainset.data, trainset.target = separate_dataset(trainset,1.0)
print (trainset.target_names)
print (len(trainset.data))
print (len(trainset.target))
ONEHOT = np.zeros((len(trainset.data),len(trainset.target_names)))
ONEHOT[np.arange(len(trainset.data)),trainset.target] = 1.0
train_X, test_X, train_Y, test_Y, train_onehot, test_onehot = train_test_split(trainset.data,
trainset.target,
ONEHOT, test_size = 0.2)
concat = ' '.join(trainset.data).split()
vocabulary_size = len(list(set(concat)))
data, count, dictionary, rev_dictionary = build_dataset(concat, vocabulary_size)
print('vocab from size: %d'%(vocabulary_size))
print('Most common words', count[4:10])
print('Sample data', data[:10], [rev_dictionary[i] for i in data[:10]])
GO = dictionary['GO']
PAD = dictionary['PAD']
EOS = dictionary['EOS']
UNK = dictionary['UNK']
def temporal_padding(x,padding=(1,1)):
return tf.pad(x, [[0, 0], [padding[0], padding[1]], [0, 0]])
def attention_block(x):
k_size = x.get_shape()[-1].value
v_size = x.get_shape()[-1].value
key = tf.layers.dense(x, units=k_size, activation=None, use_bias=False,
kernel_initializer=tf.random_normal_initializer(0, 0.01))
query = tf.layers.dense(x, units=v_size, activation=None, use_bias=False,
kernel_initializer=tf.random_normal_initializer(0, 0.01))
logits = tf.matmul(key, key, transpose_b=True)
logits = logits / np.sqrt(k_size)
weights = tf.nn.softmax(logits, name="attention_weights")
return tf.matmul(weights, query)
def convolution1d(x, num_filters, dilation_rate, k,
filter_size=3, stride=[1], pad='VALID'):
with tf.variable_scope('conv1d_%d'%(k)):
num_filters = num_filters * 2
V = tf.get_variable('V', [filter_size, int(x.get_shape()[-1]), num_filters],
tf.float32, initializer=None,trainable=True)
g = tf.get_variable('g', shape=[num_filters], dtype=tf.float32,
initializer=tf.constant_initializer(1.), trainable=True)
b = tf.get_variable('b', shape=[num_filters], dtype=tf.float32,
initializer=None, trainable=True)
W = tf.reshape(g, [1, 1, num_filters]) * tf.nn.l2_normalize(V, [0, 1])
left_pad = dilation_rate * (filter_size - 1)
x = temporal_padding(x, (left_pad, 0))
x = tf.nn.bias_add(tf.nn.convolution(x, W, pad, stride, [dilation_rate]), b)
split0, split1 = tf.split(x, num_or_size_splits=2, axis=2)
split1 = tf.sigmoid(split1)
return tf.multiply(split0, split1)
def temporalblock(input_layer, out_channels, filter_size, stride, dilation_rate,
dropout,k,highway=False):
keep_prob = 1.0 - dropout
in_channels = input_layer.get_shape()[-1]
count = 0
with tf.variable_scope('temporal_block_%d'%(k)):
conv1 = convolution1d(input_layer, out_channels, dilation_rate, count,
filter_size, [stride])
noise_shape = (tf.shape(conv1)[0], tf.constant(1), tf.shape(conv1)[2])
dropout1 = tf.nn.dropout(conv1, keep_prob, noise_shape)
dropout1 = attention_block(dropout1)
count += 1
conv2 = convolution1d(input_layer, out_channels, dilation_rate, count,
filter_size, [stride])
dropout2 = tf.nn.dropout(conv2, keep_prob, noise_shape)
dropout2 = attention_block(dropout2)
residual = None
if highway:
W_h = tf.get_variable('W_h', [1, int(input_layer.get_shape()[-1]), out_channels],
tf.float32, tf.random_normal_initializer(0, 0.01), trainable=True)
b_h = tf.get_variable('b_h', shape=[out_channels], dtype=tf.float32,
initializer=None, trainable=True)
H = tf.nn.bias_add(tf.nn.convolution(input_layer, W_h, 'SAME'), b_h)
W_t = tf.get_variable('W_t', [1, int(input_layer.get_shape()[-1]), out_channels],
tf.float32, tf.random_normal_initializer(0, 0.01), trainable=True)
b_t = tf.get_variable('b_t', shape=[out_channels], dtype=tf.float32,
initializer=None, trainable=True)
T = tf.nn.bias_add(tf.nn.convolution(input_layer, W_t, 'SAME'), b_t)
T = tf.nn.sigmoid(T)
residual = H*T + input_layer * (1.0 - T)
elif in_channels != out_channels:
W_h = tf.get_variable('W_h', [1, int(input_layer.get_shape()[-1]), out_channels],
tf.float32, tf.random_normal_initializer(0, 0.01), trainable=True)
b_h = tf.get_variable('b_h', shape=[out_channels], dtype=tf.float32,
initializer=None, trainable=True)
residual = tf.nn.bias_add(tf.nn.convolution(input_layer, W_h, 'SAME'), b_h)
else:
print("no residual convolution")
res = input_layer if residual is None else residual
return tf.nn.relu(dropout2 + res)
def temporal_convd(input_layer, num_channels, sequence_length,
kernel_size=2, dropout=0):
for i in range(len(num_channels)):
dilation_size = 2 ** i
out_channels = num_channels[i]
input_layer = temporalblock(input_layer, out_channels, kernel_size, 1, dilation_size,dropout,i)
print(input_layer.shape)
return input_layer
class Model:
def __init__(self,embedded_size,dict_size, dimension_output, learning_rate,
levels=5,size_layer=256,kernel_size=7,maxlen=50):
self.X = tf.placeholder(tf.int32, [None, maxlen])
self.Y = tf.placeholder(tf.float32, [None, dimension_output])
encoder_embeddings = tf.Variable(tf.random_uniform([dict_size, embedded_size], -1, 1))
encoder_embedded = tf.nn.embedding_lookup(encoder_embeddings, self.X)
#channel_sizes = [int(size_layer * ((i+1) / levels)) for i in reversed(range(levels))]
channel_sizes = [size_layer] * levels
tcn = temporal_convd(input_layer=encoder_embedded, num_channels=channel_sizes,
sequence_length=maxlen, kernel_size=kernel_size)
self.logits = tf.contrib.layers.fully_connected(tcn[:, -1, :], dimension_output,
activation_fn=None)
self.cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits = self.logits, labels = self.Y))
self.optimizer = tf.train.AdamOptimizer(learning_rate = learning_rate).minimize(self.cost)
correct_pred = tf.equal(tf.argmax(self.logits, 1), tf.argmax(self.Y, 1))
self.accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
embedded_size = 128
dimension_output = len(trainset.target_names)
batch_size = 128
learning_rate = 1e-3
maxlen = 50
tf.reset_default_graph()
sess = tf.InteractiveSession()
model = Model(embedded_size,len(dictionary),dimension_output,learning_rate)
sess.run(tf.global_variables_initializer())
EARLY_STOPPING, CURRENT_CHECKPOINT, CURRENT_ACC, EPOCH = 5, 0, 0, 0
while True:
lasttime = time.time()
if CURRENT_CHECKPOINT == EARLY_STOPPING:
print('break epoch:%d\n'%(EPOCH))
break
train_acc, train_loss, test_acc, test_loss = 0, 0, 0, 0
for i in range(0, (len(train_X) // batch_size) * batch_size, batch_size):
batch_x = str_idx(train_X[i:i+batch_size],dictionary,maxlen)
acc, loss, _ = sess.run([model.accuracy, model.cost, model.optimizer],
feed_dict = {model.X : batch_x, model.Y : train_onehot[i:i+batch_size]})
train_loss += loss
train_acc += acc
for i in range(0, (len(test_X) // batch_size) * batch_size, batch_size):
batch_x = str_idx(test_X[i:i+batch_size],dictionary,maxlen)
acc, loss = sess.run([model.accuracy, model.cost],
feed_dict = {model.X : batch_x, model.Y : test_onehot[i:i+batch_size]})
test_loss += loss
test_acc += acc
train_loss /= (len(train_X) // batch_size)
train_acc /= (len(train_X) // batch_size)
test_loss /= (len(test_X) // batch_size)
test_acc /= (len(test_X) // batch_size)
if test_acc > CURRENT_ACC:
print('epoch: %d, pass acc: %f, current acc: %f'%(EPOCH,CURRENT_ACC, test_acc))
CURRENT_ACC = test_acc
CURRENT_CHECKPOINT = 0
else:
CURRENT_CHECKPOINT += 1
print('time taken:', time.time()-lasttime)
print('epoch: %d, training loss: %f, training acc: %f, valid loss: %f, valid acc: %f\n'%(EPOCH,train_loss,
train_acc,test_loss,
test_acc))
EPOCH += 1
logits = sess.run(model.logits, feed_dict={model.X:str_idx(test_X,dictionary,maxlen)})
print(metrics.classification_report(test_Y, np.argmax(logits,1), target_names = trainset.target_names))
```
| github_jupyter |
```
from skimage import io
import matplotlib.pyplot as plt
%matplotlib inline
klads = [io.imread('Klad00.jpg'), io.imread('Klad01.jpg'), io.imread('Klad02.jpg')]
def plot_klads(klads):
for i in klads:
io.imshow(i)
io.show()
plot_klads(klads)
from skimage.color import rgb2gray
thresh = 0.04
def get_binary(img):
img_gray = rgb2gray(img)
img_binary = img_gray > thresh
return img_binary
print u'Порог:', thresh
klads_binary = map(get_binary, klads)
plot_klads(klads_binary)
from skimage import measure
def get_connected_components(img):
components = measure.label(img)
return components
klad_components = map(get_connected_components, klads_binary)
plot_klads(klad_components)
from skimage.measure import regionprops
thresh_eccentricity = 0.8
def get_arrows(components):
arrows = components.copy()
labels_props = regionprops(components)
arrows_labels = [label_props.label
for label_props in labels_props if label_props.eccentricity > thresh_eccentricity]
for i in range(arrows.shape[0]):
for j in range(arrows.shape[1]):
arrows[i][j] = arrows[i][j] in arrows_labels
return arrows * components
klad_arrows = map(get_arrows, klad_components)
plot_klads(klad_arrows)
def get_init_arrow((arrows, src)):
red_mask = (src[:,:,2] < 90) * (src[:,:,0] > 100)
init_arrow = arrows * red_mask
return init_arrow
klad_init_arrow = map(get_init_arrow, zip(klad_arrows, klads))
plot_klads(klad_init_arrow)
def get_next_label(label, arrows, src):
dots = (src[:,:,1] > 50) * (src[:,:,2] < 50) * 100 * (arrows > 0)
arrows_props = regionprops(arrows, dots + (arrows > 0))
arrow_props = None
for p in arrows_props:
if p.label == label:
arrow_props = p
cy, cx = arrow_props.centroid
weighed_cy, weighted_cx = arrow_props.weighted_centroid
range_x = range(int(weighted_cx), src.shape[1]) if cx < weighted_cx else range(int(weighted_cx), -1, -1)
def get_y(x):
return -(weighed_cy - cy) * (weighted_cx - x)/(weighted_cx - cx) + weighed_cy
next_items = dict()
next_label = 0.
for x in range_x:
y = get_y(x)
if y < arrows.shape[0] and arrows[y][x] != 0 and arrows[y][x] != label:
next_label = arrows[y][x]
next_items[label] = next_label
break
return next_label
klad0_arrows = klad_arrows[0]
label = 2
io.imshow(klad0_arrows == label)
io.show()
next_label = get_next_label(label, klad0_arrows, klads[0])
io.imshow(klad0_arrows == next_label)
io.show()
import numpy as np
def get_the_treasure(src):
binary = get_binary(src)
components = get_connected_components(binary)
arrows = get_arrows(components)
arrows_labels = [label_props.label
for label_props in regionprops(components) if label_props.eccentricity > thresh_eccentricity]
init_arrow = get_init_arrow((arrows, src))
cur_label = (set(init_arrow.ravel()) - {0}).pop()
while cur_label in arrows_labels:
cur_label = get_next_label(cur_label, components, src)
return components == cur_label
treasures = map(get_the_treasure, klads)
plot_klads(treasures)
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import math
import os
def printCategory(category_dict):
for category in category_dict:
print(category)
if category_dict[category] is not None:
sub_dic = category_dict[category]
printCategory(sub_dic)
def createLevelColumns(df, levels, new_columns):
for i in range(len(df)):
df_level1, df_level2, df_level3 = "NULL","NULL","NULL"
df_level1, nextLevels = findNotNullLevel(df, i, levels)
if nextLevels is not None:
df_level2, nextLevels = findNotNullLevel(df, i, nextLevels)
if nextLevels is not None:
df_level3, nextLevel = findNotNullLevel(df, i, nextLevels)
most_specific_category = "NULL"
if df_level3 != "NULL":
df_level3 = df_level3 + "__" + df_level2 + "__" + df_level1
if df_level2 != "NULL":
df_level2 = df_level2 + "__" + df_level1
if df_level3 != "NULL":
most_specific_category = df_level3
elif df_level2 != "NULL":
most_specific_category = df_level2
elif df_level1 != "NULL":
most_specific_category = df_level1
df.iloc[i, df.columns.get_loc('level1')] = df_level1
df.iloc[i, df.columns.get_loc('level2')] = df_level2
df.iloc[i, df.columns.get_loc('level3')] = df_level3
df.iloc[i, df.columns.get_loc('mostSpecificCategory')] = most_specific_category
def findNotNullLevel(df, i, levels):
for level in levels:
if df.iloc[i][level] == "NULL":
continue
else:
return df.iloc[i][level], levels[level]
return "NULL", None
def unionTwoLists(list1, list2):
for category in list1:
if category not in list2:
list2.append(category)
return list2
def checkNULL(checked_list):
for item in checked_list:
if item == "NULL":
print("Contains NULL")
return
print("Not contains NULL")
```
#### 导入数据 yearly and range
#### process为designed schema
1. GazetteerEconomy table
1.1 Index - Id
1.2 Gazetteer Code - gazetteerId
1.3 Category - categoryId
1.4 yearly - startYear = endYear
1.5 2010 data - data
1.6 Unit - unitId
2. EconomyCategory table
2.1 Category Division1 Subdivision - Agri. Subdivision - Misc. Subdivision - Service Division2 Subdivision - Househould
3. UnitCategory
4. Gazetteer
```
path = os.path.abspath(os.getcwd())
df = pd.read_csv(path + "/Data2/Education - Yearly.csv")
df2 = pd.read_csv(path + "/Data2/Education - Range.csv")
df = df.dropna(axis = 0, how = 'all')
df2 = df2.dropna(axis = 0, how = 'all')
print(df.columns)
print(df['Category'].astype('category').unique())
print(df['Division1'].astype('category').unique())
print(df['Subdivision'].astype('category').unique())
# create new columns at df and df2
new_columns = ['level1', 'level2', 'level3', 'mostSpecificCategory', 'categoryId']
for column in new_columns:
df[column] = None
df = df.where(df.notnull(), "NULL")
for column in new_columns:
df2[column] = None
df2 = df2.where(df2.notnull(), "NULL")
# create dictionary records category levles
heading = {'Category': {'Division1':{'Subdivision': None}}}
printCategory(heading)
createLevelColumns(df, heading, new_columns)
createLevelColumns(df2, heading, new_columns)
level1 = unionTwoLists([cat for cat in df['level1'].astype('category').unique()],
[cat for cat in df2['level1'].astype('category').unique()] )
level2 = unionTwoLists([cat for cat in df['level2'].astype('category').unique()],
[cat for cat in df2['level2'].astype('category').unique()] )
level3 = unionTwoLists([cat for cat in df['level3'].astype('category').unique()],
[cat for cat in df2['level3'].astype('category').unique()] )
most_specific_category = unionTwoLists([cat for cat in df['mostSpecificCategory'].astype('category').unique()],
[cat for cat in df2['mostSpecificCategory'].astype('category').unique()])
total_categories = unionTwoLists(level1, level2)
total_categories = unionTwoLists(level3, total_categories)
# get total categories
total_categories.sort()
checkNULL(total_categories)
print("total number of categories are " + str(len(total_categories)))
# temp = [ item.split('__', 1) for item in total_categories]
# get most specific category
most_specific_category.sort()
checkNULL(most_specific_category)
print("total number of recorded categories are " + str(len(most_specific_category)))
# create dict "dic_category_id" store { category_name : id}
dic_category_id = {}
count = 1
for category in total_categories:
if category != "NULL" and category not in dic_category_id:
dic_category_id[category] = count
count = count + 1
total_categories.sort()
total_categories
len(df)
# creat categoryId column at dataframe
df_categoryId = []
for i in range(len(df)):
category = df.iloc[i]['mostSpecificCategory']
if category in dic_category_id:
df_categoryId.append(dic_category_id[category])
else:
print("Not recorded category for entity " + str(i))
break;
df['categoryId'] = df_categoryId
df_categoryId = []
for i in range(len(df2)):
category = df2.iloc[i]['mostSpecificCategory']
if category in dic_category_id:
df_categoryId.append(dic_category_id[category])
else:
print("Not recorded category for entity " + str(i))
break;
df2['categoryId'] = df_categoryId
print(df.columns)
print(df2.columns)
# create economy_df
economy_df = pd.DataFrame(columns = ['gazetteerId', 'categoryId', 'startYear', 'endYear', 'data'])
years = [str(i) for i in range(1949,2020)]
dic_for_economy_df = {'gazetteerId':[], 'categoryId':[], 'startYear':[], 'endYear':[], 'data':[]}
# Process yearly data
for i in range(len(df)):# each row
for year in years: # 1949 - 2019
if df.iloc[i][year] != "NULL":
dic_for_economy_df['gazetteerId'].append(df.iloc[i]['村志代码 Gazetteer Code'])
dic_for_economy_df['categoryId'].append(df.iloc[i]['categoryId'])
dic_for_economy_df['startYear'].append(int(year))
dic_for_economy_df['endYear'].append(int(year))
dic_for_economy_df['data'].append(df.iloc[i][year])
# dic_for_economy_df['unitId'].append(dic_for_unitId[df.iloc[i]['Unit']])
# Process range data
for i in range(len(df2)):
dic_for_economy_df['gazetteerId'].append(df2.iloc[i]['村志代码 Gazetteer Code'])
dic_for_economy_df['categoryId'].append(df2.iloc[i]['categoryId'])
dic_for_economy_df['startYear'].append(df2.iloc[i]['Start Year'])
dic_for_economy_df['endYear'].append(df2.iloc[i]['End Year'])
dic_for_economy_df['data'].append(df2.iloc[i]['Data'])
# dic_for_economy_df['unitId'].append(dic_for_unitId[df.iloc[i]['Unit']])
for attribute in economy_df.columns:
economy_df[attribute] = dic_for_economy_df[attribute]
economy_df.head()
# create economyCategory_df
economyCategory_df = pd.DataFrame(columns = ['id', 'name', 'parentId'])
dic_for_ecoCategorydf = {'id':[], 'name':[], 'parentId':[]}
for category in dic_category_id:
child_parent = category.split('__', 1)
name = child_parent[0]
if len(child_parent) == 1:
dic_for_ecoCategorydf['id'].append(dic_category_id[category])
dic_for_ecoCategorydf['name'].append(name)
dic_for_ecoCategorydf['parentId'].append("NULL")
else:
parentId = dic_category_id[child_parent[1]]
dic_for_ecoCategorydf['id'].append(dic_category_id[category])
dic_for_ecoCategorydf['name'].append(name)
dic_for_ecoCategorydf['parentId'].append(parentId)
for attribute in economyCategory_df.columns:
economyCategory_df[attribute] = dic_for_ecoCategorydf[attribute]
len(economyCategory_df)
# creat economyUnitCategory_df
# economyUnitCategory_df = pd.DataFrame(columns = ['id', 'name'])
# dic_for_economyUnitCategory_df = {'id':[], 'name':[]}
# for unit_name in dic_for_unitId:
# dic_for_economyUnitCategory_df['id'].append(dic_for_unitId[unit_name])
# dic_for_economyUnitCategory_df['name'].append(unit_name)
# for attribute in economyUnitCategory_df.columns:
# economyUnitCategory_df[attribute] = dic_for_economyUnitCategory_df[attribute]
# len(economyUnitCategory_df)
economy_df.to_csv('education_df.csv', index = False, na_rep = "NULL")
economyCategory_df.to_csv('educationCategory.csv', index = False, na_rep = "NULL")
# economyUnitCategory_df.to_csv('economyUnitCategory_df.csv', index = False, na_rep = "NULL")
```
| github_jupyter |
```
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
from astropy.table import Table
from lightkurve import search_lightcurvefile
from requests.exceptions import HTTPError
from astropy.coordinates import SkyCoord
from astroquery.mast import Tesscut
from astropy.timeseries import LombScargle
from photutils import CircularAperture, aperture_photometry
import healpy as hp
import astropy.units as u
coord = SkyCoord.from_name('K2-100')
slcf = search_lightcurvefile(coord, mission='K2')
lc = slcf.download_all()
pdcsap = lc.PDCSAP_FLUX.stitch()
time = pdcsap.time
flux = pdcsap.flux
notnanflux = ~np.isnan(flux)
flux = flux[notnanflux & (time > 3422) & (time < 3445)]
time = time[notnanflux & (time > 3422) & (time < 3445)]
flux /= np.polyval(np.polyfit(time - time.mean(), flux, 1), time-time.mean())
pdcsap.plot()
# plt.errorbar(time % best_period.value, flux, 0.002, ecolor='silver');
periods = np.linspace(1, 20, 1000) * u.day
freqs = 1 / periods
powers = LombScargle(time[~np.isnan(flux)] * u.day, flux[~np.isnan(flux)]).power(freqs)
best_period = periods[powers.argmax()]
plt.plot(periods, powers)
best_period
from fleck import Star
from emcee import EnsembleSampler
from multiprocessing import Pool
from corner import corner
u_ld = [0.46, 0.11]
contrast = 0.7
phases = (time % best_period.value) / best_period.value
s = Star(contrast, u_ld, n_phases=len(time), rotation_period=best_period.value)
# init_lons = np.array([0, 40, 290])
# init_lats = [0, 60, 0]
# init_rads = [0.01, 0.3, 0.38]
init_lons = np.array([0, 350, 250])
init_lats = [0, 20, 0]
init_rads = [0.01, 0.1, 0.2]
yerr = 0.001
init_p = np.concatenate([init_lons, init_lats, init_rads])
p = init_p.copy()
lons = p[0:3]
lats = p[3:6]
rads = p[6:9]
lc = s.light_curve(lons[:, None] * u.deg, lats[:, None] * u.deg, rads[:, None],
inc_stellar=90*u.deg, times=time, time_ref=0)
lc /= np.mean(lc)
plt.plot(time, flux)
plt.plot(time, lc)
def log_likelihood(p):
lons = p[0:3]
lats = p[3:6]
rads = p[6:9]
lc = s.light_curve(lons[:, None] * u.deg, lats[:, None] * u.deg, rads[:, None],
inc_stellar=90*u.deg, times=time, time_ref=0)[:, 0]
# plt.plot(time, lc/lc.max())
# plt.plot(time, flux)
# plt.show()
return - 0.5 * np.sum((lc/np.mean(lc) - flux)**2 / yerr**2)
def log_prior(p):
lons = p[0:3]
lats = p[3:6]
rads = p[6:9]
# if (np.all(rads < 1.) and np.all(rads > 0) and np.all(lats > -90) and
# np.all(lats < 90) and np.all(lons > 0) and np.all(lons < 360)):
if (np.all(rads < 1.) and np.all(rads > 0) and np.all(lats > -60) and
np.all(lats < 60) and np.all(lons > 0) and np.all(lons < 360)):
return 0
return -np.inf
def log_probability(p):
lp = log_prior(p)
if not np.isfinite(lp):
return -np.inf
return lp + log_likelihood(p)
ndim = len(init_p)
nwalkers = 2 * ndim
nsteps = 50000
pos = []
while len(pos) < nwalkers:
trial = init_p + 0.01 * np.random.randn(ndim)
lp = log_prior(trial)
if np.isfinite(lp):
pos.append(trial)
plt.errorbar(time, flux, yerr, ecolor='silver')
for p in pos:
lons = p[0:3]
lats = p[3:6]
rads = p[6:9]
lc = s.light_curve(lons[:, None] * u.deg, lats[:, None] * u.deg, rads[:, None],
inc_stellar=90*u.deg, times=time)[:, 0]
lc /= np.mean(lc)
plt.plot(time, lc, color='k')
with Pool() as pool:
sampler = EnsembleSampler(nwalkers, ndim, log_probability, pool=pool)
sampler.run_mcmc(pos, nsteps, progress=True);
# sampler = EnsembleSampler(nwalkers, ndim, log_probability)
# sampler.run_mcmc(pos, nsteps, progress=True);
samples_burned_in = sampler.flatchain[len(sampler.flatchain)//2:, :]
np.save('data/k2100_samples.npy', samples_burned_in)
corner(samples_burned_in);
samples_burned_in = np.load('data/k2100_samples.npy')
fig, ax = plt.subplots(1, 3, figsize=(8, 1))
for i in np.random.randint(0, len(samples_burned_in), size=50):
trial = samples_burned_in[i, :]
lons = trial[0:3]
lats = trial[3:6]
rads = trial[6:9]
lc = s.light_curve(lons[:, None] * u.deg, lats[:, None] * u.deg, rads[:, None],
inc_stellar=90*u.deg, times=time, time_ref=0)[:, 0]
ax[0].plot(time, lc/lc.mean(), color='DodgerBlue', alpha=0.05)
f_S = np.sum(samples_burned_in[:, -3:]**2 / 4, axis=1)
ax[1].hist(f_S, bins=25, histtype='step', lw=2, color='k', range=[0, 0.12], density=True)
ax[0].set(xlabel='BJD - 2454833', ylabel='Flux', xticks=[2230, 2233, 2236, 2239])
ax[1].set_xlabel('$f_S$')
ax[0].plot(time, flux, ',', color='k')
NSIDE = 2**10
NPIX = hp.nside2npix(NSIDE)
m = np.zeros(NPIX)
# for lon, lat, rad in np.median(samples_burned_in, axis=0).reshape((3, 3)).T:
np.random.seed(0)
random_index = np.random.randint(samples_burned_in.shape[0]//2,
samples_burned_in.shape[0])
random_sample = samples_burned_in[random_index].reshape((3, 3)).T
for lon, lat, rad in random_sample:
t = np.radians(lat + 90)
p = np.radians(lon)
spot_vec = hp.ang2vec(t, p)
ipix_spots = hp.query_disc(nside=NSIDE, vec=spot_vec, radius=rad)
m[ipix_spots] = 0.7
cmap = plt.cm.Greys
cmap.set_under('w')
plt.axes(ax[2])
hp.mollview(m, cbar=False, title="", cmap=cmap, hold=True,
max=1.0, notext=True, flip='geo')
hp.graticule(color='silver')
fig.suptitle('K2-100')
for axis in ax:
for sp in ['right', 'top']:
axis.spines[sp].set_visible(False)
fig.savefig('plots/k2100.pdf', bbox_inches='tight')
lo, mid, hi = np.percentile(f_S, [16, 50, 84])
f"$f_S = {{{mid:g}}}^{{+{hi-mid:g}}}_{{-{mid-lo:g}}}$"
np.save('data/k2100.npy', [mid, hi-mid, mid-lo])
```
| github_jupyter |
# 1.0 McNulty Exploratory Visualizations (Draft)
```
import mcnultymod
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.basemap import Basemap
import seaborn as sns
plt.style.use('seaborn')
%matplotlib inline
# import os
from datetime import datetime, timedelta, date
auctions = pd.read_pickle('../data/auctionsclosed.pkl')
auctions.sample(5)
```
### Purchaser Type
```
pt_ratios = []
for i in range(0,2):
pt_ratios.append((auctions.purchasertype.value_counts()[i] / auctions.shape[0]) * 100)
x_h = [0]
y_h = pt_ratios[0]
x_i = [1]
y_i = pt_ratios[1]
plt.figure(figsize=(10,9))
plt.bar(x_h, y_h, color='cyan', label='Homebuyer')
plt.bar(x_i, y_i, color='#00A2FF', label='Investor')
#plt.ylim(0,1)
plt.xticks([0, 1], ['', ''])
plt.xlabel("Purchaser Type", fontsize=14)
plt.ylabel("Percentage", fontsize=14)
plt.legend(fontsize=16)
plt.title("Proportions of Purchaser Types", fontsize=24)
#plt.savefig('../img/ptypes_bar.png', dpi=200, bbox_inches = 'tight')
;
```
### Price
```
auctions.price[auctions.purchasertype == 'Homebuyer'][auctions.price < 5000].shape
auctions.price[auctions.purchasertype == 'Investor'][auctions.price < 5000].shape
x = auctions.price
# for buyer in ['Homebuyer', 'Investor']:
# x_sub = np.array(auctions.price[auctions.purchasertype == buyer])
# x.append(x_sub)
plt.figure(figsize=(16,8))
plt.hist(x, bins=20, range=[0,100000], color='#FDCA40')
# plt.xticks([n*500 for n in range(21)], [n*500 for n in range(21)])
plt.xlabel("Prices", fontsize=14)
plt.ylabel("Frequency", fontsize=14)
# plt.legend(fontsize=12)
plt.title("Histogram of Closing Prices", fontsize=24)
# plt.savefig('../img/hist_price.png', dpi=200, bbox_inches = 'tight')
;
x = []
for buyer in ['Homebuyer', 'Investor']:
x_sub = np.array(auctions.price[auctions.purchasertype == buyer])
x.append(x_sub)
plt.figure(figsize=(16,8))
plt.hist(x, bins=20, color=['cyan', '#00A2FF'], label=['Homebuyer', 'Investor'])
# plt.xticks([n*5 for n in range(21)])
plt.xlabel("Prices", fontsize=14)
plt.ylabel("Frequency", fontsize=14)
plt.legend(fontsize=12)
plt.title("Histogram of Closing Prices by Puchaser Type", fontsize=24)
#plt.savefig('../img/hist_price_sep.png', dpi=200, bbox_inches = 'tight')
;
# this was working and now it's not!!!
plt.figure(figsize=(16,8))
# pal = {purchasertype: "#00A2FF" if purchasertype == "Investor" else "cyan" for purchasertype in auctions.purchasertype.unique()}
sns.boxplot(x=auctions.price,
y=auctions.purchasertype
# data=auctions,
# color=['cyan', 'cornflowerblue'],
# palette=pal
)
plt.xlabel("Closing Price (Dollars)", fontsize=16)
plt.ylabel("Purchaser Type", fontsize=16)
plt.yticks(range(2), ['Homebuyer', 'Investor'], rotation=90)
plt.xticks([n*5000 for n in range(21)], [str(5*n)+'K' for n in range(21)])
plt.title("Closing Price by Purchaser Type", fontsize=24)
# plt.savefig('../img/purchaser_box.png', dpi=200, bbox_inches = 'tight')
;
x = []
for buyer in ['Homebuyer', 'Investor']:
x_sub = np.array(auctions.price[auctions.purchasertype == buyer])
x.append(x_sub)
plt.figure(figsize=(16,9))
plt.hist(x, bins=20, color=['cyan', '#00A2FF'], label=['Homebuyer', 'Investor'])
plt.yscale('log');
# plt.xticks([n*5 for n in range(21)])
plt.xlabel("Prices", fontsize=14)
plt.ylabel("Frequency (log scale)", fontsize=14)
plt.legend(fontsize=12)
plt.title("Histogram of Closing Prices (log)", fontsize=24)
;
```
Hard to tell anything from that... perhaps this is better for now:
```
x = []
for buyer in ['Homebuyer', 'Investor']:
x_sub = np.array(auctions.price[auctions.purchasertype == buyer])
x.append(x_sub)
fig, ax = plt.subplots(1, 2, figsize=(16,7))
fig.suptitle("Histograms of Closing Price by Buyer Type", fontsize=24)
ax[0].hist(x[0], bins=20, range=[0,100000], color='#00FFFF')
ax[0].set_xlabel("Price (Dollars)", fontsize=12)
ax[0].set_ylabel("Frequency")
ax[0].set_title("Homebuyers", fontsize=14)
ax[1].hist(x[1], bins=20, range=[0,100000], color='#00A2FF')
ax[1].set_xlabel("Price (Dollars)", fontsize=12)
ax[1].set_ylabel("Frequency")
ax[1].set_title("Investor", fontsize=14)
# plt.savefig('../img/hist_price_x2.png', dpi=200, bbox_inches = 'tight')
;
x = []
for buyer in ['Homebuyer', 'Investor']:
x_sub = np.array(auctions.price[auctions.purchasertype == buyer])
x.append(x_sub)
fig, ax = plt.subplots(1, 2, figsize=(16,7))
fig.suptitle("Histograms of Closing Price by Buyer Type (> $5000)", fontsize=18)
ax[0].hist(x[0], bins=19, range=[5000,100000], color='c')
ax[0].set_xlabel("Price (Dollars)")
ax[0].set_ylabel("Frequency")
ax[0].set_title("Homebuyers")
ax[1].hist(x[1], bins=19, range=[5000,100000], color='#00A2FF')
ax[1].set_xlabel("Price (Dollars)")
ax[1].set_ylabel("Frequency")
ax[1].set_title("Investor")
# plt.figure(figsize=(16,8))
# plt.hist(x, bins=20, color=['cyan', 'blue'], label=['Homebuyer', 'Investor']);
# plt.xticks([n*5 for n in range(21)])
# plt.xlabel("Price", fontsize=12)
# plt.ylabel("Frequency", fontsize=12)
# plt.title("Histogram of Neighborhood Occurrences", fontsize=18)
# plt.savefig('../img/hists_price_over5k.png', dpi=200, bbox_inches = 'tight')
;
price_counts_norm = []
for buyer in ['Homebuyer', 'Investor']:
buyer_count = []
total = auctions[auctions.purchasertype == buyer].shape[0]
for n in range(18):
count = (auctions[(auctions.purchasertype == buyer) &
(auctions.price >= n*5000) &
(auctions.price < (n*5000 + 5000))].shape[0]
)
buyer_count.append(count / total)
price_counts_norm.append(buyer_count)
x = np.array(range(0,18))
plt.figure(figsize=(16,9))
plt.bar(x - 0.2,
price_counts_norm[0],
color='cyan',
width=0.4,
align='center',
label='Homebuyers')
plt.bar(x + 0.2,
price_counts_norm[1],
color='#00A2FF',
width=0.4,
align='center',
label='Investor')
plt.legend()
plt.xticks(range(18), [n*5000 + 5000 for n in range(18)])
plt.xlabel("District", fontsize=12)
plt.ylabel("Price (Dollars)", fontsize=12)
plt.title("Normalized Frequency of Purchaser Types by Price ($5000 Increments)", fontsize=18)
# plt.savefig('../img/bar_districts.png', dpi=200, bbox_inches = 'tight')
;
```
### Neighborhoods
```
plt.figure(figsize=(16,8))
plt.hist(auctions.neighborhood.value_counts(), bins=18, range=[0,90], color='#44AF69');
plt.xticks([n*5 for n in range(19)])
plt.xlabel("Neighborhood Counts", fontsize=14)
plt.ylabel("Frequency", fontsize=14)
plt.title("Histogram of Neighborhood Occurrences", fontsize=24)
# plt.savefig('../img/hist_nhood.png', dpi=200, bbox_inches = 'tight')
;
```
### Concentration of Investors per Neighborhood
```
# calculate investor ratio overall
inv_ratio_all = auctions.purchasertype.value_counts()[1] / auctions.shape[0]
inv_ratio_all
# (auctions
# .groupby('neighborhood')
# .count()
# .filter(['neighborhood', 'parcelid'])
# .sort_values('parcelid', ascending=False)
# .rename(columns={'index': 'neighborhood', 'parcelid': 'n_count'})
# .reset_index()
# )
# performs same function as above with fewer lines of code
nhcounts = (auctions
.neighborhood
.value_counts()
.reset_index()
.rename(columns={'index': 'neighborhood', 'neighborhood': 'n_count'})
)
# most frequent neighborhoods
nhcounts.head(10)
def add_inv_ratios(row):
"""Calculates investor ratio for each neighborhood, use via .apply() method"""
nhood = row[0] # same as row.neighborhood
auc_sub = auctions[auctions.neighborhood == nhood]
inv_ratio = auc_sub[auc_sub.purchasertype == 'Investor'].shape[0] / auc_sub.shape[0]
return inv_ratio
nhcounts['inv_ratio'] = nhcounts.apply(add_inv_ratios, axis=1)
nhcounts.head(10)
# adds ratio of investors to neighborhoods
### THIS IS NO LONGER IN MODULE AS OF 3/23... AND WHY??
nhcounts = mcnultymod.nhood_investor_ratios(auctions, nhcounts)
nhcounts.head()
def rand_investor_ratios(row):
"""For each neighborhood, pulls random sample of n properties from auctions and calculates
investor ratio, where n is the number of properties in that neighborhood. For use with
.apply() method on nhcounts df"""
sample = auctions.sample(row[1]) # same as row.n_count
sample_inv_ratio = sample[sample.purchasertype == 'Investor'].shape[0] / row[1]
return sample_inv_ratio
ir_all = inv_ratio_all
x = nhcounts.n_count
y = nhcounts.inv_ratio
# for each neighborhood, pull a random sample of properties from auctions of equal number
y_rand = nhcounts.apply(rand_investor_ratios, axis=1)
x_lin = np.linspace(1,100, num=99)
y_avg = [ir_all] * len(x_lin)
# y_2sd_pos = ir_all + (2 * np.sqrt(x_lin * ir_all * (1-ir_all)) / x_lin)
# y_2sd_neg = ir_all - (2 * np.sqrt(x_lin * ir_all * (1-ir_all)) / x_lin)
plt.figure(figsize=(16,9))
plt.plot(x_lin, y_avg, 'r--', label='Proportion of Investors, All Properties (0.31)')
# plt.plot(x_lin, y_2sd_pos, 'g--', alpha=0.5, label='Expected Bounds of 95% of Inv. Proportions')
plt.plot(x_lin, y_2sd_neg, 'g--', alpha=0.5)
plt.plot(x, y, 'o', color='#00A2FF', alpha=0.8, label="Proportion of Investors per Neighborhood")
# plt.plot(x, y_rand, 'ko', alpha=0.5, label="Proportion of Investors per Random Sample") # color='#44AF69'
plt.legend(fontsize=12)
plt.xlabel("Properties in Neighborhood / Sample", fontsize=14)
plt.ylabel("Proportion of Investors", fontsize=14)
plt.ylim(-0.05, 1.05)
plt.title("Investor Proportions as Neighborhood Size Increases", fontsize=24);
# plt.savefig('../viz/nhood_inv.png', dpi=200, bbox_inches = 'tight')
```
### Council District
```
auctions[auctions.purchasertype == 'Homebuyer'].groupby('councildistrict').purchasertype.count()
counts_norm = []
for buyer in ['Homebuyer', 'Investor']:
counts = np.array(auctions[auctions.purchasertype == buyer]
.groupby('councildistrict')
.purchasertype
.count()
)
counts = counts / counts.sum()
counts_norm.append(counts)
x = np.array(range(1,8))
plt.figure(figsize=(16,8))
plt.bar(x - 0.2,
counts_norm[0],
color='cyan',
width=0.4,
align='center',
label='Homebuyers')
plt.bar(x + 0.2,
counts_norm[1],
color='#00A2FF',
width=0.4,
align='center',
label='Investor')
plt.legend()
# plt.xticks([-0.25,0.25], ['Democrats', 'Republicans'])
plt.xlabel("District", fontsize=12)
plt.ylabel("Normalized Frequency", fontsize=12)
plt.title("Normalized Frequency of Purchaser Types per Council District", fontsize=18)
# plt.savefig('../img/bar_districts.png', dpi=200, bbox_inches = 'tight')
;
counts_norm[0].sum()
```
### Maps!
```
inv_lats = list(auctions[auctions.purchasertype == 'Investor']['latitude'])
inv_lons = list(auctions[auctions.purchasertype == 'Investor']['longitude'])
hb_lats = list(auctions[auctions.purchasertype == 'Homebuyer']['latitude'])
hb_lons = list(auctions[auctions.purchasertype == 'Homebuyer']['longitude'])
plt.figure(figsize=(16, 10))
m = Basemap(projection='merc', # 'merc', 'lcc'
resolution='i', # 'c', 'l', 'i', 'h', 'f', None
area_thresh=0.1,
# width=36000,
# height=24000,
# lat_0=42.3313889, lon_0=-83.0458333,
llcrnrlon=-83.30, llcrnrlat=42.245,
urcrnrlon=-82.89, urcrnrlat=42.465,
)
# m.drawcountries()
# m.drawcoastlines()
# m.drawstates()
# m.fillcontinents(color='white', lake_color='aqua')
m.drawmapboundary(fill_color='white')
# m.drawrivers(color='blue')
# m.bluemarble(scale=0.2)
# m.readshapefile("../data/river/AOC_MI_DETROIT", 'river2')
# m.readshapefile("../data/tract10_mi/tract10_miv14a", 'cities') # needs reformatting
m.readshapefile("../data/mi_admin/michigan_administrative", 'admin')
# m.readshapefile("../data/mi_hwy/michigan_highway", 'hwy') # too much!!
m.readshapefile("../data/neighborhoods/geo_export_333d2406-1960-4afa-a078-0ff790c78e70", 'neighborhoods')
# m.readshapefile("../data/zipcodes/geo_export_681b42b5-2249-4750-9e61-6e2271a3974d", 'zips')
# m.readshapefile("../data/councildistricts/geo_export_cafa6f85-7226-42e7-a1e1-2f808c72737b", 'districts')
# m.readshapefile("../data/mi_coast/michigan_coastline", 'river')
xh, yh = m(hb_lons, hb_lats)
xi, yi = m(inv_lons, inv_lats)
plt.plot(xi, yi, 'o', color='#0000FF', markersize=6, alpha=0.5, label="Investor")
plt.plot(xh, yh, 'co', markersize=6, alpha=0.5, label="Homebuyer")
plt.legend(loc='lower right', fontsize=16)
plt.title("DLBA Closed Auctions", fontsize=24)
# plt.savefig('../img/map_district.png', dpi=200, bbox_inches = 'tight')
# plt.savefig('../img/map_nhood.png', dpi=200, bbox_inches = 'tight')
;
```
| github_jupyter |
```
cd E:\NEURAL TRANSFER
import os
import sys
import scipy.io
import scipy.misc
import matplotlib.pyplot as plt
from matplotlib.pyplot import imshow
from PIL import Image
import numpy as np
import tensorflow as tf
from keras.preprocessing import image
%matplotlib inline
model = load_vgg_model("imagenet-vgg-verydeep-19")
print(model)
# GRADED FUNCTION: compute_content_cost
def compute_content_cost(a_C, a_G):
"""
Computes the content cost
Arguments:
a_C -- tensor of dimension (1, n_H, n_W, n_C), hidden layer activations representing content of the image C
a_G -- tensor of dimension (1, n_H, n_W, n_C), hidden layer activations representing content of the image G
Returns:
J_content -- scalar that you compute using equation 1 above.
"""
### START CODE HERE ###
# Retrieve dimensions from a_G (≈1 line)
m, n_H, n_W, n_C = a_G.get_shape().as_list();
# Reshape a_C and a_G (≈2 lines)
a_C_unrolled = tf.reshape(a_C,shape=(m,n_H*n_W,n_C));
a_G_unrolled = tf.reshape(a_G,shape=(m,n_H*n_W,n_C));
# compute the cost with tensorflow (≈1 line)
J_content = (1/(4*n_H*n_W*n_C))*(tf.reduce_sum(tf.square(tf.subtract(a_C_unrolled,a_G_unrolled))));
### END CODE HERE ###
return J_content
# GRADED FUNCTION: gram_matrix
def gram_matrix(A):
"""
Argument:
A -- matrix of shape (n_C, n_H*n_W)
Returns:
GA -- Gram matrix of A, of shape (n_C, n_C)
"""
### START CODE HERE ### (≈1 line)
GA = tf.matmul(A,tf.transpose(A));
### END CODE HERE ###
return GA
# GRADED FUNCTION: compute_layer_style_cost
def compute_layer_style_cost(a_S, a_G):
"""
Arguments:
a_S -- tensor of dimension (1, n_H, n_W, n_C), hidden layer activations representing style of the image S
a_G -- tensor of dimension (1, n_H, n_W, n_C), hidden layer activations representing style of the image G
Returns:
J_style_layer -- tensor representing a scalar value, style cost defined above by equation (2)
"""
### START CODE HERE ###
# Retrieve dimensions from a_G (≈1 line)
m, n_H, n_W, n_C = a_G.get_shape().as_list()
# Reshape the images to have them of shape (n_C, n_H*n_W) (≈2 lines)
a_S = tf.reshape(tf.transpose(a_S),shape=(n_C,n_H*n_W)); # carefull only changing shape will
#not work u need to take transpose as well
a_G = tf.reshape(tf.transpose(a_G),shape=(n_C,n_H*n_W));
# Computing gram_matrices for both images S and G (≈2 lines)
GS = gram_matrix(a_S)
GG = gram_matrix(a_G)
# Computing the loss (≈1 line)
J_style_layer = (1/(4*n_C*n_C*(n_H*n_W)*(n_H*n_W)))*tf.reduce_sum(tf.square(tf.subtract(GS,GG)));
### END CODE HERE ###
return J_style_layer
STYLE_LAYERS = [
('conv1_1', 0.2),
('conv2_1', 0.2),
('conv3_1', 0.2),
('conv4_1', 0.2),
('conv5_1', 0.2)]
def compute_style_cost(model, STYLE_LAYERS):
"""
Computes the overall style cost from several chosen layers
Arguments:
model -- our tensorflow model
STYLE_LAYERS -- A python list containing:
- the names of the layers we would like to extract style from
- a coefficient for each of them
Returns:
J_style -- tensor representing a scalar value, style cost defined above by equation (2)
"""
# initialize the overall style cost
J_style = 0
for layer_name, coeff in STYLE_LAYERS:
# Select the output tensor of the currently selected layer
out = model[layer_name]
# Set a_S to be the hidden layer activation from the layer we have selected, by running the session on out
a_S = sess.run(out)
# Set a_G to be the hidden layer activation from same layer. Here, a_G references model[layer_name]
# and isn't evaluated yet. Later in the code, we'll assign the image G as the model input, so that
# when we run the session, this will be the activations drawn from the appropriate layer, with G as input.
a_G = out
# Compute style_cost for the current layer
J_style_layer = compute_layer_style_cost(a_S, a_G)
# Add coeff * J_style_layer of this layer to overall style cost
J_style += coeff * J_style_layer
return J_style
# GRADED FUNCTION: total_cost
def total_cost(J_content, J_style, alpha = 10, beta = 40):
"""
Computes the total cost function
Arguments:
J_content -- content cost coded above
J_style -- style cost coded above
alpha -- hyperparameter weighting the importance of the content cost
beta -- hyperparameter weighting the importance of the style cost
Returns:
J -- total cost as defined by the formula above.
"""
### START CODE HERE ### (≈1 line)
J = (alpha*J_content)+(beta*(J_style));
### END CODE HERE ###
return J
# Reset the graph
tf.reset_default_graph()
# Start interactive session
sess = tf.InteractiveSession()
##Content Image
content_image = scipy.misc.imread("Content/saksham.jpg")
content_image= reshape_and_normalize(content_image)
##Style Image
style_image = scipy.misc.imread("Styles/style2.jpg")
style_image = reshape_and_normalize(style_image)
generated_image = generate_noise_image(content_image)
model = load_vgg_model("imagenet-vgg-verydeep-19.mat")
# Assign the content image to be the input of the VGG model.
sess.run(model['input'].assign(content_image))
# Select the output tensor of layer conv4_2
out = model['conv4_2']
# Set a_C to be the hidden layer activation from the layer we have selected
a_C = sess.run(out)
# Set a_G to be the hidden layer activation from same layer. Here, a_G references model['conv4_2']
# and isn't evaluated yet. Later in the code, we'll assign the image G as the model input, so that
# when we run the session, this will be the activations drawn from the appropriate layer, with G as input.
a_G = out
# Compute the content cost
J_content = compute_content_cost(a_C, a_G)
# Assign the input of the model to be the "style" image
sess.run(model['input'].assign(style_image))
# Compute the style cost
J_style = compute_style_cost(model, STYLE_LAYERS)
### START CODE HERE ### (1 line)
J = total_cost(J_content, J_style, alpha = 10, beta = 40)
### END CODE HERE ###
# define optimizer (1 line)
optimizer = tf.train.AdamOptimizer(2.0)
# define train_step (1 line)
train_step = optimizer.minimize(J)
def model_nn(sess, input_image, num_iterations = 200):
# Initialize global variables (you need to run the session on the initializer)
### START CODE HERE ### (1 line)
init = tf.global_variables_initializer();
sess.run(init)
### END CODE HERE ###
# Run the noisy input image (initial generated image) through the model. Use assign().
### START CODE HERE ### (1 line)
sess.run(model['input'].assign(input_image))
### END CODE HERE ###
for i in range(num_iterations):
# Run the session on the train_step to minimize the total cost
### START CODE HERE ### (1 line)
sess.run([train_step])
### END CODE HERE ###
# Compute the generated image by running the session on the current model['input']
### START CODE HERE ### (1 line)
generated_image = sess.run(model['input'])
### END CODE HERE ###
# Print every 20 iteration.
if i%20 == 0:
Jt, Jc, Js = sess.run([J, J_content, J_style])
print("Iteration " + str(i) + " :")
print("total cost = " + str(Jt))
print("content cost = " + str(Jc))
print("style cost = " + str(Js))
# save current generated image in the "/output" directory
save_image("output/" + str(i)+"a" + ".png", generated_image)
# save last generated image
save_image('output/generated_image1.jpg', generated_image)
return generated_image
import os
os.mkdir('output')
model_nn(sess, generated_image)
```
| github_jupyter |
# Forecasting with a stateful RNN
## Setup
```
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
keras = tf.keras
def plot_series(time, series, format="-", start=0, end=None, label=None):
plt.plot(time[start:end], series[start:end], format, label=label)
plt.xlabel("Time")
plt.ylabel("Value")
if label:
plt.legend(fontsize=14)
plt.grid(True)
def trend(time, slope=0):
return slope * time
def seasonal_pattern(season_time):
"""Just an arbitrary pattern, you can change it if you wish"""
return np.where(season_time < 0.4,
np.cos(season_time * 2 * np.pi),
1 / np.exp(3 * season_time))
def seasonality(time, period, amplitude=1, phase=0):
"""Repeats the same pattern at each period"""
season_time = ((time + phase) % period) / period
return amplitude * seasonal_pattern(season_time)
def white_noise(time, noise_level=1, seed=None):
rnd = np.random.RandomState(seed)
return rnd.randn(len(time)) * noise_level
time = np.arange(4 * 365 + 1)
slope = 0.05
baseline = 10
amplitude = 40
series = baseline + trend(time, slope) + seasonality(time, period=365, amplitude=amplitude)
noise_level = 5
noise = white_noise(time, noise_level, seed=42)
series += noise
plt.figure(figsize=(10, 6))
plot_series(time, series)
plt.show()
split_time = 1000
time_train = time[:split_time]
x_train = series[:split_time]
time_valid = time[split_time:]
x_valid = series[split_time:]
```
## Stateful RNN Forecasting
```
def sequential_window_dataset(series, window_size):
series = tf.expand_dims(series, axis=-1) # https://www.tensorflow.org/api_docs/python/tf/expand_dims
ds = tf.data.Dataset.from_tensor_slices(series)
ds = ds.window(window_size + 1, shift=window_size, drop_remainder=True)
ds = ds.flat_map(lambda window: window.batch(window_size + 1))
ds = ds.map(lambda window: (window[:-1], window[1:]))
return ds.batch(1).prefetch(1)
for X_batch, y_batch in sequential_window_dataset(tf.range(10), 3):
print(X_batch.numpy(), y_batch.numpy())
class ResetStatesCallback(keras.callbacks.Callback):
def on_epoch_begin(self, epoch, logs):
self.model.reset_states()
keras.backend.clear_session()
tf.random.set_seed(42)
np.random.seed(42)
window_size = 30
train_set = sequential_window_dataset(x_train, window_size)
model = keras.models.Sequential([
keras.layers.SimpleRNN(100, return_sequences=True, stateful=True,
batch_input_shape=[1, None, 1]),
keras.layers.SimpleRNN(100, return_sequences=True, stateful=True),
keras.layers.Dense(1),
keras.layers.Lambda(lambda x: x * 200.0)
])
lr_schedule = keras.callbacks.LearningRateScheduler(
lambda epoch: 1e-8 * 10**(epoch / 30))
reset_states = ResetStatesCallback()
optimizer = keras.optimizers.SGD(lr=1e-8, momentum=0.9)
model.compile(loss=keras.losses.Huber(),
optimizer=optimizer,
metrics=["mae"])
history = model.fit(train_set, epochs=100,
callbacks=[lr_schedule, reset_states])
plt.semilogx(history.history["lr"], history.history["loss"])
plt.axis([1e-8, 1e-4, 0, 30])
keras.backend.clear_session()
tf.random.set_seed(42)
np.random.seed(42)
window_size = 30
train_set = sequential_window_dataset(x_train, window_size)
valid_set = sequential_window_dataset(x_valid, window_size)
model = keras.models.Sequential([
keras.layers.SimpleRNN(100, return_sequences=True, stateful=True,
batch_input_shape=[1, None, 1]),
keras.layers.SimpleRNN(100, return_sequences=True, stateful=True),
keras.layers.Dense(1),
keras.layers.Lambda(lambda x: x * 200.0)
])
optimizer = keras.optimizers.SGD(lr=1e-7, momentum=0.9)
model.compile(loss=keras.losses.Huber(),
optimizer=optimizer,
metrics=["mae"])
reset_states = ResetStatesCallback()
model_checkpoint = keras.callbacks.ModelCheckpoint(
"my_checkpoint.h5", save_best_only=True)
early_stopping = keras.callbacks.EarlyStopping(patience=50)
model.fit(train_set, epochs=500,
validation_data=valid_set,
callbacks=[early_stopping, model_checkpoint, reset_states])
model = keras.models.load_model("my_checkpoint.h5")
model.reset_states()
rnn_forecast = model.predict(series[np.newaxis, :, np.newaxis])
rnn_forecast = rnn_forecast[0, split_time - 1:-1, 0]
rnn_forecast.shape
plt.figure(figsize=(10, 6))
plot_series(time_valid, x_valid)
plot_series(time_valid, rnn_forecast)
keras.metrics.mean_absolute_error(x_valid, rnn_forecast).numpy()
```
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
from sklearn.neighbors import KNeighborsClassifier
from sklearn import svm
# grAdapt
import grAdapt
from grAdapt.models import Sequential
from grAdapt.space.datatype import Integer, Float, Categorical
```
## 1. Introduction
This is a tutorial how to use **grAdapt**. The package itself is much more powerful and there are many possibilites to use this package. This notebook briefly introduces how to optimize blackbox functions. We will take a look at tuning hyperparameters for classification tasks.
### 1.1 Data Set
For the data set, we will use the **NIST data set** as it is small enough to train a classifier fast on it. Though the nature of the developed package is to optimize of float variables, we will take a look at integer and categorical optimization.
### 1.2 Hyperparameter Optimization
For demonstration, we will tune the hyperparameters for k-Nearest Neighbor and the Support Vector Machine. The k-Nearest Neighbor algorithm requires one hyperparameter. To show that grAdapt works with multiple input dimensions, three hyperparameters of the Support Vector Machine will be tuned where each hyperparameter can have a different type than the other.
#### Load MNIST data set
```
from sklearn import datasets, svm, metrics
from sklearn.model_selection import train_test_split
# The digits dataset
digits = datasets.load_digits()
# To apply a classifier on this data, we need to flatten the image, to
# turn the data in a (samples, feature) matrix:
n_samples = len(digits.images)
data = digits.images.reshape((n_samples, -1))
print(data.shape)
# Split data into train and validation subsets
X_train, X_val, y_train, y_val = train_test_split(
data, digits.target, test_size=0.8, shuffle=False)
```
## 2. Define Blackbox
The intuition behind defining a blackbox function it to fit the classifier to the training set and then returning the validation accuracy. The validation accuracy is to be maximized and the input variables of the blackbox function are the hyperparameters to be optimized. The hyperparameters are set when creating a classifier instance.
```
def knn_opt(n):
# create KNN instance
neigh = KNeighborsClassifier(n_neighbors=n)
# fit/train
neigh.fit(X_train, y_train)
# return validation accuracy
return neigh.score(X_val, y_val)
def svm_opt(C, gamma, kernel):
# create SVM instance
clf = svm.SVC(C=C, gamma=gamma, kernel=kernel)
# fit/train
clf.fit(X_train, y_train)
# return validation accuracy
return clf.score(X_val, y_val)
```
## 3. Hyperparameter Optimization with grAdapt
In this section, we are creating a **grAdapt Sequential** model. For brevity, settings of the Sequential model are not demonstrated. We will also take a look how to define search spaces by using **grAdapt Datatypes**.
### 3.1 k-Nearest Neighbor
k-Nearest Neighbor takes one hyperparameter which is the number of neighbors *n*. Because the number of neighbors is an integer, we are doing **integer programming**.
```
# create base model
model = Sequential()
# create bounds
bounds = [Integer(low=1, high=100)]
```
Optimize *knn_opt* function by the given *bounds*. Evaluate the blackbox function at most *500* times.
```
res = model.maximize_args(knn_opt, bounds, 500, show_progressbar=True)
```
**res** is a dictionary. The history can be accessed as in the following where
*x* contains the history of the hyperparameter settings and *y* contains the history of the validation accuracy.
```
y_knn = res['y']
x_knn = res['x']
```
#### 3.1.1 Training Plot
```
plt.plot(np.arange(y_knn.shape[0]), y_knn)
plt.title('Training plot')
plt.xlabel('History')
plt.ylabel('Validation accuracy')
plt.show()
```
#### 3.1.2 Hyperparameter Plot
```
plt.scatter(np.arange(x_knn.shape[0]), x_knn, s=3)
plt.title('Training plot')
plt.xlabel('History')
plt.ylabel('KNN n')
plt.show()
```
#### 3.1.3 Best Hyperparameter
```
x_knn_sol = res['x_sol']
y_knn_sol = res['y_sol']
print(x_knn_sol)
print(y_knn_sol)
```
Internally grAdapt uses floats and thus returns floats. To obtain the optimal hyperparameter when having integer variables, we have to manually round these. The real solution is then:
```
np.round(x_knn_sol)
```
with the achieved validation accuracy of:
```
print(y_knn_sol)
```
#### 3.1.4 Results
grAdapt slowly decreases *n* as this setting improves the performance. grAdapt still escapes from the local optima but higher values led to poor performances. Thus, the chances of obtaining higher values of *n* is decreased but not vanished (to ensure a global solution). A variation of *n* can be obtained through the whole history. It is very unlikely for grAdapt to obtain the same values twice (internally).
### 3.2 Support Vector Machine
Support Vector Machine takes three hyperparameters which are *C*, *gamma* and the *kernel* function. Both *C* and *gamma* are float variables. The last hyperparameter is a categorical variable which is a list of strings.
```
# create base model
model = Sequential()
# create bounds
c_var = Float(low=1e-3, high=100)
gamma_var = Float(low=1e-3, high=1e3)
kernel_var = Categorical(['linear', 'poly', 'sigmoid', 'rbf'])
bounds = [c_var, gamma_var, kernel_var]
res_svm = model.maximize_args(svm_opt, bounds, 500, show_progressbar=True)
y_svm = res_svm['y']
x_svm = res_svm['x']
```
#### 3.2.1 Training Plot
```
plt.plot(np.arange(y_svm.shape[0]), y_svm)
plt.title('Training plot')
plt.xlabel('History')
plt.ylabel('Validation accuracy')
plt.show()
```
#### 3.2.2 Hyperparameter Plot
##### 3.2.2.1 C value
```
plt.scatter(np.arange(x_svm.shape[0]), x_svm[:,0], s=3)
plt.title('Training plot')
plt.xlabel('History')
plt.ylabel('SVM C')
plt.show()
```
##### 3.2.2.2 gamma value
```
plt.scatter(np.arange(x_svm.shape[0]), x_svm[:,1], s=3)
plt.title('Training plot')
plt.xlabel('History')
plt.ylabel('SVM gamma')
plt.show()
```
##### 3.2.2.3 kernel
```
plt.scatter(np.arange(x_svm.shape[0]), x_svm[:,2], s=3)
plt.title('Training plot')
plt.xlabel('History')
plt.ylabel('SVM kernel')
plt.show()
```
#### 3.2.3 Best Hyperparameters
```
x_svm_sol = res_svm['x_sol']
y_svm_sol = res_svm['y_sol']
print(x_svm_sol)
print(y_svm_sol)
```
Real solutions:
```
x_svm_sol = x_svm_sol.tolist()
x_svm_sol[2] = kernel_var.get_category(x_svm_sol[2])
x_svm_sol
```
#### 3.2.4 Results
Optimizing three hyperparameters at once does not seem to be a problem for grAdapt. Even if the hyperparameters are of different datatypes (float+categorical).
It can be obtained that high *C* and low *gamma* values led to good performances. grAdapt often escapes but also stays local when good solutions were found. For the categorical variable, it can not be read which kernel function led to good performances. It seems that the kernel function is less important for the specific data set.
Because each variable can influence each other, it is astonishing to see that grAdapt handles this well and has found good settings for each parameter.
| github_jupyter |
# Feathr Feature Store on Azure Demo Notebook
This notebook illustrates the use of Feature Store to create a model that predicts NYC Taxi fares. It includes these steps:
This tutorial demonstrates the key capabilities of Feathr, including:
1. Install and set up Feathr with Azure
2. Create shareable features with Feathr feature definition configs.
3. Create a training dataset via point-in-time feature join.
4. Compute and write features.
5. Train a model using these features to predict fares.
6. Materialize feature value to online store.
7. Fetch feature value in real-time from online store for online scoring.
In this tutorial, we use Feathr Feature Store to create a model that predicts NYC Taxi fares. The dataset comes from [here](https://www1.nyc.gov/site/tlc/about/tlc-trip-record-data.page). The feature flow is as below:

## Prerequisite: Use Quick Start Template to Provision Azure Resources
Feathr has native cloud integration. To use Feathr on Azure, you only need three steps:
- Get the `Principal ID` of your account by running `az ad signed-in-user show --query objectId -o tsv` in the link below (Select "Bash" if asked), and write down that value (something like `b65ef2e0-42b8-44a7-9b55-abbccddeefff`). Think this ID as something representing you when accessing Azure, and it will be used to grant permissions in the next step in the UI.
[Launch Cloud Shell](https://shell.azure.com/bash)
- Click the button below to deploy a minimal set of Feathr resources for demo purpose. You will need to fill in the `Principal ID` and `Resource Prefix`. You will need "Owner" permission of the selected subscription.
[](https://portal.azure.com/#create/Microsoft.Template/uri/https%3A%2F%2Fraw.githubusercontent.com%2Flinkedin%2Ffeathr%2Fmain%2Fdocs%2Fhow-to-guides%2Fazure_resource_provision.json)
- Run the cells below.
And the architecture is as below. In the above template, we are using Synapse as Spark provider, use Azure Data Lake Gen2 as offline store, and use Redis as online store, Azure Purview (Apache Atlas compatible) as feature reigstry.

## Prerequisite: Install Feathr
Install Feathr using pip:
`pip install -U feathr pandavro scikit-learn`
## Prerequisite: Configure the required environment with Feathr Quick Start Template
In the first step (Provision cloud resources), you should have provisioned all the required cloud resources. Run the code below to install Feathr, login to Azure to get the required credentials to access more cloud resources.
**REQUIRED STEP: Fill in the resource prefix when provisioning the resources**
```
resource_prefix = "feathr_resource_prefix"
! pip install azure-cli
! pip install feathr pandavro scikit-learn
```
Login to Azure with a device code (You will see instructions in the output):
```
! az login --use-device-code
import glob
import os
import tempfile
from datetime import datetime, timedelta
from math import sqrt
import pandas as pd
import pandavro as pdx
from feathr import FeathrClient
from feathr import BOOLEAN, FLOAT, INT32, ValueType
from feathr import Feature, DerivedFeature, FeatureAnchor
from feathr import BackfillTime, MaterializationSettings
from feathr import FeatureQuery, ObservationSettings
from feathr import RedisSink
from feathr import INPUT_CONTEXT, HdfsSource
from feathr import WindowAggTransformation
from feathr import TypedKey
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import train_test_split
from azure.identity import DefaultAzureCredential
from azure.keyvault.secrets import SecretClient
```
Get all the required credentials from Azure KeyVault
```
# Get all the required credentials from Azure Key Vault
key_vault_name=resource_prefix+"kv"
synapse_workspace_url=resource_prefix+"syws"
adls_account=resource_prefix+"dls"
adls_fs_name=resource_prefix+"fs"
purview_name=resource_prefix+"purview"
key_vault_uri = f"https://{key_vault_name}.vault.azure.net"
credential = DefaultAzureCredential(exclude_interactive_browser_credential=False)
client = SecretClient(vault_url=key_vault_uri, credential=credential)
secretName = "FEATHR-ONLINE-STORE-CONN"
retrieved_secret = client.get_secret(secretName).value
# Get redis credentials; This is to parse Redis connection string.
redis_port=retrieved_secret.split(',')[0].split(":")[1]
redis_host=retrieved_secret.split(',')[0].split(":")[0]
redis_password=retrieved_secret.split(',')[1].split("password=",1)[1]
redis_ssl=retrieved_secret.split(',')[2].split("ssl=",1)[1]
# Set the resource link
os.environ['spark_config__azure_synapse__dev_url'] = f'https://{synapse_workspace_url}.dev.azuresynapse.net'
os.environ['spark_config__azure_synapse__pool_name'] = 'spark31'
os.environ['spark_config__azure_synapse__workspace_dir'] = f'abfss://{adls_fs_name}@{adls_account}.dfs.core.windows.net/feathr_project'
os.environ['feature_registry__purview__purview_name'] = f'{purview_name}'
os.environ['online_store__redis__host'] = redis_host
os.environ['online_store__redis__port'] = redis_port
os.environ['online_store__redis__ssl_enabled'] = redis_ssl
os.environ['REDIS_PASSWORD']=redis_password
os.environ['feature_registry__purview__purview_name'] = f'{purview_name}'
feathr_output_path = f'abfss://{adls_fs_name}@{adls_account}.dfs.core.windows.net/feathr_output'
```
## Prerequisite: Configure the required environment (Don't need to update if using the above Quick Start Template)
In the first step (Provision cloud resources), you should have provisioned all the required cloud resources. If you use Feathr CLI to create a workspace, you should have a folder with a file called `feathr_config.yaml` in it with all the required configurations. Otherwise, update the configuration below.
The code below will write this configuration string to a temporary location and load it to Feathr. Please still refer to [feathr_config.yaml](https://github.com/linkedin/feathr/blob/main/feathr_project/feathrcli/data/feathr_user_workspace/feathr_config.yaml) and use that as the source of truth. It should also have more explanations on the meaning of each variable.
```
import tempfile
yaml_config = """
# Please refer to https://github.com/linkedin/feathr/blob/main/feathr_project/feathrcli/data/feathr_user_workspace/feathr_config.yaml for explanations on the meaning of each field.
api_version: 1
project_config:
project_name: 'feathr_getting_started'
required_environment_variables:
- 'REDIS_PASSWORD'
- 'AZURE_CLIENT_ID'
- 'AZURE_TENANT_ID'
- 'AZURE_CLIENT_SECRET'
offline_store:
adls:
adls_enabled: true
wasb:
wasb_enabled: true
s3:
s3_enabled: false
s3_endpoint: 's3.amazonaws.com'
jdbc:
jdbc_enabled: false
jdbc_database: 'feathrtestdb'
jdbc_table: 'feathrtesttable'
snowflake:
url: "dqllago-ol19457.snowflakecomputing.com"
user: "feathrintegration"
role: "ACCOUNTADMIN"
spark_config:
spark_cluster: 'azure_synapse'
spark_result_output_parts: '1'
azure_synapse:
dev_url: 'https://feathrazuretest3synapse.dev.azuresynapse.net'
pool_name: 'spark3'
workspace_dir: 'abfss://feathrazuretest3fs@feathrazuretest3storage.dfs.core.windows.net/feathr_getting_started'
executor_size: 'Small'
executor_num: 4
feathr_runtime_location: wasbs://public@azurefeathrstorage.blob.core.windows.net/feathr-assembly-LATEST.jar
databricks:
workspace_instance_url: 'https://adb-2474129336842816.16.azuredatabricks.net'
config_template: {'run_name':'','new_cluster':{'spark_version':'9.1.x-scala2.12','node_type_id':'Standard_D3_v2','num_workers':2,'spark_conf':{}},'libraries':[{'jar':''}],'spark_jar_task':{'main_class_name':'','parameters':['']}}
work_dir: 'dbfs:/feathr_getting_started'
feathr_runtime_location: https://azurefeathrstorage.blob.core.windows.net/public/feathr-assembly-LATEST.jar
online_store:
redis:
host: 'feathrazuretest3redis.redis.cache.windows.net'
port: 6380
ssl_enabled: True
feature_registry:
purview:
type_system_initialization: true
purview_name: 'feathrazuretest3-purview1'
delimiter: '__'
"""
tmp = tempfile.NamedTemporaryFile(mode='w', delete=False)
with open(tmp.name, "w") as text_file:
text_file.write(yaml_config)
```
## Setup necessary environment variables (Skip if using the above Quick Start Template)
You should setup the environment variables in order to run this sample. More environment variables can be set by referring to [feathr_config.yaml](https://github.com/linkedin/feathr/blob/main/feathr_project/feathrcli/data/feathr_user_workspace/feathr_config.yaml) and use that as the source of truth. It also has more explanations on the meaning of each variable.
```
# os.environ['REDIS_PASSWORD'] = ''
# os.environ['AZURE_CLIENT_ID'] = ''
# os.environ['AZURE_TENANT_ID'] = ''
# os.environ['AZURE_CLIENT_SECRET'] = ''
# # Optional envs if you are using different runtimes
# os.environ['DATABRICKS_WORKSPACE_TOKEN_VALUE'] = ''
```
# Initialize Feathr Client
```
client = FeathrClient(config_path=tmp.name)
```
## View the data
In this tutorial, we use Feathr Feature Store to create a model that predicts NYC Taxi fares. The dataset comes from [here](https://www1.nyc.gov/site/tlc/about/tlc-trip-record-data.page). The data is as below
```
import pandas as pd
pd.read_csv("https://azurefeathrstorage.blob.core.windows.net/public/sample_data/green_tripdata_2020-04_with_index.csv")
```
## Defining Features with Feathr
In Feathr, a feature is viewed as a function, mapping from entity id or key, and timestamp to a feature value. For more details on feature definition, please refer to the [Feathr Feature Definition Guide](https://github.com/linkedin/feathr/blob/main/docs/concepts/feature-definition.md)
1. The typed key (a.k.a. entity id) identifies the subject of feature, e.g. a user id, 123.
2. The feature name is the aspect of the entity that the feature is indicating, e.g. the age of the user.
3. The feature value is the actual value of that aspect at a particular time, e.g. the value is 30 at year 2022.
Note that, in some cases, such as features defined on top of request data, may have no entity key or timestamp.
It is merely a function/transformation executing against request data at runtime.
For example, the day of week of the request, which is calculated by converting the request UNIX timestamp.
### Define Sources Section with UDFs
A feature source is needed for anchored features that describes the raw data in which the feature values are computed from. See the python documentation to get the details on each input column.
```
from pyspark.sql import SparkSession, DataFrame
def feathr_udf_day_calc(df: DataFrame) -> DataFrame:
from pyspark.sql.functions import dayofweek, dayofyear, col
df = df.withColumn("fare_amount_cents", col("fare_amount")*100)
return df
batch_source = HdfsSource(name="nycTaxiBatchSource",
path="wasbs://public@azurefeathrstorage.blob.core.windows.net/sample_data/green_tripdata_2020-04_with_index.csv",
event_timestamp_column="lpep_dropoff_datetime",
preprocessing=feathr_udf_day_calc,
timestamp_format="yyyy-MM-dd HH:mm:ss")
```
### Define Anchors and Features
A feature is called an anchored feature when the feature is directly extracted from the source data, rather than computed on top of other features. The latter case is called derived feature.
```
f_trip_distance = Feature(name="f_trip_distance",
feature_type=FLOAT, transform="trip_distance")
f_trip_time_duration = Feature(name="f_trip_time_duration",
feature_type=INT32,
transform="(to_unix_timestamp(lpep_dropoff_datetime) - to_unix_timestamp(lpep_pickup_datetime))/60")
features = [
f_trip_distance,
f_trip_time_duration,
Feature(name="f_is_long_trip_distance",
feature_type=BOOLEAN,
transform="cast_float(trip_distance)>30"),
Feature(name="f_day_of_week",
feature_type=INT32,
transform="dayofweek(lpep_dropoff_datetime)"),
]
request_anchor = FeatureAnchor(name="request_features",
source=INPUT_CONTEXT,
features=features)
```
### Window aggregation features
For window aggregation features, see the supported fields below:
Note that the `agg_func` should be any of these:
| Aggregation Type | Input Type | Description |
| --- | --- | --- |
|SUM, COUNT, MAX, MIN, AVG |Numeric|Applies the the numerical operation on the numeric inputs. |
|MAX_POOLING, MIN_POOLING, AVG_POOLING | Numeric Vector | Applies the max/min/avg operation on a per entry bassis for a given a collection of numbers.|
|LATEST| Any |Returns the latest not-null values from within the defined time window |
After you have defined features and sources, bring them together to build an anchor:
Note that if the data source is from the observation data, the `source` section should be `INPUT_CONTEXT` to indicate the source of those defined anchors.
```
location_id = TypedKey(key_column="DOLocationID",
key_column_type=ValueType.INT32,
description="location id in NYC",
full_name="nyc_taxi.location_id")
agg_features = [Feature(name="f_location_avg_fare",
key=location_id,
feature_type=FLOAT,
transform=WindowAggTransformation(agg_expr="cast_float(fare_amount)",
agg_func="AVG",
window="90d")),
Feature(name="f_location_max_fare",
key=location_id,
feature_type=FLOAT,
transform=WindowAggTransformation(agg_expr="cast_float(fare_amount)",
agg_func="MAX",
window="90d")),
Feature(name="f_location_total_fare_cents",
key=location_id,
feature_type=FLOAT,
transform=WindowAggTransformation(agg_expr="fare_amount_cents",
agg_func="SUM",
window="90d")),
]
agg_anchor = FeatureAnchor(name="aggregationFeatures",
source=batch_source,
features=agg_features)
```
### Derived Features Section
Derived features are the features that are computed from other features. They could be computed from anchored features, or other derived features.
```
f_trip_time_distance = DerivedFeature(name="f_trip_time_distance",
feature_type=FLOAT,
input_features=[
f_trip_distance, f_trip_time_duration],
transform="f_trip_distance * f_trip_time_duration")
f_trip_time_rounded = DerivedFeature(name="f_trip_time_rounded",
feature_type=INT32,
input_features=[f_trip_time_duration],
transform="f_trip_time_duration % 10")
```
And then we need to build those features so that it can be consumed later. Note that we have to build both the "anchor" and the "derived" features (which is not anchored to a source).
```
client.build_features(anchor_list=[agg_anchor, request_anchor], derived_feature_list=[
f_trip_time_distance, f_trip_time_rounded])
```
## Create training data using point-in-time correct feature join
A training dataset usually contains entity id columns, multiple feature columns, event timestamp column and label/target column.
To create a training dataset using Feathr, one needs to provide a feature join configuration file to specify
what features and how these features should be joined to the observation data.
To learn more on this topic, please refer to [Point-in-time Correctness](https://github.com/linkedin/feathr/blob/main/docs/concepts/point-in-time-join.md)
```
if client.spark_runtime == 'databricks':
output_path = 'dbfs:/feathrazure_test.avro'
else:
output_path = feathr_output_path
feature_query = FeatureQuery(
feature_list=["f_location_avg_fare", "f_trip_time_rounded", "f_is_long_trip_distance", "f_location_total_fare_cents"], key=location_id)
settings = ObservationSettings(
observation_path="wasbs://public@azurefeathrstorage.blob.core.windows.net/sample_data/green_tripdata_2020-04_with_index.csv",
event_timestamp_column="lpep_dropoff_datetime",
timestamp_format="yyyy-MM-dd HH:mm:ss")
client.get_offline_features(observation_settings=settings,
feature_query=feature_query,
output_path=output_path)
client.wait_job_to_finish(timeout_sec=500)
```
## Download the result and show the result
Let's use the helper function `get_result_df` to download the result and view it:
```
def get_result_df(client: FeathrClient) -> pd.DataFrame:
"""Download the job result dataset from cloud as a Pandas dataframe."""
res_url = client.get_job_result_uri(block=True, timeout_sec=600)
tmp_dir = tempfile.TemporaryDirectory()
client.feathr_spark_laucher.download_result(result_path=res_url, local_folder=tmp_dir.name)
dataframe_list = []
# assuming the result are in avro format
for file in glob.glob(os.path.join(tmp_dir.name, '*.avro')):
dataframe_list.append(pdx.read_avro(file))
vertical_concat_df = pd.concat(dataframe_list, axis=0)
tmp_dir.cleanup()
return vertical_concat_df
df_res = get_result_df(client)
df_res
```
## Train a machine learning model
After getting all the features, let's train a machine learning model with the converted feature by Feathr:
```
# remove columns
from sklearn.ensemble import GradientBoostingRegressor
final_df = df_res
final_df.drop(["lpep_pickup_datetime", "lpep_dropoff_datetime",
"store_and_fwd_flag"], axis=1, inplace=True, errors='ignore')
final_df.fillna(0, inplace=True)
final_df['fare_amount'] = final_df['fare_amount'].astype("float64")
train_x, test_x, train_y, test_y = train_test_split(final_df.drop(["fare_amount"], axis=1),
final_df["fare_amount"],
test_size=0.2,
random_state=42)
model = GradientBoostingRegressor()
model.fit(train_x, train_y)
y_predict = model.predict(test_x)
y_actual = test_y.values.flatten().tolist()
rmse = sqrt(mean_squared_error(y_actual, y_predict))
sum_actuals = sum_errors = 0
for actual_val, predict_val in zip(y_actual, y_predict):
abs_error = actual_val - predict_val
if abs_error < 0:
abs_error = abs_error * -1
sum_errors = sum_errors + abs_error
sum_actuals = sum_actuals + actual_val
mean_abs_percent_error = sum_errors / sum_actuals
print("Model MAPE:")
print(mean_abs_percent_error)
print()
print("Model Accuracy:")
print(1 - mean_abs_percent_error)
```
## Materialize feature value into offline/online storage
While Feathr can compute the feature value from the feature definition on-the-fly at request time, it can also pre-compute
and materialize the feature value to offline and/or online storage.
We can push the generated features to the online store like below:
```
backfill_time = BackfillTime(start=datetime(
2020, 5, 20), end=datetime(2020, 5, 20), step=timedelta(days=1))
redisSink = RedisSink(table_name="nycTaxiDemoFeature")
settings = MaterializationSettings("nycTaxiTable",
backfill_time=backfill_time,
sinks=[redisSink],
feature_names=["f_location_avg_fare", "f_location_max_fare"])
client.materialize_features(settings)
client.wait_job_to_finish(timeout_sec=500)
```
We can then get the features from the online store (Redis):
## Fetching feature value for online inference
For features that are already materialized by the previous step, their latest value can be queried via the client's
`get_online_features` or `multi_get_online_features` API.
```
res = client.get_online_features('nycTaxiDemoFeature', '265', [
'f_location_avg_fare', 'f_location_max_fare'])
client.multi_get_online_features("nycTaxiDemoFeature", ["239", "265"], [
'f_location_avg_fare', 'f_location_max_fare'])
```
### Registering and Fetching features
We can also register the features with an Apache Atlas compatible service, such as Azure Purview, and share the registered features across teams:
```
client.register_features()
client.list_registered_features(project_name="feathr_getting_started")
```
| github_jupyter |
# Clustering Tutorial
This guide will show how to use Tribuo’s clustering models to find clusters in a toy dataset drawn from a mixture of Gaussians. We'll look at Tribuo's K-Means implementation and also discuss how evaluation works for clustering tasks.
## Setup
We'll load in some jars and import a few packages.
```
%jars ./tribuo-clustering-kmeans-4.0.0-jar-with-dependencies.jar
import org.tribuo.*;
import org.tribuo.util.Util;
import org.tribuo.clustering.*;
import org.tribuo.clustering.evaluation.*;
import org.tribuo.clustering.example.ClusteringDataGenerator;
import org.tribuo.clustering.kmeans.*;
import org.tribuo.clustering.kmeans.KMeansTrainer.Distance;
var eval = new ClusteringEvaluator();
```
## Dataset
Tribuo's clustering package comes with a simple data generator that emits data sampled from a mixture of 5 2-dimensional Gaussians (the centroids and variances are fixed). This generator gives the ground truth cluster IDs, so it can be used for demos like this. You can also use any of the standard data loaders to pull in clustering data.
As it conforms to the standard `Trainer` and `Model` interface used for the rest of Tribuo, the training of a clustering algorithm doesn't produce cluster assignments that are visible, to recover the assignments we need to call `model.predict(trainData)`.
We're going to sample two datasets (using different seeds) one for fitting the cluster centroids, and one to measure clustering performance.
```
var data = ClusteringDataGenerator.gaussianClusters(500, 1L);
var test = ClusteringDataGenerator.gaussianClusters(500, 2L);
```
The data generator uses the following Gaussians:
1. `N([ 0.0,0.0], [[1.0,0.0],[0.0,1.0]])`
2. `N([ 5.0,5.0], [[1.0,0.0],[0.0,1.0]])`
3. `N([ 2.5,2.5], [[1.0,0.5],[0.5,1.0]])`
4. `N([10.0,0.0], [[0.1,0.0],[0.0,0.1]])`
5. `N([-1.0,0.0], [[1.0,0.0],[0.0,0.1]])`
## Model Training
We'll first fit a K-Means using 5 centroids, a maximum of 10 iterations, using the euclidean distance and a single computation thread.
```
var trainer = new KMeansTrainer(5,10,Distance.EUCLIDEAN,1,1);
var startTime = System.currentTimeMillis();
var model = trainer.train(data);
var endTime = System.currentTimeMillis();
System.out.println("Training with 5 clusters took " + Util.formatDuration(startTime,endTime));
```
We can inspect the centroids by querying the model.
```
var centroids = model.getCentroidVectors();
for (var centroid : centroids) {
System.out.println(centroid);
}
```
These centroids line up pretty well with the Gaussian centroids. The predicted ones line up with the true ones as follows:
|Predicted|True|
|---|---|
|1|5|
|2|3|
|3|1|
|4|2|
|5|4|
Though the first one is a bit far out as it's x_1 should be -1.0 not -1.7, and there is a little wobble in the rest. Still it's pretty good considering K-Means assumes spherical gaussians and our data generator has a covariance matrix per gaussian.
## Model evaluation
Tribuo uses the normalized mutual information to measure the quality
of two clusterings. This avoids the issue that swapping the id number of any given centroid doesn't change the overall clustering. We're going to compare against the ground truth cluster labels from the data generator.
First for the training data:
```
var trainEvaluation = eval.evaluate(model,data);
trainEvaluation.toString();
```
Then for the unseen test data:
```
var testEvaluation = eval.evaluate(model,test);
testEvaluation.toString();
```
We see that as expected it's a pretty good correlation to the ground truth labels. K-Means (of the kind implemented in Tribuo) is similar to a gaussian mixture using spherical gaussians, and our data generator uses gaussians with full rank covariances, so it won't be perfect.
## Multithreading
Tribuo's K-Means supports multi-threading of both the expectation and maximisation steps in the algorithm (i.e. the finding of the new centroids, and the assignment of points to centroids). We'll run the same experiment as before, both with 5 centroids and with 20 centroids, using 4 threads, though this time we'll use 2000 points for training.
```
var mtData = ClusteringDataGenerator.gaussianClusters(2000, 1L);
var mtTrainer = new KMeansTrainer(5,10,Distance.EUCLIDEAN,4,1);
var mtStartTime = System.currentTimeMillis();
var mtModel = mtTrainer.train(mtData);
var mtEndTime = System.currentTimeMillis();
System.out.println("Training with 5 clusters on 4 threads took " + Util.formatDuration(mtStartTime,mtEndTime));
```
Now with 20 centroids:
```
var overTrainer = new KMeansTrainer(20,10,Distance.EUCLIDEAN,4,1);
var overStartTime = System.currentTimeMillis();
var overModel = overTrainer.train(mtData);
var overEndTime = System.currentTimeMillis();
System.out.println("Training with 20 clusters on 4 threads took " + Util.formatDuration(overStartTime,overEndTime));
```
We can evaluate the two models as before, using our `ClusteringEvaluator`. First with 5 centroids:
```
var mtTestEvaluation = eval.evaluate(mtModel,test);
mtTestEvaluation.toString();
```
Then with 20:
```
var overTestEvaluation = eval.evaluate(overModel,test);
overTestEvaluation.toString();
```
We see that the multi-threaded versions run in less time than the single threaded trainer, despite them using 4 times the training data. The 20 centroid model has a tighter fit of the test data, despite being overparameterised. This is common in clustering tasks where it's hard to balance the model fitting with complexity. We'll look at adding more performance metrics so users can diagnose such issues in future releases.
## Conclusion
We looked at clustering using Tribuo's K-Means implementation, comparing both the single-threaded and multi-threaded versions, then looked at the performance metrics available when there are ground truth clusterings.
We plan to further expand Tribuo's clustering functionality to incorporate other algorithms in the future. If you want to help, or have specific algorithmic requirements, file an issue on our [github page](https://github.com/oracle/tribuo).
| github_jupyter |
# Data Extraction
### 1. Get the Dataset
##### Download the Zip File from the link Given below:
https://unearthed-exploresa.s3-ap-southeast-2.amazonaws.com/Unearthed_5_SARIG_Data_Package.zip
#### 2. Extract the files in a desired folder
Open this jupyter-notebook from that folder
```
# Importing the required libraries
import pandas as pd
import numpy as np
```
We'll be using the `sarig_rs_chem_exp.csv` file for preparing our dataset. Since the size of file is around ~11 GB, we'll specify the columns that we need for analysis. We'll also be specifying the low_memory parameter.
The columns we need are ->
- 'LONGITUDE_GDA94'
- 'LATITUDE_GDA94'
- 'CHEM_CODE'
- 'VALUE'
- 'UNIT'
If you wish to have a full look at the dataset, you can always remove the `usecols` parameter. Though, its highly likely that the kernel will crash. _Hack - You can specify nrows = 10 or 100 to preview the first few rows._
```
# Mention either the complete or relative path of the sarig_rs_chem_exp.csv file
path = '/home/xavian/Downloads/The_Gawler_Challenge/GeoChem_Data/Unearthed_5_SARIG_Data_Package/SARIG_Data_Package2_Exported20052020/sarig_rs_chem_exp.csv'
data = pd.read_csv(path, low_memory=False, encoding = "cp1252", usecols = ['LONGITUDE_GDA94','LATITUDE_GDA94','CHEM_CODE', 'VALUE', 'UNIT'])
# View the first few rows of the dataset
data.head()
# Let's view all the unique UNITS
data.UNIT.unique()
data.info()
# We'll be removing all the non-numeric characters -> '<>-'
data['VALUE'] = data.VALUE.str.lstrip('<->')
# We'll be dropping all the values with AMOUNT less than 10
data=data.drop(data[data['VALUE']=='0-10'].index)
# Convert the datatype to numeric
data[['VALUE']] = data[['VALUE']].apply(pd.to_numeric)
# Let's preview
data
# Create a new column - UNIT_PPM by converting all the other units to PPM
data['UNIT_PPM'] = np.where(data.UNIT == 'ppb', data.VALUE/1000, (np.where(data.UNIT == '%', data.VALUE*10000, data.VALUE)))
# Let's drop the UNIT & VALUE Column
data = data.drop(['UNIT', 'VALUE'], axis=1)
df=data.reset_index()
df
# These are the Chem codes we'll be dropping
drop_min=[ 'LOI', 'H2O_plus', 'H2O_minus', 'Insol', 'Total',
'GoI', 'TOT/C', 'pH', 'EC', 'RADBK',
'RADTC', 'HMIN', 'H2O','CPS_gamma']
# Let's drop them!
for i in drop_min:
print(i)
df=df.drop(df[ df['CHEM_CODE'] == i].index)
df.CHEM_CODE.unique()
df
```
Make sure you have `lxml` installed for the next step. It's not installed if you are in a conda environment. Install it using `pip install lxml`
```
# To select unique chem codes at every individual location, we'll be calculating VALUE = UNIT_PPM * PRICE per KG
link = "https://en.wikipedia.org/wiki/Prices_of_chemical_elements"
tables = pd.read_html(link,header=0)[0]
# create new column names
table = tables[['Symbol', 'Name', 'Price[5]']]
table=table.drop(index=0)
table=table.rename(columns={"Price[5]": "Price in USD/Kg"})
# View the table
table
# Lets make a dictionary with symbols as keys and price as Value
s=table['Symbol'].to_list()
p=table['Price in USD/Kg'].to_list()
price_dict = {s[i]: p[i] for i in range(len(s))}
# Now lets replace the price with a range, with a fixed value. For Ex
price = {'Ti': '11.7','K': '13.6','Na': '3.43', 'Ca': '2.35', 'Ru': '10600', 'Zr': '37.1', 'Ge': '1010', 'Ir': '56200', 'Sr': '6.68', 'Ta': '312', 'Re': '4150','Nb': '85.6','La': '4.92', 'Li': '85.6','Ce': '4.71','As': '1.31', 'Ba': '0.275','Ti': '11.7', 'V': '385','F': '2.16'}
price.update(price)
# Lets check the Dictionary
print(price_dict)
chemcode_list=df.CHEM_CODE.unique().tolist()
# Lets find the CHEM_CODE whose value we don't know[which are not in the 'price' dict]
def uncommon(lst1, lst2):
lst3 = [value for value in lst1 if value not in lst2]
return lst3
lst1 = s
lst2 = a
other=uncommon(lst1, lst2)
print(other)
# Now lets make a dict which will replace the 'other' CHEM_CODE with a specific element
element=['U', 'Si', 'Al', 'Ti', 'Fe', 'Mn', 'Mg', 'Ca', 'Na', 'K', 'P', 'Fe', ' ', 'Fe', 'Cr', 'V', 'Th', 'W', 'Ta', 'Nb', ' ', 'Na', 'Ba', 'Ca', ' ', 'Ca', 'Mg', 'Ca', ' ', '-', ' ', ' ', 'Fe', 'Fe', 'Ni', 'V', 'Zn', 'Sr', 'Cu', 'Zr', 'Hf', 'Sr', '-', 'K', 'Fe', 'Co', '-', 'Cl']
element_dict = {other[i]: element[i] for i in range(len(other))}
print(element_dict)
# Now lets make a new column so that we can make change without disturbing the original data
df[['CHEM_CODE_N']]=df[['CHEM_CODE']]
# Now Lets replace the 'other' CHEM_CODE using the 'element_dict'
df=df.replace({"CHEM_CODE_N": element_dict})
# Now lets drop some more dummy CHEM_CODE
# Doing it seperately reduces its asymptotic complexity
indexNames = df[ df['CHEM_CODE_N']==" " ].index
df=df.drop(indexNames)
indexNames = df[ df['CHEM_CODE_N']=="-" ].index
df=df.drop(indexNames)
# Lets check the changes
df.CHEM_CODE_N.unique()
# Now we will make find the Price of the chemcodes of column 'CHEM_CODE_N' and make a price column in dataframe 'df'
CHEM_CODE_N_price=[]
chemcode=df.CHEM_CODE_N.tolist()
for i in chemcode:
CHEM_CODE_price.append(float(price[i]))
df['price'] = np.array(CHEM_CODE_N_price)
# Now we will make a new column VALUE with approx values of chemcodes
df['VALUE'] = df['UNIT_PPM']*df['price']
# Now lets Drop the Unnesecary columns and reset its index
df=df.drop(columns=['price'])
df=df.reset_index()
df=df.drop(columns=['index'])
# Now lets find the most valuable minerals at a point and drop the other less valueable ones[present at the same point]
most_valued_minerals_at_points=df.sort_values('VALUE', ascending=False).drop_duplicates(['LONGITUDE_GDA94','LATITUDE_GDA94']).reset_index()
```
By our Preliminary Testing and analysis of Datasets we found that 'most_valued_minerals_at_points' had a lot of Ca and hence was bias and not good for training. Hence we should trim some Ca's
```
# Lets make a DataFrame with only Ca at different points
df_Ca=df.loc[df['CHEM_CODE']=='Ca'].drop_duplicates(['LONGITUDE_GDA94','LATITUDE_GDA94'])
# Now we will sort them in descending order and make a new DataFrame with the top 100000 rows
df_Ca=df_Ca.sort_values('VALUE', ascending=False)
df_Ca=df_Ca.iloc[:100000,:]
# Now we will make a index column by using reset_index() and using the index column to drop Ca from the main DataFrame
df_Ca=df_Ca.reset_index()
df=df.drop(df_Ca['index'].tolist())
# Now we will create a final DataFrame which will be used for training
final_df = df.sort_values('VALUE', ascending=False).drop_duplicates(['LONGITUDE_GDA94','LATITUDE_GDA94']).reset_index()
# Lets see how it looks
final_df
# Now we will classify UNIT_PPM into three classes
final_df.loc[final_df['UNIT_PPM']>50000, 'UNIT_PPM_CLASS']='HIGH'
final_df.loc[final_df['UNIT_PPM']<=50000, 'UNIT_PPM_CLASS']='MED'
final_df.loc[final_df['UNIT_PPM']<=100, 'UNIT_PPM_CLASS']='LOW'
# Lets see how it looks
final_df
# Finally lets Drop the unnescessary columns
final_df=final_df.drop(columns=['CHEM_CODE_N','index'])
# Now finally lets export the DataFrame as csv so that we can use for training
final_df.to_csv('unsampled.csv')
```
| github_jupyter |
# Init
```
# %load_ext autoreload
# %autoreload 2
from pprint import pprint
import numpy as np
import qcodes as qc
from labcore.measurement import *
```
# Intro
The way we would like to measure looks something like this:
```python
>>> m = Measurement(
... sweep_object,
... setup=setup_function,
... finish=teardown_function,
... )
... m.run(**config_options)
```
The reason why we would like to define the measurement in an object is to be able to have access to main characteristics (such as the type of data acquired during the run) in advance, before any measurement code is executed. This will also be useful for serializing measurements.
The most tricky aspect here is to define what the ``sweep`` object is. Often we define the flow in an experiment by directly using for-loops and the like. But that will not allow us the desired goals. We need to come up with an object-based description that makes the essence of the flow clear without executing it.
## Main requirements for sweeps
- a sweep object can be iterated over. each step performs some actions, and returns some data.
that means, we want to be able to use it as follows. We call the dictionary that is produced for each step a **record**.
```python
>>> for data in sweep_object:
... print(data)
{variable_1: some_data, variable_2: more_data}
{variable_1: different_data, variable_2: more_different_data}
```
- a sweep object is defined by a pointer and 0 or more actions.
- A pointer is again an iterable, and the coordinates it traverses may or may not be known in advance.
- Actions are callables that may take the values the pointer is returning at each iteration as arguments. Each action is executed once for each pointer.
A simple sweep can then be defined like this:
```python
>>> sweep_object = Sweep(range(5), func_1, func_2)
```
Executing this sweep will loop over the range, and will then call ``func_1(i)`` and subsequently ``func_2(i)``, for each ``i`` in the range from 0 to 4.
- Sweeps can be combined in ways that result in nesting, zipping, or appending. Combining sweeps again results in a sweep. This will allow us to construct modular measurements from pre-defined blocks.
- After definition, but before execution, we can easily infer what records the sweep produces.
# Examples
In the following first look a prototypical measurement protocol.
After that we have a closer look at individual components.
## Jumping right in
### A prototypical, slightly non-trivial measurement protocol
Lets say we have the following task:
1. our basic measurement is to record the phase as a function of frequency (for instance with a VNA)
2. we do this as function of some other parameter that tunes something in the experiment (maybe a voltage)
3. finally, we need to average out some slow noise, so we repeat everything some number of times
4. the tuning might heat up the sample, so we want to keep an eye on it. We want to measure the fridge temperature once per repetition.
Being good with organizing our tools and prefering a modular way of putting together measurements, we have already defined the main building blocks so we can re-use them. They might even be located in some module that import for a particular class of experiments.
The functions that return data we want to save later are decorated with ``recording``, to describe what kind of data they produce. (Note: the decorator is not necessary; we can also use the function ``record_as``, as we'll see below)
```
# these are just some dummy functions that follow the recipe outlined above.
def tune_something(param):
print(f'adjusting something in-situ using param={param}')
@recording(
independent('frq', type='array', unit='Hz'),
dependent('phase', depends_on=['frq'], type='array', unit='rad')
)
def measure_frequency_trace(npoints=3):
return np.arange(1,npoints+1)*1e6, np.random.normal(size=npoints)
@recording(dependent('temperature', unit='K'))
def measure_temperature():
return np.random.normal(loc=0.1, scale=0.01)
```
We now use these functions to assemble the measurement loop. It works like this:
- The matrix multiplication operator ``@`` creates a nested loop of sweeps (the outer loop are the repetitions, the inner loop the tuning of the parameter).
- The multiplication operator ``*`` is like an inner product (or zip), i.e., results in element-wise combination. That is, for each value of ``param`` we run ``measure_frequency_trace``.
- The addition operator ``+`` appends sweeps or actions to sweeps. I.e., ``measure_temperature`` is executed once after each ``param`` sweep.
Finally, we can, even without running any measurement code, already look at the data that the loop will produce.
```
sweep = (
sweep_parameter('repetition', range(3))
@ (sweep_parameter('param', range(3), tune_something)
* measure_frequency_trace
+ measure_temperature)
)
pprint(sweep.get_data_specs())
print()
```
Running the sweep is very easy -- we can simply iterate over it. Each sweep coordinate produces a data set that by default contains everything that has been annotated as recording data (the ``sweep_parameter`` function implicitly generates a record for the parameter that is being varied).
```
for data in sweep:
print(data)
```
### A QCoDeS parameter sweep
Sweeps are often done over qcodes parameters, and the data we acquire may also come from parameters.
In this minimal example we set a parameter (``x``) to a range of values, and get data from another parameter for each set value.
```
from qcodes import Parameter
def measure_stuff():
return np.random.normal()
x = Parameter('x', set_cmd=lambda x: print(f'setting x to {x}'), initial_value=0)
data = Parameter('data', get_cmd=lambda: np.random.normal())
for record in sweep_parameter(x, range(3), get_parameter(data)):
print(record)
```
## Constructing single sweeps
The main ingredients for a single sweep is 1) a pointer iterable, and 2) a variable number of actions to execute at each iteration step.
Both pointer and actions may generate records.
The most bare example would look something like this:
```
for data in Sweep(range(3)):
print(data)
```
We loop over the iterable, but only empty records are generated.
We can use ``record_as`` to indicate that generated values should be recorded:
```
def my_func():
return 0
sweep = Sweep(
record_as(range(3), independent('x')), # this is the pointer. We specify 'x' as an independent (we control it)
record_as(my_func, 'y') # y is not declared as independent;
# dependent (on what it depends is partially determined by the sweep) is the default.
)
pprint(sweep.get_data_specs())
for data in sweep:
print(data)
```
A more convenient way of doing exactly the same thing:
```
sweep = sweep_parameter('x', range(3), record_as(my_func, 'y'))
for data in sweep:
print(data)
```
Elements can also produce records with multiple parameters:
```
def my_func():
return 1, 2
sweep = Sweep(
record_as(zip(range(3), ['a', 'b', 'c']), independent('number'), independent('string')), # a pointer with two parameters
record_as(my_func, 'one', 'two')
)
pprint(sweep.get_data_specs())
for data in sweep:
print(data)
```
## Specifying options before executing a sweep
Many functions we are using take optional parameters we only want to specify just before executing the sweep (but are constant throughout the sweep).
If we don't want to resort to global variables we can do so by using ``Sweep.set_action_opts``.
It accepts the names of action functions as keywords, and dictionaries containing keyword arguments to pass to those functions as value.
Keywords specified in this way always override key words that are passed around internally in the sweep (See further below for an explanation on how parameters are passed internally).
```
def test_fun(value, a_property=0):
print('inside test_fun:')
print(f"value: {value}, property: {a_property}")
print()
return value
def another_fun(another_value, *args, **kwargs):
print('inside another_fun:')
print(f"my value: {another_value}")
print(f"other stuff:", args, kwargs)
print()
sweep_1 = sweep_parameter('value', range(3), record_as(test_fun, dependent('data')))
sweep_2 = sweep_parameter('another_value', range(2), another_fun)
sweep = (
sweep_1
@ sweep_2
)
sweep.set_action_opts(
test_fun = dict(a_property=1),
another_fun = dict(another_value='Hello', another_property=True)
)
for data in sweep:
print("Data:", data)
print()
```
## Appending
A simple example for how to append sweeps to each other.
The `sweep_parameter` function creates a 1D sweep over a parameter that we can specify just with a name. It will automatically result in a returned record with that name when it is set.
We create two sweeps that consist of a parameter sweep and an action that returns a random number as dependent variable.
The sweep parameter will automatically be inserted as a dependency in that case.
Here, the annotation of the return data is done using ``record_as``. This allows us to use the same function twice with different return data names.
We finally attach another function call to the sweep. It is executed only once at the very end (internally it's made into a 'null sweep'). It is also not annotated at all, so won't record any data.
Note: ``Sweep.return_none`` controls whether we include data fields that have returned nothing during setting a pointer or executing an action. It can be set on the class or the instance of a particular sweep.
Setting it to true (the default) guarantees that each data spec of the sweep has an entry per sweep point, even if it is ``None``.
```
def get_random_number():
return np.random.rand()
def get_another_random_number():
print('rolling the dice!')
return np.random.rand()
# change this to see what happens.
Sweep.record_none = False
sweep_1 = sweep_parameter('x', range(3), record_as(get_random_number, dependent('y')))
sweep_2 = sweep_parameter('a', range(4), record_as(get_random_number, dependent('b')))
for data in sweep_1 + sweep_2 + get_another_random_number:
print(data)
# just to set it back for the other examples
Sweep.record_none = True
```
## Multiplying
By multiplying we refer to an inner product, i.e., the result is basically what you'd expect from ``zip``-ing two iterables.
Simplest case: we have a sweep and want to attach another action to each sweep point:
```
sweep = (
sweep_parameter('x', range(3), record_as(get_random_number, dependent('data_1')))
* record_as(get_random_number, dependent('data_2'))
)
pprint(sweep.get_data_specs())
print()
for data in sweep:
print(data)
```
If the two objects we want to combine are both sweeps, then we get zip-like behavior (and dependencies stay separate).
```
sweep = (
sweep_parameter('x', range(3), record_as(get_random_number, dependent('data_1')))
* sweep_parameter('y', range(5), record_as(get_random_number, dependent('data_2')))
)
pprint(sweep.get_data_specs())
print()
for data in sweep:
print(data)
```
## Nesting sweeps
The most simple example:
Sweep parameters (say, 'x', 'y', and 'z') against each other, and perform a measurement at each point.
For syntactic brevity we're using the matrix multiplication ('@') operator for nesting (you can also use ``labcore.measurement.sweep.nest_sweeps``).
```
@recording(dependent('my_data'))
def measure_something():
return np.random.rand()
my_sweep = (
sweep_parameter('x', range(3))
@ sweep_parameter('y', np.linspace(0,1,3))
@ sweep_parameter('z', range(-5, -2))
@ measure_something
)
for data in my_sweep:
print(data)
```
The outer loops can be more complex sweeps as well -- for example, we can also execute measurements on each nesting level.
```
def measure_something():
return np.random.rand()
sweep_1 = sweep_parameter('x', range(3), record_as(measure_something, 'a'))
sweep_2 = sweep_parameter('y', range(4), record_as(measure_something, 'b'))
for data in sweep_1 @ sweep_2 @ record_as(get_random_number, 'more_data'):
print(data)
```
## Combining operators and more complex constructs
A not uncommon task in quantum circuits:
1. sweep some parameter. We're interested in some response of the system on this parameter.
2. Hovewer, changing this parameter changes an operation point of our sample. To deal with this we need to perform an auxiliary measurement, and based on its result we set some other parameter. (A familiar case: we increase a drive power that is expected to have some effect on a qubit. But after changing the power we need to first find the readout resonator frequency because of the changed Stark shift.)
3. After this 'calibration step' we can now measure the response we're actually looking for.
Further, we want to record not only the data that is taken in the last step, but also the calibration data.
All together this leads us to a measurement structure as below. As before, the building blocks are modular and relatively self-contained, and might be part of some package.
The data this will produce is two records per sweep parameter (``x``, creatively): one that contains the calibration measurement and result, and one for the actual measurement.
```
@recording(
independent('readout_probe_points', type='array'),
dependent('readout_probe_response', type='array', depends_on=['readout_probe_points']),
dependent('readout_point') # note: we don't make this dependent on readout_probe_points
)
def calibrate_readout():
"""An example for a readout calibration."""
pts = np.linspace(0, 1, 3)
response = np.random.normal(size=3, scale=10)
result = response.max() # some analysis of the calibration measurement
return pts, response, result
def set_operation_point(value):
print(f'setting the readout point to {value}')
# the actual measurement function accepts the argument `readout_point` which will be passed from the calibration.
@recording(
independent('probe_frequency', type='array'),
dependent('probe_response', type='array', depends_on=['probe_frequency']),
)
def perform_measurement(readout_point):
set_operation_point(readout_point)
return np.arange(3)+10, np.random.normal(size=3, loc=readout_point)
# because of the way brackets are set here, we need to make a sweep out of the calibrate_readout
# function. this is easily done with the 'once' function, which creates a length-1 sweep with no additional
# return data.
sweep = (
sweep_parameter('x', range(3))
@ (once(calibrate_readout) + perform_measurement)
)
for data in sweep:
pprint(data)
pprint(sweep.get_data_specs())
```
# Looking a bit deeper
## Annotating objects for recording
These are just some snippets that show how we can annotate records and what the results are.
Without record annotations sweeps still are executed, but no data is recorded:
```
sweep_object = Sweep(range(3), lambda x: np.random.normal(size=x))
for data in sweep_object:
print(data)
```
The underlying object we use for declaring data returns is ``DataSpec``. ``ds`` is just a shortcut that points to its constructor.
``independent`` (or ``indep``) is creates a DataSpec with ``depends_on=None``, and ``dependent`` creates a DataSpec with ``depends_on=[]``. If ``depends_on`` is ``None``, it will remain independent even when the annotated object is embedded in larger structures. If it is `[]` then more dependencies are added automatically.
```
# defining some example measurement functions without short-hand notations
@recording(DataSpec('x'), DataSpec('y', depends_on=['x'], type='array'))
def measure_stuff(n, *args, **kwargs):
return n, np.random.normal(size=n)
@recording(ds('a'))
def set_stuff(x, *args, **kwargs):
return x
measure_stuff(1), set_stuff(1)
```
Generators can also be annotated to produce data.
```
@recording(ds('a'))
def make_sequence(n):
for i in range(n):
yield i
for data in make_sequence(3):
print(data)
```
Using ``record_as`` is a practical way of annotating records just before executing,
independently of an earlier function definition.
This works with functions and generators as well.
```
def get_some_data(n):
return np.random.normal(size=n)
record_as(get_some_data, ds('random_var'))(3)
```
record also provides a simple short hand for labelling regular iterables and iterators:
it automatically removes surplus values, and always returns values for all specified data,
even if the annotated object does not return in (missing values will be ``None``)
```
for data in record_as(zip(np.linspace(0,1,6), np.arange(6)), ds('x')):
print(data)
for data in record_as(zip(np.linspace(0,1,6), np.arange(6)), ds('x'), ds('y')):
print(data)
for data in record_as(np.linspace(0,1,6), ds('x'), ds('y')):
print(data)
```
## Passing parameters in a sweep
Everything that is generated by functions (or pointers and sweeps) can in principle be passed on to subsequently executed elements.
When there's no record annotations arguments may still be passed as positional arguments:
```
def test(*args, **kwargs):
print(args, kwargs)
return 100
for data in Sweep(range(3), test):
print(data)
```
Because it would get too confusing otherwise, positional arguments only get passed originating from a pointer to all actions in a single sweep. In the following example the two ``test`` functions in the first sweep are passed the same integer, whereas the function in the second sweep is only receiving (``x``, ``True``) or (``y``, ``False``). Note that the return of ``test`` is not passed to any other object.
```
for data in Sweep(range(3), test, test) * Sweep(zip(['x', 'y'], [True, False]), test):
print(data)
```
In the previous example, ``test`` received any arguments passed to it because its signature included variational positional arguments (``*args``). The situation changes when this is not the case. The function is receiving only arguments that it can accept.
```
def test_2(x=2):
print(x)
return True
for data in Sweep(zip([1,2], [3,4]), test_2):
pass
```
We are more flexible when we use keyword arguments. Here, the rule is:
1. All records produced are passed to all subsequent functions in the sweep (even across different sub-sweeps!) that accept the keyword.
2. If a pointer yields non-annotated values, these are still used as positional arguments, but only where accepted, and with higher priority given to keywords.
3. using ``lambda`` and ``record_as`` allow pretty simple translation of records and argument names to avoid conflicts.
4. some elementary control over passing behavior is provided by ``Sweep.pass_on_returns`` and ``Sweep.pass_on_none``.
1. ``Sweep.pass_on_returns`` defaults to ``True``. If set to ``False``, nothing will be passed on.
2. ``Sweep.pass_on_none`` defaults to ``False``. If set to ``False`` the behavior is such that records that are ``None`` will not be passed on further. (Because ``None`` is typically indicating that function did not return anything as data even though a record was declard using ``recording`` or ``record_as``).
3. Note: At the moment we can set those only globally using the class attribute; a likely update in the future.
Some examples:
```
def test(x, y, z=5):
print("my three arguments:", x, y, z)
return x, y, z
def print_all_args(*args, **kwargs):
print("arguments at the end of the line:", args, kwargs)
for data in sweep_parameter('x', range(3), test):
print("data:", data)
print()
sweep = (
sweep_parameter('y', range(3), record_as(test, dependent('xx'), dependent('yy'), dependent('zz')))
@ print_all_args
)
for data in sweep:
print("data:", data)
print()
sweep = (
sweep_parameter('y', range(3), record_as(test, dependent('xx'), dependent('yy'), dependent('zz')))
@ record_as(lambda xx, yy, zz: test(xx, yy, zz), dependent('some'), dependent('different'), dependent('names'))
@ print_all_args
+ print_all_args
)
for data in sweep:
print("data:", data)
Sweep.pass_on_returns = False
sweep = (
sweep_parameter('y', range(3), record_as(test, dependent('xx'), dependent('yy'), dependent('zz')))
@ print_all_args
)
for data in sweep:
print("data:", data)
# setting back to default
Sweep.pass_on_returns = True
Sweep.pass_on_none = True
sweep = (
sweep_parameter('y', range(3), record_as(test, dependent('xx'), dependent('yy'), dependent('zz')))
@ print_all_args
)
for data in sweep:
print("data:", data)
# setting back to default
Sweep.pass_on_none = False
```
| github_jupyter |
# Interpreting nodes and edges with saliency maps in GCN (sparse)
<table><tr><td>Run the latest release of this notebook:</td><td><a href="https://mybinder.org/v2/gh/stellargraph/stellargraph/master?urlpath=lab/tree/demos/interpretability/gcn-sparse-node-link-importance.ipynb" alt="Open In Binder" target="_parent"><img src="https://mybinder.org/badge_logo.svg"/></a></td><td><a href="https://colab.research.google.com/github/stellargraph/stellargraph/blob/master/demos/interpretability/gcn-sparse-node-link-importance.ipynb" alt="Open In Colab" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg"/></a></td></tr></table>
This demo shows how to use integrated gradients in graph convolutional networks to obtain accurate importance estimations for both the nodes and edges. The notebook consists of three parts:
- setting up the node classification problem for Cora citation network
- training and evaluating a GCN model for node classification
- calculating node and edge importances for model's predictions of query ("target") nodes
<a name="refs"></a>
**References**
[1] Axiomatic Attribution for Deep Networks. M. Sundararajan, A. Taly, and Q. Yan.
Proceedings of the 34th International Conference on Machine Learning, Sydney, Australia, PMLR 70, 2017
([link](https://arxiv.org/pdf/1703.01365.pdf)).
[2] Adversarial Examples on Graph Data: Deep Insights into Attack and Defense. H. Wu, C. Wang, Y. Tyshetskiy, A. Docherty, K. Lu, and L. Zhu. arXiv: 1903.01610 ([link](https://arxiv.org/abs/1903.01610)).
```
# install StellarGraph if running on Google Colab
import sys
if 'google.colab' in sys.modules:
%pip install -q stellargraph[demos]==1.3.0b
# verify that we're using the correct version of StellarGraph for this notebook
import stellargraph as sg
try:
sg.utils.validate_notebook_version("1.3.0b")
except AttributeError:
raise ValueError(
f"This notebook requires StellarGraph version 1.3.0b, but a different version {sg.__version__} is installed. Please see <https://github.com/stellargraph/stellargraph/issues/1172>."
) from None
import networkx as nx
import pandas as pd
import numpy as np
from scipy import stats
import os
import time
import stellargraph as sg
from stellargraph.mapper import FullBatchNodeGenerator
from stellargraph.layer import GCN
from tensorflow import keras
from tensorflow.keras import layers, optimizers, losses, metrics, Model, regularizers
from sklearn import preprocessing, feature_extraction, model_selection
from copy import deepcopy
import matplotlib.pyplot as plt
from stellargraph import datasets
from IPython.display import display, HTML
%matplotlib inline
```
## Loading the CORA network
(See [the "Loading from Pandas" demo](../basics/loading-pandas.ipynb) for details on how data can be loaded.)
```
dataset = datasets.Cora()
display(HTML(dataset.description))
G, subjects = dataset.load()
```
### Splitting the data
For machine learning we want to take a subset of the nodes for training, and use the rest for validation and testing. We'll use scikit-learn again to do this.
Here we're taking 140 node labels for training, 500 for validation, and the rest for testing.
```
train_subjects, test_subjects = model_selection.train_test_split(
subjects, train_size=140, test_size=None, stratify=subjects
)
val_subjects, test_subjects = model_selection.train_test_split(
test_subjects, train_size=500, test_size=None, stratify=test_subjects
)
```
### Converting to numeric arrays
For our categorical target, we will use one-hot vectors that will be fed into a soft-max Keras layer during training. To do this conversion ...
```
target_encoding = preprocessing.LabelBinarizer()
train_targets = target_encoding.fit_transform(train_subjects)
val_targets = target_encoding.transform(val_subjects)
test_targets = target_encoding.transform(test_subjects)
all_targets = target_encoding.transform(subjects)
```
### Creating the GCN model in Keras
To feed data from the graph to the Keras model we need a generator. Since GCN is a full-batch model, we use the `FullBatchNodeGenerator` class.
```
generator = FullBatchNodeGenerator(G, sparse=True)
```
For training we map only the training nodes returned from our splitter and the target values.
```
train_gen = generator.flow(train_subjects.index, train_targets)
```
Now we can specify our machine learning model: tn this example we use two GCN layers with 16-dimensional hidden node features at each layer with ELU activation functions.
```
layer_sizes = [16, 16]
gcn = GCN(
layer_sizes=layer_sizes,
activations=["elu", "elu"],
generator=generator,
dropout=0.3,
kernel_regularizer=regularizers.l2(5e-4),
)
# Expose the input and output tensors of the GCN model for node prediction, via GCN.in_out_tensors() method:
x_inp, x_out = gcn.in_out_tensors()
# Snap the final estimator layer to x_out
x_out = layers.Dense(units=train_targets.shape[1], activation="softmax")(x_out)
```
### Training the model
Now let's create the actual Keras model with the input tensors `x_inp` and output tensors being the predictions `x_out` from the final dense layer
```
model = keras.Model(inputs=x_inp, outputs=x_out)
model.compile(
optimizer=optimizers.Adam(lr=0.01), # decay=0.001),
loss=losses.categorical_crossentropy,
metrics=[metrics.categorical_accuracy],
)
```
Train the model, keeping track of its loss and accuracy on the training set, and its generalisation performance on the validation set (we need to create another generator over the validation data for this)
```
val_gen = generator.flow(val_subjects.index, val_targets)
```
Train the model
```
history = model.fit(
train_gen, shuffle=False, epochs=20, verbose=2, validation_data=val_gen
)
sg.utils.plot_history(history)
```
Evaluate the trained model on the test set
```
test_gen = generator.flow(test_subjects.index, test_targets)
test_metrics = model.evaluate(test_gen)
print("\nTest Set Metrics:")
for name, val in zip(model.metrics_names, test_metrics):
print("\t{}: {:0.4f}".format(name, val))
```
## Node and link importance via saliency maps
In order to understand why a selected node is predicted as a certain class we want to find the node feature importance, total node importance, and link importance for nodes and edges in the selected node's neighbourhood (ego-net). These importances give information about the effect of changes in the node's features and its neighbourhood on the prediction of the node, specifically:
- **Node feature importance**: Given the selected node $t$ and the model's prediction $s(c)$ for class $c$. The feature importance can be calculated for each node $v$ in the selected node's ego-net where the importance of feature $f$ for node $v$ is the change predicted score $s(c)$ for the selected node when the feature $f$ of node $v$ is perturbed.
- **Total node importance**: This is defined as the sum of the feature importances for node $v$ for all features. Nodes with high importance (positive or negative) affect the prediction for the selected node more than links with low importance.
- **Link importance**: This is defined as the change in the selected node's predicted score $s(c)$ if the link $e=(u, v)$ is removed from the graph. Links with high importance (positive or negative) affect the prediction for the selected node more than links with low importance.
Node and link importances can be used to assess the role of nodes and links in model's predictions for the node(s) of interest (the selected node). For datasets like CORA-ML, the features and edges are binary, vanilla gradients may not perform well so we use integrated gradients [[1]](#refs) to compute them.
Another interesting application of node and link importances is to identify model vulnerabilities to attacks via perturbing node features and graph structure (see [[2]](#refs)).
To investigate these importances we use the StellarGraph `saliency_maps` routines:
```
from stellargraph.interpretability.saliency_maps import IntegratedGradients
```
Select the target node whose prediction is to be interpreted
```
graph_nodes = list(G.nodes())
target_nid = 1109199
target_idx = graph_nodes.index(target_nid)
y_true = all_targets[target_idx] # true class of the target node
all_gen = generator.flow(graph_nodes)
y_pred = model.predict(all_gen)[0, target_idx]
class_of_interest = np.argmax(y_pred)
print(
"Selected node id: {}, \nTrue label: {}, \nPredicted scores: {}".format(
target_nid, y_true, y_pred.round(2)
)
)
```
Get the node feature importance by using integrated gradients
```
int_grad_saliency = IntegratedGradients(model, train_gen)
```
For the parameters of `get_node_importance` method, `X` and `A` are the feature and adjacency matrices, respectively. If `sparse` option is enabled, `A` will be the non-zero values of the adjacency matrix with `A_index` being the indices. `target_idx` is the node of interest, and `class_of_interest` is set as the predicted label of the node. `steps` indicates the number of steps used to approximate the integration in integrated gradients calculation. A larger value of `steps` gives better approximation, at the cost of higher computational overhead.
```
integrated_node_importance = int_grad_saliency.get_node_importance(
target_idx, class_of_interest, steps=50
)
integrated_node_importance.shape
print("\nintegrated_node_importance", integrated_node_importance.round(2))
print("integrate_node_importance.shape = {}".format(integrated_node_importance.shape))
print(
"integrated self-importance of target node {}: {}".format(
target_nid, integrated_node_importance[target_idx].round(2)
)
)
```
Check that number of non-zero node importance values is less or equal the number of nodes in target node's K-hop ego net (where K is the number of GCN layers in the model)
```
G_ego = nx.ego_graph(G.to_networkx(), target_nid, radius=len(gcn.activations))
print("Number of nodes in the ego graph: {}".format(len(G_ego.nodes())))
print(
"Number of non-zero elements in integrated_node_importance: {}".format(
np.count_nonzero(integrated_node_importance)
)
)
```
We now compute the link importance using integrated gradients [[1]](#refs). Integrated gradients are obtained by accumulating the gradients along the path between the baseline (all-zero graph) and the state of the graph. They provide better sensitivity for the graphs with binary features and edges compared with the vanilla gradients.
```
integrate_link_importance = int_grad_saliency.get_integrated_link_masks(
target_idx, class_of_interest, steps=50
)
integrate_link_importance_dense = np.array(integrate_link_importance.todense())
print("integrate_link_importance.shape = {}".format(integrate_link_importance.shape))
print(
"Number of non-zero elements in integrate_link_importance: {}".format(
np.count_nonzero(integrate_link_importance.todense())
)
)
```
We can now find the nodes that have the highest importance to the prediction of the selected node:
```
sorted_indices = np.argsort(integrate_link_importance_dense.flatten())
N = len(graph_nodes)
integrated_link_importance_rank = [(k // N, k % N) for k in sorted_indices[::-1]]
topk = 10
# integrate_link_importance = integrate_link_importance_dense
print(
"Top {} most important links by integrated gradients are:\n {}".format(
topk, integrated_link_importance_rank[:topk]
)
)
# Set the labels as an attribute for the nodes in the graph. The labels are used to color the nodes in different classes.
nx.set_node_attributes(G_ego, values={x[0]: {"subject": x[1]} for x in subjects.items()})
```
In the following, we plot the link and node importance (computed by integrated gradients) of the nodes within the ego graph of the target node.
For nodes, the shape of the node indicates the positive/negative importance the node has. 'round' nodes have positive importance while 'diamond' nodes have negative importance. The size of the node indicates the value of the importance, e.g., a large diamond node has higher negative importance.
For links, the color of the link indicates the positive/negative importance the link has. 'red' links have positive importance while 'blue' links have negative importance. The width of the link indicates the value of the importance, e.g., a thicker blue link has higher negative importance.
```
integrated_node_importance.max()
integrate_link_importance.max()
node_size_factor = 1e2
link_width_factor = 2
nodes = list(G_ego.nodes())
colors = pd.DataFrame(
[v[1]["subject"] for v in G_ego.nodes(data=True)], index=nodes, columns=["subject"]
)
colors = np.argmax(target_encoding.transform(colors), axis=1) + 1
fig, ax = plt.subplots(1, 1, figsize=(15, 10))
pos = nx.spring_layout(G_ego)
# Draw ego as large and red
node_sizes = [integrated_node_importance[graph_nodes.index(k)] for k in nodes]
node_shapes = ["o" if w > 0 else "d" for w in node_sizes]
positive_colors, negative_colors = [], []
positive_node_sizes, negative_node_sizes = [], []
positive_nodes, negative_nodes = [], []
node_size_scale = node_size_factor / np.max(node_sizes)
for k in range(len(nodes)):
if nodes[k] == target_idx:
continue
if node_shapes[k] == "o":
positive_colors.append(colors[k])
positive_nodes.append(nodes[k])
positive_node_sizes.append(node_size_scale * node_sizes[k])
else:
negative_colors.append(colors[k])
negative_nodes.append(nodes[k])
negative_node_sizes.append(node_size_scale * abs(node_sizes[k]))
# Plot the ego network with the node importances
cmap = plt.get_cmap("jet", np.max(colors) - np.min(colors) + 1)
nc = nx.draw_networkx_nodes(
G_ego,
pos,
nodelist=positive_nodes,
node_color=positive_colors,
cmap=cmap,
node_size=positive_node_sizes,
vmin=np.min(colors) - 0.5,
vmax=np.max(colors) + 0.5,
node_shape="o",
)
nc = nx.draw_networkx_nodes(
G_ego,
pos,
nodelist=negative_nodes,
node_color=negative_colors,
cmap=cmap,
node_size=negative_node_sizes,
vmin=np.min(colors) - 0.5,
vmax=np.max(colors) + 0.5,
node_shape="d",
)
# Draw the target node as a large star colored by its true subject
nx.draw_networkx_nodes(
G_ego,
pos,
nodelist=[target_nid],
node_size=50 * abs(node_sizes[nodes.index(target_nid)]),
node_shape="*",
node_color=[colors[nodes.index(target_nid)]],
cmap=cmap,
vmin=np.min(colors) - 0.5,
vmax=np.max(colors) + 0.5,
label="Target",
)
# Draw the edges with the edge importances
edges = G_ego.edges()
weights = [
integrate_link_importance[graph_nodes.index(u), graph_nodes.index(v)]
for u, v in edges
]
edge_colors = ["red" if w > 0 else "blue" for w in weights]
weights = link_width_factor * np.abs(weights) / np.max(weights)
ec = nx.draw_networkx_edges(G_ego, pos, edge_color=edge_colors, width=weights)
plt.legend()
plt.colorbar(nc, ticks=np.arange(np.min(colors), np.max(colors) + 1))
plt.axis("off")
plt.show()
```
We then remove the node or edge in the ego graph one by one and check how the prediction changes. By doing so, we can obtain the ground truth importance of the nodes and edges. Comparing the following figure and the above one can show the effectiveness of integrated gradients as the importance approximations are relatively consistent with the ground truth.
```
(X, _, A_index, A), _ = train_gen[0]
X_bk = deepcopy(X)
A_bk = deepcopy(A)
selected_nodes = np.array([[target_idx]], dtype="int32")
nodes = [graph_nodes.index(v) for v in G_ego.nodes()]
edges = [(graph_nodes.index(u), graph_nodes.index(v)) for u, v in G_ego.edges()]
clean_prediction = model.predict([X, selected_nodes, A_index, A]).squeeze()
predict_label = np.argmax(clean_prediction)
groud_truth_node_importance = np.zeros((N,))
for node in nodes:
# we set all the features of the node to zero to check the ground truth node importance.
X_perturb = deepcopy(X_bk)
X_perturb[:, node, :] = 0
predict_after_perturb = model.predict(
[X_perturb, selected_nodes, A_index, A]
).squeeze()
groud_truth_node_importance[node] = (
clean_prediction[predict_label] - predict_after_perturb[predict_label]
)
node_shapes = [
"o" if groud_truth_node_importance[k] > 0 else "d" for k in range(len(nodes))
]
positive_colors, negative_colors = [], []
positive_node_sizes, negative_node_sizes = [], []
positive_nodes, negative_nodes = [], []
# node_size_scale is used for better visulization of nodes
node_size_scale = node_size_factor / max(groud_truth_node_importance)
for k in range(len(node_shapes)):
if nodes[k] == target_idx:
continue
if node_shapes[k] == "o":
positive_colors.append(colors[k])
positive_nodes.append(graph_nodes[nodes[k]])
positive_node_sizes.append(
node_size_scale * groud_truth_node_importance[nodes[k]]
)
else:
negative_colors.append(colors[k])
negative_nodes.append(graph_nodes[nodes[k]])
negative_node_sizes.append(
node_size_scale * abs(groud_truth_node_importance[nodes[k]])
)
X = deepcopy(X_bk)
groud_truth_edge_importance = np.zeros((N, N))
G_edge_indices = [(A_index[0, k, 0], A_index[0, k, 1]) for k in range(A.shape[1])]
for edge in edges:
edge_index = G_edge_indices.index((edge[0], edge[1]))
origin_val = A[0, edge_index]
A[0, edge_index] = 0
# we set the weight of a given edge to zero to check the ground truth link importance
predict_after_perturb = model.predict([X, selected_nodes, A_index, A]).squeeze()
groud_truth_edge_importance[edge[0], edge[1]] = (
predict_after_perturb[predict_label] - clean_prediction[predict_label]
) / (0 - 1)
A[0, edge_index] = origin_val
fig, ax = plt.subplots(1, 1, figsize=(15, 10))
cmap = plt.get_cmap("jet", np.max(colors) - np.min(colors) + 1)
# Draw the target node as a large star colored by its true subject
nx.draw_networkx_nodes(
G_ego,
pos,
nodelist=[target_nid],
node_size=50 * abs(node_sizes[nodes.index(target_idx)]),
node_color=[colors[nodes.index(target_idx)]],
cmap=cmap,
node_shape="*",
vmin=np.min(colors) - 0.5,
vmax=np.max(colors) + 0.5,
label="Target",
)
# Draw the ego net
nc = nx.draw_networkx_nodes(
G_ego,
pos,
nodelist=positive_nodes,
node_color=positive_colors,
cmap=cmap,
node_size=positive_node_sizes,
vmin=np.min(colors) - 0.5,
vmax=np.max(colors) + 0.5,
node_shape="o",
)
nc = nx.draw_networkx_nodes(
G_ego,
pos,
nodelist=negative_nodes,
node_color=negative_colors,
cmap=cmap,
node_size=negative_node_sizes,
vmin=np.min(colors) - 0.5,
vmax=np.max(colors) + 0.5,
node_shape="d",
)
edges = G_ego.edges()
weights = [
groud_truth_edge_importance[graph_nodes.index(u), graph_nodes.index(v)]
for u, v in edges
]
edge_colors = ["red" if w > 0 else "blue" for w in weights]
weights = link_width_factor * np.abs(weights) / np.max(weights)
ec = nx.draw_networkx_edges(G_ego, pos, edge_color=edge_colors, width=weights)
plt.legend()
plt.colorbar(nc, ticks=np.arange(np.min(colors), np.max(colors) + 1))
plt.axis("off")
plt.show()
```
By comparing the above two figures, one can see that the integrated gradients are quite consistent with the brute-force approach. The main benefit of using integrated gradients is scalability. The gradient operations are very efficient to compute on deep learning frameworks with the parallelism provided by GPUs. Also, integrated gradients can give the importance of individual node features, for all nodes in the graph. Achieving this by brute-force approach is often non-trivial.
<table><tr><td>Run the latest release of this notebook:</td><td><a href="https://mybinder.org/v2/gh/stellargraph/stellargraph/master?urlpath=lab/tree/demos/interpretability/gcn-sparse-node-link-importance.ipynb" alt="Open In Binder" target="_parent"><img src="https://mybinder.org/badge_logo.svg"/></a></td><td><a href="https://colab.research.google.com/github/stellargraph/stellargraph/blob/master/demos/interpretability/gcn-sparse-node-link-importance.ipynb" alt="Open In Colab" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg"/></a></td></tr></table>
| github_jupyter |
# Pytorch Embedding NN code:
https://www.kaggle.com/sapthrishi007/jane-fastai-embedding-smoothnn5-300x5/notebook?scriptVersionId=53603141&select=Jane_EmbNN5_auc_400_400_400.pth
Apart from the 130 featres, it also creates feature embeddings from their tags in features.csv file.
## PyTorch train code:
https://www.kaggle.com/a763337092/neural-network-starter-pytorch-version
https://www.kaggle.com/a763337092/pytorch-resnet-starter-training
## TensorFlow training code:
https://www.kaggle.com/code1110/jane-street-with-keras-nn-overfit
## Blending PyTorch and TensorFlow code:
https://www.kaggle.com/a763337092/blending-tensorflow-and-pytorch
```
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
# You can write up to 20GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
```
## PyTorch ResNet part
```
import os
import time
import pickle
import random
import numpy as np
import pandas as pd
from tqdm import tqdm
from collections import namedtuple
from sklearn.metrics import log_loss, roc_auc_score
import torch
import torch.nn as nn
from torch.autograd import Variable
from torch.utils.data import DataLoader
from torch.nn import CrossEntropyLoss, MSELoss
from torch.nn.modules.loss import _WeightedLoss
import torch.nn.functional as F
import warnings
warnings.filterwarnings ("ignore")
pd.set_option('display.max_columns', 100)
pd.set_option('display.max_rows', 100)
DATA_PATH = '../input/jane-street-market-prediction/'
NFOLDS = 5
TRAIN = False
CACHE_PATH = '../input/mlp012003weights'
def save_pickle(dic, save_path):
with open(save_path, 'wb') as f:
# with gzip.open(save_path, 'wb') as f:
pickle.dump(dic, f)
def load_pickle(load_path):
with open(load_path, 'rb') as f:
# with gzip.open(load_path, 'rb') as f:
message_dict = pickle.load(f)
return message_dict
feat_cols = [f'feature_{i}' for i in range(130)]
target_cols = ['action', 'action_1', 'action_2', 'action_3', 'action_4']
f_mean = np.load(f'{CACHE_PATH}/f_mean_online.npy')
##### Making features
all_feat_cols = [col for col in feat_cols]
all_feat_cols.extend(['cross_41_42_43', 'cross_1_2'])
##### Model&Data fnc
class Model(nn.Module):
def __init__(self):
super(Model, self).__init__()
self.batch_norm0 = nn.BatchNorm1d(len(all_feat_cols))
self.dropout0 = nn.Dropout(0.2)
dropout_rate = 0.2
hidden_size = 256
self.dense1 = nn.Linear(len(all_feat_cols), hidden_size)
self.batch_norm1 = nn.BatchNorm1d(hidden_size)
self.dropout1 = nn.Dropout(dropout_rate)
self.dense2 = nn.Linear(hidden_size+len(all_feat_cols), hidden_size)
self.batch_norm2 = nn.BatchNorm1d(hidden_size)
self.dropout2 = nn.Dropout(dropout_rate)
self.dense3 = nn.Linear(hidden_size+hidden_size, hidden_size)
self.batch_norm3 = nn.BatchNorm1d(hidden_size)
self.dropout3 = nn.Dropout(dropout_rate)
self.dense4 = nn.Linear(hidden_size+hidden_size, hidden_size)
self.batch_norm4 = nn.BatchNorm1d(hidden_size)
self.dropout4 = nn.Dropout(dropout_rate)
self.dense5 = nn.Linear(hidden_size+hidden_size, len(target_cols))
self.Relu = nn.ReLU(inplace=True)
self.PReLU = nn.PReLU()
self.LeakyReLU = nn.LeakyReLU(negative_slope=0.01, inplace=True)
# self.GeLU = nn.GELU()
self.RReLU = nn.RReLU()
def forward(self, x):
x = self.batch_norm0(x)
x = self.dropout0(x)
x1 = self.dense1(x)
x1 = self.batch_norm1(x1)
# x = F.relu(x)
# x = self.PReLU(x)
x1 = self.LeakyReLU(x1)
x1 = self.dropout1(x1)
x = torch.cat([x, x1], 1)
x2 = self.dense2(x)
x2 = self.batch_norm2(x2)
# x = F.relu(x)
# x = self.PReLU(x)
x2 = self.LeakyReLU(x2)
x2 = self.dropout2(x2)
x = torch.cat([x1, x2], 1)
x3 = self.dense3(x)
x3 = self.batch_norm3(x3)
# x = F.relu(x)
# x = self.PReLU(x)
x3 = self.LeakyReLU(x3)
x3 = self.dropout3(x3)
x = torch.cat([x2, x3], 1)
x4 = self.dense4(x)
x4 = self.batch_norm4(x4)
# x = F.relu(x)
# x = self.PReLU(x)
x4 = self.LeakyReLU(x4)
x4 = self.dropout4(x4)
x = torch.cat([x3, x4], 1)
x = self.dense5(x)
return x
if True:
# device = torch.device("cuda:0")
device = torch.device("cpu")
model_list = []
tmp = np.zeros(len(feat_cols))
for _fold in range(NFOLDS):
torch.cuda.empty_cache()
model = Model()
model.to(device)
model_weights = f"{CACHE_PATH}/online_model{_fold}.pth"
model.load_state_dict(torch.load(model_weights, map_location=torch.device('cpu')))
model.eval()
model_list.append(model)
```
## TensorFlow part
```
!ls ../input/jane-street-with-keras-nn-overfit
from tensorflow.keras.layers import Input, Dense, BatchNormalization, Dropout, Concatenate, Lambda, GaussianNoise, Activation
from tensorflow.keras.models import Model, Sequential
from tensorflow.keras.losses import BinaryCrossentropy
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.callbacks import EarlyStopping
from tensorflow.keras.layers.experimental.preprocessing import Normalization
import tensorflow as tf
import tensorflow_addons as tfa
import numpy as np
import pandas as pd
from tqdm import tqdm
from random import choices
SEED = 1111
np.random.seed(SEED)
# fit
def create_mlp(
num_columns, num_labels, hidden_units, dropout_rates, label_smoothing, learning_rate
):
inp = tf.keras.layers.Input(shape=(num_columns,))
x = tf.keras.layers.BatchNormalization()(inp)
x = tf.keras.layers.Dropout(dropout_rates[0])(x)
for i in range(len(hidden_units)):
x = tf.keras.layers.Dense(hidden_units[i])(x)
x = tf.keras.layers.BatchNormalization()(x)
x = tf.keras.layers.Activation(tf.keras.activations.swish)(x)
x = tf.keras.layers.Dropout(dropout_rates[i + 1])(x)
x = tf.keras.layers.Dense(num_labels)(x)
out = tf.keras.layers.Activation("sigmoid")(x)
model = tf.keras.models.Model(inputs=inp, outputs=out)
model.compile(
optimizer=tfa.optimizers.RectifiedAdam(learning_rate=learning_rate),
loss=tf.keras.losses.BinaryCrossentropy(label_smoothing=label_smoothing),
metrics=tf.keras.metrics.AUC(name="AUC"),
)
return model
epochs = 200
batch_size = 4096
hidden_units = [160, 160, 160]
dropout_rates = [0.2, 0.2, 0.2, 0.2]
label_smoothing = 1e-2
learning_rate = 1e-3
tf.keras.backend.clear_session()
tf.random.set_seed(SEED)
clf = create_mlp(
len(feat_cols), 5, hidden_units, dropout_rates, label_smoothing, learning_rate
)
clf.load_weights('../input/jane-street-with-keras-nn-overfit/model.h5')
tf_models = [clf]
```
# EmbeddingsNN
```
N_FEAT_TAGS = 29 # No of tags in features.csv
DEVICE = device
N_FEATURES = 130
THREE_HIDDEN_LAYERS = [400, 400, 400]
class FFN (nn.Module):
def __init__(self, inputCount=130, outputCount=5, hiddenLayerCounts=[150, 150, 150],
drop_prob=0.2, nonlin=nn.SiLU (), isOpAct=False):
super(FFN, self).__init__()
self.nonlin = nonlin
self.dropout = nn.Dropout (drop_prob)
self.batchnorm0 = nn.BatchNorm1d (inputCount)
self.dense1 = nn.Linear (inputCount, hiddenLayerCounts[0])
self.batchnorm1 = nn.BatchNorm1d (hiddenLayerCounts[0])
self.dense2 = nn.Linear(hiddenLayerCounts[0], hiddenLayerCounts[1])
self.batchnorm2 = nn.BatchNorm1d (hiddenLayerCounts[1])
self.dense3 = nn.Linear(hiddenLayerCounts[1], hiddenLayerCounts[2])
self.batchnorm3 = nn.BatchNorm1d (hiddenLayerCounts[2])
self.outDense = None
if outputCount > 0:
self.outDense = nn.Linear (hiddenLayerCounts[-1], outputCount)
self.outActivtn = None
if isOpAct:
if outputCount == 1 or outputCount == 2:
self.outActivtn = nn.Sigmoid ()
elif outputCount > 0:
self.outActivtn = nn.Softmax (dim=-1)
return
def forward (self, X):
# X = self.dropout (self.batchnorm0 (X))
X = self.batchnorm0 (X)
X = self.dropout (self.nonlin (self.batchnorm1 (self.dense1 (X))))
X = self.dropout (self.nonlin (self.batchnorm2 (self.dense2 (X))))
X = self.dropout (self.nonlin (self.batchnorm3 (self.dense3 (X))))
if self.outDense:
X = self.outDense (X)
if self.outActivtn:
X = self.outActivtn (X)
return X
class Emb_NN_Model (nn.Module):
def __init__(self, three_hidden_layers=THREE_HIDDEN_LAYERS, embed_dim=(N_FEAT_TAGS), csv_file='../input/jane-street-market-prediction/features.csv'):
super (Emb_NN_Model, self).__init__()
global N_FEAT_TAGS
N_FEAT_TAGS = 29
# store the features to tags mapping as a datframe tdf, feature_i mapping is in tdf[i, :]
dtype = {'tag_0' : 'int8'}
for i in range (1, 29):
k = 'tag_' + str (i)
dtype[k] = 'int8'
t_df = pd.read_csv (csv_file, usecols=range (1,N_FEAT_TAGS+1), dtype=dtype)
t_df['tag_29'] = np.array ([1] + ([0] * (t_df.shape[0]-1)) ).astype ('int8')
self.features_tag_matrix = torch.tensor (t_df.to_numpy ())
N_FEAT_TAGS += 1
# print ('self.features_tag_matrix =', self.features_tag_matrix)
# embeddings for the tags. Each feature is taken a an embedding which is an avg. of its' tag embeddings
self.embed_dim = embed_dim
self.tag_embedding = nn.Embedding (N_FEAT_TAGS+1, embed_dim) # create a special tag if not known tag for any feature
self.tag_weights = nn.Linear (N_FEAT_TAGS, 1)
drop_prob = 0.5
self.ffn = FFN (inputCount=(130+embed_dim), outputCount=0, hiddenLayerCounts=[(three_hidden_layers[0]+embed_dim), (three_hidden_layers[1]+embed_dim), (three_hidden_layers[2]+embed_dim)], drop_prob=drop_prob)
self.outDense = nn.Linear (three_hidden_layers[2]+embed_dim, 5)
return
def features2emb (self):
"""
idx : int feature index 0 to N_FEATURES-1 (129)
"""
all_tag_idxs = torch.LongTensor (np.arange (N_FEAT_TAGS)) #.to (DEVICE) # (29,)
tag_bools = self.features_tag_matrix # (130, 29)
# print ('tag_bools.shape =', tag_bools.size())
f_emb = self.tag_embedding (all_tag_idxs).repeat (130, 1, 1) #;print ('1. f_emb =', f_emb) # (29, 7) * (130, 1, 1) = (130, 29, 7)
# print ('f_emb.shape =', f_emb.size())
f_emb = f_emb * tag_bools[:, :, None] #;print ('2. f_emb =', f_emb) # (130, 29, 7) * (130, 29, 1) = (130, 29, 7)
# print ('f_emb.shape =', f_emb.size())
# Take avg. of all the present tag's embeddings to get the embedding for a feature
s = torch.sum (tag_bools, dim=1) # (130,)
# print ('s =', s)
f_emb = torch.sum (f_emb, dim=-2) / s[:, None] # (130, 7)
# print ('f_emb =', f_emb)
# print ('f_emb.shape =', f_emb.shape)
# take a linear combination of the present tag's embeddings
# f_emb = f_emb.permute (0, 2, 1) # (130, 7, 29)
# f_emb = self.tag_weights (f_emb) #;print ('3. f_emb =', f_emb) # (130, 7, 1)
# f_emb = torch.squeeze (f_emb, dim=-1) #;print ('4. f_emb =', f_emb) # (130, 7)
return f_emb
def forward (self, cat_featrs, features):
"""
when you call `model (x ,y, z, ...)` then this method is invoked
"""
cat_featrs = None
features = features.view (-1, N_FEATURES)
f_emb = self.features2emb () #;print ('5. f_emb =', f_emb); print ('6. features =', features) # (130, 7)
# print ('features.shape =', features.shape, 'f_emb.shape =', f_emb.shape)
features_2 = torch.matmul (features, f_emb) #;print ('7. features =', features) # (1, 130) * (130, 7) = (1, 7)
# print ('features.shape =', features.shape)
# Concatenate the two features (features + their embeddings)
features = torch.hstack ((features, features_2))
x = self.ffn (features) #;print ('8. x.shape = ', x.shape, 'x =', x) # (1, 7) -> (1, 7)
# x = self.layer_normal (x + features) #;print ('9. x.shape = ', x.shape, 'x =', x) # (1, 7) -> (1, 2)
out_logits = self.outDense (x) #;print ('10. out_logits.shape = ', out_logits.shape, 'out_logits =', out_logits)
# return sigmoid probs
# out_probs = F.sigmoid (out_logits)
return out_logits
embNN_model = Emb_NN_Model ()
try:
embNN_model.load_state_dict (torch.load ("../input/embedd9203/Jane_EmbNN5_auc_400_400_400.pth"))
except:
embNN_model.load_state_dict (torch.load ("../input/embedd9203/Jane_EmbNN5_auc_400_400_400.pth", map_location='cpu'))
embNN_model = embNN_model.eval ()
```
## Inference
```
import janestreet
env = janestreet.make_env()
env_iter = env.iter_test()
if True:
for (test_df, pred_df) in tqdm(env_iter):
if test_df['weight'].item() > 0:
x_tt = test_df.loc[:, feat_cols].values
if np.isnan(x_tt.sum()):
x_tt = np.nan_to_num(x_tt) + np.isnan(x_tt) * f_mean
cross_41_42_43 = x_tt[:, 41] + x_tt[:, 42] + x_tt[:, 43]
cross_1_2 = x_tt[:, 1] / (x_tt[:, 2] + 1e-5)
feature_inp = np.concatenate((
x_tt,
np.array(cross_41_42_43).reshape(x_tt.shape[0], 1),
np.array(cross_1_2).reshape(x_tt.shape[0], 1),
), axis=1)
# torch_pred
torch_pred = np.zeros((1, len(target_cols)))
for model in model_list:
torch_pred += model(torch.tensor(feature_inp, dtype=torch.float).to(device)).sigmoid().detach().cpu().numpy() / NFOLDS
torch_pred = np.median(torch_pred)
# tf_pred
tf_pred = np.median(np.mean([model(x_tt, training = False).numpy() for model in tf_models],axis=0))
# torch embedding_NN pred
x_tt = torch.tensor (x_tt).float ().view (-1, 130)
embnn_p = np.median (torch.sigmoid (embNN_model (None, x_tt)).detach ().cpu ().numpy ().reshape ((-1, 5)), axis=1) # not logits, actually sigmoid probabilities
# avg
pred_pr = torch_pred*0.41 + tf_pred*0.41 + embnn_p*0.18
pred_df.action = np.where (pred_pr >= 0.4918, 1, 0).astype (int)
else:
pred_df.action = 0
env.predict(pred_df)
print ("Done !")
```
| github_jupyter |
```
%load_ext watermark
%watermark -d -u -a 'Andreas Mueller, Kyle Kastner, Sebastian Raschka' -v -p numpy,scipy,matplotlib,scikit-learn
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
```
# SciPy 2016 Scikit-learn Tutorial
## Manifold Learning
One weakness of PCA is that it cannot detect non-linear features. A set
of algorithms known as *Manifold Learning* have been developed to address
this deficiency. A canonical dataset used in Manifold learning is the
*S-curve*:
```
from sklearn.datasets import make_s_curve
X, y = make_s_curve(n_samples=1000)
from mpl_toolkits.mplot3d import Axes3D
ax = plt.axes(projection='3d')
ax.scatter3D(X[:, 0], X[:, 1], X[:, 2], c=y)
ax.view_init(10, -60);
```
This is a 2-dimensional dataset embedded in three dimensions, but it is embedded
in such a way that PCA cannot discover the underlying data orientation:
```
from sklearn.decomposition import PCA
X_pca = PCA(n_components=2).fit_transform(X)
plt.scatter(X_pca[:, 0], X_pca[:, 1], c=y);
```
Manifold learning algorithms, however, available in the ``sklearn.manifold``
submodule, are able to recover the underlying 2-dimensional manifold:
```
from sklearn.manifold import Isomap
iso = Isomap(n_neighbors=15, n_components=2)
X_iso = iso.fit_transform(X)
plt.scatter(X_iso[:, 0], X_iso[:, 1], c=y);
```
## Manifold learning on the digits data
We can apply manifold learning techniques to much higher dimensional datasets, for example the digits data that we saw before:
```
from sklearn.datasets import load_digits
digits = load_digits()
fig, axes = plt.subplots(2, 5, figsize=(10, 5),
subplot_kw={'xticks':(), 'yticks': ()})
for ax, img in zip(axes.ravel(), digits.images):
ax.imshow(img, interpolation="none", cmap="gray")
```
We can visualize the dataset using a linear technique, such as PCA. We saw this already provides some intuition about the data:
```
# build a PCA model
pca = PCA(n_components=2)
pca.fit(digits.data)
# transform the digits data onto the first two principal components
digits_pca = pca.transform(digits.data)
colors = ["#476A2A", "#7851B8", "#BD3430", "#4A2D4E", "#875525",
"#A83683", "#4E655E", "#853541", "#3A3120","#535D8E"]
plt.figure(figsize=(10, 10))
plt.xlim(digits_pca[:, 0].min(), digits_pca[:, 0].max() + 1)
plt.ylim(digits_pca[:, 1].min(), digits_pca[:, 1].max() + 1)
for i in range(len(digits.data)):
# actually plot the digits as text instead of using scatter
plt.text(digits_pca[i, 0], digits_pca[i, 1], str(digits.target[i]),
color = colors[digits.target[i]],
fontdict={'weight': 'bold', 'size': 9})
plt.xlabel("first principal component")
plt.ylabel("second principal component");
```
Using a more powerful, nonlinear techinque can provide much better visualizations, though.
Here, we are using the t-SNE manifold learning method:
```
from sklearn.manifold import TSNE
tsne = TSNE(random_state=42)
# use fit_transform instead of fit, as TSNE has no transform method:
digits_tsne = tsne.fit_transform(digits.data)
plt.figure(figsize=(10, 10))
plt.xlim(digits_tsne[:, 0].min(), digits_tsne[:, 0].max() + 1)
plt.ylim(digits_tsne[:, 1].min(), digits_tsne[:, 1].max() + 1)
for i in range(len(digits.data)):
# actually plot the digits as text instead of using scatter
plt.text(digits_tsne[i, 0], digits_tsne[i, 1], str(digits.target[i]),
color = colors[digits.target[i]],
fontdict={'weight': 'bold', 'size': 9})
```
t-SNE has a somewhat longer runtime that other manifold learning algorithms, but the result is quite striking. Keep in mind that this algorithm is purely unsupervised, and does not know about the class labels. Still it is able to separate the classes very well (though the classes four, one and nine have been split into multiple groups).
## Exercises
Compare the results of applying isomap to the digits dataset to the results of PCA and t-SNE. Which result do you think looks best?
```
# %load solutions/22A_isomap_digits.py
from sklearn.manifold import Isomap
iso = Isomap(n_components=2)
digits_isomap = iso.fit_transform(digits.data)
plt.figure(figsize=(10, 10))
plt.xlim(digits_isomap[:, 0].min(), digits_isomap[:, 0].max() + 1)
plt.ylim(digits_isomap[:, 1].min(), digits_isomap[:, 1].max() + 1)
for i in range(len(digits.data)):
# actually plot the digits as text instead of using scatter
plt.text(digits_isomap[i, 0], digits_isomap[i, 1], str(digits.target[i]),
color = colors[digits.target[i]],
fontdict={'weight': 'bold', 'size': 9})
```
Given how well t-SNE separated the classes, one might be tempted to use this processing for classification. Try training a K-nearest neighbor classifier on digits data transformed with t-SNE, and compare to the accuracy on using the dataset without any transformation.
```
# %load solutions/22B_tsne_classification.py
from sklearn.manifold import TSNE
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(digits.data, digits.target, random_state=1)
clf = KNeighborsClassifier()
clf.fit(X_train, y_train)
print('KNeighborsClassifier accuracy without t-SNE: {}'.format(clf.score(X_test, y_test)))
tsne = TSNE(random_state=42)
digits_tsne_train = tsne.fit_transform(X_train)
digits_tsne_test = tsne.fit_transform(X_test)
clf = KNeighborsClassifier()
clf.fit(digits_tsne_train, y_train)
print('KNeighborsClassifier accuracy with t-SNE: {}'.format(clf.score(digits_tsne_test, y_test)))
```
| github_jupyter |
```
import os
import os.path as osp
import sys
import numpy as np
from pprint import pprint
# add matplotlib before cv2, otherwise bug
import matplotlib.pyplot as plt
%matplotlib inline
plt.rcParams['figure.figsize'] = (12.0, 10.0)
# add
sys.path.insert(0, '../tools')
import _init_paths
from roi_data_layer.layer import RoIDataLayer
from datasets.factory import get_imdb
from model.train_val import get_training_roidb
from layer_utils.proposal_target_layer import proposal_target_layer
from utils.mask_utils import recover_masks
from model.config import cfg
cfg.TRAIN.BATCH_SIZE = 12
cfg.TRAIN.BBOX_NORMALIZE_TARGETS_PRECOMPUTED = True
# torch
import torch
from torch.autograd import Variable
def vis(imdb, blobs, boxes, masks):
PIXEL_MEANS = np.array([[[102.9801, 115.9465, 122.7717]]])
im_data = blobs['data'][0].copy()
im_data += PIXEL_MEANS
im_data[:,:,:] = im_data[:,:,[2,1,0]]
im_data = im_data.astype(np.uint8)
plt.imshow(im_data)
ax = plt.gca()
num_objs = boxes.shape[0]
for ix in range(num_objs):
# add mask
m = masks[ix] # uint8
color_mask = np.random.random((1,3)).tolist()[0]
img = np.ones( (m.shape[0], m.shape[1], 3) )
for i in range(3):
img[:, :, i] = color_mask[i]
ax.imshow(np.dstack( (img, m/255.*0.5) ))
# add box and label
box = boxes[ix] # float32 (x1y1x2y2,cls)
x1 = int(round(box[0]))
y1 = int(round(box[1]))
x2 = int(round(box[2]))
y2 = int(round(box[3]))
cls_ix = int(box[4])
display_txt = imdb.classes[cls_ix]
coords = (x1, y1), x2-x1+1, y2-y1+1
ax.add_patch(plt.Rectangle(*coords, fill=False, edgecolor='red', linewidth=2))
ax.text(x1, y1, display_txt, bbox={'facecolor':'red', 'alpha':0.5})
# dataset
imdb = get_imdb('coco_2014_minival')
imdb.set_proposal_method('gt')
roidb = get_training_roidb(imdb)
# Layer
data_layer = RoIDataLayer(roidb, imdb.num_classes)
# get one batch
blobs = data_layer.forward()
print(blobs.keys())
print(blobs['gt_boxes'].shape)
# Fake some inputs
num_rpn_rois = min(3, blobs['gt_boxes'].shape[0])
rpn_rois = np.hstack([np.zeros((num_rpn_rois, 1)),
blobs['gt_boxes'][:num_rpn_rois, :4]]).astype(np.float32) # float (N, 5) [0xyxy]
rpn_rois = np.vstack([rpn_rois,
np.hstack([np.zeros((num_rpn_rois, 1)), rpn_rois[:, 1:]+100.0])]).astype(np.float32)
gt_boxes = blobs['gt_boxes'] # (M, 5) [xyxycls]
gt_masks = blobs['gt_masks'] # uint8 [0, 1]
# move to gpu Variable
rpn_rois = Variable(torch.from_numpy(rpn_rois).cuda())
gt_boxes = Variable(torch.from_numpy(gt_boxes).float().cuda())
rpn_scores = Variable(torch.randn(gt_boxes.size(0)).cuda())
rois, roi_scores, labels, bbox_targets, bbox_inside_weights, bbox_outside_weights, mask_targets = \
proposal_target_layer(rpn_rois, rpn_scores, gt_boxes, gt_masks, imdb.num_classes)
imh, imw = int(blobs['im_info'][0][0]), int(blobs['im_info'][0][1])
num_pos = mask_targets.size(0)
boxes = torch.cat([rois[:, 1:5], labels.view(-1,1)], 1).data.cpu().numpy()
boxes = boxes[:num_pos]
masks = recover_masks(mask_targets.data.cpu().numpy(), boxes, imh, imw, 'bilinear')
vis(imdb, blobs, boxes, masks)
```
| github_jupyter |
# Table of Contents
<p><div class="lev1 toc-item"><a href="#Load-Libraries" data-toc-modified-id="Load-Libraries-1"><span class="toc-item-num">1 </span>Load Libraries</a></div><div class="lev1 toc-item"><a href="#Load-data/Create-data-Generators" data-toc-modified-id="Load-data/Create-data-Generators-2"><span class="toc-item-num">2 </span>Load data/Create data Generators</a></div><div class="lev1 toc-item"><a href="#AUC-callback-function" data-toc-modified-id="AUC-callback-function-3"><span class="toc-item-num">3 </span>AUC callback function</a></div><div class="lev1 toc-item"><a href="#Load-the-model-&-weights" data-toc-modified-id="Load-the-model-&-weights-4"><span class="toc-item-num">4 </span>Load the model & weights</a></div><div class="lev1 toc-item"><a href="#Training" data-toc-modified-id="Training-5"><span class="toc-item-num">5 </span>Training</a></div><div class="lev1 toc-item"><a href="#Prediction" data-toc-modified-id="Prediction-6"><span class="toc-item-num">6 </span>Prediction</a></div>
Training after specifying class weights. Also, calculating AUC after every epoch.
# Load Libraries
```
import keras
from keras.preprocessing.image import ImageDataGenerator
from keras.preprocessing import image
from keras.models import Sequential, load_model, Model
from keras.layers import Activation, Dropout, Flatten, Dense, GlobalAveragePooling2D
from keras.optimizers import SGD
from keras.callbacks import EarlyStopping, ModelCheckpoint
from keras.applications.vgg16 import VGG16
from keras_tqdm import TQDMNotebookCallback
from datetime import datetime
import os
import numpy as np
import pandas as pd
import math
pd.options.display.max_rows = 40
```
# Load data/Create data Generators
```
validgen = ImageDataGenerator()
# 600/450 _ 500/375 _ 400/300 _ 300/225
img_width = 600
img_height = 450
train_data_dir = "data/train"
validation_data_dir = "data/valid"
test_data_dir = "data/test"
batch_size_train = 16
batch_size_val = 32
val_data = validgen.flow_from_directory(
directory = validation_data_dir,
target_size = (img_height, img_width),
batch_size = 568,
class_mode = "binary",
shuffle = False).next()
train_data = validgen.flow_from_directory(
directory = train_data_dir,
target_size = (img_height, img_width),
batch_size = 1727,
class_mode = "binary",
shuffle = False).next()
datagen = ImageDataGenerator(
rotation_range = 20,
width_shift_range = 0.2,
height_shift_range = 0.2,
horizontal_flip = True)
train_gen = datagen.flow_from_directory(
directory = train_data_dir,
target_size = (img_height, img_width),
batch_size = batch_size_train,
class_mode = "binary",
shuffle = True)
train_samples = len(train_gen.filenames)
```
# AUC callback function
```
from sklearn.metrics import roc_auc_score
from sklearn.metrics import accuracy_score
from sklearn.metrics import log_loss
class auc_callback(keras.callbacks.Callback):
def __init__(self, val_data, init_epoch):
self.val_x = val_data[0]
self.val_y = val_data[1]
self.init_epoch = init_epoch
def on_train_begin(self, logs={}):
return
def on_train_end(self, logs={}):
return
def on_epoch_begin(self, epoch, logs={}):
return
def on_epoch_end(self, epoch, logs={}):
self.model.save_weights('vgg-class-weights-epoch-' + str(self.init_epoch + epoch) + '.hdf5')
val_pred = self.model.predict(self.val_x, batch_size=32, verbose=0)
val_roc = roc_auc_score(self.val_y, val_pred[:,0])
val_loss = log_loss(self.val_y, np.append(1 - val_pred, val_pred, axis=1))
val_acc = accuracy_score(self.val_y, val_pred >= 0.5)
print('\nVal AUC: ' + str(val_roc))
print('\nVal Los: ' + str(val_loss))
print('\nVal Acc: ' + str(val_acc) + '\n')
return
def on_batch_begin(self, batch, logs={}):
return
def on_batch_end(self, batch, logs={}):
return
```
# Load the model & weights
```
vgg16 = VGG16(weights = 'imagenet',include_top=False)
x = vgg16.get_layer('block5_conv3').output
x = GlobalAveragePooling2D()(x)
x = Dense(256, activation='relu')(x)
x = Dropout(0.5)(x)
x = Dense(1, activation='sigmoid')(x)
model_final = Model(inputs=vgg16.input, outputs=x)
model_final.compile(loss = 'binary_crossentropy',
optimizer = SGD(lr = 0.0001, momentum = 0.9, decay = 1e-5),
metrics = ['accuracy'])
model_final.load_weights('./weights/weights-iter-6-epoch-05.hdf5')
val_pred = model_final.predict(val_data[0], batch_size=32)
log_loss(val_data[1], np.append(1 - val_pred, val_pred, axis=1))
accuracy_score(val_data[1], val_pred >= 0.5)
roc_auc_score(val_data[1], val_pred[:,0])
```
# Training
```
model_final.compile(loss = 'binary_crossentropy',
optimizer = SGD(lr = 0.0001, momentum = 0.9, decay = 1e-5, nesterov = True),
metrics = ['accuracy'])
model_final.fit_generator(generator = train_gen,
epochs = 10,
steps_per_epoch = math.ceil(1727 / batch_size_train),
validation_data = None,
verbose = 2,
callbacks = [auc_callback(val_data, 0), TQDMNotebookCallback()],
class_weight = {0: 1090/1727, 1: 637/1727})
model_final.load_weights('./vgg-class-weights-epoch-1.hdf5')
val_pred = model_final.predict(val_data[0], batch_size=32)
log_loss(val_data[1], np.append(1 - val_pred, val_pred, axis=1))
model_final.compile(loss = 'binary_crossentropy',
optimizer = SGD(lr = 0.00001, momentum = 0.9, decay = 1e-5, nesterov = True),
metrics = ['accuracy'])
model_final.fit_generator(generator = train_gen,
epochs = 10,
steps_per_epoch = math.ceil(1727 / batch_size_train),
validation_data = None,
verbose = 2,model_final.fit_generator(generator = train_gen,
epochs = 10,
steps_per_epoch = math.ceil(1727 / batch_size_train),
validation_data = None,
verbose = 2,
callbacks = [auc_callback(val_data, 5), TQDMNotebookCallback()],
class_weight = {0: 1090/1727, 1: 637/1727})
callbacks = [auc_callback(val_data, 5), TQDMNotebookCallback()],
class_weight = {0: 1090/1727, 1: 637/1727})
model_final.load_weights('./vgg-class-weights-epoch-6.hdf5')
val_pred = model_final.predict(val_data[0], batch_size=32)
log_loss(val_data[1], np.append(1 - val_pred, val_pred, axis=1))
accuracy_score(val_data[1], val_pred >= 0.5)
roc_auc_score(val_data[1], val_pred[:,0])
model_final.compile(loss = 'binary_crossentropy',
optimizer = SGD(lr = 0.00001, momentum = 0.9, decay = 1e-5, nesterov = True),
metrics = ['accuracy'])
model_final.fit_generator(generator = train_gen,
epochs = 10,
steps_per_epoch = math.ceil(1727 / batch_size_train),
validation_data = None,
verbose = 2,
callbacks = [auc_callback(val_data, 7), TQDMNotebookCallback()],
class_weight = {0: 1090/1727, 1: 637/1727})
```
# Prediction
```
model_final.load_weights('./vgg-class-weights-epoch-5.hdf5')
val_pred = model_final.predict(val_data[0], batch_size=32)
log_loss(val_data[1], np.append(1 - val_pred, val_pred, axis=1))
batch_size_test = 32
test_gen = validgen.flow_from_directory(
directory = test_data_dir,
target_size = (img_height, img_width),
batch_size = batch_size_test,
class_mode = "binary",
shuffle = False)
test_samples = len(test_gen.filenames)
preds = model_final.predict_generator(test_gen, math.ceil(test_samples / batch_size_test))
preds_filenames = test_gen.filenames
preds_filenames = [int(x.replace("unknown/", "").replace(".jpg", "")) for x in preds_filenames]
df_result = pd.DataFrame({'name': preds_filenames, 'invasive': preds[:,0]})
df_result = df_result.sort_values("name")
df_result.index = df_result["name"]
df_result = df_result.drop(["name"], axis=1)
df_result.to_csv("submission_10.csv", encoding="utf8", index=True)
from IPython.display import FileLink
FileLink('submission_10.csv')
# Got 0.99179 on LB
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import linearsolve as ls
import matplotlib.pyplot as plt
plt.style.use('classic')
%matplotlib inline
```
# Discussion: Week 9
In this Notebook, we will investigate how the degree to which the central bank responds to changes inflation affects how exogenous shocks to demand or inflation propagate.
## The New-Keynesian Model
The most basic version of the New-Keynesian Model can be expressed as:
\begin{align}
y_t & = E_t y_{t+1} - \left( r_{t} - \bar{r}\right) + g_t\\
i_{t} & = r_{t} + E_t \pi_{t+1}\\
i_{t} & = \bar{r} + \pi^T + \phi_{\pi}\big(\pi_t - \pi^T\big) + \phi_{y}\big(y_t - \bar{y}\big)\\
\pi_t -\pi^T & = \beta \left( E_t\pi_{t+1} - \pi^T\right) + \kappa (y_t -\bar{y})+ u_t,
\end{align}
where: $y_t$ is (log) output, $r_t$ is the real interest rate, $i_t$ is the nominal interest rate, $\pi_t$ is the rate of inflation between periods $t-1$ and $t$, $\bar{r}$ is the long-run average real interest rate or the *natural rate of interest*, $\beta$ is the household's subjective discount factor, and $\pi^T$ is the central bank's inflation target. The coefficients $\phi_{\pi}$ and $\phi_{y}$ reflect the degree of intensity to which the central bank *endogenously* adjusts the nominal interest rate in response to movements in inflation and output.
The variables $g_t$ and $u_t$ represent exogenous shocks to aggregate demand and inflation. They follow AR(1) processes:
\begin{align}
g_{t+1} & = \rho_g g_{t} + \epsilon^g_{t+1}\\
u_{t+1} & = \rho_u u_{t} + \epsilon^u_{t+1}
\end{align}
I have removed the monetary policy from the model because today we are going to focus on demand and inflation shocks. We will use the following parameterization:
| $$\bar{y}$$ | $$\beta$$ | $$\bar{r}$$ | $$\kappa$$ | $$\pi^T$$ | $$\phi_{\pi}$$ | $$\phi_y$$ | $$\rho_g$$ | $$\rho_u$$ |
|-------------|-----------|----------------|------------|-----------|----------------|------------|------------|------------|
| 0 | 0.995 | $$-\log\beta$$ | 0.1 | 0.02/4 | — | 0.5/4 | 0.5 | 0.5 |
```
# Create a variable called 'parameters' that stores the model parameter values in a Pandas Series. CELL PROVIDED
parameters = pd.Series(dtype=float)
parameters['y_bar'] = 0
parameters['beta'] = 0.995
parameters['r_bar'] = -np.log(parameters.beta)
parameters['kappa'] = 0.25
parameters['pi_T'] = 0.02/4
parameters['phi_pi'] = 1.5
parameters['phi_y'] = 0.5/4
parameters['rho_g'] = 0.5
parameters['rho_u'] = 0.5
# Create variable called 'var_names' that stores the variable names in a list with state variables ordered first
var_names = ['g','u','y','pi','i','r']
# Create variable called 'shock_names' that stores an exogenous shock name for each state variable.
shock_names = ['e_g','e_u']
# Define a function that evaluates the equilibrium conditions of the model solved for zero.
def equilibrium_equations(variables_forward,variables_current,parameters):
# Parameters
p = parameters
# Current variables
cur = variables_current
# Forward variables
fwd = variables_forward
# IS equation
is_equation = fwd.y - (cur.r -p.r_bar) + cur.g - cur.y
# Fisher_equation
fisher_equation = cur.r + fwd.pi - cur.i
# Monetary policy
monetary_policy = p.r_bar + p.pi_T + p.phi_pi*(cur.pi - p.pi_T) + p.phi_y*cur.y - cur.i
# Phillips curve
phillips_curve = p.beta*(fwd.pi- p.pi_T) + p.kappa*cur.y + cur.u - (cur.pi-p.pi_T)
# Demand process
demand_process = p.rho_g*cur.g - fwd.g
# Inflation process
inflation_process = p.rho_u*cur.u - fwd.u
# Stack equilibrium conditions into a numpy array
return np.array([
is_equation,
fisher_equation,
monetary_policy,
phillips_curve,
demand_process,
inflation_process
])
# Initialize the model into a variable named 'nk_model'
nk_model = ls.model(equations = equilibrium_equations,
n_states=2,
var_names=var_names,
shock_names=shock_names,
parameters = parameters)
# Compute the steady state numerically using .compute_ss() method of nk_model
guess = [0,0,0,0.01,0.01,0.01]
nk_model.compute_ss(guess)
```
## Exercise: Demand Shocks and $\phi_{\pi}$
For $\phi_{\pi} = 1.5,2,10,100$, compute impulse responses for output and inflation to a 0.01 unit increase in $g_t$. Simulate a total of 21 periods with the arriving in period 5. Construct a figure with two axes side-by-side with output ploted on the left and inflation plotted on the right.
```
# Create a variable called 'phi_pi_values' that stores the values for phi_pi provided above.
# Create a figure with dimensions 12x4. PROVIDED
fig = plt.figure(figsize=(12,4))
# Create the left axis. PROVIDED
ax1 = fig.add_subplot(1,2,1)
# Create the right axis. PROVIDED
ax2 = fig.add_subplot(1,2,2)
# Set the title of the left axis. PROVIDED
ax1.set_title('Output')
# Set the title of the right axis. PROVIDED
ax2.set_title('Inflation')
# Set the y-axis label of the left axis. PROVIDED
ax1.set_ylabel('Percent')
# Set the y-axis label of the right axis. PROVIDED
ax2.set_ylabel('Percent')
# Construct a for loop that iterates over the values in phi_pi
# Set the phi_pi value in nk_model.parameters to the current value of phi_pi
# Approximate and solve the model. Set log_linear argument to False since the model is already linear
# Compute impulse responses to a one percent shock to demand
# Plot output
# Plot inflation impulse response
# Construct legend. PROVIDED
ax2.legend(loc='center left', bbox_to_anchor=(1, 0.5))
```
**Questions**
1. How do the computed impulse responses for output change as $\phi_{\pi}$ increases from 1.5 to 100?
2. How do the computed impulse responses for inflation change as $\phi_{\pi}$ increases from 1.5 to 100?
3. Does the central banker face a trade-off between stabilizing output fluctuations or stabilizing inflation fluctuations in the presence of a demand shock?
**Answers**
1.
1.
3.
## Exercise: Inflation Shocks and $\phi_{\pi}$
For $\phi_{\pi} = 1.5,2,10,100$, compute impulse responses for output and inflation to a 0.01/4 unit increase in $u_t$. Simulate a total of 21 periods with the arriving in period 5. Construct a figure with two axes side-by-side with output ploted on the left and inflation plotted on the right.
```
# Create a variable called 'phi_pi_values' that stores the values for phi_pi provided above.
# Create a figure with dimensions 12x4. PROVIDED
fig = plt.figure(figsize=(12,4))
# Create the left axis. PROVIDED
ax1 = fig.add_subplot(1,2,1)
# Create the right axis. PROVIDED
ax2 = fig.add_subplot(1,2,2)
# Set the title of the left axis. PROVIDED
ax1.set_title('Output')
# Set the title of the right axis. PROVIDED
ax2.set_title('Inflation')
# Set the y-axis label of the left axis. PROVIDED
ax1.set_ylabel('Percent')
# Set the y-axis label of the right axis. PROVIDED
ax2.set_ylabel('Percent')
# Construct a for loop that iterates over the values in phi_pi
# Set the phi_pi value in nk_model.parameters to the current value of phi_pi
# Approximate and solve the model. Set log_linear argument to False since the model is already linear
# Compute impulse responses to a one percent shock to supply
# Plot output
# Plot inflation impulse response
# Construct legend. PROVIDED
ax2.legend(loc='center left', bbox_to_anchor=(1, 0.5))
```
**Questions**
1. How do the computed impulse responses for output change as $\phi_{\pi}$ increases from 1.5 to 100?
2. How do the computed impulse responses for inflation change as $\phi_{\pi}$ increases from 1.5 to 100?
3. Does the central banker face a trade-off between stabilizing output fluctuations or stabilizing inflation fluctuations in the presence of an inflation shock?
**Answers**
1.
1.
3.
| github_jupyter |
## PS3-1 A Simple Neural Network
#### (a)
Recall the following equations:
\begin{align*}
z^{[1]} & = W^{[1]} x + W_0^{[1]} \\
h & = \sigma (z^{[1]}) \\
z^{[2]} & = W^{[2]} h + W_0^{[2]} \\
o & = \sigma (z^{[2]}) \\
\ell & = \frac{1}{m} \sum_{i = 1}^{m} (o^{(i)} - y^{(i)})^2 = \frac{1}{m} \sum_{i = 1}^{m} J^{(i)}
\end{align*}
For a single training example,
\begin{align*}
\frac{\partial J}{\partial w_{1,2}^{[1]}} & = \frac{\partial J}{\partial o} \frac{\partial o}{\partial z^{[2]}} \frac{\partial z^{[2]}}{\partial h_2} \frac{\partial h_2}{\partial z_2^{[1]}} \frac{\partial z_2^{[1]}}{\partial w_{1,2}^{[1]}} \\
& = 2 (o - y) \cdot o (1 - o) \cdot w_2^{[2]} \cdot h_2 (1 - h_2) \cdot x_1
\end{align*}
where $h_2 = w_{1,2}^{[1]} x_1 + w_{2,2}^{[1]} x_2 + w_{0,2}^{[1]}$.
Therefore, the gradient descent update rule for $w_{1,2}^{[1]}$ is
$$w_{1,2}^{[1]} := w_{1,2}^{[1]} - \alpha \frac{2}{m} \sum_{i = 1}^{m} (o^{(i)} - y^{(i)}) \cdot o^{(i)} (1 - o^{(i)}) \cdot w_2^{[2]} \cdot h_2^{(i)} (1 - h_2^{(i)}) \cdot x_1^{(i)}$$
where $h_2^{(i)} = w_{1,2}^{[1]} x_1^{(i)} + w_{2,2}^{[1]} x_2^{(i)} + w_{0,2}^{[1]}$.
#### (b)
It is possible. The three neurons can be treated as three independent linear classifiers. The three decision boundaries
form a triangle that classifies the outside data into class 1, and the inside ones into class 0.
\begin{align*}
w_{1,1}^{[1]} x_1 + x_{2,1}^{[1]} x_2 + w_{0,1}^{[1]} & = 0 \\
w_{1,2}^{[1]} x_1 + x_{2,2}^{[1]} x_2 + w_{0,2}^{[1]} & = 0 \\
w_{1,3}^{[1]} x_1 + x_{2,3}^{[1]} x_2 + w_{0,3}^{[1]} & = 0
\end{align*}
Plug in some data points and solve the equations, we can obtain the weights. The weights vary upon the choice of the decision boundaries.
Here is one possible solution:
```
w = {}
w['hidden_layer_0_1'] = 0.5
w['hidden_layer_1_1'] = -1
w['hidden_layer_2_1'] = 0
w['hidden_layer_0_2'] = 0.5
w['hidden_layer_1_2'] = 0
w['hidden_layer_2_2'] = -1
w['hidden_layer_0_3'] = -4
w['hidden_layer_1_3'] = 1
w['hidden_layer_2_3'] = 1
w['output_layer_0'] = -0.5
w['output_layer_1'] = 1
w['output_layer_2'] = 1
w['output_layer_3'] = 1
```
#### (c)
No, it is not possible to achieve 100% accuracy using identity function as the activation functions for $h_1$, $h_2$ and $h_3$. Because
\begin{align*}
o & = \sigma (z^{[2]}) \\
& = \sigma (W^{[2]} h + W_0^{[2]}) \\
& = \sigma (W^{[2]} (W^{[1]} x + W_0^{[1]}) + W_0^{[2]}) \\
& = \sigma (W^{[2]} W^{[1]} x + W^{[2]} W_0^{[1]} + W_0^{[2]}) \\
& = \sigma (\tilde{W} x + \tilde{W_0})
\end{align*}
where $\tilde{W} = W^{[2]} W^{[1]}$ and $\tilde{W_0} = W^{[2]} W_0^{[1]} + W_0^{[2]}$.
We can see that the resulting classifier is still linear, and it is not able to classify datasets that are not linearly separable with 100% accuracy.
| github_jupyter |
# Lecture 1: Introduction to Programming
## Agenda for the Class:
1. Python in-built datatypes
2. Basic mathematical operators and Precedence order
3. Python Interpreter vs Python for Scripting
Firstly we'll focus on the **datatypes**.
1. **Numeric**
2. **Strings**
3. **Lists**
General format for **Assigning** a variable a value:
Variable_name = Variable_Value
1. We **Do not** mention datatype while assigning a variable a value in Python. (i.e. Dynamically Typed)
2. **=** is used to assign a variable a value. ( L Value and R value)
3. A variable name must follow certain naming conventions. Example: '23', 'len' can be variable names.
4. There is no such thing as "variable declaration" or "variable initialization" in Python. It's only variable assignment
## Numeric data
```
a=1
b=3.14
# Assigning value 1 to variable a and 3.14 to variable b
```
Mathematical Operations on Variables:
1. Add ('+')
2. Multiple ('*')
3. Subtract ('-')
4. Divide ('/')
5. Modulo ('%')
6. Exponentiation (\*\*)
### **Order of Precedence**
Exponent > (Multiple, Divide, Modulo) > (Add, Subtract)
```
a = 20
b = 10
c = 15
d = 5
e = 0
e = (a + b) * c / d #( 30 * 15 ) / 5
print ("Value of (a + b) * c / d is ", e)
e = ((a + b) * c) / d # (30 * 15 ) / 5
print ("Value of ((a + b) * c) / d is ", e)
e = (a + b) * (c / d); # (30) * (15/5)
print ("Value of (a + b) * (c / d) is ", e)
e = a + (b * c) / d; # 20 + (150/5)
print ("Value of a + (b * c) / d is ", e)
```
#### In case you are using Python 2 and want floating point division (e.g: 4/3 --> 1.33333333333 and not 4/3 --> 1) :
For Python shell type in : from __future__ import print_function, division
For a ".py" file : Use that import statement in the beginning of your Python file.
## Strings
1. **Immutable** datatype
2. String enclosed within **" String"** or **'String'**
```
course_name = "Introduction to Programming"
question = "Having a good time ? ;)"
print(course_name)
print(question)
```
### Operations on Strings
1. Since strings are immutable, we can't change the value stored in a string
2. We can concatenate ('join') multiple strings.
3. Slice/substring operations
```
string_1 = "Hello World!"
n = len(string_1) # "len" gives us the number of characters in the string
print(string_1 + " has", n , "characters")
```
1. **Indexing** : Every charcater of the string can be accessed by it's position in the string.
2. Indexing starts from zero.
3. Syntax
string_name[index_number]
Example:
```
print(string_1[0])
print(string_1[1])
print(string_1[-2])
```
Negative Indexing:
string[-1] gives us the last character
string[-2] gives us the second last character
and so on...
#### Slicing operations
Syntax:
string_name[start_index,end_index]
```
print(string_1[0:2])
print(string_1[5 : len(string_1)])
print(string_1[0:4]+string_1[4:len(string_1)])
```
## Lists
1. Initializing syntax: \n
list_name = [value_1,value_2,...,value_n]
2. Behaviour Similar to strings
3. Mutable
4. Can contain multiple data types.
```
primes = [2,3,5,8,11]
print(primes)
print(primes[0])
print(len(primes))
classroom = ['L','T', 1]
print(classroom)
print((classroom[2]+ 4))
```
### What to do next?
1.Play around in the python interpreter to explore various functionalities
2.Understand the rules to name variables.
| github_jupyter |
```
import pandas as pd
import numpy as np
import json
import re
import os
import chardet
from io import BytesIO
import spacy
from spacy_langdetect import LanguageDetector
import matplotlib.pyplot as plt
%matplotlib inline
# --------- Pandas Settings ---------- #
pd.set_option('display.max_rows', 200)
pd.set_option('display.max_columns', 200)
pd.set_option('display.max_colwidth', -1)
# --------- Load Spacy Model --------- #
nlp = spacy.load("en_core_web_sm")
nlp.add_pipe(LanguageDetector(), name='language_detector', last=True)
```
## Load memegenerator.csv to pandas dataframe
```
DATA_DIR = '../data'
FILE_NAME = 'memegenerator-dataset/memegenerator.csv'
FILE_PATH = os.path.join(DATA_DIR, FILE_NAME)
SAVE_FILE_PATH = os.path.join(DATA_DIR, 'cleaned_memegenerator.json')
# Check csv encoding
rawdata = open(FILE_PATH, 'rb').read()
result = chardet.detect(rawdata)
charenc = result['encoding']
charenc
# Convert csv to dataframe
with open(FILE_PATH,'rb') as f:
df = pd.read_csv(BytesIO(f.read().decode('UTF-16').encode('UTF-8')), sep='\t+')
df.head()
# Simplifying df to show only columns of need
wanted_columns = ['Archived URL', 'Base Meme Name',
'Alternate Text', 'Display Name',
'Upper Text', 'Lower Text']
df = df[wanted_columns]
df.head()
# Lowercase column names & replace spaces with underscores
columns = list(df.columns)
columns = [x.lower() for x in columns]
columns = [x.replace(' ', '_') for x in columns]
df.columns = columns
df.head()
# Compare base_meme_name to display_name
print(df.shape)
df[df['base_meme_name'] != df['display_name']].shape
# Create function to clean the dataframe
def wrangling(df: pd.core.frame.DataFrame):
df = df.copy()
column_names = list(df.columns)
pattern = r'^[.,\/#!$%\^&\*;:{}=\-_`~()+]{4,}'
# Remove extra whitespace and lowercase everything
for i in column_names:
if i != 'archived_url':
df[i] = df[i].str.lower().str.strip()
# Get rid of pesky punctuations
df = df[~df[i].str.contains(pat=pattern, regex=True, na=False)]
return df
df = wrangling(df)
df.head()
df[df['upper_text'].isnull() == True]
# Drop those 2 useless rows above
print(df.shape)
df = df.drop([39507, 43566])
df.shape
print(df.shape)
df[df['lower_text'].isnull() == True]
# Remove all null values
print(df.shape)
df = df.dropna(axis=0)
print(df.shape)
df.head()
# Check distribution of meme text lengths
df['alternate_text'].str.len().hist()
```
## Set upper and lower bounds on the alternate_text
```
# Remove entries whose len is < 5 and > 150
df = df[(df['alternate_text'].str.len() > 4) & (df['alternate_text'].str.len() < 150)]
print(df.shape)
df.sample(10)
# Check distribution of meme text lengths AFTER bounds set
df['alternate_text'].str.len().hist();
df['base_meme_name'].value_counts()[df['base_meme_name'].value_counts()>99]
# Remove laggard punctuations
import string
column_names = ['base_meme_name', 'alternate_text', 'display_name',
'upper_text', 'lower_text']
for i in column_names:
df[i] = df[i].str.translate(str.maketrans('', '', string.punctuation))
df.head()
# Observe samples where base_meme_name != display_name
print(df[df['base_meme_name'] != df['display_name']].shape)
df[df['base_meme_name'] != df['display_name']]
df['base_meme_name'].nunique(), df['display_name'].nunique()
```
## Get data from memefly database to create new dataframe
```
meme_name_json = ["advice-dog","advice-god","advice-peeta","annoying-facebook-girl","anti-joke-chicken","archer","art-student-owl","back-in-my-day","bad-advice-cat","bad-luck-brian","batman-slapping-robin","bear-grylls","buddy-the-elf","business-cat","butthurt-dweller","chemistry-cat","chill-out-lemur","chubby-bubbles-girl","chuck-norris","close-enough","conspiracy-keanu","contradictory-chris","courage-wolf","crazy-girlfriend-praying-mantis","cute-cat","depression-dog","disaster-girl","dont-you-squidward","dr-evil-laser","dumb-blonde","ermahgerd-berks","evil-cows","eye-of-sauron","felix-baumgartner","first-day-on-the-internet-kid","first-world-problems","forever-alone","foul-bachelorette-frog","foul-bachelor-frog","futurama-fry","gangnam-style","george-bush","good-guy-greg","grumpy-cat","han-solo","hedonism-bot","high-expectations-asian-father","hipster-ariel","hipster-kitty","how-tough-are-you","idiot-nerd-girl","ill-just-wait-here","insanity-wolf","i-see-dead-people","i-should-buy-a-boat-cat","i-will-find-you-and-kill-you","joe-biden","joseph-ducreux","kill-yourself-guy","kobe","lame-pun-coon","malicious-advice-mallard","matrix-morpheus","maury-lie-detector","mckayla-maroney-not-impressed","musically-oblivious-8th-grader","obama","one-does-not-simply","oprah-you-get-a","ordinary-muslim-man","original-stoner-dog","overly-manly-man","paranoid-parrot","patriotic-eagle","pepperidge-farm-remembers","philosoraptor","pickup-line-panda","priority-peter","professor-oak","rebecca-black","rich-raven","sad-x-all-the-y","scared-cat","scene-wolf","scumbag-boss","scumbag-steve","see-nobody-cares","sheltering-suburban-mom","slenderman","slowpoke","socially-awesome-awkward-penguin","socially-awesome-penguin","socially-awkward-penguin","south-park-craig","successful-black-man","success-kid","sudden-clarity-clarence","super-cool-ski-instructor","thats-a-paddlin","that-would-be-great","the-most-interesting-cat-in-the-world","the-most-interesting-man-in-the-world","unicorn-man","unpopular-opinion-puffin","vengeance-dad","we-will-rebuild","x-all-the-y","y-u-no"]
meme_name_json
meme_name_df = pd.Series(meme_name_json)
meme_name_df = meme_name_df.to_frame(name='meme_name')
meme_name_df.head()
# Replace hyphens with spaces for merge
meme_name_df['meme_name'] = meme_name_df['meme_name'].str.replace('-', ' ')
meme_name_df.head()
```
## Merge meme_name_df and df, left on meme_name right on display_name
```
df = meme_name_df.merge(df, left_on='meme_name', right_on='display_name', how='outer')
df.sample(25)
df[df['meme_name'].isnull() == True]
```
## Check for meme name aliases in base_meme_name and display_name
```
"""
NOTE: Row 26238 has 'advice dog' in base_meme_name, but because the display_name cell
was empty when the merge happened, the meme_name column shows NaN.
"""
df[(df['base_meme_name'].str.contains('advice')) & (df['base_meme_name'].str.contains('dog'))]
```
### Verifying whether our merge worked 100% correctly
There may be instances where the merge didn't happen even though
the meme name is correct. Since we merged meme_name_df with our
original df, there may be some instances where the meme name in
base_meme_name is actually correct.
```
# Create list of all meme names from memefly database
meme_names = list(df['meme_name'].unique())
meme_names = meme_names[:108]
name_sets = []
for i in range(len(meme_names)):
split = meme_names[i].split()
name_sets.append(split)
meme_names
"""
There may be instances where the merge didn't happen even though
the meme name is correct. Since we merged meme_name_df with our
original df, there may be some instances where the meme name in
base_meme_name is actually correct.
"""
# Create dataframe containing only entries whose meme_name is NaN
nan_meme_names = df[df['meme_name'].isnull() == True]
# Create sets of the meme_name column and base_meme_name column
# Get only unique values
# We can perform arithmetic operations on sets
nan_list = set(list(nan_meme_names['base_meme_name'].unique()))
meme_list = set(meme_names)
# Create list of meme names from base_meme_name that are actually in meme_name
in_list = [i for i in nan_list if i in meme_list]
in_list = sorted(in_list)
in_list # 22 different instances of meme type that missed the merge
```
### Correct meme_name where the merge missed
We want to replace the NaN value in `meme_name` with one of the items from `in_list` if `base_meme_name` == one of the items in `in_list`
```
mask = (df['base_meme_name'].isin(in_list)) & (df['meme_name'].isnull() == True)
df['meme_name'] = np.where(mask, df['base_meme_name'], df['meme_name'])
df.loc[26148, 'meme_name'] # Was NaN prior to above code implementation
type(df.loc[55563, 'meme_name']) == float
df[(df['base_meme_name'].isin(in_list)) & (df['meme_name'].isnull() == True)]
name_sets
df[df['meme_name'].isnull() == True].shape
for name_list in name_sets:
condition = ((df['base_meme_name'].str.contains('(' + ' ?'.join(name_list) + ')',
regex=True,
flags=re.IGNORECASE))
& (df['meme_name'].isnull()==True))
df['meme_name'] = np.where(condition, ' '.join(name_list), df['meme_name'])
df[df['meme_name'].isnull() == True].shape
df[(df['base_meme_name'].str.contains(name_sets[9][0])) & (df['base_meme_name'].str.contains(name_sets[9][1])) & (df['meme_name'].isnull() == True)]
df[df['meme_name'] == 'bad luck brian'].shape
df[(df['base_meme_name'].str.contains('(' + ' ?'.join(name_sets[9]) + ')', regex=True, flags=re.IGNORECASE)) & (df['meme_name'].isnull()==True)]
condition = ((df['base_meme_name'].str.contains('(' + ' ?'.join(name_sets[9]) + ')',
regex=True,
flags=re.IGNORECASE))
& (df['meme_name'].isnull()==True))
df['meme_name'] = np.where(condition, ' '.join(name_sets[9]), df['meme_name'])
df[(df['base_meme_name'].str.contains('(' + ' ?'.join(name_sets[9]) + ')', regex=True, flags=re.IGNORECASE)) & (df['meme_name'].isnull()==True)]
df[df['meme_name'].isnull() == False].shape
df['meme_name'] = np.where(df['base_meme_name'] == 'dos equis',
'the most interesting man in the world',
df['meme_name'])
df[df['base_meme_name'] == 'dos equis']
df['meme_name'] = np.where(df['base_meme_name'] == 'all the things',
'x all the y',
df['meme_name'])
df[df['base_meme_name'] == 'all the things']
df['meme_name'] = np.where(df['base_meme_name'] == 'sudden realization ralph',
'sudden clarity clarence',
df['meme_name'])
df[df['base_meme_name'] == 'sudden realization ralph']
df['meme_name'] = np.where(df['base_meme_name'] == 'shut up and take my money fry',
'futurama fry',
df['meme_name'])
df[df['base_meme_name'] =='shut up and take my money fry']
df['meme_name'] = np.where(df['base_meme_name'] == 'batman bitchslap',
'batman slapping robin',
df['meme_name'])
df[df['base_meme_name'] == 'batman bitchslap']
df[df['meme_name'].isnull() == True].sample(100)
meme_list
```
## Remove non-english text
A lot of the memes are not in English. These will in no way benefit our model, so they should be removed. We will use SpaCy to accomplish this
```
def lang_detect(text: str) -> str:
"""
Takes a string and returns the language of its text.
Arguments
------------------------------------------------------------------
text (str).........................a string of text
Returns
-------------------------------------------------------------------
doc._.language['language'] (str)...value entry of language key from
spacy-langdetect library
"""
doc = nlp(text)
return doc._.language['language']
```
### Replacing null values with a blank string
_**IMPORTANT:**_ Use `df[df['column_name'] == '']` to apply booleean masks going forward!
```
# Replace NaNs with empty strings
# Must be done in order to use spacy-langdetect
df = df.replace(np.nan, '', regex=True)
df.isnull().sum()
df['lang'] = df['alternate_text'].apply(lang_detect)
df.sample(100)
# Filter df to show only rows with english text
df = df[df['lang'] == 'en']
print(df.shape)
df.sample(10)
# Convert empty string back to NaN
df = df.replace('', np.nan)
df.sample(10)
```
### Drop rows with NaN value in `meme_name`
```
df = df.dropna(axis=0)
print(df.shape)
df.sample(20)
```
### Group all texts by `meme_name`
```
df = df[['meme_name', 'archived_url', 'alternate_text',
'upper_text', 'lower_text']]
df.head()
# Create new column that contains a dictionary
# Key = upper_text
# Value = lower_text
df['text_parser'] = df[['upper_text', 'lower_text']].to_dict(orient='records')
df.sample(10)
grouped = df.groupby('meme_name')
alternate_txt_grouped = grouped['alternate_text'].apply(list).to_frame().reset_index()
parsed_txt_grouped = grouped['text_parser'].apply(list).to_frame().reset_index()
alternate_txt_grouped.head(1)
parsed_txt_grouped.head(1)
merged = alternate_txt_grouped.merge(parsed_txt_grouped, on='meme_name')
merged.head(1)
# Get archived_url for each meme_name
de_dups = df.drop_duplicates(subset='meme_name', keep='first').drop(columns=['alternate_text',
'upper_text',
'lower_text',
'text_parser'])
de_dups.head()
merged = de_dups.merge(merged, on='meme_name')
merged.head(1)
# Save to JSON
merged.to_json(SAVE_FILE_PATH, orient='records')
```
| github_jupyter |
# Dependencies
```
import os, warnings, shutil
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from tokenizers import ByteLevelBPETokenizer
from sklearn.utils import shuffle
from sklearn.model_selection import StratifiedKFold
from tweet_utility_scripts import *
from tweet_utility_preprocess_roberta_scripts_aux import *
import tweet_utility_preprocess_roberta_scripts_text as preprocess_text
SEED = 0
warnings.filterwarnings("ignore")
```
# Tokenizer
```
MAX_LEN = 96
base_path = '/kaggle/input/qa-transformers/roberta/'
vocab_path = base_path + 'roberta-base-vocab.json'
merges_path = base_path + 'roberta-base-merges.txt'
tokenizer = ByteLevelBPETokenizer(vocab_file=vocab_path, merges_file=merges_path, lowercase=True, add_prefix_space=True)
tokenizer.save('./')
```
# Load data
```
train_df = pd.read_csv('/kaggle/input/tweet-sentiment-extraction/train.csv')
# pre-process
train_df.dropna(inplace=True)
train_df = train_df.reset_index()
train_df.drop('index', axis=1, inplace=True)
train_df["text"] = train_df["text"].apply(lambda x: x.strip())
train_df["selected_text"] = train_df["selected_text"].apply(lambda x: x.strip())
train_df["text"] = train_df["text"].apply(lambda x: x.lower())
train_df["selected_text"] = train_df["selected_text"].apply(lambda x: x.lower())
train_df['jaccard'] = train_df.apply(lambda x: jaccard(x['text'], x['selected_text']), axis=1)
train_df['text_len'] = train_df['text'].apply(lambda x : len(x))
train_df['text_wordCnt'] = train_df['text'].apply(lambda x : len(x.split(' ')))
train_df['text_tokenCnt'] = train_df['text'].apply(lambda x : len(tokenizer.encode(x).ids))
train_df['selected_text_len'] = train_df['selected_text'].apply(lambda x : len(x))
train_df['selected_text_wordCnt'] = train_df['selected_text'].apply(lambda x : len(x.split(' ')))
train_df['selected_text_tokenCnt'] = train_df['selected_text'].apply(lambda x : len(tokenizer.encode(x).ids))
sentiment_cols = train_df['sentiment'].unique()
print('Train samples: %s' % len(train_df))
display(train_df.head())
display(train_df.describe())
```
## Tokenizer sanity check
```
for idx in range(10):
print('\nRow %d' % idx)
max_seq_len = 32
text = train_df['text'].values[idx]
selected_text = train_df['selected_text'].values[idx]
question = train_df['sentiment'].values[idx]
_, (target_start, target_end, _) = preprocess_roberta(' ' + text, selected_text, ' ' + question, tokenizer, max_seq_len)
question_encoded = tokenizer.encode(question).ids
question_size = len(question_encoded) + 3
decoded_text = decode(target_start.argmax(), target_end.argmax(), text, question_size, tokenizer)
print('text : "%s"' % text)
print('selected_text: "%s"' % selected_text)
print('decoded_text : "%s"' % decoded_text)
assert selected_text == decoded_text
```
## Data generation sanity check
```
for idx in range(5):
print('\nRow %d' % idx)
max_seq_len = 24
text = train_df['text'].values[idx]
selected_text = train_df['selected_text'].values[idx]
question = train_df['sentiment'].values[idx]
jaccard = train_df['jaccard'].values[idx]
selected_text_wordCnt = train_df['selected_text_wordCnt'].values[idx]
x_train, x_train_aux, x_train_aux_2, y_train, y_train_mask, y_train_aux = get_data(train_df[idx:idx+1], tokenizer, max_seq_len,
preprocess_fn=preprocess_roberta)
print('text : "%s"' % text)
print('jaccard : "%.4f"' % jaccard)
print('sentiment : "%s"' % question)
print('word count : "%d"' % selected_text_wordCnt)
print('input_ids : "%s"' % x_train[0][0])
print('attention_mask: "%s"' % x_train[1][0])
print('sentiment : "%d"' % x_train_aux[0])
print('sentiment OHE : "%s"' % x_train_aux_2[0])
print('selected_text : "%s"' % selected_text)
print('start : "%s"' % y_train[0][0])
print('end : "%s"' % y_train[1][0])
print('mask : "%s"' % y_train_mask[0])
print('jaccard : "%.4f"' % y_train_aux[0][0])
print('word count : "%d"' % y_train_aux[1][0])
assert len(x_train) == 2
assert len(x_train_aux) == 1
assert len(x_train_aux_2) == 1
assert len(y_train) == 2
assert len(y_train_mask) == 1
assert len(y_train_aux) == 3
assert len(x_train[0][0]) == len(x_train[1][0]) == max_seq_len
assert len(y_train[0][0]) == len(y_train[1][0]) == len(y_train_mask[0]) == max_seq_len
```
# 5-Fold split
```
folds = StratifiedKFold(n_splits=5, shuffle=True, random_state=SEED)
for fold_n, (train_idx, val_idx) in enumerate(folds.split(train_df, train_df['sentiment'])):
print('Fold: %s, Train size: %s, Validation size %s' % (fold_n+1, len(train_idx), len(val_idx)))
train_df[('fold_%s' % str(fold_n+1))] = 0
train_df[('fold_%s' % str(fold_n+1))].loc[train_idx] = 'train'
train_df[('fold_%s' % str(fold_n+1))].loc[val_idx] = 'validation'
```
## Sentiment distribution
```
for fold_n in range(folds.n_splits):
fold_n += 1
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(20, 8.7))
fig.suptitle('Fold %s' % fold_n, fontsize=22)
sns.countplot(x="sentiment", data=train_df[train_df[('fold_%s' % fold_n)] == 'train'], palette="GnBu_d", order=sentiment_cols, ax=ax1).set_title('Train')
sns.countplot(x="sentiment", data=train_df[train_df[('fold_%s' % fold_n)] == 'validation'], palette="GnBu_d", order=sentiment_cols, ax=ax2).set_title('Validation')
sns.despine()
plt.show()
```
## Word count distribution
```
for fold_n in range(folds.n_splits):
fold_n += 1
fig, (ax1, ax2) = plt.subplots(2, 1, figsize=(20, 8.7), sharex=True)
fig.suptitle('Fold %s' % fold_n, fontsize=22)
sns.distplot(train_df[train_df[('fold_%s' % fold_n)] == 'train']['text_wordCnt'], ax=ax1).set_title("Train")
sns.distplot(train_df[train_df[('fold_%s' % fold_n)] == 'validation']['text_wordCnt'], ax=ax2).set_title("Validation")
sns.despine()
plt.show()
```
# Output 5-fold set
```
train_df.to_csv('5-fold.csv', index=False)
display(train_df.head())
for fold_n in range(folds.n_splits):
fold_n += 1
base_path = 'fold_%d/' % fold_n
# Create dir
os.makedirs(base_path)
x_train, x_train_aux, x_train_aux_2, y_train, y_train_mask, y_train_aux = get_data(train_df[train_df[('fold_%s' % fold_n)] == 'train'], tokenizer,
MAX_LEN, preprocess_fn=preprocess_roberta)
x_valid, x_valid_aux, x_valid_aux_2, y_valid, y_valid_mask, y_valid_aux = get_data(train_df[train_df[('fold_%s' % fold_n)] == 'validation'], tokenizer,
MAX_LEN, preprocess_fn=preprocess_roberta)
np.save(base_path + 'x_train', np.array(x_train))
np.save(base_path + 'y_train', np.array(y_train))
np.save(base_path + 'x_valid', np.array(x_valid))
np.save(base_path + 'y_valid', np.array(y_valid))
np.save(base_path + 'x_train_aux', np.array(x_train_aux))
np.save(base_path + 'x_train_aux2', np.array(x_train_aux_2))
np.save(base_path + 'y_train_mask', np.array(y_train_mask))
np.save(base_path + 'y_train_aux', np.array(y_train_aux))
np.save(base_path + 'x_valid_aux', np.array(x_valid_aux))
np.save(base_path + 'x_valid_aux2', np.array(x_valid_aux_2))
np.save(base_path + 'y_valid_mask', np.array(y_valid_mask))
np.save(base_path + 'y_valid_aux', np.array(y_valid_aux))
# Compress logs dir
!tar -czf fold_1.tar.gz fold_1
!tar -czf fold_2.tar.gz fold_2
!tar -czf fold_3.tar.gz fold_3
!tar -czf fold_4.tar.gz fold_4
!tar -czf fold_5.tar.gz fold_5
# Delete logs dir
shutil.rmtree('fold_1')
shutil.rmtree('fold_2')
shutil.rmtree('fold_3')
shutil.rmtree('fold_4')
shutil.rmtree('fold_5')
```
# Output 5-fold set (balanced)
```
for fold_n in range(folds.n_splits):
fold_n += 1
base_path = 'balanced_fold_%d/' % fold_n
# Create dir
os.makedirs(base_path)
train_fold = train_df[train_df[('fold_%s' % fold_n)] == 'train'].copy()
valid_fold = train_df[train_df[('fold_%s' % fold_n)] == 'validation'].copy()
# Sample data by lower bound
lower_count_train = min(len(train_fold[train_fold['sentiment'] == 'neutral']),
len(train_fold[train_fold['sentiment'] == 'negative']),
len(train_fold[train_fold['sentiment'] == 'positive']))
lower_count_valid = min(len(valid_fold[valid_fold['sentiment'] == 'neutral']),
len(valid_fold[valid_fold['sentiment'] == 'negative']),
len(valid_fold[valid_fold['sentiment'] == 'positive']))
train_fold = pd.concat([train_fold[train_fold['sentiment'] == 'neutral'].sample(n=lower_count_train, random_state=SEED),
train_fold[train_fold['sentiment'] == 'negative'].sample(n=lower_count_train, random_state=SEED),
train_fold[train_fold['sentiment'] == 'positive'].sample(n=lower_count_train, random_state=SEED),
])
valid_fold = pd.concat([valid_fold[valid_fold['sentiment'] == 'neutral'].sample(n=lower_count_valid, random_state=SEED),
valid_fold[valid_fold['sentiment'] == 'negative'].sample(n=lower_count_valid, random_state=SEED),
valid_fold[valid_fold['sentiment'] == 'positive'].sample(n=lower_count_valid, random_state=SEED),
])
train_fold = shuffle(train_fold, random_state=SEED).reset_index(drop=True)
valid_fold = shuffle(valid_fold, random_state=SEED).reset_index(drop=True)
x_train, x_train_aux, x_train_aux_2, y_train, y_train_mask, y_train_aux = get_data(train_fold, tokenizer, MAX_LEN, preprocess_fn=preprocess_roberta)
x_valid, x_valid_aux, x_valid_aux_2, y_valid, y_valid_mask, y_valid_aux = get_data(valid_fold, tokenizer, MAX_LEN, preprocess_fn=preprocess_roberta)
np.save(base_path + 'x_train', np.array(x_train))
np.save(base_path + 'y_train', np.array(y_train))
np.save(base_path + 'x_valid', np.array(x_valid))
np.save(base_path + 'y_valid', np.array(y_valid))
np.save(base_path + 'x_train_aux', np.array(x_train_aux))
np.save(base_path + 'x_train_aux2', np.array(x_train_aux_2))
np.save(base_path + 'y_train_mask', np.array(y_train_mask))
np.save(base_path + 'y_train_aux', np.array(y_train_aux))
np.save(base_path + 'x_valid_aux', np.array(x_valid_aux))
np.save(base_path + 'x_valid_aux2', np.array(x_valid_aux_2))
np.save(base_path + 'y_valid_mask', np.array(y_valid_mask))
np.save(base_path + 'y_valid_aux', np.array(y_valid_aux))
# Compress logs dir
!tar -czf balanced_fold_1.tar.gz balanced_fold_1
!tar -czf balanced_fold_2.tar.gz balanced_fold_2
!tar -czf balanced_fold_3.tar.gz balanced_fold_3
!tar -czf balanced_fold_4.tar.gz balanced_fold_4
!tar -czf balanced_fold_5.tar.gz balanced_fold_5
# Delete logs dir
shutil.rmtree('balanced_fold_1')
shutil.rmtree('balanced_fold_2')
shutil.rmtree('balanced_fold_3')
shutil.rmtree('balanced_fold_4')
shutil.rmtree('balanced_fold_5')
```
## Tokenizer sanity check (no QA)
```
for idx in range(5):
print('\nRow %d' % idx)
max_seq_len = 32
text = train_df['text'].values[idx]
selected_text = train_df['selected_text'].values[idx]
_, (target_start, target_end, _) = preprocess_text.preprocess_roberta(' ' + text, selected_text, tokenizer, max_seq_len)
decoded_text = preprocess_text.decode(target_start.argmax(), target_end.argmax(), text, tokenizer)
print('text : "%s"' % text)
print('selected_text: "%s"' % selected_text)
print('decoded_text : "%s"' % decoded_text)
assert selected_text == decoded_text
```
## Data generation sanity check (no QA)
```
for idx in range(5):
print('\nRow %d' % idx)
max_seq_len = 24
text = train_df['text'].values[idx]
selected_text = train_df['selected_text'].values[idx]
jaccard = train_df['jaccard'].values[idx]
selected_text_wordCnt = train_df['selected_text_wordCnt'].values[idx]
x_train, x_train_aux, x_train_aux_2, y_train, y_train_mask, y_train_aux = preprocess_text.get_data(train_df[idx:idx+1], tokenizer, max_seq_len,
preprocess_fn=preprocess_text.preprocess_roberta)
print('text : "%s"' % text)
print('jaccard : "%.4f"' % jaccard)
print('sentiment : "%s"' % question)
print('word count : "%d"' % selected_text_wordCnt)
print('input_ids : "%s"' % x_train[0][0])
print('attention_mask: "%s"' % x_train[1][0])
print('sentiment : "%d"' % x_train_aux[0])
print('sentiment OHE : "%s"' % x_train_aux_2[0])
print('selected_text : "%s"' % selected_text)
print('start : "%s"' % y_train[0][0])
print('end : "%s"' % y_train[1][0])
print('mask : "%s"' % y_train_mask[0])
print('jaccard : "%.4f"' % y_train_aux[0][0])
print('word count : "%d"' % y_train_aux[1][0])
assert len(x_train) == 2
assert len(x_train_aux) == 1
assert len(x_train_aux_2) == 1
assert len(y_train) == 2
assert len(y_train_mask) == 1
assert len(y_train_aux) == 3
assert len(x_train[0][0]) == len(x_train[1][0]) == max_seq_len
assert len(y_train[0][0]) == len(y_train[1][0]) == len(y_train_mask[0]) == max_seq_len
```
# Output 5-fold set (no QA)
```
for fold_n in range(folds.n_splits):
fold_n += 1
base_path = 'no_qa_fold_%d/' % fold_n
# Create dir
os.makedirs(base_path)
x_train, x_train_aux, x_train_aux_2, y_train, y_train_mask, y_train_aux = preprocess_text.get_data(train_df[train_df[('fold_%s' % fold_n)] == 'train'], tokenizer,
MAX_LEN, preprocess_fn=preprocess_text.preprocess_roberta)
x_valid, x_valid_aux, x_valid_aux_2, y_valid, y_valid_mask, y_valid_aux = preprocess_text.get_data(train_df[train_df[('fold_%s' % fold_n)] == 'validation'], tokenizer,
MAX_LEN, preprocess_fn=preprocess_text.preprocess_roberta)
np.save(base_path + 'x_train', np.array(x_train))
np.save(base_path + 'y_train', np.array(y_train))
np.save(base_path + 'x_valid', np.array(x_valid))
np.save(base_path + 'y_valid', np.array(y_valid))
np.save(base_path + 'x_train_aux', np.array(x_train_aux))
np.save(base_path + 'x_train_aux2', np.array(x_train_aux_2))
np.save(base_path + 'y_train_mask', np.array(y_train_mask))
np.save(base_path + 'y_train_aux', np.array(y_train_aux))
np.save(base_path + 'x_valid_aux', np.array(x_valid_aux))
np.save(base_path + 'x_valid_aux2', np.array(x_valid_aux_2))
np.save(base_path + 'y_valid_mask', np.array(y_valid_mask))
np.save(base_path + 'y_valid_aux', np.array(y_valid_aux))
# Compress logs dir
!tar -czf no_qa_fold_1.tar.gz no_qa_fold_1
!tar -czf no_qa_fold_2.tar.gz no_qa_fold_2
!tar -czf no_qa_fold_3.tar.gz no_qa_fold_3
!tar -czf no_qa_fold_4.tar.gz no_qa_fold_4
!tar -czf no_qa_fold_5.tar.gz no_qa_fold_5
# Delete logs dir
shutil.rmtree('no_qa_fold_1')
shutil.rmtree('no_qa_fold_2')
shutil.rmtree('no_qa_fold_3')
shutil.rmtree('no_qa_fold_4')
shutil.rmtree('no_qa_fold_5')
```
# Test set EDA
```
test_df = pd.read_csv('/kaggle/input/tweet-sentiment-extraction/test.csv')
# pre-process
test_df["text"] = test_df["text"].apply(lambda x: x.strip())
test_df["text"] = test_df["text"].apply(lambda x: x.lower())
test_df['text_len'] = test_df['text'].apply(lambda x : len(x))
test_df['text_wordCnt'] = test_df['text'].apply(lambda x : len(x.split(' ')))
test_df['text_tokenCnt'] = test_df['text'].apply(lambda x : len(tokenizer.encode(x).ids))
print('Test samples: %s' % len(test_df))
display(test_df.head())
display(test_df.describe())
```
## Sentiment distribution
```
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(20, 8.7))
sns.countplot(x="sentiment", data=train_df, palette="GnBu_d", order=sentiment_cols, ax=ax1).set_title('Train')
sns.countplot(x="sentiment", data=test_df, palette="GnBu_d", order=sentiment_cols, ax=ax2).set_title('Test')
sns.despine()
plt.show()
```
## Word count distribution
```
fig, (ax1, ax2) = plt.subplots(2, 1, figsize=(20, 8.7), sharex=True)
sns.distplot(train_df['text_wordCnt'], ax=ax1).set_title("Train")
sns.distplot(test_df['text_wordCnt'], ax=ax2).set_title("Test")
sns.despine()
plt.show()
```
## Token count distribution
```
fig, (ax1, ax2) = plt.subplots(2, 1, figsize=(20, 8.7), sharex=True)
sns.distplot(train_df['text_tokenCnt'], ax=ax1).set_title("Train")
sns.distplot(test_df['text_tokenCnt'], ax=ax2).set_title("Test")
sns.despine()
plt.show()
```
# Original set
```
original_df = pd.read_csv("https://raw.githubusercontent.com/Galanopoulog/DATA607-Project-4/master/TextEmotion.csv")
# pre-process
original_df.dropna(inplace=True)
original_df = original_df.reset_index()
original_df.drop('index', axis=1, inplace=True)
original_df['content'] = original_df['content'].apply(lambda x: x.strip())
original_df['content'] = original_df['content'].apply(lambda x: x.lower())
original_df['text_len'] = original_df['content'].apply(lambda x : len(x))
original_df['text_wordCnt'] = original_df['content'].apply(lambda x : len(x.split(' ')))
original_df['text_tokenCnt'] = original_df['content'].apply(lambda x : len(tokenizer.encode(x).ids))
display(original_df.head())
display(original_df.describe())
fig, ax = plt.subplots(1, 1, figsize=(20, 6), sharex=True)
sns.distplot(original_df['text_wordCnt'], ax=ax).set_title("Train")
sns.despine()
plt.show()
fig, ax = plt.subplots(1, 1, figsize=(20, 6), sharex=True)
sns.distplot(original_df['text_tokenCnt'], ax=ax).set_title("Train")
sns.despine()
plt.show()
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import re
from transformers import pipeline, DebertaV2Tokenizer, DebertaV2ForQuestionAnswering
import torch
# ls local_data/unlabeled_contracts/2020
filepath = 'local_data/unlabeled_contracts/2020/000019119.txt'
with open(filepath, 'r', encoding="utf-8") as file:
contract = file.read().replace('\n', '')
contract
# Clean up and pre-process the text.
def pre_process_text(text):
# Simple replacement for "\n"
text = text.replace("\n", " ")
# Simple replacement for "\xa0"
text = text.replace("\xa0", " ")
# Simple replacement for "\x0c"
text = text.replace("\x0c", " ")
# Get rid of multiple dots
regex = "\ \.\ "
subst = "."
text = re.sub(regex, subst, text, 0)
# Get rid of underscores
regex = "_"
subst = " "
text = re.sub(regex, subst, text, 0)
# Get rid of multiple dashes
regex = "--+"
subst = " "
text = re.sub(regex, subst, text, 0)
# Get rid of multiple stars
regex = "\*+"
subst = "*"
text = re.sub(regex, subst, text, 0)
# Get rid of multiple whitespace
regex = "\ +"
subst = " "
text = re.sub(regex, subst, text, 0)
#Strip leading and trailing whitespace
text = text.strip()
return text
def get_questions_from_csv():
df = pd.read_csv("./data/category_descriptions.csv")
q_dict = {}
for i in range(df.shape[0]):
category = df.iloc[i, 0].split("Category: ")[1]
description = df.iloc[i, 1].split("Description: ")[1]
q_dict[category.title()] = description
return q_dict
qtype_dict = get_questions_from_csv()
labels = [l for l in qtype_dict.keys()]
questions = [q for q in qtype_dict.values()]
# labels, questions
context = pre_process_text(contract)
# qapipe = pipeline('question-answering', model='./models/deberta-v2-xlarge', tokenizer='./models/deberta-v2-xlarge')
'''
for idx, question in enumerate(questions):
answer = qapipe(question=question, context=context)
print(f'{labels[idx]}: {answer["answer"]}')
'''
tokenizer = DebertaV2Tokenizer.from_pretrained('./models/deberta-v2-xlarge')
model = DebertaV2ForQuestionAnswering.from_pretrained('./models/deberta-v2-xlarge')
def get_answers(questions, context):
answers = []
for question in questions:
print(question)
inputs = tokenizer(question, context, padding='max_length', truncation='only_second', return_tensors='pt')
input_ids = inputs['input_ids'].tolist()[0]
outputs = model(**inputs)
answer_start_scores = outputs.start_logits
answer_end_scores = outputs.end_logits
# Get the most likely beginning of answer with the argmax of the score
answer_start = torch.argmax(answer_start_scores)
# Get the most likely end of answer with the argmax of the score
answer_end = torch.argmax(answer_end_scores) + 1
answer = tokenizer.convert_tokens_to_string(tokenizer.convert_ids_to_tokens(input_ids[answer_start:answer_end]))
answers.append(answer)
print(answer)
return answers
# answers = get_answers(questions, context)
```
| github_jupyter |
# Python для анализа данных
## Pandas (join и merge)
*Ян Пиле, НИУ ВШЭ*
Давайте представим, что мы аналитики в компании, которая производит и продает скрепки. Нам нужно составить несколько отчетов для отдела продаж, чтобы посмотреть, как у них дела. Данные лежат в следующих словарях:
```
import numpy as np
import pandas as pd
# Dataframe of number of sales made by an employee
sales = {'Tony': 103,
'Sally': 202,
'Randy': 380,
'Ellen': 101,
'Fred': 82
}
# Dataframe of all employees and the region they work in
region = {'Tony': 'West',
'Sally': 'South',
'Carl': 'West',
'Archie': 'North',
'Randy': 'East',
'Ellen': 'South',
'Fred': np.nan,
'Mo': 'East',
'HanWei': np.nan,
}
```
Мы можем создать два отдельных dataframe из словарей
```
# Make dataframes
sales_df = pd.DataFrame.from_dict(sales,
orient='index',
columns=['sales'])
region_df = pd.DataFrame.from_dict(region,
orient='index',
columns=['region'])
sales_df
region_df
```
Теперь давайте объединим все наши данные в один Датафрейм. Но как нам это сделать?
Датафреймы Pandas имеют много SQL-подобных функций. Иногда не до конца понятно, использовать join или merge. Их часто используют взаимозаменяемо (выбирая то, что пришло в голову первым). Так когда же мы должны использовать каждый из этих методов, и насколько точно они отличаются друг от друга? Попробуем разобраться.
### Join
Давайте начнем с join, потому что он самый простой. У датафреймов данных есть параметр index. Это ключ вашей таблицы, и если мы знаем индекс, то мы можем легко получить строку, содержащую наши данные, используя .loc. Если вы напечатаете свой датафрейм, вы увидите индекс в крайнем левом столбце. Еще его можно получить, напрямую использовав .index:
```
sales_df.index
```
Значит индекс в sales_df - это имя продавца. Кстати, в отличие от первичного ключа таблицы SQL, индекс датафрейма не обязательно должен быть уникальным. Но уникальный индекс делает нашу жизнь проще, а время, необходимое для поиска в нашем датафрейме, короче. Учитывая индекс, мы можем найти данные строки так:
```
sales_df.loc['Tony']
```
Вернемся к join'ам. Метод join берет два датафрейма и соединяет их по индексам (технически вы можете выбрать столбец для объединения для левого датафрейма). Давайте посмотрим, что происходит, если мы объединим(aka сджойним) наши два датафрейма с помощью метода join:
```
joined_df = region_df.join(sales_df, how='left')
joined_df
```
Результат выглядит как результат SQL-join'a (по сути, это почти то же самое).
Метод join использует индекс или указанный столбец из левого датафрейма в качестве ключа джойна. Таким образом, столбец, по которому мы джойним левый датафрейм, не обязательно должен быть его индексом. Но вот для правого датафрейма ключ джойна должен быть его индексом ОБЯЗАТЕЛЬНО.
Лично мне проще воспринимать метод join как объединение на основе индекса и использовать merge, если не хочется привязываться к индексам.
В объединенном датафрейме есть несколько NaN. Так произошло , потому что не у всех сотрудников были продажи. Те, у кого не было продаж, отсутствуют в sales_df, но мы по-прежнему отображаем их, потому что мы выполнили left join (указав «how = left»), которое возвращает все строки из левого датафрейма region_df, независимо от того, есть ли совпадение в правом. Если мы не хотим отображать какие-либо NaN в нашем результате соединения, мы вместо этого можем сделать inner join (указав «how = inner»).
### Merge
На базовом уровне Merge делает более или менее то же самое, что и join. Оба метода используются для объединения двух датафреймов, но merge является более универсальным за счет более подробного описания входных данных. Давайте посмотрим, как мы можем создать тот же объединенный датафрейм с помощью merge:
```
joined_df_merge = region_df.merge(sales_df, how='left',
left_index=True,
right_index=True)
print(joined_df_merge)
joined_df_merge.reset_index(inplace=True)
joined_df_merge.rename({'index':'name'},inplace=True)
joined_df_merge['name'] = joined_df_merge['index']
joined_df_merge
import pandasql as ps
query = """
select a.name as name, a.region as region,
case when a.sales is null then 0 else a.sales end as sales,
b.sales as sales_region,
coalesce((case when a.sales is null then 0 else a.sales end)/b.sales, 0) as sales_percent
from joined_df_merge as a
left join grouped_df as b
on a.region = b.region
where a.region is not null
"""
tst = ps.sqldf(query, locals())
tst
```
Merge полезен, когда мы не хотим привязываться к индексам. Скажем, мы хотим знать, сколько, в процентном отношении, каждый сотрудник внес в продажи в своем регионе. Мы можем использовать groupby для суммирования всех продаж в каждом уникальном регионе. В приведенном ниже коде reset_index используется, чтобы превратить регион из индекса в обычную колонку.
```
grouped_df = joined_df_merge.groupby(by='region').sum()
grouped_df
grouped_df.reset_index(inplace=True)
print(grouped_df)
```
Теперь остается смержить join_df_merge с grouped_df, используя столбец region. Мы должны указать суффикс, потому что оба наших блока данных (которые мы объединяем) содержат столбец с названием sales. Входные суффиксы добавляют указанные строки к меткам столбцов с одинаковыми именами в обоих датафреймах (это удобно, потому что ничего не перепутается). В нашем случае, поскольку столбец продаж второго датафрейма фактически отражает продажи во всем регионе, мы можем добавить суффикс «_region».
```
employee_contrib = joined_df_merge.merge(grouped_df, how='left',
left_on='region',
right_on='region',
suffixes=('','_region')
)
print(employee_contrib)
```
ЭЭЭЭЭ, куда индекс исчез?
Используем set_index, чтобы вернуть его! (иначе мы не узнаем, какому сотруднику соответствует какая строка):
```
employee_contrib = employee_contrib.set_index(joined_df_merge.index)
print(employee_contrib)
```
Теперь у нас есть исходный столбец продаж и новый столбец sales_region, в котором указывается общий объем продаж в регионе. Давайте посчитаем процент продаж каждого сотрудника, а затем очистим наш датафрейм, отбросив наблюдения, без региона (Fred и HanWei), и заполняя NaN в столбце продаж нулями.
```
# Drop NAs in region column
employee_contrib = employee_contrib.dropna(subset=['region'])
# Fill NAs in sales column with 0
employee_contrib = employee_contrib.fillna({'sales': 0})
employee_contrib['%_of_sales'] = employee_contrib['sales']/employee_contrib['sales_region']
print(employee_contrib[['region','sales','%_of_sales']].sort_values(by=['region','%_of_sales']))
```
| github_jupyter |
## Train/Evaluate a DNN for AMC
This notebook demonstrates how to create a neural network and a trainer in PyTorch to learn a signal classification task. The reference dataset used is the RML2016.10A dataset for Automatic Modulation Classification
### Assumptions
- The dataset wrangling has already been completed (and is provided here)
- The classifier evaluation code (and the plotting) has already been completed
### Components Recreated in Tutorial
- Deep Neural Network Model defined in PyTorch
- Training Loop that trains for *n* epochs
### See Also
The code in this tutorial is a stripped down version of the code in ``rfml.nn.model.CNN`` and ``rfml.nn.train.StandardTrainingStrategy`` that simplifies discussion. Further detail can be provided by directly browsing the source files for those classes.
## Install the library code and dependencies
```
# Install the library code
#!pip install git+https://github.com/brysef/rfml.git@1.0
# IPython Includes (just for documentation)
from IPython.display import Image
# Ensure that the least loaded GPU is used
import setGPU
# Plotting Includes
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style("whitegrid")
# External Includes
import numpy as np
from pprint import pprint
from torch.autograd import Variable
import torch.nn as nn
from torch.optim import Adam
from torch.utils.data import DataLoader
# Internal Includes
from rfml.data import Dataset, Encoder
from rfml.data.converters import load_RML201610A_dataset
from rfml.nbutils import plot_acc_vs_snr, plot_confusion, plot_convergence, plot_IQ
from rfml.nn.eval import compute_accuracy, compute_accuracy_on_cross_sections, compute_confusion
from rfml.nn.model import Model
```
## Configuration
```
gpu = True # Set to True to use a GPU for training
fig_dir = None # Set to a file path if you'd like to save the plots generated
data_path = None # Set to a file path if you've downloaded RML2016.10A locally
```
### Loading a Dataset
The dataset used is downloaded from DeepSig Inc. and provided under a Creative Commons lic
```
dataset = load_RML201610A_dataset(path=data_path)
print(len(dataset))
pprint(dataset.get_examples_per_class())
train, test = dataset.split(frac=0.3, on=["Modulation", "SNR"])
train, val = train.split(frac=0.05, on=["Modulation", "SNR"])
print("Training Examples")
print("=================")
pprint(train.get_examples_per_class())
print("=================")
print()
print("Validation Examples")
print("=================")
pprint(val.get_examples_per_class())
print("=================")
print()
print("Testing Examples")
print("=================")
pprint(test.get_examples_per_class())
print("=================")
le = Encoder(["WBFM",
"AM-DSB",
"AM-SSB",
"CPFSK",
"GFSK",
"BPSK",
"QPSK",
"8PSK",
"PAM4",
"QAM16",
"QAM64"],
label_name="Modulation")
print(le)
# Plot a sample of the data
# You can choose a different sample by changing
idx = 10
snr = 18.0
modulation = "8PSK"
mask = (dataset.df["SNR"] == snr) & (dataset.df["Modulation"] == modulation)
sample = dataset.as_numpy(mask=mask, le=le)[0][idx,0,:]
t = np.arange(sample.shape[1])
title = "{modulation} Sample at {snr:.0f} dB SNR".format(modulation=modulation, snr=snr)
fig = plot_IQ(iq=sample, title=title)
if fig_dir is not None:
file_path = "{fig_dir}/{modulation}_{snr:.0f}dB_sample.pdf".format(fig_dir=fig_dir,
modulation=modulation,
snr=snr)
print("Saving Figure -> {file_path}".format(file_path=file_path))
fig.savefig(file_path, format="pdf", transparent=True)
plt.show()
```
### Creating a Neural Network Model
We are going to recreate a Convolutional Neural Network (CNN) based on the "VT_CNN2" Architecture. This network is based off of a network for modulation classification first
introduced in [O'Shea et al.] and later updated by [West/Oshea] and [Hauser et al.]
to have larger filter sizes.
#### Citations
##### O'Shea et al.
T. J. O'Shea, J. Corgan, and T. C. Clancy, “Convolutional radio modulation recognition networks,” in International Conference on Engineering Applications of Neural Networks, pp. 213–226, Springer,2016.
##### West/O'Shea
N. E. West and T. O’Shea, “Deep architectures for modulation recognition,” in IEEE International Symposium on Dynamic Spectrum Access Networks (DySPAN), pp. 1–6, IEEE, 2017.
##### Hauser et al.
S. C. Hauser, W. C. Headley, and A. J. Michaels, “Signal detection effects on deep neural networks utilizing raw iq for modulation classification,” in Military Communications Conference, pp. 121–127, IEEE, 2017.
```
Image(filename="_fig/CNNDiagramSmall.png")
class MyCNN(Model):
def __init__(self, input_samples: int, n_classes: int):
super().__init__(input_samples=input_samples, n_classes=n_classes)
# Batch x 1-channel x IQ x input_samples
# Modifying the first convolutional layer to not use a bias term is a
# modification made by Bryse Flowers due to the observation of vanishing
# gradients during training when ported to PyTorch (other authors used
# Keras).
self.conv1 = nn.Conv2d(
in_channels=1,
out_channels=256,
kernel_size=(1, 7),
padding=(0, 3),
bias=False,
)
self.a1 = nn.ReLU()
self.n1 = nn.BatchNorm2d(256)
self.conv2 = nn.Conv2d(
in_channels=256,
out_channels=80,
kernel_size=(2, 7),
padding=(0, 3),
bias=True,
)
self.a2 = nn.ReLU()
self.n2 = nn.BatchNorm2d(80)
# Batch x Features
self.dense1 = nn.Linear(80 * 1 * input_samples, 256)
self.a3 = nn.ReLU()
self.n3 = nn.BatchNorm1d(256)
self.dense2 = nn.Linear(256, n_classes)
def forward(self, x):
x = self.conv1(x)
x = self.a1(x)
x = self.n1(x)
x = self.conv2(x)
x = self.a2(x)
x = self.n2(x)
# Flatten the input layer down to 1-d by using Tensor operations
x = x.contiguous()
x = x.view(x.size()[0], -1)
x = self.dense1(x)
x = self.a3(x)
x = self.n3(x)
x = self.dense2(x)
return x
model = MyCNN(input_samples=128, n_classes=11)
print(model)
```
### Implementing a Training Loop
```
class MyTrainingStrategy(object):
def __init__(self, lr: float = 10e-4, n_epochs: int = 3, gpu: bool = True):
self.lr = lr
self.n_epochs = n_epochs
self.gpu = gpu
def __repr__(self):
ret = self.__class__.__name__
ret += "(lr={}, n_epochs={}, gpu={})".format(self.lr, self.n_epochs, self.gpu)
return ret
def __call__(
self, model: nn.Module, training: Dataset, validation: Dataset, le: Encoder
):
criterion = nn.CrossEntropyLoss()
if self.gpu:
model.cuda()
criterion.cuda()
optimizer = Adam(model.parameters(), lr=self.lr)
train_data = DataLoader(
training.as_torch(le=le), shuffle=True, batch_size=512
)
val_data = DataLoader(
validation.as_torch(le=le), shuffle=True, batch_size=512
)
# Save two lists for plotting a convergence graph at the end
ret_train_loss = list()
ret_val_loss = list()
for epoch in range(self.n_epochs):
train_loss = self._train_one_epoch(
model=model, data=train_data, loss_fn=criterion, optimizer=optimizer
)
print("On Epoch {} the training loss was {}".format(epoch, train_loss))
ret_train_loss.append(train_loss)
val_loss = self._validate_once(
model=model, data=val_data, loss_fn=criterion
)
print("---- validation loss was {}".format(val_loss))
ret_val_loss.append(val_loss)
return ret_train_loss, ret_val_loss
def _train_one_epoch(
self, model: nn.Module, data: DataLoader, loss_fn: nn.CrossEntropyLoss, optimizer: Adam
) -> float:
total_loss = 0.0
# Switch the model mode so it remembers gradients, induces dropout, etc.
model.train()
for i, batch in enumerate(data):
x, y = batch
# Push data to GPU if necessary
if self.gpu:
x = Variable(x.cuda())
y = Variable(y.cuda())
else:
x = Variable(x)
y = Variable(y)
# Forward pass of prediction
outputs = model(x)
# Zero out the parameter gradients, because they are cumulative,
# compute loss, compute gradients (backward), update weights
loss = loss_fn(outputs, y)
optimizer.zero_grad()
loss.backward()
optimizer.step()
total_loss += loss.item()
mean_loss = total_loss / (i + 1.0)
return mean_loss
def _validate_once(
self, model: nn.Module, data: DataLoader, loss_fn: nn.CrossEntropyLoss
) -> float:
total_loss = 0.0
# Switch the model back to test mode (so that batch norm/dropout doesn't
# take effect)
model.eval()
for i, batch in enumerate(data):
x, y = batch
if self.gpu:
x = x.cuda()
y = y.cuda()
outputs = model(x)
loss = loss_fn(outputs, y)
total_loss += loss.item()
mean_loss = total_loss / (i + 1.0)
return mean_loss
trainer = MyTrainingStrategy(gpu=gpu)
print(trainer)
```
### Putting it All Together
```
train_loss, val_loss = trainer(model=model,
training=train,
validation=val,
le=le)
title = "Training Results of {model_name} on {dataset_name}".format(model_name="MyCNN", dataset_name="RML2016.10A")
fig = plot_convergence(train_loss=train_loss, val_loss=val_loss, title=title)
if fig_dir is not None:
file_path = "{fig_dir}/training_loss.pdf"
print("Saving Figure -> {file_path}".format(file_path=file_path))
fig.savefig(file_path, format="pdf", transparent=True)
plt.show()
```
### Testing the Trained Model
```
acc = compute_accuracy(model=model, data=test, le=le)
print("Overall Testing Accuracy: {:.4f}".format(acc))
acc_vs_snr, snr = compute_accuracy_on_cross_sections(model=model,
data=test,
le=le,
column="SNR")
title = "Accuracy vs SNR of {model_name} on {dataset_name}".format(model_name="MyCNN", dataset_name="RML2016.10A")
fig = plot_acc_vs_snr(acc_vs_snr=acc_vs_snr, snr=snr, title=title)
if fig_dir is not None:
file_path = "{fig_dir}/acc_vs_snr.pdf"
print("Saving Figure -> {file_path}".format(file_path=file_path))
fig.savefig(file_path, format="pdf", transparent=True)
plt.show()
cmn = compute_confusion(model=model, data=test, le=le)
title = "Confusion Matrix of {model_name} on {dataset_name}".format(model_name="MyCNN", dataset_name="RML2016.10A")
fig = plot_confusion(cm=cmn, labels=le.labels, title=title)
if fig_dir is not None:
file_path = "{fig_dir}/confusion_matrix.pdf"
print("Saving Figure -> {file_path}".format(file_path=file_path))
fig.savefig(file_path, format="pdf", transparent=True)
plt.show()
```
| github_jupyter |
```
# Import packages
import numpy as np
import pandas as pd
import sys
from datetime import datetime
sys.path.append('../..')
import pickle
import h5py
from keras import optimizers
from keras.models import Sequential
from keras.layers import Dense, Dropout, LSTM, MaxPooling2D, Conv2D
from keras.callbacks import ModelCheckpoint, Callback
import matplotlib.pyplot as plt
import missingno as msno
from Classifier.data_processing import processor
# Load previously downloaded and processed market data from pickle. Option to redownload data if needed.
load_pickle = True
start_time = datetime(2016,1,1)
end_time = datetime(2018,2,25)
interval = 1440 # 1440 minutes = 1 day
if load_pickle:
with open('pickles/core_data.pickle', 'rb') as f:
data = pickle.load(f)
else:
data = processor.historical_download(start_time, end_time, interval)
with open('pickles/core_data.pickle', 'wb') as f:
pickle.dump(data, f)
```
## Data preparation and scrubbing
```
"""
Scrub non-viable data
A lot of this data is legacy stuff that was being tested for importance. In future none of this data will be
included in the dataset so scrubbing won't be needed
"""
def scrub(data):
# Drop incomplete data
try:
data.drop('Ltc_search_US', axis=1, inplace=True)
data.drop('Ltc_search_GB', axis=1, inplace=True)
data.drop('Ltc_search_FR', axis=1, inplace=True)
data.drop('Ltc_search_DE', axis=1, inplace=True)
data.drop('Ltc_search_RU', axis=1, inplace=True)
data.drop('Ltc_search_KR', axis=1, inplace=True)
data.drop('Eth_search_US', axis=1, inplace=True)
data.drop('Eth_search_GB', axis=1, inplace=True)
data.drop('Eth_search_FR', axis=1, inplace=True)
data.drop('Eth_search_DE', axis=1, inplace=True)
data.drop('Eth_search_RU', axis=1, inplace=True)
data.drop('Eth_search_KR', axis=1, inplace=True)
data.drop('Btc_search_US', axis=1, inplace=True)
data.drop('Btc_search_GB', axis=1, inplace=True)
data.drop('Btc_search_FR', axis=1, inplace=True)
data.drop('Btc_search_DE', axis=1, inplace=True)
data.drop('Btc_search_RU', axis=1, inplace=True)
data.drop('Btc_search_KR', axis=1, inplace=True)
data.drop('Etheur_gdax_low', axis=1, inplace=True)
data.drop('Etheur_gdax_high', axis=1, inplace=True)
data.drop('Etheur_gdax_open', axis=1, inplace=True)
data.drop('Etheur_gdax_close', axis=1, inplace=True)
data.drop('Etheur_gdax_vol', axis=1, inplace=True)
data.drop('Ltcusd_gdax_low', axis=1, inplace=True)
data.drop('Ltcusd_gdax_high', axis=1, inplace=True)
data.drop('Ltcusd_gdax_open', axis=1, inplace=True)
data.drop('Ltcusd_gdax_close', axis=1, inplace=True)
data.drop('Ltcusd_gdax_vol', axis=1, inplace=True)
data.drop('Ltceur_gdax_low', axis=1, inplace=True)
data.drop('Ltceur_gdax_high', axis=1, inplace=True)
data.drop('Ltceur_gdax_open', axis=1, inplace=True)
data.drop('Ltceur_gdax_close', axis=1, inplace=True)
data.drop('Ltceur_gdax_vol', axis=1, inplace=True)
except:
pass
# Testing only: drop google search trend data
try:
data.drop('Eth_search_worldwide', axis=1, inplace=True)
data.drop('Ltc_search_worldwide', axis=1, inplace=True)
data.drop('Btc_search_worldwide', axis=1, inplace=True)
except:
pass
# Testing only: drop LTC blockchain network data
try:
data.drop('Ltc_hashrate', axis=1, inplace=True)
data.drop('Ltc_addresses', axis=1, inplace=True)
data.drop('Ltc_supply', axis=1, inplace=True)
data.drop('Ltc_daily_trx', axis=1, inplace=True)
data.drop('Ltc_fee_per_trx', axis=1, inplace=True)
except:
pass
# Testing only: drop BTC blockchain network data
try:
data.drop('Btc_hashrate', axis=1, inplace=True)
data.drop('Btc_addresses', axis=1, inplace=True)
data.drop('Btc_supply', axis=1, inplace=True)
data.drop('Btc_daily_trx', axis=1, inplace=True)
data.drop('Btc_fee_per_trx', axis=1, inplace=True)
except:
pass
# Testing only: drop ETH blockchain network data
try:
data.drop('Eth_hashrate', axis=1, inplace=True)
data.drop('Eth_addresses', axis=1, inplace=True)
data.drop('Eth_supply', axis=1, inplace=True)
data.drop('Eth_daily_trx', axis=1, inplace=True)
data.drop('Eth_fee_per_trx', axis=1, inplace=True)
except:
pass
# Testing only: drop LTC-USD kraken market data
try:
data.drop('Ltcusd_kraken_open', axis=1, inplace=True)
data.drop('Ltcusd_kraken_high', axis=1, inplace=True)
data.drop('Ltcusd_kraken_low', axis=1, inplace=True)
data.drop('Ltcusd_kraken_close', axis=1, inplace=True)
data.drop('Ltcusd_kraken_vol', axis=1, inplace=True)
except:
pass
# Testing only: drop BTC-USD gdax market data
#try:
#data.drop('Btcusd_gdax_open', axis=1, inplace=True)
#data.drop('Btcusd_gdax_high', axis=1, inplace=True)
#data.drop('Btcusd_gdax_low', axis=1, inplace=True)
#data.drop('Btcusd_gdax_close', axis=1, inplace=True)
#data.drop('Btcusd_gdax_vol', axis=1, inplace=True)
#except:
# pass
# Testing only: drop LTC-EUR kraken market data
try:
data.drop('Ltceur_kraken_open', axis=1, inplace=True)
data.drop('Ltceur_kraken_high', axis=1, inplace=True)
data.drop('Ltceur_kraken_low', axis=1, inplace=True)
data.drop('Ltceur_kraken_close', axis=1, inplace=True)
data.drop('Ltceur_kraken_vol', axis=1, inplace=True)
except:
pass
# Testing only: drop BTC-EUR gdax market data
#try:
#data.drop('Btceur_gdax_open', axis=1, inplace=True)
#data.drop('Btceur_gdax_high', axis=1, inplace=True)
#data.drop('Btceur_gdax_low', axis=1, inplace=True)
#data.drop('Btceur_gdax_close', axis=1, inplace=True)
#data.drop('Btceur_gdax_vol', axis=1, inplace=True)
#except:
# pass
# Testing only: drop ETH-USD kraken market data
#try:
#data.drop('Ethusd_kraken_open', axis=1, inplace=True)
#data.drop('Ethusd_kraken_high', axis=1, inplace=True)
#data.drop('Ethusd_kraken_low', axis=1, inplace=True)
#data.drop('Ethusd_kraken_close', axis=1, inplace=True)
#data.drop('Ethusd_kraken_vol', axis=1, inplace=True)
#except:
# pass
# Testing only: drop ETH-USD gdax market data
#try:
#data.drop('Ethusd_gdax_open', axis=1, inplace=True)
#data.drop('Ethusd_gdax_high', axis=1, inplace=True)
#data.drop('Ethusd_gdax_low', axis=1, inplace=True)
#data.drop('Ethusd_gdax_close', axis=1, inplace=True)
#data.drop('Ethusd_gdax_vol', axis=1, inplace=True)
#except:
#pass
# Testing only: drop ETH-EUR kraken market data
#try:
#data.drop('Etheur_kraken_open', axis=1, inplace=True)
#data.drop('Etheur_kraken_high', axis=1, inplace=True)
#data.drop('Etheur_kraken_low', axis=1, inplace=True)
#data.drop('Etheur_kraken_close', axis=1, inplace=True)
#data.drop('Etheur_kraken_vol', axis=1, inplace=True)
#except:
#pass
#data.drop('Ethusd_kraken_close', axis=1, inplace=True)
#data.drop('Btceur_kraken_open', axis=1, inplace=True)
data = data.astype('float64')
data = data.interpolate()
return data
data = scrub(data)
"""
Data visualisation to understand where missing data and null values are. Commented out as all validation on this
dataset has been done many times already
"""
#%matplotlib inline
#with pd.option_context('display.max_rows', None, 'display.max_columns', None):
# print(data.head(1))
#msno.matrix(data)
#data.isnull().sum()
"""
Split data into training set and targets. Target variable selects the price we're looking to predict.
Forecast_range sets how many time periods / intervals out we're looking at. Currently set to 1 period of 1440 minutes i.e. 1 day
"""
target = "Btcusd_kraken_close"
forecast_range = 1
x, y, actuals = processor.generate_x_y(data, target=target, forecast_range=1)
"""
Normalise data
"""
x_mean = x.mean(axis=0)
x_std = x.std(axis=0)
x = (x - x_mean) / x_std
y_mean = y.mean()
y_std = y.std()
y = (y - y_mean) / y_std
"""
Split data into training vs validation sets
"""
train_x, valid_x = x[:-30], x[-30:]
train_y, valid_y = y[:-30], y[-30:]
actuals = actuals[-30:] # These are raw prices to use when calculating actual returns from growth rates
```
## Prediction using LSTM
```
# Reshape data from (num_samples, features) to (num_samples, sequence_length, features)
sequence_length = 4
def seq_data(data_x, data_y, seq_length):
seq_data_x = []
seq_data_y = []
for ii in range(len(data_x) - seq_length + 1):
seq_data_x.append(data_x[ii : ii + seq_length])
seq_data_y.append(data_y[ii + seq_length-1])
return np.array(seq_data_x), np.array(seq_data_y)
# Add the last x time periods from before the validation set starts so that the first datapoint for the validation data
# also has the relevant price history for predictions
valid_x_2 = np.concatenate((train_x[-sequence_length + 1:], valid_x)) # Give full sequence length to first validation datapoint
valid_y_2 = np.concatenate((train_y[-sequence_length + 1:], valid_y)) # Give full sequence length to first validation datapoint
# Convert to sequential data feed for LSTM
train_x_seq, train_y_seq = seq_data(train_x, train_y, sequence_length)
valid_x_seq, valid_y_seq = seq_data(valid_x_2, valid_y_2, sequence_length)
class LSTM_net:
"""
RNN using LSTM
"""
def __init__(self, input_size, learning_rate):
self.input_size = input_size
self.learning_rate = learning_rate
self.build_model()
def build_model(self):
self.model = Sequential()
self.model.add(LSTM(256, return_sequences=True,
input_shape=self.input_size))
#self.model.add(Dropout(0.2))
self.model.add(LSTM(256))
#self.model.add(Dropout(0.2))
self.model.add(Dense(1, activation='linear'))
# Define optimiser and compile
optimizer = optimizers.Adam(self.learning_rate)
self.model.compile(optimizer=optimizer, loss='mse', metrics=['accuracy'])
# Initialise weight saving callback
LSTM_checkpointer = ModelCheckpoint(filepath='./saved_models/LSTM_weights.hdf5',
verbose=1, save_best_only=True)
# Initialise training hyper parameters
learning_rate = 0.00001
input_size = (train_x_seq.shape[1], train_x_seq.shape[2])
epochs = 1500
batch_size = 64
# Initialise neural network
LSTM_network = LSTM_net(input_size, learning_rate)
# Start training
LSTM_network.model.fit(train_x_seq, train_y_seq,
batch_size=batch_size, epochs=epochs,
callbacks=[LSTM_checkpointer],
validation_data=(valid_x_seq, valid_y_seq))
# Load the model weights with the best validation loss.
LSTM_network.model.load_weights('saved_models/LSTM_weights.hdf5')
prediction = []
for ii in range(len(valid_x_seq)):
input_data = np.reshape(valid_x_seq[ii], (-1, valid_x_seq.shape[1], valid_x_seq.shape[2]))
model_output = LSTM_network.model.predict(input_data)
prediction.append(model_output.item() * y_std + y_mean)
predicted_price = ([x + 1 for x in prediction]) * actuals
%matplotlib inline
plt.plot(range(1, len(predicted_price)+1), predicted_price, label='Predicted price')
plt.plot(range(len(predicted_price)), actuals, label='Actual price')
plt.legend()
_ = plt.ylim()
# Simulate returns for validation period
position = 0
cash = 1000
for ii in range(len(predicted_price)):
action = ""
if prediction[ii] > 0:
position += cash / actuals[ii] * 0.997 # 0.997 to account for fees
cash = 0
action = "BUY"
if prediction[ii] < 0:
cash += position * actuals[ii] * 0.997
position = 0
action = "SELL"
print("Day {}: {}. Price expected to change from {} to {}. Portfolio value of {}".format(ii, action, actuals[ii], predicted_price[ii], position*actuals[ii] + cash))
```
Some results from playing with hyperparameters
Model 1:
LSTM layers = 2
Hidden nodes = 128
learning rate = 0.00001
batch size = 64
sequence_length = 3
epochs = 2000
val_loss = 0.75104
Portfolio value = $1908
Model 2:
LSTM layers = 2
Hidden nodes = 256
learning rate = 0.00001
batch size = 64
sequence_length = 4
epochs = 2000
val_loss = 0.71283
Portfolio value = 1908
Model 3:
Dropout 20%
LSTM layers = 2
Hidden nodes = 256
learning rate = 0.00001
batch size = 64
sequence_length = 4
epochs = 2000
val_loss = 0.82500
Portfolio value = 1908
Model 4 - Market data only:
LSTM layers = 2
Hidden nodes = 256
learning rate = 0.00001
batch size = 64
sequence_length = 4
epochs = 1250
val_loss = 0.77
Portfolio value = 2068
Model 5 - BTC Kraken market data only:
LSTM layers = 2
Hidden nodes = 256
learning rate = 0.00001
batch size = 64
sequence_length = 4
epochs = 1500
val_loss = 0.67853
Portfolio value = 2017
Model 6 - All BTC market data only:
LSTM layers = 2
Hidden nodes = 256
learning rate = 0.00001
batch size = 64
sequence_length = 4
epochs = 1500
val_loss = 0.83381
Portfolio value = 1961
Model 6 - All BTC and ETH market data only:
LSTM layers = 2
Hidden nodes = 256
learning rate = 0.00001
batch size = 64
sequence_length = 4
epochs = 1500
val_loss = 0.68054
Portfolio value = 2011
## Downloading and evaluating against new test data
```
start_time_n = datetime(2018,2,21)
end_time_n = datetime(2018,3,17)
interval = 1440
load_test = True
if load_test:
with open('pickles/test_data.pickle', 'rb') as f:
test_data = pickle.load(f)
else:
test_data = processor.historical_download(start_time_n, end_time_n, interval)
with open('pickles/test_data.pickle', 'wb') as f:
pickle.dump(test_data, f)
test_data = scrub(test_data)
test_data.drop('Etheur_gdax_low', axis=1, inplace=True)
test_data.drop('Etheur_gdax_high', axis=1, inplace=True)
test_data.drop('Etheur_gdax_open', axis=1, inplace=True)
test_data.drop('Etheur_gdax_close', axis=1, inplace=True)
test_data.drop('Etheur_gdax_vol', axis=1, inplace=True)
#%matplotlib inline
#with pd.option_context('display.max_rows', None, 'display.max_columns', None):
# print(data.head(1))
#msno.matrix(test_data)
test_x, test_y, test_actuals = processor.generate_x_y(test_data, target=target, forecast_range=1)
test_x = (test_x - x_mean) / x_std
test_y = (test_y - y_mean) / y_std
test_x_seq, test_y_seq = seq_data(test_x, test_y, sequence_length)
test_actuals_for_seq = test_actuals[sequence_length-1:]
# Load the model weights with the best validation loss.
LSTM_network.model.load_weights('saved_models/LSTM_weights.hdf5')
prediction = []
for ii in range(len(test_x_seq)):
input_data = np.reshape(test_x_seq[ii], (-1, test_x_seq.shape[1], test_x_seq.shape[2]))
model_output = LSTM_network.model.predict(input_data)
prediction.append(model_output.item() * y_std + y_mean)
predicted_price = ([x + 1 for x in prediction]) * test_actuals_for_seq
%matplotlib inline
plt.plot(range(1, len(predicted_price)+1), predicted_price, label='Predicted price')
plt.plot(range(len(predicted_price)), test_actuals_for_seq, label='Actual price')
plt.legend()
_ = plt.ylim()
# Simulate returns for validation period
position = 0
cash = 1000
for ii in range(len(predicted_price)):
action = ""
if prediction[ii] > 0:
position += cash / test_actuals_for_seq[ii] * 0.997 # 0.997 to account for fees
cash = 0
action = "BUY"
if prediction[ii] < 0:
cash += position * test_actuals_for_seq[ii] * 0.997
position = 0
action = "SELL"
print("Day {}: {}. Price expected to change from {} to {}. Portfolio value of {}".format(ii, action, test_actuals_for_seq[ii], predicted_price[ii], position*actuals[ii] + cash))
```
| github_jupyter |
# Arrow
Vaex supports [Arrow](https://arrow.apache.org). We will demonstrate vaex+arrow by giving a quick look at a large dataset that does not fit into memory. The NYC taxi dataset for the year 2015 contains about 150 million rows containing information about taxi trips in New York, and is about 23GB in size. You can download it here:
* https://docs.vaex.io/en/latest/datasets.html
In case you want to convert it to the arrow format, use the code below:
```python
ds_hdf5 = vaex.open('/Users/maartenbreddels/datasets/nytaxi/nyc_taxi2015.hdf5')
# this may take a while to export
ds_hdf5.export('./nyc_taxi2015.arrow')
```
Also make sure you install vaex-arrow:
```bash
$ pip install vaex-arrow
```
```
!ls -alh /Users/maartenbreddels/datasets/nytaxi/nyc_taxi2015.arrow
import vaex
```
## Opens instantly
Opening the file goes instantly, since nothing is being copied to memory. The data is only memory mapped, a technique that will only read the data when needed.
```
%time
df = vaex.open('/Users/maartenbreddels/datasets/nytaxi/nyc_taxi2015.arrow')
df
```
## Quick viz of 146 million rows
As can be seen, this dataset contains 146 million rows.
Using plot, we can generate a quick overview what the data contains. The pickup locations nicely outline Manhattan.
```
df.plot(df.pickup_longitude, df.pickup_latitude, f='log');
df.total_amount.minmax()
```
## Data cleansing: outliers
As can be seen from the total_amount columns (how much people payed), this dataset contains outliers. From a quick 1d plot, we can see reasonable ways to filter the data
```
df.plot1d(df.total_amount, shape=100, limits=[0, 100])
# filter the dataset
dff = df[(df.total_amount >= 0) & (df.total_amount < 100)]
```
## Shallow copies
This filtered dataset did not copy any data (otherwise it would have costed us about ~23GB of RAM). Shallow copies of the data are made instead and a booleans mask tracks which rows should be used.
```
dff['ratio'] = dff.tip_amount/dff.total_amount
```
## Virtual column
The new column `ratio` does not do any computation yet, it only stored the expression and does not waste any memory. However, the new (virtual) column can be used in calculations as if it were a normal column.
```
dff.ratio.mean()
```
## Result
Our final result, the percentage of the tip, can be easily calcualted for this large dataset, it did not require any excessive amount of memory.
## Interoperability
Since the data lives as Arrow arrays, we can pass them around to other libraries such as pandas, or even pass it to other processes.
```
arrow_table = df.to_arrow_table()
arrow_table
# Although you can 'convert' (pass the data) in to pandas,
# some memory will be wasted (at least an index will be created by pandas)
# here we just pass a subset of the data
df_pandas = df[:10000].to_pandas_df()
df_pandas
```
## Tutorial
If you want to learn more on vaex, take a look at the [tutorials to see what is possible](https://docs.vaex.io/en/latest/tutorial.html).
| github_jupyter |
<font size="4" style="color:red;"> **IMPORTANT: ** Only modify cells which have the following comment</font>
```python
# modify this cell
```
<font style="color:red;"> Do not add any new cells when submitting homework. For Docker users, to test out new code, use the coding **scratchpad** by clicking the triangular icon in the bottom right corner of the screen. (**hotkey:** control-B) </font>
# Exercises:
**Note: ** Make sure you have read the *What is Probability?* notebook before attempting these exercises.
In this excercise you will write code to estimate the probability that $n$ flips of a fair coin will result in number of `"heads"` between $k_1$ and $k_2$.
You should write the body of two functions:
1. <code><font color="blue">seq_sum</font>(n)</code>: generates a random sequence of coin flips and counts the number of heads.
2. <code><font color="blue">estimate_prob</font>(n,k1,k2,m)</code>: Using calls to `seq_sum`, estimate the probability of the number of heads being between $k_1$ and $k_2$.
### Notebook Setup:
The folowing magic command downloads many python packages like *numpy* and allows the notebooks to plot graphs with *matplotlib*.
<font color="red">**DO NOT**</font> import other packages. You already have all the packages you need.
```
%pylab inline
```
Specifically, you can now use `random.rand(x)` which for some $x \in N$ generates $x$ random numbers. You **will** use this command in your homework.
```
random.rand()
random.rand(4)
```
## Exercise 1:
Write a function, <code><font color ="blue">seq_sum</font>(n)</code>, which generates $n$ random coin flips from a fair coin and then returns the number of heads. A fair coin is defined to be a coin where $P($heads$)=\frac{1}{2}$
The output type should be a numpy integer, **hint:** use `random.rand()`
<font style="color:blue"> * **Code:** *</font>
```python
x = seq_sum(100)
print x
print [seq_sum(2) for x in range(20)]
```
<font style="color:magenta"> * **Output:** *</font>
```
49
[0, 1, 1, 1, 1, 2, 1, 2, 1, 1, 0, 0, 2, 1, 1, 1, 0, 0, 1, 1]
```
* Write your code for seq_sum in the cell below
```
# modify this cell
def seq_sum(n):
""" input: n, generate a sequence of n random coin flips
output: return the number of heads
Hint: For simplicity, use 1,0 to represent head,tails
"""
number_of_heads = 0;
for i in range(n):
r = random.rand();
if (r >= 0.5):
number_of_heads = number_of_heads + 1;
return number_of_heads;
```
* if the following cell runs without error you receive some points.
```
# checking function
x = seq_sum(100)
print(x)
assert unique([seq_sum(2) for x in range(0,200)]).tolist() == [0, 1, 2]
#
# AUTOGRADER TEST - DO NOT REMOVE
#
```
## Exercise 2:
Write a function, <code><font color="blue">estimate_prob</font>(n,k1,k2,m)</code>, that uses <code><font color="blue">seq_sum</font>(n)</code> to estimate the following probability:
$$ P(\; k_1 <= \text{number of heads in $n$ flips} < k_2 ) $$
The function should estimate the probability by running $m$ different trials of <code><font color="blue">seq_sum</font>(n)</code>, probably using a *`for`* loop.
In order to receive full credit **estimate_prob** <font color="red">MUST</font> call **seq_sum** (aka: seq_sum is located inside the **estimate_prob** function)
<font style="color:blue"> * **Code:** *</font>
```python
x = estimate_prob(100,45,55,1000)
print(x)
print type(x)
```
<font style="color:magenta"> * **Output:** *</font>
```
0.686
<type 'float'>
```
```
# Modify this cell
def estimate_prob(n,k1,k2,m):
"""Estimate the probability that n flips of a fair coin result in k1 to k2 heads
n: the number of coin flips (length of the sequence)
k1,k2: the trial is successful if the number of heads is
between k1 and k2-1
m: the number of trials (number of sequences of length n)
output: the estimated probability
"""
estimation = 0;
for i in range(m):
number_of_heads = seq_sum(n);
if (k1 <= number_of_heads and number_of_heads < k2):
estimation = estimation + 1;
return estimation / m;
# this is a small sanity check
# the true check for this function is further down
x = estimate_prob(100,45,55,1000)
print(x)
assert 'float' in str(type(x))
```
### Estimate vs. True Probability
We can now check how to see how close these estimates are to the true probabilities.
### Helper Functions
These helper functions are used to calculate the actual probabilities. They are used to test your code.
It is not required that you understand how they work.
```
def calc_prob(n,k1,k2):
"""Calculate the probability using a normal approximation"""
n=float(n);k1=float(k1);k2=float(k2)
z1=(k1-0.5*n)/(sqrt(n)/2)
z2=(k2-0.5*n)/(sqrt(n)/2)
return (erf(z2/sqrt(2))-erf(z1/sqrt(2)))/2
from math import erf,sqrt
def evaluate(n,q1,q2,m,r=100):
"""Run calc_range many times and test whether the estimates are consistent with calc_prob"""
k1=int(q1*n)
k2=int(q2*n)
p=calc_prob(n,k1,k2)
std=sqrt(p*(1-p)/m)
print('computed prob=%5.3f, std=%5.3f'%(p,std))
L=[estimate_prob(n,k1,k2,m) for i in range(r)]
med=np.median(L)
print('ran estimator %d times, with parameters n=%d,k1=%d,k2=%d,m=%d'%(r,n,k1,k2,m))
print('median of estimates=%5.3f, error of median estimator=%5.3f, std= %f5.3'%(med,med-p,std))
return L,med,p,std,abs((med-p)/std)
def test_report_assert(n,q1,q2,m,r=100):
k1=int(q1*n)
k2=int(q2*n)
L,med,p,std,norm_err=evaluate(n,q1,q2,m,r=100)
hist(L);
plot([p,p],plt.ylim(),'r',label='true prob')
plot([med,med],plt.ylim(),'k',label='median of %d estimates'%r)
mid_y=mean(plt.ylim())
plot([p-std,p+std],[mid_y,mid_y],'g',label='+-std')
legend();
print('normalized error of median=',norm_err,'should be <1.0')
title('r=%d,n=%d,k1=%d,k2=%d,m=%d,\nnorm_err=%4.3f'%(r,n,k1,k2,m,norm_err))
assert norm_err<1.0
```
### Testing your Functions
* We now test your functions. The graphs below show how close your estimated probability is to the true probability for various values of $k_1$ and $k_2$. You can see that your answer is never exactly the correct probability.
* For full credit, the code below must run without error.
```
# checking functions
m=100
i=1
figure(figsize=[10,12])
for n in [100,1000]:
for q1,q2 in [(0.4,0.6),(0.55,1.00),(0.47,0.499)]:
fig=subplot(3,2,i)
print('#### test no.',i)
i+=1
test_report_assert(n,q1,q2,m,r=100)
tight_layout()
# checking functions
def seq_sum(n):
#Log.append(n)
s=sum(random.rand(n)>0.5)
Log.append((n,s))
return s
n,k1,k2,m = 100,45,50,1000
for r in range(10):
Log=[]
a=estimate_prob(n,k1,k2,m)
b=float(sum([(s>=k1 and s<k2) for n,s in Log]))/m
n_correct=sum(nn==100 for nn,s in Log)
assert a==b, "estimate is incorrect. should be %4f, instead is %4f"%(b,a)
assert m==len(Log), 'should call seq_sum %d times, called it %d times'%(m,len(Log))
assert m==n_correct, 'the parameter n should be %d but sometimes it was not.'%n
print("all good!")
```
## Quiz answers
```
# This one can be 0, 1 or 2.
seq_sum(2)
avg_answer = 0;
n_tests = 100;
for i in range(n_tests):
avg_answer = avg_answer + estimate_prob(100,40,60,1000);
print(avg_answer / n_tests);
print("all good!");
```
| github_jupyter |
# Training a Machine Learning algorithm
## Load data
First, load the SDK of the Data-centric design Hub to connect to your Thing
Note: like in the Python script for data collection, you need an .env file with your thing id and token.
```
from dotenv import load_dotenv
import os
from dcd.entities.thing import Thing
# The thing ID and access token
load_dotenv()
THING_ID = os.environ['THING_ID']
THING_TOKEN = os.environ['THING_TOKEN']
my_thing = Thing(thing_id=THING_ID, token=THING_TOKEN)
my_thing.read()
```
Provide the start and end dates, defining when to look for data
```
from datetime import datetime
DATE_FORMAT = '%Y-%m-%d %H:%M:%S'
START_DATE = "2019-05-16 10:53:00"
END_DATE = "2019-05-16 10:56:00"
from_ts = datetime.timestamp(datetime.strptime(START_DATE, DATE_FORMAT)) * 1000
to_ts = datetime.timestamp(datetime.strptime(END_DATE, DATE_FORMAT)) * 1000
```
Retrieve data and label
```
FSR_PROP_NAME = "FSR"
CLASS_PROP_NAME = "Sitting Posture"
fsr = my_thing.find_property_by_name(FSR_PROP_NAME)
fsr.read(from_ts, to_ts)
data = fsr.values
sitting = my_thing.find_property_by_name(CLASS_PROP_NAME)
sitting.read(from_ts, to_ts)
label = sitting.values
```
Extract classes from the CLASS property
```
classes = []
for index, clazz in enumerate(sitting.classes):
print(index, " => ", clazz['name'])
classes.append(clazz['name'])
```
# Prepare
Split the data into training data (60%), cross validation data (20%) and test data (20%)
```
train_data = []
train_label = []
cv_data = []
cv_label = []
test_data = []
test_label = []
leftover_data = []
leftover_label = []
for index in range(len(data)):
# remove time
data[index].pop(0)
label[index].pop(0)
if index%5 == 0:
# 20% to test data
test_data.append(data[index])
test_label.append(label[index])
else:
# 80% leftover data
leftover_data.append(data[index])
leftover_label.append(label[index])
for index in range(len(leftover_data)):
if index%4 == 0:
# 20% to cross validate
cv_data.append(leftover_data[index])
cv_label.append(leftover_label[index])
else:
# 60% to train
train_data.append(leftover_data[index])
train_label.append(leftover_label[index])
```
Check the distribution
```
print("nb total data: " + str(len(data)))
print("nb total labels: " + str(len(label)))
print("nb train data: " + str(len(train_data)))
print("nb train labels: " + str(len(train_label)))
print("nb cv data: " + str(len(cv_data)))
print("nb cv labels: " + str(len(cv_label)))
print("nb test data: " + str(len(test_data)))
print("nb test labels: " + str(len(test_label)))
```
# Train
We use a k-Nearest Neighbour (kNN) algorithm
```
from sklearn.neighbors import KNeighborsClassifier
neigh = KNeighborsClassifier(n_neighbors=1)
neigh.fit(train_data, train_label)
```
# Evaluate
Import evaluation functions from scikit learn
```
from sklearn.metrics import accuracy_score
from sklearn.metrics import f1_score
from sklearn.metrics import precision_score
from sklearn.metrics import recall_score
from sklearn.metrics import confusion_matrix
from sklearn.metrics import classification_report
import numpy
```
Use the test data to evaluate the algorithm
```
predicted = neigh.predict(cv_data)
cvLabel = numpy.array(cv_label)
result = accuracy_score(cvLabel, predicted)
print("cv accuracy: {}".format(result))
```
## Cross Validation accuracy > 80%
The validation passed, we can display the test performance
```
predicted = neigh.predict(test_data)
testLabel = numpy.array(test_label)
testResult = accuracy_score(testLabel, predicted)
print("test accuracy: {}".format(testResult))
```
Confusion matrix
```
confusion_matrix(testLabel, predicted)
```
Precision score
```
precision_score(testLabel, predicted, average="macro")
```
Recall score
```
recall_score(testLabel, predicted, average="macro")
```
F1 score
```
f1_score(testLabel, predicted, average="weighted")
f1_score(testLabel, predicted, average=None)
```
Finally, we can show the classification report.
```
print(classification_report(testLabel, predicted, target_names=classes))
```
## Cross Validation accuracy < 80%
The validation failed, we can display the validation performance.
Confusion matrix
```
confusion_matrix(cvLabel, predicted)
```
Precision score
```
precision_score(cvLabel, predicted, average="macro")
```
Recall score
```
recall_score(cvLabel, predicted, average="macro")
```
F1 score
```
f1_score(cvLabel, predicted, average="weighted")
f1_score(cvLabel, predicted, average=None)
```
# Save the model in a file
```
# Where to save the model to
MODEL_FILE_NAME = "model.pickle"
# import the pickle library
import io
import pickle
with io.open(MODEL_FILE_NAME, "wb") as file:
pickle.dump(neigh, file, protocol=2)
```
| github_jupyter |
# Spherical coordinates in shenfun
The Helmholtz equation is given as
$$
-\nabla^2 u + \alpha u = f.
$$
In this notebook we will solve this equation on a unitsphere, using spherical coordinates. To verify the implementation we use a spherical harmonics function as manufactured solution.
We start the implementation by importing necessary functionality from shenfun and sympy:
```
from shenfun import *
from shenfun.la import SolverGeneric1ND
import sympy as sp
```
Define spherical coordinates $(r, \theta, \phi)$
$$
\begin{align}
x &= r \sin \theta \cos \phi \\
y &= r \sin \theta \sin \phi \\
z &= r \cos \theta
\end{align}
$$
using sympy. The radius `r` will be constant `r=1`. We create the three-dimensional position vector `rv` as a function of the two new coordinates $(\theta, \phi)$.
```
r = 1
theta, phi = psi = sp.symbols('x,y', real=True, positive=True)
rv = (r*sp.sin(theta)*sp.cos(phi), r*sp.sin(theta)*sp.sin(phi), r*sp.cos(theta))
```
We define bases with the domains $\theta \in [0, \pi]$ and $\phi \in [0, 2\pi]$. Also define a tensorproductspace, test- and trialfunction. Note that the new coordinates and the position vector are fed to the `TensorProductSpace` and not the individual spaces:
```
N, M = 64, 64
L0 = FunctionSpace(N, 'L', domain=(0, np.pi))
F1 = FunctionSpace(M, 'F', dtype='D')
T = TensorProductSpace(comm, (L0, F1), coordinates=(psi, rv, sp.Q.positive(sp.sin(theta))))
v = TestFunction(T)
u = TrialFunction(T)
```
Use one spherical harmonic function as a manufactured solution
```
#sph = sp.functions.special.spherical_harmonics.Ynm
#ue = sph(6, 3, theta, phi)
ue = sp.cos(8*(sp.sin(theta)*sp.cos(phi) + sp.sin(theta)*sp.sin(phi) + sp.cos(theta)))
```
Compute the right hand side on the quadrature mesh and take the scalar product
```
alpha = 1000
g = (-div(grad(u))+alpha*u).tosympy(basis=ue, psi=psi)
gj = Array(T, buffer=g*T.coors.sg)
g_hat = Function(T)
g_hat = inner(v, gj, output_array=g_hat)
```
Note that we can use the `shenfun` operators `div` and `grad` on a trialfunction `u`, and then switch the trialfunction for a sympy function `ue`. The operators will then make use of sympy's [derivative method](https://docs.sympy.org/latest/tutorial/calculus.html#derivatives) on the function `ue`. Here `(-div(grad(u))+alpha*u)` corresponds to the equation we are trying to solve:
```
from IPython.display import Math
Math((-div(grad(u))+alpha*u).tolatex(funcname='u', symbol_names={theta: '\\theta', phi: '\\phi'}))
#Math((grad(u)).tolatex(funcname='u', symbol_names={theta: '\\theta', phi: '\\phi'}))
```
Evaluated with `u=ue` and you get the exact right hand side `f`.
Tensor product matrices that make up the Helmholtz equation are then assembled as
```
mats = inner(v, (-div(grad(u))+alpha*u)*T.coors.sg**2)
mats[0].mats[0]
```
And the linear system of equations can be solved using the generic `SolverGeneric1ND`, that can be used for any problem that only has non-periodic boundary conditions in one dimension.
```
u_hat = Function(T)
Sol1 = SolverGeneric1ND(mats)
u_hat = Sol1(g_hat, u_hat)
```
Transform back to real space and compute the error.
```
uj = u_hat.backward()
uq = Array(T, buffer=ue)
print('Error =', np.sqrt(inner(1, (uj-uq)**2)), np.linalg.norm(uj-uq))
np.linalg.norm(u_hat - u_hat.backward().forward())
import matplotlib.pyplot as plt
%matplotlib inline
plt.spy(Sol1.MM[1].diags())
raise RuntimeError
```
## Postprocessing
Since we used quite few quadrature points in solving this problem, we refine the solution for a nicer plot. Note that `refine` simply pads Functions with zeros, which gives exactly the same accuracy, but more quadrature points in real space. `u_hat` has `NxM` quadrature points, here we refine using 3 times as many points along both dimensions
```
u_hat2 = u_hat.refine([N*3, M*3])
ur = u_hat2.backward(kind='uniform')
```
The periodic solution does not contain the periodic points twice, i.e., the computational mesh contains $0$, but not $2\pi$. It looks better if we wrap the periodic dimension all around to $2\pi$, and this is achieved with
```
xx, yy, zz = u_hat2.function_space().local_cartesian_mesh(uniform=True)
xx = np.hstack([xx, xx[:, 0][:, None]])
yy = np.hstack([yy, yy[:, 0][:, None]])
zz = np.hstack([zz, zz[:, 0][:, None]])
ur = np.hstack([ur, ur[:, 0][:, None]])
```
In the end the solution is plotted using mayavi
```
from mayavi import mlab
#mlab.init_notebook()
mlab.figure(1, bgcolor=(1, 1, 1), fgcolor=(0, 0, 0), size=(400, 400))
mlab.mesh(xx, yy, zz, scalars=ur.real, colormap='jet')
mlab.show()
```
# Biharmonic equation
A biharmonic equation is given as
$$
\nabla^4 u + \alpha u = f.
$$
This equation is extremely messy in spherical coordinates. I cannot even find it posted anywhere. Nevertheless, we can solve it trivially with shenfun, and we can also see what it looks like
```
Math((div(grad(div(grad(u))))+alpha*u).tolatex(funcname='u', symbol_names={theta: '\\theta', phi: '\\phi'}))
```
Remember that this equation uses constant radius `r=1`. We now solve the equation using the same manufactured solution as for the Helmholtz equation.
```
g = (div(grad(div(grad(u))))+alpha*u).tosympy(basis=ue, psi=psi)
gj = Array(T, buffer=g)
# Take scalar product
g_hat = Function(T)
g_hat = inner(v, gj, output_array=g_hat)
mats = inner(v, div(grad(div(grad(u)))) + alpha*u)
# Solve
u_hat = Function(T)
Sol1 = SolverGeneric1ND(mats)
u_hat = Sol1(g_hat, u_hat)
# Transform back to real space.
uj = u_hat.backward()
uq = Array(T, buffer=ue)
print('Error =', np.sqrt(dx((uj-uq)**2)))
```
Want to see what the regular 3-dimensional biharmonic equation looks like in spherical coordinates? This is extremely tedious to derive by hand, but in shenfun you can get there with the following few lines of code
```
r, theta, phi = psi = sp.symbols('x,y,z', real=True, positive=True)
rv = (r*sp.sin(theta)*sp.cos(phi), r*sp.sin(theta)*sp.sin(phi), r*sp.cos(theta))
L0 = FunctionSpace(20, 'L', domain=(0, 1))
F1 = FunctionSpace(20, 'L', domain=(0, np.pi))
F2 = FunctionSpace(20, 'F', dtype='D')
T = TensorProductSpace(comm, (L0, F1, F2), coordinates=(psi, rv, sp.Q.positive(sp.sin(theta))))
p = TrialFunction(T)
Math((div(grad(div(grad(p))))).tolatex(funcname='u', symbol_names={r: 'r', theta: '\\theta', phi: '\\phi'}))
q = TestFunction(T)
A = inner(div(grad(q)), div(grad(p)), level=2)
```
I don't know if this is actually correct, because I haven't derived it by hand and I haven't seen it printed anywhere, but at least I know the Cartesian equation is correct:
```
L0 = FunctionSpace(8, 'C', domain=(0, np.pi))
F1 = FunctionSpace(8, 'F', dtype='D')
F2 = FunctionSpace(8, 'F', dtype='D')
T = TensorProductSpace(comm, (L0, F1, F2))
p = TrialFunction(T)
Math((div(grad(div(grad(p))))).tolatex(funcname='u'))
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Using TensorBoard in Notebooks
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/tensorboard/r2/tensorboard_in_notebooks"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/tensorboard/blob/master/docs/r2/tensorboard_in_notebooks.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/tensorboard/blob/master/docs/r2/tensorboard_in_notebooks.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
</table>
TensorBoard can be used directly within notebook experiences such as [Colab](https://colab.research.google.com/) and [Jupyter](https://jupyter.org/). This can be helpful for sharing results, integrating TensorBoard into existing workflows, and using TensorBoard without installing anything locally.
## Setup
Start by installing TF 2.0 and loading the TensorBoard notebook extension:
```
!pip install -q tf-nightly-2.0-preview
# Load the TensorBoard notebook extension
%load_ext tensorboard.notebook
```
Import TensorFlow, datetime, and os:
```
import tensorflow as tf
import datetime, os
```
## TensorBoard in notebooks
Download the [FashionMNIST](https://github.com/zalandoresearch/fashion-mnist) dataset and scale it:
```
fashion_mnist = tf.keras.datasets.fashion_mnist
(x_train, y_train),(x_test, y_test) = fashion_mnist.load_data()
x_train, x_test = x_train / 255.0, x_test / 255.0
```
Create a very simple model:
```
def create_model():
return tf.keras.models.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28)),
tf.keras.layers.Dense(512, activation='relu'),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(10, activation='softmax')
])
```
Train the model using Keras and the TensorBoard callback:
```
def train_model():
model = create_model()
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
logdir = os.path.join("logs", datetime.datetime.now().strftime("%Y%m%d-%H%M%S"))
tensorboard_callback = tf.keras.callbacks.TensorBoard(logdir, histogram_freq=1)
model.fit(x=x_train,
y=y_train,
epochs=5,
validation_data=(x_test, y_test),
callbacks=[tensorboard_callback])
train_model()
```
Start TensorBoard within the notebook using [magics](https://ipython.readthedocs.io/en/stable/interactive/magics.html):
```
%tensorboard --logdir logs
```
<img class="tfo-display-only-on-site" src="images/notebook_tensorboard.png?raw=1"/>
You can now view dashboards such as scalars, graphs, histograms, and others. Some dashboards are not available yet in Colab (such as the profile plugin).
The `%tensorboard` magic has exactly the same format as the TensorBoard command line invocation, but with a `%`-sign in front of it.
You can also start TensorBoard before training to monitor it in progress:
```
%tensorboard --logdir logs
```
<img class="tfo-display-only-on-site" src="images/notebook_tensorboard_two_runs.png?raw=1"/>
The same TensorBoard backend is reused by issuing the same command. If a different logs directory was chosen, a new instance of TensorBoard would be opened. Ports are managed automatically.
Start training a new model and watch TensorBoard update automatically every 30 seconds or refresh it with the button on the top right:
```
train_model()
```
You can use the `tensorboard.notebook` APIs for a bit more control:
```
from tensorboard import notebook
notebook.list() # View open TensorBoard instances
# Control TensorBoard display. If no port is provided,
# the most recently launched TensorBoard is used
notebook.display(port=6006, height=1000)
```
<img class="tfo-display-only-on-site" src="images/notebook_tensorboard_tall.png?raw=1"/>
| github_jupyter |
NASA-HTTP
Description
These two traces contain two month's worth of all HTTP requests to the NASA Kennedy Space Center WWW server in Florida.
Format
The logs are an ASCII file with one line per request, with the following columns:
host making the request. A hostname when possible, otherwise the Internet address if the name could not be looked up.
timestamp in the format "DAY MON DD HH:MM:SS YYYY", where DAY is the day of the week, MON is the name of the month, DD is the day of the month, HH:MM:SS is the time of day using a 24-hour clock, and YYYY is the year. The timezone is -0400.
request given in quotes.
HTTP reply code.
bytes in the reply.
Measurement
The first log was collected from 00:00:00 July 1, 1995 through 23:59:59 July 31, 1995, a total of 31 days. The second log was collected from 00:00:00 August 1, 1995 through 23:59:59 Agust 31, 1995, a total of 7 days. In this two week period there were 3,461,612 requests. Timestamps have 1 second resolution. Note that from 01/Aug/1995:14:52:01 until 03/Aug/1995:04:36:13 there are no accesses recorded, as the Web server was shut down, due to Hurricane Erin.
Privacy
The logs fully preserve the originating host and HTTP request. Please do not however attempt any analysis beyond general traffic patterns.
Acknowledgements
The logs was collected by Jim Dumoulin of the Kennedy Space Center, and contributed by Martin Arlitt (mfa126@cs.usask.ca) and Carey Williamson (carey@cs.usask.ca) of the University of Saskatchewan.
Publications
This is one of six data sets analyzed in an upcoming paper by
M. Arlitt and C. Williamson, entitled ``Web Server Workload Characterization: The Search for Invariants'', to appear in the proceedings of the 1996 ACM SIGMETRICS Conference on the Measurement and Modeling of Computer Systems, Philadelphia, PA, May 23-26, 1996. An extended version of this paper is available on-line; see also the DISCUS home page and the group's publications.
Related
Permission has been granted to make four of the six data sets discussed in ``Web Server Workload Characterization: The Search for Invariants'' available. The four data sets are: Calgary-HTTP , ClarkNet-HTTP , NASA-HTTP , and Saskatchewan-HTTP .
Restrictions
The traces may be freely redistributed.
Distribution
Available from the Archive in Jul 01 to Jul 31, ASCII format, 20.7 MB gzip compressed, 205.2 MB uncompressed, and Aug 04 to Aug 31, ASCII format, 21.8 MB gzip compressed, 167.8 MB uncompressed.
```
!pip install StringIO
import pandas as pd
import StringIO
data = pd.read_csv('NASA_access_log_Jul95.txt', sep=" ", header=None)
data.columns = ["host", "timestamp", "request", "HTTP reply code", "bytes in the reply"]
pd.read_csv(StringIO(NASA_access_log_Jul95.txt), sep=' ', skipfooter=1, skiprows=1, skipinitialspace=True).drop([0])
import tarfile
tar = tarfile.open("UCB-home-IP-848278026-848292426.tr.gz")
for member in tar.getmembers():
f = tar.extractfile(member)
if f is not None:
content = f.read()
from bs4 import BeautifulSoup
def html2tsv(html, index=0):
"""Parse the index'th HTML table in ``html``. Return table as a list of
tab-separated ASCII table lines"""
soup = BeautifulSoup(html)
tables = soup.findAll('table')
table = tables[index]
out = []
for row in table.findAll('tr'):
colvals = [col.text for col in row.findAll('td')]
out.append('\t'.join(colvals))
return out
from astropy.extern.six.moves.urllib import request
from astropy.io import ascii
html = request.urlopen('http://www.almhuette-raith.at/apache-log/access.log').read() # Get web page as string
table1 = html2tsv(html, 0) # Parse the first table in the web page
#table2 = html2tsv(html, 1) # Parse the second table
#table3 = html2tsv(html, 2) # Parse the third table
dat = ascii.read('http://www.almhuette-raith.at/apache-log/access.log',
format='html',
htmldict={'table_id':2}, # Get data from the second table
fill_values=('', '-1'), # Fill blank entries with -1
header_start=0, # Row 0 is header with column names
data_start=1) # Row 1 is start of table data
```
| github_jupyter |
# Data Collection
To build the dataset of products and ingredients, I scraped the most popular skincare products from Walgreen's website and Watson's website. These are the most popular drugstores in America and across Asia, respectively. I then looked up each product's ingredients on CosDNA.com, my favorite website for analyzing cosmetics and their ingredients. This was all done with VBA in excel, which, if I were to do this again, I would definitely choose python! But, you live and learn.
```
Private Sub IE_Autiomation()
Dim i As Long
Dim ie As Object
Dim doc As Object
Dim htmlname As Object
Dim ing As Range
Dim html As Object ' HTML document
Dim myLinks As Object ' Links collection
Dim myLink As Object 'Single Link
Dim result As String
Dim myURL As String 'Web Links on worksheet
Dim LastRow As Integer ' VBA execution should stop here
Dim row As Integer
Dim j As Integer
Dim el As Object
Dim objElement As Object
Dim objCollection As Object
' GO COLLECT PRODUCT NAMES FROM DRUGSTORE WEBSITES SKINCARE SECTION
' Create InternetExplorer Object
Set ie = CreateObject("InternetExplorer.Application")
Set html = CreateObject("htmlfile")
ie.Visible = True
For Each URL In Sheets(1).Range("G2:G254")
row = URL.row
' Send the form data To URL As POST binary request
If ie.Busy = False Then
ie.navigate URL.Value
End If
' Wait while IE loading...
Do While ie.Busy
Application.Wait DateAdd("s", 1, Now)
Loop
result = ie.document.body.innerHTML
'now place all the data extracted from the web page into the new html document
html.body.innerHTML = result
Set myLinks = html.getElementsByTagName("a")
' LOOK UP PRODUCT INGREDIENTS ON COSDNA.COM
' Loop through the collected links and get a specific link defined by the conditions
For Each myLink In myLinks
If Left(myLink, 15) = "about:cosmetic_" Then
Sheets(1).Range("H" & row).Value = "http://www.cosdna.com/eng/" & Right(myLink, Len(myLink) - 6)
Exit For
End If
' Go to the next link
Next myLink
Set myLinks = Nothing
' Visit each link
Set ie = CreateObject("InternetExplorer.Application")
Set html = CreateObject("htmlfile")
ie.Visible = True
For Each URL In Sheets(1).Range("G2", Cells("G", LastRow)
row = URL.row
' Send the form data To URL As POST binary request
If ie.Busy = False Then
ie.navigate URL.Value
End If
' Wait while IE loading...
Do While ie.Busy
Application.Wait DateAdd("s", 1, Now)
Loop
' Get the ingredients from the website
Set objCollection = IE.document.getElementsByTagName("input")
Set objCollection = doc.getElementsByClassName("iStuffList")
class="iStuffList"
' Put comma delimited ingredients into excel
Set ing = Cells(9, Url.Row)
For Each el In ie.document.getElementsByClassName("iStuffList")
ing.Value = ing.Value & ", " & el.innerText
Next el
err_clear:
If Err <> 0 Then
Err.Clear
Resume Next
End If
' Clean up
Set ie = Nothing
Set objElement = Nothing
Set objCollection = Nothing
Application.StatusBar = ""
End Sub
```
| github_jupyter |
# Param search
Imports
```
import logging
from tinkoff.invest.mock_services import MockedSandboxClient
from decimal import Decimal
from tinkoff.invest.strategies.moving_average.strategy_settings import (
MovingAverageStrategySettings,
)
from tinkoff.invest import CandleInterval, MoneyValue
from tinkoff.invest.strategies.moving_average.signal_executor import (
MovingAverageSignalExecutor,
)
from tinkoff.invest.strategies.moving_average.supervisor import (
MovingAverageStrategySupervisor,
)
from tinkoff.invest.strategies.moving_average.strategy_state import (
MovingAverageStrategyState,
)
from tinkoff.invest.strategies.moving_average.strategy import MovingAverageStrategy
from tinkoff.invest.strategies.moving_average.trader import MovingAverageStrategyTrader
from datetime import timedelta, datetime, timezone
from tinkoff.invest.typedefs import ShareId, AccountId
from tinkoff.invest.strategies.base.account_manager import AccountManager
from tinkoff.invest.strategies.moving_average.plotter import (
MovingAverageStrategyPlotter,
)
logging.basicConfig(format="%(asctime)s %(levelname)s:%(message)s", level=logging.INFO)
logger = logging.getLogger(__name__)
```
Setup
```
token =
```
Settings
```
figi = ShareId("BBG0013HGFT4")
account_id = AccountId("1337007228")
settings = MovingAverageStrategySettings(
share_id=figi,
account_id=account_id,
max_transaction_price=Decimal(10000),
candle_interval=CandleInterval.CANDLE_INTERVAL_1_MIN,
long_period=timedelta(minutes=100),
short_period=timedelta(minutes=50),
std_period=timedelta(minutes=30),
)
```
Stocks for date
```
def start_datetime() -> datetime:
return datetime(year=2022, month=2, day=16, hour=17, tzinfo=timezone.utc)
real_market_data_test_from = start_datetime() - timedelta(days=1)
real_market_data_test_start = start_datetime()
real_market_data_test_end = start_datetime() + timedelta(days=3)
```
Initial balance
```
balance = MoneyValue(currency="rub", units=20050, nano=690000000)
```
Trader
```
with MockedSandboxClient(
token=token,
balance=balance,
) as mocked_services:
account_manager = AccountManager(
services=mocked_services, strategy_settings=settings
)
state = MovingAverageStrategyState()
strategy = MovingAverageStrategy(
settings=settings,
account_manager=account_manager,
state=state,
)
supervisor = MovingAverageStrategySupervisor()
signal_executor = MovingAverageSignalExecutor(
services=mocked_services,
state=state,
settings=settings,
)
moving_average_strategy_trader = MovingAverageStrategyTrader(
strategy=strategy,
settings=settings,
services=mocked_services,
state=state,
signal_executor=signal_executor,
account_manager=account_manager,
supervisor=supervisor,
)
plotter = MovingAverageStrategyPlotter(settings=settings)
initial_balance = account_manager.get_current_balance()
for i in range(50):
logger.info("Trade %s", i)
events = list(supervisor.get_events())
plotter.plot(events)
try:
moving_average_strategy_trader.trade()
except Exception:
pass
current_balance = account_manager.get_current_balance()
assert initial_balance != current_balance
logger.info("Initial balance %s", initial_balance)
logger.info("Current balance %s", current_balance)
```
| github_jupyter |
```
#default_exp core
```
# Core
> Basic functionality for unpickling objects
```
#hide
from nbdev.showdoc import *
from fastcore.test import *
import shutil
#export
from fastcore.xtras import Path
from fastcore.foundation import L
from fastcore.basics import setify, store_attr
from fastcore.meta import delegates
from importlib import import_module
import os
import pickle
#export
def _get_files(p, fs, extensions=None):
p = Path(p)
res = [p/f for f in fs if not f.startswith('.')
and ((not extensions) or f'.{f.split(".")[-1].lower()}' in extensions)]
return res
def get_files(path, extensions=None, recurse=True, folders=None, followlinks=True):
"Get all the files in `path` with optional `extensions`, optionally with `recurse`, only in `folders`, if specified."
path = Path(path)
folders=L(folders)
extensions = setify(extensions)
extensions = {e.lower() for e in extensions}
if recurse:
res = []
for i,(p,d,f) in enumerate(os.walk(path, followlinks=followlinks)): # returns (dirpath, dirnames, filenames)
if len(folders) !=0 and i==0: d[:] = [o for o in d if o in folders]
else: d[:] = [o for o in d if not o.startswith('.')]
if len(folders) !=0 and i==0 and '.' not in folders: continue
res += _get_files(p, f, extensions)
else:
f = [o.name for o in os.scandir(path) if o.is_file()]
res = _get_files(path, f, extensions)
return L(res)
#export
def _try_import(path, name):
for file in get_files(path, extensions='.py'):
try:
lib_name = str(file).split(path.name)[1]
import_statement = f'{path.name}'
for item in lib_name.split('/'):
if len(item.split('.')) > 1:
item = item.split('.')[0]
import_statement += f'{item}.'
mod = import_module(import_statement[:-1], name)
g = globals()
g[name] = getattr(mod, name)
return
except:
raise ModuleNotFoundError(f"Could not find {name} in any modules from {path.name}")
raise ModuleNotFoundError(f"Could not find {name} in any modules from {path.name}")
#hide
os.mkdir('lib')
tmp = open('lib/tmp.py', 'w')
tmp.write('def foo(a, b): return a, b')
tmp.close()
_try_import(Path('lib'), 'foo')
test_eq(foo('x', 'y'), ('x', 'y'))
shutil.rmtree('lib')
g = globals()
g.pop('foo');
del g
#hide
with ExceptionExpected(ModuleNotFoundError):
_try_import(Path('lib'), 'foo')
#export
class UnpicklerModule(pickle.Unpickler):
"Custom unpickler class for getting classes from a custom library."
@delegates(pickle.Unpickler)
def __init__(self, lib_path:Path=Path('.'), **kwargs):
store_attr()
super().__init__(**kwargs)
def find_class(self, module, name):
try:
return super().find_class(module, name)
except:
_try_import(self.lib_path, module, name)
return super().find_class(module, name)
```
| github_jupyter |
```
#import required dependencies here:
import pandas as pd
import datetime
#NOTE: before reading in any .csv files, you need to open the .csv file using a simple text editor and remove the first
#four lines (starting with '#' symbols) in order to get the csv to load correctly
#read in csv here
AQI_2018_df = pd.read_csv("./2018 Source/waqi-covid19-airqualitydata-2018H1.csv")
#check output to see if csv loaded into df correctly
AQI_2018_df.head()
#special case for the 2019 datasets - show how to merge two quarters here
#ignore this cell if you have data from any year besides 2019
#note that there are probably ~2 million rows to in the final table, so the operations might take a few seconds
#read in the different quarters - two are shown here
AQI_Info_Q1_2019 = pd.read_csv("./2019 Source/waqi-covid19-airqualitydata-2019Q1.csv")
AQI_Info_Q2_2019 = pd.read_csv("./2019 Source/waqi-covid19-airqualitydata-2019Q2.csv")
#add the quarters together using pandas' merge operation - outer join
all_quarters_df = AQI_Info_Q1_2019.merge(AQI_Info_Q2_2019, how="outer")
#subsequently merge quarters 3 and 4 into the "all_quarters_df" as shown above, but replacing the "AQI_Info_Q1_2019" df
#with the "all_quarters_df"
#now check that all the quarters have been joined correctly using an outer join by checking the table length - that is, the
#total table rows should just be the sum of the rows for each quarter's dataframe, or a merge was performed incorrectly
if(len(all_quarters_df) != (len(AQI_Info_Q1_2019) + len(AQI_Info_Q2_2019))):
print("Some rows were removed - check your merges!")
else:
print("The table lengths match, so everything was probably merged correctly!")
#group data set by date, and check the date range of the output
#it looks like for 2018 and earlier years the data might only go to the current date (~7/5) because
#those dates are comparable to the coronavirus shutdown this year
#2019 should have the full range of dates if all quarters are merged correctly
AQI_2018_df.groupby("Date").count()
#now that we have checked out the data date ranges, we should narrow the data down to the desired info
#first select by cities that we are interested in
interesting_city_list = ["Los Angeles", "Houston", "Jacksonville", "Manhattan", "Beijing", "Shanghai", "Wuhan", "New Delhi"]
interesting_cities_AQI = AQI_2018_df.loc[AQI_2018_df["City"].isin(interesting_city_list)]
#this should give some idea about how many records there are for each city we are interested in for the year
#the higher the number the more complete the data!
interesting_cities_AQI["City"].value_counts()
#get subsets of o3 and pm2.5 for each city
interesting_cities_o3_2018 = interesting_cities_AQI.loc[interesting_cities_AQI["Specie"] == "o3"]
interesting_cities_pm25_2018 = interesting_cities_AQI.loc[interesting_cities_AQI["Specie"] == "pm25"]
#can check to see if we are missing data for any days for either o3 or pm2.5
print(interesting_cities_o3_2018["City"].value_counts())
print(interesting_cities_pm25_2018["City"].value_counts())
#needed to beef up the AQI conversion table past 500 so that higher AQIs could be converted to concentrations
#used linear regression on the last 50 AQI values (450-500) to extrapolate AQI to concentration conversions for AQI values past
#500, up to 1000 - but please note that this is an assumption!
print(interesting_cities_pm25_2018["max"].max())
#import csv info for conversion table between AQI and pollutant concentrations
pm25_conversion_df = pd.read_csv("../Resources/pm25_conversion.csv")
o3_conversion_df = pd.read_csv("../Resources/o3_conversion.csv")
#conversion from aqi to concentration for pm2.5
#variance omitted because that calculation is not directly proportional to AQI and not that relevant for our project
#function AQI_to_Conc takes an o3 or an pm25 dataframe as an argument
#returns a new dataframe with the min, max, and median values converted to measures with units
#also drops the variance column for the new dataframe returned
def AQI_to_Conc(df):
#setup dictionary to choose how to convert input data
types = {"o3":o3_conversion_df, "pm25":pm25_conversion_df}
#picks conversion df dynamically
pollutant = df["Specie"].iloc[0]
conversion_df = types[pollutant]
#copy input dataframe -> will be changed by function
updated_df = df.copy()
#get the units for the concentration measurements
units = conversion_df.columns[0].replace(pollutant, "")
#add the new columns
update_columns = ["min", "max", "median"]
for col in update_columns:
#note that we need to round the AQI to get an integer for row lookup
updated_df[col + units] = updated_df[col].apply(lambda x: conversion_df[pollutant + units][round(x)])
#drop the old columns (and "variance")
updated_df = updated_df.drop(axis=1, columns=(update_columns+["variance"]))
return updated_df
#get new dataframes for the concentrations of the pollutants using the above function
#only takes the dataframe for one pollutant as an argument
interesting_cities_pm25_2018_conc = AQI_to_Conc(interesting_cities_pm25_2018)
interesting_cities_o3_2018_conc = AQI_to_Conc(interesting_cities_o3_2018)
print(interesting_cities_pm25_2018_conc.head())
interesting_cities_o3_2018_conc.head()
#output the pm25 concentration data for our cities for the year 2018
interesting_cities_pm25_2018_conc.to_csv("./2018 Cleaned/our_cities_pm25_2018.csv", index=False)
#output the o3 concentration data for our cities for the year 2018
interesting_cities_o3_2018_conc.to_csv("./2018 Cleaned/our_cities_o3_2018.csv", index=False)
```
## Stopping Point
After you have generated the dataframes that contain the concentrations for each pollutant for the cities we are looking at, and exported the CSVs for those two pollutants for those cities for your year, you are done with this first part of the project. We will need to combine the data from the first 3 years into a larger, aggregate dataframe. From this dataframe we can break down the data by month and then calculate values for each year or all three years combined.
```
#select by for a certain city, in this case we are doing Wuhan
wuhan_2018_pm25 = interesting_cities_pm25_2018_conc.loc[interesting_cities_pm25_2018_conc["City"] == "Wuhan"]
wuhan_2018_pm25.head()
#convert "Date" column values to datetime objects
wuhan_2018_pm25["Date"] = pd.to_datetime(wuhan_2018_pm25["Date"])
#re-index by date
wuhan_2018_pm25 = wuhan_2018_pm25.set_index("Date")
wuhan_2018_pm25
#example of getting data for a month
wuhan_jan_2018_pm25 = wuhan_2018_pm25.loc[wuhan_2018_pm25.index.month == 1]
wuhan_jan_2018_pm25
#can get summary stats for a month as follows
#can look at averages of medians, median of medians, average maximum for the month, maximum values for each month overall, etc.
#can also write a function later to do this for each month and output the relevant statistics to a dataframe
wuhan_jan_2018_pm25.describe()
#example line plot for wuhan for overall time period of 2018 for pm2.5
#first plot median line graph
example_plot = wuhan_2018_pm25.plot(kind="line", y="median (ug/m3)")
#sort the values by date
wuhan_2018_pm25 = wuhan_2018_pm25.sort_values(by="Date")
#then plot the max line graph on the same plot
example_plot.plot(wuhan_2018_pm25.index, wuhan_2018_pm25["max (ug/m3)"])
wuhan_2018_pm25
#example bar graph
averages_per_month = []
max_averages_per_month = []
months = []
for m in range(1, 7):
months.append(m)
curr_month_wuhan = wuhan_2018_pm25.loc[wuhan_2018_pm25.index.month == m]
averages_per_month.append(curr_month_wuhan["median (ug/m3)"].mean())
max_averages_per_month.append(curr_month_wuhan["max (ug/m3)"].mean())
bar_chart_df = pd.DataFrame({"Median Average per Month Wuhan 2018":averages_per_month,
"Max Avg Per Month":max_averages_per_month}, index=months)
bar_axes = bar_chart_df.plot(kind="bar", y="Max Avg Per Month", color="r")
bar_axes.bar([a-1 for a in months], bar_chart_df["Median Average per Month Wuhan 2018"], label="Median Avg Per Month")
bar_axes.legend(loc="best")
```
I wanted to do the shutdown times vs. previous years by month on a chart like the one above for some measure of center to get the percentage pollution decrease
| github_jupyter |
# BlazingSQL Cheat Sheets sample code
(c) 2020 NVIDIA, Blazing SQL
Distributed under Apache License 2.0
### Imports
```
import cudf
import numpy as np
from blazingsql import BlazingContext
```
### Sample Data Table
```
df = cudf.DataFrame(
[
(39, -6.88, np.datetime64('2020-10-08T12:12:01'), 'C', 'D', 'data'
, 'RAPIDS.ai is a suite of open-source libraries that allow you to run your end to end data science and analytics pipelines on GPUs.')
, (11, 4.21, None, 'A', 'D', 'cuDF'
, 'cuDF is a Python GPU DataFrame (built on the Apache Arrow columnar memory format)')
, (31, 4.71, np.datetime64('2020-10-10T09:26:43'), 'U', 'D', 'memory'
, 'cuDF allows for loading, joining, aggregating, filtering, and otherwise manipulating tabular data using a DataFrame style API.')
, (40, 0.93, np.datetime64('2020-10-11T17:10:00'), 'P', 'B', 'tabular'
, '''If your workflow is fast enough on a single GPU or your data comfortably fits in memory on
a single GPU, you would want to use cuDF.''')
, (33, 9.26, np.datetime64('2020-10-15T10:58:02'), 'O', 'D', 'parallel'
, '''If you want to distribute your workflow across multiple GPUs or have more data than you can fit
in memory on a single GPU you would want to use Dask-cuDF''')
, (42, 4.21, np.datetime64('2020-10-01T10:02:23'), 'U', 'C', 'GPUs'
, 'BlazingSQL provides a high-performance distributed SQL engine in Python')
, (36, 3.01, np.datetime64('2020-09-30T14:36:26'), 'T', 'D', None
, 'BlazingSQL is built on the RAPIDS GPU data science ecosystem')
, (38, 6.44, np.datetime64('2020-10-10T08:34:36'), 'X', 'B', 'csv'
, 'BlazingSQL lets you ETL raw data directly into GPU memory as a GPU DataFrame (GDF)')
, (17, -5.28, np.datetime64('2020-10-09T08:34:40'), 'P', 'D', 'dataframes'
, 'Dask is a flexible library for parallel computing in Python')
, (10, 8.28, np.datetime64('2020-10-03T03:31:21'), 'W', 'B', 'python'
, None)
]
, columns = ['number', 'float_number', 'datetime', 'letter', 'category', 'word', 'string']
)
bc = BlazingContext()
bc.create_table('df', df)
```
# SQL String Functions
#### CONCAT
```
query = '''
SELECT string
, 'INFO: ' || string AS r
FROM df
'''
bc.sql(query)
```
#### SUBSTRING
```
query = '''
SELECT string
, SUBSTRING(string, 0, 2) AS r
FROM df
'''
bc.sql(query)
```
#### CHAR_LENGTH
```
query = '''
SELECT string
, CHAR_LENGTH(string) AS r
FROM df
'''
bc.sql(query)
```
#### LEFT
```
query = '''
SELECT string
, LEFT(string, 3) AS r
FROM df
'''
bc.sql(query)
```
#### RIGHT
```
query = '''
SELECT string
, RIGHT(string, 3) AS r
FROM df
'''
bc.sql(query)
```
#### LTRIM
```
query = '''
SELECT string
, LTRIM(string) AS r
FROM df
'''
bc.sql(query)
```
#### RTRIM
```
query = '''
SELECT string
, RTRIM(string) AS r
FROM df
'''
bc.sql(query)
```
#### REPLACE
```
query = '''
SELECT string
, REPLACE(string, 'RAPIDS', 'NVIDIA RAPIDS') AS r
FROM df
'''
bc.sql(query)
```
#### UPPER
```
query = '''
SELECT string
, UPPER(string) AS r
FROM df
'''
bc.sql(query)
```
#### LOWER
```
query = '''
SELECT string
, LOWER(string) AS r
FROM df
'''
bc.sql(query)
```
#### REVERSE
```
query = '''
SELECT string
, REVERSE(string) AS r
FROM df
'''
bc.sql(query)
```
| github_jupyter |
```
import requests
import os
import pandas as pd
from bs4 import BeautifulSoup
import urllib
import re
def get_singers(url=None):
if url is None:
raise ValueError('You have to enter a valid URL')
else:
r = requests.get(url)
soup = BeautifulSoup(r.text, "html.parser")
table = soup.find_all("div", class_="itemListSubCategories")[0]
singers = table.find_all('a')
all_singers = []
for singer in singers:
all_singers.append([singer.text.strip(), singer.get('href')])
all_singers_df = pd.DataFrame(all_singers, columns=['name', 'link'])
return all_singers_df
def get_lyrics(song_url=None):
if song_url is None:
raise ValueError('You have to enter a valid URL')
else:
base = 'http://fnanen.net'
url = ''.join(base + song_url)
r = requests.get(url)
soup = BeautifulSoup(r.text, "html.parser")
lyrics = soup.find_all("div", class_="itemFullText")[0]
raw_lyrics = re.sub('<[^>]*>', '\n', str(lyrics))
return raw_lyrics
def get_songs_from_url(url=None, with_lyrics = True, verbose=True):
if url is None:
raise ValueError('You have to enter a valid URL')
else:
r = requests.get(url)
soup = BeautifulSoup(r.text, "html.parser")
table = soup.find_all("div", class_="itemListView")[0]
titles = table.find_all('a',attrs={'class':'', 'data-animation':'true'})
songs = []
if with_lyrics:
for title in titles:
songs.append([title.text, title.get('href'), get_lyrics(title.get('href'))])
all_songs_df = pd.DataFrame(songs, columns=['title', 'link', 'lyrics'])
if verbose:
print('Song ', title.text, ' fetched.')
else:
for title in titles:
songs.append([title.text, title.get('href')])
all_songs_df = pd.DataFrame(songs, columns=['title', 'link'])
if verbose:
print('Song ', title.text, ' fetched.')
return all_songs_df
def get_songs(url=None, with_lyrics=True, verbose=True):
base = 'http://fnanen.net'
if url is None:
raise ValueError('You have to enter a valid URL')
else:
r = requests.get(''.join(base + url))
soup = BeautifulSoup(r.text, "html.parser")
table = soup.find_all("div", class_="itemListView")[0]
titles = table.find_all('a',attrs={'class':''})
pagination = soup.find_all('div', 'k2Pagination')
songs_df = pd.DataFrame({'title':[], 'link':[],'lyrics':[]})
links = [''.join(base + url)]
if pagination != []:
pages = pagination[0].find_all('a', attrs={'title':['2','3','4']})
for p in pages:
links.append(''.join(base + p.get('href')))
for link in links:
if verbose:
print('Parsing.. ', link)
try:
songs = get_songs_from_url(link, with_lyrics=with_lyrics, verbose=verbose)
songs_df = songs_df.append(songs, ignore_index=True)
except:
print('ERROR: broken link? check ', link)
continue
return songs_df
r = requests.get('http://fnanen.net/')
soup = BeautifulSoup(r.text, "html.parser")
letters = soup.find_all("ul", class_="menu menu-vertical dropdown-hover ")[0]
subpages = [''.join('http://fnanen.net' + l.get('href')) for l in letters.find_all('a')]
subpages.remove(subpages[16])
all_songs = []
for page in subpages[17:]:
m_singers = get_singers(page)
for singer in m_singers.iterrows():
url = singer[1][1]
songs = get_songs(url, with_lyrics=True, verbose=False)
songs = songs.assign(singer=singer[1][0])
all_songs.append(songs)
print('done with ', singer[1][0], '. All songs len is: ', len(all_songs) )
all_songs = pd.concat(all_songs, ignore_index=True)
all_songs.shape
all_songs.to_pickle('all_songs.pickle')
```
| github_jupyter |
```
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from nltk.tokenize import word_tokenize
from collections import Counter
import numpy as np
import matplotlib as mpl
mpl.rcParams['figure.dpi'] = 200
# data
df_to_eda = pd.read_csv("../../data/curated/df_to_eda.csv")
# how much data we have on school_step
df_to_eda["school_step"].value_counts()
```
Como podemos ver aqui, existe uma quantidade bem interessante entre as 3 classes que iremos prever. No FUND-I temos um pouco mais da metade das outras duas classes, mas acreditamos que não será um problema.
```
df_to_eda
```
# Tamanho das sentenças
```
max = df_to_eda['question_c6'].str.len().max()
min = df_to_eda['question_c6'].str.len().min()
median = df_to_eda['question_c6'].str.len().median()
mean = df_to_eda['question_c6'].str.len().mean()
print('As questões vão de %d à %d caracteres por questão' % (min, max))
print('O valor mediano e médio de caracteres por questão é de %d e de %d, respectivamente.' %
(median, mean))
fig, ax = plt.subplots(figsize=(20, 10))
sns.histplot(df_to_eda['question_c6'].str.len(), ax = ax)
targets = ["Médio & Pré-Vestibular", "Fundamental II", "Fundamental I"]
for target in targets:
fig, ax = plt.subplots(figsize=(20, 10))
_ = sns.histplot(df_to_eda[df_to_eda["school_step"] == target]['question_c6'].str.len(), ax = ax)
plt.xlabel(target)
plt.show()
```
Curiosamente temos muitas sentenças de tamanho 1. Provavelmente esses itens não representam nada e precisam ser excluídos.
```
df_to_eda["len_sentence"] = df_to_eda["question_c6"].str.len()
# filtering values more than 50.000
df_to_eda_less_than_50k = df_to_eda[df_to_eda["len_sentence"] < 50000]
_ = sns.boxplot(x="school_step", y="len_sentence", data=df_to_eda_less_than_50k)
plt.xlabel("")
plt.ylabel("Quantidade de Caracteres")
plt.show()
df_to_eda.sort_values(["len_sentence"])
df_to_eda['question_c6'].astype("str").apply(len).sort_values()
# looking at those items with sentence length of 1
df_to_eda[df_to_eda["question_c6"].astype("str").apply(len) == 3]
# lets look at length by sentence
df_to_eda.value_counts(["len_sentence", 'school_step'])
df_to_eda[df_to_eda["question_c6"].astype("str").apply(len) == 30].value_counts(["school_step"])
df_to_eda[~df_to_eda["question_c6"].astype("str").apply(len).isin([1, 2, 3])].shape
# filtering length 1 sentence size items
df_c1 = df_to_eda[df_to_eda["question_c6"].astype("str").apply(len) != 1]
# let's check the length of these sentences again
df_c1['question_c6'].astype("str").apply(len).sort_values()
df_c1["sentence_len"] = df_c1["question_c6"].astype("str").apply(len)
df_c1
```
Sabemos que apesar de termo removido as stopwords, existem palavras específicas desse contexto que não representam muito para o problema que temos. Porém, para melhor visualizá-las, vamos plotar uma wordcloud em todas as sentenças do dataset.
```
# plot wordcloud
tokens = []
text_list = df_c1["question_c6"].values
_ = [tokens.extend(word_tokenize(str(text))) for text in text_list]
counting_tokens = Counter(tokens)
count_tokens_df = pd.DataFrame({"words": counting_tokens.keys(), "count": counting_tokens.values()})
count_tokens_df.sort_values(["count"], ascending=False)
count_tokens_df["q_groups"] = pd.qcut(count_tokens_df["count"], q=[.1, .5, .6, .7, .8, .9, .99, .999, 1])
count_tokens_df.value_counts("q_groups")
count_tokens_df["q_groups"] = count_tokens_df["q_groups"].astype("str")
count_tokens_df
bins_to_plot = count_tokens_df["q_groups"].value_counts().reset_index()
_ = sns.barplot(x="q_groups", y="index", data=bins_to_plot)
plt.xlabel("Quantidade de Observações")
plt.ylabel("Grupos")
plt.show()
```
Mais de 90% das palavras aparecem de 1 a 25 vezes, precisamos investigar essas palavras.
```
less_than_25_df = count_tokens_df[count_tokens_df["count"] <= 25]
less_than_25_df
```
| github_jupyter |
```
# @Vishal Dhiman
try:
%tensorflow_version 2.x
except Exception:
pass
from __future__ import absolute_import,print_function,division,unicode_literals
import pathlib
import tensorflow as tf
from tensorflow import keras
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
class PrintDot(keras.callbacks.Callback):
def on_epoch_end(self,epoch,logs):
if epoch % 100==0: print("")
print(".",end="")
class MPG_data:
def __init__(self):
self.epoch=0
self.column_name=['MPG','Cylinders','Displacement','Horsepower','Weight','Acceleration','Model Year','Origin']
def load_data(self):
#loading data
data_path=keras.utils.get_file("auto-mpg.data","http://archive.ics.uci.edu/ml/machine-learning-databases/auto-mpg/auto-mpg.data")
raw_data=pd.read_csv(data_path,names=self.column_name,na_values='?',comment='\t',sep=" ",skipinitialspace=True)
self.dataset=raw_data.copy()
print("\n[Out]: Data Loaded Successfully.")
def preprocessing(self):
#normalize 0-1
self.dataset = self.dataset.dropna()
origin=self.dataset.pop('Origin')
self.dataset['USA']=(origin==1)*1.0
self.dataset['Europe']=(origin==2)*1.0
self.dataset['Japan']=(origin==3)*1.0
print("\n[Out]: Preprocessing done.\n")
def data_info(self):
print("[Out]: Data Frame:\n",self.dataset.head(10))
print("\n [Out]: Data shape: ",self.dataset.shape)
print("\n [Out]: Show Missing values:\n ",self.dataset.isna().sum())
def train_test_split(self):
self.train_set=self.dataset.sample(frac=.8,random_state=0)
self.test_set=self.dataset.drop(self.train_set.index)
print("\n [OUT]:Train and Test Split Done.")
def show_data(self):
print("\n[OUT]:correlation of various features plot:")
sns.pairplot(self.train_set[['MPG','Cylinders','Displacement','Weight']],diag_kind="kde")
def data_stats(self):
self.train_lab=self.train_set.pop("MPG")
self.test_lab=self.test_set.pop("MPG")
train_stats=self.train_set.describe()
#train_stats.pop("MPG")
train_stats=train_stats.transpose()
self.train_stats=train_stats
print("\n[OUT]: Data Stats are as:\n",print(train_stats))
def norm_data(self):
self.train_set.applymap(lambda x : x-self.train_stats['mean']/self.train_stats['std'])
self.test_set.applymap(lambda x : x-self.train_stats['mean']/self.train_stats['std'])
print("\n[OUT]:Normalization done.")
def create_model(self):
#define layers
model=keras.Sequential([
keras.layers.Dense(64,activation='relu',input_shape=[len(self.train_set.keys())]),
keras.layers.Dense(64,activation='relu'),
keras.layers.Dense(1)
])
optimizer=tf.keras.optimizers.RMSprop(.001)
model.compile(loss='mse',optimizer=optimizer,metrics=['mae','mse'])
print("[Out]: Model Summary:")
model.summary()
print("[Out]: Model creation and compilation successful. ")
return model
def train_model(self,model):
Epo=1000
#early_stop=keras.callbacks.EarlyStopping(monitor="val_loss",patience=10)
hist=model.fit(self.train_set,self.train_lab,epochs=Epo,validation_split=0.2,verbose=0,callbacks=[PrintDot()])
print("[Out]: Model now trained.")
history=pd.DataFrame(hist.history)
#print(history)
history['epoch']=hist.epoch
print("\n[OUT]:History After trainig model",history.head(10))
self.plot_history(hist)
return model
def plot_history(self,hist):
history=pd.DataFrame(hist.history)
history['Epoch']=hist.epoch
plt.figure()
plt.title("Count of Eopoch vs Mae")
plt.xlabel("Epoch")
plt.ylabel("Mean Absolute Error")
plt.plot(history['Epoch'],history['mae'],label='Train error')
plt.plot(history['Epoch'],history['val_mae'],label='Val_error')
#plt.ylim([0,5])
plt.legend()
plt.figure()
plt.title("Count of Eopoch vs MSE")
plt.xlabel("Epoch")
plt.ylabel("Mean Square Error")
plt.plot(history['Epoch'],history['mse'],label='Train error')
plt.plot(history['Epoch'],history['val_mse'],label='Val_error')
#plt.ylim([0,20])
plt.legend()
plt.show()
def evaluate_model(self,model):
test_los,mae,mse=model.evaluate(self.test_set,self.test_lab,verbose=0)
print("[Out]: Model test loss:{} and test mae:{} and mse:{} ".format(test_los,mae,mse))
def model_predict(self,model):
test_pred=model.predict(self.test_set).flatten()
print("\n[OUT]:Plotting predictions")
plt.scatter(self.test_lab,test_pred)
plt.title("Precition for test")
plt.xlabel("Actual")
plt.ylabel("Predictions")
plt.axis("equal")
plt.axis("square")
plt.xlim([0,plt.xlim()[1]])
plt.ylim([0,plt.ylim()[1]])
_ = plt.plot([-100,100],[-100,100])
return test_pred
def error_distribution(self,pred):
error=pred-self.test_lab
plt.figure()
plt.hist(error,bins=25)
plt.title("Distribution error")
plt.xlabel("Prediction Error")
plt.ylabel("Count")
def save_model(self,model):
model.save("model_classification.h5")
print("[Out]: Model saved.")
obj=MPG_data()
obj.load_data()
obj.preprocessing()
obj.data_info()
obj.train_test_split()
obj.show_data()
obj.data_stats()
obj.norm_data()
model=obj.create_model()
model=obj.train_model(model)
obj.evaluate_model(model)
pred=obj.model_predict(model)
obj.error_distribution(pred)
#obj.save_model(model)
```
| github_jupyter |
```
# Copyright 2019 The Kubeflow Authors. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# Install Pipeline SDK - This only needs to be ran once in the enviroment.
!python3 -m pip install 'kfp>=0.1.31' --quiet
!pip3 install tensorflow==1.14 --upgrade
```
## KubeFlow Pipelines Serving Component
In this notebook, we will demo:
* Saving a Keras model in a format compatible with TF Serving
* Creating a pipeline to serve a trained model within a KubeFlow cluster
Reference documentation:
* https://www.tensorflow.org/tfx/serving/architecture
* https://www.tensorflow.org/beta/guide/keras/saving_and_serializing
* https://www.kubeflow.org/docs/components/serving/tfserving_new/
### Setup
```
# Set your output and project. !!!Must Do before you can proceed!!!
project = 'Your-Gcp-Project-ID' #'Your-GCP-Project-ID'
model_name = 'model-name' # Model name matching TF_serve naming requirements
import time
ts = int(time.time())
model_version = str(ts) # Here we use timestamp as version to avoid conflict
output = 'Your-Gcs-Path' # A GCS bucket for asset outputs
KUBEFLOW_DEPLOYER_IMAGE = 'gcr.io/ml-pipeline/ml-pipeline-kubeflow-deployer:1.7.0-rc.2'
model_path = '%s/%s' % (output,model_name)
model_version_path = '%s/%s/%s' % (output,model_name,model_version)
```
### Load a Keras Model
Loading a pretrained Keras model to use as an example.
```
import tensorflow as tf
model = tf.keras.applications.NASNetMobile(input_shape=None,
include_top=True,
weights='imagenet',
input_tensor=None,
pooling=None,
classes=1000)
```
### Saved the Model for TF-Serve
Save the model using keras export_saved_model function. Note that specifically for TF-Serve the output directory should be structure as model_name/model_version/saved_model.
```
tf.keras.experimental.export_saved_model(model, model_version_path)
```
### Create a pipeline using KFP TF-Serve component
```
def kubeflow_deploy_op():
return dsl.ContainerOp(
name = 'deploy',
image = KUBEFLOW_DEPLOYER_IMAGE,
arguments = [
'--model-export-path', model_path,
'--server-name', model_name,
]
)
import kfp
import kfp.dsl as dsl
# The pipeline definition
@dsl.pipeline(
name='sample-model-deployer',
description='Sample for deploying models using KFP model serving component'
)
def model_server():
deploy = kubeflow_deploy_op()
```
Submit pipeline for execution on Kubeflow Pipelines cluster
```
kfp.Client().create_run_from_pipeline_func(model_server, arguments={})
#vvvvvvvvv This link leads to the run information page. (Note: There is a bug in JupyterLab that modifies the URL and makes the link stop working)
```
| github_jupyter |

This is a map of **Digital Design in Architecture**. It is created with data from 12,000 articles published between 1975 and 2019.
You can download a [high-res PDF version here](images/map-digital-design-in-architecture.pdf).
You can use this map to learn about the various concepts that build the field of digital design in architecture and the main authors defining them.
You can also check what is not there yet.
The code i used to generate it is below.
# IDEA
I recently found this interesting [visualization of 20th Century Philosophy by Maximilian Noichl](https://homepage.univie.ac.at/noichlm94/posts/structure-of-recent-philosophy-iii/).
The chart is drawn by sampling and clustering some 50,000 peer-reviewed journal article on philosophy.
It was amazing how I was able to discover relationships and authors in a discipline, which I am not familiar with, through the data clustering that it presented!
So I wanted to do a similar thing for my field of research for two reasons:
- I feel at home working in a data-driven workflow. So it felt natural to try and help myself structure the sourcing process for the literature review for my PhD thesis using a data-driven approach as well.
- I was curious what discussion such a visualization would prompt in our computational design community.
Maximilian has generously shared his code and when I emailed him, he advised me on what to pay attention to when creating my own map. Big thanks to Max!
I used Max's code as a base to built this map. My changes were in the cluster tree chart and in the text analysis. I also documented the process on how to get the data needed for such a map.
The idea of this Jupyter notebook is that you can use it to produce your own map for your own field of interest. It is build step by step but is meant to be used in a loop where you would see the result in one step and go back few steps to change the parameters (for example the size of the research clusters - Max used 440, in my case this number was too big because I had less articles) and run again.
# INIT
Let's initialize the environment and load the needed libraries.
Check the [SETUP.md](SETUP.md) file for instructions on how to set up your development environment so that you can run this Jupyter notebook.
```
# This cell loads the needed Python libraries
import metaknowledge as mk
import pandas as pd
import numpy as np
from random import randint
import datetime
import scipy as scipy
import csv
import textwrap
%matplotlib inline
import seaborn as sns
import matplotlib.pyplot as plt
#For Tables:
from IPython.display import display
from IPython.display import Latex
pd.set_option('display.max_columns', 500)
#For R (ggplot2)
#check the rpy situation by uncommenting the following
#%run -m rpy2.situation
#use the following on macOS, comment it out on Windows
%load_ext rpy2.ipython
#on Windows rpy2 doesnt show output of R cells, use the one below if on Windows
#use the following on Windows, comment it out on macOS
#%load_ext RWinOut
```
# DATA
The method to generate this map of the research landscape uses publication data from journals and conferences. The clustering is based on the co-citation patterns, i.e. which authors cite each others' papers most often.
## Where does the data come from?
So as data we need data from publications. The method by Max Noichl used [Web-Of-Science (WOS)](https://www.webofknowledge.com/) as the data source. I cannot publish the data I used to generate the map since I don't have permission to distribute it but I can show you how to get the data on your own.
## How to download publication metadata from Web-of-Science (WOS)
Open the WOS website and use your institutional sign in to log in. For German universities like the one I am at use the `German Higher Education and Research (DFN-AAI)` option and find your university on the next page.
Now let's do a sample search. Enter `International Journal of Architectural Computing` in the search field and change the type to `Publication Name`.

<img src="images/method/wos_search.png" alt="WOS search" width="75%"/>
The search results are 149 as of 5th August 2020.
<img src="images/method/wos_results.png" alt="WOS results" width="75%"/>
Click on the `Export...` button and choose `Other File Formats`.
<img src="images/method/wos_export_formats.png" alt="WOS Export formats" width="232"/>
In the next window choose Records from 1 to 149. As content choose `Author, Title, Source, Abstract`. And as File Format choose Plain Text.
<img src="images/method/wos_export_options.png" alt="WOS Export options" width="290"/>
Click Export to download the file with the data.
## How I selected the data about publications on digital technology and architectural design
The data used in the next steps has been collected in January 2019. As more publications are being added to WOS over the years, the map will for sure change a bit.
### Step 1 — A list of established conferences
I made a list of the conferences and journals that where most often discussed at my department ([DDU at TU Darmstadt](https://www.dg.architektur.tu-darmstadt.de/fachgebiet_ddu/index.de.jsp)) and where my colleagues and I have had submitted papers to:
Conferences:
- AAG
- Acadia
- CAAD futures
- Caadria
- Design Modelling Symposium
- eCAADe
- Fabricate
- RobArch
- Sigradi
- SimAUD
Peer-reviewed journals:
- IJAC
I checked how many papers from those are available on WOS. Not many :(
As you see below, I had only 1,600 something publications, yet I needed much more to create the map.
Anyway, I downloaded the metadata from WOS.
```
#list the established conferences
df = pd.read_csv('data/01a_established_conf.csv', delimiter=';')
display(df)
```
### Step 2 — List of authors established in the field
So then I decided to start with a list of people and then derive the publication sources from it.
I made a list of the authors that are established in the field of digital design in architecture. Then I checked how many articles are listed in WOS for each author and downloaded their metadata.
```
#list the selected authors
df = pd.read_csv('data/02a_established_authors.csv', delimiter=';')
display(df)
```
If you miss your hero in this list, email me or open an issue in this GitHub repo. However, the purpose of this list is to use it to get to a list of journals, so adding or removing one name might not change the results from the steps that follow.
Then I downloaded the publication data for those authors that was available on WOS.
Next we are interested in the journals where the authors from the list have published.
```
# take a glimpse at the data we have for top authors in the field
RCAuthor = mk.RecordCollection("data/02b_established_authors")
print(RCAuthor.glimpse())
```
### Step 3 — Get a list of journals where established authors have published
I could use the publication data for the top authors to derive a list of journals. I got a list of 210 journals, conference proceedings and books. There were some interesting new sources in the list but also several problems with it. Some sources were not peer-reviewed, others didn't contain publications with citations, yet citing patterns are important for this map, some sources didn't have their focus on architecture and digital technology and so on. So the list needed cleaning up.
```
# list all journal ranked by occurence
topJU = RCAuthor.rankedSeries(tag='journal', pandasMode=False)
display(pd.DataFrame(topJU,columns=['Journal', 'Articles by top authors']))
# And export the list to a CSV file for manual review
with open("data/03_journals_from_established_authors.csv", "w") as output:
writer = csv.writer(output, lineterminator='\n')
writer.writerows(topJU)
```
### Step 4 — Clean up the journal list
I exported the list of journals where the top authors had published in. Then I checked each journal one by one and excluded the really broad ones such as Architectural Digest, as well those focusing solely on engineering, history, art or aesthethical/political critique. I also removed any books from the list, such as DIGITAL TURN IN ARCHITECTURE by Mario Carpo.
Then I merged the list with the established conferences from step 1
I was left with 17 Journals.
Let's see how many articles this gives us.
```
#the updated list of journals
df = pd.read_csv('data/04a_cleaned_up_journals.csv', delimiter=';')
display(df)
# take a glimpse at the data we have for now
RCtmp = mk.RecordCollection("data/04b_cleaned_up_journals")
print(RCtmp.glimpse())
```
A little over 19000 articles. Not bad. I wanted to check for more.
### Step 5 — Fish for more data using Scimago
So using the website [Scimago](https://www.scimagojr.com/journalrank.php?category=2216) and filtering to the field of architecture, I looked for sources that:
1. Had a high ranking
2. Focused on digital technology and architecture
3. Had data available on WOS
This resulted only in one new journal being added to the list:
TECHNE-JOURNAL OF TECHNOLOGY FOR ARCHITECTURE AND ENVIRONMENT
However the whole exersize was quite interesting to get an overview of the publication landscape in architecture. If one day I decide to make a map of the whole architecture research, not just the one relating to digital design, the insights would be useful.
So let's load the data for the map.
### Step 6 — load and clean up data
First let's load the data and take a glimpse of what we have so far.
```
date_string = datetime.datetime.now().strftime("%Y-%m-%d-%H:%M")
#read the data for all journals
RC = mk.RecordCollection("data/06_all_journals")
print(RC.glimpse())
```
An intersting thing we can do with the data at this point is to see which are the TOP30 journals where the articles cited in the articles in the collection are published. This can help find more journals to include in the dataset.
```
# list TOP30 journals from the cited works in these articles
topJUcited = []
for R in RC:
#topJUcited.append(list(set(R.getCitations().get('citeString'))))
topJUcited.append(list(set(R.getCitations().get("journal"))))
#print(topJUcited[:10])
rankedJU = {}
for jlist in topJUcited:
for ju in jlist:
rankedJU[ju] = rankedJU.get(ju, 0) + 1
sorted_by_value = sorted(rankedJU.items(), key=lambda kv: kv[1], reverse=True)
#RC.rankedSeries(tag='journal', pandasMode=False)
display(pd.DataFrame(sorted_by_value[:30], columns=['Journal','Times cited']))
```
Next we will remove the articles which have less then a certain number of citations to reduce the noise in the map. We can also remove all articles that don't list the year of their publication, or are without abstracts. This helps later when we want to understand what each cluster is researching about. Let's print a glimpse of the data so far.
```
#prune the collection from articles that miss key data
RC2 = mk.RecordCollection()
for R in RC:
randnr = randint(0, 4)
if len(R.getCitations().get("author"))>=3: # and randnr==0 apply condition in order to downsample records
#Here we kick out every paper that cites less then 3 authors. Why? because they
#are so dissimilar from the others, that they only produce noise.
try:
R['year']
#if R['year']>1961:
#R['abstract'] #Add this when working with abstracts. It removes every paper that has none.
#This can sometimes remove whole journals, that are archived without abstracts, so handle with care.
RC2.add(R)
except KeyError:
pass
else:
pass
print(RC2.glimpse())
RC = RC2
```
Ok, we lost 7,000 articles. The clean up reduced the amount of datapoints to about 12,000 which is still ok.
The glimpse shows us the most prolific researchers where we have the computational design superstar Achim Menges. We see the journals with the most publications in the dataset. And we see the top cited articles where the winner is [George Stiny's "Kindergarten Grammars: Designing with Froebel's Building Gifts"](https://journals.sagepub.com/doi/abs/10.1068/b070409).The [The Palladian Grammar](https://journals.sagepub.com/doi/abs/10.1068/b050005) again by Stiny is 3rd. [Bill Hillier's Space Syntax Book](https://www.cambridge.org/core/books/social-logic-of-space/6B0A078C79A74F0CC615ACD8B250A985) and [Horst Rittel's Dillemas in a General Theory of Planning](https://link.springer.com/article/10.1007/BF01405730) also make it in the TOP 10. So far so good.
Next let's assemble a clean list of all included journals and conferences so we can print it next to the map.
```
#list all journals where the papers in the data were published
# list all journal ranked by occurence
topJU = RC.rankedSeries(tag='journal', pandasMode=False)
#conference proceedings or journals sometimes appear as separate journals for each year or issue. lets combine them
combineConfProc = [
# list of what to search for and what to change the name to and tally it up as
[['acadia'], 'ACADIA CONFERENCE PROCEEDINGS'],
[['ecaade','fusion:'], 'ECAADE CONFERENCE PROCEEDINGS'],
[['caadria'], 'CAADRIA CONFERENCE PROCEEDINGS'],
[['simaud'], 'SIMAUD CONFERENCE PROCEEDINGS'],
[['caad futures'], 'CAAD FUTURES CONFERENCE PROCEEDINGS'],
]
journalList = []
confprocdict = {}
for j in topJU:
#if conf name found, combine in conferences
isConf = False
jouName = j[0]
for conf in combineConfProc:
for searchphrase in conf[0]:
if searchphrase.lower() in jouName.lower():
isConf = True
if not conf[1] in confprocdict:
confprocdict[conf[1]] = [conf[1],j[1]]
else:
confprocdict[conf[1]][1] = confprocdict[conf[1]][1] + j[1]
#wrapepdJuName = textwrap.fill(j[0],70)
if not isConf:
aju = [ jouName, j[1]]
journalList.append(aju)
#print(wrapepdJuName + " (" + str(j[1])+ ")")
#print()
for conf in confprocdict:
journalList.append(confprocdict[conf])
wrapper=textwrap.TextWrapper()
wrapper.width=60
wrapper.initial_indent=""
wrapper.subsequent_indent=" "
#sortedJournalList
journalList = sorted(journalList, key=lambda x: x[0][0])
journalListLabels = []
for journal in journalList:
label = wrapper.fill(journal[0] + " (" + str(journal[1]) + ")")#+"\n"
journalListLabels.append(label)
#print(label )
journalListLabel = "THE MAP IS GENERATED FROM DATA FROM THE FOLLOWING PUBLICATIONS:" + "\n"+"\n".join([w for w in journalListLabels]) + "\nDATA SNAPSHOT 23 JAN 2019"
print(journalListLabel)
```
# MAP
Now let’s do the map. The method is explained below with the code. The code for the first 4 steps (extracting the fetures, dimensionality reduction, 2D embedding with umap and clustering) is largely based on the [code shared by Max Noichl here](https://homepage.univie.ac.at/maximilian.noichl/posts/structure-of-recent-philosophy-ii/), so refer to his Jupyter Notebook for details.
## Build a multi-dimensional space of relationships
In order to arrange the publications in a 2D map (which is done with UMAP), and subsequently to determine the clusters, the publications datapoints need to be assigned `features`. The datapoints with more similar features will be placed closer to each other. Max Noichl carried out some experiments with features such as author, words in abstracts, title and so on and he recommends using the following two features:
- the cited works
- the cited authors
To extract the features we use `scikit-learn` and clean up the weakly linked points.
First let's do the cited works, then the cited authors and then fitler out weak links.
```
########### Cited Works - Features ############
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer
import re
drc = pd.DataFrame.from_dict(RC.forNLP(extraColumns=['journal','AU','FU','PD']))
d = []
citedAU = []
citestring =[]
for R in RC:
d.append(list(set(R.getCitations().get("citeString")))) #To cluster by cited author
citedAU.append(list(set(R.getCitations().get("author"))))
citestring.append(list(set(R.getCitations().get("citeString"))))
drc["citedAU"] = citedAU
drc["citestring"] = citestring
#print(d[0])
authorslist = ['§'.join(filter(None,x)) for x in list(d)]
#print(authorslist[0])
# vec = TfidfVectorizer(token_pattern=r'(?<=[^|§])[\s\w,\.:;]+(?=[$|§])')
vec = CountVectorizer(token_pattern=r'(?<=[§])[\s\w,\.:;\/\[\]-]+(?=[§])',binary=True, min_df = 3)#, min_df = 1)
Xrc = vec.fit_transform(authorslist)
########### Authors - Features ############
d = []
for R in RC:
authors = list(set(R.getCitations().get("author")))
# print(authors)
authors = filter(None, authors)
f = []
for a in authors:
f.append(' '.join([w for w in a.split(' ')if len(w)>2]))
authors = f#' '.join(f)
d.append(authors)
authorslist = [';'.join(filter(None,x)) for x in list(d)]
vec = CountVectorizer(token_pattern=r'(?<=[;])[\s\w]+(?=[;])',binary=True, min_df = 10)
XrcAu = vec.fit_transform(authorslist)
k = [XrcAu,Xrc]
XrcFull = scipy.sparse.hstack(k).tocsr()
###### Filtering #######
from scipy.sparse import coo_matrix, vstack
from scipy.sparse import csr_matrix
import scipy as scipy
row_names = np.array(drc["id"])
newdf=[]
a = 0
# index by name:
for x in range(0,XrcFull.shape[0]): #Xrc.shape[0]):
row_idx, = np.where(row_names == drc["id"][x])
if np.diff(XrcFull[row_idx].tocsr().indptr) >= 4:
if a == 0:
k = [XrcFull[row_idx]]
if a != 0:
k.append(XrcFull[row_idx])
a = a+1
newdf.append(drc.loc[x])
drc = pd.DataFrame(newdf).reset_index()
M = scipy.sparse.vstack((k))
```
## Preliminary dimensionality reduction with SVD
Here is what Max Noichl says about this next step:
> "This is strictly speaking not necessary: We could pass our vectors directly to the umap-algorithm. But when we use a lot of data, we can get a sharper image when we clear out some noise with SVD beforehand. If we were interested more in classification and less in visualization, I would suggest to skip this step, reduce with umap to ~thirty dimensions and cluster on that."
```
from sklearn.decomposition import TruncatedSVD
SVD = TruncatedSVD(n_components=350, n_iter=7, random_state=42)
XSVD = SVD.fit_transform(M)
print(SVD.explained_variance_ratio_.sum())
dSVD = pd.DataFrame(XSVD)
sSVD = dSVD[[0,1]]
sSVD.columns = ['x','y']
```
## Project to 2D space
To generate a 2D projection we creating a 2D-embedding with [UMAP](https://github.com/lmcinnes/umap).
As far as I understand the algorithm here, each feature established above is a dimension along which distance between articles is measured. So we have a space of several hundred dimensions. How to plot that, i.e. project it to a 2D space so we can vizualize it as a map?
`UMAP` let's us reduce the dimensions to two and is good for two reasons: speed and preserving relationships.
So in the next step we use `UMAP` to create a 2D-embedding of the data.
```
#if this cells issues a numba warning, ignore it.
#the umap routine should have completed anyway.
#more about the issue: https://github.com/lmcinnes/umap/issues/252
import umap
try:
drc = drc.drop('x',axis=1)
drc = drc.drop('y',axis=1)
except KeyError:
pass
n_neighbors = 7
embedding = umap.UMAP(n_neighbors = n_neighbors,#small => local, large => global: 5-50
min_dist = 0.0005, #small => local, large => global: 0.001-0.5
spread = 1.5,
metric='cosine').fit_transform(XSVD)
embedding = pd.DataFrame(embedding)
embedding.columns = ['x','y']
```
And next let's plot it to see how it looks like.
```
%%R -i embedding --width 1200 --height 800 -r 140 --bg #F5F5F5
library(hrbrthemes)
library(ggplot2)
library(fields)
embedding$density <- fields::interp.surface(
MASS::kde2d(embedding$x, embedding$y), embedding[,c("x","y")])
p <- ggplot(embedding, aes(x=embedding$x, y=embedding$y,alpha = 1/density))+
guides(alpha=FALSE)+
geom_point(color="#3366cc", pch=16,cex=1.2)+ theme_ipsum_rc()+
labs(x="", y="",
title="The 2d-reduction by UMAP",
subtitle="...based on the code by McInnes, Healy (2018) and Noichl (2018)",
caption="by Anton Savov")+
theme(panel.grid.major = element_line(colour = "lightgrey"),panel.grid.minor = element_blank())+
#comment out the next two lines to see the whole space with all the outliers
scale_x_continuous(limits=c(-25,25),labels=NULL, breaks=seq(-25,25,length=16))+
scale_y_continuous(limits=c(-15,15),labels=NULL, breaks=seq(-15,15,length=10))+
NULL
p
```
## Clustering
For the clustering we use [HDBSCAN](https://github.com/scikit-learn-contrib/hdbscan).
The `clustersize` variable needs to be adjusted based on the number of papers in the data set. For me and my collection of 12,000 records a number between 100 and 200 worked well.
It is important to note that the process of embedding into two dimensions with UMAP is stochastic, i.e. non-deterministic. In plain text that means that each time we run it it will produce slightly different results even though we use the same data. This also has influence on the clustering. In one of the runs I did a cluster size of 100 split the map into 24 clusters, in the next run the same data and cluster size resulted in 17 clusters. So bare that in mind when judging the final result presented here and also if you use this method to produce your own maps.
```
try:
drc = drc.drop('cluster',axis=1)
except KeyError:
pass
import hdbscan
#(min_cluster_size=500, min_samples=30, gen_min_span_tree=True)
#clusterer = hdbscan.HDBSCAN(min_cluster_size=455, min_samples=35, gen_min_span_tree=True)
#adjust this number based on number of papers in your data set
clustersize = 140
clusterer = hdbscan.HDBSCAN(min_cluster_size=clustersize, min_samples=50, gen_min_span_tree=True)
clusterer.fit(embedding)
XCLUST = clusterer.labels_
clusternum = len(set( clusterer.labels_))-1
dfclust = pd.DataFrame(XCLUST)
dfclust.columns = ['cluster']
print(clusternum)
### Let's play a little sound when we're done:
#import winsound
#winsound.Beep(550,300)
```
Now lets plot the map and the clusters in `ggplot`:
```
%%R -i embedding,dfclust -o myNewColors --width 1200 --height 800 -r 140 --bg #F8F4E9
# --bg #231f20
library(hrbrthemes)
library(ggplot2)
library(fields)
library(plyr)
options(warn=0)# 0 zum anschalten
#Get the cluster means:
means <- aggregate(embedding[,c("x","y")], list(dfclust$cluster), median)
means <- data.frame(means)
n=nrow(means)
means <- means[-1,]
#good place to try get consistent colors is http://colorbrewer2.org/
#Make the colors:
mycolors <- c("#6abc70", "#0abcce", "#f9f4b8", "#ffe800", "#fbab19", "#f68121", "#ed1c24", "#ca205f")
pal <- colorRampPalette(sample(mycolors))
s <- n-1
myGray <- c('#aaaaaa')
myNewColors <- sample(pal(s))
myPal <- append(myGray,myNewColors)
#get density, to avoid overplotting
embedding$density <- fields::interp.surface(
MASS::kde2d(embedding$x, embedding$y), embedding[,c("x","y")])
p <- ggplot(embedding, aes(x=embedding$x, y=embedding$y, color= as.factor(dfclust$cluster), alpha = 1/density))+
geom_point(pch=20,cex=1.6)+
theme_ipsum_rc()+
scale_color_manual(values = myPal) +
guides(alpha=FALSE, color=FALSE)+
geom_point(data=means, aes(x=means$x, y=means$y), color= myNewColors, alpha = 1,size =6)+
#annotate("text", x = means[,c("x")], y = means[,c("y")], label = means[,c("Group.1")], color="white", fontface="bold", size=3.2, parse = TRUE, hjust=0.5)+
annotate("text", x = means[,c("x")], y = means[,c("y")], label = means[,c("Group.1")], color="white", fontface="bold", size=3.2, parse = TRUE, hjust=0.5)+
labs(x="", y="",
title="The clusters found by hdbscan...",
subtitle="a density-based clustering algorithm. Embedded with UMAP in two dimensions...")+
theme(panel.grid.major = element_line(colour = "lightgrey", size=0),panel.grid.minor = element_blank())+
#theme(panel.grid.major = element_blank(),panel.grid.minor = element_blank())
#comment out the next two lines to see the whole space with all the outliers
scale_x_continuous(limits=c(-25,25),labels=NULL, breaks=seq(-25,25,length=16))+
scale_y_continuous(limits=c(-15,15),labels=NULL, breaks=seq(-15,15,length=10))+
NULL
p
#pdf("ClusteringUMap.pdf", width = 12, height = 12) # Open a new pdf file
#print(p) #write the plot to the PDF
#dev.off() #close the PDF
```
## Tree of cluster similarity
Now that we have the clusters we can look at which clusters are more similar to each other by taking a look at when did the algorithm found it necessary to split a group into two.
In the chart below, the sooner a cluster broke off of the main trunk, the more disconnected are the articles it contains from the rest of the research body.
The clusters are numbered and later we will analyse the content of each cluster so you can scroll down to the labels to check what body of work is in each cluster.
My observation is that "Shape Grammars" are usually the first cluster to split from the bulk, so probably it means they are the most distinguishable in terms of the features we use (cited authors and cited works) from every other research topic in digital design in architecture.
In general it is interesting to observe in this tree where things split from each other. If we take shape grammars out then what is the next most distinguishable field of research?
### Colours
Next we will generate colours for the cluster from a gradient pallette using the cluster id or the lambda value of when it split off from the main trunk as a sampling variable.
```
#define cluster colours by split lambda value
tree = clusterer.condensed_tree_.to_pandas()
cluster_tree = tree[tree.child_size > 1]
#display(cluster_tree)
from matplotlib import cm
from matplotlib.colors import to_hex
#the reference for the built in linear color maps in matplotlib is at:
# https://matplotlib.org/3.1.1/gallery/color/colormap_reference.html
#cluster_gradient = cm.cool
#cluster_gradient = cm.rainbow
#cluster_gradient = cm.gist_rainbow
#cluster_gradient = cm.jet
#cluster_gradient = matplotlib.colors.LinearSegmentedColormap.from_list("", ["#4D3E71","#7D4495","#AB5596","#DA6688","#F88F79","#FCC696"])
cluster_gradient = matplotlib.colors.LinearSegmentedColormap.from_list(
"",
["#FCC696","#F88F79","#DA6688","#AB5596","#7D4495","#4D3E71"]
)
#cluster_gradient = matplotlib.colors.LinearSegmentedColormap.from_list("", ["#009392","#39b185","#9ccb86","#e9e29c","#eeb479","#e88471","#cf597e"])
#cluster_gradient = cm.hsv
#get the node ids of the clusters
chosen_clusters = clusterer.condensed_tree_._select_clusters()
#get the condensed cluster tree
tree = clusterer.condensed_tree_.to_numpy()
cluster_tree = [x for x in tree if x[3]>1]
#display(cluster_tree)
lambda_vals = [x[2] for x in cluster_tree if x[1] in chosen_clusters]
#display(lambda_vals)
from scipy.interpolate import interp1d
m = interp1d([min(lambda_vals),max(lambda_vals)],[0,1])
#print(m(lambda_vals[1]))
#print(to_hex(cluster_gradient(.5)))
my_cluster_colours = [to_hex(cluster_gradient(m(x))) for x in lambda_vals]
#display(my_cluster_colours)
from rpy2 import robjects
my_cluster_colours_R = robjects.vectors.StrVector(my_cluster_colours)
#print(type(my_cluster_colours_R))
#display(my_cluster_colours_R)
#or, alterntatively, define colors by cluster numbers
from matplotlib import cm
from matplotlib.colors import to_hex, LinearSegmentedColormap
#cluster_gradient = matplotlib.colors.LinearSegmentedColormap.from_list("", ["#4D3E71","#7D4495","#AB5596","#DA6688","#F88F79","#FCC696"])
cluster_gradient = LinearSegmentedColormap.from_list(
"",
["#FCC696","#F88F79","#DA6688","#AB5596","#7D4495","#4D3E71"]
)
#cluster_gradient = matplotlib.colors.LinearSegmentedColormap.from_list("", ["#009392","#39b185","#9ccb86","#e9e29c","#eeb479","#e88471","#cf597e"])
#cluster_gradient = cm.hsv
eqspace = np.linspace(0.0, 1.0, num=17)
my_cluster_colours_2 = [to_hex(cluster_gradient(x)) for x in eqspace]
display(my_cluster_colours_2)
```
Next let's use `SankeyWidget`to plot the cluster tree:
```
from ipysankeywidget import SankeyWidget
from ipywidgets import Layout
from IPython.display import (
Image,
SVG
)
parent_nodes = [x[0] for x in cluster_tree]
links = []
cluster_nodes =[]
nodes = []
removed_parent_nodes = []
first_node = min(parent_nodes)
node = {'id': first_node, 'title': 'All articles' }
nodes.append(node)
for edge in cluster_tree:
if edge[0] in chosen_clusters or edge[0] in removed_parent_nodes:
#filter out leaves that didn't get selected as a cluster
removed_parent_nodes.append(edge[1])
else:
if edge[1] in chosen_clusters:
#this is a cluster leaf so set label, color, etc
cluster_id = chosen_clusters.index(edge[1])
cluster_label = "cluster " + str(cluster_id)
color = my_cluster_colours[cluster_id]
cluster_nodes.append(edge[1])
node = {'id': edge[1], 'title': cluster_label, 'style': 'process' }
else:
#this is a branch so use default color and remove label
color = '#16457D' #"#bfc4c6"
node = {'id': edge[1], 'title': '' }
nodes.append(node)
link = {
'source':edge[0],
'target':edge[1],
'value':edge[3],
'color':color
}
links.append(link)
#display(links)
#display(cluster_nodes)
#use the rank_sets to place all cluster leafs on the same line
rank_sets = [
{ 'type': 'same', 'nodes': cluster_nodes }
]
#use the order to force order by splitting event
#to use pass order=order in the arguments of SankeyWidget
#It creates kind of a wavey chart so I decided not to use it
'''
sorted_nodes = []
for edge in cluster_tree:
for i in [0,1]:
if not edge[i] in removed_parent_nodes and not edge[i] in sorted_nodes:
sorted_nodes.append(edge[i])
order = []
for node in sorted(sorted_nodes):
order.append([node])
'''
#sankey = SankeyWidget(links=links, rank_sets=rank_sets, nodes=nodes, layout=Layout(width="2400", height="400")).auto_save_png('test.png')
sankey = SankeyWidget(links=links, nodes=nodes, layout = Layout(width="1920", height="300")).auto_save_svg('exports/cluster_tree.svg')
sankey
#SVG(sankey.svg)
#sankey.save_svg('test.svg')
```
Here is the result of the last cell:

## What is each cluster about?
Now, let's figure out what body or work does each cluster contain. Lets look at the common words, the cited works and authors for each cluster.
### The most common words and phrases
We can analyze the titles and the abstract of the articles in a cluster and derive the most common words and phrases. Since in the field there are established threee-word-terms such as "Building Information Modeling" it makes sense to look for the most common
And since some article records don't have the abstract info in the data we downloaded from WOS we will do this analysis separate for the titles and the abstract. Plus the abstract have much more words than the titles so their content will drown whatever the titles are about.
But before we do any text analysis lets unify the spelling differences between US and UK English such as "modeling" and "modelling". And we will also remove the most common words such as "paper", as well as the terms that are common accross all clusters such as "design".
```
drc = pd.concat([drc, dfclust],axis=1)
drc = drc.dropna(subset=['cluster'])
drc = pd.concat([drc, embedding],axis=1)
#load up the UK to US spelling dictionary
#the list of words is taken from http://www.tysto.com/uk-us-spelling-list.html
#import random
uk2us = {}
with open('data/uk-to-us-dict.csv', 'r') as csvfile:
spamreader = csv.reader(csvfile, delimiter=';')
for row in spamreader:
ukword = row[0].replace(" ", "")
usword = row[1].replace(" ", "")
uk2us[ukword] = usword
#print(uk2us[random.choice(list(uk2us.keys()))])
#this function will replace words that are keys in mydict with the corresponding value
def replace_all(text, mydict):
for gb, us in mydict.items():
#text = text.replace(us, gb)
text = re.sub(r"\b%s\b" % re.escape(gb) , us, text)
return text
#kick out common abstract words of no importance
removeWords = ['|','paper','argue','account','theory',
'elsevier','reserved','rights',
'used','use','new','using',
'copyright', 'published']
#builds the text bag for each cluster for a given tag
def buildTextBags(drcval, replacedict, removelist, tag, clean=True):
fullstrsl = []
#build a map of the words to remove for later
big_regex = re.compile(r'\b%s\b' % r'\b|\b'.join(map(re.escape, removelist)))
for x in range(0,clusternum):
textBag = list(drcval.loc[drcval['cluster'] == x][tag])
#titles = list(drc.loc[drc['cluster'] == x]['title'])
#abstracts = abstracts + titles
#abstracts = ";".join(str(x) for x in abstracts).replace('|',' ').replace('paper',' ').replace('argue',' ').replace('account',' ').replace('theory',' ').replace('elsevier',' ').replace('reserved',' ').replace('rights',' ').replace('used',' ').replace('use',' ').replace('new',' ').replace('using',' ').replace('process',' ').replace('approach',' ') #kick out common abstract words of no importance
textBag = ";".join(str(x) for x in textBag)
if clean:
textBag = big_regex.sub(" ", textBag)#kick out common abstract words of no importance
#use the word replacing code here https://bytes.com/topic/python/answers/795690-replacing-string-words-list#post3162180
textBag = replace_all(textBag, replacedict) #this slows things down
fullstrsl.append(textBag)
#print(fullstrsl[0][:50])
return fullstrsl
#build the bags - kind of slow for the abstracts
clusterTextBagsTitles = buildTextBags(drc, uk2us, removeWords, 'title')
clusterTextBagsAbstracts = buildTextBags(drc, uk2us, removeWords, 'abstract')
#check if a string searchfor exists in 'searchin' only as a whole word/phrase
def string_found(searchfor, searchin):
return re.search(r'\b%s\b' % (re.escape(searchfor)), searchin) is not None
#if a phrase is found in a longer phrase reduce its score with the score of hte longer phrase
#exclude if true the word score is set to 0 if exclude is false then the longer word score is subtracted from short word score
def accountScores(longphrasescores, shortphrasescores, exclude=True):
newscores =[]
for phrase in shortphrasescores:
newphraseScore = [phrase[0],phrase[1]]
for longphrase in longphrasescores:
#if phrase is found in longphrase subtract scores
#pat = re.compile(r"\b%s\b" % re.escape(phrase[0]))
#print(phrase[0])
if string_found(phrase[0], longphrase[0]):
if exclude:
newphraseScore[1] = 0
else:
newphraseScore[1] = phrase[1]-longphrase[1]
newscores.append(newphraseScore)
return newscores
#this def below is not needed anymore
def accountScoreClusters(longscores, shortscores):
counter =0
newclusterscores =[]
for cluster in shortscores:
newclusterscores.append(accountScores(longscores[counter],cluster))
counter +=1
return newclusterscores
#tag is which tag('abstract' or 'title') to analyze for
#phrase is the phrase length in words. use 2 for bigrams;
#n is how many words to output
#treshold is the minimum number of occurences needed to include the word
#clean if True will skip the words in the removeWords list
#longphrasescores is a list of pairs
# first item is scorelist for already ranked words/phrases
# second item if true the word score is set to 0 if exclude is false then the longer word score is subtracted from short word score
#longphrasescores is used for example to reduce the score of the bigram "building information" with the score of the trigram "building information modeling", i.e. to avoid double counting
def rankWords(textBag,phrase=1, n=10,treshold=10, longphrasescores=None):
vec = CountVectorizer( stop_words='english', ngram_range=(phrase, phrase))#Choose CountVectorizer for the most common words in the cluster, TfidfVectorizer for the words with the greatest differentiation value.
X = vec.fit_transform(textBag)
#print(pd.DataFrame(X.toarray(), columns=vec.get_feature_names())) #To look into the vectors. Beware, can take a bit of RAM
clusterfeatures = pd.DataFrame(X.toarray(), columns=vec.get_feature_names())
fullscore = []
for x in range(0,clusternum):
scores = zip(vec.get_feature_names(), np.asarray(X[x,:].sum(axis=0)).ravel())
accountedScores = sorted(scores, key=lambda x: x[1], reverse=True)
accountedScores = accountedScores[0:60]
#subtract scores if phrase is found in longphrasescores
if longphrasescores is not None:
for longphrasescore in longphrasescores:
accountedScores = accountScores(longphrasescore[0][x], accountedScores,longphrasescore[1])
#sort
sorted_scores = sorted(accountedScores, key=lambda x: x[1], reverse=True)
myscores = sorted_scores[0:n]
#print(myscores)
#print("___________________")
scorelist = []
counter = 0
for s in myscores:
#print(s)
if s[1]>treshold or counter == 0:
#newWord = s[0] + ' ('+str(s[1]) + ')' # do the word and the how often it comes
newWord = [s[0],s[1]] # do the word and the how often it comes
scorelist.append(newWord)
#else:
# newWord = 'x' # skip the word if it appears less than 10 times
counter += 1
fullscore.append(scorelist)
return fullscore
#adds to dataframes of scores for one cluster together
def addClusterScores(score1,score2):
newscore = []
for clusterid in range(0,len(score1)):
clusterscore =[]
for item in score1[clusterid]:
clusterscore.append(item)
for item in score2[clusterid]:
clusterscore.append(item)
newscore.append(clusterscore)
return newscore
print("TOP 5 TRIGRAMS ABSTRACTS")
trigramScoreAbstracts = rankWords(clusterTextBagsAbstracts,phrase=3)
display(pd.DataFrame(trigramScoreAbstracts).transpose().head())
print("TOP 5 TRIGRAMS TITLES")
trigramScoreTitles = rankWords(clusterTextBagsTitles,phrase=3,longphrasescores=[[trigramScoreAbstracts,True]])
display(pd.DataFrame(trigramScoreTitles).transpose().head())
print("TOP 5 TRIGRAMS COMBINED")
trigrams = addClusterScores(trigramScoreAbstracts,trigramScoreTitles)
display(pd.DataFrame(trigrams).transpose().head())
print("TOP 5 BIGRAMS ABSTRACTS")
bigramScoreAbstracts = rankWords(clusterTextBagsAbstracts,phrase=2, n=5,longphrasescores=[[trigrams,False]])
#bigramScoreAbstracts = accountScoreClusters(trigramScoreAbstracts, bigramScoreAbstracts)
display(pd.DataFrame(bigramScoreAbstracts).transpose().head())
print("TOP 5 BIGRAMS TITLES")
bigramScoreTitles = rankWords(clusterTextBagsTitles,phrase=2, n=5,longphrasescores=[[trigrams,False],[bigramScoreAbstracts,True]])
#print(bigramScoreTitles[:10])
#bigramScoreTitles = accountScoreClusters(trigramScoreTitles, bigramScoreTitles)
#print(bigramScoreTitles[:10])
display(pd.DataFrame(bigramScoreTitles).transpose().head())
print("TOP 5 BIGRAMS COMBINED")
bigrams = addClusterScores(bigramScoreAbstracts,bigramScoreTitles)
display(pd.DataFrame(bigrams).transpose().head())
print("TOP 5 WORDS IN ABSTRACTS")
wordScoreAbstracts = rankWords(clusterTextBagsAbstracts,phrase=1, n=20, longphrasescores=[[trigrams,False],[bigrams,False]])
display(pd.DataFrame(wordScoreAbstracts).transpose().head())
print("TOP 5 WORDS IN TITLES")
wordScoreTitles = rankWords(clusterTextBagsTitles,phrase=1, n=20, longphrasescores=[[trigrams,False],[bigrams,False],[wordScoreAbstracts,True]])
display(pd.DataFrame(wordScoreTitles).transpose().head())
words = addClusterScores(wordScoreAbstracts,wordScoreTitles)
#display(pd.DataFrame(words).transpose())
#common words in the field to skip only in the single words lists
# ideally this should be automated to kick out all words that are occuring in more than 40-50% of clusters
commonFieldTerms = [
'method', 'methods', 'process', 'design', 'designs',
'research', 'based', 'approach',
'information', 'architecture', 'architectural',
'digital', 'development', 'building',
'model', 'models', 'modeling',
'result', 'results', 'study', 'data',
'proposed', 'analysis', 'different',
'space', 'spaces', 'spatial', 'bv'
]
def filterScoreList(scorelist, commonTerms):
newscorelist = []
for wscore in scorelist:
#print(wscore)
if not any(wscore[0] in s for s in commonTerms):
newscorelist.append(wscore)
return newscorelist
filteredwords = []
for wordscores in words:
filteredwords.append(filterScoreList(wordscores, commonFieldTerms))
print("TOP 5 WORDS")
display(pd.DataFrame(filteredwords).transpose().head())
```
Now let's combine all words and phrases into the cluster labels.
```
#import textwrap
allscores = addClusterScores(trigrams,bigrams)
allscores = addClusterScores(allscores,filteredwords)
display(allscores)
clusterlabels = []
clusterlabelsrows = []
clusterlabels_textblock = []
#clusterlabelsb = []
counter = 0
linelength = 140
for x in range(0,len(trigrams)):
#clustertext = "TRIGRAMS: " + textwrap.fill("; ".join([ (w[0]+'('+str(w[1])+')') for w in trigrams[x] if w[1]>10 or trigrams[x].index(w)==0]),linelength) + '\n'
#clustertext = clustertext + "BIGRAMS: " + textwrap.fill("; ".join([ (w[0]+'('+str(w[1])+')') for w in bigrams[x] if w[1]>10]),linelength)+ '\n'
#clustertext = clustertext + "WORDS: " + textwrap.fill("; ".join([ (w[0]+'('+str(w[1])+')') for w in filteredwords[x] if w[1]>10]),linelength)
trigramslabel = "; ".join([ (w[0]+' ('+str(w[1])+'x)') for w in trigrams[x] if w[1]>10 ]) #and trigrams[x].index(w)<5])
bigramslabel = "; ".join([ (w[0]+' ('+str(w[1])+'x)') for w in bigrams[x] if w[1]>10 ]) #and bigrams[x].index(w)<5])
wordslabel = "; ".join([ (w[0]+' ('+str(w[1])+'x)') for w in filteredwords[x] if w[1]>10 ]) #and filteredwords[x].index(w)<20])
#clustertext = "OFTEN USED TERMS: " + trigramslabel + ("; " if len(trigramslabel)>0 else "") + bigramslabel + ("; " if len(bigramslabel)>0 else "") + wordslabel
clustertext = trigramslabel + ("; " if len(trigramslabel)>0 else "") + bigramslabel + ("; " if len(bigramslabel)>0 else "") + wordslabel
clusterlabels_textblock.append(clustertext)
clustertext = textwrap.fill(clustertext,linelength)
clusterlabels.append(clustertext)
#this is how many lines are in this cluster's label
clusterlabelsrows.append(len(clustertext.split('\n')))
#clusterlabelsb.append("")
print("cluster: " + str(counter))
#print(textwrap.fill(clustertext,70))
print(clustertext)
print("--------------------")
counter +=1
display(pd.DataFrame(clusterlabels_textblock))
#print(clusterlabels)
#display(clusterlabels_textblock)
```
### What are the most cited papers in each cluster
Next we find the most cited papers for each cluster. This will give us the seminal works that everyone working that field always refers to. For example in the Shape Grammars cluster we have the [The Palladian Grammar](https://journals.sagepub.com/doi/abs/10.1068/b050005) by Stiny and Mitchel from 1978.
```
#THIS WILL GET US THE TOP CITED PAPERS PER CLUSTER
#fullstrsl = []
citationsDict = [None]*clusternum #per cluster
fullscoreTopCitations = []
for x in range(0,clusternum):
citations = list(drc.loc[drc['cluster'] == x]['citestring'])
citationsDict[x] = {}
citations = [item for sublist in citations for item in sublist]
for citation in citations:
if not citation in citationsDict[x]:
citationsDict[x][citation] = [citation,1]
else:
citationsDict[x][citation][1] += 1
#
rankedCitations = []
for citation in citationsDict[x]:
rankedCitations.append([citationsDict[x][citation][0],citationsDict[x][citation][1]])
sorted_scores = sorted(rankedCitations, key=lambda x: x[1], reverse=True)
myscores = sorted_scores[0:5]
scorelist = []
for s in myscores:
if s[1] > 1:
scorelist.append( [s[0],s[1]]) # do the word and the how often it comes)
#scorelist.append(s[0]+ ' ('+str(s[1]) + ')') # do the word and the how often it comes)
fullscoreTopCitations.append(scorelist)
print("TOP 5 CITATIONS")
display(pd.DataFrame(fullscoreTopCitations).transpose().head())
```
### Most cited authors in a cluster
Besides the individual works it is also very revealing who defines and drives the research in each cluster if we get a list of the most cited authors. So let's do that next. Sometimes the same author is logged differently in the WOS database, for example "Gramazio Fabio" and "Gramazio F". So first we will unify the authors list and then count the most popular.
```
#cases for merging A to author B
#1. "gramazio fabio" and "gramazio f" or "knight t" and "knight tw"
#if both have two words, word[0] is same, word[1] has same first letter, name them with the longer second word
#2. "simon ha" and "simon h a"
#if A has 3 words (with 2nd and 3rd single letter)and B has 2 words (2nd of 2 letter) or vice versa and word[0] is same and word[1] same first letter and word[2] of the one who has it is same letter as the other one's word[1]'s second letter
#3. "stiny g" and "stiny g n"
##if A has 3 words (with 2nd and 3rd single letter)and B has 2 words (2nd of 1 letter) or vice versa and word[0] is same and word[1] same first letter
def mergeSameAuthors(authorscorelist):
newscorelist = []
mergedict = {}
for author in authorscorelist:
#print(author)
if not 'none' in author[0] and not "".join(author[0].split()) == '':
names = author[0].split()
#print(names)
newscore = [author[0],author[1]]
merged = False
if not names[0] in mergedict:
mergedict[names[0]] = []
#print(newscore)
#mergedict[names[0]].append(newscore)
else:
for i in range(0, len(mergedict[names[0]])):
existingnames = mergedict[names[0]][i][0].split()
if len(names) >= 2 and len(existingnames) >= 2:
#case 1. and 2. and 3. from above
if names[1][0] == existingnames[1][0]:
#print("merging: (" + author[0] + ") into (" + mergedict[names[0]][i][0]) + ")"
mergedict[names[0]][i][1] = mergedict[names[0]][i][1] + author[1]
merged = True
if not merged:
#print("appending: " + newscore[0])
mergedict[names[0]].append(newscore)
for familynamelist in mergedict:
#print(familynamelist)
for author in mergedict[familynamelist]:
#print(author)
newscorelist.append(author)
return newscorelist
fullstrsl = []
for x in range(0,clusternum):
authors = list(drc.loc[drc['cluster'] == x]['citedAU'])
authors = [item for sublist in authors for item in sublist]
#authors = authors + list(drc.loc[drc['cluster'] == x]['author'])
authors = " §".join(str(x) for x in authors)
#print(authors[:50])
#changed here from 2 to 0 because it otherwise combines different authors with same family name as one
#especially important for chinese names where family name such as "wang" is very common
#two thing can be done here
#1. merge entries for same author when he is listed with full name and family plus initial
# such as "gramazio fabio" and "gramazio f"
#2. merge entries for same author when he is listed with first initial joined and not
# such as "simon ha" and "simon h a"
#3. (optional as it might be wrong to do it) merge entries when author is listed with one and then with two initial
# such as "stiny g" and "stiny g n"
#this might be done as follows
#authors = ' '.join( [w for w in authors.split() if len(w)>2] )
authors = ' '.join( [w for w in authors.split()])# if len(w)>0] )
#print(authors[:50])
fullstrsl.append(authors)
#print(fullstrsl[1])
vec = CountVectorizer(token_pattern=r'[\s\w\.-]+(?=[$|§])')#Choose CountVectorizer for the most common words in the cluster, TfidfVectorizer for the words with the greatest differentiation value.
X = vec.fit_transform(fullstrsl)
clusterfeatures = pd.DataFrame(X.toarray(), columns=vec.get_feature_names())
fullscoreA = []
for x in range(0,clusternum):
scores = zip(vec.get_feature_names(), np.asarray(X[x,:].sum(axis=0)).ravel())
sorted_scores = sorted(scores, key=lambda x: x[1])
#print(sorted_scores[0:6])
sorted_scores = mergeSameAuthors(sorted_scores)
#print(sorted_scores[0:6])
sorted_scores = sorted(sorted_scores, key=lambda x: x[1], reverse=True)
myscores = sorted_scores[0:10]
scorelist = []
for s in myscores:
if s[1] > 1:
scorelist.append( [s[0],s[1]]) # do the word and the how often it comes)
#scorelist.append(s[0]+ ' ('+str(s[1]) + ')') # do the word and the how often it comes)
fullscoreA.append(scorelist)
print("TOP 5 CITED AUTHORS")
display(pd.DataFrame(fullscoreA).transpose().head())
```
Great, now we have a list of authors for each cluster. Let's also count the number of articles per cluster
```
#this counts how many articles are in each cluster
articleCount = []
totalArticleCount = len(drc)
print(totalArticleCount)
for x in range(0,clusternum):
articles = len(drc.loc[drc['cluster'] == x])
articleCount.append(articles)
print(articles)
```
Let's assemble the text for the sublabels for each cluster from the most cited works, most cited authors and the number of articles.
```
clusterlabelsb = []
clusterlabelsb_textblock = []
counter = 0
cluster_authors = []
cluster_citations = []
linelength = 110
#fullscoreTopCitations
for x in range(0,len(fullscoreA)):
articleCountText = str(articleCount[x]) + ' articles'
authors = "\n".join( [ (w[0].split()[0].upper() + ',' + w[0].split()[1].upper()+' ('+str(w[1])+'x)') for w in fullscoreA[x]])
cluster_authors.append(authors)
clustertext = "MOST CITED AUTHORS: " + "; ".join( [ (w[0].split()[0].upper() + ',' + w[0].split()[1][0].upper()+'. ('+str(w[1])+'x)') for w in fullscoreA[x] ] )
citations = "\n".join([ (w[0].upper() + ' ('+str(w[1])+'x)') for w in fullscoreTopCitations[x]])
cluster_citations.append(citations)
topcitationstext = "TOP CITATIONS: " + "; ".join([ (w[0].upper() + ' ('+str(w[1])+'x)') for w in fullscoreTopCitations[x]])+''
#clustertext = clustertext + "Words: " + textwrap.fill("; ".join([ (w[0]+'('+str(w[1])+')') for w in filteredwords[x] if w[1]>10]),linelength)
#clusterlabels[x] = clusterlabels[x]+ '\n' + clustertext
clusterlabelsb_textblock.append(articleCountText + "\n" + clustertext + "\n" + topcitationstext)
clustertext = textwrap.fill( clustertext,linelength)
#clustertext = articleCountText + "\n" + clustertext + "\n" + textwrap.fill(topcitationstext,linelength) #.upper()
clustertext = clustertext + "\n" + textwrap.fill(topcitationstext,linelength) #.upper()
clusterlabelsb.append(clustertext)
#clusterlabelsb.append("")
print("cluster: " + str(counter))
#print(textwrap.fill(clustertext,70))
print(clustertext)
print("--------------------")
counter +=1
#display(clusterlabelsb_textblock)
display(cluster_authors)
with open('exports/cluster_words.txt', 'w') as f:
f.writelines("%s\n" % line for line in clusterlabels_textblock)
with open('exports/cluster_cited.txt', 'w') as f:
f.writelines("%s\n" % line for line in clusterlabelsb_textblock)
```
## Plot the map
Finally we have all the information to plot the map of articles with the clusters and their labels. For plotting we use `ggplot2` from `R` as before. It gives us a raw-looking graphic but we can export as PDF and edit it further with Affinity Designer, Adobe Illustrator or Inkscape.
```
%%R -i embedding,dfclust,my_cluster_colours_R,clusterlabels,clusterlabelsb,journalListLabel -w 594 -h 420 --units mm -r 300 --bg #F5F5F2
#--bg #d3d3d3
#-o labelpol,cltest --bg #231f20
#-h 1600 -w 1600 -r 140 --bg #fbf8f1
# Some imports:
library(hrbrthemes)
library(ggplot2)
library(fields)
#library(ggrepel) #was off in original from Noichl, used for geom_label_repel
library(ggforce)
#install.packages('ggalt')
library(ggalt)
library(stringr)
#install.packages('clue')
#library(clue)
options(warn=0)# 0 zum anschalten
#Get the cluster means:
means <- aggregate(embedding[,c("x","y")], list(dfclust$cluster), mean)
means <- data.frame(means)
#And Variance, for the labels:
test <- aggregate(embedding[,c("x")], list(dfclust$cluster), var)
test <- test[-1,]
n=nrow(means)
means <- means[-1,]
#Make the colors:
#mycolors <- c("#6abc70", "#0abcce", "#f9f4b8", "#ffe800", "#fbab19", "#f68121", "#ed1c24", "#ca205f")
#mycolors <- c("#4eeadf","#e84a31","#cc2ee7","#bbe953","#50aeec")
#saturated from ColorBrewer
#mycolors <- c("#a6cee3","#1f78b4", "#b2df8a", "#33a02c", "#fb9a99", "#e31a1c", "#fdbf6f","#ff7f00", "#cab2d6","#6a3d9a", "#ffff99","#b15928")
#pastels from ColorBrewer
#mycolors <- c("#8dd3c7", "#ffffb3", "#bebada", "#fb8072", "#80b1d3", "#fdb462", "#b3de69", "#fccde5", "#d9d9d9", "#bc80bd", "#ccebc5", "#ffed6f")
#pal <- colorRampPalette(sample(mycolors))
s <- n-1
#myGray <- c('#858583') # c('#95a5a6') # for the points outside of a cluster
myGray <- c('#16457D') # ink blue
#myNewColors <- sample(pal(s))
myNewColors <- my_cluster_colours_R
myPal <- append(myGray,myNewColors)
labels <- clusterlabels
#labelsrows <- clusterlabelsrows
labelsb <- clusterlabelsb
#circular markers:
library(gridExtra)
circle <- polygon_regular(100)
pointy_points <- function(x, y, size){
do.call(rbind, mapply(function(x,y,size,id)
data.frame(x=size*circle[,1]+x, y=size*circle[,2]+y, id=id),
x=x,y=y, size=size, id=seq_along(x), SIMPLIFY=FALSE))
}
#get density, to avoid overplotting
embedding$density <- 1/ as.numeric(fields::interp.surface(MASS::kde2d(embedding$x, embedding$y), embedding[,c("x","y")]))
# get for every label, whether it is in the + or - part of the x-axis:
xpol <-abs(means[,c("x")])/means[,c("x")]
ypol <-abs(means[,c("y")])/means[,c("y")]
polfact <- 1.5
#define label coordinates
# build a circle for the labels:
r <- 14.5
sequence <- seq(from = 1, to = s, by = 1)
angles <- 360/s*sequence
angle <-(angles*(pi/180))
xlabl <- cos(angle)*r
ylabl <- sin(angle)*r
circlecord <- cbind(xlabl,ylabl)
labelpol <-abs(circlecord[,c("xlabl")])/circlecord[,c("xlabl")]
# install.packages("stringr")
# library(stringr)
#make the label coordinates on two lines left and right of the plot
halfclust = (s+1)%/%2
col1 <- data.frame(xlabl=rep(-15.5,halfclust),ylabl=seq(-13,13,length=halfclust))
col2 <- data.frame(xlabl=rep(15.5,s-halfclust),ylabl=seq(-13,13,length=s-halfclust))
linecoords <- rbind(col1,col2)
#make the label coordinates on one line on the right side of the plot
linecoords <- data.frame(xlabl=rep(20.5,s),ylabl=seq(-19.5,19.5,length=s))
circlecord <- linecoords
coord_x=5
coord_y=5
# define circular markers:
circular_annotations <- pointy_points(means$x, means$y, size=test$x*0.25+1)
#
embedding <- cbind(embedding,dfclust)
filtered <- as.data.frame(subset(embedding, cluster >= 0))
cltest <- filtered
#Let's plot!
mapdotshape = 16 # originally this was 20 (circle with fill and border). with 3(cross symbol) it takes a lot of time to render
#this plots the points that belong to a cluster
p <- ggplot(data=filtered, aes(x=x, y=y, color= as.factor(cluster),alpha=0.5))+ #, alpha='density'))+
#pch=20 is bullet shape - https://stat.ethz.ch/R-manual/R-patched/library/graphics/html/points.html
geom_point(pch=mapdotshape,cex=1.5)+#, alpha = 0/density)+
#scale_color_manual(values = myPal)+ #this applies our colours to the cluster points
scale_x_continuous(limits=c(-25,25),labels=NULL, breaks=seq(-25,25,length=16))+
scale_y_continuous(limits=c(-15,15),labels=NULL, breaks=seq(-15,15,length=10))+
#the lines below were disabled
#geom_polygon(data=circular_annotations, aes(x,y,group=factor(id), fill = factor(id)),alpha=0.15)+
#scale_fill_manual(values = myNewColors) #+
#this hides the legends
#guides(alpha=FALSE, color=FALSE, fill=FALSE) #+
#
#this creates the coloured spots over the clusters that encircle them - takes time
#stat_density2d(n=800,h=c(1.1,1.1))
#
xlim(-15, 50) +
ylim(-20, 20) +
NULL
#end of the points plotting
#ggplot_build prepares the plot for rendering - https://ggplot2.tidyverse.org/reference/ggplot_build.html
#and also adds to p the coloured spots over the clusters by creating density contours around them
q <- ggplot_build(p+stat_density2d(n=400,h=c(1.1,1.1)))$data[[2]]
#looks like this filters out all points except those belonging to group 001 which is the first, outmost density contour
#there is a problem because if first contour is comprised of several closed curves only one of them ends up being shown on the plot
q <- q[str_detect(q$group, "001") == TRUE, ]
# z <- max(test$x)
# print(z)
#this finds the point with minimum x for each cluster in the plot render data and gets its data in z
o <- aggregate(q$x, list(q$group) , min)
z <- subset(q,subset = q$x %in% c(o$x))
#this finds the point with maximum x for each cluster in the plot render data and gets its data in zmax
omax <- aggregate(q$x, list(q$group) , max)
zmax <- subset(q,subset = q$x %in% c(omax$x))
#this creates a dataframe with the points with min or max depending on which side of the origin each cluster lies
c <- data.frame()
count <- 1
for (val in xpol) {
if(val <0 ){
c <- rbind(c, z[count,])
} else {
c <- rbind(c, zmax[count,])
}
count <- count + 1
}
#these are the x and y coords of those points (leftmost or right most per cluster)
#contactpoints <- data.frame(c$x,c$y)
contactpoints <- data.frame(zmax$x,zmax$y)
#Append every label to its best fit on that circle, using the hungarian algorithm:
require(clue)
#this gives all possible distances between the two sets of points, those on the circle and those in the cluster
distances <- rdist(circlecord,contactpoints)
#this give the indices of the points on the circle that fit closest to each cluster point
sol <- solve_LSAP(t(distances))
#this builds dataframe with the coordinates of each set of two matching points (cluster to circle)
solo <- data.frame(cbind(mx=(contactpoints[,1]), my=(contactpoints[,2]), cx=(circlecord[sol, 1]), cy=(circlecord[sol, 2])))
xcpol <-abs(solo$cx)/solo$cx
#imprts the roboto condensed font for use in theme_ft_rc later
#import_roboto_condensed()
#import_titillium_web()
#this block produces the final graph using the one produced by p as the base
r <-
p +
#this adds the points outside of a cluster
geom_point(data=subset(embedding, cluster == -1), aes(x=x, y=y),pch=mapdotshape,cex=1.5,alpha=0.5, color=myGray)+
#this draws the polygons blobby coloured spots around each cluster. it uses the precalculated data in q
#geom_polygon(data=q, aes(x,y ,group = as.factor(q$group),fill = as.factor(q$group)),alpha=0.3,linetype=1,size=0.6)+ # ,linetype=3, color="black"
#geom_polygon(data=q, aes(x,y ,group = as.factor(cluster),fill = as.factor(cluster)),alpha=0.3,linetype=1,size=0.6)+ # ,linetype=3, color="black"
#this draws the iso contours around the clusters in colour
stat_density2d(aes(colour=as.factor(cluster)),n=400,h=c(1.1,1.1),linetype=1,size=0.6,alpha=0.3)+#,color="black")+
#this draws the iso contours around the clusters in black
#stat_density2d(aes(colour="black"),n=400,h=c(1.1,1.1),linetype=1,size=0.6,alpha=0.3)+#,color="black")+
##scale_fill_manual(values = myPal)+
#this is a built in theme in hrbrthemes - https://www.rdocumentation.org/packages/hrbrthemes/versions/0.6.0/topics/theme_ipsum
theme_ipsum()+
#theme_ipsum_tw()+
#theme_modern_rc()+
## scale_fill_manual(values = myPal)
#this hides the legends
guides(alpha=FALSE, color=FALSE, fill=FALSE)+
#
labs(x=NULL, y=NULL, #x="UMAP-x", y="UMAP-y",
title="DIGITAL DESIGN IN ARCHITECTURE",
subtitle="RESEARCH LANDSCAPE MAP 1975-2019\nwith data from ~12,000 articles\nmapped by who cited who",
caption=paste("BY ANTON SAVOV"))+
#set parameters for the plot title
theme(plot.title = element_text(size=20, hjust = 0.5,colour = myGray))+ #, family="SC", face="plain"
#set parameters for the plot subtitle
theme(plot.subtitle = element_text(size=13, hjust = 0.5,colour = myGray))+ #, family="SC", face="plain"
#this modifies the grid lines
#theme(panel.grid.major = element_line(colour = "grey", linetype="dotted", size=0.55),panel.grid.minor = element_blank())+
theme(panel.grid.major = element_line(colour = myGray, linetype="dotted", size=0.55),panel.grid.minor = element_blank())+
#this sets the graph background to our background colour
theme(plot.background = element_rect(fill = "#F5F5F2"))+
#use expand_limits to create some space for more text
#https://www.rdocumentation.org/packages/ggplot2/versions/3.1.0/topics/expand_limits
#doesnt seem to affect the plot because in p we used scale_x_continuous and scale_y_continuous to set limits
#expand_limits(x = c(-30,30),y = c(-30,30))+
#expand_limits(x = c(r+10,0-r-10),y = c(r+10,0-r-10))+
#fixes x scale to y scale in 1:1 by default - https://www.rdocumentation.org/packages/ggplot2/versions/3.1.0/topics/coord_fixed
coord_fixed()+
#creates an annotation - text or geometry - https://www.rdocumentation.org/packages/ggplot2/versions/3.1.0/topics/annotate
#this one in particular does the little horizontal line extending from each cluster towards it label
#annotate("segment", x = c$x, y = c$y, xend = c$x+xcpol*0.5, yend =c$y, color= myNewColors,alpha=0.3, size = 1)+
annotate("segment", x = zmax$x, y = zmax$y, xend = zmax$x+0.5, yend =zmax$y, color= myNewColors,alpha=0.3, size = 1)+
#and this one does the diagonal connecting lines starting at the end of the short one from above and ending in the label dots
#annotate("segment", x = c$x+xcpol*0.5, y =c$y, xend = solo$cx, yend = solo$cy, color=myNewColors,alpha=0.3, size = 1)+
annotate("segment", x = zmax$x+0.5, y =zmax$y, xend = solo$cx, yend = solo$cy, color=myNewColors,alpha=0.3, size = 1)+
#this plots the dots where the label text will be, pch=16 is circle, pch=22 is square
geom_point(data=solo, aes(x=cx, y=cy), color= myNewColors, alpha = 1,pch=16,size=7, stroke = 1)+
#this labels the dots with the cluster number
#annotate("text", x = solo$cx, y = solo$cy, label = means[,c("Group.1")], color="white", fontface="bold", size=3.2, parse = TRUE, hjust=0.5)+
annotate("text", x = solo$cx, y = solo$cy, label = means[,c("Group.1")], color="black", fontface="bold", size=3.2, parse = TRUE, hjust=0.5)+
## theme(plot.margin=grid::unit(c(0,0,0,0), "mm"))+
#some left over attempt at the diagonal lines
##annotate("segment", x = (solo$mx+(test$x*0.25+1)*xcpol)+xcpol*0.3, y = solo$my, xend = solo$cx, yend = solo$cy, color= myNewColors, alpha = 0.25, size = 0.7)+
#this adds the labels, still dont know what parse means - +0.19
#annotate("text", x = solo$cx+xcpol*0.3, y = solo$cy, parse = FALSE, label = labels, color=myGray, fontface="bold", size=2, hjust=abs((xcpol+1)/2-1),vjust=0)+
annotate("text", x = solo$cx+0.7, y = solo$cy, parse = FALSE, label = labels, color=myGray, fontface="plain", size=2, hjust=0,vjust=0.8)+
#this adds the sublabels - +0.3-0.5
#annotate("text", x = solo$cx+xcpol*0.3, y = solo$cy-0.1, parse = FALSE, label = labelsb, color=myGray, fontface="italic", size=2, hjust=abs((xcpol+1)/2-1),vjust=1)+
annotate("text", x = solo$cx+17.2, y = solo$cy, parse = FALSE, label = labelsb, color=myGray, fontface="plain", size=2, hjust=0,vjust=0.8)+
#could get this repel thing to work
#geom_label_repel(data = c, aes(label = labels,
# fill = myNewColors), color = 'white',
# size = 1)
#marks the cluster attachment points for the label leaders with black dots
#geom_point(data=c, aes(x,y),color="black")+
#this sets our colours to the cluster dots
scale_color_manual(values = myNewColors)+
#this sets our colours to the cluster blobs
scale_fill_manual(values = myNewColors)+
#this sets the background to transparent
#theme(plot.background=element_rect(fill=NA, colour=NA))+
#this adds the label with the journals list - for vjust ref - https://stackoverflow.com/questions/7263849/what-do-hjust-and-vjust-do-when-making-a-plot-using-ggplot
#x = -14.5, y = -14
annotate("text", x = -25, y = 8, parse = FALSE, label = journalListLabel, color=myGray, fontface="plain", size=2, hjust=0,vjust=0)+
NULL
#end of the plotting function of the cluster map
#library(gridExtra)
#grid.arrange(p,t, ncol = 1, heights = c(1, 1))
ggsave('exports/map-raw.png', plot = r,width=594,height=420, units = "mm",dpi=300, bg = "#F5F5F2")
#pdf("exports/ClusteringUMap.pdf",width=594/25.4,height=420/25.4) #,encoding='ISOLatin2.enc')#, paper='a4')# width = 12, height = 12) # Open a new pdf file
#print(r)
#dev.off()
## my output commands to understand what is what
r
#print(n)
#class(means) # get class
#sapply(means, class) # get class of all columns
#str(means) # structure
#summary(means) # summary of dataframe
#print(str(distances)) #
#print(str(sol)) #
#print(str(solo)) #
#print(str(xcpol)) #
#print(str(c)) #
#print(head(test))
#fix(sol) # view spreadsheet like grid
#rownames(means) # row names
#colnames(means) # columns names
#nrow(means) # number of rows
#ncol(means) # number of columns
#ggsave('plot-test.png', plot = last_plot(),width=594,height=420, units = "mm",dpi=300)
```
Here is the image that is direclty output from the script above.

# RESULT
And here is again the image from the very top. The final version after the improvements done with Affinity Designer. Note that I used the algortihmically derived info for each cluster to give it a title. The titles are my interpretation and you can agree or disagree with them. In any case I would love to hear from you, email me at antonsavov@gmail.com or open an issue here.

| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Transfer learning with TensorFlow Hub
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/tutorials/images/transfer_learning_with_hub"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/tutorials/images/transfer_learning_with_hub.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/images/transfer_learning_with_hub.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/tutorials/images/transfer_learning_with_hub.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
[TensorFlow Hub](http://tensorflow.org/hub) is a way to share pretrained model components. See the [TensorFlow Module Hub](https://tfhub.dev/) for a searchable listing of pre-trained models. This tutorial demonstrates:
1. How to use TensorFlow Hub with `tf.keras`.
1. How to do image classification using TensorFlow Hub.
1. How to do simple transfer learning.
## Setup
```
from __future__ import absolute_import, division, print_function, unicode_literals
import matplotlib.pylab as plt
try:
# %tensorflow_version only exists in Colab.
!pip install tf-nightly
except Exception:
pass
import tensorflow as tf
!pip install -U tf-hub-nightly
!pip install tfds-nightly
import tensorflow_hub as hub
from tensorflow.keras import layers
```
## An ImageNet classifier
### Download the classifier
Use `hub.module` to load a mobilenet, and `tf.keras.layers.Lambda` to wrap it up as a keras layer. Any [TensorFlow 2 compatible image classifier URL](https://tfhub.dev/s?q=tf2&module-type=image-classification) from tfhub.dev will work here.
```
classifier_url ="https://tfhub.dev/google/tf2-preview/mobilenet_v2/classification/2" #@param {type:"string"}
IMAGE_SHAPE = (224, 224)
classifier = tf.keras.Sequential([
hub.KerasLayer(classifier_url, input_shape=IMAGE_SHAPE+(3,))
])
```
### Run it on a single image
Download a single image to try the model on.
```
import numpy as np
import PIL.Image as Image
grace_hopper = tf.keras.utils.get_file('image.jpg','https://storage.googleapis.com/download.tensorflow.org/example_images/grace_hopper.jpg')
grace_hopper = Image.open(grace_hopper).resize(IMAGE_SHAPE)
grace_hopper
grace_hopper = np.array(grace_hopper)/255.0
grace_hopper.shape
```
Add a batch dimension, and pass the image to the model.
```
result = classifier.predict(grace_hopper[np.newaxis, ...])
result.shape
```
The result is a 1001 element vector of logits, rating the probability of each class for the image.
So the top class ID can be found with argmax:
```
predicted_class = np.argmax(result[0], axis=-1)
predicted_class
```
### Decode the predictions
We have the predicted class ID,
Fetch the `ImageNet` labels, and decode the predictions
```
labels_path = tf.keras.utils.get_file('ImageNetLabels.txt','https://storage.googleapis.com/download.tensorflow.org/data/ImageNetLabels.txt')
imagenet_labels = np.array(open(labels_path).read().splitlines())
plt.imshow(grace_hopper)
plt.axis('off')
predicted_class_name = imagenet_labels[predicted_class]
_ = plt.title("Prediction: " + predicted_class_name.title())
```
## Simple transfer learning
Using TF Hub it is simple to retrain the top layer of the model to recognize the classes in our dataset.
### Dataset
For this example you will use the TensorFlow flowers dataset:
```
data_root = tf.keras.utils.get_file(
'flower_photos','https://storage.googleapis.com/download.tensorflow.org/example_images/flower_photos.tgz',
untar=True)
```
The simplest way to load this data into our model is using `tf.keras.preprocessing.image.ImageDataGenerator`,
All of TensorFlow Hub's image modules expect float inputs in the `[0, 1]` range. Use the `ImageDataGenerator`'s `rescale` parameter to achieve this.
The image size will be handled later.
```
image_generator = tf.keras.preprocessing.image.ImageDataGenerator(rescale=1/255)
image_data = image_generator.flow_from_directory(str(data_root), target_size=IMAGE_SHAPE)
```
The resulting object is an iterator that returns `image_batch, label_batch` pairs.
```
for image_batch, label_batch in image_data:
print("Image batch shape: ", image_batch.shape)
print("Label batch shape: ", label_batch.shape)
break
```
### Run the classifier on a batch of images
Now run the classifier on the image batch.
```
result_batch = classifier.predict(image_batch)
result_batch.shape
predicted_class_names = imagenet_labels[np.argmax(result_batch, axis=-1)]
predicted_class_names
```
Now check how these predictions line up with the images:
```
plt.figure(figsize=(10,9))
plt.subplots_adjust(hspace=0.5)
for n in range(30):
plt.subplot(6,5,n+1)
plt.imshow(image_batch[n])
plt.title(predicted_class_names[n])
plt.axis('off')
_ = plt.suptitle("ImageNet predictions")
```
See the `LICENSE.txt` file for image attributions.
The results are far from perfect, but reasonable considering that these are not the classes the model was trained for (except "daisy").
### Download the headless model
TensorFlow Hub also distributes models without the top classification layer. These can be used to easily do transfer learning.
Any [Tensorflow 2 compatible image feature vector URL](https://tfhub.dev/s?module-type=image-feature-vector&q=tf2) from tfhub.dev will work here.
```
feature_extractor_url = "https://tfhub.dev/google/tf2-preview/mobilenet_v2/feature_vector/2" #@param {type:"string"}
```
Create the feature extractor.
```
feature_extractor_layer = hub.KerasLayer(feature_extractor_url,
input_shape=(224,224,3))
```
It returns a 1280-length vector for each image:
```
feature_batch = feature_extractor_layer(image_batch)
print(feature_batch.shape)
```
Freeze the variables in the feature extractor layer, so that the training only modifies the new classifier layer.
```
feature_extractor_layer.trainable = False
```
### Attach a classification head
Now wrap the hub layer in a `tf.keras.Sequential` model, and add a new classification layer.
```
model = tf.keras.Sequential([
feature_extractor_layer,
layers.Dense(image_data.num_classes, activation='softmax')
])
model.summary()
predictions = model(image_batch)
predictions.shape
```
### Train the model
Use compile to configure the training process:
```
model.compile(
optimizer=tf.keras.optimizers.Adam(),
loss='categorical_crossentropy',
metrics=['acc'])
```
Now use the `.fit` method to train the model.
To keep this example short train just 2 epochs. To visualize the training progress, use a custom callback to log the loss and accuracy of each batch individually, instead of the epoch average.
```
class CollectBatchStats(tf.keras.callbacks.Callback):
def __init__(self):
self.batch_losses = []
self.batch_acc = []
def on_train_batch_end(self, batch, logs=None):
self.batch_losses.append(logs['loss'])
self.batch_acc.append(logs['acc'])
self.model.reset_metrics()
steps_per_epoch = np.ceil(image_data.samples/image_data.batch_size)
batch_stats_callback = CollectBatchStats()
history = model.fit_generator(image_data, epochs=2,
steps_per_epoch=steps_per_epoch,
callbacks = [batch_stats_callback])
```
Now after, even just a few training iterations, we can already see that the model is making progress on the task.
```
plt.figure()
plt.ylabel("Loss")
plt.xlabel("Training Steps")
plt.ylim([0,2])
plt.plot(batch_stats_callback.batch_losses)
plt.figure()
plt.ylabel("Accuracy")
plt.xlabel("Training Steps")
plt.ylim([0,1])
plt.plot(batch_stats_callback.batch_acc)
```
### Check the predictions
To redo the plot from before, first get the ordered list of class names:
```
class_names = sorted(image_data.class_indices.items(), key=lambda pair:pair[1])
class_names = np.array([key.title() for key, value in class_names])
class_names
```
Run the image batch through the model and convert the indices to class names.
```
predicted_batch = model.predict(image_batch)
predicted_id = np.argmax(predicted_batch, axis=-1)
predicted_label_batch = class_names[predicted_id]
```
Plot the result
```
label_id = np.argmax(label_batch, axis=-1)
plt.figure(figsize=(10,9))
plt.subplots_adjust(hspace=0.5)
for n in range(30):
plt.subplot(6,5,n+1)
plt.imshow(image_batch[n])
color = "green" if predicted_id[n] == label_id[n] else "red"
plt.title(predicted_label_batch[n].title(), color=color)
plt.axis('off')
_ = plt.suptitle("Model predictions (green: correct, red: incorrect)")
```
## Export your model
Now that you've trained the model, export it as a saved model:
```
import time
t = time.time()
export_path = "/tmp/saved_models/{}".format(int(t))
model.save(export_path, save_format='tf')
export_path
```
Now confirm that we can reload it, and it still gives the same results:
```
reloaded = tf.keras.models.load_model(export_path)
result_batch = model.predict(image_batch)
reloaded_result_batch = reloaded.predict(image_batch)
abs(reloaded_result_batch - result_batch).max()
```
This saved model can be loaded for inference later, or converted to [TFLite](https://www.tensorflow.org/lite/convert/) or [TFjs](https://github.com/tensorflow/tfjs-converter).
| github_jupyter |
```
Project: McNulty
Date: 02/22/2017
Name: Prashant Tatineni
```
# Project Overview
In this project, I attempt to predict the popularity (target variable: `interest_level`) of apartment rental listings based on listing characteristics. The data comes from a [Kaggle Competition](https://www.kaggle.com/c/two-sigma-connect-rental-listing-inquiries).
AWS and SQL were not used for joining to the dataset, as it was provided as a single file `train.json` (49,352 rows).
An additional file, `test.json` (74,659 rows) contains the same columns as `train.json`, except that the target variable, `interest_level`, is missing. Predictions of the target variable are to be made on the `test.json` file and submitted to Kaggle.
## Summary of Solution Steps
1. Load data from JSON
2. Build initial predictor variables, with `interest_level` as the target.
3. Initial run of classification models.
4. Add category indicators and aggregated features based on manager_id.
5. Run new Random Forest model.
6. Predict `interest_level` for the available test dataset.
```
# imports
import pandas as pd
import dateutil.parser
from sklearn import preprocessing
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
from sklearn.naive_bayes import BernoulliNB
from sklearn.naive_bayes import GaussianNB
from sklearn.naive_bayes import MultinomialNB
from sklearn.neural_network import MLPClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import accuracy_score
from sklearn.metrics import precision_recall_fscore_support
from sklearn.metrics import log_loss
import matplotlib.pyplot as plt
import seaborn as sns
from wordcloud import WordCloud
%matplotlib inline
```
## Step 1: Load Data
```
# Load the training dataset from Kaggle.
df = pd.read_json('data/raw/train.json')
print df.shape
df.head(2)
```
**Total number of columns is 14 + 1 target:**
- 1 target variable (**interest_level**), with classes **low, medium, high**
- 1 photo link
- lat/long, street address, display address
- listing_id, building_id, manager_id
- numerical (price, bathrooms, bedrooms)
- created date
- text (description, features)
**Features for modeling:**
- bathrooms
- bedrooms
- created date (calculate age of posting in days)
- description (number of words in description)
- features (number of features)
- photos (number of photos)
- price
- features (split into category indicators)
- manager_id (with manager skill level)
**Further opportunities for modeling:**
- description (need to text parse)
- building_id (possibly with a building popularity level)
- photos (quality)
```
# Distribution of target, interest_level
s = df.groupby('interest_level')['listing_id'].count()
s.plot.bar();
df_high = df.loc[df['interest_level'] == 'high']
df_medium = df.loc[df['interest_level'] == 'medium']
df_low = df.loc[df['interest_level'] == 'low']
plt.figure(figsize=(6,10))
plt.scatter(df_low.longitude, df_low.latitude, color='yellow', alpha=0.2, marker='.', label='Low')
plt.scatter(df_medium.longitude, df_medium.latitude, color='green', alpha=0.2, marker='.', label='Medium')
plt.scatter(df_high.longitude, df_high.latitude, color='purple', alpha=0.2, marker='.', label='High')
plt.xlim(-74.04,-73.80)
plt.ylim(40.6,40.9)
plt.legend(loc=2);
```
## Step 2: Initial Features
```
(pd.to_datetime(df['created'])).sort_values(ascending=False).head()
# The most recent records are 6/29/2016. Computing days old from 6/30/2016.
df['days_old'] = (dateutil.parser.parse('2016-06-30') - pd.to_datetime(df['created'])).apply(lambda x: x.days)
# Add other "count" features
df['num_words'] = df['description'].apply(lambda x: len(x.split()))
df['num_features'] = df['features'].apply(len)
df['num_photos'] = df['photos'].apply(len)
```
## Step 3: Modeling, First Pass
```
X = df[['bathrooms','bedrooms','price','latitude','longitude','days_old','num_words','num_features','num_photos']]
y = df['interest_level']
# Scaling is necessary for Logistic Regression and KNN
X_scaled = pd.DataFrame(preprocessing.scale(X))
X_scaled.columns = X.columns
X_train, X_test, y_train, y_test = train_test_split(X_scaled, y, random_state=42)
```
### Logistic Regression
```
lr = LogisticRegression()
lr.fit(X_train, y_train)
y_test_predicted_proba = lr.predict_proba(X_test)
log_loss(y_test, y_test_predicted_proba)
lr = LogisticRegression(solver='newton-cg', multi_class='multinomial')
lr.fit(X_train, y_train)
y_test_predicted_proba = lr.predict_proba(X_test)
log_loss(y_test, y_test_predicted_proba)
```
### KNN
```
for i in [95,100,105]:
knn = KNeighborsClassifier(n_neighbors=i)
knn.fit(X_train, y_train)
y_test_predicted_proba = knn.predict_proba(X_test)
print log_loss(y_test, y_test_predicted_proba)
```
### Random Forest
```
rf = RandomForestClassifier(n_estimators=1000, n_jobs=-1)
rf.fit(X_train, y_train)
y_test_predicted_proba = rf.predict_proba(X_test)
log_loss(y_test, y_test_predicted_proba)
```
### Naive Bayes
```
bnb = BernoulliNB()
bnb.fit(X_train, y_train)
y_test_predicted_proba = bnb.predict_proba(X_test)
log_loss(y_test, y_test_predicted_proba)
gnb = GaussianNB()
gnb.fit(X_train, y_train)
y_test_predicted_proba = gnb.predict_proba(X_test)
log_loss(y_test, y_test_predicted_proba)
```
### Neural Network
```
clf = MLPClassifier(hidden_layer_sizes=(100,50,10))
clf.fit(X_train, y_train)
y_test_predicted_proba = clf.predict_proba(X_test)
log_loss(y_test, y_test_predicted_proba)
```
## Step 4: More Features
### Splitting out categories into 0/1 dummy variables
```
# Reduce 1556 unique category text values into 34 main categories
def reduce_categories(full_list):
reduced_list = []
for i in full_list:
item = i.lower()
if 'cats allowed' in item:
reduced_list.append('cats')
if 'dogs allowed' in item:
reduced_list.append('dogs')
if 'elevator' in item:
reduced_list.append('elevator')
if 'hardwood' in item:
reduced_list.append('elevator')
if 'doorman' in item or 'concierge' in item:
reduced_list.append('doorman')
if 'dishwasher' in item:
reduced_list.append('dishwasher')
if 'laundry' in item or 'dryer' in item:
if 'unit' in item:
reduced_list.append('laundry_in_unit')
else:
reduced_list.append('laundry')
if 'no fee' in item:
reduced_list.append('no_fee')
if 'reduced fee' in item:
reduced_list.append('reduced_fee')
if 'fitness' in item or 'gym' in item:
reduced_list.append('gym')
if 'prewar' in item or 'pre-war' in item:
reduced_list.append('prewar')
if 'dining room' in item:
reduced_list.append('dining')
if 'pool' in item:
reduced_list.append('pool')
if 'internet' in item:
reduced_list.append('internet')
if 'new construction' in item:
reduced_list.append('new_construction')
if 'wheelchair' in item:
reduced_list.append('wheelchair')
if 'exclusive' in item:
reduced_list.append('exclusive')
if 'loft' in item:
reduced_list.append('loft')
if 'simplex' in item:
reduced_list.append('simplex')
if 'fire' in item:
reduced_list.append('fireplace')
if 'lowrise' in item or 'low-rise' in item:
reduced_list.append('lowrise')
if 'midrise' in item or 'mid-rise' in item:
reduced_list.append('midrise')
if 'highrise' in item or 'high-rise' in item:
reduced_list.append('highrise')
if 'pool' in item:
reduced_list.append('pool')
if 'ceiling' in item:
reduced_list.append('high_ceiling')
if 'garage' in item or 'parking' in item:
reduced_list.append('parking')
if 'furnished' in item:
reduced_list.append('furnished')
if 'multi-level' in item:
reduced_list.append('multilevel')
if 'renovated' in item:
reduced_list.append('renovated')
if 'super' in item:
reduced_list.append('live_in_super')
if 'green building' in item:
reduced_list.append('green_building')
if 'appliances' in item:
reduced_list.append('new_appliances')
if 'luxury' in item:
reduced_list.append('luxury')
if 'penthouse' in item:
reduced_list.append('penthouse')
if 'deck' in item or 'terrace' in item or 'balcony' in item or 'outdoor' in item or 'roof' in item or 'garden' in item or 'patio' in item:
reduced_list.append('outdoor_space')
return list(set(reduced_list))
df['categories'] = df['features'].apply(reduce_categories)
text = ''
for index, row in df.iterrows():
for i in row.categories:
text = text + i + ' '
plt.figure(figsize=(12,6))
wc = WordCloud(background_color='white', width=1200, height=600).generate(text)
plt.title('Reduced Categories', fontsize=30)
plt.axis("off")
wc.recolor(random_state=0)
plt.imshow(wc);
# Create indicators
X_dummies = pd.get_dummies(df['categories'].apply(pd.Series).stack()).sum(level=0)
```
### Aggregate manager_id to get features representing manager performance
Note: Need to aggregate manager performance ONLY over a training subset in order to validate against test subset. So the train-test split is being performed in this step before creating the columns for manager performance.
```
# Choose features for modeling (and sorting)
df = df.sort_values('listing_id')
X = df[['bathrooms','bedrooms','price','latitude','longitude','days_old','num_words','num_features','num_photos','listing_id','manager_id']]
y = df['interest_level']
# Merge indicators to X dataframe and sort again to match sorting of y
X = X.merge(X_dummies, how='outer', left_index=True, right_index=True).fillna(0)
X = X.sort_values('listing_id')
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
# compute ratios and count for each manager
mgr_perf = pd.concat([X_train.manager_id,pd.get_dummies(y_train)], axis=1).groupby('manager_id').mean()
mgr_perf.head(2)
mgr_perf['manager_count'] = X_train.groupby('manager_id').count().iloc[:,1]
mgr_perf['manager_skill'] = mgr_perf['high']*1 + mgr_perf['medium']*0 + mgr_perf['low']*-1
# for training set
X_train = X_train.merge(mgr_perf.reset_index(), how='left', left_on='manager_id', right_on='manager_id')
# for test set
X_test = X_test.merge(mgr_perf.reset_index(), how='left', left_on='manager_id', right_on='manager_id')
# Fill na's with mean skill and median count
X_test['manager_skill'] = X_test.manager_skill.fillna(X_test.manager_skill.mean())
X_test['manager_count'] = X_test.manager_count.fillna(X_test.manager_count.median())
# Delete unnecessary columns before modeling
del X_train['listing_id']
del X_train['manager_id']
del X_test['listing_id']
del X_test['manager_id']
del X_train['high']
del X_train['medium']
del X_train['low']
del X_test['high']
del X_test['medium']
del X_test['low']
```
## Step 5: Modeling, second pass with Random Forest
```
rf = RandomForestClassifier(n_estimators=1000, n_jobs=-1)
rf.fit(X_train, y_train)
y_test_predicted_proba = rf.predict_proba(X_test)
log_loss(y_test, y_test_predicted_proba)
y_test_predicted = rf.predict(X_test)
accuracy_score(y_test, y_test_predicted)
precision_recall_fscore_support(y_test, y_test_predicted)
rf.classes_
plt.figure(figsize=(15,5))
pd.Series(index = X_train.columns, data = rf.feature_importances_).sort_values().plot(kind = 'bar');
```
As seen here, introducing feature categories and manager performance has improved the model. In particular, `manager_skill` shows up as the dominant feature in terms of importance in this random forest model.
## Step 6: Prediction and Next opportunities
To make a prediction for submission to Kaggle, this notebook is recreated with the `test.json` dataset. The submission requires the predicted high, medium, and low probabilities for each `listing_id`.
Further opportunities to improve prediction on this dataset lie in the text descriptions and image data, which were not used thus far except as a numerical "length" value. Building popularity could also be assessed via the `building_id` variable, similar to the aggregation of the `manager_id` variable.
It will also be valuable to spend time optimizing the model used here (perhaps using GridSearch).
| github_jupyter |
# Double Exponential Weighted Moving Average
https://digscholarship.unco.edu/cgi/viewcontent.cgi?article=1428&context=dissertations
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings("ignore")
# fix_yahoo_finance is used to fetch data
import fix_yahoo_finance as yf
yf.pdr_override()
# input
symbol = 'AAPL'
start = '2018-01-01'
end = '2019-01-01'
# Read data
df = yf.download(symbol,start,end)
# View Columns
df.head()
n = 7
ewma = df['Adj Close'].ewm(ignore_na=False, min_periods=n - 1,
span=n).mean()
ewma_mean = ewma.ewm(ignore_na=False, min_periods=n- 1,
span=n).mean()
df['DEWMA'] = 2 * ewma - ewma_mean
df.head(20)
fig = plt.figure(figsize=(14,10))
ax1 = plt.subplot(2, 1, 1)
ax1.plot(df['Adj Close'])
ax1.set_title('Stock '+ symbol +' Closing Price')
ax1.set_ylabel('Price')
ax2 = plt.subplot(2, 1, 2)
ax2.plot(df['DEWMA'], label='Double Exponential Weighted Moving Average', color='red')
#ax2.axhline(y=0, color='blue', linestyle='--')
#ax2.axhline(y=0.5, color='darkblue')
#ax2.axhline(y=-0.5, color='darkblue')
ax2.grid()
ax2.set_ylabel('Double Exponential Weighted Moving Average')
ax2.set_xlabel('Date')
ax2.legend(loc='best')
```
## Candlestick with Double Exponential Weighted Moving Average
```
from matplotlib import dates as mdates
import datetime as dt
dfc = df.copy()
dfc['VolumePositive'] = dfc['Open'] < dfc['Adj Close']
#dfc = dfc.dropna()
dfc = dfc.reset_index()
dfc['Date'] = pd.to_datetime(dfc['Date'])
dfc['Date'] = dfc['Date'].apply(mdates.date2num)
dfc.head()
from mpl_finance import candlestick_ohlc
fig = plt.figure(figsize=(14,10))
ax1 = plt.subplot(2, 1, 1)
candlestick_ohlc(ax1,dfc.values, width=0.5, colorup='g', colordown='r', alpha=1.0)
ax1.xaxis_date()
ax1.xaxis.set_major_formatter(mdates.DateFormatter('%d-%m-%Y'))
ax1.grid(True, which='both')
ax1.minorticks_on()
ax1v = ax1.twinx()
colors = dfc.VolumePositive.map({True: 'g', False: 'r'})
ax1v.bar(dfc.Date, dfc['Volume'], color=colors, alpha=0.4)
ax1v.axes.yaxis.set_ticklabels([])
ax1v.set_ylim(0, 3*df.Volume.max())
ax1.set_title('Stock '+ symbol +' Closing Price')
ax1.set_ylabel('Price')
ax2 = plt.subplot(2, 1, 2)
ax2.plot(df['DEWMA'], label='Double Exponential Weighted Moving Average', color='red')
#ax2.axhline(y=0, color='blue', linestyle='--')
#ax2.axhline(y=0.5, color='darkblue')
#ax2.axhline(y=-0.5, color='darkblue')
ax2.grid()
ax2.set_ylabel('Double Exponential Weighted Moving Average')
ax2.set_xlabel('Date')
ax2.legend(loc='best')
```
| github_jupyter |
# Spatial Joins
A *spatial join* uses [binary predicates](http://toblerity.org/shapely/manual.html#binary-predicates)
such as `intersects` and `crosses` to combine two `GeoDataFrames` based on the spatial relationship
between their geometries.
A common use case might be a spatial join between a point layer and a polygon layer where you want to retain the point geometries and grab the attributes of the intersecting polygons.
```
from IPython.core.display import Image
Image(url='https://web.natur.cuni.cz/~langhamr/lectures/vtfg1/mapinfo_1/about_gis/Image23.gif')
```
## Types of spatial joins
We currently support the following methods of spatial joins. We refer to the *left_df* and *right_df* which are the correspond to the two dataframes passed in as args.
### Left outer join
In a LEFT OUTER JOIN (`how='left'`), we keep *all* rows from the left and duplicate them if necessary to represent multiple hits between the two dataframes. We retain attributes of the right if they intersect and lose right rows that don't intersect. A left outer join implies that we are interested in retaining the geometries of the left.
This is equivalent to the PostGIS query:
```
SELECT pts.geom, pts.id as ptid, polys.id as polyid
FROM pts
LEFT OUTER JOIN polys
ON ST_Intersects(pts.geom, polys.geom);
geom | ptid | polyid
--------------------------------------------+------+--------
010100000040A9FBF2D88AD03F349CD47D796CE9BF | 4 | 10
010100000048EABE3CB622D8BFA8FBF2D88AA0E9BF | 3 | 10
010100000048EABE3CB622D8BFA8FBF2D88AA0E9BF | 3 | 20
0101000000F0D88AA0E1A4EEBF7052F7E5B115E9BF | 2 | 20
0101000000818693BA2F8FF7BF4ADD97C75604E9BF | 1 |
(5 rows)
```
### Right outer join
In a RIGHT OUTER JOIN (`how='right'`), we keep *all* rows from the right and duplicate them if necessary to represent multiple hits between the two dataframes. We retain attributes of the left if they intersect and lose left rows that don't intersect. A right outer join implies that we are interested in retaining the geometries of the right.
This is equivalent to the PostGIS query:
```
SELECT polys.geom, pts.id as ptid, polys.id as polyid
FROM pts
RIGHT OUTER JOIN polys
ON ST_Intersects(pts.geom, polys.geom);
geom | ptid | polyid
----------+------+--------
01...9BF | 4 | 10
01...9BF | 3 | 10
02...7BF | 3 | 20
02...7BF | 2 | 20
00...5BF | | 30
(5 rows)
```
### Inner join
In an INNER JOIN (`how='inner'`), we keep rows from the right and left only where their binary predicate is `True`. We duplicate them if necessary to represent multiple hits between the two dataframes. We retain attributes of the right and left only if they intersect and lose all rows that do not. An inner join implies that we are interested in retaining the geometries of the left.
This is equivalent to the PostGIS query:
```
SELECT pts.geom, pts.id as ptid, polys.id as polyid
FROM pts
INNER JOIN polys
ON ST_Intersects(pts.geom, polys.geom);
geom | ptid | polyid
--------------------------------------------+------+--------
010100000040A9FBF2D88AD03F349CD47D796CE9BF | 4 | 10
010100000048EABE3CB622D8BFA8FBF2D88AA0E9BF | 3 | 10
010100000048EABE3CB622D8BFA8FBF2D88AA0E9BF | 3 | 20
0101000000F0D88AA0E1A4EEBF7052F7E5B115E9BF | 2 | 20
(4 rows)
```
## Spatial Joins between two GeoDataFrames
Let's take a look at how we'd implement these using `GeoPandas`. First, load up the NYC test data into `GeoDataFrames`:
```
%matplotlib inline
from shapely.geometry import Point
from geopandas import datasets, GeoDataFrame, read_file
from geopandas.tools import overlay
# NYC Boros
zippath = datasets.get_path('nybb')
polydf = read_file(zippath)
# Generate some points
b = [int(x) for x in polydf.total_bounds]
N = 8
pointdf = GeoDataFrame([
{'geometry': Point(x, y), 'value1': x + y, 'value2': x - y}
for x, y in zip(range(b[0], b[2], int((b[2] - b[0]) / N)),
range(b[1], b[3], int((b[3] - b[1]) / N)))])
# Make sure they're using the same projection reference
pointdf.crs = polydf.crs
pointdf
polydf
pointdf.plot()
polydf.plot()
```
## Joins
```
from geopandas.tools import sjoin
join_left_df = sjoin(pointdf, polydf, how="left")
join_left_df
# Note the NaNs where the point did not intersect a boro
join_right_df = sjoin(pointdf, polydf, how="right")
join_right_df
# Note Staten Island is repeated
join_inner_df = sjoin(pointdf, polydf, how="inner")
join_inner_df
# Note the lack of NaNs; dropped anything that didn't intersect
```
We're not limited to using the `intersection` binary predicate. Any of the `Shapely` geometry methods that return a Boolean can be used by specifying the `op` kwarg.
```
sjoin(pointdf, polydf, how="left", op="within")
```
| github_jupyter |
# Monte Carlo methods
- MC integration: estimate the value of `pi`
- MC simulation: simulate the exponential law
```
import numpy as np
import matplotlib.pyplot as plt
N = 10000
M = 0
X, Y = [], []
Z = []
E = []
V = []
np.random.seed(100)
for i in range(1, N+1):
x, y = np.random.random(2)
if pow(x, 2) + pow(y, 2) <= 1.0:
M+=1
label = 1
else:
label = 0
estimate = M/i * 4
error = (estimate - np.pi) / np.pi
variance = i / (i - 1) * (1 / i* 4 * M - estimate) if i != 1 else 0
X.append((x, y))
Y.append(label)
Z.append(estimate)
V.append(variance)
E.append(error)
if i % 1000 == 0:
print("Estimation at {:08d}: {:8f} variance: {}".format(i, estimate, variance))
print("Final estimate at {}: {}".format(N, M/N * 4))
X = np.asarray(X)
Y = np.asarray(Y)
Z = np.asarray(Z)
E = np.asarray(E)
plt.scatter(X[:, 0], X[:, 1], c=Y)
plt.show()
```
The variance of MC estimate is:

And the error:

decreases as 1/sqrt(N)
```
plt.plot(np.arange(N), E)
plt.plot(np.arange(N)[1:], 1/np.sqrt(np.arange(N)[1:]))
```
### Variance Reduction: Importance Sampling
One technique to reduce the variance of the estimate is _Importance Sampling_

where:

p(x) is the importance sampling distribution.
## Simulation of laws
Proposition 4.1 [1] gives us a way to simulate distributions.
Let's do it for an _exponential law_:
```
# exponential law
import numpy as np
lambda_ = 1.5
exp_law_monte_carlo = lambda u: - np.log(u)/lambda_
N = 5000
Y = []
for i in range(N):
y = exp_law_monte_carlo(np.random.random())
Y.append(y)
Y = np.asarray(Y)
plt.hist(Y, 100, density=True, alpha=0.75)
plt.hist(np.random.exponential(1/lambda_, N), 100, density=True, alpha=0.75)
plt.show()
```
## References
- [1] [http://cermics.enpc.fr/~bl/Halmstad/monte-carlo/lecture-1.pdf](http://cermics.enpc.fr/~bl/Halmstad/monte-carlo/lecture-1.pdf)
- [2] [Monte Carlo and quasi-Monte Carlo methods](https://pdfs.semanticscholar.org/85b5/97939f26a4f7a149887e00e83e2f5ba35c8f.pdf)
- [3] [https://en.wikipedia.org/wiki/Monte_Carlo_integration](https://en.wikipedia.org/wiki/Monte_Carlo_integration)
- [4] [Monte Carlo Methods by Kroese (notes)](https://people.smp.uq.edu.au/DirkKroese/mccourse.pdf)
| github_jupyter |
```
import dask
import dask.dataframe as dd
import warnings
import datetime
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import category_encoders as ce
from catboost import Pool
from catboost import CatBoostClassifier
from sklearn.model_selection import train_test_split, StratifiedKFold, GridSearchCV
from sklearn.metrics import confusion_matrix, classification_report, accuracy_score, recall_score, precision_score, f1_score
%matplotlib inline
sns.set(style="whitegrid")
warnings.filterwarnings("ignore")
pd.set_option('display.float_format', lambda x: '%.2f' % x)
dtypes = {
'MachineIdentifier': 'category',
'AVProductsInstalled': 'float16',
'AVProductsEnabled': 'float16',
'IsProtected': 'float16',
'Census_ProcessorCoreCount': 'float16',
'Census_SystemVolumeTotalCapacity': 'float32',
'Census_IsAlwaysOnAlwaysConnectedCapable': 'float16',
'Wdft_IsGamer': 'float16',
'AvSigVersion': 'category',
'OsBuildLab': 'category',
'Census_OSVersion': 'category',
'AppVersion': 'category',
'EngineVersion': 'category',
'Census_PowerPlatformRoleName': 'category',
'OsPlatformSubRelease': 'category',
'Census_OSInstallTypeName': 'category',
'SkuEdition': 'category',
'Census_ActivationChannel': 'category',
'Census_OSWUAutoUpdateOptionsName': 'category',
'ProductName': 'category',
'Platform': 'category',
'Census_PrimaryDiskTypeName': 'category',
'Census_DeviceFamily': 'category',
'Census_OSArchitecture': 'category',
'Processor': 'category',
'HasDetections': 'int8'
}
label = ['HasDetections']
ids = ['MachineIdentifier']
numerical_features = ['AVProductsEnabled', 'AVProductsInstalled',
'Census_ProcessorCoreCount', 'Census_SystemVolumeTotalCapacity']
binary_features = ['Census_IsAlwaysOnAlwaysConnectedCapable', 'IsProtected', 'Wdft_IsGamer']
version_features = ['AvSigVersion', 'OsBuildLab', 'Census_OSVersion', 'AppVersion', 'EngineVersion']
# < 10 categories
low_cardinality_features = ['Census_PowerPlatformRoleName', 'OsPlatformSubRelease',
'Census_OSInstallTypeName', 'SkuEdition', 'Census_ActivationChannel',
'Census_OSWUAutoUpdateOptionsName', 'ProductName',
'Platform', 'Census_PrimaryDiskTypeName', 'Census_DeviceFamily',
'Census_OSArchitecture', 'Processor']
use_columns = numerical_features + binary_features + version_features + low_cardinality_features
```
### Load data
```
train = pd.read_csv('../input/microsoft-malware-prediction/train.csv', dtype=dtypes, usecols=(use_columns + label))
# IMPORT TIMESTAMP DICTIONARY
datedict = np.load('../input/malware-timestamps/AvSigVersionTimestamps.npy')
datedict = datedict[()]
# ADD TIMESTAMPS
train['Date'] = train['AvSigVersion'].map(datedict)
print(train.shape)
```
### Reduce granularity on version features
```
for feature in version_features:
if feature in ['EngineVersion']:
train[feature] = train[feature].apply(lambda x : ".".join(x.split('.')[:3]))
elif feature in ['OsBuildLab']:
train[feature] = train[feature].apply(lambda x : ".".join(x.split('.')[:1]))
else:
train[feature] = train[feature].apply(lambda x : ".".join(x.split('.')[:2]))
```
### Raw dataset overview
```
print('Dataset number of records: %s' % train.shape[0])
print('Dataset number of columns: %s' % train.shape[1])
train.head()
```
### Pre process
```
# Remove rows with NA
train.dropna(inplace=True)
display(train[numerical_features].describe().T)
display(train[binary_features].describe().T)
display(train[version_features].describe().T)
```
#### Train/validation random split (80% train / 20% validation)
* Use last month as validation
```
print('Train set min date', train['Date'].min().date())
print('Train set max date', train['Date'].max().date())
X_train_raw = train[train['Date']<=datetime.date(2018, 9, 15)]
X_val_raw = train[train['Date']>datetime.date(2018, 9, 15)]
X_train, X_val = train_test_split(X_train_raw, test_size=0.1,random_state=1)
X_val = X_val.append(X_val_raw)
# Get labels
Y_train = X_train['HasDetections'].values
Y_val = X_val['HasDetections'].values
X_train.drop(['HasDetections'], axis=1, inplace=True)
X_val.drop(['HasDetections'], axis=1, inplace=True)
del train, X_train_raw, X_val_raw, datedict
```
### Mean encoder
```
encoder = ce.TargetEncoder(cols=(version_features + low_cardinality_features))
encoder.fit(X_train, Y_train)
X_train = encoder.fit_transform(X_train.reset_index(), Y_train)
X_val = encoder.transform(X_val.reset_index())
X_train.drop('index', axis=1, inplace=True)
X_val.drop('index', axis=1, inplace=True)
```
### Fill missing values with mean
```
X_train.fillna(X_train.mean(), inplace=True)
X_val.fillna(X_val.mean(), inplace=True)
print(X_train.shape)
print(X_val.shape)
print(X_train['Date'].max().date())
print(X_val['Date'].max().date())
X_train.drop(['Date'], axis=1, inplace=True)
X_val.drop(['Date'], axis=1, inplace=True)
X = X_train.append(X_val).values
Y = np.append(Y_train, Y_val)
del X_train, X_val, Y_train, Y_val
```
### Hyperparameter Tuning
```
# K-Fold cross validation
folds = 3
kfold = StratifiedKFold(n_splits=folds, shuffle=True, random_state=0)
```
#### Load test set
```
test = dd.read_csv('../input/microsoft-malware-prediction/test.csv', dtype=dtypes, usecols=(use_columns + ids)).compute()
```
#### Pre process test
```
for feature in version_features:
if feature in ['EngineVersion']:
test[feature] = test[feature].apply(lambda x : ".".join(x.split('.')[:3]))
elif feature in ['OsBuildLab']:
test[feature] = test[feature].apply(lambda x : ".".join(x.split('.')[:1]))
else:
test[feature] = test[feature].apply(lambda x : ".".join(x.split('.')[:2]))
test = encoder.transform(test.reset_index())
test.drop('index', axis=1, inplace=True)
submission = pd.DataFrame({"MachineIdentifier":test['MachineIdentifier']})
submission['HasDetections'] = 0
test.drop('MachineIdentifier',axis=1, inplace=True)
test.fillna(test.mean(), inplace=True)
```
#### Model training
```
counter = 0
for train_index, val_index in kfold.split(X, Y):
counter += 1
print('Fold {}\n'.format(counter))
X_train, X_val = X[train_index], X[val_index]
Y_train, Y_val = Y[train_index], Y[val_index]
max_depth = 8
n_estimators = 300
colsample_bylevel = 0.7
model = CatBoostClassifier(
od_type='Iter',
od_wait=20,
verbose=50,
loss_function='Logloss',
max_depth=max_depth,
n_estimators=n_estimators,
colsample_bylevel=colsample_bylevel
)
model.fit(
X_train, Y_train,
eval_set=(X_val, Y_val)
)
train_predictions = model.predict(X_train)
val_predictions = model.predict(X_val)
train_acc = accuracy_score(Y_train, train_predictions) * 100
val_acc = accuracy_score(Y_val, val_predictions) * 100
train_recall = recall_score(Y_train, train_predictions) * 100
val_recall = recall_score(Y_val, val_predictions) * 100
train_precision = precision_score(Y_train, train_predictions) * 100
val_precision = precision_score(Y_val, val_predictions) * 100
train_f1 = f1_score(Y_train, train_predictions) * 100
val_f1 = f1_score(Y_val, val_predictions) * 100
print('-----Train-----')
print('Acc: %.2f Precision: %.2f Recall: %.2f F1: %.2f' % (train_acc, train_precision, train_recall, train_f1))
print('-----Validation-----')
print('Acc: %.2f Precision: %.2f Recall: %.2f F1: %.2f \n' % (val_acc, val_precision, val_recall, val_f1))
# Make predictions
predictions = model.predict_proba(test)
predictions = [x[1] for x in predictions]
submission['HasDetections'] += predictions
submission['HasDetections'] /=folds
```
#### Model feature importance
```
# feature_score = pd.DataFrame(list(zip(X_train.dtypes.index, model.get_feature_importance(Pool(X_train, label=Y_train, cat_features=[])))), columns=['Feature','Score'])
# feature_score = feature_score.sort_values(by='Score', ascending=False, inplace=False, kind='quicksort', na_position='last')
# plt.rcParams["figure.figsize"] = (16, 10)
# ax = feature_score.plot('Feature', 'Score', kind='bar', color='c')
# ax.set_title("Catboost Feature Importance Ranking", fontsize = 14)
# ax.set_xlabel('')
# rects = ax.patches
# labels = feature_score['Score'].round(2)
# for rect, label in zip(rects, labels):
# height = rect.get_height()
# ax.text(rect.get_x() + rect.get_width()/2, height + 0.35, label, ha='center', va='bottom')
# plt.show()
target_names=['HasDetections = 0', 'HasDetections = 1']
print('-----Train-----')
print(classification_report(Y_train, train_predictions, target_names=target_names))
print('-----Validation-----')
print(classification_report(Y_val, val_predictions, target_names=target_names))
```
#### Confusion matrix
```
f, axes = plt.subplots(1, 2, figsize=(16, 5), sharex=True)
train_cnf_matrix = confusion_matrix(Y_train, train_predictions)
val_cnf_matrix = confusion_matrix(Y_val, val_predictions)
train_cnf_matrix_norm = train_cnf_matrix / train_cnf_matrix.sum(axis=1)[:, np.newaxis]
val_cnf_matrix_norm = val_cnf_matrix / val_cnf_matrix.sum(axis=1)[:, np.newaxis]
train_df_cm = pd.DataFrame(train_cnf_matrix_norm, index=[0, 1], columns=[0, 1])
val_df_cm = pd.DataFrame(val_cnf_matrix_norm, index=[0, 1], columns=[0, 1])
sns.heatmap(train_df_cm, annot=True, fmt='.2f', cmap="Blues", ax=axes[0]).set_title("Train")
sns.heatmap(val_df_cm, annot=True, fmt='.2f', cmap="Blues", ax=axes[1]).set_title("Validation")
plt.show()
```
#### Output results
```
submission.to_csv("submission.csv", index=False)
submission.head(10)
```
Blend predictions
```
a = pd.read_csv('../input/lightgbm-baseline-model-using-sparse-matrix/lgb_submission.csv') # 3
b = pd.read_csv('../input/xdeepfm-baseline/nffm_submission.csv') # 4
c = pd.read_csv('../input/nffm-baseline-0-690-on-lb/nffm_submission.csv') # 5
d = pd.read_csv('../input/ms-malware-starter/ms_malware.csv') # 8
e = pd.read_csv('../input/light-gbm-on-stratified-k-folds-malwares/lgb_submission.csv') # ??
predictions = submission['HasDetections'].values
submission = a[['MachineIdentifier']]
submission['HasDetections'] = (a['HasDetections']+b['HasDetections']+c['HasDetections']+d['HasDetections']+e['HasDetections']+predictions)/6
submission.to_csv('submission_ensembled.csv', index=False)
submission.head(10)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/Tessellate-Imaging/monk_v1/blob/master/study_roadmaps/1_getting_started_roadmap/2_elemental_features_of_monk/2)%20Feature%20-%20Resume%20an%20interrupted%20training.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Goals
### 1. Understand how you can resume an interrupted training from last check point
- Training can get interrupted for variety of reasons
- power failure
- connection lost to power remote desktop
### 2. Training is also somwtimes intentionally interrupted to update hyper-parameters and resume training
### 3. Steps
- Start training a classifier for 20 epochs
- Interrupt training manually at around 10 epochs
- Change batch size
- Resume training
## Table of Contents
## [0. Install](#0)
## [1. Start training a classifier for 20 epochs](#1)
## [2. Reload experiment in resume mode](#2)
## [3. Resume training the classifier](#3)
<a id='0'></a>
# Install Monk
- git clone https://github.com/Tessellate-Imaging/monk_v1.git
- cd monk_v1/installation/Linux && pip install -r requirements_cu9.txt
- (Select the requirements file as per OS and CUDA version)
```
!git clone https://github.com/Tessellate-Imaging/monk_v1.git
# Select the requirements file as per OS and CUDA version
!cd monk_v1/installation/Linux && pip install -r requirements_cu9.txt
```
## Dataset - Malarial cell images
- Credits: https://www.kaggle.com/iarunava/cell-images-for-detecting-malaria
```
! wget --load-cookies /tmp/cookies.txt "https://docs.google.com/uc?export=download&confirm=$(wget --save-cookies /tmp/cookies.txt --keep-session-cookies --no-check-certificate 'https://docs.google.com/uc?export=download&id=1mMEtGIK8UZNCrErXRJR-kutNTaN1zxjC' -O- | sed -rn 's/.*confirm=([0-9A-Za-z_]+).*/\1\n/p')&id=1mMEtGIK8UZNCrErXRJR-kutNTaN1zxjC" -O malaria_cell.zip && rm -rf /tmp/cookies.txt
! unzip -qq malaria_cell.zip
```
# Imports
- Using single mxnet-gluoncv backend for this tutorial
```
# Monk
import os
import sys
sys.path.append("monk_v1/monk/");
#Using mxnet-gluon backend
from gluon_prototype import prototype
```
<a id='1'></a>
# Start training a classifier for 20 epochs
### Creating and managing experiments
- Provide project name
- Provide experiment name
```
gtf = prototype(verbose=1);
gtf.Prototype("Malaria-Cell", "exp-resume-training");
```
### This creates files and directories as per the following structure
workspace
|
|--------Malaria-Cell
|
|
|-----exp-resume-training
|
|-----experiment-state.json
|
|-----output
|
|------logs (All training logs and graphs saved here)
|
|------models (all trained models saved here)
### Load Dataset
```
gtf.Default(dataset_path="malaria_cell",
model_name="resnet18_v2",
num_epochs=20);
#Read the summary generated once you run this cell.
```
### From the summary current batch size is 4
```
#Start Training
gtf.Train();
#Manually stop the cell after 10 iteration
```
<a id='2'></a>
# Reload experiment in resume mode
- Set resume_train flag as True
```
gtf = prototype(verbose=1);
gtf.Prototype("Malaria-Cell", "exp-resume-training", resume_train=True);
```
### Print Summary
```
gtf.Summary()
```
### Summary findings
- Training status is still False
### Update batch size
```
# This part of code will be taken up again in upcoming sections
gtf.update_batch_size(8);
gtf.Reload();
```
<a id='3'></a>
# Resume training the classifier
```
gtf.Train();
```
### It skipped epochs 1-9
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/Array/array_transformations.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/Array/array_transformations.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/Array/array_transformations.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API and geemap
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geemap](https://github.com/giswqs/geemap). The **geemap** Python package is built upon the [ipyleaflet](https://github.com/jupyter-widgets/ipyleaflet) and [folium](https://github.com/python-visualization/folium) packages and implements several methods for interacting with Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, and `Map.centerObject()`.
The following script checks if the geemap package has been installed. If not, it will install geemap, which automatically installs its [dependencies](https://github.com/giswqs/geemap#dependencies), including earthengine-api, folium, and ipyleaflet.
**Important note**: A key difference between folium and ipyleaflet is that ipyleaflet is built upon ipywidgets and allows bidirectional communication between the front-end and the backend enabling the use of the map to capture user input, while folium is meant for displaying static data only ([source](https://blog.jupyter.org/interactive-gis-in-jupyter-with-ipyleaflet-52f9657fa7a)). Note that [Google Colab](https://colab.research.google.com/) currently does not support ipyleaflet ([source](https://github.com/googlecolab/colabtools/issues/60#issuecomment-596225619)). Therefore, if you are using geemap with Google Colab, you should use [`import geemap.eefolium`](https://github.com/giswqs/geemap/blob/master/geemap/eefolium.py). If you are using geemap with [binder](https://mybinder.org/) or a local Jupyter notebook server, you can use [`import geemap`](https://github.com/giswqs/geemap/blob/master/geemap/geemap.py), which provides more functionalities for capturing user input (e.g., mouse-clicking and moving).
```
# Installs geemap package
import subprocess
try:
import geemap
except ImportError:
print('geemap package not installed. Installing ...')
subprocess.check_call(["python", '-m', 'pip', 'install', 'geemap'])
# Checks whether this notebook is running on Google Colab
try:
import google.colab
import geemap.eefolium as emap
except:
import geemap as emap
# Authenticates and initializes Earth Engine
import ee
try:
ee.Initialize()
except Exception as e:
ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
The default basemap is `Google Satellite`. [Additional basemaps](https://github.com/giswqs/geemap/blob/master/geemap/geemap.py#L13) can be added using the `Map.add_basemap()` function.
```
Map = emap.Map(center=[40,-100], zoom=4)
Map.add_basemap('ROADMAP') # Add Google Map
Map
```
## Add Earth Engine Python script
```
# Add Earth Engine dataset
import math
# This function masks the input with a threshold on the simple cloud score.
def cloudMask(img):
cloudscore = ee.Algorithms.Landsat.simpleCloudScore(img).select('cloud')
return img.updateMask(cloudscore.lt(50))
# cloudMask = function(img) {
# cloudscore = ee.Algorithms.Landsat.simpleCloudScore(img).select('cloud')
# return img.updateMask(cloudscore.lt(50))
# }
# This function computes the predictors and the response from the input.
def makeVariables(image):
# Compute time of the image in fractional years relative to the Epoch.
year = ee.Image(image.date().difference(ee.Date('1970-01-01'), 'year'))
# Compute the season in radians, one cycle per year.
season = year.multiply(2 * math.pi)
# Return an image of the predictors followed by the response.
return image.select() \
.addBands(ee.Image(1)) \
.addBands(year.rename('t')) \
.addBands(season.sin().rename('sin')) \
.addBands(season.cos().rename('cos')) \
.addBands(image.normalizedDifference().rename('NDVI')) \
.toFloat()
# Load a Landsat 5 image collection.
collection = ee.ImageCollection('LANDSAT/LT05/C01/T1_TOA') \
.filterDate('2008-04-01', '2010-04-01') \
.filterBounds(ee.Geometry.Point(-122.2627, 37.8735)) \
.map(cloudMask) \
.select(['B4', 'B3']) \
.sort('system:time_start', True)
# # This function computes the predictors and the response from the input.
# makeVariables = function(image) {
# # Compute time of the image in fractional years relative to the Epoch.
# year = ee.Image(image.date().difference(ee.Date('1970-01-01'), 'year'))
# # Compute the season in radians, one cycle per year.
# season = year.multiply(2 * Math.PI)
# # Return an image of the predictors followed by the response.
# return image.select() \
# .addBands(ee.Image(1)) # 0. constant \
# .addBands(year.rename('t')) # 1. linear trend \
# .addBands(season.sin().rename('sin')) # 2. seasonal \
# .addBands(season.cos().rename('cos')) # 3. seasonal \
# .addBands(image.normalizedDifference().rename('NDVI')) # 4. response \
# .toFloat()
# }
# Define the axes of variation in the collection array.
imageAxis = 0
bandAxis = 1
# Convert the collection to an array.
array = collection.map(makeVariables).toArray()
# Check the length of the image axis (number of images).
arrayLength = array.arrayLength(imageAxis)
# Update the mask to ensure that the number of images is greater than or
# equal to the number of predictors (the linear model is solveable).
array = array.updateMask(arrayLength.gt(4))
# Get slices of the array according to positions along the band axis.
predictors = array.arraySlice(bandAxis, 0, 4)
response = array.arraySlice(bandAxis, 4)
# Compute coefficients the easiest way.
coefficients3 = predictors.matrixSolve(response)
# Turn the results into a multi-band image.
coefficientsImage = coefficients3 \
.arrayProject([0]) \
.arrayFlatten([
['constant', 'trend', 'sin', 'cos']
])
print(coefficientsImage.getInfo())
Map.setCenter(-122.2627, 37.8735, 10)
Map.addLayer(coefficientsImage, {}, 'coefficientsImage')
```
## Display Earth Engine data layers
```
Map.addLayerControl() # This line is not needed for ipyleaflet-based Map.
Map
```
| github_jupyter |
## RDF
The radial distribution function (RDF) denoted in equations by g(r) defines the probability of finding a particle at a distance r from another tagged particle. The RDF is strongly dependent on the type of matter so will vary greatly for solids, gases and liquids.
<img src="../images/rdf.png" width="60%" height="60%">
As you might have observed the code complexity of the algorithm in $N^{2}$ . Let us get into details of the sequential code. **Understand and analyze** the code present at:
[RDF Serial Code](../../source_code/serial/rdf.cpp)
[File Reader](../../source_code/serial/dcdread.h)
[Makefile](../../source_code/serial/Makefile)
Open the downloaded file for inspection.
```
!cd ../../source_code/serial && make clean && make
```
We plan to follow the typical optimization cycle that every code needs to go through
<img src="../images/workflow.png" width="70%" height="70%">
In order analyze the application we we will make use of profiler "nsys" and add "nvtx" marking into the code to get more information out of the serial code. Before running the below cells, let's first start by divining into the profiler lab to learn more about the tools. Using Profiler gives us the hotspots and helps to understand which function is important to be made parallel.
-----
# <div style="text-align: center ;border:3px; border-style:solid; border-color:#FF0000 ; padding: 1em">[Profiling lab](../../../../../profiler/English/jupyter_notebook/profiling-c.ipynb)</div>
-----
Now, that we are familiar with the Nsight Profiler and know how to [NVTX](../../../../../profiler/English/jupyter_notebook/profiling-c.ipynb#nvtx), let's profile the serial code and checkout the output.
```
!cd ../../source_code/serial&& nsys profile -t nvtx --stats=true --force-overwrite true -o rdf_serial ./rdf
```
Once you run the above cell, you should see the following in the terminal.
<img src="../images/serial.png" width="70%" height="70%">
To view the profiler report, you would need to [Download the profiler output](../../source_code/serial/rdf_serial.qdrep) and open it via the GUI. For more information on how to open the report via the GUI, please checkout the section on [How to view the report](../../../../../profiler/English/jupyter_notebook/profiling-c.ipynb#gui-report).
From the timeline view, right click on the nvtx row and click the "show in events view". Now you can see the nvtx statistic at the bottom of the window which shows the duration of each range. In the following labs, we will look in to the profiler report in more detail.
<img src="../images/nvtx_serial.png" width="100%" height="100%">
The obvious next step is to make **Pair Calculation** algorithm parallel using different approaches to GPU Programming. Please follow the below link and choose one of the approaches to parallelise th serial code.
-----
# <div style="text-align: center ;border:3px; border-style:solid; border-color:#FF0000 ; padding: 1em">[HOME](../../../nways_MD_start.ipynb)</div>
-----
# Links and Resources
<!--[OpenACC API guide](https://www.openacc.org/sites/default/files/inline-files/OpenACC%20API%202.6%20Reference%20Guide.pdf)-->
[NVIDIA Nsight System](https://docs.nvidia.com/nsight-systems/)
<!--[NVIDIA Nsight Compute](https://developer.nvidia.com/nsight-compute)-->
<!--[CUDA Toolkit Download](https://developer.nvidia.com/cuda-downloads)-->
[Profiling timelines with NVTX](https://devblogs.nvidia.com/cuda-pro-tip-generate-custom-application-profile-timelines-nvtx/)
**NOTE**: To be able to see the Nsight System profiler output, please download Nsight System latest version from [here](https://developer.nvidia.com/nsight-systems).
Don't forget to check out additional [OpenACC Resources](https://www.openacc.org/resources) and join our [OpenACC Slack Channel](https://www.openacc.org/community#slack) to share your experience and get more help from the community.
---
## Licensing
This material is released by NVIDIA Corporation under the Creative Commons Attribution 4.0 International (CC BY 4.0).
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.