text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
# 머신 러닝 교과서 3판
# 16장 - 순환 신경망으로 순차 데이터 모델링 (2/2)
**아래 링크를 통해 이 노트북을 주피터 노트북 뷰어(nbviewer.jupyter.org)로 보거나 구글 코랩(colab.research.google.com)에서 실행할 수 있습니다.**
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://nbviewer.jupyter.org/github/rickiepark/python-machine-learning-book-3rd-edition/blob/master/ch16/ch16_part2.ipynb"><img src="https://jupyter.org/assets/main-logo.svg" width="28" />주피터 노트북 뷰어로 보기</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/rickiepark/python-machine-learning-book-3rd-edition/blob/master/ch16/ch16_part2.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />구글 코랩(Colab)에서 실행하기</a>
</td>
</table>
### 목차
- 텐서플로로 시퀀스 모델링을 위한 RNN 구현하기
- 두 번째 프로젝트: 텐서플로로 글자 단위 언어 모델 구현
- 데이터셋 전처리
- 문자 수준의 RNN 모델 만들기
- 평가 단계 - 새로운 텍스트 생성
- 트랜스포머 모델을 사용한 언어 이해
- 셀프 어텐션 메카니즘 이해하기
- 셀프 어텐션 기본 구조
- 쿼리, 키, 값 가중치를 가진 셀프 어텐션 메카니즘
- 멀티-헤드 어텐션과 트랜스포머 블록
- 요약
```
from IPython.display import Image
```
## 두 번째 프로젝트: 텐서플로로 글자 단위 언어 모델 구현
```
Image(url='https://git.io/JLdVE', width=700)
```
### 데이터셋 전처리
```
! curl -O http://www.gutenberg.org/files/1268/1268-0.txt
import numpy as np
## 텍스트 읽고 전처리하기
with open('1268-0.txt', 'r', encoding='UTF8') as fp:
text=fp.read()
start_indx = text.find('THE MYSTERIOUS ISLAND')
end_indx = text.find('End of the Project Gutenberg')
print(start_indx, end_indx)
text = text[start_indx:end_indx]
char_set = set(text)
print('전체 길이:', len(text))
print('고유한 문자:', len(char_set))
Image(url='https://git.io/JLdVz', width=700)
chars_sorted = sorted(char_set)
char2int = {ch:i for i,ch in enumerate(chars_sorted)}
char_array = np.array(chars_sorted)
text_encoded = np.array(
[char2int[ch] for ch in text],
dtype=np.int32)
print('인코딩된 텍스트 크기: ', text_encoded.shape)
print(text[:15], ' == 인코딩 ==> ', text_encoded[:15])
print(text_encoded[15:21], ' == 디코딩 ==> ', ''.join(char_array[text_encoded[15:21]]))
Image(url='https://git.io/JLdVV', width=700)
import tensorflow as tf
ds_text_encoded = tf.data.Dataset.from_tensor_slices(text_encoded)
for ex in ds_text_encoded.take(5):
print('{} -> {}'.format(ex.numpy(), char_array[ex.numpy()]))
seq_length = 40
chunk_size = seq_length + 1
ds_chunks = ds_text_encoded.batch(chunk_size, drop_remainder=True)
## inspection:
for seq in ds_chunks.take(1):
input_seq = seq[:seq_length].numpy()
target = seq[seq_length].numpy()
print(input_seq, ' -> ', target)
print(repr(''.join(char_array[input_seq])),
' -> ', repr(''.join(char_array[target])))
Image(url='https://git.io/JLdVr', width=700)
## x & y를 나누기 위한 함수를 정의합니다
def split_input_target(chunk):
input_seq = chunk[:-1]
target_seq = chunk[1:]
return input_seq, target_seq
ds_sequences = ds_chunks.map(split_input_target)
## 확인:
for example in ds_sequences.take(2):
print('입력 (x):', repr(''.join(char_array[example[0].numpy()])))
print('타깃 (y):', repr(''.join(char_array[example[1].numpy()])))
print()
# 배치 크기
BATCH_SIZE = 64
BUFFER_SIZE = 10000
tf.random.set_seed(1)
ds = ds_sequences.shuffle(BUFFER_SIZE).batch(BATCH_SIZE)# drop_remainder=True)
ds
```
### 문자 수준의 RNN 모델 만들기
```
def build_model(vocab_size, embedding_dim, rnn_units):
model = tf.keras.Sequential([
tf.keras.layers.Embedding(vocab_size, embedding_dim),
tf.keras.layers.LSTM(
rnn_units, return_sequences=True),
tf.keras.layers.Dense(vocab_size)
])
return model
charset_size = len(char_array)
embedding_dim = 256
rnn_units = 512
tf.random.set_seed(1)
model = build_model(
vocab_size = charset_size,
embedding_dim=embedding_dim,
rnn_units=rnn_units)
model.summary()
model.compile(
optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(
from_logits=True
))
model.fit(ds, epochs=20)
```
### 평가 단계 - 새로운 텍스트 생성
```
tf.random.set_seed(1)
logits = [[1.0, 1.0, 1.0]]
print('확률:', tf.math.softmax(logits).numpy()[0])
samples = tf.random.categorical(
logits=logits, num_samples=10)
tf.print(samples.numpy())
tf.random.set_seed(1)
logits = [[1.0, 1.0, 3.0]]
print('확률:', tf.math.softmax(logits).numpy()[0])
samples = tf.random.categorical(
logits=logits, num_samples=10)
tf.print(samples.numpy())
def sample(model, starting_str,
len_generated_text=500,
max_input_length=40,
scale_factor=1.0):
encoded_input = [char2int[s] for s in starting_str]
encoded_input = tf.reshape(encoded_input, (1, -1))
generated_str = starting_str
model.reset_states()
for i in range(len_generated_text):
logits = model(encoded_input)
logits = tf.squeeze(logits, 0)
scaled_logits = logits * scale_factor
new_char_indx = tf.random.categorical(
scaled_logits, num_samples=1)
new_char_indx = tf.squeeze(new_char_indx)[-1].numpy()
generated_str += str(char_array[new_char_indx])
new_char_indx = tf.expand_dims([new_char_indx], 0)
encoded_input = tf.concat(
[encoded_input, new_char_indx],
axis=1)
encoded_input = encoded_input[:, -max_input_length:]
return generated_str
tf.random.set_seed(1)
print(sample(model, starting_str='The island'))
```
* **예측 가능성 대 무작위성**
```
logits = np.array([[1.0, 1.0, 3.0]])
print('스케일 조정 전의 확률: ', tf.math.softmax(logits).numpy()[0])
print('0.5배 조정 후 확률: ', tf.math.softmax(0.5*logits).numpy()[0])
print('0.1배 조정 후 확률: ', tf.math.softmax(0.1*logits).numpy()[0])
tf.random.set_seed(1)
print(sample(model, starting_str='The island',
scale_factor=2.0))
tf.random.set_seed(1)
print(sample(model, starting_str='The island',
scale_factor=0.5))
```
# 트랜스포머 모델을 사용한 언어 이해
## 셀프 어텐션 메카니즘 이해하기
### 셀프 어텐션 기본 구조
```
Image(url='https://git.io/JLdVo', width=700)
```
### 쿼리, 키, 값 가중치를 가진 셀프 어텐션 메카니즘
## 멀티-헤드 어텐션과 트랜스포머 블록
```
Image(url='https://git.io/JLdV6', width=700)
```
...
# 요약
...
| github_jupyter |
# AU Fundamentals of Python Programming-Module2Problems (Part. A)
## M2-Q01 最大值與最小值
問題描述:
寫一個程式來找出輸入的 5 個數字的最大值和最小值,數值不限定為整數,且值可存放於 float 型態數值內。
輸入說明:
輸入5個數字
輸出說明:
輸出數列中的最大值與最小值,輸出時需附上小數點後兩位數字,最後必須有換行字元。
| Sample Input: | Sample Output: |
|:----------------|:-------------------------|
|-2 -15.2 0 9.5 100 | max=100.00 |
| |min=-15.20⏎ |
| 0 3 52.7 998 135 | max=998.00 |
| |min=0.00⏎ |
```
```
## M2-Q03 '*'反向三角形
問題描述:
讓使用者輸入一正整數 n,利用迴圈以字元 '*' 輸出高度為 n 的三角形。
輸入說明:
輸入一正整數 n。
輸出說明:
利用迴圈以字元 '*' 輸出高度為 n 的三角形,最後必須有換行字元。
| Sample Input: | Sample Output: |
|:----------------|:-------------------------|
|4 | \* |
| | \*\* |
| | \*\*\* |
| |\*\*\*\*⏎ |
```
```
## M2-Q05 十進制轉二進制
問題描述:
撰寫一個程式,使用者輸入一個非負整數,印出其 8 位元的二進制表示。
輸入說明:
輸入一個非負整數,介於 0~255 之間。
輸出說明:
以 8 位元的二進制表示,最後必須有換行字元。
| Sample Input: | Sample Output: |
|:----------------|:-------------------------|
|15 | 00001111⏎ |
|254 |11111110⏎ |
```
```
## M2-Q06 質數判別
問題描述:
試撰寫一個程式,由輸入一個整數,然後判別此數是否為質數。質數是指除了 1 和它本身之外,沒有其它的數可以整除它的數,例如:2,3,5,7 與 11 等皆為質數。
輸入說明:
輸入一個正整數。
輸出說明:
質數顯示 YES;非質數顯示 NO,最後必須有換行字元。
| Sample Input: | Sample Output: |
|:----------------|:-------------------------|
|23 | YES⏎ |
|37 |YES⏎ |
|39 |NO⏎ |
```
```
## M2-Q07 考試測驗
問題描述:
某間學校舉辦英文檢定測驗,若是學生通過該測驗,則能通過畢業門檻。老師準備的英文測驗中分為三個項目當作考試,分別是聽力、閱讀、口說。每一個項目測驗滿分皆為 100 分,總分為 300 分。想要通過測驗有兩種方式。
方式一:若是三個項目分數皆為 60 分以上(包含 60 分)即為通過測驗。
方式二:
1. 若三個項目中有其中一個項目分數未滿 60 分,但三個項目分數總合大於等於220 分,也可算通過測驗。
2. 若三個項目中有其中一個項目分數未滿 60 分,而三個項目分數總合也沒能達到 220 分,可獲得補考機會。
3. 若是三個項目中有兩個項目不及格,但另一個項目成績高於 80 分(包含 80 分),也可獲得補考的機會。
其餘皆判定為無法通過測驗。
輸入說明:
第一行為一個整數 N,代表共有 N 組測試資料。之後有 N 行,每一行有 3 個非負整數(範圍皆為 0 到 100),分別代表該名學生聽力、閱讀、口說的測驗分數。
輸出說明:
若是通過測驗,則輸出“P”。若是需要補考,則輸出“M”。若是無法通過測驗,則輸出“F”。每組答案結果輸出於一行,最後必須有換行字元。
| Sample Input: | Sample Output: |
|:----------------|:-------------------------|
|5 | P |
|95 86 100 |M |
|30 60 80 |P |
|80 55 85 | F |
|20 30 60 |M |
|10 80 10 | |
```
```
## M2-Q10 n×n 乘法表
問題描述:
輸出 n×n 乘法表。
輸入說明:
輸入一正整數 n。
輸出說明:
輸出 n×n 乘法表(0<n<=9),每個輸出數字皆以 tab 間格,最後必須有換行字元。
| Sample Input: | Sample Output: |
|:----------------|:-------------------------|
|8 | 1 2 3 4 5 6 7 8 |
| | 2 4 6 8 10 12 14 16 |
| | 3 6 9 12 15 18 21 24 |
| | 4 8 12 16 20 24 28 32 |
| | 5 10 15 20 25 30 35 40 |
| | 6 12 18 24 30 36 42 48 |
| | 7 14 21 28 35 42 49 56 |
| | 8 16 24 32 40 48 56 64⏎ |
|5 |1 2 3 4 5 |
| | 2 4 6 8 10 |
| | 3 6 9 12 15 |
| | 4 8 12 16 20 |
| | 5 10 15 20 25⏎ |
```
```
## M2-Q11 正因數
問題描述:
讓使用者輸入一正整數 n,輸出 n 的所有正因數。
輸入說明:
輸入一正整數 n。
輸出說明:
輸出 n 的所有正因數,最後必須有換行字元。
| Sample Input: | Sample Output: |
|:----------------|:-------------------------|
|5 | 1 |
| |5⏎ |
|10 | 1 |
| |2 |
| | 5 |
| |10⏎ |
```
```
## M2-Q12 實心正方形
問題描述:
試撰寫一程式,由使用者輸入一正整數 n 及字元 c,輸出以 c 字元印出邊長為 n 之實心正方形。
輸入說明:
分別輸入一正整數 n 及字元 c。
輸出說明:
輸出以 c 字元,印出邊長為 n 之實心正方形,最後必須有換行字元。
| Sample Input: | Sample Output: |
|:----------------|:-------------------------|
|3@ | \@\@\@ |
| |\@\@\@ |
| |\@\@\@⏎ |
```
```
| github_jupyter |
# Definition
The RSA algorithm involves four steps: key generation, key distribution, encryption and decryption.
A basic principle behind RSA is the observation that it is practical to find three very large positive integers e, d and n such that with modular exponentiation for all integer m:
$(m^{e})^{d}\equiv m{\pmod {n}}$
and that even knowing e and n or even m it can be extremely difficult to find d.
Additionally, for some operations it is convenient that the order of the two exponentiations can be changed and that this relation also implies:
$(m^{d})^{e}\equiv m{\pmod {n}}$
RSA involves a public key and a private key. The public key can be known by everyone and is used for encrypting messages. The intention is that messages encrypted with the public key can only be decrypted in a reasonable amount of time using the private key. The public key is represented by the integers n and e; and, the private key, by the integer d (although n is also used during the decryption process; so, it might be considered a part of the private key, too). m represents the message (previously prepared with certain technique explained below).
## Key generation
The keys for the RSA algorithm are generated the following way:
1. Choose two distinct prime numbers p and q.
* For security purposes, the integers p and q should be chosen at random, and should be similar in magnitude but 'differ in length by a few digits'[2] to make factoring harder. Prime integers can be efficiently found using a primality test.
1.Compute n = pq.
* n is used as the modulus for both the public and private keys. Its length, usually expressed in bits, is the key length.
1. Compute λ(n) = lcm(λ(p), λ(q)) = lcm(p − 1, q − 1), where λ is Carmichael's totient function. This value is kept private.
1. Choose an integer e such that 1 < e < λ(n) and gcd(e, λ(n)) = 1; i.e., e and λ(n) are coprime.
1. Determine d as d ≡ e−1 (mod λ(n)); i.e., d is the modular multiplicative inverse of e (modulo λ(n)).
* This is more clearly stated as: solve for d given d⋅e ≡ 1 (mod λ(n)).
* e having a short bit-length and small Hamming weight results in more efficient encryption – most commonly e = 216 + 1 = 65,537. However, much smaller values of e (such as 3) have been shown to be less secure in some settings.[14]
* e is released as the public key exponent.
* d is kept as the private key exponent.
```
import secrets
import math
"""
Euler's totient function
"""
def phi(n):
count = 0
for i in range(1, n + 1):
if math.gcd(i, n) == 1:
count += 1
return count
def egcd(a, b):
if a == 0:
return (b, 0, 1)
else:
g, y, x = egcd(b % a, a)
return (g, x - (b // a) * y, y)
def modinv(a, m):
g, x, y = egcd(a, m)
if g != 1:
raise Exception('modular inverse does not exist')
else:
return x % m
# Load prime numbers from 0 to 100,000
primes = []
with open('primes-to-100k.txt') as f:
primes = [int(line.split()[0]) for line in f]
RANGE = 10#len(primes)
# Choose two random primes p and q
halflen = RANGE//2
p = primes[secrets.randbelow(halflen)]
q = primes[halflen+secrets.randbelow(halflen)]
# Calculate modu
n = p*q
phi_n = (p-1)*(q-1)
# Choose e (public key) such as 1 < e < phi_n
e = phi_n
while e >= phi_n or math.gcd(e, phi_n) != 1:
e = primes[1+secrets.randbelow(RANGE)]
# Calculate d (private key) such as de ≡ 1 (mod ϕ(n)) ie: de = 1 + kϕ(n)
d = modinv(e, phi_n)
# Encrypt number
msg = 12
enc = msg ** e % n
dec = enc ** d % n
print(msg, enc, dec)
```
# References
1. Wikipedia, "RSA (cryptosystem)", https://en.wikipedia.org/wiki/RSA_(cryptosystem)
2. Art of the Problem, "Public Key Cryptography: RSA Encryption Algorithm", https://youtu.be/wXB-V_Keiu8
| github_jupyter |
# BIDMach: parameter tuning
In this notebook we'll explore automated parameter exploration by grid search.
```
import $exec.^.lib.bidmach_notebook_init
if (Mat.hasCUDA > 0) GPUmem
```
## Dataset: Reuters RCV1 V2
The dataset is the widely used Reuters news article dataset RCV1 V2. This dataset and several others are loaded by running the script <code>getdata.sh</code> from the BIDMach/scripts directory. The data include both train and test subsets, and train and test labels (cats).
```
var dir = "../data/rcv1/" // adjust to point to the BIDMach/data/rcv1 directory
tic
val train = loadSMat(dir+"docs.smat.lz4")
val cats = loadFMat(dir+"cats.fmat.lz4")
val test = loadSMat(dir+"testdocs.smat.lz4")
val tcats = loadFMat(dir+"testcats.fmat.lz4")
toc
```
First lets enumerate some parameter combinations for learning rate and time exponent of the optimizer (texp)
```
val lrates = col(0.03f, 0.1f, 0.3f, 1f) // 4 values
val texps = col(0.3f, 0.4f, 0.5f, 0.6f, 0.7f) // 5 values
```
The next step is to enumerate all pairs of parameters. We can do this using the kron operator for now, this will eventually be a custom function:
```
val lrateparams = ones(texps.nrows, 1) ⊗ lrates
val texpparams = texps ⊗ ones(lrates.nrows,1)
lrateparams \ texpparams
```
Here's the learner again:
```
val (mm, opts) = GLM.learner(train, cats, GLM.logistic)
```
To keep things simple, we'll focus on just one category and train many models for it. The "targmap" option specifies a mapping from the actual base categories to the model categories. We'll map from category six to all our models:
```
val nparams = lrateparams.length
val targmap = zeros(nparams, 103)
targmap(?,6) = 1
opts.targmap = targmap
opts.lrate = lrateparams
opts.texp = texpparams
mm.train
val (pp, popts) = GLM.predictor(mm.model, test)
```
And invoke the predict method on the predictor:
```
pp.predict
val preds = FMat(pp.preds(0))
pp.model.asInstanceOf[GLM].mats.length
```
Although ll values are printed above, they are not meaningful (there is no target to compare the prediction with).
We can now compare the accuracy of predictions (preds matrix) with ground truth (the tcats matrix).
```
val vcats = targmap * tcats // create some virtual cats
val lls = mean(ln(1e-7f + vcats ∘ preds + (1-vcats) ∘ (1-preds)),2) // actual logistic likelihood
mean(lls)
```
A more thorough measure is ROC area:
```
val rocs = roc2(preds, vcats, 1-vcats, 100) // Compute ROC curves for all categories
plot(rocs)
val aucs = mean(rocs)
```
The maxi2 function will find the max value and its index.
```
val (bestv, besti) = maxi2(aucs)
```
And using the best index we can find the optimal parameters:
```
texpparams(besti) \ lrateparams(besti)
```
> Write the optimal values in the cell below:
<b>Note:</b> although our parameters lay in a square grid, we could have enumerated any sequence of pairs, and we could have searched over more parameters. The learner infrastructure supports more intelligent model optimization (e.g. Bayesian methods).
| github_jupyter |
<table> <tr>
<td style="background-color:#ffffff;">
<a href="http://qworld.lu.lv" target="_blank"><img src="../images/qworld.jpg" width="25%" align="left"> </a></td>
<td style="background-color:#ffffff;vertical-align:bottom;text-align:right;">
prepared by <a href="http://abu.lu.lv" target="_blank">Abuzer Yakaryilmaz</a> (<a href="http://qworld.lu.lv/index.php/qlatvia/" target="_blank">QLatvia</a>)
</td>
</tr></table>
<table width="100%"><tr><td style="color:#bbbbbb;background-color:#ffffff;font-size:11px;font-style:italic;text-align:right;">This cell contains some macros. If there is a problem with displaying mathematical formulas, please run this cell to load these macros. </td></tr></table>
$ \newcommand{\bra}[1]{\langle #1|} $
$ \newcommand{\ket}[1]{|#1\rangle} $
$ \newcommand{\braket}[2]{\langle #1|#2\rangle} $
$ \newcommand{\dot}[2]{ #1 \cdot #2} $
$ \newcommand{\biginner}[2]{\left\langle #1,#2\right\rangle} $
$ \newcommand{\mymatrix}[2]{\left( \begin{array}{#1} #2\end{array} \right)} $
$ \newcommand{\myvector}[1]{\mymatrix{c}{#1}} $
$ \newcommand{\myrvector}[1]{\mymatrix{r}{#1}} $
$ \newcommand{\mypar}[1]{\left( #1 \right)} $
$ \newcommand{\mybigpar}[1]{ \Big( #1 \Big)} $
$ \newcommand{\sqrttwo}{\frac{1}{\sqrt{2}}} $
$ \newcommand{\dsqrttwo}{\dfrac{1}{\sqrt{2}}} $
$ \newcommand{\onehalf}{\frac{1}{2}} $
$ \newcommand{\donehalf}{\dfrac{1}{2}} $
$ \newcommand{\hadamard}{ \mymatrix{rr}{ \sqrttwo & \sqrttwo \\ \sqrttwo & -\sqrttwo }} $
$ \newcommand{\vzero}{\myvector{1\\0}} $
$ \newcommand{\vone}{\myvector{0\\1}} $
$ \newcommand{\stateplus}{\myvector{ \sqrttwo \\ \sqrttwo } } $
$ \newcommand{\stateminus}{ \myrvector{ \sqrttwo \\ -\sqrttwo } } $
$ \newcommand{\myarray}[2]{ \begin{array}{#1}#2\end{array}} $
$ \newcommand{\X}{ \mymatrix{cc}{0 & 1 \\ 1 & 0} } $
$ \newcommand{\I}{ \mymatrix{rr}{1 & 0 \\ 0 & 1} } $
$ \newcommand{\Z}{ \mymatrix{rr}{1 & 0 \\ 0 & -1} } $
$ \newcommand{\Htwo}{ \mymatrix{rrrr}{ \frac{1}{2} & \frac{1}{2} & \frac{1}{2} & \frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & \frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} & \frac{1}{2} } } $
$ \newcommand{\CNOT}{ \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0} } $
$ \newcommand{\norm}[1]{ \left\lVert #1 \right\rVert } $
$ \newcommand{\pstate}[1]{ \lceil \mspace{-1mu} #1 \mspace{-1.5mu} \rfloor } $
$ \newcommand{\greenbit}[1] {\mathbf{{\color{green}#1}}} $
$ \newcommand{\bluebit}[1] {\mathbf{{\color{blue}#1}}} $
$ \newcommand{\redbit}[1] {\mathbf{{\color{red}#1}}} $
$ \newcommand{\brownbit}[1] {\mathbf{{\color{brown}#1}}} $
<h2> Basics of Python: Lists </h2>
We review using Lists in Python here.
Please run each cell and check the results.
A list (or array) is a collection of objects (variables) separated by comma.
The order is important, and we can access each element in the list with its index starting from 0.
```
# here is a list holding all even numbers between 10 and 20
L = [10, 12, 14, 16, 18, 20]
# let's print the list
print(L)
# let's print each element by using its index but in reverse order
print(L[5],L[4],L[3],L[2],L[1],L[0])
# let's print the length (size) of list
print(len(L))
# let's print each element and its index in the list
# we use a for-loop, and the number of iteration is determined by the length of the list
# everthing is automatical :-)
L = [10, 12, 14, 16, 18, 20]
for i in range(len(L)):
print(L[i],"is the element in our list with the index",i)
# let's replace each number in the above list with its double value
# L = [10, 12, 14, 16, 18, 20]
# let's print the list before doubling operation
print("the list before doubling operation is",L)
for i in range(len(L)):
current_element=L[i] # get the value of the i-th element
L[i] = 2 * current_element # update the value of the i-th element
# let's shorten the code as
#L[i] = 2 * L[i]
# or
#L[i] *= 2
# let's print the list after doubling operation
print("the list after doubling operation is",L)
# after each execution of this cell, the latest values will be doubled
# so the values in the list will be exponentially increased
# let's define two lists
L1 = [1,2,3,4]
L2 = [-5,-6,-7,-8]
# two lists can be concatenated
# the result is a new list
print("the concatenation of L1 and L2 is",L1+L2)
# the order of terms is important
print("the concatenation of L2 and L1 is",L2+L1) # this is a different list than L1+L2
# we can add a new element to a list, which increases its length/size by 1
L = [10, 12, 14, 16, 18, 20]
print(L,"the current length is",len(L))
# we add two values by showing two different methods
# L.append(value) directly adds the value as a new element to the list
L.append(-4)
# we can also use concatenation operator +
L = L + [-8] # here [-8] is a list having a single element
print(L,"the new length is",len(L))
# a list can be multiplied with an integer
L = [1,2]
# we can consider the multiplication of L by an integer as a repeated summation (concatenation) of L by itself
# L * 1 is the list itself
# L * 2 is L + L (the concatenation of L with itself)
# L * 3 is L + L + L (the concatenation of L with itself twice)
# L * m is L + ... + L (the concatenation of L with itself m-1 times)
# L * 0 is the empty list
# L * i is the same as i * L
# let's print the different cases
for i in range(6):
print(i,"* L is",i*L)
# this operation can be useful when initializing a list with the same value(s)
# let's create a list of prime numbers less than 100
# here is a function that determines whether a given number is prime or not
def prime(number):
if number < 2: return False
if number == 2: return True
if number % 2 == 0: return False
for i in range(3,number,2):
if number % i == 0: return False
return True
# end of a function
# let's start with an empty list
L=[]
# what can the length of this list be?
print("my initial length is",len(L))
for i in range(2,100):
if prime(i):
L.append(i)
# alternative methods:
#L = L + [i]
#L += [i]
# print the final list
print(L)
print("my final length is",len(L))
```
For a given integer $n \geq 0$, $ S(0) = 0 $, $ S(1)=1 $, and $ S(n) = 1 + 2 + \cdots + n $.
We define list $ L(n) $ such that the element with index $n$ holds $ S(n) $.
In other words, the elements of $ L(n) $ are $ [ S(0)~~S(1)~~S(2)~~\cdots~~S(n) ] $.
Let's build the list $ L(20) $.
```
# let's define the list with S(0)
L = [0]
# let's iteratively define n and S
# initial values
n = 0
S = 0
# the number of iterations
N = 20
while n <= N: # we iterate all values from 1 to 20
n = n + 1
S = S + n
L.append(S)
# print the final list
print(L)
```
<h3> Task 1 </h3>
Fibonacci sequence starts with $ 1 $ and $ 1 $. Then, each next element is equal to the summation of the previous two elements:
$$
1, 1, 2 , 3 , 5, 8, 13, 21, 34, 55, \ldots
$$
Find the first 30 elements of the Fibonacci sequence, store them in a list, and then print the list.
You can verify the first 10 elements of your result with the above list.
```
#
# your solution is here
#
F = [1,1]
```
<a href="Python20_Basics_Lists_Solutions.ipynb#task1">click for our solution</a>
<h3> Lists of different objects </h3>
A list can have any type of values.
```
# the following list stores certain information about Asja
# name, surname, age, profession, height, weight, partner(s) if any, kid(s) if any, the creation date of list
ASJA = ['Asja','Sarkane',34,'musician',180,65.5,[],['Eleni','Fyodor'],"October 24, 2018"]
print(ASJA)
# Remark that an element of a list can be another list as well.
```
<h3> Task 2 </h3>
Define a list $ N $ with 11 elements such that $ N[i] $ is another list with four elements such that $ [i, i^2, i^3, i^2+i^3] $.
The index $ i $ should be between $ 0 $ and $ 10 $.
```
#
# your solution is here
#
```
<a href="Python20_Basics_Lists_Solutions.ipynb#task2">click for our solution</a>
<h3> Dictionaries </h3>
The outcomes of a quantum program (circuit) will be stored in a dictionary.
Therefore, we very shortly mention about the dictionary data type.
A dictionary is a set of paired elements.
Each pair is composed by a key and its value, and any value can be accessed by its key.
```
# let's define a dictionary pairing a person with her/his age
ages = {
'Asja':32,
'Balvis':28,
'Fyodor':43
}
# let print all keys
for person in ages:
print(person)
# let's print the values
for person in ages:
print(ages[person])
```
| github_jupyter |
# Mini-project I: Parameter estimation for a toy model of an EFT
The overall project goal is to reproduce various results in a paper co-authored by Daniel and Dick (and others): [*Bayesian parameter estimation for effective field theories*](https://arxiv.org/abs/1511.03618). It's a long paper, so don't try to read all of it! (At least not now.) We'll guide you to the relevant parts.
The paper uses toy models for effective field theories, namely Taylor series of some specified functions, to present guidelines for parameter estimation. This will also be a check of whether you can follow (or give you practice on) Bayesian statistics discussions in the physics literature.
You'll find summaries in section II that touch on topics we have discussed and will discuss. The function
$$
g(x) = \left(\frac12 + \tan\left(\frac{\pi}{2}x\right)\right)^2
$$
represents the true, underlying theory. It has a Taylor expansion
$$
g(x) = 0.25 + 1.57x + 2.47x^2 + 1.29 x^3 + \cdots
$$
Our model for an EFT for this "theory" is
$$
g_{\rm th}(x) \equiv \sum_{i=0}^k a_i x^i
$$
and your general task is to fit 1, 2, 3, ... of the constants $a_i$, and analyze the results.
**Your primary goal is to reproduce and interpret Table III on page 12 of the arXiv preprint. A secondary goal is to reproduce Figure 1 of the same paper.** You should use the emcee sampler and corner to make plots.
This a less-guided set of tasks than the ones we've done so far, which will have you put together ideas and tools we've discussed. You'll work on the mini-project the first Thursday and Friday afternoons in the exercise session and we'll do a recap early next week. There is nothing to hand in but we'll be happy to review what you come up with.
<div style="float:center;"><img src="summary_of_project.png" width=700px></div>
### Learning goals:
* Apply and extend the Bayesian parameter estimation ideas and techniques from the course.
* Explore the impact of control features: dependence on how much data is used and how precise it is; apply an *informative* prior.
* Learn about some diagnostics for Bayesian parameter estimation.
* Try out sampling on a controlled problem.
### Suggestions for how to proceed:
* Follow the lead of the notebooks [Intro notebook revisited [ipynb]](https://github.com/NuclearTalent/Bayes2019/blob/master/topics/bayesian-parameter-estimation/parameter_estimation_in_bayesTALENT_intro.ipynb) and [Fitting a straight line II [ipynb]](https://github.com/NuclearTalent/Bayes2019/blob/master/topics/why-bayes-is-better/parameter_estimation_fitting_straight_line_II.ipynb).
* Define a function for the exact result plus noise, noting from the arXiv paper what type of noise is added and where the points are located (i.e., what values of $x$).
* Define functions for the two choices of prior and for the likelihood.
* Call emcee to sample the posteriors.
* Use corner to create plots. You can read the answers for the tables from the corner plots.
* Don't try to do too much in your code at first (start with the lowest order in Table III).
* Fill in the rest of Table III.
* Generate figures for the lowest orders analogous to Figure 1 and then reproduce Figure 1.
### Comments and suggestions
* The 5% error is a *relative* error, meaning it is 0.05 times the data at that point. This means if you generate a Gaussian random number `err` distributed with standard deviation 0.05, the value of sigma for the log likelihood is `sigma[i] = data[i] * err` (use the data, not the theory at `i`).
* The `show_titles=True` option to corner will show central results and one-$\sigma$ error limits on the projected posterior plots.
* The `quantiles=[0.16, 0.5, 0.84]`option to corner adds the dashed vertical lines to the marginal posteriors on the diagonal. You can obviously change the quantiles if you want another credibility region.
* The python command `np.percentile(samples, [16, 50, 84],axis=0)` might be useful to extract numerical values for the credibility region and the mean from a python array `samples`of shape (nsamples,ndimensions).
* The example on [Fitting a Model to Data](https://emcee.readthedocs.io/en/v2.2.1/user/line/) from the emcee documentation may be useful to supplement the examples in the TALENT notebooks.
### Additional subtasks (for now and the future)
* Reproduce Figures 3 and 4, showing the predictions with error bands for the two priors compared to the true function. You can use Matplotlib's `fill_between(x, y-error, y+error)` to make bands. (Use the `alpha` keyword for `fill_between`, e.g., `alpha=0.5`, to make the bands more transparent.)
* Reproduce Figures 5 (alternative prior and "returning the prior"), 6 (posterior for $\overline a$), and 7 ("relaxing to least squares").
* Reproduce Figure 9 (sensitivity to choice of $x_{\rm max}$)
* Repeat analysis with same function but different data precision and/or quantity (number of data points).
* Repeat analysis with a different function from the paper or invent your own function and analyze.
| github_jupyter |
```
import random
prime = 0x0800000000000011000000000000000000000000000000000000000000000001
def div_mod(a, b, m):
return (a * pow(b, m - 2, m)) % m
def rand():
return random.randrange(0, prime)
def limbs(n):
limbs = []
for i in range(8):
limbs += [n % 2**32]
n >>= 32
return limbs
def constant(n):
limbs = []
for i in range(8):
limbs += [n % 2**32]
n >>= 32
return ', '.join(['0x{:08x}'.format(i) for i in limbs])
def U256(n):
return 'u256h!("{:064x}")'.format(n)
def gcdex(a, b):
p = b
x, y, r, s = 1, 0, 0, 1
while b:
(c, q) = (a % b, a // b)
(a, b, r, s, x, y) = (b, c, x-q*r, y-q*s, r, s)
return (x, y, a)
def modinv(n, m):
a, b, c = gcdex(n, m)
assert c == 1
return a % m
n = rand() % prime
print(U256(n))
print(U256(modinv(n, prime)))
(0x018a5cc4c55ac5b050a0831b65e827e5e39fd4515e4e094961c61509e7870814 *
0x0713ccbc2d1786937a9854a7c169625681304f782ec8426660f1ba26988fc815) % prime
u = 0x002d7b8fcb8a16f362c38a1fb043372572e3c59bfd426cef6558622707f7be71
v = 0x00ec3343d2e8797d8567ab583e969da97ecfb087d137bd999f0e45d9677037ec
prime * u - n * v
v * (prime - n) % prime
def U2562(n):
return 'u256h!("{:064x}")'.format(n)
prime = 3618502788666131213697322783095070105623107215331596699973092056135872020481
U256(1479382000703807775820380567681950527890950161973239379326216893866213995534)
print(constant(prime - 2))
print(U256(prime - 2))
print(U2562(prime - 2))
constant(2**251 + 2**196 + 2**192 - 1)
constant(prime)
constant(prime-2)
R = 2**264 % prime
R2 = (R * R) % prime
R3 = (R2 * R) % prime
# -(m^{-1} mod m) mod m
x = rand()
r = modinv(x, prime)
print(U256(x))
print(U256(r))
a = rand()
b = rand()
m = rand() >> 23
print(U256(a))
print(U256(b))
print(U256(m))
e = (a * b) % m
print(U256(e))
r = 0x000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b016000000000000000000000000000000000
r
a * b
print(r % m)
print(e)
print(m)
B = 2**256
def bogo_reduce(lo, hi, m):
print(hi)
if hi == 0:
return lo % m
a = B % m
rlo = (a * hi) % B
rhi = (a * hi) // B
lo = (lo + rlo) % m
return bogo_reduce(lo, rhi, m)
a = rand()
b = rand()
m = prime
bogo_reduce(a, b, m)
(a + b * B) % m
X = rand()
Y = rand()
Z = (X * pow(Y, prime - 2, prime)) % prime
constant(X)
constant(Y)
constant(Z)
X
Y
Z
3224672152922917067672837989388327988581742594340864239845412773431323678197
4174868897963978531447481309916031480477542228864300399314946096445175879634
Z < prime
prime
def ec_add(point1, point2, p):
"""
Gets two points on an elliptic curve mod p and returns their sum.
Assumes the points are given in affine form (x, y) and have different x coordinates.
"""
assert (point1[0] - point2[0]) % p != 0
m = div_mod(point1[1] - point2[1], point1[0] - point2[0], p)
x = (m * m - point1[0] - point2[0]) % p
y = (m * (point1[0] - x) - point1[1]) % p
return x, y
ax = rand()
ay = rand()
bx = rand()
by = rand()
cx, cy = ec_add((ax, ay), (bx, by), prime)
constant(ax)
constant(ay)
constant(bx)
constant(by)
constant(cx)
constant(cy)
def ec_double(point, alpha, p):
"""
Doubles a point on an elliptic curve with the equation y^2 = x^3 + alpha*x + beta mod p.
Assumes the point is given in affine form (x, y) and has y != 0.
"""
assert point[1] % p != 0
m = div_mod(3 * point[0] * point[0] + alpha, 2 * point[1], p)
x = (m * m - 2 * point[0]) % p
y = (m * (point[0] - x) - point[1]) % p
return x, y
ax = rand()
ay = rand()
bx, by = ec_double((ax, ay), 1, prime)
constant(ax)
constant(ay)
constant(bx)
constant(by)
def ec_mult(m, point, alpha, p):
"""
Multiplies by m a point on the elliptic curve with equation y^2 = x^3 + alpha*x + beta mod p.
Assumes the point is given in affine form (x, y) and that 0 < m < order(point).
"""
if m == 1:
return point
if m % 2 == 0:
return ec_mult(m // 2, ec_double(point, alpha, p), alpha, p)
return ec_add(ec_mult(m - 1, point, alpha, p), point, p)
ax = rand()
ay = rand()
b = rand()
cx, cy = ec_mult(b, (ax, ay), 1, prime)
constant(ax)
constant(ay)
constant(b)
constant(cx)
constant(cy)
constant(0x800000000000010ffffffffffffffffb781126dcae7b2321e66a241adc64d2f)
a = rand()
b = rand()
p = (rand(), rand())
lhs = ec_add(ec_mult(a, p, 1, prime), ec_mult(b, p, 1, prime), prime)
rhs = ec_mult(a + b, p, 1, prime)
lhs == rhs
D = [
[285, 0x84b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c000000000000000000000000000000000000000000000000000000000000000000000000, 0x000d7d418f2575f2fd73563a65410a6ca4bcd239abf3fab0d1e31ce288448e3f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[284, 0x425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b016000000000000000000000000000000000000000000000000000000000000000000000000, 0x000d7d418f2575f2fd73563a65410a6ca4bcd239abf3fab0d1e31ce288448e3f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[283, 0x212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b000000000000000000000000000000000000000000000000000000000000000000000000, 0x000d7d418f2575f2fd73563a65410a6ca4bcd239abf3fab0d1e31ce288448e3f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[282, 0x10969a86220d2f8df39776207e6ea153420e2218781224f105c02c05800000000000000000000000000000000000000000000000000000000000000000000000, 0x000d7d418f2575f2fd73563a65410a6ca4bcd239abf3fab0d1e31ce288448e3f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[281, 0x084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c00000000000000000000000000000000000000000000000000000000000000000000000, 0x000d7d418f2575f2fd73563a65410a6ca4bcd239abf3fab0d1e31ce288448e3f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[280, 0x0425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b01600000000000000000000000000000000000000000000000000000000000000000000000, 0x000d7d418f2575f2fd73563a65410a6ca4bcd239abf3fab0d1e31ce288448e3f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[279, 0x0212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b00000000000000000000000000000000000000000000000000000000000000000000000, 0x000d7d418f2575f2fd73563a65410a6ca4bcd239abf3fab0d1e31ce288448e3f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[278, 0x010969a86220d2f8df39776207e6ea153420e2218781224f105c02c0580000000000000000000000000000000000000000000000000000000000000000000000, 0x000d7d418f2575f2fd73563a65410a6ca4bcd239abf3fab0d1e31ce288448e3f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[277, 0x0084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c0000000000000000000000000000000000000000000000000000000000000000000000, 0x000d7d418f2575f2fd73563a65410a6ca4bcd239abf3fab0d1e31ce288448e3f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[276, 0x00425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b0160000000000000000000000000000000000000000000000000000000000000000000000, 0x000d7d418f2575f2fd73563a65410a6ca4bcd239abf3fab0d1e31ce288448e3f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[275, 0x00212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b0000000000000000000000000000000000000000000000000000000000000000000000, 0x000d7d418f2575f2fd73563a65410a6ca4bcd239abf3fab0d1e31ce288448e3f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[274, 0x0010969a86220d2f8df39776207e6ea153420e2218781224f105c02c058000000000000000000000000000000000000000000000000000000000000000000000, 0x000d7d418f2575f2fd73563a65410a6ca4bcd239abf3fab0d1e31ce288448e3f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[273, 0x00084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c000000000000000000000000000000000000000000000000000000000000000000000, 0x000531f44c146f5b36798a7f5501d31bfb1bcb289fb7f19e59603ccc85848e3f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[272, 0x000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b016000000000000000000000000000000000000000000000000000000000000000000000, 0x00010c4daa8bec0f52fca4a1cce23773a64b47a01999ed151d1eccc184248e3f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[271, 0x000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b000000000000000000000000000000000000000000000000000000000000000000000, 0x00010c4daa8bec0f52fca4a1cce23773a64b47a01999ed151d1eccc184248e3f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[270, 0x00010969a86220d2f8df39776207e6ea153420e2218781224f105c02c05800000000000000000000000000000000000000000000000000000000000000000000, 0x000002e40229cb3c5a1d6b2a6ada5089911726bdf8126bf2ce0e70bec3cc8e3f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[269, 0x000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c00000000000000000000000000000000000000000000000000000000000000000000, 0x000002e40229cb3c5a1d6b2a6ada5089911726bdf8126bf2ce0e70bec3cc8e3f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[268, 0x0000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b01600000000000000000000000000000000000000000000000000000000000000000000, 0x000002e40229cb3c5a1d6b2a6ada5089911726bdf8126bf2ce0e70bec3cc8e3f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[267, 0x0000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b00000000000000000000000000000000000000000000000000000000000000000000, 0x000002e40229cb3c5a1d6b2a6ada5089911726bdf8126bf2ce0e70bec3cc8e3f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[266, 0x000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c0580000000000000000000000000000000000000000000000000000000000000000000, 0x000002e40229cb3c5a1d6b2a6ada5089911726bdf8126bf2ce0e70bec3cc8e3f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[265, 0x0000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c0000000000000000000000000000000000000000000000000000000000000000000, 0x000002e40229cb3c5a1d6b2a6ada5089911726bdf8126bf2ce0e70bec3cc8e3f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[264, 0x00000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b0160000000000000000000000000000000000000000000000000000000000000000000, 0x000002e40229cb3c5a1d6b2a6ada5089911726bdf8126bf2ce0e70bec3cc8e3f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[263, 0x00000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b0000000000000000000000000000000000000000000000000000000000000000000, 0x000000d12ed906fab42bacb77c1640bbbcecbe7c33cf5cf089705006be4bde3f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[262, 0x0000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c058000000000000000000000000000000000000000000000000000000000000000000, 0x000000d12ed906fab42bacb77c1640bbbcecbe7c33cf5cf089705006be4bde3f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[261, 0x00000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c000000000000000000000000000000000000000000000000000000000000000000, 0x0000004c7a04d5ea4aaf3d1ac0653cc847e2246bc2be992ff848c7d8bcebb23f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[260, 0x000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b016000000000000000000000000000000000000000000000000000000000000000000, 0x0000000a1f9abd6215f1054c628cbace8d5cd7638a36374fafb503c1bc3b9c3f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[259, 0x000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b000000000000000000000000000000000000000000000000000000000000000000, 0x0000000a1f9abd6215f1054c628cbace8d5cd7638a36374fafb503c1bc3b9c3f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[258, 0x00000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c05800000000000000000000000000000000000000000000000000000000000000000, 0x0000000a1f9abd6215f1054c628cbace8d5cd7638a36374fafb503c1bc3b9c3f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[257, 0x000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c00000000000000000000000000000000000000000000000000000000000000000, 0x00000001d44d7a510f593e5296d1aa8f560c2dc283252b13a6a28b3edc25997f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[256, 0x0000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b01600000000000000000000000000000000000000000000000000000000000000000, 0x00000001d44d7a510f593e5296d1aa8f560c2dc283252b13a6a28b3edc25997f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[255, 0x0000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b00000000000000000000000000000000000000000000000000000000000000000, 0x00000001d44d7a510f593e5296d1aa8f560c2dc283252b13a6a28b3edc25997f4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[254, 0x000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c0580000000000000000000000000000000000000000000000000000000000000000, 0x00000000cae3d1eeee8645735d5a48876f22188e6243098c25803c2e8022d9274d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[253, 0x0000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c0000000000000000000000000000000000000000000000000000000000000000, 0x00000000462efdbdde1cc903c09e97837bad0df451d1f8c864ef14a6522178fb4d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[252, 0x00000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b0160000000000000000000000000000000000000000000000000000000000000000, 0x0000000003d493a555e80acbf240bf0181f288a74999706684a680e23b20c8e54d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[251, 0x00000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b0000000000000000000000000000000000000000000000000000000000000000, 0x0000000003d493a555e80acbf240bf0181f288a74999706684a680e23b20c8e54d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[250, 0x0000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c058000000000000000000000000000000000000000000000000000000000000000, 0x0000000003d493a555e80acbf240bf0181f288a74999706684a680e23b20c8e54d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[249, 0x00000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c000000000000000000000000000000000000000000000000000000000000000, 0x0000000003d493a555e80acbf240bf0181f288a74999706684a680e23b20c8e54d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[248, 0x000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b016000000000000000000000000000000000000000000000000000000000000000, 0x0000000003d493a555e80acbf240bf0181f288a74999706684a680e23b20c8e54d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[247, 0x000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b000000000000000000000000000000000000000000000000000000000000000, 0x0000000001c1c05491a664da33cdd03d7224b47ce157ac2375a43c441a68c3649d42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[246, 0x00000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c05800000000000000000000000000000000000000000000000000000000000000, 0x0000000000b856ac2f8591e1549458db6a3dca67ad36ca01ee2319f50a0cc0a44542ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[245, 0x000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c00000000000000000000000000000000000000000000000000000000000000, 0x000000000033a1d7fe752864e4f79d2a664a555d132658f12a6288cd81debf441942ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[244, 0x0000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b01600000000000000000000000000000000000000000000000000000000000000, 0x000000000033a1d7fe752864e4f79d2a664a555d132658f12a6288cd81debf441942ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[243, 0x0000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b00000000000000000000000000000000000000000000000000000000000000, 0x00000000001274a2f2310e05c9106e3e254d781a6ca23cacf97264839fd33eec0e42ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[242, 0x000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c0580000000000000000000000000000000000000000000000000000000000000, 0x000000000001de086c0f00d63b1cd6c804cf097919602e8ae0fa525eaecd7ec008c2ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[241, 0x0000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c0000000000000000000000000000000000000000000000000000000000000, 0x000000000001de086c0f00d63b1cd6c804cf097919602e8ae0fa525eaecd7ec008c2ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[240, 0x00000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b0160000000000000000000000000000000000000000000000000000000000000, 0x000000000001de086c0f00d63b1cd6c804cf097919602e8ae0fa525eaecd7ec008c2ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[239, 0x00000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b0000000000000000000000000000000000000000000000000000000000000, 0x000000000001de086c0f00d63b1cd6c804cf097919602e8ae0fa525eaecd7ec008c2ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[238, 0x0000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c058000000000000000000000000000000000000000000000000000000000000, 0x000000000000d49ec3ace003423d9d50a2c7228f042c0da8bf72d13c5fbd22bd486aab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[237, 0x00000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c000000000000000000000000000000000000000000000000000000000000, 0x0000000000004fe9ef7bcf99c5ce0094f1c32f19f991fd37aeaf10ab3834f4bbe83eab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[236, 0x000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b016000000000000000000000000000000000000000000000000000000000000, 0x0000000000000d8f85634765079632371941355f7444f4ff264d3062a470ddbb3828ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[235, 0x000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b000000000000000000000000000000000000000000000000000000000000, 0x0000000000000d8f85634765079632371941355f7444f4ff264d3062a470ddbb3828ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[234, 0x00000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c05800000000000000000000000000000000000000000000000000000000000, 0x0000000000000d8f85634765079632371941355f7444f4ff264d3062a470ddbb3828ab9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[233, 0x000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c00000000000000000000000000000000000000000000000000000000000, 0x00000000000005443820365e6fcf386b5e30f628239b53f81540f45991f85adb2225eb9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[232, 0x0000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b01600000000000000000000000000000000000000000000000000000000000, 0x000000000000011e917eaddb23ebbb8580a8d68c7b4683748cbad65508bc196b17248b9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[231, 0x0000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b00000000000000000000000000000000000000000000000000000000000, 0x000000000000011e917eaddb23ebbb8580a8d68c7b4683748cbad65508bc196b17248b9e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[230, 0x000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c0580000000000000000000000000000000000000000000000000000000000, 0x000000000000001527d64bba50f2dc4c0946cea591314f53aa994ed3e66d090f1464339e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[229, 0x0000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c0000000000000000000000000000000000000000000000000000000000, 0x000000000000001527d64bba50f2dc4c0946cea591314f53aa994ed3e66d090f1464339e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[228, 0x00000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b0160000000000000000000000000000000000000000000000000000000000, 0x000000000000001527d64bba50f2dc4c0946cea591314f53aa994ed3e66d090f1464339e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[227, 0x00000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b0000000000000000000000000000000000000000000000000000000000, 0x000000000000001527d64bba50f2dc4c0946cea591314f53aa994ed3e66d090f1464339e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[226, 0x0000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c058000000000000000000000000000000000000000000000000000000000, 0x0000000000000004913bc59843c34e5871d0ae27228ffc119c77365bd448180954382e1e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[225, 0x00000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c000000000000000000000000000000000000000000000000000000000, 0x0000000000000004913bc59843c34e5871d0ae27228ffc119c77365bd448180954382e1e5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[224, 0x000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b016000000000000000000000000000000000000000000000000000000000, 0x00000000000000006b95240fc0776adb8bf3260786e7a74118eeb03dcfbedbc7e42d2cbe5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[223, 0x000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b000000000000000000000000000000000000000000000000000000000, 0x00000000000000006b95240fc0776adb8bf3260786e7a74118eeb03dcfbedbc7e42d2cbe5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[222, 0x00000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c05800000000000000000000000000000000000000000000000000000000, 0x00000000000000006b95240fc0776adb8bf3260786e7a74118eeb03dcfbedbc7e42d2cbe5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[221, 0x000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c00000000000000000000000000000000000000000000000000000000, 0x00000000000000006b95240fc0776adb8bf3260786e7a74118eeb03dcfbedbc7e42d2cbe5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[220, 0x0000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b01600000000000000000000000000000000000000000000000000000000, 0x0000000000000000293ab9f73842aca3bd954d858d2d21f410b627dbef764803cd2c7ca85ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[219, 0x0000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b00000000000000000000000000000000000000000000000000000000, 0x0000000000000000080d84eaf4284d87d6666144904fdf4d8c99e3aaff51fe21c1ac249d5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[218, 0x000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c0580000000000000000000000000000000000000000000000000000000, 0x0000000000000000080d84eaf4284d87d6666144904fdf4d8c99e3aaff51fe21c1ac249d5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[217, 0x0000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c0000000000000000000000000000000000000000000000000000000, 0x0000000000000000080d84eaf4284d87d6666144904fdf4d8c99e3aaff51fe21c1ac249d5ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[216, 0x00000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b0160000000000000000000000000000000000000000000000000000000, 0x000000000000000003e7de496ba501a4598083bc70b436f8bc165b24e14d74e5803c199bfed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[215, 0x00000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b0000000000000000000000000000000000000000000000000000000, 0x000000000000000001d50af8a7635bb29b0d94f860e662ce53d496e1d24b30475f84141b4ed584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[214, 0x0000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c058000000000000000000000000000000000000000000000000000000, 0x000000000000000000cba150454288b9bbd41d9658ff78b91fb3b4c04aca0df84f28115af6d584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[213, 0x00000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c000000000000000000000000000000000000000000000000000000, 0x00000000000000000046ec7c14321f3d4c3761e5550c03ae85a343af87097cd0c6fa0ffacad584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[212, 0x000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b016000000000000000000000000000000000000000000000000000000, 0x000000000000000000049211fba9ea7f1469040cd3124929389b0b272529343d02e30f4ab4d584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[211, 0x000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b000000000000000000000000000000000000000000000000000000, 0x000000000000000000049211fba9ea7f1469040cd3124929389b0b272529343d02e30f4ab4d584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[210, 0x00000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c05800000000000000000000000000000000000000000000000000000, 0x000000000000000000049211fba9ea7f1469040cd3124929389b0b272529343d02e30f4ab4d584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[209, 0x000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c00000000000000000000000000000000000000000000000000000, 0x000000000000000000049211fba9ea7f1469040cd3124929389b0b272529343d02e30f4ab4d584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[208, 0x0000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b01600000000000000000000000000000000000000000000000000000, 0x000000000000000000006c6b5a21673330ec1e2f4af2ad80e3ca879e9f0b2fb3c6a19f3fb37584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[207, 0x0000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b00000000000000000000000000000000000000000000000000000, 0x000000000000000000006c6b5a21673330ec1e2f4af2ad80e3ca879e9f0b2fb3c6a19f3fb37584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[206, 0x000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c0580000000000000000000000000000000000000000000000000000, 0x000000000000000000006c6b5a21673330ec1e2f4af2ad80e3ca879e9f0b2fb3c6a19f3fb37584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[205, 0x0000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c0000000000000000000000000000000000000000000000000000, 0x000000000000000000006c6b5a21673330ec1e2f4af2ad80e3ca879e9f0b2fb3c6a19f3fb37584a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[204, 0x00000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b0160000000000000000000000000000000000000000000000000000, 0x000000000000000000002a10f008defe72b44fd17270b3c65e7d7f6616a94f6b32dd883f035f84a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[203, 0x00000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b0000000000000000000000000000000000000000000000000000, 0x0000000000000000000008e3bafc9ae4139868a2862fb6e91bd6fb49d2785f46e8fb7cbeab5484a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[202, 0x0000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c058000000000000000000000000000000000000000000000000000, 0x0000000000000000000008e3bafc9ae4139868a2862fb6e91bd6fb49d2785f46e8fb7cbeab5484a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[201, 0x00000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c000000000000000000000000000000000000000000000000000, 0x0000000000000000000000986db989dd7bd16ed6cb1f77b1cb2d5a42c16c233dd682f9de9551c4a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[200, 0x000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b016000000000000000000000000000000000000000000000000000, 0x0000000000000000000000986db989dd7bd16ed6cb1f77b1cb2d5a42c16c233dd682f9de9551c4a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[199, 0x000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b000000000000000000000000000000000000000000000000000, 0x0000000000000000000000986db989dd7bd16ed6cb1f77b1cb2d5a42c16c233dd682f9de9551c4a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[198, 0x00000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c05800000000000000000000000000000000000000000000000000, 0x0000000000000000000000986db989dd7bd16ed6cb1f77b1cb2d5a42c16c233dd682f9de9551c4a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[197, 0x000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c00000000000000000000000000000000000000000000000000, 0x000000000000000000000013b8e558cd1254ff3a0f6e73be5622c032505b5f7d455b71b093f198a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[196, 0x0000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b01600000000000000000000000000000000000000000000000000, 0x000000000000000000000013b8e558cd1254ff3a0f6e73be5622c032505b5f7d455b71b093f198a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[195, 0x0000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b00000000000000000000000000000000000000000000000000, 0x000000000000000000000013b8e558cd1254ff3a0f6e73be5622c032505b5f7d455b71b093f198a621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[194, 0x000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c0580000000000000000000000000000000000000000000000000, 0x000000000000000000000003224ad2ab0525714677f8533fe7816cf042394705333680aad3c5932621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[193, 0x0000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c0000000000000000000000000000000000000000000000000, 0x000000000000000000000003224ad2ab0525714677f8533fe7816cf042394705333680aad3c5932621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[192, 0x00000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b0160000000000000000000000000000000000000000000000000, 0x000000000000000000000003224ad2ab0525714677f8533fe7816cf042394705333680aad3c5932621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[191, 0x00000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b0000000000000000000000000000000000000000000000000, 0x0000000000000000000000010f7781e6c37f7f8805098f3019ad4288007503f630f1e28a1bc0127621fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[190, 0x0000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c058000000000000000000000000000000000000000000000000, 0x000000000000000000000000060dd984a2ac86a8cb922d2832c32d53df92e26eafcf9379bfbd521e21fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[189, 0x00000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c000000000000000000000000000000000000000000000000, 0x000000000000000000000000060dd984a2ac86a8cb922d2832c32d53df92e26eafcf9379bfbd521e21fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[188, 0x000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b016000000000000000000000000000000000000000000000000, 0x000000000000000000000000060dd984a2ac86a8cb922d2832c32d53df92e26eafcf9379bfbd521e21fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[187, 0x000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b000000000000000000000000000000000000000000000000, 0x000000000000000000000000060dd984a2ac86a8cb922d2832c32d53df92e26eafcf9379bfbd521e21fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[186, 0x00000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c05800000000000000000000000000000000000000000000000, 0x000000000000000000000000060dd984a2ac86a8cb922d2832c32d53df92e26eafcf9379bfbd521e21fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[185, 0x000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c00000000000000000000000000000000000000000000000, 0x000000000000000000000000060dd984a2ac86a8cb922d2832c32d53df92e26eafcf9379bfbd521e21fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[184, 0x0000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b01600000000000000000000000000000000000000000000000, 0x00000000000000000000000001e832e31a293ac54eac4fa0132784ff0f0f59e891cb0a3d7e4d471cc1fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[183, 0x0000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b00000000000000000000000000000000000000000000000, 0x00000000000000000000000001e832e31a293ac54eac4fa0132784ff0f0f59e891cb0a3d7e4d471cc1fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[182, 0x000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c0580000000000000000000000000000000000000000000000, 0x00000000000000000000000000dec93ab80867cc6f72d83e0b409ae9daee77c70a49e7ee6df1445c69fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[181, 0x0000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c0000000000000000000000000000000000000000000000, 0x000000000000000000000000005a146686f7fe4fffd61c8d074d25df40de06b6468956c6e5c342fc3dfc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[180, 0x00000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b0160000000000000000000000000000000000000000000000, 0x0000000000000000000000000017b9fc6e6fc991c807beb485536b59f3d5ce2de4a90e3321ac424c27fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[179, 0x00000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b0000000000000000000000000000000000000000000000, 0x0000000000000000000000000017b9fc6e6fc991c807beb485536b59f3d5ce2de4a90e3321ac424c27fc66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[178, 0x0000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c058000000000000000000000000000000000000000000000, 0x00000000000000000000000000072361e84dbc623a14273e64d4fcb8a093c00bcc30fc0e30a68220227c66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[177, 0x00000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c000000000000000000000000000000000000000000000, 0x00000000000000000000000000072361e84dbc623a14273e64d4fcb8a093c00bcc30fc0e30a68220227c66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[176, 0x000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b016000000000000000000000000000000000000000000000, 0x0000000000000000000000000002fdbb46c5391656974160dcb561104bc33c834612f784f4651215211c66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[175, 0x000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b000000000000000000000000000000000000000000000, 0x0000000000000000000000000000eae7f600f77064d8ce7218a5933c215afabf0303f54056445a0fa06c66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[174, 0x00000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c05800000000000000000000000000000000000000000000, 0x0000000000000000000000000000eae7f600f77064d8ce7218a5933c215afabf0303f54056445a0fa06c66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[173, 0x000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c00000000000000000000000000000000000000000000, 0x0000000000000000000000000000663321cfe706e86931b667a19fc716c0ea4df24034af2ebc2c0e404066231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[172, 0x0000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b01600000000000000000000000000000000000000000000, 0x000000000000000000000000000023d8b7b75ed22a3163588f1fa60c9173e21569de54669af8150d902a66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[171, 0x0000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b00000000000000000000000000000000000000000000, 0x000000000000000000000000000002ab82ab1ab7cb157c29a2dea92f4ecd5df925ad64425116098d381f66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[170, 0x000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c0580000000000000000000000000000000000000000000, 0x000000000000000000000000000002ab82ab1ab7cb157c29a2dea92f4ecd5df925ad64425116098d381f66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[169, 0x0000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c0000000000000000000000000000000000000000000, 0x000000000000000000000000000002ab82ab1ab7cb157c29a2dea92f4ecd5df925ad64425116098d381f66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[168, 0x00000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b0160000000000000000000000000000000000000000000, 0x000000000000000000000000000002ab82ab1ab7cb157c29a2dea92f4ecd5df925ad64425116098d381f66231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[167, 0x00000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b0000000000000000000000000000000000000000000, 0x00000000000000000000000000000098af5a56762523bdb6b41a99617aa2f5b7616a55400c77e8d5329eb6231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[166, 0x0000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c058000000000000000000000000000000000000000000, 0x00000000000000000000000000000098af5a56762523bdb6b41a99617aa2f5b7616a55400c77e8d5329eb6231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[165, 0x00000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c000000000000000000000000000000000000000000, 0x00000000000000000000000000000013fa862565bba74e19f869956e05985ba6f059917f7b5060a7313e8a231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[164, 0x000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b016000000000000000000000000000000000000000000, 0x00000000000000000000000000000013fa862565bba74e19f869956e05985ba6f059917f7b5060a7313e8a231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[163, 0x000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b000000000000000000000000000000000000000000, 0x00000000000000000000000000000013fa862565bba74e19f869956e05985ba6f059917f7b5060a7313e8a231d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[162, 0x00000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c05800000000000000000000000000000000000000000, 0x0000000000000000000000000000000363eb9f43ae77c02660f374ef96f70864e2377907692b6fa1711284a31d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[161, 0x000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c00000000000000000000000000000000000000000, 0x0000000000000000000000000000000363eb9f43ae77c02660f374ef96f70864e2377907692b6fa1711284a31d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[160, 0x0000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b01600000000000000000000000000000000000000000, 0x0000000000000000000000000000000363eb9f43ae77c02660f374ef96f70864e2377907692b6fa1711284a31d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[159, 0x0000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b00000000000000000000000000000000000000000, 0x0000000000000000000000000000000151184e7f6cd1ce67ee04b0dfc922ddfca07335f866e6d180b90d03f31d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[158, 0x000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c0580000000000000000000000000000000000000000, 0x0000000000000000000000000000000047aea61d4bfed588b48d4ed7e238c8c87f911470e5c482705d0a439b1d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[157, 0x0000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c0000000000000000000000000000000000000000, 0x0000000000000000000000000000000047aea61d4bfed588b48d4ed7e238c8c87f911470e5c482705d0a439b1d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[156, 0x00000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b0160000000000000000000000000000000000000000, 0x0000000000000000000000000000000005543c04c3ca1750e62f7655e87e437b77588c0f057beeac460993851d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[155, 0x00000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b0000000000000000000000000000000000000000, 0x0000000000000000000000000000000005543c04c3ca1750e62f7655e87e437b77588c0f057beeac460993851d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[154, 0x0000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c058000000000000000000000000000000000000000, 0x0000000000000000000000000000000005543c04c3ca1750e62f7655e87e437b77588c0f057beeac460993851d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[153, 0x00000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c000000000000000000000000000000000000000, 0x0000000000000000000000000000000005543c04c3ca1750e62f7655e87e437b77588c0f057beeac460993851d5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[152, 0x000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b016000000000000000000000000000000000000000, 0x00000000000000000000000000000000012e95633b46cb6d694998cdc8e29b26a6d50388e777657004998883bd5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[151, 0x000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b000000000000000000000000000000000000000, 0x00000000000000000000000000000000012e95633b46cb6d694998cdc8e29b26a6d50388e777657004998883bd5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[150, 0x00000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c05800000000000000000000000000000000000000, 0x0000000000000000000000000000000000252bbad925f8748a10216bc0fbb11172b421675ff64320f43d85c3655e4bbdd963f69888f6d51cc9fc71b8164801f2],
[149, 0x000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c00000000000000000000000000000000000000, 0x0000000000000000000000000000000000252bbad925f8748a10216bc0fbb11172b421675ff64320f43d85c3655e4bbdd963f69888f6d51cc9fc71b8164801f2],
[148, 0x0000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b01600000000000000000000000000000000000000, 0x0000000000000000000000000000000000252bbad925f8748a10216bc0fbb11172b421675ff64320f43d85c3655e4bbdd963f69888f6d51cc9fc71b8164801f2],
[147, 0x0000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b00000000000000000000000000000000000000, 0x000000000000000000000000000000000003fe85cce1de156e28f27f7ffed3cecc3005232f061ed71232056b5a5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[146, 0x000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c0580000000000000000000000000000000000000, 0x000000000000000000000000000000000003fe85cce1de156e28f27f7ffed3cecc3005232f061ed71232056b5a5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[145, 0x0000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c0000000000000000000000000000000000000, 0x000000000000000000000000000000000003fe85cce1de156e28f27f7ffed3cecc3005232f061ed71232056b5a5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[144, 0x00000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b0160000000000000000000000000000000000000, 0x000000000000000000000000000000000003fe85cce1de156e28f27f7ffed3cecc3005232f061ed71232056b5a5e4bbdd963f69888f6d51cc9fc71b8164801f2],
[143, 0x00000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b0000000000000000000000000000000000000, 0x000000000000000000000000000000000001ebb27c1d9c6f7c6a7f90bbef05faa1c7c35eebf71c9274114d65d9ae4bbdd963f69888f6d51cc9fc71b8164801f2],
[142, 0x0000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c058000000000000000000000000000000000000, 0x000000000000000000000000000000000000e248d3bb7b9c838b461959e71f108c93a27cca6f9b702500f16319564bbdd963f69888f6d51cc9fc71b8164801f2],
[141, 0x00000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c000000000000000000000000000000000000, 0x0000000000000000000000000000000000005d93ff8a6b33071ba95da8e32b9b81f9920bb9abdadefd78c361b92a4bbdd963f69888f6d51cc9fc71b8164801f2],
[140, 0x000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b016000000000000000000000000000000000000, 0x0000000000000000000000000000000000001b399571e2fe48e3daffd06131e0fcac89d33149fa9669b4ac6109144bbdd963f69888f6d51cc9fc71b8164801f2],
[139, 0x000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b000000000000000000000000000000000000, 0x0000000000000000000000000000000000001b399571e2fe48e3daffd06131e0fcac89d33149fa9669b4ac6109144bbdd963f69888f6d51cc9fc71b8164801f2],
[138, 0x00000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c05800000000000000000000000000000000000, 0x0000000000000000000000000000000000000aa2faebc0f11955e7685a40b3725b5947c50f31828444c3a6a0dd0ecbbdd963f69888f6d51cc9fc71b8164801f2],
[137, 0x000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c00000000000000000000000000000000000, 0x0000000000000000000000000000000000000257ada8afea818eed9c9f30743b0aafa6bdfe25467b324b23c0c70c0bbdd963f69888f6d51cc9fc71b8164801f2],
[136, 0x0000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b01600000000000000000000000000000000000, 0x0000000000000000000000000000000000000257ada8afea818eed9c9f30743b0aafa6bdfe25467b324b23c0c70c0bbdd963f69888f6d51cc9fc71b8164801f2],
[135, 0x0000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b00000000000000000000000000000000000, 0x0000000000000000000000000000000000000044da57eba8db9d2f29b06c646d36853e7c39e23778edad0308c18b5bbdd963f69888f6d51cc9fc71b8164801f2],
[134, 0x000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c0580000000000000000000000000000000000, 0x0000000000000000000000000000000000000044da57eba8db9d2f29b06c646d36853e7c39e23778edad0308c18b5bbdd963f69888f6d51cc9fc71b8164801f2],
[133, 0x0000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c0000000000000000000000000000000000, 0x0000000000000000000000000000000000000044da57eba8db9d2f29b06c646d36853e7c39e23778edad0308c18b5bbdd963f69888f6d51cc9fc71b8164801f2],
[132, 0x00000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b0160000000000000000000000000000000000, 0x00000000000000000000000000000000000000027fedd320a6def75b5293e2737bfff1740159d598a5193ef1c0db45bdd963f69888f6d51cc9fc71b8164801f2],
[131, 0x00000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b0000000000000000000000000000000000, 0x00000000000000000000000000000000000000027fedd320a6def75b5293e2737bfff1740159d598a5193ef1c0db45bdd963f69888f6d51cc9fc71b8164801f2],
[130, 0x0000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c058000000000000000000000000000000000, 0x00000000000000000000000000000000000000027fedd320a6def75b5293e2737bfff1740159d598a5193ef1c0db45bdd963f69888f6d51cc9fc71b8164801f2],
[129, 0x00000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c000000000000000000000000000000000, 0x00000000000000000000000000000000000000027fedd320a6def75b5293e2737bfff1740159d598a5193ef1c0db45bdd963f69888f6d51cc9fc71b8164801f2],
[128, 0x000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b016000000000000000000000000000000000, 0x00000000000000000000000000000000000000027fedd320a6def75b5293e2737bfff1740159d598a5193ef1c0db45bdd963f69888f6d51cc9fc71b8164801f2],
[127, 0x000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b000000000000000000000000000000000, 0x00000000000000000000000000000000000000006d1a825c6539059cdfa51e63ae2bc70bbf959289a2d4a0d108d5c50dd963f69888f6d51cc9fc71b8164801f2],
[126, 0x00000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c05800000000000000000000000000000000, 0x00000000000000000000000000000000000000006d1a825c6539059cdfa51e63ae2bc70bbf959289a2d4a0d108d5c50dd963f69888f6d51cc9fc71b8164801f2],
[125, 0x000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c00000000000000000000000000000000, 0x00000000000000000000000000000000000000006d1a825c6539059cdfa51e63ae2bc70bbf959289a2d4a0d108d5c50dd963f69888f6d51cc9fc71b8164801f2],
[124, 0x0000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b01600000000000000000000000000000000, 0x00000000000000000000000000000000000000002ac01843dd044765114745e1b47141beb75d0a27c28c0d0cf1d514f7d963f69888f6d51cc9fc71b8164801f2],
[123, 0x0000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b00000000000000000000000000000000, 0x00000000000000000000000000000000000000000992e33798e9e8492a1859a0b793ff183340c5f6d267c32ae654bcecd963f69888f6d51cc9fc71b8164801f2],
[122, 0x000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c0580000000000000000000000000000000, 0x00000000000000000000000000000000000000000992e33798e9e8492a1859a0b793ff183340c5f6d267c32ae654bcecd963f69888f6d51cc9fc71b8164801f2],
[121, 0x0000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c0000000000000000000000000000000, 0x0000000000000000000000000000000000000000014795f487e35082304c9e90785cae6e9239b4ea965eb0b26374a6ea1963f69888f6d51cc9fc71b8164801f2],
[120, 0x00000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b0160000000000000000000000000000000, 0x0000000000000000000000000000000000000000014795f487e35082304c9e90785cae6e9239b4ea965eb0b26374a6ea1963f69888f6d51cc9fc71b8164801f2],
[119, 0x00000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b0000000000000000000000000000000, 0x0000000000000000000000000000000000000000014795f487e35082304c9e90785cae6e9239b4ea965eb0b26374a6ea1963f69888f6d51cc9fc71b8164801f2],
[118, 0x0000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c058000000000000000000000000000000, 0x0000000000000000000000000000000000000000003e2c4c25c27d895113272e7075c4595e18d2c90edd8e635318a429c163f69888f6d51cc9fc71b8164801f2],
[117, 0x00000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c000000000000000000000000000000, 0x0000000000000000000000000000000000000000003e2c4c25c27d895113272e7075c4595e18d2c90edd8e635318a429c163f69888f6d51cc9fc71b8164801f2],
[116, 0x000000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b016000000000000000000000000000000, 0x0000000000000000000000000000000000000000003e2c4c25c27d895113272e7075c4595e18d2c90edd8e635318a429c163f69888f6d51cc9fc71b8164801f2],
[115, 0x000000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b000000000000000000000000000000, 0x0000000000000000000000000000000000000000001cff17197e632a352bf8422f78e716b794b684dded6a19710d23d1b663f69888f6d51cc9fc71b8164801f2],
[114, 0x00000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c05800000000000000000000000000000, 0x0000000000000000000000000000000000000000000c687c935c55faa73860cc0efa78756452a862c57557f4800763a5b0e3f69888f6d51cc9fc71b8164801f2],
[113, 0x000000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c00000000000000000000000000000, 0x000000000000000000000000000000000000000000041d2f504b4f62e03e9510febb4124bab1a151b9394ee20784838fae23f69888f6d51cc9fc71b8164801f2],
[112, 0x0000000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b01600000000000000000000000000000, 0x000000000000000000000000000000000000000000041d2f504b4f62e03e9510febb4124bab1a151b9394ee20784838fae23f69888f6d51cc9fc71b8164801f2],
[111, 0x0000000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b00000000000000000000000000000, 0x000000000000000000000000000000000000000000020a5bff870dbcee8022223aab735090495f8d762a4c9d6963cb8a2d73f69888f6d51cc9fc71b8164801f2],
[110, 0x000000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c0580000000000000000000000000000, 0x0000000000000000000000000000000000000000000100f25724ece9f5a0e8aad8a38c667b153eab54a2cb7b1a536f876d1bf69888f6d51cc9fc71b8164801f2],
[109, 0x0000000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c0000000000000000000000000000, 0x000000000000000000000000000000000000000000007c3d82f3dc8079314bef279f98f1707b2e3a43df0ae9f2cb41860ceff69888f6d51cc9fc71b8164801f2],
[108, 0x00000000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b0160000000000000000000000000000, 0x0000000000000000000000000000000000000000000039e318db544bbaf97d914f1d9f36eb2e2601bb7d2aa15f072a855cd9f69888f6d51cc9fc71b8164801f2],
[107, 0x00000000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b0000000000000000000000000000, 0x0000000000000000000000000000000000000000000018b5e3cf10315bdd966262dca259a887a1e5774c3a7d15251f0504cef69888f6d51cc9fc71b8164801f2],
[106, 0x0000000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c058000000000000000000000000000, 0x00000000000000000000000000000000000000000000081f4948ee242c4fa2caecbc23eb07345fd75533c26af0341944d8c9769888f6d51cc9fc71b8164801f2],
[105, 0x00000000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c000000000000000000000000000, 0x00000000000000000000000000000000000000000000081f4948ee242c4fa2caecbc23eb07345fd75533c26af0341944d8c9769888f6d51cc9fc71b8164801f2],
[104, 0x000000000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b016000000000000000000000000000, 0x0000000000000000000000000000000000000000000003f9a2a765a0e06c25e50f34044f5edf8f53ccada46666f7d7d4cdc8169888f6d51cc9fc71b8164801f2],
[103, 0x000000000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b000000000000000000000000000, 0x0000000000000000000000000000000000000000000001e6cf56a15f3a7a6772206ff4818ab52712086a95642259b71cc847669888f6d51cc9fc71b8164801f2],
[102, 0x00000000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c05800000000000000000000000000, 0x0000000000000000000000000000000000000000000000dd65ae3f3e67818838a90dec9aa09ff2f126490de3000aa6c0c5870e9888f6d51cc9fc71b8164801f2],
[101, 0x000000000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c00000000000000000000000000, 0x000000000000000000000000000000000000000000000058b0da0e2dfe05189bed5ce8a72b9558e0b5384a226ee31e92c426e29888f6d51cc9fc71b8164801f2],
[100, 0x0000000000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b01600000000000000000000000000, 0x000000000000000000000000000000000000000000000016566ff5a5c946e0cd8f8466ad71100bd87cafe842264f5a7bc376cc9888f6d51cc9fc71b8164801f2],
[99, 0x0000000000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b00000000000000000000000000, 0x000000000000000000000000000000000000000000000016566ff5a5c946e0cd8f8466ad71100bd87cafe842264f5a7bc376cc9888f6d51cc9fc71b8164801f2],
[98, 0x000000000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c0580000000000000000000000000, 0x000000000000000000000000000000000000000000000005bfd56f83bc1752d9f80e462f026eb8966e8dcfca142a6976034ac71888f6d51cc9fc71b8164801f2],
[97, 0x0000000000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c0000000000000000000000000, 0x000000000000000000000000000000000000000000000005bfd56f83bc1752d9f80e462f026eb8966e8dcfca142a6976034ac71888f6d51cc9fc71b8164801f2],
[96, 0x00000000000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b0160000000000000000000000000, 0x0000000000000000000000000000000000000000000000019a2ecdfb38cb6f5d1230be0f66c663c5eb0549ac0fa12d34933fc5b888f6d51cc9fc71b8164801f2],
[95, 0x00000000000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b0000000000000000000000000, 0x0000000000000000000000000000000000000000000000019a2ecdfb38cb6f5d1230be0f66c663c5eb0549ac0fa12d34933fc5b888f6d51cc9fc71b8164801f2],
[94, 0x0000000000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c058000000000000000000000000, 0x00000000000000000000000000000000000000000000000090c5259917f8767dd8b95c077fdc4e91ca2328248e7ede24373d056088f6d51cc9fc71b8164801f2],
[93, 0x00000000000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c000000000000000000000000, 0x0000000000000000000000000000000000000000000000000c105168078efa0e3bfdab038c6743f7b9b21760cdedb69c093ba53488f6d51cc9fc71b8164801f2],
[92, 0x000000000000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b016000000000000000000000000, 0x0000000000000000000000000000000000000000000000000c105168078efa0e3bfdab038c6743f7b9b21760cdedb69c093ba53488f6d51cc9fc71b8164801f2],
[91, 0x000000000000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b000000000000000000000000, 0x0000000000000000000000000000000000000000000000000c105168078efa0e3bfdab038c6743f7b9b21760cdedb69c093ba53488f6d51cc9fc71b8164801f2],
[90, 0x00000000000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c05800000000000000000000000, 0x0000000000000000000000000000000000000000000000000c105168078efa0e3bfdab038c6743f7b9b21760cdedb69c093ba53488f6d51cc9fc71b8164801f2],
[89, 0x000000000000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c00000000000000000000000, 0x00000000000000000000000000000000000000000000000003c50424f68862474231eff34d2ff34e18ab065491e4a423865b8f31c8f6d51cc9fc71b8164801f2],
[88, 0x0000000000000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b01600000000000000000000000, 0x00000000000000000000000000000000000000000000000003c50424f68862474231eff34d2ff34e18ab065491e4a423865b8f31c8f6d51cc9fc71b8164801f2],
[87, 0x0000000000000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b00000000000000000000000, 0x00000000000000000000000000000000000000000000000001b230d43246bc5583bf012f3d621f23b069421182e25f8565a389b118f6d51cc9fc71b8164801f2],
[86, 0x000000000000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c0580000000000000000000000, 0x00000000000000000000000000000000000000000000000000a8c72bd025e95ca48589cd357b350e7c485feffb613d36554786f0c0f6d51cc9fc71b8164801f2],
[85, 0x0000000000000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c0000000000000000000000, 0x000000000000000000000000000000000000000000000000002412579f157fe034e8ce1c3187c003e237eedf37a0ac0ecd19859094f6d51cc9fc71b8164801f2],
[84, 0x00000000000000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b0160000000000000000000000, 0x000000000000000000000000000000000000000000000000002412579f157fe034e8ce1c3187c003e237eedf37a0ac0ecd19859094f6d51cc9fc71b8164801f2],
[83, 0x00000000000000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b0000000000000000000000, 0x0000000000000000000000000000000000000000000000000002e52292d1658119019f2ff08ae2c13bb3d29b06b087c4eb0e053889f6d51cc9fc71b8164801f2],
[82, 0x0000000000000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c058000000000000000000000, 0x0000000000000000000000000000000000000000000000000002e52292d1658119019f2ff08ae2c13bb3d29b06b087c4eb0e053889f6d51cc9fc71b8164801f2],
[81, 0x00000000000000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c000000000000000000000, 0x0000000000000000000000000000000000000000000000000002e52292d1658119019f2ff08ae2c13bb3d29b06b087c4eb0e053889f6d51cc9fc71b8164801f2],
[80, 0x000000000000000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b016000000000000000000000, 0x0000000000000000000000000000000000000000000000000002e52292d1658119019f2ff08ae2c13bb3d29b06b087c4eb0e053889f6d51cc9fc71b8164801f2],
[79, 0x000000000000000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b000000000000000000000, 0x0000000000000000000000000000000000000000000000000000d24f420d23db27432c412c7b14ed114b90d6c3a185804ced4d330946d51cc9fc71b8164801f2],
[78, 0x00000000000000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c05800000000000000000000, 0x0000000000000000000000000000000000000000000000000000d24f420d23db27432c412c7b14ed114b90d6c3a185804ced4d330946d51cc9fc71b8164801f2],
[77, 0x000000000000000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c00000000000000000000, 0x00000000000000000000000000000000000000000000000000004d9a6ddc1371aad38f857b77217806b18065b2ddc4ef25651f31a91ad51cc9fc71b8164801f2],
[76, 0x0000000000000000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b01600000000000000000000, 0x00000000000000000000000000000000000000000000000000000b4003c38b3cec9bc127a2f527bd8164782d2a7be4a691a10830f904d51cc9fc71b8164801f2],
[75, 0x0000000000000000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b00000000000000000000, 0x00000000000000000000000000000000000000000000000000000b4003c38b3cec9bc127a2f527bd8164782d2a7be4a691a10830f904d51cc9fc71b8164801f2],
[74, 0x000000000000000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c0580000000000000000000, 0x00000000000000000000000000000000000000000000000000000b4003c38b3cec9bc127a2f527bd8164782d2a7be4a691a10830f904d51cc9fc71b8164801f2],
[73, 0x0000000000000000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c0000000000000000000, 0x000000000000000000000000000000000000000000000000000002f4b6807a3654d4c75be7e4e88630bad726196fa89d7f288550e302151cc9fc71b8164801f2],
[72, 0x00000000000000000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b0160000000000000000000, 0x000000000000000000000000000000000000000000000000000002f4b6807a3654d4c75be7e4e88630bad726196fa89d7f288550e302151cc9fc71b8164801f2],
[71, 0x00000000000000000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b0000000000000000000, 0x000000000000000000000000000000000000000000000000000000e1e32fb5f4aee308e8f920d8b85c906ee4552c999b3a8a6498dd81651cc9fc71b8164801f2],
[70, 0x0000000000000000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c058000000000000000000, 0x000000000000000000000000000000000000000000000000000000e1e32fb5f4aee308e8f920d8b85c906ee4552c999b3a8a6498dd81651cc9fc71b8164801f2],
[69, 0x00000000000000000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c000000000000000000, 0x0000000000000000000000000000000000000000000000000000005d2e5b84e44566994c3d6fd4c4e785d4d3e41bd5daa962dc6adc21391cc9fc71b8164801f2],
[68, 0x000000000000000000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b016000000000000000000, 0x0000000000000000000000000000000000000000000000000000001ad3f16c5c10a8617ddf9752cb2d0087cbab9373fa60cf1853db71231cc9fc71b8164801f2],
[67, 0x000000000000000000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b000000000000000000, 0x0000000000000000000000000000000000000000000000000000001ad3f16c5c10a8617ddf9752cb2d0087cbab9373fa60cf1853db71231cc9fc71b8164801f2],
[66, 0x00000000000000000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c05800000000000000000, 0x0000000000000000000000000000000000000000000000000000000a3d56e63a0378d38a4821324cbe5f34899d715b824eaa274e1b451d9cc9fc71b8164801f2],
[65, 0x000000000000000000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c00000000000000000, 0x00000000000000000000000000000000000000000000000000000001f209a328fce10c907c66220d870e8ae896604f464597aecb3b2f1adcc9fc71b8164801f2],
[64, 0x0000000000000000000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b01600000000000000000, 0x00000000000000000000000000000000000000000000000000000001f209a328fce10c907c66220d870e8ae896604f464597aecb3b2f1adcc9fc71b8164801f2],
[63, 0x0000000000000000000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b00000000000000000, 0x00000000000000000000000000000000000000000000000000000001f209a328fce10c907c66220d870e8ae896604f464597aecb3b2f1adcc9fc71b8164801f2],
[62, 0x000000000000000000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c0580000000000000000, 0x00000000000000000000000000000000000000000000000000000000e89ffac6dc0e13b142eec005a02475b4757e2dbec4755fbadf2c5a84c9fc71b8164801f2],
[61, 0x0000000000000000000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c0000000000000000, 0x0000000000000000000000000000000000000000000000000000000063eb2695cba49741a6330f01acaf6b1a650d1cfb03e43832b12afa58c9fc71b8164801f2],
[60, 0x00000000000000000000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b0160000000000000000, 0x000000000000000000000000000000000000000000000000000000002190bc7d436fd909d7d5367fb2f4e5cd5cd49499239ba46e9a2a4a42c9fc71b8164801f2],
[59, 0x00000000000000000000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b0000000000000000, 0x0000000000000000000000000000000000000000000000000000000000638770ff5579edf0a64a3eb617a326d8b8506833775a8c8ea9f237c9fc71b8164801f2],
[58, 0x0000000000000000000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c058000000000000000, 0x0000000000000000000000000000000000000000000000000000000000638770ff5579edf0a64a3eb617a326d8b8506833775a8c8ea9f237c9fc71b8164801f2],
[57, 0x00000000000000000000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c000000000000000, 0x0000000000000000000000000000000000000000000000000000000000638770ff5579edf0a64a3eb617a326d8b8506833775a8c8ea9f237c9fc71b8164801f2],
[56, 0x000000000000000000000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b016000000000000000, 0x0000000000000000000000000000000000000000000000000000000000638770ff5579edf0a64a3eb617a326d8b8506833775a8c8ea9f237c9fc71b8164801f2],
[55, 0x000000000000000000000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b000000000000000, 0x0000000000000000000000000000000000000000000000000000000000638770ff5579edf0a64a3eb617a326d8b8506833775a8c8ea9f237c9fc71b8164801f2],
[54, 0x00000000000000000000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c05800000000000000, 0x0000000000000000000000000000000000000000000000000000000000638770ff5579edf0a64a3eb617a326d8b8506833775a8c8ea9f237c9fc71b8164801f2],
[53, 0x000000000000000000000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c00000000000000, 0x0000000000000000000000000000000000000000000000000000000000638770ff5579edf0a64a3eb617a326d8b8506833775a8c8ea9f237c9fc71b8164801f2],
[52, 0x0000000000000000000000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b01600000000000000, 0x0000000000000000000000000000000000000000000000000000000000212d06e6cd452fb8d7ec66341de8a18bb017dfd19711f8ca92f187b3fc71b8164801f2],
[51, 0x0000000000000000000000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b00000000000000, 0x0000000000000000000000000000000000000000000000000000000000212d06e6cd452fb8d7ec66341de8a18bb017dfd19711f8ca92f187b3fc71b8164801f2],
[50, 0x000000000000000000000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c0580000000000000, 0x000000000000000000000000000000000000000000000000000000000010966c60ab38002ae454f0139f7a00386e09bdb91effd3d98d315bae7c71b8164801f2],
[49, 0x0000000000000000000000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c0000000000000, 0x0000000000000000000000000000000000000000000000000000000000084b1f1d9a316863ea8935036042af8ecd02acace2f6c1610a5145abbc71b8164801f2],
[48, 0x00000000000000000000000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b0160000000000000, 0x00000000000000000000000000000000000000000000000000000000000425787c11ae1c806da3577b40a70739fc7f2426c4f23824c8e13aaa5c71b8164801f2],
[47, 0x00000000000000000000000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b0000000000000, 0x00000000000000000000000000000000000000000000000000000000000212a52b4d6c768eaf3068b730d9330f943d5fe3b5eff386a8293529ac71b8164801f2],
[46, 0x0000000000000000000000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c058000000000000, 0x000000000000000000000000000000000000000000000000000000000001093b82eb4ba395cff6f15528f248fa601c7dc22e6ed13797cd32695471b8164801f2],
[45, 0x00000000000000000000000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c000000000000, 0x0000000000000000000000000000000000000000000000000000000000008486aeba3b3a19605a35a424fed3efc60c0cb16aae40100f9f31092871b8164801f2],
[44, 0x000000000000000000000000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b016000000000000, 0x000000000000000000000000000000000000000000000000000000000000422c44a1b3055b288bd7cba305196a7903d42908cdf77c4b8830591271b8164801f2],
[43, 0x000000000000000000000000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b000000000000, 0x00000000000000000000000000000000000000000000000000000000000020ff0f956eeafc0ca4a8df62083c27d27fb7e4d7ddd332697cb0010771b8164801f2],
[42, 0x00000000000000000000000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c05800000000000, 0x0000000000000000000000000000000000000000000000000000000000001068750f4cddcc7eb111694189cd867f3da9c2bf65c10d7876efd501f1b8164801f2],
[41, 0x000000000000000000000000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c00000000000, 0x000000000000000000000000000000000000000000000000000000000000081d27cc3bd734b7b745ae314a9635d59ca2b1b329b7fafff40fbeff31b8164801f2],
[40, 0x0000000000000000000000000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b01600000000000, 0x00000000000000000000000000000000000000000000000000000000000003f7812ab353e8d43a5fd0a92afa8d80cc1f292d0bb371c3b29fb3fdd1b8164801f2],
[39, 0x0000000000000000000000000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b00000000000, 0x00000000000000000000000000000000000000000000000000000000000001e4add9ef1242e27bece1e51b2cb95663dd64e9fcb12d2591e7ae7d21b8164801f2],
[38, 0x000000000000000000000000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c0580000000000, 0x00000000000000000000000000000000000000000000000000000000000000db44318cf16fe99cb36a831345cf412fbc82c875300ad6818babbcc9b8164801f2],
[37, 0x0000000000000000000000000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c0000000000, 0x00000000000000000000000000000000000000000000000000000000000000568f5d5be1066d2d16aed20f525a3695ac11b7b16f79aef95daa5c9db8164801f2],
[36, 0x00000000000000000000000000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b0160000000000, 0x000000000000000000000000000000000000000000000000000000000000001434f34358d1aef54850f98d589fb148a3d92f4f8f311b3546a9ac87b8164801f2],
[35, 0x00000000000000000000000000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b0000000000, 0x000000000000000000000000000000000000000000000000000000000000001434f34358d1aef54850f98d589fb148a3d92f4f8f311b3546a9ac87b8164801f2],
[34, 0x0000000000000000000000000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c058000000000, 0x00000000000000000000000000000000000000000000000000000000000000039e58bd36c47f6754b9836cda310ff561cb0d37171ef64440e9808238164801f2],
[33, 0x00000000000000000000000000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c000000000, 0x00000000000000000000000000000000000000000000000000000000000000039e58bd36c47f6754b9836cda310ff561cb0d37171ef64440e9808238164801f2],
[32, 0x000000000000000000000000000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b016000000000, 0x00000000000000000000000000000000000000000000000000000000000000039e58bd36c47f6754b9836cda310ff561cb0d37171ef64440e9808238164801f2],
[31, 0x000000000000000000000000000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b000000000, 0x00000000000000000000000000000000000000000000000000000000000000018b856c7282d975964694a8ca633bcaf98948f4081cb1a620317b0188164801f2],
[30, 0x00000000000000000000000000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c05800000000, 0x0000000000000000000000000000000000000000000000000000000000000000821bc41062067cb70d1d46c27c51b5c56866d2809b8f570fd5784130164801f2],
[29, 0x000000000000000000000000000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c00000000, 0x0000000000000000000000000000000000000000000000000000000000000000821bc41062067cb70d1d46c27c51b5c56866d2809b8f570fd5784130164801f2],
[28, 0x0000000000000000000000000000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b01600000000, 0x00000000000000000000000000000000000000000000000000000000000000003fc159f7d9d1be7f3ebf6e4082973078602e4a1ebb46c34bbe77911a164801f2],
[27, 0x0000000000000000000000000000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b00000000, 0x00000000000000000000000000000000000000000000000000000000000000001e9424eb95b75f63579081ff85b9edd1dc1205edcb227969b2f7390f164801f2],
[26, 0x000000000000000000000000000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c0580000000, 0x00000000000000000000000000000000000000000000000000000000000000000dfd8a6573aa2fd563f90bdf074b4c7e9a03e3d553105478ad370d09964801f2],
[25, 0x0000000000000000000000000000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c0000000, 0x000000000000000000000000000000000000000000000000000000000000000005b23d2262a3980e6a2d50cec813fbd4f8fcd2c9170742002a56f706d64801f2],
[24, 0x00000000000000000000000000000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b0160000000, 0x0000000000000000000000000000000000000000000000000000000000000000018c9680da204c2aed477346a878538028794a42f902b8c3e8e6ec05764801f2],
[23, 0x00000000000000000000000000000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b0000000, 0x0000000000000000000000000000000000000000000000000000000000000000018c9680da204c2aed477346a878538028794a42f902b8c3e8e6ec05764801f2],
[22, 0x0000000000000000000000000000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c058000000, 0x000000000000000000000000000000000000000000000000000000000000000000832cd877ff79320e0dfbe4a091696af458682171819674d88ae9451e4801f2],
[21, 0x00000000000000000000000000000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c000000, 0x000000000000000000000000000000000000000000000000000000000000000000832cd877ff79320e0dfbe4a091696af458682171819674d88ae9451e4801f2],
[20, 0x000000000000000000000000000000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b016000000, 0x00000000000000000000000000000000000000000000000000000000000000000040d26e5f774473d63f9e0c1e97aee5a7502f990fa14de11473e895084801f2],
[19, 0x000000000000000000000000000000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b000000, 0x0000000000000000000000000000000000000000000000000000000000000000001fa53953332a14ba586f1fdd9ad1a300cc1354deb129973268683cfd4801f2],
[18, 0x00000000000000000000000000000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c05800000, 0x0000000000000000000000000000000000000000000000000000000000000000000f0e9ecd111ce52c64d7a9bd1c6301ad8a0532c63917724162a810f7c801f2],
[17, 0x000000000000000000000000000000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c00000, 0x00000000000000000000000000000000000000000000000000000000000000000006c3518a00164d656b0beeacdd2bb103e8fe21b9fd0e5fc8dfc7faf50801f2],
[16, 0x0000000000000000000000000000000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b01600000, 0x000000000000000000000000000000000000000000000000000000000000000000029daae877930181ee261124bd9008af187a9933df09d68c9e57eff3a801f2],
[15, 0x0000000000000000000000000000000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b00000, 0x000000000000000000000000000000000000000000000000000000000000000000008ad797b3515b902fb32260adc23484b038d4f0d00791ee7d9fea72f801f2],
[14, 0x000000000000000000000000000000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c0580000, 0x000000000000000000000000000000000000000000000000000000000000000000008ad797b3515b902fb32260adc23484b038d4f0d00791ee7d9fea72f801f2],
[13, 0x0000000000000000000000000000000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c0000, 0x000000000000000000000000000000000000000000000000000000000000000000000622c38240f213c01666afa9cebf7a162863e00c4700c6f571e912cc01f2],
[12, 0x00000000000000000000000000000000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b0160000, 0x000000000000000000000000000000000000000000000000000000000000000000000622c38240f213c01666afa9cebf7a162863e00c4700c6f571e912cc01f2],
[11, 0x00000000000000000000000000000000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b0000, 0x000000000000000000000000000000000000000000000000000000000000000000000622c38240f213c01666afa9cebf7a162863e00c4700c6f571e912cc01f2],
[10, 0x0000000000000000000000000000000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c058000, 0x000000000000000000000000000000000000000000000000000000000000000000000622c38240f213c01666afa9cebf7a162863e00c4700c6f571e912cc01f2],
[9, 0x00000000000000000000000000000000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c000, 0x000000000000000000000000000000000000000000000000000000000000000000000622c38240f213c01666afa9cebf7a162863e00c4700c6f571e912cc01f2],
[8, 0x000000000000000000000000000000000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b016000, 0x0000000000000000000000000000000000000000000000000000000000000000000001fd1ce0b86ec7dc9980d221af23d1c157e0578628fc3db9307907caa1f2],
[7, 0x000000000000000000000000000000000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b000, 0x0000000000000000000000000000000000000000000000000000000000000000000001fd1ce0b86ec7dc9980d221af23d1c157e0578628fc3db9307907caa1f2],
[6, 0x00000000000000000000000000000000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c05800, 0x0000000000000000000000000000000000000000000000000000000000000000000000f3b338564df4e3ba475abfa73ce7ac23bf7564a17b1b6a201d050a49f2],
[5, 0x000000000000000000000000000000000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c00, 0x00000000000000000000000000000000000000000000000000000000000000000000006efe64253d8b674aaa9f0ea34972a189af0453ddba8a4297ef03aa1df2],
[4, 0x0000000000000000000000000000000000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b01600, 0x00000000000000000000000000000000000000000000000000000000000000000000002ca3fa0cb556a912dc4136214fb81c3ca6cbcb7bda41aed3d802fa07f2],
[3, 0x0000000000000000000000000000000000000000000000000000000000000000000000212d350c441a5f1be72eec40fcdd42a6841c4430f02449e20b80580b00, 0x00000000000000000000000000000000000000000000000000000000000000000000000b76c500713c49f6f51249e052dad99622af874aea1d64f1cc82a1fcf2],
[2, 0x000000000000000000000000000000000000000000000000000000000000000000000010969a86220d2f8df39776207e6ea153420e2218781224f105c02c0580, 0x00000000000000000000000000000000000000000000000000000000000000000000000b76c500713c49f6f51249e052dad99622af874aea1d64f1cc82a1fcf2],
[1, 0x0000000000000000000000000000000000000000000000000000000000000000000000084b4d43110697c6f9cbbb103f3750a9a107110c3c09127882e01602c0, 0x0000000000000000000000000000000000000000000000000000000000000000000000032b77bd6035b22ffb468ed013a388ec81a8763eae14527949a28bfa32],
[0, 0x00000000000000000000000000000000000000000000000000000000000000000000000425a6a188834be37ce5dd881f9ba854d08388861e04893c41700b0160, 0x0000000000000000000000000000000000000000000000000000000000000000000000032b77bd6035b22ffb468ed013a388ec81a8763eae14527949a28bfa32]
]
for [i, t, r] in D:
print(i, t % m, r % m)
assert t % m == 0
assert r % m == (a * b) % m
a = rand() % prime
b = rand() % prime
print(U2562(a))
print(U2562(b))
print(U2562((a + b) % prime ))
def cpoint(x, y):
return 'CurvePoint{\n x: FieldElement(' + U2562(x) + '),\n y: FieldElement(' + U2562(y) + '),\n}'
cpoint(a, b)
constant(0x14023b44fbb1e6f2a79c929c6da775be3c4b9e043d439385b5050fdc69177e3)
x = 0x0548c135e26faa9c977fb2eda057b54b2e0baa9a77a0be7c80278f4f03462d4c
y = 0x024385f6bebc1c496e09955db534ef4b1eaff9a78e27d4093cfa8f7c8f886f6b
r = x * y * modinv(2**256, prime) % prime
hex(r)
def mul_red(x, y):
A = 0
for i in range(4):
a0 = A % 2**64
xi = (x >> (64 * i)) % 2**64
y0 = y % 2**64
u = ((a0 + xi * y0) * (-1)) % 2**64
print('u = ', hex(u))
A += xi * y
print('A = ', hex(A))
A += u * prime
print('A = ', hex(A))
A >>= 64
print('A = ', hex(A))
mul_red(x, y)
ui = (0x0000000000000000 + 0x80278f4f03462d4c * 0x3cfa8f7c8f886f6b) * 0xffffffffffffffff
ui %= 2**64
hex(ui)
A = 0x292309a7824ead8f4857284de8df558cba373e8fe0ae7f04f3d11ef358d1fc40000000000000000
A = 0x59d9aa979919b6907dc3904d75a1e63384ffab48f66508cec166da1d7fac41a0000000000000000
B = 0x02916bc86cea6d5ecdf89ac9f418566b384ffab48f66508cec166da1d7fac4199e7a23de6b1a3f88
A - B
u = (0x4f3d11ef358d1fc4 + 0x2e0baa9a77a0be7c * 0x3cfa8f7c8f886f6b) * 0xffffffffffffffff
hex(u % 2**64)
hex(21888242871839275222246405745257275088696311157297823662689037894645226208583)
0x0800000000000011000000000000000000000000000000000000000000000001
2 + 2*252
2 + 2 * 252
1000 / 2.6
500 / 120
2154686749748910716 < 2**60
rand() % 2**53
```
| github_jupyter |
# 4.3 Linear Discriminant Analysis
Suppose $f_k(x)$ is the class-conditional density of X, and let $\pi_k$ be the prior-probability, with $\sum \pi_k = 1$.
The Bayes theorem gives us (4.7):
$$
Pr(G = k, X = x) = \cfrac{f_k(x)\pi_k}{\sum_{l=1}^K f_l(x)\pi_l}
$$
Suppose that we model each class density as multivariate Gaussian (4.8):
$$
f_k(x) = \cfrac{1}{(2\pi)^{p/2} |\Sigma_k|^{1/2}} e^{-\frac{1}{2}(x-\mu_k)^T\Sigma_k^{-1}(x-\mu_k)}
$$
Linear discriminant analysis (LDA) arises when $\Sigma_k = \Sigma \text{ }\forall k$. The log-ration between two classes $k \text{ and } l$ is (4.9):
$$
\begin{align}
log \cfrac{PR(G=k|X=x)}{PR(G=l|X=x)} &= log \cfrac{f_k(x)}{f_l(x)} + log \cfrac{\pi_k}{\pi_l}\\
&= log \cfrac{\pi_k}{\pi_l} - \frac{1}{2}(\mu_k+\mu_l)^T\Sigma^{-1}(\mu_k - \mu_l)
+ x^T\Sigma^{-1}(\mu_k-\mu_l),
\end{align}
$$
an equation is linear in x.
From (4.9) we see that the linear discriminant functions (4.10):
$$
\delta_k(x) = x^T\Sigma^{-1}\mu_k - \frac{1}{2}\mu_k^T\Sigma^{-1}\mu_k + log \pi_k
$$
are an equivalent description of the decision rule, with $G(x) = argmax_k \delta_k(x)$.
In practice we do not know the parameters of the Gaussian distributions, and will need to estimate them using the training data:
- $\hat{\pi_k} = N_k / N, N_k$ is the number of class-k observations;
- $\hat{\mu}_k = \sum_{g_i = k} x_i / N_k$
- $\hat{\Sigma} = \sum_{k=1}^K\sum_{g_i=k} (x_i - \hat{\mu}_k)(x_i-\hat{\mu}_k)^T / (N - K)$
With two classes, the LDA rule classifies to class 2 if (4.11):
$$
x^T\hat{\Sigma}^{-1}(\hat{\mu}_2 - \hat{\mu}_1) >
\frac{1}{2}\hat{\mu}_2^T\hat{\Sigma}^{-1}\hat{\mu}_2
- \frac{1}{2}\hat{\mu}_1^T\hat{\Sigma}^{-1}\hat{\mu}_1
+ log(N_1/N)
- log(N_2/N)
$$
Suppose we code the targets in the 2-classes as +1 and -1. It is easy to show that the coefficient vector from least squares is proportional to the LDA direction given in (4.11). However unless $N_1 = N_2$ the intercepts are different.
(**TODO**: solve exercise 4.11)
Since LDA direction via least squares does not use a Gaussian assumption, except the derivation of the intercept or cut-point via (4.11). Thus it makes sense to choose the cut-point that minimizes the training error.
With more than two classes, LDA is not the same as linear regression and it avoids the masking problems.
**Quadratic discriminant functions**
If $\Sigma_k$ are assumed to be equal, then we get *quadratic discriminant functions (QDA)* (4.12):
$$
\delta_k(x)=\cfrac{1}{2}log|\Sigma_k| - \cfrac{1}{2}(x-\mu_k)^T\Sigma_k^{-1}(x-\mu_k)+log \pi_k
$$
| github_jupyter |
# Mask R-CNN - Inspect Custom Trained Model
Code and visualizations to test, debug, and evaluate the Mask R-CNN model.
```
import os
import cv2
import sys
import random
import math
import re
import time
import numpy as np
import tensorflow as tf
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.patches as patches
import skimage
import glob
# Root directory of the project
ROOT_DIR = os.getcwd()
# Import Mask RCNN
sys.path.append(ROOT_DIR) # To find local version of the library
from mrcnn import utils
from mrcnn import visualize
from mrcnn.visualize import display_images
import mrcnn.model as modellib
from mrcnn.model import log
import custom
%matplotlib inline
# Directory to save logs and trained model
MODEL_DIR = os.path.join(ROOT_DIR, "logs")
custom_WEIGHTS_PATH = "mask_rcnn_damage_0010.h5" # TODO: update this path
```
## Configurations
```
config = custom.CustomConfig()
custom_DIR = os.path.join(ROOT_DIR, "customImages")
# Override the training configurations with a few
# changes for inferencing.
class InferenceConfig(config.__class__):
# Run detection on one image at a time
GPU_COUNT = 1
IMAGES_PER_GPU = 1
config = InferenceConfig()
config.display()
```
## Notebook Preferences
```
# Device to load the neural network on.
# Useful if you're training a model on the same
# machine, in which case use CPU and leave the
# GPU for training.
DEVICE = "/cpu:0" # /cpu:0 or /gpu:0
# Inspect the model in training or inference modes
# values: 'inference' or 'training'
# TODO: code for 'training' test mode not ready yet
TEST_MODE = "inference"
def get_ax(rows=1, cols=1, size=16):
"""Return a Matplotlib Axes array to be used in
all visualizations in the notebook. Provide a
central point to control graph sizes.
Adjust the size attribute to control how big to render images
"""
_, ax = plt.subplots(rows, cols, figsize=(size*cols, size*rows))
return ax
```
## Load Validation Dataset
```
# Load validation dataset
dataset = custom.CustomDataset()
dataset.load_custom(custom_DIR, "val")
# Must call before using the dataset
dataset.prepare()
print("Images: {}\nClasses: {}".format(len(dataset.image_ids), dataset.class_names))
```
## Load Model
```
# Create model in inference mode
with tf.device(DEVICE):
model = modellib.MaskRCNN(mode="inference", model_dir=MODEL_DIR,
config=config)
# load the last model you trained
# weights_path = model.find_last()[1]
# Load weights
print("Loading weights ", custom_WEIGHTS_PATH)
model.load_weights(custom_WEIGHTS_PATH, by_name=True)
from importlib import reload # was constantly changin the visualization, so I decided to reload it instead of notebook
reload(visualize)
```
# Run Detection on Images
```
image_id = random.choice(dataset.image_ids)
image, image_meta, gt_class_id, gt_bbox, gt_mask =\
modellib.load_image_gt(dataset, config, image_id, use_mini_mask=False)
info = dataset.image_info[image_id]
print("image ID: {}.{} ({}) {}".format(info["source"], info["id"], image_id,
dataset.image_reference(image_id)))
# Run object detection
results = model.detect([image], verbose=1)
# Display results
ax = get_ax(1)
r = results[0]
visualize.display_instances(image, r['rois'], r['masks'], r['class_ids'],
dataset.class_names, r['scores'], ax=ax,
title="Predictions")
log("gt_class_id", gt_class_id)
log("gt_bbox", gt_bbox)
log("gt_mask", gt_mask)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/Prim9000/Thai_TTS/blob/main/Thai_TTS_Training.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Extracting the Dataset
```
import os
import shutil
def download():
url = "https://github.com/korakot/corpus/releases/download/v1.0/AIFORTHAI-TSync2Corpus.zip"
print("NECTEC licensed TSync2 under CC-BY-NC-SA")
print("Start downloading: .. ")
os.system(f"wget {url}")
os.system("unzip AIFORTHAI-TSync2Corpus.zip")
os.system("rm AIFORTHAI-TSync2Corpus.zip")
shutil.move('/content/TSync2/wav','/content/wav')
shutil.move('/content/TSync2/wrd_ph','/content/wrd_ph')
os.system("rm /content/TSync2")
print("Finished")
download()
!unzip /content/drive/MyDrive/TSync2/TSync2.zip
```
# Trim silence sampling rate = 220505
```
import matplotlib.pyplot as plt
import os
import librosa
import shutil
import soundfile as sf
from tqdm.auto import tqdm
def trim(directory,filename,sr=22050, threshold=20):
new_filename = "{}.wav".format(filename[:-4])
signal, sr = librosa.load(os.path.join(directory,filename), sr=sr)
trimed, index = librosa.effects.trim(signal, top_db=threshold)
sf.write(os.path.join(directory, new_filename), trimed, samplerate=sr)
shutil.move(os.path.join(directory, new_filename) , os.path.join('/content/wav', new_filename))
```
Trimming the silence (this would take a while)
```
source = '/content/wav/'
for root, dirnames, filenames in os.walk(source):
for filename in filenames:
try:
trim(source,filename)
except:
pass
```
# Tacotron2
```
%cd /content/
%tensorflow_version 1.x
import os
from os.path import exists, join, basename, splitext
git_repo_url = 'https://github.com/Prim9000/tacotron2.git'
project_name = splitext(basename(git_repo_url))[0]
if not exists(project_name):
# clone and install
!git clone -q --recursive {git_repo_url}
!cd {project_name}/waveglow && git checkout 9168aea
!pip install -q librosa unidecode
import sys
sys.path.append(join(project_name, 'waveglow/'))
sys.path.append(project_name)
import time
import matplotlib
import matplotlib.pylab as plt
plt.rcParams["axes.grid"] = False
def download_from_google_drive(file_id, file_name):
# download a file from the Google Drive link
!rm -f ./cookie
!curl -c ./cookie -s -L "https://drive.google.com/uc?export=download&id={file_id}" > /dev/null
confirm_text = !awk '/download/ {print $NF}' ./cookie
confirm_text = confirm_text[0]
!curl -Lb ./cookie "https://drive.google.com/uc?export=download&confirm={confirm_text}&id={file_id}" -o {file_name}
tacotron2_pretrained_model = 'tacotron2_statedict.pt'
if not exists(tacotron2_pretrained_model):
# download the Tacotron2 pretrained model
download_from_google_drive('1c5ZTuT7J08wLUoVZ2KkUs_VdZuJ86ZqA', tacotron2_pretrained_model)
waveglow_pretrained_model = 'waveglow_old.pt'
if not exists(waveglow_pretrained_model):
# download the Waveglow pretrained model
download_from_google_drive('1WsibBTsuRg_SF2Z6L6NFRTT-NjEy1oTx', waveglow_pretrained_model)
import IPython.display as ipd
import numpy as np
import torch
from hparams import create_hparams
from model import Tacotron2
from layers import TacotronSTFT
from audio_processing import griffin_lim
from text import text_to_sequence
from denoiser import Denoiser
def plot_data(data, figsize=(16, 4)):
fig, axes = plt.subplots(1, len(data), figsize=figsize)
for i in range(len(data)):
axes[i].imshow(data[i], aspect='auto', origin='bottom',
interpolation='none', cmap='viridis')
torch.set_grad_enabled(False)
# initialize Tacotron2 with the pretrained model
hparams = create_hparams()
hparams.sampling_rate = 22050
model = Tacotron2(hparams)
model.load_state_dict(torch.load(tacotron2_pretrained_model)['state_dict'])
_ = model.cuda().eval()#.half()
# initialize Waveglow with the pretrained model
# waveglow = torch.load(waveglow_pretrained_model)['model']
# WORKAROUND for: https://github.com/NVIDIA/tacotron2/issues/182
import json
from glow import WaveGlow
waveglow_config = json.load(open('%s/waveglow/config.json' % project_name))['waveglow_config']
waveglow = WaveGlow(**waveglow_config)
waveglow.load_state_dict(torch.load(waveglow_pretrained_model)['model'].state_dict())
_ = waveglow.cuda().eval()#.half()
for k in waveglow.convinv:
k.float()
denoiser = Denoiser(waveglow)
%cd /content/tacotron2
shutil.move('/content/tacotron2_statedict.pt','/content/tacotron2/tacotron2_statedict.pt')
from scipy.io import wavfile
samplerate, data = wavfile.read('/content/wav/tsync2_noon_0_1228.wav')
```
# Training
```
%load_ext tensorboard
%tensorboard --logdir=outdir/logdir
!gdown --id 1tukxLX1Ul2O3zpTztX18K6WWfMrPiXJ2
# Training from the beginning
#!python train.py --output_directory=outdir --log_directory=logdir -c tacotron2_statedict.pt --warm_start
# Training from scratch (cold start)
#!python train.py --output_directory=outdir --log_directory=logdir
!python train.py --output_directory=outdir --log_directory=logdir -c /content/tacotron2/checkpoint_10000 --warm_start
```
# Synthesizing Speech
```
import matplotlib
%matplotlib inline
import matplotlib.pylab as plt
import IPython.display as ipd
import sys
sys.path.append('waveglow/')
import numpy as np
import torch
from hparams import create_hparams
from model import Tacotron2
from layers import TacotronSTFT, STFT
from audio_processing import griffin_lim
from train import load_model
from text import text_to_sequence
from denoiser import Denoiser
def plot_data(data, figsize=(16, 4)):
fig, axes = plt.subplots(1, len(data), figsize=figsize)
for i in range(len(data)):
axes[i].imshow(data[i], aspect='auto', origin='bottom',
interpolation='none')
hparams = create_hparams()
hparams.sampling_rate = 22050
```
### Change your checkpoint
```
#change your checkpoint path here
checkpoint_path ='/content/tacotron2/checkpoint_10000' # '/content/tacotron2/outdir/checkpoint_0'
model = load_model(hparams)
model.load_state_dict(torch.load(checkpoint_path)['state_dict'])
_ = model.cuda().eval().half()
waveglow_path = '/content/waveglow_old.pt'
waveglow = torch.load(waveglow_path)['model']
waveglow.cuda().eval().half()
for k in waveglow.convinv:
k.float()
denoiser = Denoiser(waveglow)
!pip install pythainlp
```
## Change your text
```
text = 'ยินดีที่ได้รู้จัก นี่คือเสียงจากปัญญาประดิษฐ์'
from pythainlp import word_tokenize
def text_process(text):
final = text
final = word_tokenize(final)
final = " ".join(word for word in final)
final += " ."
return final
text = text_process(text)
text
sequence = np.array(text_to_sequence(text, ['english_cleaners']))[None, :]
sequence = torch.autograd.Variable(
torch.from_numpy(sequence)).cuda().long()
mel_outputs, mel_outputs_postnet, _, alignments = model.inference(sequence)
%matplotlib inline
plot_data((mel_outputs.float().data.cpu().numpy()[0],
mel_outputs_postnet.float().data.cpu().numpy()[0],
alignments.float().data.cpu().numpy()[0].T))
```
## The Synthesized Speech
```
with torch.no_grad():
audio = waveglow.infer(mel_outputs_postnet, sigma=0.666)
ipd.Audio(audio[0].data.cpu().numpy(), rate=hparams.sampling_rate)
```
Add Denoiser
```
audio_denoised = denoiser(audio, strength=0.01)[:, 0]
ipd.Audio(audio_denoised.cpu().numpy(), rate=hparams.sampling_rate)
```
| github_jupyter |
```
# Erasmus+ ICCT project (2018-1-SI01-KA203-047081)
# Toggle cell visibility
from IPython.display import HTML
tag = HTML('''<script>
code_show=true;
function code_toggle() {
if (code_show){
$('div.input').hide()
} else {
$('div.input').show()
}
code_show = !code_show
}
$( document ).ready(code_toggle);
</script>
Toggle cell visibility <a href="javascript:code_toggle()">here</a>.''')
display(tag)
# Hide the code completely
# from IPython.display import HTML
# tag = HTML('''<style>
# div.input {
# display:none;
# }
# </style>''')
# display(tag)
%matplotlib notebook
import matplotlib.pyplot as plt
import numpy as np
import sympy as sym
import scipy.signal as signal
from ipywidgets import widgets, interact
```
## PID krmilnik - zaprtozančni sistem
Proporcionalno-integrirni-diferencirni (PID) krmilni algoritem je najpogosteje uporabljen krmilni algoritem. Njegovo prenosno funkcijo zapišemo kot:
\begin{equation}
P(s)=K_p \cdot \left( 1 + \frac{1}{T_i s} + T_d s \right).
\end{equation}
Prenosna funkcija je sestavljena iz vsote proporcionalne, integrirne in diferencirne komponente. Ni nujno, da so v izbranem krmilniku prisotne vse tri komponente; če ni diferencirne ali integrirne komponente, govorimo tako o PI oz. PD krmilniku. V tem interaktivnem primeru je prikazan odziv P, Pi, PD in PID krmilnika na enotsko skočno, enotsko impulzno in sinusno funkcijo ter enotsko rampo. Krmilnik je v tem primeru del krmilnega sistema s povratno zvezo. Objekt je lahko proporcionalni objekt ničtega, prvega ali drugega reda, ali pa integrirni objekt ničtega ali prvega reda.
Spodnji grafi prikazujejo.
1. Odziv zaprtozančnega sistema na izbran vstopni signal, izbran tip objekta in izbrani krmilnik (levi graf).
2. Lego ničel in polov prenosne funkcije rezultirajočega zaprtozancnega sistema.
---
### Kako upravljati s tem interaktivnim primerom?
1. Izberi vstopni signal s preklapljanjem med *enotsko skočno funkcijo*, *enotsko impulzno funkcijo*, *enotsko rampo* in *sinusno funkcijo*.
2. Izberi tip objekta: *P0* (proporcionalni objekt ničtega reda), *P1* (proporcionalni objekt prvega reda), *I0* (integrirni objekt ničtega reda) ali *I1* (integrirni objekt prvega reda). Prenosna funkcija objekta P0 je $k_p$ (v tem interaktivnem primeru $k_p=2$), PI objekta $\frac{k_p}{\tau s+1}$ (v tem interaktivnem primeru $k_p=1$ and $\tau=2$), IO objekta $\frac{k_i}{s}$ (v tem interaktivnem primeru $k_i=\frac{1}{10}$) in I1 objekta $\frac{k_i}{s(\tau s +1)}$ (v tem interaktivnem primeru $k_i=1$ in $\tau=10$).
3. Izberi tip krmilnega algoritma s klikom na *P*, *PI*, *PD* ali *PID* gumb.
4. Z uporabo drsnikov spreminjaj vrednosti koeficientov proporcionalnega ($K_p$), integrirnega ($T_i$) in diferencirnega ($T_d$) ojačnja.
5. Z uporabo drsnika $t_{max}$ lahko spreminjaš interval vrednosti prikazanih na x osi.
<!-- A proportional–integral–derivative (PID) control algorithm is by far the most common control algorithm. Its transfer function is equal to:
\begin{equation}
P(s)=K_p \cdot \left( 1 + \frac{1}{T_i s} + T_d s \right).
\end{equation}
It is made as a sum of proportional, integral and derivative channels. Not all of them have to be present, so PI or PD control algorithms are also used. In this example the response of a P, PI, PD or PID controller is shown for unit step, unit impulse, unit ramp or sine input. The controller is in this case part of a feedback control system. The object can either be a proportional of the zeroth, first or second order, or an integral of zeroth or first order.
The plots below show:
1. The response of the closed-loop system for the selected input, object type and the selected controller (left figure).
2. The position of the zeros and poles of the transfer function of the resulting closed-loop system.
---
### How to use this notebook?
1. Toggle between *unit step function*, *unit impulse function*, *unit ramp function*, and *sine* to select the input signal.
2. Click on *P0*, *P1*, *I0* or *I1* to toggle between the following objects: proportional of the zeroth, first or second order, or an integral one of zeroth or first order. The transfer function of P0 object is $k_p$ (in this example $k_p=2$), of PI object $\frac{k_p}{\tau s+1}$ (in this example $k_p=1$ and $\tau=2$), of IO object $\frac{k_i}{s}$ (in this example $k_i=\frac{1}{10}$) and of I1 object $\frac{k_i}{s(\tau s +1}$ (in this example $k_i=1$ and $\tau=10$).
3. Click on the *P*, *PI*, *PD* and *PID* buttons to toogle between proportional, proportional-integral, proportional-derivative and proportional–integral–derivative control algorithm type.
4. Move the sliders to change the values of proportional ($K_p$), integral ($T_i$) and derivative ($T_d$) PID tunning coefficients.
5. Move the slider $t_{max}$ to change the maximum value of the time on x axis of the Time response plot. -->
```
A = 10
a=0.1
s, P, I, D = sym.symbols('s, P, I, D')
obj = 1/(A*s)
PID = P + P/(I*s) + P*D*s#/(a*D*s+1)
system = obj*PID/(1+obj*PID)
num = [sym.fraction(system.factor())[0].expand().coeff(s, i) for i in reversed(range(1+sym.degree(sym.fraction(system.factor())[0], gen=s)))]
den = [sym.fraction(system.factor())[1].expand().coeff(s, i) for i in reversed(range(1+sym.degree(sym.fraction(system.factor())[1], gen=s)))]
# make figure
fig = plt.figure(figsize=(9.8, 4),num='PID krmilnik - zaprtozančni sistem')
plt.subplots_adjust(wspace=0.3)
# add axes
ax = fig.add_subplot(121)
ax.grid(which='both', axis='both', color='lightgray')
ax.set_title('Časovni odziv')
ax.set_xlabel('$t$ [s]')
ax.set_ylabel('vhod, izhod')
ax.axhline(linewidth=.5, color='k')
ax.axvline(linewidth=.5, color='k')
rlocus = fig.add_subplot(122)
input_type = 'enotska skočna funkcija'
# plot step function and responses (initalisation)
input_plot, = ax.plot([],[],'C0', lw=1, label='vstopni signal')
response_plot, = ax.plot([],[], 'C1', lw=2, label='izstopni signal')
ax.legend()
rlocus_plot, = rlocus.plot([], [], 'r')
plt.show()
def update_plot(KP, TI, TD, Time_span):
global num, den, input_type
num_temp = [float(i.subs(P,KP).subs(I,TI).subs(D,TD)) for i in num]
den_temp = [float(i.subs(P,KP).subs(I,TI).subs(D,TD)) for i in den]
system = signal.TransferFunction(num_temp, den_temp)
zeros = np.roots(num_temp)
poles = np.roots(den_temp)
rlocus.clear()
rlocus.scatter([np.real(i) for i in poles], [np.imag(i) for i in poles], marker='x', color='g', label='pol')
rlocus.scatter([np.real(i) for i in zeros], [np.imag(i) for i in zeros], marker='o', color='g', label='ničla')
rlocus.set_title('Diagram lege ničel in polov')
rlocus.set_xlabel('Re')
rlocus.set_ylabel('Im')
rlocus.grid(which='both', axis='both', color='lightgray')
time = np.linspace(0, Time_span, 300)
if input_type == 'enotska skočna funkcija':
u = np.ones_like(time)
u[0] = 0
time, response = signal.step(system, T=time)
elif input_type == 'enotska impulzna funkcija':
u = np.zeros_like(time)
u[0] = 10
time, response = signal.impulse(system, T=time)
elif input_type == 'sinusna funkcija':
u = np.sin(time*2*np.pi)
time, response, _ = signal.lsim(system, U=u, T=time)
elif input_type == 'enotska rampa':
u = time
time, response, _ = signal.lsim(system, U=u, T=time)
else:
raise Exception("Error in the program. Please restart simulation.")
response_plot.set_data(time, response)
input_plot.set_data(time, u)
rlocus.axhline(linewidth=.3, color='k')
rlocus.axvline(linewidth=.3, color='k')
rlocus.legend()
ax.set_ylim([min([np.min(u), min(response),-.1]),min(100,max([max(response)*1.05, 1, 1.05*np.max(u)]))])
ax.set_xlim([-0.1,max(time)])
plt.show()
controller_ = PID
object_ = obj
def calc_tf():
global num, den, controller_, object_
system_func = object_*controller_/(1+object_*controller_)
num = [sym.fraction(system_func.factor())[0].expand().coeff(s, i) for i in reversed(range(1+sym.degree(sym.fraction(system_func.factor())[0], gen=s)))]
den = [sym.fraction(system_func.factor())[1].expand().coeff(s, i) for i in reversed(range(1+sym.degree(sym.fraction(system_func.factor())[1], gen=s)))]
update_plot(Kp_widget.value, Ti_widget.value, Td_widget.value, time_span_widget.value)
def transfer_func(controller_type):
global controller_
proportional = P
integral = P/(I*s)
differential = P*D*s/(a*D*s+1)
if controller_type =='P':
controller_func = proportional
Kp_widget.disabled=False
Ti_widget.disabled=True
Td_widget.disabled=True
elif controller_type =='PI':
controller_func = proportional+integral
Kp_widget.disabled=False
Ti_widget.disabled=False
Td_widget.disabled=True
elif controller_type == 'PD':
controller_func = proportional+differential
Kp_widget.disabled=False
Ti_widget.disabled=True
Td_widget.disabled=False
else:
controller_func = proportional+integral+differential
Kp_widget.disabled=False
Ti_widget.disabled=False
Td_widget.disabled=False
controller_ = controller_func
calc_tf()
def transfer_func_obj(object_type):
global object_
if object_type == 'P0':
object_ = 2
elif object_type == 'P1':
object_ = 1/(2*s+1)
elif object_type == 'I0':
object_ = 1/(10*s)
elif object_type == 'I1':
object_ = 1/(s*(10*s+1))
calc_tf()
style = {'description_width': 'initial'}
def buttons_controller_clicked(event):
controller = buttons_controller.options[buttons_controller.index]
transfer_func(controller)
buttons_controller = widgets.ToggleButtons(
options=['P', 'PI', 'PD', 'PID'],
description='Izberi tip krmilnega algoritma:',
disabled=False,
style=style)
buttons_controller.observe(buttons_controller_clicked)
def buttons_object_clicked(event):
object_ = buttons_object.options[buttons_object.index]
transfer_func_obj(object_)
buttons_object = widgets.ToggleButtons(
options=['P0', 'P1', 'I0', 'I1'],
description='Izberi tip objekta:',
disabled=False,
style=style)
buttons_object.observe(buttons_object_clicked)
def buttons_input_clicked(event):
global input_type
input_type = buttons_input.options[buttons_input.index]
update_plot(Kp_widget.value, Ti_widget.value, Td_widget.value, time_span_widget.value)
buttons_input = widgets.ToggleButtons(
options=['enotska skočna funkcija','enotska impulzna funkcija', 'enotska rampa', 'sinusna funkcija'],
description='Izberi vstopni signal:',
disabled=False,
style = {'description_width': 'initial','button_width':'180px'})
buttons_input.observe(buttons_input_clicked)
Kp_widget = widgets.IntSlider(value=10,min=1,max=50,step=1,description=r'\(K_p\)',
disabled=False,continuous_update=True,orientation='horizontal',readout=True,readout_format='.1d')
Ti_widget = widgets.FloatLogSlider(value=1.,min=-3,max=1.1,step=.001,description=r'\(T_{i} \)',
disabled=False,continuous_update=True,orientation='horizontal',readout=True,readout_format='.3f')
Td_widget = widgets.FloatLogSlider(value=1.,min=-3,max=1.1,step=.001,description=r'\(T_{d} \)',
disabled=False,continuous_update=True,orientation='horizontal',readout=True,readout_format='.3f')
time_span_widget = widgets.FloatSlider(value=10.,min=.5,max=50.,step=0.1,description=r'\(t_{max} \)',
disabled=False,continuous_update=True,orientation='horizontal',readout=True,readout_format='.1f')
transfer_func(buttons_controller.options[buttons_controller.index])
transfer_func_obj(buttons_object.options[buttons_object.index])
display(buttons_input)
display(buttons_object)
display(buttons_controller)
interact(update_plot, KP=Kp_widget, TI=Ti_widget, TD=Td_widget, Time_span=time_span_widget);
```
| github_jupyter |
# Introduction to Numpy
## NDArray
```
import numpy as np
data = np.array([10, 11, 12, 13])
print(type(data)) # data is of object type ndarray
```
## Numpy dimensions
```
data = np.array([([1, 2]), ([3, 4])])
# print the shape of object "data"
print(data.shape)
```
```
var = np.array(
[([11, 22, 33, 44, 45, 46]), ([36, 37, 38, 39, 40, 41]), ([61, 62, 63, 64, 65, 66])]
)
# print the shape of object "var"
var.shape
```
## Storage List Vs Numpy Array
```
a = [1, 2, 3, 4]
b = np.array([1, 2, 3, 4])
# Size of the numpy array
print("Size of Numpy Array ", b.itemsize * b.size, "bytes")
# Size of the list
import sys
print("Size of the Python List :", sys.getsizeof(3) * len(a), "bytes")
```
## Speed List Vs Numpy Array
```
import time
a1 = range(10000000)
a2 = range(10000000)
b1 = np.arange(10000000)
b2 = np.arange(10000000)
start = time.time()
result = [(x + y) for x, y in zip(a1, a2)]
print((time.time() - start) * 1000, "milliseconds")
start = time.time()
output = b1 + b2
print((time.time() - start) * 1000, "milliseconds")
```
## Python Lists
```
# Declare a list with arbitrary datatypes
# int, string, list and dictionary
a = [1, "Hello", 3.14, [2, 3, 4], {"a": 1, "b": 2}]
# Length of the list
print(len(a))
# Acessing the elements using index values
print(a[2])
```
## Numpy arrays
```
# import the numpy module
import numpy as np
# Declare a Numpy array and store it in variable b
b = np.array([1, 2, 3, 4, 5])
# Fetch the length of b (count of total elements in b)
print(len(b))
# Find the index value of third element in the array
# (index starts at 0 (zero))
print(b[2])
```
## Be careful while copying arrays
```
x = np.array([42, 55, 66])
print("x List:", x)
y = x
print("y List:", y)
y[0] = 100
print("y List:", y)
print("x List:", x) # now the value of x is also changed
x = np.array([42, 55, 66])
y = x.copy()
y[0] = 100
print("x List:", x)
print("y List:", y)
```
## Mathematical Operation
```
import numpy as np
a = np.array([1, 2, 3, 4])
b = np.array([5, 6, 7, 8])
print("Addition :", a + b)
import numpy as np
a = np.array([1, 2, 3, 4])
b = np.array([5, 6, 7, 8])
print("Subtraction :", a - b)
print("Multiplication :", a * b)
print("Division :", a / b)
```
## Trigometrial functions
```
a = [1, 2, 3, 4]
# Sin function
print("Sin values:", np.sin(a))
# Cos function
print("Cos values:", np.cos(a))
```
## Statistics
```
data = np.array([1, 23, 52, 3, 6.2, 72, 8, 19, 0, 38, 4, 57, 2, 4])
print(np.mean(data))
print(np.median(data))
print(np.std(data))
```
## Min, Max, Sum
```
x = np.array([[71, 12, 36], [42, 55, 26]])
print(" Minimum value :", np.min(x))
print(" Maximum value :", np.max(x))
print("Addition :", np.sum(x))
print("Multiplication :", np.product(x))
```
## Axis - 0 , 1
```
x = np.array([[71, 12, 36, 22], [42, 55, 26, 75], [12, 35, 56, 25]])
print(x)
print(x.shape)
```
#### Variable 'x' has 3 rows and 4 columns
```
print("Column-wise minimum values :", np.min(x, axis=0))
print("Column-wise maximum values :", np.max(x, axis=0))
print("Row-wise minimum values :", np.min(x, axis=1))
print("Row-wise maximum values :", np.max(x, axis=1))
print("Row-wise sum :", np.sum(x, axis=1))
print("Column-wise sum :", np.sum(x, axis=0))
```
## Reshape arrays
```
x1 = np.array([[1, 2, 3, 4], [5, 6, 7, 8]])
print(x1)
# Shape ( 2 rows and 4 columns)
print(x1.shape)
# Reshape to 4 rows and 2 column
x2 = x1.reshape(4, 2)
print(x2)
# Reshape to 8 rows and 1 column
x3 = x1.reshape(8, 1)
print(x3)
```
## Vertical stacking
```
v1 = np.array([1, 2, 3, 4])
v2 = np.array([5, 6, 7, 8])
v3 = np.vstack([v1, v2])
print(v3)
v1 = np.array([1,2,3,4])
v2 = np.array([5,6,8])
print(np.vstack([v1,v2]))
```
## Horizontal stacking
```
v1 = np.array([1, 2, 3, 4])
v2 = np.array([5, 6, 7, 8])
np.hstack([v1, v2])
```
## Fancy Indexing
```
import numpy as np
a = np.arange(30).reshape(6, 5)
print(a)
# a[rows, columns]
# displays the second,third and fifth row
# index starting at 0 and all the columns
a_index_2_3_5 = a[[2, 3, 5], :]
print(a_index_2_3_5)
# displays the elements of 3 and 5 row
# of columns 2 and 5
a_3_5 = a[[3, 5], 2:5]
print(a_3_5)
```
## Indexing with boolean Arrays
```
data = np.array(
[[1, 2, 3, 4, 5, 6, 7], [87, 3, 4, 28, 4, 5, 73], [91, 42, 53, 39, 13, 58, 33]],
dtype="int32",
)
data > 30 # boolean indexing
# Boolean indexing with Single condition
data_gt_zero = data[data > 30]
print(data_gt_zero)
# Boolean indexing with Multiple Conditions
data_gt_30_lt_100 = data[((data > 30) & (data < 100))]
print(data_gt_30_lt_100)
```
## Broadcasting
```
import numpy as np
a = np.array([11, 12, 13, 14])
b = np.array([10, 20, 30, 40])
print(a.shape)
print(b.shape)
c = a * b
print(c)
```
### Different shapes
```
x = np.arange(6).reshape(2, 3)
y = np.array([[1, 2, 3]])
print(x.shape)
print(y.shape)
print(x + y)
```
### Different shape conditions not met
```
a = np.arange(10).reshape(2, 5)
b = np.array([[1, 2, 3, 4, 5, 6, 7, 8, 9, 10]])
print(a)
print(b)
print(a.shape, b.shape)
a+b
```
| github_jupyter |
```
# dependencies
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.colors as mcolors
import random
import math
import time
from sklearn.model_selection import RandomizedSearchCV, train_test_split
from sklearn.svm import SVR
from sklearn.metrics import mean_squared_error, mean_absolute_error
import datetime
import operator
plt.style.use('seaborn')
%matplotlib inline
# Loading all the dataset
confirmed_cases=pd.read_csv("./static/output/df_confirmed.csv")
# Display the head of the dataset
confirmed_cases.head()
# Extracting all the columns using the .heys() function
cols = confirmed_cases.keys()
cols
# Extracting only the dates columns that have information of confirmed, deaths and recovered cases
confirmed = confirmed_cases.loc[:,cols[5]:cols[-2]]
# check the head of the outbreak cases
confirmed.head()
total=confirmed.sum(axis=0)
total
change=total.pct_change()
total_df=total.to_frame()
change=change.to_list()
total_df=total_df.reset_index()
total_df=total_df.rename(columns={"index":"date",0: "confirmed"})
total_df["pct_change"]=change
total_df
total_0_df=total_df.loc[total_df['confirmed'] != 0]
plt.bar(total_0_df["date"],total_0_df["confirmed"])
degrees = 45
plt.xticks(rotation=degrees)
plt.bar(total_0_df["date"],total_0_df["pct_change"])
degrees = 45
plt.xticks(rotation=degrees)
# Finding the total confirmed cases, death cases and the recovered cases and append the to an 4 empty lists
# Also, calculate the total mortality rate which is the death_sum/confirmed cases
dates = confirmed.keys()
world_cases= []
total_deaths= []
mortality_rate= []
total_recovered= []
for i in dates:
confirmed_sum=confirmed[i].sum()
# death_sum = deaths[i].sum()
# recovered_sum = recoveries[i].sum()
world_cases.append(confirmed_sum)
# total_deaths.append(death_sum)
# mortality_rate.append(death_sum/confirmed_sum)
# total_recovered.append(recovered_sum)
# Display each of the newly created variables
confirmed_sum
# death_sum
# recovered_sum
# world_cases
# Convert all the dates and the cases in the form of a numpy array
days_since_1_22 =np.array([i for i in range(len(dates))]).reshape(-1,1)
# world_cases = np.array(world_cases).reshape(-1,1)
# total_deaths = np.array(total_deaths).reshape(-1,1)
# total_recovered = np.array(total_recovered).reshape(-1,1)
days_since_1_22
# world_cases
# total_deaths
# total_recovered
# Future forecasting for the next 10 days
days_in_future = 10
future_forecast = np.array([i for i in range(len(dates)+days_in_future)]).reshape(-1,1)
adjusted_dates = future_forecast[:-10]
future_forecast
# Convert all the integers into datetime for better visualization
start = '1/22/2020'
start_date = datetime.datetime.strptime(start,'%m/%d/%Y')
future_forecast_dates = []
for i in range(len(future_forecast)):
future_forecast_dates.append(((start_date+datetime.timedelta(days=i)).strftime('%m/%d/%Y')))
# For visualization with the lates data of 22 of march
latest_confirmed = confirmed_cases[dates[-1]]
# latest_deaths = deaths_reported[dates[-1]]
# latest_recoveries = recovered_reported[dates[-1]]
latest_confirmed
# latest_deaths
# latest_recoveries
# Find the list of unique states
unique_states = list(confirmed_cases['state_name'].unique())
unique_states
# The next line of code will basically calculate the total number of confirmed cases by each state
state_confirmed_cases = []
no_cases = []
for i in unique_states:
cases = latest_confirmed[confirmed_cases['state_name']==i].sum()
if cases > 0:
state_confirmed_cases.append(cases)
else:
no_cases.append(i)
for i in no_cases:
unique_states.remove(i)
unique_states = [k for k,v in sorted(zip(unique_states,state_confirmed_cases),key=operator.itemgetter(1),reverse=True)]
for i in range(len(unique_states)):
state_confirmed_cases[i] = latest_confirmed[confirmed_cases['state_name']==unique_states[i]].sum()
# number of cases per state
print('Confirmed Cases by States:')
for i in range(len(unique_states)):
print(f'{unique_states[i]}:{state_confirmed_cases[i]} cases')
# handling nan values if there is any
nan_indices = []
for i in range(len(unique_states)):
if type(unique_states[i])== float:
nan_indices.append(i)
unique_states = list(unique_states)
state_confirmed_cases = list(state_confirmed_cases)
for i in nan_indices:
unique_states.pop(i)
state_confirmed_cases.pop(i)
# Plot a bar graph to see the total confirmed cases across different countries
plt.figure(figsize = (32,32))
plt.barh(unique_states,state_confirmed_cases)
plt.title('Number of Covid-19 Confirmed Cases in States')
plt.xlabel('Number of Covid19 Confirmed Cases')
plt.show()
# Only show 10 States with the most confirmed cases, the rest are grouped into the category named others
visual_unique_states = []
visual_confirmed_cases = []
others = np.sum(state_confirmed_cases[10:])
for i in range(len(state_confirmed_cases[:10])):
visual_unique_states.append(unique_states[i])
visual_confirmed_cases.append(state_confirmed_cases[i])
visual_unique_states.append('Others')
visual_confirmed_cases.append(others)
# Visualize the 10 states
plt.figure(figsize =(32,18))
plt.barh(visual_unique_states,visual_confirmed_cases)
plt.title('Number of Covid-19 Confirmed Cases in States',size=20)
plt.show()
y = confirmed["3/22/20"]
X = confirmed.drop("3/22/20",axis=1)
x_train_confirmed, x_test_confirmed, y_train_confirmed, y_test_confirmed = train_test_split(X,y, random_state=42)
# Building the SVM model
kernel= ['poly','sigmoid','rbf']
c = [0.01,0.1,1,10]
gamma = [0.01,0.1,1]
epsilon = [0.01,0.1,1]
shrinking = [True, False]
svm_grid = {'kernel':kernel,'C':c,'gamma':gamma,'epsilon':epsilon,'shrinking':shrinking}
svm = SVR()
svm_search = RandomizedSearchCV(svm,svm_grid,scoring = 'neg_mean_squared_error',cv=3,return_train_score=True,n_jobs=-1,n_iter=40,verbose=1)
svm_search.fit(x_train_confirmed,y_train_confirmed)
svm_search.best_params_
X.shape
svm_confirmed = svm_search.best_estimator_
svm_pred = svm_confirmed.predict(x_test_confirmed)
svm_confirmed
svm_pred
```
| github_jupyter |
# Energy System Modelling - Tutorial II
**Imports**
```
import numpy as np
import numpy.linalg
import pandas as pd
import matplotlib.pyplot as plt
plt.style.use('bmh')
%matplotlib inline
```
***
## Problem II.1
**(a) Compile the nodes list and the edge list.**
> **Remark:** While graph-theoretically both lists are unordered sets, let's agree on an ordering now which can serve as basis for the matrices in the following exercises: we sort everything in ascending numerical order, i.e. node 1 before node 2 and edge (1,2) before (1,4) before (2,3).
```
nodes = [0, 1, 2, 3, 4, 5]
edges = [(0, 1), (1, 2), (1, 4),
(1, 5), (2, 3), (2, 4),
(3, 4), (4, 5)]
```
***
**(b) Determine the order and the size of the network.**
```
N = len(nodes)
E = len(edges)
print("Order: {}\nSize: {}".format(N, E))
```
***
**(c) Compute the adjacency matrix $A$ and check that it is symmetric.**
> In graph theory and computer science, an adjacency matrix is a square matrix used to represent a finite graph. The elements of the matrix indicate whether pairs of vertices are adjacent or not in the graph.
Adjacency Matrix:
```
A = np.zeros((N, N))
for u, v in edges:
A[u, v] += 1
A[v, u] += 1
A
```
Check for symmetry:
```
(A == A.T).all()
```
***
**(d) Find the degree $k_n$ of each node $n$ and compute the average degree of the network.**
> In graph theory, the degree (or valency) of a vertex of a graph is the number of edges incident to the vertex, with loops counted twice.
```
k = A.sum(axis=1)
k
k.mean()
```
***
**(e) Determine the incidence matrix $K$ by assuming the links are always directed from smaller-numbered node to larger-numbered node, i.e. from node 2 to node 3, instead of from 3 to 2.**
> The unoriented incidence matrix (or simply incidence matrix) of an undirected graph is a $n \times m$ matrix $B$, where n and m are the numbers of vertices and edges respectively, such that $B_{i,j} = 1$ if the vertex $v_i$ and edge $e_j$ are incident and 0 otherwise.
```
K = np.zeros((N,E))
for i, (u, v) in enumerate(edges):
K[u,i] = 1
K[v,i] = -1
K
```
***
**(f) Compute the Laplacian $L$ of the network using $k_n$ and $A$. Remember that the Laplacian can also be computed as $L=KK^T$ and check that the two definitions agree.**
> The **Laplacian** (also: admittance matrix, Kirchhoff matrix, discrete Laplacian) is a matrix representation of a graph. It is defined as the difference of degree matrix and adjacency matrix. The **degree matrix** is a diagonal matrix which contains information about the degree of each vertex.
```
D = np.diag(A.sum(axis=1))
L = D - A
L
np.array_equal(L, K.dot(K.T))
```
***
**(g) Find the diameter of the network by looking at the graph.**
> The diameter of a network is the longest of all the calculated shortest paths in a network. It is the shortest distance between the two most distant nodes in the network. In other words, once the shortest path length from every node to all other nodes is calculated, the diameter is the longest of all the calculated path lengths. The diameter is representative of the linear size of a network.
By looking: Between nodes $0$ and $3$, f.ex. $0 \to 1 \to 2 \to 3$
***
## Problem II.2
If you map the nodes to `0=DK, 1=DE, 2=CH, 3=IT, 4=AT, 5=CZ` the network represents a small part of the European electricity network (albeit very simplified). In the repository, you can find the power imbalance time series for the six countries for January 2017 in hourly MW at `./data/imbalance.csv`. They have been derived from physical flows as published by [ENTSO-E](https://transparency.entsoe.eu/transmission-domain/physicalFlow/show)
The linear power flow is given by
$$p_i = \sum_j \tilde{L}_{i,j}\theta_j \qquad \text{and} \qquad f_l = \frac{1}{x_l} \sum_i K_{i,l}\theta_i, \qquad \text{where} \qquad \tilde{L}_{i,j}= \sum_l K_{i,l}\frac{1}{x_l} K_{j,l}$$
is the weighted Laplacian. For simplicity, we assume identity reactance on all links $x_l = 1$.
***
**Read data**
```
imbalance = pd.read_csv('data/imbalance.csv', index_col=0, parse_dates=True)
imbalance.head()
```
\begin{equation}
p_u = \sum_v L_{u,v} \theta_v
\end{equation}
***
**(a) Compute the voltage angles $\theta_j$ and flows $f_l$ for the first hour in the dataset with the convention of $\theta_0 = 0$; i.e. the slack bus is at node 0.**
> **Remark:** Linear equation systems are solved efficiently using [`numpy.linalg.solve`](https://docs.scipy.org/doc/numpy/reference/generated/numpy.linalg.solve.html).
Calculate the *voltage angles* first.
> Note, that we define the node for Denmark as slack and therefore force $\theta_{DK}=0$.
```
imbalance.iloc[0].values[1:]
theta = np.r_[0, np.linalg.solve(L[1:,1:], imbalance.iloc[0].values[1:])]
theta
```
Then, calculate the *flows*:
```
flows = K.T.dot(theta)
flows
```
***
**(b) Determine the average flow on each link for January 2017 and draw it as a directed network**
> **Hint:** You may want to make use of the function `np.vstack`.
```
flows = K.T.dot(np.vstack([np.zeros((1, len(imbalance))), np.linalg.solve(L[1:,1:], imbalance.values[:,1:].T)]))
flows.shape
avg_flows = flows.mean(axis=1)
avg_flows
```
| github_jupyter |
# Polynomial Regression
This Code template is for the regression analysis using a Polynomial Regression. Polynomial Regression can be performed in python using Polynomial Features along with Linear Regression in a pipeline.
### Required Packages
```
import warnings
import numpy as np
import pandas as pd
import seaborn as se
import matplotlib.pyplot as plt
from sklearn.preprocessing import PolynomialFeatures
from sklearn.model_selection import train_test_split
from sklearn.metrics import r2_score, mean_absolute_error, mean_squared_error
from sklearn.linear_model import LinearRegression
from sklearn.pipeline import make_pipeline
warnings.filterwarnings('ignore')
```
### Initialization
Filepath of CSV file
```
#filepath
file_path= ""
```
List of features which are required for model training .
```
#x_values
features=[]
```
Target feature for prediction.
```
#y_value
target=''
```
### Data Fetching
Pandas is an open-source, BSD-licensed library providing high-performance, easy-to-use data manipulation and data analysis tools.
We will use panda's library to read the CSV file using its storage path.And we use the head function to display the initial row or entry.
```
df=pd.read_csv(file_path)
df.head()
```
### Feature Selections
It is the process of reducing the number of input variables when developing a predictive model. Used to reduce the number of input variables to both reduce the computational cost of modelling and, in some cases, to improve the performance of the model.
We will assign all the required input features to X and target/outcome to Y.
```
X=df[features]
Y=df[target]
```
### Data Preprocessing
Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
```
def NullClearner(df):
if(isinstance(df, pd.Series) and (df.dtype in ["float64","int64"])):
df.fillna(df.mean(),inplace=True)
return df
elif(isinstance(df, pd.Series)):
df.fillna(df.mode()[0],inplace=True)
return df
else:return df
def EncodeX(df):
return pd.get_dummies(df)
```
Calling preprocessing functions on the feature and target set.
```
x=X.columns.to_list()
for i in x:
X[i]=NullClearner(X[i])
X=EncodeX(X)
Y=NullClearner(Y)
X.head()
```
#### Correlation Map
In order to check the correlation between the features, we will plot a correlation matrix. It is effective in summarizing a large amount of data where the goal is to see patterns.
```
f,ax = plt.subplots(figsize=(18, 18))
matrix = np.triu(X.corr())
se.heatmap(X.corr(), annot=True, linewidths=.5, fmt= '.1f',ax=ax, mask=matrix)
plt.show()
```
### Data Splitting
The train-test split is a procedure for evaluating the performance of an algorithm. The procedure involves taking a dataset and dividing it into two subsets. The first subset is utilized to fit/train the model. The second subset is used for prediction. The main motive is to estimate the performance of the model on new data.
```
x_train,x_test,y_train,y_test=train_test_split(X,Y,test_size=0.2,random_state=123)
```
### Model
Polynomial regression is a special case of linear regression where we fit a polynomial equation on the data with a curvilinear relationship between the target variable and the independent variables.
In a curvilinear relationship, the value of the target variable changes in a non-uniform manner with respect to the predictor (s).
With scikit learn, it is possible to create one in a pipeline combining these two steps (Polynomialfeatures and LinearRegression).
PolynomialFeatures() function generate a new feature matrix consisting of all polynomial combinations of the features with degree less than or equal to the specified degree. For example, if an input sample is two dimensional and of the form [a, b], the degree-2 polynomial features are [1, a, b, a^2, ab, b^2].The 1-degree polynomial is a simple linear regression; therefore, the value of degree must be greater than 1.
#### Polynomial Feature Tuning parameters
> **degree** -> The degree of the polynomial features.
> **include_bias** -> If True (default), then include a bias column, the feature in which all polynomial powers are zero (i.e. a column of ones - acts as an intercept term in a linear model).
```
model=make_pipeline(PolynomialFeatures(),LinearRegression(n_jobs=-1))
model.fit(x_train,y_train)
```
#### Model Accuracy
We will use the trained model to make a prediction on the test set.Then use the predicted value for measuring the accuracy of our model.
```
print("Accuracy score {:.2f} %\n".format(model.score(x_test,y_test)*100))
```
> **r2_score**: The **r2_score** function computes the percentage variablility explained by our model, either the fraction or the count of correct predictions.
> **mae**: The **mean abosolute error** function calculates the amount of total error(absolute average distance between the real data and the predicted data) by our model.
> **mse**: The **mean squared error** function squares the error(penalizes the model for large errors) by our model.
```
y_pred=model.predict(x_test)
print("R2 Score: {:.2f} %".format(r2_score(y_test,y_pred)*100))
print("Mean Absolute Error {:.2f}".format(mean_absolute_error(y_test,y_pred)))
print("Mean Squared Error {:.2f}".format(mean_squared_error(y_test,y_pred)))
```
#### Prediction Plot
First, we make use of a plot to plot the actual observations, with x_train on the x-axis and y_train on the y-axis.
For the regression line, we will use x_train on the x-axis and then the predictions of the x_train observations on the y-axis.
```
plt.figure(figsize=(14,10))
plt.plot(range(20),y_test[0:20], color = "green")
plt.plot(range(20),model.predict(x_test[0:20]), color = "red")
plt.legend(["Actual","prediction"])
plt.title("Predicted vs True Value")
plt.xlabel("Record number")
plt.ylabel(target)
plt.show()
```
#### Creator: Thilakraj Devadiga , Github: [Profile](https://github.com/Thilakraj1998)
| github_jupyter |
<table border="0">
<tr>
<td>
<img src="https://ictd2016.files.wordpress.com/2016/04/microsoft-research-logo-copy.jpg" style="width 30px;" />
</td>
<td>
<img src="https://www.microsoft.com/en-us/research/wp-content/uploads/2016/12/MSR-ALICE-HeaderGraphic-1920x720_1-800x550.jpg" style="width 100px;"/></td>
</tr>
</table>
# Deep IV: Use Case and Examples
Deep IV uses deep neural networks in a two-stage instrumental variable (IV) estimation of causal effects, as described in [this ICML publication](http://proceedings.mlr.press/v70/hartford17a/hartford17a.pdf) or in the `econml` [specification](https://econml.azurewebsites.net/spec/estimation/deepiv.html). In the EconML SDK, we have implemented Deep IV estimation on top of the Keras framework for building and training neural networks. In this notebook, we'll demonstrate how to use the SDK to apply Deep IV to synthetic data.
### Data
Deep IV works in settings where we have several different types of observations:
* Covariates, which we will denote with `X`
* Instruments, which we will denote with `Z`
* Treatments, which we will denote with `T`
* Responses, which we will denote with `Y`
The main requirement is that `Z` is a set of valid instruments; in particular `Z` should affect the responses `Y` only through the treatments `T`. We assume that `Y` is an arbitrary function of `T` and `X`, plus an additive error term, and that `T` is an arbitrary function of `Z` and `X`. Deep IV then allows us to estimate `Y` given `T` and `X`.
### Estimation
To do this, the Deep IV estimator uses a two-stage approach that involves solving two subproblems:
1. It estimates the *distribution* of the treatment `T` given `Z` and `X`, using a mixture density network.
2. It estimates the dependence of the response `Y` on `T` and `X`.
Both of these estimates are performed using neural networks. See the paper for a more complete description of the setup and estimation approach.
### Using the SDK
In the `econml` package, our Deep IV estimator is built on top of the Keras framework; we support either the Tensorflow or the Theano backends. There are three steps to using the `DeepIVEstimator`:
1. Construct an instance.
* The `m` and `h` arguments to the initializer specify deep neural network models for estimating `T` and `Y` as described above. They are each *functions* that take two Keras inputs and return a Keras model (the inputs are `z` and `x` in the case of `m` and the output's shape should match `t`'s; the inputs are `t` and `x` in the case of `h` and the output's shape should match `y`'s). Note that the `h` function will be called multiple times, but should reuse the same weights - see below for a concrete example of how to achieve this using the Keras API.
* The `n_samples`, `use_upper_bound_loss`, and `n_gradient_samples` arguments together determine how the loss for the response model will be computed.
* If `use_upper_bound_loss` is `False` and `n_gradient_samples` is zero, then `n_samples` samples will be averaged to approximate the response - this will provide an unbiased estimate of the correct loss only in the limit as the number of samples goes to infinity.
* If `use_upper_bound_loss` is `False` and `n_gradient_samples` is nonzero, then we will average `n_samples` samples to approximate the response a first time and average `n_gradient_samples` samples to approximate it a second time - combining these allows us to provide an unbiased estimate of the true loss.
* If `use_upper_bound_loss` is `True`, then `n_gradient_samples` must be `0`; `n_samples` samples will be used to get an unbiased estimate of an upper bound of the true loss - this is equivalent to adding a regularization term penalizing the variance of the response model (see the `econml` specification linked above for a derivation of this fact).
2. Call `fit` with training samples of `Y`, `T`, `X`, and `Z`; this will train both sub-models.
3. Call `effect` or `predict` depending on what output you want. `effect` calculates the difference in outcomes based on the features and two different treatments, while `predict` predicts the outcome based on a single treatment.
The remainder of this notebook will walk through a concete example.
```
from econml.deepiv import DeepIVEstimator
import keras
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
```
## Synthetic data
To demonstrate the Deep IV approach, we'll construct a synthetic dataset obeying the requirements set out above. In this case, we'll take `X`, `Z`, `T` to come from the following distribution:
```
n = 5000
# Initialize exogenous variables; normal errors, uniformly distributed covariates and instruments
e = np.random.normal(size=(n,))
x = np.random.uniform(low=0.0, high=10.0, size=(n,))
z = np.random.uniform(low=0.0, high=10.0, size=(n,))
# Initialize treatment variable
t = np.sqrt((x+2) * z) + e
# Show the marginal distribution of t
plt.hist(t)
plt.xlabel("t")
plt.show()
plt.scatter(z[x < 1], t[x < 1], label='low X')
plt.scatter(z[(x > 4.5) * (x < 5.5)], t[(x > 4.5) * (x < 5.5)], label='moderate X')
plt.scatter(z[x > 9], t[x > 9], label='high X')
plt.legend()
plt.xlabel("z")
plt.ylabel("t")
plt.show()
```
Here, we'll imagine that `Z` and `X` are causally affecting `T`; as you can see in the plot above, low or high values of `Z` drive moderate values of `T` and moderate values of `Z` cause `T` to have a bi-modal distribution when `X` is high, but a unimodal distribution centered on 0 when `X` is low. The instrument is positively correlated with the treatment and treatments tend to be bigger at high values of x. The instrument has higher power at higher values of x
```
# Outcome equation
y = t*t / 10 - x*t / 10 + e
# The endogeneity problem is clear, the latent error enters both treatment and outcome equally
plt.scatter(t,z, label ='raw data')
tticks = np.arange(-2,12)
yticks2 = tticks*tticks/10 - 0.2 * tticks
yticks5 = tticks*tticks/10 - 0.5 * tticks
yticks8 = tticks*tticks/10 - 0.8 * tticks
plt.plot(tticks,yticks2, 'r--', label = 'truth, x=2')
plt.plot(tticks,yticks5, 'g--', label = 'truth, x=5')
plt.plot(tticks,yticks8, 'y--', label = 'truth, x=8')
plt.xlabel("t")
plt.ylabel("y")
plt.legend()
plt.show()
```
`Y` is a non-linear function of `T` and `X` with no direct dependence on `Z` plus additive noise (as required). We want to estimate the effect of particular `T` and `X` values on `Y`.
The plot makes it clear that looking at the raw data is highly misleading as to the treatment effect. Moreover the treatment effects are both non-linear and heterogeneous in x, so this is a hard problem!
## Defining the neural network models
Now we'll define simple treatment and response models using the Keras `Sequential` model built up of a series of layers. Each model will have an `input_shape` of 2 (to match the sums of the dimensions of `X` plus `Z` in the treatment case and `T` plus `X` in the response case).
```
treatment_model = keras.Sequential([keras.layers.Dense(128, activation='relu', input_shape=(2,)),
keras.layers.Dropout(0.17),
keras.layers.Dense(64, activation='relu'),
keras.layers.Dropout(0.17),
keras.layers.Dense(32, activation='relu'),
keras.layers.Dropout(0.17)])
response_model = keras.Sequential([keras.layers.Dense(128, activation='relu', input_shape=(2,)),
keras.layers.Dropout(0.17),
keras.layers.Dense(64, activation='relu'),
keras.layers.Dropout(0.17),
keras.layers.Dense(32, activation='relu'),
keras.layers.Dropout(0.17),
keras.layers.Dense(1)])
```
Now we'll instantiate the `DeepIVEstimator` class using these models. Defining the response model *outside* of the lambda passed into constructor is important, because (depending on the settings for the loss) it can be used multiple times in the second stage and we want the same weights to be used every time.
```
keras_fit_options = { "epochs": 30,
"validation_split": 0.1,
"callbacks": [keras.callbacks.EarlyStopping(patience=2, restore_best_weights=True)]}
deepIvEst = DeepIVEstimator(n_components = 10, # number of gaussians in our mixture density network
m = lambda z, x : treatment_model(keras.layers.concatenate([z,x])), # treatment model
h = lambda t, x : response_model(keras.layers.concatenate([t,x])), # response model
n_samples = 1, # number of samples to use to estimate the response
use_upper_bound_loss = False, # whether to use an approximation to the true loss
n_gradient_samples = 1, # number of samples to use in second estimate of the response (to make loss estimate unbiased)
optimizer='adam', # Keras optimizer to use for training - see https://keras.io/optimizers/
first_stage_options = keras_fit_options, # options for training treatment model
second_stage_options = keras_fit_options) # options for training response model
```
## Fitting and predicting using the model
Now we can fit our model to the data:
```
deepIvEst.fit(Y=y,T=t,X=x,Z=z)
```
And now we can create a new set of data and see whether our predicted effect matches the true effect `T*T-X*X`:
```
n_test = 500
for i, x in enumerate([2, 5, 8]):
t = np.linspace(0,10,num = 100)
y_true = t*t / 10 - x*t/10
y_pred = deepIvEst.predict(t, np.full_like(t, x))
plt.plot(t, y_true, label='true y, x={0}'.format(x),color='C'+str(i))
plt.plot(t, y_pred, label='pred y, x={0}'.format(x),color='C'+str(i),ls='--')
plt.xlabel('t')
plt.ylabel('y')
plt.legend()
plt.show()
```
You can see that despite the fact that the response surface varies with x, our model was able to fit the data reasonably well. Where is does worst is where the instrument has the least power, which is in the low x case. There it fits a straight line rather than a quadratic, which suggests that the regularization at least is perfoming well.
| github_jupyter |
<div align="right"><i>COM418 - Computers and Music</i></div>
<div align="right"><a href="https://people.epfl.ch/paolo.prandoni">Paolo Prandoni</a>, <a href="https://www.epfl.ch/labs/lcav/">LCAV, EPFL</a></div>
<p style="font-size: 30pt; font-weight: bold; color: #B51F1F;">Practical filters for Audio Processing</p>
```
%matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
from IPython.display import Audio
from scipy import signal
import import_ipynb
from FilterUtils import *
plt.rcParams['figure.figsize'] = 14, 4
matplotlib.rcParams.update({'font.size': 14})
DEFAULT_SF = 16000
```
# Introduction
In this notebook we will explore a complete set of "recipes" to design second-order digital IIR filters. The transfer function of a generic second-order section (also known as a **biquad**) has the canonical form
$$
H(z) = \frac{b_0 + b_1 z^{-1} + b_{2}z^{-2}}{1 + a_1 z^{-1} + a_{2}z^{-2}}
$$
The routines defined in the rest of this notebook will allow you to compute the values of the five biquad parameters in order to implement a variety of different filter prototypes according to the desired specifications. We will also explore how to cascade second-order sections to implement higher-order filters with improved characteristics.
## Common practices
Although ultimately we will design digital filters, in audio applications it is important to become familiar with the main ideas behind _analog_ filter design and analysis; indeed, audio recording and production techniques have been developed and fine-tuned well before the advent of DSP and, even in today's world of DAWs, the language and many of the practices in current use still reflect the analog conventions of yore.
In particular:
* filter specifications are almost always expressed in terms of real-world frequencies in Hz rather than as normalized frequencies over $[-\pi, \pi]$; this of course implies that the underlying sampling frequency is known
* plots of the magnitude response will usually be shown on a log-log graph, generally using a decibel scale for the amplitde and a decade scale for frequencies; this mirrors the way in which human audio perception is approximately logarithmic both in frequency and in scale.
The companion notebook ``FilterUtils.ipynb`` implements a set of plotting routines that take these conventions into account. For example, this is how the magnitude response of the same leaky integrator looks like in the typical representations used in communication systems vs. audio equalization:
```
lam = 0.9
filter_props([1 - lam], [1, -lam])
analog_response([1 - lam], [1, -lam], DEFAULT_SF, dB=-50)
```
# The biquad design strategy
The next section provides a set of functions to compute the five biquad filter coefficients for a desired response (lowpass, bandpass, etc) and an associated set of specifications (cutoff, attenuation, etc). Rather than tackling the search for the coefficients as an abstract optimization problem, each recipe starts from a well-known _analog_ second-order filter with the desired characteristic, and then converts it to a discrete-time filter using a mapping called the _bilinear transform_ .
The reason for this approach is that the classic analog filter prototypes (Butterworth, Chebyshev, etc.) (and the topologies used for their implementation) are extremely well understood and have proven extremely reliable over more than a century of research and practical experimentation.
## The analog prototypes
<img src="img/rlc.png" alt="rlc" style="float: right; width: 200px; margin: 0 10px 10px 30px;"/>
Historically, the development of electronic filters began with the design of **passive** analog filters, that is, filters using only resistors, capacitors and inductors; indeed, an RLC circuit, that is, a network containing a resistor, a capacitor and an inductor, can implement the prototypical analog second-order section. Since these filters have no active elements that can provide signal amplification, the power of their output is at most equal to (but, in practice, smaller than) the power of the input. This implicitly guarantees the stability of these systems although, at least in theory, strong resonances can appear in the frequency response.
Analog filters work by exploiting the frequency-dependent reactance of capacitors and inductors. The input-output characteristic for linear circuits using these electronic components is described by linear differential equations, which implies the existence of some form of _feedback_ in the circuits themselves. As a consequence, when we convert the analog prototypes to digital realizations, we invariably end up with IIR filters.
Passive filters operating in the frequency range of audio applications require the use of bulky inductors and therefore **active** filters are usually preferred in the analog domain. In the digital domain, on the other hand, we are in fact free to use arbitrary gain factors (it's just multiplications!) and so the resulting transfer functions can approximate either type of design.
## The bilinear transform
The cookbook recipes below are obtained by mapping second-order analog filters prototypes to equivalent digital realization via the _bilinear transform_ . We will not go into full details but, as a quick reference, here are the main ideas behind the method.
An analog filter is described by a transfer function $H(s)$, with $s\in \mathbb{C}$, which is the Laplace transform of the filter's continuous-time impulse response. The key facts about $H(s)$ are:
* filter stability requires that all the poles of $H(s)$ lie in the left half of the complex plane (i.e. their real part must be negative)
* the filter's frequency response is given by $H(j\Omega)$, that is, by the values of $H(s)$ along the imaginary axis
<img src="img/bilinear.png" alt="rlc" style="float: right; width: 300px; margin: 10px 0;"/>
The bilinear transform maps the complex $z$-plane (discrete time) to the complex $s$-plane (continuous time) as
$$
s \leftarrow c \frac{1 - z^{-1}}{1 + z^{-1}} = \Phi_{c}(z)
$$
where $c$ is a real-valued constant. Given a stable analog filter $H(s)$, the transfer function of its discrete-time version is $H_d(z) = H(\Phi_{c}(z))$ and it is relatively easy to verify that
* the inside of the unit circle on the $z$-plane is mapped to the left half of the $s$-plane, which preserves stability
* the unit circle on the $z$-plane is mapped to the imaginary axis of the $s$-plane
The last property allows us to determine the frequency response of the digital filter as $H_d(e^{j\omega}) = H(\Phi_{c}(e^{j\omega})) = H(j\,c\tan(\omega/2))$, or:
$$
\Omega \leftarrow c\tan(\omega/2) %\omega \leftarrow 2\arctan(\Omega/c)
$$
we can see that $\omega=0$ is mapped to $\Omega=0$, $\omega=\pi/2$ is mapped to $\Omega=c$, and $\omega=\pi$ is mapped to $\Omega=\infty$, which reveals the high nonlinearity of the frequency mapping. We usually need to precisely control a least one notable point $H_d(e^{j\omega_0})$ in the frequency response of the discrete-time filter; for example, in a resonator, we need to place the magnitude peak at a specific frequency $\omega_0$. To achieve this, we design the analog filter so that $H(1j) = H_d(e^{j\omega_0})$ and then we set $c = 1/\tan(\omega_0/2)$ in the bilinear operator; this adjustment, called **pre-warping** , is used in the recipes below.
To illustrate the principle, here is a simple transfer function $H(s)$ that provide a triangular response centered at $\Omega=1$; the default width is $1/2$ and the width can be optionally scaled.
along the imaginary axis; we can parametrize the width of the bell and its center position is at by default:
```
if __name__ == '__main__':
def H(f, scale=1):
return np.maximum(1 - 4 * np.abs(np.imag(f) - 1) / scale, 0)
f = np.linspace(0, 3, 1000)
plt.plot(f, H(1j * f));
```
Using the bilinear transform with pre-warping, we can move the equivalent discrete-time frequency response over the $[0, \pi]$ interval.
```
if __name__ == '__main__':
def BL(z, c=1):
return c * (1 - 1/z) / (1 + 1/z)
if __name__ == '__main__':
w = np.linspace(0, np.pi, 1000)
center_freqs = np.pi * np.arange(0.1, 0.9, 0.15)
for w0 in center_freqs:
c = 1 / np.tan(w0 / 2)
plt.plot(w, H(BL(np.exp(1j * w), c=c)))
```
Note that the nonlinear mapping between frequency axes has two consequences on the the discrete-time frequency response:
* at low and high digital frequencies the response becomes more narrow; this can be compensated for by scaling the analog prototype
* as we move to higher frequencies, the response is less and less symmetric; this is much harder to compensate for because it would require a different analog design and it is therefore an accepted tradeoff.
The following example tries to keep the width of the response uniform
```
if __name__ == '__main__':
for w0 in center_freqs:
c = 1 / np.tan(w0 / 2)
scaling_factor = (c * c + 1) / (2 * c)
plt.plot(w, H(BL(np.exp(1j * w), c=c), scale=scaling_factor))
```
**Exercise**: how was the scaling factor derived? Can you improve on it?
Using the bilinear transform with pre-warping, we can move the equivalent discrete-time frequency response over the $[0, \pi]$ interval
# What about FIRs?
FIR filters are a great tool in digital signal processing; as opposed to IIR (which can be seen as a digital adaptation of electronic filters) FIRs offer:
* unconditional stability
* the possibility of a linear phase response
* a great design algorithm (Parks-McClellan) even for arbitrary responses
The price for stability and linear phase is a much higher computational cost: for the same specifications, an FIR filter will require up to a hundred times more operations per sample with respect to an IIR implementation. Linear phase, however, is not terribly relevant in audio applications because of the limited phase sensitivity in the human auditory system. On the other hand, especially in real-time applications, the primary goal in audio processing is to minimize the overall processing delay; since linear phase FIRs have a symmetric impulse response, and since a well-performing filter will have a very long impulse response, the associated delay often makes FIRs difficult to use. Even if we give up linear phase and implement an asymmetric, minimum-phase FIR, the computational cost may be too high.
There are countless audiophile blogs debating the merits and demerits of FIRs in audio applications. Some of the purported negatives that are often quoted include
* FIRs sound "cold"
* linear phase FIRs cause pre-echos (because of their symmetric impulse response)
* minimum-phase FIRs exhibit excessive ringing in the impulse response
It must be said that these artefacts, if they can be noticed at all, are anyway extremely subtle and unlikely to compromise overall sound quality in a significant way. The major obstacle to the use of FIRs remains their inherent processing delay.
# The cookbook
In the following, we define a set of functions that return the five biquad coefficients for the most common types of audio filtering applications. Many of the formulas have been adapted from Robert Bristow-Johnson's famous [cookbook](https://webaudio.github.io/Audio-EQ-Cookbook/audio-eq-cookbook.html).
Each function returns ``b`` and ``a``, two arrays of three floats each containing the coefficients of the transfer function
$$
H(z) = \frac{b_0 + b_1 z^{-1} + b_{2}z^{-2}}{1 + a_1 z^{-1} + a_{2}z^{-2}} \qquad (a_0 = 1)
$$
## Lowpass
A second-order lowpass filter section will have a passband with approximately unit gain (0 dB) and a monotonically decreasing stopband. It is defined by two parameters:
1. the "quality factor" $Q$, which determines the shape of the magnitude response; by default $Q = \sqrt{1/2}$, which yields a Butterworth characteristic (i.e. a monotonically decreasing response).
1. the _corner frequency_ $f_c$ (also called the _cutoff_ frequency); the magnitude response will be equal to the quality factor $Q$ at $f_c$ and will decrease monotonically afterwards. For $Q = \sqrt{1/2}$, the attenuation at $f_c$ is equal to $20\log_{10}(\sqrt{1/2}) \approx -3$ dB, which yields a Butterworth (maximally flat) characteristic.
```
def LPF(fc, sf, Q=(1/np.sqrt(2))):
"""Biquad lowpass filter"""
w = 2 * np.pi * fc / sf
alpha = np.sin(w) / (2 * Q)
c = np.cos(w)
a = np.array([1 + alpha, -2 * c, 1 - alpha])
b = np.array([(1 - c) / 2, 1 - c, (1 - c) / 2])
return b / a[0], a / a[0]
if __name__ == '__main__':
CUTOFF = 1000
b, a = LPF(CUTOFF, DEFAULT_SF)
analog_response(b, a, DEFAULT_SF, dB=-50)
plt.axhline(y=-3, linewidth=0.5, color='r')
plt.axvline(x=CUTOFF, linewidth=0.5, color='r')
if __name__ == '__main__':
filter_props(b, a)
plt.gcf().get_axes()[0].axhline(y=np.sqrt(0.5), linewidth=0.5, color='r')
plt.gcf().get_axes()[0].axvline(x=(2 * np.pi * CUTOFF / DEFAULT_SF), linewidth=0.5, color='r')
```
When $Q = 1/\sqrt{2}$, as we said, the lowpass section corresponds to a Butterworth filter, that is, a filter with a maximally flat passband and a monotonically decreasing stopband. For higher $Q$ values the magnitude response exhibits a peak around $f_c$ which, in the time domain, corresponds to a damped oscillatory impulse response as shown in the following examples; for lower $Q$ values, the roll-off of the magnitude response will be less steep.
While these $Q$ values are clearly not a good choice for a single-stage lowpass, values other than $1/\sqrt{2}$ become useful when cascading multiple sections, as we will see later.
```
if __name__ == '__main__':
_, (fr, ir) = plt.subplots(2, figsize=(16,9))
CUTOFF = 100
Q = [0.1, 0.5, 1/np.sqrt(2), 5, 20]
for n, q in enumerate(Q):
b, a = LPF(CUTOFF, DEFAULT_SF, Q=q)
analog_response(b, a, DEFAULT_SF, dB=-50, axis=fr, color=f'C{n}')
ir.plot(signal.lfilter(b, a, np.r_[1, np.zeros(2000)]))
```
## Highpass
A highpass filter is simply the complementary filter to a lowpass, with the same roles for $f_c$ and $Q$.
```
def HPF(fc, sf, Q=(1/np.sqrt(2))):
"""Biquad highpass filter"""
w = 2 * np.pi * fc / sf
alpha = np.sin(w) / (2 * Q)
c = np.cos(w)
a = np.array([1 + alpha, -2 * c, 1 - alpha])
b = np.array([(1 + c) / 2, -1 - c, (1 + c) / 2])
return b / a[0], a / a[0]
if __name__ == '__main__':
CUTOFF = 2500
b, a = HPF(CUTOFF, DEFAULT_SF)
analog_response(b, a, DEFAULT_SF, dB=-50)
plt.axhline(y=-3, linewidth=0.5, color='r')
plt.axvline(x=CUTOFF, linewidth=0.5, color='r')
if __name__ == '__main__':
filter_props(b, a)
plt.gcf().get_axes()[0].axhline(y=np.sqrt(0.5), linewidth=0.5, color='r')
plt.gcf().get_axes()[0].axvline(x=(2 * np.pi * CUTOFF / DEFAULT_SF), linewidth=0.5, color='r')
```
## Bandpass
A second-order bandpass filter section will have approximately unit gain (0 dB) in the passband and will decrease monotonically to zero in the stopband. It is defined by two parameters:
1. the center frequency $f_c$, where the gain is unitary
1. the bandwidth $b = (f_+ - f_-)$, where $f_- < f_c < f_+$ are the first frequencies, left and right of $f_c$ where the attenuation reaches $-3$ dB. For the reasons explained above, note that the passband is almost but not exactly symmetric around $f_c$, with the asymmetry more pronounced towards the high end of the spectrum.
```
def BPF(fc, bw, sf):
"""Biquad bandpass filter"""
w = 2 * np.pi * fc / sf
alpha = np.tan(np. pi * bw / sf)
c = np.cos(w)
b = np.array([alpha, 0, -alpha])
a = np.array([1 + alpha, -2 * c, 1 - alpha])
return b / a[0], a / a[0]
if __name__ == '__main__':
CENTER, BANDWIDTH = 1000, 400
b, a = BPF(CENTER, BANDWIDTH, DEFAULT_SF)
analog_response(b, a, DEFAULT_SF, dB=-40)
plt.axhline(y=-3, linewidth=0.5, color='r')
plt.axvline(x=CENTER, linewidth=0.5, color='r')
plt.axvline(x=CENTER - BANDWIDTH / 2, linewidth=0.5, color='r')
plt.axvline(x=CENTER + BANDWIDTH / 2, linewidth=0.5, color='r')
if __name__ == '__main__':
filter_props(b, a)
plt.gcf().get_axes()[0].axhline(y=np.sqrt(0.5), linewidth=0.5, color='r')
plt.gcf().get_axes()[0].axvline(x=(2 * np.pi * CENTER / DEFAULT_SF), linewidth=0.5, color='r')
```
## Resonator
When the bandwith is very small, the second order bandpass becomes a constant-gain resonator:
```
if __name__ == '__main__':
_, ax = plt.subplots()
BANDWIDTH = 10
FC = [100, 1000, 2000, 4000, 6000]
for n, fc in enumerate(FC):
b, a = BPF(fc, BANDWIDTH, DEFAULT_SF)
frequency_response(b, a, dB=-50, half=True, axis=ax)
```
## Notch
A notch filter is the complementary filter to a resonator; its attenuation reaches $-\infty$ at $f_c$ and its bandwidth is usually kept very small in order to selectively remove only a given frequency; this is achieved by placing a pair of complex-conjugate zeros _on_ the unit circle and by placing two poles very close to the zeros.
```
def notch(fc, bw, sf):
"""Biquad notch filter"""
w = 2 * np.pi * fc / sf
alpha = np.tan(np. pi * bw / sf)
c = np.cos(w)
b = np.array([1, -2 * c, 1])
a = np.array([1 + alpha, -2 * c, 1 - alpha])
return b / a[0], a / a[0]
if __name__ == '__main__':
CENTER, BANDWIDTH = 2000, 100
b, a = notch(CENTER, BANDWIDTH, DEFAULT_SF)
analog_response(b, a, DEFAULT_SF, dB=-40)
plt.axhline(y=-6, linewidth=0.5, color='r')
plt.axvline(x=CENTER, linewidth=0.5, color='r')
if __name__ == '__main__':
filter_props(b, a)
plt.gcf().get_axes()[0].axhline(y=np.sqrt(0.5), linewidth=0.5, color='r')
plt.gcf().get_axes()[0].axvline(x=(2 * np.pi * CENTER / DEFAULT_SF), linewidth=0.5, color='r')
```
## Shelves
Shelving filters are used to amplify either the low or the high end of a signal's spectrum. A high shelf, for instanance, provides an arbitrary gain for high frequencies and has approximately unit gain in the low end of the spectrum. Shelving filters, high or low, are defined by the following parameters:
1. the desired _shelf gain_ in dB
1. the midpoint frequency $f_c$, which corresponds to the frequency in the transition band where the gain reaches half its value.
1. the "quality factor" $Q$, which determines the steepnes off the transition band; as for lowpass filters, the default value $Q = 1/\sqrt{2}$ yields the steepest transition band while avoiding resonances.
A common use case for shelving filters is in consumer audio appliances, where the standard "Bass" and "Treble" tone knobs control the gain of two complementary shelves with fixed midpoint frequency.
```
def LSH(fc, gain, sf, Q=(1/np.sqrt(2))):
"""Biquad low shelf"""
w = 2 * np.pi * fc / sf
A = 10 ** (gain / 40)
alpha = np.sin(w) / (2 * Q)
c = np.cos(w)
b = np.array([A * ((A + 1) - (A - 1) * c + 2 * np.sqrt(A) * alpha),
2 * A * ((A - 1) - (A + 1) * c),
A * ((A + 1) - (A - 1) * c - 2 * np.sqrt(A) * alpha)])
a = np.array([(A + 1) + (A - 1) * c + 2 * np.sqrt(A) * alpha,
-2 * ((A - 1) + (A + 1) * c),
(A + 1) + (A - 1) * c - 2 * np.sqrt(A) * alpha])
return b / a[0], a / a[0]
if __name__ == '__main__':
MIDPOINT, GAIN_DB = 200, 40
b, a = LSH(MIDPOINT, GAIN_DB, DEFAULT_SF)
analog_response(b, a, DEFAULT_SF, dB=-40)
plt.axhline(y=GAIN_DB / 2, linewidth=0.5, color='r')
plt.axvline(x=MIDPOINT, linewidth=0.5, color='r')
if __name__ == '__main__':
filter_props(b, a)
def HSH(fc, gain, sf, Q=(1/np.sqrt(2))):
"""Biquad high shelf"""
w = 2 * np.pi * fc / sf
A = 10 ** (gain / 40)
alpha = np.sin(w) / (2 * Q)
c = np.cos(w)
b = np.array([A * ((A + 1) + (A - 1) * c + 2 * np.sqrt(A) * alpha),
-2 * A * ((A - 1) + (A + 1) * c),
A * ((A + 1) + (A - 1) * c - 2 * np.sqrt(A) * alpha)])
a = np.array([(A + 1) - (A - 1) * c + 2 * np.sqrt(A) * alpha,
2 * ((A - 1) - (A + 1) * c),
(A + 1) - (A - 1) * c - 2 * np.sqrt(A) * alpha])
return b / a[0], a / a[0]
if __name__ == '__main__':
MIDPOINT, GAIN_DB = 2000, 40
b, a = HSH(MIDPOINT, GAIN_DB, DEFAULT_SF)
analog_response(b, a, DEFAULT_SF, dB=-40)
plt.axhline(y=GAIN_DB / 2, linewidth=0.5, color='r')
plt.axvline(x=MIDPOINT, linewidth=0.5, color='r')
if __name__ == '__main__':
filter_props(b, a)
```
## Peaking EQ
A peaking equalizer filter is the fundamental ingrediend in multiband parametric equalization. Each filter provides an arbitrary boost or attenuation for a given frequency band centered around a peak freqency and flattens to unit gain elsewhere. The filter is defined by the following parameters:
1. the desired gain in dB (which can be negative)
1. the peak frequency $f_c$, where the desired gain is attained
1. the bandwidth of the filter, defined as the interval around $f_c$ where the gain is greater (or smaller, for attenuators) than half the desired gain in dB; for instance, if the desired gain is 40dB, all frequencies within the filter's bandwidth will be boosted by at least 20dB. Note that the bandwdidth is not exactly symmetrical around $f_c$
```
def PEQ(fc, bw, gain, sf):
"""Biquad bandpass filter """
w = 2 * np.pi * fc / sf
A = 10 ** (gain / 40)
alpha = np.tan(np. pi * bw / sf)
c = np.cos(w)
b = np.array([1 + alpha * A, -2 * c, 1 - alpha * A])
a = np.array([1 + alpha / A, -2 * c, 1 - alpha / A])
return b / a[0], a / a[0]
if __name__ == '__main__':
CENTER, BW, GAIN_DB = 800, 400, 40
b, a = PEQ(CENTER, BW, GAIN_DB, DEFAULT_SF)
analog_response(b, a, DEFAULT_SF, dB=-40)
plt.axhline(y=GAIN_DB / 2, linewidth=0.5, color='r')
plt.axvline(x=CENTER, linewidth=0.5, color='r')
if __name__ == '__main__':
filter_props(b, a)
```
Note that peaking EQ filters with opposite gains are perfectly complementary:
```
if __name__ == '__main__':
CENTER, BW, GAIN_DB = 800, 400, 40
b, a = PEQ(CENTER, BW, GAIN_DB, DEFAULT_SF)
y = signal.lfilter(b, a, np.r_[1, np.zeros(200)])
plt.plot(y)
b, a = PEQ(CENTER, BW, -GAIN_DB, DEFAULT_SF)
y = signal.lfilter(b, a, y)
plt.plot(y)
```
# Cascades of biquads
The performance of a single biquad filter may not be adequate for a given application: a second-order lowpass filter, for instance, may not provide a sufficent amount of rejection in the stopband because of its rather slow roll-off characteristic; or we may want to design an equalizer with multiple peaks and dips. In all cases, we usually want to implement the final design as a cascade of biquad sections, because of their inherent numerical robustness.
## Factorization of higher-order filters
The first solution if a filter does not meet the requires specifications is to design a higher-order filter, possibly using different filter "recipes"; in the case of bandpass filters, for instance, we could try a Chebyshev or elliptic design. The resulting high-order transfer function can be then factored into a cascade of second-order sections (or, in the case of an odd-order filter, a cascade of second order-sections followed by a first-order filter):
$$
H(z) = \frac{b_0 + b_1 z^{-1} + \ldots + b_{N-1}z^{-N+1}}{a_0 + a_1 z^{-1} + \ldots + a_{N-1}z^{-N+1}} = \prod_{k=0}^{N/2} \frac{b_{k,0} + b_{k,1} z^{-1} + b_{k,2}z^{-2}}{1 + a_{k,1} z^{-1} + a_{k,2}z^{-2}}
$$
The biquad elements returned by the factorization are not related to the "cookbook" prototypes of the previous section and therefore this method is simply an implementation strategy that focuses on second-order structures; the design algorithm, in other words, will be dependent on the particular type of filter. Nevertheless, both the design and the factorization are usually available in numerical packages such as Scipy, for instance and, in the following example, we illustrate the difference between a 6th-order elliptic lowpass and a single second-order butterworth. First we will use the full high-order realization and then we will show how a cascade of three second-order sections implements the same characteristic.
Note that, when cascading transfer functions, the equivalent higher-order filter coefficients can be obtained simply by polynomial multiplication.
```
if __name__ == '__main__':
_, ax = plt.subplots()
CUTOFF = 1000
b, a = LPF(CUTOFF, DEFAULT_SF)
analog_response(b, a, DEFAULT_SF, dB=-60, axis=ax, color=f'C0')
eb, ea = signal.ellip(6, 1, 40, CUTOFF, fs=DEFAULT_SF)
analog_response(eb, ea, DEFAULT_SF, dB=-60, axis=ax, color=f'C3')
plt.axvline(x=CUTOFF, linewidth=0.5, color='r')
plt.axhline(y=-3, linewidth=0.5, color='r')
if __name__ == '__main__':
_, ax = plt.subplots()
CUTOFF = 1000
b, a = LPF(CUTOFF, DEFAULT_SF)
# this returns an array of second-order filter coefficients. Each row corresponds to a section,
# with the first three columns providing the numerator coefficients and the last three providing the denominator
soe = signal.ellip(6, 1, 40, CUTOFF, fs=DEFAULT_SF, output='sos')
cb, ca = [1], [1]
for n in range(0, 3):
b, a = soe[n][0:3], soe[n][3:6]
analog_response(b, a, DEFAULT_SF, dB=-60, axis=ax, color=f'C{n}:')
cb = np.polymul(b, cb)
ca = np.polymul(a, ca)
analog_response(cb, ca, DEFAULT_SF, dB=-60, axis=ax, color='C3')
```
## Cascading lowpass and highpass biquads
A cascade of $N$ identical sections with transfer function $H(z)$ will yield the overall transfer function $H_c(z) = H^N(z)$ and thus the stopband attenuation in decibels will increase $N$-fold. For instance, the following example shows the cumulative magnitude responses obtained by cascading up to five identical second-order Butterworth lowpass sections:
```
if __name__ == '__main__':
_, ax = plt.subplots()
CUTOFF = 1000
b, a = LPF(CUTOFF, DEFAULT_SF)
cb, ca = b, a
for n in range(0, 5):
analog_response(cb, ca, DEFAULT_SF, dB=-60, axis=ax, color=f'C{n}')
ca = np.polymul(a, ca)
cb = np.polymul(b, cb)
plt.axvline(x=CUTOFF, linewidth=0.5, color='r')
plt.axhline(y=-3, linewidth=0.5, color='r')
plt.axhline(y=-15, linewidth=0.5, color='r')
```
As shown by the previous plot, a cascade of identical maximally flat lowpass sections yields a steeper roll-off and preserves the monotonicity of the response. However, since the passband of each filter is not perfectly flat, the $-3~\mathrm{dB}$ cutoff frequency of the cascade becomes smaller with each added section and the effective bandwidth of the filter is reduced. In the previous example, the original $-3~\mathrm{dB}$ cutoff frequency was $f_c = 1000~\mathrm{Hz}$ but the magnitude response of the cascade at $f_c$ is $-15~\mathrm{dB}$ whereas the actual $-3~\mathrm{dB}$ point has shifted close to $600~\mathrm{Hz}$.
If our goal is to obtain a cascade with a maximally flat (Butterworth) response with a given $f_c$, an obvious approach is simply to factorize the transfer function of a high-order Butterworth as explained in the previous section. There is however a clever and simpler design strategy that is based on the geometric arrangement of the poles of an analog Butterworth filter of order $N$:
* the $N$ complex-conjugate poles are equally spaced along a circular contour centered on the origin of the $s$-plane
* the angle between poles is equal to $\pi/N$
With this, the pole angles in the upper $s$-plane are given by
$$
\theta_n = \frac{\pi}{2N} + n\frac{\pi}{N} = \frac{(2n+1)\pi}{2N}, \qquad n = 0, \ldots, N/2
$$
```
if __name__ == '__main__':
fig, sp = plt.subplots(1, 4, gridspec_kw={'wspace': 1})
for n in range(0, 4):
sp[n].plot(np.cos(np.linspace(0, 2 * np.pi, 100)), np.sin(np.linspace(0, 2 * np.pi, 100)), 'k:')
p = np.roots(signal.butter(2 * (n + 1), 1, analog=True)[1])
sp[n].plot(p.real, p.imag, 'C3x', ms=10, markeredgewidth=3.0)
sp[n].axis('square')
sp[n].set_xlim(-1.2, 1.2)
sp[n].set_ylim(-1.2, 1.2)
```
Now, a generic second-order analog filter will have a single pair of complex-conjugate poles at $p_{1,2} = \rho e^{\pm \theta}$ on the $s$-plane and, by cascading identical sections, we will only manage to increase the poles' multiplicity but we will not be able to change their position. In order to achieve a Butterworth pole configuration we will thus need to adjust the pole angle for each section; this is a simple task because it turns out that a second-order filter's quality factor $Q$ is related to the pole angle as
$$
1/Q = 2\cos \theta
$$
which means that we can choose the suitable pole angle for each section simply by setting $Q_n = 1/(2\cos \theta_n)$. We can now design $N/2$ discrete-time biquads with the same $Q_n$ values to obtain the desired result.
Below is the example for a cascade of five lowpass sections (i.e. a 10th-order filter) compared to a single biquad, both with cutoff $f_c = 1000~\mathrm{Hz}$; notice how the $-3~\mathrm{dB}$ point has not moved in spite of the much steeper rolloff.
```
if __name__ == '__main__':
_, ax = plt.subplots()
CUTOFF = 1000
b, a = LPF(CUTOFF, DEFAULT_SF)
analog_response(b, a, DEFAULT_SF, dB=-60, axis=ax, color='C0')
cb, ca, sections = [1], [1], 5
for n in range(0, sections):
iq = 2 * np.cos((2 * n + 1) * np.pi / (4 * sections))
b, a = LPF(CUTOFF, DEFAULT_SF, Q=1/iq)
ca = np.polymul(a, ca)
cb = np.polymul(b, cb)
analog_response(cb, ca, DEFAULT_SF, dB=-60, axis=ax, color='C1')
plt.axvline(x=CUTOFF, linewidth=0.5, color='r')
plt.axhline(y=-3, linewidth=0.5, color='r')
```
The resulting digital filter has its poles arranged on a circular contour centered in $z=1$ if the cutoff frequency is less than $\pi/2$ and centered on $z=-1$ otherwise.
```
if __name__ == '__main__':
filter_props(cb, ca, DEFAULT_SF, dB=-60)
```
Finally, the following plot shows the individual magnitude responses of the five sections. You can observe that the required $Q_n$ values lead to some biquad sections with a clear peak at the cutoff frequency, although the overall response is monotonic:
```
if __name__ == '__main__':
_, ax = plt.subplots()
CUTOFF = 1000
cb, ca, sections = [1], [1], 5
for n in range(0, sections):
iq = 2 * np.cos((2 * n + 1) * np.pi / (4 * sections))
b, a = LPF(CUTOFF, DEFAULT_SF, Q=1/iq)
analog_response(b, a, DEFAULT_SF, dB=-0, axis=ax, color=f'C{n+2}:')
ca = np.polymul(a, ca)
cb = np.polymul(b, cb)
analog_response(cb, ca, DEFAULT_SF, dB=-60, axis=ax, color='C1')
plt.axvline(x=CUTOFF, linewidth=0.5, color='r')
plt.axhline(y=-3, linewidth=0.5, color='r')
```
## Combining shelving filters
Shelving filters may be combined to create filters to boost a particular frequency range
```
if __name__ == '__main__':
cb, ca = LSH(1000, 20, DEFAULT_SF)
b, a = HSH(10, 20, DEFAULT_SF)
cb = np.polymul(b, cb)
ca = np.polymul(a, ca)
# normalize
analog_response(cb / 10, ca, DEFAULT_SF, dB=-50, points=10001)
```
## Parametric equalization
Peaking equalizers with distinct bandwidths can be cascaded to obtain an arbitrary equalization curve for the entire range of input frequencies; indeed, this is the technique behind so-called _parametric equalizers_ where a bank of logarithmically spaced peaking eq's with independent gain controls allow the user to easily define a global equalization response.
```
if __name__ == '__main__':
cb, ca = np.ones(1), np.ones(1)
for n, g in enumerate([20, -10, 40]):
b, a = PEQ(10 ** (n+1), 10 ** (n + 1), g, DEFAULT_SF)
cb = np.polymul(b, cb)
ca = np.polymul(a, ca)
analog_response(cb, ca, DEFAULT_SF, dB=-50, points=10001)
```
# Examples
# References
https://webaudio.github.io/Audio-EQ-Cookbook/audio-eq-cookbook.html
| github_jupyter |
<a href="https://colab.research.google.com/github/satyajitghana/TSAI-DeepVision-EVA4.0/blob/master/14_RCNN/01_DenseDepth_DatasetCreation.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Depth Project - Dataset Creation
```
from google.colab import drive
drive.mount('/content/gdrive')
cd gdrive/My\ Drive/DepthProject
!ls -l
```
## Note
I came up with the conclusion that writing the images directly to google drive is very slow, instead write them into a .zip file directly, which i did on my local machine, this notebook however shows the concept of how i had tried different methods
## This is the write the image to folder
### **PROCEED WITH ATMOST CAUTION**
```
!rm -r depth_dataset_cleaned/
```
### Count the number of processed folders, should be 100
```
!ls depth_dataset_cleaned/fg_bg/ | wc -l
```
## Count the Processed files on each folder, should be 4000 in each
```
! find ./depth_dataset_cleaned/fg_bg/ -type d | awk '{print "echo -n \""$0" \";ls -l "$0" | grep -v total | wc -l" }' | sh
!ls depth_dataset_cleaned/fg_bg/
```
## Unzip the Entire DATASET
This will take a long time, ~3-4 hours, instead use the partial dataset and then create own
```
!unzip -n depth_dataset_cleaned.zip
```
## Unzip Partial, Create the Dataset later
```
!unzip depth_dataset_cleaned_raw.zip -d depth_dataset_cleaned/
```
# Create the Dataset
```
import glob
import PIL
from PIL import Image
import os
import cv2
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from tqdm.auto import tqdm
from pathlib import Path
sns.set()
!ls depth_dataset_cleaned/
fgc_images = [f for f in glob.glob('depth_dataset_cleaned/fg/*.*')]
bgc_images = [f for f in glob.glob('depth_dataset_cleaned/bg/*.*')]
fgc_mask_images = [f for f in glob.glob('depth_dataset_cleaned/fg_mask/*.*')]
last_idx = 15
```
This was my attempt on creating the files direcctly into folders, this does work, however is a tedious process
```
idx = 0
for bidx, bg_image in enumerate(tqdm(bgc_images)):
if (bidx < last_idx):
continue
Path(f'depth_dataset_cleaned/labels/').mkdir(parents=True, exist_ok=True)
label_info = open(f"depth_dataset_cleaned/labels/bg_{bidx:03d}_label_info.txt","w+")
idx = 4000 * bidx
print(f'Processing BG {bidx}')
Path(f'depth_dataset_cleaned/fg_bg/bg_{bidx:03d}').mkdir(parents=True, exist_ok=True)
Path(f'depth_dataset_cleaned/fg_bg_mask/bg_{bidx:03d}').mkdir(parents=True, exist_ok=True)
for fidx, fg_image in enumerate(tqdm(fgc_images)):
# do the add fg to bg 20 times
for i in range(20):
# do this twice, one with flip once without
for should_flip in [True, False]:
background = Image.open(bg_image)
foreground = Image.open(fg_image)
fg_mask = Image.open(fgc_mask_images[fidx])
if should_flip:
foreground = foreground.transpose(PIL.Image.FLIP_LEFT_RIGHT)
fg_mask = fg_mask.transpose(PIL.Image.FLIP_LEFT_RIGHT)
b_width, b_height = background.size
f_width, f_height = foreground.size
max_y = b_height - f_height
max_x = b_width - f_width
pos_x = np.random.randint(low=0, high=max_x, size=1)[0]
pos_y = np.random.randint(low=0, high=max_y, size=1)[0]
background.paste(foreground, (pos_x, pos_y), foreground)
mask_bg = Image.new('L', background.size)
fg_mask = fg_mask.convert('L')
mask_bg.paste(fg_mask, (pos_x, pos_y), fg_mask)
background.save(f'depth_dataset_cleaned/fg_bg/bg_{bidx:03d}/fg_{fidx:03d}_bg_{bidx:03d}_{idx:06d}.jpg', optimize=True, quality=30)
mask_bg.save(f'depth_dataset_cleaned/fg_bg_mask/bg_{bidx:03d}/fg_{fidx:03d}_bg_{bidx:03d}_mask_{idx:06d}.jpg', optimize=True, quality=30)
label_info.write(f'fg_bg/bg_{bidx:03d}/fg_{fidx:03d}_bg_{bidx:03d}_{idx:06d}.jpg\tfg_bg_mask/bg_{bidx:03d}/fg_{fidx:03d}_bg_{bidx:03d}_mask_{idx:06d}.jpg\t{pos_x}\t{pos_y}\n')
idx = idx + 1
label_info.close()
last_idx = bidx
```
## This is how i created the .zip file
the output is removed, because the actual run was made on local machine
```
idx = 0
# for each background image
for bidx, bg_image in enumerate(tqdm(bgc_images)):
# output zip file, open in append mode
out_zip = ZipFile('fg_bg.zip', mode='a', compression=zipfile.ZIP_STORED)
# labels for the craeted images
label_info = open(f't_label.txt', 'w+')
idx = 4000 * bidx
print(f'Processing BG {bidx}')
for fidx, fg_image in enumerate(tqdm(fgc_images)):
# do the add fg to bg 20 times
for i in range(20):
# do this twice, one with flip once without
for should_flip in [True, False]:
# open the bg and fg images
background = Image.open(bg_image)
foreground = Image.open(fg_image)
fg_mask = Image.open(fgc_mask_images[fidx])
# if the fg image should be flipped
if should_flip:
foreground = foreground.transpose(PIL.Image.FLIP_LEFT_RIGHT)
fg_mask = fg_mask.transpose(PIL.Image.FLIP_LEFT_RIGHT)
# choose a random point on the bg to paste the fg image
b_width, b_height = background.size
f_width, f_height = foreground.size
max_y = b_height - f_height
max_x = b_width - f_width
pos_x = np.random.randint(low=0, high=max_x, size=1)[0]
pos_y = np.random.randint(low=0, high=max_y, size=1)[0]
background.paste(foreground, (pos_x, pos_y), foreground)
mask_bg = Image.new('L', background.size)
fg_mask = fg_mask.convert('L')
mask_bg.paste(fg_mask, (pos_x, pos_y), fg_mask)
label_info.write(f'fg_bg/bg_{bidx:03d}/fg_{fidx:03d}_bg_{bidx:03d}_{idx:06d}.jpg\tfg_bg_mask/bg_{bidx:03d}/fg_{fidx:03d}_bg_{bidx:03d}_mask_{idx:06d}.jpg\t{pos_x}\t{pos_y}\n')
# save the background and the mask as temp .jpg files
background.save('b_temp.jpg', optimize=True, quality=30)
mask_bg.save('m_temp.jpg', optimize=True, quality=30)
# save the files to .zip file
out_zip.write('b_temp.jpg', f'depth_dataset_cleaned/fg_bg/bg_{bidx:03d}/fg_{fidx:03d}_bg_{bidx:03d}_{idx:06d}.jpg')
out_zip.write('m_temp.jpg', f'depth_dataset_cleaned/fg_bg_mask/bg_{bidx:03d}/fg_{fidx:03d}_bg_{bidx:03d}_mask_{idx:06d}.jpg')
idx = idx + 1
label_info.close()
# write the labels file to zip
out_zip.write('t_labels.txt', f'depth_dataset_cleaned/labels/bg_{bidx:03d}_label_info.txt')
# important: close the zip, else it gets corrupted
out_zip.close()
```
| github_jupyter |
# The Pipeline
1. Question & required data
2. Acquire the data
3. Data Analysis
4. Prepare the data for the Deep Learning Model
5. Building & training the model
6. Testing the model
7. Interpreting the model
8. Improving the model
# Problem introduction: predicting whether a bank customer will leave the bank
## Question & required data
The dataset consists of a bunch of features describing the customers of a bank. The question that is posed is that, given these features could we detect some patterns that allow us to predict if a customer is about to leave a bank?
It would then be possible to direct marketing efforts to the customers if it would be beneficial to retain them.
### ML problem characteristics
1. Task: **classification**
2. Attribute Types: **Categorical, continuous and binary**
3. **14** attributes
4. **10k** instances
5. ML model: **Deep neural net**
6. Deep learning library: **Keras (Tensorflow)**
7. Hyperparameter tuning: **Talos**
#### The feature vector
1. *RowNumber*
2. *CustomerId:* unique id to identify a customer
3. *Surname*
4. *CreditScore:* a creditscore compiled by the bank for each customer
5. *Geography*
6. *Gender*
7. *Age*
8. *Tenure:* how long has the customer been with the bank
9. *Balance*
10. *NumOfProducts:* how many products of the bank (credit/debit cards etc.) does the customer have
11. *HasCrCard:* does the customer have a credit card
12. *IsActiveMember:* is the customer an active member of the bank
13. *EstimatedSalary:* the customer's estimated salary
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from collections import Counter
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.metrics import accuracy_score
import tensorflow as tf
from tensorflow import keras
from keras.models import Sequential
from keras.layers import Dropout, Dense
import talos
sns.set()
tf.__version__
data = pd.read_csv('dataset/exit_prediction.csv')
print('Our dataset holds {} records and {} attributes'.format(*data.shape))
data.head()
```
*RowNumber*, *CustomerId* & *Surname* won't actually be useful for the deep learning model that we will build in the end so we can just go ahead and drop them.
```
data.drop(labels=['RowNumber', 'CustomerId', 'Surname'], axis=1, inplace=True)
```
## Data Analysis
1. [Tidy](https://cran.r-project.org/web/packages/tidyr/vignettes/tidy-data.html) Data: by default
2. **Acquaintance with the data**
3. Outliers
4. Missing Data
5. Class-Imbalance problem
6. Feature correlation
### Acquaintance with the data
Lets visualize a subset of the feature vector for this purpose
```
plt.figure(figsize=(15,13))
selected_features = data[['CreditScore', 'Geography', 'Age', 'Tenure', 'Balance', 'NumOfProducts',
'EstimatedSalary', 'HasCrCard', 'Exited']]
for i in range(1, selected_features.shape[1]+1):
plt.subplot(3,3,i)
figure=plt.gca()
figure.set_title(selected_features.columns[i-1])
plt.hist(selected_features.iloc[:, i-1])
if selected_features.columns[i-1] in ['EstimatedSalary', 'HasCrCard', 'Exited']:
plt.xlabel('Measure')
if selected_features.columns[i-1] in ['CreditScore', 'Tenure', 'EstimatedSalary']:
plt.ylabel('Count')
plt.tight_layout(pad=2)
```
### Outliers
It appears that we have no outliers. This observation is also confirmed by the above plot
```
data.describe()
```
### Missing Data
Our dataset seems to have no missing values. We cannot call the values of the *Balance = 0* as missing values since this can be a value that the balance can indeed take.
For an approach of taking care of missing values see the other project on Autism Predictor
```
print('Does our dataset contain missing values?', data.isnull().any().any())
```
### Class-Imbalance problem
Our data seems indeed to have imbalanced classes. Obviously, most of the people did not leave the bank.
```
print(data['Exited'].value_counts()/data.shape[0])
sns.countplot(x='Exited', data=data, palette='hls'); plt.show()
```
**Are the classes well separated?**
If our classes are well separated, then the class-imbalance problem is not really a problem. We can simply visualize a couple of the attributes for this purpose.
```
fig, ((ax1, ax2, ax3)) = plt.subplots(nrows=1, ncols=3, figsize=(15,5))
att1, att2, att3 = np.random.choice(data.columns.delete([13]), 3, replace=False)
att1_data = list(data[att1].groupby(data['Exited']))
ax1.plot(att1_data[0][1], 'g^', label=att1_data[0][0])
ax1.plot(att1_data[1][1], 'ro', label=att1_data[1][0])
ax1.set_xlabel('Count'); ax1.set_ylabel('Measure'); ax1.set_title(att1_data[0][1].name)
ax1.legend()
att2_data = list(data[att2].groupby(data['Exited']))
ax2.plot(att2_data[0][1], 'g^', label=att2_data[0][0])
ax2.plot(att2_data[1][1], 'ro', label=att2_data[1][0])
ax2.set_xlabel('Count'); ax2.set_ylabel(''); ax2.set_title(att2_data[0][1].name)
ax2.legend()
att3_data = list(data[att3].groupby(data['Exited']))
ax3.plot(att3_data[1][1], 'ro', label=att3_data[1][0])
ax3.plot(att3_data[0][1], 'g^', label=att3_data[0][0])
ax3.set_xlabel('Count'); ax3.set_ylabel(''); ax3.set_title(att3_data[0][1].name)
ax3.legend(); plt.show()
```
It seems that our classes are not well separated. There are techniques to fight the class-imbalance problem but in this case we are just going to continue like this.
### One-Hot Encoding
Convert categorical data to numerical. We have two categorical features in our case: *geography* & *gender*
```
data = pd.get_dummies(data)
data.head(2)
```
### Feature correlation
We want the features to be independent of each-other. Most ML algorithms work on this assumption.
#### Correlation of features with the class label
This will also give us an impression into which variable is the more important for predicting the class variable.
None of the attributes seems to be significantly correlated with the class.
```
data.drop('Exited', axis=1).corrwith(data['Exited']).plot.bar(figsize=(12,5),
title='Correlation with the class variable', rot=45, fontsize=11)
plt.show()
```
We can also check the influence of the attributes on the class by aggregating the mean values after grouping by the class label. The below result confirms the correlation plot above.
```
class_group = data.groupby('Exited')
print(class_group.agg(np.mean))
```
#### Correlation of features with each-other
```
sns.set(style='white', font_scale=1.5)
corr_matrix = data.corr()
# create a mask for the upper triangle so that we can ignore it when later building the heatmap
mask = np.zeros_like(corr_matrix, dtype=np.bool)
mask[np.triu_indices_from(mask)] = True
# color map for the different values of the correlation matrix
cmap = sns.diverging_palette(220, 10, as_cmap=True)
plt.figure(figsize=(11,8))
plt.title('Correlation matrix for Features')
sns.heatmap(corr_matrix, square=True, mask=mask, cmap=cmap, center=0, linewidths=1.0, cbar_kws={'shrink':0.8})
plt.show()
```
We can see some correlations, like for instance the negative correlation between the number of products and the balance but they are less than 0.3 and so do not pose any problems for the algorithm. We may further proceed like this.
## Prepare the data for the Neural Net Model
1. Class & Features separation
2. Train/Test split
3. Scaling/Standardization if needed
### Class & Feature separation
```
labels = data.pop('Exited')
features = data
print('Shape of features', features.shape)
print('Shape of labels', labels.shape)
features.head(2)
```
### Train/Test split
```
train_features, test_features, train_labels, test_labels = train_test_split(features, labels, test_size=0.3,
random_state=1, stratify=labels)
print('Shape of training data', train_features.shape)
print('Shape of testing data', test_features.shape)
```
### Feature Scaling
```
scaler = StandardScaler()
to_scale = ['CreditScore', 'Age', 'Balance', 'EstimatedSalary']
train_features_sc = pd.DataFrame(scaler.fit_transform(train_features[to_scale]))
train_features_sc.columns = train_features[to_scale].columns
train_features_sc.index = train_features[to_scale].index.values
train_features[to_scale] = train_features_sc
test_features_sc = pd.DataFrame(scaler.fit_transform(test_features[to_scale]))
test_features_sc.columns = test_features[to_scale].columns
test_features_sc.index = test_features[to_scale].index.values
test_features[to_scale] = test_features_sc
```
## Build & train the Neural Net model
We shall first build a basic Neural Net model using Keras. On the next chapter we shall see how to tune its parameters using talos in a bid to improve the model's performance.
### Build the Deep Neural Net
We can instantiate different aspects of the neural net either by passing a string identifier (and in this case the default parameters for the optimizer can be used) or by building an object from the class. The latter should be the used way as it gives us the possibility to tweak stuff.
[Docs](https://keras.io/api/layers/core_layers/dense/) on the dense layers
[Docs](https://keras.io/api/models/model_training_apis/) for the compile method
[Docs](https://keras.io/api/optimizers/) on optimizers
[Docs](https://keras.io/api/losses/) on the loss function
- *from_logits=True*: will significantly boost testing accuracy at a cost of training
- *label_smoothing=1*: can be played with in the range of [0,1]
- *reduction*: different options include, 'sum', keras.losses.Reduction.NONE
[Docs](https://keras.io/api/models/model_training_apis/) on the fit method
1. *class_weight*: can be useful to tell the model to "pay more attention" to samples from an under-represented class
```
ann = Sequential()
ann.add(Dense(units=100, activation='relu', use_bias=True))
ann.add(Dense(units=64, activation='relu', use_bias=True))
ann.add(Dense(units=32, activation='relu', use_bias=True))
ann.add(Dense(units=32, activation='relu', use_bias=True))
# Use sigmoid in the final layer since we have a binary classification problem
ann.add(Dense(units=1, activation='sigmoid', use_bias=False))
optimizer = keras.optimizers.Adam(learning_rate=0.001, beta_1=0.9, beta_2=0.999)
# binary cross entropy as loss since we have a binary classification problem
loss = keras.losses.BinaryCrossentropy(name='binary_crossentropy', from_logits=False,
label_smoothing=0.05, reduction='auto')
ann.compile(optimizer=optimizer, loss=loss, metrics=['accuracy'])
def get_class_weights(labels):
counter = Counter(labels)
majority = max(counter.values())
return {cls: float(majority/count) for cls, count in counter.items()}
get_class_weights(labels)
c_weight = {0:1, 1:4}
ann.fit(x=train_features.values, y=train_labels.values, epochs=100, verbose=2, shuffle=True, batch_size=70,
use_multiprocessing=True, )
```
## Test the Model
```
print('Shape of testing dataset', test_features.shape)
predictions = ((ann.predict(test_features, batch_size=70)) > 0.5)
print('Our models accuracy on test data: ', round(100*accuracy_score(test_labels, predictions), 2))
```
## Interpret the Model
Our Deep Neural Network performed good. It achieved an ~83% accuracy on the testing data. This was achieved only by utilizing a few hyperparameters of the neural network.
We shall next see if we can improve this number by tuning the various hyperparameters that a neural network can take. We shall do this using talos.
| github_jupyter |
# **MobileNetV2-Alpaca-Classifier**
Alpaca Classifier Using Transfer Learning with MobileNetV2 trained on ImageNet
Github Repo : [MobileNetV2-Alpaca-Classifier](https://github.com/sid4sal/MobileNetV2-Alpaca-Classifier)
### 1. Import Packages
Import the required packages
```
import matplotlib.pyplot as plt
import numpy as np
import os
import tensorflow as tf
import tensorflow.keras.layers as tfl
from tensorflow.keras.preprocessing import image_dataset_from_directory
from tensorflow.keras.layers.experimental.preprocessing import RandomFlip, RandomRotation
```
### 2. Load Dataset
Dataset used is uploaded on kaggle : [Download Dataset](https://www.kaggle.com/sid4sal/alpaca-dataset-small)
Dataset contains JPG Images of Alpaca and Not Alpaca (Images not containing alpaca but has subjects similar to alpaca)
```
from google.colab import drive
drive.mount('/content/drive')
BATCH_SIZE = 32
IMG_SIZE = (160, 160)
directory = '/content/drive/My Drive/MobileNetV2-Alpaca-Classifier/dataset/'
train_dataset = image_dataset_from_directory(directory,
shuffle=True,
batch_size=BATCH_SIZE,
image_size=IMG_SIZE,
validation_split=0.2,
subset='training',
seed=42)
validation_dataset = image_dataset_from_directory(directory,
shuffle=True,
batch_size=BATCH_SIZE,
image_size=IMG_SIZE,
validation_split=0.2,
subset='validation',
seed=42)
```
### 3. Augment and Preprocess the Data
Our dataset is small, even for transfer learning, so data augmentation is needed.
```
AUTOTUNE = tf.data.experimental.AUTOTUNE
train_dataset = train_dataset.prefetch(buffer_size=AUTOTUNE)
def data_augmenter():
data_augmentation = tf.keras.Sequential()
data_augmentation.add(RandomFlip('horizontal'))
data_augmentation.add(RandomRotation(0.2))
return data_augmentation
preprocess_input = tf.keras.applications.mobilenet_v2.preprocess_input
```
### 4. Build and Train the Model
#### Build the Model
Define the alpaca_model and use the MobileNetV2 as base model with ImageNet weights (i.e trained on ImageNet Dataset)
Freeze all the layers before the `fine_tune_at` layer.
```
def alpaca_model(image_shape=IMG_SIZE, data_augmentation=data_augmenter(), fine_tune_at=120):
input_shape = image_shape + (3,)
base_model = tf.keras.applications.MobileNetV2(input_shape=input_shape,
include_top=False,
weights='imagenet')
base_model.trainable = True
for layer in base_model.layers[:fine_tune_at]:
layer.trainable = False
inputs = tf.keras.Input(shape=input_shape)
x = data_augmentation(inputs)
x = preprocess_input(x)
x = base_model(x, training=False)
x = tfl.GlobalAveragePooling2D()(x)
x = tfl.Dropout(0.2)(x)
prediction_layer = tfl.Dense(1)
outputs = prediction_layer(x)
model = tf.keras.Model(inputs, outputs)
return model
```
#### Set the Hyperparameters
There are 3 important parameters:-
* learning_rate
* epochs
* fine_tune_at :
Train (unfreeze) all the layers after this layer, i.e freeze all te layers before this layer.
Get the number of layers in the base model (MobileNetV2) excluding softmax layer, for setting the 'fine_tune_at' paramater
```
base_model = tf.keras.applications.MobileNetV2(input_shape=IMG_SIZE + (3,),
include_top=False,
weights='imagenet')
print("Number of layers in the base model: ", len(base_model.layers))
learning_rate = 0.0001
fine_tune_at = 120
epochs = 8
data_augmentation = data_augmenter()
```
#### Compile and Train the Model
```
model = alpaca_model(IMG_SIZE, data_augmentation, fine_tune_at)
model.compile(optimizer=tf.keras.optimizers.Adam(learning_rate=learning_rate),
loss=tf.keras.losses.BinaryCrossentropy(from_logits=True),
metrics=['accuracy'])
model_train = model.fit(train_dataset, validation_data=validation_dataset, epochs=epochs)
```
### 5. Plot the Accuracy and Loss
```
acc = [0.] + model_train.history['accuracy']
val_acc = [0.] + model_train.history['val_accuracy']
loss = model_train.history['loss']
val_loss = model_train.history['val_loss']
plt.figure(figsize=(8, 8))
plt.subplot(2, 1, 1)
plt.plot(acc, label='Training Accuracy')
plt.plot(val_acc, label='Validation Accuracy')
plt.legend(loc='lower right')
plt.ylabel('Accuracy')
plt.ylim([min(plt.ylim()),1])
plt.title('Training and Validation Accuracy')
plt.subplot(2, 1, 2)
plt.plot(loss, label='Training Loss')
plt.plot(val_loss, label='Validation Loss')
plt.legend(loc='upper right')
plt.ylabel('Cross Entropy')
plt.ylim([0,1.0])
plt.title('Training and Validation Loss')
plt.xlabel('epoch')
plt.show()
```
### 6. Predictions
Test the predictions made by the Alpaca Model on validation set.
```
class_names = validation_dataset.class_names
plt.figure(figsize=(10, 10))
for images, labels in validation_dataset.take(1):
image_var = tf.Variable(images)
pred = model.predict(images)
for i in range(9):
ax = plt.subplot(3, 3, i + 1)
plt.imshow(images[i].numpy().astype("uint8"))
p = "alpaca" if pred[i] < 0 else "not alpaca"
plt.title("pred:{} label:{}".format(p,class_names[labels[i]]))
plt.axis("off")
```
| github_jupyter |
```
# -*- coding: utf-8 -*-
"""
EVCで変換する.
詳細 : https://pdfs.semanticscholar.org/cbfe/71798ded05fb8bf8674580aabf534c4dbb8bc.pdf
Converting by EVC.
Check detail : https://pdfs.semanticscholar.org/cbfe/71798ded05fb8bf8674580abf534c4dbb8bc.pdf
"""
from __future__ import division, print_function
import os
from shutil import rmtree
import argparse
import glob
import pickle
import time
import numpy as np
from numpy.linalg import norm
from sklearn.decomposition import PCA
from sklearn.mixture import GMM # sklearn 0.20.0から使えない
from sklearn.preprocessing import StandardScaler
import scipy.signal
import scipy.sparse
%matplotlib inline
import matplotlib.pyplot as plt
import IPython
from IPython.display import Audio
import soundfile as sf
import wave
import pyworld as pw
import librosa.display
from dtw import dtw
import warnings
warnings.filterwarnings('ignore')
"""
Parameters
__Mixtured : GMM混合数
__versions : 実験セット
__convert_source : 変換元話者のパス
__convert_target : 変換先話者のパス
"""
# parameters
__Mixtured = 40
__versions = 'pre-stored0.1.3'
__convert_source = 'input/EJM10/V01/T01/TIMIT/000/*.wav'
__convert_target = 'adaptation/EJM05/V01/T01/ATR503/A/*.wav'
# settings
__same_path = './utterance/' + __versions + '/'
__output_path = __same_path + 'output/EJM05/' # EJF01, EJF07, EJM04, EJM05
Mixtured = __Mixtured
pre_stored_pickle = __same_path + __versions + '.pickle'
pre_stored_source_list = __same_path + 'pre-source/**/V01/T01/**/*.wav'
pre_stored_list = __same_path + "pre/**/V01/T01/**/*.wav"
#pre_stored_target_list = "" (not yet)
pre_stored_gmm_init_pickle = __same_path + __versions + '_init-gmm.pickle'
pre_stored_sv_npy = __same_path + __versions + '_sv.npy'
save_for_evgmm_covarXX = __output_path + __versions + '_covarXX.npy'
save_for_evgmm_covarYX = __output_path + __versions + '_covarYX.npy'
save_for_evgmm_fitted_source = __output_path + __versions + '_fitted_source.npy'
save_for_evgmm_fitted_target = __output_path + __versions + '_fitted_target.npy'
save_for_evgmm_weights = __output_path + __versions + '_weights.npy'
save_for_evgmm_source_means = __output_path + __versions + '_source_means.npy'
for_convert_source = __same_path + __convert_source
for_convert_target = __same_path + __convert_target
converted_voice_npy = __output_path + 'sp_converted_' + __versions
converted_voice_wav = __output_path + 'sp_converted_' + __versions
mfcc_save_fig_png = __output_path + 'mfcc3dim_' + __versions
f0_save_fig_png = __output_path + 'f0_converted' + __versions
converted_voice_with_f0_wav = __output_path + 'sp_f0_converted' + __versions
EPSILON = 1e-8
class MFCC:
"""
MFCC() : メル周波数ケプストラム係数(MFCC)を求めたり、MFCCからスペクトルに変換したりするクラス.
動的特徴量(delta)が実装途中.
ref : http://aidiary.hatenablog.com/entry/20120225/1330179868
"""
def __init__(self, frequency, nfft=1026, dimension=24, channels=24):
"""
各種パラメータのセット
nfft : FFTのサンプル点数
frequency : サンプリング周波数
dimension : MFCC次元数
channles : メルフィルタバンクのチャンネル数(dimensionに依存)
fscale : 周波数スケール軸
filterbankl, fcenters : フィルタバンク行列, フィルタバンクの頂点(?)
"""
self.nfft = nfft
self.frequency = frequency
self.dimension = dimension
self.channels = channels
self.fscale = np.fft.fftfreq(self.nfft, d = 1.0 / self.frequency)[: int(self.nfft / 2)]
self.filterbank, self.fcenters = self.melFilterBank()
def hz2mel(self, f):
"""
周波数からメル周波数に変換
"""
return 1127.01048 * np.log(f / 700.0 + 1.0)
def mel2hz(self, m):
"""
メル周波数から周波数に変換
"""
return 700.0 * (np.exp(m / 1127.01048) - 1.0)
def melFilterBank(self):
"""
メルフィルタバンクを生成する
"""
fmax = self.frequency / 2
melmax = self.hz2mel(fmax)
nmax = int(self.nfft / 2)
df = self.frequency / self.nfft
dmel = melmax / (self.channels + 1)
melcenters = np.arange(1, self.channels + 1) * dmel
fcenters = self.mel2hz(melcenters)
indexcenter = np.round(fcenters / df)
indexstart = np.hstack(([0], indexcenter[0:self.channels - 1]))
indexstop = np.hstack((indexcenter[1:self.channels], [nmax]))
filterbank = np.zeros((self.channels, nmax))
for c in np.arange(0, self.channels):
increment = 1.0 / (indexcenter[c] - indexstart[c])
# np,int_ は np.arangeが[0. 1. 2. ..]となるのをintにする
for i in np.int_(np.arange(indexstart[c], indexcenter[c])):
filterbank[c, i] = (i - indexstart[c]) * increment
decrement = 1.0 / (indexstop[c] - indexcenter[c])
# np,int_ は np.arangeが[0. 1. 2. ..]となるのをintにする
for i in np.int_(np.arange(indexcenter[c], indexstop[c])):
filterbank[c, i] = 1.0 - ((i - indexcenter[c]) * decrement)
return filterbank, fcenters
def mfcc(self, spectrum):
"""
スペクトルからMFCCを求める.
"""
mspec = []
mspec = np.log10(np.dot(spectrum, self.filterbank.T))
mspec = np.array(mspec)
return scipy.fftpack.realtransforms.dct(mspec, type=2, norm="ortho", axis=-1)
def delta(self, mfcc):
"""
MFCCから動的特徴量を求める.
現在は,求める特徴量フレームtをt-1とt+1の平均としている.
"""
mfcc = np.concatenate([
[mfcc[0]],
mfcc,
[mfcc[-1]]
]) # 最初のフレームを最初に、最後のフレームを最後に付け足す
delta = None
for i in range(1, mfcc.shape[0] - 1):
slope = (mfcc[i+1] - mfcc[i-1]) / 2
if delta is None:
delta = slope
else:
delta = np.vstack([delta, slope])
return delta
def imfcc(self, mfcc, spectrogram):
"""
MFCCからスペクトルを求める.
"""
im_sp = np.array([])
for i in range(mfcc.shape[0]):
mfcc_s = np.hstack([mfcc[i], [0] * (self.channels - self.dimension)])
mspectrum = scipy.fftpack.idct(mfcc_s, norm='ortho')
# splrep はスプライン補間のための補間関数を求める
tck = scipy.interpolate.splrep(self.fcenters, np.power(10, mspectrum))
# splev は指定座標での補間値を求める
im_spectrogram = scipy.interpolate.splev(self.fscale, tck)
im_sp = np.concatenate((im_sp, im_spectrogram), axis=0)
return im_sp.reshape(spectrogram.shape)
def trim_zeros_frames(x, eps=1e-7):
"""
無音区間を取り除く.
"""
T, D = x.shape
s = np.sum(np.abs(x), axis=1)
s[s < 1e-7] = 0.
return x[s > eps]
def analyse_by_world_with_harverst(x, fs):
"""
WORLD音声分析合成器で基本周波数F0,スペクトル包絡,非周期成分を求める.
基本周波数F0についてはharvest法により,より精度良く求める.
"""
# 4 Harvest with F0 refinement (using Stonemask)
frame_period = 5
_f0_h, t_h = pw.harvest(x, fs, frame_period=frame_period)
f0_h = pw.stonemask(x, _f0_h, t_h, fs)
sp_h = pw.cheaptrick(x, f0_h, t_h, fs)
ap_h = pw.d4c(x, f0_h, t_h, fs)
return f0_h, sp_h, ap_h
def wavread(file):
"""
wavファイルから音声トラックとサンプリング周波数を抽出する.
"""
wf = wave.open(file, "r")
fs = wf.getframerate()
x = wf.readframes(wf.getnframes())
x = np.frombuffer(x, dtype= "int16") / 32768.0
wf.close()
return x, float(fs)
def preEmphasis(signal, p=0.97):
"""
MFCC抽出のための高域強調フィルタ.
波形を通すことで,高域成分が強調される.
"""
return scipy.signal.lfilter([1.0, -p], 1, signal)
def alignment(source, target, path):
"""
タイムアライメントを取る.
target音声をsource音声の長さに合うように調整する.
"""
# ここでは814に合わせよう(targetに合わせる)
# p_p = 0 if source.shape[0] > target.shape[0] else 1
#shapes = source.shape if source.shape[0] > target.shape[0] else target.shape
shapes = source.shape
align = np.array([])
for (i, p) in enumerate(path[0]):
if i != 0:
if j != p:
temp = np.array(target[path[1][i]])
align = np.concatenate((align, temp), axis=0)
else:
temp = np.array(target[path[1][i]])
align = np.concatenate((align, temp), axis=0)
j = p
return align.reshape(shapes)
covarXX = np.load(save_for_evgmm_covarXX)
covarYX = np.load(save_for_evgmm_covarYX)
fitted_source = np.load(save_for_evgmm_fitted_source)
fitted_target = np.load(save_for_evgmm_fitted_target)
weights = np.load(save_for_evgmm_weights)
source_means = np.load(save_for_evgmm_source_means)
"""
声質変換に用いる変換元音声と目標音声を読み込む.
"""
timer_start = time.time()
source_mfcc_for_convert = []
source_sp_for_convert = []
source_f0_for_convert = []
source_ap_for_convert = []
fs_source = None
for name in sorted(glob.iglob(for_convert_source, recursive=True)):
print("source = ", name)
x_source, fs_source = sf.read(name)
f0_source, sp_source, ap_source = analyse_by_world_with_harverst(x_source, fs_source)
mfcc_source = MFCC(fs_source)
#mfcc_s_tmp = mfcc_s.mfcc(sp)
#source_mfcc_for_convert = np.hstack([mfcc_s_tmp, mfcc_s.delta(mfcc_s_tmp)])
source_mfcc_for_convert.append(mfcc_source.mfcc(sp_source))
source_sp_for_convert.append(sp_source)
source_f0_for_convert.append(f0_source)
source_ap_for_convert.append(ap_source)
target_mfcc_for_fit = []
target_f0_for_fit = []
target_ap_for_fit = []
for name in sorted(glob.iglob(for_convert_target, recursive=True)):
print("target = ", name)
x_target, fs_target = sf.read(name)
f0_target, sp_target, ap_target = analyse_by_world_with_harverst(x_target, fs_target)
mfcc_target = MFCC(fs_target)
#mfcc_target_tmp = mfcc_target.mfcc(sp_target)
#target_mfcc_for_fit = np.hstack([mfcc_t_tmp, mfcc_t.delta(mfcc_t_tmp)])
target_mfcc_for_fit.append(mfcc_target.mfcc(sp_target))
target_f0_for_fit.append(f0_target)
target_ap_for_fit.append(ap_target)
# 全部numpy.arrrayにしておく
source_data_mfcc = np.array(source_mfcc_for_convert)
source_data_sp = np.array(source_sp_for_convert)
source_data_f0 = np.array(source_f0_for_convert)
source_data_ap = np.array(source_ap_for_convert)
target_mfcc = np.array(target_mfcc_for_fit)
target_f0 = np.array(target_f0_for_fit)
target_ap = np.array(target_ap_for_fit)
print("Load Input and Target Voice time = ", time.time() - timer_start , "[sec]")
def convert(source, covarXX, fitted_source, fitted_target, covarYX, weights, source_means):
"""
声質変換を行う.
"""
Mixtured = 40
D = source.shape[0]
E = np.zeros((Mixtured, D))
for m in range(Mixtured):
xx = np.linalg.solve(covarXX[m], source - fitted_source[m])
E[m] = fitted_target[m] + np.dot(covarYX[m], xx)
px = GMM(n_components = Mixtured, covariance_type = 'full')
px.weights_ = weights
px.means_ = source_means
px.covars_ = covarXX
posterior = px.predict_proba(np.atleast_2d(source))
return np.dot(posterior, E)
def calc_std_mean(input_f0):
"""
F0変換のために標準偏差と平均を求める.
"""
logF0 = np.ma.log(input_f0) # 0要素にlogをするとinfになるのでmaskする
fixed_logF0 = np.ma.fix_invalid(logF0).data # maskを取る
return np.std(fixed_logF0), np.mean(fixed_logF0) # 標準偏差と平均を返す
"""
距離を測るために,正しい目標音声を読み込む
"""
source_mfcc_for_measure_target = []
source_sp_for_measure_target = []
source_f0_for_measure_target = []
source_ap_for_measure_target = []
for name in sorted(glob.iglob(for_measure_target, recursive=True)):
print("measure_target = ", name)
x_measure_target, fs_measure_target = sf.read(name)
f0_measure_target, sp_measure_target, ap_measure_target = analyse_by_world_with_harverst(x_measure_target, fs_measure_target)
mfcc_measure_target = MFCC(fs_measure_target)
#mfcc_s_tmp = mfcc_s.mfcc(sp)
#source_mfcc_for_convert = np.hstack([mfcc_s_tmp, mfcc_s.delta(mfcc_s_tmp)])
source_mfcc_for_measure_target.append(mfcc_measure_target.mfcc(sp_measure_target))
source_sp_for_measure_target.append(sp_measure_target)
source_f0_for_measure_target.append(f0_measure_target)
source_ap_for_measure_target.append(ap_measure_target)
measure_target_data_mfcc = np.array(source_mfcc_for_measure_target)
measure_target_data_sp = np.array(source_sp_for_measure_target)
measure_target_data_f0 = np.array(source_f0_for_measure_target)
measure_target_data_ap = np.array(source_ap_for_measure_target)
def calc_mcd(source, convert, target):
"""
変換する前の音声と目標音声でDTWを行う.
その後,変換後の音声と目標音声とのMCDを計測する.
"""
dist, cost, acc, path = dtw(source, target, dist=lambda x, y: norm(x-y, ord=1))
aligned = alignment(source, target, path)
return 10.0 / np.log(10) * np.sqrt(2 * np.sum(np.square(aligned - convert)))
"""
変換を行う.
"""
timer_start = time.time()
# 事前に目標話者の標準偏差と平均を求めておく
temp_f = None
for x in range(len(target_f0)):
temp = target_f0[x].flatten()
if temp_f is None:
temp_f = temp
else:
temp_f = np.hstack((temp_f, temp))
target_std, target_mean = calc_std_mean(temp_f)
# 変換
output_mfcc = []
filer = open(mcd_text, 'a')
for i in range(len(source_data_mfcc)):
print("voice no = ", i)
# convert
source_temp = source_data_mfcc[i]
output_mfcc = np.array([convert(source_temp[frame], covarXX, fitted_source, fitted_target, covarYX, weights, source_means)[0] for frame in range(source_temp.shape[0])])
# syntehsis
source_sp_temp = source_data_sp[i]
source_f0_temp = source_data_f0[i]
source_ap_temp = source_data_ap[i]
output_imfcc = mfcc_source.imfcc(output_mfcc, source_sp_temp)
y_source = pw.synthesize(source_f0_temp, output_imfcc, source_ap_temp, fs_source, 5)
np.save(converted_voice_npy + "s{0}.npy".format(i), output_imfcc)
sf.write(converted_voice_wav + "s{0}.wav".format(i), y_source, fs_source)
# save figure spectram
range_s = output_imfcc.shape[0]
scale = [x for x in range(range_s)]
MFCC_sample_s = [source_temp[x][0] for x in range(range_s)]
MFCC_sample_c = [output_mfcc[x][0] for x in range(range_s)]
plt.subplot(311)
plt.plot(scale, MFCC_sample_s)
plt.plot(scale, MFCC_sample_c)
plt.xlabel("Flame")
plt.ylabel("amplitude MFCC")
MFCC_sample_s = [source_temp[x][1] for x in range(range_s)]
MFCC_sample_c = [output_mfcc[x][1] for x in range(range_s)]
plt.subplot(312)
plt.plot(scale, MFCC_sample_s)
plt.plot(scale, MFCC_sample_c)
plt.xlabel("Flame")
plt.ylabel("amplitude MFCC")
MFCC_sample_s = [source_temp[x][2] for x in range(range_s)]
MFCC_sample_c = [output_mfcc[x][2] for x in range(range_s)]
plt.subplot(313)
plt.plot(scale, MFCC_sample_s)
plt.plot(scale, MFCC_sample_c)
plt.xlabel("Flame")
plt.ylabel("amplitude MFCC")
plt.savefig(mfcc_save_fig_png + "s{0}.png".format(i) , format='png', dpi=300)
plt.close()
# synthesis with conveted f0
source_std, source_mean = calc_std_mean(source_f0_temp)
std_ratio = target_std / source_std
log_conv_f0 = std_ratio * (source_f0_temp - source_mean) + target_mean
conv_f0 = np.maximum(log_conv_f0, 0)
#conv_f0 = np.exp(log_conv_f0)
#print(conv_f0)
np.save(converted_voice_npy + "f{0}.npy".format(i), conv_f0)
y_conv = pw.synthesize(conv_f0, output_imfcc, source_ap_temp, fs_source, 5)
sf.write(converted_voice_with_f0_wav + "sf{0}.wav".format(i) , y_conv, fs_source)
# save figure f0
F0_s = [source_f0_temp[x] for x in range(range_s)]
F0_c = [conv_f0[x] for x in range(range_s)]
plt.plot(scale, F0_s)
plt.plot(scale, F0_c)
plt.xlabel("Frame")
plt.ylabel("amplitude")
plt.savefig(f0_save_fig_png + "f{0}.png".format(i), format='png', dpi=300)
plt.close()
# calc MCD
measure_temp = measure_target_data_mfcc[i]
mcd = calc_mcd(source_temp, output_mfcc, measure_temp)
filer.write("MCD No.{0} = {1} , shape = {2}\n".format(i, mcd, source_temp.shape))
filer.close()
print("Make Converted Spectram time = ", time.time() - timer_start , "[sec]")
```
| github_jupyter |
# A TensorFlow Word2Vec Model for Word Similarity Prediction
```
import urllib.request
import collections
import math
import os
import random
import zipfile
import datetime as dt
import numpy as np
import tensorflow as tf
```
## Background
Word2Vec is a model that was created by [Mikolov et al.](http://papers.nips.cc/paper/5021-distributed-representations-of-words-and-phrases-and-their-compositionality.pdf). It uses the concept of "word embeddings", which is a way to represent relationships between words using vectors. This makes it a useful tool to find words that are similar to eachother.
Here is an example of an embedding matrix taken from the TensorFlow tutorial:
## Data
The data used here is a cleaned version of the first 10^9 bytes of an English Wikipedia dump performed on Mar. 3, 2006. See [this site](https://cs.fit.edu/~mmahoney/compression/textdata.html) for more information.
```
def maybe_download(filename, url, expected_bytes):
"""Download a file if not present, and make sure it's the right size."""
if not os.path.exists(filename):
filename, _ = urllib.request.urlretrieve(url + filename, filename)
statinfo = os.stat(filename)
if statinfo.st_size == expected_bytes:
print('Found and verified', filename)
else:
print(statinfo.st_size)
raise Exception(
'Failed to verify ' + filename + '. Can you get to it with a browser?')
return filename
url = 'http://mattmahoney.net/dc/'
filename = maybe_download('text8.zip', url, 31344016)
# Read the data into a list of strings.
def read_data(filename):
"""Extract the first file enclosed in a zip file as a list of words."""
with zipfile.ZipFile(filename) as f:
data = tf.compat.as_str(f.read(f.namelist()[0])).split()
return data
vocabulary = read_data(filename)
print(vocabulary[:7])
['anarchism', 'originated', 'as', 'a', 'term', 'of', 'abuse']
def build_dataset(words, n_words):
"""Process raw inputs into a dataset."""
count = [['UNK', -1]]
count.extend(collections.Counter(words).most_common(n_words - 1))
dictionary = dict()
for word, _ in count:
dictionary[word] = len(dictionary)
data = list()
unk_count = 0
for word in words:
if word in dictionary:
index = dictionary[word]
else:
index = 0 # dictionary['UNK']
unk_count += 1
data.append(index)
count[0][1] = unk_count
reversed_dictionary = dict(zip(dictionary.values(), dictionary.keys()))
return data, count, dictionary, reversed_dictionary
def collect_data(vocabulary_size=10000):
"""Read data and create the dictionary"""
url = 'http://mattmahoney.net/dc/'
filename = maybe_download('text8.zip', url, 31344016)
vocabulary = read_data(filename)
print(vocabulary[:7])
data, count, dictionary, reverse_dictionary = build_dataset(vocabulary,
vocabulary_size)
del vocabulary # Hint to reduce memory.
return data, count, dictionary, reverse_dictionary
data_index = 0
def generate_batch(data, batch_size, num_skips, skip_window):
"""Generate batch data"""
global data_index
assert batch_size % num_skips == 0
assert num_skips <= 2 * skip_window
batch = np.ndarray(shape=(batch_size), dtype=np.int32)
context = np.ndarray(shape=(batch_size, 1), dtype=np.int32)
span = 2 * skip_window + 1 # [ skip_window input_word skip_window ]
buffer = collections.deque(maxlen=span)
for _ in range(span):
buffer.append(data[data_index])
data_index = (data_index + 1) % len(data)
for i in range(batch_size // num_skips):
target = skip_window # input word at the center of the buffer
targets_to_avoid = [skip_window]
for j in range(num_skips):
while target in targets_to_avoid:
target = random.randint(0, span - 1)
targets_to_avoid.append(target)
batch[i * num_skips + j] = buffer[skip_window] # this is the input word
context[i * num_skips + j, 0] = buffer[target] # these are the context words
buffer.append(data[data_index])
data_index = (data_index + 1) % len(data)
# Backtrack a little bit to avoid skipping words in the end of a batch
data_index = (data_index + len(data) - span) % len(data)
return batch, context
vocabulary_size = 10000
data, count, dictionary, reverse_dictionary = collect_data(vocabulary_size=vocabulary_size)
```
## TensorFlow Model
```
graph = tf.Graph()
batch_size = 128
embedding_size = 128 # Dimension of the embedding vector.
skip_window = 1 # How many words to consider left and right.
num_skips = 2 # How many times to reuse an input to generate a context.
# We pick a random validation set to sample nearest neighbors. Here we limit the
# validation samples to the words that have a low numeric ID, which by
# construction are also the most frequent.
valid_size = vocabulary_size # Random set of words to evaluate similarity on.
valid_window = 100 # Only pick dev samples in the head of the distribution.
valid_examples = np.arange(valid_size) # np.random.choice(valid_window, valid_size, replace=False)
num_sampled = 64 # Number of negative examples to sample.
```
There is a fast scheme called Noise Contrastive Estimation (NCE). Instead of taking the probability of the context word compared to all of the possible context words in the vocabulary, this method randomly samples 2-20 possible context words and evaluates the probability only from these.
```
with graph.as_default():
# Input data.
train_inputs = tf.placeholder(tf.int32, shape=[batch_size])
train_context = tf.placeholder(tf.int32, shape=[batch_size, 1])
valid_dataset = tf.constant(valid_examples, dtype=tf.int32)
# Look up embeddings for inputs.
embeddings = tf.Variable(
tf.random_uniform([vocabulary_size, embedding_size], -1.0, 1.0))
embed = tf.nn.embedding_lookup(embeddings, train_inputs)
# Construct the variables for the NCE loss
nce_weights = tf.Variable(
tf.truncated_normal([vocabulary_size, embedding_size],
stddev=1.0 / math.sqrt(embedding_size)))
nce_biases = tf.Variable(tf.zeros([vocabulary_size]))
nce_loss = tf.reduce_mean(
tf.nn.nce_loss(weights=nce_weights,
biases=nce_biases,
labels=train_context,
inputs=embed,
num_sampled=num_sampled,
num_classes=vocabulary_size))
optimizer = tf.train.GradientDescentOptimizer(1.0).minimize(nce_loss)
# Compute the cosine similarity between minibatch examples and all embeddings.
norm = tf.sqrt(tf.reduce_sum(tf.square(embeddings), 1, keep_dims=True))
normalized_embeddings = embeddings / norm
valid_embeddings = tf.nn.embedding_lookup(
normalized_embeddings, valid_dataset)
similarity = tf.matmul(
valid_embeddings, normalized_embeddings, transpose_b=True)
# Add variable initializer.
init = tf.global_variables_initializer()
```
## Run the Model
```
def train(graph, num_steps):
with tf.Session(graph=graph) as session:
with session.as_default():
# We must initialize all variables before we use them.
init.run()
print('Initialized')
average_loss = 0
for step in range(num_steps):
batch_inputs, batch_context = generate_batch(data,
batch_size, num_skips, skip_window)
feed_dict = {train_inputs: batch_inputs, train_context: batch_context}
# We perform one update step by evaluating the optimizer op (including it
# in the list of returned values for session.run()
_, loss_val = session.run([optimizer, nce_loss], feed_dict=feed_dict)
average_loss += loss_val
if step % 1000 == 0:
if step > 0:
average_loss /= 2000
# The average loss is an estimate of the loss over the last 2000 batches.
print('Average loss at step ', step, ': ', average_loss)
average_loss = 0
# Note that this is expensive (~20% slowdown if computed every 500 steps)
if step % 1000 == 0:
sim = similarity.eval()
for i in range(valid_size):
valid_word = reverse_dictionary[valid_examples[i]]
top_k = 8 # number of nearest neighbors
nearest = (-sim[i, :]).argsort()[1:top_k + 1]
log_str = 'Nearest to %s:' % valid_word
for k in range(top_k):
close_word = reverse_dictionary[nearest[k]]
log_str = '%s %s,' % (log_str, close_word)
print(log_str)
final_embeddings = normalized_embeddings.eval()
saver = tf.train.Saver()
saver.save(session, os.path.join("model.ckpt"))
```
### Training
```
num_steps = 10000
softmax_start_time = dt.datetime.now()
train(graph, num_steps=num_steps)
softmax_end_time = dt.datetime.now()
print("Training took {} minutes to run {} iterations".format(
(softmax_end_time-softmax_start_time).total_seconds()/60, str(num_steps)))
```
### Predict similarity
```
def predict_sim(input_word, model_path):
# Reinitialize things
with graph.as_default():
# Input data.
train_inputs = tf.placeholder(tf.int32, shape=[batch_size])
train_context = tf.placeholder(tf.int32, shape=[batch_size, 1])
valid_dataset = tf.constant(valid_examples, dtype=tf.int32)
# Look up embeddings for inputs.
embeddings = tf.Variable(
tf.random_uniform([vocabulary_size, embedding_size], -1.0, 1.0))
embed = tf.nn.embedding_lookup(embeddings, train_inputs)
# Compute the cosine similarity between minibatch examples and all embeddings.
norm = tf.sqrt(tf.reduce_sum(tf.square(embeddings), 1, keep_dims=True))
normalized_embeddings = embeddings / norm
valid_embeddings = tf.nn.embedding_lookup(
normalized_embeddings, valid_dataset)
similarity = tf.matmul(
valid_embeddings, normalized_embeddings, transpose_b=True)
# Add variable initializer.
init = tf.global_variables_initializer()
with tf.Session(graph=graph) as session:
saver = tf.train.Saver()
saver.restore(session,
os.path.join(model_path, "model.ckpt"))
sim = similarity.eval()
if input_word in dictionary:
idx = dictionary[input_word]
valid_word = reverse_dictionary[idx]
top_k = 3 # number of nearest neighbors
nearest = (-sim[idx, :]).argsort()[1:top_k + 1]
log_str = 'Nearest to %s:' % valid_word
for k in range(top_k):
close_word = reverse_dictionary[nearest[k]]
log_str = '%s %s' % (log_str, close_word)
print(log_str)
else:
return 'Word not present in dictionary. Try a different one.'
```
Let's test the trained model and see if it can predict similar words.
```
# Define location of saved model
model_path = os.getcwd()
graph = tf.Graph()
predict_sim('science', model_path)
```
## Exercises:
1. Following this tutorial, create a TensorFlow model and train it.
2. Using the model you created, adjust the hyperparatmeters and see if the model training improves.
## Advanced Exercises:
1. Download a separate dataset from the internet. Reformat so that it can be understood by TensorFlow. Train a new TensorFlow model and see if it performs better.
| github_jupyter |
TSG020- Describe nodes (Kubernetes)
===================================
Steps
-----
### Common functions
Define helper functions used in this notebook.
```
# Define `run` function for transient fault handling, suggestions on error, and scrolling updates on Windows
import sys
import os
import re
import platform
import shlex
import shutil
import datetime
from subprocess import Popen, PIPE
from IPython.display import Markdown
retry_hints = {} # Output in stderr known to be transient, therefore automatically retry
error_hints = {} # Output in stderr where a known SOP/TSG exists which will be HINTed for further help
install_hint = {} # The SOP to help install the executable if it cannot be found
def run(cmd, return_output=False, no_output=False, retry_count=0, base64_decode=False, return_as_json=False, regex_mask=None):
"""Run shell command, stream stdout, print stderr and optionally return output
NOTES:
1. Commands that need this kind of ' quoting on Windows e.g.:
kubectl get nodes -o jsonpath={.items[?(@.metadata.annotations.pv-candidate=='data-pool')].metadata.name}
Need to actually pass in as '"':
kubectl get nodes -o jsonpath={.items[?(@.metadata.annotations.pv-candidate=='"'data-pool'"')].metadata.name}
The ' quote approach, although correct when pasting into Windows cmd, will hang at the line:
`iter(p.stdout.readline, b'')`
The shlex.split call does the right thing for each platform, just use the '"' pattern for a '
"""
MAX_RETRIES = 5
output = ""
retry = False
# When running `azdata sql query` on Windows, replace any \n in """ strings, with " ", otherwise we see:
#
# ('HY090', '[HY090] [Microsoft][ODBC Driver Manager] Invalid string or buffer length (0) (SQLExecDirectW)')
#
if platform.system() == "Windows" and cmd.startswith("azdata sql query"):
cmd = cmd.replace("\n", " ")
# shlex.split is required on bash and for Windows paths with spaces
#
cmd_actual = shlex.split(cmd)
# Store this (i.e. kubectl, python etc.) to support binary context aware error_hints and retries
#
user_provided_exe_name = cmd_actual[0].lower()
# When running python, use the python in the ADS sandbox ({sys.executable})
#
if cmd.startswith("python "):
cmd_actual[0] = cmd_actual[0].replace("python", sys.executable)
# On Mac, when ADS is not launched from terminal, LC_ALL may not be set, which causes pip installs to fail
# with:
#
# UnicodeDecodeError: 'ascii' codec can't decode byte 0xc5 in position 4969: ordinal not in range(128)
#
# Setting it to a default value of "en_US.UTF-8" enables pip install to complete
#
if platform.system() == "Darwin" and "LC_ALL" not in os.environ:
os.environ["LC_ALL"] = "en_US.UTF-8"
# When running `kubectl`, if AZDATA_OPENSHIFT is set, use `oc`
#
if cmd.startswith("kubectl ") and "AZDATA_OPENSHIFT" in os.environ:
cmd_actual[0] = cmd_actual[0].replace("kubectl", "oc")
# To aid supportability, determine which binary file will actually be executed on the machine
#
which_binary = None
# Special case for CURL on Windows. The version of CURL in Windows System32 does not work to
# get JWT tokens, it returns "(56) Failure when receiving data from the peer". If another instance
# of CURL exists on the machine use that one. (Unfortunately the curl.exe in System32 is almost
# always the first curl.exe in the path, and it can't be uninstalled from System32, so here we
# look for the 2nd installation of CURL in the path)
if platform.system() == "Windows" and cmd.startswith("curl "):
path = os.getenv('PATH')
for p in path.split(os.path.pathsep):
p = os.path.join(p, "curl.exe")
if os.path.exists(p) and os.access(p, os.X_OK):
if p.lower().find("system32") == -1:
cmd_actual[0] = p
which_binary = p
break
# Find the path based location (shutil.which) of the executable that will be run (and display it to aid supportability), this
# seems to be required for .msi installs of azdata.cmd/az.cmd. (otherwise Popen returns FileNotFound)
#
# NOTE: Bash needs cmd to be the list of the space separated values hence shlex.split.
#
if which_binary == None:
which_binary = shutil.which(cmd_actual[0])
# Display an install HINT, so the user can click on a SOP to install the missing binary
#
if which_binary == None:
print(f"The path used to search for '{cmd_actual[0]}' was:")
print(sys.path)
if user_provided_exe_name in install_hint and install_hint[user_provided_exe_name] is not None:
display(Markdown(f'HINT: Use [{install_hint[user_provided_exe_name][0]}]({install_hint[user_provided_exe_name][1]}) to resolve this issue.'))
raise FileNotFoundError(f"Executable '{cmd_actual[0]}' not found in path (where/which)")
else:
cmd_actual[0] = which_binary
start_time = datetime.datetime.now().replace(microsecond=0)
cmd_display = cmd
if regex_mask is not None:
regex = re.compile(regex_mask)
cmd_display = re.sub(regex, '******', cmd)
print(f"START: {cmd_display} @ {start_time} ({datetime.datetime.utcnow().replace(microsecond=0)} UTC)")
print(f" using: {which_binary} ({platform.system()} {platform.release()} on {platform.machine()})")
print(f" cwd: {os.getcwd()}")
# Command-line tools such as CURL and AZDATA HDFS commands output
# scrolling progress bars, which causes Jupyter to hang forever, to
# workaround this, use no_output=True
#
# Work around a infinite hang when a notebook generates a non-zero return code, break out, and do not wait
#
wait = True
try:
if no_output:
p = Popen(cmd_actual)
else:
p = Popen(cmd_actual, stdout=PIPE, stderr=PIPE, bufsize=1)
with p.stdout:
for line in iter(p.stdout.readline, b''):
line = line.decode()
if return_output:
output = output + line
else:
if cmd.startswith("azdata notebook run"): # Hyperlink the .ipynb file
regex = re.compile(' "(.*)"\: "(.*)"')
match = regex.match(line)
if match:
if match.group(1).find("HTML") != -1:
display(Markdown(f' - "{match.group(1)}": "{match.group(2)}"'))
else:
display(Markdown(f' - "{match.group(1)}": "[{match.group(2)}]({match.group(2)})"'))
wait = False
break # otherwise infinite hang, have not worked out why yet.
else:
print(line, end='')
if wait:
p.wait()
except FileNotFoundError as e:
if install_hint is not None:
display(Markdown(f'HINT: Use {install_hint} to resolve this issue.'))
raise FileNotFoundError(f"Executable '{cmd_actual[0]}' not found in path (where/which)") from e
exit_code_workaround = 0 # WORKAROUND: azdata hangs on exception from notebook on p.wait()
if not no_output:
for line in iter(p.stderr.readline, b''):
try:
line_decoded = line.decode()
except UnicodeDecodeError:
# NOTE: Sometimes we get characters back that cannot be decoded(), e.g.
#
# \xa0
#
# For example see this in the response from `az group create`:
#
# ERROR: Get Token request returned http error: 400 and server
# response: {"error":"invalid_grant",# "error_description":"AADSTS700082:
# The refresh token has expired due to inactivity.\xa0The token was
# issued on 2018-10-25T23:35:11.9832872Z
#
# which generates the exception:
#
# UnicodeDecodeError: 'utf-8' codec can't decode byte 0xa0 in position 179: invalid start byte
#
print("WARNING: Unable to decode stderr line, printing raw bytes:")
print(line)
line_decoded = ""
pass
else:
# azdata emits a single empty line to stderr when doing an hdfs cp, don't
# print this empty "ERR:" as it confuses.
#
if line_decoded == "":
continue
print(f"STDERR: {line_decoded}", end='')
if line_decoded.startswith("An exception has occurred") or line_decoded.startswith("ERROR: An error occurred while executing the following cell"):
exit_code_workaround = 1
# inject HINTs to next TSG/SOP based on output in stderr
#
if user_provided_exe_name in error_hints:
for error_hint in error_hints[user_provided_exe_name]:
if line_decoded.find(error_hint[0]) != -1:
display(Markdown(f'HINT: Use [{error_hint[1]}]({error_hint[2]}) to resolve this issue.'))
# Verify if a transient error, if so automatically retry (recursive)
#
if user_provided_exe_name in retry_hints:
for retry_hint in retry_hints[user_provided_exe_name]:
if line_decoded.find(retry_hint) != -1:
if retry_count < MAX_RETRIES:
print(f"RETRY: {retry_count} (due to: {retry_hint})")
retry_count = retry_count + 1
output = run(cmd, return_output=return_output, retry_count=retry_count)
if return_output:
if base64_decode:
import base64
return base64.b64decode(output).decode('utf-8')
else:
return output
elapsed = datetime.datetime.now().replace(microsecond=0) - start_time
# WORKAROUND: We avoid infinite hang above in the `azdata notebook run` failure case, by inferring success (from stdout output), so
# don't wait here, if success known above
#
if wait:
if p.returncode != 0:
raise SystemExit(f'Shell command:\n\n\t{cmd_display} ({elapsed}s elapsed)\n\nreturned non-zero exit code: {str(p.returncode)}.\n')
else:
if exit_code_workaround !=0 :
raise SystemExit(f'Shell command:\n\n\t{cmd_display} ({elapsed}s elapsed)\n\nreturned non-zero exit code: {str(exit_code_workaround)}.\n')
print(f'\nSUCCESS: {elapsed}s elapsed.\n')
if return_output:
if base64_decode:
import base64
return base64.b64decode(output).decode('utf-8')
else:
return output
# Hints for tool retry (on transient fault), known errors and install guide
#
retry_hints = {'python': [ ], 'azdata': ['Endpoint sql-server-master does not exist', 'Endpoint livy does not exist', 'Failed to get state for cluster', 'Endpoint webhdfs does not exist', 'Adaptive Server is unavailable or does not exist', 'Error: Address already in use', 'Login timeout expired (0) (SQLDriverConnect)', 'SSPI Provider: No Kerberos credentials available', ], 'kubectl': ['A connection attempt failed because the connected party did not properly respond after a period of time, or established connection failed because connected host has failed to respond', ], }
error_hints = {'python': [['Library not loaded: /usr/local/opt/unixodbc', 'SOP012 - Install unixodbc for Mac', '../install/sop012-brew-install-odbc-for-sql-server.ipynb'], ['WARNING: You are using pip version', 'SOP040 - Upgrade pip in ADS Python sandbox', '../install/sop040-upgrade-pip.ipynb'], ], 'azdata': [['Please run \'azdata login\' to first authenticate', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['The token is expired', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['Reason: Unauthorized', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['Max retries exceeded with url: /api/v1/bdc/endpoints', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['Look at the controller logs for more details', 'TSG027 - Observe cluster deployment', '../diagnose/tsg027-observe-bdc-create.ipynb'], ['provided port is already allocated', 'TSG062 - Get tail of all previous container logs for pods in BDC namespace', '../log-files/tsg062-tail-bdc-previous-container-logs.ipynb'], ['Create cluster failed since the existing namespace', 'SOP061 - Delete a big data cluster', '../install/sop061-delete-bdc.ipynb'], ['Failed to complete kube config setup', 'TSG067 - Failed to complete kube config setup', '../repair/tsg067-failed-to-complete-kube-config-setup.ipynb'], ['Data source name not found and no default driver specified', 'SOP069 - Install ODBC for SQL Server', '../install/sop069-install-odbc-driver-for-sql-server.ipynb'], ['Can\'t open lib \'ODBC Driver 17 for SQL Server', 'SOP069 - Install ODBC for SQL Server', '../install/sop069-install-odbc-driver-for-sql-server.ipynb'], ['Control plane upgrade failed. Failed to upgrade controller.', 'TSG108 - View the controller upgrade config map', '../diagnose/tsg108-controller-failed-to-upgrade.ipynb'], ['NameError: name \'azdata_login_secret_name\' is not defined', 'SOP013 - Create secret for azdata login (inside cluster)', '../common/sop013-create-secret-for-azdata-login.ipynb'], ['ERROR: No credentials were supplied, or the credentials were unavailable or inaccessible.', 'TSG124 - \'No credentials were supplied\' error from azdata login', '../repair/tsg124-no-credentials-were-supplied.ipynb'], ['Please accept the license terms to use this product through', 'TSG126 - azdata fails with \'accept the license terms to use this product\'', '../repair/tsg126-accept-license-terms.ipynb'], ], 'kubectl': [['no such host', 'TSG010 - Get configuration contexts', '../monitor-k8s/tsg010-get-kubernetes-contexts.ipynb'], ['No connection could be made because the target machine actively refused it', 'TSG056 - Kubectl fails with No connection could be made because the target machine actively refused it', '../repair/tsg056-kubectl-no-connection-could-be-made.ipynb'], ], }
install_hint = {'azdata': [ 'SOP063 - Install azdata CLI (using package manager)', '../install/sop063-packman-install-azdata.ipynb' ], 'kubectl': [ 'SOP036 - Install kubectl command line interface', '../install/sop036-install-kubectl.ipynb' ], }
print('Common functions defined successfully.')
```
### Get the Kubernetes namespace for the big data cluster
Get the namespace of the Big Data Cluster use the kubectl command line
interface .
**NOTE:**
If there is more than one Big Data Cluster in the target Kubernetes
cluster, then either:
- set \[0\] to the correct value for the big data cluster.
- set the environment variable AZDATA\_NAMESPACE, before starting
Azure Data Studio.
```
# Place Kubernetes namespace name for BDC into 'namespace' variable
if "AZDATA_NAMESPACE" in os.environ:
namespace = os.environ["AZDATA_NAMESPACE"]
else:
try:
namespace = run(f'kubectl get namespace --selector=MSSQL_CLUSTER -o jsonpath={{.items[0].metadata.name}}', return_output=True)
except:
from IPython.display import Markdown
print(f"ERROR: Unable to find a Kubernetes namespace with label 'MSSQL_CLUSTER'. SQL Server Big Data Cluster Kubernetes namespaces contain the label 'MSSQL_CLUSTER'.")
display(Markdown(f'HINT: Use [TSG081 - Get namespaces (Kubernetes)](../monitor-k8s/tsg081-get-kubernetes-namespaces.ipynb) to resolve this issue.'))
display(Markdown(f'HINT: Use [TSG010 - Get configuration contexts](../monitor-k8s/tsg010-get-kubernetes-contexts.ipynb) to resolve this issue.'))
display(Markdown(f'HINT: Use [SOP011 - Set kubernetes configuration context](../common/sop011-set-kubernetes-context.ipynb) to resolve this issue.'))
raise
print(f'The SQL Server Big Data Cluster Kubernetes namespace is: {namespace}')
```
### Describe all nodes
```
run(f'kubectl describe nodes')
print("Notebook execution is complete.")
```
| github_jupyter |
```
import pickle
import numpy
import matplotlib.pyplot as plt
f_all = open('movie_all_data_item', 'rb')
all_data = pickle.load(f_all)
f_all.close()
f_train = open('movie_train_data_item', 'rb')
train_data = pickle.load(f_train)
f_train.close()
f_test = open('movie_test_data_item', 'rb')
test_data = pickle.load(f_test)
f_test.close()
print len(all_data)
print len(train_data)
print len(test_data)
#train_data=train_data[:300000]
#test_data=test_data[:300000]
for datum in all_data:
if datum['helpful'][0]>datum['helpful'][1]:
all_data.remove(datum)
print '123'
item_time = {}
for datum in all_data:
if datum['asin'] not in item_time.keys():
item_time[datum['asin']] = []
item_time[datum['asin']].append(datum['unixReviewTime'])
min_review_time={}
items_index=[]
for item in item_time.keys():
items_index.append(item)
min_review_time[item] = min(item_time[item])
review_duration = []
for item in item_time.keys():
review_duration.append(float(max(item_time[item])-min(item_time[item]))/(3600*24*30))
len(review_duration)
n,bins,patches = plt.hist(review_duration,48, facecolor='green')
plt.xlim([-5,115])
plt.xlabel("last review time - first review time (in months)")
plt.ylabel("number of items")
plt.title("histogram of [last review time - first review time]")
plt.xticks(numpy.arange(0,120,12))
plt.savefig('Figure_durationOfReview')
plt.show()
all_data[0]
# split data based on [0,0] or not
zero_helpful = []
non_zero_helpful = []
for datum in all_data:
if datum['helpful'] == [0,0]:
zero_helpful.append(datum)
else:
non_zero_helpful.append(datum)
zero_helpful_index=[]
zero_helpful_rating=[]
count=[0]*6
for datum in zero_helpful:
zero_helpful_index.append(datum['asin'])
zero_helpful_rating.append(datum['overall'])
if datum['overall'] == 0:
count[0] += 1
if datum['overall'] == 1:
count[1] += 1
if datum['overall'] == 2:
count[2] += 1
if datum['overall'] == 3:
count[3] += 1
if datum['overall'] == 4:
count[4] += 1
if datum['overall'] == 5:
count[5] += 1
print count
print len(zero_helpful)
print len(non_zero_helpful)
n,bins,patches = plt.hist(zero_helpful_rating,50,facecolor='green')
plt.xlim([0,6])
plt.xlabel("rating score")
plt.ylabel("number of items")
plt.title("rating histogram of items with zero helpful feature")
plt.xticks(numpy.arange(0,6,1))
plt.savefig('norm_zero_help_rating_histogram')
plt.show()
relative_time_non_zero = []
relative_time_zero = []
for datum in non_zero_helpful:
relative_time_non_zero.append(float(datum['unixReviewTime']-min(item_time[datum['asin']]))/(3600*24*30))
for datum in zero_helpful:
relative_time_zero.append(float(datum['unixReviewTime']-min(item_time[datum['asin']]))/(3600*24*30))
n,bins,patches = plt.hist(relative_time_non_zero,50,facecolor='green')
plt.xlim([0,210])
plt.xlabel("TSFR / month")
plt.ylabel("number of reviews")
plt.title("Histogram of TSFR for reviews with non zero HR")
plt.xticks(numpy.arange(0,220,20))
plt.savefig('non_zero_TSFR_histogram')
plt.show()
n,bins,patches = plt.hist(relative_time_zero,50,facecolor='green')
plt.xlim([0,210])
plt.xlabel("TSFR / month")
plt.ylabel("number of reviews")
plt.title("Histogram of TSFR for reviews with zero HR")
plt.xticks(numpy.arange(0,220,20))
plt.savefig('zero_TSFR_histogram')
plt.show()
test_length_non_zero = []
test_length_time_zero = []
for datum in non_zero_helpful:
test_length_non_zero.append(len(datum['reviewText']))
for datum in zero_helpful:
test_length_time_zero.append(len(datum['reviewText']))
n,bins,patches = plt.hist(test_length_non_zero,500,facecolor='green')
plt.xlim([0,8500])
plt.xlabel("RL")
plt.ylabel("number of reviews")
plt.title("Histogram of RL for reviews with non zero HR")
plt.xticks(numpy.arange(0,8500,1000))
plt.savefig('non_zero_RL_histogram')
plt.show()
n,bins,patches = plt.hist(test_length_time_zero,500,facecolor='green')
plt.xlim([0,5000])
plt.xlabel("RL")
plt.ylabel("number of reviews")
plt.title("Histogram of RL for reviews with zero HR")
plt.xticks(numpy.arange(0,5000,500))
plt.savefig('zero_RL_histogram')
plt.show()
non_zero_helpful_index=[]
non_zero_helpful_rating=[]
non_zero_helpful_ratio=[]
non_zero_helpful_total=[]
count_rating_1_ratio=[]
count_rating_2_ratio=[]
count_rating_3_ratio=[]
count_rating_4_ratio=[]
count_rating_5_ratio=[]
count_rating_1_total=[]
count_rating_2_total=[]
count_rating_3_total=[]
count_rating_4_total=[]
count_rating_5_total=[]
count = [0]*6
non_zero_time_feature=[]
for datum in non_zero_helpful:
non_zero_time_feature.append(float(datum['unixReviewTime']-min_review_time[datum['asin']])/(3600*24*30))
non_zero_helpful_index.append(datum['asin'])
non_zero_helpful_rating.append(datum['overall'])
non_zero_helpful_ratio.append(float(datum['helpful'][0])/datum['helpful'][1])
non_zero_helpful_total.append(datum['helpful'][1])
if datum['overall'] == 0:
count[0] += 1
if datum['overall'] == 1:
count[1] += 1
count_rating_1_ratio.append(float(datum['helpful'][0])/datum['helpful'][1])
count_rating_1_total.append(datum['helpful'][1])
if datum['overall'] == 2:
count[2] += 1
count_rating_2_ratio.append(float(datum['helpful'][0])/datum['helpful'][1])
count_rating_2_total.append(datum['helpful'][1])
if datum['overall'] == 3:
count[3] += 1
count_rating_3_ratio.append(float(datum['helpful'][0])/datum['helpful'][1])
count_rating_3_total.append(datum['helpful'][1])
if datum['overall'] == 4:
count[4] += 1
count_rating_4_ratio.append(float(datum['helpful'][0])/datum['helpful'][1])
count_rating_4_total.append(datum['helpful'][1])
if datum['overall'] == 5:
count[5] += 1
count_rating_5_ratio.append(float(datum['helpful'][0])/datum['helpful'][1])
count_rating_5_total.append(datum['helpful'][1])
print count
print sum(count)
print len(count_rating_1_total+count_rating_2_total+count_rating_3_total+count_rating_4_total+count_rating_5_total)
print max(count_rating_3_total)
n,bins,patches = plt.hist(non_zero_helpful_rating,50, facecolor='green')
plt.xlim([0,6])
plt.xlabel("rating score")
plt.ylabel("number of items")
plt.title("rating histogram of items with non-zero helpful feature")
plt.xticks(numpy.arange(0,6,1))
plt.savefig('non_zero_help_rating_histogram')
plt.show()
n,bins,patches = plt.hist(non_zero_helpful_ratio,10, facecolor='green')
plt.xlim([0,1.1])
plt.xlabel("helpful ratio")
plt.ylabel("number of items")
plt.title("helpful ratio histogram of items with non-zero helpful feature")
plt.xticks(numpy.arange(0,1.1,0.1))
plt.savefig('non_zero_help_ratio_histogram')
plt.show()
n,bins,patches = plt.hist(non_zero_helpful_total,bins = range(0,50,5), facecolor='green')
plt.xlim([-1,50])
plt.xlabel("total helpful evaluation number")
plt.ylabel("number of items")
plt.title("total helpful evaluation number histogram\n for items with non-zero helpful feature")
plt.xticks(numpy.arange(0,47,5))
plt.savefig('non_zero_help_total_evaluation_number_histogram')
plt.show()
plt.scatter(non_zero_helpful_rating,non_zero_helpful_ratio,marker='o',color='b',alpha=0.5)
plt.show()
#plt.boxplot(non_zero_helpful_rating, notch=False, vert=True, patch_artist=True)
plt.xticks(numpy.arange(-1,6,1), ['0.0', '1.0', '2.0', '3.0', '4.0', '5.0'])
rating_ratio = numpy.array([count_rating_1_ratio,count_rating_2_ratio,count_rating_3_ratio,count_rating_4_ratio,count_rating_5_ratio])
plt.boxplot(rating_ratio, notch=False, vert=True, patch_artist=True,showmeans=True)
#plt.boxplot(count_rating_1_ratio, notch=False, vert=True, patch_artist=True)
#plt.boxplot(count_rating_2_ratio, notch=False, vert=True, patch_artist=True)
#plt.boxplot(count_rating_3_ratio, notch=False, vert=True, patch_artist=True)
#plt.boxplot(count_rating_4_ratio, notch=False, vert=True, patch_artist=True)
#plt.boxplot(count_rating_5_ratio, notch=False, vert=True, patch_artist=True)
plt.ylabel('helpful ratio')
plt.xlabel('rating')
t = plt.title('helpful ratio versus rating score \nbox plot')
plt.savefig("helpful_ratio_vs_rating_box_plot")
plt.show()
plt.xticks(numpy.arange(-1,6,1), ['0.0', '1.0', '2.0', '3.0', '4.0', '5.0'])
rating_total = numpy.array([count_rating_1_total,count_rating_2_total,count_rating_3_total,count_rating_4_total,count_rating_5_total])
plt.boxplot(rating_total, notch=False, vert=True, patch_artist=True, showmeans=True)
plt.ylabel('total helpful evaluation number')
plt.xlabel('rating')
t = plt.title('total helpful evaluation number versus rating score \nbox plot')
plt.savefig("helpful_total_vs_rating_box_plot")
plt.show()
plt.xticks(numpy.arange(0,6,1), ['0.0', '1.0', '2.0', '3.0', '4.0', '5.0'])
rating_ratio = numpy.array([count_rating_1_ratio,count_rating_2_ratio,count_rating_3_ratio,count_rating_4_ratio,count_rating_5_ratio])
plt.violinplot(rating_ratio, vert=True, showmeans=True)
plt.xlabel('OR')
plt.ylabel('HR')
t = plt.title('HR vs OR \nviolin plot')
plt.savefig("HR_vs_OR_violin_plot")
plt.show()
plt.scatter(non_zero_time_feature,non_zero_helpful_ratio,marker='o',color='b',alpha=0.5)
plt.xlabel('time after first review (month)')
plt.ylabel('helpful ratio')
plt.title('helpful ratio versus time after first review')
plt.savefig("helpful_ratio_vs_time_feature_plot")
plt.show()
plt.scatter(non_zero_time_feature,non_zero_helpful_total,marker='o',color='b',alpha=0.5)
plt.xlabel('time after first review (month)')
plt.ylabel('total helpful evaluation number')
plt.title('total helpful evaluation number versus time after first review')
plt.savefig("helpful_total_vs_time_feature_plot")
plt.show()
# create time feature label
time_feature=[]
for datum in all_data:
time_feature.append(float(datum['unixReviewTime']-min_review_time[datum['asin']])/(3600*24*30))
print time_feature[0]
for datum in all_data:
if datum['helpful'][1]!=0 and float(datum['helpful'][0])/datum['helpful'][1] > 1:
all_data.remove(datum)
type(all_data)
non_zero_RL = []
non_zero_TSFR = []
non_zero_rating = []
non_zero_HR = []
for datum in non_zero_helpful:
non_zero_RL.append(len(datum['reviewText']))
non_zero_TSFR.append(float(datum['unixReviewTime']-min_review_time[datum['asin']])/(3600*24*30))
non_zero_rating.append(datum['overall'])
non_zero_HR.append(float(datum['helpful'][0])/datum['helpful'][1])
plt.scatter(non_zero_TSFR,non_zero_HR,marker='o',color='b',alpha=0.5)
plt.xlabel('TSFR / month')
plt.ylabel('HR')
plt.title('HR vs TSFR')
plt.savefig("HR_vs_TSFR")
plt.show()
plt.scatter(non_zero_RL,non_zero_HR,marker='o',color='b',alpha=0.5)
plt.xlabel('RL')
plt.ylabel('HR')
plt.title('HR vs RL')
plt.savefig("HR_vs_RL")
plt.show()
```
### Filter 0<=HR<=0.2 and 0.8<=HR<=1.0
```
filtered_non_zero_helpful = []
for datum in non_zero_helpful:
if float(datum['helpful'][0])/datum['helpful'][1]<=0.2 and float(datum['helpful'][0])/datum['helpful'][1]>=0:
filtered_non_zero_helpful.append(datum)
if float(datum['helpful'][0])/datum['helpful'][1]<=1.0 and float(datum['helpful'][0])/datum['helpful'][1]>=0.8:
filtered_non_zero_helpful.append(datum)
OR = []
for datum in filtered_non_zero_helpful:
OR.append(datum['overall'])
HR = []
rate_1_HR=[]
rate_2_HR=[]
rate_3_HR=[]
rate_4_HR=[]
rate_5_HR=[]
for datum in filtered_non_zero_helpful:
HR.append(float(datum['helpful'][0])/datum['helpful'][1])
if datum['overall'] == 1.0:
rate_1_HR.append(float(datum['helpful'][0])/datum['helpful'][1])
if datum['overall'] == 2.0:
rate_2_HR.append(float(datum['helpful'][0])/datum['helpful'][1])
if datum['overall'] == 3.0:
rate_3_HR.append(float(datum['helpful'][0])/datum['helpful'][1])
if datum['overall'] == 4.0:
rate_4_HR.append(float(datum['helpful'][0])/datum['helpful'][1])
if datum['overall'] == 5.0:
rate_5_HR.append(float(datum['helpful'][0])/datum['helpful'][1])
plt.scatter(OR,HR,marker='o',color='b',alpha=0.5)
plt.xlabel('OR')
plt.ylabel('HR')
plt.title('HR versus OR for 0<=HR<=0.2 or 0.8<=HR<=1.0')
plt.savefig("HR_vs_OR")
plt.show()
plt.xticks(numpy.arange(0,6,1), ['0.0', '1.0', '2.0', '3.0', '4.0', '5.0'])
rating_ratio = numpy.array([rate_1_HR,rate_2_HR,rate_3_HR,rate_4_HR,rate_5_HR])
plt.violinplot(rating_ratio, vert=True, showmeans=True)
plt.xlabel('OR')
plt.ylabel('HR')
t = plt.title('HR versus OR for 0<=HR<=0.2 or 0.8<=HR<=1.0')
plt.savefig("HR_vs_OR")
plt.show()
filtered_relative_time_non_zero = []
for datum in filtered_non_zero_helpful:
filtered_relative_time_non_zero.append(float(datum['unixReviewTime']-min(item_time[datum['asin']]))/(3600*24*30))
filtered_test_length_non_zero = []
for datum in filtered_non_zero_helpful:
filtered_test_length_non_zero.append(len(datum['reviewText']))
n,bins,patches = plt.hist(filtered_relative_time_non_zero,50,facecolor='green')
plt.xlim([0,210])
plt.xlabel("TSFR / month")
plt.ylabel("number of reviews")
plt.title("Histogram of TSFR for reviews with 0<=HR<=0.2 or 0.8<=HR<=1.0")
plt.xticks(numpy.arange(0,220,20))
plt.savefig('filtered_non_zero_TSFR_histogram')
plt.show()
n,bins,patches = plt.hist(filtered_test_length_non_zero,500,facecolor='green')
plt.xlim([0,8000])
plt.xlabel("RL")
plt.ylabel("number of reviews")
plt.title("Histogram of RL for reviews with 0<=HR<=0.2 or 0.8<=HR<=1.0")
plt.xticks(numpy.arange(0,8000,1000))
plt.savefig('filtered_non_zero_RL_histogram')
plt.show()
for datum in all_data[:20]:
print datum
reviewer = []
count=0
for datum in all_data:
count+=1
if count % 1000 ==0:
print count
if not datum['reviewerID'] in reviewer:
reviewer.append(datum['reviewerID'])
print len(reviewer)
```
| github_jupyter |
**Chapter 5 – Support Vector Machines**
_This notebook contains all the sample code and solutions to the exercises in chapter 5._
<table align="left">
<td>
<a target="_blank" href="https://colab.research.google.com/github/ageron/handson-ml2/blob/master/05_support_vector_machines.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
</table>
# Setup
First, let's import a few common modules, ensure MatplotLib plots figures inline and prepare a function to save the figures. We also check that Python 3.5 or later is installed (although Python 2.x may work, it is deprecated so we strongly recommend you use Python 3 instead), as well as Scikit-Learn ≥0.20.
```
# Python ≥3.5 is required
import sys
assert sys.version_info >= (3, 5)
# Scikit-Learn ≥0.20 is required
import sklearn
assert sklearn.__version__ >= "0.20"
# Common imports
import numpy as np
import os
# to make this notebook's output stable across runs
np.random.seed(42)
# To plot pretty figures
%matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rc('axes', labelsize=14)
mpl.rc('xtick', labelsize=12)
mpl.rc('ytick', labelsize=12)
# Where to save the figures
PROJECT_ROOT_DIR = "."
CHAPTER_ID = "svm"
IMAGES_PATH = os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID)
os.makedirs(IMAGES_PATH, exist_ok=True)
def save_fig(fig_id, tight_layout=True, fig_extension="png", resolution=300):
path = os.path.join(IMAGES_PATH, fig_id + "." + fig_extension)
print("Saving figure", fig_id)
if tight_layout:
plt.tight_layout()
plt.savefig(path, format=fig_extension, dpi=resolution)
```
# Large margin classification
The next few code cells generate the first figures in chapter 5. The first actual code sample comes after:
```
from sklearn.svm import SVC
from sklearn import datasets
iris = datasets.load_iris()
X = iris["data"][:, (2, 3)] # petal length, petal width
y = iris["target"]
setosa_or_versicolor = (y == 0) | (y == 1)
X = X[setosa_or_versicolor]
y = y[setosa_or_versicolor]
# SVM Classifier model
svm_clf = SVC(kernel="linear", C=float("inf"))
svm_clf.fit(X, y)
# Bad models
x0 = np.linspace(0, 5.5, 200)
pred_1 = 5*x0 - 20
pred_2 = x0 - 1.8
pred_3 = 0.1 * x0 + 0.5
def plot_svc_decision_boundary(svm_clf, xmin, xmax):
w = svm_clf.coef_[0]
b = svm_clf.intercept_[0]
# At the decision boundary, w0*x0 + w1*x1 + b = 0
# => x1 = -w0/w1 * x0 - b/w1
x0 = np.linspace(xmin, xmax, 200)
decision_boundary = -w[0]/w[1] * x0 - b/w[1]
margin = 1/w[1]
gutter_up = decision_boundary + margin
gutter_down = decision_boundary - margin
svs = svm_clf.support_vectors_
plt.scatter(svs[:, 0], svs[:, 1], s=180, facecolors='#FFAAAA')
plt.plot(x0, decision_boundary, "k-", linewidth=2)
plt.plot(x0, gutter_up, "k--", linewidth=2)
plt.plot(x0, gutter_down, "k--", linewidth=2)
fig, axes = plt.subplots(ncols=2, figsize=(10,2.7), sharey=True)
plt.sca(axes[0])
plt.plot(x0, pred_1, "g--", linewidth=2)
plt.plot(x0, pred_2, "m-", linewidth=2)
plt.plot(x0, pred_3, "r-", linewidth=2)
plt.plot(X[:, 0][y==1], X[:, 1][y==1], "bs", label="Iris versicolor")
plt.plot(X[:, 0][y==0], X[:, 1][y==0], "yo", label="Iris setosa")
plt.xlabel("Petal length", fontsize=14)
plt.ylabel("Petal width", fontsize=14)
plt.legend(loc="upper left", fontsize=14)
plt.axis([0, 5.5, 0, 2])
plt.sca(axes[1])
plot_svc_decision_boundary(svm_clf, 0, 5.5)
plt.plot(X[:, 0][y==1], X[:, 1][y==1], "bs")
plt.plot(X[:, 0][y==0], X[:, 1][y==0], "yo")
plt.xlabel("Petal length", fontsize=14)
plt.axis([0, 5.5, 0, 2])
save_fig("large_margin_classification_plot")
plt.show()
```
# Sensitivity to feature scales
```
Xs = np.array([[1, 50], [5, 20], [3, 80], [5, 60]]).astype(np.float64)
ys = np.array([0, 0, 1, 1])
svm_clf = SVC(kernel="linear", C=100)
svm_clf.fit(Xs, ys)
plt.figure(figsize=(9,2.7))
plt.subplot(121)
plt.plot(Xs[:, 0][ys==1], Xs[:, 1][ys==1], "bo")
plt.plot(Xs[:, 0][ys==0], Xs[:, 1][ys==0], "ms")
plot_svc_decision_boundary(svm_clf, 0, 6)
plt.xlabel("$x_0$", fontsize=20)
plt.ylabel("$x_1$ ", fontsize=20, rotation=0)
plt.title("Unscaled", fontsize=16)
plt.axis([0, 6, 0, 90])
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
X_scaled = scaler.fit_transform(Xs)
svm_clf.fit(X_scaled, ys)
plt.subplot(122)
plt.plot(X_scaled[:, 0][ys==1], X_scaled[:, 1][ys==1], "bo")
plt.plot(X_scaled[:, 0][ys==0], X_scaled[:, 1][ys==0], "ms")
plot_svc_decision_boundary(svm_clf, -2, 2)
plt.xlabel("$x_0$", fontsize=20)
plt.ylabel("$x'_1$ ", fontsize=20, rotation=0)
plt.title("Scaled", fontsize=16)
plt.axis([-2, 2, -2, 2])
save_fig("sensitivity_to_feature_scales_plot")
```
# Sensitivity to outliers
```
X_outliers = np.array([[3.4, 1.3], [3.2, 0.8]])
y_outliers = np.array([0, 0])
Xo1 = np.concatenate([X, X_outliers[:1]], axis=0)
yo1 = np.concatenate([y, y_outliers[:1]], axis=0)
Xo2 = np.concatenate([X, X_outliers[1:]], axis=0)
yo2 = np.concatenate([y, y_outliers[1:]], axis=0)
svm_clf2 = SVC(kernel="linear", C=10**9)
svm_clf2.fit(Xo2, yo2)
fig, axes = plt.subplots(ncols=2, figsize=(10,2.7), sharey=True)
plt.sca(axes[0])
plt.plot(Xo1[:, 0][yo1==1], Xo1[:, 1][yo1==1], "bs")
plt.plot(Xo1[:, 0][yo1==0], Xo1[:, 1][yo1==0], "yo")
plt.text(0.3, 1.0, "Impossible!", fontsize=24, color="red")
plt.xlabel("Petal length", fontsize=14)
plt.ylabel("Petal width", fontsize=14)
plt.annotate("Outlier",
xy=(X_outliers[0][0], X_outliers[0][1]),
xytext=(2.5, 1.7),
ha="center",
arrowprops=dict(facecolor='black', shrink=0.1),
fontsize=16,
)
plt.axis([0, 5.5, 0, 2])
plt.sca(axes[1])
plt.plot(Xo2[:, 0][yo2==1], Xo2[:, 1][yo2==1], "bs")
plt.plot(Xo2[:, 0][yo2==0], Xo2[:, 1][yo2==0], "yo")
plot_svc_decision_boundary(svm_clf2, 0, 5.5)
plt.xlabel("Petal length", fontsize=14)
plt.annotate("Outlier",
xy=(X_outliers[1][0], X_outliers[1][1]),
xytext=(3.2, 0.08),
ha="center",
arrowprops=dict(facecolor='black', shrink=0.1),
fontsize=16,
)
plt.axis([0, 5.5, 0, 2])
save_fig("sensitivity_to_outliers_plot")
plt.show()
```
# Large margin *vs* margin violations
This is the first code example in chapter 5:
```
import numpy as np
from sklearn import datasets
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.svm import LinearSVC
iris = datasets.load_iris()
X = iris["data"][:, (2, 3)] # petal length, petal width
y = (iris["target"] == 2).astype(np.float64) # Iris virginica
svm_clf = Pipeline([
("scaler", StandardScaler()),
("linear_svc", LinearSVC(C=1, loss="hinge", random_state=42)),
])
svm_clf.fit(X, y)
svm_clf.predict([[5.5, 1.7]])
```
Now let's generate the graph comparing different regularization settings:
```
scaler = StandardScaler()
svm_clf1 = LinearSVC(C=1, loss="hinge", random_state=42)
svm_clf2 = LinearSVC(C=100, loss="hinge", random_state=42)
scaled_svm_clf1 = Pipeline([
("scaler", scaler),
("linear_svc", svm_clf1),
])
scaled_svm_clf2 = Pipeline([
("scaler", scaler),
("linear_svc", svm_clf2),
])
scaled_svm_clf1.fit(X, y)
scaled_svm_clf2.fit(X, y)
# Convert to unscaled parameters
b1 = svm_clf1.decision_function([-scaler.mean_ / scaler.scale_])
b2 = svm_clf2.decision_function([-scaler.mean_ / scaler.scale_])
w1 = svm_clf1.coef_[0] / scaler.scale_
w2 = svm_clf2.coef_[0] / scaler.scale_
svm_clf1.intercept_ = np.array([b1])
svm_clf2.intercept_ = np.array([b2])
svm_clf1.coef_ = np.array([w1])
svm_clf2.coef_ = np.array([w2])
# Find support vectors (LinearSVC does not do this automatically)
t = y * 2 - 1
support_vectors_idx1 = (t * (X.dot(w1) + b1) < 1).ravel()
support_vectors_idx2 = (t * (X.dot(w2) + b2) < 1).ravel()
svm_clf1.support_vectors_ = X[support_vectors_idx1]
svm_clf2.support_vectors_ = X[support_vectors_idx2]
fig, axes = plt.subplots(ncols=2, figsize=(10,2.7), sharey=True)
plt.sca(axes[0])
plt.plot(X[:, 0][y==1], X[:, 1][y==1], "g^", label="Iris virginica")
plt.plot(X[:, 0][y==0], X[:, 1][y==0], "bs", label="Iris versicolor")
plot_svc_decision_boundary(svm_clf1, 4, 5.9)
plt.xlabel("Petal length", fontsize=14)
plt.ylabel("Petal width", fontsize=14)
plt.legend(loc="upper left", fontsize=14)
plt.title("$C = {}$".format(svm_clf1.C), fontsize=16)
plt.axis([4, 5.9, 0.8, 2.8])
plt.sca(axes[1])
plt.plot(X[:, 0][y==1], X[:, 1][y==1], "g^")
plt.plot(X[:, 0][y==0], X[:, 1][y==0], "bs")
plot_svc_decision_boundary(svm_clf2, 4, 5.99)
plt.xlabel("Petal length", fontsize=14)
plt.title("$C = {}$".format(svm_clf2.C), fontsize=16)
plt.axis([4, 5.9, 0.8, 2.8])
save_fig("regularization_plot")
```
# Non-linear classification
```
X1D = np.linspace(-4, 4, 9).reshape(-1, 1)
X2D = np.c_[X1D, X1D**2]
y = np.array([0, 0, 1, 1, 1, 1, 1, 0, 0])
plt.figure(figsize=(10, 3))
plt.subplot(121)
plt.grid(True, which='both')
plt.axhline(y=0, color='k')
plt.plot(X1D[:, 0][y==0], np.zeros(4), "bs")
plt.plot(X1D[:, 0][y==1], np.zeros(5), "g^")
plt.gca().get_yaxis().set_ticks([])
plt.xlabel(r"$x_1$", fontsize=20)
plt.axis([-4.5, 4.5, -0.2, 0.2])
plt.subplot(122)
plt.grid(True, which='both')
plt.axhline(y=0, color='k')
plt.axvline(x=0, color='k')
plt.plot(X2D[:, 0][y==0], X2D[:, 1][y==0], "bs")
plt.plot(X2D[:, 0][y==1], X2D[:, 1][y==1], "g^")
plt.xlabel(r"$x_1$", fontsize=20)
plt.ylabel(r"$x_2$ ", fontsize=20, rotation=0)
plt.gca().get_yaxis().set_ticks([0, 4, 8, 12, 16])
plt.plot([-4.5, 4.5], [6.5, 6.5], "r--", linewidth=3)
plt.axis([-4.5, 4.5, -1, 17])
plt.subplots_adjust(right=1)
save_fig("higher_dimensions_plot", tight_layout=False)
plt.show()
from sklearn.datasets import make_moons
X, y = make_moons(n_samples=100, noise=0.15, random_state=42)
def plot_dataset(X, y, axes):
plt.plot(X[:, 0][y==0], X[:, 1][y==0], "bs")
plt.plot(X[:, 0][y==1], X[:, 1][y==1], "g^")
plt.axis(axes)
plt.grid(True, which='both')
plt.xlabel(r"$x_1$", fontsize=20)
plt.ylabel(r"$x_2$", fontsize=20, rotation=0)
plot_dataset(X, y, [-1.5, 2.5, -1, 1.5])
plt.show()
from sklearn.datasets import make_moons
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import PolynomialFeatures
polynomial_svm_clf = Pipeline([
("poly_features", PolynomialFeatures(degree=3)),
("scaler", StandardScaler()),
("svm_clf", LinearSVC(C=10, loss="hinge", random_state=42))
])
polynomial_svm_clf.fit(X, y)
def plot_predictions(clf, axes):
x0s = np.linspace(axes[0], axes[1], 100)
x1s = np.linspace(axes[2], axes[3], 100)
x0, x1 = np.meshgrid(x0s, x1s)
X = np.c_[x0.ravel(), x1.ravel()]
y_pred = clf.predict(X).reshape(x0.shape)
y_decision = clf.decision_function(X).reshape(x0.shape)
plt.contourf(x0, x1, y_pred, cmap=plt.cm.brg, alpha=0.2)
plt.contourf(x0, x1, y_decision, cmap=plt.cm.brg, alpha=0.1)
plot_predictions(polynomial_svm_clf, [-1.5, 2.5, -1, 1.5])
plot_dataset(X, y, [-1.5, 2.5, -1, 1.5])
save_fig("moons_polynomial_svc_plot")
plt.show()
from sklearn.svm import SVC
poly_kernel_svm_clf = Pipeline([
("scaler", StandardScaler()),
("svm_clf", SVC(kernel="poly", degree=3, coef0=1, C=5))
])
poly_kernel_svm_clf.fit(X, y)
poly100_kernel_svm_clf = Pipeline([
("scaler", StandardScaler()),
("svm_clf", SVC(kernel="poly", degree=10, coef0=100, C=5))
])
poly100_kernel_svm_clf.fit(X, y)
fig, axes = plt.subplots(ncols=2, figsize=(10.5, 4), sharey=True)
plt.sca(axes[0])
plot_predictions(poly_kernel_svm_clf, [-1.5, 2.45, -1, 1.5])
plot_dataset(X, y, [-1.5, 2.4, -1, 1.5])
plt.title(r"$d=3, r=1, C=5$", fontsize=18)
plt.sca(axes[1])
plot_predictions(poly100_kernel_svm_clf, [-1.5, 2.45, -1, 1.5])
plot_dataset(X, y, [-1.5, 2.4, -1, 1.5])
plt.title(r"$d=10, r=100, C=5$", fontsize=18)
plt.ylabel("")
save_fig("moons_kernelized_polynomial_svc_plot")
plt.show()
def gaussian_rbf(x, landmark, gamma):
return np.exp(-gamma * np.linalg.norm(x - landmark, axis=1)**2)
gamma = 0.3
x1s = np.linspace(-4.5, 4.5, 200).reshape(-1, 1)
x2s = gaussian_rbf(x1s, -2, gamma)
x3s = gaussian_rbf(x1s, 1, gamma)
XK = np.c_[gaussian_rbf(X1D, -2, gamma), gaussian_rbf(X1D, 1, gamma)]
yk = np.array([0, 0, 1, 1, 1, 1, 1, 0, 0])
plt.figure(figsize=(10.5, 4))
plt.subplot(121)
plt.grid(True, which='both')
plt.axhline(y=0, color='k')
plt.scatter(x=[-2, 1], y=[0, 0], s=150, alpha=0.5, c="red")
plt.plot(X1D[:, 0][yk==0], np.zeros(4), "bs")
plt.plot(X1D[:, 0][yk==1], np.zeros(5), "g^")
plt.plot(x1s, x2s, "g--")
plt.plot(x1s, x3s, "b:")
plt.gca().get_yaxis().set_ticks([0, 0.25, 0.5, 0.75, 1])
plt.xlabel(r"$x_1$", fontsize=20)
plt.ylabel(r"Similarity", fontsize=14)
plt.annotate(r'$\mathbf{x}$',
xy=(X1D[3, 0], 0),
xytext=(-0.5, 0.20),
ha="center",
arrowprops=dict(facecolor='black', shrink=0.1),
fontsize=18,
)
plt.text(-2, 0.9, "$x_2$", ha="center", fontsize=20)
plt.text(1, 0.9, "$x_3$", ha="center", fontsize=20)
plt.axis([-4.5, 4.5, -0.1, 1.1])
plt.subplot(122)
plt.grid(True, which='both')
plt.axhline(y=0, color='k')
plt.axvline(x=0, color='k')
plt.plot(XK[:, 0][yk==0], XK[:, 1][yk==0], "bs")
plt.plot(XK[:, 0][yk==1], XK[:, 1][yk==1], "g^")
plt.xlabel(r"$x_2$", fontsize=20)
plt.ylabel(r"$x_3$ ", fontsize=20, rotation=0)
plt.annotate(r'$\phi\left(\mathbf{x}\right)$',
xy=(XK[3, 0], XK[3, 1]),
xytext=(0.65, 0.50),
ha="center",
arrowprops=dict(facecolor='black', shrink=0.1),
fontsize=18,
)
plt.plot([-0.1, 1.1], [0.57, -0.1], "r--", linewidth=3)
plt.axis([-0.1, 1.1, -0.1, 1.1])
plt.subplots_adjust(right=1)
save_fig("kernel_method_plot")
plt.show()
x1_example = X1D[3, 0]
for landmark in (-2, 1):
k = gaussian_rbf(np.array([[x1_example]]), np.array([[landmark]]), gamma)
print("Phi({}, {}) = {}".format(x1_example, landmark, k))
rbf_kernel_svm_clf = Pipeline([
("scaler", StandardScaler()),
("svm_clf", SVC(kernel="rbf", gamma=5, C=0.001))
])
rbf_kernel_svm_clf.fit(X, y)
from sklearn.svm import SVC
gamma1, gamma2 = 0.1, 5
C1, C2 = 0.001, 1000
hyperparams = (gamma1, C1), (gamma1, C2), (gamma2, C1), (gamma2, C2)
svm_clfs = []
for gamma, C in hyperparams:
rbf_kernel_svm_clf = Pipeline([
("scaler", StandardScaler()),
("svm_clf", SVC(kernel="rbf", gamma=gamma, C=C))
])
rbf_kernel_svm_clf.fit(X, y)
svm_clfs.append(rbf_kernel_svm_clf)
fig, axes = plt.subplots(nrows=2, ncols=2, figsize=(10.5, 7), sharex=True, sharey=True)
for i, svm_clf in enumerate(svm_clfs):
plt.sca(axes[i // 2, i % 2])
plot_predictions(svm_clf, [-1.5, 2.45, -1, 1.5])
plot_dataset(X, y, [-1.5, 2.45, -1, 1.5])
gamma, C = hyperparams[i]
plt.title(r"$\gamma = {}, C = {}$".format(gamma, C), fontsize=16)
if i in (0, 1):
plt.xlabel("")
if i in (1, 3):
plt.ylabel("")
save_fig("moons_rbf_svc_plot")
plt.show()
```
# Regression
```
np.random.seed(42)
m = 50
X = 2 * np.random.rand(m, 1)
y = (4 + 3 * X + np.random.randn(m, 1)).ravel()
from sklearn.svm import LinearSVR
svm_reg = LinearSVR(epsilon=1.5, random_state=42)
svm_reg.fit(X, y)
svm_reg1 = LinearSVR(epsilon=1.5, random_state=42)
svm_reg2 = LinearSVR(epsilon=0.5, random_state=42)
svm_reg1.fit(X, y)
svm_reg2.fit(X, y)
def find_support_vectors(svm_reg, X, y):
y_pred = svm_reg.predict(X)
off_margin = (np.abs(y - y_pred) >= svm_reg.epsilon)
return np.argwhere(off_margin)
svm_reg1.support_ = find_support_vectors(svm_reg1, X, y)
svm_reg2.support_ = find_support_vectors(svm_reg2, X, y)
eps_x1 = 1
eps_y_pred = svm_reg1.predict([[eps_x1]])
def plot_svm_regression(svm_reg, X, y, axes):
x1s = np.linspace(axes[0], axes[1], 100).reshape(100, 1)
y_pred = svm_reg.predict(x1s)
plt.plot(x1s, y_pred, "k-", linewidth=2, label=r"$\hat{y}$")
plt.plot(x1s, y_pred + svm_reg.epsilon, "k--")
plt.plot(x1s, y_pred - svm_reg.epsilon, "k--")
plt.scatter(X[svm_reg.support_], y[svm_reg.support_], s=180, facecolors='#FFAAAA')
plt.plot(X, y, "bo")
plt.xlabel(r"$x_1$", fontsize=18)
plt.legend(loc="upper left", fontsize=18)
plt.axis(axes)
fig, axes = plt.subplots(ncols=2, figsize=(9, 4), sharey=True)
plt.sca(axes[0])
plot_svm_regression(svm_reg1, X, y, [0, 2, 3, 11])
plt.title(r"$\epsilon = {}$".format(svm_reg1.epsilon), fontsize=18)
plt.ylabel(r"$y$", fontsize=18, rotation=0)
#plt.plot([eps_x1, eps_x1], [eps_y_pred, eps_y_pred - svm_reg1.epsilon], "k-", linewidth=2)
plt.annotate(
'', xy=(eps_x1, eps_y_pred), xycoords='data',
xytext=(eps_x1, eps_y_pred - svm_reg1.epsilon),
textcoords='data', arrowprops={'arrowstyle': '<->', 'linewidth': 1.5}
)
plt.text(0.91, 5.6, r"$\epsilon$", fontsize=20)
plt.sca(axes[1])
plot_svm_regression(svm_reg2, X, y, [0, 2, 3, 11])
plt.title(r"$\epsilon = {}$".format(svm_reg2.epsilon), fontsize=18)
save_fig("svm_regression_plot")
plt.show()
np.random.seed(42)
m = 100
X = 2 * np.random.rand(m, 1) - 1
y = (0.2 + 0.1 * X + 0.5 * X**2 + np.random.randn(m, 1)/10).ravel()
```
**Note**: to be future-proof, we set `gamma="scale"`, as this will be the default value in Scikit-Learn 0.22.
```
from sklearn.svm import SVR
svm_poly_reg = SVR(kernel="poly", degree=2, C=100, epsilon=0.1, gamma="scale")
svm_poly_reg.fit(X, y)
from sklearn.svm import SVR
svm_poly_reg1 = SVR(kernel="poly", degree=2, C=100, epsilon=0.1, gamma="scale")
svm_poly_reg2 = SVR(kernel="poly", degree=2, C=0.01, epsilon=0.1, gamma="scale")
svm_poly_reg1.fit(X, y)
svm_poly_reg2.fit(X, y)
fig, axes = plt.subplots(ncols=2, figsize=(9, 4), sharey=True)
plt.sca(axes[0])
plot_svm_regression(svm_poly_reg1, X, y, [-1, 1, 0, 1])
plt.title(r"$degree={}, C={}, \epsilon = {}$".format(svm_poly_reg1.degree, svm_poly_reg1.C, svm_poly_reg1.epsilon), fontsize=18)
plt.ylabel(r"$y$", fontsize=18, rotation=0)
plt.sca(axes[1])
plot_svm_regression(svm_poly_reg2, X, y, [-1, 1, 0, 1])
plt.title(r"$degree={}, C={}, \epsilon = {}$".format(svm_poly_reg2.degree, svm_poly_reg2.C, svm_poly_reg2.epsilon), fontsize=18)
save_fig("svm_with_polynomial_kernel_plot")
plt.show()
```
# Under the hood
```
iris = datasets.load_iris()
X = iris["data"][:, (2, 3)] # petal length, petal width
y = (iris["target"] == 2).astype(np.float64) # Iris virginica
from mpl_toolkits.mplot3d import Axes3D
def plot_3D_decision_function(ax, w, b, x1_lim=[4, 6], x2_lim=[0.8, 2.8]):
x1_in_bounds = (X[:, 0] > x1_lim[0]) & (X[:, 0] < x1_lim[1])
X_crop = X[x1_in_bounds]
y_crop = y[x1_in_bounds]
x1s = np.linspace(x1_lim[0], x1_lim[1], 20)
x2s = np.linspace(x2_lim[0], x2_lim[1], 20)
x1, x2 = np.meshgrid(x1s, x2s)
xs = np.c_[x1.ravel(), x2.ravel()]
df = (xs.dot(w) + b).reshape(x1.shape)
m = 1 / np.linalg.norm(w)
boundary_x2s = -x1s*(w[0]/w[1])-b/w[1]
margin_x2s_1 = -x1s*(w[0]/w[1])-(b-1)/w[1]
margin_x2s_2 = -x1s*(w[0]/w[1])-(b+1)/w[1]
ax.plot_surface(x1s, x2, np.zeros_like(x1),
color="b", alpha=0.2, cstride=100, rstride=100)
ax.plot(x1s, boundary_x2s, 0, "k-", linewidth=2, label=r"$h=0$")
ax.plot(x1s, margin_x2s_1, 0, "k--", linewidth=2, label=r"$h=\pm 1$")
ax.plot(x1s, margin_x2s_2, 0, "k--", linewidth=2)
ax.plot(X_crop[:, 0][y_crop==1], X_crop[:, 1][y_crop==1], 0, "g^")
ax.plot_wireframe(x1, x2, df, alpha=0.3, color="k")
ax.plot(X_crop[:, 0][y_crop==0], X_crop[:, 1][y_crop==0], 0, "bs")
ax.axis(x1_lim + x2_lim)
ax.text(4.5, 2.5, 3.8, "Decision function $h$", fontsize=16)
ax.set_xlabel(r"Petal length", fontsize=16, labelpad=10)
ax.set_ylabel(r"Petal width", fontsize=16, labelpad=10)
ax.set_zlabel(r"$h = \mathbf{w}^T \mathbf{x} + b$", fontsize=18, labelpad=5)
ax.legend(loc="upper left", fontsize=16)
fig = plt.figure(figsize=(11, 6))
ax1 = fig.add_subplot(111, projection='3d')
plot_3D_decision_function(ax1, w=svm_clf2.coef_[0], b=svm_clf2.intercept_[0])
save_fig("iris_3D_plot")
plt.show()
```
# Small weight vector results in a large margin
```
def plot_2D_decision_function(w, b, ylabel=True, x1_lim=[-3, 3]):
x1 = np.linspace(x1_lim[0], x1_lim[1], 200)
y = w * x1 + b
m = 1 / w
plt.plot(x1, y)
plt.plot(x1_lim, [1, 1], "k:")
plt.plot(x1_lim, [-1, -1], "k:")
plt.axhline(y=0, color='k')
plt.axvline(x=0, color='k')
plt.plot([m, m], [0, 1], "k--")
plt.plot([-m, -m], [0, -1], "k--")
plt.plot([-m, m], [0, 0], "k-o", linewidth=3)
plt.axis(x1_lim + [-2, 2])
plt.xlabel(r"$x_1$", fontsize=16)
if ylabel:
plt.ylabel(r"$w_1 x_1$ ", rotation=0, fontsize=16)
plt.title(r"$w_1 = {}$".format(w), fontsize=16)
fig, axes = plt.subplots(ncols=2, figsize=(9, 3.2), sharey=True)
plt.sca(axes[0])
plot_2D_decision_function(1, 0)
plt.sca(axes[1])
plot_2D_decision_function(0.5, 0, ylabel=False)
save_fig("small_w_large_margin_plot")
plt.show()
from sklearn.svm import SVC
from sklearn import datasets
iris = datasets.load_iris()
X = iris["data"][:, (2, 3)] # petal length, petal width
y = (iris["target"] == 2).astype(np.float64) # Iris virginica
svm_clf = SVC(kernel="linear", C=1)
svm_clf.fit(X, y)
svm_clf.predict([[5.3, 1.3]])
```
# Hinge loss
```
t = np.linspace(-2, 4, 200)
h = np.where(1 - t < 0, 0, 1 - t) # max(0, 1-t)
plt.figure(figsize=(5,2.8))
plt.plot(t, h, "b-", linewidth=2, label="$max(0, 1 - t)$")
plt.grid(True, which='both')
plt.axhline(y=0, color='k')
plt.axvline(x=0, color='k')
plt.yticks(np.arange(-1, 2.5, 1))
plt.xlabel("$t$", fontsize=16)
plt.axis([-2, 4, -1, 2.5])
plt.legend(loc="upper right", fontsize=16)
save_fig("hinge_plot")
plt.show()
```
# Extra material
## Training time
```
X, y = make_moons(n_samples=1000, noise=0.4, random_state=42)
plt.plot(X[:, 0][y==0], X[:, 1][y==0], "bs")
plt.plot(X[:, 0][y==1], X[:, 1][y==1], "g^")
import time
tol = 0.1
tols = []
times = []
for i in range(10):
svm_clf = SVC(kernel="poly", gamma=3, C=10, tol=tol, verbose=1)
t1 = time.time()
svm_clf.fit(X, y)
t2 = time.time()
times.append(t2-t1)
tols.append(tol)
print(i, tol, t2-t1)
tol /= 10
plt.semilogx(tols, times, "bo-")
plt.xlabel("Tolerance", fontsize=16)
plt.ylabel("Time (seconds)", fontsize=16)
plt.grid(True)
plt.show()
```
## Linear SVM classifier implementation using Batch Gradient Descent
```
# Training set
X = iris["data"][:, (2, 3)] # petal length, petal width
y = (iris["target"] == 2).astype(np.float64).reshape(-1, 1) # Iris virginica
from sklearn.base import BaseEstimator
class MyLinearSVC(BaseEstimator):
def __init__(self, C=1, eta0=1, eta_d=10000, n_epochs=1000, random_state=None):
self.C = C
self.eta0 = eta0
self.n_epochs = n_epochs
self.random_state = random_state
self.eta_d = eta_d
def eta(self, epoch):
return self.eta0 / (epoch + self.eta_d)
def fit(self, X, y):
# Random initialization
if self.random_state:
np.random.seed(self.random_state)
w = np.random.randn(X.shape[1], 1) # n feature weights
b = 0
m = len(X)
t = y * 2 - 1 # -1 if t==0, +1 if t==1
X_t = X * t
self.Js=[]
# Training
for epoch in range(self.n_epochs):
support_vectors_idx = (X_t.dot(w) + t * b < 1).ravel()
X_t_sv = X_t[support_vectors_idx]
t_sv = t[support_vectors_idx]
J = 1/2 * np.sum(w * w) + self.C * (np.sum(1 - X_t_sv.dot(w)) - b * np.sum(t_sv))
self.Js.append(J)
w_gradient_vector = w - self.C * np.sum(X_t_sv, axis=0).reshape(-1, 1)
b_derivative = -C * np.sum(t_sv)
w = w - self.eta(epoch) * w_gradient_vector
b = b - self.eta(epoch) * b_derivative
self.intercept_ = np.array([b])
self.coef_ = np.array([w])
support_vectors_idx = (X_t.dot(w) + t * b < 1).ravel()
self.support_vectors_ = X[support_vectors_idx]
return self
def decision_function(self, X):
return X.dot(self.coef_[0]) + self.intercept_[0]
def predict(self, X):
return (self.decision_function(X) >= 0).astype(np.float64)
C=2
svm_clf = MyLinearSVC(C=C, eta0 = 10, eta_d = 1000, n_epochs=60000, random_state=2)
svm_clf.fit(X, y)
svm_clf.predict(np.array([[5, 2], [4, 1]]))
plt.plot(range(svm_clf.n_epochs), svm_clf.Js)
plt.axis([0, svm_clf.n_epochs, 0, 100])
print(svm_clf.intercept_, svm_clf.coef_)
svm_clf2 = SVC(kernel="linear", C=C)
svm_clf2.fit(X, y.ravel())
print(svm_clf2.intercept_, svm_clf2.coef_)
yr = y.ravel()
fig, axes = plt.subplots(ncols=2, figsize=(11, 3.2), sharey=True)
plt.sca(axes[0])
plt.plot(X[:, 0][yr==1], X[:, 1][yr==1], "g^", label="Iris virginica")
plt.plot(X[:, 0][yr==0], X[:, 1][yr==0], "bs", label="Not Iris virginica")
plot_svc_decision_boundary(svm_clf, 4, 6)
plt.xlabel("Petal length", fontsize=14)
plt.ylabel("Petal width", fontsize=14)
plt.title("MyLinearSVC", fontsize=14)
plt.axis([4, 6, 0.8, 2.8])
plt.legend(loc="upper left")
plt.sca(axes[1])
plt.plot(X[:, 0][yr==1], X[:, 1][yr==1], "g^")
plt.plot(X[:, 0][yr==0], X[:, 1][yr==0], "bs")
plot_svc_decision_boundary(svm_clf2, 4, 6)
plt.xlabel("Petal length", fontsize=14)
plt.title("SVC", fontsize=14)
plt.axis([4, 6, 0.8, 2.8])
from sklearn.linear_model import SGDClassifier
sgd_clf = SGDClassifier(loss="hinge", alpha=0.017, max_iter=1000, tol=1e-3, random_state=42)
sgd_clf.fit(X, y.ravel())
m = len(X)
t = y * 2 - 1 # -1 if t==0, +1 if t==1
X_b = np.c_[np.ones((m, 1)), X] # Add bias input x0=1
X_b_t = X_b * t
sgd_theta = np.r_[sgd_clf.intercept_[0], sgd_clf.coef_[0]]
print(sgd_theta)
support_vectors_idx = (X_b_t.dot(sgd_theta) < 1).ravel()
sgd_clf.support_vectors_ = X[support_vectors_idx]
sgd_clf.C = C
plt.figure(figsize=(5.5,3.2))
plt.plot(X[:, 0][yr==1], X[:, 1][yr==1], "g^")
plt.plot(X[:, 0][yr==0], X[:, 1][yr==0], "bs")
plot_svc_decision_boundary(sgd_clf, 4, 6)
plt.xlabel("Petal length", fontsize=14)
plt.ylabel("Petal width", fontsize=14)
plt.title("SGDClassifier", fontsize=14)
plt.axis([4, 6, 0.8, 2.8])
```
# Exercise solutions
## 1. to 7.
See appendix A.
# 8.
_Exercise: train a `LinearSVC` on a linearly separable dataset. Then train an `SVC` and a `SGDClassifier` on the same dataset. See if you can get them to produce roughly the same model._
Let's use the Iris dataset: the Iris Setosa and Iris Versicolor classes are linearly separable.
```
from sklearn import datasets
iris = datasets.load_iris()
X = iris["data"][:, (2, 3)] # petal length, petal width
y = iris["target"]
setosa_or_versicolor = (y == 0) | (y == 1)
X = X[setosa_or_versicolor]
y = y[setosa_or_versicolor]
from sklearn.svm import SVC, LinearSVC
from sklearn.linear_model import SGDClassifier
from sklearn.preprocessing import StandardScaler
C = 5
alpha = 1 / (C * len(X))
lin_clf = LinearSVC(loss="hinge", C=C, random_state=42)
svm_clf = SVC(kernel="linear", C=C)
sgd_clf = SGDClassifier(loss="hinge", learning_rate="constant", eta0=0.001, alpha=alpha,
max_iter=1000, tol=1e-3, random_state=42)
scaler = StandardScaler()
X_scaled = scaler.fit_transform(X)
lin_clf.fit(X_scaled, y)
svm_clf.fit(X_scaled, y)
sgd_clf.fit(X_scaled, y)
print("LinearSVC: ", lin_clf.intercept_, lin_clf.coef_)
print("SVC: ", svm_clf.intercept_, svm_clf.coef_)
print("SGDClassifier(alpha={:.5f}):".format(sgd_clf.alpha), sgd_clf.intercept_, sgd_clf.coef_)
```
Let's plot the decision boundaries of these three models:
```
# Compute the slope and bias of each decision boundary
w1 = -lin_clf.coef_[0, 0]/lin_clf.coef_[0, 1]
b1 = -lin_clf.intercept_[0]/lin_clf.coef_[0, 1]
w2 = -svm_clf.coef_[0, 0]/svm_clf.coef_[0, 1]
b2 = -svm_clf.intercept_[0]/svm_clf.coef_[0, 1]
w3 = -sgd_clf.coef_[0, 0]/sgd_clf.coef_[0, 1]
b3 = -sgd_clf.intercept_[0]/sgd_clf.coef_[0, 1]
# Transform the decision boundary lines back to the original scale
line1 = scaler.inverse_transform([[-10, -10 * w1 + b1], [10, 10 * w1 + b1]])
line2 = scaler.inverse_transform([[-10, -10 * w2 + b2], [10, 10 * w2 + b2]])
line3 = scaler.inverse_transform([[-10, -10 * w3 + b3], [10, 10 * w3 + b3]])
# Plot all three decision boundaries
plt.figure(figsize=(11, 4))
plt.plot(line1[:, 0], line1[:, 1], "k:", label="LinearSVC")
plt.plot(line2[:, 0], line2[:, 1], "b--", linewidth=2, label="SVC")
plt.plot(line3[:, 0], line3[:, 1], "r-", label="SGDClassifier")
plt.plot(X[:, 0][y==1], X[:, 1][y==1], "bs") # label="Iris versicolor"
plt.plot(X[:, 0][y==0], X[:, 1][y==0], "yo") # label="Iris setosa"
plt.xlabel("Petal length", fontsize=14)
plt.ylabel("Petal width", fontsize=14)
plt.legend(loc="upper center", fontsize=14)
plt.axis([0, 5.5, 0, 2])
plt.show()
```
Close enough!
# 9.
_Exercise: train an SVM classifier on the MNIST dataset. Since SVM classifiers are binary classifiers, you will need to use one-versus-all to classify all 10 digits. You may want to tune the hyperparameters using small validation sets to speed up the process. What accuracy can you reach?_
First, let's load the dataset and split it into a training set and a test set. We could use `train_test_split()` but people usually just take the first 60,000 instances for the training set, and the last 10,000 instances for the test set (this makes it possible to compare your model's performance with others):
```
from sklearn.datasets import fetch_openml
mnist = fetch_openml('mnist_784', version=1, cache=True)
X = mnist["data"]
y = mnist["target"].astype(np.uint8)
X_train = X[:60000]
y_train = y[:60000]
X_test = X[60000:]
y_test = y[60000:]
```
Many training algorithms are sensitive to the order of the training instances, so it's generally good practice to shuffle them first. However, the dataset is already shuffled, so we do not need to do it.
Let's start simple, with a linear SVM classifier. It will automatically use the One-vs-All (also called One-vs-the-Rest, OvR) strategy, so there's nothing special we need to do. Easy!
**Warning**: this may take a few minutes depending on your hardware.
```
lin_clf = LinearSVC(random_state=42)
lin_clf.fit(X_train, y_train)
```
Let's make predictions on the training set and measure the accuracy (we don't want to measure it on the test set yet, since we have not selected and trained the final model yet):
```
from sklearn.metrics import accuracy_score
y_pred = lin_clf.predict(X_train)
accuracy_score(y_train, y_pred)
```
Okay, 89.5% accuracy on MNIST is pretty bad. This linear model is certainly too simple for MNIST, but perhaps we just needed to scale the data first:
```
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train.astype(np.float32))
X_test_scaled = scaler.transform(X_test.astype(np.float32))
```
**Warning**: this may take a few minutes depending on your hardware.
```
lin_clf = LinearSVC(random_state=42)
lin_clf.fit(X_train_scaled, y_train)
y_pred = lin_clf.predict(X_train_scaled)
accuracy_score(y_train, y_pred)
```
That's much better (we cut the error rate by about 25%), but still not great at all for MNIST. If we want to use an SVM, we will have to use a kernel. Let's try an `SVC` with an RBF kernel (the default).
**Note**: to be future-proof we set `gamma="scale"` since it will be the default value in Scikit-Learn 0.22.
```
svm_clf = SVC(gamma="scale")
svm_clf.fit(X_train_scaled[:10000], y_train[:10000])
y_pred = svm_clf.predict(X_train_scaled)
accuracy_score(y_train, y_pred)
```
That's promising, we get better performance even though we trained the model on 6 times less data. Let's tune the hyperparameters by doing a randomized search with cross validation. We will do this on a small dataset just to speed up the process:
```
from sklearn.model_selection import RandomizedSearchCV
from scipy.stats import reciprocal, uniform
param_distributions = {"gamma": reciprocal(0.001, 0.1), "C": uniform(1, 10)}
rnd_search_cv = RandomizedSearchCV(svm_clf, param_distributions, n_iter=10, verbose=2, cv=3)
rnd_search_cv.fit(X_train_scaled[:1000], y_train[:1000])
rnd_search_cv.best_estimator_
rnd_search_cv.best_score_
```
This looks pretty low but remember we only trained the model on 1,000 instances. Let's retrain the best estimator on the whole training set (run this at night, it will take hours):
```
rnd_search_cv.best_estimator_.fit(X_train_scaled, y_train)
y_pred = rnd_search_cv.best_estimator_.predict(X_train_scaled)
accuracy_score(y_train, y_pred)
```
Ah, this looks good! Let's select this model. Now we can test it on the test set:
```
y_pred = rnd_search_cv.best_estimator_.predict(X_test_scaled)
accuracy_score(y_test, y_pred)
```
Not too bad, but apparently the model is overfitting slightly. It's tempting to tweak the hyperparameters a bit more (e.g. decreasing `C` and/or `gamma`), but we would run the risk of overfitting the test set. Other people have found that the hyperparameters `C=5` and `gamma=0.005` yield even better performance (over 98% accuracy). By running the randomized search for longer and on a larger part of the training set, you may be able to find this as well.
## 10.
_Exercise: train an SVM regressor on the California housing dataset._
Let's load the dataset using Scikit-Learn's `fetch_california_housing()` function:
```
from sklearn.datasets import fetch_california_housing
housing = fetch_california_housing()
X = housing["data"]
y = housing["target"]
```
Split it into a training set and a test set:
```
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
```
Don't forget to scale the data:
```
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train)
X_test_scaled = scaler.transform(X_test)
```
Let's train a simple `LinearSVR` first:
```
from sklearn.svm import LinearSVR
lin_svr = LinearSVR(random_state=42)
lin_svr.fit(X_train_scaled, y_train)
```
Let's see how it performs on the training set:
```
from sklearn.metrics import mean_squared_error
y_pred = lin_svr.predict(X_train_scaled)
mse = mean_squared_error(y_train, y_pred)
mse
```
Let's look at the RMSE:
```
np.sqrt(mse)
```
In this training set, the targets are tens of thousands of dollars. The RMSE gives a rough idea of the kind of error you should expect (with a higher weight for large errors): so with this model we can expect errors somewhere around $10,000. Not great. Let's see if we can do better with an RBF Kernel. We will use randomized search with cross validation to find the appropriate hyperparameter values for `C` and `gamma`:
```
from sklearn.svm import SVR
from sklearn.model_selection import RandomizedSearchCV
from scipy.stats import reciprocal, uniform
param_distributions = {"gamma": reciprocal(0.001, 0.1), "C": uniform(1, 10)}
rnd_search_cv = RandomizedSearchCV(SVR(), param_distributions, n_iter=10, verbose=2, cv=3, random_state=42)
rnd_search_cv.fit(X_train_scaled, y_train)
rnd_search_cv.best_estimator_
```
Now let's measure the RMSE on the training set:
```
y_pred = rnd_search_cv.best_estimator_.predict(X_train_scaled)
mse = mean_squared_error(y_train, y_pred)
np.sqrt(mse)
```
Looks much better than the linear model. Let's select this model and evaluate it on the test set:
```
y_pred = rnd_search_cv.best_estimator_.predict(X_test_scaled)
mse = mean_squared_error(y_test, y_pred)
np.sqrt(mse)
```
| github_jupyter |
# T003 · Molecular filtering: unwanted substructures
Authors:
- Maximilian Driller, CADD seminar, 2017, Charité/FU Berlin
- Sandra Krüger, CADD seminar, 2018, Charité/FU Berlin
__Talktorial T003__: This talktorial is part of the TeachOpenCADD pipeline described in the first TeachOpenCADD publication ([_J. Cheminform._ (2019), **11**, 1-7](https://jcheminf.biomedcentral.com/articles/10.1186/s13321-019-0351-x)), comprising of talktorials T001-T010.
## Aim of this talktorial
There are some substructures we prefer not to include into our screening library. In this talktorial, we learn about different types of such unwanted substructures and how to find, highlight and remove them with RDKit.
### Contents in Theory
* Unwanted substructures
* Pan Assay Interference Compounds (PAINS)
### Contents in Practical
* Load and visualize data
* Filter for PAINS
* Filter for unwanted substructures
* Highlight substructures
* Substructure statistics
### References
* Pan Assay Interference compounds ([wikipedia](https://en.wikipedia.org/wiki/Pan-assay_interference_compounds), [_J. Med. Chem._ (2010), **53**, 2719-2740](https://pubs.acs.org/doi/abs/10.1021/jm901137j))
* Unwanted substructures according to Brenk *et al.* ([_Chem. Med. Chem._ (2008), **3**, 435-44](https://onlinelibrary.wiley.com/doi/full/10.1002/cmdc.200700139))
* Inspired by a Teach-Discover-Treat tutorial ([repository](https://github.com/sriniker/TDT-tutorial-2014/blob/master/TDT_challenge_tutorial.ipynb))
* RDKit ([repository](https://github.com/rdkit/rdkit), [documentation](https://www.rdkit.org/docs/index.html))
## Theory
### Unwanted substructures
Substructures can be unfavorable, e.g., because they are toxic or reactive, due to unfavorable pharmacokinetic properties, or because they likely interfere with certain assays.
Nowadays, drug discovery campaigns often involve [high throughput screening](https://en.wikipedia.org/wiki/High-throughput_screening). Filtering unwanted substructures can support assembling more efficient screening libraries, which can save time and resources.
Brenk *et al.* ([_Chem. Med. Chem._ (2008), **3**, 435-44](https://onlinelibrary.wiley.com/doi/full/10.1002/cmdc.200700139)) have assembled a list of unfavorable substructures to filter their libraries used to screen for compounds to treat neglected diseases. Examples of such unwanted features are nitro groups (mutagenic), sulfates and phosphates (likely resulting in unfavorable pharmacokinetic properties), 2-halopyridines and thiols (reactive). This list of undesired substructures was published in the above mentioned paper and will be used in the practical part of this talktorial.
### Pan Assay Interference Compounds (PAINS)
[PAINS](https://en.wikipedia.org/wiki/Pan-assay_interference_compounds) are compounds that often occur as hits in HTS even though they actually are false positives. PAINS show activity at numerous targets rather than one specific target. Such behavior results from unspecific binding or interaction with assay components. Baell *et al.* ([_J. Med. Chem._ (2010), **53**, 2719-2740](https://pubs.acs.org/doi/abs/10.1021/jm901137j)) focused on substructures interfering in assay signaling. They described substructures which can help to identify such PAINS and provided a list which can be used for substructure filtering.

Figure 1: Specific and unspecific binding in the context of PAINS. Figure taken from [Wikipedia](https://commons.wikimedia.org/wiki/File:PAINS_Figure.tif).
## Practical
### Load and visualize data
First, we import the required libraries, load our filtered dataset from **Talktorial T002** and draw the first molecules.
```
from pathlib import Path
import pandas as pd
from tqdm.auto import tqdm
from rdkit import Chem
from rdkit.Chem import PandasTools
from rdkit.Chem.FilterCatalog import FilterCatalog, FilterCatalogParams
# define paths
HERE = Path(_dh[-1])
DATA = HERE / "data"
# load data from Talktorial T2
egfr_data = pd.read_csv(
HERE / "../T002_compound_adme/data/EGFR_compounds_lipinski.csv",
index_col=0,
)
# Drop unnecessary information
print("Dataframe shape:", egfr_data.shape)
egfr_data.drop(columns=["molecular_weight", "n_hbd", "n_hba", "logp"], inplace=True)
egfr_data.head()
# Add molecule column
PandasTools.AddMoleculeColumnToFrame(egfr_data, smilesCol="smiles")
# Draw first 3 molecules
Chem.Draw.MolsToGridImage(
list(egfr_data.head(3).ROMol),
legends=list(egfr_data.head(3).molecule_chembl_id),
)
```
### Filter for PAINS
The PAINS filter is already implemented in RDKit ([documentation](http://rdkit.org/docs/source/rdkit.Chem.rdfiltercatalog.html)). Such pre-defined filters can be applied via the `FilterCatalog` class. Let's learn how it can be used.
```
# initialize filter
params = FilterCatalogParams()
params.AddCatalog(FilterCatalogParams.FilterCatalogs.PAINS)
catalog = FilterCatalog(params)
# search for PAINS
matches = []
clean = []
for index, row in tqdm(egfr_data.iterrows(), total=egfr_data.shape[0]):
molecule = Chem.MolFromSmiles(row.smiles)
entry = catalog.GetFirstMatch(molecule) # Get the first matching PAINS
if entry is not None:
# store PAINS information
matches.append(
{
"chembl_id": row.molecule_chembl_id,
"rdkit_molecule": molecule,
"pains": entry.GetDescription().capitalize(),
}
)
else:
# collect indices of molecules without PAINS
clean.append(index)
matches = pd.DataFrame(matches)
egfr_data = egfr_data.loc[clean] # keep molecules without PAINS
# NBVAL_CHECK_OUTPUT
print(f"Number of compounds with PAINS: {len(matches)}")
print(f"Number of compounds without PAINS: {len(egfr_data)}")
```
Let's have a look at the first 3 identified PAINS.
```
Chem.Draw.MolsToGridImage(
list(matches.head(3).rdkit_molecule),
legends=list(matches.head(3)["pains"]),
)
```
### Filter and highlight unwanted substructures
Some lists of unwanted substructures, like PAINS, are already implemented in RDKit. However, it is also possible to use an external list and get the substructure matches manually.
Here, we use the list provided in the supporting information from Brenk *et al.* ([_Chem. Med. Chem._ (2008), **3**, 535-44](https://onlinelibrary.wiley.com/doi/full/10.1002/cmdc.200700139)).
```
substructures = pd.read_csv(DATA / "unwanted_substructures.csv", sep="\s+")
substructures["rdkit_molecule"] = substructures.smarts.apply(Chem.MolFromSmarts)
print("Number of unwanted substructures in collection:", len(substructures))
# NBVAL_CHECK_OUTPUT
```
Let's have a look at a few substructures.
```
Chem.Draw.MolsToGridImage(
mols=substructures.rdkit_molecule.tolist()[2:5],
legends=substructures.name.tolist()[2:5],
)
```
Search our filtered dataframe for matches with these unwanted substructures.
```
# search for unwanted substructure
matches = []
clean = []
for index, row in tqdm(egfr_data.iterrows(), total=egfr_data.shape[0]):
molecule = Chem.MolFromSmiles(row.smiles)
match = False
for _, substructure in substructures.iterrows():
if molecule.HasSubstructMatch(substructure.rdkit_molecule):
matches.append(
{
"chembl_id": row.molecule_chembl_id,
"rdkit_molecule": molecule,
"substructure": substructure.rdkit_molecule,
"substructure_name": substructure["name"],
}
)
match = True
if not match:
clean.append(index)
matches = pd.DataFrame(matches)
egfr_data = egfr_data.loc[clean]
# NBVAL_CHECK_OUTPUT
print(f"Number of found unwanted substructure: {len(matches)}")
print(f"Number of compounds without unwanted substructure: {len(egfr_data)}")
```
### Highlight substructures
Let's have a look at the first 3 identified unwanted substructures. Since we have access to the underlying SMARTS patterns we can highlight the substructures within the RDKit molecules.
```
to_highlight = [
row.rdkit_molecule.GetSubstructMatch(row.substructure) for _, row in matches.head(3).iterrows()
]
Chem.Draw.MolsToGridImage(
list(matches.head(3).rdkit_molecule),
highlightAtomLists=to_highlight,
legends=list(matches.head(3).substructure_name),
)
```
### Substructure statistics
Finally, we want to find the most frequent substructure found in our data set. The Pandas `DataFrame` provides convenient methods to group containing data and to retrieve group sizes.
```
# NBVAL_CHECK_OUTPUT
groups = matches.groupby("substructure_name")
group_frequencies = groups.size()
group_frequencies.sort_values(ascending=False, inplace=True)
group_frequencies.head(10)
```
## Discussion
In this talktorial we learned two possibilities to perform a search for unwanted substructures with RDKit:
* The `FilterCatalog` class can be used to search for predefined collections of substructures, e.g., PAINS.
* The `HasSubstructMatch()` function to perform manual substructure searches.
Actually, PAINS filtering could also be implemented via manual substructure searches with `HasSubstructMatch()`. Furthermore, the substructures defined by Brenk *et al.* ([_Chem. Med. Chem._ (2008), **3**, 535-44](https://onlinelibrary.wiley.com/doi/full/10.1002/cmdc.200700139)) are already implemented as a `FilterCatalog`. Additional pre-defined collections can be found in the RDKit [documentation](http://rdkit.org/docs/source/rdkit.Chem.rdfiltercatalog.html).
So far, we have been using the `HasSubstructMatch()` function, which only yields one match per compound. With the `GetSubstructMatches()` function ([documentation](https://www.rdkit.org/docs/source/rdkit.Chem.rdchem.html)) we have the opportunity to identify all occurrences of a particular substructure in a compound.
In case of PAINS, we have only looked at the first match per molecule (`GetFirstMatch()`). If we simply want to filter out all PAINS this is enough. However, we could also use `GetMatches()` in order to see all critical substructures of a molecule.
Detected substructures can be handled in two different fashions:
* Either, the substructure search is applied as a filter and the compounds are excluded from further testing to save time and money.
* Or, they can be used as warnings, since ~5 % of FDA-approved drugs were found to contain PAINS ([_ACS. Chem. Biol._ (2018), **13**, 36-44](https://pubs.acs.org/doi/10.1021/acschembio.7b00903)). In this case experts can judge manually, if an identified substructure is critical or not.
## Quiz
* Why should we consider removing "PAINS" from a screening library? What is the issue with these compounds?
* Can you find situations when some unwanted substructures would not need to be removed?
* How are the substructures we used in this tutorial encoded?
| github_jupyter |
```
from recOrder.compute.qlipp_compute import initialize_reconstructor, reconstruct_qlipp_stokes, reconstruct_phase3D, \
reconstruct_phase2D, reconstruct_qlipp_birefringence
import numpy as np
```
## Setup Fake Data and Initialize Reconstructor
```
data = np.random.random((1, 41, 256, 256)) # (C, Z, Y, X) where C=0 is BF data
image_dim = (data.shape[-2], data.shape[-1]) # (Y, X)
NA_obj = 0.4 # Numerical Aperture of Objective
NA_illu = 0.2 # Numerical Aperture of Condenser
wavelength = 532 # wavelength in nm
n_objective_media = 1.0 # refractive index of objective immersion media
mag = 20 # magnification
n_slices = data.shape[-3] # number of slices in z-stack
z_step_um = 0.25 # z-step size in um
pad_z = 0 # slices to pad for phase reconstruction boundary artifacts
pixel_size_um = 6.5 # pixel size in um
mode = '3D' # phase reconstruction mode, '2D' or '3D'
use_gpu = False
gpu_id = 0
# Initialize Reconstructor
reconstructor = initialize_reconstructor(pipeline='PhaseFromBF',
image_dim=image_dim,
NA_obj=NA_obj,
NA_illu=NA_illu,
wavelength_nm=wavelength,
n_obj_media=n_objective_media,
mag=mag,
n_slices=n_slices,
z_step_um=z_step_um,
pad_z=pad_z,
pixel_size_um=pixel_size_um,
mode=mode,
use_gpu=use_gpu,
gpu_id=gpu_id)
```
## Reconstruct Phase3D from BF
```
S0 = np.transpose(data[0], (1, 2, 0)) # Need to transpose BF into (Y, X, Z)
phase3D = reconstruct_phase3D(S0, reconstructor, method='Tikhonov', reg_re=1e-4)
print(f'Shape of 3D phase data: {np.shape(phase3D)}')
```
## Reconstruct Phase2D from BF
we need a new reconstructor to compute phase 2D
```
mode = '2D'
# Initialize Reconstructor
reconstructor = initialize_reconstructor(pipeline='PhaseFromBF',
image_dim=image_dim,
NA_obj=NA_obj,
NA_illu=NA_illu,
wavelength_nm=wavelength,
n_obj_media=n_objective_media,
mag=mag,
n_slices=n_slices,
z_step_um=z_step_um,
pad_z=pad_z,
pixel_size_um=pixel_size_um,
mode=mode,
use_gpu=use_gpu,
gpu_id=gpu_id)
phase2D = reconstruct_phase2D(S0, reconstructor, method='Tikhonov', reg_p=1e-4)
print(f'Shape of 2D phase data: {np.shape(phase2D)}')
```
| github_jupyter |
# Workshop PR02: Machine Learning Kickoff
## Agenda
- Preparing the data for modelling
- Introduction to ML frameworks and algorithms
- Supervised Learning: sklearn
- Deep Learning: tensorflow
## Previously on the last workshop
From last workshop we know how to do the following:
- Read the data: IEEE fraud detection dateset [(download it here)](https://www.kaggle.com/c/ieee-fraud-detection/data)
- Join the identify and transaction dataset together
## Exercise
- find out whether feature selection would boost performance
If you have forgotten, these are from the last workshop:
```
import pandas as pd
import numpy as np
# read csv file into a dataframe
df_id_train = pd.read_csv("train_identity.csv")
df_tran_train = pd.read_csv("train_transaction.csv")
df_id_test = pd.read_csv("test_identity.csv")
df_tran_test = pd.read_csv("test_transaction.csv")
# joining table
df_train = pd.merge(df_tran_train,df_id_train, on='TransactionID' ,how='left')
df_train.info()
# target dataframe
Y_train = df_train['isFraud']
Y_train = pd.DataFrame(Y_train)
Y_train.head()
# dropping the irrelevant data for training
list = ['isFraud','TransactionID','DeviceInfo']
X_train = df_train.drop(list, axis=1)
X_train.head()
X_train.info()
```
## Prepping the data
- Encode string data into float
- Remove low quality data/engineer high quality data (Feature Selection/Engineering)
### String Encoding
There are many (30) features in string format, so we need to encode them into float before we can use them for training.
```
obj_df = X_train.select_dtypes(include=['object']).copy()
int_df = X_train.select_dtypes(include=['int64']).copy()
float_df = X_train.select_dtypes(include=['float64']).copy()
for column in obj_df.head(0):
obj_df[column] = obj_df[column].astype('category')
obj_df[column] = obj_df[column].cat.codes
X_train = pd.concat([obj_df,int_df,float_df],axis=1, sort=False)
X_train.head()
```
Now we can see which objects are encoded into int. Note: -1 means NaN.
```
obj_df.info()
```
Now to be uniform, we'll change all NaN into -1 in other non-object columns as well.
```
X_train.fillna(value=-1,inplace=True)
print(X_train.isnull().values.any()) # False means there isn't any NaN
```
### Feature Selection
There are way too many features (over 400!) to train the model efficiently, so we're going to narrow down to the more important features. Now there are many ways to shrink our high-dimensional dataset, and we will show the followings:
- Variance Threshold
- Univariate Feature Selection
- Recursive Feature Elimination
- Select From Model (linear-based, tree-based)
- As a part of pipeline
Mote detail: https://scikit-learn.org/stable/modules/feature_selection.html
#### 1) Variance Threshols
Removes the features that have low variance. For example, the default is to remove all features with the same entries.
```
from sklearn.feature_selection import VarianceThreshold
# for boolean features, we want to remove all features that
# are either one or zero (on or off) in more than 80% of the samples
VT = VarianceThreshold(threshold=(.8 * (1 - .8))) # variance of Bernoulli is p(1-p)
# for multinomial variables, variance depends on the number of categories,
# but for convenience we'll use the same formula
X_train_VT = VT.fit_transform(X_train)
X_train_VT.shape
```
OK now we have reduced to 393 features. Before we move to the next method of feature selection, let's fit into a model (we use Random Forest here) and see how it goes.
```
from sklearn.ensemble import RandomForestClassifier
random_forest = RandomForestClassifier()
random_forest.fit(X_train_VT, Y_train)
random_forest.score(X_train_VT, Y_train)
random_forest.fit(X_train, Y_train) # this is without feature selection
random_forest.score(X_train, Y_train)
```
Not bad eh, but don't get too excited now since this is only train error. When we move to the next workshop we will show how we should train models properly by using test error.
#### 2) Univariate Feature Selection
Univariate feature selection selects the best features based on univariate statistical tests. It can be seen as a preprocessing step to an estimator. What statistical test is used to score the features depends on whether it's a regression or classification problem.
- For regression: `f_regression` ,`mutual_info_regression`
- For classification: `chi2`, `f_classif`, `mutual_info_classif`
Also you can choose how to select the features based on the scores:
- `SelectKBest`: select the k highest scoring features
- `SelectPercentile`: select the user-specified highest scoring percentage of features
- Common univariate statistical tests for each feature: false positive rate `SelectFpr`, false discovery rate `SelectFdr`, or family wise error `SelectFwe`
- `GenericUnivariateSelect`: "allows to perform univariate feature selection with a configurable strategy. This allows to select the best univariate selection strategy with hyper-parameter search estimator"
```
from sklearn.feature_selection import SelectKBest, f_classif
# select the 200 best features based on ANOVA F-value
X_train_Kbest = SelectKBest(f_classif, k=200).fit_transform(X_train, Y_train)
X_train_Kbest.shape
# so what are the first 20 features
X_train.columns[SelectKBest(f_classif, k=20).fit(X_train, Y_train).get_support()]
```
Now apart from choosing the K highest scores features, we can also choose according to a percentile of the highest scores (e.g. top 50% of the best features).
```
from sklearn.feature_selection import SelectPercentile
X_train_Pbest = SelectPercentile(f_classif, percentile=50).fit_transform(X_train, Y_train)
X_train_Pbest.shape
```
So 215 features are chosen based on the 50% percentile.
#### 3) Recursive Feature Elimination
Given an external estimator that assigns weights to features (e.g., the coefficients of a linear model), recursive feature elimination ([RFE](https://scikit-learn.org/stable/modules/generated/sklearn.feature_selection.RFE.html#sklearn.feature_selection.RFE) or [RFECV](https://scikit-learn.org/stable/modules/generated/sklearn.feature_selection.RFECV.html#sklearn.feature_selection.RFECV)) selects features by recursively considering smaller and smaller sets of features untill the desired number of features is reached:
1. The chosen estimator is trained on the initial set of features and the importance of each feature is obtained either through a `coef_` attribute or through a `feature_importances_` attribute.
2. The least important features are pruned from current set of features.
3. Repeat 1 and 2 on the pruned set until the desired number of features is obbtained.
RFECV performs RFE in a cross-validation loop to find the optimal number of features, i.e. it removes all the features and calculates the CV error.
```
# this cell is optional: run this if you don't want to read the future warning messages
# import warnings filter
from warnings import simplefilter
# ignore all future warnings
simplefilter(action='ignore', category=FutureWarning)
# this is to stop the format warning
Y_train = np.array(Y_train).ravel()
```
The following cell might take a while (5 minutes), while it's running have a break! Or skip this part.
```
from sklearn.feature_selection import RFE, RFECV
from sklearn.linear_model import SGDClassifier
import matplotlib.pyplot as plt
estimator = SGDClassifier()
rfecv = RFECV(estimator,verbose=True,
cv=3,step=0.1)
#step means the number/percentage of features to be removed each loop
X_train_refcv = rfecv.fit_transform(X_train, Y_train)
print("Optimal number of features : %d" % rfecv.n_features_)
# Plot number of features VS. cross-validation scores
plt.figure()
plt.xlabel("Number of features selected")
plt.ylabel("Cross validation score (num of correct classifications)")
plt.plot(range(0, 40*len(rfecv.grid_scores_),40), rfecv.grid_scores_)
plt.show()
```
#### 4) Feature Selection using `SelectFromModel`
As the name suggests, features are selected from a model that assigns importance to features. `SelectFromModel` is a meta-transformer that can be used along with any estimator that has a `coef_` or `feature_importances_` attribute after fitting. The unimportant features are remoced if the corresponding `coef_` or `feature_importances_` values are below the certain threshold. Apart from specifying the threshold numerically, there are built-in heuristics for finding a threshold using a string argument. Available heuristics are “mean”, “median” and float multiples of these like “0.1*mean”.
```
from sklearn.feature_selection import SelectFromModel
from sklearn.linear_model import SGDClassifier
model_sgd = SGDClassifier() # model
sfm = SelectFromModel(model_sgd) # assigning model
sfm.fit(X_train, Y_train) # fitting data
X_train_sgd = sfm.transform(X_train) # transform full dataset to reduced dataset
features_selected = X_train.columns[sfm.get_support()] # get the selected features
print(X_train_sgd.shape)
print(features_selected)
from sklearn.ensemble import RandomForestClassifier
model_rf = RandomForestClassifier().fit(X_train, Y_train)
X_train_rf = SelectFromModel(model_rf,prefit=True).transform(X_train)
X_train_rf.shape
```
#### 5) Feature Selection as Part of a Pipeline
Pipeline of transforms with a final estimator.
Feature selection is usually used as a pre-processing step before doing the actual learning. The recommended way to do this in scikit-learn is to use a `sklearn.pipeline.Pipeline`.
The following part is straight from [this link](https://scikit-learn.org/stable/modules/generated/sklearn.pipeline.Pipeline.html#sklearn.pipeline.Pipeline):
Sequentially apply a list of transforms and a final estimator. Intermediate steps of the pipeline must be ‘transforms’, that is, they must implement fit and transform methods. The final estimator only needs to implement fit. The transformers in the pipeline can be cached using `memory` argument.
The purpose of the pipeline is to assemble several steps that can be cross-validated together while setting different parameters. For this, it enables setting parameters of the various steps using their names and the parameter name separated by a ‘__’. A step’s estimator may be replaced entirely by setting the parameter with its name to another estimator, or a transformer removed by setting it to ‘passthrough’ or `None`.
```
from sklearn.pipeline import Pipeline
clf = Pipeline([
('feature_selection', SelectFromModel(model_sgd)),
#feature_selection can be any method mentioned above
('classification', RandomForestClassifier())
#to be consistent we use random forest classifier again
])
clf.fit(X_train, Y_train)
clf.score(X_train, Y_train)
```
That's it for this workshop!
So we have learned some methods of dealing with high dimemsional data, but also bear in mind that sometimes when we have smaller dataset (less than 10 features or so), instead of **feature selection** we need to do **feature engineering**, which could be as simple as creating simple features such as speed (if we are given time and distance), or it could be as complicated as obtaining new data such as distance to the city (if we are given locations).
The reason why we spend some much effort on prepping the data is because of the well-known "garbage-in-garbage-out", as well as the intention of training the model more efficiently and avoiding over-fitting.
Note that we have only shown classification example, for regression problems you might need to alter something (e.g. we can use lasso for regression) and the links provided should contain enough information to get you started. There are also other techniques for data reduction such as [PCA](https://scikit-learn.org/stable/modules/generated/sklearn.decomposition.PCA.html#sklearn.decomposition.PCA) but we'll just leave that part for you to explore as this could be done quiet easily.
Next workshop we will dive into model training. (Yay!)
| github_jupyter |
```
import numpy as np
import json
import pandas as pd
from scipy import interpolate
from gensim.models import KeyedVectors
import pickle
import re
from nltk.corpus import stopwords
from nltk import word_tokenize
from nltk import pos_tag
from string import punctuation,digits
import os
import pickle
from sklearn import preprocessing
import tensorflow as tf
from scipy.interpolate import interp1d
from keras.utils import to_categorical
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
a=[]
for i in list(punctuation):
if i=='%':
continue
else:
a.append(i)
def remove_punctuation(s):
list_punctuation = a
for i in list_punctuation:
s = s.replace(i,'')
return s
def clean_sentence(sentence):
sentence = sentence.lower()
sentence = re.sub(r'(\W)\1{2,}', r'\1', sentence)
sentence = re.sub(r'(\w)\1{2,}', r'\1\1', sentence)
sentence = re.sub(r'^https?:\/\/.*[\r\n]*', r'', sentence)
sentence = re.sub(r'(?P<url>http?://[^\s]+)', r'', sentence)
sentence = re.sub(r"\@(\w+)", "", sentence)
sentence = re.sub(r"$ ", "", sentence)
sentence = sentence.replace('#',' ')
sentence = sentence.replace("'s",' ')
sentence = sentence.replace("-",' ')
tokens = sentence.split()
tokens = [remove_punctuation(w) for w in tokens]
stop_words = set(stopwords.words('english'))
tokens = [w for w in tokens if not w in stop_words]
#remove_digits = str.maketrans('', '', digits)
tokens = [w.translate(tokens) for w in tokens]
tokens = [w.strip() for w in tokens]
tokens = [w for w in tokens if w!=""]
tokens = [i.replace("ocompany","OCOMPANY") for i in tokens]
tokens = [i.replace("company","COMPANY") for i in tokens]
tokens = ' '.join(tokens)
return tokens
def extract_domain_specific_features(text):
features=[]
for i in text.split():
match=re.search("(^|[ \t])([-+$]?(\d+|\.\d+|\d+\.\d*))($|[^+-.]*)", i)
if match:
features.append(match.group(0))
return features
print(extract_domain_specific_features("Hello $123 this"))
typos={'NewsMorrisons':'News Morrissons',
'Taylor Wimpey':'Tailor Wimpey',
'AB InBev':'AB InBrev',
'Aberdeen Asset Management':'Aberdeen Asset Managment',
'International Business Machines Corp. ':'IBM',
}
target_typos={
'shoeders':'schroders',
# 'jp morgan chase':'jpmorgan chase',
'gs':'companiesg4s',
''
}
def prepare_data(fname):
with open(fname, encoding='utf-8') as f:
foo = json.load(f)
sentence_l=[]
target_l=[]
aspect_l=[]
sentiment_l=[]
snippet_l=[]
features=[]
for key in foo.keys():
for info in foo[key]['info']:
#print(info)
sentence=foo[key]['sentence']
z=sentence
target= info['target']
c=[]
#print(sentence)
sentence = sentence.replace("'s",' ')
for typo,correct in typos.items():
if typo in sentence:
#print(sentence)
sentence=re.sub(typo,correct,sentence)
#print(sentence)
#sentence=re.sub(target.lower(),'COMPANY',sentence)
#sentence = re.sub(r"[$]?(\w+)", "OCOMPANY", sentence)
features.append(extract_domain_specific_features(sentence))
sentence=sentence.lower()
p=[]
for i in sentence.partition(target.lower()):
if i==target.lower():
p.append('COMPANY')
else:
p.append(i)
p=" ".join(p)
p=p.strip()
l=[]
for i in p.split(" "):
if i.startswith('$'):
if not i.lstrip('$').isalpha():
l.append('')
elif not i.lstrip('$').isnumeric():
l.append("OCOMPANY")
else:
l.append(i)
l=" ".join(l)
l=l.strip()
sentence = [clean_sentence(x) for x in l.split(" ")]
#print(sentence)
sentence=' '.join(sentence)
if 'COMPANY' not in sentence.split():
k=[]
#print("STILL NOT SOLVED 1")
#print(sentence,"AND",target)
#if sentence=='xli potential intermediate top uptrend charts tdf':
# sentence=re.sub('xli','COMPANY',sentence)
new=target.lower()[:3]
for i in sentence.split():
#print(i)
if i.startswith(new):
k.append('COMPANY')
else :
k.append(i)
k=' '.join(k)
sentence=k
#print(z,"AND",target)
#print(sentence)
#if 'COMPANY' not in sentence.split(" ") :
#print("NOT SOLVED :",sentence, " AND ",target.lower())
# sentence=re.sub('OCOMPANY','COMPANY',sentence)
# if 'COMPANY' in sentence.split(" ") : print("SOLVED :",sentence)
if 'COMPANY' not in sentence.split(" "):
# print("STILL NOT SOLVED 2")
# print(sentence,"AND",target)
if sentence.strip().startswith('gemalto shares slump third profit'):
sentence=re.sub('gemalto','COMPANY',sentence)
if sentence.strip().startswith('coca cola shares rise earnings beat expectations'):
sentence=re.sub('coca cola','COMPANY',sentence)
if sentence.strip().startswith('alphabet revenue 21 percent'):
sentence=re.sub('alphabet','COMPANY',sentence)
if sentence.startswith('price comparison site slumps 4 y'):
#sentence=re.sub('companiescoutts','companies coutts',sentence)
sentence='COMPANY '+sentence
'''
for typo,correct in target_typos.items():
if typo ==target.lower():
sentence=re.sub(correct,'COMPANY',sentence)
'''
#if 'COMPANY' not in sentence.split(" ") :
# sentence='COMPANY '+sentence+ " AND ALS0 "+ target
if 'COMPANY' not in sentence.split(" ") : print('NOT solved',z,"#####", target,"@@@@",sentence)
if sentence.strip().startswith('industry newsrevenue earnings take'):
sentence=re.sub('newsrevenue','news revenue',sentence)
snippet=info['snippets'].lstrip('[')
snippet=snippet.rstrip(']')
snippet=snippet.lower()
#print(snippet)
sentiment_score = info['sentiment_score']
#print(sentiment_score)
aspect= info['aspects']
#print(aspect)
asp=aspect[0].lstrip("['")
asp=asp.rstrip("']")
#print(asp)
l=asp.split("/")
#print(l)
aspect=l[1]
#print(aspect)
sentence=re.sub(' +', ' ',sentence)
sentence=sentence.strip()
#print(sentence)
sentence_l.append(sentence)
target_l.append(target)
sentiment_l.append(sentiment_score)
aspect_l.append(aspect)
s=snippet.lstrip('\'')
s=s.rstrip('\'')
#print(s)
snippet_l.append(s)
return sentence_l,target_l,sentiment_l,aspect_l,features,snippet_l
print("preparing Finance dataset...")
fname = {
'finance': {
'train_headline': r'D:\Datasets\FinanceHeadlineDataset\train\headline.json',
'train_post': r'D:\Datasets\FinanceHeadlineDataset\train\post.json',
'test_headline': r'D:\Datasets\FinanceHeadlineDataset\test\headline.json',
'test_post': r'D:\Datasets\FinanceHeadlineDataset\test\post.json',
'validation_post' : r'D:/PythonCodes/Sentiment-Analysis/Data/validation_post_1.json',
'validation_headline' :r'D:/PythonCodes/Sentiment-Analysis/Data/validation_headline_1.json'
},
'Data_Augmentation':{
'sentiment':{
'train_headline':r'D:\PythonCodes\Sentiment-Analysis\Data\Headline_Train.json',
'train_post':r'D:\PythonCodes\Sentiment-Analysis\Data\Microblog_Trainingdata.json',
'validation_headline':'D:\PythonCodes\Sentiment-Analysis\Data\Headline_Trialdata.json',
'validation_post':'D:\PythonCodes\Sentiment-Analysis\Data\Microblog_Trialdata.json'
},
'aspect':{
'train_data':'train_data_augmented.dat'
}
}
}
```
# TRAIN
```
#headlines=pd.DataFrame(data={'sentence':h_sentence,'sentiment':h_sentiment,'aspect':h_aspect,'snippet':h_snippet,'features':h_features})
#pickle.dump(headlines,open('headlines.data',"wb"))
headlines=pickle.load(open('headlines.data',"rb"))
headlines.head()
post=pickle.load(open('post.dat',"rb"))
post.iloc[650]
#h_sentence,h_target,h_sentiment,h_aspect,h_snippet,h_features=prepare_data(fname['finance']['train_headline'])
#p_sentence,p_target,p_sentiment,p_aspect,p_snippet,p_features=prepare_data(fname['finance']['train_post'])
#p_sentence[:1],p_target[:1]
#post=pd.DataFrame(data={'sentence':p_sentence,'sentiment':p_sentiment,'aspect':p_aspect,'snippet':p_snippet,'features':p_features})
#pickle.dump(post,open('post.dat',"wb"))
for i in p_sentence:
h_sentence.append(i)
for i in p_sentiment:
h_sentiment.append(i)
for i in p_aspect:
h_aspect.append(i)
for i in p_snippet:
h_snippet.append(i)
train_data=pd.DataFrame(data={'sentence': h_sentence,'sentiment':h_sentiment,'aspect':h_aspect,'snippet':h_snippet})
pickle.dump(train_data,open("train_data_initial.dat","wb"))
train_data.head()
p_sentence[:2],p_target[:2],p_sentiment[:2],p_aspect[:2],p_snippet[:2],p_features[:2]
```
# TEST
```
t_h_sentence,t_h_target,t_h_sentiment,t_h_aspect,t_h_features,t_h_snippet=prepare_data(fname['finance']['validation_headline'])
head=pd.DataFrame(data={'sentence':t_h_sentence,'sentiment':t_h_sentiment,'aspect':t_h_aspect,'snippets':t_h_snippet})
pickle.dump(head,open('test_head.dat',"wb"))
post=pd.DataFrame(data={'sentence':t_p_sentence,'sentiment':t_p_sentiment,'aspect':t_p_aspect,'snippets':snippet})
pickle.dump(post,open('test_post.dat',"wb"))
t_p_sentence,t_p_target,t_p_sentiment,t_p_aspect,t_p_features,snippet=prepare_data(fname['finance']['validation_post'])
t_p_sentence
headlines = pickle.load(open('test_head.dat',"rb"))
test_sentence=[]
test_sentiment=[]
test_aspect=[]
test_sentence,test_sentiment,test_aspect=headlines['sentence'],headlines['sentiment'],headlines['aspect']
for i in headlines['sentence']:
test_sentence.append(i)
for i in headlines['sentiment']:
test_sentiment.append(i)
for i in headlines['aspect']:
test_aspect.append(i)
len(test_sentence)
for i in t_p_sentence:
test_sentence.append(i)
for i in t_p_sentiment:
test_sentiment.append(i)
for i in t_p_aspect:
test_aspect.append(i)
Validation_data=pd.DataFrame(data={'sentence': test_sentence,'sentiment':test_sentiment,'aspect':test_aspect})
Validation_data.head()
pickle.dump(Validation_data,open("Final_ValidationData.dat","wb"))
t_p_sentence[:1],t_p_target[:1],t_p_sentiment[:1],t_p_aspect[:1],t_p_features[:1]
```
# DATA AUGMENT
```
h_sentence_1=[]
h_target_1=[]
h_sentiment_1=[]
p_sentence_1=[]
p_target_1=[]
p_sentiment_1=[]
def prepare_data1(fname):
with open(fname, encoding='utf-8') as f:
foo = json.load(f)
sentence_l=[]
target_l=[]
sentiment_l=[]
features=[]
for info in foo:
sentence=info['title']
target= info['company']
features.append(extract_domain_specific_features(sentence))
#print(sentence," +Tar+ ",target)
m=[]
for i in sentence.partition(target):
if i==target:
m.append('COMPANY')
else : m.append(i)
m=' '.join(m)
k=[]
for i in m.split():
if i.lstrip('$')==target[0]:
k.append('COMPANY')
elif i ==target[0]:
k.append('COMPANY')
else:
k.append(i)
k=' '.join(k)
sentence = [clean_sentence(x) for x in k.split(" ")]
#print(sentence)
sentence=' '.join(sentence)
sentiment_score = info['sentiment']
#print(sentiment_score)
sentence=re.sub(' +', ' ',sentence)
sentence=sentence.strip()
sentence_l.append(sentence)
target_l.append(target)
sentiment_l.append(sentiment_score)
return sentence_l,target_l,sentiment_l,features
h_sentence_1,h_target_1,h_sentiment_1,h_features_1=prepare_data1(fname['Data_Augmentation']['sentiment']['validation_headline'])
h_sentence_1[:5],h_target_1[:5],h_sentiment_1[:5],h_features_1[:5]
h_sentence_2,h_target_2,h_sentiment_2,h_features_2=prepare_data1(fname['Data_Augmentation']['sentiment']['train_headline'])
def prepare_data2(fname):
with open(fname, encoding='utf-8') as f:
foo = json.load(f)
sentence_l=[]
target_l=[]
sentiment_l=[]
features=[]
for info in foo:
sentence=info['spans'][0]
#print(sentence)
target= info['cashtag']
features.append(extract_domain_specific_features(sentence))
m=[]
for i in sentence.partition(target):
if i==target:
m.append('COMPANY')
else : m.append(i)
m=' '.join(m)
#target= info['cashtag'].lstrip("$")
#print(target)
sentiment_score = info['sentiment score']
#print(sentiment_score)
sentence_l.append(m)
target_l.append(target)
sentiment_l.append(sentiment_score)
return sentence_l,target_l,sentiment_l,features
p_sentence_1,p_target_1,p_sentiment_1,p_features_1=prepare_data2(fname['Data_Augmentation']['sentiment']['validation_post'])
p_sentence_1[:1],p_target_1[:1],p_sentiment_1[:1],p_features_1[:1]
p_sentence_2,p_target_2,p_sentiment_2,p_features_2=prepare_data2(fname['Data_Augmentation']['sentiment']['train_post'])
len(h_sentence), len(p_sentence)
len(h_sentence_1), len(p_sentence_1)
len(h_sentence_2), len(p_sentence_2)
```
# Combine ALL
```
#Sentence
for i in h_sentence_1:
h_sentence.append(i)
for i in h_sentence_2:
h_sentence.append(i)
for i in p_sentence_1:
p_sentence.append(i)
for i in p_sentence_2:
p_sentence.append(i)
#Sentiment
for i in h_sentiment_1:
h_sentiment.append(i)
for i in h_sentiment_2:
h_sentiment.append(i)
for i in p_sentiment_1:
p_sentiment.append(i)
for i in p_sentiment_2:
p_sentiment.append(i)
#Features
for i in h_features_1:
h_features.append(i)
for i in h_features_2:
h_features.append(i)
for i in p_features_1:
p_features.append(i)
for i in p_features_2:
p_features.append(i)
len(h_sentence), len(p_sentence)
for i in p_sentence:
h_sentence.append(i)
for i in p_sentiment:
h_sentiment.append(i)
for i in p_features:
h_features.append(i)
len(h_sentence)
data={'sentence':h_sentence,
'sentiment':h_sentiment,
'features':h_features}
pickle.dump(data,open("Final_TrainData.dat","wb"))
data=pickle.load(open("Final_TrainData.dat","rb"))
len(data['sentence'])
```
# Word Embedding
```
def load_google_word2vec(file_name):
return KeyedVectors.load_word2vec_format(file_name, binary=True)
def build_embedding_matrix(vocab_size, embed_dim, tokenizer ):
embedding_matrix_file_name='D:/PythonCodes/Sentiment-Analysis/Data/embedding_matrix_sentiment.dat'
if os.path.exists(embedding_matrix_file_name):
print('loading embedding_matrix:', embedding_matrix_file_name)
embedding_matrix = pickle.load(open(embedding_matrix_file_name, 'rb'))
else:
print('loading word vectors...')
fname = 'D:\PythonCodes\Jupyter notebooks\Word Embeddings\GoogleNews-vectors-negative300.bin'
model=load_google_word2vec(fname)
embedding_matrix = np.zeros((vocab_size, embed_dim))
for word, i in tokenizer.word_index.items():
try:
embedding_vector = model[word]
except KeyError:
embedding_vector = None
if embedding_vector is not None:
embedding_matrix[i]=embedding_vector
pickle.dump(embedding_matrix, open(embedding_matrix_file_name, 'wb'))
return embedding_matrix
embedding_matrix_file_name='D:/PythonCodes/Sentiment-Analysis/Data/embedding_matrix_sentiment.dat'
if os.path.exists(embedding_matrix_file_name):
print('loading embedding_matrix:', embedding_matrix_file_name)
embedding_matrix = pickle.load(open(embedding_matrix_file_name, 'rb'))
len(embedding_matrix)
```
# SENTIMENT RESCALING
```
def rescale(series,old_range,new_range):
m = interp1d(old_range,new_range)
return [float(m(x)) for x in series]
sentiment = rescale(data['sentiment'],[-1,1],[0,1])
head_sentiment = rescale(t_h_sentiment,[-1,1],[0,1])
post_sentiment = rescale(t_p_sentiment,[-1,1],[0,1])
```
# TOKENIZING
```
def create_tokenizer(lines):
tokenizer = Tokenizer()
tokenizer.fit_on_texts(lines)
return tokenizer
# encode a list of lines
def encode_text(tokenizer, lines, length):
encoded = tokenizer.texts_to_sequences(lines)
padded = pad_sequences(encoded, maxlen=length, padding='post')
return padded
import random
c = list(zip(data['sentiment'], data['sentence'],data['features']))
random.shuffle(c)
data['sentiment'], data['sentence'],data['features']= zip(*c)
tokenizer = create_tokenizer(data['sentence'])
vocab_size = len(tokenizer.word_index) + 1
vocab_size
data['sentence'][:10]
target=[]
for i in data['sentence'] :
if "COMPANY" in i :
target.append('COMPANY')
else:
target.append("")
data['sentence'][5]
trainX[5]
max_length=15
trainX= encode_text(tokenizer, data['sentence'], max_length)
target= encode_text(tokenizer, target, 1)
trainY= sentiment
target=np.tile(target,15)
embedding_matrix = build_embedding_matrix(vocab_size, 300,tokenizer)
len(eval('embedding_matrix'))
head_X= encode_text(tokenizer, t_h_sentence, max_length)
postX =encode_text(tokenizer, t_p_sentence, max_length)
data_augmented_test={'data':data}
pickle.dump(data_augmented_test,open("Sentiment.dat","wb"))
data_for_sentiment={
'trainX': trainX,
'target':target,
'trainY':trainY,
'trainFeatures':h_features,
'HEAD_testX':head_X,
'HEAD_testY':head_sentiment,
'POST_testX':postX,
'POST_testY':post_sentiment,
'embedding_matrix':embedding_matrix,
'vocab_size':vocab_size
}
pickle.dump(data_for_sentiment,open("ALLdataForSentiment.dat","wb"))
```
| github_jupyter |
```
import igraph
import xml.etree.ElementTree as ElementTree
import numpy as np
from tqdm.notebook import tqdm
from scipy.spatial.transform import Rotation
from joblib import Parallel, delayed
import pickle
%matplotlib notebook
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
# define a mapping from single letter amino acid codes to three letter abbreviations
mappings = ('ala:A|arg:R|asn:N|asp:D|cys:C|gln:Q|glu:E|gly:G|his:H|ile:I|leu:L|lys:K|met:M|phe:F|pro:P|ser:S|thr:T|trp:W|tyr:Y|val:V').upper().split('|')
letter2aa = dict([m.split(':')[::-1] for m in mappings])
# load an amino acid sequence from a fasta file
with open('proteins/6ct4.fasta') as file:
fasta = file.read().strip().split('\n')[-1]
res_sequence = 'MALEK'
#res_sequence = fasta
pos_init_range = 3
print('protein fasta:', res_sequence)
atoms = []
bonds = []
external_bond_indices = []
# load the amber forcefield
forcefield = ElementTree.parse('forcefields/amber99sb.xml').getroot()
# iterate over the amino acids in the sequence and load details from the forcefield
for i, res in enumerate(tqdm(res_sequence, desc='loading atom and force definitions from the forcefield', ncols=850)):
external_bond_indices.append([])
atom_count = len(atoms)
# get the three letter code from the current amino acid letter
aa_name = letter2aa[res]
if i == 0:
# add an N to the first residue to signal that hydrogen has to be added to the amine group
aa_name = 'N' + aa_name
elif i == len(res_sequence) - 1:
# add a C to the last residue to signal that a terminal oxygen has to be added to the carboxyl group
aa_name = 'C' + aa_name
# iterate over the children for the current residue defined in the forcefield
for obj in forcefield.find(f'Residues/Residue[@name=\'{aa_name}\']'):
if obj.tag == 'Atom':
# processing an atom of the current residue
name = obj.get('name')
type_id = int(obj.get('type'))
# get the type traits for the current atom
atom_traits = forcefield[0][type_id].attrib
atom_class = atom_traits['class']
element = atom_traits['element']
mass = float(atom_traits['mass'])
# extract information about nonbonded forces for this atom
nonbonded_traits = forcefield[5][type_id].attrib
charge = float(nonbonded_traits.get('charge'))
sigma = float(nonbonded_traits.get('sigma'))
epsilon = float(nonbonded_traits.get('epsilon'))
# initialize the position randomly, could add more options here
pos = np.random.uniform(0, pos_init_range, size=3)
# add a dictionary containing all information about the atom to the atom list
atoms.append(dict(name=name, type_id=type_id, atom_class=atom_class, element=element,
index=len(atoms), residue_index=i, residue=aa_name, mass=mass,
charge=charge, sigma=sigma, epsilon=epsilon, pos=pos))
elif obj.tag == 'Bond':
# processing a harmonic bond in the current residue
from_index = atom_count + int(obj.get('from'))
to_index = atom_count + int(obj.get('to'))
# get the atoms between the bond exists, atoms for the current residue should be fully loaded
# at this point due to the ordering in the forcefield xml
from_atom = atoms[from_index]
to_atom = atoms[to_index]
# find the bond force definition for the current bond in the forcefield
bond = forcefield.find(f'HarmonicBondForce/Bond[@class1=\'{from_atom["atom_class"]}\'][@class2=\'{to_atom["atom_class"]}\']')
if bond is None:
# try again with the atom classes in the reversed order
bond = forcefield.find(f'HarmonicBondForce/Bond[@class1=\'{to_atom["atom_class"]}\'][@class2=\'{from_atom["atom_class"]}\']')
# add a dictionary containing all information about the current bond the the bond list
bonds.append(dict(from_index=from_index, to_index=to_index, length=float(bond.get('length')),
k=float(bond.get('k')), bond_type='internal'))
elif obj.tag == 'ExternalBond':
# add the atom indices of external bonding sites to the list
from_index = atom_count + int(obj.get('from'))
external_bond_indices[-1].append(dict(from_index=from_index))
if i > 0:
# create an external bond definition (bond between two residues)
from_index = external_bond_indices[i-1][-1]['from_index']
to_index = external_bond_indices[i][0]['from_index']
# get the involved atoms
from_atom = atoms[from_index]
to_atom = atoms[to_index]
# find the force definition for the current bond
bond = forcefield.find(f'HarmonicBondForce/Bond[@class1=\'{from_atom["atom_class"]}\'][@class2=\'{to_atom["atom_class"]}\']')
if bond is None:
# try again with the atom classes in the reversed order
bond = forcefield.find(f'HarmonicBondForce/Bond[@class1=\'{to_atom["atom_class"]}\'][@class2=\'{from_atom["atom_class"]}\']')
# add the external bond to the bond list
bonds.append(dict(from_index=from_index, to_index=to_index, length=float(bond.get('length')),
k=float(bond.get('k')), bond_type='external'))
graph = igraph.Graph()
# add all atoms to the graph as vertices
for atom in atoms:
v = graph.add_vertex(atom['name'])
v.update_attributes(atom)
# all all harmonic distance bonds between atoms to the graph as edges
for bond in bonds:
e = graph.add_edge(graph.vs[bond['from_index']], graph.vs[bond['to_index']])
e.update_attributes(bond)
angle_bonds = []
for e in graph.es:
# get the two vertices of the current edge
bond_tuple = [e.source, e.target]
# iterate over the source vertex neighbours
for vertex in e.source_vertex.neighbors():
if vertex.index in bond_tuple:
# skip the vertex if it is part of the current edge
continue
# define the vertex triplet for the current angle bond
triplet = [vertex.index] + bond_tuple
classes = [graph.vs[triplet[0]]['atom_class'],
graph.vs[triplet[1]]['atom_class'],
graph.vs[triplet[2]]['atom_class']]
# find the angle force definition for the current triplet in the forcefield
bond = forcefield.find(f'HarmonicAngleForce/Angle[@class1=\'{classes[0]}\'][@class2=\'{classes[1]}\'][@class3=\'{classes[2]}\']')
if bond is None:
# reverse the atom order and try again
bond = forcefield.find(f'HarmonicAngleForce/Angle[@class1=\'{classes[2]}\'][@class2=\'{classes[1]}\'][@class3=\'{classes[0]}\']')
# add a dictionary containing all bond information to the list
angle = float(bond.get('angle'))
k = float(bond.get('k'))
angle_bonds.append(dict(index1=triplet[0], index2=triplet[1], index3=triplet[2], angle=angle, k=k))
nonbonded = []
for i, a1 in enumerate(tqdm(graph.vs, desc='loading nonbonded atom interactions', ncols=850)):
for a2 in graph.vs[i+1:]:
sigma = (a1['sigma'] + a2['sigma']) / 2
sigma7 = sigma ** 7
sigma13 = sigma ** 13
epsilon = 4 * np.sqrt(a1['epsilon'] * a2['epsilon'])
coulomb = a1['charge'] * a2['charge'] # this should be mutliplied by the Coulomb constant
nb = lambda r: epsilon * ((sigma / r) ** 12 - (sigma / r) ** 6)# + (coulomb / (r ** 2))
xs = np.linspace(2e-1, 1, 10000)
arg = np.argmin(nb(xs))
nonbonded.append(dict(index1=a1.index, index2=a2.index,
sigma7=sigma7, sigma13=sigma13,
epsilon=epsilon, coulomb=coulomb, length=xs[arg]))
def bfs(graph, vid, max_dist=None):
# run a breadth first search up to a maximum distance and save the detection order and parents
order, parents = [], []
for v, dist, parent in graph.bfsiter(vid, advanced=True):
if max_dist is not None and dist > max_dist:
# reached maximum distance, stop the search
break
order.append(v.index)
parents.append(parent.index if parent is not None else -1)
return (order, parents)
def remove_bfs_vertex(vid, order, parents):
# remove the current vertex
if vid in order:
order[np.argwhere(order == vid)[0,0]] = -1
# recursive calls to remove all vertices of which the current one is a parent
for index in np.argwhere(parents == vid):
remove_bfs_vertex(order[index], order, parents)
# maximum distance from the bond to be influenced
max_dist = None
# generate influence lists for each harmonic DISTANCE bond, containing the connected vertices up to max_dist
for e in tqdm(graph.es, desc='computing influence radii for harmonic distance bonds', ncols=850):
# run a breadth-first-search to get all connected vertices on the SOURCE side of the bond
result = bfs(graph, e.source, max_dist=max_dist)
order, parents = np.array(result[0]), np.array(result[1])
# remove all vertices "behind" the bond
remove_bfs_vertex(e.target, order, parents)
e.update_attributes(source_influence=list(order[order != -1]))
# run a breadth-first-search to get all connected vertices on the TARGET side of the bond
result = bfs(graph, e.target, max_dist=max_dist)
order, parents = np.array(result[0]), np.array(result[1])
# remove all vertices "behind" the bond
remove_bfs_vertex(e.source, order, parents)
e.update_attributes(target_influence=list(order[order != -1]))
# generate influence lists for each harmonic ANGLE bond, containing the connected vertices up to max_dist
for angle in tqdm(angle_bonds, desc='computing influence radii for harmonic angle bonds', ncols=850):
result = bfs(graph, angle['index1'], max_dist=max_dist)
order, parents = np.array(result[0]), np.array(result[1])
remove_bfs_vertex(angle['index2'], order, parents)
angle['idx1_influence'] = list(order[order != -1])
result = bfs(graph, angle['index3'], max_dist=max_dist)
order, parents = np.array(result[0]), np.array(result[1])
remove_bfs_vertex(angle['index2'], order, parents)
angle['idx3_influence'] = list(order[order != -1])
length_summ = []
angle_summ = []
nonbonded_summ = []
length_step_size = 0.1
angle_step_size = 0.1
nonbonded_step_size = 0#0.001
steps = 50
def process_dist_bond(e, source_pos, target_pos):
# initialize offsets for each vertex
offsets = np.zeros((len(graph.vs), 3))
# get the vector and distance from the source to the target
vec = target_pos - source_pos
dist = np.linalg.norm(vec)
# compute the current error in the distance
error = dist - e['length']
# iterate over all vertices connected to the source vertex
for v in e['source_influence']:
offsets[v] += vec / dist * error / 2
# iterate over all vertices connected to the target vertex
for v in e['target_influence']:
offsets[v] -= vec / dist * error / 2
return offsets, abs(error)
def process_angle_bond(a):
points = [graph.vs[a['index1']]['pos'], graph.vs[a['index2']]['pos'], graph.vs[a['index3']]['pos']]
# initialize offsets for each vertex
offsets = np.zeros((len(graph.vs), 3))
# get the angle between the three points
s1 = points[0] - points[1]
s2 = points[2] - points[1]
angle = np.arccos(np.dot(s1, s2) / (np.linalg.norm(s1) * np.linalg.norm(s2)))
# get the current angle's error
angle_error = angle - a['angle']
# get the rotation axis by computing the face normal of the three points
rot_axis = np.cross(s1, s2)
# define the rotation matrix
rot = Rotation.from_rotvec((angle_error / 2 * angle_step_size) * (rot_axis / np.linalg.norm(rot_axis)))
origin = points[1]
# iterate over all vertices connected to angle vertex 1
for v in a['idx1_influence']:
pos = graph.vs[v]['pos']
offsets[v] += rot.apply(pos - origin) + origin - pos
# iterate over all vertices connected to angle vertex 2
for v in a['idx3_influence']:
pos = graph.vs[v]['pos']
offsets[v] += rot.inv().apply(pos - origin) + origin - pos
return offsets, abs(angle_error)
def process_nonbonded(nb):
p1 = graph.vs[nb['index1']]['pos']
p2 = graph.vs[nb['index2']]['pos']
vec = p2 - p1
r = np.sqrt(np.sum(vec ** 2))
'''
r3 = r ** 3
r7 = r ** 7
r13 = r ** 13
lj = nb['epsilon'] * (6 * nb['sigma7'] / r7 - 12 * nb['sigma13'] / r13)
coulomb = -2 * nb['coulomb'] / r3
err = lj# + coulomb
if r < 0.05 or r > 2:
err = 0
'''
err = nb['epsilon'] * (r - nb['length'])
return (nb['index1'], vec * err), (nb['index2'], vec * -err)
t = tqdm(total=steps, unit='step', desc='optimizing protein structure', ncols=850)
for step in range(steps):
total_length_error = 0
total_angle_error = 0
total_nonbonded_error = 0
# initialize offsets for each vertex (length and angle)
offsets = np.zeros((len(graph.vs), 3))
# compute the vertex offsets for each harmonic DISTANCE bond
result = Parallel(n_jobs=-1, backend='multiprocessing')(
delayed(process_dist_bond)(e.attributes(), e.source_vertex['pos'], e.target_vertex['pos']) for e in graph.es)
# unpack the results
off, err = zip(*result)
offsets += np.array(off).sum(axis=0) * length_step_size
total_length_error += np.sum(err)
# normalize the total length error
total_length_error /= len(graph.es)
# compute the vertex offsets for each harmonic ANGLE bond
result = Parallel(n_jobs=-1, backend='multiprocessing')(delayed(process_angle_bond)(a) for a in angle_bonds)
# unpack the results
off, err = zip(*result)
offsets += np.array(off).sum(axis=0)
total_angle_error += np.sum(err)
# normalize the total angle error
total_angle_error /= len(angle_bonds)
# compute the vertex offsets for each NON-BONDED atom pair
result = Parallel(n_jobs=-1, backend='multiprocessing')(delayed(process_nonbonded)(nb) for nb in nonbonded)
for curr in result:
for index, off in curr:
offsets[index] += off * nonbonded_step_size
total_nonbonded_error += abs(np.sum(off ** 2))
# normalize the total angle error
total_nonbonded_error /= len(nonbonded)
# apply the offset to the vertex positions
for v, offset in zip(graph.vs, offsets):
v['pos'] += offset
length_summ.append(total_length_error)
angle_summ.append(total_angle_error)
nonbonded_summ.append(total_nonbonded_error)
status = dict(length=f'{total_length_error:.3g}',
angle=f'{total_angle_error:.3g}',
nonbonded=f'{total_nonbonded_error:.3g}')
t.set_postfix(status)
t.update()
fig, axes = plt.subplots(3, 1, figsize=(9.5, 8))
axes[0].set_title('length')
axes[0].plot(length_summ)
axes[1].set_title('angle')
axes[1].plot(angle_summ)
axes[2].set_title('nonbonded')
axes[2].plot(nonbonded_summ)
plt.tight_layout()
plt.show()
def draw_graph(g, error_scale=3, draw_names='backbone'):
fig = plt.figure(figsize=(9, 6))
ax = fig.add_subplot(111, projection='3d')
# create the atom name list if backbone visualization was chosen
if draw_names == 'backbone':
draw_names = ['N', 'CA', 'C']
# draw the edges
for edge in g.es:
name1 = edge.source_vertex['name']
name2 = edge.target_vertex['name']
# filter out atoms that are not in the draw_names list
if draw_names is not None and (name1 not in draw_names or name2 not in draw_names):
continue
pos1 = edge.source_vertex['pos']
pos2 = edge.target_vertex['pos']
# get the length error of the bond and get the corresponding color value
error = np.minimum(1, np.abs(np.linalg.norm(pos2 - pos1) - e['length']) * error_scale)
ax.plot([pos1[0], pos2[0]], [pos1[1], pos2[1]], [pos1[2], pos2[2]], c=(error, (1 - error), 0))
# create a dict with residue names as keys and the corresponding atoms as values
vertex_dict = {}
for vertex in g.vs:
res = f'{vertex["residue_index"] + 1} - {vertex["residue"]}'
if res in vertex_dict:
vertex_dict[res].append(vertex)
else:
vertex_dict[res] = [vertex]
# draw vertices with names defined by draw_names
for res, vertices in vertex_dict.items():
pos = np.array([vertex['pos'] for vertex in vertices if draw_names is None or vertex['name'] in draw_names])
ax.plot(pos[:,0], pos[:,1], pos[:,2], 'o', label=res)
plt.legend()
plt.tight_layout()
plt.show()
draw_graph(graph, draw_names=None)
with open('proteins/malek.pickle', 'wb') as file:
pickle.dump([graph, angle_bonds, nonbonded], file)
```
| github_jupyter |
```
%matplotlib inline
import os
os.chdir("..")
os.getcwd()
from pymongo import MongoClient
from sklearn.decomposition import PCA
from scipy.spatial.distance import cosine
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
from matplotlib import rcParams
import matplotlib.patches as patches
from PIL import Image
from PIL import ImageOps
import torch.utils.data as data
from keras.preprocessing import image
from tqdm import tqdm_notebook
from io import BytesIO
from torchvision import transforms
from torch.autograd import Variable
from sklearn.cluster import KMeans
from sklearn.externals import joblib
import os
from utils.image_utils import scale_image
import os
import glob
import requests
import numpy as np
import pandas as pd
import torch
from utils.composite_model_utils import load_extractor_model
from ai.feature_extraction import vectorize, get_deep_color_top_n
import logging
DATA_PATH = '/run/media/twoaday/data-storag/data-sets/where2buyit/photos'
MODEL_PATH = '/home/twoaday/ai/models/fashion-vectorizer-converted.pth.tar'
NUM_CLUSTERS = 50
KNN_DF_MODEL_PATH = '/home/twoaday/ai/models/knn-deep-features-fashion.m'
PCA_DF_MODEL_PATH = '/home/twoaday/ai/models/pca-2-deep-features-fashion.m'
PCA_CF_MODEL_PATH = '/home/twoaday/ai/models/pca-2-color-features-fashion.m'
mongo_client = MongoClient()
products_collection = mongo_client["deep-fashion"]["products"]
extractor = load_extractor_model(MODEL_PATH)
result = [y for x in tqdm_notebook(os.walk(DATA_PATH), 'Walking through folders')
for y in glob.glob(os.path.join(x[0], '*.jpg'))]
df_data_set = []
for file in tqdm_notebook(result, desc='Parsing files'):
s = file.split('/')
name, product, category = s[-1].replace('.jpg', ''), s[-2], s[-3]
df_data_set.append([name, product, category, file])
df_data_set = pd.DataFrame(df_data_set)
df_data_set.columns = ['name', 'product', 'category', 'file']
df_query_set = df_data_set.loc[df_data_set.name == 'query']
df_data_set = df_data_set.loc[df_data_set.name != 'query']
df_data_set.count()
rows = []
for _, row in tqdm_notebook(df_data_set.iterrows(), desc='Vectorizing', total=len(df_data_set)):
try:
deep_feat, color_feat = vectorize(extractor, row['file'])
rows.append([row['name'],
row['product'],
row['category'],
row['file'],
deep_feat,
color_feat])
except Exception as exp:
logging.error('Can not vectorize {0}'.format(row['file']), exp)
knn = KMeans(n_clusters=NUM_CLUSTERS, n_jobs=8).fit([r[4] for r in rows])
joblib.dump(knn, KNN_DF_MODEL_PATH)
clusters = knn.predict([r[4] for r in rows])
rows = [r + [c] for r, c in tqdm_notebook(zip(rows, clusters), desc='Appending clusters')]
df_pca = PCA(n_components=2).fit([r[4] for r in rows])
joblib.dump(df_pca, PCA_DF_MODEL_PATH)
cf_pca = PCA(n_components=2).fit([r[5] for r in rows])
joblib.dump(df_pca, PCA_CF_MODEL_PATH)
df_coords = df_pca.transform([r[4] for r in rows])
color_coords = cf_pca.transform([r[5] for r in rows])
for r, dfc, cfc in tqdm_notebook(zip(rows, df_coords, color_coords),
desc='Generating collection',
total=len(rows)):
try:
if len(r) == 7:
obj = {'imageName' : r[0],
'product': r[1],
'category': r[2],
'filePath': r[3],
'cluster': int(r[6]),
'deepFeatures': [float(dfc[0]), float(dfc[1])],
'colorFeatures': [float(cfc[0]), float(cfc[1])]
}
products_collection.insert_one(obj)
except Exception as exp:
logging.error(exp)
def search(row):
deep_feat, color_feat = vectorize(extractor, row['file'])
df_coord = df_pca.transform([deep_feat])[0]
color_coord = cf_pca.transform([color_feat])[0]
cn = knn.predict([deep_feat])[0]
query = {
'cluster':int(cn),
'deepFeatures' :{
'$near': {
'$geometry' : {
'type' : 'Point' ,
'coordinates' : [float(df_coord[0]), float(df_coord[1])]
}
}
}
}
return products_collection.find(query).limit(3)
for _, r in df_query_set.sample(1).iterrows():
search_result = [o['filePath'] for o in search(r)]
# figure size in inches optional
rcParams['figure.figsize'] = 11, 8
# display images
fig, ax = plt.subplots(1,4)
ax[0].imshow(mpimg.imread(r['file']))
indx = 1
for img_path in search_result:
ax[indx].imshow(mpimg.imread(img_path))
indx += 1
for _, anchor in df_query_set.sample(1).iterrows():
deep_feat, color_feat = vectorize(extractor, anchor['file'])
rs = [r for r in rows if r[2] == anchor['category']]
similar = get_deep_color_top_n(deep_feat,
color_feat,
[r[4] for r in rs],
[r[5] for r in rs],
[r[3] for r in rs],
5)
# figure size in inches optional
rcParams['figure.figsize'] = 15, 11
# display images
fig, ax = plt.subplots(1, 6)
ax[0].imshow(mpimg.imread(anchor['file']))
indx = 1
for sm_img in similar:
ax[indx].imshow(mpimg.imread(sm_img[0]))
indx += 1
print(anchor['category'])
print('distances:')
for sm_img in similar:
print(sm_img[1])
```
| github_jupyter |
```
# Copyright 2019 The TensorFlow Hub Authors. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
```
# CropNet: Cassava Disease Detection
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/hub/tutorials/cropnet_cassava"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/hub/blob/master/examples/colab/cropnet_cassava.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/hub/blob/master/examples/colab/cropnet_cassava.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/hub/examples/colab/cropnet_cassava.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
<td>
<a href="https://tfhub.dev/google/cropnet/classifier/cassava_disease_V1/2"><img src="https://www.tensorflow.org/images/hub_logo_32px.png" />See TF Hub model</a>
</td>
</table>
This notebook shows how to use the CropNet [cassava disease classifier](https://tfhub.dev/google/cropnet/classifier/cassava_disease_V1/2) model from **TensorFlow Hub**. The model classifies images of cassava leaves into one of 6 classes: *bacterial blight, brown streak disease, green mite, mosaic disease, healthy, or unknown*.
This colab demonstrates how to:
* Load the https://tfhub.dev/google/cropnet/classifier/cassava_disease_V1/2 model from **TensorFlow Hub**
* Load the [cassava](https://www.tensorflow.org/datasets/catalog/cassava) dataset from **TensorFlow Datasets (TFDS)**
* Classify images of cassava leaves into 4 distinct cassava disease categories or as healthy or unknown.
* Evaluate the *accuracy* of the classifier and look at how *robust* the model is when applied to out of domain images.
## Imports and setup
```
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
import tensorflow_datasets as tfds
import tensorflow_hub as hub
#@title Helper function for displaying examples
def plot(examples, predictions=None):
# Get the images, labels, and optionally predictions
images = examples['image']
labels = examples['label']
batch_size = len(images)
if predictions is None:
predictions = batch_size * [None]
# Configure the layout of the grid
x = np.ceil(np.sqrt(batch_size))
y = np.ceil(batch_size / x)
fig = plt.figure(figsize=(x * 6, y * 7))
for i, (image, label, prediction) in enumerate(zip(images, labels, predictions)):
# Render the image
ax = fig.add_subplot(x, y, i+1)
ax.imshow(image, aspect='auto')
ax.grid(False)
ax.set_xticks([])
ax.set_yticks([])
# Display the label and optionally prediction
x_label = 'Label: ' + name_map[class_names[label]]
if prediction is not None:
x_label = 'Prediction: ' + name_map[class_names[prediction]] + '\n' + x_label
ax.xaxis.label.set_color('green' if label == prediction else 'red')
ax.set_xlabel(x_label)
plt.show()
```
## Dataset
Let's load the *cassava* dataset from TFDS
```
dataset, info = tfds.load('cassava', with_info=True)
```
Let's take a look at the dataset info to learn more about it, like the description and citation and information about how many examples are available
```
info
```
The *cassava* dataset has images of cassava leaves with 4 distinct diseases as well as healthy cassava leaves. The model can predict all of these classes as well as sixth class for "unknown" when the model is not confident in it's prediction.
```
# Extend the cassava dataset classes with 'unknown'
class_names = info.features['label'].names + ['unknown']
# Map the class names to human readable names
name_map = dict(
cmd='Mosaic Disease',
cbb='Bacterial Blight',
cgm='Green Mite',
cbsd='Brown Streak Disease',
healthy='Healthy',
unknown='Unknown')
print(len(class_names), 'classes:')
print(class_names)
print([name_map[name] for name in class_names])
```
Before we can feed the data to the model, we need to do a bit of preprocessing. The model expects 224 x 224 images with RGB channel values in [0, 1]. Let's normalize and resize the images.
```
def preprocess_fn(data):
image = data['image']
# Normalize [0, 255] to [0, 1]
image = tf.cast(image, tf.float32)
image = image / 255.
# Resize the images to 224 x 224
image = tf.image.resize(image, (224, 224))
data['image'] = image
return data
```
Let's take a look at a few examples from the dataset
```
batch = dataset['validation'].map(preprocess_fn).batch(25).as_numpy_iterator()
examples = next(batch)
plot(examples)
```
## Model
Let's load the classifier from TF-Hub and get some predictions and see the predictions of the model is on a few examples
```
classifier = hub.KerasLayer('https://tfhub.dev/google/cropnet/classifier/cassava_disease_V1/2')
probabilities = classifier(examples['image'])
predictions = tf.argmax(probabilities, axis=-1)
plot(examples, predictions)
```
## Evaluation & robustness
Let's measure the *accuracy* of our classifier on a split of the dataset. We can also look at the *robustness* of the model by evaluating its performance on a non-cassava dataset. For image of other plant datasets like iNaturalist or beans, the model should almost always return *unknown*.
```
#@title Parameters {run: "auto"}
DATASET = 'cassava' #@param {type:"string"} ['cassava', 'beans', 'i_naturalist2017']
DATASET_SPLIT = 'test' #@param {type:"string"} ['train', 'test', 'validation']
BATCH_SIZE = 32 #@param {type:"integer"}
MAX_EXAMPLES = 1000 #@param {type:"integer"}
def label_to_unknown_fn(data):
data['label'] = 5 # Override label to unknown.
return data
# Preprocess the examples and map the image label to unknown for non-cassava datasets.
ds = tfds.load(DATASET, split=DATASET_SPLIT).map(preprocess_fn).take(MAX_EXAMPLES)
dataset_description = DATASET
if DATASET != 'cassava':
ds = ds.map(label_to_unknown_fn)
dataset_description += ' (labels mapped to unknown)'
ds = ds.batch(BATCH_SIZE)
# Calculate the accuracy of the model
metric = tf.keras.metrics.Accuracy()
for examples in ds:
probabilities = classifier(examples['image'])
predictions = tf.math.argmax(probabilities, axis=-1)
labels = examples['label']
metric.update_state(labels, predictions)
print('Accuracy on %s: %.2f' % (dataset_description, metric.result().numpy()))
```
## Learn more
* Learn more about the model on TensorFlow Hub: https://tfhub.dev/google/cropnet/classifier/cassava_disease_V1/2
* Learn how to build a custom image classifier running on a mobile phone with [ML Kit](https://developers.google.com/ml-kit/custom-models#tfhub) with the [TensorFlow Lite version of this model](https://tfhub.dev/google/lite-model/cropnet/classifier/cassava_disease_V1/1).
| github_jupyter |
# Simple Vectorfield Model
*mZargham*
Demonstration of hyperbolic coordinates and vectorfield interpretation of constant product market maker activity
```
from cadCAD.configuration import Experiment
from cadCAD.configuration.utils import config_sim
from cadCAD.configuration import Configuration
from cadCAD.engine import ExecutionMode, ExecutionContext, Executor
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
MONTE_CARLO_RUNS = 1
SIMULATION_TIMESTEPS = 100
x0 = 20
y0 = 100
#cartersian to hyperbolic
def xy2uv(x,y):
u = np.log(np.sqrt(x/y))
v = np.sqrt(x*y)
return (u,v)
#hyperbolic to cartesian
def uv2xy(u,v):
x = v*np.exp(u)
y = v*np.exp(-u)
return (x,y)
u0, v0 = xy2uv(x0,y0)
genesis_states = {
'cartesian': (x0,y0),
'hyperbolic': (u0,v0)
}
```
This snippet is intended to demonstrate the impact of latent market forces on an AMM using a simple vectorfield representation.
For simplicity we will construct a simple vector field around an equilibrium characterized by $P(x,y) = \frac{x}{y} = P^*$.
While our market is characterized by Quadrant I of the Cartesian plane, we are working with the idealized constant product market maker so there is an invariant function $V(x,y) = x \cdot y = V^*$
The resulting dynamical system can be expressed succinctly in hyperbolic coordinates
$u = \ln \sqrt{\frac{x}{y}} = \ln \sqrt{P(x,y)}$
$v = \sqrt{x\cdot y} = \sqrt{V(x,y)}$
By excluding the add/remove liquidity mechanism we can characterize the vectorfield using only the $u$ dimension as fixed at $v = v_0 = \sqrt{V^*}$
$\Delta u = -k(u-u^*)$
$\Delta {v} = 0$
where $u^* = \ln \sqrt{P^*}$ and $k$ is gain coeficient.
we can construct a vectorfield representation of this in cartesian coordinates as follows
```
# vectorfield
k=.5
pstar = 1
ustar = np.log(np.sqrt(pstar))
x,y = np.meshgrid(np.linspace(1,2001,21),np.linspace(1,2001,21))
u, v = xy2uv(x,y)
du = -k*(u-ustar)
dv = np.zeros_like(v)
xplus, yplus = uv2xy(u+du,v+dv)
dx = (xplus-x)
dy = (yplus-y)
plt.rcParams.update({'font.size': 22})
plt.xlabel('quatity of token x')
plt.ylabel('quantity of token y')
plt.plot([0,2000], [0,2000], np.linspace(1,2001, 201), 2000/np.linspace(1,2001, 201))
plt.legend(["x=y", "x*y=2000"])
plt.quiver(x,y,dx,dy)
plt.title('AMM vectorfield for x~y')
f = plt.gcf()
f.set_figwidth(12)
f.set_figheight(12)
def applyField(params,
substep,
state_history,
prev_state):
pstar = params['pstar']
ustar = np.log(np.sqrt(pstar))
u, __ = prev_state['hyperbolic']
error = u-ustar
du = -params['field']*error + params['noise']*np.random.randn()
return {'du':du}
def update_hyperbolic(params,
substep,
state_history,
prev_state,
policy_input):
u, v = prev_state['hyperbolic']
u = u+policy_input['du']
point = (u, v)
return ('hyperbolic', point)
def update_cartesian(params,
substep,
state_history,
prev_state,
policy_input):
u, v = prev_state['hyperbolic']
u = u+policy_input['du']
x,y = uv2xy(u,v)
point = (x, y)
return ('cartesian', point)
sys_params = {
'pstar': [1],
'field': [.25],
'noise' : [.05]
}
partial_state_update_blocks = [
{
'policies': {applyField
},
'variables': {
'cartesian': update_cartesian,
'hyperbolic': update_hyperbolic
}
}
]
sim_config = {
'N': MONTE_CARLO_RUNS,
'T': range(SIMULATION_TIMESTEPS),
'M': sys_params
}
sim_params = config_sim(sim_config)
print(sim_params)
exp = Experiment()
exp.append_configs(
sim_configs=sim_params,
initial_state=genesis_states,
partial_state_update_blocks=partial_state_update_blocks
)
exec_mode = ExecutionMode()
local_mode_ctx = ExecutionContext(context=exec_mode.local_mode)
simulation = Executor(exec_context=local_mode_ctx,
configs=exp.configs)
raw_system_events, tensor_field, sessions = simulation.execute()
df = pd.DataFrame(raw_system_events)
df.head()
df['x'] = df.cartesian.apply(lambda z: z[0])
df['y'] = df.cartesian.apply(lambda z: z[1])
df['dx']= df.x.diff()
df['dy']= df.y.diff()
plt.plot(np.linspace(df.x.min(),df.x.max(), 101), x0*y0/np.linspace(df.x.min(),df.x.max(),101))
plt.plot(df.x,df.y, '.')
plt.grid('square')
plt.title("CPMM Cartesian Coordinates")
plt.xlabel('x')
plt.ylabel('y')
df['u'] = df.hyperbolic.apply(lambda z: z[0])
df['v'] = df.hyperbolic.apply(lambda z: z[1])
df['du']= df.u.diff()
df['dv']= df.v.diff()
plt.plot(np.linspace(df.u.min(),df.u.max(), 101), v0*np.ones_like(np.linspace(df.u.min(),df.u.max(), 101)))
plt.plot(df.u,df.v, '.')
plt.grid('square')
plt.title("CPMM Hyperbolic Coordinates")
plt.xlabel('u')
plt.ylabel('v')
plt.hist(df.y/df.x, bins=np.linspace(0,5,31), alpha=.5)
plt.hist(-df.y.diff()/df.x.diff(), bins=np.linspace(0,5,31), alpha=.5)
plt.vlines(1,0,30, 'g')
f = plt.gcf()
f.set_figwidth(12)
f.set_figheight(12)
plt.legend(["p-star", "spot price", "realized price"])
plt.plot(df.index,df.x*df.y)
plt.title("Constant Product")
plt.ylabel("xy")
plt.xlabel("index")
plt.plot(df.index,df.x, df.index, df.y)
plt.title("Cartesian Trajectory")
plt.ylabel("state")
plt.xlabel("index")
plt.legend(["x", "y"])
plt.plot(df.index,df.dx, df.index, df.dy)
plt.title("Cartesian State Changes")
plt.ylabel("state changes")
plt.xlabel("index")
plt.legend(["dx", "dy"])
plt.plot(df.index,df.u)
ax1=plt.gca()
plt.xlabel("index")
ax2 = ax1.twinx()
ax2.plot(df.index, df.v, 'orange')
plt.title("Hyperbolic Trajectory")
ax1.set_ylabel('u', color='b')
ax2.set_ylabel('v', color='orange')
plt.plot(df.index,df.du, df.index, df.dv)
plt.title("Hyperbolic State change")
plt.ylabel("state changes")
plt.xlabel("index")
plt.legend(["du", "dv"])
```
| github_jupyter |
# ARCLYTICS User Analytics
This Jupyter Notebook is for doing some quick and dirty tests for running data
analytics operations on the MongoDB and Redis data persistence. The purpose of
which is to then put into a pipeline and then create endpoints for each type
of query and analysis that can be done.
```
# Plotly imports
import chart_studio
import chart_studio.plotly as py
import plotly.graph_objects as go
import plotly.io as pio
import plotly.express as px
from plotly.subplots import make_subplots
chart_studio.tools.set_credentials_file(
username='codeninja55',
api_key='mLp691cLJDdKaNgJykR4'
)
chart_studio.tools.set_config_file(
world_readable=True,
sharing='public'
)
# imports
import datetime
from os import environ as env
from pymongo import MongoClient
import pandas as pd
from redis import Redis
import json
conn = MongoClient(env.get('MONGO_URI'))
db_name = 'arc_dev'
collection = 'users'
db = conn[db_name]
db
```
## Search Testing
```
pipeline = [
{
"$lookup": {
"from": "users",
"localField": "user",
"foreignField": "_id",
"as": "user"
}
},
{ "$unwind": "$user" },
{ "$skip": 100 },
{ "$sort": { "category": 1 } },
{ "$limit": 10 },
{
"$project": {
"_id": 0,
"category": 1,
"rating": 1,
"comment": 1,
"created_date": 1,
"user.email": 1,
}
}
]
res = db['feedback'].aggregate(pipeline)
data = list(res)
data[0]['user']['email']
pipeline = [
{
"$lookup": {
"from": "users",
"localField": "user",
"foreignField": "_id",
"as": "user"
}
},
{ "$unwind": "$user" },
{
"$match": {
"user.email": "andrew@neuraldev.io"
}
},
{ "$sort": { "category": 1 } },
{ "$limit": 2 },
{
"$project": {
"_id": 0,
"category": 1,
"rating": 1,
"comment": 1,
"created_date": 1,
"user.email": 1,
}
}
]
res = db['feedback'].aggregate(pipeline)
data = list(res)
data
pipeline = [
{
"$match": {
"$text": {
"$search": "Appearance",
"$language": "en",
"$caseSensitive": False
}
}
},
{
"$lookup": {
"from": "users",
"localField": "user",
"foreignField": "_id",
"as": "user"
}
},
{ "$unwind": "$user" },
{ "$sort": { "category": 1 } },
{ "$limit": 2 },
{
"$project": {
"_id": 0,
"category": 1,
"rating": 1,
"comment": 1,
"created_date": 1,
"user.email": 1,
}
}
]
res = db['feedback'].aggregate(pipeline)
data = list(res)
data
cursor = db[collection].find(
{
'profile': {'$exists': True}
},
projection={'password': 0, '_id': False}
)
df = pd.DataFrame(list(cursor))
df.head()
```
## User Profile Data
```
pipeline = [
{'$unwind': '$profile'},
{'$project': {'profile': 1, '_id': False}},
]
res = db[collection].aggregate(pipeline)
list(res)
pipeline = [
{'$unwind': '$profile'},
{'$project': {
'aim': '$profile.aim',
'highest_education': '$profile.highest_education',
'sci_tech_exp': '$profile.sci_tech_exp',
'phase_transform_exp': '$profile.phase_transform_exp',
'_id': 0
}
},
]
res = db[collection].aggregate(pipeline)
profile_df = pd.DataFrame(list(res))
profile_df['aim'].value_counts()
list(profile_df['aim'].unique())
list(profile_df['aim'].value_counts())
# layout = go.Layout(
# title='User Profile Aim',
# xaxis=dict(title='User Aims'),
# yaxis=dict(title='Count')
# )
# fig = go.Figure(layout=layout)
fig = make_subplots(
rows=2,
cols=2,
subplot_titles=[
'Aim',
'Highest Education',
'Science Tech. Experience',
'Phase Transform Experience'
]
)
trace_aim = go.Bar(x=list(profile_df['aim'].unique()), y=list(profile_df['aim'].value_counts()))
trace_edu = go.Bar(x=list(profile_df['highest_education'].unique()), y=list(profile_df['highest_education'].value_counts()))
trace_sci = go.Bar(x=list(profile_df['sci_tech_exp'].unique()), y=list(profile_df['sci_tech_exp'].value_counts()))
trace_pha = go.Bar(x=list(profile_df['phase_transform_exp'].unique()), y=list(profile_df['phase_transform_exp'].value_counts()))
fig.add_trace(trace_aim, row=1, col=1)
fig.add_trace(trace_edu, row=1, col=2)
fig.add_trace(trace_sci, row=2, col=1)
fig.add_trace(trace_pha, row=2, col=2)
fig.update_layout(
# height=800,
# width=1200,
showlegend=False,
title_text="User Profile Answers"
)
py.iplot(fig, filename='user_profile_bar')
# pio.write_image(fig, file='user_profile_aim.png')
```
## Count
```
# Total user count
db[collection].estimated_document_count()
# Total saved simulations count
db['saved_simulations'].estimated_document_count()
# Total feedback count
db['feedback'].estimated_document_count()
# Total shares
db['shared_simulations'].estimated_document_count()
# Total simulations
pipeline = [
{
'$group': {
'_id': None,
'total': {
'$sum': '$simulations_count'
}
}
}
]
cursor = db[collection].aggregate(pipeline)
# count_df = pd.DataFrame(list(cursor))
# count_df
list(cursor)[0]['total']
# Total saved user alloys
pipeline = [
{
'$group': {
'_id': None,
'total': {
'$sum': {'$size': '$saved_alloys'}
}
}
}
]
cursor = db[collection].aggregate(pipeline)
list(cursor)[0]['total']
# Total ratings average
pipeline = [
{'$unwind': '$ratings'},
{
'$group': {
'_id': None,
'count': { '$sum': 1 },
'average': {'$avg': {'$sum': '$ratings.rating'}}
}
}
]
cursor = db[collection].aggregate(pipeline)
list(cursor)[0]
```
## Live Login Data
```
cursor = db[collection].find(
{
'last_login': {'$exists': 1}
},
projection={'password': 0, '_id': False}
)
df = pd.DataFrame(list(cursor))
df.head()
pipeline = [
{'$unwind': '$login_data'},
{'$project': {'_id': 0, 'login_data': 1, 'email': 1}},
{'$sort': {'login_data.created_datetime': 1}}
]
res = db[collection].aggregate(pipeline)
# login_df = pd.DataFrame(list(res))
list(res)
pipeline = [
{'$unwind': '$login_data'},
{'$project': {
'_id': 0,
'created_datetime': '$login_data.created_datetime',
}
},
]
res = db[collection].aggregate(pipeline)
list(res)
# Using graph_objects
import plotly.graph_objects as go
import pandas as pd
df = pd.read_csv('https://raw.githubusercontent.com/plotly/datasets/master/finance-charts-apple.csv')
df['Date'].head(n=10)
df['AAPL.High'].head(n=10)
fig = go.Figure([go.Scatter(x=df['Date'], y=df['AAPL.High'])])
fig.show()
import plotly.graph_objects as go
import datetime
x = [datetime.datetime(year=2013, month=10, day=4),
datetime.datetime(year=2013, month=11, day=5),
datetime.datetime(year=2013, month=12, day=6)]
fig = go.Figure(data=[go.Scatter(x=x, y=[1, 3, 6])])
# Use datetime objects to set xaxis range
fig.update_layout(xaxis_range=[datetime.datetime(2013, 10, 17),
datetime.datetime(2013, 11, 20)])
fig.show()
pipeline = [
{'$unwind': '$login_data'},
{'$project': {
'_id': 0,
'timestamp': '$login_data.created_datetime',
'user': '$email',
}
},
]
res = db[collection].aggregate(pipeline)
# dt_idx = pd.to_datetime()
df = pd.DataFrame(list(res))
df['timestamp'] = pd.to_datetime(df['timestamp'])
# df.set_index('timestamp', inplace=True)
df = df.groupby(pd.Grouper(key='timestamp', freq='1min')).count().dropna()
# df = df.groupby(pd.Grouper(key='timestamp', freq='60s'))
# res = (pd.DataFrame(df.index[1:]) - pd.DataFrame(df.index[:-1]))
# df = df.to_frame().reset_index()
# df.resample('T').count()
# res['timestamp'].value_counts()
df
fig = go.Figure()
trace = go.Scatter(x=df.index, y=df['user'])
fig.add_trace(trace)
fig.update_layout(
showlegend=False,
title_text="Logged in Users",
xaxis_range=[
datetime.datetime(2019, 10, 4),
datetime.datetime(2019, 10, 5)
],
xaxis_rangeslider_visible=True
)
py.iplot(fig, filename='user_login_timestamps')
```
## Logged In User Map
```
redis_uri = env.get('REDIS_URI')
client = Redis(redis_uri)
client
keys = client.keys(pattern=u'session*')
keys
for byte_key in keys:
key = byte_key.decode('utf-8')
print()
sess_store = json.loads(client.get(key))
print(sess_store)
gapminder = px.data.gapminder().query("year == 2007")
t = gapminder['iso_alpha'].value_counts().to_dict()
t['AUS']
df = pd.read_csv('https://raw.githubusercontent.com/plotly/datasets/master/2014_world_gdp_with_codes.csv')
fig = go.Figure(data=go.Choropleth(
locations = df['CODE'],
z = df['GDP (BILLIONS)'],
text = df['COUNTRY'],
colorscale = 'Blues',
autocolorscale=False,
reversescale=True,
marker_line_color='darkgray',
marker_line_width=0.5,
colorbar_tickprefix = '$',
colorbar_title = 'GDP<br>Billions US$',
))
# fig.update_layout(
# title_text='2014 Global GDP',
# geo=dict(
# showframe=False,
# showcoastlines=False,
# projection_type='equirectangular'
# ),
# annotations = [dict(
# x=0.55,
# y=0.1,
# xref='paper',
# yref='paper',
# text='Source: <a href="https://www.cia.gov/library/publications/the-world-factbook/fields/2195.html">\
# CIA World Factbook</a>',
# showarrow = False
# )]
# )
# fig.show()
df['CODE'].value_counts()
pipeline = [
{'$unwind': '$login_data'},
{'$project': {
'_id': 0,
'created_datetime': '$login_data.created_datetime',
'ip_address': '$login_data.ip_address',
'state': '$login_data.state',
'country': '$login_data.country',
'iso_code': '$login_data.country_iso_code',
'continent': '$login_data.continent',
'accuracy_radius': '$login_data.accuracy_radius',
'timezone': '$login_data.timezone',
'latitude': {'$arrayElemAt': [ '$login_data.geo_point.coordinates', 0 ]},
'longitude': {'$arrayElemAt': [ '$login_data.geo_point.coordinates', 1 ]},
}
},
]
res = db[collection].aggregate(pipeline)
df = pd.DataFrame(list(res))
df.dropna(subset=['country', 'continent'], axis=0, inplace=True)
# cnt = df['country'].value_counts().to_dict()
# df['count'] = df[ df['country'] == cnt[] ]
df = df.groupby(
['latitude', 'longitude', 'country', 'continent']
).size().to_frame('count').reset_index()
# print(df.count())
df.head(n=10)
df['country'].tolist()
s_df = df[df['country'] == 'Singapore']
s_df['ip_address'].value_counts()
# df_group.set_index('continent', inplace=True)
df_group.reset_index()
df_group.head()
# fig = go.Figure()
# trace = go.Scatter(x=df.index, y=df['user'])
# fig.add_trace(trace)
# fig.update_layout(
# showlegend=False,
# title_text="Logged in Users",
# xaxis_range=[
# datetime.datetime(2019, 10, 4),
# datetime.datetime(2019, 10, 5)
# ],
# xaxis_rangeslider_visible=True
# )
# py.iplot(fig, filename='user_login_timestamps')
fig = px.scatter_geo(
df,
locations='iso_code',
color='continent',
hover_name='country',
size='count',
projection="natural earth"
)
py.iplot(fig, filename='user_login_map')
# mapbox_access_token = open(
# "/home/codeninja/Arclytics/arclytics_sim/.mapbox_token"
# ).read()
mapbox_access_token = 'pk.eyJ1IjoiY29kZW5pbmphNTUiLCJhIjoiY2sxZG5kb2JvMDV3dzNsbXV6dmhwd2xkaCJ9.3yH0KfKaMVn0MHNqgq7g5g'
fig = go.Figure(go.Densitymapbox(
lat=df['latitude'],
lon=df['longitude'],
z=df['count'],
radius=10,
# mode='markers',
# marker=go.scattermapbox.Marker(
# size=8,
# color='rgb(254, 67, 54)',
# opacity=0.8
# ),
text=df['count'],
))
fig.update_layout(
hovermode='closest',
mapbox=go.layout.Mapbox(
accesstoken=mapbox_access_token,
# bearing=0,
center=go.layout.mapbox.Center(
lat=0,
lon=180
),
# pitch=0,
zoom=1
)
)
fig.update_layout(margin={"r":0,"t":0,"l":0,"b":0})
# fig.show()
py.iplot(fig, filename='user_login_map')
```
| github_jupyter |
```
# Built-in libraries
from datetime import datetime, timedelta
# NumPy, SciPy and Pandas
import pandas as pd
import numpy as np
def hourly_dataset(name):
"""
Constants for time period with maximum number of buildings measured simultaneously in the BDG dataset.
For more details, go to old_files/RawFeatures_BDG.ipynb
"""
BDG_STARTDATE = datetime.strptime('01/01/15 00:00', '%d/%m/%y %H:%M')
BDG_ENDDATE = datetime.strptime('30/11/15 23:00','%d/%m/%y %H:%M')
# Building Data Genome dataset
if name == 'BDG':
df = pd.read_csv('../data/raw/temp_open_utc_complete.csv', parse_dates=True,
infer_datetime_format=True, index_col=0)
# truncate the dataframe based on a pre-calculated time period, if needed
startDate = BDG_STARTDATE
endDate = BDG_ENDDATE
df = df[(df.index >= startDate) & (df.index <= endDate)]
# Washington D.C. dataset
elif name == 'DGS':
df = pd.read_csv('../data/raw/DGS_322_Buildings-15m-By_Building-DST-gap-filled-3-2-18-508pm.csv',
parse_dates=[['Building ID', 'Unnamed: 1']], infer_datetime_format=True)
# get rid of temperature column
del df['Unnamed: 2']
# update column names to match the row of building names
new_column_names = df.iloc[0,:]
df.columns = new_column_names
# get rid of rows with metadata and update index
df = df.drop([0,1,2], axis=0)
df = df.rename(columns = {'Building nan':'timestamp'})
df.index = df['timestamp'].astype('datetime64[ns]')
del df['timestamp']
df = df.astype(float)
# since the dataset is made from 15min interval readings, resample to 1 hr
df = df.resample('1H').sum()
else:
print("Please choose a valid dataset")
exit()
# save the file to csv before exit
df.to_csv('../data/processed/{}_dataset.csv'.format(name))
return df
from collections import Counter
def resampleDGS():
df = pd.read_csv("../data/processed/DGS_dataset.csv", parse_dates=True, infer_datetime_format=True, index_col=0)
og_index = df.index.values
df = df.T
df_meta = pd.read_csv('../data/raw/dgs_metadata.csv')
df_aux = pd.read_csv("../data/raw/DGS_322_Buildings-15m-By_Building-DST-gap-filled-3-2-18-508pm.csv")
# get labels for all buildings
df_aux = df_aux.T
df_aux_og = df_aux.copy()
df_label = df_aux[df_aux.iloc[:, 0].isin(df.index.values)] # get id based on names
df_label = df_meta[df_meta['id'].isin(df_label.index.values)] # get label based on id
# print(c.value_counts())
cnt = Counter(df_label['espm_type_name'
])
for i in df_label['espm_type_name']:
print(cnt[i])
df_label = df_label[(df_label['espm_type_name'] == 'K-12 School') |
(df_label['espm_type_name'] == 'Other - Recreation') |
(df_label['espm_type_name'] == 'Fire Station') |
(df_label['espm_type_name'] == 'Office') |
(df_label['espm_type_name'] == 'Library') |
(df_label['espm_type_name'] == 'Other - Public Services') |
(df_label['espm_type_name'] == 'Police Station')]
# print(df_label['espm_type_name'].value_counts())
df_aux_og = df_aux_og.drop(df_aux_og.index[0:3])
df_aux_og.index = list(map(int, df_aux_og.index.values))
df_bdg_name = df_aux_og[df_aux_og.index.isin(df_label['id'])]
df = df[df.index.isin(df_bdg_name.iloc[:, 0])]
df = df.T
df.index = og_index
# df.to_csv('../data/processed/DGS_dataset.csv')
# load building gnome dataset (BDG)
df_BDG = hourly_dataset('BDG')
# load dc building dataset (DC)
df_DGS = hourly_dataset('DGS')
resampleDGS()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_06_5_yolo.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# T81-558: Applications of Deep Neural Networks
**Module 6: Convolutional Neural Networks (CNN) for Computer Vision**
* Instructor: [Jeff Heaton](https://sites.wustl.edu/jeffheaton/), McKelvey School of Engineering, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
* For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
# Module 6 Material
* Part 6.1: Image Processing in Python [[Video]](https://www.youtube.com/watch?v=4Bh3gqHkIgc&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_06_1_python_images.ipynb)
* Part 6.2: Keras Neural Networks for Digits and Fashion MNIST [[Video]](https://www.youtube.com/watch?v=-SA8BmGvWYE&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_06_2_cnn.ipynb)
* Part 6.3: Implementing a ResNet in Keras [[Video]](https://www.youtube.com/watch?v=qMFKsMeE6fM&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_06_3_resnet.ipynb)
* Part 6.4: Using Your Own Images with Keras [[Video]](https://www.youtube.com/watch?v=VcFja1fUNSk&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_06_4_keras_images.ipynb)
* **Part 6.5: Recognizing Multiple Images with YOLO Darknet** [[Video]](https://www.youtube.com/watch?v=oQcAKvBFli8&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_06_5_yolo.ipynb)
# Google CoLab Instructions
The following code ensures that Google CoLab is running the correct version of TensorFlow.
Running the following code will map your GDrive to ```/content/drive```.
```
try:
from google.colab import drive
drive.mount('/content/drive', force_remount=True)
COLAB = True
print("Note: using Google CoLab")
%tensorflow_version 2.x
except:
print("Note: not using Google CoLab")
COLAB = False
# Nicely formatted time string
def hms_string(sec_elapsed):
h = int(sec_elapsed / (60 * 60))
m = int((sec_elapsed % (60 * 60)) / 60)
s = sec_elapsed % 60
return f"{h}:{m:>02}:{s:>05.2f}"
```
# Part 6.5: Recognizing Multiple Images with Darknet
Programmers typically design convolutional neural networks to classify a single item centered in an image. However, as humans, we can recognize many items in our field of view in real-time. It is advantageous to be able to recognize multiple items in a single image. One of the most advanced means of doing this is YOLO DarkNet (not to be confused with the Internet [Darknet](https://en.wikipedia.org/wiki/Darknet). YOLO [[Cite:redmon2016you]](https://arxiv.org/abs/1506.02640) is an acronym for You Only Look Once. The fact that YOLO must only look once speaks to the efficiency of the algorithm. In this context, to "look" means to perform one scan over the image. Figure 6.YOLO shows YOLO tagging in action.
**Figure 6.YOLO: YOLO Tagging**

It is also possible to run YOLO on live video streams. Figure 6.VIDEO is from the YouTube Video for this module.
**Figure 6.VIDEO: YOLO Video Tagging**

As you can see, it is classifying many things in this video. My collection of books behind me is adding considerable "noise," as DarkNet tries to classify every book behind me. If you watch the video, you can see that it is less than perfect. The coffee mug that I pick up gets classified as a cell phone and, at times, a remote. The small yellow object behind me on the desk is a small toolbox (not a remote). However, it gets classified as a book at times and a remote at other times. Currently, this algorithm classifies each frame on its own. The program could achieve greater accuracy if it analyzed multiple images from a video stream. Consider when you see an object coming towards you, if it changes angles, you might form a better opinion of what it was. If that same object now changes to an unfavorable angle, you still know what it is, based on previous information.
### How Does DarkNet/YOLO Work?
YOLO begins by resizing the image to a $S \times S$ grid. YOLO runs a single convolutional neural network against this grid that predicts bounding boxes and what might be contained by those boxes. Each bounding box also has a confidence in which item it believes the box contains. YOLO is a regular convolution network, just like we've seen previously. The only difference is that a YOLO CNN outputs multiple prediction bounding boxes. At a high level, Figure 6.YOLO-DET illustrates this.
**Figure 6.YOLO-DET: The YOLO Detection System**

The output of the YOLO convolutional neural networks is essentially a multiple regression. YOLO generated the following values for each of the bounding rectangles.
* **x** - The x-coordinate of the center of a bounding rectangle.
* **y** - The y-coordinate of the center of a bounding rectangle.
* **w** - The width of each bounding rectangle.
* **h** - The height of each bounding rectangle.
* **labels** - The relative probabilities of each of the labels (1 value for each label)
* **confidence** - The confidence in this rectangle.
The output layer of a Keras neural network is a Tensor. In the case of YOLO, this output tensor is 3D and is of the following dimensions.
$ S \times S \times (B \cdot 5 + C) $
The constants in the above expression are:
* *S* - The dimensions that YOLO overlays across the source image.
* *B* - The number of potential bounding rectangles generated for each grid cell.
* *C* - The number of class labels that here are.
The value 5 in the above expression is simply the count of non-label components of each bounding rectangle ($x$, $y$, $h$, $w$, $confidence$.
Because there are $S^2 \cdot B$ total potential bounding rectangles, the image is nearly full. Because of this, it is essential to drop all rectangles below some threshold of confidence. The image below demonstrates this.
The actual structure of the convolutional neural network behind YOLO is relatively simple, as the following figure illustrates. Because there is only one convolutional neural network, and it "only looks once," the performance is not impacted by how many objects are detected. Figure 6.YOLO-STRUCT shows the YOLO structure.
**Figure 6.YOLO-STRUCT: YOLO Structure**

Figure 6.YOLO-DEMO shows some additional recognitions performed by a YOLO.
**Figure 6.YOLO-DEMO: YOLO Structure**

### Using YOLO in Python
To make use of YOLO in Python, you have several options:
* **[DarkNet](https://pjreddie.com/darknet/yolo/)** - The original implementation of YOLO, written in C.
* **[yolov3-tf2](https://github.com/zzh8829/yolov3-tf2)** - An unofficial Python package that implements YOLO in Python, using TensorFlow 2.0.
The code provided in this notebook works equally well when run either locally or from Google CoLab. In either case, the programmer should use TensorFlow 2.0.
### Installing YoloV3-TF2
YoloV3-TF2 is not available directly through either PIP or CONDA. Additionally, YoloV3-TF2 is not installed in Google CoLab by default. Therefore, whether you wish to use YoloV3-TF2 through CoLab or run it locally, you need to go through several steps to install it. This section describes the process of installing YoloV3-TF2. The same steps apply to either CoLab or a local install. For CoLab, you must repeat these steps each time the system restarts your virtual environment. For a local install, you must perform these steps only once for your virtual Python environment. If you are installing locally, make sure to install to the same virtual environment that you created for this course. The following command installs YoloV3-TF2 directly from it's GitHub repository.
```
import sys
!{sys.executable} -m pip install git+https://github.com/
zzh8829/yolov3-tf2.git@master
```
Before you can make use of YoloV3-TF2 there are several files you must obtain:
* **yolov3.weights** - These are the pre-trained weights provided by the author of YOLO.
* **convert.py** - This is a Python script that converts **yolov3.weights** into a TensorFlow compatible weight format.
* **coco.names** - The names of the 80 items that the **yolov3.weights** neural network was trained to recognize.
* **yolov3.tf** - The YOLO weights converted to a format that TensorFlow can use directly.
The code provided below obtains these files. The script stores these files to either your GDrive, if you are using CoLab, or a local folder named "data" if you are running locally.
Researchers have trained YOLO on a variety of different computer image datasets. The version of YOLO weights used in this course is from the dataset Common Objects in Context (COCO). [[Cite: lin2014microsoft]](https://arxiv.org/abs/1405.0312) This dataset contains images labeled into 80 different classes. COCO is the source of the file coco.txt that used in this module.
Developers have also adapted YOLO for mobile devices by creating the YOLO Tiny pre-trained weights that use a much smaller convolutional neural network and still achieve acceptable levels of quality. Though YoloV3-TF2 can work with either YOLO Tiny or regular YOLO we are not using the tiny weights for this course.
```
import tensorflow as tf
import os
if COLAB:
ROOT = '/content/drive/My Drive/projects/t81_558_dlearning/yolo'
else:
ROOT = os.path.join(os.getcwd(),'data')
filename_darknet_weights = tf.keras.utils.get_file(
os.path.join(ROOT,'yolov3.weights'),
origin='https://pjreddie.com/media/files/yolov3.weights')
TINY = False
filename_convert_script = tf.keras.utils.get_file(
os.path.join(os.getcwd(),'convert.py'),
origin='https://raw.githubusercontent.com/zzh8829/'\
"yolov3-tf2/master/convert.py')
filename_classes = tf.keras.utils.get_file(
os.path.join(ROOT,'coco.names'),
origin='https://raw.githubusercontent.com/zzh8829/'\
'yolov3-tf2/master/data/coco.names')
filename_converted_weights = os.path.join(ROOT,'yolov3.tf')
```
### Transfering Weights
In the course, we transfer already trained weights into our YOLO networks. It can take considerable time to train a YOLO network from scratch. If you would like to train a YOLO network to recognize images other than the COLO provided images, then you may need to train your own YOLO information. If training from scratch is something you need to do, there is further information on this at the YoloV3-TF2 GitHub repository.
The weights provided by the original authors of YOLO is not directly compatible with TensorFlow. Because of this, it is necessary first to convert the YOLO provided weights into a TensorFlow compatible format. The following code does this conversion. This process does not need to be repeated by the program. Once the conversion script processes YOLO weights the saved to the yolov3.tf YOLO can reuse these converted wights. The following code performs this conversion.
```
import sys
!{sys.executable} "{filename_convert_script}" --weights
"{filename_darknet_weights}" --output "{filename_converted_weights}"
```
The conversion script is no longer needed once this script converts the YOLO weights have to a TensorFlow format. Because this executable file resides in the same directory as the course files, we delete it at this point.
```
import os
os.remove(filename_convert_script)
```
Now that we have all of the files needed for YOLO, we are ready to use it to recognize components of an image.
### Running DarkFlow (YOLO)
The YoloV3-TF2 library can easily integrate with Python applications. The initialization of the library consists of three steps. First, it is essential to import all of the needed packages for the library. Next, the Python program must define all of the YOLO configurations through the Keras flags architecture. The Keras flag system primarily works from the command line; however, it also allows configuration programmatically in an application. For this example, we configure the package programmatically. Finally, we must scan available devices so that our application takes advantage of any GPUs. The following code performs all three of these steps.
```
import time
from absl import app, flags, logging
from absl.flags import FLAGS
import cv2
import numpy as np
import tensorflow as tf
from yolov3_tf2.models import (YoloV3, YoloV3Tiny)
from yolov3_tf2.dataset import transform_images, load_tfrecord_dataset
from yolov3_tf2.utils import draw_outputs
import sys
from PIL import Image, ImageFile
import requests
# Flags are used to define several options for YOLO.
flags.DEFINE_string('classes', filename_classes, 'path to classes file')
flags.DEFINE_string('weights', filename_converted_weights, 'path to weights file')
flags.DEFINE_boolean('tiny', False, 'yolov3 or yolov3-tiny')
flags.DEFINE_integer('size', 416, 'resize images to')
flags.DEFINE_string('tfrecord', None, 'tfrecord instead of image')
flags.DEFINE_integer('num_classes', 80, 'number of classes in the model')
FLAGS([sys.argv[0]])
# Locate devices to run YOLO on (e.g. GPU)
physical_devices = tf.config.experimental.list_physical_devices('GPU')
if len(physical_devices) > 0:
tf.config.experimental.set_memory_growth(physical_devices[0], True)
```
It is important to understand that Keras flags can only be defined once. If you are going to classify more than one image, make sure that you do not define the flags additional times.
The following code initializes a YoloV3-TF2 classification object. The weights are loaded, and the object is ready for use as the **yolo** variable. It is not necessary to reload the weights and obtain a new **yolo** variable for each classification.
```
# This example does not use the "Tiny version"
if FLAGS.tiny:
yolo = YoloV3Tiny(classes=FLAGS.num_classes)
else:
yolo = YoloV3(classes=FLAGS.num_classes)
# Load weights and classes
yolo.load_weights(FLAGS.weights).expect_partial()
print('weights loaded')
class_names = [c.strip() for c in open(FLAGS.classes).readlines()]
print('classes loaded')
```
Next, we obtain an image to classify. For this example, the program loads the image from a URL. YoloV3-TF2 expects that the image is in the format of a Numpy array. An image file, such as JPEG or PNG, is converted into this raw Numpy format by calling the TensorFlow **decode_image** function. YoloV3-TF2 can obtain images from other sources, so long as the program first decodes them to raw Numpy format. The following code obtains the image in this format.
```
# Read image to classify
url = "https://raw.githubusercontent.com/jeffheaton/"\
"t81_558_deep_learning/master/images/cook.jpg"
response = requests.get(url)
img_raw = tf.image.decode_image(response.content, channels=3)
```
At this point, we can classify the image that was just loaded. The program should preprocess the image so that it is the size expected by YoloV3-TF2. Your program also sets the confidence threshold at this point. Any sub-image recognized with confidence below this value is not returned by YOLO.
```
# Preprocess image
img = tf.expand_dims(img_raw, 0)
img = transform_images(img, FLAGS.size)
# Desired threshold (any sub-image below this confidence
# level will be ignored.)
FLAGS.yolo_score_threshold = 0.5
# Recognize and report results
t1 = time.time()
boxes, scores, classes, nums = yolo(img)
t2 = time.time()
print(f"Prediction time: {hms_string(t2 - t1)}")
```
It is important to note that the **yolo** class instantiated here is a callable object, which means that it can fill the role of both an object and a function. Acting as a function, *yolo* returns three arrays named **boxes**, **scores**, and **classes** that are of the same length. The function returns all sub-images found with a score above the minimum threshold. Additionally, the **yolo** function returns an array named called **nums**. The first element of the **nums** array specifies how many sub-images YOLO found to be above the score threshold.
* **boxes** - The bounding boxes for each of the sub-images detected in the image sent to YOLO.
* **scores** - The confidence for each of the sub-images detected.
* **classes** - The string class names for each of the items. These are COCO names such as "person" or "dog."
* **nums** - The number of images above the threshold.
Your program should use these values to perform whatever actions you wish as a result of the input image. The following code simply displays the images detected above the threshold.
```
print('detections:')
for i in range(nums[0]):
cls = class_names[int(classes[0][i])]
score = np.array(scores[0][i])
box = np.array(boxes[0][i])
print(f"\t{cls}, {score}, {box}")
```
Your program should use these values to perform whatever actions you wish as a result of the input image. The following code simply displays the images detected above the threshold.
YoloV3-TF2 includes a function named **draw_outputs** that allows the sub-image detections to visualized. The following image shows the output of the draw_outputs function. You might have first seen YOLO demonstrated as an image with boxes and labels around the sub-images. A program can produce this output with the arrays returned by the **yolo** function.
```
# Display image using YOLO library's built in function
img = img_raw.numpy()
img = draw_outputs(img, (boxes, scores, classes, nums), class_names)
#cv2.imwrite(FLAGS.output, img) # Save the image
display(Image.fromarray(img, 'RGB')) # Display the image
```
# Module 6 Assignment
You can find the first assignment here: [assignment 6](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/assignments/assignment_yourname_class6.ipynb)
| github_jupyter |
# Chapter 5
# Introduction to Numpy
```
!pip install -q numpy
```
Numpy mean numerical python.
It is use for numerical operation and manipulation. Especially in array or matrix.
| 1 | 2 | 3 |
| --- | --- | ---|
| 4 | 5 | 6 |
| 7 | 8 | 9 |
order 3 x 3
```
# import package numpy and give it an alias np
import numpy as np
a = np.array(
[
[1,2,3],
[4,5,6],
[7,8,9]
]
)
print(a)
print("dim = ", a.ndim)
print("shape = ", a.shape)
print("datatype = ", a.dtype)
print("size = ", a.size)
print()
alist = [
[1,2,3],
[4,5,6],
[7,8,9]
]
print(alist)
print(len(alist))
# print(size(alist))
# print(shape(alist))
print(type(a))
print(type(alist))
import numpy as np
blist = [31, 32, 33]
b = np.array(blist)
print(blist)
print()
print(b)
print("dim = ", b.ndim)
print("shape = ", b.shape)
print("datatype = ", b.dtype)
print("size = ", b.size)
print()
import numpy as np
clist = [1, 'Tue', 3, 'Wed']
c = np.array(clist)
print(clist)
print()
print(c)
print("dim = ", c.ndim)
print("shape = ", c.shape)
print("datatype = ", c.dtype)
print("size = ", c.size)
print()
```
Class activity
| 11 | 12 | 13 |
| --- | --- | ---|
order 1 x 3
or row matrix
$$
\begin{pmatrix}
11 & 12 & 13
\end{pmatrix}
$$
### Without specifying the datatype.
```
# import package numpy and give it an alias np
import numpy as np
a = np.array(
[
[11, 12, 13]
]
)
print(a)
print("dim = ", a.ndim)
print("shape = ", a.shape)
print("datatype = ", a.dtype)
print("size = ", a.size)
print()
```
### Specifying datatype int64
```
# import package numpy and give it an alias np
import numpy as np
a = np.array(
[
[11, 12, 13]
], dtype='int64'
)
print(a)
print("dim = ", a.ndim)
print("shape = ", a.shape)
print("datatype = ", a.dtype)
print("size = ", a.size)
print()
```
### Specifying datatype float
```
# import package numpy and give it an alias np
import numpy as np
a = np.array(
[
[11, 12, 13]
], dtype='float'
)
print(a)
print("dim = ", a.ndim)
print("shape = ", a.shape)
print("datatype = ", a.dtype)
print("size = ", a.size)
print()
```
### Specifying datatype float32
```
# import package numpy and give it an alias np
import numpy as np
a = np.array(
[
[11, 12, 13]
], dtype='float32'
)
print(a)
print("dim = ", a.ndim)
print("shape = ", a.shape)
print("datatype = ", a.dtype)
print("size = ", a.size)
print()
```
### float value
```
# import package numpy and give it an alias np
import numpy as np
a = np.array(
[
[11.0, 12.0, 13.0]
]
)
print(a)
print("dim = ", a.ndim)
print("shape = ", a.shape)
print("datatype = ", a.dtype)
print("size = ", a.size)
print()
```
## Class activity
| 21 |
|---|
| 22 |
| 31 |
| 0 |
order 4 x 1
or column matrix
$$
\begin{pmatrix}
21 \\
22 \\
31 \\
0 \\
\end{pmatrix}
$$
```
# import package numpy and give it an alias np
import numpy as np
a = np.array(
[
[21],
[22],
[31],
[0],
], dtype='uint8'
)
print(a)
print("dim = ", a.ndim)
print("shape = ", a.shape)
print("datatype = ", a.dtype)
print("size = ", a.size)
print()
```
Minimum value in 8 bits
|7|6|5|4|3|2|1|0|
|---|---|---|---|---|---|---|---|
|0|0|0|0|0|0|0|0|
Maximum value in 8 bits
|7|6|5|4|3|2|1|0|
|---|---|---|---|---|---|---|---|
|1|1|1|1|1|1|1|1|
uint8 mean unsigned integer in 8 bits
min = 0
max = 255
int8 mean signed integer in 8 bits
min = -128
max = 127
## Installation of numpy, scipy and pandas
```
!pip install -q numpy
def array_properties(a):
print("a = \n", a)
print("dim = ", a.ndim)
print("shape = ", a.shape)
print("datatype = ", a.dtype)
print("size = ", a.size)
print()
```
## Creating array
```
import numpy as np
a = np.array([1, 2, 3])
print(a)
print()
print("a = \n", a)
print("dim = ", a.ndim)
print("shape = ", a.shape)
print("datatype = ", a.dtype)
print("size = ", a.size)
print()
import numpy as np
a_3x3 = np.array([[1, 1, 1], [2, 2, 2], [3, 3, 3]])
array_properties(a_3x3)
```
## arange
integer array creation.
range used regularly in for loop
```
for n in range(10):
print(n)
start = 1
stop = 10 + 1
for n in range(start,stop):
print(n, end=', ')
start = 1
stop = 10 + 1
step = 2
for n in range(start, stop, step):
print(n, end=', ')
import numpy as np
seq_a = np.arange(1, 10)
array_properties(seq_a)
import numpy as np
ar10 = np.arange(10)
array_properties(ar10)
for n in ar10:
print(n, end=' | ')
start = 1
stop = 10 + 1
step = 2
ar10b = np.arange(start, stop, step)
array_properties(ar10b)
for n in ar10b:
print(n, end=' | ')
start = 1
stop = 10 + 1
step = 2
ar10b = np.arange(start, stop, step)
ar10b = np.uint8(ar10b)
array_properties(ar10b)
for n in ar10b:
print(n, end=' | ')
print()
print()
ar10b = np.float32(ar10b)
array_properties(ar10b)
```
## Class Activity
Create an array of integers [0, 5, 15, ..., 100]
## linspace
create array of floating point value of a specific size.
```
import numpy as np
seq_a2 = np.linspace(1, 10, 15)
array_properties(seq_a2)
```
|1|2|3|4|5|6|7|8|9|10|
|---|---|---|---|---|---|---|---|---|---|
|0|1|2|3|4|5|6|7|8|9|
|0|x|x|x|x|x|x|x|x|10|
0 --> 0
1 --> x
2 --> x
9 --> 10
```
x = 10/9 * 9
x
import numpy as np
start = 0
stop = 10
size = 10
arls_0_10_10 = np.linspace(start, stop, size)
array_properties(arls_0_10_10)
import numpy as np
start = 0
stop = 10
size = 11
arls_0_10_11 = np.linspace(start, stop, size)
array_properties(arls_0_10_11)
```
## zeros
create array of zeros
```
import numpy as np
zer_10 = np.zeros((1,10))
array_properties(zer_10)
import numpy as np
zer_10a = np.zeros((1,10), dtype='float32')
array_properties(zer_10a)
import numpy as np
zer_10 = np.zeros((1,10), dtype=np.float32)
array_properties(zer_10)
import numpy as np
zer_10c = np.zeros((1,10), dtype='int')
array_properties(zer_10c)
import numpy as np
zer_10 = np.zeros((1,10), dtype=np.uint8)
array_properties(zer_10)
import numpy as np
zeros_arr = np.zeros((2, 4))
array_properties(zeros_arr)
import numpy as np
zer_4_5_3 = np.zeros((4,5,3), dtype=np.uint8)
array_properties(zer_4_5_3)
import numpy as np
zer_4_5_3_2 = np.zeros((4,5,3,2), dtype=np.uint8)
array_properties(zer_4_5_3_2)
```
## ones
Create array of ones
```
import numpy as np
# specify the shape of the array in the `np.ones` function.
ones_arr = np.ones((4, 2))
array_properties(ones_arr)
import numpy as np
shape = (1,10)
ones_10 = np.ones(shape)
array_properties(ones_10)
import numpy as np
shape = (10,)
ones_10 = np.ones(shape)
array_properties(ones_10)
```
## Class Activity
Create an array of dim 3 filled with ones.
# Recap
1. Create array from list value
2. convert list to array.
3. Convert arrays of diffrent types (int32, int64, float64, float32, uint8, <U11)
4. Create array of zeros
5. Create array of ones
```
import numpy as np
emp_arr = np.empty((4, 4))
array_properties(emp_arr)
```
## Class Activity
Create empty array of 1x10
```
import numpy as np
emp_1x10 = np.empty((1, 10))
array_properties(emp_1x10)
```
## Reshape
Create an array of 1x10 then reshape to 2x5
```
import numpy as np
ar1x10 = np.arange(1, 11)
array_properties(ar1x10)
# reshape class method
ar2x5 = ar1x10.reshape((2,5))
array_properties(ar2x5)
# reshape function of np
ar2x5 = np.reshape(ar1x10, (2,5))
array_properties(ar2x5)
import numpy as np
arls10 = np.linspace(10, 55, 10)
array_properties(arls10)
# reshape into wrong container
ar3x3 = ar1x10.reshape((3,3))
array_properties(ar3x3)
import numpy as np
a1 = np.arange(1, 13)
array_properties(a1)
a2 = np.reshape(a1, (3, 4))
array_properties(a2)
```
Reshape knowing only one dim size.
```
import numpy as np
arls10 = np.linspace(10, 55, 10)
array_properties(arls10)
# reshape into 5xunknown
ar5x_ = ar1x10.reshape((5,-1))
array_properties(ar5x_)
# reshape into unknown x5
ar_x5 = ar1x10.reshape((-1,5))
array_properties(ar_x5)
import numpy as np
arls10 = np.linspace(10, 55, 10)
array_properties(arls10)
# reshape into 3xunknown
ar3x_ = ar1x10.reshape((3,-1))
array_properties(ar3x_)
import numpy as np
a1 = np.arange(1, 13, 1.5)
array_properties(a1)
a2 = np.reshape(a1, (4, -1))
array_properties(a2)
import numpy as np
a1 = np.linspace(1, 10, 15).reshape(5, -1)
array_properties(a1)
```
## Class Activity
Create an array of shape 4x5 and reshape into 2x-1.
```
import numpy as np
ar20 = np.linspace(1,21,20)
array_properties(ar20)
ar4x5 = ar20.reshape((4,5))
array_properties(ar4x5)
ar2x_ = ar4x5.reshape((2,-1))
array_properties(ar2x_)
```
## Class Activity
Create an array of 24 elements or size.
Reshape into multiples of 3 and 4 in row dimension.
```
import numpy as np
# create a 1 dim array of size 24
# reshape into array o 3x-1
# reshape into array of 4x-1
```
## Class Activity
Create an array of unsugned integer 8 bits of size 300 and reshape into (10,10,-1).
```
ar300 = np.arange(0,300,dtype='uint8')
print(ar300.shape)
ar10x10x_ = ar300.reshape((10,10,-1))
print(ar10x10x_.shape)
# 10,-1,3
ar10x_x3 = ar300.reshape((10,-1,3))
print(ar10x_x3.shape)
# -1,10,3
ar_x10x3 = ar300.reshape((-1,10,3))
print(ar_x10x3.shape)
# -1,10,3
ar_x_x3 = ar300.reshape((-1,-1,3))
print(ar_x_x3.shape)
```
## random
`rand` will geneartion random values between 0 - 1. It accepts the shape of the array to be created.
```
import numpy as np
# random array between 0-1 with shape (4,5)
a1 = np.random.rand(4, 5)
array_properties(a1)
```
Class Activity
Create an array with shape (3,7) of values between 0-1.
## randint
Creates an array of integer.
It accepts start, stop and shape.
`randint(start, stop, shape)`
```
import numpy as np
a1 = np.random.randint(0, 10, (4,5))
array_properties(a1)
```
## Class Activity
Create an array of unsigned integer of 8 bits with shape (100, 100, 3) and filled with random integer values.
## Accessing an array element
### 1 D Array
```
import numpy as np
a1 = np.arange(1, 13)
array_properties(a1)
```
|index|0|1|2|3|4|5|6|7|8|9|10|11|
|---|---|---|---|---|---|---|---|---|---|---|---|---|
|value|1|2|3|4|5|6|7|8|9|10|11|12|
|special index|-12|-11|-10|-9|-8|-7|-6|-5|-4|-3|-2|-1|
```
print('first: ', a1[0])
print('second: ', a1[1])
print('third: ', a1[2])
print('ninth: ', a1[9-1])
print('last: ', a1[11])
print('last: ', a1[-1])
# a1[istart:istop]
# stop or istop excluded
print('index 0 to 4: ', a1[0:4])
# omitte the istart
print('index 0 to 4: ', a1[:4])
print('index 3 to 6: ', a1[3:6])
print('index 0 to 10: ', a1[0:10])
print('index 0 to 10: ', a1[:10])
# omitt istop equal last index
print('index 3 to last: ', a1[3:12])
print('index 3 to last: ', a1[3:])
# from istart to second to the last value
# index -1 equal last index.
print('index 3 to second to the last: ', a1[3:-1])
print('last 2 numbers: ', a1[10:])
print('last 2 numbers: ', a1[-2:])
# get odd positions
# a1[istart : istop : istep]
print('odd positions: ', a1[0::2])
print('even positions: ', a1[1::2])
import numpy as np
N = 12
a1 = np.arange(N+1)
array_properties(a1)
```
Use for loop on array.
```
for i in range(N+1):
print(f'{i}: {a1[i]}')
```
# Assignment
1. Create an array of shape (10, 10) with uint8 random values.
2. print the shape, size and dimension,
3. Compute the sum of the third row to the last.
4. Compute the average of the columns
### 2 D Array
```
import numpy as np
a14 = np.arange(14+1)
print('size of a14=', a14.size)
if (a14.size % 4) == 0:
reshape_a = a14.reshape((4,-1))
print(reshape_a.shape)
# divide result in float
print('16 / 3=', 16 / 3)
# remainder
print('16 % 3=', 16 % 3)
# whole or integer
print('16 // 3', 16 // 3, sep='=')
print('16 / 4=', 16 / 4)
print('16 % 4=', 16 % 4)
print('16 // 4=', 16 // 4)
import numpy as np
a1 = np.arange(1, 17)
a2 = np.resize(a1, (4,4))
array_properties(a2)
print('Element in row=1, column=1: ', a2[0,0])
print('Element in row=3, column=1: ', a2[2,0])
print('Element in row=2, column=3: ', a2[1,2])
array_properties(a2)
# row & index
row = 1
i = row - 1
# column & index
col = 2
j = col - 1
print(f'Element in row={row}, column={col}: {a2[i,j]}')
# get all elements in row 2
print(f'Elements in row= 2: {a2[1,:]}')
# alternative: get all elements in row 2 or dimension 1
print(f'Elements in row= 2: {a2[1]}')
# get all value in column 2 or dimension
print('Elements in col= 2: ', a2[:,1])
# alternative: wrong get all value in column 2 or dimension 2
print('Elements in col= 2: ', a2[,1])
print(f'Elements in rows= 1 to 3 and columns= 2 to 4: \n', a2[0:3,1:3])
print()
print(f'Elements in row index= 0 to 2 and column index= 1 to 2: \n', a2[0:3,1:3])
array_properties(a2)
rows, columns = a2.shape
for i in range(rows):
for j in range(columns):
print(f'{i},{j}: {a2[i,j]}', end=', ')
print()
print()
rows, columns = a2.shape
for i in range(rows):
for j in range(columns):
print(a2[i,j], end=', ')
print()
```
### 3D Array
```
import numpy as np
a3 = np.arange(1, 9).reshape((2,2,2))
array_properties(a3)
print(a3[0,1,1])
#
print(a3[0,1,:])
print(a3[0,1])
#
print(a3[0,:,1])
print(a3[:,1,1])
a3D = np.array([
[
[111,112,113],
[121,122,123],
],
[
[211,212,213],
[221,222,223],
]
])
array_properties(a3D)
print('Element 1,2,2: ', a3D[0,1,1])
array_properties(a3)
rows_3d, rows, columns = a3.shape
for i in range(rows_3d):
for j in range(rows):
for k in range(columns):
print(f'{i},{j},{k}: {a3[i,j,k]}', end=', ')
print()
print()
print()
rows_3d, rows, columns = a3.shape
for i in range(rows_3d):
for j in range(rows):
for k in range(columns):
print(a3[i,j,k], end=', ')
print()
print()
```
## Changing Array Element
```
import numpy as np
a3x3 = np.arange(1,10).reshape((3,-1))
print(a3x3)
```
||0|1|2|
|---|---|---|--|
|0|1|2|3|
|1|4|5|6|
|2|7|8|9|
```
# print value 6
print(a3x3[1,2])
# print 3
print(a3x3[0,2])
# change 3 to 33
a3x3[0,2] = 33
print(a3x3)
```
||0|1|2|
|---|---|---|--|
|0|1|2|33|
|1|4|5|6|
|2|7|8|9|
```
# change value 8 to 18
print(a3x3[2,1])
a3x3[2,1] = 18
print(a3x3)
import numpy as np
a1 = np.arange(9)
array_properties(a1)
print('Third element: ', a1[2])
a1[2] = a1[2] ** 2
print('Third element squared: ', a1[2])
```
||0|1|2|3|4|5|6|7|8|
|---|---|---|---|---|---|---|---|---|---|
|0|0|1|2|3|4|5|6|7|8|
```
# 3 ^ 2 = 9
3 ** 2
import numpy as np
a2 = np.arange(1,10).reshape(3,3)
array_properties(a2)
print('2nd row, 2nd column element: ', a2[1,1])
a2[1,1] = a2[1,1] ** 2
print('2nd row, 2nd column element squared: ', a2[1,1])
print(a2)
print('Third row: ', a2[2,:])
a2[2] = a2[2,:] * 2
print('Third row doubled: ', a2[2,:])
print(a2)
print('First row: ', a2[0,:])
a2[0] = a2[0,:] + a2[1,:]
print('First row increased by second row: ', a2[0,:])
print(a2)
print('Second row: ', a2[1,:])
a2[1] = a2[1,:] - a2[2,:]
print('Second row decreased by third row: ', a2[1,:])
print(a2)
x = 3
y=4
print('x=', x, 'y=', y)
x = x - y
print('x=', x, 'y=', y)
```
## Class Activity
1. Create an array a of shape (2, 5) and another array b of shape (3, 5).
2. Create another array c = a
3. Change array c row 2 to the sum of array a row 2 and array b row 2.
| github_jupyter |
# > Desafio
Integrantes: \
-Hugo Rocha -- 201610531-K \
-Gabriel Vergara -- 201510519-7
Equipo: RNG
[Video de la defensa](https://youtu.be/Lo9bp2NuXZQ)
El presente es un trabajo original, desarrollado por los autores en conformidad con todas reglas de codigo de honor y honestidad. Sumado a esto ultimo
se deja en claro que la contribucion de ambos participantes en el presente trabajo fue la siguiente:
Gabriel Vergara:\
-Exploratorio. \
-Testeo inicial modelos de regresion.
Hugo Rocha:
-preprocesamiento. \
-Cross Validation para elecciond e modelos y parametros.
Ambos trabajamos de manera colaborativa en la formulacion de las conclusiones obtenidas en el presente trabajo.
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import plotly.graph_objs as go
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import plotly.graph_objs as go
import re
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split, cross_val_score
from sklearn.metrics import roc_curve, auc, confusion_matrix
from sklearn.metrics import accuracy_score
from sklearn.metrics import precision_score
from sklearn.metrics import confusion_matrix, ConfusionMatrixDisplay
from sklearn.linear_model import LinearRegression
from sklearn.linear_model import LogisticRegression
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis
import scipy.stats as stats
from scipy.stats import kurtosis,skew
import statsmodels.api as sm
import warnings
warnings.filterwarnings("ignore")
def rankear_hoteles(hotel_details):
lista = []
for i in hotel_details:
hoteles = i.split('|')
aux = []
aux2 = []
for j in range(len(hoteles)):
if hoteles[0] != 'Not Available':
aux.append(hoteles[j].split(':')[1])
aux2.append(hoteles[j].split(':')[0])
lista.append([aux2,aux])
hoteles_todos = []
scores_todos = []
for i in lista:
hoteles_todos = [*hoteles_todos,*i[0]]
scores_todos = [*scores_todos,*i[1]]
d = {'hoteles': hoteles_todos, 'scores': scores_todos}
df = pd.DataFrame(data=d)
hoteles_unicos = df['hoteles'].unique()
score = []
for i in range(len(hoteles_unicos)):
hotel_idx = df['hoteles'] == hoteles_unicos[i]
n = np.mean(pd.to_numeric(df[hotel_idx]['scores']))
score.append(n)
hotel_rank = {'hotel': hoteles_unicos, 'scores': score}
Hotel_rank = pd.DataFrame(data=hotel_rank)
media = np.mean(Hotel_rank['scores'])
Hotel_rank['scores'] = Hotel_rank['scores'].replace(np.nan, media)
return Hotel_rank
def hotel_rank_mean(hotel_details,Hotel_rank):
hoteles = hotel_details.split('|')
hoteles
aux = []
for j in range(len(hoteles)):
if hoteles[0] != 'Not Available':
aux.append(hoteles[j].split(':')[0])
valores = []
for i in range(len(aux)):
valores.append(Hotel_rank[ Hotel_rank['hotel'] == aux[i]]['scores'])
return np.mean(valores)
def noches(itinerario):
u=itinerario.split('. ')
n = 0
for j in range(len(u)):
n = n + float(u[j].split('N')[0])
return n
def noches_ponderadas(itinerario,hotel_details,Hotel_rank):
u=itinerario.split('. ')
n = 0
hoteles = hotel_details.split('|')
aux = []
#print(u)
#print(hoteles)
for j in range(len(hoteles)):
if hoteles[0] != 'Not Available':
aux.append(hoteles[j].split(':')[0])
valores = []
for i in range(len(aux)):
valores.append(Hotel_rank[ Hotel_rank['hotel'] == aux[i]]['scores'])
if valores == []:
valores = np.mean(Hotel_rank['scores'])
for j in range(len(u)):
n = n + float(u[j].split('N')[0])*valores
else:
if len(u) == len(valores):
for j in range(len(u)):
#print(u[j].split('N')[0])
#print(valores[j])
#print(float(u[j].split('N')[0])*valores[j])
n = n + float(float(u[j].split('N')[0])*valores[j])
else:
#print(valores[0])
#print(u)
for j in range(len(u)):
#print(u[j].split('N')[0])
#print(valores[j])
#print(float(u[j].split('N')[0])*valores[j])
n = n + float(float(u[j].split('N')[0])*valores[0])
#j
return n
def dest_numb(destinos):
d=destinos.split(sep='|')
return len(d)
def mes(Travel_Date):
d=Travel_Date.split(sep='-')
return d[1]
def ano(Travel_Date):
d=Travel_Date.split(sep='-')
return d[2]
def paquete_numero(paquetes):
aux = paquetes
aux = aux.apply(lambda x: x.replace("Budget", "0"))
aux = aux.apply(lambda x: x.replace("Standard", "1"))
aux = aux.apply(lambda x: x.replace("Deluxe", "1"))
aux = aux.apply(lambda x: x.replace("Luxury", "2"))
aux = aux.apply(lambda x: x.replace("Premium", "2"))
return aux
def destinos(destinos):
lista_destinos = []
for i in destinos:
destino = i.split('|')
for j in range(len(destino)):
if destino[0] != 'Not Available':
lista_destinos.append(destino[j])
d = {'destino': lista_destinos}
df = pd.DataFrame(data=d)
destinos_unicos = df['destino'].unique()
return destinos_unicos
def obtener_destinos(destino, Destinos):
destino = destino.split('|')
#print(destino)
aux = np.zeros(len(Destinos)).astype(int)
for i in range(len(destino)):
aux = aux + (Destinos == destino[i]).astype(int)
#d = {Destinos: aux}
#df = pd.DataFrame(data=d)
return aux #pd.Series(aux,index=Destinos)
def todos_destinos(total_destinos):
lista = []
Destinos = destinos(total_destinos)
for i in total_destinos:
lista.append(obtener_destinos(i, Destinos))
ddf = pd.DataFrame(lista)
ddf.columns = Destinos
return ddf
def regla_cancelacion(Cancellation_Rules):
indice = Cancellation_Rules.unique()
Cancellation_Rules.replace({indice[0]: "Rule 1", "b": "y",
indice[1]: "Rule 2",
indice[2]: "Rule 3",
indice[3]: "Rule 4",
indice[4]: "Rule 5",
indice[5]: "Rule 6",
indice[6]: "Rule 7",
indice[7]: "Rule 8",}, inplace=True)
return Cancellation_Rules
def areglo_hoteles(Hotel_Details):
Hotel_Details = Hotel_Details.apply(lambda x: x.replace("Four", "4"))
Hotel_Details = Hotel_Details.apply(lambda x: x.replace("Two", "2"))
Hotel_Details = Hotel_Details.apply(lambda x: x.replace("Three", "3"))
Hotel_Details = Hotel_Details.apply(lambda x: x.replace("Five", "5"))
Hotel_Details = Hotel_Details.apply(lambda x: x.replace("The Lodhi:A member of The Leading Hotels Of The World", "The Lodhi"))
return Hotel_Details
def procesar_data(data_train):
data_train['Cancellation Rules'] = regla_cancelacion(data_train['Cancellation Rules'])
data_train['Hotel Details'] = areglo_hoteles(data_train['Hotel Details'])
Hotel_rank = rankear_hoteles(data_train['Hotel Details'])
data_train['total_noches'] = data_train['Itinerary'].apply(lambda row: noches(row))
data_train['hotel_qual_prom'] = data_train['Hotel Details'].apply(lambda row: hotel_rank_mean(row,Hotel_rank))
data_train['noches_pond_hotel'] = data_train.apply(lambda x: noches_ponderadas(x['Itinerary'], x['Hotel Details'],Hotel_rank), axis=1)
data_train['dest_numb'] = data_train['Destination'].apply(lambda row: dest_numb(row))
data_train['mes'] = data_train['Travel Date'].apply(lambda row: mes(row))
data_train['ano'] = data_train['Travel Date'].apply(lambda row: ano(row))
data_train['paquete'] = data_train['Package Type']
data_train['paquete'] = data_train['paquete'].apply(lambda x: x.replace("Luxury", "Premium"))
data_train['paquete'] = data_train['paquete'].apply(lambda x: x.replace("Deluxe", "Standard"))
data_train['Quarter'] = pd.to_datetime(data_train['Travel Date'].values, format='%d-%m-%Y').astype('period[Q]')
data_train['Quarter']= data_train['Quarter'].astype(str).apply(lambda x: x.replace("2021", ""))
data_train['Quarter'] = data_train['Quarter'].astype(str).apply(lambda x: x.replace("2022", ""))
# creando dummies
Package_Type=pd.get_dummies(data_train['paquete'])
Cancellation_Rules=pd.get_dummies(data_train['Cancellation Rules'])
Ciudades = pd.get_dummies(data_train['Start City'])
Años = pd.get_dummies(data_train['ano'])
Quarter = pd.get_dummies(data_train['Quarter'])
media = np.mean(Hotel_rank['scores'])
data_train['hotel_qual_prom'] = data_train['hotel_qual_prom'].fillna(media)
Destinos = todos_destinos(data_train['Destination'])
paquete_numeros = paquete_numero(data_train['Package Type'])
Sightseeing = todos_destinos(data_train['Sightseeing Places Covered']) # altamente dimensional
#data_train=data_train.drop(['Unnamed: 0','Itinerary','Destination',
# 'Package Name','Places Covered',
# 'Travel Date','Hotel Details','Start City',
# 'Airline','Sightseeing Places Covered',
# 'Package Type','Cancellation Rules',
# 'mes','ano','Quarter','paquete',
# ],axis=1)
data_train = pd.concat([data_train, Package_Type,Cancellation_Rules,
Años,Quarter,paquete_numeros,
Destinos,Sightseeing], axis=1, join="inner") #,Sightseeing
return data_train,Destinos,Sightseeing
def mape(actual, pred):
actual, pred = np.array(actual), np.array(pred)
return np.mean(np.abs((actual - pred) / actual)) * 100
def box_cox(y,lam):
if lam == 0:
return np.log(y)
else:
return (y**lam-1)/lam
def inv_box_cox(y,l):
if l == 0:
return np.exp(y)
else:
return (y*l+1)**(1/l)
data_train = pd.read_csv("travel_packages_train.csv")
data_test = pd.read_csv("travel_packages_test.csv")
```
# Procesar Data
```
datos_train,destinos_train,Sightseeing_train = procesar_data(data_train)
datos_test,destinos_test,Sightseeing_test = procesar_data(data_test)
# para dar formato al test
train_aux = destinos_train*0
train_aux[destinos_test.columns] = destinos_test
train_aux= train_aux[0:datos_test.shape[0]]
datos_test = datos_test.drop(list(destinos_test.columns), axis = 1)
datos_test = pd.concat([datos_test,train_aux], axis=1, join="inner")
# para dar formato al test
train_aux = Sightseeing_train*0
train_aux[Sightseeing_test.columns] = Sightseeing_test
train_aux= train_aux[0:datos_test.shape[0]]
datos_test = datos_test.drop(list(Sightseeing_test.columns), axis = 1)
datos_test = pd.concat([datos_test,train_aux], axis=1, join="inner")
```
# Exploratorio
```
sns.displot(data_train, x="PPPrice")
print( 'excess kurtosis of normal distribution (should be 0): {}'.format( kurtosis(data_train['PPPrice']) ))
print( 'skewness of normal distribution (should be 0): {}'.format( skew(data_train['PPPrice']) ))
sns.displot(box_cox(data_train['PPPrice'],-0.1))
print( 'excess kurtosis of normal distribution (should be 0): {}'.format( kurtosis(box_cox(data_train['PPPrice'],-0.1)) ))
print( 'skewness of normal distribution (should be 0): {}'.format( skew(box_cox(data_train['PPPrice'],-0.1)) ))
data_train.boxplot(column=['PPPrice'],by="dest_numb")
data_train.boxplot(column=['PPPrice'],by="total_noches")
data_train.boxplot(column=['PPPrice'],by="Package Type")
plt.scatter(data_train['hotel_qual_prom'],data_train['PPPrice'])
plt.scatter(data_train['noches_pond_hotel'],data_train['PPPrice'])
data_train.boxplot(column=['PPPrice'],by="Package Type")
data_train.boxplot(column=['PPPrice'],by="paquete")
data_train.boxplot(column=['PPPrice'],by="ano")
data_train.boxplot(column=['PPPrice'],by="mes")
data_train.boxplot(column=['PPPrice'],by="Quarter")
```
# Entrenamiento
```
datos_train=datos_train.drop(['Unnamed: 0','Itinerary','Destination',
'Package Name','Places Covered',
'Travel Date','Hotel Details','Start City',
'Airline',
'Package Type','Cancellation Rules','Quarter','paquete',
],axis=1)
datos_train
datos=datos_train.drop(['Meals','Sightseeing Places Covered','ano','mes','hotel_qual_prom','Rule 2','Rule 3',
'Rule 4','Rule 5','Rule 6','Rule 7',
'Rule 8','2021','Q1','Q2','Q3',#' Gwalior fort ',
'Budget'],axis=1)
datos
datos['dest_num_square']=datos['dest_numb']**2
datos['total_noches_sqr']=(datos['total_noches'])**(1/2)
datos['noches_pond_hotel_log']=np.log(datos['noches_pond_hotel'])
y = datos['PPPrice']
X = datos.drop(['PPPrice',' Gwalior fort '], axis=1)
X_train, X_val, y_train, y_val = train_test_split(X,y,test_size=0.2, random_state=44)
```
## Prueba de modelos
### OLS
```
XX = datos.drop(['PPPrice','Q4'], axis=1)#,Sightseeing_train.columns
listas = list(set(list(Sightseeing_train.columns)) )
XX = XX.drop(listas, axis = 1)
XX_train, XX_val, yy_train, yy_val = train_test_split(XX,y,test_size=0.2, random_state=44)
reg = LinearRegression().fit(XX_train, yy_train)
yy_pred = reg.predict(XX_val)
mape(yy_val,yy_pred)
residuos = yy_train - reg.predict(XX_train)
sns.displot(residuos)
print( 'excess kurtosis of normal distribution (should be 0): {}'.format( kurtosis(residuos) ))
print( 'skewness of normal distribution (should be 0): {}'.format( skew(residuos) ))
shapiro_test = stats.shapiro(residuos)
shapiro_test
```
#### box.cox-ols
```
l = -0.1
reg_cox = LinearRegression().fit(X_train, box_cox(y_train,l))
y_pred = reg_cox.predict(X_val)
mape(y_val,inv_box_cox(y_pred,l))
residuos = box_cox(y_train,l) - reg_cox.predict(X_train)
sns.displot(residuos)
print( 'excess kurtosis of normal distribution (should be 0): {}'.format( kurtosis(residuos) ))
print( 'skewness of normal distribution (should be 0): {}'.format( skew(residuos) ))
```
### LASSO
```
from sklearn.linear_model import Lasso
model = Lasso(alpha= 1)
model.fit(X_train, y_train)
y_pred = model.predict(X_val)
mape(y_val,y_pred)
residuos = y_train - model.predict(X_train)
sns.displot(residuos)
print( 'excess kurtosis of normal distribution (should be 0): {}'.format( kurtosis(residuos) ))
print( 'skewness of normal distribution (should be 0): {}'.format( skew(residuos) ))
```
#### box-cox-lasso
```
l = -0.1
model_cox = Lasso(alpha=0.01)
model.fit(X_train, box_cox(y_train,l))
y_pred = model.predict(X_val)
mape(y_val,inv_box_cox(y_pred,l))
residuos = box_cox(y_train,l) - model.predict(X_train)
sns.displot(residuos)
print( 'excess kurtosis of normal distribution (should be 0): {}'.format( kurtosis(residuos) ))
print( 'skewness of normal distribution (should be 0): {}'.format( skew(residuos) ))
```
### Ridge
```
from sklearn.linear_model import Ridge
model1 = Ridge(alpha=1)
model.fit(X_train, y_train)
y_pred = model.predict(X_val)
mape(y_val,y_pred)
residuos = y_train - model.predict(X_train)
sns.displot(residuos)
print( 'excess kurtosis of normal distribution (should be 0): {}'.format( kurtosis(residuos) ))
print( 'skewness of normal distribution (should be 0): {}'.format( skew(residuos) ))
```
#### box-cox - ridge
```
l = -0.1
model = Ridge(alpha=10)
model.fit(X_train, box_cox(y_train,l))
y_pred = model.predict(X_val)
mape(y_val,inv_box_cox(y_pred,l))
residuos = box_cox(y_train,l) - model.predict(X_train)
sns.displot(residuos)
print( 'excess kurtosis of normal distribution (should be 0): {}'.format( kurtosis(residuos) ))
print( 'skewness of normal distribution (should be 0): {}'.format( skew(residuos) ))
```
# Validación Cruzada
```
import sklearn
from sklearn.model_selection import cross_validate
scores_RL = cross_validate(LinearRegression(), X, y,scoring='neg_mean_absolute_percentage_error', cv=32)
scores_Las = cross_validate(Lasso(alpha=1.0), X, y,scoring='neg_mean_absolute_percentage_error', cv=32)
scores_Ri = cross_validate(Ridge(alpha=1.0), X, y,scoring='neg_mean_absolute_percentage_error', cv=32)
scores_reg_lin = scores_RL['test_score']*(-100)
scores_reg_lin = scores_reg_lin[scores_reg_lin < 40 ]
scores_reg_lass = scores_Las['test_score']*(-100)
scores_reg_lass = scores_reg_lass[scores_reg_lass < 40 ]
scores_reg_ridge = scores_Ri['test_score']*(-100)
scores_reg_ridge = scores_reg_ridge[scores_reg_ridge < 40 ]
print('Los resultados para cada modelo como rendiento promedio en MAPE son; Regresion Lineal: ', scores_reg_lin.mean(), ' Lasso: ',scores_reg_lass.mean(), ' Ridge: ',scores_reg_ridge.mean())
```
## Validacion de parametros
Haciendo uso del modelo OLS que dio el mejor rendimiento en la validacion cruzada procedemos a hallar el mejor parametro para box cox
```
from sklearn.model_selection import KFold
X_=np.array(X)
y_=np.array(y)
rendimiento=[]
params=np.linspace(0.1, 1.0, num=10)
for l in params:
rend=[]
kf = KFold(n_splits=10)
kf.get_n_splits(X_)
for train_index, val_index in kf.split(X_):
X_train, X_val = X_[train_index], X_[val_index]
y_train, y_val = y_[train_index], y_[val_index]
model = Lasso(alpha= l)
model.fit(X_train, y_train)
y_pred = model.predict(X_val)
rend.append(mape(y_val,y_pred))
rendimiento.append((l,np.mean(rend)))
rendimiento
rendimiento=[]
params=np.linspace(0.1, 10.0, num=10)
for l in params:
rend=[]
kf = KFold(n_splits=10)
kf.get_n_splits(X_)
for train_index, val_index in kf.split(X_):
X_train, X_val = X_[train_index], X_[val_index]
y_train, y_val = y_[train_index], y_[val_index]
model = Ridge(alpha= l)
model.fit(X_train, y_train)
y_pred = model.predict(X_val)
rend.append(mape(y_val,y_pred))
rendimiento.append((l,np.mean(rend)))
rendimiento
```
# Submit
```
test=datos_test.drop(['Unnamed: 0','Itinerary','Destination',
'Package Name','Places Covered',
'Travel Date','Hotel Details','Start City',
'Airline',
'Package Type','Cancellation Rules','Quarter','paquete',
'Meals','Sightseeing Places Covered','ano','mes','hotel_qual_prom','Rule 2','Rule 3',
'Rule 4','Rule 5','Rule 6','Rule 7',
'Rule 8','2021','Q1','Q2','Q3',' Gwalior fort ',
'Budget',' Delhi to Udaipur by Bus (Departure between 5 pm - 10 pm) ' ,' Dinner at Rambagh Palace - MMT ',' Udaipur to Delhi by Bus (Departure between 5 pm - 10 pm) '],axis=1)
test['dest_num_square']=test['dest_numb']**2
test['total_noches_sqr']=(test['total_noches'])**(1/2)
test['noches_pond_hotel_log']=np.log(test['noches_pond_hotel'])
reg = LinearRegression().fit(X, box_cox(y,-0.1))
y_pred = reg.predict(test)
y_pred=inv_box_cox(y_pred,-0.1)
submit = pd.read_csv("sample_submission.csv")
submit['PPPrice']=y_pred
submit.set_index('Index',inplace = True)
submit
submit.to_csv('sample_submission_total_final.csv')
```
| github_jupyter |
```
import os
import sys
module_path = os.path.abspath(os.path.join('..'))
if module_path not in sys.path:
sys.path.append(module_path)
import ipywidgets as widgets
import ipython_blocking
import ipywidgets as widgets
import cv2
from IPython.display import display, Image
from ipywidgets import interact, interactive, fixed, interact_manual
import PIL.Image
import io
import os
import sys
from IPython.display import display
module_path = os.path.abspath(os.path.join('..'))
if module_path not in sys.path:
sys.path.append(module_path)
button = widgets.ToggleButton(value=False,description="Click Me!")
# output = widgets.Output()
def on_button_clicked(b):
if button.value:
button.description = "Hey"
else:
button.description = "Click Me!"
# with output:
# print("Button clicked.")
button.observe(on_button_clicked)
display(button)
%blockrun button
#Test 1 of out-of-esper
def img_to_widget(img):
# height, width, _ = img.shape
f = io.BytesIO()
PIL.Image.fromarray(img).save(f, 'png')
return widgets.Image(value=f.getvalue(), height=height,
width=width)
def query_faces(ids):
faces = Face.objects.filter(id__in=ids)
return faces.values(
'id', 'bbox_y1', 'bbox_y2', 'bbox_x1', 'bbox_x2',
'frame__number', 'frame__video__id', 'frame__video__fps',
'shot__min_frame', 'shot__max_frame')
def get_img_checkbox():
img_checkbox = widgets.ToggleButton(
layout=widgets.Layout(width='auto'),
value=False,
description='hey',
disabled=False,
button_style='',
icon=''
)
def on_toggle(b):
if img_checkbox.value:
img_checkbox.button_style = 'danger'
img_checkbox.icon = 'check'
else:
img_checkbox.button_style = ''
img_checkbox.icon = ''
img_checkbox.observe(on_toggle, names='value')
return img_checkbox
def display_fancy_img():
PATH = "/Users/qlc23/Desktop/pic1.jpeg"
URL = "https://66.media.tumblr.com/6da6e8d2402a45357a6659a8122a004d/tumblr_p3a90iN5dk1u5hnjwo2_500.jpg"
# image = Image(url=URL)
file = open("ove_cat.jpg", "rb")
image = file.read()
img = widgets.Image(
value=image,
format='jpg',
width=300,
height=300,
)
inter = widgets.ToggleButton(
layout=widgets.Layout(width='auto'),
value=False,
description='hey',
disabled=False,
button_style='',
icon=''
)
image_checkbox = get_img_checkbox()
w = widgets.VBox([img, image_checkbox])
display(w)
# display(img)
# display(inter)
display_fancy_img()
def img_to_widget(img):
height, width, _ = img.shape
f = io.BytesIO()
PIL.Image.fromarray(img).save(f, 'png')
return widgets.Image(value=f.getvalue(), height=height,
width=width)
def _get_img_checkbox():
img_checkbox = widgets.ToggleButton(
layout=widgets.Layout(width='auto'),
value=False,
description='',
disabled=False,
button_style='',
icon=''
)
def on_toggle(b):
if img_checkbox.value:
img_checkbox.button_style = 'danger'
img_checkbox.icon = 'check'
else:
img_checkbox.button_style = ''
img_checkbox.icon = ''
img_checkbox.observe(on_toggle, names='value')
return img_checkbox
def _show_clusters(cluster_ids):
checkboxes = []
vboxes = []
for cluster_id in sorted(cluster_ids):
print('Cluster {} ({} faces)'.format(cluster_id, len(clusters[cluster_id])))
img_widget = img_to_widget(
cv2.cvtColor(img_crop, cv2.COLOR_BGR2RGB))
img_checkbox = get_img_checkbox()
checkboxes.append(img_checkbox)
vboxes.append(widgets.VBox([img_widget, img_checkbox]))
for cluster_id in sorted(cluster_ids):
imshow(cluster_images[cluster_id])
plt.show()
display(_get_img_checkbox)
images_per_row = 8
for i in range(0, len(vboxes), images_per_row):
display(widgets.HBox(vboxes[i:i + images_per_row]))
def _delete_clusters_display():#cluster_ids, discarded_clusters):
arr = [1]
discarded_clusters= set()
delete_button = widgets.Button(description='Delete', button_style='success')
for cluster_id in range(1,5): #sorted(cluster_ids):
# print('Cluster {} ({} faces)'.format(cluster_id, len(clusters[cluster_id])))
# imshow(cluster_images[cluster_id])
# plt.show()
output = widgets.Output()
display(delete_button, output)
def on_delete(b):
cluster_to_discard = 1 #int(b.description.split(' ')[1])
arr.append('Hello')
print(arr)
discarded_clusters.add(cluster_to_discard)
print(discarded_clusters)
# remaining_clusters = _get_remaining_clusters(meta_clusters) - discarded_clusters
# clear_output()
# _show_clusters(remaining_clusters)
delete_button.on_click(on_delete)
_delete_clusters_display()
a = widgets.IntSlider(description='a')
b = widgets.IntSlider(description='b')
c = widgets.IntSlider(description='c')
def f(a, b, c):
print('{}*{}*{}={}'.format(a, b, c, a*b*c))
out = widgets.interactive_output(f, {'a': a, 'b': b, 'c': c})
widgets.HBox([widgets.VBox([a, b, c]), out])
widgets.IntSlider(
value=7,
min=0,
max=10,
step=1,
description='Test:',
disabled=False,
continuous_update=False,
orientation='horizontal',
readout=True,
readout_format='d'
)
box = widgets.Checkbox(False, description='checker')
out = widgets.Output()
@out.capture()
def changed(b):
arr.append(1)
print(arr)
arr = []
box.observe(changed)
display(box)
display(out)
w = widgets.ToggleButton(
value=False,
description='Click me',
disabled=False,
button_style='', # 'success', 'info', 'warning', 'danger' or ''
tooltip='Description',
icon='check'
)
display(w)
out = widgets.Output(layout={'border': '1px solid black'})
with out:
for i in range(10):
print(i, 'Hello world!')
out
a = widgets.IntSlider(description='a')
b = widgets.IntSlider(description='b')
c = widgets.IntSlider(description='c')
def f(a, b, c):
print('{}*{}*{}={}'.format(a, b, c, a*b*c))
out = widgets.interactive_output(f, {'a': a, 'b': b, 'c': c})
widgets.HBox([widgets.VBox([a, b, c]), out])
debug_view = widgets.Output(layout={'border': '1px solid black'})
@debug_view.capture(clear_output=True)
def bad_callback(event):
print('This is about to explode')
return 1.0 / 0.0
button = widgets.Button(
description='click me to raise an exception',
layout={'width': '300px'}
)
button.on_click(bad_callback)
button
```
| github_jupyter |
```
# Hakan Can İpek, 528211011, Dec 6, 2021
from csv import reader
from collections import Counter
def problem1(path):
manufacturerSet = set()
soldCarsIn2010 = dict()
with open(path, 'r', newline='') as read_obj:
csv_reader = reader(read_obj, delimiter = ',')
header = next(csv_reader)
# Pass reader object to list() to get a list of lists
list_of_rows = list(csv_reader)
for x in list_of_rows:
if (x[2] == '\xa0Mercedes-Benz '):
x[2] = 'Mercedes-Benz '
manufacturerSet.add(x[2])
if (x[0] == '2010'):
if (x[2] in soldCarsIn2010):
val = soldCarsIn2010[x[2]]
val += int(x[4])
soldCarsIn2010[x[2]] = val
else:
soldCarsIn2010[x[2]] = int(x[4])
print("the number of unique manufacturers in this dataset is :", len(manufacturerSet))
k = Counter(soldCarsIn2010)
high = k.most_common(1)
print('\"', high[0][0],'\" has the highest sales in 2010, which is ', high[0][1], sep="")
problem1("C:\\Users\\ipekh\\Desktop\\Big Data Homeworks\\Advances in Data Science\\midterm_data\\norway_new_car_sales_by_model.csv")
import os
import pathlib
import timeit
def problem3(path):
start = timeit.default_timer()
totalModifyCount = 0
for root, dirs, files in os.walk(path):
for x in files:
from re import search
rx = search(".pdbqt", x)
if rx:
nx = root + "\\" + x
newFileName = ""
from csv import reader
with open(nx, 'r', newline='') as read_obj:
csv_reader = reader(read_obj)
list_of_rows = list(csv_reader)
name = list_of_rows[0][0]
newFileName = name[15:]
newFileName += ".pdbqt"
os.rename(nx, root + "\\" + newFileName)
totalModifyCount += 1
stop = timeit.default_timer()
print('the number of processed files:', totalModifyCount)
print('estimated hours to process three million files:', stop - start)
problem3("C:\\Users\\ipekh\\Desktop\\Big Data Homeworks\\Advances in Data Science\\midterm_data\\mol_files")
class Rating:
def __init__(this, _list):
this.id = _list[0]
this.movieId = _list[2]
this.stars = _list[4]
this.epoch = _list[6]
def getData(this):
return '{0}::{1}::{2}::{3}'.format(this.id, this.movieId, this.stars, this.epoch)
class User:
def __init__(this, _list):
this.id = _list[0]
this.gender = _list[2]
this.age = _list[4]
this.occ = _list[6]
this.zip = _list[8]
def getData(this):
return '{0}::{1}::{2}::{3}'.format(this.gender, this.age, this.occ, this.zip)
class Movie:
def __init__(this, _list):
this.id = _list[0]
this.title = _list[2][:-7]
this.genres = _list[4]
this.year = _list[2][-5:-1]
def getData(this):
return '{0}::{1}::{2}'.format(this.title, this.genres, this.year)
import timeit
from csv import reader
def problem4(moviePath, userPath, ratingPath, mergedPath):
start = timeit.default_timer()
movies = dict()
users = dict()
ratings = dict()
with open(ratingPath, 'r', newline='') as read_obj:
csv_reader = reader(read_obj, delimiter = ':')
list_of_rows = list(csv_reader)
for x in list_of_rows:
rate = Rating(x)
if rate.id in ratings:
_l = ratings[rate.id]
_l.append(rate)
else: _l = [rate]
ratings[rate.id] = _l
with open(userPath, 'r', newline='') as read_obj:
csv_reader = reader(read_obj, delimiter = ':')
list_of_rows = list(csv_reader)
for x in list_of_rows:
user = User(x)
users[user.id] = user
with open(moviePath, 'r', newline='') as read_obj:
csv_reader = reader(read_obj, delimiter = ':')
list_of_rows = list(csv_reader)
for x in list_of_rows:
movie = Movie(x)
movies[movie.id] = movie
mergedList = list()
for userId, user in users.items():
for rate in ratings[userId]:
merged = rate.getData() + "::" + user.getData() + "::" + movies[rate.movieId].getData()
mergedList.append(merged)
stop = timeit.default_timer()
print('time before write:', stop - start)
with open(mergedPath, 'w', newline = '') as mergeFile:
for x in mergedList:
mergeFile.write(x)
mergeFile.write('\n')
stop = timeit.default_timer()
print('time after write:', stop - start)
problem4("C:\\Users\\ipekh\\Desktop\\Big Data Homeworks\\Advances in Data Science\\midterm_data\\ml-1m\\movies.dat", "C:\\Users\\ipekh\\Desktop\\Big Data Homeworks\\Advances in Data Science\\midterm_data\\ml-1m\\users.dat", "C:\\Users\\ipekh\\Desktop\\Big Data Homeworks\\Advances in Data Science\\midterm_data\\ml-1m\\ratings.dat", "merged.dat")
problem4():
import pandas as pd
movies=pd.read_table(r"C:\Users\ipekh\Desktop\Big Data Homeworks\Advances in Data Science\midterm_data\ml-1m\movies.dat", sep="::", header=None, names=["MovieID", "Title", "Genres"], engine='python')
ratings=pd.read_table(r"C:\Users\ipekh\Desktop\Big Data Homeworks\Advances in Data Science\midterm_data\ml-1m\ratings.dat", sep="::", header=None, engine='python', names=["UserID", "MovieID", "Rating", "Timestamp"])
users=pd.read_table(r"C:\Users\ipekh\Desktop\Big Data Homeworks\Advances in Data Science\midterm_data\ml-1m\users.dat", sep="::", header=None, engine='python', names=["UserID", "Gender", "Age", "Occupation", "Zip"])
movies["year"] = movies["Title"].str[-5:-1]
movies["title"] = movies["Title"].str[:-7]
movies=movies.drop(["Title"], axis=1)
ratings["Timestamp"] = pd.to_datetime(ratings["Timestamp"], unit='s')
merge1= pd.merge(movies, ratings ,how="left", on=["MovieID", "MovieID"])
finaltable= pd.merge(merge1, users ,how="left", on=["UserID", "UserID"])
x = finaltable.to_string(header=False,
index=False,
index_names=False).split(sep="\n")
text = ['::'.join(ele.split()) for ele in x];text
problem4()
```
| github_jupyter |
### [Oregon Curriculum Network](http://www.4dsolutions.net/ocn) <br />
[Discovering Math with Python](Introduction.ipynb)
# Chapter 11: QUATERNIONS
Quaternions were invented by Sir William Rowan Hamilton around 1843 and were considered a breakthrough.
In subsequent chapters Willard Gibbs and Oliver Heaviside came up with a vector concept that proved easier to use for many of the physics applications for which quaternions had originally been proposed.
However quaternions and vectors together have become chief tools in accomplishing rotation, in computer graphics and games, robotics, rocketry. We may have entered the realm of that proverbially most difficult of disciplines: rocket science.
Quaternions have some advantages over rotation matrices. It's easier to slice up a rotation in a process called [SLERP](https://en.wikipedia.org/wiki/Slerp).
You might think of quaternions as "vectors on steroids" if that helps. They have some properties in common with vectors, and are conceived to have a "vector part" somewhat like complex numbers have a "real part". However, like complex numbers, they're considered numbers in their own right, actually a superset of the complex, which in turn contain the reals and so on down to N, the counting numbers (natural numbers).
Unit quaternions (w, x, y, z) with w\*\*2 + x\*\*2 + y\*\*2 + z\*\*2 == 1, form a group under multiplication. Any two unit quaternions, when multiplied, produce a unit quaternion (Closure), and every unit quaternion has an inverse (w, -x, -y, -z), such that q \* q\*\*-1 gives the unit quaternion (1, 0, 0, 0).
Indeed the elements i, j, k from which quaternions are made, abet the i of complex number fame, with two more 2nd roots of -1. All three, and their three inverses, engage in a kind of dance.
Group elements are: {i, j, k, 1, -1, -i, -j, -k}. Every product of two of these elements, is in this set (closure); associativity holds; every element has an inverse such that the two give a product of 1, and 1 serves as the neutral (identity) element.
<a data-flickr-embed="true" href="https://www.flickr.com/photos/kirbyurner/36363327133/in/dateposted-public/" title="Homework Assignment"><img src="https://farm5.staticflickr.com/4353/36363327133_1962435626.jpg" width="500" height="375" alt="Homework Assignment"></a><script async src="//embedr.flickr.com/assets/client-code.js" charset="utf-8"></script>
Just the identities we've been given: i\*\*2 = j\*\*2 = k\*\*2 = i \* j \* k = -1 are suffient to derive the above Cayley Table. This table, in turn enables use to flesh out the \_\_mul\_\_ method, by taking sixteen products (w, x, y, z times each w', x', y', z') and substituting for products such as -i \* k and k \* j. Some of the derivations for are table are shown above.
What we'll plan to do, when rotating point vectors of polyhedron P, by n degrees around unit rotation vector q, is initialize the corresponding unit quaternion rQ using function rotation(n, q) and then multiply every point vector of P in a "sandwich" between rQ and its inverse:
new_Pv = rQ \* Pv \* ~rQ (how we apply rotations to each position vector Pv).
Remember our Polyhedron objects, as data structures, start with a set of faces pegged to vertexes, expressed as a dict of position Vectors. We used both xyz and quadray notation to map the same set of points.
Below is the source code for doing that. Understanding multiplication involves developing the multiplication table for i, j, k which play the role of basis vectors in this language game. i\*\*2 == j\*\*2 == k\*\*2 == -1, and i \* j \* k == -1 as well.
```
from math import cos, sin, radians, pi
from qrays import Vector
import unittest
class Quaternion:
def __init__(self, w, x, y, z):
"""
w is the scalar part;
x,y,z comprise a vector part as
the coefficients of i,j,k respectively
where i,j,k are the three basis vectors
described by Hamilton such that
i**2 == j**2 == k**2 == -1, and
i * j * k == -1 as well.
"""
self.w = w
self.x = x
self.y = y
self.z = z
def __mul__(self, other):
"""
Derived by inter-multiplying all four terms in
each Quaternion to get 16 products and then
simplifying according to such rules as ij = k,
ji = -k.
See: https://youtu.be/jlskQDR8-bY Mathoma
'Quaternions explained briefly'
"""
a, b, c, d = self.w, self.x, self.y, self.z
e, f, g, h = other.w, other.x, other.y, other.z
w = (a*e - b*f - c*g - d*h)
x = (a*f + b*e + c*h - d*g)
y = (a*g - b*h + c*e + d*f)
z = (a*h + b*g - c*f + d*e)
return Quaternion(w, x, y, z)
def __invert__(self):
return Quaternion( self.w,
-self.x,
-self.y,
-self.z)
def vector(self):
return Vector((self.x, self.y, self.z))
def __eq__(self, other):
tolerance = 1e-8
return (abs(self.w - other.w) < tolerance and
abs(self.x - other.x) < tolerance and
abs(self.y - other.y) < tolerance and
abs(self.z - other.z) < tolerance)
def __repr__(self):
return "Quaternion({},{},{},{})".format(self.w,
self.x,
self.y,
self.z)
def rotator(uV, α):
w = cos(α/2)
x = uV.x * sin(α/2)
y = uV.y * sin(α/2)
z = uV.z * sin(α/2)
return Quaternion(w, x, y, z)
class TestQuaternion(unittest.TestCase):
def test_inverse(self):
Q = Quaternion(0, 0, -1, 0)
inverse = ~Q
self.assertEqual(Q * inverse, Quaternion(1,0,0,0))
def test_x(self):
rV = Quaternion(0,1,0,0)
rQ = rotator(Vector((0,1,0)), pi)
newQ = rQ * rV * ~rQ
self.assertTrue(newQ.vector() == Vector((-1, 0, 0)))
def test_360(self):
rV = Quaternion(0,1,0,0)
one_deg = radians(1)
rQ = rotator(Vector((0,1,0)), one_deg)
for _ in range(360):
rV = rQ * rV * ~rQ
self.assertTrue(rV.vector() == Vector((1, 0, 0)))
a = TestQuaternion()
# unittest.TextTestRunner().run(a)
suite = unittest.TestLoader().loadTestsFromModule(a)
unittest.TextTestRunner().run(suite)
```
Back to Chapter 10: [Complex Numbers](Complex%20Numbers.ipynb) <br />
Continue to Chapter 12: [The Mandelbrot Set](Mandelbrot%20Set.ipynb)<br />
[Introduction / Table of Contents](Introduction.ipynb)
| github_jupyter |
## Data Visualization with Bokeh
## About Me
## Talk Structure
* Basic Plotting & Styling
* Layouts
* Linking Multiple Grids
* Data Shader
* Embedding into Django
* Running with Bokeh Server
* CustomJS
```
import numpy as np
from bokeh.io import output_notebook, show
from bokeh.plotting import figure
from bokeh.models import Diamond
output_notebook()
x, y = [1, 2, 3, 4, 5], [6, 7, 2, 4, 5]
# My First Scatter Plot:
# create a new plot with default tools, using figure
p = figure(height=400)
# add a circle renderer with x and y coordinates
p.circle(x, y)
# radius=0.4 - diameter to graph scale
# size=45
# size=[10, 20, 40, 80] # 'vectorize' sizes:
show(p) # show the results
p = figure(plot_height=200, sizing_mode="scale_width")
p.title.text = 'First Plot'
p.background_fill_color = 'beige'
p.outline_line_width = 7
p.outline_line_alpha = 0.3
p.outline_line_color = "navy"
r = p.square(x, y, color="firebrick")
r.glyph.size = 50
r.glyph.fill_alpha = 0.2
r.glyph.line_color = "firebrick"
r.glyph.line_dash = [5, 1]
r.glyph.line_width = 2
show(p) # show the results
```
## Modify Axis Styling
```
from math import pi
from datetime import date
p = figure(plot_height=200, sizing_mode="scale_width", tools='', x_axis_type='datetime')
dates = [date(2018, 1, 1), date(2018, 2, 1), date(2018, 3, 1), date(2018, 4, 1), date(2018, 5, 1)]
p.square(dates, y, size=50)
# change just some things about the x-axes
p.xaxis.axis_label = "Dates"
p.xaxis.axis_line_width = 3
p.xaxis.axis_line_color = "red"
p.xaxis.axis_label_text_font_size = "14pt"
p.xaxis.major_label_orientation = pi / 3
p.xaxis.major_label_text_font_size = "14pt"
# change just some things about the y-axes
p.yaxis.axis_label = "Pressure"
p.yaxis.major_label_text_color = "orange"
p.yaxis.major_label_orientation = "vertical"
p.yaxis.axis_line_color = "#00ff00"
# change things on all axes
p.axis.minor_tick_in = -3
p.axis.minor_tick_out = 6
show(p) # show the results
```
## Selecting Glyphs
```
p = figure(plot_height=200, sizing_mode="scale_width", tools='tap', title='Select a Square')
renderer = p.circle(x, y,
size=75,
# radius=0.5, # GOT-CHA - Hit testing doesn't quite work with radius on Circles
# set visual properties for selected glyphs
selection_color="firebrick",
# set visual properties for non-selected glyphs
nonselection_fill_alpha=0.2,
nonselection_fill_color="grey",
nonselection_line_color="firebrick",
nonselection_line_alpha=1.0)
show(p)
# Hold Shift / Ctrl for multi-select
```
## Glyph Types:
https://bokeh.pydata.org/en/latest/docs/reference/models/markers.html
<img src="images/bokeh_models_markers.png" width="200"/>
## Hovering Over Data
```
from bokeh.plotting import ColumnDataSource
source = ColumnDataSource(data=dict(
x=x,
y=y,
desc=['A', 'b', 'C', 'd', 'E'],
))
TOOLTIPS = [
("index", "$index"),
("(x,y)", "($x, $y{0})"),
("desc", "@desc"),
]
p = figure(plot_width=400, plot_height=400, tooltips=TOOLTIPS,
title="Mouse over the dots")
p.circle('x', 'y', size=40, source=source)
show(p)
from bokeh.models.tools import HoverTool
from bokeh.sampledata.glucose import data
subset = data.loc['2010-10-06']
x, y = subset.index.to_series(), subset['glucose']
# Basic plot setup
p = figure(width=600, height=300, x_axis_type="datetime", title='Hover over points')
p.line(x, y, line_dash="4 4", line_width=1, color='gray')
cr = p.circle(x, y, size=20,
fill_color="grey", hover_fill_color="firebrick",
fill_alpha=0.05, hover_alpha=0.3,
line_color=None, hover_line_color="white")
p.add_tools(HoverTool(tooltips=None, renderers=[cr], mode='hline'))
show(p)
from bokeh.sampledata.stocks import AAPL
from bokeh.models import ColumnDataSource
tmp = AAPL
tmp['adj close'] = AAPL['adj_close']
tmp['date'] = np.array(AAPL['date'], dtype=np.datetime64) # convert date strings to real datetimes
p = figure(x_axis_type="datetime", title="AAPL", plot_height=250, sizing_mode='scale_width')
p.xgrid.grid_line_color=None
p.ygrid.grid_line_alpha=0.5
p.xaxis.axis_label = 'Time'
p.yaxis.axis_label = 'Value'
p.line('date', 'adj close', source=ColumnDataSource(data=tmp), line_dash="dashed", line_color='grey')
ht = HoverTool(
tooltips=[
( 'date', '@date{%F}' ),
( 'close', '$@{adj close}{%0.2f}' ), # use @{ } for field names with spaces
( 'volume', '@volume{0.00 a}' ),
],
formatters={
'date' : 'datetime', # use 'datetime' formatter for 'date' field
'adj close' : 'printf', # use 'printf' formatter for 'adj close' field
# use default 'numeral' formatter for other fields
},
# display a tooltip whenever the cursor is vertically in line with a glyph
mode='vline'
)
p.add_tools(ht)
show(p)
```
## Annotations
```
from bokeh.models.annotations import Arrow
from bokeh.models.arrow_heads import OpenHead, NormalHead, VeeHead # Arrow head Types
from bokeh.sampledata.stocks import AAPL
from bokeh.models import ColumnDataSource
from bokeh.models.annotations import BoxAnnotation
tmp = AAPL
tmp['adj close'] = AAPL['adj_close']
tmp['date'] = np.array(AAPL['date'], dtype=np.datetime64) # convert date strings to real datetimes
p = figure(x_axis_type="datetime", title="AAPL", plot_height=450, sizing_mode='scale_width')
p.xgrid.grid_line_color=None
p.ygrid.grid_line_alpha=0.5
p.xaxis.axis_label = 'Time'
p.yaxis.axis_label = 'Value'
p.line('date', 'adj close', source=ColumnDataSource(data=tmp), line_dash="dashed", line_color='grey')
# x_start and _end need to be np.datetime64 data types to match the underlying data source
p.add_layout(Arrow(end=NormalHead(fill_color="red"),
x_start=np.datetime64('2000-11-14'), y_start=20, x_end=np.datetime64('2012-08-01'), y_end=680))
# region that always fills the top of the plot
upper = BoxAnnotation(bottom=600, fill_alpha=0.1, fill_color='olive')
p.add_layout(upper)
# region that always fills the bottom of the plot
lower = BoxAnnotation(top=200, fill_alpha=0.1, fill_color='black')
p.add_layout(lower)
# a finite region
center = BoxAnnotation(top=500, bottom=300, left=np.datetime64('2012-01-01'), right=np.datetime64('2014-01-01'),
fill_alpha=0.1, fill_color='navy')
p.add_layout(center)
show(p)
# Can Also add lines, circles, elipsis, or other polygons
```
## Linking Graphs
```
from bokeh.layouts import gridplot
x = list(range(11))
y0, y1, y2 = x, [10-i for i in x], [abs(i-5) for i in x]
plot_options = dict(width=250, plot_height=250, tools='pan,wheel_zoom,box_select,reset')
# create a new plot
s1 = figure(**plot_options)
s1.circle(x, y0, size=10, color="navy")
# create a new plot and share both ranges
s2 = figure(x_range=s1.x_range, y_range=s1.y_range, **plot_options)
s2.triangle(x, y1, size=10, color="firebrick")
# create a new plot and share only one range
s3 = figure(x_range=s1.x_range, **plot_options)
s3.square(x, y2, size=10, color="olive")
p = gridplot([[s1, s2, s3]])
# show the results
show(p)
from bokeh.models import ColumnDataSource
x = list(range(-20, 21))
y0, y1 = [abs(xx) for xx in x], [xx**2 for xx in x]
# create a column data source for the plots to share
source = ColumnDataSource(data=dict(x=x, y0=y0, y1=y1))
TOOLS = "wheel_zoom,box_select,lasso_select,reset"
# create a new plot and add a renderer
left = figure(tools=TOOLS, width=300, height=300)
left.circle('x', 'y0', source=source)
# create another new plot and add a renderer
right = figure(tools=TOOLS, width=300, height=300)
right.circle('x', 'y1', source=source)
p = gridplot([[left, right]])
show(p)
```
## Bar Charts
```
from bokeh.models import ColumnDataSource
from bokeh.palettes import Spectral6
fruits = ['Apples', 'Pears', 'Nectarines', 'Plums', 'Grapes', 'Strawberries']
counts = [5, 3, 4, 2, 4, 6]
source = ColumnDataSource(data=dict(fruits=fruits, counts=counts, color=Spectral6))
p = figure(x_range=fruits, plot_height=250, y_range=(0, 9), title="Fruit Counts")
p.vbar(x='fruits', top='counts', width=0.9, color='color', legend="fruits", source=source)
p.xgrid.grid_line_color = None
p.legend.orientation = "horizontal"
p.legend.location = "top_center"
show(p)
```
## Stacked Bar Chart
```
from bokeh.palettes import GnBu3, OrRd3
years = ['2015', '2016', '2017']
exports = {'fruits' : fruits,
'2015' : [2, 1, 4, 3, 2, 4],
'2016' : [5, 3, 4, 2, 4, 6],
'2017' : [3, 2, 4, 4, 5, 3]}
imports = {'fruits' : fruits,
'2015' : [-1, 0, -1, -3, -2, -1],
'2016' : [-2, -1, -3, -1, -2, -2],
'2017' : [-1, -2, -1, 0, -2, -2]}
p = figure(y_range=fruits, plot_height=250, x_range=(-16, 16), title="Fruit import/export, by year")
p.hbar_stack(years, y='fruits', height=0.9, color=GnBu3, source=ColumnDataSource(exports),
legend=["%s exports" % x for x in years])
p.hbar_stack(years, y='fruits', height=0.9, color=OrRd3, source=ColumnDataSource(imports),
legend=["%s imports" % x for x in years])
p.y_range.range_padding = 0.1
p.ygrid.grid_line_color = None
p.legend.location = "center_left"
show(p)
```
## Grouped Bar Chart
```
from bokeh.models import FactorRange
from bokeh.transform import factor_cmap
fruits = ['Apples', 'Pears', 'Nectarines', 'Plums', 'Grapes', 'Strawberries']
years = ['2015', '2016', '2017']
data = {'fruits' : fruits,
'2015' : [2, 1, 4, 3, 2, 4],
'2016' : [5, 3, 3, 2, 4, 6],
'2017' : [3, 2, 4, 4, 5, 3]}
# this creates [ ("Apples", "2015"), ("Apples", "2016"), ("Apples", "2017"), ("Pears", "2015), ... ]
x = [ (fruit, year) for fruit in fruits for year in years ]
counts = sum(zip(data['2015'], data['2016'], data['2017']), ()) # like an hstack
source = ColumnDataSource(data=dict(x=x, counts=counts))
p = figure(x_range=FactorRange(*x), plot_height=250, title="Fruit Counts by Year")
p.vbar(x='x', top='counts', width=0.9, source=source, line_color="white",
# use the palette to colormap based on the the x[1:2] values
fill_color=factor_cmap('x', palette=['firebrick', 'olive', 'navy'], factors=years, start=1, end=2))
p.y_range.start = 0
p.x_range.range_padding = 0.1
p.xaxis.major_label_orientation = 1
p.xgrid.grid_line_color = None
show(p)
```
## Bar Chart Jitter
```
from bokeh.sampledata.commits import data
# Github Commits
data.head()
from bokeh.transform import jitter
DAYS = ['Sun', 'Sat', 'Fri', 'Thu', 'Wed', 'Tue', 'Mon']
source = ColumnDataSource(data)
p = figure(plot_width=800, plot_height=300, y_range=DAYS, x_axis_type='datetime',
title="Commits by Time of Day (US/Central) 2012—2016")
p.circle(x='time', y='day', source=source, alpha=0.3)
# jitter('day', width=0.6, range=p.y_range)
p.xaxis[0].formatter.days = ['%Hh']
p.x_range.range_padding = 0
p.ygrid.grid_line_color = None
show(p)
```
## Datashader
```
import pandas as pd
import numpy as np
import datashader as ds
import datashader.transfer_functions as tf
np.random.seed(1)
num = 10000
dists = {
cat: pd.DataFrame(
dict(
x=np.random.normal(x,s,num),
y=np.random.normal(y,s,num),
val=val,
cat=cat,
color=color))
for x,y,s,val,cat,color in [
(2,2, 0.01,10,'d1', 'red'), (2,-2,0.1,20,'d2', 'blue'),
(-2,-2,0.5,30,'d3', 'green'), (-2,2,1.0,40,'d4', 'purple'), (0,0,3,50,'d5', 'orange')
]
}
df = pd.concat(dists, ignore_index=True)
df["cat"] = df["cat"].astype("category")
df.tail()
df.head()
```
## Normal Bokeh View - Very Sluggish Response
```
p = figure(x_range=(-5,5), y_range=(-5,5))
data_source = ColumnDataSource(data=df)
p.circle(x='x', y='y', source=data_source, fill_alpha=0.001, color='color')
show(p)
```
## Datashader callback - real-time response, same data
```
import bokeh.plotting as bp
from datashader.bokeh_ext import InteractiveImage
bp.output_notebook()
p = bp.figure(tools='pan,wheel_zoom,reset,box_zoom', x_range=(-5,5), y_range=(-5,5))
def image_callback(x_range, y_range, w, h):
canvas = ds.Canvas(plot_width=w, plot_height=h, x_range=x_range, y_range=y_range)
agg = canvas.points(df, 'x', 'y', ds.count_cat('cat'))
img = tf.shade(agg) #, color_key)
# 0 - 1 for threshold
return tf.dynspread(img, threshold=1)
InteractiveImage(p, image_callback)
```
## Embedding into Django
Run server from the `django_proj` folder with `python manage.py runserver`
## Bonus - CustomJS (If time / requested)
```
# Exercise: Create a plot that updates based on a Select widget
from bokeh.layouts import column, row, gridplot
from bokeh.models import CustomJS, ColumnDataSource, Select
x = [x*0.005 for x in range(0, 201)]
source = ColumnDataSource(data=dict(x=x, y=x))
plot = figure(plot_width=400, plot_height=400)
plot.line('x', 'y', source=source, line_width=3, line_alpha=0.6)
# (title="Option:", value="one", options=["one", 'two', 'three', 'four'])
select = Select(title="Power", value='0.005', options=['0.005', '0.1005', '0.2005', '0.3005'])
update_curve = CustomJS(args=dict(source=source, select=select), code="""
var data = source.data;
var f = select.value;
x = data['x']
y = data['y']
for (i = 0; i < x.length; i++) {
y[i] = Math.pow(x[i], f)
}
console.log("hello world")
// necessary becasue we mutated source.data in-place
source.change.emit();
""")
select.js_on_change('value', update_curve)
show(gridplot([[select, plot]]))
```
| github_jupyter |
Name: Srinivas Jakkula
I have provided explanation of each cell in the form or comments or some kind of description in the above or below cells for all the code
Initial Objectives give as per project proposal.
NOTE: All objectives are met in this project and work items are in different order compthis scope
I want to showcase the daily covid-19 vaccination progress across all countries for the data available in a plot for easy understanding of trends for all countries using line plots. --- Done
I want to provide a chart with the details of total vaccinations done for each country for all the data available and also provide details about how many people in each country are vaccinated per million to see how fast the vaccination is progressing across countries ---Done
I want to find if all the vaccinations available are used on the same day by comparing vaccination available vs people vaccinated using fields “total_vaccinations” and “people_vaccinated” per day. ---- Done
I want to find what vaccines are used in each country and also want to find which vaccine is more distributed across the world to see the production capability of the companies for these vaccines. ---Done
```
# Importing the required modules for COVID-19 vaccination Data
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
```
Reading the data file and creating dataframe
```
# dataset downloaded from kaggle (link for the dataset is: https://www.kaggle.com/gpreda/covid-world-vaccination-progress)
# Reading the dataset to a data frame
vaccine_df = pd.read_csv("country_vaccinations.csv")
# Displaying some basic information about the data frame like, shape, size, head and tail
# And also describing the data frame
print(vaccine_df.shape)
print(vaccine_df.size)
print(vaccine_df.head())
print(vaccine_df.tail())
vaccine_df.describe()
# Displaying some information of the data frame
vaccine_df.info()
# Printing column names in the data frame
print(vaccine_df.columns)
# Printing count of columns
print(len(vaccine_df.columns))
# Printing the unique country names
CountryList = vaccine_df['country'].unique()
print(CountryList)
print(f"Number of conutries: {len(CountryList)}")
#Check for missing data from each column
vaccine_df.isna().sum()
```
Based on above data, we see that there are some NaN values in the some of the columns.
```
# Finding th vacines used in each country
country_by_vaccine = pd.DataFrame(vaccine_df.groupby('country').vaccines.unique())
print(country_by_vaccine)
# Finding the unique columns
# Finding the number of unique values in each column.
vaccine_df.nunique()
#Dropping columns that are not required, here source_name and source_website
vaccine_df.drop(['source_name', 'source_website'], axis = 1, inplace = True)
vaccine_df.head()
# Examine the names and the count of each country.
vaccine_df['country'].value_counts()
# Based on this data, it seems like UK data is given 5 time with names United Kingdom, Northern Ireland
# Scotland, England, and Wales
#In country column, England,Scotland,Northern Ireland and Wales are part of UK, let us drop rows except UK
index_names = vaccine_df[vaccine_df.country.isin(['England', 'Scotland', 'Wales', 'Northern Ireland'])].index
vaccine_df.drop(index_names, inplace = True)
print(vaccine_df.head())
vaccine_df.shape
# Examine the names and the count of each country.
vaccine_df['country'].value_counts()
# After droping above, showing the details number of unique values in column
vaccine_df.nunique()
# Find the maximum number of total vaccinations for each country and display them in descending order.
total_vaccine_per_country = vaccine_df.groupby(['country'])['total_vaccinations'].max().reset_index()
total_vaccine_per_country_df2 = total_vaccine_per_country.sort_values(by='total_vaccinations', ascending = False, ignore_index = True)
total_vaccine_per_country_df2
plt.figure(figsize=(18,8))
plt.bar(np.arange(len(total_vaccine_per_country_df2)), total_vaccine_per_country_df2["total_vaccinations"], align="center")
plt.xticks(np.arange(len(total_vaccine_per_country_df2)),total_vaccine_per_country_df2['country'])
plt.xticks(rotation=90)
plt.xlabel("Country",fontsize=12)
plt.ylabel("Total Vaccinations",fontsize=12)
plt.title("Total Vaccinations per country",fontsize=12)
# As I am printing all 150 countries data, X axis is not visible clearly.
# Due to this, I have drawn top 20 countries details in next plot.
top_20_countries = total_vaccine_per_country_df2.iloc[0:21]
print(top_20_countries.head())
plt.figure(figsize=(18,8))
plt.bar(np.arange(len(top_20_countries)), top_20_countries["total_vaccinations"], align="center", color="green")
plt.xticks(np.arange(len(top_20_countries)),top_20_countries['country'])
plt.xticks(rotation=90)
plt.xlabel("Country",fontsize=8)
plt.ylabel("Total Vaccinations",fontsize=8)
plt.title("Total Vaccinations per country",fontsize=12)
# View the number of people who have received the vaccine, either completely or partially.
# The data is sorted in descending order based on the number of people who are fully vaccinated.
people_vaccinated1 = vaccine_df.groupby(['country'])['people_vaccinated'].max().reset_index()
people_vaccinated_df2 = people_vaccinated1.sort_values(by='people_vaccinated', ascending = False, ignore_index = True).style.background_gradient(cmap = 'Blues')
people_vaccinated_df2
# new_df = people_vaccinated_df2.iloc[0:21]
# Finding the people_vaccinated for each country and display them in descending order.
people_vaccinated2 = vaccine_df.groupby(['country'])['people_vaccinated'].max().reset_index()
people_vaccinated2_df2 = people_vaccinated2.sort_values(by='people_vaccinated', ascending = False, ignore_index = True)
#people_vaccinated2_df2
new_df = people_vaccinated2_df2.iloc[:20]
print(new_df)
plt.figure(figsize=(18,8))
plt.bar(np.arange(len(new_df)), new_df["people_vaccinated"], align="center", color="yellow")
plt.xticks(np.arange(len(new_df)),new_df['country'])
plt.xticks(rotation=90)
plt.xlabel("Country",fontsize=8)
plt.ylabel("People Vaccinated",fontsize=8)
plt.title("People Vaccinated per country",fontsize=12)
# Which country is using what vaccine/s?
country_by_vaccine = pd.DataFrame(vaccine_df.groupby('country').vaccines.unique())
print(country_by_vaccine)
# Getting the details of total vaccinations available and people vaccinated details for all countries
print(total_vaccine_per_country_df2.head())
print(people_vaccinated2_df2.head())
merged_df = pd.merge(total_vaccine_per_country_df2, people_vaccinated2_df2, on = "country", how = "inner")
print(merged_df.head())
print(merged_df.shape)
# Plotting total vaccinations vs people vaccinated.
merged_20 = merged_df.iloc[:20]
merged_20.plot(x="country", y=["total_vaccinations", "people_vaccinated"], kind="bar",figsize=(9,8))
plt.show()
```
Based on above picture, we can see that in most countries, all vaccines are not being used. And also as there is no people_vaccined data for China, hence there is no data ploted for that specific country in the above plot.
```
# Daily vaccinations of countries to see the progress
# Slice the original dataset to get the 'date', country', and 'daily_vaccinations' in a single dataframe.
df9 = vaccine_df[["date", "country", "daily_vaccinations"]]
df9.head(10)
# Collecting only US data
df_us = df9.loc[df9['country'] == "United States"]
df_us
# Plotting only US data
plt.figure(figsize=(16,8))
plt.style.use('seaborn-whitegrid')
plt.plot(df_us.date,df_us[['daily_vaccinations']], color='green')
plt.xticks(rotation=90)
plt.xlabel("Date",fontsize=12)
plt.ylabel("Daily Vaccination",fontsize=12)
plt.title("US Daily Vaccination",fontsize=16)
plt.show()
# Plotting all countries daily vaccinations
# daily_vac_20_countries = df9.head(10)
CList = df9['country'].unique()
print(CList)
plt.figure(figsize=(16,8))
plt.style.use('seaborn-whitegrid')
for ele in CList:
c_data = df9.loc[df9['country'] == ele]
plt.plot(c_data.date,c_data[['daily_vaccinations']])
plt.xticks(rotation=90)
plt.xlabel("Date",fontsize=12)
plt.ylabel("Daily Vaccination",fontsize=12)
plt.title("All Countries Daily Vaccination",fontsize=16)
plt.legend(CList)
plt.show()
```
As we are plotting all 146 countries the legend is so lengthy.
Here I am not plotting for only few countries. I am using that logic for some other tasks that I am trying to solve in this project
```
# Finding the correlation
correlation = vaccine_df.corr()
print(correlation)
plt.pcolor(correlation)
plt.yticks(np.arange(0.5, len(correlation.index), 1), correlation.index)
plt.xticks(np.arange(0.5, len(correlation.columns), 1), correlation.columns)
plt.show()
# Heat map using seaborn heatmap
import seaborn as sns
sns.heatmap(correlation, annot=True)
```
##People fully vaccinated - top 5 countries
```
#People fully vaccinated - top 5 countries
fully_vaccinated = vaccine_df.groupby('iso_code', as_index = False).people_fully_vaccinated.max()
fully_vaccinated_sort_top5 = fully_vaccinated.sort_values(by = 'people_fully_vaccinated', ascending = False).head()
print(fully_vaccinated_sort_top5.shape)
print(fully_vaccinated_sort_top5)
# Ploting a bar chart of fully vacinated people from top 5 countries
plt.figure(figsize=(18,8))
plt.bar(np.arange(len(fully_vaccinated_sort_top5)), fully_vaccinated_sort_top5["people_fully_vaccinated"], align="center")
plt.xticks(np.arange(len(fully_vaccinated_sort_top5)),fully_vaccinated_sort_top5['iso_code'])
plt.xticks(rotation=90)
plt.xlabel("iso_code",fontsize=12)
plt.ylabel("people_fully_vaccinated",fontsize=12)
plt.title("total vaccinations done for top 5 country Details",fontsize=12)
plt.show()
fully_vaccinated_sort = fully_vaccinated.sort_values(by = 'people_fully_vaccinated', ascending = False)
print(fully_vaccinated_sort.shape)
print(fully_vaccinated_sort)
# Dropping NaN values from the fully vaccinated data frame
fully_vaccinated_sort = fully_vaccinated_sort.dropna()
fully_vaccinated_sort
# Ploting a bar chart of fully vacinated people from all countries
plt.figure(figsize=(18,8))
plt.bar(np.arange(len(fully_vaccinated_sort)), fully_vaccinated_sort["people_fully_vaccinated"], align="center", color="green")
plt.xticks(np.arange(len(fully_vaccinated_sort)),fully_vaccinated_sort['iso_code'])
plt.xticks(rotation=90)
plt.xlabel("Country",fontsize=12)
plt.ylabel("people_fully_vaccinated",fontsize=12)
plt.title("total vaccinations done for top 5 country Details",fontsize=12)
# plt.legend(Gender,loc=2)
plt.show()
# What country is vaccinated a larger percent from its population?
# all the paramters are based on total vaccinations, people vaccinated so lets drop those which are null
people_vacc_df = vaccine_df[['country', 'total_vaccinations_per_hundred']]
people_vacc_df.head()
# mean of all the values of total_vaccinations_per_hundred for each country
# total vaccinations will not be a good measure as US, UK has greater value of it but their population size is also higher
people_vacc_grouped = people_vacc_df.groupby("country").mean()
print(people_vacc_grouped.head())
# people_vacc_grouped.tail()
# Sorting the data based on total_vaccinations_per_hundred from higher to lower
people_vacc_grouped.sort_values(by="total_vaccinations_per_hundred", ascending=False, inplace=True)
print(people_vacc_grouped.head())
people_vacc_grouped
# Based on above data We can observe that Israel, UAE, etc. have a higher ratio of total vaccinations
# per hundred as compared to USA, UK, China
# total_vaccinations_per_hundred
people_vacc_grouped_20 = people_vacc_grouped.iloc[:21]
print(people_vacc_grouped_20.shape)
print(people_vacc_grouped_20.tail())
plt.figure(figsize=(18,8))
plt.bar(np.arange(len(people_vacc_grouped_20)), people_vacc_grouped_20["total_vaccinations_per_hundred"], align="center", color="green")
plt.xticks(np.arange(len(people_vacc_grouped_20)),people_vacc_grouped_20.index)
plt.xticks(rotation=90)
plt.xlabel("Country",fontsize=12)
plt.ylabel("total_vaccinations_per_hundred",fontsize=12)
plt.title("total_vaccinations_per_hundred of top 20 country Details",fontsize=12)
# plt.legend(Gender,loc=2)
plt.show()
# Finding the distribuation of each vaccine
vaccine_df['vaccines'].value_counts()
```
We find that the 'Moderna, Oxford/AstraZeneca, Pfizer/BioNTech' combination has the highest number of doses, while 'Moderna, Oxford/AstraZeneca' has the least number of doses.
This should imply that the 'Moderna, Oxford/AstraZeneca, Pfizer/BioNTech' combination is used in the most number of countries
With all this data analysis, I could able to show the data analysis and visualzation of covid-19 vaccination progress and show the work to achive my work related to initial objectives of this project as given below
I want to showcase the daily covid-19 vaccination progress across all countries for the data available in a plot for easy understanding of trends for all countries using line plots. --- Done
I want to provide a chart with the details of total vaccinations done for each country for all the data available and also provide details about how many people in each country are vaccinated per million to see how fast the vaccination is progressing across countries ---Done
I want to find if all the vaccinations available are used on the same day by comparing vaccination available vs people vaccinated using fields “total_vaccinations” and “people_vaccinated” per day. ---- Done
I want to find what vaccines are used in each country and also want to find which vaccine is more distributed across the world to see the production capability of the companies for these vaccines. ---Done
| github_jupyter |
# Writing Down Qubit States
```
from qiskit import *
```
In the previous chapter we saw that there are multiple ways to extract an output from a qubit. The two methods we've used so far are the z and x measurements.
```
# z measurement of qubit 0
measure_z = QuantumCircuit(1,1)
measure_z.measure(0,0);
# x measurement of qubit 0
measure_x = QuantumCircuit(1,1)
measure_x.h(0)
measure_x.measure(0,0);
```
Sometimes these measurements give results with certainty. Sometimes their outputs are random. This all depends on which of the infinitely many possible states our qubit is in. We therefore need a way to write down these states and figure out what outputs they'll give. For this we need some notation, and we need some math.
### The z basis
If you do nothing in a circuit but a measurement, you are certain to get the outcome `0`. This is because the qubits always start in a particular state, whose defining property is that it is certain to output a `0` for a z measurement.
We need a name for this state. Let's be unimaginative and call it $0$ . Similarly, there exists a qubit state that is certain to output a `1`. We'll call this $1$.
These two states are completely mutually exclusive. Either the qubit definitely outputs a ```0```, or it definitely outputs a ```1```. There is no overlap.
One way to represent this with mathematics is to use two orthogonal vectors.
$$
|0\rangle = \begin{pmatrix} 1 \\\\\\ 0 \end{pmatrix} \, \, \, \, |1\rangle =\begin{pmatrix} 0 \\\\\\ 1 \end{pmatrix}.
$$
This is a lot of notation to take in all at once. First let's unpack the weird $|$ and $\rangle$ . Their job is essentially just to remind us that we are talking about the vectors that represent qubit states labelled $0$ and $1$. This helps us distinguish them from things like the bit values ```0``` and ```1``` or the numbers 0 and 1. It is part of the bra-ket notation, introduced by Dirac.
If you are not familiar with vectors, you can essentially just think of them as lists of numbers which we manipulate using certain rules. If you are familiar with vectors from your high school physics classes, you'll know that these rules make vectors well-suited for describing quantities with a magnitude and a direction. For example, velocity of an object is described perfectly with a vector. However, the way we use vectors for quantum states is slightly different to this. So don't hold on too hard to your previous intuition. It's time to do something new!
In the example above, we wrote the vector as a vertical list of numbers. We call these _column vectors_. In Dirac notation, they are also called _kets_.
Horizontal lists are called _row vectors_. In Dirac notation they are _bras_. They are represented with a $\langle$ and a $|$.
$$
\langle 0| = \begin{pmatrix} 1 & 0\end{pmatrix} \, \, \, \, \langle 1| =\begin{pmatrix} 0 & 1 \end{pmatrix}.
$$
The rules on how to manipulate vectors define what it means to add or multiply them. For example, to add two vectors we need them to be the same type (either both column vectors, or both row vectors) and the same length. Then we add each element in one list to the corresponding element in the other. For a couple of arbitrary vectors that we'll call $a$ and $b$, this works as follows.
$$
\begin{pmatrix} a_0 \\\\ a_1 \end{pmatrix} +\begin{pmatrix} b_0 \\\\ b_1 \end{pmatrix}=\begin{pmatrix} a_0+b_0 \\\\ a_1+b_1 \end{pmatrix}.
$$
To multiple a vector by a number, we simply multiply every element in the list by that number:
$$
x \times\begin{pmatrix} a_0 \\\\ a_1 \end{pmatrix} = \begin{pmatrix} x \times a_0 \\\\ x \times a_1 \end{pmatrix}
$$
Multiplying a vector with another vector is a bit more tricky, since there are multiple ways we can do it. One is called the 'inner product', and works as follows.
$$
\begin{pmatrix} a_0 & a_1 \end{pmatrix} \begin{pmatrix} b_0 \\\\ b_1 \end{pmatrix}= a_0~b_0 + a_1~b_1.
$$
Note that the right hand side of this equation contains only normal numbers being multipled and added in a normal way. The inner product of two vectors therefore yields just a number. As we'll see, we can interpret this as a measure of how similar the vectors are.
The inner product requires the first vector to be a bra and the second to be a ket. In fact, this is where their names come from. Dirac wanted to write the inner product as something like $\langle a | b \rangle$, which looks like the names of the vectors enclosed in brackets. Then he worked backwards to split the _bracket_ into a _bra_ and a _ket_.
If you try out the inner product on the vectors we already know, you'll find
$$
\langle 0 | 0\rangle = \langle 1 | 1\rangle = 1,\\\\
\langle 0 | 1\rangle = \langle 1 | 0\rangle = 0.
$$
Here we are using a concise way of writing the inner products where, for example, $\langle 0 | 1 \rangle$ is the inner product of $\langle 0 |$ with $| 1 \rangle$. The top line shows us that the inner product of these states with themselves always gives a 1. When done with two orthogonal states, as on the bottom line, we get the outcome 0. These two properties will come in handy later on.
### The x basis - part 1
So far we've looked at states for which the z measurement has a certain outcome. But there are also states for which the outcome of a z measurement is equally likely to be `0` or `1`. What might these look like in the language of vectors?
A good place to start would be something like $|0\rangle + |1\rangle$ , since this includes both $|0\rangle$ and $|1\rangle$ with no particular bias towards either. But let's hedge our bets a little and multiply it by some number $x$ .
$$
x ~ (|0\rangle + |1\rangle) = \begin{pmatrix} x \\\\ x \end{pmatrix}
$$
We can choose the value of $x$ to make sure that the state plays nicely in our calculations. For example, think about the inner product,
$$
\begin{pmatrix} x & x \end{pmatrix} \times \begin{pmatrix} x \\\\ x \end{pmatrix}= 2x^2.
$$
We can get any value for the inner product that we want, just by choosing the appropriate value of $x$.
As mentioned earlier, we are going to use the inner product as a measure of how similar two vectors are. With this interpretation in mind, it is natural to require that the inner product of any state with itself gives the value $1$. This is already acheived for the inner products of $|0\rangle$ and $|1\rangle$ with themselves, so let's make it true for all other states too.
This condition is known as the normalization condition. In this case, it means that $x=\frac{1}{\sqrt{2}}$. Now we know what our new state is, so here's a few ways of writing it down.
$$
\begin{pmatrix} \frac{1}{\sqrt{2}} \\\\ \frac{1}{\sqrt{2}} \end{pmatrix} = \frac{1}{\sqrt{2}}\begin{pmatrix} 1 \\\\ 1 \end{pmatrix} = \frac{ |0\rangle + |1\rangle}{\sqrt{2}}
$$
This state is essentially just $|0\rangle$ and $|1\rangle$ added together and then normalized, so we will give it a name to reflect that origin. We call it $|+\rangle$ .
### The Born rule
Now we've got three states that we can write down as vectors. We can also calculate inner products for them. For example, the inner product of each with $\langle 0 |$ is
$$
\langle 0 | 0\rangle = 1 \\\\ \langle 0 | 1\rangle = 0 \\\\ \, \, \, \, \langle 0 | +\rangle = \frac{1}{\sqrt{2}}.
$$
We also know the probabilities of getting various outcomes from a z measurement for these states. For example, let's use $p^z_0$ to denote the probability of the result `0` for a z measurement. The values this has for our three states are
$$
p_0^z( | 0\rangle) = 1,\\\\ p_0^z( | 1\rangle) = 0, \\\\ p_0^z( | +\rangle) = \frac{1}{2}.
$$
As you might have noticed, there's a lot of similarlity between the numbers we get from the inner products and those we get for the probabilities. Specifically, the three probabilities can all be written as the square of the inner products:
$$
p_0^z(|a\rangle) = (~\langle0|a\rangle~)^2.
$$
Here $|a\rangle$ represents any generic qubit state.
This property doesn't just hold for the `0` outcome. If we compare the inner products with $\langle 1 |$ with the probabilities of the `1` outcome, we find a similar relation.
$$
\\\\
p_1^z(|a\rangle) = (~\langle1|a\rangle~)^2.
$$
The same also holds true for other types of measurement. All probabilities in quantum mechanics can be expressed in this way. It is known as the *Born rule*.
### Global and relative phases
Vectors are how we use math to represent the state of a qubit. With them we can calculate the probabilities of all the possible things that could ever be measured. These probabilities are essentially all that is physically relevant about a qubit. It is by measuring them that we can determine or verify what state our qubits are in. Any aspect of the state that doesn't affect the probabilities is therefore just a mathematical curiosity.
Let's find an example. Consider a state that looks like this:
$$
|\tilde 0\rangle = \begin{pmatrix} -1 \\\\ 0 \end{pmatrix} = -|0\rangle.
$$
This is equivalent to multiplying the state $|0\rangle$ by $-1$. It means that every inner product we could calculate with $|\tilde0\rangle$ is the same as for $|0\rangle$, but multplied by $-1$.
$$
\langle a|\tilde 0\rangle = -\langle a| 0\rangle
$$
As you probably know, any negative number squares to the same value as its positive counterpart: $(-x)^2 =x^2$.
Since we square inner products to get probabilities, this means that any probability we could ever calculate for $|\tilde0\rangle$ will give us the same value as for $|0\rangle$. If the probabilities of everything are the same, there is no observable difference between $|\tilde0\rangle$ and $|0\rangle$; they are just different ways of representing the same state.
This is known as the irrelevance of the global phase. Quite simply, this means that multplying the whole of a quantum state by $-1$ gives us a state that will look different mathematically, but which is actually completely equivalent physically.
The same is not true if the phase is *relative* rather than *global*. This would mean multiplying only part of the state by $-1$ , for example:
$$
\begin{pmatrix} a_0 \\\\ a_1 \end{pmatrix} \rightarrow \begin{pmatrix} a_0 \\\\ -a_1 \end{pmatrix}.
$$
Doing this with the $|+\rangle$ state gives us a new state. We'll call it $|-\rangle$.
$$
|-\rangle = \frac{1}{\sqrt{2}}\begin{pmatrix} 1 \\\\ -1 \end{pmatrix} = \frac{ |0\rangle - |1\rangle}{\sqrt{2}}
$$
The values $p_0^z$ and $p_1^z$ for $|-\rangle$ are the same as for $|+\rangle$. These two states are thus indistinguishable when we make only z measurements. But there are other ways to distinguish them. To see how, consider the inner product of $|+\rangle$ and $|-\rangle$.
$$
\langle-|+\rangle = \langle+|-\rangle = 0
$$
The inner product is 0, just as it is for $|0\rangle$ and $|1\rangle$. This means that the $|+\rangle$ and $|-\rangle$ states are orthogonal: they represent a pair of mutually exclusive possible ways for a qubit to be a qubit.
### The x basis - part 2
Whenever we find a pair of orthogonal qubit states, we can use it to define a new kind of measurement.
First, let's apply this to the case we know well: the z measurement. This asks a qubit whether it is $|0\rangle$ or $|1\rangle$. If it is $|0\rangle$, we get the result `0`. For $|1\rangle$ we get `1`. Anything else, such as $|+\rangle$, is treated as a superposition of the two.
$$
|+\rangle = \frac{|0\rangle+|1\rangle}{\sqrt{2}}.
$$
For a superposition, the qubit needs to randomly choose between the two possibilities according to the Born rule.
We can similarly define a measurement based on $|+\rangle$ and $|-\rangle$. This asks a qubit whether it is $|+\rangle$ or $|-\rangle$. If it is $|+\rangle$, we get the result `0`. For $|-\rangle$ we get `1`. Anything else is treated as a superposition of the two. This includes the states $|0\rangle$ and $|1\rangle$, which we can write as
$$
|0\rangle = \frac{|+\rangle+|-\rangle}{\sqrt{2}}, \, \, \, \, |1\rangle = \frac{|+\rangle-|-\rangle}{\sqrt{2}}.
$$
For these, and any other superpositions of $|+\rangle$ and $|-\rangle$, the qubit chooses its outcome randomly with probabilities
$$
p_0^x(|a\rangle) = (~\langle+|a\rangle~)^2,\\\\
p_1^x(|a\rangle) = (~\langle-|a\rangle~)^2.
$$
This is the x measurement.
### The conservation of certainty
Qubits in quantum circuits always start out in the state $|0\rangle$. By applying different operations, we can make them explore other states.
Try this out yourself using a single qubit, creating circuits using operations from the following list, and then doing the x and z measurements in the way described at the top of the page.
```
qc = QuantumCircuit(1)
qc.h(0) # the hadamard
qc.x(0) # x gate
qc.y(0) # y gate
qc.z(0) # z gate
# for the following, replace theta by any number
theta = 3.14159/4
qc.ry(theta,0); # y axis rotation
```
You'll find examples where the z measurement gives a certain result, but the x is completely random. You'll also find examples where the opposite is true. Furthermore, there are many examples where both are partially random. With enough experimentation, you might even uncover the rule that underlies this behavior:
$$
(p^z_0-p^z_1)^2 + (p^x_0-p^x_1)^2 = 1.
$$
This is a version of Heisenberg's famous uncertainty principle. The $(p^z_0-p^z_1)^2$ term measures how certain the qubit is about the outcome of a z measurement. The $(p^x_0-p^x_1)^2$ term measures the same for the x measurement. Their sum is the total certainty of the two combined. Given that this total always takes the same value, we find that the amount of information a qubit can be certain about is a limited and conserved resource.
Here is a program to calculate this total certainty. As you should see, whatever gates from the above list you choose to put in `qc`, the total certainty comes out as $1$ (or as near as possible given statistical noise).
```
shots = 2**14 # number of samples used for statistics
uncertainty = 0
for measure_circuit in [measure_z, measure_x]:
# run the circuit with a the selected measurement and get the number of samples that output each bit value
counts = execute(qc+measure_circuit,Aer.get_backend('qasm_simulator'),shots=shots).result().get_counts()
# calculate the probabilities for each bit value
probs = {}
for output in ['0','1']:
if output in counts:
probs[output] = counts[output]/shots
else:
probs[output] = 0
uncertainty += ( probs['0'] - probs['1'] )**2
# print the total uncertainty
print('The total uncertainty is',uncertainty )
```
Now we have found this rule, let's try to break it! Then we can hope to get a deeper understanding of what is going on. We can do this by simply implementing the operation below, and then recalculating the total uncertainty.
```
# for the following, replace theta by any number
theta = 3.14159/2
qc.rx(theta,0); # x axis rotation
```
For a circuit with a single `rx` with $\theta=\pi/2$, we will find that $(p^z_0-p^z_1)^2 + (p^x_0-p^x_1)^2=0$. This operation seems to have reduced our total certainty to zero.
All is not lost, though. We simply need to perform another identical `rx` gate to our circuit to go back to obeying $(p^z_0-p^z_1)^2 + (p^x_0-p^x_1)^2=1$. This shows that the operation does not destroy our certainty; it simply moves it somewhere else and then back again. So let's find that somewhere else.
### The y basis - part 1
There are infinitely many ways to measure a qubit, but the z and x measurements have a special relationship with each other. We say that they are *mutually unbiased*. This simply means that certainty for one implies complete randomness for the other.
At the end of the last section, it seemed that we were missing a piece of the puzzle. We need another type of measurement to plug the gap in our total certainty, and it makes sense to look for one that is also mutually unbiased with x and z.
The first step is to find a state that seems random to both x and z measurements. Let's call it $|\circlearrowleft\rangle$, for no apparent reason.
$$
|\circlearrowleft\rangle = c_0 | 0 \rangle + c_1 | 1 \rangle
$$
Now the job is to find the right values for $c_0$ and $c_1$. You could try to do this with standard positive and negative numbers, but you'll never be able to find a state that is completely random for both x and z measurements. To achieve this, we need to use complex numbers.
### Complex numbers
Hopefully you've come across complex numbers before, but here is a quick reminder.
Normal numbers, such as the ones we use for counting bananas, are known as *real numbers*. We cannot solve all possible equations using only real numbers. For example, there is no real number that serves as the square root of $-1$. To deal with this issue, we need more numbers, which we call *complex numbers*.
To define complex numbers we start by accepting the fact that $-1$ has a square root, and that its name is $i$. Any complex number can then be written
$$
x = x_r + i~x_i .
$$
Here $x_r$ and $x_i$ are both normal numbers \(positive or negative\), where $x_r$ is known as the real part and $x_i$ as the imaginary part.
For every complex number $x$ there is a corresponding complex conjugate $x^*$
$$
x^* = x_r - i~x_i .
$$
Multiplying $x$ by $x^*$ gives us a real number. It's most useful to write this as
$$
|x| = \sqrt{x~x^*}.
$$
Here $|x|$ is known as the magnitude of $x$ \(or, equivalently, of $x^*$ \).
If we are going to allow the numbers in our quantum states to be complex, we'll need to upgrade some of our equations.
First, we need to ensure that the inner product of a state with itself is always 1. To do this, the bra and ket versions of the same state must be defined as follows:
$$
|a\rangle = \begin{pmatrix} a_0 \\\\ a_1 \end{pmatrix}, ~~~ \langle a| = \begin{pmatrix} a_0^* & a_1^* \end{pmatrix}.
$$
Then we just need a small change to the Born rule, where we square the magnitudes of inner products, rather than just the inner products themselves.
$$
p_0^z(|a\rangle) = |~\langle0|a\rangle~|^2,\\\\
p_1^z(|a\rangle) = |~\langle1|a\rangle~|^2,\\\\
p_0^x(|a\rangle) = |~\langle+|a\rangle~|^2,\\\\
p_1^x(|a\rangle) = |~\langle-|a\rangle~|^2.
$$
The irrelevance of the global phase also needs an upgrade. Previously, we only talked about multiplying by -1. In fact, we can multiply a state by any complex number whose magnitude is 1. This will give us a state that will look different, but which is actually completely equivalent. This includes multiplying by $i$, $-i$ or infinitely many other possibilities.
### The y basis - part 2
Now that we have complex numbers, we can define the following pair of states.
$$
|\circlearrowright\rangle = \frac{ | 0 \rangle + i | 1 \rangle}{\sqrt{2}}, ~~~~ |\circlearrowleft\rangle = \frac{ | 0 \rangle -i | 1 \rangle}{\sqrt{2}}
$$
You can verify yourself that they both give random outputs for x and z measurements. They are also orthogonal to each other. They therefore define a new measurement, and that basis is mutally unbiased with x and z. This is the third and final fundamental measurement for a single qubit. We call it the y measurement, and can implement it with
```
# y measurement of qubit 0
measure_y = QuantumCircuit(1,1)
measure_y.sdg(0)
measure_y.h(0)
measure_y.measure(0,0);
```
With the x, y and z measurements, we now have everything covered. Whatever operations we apply, a single isolated qubit will always obey
$$
(p^z_0-p^z_1)^2 + (p^y_0-p^y_1)^2 + (p^x_0-p^x_1)^2 = 1.
$$
To see this, we can incorporate the y measurement into our measure of total certainty.
```
shots = 2**14 # number of samples used for statistics
uncertainty = 0
for measure_circuit in [measure_z, measure_x, measure_y]:
# run the circuit with a the selected measurement and get the number of samples that output each bit value
counts = execute(qc+measure_circuit,Aer.get_backend('qasm_simulator'),shots=shots).result().get_counts()
# calculate the probabilities for each bit value
probs = {}
for output in ['0','1']:
if output in counts:
probs[output] = counts[output]/shots
else:
probs[output] = 0
uncertainty += ( probs['0'] - probs['1'] )**2
# print the total uncertainty
print('The total uncertainty is',uncertainty )
```
For more than one qubit, this relation will need another upgrade. This is because the qubits can spend their limited certainty on creating correlations that can only be detected when multiple qubits are measured. The fact that certainty is conserved remains true, but it can only be seen when looking at all the qubits together.
Before we move on to entanglement, there is more to explore about just a single qubit. As we'll see in the next section, the conservation of certainty leads to a particularly useful way of visualizing single-qubit states and gates.
```
import qiskit
qiskit.__qiskit_version__
```
| github_jupyter |
## Import Librarys
```
import numpy as np
import os
import cv2
import time
from model.yolo_model import YOLO
```
## Create some important functions
**Define the process image function**
```
def process_image(img):
# resize
image = cv2.resize(img, (416, 416), interpolation=cv2.INTER_CUBIC)
image = np.array(image, dtype="float32")
# normalize
image /= 255.
image = np.expand_dims(image, axis=0)
return image
```
**Draw rectangle and detect function on image**
```
def draw(image, boxes, scores, classes, all_classes):
for box, score, cl in zip(boxes, scores, classes):
x, y, w, h = box
top = max(0, np.floor(x+0.5).astype(int))
left = max(0, np.floor(y+0.5).astype(int))
right = min(image.shape[1], np.floor(x+w+0.5).astype(int))
bottom = min(image.shape[0], np.floor(y+h+0.5).astype(int))
cv2.rectangle(image, (top, left), (right, bottom), (255, 0, 0), 2)
cv2.putText(image, '{0} {1:.2f}'.format(all_classes[cl], score),
(top, left - 6),
cv2.FONT_HERSHEY_SIMPLEX,
0.6, (0, 0, 255), 1,
cv2.LINE_AA)
print("box coordinate x, y, w, h: {0}".format(box))
print()
def detect_image(image, yolo, all_classes):
pimage = process_image(image)
start = time.time()
boxes, classes, scores = yolo.predict(pimage, image.shape)
end = time.time()
print("time: {0:.2f}".format(end-start))
if boxes is not None:
draw(image, boxes, scores, classes, all_classes)
return image
def detect_video(video, yolo, all_classes):
video_path = os.path.join("/Users/neemiasbsilva/Downloads/Computer-Vision-with-Python/06-Deep-Learning-Computer-Vision/06-YOLOv3/videos", "test", video)
camera = cv2.VideoCapture(video_path)
cv2.namedWindow("detection", cv2.WINDOW_AUTOSIZE)
# Prepare for saving the detected video
sz = (int(camera.get(cv2.CAP_PROP_FRAME_WIDTH)),
int(camera.get(cv2.CAP_PROP_FRAME_HEIGHT)))
fourcc = cv2.VideoWriter_fourcc(*'mpeg')
vout = cv2.VideoWriter()
vout.open(os.path.join("/Users/neemiasbsilva/Downloads/Computer-Vision-with-Python/06-Deep-Learning-Computer-Vision/06-YOLOv3/videos", "res", video), fourcc, 20, sz, True)
while True:
res, frame = camera.read()
if not res:
break
image = detect_image(frame, yolo, all_classes)
cv2.imshow("detection", image)
# Save the video frame by frame
vout.write(image)
if cv2.waitKey(110) & 0xff == 27:
break
vout.release()
camera.release()
```
**Get classes name of coco dataset**
```
def get_classes(file):
with open(file) as f:
class_names = f.readlines()
class_names = [c.strip() for c in class_names]
return class_names
```
## Detecting image
```
yolo = YOLO(0.6, 0.5)
file = "coco_classes.txt"
all_classes = get_classes(file)
f = "person-car-02.jpg"
path = "images/"+f
image = cv2.imread(path)
print(image.shape)
image = detect_image(image, yolo, all_classes)
cv2.imwrite("images/result/"+f, image)
```
| github_jupyter |
# $\ell_1$ trend filtering
**Reference:** S.-J. Kim, K. Koh, S. Boyd, and D. Gorinevsky. [*$\ell_1$ Trend Filtering*.](http://stanford.edu/~boyd/papers/l1_trend_filter.html) SIAM Review, 51(2):339-360, 2009.
## Introduction
The problem of estimating underlying trends in time series data arises in a variety of disciplines. The $\ell_1$ trend filtering method produces trend estimates $z$ that are piecewise linear from the time series $y$.
The $\ell_1$ trend estimation problem can be formulated as
$$\text{minimize}~ \frac{1}{2}\|y - z\|_2^2 + \alpha \|Dz\|_1,$$
with variable $z \in \mathbf{R}^q$, problem data $y \in \mathbf{R}^q$, and smoothing parameter $\alpha \geq 0$. Here $D \in \mathbf{R}^{(q-2) \times q}$ is the second difference matrix
$$D = \left[\begin{array}{ccccccc}
1 & -2 & 1 & 0 & \ldots & 0 &0 \\
0 & 1 & -2 & 1 & \ldots & 0 & 0 \\
\vdots & \vdots & \ddots & \ddots & \ddots & \vdots& \vdots \\
0 & 0 & \ldots &1 & -2 & 1 & 0 \\
0 & 0 & \ldots & 0 & 1 & -2 & 1
\end{array}\right].$$
## Reformulate and Solve Problem
This problem can be written in standard form by letting
$$f_1(x_1) = \frac{1}{2}\|y - x_1\|_2^2, \quad f_2(x_2) = \alpha \|x_2\|_1,$$
$$A_1 = D, \quad A_2 = -I, \quad b = 0,$$
where the variables $x_1 \in \mathbf{R}^q$ and $x_2 \in \mathbf{R}^{q-2}$. We solve an instance where $y$ is a snapshot of the S&P 500 price for $q = 2000$ time steps and $\alpha = 0.01\|y\|_{\infty}$.
```
import numpy as np
from scipy import sparse
from a2dr import a2dr
from a2dr.proximal import *
# Load time series data: S&P 500 price log.
y = np.loadtxt(open("data/snp500.txt", "rb"), delimiter = ",")
q = y.size
alpha = 0.01*np.linalg.norm(y, np.inf)
# Form second difference matrix.
D = sparse.lil_matrix(sparse.eye(q))
D.setdiag(-2, k = 1)
D.setdiag(1, k = 2)
D = D[:(q-2),:]
# Convert problem to standard form.
prox_list = [lambda v, t: prox_sum_squares(v, t = 0.5*t, offset = y),
lambda v, t: prox_norm1(v, t = alpha*t)]
A_list = [D, -sparse.eye(q-2)]
b = np.zeros(q-2)
# Solve with A2DR.
a2dr_result = a2dr(prox_list, A_list, b)
# Save solution.
z_star = a2dr_result["x_vals"][0]
print("Solve time:", a2dr_result["solve_time"])
print("Number of iterations:", a2dr_result["num_iters"])
```
## Plot Results
```
import matplotlib.pyplot as plt
# Show plots inline in ipython.
%matplotlib inline
# Plot properties.
plt.rc("text", usetex = True)
plt.rc("font", family = "serif")
font = {"weight" : "normal",
"size" : 16}
plt.rc("font", **font)
# Plot estimated trend with original signal.
plt.figure(figsize = (6, 6))
plt.plot(np.arange(1,q+1), y, "k:", linewidth = 1.0)
plt.plot(np.arange(1,q+1), z_star, "b-", linewidth = 2.0)
plt.xlabel("Time")
```
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
from sklearn.metrics.pairwise import euclidean_distances
import sklearn.metrics as me
from __future__ import division
%matplotlib inline
layer_1 = np.array([[1,7],[5,7],[6,7], [9,8]])
layer_2 = np.array([[1,1],[5,1],[9,1]])
layer_3 = np.array([[1,1],[3,2],[7,4]])
dip_pos_1 = np.array([2,4])
dip_angle_1 = 135
dip_pos_1_v = np.array([np.cos(np.deg2rad(dip_angle_1))*1,
np.sin(np.deg2rad(dip_angle_1))]) + dip_pos_1
dip_pos_2 = np.array([6,3])
dip_angle_2 = 45
dip_pos_2_v = np.array([np.cos(np.deg2rad(dip_angle_2))*1,
np.sin(np.deg2rad(dip_angle_2))]) + dip_pos_2
plt.arrow(dip_pos_1[0],dip_pos_1[1], dip_pos_1_v[0]-dip_pos_1[0],
dip_pos_1_v[1]-dip_pos_1[1], head_width = 0.2)
plt.arrow(dip_pos_2[0],dip_pos_2[1],dip_pos_2_v[0]-dip_pos_2[0],
dip_pos_2_v[1]-dip_pos_2[1], head_width = 0.2)
plt.plot(layer_1[:,0],layer_1[:,1], "o")
plt.plot(layer_2[:,0],layer_2[:,1], "o")
plt.xlim(0,10)
plt.ylim(0,10)
print (dip_pos_1_v, dip_pos_2_v, layer_1)
layers = np.asarray([layer_1,layer_2])
dips = np.asarray([dip_pos_1,dip_pos_2])
#layers = [np.random.uniform(0,10,(100,2)) for i in range(2)]
#dips = np.random.uniform(0,10, (30,2))
#dips_angles = np.random.normal(90,10, 8)
import pdb;
# =================================0
# THE INCREMENTS OF POTENTIAL
def cov_cubic_f(r,a = 6, c_o = 1):
ans_d0 = c_o*(1-7*(r/a)**2+35/4*(r/a)**3-7/2*(r/a)**5+3/4*(r/a)**7)
ans_d0[r>a] = 0
return ans_d0
def cov_cubic_d1_f(r,a = 6., c_o = 1):
ans_d1 = (-7* (a - r)**3 *r* (8* a**2 + 9 *a* r + 3* r**2)* (c_o))/(4* a**7)
ans_d1[r>a] = 0.
return ans_d1
def cov_cubic_d2_f(r, a = 6, c_o = 1):
ans_d2 = (-7 * (4.* a**5. - 15. *a**4. * r + 20. *( a**2)*(r**3) - 9* r**5) *
(c_o))/(2*a**7)
ans_d2[r>a] = 0
return ans_d2
def cov_cubic_layer(X,Y, a = 6., c_o = 1., verbose = 0):
"""x = Array: Position of the measured points"
a = Range of the spherical semivariogram
C_o = Nugget, variance
"""
# Creation of r vector
r_m = np.asarray(euclidean_distances(X,Y))
# Initializing
# Applying the functio
C_h = c_o*(1.-7.*(r_m/a)**2.+35./4.*(r_m/a)**3.
-7./2.*(r_m/a)**5.+3./4.*(r_m/a)**7.)
C_h[r_m>a] = 0
if verbose !=0:
print(r_m>a)
print ("Our lag matrix is", r_m)
print("Our covariance matrix is",C_h)
return C_h
def C_I(layers, a = 6.):
#print "layers", layers
layers = np.asarray(layers)
#print "layers", len(layers)
for r in range(len(layers)):
for s in range(len(layers)):
# print "layers2", layers[r][1:],layers[s][1:]
# print "nagnoagjja", layers[s][0].reshape(1,-1),layers[r][1:],
a_p = cov_cubic_layer(layers[r][1:],layers[s][1:], a = a)
b_p = cov_cubic_layer(layers[s][0].reshape(1,-1),
layers[r][1:], a = a).transpose()
test = cov_cubic_layer(layers[r][1:],layers[s][0].reshape(1,-1), a =a)
c_p = cov_cubic_layer(layers[s][1:],
layers[r][0].reshape(1,-1),a=a).transpose()
d_p = cov_cubic_layer(layers[r][0].reshape(1,-1),
layers[s][0].reshape(1,-1), a=a)
#pdb.set_trace()
#print "s", s,"r", r
if s == 0:
C_I_row = a_p-b_p-c_p+d_p
else:
C_I_row = np.hstack((C_I_row, a_p-b_p-c_p+d_p))
if r == 0:
C_I = C_I_row
else:
C_I = np.vstack((C_I,C_I_row))
# C_I += 0.00000001
return C_I
C_I(layers)
#=====================
# THE GRADIENTS
def h_f(dips, direct):
#pdb.set_trace()
if direct == "x":
#print (np.subtract.outer(dips[:,0],dips[:,0]))
#print (dips[:,0] , dips[:,0].reshape((dips[:,0].shape[0],1)))
# print ("hx",dips[:,0] - dips[:,0].reshape((dips[:,0].shape[0],1)))
return (np.subtract.outer(dips[:,0],dips[:,0]))
if direct == "y":
# print ("hy",dips[:,1] - dips[:,1].reshape((dips[:,1].shape[0],1)))
return (np.subtract.outer(dips[:,1],dips[:,1]))
def C_G(dips, sig_z = 1, a = 6., C_000 = -14*1/6**2-0.2):
dips = np.asarray(dips)
if len(dips) == 1:
lalolu = np.ones((2,2))*C_0
lalolu[0,1 ] = 0
lalolu[1,0] = 0
return lalolu
r = me.euclidean_distances(dips)
# print ("R",r)
C_0 = 1
for i in "xy":
for j in "xy":
if i == j and j == "x":
h0 = h_f(dips, direct = i)
C_G_row = (C_0*(h0**2/r**3-1/r)*cov_cubic_d1_f(r, a = a) -
(h0/r)**2*cov_cubic_d2_f(r, a = a))
# print ("teta", C_0*(h0**2/r**3-1/r)*cov_cubic_d1_f(r, a = a) -
# (h0/r)**2*cov_cubic_d2_f(r, a = a))
h1 = h_f(dips, direct = i)
h2 = h_f(dips, direct = j)
# print ("teta 3" , ((C_0*(h0*h0/r**2)*
# ((1/r)*cov_cubic_d1_f(r, a = a)
# -cov_cubic_d2_f(r, a = a)))))
# pdb.set_trace()
# print "0"
elif i == j:
# print ("a",h0**2)
h0 = h_f(dips, direct = i)
C_G_row = np.hstack((C_G_row, (
C_0*(h0**2/r**3-1/r)
*cov_cubic_d1_f(r, a = a) -
(h0/r)**2*cov_cubic_d2_f(r, a = a))))
# pdb.set_trace()
else:
if j == "x":
"""
print ("cov_d1",cov_cubic_d1_f(r, a = a))
print ("cov_d2",cov_cubic_d2_f(r, a = a))
print ("a",h1*h2)
print ("b", C_0*(h1*h2/r**2) )
print ("c", r**2)
"""
h1 = h_f(dips, direct = i)
h2 = h_f(dips, direct = j)
C_G_row = ((C_0*(h1*h2/r**2)*
((1/r)*cov_cubic_d1_f(r, a = a)
-cov_cubic_d2_f(r, a = a))))
# pdb.set_trace()
# print "2"
else:
h1 = h_f(dips, direct = i)
h2 = h_f(dips, direct = j)
# print ("a",h1*h2)
C_G_row = np.hstack((C_G_row, (C_0*(h1*h2/r**2)*
((1/r)*cov_cubic_d1_f(r, a = a)
-cov_cubic_d2_f(r, a = a)))))
# pdb.set_trace()
# print "3"
if i == "x":
C_G = C_G_row
else:
C_G = np.vstack((C_G, C_G_row))
# C_G[C_G == 0] = 0.0000000000000000000001
sol_CG = np.nan_to_num(C_G)
#sol_CG = C_G
# sol_CG[sol_CG== 0] = C_0
g,h = np.indices(np.shape(sol_CG))
sol_CG[g==h] = C_000
# sol_CG[g+2==h] = 0.01
# sol_CG[g-2==h] = 0.01
# sol_CG[sol_CG == 0] = C_0
# sol_CG[2,0] = -C_0
# sol_CG[3,1] = -C_0
# print (sol_CG)
return sol_CG
C_G(dips)
#========================================
#THE INTERACTION GRADIENTS/INTERFACES
def h_f_GI(dips, layers, direct):
if direct == "x":
return (np.subtract.outer(dips[:,0],layers[:,0]))
if direct == "y":
return (np.subtract.outer(dips[:,1],layers[:,1]))
def C_GI(dips,layers, sig_z = 1., a = 6., C_01 = 1, verbose = 0):
dips = np.asarray(dips)
layers = np.asarray(layers)
C_00 = C_01
for k in range(len(layers)):
for i in "xy":
r = me.euclidean_distances(dips,layers[k])
h1 = h_f_GI(dips,layers[k], i)
Cov_d1 = cov_cubic_d1_f(r, a = a)
# pdb.set_trace()
if verbose != 0:
print ("dips", dips)
print ("layers", layers)
print ("h1", h1, h1[:,0])
print ("")
print ("r", r, r[:,0])
print ("")
print ("Cov_d1", Cov_d1)
if i == "x":
cov_1 = C_00* h1[:,0] / r[:,0] * Cov_d1[:,0]
cov_j = C_00* h1[:,1:] / r[:,1:] * Cov_d1[:,1:]
# C_GI_row = alpha * sig_z / a**2 * h1 / r * Cov_d1
# print "cov_j, cov_1", cov_j, cov_1.reshape(-1,1), "h1",h1
C_GI_row = cov_j.transpose()-cov_1#.transpose()
# pdb.set_trace()
else:
cov_1 = C_00* h1[:,0] / r[:,0] * Cov_d1[:,0]
cov_j = C_00* h1[:,1:] / r[:,1:] * Cov_d1[:,1:]
#C_GI_row = np.hstack((C_GI_row,
# alpha * sig_z / a**2 * h1 / r * Cov_d1))
#pdb.set_trace()
C_GI_row = np.hstack((C_GI_row, cov_j.transpose()-cov_1))
# pdb.set_trace()
#.reshape(-1,1)))
if k==0:
C_GI = C_GI_row
else:
#pdb.set_trace()
C_GI = np.vstack((C_GI,C_GI_row))
return C_GI
np.set_printoptions(precision=2)
a = C_GI(dips,layers, verbose =0)
a
#======================
# The condition of universality
# GRADIENTS
def U_G(dips):
dips = np.asarray(dips)
n = len(dips)
# x
U_G = np.array([np.ones(n), #x
np.zeros(n),]) #y
# dips[:,0]*2, #xx
# np.zeros(n), #yy
# dips[:,1],]) #xy
# y
U_G = np.hstack((U_G,np.array([np.zeros(n),
np.ones(n),])))
# np.zeros(n),
# 2*dips[:,1]
# ,dips[:,0]])))
return U_G
U_G(dips)
#======================
# The condition of universality
# Interfaces
def U_I(layers):
layers = np.asarray(layers)
for e,l in enumerate(layers):
if e == 0:
U_I = np.array([(l[1:,0]-l[0,0]), # x
(l[1:,1]-l[0,1]),]) # y
# np.square(l[1:,0])- np.square(l[0,0]), # xx
# np.square(l[1:,1])- np.square(l[0,1]), # yy
#(l[1:,0]* l[1:,1])-(l[0,0]*l[0,1])]) #xy
else:
U_I = np.hstack((U_I, np.array([(l[1:,0]-l[0,0]), # x
(l[1:,1]-l[0,1]),]))) # y
# np.square(l[1:,0])- np.square(l[0,0]), # xx
# np.square(l[1:,1])- np.square(l[0,1]), # yy
# (l[1:,0]* l[1:,1])-(l[0,0]*l[0,1])]))) #xy
return U_I
U_I(layers);
theano_CG = np.array([[-0.58888888, -0.13136305, 0. , 0.03284076],
[-0.13136305, -0.58888888, 0.03284076, 0. ],
[ 0. , -0.06287594, -0.58888888, -0.10392689],
[ 0.03284076, 0. , -0.10392689, -0.58888888]]
)
theano_CG
```
## A matrix
```
def A_matrix(layers,dips, sig_z = 1., a = 6., C_0 = -14*1/6**2-0.2,
C_01 = 1, verbose = 0):
#CG = theano_CG
CG = C_G(dips)
CGI = C_GI(dips,layers,a = a, C_01=C_01)
CI = C_I(layers, a = a)
UG = U_G(dips)
UI = U_I(layers)
# print np.shape(UI)[0]
zeros = np.zeros((np.shape(UI)[0],np.shape(UI)[0]))
#print CG,CGI.transpose(),UG.transpose()
A1 = np.hstack((-CG,CGI.transpose(),UG.transpose()))
A2 = np.hstack((CGI,CI,UI.transpose()))
A3 = np.hstack((UG,UI,zeros))
A = np.vstack((A1,A2,A3))
return A
np.set_printoptions(precision = 2, linewidth= 130, suppress = True)
aa = A_matrix(layers, dips)
np.shape(aa)
#aa
```
### Dual Kriging
```
def G_f(dips, dips_v):
a_g = np.asarray(dips)
b_g = np.asarray(dips_v)
# print a, a[:,0]
# print b,b[:,0]
Gx = b_g[:,0] - a_g[:,0] # x
Gy = b_g[:,1] -a_g[:,1] # y
G = np.hstack((Gx,Gy))
# G = np.array([-0.71,0.34,0.71,0.93])
return G
def b(dips,dips_v,n):
n -= len(dips)*2 # because x and y direction
G = G_f(dips,dips_v)
b = np.hstack((G, np.zeros(n)))
return b
```
## Estimator normal
```
aa = A_matrix(layers, dips)
bb = b([dip_pos_1, dip_pos_2],
[dip_pos_1_v,dip_pos_2_v], len(aa))
# bb[1] = 0
print (bb)
sol = np.linalg.solve(aa,bb)
aa
bb
sol
x = [1,1]
def estimator(x, dips, layers, sol, sig_z = 1., a = 6., C_01 = 1, verbose = 0):
x = np.asarray(x).reshape(1,-1)
dips = np.asarray(dips)
layers = np.asarray(layers)
C_01 = C_01
n = 0
m = len(dips)
# print layers
# print x.reshape(1,-1), dips
r_i = me.euclidean_distances(dips,x)
hx = h_f_GI(dips, x, "x")
Cov_d1 = cov_cubic_d1_f(r_i, a = a)
KzGx = sol[:m] * np.squeeze( C_01*hx / r_i * Cov_d1)
hy = h_f_GI(dips, x, "y")
KzGy = sol[m:2*m] * np.squeeze( C_01 * hy / r_i * Cov_d1)
# KzGx[KzGx == 0] = -0.01
# KzGy[KzGy == 0] = -0.01
# print "KzGx", KzGx, sol[:m]
for s in range(len(layers)):
n += len(layers[s][1:])
a_l = cov_cubic_layer(x, layers[s][1:], a = a)
b_l = cov_cubic_layer(x, layers[s][0].reshape(1,-1), a = a)
aux = a_l-b_l
# aux[aux==0] = 0.000001
if s == 0:
L = np.array(sol[2*m:2*m+n]*(aux))
else:
L = np.hstack((L,sol[2*m+n2:2*m+n]*(aux)))
n2 = n
L = np.squeeze(L)
univ = (sol[2*m+n]*x[0,0] + # x
sol[2*m+n+1] * x[0,1] ) # y
# + sol[2*m+n+2]* x[0,0]**2 # xx
# + sol[2*m+n+3] * x[0,1]**2 # yy
# + sol[2*m+n+4] * x[0,0]*x[0,1]) #xy
if verbose != 0:
print (KzGx, KzGy, L ,univ)
print (Cov_d1, r_i)
print ("")
print (hx, hx/r_i)
print ("angaglkagm",hy/r_i, sol[m:2*m])
z_star = np.sum(KzGx)+np.sum(KzGy)+np.sum(L)+univ
return z_star
pot = np.zeros((100,100))
for i in range(100):
for j in range(100):
pot[i,j] = estimator([i/10.,j/10.],[dip_pos_1, dip_pos_2],
[layer_1, layer_2]
, sol, verbose = 0, C_01 = 1,
a = 6.)
plt.arrow(dip_pos_1[0],dip_pos_1[1], dip_pos_1_v[0]-dip_pos_1[0],
dip_pos_1_v[1]-dip_pos_1[1], head_width = 0.2)
plt.arrow(dip_pos_2[0],dip_pos_2[1],dip_pos_2_v[0]-dip_pos_2[0],
dip_pos_2_v[1]-dip_pos_2[1], head_width = 0.2)
plt.plot(layer_1[:,0],layer_1[:,1], "o")
plt.plot(layer_2[:,0],layer_2[:,1], "o")
plt.plot(layer_1[:,0],layer_1[:,1], )
plt.plot(layer_2[:,0],layer_2[:,1], )
plt.contour(pot.transpose(),30,extent = (0,10,0,10) )
plt.colorbar()
plt.xlim(0,10)
plt.ylim(0,10)
plt.title("GeoMigueller v 0.1")
print (dip_pos_1_v, dip_pos_2_v, layer_1)
```
## La Buena
```
%matplotlib inline
def pla(angle1,angle2, C_0 = -14*1/6**2-0.2, C_01 = 1):
layer_1 = np.array([[1,7],[5,6], [6,8], [9,9] ])
layer_2 = np.array([[1,2],[5,3], [9,7]])
layer_3 = np.array([[1,1],[3,2],[7,4]])
dip_pos_1 = np.array([3,4])
dip_angle_1 = angle1
dip_pos_1_v = np.array([np.cos(np.deg2rad(dip_angle_1))*1,
np.sin(np.deg2rad(dip_angle_1))]) + dip_pos_1
dip_pos_2 = np.array([6,6])
dip_angle_2 = angle2
dip_pos_2_v = np.array([np.cos(np.deg2rad(dip_angle_2))*1,
np.sin(np.deg2rad(dip_angle_2))]) + dip_pos_2
dip_pos_3 = np.array([9,5])
dip_angle_3 = 90
dip_pos_3_v = np.array([np.cos(np.deg2rad(dip_angle_3))*1,
np.sin(np.deg2rad(dip_angle_3))]) + dip_pos_3
#print b([dip_pos_1,dip_pos_2], [dip_pos_1_v,dip_pos_2_v],13)
aa = A_matrix([layer_1,layer_2, layer_3],
[dip_pos_1,dip_pos_2, dip_pos_3], a = 6.,
C_0= C_0,
C_01 = C_01)
bb = b([dip_pos_1, dip_pos_2, dip_pos_3],
[dip_pos_1_v,dip_pos_2_v, dip_pos_3_v], len(aa))
# bb[1] = 0
print (bb)
sol = np.linalg.solve(aa,bb)
#sol[:-2] = 0
#print aa
print( sol)
pot = np.zeros((50,50))
for i in range(50):
for j in range(50):
pot[i,j] = estimator([i/5.,j/5.],[dip_pos_1, dip_pos_2, dip_pos_3],
[layer_1, layer_2, layer_3]
, sol, verbose = 0, C_01 = C_01,
a = 6.)
plt.arrow(dip_pos_1[0],dip_pos_1[1], dip_pos_1_v[0]-dip_pos_1[0],
dip_pos_1_v[1]-dip_pos_1[1], head_width = 0.2)
plt.arrow(dip_pos_2[0],dip_pos_2[1],dip_pos_2_v[0]-dip_pos_2[0],
dip_pos_2_v[1]-dip_pos_2[1], head_width = 0.2)
plt.arrow(dip_pos_3[0],dip_pos_3[1],dip_pos_3_v[0]-dip_pos_3[0],
dip_pos_3_v[1]-dip_pos_3[1], head_width = 0.2)
plt.plot(layer_1[:,0],layer_1[:,1], "o")
plt.plot(layer_2[:,0],layer_2[:,1], "o")
plt.plot(layer_3[:,0],layer_3[:,1], "o")
plt.plot(layer_1[:,0],layer_1[:,1], )
plt.plot(layer_2[:,0],layer_2[:,1], )
plt.plot(layer_3[:,0],layer_3[:,1], )
plt.contour(pot.transpose(),30,extent = (0,10,0,10) )
plt.colorbar()
plt.xlim(0,10)
plt.ylim(0,10)
plt.title("GeoMigueller v 0.1")
print (dip_pos_1_v, dip_pos_2_v, layer_1)
return pot
jhjs2 = pla(120,130,C_0=-0.5, C_01 = 1)
# jhjs = pla(120,-30, C_01 = 0.9)
jhjs = pla(120,-30)
jh = pla(120,0)
jhjs = pla(-2,0)
137.769228/3.184139677, -106.724083/-2.9572844241540727772132
3.16/0.15
59.12/3.15, 3.16/0.1425
51.109568/3.15, 2.669329/0.1425
45.047943/3.15, 2.29186/0.1425
layer_1 = np.array([[1,7],[5,7],[6,7], [9,8], ])
layer_2 = np.array([[1,1],[5,1],[9,1], ])
layer_3 = np.array([[1,1],[3,2],[7,4]])
dip_pos_1 = np.array([2,4])
dip_angle_1 = 45
dip_pos_1_v = np.array([np.cos(np.deg2rad(dip_angle_1))*1,
np.sin(np.deg2rad(dip_angle_1))]) + dip_pos_1
dip_pos_2 = np.array([9,7])
dip_angle_2 = 90
dip_pos_2_v = np.array([np.cos(np.deg2rad(dip_angle_2))*1,
np.sin(np.deg2rad(dip_angle_2))]) + dip_pos_2
dip_pos_3 = np.array([5,5])
dip_angle_3 = 90
dip_pos_3_v = np.array([np.cos(np.deg2rad(dip_angle_3))*1,
np.sin(np.deg2rad(dip_angle_3))]) + dip_pos_3
#print b([dip_pos_1,dip_pos_2], [dip_pos_1_v,dip_pos_2_v],13)
aa = A_matrix([layer_1,layer_2], [dip_pos_1,dip_pos_2], a = 6., alpha = 14)
bb = b([dip_pos_1,dip_pos_2], [dip_pos_1_v,dip_pos_2_v], 11)
print bb
sol = np.linalg.solve(aa,bb)
#sol[:-2] = 0
#print aa
print sol
pot = np.zeros((50,50))
for i in range(50):
for j in range(50):
pot[i,j] = estimator([i/5.,j/5.],[dip_pos_1,dip_pos_2],
[layer_1,layer_2], sol, verbose = 0, alpha = 14,
a = 6.)
plt.arrow(dip_pos_1[0],dip_pos_1[1], dip_pos_1_v[0]-dip_pos_1[0],
dip_pos_1_v[1]-dip_pos_1[1], head_width = 0.2)
plt.arrow(dip_pos_2[0],dip_pos_2[1],dip_pos_2_v[0]-dip_pos_2[0],
dip_pos_2_v[1]-dip_pos_2[1], head_width = 0.2)
#plt.arrow(dip_pos_3[0],dip_pos_3[1],dip_pos_3_v[0]-dip_pos_3[0],
# dip_pos_3_v[1]-dip_pos_3[1], head_width = 0.2)
plt.plot(layer_1[:,0],layer_1[:,1], "o")
plt.plot(layer_2[:,0],layer_2[:,1], "o")
#plt.plot(layer_3[:,0],layer_3[:,1], "o")
plt.plot(layer_1[:,0],layer_1[:,1], )
plt.plot(layer_2[:,0],layer_2[:,1], )
plt.contour(pot.transpose(),20,extent = (0,10,0,10) )
plt.colorbar()
plt.xlim(0,10)
plt.ylim(0,10)
print dip_pos_1_v, dip_pos_2_v, layer_1
np.cos(np.deg2rad(45))
layer_1 = np.array([[1,7],[5,7],[6,7], [9,7], ])
layer_2 = np.array([[1,1],[5,1],[9,1], ])
layer_3 = np.array([[1,1],[3,2],[7,4]])
dip_pos_1 = np.array([2,4])
dip_angle_1 = 100
dip_pos_1_v = np.array([np.cos(np.deg2rad(dip_angle_1))*1,
np.sin(np.deg2rad(dip_angle_1))]) + dip_pos_1
dip_pos_2 = np.array([8,5])
dip_angle_2 = 70
dip_pos_2_v = np.array([np.cos(np.deg2rad(dip_angle_2))*1,
np.sin(np.deg2rad(dip_angle_2))]) + dip_pos_2
dip_pos_3 = np.array([8,5])
dip_angle_3 = 90
dip_pos_3_v = np.array([np.cos(np.deg2rad(dip_angle_3))*1,
np.sin(np.deg2rad(dip_angle_3))]) + dip_pos_3
#print b([dip_pos_1,dip_pos_2], [dip_pos_1_v,dip_pos_2_v],13)
aa = A_matrix([layer_1,layer_2], [dip_pos_1,dip_pos_2], a = 6., alpha = 14)
bb = b([dip_pos_1,dip_pos_2], [dip_pos_1_v,dip_pos_2_v], 11)
print bb
sol = np.linalg.solve(aa,bb)
#sol[:-2] = 0
#print aa
print sol
pot = np.zeros((50,50))
for i in range(50):
for j in range(50):
pot[i,j] = estimator([i/5.,j/5.],[dip_pos_1,dip_pos_2],
[layer_1,layer_2], sol, verbose = 0, alpha = 14,
a = 6.)
plt.arrow(dip_pos_1[0],dip_pos_1[1], dip_pos_1_v[0]-dip_pos_1[0],
dip_pos_1_v[1]-dip_pos_1[1], head_width = 0.2)
plt.arrow(dip_pos_2[0],dip_pos_2[1],dip_pos_2_v[0]-dip_pos_2[0],
dip_pos_2_v[1]-dip_pos_2[1], head_width = 0.2)
#plt.arrow(dip_pos_3[0],dip_pos_3[1],dip_pos_3_v[0]-dip_pos_3[0],
# dip_pos_3_v[1]-dip_pos_3[1], head_width = 0.2)
plt.plot(layer_1[:,0],layer_1[:,1], "o")
plt.plot(layer_2[:,0],layer_2[:,1], "o")
#plt.plot(layer_3[:,0],layer_3[:,1], "o")
plt.plot(layer_1[:,0],layer_1[:,1], )
plt.plot(layer_2[:,0],layer_2[:,1], )
plt.contour(pot.transpose(),20,extent = (0,10,0,10) )
plt.colorbar()
plt.xlim(0,10)
plt.ylim(0,10)
print dip_pos_1_v, dip_pos_2_v, layer_1
%matplotlib inline
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import cm
from matplotlib.ticker import LinearLocator, FormatStrFormatter
fig = plt.figure()
ax = fig.gca(projection='3d')
X = np.arange(0, 10, 0.1)
Y = np.arange(0, 10, 0.1)
X, Y = np.meshgrid(X, Y)
Z = pot.transpose()
surf = ax.plot_surface(X, Y, Z, rstride=1, cstride=1, cmap=cm.coolwarm,
linewidth=0, antialiased=False)
ax.set_xlabel("x")
ax.set_ylabel("y")
print "layer1",(pot.transpose()[1,7],pot.transpose()[3,4],pot.transpose()[8,5],
pot.transpose()[9,7])
print "layer2",pot.transpose()[1,3],pot.transpose()[3,4]
print "layer3",pot.transpose()[1,1],pot.transpose()[3,1],pot.transpose()[7,4]
layer_1 = np.array([[5,5],[3,5]])
layer_2 = np.array([[1,3],[5,3],[7,3],[9,3]])
dip_pos_1 = np.array([2,4])
dip_angle_1 = 90
dip_pos_1_v = np.array([np.cos(np.deg2rad(dip_angle_1))*1,
np.sin(np.deg2rad(dip_angle_1))]) + dip_pos_1
dip_pos_2 = np.array([6,4])
dip_angle_2 = 90
dip_pos_2_v = np.array([np.cos(np.deg2rad(dip_angle_2))*1,
np.sin(np.deg2rad(dip_angle_2))]) + dip_pos_2
#print b([dip_pos_1,dip_pos_2], [dip_pos_1_v,dip_pos_2_v],13)
bb = b([dip_pos_1], [dip_pos_1_v], 15 )
sol = np.linalg.solve(aa,bb)
print sol
pot = np.zeros((20,20))
for i in range(20):
for j in range(20):
pot[i,j] = estimator([i/2.,j/2.],[dip_pos_1,dip_pos_2],
[layer_1,], sol, verbose = 0)
plt.arrow(dip_pos_1[0],dip_pos_1[1], dip_pos_1_v[0]-dip_pos_1[0],
dip_pos_1_v[1]-dip_pos_1[1], head_width = 0.2)
plt.arrow(dip_pos_2[0],dip_pos_2[1],dip_pos_2_v[0]-dip_pos_2[0],
dip_pos_2_v[1]-dip_pos_2[1], head_width = 0.2)
plt.plot(layer_1[:,0],layer_1[:,1], "o")
plt.plot(layer_2[:,0],layer_2[:,1], "o")
plt.plot(layer_1[:,0],layer_1[:,1], )
plt.plot(layer_2[:,0],layer_2[:,1], )
plt.contour(pot,20, extent = (0,10,0,10) )
plt.colorbar()
plt.xlim(0,10)
plt.ylim(0,10)
print dip_pos_1_v, dip_pos_2_v, layer_1
plt.arrow?
```
### Normal Universal cookriging
```
def G_f(dips,x):
dips = np.asarray(dips)
a = np.asarray(dips)
b = np.asarray(x)
# print a, a[:,0]
# print b,b[:,0]
Gx = b[0] - a[:,0]
Gy = b[1] -a[:,1]
G = np.hstack((Gx,Gy))
return G
def b(x, dips,n):
n -= len(dips)*2 # because x and y direction
G = G_f(dips,x)
b = np.hstack((G, np.zeros(n)))
return b,G
b([1,1],[dip_pos_1,dip_pos_2],13)
bb,g = b([1,1],[dip_pos_1,dip_pos_2],13)
len(bb)
sol = np.linalg.solve(aa,bb)
sol
dip_pos_1, dip_pos_2
z1 = dip_pos_1_v - dip_pos_1
z2 = dip_pos_2_v - dip_pos_2
print z1, z2
g
#=====================
# THE GRADIENTS
def h_f(dips, direct):
if direct == "x":
return np.abs(np.subtract.outer(dips[:,0],dips[:,0]))
if direct == "y":
return np.abs(np.subtract.outer(dips[:,1],dips[:,1]))
def C_G(dips, sig_z = 1., a = 6., nugget= 0.01):
dips = np.asarray(dips)
r = me.euclidean_distances(dips)
for i in "xy":
for j in "xy":
if j == "x":
h1 = h_f(dips, direct = i)
h2 = h_f(dips, direct = j)
# print h1,h2
C_G_row = (sig_z*h1*h2/a**2/r**2*
(1/r*cov_cubic_d1_f(r)-cov_cubic_d2_f(r)))
# print 1/r*cov_cubic_d1_f(r), cov_cubic_d2_f(r)
else:
h1 = h_f(dips, direct = i)
h2 = h_f(dips, direct = j)
C_G_row = np.hstack((C_G_row, (sig_z*h1*h2/a**2/r**2*
(1/r*cov_cubic_d1_f(r)-cov_cubic_d2_f(r)))))
if i == "x":
C_G = C_G_row
else:
C_G = np.vstack((C_G, C_G_row))
return np.nan_to_num(C_G)
```
## Estimator geomodeller (maybe)
```
def estimator(x, dips, layers, sol, sig_z = 1., a = 6., alpha = 1, verbose = 0):
x = np.asarray(x).reshape(1,-1)
dips = np.asarray(dips)
layers = np.asarray(layers)
n = 0
m = len(dips)
# print layers
# print x.reshape(1,-1), dips
r_i = me.euclidean_distances(dips,x)
hx = h_f_GI(dips, x, "x")
Cov_d1 = cov_cubic_d1_f(r_i)
KzGx = sol[:m] * np.squeeze(alpha * sig_z / a**2 * hx / r_i * Cov_d1)
hy = h_f_GI(dips, x, "y")
KzGy = sol[m:2*m] * np.squeeze(alpha * sig_z / a**2 * hy / r_i * Cov_d1)
for s in range(len(layers)):
n += len(layers[s][1:])
a = cov_cubic_layer(x, layers[s][1:])
b = cov_cubic_layer(x, layers[s][0].reshape(1,-1))
# print a,b
if s == 0:
L = np.array(sol[2*m:2*m+n]*(a-b))
else:
L = np.hstack((L,sol[2*m+n2:2*m+n]*(a-b)))
n2 = n
L = np.squeeze(L)
# print m,n
univ = (sol[2*m+n]*x[0,0]**2 + sol[2*m+n+1] * x[0,1]**2
+ sol[2*m+n+2]* x[0,0]*x[0,1]
+ sol[2*m+n+3] * x[0,0]
+ sol[2*m+n+4] * x[0,1])
if verbose != 0:
print KzGx, KzGy, L, univ
z_star = np.sum(KzGx)+np.sum(KzGy)+np.sum(L)+univ
return z_star
#========================================
#THE INTERACTION GRADIENTS/INTERFACES
def h_f_GI(dips, layers, direct):
if direct == "x":
return (np.subtract.outer(dips[:,0],layers[:,0]))
if direct == "y":
return (np.subtract.outer(dips[:,1],layers[:,1]))
def C_GI(dips,layers, sig_z = 1., a = 6., alpha = 14, verbose = 0):
dips = np.asarray(dips)
layers = np.asarray(layers)
for k in range(len(layers)):
for i in "xy":
r = me.euclidean_distances(dips,layers[k])
h1 = h_f_GI(dips,layers[k], i)
Cov_d1 = cov_cubic_d1_f(r)
if verbose != 0:
print "dips", dips
print "layers", layers
print "h1", h1, h1[:,0]
print ""
print "r", r, r[:,0]
print ""
print "Cov_d1", Cov_d1
if i == "x":
cov_1 = alpha * sig_z / a**2 * h1[:,0] / r[:,0] * Cov_d1[:,0]
cov_j = alpha * sig_z / a**2 * h1[:,1:] / r[:,1:] * Cov_d1[:,1:]
# C_GI_row = alpha * sig_z / a**2 * h1 / r * Cov_d1
#print "cov_j, cov_1", cov_j, cov_1.reshape(-1,1)
# pdb.set_trace()
C_GI_row = cov_j.transpose()-cov_1#.transpose()
else:
cov_1 = alpha * sig_z / a**2 * h1[:,0] / r[:,0] * Cov_d1[:,0]
cov_j = alpha * sig_z / a**2 * h1[:,1:] / r[:,1:] * Cov_d1[:,1:]
#C_GI_row = np.hstack((C_GI_row,
# alpha * sig_z / a**2 * h1 / r * Cov_d1))
#pdb.set_trace()
C_GI_row = np.hstack((C_GI_row, cov_j.transpose()-cov_1))
#.reshape(-1,1)))
if k==0:
C_GI = C_GI_row
else:
#pdb.set_trace()
C_GI = np.vstack((C_GI,C_GI_row))
return C_GI
```
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Forecasting with an RNN
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/examples/blob/master/courses/udacity_intro_to_tensorflow_for_deep_learning/l08c06_forecasting_with_rnn.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/examples/blob/master/courses/udacity_intro_to_tensorflow_for_deep_learning/l08c06_forecasting_with_rnn.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
</table>
## Setup
```
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
keras = tf.keras
def plot_series(time, series, format="-", start=0, end=None, label=None):
plt.plot(time[start:end], series[start:end], format, label=label)
plt.xlabel("Time")
plt.ylabel("Value")
if label:
plt.legend(fontsize=14)
plt.grid(True)
def trend(time, slope=0):
return slope * time
def seasonal_pattern(season_time):
"""Just an arbitrary pattern, you can change it if you wish"""
return np.where(season_time < 0.4,
np.cos(season_time * 2 * np.pi),
1 / np.exp(3 * season_time))
def seasonality(time, period, amplitude=1, phase=0):
"""Repeats the same pattern at each period"""
season_time = ((time + phase) % period) / period
return amplitude * seasonal_pattern(season_time)
def white_noise(time, noise_level=1, seed=None):
rnd = np.random.RandomState(seed)
return rnd.randn(len(time)) * noise_level
def window_dataset(series, window_size, batch_size=32,
shuffle_buffer=1000):
dataset = tf.data.Dataset.from_tensor_slices(series)
dataset = dataset.window(window_size + 1, shift=1, drop_remainder=True)
dataset = dataset.flat_map(lambda window: window.batch(window_size + 1))
dataset = dataset.shuffle(shuffle_buffer)
dataset = dataset.map(lambda window: (window[:-1], window[-1]))
dataset = dataset.batch(batch_size).prefetch(1)
return dataset
def model_forecast(model, series, window_size):
ds = tf.data.Dataset.from_tensor_slices(series)
ds = ds.window(window_size, shift=1, drop_remainder=True)
ds = ds.flat_map(lambda w: w.batch(window_size))
ds = ds.batch(32).prefetch(1)
forecast = model.predict(ds)
return forecast
time = np.arange(4 * 365 + 1)
slope = 0.05
baseline = 10
amplitude = 40
series = baseline + trend(time, slope) + seasonality(time, period=365, amplitude=amplitude)
noise_level = 5
noise = white_noise(time, noise_level, seed=42)
series += noise
plt.figure(figsize=(10, 6))
plot_series(time, series)
plt.show()
split_time = 1000
time_train = time[:split_time]
x_train = series[:split_time]
time_valid = time[split_time:]
x_valid = series[split_time:]
```
## Simple RNN Forecasting
```
keras.backend.clear_session()
tf.random.set_seed(42)
np.random.seed(42)
window_size = 30
train_set = window_dataset(x_train, window_size, batch_size=128)
model = keras.models.Sequential([
keras.layers.Lambda(lambda x: tf.expand_dims(x, axis=-1),
input_shape=[None]),
keras.layers.SimpleRNN(100, return_sequences=True),
keras.layers.SimpleRNN(100),
keras.layers.Dense(1),
keras.layers.Lambda(lambda x: x * 200.0) #scale
])
lr_schedule = keras.callbacks.LearningRateScheduler(
lambda epoch: 1e-7 * 10**(epoch / 20))
optimizer = keras.optimizers.SGD(lr=1e-7, momentum=0.9)
model.compile(loss=keras.losses.Huber(),
optimizer=optimizer,
metrics=["mae"])
history = model.fit(train_set, epochs=100, callbacks=[lr_schedule])
plt.semilogx(history.history["lr"], history.history["loss"])
plt.axis([1e-7, 1e-4, 0, 30])
keras.backend.clear_session()
tf.random.set_seed(42)
np.random.seed(42)
window_size = 30
train_set = window_dataset(x_train, window_size, batch_size=128)
valid_set = window_dataset(x_valid, window_size, batch_size=128)
model = keras.models.Sequential([
keras.layers.Lambda(lambda x: tf.expand_dims(x, axis=-1),
input_shape=[None]),
keras.layers.SimpleRNN(100, return_sequences=True),
keras.layers.SimpleRNN(100),
keras.layers.Dense(1),
keras.layers.Lambda(lambda x: x * 200.0)
])
optimizer = keras.optimizers.SGD(lr=1.5e-6, momentum=0.9)
model.compile(loss=keras.losses.Huber(),
optimizer=optimizer,
metrics=["mae"])
early_stopping = keras.callbacks.EarlyStopping(patience=50)
model_checkpoint = keras.callbacks.ModelCheckpoint(
"my_checkpoint", save_best_only=True)
model.fit(train_set, epochs=500,
validation_data=valid_set,
callbacks=[early_stopping, model_checkpoint])
model = keras.models.load_model("my_checkpoint")
rnn_forecast = model_forecast(
model,
series[split_time - window_size:-1],
window_size)[:, 0]
plt.figure(figsize=(10, 6))
plot_series(time_valid, x_valid)
plot_series(time_valid, rnn_forecast)
keras.metrics.mean_absolute_error(x_valid, rnn_forecast).numpy()
```
## Sequence-to-Sequence Forecasting
```
def seq2seq_window_dataset(series, window_size, batch_size=32,
shuffle_buffer=1000):
series = tf.expand_dims(series, axis=-1)
ds = tf.data.Dataset.from_tensor_slices(series)
ds = ds.window(window_size + 1, shift=1, drop_remainder=True)
ds = ds.flat_map(lambda w: w.batch(window_size + 1))
ds = ds.shuffle(shuffle_buffer)
ds = ds.map(lambda w: (w[:-1], w[1:]))
return ds.batch(batch_size).prefetch(1)
for X_batch, Y_batch in seq2seq_window_dataset(tf.range(10), 3,
batch_size=1):
print("X:", X_batch.numpy())
print("Y:", Y_batch.numpy())
keras.backend.clear_session()
tf.random.set_seed(42)
np.random.seed(42)
window_size = 30
train_set = seq2seq_window_dataset(x_train, window_size,
batch_size=128)
model = keras.models.Sequential([
keras.layers.SimpleRNN(100, return_sequences=True,
input_shape=[None, 1]),
keras.layers.SimpleRNN(100, return_sequences=True),
keras.layers.Dense(1),
keras.layers.Lambda(lambda x: x * 200)
])
lr_schedule = keras.callbacks.LearningRateScheduler(
lambda epoch: 1e-7 * 10**(epoch / 30))
optimizer = keras.optimizers.SGD(lr=1e-7, momentum=0.9)
model.compile(loss=keras.losses.Huber(),
optimizer=optimizer,
metrics=["mae"])
history = model.fit(train_set, epochs=100, callbacks=[lr_schedule])
plt.semilogx(history.history["lr"], history.history["loss"])
plt.axis([1e-7, 1e-4, 0, 30])
keras.backend.clear_session()
tf.random.set_seed(42)
np.random.seed(42)
window_size = 30
train_set = seq2seq_window_dataset(x_train, window_size,
batch_size=128)
valid_set = seq2seq_window_dataset(x_valid, window_size,
batch_size=128)
model = keras.models.Sequential([
keras.layers.SimpleRNN(100, return_sequences=True,
input_shape=[None, 1]),
keras.layers.SimpleRNN(100, return_sequences=True),
keras.layers.Dense(1),
keras.layers.Lambda(lambda x: x * 200.0)
])
optimizer = keras.optimizers.SGD(lr=1e-6, momentum=0.9)
model.compile(loss=keras.losses.Huber(),
optimizer=optimizer,
metrics=["mae"])
early_stopping = keras.callbacks.EarlyStopping(patience=10)
model.fit(train_set, epochs=500,
validation_data=valid_set,
callbacks=[early_stopping])
rnn_forecast = model_forecast(model, series[..., np.newaxis], window_size)
rnn_forecast = rnn_forecast[split_time - window_size:-1, -1, 0]
plt.figure(figsize=(10, 6))
plot_series(time_valid, x_valid)
plot_series(time_valid, rnn_forecast)
keras.metrics.mean_absolute_error(x_valid, rnn_forecast).numpy()
```
| github_jupyter |
## Feature Engineering for Customer Revenue Prediction
The purpose of this Kernel is to ingest the raw data, extract features, and engineer new ones to provide community with a ready-to-model flat dataset. The kernel is split into three part:
1. [Data Ingestion](#Data Ingestion)
2. [Feature Extraction](#Feature Extraction)
3. Feature Engineering - WIP
<a id='Data Ingestion'></a>
### 1. Data Ingestion
First, I import the libraries required to extraction.
```
# Import libraries
import pandas as pd
import numpy as np
import json
from pandas.io.json import json_normalize
```
Next, I use the function defined by [Julian Peller](https://www.kaggle.com/julian3833) to read ingest the raw data. Note that the function will automatically process nested JSON data elements and populate each into a separate column.
```
# Read in the raw traininig dataset
# Credit: https://www.kaggle.com/julian3833/1-quick-start-read-csv-and-flatten-json-fields
JSON_COLUMNS = ['device', 'geoNetwork', 'totals', 'trafficSource']
def load_df(csv_path='../train.csv'):
df = pd.read_csv(csv_path,
converters={column: json.loads for column in JSON_COLUMNS},
dtype={'fullVisitorId': 'str'},
parse_dates=['date']) # Note: added this line to Julian's code to parse dates on ingestion. It slows the process a bit
for column in JSON_COLUMNS:
column_as_df = json_normalize(df[column])
column_as_df.columns = [f"{column}.{subcolumn}" for subcolumn in column_as_df.columns]
df = df.drop(column, axis=1).merge(column_as_df, right_index=True, left_index=True)
return df
raw_train = load_df("../input/train.csv")
```
Similar to other Kernels, I drop the columns that are either all missing values or have just one unique value filled in (looking at you _trafficSource.campaign_ :) )
```
# Drop columns with just one value or all unknown
cols_to_drop = [col for col in raw_train.columns if raw_train[col].nunique() == 1]
raw_train.drop(columns = cols_to_drop, inplace=True)
# Drop campaign colum as it only has one non-null value
raw_train.drop(['trafficSource.campaign'], axis=1, inplace=True)
```
To make column names nicer, I'll rename a few to avoid the nesting dot notation. To do this I will split each column name on the dot (.) character and select the last string in the list to be the new column name.
```
# Rename long column names to be more concise
raw_train.rename(columns={col_name: col_name.split('.')[-1] for col_name in raw_train.columns}, inplace = True)
```
At the end, I am left with 30 columns, in a dataset of 903,653 rows. One of the columns is the dependent variable we are interested in predicting the log of accross all user visits - _transactionRevenue_.
```
print('Number of columns: {:}\nNumber of rows: {:}'.format(raw_train.shape[1], raw_train.shape[0]))
# Fill transactionRevenue with zeroes and convert its type to numeric
raw_train['transactionRevenue'].fillna(0, inplace=True)
raw_train['transactionRevenue'] = pd.to_numeric(raw_train['transactionRevenue'])
```
<a id='Feature Extraction'></a>
### 2. Feature Extraction & Cleaning
In this section I will constuct a dataset with the grain of sessionId, which in turn is just a concatenation of _fullVisitorId_ and _visitId_. The following table gives the names of the features I extract, the type of extraction (numeric or converting to dummy variables), and the source column. You can learn more about the meaning of each column [here](https://support.google.com/analytics/answer/3437719?hl=en).
| Feature | Type | Description | Source Column |
|:------|:------|:---|:---|
|Month | Numeric | Month of the visit. Values 1 to 12. | _date_ |
|Week | Numeric | Week of the year of the visit. Values 1 to 52. | _date_ |
|Weekday| Numeric | Day of the week of the visit. Values 1 to 7. | _date_|
|Hour| Numeric | Hour of day of the start of the visit. Values 0 to 24. | _visitStartTime_|
|Channel_X| Dummy | Three dummy columns indicating the channel the visit came in from. Values 0 or 1. | _channelGrouping_ |
|visitNumber| Numeric | The count of the current visit for this user. Values 1 to n. | _vistiNumber_ |
|Browser_X| Dummy | Dummy columns for each of the major browsers and additional column for "other". Values 0 or 1. | _browser_ |
|Device| Dummy | Three dummy columns indicating type of device. Values 'desktop', 'mobile', 'tablet'. | _deviceCategory_ |
|OS| Dummy | Dummy columns for each major operating system and additional column for "other". Values 0 or 1. |_operatingSystem_|
|SubContinent| Dummy | Dummy columns for each subcontient, combinig some of the smaller ones. Values 0 or 1. |_subContinent_|
|pageViews| Numeric | Number of pageviews that are generated by user thus far. Every row in the dataset adds 1 to user count. Values 1 to n. |_pageviews_|
|hits| Numeric| Superset of user activity count that also includes _pageviews_. Captures district interactions between user and webpage. Values 1 to n. |_hits_|
|Medium| Dummy | Dummy columns to indicate what type of marketing brought the user to the site. Values 0 or 1.|_medium_|
Note that the following columns I did not include in the feature extraction, and the reasons are as follows:
* _isMobile_ is already encoded as either mobile or tablet, so there is little need for a separate column.
* _sessionId_ is nothing more than a concatenation of _fullVisitorId_ and _visitId_
* _Year_ - there is only 12 months worth of data in both train and test datasets, so Month and Week are plenty to capture the timeframe.
* _city_ has almost 650 unique values, which is too many to encode as dummy variables. In the Feature engineering part I'll see what can be done with this information.
* _continent_ is a superset of _subContinent_ which I'm using as the base location variable. So, _continent_ is redundant.
* _country_ likely provides a good amount of information about the demographic of the user, but I'll tackle than in feature engineering section.
* _metro_ is rarely filled in and is highly correlated with _city_, so it's a good candidate to drop entirely.
* _networkDomain_ could be interesting from feature engineering perspective, but I'll drop it for now.
* _region_ can be through of as a "state" or "province", but I will again leave that for feature engineering.
* _adNetwork_ only appears to be filled in when _adContent_ is filled in. Since the vast majority of _adNetwork_ values are "Google Search", this becomes a redundant feature.
* _gclId_ seems to be an internal ID used by google for tracking purposes. I don't think it will be of any use for modeling.
* _page_ onlye appears to be filled in when _adContent_ is filled in. Vast majority of values are 1 (i.e. ad appeared on page 1). I will drop the column for now.
* _slot_ means either an ad on top of the screen or RHS (right hand side) ad. I'll drop it for now.
* _keyword_ may be useful in the feature engineering, but in its raw form it's too cumbersome to include.
* _referralPath_ requires some digging and feature engineering, but it will probably be a useful column to explore.
* _source_ same as above - requires some digging and feature engineering.
Phew, that took a while to writeup. Hopefully, it'll save you some work and you would have learned something in the process. Let's get "extracting"! I will use the existing dataframe as the base for pulling out the features listed in the table above. I will not drop the columns that would not be used for model training (e.g. date), but I'll add comented out code at the bottom of this Kernel to remove them if you wish.
Start with the date fields that are relatively straightforward.
```
# Get the month value from date
raw_train['Month'] = raw_train['date'].dt.month
# Get the week value from date
raw_train['Week'] = raw_train['date'].dt.week
# Get the weekday value from date
raw_train['Weekday'] = raw_train['date'].dt.weekday
# Get the hour value from visitStartTime
raw_train['Hour'] = pd.to_datetime(raw_train['visitStartTime'], unit='s').dt.hour
```
Next, let's dummify the _challenGrouping_ and _device_ variables. This is a grouping of the sources of web traffic that lead to the GStore and the device that the user is on. The dummifying operation is pretty straightforward here.
```
# Dummify challenGrouping into 8 separate binary columns
raw_train = pd.get_dummies(raw_train, columns = ['channelGrouping', 'deviceCategory'])
```
Who would have though that there are so many different browsers out there. I mean, have you ever heard of [Puffin](https://www.puffinbrowser.com/) or [Lunascape](https://www.lunascape.tv/)? For the purposes of training a model, I believe it will suffice to lump all of the little known browsers into one bucket called "Other" and call it a day. The major browsers I leave in their own dummy columns are Chrome, Safari, Firefox, IE, Edge, Android, Safari, Opera, UC Browser (marker for Asian market), Coc coc (marker for Vietnameese market).
```
# Group all little known broswers into "Other" bucket
raw_train.loc[~raw_train['browser'].isin(['Chrome', 'Safari', 'Firefox', 'Internet Explorer',
'Edge', 'Android Webview', 'Safari (in-app)', 'Opera Mini',
'Opera', 'UC Browser', 'Coc Coc']), ['browser']] = 'Other'
# Dummify browser into separage binary columns
raw_train = pd.get_dummies(raw_train, columns = ['browser'])
```
For operating system, again I will take just the top 7 values and set the rest to "Other".
```
# Group all less common operating systems into "Other" bucket, including where it's (not set)
raw_train.loc[~raw_train['operatingSystem'].isin(['Windows', 'Macintosh', 'Android', 'iOS',
'Linux', 'Chrome OS', 'Windows Phone']), ['operatingSystem']] = 'Other'
# Dummify operatingSytem into separate binary columns
raw_train = pd.get_dummies(raw_train, columns = ['operatingSystem'])
```
Similarly, I'll combine Polynesia, Micronesia, Melanesia into "Other", cutting down on a few unnecessary dummy columns.
```
# Group all less populated parts of the world into "Other" bucket, including where it's (not set)
raw_train.loc[raw_train['subContinent'].isin(['Polynesia', 'Micronesian Region',
'Melanesia', '(not set)']), ['subContinent']] = 'Other'
# Dummify subContinent into separate binary columns
raw_train = pd.get_dummies(raw_train, columns = ['subContinent'])
```
Finally, let's look at the marketing medium. Here we have values like CPM (cost per thousand impressions), CPC (cost per click), affiliate (from affiliate site), referral (more targeted link (e.g. share)), organic (user finds page themselves). I will turn each of these into dummy variables and combine (not set) and (none) into "Other" bucket.
```
# Group unknown marketing mediums into "Other" bucket
raw_train.loc[raw_train['medium'].isin(['(not set)', '(none)']), ['medium']] = 'Other'
# Dummify operatingSytem into separate binary columns
raw_train = pd.get_dummies(raw_train, columns = ['medium'])
```
With that the dataset grew quite a bit in its width: from 30 columns to 85. However, remember that there are some columns I won't be using at this time (they require further engineering). So, I'll leave you with the code that will drop these columns from the dataframe. That way there is a clean dataset to plug and plan into the model. Of course, normalization will need to be done on the variables and the log transform on the target variable
```
print('Number of columns: {:}'.format(raw_train.shape[1]))
# Drop columns that will not be used at this point in time
raw_train.drop(['date', 'isMobile', 'sessionId', 'visitStartTime',
'city', 'continent', 'country', 'metro', 'networkDomain',
'region', 'adContent', 'adNetworkType', 'gclId', 'page',
'slot', 'keyword', 'referralPath', 'source'], axis=1, inplace = True)
```
### 3. Feature Engineering - Work in Progress...
In this section I will derive more advanced features such as the following:
1. Hours/minutes/seconds since last visit
2. Combine continent and country into a useful set of dummy variables
3. Group adContnet into useful categories
4. Further explore keyword, medium, referral path, and source
| github_jupyter |
## Implementation of Salisman's Don't Overfit submission
From [Kaggle](http://www.kaggle.com/c/overfitting)
>In order to achieve this we have created a simulated data set with 200 variables and 20,000 cases. An ‘equation’ based on this data was created in order to generate a Target to be predicted. Given the all 20,000 cases, the problem is very easy to solve – but you only get given the Target value of 250 cases – the task is to build a model that gives the best predictions on the remaining 19,750 cases.
```
import gzip
import requests
import zipfile
url = "https://dl.dropbox.com/s/lnly9gw8pb1xhir/overfitting.zip"
results = requests.get(url)
import StringIO
z = zipfile.ZipFile(StringIO.StringIO(results.content))
# z.extractall()
z.extractall()
z.namelist()
d = z.open('overfitting.csv')
d.readline()
import numpy as np
M = np.fromstring(d.read(), sep=",")
len(d.read())
np.fromstring?
data = np.loadtxt("overfitting.csv", delimiter=",", skiprows=1)
print """
There are also 5 other fields,
case_id - 1 to 20,000, a unique identifier for each row
train - 1/0, this is a flag for the first 250 rows which are the training dataset
Target_Practice - we have provided all 20,000 Targets for this model, so you can develop your method completely off line.
Target_Leaderboard - only 250 Targets are provided. You submit your predictions for the remaining 19,750 to the Kaggle leaderboard.
Target_Evaluate - again only 250 Targets are provided. Those competitors who beat the 'benchmark' on the Leaderboard will be asked to make one further submission for the Evaluation model.
"""
data.shape
ix_training = data[:, 1] == 1
ix_testing = data[:, 1] == 0
training_data = data[ix_training, 5:]
testing_data = data[ix_testing, 5:]
training_labels = data[ix_training, 2]
testing_labels = data[ix_testing, 2]
print "training:", training_data.shape, training_labels.shape
print "testing: ", testing_data.shape, testing_labels.shape
```
## Develop Tim's model
He mentions that the X variables are from a Uniform distribution. Let's investigate this:
```
figsize(12, 4)
hist(training_data.flatten())
print training_data.shape[0] * training_data.shape[1]
```
looks pretty right
```
import pymc as pm
to_include = pm.Bernoulli("to_include", 0.5, size=200)
coef = pm.Uniform("coefs", 0, 1, size=200)
@pm.deterministic
def Z(coef=coef, to_include=to_include, data=training_data):
ym = np.dot(to_include * training_data, coef)
return ym - ym.mean()
@pm.deterministic
def T(z=Z):
return 0.45 * (np.sign(z) + 1.1)
obs = pm.Bernoulli("obs", T, value=training_labels, observed=True)
model = pm.Model([to_include, coef, Z, T, obs])
map_ = pm.MAP(model)
map_.fit()
mcmc = pm.MCMC(model)
mcmc.sample(100000, 90000, 1)
(np.round(T.value) == training_labels).mean()
t_trace = mcmc.trace("T")[:]
(np.round(t_trace[-500:-400, :]).mean(axis=0) == training_labels).mean()
t_mean = np.round(t_trace).mean(axis=1)
imshow(t_trace[-10000:, :], aspect="auto")
colorbar()
figsize(23, 8)
coef_trace = mcmc.trace("coefs")[:]
imshow(coef_trace[-10000:, :], aspect="auto", cmap=pyplot.cm.RdBu, interpolation="none")
include_trace = mcmc.trace("to_include")[:]
figsize(23, 8)
imshow(include_trace[-10000:, :], aspect="auto", interpolation="none")
```
| github_jupyter |
# Chapter 22
*Modeling and Simulation in Python*
Copyright 2021 Allen Downey
License: [Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International](https://creativecommons.org/licenses/by-nc-sa/4.0/)
```
# check if the libraries we need are installed
try:
import pint
except ImportError:
!pip install pint
import pint
try:
from modsim import *
except ImportError:
!pip install modsimpy
from modsim import *
```
### Vectors
A `Vector` object represents a vector quantity. In the context of mechanics, vector quantities include position, velocity, acceleration, and force, all of which might be in 2D or 3D.
You can define a `Vector` object without units, but if it represents a physical quantity, you will often want to attach units to it.
I'll start by grabbing the units we'll need.
```
m = UNITS.meter
s = UNITS.second
kg = UNITS.kilogram
```
Here's a two dimensional `Vector` in meters.
```
A = Vector(3, 4) * m
```
We can access the elements by name.
```
A.x
A.y
```
The magnitude is the length of the vector.
```
A.mag
```
The angle is the number of radians between the vector and the positive x axis.
```
A.angle
```
If we make another `Vector` with the same units,
```
B = Vector(1, 2) * m
```
We can add `Vector` objects like this
```
A + B
```
And subtract like this:
```
A - B
```
We can compute the Euclidean distance between two Vectors.
```
A.dist(B)
```
And the difference in angle
```
A.diff_angle(B)
```
If we are given the magnitude and angle of a vector, what we have is the representation of the vector in polar coordinates.
```
mag = A.mag
angle = A.angle
```
We can use `pol2cart` to convert from polar to Cartesian coordinates, and then use the Cartesian coordinates to make a `Vector` object.
In this example, the `Vector` we get should have the same components as `A`.
```
x, y = pol2cart(angle, mag)
Vector(x, y)
```
Another way to represent the direction of `A` is a unit vector, which is a vector with magnitude 1 that points in the same direction as `A`. You can compute a unit vector by dividing a vector by its magnitude:
```
A / A.mag
```
Or by using the `hat` function, so named because unit vectors are conventionally decorated with a hat, like this: $\hat{A}$:
```
A.hat()
```
**Exercise:** Create a `Vector` named `a_grav` that represents acceleration due to gravity, with x component 0 and y component $-9.8$ meters / second$^2$.
```
# Solution goes here
```
### Degrees and radians
Pint provides units to represent degree and radians.
```
degree = UNITS.degree
radian = UNITS.radian
```
If you have an angle in degrees,
```
angle = 45 * degree
angle
```
You can convert to radians.
```
angle_rad = angle.to(radian)
```
If it's already in radians, `to` does the right thing.
```
angle_rad.to(radian)
```
You can also convert from radians to degrees.
```
angle_rad.to(degree)
```
As an alterative, you can use `np.deg2rad`, which works with Pint quantities, but it also works with simple numbers and NumPy arrays:
```
np.deg2rad(angle)
```
**Exercise:** Create a `Vector` named `a_force` that represents acceleration due to a force of 0.5 Newton applied to an object with mass 0.3 kilograms, in a direction 45 degrees up from the positive x-axis.
Add `a_force` to `a_grav` from the previous exercise. If that addition succeeds, that means that the units are compatible. Confirm that the total acceleration seems to make sense.
```
# Solution goes here
# Solution goes here
```
### Baseball
Here's a `Params` object that contains parameters for the flight of a baseball.
```
t_end = 10 * s
dt = t_end / 100
params = Params(x = 0 * m,
y = 1 * m,
g = 9.8 * m/s**2,
mass = 145e-3 * kg,
diameter = 73e-3 * m,
rho = 1.2 * kg/m**3,
C_d = 0.33,
angle = 45 * degree,
velocity = 40 * m / s,
t_end=t_end, dt=dt)
```
And here's the function that uses the `Params` object to make a `System` object.
```
def make_system(params):
"""Make a system object.
params: Params object with angle, velocity, x, y,
diameter, duration, g, mass, rho, and C_d
returns: System object
"""
angle, velocity = params.angle, params.velocity
# convert angle to degrees
theta = np.deg2rad(angle)
# compute x and y components of velocity
vx, vy = pol2cart(theta, velocity)
# make the initial state
R = Vector(params.x, params.y)
V = Vector(vx, vy)
init = State(R=R, V=V)
# compute area from diameter
diameter = params.diameter
area = np.pi * (diameter/2)**2
return System(params, init=init, area=area)
```
Here's how we use it:
```
system = make_system(params)
```
Here's a function that computes drag force using vectors:
```
def drag_force(V, system):
"""Computes drag force in the opposite direction of `v`.
V: velocity Vector
system: System object with rho, C_d, area
returns: Vector drag force
"""
rho, C_d, area = system.rho, system.C_d, system.area
mag = rho * V.mag**2 * C_d * area / 2
direction = -V.hat()
f_drag = direction * mag
return f_drag
```
We can test it like this.
```
V_test = Vector(10, 10) * m/s
drag_force(V_test, system)
```
Here's the slope function that computes acceleration due to gravity and drag.
```
def slope_func(state, t, system):
"""Computes derivatives of the state variables.
state: State (x, y, x velocity, y velocity)
t: time
system: System object with g, rho, C_d, area, mass
returns: sequence (vx, vy, ax, ay)
"""
R, V = state
mass, g = system.mass, system.g
a_drag = drag_force(V, system) / mass
a_grav = Vector(0, -g)
A = a_grav + a_drag
return V, A
```
Always test the slope function with the initial conditions.
```
slope_func(system.init, 0, system)
```
We can use an event function to stop the simulation when the ball hits the ground:
```
def event_func(state, t, system):
"""Stop when the y coordinate is 0.
state: State object
t: time
system: System object
returns: y coordinate
"""
R, V = state
return R.y
event_func(system.init, 0, system)
```
Now we can call `run_ode_solver`
```
results, details = run_ode_solver(system, slope_func, events=event_func)
details
```
The final label tells us the flight time.
```
flight_time = get_last_label(results) * s
```
The final value of `x` tells us the how far the ball landed from home plate:
```
R_final = get_last_value(results.R)
x_dist = R_final.x
```
### Visualizing the results
The simplest way to visualize the results is to plot x and y as functions of time.
```
xs = results.R.extract('x')
ys = results.R.extract('y')
xs.plot()
ys.plot()
decorate(xlabel='Time (s)',
ylabel='Position (m)')
```
We can plot the velocities the same way.
```
vx = results.V.extract('x')
vy = results.V.extract('y')
vx.plot(label='vx')
vy.plot(label='vy')
decorate(xlabel='Time (s)',
ylabel='Velocity (m/s)')
```
The x velocity slows down due to drag.
The y velocity drops quickly while drag and gravity are in the same direction, then more slowly after the ball starts to fall.
Another way to visualize the results is to plot y versus x. The result is the trajectory of the ball through its plane of motion.
```
def plot_trajectory(results):
xs = results.R.extract('x')
ys = results.R.extract('y')
plot(xs, ys, color='C2', label='trajectory')
decorate(xlabel='x position (m)',
ylabel='y position (m)')
plot_trajectory(results)
```
### Animation
One of the best ways to visualize the results of a physical model is animation. If there are problems with the model, animation can make them apparent.
The ModSimPy library provides `animate`, which takes as parameters a `TimeSeries` and a draw function.
The draw function should take as parameters a `State` object and the time. It should draw a single frame of the animation.
Inside the draw function, you almost always have to call `set_xlim` and `set_ylim`. Otherwise `matplotlib` auto-scales the axes, which is usually not what you want.
```
xs = results.R.extract('x')
ys = results.R.extract('y')
def draw_func(state, t):
set_xlim(xs)
set_ylim(ys)
x, y = state.R
plot(x, y, 'bo')
decorate(xlabel='x position (m)',
ylabel='y position (m)')
animate(results, draw_func)
```
**Exercise:** Delete the lines that set the x and y axes (or [comment them out](https://en.wiktionary.org/wiki/comment_out)) and see what the animation does.
### Under the hood
`Vector` is a function that returns a `ModSimVector` object.
```
V = Vector(3, 4)
type(V)
```
A `ModSimVector` is a specialized kind of Pint `Quantity`.
```
isinstance(V, Quantity)
```
There's one gotcha you might run into with Vectors and Quantities. If you multiply a `ModSimVector` and a `Quantity`, you get a `ModSimVector`:
```
V1 = V * m
type(V1)
```
But if you multiply a `Quantity` and a `Vector`, you get a `Quantity`:
```
V2 = m * V
type(V2)
```
With a `ModSimVector` you can get the coordinates using dot notation, as well as `mag`, `mag2`, and `angle`:
```
V1.x, V1.y, V1.mag, V1.angle
```
With a `Quantity`, you can't. But you can use indexing to get the coordinates:
```
V2[0], V2[1]
```
And you can use vector functions to get the magnitude and angle.
```
vector_mag(V2), vector_angle(V2)
```
And often you can avoid the whole issue by doing the multiplication with the `ModSimVector` on the left.
### Exercises
**Exercise:** Run the simulation with and without air resistance. How wrong would we be if we ignored drag?
```
# Hint
system_no_drag = System(system, C_d=0)
# Solution goes here
# Solution goes here
# Solution goes here
# Solution goes here
# Solution goes here
```
**Exercise:** The baseball stadium in Denver, Colorado is 1,580 meters above sea level, where the density of air is about 1.0 kg / meter$^3$. How much farther would a ball hit with the same velocity and launch angle travel?
```
# Hint
system2 = System(system, rho=1.0*kg/m**3)
# Solution goes here
# Solution goes here
```
**Exercise:** The model so far is based on the assumption that coefficient of drag does not depend on velocity, but in reality it does. The following figure, from Adair, [*The Physics of Baseball*](https://books.google.com/books/about/The_Physics_of_Baseball.html?id=4xE4Ngpk_2EC), shows coefficient of drag as a function of velocity.
<img src="data/baseball_drag.png" width="400">
I used [an online graph digitizer](https://automeris.io/WebPlotDigitizer/) to extract the data and save it in a CSV file. Here's how we can read it:
```
import os
filename = 'baseball_drag.csv'
if not os.path.exists(filename):
!wget https://raw.githubusercontent.com/AllenDowney/ModSimPy/master/data/baseball_drag.csv
baseball_drag = pd.read_csv(filename)
mph = Quantity(baseball_drag['Velocity in mph'], UNITS.mph)
mps = mph.to(m/s)
baseball_drag.index = magnitude(mps)
baseball_drag.index.name = 'Velocity in meters per second'
baseball_drag
```
Modify the model to include the dependence of `C_d` on velocity, and see how much it affects the results. Hint: use `interpolate`.
```
# Solution goes here
# Solution goes here
C_d = drag_interp(43 * m / s)
# Solution goes here
# Solution goes here
# Solution goes here
# Solution goes here
# Solution goes here
# Solution goes here
# Solution goes here
# Solution goes here
```
| github_jupyter |
```
# Comparing fiTQun's results with the fully supervised ResNet-18 classifier on the varying position dataset
# Naming convention: first particle type is which file it is from, last particletype is what the hypothesis is
## Imports
import sys
import os
import time
import math
import random
import pdb
import h5py
# Add the path to the parent directory to augment search for module
par_dir = os.path.abspath(os.path.join(os.getcwd(), os.pardir))
if par_dir not in sys.path:
sys.path.append(par_dir)
# Plotting import
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# Import the utils for plotting the metrics
from plot_utils import plot_utils
from plot_utils import notebook_utils_2
from sklearn.metrics import roc_curve, auc
# Fix the colour scheme for each particle type
COLOR_DICT = {"gamma":"red", "e":"blue", "mu":"green"}
def moving_average(a, n=3) :
ret = np.cumsum(a, dtype=float)
ret[n:] = ret[n:] - ret[:-n]
return ret[n - 1:] / n
def plot_multiple_ROC(fprs, tprs, thresholds, label_0, label_1, lbound, ubound, interval):
min_energy = 0
max_energy = 1000
fig, ax = plt.subplots(figsize=(16,9),facecolor="w")
ax.tick_params(axis="both", labelsize=20)
model_colors = [np.random.rand(3,) for i in fprs]
for j in np.arange(len(fprs)):
fpr = fprs[j]
tpr = tprs[j]
threshold = thresholds[j]
roc_auc = auc(fpr, tpr)
inv_fpr = []
for i in fpr:
inv_fpr.append(1/i) if i != 0 else inv_fpr.append(1/1e-5)
tnr = 1. - fpr
# TNR vs TPR plot
ax.plot(tpr, inv_fpr, color=model_colors[j],
label=r"Interval ${1:0.3f}$: $\{0}$, AUC ${1:0.3f}$".format((j+1),label_0, roc_auc) if label_0 is not "e" else r"${0}$, AUC ${1:0.3f}$".format(label_0, roc_auc),
linewidth=1.0, marker=".", markersize=4.0, markerfacecolor=model_colors[j])
# Show coords of individual points near x = 0.2, 0.5, 0.8
todo = {0.2: True, 0.5: True, 0.8: True}
for xy in zip(tpr, inv_fpr, tnr):
xy = (round(xy[0], 4), round(xy[1], 4), round(xy[2], 4))
xy_plot = (round(xy[0], 4), round(xy[1], 4))
for point in todo.keys():
if xy[0] >= point and todo[point]:
#ax.annotate('(%s, %s, %s)' % xy, xy=xy_plot, textcoords='data', fontsize=18, bbox=dict(boxstyle="square", fc="w"))
todo[point] = False
ax.grid(True, which='both', color='grey')
xlabel = r"$\{0}$ signal efficiency".format(label_0) if label_0 is not "e" else r"${0}$ signal efficiency".format(label_0)
ylabel = r"$\{0}$ background rejection".format(label_1) if label_1 is not "e" else r"${0}$ background rejection".format(label_1)
ax.set_xlabel(xlabel, fontsize=20)
ax.set_ylabel(ylabel, fontsize=20)
ax.set_title(r"${0} \leq E < {1}$".format(round(lbound,2), round(ubound,2)), fontsize=20)
ax.legend(loc="upper right", prop={"size":20})
plt.margins(0.1)
plt.yscale("log")
plt.savefig(('/home/ttuinstr/VAE/debugging/ROC_' + str(interval) + '.png'), bbox_inches='tight')
plt.show()
plt.clf() # Clear the current figure
plt.close() # Close the opened window
return fpr, tpr, threshold, roc_auc
def plot_rej_energy(fprs, tprs, thresholds, label_0, label_1, lbound, ubound, interval, efficiency, bins):
min_energy = 0
max_energy = 1000
fig, ax = plt.subplots(figsize=(16,9),facecolor="w")
ax.tick_params(axis="both", labelsize=20)
model_colors = [np.random.rand(3,) for i in fprs]
eff_invfpr = np.array([])
for j in np.arange(len(fprs)):
fpr = fprs[j]
tpr = tprs[j]
threshold = thresholds[j]
roc_auc = auc(fpr, tpr)
inv_fpr = np.array([])
for i in fpr:
if i != 0:
inv_fpr = np.append(inv_fpr, (1/i))
else:
inv_fpr = np.append(inv_fpr, (1/1e-5))
tnr = 1. - fpr
eff_index = np.where(np.around(tpr, decimals=2) == 0.8)[0].astype(int)
eff_invfpr = np.append(eff_invfpr, inv_fpr[eff_index].mean())
# TNR vs Energy bin plot
label = 0
ax.bar(bins, height=eff_invfpr, width=interval, color="green", align='edge')
ax.legend()
ax.set_ylabel("Muon background rejection", fontsize=20)
plt.xlabel("Energy (MeV)", fontsize=20)
ax.set_title("Rejection vs. Energy Level at " + str(efficiency) + " Efficiency", fontsize=20)
plt.yscale("log")
plt.savefig(('/home/ttuinstr/VAE/debugging/RejectionEnergyBinning_' + str(efficiency) + "_" + str(interval) + '.png'), bbox_inches='tight')
plt.show()
plt.clf() # Clear the current figure
plt.close() # Close the opened window
return eff_invfpr, threshold, roc_auc
def plot_rej_pos(fprs, tprs, thresholds, label_0, label_1, lbound, ubound, interval, efficiency, bins):
min_pos = 0
max_pos = 720
fig, ax = plt.subplots(figsize=(16,9),facecolor="w")
ax.tick_params(axis="both", labelsize=20)
model_colors = [np.random.rand(3,) for i in fprs]
eff_invfpr = np.array([])
for j in np.arange(len(fprs)):
fpr = fprs[j]
tpr = tprs[j]
threshold = thresholds[j]
roc_auc = auc(fpr, tpr)
inv_fpr = np.array([])
for i in fpr:
if i != 0:
inv_fpr = np.append(inv_fpr, (1/i))
else:
inv_fpr = np.append(inv_fpr, (1/1e-5))
tnr = 1. - fpr
eff_index = np.where(np.around(tpr, decimals=2) == 0.8)[0].astype(int)
eff_invfpr = np.append(eff_invfpr, inv_fpr[eff_index].mean())
# TNR vs Energy bin plot
label = 0
ax.bar(bins, height=eff_invfpr, width=-interval, color="green", align='edge')
ax.legend()
ax.set_ylabel("Muon background rejection", fontsize=20)
plt.xlabel("Distance from Center of Tank (cm)", fontsize=20)
ax.set_title("Rejection vs. Distance from Center at " + str(efficiency) + " Efficiency", fontsize=20)
plt.yscale("log")
plt.savefig(('/home/ttuinstr/VAE/debugging/RejectionEnergyBinning_' + str(efficiency) + "_" + str(interval) + '.png'), bbox_inches='tight')
plt.show()
plt.clf() # Clear the current figure
plt.close() # Close the opened window
return eff_invfpr, threshold, roc_auc
# returns the max and min angle at which cherenkov radiation will hit the tank
# and the max and min distance of the cherenkov ring to the wall of the barrel
def find_bounds(pos, ang, energy):
# Input parameters:
# pos - array of position of particles
# ang - array polar and azimuth angles of particle
# label - array type of particle
# energy - array particle energy
# ***all parameters should have same size of first dimension
# This is (I believe) what John told me the max Cherenkov emission angle would be
max_ang = abs(np.arccos(1/(1.33)))*(1 + abs(pos[:,1]/max(pos[:,1]))*0.5)
theta = ang[:,1]
phi = ang[:,0]
# radius and height of barrel
r = 400
tank_height = 520
# position of particle in barrel
end = np.array([pos[:,0], pos[:,2]]).transpose()
# *********************
# Calculate intersection with the wall
# This is done for the particle as well as for where the left and right edges of the max Cherenkov ring
# will hit the wall of the tank (this can later be used to restrict for particles hitting the wall at an angle)
#
# 1) Calculate intersection of right edge of ring with wall
# a point along the direction vector outside of the barrel
start = end + 1000*(np.array([np.cos(theta + max_ang), np.sin(theta + max_ang)]).transpose())
# finding the intersection of particle with barrel
a = (end[:,0] - start[:,0])**2 + (end[:,1] - start[:,1])**2
b = 2*(end[:,0] - start[:,0])*(start[:,0]) + 2*(end[:,1] - start[:,1])*(start[:,1])
c = start[:,0]**2 + start[:,1]**2 - r**2
t = (-b - (b**2 - 4*a*c)**0.5)/(2*a)
intersection = np.array([(end[:,0]-start[:,0])*t,(end[:,1]-start[:,1])*t]).transpose() + start
# distance to wall
length = end - intersection
length1 = (length[:,0]**2 + length[:,1]**2)**0.5
# 2) Calculate intersection of left edge of ring with wall
# a point along the direction vector outside of the barrel
start = end + 1000*(np.array([np.cos(theta - max_ang), np.sin(theta - max_ang)]).transpose())
# finding intersection of particle with barrel
a = (end[:,0] - start[:,0])**2 + (end[:,1] - start[:,1])**2
b = 2*(end[:,0] - start[:,0])*(start[:,0]) + 2*(end[:,1] - start[:,1])*(start[:,1])
c = start[:,0]**2 + start[:,1]**2 - r**2
t = (-b - (b**2 - 4*a*c)**0.5)/(2*a)
intersection = np.array([(end[:,0]-start[:,0])*t,(end[:,1]-start[:,1])*t]).transpose() + start
# distance to wall
length = end - intersection
length2 = (length[:,0]**2 + length[:,1]**2)**0.5
# 3) Calculate intersection of particle with wall
# a point along the particle's direction vector outside of the barrel
start = end + 1000*(np.array([np.cos(theta), np.sin(theta)]).transpose())
# finding intersection of particle with barrel
a = (end[:,0] - start[:,0])**2 + (end[:,1] - start[:,1])**2
b = 2*(end[:,0] - start[:,0])*(start[:,0]) + 2*(end[:,1] - start[:,1])*(start[:,1])
c = start[:,0]**2 + start[:,1]**2 - r**2
t = (-b - (b**2 - 4*a*c)**0.5)/(2*a)
intersection = np.array([(end[:,0]-start[:,0])*t,(end[:,1]-start[:,1])*t]).transpose() + start
# distance to wall
length = end - intersection
length3 = (length[:,0]**2 + length[:,1]**2)**0.5
# find maximum distance to wall
length = np.maximum(np.maximum(length1,length2), length3)
# find upper and lower bound of angle where Cherenkov ring is contained in barrel
top_ang = math.pi/2 - np.arctan((tank_height - pos[:,2])/ length)
bot_ang = math.pi/2 + np.arctan(abs(-tank_height - pos[:,2])/length)
lb = top_ang + max_ang
ub = bot_ang - max_ang
# returning:
# - upper bound and lower bound of polar angle that will be within the barrel
# - minimum and maximum distance of the emission ring to the wall of the barrel
return np.array([lb, ub, np.minimum(np.minimum(length1,length2), length3), length]).transpose()
# Import test events from h5 file
filtered_index = "/fast_scratch/WatChMaL/data/IWCD_fulltank_300_pe_idxs.npz"
filtered_indices = np.load(filtered_index, allow_pickle=True)
test_filtered_indices = filtered_indices['test_idxs']
original_data_path = "/data/WatChMaL/data/IWCDmPMT_4pi_fulltank_9M.h5"
f = h5py.File(original_data_path, "r")
original_eventids = np.array(f['event_ids'])
original_rootfiles = np.array(f['root_files'])
original_energies = np.array(f['energies'])
original_positions = np.array(f['positions'])
original_angles = np.array(f['angles'])
filtered_eventids = original_eventids[test_filtered_indices]
filtered_rootfiles = original_rootfiles[test_filtered_indices]
filtered_energies = original_energies[test_filtered_indices]
filtered_positions = original_positions[test_filtered_indices]
filtered_angles = original_angles[test_filtered_indices]
# Map test events to fiTQun events
# First, separate event types
e_test_indices = np.load('/home/ttuinstr/VAE/debugging/test_indices_e.npz')
e_test_indices = e_test_indices['arr_0'].astype(int)
mu_test_indices = np.load('/home/ttuinstr/VAE/debugging/test_indices_mu.npz')
mu_test_indices = mu_test_indices['arr_0'].astype(int)
gamma_test_indices = np.load('/home/ttuinstr/VAE/debugging/test_indices_gamma.npz')
gamma_test_indices = gamma_test_indices['arr_0'].astype(int)
e_positions = filtered_positions[e_test_indices]
mu_positions = filtered_positions[mu_test_indices]
gamma_positions = filtered_positions[gamma_test_indices]
e_angles = filtered_angles[e_test_indices]
mu_angles = filtered_angles[mu_test_indices]
gamma_angles = filtered_angles[gamma_test_indices]
e_energies = filtered_energies[e_test_indices]
mu_energies = filtered_energies[mu_test_indices]
gamma_energies = filtered_energies[gamma_test_indices]
# Match events in event types to fiTQun results
e_map_indices = np.load('/home/ttuinstr/VAE/debugging/map_indices_e.npz')
e_map_indices = e_map_indices['arr_0'].astype(int)
mu_map_indices = np.load('/home/ttuinstr/VAE/debugging/map_indices_mu.npz')
mu_map_indices = mu_map_indices['arr_0'].astype(int)
gamma_map_indices = np.load('/home/ttuinstr/VAE/debugging/map_indices_gamma.npz')
gamma_map_indices = gamma_map_indices['arr_0'].astype(int)
e_positions = e_positions[e_map_indices]
mu_positions = mu_positions[mu_map_indices]
gamma_positions = gamma_positions[gamma_map_indices]
e_angles = e_angles[e_map_indices]
mu_angles = mu_angles[mu_map_indices]
gamma_angles = gamma_angles[gamma_map_indices]
e_energies = e_energies[e_map_indices]
mu_energies = mu_energies[mu_map_indices]
gamma_energies = gamma_energies[gamma_map_indices]
# File paths for fiTQun results
fiTQun_e_path = "/fast_scratch/WatChMaL/data/IWCDmPMT_4pi_fulltank_fiTQun_e-.npz"
fiTQun_mu_path = "/fast_scratch/WatChMaL/data/IWCDmPMT_4pi_fulltank_fiTQun_mu-.npz"
fiTQun_gamma_path = "/fast_scratch/WatChMaL/data/IWCDmPMT_4pi_fulltank_fiTQun_gamma.npz"
# Load fiTQun results
f_e = np.load(fiTQun_e_path, allow_pickle=True)
f_mu = np.load(fiTQun_mu_path, allow_pickle=True)
f_gamma = np.load(fiTQun_gamma_path, allow_pickle=True)
list(f_e.keys())
# Load the results
# Remove events with a non-zero flag (this filtering will be applied to the other results as well)
# * A non-zero flag value usually implies that either the reconstruction is known to have failed
# or the particle exited the tank and so would not be included in actual physics analysis
e_flag = np.array(f_e['flag'])
e_indices = np.where((e_flag[:,0] == 0) & (e_flag[:,1] == 0))
mu_flag = np.array(f_mu['flag'])
mu_indices = np.where((mu_flag[:,0] == 0) & (mu_flag[:,1] == 0))
gamma_flag = np.array(f_gamma['flag'])
gamma_indices = np.where((gamma_flag[:,0] == 0) & (gamma_flag[:,1] == 0))
# Get the filename for each event
e_file = np.array(f_e['filename'])
e_file = e_file[e_indices]
mu_file = np.array(f_mu['filename'])
mu_file = mu_file[mu_indices]
gamma_file = np.array(f_gamma['filename'])
gamma_file = gamma_file[gamma_indices]
# Get the event ID for each event
e_eventid = np.array(f_e['eventid'])
e_eventid = e_eventid[e_indices]
mu_eventid = np.array(f_mu['eventid'])
mu_eventid = mu_eventid[mu_indices]
gamma_eventid = np.array(f_gamma['eventid'])
gamma_eventid = gamma_eventid[gamma_indices]
# Get the nLL for each event
# The first nLL value is for electron hypothesis and second nLL is for the muon hypothesis
e_nLL = np.array(f_e['nLL'])
e_nLL = e_nLL[e_indices]
mu_nLL = np.array(f_mu['nLL'])
mu_nLL = mu_nLL[mu_indices]
gamma_nLL = np.array(f_gamma['nLL'])
gamma_nLL = gamma_nLL[gamma_indices]
# Get the position for each event
e_dir = np.array(f_e['direction'])
e_dir = e_dir[e_indices]
mu_dir = np.array(f_mu['direction'])
mu_dir = mu_dir[mu_indices]
gamma_dir = np.array(f_gamma['direction'])
gamma_dir = gamma_dir[gamma_indices]
# Get the direction for each event
e_pos = np.array(f_e['position'])
e_pos = e_pos[e_indices]
mu_pos = np.array(f_mu['position'])
mu_pos = mu_pos[mu_indices]
gamma_pos = np.array(f_gamma['position'])
gamma_pos = gamma_pos[gamma_indices]
# Get the momentum for each event
e_mom = np.array(f_e['momentum'])
e_mom = e_mom[e_indices]
mu_mom = np.array(f_mu['momentum'])
mu_mom = mu_mom[mu_indices]
gamma_mom = np.array(f_gamma['momentum'])
gamma_mom = gamma_mom[gamma_indices]
# Get the time for each event
e_time = np.array(f_e['time'])
e_time = e_time[e_indices]
mu_time = np.array(f_mu['time'])
mu_time = mu_time[mu_indices]
gamma_time = np.array(f_gamma['time'])
gamma_time = gamma_time[gamma_indices]
# Find the raw nLL differences
# nLL differences
e_nLL_diff_e = e_nLL[:,1] - e_nLL[:,0]
mu_nLL_diff_e = mu_nLL[:,1] - mu_nLL[:,0]
gamma_nLL_diff_e = gamma_nLL[:,1] - gamma_nLL[:,0]
e_nLL_diff_mu = e_nLL[:,0] - e_nLL[:,1]
mu_nLL_diff_mu = mu_nLL[:,0] - mu_nLL[:,1]
gamma_nLL_diff_mu = gamma_nLL[:,0] - gamma_nLL[:,1]
# labels
e_labels_mu = np.zeros(e_nLL_diff_mu.shape)
mu_labels_mu = np.ones(mu_nLL_diff_mu.shape)
e_labels_e = np.ones(e_nLL_diff_e.shape)
mu_labels_e = np.zeros(mu_nLL_diff_e.shape)
# concatenate labels and differences from mu and e events
diff_mu = np.concatenate((mu_nLL_diff_mu, e_nLL_diff_mu), axis=0)
labels_mu = np.concatenate((mu_labels_mu, e_labels_mu), axis=0)
diff_e = np.concatenate((mu_nLL_diff_e, e_nLL_diff_e), axis=0)
labels_e = np.concatenate((mu_labels_e, e_labels_e), axis=0)
# Find indices of events that hit within the barrel
e_contained_indices = np.arange(e_positions.shape[0])
bound = find_bounds(e_positions[:,0,:], e_angles[:,:],e_energies[:,0])
c = np.ma.masked_where(bound[e_contained_indices,2] < 200, e_contained_indices)
c = np.ma.masked_where(bound[e_contained_indices,2] > 400, c)
c = np.ma.masked_where((e_positions[e_contained_indices,0,0]**2 + e_positions[e_contained_indices,0,2]**2 + e_positions[e_contained_indices,0,1]**2)**0.5 > 400, c)
c = np.ma.masked_where(((e_angles[e_contained_indices,0] > bound[e_contained_indices,1]) | (e_angles[e_contained_indices,0] < bound[e_contained_indices,0])), c)
e_contained_indices = c.compressed()
mu_contained_indices = np.arange(mu_positions.shape[0])
bound = find_bounds(mu_positions[:,0,:], mu_angles[:,:], mu_energies[:,0])
c = np.ma.masked_where(bound[mu_contained_indices,2] < 200, mu_contained_indices)
c = np.ma.masked_where(bound[mu_contained_indices,2] > 400, c)
c = np.ma.masked_where((mu_positions[mu_contained_indices,0,0]**2 + mu_positions[mu_contained_indices,0,2]**2 + mu_positions[mu_contained_indices,0,1]**2)**0.5 > 400, c)
c = np.ma.masked_where(((mu_angles[mu_contained_indices,0] > bound[mu_contained_indices,1]) | (mu_angles[mu_contained_indices,0] < bound[mu_contained_indices,0])), c)
mu_contained_indices = c.compressed()
print(e_positions.shape)
print(e_pos.shape)
print(e_contained_indices.max())
# Use contained indices to filter events
e_pos = e_pos[e_contained_indices]
e_mom = e_mom[e_contained_indices]
mu_pos = mu_pos[mu_contained_indices]
mu_mom = mu_mom[mu_contained_indices]
e_positions = e_positions[e_contained_indices]
e_energies = e_energies[e_contained_indices]
mu_positions = mu_positions[mu_contained_indices]
mu_energies = mu_energies[mu_contained_indices]
# Take slices of events based on interval size for (reconstructed) position
diff_e_slices = []
labels_e_slices = []
e_nLL_diff_e_slices = []
mu_nLL_diff_e_slices = []
e_mom_slices = []
mu_mom_slices = []
tank_d = 7.42*100
tank_h = 10.42*100
interval = 50
print(int(int(tank_d/2)/interval))
for i in np.arange(int(int(tank_d/2)/interval)):
lb = i*interval
ub = (i+1)*interval
e_slice_indices = np.where(((e_pos[:,0,0]**2 + e_pos[:,0,1]**2 + e_pos[:,0,2]**2)**0.5 > lb) & ((e_pos[:,0,0]**2 + e_pos[:,0,1]**2 + e_pos[:,0,2]**2)**0.5 < ub))[0]
mu_slice_indices = np.where(((mu_pos[:,0,0]**2 + mu_pos[:,0,1]**2 + mu_pos[:,0,2]**2)**0.5 > lb) & ((mu_pos[:,0,0]**2 + mu_pos[:,0,1]**2 + mu_pos[:,0,2]**2)**0.5 < ub))[0]
e_nLL_slice = e_nLL[e_slice_indices,:]
mu_nLL_slice = mu_nLL[mu_slice_indices,:]
e_mom_slices.append(e_mom[e_slice_indices])
mu_mom_slices.append(mu_mom[mu_slice_indices])
e_nLL_diff_e_slices.append(e_nLL_slice[:,1] - e_nLL_slice[:,0])
mu_nLL_diff_e_slices.append(mu_nLL_slice[:,1] - mu_nLL_slice[:,0])
e_labels_e_slice = np.ones(e_nLL_diff_e_slices[i].shape)
mu_labels_e_slice = np.zeros(mu_nLL_diff_e_slices[i].shape)
diff_e_slices.append(np.concatenate((mu_nLL_diff_e_slices[i], e_nLL_diff_e_slices[i]), axis=0))
labels_e_slices.append(np.concatenate((mu_labels_e_slice, e_labels_e_slice), axis=0))
# Take slices of events based on interval size for (true) position
diff_e_slices = []
labels_e_slices = []
e_nLL_diff_e_slices = []
mu_nLL_diff_e_slices = []
e_mom_slices = []
mu_mom_slices = []
tank_d = 7.42*100
tank_h = 10.42*100
interval = 50
print(int(int(tank_d/2)/interval))
for i in np.arange(int(int(tank_d/2)/interval)):
lb = i*interval
ub = (i+1)*interval
e_slice_indices = np.where(((e_positions[:,0,0]**2 + e_positions[:,0,1]**2 + e_positions[:,0,2]**2)**0.5 > lb) & ((e_positions[:,0,0]**2 + e_positions[:,0,1]**2 + e_positions[:,0,2]**2)**0.5 < ub))[0]
mu_slice_indices = np.where(((mu_positions[:,0,0]**2 + mu_positions[:,0,1]**2 + mu_positions[:,0,2]**2)**0.5 > lb) & ((mu_positions[:,0,0]**2 + mu_pos[:,0,1]**2 + mu_positions[:,0,2]**2)**0.5 < ub))[0]
e_nLL_slice = e_nLL[e_slice_indices,:]
mu_nLL_slice = mu_nLL[mu_slice_indices,:]
e_mom_slices.append(e_mom[e_slice_indices])
mu_mom_slices.append(mu_mom[mu_slice_indices])
e_nLL_diff_e_slices.append(e_nLL_slice[:,1] - e_nLL_slice[:,0])
mu_nLL_diff_e_slices.append(mu_nLL_slice[:,1] - mu_nLL_slice[:,0])
e_labels_e_slice = np.ones(e_nLL_diff_e_slices[i].shape)
mu_labels_e_slice = np.zeros(mu_nLL_diff_e_slices[i].shape)
diff_e_slices.append(np.concatenate((mu_nLL_diff_e_slices[i], e_nLL_diff_e_slices[i]), axis=0))
labels_e_slices.append(np.concatenate((mu_labels_e_slice, e_labels_e_slice), axis=0))
# Make and plot ROC curves of event slices on same figure
#fpr, tpr, threshold = roc_curve(labels_e_slice, diff_e_slice)
#roc_metrics = plot_ROC(fpr, tpr, threshold, "e", "mu", lb, ub)
fprs = []
tprs = []
thresholds = []
for i in np.arange(len(diff_e_slices)):
fpr, tpr, threshold = roc_curve(labels_e_slices[i], diff_e_slices[i])
fprs.append(fpr)
tprs.append(tpr)
thresholds.append(threshold)
#roc_metrics = plot_multiple_ROC(fprs, tprs, thresholds, "e", "mu", 0, 1000, 50)
# Make and plot rejection vs. energy bin with fixed efficiency
efficiency = 0.8
fprs = []
tprs = []
thresholds = []
for i in np.arange(len(labels_e_slices)):
fpr, tpr, threshold = roc_curve(labels_e_slices[i], diff_e_slices[i])
fprs.append(fpr)
tprs.append(tpr)
thresholds.append(threshold)
# Set up the bins for the histogram
bins = []
for i in np.arange(len(fprs)):
bins.append(i*interval)
print(len(bins))
print(bins)
curve_metrics = plot_rej_pos(fprs, tprs, thresholds, "e", "mu", 0, 1000, 50, efficiency, bins)
# Take slices of events based on interval size for (reconstructed) energy
diff_e_slices = []
labels_e_slices = []
e_nLL_diff_e_slices = []
mu_nLL_diff_e_slices = []
e_mom_slices = []
mu_mom_slices = []
interval = 50
print(int(1000/interval))
for i in np.arange(int(1000/interval)):
lb = i*interval
ub = (i+1)*interval
e_nLL_slice = e_nLL[np.where(((e_mom[:,0] >lb) & (e_mom[:,0] <ub)))[0],:]
mu_nLL_slice = mu_nLL[np.where(((mu_mom[:,0] >lb) & (mu_mom[:,0] <ub)))[0],:]
e_mom_slices.append(e_mom[np.where(((e_mom[:,0] >lb) & (e_mom[:,0] <ub)))[0],:])
mu_mom_slices.append(mu_mom[np.where(((mu_mom[:,0] >lb) & (mu_mom[:,0] <ub)))[0],:])
e_nLL_diff_e_slices.append(e_nLL_slice[:,1] - e_nLL_slice[:,0])
mu_nLL_diff_e_slices.append(mu_nLL_slice[:,1] - mu_nLL_slice[:,0])
e_labels_e_slice = np.ones(e_nLL_diff_e_slices[i].shape)
mu_labels_e_slice = np.zeros(mu_nLL_diff_e_slices[i].shape)
diff_e_slices.append(np.concatenate((mu_nLL_diff_e_slices[i], e_nLL_diff_e_slices[i]), axis=0))
labels_e_slices.append(np.concatenate((mu_labels_e_slice, e_labels_e_slice), axis=0))
# Take slices of events based on interval size for (true) energy
diff_e_slices = []
labels_e_slices = []
e_nLL_diff_e_slices = []
mu_nLL_diff_e_slices = []
e_mom_slices = []
mu_mom_slices = []
interval = 50
print(int(1000/interval))
for i in np.arange(int(1000/interval)):
lb = i*interval
ub = (i+1)*interval
e_nLL_slice = e_nLL[np.where(((e_energies[:,0] >lb) & (e_energies[:,0] <ub)))[0],:]
mu_nLL_slice = mu_nLL[np.where(((mu_energies[:,0] >lb) & (mu_energies[:,0] <ub)))[0],:]
e_mom_slices.append(e_mom[np.where(((e_energies[:,0] >lb) & (e_energies[:,0] <ub)))[0],:])
mu_mom_slices.append(mu_mom[np.where(((mu_energies[:,0] >lb) & (mu_energies[:,0] <ub)))[0],:])
e_nLL_diff_e_slices.append(e_nLL_slice[:,1] - e_nLL_slice[:,0])
mu_nLL_diff_e_slices.append(mu_nLL_slice[:,1] - mu_nLL_slice[:,0])
e_labels_e_slice = np.ones(e_nLL_diff_e_slices[i].shape)
mu_labels_e_slice = np.zeros(mu_nLL_diff_e_slices[i].shape)
diff_e_slices.append(np.concatenate((mu_nLL_diff_e_slices[i], e_nLL_diff_e_slices[i]), axis=0))
labels_e_slices.append(np.concatenate((mu_labels_e_slice, e_labels_e_slice), axis=0))
# Make and plot rejection vs. energy bin with fixed efficiency
efficiency = 0.8
fprs = []
tprs = []
thresholds = []
for i in np.arange(len(labels_e_slices)):
fpr, tpr, threshold = roc_curve(labels_e_slices[i], diff_e_slices[i])
fprs.append(fpr)
tprs.append(tpr)
thresholds.append(threshold)
# Set up the bins for the histogram
bins = []
for i in np.arange(len(fprs)):
bins.append(i*interval)
print(len(bins))
print(bins)
curve_metrics = plot_rej_energy(fprs, tprs, thresholds, "e", "mu", 0, 1000, 50, efficiency, bins)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/mitpalorg/police-accountability-lab/blob/master/r_propub_2020.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
propub <- read.csv(url("https://raw.githubusercontent.com/mitpalorg/police-accountability-lab/master/allegations_20200726939.csv"))
head(propub)
a <- propub[,'unique_mos_id']
plot(ecdf(a), main=NA)
library(tidyverse)
b <- propub %>% filter(fado_type=='Force') %>%
select(unique_mos_id)
c <- data.frame(cbind( Freq=table(b), Cumul=cumsum(table(b)), relative=prop.table(table(b))))
library(ggplot2)
#ggplot(data = data.frame(c$Cumul)) + geom_point()
plot(ecdf(c$Cumul), main=NA)
c
d <- propub %>% filter(fado_type=='Force')
options(repr.plot.width = 12, repr.plot.height = 10)
g <- ggplot(d, aes(mos_age_incident, complainant_age_incident, col=complainant_ethnicity))
g + geom_jitter(width = .5, size=.5) +
labs(subtitle="Maturity? Complainant v. Officer Age",
y="Complainant",
x="Officer",
title="Jittered Points")
var <- d$board_disposition # the categorical data
## Prep data (nothing to change here)
nrows <- 10
df <- expand.grid(y = 1:nrows, x = 1:nrows)
categ_table <- round(table(var) * ((nrows*nrows)/(length(var))))
categ_table
df$category <- factor(rep(names(categ_table), categ_table))
# NOTE: if sum(categ_table) is not 100 (i.e. nrows^2), it will need adjustment to make the sum to 100.
## Plot
ggplot(df, aes(x = df$x, y = df$y, fill = category)) +
geom_tile(color = "black", size = 0.5) +
scale_x_continuous(expand = c(0, 0)) +
scale_y_continuous(expand = c(0, 0), trans = 'reverse') +
scale_fill_brewer(palette = "Set3") +
labs(title="Waffle Chart", subtitle="'Class' of vehicles",
caption="Source: mpg") +
theme(panel.border = element_rect(size = 2),
plot.title = element_text(size = rel(1.2)),
axis.text = element_blank(),
axis.title = element_blank(),
axis.ticks = element_blank(),
legend.title = element_blank(),
legend.position = "right")
library(plyr)
library(scales)
library(zoo)
df <- read.csv("https://raw.githubusercontent.com/selva86/datasets/master/yahoo.csv")
df$date <- as.Date(df$date) # format date
df <- df[df$year >= 2012, ] # filter reqd years
# Create Month Week
df$yearmonth <- as.yearmon(df$date)
df$yearmonthf <- factor(df$yearmonth)
df <- ddply(df,.(yearmonthf), transform, monthweek=1+week-min(week)) # compute week number of month
df <- df[, c("year", "yearmonthf", "monthf", "week", "monthweek", "weekdayf", "VIX.Close")]
head(df)
#> year yearmonthf monthf week monthweek weekdayf VIX.Close
#> 1 2012 Jan 2012 Jan 1 1 Tue 22.97
#> 2 2012 Jan 2012 Jan 1 1 Wed 22.22
#> 3 2012 Jan 2012 Jan 1 1 Thu 21.48
#> 4 2012 Jan 2012 Jan 1 1 Fri 20.63
#> 5 2012 Jan 2012 Jan 2 2 Mon 21.07
#> 6 2012 Jan 2012 Jan 2 2 Tue 20.69
# Plot
ggplot(df, aes(monthweek, weekdayf, fill = VIX.Close)) +
geom_tile(colour = "white") +
facet_grid(year~monthf) +
scale_fill_gradient(low="red", high="green") +
labs(x="Week of Month",
y="",
title = "Time-Series Calendar Heatmap",
subtitle="Yahoo Closing Price",
fill="Close")
install.packages("ggmap", repos = "http://cran.rstudio.com/")
devtools::install_github("hrbrmstr/ggalt")
install.packages("waffle", repos = "https://cinc.rud.is")
library(ggmap)
library(ggalt)
chennai <- geocode("Chennai") # get longitude and latitude
# Get the Map ----------------------------------------------
# Google Satellite Map
chennai_ggl_sat_map <- qmap("chennai", zoom=12, source = "google", maptype="satellite")
# Google Road Map
chennai_ggl_road_map <- qmap("chennai", zoom=12, source = "google", maptype="roadmap")
# Google Hybrid Map
chennai_ggl_hybrid_map <- qmap("chennai", zoom=12, source = "google", maptype="hybrid")
# Open Street Map
chennai_osm_map <- qmap("chennai", zoom=12, source = "osm")
# Get Coordinates for Chennai's Places ---------------------
chennai_places <- c("Kolathur",
"Washermanpet",
"Royapettah",
"Adyar",
"Guindy")
places_loc <- geocode(chennai_places) # get longitudes and latitudes
# Google Hybrid Map ----------------------------------------
chennai_ggl_hybrid_map + geom_point(aes(x=lon, y=lat),
data = places_loc,
alpha = 0.7,
size = 7,
color = "tomato") +
geom_encircle(aes(x=lon, y=lat),
data = places_loc, size = 2, color = "blue")
```
| github_jupyter |
## Exploratory analysis of the US Airport Dataset
This dataset contains data for 25 years[1995-2015] of flights between various US airports and metadata about these routes. Taken from Bureau of Transportation Statistics, United States Department of Transportation.
Let's see what can we make out of this!
```
%matplotlib inline
import networkx as nx
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import warnings
warnings.filterwarnings('ignore')
pass_air_data = pd.read_csv('datasets/passengers.csv')
```
In the `pass_air_data` dataframe we have the information of number of people that fly every year on a particular route on the list of airlines that fly that route.
```
pass_air_data.head()
# Create a MultiDiGraph from this dataset
passenger_graph = nx.from_pandas_edgelist(pass_air_data, source='ORIGIN', target='DEST', edge_attr=['YEAR', 'PASSENGERS', 'UNIQUE_CARRIER_NAME'], create_using=nx.MultiDiGraph())
```
### Cleveland to Chicago, how many people fly this route?
```
passenger_graph['CLE']['ORD'][25]
temp = [(i['YEAR'], i['PASSENGERS'])for i in dict(passenger_graph['CLE']['ORD']).values()]
x, y = zip(*temp)
plt.plot(x, y)
plt.show()
```
## Exercise
Find the busiest route in 1990 and in 2015 according to number of passengers, and plot the time series of number of passengers on these routes.
You can use the DataFrame instead of working with the network. It will be faster ;)
[5 mins]
```
temp = pass_air_data.groupby(['YEAR'])['PASSENGERS'].transform(max) == pass_air_data['PASSENGERS']
pass_air_data[temp][pass_air_data.YEAR.isin([1990, 2015])]
pass_air_data[(pass_air_data['ORIGIN'] == 'LAX') & (pass_air_data['DEST'] == 'HNL')].plot('YEAR', 'PASSENGERS')
pass_air_data[(pass_air_data['ORIGIN'] == 'LAX') & (pass_air_data['DEST'] == 'SFO')].plot('YEAR', 'PASSENGERS')
```
So let's have a look at the important nodes in this network, i.e. important airports in this network. We'll use pagerank, betweenness centrality and degree centrality.
```
# nx.pagerank(passenger_graph)
def year_network(G, year):
temp_g = nx.DiGraph()
for i in G.edges(data=True):
if i[2]['YEAR'] == year:
temp_g.add_edge(i[0], i[1], weight=i[2]['PASSENGERS'])
return temp_g
pass_2015 = year_network(passenger_graph, 2015)
len(pass_2015)
len(pass_2015.edges())
# Load in the GPS coordinates of all the airports
lat_long = pd.read_csv('datasets/GlobalAirportDatabase.txt', delimiter=':', header=None)
lat_long[lat_long[1].isin(list(pass_2015.nodes()))]
pos_dict = {}
for airport in lat_long[lat_long[1].isin(list(pass_2015.nodes()))].iterrows():
pos_dict[airport[1][1]] = (airport[1][15], airport[1][14])
pos_dict['AUS']
```
## Exercise
Using the position dictionary `pos_dict` create a plot of the airports, only the nodes not the edges.
- As we don't have coordinates for all the airports we have to create a subgraph first.
- Use `nx.subgraph(Graph, iterable of nodes)` to create the subgraph
- Use `nx.draw_networkx_nodes(G, pos)` to map the nodes.
or
- Just use a scatter plot :)
```
plt.figure(figsize=(20, 9))
G = nx.subgraph(pass_2015, pos_dict.keys())
nx.draw_networkx_nodes(G, pos=pos_dict, node_size=10, alpha=0.6, node_color='b')
# nx.draw_networkx_edges(G, pos=pos_dict, width=0.1, arrows=False)
plt.show()
plt.figure(figsize=(20, 9))
x = [i[0] for i in pos_dict.values()]
y = [i[1] for i in pos_dict.values()]
plt.scatter(x, y)
```
### What about degree distribution of this network?
```
plt.hist(list(nx.degree_centrality(pass_2015).values()))
plt.show()
```
Let's plot a log log plot to get a better overview of this.
```
d = {}
for i, j in dict(nx.degree(pass_2015)).items():
if j in d:
d[j] += 1
else:
d[j] = 1
x = np.log2(list((d.keys())))
y = np.log2(list(d.values()))
plt.scatter(x, y, alpha=0.4)
plt.show()
```
### Directed Graphs

```
G = nx.DiGraph()
G.add_edge(1, 2, weight=1)
# print(G.edges())
# G[1][2]
# G[2][1]
# G.is_directed()
# type(G)
G.add_edges_from([(1, 2), (3, 2), (4, 2), (5, 2), (6, 2), (7, 2)])
nx.draw_circular(G, with_labels=True)
G.in_degree()
nx.pagerank(G)
G.add_edge(5, 6)
nx.draw_circular(G, with_labels=True)
nx.pagerank(G)
G.add_edge(2, 8)
nx.draw_circular(G, with_labels=True)
nx.pagerank(G)
```
### Moving back to Airports
```
sorted(nx.pagerank(pass_2015, weight=None).items(), key=lambda x:x[1], reverse=True)[:10]
sorted(nx.betweenness_centrality(pass_2015).items(), key=lambda x:x[1], reverse=True)[0:10]
sorted(nx.degree_centrality(pass_2015).items(), key=lambda x:x[1], reverse=True)[0:10]
```
'ANC' is the airport code of Anchorage airport, a place in Alaska, and according to pagerank and betweenness centrality it is the most important airport in this network Isn't that weird? Thoughts?
related blog post: https://toreopsahl.com/2011/08/12/why-anchorage-is-not-that-important-binary-ties-and-sample-selection/
Let's look at weighted version, i.e taking into account the number of people flying to these places.
```
sorted(nx.betweenness_centrality(pass_2015, weight='weight').items(), key=lambda x:x[1], reverse=True)[0:10]
sorted(nx.pagerank(pass_2015, weight='weight').items(), key=lambda x:x[1], reverse=True)[0:10]
```
## How reachable is this network?
We calculate the average shortest path length of this network, it gives us an idea about the number of jumps we need to make around the network to go from one airport to any other airport in this network.
```
# nx.average_shortest_path_length(pass_2015)
```
Wait, What??? This network is not connected. That seems like a really stupid thing to do.
```
list(nx.weakly_connected_components(pass_2015))[1:]
```
### SPB, SSB, AIK anyone?
```
pass_air_data[(pass_air_data['YEAR'] == 2015) & (pass_air_data['ORIGIN'] == 'AIK')]
pass_2015.remove_nodes_from(['SPB', 'SSB', 'AIK'])
nx.is_weakly_connected(pass_2015)
nx.is_strongly_connected(pass_2015)
```
### Strongly vs weakly connected graphs.
```
G = nx.DiGraph()
G.add_edge(1, 2)
G.add_edge(2, 3)
G.add_edge(3, 1)
nx.draw(G)
G.add_edge(3, 4)
nx.draw(G)
nx.is_strongly_connected(G)
# list(nx.strongly_connected_components(pass_2015))
nx.strongly_connected_components(pass_2015)
pass_air_data[(pass_air_data['YEAR'] == 2015) & (pass_air_data['DEST'] == 'TSP')]
pass_2015_strong = pass_2015.subgraph(
max(nx.strongly_connected_components(pass_2015), key=len))
# (pass_2015_strong)
nx.average_shortest_path_length(pass_2015_strong)
```
#### Exercise! (Actually this is a game :D)
How can we decrease the avg shortest path length of this network?
Think of an effective way to add new edges to decrease the avg shortest path length.
Let's see if we can come up with a nice way to do this, and the one who gets the highest decrease wins!!!
The rules are simple:
- You can't add more than 2% of the current edges( ~500 edges)
[10 mins]
```
# unfreeze the graph
pass_2015_strong = nx.DiGraph(pass_2015_strong)
sort_degree = sorted(nx.degree_centrality(pass_2015_strong).items(), key=lambda x:x[1], reverse=True)
top_count = 0
for n, v in sort_degree:
count = 0
for node, val in sort_degree:
if node != n:
if node not in pass_2015_strong.adj[n]:
pass_2015_strong.add_edge(n, node)
count += 1
if count == 25:
break
top_count += 1
if top_count == 20:
break
nx.average_shortest_path_length(pass_2015_strong)
```
### What about airlines? Can we find airline specific reachability?
```
passenger_graph['JFK']['SFO'][25]
def str_to_list(a):
return a[1:-1].split(', ')
for i in str_to_list(passenger_graph['JFK']['SFO'][25]['UNIQUE_CARRIER_NAME']):
print(i)
%%time
for origin, dest in passenger_graph.edges():
for key in passenger_graph[origin][dest]:
passenger_graph[origin][dest][key]['airlines'] = str_to_list(passenger_graph[origin][dest][key]['UNIQUE_CARRIER_NAME'])
```
### Exercise
Play around with United Airlines network.
- Extract a network for United Airlines flights from the metagraph `passenger_graph` for the year 2015
- Make sure it's a weighted network, where weight is the number of passengers.
- Find the number of airports and connections in this network
- Find the most important airport, according to PageRank and degree centrality.
```
united_network = nx.DiGraph()
for origin, dest in passenger_graph.edges():
if 25 in passenger_graph[origin][dest]:
if "'United Air Lines Inc.'" in passenger_graph[origin][dest][25]['airlines']:
united_network.add_edge(origin, dest, weight=passenger_graph[origin][dest][25]['PASSENGERS'])
len(united_network)
len(united_network.edges())
sorted(nx.pagerank(united_network, weight='weight').items(), key=lambda x:x[1], reverse=True)[0:10]
sorted(nx.degree_centrality(united_network).items(), key=lambda x:x[1], reverse=True)[0:10]
```
### Exercise
We are in Cleveland so what should we do?
Obviously we will make a time series of number of passengers flying out of Cleveland with United Airlines over the years.
There are 2 ways of doing it.
- Create a new multidigraph specifically for this exercise.
OR
- exploit the `pass_air_data` dataframe.
```
pass_air_data[(pass_air_data.ORIGIN == 'CLE') &
(pass_air_data.UNIQUE_CARRIER_NAME.str.contains('United Air Lines Inc.'))
].groupby('YEAR')['PASSENGERS'].sum().plot()
```
| github_jupyter |
# Introduction to the pandapower control module
This tutorial introduces the pandapower controle module with the example of tap changer control. For this, we first load the MV oberrhein network that contains two 110/20 kV transformers:
```
# Importing necessary packages
import pandapower as pp
from pandapower.networks import mv_oberrhein
net = mv_oberrhein()
net.trafo
```
If we run a power flow, we can see the voltage at the low voltage side of the transformers:
```
pp.runpp(net)
net.res_trafo.vm_lv_pu
```
Both transformers include a tap changer with a range of -9 to +9, which are set to positions -2 and -3 respectively:
```
net.trafo["tap_pos"]
```
The tap position is constant within a power flow calculation. A controller can now be used to control the tap changer position depending on the bus voltage.
### Discrete Tap Control
The DiscreteTapControl from the pandapower control package receives a deadband of permissable voltage and uses the tap changer to keep the voltage within this voltage band. We define such a controller for the first transformer in the oberrhein network with a deadband of 0.99 to 1.01pu:
```
import pandapower.control as control
trafo_controller = control.DiscreteTapControl(net=net, tid=114, vm_lower_pu=0.99, vm_uppe_pur=1.01)
```
The initiated controller automatically registers in the net. It can be found in the controller table:
```
net.controller
```
We now run a controlled power flow by setting **run_control=True** within the runpp arguments and check the transformer voltage:
```
# running a control-loop
pp.runpp(net, run_control=True)
net.res_trafo.vm_lv_pu
```
The voltage at transformer 114 is now within the given range. If we checke the transformer table, we can see that the tap position of the first transformer as been changed from -2 to -1:
```
net.trafo["tap_pos"]
```
### Continous Tap Control
It is also possible to control transformer with a **ContiniousTapControl** strategy. Instead of a range, this type of controller is able to achieve an exact output voltage. For this it assumes tap positions as floating numbers. We define such a controller for the second transformer in the network:
```
trafo_controller = control.ContinuousTapControl(net=net, tid=142, vm_set_pu=0.98, tol=1e-6)
```
If we now run the result, the low voltage side of the second transformer is controlled to exactly 0.98 pu:
```
pp.runpp(net, run_control=True)
net.res_trafo.vm_lv_pu
```
The tap position is set to -0.07:
```
net.trafo["tap_pos"]
```
While this obviously would not possible in real transformers, it can be useful to assume continous taps in large scale studies to avoid big steps in the results.
| github_jupyter |
## Dependencies
```
import json, warnings, shutil
from tweet_utility_scripts import *
from tweet_utility_preprocess_roberta_scripts import *
from transformers import TFRobertaModel, RobertaConfig
from tokenizers import ByteLevelBPETokenizer
from tensorflow.keras.models import Model
from tensorflow.keras import optimizers, metrics, losses, layers
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint, LearningRateScheduler
SEED = 0
seed_everything(SEED)
warnings.filterwarnings("ignore")
```
# Load data
```
database_base_path = '/kaggle/input/tweet-dataset-split-roberta-base-96/'
k_fold = pd.read_csv(database_base_path + '5-fold.csv')
display(k_fold.head())
# Unzip files
!tar -xvf /kaggle/input/tweet-dataset-split-roberta-base-96/fold_1.tar.gz
!tar -xvf /kaggle/input/tweet-dataset-split-roberta-base-96/fold_2.tar.gz
!tar -xvf /kaggle/input/tweet-dataset-split-roberta-base-96/fold_3.tar.gz
!tar -xvf /kaggle/input/tweet-dataset-split-roberta-base-96/fold_4.tar.gz
!tar -xvf /kaggle/input/tweet-dataset-split-roberta-base-96/fold_5.tar.gz
```
# Model parameters
```
vocab_path = database_base_path + 'vocab.json'
merges_path = database_base_path + 'merges.txt'
base_path = '/kaggle/input/qa-transformers/roberta/'
config = {
"MAX_LEN": 96,
"BATCH_SIZE": 32,
"EPOCHS": 6,
"LEARNING_RATE": 3e-5,
"ES_PATIENCE": 1,
"question_size": 4,
"N_FOLDS": 5,
"base_model_path": base_path + 'roberta-base-tf_model.h5',
"config_path": base_path + 'roberta-base-config.json'
}
with open('config.json', 'w') as json_file:
json.dump(json.loads(json.dumps(config)), json_file)
```
# Tokenizer
```
tokenizer = ByteLevelBPETokenizer(vocab_file=vocab_path, merges_file=merges_path,
lowercase=True, add_prefix_space=True)
tokenizer.save('./')
```
## Learning rate schedule
```
LR_MIN = 1e-6
LR_MAX = config['LEARNING_RATE']
LR_EXP_DECAY = .5
@tf.function
def lrfn(epoch):
lr = LR_MAX * LR_EXP_DECAY**epoch
if lr < LR_MIN:
lr = LR_MIN
return lr
rng = [i for i in range(config['EPOCHS'])]
y = [lrfn(x) for x in rng]
fig, ax = plt.subplots(figsize=(20, 6))
plt.plot(rng, y)
print("Learning rate schedule: {:.3g} to {:.3g} to {:.3g}".format(y[0], max(y), y[-1]))
```
# Model
```
module_config = RobertaConfig.from_pretrained(config['config_path'], output_hidden_states=False)
def model_fn(MAX_LEN):
input_ids = layers.Input(shape=(MAX_LEN,), dtype=tf.int32, name='input_ids')
attention_mask = layers.Input(shape=(MAX_LEN,), dtype=tf.int32, name='attention_mask')
base_model = TFRobertaModel.from_pretrained(config['base_model_path'], config=module_config, name="base_model")
last_hidden_state, _ = base_model({'input_ids': input_ids, 'attention_mask': attention_mask})
x_start = layers.Dropout(.1)(last_hidden_state)
x_start = layers.Dense(1)(x_start)
x_start = layers.BatchNormalization()(x_start)
x_start = layers.Flatten()(x_start)
y_start = layers.Activation('softmax', name='y_start')(x_start)
x_end = layers.Dropout(.1)(last_hidden_state)
x_end = layers.Dense(1)(x_end)
x_end = layers.BatchNormalization()(x_end)
x_end = layers.Flatten()(x_end)
y_end = layers.Activation('softmax', name='y_end')(x_end)
model = Model(inputs=[input_ids, attention_mask], outputs=[y_start, y_end])
optimizer = optimizers.Adam(lr=config['LEARNING_RATE'])
model.compile(optimizer, loss={'y_start': losses.CategoricalCrossentropy(label_smoothing=0.2),
'y_end': losses.CategoricalCrossentropy(label_smoothing=0.2)},
metrics={'y_start': metrics.CategoricalAccuracy(),
'y_end': metrics.CategoricalAccuracy()})
return model
```
# Train
```
history_list = []
for n_fold in range(config['N_FOLDS']):
n_fold +=1
print('\nFOLD: %d' % (n_fold))
# Load data
base_data_path = 'fold_%d/' % (n_fold)
x_train = np.load(base_data_path + 'x_train.npy')
y_train = np.load(base_data_path + 'y_train.npy')
x_valid = np.load(base_data_path + 'x_valid.npy')
y_valid = np.load(base_data_path + 'y_valid.npy')
### Delete data dir
shutil.rmtree(base_data_path)
# Train model
model_path = 'model_fold_%d.h5' % (n_fold)
model = model_fn(config['MAX_LEN'])
es = EarlyStopping(monitor='val_loss', mode='min', patience=config['ES_PATIENCE'],
restore_best_weights=True, verbose=1)
checkpoint = ModelCheckpoint(model_path, monitor='val_loss', mode='min',
save_best_only=True, save_weights_only=True)
lr_schedule = LearningRateScheduler(lrfn)
history = model.fit(list(x_train), list(y_train),
validation_data=(list(x_valid), list(y_valid)),
batch_size=config['BATCH_SIZE'],
callbacks=[checkpoint, es, lr_schedule],
epochs=config['EPOCHS'],
verbose=1).history
history_list.append(history)
model.save_weights('last_' + model_path)
# Make predictions
train_preds = model.predict(list(x_train))
valid_preds = model.predict(list(x_valid))
k_fold.loc[k_fold['fold_%d' % (n_fold)] == 'train', 'start_fold_%d' % (n_fold)] = train_preds[0].argmax(axis=-1)
k_fold.loc[k_fold['fold_%d' % (n_fold)] == 'train', 'end_fold_%d' % (n_fold)] = train_preds[1].argmax(axis=-1)
k_fold.loc[k_fold['fold_%d' % (n_fold)] == 'validation', 'start_fold_%d' % (n_fold)] = valid_preds[0].argmax(axis=-1)
k_fold.loc[k_fold['fold_%d' % (n_fold)] == 'validation', 'end_fold_%d' % (n_fold)] = valid_preds[1].argmax(axis=-1)
k_fold['end_fold_%d' % (n_fold)] = k_fold['end_fold_%d' % (n_fold)].astype(int)
k_fold['start_fold_%d' % (n_fold)] = k_fold['start_fold_%d' % (n_fold)].astype(int)
k_fold['end_fold_%d' % (n_fold)].clip(0, k_fold['text_len'], inplace=True)
k_fold['start_fold_%d' % (n_fold)].clip(0, k_fold['end_fold_%d' % (n_fold)], inplace=True)
k_fold['prediction_fold_%d' % (n_fold)] = k_fold.apply(lambda x: decode(x['start_fold_%d' % (n_fold)], x['end_fold_%d' % (n_fold)], x['text'], config['question_size'], tokenizer), axis=1)
k_fold['prediction_fold_%d' % (n_fold)].fillna(k_fold["text"], inplace=True)
k_fold['jaccard_fold_%d' % (n_fold)] = k_fold.apply(lambda x: jaccard(x['selected_text'], x['prediction_fold_%d' % (n_fold)]), axis=1)
```
# Model loss graph
```
sns.set(style="whitegrid")
for n_fold in range(config['N_FOLDS']):
print('Fold: %d' % (n_fold+1))
plot_metrics(history_list[n_fold])
```
# Model evaluation
```
display(evaluate_model_kfold(k_fold, config['N_FOLDS']).style.applymap(color_map))
```
# Visualize predictions
```
display(k_fold[[c for c in k_fold.columns if not (c.startswith('textID') or
c.startswith('text_len') or
c.startswith('selected_text_len') or
c.startswith('text_wordCnt') or
c.startswith('selected_text_wordCnt') or
c.startswith('fold_') or
c.startswith('start_fold_') or
c.startswith('end_fold_'))]].head(15))
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
import matplotlib.pyplot as plt
import numpy as np
from mbench.intervention.efficacy import Converter_2022
from mbench.util import np_looper
import matplotlib.pyplot as plt
converter = Converter_2022(verbose=True)
x = np.linspace(0, 1, 300)
mortality_pyrethroid_hut_trail = [converter.mortality_bioassay_to_hut_trail(_) for _ in x]
plt.plot(x, mortality_pyrethroid_hut_trail)
plt.xlabel("mortality pyrethroid bioassay")
plt.ylabel("mortality pyrethroid bioassay")
mortality_pbo_hut_trail = [converter.mortality_hut_trail_from_pyrethroid_to_pbo(_) for _ in x]
plt.plot(x, mortality_pbo_hut_trail)
plt.xlabel("mortality pyrethroid hut trail")
plt.ylabel("mortality PBO hut trail")
p_of_deterred = [converter.ratio_of_mosquitoes_entering_hut_to_without_net(_) for _ in x]
plt.plot(x, p_of_deterred)
plt.ylabel("p of deterred")
plt.xlabel("mortality hut trail")
p_of_feed = [converter.proportion_of_mosquitoes_successfully_feed_upon_entering(_) for _ in x]
plt.plot(x, p_of_feed)
plt.ylabel("p of feed")
plt.xlabel("mortality hut trail")
p_entering_regular_ratio = [
converter.ratio_of_mosquitoes_entering_hut_to_without_net(_) for _ in mortality_pyrethroid_hut_trail
]
p_entering_pbo_ratio = [
converter.ratio_of_mosquitoes_entering_hut_to_without_net(_) for _ in mortality_pbo_hut_trail
]
plt.plot(x, p_entering_regular_ratio)
plt.plot(x, p_entering_pbo_ratio)
plt.xlabel("mortality pyrethroid bioassay")
plt.ylabel("ratio of mosquitoes entering hut to without net")
plt.legend(
labels=['regular', 'PBO'],
loc='best'
)
plt.hlines(1.0, 0, 1, colors='k', linestyles='dotted')
plt.hlines(0.0, 0, 1, colors='k', linestyles='dotted')
plt.show()
r10 = []
r11 = []
d10 = []
f10 = []
r20 = []
r21 = []
d20 =[]
f20 = []
for _ in np.nditer(x):
((_r10, _r11, _d10, _f10), (_r20, _r21, _d20, _f20), *others) = converter.bioassay_to_rds(_)
r10.append(_r10)
r11.append(_r11)
d10.append(_d10)
f10.append(_f10)
r20.append(_r20)
r21.append(_r21)
d20.append(_d20)
f20.append(_f20)
r10 = np.array(r10)
r11 = np.array(r11)
d10 = np.array(d10)
f10 = np.array(f10)
r20 = np.array(r20)
r21 = np.array(r21)
d20 = np.array(d20)
f20 = np.array(f20)
# ((r10, r11, d10, f10), (r20, r21, d20, f20)) = ((1,2,3,4),(1,2,3,4))
plt.stackplot(x, f10, d10, r10, labels=['f','d','r'])
plt.xlabel('mortality')
plt.ylabel('P')
plt.legend(['Blood fed','killed', 'repeating'])
plt.title('Relationships between Mortality in pyrethroid bioassay and efficacy in pyrethroid hut trial')
plt.stackplot(x, f20, d20, r20, labels=['f','d','r'])
plt.xlabel('mortality')
plt.ylabel('P')
plt.legend(['Blood fed','killed', 'repeating'])
plt.title('Relationships between Mortality in pyrethroid bioassay and efficacy in PBO hut trial')
converter.bioassay_to_rds(mortality_pyrethroid_bioassay=.8)
converter.proportion_of_mosquitoes_successfully_feed_upon_entering(0)
```
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import random
import time
sns.set()
def get_vocab(file, lower = False):
with open(file, 'r') as fopen:
data = fopen.read()
if lower:
data = data.lower()
vocab = list(set(data))
return data, vocab
def embed_to_onehot(data, vocab):
onehot = np.zeros((len(data), len(vocab)), dtype = np.float32)
for i in range(len(data)):
onehot[i, vocab.index(data[i])] = 1.0
return onehot
text, text_vocab = get_vocab('consumer.h', lower = False)
onehot = embed_to_onehot(text, text_vocab)
learning_rate = 0.0001
batch_size = 64
sequence_length = 12
epoch = 1000
num_layers = 2
size_layer = 128
possible_batch_id = range(len(text) - sequence_length - 1)
dimension = onehot.shape[1]
epsilon = 1e-8
beta1 = 0.9
beta2 = 0.999
U = np.random.randn(size_layer, dimension) / np.sqrt(size_layer)
U_g = np.zeros(U.shape)
U_g2 = np.zeros(U.shape)
Wf = np.random.randn(size_layer, size_layer) / np.sqrt(size_layer)
Wf_g = np.zeros(Wf.shape)
Wf_g2 = np.zeros(Wf.shape)
Wi = np.random.randn(size_layer, size_layer) / np.sqrt(size_layer)
Wi_g = np.zeros(Wi.shape)
Wi_g2 = np.zeros(Wi.shape)
Wc = np.random.randn(size_layer, size_layer) / np.sqrt(size_layer)
Wc_g = np.zeros(Wc.shape)
Wc_g2 = np.zeros(Wc.shape)
Wo = np.random.randn(size_layer, size_layer) / np.sqrt(size_layer)
Wo_g = np.zeros(Wo.shape)
Wo_g2 = np.zeros(Wo.shape)
V = np.random.randn(dimension, size_layer) / np.sqrt(dimension)
V_g = np.zeros(V.shape)
V_g2 = np.zeros(V.shape)
def tanh(x, grad=False):
if grad:
output = np.tanh(x)
return (1.0 - np.square(output))
else:
return np.tanh(x)
def sigmoid(x, grad=False):
if grad:
return sigmoid(x) * (1 - sigmoid(x))
else:
return 1 / (1 + np.exp(-x))
def softmax(x):
exp_scores = np.exp(x - np.max(x))
return exp_scores / (np.sum(exp_scores, axis=1, keepdims=True) + 1e-8)
def derivative_softmax_cross_entropy(x, y):
delta = softmax(x)
delta[range(X.shape[0]), y] -= 1
return delta
def forward_multiply_gate(w, x):
return np.dot(w, x)
def backward_multiply_gate(w, x, dz):
dW = np.dot(dz.T, x)
dx = np.dot(w.T, dz.T)
return dW, dx
def forward_add_gate(x1, x2):
return x1 + x2
def backward_add_gate(x1, x2, dz):
dx1 = dz * np.ones_like(x1)
dx2 = dz * np.ones_like(x2)
return dx1, dx2
def cross_entropy(Y_hat, Y, epsilon=1e-12):
Y_hat = np.clip(Y_hat, epsilon, 1. - epsilon)
N = Y_hat.shape[0]
return -np.sum(np.sum(Y * np.log(Y_hat+1e-9))) / N
def forward_recurrent(x, c_state, h_state, U, Wf, Wi, Wc, Wo, V):
mul_u = forward_multiply_gate(x, U.T)
mul_Wf = forward_multiply_gate(h_state, Wf.T)
add_Wf = forward_add_gate(mul_u, mul_Wf)
f = sigmoid(add_Wf)
mul_Wi = forward_multiply_gate(h_state, Wi.T)
add_Wi = forward_add_gate(mul_u, mul_Wi)
i = sigmoid(add_Wi)
mul_Wc = forward_multiply_gate(h_state, Wc.T)
add_Wc = forward_add_gate(mul_u, mul_Wc)
c_hat = tanh(add_Wc)
C = c_state * f + i * c_hat
mul_Wo = forward_multiply_gate(h_state, Wo.T)
add_Wo = forward_add_gate(mul_u, mul_Wo)
o = sigmoid(add_Wo)
h = o * tanh(C)
mul_v = forward_multiply_gate(h, V.T)
return (mul_u, mul_Wf, add_Wf, mul_Wi, add_Wi, mul_Wc, add_Wc, C, mul_Wo, add_Wo, h, mul_v, i, o, c_hat)
def backward_recurrent(x, c_state, h_state, U, Wf, Wi, Wc, Wo, V, d_mul_v, saved_graph):
mul_u, mul_Wf, add_Wf, mul_Wi, add_Wi, mul_Wc, add_Wc, C, mul_Wo, add_Wo, h, mul_v, i, o, c_hat = saved_graph
dV, dh = backward_multiply_gate(V, h, d_mul_v)
dC = tanh(C, True) * o * dh.T
do = tanh(C) * dh.T
dadd_Wo = sigmoid(add_Wo, True) * do
dmul_u1, dmul_Wo = backward_add_gate(mul_u, mul_Wo, dadd_Wo)
dWo, dprev_state = backward_multiply_gate(Wo, h_state, dmul_Wo)
dc_hat = dC * i
dadd_Wc = tanh(add_Wc, True) * dc_hat
dmul_u2, dmul_Wc = backward_add_gate(mul_u, mul_Wc, dadd_Wc)
dWc, dprev_state = backward_multiply_gate(Wc, h_state, dmul_Wc)
di = dC * c_hat
dadd_Wi = sigmoid(add_Wi, True) * di
dmul_u3, dmul_Wi = backward_add_gate(mul_u, mul_Wi, dadd_Wi)
dWi, dprev_state = backward_multiply_gate(Wi, h_state, dmul_Wi)
df = dC * c_state
dadd_Wf = sigmoid(add_Wf, True) * df
dmul_u4, dmul_Wf = backward_add_gate(mul_u, mul_Wf, dadd_Wf)
dWf, dprev_state = backward_multiply_gate(Wf, h_state, dmul_Wf)
dU, dx = backward_multiply_gate(U, x, dmul_u4)
return (dU, dWf, dWi, dWc, dWo, dV)
for i in range(epoch):
batch_x = np.zeros((batch_size, sequence_length, dimension))
batch_y = np.zeros((batch_size, sequence_length, dimension))
batch_id = random.sample(possible_batch_id, batch_size)
prev_c = np.zeros((batch_size, size_layer))
prev_h = np.zeros((batch_size, size_layer))
for n in range(sequence_length):
id1 = [k + n for k in batch_id]
id2 = [k + n + 1 for k in batch_id]
batch_x[:,n,:] = onehot[id1, :]
batch_y[:,n,:] = onehot[id2, :]
layers = []
out_logits = np.zeros((batch_size, sequence_length, dimension))
for n in range(sequence_length):
layers.append(forward_recurrent(batch_x[:,n,:], prev_c, prev_h, U, Wf, Wi, Wc, Wo, V))
prev_c = layers[-1][7]
prev_h = layers[-1][10]
out_logits[:, n, :] = layers[-1][-4]
probs = softmax(out_logits.reshape((-1, dimension)))
y = np.argmax(batch_y.reshape((-1, dimension)),axis=1)
accuracy = np.mean(np.argmax(probs,axis=1) == y)
loss = cross_entropy(probs, batch_y.reshape((-1, dimension)))
delta = probs
delta[range(y.shape[0]), y] -= 1
delta = delta.reshape((batch_size, sequence_length, dimension))
dU = np.zeros(U.shape)
dV = np.zeros(V.shape)
dWf = np.zeros(Wf.shape)
dWi = np.zeros(Wi.shape)
dWc = np.zeros(Wc.shape)
dWo = np.zeros(Wo.shape)
prev_c = np.zeros((batch_size, size_layer))
prev_h = np.zeros((batch_size, size_layer))
for n in range(sequence_length):
d_mul_v = delta[:, n, :]
dU_t, dWf_t, dWi_t, dWc_t, dWo_t, dV_t = backward_recurrent(batch_x[:,n,:], prev_c, prev_h, U, Wf, Wi,
Wc, Wo, V, d_mul_v, layers[n])
prev_c = layers[n][7]
prev_h = layers[n][10]
dU += dU_t
dV += dV_t
dWf += dWf_t
dWi += dWi_t
dWc += dWc_t
dWo += dWo_t
U_g += beta1 * U_g + (1-beta1) * dU
g_hat = U_g / (1-beta1)
U_g2 += beta2 * U_g2 + (1-beta2) * np.square(dU)
g2_hat = U_g2 / (1-beta2)
U += -learning_rate * g_hat / np.sqrt(g2_hat + epsilon)
V_g += beta1 * V_g + (1-beta1) * dV
g_hat = V_g / (1-beta1)
V_g2 += beta2 * V_g2 + (1-beta2) * np.square(dV)
g2_hat = V_g2 / (1-beta2)
V += -learning_rate * g_hat / np.sqrt(g2_hat + epsilon)
Wf_g += beta1 * Wf_g + (1-beta1) * dWf
g_hat = Wf_g / (1-beta1)
Wf_g2 += beta2 * Wf_g2 + (1-beta2) * np.square(dWf)
g2_hat = Wf_g2 / (1-beta2)
Wf += -learning_rate * g_hat / np.sqrt(g2_hat + epsilon)
Wi_g += beta1 * Wi_g + (1-beta1) * dWi
g_hat = Wi_g / (1-beta1)
Wi_g2 += beta2 * Wf_g2 + (1-beta2) * np.square(dWi)
g2_hat = Wi_g2 / (1-beta2)
Wi += -learning_rate * g_hat / np.sqrt(g2_hat + epsilon)
Wc_g += beta1 * Wc_g + (1-beta1) * dWc
g_hat = Wc_g / (1-beta1)
Wc_g2 += beta2 * Wc_g2 + (1-beta2) * np.square(dWc)
g2_hat = Wc_g2 / (1-beta2)
Wc += -learning_rate * g_hat / np.sqrt(g2_hat + epsilon)
Wo_g += beta1 * Wo_g + (1-beta1) * dWo
g_hat = Wo_g / (1-beta1)
Wo_g2 += beta2 * Wo_g2 + (1-beta2) * np.square(dWo)
g2_hat = Wo_g2 / (1-beta2)
Wo += -learning_rate * g_hat / np.sqrt(g2_hat + epsilon)
if (i+1) % 50 == 0:
print('epoch %d, loss %f, accuracy %f'%(i+1, loss, accuracy))
```
| github_jupyter |
```
# Run this script only when you haven't downloaded data yet.
# !./prepare_input.sh
import pandas as pd
import numpy as np
import lib.Model as md
import lib.merge_spray as merge_spray
raw_train_data = pd.read_csv('./input/train.csv', parse_dates=['Date'])
raw_test_data = pd.read_csv('./input/test.csv', parse_dates=['Date'])
weather_data = pd.read_csv('./input/weather.csv')
spray_data = pd.read_csv('./input/spray.csv', parse_dates=['Date'])
def dummy_species(data): # TODO 高速化
# 蚊の種類をone-hot形式にする
sp_keys = {'CULEX ERRATICUS' , 'CULEX PIPIENS', 'CULEX RESTUANS',
'CULEX SALINARIUS', 'CULEX TARSALIS', 'CULEX TERRITANS' , 'CULEX PIPIENS/RESTUANS'}
sp_map = {'CULEX ERRATICUS' : np.array([1, 0, 0, 0, 0, 0]),
'CULEX PIPIENS' : np.array([0, 1, 0, 0 , 0, 0]),
'CULEX RESTUANS' : np.array([0, 0, 1, 0, 0, 0]),
'CULEX SALINARIUS' : np.array([0, 0, 0, 1, 0, 0]),
'CULEX TARSALIS' : np.array([0, 0, 0, 0, 1, 0]),
'CULEX TERRITANS' : np.array([0, 0, 0, 0, 0, 1]),
'CULEX PIPIENS/RESTUANS' : np.array([0, 1, 1, 0, 0, 0])}
dummies = np.empty((0, len(sp_keys)-1), int)
for i in data:
if i not in sp_keys:
dummies = np.append(dummies, [np.zeros(len(sp_keys)-1)], axis=0)
else:
dummies = np.append(dummies, [np.array(sp_map[i])], axis=0)
return pd.DataFrame(dummies)
def dummy_trap(data): # TODO 高速化
# trapの種類をone-hot形式にする
trap_keys = list(set(raw_train_data['Trap']))
trap_dummy = pd.get_dummies(trap_keys)
dummies = np.empty((0, len(trap_keys)), int)
for i in data:
if i not in trap_keys:
dummies = np.append(dummies, [np.zeros(len(trap_keys))], axis=0)
else:
dummies = np.append(dummies, [np.array(trap_dummy[i])], axis=0)
return pd.DataFrame(dummies)
def preprocessing(data, weather, spray):
# pandasのデータで前処理をしてnumpy配列を返す
# 天気情報の前処理
formated_weather = weather.drop_duplicates(subset='Date').drop(['Station', 'CodeSum', 'Depth', 'Water1'], axis=1)
formated_weather = formated_weather.convert_objects(convert_numeric=True).fillna(0) # TODO 欠損値の扱い
# TODO スプレーデータの前処理
data = data.assign(ClosestSprayKmIn2Days=lambda df: merge_spray.min_dist(df, spray))
# メインデータの前処理
#if 'NumMosquitos' in data: # TODO 重複データの扱い
# data = data.drop_duplicates(subset=['Date', 'Species', 'Latitude', 'Longitude', 'Trap'])
species = dummy_species(data['Species'])
trap = dummy_trap(data['Trap'])
merged_data = pd.merge(data, formated_weather, on='Date')
merged_data = pd.merge(merged_data, species, right_index=True, left_index=True)
merged_data = pd.merge(merged_data, trap, right_index=True, left_index=True)
if 'NumMosquitos' in merged_data:
preprocessed_data = merged_data.drop(['Date', 'Address', 'Species', 'Block', 'Street', 'Trap', 'AddressNumberAndStreet', 'AddressAccuracy', 'NumMosquitos', 'WnvPresent'], axis=1)
return np.array(preprocessed_data), np.array(data['NumMosquitos']), np.array(data['WnvPresent'])
else:
preprocessed_data = merged_data.drop(['Id','Date', 'Address', 'Species', 'Block', 'Street', 'Trap', 'AddressNumberAndStreet', 'AddressAccuracy'], axis=1)
return np.array(preprocessed_data), np.array([]), np.array([])
print("Make training data")
train_data, num_mosquitos, label = preprocessing(raw_train_data, weather_data, spray_data)
print("Make test data")
test_data, tmp1, tmp2 = preprocessing(raw_test_data, weather_data, spray_data)
# SVM
model_svm = md.SVM(optimization=True)
model_svm.fit(train_data, label)
result_svm = model_svm.predict(test_data)
# RandomForest
model_rf = md.RandomForest()
model_rf.fit(train_data, label)
result_rf = model_rf.predict(test_data)
# LightGBM
model_lgbm = md.LightGBM()
model_lgbm.fit(train_data, label)
result_lgbm = model_lgbm.predict(test_data)
# GLM
model_glm = md.GLM()
model_glm.fit(train_data, label, num_mosquitos)
result_glm = model_glm.predict(test_data)
submitData_svm = pd.DataFrame.from_dict({"Id": raw_test_data['Id'], "WnvPresent": result_svm})
submitData_rf = pd.DataFrame.from_dict({"Id": raw_test_data['Id'], "WnvPresent": result_rf})
submitData_lgbm = pd.DataFrame.from_dict({"Id": raw_test_data['Id'], "WnvPresent": result_lgbm})
submitData_glm = pd.DataFrame.from_dict({"Id": raw_test_data['Id'], "WnvPresent": result_glm})
submitData_svm.to_csv('submission_svm.csv', index=0)
submitData_rf.to_csv('submission_rf.csv', index=0)
submitData_lgbm.to_csv('submission_lgbm.csv', index=0)
submitData_glm.to_csv('submission_glm.csv', index=0)
"""
private public
svm 0.515 0.510
rf 0.670 0.713
lgbm 0.715 0.73
glm 0.474 0.488
"""
```
| github_jupyter |
# 1章 Python入門
PyTorchを使ったディープラーニング・プログラミングで重要になる概念だけを抜き出して説明する
```
# 必要ライブラリの導入
!pip install japanize_matplotlib | tail -n 1
# 必要ライブラリのインポート
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import japanize_matplotlib
# warning表示off
import warnings
warnings.simplefilter('ignore')
# デフォルトフォントサイズ変更
plt.rcParams['font.size'] = 14
# デフォルトグラフサイズ変更
plt.rcParams['figure.figsize'] = (6,6)
# デフォルトで方眼表示ON
plt.rcParams['axes.grid'] = True
# numpyの表示桁数設定
np.set_printoptions(suppress=True, precision=5)
```
## 1.2 コンテナ変数にご用心
Pythonでは、変数は単に実際のデータ構造へのポインタに過ぎない。
Numpy配列などでは、このことを意識しないと思わぬ結果を招く場合がある。
### NumPy変数間
```
# Numpy配列 x1 を定義
x = np.array([5, 7, 9])
# 変数yにxを代入する
# このとき、実体は共通なまま
y = x
# 結果確認
print(x)
print(y)
# ここでxの特定の要素の値を変更する
x[1] = -1
# すると、yも連動して値が変わる
print(x)
print(y)
# yも同時に変化して困る場合は、代入時にcopy関数を利用する
x = np.array([5, 7, 9])
y = x.copy()
# すると、xの特定の要素値の変更がyに影響しなくなる
x[1] = -1
print(x)
print(y)
```
### テンソルとNumPy間
```
import torch
# x1: shape=[5] となるすべて値が1テンソル
x1 = torch.ones(5)
# 結果確認
print(x1)
# x2 x1から生成したNumPy
x2 = x1.data.numpy()
# 結果確認
print(x2)
# x1の値を変更
x1[1] = -1
# 連動してx2の値も変わる
print(x1)
print(x2)
# 安全な方法
# x1 テンソル
x1 = torch.ones(5)
# x2 x1から生成したNumPy
x2 = x1.data.numpy().copy()
x1[1] = -1
# 結果確認
print(x1)
print(x2)
```
## 1.3 数学上の合成関数とPythonの合成関数
数学上の合成関数がPythonでどう実装されるか確認する
$f(x) = 2x^2 + 2$を関数として定義する
```
def f(x):
return (2 * x**2 + 2)
# xをnumpy配列で定義
x = np.arange(-2, 2.1, 0.25)
print(x)
# f(x)の結果をyに代入
y = f(x)
print(y)
# 関数のグラフ表示
plt.plot(x, y)
plt.show()
# 3つの基本関数の定義
def f1(x):
return(x**2)
def f2(x):
return(x*2)
def f3(x):
return(x+2)
# 合成関数を作る
x1 = f1(x)
x2 = f2(x1)
y = f3(x2)
# 合成関数の値の確認
print(y)
# 合成関数のグラフ表示
plt.plot(x, y)
plt.show()
```
## 1.4 数学上の微分とPythonでの数値微分実装
Pythonでは、関数もまた、変数名は単なるポインタで、実体は別にある。
このことを利用すると、「関数を引数とする関数」を作ることが可能になる。
ここで関数を数値微分する関数``diff``を定義する。
数値微分の計算には、普通の微分の定義式よりいい近似式である $f'(x) = \dfrac{f(x+h)-f(x-h)}{2h}$を利用する。
```
# 関数を微分する関数fdiffの定義
def fdiff(f):
# 関数fを引数に微分した結果の関数をdiffとして定義
def diff(x):
h = 1e-6
return (f(x+h) - f(x-h)) / (2*h)
# fdiffの戻りは微分した結果の関数diff
return diff
```
2次関数fに対して、今作った関数fdiffを適用して、数値微分計算をしてみる。
```
# 2次関数の数値微分
# fの微分結果の関数diffを取得
diff = fdiff(f)
# 微分結果を計算しy_dashに代入
y_dash = diff(x)
# 結果確認
print(y_dash)
# 結果のグラフ表示
plt.plot(x, y, label=r'y = f(x)', c='b')
plt.plot(x, y_dash, label=r"y = f '(x)", c='k')
plt.legend()
plt.show()
```
シグモイド関数 $g(x) = \dfrac{1}{1 + \exp(-x)}$に対して同じことをやってみる。
```
# シグモイド関数の定義
def g(x):
return 1 / (1 + np.exp(-x))
# シグモイド関数の計算
y = g(x)
print(y)
# 関数のグラフ表示
plt.plot(x, y)
plt.show()
# シグモイド関数の数値微分
# gを微分した関数を取得
diff = fdiff(g)
# diffを用いて微分結果y_dashを計算
y_dash = diff(x)
# 結果確認
print(y_dash)
# 結果のグラフ表示
plt.plot(x, y, label=r'y = f(x)', c='b')
plt.plot(x, y_dash, label=r"y = f '(x)", c='k')
plt.legend()
plt.show()
```
シグモイド関数の微分結果は$y(1-y)$となることがわかっている。
これはyの二次関数で、$y=\dfrac{1}{2}$の時に最大値$\dfrac{1}{4}$を取る。
上のグラフはその結果と一致していて、数値微分が正しくできていることがわかる。
## 1.5 オブジェクト指向プログラミング入門
```
# グラフ描画用ライブラリ
import matplotlib.pyplot as plt
# 円描画に必要なライブラリ
import matplotlib.patches as patches
# クラス Point の定義
class Point:
# インスタンス生成時にxとyの2つの引数を持つ
def __init__(self, x, y):
# インスタンスの属性xに第一引数をセットする
self.x = x
# インスタンスの属性yに第二引数をセットする
self.y = y
# 描画関数 drawの定義 (引数はなし)
def draw(self):
# (x, y)に点を描画する
plt.plot(self.x, self.y, marker='o', markersize=10, c='k')
# クラスPointからインスタンス変数p1とp2を生成する
p1 = Point(2,3)
p2 = Point(-1, -2)
# p1とp2の属性x, yの参照
print(p1.x, p1.y)
print(p2.x, p2.y)
# p1とp2のdraw関数を呼び出し、2つの点を描画する
p1.draw()
p2.draw()
plt.xlim(-4, 4)
plt.ylim(-4, 4)
plt.show()
# Pointの子クラスCircleの定義その1
class Circle1(Point):
# Circleはインスタンス生成時に引数x,y,rを持つ
def __init__(self, x, y, r):
# xとyは、親クラスの属性として設定
super().__init__(x, y)
# rは、Circleの属性として設定
self.r = r
# この段階でdraw関数は定義しない
# クラスCircleからインスタンス変数c1_1を生成する
c1_1 = Circle1(1, 0, 2)
# c1_1の属性の確認
print(c1_1.x, c1_1.y, c1_1.r)
# p1, p2, c1_1 のそれぞれのfraw関数を呼び出す
ax = plt.subplot()
p1.draw()
p2.draw()
c1_1.draw()
plt.xlim(-4, 4)
plt.ylim(-4, 4)
plt.show()
```
この段階でdraw関数は親で定義した関数が呼ばれていることがわかる
```
# Pointの子クラスCircleの定義その2
class Circle2(Point):
# Circleはインスタンス生成時に引数x,y,rを持つ
def __init__(self, x, y, r):
# xとyは、親クラスの属性として設定
super().__init__(x, y)
# rは、Circleの属性として設定
self.r = r
# draw関数は、子クラス独自に円の描画を行う
def draw(self):
# 円の描画
c = patches.Circle(xy=(self.x, self.y), radius=self.r, fc='b', ec='k')
ax.add_patch(c)
# クラスCircle2からインスタンス変数c2_1を生成する
c2_1 = Circle2(1, 0, 2)
# p1, p2, c2_1 のそれぞれのfraw関数を呼び出す
ax = plt.subplot()
p1.draw()
p2.draw()
c2_1.draw()
plt.xlim(-4, 4)
plt.ylim(-4, 4)
plt.show()
```
親のdarw関数の代わりに子のdraw関数が呼ばれたことがわかる
では、この関数と親の関数を両方呼びたいときはどうしたらいいか
```
# Pointの子クラスCircleの定義その3
class Circle3(Point):
# Circleはインスタンス生成時に引数x,y,rを持つ
def __init__(self, x, y, r):
# xとyは、親クラスの属性として設定
super().__init__(x, y)
# rは、Circleの属性として設定
self.r = r
# Circleのdraw関数は、親の関数呼び出しの後で、円の描画も独自に行う
def draw(self):
# 親クラスのdraw関数呼び出し
super().draw()
# 円の描画
c = patches.Circle(xy=(self.x, self.y), radius=self.r, fc='b', ec='k')
ax.add_patch(c)
# クラスCircle3からインスタンス変数c3_1を生成する
c3_1 = Circle3(1, 0, 2)
# p1, p2, c3_1 のそれぞれのfraw関数を呼び出す
ax = plt.subplot()
p1.draw()
p2.draw()
c3_1.draw()
plt.xlim(-4, 4)
plt.ylim(-4, 4)
plt.show()
```
無事、両方を呼び出すことができた
## 1.6 インスタンスを関数として呼び出し可能にする
```
# 関数クラスHの定義
class H:
def __call__(self, x):
return 2*x**2 + 2
# hが関数として動作することを確認する
# numpy配列としてxの定義
x = np.arange(-2, 2.1, 0.25)
print(x)
# Hクラスのインスタンスとしてhを生成
h = H()
# 関数hの呼び出し
y = h(x)
print(y)
# グラフ描画
plt.plot(x, y)
plt.show()
```
| github_jupyter |
# Gaussian Process Distribution of Relaxation Times.
## In this tutorial we will show use the GP-DRT method to analyze actual experimental data
The impedance data in the csv file named `EIS_experiment.csv`. The file has three columns. The first column is the frequency, the second one the real part of the impedance. The third column is the imaginary part of impedance. To use this tutorial for your own data, we recommend the frequencies go are sorted ascendingly.
```
from math import cos, pi, sin
import GP_DRT
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from scipy.optimize import minimize
%matplotlib inline
```
## 1) Read in the impedance data from the csv file
### IMPORTANT: the frequency value should be sorted ascendingly (from low to high)
```
Z_data = pd.read_csv("EIS_experiment.csv")
freq_vec, Z_exp = (
Z_data["freq"].values,
Z_data["Z_real"].values + 1j * Z_data["Z_imag"].values,
)
# define the frequency range
N_freqs = len(freq_vec)
xi_vec = np.log(freq_vec)
tau = 1 / freq_vec
# define the frequency range used for prediction, we choose a wider range to better display the DRT
freq_vec_star = np.logspace(-4.0, 6.0, num=101, endpoint=True)
xi_vec_star = np.log(freq_vec_star)
# finer mesh for plotting only
freq_vec_plot = np.logspace(-4.0, 6.0, num=1001, endpoint=True)
```
## 2) Show the impedance spectrum as a Nyquist plot
```
# Nyquist plot of the EIS spectrum
plt.plot(
np.real(Z_exp),
-np.imag(Z_exp),
"o",
markersize=10,
fillstyle="none",
color="red",
label="experiment",
)
plt.plot(
np.real(Z_exp[40:80:10]),
-np.imag(Z_exp[40:80:10]),
"o",
markersize=10,
color="black",
)
plt.rc("text", usetex=False)
plt.rc("font", family="serif", size=15)
plt.rc("xtick", labelsize=15)
plt.rc("ytick", labelsize=15)
plt.legend(frameon=False, fontsize=15)
plt.axis("scaled")
# this depends on the data used - if you wish to use your own data you may need to modify this
plt.xlim(1.42, 1.52)
plt.ylim(-0.001, 0.051)
plt.xticks(np.arange(1.42, 1.521, 0.02))
plt.yticks(np.arange(0.00, 0.051, 0.01))
plt.gca().set_aspect("equal", adjustable="box")
plt.xlabel(r"$Z_{\rm re}/\Omega$", fontsize=20)
plt.ylabel(r"$-Z_{\rm im}/\Omega$", fontsize=20)
# label the frequencies - if you wish to use your own data you may need to modify this
label_index = range(40, 80, 10)
move = [[-0.005, 0.008], [-0.005, 0.008], [-0.005, 0.008], [-0.005, 0.01]]
for k, ind in enumerate(label_index):
power = int(np.log10(freq_vec[ind]))
num = freq_vec[ind] / (10 ** (power))
plt.annotate(
r"${0:.1f}\times 10^{1}$".format(num, power),
xy=(np.real(Z_exp[ind]), -np.imag(Z_exp[ind])),
xytext=(np.real(Z_exp[ind]) + move[k][0], move[k][1] - np.imag(Z_exp[ind])),
arrowprops=dict(arrowstyle="-", connectionstyle="arc"),
)
plt.show()
```
## 3) Compute the optimal hyperparameters
### Note: the intial parameters may need to be adjusted according to the specific problem
```
# initial parameters parameter to maximize the marginal log-likelihood as shown in eq (31)
sigma_n = 1.0e-4
sigma_f = 1.0e-3
ell = 1.0
theta_0 = np.array([sigma_n, sigma_f, ell])
seq_theta = np.copy(theta_0)
def print_results(theta):
global seq_theta
seq_theta = np.vstack((seq_theta, theta))
print("{0:.7f} {1:.7f} {2:.7f}".format(theta[0], theta[1], theta[2]))
GP_DRT.NMLL_fct(theta_0, Z_exp, xi_vec)
GP_DRT.grad_NMLL_fct(theta_0, Z_exp, xi_vec)
print("sigma_n, sigma_f, ell")
# minimize the NMLL $L(\theta)$ w.r.t sigma_n, sigma_f, ell using the BFGS method as implemented in scipy
res = minimize(
GP_DRT.NMLL_fct,
theta_0,
args=(Z_exp, xi_vec),
method="BFGS",
jac=GP_DRT.grad_NMLL_fct,
callback=print_results,
options={"disp": True},
)
# collect the optimized parameters
sigma_n, sigma_f, ell = res.x
```
## 4) Core of the GP-DRT
### 4a) Compute matrices
```
# calculate the matrices shown in eq (18)
K = GP_DRT.matrix_K(xi_vec, xi_vec, sigma_f, ell)
L_im_K = GP_DRT.matrix_L_im_K(xi_vec, xi_vec, sigma_f, ell)
L2_im_K = GP_DRT.matrix_L2_im_K(xi_vec, xi_vec, sigma_f, ell)
Sigma = (sigma_n ** 2) * np.eye(N_freqs)
```
### 4b) Factorize the matrices and solve the linear equations
```
# the matrix $\mathcal L^2_{\rm im} \mathbf K + \sigma_n^2 \mathbf I$ whose inverse is needed
K_im_full = L2_im_K + Sigma
# Cholesky factorization, L is a lower-triangular matrix
L = np.linalg.cholesky(K_im_full)
# solve for alpha
alpha = np.linalg.solve(L, Z_exp.imag)
alpha = np.linalg.solve(L.T, alpha)
# estimate the gamma of eq (21a), the minus sign, which is not included in L_im_K, refers to eq (65)
gamma_fct_est = -np.dot(L_im_K.T, alpha)
# covariance matrix
inv_L = np.linalg.inv(L)
inv_K_im_full = np.dot(inv_L.T, inv_L)
# estimate the sigma of gamma for eq (21b)
cov_gamma_fct_est = K - np.dot(L_im_K.T, np.dot(inv_K_im_full, L_im_K))
sigma_gamma_fct_est = np.sqrt(np.diag(cov_gamma_fct_est))
```
### 4c) Predict the imaginary part of the GP-DRT and impedance
```
# initialize the imaginary part of impedance vector
Z_im_vec_star = np.empty_like(xi_vec_star)
Sigma_Z_im_vec_star = np.empty_like(xi_vec_star)
gamma_vec_star = np.empty_like(xi_vec_star)
Sigma_gamma_vec_star = np.empty_like(xi_vec_star)
# calculate the imaginary part of impedance at each $\xi$ point for the plot
for index, val in enumerate(xi_vec_star):
xi_star = np.array([val])
# compute matrices shown in eq (18), k_star corresponds to a new point
k_star = GP_DRT.matrix_K(xi_vec, xi_star, sigma_f, ell)
L_im_k_star = GP_DRT.matrix_L_im_K(xi_vec, xi_star, sigma_f, ell)
L2_im_k_star = GP_DRT.matrix_L2_im_K(xi_vec, xi_star, sigma_f, ell)
k_star_star = GP_DRT.matrix_K(xi_star, xi_star, sigma_f, ell)
L_im_k_star_star = GP_DRT.matrix_L_im_K(xi_star, xi_star, sigma_f, ell)
L2_im_k_star_star = GP_DRT.matrix_L2_im_K(xi_star, xi_star, sigma_f, ell)
# compute Z_im_star mean and standard deviation using eq (26)
Z_im_vec_star[index] = np.dot(L2_im_k_star.T, np.dot(inv_K_im_full, Z_exp.imag))
Sigma_Z_im_vec_star[index] = L2_im_k_star_star - np.dot(
L2_im_k_star.T, np.dot(inv_K_im_full, L2_im_k_star)
)
# compute Z_im_star mean and standard deviation
gamma_vec_star[index] = -np.dot(L_im_k_star.T, np.dot(inv_K_im_full, Z_exp.imag))
Sigma_gamma_vec_star[index] = k_star_star - np.dot(
L_im_k_star.T, np.dot(inv_K_im_full, L_im_k_star)
)
```
### 4d) Plot the obtained GP-DRT
```
# plot the DRT and its confidence region
plt.semilogx(freq_vec_star, gamma_vec_star, linewidth=4, color="red", label="GP-DRT")
plt.fill_between(
freq_vec_star,
gamma_vec_star - 3 * np.sqrt(abs(Sigma_gamma_vec_star)),
gamma_vec_star + 3 * np.sqrt(abs(Sigma_gamma_vec_star)),
color="0.4",
alpha=0.3,
)
plt.rc("text", usetex=False)
plt.rc("font", family="serif", size=15)
plt.rc("xtick", labelsize=15)
plt.rc("ytick", labelsize=15)
plt.axis([1e-4, 1e6, -0.01, 0.025])
plt.yticks(np.arange(-0.01, 0.025, 0.01))
plt.legend(frameon=False, fontsize=15)
plt.xlabel(r"$f/{\rm Hz}$", fontsize=20)
plt.ylabel(r"$\gamma/\Omega$", fontsize=20)
plt.show()
```
### 4e) Plot the imaginary part of the GP-DRT impedance together with the experimental one
```
plt.semilogx(
freq_vec, -Z_exp.imag, "o", markersize=10, color="black", label="synth exp"
)
plt.semilogx(freq_vec_star, -Z_im_vec_star, linewidth=4, color="red", label="GP-DRT")
plt.fill_between(
freq_vec_star,
-Z_im_vec_star - 3 * np.sqrt(abs(Sigma_Z_im_vec_star)),
-Z_im_vec_star + 3 * np.sqrt(abs(Sigma_Z_im_vec_star)),
alpha=0.3,
)
plt.rc("text", usetex=False)
plt.rc("font", family="serif", size=15)
plt.rc("xtick", labelsize=15)
plt.rc("ytick", labelsize=15)
plt.axis([1e-3, 1e5, -0.01, 0.03])
plt.legend(frameon=False, fontsize=15)
plt.xlabel(r"$f/{\rm Hz}$", fontsize=20)
plt.ylabel(r"$-Z_{\rm im}/\Omega$", fontsize=20)
plt.show()
```
| github_jupyter |
# Logistic Regression
In this lesson, we're going to implement logistic regression for a classification task where we want to probabilistically determine the outcome for a given set of inputs. We will understand the basic math behind it, implement it in just NumPy and then in PyTorch.
<div align="left">
<a href="https://github.com/madewithml/lessons/blob/master/notebooks/02_Basics/02_Logistic_Regression/02_PT_Logistic_Regression.ipynb" role="button"><img class="notebook-badge-image" src="https://img.shields.io/static/v1?label=&message=View%20On%20GitHub&color=586069&logo=github&labelColor=2f363d"></a>
<a href="https://colab.research.google.com/github/madewithml/lessons/blob/master/notebooks/02_Basics/02_Logistic_Regression/02_PT_Logistic_Regression.ipynb"><img class="notebook-badge-image" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"></a>
</div>
# Overview
Logistic regression is just an extension on linear regression (both are generalized linear methods). We will still learn to model a line (plane) that models $y$ given $X$. Except now we are dealing with classification problems as opposed to regression problems so we want classification probabilities. We'll be using the softmax operation to normalize our logits ($XW$) to derive probabilities.
Our goal is to learn a logistic model $\hat{y}$ that models $y$ given $X$.
$ \hat{y} = \frac{e^{XW_y}}{\sum_j e^{XW}} $
* $\hat{y}$ = prediction | $\in \mathbb{R}^{NX1}$ ($N$ is the number of samples)
* $X$ = inputs | $\in \mathbb{R}^{NXD}$ ($D$ is the number of features)
* $W$ = weights | $\in \mathbb{R}^{DXC}$ ($C$ is the number of classes)
This function is known as the multinomial logistic regression or the softmax classifier. The softmax classifier will use the linear equation ($z=XW$) and normalize it (using the softmax function) to produce the probabiltiy for class y given the inputs.
**NOTE**: We'll leave the bias weights out for now to avoid complicating the backpropagation calculation.
* **Objective:** Predict the probability of class $y$ given the inputs $X$. The softmax classifier normalizes the linear outputs to determine class probabilities.
* **Advantages:**
* Can predict class probabilities given a set on inputs.
* **Disadvantages:**
* Sensitive to outliers since objective is to minimize cross entropy loss. Support vector machines ([SVMs](https://towardsdatascience.com/support-vector-machine-vs-logistic-regression-94cc2975433f)) are a good alternative to counter outliers.
* **Miscellaneous:** Softmax classifier is going to used widely in neural network architectures as the last layer since it produces class probabilities.
# Data
## Load data
We'll used some synthesized data to train our models on. The task is to determine whether a tumor will be benign (harmless) or malignant (harmful) based on leukocyte (white blood cells) count and blood pressure. Note that this is a synethic dataset that has no clinical relevance.
```
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from pandas.plotting import scatter_matrix
import urllib
SEED = 1234
DATA_FILE = 'tumors.csv'
# Set seed for reproducibility
np.random.seed(SEED)
# Download data from GitHub to this notebook's local drive
url = "https://raw.githubusercontent.com/madewithml/lessons/master/data/tumors.csv"
response = urllib.request.urlopen(url)
html = response.read()
with open(DATA_FILE, 'wb') as fp:
fp.write(html)
# Read from CSV to Pandas DataFrame
df = pd.read_csv(DATA_FILE, header=0)
df.head()
# Define X and y
X = df[['leukocyte_count', 'blood_pressure']].values
y = df['tumor_class'].values
# Plot data
colors = {'benign': 'red', 'malignant': 'blue'}
plt.scatter(X[:, 0], X[:, 1], c=[colors[_y] for _y in y], s=25, edgecolors='k')
plt.xlabel('leukocyte count')
plt.ylabel('blood pressure')
plt.legend(['malignant ', 'benign'], loc="upper right")
plt.show()
```
## Split data
```
import collections
from sklearn.model_selection import train_test_split
TRAIN_SIZE = 0.7
VAL_SIZE = 0.15
TEST_SIZE = 0.15
SHUFFLE = True
```
Splitting the dataset for classification tasks requires more than just randomly splitting into train, validation and test sets. We want to ensure that each split has a similar class distribution so that we can learn to predict well across all the classes. We can do this by specifying the `stratify` argument in [`train_test_split`](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html) which will be set to the class labels.
```
def train_val_test_split(X, y, val_size, test_size, shuffle):
"""Split data into train/val/test datasets.
"""
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=test_size, stratify=y, shuffle=shuffle) # notice the `stratify=y`
X_train, X_val, y_train, y_val = train_test_split(
X_train, y_train, test_size=val_size, stratify=y_train, shuffle=shuffle) # notice the `stratify=y_train`
return X_train, X_val, X_test, y_train, y_val, y_test
# Create data splits
X_train, X_val, X_test, y_train, y_val, y_test = train_val_test_split(
X=X, y=y, val_size=VAL_SIZE, test_size=TEST_SIZE, shuffle=SHUFFLE)
class_counts = dict(collections.Counter(y))
print (f"X_train: {X_train.shape}, y_train: {y_train.shape}")
print (f"X_val: {X_val.shape}, y_val: {y_val.shape}")
print (f"X_test: {X_test.shape}, y_test: {y_test.shape}")
print (f"Sample point: {X_train[0]} → {y_train[0]}")
print (f"Classes: {class_counts}")
```
## Label encoder
You'll notice that our class labels are text. We need to encode them into integers so we can use them in our models. We're going to the scikit-learn's [LabelEncoder](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.LabelEncoder.html#sklearn.preprocessing.LabelEncoder) to do this.
```
from sklearn.preprocessing import LabelEncoder
# Output vectorizer
y_tokenizer = LabelEncoder()
# Fit on train data
y_tokenizer = y_tokenizer.fit(y_train)
classes = y_tokenizer.classes_
print (f"classes: {classes}")
# Convert labels to tokens
print (f"y_train[0]: {y_train[0]}")
y_train = y_tokenizer.transform(y_train)
y_val = y_tokenizer.transform(y_val)
y_test = y_tokenizer.transform(y_test)
print (f"y_train[0]: {y_train[0]}")
# Class weights
counts = collections.Counter(y_train)
class_weights = {_class: 1.0/count for _class, count in counts.items()}
print (f"class counts: {counts},\nclass weights: {class_weights}")
```
**NOTE**: Class weights are useful for weighting the loss function during training. It tells the model to focus on samples from an under-represented class. The loss section below will show how to incorporate these weights.
## Standardize data
We need to standardize our data (zero mean and unit variance) in order to optimize quickly. We're only going to standardize the inputs X because our outputs y are class values.
```
from sklearn.preprocessing import StandardScaler
# Standardize the data (mean=0, std=1) using training data
X_scaler = StandardScaler().fit(X_train)
# Apply scaler on training and test data (don't standardize outputs for classification)
X_train = X_scaler.transform(X_train)
X_val = X_scaler.transform(X_val)
X_test = X_scaler.transform(X_test)
# Check (means should be ~0 and std should be ~1)
print (f"X_train[0]: mean: {np.mean(X_train[:, 0], axis=0):.1f}, std: {np.std(X_train[:, 0], axis=0):.1f}")
print (f"X_train[1]: mean: {np.mean(X_train[:, 1], axis=0):.1f}, std: {np.std(X_train[:, 1], axis=0):.1f}")
print (f"X_val[0]: mean: {np.mean(X_val[:, 0], axis=0):.1f}, std: {np.std(X_val[:, 0], axis=0):.1f}")
print (f"X_val[1]: mean: {np.mean(X_val[:, 1], axis=0):.1f}, std: {np.std(X_val[:, 1], axis=0):.1f}")
print (f"X_test[0]: mean: {np.mean(X_test[:, 0], axis=0):.1f}, std: {np.std(X_test[:, 0], axis=0):.1f}")
print (f"X_test[1]: mean: {np.mean(X_test[:, 1], axis=0):.1f}, std: {np.std(X_test[:, 1], axis=0):.1f}")
```
# NumPy
Now that we have our data prepared, we'll first implement logistic regression using just NumPy. This will let us really understand the underlying operations. It's normal to find the math and code in this section slightly complex. You can still read each of the steps to build intuition for when we implement this using PyTorch.
Our goal is to learn a logistic model $\hat{y}$ that models $y$ given $X$.
$ \hat{y} = \frac{e^{XW_y}}{\sum e^{XW}} $
* $\hat{y}$ = prediction | $\in \mathbb{R}^{NX1}$ ($N$ is the number of samples)
* $X$ = inputs | $\in \mathbb{R}^{NXD}$ ($D$ is the number of features)
* $W$ = weights | $\in \mathbb{R}^{DXC}$ ($C$ is the number of classes)
We are going to use multinomial logistic regression even though our task only involves two classes because you can generalize the softmax classifier to any number of classes.
## Initialize weights
1. Randomly initialize the model's weights $W$.
```
INPUT_DIM = X_train.shape[1] # X is 2-dimensional
NUM_CLASSES = len(classes) # y has two possibilities (begign or malignant)
# Initialize random weights
W = 0.01 * np.random.randn(INPUT_DIM, NUM_CLASSES)
b = np.zeros((1, NUM_CLASSES))
print (f"W: {W.shape}")
print (f"b: {b.shape}")
```
## Model
2. Feed inputs $X$ into the model to receive the logits ($z=XW$). Apply the softmax operation on the logits to get the class probabilies $\hat{y}$ in one-hot encoded form. For example, if there are three classes, the predicted class probabilities could look like [0.3, 0.3, 0.4].
* $ \hat{y} = softmax(z) = softmax(XW) = \frac{e^{XW_y}}{\sum_j e^{XW}} $
```
# Forward pass [NX2] · [2X2] + [1,2] = [NX2]
logits = np.dot(X_train, W) + b
print (f"logits: {logits.shape}")
print (f"sample: {logits[0]}")
# Normalization via softmax to obtain class probabilities
exp_logits = np.exp(logits)
y_hat = exp_logits / np.sum(exp_logits, axis=1, keepdims=True)
print (f"y_hat: {y_hat.shape}")
print (f"sample: {y_hat[0]}")
```
## Loss
3. Compare the predictions $\hat{y}$ (ex. [0.3, 0.3, 0.4]]) with the actual target values $y$ (ex. class 2 would look like [0, 0, 1]) with the objective (cost) function to determine loss $J$. A common objective function for logistics regression is cross-entropy loss.
* $J(\theta) = - \sum_i ln(\hat{y_i}) = - \sum_i ln (\frac{e^{X_iW_y}}{\sum_j e^{X_iW}}) $
```
# Loss
correct_class_logprobs = -np.log(y_hat[range(len(y_hat)), y_train])
loss = np.sum(correct_class_logprobs) / len(y_train)
print (f"loss: {loss:.2f}")
```
## Gradients
4. Calculate the gradient of loss $J(\theta)$ w.r.t to the model weights. Let's assume that our classes are mutually exclusive (a set of inputs could only belong to one class).
* $\frac{\partial{J}}{\partial{W_j}} = \frac{\partial{J}}{\partial{\hat{y}}}\frac{\partial{\hat{y}}}{\partial{W_j}} = - \frac{1}{\hat{y}}\frac{\partial{\hat{y}}}{\partial{W_j}} = - \frac{1}{\frac{e^{XW_y}}{\sum_j e^{XW}}}\frac{\sum_j e^{XW}e^{XW_y}0 - e^{XW_y}e^{XW_j}X}{(\sum_j e^{XW})^2} = \frac{Xe^{XW_j}}{\sum_j e^{XW}} = X\hat{y}$
* $\frac{\partial{J}}{\partial{W_y}} = \frac{\partial{J}}{\partial{\hat{y}}}\frac{\partial{\hat{y}}}{\partial{W_y}} = - \frac{1}{\hat{y}}\frac{\partial{\hat{y}}}{\partial{W_y}} = - \frac{1}{\frac{e^{XW_y}}{\sum_j e^{XW}}}\frac{\sum_j e^{XW}e^{XW_y}X - e^{W_yX}e^{XW_y}X}{(\sum_j e^{XW})^2} = \frac{1}{\hat{y}}(X\hat{y} - X\hat{y}^2) = X(\hat{y}-1)$
```
# Backpropagation
dscores = y_hat
dscores[range(len(y_hat)), y_train] -= 1
dscores /= len(y_train)
dW = np.dot(X_train.T, dscores)
db = np.sum(dscores, axis=0, keepdims=True)
```
## Update weights
5. Update the weights 𝑊 using a small learning rate 𝛼. The updates willpenalize the probabiltiy for the incorrect classes (j) and encourage a higher probability for the correct class (y).
* $W_j = W_j - \alpha\frac{\partial{J}}{\partial{W_j}}$
```
LEARNING_RATE = 1e-1
# Update weights
W += -LEARNING_RATE * dW
b += -LEARNING_RATE * db
```
## Training
```
NUM_EPOCHS = 50
```
6. Repeat steps 2 - 5 to minimize the loss and train the model.
```
# Initialize random weights
W = 0.01 * np.random.randn(INPUT_DIM, NUM_CLASSES)
b = np.zeros((1, NUM_CLASSES))
# Training loop
for epoch_num in range(NUM_EPOCHS):
# Forward pass [NX2] · [2X2] = [NX2]
logits = np.dot(X_train, W) + b
# Normalization via softmax to obtain class probabilities
exp_logits = np.exp(logits)
y_hat = exp_logits / np.sum(exp_logits, axis=1, keepdims=True)
# Loss
correct_class_logprobs = -np.log(y_hat[range(len(y_hat)), y_train])
loss = np.sum(correct_class_logprobs) / len(y_train)
# show progress
if epoch_num%10 == 0:
# Accuracy
y_pred = np.argmax(logits, axis=1)
accuracy = np.mean(np.equal(y_train, y_pred))
print (f"Epoch: {epoch_num}, loss: {loss:.3f}, accuracy: {accuracy:.3f}")
# Backpropagation
dscores = y_hat
dscores[range(len(y_hat)), y_train] -= 1
dscores /= len(y_train)
dW = np.dot(X_train.T, dscores)
db = np.sum(dscores, axis=0, keepdims=True)
# Update weights
W += -LEARNING_RATE * dW
b += -LEARNING_RATE * db
```
Since we're taking the argmax, we can just calculate logits since normalization won't change which index has the higher value.
```
class LogisticRegressionFromScratch():
def predict(self, x):
logits = np.dot(x, W) + b
exp_logits = np.exp(logits)
y_hat = exp_logits / np.sum(exp_logits, axis=1, keepdims=True)
return y_hat
# Evaluation
model = LogisticRegressionFromScratch()
logits_train = model.predict(X_train)
pred_train = np.argmax(logits_train, axis=1)
logits_test = model.predict(X_test)
pred_test = np.argmax(logits_test, axis=1)
# Training and test accuracy
train_acc = np.mean(np.equal(y_train, pred_train))
test_acc = np.mean(np.equal(y_test, pred_test))
print (f"train acc: {train_acc:.2f}, test acc: {test_acc:.2f}")
def plot_multiclass_decision_boundary(model, X, y, savefig_fp=None):
"""Plot the multiclass decision boundary for a model that accepts 2D inputs.
Arguments:
model {function} -- trained model with function model.predict(x_in).
X {numpy.ndarray} -- 2D inputs with shape (N, 2).
y {numpy.ndarray} -- 1D outputs with shape (N,).
"""
# Axis boundaries
x_min, x_max = X[:, 0].min() - 0.1, X[:, 0].max() + 0.1
y_min, y_max = X[:, 1].min() - 0.1, X[:, 1].max() + 0.1
xx, yy = np.meshgrid(np.linspace(x_min, x_max, 101),
np.linspace(y_min, y_max, 101))
# Create predictions
x_in = np.c_[xx.ravel(), yy.ravel()]
y_pred = model.predict(x_in)
y_pred = np.argmax(y_pred, axis=1).reshape(xx.shape)
# Plot decision boundary
plt.contourf(xx, yy, y_pred, cmap=plt.cm.Spectral, alpha=0.8)
plt.scatter(X[:, 0], X[:, 1], c=y, s=40, cmap=plt.cm.RdYlBu)
plt.xlim(xx.min(), xx.max())
plt.ylim(yy.min(), yy.max())
# Plot
if savefig_fp:
plt.savefig(savefig_fp, format='png')
# Visualize the decision boundary
plt.figure(figsize=(12,5))
plt.subplot(1, 2, 1)
plt.title("Train")
plot_multiclass_decision_boundary(model=model, X=X_train, y=y_train)
plt.subplot(1, 2, 2)
plt.title("Test")
plot_multiclass_decision_boundary(model=model, X=X_test, y=y_test)
plt.show()
```
Credit for the plotting functions and the intuition behind all this is due to [CS231n](http://cs231n.github.io/neural-networks-case-study/), one of the best courses for machine learning. Now let's implement logistic regression with PyTorch.
# PyTorch
Now that we've implemented logistic regression with Numpy, let's do the same with PyTorch.
```
import torch
# Set seed for reproducibility
torch.manual_seed(SEED)
```
## Model
We will be using Linear layers to recreate the same model.
```
from torch import nn
import torch.nn.functional as F
from torchsummary import summary
class LogisticRegression(nn.Module):
def __init__(self, input_dim, num_classes):
super(LogisticRegression, self).__init__()
self.fc1 = nn.Linear(input_dim, num_classes)
def forward(self, x_in, apply_softmax=False):
y_pred = self.fc1(x_in)
if apply_softmax:
y_pred = F.softmax(y_pred, dim=1)
return y_pred
# Initialize model
model = LogisticRegression(input_dim=INPUT_DIM, num_classes=NUM_CLASSES)
print (model.named_parameters)
summary(model, input_size=(INPUT_DIM,))
```
## Loss
Our loss will be the categorical crossentropy.
```
loss_fn = nn.CrossEntropyLoss()
y_pred = torch.randn(3, NUM_CLASSES, requires_grad=False)
y_true = torch.empty(3, dtype=torch.long).random_(NUM_CLASSES)
print (y_true)
loss = loss_fn(y_pred, y_true)
print(f'Loss: {loss.numpy()}')
```
In our case, we will also incorporate the class weights into our loss function to counter any class imbalances.
```
# Loss
weights = torch.Tensor([class_weights[key] for key in sorted(class_weights.keys())])
loss_fn = nn.CrossEntropyLoss(weight=weights)
```
## Metrics
```
# Accuracy
def accuracy_fn(y_pred, y_true):
n_correct = torch.eq(y_pred, y_true).sum().item()
accuracy = (n_correct / len(y_pred)) * 100
return accuracy
y_pred = torch.Tensor([0, 0, 1])
y_true = torch.Tensor([1, 1, 1])
print(f'Accuracy: {accuracy_fn(y_pred, y_true):.1f}')
```
## Optimizer
```
from torch.optim import Adam
# Optimizer
optimizer = Adam(model.parameters(), lr=LEARNING_RATE)
```
## Training
```
# Convert data to tensors
X_train = torch.Tensor(X_train)
y_train = torch.LongTensor(y_train)
X_val = torch.Tensor(X_val)
y_val = torch.LongTensor(y_val)
X_test = torch.Tensor(X_test)
y_test = torch.LongTensor(y_test)
# Training
for epoch in range(NUM_EPOCHS):
# Forward pass
y_pred = model(X_train)
# Loss
loss = loss_fn(y_pred, y_train)
# Zero all gradients
optimizer.zero_grad()
# Backward pass
loss.backward()
# Update weights
optimizer.step()
if epoch%10==0:
predictions = y_pred.max(dim=1)[1] # class
accuracy = accuracy_fn(y_pred=predictions, y_true=y_train)
print (f"Epoch: {epoch} | loss: {loss:.2f}, accuracy: {accuracy:.1f}")
```
**NOTE**: We used the `class_weights` from earlier which will be used with our loss function to account for any class imbalances (our dataset doesn't suffer from this issue).
## Evaluation
```
import itertools
import matplotlib.pyplot as plt
from sklearn.metrics import accuracy_score
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
def plot_multiclass_decision_boundary(model, X, y):
x_min, x_max = X[:, 0].min() - 0.1, X[:, 0].max() + 0.1
y_min, y_max = X[:, 1].min() - 0.1, X[:, 1].max() + 0.1
xx, yy = np.meshgrid(np.linspace(x_min, x_max, 101), np.linspace(y_min, y_max, 101))
cmap = plt.cm.Spectral
X_test = torch.from_numpy(np.c_[xx.ravel(), yy.ravel()]).float()
y_pred = model(X_test, apply_softmax=True)
_, y_pred = y_pred.max(dim=1)
y_pred = y_pred.reshape(xx.shape)
plt.contourf(xx, yy, y_pred, cmap=plt.cm.Spectral, alpha=0.8)
plt.scatter(X[:, 0], X[:, 1], c=y, s=40, cmap=plt.cm.RdYlBu)
plt.xlim(xx.min(), xx.max())
plt.ylim(yy.min(), yy.max())
def plot_confusion_matrix(y_true, y_pred, classes, cmap=plt.cm.Blues):
"""Plot a confusion matrix using ground truth and predictions."""
# Confusion matrix
cm = confusion_matrix(y_true, y_pred)
cm_norm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
# Figure
fig = plt.figure()
ax = fig.add_subplot(111)
cax = ax.matshow(cm, cmap=plt.cm.Blues)
fig.colorbar(cax)
# Axis
plt.title("Confusion matrix")
plt.ylabel("True label")
plt.xlabel("Predicted label")
ax.set_xticklabels([''] + classes)
ax.set_yticklabels([''] + classes)
ax.xaxis.set_label_position('bottom')
ax.xaxis.tick_bottom()
# Values
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, f"{cm[i, j]:d} ({cm_norm[i, j]*100:.1f}%)",
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
# Display
plt.show()
# Predictions
pred_train = model(X_train, apply_softmax=True)
pred_test = model(X_test, apply_softmax=True)
print (f"sample probability: {pred_test[0]}")
pred_train = pred_train.max(dim=1)[1]
pred_test = pred_test.max(dim=1)[1]
print (f"sample class: {pred_test[0]}")
# Accuracy
train_acc = accuracy_score(y_train, pred_train)
test_acc = accuracy_score(y_test, pred_test)
print (f"train acc: {train_acc:.2f}, test acc: {test_acc:.2f}")
# Visualize the decision boundary
plt.figure(figsize=(12,5))
plt.subplot(1, 2, 1)
plt.title("Train")
plot_multiclass_decision_boundary(model=model, X=X_train, y=y_train)
plt.subplot(1, 2, 2)
plt.title("Test")
plot_multiclass_decision_boundary(model=model, X=X_test, y=y_test)
plt.show()
```
So far we looked at accuracy as the metric that determines our mode's level of performance. But we have several other options when it comes to evaluation metrics.
<div align="left">
<img src="https://raw.githubusercontent.com/madewithml/images/master/02_Basics/02_Logistic_Regression/metrics.png" width="350">
</div>
<small>Image credit: "Precisionrecall" by Walber</small>
The metric we choose really depends on the situation.
positive - true, 1, tumor, issue, etc., negative - false, 0, not tumor, not issue, etc.
$\text{accuracy} = \frac{TP+TN}{TP+TN+FP+FN}$
$\text{recall} = \frac{TP}{TP+FN}$ → (how many of the actual issues did I catch)
$\text{precision} = \frac{TP}{TP+FP}$ → (out of all the things I said were issues, how many were actually issues)
$F_1 = 2 * \frac{\text{precision } * \text{ recall}}{\text{precision } + \text{ recall}}$
where:
* TP: # of samples predicted to be positive and were actually positive
* TN: # of samples predicted to be negative and were actually negative
* FP: # of samples predicted to be positive but were actually negative
* FN: # of samples predicted to be negative but were actually positive
```
# Classification report
plot_confusion_matrix(y_true=y_test, y_pred=pred_test, classes=classes)
print (classification_report(y_test, pred_test))
```
## Inference
```
# Inputs for inference
X_infer = pd.DataFrame([{'leukocyte_count': 13, 'blood_pressure': 12}])
X_infer.head()
# Standardize
X_infer = X_scaler.transform(X_infer)
print (X_infer)
# Predict
y_infer = model(torch.Tensor(X_infer), apply_softmax=True)
prob, _class = y_infer.max(dim=1)
print (f"The probability that you have a {classes[_class.detach().numpy()[0]]} tumor is {prob.detach().numpy()[0]*100.0:.0f}%")
```
# Unscaled weights
Note that only X was standardized.
$\hat{y}_{unscaled} = b_{scaled} + \sum_{j=1}^{k}W_{{scaled}_j}x_{{scaled}_j}$
* $x_{scaled} = \frac{x_j - \bar{x}_j}{\sigma_j}$
$\hat{y}_{unscaled} = b_{scaled} + \sum_{j=1}^{k} W_{{scaled}_j} (\frac{x_j - \bar{x}_j}{\sigma_j}) $
$\hat{y}_{unscaled} = (b_{scaled} - \sum_{j=1}^{k} W_{{scaled}_j}\frac{\bar{x}_j}{\sigma_j}) + \sum_{j=1}^{k} (\frac{W_{{scaled}_j}}{\sigma_j})x_j$
In the expression above, we can see the expression $\hat{y}_{unscaled} = W_{unscaled}x + b_{unscaled} $
* $W_{unscaled} = \sum_{j=1}^{k} (\frac{W_{{scaled}_j}}{\sigma_j}) $
* $b_{unscaled} = b_{scaled} - \sum_{j=1}^{k} W_{{scaled}_j}\frac{\bar{x}_j}{\sigma_j}$
```
# Unstandardize weights
W = model.fc1.weight.data.numpy()
b = model.fc1.bias.data.numpy()
W_unscaled = W / X_scaler.scale_
b_unscaled = b - np.sum((W_unscaled * X_scaler.mean_))
print (W_unscaled)
print (b_unscaled)
```
---
Share and discover ML projects at <a href="https://madewithml.com/">Made With ML</a>.
<div align="left">
<a class="ai-header-badge" target="_blank" href="https://github.com/madewithml/lessons"><img src="https://img.shields.io/github/stars/madewithml/lessons.svg?style=social&label=Star"></a>
<a class="ai-header-badge" target="_blank" href="https://www.linkedin.com/company/madewithml"><img src="https://img.shields.io/badge/style--5eba00.svg?label=LinkedIn&logo=linkedin&style=social"></a>
<a class="ai-header-badge" target="_blank" href="https://twitter.com/madewithml"><img src="https://img.shields.io/twitter/follow/madewithml.svg?label=Follow&style=social"></a>
</div>
| github_jupyter |
```
# default_exp torch
```
# Pytorch Errors
> All the possible errors that fastdebug can support and verbosify involving Pytorch
```
#hide
from nbdev.showdoc import *
from fastcore.test import test_eq
#export
import torch
import re
from fastai.callback.hook import Hook
from fastai.torch_core import to_detach
from fastai.layers import flatten_model
from fastcore.basics import store_attr
```
## Errors
While some errrors are specifically designed for the [fastai](https://docs.fast.ai) library, the general idea still holds true in raw `Pytorch` as well.
```
#export
def device_error(e:Exception, a:str, b:str) -> Exception:
"""
Verbose error for if `a` and `b` are on different devices
Should be used when checking if a model is on the same device, or two tensors
"""
inp, weight, _ = e.args[0].replace('( ', '').split(')')
inp = inp.replace('Input type', f'{a} has type: \t\t')
weight = weight.replace(' and weight type', f'{b} have type: \t')
err = f'Mismatch between weight types\n\n{inp})\n{weight})\n\nBoth should be the same.'
e.args = [err]
raise e
```
The device error provides a much more readable error when `a` and `b` were on two different devices. An situation is below:
```python
inp = torch.rand().cuda()
model = model.cpu()
try:
_ = model(inp)
except Exception as e:
device_error(e, 'Input type', 'Model weights')
```
And our new log:
```bash
---------------------------------------------------------------------------
RuntimeError Traceback (most recent call last)
<ipython-input-28-981e0ace9c38> in <module>()
2 model(x)
3 except Exception as e:
----> 4 device_error(e, 'Input type', 'Model weights')
10 frames
/usr/local/lib/python3.7/dist-packages/torch/tensor.py in __torch_function__(cls, func, types, args, kwargs)
993
994 with _C.DisableTorchFunction():
--> 995 ret = func(*args, **kwargs)
996 return _convert(ret, cls)
997
RuntimeError: Mismatch between weight types
Input type has type: (torch.cuda.FloatTensor)
Model weights have type: (torch.FloatTensor)
Both should be the same.
```
```
#export
def hook_fn(m, i):
"Simple hook fn to return the layer"
return m
#export
class PreHook(Hook):
"Creates and registers a hook on `m` with `hook_func` as a forward pre_hook"
def __init__(self, m, hook_func, is_forward=True, detach=True, cpu=False, gather=False):
store_attr('hook_func,detach,cpu,gather')
f = m.register_forward_pre_hook if is_forward else m.register_backward_pre_hook
self.hook = f(self.hook_fn)
self.stored,self.removed = None, False
def hook_fn(self, module, inp):
"Applies `hook_fn` to `module` and `inp`"
if self.detach:
inp = to_detach(inp, cpu=self.cpu, gather=self.gather)
self.stored = self.hook_func(module, inp)
#export
class ForwardHooks():
"Create several forward-hooks on the modules in `ms` with `hook_func`"
def __init__(self, ms, hook_func, is_forward=True, detach=True, cpu=False):
self.hooks = []
for i, m in enumerate(flatten_model(ms)):
self.hooks.append(PreHook(m, hook_func, is_forward, detach, cpu))
#export
def hook_outputs(modules, detach=True, cpu=False, grad=False):
"Return `Hooks` that store activations of all `modules` in `self.stored`"
return ForwardHooks(modules, hook_fn, detach=detach, cpu=cpu, is_forward=not grad)
```
By using forward hooks, we can locate our problem layers when they arrive rather than trying to figure out which one it is through a list of confusing errors.
For this tutorial and testing we'll purposefully write a broken model:
```
from torch import nn
m = nn.Sequential(
nn.Conv2d(3,3,1),
nn.ReLU(),
nn.Linear(3,2)
)
#export
def layer_error(e:Exception, model, *inp) -> Exception:
"""
Verbose error for when there is a size mismatch between some input and the model.
`model` should be any torch model
`inp` is the input that went to the model
"""
args = e.args[0].replace("Expected", "Model expected")
hooks = hook_outputs(model)
try:
_ = model(*inp)
except:
pass
finally:
layers,num = [], 0
for i, layer in enumerate(hooks.hooks):
if layer.stored is not None:
layers.append(layer.stored)
num += 1
layer = layers[-1]
[h.remove() for h in hooks.hooks]
e.args = [f'Size mismatch between input tensors and what the model expects\n{"-"*76}\nLayer: {i}, {layer}\nError: {args}']
raise e
```
`layer_error` can be used anywhere that you want to check that the inputs are right for some model.
Let's use our `m` model from earlier to show an example:
```
#failing
inp = torch.rand(5,2, 3)
try:
m(inp)
except Exception as e:
layer_error(e, m, inp)
```
This will also work with multi-input and multi-output models:
```
class DoubleInputModel(nn.Sequential):
def __init__(self):
super().__init__()
self.layers = nn.Sequential(nn.Conv2d(3,3,1),
nn.ReLU(),
nn.Linear(3,2))
def forward(self, a, b):
return self.layers(a), self.layers(b)
model = DoubleInputModel()
#failing
inp = torch.rand(5,2, 3)
try:
model(inp, inp)
except Exception as e:
layer_error(e, model, inp, inp)
```
Much more readable!
| github_jupyter |
<a href="https://colab.research.google.com/github/aly202012/Teaching/blob/master/Classification.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
Binary Classification
```
import pandas as pd
# load the training dataset
diabetes = pd.read_csv('diabetes.csv')
diabetes.head()
#diabetes.shape
# (2000, 9)
# هنا يتم الفصل بين المتغيرات وبين الهدف
# Separate features and labels
features = ['Pregnancies','Glucose','BloodPressure','SkinThickness','Insulin','BMI','DiabetesPedigreeFunction','Age']
label = 'Outcome'
X, y = diabetes[features].values, diabetes[label].values
# for n in range(0,10):
# هنا تم التحويل او العرض علي شكل ليست
for n in range(10):
print("Patient", str(n+1), "\n Features:",list(X[n]), "\n Label:", y[n])
from matplotlib import pyplot as plt
#%matplotlib inline
features = ['Pregnancies','Glucose','BloodPressure','SkinThickness','Insulin','BMI','DiabetesPedigreeFunction','Age']
for col in features:
diabetes.boxplot(column=col, by='Outcome', figsize=(6,6))
plt.title(col)
plt.show()
# بدايه تقسيم البيانات
from sklearn.model_selection import train_test_split
# Split data 70%-30% into training set and test set
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.30, random_state=0)
print ('Training cases: %d\nTest cases: %d' % (X_train.shape[0], X_test.shape[0]))
# نلاحظ ان عدد الصفوف لدينا يساوي 2000
# بدايه تدريب النموذج
# Train the model
from sklearn.linear_model import LogisticRegression
# Set regularization rate
reg = 0.01
# train a logistic regression model on the training set
model = LogisticRegression(C=1/reg, solver="liblinear").fit(X_train, y_train)
print (model)
# بدايه الاختبار للموذج والتي تم الاحتفاظ بها في الاكس تست
predictions = model.predict(X_test)
print('Predicted labels: ', predictions)
print('Actual labels: ' ,y_test)
# لمقارنه التنبؤات بشكل افضل
from sklearn.metrics import accuracy_score
print('Accuracy: ', accuracy_score(y_test, predictions))
# لحسن الحظ ، هناك بعض المقاييس الأخرى التي تكشف المزيد عن كيفية أداء نموذجنا. يتضمن
# Scikit-Learn القدرة على إنشاء تقرير تصنيف يوفر رؤية أكثر من الدقة الأولية وحدها.
from sklearn. metrics import classification_report
print(classification_report(y_test, predictions))
#Precision: Of the predictions the model made for this class, what proportion were correct?
#Recall: Out of all of the instances of this class in the test dataset, how many did the model identify?
#F1-Score: An average metric that takes both precision and recall into account.
#Support: How many instances of this class are there in the test dataset?
#الدقة : ما هي النسبة الصحيحة من التنبؤات التي قدمها النموذج لهذه الفئة؟
# أذكر : من بين جميع مثيلات هذه الفئة في مجموعة بيانات الاختبار ، كم عددًا حددها النموذج؟
#F1- نقاط: مقياس متوسط يأخذ الدقة والاسترجاع في الاعتبار.
# الدعم : كم عدد حالات هذه الفئة الموجودة في مجموعة بيانات الاختبار؟
from sklearn.metrics import precision_score, recall_score
print("Overall Precision:",precision_score(y_test, predictions))
print("Overall Recall:",recall_score(y_test, predictions))
# مصفوفه التشتت
from sklearn.metrics import confusion_matrix
# Print the confusion matrix
cm = confusion_matrix(y_test, predictions)
print (cm)
# احتمال ان الاجابه صحيحه ما بين خانتين
y_scores = model.predict_proba(X_test)
print(y_scores)
```
تتمثل إحدى الطرق الشائعة لتقييم المصنف في فحص المعدل الإيجابي الحقيقي (وهو اسم آخر للتذكر) والمعدل الإيجابي الخاطئ لمجموعة من العتبات المحتملة. ثم يتم رسم هذه الأسعار مقابل جميع العتبات الممكنة لتشكيل مخطط يُعرف باسم مخطط خصائص عامل التشغيل المستلم (ROC) ، مثل هذا:
A common way to evaluate a classifier is to examine the true positive rate (which is another name for recall) and the false positive rate for a range of possible thresholds. These rates are then plotted against all possible thresholds to form a chart known as a received operator characteristic (ROC) chart, like this:
```
from sklearn.metrics import roc_curve
from sklearn.metrics import confusion_matrix
import matplotlib
import matplotlib.pyplot as plt
%matplotlib inline
# calculate ROC curve
fpr, tpr, thresholds = roc_curve(y_test, y_scores[:,1])
# plot ROC curve
fig = plt.figure(figsize=(6, 6))
# Plot the diagonal 50% line
plt.plot([0, 1], [0, 1], 'k--')
# Plot the FPR and TPR achieved by our model
plt.plot(fpr, tpr)
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('ROC Curve')
plt.show()
```
المنطقة الواقعة تحت المنحنى (AUC) هي قيمة بين 0 و 1 تحدد الأداء العام للنموذج. كلما كانت هذه القيمة أقرب إلى 1 ، كان النموذج أفضل. مرة أخرى ، يتضمن scikit-Learn وظيفة لحساب هذا المقياس.
```
from sklearn.metrics import roc_auc_score
auc = roc_auc_score(y_test,y_scores[:,1])
print('AUC: ' + str(auc))
```
In practice, it's common to perform some preprocessing of the data to make it easier for the algorithm to fit a model to it. There's a huge range of preprocessing transformations you can perform to get your data ready for modeling, but we'll limit ourselves to a few common techniques:
Scaling numeric features so they're on the same scale. This prevents features with large values from producing coefficients that disproportionately affect the predictions.
Encoding categorical variables. For example, by using a one hot encoding technique you can create individual binary (true/false) features for each possible category value.
```
# لرفع الكفاءه للخوارزم يجب عمل معالجه للبيانات بشكل افضل واكثر
# Train the model
from sklearn.compose import ColumnTransformer
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler, OneHotEncoder
from sklearn.linear_model import LogisticRegression
import numpy as np
# Define preprocessing for numeric columns (normalize them so they're on the same scale)
numeric_features = [0,1,2,3,4,5,6]
numeric_transformer = Pipeline(steps=[
('scaler', StandardScaler())])
# Define preprocessing for categorical features (encode the Age column)
categorical_features = [7]
categorical_transformer = Pipeline(steps=[
('onehot', OneHotEncoder(handle_unknown='ignore'))])
# Combine preprocessing steps
preprocessor = ColumnTransformer(
transformers=[
('num', numeric_transformer, numeric_features),
('cat', categorical_transformer, categorical_features)])
# Create preprocessing and training pipeline
pipeline = Pipeline(steps=[('preprocessor', preprocessor),
('logregressor', LogisticRegression(C=1/reg, solver="liblinear"))])
# fit the pipeline to train a logistic regression model on the training set
model = pipeline.fit(X_train, (y_train))
print (model)
```
دعنا نستخدم النموذج الذي تم تدريبه بواسطة خط الأنابيب هذا للتنبؤ بتسميات مجموعة الاختبار الخاصة بنا ، ومقارنة مقاييس الأداء بالنموذج الأساسي الذي أنشأناه سابقًا.
```
# Get predictions from test data
predictions = model.predict(X_test)
y_scores = model.predict_proba(X_test)
# Get evaluation metrics
cm = confusion_matrix(y_test, predictions)
print ('Confusion Matrix:\n',cm, '\n')
print('Accuracy:', accuracy_score(y_test, predictions))
print("Overall Precision:",precision_score(y_test, predictions))
print("Overall Recall:",recall_score(y_test, predictions))
auc = roc_auc_score(y_test,y_scores[:,1])
print('AUC: ' + str(auc))
# calculate ROC curve
fpr, tpr, thresholds = roc_curve(y_test, y_scores[:,1])
# plot ROC curve
fig = plt.figure(figsize=(6, 6))
# Plot the diagonal 50% line
plt.plot([0, 1], [0, 1], 'k--')
# Plot the FPR and TPR achieved by our model
plt.plot(fpr, tpr)
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('ROC Curve')
plt.show()
# Try a different algorithm
```
Support Vector Machine algorithms: Algorithms that define a hyperplane that separates classes.
Tree-based algorithms: Algorithms that build a decision tree to reach a prediction
Ensemble algorithms: Algorithms that combine the outputs of multiple base algorithms to improve generalizability.
```
from sklearn.ensemble import RandomForestClassifier
# Create preprocessing and training pipeline
pipeline = Pipeline(steps=[('preprocessor', preprocessor),
('logregressor', RandomForestClassifier(n_estimators=100))])
# fit the pipeline to train a random forest model on the training set
model = pipeline.fit(X_train, (y_train))
print (model)
# القاء نظره علي مقاييس الاداء
predictions = model.predict(X_test)
y_scores = model.predict_proba(X_test)
cm = confusion_matrix(y_test, predictions)
print ('Confusion Matrix:\n',cm, '\n')
print('Accuracy:', accuracy_score(y_test, predictions))
print("Overall Precision:",precision_score(y_test, predictions))
print("Overall Recall:",recall_score(y_test, predictions))
auc = roc_auc_score(y_test,y_scores[:,1])
print('\nAUC: ' + str(auc))
# calculate ROC curve
fpr, tpr, thresholds = roc_curve(y_test, y_scores[:,1])
# plot ROC curve
fig = plt.figure(figsize=(6, 6))
# Plot the diagonal 50% line
plt.plot([0, 1], [0, 1], 'k--')
# Plot the FPR and TPR achieved by our model
plt.plot(fpr, tpr)
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('ROC Curve')
plt.show()
```
Use the Model for Inferencing
استخدام النموذج للاستدلال
الآن بعد أن أصبح لدينا نموذجًا مدربًا مفيدًا بشكل معقول ، يمكننا حفظه لاستخدامه لاحقًا للتنبؤ بتسميات البيانات الجديدة:
```
import joblib
# Save the model as a pickle file
filename = 'diabetes_model.pkl'
joblib.dump(model, filename)
# When we have some new observations for which the label is unknown,
# we can load the model and use it to predict values for the unknown label:
# Load the model from the file
model = joblib.load(filename)
# predict on a new sample
# The model accepts an array of feature arrays (so you can predict the classes of multiple patients in a single call)
# We'll create an array with a single array of features, representing one patient
X_new = np.array([[2,180,74,24,21,23.9091702,1.488172308,22]])
print ('New sample: {}'.format(list(X_new[0])))
# Get a prediction
pred = model.predict(X_new)
# The model returns an array of predictions - one for each set of features submitted
# In our case, we only submitted one patient, so our prediction is the first one in the resulting array.
print('Predicted class is {}'.format(pred[0]))
# Multiclass Classification
import pandas as pd
# load the training dataset
penguins = pd.read_csv('penguins.csv')
# Display a random sample of 10 observations
sample = penguins.sample(10)
sample
# (10, 9)
# ما تبقي من الكود هو لمجموعه من البيانات متعدده التصنيف
# سبق وتدربنا علي هذا النوع كثيرا
# لذا لن نكمل
```
| github_jupyter |
# Overlays
Spatial overlays allow you to compare two GeoDataFrames containing polygon or multipolygon geometries
and create a new GeoDataFrame with the new geometries representing the spatial combination *and*
merged properties. This allows you to answer questions like
> What are the demographics of the census tracts within 1000 ft of the highway?
The basic idea is demonstrated by the graphic below but keep in mind that overlays operate at the dataframe level,
not on individual geometries, and the properties from both are retained
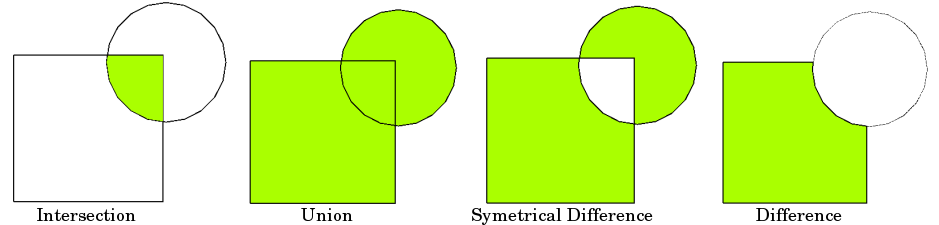
Now we can load up two GeoDataFrames containing (multi)polygon geometries...
```
%matplotlib inline
from shapely.geometry import Point
from geopandas import datasets, GeoDataFrame, read_file
from geopandas.tools import overlay
# NYC Boros
zippath = datasets.get_path('nybb')
polydf = read_file(zippath)
# Generate some circles
b = [int(x) for x in polydf.total_bounds]
N = 10
polydf2 = GeoDataFrame([
{'geometry': Point(x, y).buffer(10000), 'value1': x + y, 'value2': x - y}
for x, y in zip(range(b[0], b[2], int((b[2] - b[0]) / N)),
range(b[1], b[3], int((b[3] - b[1]) / N)))])
```
The first dataframe contains multipolygons of the NYC boros
```
polydf.plot()
```
And the second GeoDataFrame is a sequentially generated set of circles in the same geographic space. We'll plot these with a [different color palette](https://matplotlib.org/examples/color/colormaps_reference.html).
```
polydf2.plot(cmap='tab20b')
```
The `geopandas.tools.overlay` function takes three arguments:
* df1
* df2
* how
Where `how` can be one of:
['intersection',
'union',
'identity',
'symmetric_difference',
'difference']
So let's identify the areas (and attributes) where both dataframes intersect using the `overlay` tool.
```
from geopandas.tools import overlay
newdf = overlay(polydf, polydf2, how="intersection")
newdf.plot(cmap='tab20b')
```
And take a look at the attributes; we see that the attributes from both of the original GeoDataFrames are retained.
```
polydf.head()
polydf2.head()
newdf.head()
```
Now let's look at the other `how` operations:
```
newdf = overlay(polydf, polydf2, how="union")
newdf.plot(cmap='tab20b')
newdf = overlay(polydf, polydf2, how="identity")
newdf.plot(cmap='tab20b')
newdf = overlay(polydf, polydf2, how="symmetric_difference")
newdf.plot(cmap='tab20b')
newdf = overlay(polydf, polydf2, how="difference")
newdf.plot(cmap='tab20b')
```
| github_jupyter |
# Using AXI GPIO with PYNQ
## Goal
The aim of this notebook is to show how to use AXI GPIO from PYNQ.
Multiple AXI GPIO controllers can be implemented in the programmable logic and used to control internal or external GPIO signals.
## Hardware design
This example uses a bitstream that connects three AXI GPIO controllers to the LEDs, buttons, and switches and can be used with the PYNQ-Z1 or PYNQ-Z2 board. (Each AXI GPIO controller has 2 channels, so multiple peripherals could be controlled from one AXI GPIO IP, but for simplicity and demonstration purposes, separate AXI GPIO controllers are used.

### 1. Download the tutorial overlay
The `axi_gpio.bit` and `axi_gpio.hwh` files can be found in the bitstreams directory local to this folder.
The bitstream can be downloaded by passing the relative path to the Overlay class.
* Check the bitstream and .tcl exists in the bitstream directory
```
!dir ./bitstream/axi_gpio.*
```
* Download the bitstream
```
from pynq import Overlay
axi_gpio_design = Overlay("./bitstream/axi_gpio.bit")
```
Check the IP Dictionary for this design. The IP dictionary lists AXI IP in the design, and for this example will list the AXI GPIO controllers for the buttons, LEDs, and switches. The Physical address, the address range and IP type will be listed. If any interrupts, or GPIO were connected to the PS, they would also be reported.
```
axi_gpio_design.ip_dict
hex(axi_gpio_design.ip_dict["buttons"]["phys_addr"])
```
## AxiGPIO class
The PYNQ AxiGPIO class will be used to access the AXI GPIO controllers.
### 1. Controlling the switches and push-buttons
The instances can be found and referenced from the IP dictionary.
```
from pynq.lib import AxiGPIO
buttons_instance = axi_gpio_design.ip_dict['buttons']
buttons = AxiGPIO(buttons_instance).channel1
buttons.read()
```
The buttons controller is connected to all four user push-buttons on the board (BTN0 to BTN3). Try pressing any combination of the buttons and rerunning the cell above.
The AXI GPIO controller for the switches can be used in a similar way:
```
switches_instance = axi_gpio_design.ip_dict['switches']
switches = AxiGPIO(switches_instance).channel1
print(f"Switches: {switches.read()}")
```
### 2. Controlling the LEDs
The LEDs can be used in a similar way.
```
led_instance = axi_gpio_design.ip_dict['leds']
led = AxiGPIO(led_instance).channel1
```
The outputs can be addressed using a slice.
```
led[0:4].write(0x1)
from time import sleep
led[0:4].write(0x3)
sleep(1)
led[0:4].write(0x7)
sleep(1)
led[0:4].write(0xf)
```
* Turn off the LEDs
```
led[0:4].off()
```
### 3 Putting it together
Run a loop to set the LEDs to the value of the pushbuttons.
Before executing the next cell, make sure Switch 0 (SW0) is "on". While the loop is running, press a push-button and notice the corresponding LED turns on. To exist the loop, change Switch 0 to off.
```
while((switches.read() & 0x1) == 1):
led[0:4].write(buttons.read())
```
| github_jupyter |
```
import numpy as np
import pandas as pd
pd.set_option('display.float_format', lambda x: '%.3f' % x)
pd.options.mode.chained_assignment = None
import xgboost as xgb
from xgboost import XGBClassifier
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
%matplotlib inline
import matplotlib
#matplotlib.use('agg')
matplotlib.style.use('ggplot')
import pickle as pkl
from matplotlib import pyplot as plt
from collections import Counter
from functools import reduce
import random
random.seed(1991)
import glob
novel_compounds_list = pkl.load( open( "/data/dharp/compounding/datasets/novel_compounds_list.pkl", "rb" ) )
m, h = zip(*novel_compounds_list)
heads_list=list(set(h))
modifiers_list=list(set(m))
constituents=pd.read_pickle("/data/dharp/compounding/datasets/constituents_CompoundAgnostic_DecadeAgnostic_300.pkl")
#constituents.index.names=['constituent','decade']
#constituents.reset_index(inplace=True)
constituents.info()
#constituents=constituents.drop(['decade'],axis=1).groupby(['constituent']).mean()
constituents.info()
modifiers=constituents.loc[constituents.index.isin(modifiers_list)]
modifiers.index.names=['modifier']
modifiers.info()
modifiers.head()
heads=constituents.loc[constituents.index.isin(heads_list)]
heads.index.names=['head']
heads.info()
heads.head()
novel_compounds=pd.DataFrame(novel_compounds_list)
novel_compounds.columns=['modifier','head']
novel_compounds
positive_df=pd.merge(novel_compounds,heads.reset_index(),on=["head"])
positive_df=pd.merge(positive_df,modifiers.reset_index(),on=["modifier"])
positive_df['Plausibility']=True
positive_df
def neg_df_creator(file):
pkl_file=pkl.load( open(file,'rb'))
df=pd.DataFrame(pkl_file)
df.columns=['modifier','head']
negative_df=pd.merge(df,heads.reset_index(),on=["head"])
negative_df=pd.merge(negative_df,modifiers.reset_index(),on=["modifier"])
negative_df['Plausibility']=False
return negative_df
def df_joiner(files):
df_list=[]
for file in files:
neg_df=neg_df_creator(file)
whole_df=pd.concat([neg_df,positive_df])
df_list.append(whole_df)
return df_list
corrupt_modifier_files=[]
for file in glob.glob("/data/dharp/compounding/datasets/corrupt_modifier*"):
corrupt_modifier_files.append(file)
corrupt_modifiers=df_joiner(corrupt_modifier_files)
corrupt_head_files=[]
for file in glob.glob("/data/dharp/compounding/datasets/corrupt_head*"):
corrupt_head_files.append(file)
corrupt_heads=df_joiner(corrupt_head_files)
XGBClassifier()
acc_ch=[]
for i,corrupt_head in enumerate(corrupt_heads):
data=corrupt_head.drop(['modifier','head'],axis=1)
X, Y = data.iloc[:,:-1],data.iloc[:,-1]
X_train, X_test, y_train, y_test = train_test_split(X, Y, test_size=0.2, random_state=1991)
# fit model no training data
model = XGBClassifier(njobs=-1)
model.fit(X_train, y_train)
# make predictions for test data
y_pred = model.predict(X_test)
predictions = [round(value) for value in y_pred]
# evaluate predictions
print("DF",i+1)
accuracy = accuracy_score(y_test, predictions)
acc_ch.append(accuracy)
print("Accuracy: %.2f%%" % (accuracy * 100.0))
round(np.mean(acc_ch)*100,2)
round(np.std(acc_ch)*100,2)
acc_cm=[]
for i,corrupt_head in enumerate(corrupt_modifiers):
data=corrupt_head.drop(['modifier','head'],axis=1)
X, Y = data.iloc[:,:-1],data.iloc[:,-1]
X_train, X_test, y_train, y_test = train_test_split(X, Y, test_size=0.2, random_state=1991)
# fit model no training data
model = XGBClassifier(njobs=-1)
model.fit(X_train, y_train)
# make predictions for test data
y_pred = model.predict(X_test)
predictions = [round(value) for value in y_pred]
# evaluate predictions
accuracy = accuracy_score(y_test, predictions)
acc_cm.append(accuracy)
print("DF",i+1)
print("Accuracy: %.2f%%" % (accuracy * 100.0))
round(np.mean(acc_cm)*100,2)
round(np.std(acc_cm)*100,2)
```
| github_jupyter |
# Figures
```
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import gridspec
%matplotlib inline
plt.rcParams["font.family"] = "DejaVu Sans"
```
<h2 id="tocheading">Table of Contents</h2>
<div id="toc"></div>
```
%%javascript
$.getScript('https://kmahelona.github.io/ipython_notebook_goodies/ipython_notebook_toc.js')
```
## Some general stuff
- Axis labels start with a lowercase letter, e.g. _altitude_ not _Altitude_
## Bayes' Theorem Visualization
A 2-dimensional example of a Bayesian update. Figure 1.
```
import scipy.stats as stats
x, y = np.meshgrid(np.linspace(-1, 1, 201), np.linspace(-1, 1, 201))
pos = np.dstack([x, y])
n = 81
xx = np.empty([n, n], dtype=float)
yy = np.empty([n, n], dtype=float)
prior = np.empty([n, n], dtype=float)
likelihood = np.empty([n, n], dtype=float)
for i, x in enumerate(np.linspace(-1, 1, n)):
for j, y in enumerate(np.linspace(-1, 1, n)):
xx[i,j] = x
yy[i,j] = y
prior[i,j] = stats.multivariate_normal([0., 0.], [[0.2, 0.], [0., 0.2]]).pdf([x, y])
likelihood[i,j] = stats.multivariate_normal([0.1*x - 0.3, -0.3*y], [[0.6, 0.58], [0.58, 0.6]]).pdf([x, y])
posterior = likelihood*prior
prior = prior/np.max(prior)
likelihood = likelihood/np.max(likelihood)
posterior = posterior/np.max(posterior)
levels = 7
cmap = plt.get_cmap("Greys", levels)
norm = plt.Normalize(0.01, 1)
fig, (ax1, ax2, ax3) = plt.subplots(1, 3, figsize=(9, 3.2))
ax1.contour(xx, yy, prior, levels, cmap=cmap, norm=norm, zorder=-20)
ax1.contourf(xx, yy, prior, levels, cmap=cmap, norm=norm, zorder=-10)
ax1.set_title("Prior distribution", size=11)
ax2.contour(xx, yy, likelihood, levels, cmap=cmap, norm=norm, zorder=-20)
ax2.contourf(xx, yy, likelihood, levels, cmap=cmap, norm=norm, zorder=-10)
ax2.set_title("Likelihood function", size=11)
ax3.contour(xx, yy, posterior, levels, cmap=cmap, norm=norm, zorder=-20)
ax3.contourf(xx, yy, posterior, levels, cmap=cmap, norm=norm, zorder=-10)
ax3.set_title("Posterior distribution", size=11)
for ax in [ax1, ax2, ax3]:
ax.set_xticks([0])
ax.set_xticklabels([])
ax.set_yticks([0])
ax.set_yticklabels([])
fig.tight_layout()
#fig.savefig("../tex/figures/bayes_theorem.pdf")
```
## Bayesian Linear Regression
Regression model predictions of 3 different models illustrating the limits of the posterior distribution's uncertainty estimate. Figure 3.
```
from regression import LinearRegression
def flatten(*args):
return (arg.flatten() for arg in args)
np.random.seed(2)
cov = 0.2**2
x = np.vstack([
np.random.normal(1.8, size=[6,1]),
np.random.normal(-1.8, size=[6,1])
])
y = np.sin(x) + np.random.normal(scale=np.sqrt(cov), size=x.shape)
x_ref = np.linspace(-5, 5, 100).reshape(-1, 1)
bases = [
lambda x: np.hstack([1., x, x**2, x**3, x**4, x**5]),
lambda x: np.hstack([1., x]),
lambda x: np.hstack([np.exp(-0.5*(x-μ)**2) for μ in range(-3, 4)])
]
alphas = [0.1, 0.1, 1]
fig, (ax1, ax2, ax3) = plt.subplots(1, 3, figsize=(8, 2.7))
for ax, basis, alpha in zip([ax1, ax2, ax3], bases, alphas):
l = LinearRegression(basis, β=1/cov, α=alpha)
l.fit(x, y)
yy, std = flatten(*l.predict(x_ref, samples="std"))
ax.plot(x_ref, yy, color="k", linewidth=2, zorder=-10)
ax.fill_between(x_ref.flatten(), yy-std, yy+std, color="#BBBBBB", zorder=-40)
ax.fill_between(x_ref.flatten(), yy-std*3, yy+std*3, color="#E0E0E0", zorder=-50)
ax.scatter(*flatten(x, y), 40, edgecolor="#000000", facecolor="#FFFFFF", linewidth=1.2)
ax.set_xlim(-5, 5)
ax.set_ylim(-2, 2)
ax.set_yticks([-2, -1, 0, 1, 2])
ax.set_yticklabels(["-2", "-1", "0", "1", "2"])
ax.set_xlabel("predictor")
ax.label_outer()
ax1.set_ylabel("target")
fig.tight_layout()
#fig.savefig("../tex/figures/bayesian_regression.pdf")
```
## Absorption in the Microwave Region
Absorption coefficients in the microwave region for an example atmospheric layer. Figure 4.
```
import mwrt
import formulas as fml
νs = np.linspace(10, 80, 1000)
T = 273.15
θ = 300/T
p = 850.
rh = 100.
esat = fml.esat(T=T)
e = fml.e(esat=esat, RH=rh)
qliq = 0.0001
ylim = 8.0e-7, 1.0e-2
hatpro_o2 = np.array([22.24, 23.04, 23.84, 25.44, 26.24, 27.84, 31.40])
hatpro_hu = np.array([51.26, 52.28, 53.86, 54.94, 56.66, 57.30, 58.00])
def as_absorp(f):
def absorp(ν, *args, **kwargs):
return 4*np.pi*ν*1.0e9/299792458.*np.imag(f(ν, *args, **kwargs))
return absorp
gas_absorp = as_absorp(mwrt.liebe93.refractivity_gaseous)
h2o_absorp = as_absorp(mwrt.liebe93.refractivity_H2O)
cld_absorp = as_absorp(mwrt.tkc.refractivity_lwc)
α_gas = gas_absorp(νs, θ, p-e, e)
α_h2o = h2o_absorp(νs, θ, p-e, e)
α_dry = α_gas - α_h2o
α_cld = qliq * fml.ρ(T=T, p=p, e=e) * cld_absorp(νs, θ)
α_ho2 = gas_absorp(hatpro_o2, θ, p-e, e) + qliq * fml.ρ(T=T, p=p, e=e) * cld_absorp(hatpro_o2, θ)
α_hhu = gas_absorp(hatpro_hu, θ, p-e, e) + qliq * fml.ρ(T=T, p=p, e=e) * cld_absorp(hatpro_hu, θ)
fig, ax = plt.subplots(1, 1, figsize=(8, 3.5))
ax.semilogy(νs, α_gas + α_cld, color="#000000", linewidth=2.5, zorder=-10, label="total")
ax.semilogy(νs, α_cld, linewidth=1.5, color="#666666", zorder=-50, label="cloud")
ax.semilogy(νs, α_dry, linewidth=1.5, color="#33a02c", zorder=-40, label="dry")
ax.semilogy(νs, α_h2o, linewidth=1.5, color="#1f78b4", zorder=-30, label="H₂O")
ax.scatter(hatpro_o2, α_ho2*1.55, 90, marker="|", zorder=-5, color="#000000")
ax.scatter(hatpro_o2, α_ho2*1.35, 20, marker="v", zorder=-5, color="#000000")
ax.scatter(hatpro_hu, α_hhu*1.55, 90, marker="|", zorder=-5, color="#000000")
ax.scatter(hatpro_hu, α_hhu*1.35, 20, marker="v", zorder=-5, color="#000000")
ax.legend(loc="upper left", ncol=2)
ax.text(24, 1.3e-4, "K band", ha="center", fontsize=12)
ax.text(60, 6.0e-4, "V band", ha="center", fontsize=12)
ax.set_xlabel("frequency [GHz]")
ax.set_ylabel("absorption [1/m]")
ax.set_ylim(*ylim)
ax.set_xlim(min(νs), max(νs))
fig.tight_layout()
#fig.savefig("../tex/figures/absorption.pdf")
```
## Verification of Gaussian Assumption
How good is the Gaussian assumption for temperature and total water content? An example. Figure 5.
```
from scipy.stats import norm
from db_tools import read_csv_profiles
T = read_csv_profiles("../data/unified/T_raso.csv")
qvap = read_csv_profiles("../data/unified/qvap_raso.csv")
qliq = read_csv_profiles("../data/unified/qliq_raso.csv")
q = qvap + qliq
lnq = np.log(q)
level = "z=1366m"
fig, (ax1, ax2, ax3) = plt.subplots(1, 3, figsize=(7.5, 2.3))
counts, *_ = ax1.hist(T[level].values, bins=np.linspace(250, 300, 15), edgecolor="#666666", color="#BBBBBB", linewidth=1.5)
grd = np.linspace(248, 302, 70)
pdf = norm(*norm.fit(T[level].values)).pdf(grd)
ax1.plot(grd, pdf/pdf.max()*counts.max(), color="k", linewidth=1.5)
ax1.set_xticks([250, 260, 270, 280, 290, 300])
ax1.set_xlim(248, 302)
ax1.set_title("T [K]", loc="right", size=10)
counts, *_ = ax2.hist(q[level].values, bins=np.linspace(0, 0.015, 20), edgecolor="#666666", color="#BBBBBB", linewidth=1.5)
ax2.set_xticks([0., 0.005, 0.01, 0.015])
grd = np.linspace(-0.001, 0.016, 70)
pdf = norm(*norm.fit(q[level].values)).pdf(grd)
ax2.plot(grd, pdf/pdf.max()*counts.max(), color="k", linewidth=1.5)
ax2.set_xlim(-0.001, 0.016)
ax2.set_title("q [kg/kg]", loc="right", size=10)
counts, *_ = ax3.hist(lnq[level].values, bins=np.linspace(-8, -3.5, 20), edgecolor="#666666", color="#BBBBBB", linewidth=1.5)
grd = np.linspace(-8.2, -3.2, 70)
pdf = norm(*norm.fit(lnq[level].values)).pdf(grd)
ax3.plot(grd, pdf/pdf.max()*counts.max(), color="k", linewidth=1.5)
ax3.set_xticks([-8, -7, -6, -5, -4])
ax3.set_xlim(-8.2, -3.2)
ax3.set_title("ln(q)", loc="right", size=10)
for ax in [ax1, ax2, ax3]:
ax.set_title("1366 m", size=10, loc="left")
ax.set_yticks([])
ax1.set_ylabel("normalized counts")
fig.tight_layout(pad=0.3)
#fig.savefig("../tex/figures/gauss_verification.pdf")
```
## Weighting Functions
```
from mwrt import MWRTM, LinearInterpolation
from optimal_estimation import VirtualHATPRO
import formulas as fml
z = np.logspace(np.log10(612), np.log10(12612), 150)
T = 288.15 - z * 0.0065
T[z>11000] = 216.65
p = fml.p(z=z, T=T, q=0, p0=940)
rh = 0.1 + (T-216.65)/(T[0] - 216.65) * 0.7
lnq = np.log(fml.qvap(p=p, T=T, RH=rh))
model_grid = np.logspace(np.log10(612), np.log10(12612), 3000)
itp = LinearInterpolation(source=z, target=model_grid)
```
### Multiple Frequencies at Zenith
Weighting functions associated with all HATPRO channels at zenith for an example atmosphere. Figure 6.
```
kband = VirtualHATPRO.absorptions[:7]
vband = VirtualHATPRO.absorptions[7:]
titles = ["V band, temperature",
"K band, humidity"]
colors = ["#"+c*6 for c in ["0", "2", "4", "6", "8", "A", "B"]]
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(8, 3.5))
for ax, title in zip([ax1, ax2], titles):
faps = vband if title.startswith("V") else kband
res = MWRTM.simulate_radiometer(itp, faps, angles=[0.], p=p, T=T, lnq=lnq)
jac = res.dT if title.startswith("V") else res.dlnq
jac[:,0] *= 2
jac[:,-1] *= 2
for row, color in zip(jac, colors):
ax.plot(row/np.max(jac), (z-612)/1000, color=color, linewidth=2)
ax.set_xticks([0, 0.2, 0.4, 0.6, 0.8, 1.])
ax.set_ylim(0,8)
ax.label_outer()
ax.set_title(title, loc="right", size=11)
ax1.set_ylabel("height above ground [km]")
ax1.set_xlim(-0.18, 1.05)
fig.tight_layout()
#fig.savefig("../tex/figures/jacobian_frequency.pdf")
```
### Same Frequency at Elevations
Weighting functions of 2 HATPRO channels at different elevation angles for an example atmosphere. Figure 7.
```
freqs = [54.94, 58.00]
absorp = [VirtualHATPRO.absorptions[-4], VirtualHATPRO.absorptions[-1]]
angles = [0., 60., 65., 70., 75., 80., 85.]
colors = ["#1f78b4"] + ["#"+c*6 for c in ["0", "2", "4", "6", "8", "A", "B"]]
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(8, 3.5))
for ax, freq, fap in zip([ax1, ax2], freqs, absorp):
model = MWRTM(itp, fap)
res = model(angles=angles, p=p, T=T, lnq=lnq)
res.dT[:,0] *= 2
for row, color in zip(res.dT, colors):
ax.plot(row/np.max(np.abs(res.dT)), (z-612)/1000, color=color, linewidth=2)
ax.set_ylim(0, 1.)
ax.label_outer()
ax.set_title("{:>5.2f} GHz".format(freq), loc="right", size=11)
ax1.set_ylabel("height above ground [km]")
ax1.set_xlim(0, 0.55)
ax1.set_xticks([0, 0.1, 0.2, 0.3, 0.4, 0.5])
ax2.set_xlim(0, 0.35)
ax2.set_xticks([0, 0.1, 0.2, 0.3])
fig.tight_layout()
#fig.savefig("../tex/figures/jacobian_angle.pdf")
```
## Model Intercomparison
RTM comparison with the Innsbruck raso climatology. Figure 8.
```
from db_tools import read_csv_profiles
mwrtm = read_csv_profiles("../data/unified/training/TB_mwrtm.csv")
mwrtmfap = read_csv_profiles("../data/unified/training/TB_mwrtm_fap.csv")
monortm = read_csv_profiles("../data/unified/training/TB_monortm.csv")
igmk = read_csv_profiles("../data/unified/training/TB_igmk.csv")
cloudy_raso = read_csv_profiles("../data/unified/training/cloudy_raso.csv")["cloudy"]
cloudy_igmk = read_csv_profiles("../data/unified/training/cloudy_igmk.csv")["cloudy"]
zenith = [col for col in mwrtm.columns if col.endswith("_00.0")]
data1 = mwrtm.loc[~cloudy_raso,zenith]
data2 = monortm.loc[~cloudy_raso,zenith]
data3 = igmk.loc[~cloudy_igmk,zenith]
data4 = mwrtmfap.loc[~cloudy_raso,zenith]
grid = np.arange(0, data1.shape[1])
mean12 = (data1 - data2).mean().values
mean13 = (data1 - data3).mean().values
mean14 = (data1 - data4).mean().values
mn = np.random.multivariate_normal
std12 = mn(mean=np.zeros_like(grid), cov=(data1 - data2).cov().values, size=1000).std(axis=0)
std13 = mn(mean=np.zeros_like(grid), cov=(data1 - data3).cov().values, size=1000).std(axis=0)
std14 = mn(mean=np.zeros_like(grid), cov=(data1 - data4).cov().values, size=1000).std(axis=0)
freqs = ["{:>5.2f}".format(int(col[3:8])/1000) for col in data1.columns]
gs = gridspec.GridSpec(2, 1, height_ratios=[3, 1])
fig = plt.figure(figsize=(7.5, 5.5))
ax1 = fig.add_subplot(gs[0])
ax2 = fig.add_subplot(gs[1])
ax1.bar(grid-0.375, mean12, width=0.25, color="#666666", zorder=-20, label="MWRTM - MonoRTM")
ax1.bar(grid-0.125, mean13, width=0.25, color="#BBBBBB", zorder=-20, label="MWRTM - Rosenkranz")
ax1.bar(grid+0.125, mean14, width=0.25, color="#FFFFFF", zorder=-20, label="MWRTM - MWRTM/FAP")
for ax in [ax1, ax2]:
ax.set_xticks(grid)
ax.set_xticklabels(["{:>5.2f}".format(int(col[3:8])/1000) for col in data1.columns], size=9)
ax.tick_params(bottom="off", top="off")
ax.set_yticks([-1.5, -1, -0.5, 0, 0.5, 1])
ax.set_title("1581 clear sky cases", size=10, loc="right")
ax1.hlines(0, -0.5, 13.5, color="#000000", zorder=-50)
ax1.vlines(grid+0.5, -2.6, 2.1, color="#E0E0E0", zorder=-55)
ax1.set_ylim(-1.8, 1.3)
ax1.set_title("mean model differences", size=10, loc="left")
ax1.legend(loc="lower left", fontsize=11);
ax1.set_ylabel("brightness temperature [K]")
ax2.bar(grid-0.375, std12, width=0.25, color="#666666", zorder=-20)
ax2.bar(grid-0.125, std13, width=0.25, color="#BBBBBB", zorder=-20)
ax2.bar(grid+0.125, std14, width=0.25, color="#FFFFFF", zorder=-20)
ax2.set_ylabel("br. temp. [K]")
ax2.set_xlabel("channel frequency [GHz]")
ax2.set_title("standard deviation of model differences", size=10, loc="left")
ax2.set_ylim(0, 0.9)
ax2.vlines(grid+0.5, 0, 0.9, color="#E0E0E0", zorder=-55)
fig.tight_layout()
#fig.savefig("../tex/figures/model_comparison.pdf")
```
## Model Bias
RTM vs. HATPRO with actual radiometer measurements. Figure 12.
```
from db_tools import read_csv_mean
mwrtm = read_csv_mean("../data/unified/priors/TB_mwrtm_bias.csv")
mwrtmfap = read_csv_mean("../data/unified/priors/TB_mwrtm_fap_bias.csv")
monortm = read_csv_mean("../data/unified/priors/TB_monortm_bias.csv")
zenith = [col for col in mwrtm.index if col.endswith("_00.0")]
freqs = ["{:>5.2f}".format(int(col[3:8])/1000) for col in zenith]
grid = np.arange(0, len(zenith))
fig, ax = plt.subplots(1, 1, figsize=(7.5, 3.5))
ax.bar(grid-0.375, monortm.ix[zenith], width=0.25, color="#666666", zorder=-20, label="MonoRTM")
ax.bar(grid-0.125, mwrtm.ix[zenith], width=0.25, color="#BBBBBB", zorder=-20, label="MWRTM")
ax.bar(grid+0.125, mwrtmfap.ix[zenith], width=0.25, color="#FFFFFF", zorder=-20, label="MWRTM/FAP")
ax.set_xticks(grid)
ax.set_xticklabels(["{:>5.2f}".format(int(col[3:8])/1000) for col in zenith], size=9)
ax.tick_params(bottom="off", top="off")
#ax.set_yticks([-1.5, -1, -0.5, 0, 0.5, 1])
ax.set_title("10 clear sky cases", size=10, loc="right")
ax.hlines(0, -0.5, 13.5, color="#000000", zorder=-50)
ax.vlines(grid+0.5, -0.7, 3.7, color="#E0E0E0", zorder=-55)
ax.set_ylim(-0.7, 3.7)
ax.set_title("RTM - HATPRO bias", size=10, loc="left")
ax.legend(loc="upper right", fontsize=11);
ax.set_ylabel("brightness temperature [K]")
ax.set_xlabel("channel frequency [GHz]")
fig.tight_layout()
#fig.savefig("../tex/figures/model_bias.pdf")
```
## Prior Distributions
```
import datetime as dt
from db_tools import iter_profiles, read_csv_covariance, read_csv_mean
from optimal_estimation import rgrid
```
### COSMO7
Example of a COSMO-7 prior distribution. Figure 11.
```
profiles = iter_profiles("../data/unified/priors/<VAR>_cosmo7+00+06_mean.csv")
Tcov = read_csv_covariance("../data/unified/priors/T_cosmo7+00+06_cov.csv")
lnqcov = read_csv_covariance("../data/unified/priors/lnq_cosmo7+00+06_cov.csv")
for valid, df in profiles:
if valid == dt.datetime(2015, 9, 11, 3, 48):
break
T = df["T"].values
T_rand = np.random.multivariate_normal(mean=T, cov=Tcov.values, size=1000)
T_std = np.std(T_rand, axis=0)
lnq = df["lnq"].values
lnq_rand = np.random.multivariate_normal(mean=lnq, cov=lnqcov.values, size=1000)
lnq_std = np.std(lnq_rand, axis=0)
z = (rgrid - rgrid[0])/1000
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(8, 3))
ax1.plot(T, z, color="#000000", linewidth=2, label="mean", zorder=-10)
ax1.fill_betweenx(z, T-T_std, T+T_std, color="#BBBBBB", label="1σ", zorder=-20)
ax1.fill_betweenx(z, T-2*T_std, T+2*T_std, color="#E0E0E0", label="2σ", zorder=-30)
ax1.set_ylim(0, 6)
ax1.set_xlim(243, 292)
ax1.set_title("COSMO-7 prior distribution", loc="left", size=11)
ax1.set_ylabel("height above ground [km]")
ax1.set_xlabel("temperature [K]")
ax2.plot(np.exp(lnq), z, color="#000000", linewidth=2, label="mean", zorder=-10)
ax2.fill_betweenx(z, np.exp(lnq-lnq_std), np.exp(lnq+lnq_std), color="#BBBBBB", label="1σ", zorder=-20)
ax2.fill_betweenx(z, np.exp(lnq-2*lnq_std), np.exp(lnq+2*lnq_std), color="#E0E0E0", label="2σ", zorder=-30)
ax2.set_xticks([0, 0.002, 0.004, 0.006, 0.008, 0.01])
ax2.set_xticklabels(["0", "2", "4", "6", "8", "10"])
ax2.set_ylim(0, 6)
ax2.set_xlim(0, 0.0102)
ax2.set_xlabel("specific water content [g/kg]")
ax2.set_title(valid.strftime("%Y-%m-%d %H:%M UTC"), loc="right", size=11)
ax2.label_outer()
fig.tight_layout(pad=0.4)
#fig.savefig("../tex/figures/cosmo7_prior.pdf")
```
### Radiosonde Climatology
Mean and standard deviation of the radiosonde climatology. Figure 10.
```
T = read_csv_mean("../data/unified/priors/T_rasoclim_mean.csv").values
Tcov = read_csv_covariance("../data/unified/priors/T_rasoclim_cov.csv").values
lnq = read_csv_mean("../data/unified/priors/lnq_rasoclim_mean.csv").values
lnqcov = read_csv_covariance("../data/unified/priors/lnq_rasoclim_cov.csv").values
T_rand = np.random.multivariate_normal(mean=T, cov=Tcov, size=1000)
T_std = np.std(T_rand, axis=0)
lnq_rand = np.random.multivariate_normal(mean=lnq, cov=lnqcov, size=1000)
lnq_std = np.std(lnq_rand, axis=0)
z = (rgrid - rgrid[0])/1000
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(8, 3))
ax1.plot(T, z, color="#000000", linewidth=2, label="mean", zorder=-10)
ax1.fill_betweenx(z, T-T_std, T+T_std, color="#BBBBBB", label="1σ", zorder=-20)
ax1.fill_betweenx(z, T-2*T_std, T+2*T_std, color="#E0E0E0", label="2σ", zorder=-30)
ax1.set_ylim(0, 12)
ax1.set_xlim(202, 297)
ax1.set_title("Radiosonde climatology", loc="left", size=11)
ax1.set_ylabel("height above ground [km]")
ax1.set_xlabel("temperature [K]")
ax2.plot(np.exp(lnq), z, color="#000000", linewidth=2, label="mean", zorder=-10)
ax2.fill_betweenx(z, np.exp(lnq-lnq_std), np.exp(lnq+lnq_std), color="#BBBBBB", label="1σ", zorder=-20)
ax2.fill_betweenx(z, np.exp(lnq-2*lnq_std), np.exp(lnq+2*lnq_std), color="#E0E0E0", label="2σ", zorder=-30)
ax2.set_xticks([0, 0.002, 0.004, 0.006, 0.008])
ax2.set_xticklabels(["0", "2", "4", "6", "8"])
ax2.set_ylim(0, 12)
ax2.set_xlim(0., 0.0092)
ax2.set_xlabel("specific water content [g/kg]")
ax2.set_title("3561 profiles", loc="right", size=11)
ax2.label_outer()
fig.tight_layout(pad=0.4)
#fig.savefig("../tex/figures/raso_prior.pdf")
```
| github_jupyter |
# 05.04 - PARTICIPATE IN KAGGLE
```
!wget --no-cache -O init.py -q https://raw.githubusercontent.com/rramosp/20201.xai4eng/master/content/init.py
import init; init.init(force_download=False); init.get_weblink()
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
from sklearn.metrics import accuracy_score
from sklearn.model_selection import train_test_split
import local.lib.mlutils
import pandas as pd
from tensorflow.keras import Sequential
from tensorflow.keras.layers import Dense, Dropout, Flatten
%matplotlib inline
```
## We use Titanic data in [Kaggle](http://www.kaggle.com)
- Register to [Kaggle](http://www.kaggle.com)
- Enter the competition [Titanic Data at Kaggle](https://www.kaggle.com/c/titanic)
- Download the `train.csv` and `test.csv` files
- **UPLOAD THE FILES** to your notebook environment (in Colab, open the Files tab and upload)
```
d = pd.read_csv("train.csv")
print (d.shape)
d.head()
```
**Understand `NaN` values are present**
```
for i in d.columns:
print ("%20s"%i, np.sum(d[i].isna()))
d.Embarked.value_counts()
plt.hist(d.Age.dropna().values, bins=30);
```
**Remove uninformative columns**
```
del(d["PassengerId"])
del(d["Name"])
del(d["Ticket"])
del(d["Cabin"])
```
**Fix `NaN` values**
- observe the different filling policies we decide to have
```
d["Embarked"] = d.Embarked.fillna("N")
d["Age"] = d.Age.fillna(d.Age.mean())
d.head()
plt.hist(d.Age.dropna().values, bins=30);
```
**Turn categorical columns to a `one_hot` encoding**
```
def to_onehot(x):
values = np.unique(x)
r = np.r_[[np.argwhere(i==values)[0][0] for i in x]]
return np.eye(len(values))[r].astype(int)
k = to_onehot(d.Embarked.values)
k[:5]
def replace_columns_with_onehot(d, col):
k = to_onehot(d[col].values)
r = pd.DataFrame(k, columns=["%s_%d"%(col, i) for i in range(k.shape[1])], index=d.index).join(d)
del(r[col])
return r
d.head()
d = replace_columns_with_onehot(d, "Embarked")
d.head()
d = replace_columns_with_onehot(d, "Sex")
d.head()
d.shape, d.values.sum()
```
### Put all transformations together
```
def clean_titanic(d):
del(d["PassengerId"])
del(d["Name"])
del(d["Ticket"])
del(d["Cabin"])
d["Embarked"] = d.Embarked.fillna("N")
d["Fare"] = d.Fare.fillna(d.Fare.mean())
d["Age"] = d.Age.fillna(d.Age.mean())
d = replace_columns_with_onehot(d, "Embarked")
d = replace_columns_with_onehot(d, "Sex")
return d
```
**transform train and test data together**
- observe that test data **does not have** a `Survival` column. This is the result to submit to Kaggle
```
dtr = pd.read_csv("train.csv")
dts = pd.read_csv("test.csv")
lentr = len(dtr)
dtr.shape, dts.shape
dts.head()
```
**get data ready for training**
```
source_cols = [i for i in dtr.columns if i!="Survived"]
all_data = pd.concat((dtr[source_cols], dts[source_cols]))
all_data.index = range(len(all_data))
all_data = clean_titanic(all_data)
Xtr, ytr = all_data.iloc[:lentr].values, dtr["Survived"].values
Xts = all_data.iloc[lentr:].values
print (Xtr.shape, ytr.shape)
print (Xts.shape)
```
**cross validate for model selection**
```
from sklearn.ensemble import RandomForestClassifier
from sklearn.svm import SVC
from sklearn.model_selection import cross_val_score
rf = RandomForestClassifier()
print (cross_val_score(rf, Xtr, ytr))
svc = SVC()
print (cross_val_score(svc, Xtr, ytr))
```
**now train with full dataset and generate submission for Kaggle**
```
rf.fit(Xtr, ytr)
preds_ts = rf.predict(Xts)
preds_ts
```
**get predictions ready to submit to Kaggle**
- see https://www.kaggle.com/c/titanic#evaluation for file format
```
submission = pd.DataFrame([dts.PassengerId, pd.Series(preds_ts, name="Survived")]).T
submission.head()
submission.to_csv("titanic_kaggle.csv", index=False)
!head titanic_kaggle.csv
```
| github_jupyter |
## ROC Curve and ROC-AUC in Multiclass
For multiclass classification, we have 2 options:
- determine a ROC curve for each class.
- determine the overall ROC curve as the micro-average of all classes.
Let's see how to do both.
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.datasets import load_wine
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn.multiclass import OneVsRestClassifier
# to convert the 1-D target vector in to a matrix
from sklearn.preprocessing import label_binarize
from sklearn.metrics import roc_curve, roc_auc_score
from yellowbrick.classifier import ROCAUC
```
## Load data (multiclass)
```
# load data
data = load_wine()
data = pd.concat([
pd.DataFrame(data.data, columns=data.feature_names),
pd.DataFrame(data.target, columns=['target']),
], axis=1)
data.head()
# target distribution:
# multiclass and (fairly) balanced
data.target.value_counts(normalize=True)
# separate dataset into train and test
X_train, X_test, y_train, y_test = train_test_split(
data.drop(labels=['target'], axis=1), # drop the target
data['target'], # just the target
test_size=0.3,
random_state=0)
X_train.shape, X_test.shape
# the target is a vector with the 3 classes
y_test[0:10]
```
## Train ML models
The dataset we are using is very, extremely simple, so I am creating dumb models intentionally, that is few trees and very shallow for the random forests and few iterations for the logit. This is, so that we can get the most out of the PR curves by inspecting them visually.
### Random Forests
The Random Forests in sklearn are not trained as a 1 vs Rest. So in order to produce a 1 vs rest probability vector for each class, we need to wrap this estimator with another one from sklearn:
- [OneVsRestClassifier](https://scikit-learn.org/stable/modules/generated/sklearn.multiclass.OneVsRestClassifier.html)
```
# set up the model, wrapped by the OneVsRestClassifier
rf = OneVsRestClassifier(
RandomForestClassifier(
n_estimators=10, random_state=39, max_depth=1, n_jobs=4,
)
)
# train the model
rf.fit(X_train, y_train)
# produce the predictions (as probabilities)
y_train_rf = rf.predict_proba(X_train)
y_test_rf = rf.predict_proba(X_test)
# note that the predictions are an array of 3 columns
# first column: the probability of an observation of being of class 0
# second column: the probability of an observation of being of class 1
# third column: the probability of an observation of being of class 2
y_test_rf[0:10, :]
pd.DataFrame(y_test_rf).sum(axis=1)[0:10]
# The final prediction is that of the biggest probabiity
rf.predict(X_test)[0:10]
```
### Logistic Regression
The Logistic regression supports 1 vs rest automatically though its multi_class parameter:
```
# set up the model
logit = LogisticRegression(
random_state=0, multi_class='ovr', max_iter=10,
)
# train
logit.fit(X_train, y_train)
# obtain the probabilities
y_train_logit = logit.predict_proba(X_train)
y_test_logit = logit.predict_proba(X_test)
# note that the predictions are an array of 3 columns
# first column: the probability of an observation of being of class 0
# second column: the probability of an observation of being of class 1
# third column: the probability of an observation of being of class 2
y_test_logit[0:10, :]
# The final prediction is that of the biggest probabiity
logit.predict(X_test)[0:10]
```
## ROC Curve
### Per class with Sklearn
```
# with label_binarize we transform the target vector
# into a multi-label matrix, so that it matches the
# outputs of the models
# then we have 1 class per column
y_test = label_binarize(y_test, classes=[0, 1, 2])
y_test[0:10, :]
# now we determine the tpr and fpr at different thresholds
# considering only the probability vector for class 2 and the true
# target for class 2
# so we treat the problem as class 2 vs rest
fpr, tpr, thresholds = roc_curve(y_test[:, 2], y_test_rf[:, 2])
# false positive rate
fpr
# true positive rate
tpr
# threhsolds examined
thresholds
```
Go ahead and examine the precision and recall for the other classes see how these values change.
```
# now let's do these for all classes and capture the results in
# dictionaries, so we can plot the values afterwards
# determine the tpr and fpr
# at various thresholds of probability
# in a 1 vs all fashion, for each class
fpr_rf = dict()
tpr_rf = dict()
# for each class
for i in range(3):
# determine tpr and fpr at various thresholds
# in a 1 vs all fashion
fpr_rf[i], tpr_rf[i], _ = roc_curve(
y_test[:, i], y_test_rf[:, i])
fpr_rf
# plot the curves for each class
for i in range(3):
plt.plot(fpr_rf[i], tpr_rf[i], label='class {}'.format(i))
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic example')
plt.legend(loc="lower right")
plt.show()
# and now for the logistic regression
fpr_lg = dict()
tpr_lg = dict()
# for each class
for i in range(3):
# determine precision and recall at various thresholds
# in a 1 vs all fashion
fpr_lg[i], tpr_lg[i], _ = roc_curve(
y_test[:, i], y_test_logit[:, i])
plt.plot(fpr_lg[i], tpr_lg[i], label='class {}'.format(i))
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic example')
plt.legend(loc="lower right")
plt.show()
# and now, just because it is a bit difficult to compare
# between models, we plot the ROC curves class by class,
# but the 2 models in the same plot
# for each class
for i in range(3):
plt.plot(fpr_lg[i], tpr_lg[i], label='logit class {}'.format(i))
plt.plot(fpr_rf[i], tpr_rf[i], label='rf class {}'.format(i))
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic example')
plt.legend(loc="lower right")
plt.show()
```
We see that the Random Forest does a better job for all classes.
### Micro-average with sklearn
In order to do this, we concatenate all the probability vectors 1 after the other, and so we do with the real values.
```
# probability vectors for all classes in 1-d vector
y_test_rf.ravel()
# see that the unravelled prediction vector has 3 times the size
# of the origina target
len(y_test), len(y_test_rf.ravel())
# A "micro-average": quantifying score on all classes jointly
# for random forests
# Compute micro-average ROC curve and ROC area
fpr_rf["micro"], tpr_rf["micro"], _ = roc_curve(
y_test.ravel(), y_test_rf.ravel(),
)
# for logistic regression
# Compute micro-average ROC curve and ROC area
fpr_lg["micro"], tpr_lg["micro"], _ = roc_curve(
y_test.ravel(), y_test_logit.ravel(),
)
# now we plot them next to each other
i = "micro"
plt.plot(fpr_lg[i], tpr_lg[i], label='logit micro {}')
plt.plot(fpr_rf[i], tpr_rf[i], label='rf micro {}')
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic example')
plt.legend(loc="lower right")
plt.show()
```
## ROC-AUC with sklearn
```
macro_roc_auc_ovo = roc_auc_score(
y_test, y_test_rf, multi_class="ovo", average="macro")
weighted_roc_auc_ovo = roc_auc_score(
y_test, y_test_rf, multi_class="ovo", average="weighted")
macro_roc_auc_ovr = roc_auc_score(
y_test, y_test_rf, multi_class="ovr", average="macro")
weighted_roc_auc_ovr = roc_auc_score(
y_test, y_test_rf, multi_class="ovr", average="weighted")
print("One-vs-One ROC AUC scores:\n{:.6f} (macro),\n{:.6f} "
"(weighted)"
.format(macro_roc_auc_ovo, weighted_roc_auc_ovo))
print("One-vs-Rest ROC AUC scores:\n{:.6f} (macro),\n{:.6f} "
"(weighted)"
.format(macro_roc_auc_ovr, weighted_roc_auc_ovr))
macro_roc_auc_ovo = roc_auc_score(
y_test, y_test_logit, multi_class="ovo", average="macro")
weighted_roc_auc_ovo = roc_auc_score(
y_test, y_test_logit, multi_class="ovo", average="weighted")
macro_roc_auc_ovr = roc_auc_score(
y_test, y_test_logit, multi_class="ovr", average="macro")
weighted_roc_auc_ovr = roc_auc_score(
y_test, y_test_logit, multi_class="ovr", average="weighted")
print("One-vs-One ROC AUC scores:\n{:.6f} (macro),\n{:.6f} "
"(weighted)"
.format(macro_roc_auc_ovo, weighted_roc_auc_ovo))
print("One-vs-Rest ROC AUC scores:\n{:.6f} (macro),\n{:.6f} "
"(weighted)"
.format(macro_roc_auc_ovr, weighted_roc_auc_ovr))
```
## ROC curve and ROC-AUC
### Per class with Yellobrick
**Note:**
In the cells below, we are passing to Yellobrick classes a model that is already fit. When we fit() the Yellobrick class, it will check if the model is fit, in which case it will do nothing.
If we pass a model that is not fit, and a multiclass target, Yellowbrick will wrap the model automatically with a 1 vs Rest classifier.
Check Yellobrick's documentation for more details.
```
rf = RandomForestClassifier(
n_estimators=10, random_state=39, max_depth=1, n_jobs=4,
)
logit = LogisticRegression(
random_state=0, multi_class='ovr', max_iter=10,
)
# let's reconstitute the original format of
# the target for this calculations
y_test = np.argmax(y_test, axis=1)
y_test
visualizer = ROCAUC(
rf, per_class=True, cmap="cool", micro=False,
)
visualizer.fit(X_train, y_train) # Fit the training data to the visualizer
visualizer.score(X_test, y_test) # Evaluate the model on the test data
visualizer.show() # Finalize and show the figure
visualizer = ROCAUC(
logit, per_class=True, cmap="cool", micro=False, cv=0.05,
)
visualizer.fit(X_train, y_train) # Fit the training data to the visualizer
visualizer.score(X_test, y_test) # Evaluate the model on the test data
visualizer.show() # Finalize and show the figure
```
### Micro yellowbrick
- [ROCAUC](https://www.scikit-yb.org/en/latest/api/classifier/rocauc.html)
```
visualizer = ROCAUC(
rf, per_class = False, cmap="cool", micro=True,
)
visualizer.fit(X_train, y_train) # Fit the training data to the visualizer
visualizer.score(X_test, y_test) # Evaluate the model on the test data
visualizer.show() # Finalize and show the figure
visualizer = ROCAUC(
logit, per_class = False, cmap="cool", micro=True, cv=0.05,
)
visualizer.fit(X_train, y_train) # Fit the training data to the visualizer
visualizer.score(X_test, y_test) # Evaluate the model on the test data
visualizer.show() # Finalize and show the figure
```
| github_jupyter |
```
import numpy as np
from keras.models import *
from keras import backend as K
from keras.preprocessing.image import ImageDataGenerator
from model_provider import getModel
from datahandler import DataHandler
from kfold_data_loader import *
from params import *
import os
import cv2
import skimage.io as io
from tqdm import tqdm
from medpy.io import save
from math import ceil, floor
from matplotlib import pyplot as plt
from sklearn.metrics import f1_score, jaccard_similarity_score
from scipy.ndimage import _ni_support
from scipy.ndimage.morphology import distance_transform_edt, binary_erosion,\
generate_binary_structure
from skimage.morphology import cube, binary_closing
from skimage.measure import label
import warnings
warnings.filterwarnings("ignore")
plt.gray()
def destiny_directory(model_name, dice_score, post_processing = False):
if post_processing:
pre = './data/eval_pp/'+model_name+'/'
else:
pre = './data/eval/'+model_name+'/'
if dice_score >= 98:
return pre + 'dice_98_100/'
elif dice_score >= 96:
return pre + 'dice_96_98/'
elif dice_score >= 94:
return pre + 'dice_94_96/'
elif dice_score >= 92:
return pre + 'dice_92_94/'
elif dice_score >= 90:
return pre + 'dice_90_92/'
elif dice_score >= 88:
return pre + 'dice_88_90/'
elif dice_score >= 85:
return pre + 'dice_85_88'
elif dice_score >= 80:
return pre + 'dice_80_85/'
elif dice_score >= 70:
return pre + 'dice_70_80/'
elif dice_score >= 60:
return pre + 'dice_60_70/'
else:
return pre + 'dice_less_60'
def getGenerator(images, bs=1):
image_datagen = ImageDataGenerator(rescale=1./255)
image_datagen.fit(images, augment = True)
image_generator = image_datagen.flow(x = images, batch_size=bs,
shuffle = False)
return image_generator
def getDiceScore(ground_truth, prediction):
#convert to boolean values and flatten
ground_truth = np.asarray(ground_truth, dtype=np.bool).flatten()
prediction = np.asarray(prediction, dtype=np.bool).flatten()
return f1_score(ground_truth, prediction)
def hd(result, reference, voxelspacing=None, connectivity=1):
hd1 = __surface_distances(result, reference, voxelspacing, connectivity).max()
hd2 = __surface_distances(reference, result, voxelspacing, connectivity).max()
hd = max(hd1, hd2)
return hd
def hd95(result, reference, voxelspacing=None, connectivity=1):
hd1 = __surface_distances(result, reference, voxelspacing, connectivity)
hd2 = __surface_distances(reference, result, voxelspacing, connectivity)
hd95 = np.percentile(np.hstack((hd1, hd2)), 95)
return hd95
def __surface_distances(result, reference, voxelspacing=None, connectivity=1):
result = np.atleast_1d(result.astype(np.bool))
reference = np.atleast_1d(reference.astype(np.bool))
if voxelspacing is not None:
voxelspacing = _ni_support._normalize_sequence(voxelspacing, result.ndim)
voxelspacing = np.asarray(voxelspacing, dtype=np.float64)
if not voxelspacing.flags.contiguous:
voxelspacing = voxelspacing.copy()
footprint = generate_binary_structure(result.ndim, connectivity)
if 0 == np.count_nonzero(result):
raise RuntimeError('The first supplied array does not contain any binary object.')
if 0 == np.count_nonzero(reference):
raise RuntimeError('The second supplied array does not contain any binary object.')
result_border = result ^ binary_erosion(result, structure=footprint, iterations=1)
reference_border = reference ^ binary_erosion(reference, structure=footprint, iterations=1)
dt = distance_transform_edt(~reference_border, sampling=voxelspacing)
sds = dt[result_border]
return sds
image_files, mask_files = load_data_files('data/kfold_data/')
print(len(image_files))
print(len(mask_files))
skf = getKFolds(image_files, mask_files, n=10)
kfold_indices = []
for train_index, val_index in skf.split(image_files, mask_files):
kfold_indices.append({'train': train_index, 'val': val_index})
def predictMask(model, image):
image_gen = getGenerator(image)
return model.predict_generator(image_gen, steps=len(image))
def prepareForSaving(image):
#image = np.swapaxes(image, -1, 0)
image = np.moveaxis(image, 0, -1)
return image
def predictAll(model, model_name, data, num_data=0, post_processing = False):
dice_scores = []
names = []
hd_scores = []
hd95_scores = []
for image_file, mask_file in tqdm(data, total=num_data):
fname = image_file[image_file.rindex('/')+1 : image_file.rindex('.')]
image, hdr = dh.getImageData(image_file)
gt_mask, _ = dh.getImageData(mask_file, is_mask=True)
assert image.shape == gt_mask.shape
if image.shape[1] != 256:
continue
pred_mask = predictMask(model, image)
pred_mask[pred_mask>=0.5] = 1
pred_mask[pred_mask<0.5] = 0
pred_mask = np.squeeze(pred_mask)
#closing and defrag squeze of mask
if post_processing:
pred_mask = binary_closing(pred_mask, cube(2))
try:
labels = label(pred_mask)
pred_mask = (labels == np.argmax(np.bincount(labels.flat)[1:])+1).astype(int)
except:
pred_mask = pred_mask
pred_mask = np.array(pred_mask, dtype=np.uint16)
gt_mask = np.squeeze(gt_mask)
dice_score = getDiceScore(gt_mask, pred_mask)
if dice_score == 0:
dice_scores.append(dice_score)
hd_scores.append(200)
hd95_scores.append(200)
names.append(fname)
save_path = destiny_directory(model_name, int_dice_score,
post_processing = post_processing)
pred_mask = prepareForSaving(pred_mask)
save(pred_mask, os.path.join(save_path, fname + '_' + model_name + '_'
+ str(int_dice_score) + '.nii'), hdr)
continue
names.append(fname)
dice_scores.append(dice_score)
hd_score = hd(gt_mask, pred_mask)
hd_scores.append(hd_score)
hd95_score = hd95(gt_mask, pred_mask)
hd95_scores.append(hd95_score)
int_dice_score = floor(dice_score * 100)
save_path = destiny_directory(model_name, int_dice_score,
post_processing = post_processing)
pred_mask = prepareForSaving(pred_mask)
'''image = prepareForSaving(image)
gt_mask = prepareForSaving(gt_mask)'''
save(pred_mask, os.path.join(save_path, fname + '_' + model_name + '_'
+ str(int_dice_score) + '.nii'), hdr)
'''save(image, os.path.join(save_path, fname + '_img.nii'), hdr)'''
'''save(gt_mask, os.path.join(save_path, fname + '_mask.nii'), hdr)'''
return dice_scores, hd_scores, hd95_scores, names
model_types = ['unet', 'vgg19_fcn_upconv', 'vgg19FCN', 'unet_se']
for post_processing in [False, True]:
for model_type in model_types:
print()
if post_processing:
print('pp')
else:
print('no pp')
dh = DataHandler()
all_dice = []
all_hd = []
all_hd95 = []
all_names = []
for i in range(len(kfold_indices)):
exp_name = 'kfold_%s_dice_DA_K%d'%(model_type, i)
#get parameters
params = getParams(exp_name, unet_type=model_type)
val_img_files = np.take(image_files, kfold_indices[i]['val'])
val_mask_files = np.take(mask_files, kfold_indices[i]['val'])
model = getModel(model_type)
print('loading weights from %s'%params['checkpoint']['name'])
model.load_weights(params['checkpoint']['name'])
data = zip(val_img_files, val_mask_files)
dice_score, hd_score, hd95_score, names = predictAll(model, model_type, data,
num_data=len(val_mask_files),
post_processing = post_processing)
print('Finished K%d'%i)
all_dice += dice_score
all_hd += hd_score
all_hd95 += hd95_score
all_names.extend(names)
if post_processing:
report_name = 'data/eval_pp/' + model_type + '/' + model_type + '_report.txt'
else:
report_name = 'data/eval/' + model_type + '/' + model_type + '_report.txt'
with open(report_name, 'w+') as f:
for i in range(len(all_dice)):
f.write("%s, %f, %f, %f\n"%(all_names[i],
all_dice[i],
all_hd[i],
all_hd95[i]))
f.write('\n')
f.write('Final results for %s\n'%model_type)
f.write('dice %f\n'%np.mean(all_dice))
f.write('hd %f\n'%np.mean(all_hd))
f.write('hd95 %f\n'%np.mean(all_hd95))
```
| github_jupyter |
# NYC Capital Projects
## Notebook 06: Merge Engineered Features Into Final NYC Capital Projects Datasets
This notebook merges the cleansed NYC capital projects 3-year interval training and test datasets with the engineered features generated in the prior several notebooks. Those engineered features include the $k$-means-generated reference class labels; the PCA-, autoencoder-, and UMAP-generated 2-dimensional encoded BERT embeddings; and the UMAP+HDBSCAN reference class labels. The resulting merged training and test datasets will be used for predictive modeling in all subsequent notebooks for this project.
### Project authors
- [An Hoang](https://github.com/hoangthienan95)
- [Mark McDonald](https://github.com/mcdomx)
- [Mike Sedelmeyer](https://github.com/sedelmeyer)
### Inputs:
The following files are required to successfully run this notebook.
- ``../data/interim/NYC_capital_projects_3yr_train.csv``
The training split of our 3-year interval subsetted project data.
- ``../data/interim/NYC_capital_projects_3yr_test.csv``
The test split of our 3-year interval subsetted project data.
- ``../data/interim/kmeans3_attribute_labels_train.csv``
The $k$-means labels corresponding to each project record in our 3-year training split.
- ``../data/interim/kmeans3_attribute_labels_test.csv``
The $k$-means labels corresponding to each project record in our 3-year test split.
- ``../data/interim/ae_pca_encoded_embed_train.csv``
The PCA and autoencoder 2-dimensional feature values corresponding to each project record in our 3-year training split.
- ``../data/interim/ae_pca_encoded_embed_test.csv``
The PCA and autoencoder 2-dimensional feature values corresponding to each project record in our 3-year test split.
- ``../data/interim/UMAP_embeddings_NYC_capital_projects_3yr_train.csv``
The UMAP 2-dimensional encoded embeddings and the UMAP+HDBSCAN reference class cluster labels corresponding to each project record in our 3-year training split.
- ``../data/interim/UMAP_embeddings_NYC_capital_projects_3yr_test.csv``
The UMAP 2-dimensional encoded embeddings and the UMAP+HDBSCAN reference class cluster labels corresponding to each project record in our 3-year test split.
### Outputs:
The following files are generated by executing the code in this notebook.
- ``../data/processed/NYC_capital_projects_3yr_final_train.csv``
The final training dataset, including all engineered features, for use in subsequent prediction models.
- ``../data/processed/NYC_capital_projects_3yr_final_test.csv``
The final test dataset, including all engineered features, for use in subsequent prediction models.
# Notebook contents
1. [Imports](#Imports)
2. [Read and merge datasets](#Read-and-merge-datasets)
3. [Save merged datasets](#Save-merged-datasets)
# Imports
[Return to top](#Notebook-contents)
```
from functools import reduce
import glob
import os
from pickle import dump, load
import numpy as np
import pandas as pd
from tqdm.auto import tqdm
```
# Read and merge datasets
Load all required datasets and merge all engineered features with the NYC Capital Projects train and test datasets
[Return to top](#Notebook-contents)
```
files_needed = set(
[
'NYC_capital_projects_3yr_test',
'NYC_capital_projects_3yr_train',
'ae_pca_encoded_embed_test',
'ae_pca_encoded_embed_train',
'UMAP_embeddings_NYC_capital_projects_3yr_test',
'UMAP_embeddings_NYC_capital_projects_3yr_train',
'kmeans3_attribute_labels_test',
'kmeans3_attribute_labels_train',
]
)
files_needed_paths = [f"../data/interim/{file}.csv" for file in files_needed]
savepath_train = "../data/processed/NYC_capital_projects_3yr_final_train.csv"
savepath_test = "../data/processed/NYC_capital_projects_3yr_final_test.csv"
# check to ensure target files exist to prevent runtime errors
path_errors = []
for filepath in files_needed_paths:
if (not os.path.isfile(filepath)) and (not os.path.isdir(filepath)):
path_errors.append(filepath)
if len(path_errors)==0:
print("OK - all 'files_needed_paths' point to existing files!")
else:
raise ValueError(
"The following target paths do not exist...\n\n\t{}\n"\
"".format(path_errors)
)
data_dict = {"train":{}, "test":{}}
print('Dataframes added to data dictionary:\n')
for file in sorted(files_needed_paths):
file_name, extension = file.split("/")[-1].split(".")
if file_name.startswith("NYC"):
date_cols = [
'Design_Start',
'Final_Change_Date',
'Schedule_Start',
'Schedule_End',
]
drop_col = "Unnamed: 0"
#umap
else:
date_cols = []
drop_col = []
df = pd.read_csv(file, parse_dates=date_cols).drop(columns=drop_col)
if file_name.split("_")[-1] == "train":
data_dict["train"][file_name] = df
elif file_name.split("_")[-1] == "test":
data_dict["test"][file_name] = df
else:
data_dict[file_name] = df
print(f'\t{file_name}')
data_dict["train"].keys()
data_dict["test"].keys()
data_dict["train"]['UMAP_embeddings_NYC_capital_projects_3yr_train'].columns[
data_dict["train"]['UMAP_embeddings_NYC_capital_projects_3yr_train'].columns.str.contains("label")
]
# filters so only keep the 2D features
umap_df_train = data_dict["train"]['UMAP_embeddings_NYC_capital_projects_3yr_train']
umap_df_train['attribute_clustering_label'] = umap_df_train['attribute_clustering_label'].astype("str")
data_dict["train"]['UMAP_embeddings_NYC_capital_projects_3yr_train'] = umap_df_train[
["PID"] + list(
umap_df_train.columns[
umap_df_train.columns.str.startswith("umap_attributes_2D") |
umap_df_train.columns.str.startswith("umap_descr_2D")
]
) + ['attribute_clustering_label']
]
# filters so only keep the 2D features
umap_df_test = data_dict["test"]['UMAP_embeddings_NYC_capital_projects_3yr_test']
umap_df_test['attribute_clustering_label'] = umap_df_test['attribute_clustering_label'].astype("str")
data_dict["test"]['UMAP_embeddings_NYC_capital_projects_3yr_test'] = umap_df_test[
["PID"] + list(
umap_df_test.columns[
umap_df_test.columns.str.startswith("umap_attributes_2D") |
umap_df_test.columns.str.startswith("umap_descr_2D")
]
) + ['attribute_clustering_label']
]
df_train_merged = reduce(lambda left,right: pd.merge(left.copy(),right.copy(),on='PID',
how='left'), data_dict["train"].values())
assert df_train_merged.isnull().sum().sum() == 0
assert df_train_merged.shape == (134,53)
df_test_merged = reduce(lambda left,right: pd.merge(left.copy(),right.copy(),on='PID',
how='left'), data_dict["test"].values())
assert df_test_merged.isnull().sum().sum() == 0
assert df_test_merged.shape == (15,53)
df_train_merged.info()
df_test_merged.info()
```
# Save merged datasets
[Return to top](#Notebook-contents)
```
df_train_merged.to_csv(savepath_train)
df_test_merged.to_csv(savepath_test)
```
| github_jupyter |
```
import numpy
from keras.callbacks import TensorBoard
from keras.models import Sequential, load_model
from keras.preprocessing.image import ImageDataGenerator
from keras.layers import Dense
from keras.layers import Dropout
from keras.layers import Flatten
from keras.constraints import maxnorm
from keras.optimizers import SGD
from keras.layers.convolutional import Conv2D
from keras.layers.convolutional import MaxPooling2D
from keras.utils import np_utils
import pandas as pd
import matplotlib.pyplot as plt
%matplotlib inline
# load dataset
# only will be taking first 10000 images to make training faster
df_train = pd.read_csv('train.csv')[:10000]
# normalize pixels (only for X)
X = df_train.ix[:,1:].astype('float32') / 255.0
y = df_train.ix[:,0]
X.head()
print("Looking at digit", y.ix[0,0])
plt.imshow(X.ix[0,:].values.reshape(28,28),cmap='gray')
plt.axis('off')
plt.colorbar()
# pd.get_dummies(y_train) gives us a dataframe
# this one by keras will give in array form so might as well use this
y = np_utils.to_categorical(y)
num_classes = y.shape[1]
y
# keras doesnt deal with dataframes, only matrixes
# reshape to 4d; (samples, rows, cols, channels) if data_format='channels_last'.
X = X.as_matrix().reshape(X.shape[0], 28, 28, 1)
X.shape
# simple CNN
simple = Sequential()
simple.add(Conv2D(32, (3,3), padding='same', input_shape=(28,28,1), activation='relu', kernel_constraint=maxnorm(3)))
simple.add(Dropout(0.2))
simple.add(Conv2D(32, (3,3), padding='same', activation='relu', kernel_constraint=maxnorm(3)))
simple.add(MaxPooling2D())
simple.add(Flatten())
simple.add(Dense(512, activation='relu', kernel_constraint=maxnorm(3)))
simple.add(Dropout(0.5))
simple.add(Dense(num_classes, activation='softmax'))
# compile the model
simple.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
model.fit(X, y, validation_split=0.2, callbacks=[TensorBoard('./logs/run1')])
# a CNN
model = Sequential()
model.add(Conv2D(32, (3,3), padding='same', input_shape=(28,28,1), activation='relu', kernel_constraint=maxnorm(3)))
model.add(Dropout(0.2))
model.add(Conv2D(32, (3,3), padding='same', activation='relu', kernel_constraint=maxnorm(3)))
model.add(MaxPooling2D())
model.add(Conv2D(32, (3,3), padding='same', input_shape=(28,28,1), activation='relu', kernel_constraint=maxnorm(3)))
model.add(Dropout(0.2))
model.add(Conv2D(32, (3,3), padding='same', activation='relu', kernel_constraint=maxnorm(3)))
model.add(MaxPooling2D())
model.add(Conv2D(32, (3,3), padding='same', input_shape=(28,28,1), activation='relu', kernel_constraint=maxnorm(3)))
model.add(Dropout(0.2))
model.add(Conv2D(32, (3,3), padding='same', activation='relu', kernel_constraint=maxnorm(3)))
model.add(MaxPooling2D())
model.add(Flatten())
model.add(Dense(512, activation='relu', kernel_constraint=maxnorm(3)))
model.add(Dropout(0.5))
model.add(Dense(256, activation='relu', kernel_constraint=maxnorm(3)))
model.add(Dropout(0.5))
model.add(Dense(num_classes, activation='softmax'))
# compile the model (rmsprop)
model.compile(loss='categorical_crossentropy', optimizer='rmsprop', metrics=['accuracy'])
history = model.fit(X, y, validation_split=0.2, callbacks=[TensorBoard('./logs/run2')])
# compile the model (adam)
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
model.fit(X, y, validation_split=0.2, callbacks=[TensorBoard('./logs/run2')])
model.save('my_model.h5')
history_dict = history.history
history_dict
```
### simple visual for train/val loss
### more stuff on tensorboard
```
loss_values = history_dict['loss']
val_loss_values = history_dict['val_loss']
epochs = range(1, len(loss_values) + 1)
# "bo" is for "blue dot"
plt.plot(epochs, loss_values, 'bo')
# b+ is for "blue crosses"
plt.plot(epochs, val_loss_values, 'b+')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.show()
acc_values = history_dict['acc']
val_acc_values = history_dict['val_acc']
plt.plot(epochs, acc_values, 'bo')
plt.plot(epochs, val_acc_values, 'b+')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.show()
```
| github_jupyter |
## Initialize the ckan environment and requests session
```
from os import path, environ
import requests
from dataflows import Flow, load
from datapackage_pipelines_ckanext.helpers import get_plugin_configuration
config = get_plugin_configuration('odata_org_il')
data_path = config['data_path']
CKAN_API_KEY = environ.get('CKAN_API_KEY')
CKAN_URL = environ.get('CKAN_URL')
assert CKAN_API_KEY and CKAN_URL
CKAN_AUTH_HEADERS = {'Authorization': CKAN_API_KEY}
session = requests.session()
session.headers.update(CKAN_AUTH_HEADERS)
```
## Source entity data from foi site
```
from dataflows import Flow, load
import yaml
def process(rows):
for row in rows:
if int(row['nid']) == 446:
print(yaml.dump(row, default_flow_style=False, allow_unicode=True))
yield row
source_entity = Flow(
load(data_path+'/new_foi_offices/datapackage.json'),
process
).results()[0][0]
```
## Matching ckan group from foi_groups_matching excel resource
```
from dataflows import Flow, load
import yaml
def process(rows):
for i, row in enumerate(rows):
if row['entity_id'] == f'foi-office-{source_entity["nid"]}':
yield row
foi_group_matching_resource = Flow(load(data_path+'/foi_groups_matching/datapackage.json')).results()[0][0]
foi_group_matching_source_entity = [row for row in foi_group_matching_resource if row['entity_id'] == f'foi-office-{source_entity["nid"]}'][0]
print(yaml.dump(foi_group_matching_source_entity, default_flow_style=False, allow_unicode=True))
```
## Load existing entities and find matching group
```
from dataflows import Flow, load
from load_existing_entities import get_existing_entities_resource, get_existing_entities_resource_descriptor
from collections import defaultdict
stats = defaultdict(int)
existing_entities_resource = Flow(load(({'resources': [get_existing_entities_resource_descriptor()]},
[get_existing_entities_resource(stats)]))
).results()[0][0]
existing_entity = [row for row in existing_entities_resource if row['group_id'] == foi_group_matching_source_entity['Column3']][0]
print(yaml.dump(existing_entity, default_flow_style=False, allow_unicode=True))
print(f'num existing entities = {len(existing_entities)}')
print(dict(stats))
```
## Run dry run to update_foi_offices_entities manually only for this group
```
from update_foi_offices_entities import get_foi_offices_resource, get_existing_entities, get_foi_groups_matching
from collections import defaultdict
import yaml
stats = defaultdict(int)
existing_entities = {}
for row in get_existing_entities(existing_entities_resource, existing_entities, stats):
pass
for row in get_foi_groups_matching(foi_group_matching_resource, existing_entities, stats):
pass
for row in get_foi_offices_resource([source_entity], existing_entities, stats, True):
print(yaml.dump(row, default_flow_style=False, allow_unicode=True))
```
## Before updating - save the group datasets, otherwise they will be disconnected from group
```
raise NotImplementedError()
```
## Do the update
```
for row in get_foi_offices_resource([source_entity], existing_entities, stats, False):
print(yaml.dump(row, default_flow_style=False, allow_unicode=True))
```
## Restore the datasets
```
from os import path, environ
import requests
from dataflows import Flow, load
from datapackage_pipelines_ckanext.helpers import get_plugin_configuration
def restore_group_datasets(row):
group_id = row['group_id']
if group_id == existing_entity['group_id']:
for dataset_id in row['dataset_ids']:
res = session.post('{}/api/3/action/member_create'.format(CKAN_URL),
json=dict(id=group_id,
object=dataset_id,
object_type='package',
capacity='')).json()
assert res and res['success']
Flow(
load(path.join(data_path, 'dump_group_datasets/datapackage.json'), resources=['group_datasets']),
restore_group_datasets
).process()
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn import preprocessing
from sklearn import metrics
import os, sys
from time import time
from phm08ds.models import experiment
```
## Load Dataset
```
folderpath = '../../../data/interim/'
data_op_03 = pd.read_csv(folderpath + 'data_op_03.csv')
data_op_03.head()
```
## Data preprocessing
### Get rid of informations there are not sensor readings
Wang (2008) reports Sensor 15 has importat information. However, there are no relevant informations of this sensor. The data seems to be corrupted like this:
Let's remove it from our database creating an object transformer.
```
from phm08ds.features.feature_selection import RemoveSensor
tf_remove_sensor_15 = RemoveSensor(sensors=[15])
data_op_03 = tf_remove_sensor_15.fit_transform(data_op_03)
data_op_03.head()
```
Before feeding to the classifier, let's remove unwanted information, such as unit, time_step and operational settings.
```
from phm08ds.features.feature_selection import RemoveInfo
tf_remove_info = RemoveInfo()
data_with_features = tf_remove_info.fit_transform(data_op_03)
data_with_features.head()
```
We need to normalize our data. Let's use Z-score standardization.
```
from sklearn.preprocessing import StandardScaler
tf_std_scaller = preprocessing.StandardScaler()
data_with_features_std = tf_std_scaller.fit_transform(data_with_features.drop(labels='Health_state', axis=1))
data_with_features_std
labels = np.array(data_with_features['Health_state'])
labels
```
# Classification steps
## Load Experiment model
```
from phm08ds.models import experiment
```
## Define classifiers and its specifications
```
from sklearn.neural_network import MLPClassifier
from sklearn.svm import SVC
# SVM
svm_linear_clf = SVC(kernel='linear')
svm_rbf_clf = SVC(kernel='rbf')
svm_poly_clf = SVC(kernel='poly')
svm_sigmoid_clf = SVC(kernel='sigmoid')
```
## Put all clf in a dictionary:
```
classifiers = {'SVM-Linear': svm_linear_clf, 'SVM-RBF': svm_rbf_clf, 'SVM-Poly': svm_poly_clf, 'SVM-Sigmoid': svm_sigmoid_clf}
```
Since we are using SVM and MLP we need to extract all power from those methods. Let's perform a Random Search to parameters optimizations.
### Hyperparameter tunning
```
from sklearn.pipeline import Pipeline
data_preprocessing = Pipeline([('remove_sensor_15', tf_remove_sensor_15),
('remove_info', tf_remove_info),
('std_scaler', tf_std_scaller)
])
from sklearn.model_selection import RandomizedSearchCV
random_search = dict((k,[]) for k in classifiers.keys())
param_dist_dict = {
'SVM-Linear': {'C': [2**i for i in range(-5,15)]},
'SVM-RBF': {'gamma': [2**i for i in range(-15,3)], 'C': [2**i for i in range(-5,15)]},
'SVM-Poly': {'gamma': [2**i for i in range(-15,3)], 'C': [2**i for i in range(-5,15)]},
'SVM-Sigmoid': {'gamma': [2**i for i in range(-15,3)], 'C': [2**i for i in range(-5,15)]}
}
for clf in param_dist_dict.keys():
start = time()
random_search[clf] = RandomizedSearchCV(classifiers[clf], param_dist_dict[clf], cv=10, n_iter=5, verbose=5, n_jobs=1, scoring='accuracy')
random_search[clf].fit(data_with_features_std, labels)
experiment.save_models(random_search, name='clf_svm')
experiment.save_pipeline(data_preprocessing)
print('Elapsed time:')
print(time() - start)
```
## Savel results, models and pipeline to a .pkl file
```
from sklearn.pipeline import Pipeline
data_preprocessing = Pipeline([('remove_sensor_15', tf_remove_sensor_15),
('remove_info', tf_remove_info),
('std_scaler', tf_std_scaller)
])
experiment.save_models(random_search, name='clf_svm')
experiment.save_pipeline(data_preprocessing)
```
| github_jupyter |
```
import pandas as pd
import holoviews as hv
import geoviews as gv
import geoviews.feature as gf
import cartopy
import cartopy.feature as cf
from geoviews import opts
from cartopy import crs as ccrs
gv.extension('matplotlib', 'bokeh')
gv.output(dpi=120, fig='svg')
```
Cartopy and shapely make working with geometries and shapes very simple, and GeoViews provides convenient wrappers for the various geometry types they provide. In addition to Path and Polygons types, which draw geometries from lists of arrays or a geopandas DataFrame, GeoViews also provides the ``Feature`` and ``Shape`` types, which wrap cartopy Features and shapely geometries respectively.
### Feature
The Feature Element provides a very convenient means of overlaying a set of basic geographic features on top of or behind a plot. The ``cartopy.feature`` module provides various ways of loading custom features, however geoviews provides a number of default features which we have imported as ``gf``, amongst others this includes coastlines, country borders, and land masses. Here we demonstrate how we can plot these very easily, either in isolation or overlaid:
```
(gf.ocean + gf.land + gf.ocean * gf.land * gf.coastline * gf.borders).cols(3)
```
These deafult features simply wrap around cartopy Features, therefore we can easily load a custom ``NaturalEarthFeature`` such as graticules at 30 degree intervals:
```
graticules = cf.NaturalEarthFeature(
category='physical',
name='graticules_30',
scale='110m')
(gf.ocean() * gf.land() * gv.Feature(graticules, group='Lines') * gf.borders * gf.coastline).opts(
opts.Feature('Lines', projection=ccrs.Robinson(), facecolor='none', edgecolor='gray'))
```
The scale of features may be controlled using the ``scale`` plot option, the most common options being `'10m'`, `'50m'` and `'110m'`. Cartopy will downloaded the requested resolution as needed.
```
gv.output(backend='bokeh')
(gf.ocean * gf.land.options(scale='110m', global_extent=True) * gv.Feature(graticules, group='Lines') +
gf.ocean * gf.land.options(scale='50m', global_extent=True) * gv.Feature(graticules, group='Lines'))
```
Zoom in using the bokeh zoom widget and you should see that the right hand panel is using a higher resolution dataset for the land feature.
Instead of displaying a ``Feature`` directly it is also possible to request the geometries inside a ``Feature`` using the ``Feature.geoms`` method, which also allows specifying a ``scale`` and a ``bounds`` to select a subregion:
```
gf.land.geoms('50m', bounds=(-10, 40, 10, 60))
```
When working interactively with higher resolution datasets it is sometimes necessary to dynamically update the geometries based on the current viewport. The ``resample_geometry`` operation is an efficient way to display only polygons that intersect with the current viewport and downsample polygons on-the-fly.
```
gv.operation.resample_geometry(gf.coastline.geoms('10m')).opts(width=400, height=400, color='black')
```
Try zooming into the plot above and you will see the coastline geometry resolve to a higher resolution dynamically (this requires a live Python kernel).
### Shape
The ``gv.Shape`` object wraps around any shapely geometry, allowing finer grained control over each polygon. We can, for example, access the geometries on the ``LAND`` feature and display them individually. Here we will get the geometry corresponding to the Australian continent and display it using shapely's inbuilt SVG repr (not yet a HoloViews plot, just a bare SVG displayed by Jupyter directly):
```
land_geoms = gf.land.geoms(as_element=False)
land_geoms[21]
```
Instead of letting shapely render it as an SVG, we can now wrap it in the ``gv.Shape`` object and let matplotlib or bokeh render it, alone or with other GeoViews or HoloViews objects:
```
australia = gv.Shape(land_geoms[21])
alice_springs = gv.Text(133.870,-21.5, 'Alice Springs')
australia * gv.Points([(133.870,-23.700)]).opts(color='black', width=400) * alice_springs
```
We can also supply a list of geometries directly to a Polygons or Path element:
```
gv.Polygons(land_geoms) + gv.Path(land_geoms)
```
This makes it possible to create choropleth maps, where each part of the geometry is assigned a value that will be used to color it. However, constructing a choropleth by combining a bunch of shapes can be a lot of effort and is error prone. For that reason, the Shape Element provides convenience methods to load geometries from a shapefile. Here we load the boundaries of UK electoral districts directly from an existing shapefile:
```
hv.output(backend='matplotlib')
shapefile = '../assets/boundaries/boundaries.shp'
gv.Shape.from_shapefile(shapefile, crs=ccrs.PlateCarree())
```
To combine these shapes with some actual data, we have to be able to merge them with a dataset. To do so we can inspect the records the cartopy shapereader loads:
```
shapes = cartopy.io.shapereader.Reader(shapefile)
list(shapes.records())[0]
```
As we can see, the record contains a ``MultiPolygon`` together with a standard geographic ``code``, which we can use to match up the geometries with a dataset. To continue we will require a dataset that is also indexed by these codes. For this purpose we load a dataset of the 2016 EU Referendum result in the UK:
```
referendum = pd.read_csv('../assets/referendum.csv')
referendum = hv.Dataset(referendum)
referendum.data.head()
```
The ``from_records`` function optionally also supports merging the records and dataset directly. To merge them, supply the name of the shared attribute on which the merge is based via the ``on`` argument. If the name of attribute in the records and the dimension in the dataset match exactly, you can simply supply it as a string, otherwise supply a dictionary mapping between the attribute and column name. In this case we want to color the choropleth by the `'leaveVoteshare'`, which we define via the `value` argument.
Additionally we can request one or more indexes using the ``index`` argument. Finally we will declare the coordinate reference system in which this data is stored, which will in most cases be the simple Plate Carree projection. We can then view the choropleth, with each shape colored by the specified value (the percentage who voted to leave the EU):
```
hv.output(backend='bokeh')
gv.Shape.from_records(shapes.records(), referendum, on='code', value='leaveVoteshare',
index=['name', 'regionName']).opts(tools=['hover'], width=350, height=500)
```
## GeoPandas
GeoPandas extends the datatypes used by pandas to allow spatial operations on geometric types, which makes it a very convenient way of working with geometries with associated variables. GeoViews ``Path``, ``Contours`` and ``Polygons`` Elements natively support projecting and plotting of
geopandas DataFrames using both ``matplotlib`` and ``bokeh`` plotting extensions. We will load the example dataset of the world which also includes some additional data about each country:
```
import geopandas as gpd
world = gpd.read_file(gpd.datasets.get_path('naturalearth_lowres'))
world.head()
```
We can simply pass the GeoPandas DataFrame to a Polygons, Path or Contours element and it will plot the data for us. The ``Contours`` and ``Polygons`` will automatically color the data by the first specified value dimension defined by the ``vdims`` keyword (the geometries may be colored by any dimension using the ``color_index`` plot option):
```
gv.Polygons(world, vdims='pop_est').opts(projection=ccrs.Robinson(), width=600, tools=['hover'])
```
Geometries can be displayed using both matplotlib and bokeh, here we will switch to bokeh allowing us to color by a categorical variable (``continent``) and activating the hover tool to reveal information about the plot.
```
gv.Polygons(world, vdims=['continent', 'name', 'pop_est']).opts(
cmap='tab20', width=600, height=400, tools=['hover'], infer_projection=True)
```
The "Working with Bokeh" GeoViews notebook shows how to enable hover data that displays information about each of these shapes interactively.
| github_jupyter |
Copyright (c) Microsoft Corporation. All rights reserved.
Licensed under the MIT License.

# Azure Machine Learning Pipelines with Data Dependency
In this notebook, we will see how we can build a pipeline with implicit data dependency.
## Prerequisites and Azure Machine Learning Basics
If you are using an Azure Machine Learning Notebook VM, you are all set. Otherwise, make sure you go through the [configuration Notebook](https://aka.ms/pl-config) first if you haven't. This sets you up with a working config file that has information on your workspace, subscription id, etc.
### Azure Machine Learning and Pipeline SDK-specific Imports
```
import azureml.core
from azureml.core import Workspace, Experiment, Datastore
from azureml.core.compute import AmlCompute
from azureml.core.compute import ComputeTarget
from azureml.widgets import RunDetails
# Check core SDK version number
print("SDK version:", azureml.core.VERSION)
from azureml.data.data_reference import DataReference
from azureml.pipeline.core import Pipeline, PipelineData
from azureml.pipeline.steps import PythonScriptStep
print("Pipeline SDK-specific imports completed")
```
### Initialize Workspace
Initialize a [workspace](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.workspace(class%29) object from persisted configuration.
```
ws = Workspace.from_config()
print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\n')
# Default datastore (Azure blob storage)
# def_blob_store = ws.get_default_datastore()
def_blob_store = Datastore(ws, "workspaceblobstore")
print("Blobstore's name: {}".format(def_blob_store.name))
```
### Source Directory
The best practice is to use separate folders for scripts and its dependent files for each step and specify that folder as the `source_directory` for the step. This helps reduce the size of the snapshot created for the step (only the specific folder is snapshotted). Since changes in any files in the `source_directory` would trigger a re-upload of the snapshot, this helps keep the reuse of the step when there are no changes in the `source_directory` of the step.
```
# source directory
source_directory = 'data_dependency_run_train'
print('Sample scripts will be created in {} directory.'.format(source_directory))
```
### Required data and script files for the the tutorial
Sample files required to finish this tutorial are already copied to the project folder specified above. Even though the .py provided in the samples don't have much "ML work," as a data scientist, you will work on this extensively as part of your work. To complete this tutorial, the contents of these files are not very important. The one-line files are for demostration purpose only.
### Compute Targets
See the list of Compute Targets on the workspace.
```
cts = ws.compute_targets
for ct in cts:
print(ct)
```
#### Retrieve or create an Aml compute
Azure Machine Learning Compute is a service for provisioning and managing clusters of Azure virtual machines for running machine learning workloads. Let's get the default Aml Compute in the current workspace. We will then run the training script on this compute target.
```
from azureml.core.compute_target import ComputeTargetException
aml_compute_target = "cpu-cluster"
try:
aml_compute = AmlCompute(ws, aml_compute_target)
print("found existing compute target.")
except ComputeTargetException:
print("creating new compute target")
provisioning_config = AmlCompute.provisioning_configuration(vm_size = "STANDARD_D2_V2",
min_nodes = 1,
max_nodes = 4)
aml_compute = ComputeTarget.create(ws, aml_compute_target, provisioning_config)
aml_compute.wait_for_completion(show_output=True, min_node_count=None, timeout_in_minutes=20)
print("Aml Compute attached")
# For a more detailed view of current Azure Machine Learning Compute status, use get_status()
# example: un-comment the following line.
# print(aml_compute.get_status().serialize())
```
**Wait for this call to finish before proceeding (you will see the asterisk turning to a number).**
Now that you have created the compute target, let's see what the workspace's compute_targets() function returns. You should now see one entry named 'amlcompute' of type AmlCompute.
## Building Pipeline Steps with Inputs and Outputs
As mentioned earlier, a step in the pipeline can take data as input. This data can be a data source that lives in one of the accessible data locations, or intermediate data produced by a previous step in the pipeline.
### Datasources
Datasource is represented by **[DataReference](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.data.data_reference.datareference?view=azure-ml-py)** object and points to data that lives in or is accessible from Datastore. DataReference could be a pointer to a file or a directory.
```
# Reference the data uploaded to blob storage using DataReference
# Assign the datasource to blob_input_data variable
# DataReference(datastore,
# data_reference_name=None,
# path_on_datastore=None,
# mode='mount',
# path_on_compute=None,
# overwrite=False)
blob_input_data = DataReference(
datastore=def_blob_store,
data_reference_name="test_data",
path_on_datastore="20newsgroups/20news.pkl")
print("DataReference object created")
```
### Intermediate/Output Data
Intermediate data (or output of a Step) is represented by **[PipelineData](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-core/azureml.pipeline.core.pipelinedata?view=azure-ml-py)** object. PipelineData can be produced by one step and consumed in another step by providing the PipelineData object as an output of one step and the input of one or more steps.
#### Constructing PipelineData
- **name:** [*Required*] Name of the data item within the pipeline graph
- **datastore_name:** Name of the Datastore to write this output to
- **output_name:** Name of the output
- **output_mode:** Specifies "upload" or "mount" modes for producing output (default: mount)
- **output_path_on_compute:** For "upload" mode, the path to which the module writes this output during execution
- **output_overwrite:** Flag to overwrite pre-existing data
```
# Define intermediate data using PipelineData
# Syntax
# PipelineData(name,
# datastore=None,
# output_name=None,
# output_mode='mount',
# output_path_on_compute=None,
# output_overwrite=None,
# data_type=None,
# is_directory=None)
# Naming the intermediate data as processed_data1 and assigning it to the variable processed_data1.
processed_data1 = PipelineData("processed_data1",datastore=def_blob_store)
print("PipelineData object created")
```
### Pipelines steps using datasources and intermediate data
Machine learning pipelines can have many steps and these steps could use or reuse datasources and intermediate data. Here's how we construct such a pipeline:
#### Define a Step that consumes a datasource and produces intermediate data.
In this step, we define a step that consumes a datasource and produces intermediate data.
**Open `train.py` in the local machine and examine the arguments, inputs, and outputs for the script. That will give you a good sense of why the script argument names used below are important.**
#### Specify conda dependencies and a base docker image through a RunConfiguration
This step uses a docker image and scikit-learn, use a [**RunConfiguration**](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.runconfiguration?view=azure-ml-py) to specify these requirements and use when creating the PythonScriptStep.
```
from azureml.core.runconfig import RunConfiguration
from azureml.core.conda_dependencies import CondaDependencies
from azureml.core.runconfig import DEFAULT_CPU_IMAGE
# create a new runconfig object
run_config = RunConfiguration()
# enable Docker
run_config.environment.docker.enabled = True
# set Docker base image to the default CPU-based image
run_config.environment.docker.base_image = DEFAULT_CPU_IMAGE
# use conda_dependencies.yml to create a conda environment in the Docker image for execution
run_config.environment.python.user_managed_dependencies = False
# specify CondaDependencies obj
run_config.environment.python.conda_dependencies = CondaDependencies.create(conda_packages=['scikit-learn'])
# step4 consumes the datasource (Datareference) in the previous step
# and produces processed_data1
trainStep = PythonScriptStep(
script_name="train.py",
arguments=["--input_data", blob_input_data, "--output_train", processed_data1],
inputs=[blob_input_data],
outputs=[processed_data1],
compute_target=aml_compute,
source_directory=source_directory,
runconfig=run_config
)
print("trainStep created")
```
#### Define a Step that consumes intermediate data and produces intermediate data
In this step, we define a step that consumes an intermediate data and produces intermediate data.
**Open `extract.py` in the local machine and examine the arguments, inputs, and outputs for the script. That will give you a good sense of why the script argument names used below are important.**
```
# step5 to use the intermediate data produced by step4
# This step also produces an output processed_data2
processed_data2 = PipelineData("processed_data2", datastore=def_blob_store)
source_directory = "data_dependency_run_extract"
extractStep = PythonScriptStep(
script_name="extract.py",
arguments=["--input_extract", processed_data1, "--output_extract", processed_data2],
inputs=[processed_data1],
outputs=[processed_data2],
compute_target=aml_compute,
source_directory=source_directory)
print("extractStep created")
```
#### Define a Step that consumes intermediate data and existing data and produces intermediate data
In this step, we define a step that consumes multiple data types and produces intermediate data.
This step uses the output generated from the previous step as well as existing data on a DataStore. The location of the existing data is specified using a [**PipelineParameter**](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-core/azureml.pipeline.core.pipelineparameter?view=azure-ml-py) and a [**DataPath**](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.data.datapath.datapath?view=azure-ml-py). Using a PipelineParameter enables easy modification of the data location when the Pipeline is published and resubmitted.
**Open `compare.py` in the local machine and examine the arguments, inputs, and outputs for the script. That will give you a good sense of why the script argument names used below are important.**
```
# Reference the data uploaded to blob storage using a PipelineParameter and a DataPath
from azureml.pipeline.core import PipelineParameter
from azureml.data.datapath import DataPath, DataPathComputeBinding
datapath = DataPath(datastore=def_blob_store, path_on_datastore='20newsgroups/20news.pkl')
datapath_param = PipelineParameter(name="compare_data", default_value=datapath)
data_parameter1 = (datapath_param, DataPathComputeBinding(mode='mount'))
# Now define the compare step which takes two inputs and produces an output
processed_data3 = PipelineData("processed_data3", datastore=def_blob_store)
source_directory = "data_dependency_run_compare"
compareStep = PythonScriptStep(
script_name="compare.py",
arguments=["--compare_data1", data_parameter1, "--compare_data2", processed_data2, "--output_compare", processed_data3],
inputs=[data_parameter1, processed_data2],
outputs=[processed_data3],
compute_target=aml_compute,
source_directory=source_directory)
print("compareStep created")
```
#### Build the pipeline
```
pipeline1 = Pipeline(workspace=ws, steps=[compareStep])
print ("Pipeline is built")
pipeline_run1 = Experiment(ws, 'Data_dependency_sample').submit(pipeline1)
print("Pipeline is submitted for execution")
RunDetails(pipeline_run1).show()
```
#### Wait for pipeline run to complete
```
pipeline_run1.wait_for_completion(show_output=True)
```
### See Outputs
See where outputs of each pipeline step are located on your datastore.
***Wait for pipeline run to complete, to make sure all the outputs are ready***
```
# Get Steps
for step in pipeline_run1.get_steps():
print("Outputs of step " + step.name)
# Get a dictionary of StepRunOutputs with the output name as the key
output_dict = step.get_outputs()
for name, output in output_dict.items():
output_reference = output.get_port_data_reference() # Get output port data reference
print("\tname: " + name)
print("\tdatastore: " + output_reference.datastore_name)
print("\tpath on datastore: " + output_reference.path_on_datastore)
```
### Download Outputs
We can download the output of any step to our local machine using the SDK.
```
# Retrieve the step runs by name 'train.py'
train_step = pipeline_run1.find_step_run('train.py')
if train_step:
train_step_obj = train_step[0] # since we have only one step by name 'train.py'
train_step_obj.get_output_data('processed_data1').download("./outputs") # download the output to current directory
```
# Next: Publishing the Pipeline and calling it from the REST endpoint
See this [notebook](https://aka.ms/pl-pub-rep) to understand how the pipeline is published and you can call the REST endpoint to run the pipeline.
| github_jupyter |
# Bungee Dunk Revisited
*Modeling and Simulation in Python*
Copyright 2021 Allen Downey
License: [Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International](https://creativecommons.org/licenses/by-nc-sa/4.0/)
```
# install Pint if necessary
try:
import pint
except ImportError:
!pip install pint
# download modsim.py if necessary
from os.path import exists
filename = 'modsim.py'
if not exists(filename):
from urllib.request import urlretrieve
url = 'https://raw.githubusercontent.com/AllenDowney/ModSim/main/'
local, _ = urlretrieve(url+filename, filename)
print('Downloaded ' + local)
# import functions from modsim
from modsim import *
```
In the previous case study, we simulated a bungee jump with a model that took into account gravity, air resistance, and the spring force of the bungee cord, but we ignored the weight of the cord.
It is tempting to say that the weight of the cord doesn't matter because it falls along with the jumper. But that intuition is incorrect, as explained by [Heck, Uylings, and Kędzierska](http://iopscience.iop.org/article/10.1088/0031-9120/45/1/007). As the cord falls, it transfers energy to the jumper. They derive a differential equation that relates the acceleration of the jumper to position and velocity:
$a = g + \frac{\mu v^2/2}{\mu(L+y) + 2L}$
where $a$ is the net acceleration of the jumper, $g$ is acceleration due to gravity, $v$ is the velocity of the jumper, $y$ is the position of the jumper relative to the starting point (usually negative), $L$ is the length of the cord, and $\mu$ is the mass ratio of the cord and jumper.
If you don't believe this model is correct, [this video might convince you](https://www.youtube.com/watch?v=X-QFAB0gEtE).
Following the previous case study, we'll model the jump with the following assumptions:
1. Initially the bungee cord hangs from a crane with the attachment point 80 m above a cup of tea.
2. Until the cord is fully extended, it applies a force to the jumper as explained above.
3. After the cord is fully extended, it obeys [Hooke's Law](https://en.wikipedia.org/wiki/Hooke%27s_law); that is, it applies a force to the jumper proportional to the extension of the cord beyond its resting length.
4. The jumper is subject to drag force proportional to the square of their velocity, in the opposite of their direction of motion.
First I'll create a `Param` object to contain the quantities we'll need:
1. Let's assume that the jumper's mass is 75 kg and the cord's mass is also 75 kg, so `mu=1`.
2. The jumpers's frontal area is 1 square meter, and terminal velocity is 60 m/s. I'll use these values to back out the coefficient of drag.
3. The length of the bungee cord is `L = 25 m`.
4. The spring constant of the cord is `k = 40 N / m` when the cord is stretched, and 0 when it's compressed.
I adopt the coordinate system and most of the variable names from [Heck, Uylings, and Kędzierska](http://iopscience.iop.org/article/10.1088/0031-9120/45/1/007).
```
params = Params(y_attach = 80, # m,
v_init = 0, # m / s,
g = 9.8, # m/s**2,
M = 75, # kg,
m_cord = 75, # kg
area = 1, # m**2,
rho = 1.2, # kg/m**3,
v_term = 60, # m / s,
L = 25, # m,
k = 40, # N / m
)
```
Now here's a version of `make_system` that takes a `Params` object as a parameter.
`make_system` uses the given value of `v_term` to compute the drag coefficient `C_d`.
It also computes `mu` and the initial `State` object.
```
def make_system(params):
"""Makes a System object for the given params.
params: Params object
returns: System object
"""
M, m_cord = params.M, params.m_cord
g, rho, area = params.g, params.rho, params.area
v_init, v_term = params.v_init, params.v_term
# back out the coefficient of drag
C_d = 2 * M * g / (rho * area * v_term**2)
mu = m_cord / M
init = State(y=params.y_attach, v=v_init)
t_end = 8
return System(params, C_d=C_d, mu=mu,
init=init, t_end=t_end)
```
Let's make a `System`
```
system1 = make_system(params)
```
`drag_force` computes drag as a function of velocity:
```
def drag_force(v, system):
"""Computes drag force in the opposite direction of `v`.
v: velocity
returns: drag force in N
"""
rho, C_d, area = system.rho, system.C_d, system.area
f_drag = -np.sign(v) * rho * v**2 * C_d * area / 2
return f_drag
```
Here's drag force at 20 m/s.
```
drag_force(20, system1)
```
The following function computes the acceleration of the jumper due to tension in the cord.
$a_{cord} = \frac{\mu v^2/2}{\mu(L+y) + 2L}$
```
def cord_acc(y, v, system):
"""Computes the force of the bungee cord on the jumper:
y: height of the jumper
v: velocity of the jumpter
returns: acceleration in m/s
"""
L, mu = system.L, system.mu
a_cord = -v**2 / 2 / (2*L/mu + (L+y))
return a_cord
```
Here's acceleration due to tension in the cord if we're going 20 m/s after falling 20 m.
```
y = -20
v = -20
cord_acc(y, v, system1)
```
Now here's the slope function:
```
def slope_func1(t, state, system):
"""Compute derivatives of the state.
state: position, velocity
t: time
system: System object containing g, rho,
C_d, area, and mass
returns: derivatives of y and v
"""
y, v = state
M, g = system.M, system.g
a_drag = drag_force(v, system) / M
a_cord = cord_acc(y, v, system)
dvdt = -g + a_cord + a_drag
return v, dvdt
```
As always, let's test the slope function with the initial params.
```
slope_func1(0, system1.init, system1)
```
We'll need an event function to stop the simulation when we get to the end of the cord.
```
def event_func1(t, state, system):
"""Run until y=-L.
state: position, velocity
t: time
system: System object containing g, rho,
C_d, area, and mass
returns: difference between y and y_attach-L
"""
y, v = state
return y - (system.y_attach - system.L)
```
We can test it with the initial conditions.
```
event_func1(0, system1.init, system1)
```
And then run the simulation.
```
results1, details1 = run_solve_ivp(system1, slope_func1,
events=event_func1)
details1.message
```
Here's the plot of position as a function of time.
```
def plot_position(results, **options):
results.y.plot(**options)
decorate(xlabel='Time (s)',
ylabel='Position (m)')
plot_position(results1)
```
We can use `min` to find the lowest point:
```
min(results1.y)
```
As expected, Phase 1 ends when the jumper reaches an altitude of 55 m.
## Phase 2
Once the jumper has falled more than the length of the cord, acceleration due to energy transfer from the cord stops abruptly. As the cord stretches, it starts to exert a spring force. So let's simulate this second phase.
`spring_force` computes the force of the cord on the jumper:
```
def spring_force(y, system):
"""Computes the force of the bungee cord on the jumper:
y: height of the jumper
Uses these variables from system:
y_attach: height of the attachment point
L: resting length of the cord
k: spring constant of the cord
returns: force in N
"""
L, k = system.L, system.k
distance_fallen = system.y_attach - y
extension = distance_fallen - L
f_spring = k * extension
return f_spring
```
The spring force is 0 until the cord is fully extended. When it is extended 1 m, the spring force is 40 N.
```
spring_force(55, system1)
spring_force(56, system1)
```
The slope function for Phase 2 includes the spring force, and drops the acceleration due to the cord.
```
def slope_func2(t, state, system):
"""Compute derivatives of the state.
state: position, velocity
t: time
system: System object containing g, rho,
C_d, area, and mass
returns: derivatives of y and v
"""
y, v = state
M, g = system.M, system.g
a_drag = drag_force(v, system) / M
a_spring = spring_force(y, system) / M
dvdt = -g + a_drag + a_spring
return v, dvdt
```
The initial state for Phase 2 is the final state from Phase 1.
```
t_final = results1.index[-1]
t_final
state_final = results1.iloc[-1]
state_final
```
And that gives me the starting conditions for Phase 2.
```
system2 = System(system1, t_0=t_final, init=state_final)
```
Here's how we run Phase 2, setting the direction of the event function so it doesn't stop the simulation immediately.
```
results2, details2 = run_solve_ivp(system2, slope_func2)
details2.message
t_final = results2.index[-1]
t_final
```
We can plot the results on the same axes.
```
plot_position(results1, label='Phase 1')
plot_position(results2, label='Phase 2')
```
And get the lowest position from Phase 2.
```
min(results2.y)
```
To see how big the effect of the cord is, I'll collect the previous code in a function.
```
def run_two_phases(params):
system1 = make_system(params)
results1, details1 = run_solve_ivp(system1, slope_func1,
events=event_func1)
t_final = results1.index[-1]
state_final = results1.iloc[-1]
system2 = system1.set(t_0=t_final, init=state_final)
results2, details2 = run_solve_ivp(system2, slope_func2)
return results1.append(results2)
```
Now we can run both phases and get the results in a single `TimeFrame`.
```
results = run_two_phases(params)
plot_position(results)
params_no_cord = params.set(m_cord=1)
results_no_cord = run_two_phases(params_no_cord);
plot_position(results, label='m_cord = 75 kg')
plot_position(results_no_cord, label='m_cord = 1 kg')
min(results_no_cord.y)
diff = min(results.y) - min(results_no_cord.y)
diff
```
The difference is about a meter, which could certainly be the difference between a successful bungee dunk and a bad day.
| github_jupyter |
# Heterogeneous Synapses
In this example, we demonstrate how to build an `Ensemble` that uses different synapses per dimension in the vector space, or a different synapse per neuron.
For the most general case, **``HeteroSynapse``** is a function for use within a `Node`. It accepts some vector as input, and outputs a filtered version of this vector. The dimensionality of the output vector depends on the number of elements in the list `synapses` and the boolean value `elementwise`:
- If `elementwise == False`, then each synapse is applied to every input dimension, resulting in an output vector that is `len(synapses)` times larger.
- If `elementwise == True`, then each synapse is applied separately to a single input dimension, resulting in an output vector that is size `len(synapses)`, which must also be the same as the input dimension.
### Neuron Example
We first sample 100 neurons and 100 synapses randomly.
```
import numpy as np
import nengo
import nengolib
from nengolib.stats import sphere
from nengolib.synapses import HeteroSynapse
n_neurons = 100
dt = 0.001
T = 0.1
dims_in = 2
taus = nengo.dists.Uniform(0.001, 0.1).sample(n_neurons)
synapses = [nengo.Lowpass(tau) for tau in taus]
encoders = sphere.sample(n_neurons, dims_in)
```
Now we create two identical ensembles, one to hold the expected result, and one to compare this with the actual result from using `HeteroSynapse`. The former is computed via brute-force, by creating a separate connection for each synapse. The latter requires a single connection to the special node.
When `elementwise = False`, each input dimension is effectively broadcast to all of the neurons with a different synapse per neuron. We also note that since we are connecting directly to the neurons, we must embed the encoders in the transformation.
```
hs = HeteroSynapse(synapses, dt)
def embed_encoders(x):
# Reshapes the vectors to be the same dimensionality as the
# encoders, and then takes the dot product row by row.
# See http://stackoverflow.com/questions/26168363/ for a more
# efficient solution.
return np.sum(encoders * hs.from_vector(x), axis=1)
with nengolib.Network() as model:
# Input stimulus
stim = nengo.Node(size_in=dims_in)
for i in range(dims_in):
nengo.Connection(
nengo.Node(output=nengo.processes.WhiteSignal(T, high=10)),
stim[i], synapse=None)
# HeteroSynapse node
syn = nengo.Node(size_in=dims_in, output=hs)
# For comparing results
x = [nengo.Ensemble(n_neurons, dims_in, seed=0, encoders=encoders)
for _ in range(2)] # expected, actual
# Expected
for i, synapse in enumerate(synapses):
t = np.zeros_like(encoders)
t[i, :] = encoders[i, :]
nengo.Connection(stim, x[0].neurons, transform=t, synapse=synapse)
# Actual
nengo.Connection(stim, syn, synapse=None)
nengo.Connection(syn, x[1].neurons, function=embed_encoders, synapse=None)
# Probes
p_exp = nengo.Probe(x[0].neurons, synapse=None)
p_act = nengo.Probe(x[1].neurons, synapse=None)
# Check correctness
sim = nengo.Simulator(model, dt=dt)
sim.run(T)
assert np.allclose(sim.data[p_act], sim.data[p_exp])
```
### Vector Example
This example applies 2 synapses to their respective dimensions in a 2D-vector. We first initialize our parameters to use 20 neurons and 2 randomly chosen synapses.
```
n_neurons = 20
dt = 0.0005
T = 0.1
dims_in = 2
synapses = [nengo.Alpha(0.1), nengo.Lowpass(0.005)]
assert dims_in == len(synapses)
encoders = sphere.sample(n_neurons, dims_in)
```
Similar to the last example, we create two ensembles, one to obtain the expected result for verification, and another to be computed using the `HeteroSynapse` node.
```
with nengolib.Network() as model:
# Input stimulus
stim = nengo.Node(size_in=dims_in)
for i in range(dims_in):
nengo.Connection(
nengo.Node(output=nengo.processes.WhiteSignal(T, high=10)),
stim[i], synapse=None)
# HeteroSynapse Nodes
syn_elemwise = nengo.Node(
size_in=dims_in,
output=HeteroSynapse(synapses, dt, elementwise=True))
# For comparing results
x = [nengo.Ensemble(n_neurons, dims_in, seed=0, encoders=encoders)
for _ in range(2)] # expected, actual
# Expected
for j, synapse in enumerate(synapses):
nengo.Connection(stim[j], x[0][j], synapse=synapse)
# Actual
nengo.Connection(stim, syn_elemwise, synapse=None)
nengo.Connection(syn_elemwise, x[1], synapse=None)
# Probes
p_exp = nengo.Probe(x[0], synapse=None)
p_act_elemwise = nengo.Probe(x[1], synapse=None)
# Check correctness
sim = nengo.Simulator(model, dt=dt)
sim.run(T)
assert np.allclose(sim.data[p_act_elemwise], sim.data[p_exp])
```
### Multiple Vector Example
As a final example, to demonstrate the generality of this approach, we consider the situation where we wish to apply a number of different synapses to every dimension. For instance, with a 2D input vector, we pick 3 synapses to apply to every dimension, such that our ensemble will represent a 6D-vector (one for each dimension/synapse pair).
```
n_neurons = 20
dt = 0.0005
T = 0.1
dims_in = 2
synapses = [nengo.Alpha(0.1), nengo.Lowpass(0.005), nengo.Alpha(0.02)]
dims_out = len(synapses)*dims_in
encoders = sphere.sample(n_neurons, dims_out)
```
We also demonstrate that this can be achieved in two different ways. The first is with `elementwise=False`, by a broadcasting similar to the first example. The second is with `elementwise=True`, by replicating each synapse to align with each dimension, and then proceeding similar to the second example.
```
with nengolib.Network() as model:
# Input stimulus
stim = nengo.Node(size_in=dims_in)
for i in range(dims_in):
nengo.Connection(
nengo.Node(output=nengo.processes.WhiteSignal(T, high=10)),
stim[i], synapse=None)
# HeteroSynapse Nodes
syn_dot = nengo.Node(
size_in=dims_in, output=HeteroSynapse(synapses, dt))
syn_elemwise = nengo.Node(
size_in=dims_out,
output=HeteroSynapse(np.repeat(synapses, dims_in), dt, elementwise=True))
# For comparing results
x = [nengo.Ensemble(n_neurons, dims_out, seed=0, encoders=encoders)
for _ in range(3)] # expected, actual 1, actual 2
# Expected
for j, synapse in enumerate(synapses):
nengo.Connection(stim, x[0][j*dims_in:(j+1)*dims_in], synapse=synapse)
# Actual (method #1 = matrix multiplies)
nengo.Connection(stim, syn_dot, synapse=None)
nengo.Connection(syn_dot, x[1], synapse=None)
# Actual (method #2 = elementwise)
for j in range(len(synapses)):
nengo.Connection(stim, syn_elemwise[j*dims_in:(j+1)*dims_in], synapse=None)
nengo.Connection(syn_elemwise, x[2], synapse=None)
# Probes
p_exp = nengo.Probe(x[0], synapse=None)
p_act_dot = nengo.Probe(x[1], synapse=None)
p_act_elemwise = nengo.Probe(x[2], synapse=None)
# Check correctness
sim = nengo.Simulator(model, dt=dt)
sim.run(T)
assert np.allclose(sim.data[p_act_dot], sim.data[p_exp])
assert np.allclose(sim.data[p_act_elemwise], sim.data[p_exp])
```
| github_jupyter |
```
# default_exp models.transfermodels
```
# models.transfermodels
> API details.
```
# export
import os
os.environ["CUDA_VISIBLE_DEVICES"] = ""
import numpy as np
import torch
from torch import nn
from fastrenewables.tabular.model import *
from fastrenewables.timeseries.model import *
from fastai.tabular.all import *
from torch.autograd import Variable
from sklearn.datasets import make_regression
from fastai.learner import *
from fastrenewables.utils_pytorch import *
import copy
from fastrenewables.timeseries.model import *
from fastrenewables.baselines import BayesLinReg
from fastrenewables.tabular.learner import convert_to_tensor
from fastrenewables.losses import *
def generate_single_dataset(n_samples, start, end, bias, coef, noise_factor=0.3):
X = np.random.uniform(low=start, high=end,size=n_samples)
y = np.sin(X*coef*2*np.pi) + np.random.randn(X.shape[0])*noise_factor+bias
return X,y
def generate_all_tasks(n_samples=100):
starts = [0, 0]
ends = [4.1, 3.9]
coefs = [3.4, 4]
biases = [0.1, 0.1]
n_samples = [n_samples, 30]
df_tasks = []
for task_id in range(len(starts)):
start, end, bias, coef = starts[task_id], ends[task_id], coefs[task_id], biases[task_id]
X,y = generate_single_dataset(n_samples[task_id], start, end, bias, coef, noise_factor=0.05)
df_task = pd.DataFrame({"X": X.ravel(), "y":y.ravel()})
df_task["TaskID"] = task_id
df_tasks.append(df_task)
return pd.concat(df_tasks)
def get_source_task(df):
df_source = df[df.TaskID == 0]
dls = TabularDataLoaders.from_df(df_source, cont_names="X", y_names="y",
deivce="cpu", procs=Normalize, bs=10)
return dls
def get_target_task(df):
df_target = df[df.TaskID == 1]
dls = TabularDataLoaders.from_df(df_target, cont_names="X", y_names="y",
deivce="cpu", procs=Normalize, bs=10)
return dls
df = generate_all_tasks()
plt.scatter(df.X, df.y, c=df.TaskID)
dls_source = get_source_task(df)
dls_target = get_target_task(df)
set_seed(10)
source_model = MultiLayerPerceptron([1, 100, 10, 5, 1], use_bn=True, bn_cont=False)
learn_source = Learner(dls_source, source_model, metrics=rmse)
learn_source.fit(25, lr=0.01)
targets, preds = learn_source.get_preds(ds_idx=0)
plt.scatter(dls_source.train_ds.items.X, preds, label="preds")
plt.scatter(dls_source.train_ds.items.X, targets, label="targets")
plt.legend()
plt.show()
# hide
# export
def _create_matrices(n_features, alpha):
w_mean = torch.zeros(n_features)
w_precision = torch.eye(n_features) / alpha
return w_mean, w_precision
# minimal check if we can create the matricies
mean, precision = _create_matrices(10, 10)
test_eq(0, mean.sum())
test_eq(0.1, precision[0,0])
# export
class LinearTransferModel(nn.Module):
def __init__(self, source_model, num_layers_to_remove=1,
name_layers_or_function_to_remove="layers",
use_original_weights=True,
prediction_model=BayesLinReg(alpha=1, beta=1, empirical_bayes=False)):
super().__init__()
self.are_weights_initialized = False
self.num_layers_to_remove = num_layers_to_remove
self.ts_length = 1
self.source_model = copy.deepcopy(source_model)
self._prediction_model = prediction_model
self.prediction_models = []
if use_original_weights:
self._prediction_model.empirical_bayes=False
if callable(name_layers_or_function_to_remove):
name_layers_or_function_to_remove(self.source_model, num_layers_to_remove)
elif type(name_layers_or_function_to_remove) == str:
layers = getattrs(self.source_model, name_layers_or_function_to_remove, default=None)[0]
if layers is None:
raise ValueError(f"Could not find layers by given name {name_layers_or_function_to_remove}.")
elif isinstance(layers, torch.nn.modules.container.Sequential):
setattr(self.source_model, name_layers_or_function_to_remove, layers[0:-self.num_layers_to_remove])
else:
raise ValueError(f"Only sequential layers are supported.")
else:
ValueError("Unknown type for name_layers_or_function_to_remove")
if num_layers_to_remove != 1 and use_original_weights:
raise ValueError("Can only reuse weights when using the last layers due to the dimension.")
elif num_layers_to_remove == 1 and use_original_weights:
for element in layers[-1]:
if isinstance(element, nn.Linear):
# create mean matrix including bias
w_mean = copy.copy(element.weight.data)
bias = copy.copy(element.bias.data)
w_mean = w_mean.reshape(w_mean.shape[1])
w_mean = to_np(torch.cat([bias, w_mean]))
# create precision and variance matrix
self.n_features = w_mean.shape[0]
model = self._create_single_model(self.n_features)
model.w_mean = w_mean
self.prediction_models.append(model)
self.are_weights_initialized = True
if not self.are_weights_initialized:
raise ValueError(f"Could not find linear layer in last layer {self.layers[-1]}")
freeze(self.source_model)
# fake param so that it can be used with pytorch trainers
self.fake_param=nn.Parameter(torch.zeros((1,1), dtype=torch.float))
self.fake_param.requires_grad =True
def _create_single_model(self,n_features):
model = copy.copy(self._prediction_model)
model._create_matrices(np.ones(n_features).reshape(1, n_features))
model.w_covariance = np.linalg.inv(model.w_precision)
return model
@property
def alpha(self):
return self._prediction_model.alpha
@property
def beta(self):
return self._prediction_model.beta
@alpha.setter
def alpha(self, alpha):
self._prediction_model.alpha = alpha
@beta.setter
def beta(self, beta):
self._prediction_model.beta = beta
def correct_shape(self, x):
n_samples = x.shape[0]
return x.reshape(n_samples, -1)
def transform(self, cats, conts, as_np=False):
x_transformed = self.source_model(cats, conts)
x_transformed = self.correct_shape(x_transformed)
if as_np: return to_np(x_transformed)
else: return x_transformed
def forward(self, cats, conts):
n_samples = conts.shape[0]
self.ts_length = 1
if len(conts.shape) == 3:
self.ts_length = conts.shape[2]
x_transformed = self.transform(cats, conts)
if not self.are_weights_initialized:
self.n_features = x_transformed.shape[1]+1
for idx in range(self.ts_length):
model = self._create_single_model(self.n_features)
self.prediction_models.append(model)
self.are_weights_initialized=True
if self.training:
return x_transformed
else:
preds = self.pred_transformed_X(x_transformed)
return preds
def update(self, X, y):
X = to_np(X)
y = to_np(y)
y = self.correct_shape(y)
for idx, prediction_model in enumerate(self.prediction_models):
prediction_model.fit(X, y[:,idx].ravel())
return self
def predict(self, cats, conts):
x_transformed = self.transform(cats, conts, as_np=True)
return self.pred_transformed_X(x_transformed)
def predict_proba(self, cats, conts):
x_transformed = self.transform(cats, conts, as_np=True)
return self.pred_transformed_X(x_transformed, include_std=True)
def pred_transformed_X(self, x_transformed, include_std=False):
y_pred_means = np.zeros((len(x_transformed), len(self.prediction_models)))
y_pred_stds = np.zeros((len(x_transformed), len(self.prediction_models)))
for idx, prediction_model in enumerate(self.prediction_models):
y_pred_mean, y_pred_std = prediction_model.predict_proba(x_transformed)
y_pred_means[:,idx] = y_pred_mean
y_pred_stds[:,idx] = y_pred_std
if include_std:
return torch.tensor(y_pred_means, dtype=torch.float32), torch.tensor(y_pred_stds, dtype=torch.float32),
else:
return torch.tensor(y_pred_means, dtype=torch.float32)
def loss_func(self, x_transformed, ys):
ys = self.correct_shape(ys)
if self.training:
self.update(x_transformed, ys)
fake_loss = torch.tensor([0], dtype=torch.float)
fake_loss.requires_grad=True
return self.fake_param + fake_loss
else:
# in case of validation return MSE
return ((x_transformed-ys)**2).mean()
def log_posterior(self, cats, conts, ys):
ys = to_np(self.correct_shape(ys))
x_transformed = self.transform(cats, conts, as_np=True)
posteriors = np.zeros((len(self.prediction_models),1))
for idx, pred_model in enumerate(self.prediction_models):
log_posterior = pred_model.log_posterior(x_transformed, ys[:,idx].ravel())
posteriors[idx] = log_posterior
return posteriors
def log_evidence(self, cats, conts, ys, logme=False):
evidences = []
ys = to_np(self.correct_shape(ys))
x_transformed = self.transform(cats, conts, as_np=True)
for idx, pred_model in enumerate(self.prediction_models):
ev = pred_model.log_evidence(x_transformed, ys[:,idx].ravel())
evidences.append(ev)
evidences = np.array(evidences, dtype=np.float)
if logme:
evidences = evidences / len(conts)
return evidences.mean()
# hide
cats, conts, targets = dls_target.one_batch()
# test if we get the same results as with the original model
with torch.no_grad():
preds_source = source_model(cats, conts)
ltmodel = LinearTransferModel(source_model, num_layers_to_remove=1, use_original_weights=True)
preds_target = ltmodel.predict(cats, conts)
test_close(preds_source, preds_target)
preds_target = ltmodel.predict(cats, conts)
test_close(preds_source, preds_target)
preds_target = ltmodel.predict_proba(cats, conts)[0]
test_close(preds_source, preds_target)
```
# Synthetic data
## Linear Transfer Model
```
target_model = LinearTransferModel(source_model, num_layers_to_remove=1,
use_original_weights=True,
)
target_model = target_model.eval()
```
Create forecats and check if the source model and the untrained model create the same results
```
cats, conts, targets = dls_source.one_batch()
with torch.no_grad():
preds_source = source_model(cats, conts).ravel()
preds_target = target_model(cats.to("cpu"), conts.to("cpu"))
# targets, preds = learn_source.get_preds(ds_idx=0)
plt.scatter(conts, preds_target, label="preds target model - untrained", alpha=0.2)
plt.scatter(conts, preds_source.detach(), label="preds source model", alpha=0.2)
plt.scatter(conts, targets, label="targets")
plt.legend()
plt.show()
target_model = LinearTransferModel(source_model, num_layers_to_remove=1,
use_original_weights=True,
)
target_model.training = True
cats, conts, targets = convert_to_tensor(dls_target.train_ds)
preds_before_training = target_model.predict(cats, conts)
x_transformed = target_model.forward(cats, conts)
target_model = target_model.update(x_transformed, targets.ravel())
target_model.training = False
preds_after_training = target_model.forward(cats, conts)
((targets.reshape(-1)-preds_after_training.reshape(-1))**2).mean()**0.5
plt.scatter(conts, targets, label="targets")
plt.scatter(conts, preds_before_training, label="before training")
plt.scatter(conts, preds_after_training, label="after training")
plt.legend()
```
Lets assure that it also works with pytorch trainings such as the fastai training loop
```
target_model = LinearTransferModel(source_model,
num_layers_to_remove=1,
use_original_weights=True,
)
target_model = LinearTransferModel(source_model,
num_layers_to_remove=1,
use_original_weights=True,
)
target_learner = Learner(dls_target, target_model, loss_func=target_model.loss_func, metrics=rmse,)
target_learner.fit(1)
```
We can also not using the last layer as initializaition
```
target_model = LinearTransferModel(source_model,
num_layers_to_remove=1,
use_original_weights=False,
prediction_model=BayesLinReg(alpha=1, beta=1, empirical_bayes=False))
target_learner = Learner(dls_target, target_model, loss_func=target_model.loss_func, metrics=rmse,)
target_learner.fit(1)
```
Due to the few data samples, the empircal bayes is not working well.
```
target_model = LinearTransferModel(source_model,
num_layers_to_remove=1,
use_original_weights=False,
prediction_model=BayesLinReg(alpha=1, beta=1, empirical_bayes=True))
target_learner = Learner(dls_target, target_model, loss_func=target_model.loss_func, metrics=rmse,)
target_learner.fit(1)
```
Or remove multiple layers
```
target_model = LinearTransferModel(source_model,
num_layers_to_remove=2,
use_original_weights=False,
prediction_model=BayesLinReg(alpha=10, beta=10, empirical_bayes=False)
)
target_learner = Learner(dls_target, target_model, loss_func=target_model.loss_func, metrics=rmse,)
target_learner.fit(1)
```
## B Tuning
```
#export
class BTuningModel(nn.Module):
def __init__(self, source_model, b_tuning_models):
"""Based on Ranking and Tuning Pre-trained Models: A New Paradigm of Exploiting Model Hubs"""
super().__init__()
self.source_model = source_model
self.b_tuning_models = b_tuning_models
for model in self.b_tuning_models:
model.fake_param.requires_grad=False
model.eval()
def forward(self, cats, conts):
yhat_source_model = self.source_model(cats, conts)
yhat_b_tuning_models = []
n_samples = len(conts)
is_ts = True if len(yhat_source_model.shape) == 3 else False
for b_tuning_model in self.b_tuning_models:
cur_y_hat = b_tuning_model(cats, conts)
if is_ts:
cur_y_hat = cur_y_hat.reshape(n_samples, 1, -1)
yhat_b_tuning_models.append(cur_y_hat)
yhat_b_tuning_models = torch.cat(yhat_b_tuning_models, axis=1)
# if is_ts and len(yhat_b_tuning_models.shape) == 2:
# yhat_b_tuning_models = yhat_b_tuning_models[:, np.newaxis, :]
yhat_all = torch.cat([yhat_source_model, yhat_b_tuning_models], axis=1)
return yhat_all
```
# Test on real world data
## MLP
```
from fastrenewables.tabular.core import *
from fastrenewables.tabular.data import *
from fastrenewables.tabular.model import *
from fastrenewables.tabular.learner import *
cont_names = ['T_HAG_2_M', 'RELHUM_HAG_2_M', 'PS_SFC_0_M', 'ASWDIFDS_SFC_0_M',
'ASWDIRS_SFC_0_M', 'WindSpeed58m',
'SinWindDirection58m', 'CosWindDirection58m', 'WindSpeed60m',
'SinWindDirection60m', 'CosWindDirection60m', 'WindSpeed58mMinus_t_1',
'SinWindDirection58mMinus_t_1', 'CosWindDirection58mMinus_t_1',
'WindSpeed60mMinus_t_1', 'SinWindDirection60mMinus_t_1',
'CosWindDirection60mMinus_t_1', 'WindSpeed58mPlus_t_1',
'SinWindDirection58mPlus_t_1', 'CosWindDirection58mPlus_t_1',
'WindSpeed60mPlus_t_1', 'SinWindDirection60mPlus_t_1',
'CosWindDirection60mPlus_t_1']
cat_names = ['TaskID', 'Month', 'Day', 'Hour']
set_seed(23, reproducible=True)
dls_source = RenewableDataLoaders.from_files(glob.glob("../data/*.h5")[0], y_names="PowerGeneration",
pre_procs=[FilterYear(year=2020),
AddSeasonalFeatures(as_cont=False)],
cat_names=cat_names, cont_names=cont_names)
dls_source2 = RenewableDataLoaders.from_files(glob.glob("../data/*.h5")[1], y_names="PowerGeneration",
pre_procs=[FilterYear(year=2020),
AddSeasonalFeatures(as_cont=False)],
cat_names=cat_names, cont_names=cont_names)
dls_target = RenewableDataLoaders.from_files(glob.glob("../data/*.h5")[2], y_names="PowerGeneration",
pre_procs=[FilterYear(year=2020),
FilterMonths(months=[4]),
FilterDays(14),
AddSeasonalFeatures(as_cont=False)],
cat_names=cat_names, cont_names=cont_names)
dls_target_test = RenewableDataLoaders.from_files(glob.glob("../data/*.h5")[2], y_names="PowerGeneration",
pre_procs=[FilterYear(year=2020, drop=False),
AddSeasonalFeatures(as_cont=False)],
cat_names=cat_names, cont_names=cont_names)
learn_source = renewable_learner(dls_source, metrics=rmse)
learn_source.fit_one_cycle(5)
learn_source2 = renewable_learner(dls_source2, metrics=rmse)
learn_source2.fit_one_cycle(5)
```
When using empirical bayes (MacKay or FixedPoint) we need to set the size of a batch equal to the length of the complete data. As these algorithms are not supporting a batch wise optimization.
```
target_model = LinearTransferModel(learn_source.model, 1,
use_original_weights=True,
prediction_model=BayesLinReg(alpha=10, beta=10, empirical_bayes=False))
target_learner = RenewableLearner(dls_target, target_model, loss_func=target_model.loss_func, metrics=rmse,)
target_learner.fit(1)
preds, targets = target_learner.predict(ds_idx=1, filter=True, flatten=True)
plt.scatter(dls_target.valid_ds.items.WindSpeed58m, targets)
plt.scatter(dls_target.valid_ds.items.WindSpeed58m, preds)
target_model = LinearTransferModel(learn_source.model, 1,
use_original_weights=True,
prediction_model=BayesLinReg(alpha=10, beta=10, empirical_bayes=True))
target_learner = RenewableLearner(dls_target, target_model, loss_func=target_model.loss_func, metrics=rmse,)
target_learner.dls[0].bs=len(target_learner.dls.train_ds)
target_learner.fit(1)
target_model2 = LinearTransferModel(learn_source2.model, 1,
use_original_weights=True,
prediction_model=BayesLinReg(alpha=10, beta=10, empirical_bayes=True))
target_learner = RenewableLearner(dls_target, target_model, loss_func=target_model.loss_func, metrics=rmse,)
target_learner.dls[0].bs=len(target_learner.dls.train_ds)
target_learner.fit(1)
```
## B Tunning
```
target_learner = RenewableLearner(dls_target, copy.deepcopy(learn_source.model), metrics=rmse)
target_learner.fit(5, lr=5e-4)
preds, targets = target_learner.predict(test_dl=dls_target_test, filter=False, flatten=False)
((targets.ravel()-preds.ravel())**2).mean()**0.5
dls_target_test
source_model = copy.deepcopy(learn_source.model)
btuning_model = BTuningModel(source_model, [target_model,target_model2])
btuning_loss = BTuningLoss(lambd=1)
target_learner = RenewableLearner(dls_target, btuning_model, loss_func=btuning_loss, metrics=btuning_rmse)
preds, targets = target_learner.predict(test_dl=dls_target_test, filter=False, flatten=False)
print(btuning_rmse(preds, targets))
target_learner.fit(5, lr=5e-4)
preds, targets = target_learner.predict(test_dl=dls_target_test, filter=False, flatten=False)
btuning_rmse(preds, targets)
```
## TCN
### Source Model
```
from fastrenewables.timeseries.core import *
from fastrenewables.tabular.core import *
from fastrenewables.timeseries.data import *
from fastrenewables.timeseries.model import *
from fastrenewables.timeseries.learner import *
def get_dls(y_names="PowerGeneration"):
pd.options.mode.chained_assignment=None
dls_source = RenewableTimeSeriesDataLoaders.from_files(glob.glob("../data/*.h5")[0:1],
y_names=y_names,
cat_names=cat_names,
cont_names=cont_names,
pre_procs=[CreateTimeStampIndex(index_col_name="TimeUTC"),
FilterYear(year=2020),
AddSeasonalFeatures(as_cont=False),
FilterInconsistentSamplesPerDay],
procs=[Categorify, Normalize],
bs=12,
y_block=RegressionBlock(),
)
dls_target = RenewableTimeSeriesDataLoaders.from_files(glob.glob("../data/*.h5")[2],
y_names=y_names,
cat_names=cat_names,
cont_names=cont_names,
pre_procs=[CreateTimeStampIndex(index_col_name="TimeUTC"),
FilterYear(year=2020),
AddSeasonalFeatures(as_cont=False),
FilterInconsistentSamplesPerDay,
FilterMonths(months=[1,2,3,4])],
procs=[Categorify, Normalize],
bs=12,
y_block=RegressionBlock(),
)
dls_target_test = RenewableTimeSeriesDataLoaders.from_files(glob.glob("../data/*.h5")[2],
y_names=y_names,
cat_names=cat_names,
cont_names=cont_names,
pre_procs=[CreateTimeStampIndex(index_col_name="TimeUTC"),
FilterYear(year=2020, drop=False),
AddSeasonalFeatures(as_cont=False),
FilterInconsistentSamplesPerDay,
FilterMonths(months=[1,2,3,4])],
procs=[Categorify, Normalize],
bs=12,
y_block=RegressionBlock(),
)
return dls_source, dls_target,dls_target_test
set_seed(23, reproducible=True)
dls_source, dls_target, dls_target_test = get_dls()
n_features = len(dls_source.train_ds.cont_names)
learner = renewable_timeseries_learner(dls_source, metrics=rmse, layers=[n_features, 200, 100, 50, 25, 5, 1])
learner.fit_one_cycle(5)
preds, targets = learner.predict(ds_idx=1, filter=True)
id_ws = 5
```
We catch the the wind speed from the validation data to check the forecasts.
```
windspeed = dls_source.valid_ds.conts[:,id_ws,:]
windspeed = windspeed.reshape(-1,1)
plt.scatter(windspeed, targets)
plt.scatter(windspeed, preds)
```
### Target Model
```
# export
def reduce_layers_tcn_model(tcn_model, num_layers=0):
tcn_model.layers.temporal_blocks = tcn_model.layers.temporal_blocks[:-num_layers]
```
The problem is that we have a lot of features while less samples compared to the linear model.
```
target_model = LinearTransferModel(learner.model, 1,
reduce_layers_tcn_model,
use_original_weights=False,
prediction_model=BayesLinReg(10, 10, empirical_bayes=False),
)
target_learner = RenewableTimeseriesLearner(dls_target, target_model, loss_func=target_model.loss_func, metrics=rmse,)
target_learner.fit(1)
preds, targets = target_learner.predict(ds_idx=1, filter=True, flatten=True)
windspeed = dls_target.valid_ds.conts[:,id_ws,:]
windspeed = windspeed.reshape(-1,1)
plt.scatter(windspeed, targets)
plt.scatter(windspeed, preds)
cats, conts, ys = convert_to_tensor_ts(dls_target.train_ds)
target_model.log_posterior(cats, conts, ys).shape
target_model = LinearTransferModel(learner.model, 1,
reduce_layers_tcn_model,
use_original_weights=False,
prediction_model=BayesLinReg(1, 1, empirical_bayes=True),
)
target_learner = RenewableTimeseriesLearner(dls_target, target_model, loss_func=target_model.loss_func, metrics=rmse,)
target_learner.dls[0].bs=len(target_learner.dls.train_ds)
target_learner.fit(1)
target_model = LinearTransferModel(learner.model, 1,
reduce_layers_tcn_model,
use_original_weights=False,
prediction_model=BayesLinReg(1, 1, use_fixed_point=True),
)
target_learner = RenewableTimeseriesLearner(dls_target, target_model, loss_func=target_model.loss_func, metrics=rmse,)
target_learner.dls[0].bs=len(target_learner.dls.train_ds)
target_learner.fit(1)
preds, targets = target_learner.predict(test_dl=dls_target_test)
((targets-preds)**2).mean()**0.5
preds, targets = target_learner.predict(ds_idx=1, filter=True, flatten=True)
windspeed = dls_target.valid_ds.conts[:,id_ws,:]
windspeed = dls_target.valid_ds.conts[:,id_ws,:]
windspeed = windspeed.reshape(-1,1)
plt.scatter(windspeed.ravel(), targets)
plt.scatter(windspeed.ravel(), preds)
source_model = copy.deepcopy(learner.model)
btuning_model = BTuningModel(source_model, [target_model])
btuning_loss = BTuningLoss(lambd=1)
target_learner = RenewableTimeseriesLearner(dls_target, btuning_model, loss_func=btuning_loss, metrics=btuning_rmse)
preds, targets = target_learner.predict(test_dl=dls_target_test, filter=False, flatten=False)
print(btuning_rmse(preds, targets))
target_learner.fit(5, lr=5e-4)
preds, targets = target_learner.predict(test_dl=dls_target_test, filter=False, flatten=False)
btuning_rmse(preds, targets)
```
| github_jupyter |
# Introduction
This example demonstrates how to convert a network from [Caffe's Model Zoo](https://github.com/BVLC/caffe/wiki/Model-Zoo) for use with Lasagne. We will be using the [NIN model](https://gist.github.com/mavenlin/e56253735ef32c3c296d) trained for CIFAR10.
We will create a set of Lasagne layers corresponding to the Caffe model specification (prototxt), then copy the parameters from the caffemodel file into our model.
# Final product
If you just want to try the final result, you can download the pickled weights [here](https://s3.amazonaws.com/lasagne/recipes/pretrained/cifar10/model.pkl)
# Converting from Caffe to Lasagne
### Download the required files
First we download `cifar10_nin.caffemodel` and `model.prototxt`. The supplied `train_val.prototxt` was modified to replace the data layers with an input specification, and remove the unneeded loss/accuracy layers.
```
!wget https://www.dropbox.com/s/blrajqirr1p31v0/cifar10_nin.caffemodel
!wget https://gist.githubusercontent.com/ebenolson/91e2cfa51fdb58782c26/raw/b015b7403d87b21c6d2e00b7ec4c0880bbeb1f7e/model.prototxt
```
### Import Caffe
To load the saved parameters, we'll need to have Caffe's Python bindings installed.
```
import caffe
```
### Load the pretrained Caffe network
```
net_caffe = caffe.Net('model.prototxt', 'cifar10_nin.caffemodel', caffe.TEST)
```
### Import Lasagne
```
import lasagne
from lasagne.layers import InputLayer, DropoutLayer, FlattenLayer
from lasagne.layers.dnn import Conv2DDNNLayer as ConvLayer
from lasagne.layers import Pool2DLayer as PoolLayer
from lasagne.utils import floatX
```
### Create a Lasagne network
Layer names match those in `model.prototxt`
```
net = {}
net['input'] = InputLayer((None, 3, 32, 32))
net['conv1'] = ConvLayer(net['input'], num_filters=192, filter_size=5, pad=2)
net['cccp1'] = ConvLayer(net['conv1'], num_filters=160, filter_size=1)
net['cccp2'] = ConvLayer(net['cccp1'], num_filters=96, filter_size=1)
net['pool1'] = PoolLayer(net['cccp2'], pool_size=3, stride=2, mode='max', ignore_border=False)
net['drop3'] = DropoutLayer(net['pool1'], p=0.5)
net['conv2'] = ConvLayer(net['drop3'], num_filters=192, filter_size=5, pad=2)
net['cccp3'] = ConvLayer(net['conv2'], num_filters=192, filter_size=1)
net['cccp4'] = ConvLayer(net['cccp3'], num_filters=192, filter_size=1)
net['pool2'] = PoolLayer(net['cccp4'], pool_size=3, stride=2, mode='average_exc_pad', ignore_border=False)
net['drop6'] = DropoutLayer(net['pool2'], p=0.5)
net['conv3'] = ConvLayer(net['drop6'], num_filters=192, filter_size=3, pad=1)
net['cccp5'] = ConvLayer(net['conv3'], num_filters=192, filter_size=1)
net['cccp6'] = ConvLayer(net['cccp5'], num_filters=10, filter_size=1)
net['pool3'] = PoolLayer(net['cccp6'], pool_size=8, mode='average_exc_pad', ignore_border=False)
net['output'] = lasagne.layers.FlattenLayer(net['pool3'])
```
### Copy the parameters from Caffe to Lasagne
```
layers_caffe = dict(zip(list(net_caffe._layer_names), net_caffe.layers))
for name, layer in net.items():
try:
layer.W.set_value(layers_caffe[name].blobs[0].data)
layer.b.set_value(layers_caffe[name].blobs[1].data)
except AttributeError:
continue
```
# Trying it out
Let's see if that worked.
### Import numpy and set up plotting
```
import numpy as np
import pickle
import matplotlib.pyplot as plt
%matplotlib inline
```
### Download some test data
Since the network expects ZCA whitened and normalized input, we'll download a preprocessed portion (1000 examples) of the CIFAR10 test set.
```
!wget https://s3.amazonaws.com/lasagne/recipes/pretrained/cifar10/cifar10.npz
data = np.load('cifar10.npz')
```
### Make predictions on the test data
```
prob = np.array(lasagne.layers.get_output(net['output'], floatX(data['whitened']), deterministic=True).eval())
predicted = np.argmax(prob, 1)
```
### Check our accuracy
We expect around 90%
```
accuracy = np.mean(predicted == data['labels'])
print(accuracy)
```
### Double check
Let's compare predictions against Caffe
```
net_caffe.blobs['data'].reshape(1000, 3, 32, 32)
net_caffe.blobs['data'].data[:] = data['whitened']
prob_caffe = net_caffe.forward()['pool3'][:,:,0,0]
np.allclose(prob, prob_caffe)
```
### Graph some images and predictions
```
def make_image(X):
im = np.swapaxes(X.T, 0, 1)
im = im - im.min()
im = im * 1.0 / im.max()
return im
plt.figure(figsize=(16, 5))
for i in range(0, 10):
plt.subplot(1, 10, i+1)
plt.imshow(make_image(data['raw'][i]), interpolation='nearest')
true = data['CLASSES'][data['labels'][i]]
pred = data['CLASSES'][predicted[i]]
color = 'green' if true == pred else 'red'
plt.text(0, 0, true, color='black', bbox=dict(facecolor='white', alpha=1))
plt.text(0, 32, pred, color=color, bbox=dict(facecolor='white', alpha=1))
plt.axis('off')
```
### Save our model
Let's save the weights in pickle format, so we don't need Caffe next time
```
import pickle
values = lasagne.layers.get_all_param_values(net['output'])
pickle.dump(values, open('model.pkl', 'w'))
```
| github_jupyter |
# Functions to select best pitch to throw in given baseball situation
```
# Import statements
import pandas as pd
```
### Function to combine pitcher and batter outcome histories
```
def combine_histories(pitcher_history: pd.DataFrame, batter_history: pd.DataFrame,
method: str = 'product', extra_pitches: bool = False) -> pd.DataFrame:
'''
Combines outcome histories to produce outcome metrics for pitch selections. Returns pandas dataframe.
Method argument can be 'product', 'min', or 'max'. Describes how cells are combined to get result.
Extra pitches specifies if pitches in pitcher's history but not batter's are included.
Method and extra pitches args default to 'product' and False, respectively.
'''
# Check if dataframes requires numeric conversion
# Pitcher
if pitcher_history.Ball.dtype.kind is 'O': # Object dtype
pitcher_history = bb_webscraper.perc_to_dec(pitcher_history,ignore_cols='Count')
# Batter
if batter_history.Ball.dtype.kind is 'O': # Object dtype
batter_history = bb_webscraper.perc_to_dec(batter_history,ignore_cols='Count')
# Product method
combined_history = pitcher_history # Placeholder df
combined_history.iloc[:,1:] = pitcher_history.iloc[:,1:].multiply(batter_history.iloc[:,1:]) # Product
combined_history.Count += batter_history.Count # Total pitch counts
combined_history.dropna() # Drop rows/cols where pitch was not in both players' data
# Return resulting dataframe
return(combined_history)
```
### Pitch Selector Function
```
def pitch_selector(pitcher: str, batter: str, balls: int, strikes: int, outs: int,
first: int, second: int, third: int, inning: int = None, season: int = None,
runs_scored:int = 0, runs_allowed:int = 0) -> dict:
'''
Takes in game state info, calculates best pitch to throw given game state, pitcher history,
and batter history, then returns dict of best pitch, location, and other important variables.
First, second, and third should be 1 for occupied, 0 for empty.
Note: Need to think of how to pass extra parameters to internal functions.
>>> pitch_selector('Dallas Keuchel', 'Albert Almora', balls=1, strikes=2, outs=1,
first=1, second=0, third=0, inning=3, season=2019)['Pitch']
'Changeup'
'''
# Initialize return dict
results = {}
# Scrape pitcher and batter histories in given game state
pitcher_history = bb_webscraper.scrape_brooksbb(player=pitcher, batter_hand='R', season=season,
pitcher_or_batter='pitcher',table_type='po',
params_dict={'balls':balls,'strikes':strikes,'1b':first,
'2b':second,'3b':third})
#print("Pitcher Data: \n", pitcher_history)
batter_history = bb_webscraper.scrape_brooksbb(player=batter, pitcher_hand='R', season=season,
pitcher_or_batter='batter',table_type='po',
params_dict={'balls':balls,'strikes':strikes,'1b':first,
'2b':second,'3b':third})
#print("Batter Data: \n", batter_history)
# Create game state df and decide desired outcome
game_state = pd.DataFrame(data={"Inning":inning, "Outs":outs, "Strikes":strikes, "Balls":balls,
"First":bool(first), "Second":bool(second), "Third":bool(third),
"R":0, "RA":0}, index=[0])
game_state['Desired Outcome'] = outcomes.desired_outcomes(game_state)
desired_outcome = game_state.loc[0,'Desired Outcome']
#print("Desired Outcome:", desired_outcome)
# Combine pitcher and batter histories for outcome metrics
combined_histories = combine_histories(pitcher_history,batter_history)
#print("Combined Histories: \n", combined_histories)
# Choose pitch with greatest chance of producign desired result based on combined histories
best_pitch = combined_histories[desired_outcome].idxmax()
# Choose best pitch location !!!WORK IN PROGRESS!!!
location = outcomes.pitch_for_outcome().loc[desired_outcome,'Location']
# Fill result dict and return it
results['Pitch'] = best_pitch
results['Location'] = location
results['Desired Outcome'] = desired_outcome
return(results)
# Given game state, i.e. 1 row of info...
# 1. Decide desired outcome
# 2. Retrieve pitcher and batter outcome histories (maybe add pitcher/batter names to df)
# 3. Left join data so only pitches in pitcher's repertoire are considered (pd.merge)
# a. Make sure data is converted from string percents to decimal values for numerical work
# b. Maybe multiply dfs elementwise upon merging to combine batter/pitcher probabilities
# -> e.g. 'Fourseam, Whiffs' is 0.25 for pitcher, 0.4 for batter, 0.1 resulting
# -> Weighing the multiplication by pitches thrown might be good, but requires
# exponentiation and some basic probability concepts. Start without weights for now.
# -> Maybe taking the min or max is smarter... try prod,weights,min,max and compare? :)
# c. Note: If we already know the outcome we want, might just need 1 column from each outcome df
# 4. Choose pitch with maximum chances of desired result.
```
| github_jupyter |
```
from google.colab import drive
drive.mount('/content/drive')
!pip install -U pywsd
!pip install -U wn==0.0.23
!pip install XlsxWriter
```
# partOf: wsd measurement
```
import pandas as pd
from tabulate import tabulate
from pywsd.cosine import cosine_similarity
from nltk.stem import PorterStemmer
from nltk.corpus import stopwords
from nltk.tokenize import word_tokenize
from nltk.stem.wordnet import WordNetLemmatizer
from sklearn import metrics
from sklearn.metrics import confusion_matrix
# nltk.download('averaged_perceptron_tagger')
# nltk.download('punkt')
# nltk.download('stopwords')
# nltk.download('wordnet')
stemming = PorterStemmer()
stops = set(stopwords.words("english"))
lem = WordNetLemmatizer()
class wsd_partof:
def __init__(self):
pass
def fulldataset(self, dataFile, inputSRS):
xl = pd.ExcelFile(dataFile)
dfs = {sh:xl.parse(sh) for sh in xl.sheet_names}
kalimat = dfs[inputSRS]
kalimat_semua = kalimat.head(len(kalimat))
return kalimat_semua
def preprocessing(self, dataFile):
xl = pd.ExcelFile(dataFile)
for sh in xl.sheet_names:
df = xl.parse(sh)
print('Processing: [{}] ...'.format(sh))
print(df.head())
# cleaning text
def apply_cleaning_function_to_list(self, X):
cleaned_X = []
for element in X:
cleaned_X.append(wsd_partof.clean_text(self, raw_text= element))
return cleaned_X
def clean_text(self, raw_text):
text = raw_text.lower()
tokens = word_tokenize(text)
token_words = [w for w in tokens if w.isalpha()]
lemma_words = [lem.lemmatize(w) for w in token_words]
meaningful_words = [w for w in lemma_words if not w in stops]
joined_words = ( " ".join(meaningful_words))
return joined_words
if __name__ == "__main__":
try:
myWsd_partof = wsd_partof()
# myWsd_partof.preprocessing(dataFile= r'/content/drive/MyDrive/dataset/dataset_2_split.xlsx')
file1 = r'/content/drive/MyDrive/dataset/dataset_2.xlsx'
dataSRS = '2005 - Grid 3D'
a = myWsd_partof.fulldataset(dataFile= file1, inputSRS= dataSRS)
list_req1 = list(a['Requirement Statement'])
id_req1 = list(a['ID'])
cleaned1 = myWsd_partof.apply_cleaning_function_to_list(X= list_req1)
file2 = r'/content/drive/MyDrive/dataset/dataset_2_split.xlsx'
b = myWsd_partof.fulldataset(dataFile= file2, inputSRS= dataSRS)
list_req2 = list(b['Requirement Statement'])
id_req2 = list(b['ID'])
cleaned2 = myWsd_partof.apply_cleaning_function_to_list(X= list_req2)
hasil_wsd = []
for num in cleaned1:
text = [cosine_similarity(num, angka) for angka in cleaned2]
hasil_wsd.append(text)
data_raw = pd.DataFrame(hasil_wsd, index= id_req1, columns= id_req2)
print("Hasil pengukuran semantik antar kebutuhan atomik dan non atomik {}".format(dataSRS))
print(tabulate(data_raw, headers = 'keys', tablefmt = 'psql'))
# thresholding
threshold = 0.1
d = data_raw.values >= threshold
d1 = pd.DataFrame(d, index= id_req1, columns= id_req2)
mask = d1.isin([True])
d2 = d1.where(mask, other= 0)
mask2 = d1.isin([False])
d3 = d2.where(mask2, other= 1)
print("\nHasil ukur semantik diatas threshold {}".format(threshold))
print(tabulate(d3, headers = 'keys', tablefmt = 'psql'))
file3 = r'/content/drive/MyDrive/dataset/wsd/wsd_groundtruth.xlsx'
dataGT = 'grid3d_eval'
b3 = fulldataset(data= file3, inputSRS= dataGT)
b3 = b3.drop(['Index'], axis= 1)
b3.index= d3.index
print("\nData Hasil Ground Truth {}".format(dataGT))
print(tabulate(b3, headers = 'keys', tablefmt = 'psql'))
y_actual = d3.values.astype(int)
y_predicted = b3.values.astype(int)
print("akurasi", metrics.accuracy_score(y_true= d3_list, y_pred= b3_list))
print("presion", metrics.precision_score(y_true= y_actual, y_pred= y_predicted, average= 'macro'))
print("recall", metrics.recall_score(y_true= y_actual, y_pred= y_predicted, average= 'macro'))
print("metrics {}".format(metrics.classification_report(y_true= y_actual, y_pred= y_predicted)))
except OSError as err:
print("OS error: {0}".format(err))
import xlsxwriter
import pandas as pd
dfs = {
'tabel_dataset' : data_raw,
'tabel_threshold' : d3,
'tabel_groundtruth' : b3,
}
writer = pd.ExcelWriter('/content/mydrive/MyDrive/dataset/wsd/data_wsd.xlsx')
for name,dataframe in dfs.items():
dataframe.to_excel(writer,name,index=False)
writer.save()
```
# manual
```
from sklearn import metrics
from sklearn.metrics import confusion_matrix
import pandas as pd
#define array of actual values
y_actual = [
1, 1, 1, 1, 1, 1, 0, 0, 0, 0, 0, 0, 1, 1, 1, 1, 1, 0, 0, 0, 1, 1, 1, 1, 1, 0, 0, 1, 1, 1, 1, 1, 0, 1, 1, 1, 1, 1, 0, 0, 0, 1, 1, 1, 1, 1, 0, 0, 0, 0, 1, 1, 1, 1, 1, 0, 0, 0, 0, 1, 1, 1, 1, 1, 0, 0, 0, 1, 1, 1, 1, 1, 0, 1, 1, 1, 1, 1, 0, 0, 0, 1, 1, 1, 1, 1, 0, 0, 0, 1, 1, 1, 1, 1, 0, 0, 0, 0, 1, 1, 1, 1, 1, 0, 0, 0, 1, 1, 1, 1, 1,
0, 0, 0, 0, 0, 0, 1, 1, 1, 1, 1, 0, 0, 0, 0, 0, 0, 1, 1, 1, 0, 0, 0, 0, 0, 1, 1, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 1, 1, 1, 0, 0, 0, 0, 0, 1, 1, 1, 1, 0, 0, 0, 0, 0, 1, 1, 1, 1, 0, 0, 0, 0, 0, 1, 1, 1, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 1, 1, 1, 0, 0, 0, 0, 0, 1, 1, 1, 0, 0, 0, 0, 0, 1, 1, 1, 1, 0, 0, 0, 0, 0, 1, 1, 1, 0, 0, 0, 0, 0,
0, 0, 1, 1, 0, 0, 1, 0, 0, 0, 0, 1, 0, 1, 1, 0, 0, 1, 0, 0, 0, 1, 1, 0, 0, 1, 0, 0, 1, 1, 0, 0, 1, 0, 1, 1, 0, 0, 1, 0, 0, 0, 1, 1, 0, 0, 1, 0, 0, 0, 0, 1, 1, 0, 0, 1, 0, 0, 0, 0, 1, 1, 0, 0, 1, 0, 0, 0, 1, 1, 0, 0, 1, 0, 1, 1, 0, 0, 1, 0, 0, 0, 1, 1, 0, 0, 1, 0, 0, 0, 1, 1, 0, 0, 1, 0, 0, 0, 0, 1, 1, 0, 0, 1, 0, 0, 0, 1, 1, 0, 0,
0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1,
0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 1, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 1, 0, 0, 0, 1, 0, 0, 0, 1, 0, 0, 1, 0, 0, 0, 1, 0, 1, 0, 0, 0, 1, 0, 0, 0, 1, 0, 0, 0, 1, 0, 0, 0, 0, 1, 0, 0, 0, 1, 0, 0, 0, 0, 1, 0, 0, 0, 1, 0, 0, 0, 1, 0, 0, 0, 1, 0, 1, 0, 0, 0, 1, 0, 0, 0, 1, 0, 0, 0, 1, 0, 0, 0, 1, 0, 0, 0, 1, 0, 0, 0, 0, 1, 0, 0, 0, 1, 0, 0, 0, 1, 0, 0, 0, 1,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 0, 0, 0, 0, 0, 0, 1, 1, 0, 0, 0, 0, 0, 1, 1, 0, 0, 0, 0, 1, 1, 0, 0, 0, 0, 0, 0, 1, 1, 0, 0, 0, 0, 0, 0, 0, 1, 1, 0, 0, 0, 0, 0, 0, 0, 1, 1, 0, 0, 0, 0, 0, 0, 1, 1, 0, 0, 0, 0, 1, 1, 0, 0, 0, 0, 0, 0, 1, 1, 0, 0, 0, 0, 0, 0, 1, 1, 0, 0, 0, 0, 0, 0, 0, 1, 1, 0, 0, 0, 0, 0, 0, 1, 1, 0, 0,
]
#define array of predicted values
y_predicted = [
1, 1, 1, 1, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 1, 1, 1, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 1, 1, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 1, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 1, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1,
]
tn, fp, fn, tp = confusion_matrix(y_true= y_actual, y_pred= y_predicted).ravel()
print("false positif : ",fp)
print("false negative : ",fn)
print("true positive : ", tp)
print("true negative : ", tn)
print("akurasi", metrics.accuracy_score(y_true= y_actual, y_pred= y_predicted))
print("recall", metrics.recall_score(y_true= y_actual, y_pred= y_predicted))
print("presion", metrics.precision_score(y_true= y_actual, y_pred= y_predicted))
d3.reset_index(drop=True, inplace=True)
d3
from pywsd import disambiguate
from pywsd.similarity import similarity_by_path
file1 = r'/content/drive/MyDrive/dataset/dataset_2.xlsx'
dataSRS = '2004 - colorcast'
a = fulldataset(data= file1, inputSRS= dataSRS)
list_req1 = list(a['Requirement Statement'])
id_req1 = list(a['ID'])
cleaned1 = apply_cleaning_function_to_list(list_req1)
file2 = r'/content/drive/MyDrive/dataset/dataset_2_split.xlsx'
b = fulldataset(data= file2, inputSRS= dataSRS)
list_req2 = list(b['Requirement Statement'])
id_req2 = list(b['ID'])
cleaned2 = apply_cleaning_function_to_list(list_req2)
word1 = [disambiguate(x) for x in cleaned1 if x is not None]
word1_synset = [[n[1] for n in y if n[1] is not None] for y in word1]
word1_kata = [[n[0] for n in y if n[0] is not None] for y in word1]
word2 = [disambiguate(x) for x in cleaned2 if x is not None]
word2_synset = [[n[1] for n in y if n[1] is not None] for y in word2]
word2_kata = [[n[0] for n in y if n[1] is not None] for y in word2]
id1 = 7
id2 = 19
data_list = []
for idx, num in zip(word1_kata[id1], word1_synset[id1]):
a = [(similarity_by_path(num, angka, option= "wup")) for idy, angka in enumerate(word2_synset[id2])]
data_list.append(a)
df_sem = pd.DataFrame(data_list, index= word1_synset[id1], columns= word2_synset[id2])
df_sem
import numpy as np
A=np.array(data_list)
B=np.array(data_list)
def csm(A,B,corr):
if corr:
B=B-B.mean(axis=1)[:,np.newaxis]
A=A-A.mean(axis=1)[:,np.newaxis]
num=np.dot(A,B.T)
p1=np.sqrt(np.sum(A**2,axis=1))[:,np.newaxis]
p2=np.sqrt(np.sum(B**2,axis=1))[np.newaxis,:]
return num/(p1*p2)
hasil = (csm(A,B,True))
pd.DataFrame(hasil, )
import math
import re
from collections import Counter
WORD = re.compile(r"\w+")
def get_cosine(vec1, vec2):
intersection = set(vec1.keys()) & set(vec2.keys())
numerator = sum([vec1[x] * vec2[x] for x in intersection])
sum1 = sum([vec1[x] ** 2 for x in list(vec1.keys())])
sum2 = sum([vec2[x] ** 2 for x in list(vec2.keys())])
denominator = math.sqrt(sum1) * math.sqrt(sum2)
if not denominator:
return 0.0
else:
return float(numerator) / denominator
def text_to_vector(text):
words = WORD.findall(text)
return Counter(words)
text1 = "This is a foo bar sentence ."
text2 = "This sentence is similar to a foo bar sentence ."
vector1 = text_to_vector(text1)
vector2 = text_to_vector(text2)
print(vector1)
print(vector2)
cosine = get_cosine(vector1, vector2)
print("Cosine:", cosine)
from sklearn import metrics
from sklearn.metrics import accuracy_score, confusion_matrix
import pandas as pd
#define array of actual values
y_actual = [
]
#define array of predicted values
y_predicted = [
1,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,
0,0,1,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,
0,0,0,0,1,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,
0,0,0,0,0,0,1,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,
0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,
0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,
0,0,0,0,0,0,0,0,0,0,1,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,
0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,
0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,
0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,
0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,
0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,
0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,
0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,
0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,
0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,
0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,
0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,
0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,
0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,
0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,
0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,
0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,1,1,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,
0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,
0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,
0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,
0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,
0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,
0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,
0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,1,0,0,0,0,0,0,0,0,
0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,
0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,
0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,
0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,1,0,0,0,
0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,1,0,
0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,
]
# tn, fp, fn, tp = confusion_matrix(predict_train,y_train).ravel()
tn, fp, fn, tp = confusion_matrix(y_predicted,y_actual).ravel()
print("True Negatives : ",tn)
print("False Positives : ",fp)
print("False Negatives : ", fn)
print("True Positives : ", tp)
list1= set(['product', 'plot', 'data', 'point', 'scientific', 'correct', 'manner', 'grid', 'axis', 'label', 'correct', 'accord', 'input', 'data', 'file',
'data', 'point', 'colour', 'accord', 'cluster', 'number', 'contain', 'file', 'single', 'click', 'mouse', 'data', 'point', 'bring', 'name', 'mouse', 'data',
'point', 'cause', 'application', 'display', 'detail', 'product', 'allow', 'multiple', 'point', 'click', 'name', 'display', 'product', 'allow', 'grid', 'orient',
'user', 'rotation', 'zoom', 'move', 'function', 'employ', 'data' , 'file', 'contain', 'name', 'point', 'parameter', 'plot', 'single', 'designate', 'colour',
'attribute', 'used', 'comparison', 'description', 'point', 'large', 'enough', 'see', 'select', 'point', 'big', 'distort', 'overall', 'pattern', 'spread',
'axis', 'clearly', 'label', 'easily', 'recognise', 'grid', 'orient', 'different', 'position', 'application', 'colour', 'screen', 'shot', 'print',
'clear', 'black', 'white', 'background', 'application', 'intuitive', 'require', 'specialist', 'training', 'program', 'start', 'within', 'second', 'depends',
'number', 'data', 'point', 'plotted', 'interaction', 'data', 'point', 'delay', 'long', 'second', 'response', 'change', 'orientation', 'fast', 'enough',
'avoid', 'interrupt', 'user', 'flow'])
list2 = set(['product', 'allow', 'grid', 'orient', 'user', 'rotation', 'function', 'employed', 'zoom', 'function', 'employed', 'move', 'function', 'employed', 'data',
'file', 'contain', 'name', 'data', 'point', 'data', 'file', 'contain', 'parameter', 'data', 'point', 'plot', 'data', 'file', 'contain', 'single', 'parameter',
'designate', 'colour', 'point', 'attribute', 'used', 'comparison', 'description', 'data', 'point', 'point', 'large', 'enough', 'see', 'point', 'large', 'enough',
'select', 'axis', 'clearly', 'labelled', 'axis', 'easily', 'recognised', 'grid', 'oriented', 'different', 'position', 'application', 'coloured', 'screen',
'shot', 'printed', 'clearly', 'black', 'white', 'background', 'application', 'intuitive', 'application', 'require', 'specialist', 'training', 'program', 'start',
'within', 'second', 'depends', 'number', 'data', 'point', 'plot'])
from pywsd import disambiguate
from pywsd.similarity import similarity_by_path
word1 = [disambiguate(x) for x in list1]
word1_synset = [[n[1] for n in y] for y in word1]
word1_kata = [[n[0] for n in y] for y in word1]
word2 = [disambiguate(x) for x in list2]
word2_synset = [[n[1] for n in y] for y in word2]
word2_kata = [[n[0] for n in y] for y in word2]
data_list= []
for idx, num in zip(word1_kata, word1_synset):
a = [similarity_by_path(num[0], angka[0], option= "wup") for idy, angka in zip(word2_kata, word2_synset)]
data_list.append(a)
df_list = pd.DataFrame(data_list, index= list1, columns= list2)
df_list['product']['data']
```
# Another method
```
import numpy as np
import nltk
from nltk.corpus import wordnet as wn
import pandas as pd
def convert_tag(tag):
"""Convert the tag given by nltk.pos_tag to the tag used by wordnet.synsets"""
tag_dict = {'N': 'n', 'J': 'a', 'R': 'r', 'V': 'v'}
try:
return tag_dict[tag[0]]
except KeyError:
return None
def doc_to_synsets(doc):
"""
Returns a list of synsets in document.
Tokenizes and tags the words in the document doc.
Then finds the first synset for each word/tag combination.
If a synset is not found for that combination it is skipped.
Args:
doc: string to be converted
Returns:
list of synsets
Example:
doc_to_synsets('Fish are nvqjp friends.')
Out: [Synset('fish.n.01'), Synset('be.v.01'), Synset('friend.n.01')]
"""
tokens = nltk.word_tokenize(doc)
pos = nltk.pos_tag(tokens)
tags = [tag[1] for tag in pos]
wntag = [convert_tag(tag) for tag in tags]
ans = list(zip(tokens,wntag))
sets = [wn.synsets(x,y) for x,y in ans]
final = [val[0] for val in sets if len(val) > 0]
return final
def similarity_score(s1, s2):
"""
Calculate the normalized similarity score of s1 onto s2
For each synset in s1, finds the synset in s2 with the largest similarity value.
Sum of all of the largest similarity values and normalize this value by dividing it by the
number of largest similarity values found.
Args:
s1, s2: list of synsets from doc_to_synsets
Returns:
normalized similarity score of s1 onto s2
Example:
synsets1 = doc_to_synsets('I like cats')
synsets2 = doc_to_synsets('I like dogs')
similarity_score(synsets1, synsets2)
Out: 0.73333333333333339
"""
s =[]
for i1 in s1:
r = []
scores = [x for x in [i1.path_similarity(i2) for i2 in s2] if x is not None]
if scores:
s.append(max(scores))
return sum(s)/len(s)
def document_path_similarity(doc1, doc2):
"""Finds the symmetrical similarity between doc1 and doc2"""
synsets1 = doc_to_synsets(doc1)
synsets2 = doc_to_synsets(doc2)
return (similarity_score(synsets1, synsets2) + similarity_score(synsets2, synsets1)) / 2
doc_to_synsets('Fish are nvqjp friends.')
synsets1 = doc_to_synsets('I like cats')
synsets2 = doc_to_synsets('I like dogs')
similarity_score(synsets1, synsets2)
doc1 = "I like you"
doc2 = "I love you"
document_path_similarity(doc1, doc2)
def test_document_path_similarity():
doc1 = 'This is a function to test document_path_similarity.'
doc2 = 'Use this function to see if your code in doc_to_synsets \
and similarity_score is correct!'
return document_path_similarity(doc1, doc2)
test_document_path_similarity()
def most_similar_docs():
paraphrases = cleaned1
paraphrases['similarity_score'] = paraphrases.apply(lambda x:document_path_similarity(x['D1'], x['D2']), axis=1)
return (paraphrases.sort_values('similarity_score', ascending=False).iloc[0]['D1'], paraphrases.sort_values('similarity_score', ascending=False).iloc[0]['D2'], paraphrases.sort_values('similarity_score', ascending=False).iloc[0]['similarity_score'])
most_similar_docs()
def label_accuracy():
from sklearn.metrics import accuracy_score
paraphrases['similarity_score'] = paraphrases.apply(lambda x:document_path_similarity(x['D1'], x['D2']), axis=1)
paraphrases['predicted'] = np.where(paraphrases['similarity_score'] > 0.75, 1, 0)
return accuracy_score(paraphrases['Quality'], paraphrases['predicted'])
```
| github_jupyter |
# Decision Tree Classification with Standard Scalar
This Code template is for the Classification task using simple DecisionTreeClassifier based on the Classification and Regression Trees algorithm along with Feature scaling technique StandardScaler.
### Required Packages
```
!pip install imblearn
import numpy as np
import pandas as pd
import seaborn as se
import warnings
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from imblearn.over_sampling import RandomOverSampler
from sklearn.tree import DecisionTreeClassifier,plot_tree
from sklearn.preprocessing import LabelEncoder
from sklearn.metrics import classification_report,plot_confusion_matrix
warnings.filterwarnings('ignore')
```
### Initialization
Filepath of CSV file
```
file_path= ""
```
List of features which are required for model training .
```
features=[]
```
Target feature for prediction.
```
target=''
```
### Data Fetching
Pandas is an open-source, BSD-licensed library providing high-performance, easy-to-use data manipulation and data analysis tools.
We will use panda's library to read the CSV file using its storage path.And we use the head function to display the initial row or entry.
```
df=pd.read_csv(file_path);
df.head()
```
### Feature Selections
It is the process of reducing the number of input variables when developing a predictive model. Used to reduce the number of input variables to both reduce the computational cost of modelling and, in some cases, to improve the performance of the model.
We will assign all the required input features to X and target/outcome to Y.
```
X=df[features]
Y=df[target]
```
### Data Preprocessing
Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
```
def NullClearner(df):
if(isinstance(df, pd.Series) and (df.dtype in ["float64","int64"])):
df.fillna(df.mean(),inplace=True)
return df
elif(isinstance(df, pd.Series)):
df.fillna(df.mode()[0],inplace=True)
return df
else:return df
def EncodeX(df):
return pd.get_dummies(df)
def EncodeY(df):
if len(df.unique())<=2:
return df
else:
un_EncodedT=np.sort(pd.unique(df), axis=-1, kind='mergesort')
df=LabelEncoder().fit_transform(df)
EncodedT=[xi for xi in range(len(un_EncodedT))]
print("Encoded Target: {} to {}".format(un_EncodedT,EncodedT))
return df
```
Calling preprocessing functions on the feature and target set.
```
x=X.columns.to_list()
for i in x:
X[i]=NullClearner(X[i])
X=EncodeX(X)
Y=EncodeY(NullClearner(Y))
X.head()
```
#### Correlation Map
In order to check the correlation between the features, we will plot a correlation matrix. It is effective in summarizing a large amount of data where the goal is to see patterns.
```
f,ax = plt.subplots(figsize=(18, 18))
matrix = np.triu(X.corr())
se.heatmap(X.corr(), annot=True, linewidths=.5, fmt= '.1f',ax=ax, mask=matrix)
```
#### Distribution Of Target Variable
```
plt.figure(figsize = (10,6))
se.countplot(Y)
```
### Data Splitting
The train-test split is a procedure for evaluating the performance of an algorithm. The procedure involves taking a dataset and dividing it into two subsets. The first subset is utilized to fit/train the model. The second subset is used for prediction. The main motive is to estimate the performance of the model on new data.
```
x_train,x_test,y_train,y_test=train_test_split(X,Y,test_size=0.2,random_state=123)
```
<h3>Data Scaling </h3><br>
Standardizing a dataset involves rescaling the distribution of values so that the mean of observed values is 0 and the standard deviation is 1.
Like normalization, standardization can be useful, when your data has input values with differing scales. Standardization assumes that your observations fit a Gaussian distribution (bell curve) with a well-behaved mean and standard deviation.
```
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
x_train = scaler.fit_transform(x_train)
x_test = scaler.fit_transform(x_test)
```
#### Handling Target Imbalance
The challenge of working with imbalanced datasets is that most machine learning techniques will ignore, and in turn have poor performance on, the minority class, although typically it is performance on the minority class that is most important.
One approach to addressing imbalanced datasets is to oversample the minority class. The simplest approach involves duplicating examples in the minority class.We will perform overspampling using imblearn library.
```
x_train,y_train = RandomOverSampler(random_state=123).fit_resample(x_train, y_train)
```
### Model
Decision tree is the most powerful and popular tool for classification and prediction. A Decision tree is a flowchart like tree structure, where each internal node denotes a test on an attribute, each branch represents an outcome of the test, and each leaf node holds a outcome label.
As with other classifiers, DecisionTreeClassifier takes as input two arrays: an array X, sparse or dense, of shape (n_samples, n_features) holding the training samples, and an array Y of integer values, shape (n_samples,), holding the class labels for the training samples.
It is capable of both binary ([-1,1] or [0,1]) classification and multiclass ([0, …,K-1]) classification.
#### Model Tuning Parameter
> - criterion -> The function to measure the quality of a split. Supported criteria are “gini” for the Gini impurity and “entropy” for the information gain.
> - max_depth -> The maximum depth of the tree. If None, then nodes are expanded until all leaves are pure or until all leaves contain less than min_samples_split samples.
> - max_leaf_nodes -> Grow a tree with max_leaf_nodes in best-first fashion. Best nodes are defined as relative reduction in impurity. If None then unlimited number of leaf nodes.
> - max_features -> The number of features to consider when looking for the best split: **{auto , sqrt, log2}**
```
model = DecisionTreeClassifier(random_state=123)
model.fit(x_train,y_train)
```
#### Model Accuracy
score() method return the mean accuracy on the given test data and labels.
In multi-label classification, this is the subset accuracy which is a harsh metric since you require for each sample that each label set be correctly predicted.
```
print("Accuracy score {:.2f} %\n".format(model.score(x_test,y_test)*100))
```
#### Confusion Matrix
A confusion matrix is utilized to understand the performance of the classification model or algorithm in machine learning for a given test set where results are known.
Plotting confusion matrix for the predicted values versus actual values.
```
plot_confusion_matrix(model,x_test,y_test,cmap=plt.cm.Blues)
```
#### Classification Report
A Classification report is used to measure the quality of predictions from a classification algorithm. How many predictions are True, how many are False.
* where:
- Precision:- Accuracy of positive predictions.
- Recall:- Fraction of positives that were correctly identified.
- f1-score:- percent of positive predictions were correct
- support:- Support is the number of actual occurrences of the class in the specified dataset.
```
print(classification_report(y_test,model.predict(x_test)))
```
#### Feature Importances.
The Feature importance refers to techniques that assign a score to features based on how useful they are for making the prediction.
```
plt.figure(figsize=(8,6))
n_features = len(X.columns)
plt.barh(range(n_features), model.feature_importances_, align='center')
plt.yticks(np.arange(n_features), X.columns)
plt.xlabel("Feature importance")
plt.ylabel("Feature")
plt.ylim(-1, n_features)
```
#### Tree Plot
Plot a decision tree.The visualization is fit automatically to the size of the axis. Use the figsize or dpi arguments of plt.figure to control the size of the rendering.
```
fig, axes = plt.subplots(nrows = 1,ncols = 1,figsize = (3,3), dpi=400)
cls_target = [str(x) for x in pd.unique(y_train)]
cls_target.sort()
plot_tree(model,feature_names = X.columns, class_names=cls_target,filled = True)
fig.savefig('./tree.png')
```
#### Creator: Anu Rithiga B , Github: [Profile - Iamgrootsh7](https://github.com/iamgrootsh7)
| github_jupyter |
```
!pip install keras-rectified-adam
from keras.datasets import cifar10
from keras.utils import to_categorical
# データの読み込み
(X_train, y_train), (X_test, y_test) = cifar10.load_data()
x_train, x_val = X_train / 255.0, X_test / 255.0
y_train, y_val = to_categorical(y_train), to_categorical(y_test)
from keras.layers import Input, Conv2D, GlobalAveragePooling2D, BatchNormalization
from keras.layers import Activation, Dense
from keras.models import Model
from keras.optimizers import Adam
from keras_radam import RAdam
from keras.preprocessing.image import ImageDataGenerator
import numpy as np
import matplotlib.pyplot as plt
def train(opt):
#cnnの学習
input_ = Input(shape=(32, 32, 3))
c = Conv2D(64, (1, 1), padding="same")(input_)
c = BatchNormalization()(c)
c = Activation("relu")(c)
c = Conv2D(64, (3, 3), padding="same")(c)
c = BatchNormalization()(c)
c = Activation("relu")(c)
c = Conv2D(64, (3, 3), padding="same")(c)
c = BatchNormalization()(c)
c = Activation("relu")(c)
c = Conv2D(64, (3, 3), strides=2)(c)
c = BatchNormalization()(c)
c = Activation("relu")(c)
c = Conv2D(128, (3, 3), padding="same")(c)
c = BatchNormalization()(c)
c = Activation("relu")(c)
c = Conv2D(128, (3, 3), padding="same")(c)
c = BatchNormalization()(c)
c = Activation("relu")(c)
c = Conv2D(128, (3, 3), strides=2)(c)
c = BatchNormalization()(c)
c = Activation("relu")(c)
c = Conv2D(256, (3, 3), padding="same")(c)
c = BatchNormalization()(c)
c = Activation("relu")(c)
c = Conv2D(256, (3, 3), padding="same")(c)
c = BatchNormalization()(c)
c = Activation("relu")(c)
c = Conv2D(256, (3, 3), strides=2)(c)
c = BatchNormalization()(c)
c = Activation("relu")(c)
c = GlobalAveragePooling2D()(c)
c = Dense(10, activation='softmax')(c)
model = Model(input_, c)
model.compile(loss='categorical_crossentropy',
optimizer=opt,
metrics=['accuracy'])
# Data Augmentation
datagen = ImageDataGenerator(rotation_range=20,
width_shift_range=0.2,
height_shift_range=0.2,
horizontal_flip=True)
datagen.fit(x_train)
#cnnの学習
hist = model.fit_generator(datagen.flow(x_train, y_train, batch_size=128),
steps_per_epoch=x_train.shape[0] /128,
validation_data=(x_val, y_val),
epochs=100,
verbose=1)
print("val_acc is ",np.max(hist.history['val_acc']))
return hist.history['val_acc']
acc_Radam = train(RAdam())
acc_adam = train(Adam())
plt.figure(figsize=(9,6))
plt.rcParams["font.size"] = 18
plt.title("CIFAR10 validation accuracy")
plt.plot(acc_Radam,label="RAdam(acc=0.91)")
plt.plot(acc_adam,label="Adam(acc=0.90)")
plt.ylim(0.7,0.95)
plt.legend()
plt.show()
```
| github_jupyter |
```
from itertools import product
import numpy as np
from scipy import optimize
from scipy import interpolate
import sympy as sm
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
plt.style.use('seaborn-whitegrid')
import seaborn as sns
%load_ext autoreload
%autoreload 2
import ASAD
```
# 1. Human capital accumulation
The **parameters** of the model are:
```
rho = 2
beta = 0.96
gamma = 0.1
b = 1
w = 2
Delta = 0.1
```
The **relevant levels of human capital** are:
```
h_vec = np.linspace(0.1,1.5,100)
```
The **basic functions** are:
```
def consumption_utility(c,rho):
""" utility of consumption
Args:
c (float): consumption
rho (float): CRRA parameter
Returns:
(float): utility of consumption
"""
return c**(1-rho)/(1-rho)
def labor_disutility(l,gamma):
""" disutility of labor
Args:
l (int): labor supply
gamma (float): disutility of labor parameter
Returns:
(float): disutility of labor
"""
return gamma*l
def consumption(h,l,w,b):
""" consumption
Args:
h (float): human capital
l (int): labor supply
w (float): wage rate
b (float): unemployment benefits
Returns:
(float): consumption
"""
if l == 1:
return w*h
else:
return b
```
The **value-of-choice functions** are:
```
def v2(l2,h2,b,w,rho,gamma):
""" value-of-choice in period 2
Args:
l2 (int): labor supply
h2 (float): human capital
w (float): wage rate
b (float): unemployment benefits
rho (float): CRRA parameter
gamma (float): disutility of labor parameter
Returns:
(float): value-of-choice in period 2
"""
c2 = consumption(h2,l2,w,b)
return consumption_utility(c2,rho)-labor_disutility(l2,gamma)
def v1(l1,h1,b,w,rho,gamma,Delta,v2_interp,eta=1):
""" value-of-choice in period 1
Args:
l1 (int): labor supply
h1 (float): human capital
w (float): wage rate
b (float): unemployment benefits
rho (float): CRRA parameter
gamma (float): disutility of labor parameter
Delta (float): potential stochastic experience gain
v2_interp (RegularGridInterpolator): interpolator for value-of-choice in period 2
eta (float,optional): scaling of determistic experience gain
Returns:
(float): value-of-choice in period 1
"""
# a. v2 value, if no experience gain
h2_low = h1 + eta*l1 + 0
v2_low = v2_interp([h2_low])[0]
# b. v2 value, if experience gain
h2_high = h1 + eta*l1 + Delta
v2_high = v2_interp([h2_high])[0]
# c. expected v2 value
v2 = 0.5*v2_low + 0.5*v2_high
# d. consumption
c1 = consumption(h1,l1,w,b)
# e. total value
return consumption_utility(c1,rho) - labor_disutility(l1,gamma) + beta*v2
```
A **general solution function** is:
```
def solve(h_vec,obj_func):
""" solve for optimal labor choice
Args:
h_vec (ndarray): human capital
obj_func (callable): objective function
Returns:
l_vec (ndarray): labor supply choices
v_vec (ndarray): implied values-of-choices
"""
# a. grids
v_vec = np.empty(h_vec.size)
l_vec = np.empty(h_vec.size)
# b. solve for each h in grid
for i,h in enumerate(h_vec):
# i. values of choices
v_nowork = obj_func(0,h)
v_work = obj_func(1,h)
# ii. maximum
if v_nowork > v_work:
v_vec[i] = v_nowork
l_vec[i] = 0
else:
v_vec[i] = v_work
l_vec[i] = 1
return l_vec,v_vec
```
A **general plotting funcition** is:
```
def plot(h_vec,l_vec,v_vec,t):
""" plot optimal choices and value function
Args:
h_vec (ndarray): human capital
l_vec (ndarray): labor supply choices
v_vec (ndarray): implied values-of-choices
t (int): period
"""
# a. labor supply function
fig = plt.figure(figsize=(10,4))
ax = fig.add_subplot(1,2,1)
ax.plot(h_vec,l_vec,label='labor supply')
# income
ax.plot(h_vec,w*h_vec,'--',label='wage income')
ax.plot(h_vec,b*np.ones(h_vec.size),'--',label='unemployment benefits')
# working with income loss
I = (l_vec == 1) & (w*h_vec < b)
if np.any(I):
ax.fill_between(h_vec[I],w*h_vec[I],b*np.ones(I.sum()),label='working with income loss')
ax.set_xlabel(f'$h_{t}$')
ax.set_ylabel(f'$l_{t}$')
ax.set_title(f'labor supply function in period {t}')
ax.legend()
# b. value function
ax = fig.add_subplot(1,2,2)
ax.plot(h_vec,v_vec,label='value function')
ax.set_xlabel(f'$h_{t}$')
ax.set_ylabel(f'$v_{t}$')
ax.set_title(f'value function in period {t}')
ax.legend()
```
## Question 1
The solution in the **second period** is:
```
# a. solve
obj_func = lambda l2,h2: v2(l2,h2,b,w,rho,gamma)
l2_vec,v2_vec = solve(h_vec,obj_func)
# b. plot
plot(h_vec,l2_vec,v2_vec,2)
```
## Question 2
The solution in the **first period** is:
```
# a. create interpolator
v2_interp = interpolate.RegularGridInterpolator((h_vec,), v2_vec,
bounds_error=False,fill_value=None)
# b. solve
obj_func = lambda l1,h1: v1(l1,h1,b,w,rho,gamma,Delta,v2_interp)
l1_vec,v1_vec = solve(h_vec,obj_func)
# c. plot
plot(h_vec,l1_vec,v1_vec,1)
```
## Question 3
1. In **period 2**, the worker only works if her potential wage income ($wh_2$) is higher than the unemployment benefits ($b$).
2. In **period 1**, the worker might work even when she looses income in the current period compared to getting unemployment benefits. The explanation is that she accumulates human capital by working which increase her utility in period 2.
To explain this further, consider the following **alternative problem**:
$$
\begin{aligned}
v_{1}(h_{1}) &= \max_{l_{1}} \frac{c_1^{1-\rho}}{1-\rho} - \gamma l_1 + \beta\mathbb{E}_{1}\left[v_2(h_2)\right] \\
\text{s.t.} \\
c_{1}& = \begin{cases}
w h_1 &
\text{if }l_1 = 1 \\
b & \text{if }l_1 = 0
\end{cases} \\
h_2 &= h_1 + \eta l_1 + \begin{cases}
0 & \text{with prob. }0.5\\
\Delta & \text{with prob. }0.5
\end{cases}\\
l_{1} &\in \{0,1\}\\
\end{aligned}
$$
where $\eta$ scales the deterministic experience gain from working. Before we had $\eta = 1$.
If we instead set $\eta = 0$, then the worker will only works in period 1 if $wh_2 > b$ by a margin compensating her for the utility loss of working.
```
# a. solve
obj_func = lambda l1,h1: v1(l1,h1,b,w,rho,gamma,Delta,v2_interp,eta=0)
l1_vec,v1_vec = solve(h_vec,obj_func)
# b. plot
plot(h_vec,l1_vec,v1_vec,1)
```
# 2. AS-AD model
```
par = {}
par['alpha'] = 5.76
par['h'] = 0.5
par['b'] = 0.5
par['phi'] = 0
par['gamma'] = 0.075
par['delta'] = 0.80
par['omega'] = 0.15
par['sigma_x'] = 3.492
par['sigma_c'] = 0.2
```
## Question 1
Construct the **AD-curve:**
```
y = sm.symbols('y_t')
v = sm.symbols('v_t')
alpha = sm.symbols('alpha')
h = sm.symbols('h')
b = sm.symbols('b')
AD = 1/(h*alpha)*(v-(1+b*alpha)*y)
AD
```
Construct the **SRAS-curve:**
```
phi = sm.symbols('phi')
gamma = sm.symbols('gamma')
pilag = sm.symbols('\pi_{t-1}')
ylag = sm.symbols('y_{t-1}')
s = sm.symbols('s_t')
slag = sm.symbols('s_{t-1}')
SRAS = pilag + gamma*y- phi*gamma*ylag + s - phi*slag
SRAS
```
**Find solution:**
```
y_eq = sm.solve(sm.Eq(AD,SRAS),y)
y_eq[0]
pi_eq = AD.subs(y,y_eq[0])
pi_eq
sm.init_printing(pretty_print=False)
```
## Question 2
Create **Python functions**:
```
AD_func = sm.lambdify((y,v,alpha,h,b),AD)
SRAS_func = sm.lambdify((y,s,ylag,pilag,slag,phi,gamma),SRAS)
y_eq_func = sm.lambdify((ylag,pilag,v,s,slag,alpha,h,b,phi,gamma),y_eq[0])
pi_eq_func = sm.lambdify((ylag,pilag,v,s,slag,alpha,h,b,phi,gamma),pi_eq)
```
**Illustrate equilibrium:**
```
# a. lagged values and shocks
y0_lag = 0.0
pi0_lag = 0.0
s0 = 0.0
s0_lag = 0.0
# b. current output
y_vec = np.linspace(-0.2,0.2,100)
# c. figure
fig = plt.figure()
ax = fig.add_subplot(1,1,1)
# SRAS
pi_SRAS = SRAS_func(y_vec,s0,y0_lag,pi0_lag,s0_lag,par['phi'],par['gamma'])
ax.plot(y_vec,pi_SRAS,label='SRAS')
# ADs
for v0 in [0, 0.1]:
pi_AD = AD_func(y_vec,v0,par['alpha'],par['h'],par['b'])
ax.plot(y_vec,pi_AD,label=f'AD ($v_0 = {v0})$')
# equilibrium
eq_y = y_eq_func(y0_lag,pi0_lag,v0,s0,s0_lag,par['alpha'],par['h'],par['b'],par['phi'],par['gamma'])
eq_pi =pi_eq_func(y0_lag,pi0_lag,v0,s0,s0_lag,par['alpha'],par['h'],par['b'],par['phi'],par['gamma'])
ax.scatter(eq_y,eq_pi,color='black',zorder=3)
ax.set_xlabel('$y_t$')
ax.set_ylabel('$\pi_t$')
ax.legend();
```
## Question 3
**Allocate memory and draw random shocks**:
```
def prep_sim(par,T,seed=1986):
""" prepare simulation
Args:
par (dict): model parameters
T (int): number of periods to simulate
seed (int,optional): seed for random numbers
Returns:
sim (dict): container for simulation results
"""
# a. set seed
if not seed == None:
np.random.seed(seed)
# b. allocate memory
sim = {}
sim['y'] = np.zeros(T)
sim['pi'] = np.zeros(T)
sim['v'] = np.zeros(T)
sim['s'] = np.zeros(T)
# c. draw random shocks
sim['x_raw'] = np.random.normal(loc=0,scale=1,size=T)
sim['c_raw'] = np.random.normal(loc=0,scale=1,size=T)
return sim
```
**Simualte** for $T$ periods:
```
def simulate(par,sim,T):
""" run simulation
Args:
par (dict): model parameters
sim (dict): container for simulation results
T (int): number of periods to simulate
"""
for t in range(1,T):
# a. shocks
sim['v'][t] = par['delta']*sim ['v'][t-1] + par['sigma_x']*sim['x_raw'][t]
sim['s'][t] = par['omega']*sim ['s'][t-1] + par['sigma_c']*sim['c_raw'][t]
# b. output
sim['y'][t] = y_eq_func(sim['y'][t-1],sim['pi'][t-1],sim['v'][t],sim['s'][t],sim['s'][t-1],
par['alpha'],par['h'],par['b'],par['phi'],par['gamma'])
# c. inflation
sim['pi'][t] = pi_eq_func(sim['y'][t-1],sim['pi'][t-1],sim['v'][t],sim['s'][t],sim['s'][t-1],
par['alpha'],par['h'],par['b'],par['phi'],par['gamma'])
# a. settings
T = 101
# b. prepare simulation
sim = prep_sim(par,T)
# c. overview shocks
sim['x_raw'][:] = 0
sim['c_raw'][:] = 0
sim ['x_raw'][1] = 0.1/par['sigma_x']
# d. run simulation
simulate(par,sim,T)
```
**$(y_t,\pi_t)$-diagram:**
```
fig = plt.figure()
ax = fig.add_subplot(1,1,1)
ax.plot(sim['y'][1:],sim['pi'][1:],ls='-',marker='o')
ax.set_xlabel('$y_t$')
ax.set_ylabel('$\pi_t$');
for i in range(1,7):
ax.text(sim['y'][i],sim['pi'][i]+0.0002,f't = {i}')
```
**Time paths:**
```
fig = plt.figure()
ax = fig.add_subplot(1,1,1)
ax.plot(np.arange(0,T-1),sim['y'][1:],label='$y_t$, output gap')
ax.plot(np.arange(0,T-1),sim['pi'][1:],label='$\pi_t$, inflation gap')
ax.legend();
```
## Question 4
```
# a. simulate
T = 1000
sim = prep_sim(par,T)
simulate(par,sim,T)
# b. figure
fig = plt.figure()
ax = fig.add_subplot(1,1,1)
ax.plot(np.arange(T),sim['y'],label='$y_t$, output gap')
ax.plot(np.arange(T),sim['pi'],label='$\pi_t$, inflation gap')
ax.legend();
# c. print
def print_sim(sim):
print(f'std. of output gap: {np.std(sim["y"]):.4f}')
print(f'std. of inflation gap: {np.std(sim["pi"]):.4f}')
print(f'correlation of output and inflation gap: {np.corrcoef(sim["y"],sim["pi"])[0,1]:.4f}')
print(f'1st order autocorrelation of output gap: {np.corrcoef(sim["y"][1:],sim["y"][:-1])[0,1]:.4f}')
print(f'1st order autocorrelation of inflation gap: {np.corrcoef(sim["pi"][1:],sim["pi"][:-1])[0,1]:.4f}')
print_sim(sim)
```
### Quesiton 5
**The initial plot:**
```
# a. calculate correlations
K = 100
phis = np.linspace(0.01,0.99,K)
corr_y_pi = np.empty(K)
est = par.copy()
for i,phi in enumerate(phis):
# i. update
est['phi'] = phi
# ii. simulate
simulate(est,sim,T)
# iii. save
corr_y_pi[i] = np.corrcoef(sim["y"],sim["pi"])[0,1]
# b. plot
fig = plt.figure()
ax = fig.add_subplot(1,1,1)
ax.plot(phis,corr_y_pi)
ax.set_xlabel('$\phi$')
ax.set_ylabel('corr($y_t,\pi_t$)');
```
**The optimization:**
```
# a. copy parameters
est = par.copy()
# b. objective funciton
def obj_func(x,est,sim,T,sim_func):
""" calculate objective function for estimation of phi
Args:
x (float): trial value for phi
est (dict): model parameters
sim (dict): container for simulation results
T (int): number of periods to simulate
sim_func (callable): simulation function
Returns:
obj (float): objective value
"""
# i. phi
est['phi'] = x
# ii. simulate
sim_func(est,sim,T)
# iii. calcualte objective
obj = (np.corrcoef(sim["y"],sim["pi"])[0,1]-0.31)**2
return obj
# c. optimize
result = optimize.minimize_scalar(obj_func,args=(est,sim,T,simulate),
bounds=(0+1e-8,1-1e-8),method='bounded')
# d. result
est['phi'] = result.x
print(f'result: phi = {result.x:.3f}')
# e. statistics
print('')
simulate(est,sim,T)
print_sim(sim)
```
### Advanced
**Problem:** The estimate for $\phi$ above depends on the seed chosen for the random number generator. This can be illustrated by re-doing the estimation for different seeds:
```
seeds = [1997,1,17,2018,999] # "randomly chosen seeds"
for seed in seeds:
# a. prepare simulate
sim_alt = prep_sim(par,T,seed)
# b. run optimizer
result = optimize.minimize_scalar(obj_func,args=(est,sim_alt,T,simulate),bounds=(0+1e-8,1-1e-8),method='bounded')
result_alt = optimize.minimize_scalar(obj_func,args=(est,sim_alt,T,ASAD.simulate),bounds=(0+1e-8,1-1e-8),method='bounded')
print(f'seed = {seed:4d}: phi = {result.x:.3f} [{result_alt.x:.3f}]')
```
**Solution:** To reduce this problem, we need to simulate more than 1,000 periods. To do so it is beneficial to use the fast simulation function provided in **ASAD.py** (optimized using numba):
1. The results in the square brackets above show that this simulation function gives the same results.
2. The results below show that when we simulate 1,000,000 periods the estimate of $\phi$ is approximately 0.983-0.984 irrespective of the seed.
```
T_alt = 1_000_000
for seed in [1997,1,17,2018,999]:
# a. simulate
sim_alt = prep_sim(par,T_alt,seed)
# b. run optimizer
result = optimize.minimize_scalar(obj_func,args=(est,sim_alt,T_alt,ASAD.simulate),bounds=(0+1e-8,1-1e-8),method='bounded')
print(f'seed = {seed:4d}: phi = {result.x:.3f}')
```
## Question 6
```
# a. copy parameters
est = par.copy()
# b. objective function
def obj_func_all(x,est,sim,T,sim_func):
""" calculate objective function for estimation of phi, sigma_c and sigma_c
Args:
x (ndarray): trial values for [phi,sigma_x,sigma_c]
est (dict): model parameters
sim (dict): container for simulation results
T (int): number of periods to simulate
sim_func (callable): simulation function
Returns:
obj (float): objective value
"""
# i. phi with penalty
penalty = 0
if x[0] < 1e-8:
phi = 1e-8
penalty += (1-1e-8-x[0])**2
elif x[0] > 1-1e-8:
phi = 1-1e-8
penalty += (1-1e-8-x[0])**2
else:
phi = x[0]
est['phi'] = phi
# ii. standard deviations (forced to be non-negative)
est['sigma_x'] = np.sqrt(x[1]**2)
est['sigma_c'] = np.sqrt(x[2]**2)
# iii. simulate
sim_func(est,sim,T)
# iv. calcualte objective
obj = 0
obj += (np.std(sim['y'])-1.64)**2
obj += (np.std(sim['pi'])-0.21)**2
obj += (np.corrcoef(sim['y'],sim['pi'])[0,1]-0.31)**2
obj += (np.corrcoef(sim['y'][1:],sim['y'][:-1])[0,1]-0.84)**2
obj += (np.corrcoef(sim['pi'][1:],sim['pi'][:-1])[0,1]-0.48)**2
return obj + penalty
# c. optimize
x0 = [0.98,par['sigma_x'],par['sigma_c']]
result = optimize.minimize(obj_func_all,x0,args=(est,sim,T,simulate))
print(result)
# d. update and print estimates
est['phi'] = result.x[0]
est['sigma_x'] = np.sqrt(result.x[1]**2)
est['sigma_c'] = np.sqrt(result.x[2]**2)
est_str = ''
est_str += f'phi = {est["phi"]:.3f}, '
est_str += f'sigma_x = {est["sigma_x"]:.3f}, '
est_str += f'sigma_c = {est["sigma_c"]:.3f}'
print(f'\n{est_str}\n')
# e. statistics
sim = prep_sim(est,T)
simulate(est,sim,T)
print_sim(sim)
```
### Advanced
**Same problem:** Different seeds give different results.
```
for seed in seeds:
# a. prepare simulation
sim_alt = prep_sim(par,T,seed)
# b. run optimizer
est = par.copy()
x0 = [0.98,par['sigma_x'],par['sigma_c']]
result = optimize.minimize(obj_func_all,x0,args=(est,sim_alt,T,simulate))
# c. update and print estimates
est['phi'] = result.x[0]
est['sigma_x'] = np.sqrt(result.x[1]**2)
est['sigma_c'] = np.sqrt(result.x[2]**2)
est_str = ''
est_str += f' phi = {est["phi"]:.3f},'
est_str += f' sigma_x = {est["sigma_x"]:.3f},'
est_str += f' sigma_c = {est["sigma_c"]:.3f}'
print(f'seed = {seed:4d}: {est_str}')
```
**Same solution:** Simulate more periods (and use the faster simulation function).
```
T_alt = 1_000_000
for seed in seeds:
# a. simulate
sim_alt = prep_sim(par,T_alt,seed)
# b. run optimizer
est = par.copy()
x0 = [0.98,par['sigma_x'],par['sigma_c']]
result = optimize.minimize(obj_func_all,x0,args=(est,sim_alt,T_alt,ASAD.simulate))
# c. update and print estimates
est['phi'] = result.x[0]
est['sigma_x'] = np.sqrt(result.x[1]**2)
est['sigma_c'] = np.sqrt(result.x[2]**2)
est_str = ''
est_str += f' phi = {est["phi"]:.3f},'
est_str += f' sigma_x = {est["sigma_x"]:.3f},'
est_str += f' sigma_c = {est["sigma_c"]:.3f}'
print(f'seed = {seed:4d}: {est_str}')
```
# 3. Exchange economy
```
# a. parameters
N = 50_000
mu = np.array([3,2,1])
Sigma = np.array([[0.25, 0, 0], [0, 0.25, 0], [0, 0, 0.25]])
zeta = 1
gamma = 0.8
# b. random draws
seed = 1986
np.random.seed(seed)
# preferences
alphas = np.exp(np.random.multivariate_normal(mu,Sigma,size=N))
betas = alphas/np.reshape(np.sum(alphas,axis=1),(N,1))
# endowments
e1 = np.random.exponential(zeta,size=N)
e2 = np.random.exponential(zeta,size=N)
e3 = np.random.exponential(zeta,size=N)
```
## Question 1
```
# a. calculate
budgetshares = betas
# b. histograms
fig = plt.figure()
ax = fig.add_subplot(1,1,1)
for i in range(3):
ax.hist(budgetshares[:,i],bins=100,alpha=0.7,density=True)
```
## Question 2
```
def excess_demands(budgetshares,p1,p2,e1,e2,e3):
""" calculate excess demands for good 1, 2 and 3
Args:
budgetshares (ndarray): budgetshares for each good for all consumers
p1 (float): price of good 1
p2 (float): price of good 2
e1 (ndrarray): endowments of good 1
e2 (ndrarray): endowments of good 2
e3 (ndrarray): endowments of good 3
Returns:
ed_1 (float): excess demands for good 1
ed_2 (float): excess demands for good 2
ed_3 (float): excess demands for good 3
"""
# a. income
I = p1*e1+p2*e2+1*e3
# b. demands
demand_1 = np.sum(budgetshares[:,0]*I/p1)
demand_2 = np.sum(budgetshares[:,1]*I/p2)
demand_3 = np.sum(budgetshares[:,2]*I) # p3 = 1, numeraire
# b. supply
supply_1 = np.sum(e1)
supply_2 = np.sum(e2)
supply_3 = np.sum(e3)
# c. excess demand
ed_1 = demand_1-supply_1
ed_2 = demand_2-supply_2
ed_3 = demand_3-supply_3
return ed_1,ed_2,ed_3
# a. calculate on grid
K = 50
p1_vec = np.linspace(4,8,K)
p2_vec = np.linspace(0.5,8,K)
p1_mat,p2_mat = np.meshgrid(p1_vec,p2_vec,indexing='ij')
ed1_mat = np.empty((K,K))
ed2_mat = np.empty((K,K))
ed3_mat = np.empty((K,K))
for (i,p1),(j,p2) in product(enumerate(p1_vec),enumerate(p2_vec)):
ed1_mat[i,j],ed2_mat[i,j],ed3_mat[i,j] = excess_demands(budgetshares,p1,p2,e1,e2,e3)
# b. plot
fig = plt.figure()
ax = fig.add_subplot(1,1,1,projection='3d')
ax.plot_wireframe(p1_mat,p2_mat,ed1_mat/N)
ax.plot_surface(p1_mat,p2_mat,np.zeros((K,K)),alpha=0.5,color='black',zorder=99)
ax.set_xlabel('$p_1$')
ax.set_ylabel('$p_2$')
ax.set_title('excess demand for good 1')
ax.invert_xaxis()
fig.tight_layout()
fig = plt.figure()
ax = fig.add_subplot(1,1,1,projection='3d')
ax.plot_wireframe(p1_mat,p2_mat,ed2_mat/N)
ax.plot_surface(p1_mat,p2_mat,np.zeros((K,K)),alpha=0.5,color='black',zorder=99)
ax.set_title('excess demand for good 2')
ax.set_xlabel('$p_1$')
ax.set_ylabel('$p_2$')
ax.invert_xaxis();
fig.tight_layout()
fig = plt.figure()
ax = fig.add_subplot(1,1,1,projection='3d')
ax.plot_wireframe(p1_mat,p2_mat,ed3_mat/N)
ax.plot_surface(p1_mat,p2_mat,np.zeros((K,K)),alpha=0.5,color='black',zorder=99)
ax.set_title('excess demand for good 3')
ax.set_xlabel('$p_1$')
ax.set_ylabel('$p_2$')
ax.invert_xaxis();
fig.tight_layout()
```
## Questions 3
**Function for finding the equilibrium:**
```
def find_equilibrium(budgetshares,p1,p2,e1,e2,e3,kappa=0.5,eps=1e-8,maxiter=5000):
""" find equilibrium prices
Args:
budgetshares (ndarray): budgetshares for each good for all consumers
p1 (float): price of good 1
p2 (float): price of good 2
e1 (ndrarray): endowments of good 1
e2 (ndrarray): endowments of good 2
e3 (ndrarray): endowments of good 3
kappa (float,optional): adjustment aggresivity parameter
eps (float,optional): tolerance for convergence
maxiter (int,optinal): maximum number of iteration
Returns:
p1 (ndarray): equilibrium price for good 1
p2 (ndarray): equilibrium price for good 2
"""
it = 0
while True:
# a. step 1: excess demands
ed_1,ed_2,_ed3 = excess_demands(budgetshares,p1,p2,e1,e2,e3)
# b: step 2: stop?
if (np.abs(ed_1) < eps and np.abs(ed_2) < eps) or it >= maxiter:
print(f'(p1,p2) = [{p1:.4f} {p2:.4f}] -> excess demands = [{ed_1:.4f} {ed_2:.4f}] (iterations: {it})')
break
# c. step 3: update p1 and p2
N = budgetshares.shape[0]
p1 = p1 + kappa*ed_1/N
p2 = p2 + kappa*ed_2/N
# d. step 4: return
it += 1
return p1,p2
```
**Apply algorithm:**
```
# a. guess prices are equal to the average beta
betas_mean = np.mean(betas,axis=0)
p1_guess,p2_guess,_p3_guess = betas_mean/betas_mean[-1]
# b. find equilibrium
p1,p2 = find_equilibrium(budgetshares,p1_guess,p2_guess,e1,e2,e3)
```
**Check excess demands:**
```
assert np.all(np.abs(np.array(excess_demands(budgetshares,p1,p2,e1,e2,e3)) < 1e-6))
```
## Questions 4
```
# a. income
I = p1*e1+p2*e2+1*e3
# b. baseline utility
x = budgetshares*np.reshape(I,(N,1))/np.array([p1,p2,1])
base_utility = np.prod(x**betas,axis=1)
# c. plot utility
fig = plt.figure()
ax = fig.add_subplot(1,1,1)
for gamma_now in [gamma,1.0,1.2]:
utility = base_utility**gamma_now
ax.hist(utility,bins=100,density=True,
label=f'$\gamma = {gamma_now:.2f}$: mean(u) = {np.mean(utility):.3f}, var(u) = {np.var(utility):.3f}')
ax.legend();
```
### Question 5
```
# a. equalize endowments
e1_equal = np.repeat(np.mean(e1),N)
e2_equal = np.repeat(np.mean(e2),N)
e3_equal = np.repeat(np.mean(e3),N)
print(f'e_equal = [{e1_equal[0]:.2f},{e2_equal[0]:.2f},{e3_equal[0]:.2f}]')
# b. find equilibrium
p1_equal,p2_equal = find_equilibrium(budgetshares,p1_guess,p2_guess,e1_equal,e2_equal,e3_equal)
```
**Check excess demands:**
```
assert np.all(np.abs(np.array(excess_demands(budgetshares,p1_equal,p2_equal,e1_equal,e2_equal,e3_equal))< 1e-6))
```
**Plot utility:**
```
# a. income
I_equal = p1_equal*e1_equal+p2_equal*e2_equal+1*e3_equal
# b. baseline utility
x = budgetshares*np.reshape(I_equal,(N,1))/np.array([p1_equal,p2_equal,1])
base_utility_equal = np.prod(x**betas,axis=1)
# c. plot utility
fig = plt.figure()
ax = fig.add_subplot(1,1,1)
for gamma_now in [gamma,1.0,1.2]:
utility = base_utility_equal**gamma_now
ax.hist(utility,bins=100,density=True,
label=f'$\gamma = {gamma_now:.2f}$: mean(u) = {np.mean(utility):.3f}, var(u) = {np.var(utility):.3f}')
ax.legend();
```
**Compare prices with baseline:**
```
print(f'baseline: p1 = {p1:.4f}, p2 = {p2:.4f}')
print(f' equal: p1 = {p1_equal:.4f}, p2 = {p2_equal:.4f}')
```
**Conclusions:** The relative prices of good 1 and good 2 *increase* slightly when endowments are equalized. (This can, however, be shown to disappear when $N \rightarrow \infty$.)
Economic behavior (demand and supply), and therefore equilibrium prices, are independent of $\gamma$, which thus only affects utility.
Irrespective of $\gamma$ we have:
1. Equalization of endowments implies a utility floor of approximately 1 because everyone then get approximately one unit of each good.
2. The variance of utility always decreases when endowments are equalized.
3. The remaining inequality in utility when endowments are equalized must be driven by preferences (see below).
The effect on the mean of utility of equalizing endowments depends on whether $\gamma$ is below, equal to or above one:
1. For $\gamma = 0.8 < 1.0$ the mean utility *increases* ("decreasing returns to scale").
2. For $\gamma = 1.0$ the mean utility is *unchanged* ("constant returns to scale").
3. For $\gamma = 1.2 > 1.0$ the utility mean *decreases* ("increasing returns to scale").
**Additional observation:** When endowments are equalized those with high utility have preferences which differ from the mean.
```
for i in range(3):
sns.jointplot(betas[:,i],base_utility_equal,kind='hex').set_axis_labels(f'$\\beta^j_{i+1}$','utility')
```
| github_jupyter |
# How to use the initial conditions
In this tutorial you learn about the initial conditions and how you can use them to create realistic distributions of infected and immune individuals at the start of a simulation.
## Explanation
Briefly, the initial conditions allow you to shape the distribution of initial infections and immunity in the population. You can
- set the number of initial infections.
- increase the number of infections by some factor to reduce underreporting, for example, due to asymptomatic cases. You can also keep shares between subgroups constant.
- let infections evolve over some periods to have courses of diseases in every stage.
- assume pre-existing immunity in the population.
In scenarios where many individuals have already been infected and the disease has spread across the population for a longer time, courses of the diseases are more heterogenous. Thus, you should start a simulation with some kind of "warm-start". That is what the initial conditions are for.
The ``initial_conditions`` can be passed to the [get_simulate_func](../autoapi/sid/index.rst#sid.get_simulate_func). It is a dictionary with the following keys.
```python
initial_conditions = {
"assort_by": None,
"burn_in_periods": 14,
"growth_rate": 1.3,
"initial_infections": 0.05,
"initial_immunity": None,
"known_cases_multiplier": 1.3
}
```
The entries have the following meaning:
- ``"initial_infections"`` is used to set the initial infections in the population. You can use an integer for the number of infected people, a float between 0 and 1 for the share and a series with values for each person.
- ``ìnitial_immunity`` can be given as an integer or a float identical to ``initial_infections`` to allow for pre-existing immunity in the population. Note that infected individuals are also immune. For a 10% pre-existing immunity with 2% currently infected people, set the key to 0.12. By default, ``initial_immunity`` is ``None`` which means no pre-existing immunity.
- ``"known_cases_multiplier"`` can be used to scale the infections indicated by ``initial_infections``. Normally, the number of initial infections relies on official resources which only cover the known, tested cases instead of the real number of infections. Assuming a asymptotic course of disease in a third of cases, the gap between the known cases and the real number of cases can be substantial.
- ``"assort_by"`` can be a variable name or a list of variable names which are used to form groups via ``.groupby``. While scaling the number of infections with the ``"known_cases_multiplier"``, the relative share of infections between groups stays the same.
- ``"burn_in_periods"`` are the days or periods during which infections are started. The shorter the burn-in period the lesser heterogenous the pattern is.
- ``"growth_rate"`` is the exponential growth rate which governs the occurrence of infections over the burn-in periods. For example, a growth rate of two leads to a duplication of infections for every day of the burn-in period.
## Example
### Preparation
Now, let us visualize the effects of the initial conditions. Note that the following example uses internal functions of sid which are not part of the public API and, thus, are not guaranteed to be stable and should not be used in general.
For the example, we need to generate some inputs which are explained below.
```
import itertools
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
import sid
```
First, we get the epidemiological parameters from sid. Further we need to load parameters related to the immunity level and waning. These are not found in the ``epidemiological_parameters`` as the immunity parameters are not calibrated to the literature yet, but set here as a rule of thumb.
```
params = sid.load_epidemiological_parameters()
immunity_params = pd.read_csv(
"../tutorials/immunity_params.csv", index_col=["category", "subcategory", "name"]
)
params = pd.concat([params, immunity_params])
```
Next, we create artifical individuals which belong to an age group and a region.
```
n_people = 100_000
seed = itertools.count()
available_ages = [
"0-9",
"10-19",
"20-29",
"30-39",
"40-49",
"50-59",
"60-69",
"70-79",
"80-100",
]
ages = np.random.choice(available_ages, size=n_people)
regions = np.random.choice(["North", "East", "South", "West"], size=n_people)
states = pd.DataFrame({"age_group": ages, "region": regions}).astype("category")
virus_strains = {"names": ["base_strain"]}
# Early processing of states and drawing courses of diseases which is necessary for
# the following exploration. Does not need to be used by users in general.
states = sid.simulate._process_initial_states(
states, assort_bys={0: ["age_group", "region"]}, virus_strains=virus_strains
)
states = sid.pathogenesis.draw_course_of_disease(states, params, next(seed))
```
Now, we want to specify the initial conditions and assume that 24% of individuals are infected. Furthermore, initial infections should strongly vary by regions. ``"North"`` and ``"South"`` have twice as many infections as ``"East"`` and ``"West"``. We assume that the actual number of infected individuals is 20% higher and infection number should be preserved between regions. We also require that infections double every day over a period of 14 days.
```
n_infections = 24_000
prob = n_infections / n_people
prob_high = prob * 4 / 3
prob_low = prob * 2 / 3
probabilities = states["region"].replace(
{"North": prob_high, "South": prob_high, "East": prob_low, "West": prob_low}
)
initial_infections = np.random.uniform(0, 1, size=len(probabilities)) <= probabilities
initial_infections = pd.Series(initial_infections)
initial_conditions = {
"initial_infections": initial_infections,
"initial_immunity": None,
"known_cases_multiplier": 1.2,
"assort_by": "region",
"growth_rate": 2,
"burn_in_periods": pd.date_range("2021-06-17", "2021-06-30"),
"virus_shares": {"base_strain": 1.0},
}
```
At last, we apply the function which handles the initial conditions and changes the states.
```
default_virus_strains = sid.parse_model.parse_virus_strains(None, params=params)
parsed_initial_conditions = sid.parse_model.parse_initial_conditions(
initial_conditions,
start_date_simulation=pd.Timestamp("2021-07-01"),
virus_strains=default_virus_strains,
)
states = sid.initial_conditions.sample_initial_distribution_of_infections_and_immunity(
states=states,
params=params,
initial_conditions=parsed_initial_conditions,
seed=seed,
testing_demand_models={},
testing_allocation_models={},
testing_processing_models={},
virus_strains=default_virus_strains,
vaccination_models={},
derived_state_variables={},
)
```
### Analysis
Next, we analyze the impact of the initial conditions. First, let us look at the total number of infections. We started with 24,000 infections and increased the number by 20% which are roughly 28,800 infections.
```
states["ever_infected"].sum()
```
We wanted to preserve the regional differences in positive cases where North and South have twice the infections of East and West.
```
states.groupby("region")["ever_infected"].mean().round(2)
```
Lastly, we wanted infections to increase every day by a factor of 2.
```
fig, ax = plt.subplots(figsize=(6, 4))
infections_by_day = states.query("ever_infected").groupby("cd_ever_infected").size()
infections_by_day.index = -1 * infections_by_day.index
infections_by_day.plot(kind="bar", ax=ax)
ax.set_ylabel("New Infections")
ax.set_xlabel("Days Since Infection")
sns.despine()
plt.show()
```
| github_jupyter |
### Data Frame Plots
documentation: http://pandas.pydata.org/pandas-docs/stable/visualization.html
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('ggplot')
```
The plot method on Series and DataFrame is just a simple wrapper around plt.plot()
If the index consists of dates, it calls gcf().autofmt_xdate() to try to format the x-axis nicely as show in the plot window.
```
ts = pd.Series(np.random.randn(1000), index=pd.date_range('1/1/2000', periods=1000))
ts = ts.cumsum()
ts.plot()
plt.show()
```
On DataFrame, plot() is a convenience to plot all of the columns, and include a legend within the plot.
```
df = pd.DataFrame(np.random.randn(1000, 4), index=pd.date_range('1/1/2016', periods=1000), columns=list('ABCD'))
df = df.cumsum()
plt.figure()
df.plot()
plt.show()
```
You can plot one column versus another using the x and y keywords in plot():
```
df3 = pd.DataFrame(np.random.randn(1000, 2), columns=['B', 'C']).cumsum()
df3['A'] = pd.Series(list(range(len(df))))
df3.plot(x='A', y='B')
plt.show()
df3.tail()
```
### Plots other than line plots
Plotting methods allow for a handful of plot styles other than the default Line plot. These methods can be provided as the kind keyword argument to plot(). These include:
- ‘bar’ or ‘barh’ for bar plots
- ‘hist’ for histogram
- ‘box’ for boxplot
- ‘kde’ or 'density' for density plots
- ‘area’ for area plots
- ‘scatter’ for scatter plots
- ‘hexbin’ for hexagonal bin plots
- ‘pie’ for pie plots
For example, a bar plot can be created the following way:
```
plt.figure()
df.ix[5].plot(kind='bar')
plt.axhline(0, color='k')
plt.show()
df.ix[5]
```
### stack bar chart
```
df2 = pd.DataFrame(np.random.rand(10, 4), columns=['a', 'b', 'c', 'd'])
df2.plot.bar(stacked=True)
plt.show()
```
### horizontal bar chart
```
df2.plot.barh(stacked=True)
plt.show()
```
### box plot
```
df = pd.DataFrame(np.random.rand(10, 5), columns=['A', 'B', 'C', 'D', 'E'])
df.plot.box()
plt.show()
```
### area plot
```
df = pd.DataFrame(np.random.rand(10, 4), columns=['a', 'b', 'c', 'd'])
df.plot.area()
plt.show()
```
### Plotting with Missing Data
Pandas tries to be pragmatic about plotting DataFrames or Series that contain missing data. Missing values are dropped, left out, or filled depending on the plot type.
| Plot Type | NaN Handling | |
|----------------|-------------------------|---|
| Line | Leave gaps at NaNs | |
| Line (stacked) | Fill 0’s | |
| Bar | Fill 0’s | |
| Scatter | Drop NaNs | |
| Histogram | Drop NaNs (column-wise) | |
| Box | Drop NaNs (column-wise) | |
| Area | Fill 0’s | |
| KDE | Drop NaNs (column-wise) | |
| Hexbin | Drop NaNs | |
| Pie | Fill 0’s | |
If any of these defaults are not what you want, or if you want to be explicit about how missing values are handled, consider using fillna() or dropna() before plotting.
### density plot
```
ser = pd.Series(np.random.randn(1000))
ser.plot.kde()
plt.show()
```
### lag plot
Lag plots are used to check if a data set or time series is random. Random data should not exhibit any structure in the lag plot. Non-random structure implies that the underlying data are not random.
```
from pandas.tools.plotting import lag_plot
plt.figure()
data = pd.Series(0.1 * np.random.rand(1000) + 0.9 * np.sin(np.linspace(-99 * np.pi, 99 * np.pi, num=1000)))
lag_plot(data)
plt.show()
```
### matplotlib gallery
documentation: http://matplotlib.org/gallery.html
| github_jupyter |
## Multi-label Classification with scikit-multilearn
### 1. Introduction
We typically group supervised machine learning problems into classification and regression problems. Within the classification problems sometimes, multiclass classification models are encountered where the classification is not binary but we have to assign a class from `n` choices.
In multi-label classification, instead of one target variable $y$, we have multiple target variables $y_1$, $y_2$, ..., $y_n$. For example there can be multiple objects in an image and we need to correctly classify them all or we are attempting predict which combination of a product that a customer would buy.
Certain decision tree based algorithms in [Scikit-Learn](http://scikit-learn.org/stable/modules/multiclass.html) are naturally able to handle multi-label classification. In this post we explore the [scikit-multilearn library](http://scikit.ml/) which leverages Scikit-Learn and is built specifically for multi-label problems.
### 2. Datasets
We use the **MediaMill** [dataset](https://ivi.fnwi.uva.nl/isis/mediamill/challenge/data.php) to explore different multi-label algorithms available in Scikit-Multilearn. Our goal is not to optimize classifier performance but to explore the various algorithms applicable to multi-label classification problems. The dataset is reasonable with over 30k train points and 12k test points. There are 120 features and 101 labels.
This dataset was chosen in order to work with a fairly large dataset to illustrate difficulties in multi-label classification instead of a toy example. In particular when there are $N$ labels, the search space increases exponentially to $2^N$. A list of multi-label datasets can be found at Manik Varma's [Extreme Classification Repository](http://manikvarma.org/downloads/XC/XMLRepository.html). The data is provided in sparse format and the authors only provide Matlab scripts to convert them; some data wrangling is needed in python to handle them.
```
from scipy import sparse
import gc
f = open(r'C:\Users\david\Downloads\Mediamill\Mediamill_data.txt',
'r',encoding='utf-8')
#f = open(r'C:\Users\kaoyuant\Downloads\Mediamill\Mediamill_data.txt',
# 'r',encoding='utf-8')
size = f.readline()
nrows, nfeature,nlabel = [int(s) for s in size.split()]
x_m = [[] for i in range(nrows)]
pos = [[] for i in range(nrows)]
y_m = [[] for i in range(nrows)]
for i in range(nrows):
line = f.readline()
temp=[s for s in line.split(sep=' ')]
pos[i]=[int(s.split(':')[0]) for s in temp[1:]]
x_m[i]=[float(s.split(':')[1]) for s in temp[1:]]
for s in temp[0].split(','):
try:
int(s)
y_m[i]=[ int(s) for s in temp[0].split(',')]
except:
y_m[i]=[]
f = open(r'C:\Users\david\Downloads\Mediamill\Mediamill_trSplit.txt',
'r',encoding='utf-8')
#f = open(r'C:\Users\kaoyuant\Downloads\Mediamill\Mediamill_trSplit.txt',
# 'r',encoding='utf-8')
train=f.readlines()
f = open(r'C:\Users\david\Downloads\Mediamill\Mediamill_tstSplit.txt',
'r',encoding='utf-8')
#f = open(r'C:\Users\kaoyuant\Downloads\Mediamill\Mediamill_tstSplit.txt',
# 'r',encoding='utf-8')
test=f.readlines()
select=0
train_=[int(s.split()[select])-1 for s in train]
test_=[int(s.split()[select])-1 for s in test]
xm_train=[x_m[i] for i in train_]
ym_train=[y_m[i] for i in train_]
xm_test=[x_m[i] for i in test_]
ym_test=[y_m[i] for i in test_]
x_train=sparse.lil_matrix((len(train_),nfeature))
for i in range(len(train_)):
for j in range(len(pos[i])):
x_train[i,pos[i][j]]=xm_train[i][j]
x_test=sparse.lil_matrix((len(test_),nfeature))
for i in range(len(test_)):
for j in range(len(pos[i])):
x_test[i,pos[i][j]]=xm_test[i][j]
del x_m, xm_train, pos,xm_test
gc.collect()
y_train=sparse.lil_matrix((len(train_),nlabel))
for i in range(len(train_)):
for j in ym_train[i]:
y_train[i,j]=1
y_test=sparse.lil_matrix((len(test_),nlabel))
for i in range(len(test_)):
for j in ym_test[i]:
y_test[i,j]=1
del y_m, ym_train, ym_test
gc.collect()
```
### 3. Label Graph
When the label space is large, we can try to explore it using graph methods. Each label is a node in the graph and an edge exists when labels co-occur, weighted by the frequency of co-occurence.
```
from skmultilearn.cluster import LabelCooccurrenceGraphBuilder
graph_builder = LabelCooccurrenceGraphBuilder(weighted=True,
include_self_edges=False)
label_names=[i for i in range(nlabel)]
edge_map = graph_builder.transform(y_train)
print("{} labels, {} edges".format(len(label_names), len(edge_map)))
```
Once we have constructed the label graph, we can apply graph algorithms on it to explore and understand it.
One possibility is to cluster similar labels together so that they are processed together by the multilabel classification algorithms. Community detection methods such as the [Louvain algorithm](https://en.wikipedia.org/wiki/Louvain_Modularity) allow us to cluster the label graph. This is implemented in the `NetworkXLabelGraphClusterer` with the parameter `methods = louvain`.
```
from skmultilearn.cluster import NetworkXLabelGraphClusterer
# we define a helper function for visualization purposes
def to_membership_vector(partition):
return {
member : partition_id
for partition_id, members in enumerate(partition)
for member in members
}
clusterer = NetworkXLabelGraphClusterer(graph_builder, method='louvain')
partition = clusterer.fit_predict(x_train,y_train)
membership_vector = to_membership_vector(partition)
print('There are', len(partition),'clusters')
```
Our graph can be visualized with NetworkX. We use the force or spring layout for better visibility.
```
import networkx as nx
names_dict = dict(enumerate(x for x in label_names))
import matplotlib.pyplot as plt
%matplotlib inline
nx.draw(
clusterer.graph_,
pos=nx.spring_layout(clusterer.graph_,k=4),
labels=names_dict,
with_labels = True,
width = [10*x/y_train.shape[0] for x in clusterer.weights_['weight']],
node_color = [membership_vector[i] for i in range(y_train.shape[1])],
cmap=plt.cm.viridis,
node_size=250,
font_size=10,
font_color='white',
alpha=0.8
)
```
We get 3 clusters. It might make sense to group them together when we send them to the multilabel algorithm. Certain labels have lower centrality, indicating less influence from other labels. It is possible to predict these labels first before predicting the more central labels when using the chained classifier algorithm (see below).
### 4. Metric
Before going into the details of each multilabel classification method, we select a metric to gauge how well the algorithm is performing. Similar to a classification problem it is possible to use `Hamming Loss`, `Accuracy`, `Precision`, `Jaccard Similarity`, `Recall`, and `F1 Score`. These are available from Scikit-Learn.
Going forward we'll chose the `F1 Score` as it averages both `Precision` and `Recall`. We set the parameter `average = micro` to calculate metrics globally. There are many labels and some labels are not predicted; using `average = weighted` will result in the score for certain labels to be set to `0` before averaging.
It is also helpful to plot the confusion matrix to understand how the classifier is performing, but in our case there are too many labels to visualize.
### 5. Multilabel Classifiers - Algorithm Adaptation
Algorithm Adaptation, as indicated by it's name, extend single label classification to the multi-label context, usually by changing the cost or decision functions.
#### 5a. Algorithm Adaptation - MLkNN
Multi-label K Nearest Neighbours uses k-Nearest Neighbors to find nearest examples to a test class and uses Bayesian inference to predict labels. This is a distance based method and works well when there is a relationship between distance and labels. Parameters `k` and `s` need to be determined. This can be done using the `GridSearchCV` function from sklearn.
```
from skmultilearn.adapt import MLkNN
from sklearn.model_selection import GridSearchCV
import time
parameters = {'k': range(1,3), 's': [0.5, 0.7, 1.0]}
score = 'f1_micro'
start=time.time()
classifier = GridSearchCV(MLkNN(), parameters, scoring=score)
classifier.fit(x_train, y_train)
print('training time taken: ',round(time.time()-start,0),'seconds')
print('best parameters :', classifier.best_params_, 'best score: ',
clf.best_score_)
print('best parameters :', classifier.best_params_,
'best score: ',classifier.best_score_)
```
#### 5b. Algorithm Adaptation - BRkNNaClassifier
Short hand for Binary Relevance k-Nearest Neighbours. A k-Nearest Neighbour is trained per label. This requires a lot of compute for a large label space.
```
from skmultilearn.adapt import BRkNNaClassifier
parameters = {'k': range(3,5)}
score = 'f1_micro'
start=time.time()
classifier = GridSearchCV(BRkNNaClassifier(), parameters, scoring=score)
classifier.fit(x_train, y_train)
print('training time taken: ',round(time.time()-start,0),'seconds')
print('best parameters :', classifier.best_params_,
'best score: ',classifier.best_score_)
```
### 6. Multilabel Classifiers - Problem Transformation
#### 6a. Problem Transformation : Binary Relevance
Binary relevance is simple; each target variable ($y_1$, $y_2$,..,$y_n$) is treated independently and we are reduced to $n$ classification problems. `Scikit-Multilearn` implements this for us, saving us the hassle of splitting the dataset and training each of them separately.
This classifier can generalize beyond labels present in the training set. However it is very slow if the label space is large.
```
from skmultilearn.problem_transform import BinaryRelevance
from sklearn.ensemble import RandomForestClassifier
import time
start=time.time()
classifier = BinaryRelevance(
classifier = RandomForestClassifier(),
require_dense = [False, True]
)
classifier.fit(x_train, y_train)
print('training time taken: ',round(time.time()-start,0),'seconds')
start=time.time()
y_hat=classifier.predict(x_test)
print('prediction time taken: ',round(time.time()-start,0),'seconds')
import sklearn.metrics as metrics
br_f1=metrics.f1_score(y_test, y_hat, average='micro')
br_hamm=metrics.hamming_loss(y_test,y_hat)
print('Binary Relevance F1-score:',round(br_f1,3))
print('Binary Relevance Hamming Loss:',round(br_hamm,3))
```
#### 6b. Problem Transformation - Label Powerset
This method transforms the problem into a multiclass classification problem; the target variables ($y_1$, $y_2$,..,$y_n$) are combined and each combination is treated as a unique class. This method will produce many classes.
This transformation reduces the problem to only one classifier but, all possible labels need to be present in the training set.
```
from skmultilearn.problem_transform import LabelPowerset
classifier = LabelPowerset(
classifier = RandomForestClassifier(),
require_dense = [False, True]
)
start=time.time()
classifier.fit(x_train, y_train)
print('training time taken: ',round(time.time()-start,0),'seconds')
start=time.time()
y_hat=classifier.predict(x_test)
print('prediction time taken: ',round(time.time()-start,0),'seconds')
lp_f1=metrics.f1_score(y_test, y_hat, average='micro')
lp_hamm=metrics.hamming_loss(y_test,y_hat)
print('Label Powerset F1-score:',round(lp_f1,3))
print('Label Powerset Hamming Loss:',round(lp_hamm,3))
```
#### 6c. Problem Transformation - Classifier Chains
Classifier chains are akin to binary relevance, however the target variables ($y_1$, $y_2$,.., $y_n$) are not fully independent. The features ($x_1$, $x_2$,.., $x_m$) are initially used to predict $y_1$. Next ($x_1$, $x_2$,.., $x_m$, $y_1$) is used to predict $y_2$. At the $n^{th}$ step, ($x_1$, $x_2$,.., $x_m$, $y_1$,.., $y_{n-1}$) predicts $y_n$. The ordering in which the labels are predicted can be determined by the user and can greatly influence the results.
This classifier takes label dependencies into account and generalizes to label combinations not present in the training data. However the quality of the classifier is heavily dependent on the ordering; there are $n!$ possible orderings and this method is slow if the label space is large.
```
from skmultilearn.problem_transform import ClassifierChain
classifier = ClassifierChain(
classifier = RandomForestClassifier(),
require_dense = [False, True],
order=[i for i in range(nlabel)]
)
start=time.time()
classifier.fit(x_train,y_train)
print('training time taken: ',round(time.time()-start,0),'seconds')
start=time.time()
y_hat=classifier.predict(x_test)
print('prediction time taken: ',round(time.time()-start,0),'seconds')
cc_f1=metrics.f1_score(y_test, y_hat, average='micro')
cc_hamm=metrics.hamming_loss(y_test,y_hat)
print('Classifier Chain F1-score:',round(cc_f1,3))
print('Classifier Chain Hamming Loss:',round(cc_hamm,3))
```
Based on our construction of the label space Graph, we try to order the labels by increasing betweeness centrality; poorly connected labels will first be trained and then used to classify other labels.
```
from operator import itemgetter
sorted_deg = [s[0] for s in sorted(nx.betweenness_centrality(clusterer.graph_).items(),
key=itemgetter(1))]
classifier = ClassifierChain(
classifier = RandomForestClassifier(),
require_dense = [False, True],
order=sorted_deg
)
start=time.time()
classifier.fit(x_train,y_train)
print('training time taken: ',round(time.time()-start,0),'seconds')
start=time.time()
y_hat=classifier.predict(x_test)
print('prediction time taken: ',round(time.time()-start,0),'seconds')
cco_f1=metrics.f1_score(y_test, y_hat, average='micro')
cco_hamm=metrics.hamming_loss(y_test,y_hat)
print('Classifier Chain Ordered F1-score:',round(cco_f1,3))
print('Classifier Chain Ordered Hamming Loss:',round(cco_hamm,3))
```
Classifying by increasing betweeness centrality produces very poor results as seen above. If we were to order the chain by decreasing betweeness centrality we still get poor results.
```
rev_sorted_deg = [s[0] for s in sorted(nx.betweenness_centrality(clusterer.graph_).items(),
key=itemgetter(1),
reverse=True)]
classifier = ClassifierChain(
classifier = RandomForestClassifier(),
require_dense = [False, True],
order=rev_sorted_deg
)
start=time.time()
classifier.fit(x_train,y_train)
print('training time taken: ',round(time.time()-start,0),'seconds')
start=time.time()
y_hat=classifier.predict(x_test)
print('prediction time taken: ',round(time.time()-start,0),'seconds')
ccr_f1=metrics.f1_score(y_test, y_hat, average='micro')
ccr_hamm=metrics.hamming_loss(y_test,y_hat)
print('Classifier Chain Reverse Ordered F1-score:',round(ccr_f1,3))
print('Classifier Chain Reverse Ordered Hamming Loss:',round(ccr_hamm,3))
```
These results illustrate the dependence on the order of the labels. Unfortunately we did not get the improvement hoped for by using the ordering derived from the label graph.
### 7. Multilabel Classifiers - Ensembles of Classifiers
This class uses [ensemble learning](https://en.wikipedia.org/wiki/Ensemble_learning) with the base classifier being a multi-label classifier.
#### 7a. Ensembles of Classifiers - LabelSpacePartitioningClassifier
The label space is partitioned into separate sub label spaces, for example by constructing a label graph and applying a graph clustering/community detection algorithm like we did in section 3. A base multi-label subclassifier is trained on each subspace. The result is predicted with each of the subclassifiers and we take the **sum**
This method adapts the classifiers according to the label space and requires less classifiers than binary relevance. On the downside, the label combination has to be present in the training dataset in order to be predicted and partitioning might prevent correct classification of certain label combinations.
```
from skmultilearn.ensemble import LabelSpacePartitioningClassifier
classifier = LabelSpacePartitioningClassifier(
classifier = BinaryRelevance(
classifier = RandomForestClassifier(),
require_dense = [False, True]
),
clusterer = NetworkXLabelGraphClusterer(graph_builder, method='louvain')
)
start=time.time()
classifier.fit(x_train,y_train)
print('training time taken: ',round(time.time()-start,0),'seconds')
start=time.time()
y_hat=classifier.predict(x_test)
print('prediction time taken: ',round(time.time()-start,0),'seconds')
part_f1=metrics.f1_score(y_test, y_hat, average='micro')
part_hamm=metrics.hamming_loss(y_test,y_hat)
print('Label Space Partitioning Classifier F1-score:',round(part_f1,3))
print('Label Space Partitioning Classifier Hamming Loss:',round(part_hamm,3))
```
#### 7b. Ensembles of Classifiers - Majority Voting Classifier
This is similar to the Label Space Partitioning Classifier but the **majority vote** is used instead.
```
from skmultilearn.ensemble import MajorityVotingClassifier
classifier = LabelSpacePartitioningClassifier(
classifier = BinaryRelevance(
classifier = RandomForestClassifier(),
require_dense = [False, True]
),
clusterer = NetworkXLabelGraphClusterer(graph_builder, method='louvain')
)
start=time.time()
classifier.fit(x_train,y_train)
print('training time taken: ',round(time.time()-start,0),'seconds')
start=time.time()
y_hat=classifier.predict(x_test)
print('prediction time taken: ',round(time.time()-start,0),'seconds')
majo_f1=metrics.f1_score(y_test, y_hat, average='micro')
majo_hamm=metrics.hamming_loss(y_test,y_hat)
print('Majority Voting Classifier F1-score:',round(majo_f1,3))
print('Majority Voting Classifier Hamming Loss:',round(majo_hamm,3))
```
### 8. Conclusion
Multi-label classification methods allow us to classify data sets with more than 1 target variable and is an area of active research. There are various methods which should be used depending on the dataset on hand. A variety of base classifiers can be chosen; Random Forest was used for simplicity and to minimize calculation time.
The **MediaMill** dataset is among the smallest in the [Extreme Classification Repository](http://manikvarma.org/downloads/XC/XMLRepository.html). The methods explored above would be far too slow if applied on datasets with label dimensionality of hundreds of thousands or millions. Datasets with a large number of possible labels require different approaches which will be explored in a separate post.
| github_jupyter |
# Updating JSON Documents
Updated: 2019-10-03
## Document Maintenance
The current ISO SQL JSON standard does not provide any definition for an SQL function to update or delete objects or values within a JSON document. From the ISO perspective, the only way to update a JSON document is to extract it from the database, modify it with an application, and then replace the entire document back into a table. This does not help a DBA or developer easily make quick fixes to an individual document or apply table-wide changes to existing documents.
The Db2 JSON SYSTOOLS functions that were developed as a hidden part of Db2's NoSQL API support (based on the MongoDB wire protocol) were documented in Db2 Version 11.1.2.2 under the "SQL access to JSON documents" section and were also added to the system catalogs at that time. While these functions do not conform to the ISO SQL JSON standard, they do provide some functionality that is currently not available with the new ISO JSON functions with the one of specific interest to us in this notebook being the `JSON_UPDATE` function.
**Note:** To use the SYSTOOLS `JSON_UPDATE` function, you must store the data as BSON in a BLOB column.
### JSON_UPDATE
The `JSON_UPDATE` function is part of the SYSTOOLS schema, so it requires the user or application be granted EXECUTE privilege on the function as well as either explicitly qualify any reference to the function with the SYSTOOLS schema (or add SYSTOOLS to the `CURRENT PATH` special register).
The syntax of the JSON_UPDATE function is:
```sql
JSON_UPDATE(document, '{$set : {field:value}}')
'{$unset: {field:null}}'
```
The arguments are:
* document – BSON document
* action – the action we want to take which consists of:
- operation (`$set` or `$unset`)
- key – The key we are looking for
- value – The value we want to set the field to
There are three possible outcomes from using the `JSON_UPDATE` statement:
* If the field is found, the existing value is replaced with the new one when the `$set` is specified
* If the field is not found, the field:value pair is added to the document when `$set` is specified
* If you use the `$unset` keyword and set the value to `null`, the field is removed from the document
There are some significant differences between the arguments used with `JSON_UPDATE` compared to the ISO SQL JSON functions. The first difference is that the document must be in BSON format. This excludes direct access to any JSON documents that you may have stored as character strings. In addition, if a table column is used as the source, the data type of the column must be BLOB.
The field that defines the JSON object also has a different format. With the new ISO JSON functions, you can specify a path to find the target without the root character used by those JSON functions (I.e. `$.`).
You can always convert your documents to BSON format using the new `JSON_TO_BSON` conversion function (and restore them to JSON afterwards) if you find the `JSON_UPDATE` function to be useful. See the example at the end of this section.
### Adding or Updating a New Key-Value Pair
We will create a BOOK table contains a JSON document with the following information.
**Note:** To represent `null` in a Python dictionary (JSON), the value `None` is used instead.
```
book_info = \
{
"authors": {
"primary" : {"first_name":"Paul", "last_name":"Bird"},
"secondary" : {"first_name":"George","last_name":"Baklarz"}
},
"foreword": {
"primary" : {"first_name":"Thomas","last_name":"Hronis"}
},
"formats": ["Hardcover","Paperback",None,"PDF"]
}
```
### Load Db2 Extensions
The Db2 Jupyter extensions need to be loaded in order to run any of the examples in this notebook. In addition, a `CONNECT` command needs to be issued to connect to the local Db2 database. The default `SAMPLE` database is assumed to exist on the local system. If not, you need to modify the `CONNECT` command to use the appropriate userid, database, and host parameters.
```
%run ../db2.ipynb
%run ../connection.ipynb
```
To simplify updating of JSON values, the table should be defined with a BLOB column.
```
%%sql
DROP TABLE BOOKS;
CREATE TABLE BOOKS(INFO BLOB(2000) INLINE LENGTH 2000);
```
Any inserts into this table would need to ensure that the data is converted to BSON format.
```
%sql INSERT INTO BOOKS VALUES JSON_TO_BSON(:book_info)
```
You can use the `JSON_UPDATE` function with regular JSON (character) data, but you will first need to convert this data to BSON, execute the UPDATE statement, and then convert it back to character JSON.
To add a new field to the record, the `JSON_UPDATE` function needs to specify the target key and the new or replacement value, including the full nesting within the document. Since there is no existing key that matches, the following SQL will add a new field called *publish_date* with the date that the book was made available.
```
%%sql
UPDATE BOOKS
SET INFO = SYSTOOLS.JSON_UPDATE(INFO,
'{ $set: {"publish_date": "2018-12-31"}}');
```
The resulting document now contains the new field.
```
%sql -j SELECT BSON_TO_JSON(INFO) FROM BOOKS;
```
If the publish_date field already existed, then the current value for the key would have been replaced by the new value. In the following example, the `JSON_UPDATE` function would replace the date with the new value.
```
%%sql -j
UPDATE BOOKS
SET INFO = SYSTOOLS.JSON_UPDATE(INFO,
'{ $set: {"publish_date": "2018-11-30"}}');
SELECT BSON_TO_JSON(INFO) FROM BOOKS;
```
To update a column that contained character-based JSON, you would need to add appropriate functions that convert the data to and from BSON in order for the update to work.
```sql
UPDATE BOOKS
SET INFO =
BSON_TO_JSON(
SYSTOOLS.JSON_UPDATE(
JSON_TO_BSON(INFO),
'{ $set: {"publish_date": "2018-12-31"}}'
)
);
```
### Adding or Updating a New Array Value
Adding a new value to an array requires some care. The formats field contains four different ways that a book is available for reading. If we want to add a new format (Audio Book), it would be tempting to use the same syntax that was used for adding a new publish date.
```
%%sql
UPDATE BOOKS
SET INFO =
SYSTOOLS.JSON_UPDATE(INFO,'{ $set: {"formats": "Audio Book"}}');
```
Unfortunately, this ends up wiping out the array and replacing it with just a single value:
```
%sql -j SELECT JSON_QUERY(INFO,'$.formats') FROM BOOKS
```
Using the array specification would seem to be a better approach, but the `JSON_UPDATE` function does not use the ISO SQL JSON path method of referring to an array item. To refer to an element in an array, you must append a dot (`.`) after the array name followed by the array index value. So rather than specifying `formats[0]`, the path would be `formats.0`.
We reset the book in our table and then perform the update. This SQL will replace element zero of the array (Hardcover) with "Audio Book".
```
%%sql
DELETE FROM BOOKS;
INSERT INTO BOOKS VALUES JSON_TO_BSON(:book_info);
UPDATE BOOKS
SET INFO =
SYSTOOLS.JSON_UPDATE(INFO,'{ $set: {"formats.0": "Audio Book"}}');
```
We check the contents of the array to see what the contents are.
```
%sql -j SELECT JSON_QUERY(INFO,'$.formats') FROM BOOKS
```
The only way to insert a new value into the array is to pick an index value that is greater than what the list could possibly be. If we reset the table back to the original value that we started with and then issue the following SQL, the formats field will contain the new value.
```
%%sql -j
DELETE FROM BOOKS;
INSERT INTO BOOKS VALUES JSON_TO_BSON(:book_info);
UPDATE BOOKS
SET INFO =
SYSTOOLS.JSON_UPDATE(INFO,'{ $set: {"formats.999": "Audio Book"}}');
SELECT JSON_QUERY(INFO,'$.formats') FROM BOOKS
```
Note that the new array element will be placed at the end as the specified index is 999 which is greater than the current size of the array, but the new element will have the array index value of 4 (JSON arrays start at index 0!) not the 999 specified in the `JSON_UPDATE` call.
### Removing a Field
To remove a field from a document you must use the following syntax:
```sql
JSON_UPDATE(document, '{$unset: {field:null}}')
```
The field must be set to `null` to remove it from the document and the operation is now `$unset` (not the `$set` we used before). Our modified BOOKS table contains the publish_date which now will be removed.
```
%%sql -j
UPDATE BOOKS
SET INFO =
SYSTOOLS.JSON_UPDATE(INFO,'{ $unset: {"publish_date": null}}');
SELECT BSON_TO_JSON(INFO) FROM BOOKS;
```
It is not actually possible to remove an item from an array, but it is possible to set the specific array value to `null`. Again, you must use the SYSTOOLS functions approach to array specification instead of the JSON SQL path expression we have discussed in previous chapters.
This SQL will set the "Audio Books" array item to null in the list but will not actually remove it. Here we have to specify the specific array index value that we want to remove (which is 4).
```
%%sql -j
UPDATE BOOKS
SET INFO =
SYSTOOLS.JSON_UPDATE(INFO,'{ $unset: {"formats.4": null}}');
SELECT JSON_QUERY(INFO,'$.formats') FROM BOOKS;
```
You can't remove the null value from the array. `JSON_UPDATE` does not remove the actual array entry when a delete occurs in order that the index values for subsequent elements within an array will be preserved. Although, in this example there are no entries after the one affected, `JSON_UPDATE` does not try to be too clever about this, it just does not remove them.
### Updating JSON documents stored as Character Strings
The `JSON_UPDATE` function requires that the document be stored as a BSON object in a BLOB column. If your documents are currently stored as character string, then you will need to add some additional logic around the `UPDATE` statement.
The BOOKS table was recreated in the following format.
```
%%sql
DROP TABLE BOOKS;
CREATE TABLE BOOKS(INFO VARCHAR(2000));
INSERT INTO BOOKS VALUES (:book_info);
```
To add the new publish_date field to the record, we would use the following `UPDATE` statement.
```
%%sql
UPDATE BOOKS
SET INFO =
BSON_TO_JSON(
SYSTOOLS.JSON_UPDATE(JSON_TO_BSON(INFO),
'{ $set: {"formats.999": "Audio Book"}}')
);
```
As of Db2 11.1.4.4, the JSON SYSTOOLS functions are compatible with the BSON storage format used by the ISO SQL JSON functions so that is why the `BSON_TO_JSON` and `JSON_TO_BSON` functions are used rather than the original SYSTOOLS conversion functions.
We check the contents of the book document to make sure our Audio book has been added.
```
%%sql -j
SELECT JSON_QUERY(INFO,'$.formats') FROM BOOKS
```
## Summary
The ISO SQL JSON functions currently do not provide a mechanism for adding, updating, or deleting objects or elements within a JSON document. Without this capability, applications will need to retrieve entire documents, modify them, and then re-store them back into the database.
Db2 includes a JSON SYSTOOLS function called `JSON_UPDATE` that allows for the update of key:value pairs within a JSON document. It has some restrictions on the format that the document must be in and uses a slightly different JSON path expression that the standard uses. However, in situations where simple updates or quick fixes are required, this function may be sufficient. The only drawback is that this function is not part of the ISO SQL standard and may be discontinued at a future date once a replacement is made available.
#### Credits: IBM 2019, George Baklarz [baklarz@ca.ibm.com]
| github_jupyter |
<center><img src="https://ioam.github.io/imagen/_images/patterntypes_small.png"/></center>
The ImaGen package provides comprehensive support for creating resolution-independent two-dimensional pattern distributions. ImaGen consists of a large library of spatial patterns, including mathematical functions, geometric primitives, and images read from files, along with many ways to combine or select from any other patterns. These patterns can be used in any Python program that needs configurable patterns or streams of patterns. Basically, as long as the code can accept a Python callable and will call it each time it needs a new pattern, users can then specify any pattern possible in ImaGen's simple declarative pattern language, and the downstream code need not worry about any of the details about how the pattern is specified or generated. This approach gives users full flexibility about which patterns they wish to use, while relieving the downstream code from having to implement anything about patterns. The detailed examples below should help make this clear.
## Usage
To create a pattern, just ``import imagen``, then instantiate one of ImaGen's ``PatternGenerator`` classes. Each of these classes support various parameters, which are each described in the [Reference Manual](Reference_Manual) or via ``help(``pattern-object-or-class``)``. Any parameter values specified on instantiation become the defaults for that object:
```
import imagen as ig
line=ig.Line(xdensity=5, ydensity=5, smoothing=0)
```
Then whenever the ``line`` object is called, you'll get a new [NumPy](http://numpy.org) array:
```
line()
```
Here the parameters ``xdensity`` and ``ydensity`` specified that a continuous 1.0×1.0 region in (x,y) space should be sampled on a 5×5 grid. The ``line`` object can now be called repeatedly to get new arrays of data, with any parameter values specified to override those declared above:
```
import numpy as np
np.set_printoptions(1)
line(smoothing=0.1,orientation=0.8,thickness=0.4)
```
As you can see, the results are in the form of a Numpy array, and here we have used very small arrays to avoid generating a lot of numeric output. For larger arrays, it is convenient to view them as images, which you can do easily if you have the ``PIL`` or ``pillow`` libraries installed:
```
line.set_param(xdensity=72,ydensity=72,orientation=np.pi/4, thickness=0.1, smoothing=0.02)
line.pil(orientation=3*np.pi/4)
```
As you can see, the `.pil()` method accepts the same options as supported elsewhere, making it simple to create the image that you need to generate. `.pil()` returns a PIL image, which can be exported to files on disk using the methods described in the PIL documentation.
ImaGen depends only on NumPy, Param, and HoloViews, none of which have any other required dependencies, and it is thus easy to incorporate ImaGen into your own code to generate or use patterns freely. That said, HoloViews supports various plotting libraries, including Matplotlib and Bokeh, and if you have one of those installed then imagen provides a convenient way to plot the pattern objects with axes and labels:
```
import holoviews as hv
hv.notebook_extension("matplotlib")
line[:]
```
We will use this plotting interface to show off the remaining patterns, but please remember that the main purpose of ImaGen is to generate arrays for use in other programs, not simply to draw pretty patterns for plotting!
### Dynamic parameter values
As you can see above, ``PatternGenerator`` objects return different patterns depending on their parameter values. An important feature of these parameter values is that any of them can be set to "dynamic" values, which will then result in a different pattern each time (see the [Param package](http://ioam.github.io/param) and its ``numbergen`` module for details). With dynamic parameters, ``PatternGenerators`` provide streams of patterns, not just individual patterns. For example, let's define a ``SineGrating`` object with a random orientation, collect four of them at different times (using the ``.anim()`` method), and lay them out next to each other (using the ``NdLayout`` class from HoloViews):
```
import numbergen as ng
from holoviews import NdLayout
import param
param.Dynamic.time_dependent=True
NdLayout(ig.SineGrating(orientation=np.pi*ng.UniformRandom()).anim(3))
```
As you can see, each time the sine grating was rendered, the pattern differed, because the parameter value for orientation was chosen randomly. Of course, you can set any combination of patterns to dynamic values, to get arbitrarily complex variation over time:
```
%%opts Image (cmap='gray')
sine_disk = ig.SineGrating(orientation=np.pi*ng.UniformRandom(),
scale=0.25*ng.ExponentialDecay(time_constant=3),
frequency=4+7*ng.UniformRandom(),
x=0.3*ng.NormalRandom(seed=1),
y=0.2*ng.UniformRandom(seed=2)-0.1,
mask_shape=ig.Disk(size=0.5,smoothing=0.01))
NdLayout(sine_disk.anim(3))
```
### Composite patterns
As you can see above, ``PatternGenerator`` objects can also be used as a ``mask`` for another ``PatternGenerator``, which is one simple way to combine them.
``PatternGenerator``s can also be combined directly with each other to create ``Composite`` ``PatternGenerator``s, which can make any possible 2D pattern. For instance, we can easily sum 10 oriented ``Gaussian`` patterns, each with random positions and orientations, giving a different overall pattern at each time:
```
gs = ig.Composite(operator=np.add,
generators=[ig.Gaussian(size=0.15,
x=ng.UniformRandom(seed=i+1)-0.5,
y=ng.UniformRandom(seed=i+2)-0.5,
orientation=np.pi*ng.UniformRandom(seed=i+3))
for i in range(10)])
NdLayout(gs.anim(4)).cols(5)
```
Once it has been defined, a ``Composite`` pattern works just like any other pattern, so that it can be placed, rotated, combined with others, etc., allowing you to build up arbitrarily complex objects out of simple primitives. Here we created a ``Composite`` pattern explicitly, but it's usually easier to create them by simply using any of the usual Python operators (``+``, ``-``, ``*``, ``/``, ``**``, ``%``, ``&`` (min), and ``|`` (max)) as in the examples below.
For instance, here's an example using ``np.maximum`` (via the ``|`` operator on ``PatternGenerator``s), rotating the composite pattern together as a unit. We also leave it as a HoloViews animation rather than laying it out over space:
```
%%opts Image.Pattern (cmap='Blues_r')
l1 = ig.Line(orientation=-np.pi/4)
l2 = ig.Line(orientation=+np.pi/4)
cross = l1 | l2
cross.orientation=ng.ScaledTime()*(np.pi/-20)
l1.anim(20) + l2.anim(20) + cross.anim(20)
```
The ``.anim()`` method collects results at different times conveniently. What it's doing repeatedly is getting a copy of each pattern, then running ``param.Dynamic.time_fn.advance(1.0)`` to advance the nominal time, then getting another copy of each pattern until 20 different times have been sampled. The values are "time dependent" (because we set them to be so above), so that any randomness changes only when the time changes, and the randomness is computed as a function of time. That way, regardless of the order you generate the patterns, or even if you go back and forward in time, you will always get the same results at a given nominal time. In your own code, you can turn off time dependence (``param.Dynamic.time_dependent=False``), in which case new parameter values will be generated for every call to the ``PatternGenerator``. Or, if you are working in a domain that has a clear temporal component, such as simulation, you can set ``param.Dynamic.time_fn`` to a function based on your own nominal time, advancing it as appropriate. You can even set that function to real time, in which case you'll get completely unpredictable randomness, which may be appropriate in some circumstances. Whenever there is some notion of time that governs the patterns you want to see, setting ``time_dependent=True`` is a good idea, so that you have precise control over the randomness to ensure reproducible results.
We used one operator above to make the cross image, but we can combine operators in any combination, here to build a cartoon face and add the result to a sweeping ``Line`` pattern masked with a ``Disk``, creating an animated GIF of the results with HoloViews:
```
%opts Image (cmap='gray')
import param
param.Dynamic.time_fn.advance(1)
print("The current nominal time value is %s" % param.Dynamic.time_fn())
%%output backend='matplotlib' holomap='gif'
lefteye = ig.Disk(aspect_ratio=0.7, x=0.04, y=0.10, size=0.08,smoothing=0.005)
leftpupil = ig.Disk(aspect_ratio=1.0, x=0.03, y=0.08, size=0.04,smoothing=0.005)
righteye = ig.Disk(aspect_ratio=0.7, x=0.04, y=-0.1, size=0.08,smoothing=0.005)
rightpupil = ig.Disk(aspect_ratio=1.0, x=0.03, y=-0.08,size=0.04,smoothing=0.005)
nose = ig.Gaussian(aspect_ratio=0.8, x=-0.1, y=0.00, size=0.04)
mouth = ig.Gaussian(aspect_ratio=0.8, x=-0.2, y=0.00, size=0.06)
head = ig.Disk( aspect_ratio=1.5, x=-0.02,y=0.00, size=0.40, scale=0.70,smoothing=0.005)
face=head + lefteye - 1.6*leftpupil + righteye - 1.6*rightpupil - 0.5*nose - 0.8*mouth
face.set_param(x=0.2, y=0.1, offset=0.5, size=0.75)
face.orientation=ng.ScaledTime()*np.pi/20
line = ig.Line(y=0.6-ng.ScaledTime()*0.03)
disk = ig.Disk(smoothing=0.01, size=0.4, x=-0.2, y=-0.2)
(face + line*disk).anim(39)
```
### Image patterns
ImaGen can load and manipulate photographic images just like other patterns, apart from them not being resolution independent. For full functionality this requires the optional PIL or Pillow library, but support for Numpy arrays as images is provided with no further dependencies. For instance, if you have a database of images (here consisting of only one image for simplicity), you can repeatedly select an image at random from the database using ``Selector``, rotate it randomly if desired, and select a random patch of the image at each time:
```
from imagen.image import FileImage
inputs=[FileImage(filename=f, size=6.0,
x=ng.UniformRandom(lbound=-2,ubound=2),
y=ng.UniformRandom(lbound=-2,ubound=2),
orientation=ng.NormalRandom(sigma=0.1*np.pi))
for f in ["images/ellen_arthur.pgm"]]
random_selection=ig.Selector(generators=inputs)
NdLayout(random_selection.anim(5)).cols(6)
```
### Applying functions to generated patterns
Once the pattern has been generated, but before it is returned, you can apply any function to the data that you like, via the ``output_fns`` parameter. A variety of useful ``TransferFn``s are supplied for use as ``output_fns``, such as thresholding functions, normalizing functions (L0, L1, L2, L-infinity, etc.), and convolutions. Any number of these or your own functions (anything that can operate on a 2D Numpy array) can be applied, in order:
```
import imagen.transferfn as tf
from imagen.transferfn.sheet_tf import Convolve
(FileImage()[:] + \
FileImage(output_fns=[tf.BinaryThreshold()])[:] + \
FileImage(output_fns=[Convolve()])[:] + \
FileImage(output_fns=[Convolve(),tf.BinaryThreshold(threshold=0.45)])[:] + \
FileImage(output_fns=[Convolve(kernel_pattern=ig.DifferenceOfGaussians(size=0.12)),
tf.BinaryThreshold()])[:]).cols(5)
```
### Multi-channel patterns
The above examples all show "single-channel" ``PatternGenerator`` objects, which are very general and usable for a huge variety of applications, as they are simply Numpy arrays.
``PatternGenerator`` objects can have any number of channels, with each channel generating a Numpy array of the same size. Multi-channel patterns are used less often, but are particularly useful for generating color images. Color images loaded by the ``FileImage`` pattern will have four channels, one for the monochrome image (as above), and the other three for the red, green, and blue channels (accessed using *object*``.channels()``). RGB images can also be constructed by colorizing a monochrome pattern, or out of combinations of any of the other patterns, using the ``ComposeChannels`` object:
```
from imagen.image import ScaleChannels
ig.ComposeChannels(generators=[ig.Spiral(smoothing=0.02),ig.Spiral(),ig.Spiral(scale=0)])[:] + \
ig.ComposeChannels(generators=[ig.Line(orientation=np.pi/2),ig.Ring(),ig.SquareGrating()])[:]
```
Three- or four-channel patterns support ``.pil()``, generating RGB or RGBA color images.
```
ig.ComposeChannels(generators=[ig.Line(orientation=np.pi/2),ig.Ring(),ig.SquareGrating(),ig.Disk()]).pil()
```
## Pattern types provided
Below are shown examples of each of the pattern types currently provided, using their default parameter values. Very many different parameter values can be chosen, to produce a much wider range of patterns, and of course new patterns can be created as ``Composite`` patterns as shown above.
```
%opts Layout [sublabel_format="" horizontal_spacing=.1 vertical_spacing=.1]
%opts Image (cmap='gray') [xaxis=None yaxis=None show_frame=True] {+axiswise}
from imagen import *
from imagen.random import *
from imagen.image import *
pattern_classes = [x for _, x in sorted(locals().items()) if isinstance(x,type)
and issubclass(x,PatternGenerator) and not x.abstract]
all_patterns = [x()[:] for x in pattern_classes[10:]]
hv.Layout(all_patterns).cols(5)
```
## Extending ImaGen
New ``Composite`` patterns can be created easily without writing new classes, as shown above. If you want to create a new non-composite type, you can simply define a new class inheriting from ``PatternGenerator``, then override ``self.function()`` to draw the pattern, and declare any new parameter(s) used by ``self.function()``. The new pattern can then be rotated, scaled, translated, etc. automatically, with no further coding, it will support dynamic pattern streams automatically, and it can be combined with any existing or new pattern to make new ``Composite`` patterns. If you don't want the automatic scaling, rotating, etc. (e.g. for a whole-field pattern like a new type of random distribution), you can override ``self.__call__`` instead of ``self.function()``, which allows you to do anything that returns a Numpy array of the requested size. See the many classes in ``imagen/__init__.py`` and ``imagen/random.py`` for examples of each approach.
| github_jupyter |
# NEU Surface Defect Dataset
The NEU Surface Defect Database was developed by K. Song and Y. Yan. [The original link appears to be broken.](http://faculty.neu.edu.cn/yunhyan/NEU_surface_defect_database.html) Fortunately, we have a copy of it stored on Box. The original paper describing the dataset and some methods for performing classification is [here](https://doi.org/10.1016/j.apsusc.2013.09.002).
K. Song and Y. Yan, “A noise robust method based on completed local binary patterns for hot-rolled steel strip surface defects,” Applied Surface Science, vol. 285, pp. 858-864, Nov. 2013.
### Dataset Details
The original dataset contains 1800 images of 6 types of surface defects in hot-rolled steel strip. The images we will be working with are RGB bitmap images with the class of the defect noted as a two-character code in the filename. The codes and defect types are noted below:
- Cr: Crazing
- In: Inclusion
- Pa: Patches
- PS: Pitted Surface
- RS: Rolled-in Scale
- Sc: Scratch
### Exercise Goals
1. Prepare dataset for analysis
2. Show example images and classes
3. Save images and class labels in a convenient manner for future use.
___
## 1. Prepare Dataset for Analysis
In this section we'll cover some basic IO for reading the data, exploring the dataset, and making convenient data structures for further analysis.
```
# let's quickly verify that our kernel is running
import sys
print(sys.version)
print("Hello Jupyter")
!pip install matplotlib
# import libraries that we'll use
import os
import glob
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import PIL
from sklearn.utils import shuffle
# Jupyter magic command
%matplotlib inline
# Let's make a helper function to help us find data files
def get_files(file_directory, extension='*.bmp'):
"""
Arguments:
file_directory: path to directory to search for files
extension: desired file type, default *.bmp (bitmap)
Return:
files: list of files in file_directory with extension
"""
files = glob.glob(os.path.join(file_directory, extension))
return files
os.path.join?
glob.glob?
# path to data
DATA_PATH = os.path.join('..', 'data', 'NEU-CLS')
print(DATA_PATH)
```
Let's go ahead and make a list of all of our images. As we've discussed, this dataset has 1800 images so we expect this list to have length of 1800.
```
image_paths = get_files(file_directory=DATA_PATH, extension='*.bmp')
print(len(image_paths))
# let's see what the first couple of images are called
print(image_paths[0:2])
```
We can see that each image path is indeed a path to a particular image. Let's open an image to see how it looks.
```
print(image_paths[0])
img = PIL.Image.open(image_paths[0])
img
# and we can check the size of the image
print(img.size)
type(img)
```
### Examine data structure
It looks like this dataset is 1800 bitmap images of size (200, 200). We can see the class label in the filename, but ideally we'd like to structure our data such that we have something like
```
x: data
y: labels
```
Let's make our list of labels, y
```
image_path = image_paths[0]
print(image_path)
_, filename = os.path.split(image_path)
print(_)
print(filename)
# All of the images are labeled with the pattern YY_#.bmp
# We can use this to our advantage to parse the label from the filename
# let's look at just one image for now
image_path = image_paths[1]
_, filename = os.path.split(image_path)
print(filename)
```
Great! It looks pretty simple to pull the filename from the image. And there are a couple of ways that we can extract the label from here.
1. We can take the first two characters, or
2. We can split the string using the underscore as our marker
```
type(filename)
filename
filename.split?
print(filename[:2])
print(filename.split('_'))
print(filename.split('_')[0]) # index 0: get the first entry from this list
```
---
**Discuss**
Why would we want to use one method over the other? Does it really matter in this case?
---
Now that we have a good strategy to extract labels, let's store these as a convenient variable, `y`. Again, there is more than one way to go about this. I'll show you a couple of ways.
1. For Loop (perhaps the more intuitive way)
2. List comprehension (a cool thing we can do in Python)
```
# For Loop Example (5 lines)
y = [] # make an empty list
type(y)
for image_path in image_paths:
_, filename = os.path.split(image_path) # note the underscore is a commonly used dummy variable
label, _ = filename.split('_') # and sometimes it can make code a bit more readable
y.append(label)
print(len(y))
print(y[:10])
```
Simple enough. Now let's start over. We'll delete the variable y and try again.
```
del y
# List comprehension example (1 line)
# Performs same task as for loop example above
y = [os.path.split(image_path)[1].split('_')[0] for image_path in image_paths]
print(len(y))
print(y[:10])
```
List comprehensions are great for consolidating code, but they can quickly become hard to read if you squish too many steps into the one line. That being said, I'm a big fan of list comprehensions. Just be sure to describe what they do.
### Make a DataFrame with Pandas
It's worth noting that this dataset is small and can easily be read into memory even on my laptop. But as we move forward we may not be so lucky. Let's try one method that will make it easy for us to manage our data: Pandas!
```
# we've already import pandas as pd
df = pd.DataFrame(data=list(zip(image_paths, y)),
columns=['image_path', 'label'])
# we can check the first few lines our dataframe
df.head()
df.shape
# We can even save our dataframe so that we don't have to do all of this in the future
DF_PATH = os.path.join('..', 'data', 'NEU_dataframe.pkl')
df.to_pickle(DF_PATH)
del df
df
# loading the dataframe is simple too
df = pd.read_pickle(DF_PATH)
df.head()
```
And that's it! That's pretty much all we need to do to structure our data for further exploration! Without all of the extra fluff that I've written to type everything out, it really is as easy as
```python
image_paths = get_files(DATA_PATH)
y = [os.path.split(image_path)[1].split('_')[0] for image_path in image_paths]
df = pd.DataFrame(data=list(zip(image_paths, y)),
columns=['image_path', 'label'])
```
## 2. Visualize the Data
To this point our focus has been on the filenames and the labels of the images, but now we're going to look a bit more at the images directly. This is an important step to understand for ourselves what types of things we might want to consider in future decisions.
To do this, let's do a few things:
1. Shuffle the data in the dataframe
2. Use Pandas to group the data by the various classes we have
3. Randomly sample from each group
4. Show a handful of images from each class
```
# 1. Shuffle the data in the dataframe
df = shuffle(df)
# 2. Use Pandas to group the data by label
subset = df.groupby('label', as_index=False)
# 2. Let's quickly double-check the number of groups in this subset (we expect 6)
n_groups = len(subset)
print(n_groups)
df.loc?
# 3. Randomly sample from each group
# We'll use a cool trick called a lambda function
sample_count = 10 # pick 10 samples
replace = True # sample with replacement
L = lambda x: x.loc[np.random.choice(x.index, sample_count, replace),:]
subset = subset.apply(L)
# 4. Show a handful of images from each class
# 6 groups (rows)
# 10 samples per group (columns)
fig, axes = plt.subplots(ncols=sample_count, nrows=n_groups, figsize=(10,10))
for ax, image_path, label in zip(axes.flatten(), subset.image_path, subset.label):
image = PIL.Image.open(image_path)
ax.imshow(image, cmap='gray')
ax.set_title(label)
ax.axis('off')
```
# 3. Save images and class labels in a convenient manner for future use.
As it turns out, we've already done this when we saved our dataframe! You can load the dataframe in a notebook or a script. This dataframe gives you a path to a file for you to read as well as the corresponding class. We'll see in future exercises how there are other ways to accomplish this goal. And of course, you *could* read all of the images into memory and store as a numpy array and store it, but that doesn't really work as your datasets get bigger.
Remember that we shuffled the dataframe **after** we saved it. So you'll want to shuffle it after reloading it and before training your classifier.
Loading the dataframe:
```python
import os
import pandas as pd
DF_PATH = os.path.join('..', 'data', 'NEU_dataframe.pkl')
df = pd.read_pickle(DF_PATH)
```
Remember that we shuffled the dataframe **after** we saved it. So you'll want to shuffle it after reloading it and before training your classifier.
---
# Summary
That's it! Now we can study each of the classes in this dataset and think about some strategies to pursue. Of course, in your exercise, you'll be asked to perform a classification task on this dataset. We've seen for ourselves that this dataset has 6 groups. We can see that the images vary widely in grayscale intensity, so we might want to consider ways of managing brightness as a possible feature; maybe it's unwanted or maybe we want it. There also seems to be some rotations within classes, especially scratches and inclusions.
| github_jupyter |
```
import matplotlib.pyplot as plt
import numpy as np
import cv2
doc_image = cv2.imread('doc1.jpg')
plt.imshow(doc_image)
plt.show()
min_w = 200
height, width, _ = doc_image.shape
scale = min(10., width * 1. / min_w)
height_rescale = int(height * 1. / scale)
width_rescale = int(width * 1. / scale)
height_rescale, width_rescale
doc_copy = np.copy(doc_image)
doc_copy = cv2.resize(doc_copy, (width_rescale, height_rescale))
plt.imshow(doc_copy)
plt.show()
gray = cv2.cvtColor(doc_copy, cv2.COLOR_BGR2GRAY)
high_threshold = cv2.threshold(gray, 0, 255, cv2.THRESH_BINARY | cv2.THRESH_OTSU)[0]
low_threshold = high_threshold * 0.5
canny = cv2.Canny(gray, low_threshold, high_threshold*1.7)
plt.imshow(canny)
plt.show()
lines = cv2.HoughLinesP(canny, 1, np.pi / 180, width_rescale // 3, None, width_rescale / 3, 20)
gray_rgb = cv2.cvtColor(canny, cv2.COLOR_GRAY2RGB)
class Line:
def __init__(self, l):
self.point = l
x1, y1, x2, y2 = l
self.c_x = (x1 + x2) / 2
self.c_y = (y1 + y2) / 2
horizontal, vertical = [], []
for line in lines[0]:
x1, y1, x2, y2 = line
if abs(x1 - x2) > abs(y1 - y2):
horizontal.append(Line(line))
else:
vertical.append(Line(line))
cv2.line(gray_rgb, (x1, y1), (x2, y2), (0, 0, 255), 1)
plt.imshow(gray_rgb)
plt.show()
if len(horizontal) < 2:
if not horizontal or horizontal[0].c_y > height_rescale / 2:
horizontal.append(Line((0, 0, width_rescale - 1, 0)))
if not horizontal or horizontal[0].c_y <= height_rescale / 2:
horizontal.append(Line((0, height_rescale - 1, width_rescale - 1, height_rescale - 1)))
if len(vertical) < 2:
if not vertical or vertical[0].c_x > width_rescale / 2:
vertical.append(Line((0, 0, 0, height_rescale - 1)))
if not vertical or vertical[0].c_x <= width_rescale / 2:
vertical.append(Line((width_rescale - 1, 0, width_rescale - 1, height_rescale - 1)))
horizontal.sort(key=lambda l: l.c_y)
vertical.sort(key=lambda l: l.c_x)
def intersection(l1, l2):
x1, y1, x2, y2 = l1.point
x3, y3, x4, y4 = l2.point
a1, b1 = y2 - y1, x1 - x2
c1 = a1 * x1 + b1 * y1
a2, b2 = y4 - y3, x3 - x4
c2 = a2 * x3 + b2 * y3
det = a1 * b2 - a2 * b1
assert det, 'lines are parallel'
return (1. * (b2 * c1 - b1 * c2) / det, 1. * (a1 * c2 - a2 * c1) / det)
for l in [horizontal[0], vertical[0], horizontal[-1], vertical[-1]]:
x1, y1, x2, y2 = l.point
cv2.line(gray_rgb, (x1, y1), (x2, y2), (0, 255, 255), 1)
plt.imshow(gray_rgb)
plt.show()
img_pts = [intersection(horizontal[0], vertical[0]), intersection(horizontal[0], vertical[-1]),
intersection(horizontal[-1], vertical[0]), intersection(horizontal[-1], vertical[-1])]
for i, p in enumerate(img_pts):
x, y = p
img_pts[i] = (x * scale, y * scale)
cv2.circle(gray_rgb, (int(x), int(y)), 1, (255, 255, 0), 3)
w_a4, h_a4 = 1654, 2339
dst_pts = np.array(((0, 0), (w_a4 - 1, 0), (0, h_a4 - 1), (w_a4 - 1, h_a4 - 1)), np.float32)
img_pts = np.array(img_pts, np.float32)
transmtx = cv2.getPerspectiveTransform(img_pts, dst_pts)
wrapped_img = cv2.warpPerspective(doc_image, transmtx, (w_a4, h_a4))
plt.imshow(wrapped_img)
plt.show()
```
| github_jupyter |
# Kmeans over a set of GeoTiffs
This notebook loads a set of GeoTiffs into a **RDD** of Tiles, with each Tile being a band in the GeoTiff. Each GeoTiff file contains **SpringIndex-** or **LastFreeze-** value for one year over the entire USA.
Kmeans takes years as dimensions. Hence, the matrix has cells as rows and the years as columns. To cluster on all years, the matrix needs to be transposed. The notebook has two flavors of matrix transpose, locally by the Spark-driver or distributed using the Spark-workers. Once transposed the matrix is converted to a **RDD** of dense vectors to be used by **Kmeans** algorithm from **Spark-MLlib**. The end result is a grid where each cell has a cluster ID which is then saved into a SingleBand GeoTiff. By saving the result into a GeoTiff, the reader can plot it using a Python notebook as the one defined in the [python examples](../examples/python).
<span style="color:red">In this notebook the reader only needs to modify the variables in **Mode of Operation Setup**</span>.
## Dependencies
```
import sys.process._
import java.io.{ByteArrayInputStream, ByteArrayOutputStream, ObjectInputStream, ObjectOutputStream}
import geotrellis.proj4.CRS
import geotrellis.raster.{CellType, ArrayTile, DoubleArrayTile, Tile, UByteCellType}
import geotrellis.raster.io.geotiff._
import geotrellis.raster.io.geotiff.writer.GeoTiffWriter
import geotrellis.raster.io.geotiff.{GeoTiff, SinglebandGeoTiff}
import geotrellis.spark.io.hadoop._
import org.apache.hadoop.io._
import geotrellis.vector.{Extent, ProjectedExtent}
import org.apache.spark.broadcast.Broadcast
import org.apache.spark.mllib.clustering.{KMeans, KMeansModel}
import org.apache.spark.mllib.linalg.distributed.{CoordinateMatrix, MatrixEntry, RowMatrix}
import org.apache.spark.mllib.linalg.{Vector, Vectors}
import org.apache.spark.rdd.RDD
import org.apache.spark.{SparkConf, SparkContext}
import org.apache.hadoop.io.{IOUtils, SequenceFile}
import org.apache.hadoop.io.SequenceFile.Writer
//Spire is a numeric library for Scala which is intended to be generic, fast, and precise.
import spire.syntax.cfor._
```
## Mode of operation
Here the user can define the mode of operation.
* **rdd_offline_mode**: If false it means the notebook will create all data from scratch and store grid0, grid0_index, protected_extent and num_cols_rows (from grid0) into HDFS. Otherwise, these data structures are read from HDFS.
* **matrix_offline_mode**: If false it means the notebook will create a mtrix, transposed it and save it to HDFS. Otherwise, these data structures are read from HDFS.
* **kmeans_offline_mode**: If false it means the notebook will train kmeans and run kemans and store kmeans model into HDFS. Otherwise, these data structures are read from HDFS.
It is also possible to define which directory of GeoTiffs is to be used and on which **band** to run Kmeans. The options are
* **BloomFinal** or **LeafFinal** which are multi-band (**4 bands**)
* **DamageIndex** and **LastFreeze** which are single-band and if set band_num higher, it will reset to 0
For kmeans the user can define the **number of iterations** and **number of clusters** as an inclusive range. Such range is defined using **minClusters**, **maxClusters**, and **stepClusters**. These variables will set a loop starting at **minClusters** and stopping at **maxClusters** (inclusive), iterating **stepClusters** at the time. <span style="color:red">Note that when using a range **kemans offline mode** is not possible and it will be reset to **online mode**</span>.
### Mode of Operation setup
<a id='mode_of_operation_setup'></a>
```
//Operation mode
var rdd_offline_mode = true
var matrix_offline_mode = true
var kmeans_offline_mode = true
//GeoTiffs to be read from "hdfs:///user/hadoop/spring-index/"
var dir_path = "hdfs:///user/hadoop/spring-index/"
var offline_dir_path = "hdfs:///user/pheno/spring-index/"
var geoTiff_dir = "LeafFinal"
var geoTiff_2_dir = "BloomFinal"
var band_num = 3
//Years between (inclusive) 1980 - 2015
var model_first_year = 1989
var model_last_year = 2014
//Mask
val toBeMasked = true
val mask_path = "hdfs:///user/hadoop/usa_mask.tif"
//Kmeans number of iterations and clusters
var numIterations = 75
var minClusters = 210
var maxClusters = 500
var stepClusters = 10
var save_kmeans_model = false
```
<span style="color:red">DON'T MODIFY ANY PIECE OF CODE FROM HERE ON!!!</span>.
### Mode of operation validation
```
//Validation, do not modify these lines.
var single_band = false
if (geoTiff_dir == "BloomFinal" || geoTiff_dir == "LeafFinal") {
single_band = false
} else if (geoTiff_dir == "LastFreeze" || geoTiff_dir == "DamageIndex") {
single_band = true
if (band_num > 0) {
println("Since LastFreezze and DamageIndex are single band, we will use band 0!!!")
band_num = 0
}
} else {
println("Directory unknown, please set either BloomFinal, LeafFinal, LastFreeze or DamageIndex!!!")
}
if (minClusters > maxClusters) {
maxClusters = minClusters
stepClusters = 1
}
if (stepClusters < 1) {
stepClusters = 1
}
//Paths to store data structures for Offline runs
var mask_str = ""
if (toBeMasked)
mask_str = "_mask"
var grid0_path = offline_dir_path + geoTiff_dir + "/grid0" + mask_str
var grid0_index_path = offline_dir_path + geoTiff_dir + "/grid0_index" + mask_str
var grids_noNaN_path = offline_dir_path + geoTiff_dir + "/grids_noNaN" + mask_str
var metadata_path = offline_dir_path + geoTiff_dir + "/metadata" + mask_str
var grids_matrix_path = offline_dir_path + geoTiff_dir + "/grids_matrix" + mask_str
//Check offline modes
var conf = sc.hadoopConfiguration
var fs = org.apache.hadoop.fs.FileSystem.get(conf)
val rdd_offline_exists = fs.exists(new org.apache.hadoop.fs.Path(grid0_path))
val matrix_offline_exists = fs.exists(new org.apache.hadoop.fs.Path(grids_matrix_path))
if (rdd_offline_mode != rdd_offline_exists) {
println("\"Load GeoTiffs\" offline mode is not set properly, i.e., either it was set to false and the required file does not exist or vice-versa. We will reset it to " + rdd_offline_exists.toString())
rdd_offline_mode = rdd_offline_exists
}
if (matrix_offline_mode != matrix_offline_exists) {
println("\"Matrix\" offline mode is not set properly, i.e., either it was set to false and the required file does not exist or vice-versa. We will reset it to " + matrix_offline_exists.toString())
matrix_offline_mode = matrix_offline_exists
}
if (!fs.exists(new org.apache.hadoop.fs.Path(mask_path))) {
println("The mask path: " + mask_path + " is invalid!!!")
}
//Years
val model_years = 1980 to 2015
if (!model_years.contains(model_first_year) || !(model_years.contains(model_last_year))) {
println("Invalid range of years for " + geoTiff_dir + ". I should be between " + model_first_year + " and " + model_last_year)
System.exit(0)
}
var model_years_range = (model_years.indexOf(model_first_year), model_years.indexOf(model_last_year))
var num_kmeans :Int = 1
if (minClusters != maxClusters) {
num_kmeans = ((maxClusters - minClusters) / stepClusters) + 1
}
println(num_kmeans)
var kmeans_model_paths :Array[String] = Array.fill[String](num_kmeans)("")
var wssse_path :String = offline_dir_path + geoTiff_dir + "/" + numIterations +"_wssse"
var geotiff_hdfs_paths :Array[String] = Array.fill[String](num_kmeans)("")
var geotiff_tmp_paths :Array[String] = Array.fill[String](num_kmeans)("")
if (num_kmeans > 1) {
var numClusters_id = 0
cfor(minClusters)(_ <= maxClusters, _ + stepClusters) { numClusters =>
kmeans_model_paths(numClusters_id) = offline_dir_path + geoTiff_dir + "/kmeans_model_" + numClusters + "_" + numIterations
//Check if the file exists
val kmeans_exist = fs.exists(new org.apache.hadoop.fs.Path(kmeans_model_paths(numClusters_id)))
if (kmeans_exist && !kmeans_offline_mode) {
println("The kmeans model path " + kmeans_model_paths(numClusters_id) + " exists, please remove it.")
} else if (!kmeans_exist && kmeans_offline_mode) {
kmeans_offline_mode = false
}
geotiff_hdfs_paths(numClusters_id) = offline_dir_path + geoTiff_dir + "/clusters_" + numClusters + "_" + numIterations + ".tif"
geotiff_tmp_paths(numClusters_id) = "/tmp/clusters_" + geoTiff_dir + "_" + numClusters + "_" + numIterations + ".tif"
if (fs.exists(new org.apache.hadoop.fs.Path(geotiff_hdfs_paths(numClusters_id)))) {
println("There is already a GeoTiff with the path: " + geotiff_hdfs_paths(numClusters_id) + ". Please make either a copy or move it to another location, otherwise, it will be over-written.")
}
numClusters_id += 1
}
kmeans_offline_mode = false
} else {
kmeans_model_paths(0) = offline_dir_path + geoTiff_dir + "/kmeans_model_" + minClusters + "_" + numIterations
val kmeans_offline_exists = fs.exists(new org.apache.hadoop.fs.Path(kmeans_model_paths(0)))
if (kmeans_offline_mode != kmeans_offline_exists) {
println("\"Kmeans\" offline mode is not set properly, i.e., either it was set to false and the required file does not exist or vice-versa. We will reset it to " + kmeans_offline_exists.toString())
kmeans_offline_mode = kmeans_offline_exists
}
geotiff_hdfs_paths(0) = offline_dir_path + geoTiff_dir + "/clusters_" + minClusters + "_" + numIterations + ".tif"
geotiff_tmp_paths(0) = "/tmp/clusters_" + geoTiff_dir + "_" + minClusters + "_" + numIterations + ".tif"
if (fs.exists(new org.apache.hadoop.fs.Path(geotiff_hdfs_paths(0)))) {
println("There is already a GeoTiff with the path: " + geotiff_hdfs_paths(0) + ". Please make either a copy or move it to another location, otherwise, it will be over-written.")
}
}
```
## Functions to (de)serialize any structure into Array[Byte]
```
def serialize(value: Any): Array[Byte] = {
val out_stream: ByteArrayOutputStream = new ByteArrayOutputStream()
val obj_out_stream = new ObjectOutputStream(out_stream)
obj_out_stream.writeObject(value)
obj_out_stream.close
out_stream.toByteArray
}
def deserialize(bytes: Array[Byte]): Any = {
val obj_in_stream = new ObjectInputStream(new ByteArrayInputStream(bytes))
val value = obj_in_stream.readObject
obj_in_stream.close
value
}
```
## Load GeoTiffs
Using GeoTrellis all GeoTiffs of a directory will be loaded into a RDD. Using the RDD, we extract a grid from the first file to lated store the Kmeans cluster_IDS, we build an Index for populate such grid and we filter out here all NaN values.
```
val t0 = System.nanoTime()
//Global variables
var projected_extent = new ProjectedExtent(new Extent(0,0,0,0), CRS.fromName("EPSG:3857"))
var grid0: RDD[(Long, Double)] = sc.emptyRDD
var grid0_index: RDD[Long] = sc.emptyRDD
var grids_noNaN_RDD: RDD[Array[Double]] = sc.emptyRDD
var num_cols_rows :(Int, Int) = (0, 0)
var cellT :CellType = UByteCellType
var grids_RDD :RDD[Array[Double]] = sc.emptyRDD
var mask_tile0 :Tile = new SinglebandGeoTiff(geotrellis.raster.ArrayTile.empty(cellT, num_cols_rows._1, num_cols_rows._2), projected_extent.extent, projected_extent.crs, Tags.empty, GeoTiffOptions.DEFAULT).tile
//Load Mask
if (toBeMasked) {
val mask_tiles_RDD = sc.hadoopGeoTiffRDD(mask_path).values
val mask_tiles_withIndex = mask_tiles_RDD.zipWithIndex().map{case (e,v) => (v,e)}
mask_tile0 = (mask_tiles_withIndex.filter(m => m._1==0).values.collect())(0)
}
//Local variables
val pattern: String = "tif"
val filepath: String = dir_path + geoTiff_dir
val filepath_2: String = dir_path + geoTiff_2_dir
if (rdd_offline_mode) {
grids_noNaN_RDD = sc.objectFile(grids_noNaN_path)
grid0 = sc.objectFile(grid0_path)
grid0_index = sc.objectFile(grid0_index_path)
val metadata = sc.sequenceFile(metadata_path, classOf[IntWritable], classOf[BytesWritable]).map(_._2.copyBytes()).collect()
projected_extent = deserialize(metadata(0)).asInstanceOf[ProjectedExtent]
num_cols_rows = (deserialize(metadata(1)).asInstanceOf[Int], deserialize(metadata(2)).asInstanceOf[Int])
} else {
if (single_band) {
//Lets load a Singleband GeoTiffs and return RDD just with the tiles.
var tiles_1_RDD :RDD[Tile] = sc.hadoopGeoTiffRDD(filepath, pattern).values
var tiles_2_RDD :RDD[Tile] = sc.hadoopGeoTiffRDD(filepath_2, pattern).values
//Retrive the numbre of cols and rows of the Tile's grid
val tiles_withIndex = tiles_1_RDD.zipWithIndex().map{case (e,v) => (v,e)}
val tile0 = (tiles_withIndex.filter(m => m._1==0).values.collect())(0)
num_cols_rows = (tile0.cols,tile0.rows)
cellT = tile0.cellType
val tiles_RDD = sc.union([tiles_1_RDD, tiles_2_RDD])
if (toBeMasked) {
val mask_tile_broad :Broadcast[Tile] = sc.broadcast(mask_tile0)
grids_RDD = tiles_RDD.map(m => m.localInverseMask(mask_tile_broad.value, 1, 0).toArrayDouble())
} else {
grids_RDD = tiles_RDD.map(m => m.toArrayDouble())
}
} else {
//Lets load Multiband GeoTiffs and return RDD just with the tiles.
val tiles_1_RDD = sc.hadoopMultibandGeoTiffRDD(filepath, pattern).values
val tiles_2_RDD = sc.hadoopMultibandGeoTiffRDD(filepath_2, pattern).values
//Retrive the numbre of cols and rows of the Tile's grid
val tiles_withIndex = tiles_1_RDD.zipWithIndex().map{case (e,v) => (v,e)}
val tile0 = (tiles_withIndex.filter(m => m._1==0).values.collect())(0)
num_cols_rows = (tile0.cols,tile0.rows)
cellT = tile0.cellType
val tiles_RDD = sc.union([tiles_1_RDD, tiles_2_RDD])
//Lets read the average of the Spring-Index which is stored in the 4th band
val band_numB :Broadcast[Int] = sc.broadcast(band_num)
if (toBeMasked) {
val mask_tile_broad :Broadcast[Tile] = sc.broadcast(mask_tile0)
grids_RDD = tiles_RDD.map(m => m.band(band_numB.value).localInverseMask(mask_tile_broad.value, 1, 0).toArrayDouble().map(m => if (m == 0.0) Double.NaN else m))
} else {
grids_RDD = tiles_RDD.map(m => m.band(band_numB.value).toArrayDouble())
}
}
//Retrieve the ProjectExtent which contains metadata such as CRS and bounding box
val projected_extents_withIndex = sc.hadoopGeoTiffRDD(filepath, pattern).keys.zipWithIndex().map{case (e,v) => (v,e)}
projected_extent = (projected_extents_withIndex.filter(m => m._1 == 0).values.collect())(0)
//Get Index for each Cell
val grids_withIndex = grids_RDD.zipWithIndex().map { case (e, v) => (v, e) }
grid0_index = grids_withIndex.filter(m => m._1 == 0).values.flatMap(m => m).zipWithIndex.filter(m => !m._1.isNaN).map { case (v, i) => (i) }
//Get the Tile's grid
grid0 = grids_withIndex.filter(m => m._1 == 0).values.flatMap( m => m).zipWithIndex.map{case (v,i) => (i,v)}
//Lets filter out NaN
grids_noNaN_RDD = grids_RDD.map(m => m.filter(!_.isNaN))
//Store data in HDFS
grid0.saveAsObjectFile(grid0_path)
grid0_index.saveAsObjectFile(grid0_index_path)
grids_noNaN_RDD.saveAsObjectFile(grids_noNaN_path)
val grids_noNaN_RDD_withIndex = grids_noNaN_RDD.zipWithIndex().map { case (e, v) => (v, e) }
grids_noNaN_RDD = grids_noNaN_RDD_withIndex.filterByRange(model_years_range._1, model_years_range._2).values
val writer: SequenceFile.Writer = SequenceFile.createWriter(conf,
Writer.file(metadata_path),
Writer.keyClass(classOf[IntWritable]),
Writer.valueClass(classOf[BytesWritable])
)
writer.append(new IntWritable(1), new BytesWritable(serialize(projected_extent)))
writer.append(new IntWritable(2), new BytesWritable(serialize(num_cols_rows._1)))
writer.append(new IntWritable(3), new BytesWritable(serialize(num_cols_rows._2)))
writer.hflush()
writer.close()
}
val t1 = System.nanoTime()
println("Elapsed time: " + (t1 - t0) + "ns")
```
## Matrix
We need to do a Matrix transpose to have clusters per cell and not per year. With a GeoTiff representing a single year, the loaded data looks liks this:
```
bands_RDD.map(s => Vectors.dense(s)).cache()
//The vectors are rows and therefore the matrix will look like this:
[
Vectors.dense(0.0, 1.0, 2.0),
Vectors.dense(3.0, 4.0, 5.0),
Vectors.dense(6.0, 7.0, 8.0),
Vectors.dense(9.0, 0.0, 1.0)
]
```
To achieve that we convert the **RDD[Vector]** into a distributed Matrix, a [**CoordinateMatrix**](https://spark.apache.org/docs/latest/mllib-data-types.html#coordinatematrix), which as a **transpose** method.
```
val t0 = System.nanoTime()
//Global variables
var grids_matrix: RDD[Vector] = sc.emptyRDD
if (matrix_offline_mode) {
grids_matrix = sc.objectFile(grids_matrix_path)
} else {
val mat :RowMatrix = new RowMatrix(grids_noNaN_RDD.map(m => Vectors.dense(m)))
// Split the matrix into one number per line.
val byColumnAndRow = mat.rows.zipWithIndex.map {
case (row, rowIndex) => row.toArray.zipWithIndex.map {
case (number, columnIndex) => new MatrixEntry(rowIndex, columnIndex, number)
}
}.flatMap(x => x)
val matt: CoordinateMatrix = new CoordinateMatrix(byColumnAndRow)
val matt_T = matt.transpose()
//grids_matrix = matt_T.toRowMatrix().rows
grids_matrix = matt_T.toIndexedRowMatrix().rows.sortBy(_.index).map(_.vector)
grids_matrix.saveAsObjectFile(grids_matrix_path)
}
val t1 = System.nanoTime()
println("Elapsed time: " + (t1 - t0) + "ns")
```
## Kmeans
We use Kmeans from Sparl-MLlib. The user should only modify the variables on Kmeans setup.
### Kmeans Training
```
val t0 = System.nanoTime()
//Global variables
var kmeans_models :Array[KMeansModel] = new Array[KMeansModel](num_kmeans)
var wssse_data :List[(Int, Int, Double)] = List.empty
if (kmeans_offline_mode) {
var numClusters_id = 0
cfor(minClusters)(_ <= maxClusters, _ + stepClusters) { numClusters =>
if (!fs.exists(new org.apache.hadoop.fs.Path(kmeans_model_paths(numClusters_id)))) {
println("One of the files does not exist, we will abort!!!")
System.exit(0)
} else {
kmeans_models(numClusters_id) = KMeansModel.load(sc, kmeans_model_paths(numClusters_id))
}
numClusters_id += 1
}
val wssse_data_RDD :RDD[(Int, Int, Double)] = sc.objectFile(wssse_path)
wssse_data = wssse_data_RDD.collect().toList
} else {
var numClusters_id = 0
if (fs.exists(new org.apache.hadoop.fs.Path(wssse_path))) {
val wssse_data_RDD :RDD[(Int, Int, Double)] = sc.objectFile(wssse_path)
wssse_data = wssse_data_RDD.collect().toList
}
grids_matrix.cache()
cfor(minClusters)(_ <= maxClusters, _ + stepClusters) { numClusters =>
println(numClusters)
kmeans_models(numClusters_id) = {
KMeans.train(grids_matrix, numClusters, numIterations)
}
// Evaluate clustering by computing Within Set Sum of Squared Errors
val WSSSE = kmeans_models(numClusters_id).computeCost(grids_matrix)
println("Within Set Sum of Squared Errors = " + WSSSE)
wssse_data = wssse_data :+ (numClusters, numIterations, WSSSE)
//Save kmeans model
if (save_kmeans_model) {
if (!fs.exists(new org.apache.hadoop.fs.Path(kmeans_model_paths(numClusters_id)))) {
kmeans_models(numClusters_id).save(sc, kmeans_model_paths(numClusters_id))
}
}
numClusters_id += 1
}
//Un-persist it to save memory
grids_matrix.unpersist()
if (fs.exists(new org.apache.hadoop.fs.Path(wssse_path))) {
println("We will delete the wssse file")
try { fs.delete(new org.apache.hadoop.fs.Path(wssse_path), true) } catch { case _ : Throwable => { } }
}
println("Lets create it with the new data")
sc.parallelize(wssse_data, 1).saveAsObjectFile(wssse_path)
}
val t1 = System.nanoTime()
println("Elapsed time: " + (t1 - t0) + "ns")
```
### Inspect WSSSE
```
val t0 = System.nanoTime()
//current
println(wssse_data)
//from disk
if (fs.exists(new org.apache.hadoop.fs.Path(wssse_path))) {
var wssse_data_tmp :RDD[(Int, Int, Double)] = sc.objectFile(wssse_path)//.collect()//.toList
println(wssse_data_tmp.collect().toList)
}
val t1 = System.nanoTime()
println("Elapsed time: " + (t1 - t0) + "ns")
```
### Run Kmeans clustering
Run Kmeans and obtain the clusters per each cell.
```
val t0 = System.nanoTime()
//Cache it so kmeans is more efficient
grids_matrix.cache()
var kmeans_res: Array[RDD[Int]] = Array.fill(num_kmeans)(sc.emptyRDD)
var numClusters_id = 0
cfor(minClusters)(_ <= maxClusters, _ + stepClusters) { numClusters =>
kmeans_res(numClusters_id) = kmeans_models(numClusters_id).predict(grids_matrix)
numClusters_id += 1
}
//Un-persist it to save memory
grids_matrix.unpersist()
val t1 = System.nanoTime()
println("Elapsed time: " + (t1 - t0) + "ns")
```
#### Sanity test
It can be skipped, it only shows the cluster ID for the first 50 cells
```
val t0 = System.nanoTime()
val kmeans_res_out = kmeans_res(0).take(150)
kmeans_res_out.foreach(print)
println(kmeans_res_out.size)
val t1 = System.nanoTime()
println("Elapsed time: " + (t1 - t0) + "ns")
```
## Build GeoTiff with Kmeans cluster_IDs
The Grid with the cluster IDs is stored in a SingleBand GeoTiff and uploaded to HDFS.
### Assign cluster ID to each grid cell and save the grid as SingleBand GeoTiff
To assign the clusterID to each grid cell it is necessary to get the indices of gird cells they belong to. The process is not straight forward because the ArrayDouble used for the creation of each dense Vector does not contain the NaN values, therefore there is not a direct between the indices in the Tile's grid and the ones in **kmeans_res** (kmeans result).
To join the two RDDS the knowledge was obtaing from a stackoverflow post on [how to perform basic joins of two rdd tables in spark using python](https://stackoverflow.com/questions/31257077/how-do-you-perform-basic-joins-of-two-rdd-tables-in-spark-using-python).
```
val t0 = System.nanoTime()
var numClusters_id = 0
cfor(minClusters)(_ <= maxClusters, _ + stepClusters) { numClusters =>
//Merge two RDDs, one containing the clusters_ID indices and the other one the indices of a Tile's grid cells
val cluster_cell_pos = ((kmeans_res(numClusters_id).zipWithIndex().map{ case (v,i) => (i,v)}).join(grid0_index.zipWithIndex().map{ case (v,i) => (i,v)})).map{ case (k,(v,i)) => (v,i)}
//Associate a Cluster_IDs to respective Grid_cell
val grid_clusters = grid0.leftOuterJoin(cluster_cell_pos.map{ case (c,i) => (i.toLong, c)})
//Convert all None to NaN
val grid_clusters_res = grid_clusters.sortByKey(true).map{case (k, (v, c)) => if (c == None) (k, Double.NaN) else (k, c.get.toDouble)}
//Define a Tile
val cluster_cells :Array[Double] = grid_clusters_res.values.collect()
val cluster_cellsD = DoubleArrayTile(cluster_cells, num_cols_rows._1, num_cols_rows._2)
val cluster_tile = geotrellis.raster.DoubleArrayTile.empty(num_cols_rows._1, num_cols_rows._2)
cfor(0)(_ < num_cols_rows._1, _ + 1) { col =>
cfor(0)(_ < num_cols_rows._2, _ + 1) { row =>
val v = cluster_cellsD.getDouble(col, row)
if (v != Double.NaN)
cluster_tile.setDouble(col, row, v)
}
}
val geoTif = new SinglebandGeoTiff(cluster_tile, projected_extent.extent, projected_extent.crs, Tags.empty, GeoTiffOptions(compression.DeflateCompression))
//Save to /tmp/
GeoTiffWriter.write(geoTif, geotiff_tmp_paths(numClusters_id))
//Upload to HDFS
var cmd = "hadoop dfs -copyFromLocal -f " + geotiff_tmp_paths(numClusters_id) + " " + geotiff_hdfs_paths(numClusters_id)
Process(cmd)!
//Remove from /tmp/
cmd = "rm -fr " + geotiff_tmp_paths(numClusters_id)
Process(cmd)!
numClusters_id += 1
}
val t1 = System.nanoTime()
println("Elapsed time: " + (t1 - t0) + "ns")
```
# [Visualize results](plot_kmeans_clusters.ipynb) --------------- [Plot WSSE](kmeans_wsse.ipynb)
| github_jupyter |
# RNN with TensorFlow API
```
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
%matplotlib inline
```
### The Data
```
# Create a dataset
class TimeSeriesData():
def __init__(self,num_points,xmin,xmax):
self.xmin = xmin
self.xmax = xmax
self.num_points = num_points
self.resolution = (xmax - xmin) / num_points
self.x_data = np.linspace(xmin, xmax, num_points)
self.y_true = np.sin(self.x_data)
def ret_true(self, x_series):
return np.sin(x_series)
def next_batch(self, batch_size, steps, return_batch_ts = False):
# Grab a random starting point for each batch
rand_start = np.random.rand(batch_size, 1)
# Convert to be on time series
ts_start = rand_start * (self.xmax - self.xmin - (steps * self.resolution) )
# Create batch Time Series on t axis
batch_ts = ts_start + np.arange(0.0, steps + 1) * self.resolution
# Create Y data for time series in the batches
y_batch = np.sin(batch_ts)
# Format for RNN
if return_batch_ts:
return y_batch[:, :-1].reshape(-1, steps, 1), y_batch[:, 1:].reshape(-1, steps, 1) , batch_ts
else:
return y_batch[:, :-1].reshape(-1, steps, 1), y_batch[:, 1:].reshape(-1, steps, 1)
ts_data = TimeSeriesData(250, 0, 10)
plt.plot(ts_data.x_data, ts_data.y_true)
# Num of steps in batch (also used for prediction steps into the future)
num_time_steps = 30
y1, y2, ts = ts_data.next_batch(1, num_time_steps, True)
y1.flatten()
plt.plot(ts.flatten()[1:], y2.flatten(), '*')
plt.plot(ts_data.x_data,
ts_data.y_true,
label = 'Sin(t)')
plt.plot(ts.flatten()[1:],
y2.flatten(),
'*',
label = 'Single Training Instance')
plt.legend()
plt.tight_layout()
```
### A Training Instance and what to predict
We are trying to predict a time series shifted over by t+1
```
train_inst = np.linspace(5,5 + ts_data.resolution * (num_time_steps + 1), num_time_steps + 1)
plt.title("A training instance", fontsize=14)
plt.plot(train_inst[:-1], ts_data.ret_true(train_inst[:-1]),
"bo",
markersize = 15,
alpha = 0.5 ,
label = "instance")
plt.plot(train_inst[1:], ts_data.ret_true(train_inst[1:]),
"ko",
markersize = 7,
label = "target")
```
___________
# Creating the Model
```
tf.reset_default_graph()
```
### Constants
```
# Just one feature, the time series
num_inputs = 1
# 100 neuron layer, play with this
num_neurons = 100
# Just one output, predicted time series
num_outputs = 1
# learning rate, 0.0001 default, but you can play with this
learning_rate = 0.0001
# how many iterations to go through (training steps), you can play with this
num_train_iterations = 2000
# Size of the batch of data
batch_size = 1
```
### Placeholders
```
X = tf.placeholder(tf.float32, [None, num_time_steps, num_inputs])
y = tf.placeholder(tf.float32, [None, num_time_steps, num_outputs])
```
____
____
### RNN Cell Layer
Play around with the various cells in this section, compare how they perform against each other.
```
cell = tf.contrib.rnn.OutputProjectionWrapper(
tf.contrib.rnn.BasicRNNCell(num_units = num_neurons,
activation = tf.nn.relu),
output_size = num_outputs)
# cell = tf.contrib.rnn.OutputProjectionWrapper(
# tf.contrib.rnn.BasicLSTMCell(num_units=num_neurons, activation=tf.nn.relu),
# output_size=num_outputs)
# n_neurons = 100
# n_layers = 3
# cell = tf.contrib.rnn.MultiRNNCell([tf.contrib.rnn.BasicRNNCell(num_units=n_neurons)
# for layer in range(n_layers)])
# cell = tf.contrib.rnn.BasicLSTMCell(num_units=num_neurons, activation=tf.nn.relu)
# n_neurons = 100
# n_layers = 3
# cell = tf.contrib.rnn.MultiRNNCell([tf.contrib.rnn.BasicLSTMCell(num_units=n_neurons)
# for layer in range(n_layers)])
```
_____
_____
### Dynamic RNN Cell
```
outputs, states = tf.nn.dynamic_rnn(cell, X, dtype = tf.float32)
```
### Loss Function and Optimizer
```
loss = tf.reduce_mean(tf.square(outputs - y)) # MSE
optimizer = tf.train.AdamOptimizer(learning_rate = learning_rate)
train = optimizer.minimize(loss)
```
#### Init Variables
```
init = tf.global_variables_initializer()
```
## Session
```
# ONLY FOR GPU USERS:
# https://stackoverflow.com/questions/34199233/how-to-prevent-tensorflow-from-allocating-the-totality-of-a-gpu-memory
gpu_options = tf.GPUOptions(per_process_gpu_memory_fraction = 0.75)
saver = tf.train.Saver()
with tf.Session(config=tf.ConfigProto(gpu_options = gpu_options)) as sess:
sess.run(init)
for iteration in range(num_train_iterations):
X_batch, y_batch = ts_data.next_batch(batch_size, num_time_steps)
sess.run(train, feed_dict = {X: X_batch,
y: y_batch})
if iteration % 100 == 0:
mse = loss.eval(feed_dict = {X: X_batch, y: y_batch})
print(iteration, "\tMSE:", mse)
# Save Model for Later
saver.save(sess, "./checkpoints/rnn_time_series_model")
```
### Predicting a time series t+1
```
with tf.Session() as sess:
saver.restore(sess, "./checkpoints/rnn_time_series_model")
X_new = np.sin(np.array(train_inst[:-1].reshape(-1, num_time_steps, num_inputs)))
y_pred = sess.run(outputs, feed_dict = {X: X_new})
plt.title("Testing Model")
# Training Instance
plt.plot(train_inst[:-1], np.sin(train_inst[:-1]),
"bo",
markersize = 15,
alpha = 0.5,
label = "Training Instance")
# Target to Predict
plt.plot(train_inst[1:], np.sin(train_inst[1:]),
"ko",
markersize = 10,
label = "target")
# Models Prediction
plt.plot(train_inst[1:], y_pred[0,:,0],
"r.",
markersize = 10,
label = "prediction")
plt.xlabel("Time")
plt.legend()
plt.tight_layout()
```
# Generating New Sequences
** Note: Can give wacky results sometimes, like exponential growth**
```
with tf.Session() as sess:
saver.restore(sess, "./checkpoints/rnn_time_series_model")
# SEED WITH ZEROS
zero_seq_seed = [0. for i in range(num_time_steps)]
for iteration in range(len(ts_data.x_data) - num_time_steps):
X_batch = np.array(zero_seq_seed[-num_time_steps:]).reshape(1, num_time_steps, 1)
y_pred = sess.run(outputs, feed_dict={X: X_batch})
zero_seq_seed.append(y_pred[0, -1, 0])
plt.plot(ts_data.x_data, zero_seq_seed,
"b-")
plt.plot(ts_data.x_data[:num_time_steps],
zero_seq_seed[:num_time_steps],
"r",
linewidth = 3)
plt.xlabel("Time")
plt.ylabel("Value")
with tf.Session() as sess:
saver.restore(sess, "./checkpoints/rnn_time_series_model")
# SEED WITH Training Instance
training_instance = list(ts_data.y_true[:30])
for iteration in range(len(training_instance) - num_time_steps):
X_batch = np.array(training_instance[-num_time_steps:]).reshape(1, num_time_steps, 1)
y_pred = sess.run(outputs, feed_dict={X: X_batch})
training_instance.append(y_pred[0, -1, 0])
plt.plot(ts_data.x_data, ts_data.y_true, "b-")
plt.plot(ts_data.x_data[:num_time_steps],
training_instance[:num_time_steps],
"r-",
linewidth = 3)
plt.xlabel("Time")
```
# Great Job!
| github_jupyter |
# Binary options put, call statistical research
This Notebook analyse forex data from EURUSD pair M1 timeframe for the binary options market
the data is from the duration of january 1st 2020 to december 31st 2020, the aim of this research is to find a pattern
the placing trades in the binary options markert, in other to accurately predict where a market will close, from
the previous candles, if it closes above the open price of the candle in question it is a call option, if it closes below it is a put option, when the open price and the close price is the same this is a no trade option which is statistically rare, so this won't be a focal point in this study
```
#import the relevant packages
import pandas as pd
import numpy as np
import seaborn as sns
sns.set(color_codes=True)
#read the data from a csv file
data = pd.read_csv("EURUSD_M1_202001020600_202012310000.csv", sep="\t")
data
#drop the <VOL>, column as it is not needed in this research
data.drop("<VOL>", inplace=True, axis=1)
data
#set call when open price is less than close price
data["<UP>"] = data["<OPEN>"] < data["<CLOSE>"]
#set put when open price is greater than close price
data["<DOWN>"] = data["<OPEN>"] > data["<CLOSE>"]
#no trade when open price is equal to close price
data["<NO_MOVE>"] = data["<OPEN>"] == data["<CLOSE>"]
data
#convert these columns to type int so they appear as one hot varibles
data[["<UP>", "<DOWN>", "<NO_MOVE>"]] = data[["<UP>", "<DOWN>", "<NO_MOVE>"]].astype("int32")
data
```
```
open_close = (data["<OPEN>"] - data["<CLOSE>"]) * 100000
open_low = (data["<OPEN>"] - data["<LOW>"]) * 100000
open_high = (data["<OPEN>"] - data["<HIGH>"]) * 100000
close_low = (data["<CLOSE>"] - data["<LOW>"]) * 100000
close_high = (data["<CLOSE>"] - data["<HIGH>"]) * 100000
high_low = (data["<HIGH>"] - data["<LOW>"]) * 100000
#create a dataframe of the subtracted columns
move_df = pd.DataFrame({"open_close" : open_close,
"open_low" : open_low,
"open_high" : open_high,
"close_low" : close_low,
"close_high" : close_high,
"high_low" : high_low, })
move_df
```
### create a future dataframe to store the next data i.e the candle in question
```
future = data.iloc[1:,8:11]
future.columns = ["up*", "down*", "no_move*"]
future.reset_index(inplace=True)
future.drop(["index"], axis = 1, inplace=True)
#concat move_df and future to create a new dataframe
dataset = pd.concat([move_df, data.iloc[:,2:6], future], axis = 1)
dataset
dataset.dropna(inplace=True)
dataset
#get the absolute value of all the data
absolute_data = abs(dataset)
absolute_data
absolute_data.describe()
absolute_data.quantile([.95, .99, .995, .999])
#display frequencies of up, down and no move
dataset[["up*", "down*", "no_move*"]].value_counts()
```
# 95 percentile
# Open_Close Analysis
<li> <b> Anaylyse candles after a prior large drop in prices up to 3.2 pips in a minute open_close </b> </li>
```
big_drop_oc_95 = dataset[dataset["open_close"] > 32]
big_drop_oc_95
#get the number of up and down movement in price after a big downward movement in price
big_drop_oc_95[["up*", "down*", "no_move*"]].value_counts()
```
## Result
From the information above we had 4761 upward movement in price and 4384 downward movement in price total <br/>
and 202 no movement in price <br/>
candles = 4761 + 4384 + 202 = 9347 <br/>
percentage up = (100 * 4761)/(9347) = 50.94% <br/>
percentage down = (100 * 4384)/(9347) = 46.90% <br/>
percentage no move = (100 * 202)/(9347) = 2.16% <br/>
so after a 3.2 pips rise in price for a one minute time frame of EURUSD there was a 50.94% chance that the next candle would be bullish and 46.90% chance of a bearish candle and 2.16% chance of no movement
<li> <b> Anaylyse candles after a prior large drop in prices up to 3.2 pips in a minute open_close </b> </li>
```
big_rise_oc_95 = dataset[dataset["open_close"] < -32]
big_rise_oc_95
big_rise_oc_95[["up*", "down*", "no_move*"]].value_counts()
```
## Result
From the information above we had 4448 upward movement in price and 4304 downward movement in price total <br/>
and 191 no move that is open and close are equal
candles = 4448 + 4304 + 191 = 8943 <br/>
percentage up = (100 * 4448)/(8943) = 49.74% <br/>
percentage down = (100 * 4304)/(8943) = 48.13% <br/>
percentage no move = (100 * 191)/(8943) = 2.13% <br/>
so after a 3.2 pips rise in price for a one minute time frame of EURUSD there was a 49.74% chance that the next candle would be bullish and 48.13% chance of a bearish candle and 1.58% chance open and close are equal
### Anaylyse candles after a prior large drop in prices up to 3.1 pips in a minute open_low
```
big_drop_ol_95 = dataset[dataset["open_low"] > 31]
big_drop_ol_95
#get the number of up and down movement in price after a big downward movement in price
big_drop_ol_95[["up*", "down*", "no_move*"]].value_counts()
```
## Result
From the information above we had 9413 upward movement in price and 8860 downward movement in price total <br/>
and 442 no move that is open and close are equal
candles = 9413 + 8860 + 442 = 18715 <br/>
percentage up = (100 * 9413)/(18715) = 50.30% <br/>
percentage down = (100 * 8860)/(18715) = 47.34% <br/>
percentage no move = (100 * 442)/(18715) = 2.36% <br/>
so after a 3.1 pips drop in price for a one minute time frame of EURUSD there was a 49.74% chance that the next candle would be bullish and 48.13% chance of a bearish candle and 1.58% chance open and close are equal
### Anaylyse candles after a prior large drop in prices up to 3.0 pips in a minute open_low
```
big_rise_oh_95 = dataset[dataset["open_high"] < -30]
big_rise_oh_95
#get the number of up and down movement in price after a big upward movement in price
big_rise_oh_95[["up*", "down*", "no_move*"]].value_counts()
```
## Result
From the information above we had 9152 upward movement in price and 9146 downward movement in price total <br/>
and 479 no move that is open and close are equal
candles = 9152 + 9146 + 479 = 18777 <br/>
percentage up = (100 * 9152)/(18777) = 48.74% <br/>
percentage down = (100 * 9146)/(18777) = 48.71% <br/>
percentage no move = (100 * 479)/(18777) = 2.55% <br/>
so after a 3.1 pips drop in price for a one minute time frame of EURUSD there was a 48.74% chance that the next candle would be bullish and 48.71% chance of a bearish candle and 1.58% chance open and close are equal
# Close_Low Analysis
<b><li> Anaylyse candles after a prior large move in prices up to 3.0 pips in a minute close_low that is bearish </li></b>
```
big_move_cl_bear_95 = dataset[dataset["close_low"] > 30]
big_move_cl_bear_95 = big_move_cl_bear_95[big_move_cl_bear_95["open_close"] > 0]
big_move_cl_bear_95
big_move_cl_bear_95[["up*", "down*", "no_move*"]].value_counts()
```
## Result
From the information above we had 688 upward movement in price and 651 downward movement in price total <br/>
and 27 no move that is open and close are equal
candles = 688 + 651 + 27 = 1366 <br/>
percentage up = (100 * 688)/(1366) = 50.37% <br/>
percentage down = (100 * 651)/(1366) = 47.66% <br/>
percentage no move = (100 * 27)/(1366) = 1.98% <br/>
so after a 3.0 pips drop in price for a one minute time frame of EURUSD there was a 50.37% chance that the next candle would be bullish and 47.66% chance of a bearish candle and 1.98% chance open and close are equal
<b><li> Anaylyse candles after a prior large move in prices up to 3.0 pips in a minute close_low that is bullish </li></b>
```
big_move_cl_bull_95 = dataset[dataset["close_low"] > 30]
big_move_cl_bull_95 = big_move_cl_bull_95[big_move_cl_bull_95["open_close"] < 0]
big_move_cl_bull_95
big_move_cl_bull_95[["up*", "down*", "no_move*"]].value_counts()
```
## Result
From the information above we had 7809 upward movement in price and 8012 downward movement in price total <br/>
and 398 no move that is open and close are equal
candles = 7809 + 8012 + 398 = 16219<br/>
percentage up = (100 * 7809)/(16219) = 48.15% <br/>
percentage down = (100 * 8012)/(16219) = 49.40% <br/>
percentage no move = (100 * 398)/(16219) = 2.45% <br/>
so after a 3.0 pips drop in price for a one minute time frame of EURUSD there was a 48.15% chance that the next candle would be bullish and 49.40% chance of a bullish candle and 2.45% chance open and close are equal
# Close_High Analysis
<b> <li> Anaylyse candles after a prior large move in prices up to 3.0 pips in a minute close_high when the candle is bearish </li> </b>
```
big_drop_ch_95 = dataset[dataset["close_high"] < -30]
big_drop_ch_95 = big_drop_ch_95[big_drop_ch_95["open_close"] > 0]
big_drop_ch_95
big_drop_ch_95[["up*", "down*", "no_move*"]].value_counts()
```
## Result
From the information above we had 8407 upward movement in price and 7832 downward movement in price total <br/>
and 358 no move that is open and close are equal
candles = 8407 + 7832 + 358 = 16597 <br/>
percentage up = (100 * 8407)/(16597) = 50.65% <br/>
percentage down = (100 * 7832)/(16597) = 47.19% <br/>
percentage no move = (100 * 358)/(16597) = 2.16% <br/>
so after a 3.0 pips drop in price for a one minute time frame of EURUSD there was a 50.65% chance that the next candle would be bullish and 47.19% chance of a bearish candle and 2.16% chance open and close are equal
### Anaylyse candles after a prior large move in prices up to 3.0 pips in a minute close_high when the candle is bullish
```
big_rise_ch_95 = dataset[dataset["close_high"] < -30]
big_rise_ch_95 = big_rise_ch_95[big_rise_ch_95["open_close"] < 0]
big_rise_ch_95
big_rise_ch_95[["up*", "down*", "no_move*"]].value_counts()
```
## Result
From the information above we had 714 upward movement in price and 699 downward movement in price total <br/>
and 28 no move that is open and close are equal
candles = 714 + 699 + 28 = 1441 <br/>
percentage up = (100 * 714)/(1441) = 49.55% <br/>
percentage down = (100 * 699)/(1441) = 48.51% <br/>
percentage no move = (100 * 28)/(1441) = 1.94% <br/>
so after a 3.0 pips move in price for a one minute time frame of EURUSD there was a 49.55% chance that the next candle would be bullish and 48.51% chance of a bearish candle and 1.94% chance open and close are equal
### Anaylyse candles after a prior large drop in prices up to 4.6 pips in a minute high_low
```
big_drop_hl_95 = dataset[dataset["high_low"] > 46]
big_drop_hl_95 = big_drop_hl_95[big_drop_hl_95["open_close"] > 0]
big_drop_hl_95
big_drop_hl_95[["up*", "down*", "no_move*"]].value_counts()
```
## Result
From the information above we had 4900 upward movement in price and 4462 downward movement in price total <br/>
and 173 no move that is open and close are equal
candles = 4900 + 4462 + 173 = 9535 <br/>
percentage up = (100 * 4900)/(9535) = 51.39% <br/>
percentage down = (100 * 4462)/(9535) = 46.80% <br/>
percentage no move = (100 * 173)/(9535) = 1.81% <br/>
so after a 4.6 pips drop in price for a one minute time frame of EURUSD there was a 51.39% chance that the next candle would be bullish and 46.80% chance of a bearish candle and 1.81% chance open and close are equal
### Anaylyse candles after a prior large rise in prices up to 4.6 pips in a minute high_low
```
big_rise_hl_95 = dataset[dataset["high_low"] > 46]
big_rise_hl_95 = big_rise_hl_95[big_rise_hl_95["open_close"] < 0]
big_rise_hl_95
big_rise_hl_95[["up*", "down*", "no_move*"]].value_counts()
```
## Result
From the information above we had 4532 upward movement in price and 4661 downward movement in price total <br/>
and 183 no move that is open and close are equal
candles = 4532 + 4661 + 183 = 9376 <br/>
percentage up = (100 * 4661)/(9376) = 49.71% <br/>
percentage down = (100 * 4532)/(9376) = 48.34% <br/>
percentage no move = (100 * 183)/(9376) = 1.95% <br/>
so after a 4.6 pips drop in price for a one minute time frame of EURUSD there was a 49.71% chance that the next candle would be bullish and 48.34% chance of a bearish candle and 1.95% chance open and close are equal
| github_jupyter |
# Quantum Fourier Transform
The classical Fourier transform is an important tool in wave and signal analysis that breaks a function into components, each with a different frequency.
Its discrete analogue, the Discrete Fourier transform acts on $n$ complex numbers $x_0,\ldots,x_{N-1}$ and transforms it to another sequence of $n$ complex numbers $\tilde x_0,\ldots,\tilde x_{N-1}$ via:
$$\tilde x_k = \sum_{y=0}^{N-1}e^{-\frac{2\pi ikn}N} \cdot x_k$$
The quantum Fourier transform (commonly abbreviated as QFT) on $n$ qubits (with $N=2^n$), achieves an analogous effect, and is given by the following equation (for every basis state $x\in\{0,1\}^n$):
$$\text{QFT}(\lvert x\rangle) = \frac 1{\sqrt N}\sum_{y=0}^{N-1}e^{\frac{2\pi ixy}N}\lvert y\rangle\qquad \qquad (1)$$
Here, we abuse notation and associate a bitstring $x\in\{0,1\}^n$ to the integer $\sum_{j=0}^{n-1}2^{n-1-j}x_j$.
Effectively, one can think of QFT as a change of basis from the computational basis to the "Fourier basis".
## Factorization of QFT
It turns out (after some algebraic manipulations) that one is able to factorize the sum in Eq. $(1)$ into a product of pure states
$$\text{QFT}(\lvert x\rangle) = \frac 1{\sqrt N}\left(\vert 0\rangle + e^{\frac{2\pi ix}{2^1}}\vert 1\rangle\right)\otimes \left(\vert 0\rangle + e^{\frac{2\pi ix}{2^2}}\vert 1\rangle\right)\otimes \cdots \otimes \left(\vert 0\rangle + e^{\frac{2\pi ix}{2^n}}\vert 1\rangle\right).$$
This formulation tells us exactly how to implement a quantum circuit achieving QFT.
## Controlled Phase Rotations
Before looking at the circuit structure though, we define the gate
$$R_k = \begin{pmatrix}1 & 0\\ 0 & e^{\frac{2\pi i}{2^k}}\end{pmatrix},$$
and recall that a controlled version of this gate will change the phase of the target qubit depending on the value of the control qubit.
In other words, $CR_k(\lvert 0\rangle|x\rangle) = |0\rangle \lvert x\rangle$ and
\begin{aligned}CR_k(\lvert 1\rangle \lvert x\rangle) = \lvert 1\rangle\otimes R_k(\lvert x \rangle) &=\lvert 1\rangle\otimes R_k\left( \langle 0\lvert x\rangle \lvert 0\rangle + \langle 1 \lvert x\rangle \lvert 1\rangle\right)
\\ & = \lvert 1\rangle\otimes \left( \langle 0 \lvert x\rangle \lvert 0\rangle + \langle 1 \lvert x\rangle e^{\frac{2\pi i}{2^k}} \lvert 1\rangle\right)\end{aligned}
More compactly,
$$CR_k(\lvert y\rangle \lvert x\rangle) = \lvert y\rangle \otimes \left( \langle 0 \vert x\rangle \lvert 0\rangle + \langle 1 \vert x\rangle e^{\frac{2\pi iy}{2^k}} \lvert 1\rangle\right).$$
Therefore, $$CR_k = \begin{pmatrix}1 & 0 & 0 & 0\\ 0 & 1 & 0 & 0\\ 0& 0& 1 & 0\\ 0 & 0&0 &e^{\frac{2\pi i}{2^k}}\end{pmatrix} = \text{CPhase}\left(\frac{2\pi}{2^k}\right).$$
## Circuit Structure
Now we have all the ingredients required to construct a circuit achieving QFT.
Below is an example circuit on $n=4$ qubits:

We shall work our way through the states $\lvert \varphi_1\rangle$ through $\lvert \varphi_4\rangle$.
In $\lvert \varphi_1\rangle$, let us analyze the state of the first qubit on the basis states $x_0\in\{0,1\}$. Then $\lvert x_0\rangle$ is mapped to (let us ignore normalizations for simplifying calculations):
+ $H$: $\lvert 0\rangle + e^{\pi i x_0}\lvert 1\rangle$,
+ $R_2$: $\lvert 0\rangle + e^{\pi i x_0 + \frac{2\pi i x_1}{2^2}}\lvert 1\rangle$
+ $R_3$: $\lvert 0\rangle + e^{\pi i x_0 + \frac{2\pi i x_1}{2^2} + \frac{2\pi i x_2}{2^3}}\lvert 1\rangle$
+ $R_4$: $\lvert 0\rangle + e^{\pi i x_0 + \frac{2\pi i x_1}{2^2} + \frac{2\pi i x_2}{2^3} + \frac{2\pi i x_3}{2^4}}\lvert 1\rangle = \lvert 0\rangle + e^{\frac{2\pi i}{2^4} (8x_0 + 4x_1 + 2x_2 + x_3)}\lvert 1\rangle = |0\rangle + e^{\frac{2\pi i x}{2^4}}\lvert 1\rangle$.
Hence, $\lvert \varphi_1\rangle = \left(\lvert 0\rangle + e^{\frac{2\pi i x}{2^4}}\lvert 1\rangle\right)\otimes \lvert x_1x_2x_3\rangle$.
It is now easy to see via an inductive argument that $\lvert \varphi_2\rangle$ changes qubit $\lvert x_1\rangle$ to:
+ $\lvert 0\rangle + e^{\frac{2\pi i}{2^3}(4x_1 + 2x_2 + x_3)}\lvert 1\rangle$.
However, as $e^{2\pi ix_0} = 1$, we have $e^{\frac{2\pi i}{2^3}(4x_1 + 2x_2 + x_3)}=e^{\frac{2\pi i}{2^3}(8x_0 + 4x_1 + 2x_2 + x_3)}= e^{\frac{2\pi ix}{2^3}}$.
So, $\lvert \varphi_2\rangle = \left(\lvert 0\rangle + e^{\frac{2\pi i x}{2^4}}\lvert 1\rangle\right)\otimes \left(\lvert 0\rangle + e^{\frac{2\pi i x}{2^3}}\lvert 1\rangle\right)\otimes \lvert x_2x_3\rangle$.
Continuing, we can check that $\vert \varphi_4 \rangle$ gives us the reverse order of the qubits that we wanted, and hence we do a double swap to complete the circuit schematic of QFT.
## Writing QFT in Blueqat-SDK
```
# Install Blueqat-SDK if you haven't already!
# !pip install blueqat
# Import Modules
from blueqat import Circuit
import math
# Function to apply qft on a list of qubits of circuit
def apply_qft(circuit: Circuit(), qubits):
num_qubits = len(qubits)
for i in range(num_qubits):
circuit.h[qubits[i]]
for j in range(i+1, num_qubits):
circuit.cphase(math.pi/(2 ** (j-i)))[qubits[j],qubits[i]] # Apply gate CR_{j-i}(qubit j, qubit i)
# Reverse the order of qubits at the end
for i in range(int(num_qubits/2)):
circuit.swap(qubits[i],qubits[num_qubits-i-1])
```
### Testing on a circuit
```
n = 4 # number of qubits to apply QFT on
qc = Circuit()
qc.x[:] # Prepare the state |1111>
apply_qft(qc, range(n))
qc.run()
```
The QFT only changes phases of individual qubits, and hence cannot be physically observed via measurements. Below is a visual representation of the above statevector obtained from $\text{QFT}(|1111\rangle)$:

## Applications of QFT
QFT has wide applications in Algorithms design, and is the building block for many influential quantum algorithms such as Quantum Phase Estimation [[1]] , Shor's Algorithm [[2]], and the Hidden Subgroup Problem [[3]].
[1]: https://en.wikipedia.org/wiki/Quantum_phase_estimation_algorithm
[2]: https://en.wikipedia.org/wiki/Shor's_algorithm
[3]: https://en.wikipedia.org/wiki/Hidden_subgroup_problem
| github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from tpot import TPOTClassifier
from sklearn.datasets import load_digits
from sklearn.model_selection import train_test_split
import sklearn.metrics
import os
# Any results you write to the current directory are saved as output.
import timeit
pd.options.display.max_columns = 500
pd.options.display.width = 500
digits = load_digits()
X_train, X_test, y_train, y_test = train_test_split(digits.data, digits.target,
train_size=0.75, test_size=0.25, random_state=50)
tpot = TPOTClassifier(verbosity=3,
scoring="accuracy",
random_state=50,
n_jobs=-1,
generations=20,
periodic_checkpoint_folder="intermediate_algos",
population_size=60,
early_stop=10)
times = []
scores = []
winning_pipes = []
for x in range(1):
start_time = timeit.default_timer()
tpot.fit(X_train, y_train)
elapsed = timeit.default_timer() - start_time
times.append(elapsed)
winning_pipes.append(tpot.fitted_pipeline_)
scores.append(tpot.score(X_test, y_test))
tpot.export('tpot_mnist_pipeline1.py')
times = [time/60 for time in times]
print('Times:', times)
print('Scores:', scores)
print('Winning pipelines:', winning_pipes)
print('Times:', times)
print('Scores:', scores)
print('Winning pipelines:', winning_pipes)
import h2o
print(h2o.__version__)
from h2o.automl import H2OAutoML
h2o.init(max_mem_size='2G')
%%time
train = h2o.import_file(r"C:\Users\HP\Desktop\Oreilly\digit-recognizer/train.csv")
test = h2o.import_file(r"C:\Users\HP\Desktop\Oreilly\digit-recognizer/test.csv")
train.head()
x = train.columns[1:]
y = 'label'
# For binary classification, response should be a factor
train[y] = train[y].asfactor()
aml = H2OAutoML(max_models=30, seed=45, max_runtime_secs=28800)
aml.train(x=x, y=y, training_frame=train)
# View the AutoML Leaderboard
lb = aml.leaderboard
lb.head(rows=lb.nrows) # Print all rows instead of default (10 rows)
aml.leader
preds = aml.predict(test)
preds['p1'].as_data_frame().values.shape
preds
from tensorflow.keras.datasets import mnist
import autokeras as ak
# Prepare the dataset.
(x_train, y_train), (x_test, y_test) = mnist.load_data()
print(x_train.shape) # (60000, 28, 28)
print(y_train.shape) # (60000,)
print(y_train[:3]) # array([7, 2, 1], dtype=uint8)
# Initialize the ImageClassifier.
clf = ak.ImageClassifier(max_trials=1)
clf.fit(x_train, y_train, epochs=1)
# Evaluate on the testing data.
print("Accuracy: {accuracy}".format(accuracy=clf.evaluate(x_test, y_test)))
import torch
import torch.nn as nn
import torch.nn.functional as F
import torchvision
import torchvision.transforms as transforms
from tqdm.auto import tqdm
transform = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.1307,), (0.3081,))
])
trainset = torchvision.datasets.MNIST(root='./data', train=True, download=True, transform=transform)
testset = torchvision.datasets.MNIST(root='./data', train=False, download=True, transform=transform)
def train_mnist(args, reporter):
# get variables from args
lr = args.lr
wd = args.wd
epochs = args.epochs
net = args.net
print('lr: {}, wd: {}'.format(lr, wd))
device = 'cuda' if torch.cuda.is_available() else 'cpu'
# Model
net = net.to(device)
if device == 'cuda':
net = nn.DataParallel(net)
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.SGD(net.parameters(), lr=args.lr, momentum=0.9, weight_decay=wd)
# datasets and dataloaders
trainset = torchvision.datasets.MNIST(root='./data', train=True, download=False, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=128, shuffle=True, num_workers=2)
testset = torchvision.datasets.MNIST(root='./data', train=False, download=False, transform=transform)
testloader = torch.utils.data.DataLoader(testset, batch_size=128, shuffle=False, num_workers=2)
# Training
def train(epoch):
net.train()
train_loss, correct, total = 0, 0, 0
for batch_idx, (inputs, targets) in enumerate(trainloader):
inputs, targets = inputs.to(device), targets.to(device)
optimizer.zero_grad()
outputs = net(inputs)
loss = criterion(outputs, targets)
loss.backward()
optimizer.step()
def test(epoch):
net.eval()
test_loss, correct, total = 0, 0, 0
with torch.no_grad():
for batch_idx, (inputs, targets) in enumerate(testloader):
inputs, targets = inputs.to(device), targets.to(device)
outputs = net(inputs)
loss = criterion(outputs, targets)
test_loss += loss.item()
_, predicted = outputs.max(1)
total += targets.size(0)
correct += predicted.eq(targets).sum().item()
acc = 100.*correct/total
# 'epoch' reports the number of epochs done
reporter(epoch=epoch+1, accuracy=acc)
for epoch in tqdm(range(0, epochs)):
train(epoch)
test(epoch)
import autogluon.core as ag
@ag.obj(
hidden_conv=ag.space.Int(6, 12),
hidden_fc=ag.space.Categorical(80, 120, 160),
)
class Net(nn.Module):
def __init__(self, hidden_conv, hidden_fc):
super().__init__()
self.conv1 = nn.Conv2d(1, hidden_conv, 5)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(hidden_conv, 16, 5)
self.fc1 = nn.Linear(16 * 4 * 4, hidden_fc)
self.fc2 = nn.Linear(hidden_fc, 84)
self.fc3 = nn.Linear(84, 10)
def forward(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
x = x.view(-1, 16 * 4 * 4)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.fc3(x)
return x
@ag.args(
lr = ag.space.Real(0.01, 0.2, log=True),
wd = ag.space.Real(1e-4, 5e-4, log=True),
net = Net(),
epochs=1,
)
def ag_train_mnist(args, reporter):
return train_mnist(args, reporter)
myscheduler = ag.scheduler.FIFOScheduler(
ag_train_mnist,
resource={'num_cpus': 4, 'num_gpus': 0},
num_trials=1,
time_attr='epoch',
reward_attr='accuracy')
print(myscheduler)
myscheduler.run()
myscheduler.join_jobs()
myscheduler.get_training_curves(plot=True,use_legend=False)
print('The Best Configuration and Accuracy are: {}, {}'.format(myscheduler.get_best_config(), myscheduler.get_best_reward()))
myscheduler = ag.scheduler.FIFOScheduler(
ag_train_mnist,
resource={'num_cpus': 4, 'num_gpus': 0},
searcher='bayesopt',
num_trials=1,
time_attr='epoch',
reward_attr='accuracy')
print(myscheduler)
myscheduler.run()
myscheduler.join_jobs()
myscheduler = ag.scheduler.HyperbandScheduler(
ag_train_mnist,
resource={'num_cpus': 4, 'num_gpus': 1},
searcher='bayesopt',
num_trials=2,
time_attr='epoch',
reward_attr='accuracy',
grace_period=1,
reduction_factor=3,
brackets=1)
print(myscheduler)
myscheduler.run()
myscheduler.join_jobs()
```
| github_jupyter |
# Exercise 10
- Fraud Detection Dataset from Microsoft Azure: [data](http://gallery.cortanaintelligence.com/Experiment/8e9fe4e03b8b4c65b9ca947c72b8e463)
Fraud detection is one of the earliest industrial applications of data mining and machine learning. Fraud detection is typically handled as a binary classification problem, but the class population is unbalanced because instances of fraud are usually very rare compared to the overall volume of transactions. Moreover, when fraudulent transactions are discovered, the business typically takes measures to block the accounts from transacting to prevent further losses.
```
import pandas as pd
import zipfile
with zipfile.ZipFile('../datasets/fraud_detection.csv.zip', 'r') as z:
f = z.open('15_fraud_detection.csv')
data = pd.io.parsers.read_table(f, index_col=0, sep=',')
data.head()
X = data.drop(['Label'], axis=1)
y = data['Label']
y.value_counts(normalize=True)
```
# Exercice 10.1
Estimate a Logistic Regression, GaussianNB, K-nearest neighbors and a Decision Tree **Classifiers**
Evaluate using the following metrics:
* Accuracy
* F1-Score
* F_Beta-Score (Beta=10)
Comment about the results
Combine the classifiers and comment
```
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.neighbors import KNeighborsClassifier
models = {'lr': LogisticRegression(),
'dt': DecisionTreeClassifier(),
'nb': GaussianNB(),
'nn': KNeighborsClassifier()}
from sklearn.cross_validation import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=2)
# Train all the models
for model in models.keys():
models[model].fit(X_train, y_train)
# predict test for each model
y_pred = pd.DataFrame(index=X_test.index, columns=models.keys())
for model in models.keys():
y_pred[model] = models[model].predict(X_test)
y_pred
y_pred_ensemble1 = (y_pred.mean(axis=1) > 0.5).astype(int)
y_pred_ensemble1.mean()
from sklearn.metrics import accuracy_score, f1_score, recall_score, precision_score
stats = {'acc': accuracy_score,
'f1': f1_score,
'rec': recall_score,
'pre': precision_score}
res = pd.DataFrame(index=models.keys(), columns=stats.keys())
for model in models.keys():
for stat in stats.keys():
res.loc[model, stat] = stats[stat](y_test, y_pred[model])
res
res.loc['ensemble1'] = 0
for stat in stats.keys():
res.loc['ensemble1', stat] = stats[stat](y_test, y_pred_ensemble1)
res
```
# Exercice 10.2
Apply random-undersampling with a target percentage of 0.5
how does the results change
# Exercice 10.3
For each model estimate a BaggingClassifier of 100 models using the under sampled datasets
# Exercice 10.4
Using the under-sampled dataset
Evaluate a RandomForestClassifier and compare the results
change n_estimators=100, what happened
| github_jupyter |
# An Introduction to Python
In this lesson we will learn how to work with arthritis inflammation datasets in Python. However,
before we discuss how to deal with many data points, let's learn how to work with
single data values.
## Variables
Any Python interpreter can be used as a calculator:
```
3 + 5 * 4
```
This is great but not very interesting. To do anything useful with data, we need
to assign its value to a _variable_. In Python, we can assign a value to a
variable, using the equals sign `=`. For example, to assign value `60` to a
variable `weight_kg`, we would execute:
```
weight_kg = 60
```
From now on, whenever we use `weight_kg`, Python will substitute the value we assigned to
it. In essence, **a variable is just a name for a value**.
In Python, variable names:
- can include letters, digits, and underscores
- cannot start with a digit
- are case sensitive.
This means that, for example:
- `weight0` is a valid variable name, whereas `0weight` is not
- `weight` and `Weight` are different variables
## Types of data
Python knows various types of data. Three common ones are:
* integer numbers
* floating point numbers, and
* strings.
In the example above, variable `weight_kg` has an integer value of `60`.
To create a variable with a floating point value, we can execute:
```
weight_kg = 60.0
```
And to create a string we simply have to add single or double quotes around some text, for example:
```
weight_kg_text = 'weight in kilograms:'
```
## Using Variables in Python
To display the value of a variable to the screen in Python, we can use the `print` function:
```
print(weight_kg)
```
We can display multiple things at once using only one `print` command:
```
print(weight_kg_text, weight_kg)
```
Moreover, we can do arithmetics with variables right inside the `print` function:
```
print('weight in pounds:', 2.2 * weight_kg)
```
The above command, however, did not change the value of `weight_kg`:
```
print(weight_kg)
```
To change the value of the `weight_kg` variable, we have to
**assign** `weight_kg` a new value using the equals `=` sign:
```
weight_kg = 65.0
print('weight in kilograms is now:', weight_kg)
```
## Variables as Sticky Notes
A variable is analogous to a sticky note with a name written on it:
assigning a value to a variable is like putting that sticky note on a particular value.

This means that assigning a value to one variable does **not** change the values of other variables.
For example, let's store the subject's weight in pounds in its own variable:
```
# There are 2.2 pounds per kilogram
weight_lb = 2.2 * weight_kg
print(weight_kg_text, weight_kg, 'and in pounds:', weight_lb)
```

Let's now change `weight_kg`:
```
weight_kg = 100.0
print('weight in kilograms is now:', weight_kg, 'and weight in pounds is still:', weight_lb)
```

Since `weight_lb` doesn't remember where its value came from,
it isn't automatically updated when `weight_kg` changes.
Words are useful, but what's more useful are the sentences and stories we build
with them. Similarly, while a lot of powerful, general tools are built into
languages like Python, specialized tools built up from these basic units live in
libraries that can be called upon when needed.
## Loading data into Python
In order to load our inflammation data, we need to access
(import in Python terminology) a library called
[NumPy](http://docs.scipy.org/doc/numpy/ "NumPy Documentation"). In general you should use this
library if you want to do fancy things with numbers, especially if you have matrices or arrays. We
can import NumPy using:
```
import numpy
```
Importing a library is like getting a piece of lab equipment out of a storage locker and setting it
up on the bench. Libraries provide additional functionality to the basic Python package, much like
a new piece of equipment adds functionality to a lab space. Just like in the lab, importing too
many libraries can sometimes complicate and slow down your programs - so we only import what we
need for each program. Once we've imported the library, we can ask the library to read our data
file for us:
```
numpy.loadtxt(fname='inflammation-01.csv', delimiter=',')
```
The expression `numpy.loadtxt(...)` is a function call that asks Python to run
the function `loadtxt` which belongs to the `numpy` library. This dotted
notation is used everywhere in Python: the thing that appears before the dot
contains the thing that appears after.
As an example, John Smith is the John that belongs to the Smith family. We could
use the dot notation to write his name `smith.john`, just as `loadtxt` is a
function that belongs to the `numpy` library.
`numpy.loadtxt` has two parameters: the name of the file we want to read and the
delimiter that separates values on a line. These both need to be character
strings (or strings for short), so we put them in quotes.
Since we haven't told it to do anything else with the function's output,
the notebook displays it.
In this case,
that output is the data we just loaded.
By default,
only a few rows and columns are shown
(with `...` to omit elements when displaying big arrays). To save space, Python
displays numbers as `1.` instead of `1.0`
when there's nothing interesting after the decimal point.
Our call to `numpy.loadtxt` read our file
but didn't save the data in memory.
To do that,
we need to assign the array to a variable. Just as we can assign a single value to a variable, we
can also assign an array of values to a variable using the same syntax. Let's re-run
`numpy.loadtxt` and save the returned data:
```
data = numpy.loadtxt(fname='inflammation-01.csv', delimiter=',')
```
If we want to check that the data have been loaded,
we can print the variable's value:
```
print(data)
```
Now that the data are in memory,
we can manipulate them.
First,
let's ask what type of thing `data` refers to:
```
print(type(data))
```
The output tells us that `data` currently refers to
an N-dimensional array, the functionality for which is provided by the NumPy library.
These data correspond to arthritis patients' inflammation.
The rows are the individual patients, and the columns
are their daily inflammation measurements.
<section class="callout panel panel-warning">
<div class="panel-heading">
<h2><span class="fa fa-thumb-tack"></span> Data Type</h2>
</div>
<div class="panel-body">
<p>A Numpy array contains one or more elements
of the same type. The <code>type</code> function will only tell you that
a variable is a NumPy array but won't tell you the type of
thing inside the array.
We can find out the type
of the data contained in the NumPy array.</p>
<div class="codehilite"><pre><span></span><span class="k">print</span><span class="p">(</span><span class="n">data</span><span class="o">.</span><span class="n">dtype</span><span class="p">)</span>
</pre></div>
<div class="codehilite"><pre><span></span>dtype('float64')
</pre></div>
<p>This tells us that the NumPy array's elements are
floating-point numbers.</p>
</div>
</section>
With the following command, we can see the array's shape:
```
print(data.shape)
```
The output tells us that the `data` array variable contains 60 rows and 40 columns. When we
created the variable `data` to store our arthritis data, we didn't just create the array; we also
created information about the array, called members or
attributes. This extra information describes `data` in the same way an adjective describes a noun.
`data.shape` is an attribute of `data` which describes the dimensions of `data`. We use the same
dotted notation for the attributes of variables that we use for the functions in libraries because
they have the same part-and-whole relationship.
If we want to get a single number from the array, we must provide an
index in square brackets after the variable name, just as we
do in math when referring to an element of a matrix. Our inflammation data has two dimensions, so
we will need to use two indices to refer to one specific value:
```
print('first value in data:', data[0, 0])
print('middle value in data:', data[30, 20])
print(data[0:4, 0:10])
```
The expression `data[30, 20]` accesses the element at row 30, column 20. While this expression may
not surprise you,
`data[0, 0]` might.
Programming languages like Fortran, MATLAB and R start counting at 1
because that's what human beings have done for thousands of years.
Languages in the C family (including C++, Java, Perl, and Python) count from 0
because it represents an offset from the first value in the array (the second
value is offset by one index from the first value). This is closer to the way
that computers represent arrays (if you are interested in the historical
reasons behind counting indices from zero, you can read
[Mike Hoye's blog post](http://exple.tive.org/blarg/2013/10/22/citation-needed/)).
As a result,
if we have an M×N array in Python,
its indices go from 0 to M-1 on the first axis
and 0 to N-1 on the second.
It takes a bit of getting used to,
but one way to remember the rule is that
the index is how many steps we have to take from the start to get the item we want.

## In the Corner
What may also surprise you is that when Python displays an array,
it shows the element with index `[0, 0]` in the upper left corner
rather than the lower left.
This is consistent with the way mathematicians draw matrices
but different from the Cartesian coordinates.
The indices are (row, column) instead of (column, row) for the same reason,
which can be confusing when plotting data.
## Slicing data
An index like `[30, 20]` selects a single element of an array,
but we can select whole sections as well.
For example,
we can select the first ten days (columns) of values
for the first four patients (rows) like this:
The slice `0:4` means, "Start at index 0 and go up to, but not
including, index 4."Again, the up-to-but-not-including takes a bit of getting used to, but the
rule is that the difference between the upper and lower bounds is the number of values in the slice.
We don't have to start slices at 0:
```
print(data[5:10, 0:10])
```
We also don't have to include the upper and lower bound on the slice. If we don't include the lower
bound, Python uses 0 by default; if we don't include the upper, the slice runs to the end of the
axis, and if we don't include either (i.e., if we just use ':' on its own), the slice includes
everything:
```
small = data[:3, 36:]
print('small is:')
print(small)
```
The above example selects rows 0 through 2 and columns 36 through to the end of the array.
Arrays also know how to perform common mathematical operations on their values. The simplest
operations with data are arithmetic: addition, subtraction, multiplication, and division. When you
do such operations on arrays, the operation is done element-by-element. Thus:
```
doubledata = data * 2.0
```
will create a new array `doubledata`
each element of which is twice the value of the corresponding element in `data`:
```
print('original:')
print(data[:3, 36:])
print('doubledata:')
print(doubledata[:3, 36:])
```
If, instead of taking an array and doing arithmetic with a single value (as above), you did the
arithmetic operation with another array of the same shape, the operation will be done on
corresponding elements of the two arrays. Thus:
```
tripledata = doubledata + data
```
will give you an array where `tripledata[0,0]` will equal `doubledata[0,0]` plus `data[0,0]`,
and so on for all other elements of the arrays.
```
print('tripledata:')
print(tripledata[:3, 36:])
```
Often, we want to do more than add, subtract, multiply, and divide array elements. NumPy knows how
to do more complex operations, too. If we want to find the average inflammation for all patients on
all days, for example, we can ask NumPy to compute `data`'s mean value:
```
print(numpy.mean(data))
```
`mean` is a function that takes an array as an argument.
<section class="callout panel panel-warning">
<div class="panel-heading">
<h2><span class="fa fa-thumb-tack"></span> Not All Functions Have Input</h2>
</div>
<div class="panel-body">
<p>Generally, a function uses inputs to produce outputs.
However, some functions produce outputs without
needing any input. For example, checking the current time
doesn't require any input.</p>
<div class="codehilite"><pre><span></span><span class="kn">import</span> <span class="nn">time</span>
<span class="k">print</span><span class="p">(</span><span class="n">time</span><span class="o">.</span><span class="n">ctime</span><span class="p">())</span>
</pre></div>
<div class="codehilite"><pre><span></span>'Sat Mar 26 13:07:33 2016'
</pre></div>
<p>For functions that don't take in any arguments,
we still need parentheses (<code>()</code>)
to tell Python to go and do something for us.</p>
</div>
</section>
NumPy has lots of useful functions that take an array as input.
Let's use three of those functions to get some descriptive values about the dataset.
We'll also use multiple assignment,
a convenient Python feature that will enable us to do this all in one line.
```
maxval, minval, stdval = numpy.max(data), numpy.min(data), numpy.std(data)
print('maximum inflammation:', maxval)
print('minimum inflammation:', minval)
print('standard deviation:', stdval)
```
Here we've assigned the return value from `numpy.max(data)` to the variable `maxval`, the value
from `numpy.min(data)` to `minval`, and so on.
<section class="callout panel panel-warning">
<div class="panel-heading">
<h2><span class="fa fa-thumb-tack"></span> Mystery Functions in IPython</h2>
</div>
<div class="panel-body">
<p>How did we know what functions NumPy has and how to use them?
If you are working in the IPython/Jupyter Notebook, there is an easy way to find out.
If you type the name of something followed by a dot, then you can use tab completion
(e.g. type <code>numpy.</code> and then press tab)
to see a list of all functions and attributes that you can use. After selecting one, you
can also add a question mark (e.g. <code>numpy.cumprod?</code>), and IPython will return an
explanation of the method! This is the same as doing <code>help(numpy.cumprod)</code>.</p>
</div>
</section>
When analyzing data, though,
we often want to look at variations in statistical values,
such as the maximum inflammation per patient
or the average inflammation per day.
One way to do this is to create a new temporary array of the data we want,
then ask it to do the calculation:
```
patient_0 = data[0, :] # 0 on the first axis (rows), everything on the second (columns)
print('maximum inflammation for patient 0:', patient_0.max())
```
Everything in a line of code following the '#' symbol is a
comment that is ignored by Python.
Comments allow programmers to leave explanatory notes for other
programmers or their future selves.
We don't actually need to store the row in a variable of its own.
Instead, we can combine the selection and the function call:
```
print('maximum inflammation for patient 2:', numpy.max(data[2, :]))
```
What if we need the maximum inflammation for each patient over all days (as in the
next diagram on the left) or the average for each day (as in the
diagram on the right)? As the diagram below shows, we want to perform the
operation across an axis:

To support this functionality,
most array functions allow us to specify the axis we want to work on.
If we ask for the average across axis 0 (rows in our 2D example),
we get:
```
print(numpy.mean(data, axis=0))
```
As a quick check,
we can ask this array what its shape is:
```
print(numpy.mean(data, axis=0).shape)
```
The expression `(40,)` tells us we have an N×1 vector,
so this is the average inflammation per day for all patients.
If we average across axis 1 (columns in our 2D example), we get:
```
print(numpy.mean(data, axis=1))
```
which is the average inflammation per patient across all days.
## Visualizing data
The mathematician Richard Hamming once said, "The purpose of computing is insight, not numbers," and
the best way to develop insight is often to visualize data. Visualization deserves an entire
lecture of its own, but we can explore a few features of Python's `matplotlib` library here. While
there is no official plotting library, `matplotlib` is the _de facto_ standard. First, we will
import the `pyplot` module from `matplotlib` and use two of its functions to create and display a
heat map of our data:
```
%matplotlib inline
import matplotlib.pyplot
image = matplotlib.pyplot.imshow(data)
matplotlib.pyplot.show()
```
Blue pixels in this heat map represent low values, while yellow pixels represent high values. As we
can see, inflammation rises and falls over a 40-day period.
```
ave_inflammation = numpy.mean(data, axis=0)
ave_plot = matplotlib.pyplot.plot(ave_inflammation)
matplotlib.pyplot.show()
```
Here, we have put the average per day across all patients in the variable `ave_inflammation`, then
asked `matplotlib.pyplot` to create and display a line graph of those values. The result is a
roughly linear rise and fall, which is suspicious: we might instead expect a sharper rise and slower
fall. Let's have a look at two other statistics:
```
max_plot = matplotlib.pyplot.plot(numpy.max(data, axis=0))
matplotlib.pyplot.show()
min_plot = matplotlib.pyplot.plot(numpy.min(data, axis=0))
matplotlib.pyplot.show()
```
The maximum value rises and falls smoothly, while the minimum seems to be a step function. Neither
trend seems particularly likely, so either there's a mistake in our calculations or something is
wrong with our data. This insight would have been difficult to reach by examining the numbers
themselves without visualization tools.
<section class="callout panel panel-warning">
<div class="panel-heading">
<h2><span class="fa fa-thumb-tack"></span> Scientists Dislike Typing</h2>
</div>
<div class="panel-body">
<p>We have just been importing NumPy and matplotlib using <code>import numpy</code> and <code>import matplotlib.pyplot</code>.</p>
<p>From here on we are going to shorten these imports, you can either adopt this or continue using the long version. The reason we are going to start using the shortened versions is that that is the way that the majority of open source scientific python libraries use these imports, so we want to get you used to them now.</p>
<p>When working with other people, it is important to agree on a convention of how common libraries
are imported.</p>
</div>
</section>
```
import numpy as np
import matplotlib.pyplot as plt
```
### Grouping plots
You can group similar plots in a single figure using subplots.
This script below uses a number of new commands. The function `plt.figure()`
creates a space into which we will place all of our plots. The parameter `figsize`
tells Python how big to make this space. Each subplot is placed into the figure using
its `add_subplot` method. The `add_subplot` method takes 3
parameters. The first denotes how many total rows of subplots there are, the second parameter
refers to the total number of subplot columns, and the final parameter denotes which subplot
your variable is referencing (left-to-right, top-to-bottom). Each subplot is stored in a
different variable (`axes1`, `axes2`, `axes3`). Once a subplot is created, the axes can
be titled using the `set_xlabel()` command (or `set_ylabel()`).
Here are our three plots side by side:
```
data = np.loadtxt(fname='inflammation-01.csv', delimiter=',')
fig = plt.figure(figsize=(10.0, 3.0))
axes1 = fig.add_subplot(1, 3, 1)
axes2 = fig.add_subplot(1, 3, 2)
axes3 = fig.add_subplot(1, 3, 3)
axes1.set_ylabel('average')
axes1.plot(numpy.mean(data, axis=0))
axes2.set_ylabel('max')
axes2.plot(numpy.max(data, axis=0))
axes3.set_ylabel('min')
axes3.plot(numpy.min(data, axis=0))
fig.tight_layout()
plt.show()
```
The call to `loadtxt` reads our data,
and the rest of the program tells the plotting library
how large we want the figure to be,
that we're creating three subplots,
what to draw for each one,
and that we want a tight layout.
(If we leave out that call to `fig.tight_layout()`,
the graphs will actually be squeezed together more closely.)
<section class="challenge panel panel-success">
<div class="panel-heading">
<h2><span class="fa fa-pencil"></span> Challenge: Check Your Understanding</h2>
</div>
<div class="panel-body">
<p>What values do the variables <code>mass</code> and <code>age</code> have after each statement in the following program?
Test your answers by executing the commands.</p>
<div class="codehilite"><pre><span></span><span class="n">mass</span> <span class="o">=</span> <span class="mf">47.5</span>
<span class="n">age</span> <span class="o">=</span> <span class="mi">122</span>
<span class="n">mass</span> <span class="o">=</span> <span class="n">mass</span> <span class="o">*</span> <span class="mf">2.0</span>
<span class="n">age</span> <span class="o">=</span> <span class="n">age</span> <span class="o">-</span> <span class="mi">20</span>
</pre></div>
</div>
</section>
<section class="challenge panel panel-success">
<div class="panel-heading">
<h2><span class="fa fa-pencil"></span> Challenge: Sorting Out References</h2>
</div>
<div class="panel-body">
<p>What does the following program print out?</p>
<div class="codehilite"><pre><span></span>first, second = 'Grace', 'Hopper'
third, fourth = second, first
print(third, fourth)
</pre></div>
</div>
</section>
<section class="solution panel panel-primary">
<div class="panel-heading">
<h2><span class="fa fa-eye"></span> Solution</h2>
</div>
<div class="panel-body">
<div class="codehilite"><pre><span></span>Hopper Grace
</pre></div>
</div>
</section>
<section class="challenge panel panel-success">
<div class="panel-heading">
<h2><span class="fa fa-pencil"></span> Challenge: Slicing Strings</h2>
</div>
<div class="panel-body">
<p>A section of an array is called a slice.
We can take slices of character strings as well:</p>
<div class="codehilite"><pre><span></span><span class="n">element</span> <span class="o">=</span> <span class="s1">'oxygen'</span>
<span class="k">print</span><span class="p">(</span><span class="s1">'first three characters:'</span><span class="p">,</span> <span class="n">element</span><span class="p">[</span><span class="mi">0</span><span class="p">:</span><span class="mi">3</span><span class="p">])</span>
<span class="k">print</span><span class="p">(</span><span class="s1">'last three characters:'</span><span class="p">,</span> <span class="n">element</span><span class="p">[</span><span class="mi">3</span><span class="p">:</span><span class="mi">6</span><span class="p">])</span>
</pre></div>
<div class="codehilite"><pre><span></span>first three characters: oxy
last three characters: gen
</pre></div>
<p>What is the value of <code>element[:4]</code>?
What about <code>element[4:]</code>?
Or <code>element[:]</code>?</p>
</div>
</section>
<section class="solution panel panel-primary">
<div class="panel-heading">
<h2><span class="fa fa-eye"></span> Solution</h2>
</div>
<div class="panel-body">
<div class="codehilite"><pre><span></span>oxyg
en
oxygen
</pre></div>
</div>
</section>
<section class="challenge panel panel-success">
<div class="panel-heading">
<h2><span class="fa fa-pencil"></span> Challenge:</h2>
</div>
<div class="panel-body">
<p>What is <code>element[-1]</code>?
What is <code>element[-2]</code>?</p>
</div>
</section>
<section class="solution panel panel-primary">
<div class="panel-heading">
<h2><span class="fa fa-eye"></span> Solution</h2>
</div>
<div class="panel-body">
<div class="codehilite"><pre><span></span>n
e
</pre></div>
</div>
</section>
<section class="challenge panel panel-success">
<div class="panel-heading">
<h2><span class="fa fa-pencil"></span> Challenge:</h2>
</div>
<div class="panel-body">
<p>Given those answers, explain what <code>element[1:-1]</code> does.</p>
</div>
</section>
<section class="solution panel panel-primary">
<div class="panel-heading">
<h2><span class="fa fa-eye"></span> Solution</h2>
</div>
<div class="panel-body">
<p>Creates a substring from index 1 up to (not including) the final index,
effectively removing the first and last letters from 'oxygen'</p>
</div>
</section>
<section class="challenge panel panel-success">
<div class="panel-heading">
<h2><span class="fa fa-pencil"></span> Challenge: Thin Slices</h2>
</div>
<div class="panel-body">
<p>The expression <code>element[3:3]</code> produces an empty string,
i.e., a string that contains no characters.
If <code>data</code> holds our array of patient data,
what does <code>data[3:3, 4:4]</code> produce?
What about <code>data[3:3, :]</code>?</p>
</div>
</section>
<section class="solution panel panel-primary">
<div class="panel-heading">
<h2><span class="fa fa-eye"></span> Solution</h2>
</div>
<div class="panel-body">
<div class="codehilite"><pre><span></span>[]
[]
</pre></div>
</div>
</section>
<section class="challenge panel panel-success">
<div class="panel-heading">
<h2><span class="fa fa-pencil"></span> Challenge: Plot Scaling</h2>
</div>
<div class="panel-body">
<p>Why do all of our plots stop just short of the upper end of our graph?</p>
</div>
</section>
<section class="solution panel panel-primary">
<div class="panel-heading">
<h2><span class="fa fa-eye"></span> Solution</h2>
</div>
<div class="panel-body">
<p>Because matplotlib normally sets x and y axes limits to the min and max of our data
(depending on data range)</p>
<p>If we want to change this, we can use the <code>set_ylim(min, max)</code> method of each 'axes',
for example:</p>
<div class="codehilite"><pre><span></span><span class="n">axes3</span><span class="o">.</span><span class="n">set_ylim</span><span class="p">(</span><span class="mi">0</span><span class="p">,</span><span class="mi">6</span><span class="p">)</span>
</pre></div>
</div>
</section>
<section class="challenge panel panel-success">
<div class="panel-heading">
<h2><span class="fa fa-pencil"></span> Challenge:</h2>
</div>
<div class="panel-body">
<p>Update your plotting code to automatically set a more appropriate scale.
(Hint: you can make use of the <code>max</code> and <code>min</code> methods to help.)</p>
</div>
</section>
<section class="solution panel panel-primary">
<div class="panel-heading">
<h2><span class="fa fa-eye"></span> Solution</h2>
</div>
<div class="panel-body">
<div class="codehilite"><pre><span></span><span class="c1"># One method</span>
<span class="n">axes3</span><span class="o">.</span><span class="n">set_ylabel</span><span class="p">(</span><span class="s1">'min'</span><span class="p">)</span>
<span class="n">axes3</span><span class="o">.</span><span class="n">plot</span><span class="p">(</span><span class="n">numpy</span><span class="o">.</span><span class="n">min</span><span class="p">(</span><span class="n">data</span><span class="p">,</span> <span class="n">axis</span><span class="o">=</span><span class="mi">0</span><span class="p">))</span>
<span class="n">axes3</span><span class="o">.</span><span class="n">set_ylim</span><span class="p">(</span><span class="mi">0</span><span class="p">,</span><span class="mi">6</span><span class="p">)</span>
</pre></div>
<div class="codehilite"><pre><span></span><span class="c1"># A more automated approach</span>
<span class="n">min_data</span> <span class="o">=</span> <span class="n">numpy</span><span class="o">.</span><span class="n">min</span><span class="p">(</span><span class="n">data</span><span class="p">,</span> <span class="n">axis</span><span class="o">=</span><span class="mi">0</span><span class="p">)</span>
<span class="n">axes3</span><span class="o">.</span><span class="n">set_ylabel</span><span class="p">(</span><span class="s1">'min'</span><span class="p">)</span>
<span class="n">axes3</span><span class="o">.</span><span class="n">plot</span><span class="p">(</span><span class="n">min_data</span><span class="p">)</span>
<span class="n">axes3</span><span class="o">.</span><span class="n">set_ylim</span><span class="p">(</span><span class="n">numpy</span><span class="o">.</span><span class="n">min</span><span class="p">(</span><span class="n">min_data</span><span class="p">),</span> <span class="n">numpy</span><span class="o">.</span><span class="n">max</span><span class="p">(</span><span class="n">min_data</span><span class="p">)</span> <span class="o">*</span> <span class="mf">1.1</span><span class="p">)</span>
</pre></div>
</div>
</section>
<section class="challenge panel panel-success">
<div class="panel-heading">
<h2><span class="fa fa-pencil"></span> Challenge: Drawing Straight Lines</h2>
</div>
<div class="panel-body">
<p>In the center and right subplots above, we expect all lines to look like step functions because
non-integer value are not realistic for the minimum and maximum values. However, you can see
that the lines are not always vertical or horizontal, and in particular the step function
in the subplot on the right looks slanted. Why is this?</p>
</div>
</section>
<section class="solution panel panel-primary">
<div class="panel-heading">
<h2><span class="fa fa-eye"></span> Solution</h2>
</div>
<div class="panel-body">
<p>Because matplotlib interpolates (draws a straight line) between the points.
One way to do avoid this is to use the Matplotlib <code>drawstyle</code> option:</p>
</div>
</section>
```
import numpy as np
import matplotlib.pyplot as plt
data = numpy.loadtxt(fname='inflammation-01.csv', delimiter=',')
fig = plt.figure(figsize=(10.0, 3.0))
axes1 = fig.add_subplot(1, 3, 1)
axes2 = fig.add_subplot(1, 3, 2)
axes3 = fig.add_subplot(1, 3, 3)
axes1.set_ylabel('average')
axes1.plot(numpy.mean(data, axis=0), drawstyle='steps-mid')
axes2.set_ylabel('max')
axes2.plot(numpy.max(data, axis=0), drawstyle='steps-mid')
axes3.set_ylabel('min')
axes3.plot(numpy.min(data, axis=0), drawstyle='steps-mid')
fig.tight_layout()
plt.show()
```
<section class="challenge panel panel-success">
<div class="panel-heading">
<h2><span class="fa fa-pencil"></span> Challenge: Make Your Own Plot</h2>
</div>
<div class="panel-body">
<p>Create a plot showing the standard deviation (<code>numpy.std</code>)
of the inflammation data for each day across all patients.</p>
</div>
</section>
<section class="solution panel panel-primary">
<div class="panel-heading">
<h2><span class="fa fa-eye"></span> Solution</h2>
</div>
</section>
```
std_plot = plt.plot(numpy.std(data, axis=0))
plt.show()
```
<section class="challenge panel panel-success">
<div class="panel-heading">
<h2><span class="fa fa-pencil"></span> Challenge: Moving Plots Around</h2>
</div>
<div class="panel-body">
<p>Modify the program to display the three plots on top of one another
instead of side by side.</p>
</div>
</section>
<section class="solution panel panel-primary">
<div class="panel-heading">
<h2><span class="fa fa-eye"></span> Solution</h2>
</div>
</section>
```
import numpy as np
import matplotlib.pyplot as plt
data = numpy.loadtxt(fname='inflammation-01.csv', delimiter=',')
# change figsize (swap width and height)
fig = plt.figure(figsize=(3.0, 10.0))
# change add_subplot (swap first two parameters)
axes1 = fig.add_subplot(3, 1, 1)
axes2 = fig.add_subplot(3, 1, 2)
axes3 = fig.add_subplot(3, 1, 3)
axes1.set_ylabel('average')
axes1.plot(numpy.mean(data, axis=0))
axes2.set_ylabel('max')
axes2.plot(numpy.max(data, axis=0))
axes3.set_ylabel('min')
axes3.plot(numpy.min(data, axis=0))
fig.tight_layout()
plt.show()
```
## Stacking Arrays
Arrays can be concatenated and stacked on top of one another,
using NumPy's `vstack` and `hstack` functions for vertical and horizontal stacking, respectively.
```
import numpy as np
A = numpy.array([[1,2,3], [4,5,6], [7, 8, 9]])
print('A = ')
print(A)
B = numpy.hstack([A, A])
print('B = ')
print(B)
C = numpy.vstack([A, A])
print('C = ')
print(C)
```
<section class="challenge panel panel-success">
<div class="panel-heading">
<h2><span class="fa fa-pencil"></span> Challenge:</h2>
</div>
<div class="panel-body">
<p>Write some additional code that slices the first and last columns of <code>A</code>,
and stacks them into a 3x2 array.
Make sure to <code>print</code> the results to verify your solution.</p>
</div>
</section>
<section class="solution panel panel-primary">
<div class="panel-heading">
<h2><span class="fa fa-eye"></span> Solution</h2>
</div>
<div class="panel-body">
<p>A 'gotcha' with array indexing is that singleton dimensions
are dropped by default. That means <code>A[:, 0]</code> is a one dimensional
array, which won't stack as desired. To preserve singleton dimensions,
the index itself can be a slice or array. For example, <code>A[:, :1]</code> returns
a two dimensional array with one singleton dimension (i.e. a column
vector).</p>
</div>
</section>
```
D = np.hstack((A[:, :1], A[:, -1:]))
print('D = ')
print(D)
```
<section class="solution panel panel-primary">
<div class="panel-heading">
<h2><span class="fa fa-eye"></span> Solution</h2>
</div>
<div class="panel-body">
<p>An alternative way to achieve the same result is to use Numpy's
delete function to remove the second column of A.</p>
</div>
</section>
```
D = np.delete(A, 1, 1)
print('D = ')
print(D)
```
## Change In Inflammation
This patient data is _longitudinal_ in the sense that each row represents a
series of observations relating to one individual. This means that
the change in inflammation over time is a meaningful concept.
The `numpy.diff()` function takes a NumPy array and returns the
differences between two successive values along a specified axis. For
example, a NumPy array that looks like this:
```python
npdiff = numpy.array([ 0, 2, 5, 9, 14])
```
Calling `numpy.diff(npdiff)` would do the following calculations and
put the answers in another array.
```python
[ 2 - 0, 5 - 2, 9 - 5, 14 - 9 ]
```
```python
numpy.diff(npdiff)
```
```python
array([2, 3, 4, 5])
```
<section class="challenge panel panel-success">
<div class="panel-heading">
<h2><span class="fa fa-pencil"></span> Challenge:</h2>
</div>
<div class="panel-body">
<p>Which axis would it make sense to use this function along?</p>
</div>
</section>
<section class="solution panel panel-primary">
<div class="panel-heading">
<h2><span class="fa fa-eye"></span> Solution</h2>
</div>
<div class="panel-body">
<p>Since the row axis (0) is patients, it does not make sense to get the
difference between two arbitrary patients. The column axis (1) is in
days, so the difference is the change in inflammation -- a meaningful
concept.</p>
</div>
</section>
```
numpy.diff(data, axis=1)
```
<section class="challenge panel panel-success">
<div class="panel-heading">
<h2><span class="fa fa-pencil"></span> Challenge:</h2>
</div>
<div class="panel-body">
<p>If the shape of an individual data file is <code>(60, 40)</code> (60 rows and 40
columns), what would the shape of the array be after you run the <code>diff()</code>
function and why?</p>
</div>
</section>
<section class="solution panel panel-primary">
<div class="panel-heading">
<h2><span class="fa fa-eye"></span> Solution</h2>
</div>
<div class="panel-body">
<p>The shape will be <code>(60, 39)</code> because there is one fewer difference between
columns than there are columns in the data.</p>
</div>
</section>
<section class="challenge panel panel-success">
<div class="panel-heading">
<h2><span class="fa fa-pencil"></span> Challenge:</h2>
</div>
<div class="panel-body">
<p>How would you find the largest change in inflammation for each patient? Does
it matter if the change in inflammation is an increase or a decrease?</p>
</div>
</section>
<section class="solution panel panel-primary">
<div class="panel-heading">
<h2><span class="fa fa-eye"></span> Solution</h2>
</div>
<div class="panel-body">
<p>By using the <code>numpy.max()</code> function after you apply the <code>numpy.diff()</code>
function, you will get the largest difference between days.</p>
</div>
</section>
```
numpy.max(numpy.diff(data, axis=1), axis=1)
```
<section class="challenge panel panel-success">
<div class="panel-heading">
<h2><span class="fa fa-pencil"></span> Challenge:</h2>
</div>
<div class="panel-body">
<p>If inflammation values <em>decrease</em> along an axis, then the difference from
one element to the next will be negative. If
you are interested in the <strong>magnitude</strong> of the change and not the
direction, the <code>numpy.absolute()</code> function will provide that.
Notice the difference if you get the largest <em>absolute</em> difference
between readings.</p>
</div>
</section>
```
numpy.max(numpy.absolute(numpy.diff(data, axis=1)), axis=1)
```
---
The material in this notebook is derived from the Software Carpentry lessons
© [Software Carpentry](http://software-carpentry.org/) under the terms
of the [CC-BY 4.0](https://creativecommons.org/licenses/by/4.0/) license.
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.