text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
<a href="https://colab.research.google.com/github/ronva-h/technology_fundamentals/blob/main/C3%20Machine%20Learning%20I/LABS_PROJECT/Tech%20Fun%20C3%20P3%20Game%20AI%2C%20Statistics.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Technology Fundamentals Course 3, Project Part 3: Statistical Analysis
**Instructor**: Wesley Beckner
**Contact**: wesleybeckner@gmail.com
**Teaching Assistants**: Varsha Bang, Harsha Vardhan
**Contact**: vbang@uw.edu, harshav@uw.edu
<br>
---
<br>
Today we are going to perform statistical analysis on data generated from our tictactoe program!
<br>
---
<br>
<a name='x.0'></a>
## 2.0 Preparing Environment and Importing Data
[back to top](#top)
<a name='x.0.1'></a>
### 2.0.1 Import Packages
[back to top](#top)
```
import random
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
class TicTacToe:
# can preset winner and starting player
def __init__(self, winner='', start_player=''):
self.winner = winner
self.start_player = start_player
self.board = {1: ' ',
2: ' ',
3: ' ',
4: ' ',
5: ' ',
6: ' ',
7: ' ',
8: ' ',
9: ' ',}
self.win_patterns = [[1,2,3], [4,5,6], [7,8,9],
[1,4,7], [2,5,8], [3,6,9],
[1,5,9], [7,5,3]]
# the other functions are now passed self
def visualize_board(self):
print(
"|{}|{}|{}|\n|{}|{}|{}|\n|{}|{}|{}|\n".format(*self.board.values())
)
def check_winning(self):
for pattern in self.win_patterns:
values = [self.board[i] for i in pattern]
if values == ['X', 'X', 'X']:
self.winner = 'X' # we update the winner status
return "'X' Won!"
elif values == ['O', 'O', 'O']:
self.winner = 'O'
return "'O' Won!"
return ''
def check_stalemate(self):
if (' ' not in self.board.values()) and (self.check_winning() == ''):
self.winner = 'Stalemate'
return "It's a stalemate!"
class GameEngine(TicTacToe):
def __init__(self, setup='auto'):
super().__init__()
self.setup = setup
def setup_game(self):
if self.setup == 'user':
players = int(input("How many Players? (type 0, 1, or 2)"))
self.player_meta = {'first': {'label': 'X',
'type': 'ai'},
'second': {'label': 'O',
'type': 'human'}}
if players == 1:
first = input("who will go first? (X, (AI), or O (Player))")
if first == 'O':
self.player_meta = {'second': {'label': 'X',
'type': 'ai'},
'first': {'label': 'O',
'type': 'human'}}
elif players == 0:
first = random.choice(['X', 'O'])
if first == 'O':
self.player_meta = {'second': {'label': 'X',
'type': 'ai'},
'first': {'label': 'O',
'type': 'ai'}}
else:
self.player_meta = {'first': {'label': 'X',
'type': 'ai'},
'second': {'label': 'O',
'type': 'ai'}}
elif self.setup == 'auto':
first = random.choice(['X', 'O'])
if first == 'O':
self.start_player = 'O'
self.player_meta = {'second': {'label': 'X',
'type': 'ai'},
'first': {'label': 'O',
'type': 'ai'}}
else:
self.start_player = 'X'
self.player_meta = {'first': {'label': 'X',
'type': 'ai'},
'second': {'label': 'O',
'type': 'ai'}}
def play_game(self):
while True:
for player in ['first', 'second']:
self.visualize_board()
player_label = self.player_meta[player]['label']
player_type = self.player_meta[player]['type']
if player_type == 'human':
move = input("{}, what's your move?".format(player_label))
# we're going to allow the user to quit the game from the input line
if move in ['q', 'quit']:
self.winner = 'F'
print('quiting the game')
break
move = int(move)
if self.board[move] != ' ':
while True:
move = input("{}, that position is already taken! "\
"What's your move?".format(player))
move = int(move)
if self.board[move] != ' ':
continue
else:
break
else:
while True:
move = random.randint(1,9)
if self.board[move] != ' ':
continue
print('test')
else:
break
self.board[move] = player_label
# the winner varaible will now be check within the board object
self.check_winning()
self.check_stalemate()
if self.winner == '':
continue
elif self.winner == 'Stalemate':
print(self.check_stalemate())
self.visualize_board()
break
else:
print(self.check_winning())
self.visualize_board()
break
if self.winner != '':
return self
```
<a name='x.0.1'></a>
### 2.0.2 Load Dataset
[back to top](#top)
```
data = {}
for i in range(1000):
game = GameEngine()
game.setup_game()
board = game.play_game()
data['game {}'.format(i)] = {'board': board.board,
'winner': board.winner,
'starting player': board.start_player}
```
## 3.1 Clean Data
We will first need to organize the data into a parsable format.
### Q1
What is the object `data` and what does it contain?
* what are the keys of data?
* what are the keys of each game?
```
# inspect data below by grabbing the first key in data
# what are the three different keys within each game?
data['game 0']
```
### Q2
Using those keys, iterate through every `game` in `data` and append the board, the winner, and the starting player to separate lists. Call these lists: `boards, winners, and starters`
```
boards = []
winners = []
starters = []
for game in data:
# YOUR CODE HERE
```
### Q3
Make a dataframe out of the list `boards` and call it `df`. Make a series out of the list `winners`. Make a series out of the list `starters`.
Make a new column of `df` called "Winner" and set it equal to the pandas Series of the winners.
Make a new column of `df` called "Starter" and set it equal to the pandas Series of the starters.
```
# YOUR CODE HERE
```
## 3.2 Inferential Analysis
We're going to use Bayes Rule or Bayesian Inference to make a probability of winning based on positions of the board. The formula is:
$ P(A|B) = \frac{P(B|A) * P(A)}{P(B)} = \frac{P(A \cap B)}{P(B)}$
Where $\cap$ is the intersection of $A$ and $B$.
The example we will use is the following: _what is the probability of 'O' being the winner, given that they've played the center piece._
$B$ = 'O' played the center piece
$A$ = 'O' won the game
So what is probability? We will define it in terms of frequencies. So if we are for instance asking what is the probability of player 'O' being in the center piece, it would be defined as:
$ P(B) = \frac{|O_c|} {|O_c| + |X_c| + |empty|}$
Where the pipes, `| |`, or cardinality represent the count of the indicated observation or set. In this case $O_c$ (O being in the center) and $X_c$ (X being in the center).
```
Oc_Xc_empty = df[5].value_counts().sum()
Oc_Xc_empty
# example of assessing the probability of B, O playing the center piece
player = 'O'
Oc = (df[5] == player).value_counts()
Oc_Xc_empty = df[5].value_counts().sum()
Oc/Oc_Xc_empty
# we can also clean this up and replace the denominator with the whole
# observation space (which is just the total number of games, df.shape[0]).
# example of assesing probabiliy of A
(df['Winner'] == 'O').value_counts()/df.shape[0]
```
The $P(B|A) * P(A)$ is the intersection of $B$ and $A$. The intersection is defined as the two events occuring together.
Continuing with the example, the probablity of 'O' playing the center piece AND 'O' being the winner is _the number of times these observations occured together divided by the whole observation space_:
```
# in this view, the total times A and B occured together is 247
player = 'O'
df.loc[(df['Winner'] == player) & (df[5] == player)].shape[0]
# the total observation space is 1000 (1000 games)
df.shape[0]
```
And so we get:
$P(B|A) * P(A) = \frac{247} {1000} = 0.247 $
In code:
```
df.loc[(df['Winner'] == player) & (df[5] == player)].shape[0]/df.shape[0]
```
### 3.2.1 Behavioral Analysis of the Winner
#### Q4
define the 3 different board piece types and label them `middle`, `side`, and `corner`. Middle should be an int and the other two should be lists.
```
# define the 3 different board piece types
# middle =
# side =
# corner =
```
#### 3.2.1.1 What is the probability of winning after playing the middle piece?
#### Q5
```
# A intersect B: X played middle and X won / tot games
# B: X played middle / tot games
player = 'X'
# define the intersection of A AND B, A_B
# A_B =
# define prob B
# B =
# return A_B over B (The prob B given A)
A_B / B
```
#### Q6
```
# A intersect B: X played middle and X won / tot games
# B: X played middle / tot games
player = 'O'
# define the intersection of A AND B, A_B
# A_B =
# define prob B
# B =
# return A_B over B (The prob B given A)
A_B / B
```
#### 3.2.1.2 What is the probability of winning after playing a side piece?
#### Q7
```
# A intersect B: O played side and O won / tot games
# B: O played side / tot games
player = 'O'
A_B = df.loc[(df[side].T.apply(lambda x: player in x.values)) &
(df['Winner'] == player)].shape[0] / df.shape[0]
B = df.loc[(df[side].T.apply(lambda x: player in x.values))].shape[0] /\
df.shape[0]
A_B / B
# A intersect B: X played side and X won / tot games
# B: X played side / tot games
# player = # SET PLAYER
# A_B = df.loc[(df[<SET PIECE>].T.apply(lambda x: player in x.values)) &
# (df['Winner'] == player)].shape[0] / df.shape[0]
# B = df.loc[(df[<SET PIECE>].T.apply(lambda x: player in x.values))].shape[0] /\
# df.shape[0]
A_B / B
```
#### 3.2.1.3 What is the probability of winning after playing a corner piece?
### Q8
```
# A intersect B: O played corner and O won / tot games
# B: O played corner / tot games
# player = # SET PLAYER
# A_B = df.loc[(df[<SET PIECE>].T.apply(lambda x: player in x.values)) &
# (df['Winner'] == player)].shape[0] / df.shape[0]
# B = df.loc[(df[<SET PIECE>].T.apply(lambda x: player in x.values))].shape[0] /\
# df.shape[0]
A_B / B
```
### Q9
```
# A intersect B: X played corner and X won / tot games
# B: X played corner / tot games
# player = # SET PLAYER
# A_B = df.loc[(df[<SET PIECE>].T.apply(lambda x: player in x.values)) &
# (df['Winner'] == player)].shape[0] / df.shape[0]
# B = df.loc[(df[<SET PIECE>].T.apply(lambda x: player in x.values))].shape[0] /\
# df.shape[0]
A_B / B
```
Are these results surprising to you? Why? This [resource](https://www.cs.jhu.edu/~jorgev/cs106/ttt.pdf) may be illustrative.
## 3.3 Improving the Analysis
In this analysis, we only tracked what moves were made, not the order they were made in. It really limited our assessment! How might we change our recording of the games to track order of moves as well? Do we need to track all the moves or just the first and the winner?
| github_jupyter |
```
import os
os.environ['TRKXINPUTDIR']="/global/cfs/cdirs/m3443/data/trackml-kaggle/train_all"
os.environ['TRKXOUTPUTDIR']= "/global/cfs/projectdirs/m3443/usr/caditi97/iml2020/outtest"
import sys
import pkg_resources
import yaml
import pprint
import random
random.seed(1234)
import numpy as np
import pandas as pd
import itertools
import matplotlib.pyplot as plt
import tqdm
from os import listdir
from os.path import isfile, join
import matplotlib.cm as cm
# %matplotlib widget
# 3rd party
import torch
import torch.nn.functional as F
from torch_geometric.data import Data
from trackml.dataset import load_event
from pytorch_lightning import Trainer
from pytorch_lightning.callbacks import ModelCheckpoint
import seaborn as sns
from sklearn.manifold import TSNE
sys.path.append('/global/homes/c/caditi97/exatrkx-iml2020/exatrkx/src/')
# local import
from exatrkx import config_dict # for accessing predefined configuration files
from exatrkx import outdir_dict # for accessing predefined output directories
from exatrkx.src import utils_dir
from exatrkx.src import utils_robust
from utils_robust import *
# for preprocessing
from exatrkx import FeatureStore
from exatrkx.src import utils_torch
# for embedding
from exatrkx import LayerlessEmbedding
from exatrkx.src import utils_torch
from torch_cluster import radius_graph
from utils_torch import build_edges
from embedding.embedding_base import *
from processing.cell_direction_utils.utils import load_detector
from processing.utils import *
# for filtering
from exatrkx import VanillaFilter
# for GNN
import tensorflow as tf
from graph_nets import utils_tf
from exatrkx import SegmentClassifier
import sonnet as snt
# for labeling
from exatrkx.scripts.tracks_from_gnn import prepare as prepare_labeling
from exatrkx.scripts.tracks_from_gnn import clustering as dbscan_clustering
# track efficiency
from trackml.score import _analyze_tracks
from exatrkx.scripts.eval_reco_trkx import make_cmp_plot, pt_configs, eta_configs
from functools import partial
embed_ckpt_dir = '/global/cfs/cdirs/m3443/data/lightning_models/embedding/checkpoints/epoch=10.ckpt'
filter_ckpt_dir = '/global/cfs/cdirs/m3443/data/lightning_models/filtering/checkpoints/epoch=92.ckpt'
gnn_ckpt_dir = '/global/cfs/cdirs/m3443/data/lightning_models/gnn'
plots_dir = '/global/homes/c/caditi97/exatrkx-iml2020/exatrkx/src/plots/run1000' # needs to change...
ckpt_idx = -1 # which GNN checkpoint to load
dbscan_epsilon, dbscan_minsamples = 0.25, 2 # hyperparameters for DBScan
min_hits = 5 # minimum number of hits associated with a particle to define "reconstructable particles"
frac_reco_matched, frac_truth_matched = 0.5, 0.5 # parameters for track matching
device = 'cuda' if torch.cuda.is_available() else 'cpu'
e_ckpt = torch.load(embed_ckpt_dir, map_location=device)
e_config = e_ckpt['hyper_parameters']
e_config['clustering'] = 'build_edges'
e_config['knn_val'] = 500
e_config['r_val'] = 1.7
e_model = LayerlessEmbedding(e_config).to(device)
e_model.load_state_dict(e_ckpt["state_dict"])
e_model.eval()
import pickle
def load_event(event_path):
with open(event_path, 'rb') as f:
hits, truth = pickle.load(f)
return hits, truth
def get_pid(num, data):
total_un = data['pid'].unique()
total_unq = []
for u in total_un:
if (u!=0):
total_unq.append(u)
unq_pid = np.random.permutation(total_unq)[:num]
#unq_pid = total_unq[num]
return unq_pid
def select_tracks(unq_pid, data):
trk_hits = np.where(np.isin(data['pid'].detach().numpy(),unq_pid))
trk_idx = trk_hits[0]
trk_idx = trk_idx.flatten()
hid = data.hid[trk_idx]
pid = data.pid[trk_idx]
X = data.x[trk_idx]
cell_data = data.cell_data[trk_idx]
a = data.layerless_true_edges[0][trk_idx].numpy()
b = data.layerless_true_edges[1][trk_idx].numpy()
layerless = [a,b]
layerless = torch.from_numpy(np.asarray(layerless))
layers = data.layers[trk_idx]
new_data = Data(x=X, pid=pid, hid=hid, cell_data=cell_data, layers=layers, layerless_true_edges=layerless)
return new_data
# embedding
def embedding_hits(e_model,data):
with torch.no_grad():
# had to move everything to device
spatial = e_model(torch.cat([data.cell_data.to(device), data.x.to(device)], axis=-1))
spatial_np = spatial.detach().cpu().numpy()
e_spatial = utils_torch.build_edges(spatial, e_model.hparams['r_val'], e_model.hparams['knn_val'])
e_spatial_np = e_spatial.detach().cpu().numpy()
return e_spatial_np
misalignments = [0.001,0.0025,0.005,0.0075,0.01,0.012,0.015,0.017,0.02,0.1,0.4,0.6,0.8,1]
cell_features = ['cell_count', 'cell_val', 'leta', 'lphi', 'lx', 'ly', 'lz', 'geta', 'gphi']
feature_scale = [1000, np.pi, 1000]
detector_orig, detector_proc = load_detector('/global/cfs/projectdirs/m3443/usr/caditi97/iml2020/misaligned/volumes_shifted/detectors.csv')
evtid = 1001
num =1
og_event_file = f'/global/cfs/cdirs/m3443/data/trackml-kaggle/train_10evts/event00000{evtid}'
d = []
hits, particles, truth = trackml.dataset.load_event(og_event_file, parts=['hits', 'particles', 'truth'])
hits = hits.merge(truth, on='hit_id', how='left')
hits = hits.merge(particles, on='particle_id', how='left')
n_pids = 1
pids = particles[(particles.nhits) > 5]
np.random.seed(456)
rnd = np.random.randint(0, pids.shape[0], n_pids)
print("random idx: ", rnd)
sel_pids = pids.particle_id.values[rnd]
data = prepare_event(og_event_file, detector_orig=detector_orig, detector_proc=detector_proc,
cell_features=cell_features, output_dir=None,
pt_min=0, adjacent=True, endcaps=True, layerless=True, layerwise=False,
noise=0, cell_information=True, vol=None, savefile=False, pids=sel_pids)
e_spatial_np = embedding_hits(e_model,data)
fig = plt.figure(figsize=(8,8))
ax = fig.add_subplot(111)
# for pid in sel_pids:
# ax.scatter(hits[hits.particle_id == pid].x.values, hits[hits.particle_id == pid].y.values)
# add edges
e_spatial_np_t = e_spatial_np.T
for iedge in range(e_spatial_np.shape[1]):
ax.plot(hits.iloc[e_spatial_np_t[iedge]].x.values, hits.iloc[e_spatial_np_t[iedge]].y.values, color='k', alpha=0.3, lw=2.)
ax.set_xlabel('X')
ax.set_ylabel('Y')
```
| github_jupyter |
# The Mesh
The finite element mesh is a fundamental construct for Underworld modelling. It will generally determine your domain geometry, and the resolution of the finite element system. For parallel simulations, the mesh topology will also determine the domain decomposition for the problem. Currently `underworld` only provides curvilinear mesh capabilities.
#### Overview:
1. Creating mesh objects.
1. Element types.
1. Deforming the mesh.
1. Loading and saving the mesh.
1. Special sets.
1. Mesh variables
1. Setting values on a mesh variables.
1. Gradients of mesh variable fields.
1. Loading and saving mesh variable data.
**Keywords:** mesh variables, finite elements, load, save, initial conditions
## Creating the mesh
First create an 2x2 element mesh. By default the mesh will be of rectangular geometry, with domain extents specified via the `minCoord` and `maxCoord` constructor parameters.
```
# This example creates a mesh, and visualises it.
import underworld as uw
import underworld.visualisation as vis
mesh = uw.mesh.FeMesh_Cartesian( elementRes = (2, 2),
minCoord = (0.0, 0.0),
maxCoord = (2.0, 1.0) )
# visualising the result
figMesh = vis.Figure(figsize=(800,400))
figMesh.append( vis.objects.Mesh(mesh, nodeNumbers=True, pointsize=10) )
figMesh.show()
```
As is usually the pattern for data carrying objects in Underworld, underlying data is available as `numpy` arrays via the `data` mesh instances attribute. In the case of mesh objects, this data corresponds to mesh node coordinate information.
```
# This example simply prints mesh node information.
import underworld as uw
mesh = uw.mesh.FeMesh_Cartesian(elementRes=(2, 2))
for item in mesh.data:
print(item)
```
Note that data in `underworld` is always published as arrays of shape `(num_items, count)`, where `num_items` is the number of items in the dataset (in this case, 3x3 nodes, so `9`), and `count` is the size of each data item (so `1` for scalar data, `2` or `3` for vectors in 2- and 3-dimensions respectively, etc).
In parallel, each process will be assigned responsibility for a subsection of the domain, and as such the length (`num_items`) of the arrays will generally be different across processes. As such, while it might be tempting to use `numpy.reshape` to reshape your arrays to reflect the cartesian mesh topology, this approach will **not** work in parallel and is therefore not usually recommended. You should instead traverse data arrays as 1-d lists (each item of size `count`). Following this pattern will usually result in models which can safely be run in serial or parallel.
## Element Types
Underworld supports two primary element types: a linear element type (`Q1`) and a quadratic element type (`Q2`). Interpolation using these element types will provide continuous results across element boundaries, though note that interpolant derivatives will be discontinous across boundaries. Underworld also supports *mixed* element types, where a secondary element type may also be specified. This formulation is critical for stable solutions to Stokes type problems, with velocity fields being represented by the primary element type, and pressure fields by the secondary type. Secondary element types are not continuous across element boundaries. Supported secondary types are constant (`dQ0`), linear (`dPc1`) and bilinear (`dQ1`). Refer to http://femtable.org for further details on element types.
Mesh element types are specified as a textual argument to the `elementType` constructor parameter. A primary type must always be provided (`Q1` or `Q2`), and where a secondary type is required, the primary/secondary types are provided as a pair (`Q1/dQ0`, `Q2/dPc1` or `Q2/dQ1`). For mixed type elements, two mesh objects are generated by the constructor, one for the primary and one for the secondary elements. The primary mesh will be the object returned directly by the constructor, with the secondary mesh available via the primary mesh's `subMesh` attribute.
Note that where constructor parameters are omitted, default values are used. Defaults values are baked into the `underworld` API and may be determined in Jupyter by pressing `shift`-`tab` with the cursor inside the constructor parenthesis, or by using the Python `help()` function (with the class/function of interest as the argument). Default values are useful for quickly mocking up models, but may change without notice so should not be relied upon.
Element boundaries (and therefore domain geometry and resolution) is entirely determined by the primary mesh, with the location of the secondary mesh nodes slaved to the primary mesh nodes. Secondary mesh nodes are visualised below.
```
# Create a mesh with submesh
import underworld as uw
import underworld.visualisation as vis
mesh = uw.mesh.FeMesh_Cartesian( elementType="Q2/dpc1", maxCoord=(2.0, 1.0) )
submesh = mesh.subMesh
figMesh = vis.Figure(figsize=(800,400))
figMesh.append( vis.objects.Mesh(mesh) )
figMesh.append( vis.objects.Mesh(submesh, nodeNumbers=True, pointsize=10) )
figMesh.show()
```
## Deforming the mesh
It is often desirable to deform the mesh to either conform the domain to some geometry, or to implement mesh refinement by bunching nodes in certain domain regions. The user is free to modify the mesh as desired, though care must be taken to ensure elements are not improperly transformed (Jacobians should not become singular) and to avoid tangling (face normals should be consistent).
To deform the mesh, the user will directly modify the node coordinates via the usual `data` array. Note however that modifications are only possible from within the `deform_mesh()` context manager, as otherwise the array is read-only.
```
# A trivial deformation
import underworld as uw
import underworld.visualisation as vis
mesh = uw.mesh.FeMesh_Cartesian(elementRes=(2, 2), maxCoord=(2.0, 1.0) )
with mesh.deform_mesh():
mesh.data[4][0] += 0.05
figMesh = vis.Figure(figsize=(800,400))
figMesh.append( vis.objects.Mesh(mesh, nodeNumbers=True, pointsize=10) )
figMesh.show()
```
Note that in the above example, for simplicity, we've simply moved an arbitrary mesh node. Usually you will not want to construct your logic in ways which rely on the node numbers themselves, as this will usually not be consistent across parallel simulations, and/or as you vary model resolution.
We now look at a more realistic example where we deform the mesh to increase the resolution at the top of the domain. There are countless ways this may be achieved, but here we use the simple transformation $z \rightarrow \sqrt{z}$:
```
# A more useful deformation
import underworld as uw
import underworld.visualisation as vis
mesh = uw.mesh.FeMesh_Cartesian(elementRes=(32,32), maxCoord=(2.0, 1.0))
with mesh.deform_mesh():
for index, coord in enumerate(mesh.data):
mesh.data[index][1] = mesh.data[index][1]**0.5
figMesh = vis.Figure(figsize=(800,400))
figMesh.append(vis.objects.Mesh(mesh))
figMesh.show()
```
## Saving and loading the mesh
Your mesh data may be loaded or saved using the mesh instance `load()` and `save()` method respectively. All data in Underworld is written as HDF5 format binary files. There are numerous tools available for interrogating HDF5 files, but in Underworld we primarily leverage the `h5py` Python package. We recommend that users also familiarise themselves with this package to allow querying of Underworld generated data, and also as an excellent option for directly writing their own simulation data.
You will wish to save data for analysis purposes, but also for simulation restart. Note that Underworld does not provide any explicit *restart* functionality, and you will instead need to use object `load()` methods to return each object to their previous states. In Underworld, the pattern is always to first *recreate* vanilla equivalent objects, and *then* reload the data. So in the case of the mesh above, you must first recreate a standard mesh object, and it must be of identical resolution and element type. Finally you will reload the data.
```
# In this example we'll walk through some of the mesh loading mechanisms
import underworld as uw
mesh = uw.mesh.FeMesh_Cartesian()
# trivially deform
with mesh.deform_mesh():
mesh.data[3]+=0.1
# save
mesh.save('deformedMesh.h5')
# create second mesh
mesh2 = uw.mesh.FeMesh_Cartesian()
# confirm that it is different to first
import numpy as np
result = np.allclose(mesh.data,mesh2.data)
if result==True: # raise error if result not as expected
raise RuntimeError("Mesh objects should not match. Something wrong has happenend")
# now load data onto second mesh, and re-test
mesh2.load('deformedMesh.h5')
result = np.allclose(mesh.data,mesh2.data)
if result==False: # raise error if result not as expected
raise RuntimeError("Mesh objects should match. Something wrong has happenend")
# let's remove the saved file as it is no longer required.
if uw.mpi.rank==0:
import os;
os.remove('deformedMesh.h5')
```
<br>
## Special sets
Special sets are sets of nodes which are in some way special to a given mesh object. For instance, for the cartesian mesh you will often wish to tie special values to the walls to form your problem boundary conditions. The list of special sets is provided via the `specialSets` list object. The sets are named with respect to the `(I,J,K)` cartesian indexing of the mesh, with the `I` index usually used for the horizontal (left/right), `J` for the vertical, and `K` for horizontal (front/back). Note that this is just a matter of convention, and you are free to chose how the cartesian coordinates map within your model. So, for example, `MinI_VertexSet` specifies the set of nodes belonings to what we would normally consider to be the left wall, and similarly `MaxI_VertexSet` the right wall. We also provide the alias (`Left_VertexSet`, `Right_VertexSet`, etc) for the most common use case.
```
import underworld as uw
mesh = uw.mesh.FeMesh_Cartesian(elementRes=(2,2))
print("Available special sets for this mesh are:")
print(mesh.specialSets.keys())
leftset = mesh.specialSets['MinI_VertexSet']
print("\nIndices in the left set wall set:")
print(leftset)
```
You can confirm that these are indeed the left nodes with reference to the mesh visualisation at the top of the page. Note that the special set indices always specify parallel **local** indices. In almost all instances, you will interact with only local data and identifiers. This pattern is critical to successful parallel operation, and is usually a natural way of constructing your models. You should also note that the above visualisation shows the global node numbering, though for these serial demonstrations the values are identical. Your checkpointed mesh (and mesh variable) data will however be ordered according to its global numbering. If for some reason you need to determine the global identifiers, they are stored in the `data_elgId` array.
## Mesh Variables
Mesh variables are used to encode spatial data across the mesh. Underworld mesh variables assign data for each node of the mesh, so for example you might assign a temperature value at each mesh node. Mesh variables also leverage the mesh element shape functions to form interpolations within elements, allowing our discrete datasets to have continuous analogues formed.
Mesh variables may be used for any number of purposes when constructing models, but most importantly they form the unknowns for the finite element numerical systems you will be solving.
We create a mesh variable by using the mesh's `add_variable()` method, and specifying the `nodeDofCount` (node degree of freedom count). So for example, setting `nodeDofCount=1` yields a scalar mesh variable, having a single data value recorded at each node of the mesh, as you may require for perhaps a temperature field.
## Setting values on the MeshVariable
As you will find with most Underworld data carrying objects, you will access the data directly via the `data` attribute. Our data is stored there as a numpy array, and you may therefore accessed or modified it using standard Numpy operations.
Let's initialise a variable with a function based on its spatial coordinates
$$
T = 100\exp(1-z)
$$
This variable may be used to represent a temperature field. We will walk over the mesh vertex data to access coordinate information. As the temperature field array is (by design) a 1-1 map to the vertex array, we know the index of the mesh vertex will be the index of the associated temperature datum. Note that the Python *built-in* `enumerate()` acts to return **both** the index of a piece of data in a list (or array), and the data itself.
```
# create a mesh & meshvariable, and then initialise with data.
import underworld as uw
import underworld.visualisation as vis
import math
mesh = uw.mesh.FeMesh_Cartesian( maxCoord=(2., 1.) )
# now create our mesh variable
temperatureField = mesh.add_variable( nodeDofCount=1 )
pi = math.pi
for index, coord in enumerate(mesh.data):
temperatureField.data[index] = 100.*math.exp(1.-coord[1])
# vis results
fig = vis.Figure(figsize=(800,400))
fig.append( vis.objects.Surface(mesh, temperatureField) )
fig.append( vis.objects.Mesh(mesh) )
fig.show()
```
To describe for example a velocity, a vector mesh variable is required. We set `nodeDofCount=2` accordingly, and initialise to:
$$
\mathbf{v} = \left( z_0 - z, x - x_0 \right)
$$
```
# vector fields
import underworld as uw
import underworld.visualisation as vis
import math
mesh = uw.mesh.FeMesh_Cartesian( maxCoord=(2., 1.) )
velocityField = mesh.add_variable( nodeDofCount=2 )
coordmid = (1., 0.5)
for index, coord in enumerate(mesh.data):
vx = coordmid[1] - coord[1]
vz = coord[0] - coordmid[0]
velocityField.data[index] = (vx, vz)
# visualise both the velocity and temperature
fig = vis.Figure(figsize=(800,400))
fig.append( vis.objects.VectorArrows(mesh, velocityField, scaling=0.2, arrowHead=0.2) )
fig.show()
```
## Mesh variables gradients
Mesh variable gradients are calculated according to:
$$
\nabla A(\mathbf{r}) = A_i\nabla\Phi_i(\mathbf{r})
$$
Note that gradients derived in this way are not generally continuous across element boundaries. Note that we access the different gradient components via the square bracket operator, with ordering:
$$
[ \frac{\partial T}{\partial x}, \frac{\partial T}{\partial y}, \frac{\partial T}{\partial z} ]
$$
or for a vector field:
$$
[ \frac{\partial v_x}{\partial x}, \frac{\partial v_x}{\partial y}, \frac{\partial v_x}{\partial z},
\frac{\partial v_y}{\partial x}, \frac{\partial v_y}{\partial y}, \frac{\partial v_y}{\partial z},
\frac{\partial v_z}{\partial x}, \frac{\partial v_z}{\partial y}, \frac{\partial v_z}{\partial z} ]
$$
The gradient of the field is accessible via the ``fn_gradient`` attribute on the mesh variable.
## Loading and saving variables
The procedure for loading and saving variables is very similar to that used for the mesh, and uses the usual `save()` and `load()` methods. For model restarts, the normal pattern applies, with the user required to generate unmodified objects first, and then reloading the required data.
```
# mesh variable loading & saving
import underworld as uw
import numpy as np
mesh = uw.mesh.FeMesh_Cartesian()
meshvariable = mesh.add_variable(1)
# this might be a pattern you use in your models.
# set `restart` to required value, but note that
# you *must* first run with `True` to ensure that
# a file has been created!
restart=False
restartFile = "meshvariable.h5"
if not restart:
# not restarting, so init as required.
meshvariable.data[:] = 1.234
else:
meshvariable.load(restartFile)
# confirm that data is correct at this point!
result = np.allclose(meshvariable.data,1.234)
if result==False: # raise error if result not as expected
raise RuntimeError("Mesh variable does not contain expected values.")
# now save. generally this will occur after
# you've done something interesting, but we
# don't have time for that in this example.
ignore = meshvariable.save(restartFile)
# let's remove the saved file as it is no longer required.
if uw.mpi.rank==0:
import os;
os.remove(restartFile)
```
| github_jupyter |
# Grid-search results for GPs with RNNs
```
import pickle
import glob
import os
import pandas
import numpy as np
import json
with open('./datasets.json', 'r') as f:
datasets = json.load(f).keys()
models = ['GRU', 'LSTM', 'SigGRU', 'SigLSTM']
column_order = ['H8_D1', 'H8_D0', 'H32_D1', 'H32_D0', 'H128_D1', 'H128_D0']
results_folder_template = './gridsearch/GP{}'
results_dict = {m : {'acc' : {}, 'nlpp' : {}} for m in models}
```
### Parse results
```
for j, model in enumerate(models):
results_folder = results_folder_template.format(model)
for i, dataset in enumerate(datasets):
print('Parsing results for model: {} ({}/{}), dataset: {} ({:2d}/{})...'.format(model, j+1, len(models), dataset, i+1, len(datasets)))
# rint('Parsing results for model: ({}/{}) dataset: ({:2d}/{})...'.format(j+1, len(models), i+1, len(datasets)))
for num_hidden in [8, 32, 128]:
for use_dropout in [0, 1]:
experiment_name = '{}_H{}_D{}'.format(dataset, num_hidden, use_dropout)
file_path = os.path.join(results_folder, experiment_name + '.pkl')
if os.path.exists(file_path):
with open(file_path, 'rb') as f:
saved = pickle.load(f)
test_acc = saved['results']['test_acc']
test_nlpp = saved['results']['test_nlpp']
else:
print('Warning: result file missing {}'.format(file_path))
test_acc = float('nan')
test_nlpp = float('nan')
key = 'H{}_D{}'.format(num_hidden, use_dropout)
if key not in results_dict[model]['acc']:
results_dict[model]['acc'][key] = []
results_dict[model]['acc'][key].append(test_acc)
if key not in results_dict[model]['nlpp']:
results_dict[model]['nlpp'][key] = []
results_dict[model]['nlpp'][key].append(test_nlpp)
```
### Results of grid-search over 6 architectures
```
tol = 0.1
def highlight_best_nlpp(s):
min_s = s.min()
l = ['' for v in s]
for i in range(len(s)):
if np.abs(s[i] - min_s) / min_s <= tol:
l[i] = 'font-weight: bold'
break
return l
architectures = {model : {} for model in models}
from IPython.display import display
for model in models:
model_full = '# GP{} #'.format(model)
print()
print('#'*len(model_full))
print(model_full)
print('#'*len(model_full))
print()
print('Accuracy:'.format(model))
df = pandas.DataFrame.from_dict(results_dict[model]['acc']).rename(index={i : name for i, name in enumerate(datasets)})
df = df[column_order]
display(df)
print()
print('Negative log-predictive probability:'.format(model))
df = pandas.DataFrame.from_dict(results_dict[model]['nlpp']).rename(index={i : name for i, name in enumerate(datasets)})
df = df[column_order]
# get best architecture
for row in df.iterrows():
dataset = row[0]
results = row[1]
labels = results.index
nlpps = list(results)
min_nlpp = np.min(nlpps)
for i in range(len(nlpps)):
if np.abs(nlpps[i] - min_nlpp) / min_nlpp <= tol:
keys = labels[i].split('_')
architectures[model][dataset] = {x[0] : int(x[1:]) for x in keys}
break
df = df.style.apply(highlight_best_nlpp, axis=1)
display(df)
pandas.DataFrame.from_dict({m : ['H={}, D={}'.format(architectures[m][d]['H'], architectures[m][d]['D']) for d in datasets] for m in models}).rename(index={i : name for i, name in enumerate(datasets)})
with open('./architectures.json', 'w') as f:
json.dump(architectures, f)
architectures
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# TFRecord и tf.Example
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/tutorials/load_data/tfrecord"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />Смотрите на TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/ru/tutorials/load_data/tfrecord.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Запустите в Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/ru/tutorials/load_data/tfrecord.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />Изучайте код на GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/ru/tutorials/load_data/tfrecord.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Скачайте ноутбук</a>
</td>
</table>
Note: Вся информация в этом разделе переведена с помощью русскоговорящего Tensorflow сообщества на общественных началах. Поскольку этот перевод не является официальным, мы не гарантируем что он на 100% аккуратен и соответствует [официальной документации на английском языке](https://www.tensorflow.org/?hl=en). Если у вас есть предложение как исправить этот перевод, мы будем очень рады увидеть pull request в [tensorflow/docs](https://github.com/tensorflow/docs) репозиторий GitHub. Если вы хотите помочь сделать документацию по Tensorflow лучше (сделать сам перевод или проверить перевод подготовленный кем-то другим), напишите нам на [docs-ru@tensorflow.org list](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs-ru).
Чтобы эффективно читать данные будет полезно сериализовать ваши данные и держать их в наборе файлов (по 100-200MB каждый) каждый из которых может быть прочитан построчно. Это особенно верно если данные передаются по сети. Также это может быть полезно для кеширования и предобработки данных.
Формат TFRecord это простой формат для хранения последовательности двоичных записей.
[Protocol buffers](https://developers.google.com/protocol-buffers/) это кросс-платформенная, кросс-языковая библиотека для эффективной сериализации структурированных данных.
Сообщения протокола обычно определяются файлами `.proto`. Это часто простейший способ понять тип сообщения.
Сообщение `tf.Example` (или protobuf) гибкий тип сообщений, который преедставляет сопоставление `{"string": value}`. Он разработан для использования с TensorFlow и используется в высокоуровневых APIs таких как [TFX](https://www.tensorflow.org/tfx/).
Этот урок покажет как создавать, парсить и использовать сообщение `tf.Example`, а затем сериализовать читать и писать сообщения `tf.Example` в/из файлов `.tfrecord`.
Замечание: Хотя эти структуры полезны, они необязательны. Нет необходимости конвертировать существующий код для использования TFRecords если вы не используете [`tf.data`](https://www.tensorflow.org/guide/datasets) и чтение данных все еще узкое место обучения. См. [Производительность конвейера входных данных](https://www.tensorflow.org/guide/performance/datasets) для советов по производительности датасета.
## Setup
```
import tensorflow as tf
import numpy as np
import IPython.display as display
```
## `tf.Example`
### Типы данных для `tf.Example`
Фундаментально `tf.Example` это соответствие `{"string": tf.train.Feature}`.
Вид сообщений `tf.train.Feature` допускает один из следующих трех типов (См. [файл `.proto`](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/core/example/feature.proto) для справки). Большинство других общих типов может быть сведено к одному из этих трех:
1. `tf.train.BytesList` (можно привести следующие типы)
- `string`
- `byte`
1. `tf.train.FloatList` (можно привести следующие типы)
- `float` (`float32`)
- `double` (`float64`)
1. `tf.train.Int64List` (можно привести следующие типы)
- `bool`
- `enum`
- `int32`
- `uint32`
- `int64`
- `uint64`
Чтобы преобразовать стандартный тип TensorFlow в `tf.Example`-совместимый` tf.train.Feature`, вы можете использовать приведенные ниже функции. Обратите внимание, что каждая функция принимает на вход скалярное значение и возвращает `tf.train.Feature` содержащий один из трех вышеприведенных `list` типов:
```
# Следующая функция может быть использована чтобы преобразовать значение в тип совместимый с
# с tf.Example.
def _bytes_feature(value):
"""Преобразует string / byte в bytes_list."""
if isinstance(value, type(tf.constant(0))):
value = value.numpy() # BytesList не будет распаковывать строку из EagerTensor.
return tf.train.Feature(bytes_list=tf.train.BytesList(value=[value]))
def _float_feature(value):
"""Преобразует float / double в float_list."""
return tf.train.Feature(float_list=tf.train.FloatList(value=[value]))
def _int64_feature(value):
"""Преобразует bool / enum / int / uint в int64_list."""
return tf.train.Feature(int64_list=tf.train.Int64List(value=[value]))
```
Замечание: Для простоты этот пример использует только скалярные входные данные. Простейший способ обработки нескалярных признаков - использование `tf.serialize_tensor` для конвертации тензоров в двоичнеые строки. Стоки являются скалярами в тензорфлоу. Используйте `tf.parse_tensor` для обратной конвертации двоичных сток в тензор.
Ниже приведены несколько примеров того как работают эти функции. Обратите внимание на различные типы ввода и стандартизированные типы вывода. Если входной тип функции не совпадает с одним из приводимых типов указанных выше, функция вызовет исключение (например `_int64_feature(1.0)` выдаст ошибку поскольку `1.0` это значение с плавающей точкой и должно быть использовано с функцией `_float_feature`):
```
print(_bytes_feature(b'test_string'))
print(_bytes_feature(u'test_bytes'.encode('utf-8')))
print(_float_feature(np.exp(1)))
print(_int64_feature(True))
print(_int64_feature(1))
```
Все proto сообщения могут быть сериализованы в двоичную строку с использованием метода `.SerializeToString`:
```
feature = _float_feature(np.exp(1))
feature.SerializeToString()
```
### Создание сообщения `tf.Example`
Допустим вы хотите создать сообщение `tf.Example` из существующих данных. На практике данные могут прийти откуда угодно, но процедура создания сообщения `tf.Example` из одного наблюдения будет той же:
1. В рамках каждого наблюдения каждое значение должно быть преобразовано в `tf.train.Feature` содержащее одно из 3 совместимых типов, с использованием одной из вышеприведенных функций.
2. Вы создаете отображение (словарь) из строки названий признаков в закодированное значение признака выполненное на шаге #1.
3. Отображение (map) созданное на шаге 2 конвертируется в [`Features` message](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/core/example/feature.proto#L85).
В этом уроке вы создадите датасет с использованием NumPy.
У этого датасета будет 4 признака:
* булев признак, `False` или `True` с равной вероятностью
* целочисленный признак - равномерно случайно выбранный из `[0, 5]`
* строковый признак сгенерированный из табицы строк с использованием целочисленного признака в качестве индекса
* признак с плавающей точкой из стандартного нормального распределения
Рассмотрим выборку состающую из 10 000 независимых, одинаково распределенных наблюдений из каждого вышеприведенного распределения:
```
# Число наблюдений в датасете.
n_observations = int(1e4)
# Булев признак, принимающий значения False или True.
feature0 = np.random.choice([False, True], n_observations)
# Целочисленный признак, случайное число от 0 до 4.
feature1 = np.random.randint(0, 5, n_observations)
# Строковый признак
strings = np.array([b'cat', b'dog', b'chicken', b'horse', b'goat'])
feature2 = strings[feature1]
# Признак с плавающей точкой, из стандартного нормального распределения
feature3 = np.random.randn(n_observations)
```
Каждый из этих признаков может быть приведен к `tf.Example`-совместимому типу с использованием одного из `_bytes_feature`, `_float_feature`, `_int64_feature`. Вы можете затем создать `tf.Example`-сообщение из этих закодированных признаков:
```
def serialize_example(feature0, feature1, feature2, feature3):
"""
Создает tf.Example-сообщение готовое к записи в файл.
"""
# Создает словарь отображение имен признаков в tf.Example-совместимые
# типы данных.
feature = {
'feature0': _int64_feature(feature0),
'feature1': _int64_feature(feature1),
'feature2': _bytes_feature(feature2),
'feature3': _float_feature(feature3),
}
# Создает Features message с использованием tf.train.Example.
example_proto = tf.train.Example(features=tf.train.Features(feature=feature))
return example_proto.SerializeToString()
```
Возьмем, например, одно наблюдение из датасета, `[False, 4, bytes('goat'), 0.9876]`. Вы можете создать и распечатать `tf.Example`-сообщение для этого наблюдения с использованием `create_message()`. Каждое наблюдение может быть записано в виде `Features`-сообщения как указано выше. Note that the `tf.Example`-[сообщение](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/core/example/example.proto#L88) это всего лишь обертка вокруг `Features`-сообщения:
```
# Это пример наблюдения из набора данных.
example_observation = []
serialized_example = serialize_example(False, 4, b'goat', 0.9876)
serialized_example
```
Для декодирования сообщения используйте метод `tf.train.Example.FromString`.
```
example_proto = tf.train.Example.FromString(serialized_example)
example_proto
```
## Детали формата TFRecords
Файл TFRecord содержит последовательность записей. Файл может быть прочитан только линейно.
Каждая запись содержит строку байтов для данных плюс длину данных и CRC32C (32-bit CRC использующий полином Кастаньоли) хеши для проверки целостности.
Каждая запись хранится в следующих форматах:
uint64 length
uint32 masked_crc32_of_length
byte data[length]
uint32 masked_crc32_of_data
Записи сцеплены друг с другом и организуют файл.. CRCs
[описаны тут](https://en.wikipedia.org/wiki/Cyclic_redundancy_check), и
маска CRC выглядит так:
masked_crc = ((crc >> 15) | (crc << 17)) + 0xa282ead8ul
Замечание: Не обязательно использовать `tf.Example` в файлах TFRecord. `tf.Example` это всего лишь метод сериализации словарей в байтовые строки. Строки текста, закодированные данные изображений, или сериализованные тензоры (с использованием `tf.io.serialize_tensor`, и
`tf.io.parse_tensor` при загрузке). См. модуль `tf.io` для дополнительных возможностей.
## Файлы TFRecord с использованием `tf.data`
Модуль `tf.data` также предоставляет инструменты для чтения и записи данных в TensorFlow.
### Запись файла TFRecord
Простейший способ помещения данных в датасет это использование метода `from_tensor_slices`.
Примененный к массиву он возвращает датасет скаляров:
```
tf.data.Dataset.from_tensor_slices(feature1)
```
Примененный к кортежу массивов он возвращает датасет кортежей:
```
features_dataset = tf.data.Dataset.from_tensor_slices((feature0, feature1, feature2, feature3))
features_dataset
# Используйте `take(1)` чтобы взять только один пример из датасета.
for f0,f1,f2,f3 in features_dataset.take(1):
print(f0)
print(f1)
print(f2)
print(f3)
```
Используйте метод `tf.data.Dataset.map` чтобы применить функцию к каждому элементу `Dataset`.
«Функция отображения должна работать в графовом режиме TensorFlow - она должна принимать и возвращать` tf.Tensors`. Не тензорная функция, такая как `create_example`, может быть заключена в` tf.py_function`, для совместимости.
Использование `tf.py_function` требует указания размерности и информации о типе, которая в противном случае недоступна:
```
def tf_serialize_example(f0,f1,f2,f3):
tf_string = tf.py_function(
serialize_example,
(f0,f1,f2,f3), # передайте эти аргументы в верхнюю функцию.
tf.string) # возвращаемый тип `tf.string`.
return tf.reshape(tf_string, ()) # Результатом является скаляр
tf_serialize_example(f0,f1,f2,f3)
```
Примените эту функцию к каждому элементу датасета:
```
serialized_features_dataset = features_dataset.map(tf_serialize_example)
serialized_features_dataset
def generator():
for features in features_dataset:
yield serialize_example(*features)
serialized_features_dataset = tf.data.Dataset.from_generator(
generator, output_types=tf.string, output_shapes=())
serialized_features_dataset
```
И запишите их в файл TFRecord:
```
filename = 'test.tfrecord'
writer = tf.data.experimental.TFRecordWriter(filename)
writer.write(serialized_features_dataset)
```
### Чтение TFRecord файла
Вы можете также прочитать TFRecord файл используя класс `tf.data.TFRecordDataset`.
Больше информации об использовании TFRecord файлов с использованием `tf.data` может быть найдено [тут](https://www.tensorflow.org/guide/datasets#consuming_tfrecord_data)..
Использование `TFRecordDataset`-ов может быть полезно для стандартизации входных данных и оптимизации производительности.
```
filenames = [filename]
raw_dataset = tf.data.TFRecordDataset(filenames)
raw_dataset
```
На этом этапе датасет содержит сериализованные сообщения `tf.train.Example`. При их итерации возвращаются скалярные строки тензоров.
Используйте метод `.take` чтобы показать только первые 10 записей.
Замечание: итерация по `tf.data.Dataset` работает только при включенном eager execution.
```
for raw_record in raw_dataset.take(10):
print(repr(raw_record))
```
Эти тензоры может распарсить используя нижеприведенную функцию. Заметьте что `feature_description` обязателен тут поскольку датасеты используют графовое исполнение и нуждаются в этом описании для построения своей размерностной и типовой сигнатуры:
```
# Создайте описание этих признаков
feature_description = {
'feature0': tf.io.FixedLenFeature([], tf.int64, default_value=0),
'feature1': tf.io.FixedLenFeature([], tf.int64, default_value=0),
'feature2': tf.io.FixedLenFeature([], tf.string, default_value=''),
'feature3': tf.io.FixedLenFeature([], tf.float32, default_value=0.0),
}
def _parse_function(example_proto):
# Разберите `tf.Example` proto используя вышеприведенный словарь.
return tf.io.parse_single_example(example_proto, feature_description)
```
Альтернативно, используйте `tf.parse example` чтобы распарсить весь пакет за раз. Примените эту функцию к кажому элементу датасета используя метод `tf.data.Dataset.map`:
```
parsed_dataset = raw_dataset.map(_parse_function)
parsed_dataset
```
Используйте eager execution чтобы показывать наблюдения в датасете. В этом наборе данных 10,000 наблюдений, но вы выведете только первые 10. Данные показываются как словарь признаков. Каждое наблюдение это `tf.Tensor`, и элемент `numpy`этого тензора показывает значение признака:
```
for parsed_record in parsed_dataset.take(10):
print(repr(parsed_record))
```
Здесь функция `tf.parse_example` распаковывает поля `tf.Example` в стандартные тензоры.
## TFRecord файлы в Python
Модуль `tf.io` также содержит чисто Python функции для чтения и записи файлов TFRecord.
### Запись TFRecord файла
Далее запишем эти 10 000 наблюдений в файл `test.tfrecord`. Каждое наблюдения конвертируется в `tf.Example`-сообщение и затем пишется в файл. Вы можете после проверить, что файл `test.tfrecord` был создан:
```
# Запишем наблюдения `tf.Example` в файл.
with tf.io.TFRecordWriter(filename) as writer:
for i in range(n_observations):
example = serialize_example(feature0[i], feature1[i], feature2[i], feature3[i])
writer.write(example)
!du -sh {filename}
```
### Чтение TFRecord файла
Эти сериализованные тензоры могут быть легко распарсены с использование `tf.train.Example.ParseFromString`:
```
filenames = [filename]
raw_dataset = tf.data.TFRecordDataset(filenames)
raw_dataset
for raw_record in raw_dataset.take(1):
example = tf.train.Example()
example.ParseFromString(raw_record.numpy())
print(example)
```
## Упражнение: Чтение и запись данных изображений
Это пример того как читать и писать данные изображений используя TFRecords. Цель этого показать как, от начала до конца, ввести данные (в этом случае изображение) и записать данные в TFRecord файл, затем прочитать файл и показать изображение.
Это будет полезно если, например, вы хотите использовать несколько моделей на одних и тех же входных данных. Вместо хранения сырых данных изображений, они могут быть предобработанны в формат TFRecords, и затем могут быть использованы во всех дальнейших обработках и моделированиях.
Сперва давайте скачаем [это изображение](https://commons.wikimedia.org/wiki/File:Felis_catus-cat_on_snow.jpg) кота и покажем [это фото](https://upload.wikimedia.org/wikipedia/commons/f/fe/New_East_River_Bridge_from_Brooklyn_det.4a09796u.jpg) строительства моста Williamsburg, NYC.
### Получите изображения
```
cat_in_snow = tf.keras.utils.get_file('320px-Felis_catus-cat_on_snow.jpg', 'https://storage.googleapis.com/download.tensorflow.org/example_images/320px-Felis_catus-cat_on_snow.jpg')
williamsburg_bridge = tf.keras.utils.get_file('194px-New_East_River_Bridge_from_Brooklyn_det.4a09796u.jpg','https://storage.googleapis.com/download.tensorflow.org/example_images/194px-New_East_River_Bridge_from_Brooklyn_det.4a09796u.jpg')
display.display(display.Image(filename=cat_in_snow))
display.display(display.HTML('Image cc-by: <a "href=https://commons.wikimedia.org/wiki/File:Felis_catus-cat_on_snow.jpg">Von.grzanka</a>'))
display.display(display.Image(filename=williamsburg_bridge))
display.display(display.HTML('<a "href=https://commons.wikimedia.org/wiki/File:New_East_River_Bridge_from_Brooklyn_det.4a09796u.jpg">From Wikimedia</a>'))
```
### Write the TFRecord file
Как и ранее закодируйте признаки как типы совместимые с `tf.Example`. Здесь хранится необработанные данные изображения в формате string, так же как и высота, ширина, глубина и произвольный признак `label`. Последнее используется когда вы пишете файл чтобы различать изображение кота и моста. Используйте `0` изображения кота, и `1` для моста:
```
image_labels = {
cat_in_snow : 0,
williamsburg_bridge : 1,
}
# Это пример использования только изображения кота.
image_string = open(cat_in_snow, 'rb').read()
label = image_labels[cat_in_snow]
# Создайте библиотеку с признаками которые могут быть релевантны.
def image_example(image_string, label):
image_shape = tf.image.decode_jpeg(image_string).shape
feature = {
'height': _int64_feature(image_shape[0]),
'width': _int64_feature(image_shape[1]),
'depth': _int64_feature(image_shape[2]),
'label': _int64_feature(label),
'image_raw': _bytes_feature(image_string),
}
return tf.train.Example(features=tf.train.Features(feature=feature))
for line in str(image_example(image_string, label)).split('\n')[:15]:
print(line)
print('...')
```
Заметьте что все признаки сейчас содержатся в `tf.Example`-сообщении. Далее функционализируйте вышеприведенный код и запишите пример сообщений в файл с именем `images.tfrecords`:
```
# Запишем файлы изображений в `images.tfrecords`.
# Сперва, преобразуем два изображения в `tf.Example`-сообщения.
# Затем запишем их в `.tfrecords` файл.
record_file = 'images.tfrecords'
with tf.io.TFRecordWriter(record_file) as writer:
for filename, label in image_labels.items():
image_string = open(filename, 'rb').read()
tf_example = image_example(image_string, label)
writer.write(tf_example.SerializeToString())
!du -sh {record_file}
```
### Чтение TFRecord файла
У вас сейчас есть файл `images.tfrecords` и вы можете проитерировать записи в нем чтобы прочитать то что вы в него записали. Поскольку этот пример содержит только изображение единственное свойство которое вам нужно это необработанная строка изображения. Извлеките ее используя геттеры описанные выше, а именно `example.features.feature['image_raw'].bytes_list.value[0]`. Вы можете также использовать метки чтобы определить, которая запись является котом, и которая мостом:
```
raw_image_dataset = tf.data.TFRecordDataset('images.tfrecords')
# Создадим словарь описывающий свойства.
image_feature_description = {
'height': tf.io.FixedLenFeature([], tf.int64),
'width': tf.io.FixedLenFeature([], tf.int64),
'depth': tf.io.FixedLenFeature([], tf.int64),
'label': tf.io.FixedLenFeature([], tf.int64),
'image_raw': tf.io.FixedLenFeature([], tf.string),
}
def _parse_image_function(example_proto):
# Распарсим входной tf.Example proto используя вышесозданный словарь.
return tf.io.parse_single_example(example_proto, image_feature_description)
parsed_image_dataset = raw_image_dataset.map(_parse_image_function)
parsed_image_dataset
```
Восстановим изображение из TFRecord файла:
```
for image_features in parsed_image_dataset:
image_raw = image_features['image_raw'].numpy()
display.display(display.Image(data=image_raw))
```
| github_jupyter |
This example illustrates the prior and posterior of a GPR with different kernels. Mean, standard deviation, and 10 samples are shown for both prior and posterior.
#### New to Plotly?
Plotly's Python library is free and open source! [Get started](https://plot.ly/python/getting-started/) by downloading the client and [reading the primer](https://plot.ly/python/getting-started/).
<br>You can set up Plotly to work in [online](https://plot.ly/python/getting-started/#initialization-for-online-plotting) or [offline](https://plot.ly/python/getting-started/#initialization-for-offline-plotting) mode, or in [jupyter notebooks](https://plot.ly/python/getting-started/#start-plotting-online).
<br>We also have a quick-reference [cheatsheet](https://images.plot.ly/plotly-documentation/images/python_cheat_sheet.pdf) (new!) to help you get started!
### Version
```
import sklearn
sklearn.__version__
```
### Imports
This tutorial imports [GaussianProcessRegressor](http://scikit-learn.org/stable/modules/generated/sklearn.gaussian_process.GaussianProcessRegressor.html#sklearn.gaussian_process.GaussianProcessRegressor), [RBF](http://scikit-learn.org/stable/modules/generated/sklearn.gaussian_process.kernels.RBF.html#sklearn.gaussian_process.kernels.RBF), [Matern](http://scikit-learn.org/stable/modules/generated/sklearn.gaussian_process.kernels.Matern.html#sklearn.gaussian_process.kernels.Matern), [RationalQuadratic](http://scikit-learn.org/stable/modules/generated/sklearn.gaussian_process.kernels.RationalQuadratic.html#sklearn.gaussian_process.kernels.RationalQuadratic), [ExpSineSquared](http://scikit-learn.org/stable/modules/generated/sklearn.gaussian_process.kernels.ExpSineSquared.html#sklearn.gaussian_process.kernels.ExpSineSquared), [DotProduct](http://scikit-learn.org/stable/modules/generated/sklearn.gaussian_process.kernels.DotProduct.html#sklearn.gaussian_process.kernels.DotProduct) and [ConstantKernel](http://scikit-learn.org/stable/modules/generated/sklearn.gaussian_process.kernels.ConstantKernel.html#sklearn.gaussian_process.kernels.ConstantKernel).
```
import plotly.plotly as py
import plotly.graph_objs as go
from plotly import tools
import numpy as np
from sklearn.gaussian_process import GaussianProcessRegressor
from sklearn.gaussian_process.kernels import (RBF, Matern, RationalQuadratic,
ExpSineSquared, DotProduct,
ConstantKernel)
```
### Calculations
```
kernels = [1.0 * RBF(length_scale=1.0, length_scale_bounds=(1e-1, 10.0)),
1.0 * RationalQuadratic(length_scale=1.0, alpha=0.1),
1.0 * ExpSineSquared(length_scale=1.0, periodicity=3.0,
length_scale_bounds=(0.1, 10.0),
periodicity_bounds=(1.0, 10.0)),
ConstantKernel(0.1, (0.01, 10.0))
* (DotProduct(sigma_0=1.0, sigma_0_bounds=(0.0, 10.0)) ** 2),
1.0 * Matern(length_scale=1.0, length_scale_bounds=(1e-1, 10.0),
nu=1.5)]
color = 2 * ['red', 'green', 'blue', 'cyan', 'magenta', 'orange']
```
### Plot Results
```
plots = []
titles = []
for fig_index, kernel in enumerate(kernels):
# Specify Gaussian Process
plots.append([[], []])
gp = GaussianProcessRegressor(kernel=kernel)
# Plot prior
X_ = np.linspace(0, 5, 100)
y_mean, y_std = gp.predict(X_[:, np.newaxis], return_std=True)
p1 = go.Scatter(x=X_, y=y_mean,
showlegend=False,
mode='lines',
line=dict(color='black')
)
p2 = go.Scatter(x=X_, y=y_mean + y_std,
mode='lines',
showlegend=False,
line=dict(color='black')
)
p3 = go.Scatter(x=X_, y=y_mean - y_std,
mode='lines',
showlegend=False,
line=dict(color='black'),
fill = 'tonexty'
)
plots[fig_index][0].append(p2)
plots[fig_index][0].append(p3)
plots[fig_index][0].append(p1)
y_samples = gp.sample_y(X_[:, np.newaxis], 10)
k = []
for col in range(0, len(y_samples[0])):
k.append([])
for row in range(0, len(y_samples)):
k[col].append(y_samples[row][col])
for l in range(0, 10):
p4 = go.Scatter(x=X_, y=k[l],
showlegend=False,
mode='lines',
line=dict(color=color[l], width=1),
)
plots[fig_index][0].append(p4)
titles.append("Prior <br>(kernel: %s)" % kernel)
# Generate data and fit GP
rng = np.random.RandomState(4)
X = rng.uniform(0, 5, 10)[:, np.newaxis]
y = np.sin((X[:, 0] - 2.5) ** 2)
gp.fit(X, y)
# Plot posterior
y_mean, y_std = gp.predict(X_[:, np.newaxis], return_std=True)
p1 = go.Scatter(x=X_, y=y_mean,
showlegend=False,
mode='lines',
line=dict(color='black')
)
p2 = go.Scatter(x=X_, y=y_mean + y_std,
showlegend=False,
mode='lines',
line=dict(color='black')
)
p3 = go.Scatter(x=X_, y=y_mean - y_std,
mode='lines',
showlegend=False,
line=dict(color='black'),
fill = 'tonexty'
)
plots[fig_index][1].append(p2)
plots[fig_index][1].append(p3)
plots[fig_index][1].append(p1)
y_samples = gp.sample_y(X_[:, np.newaxis], 10)
k = []
for col in range(0, len(y_samples[0])):
k.append([])
for row in range(0, len(y_samples)):
k[col].append(y_samples[row][col])
for l in range(0, 10):
p4 = go.Scatter(x=X_, y=k[l],
showlegend=False,
mode='lines',
line=dict(color=color[l], width=1),
)
plots[fig_index][1].append(p4)
p5 = go.Scatter(x=X[:, 0], y=y,
showlegend=False,
mode='markers',
line=dict(color='red'),
)
plots[fig_index][1].append(p5)
titles.append("Posterior <br>(kernel: %s)<br>Log-Likelihood: %.3f"
% (gp.kernel_, gp.log_marginal_likelihood(gp.kernel_.theta)))
```
Create Plotly subplots
```
def create_subplots(plots, titles):
fig = tools.make_subplots(rows=1, cols=2,
subplot_titles=tuple(titles),
print_grid=False)
for j in range(0, len(plots[0])):
fig.append_trace(plots[0][j], 1, 1)
for k in range(0, len(plots[1])):
fig.append_trace(plots[1][k], 1, 2)
for i in map(str, range(1, 3)):
y = 'yaxis' + i
x = 'xaxis' + i
fig['layout'][y].update(showticklabels=False, ticks='',
zeroline=False, showgrid=False)
fig['layout'][x].update(showticklabels=False, ticks='',
zeroline=False, showgrid=False)
fig['layout'].update(hovermode='closest',
margin=dict(l=0, b=10,
r=0, ))
return fig
```
### RBF Kernels
```
fig = create_subplots(plots[0], titles[0 : 2])
py.iplot(fig)
```
### RationalQuadratic Kernels
```
fig = create_subplots(plots[1], titles[2 : 4])
py.iplot(fig)
```
### ExpSineSquared Kernel
```
fig = create_subplots(plots[2], titles[4 : 6])
py.iplot(fig)
```
### DotProduct Kernel
```
fig = create_subplots(plots[3], titles[6 : 8])
py.iplot(fig)
```
### Matern
```
fig = create_subplots(plots[4], titles[8 : 10])
py.iplot(fig)
```
### License
Authors:
Jan Hendrik Metzen <jhm@informatik.uni-bremen.de>
License:
BSD 3 clause
```
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
! pip install git+https://github.com/plotly/publisher.git --upgrade
import publisher
publisher.publish(
'Illustration of Prior and Posterior Gaussian Process for Different Kernels.ipynb', 'scikit-learn/plot-gpr-prior-posterior/', 'Illustration of Prior and Posterior Gaussian Process for Different Kernels | plotly',
' ',
title = 'Illustration of Prior and Posterior Gaussian Process for Different Kernels | plotly',
name = 'Illustration of Prior and Posterior Gaussian Process for Different Kernels',
has_thumbnail='true', thumbnail='thumbnail/prior.jpg',
language='scikit-learn', page_type='example_index',
display_as='gaussian-process', order=5,
ipynb= '~Diksha_Gabha/3101')
```
| github_jupyter |
```
# Imports
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from IPython.display import display, Markdown, Latex
plt.rcParams['figure.figsize'] = (15.0, 8.0) # set default size of plots
plt.rcParams['figure.facecolor'] = 'white'
pd.set_option('display.max_rows', None)
matplotlib.rcParams.update({'font.size': 15})
def read_file(name, database_type):
filename = name + "-" + str(database_type) + ".csv"
return pd.read_csv(filename, names = ["time"])
def compute_mean(data):
return data['time'].median()
def compute_stddev(data):
return data['time'].std()
def as_kb_string(kb):
return f"{kb:,}"
def as_ms_string(value):
return f"{value:,}"
def to_kb_string(b):
return f"{round(b/1024, 0):,}".replace(".0", "")
def r2(integer):
return f"{round(integer, 3):,}"
compilation_native_dataset = read_file('compilation-native', 0)
compilation_sgx_dataset = read_file('compilation-sgx', 0)
compilation_wasm_dataset = read_file('compilation-wasm', 0)
compilation_wasm_bin_dataset = read_file('compilation-wasm-bin', 0)
compilation_wasm_sgx_bin_dataset = read_file('compilation-wasm-sgx-bin', 0)
launch_time_native_dataset = read_file('launch-time-native', 0)
launch_time_sgx_dataset = read_file('launch-time-sgx', 0)
launch_time_wasm_dataset = read_file('launch-time-wasm', 0)
launch_time_wasm_sgx_dataset = read_file('launch-time-wasm-sgx', 0)
optimization_sgx_lkl_disk_image_dataset = read_file('optimization-sgx-disk-image', 0)
optimization_wasm_aot_dataset = read_file('optimization-wasm-aot', 0)
memory_native_in_memory_dataset = read_file('resident-memory-native', 0)
memory_native_in_file_dataset = read_file('resident-memory-native', 1)
memory_sgx_in_memory_dataset = read_file('resident-memory-sgx', 0)
memory_sgx_in_file_dataset = read_file('resident-memory-sgx', 1)
memory_wasm_in_memory_dataset = read_file('resident-memory-wasm', 0)
memory_wasm_in_file_dataset = read_file('resident-memory-wasm', 1)
memory_wasm_sgx_in_memory_dataset = read_file('resident-memory-wasm-sgx', 0)
memory_wasm_sgx_in_file_dataset = read_file('resident-memory-wasm-sgx', 1)
size_native_dataset = read_file("size-native", 0)
size_sgx_lkl_bin_dataset = read_file("size-sgx-lkl-bin", 0)
size_sgx_lkl_enclave_dataset = read_file("size-sgx-lkl-enclave", 0)
size_sgx_lkl_disk_image_dataset = read_file("size-sgx-disk-image", 0)
size_wasm_dataset = read_file("size-wasm", 0)
size_wasm_aot_dataset = read_file("size-wasm-aot", 0)
size_wasm_bin_dataset = read_file("size-wasm-bin", 0)
size_wasm_sgx_bin_dataset = read_file("size-wasm-sgx-bin", 0)
size_wasm_sgx_enclave_dataset = read_file("size-wasm-sgx-enclave", 0)
display(Markdown("## Compilation and optimisation time [ms]"))
l = []
l.append("||Native|SGX-LKL|WAMR|TWINE|")
l.append("|:-|-|-|-|-|")
compilation_native_average = as_ms_string(int(compute_mean(compilation_native_dataset) * 1000))
compilation_native_stddev = str(int(compute_stddev(compilation_native_dataset) * 1000))
l.append("|**SQLite (.so)**|" + \
compilation_native_average + \
" (σ = " + compilation_native_stddev + ")|" + \
compilation_native_average + \
" (σ = " + compilation_native_stddev + ")||||")
optimization_sgx_lkl_disk_image_average = as_ms_string(int(compute_mean(optimization_sgx_lkl_disk_image_dataset) * 1000))
optimization_sgx_lkl_disk_image_stddev = str(int(compute_stddev(optimization_sgx_lkl_disk_image_dataset) * 1000))
l.append("|**Disk image**||" + \
optimization_sgx_lkl_disk_image_average + \
" (σ = " + optimization_sgx_lkl_disk_image_stddev + ")|||")
compilation_sgx_average = as_ms_string(int(compute_mean(compilation_sgx_dataset) * 1000))
compilation_sgx_stddev = str(int(compute_stddev(compilation_sgx_dataset) * 1000))
compilation_wasm_bin_average = as_ms_string(int(compute_mean(compilation_wasm_bin_dataset) * 1000))
compilation_wasm_bin_stddev = str(int(compute_stddev(compilation_wasm_bin_dataset) * 1000))
compilation_wasm_sgx_bin_average = as_ms_string(int(compute_mean(compilation_wasm_sgx_bin_dataset) * 1000))
compilation_wasm_sgx_bin_stddev = str(int(compute_stddev(compilation_wasm_sgx_bin_dataset) * 1000))
l.append("|**Runtime (.so)**||" +
compilation_sgx_average + \
" (σ = " + compilation_sgx_stddev + ")|" + \
compilation_wasm_bin_average + \
" (σ = " + compilation_wasm_bin_stddev + ")|" + \
compilation_wasm_sgx_bin_average + \
" (σ = " + compilation_wasm_sgx_bin_stddev + ")|")
compilation_wasm_average = as_ms_string(int(compute_mean(compilation_wasm_dataset) * 1000))
compilation_wasm_stddev = str(int(compute_stddev(compilation_wasm_dataset) * 1000))
l.append("|**SQLite Wasm**|||" + \
compilation_wasm_average + \
" (σ = " + compilation_wasm_stddev + ")|" + \
compilation_wasm_average + \
" (σ = " + compilation_wasm_stddev + ")|")
optimization_wasm_aot_average = as_ms_string(int(compute_mean(optimization_wasm_aot_dataset) * 1000))
optimization_wasm_aot_stddev = str(int(compute_stddev(optimization_wasm_aot_dataset) * 1000))
l.append("|**SQLite AoT**|||" + \
optimization_wasm_aot_average + \
" (σ = " + optimization_wasm_aot_stddev + ")|" + \
optimization_wasm_aot_average + \
" (σ = " + optimization_wasm_aot_stddev + ")|")
display(Markdown('\n'.join(l)))
display(Markdown("## Launch time [ms]"))
l = []
l.append("|Native|SGX|WAMR|TWINE|")
l.append("|-|-|-|-|")
launch_time_native_average = as_ms_string(int(compute_mean(launch_time_native_dataset)))
launch_time_native_stddev = str(int(compute_stddev(launch_time_native_dataset)))
launch_time_sgx_average = as_ms_string(int(compute_mean(launch_time_sgx_dataset)))
launch_time_sgx_stddev = str(int(compute_stddev(launch_time_sgx_dataset)))
launch_time_wasm_average = as_ms_string(int(compute_mean(launch_time_wasm_dataset)))
launch_time_wasm_stddev = str(int(compute_stddev(launch_time_wasm_dataset)))
launch_time_wasm_sgx_average = as_ms_string(int(compute_mean(launch_time_wasm_sgx_dataset)))
launch_time_wasm_sgx_stddev = str(int(compute_stddev(launch_time_wasm_sgx_dataset)))
l.append("|" + launch_time_native_average + \
" (σ = " + launch_time_native_stddev + ")" + \
"|" + launch_time_sgx_average + \
" (σ = " + launch_time_sgx_stddev + ")" + \
"|" + launch_time_wasm_average + \
" (σ = " + launch_time_wasm_stddev + ")" + \
"|" + launch_time_wasm_sgx_average + \
" (σ = " + launch_time_wasm_sgx_stddev + ")")
display(Markdown('\n'.join(l)))
display(Markdown("## Resident memory [kB]"))
# The size of the enclave are known at build time
# The size of SGX-LKL enclave is given in the file enclave_config.json
memory_sgx_enclave_size = as_kb_string(261120)
# The size of Twine enclave is given in the SGX XML configuration file
memory_wasm_sgx_enclave_size = as_kb_string(209920)
l = []
l.append("with 175k records in the database.")
l.append("")
l.append("||Native|SGX-LKL|WAMR|TWINE|")
l.append("|:-|-|-|-|-|")
memory_native_in_memory_average = as_kb_string(int(compute_mean(memory_native_in_memory_dataset)))
memory_native_in_memory_stddev = str(int(compute_stddev(memory_native_in_memory_dataset)))
memory_sgx_in_memory_average = as_kb_string(int(compute_mean(memory_sgx_in_memory_dataset)))
memory_sgx_in_memory_stddev = str(int(compute_stddev(memory_sgx_in_memory_dataset)))
memory_wasm_in_memory_average = as_kb_string(int(compute_mean(memory_wasm_in_memory_dataset)))
memory_wasm_in_memory_stddev = str(int(compute_stddev(memory_wasm_in_memory_dataset)))
memory_wasm_sgx_in_memory_average = as_kb_string(int(compute_mean(memory_wasm_sgx_in_memory_dataset)))
memory_wasm_sgx_in_memory_stddev = str(int(compute_stddev(memory_wasm_sgx_in_memory_dataset)))
l.append("|**in-memory (untrusted)**|" + \
memory_native_in_memory_average + \
" (σ = " + memory_native_in_memory_stddev + ")|" + \
memory_sgx_in_memory_average + \
" (σ = " + memory_sgx_in_memory_stddev + ")|" + \
memory_wasm_in_memory_average + \
" (σ = " + memory_wasm_in_memory_stddev + ")|" + \
memory_wasm_sgx_in_memory_average + \
" (σ = " + memory_wasm_sgx_in_memory_stddev + ")|")
l.append("|**in-memory (trusted)**|" + \
"|" + \
memory_sgx_enclave_size + " (fixed)|" + \
"|" + \
memory_wasm_sgx_enclave_size + " (fixed)|")
memory_native_in_file_average = as_kb_string(int(compute_mean(memory_native_in_file_dataset)))
memory_native_in_file_stddev = str(int(compute_stddev(memory_native_in_file_dataset)))
memory_sgx_in_file_average = as_kb_string(int(compute_mean(memory_sgx_in_file_dataset)))
memory_sgx_in_file_stddev = str(int(compute_stddev(memory_sgx_in_file_dataset)))
memory_wasm_in_file_average = as_kb_string(int(compute_mean(memory_wasm_in_file_dataset)))
memory_wasm_in_file_stddev = str(int(compute_stddev(memory_wasm_in_file_dataset)))
memory_wasm_sgx_in_file_average = as_kb_string(int(compute_mean(memory_wasm_sgx_in_file_dataset)))
memory_wasm_sgx_in_file_stddev = str(int(compute_stddev(memory_wasm_sgx_in_file_dataset)))
l.append("|**in-file (untrusted)**|" + \
memory_native_in_file_average + \
" (σ = " + memory_native_in_file_stddev + ")|" + \
memory_sgx_in_file_average + \
" (σ = " + memory_sgx_in_file_stddev + ")|" + \
memory_wasm_in_file_average + \
" (σ = " + memory_wasm_in_file_stddev + ")|" + \
memory_wasm_sgx_in_file_average + \
" (σ = " + memory_wasm_sgx_in_file_stddev + ")|")
l.append("|**in-file (trusted)**|" + \
"|" + \
memory_sgx_enclave_size + " (fixed)|" + \
"|" + \
memory_wasm_sgx_enclave_size + " (fixed)|")
display(Markdown('\n'.join(l)))
display(Markdown("## Size of the binary [kB]"))
l = []
l.append("||Native|SGX-LKL|WAMR|TWINE|")
l.append("|:-|-|-|-|-|")
size_native_average = to_kb_string(int(compute_mean(size_native_dataset)))
size_native_stddev = str(int(compute_stddev(size_native_dataset) / 1024))
size_sgx_lkl_bin_average = to_kb_string(int(compute_mean(size_sgx_lkl_bin_dataset)))
size_sgx_lkl_bin_stddev = str(int(compute_stddev(size_sgx_lkl_bin_dataset) / 1024))
size_wasm_bin_average = to_kb_string(int(compute_mean(size_wasm_bin_dataset)))
size_wasm_bin_stddev = str(int(compute_stddev(size_wasm_bin_dataset) / 1024))
size_wasm_sgx_bin_average = to_kb_string(int(compute_mean(size_wasm_sgx_bin_dataset)))
size_wasm_sgx_bin_stddev = str(int(compute_stddev(size_wasm_sgx_bin_dataset) / 1024))
l.append("|**Executable**|" + \
size_native_average + \
" (σ = " + size_native_stddev + ")|" + \
size_sgx_lkl_bin_average + \
" (σ = " + size_sgx_lkl_bin_stddev + ")|" + \
size_wasm_bin_average + \
" (σ = " + size_wasm_bin_stddev + ")|" + \
size_wasm_sgx_bin_average + \
" (σ = " + size_wasm_sgx_bin_stddev + ")|")
size_sgx_lkl_enclave_average = to_kb_string(int(compute_mean(size_sgx_lkl_enclave_dataset)))
size_sgx_lkl_enclave_stddev = str(int(compute_stddev(size_sgx_lkl_enclave_dataset) / 1024))
size_wasm_sgx_enclave_average = to_kb_string(int(compute_mean(size_wasm_sgx_enclave_dataset)))
size_wasm_sgx_enclave_stddev = str(int(compute_stddev(size_wasm_sgx_enclave_dataset) / 1024))
l.append("|**Enclave**||" + \
size_sgx_lkl_enclave_average + \
" (σ = " + size_sgx_lkl_enclave_stddev + ")||" + \
size_wasm_sgx_enclave_average + \
" (σ = " + size_wasm_sgx_enclave_stddev + ")|")
size_sgx_lkl_disk_image_average = to_kb_string(int(compute_mean(size_sgx_lkl_disk_image_dataset)))
size_sgx_lkl_disk_image_stddev = str(int(compute_stddev(size_sgx_lkl_disk_image_dataset) / 1024))
l.append("|**Disk image**||" + \
size_sgx_lkl_disk_image_average + \
" (σ = " + size_sgx_lkl_disk_image_stddev + ")||||")
size_wasm_average = to_kb_string(int(compute_mean(size_wasm_dataset)))
size_wasm_stddev = str(int(compute_stddev(size_wasm_dataset) / 1024))
size_wasm_average = to_kb_string(int(compute_mean(size_wasm_dataset)))
size_wasm_stddev = str(int(compute_stddev(size_wasm_dataset) / 1024))
l.append("|**Wasm**|||" + \
size_wasm_average + \
" (σ = " + size_wasm_stddev + ")|" + \
size_wasm_average + \
" (σ = " + size_wasm_stddev + ")|")
size_wasm_aot_average = to_kb_string(int(compute_mean(size_wasm_aot_dataset)))
size_wasm_aot_stddev = str(int(compute_stddev(size_wasm_aot_dataset) / 1024))
size_wasm_aot_average = to_kb_string(int(compute_mean(size_wasm_aot_dataset)))
size_wasm_aot_stddev = str(int(compute_stddev(size_wasm_aot_dataset) / 1024))
l.append("|**Wasm (AoT)**|||" + \
size_wasm_aot_average + \
" (σ = " + size_wasm_aot_stddev + ")|" + \
size_wasm_aot_average + \
" (σ = " + size_wasm_aot_stddev + ")|")
display(Markdown('\n'.join(l)))
#
## Additional values used in the paper
#
launch_time_twine_vs_sgx = r2(compute_mean(launch_time_sgx_dataset) / compute_mean(launch_time_wasm_sgx_dataset))
display(Markdown("launch_time_twine_vs_sgx: " + launch_time_twine_vs_sgx + "x\n"))
#
## Export to LaTeX
#
f = open("cost-factors-export.tex", "w")
f.write(f"\def\compilationNativeAverage{{{compilation_native_average}}}\n")
f.write(f"\def\compilationSgxAverage{{{compilation_sgx_average}}}\n")
f.write(f"\def\compilationWasmBinAverage{{{compilation_wasm_bin_average}}}\n")
f.write(f"\def\compilationWasmSgxBinAverage{{{compilation_wasm_sgx_bin_average}}}\n")
f.write(f"\def\compilationWasmAverage{{{compilation_wasm_average}}}\n")
f.write(f"\def\optimizationSgxLklDiskImageAverage{{{optimization_sgx_lkl_disk_image_average}}}\n")
f.write(f"\def\optimizationWasmAotAverage{{{optimization_wasm_aot_average}}}\n")
f.write(f"\def\launchTimeNativeAverage{{{launch_time_native_average}}}\n")
f.write(f"\def\launchTimeSgxAverage{{{launch_time_sgx_average}}}\n")
f.write(f"\def\launchTimeWasmAverage{{{launch_time_wasm_average}}}\n")
f.write(f"\def\launchTimeWasmSgxAverage{{{launch_time_wasm_sgx_average}}}\n")
f.write(f"\def\memoryNativeInMemoryAverage{{{memory_native_in_memory_average}}}\n")
f.write(f"\def\memorySgxInMemoryAverage{{{memory_sgx_in_memory_average}}}\n")
f.write(f"\def\memoryWasmInMemoryAverage{{{memory_wasm_in_memory_average}}}\n")
f.write(f"\def\memoryWasmSgxInMemoryAverage{{{memory_wasm_sgx_in_memory_average}}}\n")
f.write(f"\def\memorySgxEnclaveSize{{{memory_sgx_enclave_size}}}\n")
f.write(f"\def\memoryWasmSgxEnclaveSize{{{memory_wasm_sgx_enclave_size}}}\n")
f.write(f"\def\sizeNativeAverage{{{size_native_average}}}\n")
f.write(f"\def\sizeSgxLklBinAverage{{{size_sgx_lkl_bin_average}}}\n")
f.write(f"\def\sizeWasmBinAverage{{{size_wasm_bin_average}}}\n")
f.write(f"\def\sizeWasmSgxBinAverage{{{size_wasm_sgx_bin_average}}}\n")
f.write(f"\def\sizeSgxLklEnclaveAverage{{{size_sgx_lkl_enclave_average}}}\n")
f.write(f"\def\sizeWasmSgxEnclaveAverage{{{size_wasm_sgx_enclave_average}}}\n")
f.write(f"\def\sizeSgxLklDiskImageAverage{{{size_sgx_lkl_disk_image_average}}}\n")
f.write(f"\def\sizeWasmAverage{{{size_wasm_average}}}\n")
f.write(f"\def\sizeWasmAotAverage{{{size_wasm_aot_average}}}\n")
f.write(f"\def\launchTimeTwineVsSgx{{{launch_time_twine_vs_sgx}}}\n")
f.close()
```
| github_jupyter |
# LeNet Lab Solution

Source: Yan LeCun
## Load Data
Load the MNIST data, which comes pre-loaded with TensorFlow.
You do not need to modify this section.
```
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("MNIST_data/", reshape=False)
X_train, y_train = mnist.train.images, mnist.train.labels
X_validation, y_validation = mnist.validation.images, mnist.validation.labels
X_test, y_test = mnist.test.images, mnist.test.labels
assert(len(X_train) == len(y_train))
assert(len(X_validation) == len(y_validation))
assert(len(X_test) == len(y_test))
print()
print("Image Shape: {}".format(X_train[0].shape))
print()
print("Training Set: {} samples".format(len(X_train)))
print("Validation Set: {} samples".format(len(X_validation)))
print("Test Set: {} samples".format(len(X_test)))
```
The MNIST data that TensorFlow pre-loads comes as 28x28x1 images.
However, the LeNet architecture only accepts 32x32xC images, where C is the number of color channels.
In order to reformat the MNIST data into a shape that LeNet will accept, we pad the data with two rows of zeros on the top and bottom, and two columns of zeros on the left and right (28+2+2 = 32).
You do not need to modify this section.
```
import numpy as np
# Pad images with 0s
X_train = np.pad(X_train, ((0,0),(2,2),(2,2),(0,0)), 'constant')
X_validation = np.pad(X_validation, ((0,0),(2,2),(2,2),(0,0)), 'constant')
X_test = np.pad(X_test, ((0,0),(2,2),(2,2),(0,0)), 'constant')
print("Updated Image Shape: {}".format(X_train[0].shape))
```
## Visualize Data
View a sample from the dataset.
You do not need to modify this section.
```
import random
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
index = random.randint(0, len(X_train))
image = X_train[index].squeeze()
plt.figure(figsize=(1,1))
plt.imshow(image, cmap="gray")
print(y_train[index])
```
## Preprocess Data
Shuffle the training data.
You do not need to modify this section.
```
from sklearn.utils import shuffle
X_train, y_train = shuffle(X_train, y_train)
```
## Setup TensorFlow
The `EPOCH` and `BATCH_SIZE` values affect the training speed and model accuracy.
You do not need to modify this section.
```
import tensorflow as tf
EPOCHS = 10
BATCH_SIZE = 128
```
## SOLUTION: Implement LeNet-5
Implement the [LeNet-5](http://yann.lecun.com/exdb/lenet/) neural network architecture.
This is the only cell you need to edit.
### Input
The LeNet architecture accepts a 32x32xC image as input, where C is the number of color channels. Since MNIST images are grayscale, C is 1 in this case.
### Architecture
**Layer 1: Convolutional.** The output shape should be 28x28x6.
**Activation.** Your choice of activation function.
**Pooling.** The output shape should be 14x14x6.
**Layer 2: Convolutional.** The output shape should be 10x10x16.
**Activation.** Your choice of activation function.
**Pooling.** The output shape should be 5x5x16.
**Flatten.** Flatten the output shape of the final pooling layer such that it's 1D instead of 3D. The easiest way to do is by using `tf.contrib.layers.flatten`, which is already imported for you.
**Layer 3: Fully Connected.** This should have 120 outputs.
**Activation.** Your choice of activation function.
**Layer 4: Fully Connected.** This should have 84 outputs.
**Activation.** Your choice of activation function.
**Layer 5: Fully Connected (Logits).** This should have 10 outputs.
### Output
Return the result of the 2nd fully connected layer.
```
from tensorflow.contrib.layers import flatten
def LeNet(x):
# Hyperparameters
mu = 0
sigma = 0.1
# SOLUTION: Layer 1: Convolutional. Input = 32x32x1. Output = 28x28x6.
conv1_W = tf.Variable(tf.truncated_normal(shape=(5, 5, 1, 6), mean = mu, stddev = sigma))
conv1_b = tf.Variable(tf.zeros(6))
conv1 = tf.nn.conv2d(x, conv1_W, strides=[1, 1, 1, 1], padding='VALID') + conv1_b
# SOLUTION: Activation.
conv1 = tf.nn.relu(conv1)
# SOLUTION: Pooling. Input = 28x28x6. Output = 14x14x6.
conv1 = tf.nn.max_pool(conv1, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='VALID')
# SOLUTION: Layer 2: Convolutional. Output = 10x10x16.
conv2_W = tf.Variable(tf.truncated_normal(shape=(5, 5, 6, 16), mean = mu, stddev = sigma))
conv2_b = tf.Variable(tf.zeros(16))
conv2 = tf.nn.conv2d(conv1, conv2_W, strides=[1, 1, 1, 1], padding='VALID') + conv2_b
# SOLUTION: Activation.
conv2 = tf.nn.relu(conv2)
# SOLUTION: Pooling. Input = 10x10x16. Output = 5x5x16.
conv2 = tf.nn.max_pool(conv2, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='VALID')
# SOLUTION: Flatten. Input = 5x5x16. Output = 400.
fc0 = flatten(conv2)
# SOLUTION: Layer 3: Fully Connected. Input = 400. Output = 120.
fc1_W = tf.Variable(tf.truncated_normal(shape=(400, 120), mean = mu, stddev = sigma))
fc1_b = tf.Variable(tf.zeros(120))
fc1 = tf.matmul(fc0, fc1_W) + fc1_b
# SOLUTION: Activation.
fc1 = tf.nn.relu(fc1)
# SOLUTION: Layer 4: Fully Connected. Input = 120. Output = 84.
fc2_W = tf.Variable(tf.truncated_normal(shape=(120, 84), mean = mu, stddev = sigma))
fc2_b = tf.Variable(tf.zeros(84))
fc2 = tf.matmul(fc1, fc2_W) + fc2_b
# SOLUTION: Activation.
fc2 = tf.nn.relu(fc2)
# SOLUTION: Layer 5: Fully Connected. Input = 84. Output = 10.
fc3_W = tf.Variable(tf.truncated_normal(shape=(84, 10), mean = mu, stddev = sigma))
fc3_b = tf.Variable(tf.zeros(10))
logits = tf.matmul(fc2, fc3_W) + fc3_b
return logits
```
## Features and Labels
Train LeNet to classify [MNIST](http://yann.lecun.com/exdb/mnist/) data.
`x` is a placeholder for a batch of input images.
`y` is a placeholder for a batch of output labels.
You do not need to modify this section.
```
x = tf.placeholder(tf.float32, (None, 32, 32, 1))
y = tf.placeholder(tf.int32, (None))
one_hot_y = tf.one_hot(y, 10)
```
## Training Pipeline
Create a training pipeline that uses the model to classify MNIST data.
You do not need to modify this section.
```
rate = 0.001
logits = LeNet(x)
cross_entropy = tf.nn.softmax_cross_entropy_with_logits(logits, one_hot_y)
loss_operation = tf.reduce_mean(cross_entropy)
optimizer = tf.train.AdamOptimizer(learning_rate = rate)
training_operation = optimizer.minimize(loss_operation)
```
## Model Evaluation
Evaluate how well the loss and accuracy of the model for a given dataset.
You do not need to modify this section.
```
correct_prediction = tf.equal(tf.argmax(logits, 1), tf.argmax(one_hot_y, 1))
accuracy_operation = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
saver = tf.train.Saver()
def evaluate(X_data, y_data):
num_examples = len(X_data)
total_accuracy = 0
sess = tf.get_default_session()
for offset in range(0, num_examples, BATCH_SIZE):
batch_x, batch_y = X_data[offset:offset+BATCH_SIZE], y_data[offset:offset+BATCH_SIZE]
accuracy = sess.run(accuracy_operation, feed_dict={x: batch_x, y: batch_y})
total_accuracy += (accuracy * len(batch_x))
return total_accuracy / num_examples
```
## Train the Model
Run the training data through the training pipeline to train the model.
Before each epoch, shuffle the training set.
After each epoch, measure the loss and accuracy of the validation set.
Save the model after training.
You do not need to modify this section.
```
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
num_examples = len(X_train)
print("Training...")
print()
for i in range(EPOCHS):
X_train, y_train = shuffle(X_train, y_train)
for offset in range(0, num_examples, BATCH_SIZE):
end = offset + BATCH_SIZE
batch_x, batch_y = X_train[offset:end], y_train[offset:end]
sess.run(training_operation, feed_dict={x: batch_x, y: batch_y})
validation_accuracy = evaluate(X_validation, y_validation)
print("EPOCH {} ...".format(i+1))
print("Validation Accuracy = {:.3f}".format(validation_accuracy))
print()
saver.save(sess, 'lenet')
print("Model saved")
```
## Evaluate the Model
Once you are completely satisfied with your model, evaluate the performance of the model on the test set.
Be sure to only do this once!
If you were to measure the performance of your trained model on the test set, then improve your model, and then measure the performance of your model on the test set again, that would invalidate your test results. You wouldn't get a true measure of how well your model would perform against real data.
You do not need to modify this section.
```
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('.'))
test_accuracy = evaluate(X_test, y_test)
print("Test Accuracy = {:.3f}".format(test_accuracy))
```
| github_jupyter |
# 4. Predict-Image-Similar-FCNN-Binary[batch]
For landmark-recognition-2019 algorithm validation
## Run name
```
import time
project_name = 'Google-LandMark-Rec2019'
model_name = 'Google-LandMark-Rec2019_3-Image-Similar-FCNN-Binary_20190511-093300_8406'
step_name = '4-Predict[batch]_%s' % model_name
time_str = time.strftime("%Y%m%d-%H%M%S", time.localtime())
run_name = step_name + '_' + time_str
print('run_name: ' + run_name)
t0 = time.time()
```
## Important params
```
import multiprocessing
cpu_amount = multiprocessing.cpu_count()
print('cpu_amount: ', cpu_amount)
small_debug_rows = 10
big_debug_rows = 20
predict_batch_size = 10
print('small_debug_rows:\t', small_debug_rows)
print('big_debug_rows:\t', big_debug_rows)
print('predict_batch_size:\t', predict_batch_size)
```
## Import PKGs
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
%matplotlib inline
from IPython.display import display
import os
import sys
import gc
import math
import shutil
import zipfile
import pickle
import h5py
from tqdm import tqdm
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix, accuracy_score
import keras
from keras.utils import Sequence
from keras.layers import *
from keras.models import *
from keras.applications import *
from keras.optimizers import *
from keras.regularizers import *
from keras.preprocessing.image import *
from keras.applications.inception_v3 import preprocess_input
import tensorflow as tf
import keras.backend.tensorflow_backend as KTF
config = tf.ConfigProto()
config.gpu_options.per_process_gpu_memory_fraction = 0.5
session = tf.Session(config=config)
KTF.set_session(session )
```
## Project folders
```
cwd = os.getcwd()
feature_folder = os.path.join(cwd, 'feature')
input_folder = os.path.join(cwd, 'input')
output_folder = os.path.join(cwd, 'output')
model_folder = os.path.join(cwd, 'model')
org_train_folder = os.path.join(input_folder, 'org_train')
org_test_folder = os.path.join(input_folder, 'org_test')
train_folder = os.path.join(input_folder, 'data_train')
val_folder = os.path.join(input_folder, 'data_val')
test_folder = os.path.join(input_folder, 'data_test')
test_sub_folder = os.path.join(test_folder, 'test')
vgg16_feature_file = os.path.join(feature_folder, 'feature_wrapper_171023.h5')
train_csv_file = os.path.join(input_folder, 'train.csv')
test_csv_file = os.path.join(input_folder, 'test.csv')
sample_submission_folder = os.path.join(input_folder, 'sample_submission.csv')
print(vgg16_feature_file)
print(train_csv_file)
print(test_csv_file)
print(sample_submission_folder)
```
## Functions
```
def pickle_dump(data, file):
with open(file, 'wb') as f:
pickle.dump(data, f)
def pickle_load(file):
with open(file, 'rb') as f:
data = pickle.load(f)
return data
a = list(range(10))
print(a)
demo_file = os.path.join(os.getcwd(), 'temp', 'pickle_demo.pkl')
print(demo_file)
pickle_dump(a, demo_file)
new_a = pickle_load(demo_file)
print(new_a)
%%time
def get_whole_classes_label(class_indices_data, item_classes_data):
indices2class = {}
for i, key in enumerate(class_indices_data.keys()):
# print(key, class_indices_data[key])
indices2class[class_indices_data[key]] = key
# if i >= 5:
# break
# for i, key in enumerate(class_indices_data.keys()):
# print(key, class_indices_data[key], '-->', class_indices_data[key], indices2class[class_indices_data[key]])
# if i >= 10:
# break
print(item_classes_data.shape)
whole_classes_data = np.zeros(item_classes_data.shape)
for i, class_index in enumerate(item_classes_data):
whole_classes_data[i] = indices2class[class_index]
# print(i, class_index, '-->', indices2class[class_index])
# if i >= 10:
# break
return whole_classes_data
# get_whole_classes_label(class_indices_train, item_classes_train)
```
## Load feature
```
!ls ./feature -hl
def load_feature(data_set_name, model_name, date_str, feature_amount):
# data_set_name = 'train'
# model_name = 'VGG19'
# date_str = '171023'
x_data_arr = []
classes_data_arr = []
index_data_arr = []
for i in range(feature_amount):
folder_name = 'data_%s_%02d' % (data_set_name, i)
class_indices_file = os.path.join(cwd, 'feature', 'feature_{0}_{1}_{2}_class_indices.pkl'.format(model_name, folder_name, date_str))
h5py_file_name = os.path.join(cwd, 'feature', 'feature_{0}_{1}_{2}.h5'.format(model_name, folder_name, date_str))
if not os.path.exists(class_indices_file):
print('File not exists', class_indices_file)
continue
if not os.path.exists(h5py_file_name):
print('File not exists', h5py_file_name)
continue
print(class_indices_file)
print(h5py_file_name)
class_indices_data = pickle_load(class_indices_file)
print(len(class_indices_data))
with h5py.File(h5py_file_name, 'r') as h:
item_x_data = np.array(h['x_data_%s_%02d' % (data_set_name, i)])
item_classes_data = np.array(h['classes_data_%s_%02d' % (data_set_name, i)])
item_index_data = np.array(h['index_data_%s_%02d' % (data_set_name, i)])
print(item_x_data.shape)
x_data_arr.append(item_x_data)
item_y_data = get_whole_classes_label(class_indices_data, item_classes_data)
print(item_y_data.shape, item_y_data[:10])
classes_data_arr.append(item_y_data)
index_data_arr.append(item_index_data)
x_data = np.concatenate(x_data_arr, axis=0)
y_data = np.concatenate(classes_data_arr, axis=0)
idx_data = np.concatenate(index_data_arr, axis=0)
print('*' * 60)
print(x_data.shape)
print(y_data.shape)
print(idx_data.shape)
return x_data, y_data, idx_data
x_train, y_train, idx_train = load_feature('train', 'VGG19', '171023', 20)
x_val, y_val, idx_val = load_feature('val', 'VGG19', '171023', 4)
```
## Load model
```
model_file = os.path.join(model_folder, '%s.h5' % model_name)
print(model_file)
model = load_model(model_file)
model.summary()
```
## Predict
### single item
```
%%time
idx = 0
item_main_x = np.array([x_train[idx]]*x_train.shape[0])
item_x_train = {
'main_input': item_main_x,
'library_input': x_train
}
y_proba = model.predict(item_x_train, batch_size=1024)
print(int(y_train[idx]), np.argmax(y_proba), '-->', int(y_train[np.argmax(y_proba)]), [int(y_train[item[0]]) for item in np.argsort(y_proba, axis=0)[-10:]])
```
## batch predict
```
%%time
batch_libary_x = np.tile(x_train, (3, 1))
print(batch_libary_x.shape)
def get_batch_pred(batch_x_data, batch_libary_x, topn, batch_size=1024):
print(batch_x_data.shape)
print(libary_x.shape)
# def get_single_pred(idx, main_x, libary_x, topn, batch_size=1024):
batch_main_x_arr = []
for item_x_data in batch_x_data:
expand_np = np.expand_dims(item_x_data, axis=0)
tile_np = np.tile(expand_np, (libary_x.shape[0], 1))
batch_main_x_arr.append(tile_np)
batch_main_x = np.concatenate(batch_main_x_arr, axis=0)
print(batch_main_x.shape)
item_x = {
'main_input': batch_main_x,
'library_input': batch_libary_x
}
# y_proba = model.predict(item_x, batch_size=batch_size)
# top1_pred = int(y_train[np.argmax(y_proba)])
# topn_pred_arr = [int(y_train[item[0]]) for item in np.argsort(y_proba, axis=0)[-topn:]]
# weighted_topn_pred_arr, class_score_arr = get_weight_topn(y_proba, topn, y_train)
# weighted_top1_pred = weighted_topn_pred_arr[-1]
# return top1_pred, topn_pred_arr, weighted_top1_pred, weighted_topn_pred_arr, class_score_arr
idx = 0
topn = 10
# train
print(get_batch_pred(x_train[:3], batch_libary_x, topn, batch_size=1024))
# val
# print(int(y_val[idx]), get_single_pred(idx, x_val, x_train, topn, batch_size=1024))
np.expand_dims(x_train[0], axis=0).shape
a = np.repeat(x_train[:2], 10000, axis=0)
print(a.shape)
np.tile?
orig = np.array([[1, 2], [3, 4], [5, 6]])
print(orig.shape)
print(orig)
print('*' * 80)
a = np.tile(orig, (3, 1))
print(a, a.shape)
x_train[:1].shape
assert 0
```
### topn
```
topn = 10
amount = small_debug_rows
top1_count = 0
topn_count = 0
for idx in range(amount):
item_main_x = np.array([x_train[idx]]*x_train.shape[0])
item_x_train = {
'main_input': item_main_x,
'library_input': x_train
}
y_proba = model.predict(item_x_train, batch_size=1024)
item_y_train = int(y_train[idx])
top1_pred = int(y_train[np.argmax(y_proba)])
topn_pred_arr = [int(y_train[item[0]]) for item in np.argsort(y_proba, axis=0)[-topn:]]
if item_y_train == top1_pred:
top1_count += 1
if item_y_train in topn_pred_arr:
topn_count += 1
print(item_y_train, '-->', top1_pred, topn_pred_arr)
print('*' * 80)
print(top1_count, '%.2f' % (top1_count/amount))
print(topn_count, '%.2f' % (topn_count/amount))
```
### weighted topn
```
def get_weight_topn(y_proba, topn, y_data):
y_proba_argsorted = [(y_data[item[0]], y_proba[item[0]][0]) for item in np.argsort(y_proba, axis=0)[-topn:]]
class_score_dict = {}
for item_class, item_proba in y_proba_argsorted:
if item_class in class_score_dict:
class_score_dict[item_class] += item_proba
else:
class_score_dict[item_class] = item_proba
class_score_arr = list(class_score_dict.items())
class_score_arr = sorted(class_score_arr, key=lambda x: x[1])
topn_pred_arr = [int(item[0]) for item in class_score_arr]
class_score_arr = [round(item[1], 3) for item in class_score_arr]
return topn_pred_arr, class_score_arr
y_proba = np.array([[0.1], [0.2], [0.3], [0.2], [0.2], [0.2]])
topn = 6
y_data = np.array([0, 0, 1, 1, 2, 2])
print(y_proba, type(y_proba))
print(topn)
print(y_data, type(y_data))
get_weight_topn(y_proba, topn, y_data)
topn = 10
amount = small_debug_rows
top1_count = 0
topn_1_count = 0
topn_count = 0
for idx in range(amount):
item_main_x = np.array([x_train[idx]]*x_train.shape[0])
item_x_train = {
'main_input': item_main_x,
'library_input': x_train
}
y_proba = model.predict(item_x_train, batch_size=1024)
item_y_train = int(y_train[idx])
top1_pred = int(y_train[np.argmax(y_proba)])
# topn_pred_arr = [y_train[item[0]] for item in np.argsort(y_proba, axis=0)[-topn:]]
topn_pred_arr, class_score_arr = get_weight_topn(y_proba, topn, y_train)
topn_1_pred = topn_pred_arr[-1]
if item_y_train == top1_pred:
top1_count += 1
if item_y_train == topn_1_pred:
topn_1_count += 1
if item_y_train in topn_pred_arr:
topn_count += 1
print(item_y_train, '-->', top1_pred, '\t', topn_1_pred, topn_pred_arr, class_score_arr)
print('*' * 80)
print(top1_count, '%.2f' % (top1_count/amount))
print(topn_1_count, '%.2f' % (topn_1_count/amount))
print(topn_count, '%.2f' % (topn_count/amount))
```
### encapsolution
```
%%time
def get_single_pred(idx, main_x, libary_x, topn, batch_size=1024):
time_arr = []
time_arr.append(time.strftime("%Y%m%d-%H%M%S", time.localtime()))
# item_main_x = np.array([main_x[idx]]*libary_x.shape[0])
expand_np = np.expand_dims(main_x[idx], axis=0)
item_main_x = np.tile(expand_np, (libary_x.shape[0], 1))
time_arr.append(time.strftime("%Y%m%d-%H%M%S", time.localtime()))
item_x = {
'main_input': item_main_x,
'library_input': libary_x
}
time_arr.append(time.strftime("%Y%m%d-%H%M%S", time.localtime()))
y_proba = model.predict(item_x, batch_size=batch_size)
time_arr.append(time.strftime("%Y%m%d-%H%M%S", time.localtime()))
top1_pred = int(y_train[np.argmax(y_proba)])
topn_pred_arr = [int(y_train[item[0]]) for item in np.argsort(y_proba, axis=0)[-topn:]]
weighted_topn_pred_arr, class_score_arr = get_weight_topn(y_proba, topn, y_train)
weighted_top1_pred = weighted_topn_pred_arr[-1]
time_arr.append(time.strftime("%Y%m%d-%H%M%S", time.localtime()))
# for item in time_arr:
# print(item)
return top1_pred, topn_pred_arr, weighted_top1_pred, weighted_topn_pred_arr, class_score_arr
idx = 0
topn = 10
# train
print(int(y_train[idx]), get_single_pred(idx, x_train, x_train, topn, batch_size=1024))
# val
# print(int(y_val[idx]), get_single_pred(idx, x_val, x_train, topn, batch_size=1024))
topn = 10
amount = big_debug_rows
top1_count = 0
topn_1_count = 0
topn_count = 0
for idx in range(amount):
top1_pred, topn_pred_arr, weighted_top1_pred, weighted_topn_pred_arr, class_score_arr = get_single_pred(idx, x_train, x_train, topn, batch_size=1024)
item_y_data = int(y_train[idx])
if item_y_data == top1_pred:
top1_count += 1
if item_y_data == weighted_top1_pred:
topn_1_count += 1
if item_y_data in topn_pred_arr:
topn_count += 1
print(item_y_data, '-->', top1_pred, weighted_top1_pred, '\t', weighted_topn_pred_arr, class_score_arr)
print('*' * 80)
print(top1_count, '%.2f' % (top1_count/amount))
print(topn_1_count, '%.2f' % (topn_1_count/amount))
print(topn_count, '%.2f' % (topn_count/amount))
topn = 10
amount = big_debug_rows
top1_count = 0
topn_1_count = 0
topn_count = 0
for idx in range(amount):
top1_pred, topn_pred_arr, weighted_top1_pred, weighted_topn_pred_arr, class_score_arr = get_single_pred(idx, x_val, x_train, topn, batch_size=1024)
item_y_data = int(y_val[idx])
if item_y_data == top1_pred:
top1_count += 1
if item_y_data == weighted_top1_pred:
topn_1_count += 1
if item_y_data in topn_pred_arr:
topn_count += 1
print(item_y_data, '-->', top1_pred, weighted_top1_pred, '\t', weighted_topn_pred_arr, class_score_arr)
print('*' * 80)
print(top1_count, '%.2f' % (top1_count/amount))
print(topn_1_count, '%.2f' % (topn_1_count/amount))
print(topn_count, '%.2f' % (topn_count/amount))
acc10000 = int(topn_count/amount*10000)
# acc10000 = 10000 # for test
# print(acc10000)
if acc10000 == 10000:
run_name_acc = '%s_%05d' % (run_name, acc10000)
else:
run_name_acc = '%s_%04d' % (run_name, acc10000)
print(run_name_acc)
print('Time elapsed: %.1fs' % (time.time() - t0))
print(run_name_acc)
```
| github_jupyter |
# Regression Week 1: Simple Linear Regression
In this notebook we will use data on house sales in King County to predict house prices using simple (one input) linear regression. You will:
* Write a function to compute the Simple Linear Regression weights using the closed form solution
* Write a function to make predictions of the output given the input feature
* Turn the regression around to predict the input given the output
* Compare two different models for predicting house prices
In this notebook you will be provided with some already complete code as well as some code that you should complete yourself in order to answer quiz questions. The code we provide to complete is optional and is there to assist you with solving the problems but feel free to ignore the helper code and write your own.
```
# import graphlab
import pandas as pd
import numpy as np
```
# Load house sales data
Dataset is from house sales in King County, the region where the city of Seattle, WA is located.
```
dtype_dict = {'bathrooms':float, 'waterfront':int, 'sqft_above':int, 'sqft_living15':float, 'grade':int, 'yr_renovated':int,
'price':float, 'bedrooms':float, 'zipcode':str, 'long':float, 'sqft_lot15':float, 'sqft_living':float,
'floors':str, 'condition':int, 'lat':float, 'date':str, 'sqft_basement':int, 'yr_built':int, 'id':str,
'sqft_lot':int, 'view':int}
type(dtype_dict)
# sales = graphlab.SFrame('kc_house_data.gl/')
df_house_data = pd.read_csv("../../../../data/coursera_regression/kc_house_data.csv", dtype= dtype_dict)
df_house_train = pd.read_csv("../../../../data/coursera_regression/kc_house_train_data.csv", dtype= dtype_dict)
df_house_test = pd.read_csv("../../../../data/coursera_regression/kc_house_test_data.csv", dtype= dtype_dict)
df_house_data.head()
df_house_train.head()
df_house_test.head()
df_house_data.shape, df_house_train.shape, df_house_test.shape
# Let's compute the mean of the House Prices in King County in 2 different ways.
np.mean(df_house_data["price"]), np.mean(df_house_train["price"]), np.mean(df_house_test["price"])
# #Mean Value
(17384*539366.62793373212 + 4229*543054.0430361788)/(17384+4229)
```
# Build a generic simple linear regression function
```
def simple_linear_regression(input_feature, output):
# compute the sum of input_feature and output
sum_x = input_feature.sum()
sum_y = output.sum()
# compute the product of the output and the input_feature and its sum
prod_xy = (output * input_feature).sum()
# compute the squared value of the input_feature and its sum
prod_xx = (input_feature * input_feature).sum()
N = (output.size)
# use the formula for the slope
slope = ((prod_xy-((sum_x*sum_y)/N)) / (prod_xx -((sum_x*sum_x)/N)))
# use the formula for the intercept
intercept = sum_y/N - (slope*(sum_x / N))
return (intercept, slope)
sqft_intercept, sqft_slope = simple_linear_regression(df_house_train["sqft_living"], df_house_train["price"])
print("Intercept: ", str(sqft_intercept))
print("Slope: ", str(sqft_slope))
```
# Predicting Values
```
def get_regression_predictions(input_feature, intercept, slope):
# calculate the predicted values:
predicted_values = intercept + slope*input_feature
return predicted_values
```
Now that we can calculate a prediction given the slope and intercept let's make a prediction. Use (or alter) the following to find out the estimated price for a house with 2650 squarefeet according to the squarefeet model we estiamted above.
**Quiz Question: Using your Slope and Intercept from (4), What is the predicted price for a house with 2650 sqft?**
```
my_house_sqft = 2650
estimated_price = get_regression_predictions(my_house_sqft, sqft_intercept, sqft_slope)
print("The estimated price for a house with %d squarefeet is $%.2f" % (my_house_sqft, estimated_price))
```
# Residual Sum of Squares
Now that we have a model and can make predictions let's evaluate our model using Residual Sum of Squares (RSS). Recall that RSS is the sum of the squares of the residuals and the residuals is just a fancy word for the difference between the predicted output and the true output.
Complete the following (or write your own) function to compute the RSS of a simple linear regression model given the input_feature, output, intercept and slope:
```
def get_residual_sum_of_squares(input_feature, output, intercept, slope):
# First get the predictions
predictions = get_regression_predictions(input_feature, intercept, slope)
# then compute the residuals (since we are squaring it doesn't matter which order you subtract)
# square the residuals and add them up
RSS = ((output - predictions)**2).sum()
return(RSS)
```
Let's test our get_residual_sum_of_squares function by applying it to the test model where the data lie exactly on a line. Since they lie exactly on a line the residual sum of squares should be zero!
```
print(get_residual_sum_of_squares(df_house_train["sqft_living"], df_house_train["price"], sqft_intercept, sqft_slope)) # should be 0.0
```
Now use your function to calculate the RSS on training data from the squarefeet model calculated above.
**Quiz Question: According to this function and the slope and intercept from the squarefeet model What is the RSS for the simple linear regression using squarefeet to predict prices on TRAINING data?**
```
rss_prices_on_sqft = get_residual_sum_of_squares(df_house_train["sqft_living"], df_house_train["price"], sqft_intercept, sqft_slope)
print("The RSS of predicting Prices based on Square Feet is : ", rss_prices_on_sqft)
```
# Predict the squarefeet given price
What if we want to predict the squarefoot given the price? Since we have an equation y = a + b\*x we can solve the function for x. So that if we have the intercept (a) and the slope (b) and the price (y) we can solve for the estimated squarefeet (x).
Complete the following function to compute the inverse regression estimate, i.e. predict the input_feature given the output.
```
def inverse_regression_predictions(output, intercept, slope):
# solve output = intercept + slope*input_feature for input_feature. Use this equation to compute the inverse predictions:
estimated_feature = ((output - intercept) / slope)
return estimated_feature
```
Now that we have a function to compute the squarefeet given the price from our simple regression model let's see how big we might expect a house that costs $800,000 to be.
**Quiz Question: According to this function and the regression slope and intercept from (3) what is the estimated square-feet for a house costing $800,000?**
```
my_house_price = 800000
estimated_squarefeet = inverse_regression_predictions(my_house_price, sqft_intercept, sqft_slope)
print("The estimated squarefeet for a house worth $%.2f is %d", (my_house_price, estimated_squarefeet))
```
# New Model: estimate prices from bedrooms
We have made one model for predicting house prices using squarefeet, but there are many other features in the sales SFrame.
Use your simple linear regression function to estimate the regression parameters from predicting Prices based on number of bedrooms. Use the training data!
```
# Estimate the slope and intercept for predicting 'price' based on 'bedrooms'
sqft_intercept, sqft_slope = simple_linear_regression(df_house_train['bedrooms'], df_house_train['price'])
print(sqft_intercept, sqft_slope)
```
# Test your Linear Regression Algorithm
Now we have two models for predicting the price of a house. How do we know which one is better? Calculate the RSS on the TEST data (remember this data wasn't involved in learning the model). Compute the RSS from predicting prices using bedrooms and from predicting prices using squarefeet.
**Quiz Question: Which model (square feet or bedrooms) has lowest RSS on TEST data? Think about why this might be the case.**
```
# Compute RSS when using bedrooms on TEST data:
sqft_intercept, sqft_slope = simple_linear_regression(df_house_train['bedrooms'], df_house_train['price'])
rss_prices_on_bedrooms = get_residual_sum_of_squares(df_house_test['bedrooms'], df_house_test['price'], sqft_intercept, sqft_slope)
print("The RSS of predicting Prices based on Bedrooms is : ",rss_prices_on_bedrooms)
# Compute RSS when using squarefeet on TEST data:
sqft_intercept, sqft_slope = simple_linear_regression(df_house_train['sqft_living'], df_house_train['price'])
rss_prices_on_sqft = get_residual_sum_of_squares(df_house_test['sqft_living'], df_house_test['price'], sqft_intercept, sqft_slope)
print("The RSS of predicting Prices based on Square Feet is : ", rss_prices_on_sqft)
```
| github_jupyter |
# Residual Networks
Welcome to the second assignment of this week! You will learn how to build very deep convolutional networks, using Residual Networks (ResNets). In theory, very deep networks can represent very complex functions; but in practice, they are hard to train. Residual Networks, introduced by [He et al.](https://arxiv.org/pdf/1512.03385.pdf), allow you to train much deeper networks than were previously practically feasible.
**In this assignment, you will:**
- Implement the basic building blocks of ResNets.
- Put together these building blocks to implement and train a state-of-the-art neural network for image classification.
## <font color='darkblue'>Updates</font>
#### If you were working on the notebook before this update...
* The current notebook is version "2a".
* You can find your original work saved in the notebook with the previous version name ("v2")
* To view the file directory, go to the menu "File->Open", and this will open a new tab that shows the file directory.
#### List of updates
* For testing on an image, replaced `preprocess_input(x)` with `x=x/255.0` to normalize the input image in the same way that the model's training data was normalized.
* Refers to "shallower" layers as those layers closer to the input, and "deeper" layers as those closer to the output (Using "shallower" layers instead of "lower" or "earlier").
* Added/updated instructions.
This assignment will be done in Keras.
Before jumping into the problem, let's run the cell below to load the required packages.
```
import numpy as np
from keras import layers
from keras.layers import Input, Add, Dense, Activation, ZeroPadding2D, BatchNormalization, Flatten, Conv2D, AveragePooling2D, MaxPooling2D, GlobalMaxPooling2D
from keras.models import Model, load_model
from keras.preprocessing import image
from keras.utils import layer_utils
from keras.utils.data_utils import get_file
from keras.applications.imagenet_utils import preprocess_input
import pydot
from IPython.display import SVG
from keras.utils.vis_utils import model_to_dot
from keras.utils import plot_model
from resnets_utils import *
from keras.initializers import glorot_uniform
import scipy.misc
from matplotlib.pyplot import imshow
%matplotlib inline
import keras.backend as K
K.set_image_data_format('channels_last')
K.set_learning_phase(1)
```
## 1 - The problem of very deep neural networks
Last week, you built your first convolutional neural network. In recent years, neural networks have become deeper, with state-of-the-art networks going from just a few layers (e.g., AlexNet) to over a hundred layers.
* The main benefit of a very deep network is that it can represent very complex functions. It can also learn features at many different levels of abstraction, from edges (at the shallower layers, closer to the input) to very complex features (at the deeper layers, closer to the output).
* However, using a deeper network doesn't always help. A huge barrier to training them is vanishing gradients: very deep networks often have a gradient signal that goes to zero quickly, thus making gradient descent prohibitively slow.
* More specifically, during gradient descent, as you backprop from the final layer back to the first layer, you are multiplying by the weight matrix on each step, and thus the gradient can decrease exponentially quickly to zero (or, in rare cases, grow exponentially quickly and "explode" to take very large values).
* During training, you might therefore see the magnitude (or norm) of the gradient for the shallower layers decrease to zero very rapidly as training proceeds:
<img src="images/vanishing_grad_kiank.png" style="width:450px;height:220px;">
<caption><center> <u> <font color='purple'> **Figure 1** </u><font color='purple'> : **Vanishing gradient** <br> The speed of learning decreases very rapidly for the shallower layers as the network trains </center></caption>
You are now going to solve this problem by building a Residual Network!
## 2 - Building a Residual Network
In ResNets, a "shortcut" or a "skip connection" allows the model to skip layers:
<img src="images/skip_connection_kiank.png" style="width:650px;height:200px;">
<caption><center> <u> <font color='purple'> **Figure 2** </u><font color='purple'> : A ResNet block showing a **skip-connection** <br> </center></caption>
The image on the left shows the "main path" through the network. The image on the right adds a shortcut to the main path. By stacking these ResNet blocks on top of each other, you can form a very deep network.
We also saw in lecture that having ResNet blocks with the shortcut also makes it very easy for one of the blocks to learn an identity function. This means that you can stack on additional ResNet blocks with little risk of harming training set performance.
(There is also some evidence that the ease of learning an identity function accounts for ResNets' remarkable performance even more so than skip connections helping with vanishing gradients).
Two main types of blocks are used in a ResNet, depending mainly on whether the input/output dimensions are same or different. You are going to implement both of them: the "identity block" and the "convolutional block."
### 2.1 - The identity block
The identity block is the standard block used in ResNets, and corresponds to the case where the input activation (say $a^{[l]}$) has the same dimension as the output activation (say $a^{[l+2]}$). To flesh out the different steps of what happens in a ResNet's identity block, here is an alternative diagram showing the individual steps:
<img src="images/idblock2_kiank.png" style="width:650px;height:150px;">
<caption><center> <u> <font color='purple'> **Figure 3** </u><font color='purple'> : **Identity block.** Skip connection "skips over" 2 layers. </center></caption>
The upper path is the "shortcut path." The lower path is the "main path." In this diagram, we have also made explicit the CONV2D and ReLU steps in each layer. To speed up training we have also added a BatchNorm step. Don't worry about this being complicated to implement--you'll see that BatchNorm is just one line of code in Keras!
In this exercise, you'll actually implement a slightly more powerful version of this identity block, in which the skip connection "skips over" 3 hidden layers rather than 2 layers. It looks like this:
<img src="images/idblock3_kiank.png" style="width:650px;height:150px;">
<caption><center> <u> <font color='purple'> **Figure 4** </u><font color='purple'> : **Identity block.** Skip connection "skips over" 3 layers.</center></caption>
Here are the individual steps.
First component of main path:
- The first CONV2D has $F_1$ filters of shape (1,1) and a stride of (1,1). Its padding is "valid" and its name should be `conv_name_base + '2a'`. Use 0 as the seed for the random initialization.
- The first BatchNorm is normalizing the 'channels' axis. Its name should be `bn_name_base + '2a'`.
- Then apply the ReLU activation function. This has no name and no hyperparameters.
Second component of main path:
- The second CONV2D has $F_2$ filters of shape $(f,f)$ and a stride of (1,1). Its padding is "same" and its name should be `conv_name_base + '2b'`. Use 0 as the seed for the random initialization.
- The second BatchNorm is normalizing the 'channels' axis. Its name should be `bn_name_base + '2b'`.
- Then apply the ReLU activation function. This has no name and no hyperparameters.
Third component of main path:
- The third CONV2D has $F_3$ filters of shape (1,1) and a stride of (1,1). Its padding is "valid" and its name should be `conv_name_base + '2c'`. Use 0 as the seed for the random initialization.
- The third BatchNorm is normalizing the 'channels' axis. Its name should be `bn_name_base + '2c'`.
- Note that there is **no** ReLU activation function in this component.
Final step:
- The `X_shortcut` and the output from the 3rd layer `X` are added together.
- **Hint**: The syntax will look something like `Add()([var1,var2])`
- Then apply the ReLU activation function. This has no name and no hyperparameters.
**Exercise**: Implement the ResNet identity block. We have implemented the first component of the main path. Please read this carefully to make sure you understand what it is doing. You should implement the rest.
- To implement the Conv2D step: [Conv2D](https://keras.io/layers/convolutional/#conv2d)
- To implement BatchNorm: [BatchNormalization](https://faroit.github.io/keras-docs/1.2.2/layers/normalization/) (axis: Integer, the axis that should be normalized (typically the 'channels' axis))
- For the activation, use: `Activation('relu')(X)`
- To add the value passed forward by the shortcut: [Add](https://keras.io/layers/merge/#add)
```
# GRADED FUNCTION: identity_block
def identity_block(X, f, filters, stage, block):
"""
Implementation of the identity block as defined in Figure 4
Arguments:
X -- input tensor of shape (m, n_H_prev, n_W_prev, n_C_prev)
f -- integer, specifying the shape of the middle CONV's window for the main path
filters -- python list of integers, defining the number of filters in the CONV layers of the main path
stage -- integer, used to name the layers, depending on their position in the network
block -- string/character, used to name the layers, depending on their position in the network
Returns:
X -- output of the identity block, tensor of shape (n_H, n_W, n_C)
"""
# defining name basis
conv_name_base = 'res' + str(stage) + block + '_branch'
bn_name_base = 'bn' + str(stage) + block + '_branch'
# Retrieve Filters
F1, F2, F3 = filters
# Save the input value. You'll need this later to add back to the main path.
X_shortcut = X
# First component of main path
X = Conv2D(filters = F1, kernel_size = (1, 1), strides = (1,1), padding = 'valid', name = conv_name_base + '2a', kernel_initializer = glorot_uniform(seed=0))(X)
X = BatchNormalization(axis = 3, name = bn_name_base + '2a')(X)
X = Activation('relu')(X)
### START CODE HERE ###
X = Conv2D(filters = F2, kernel_size = (f, f), strides = (1,1), padding = 'same', name = conv_name_base + '2b', kernel_initializer = glorot_uniform(seed=0))(X)
X = BatchNormalization(axis = 3, name = bn_name_base + '2b')(X)
X = Activation('relu')(X)
# Third component of main path (≈2 lines)
X = Conv2D(filters = F3, kernel_size = (1, 1), strides = (1,1), padding = 'valid', name = conv_name_base + '2c', kernel_initializer = glorot_uniform(seed=0))(X)
X = BatchNormalization(axis = 3, name = bn_name_base + '2c')(X)
# Final step: Add shortcut value to main path, and pass it through a RELU activation (≈2 lines)
X = Add()([X, X_shortcut])
X = Activation('relu')(X)
### END CODE HERE ###
return X
tf.reset_default_graph()
with tf.Session() as test:
np.random.seed(1)
A_prev = tf.placeholder("float", [3, 4, 4, 6])
X = np.random.randn(3, 4, 4, 6)
A = identity_block(A_prev, f = 2, filters = [2, 4, 6], stage = 1, block = 'a')
test.run(tf.global_variables_initializer())
out = test.run([A], feed_dict={A_prev: X, K.learning_phase(): 0})
print("out = " + str(out[0][1][1][0]))
```
**Expected Output**:
<table>
<tr>
<td>
**out**
</td>
<td>
[ 0.94822985 0. 1.16101444 2.747859 0. 1.36677003]
</td>
</tr>
</table>
## 2.2 - The convolutional block
The ResNet "convolutional block" is the second block type. You can use this type of block when the input and output dimensions don't match up. The difference with the identity block is that there is a CONV2D layer in the shortcut path:
<img src="images/convblock_kiank.png" style="width:650px;height:150px;">
<caption><center> <u> <font color='purple'> **Figure 4** </u><font color='purple'> : **Convolutional block** </center></caption>
* The CONV2D layer in the shortcut path is used to resize the input $x$ to a different dimension, so that the dimensions match up in the final addition needed to add the shortcut value back to the main path. (This plays a similar role as the matrix $W_s$ discussed in lecture.)
* For example, to reduce the activation dimensions's height and width by a factor of 2, you can use a 1x1 convolution with a stride of 2.
* The CONV2D layer on the shortcut path does not use any non-linear activation function. Its main role is to just apply a (learned) linear function that reduces the dimension of the input, so that the dimensions match up for the later addition step.
The details of the convolutional block are as follows.
First component of main path:
- The first CONV2D has $F_1$ filters of shape (1,1) and a stride of (s,s). Its padding is "valid" and its name should be `conv_name_base + '2a'`. Use 0 as the `glorot_uniform` seed.
- The first BatchNorm is normalizing the 'channels' axis. Its name should be `bn_name_base + '2a'`.
- Then apply the ReLU activation function. This has no name and no hyperparameters.
Second component of main path:
- The second CONV2D has $F_2$ filters of shape (f,f) and a stride of (1,1). Its padding is "same" and it's name should be `conv_name_base + '2b'`. Use 0 as the `glorot_uniform` seed.
- The second BatchNorm is normalizing the 'channels' axis. Its name should be `bn_name_base + '2b'`.
- Then apply the ReLU activation function. This has no name and no hyperparameters.
Third component of main path:
- The third CONV2D has $F_3$ filters of shape (1,1) and a stride of (1,1). Its padding is "valid" and it's name should be `conv_name_base + '2c'`. Use 0 as the `glorot_uniform` seed.
- The third BatchNorm is normalizing the 'channels' axis. Its name should be `bn_name_base + '2c'`. Note that there is no ReLU activation function in this component.
Shortcut path:
- The CONV2D has $F_3$ filters of shape (1,1) and a stride of (s,s). Its padding is "valid" and its name should be `conv_name_base + '1'`. Use 0 as the `glorot_uniform` seed.
- The BatchNorm is normalizing the 'channels' axis. Its name should be `bn_name_base + '1'`.
Final step:
- The shortcut and the main path values are added together.
- Then apply the ReLU activation function. This has no name and no hyperparameters.
**Exercise**: Implement the convolutional block. We have implemented the first component of the main path; you should implement the rest. As before, always use 0 as the seed for the random initialization, to ensure consistency with our grader.
- [Conv2D](https://keras.io/layers/convolutional/#conv2d)
- [BatchNormalization](https://keras.io/layers/normalization/#batchnormalization) (axis: Integer, the axis that should be normalized (typically the features axis))
- For the activation, use: `Activation('relu')(X)`
- [Add](https://keras.io/layers/merge/#add)
```
# GRADED FUNCTION: convolutional_block
def convolutional_block(X, f, filters, stage, block, s = 2):
"""
Implementation of the convolutional block as defined in Figure 4
Arguments:
X -- input tensor of shape (m, n_H_prev, n_W_prev, n_C_prev)
f -- integer, specifying the shape of the middle CONV's window for the main path
filters -- python list of integers, defining the number of filters in the CONV layers of the main path
stage -- integer, used to name the layers, depending on their position in the network
block -- string/character, used to name the layers, depending on their position in the network
s -- Integer, specifying the stride to be used
Returns:
X -- output of the convolutional block, tensor of shape (n_H, n_W, n_C)
"""
# defining name basis
conv_name_base = 'res' + str(stage) + block + '_branch'
bn_name_base = 'bn' + str(stage) + block + '_branch'
# Retrieve Filters
F1, F2, F3 = filters
# Save the input value
X_shortcut = X
##### MAIN PATH #####
# First component of main path
X = Conv2D(F1, (1, 1), strides = (s,s), name = conv_name_base + '2a', kernel_initializer = glorot_uniform(seed=0))(X)
X = BatchNormalization(axis = 3, name = bn_name_base + '2a')(X)
X = Activation('relu')(X)
### START CODE HERE ###
# Second component of main path (≈3 lines)
# Second component of main path (≈3 lines)
X = Conv2D(F2, (f, f), strides = (1,1), padding='same', name = conv_name_base + '2b', kernel_initializer = glorot_uniform(seed=0))(X)
X = BatchNormalization(axis = 3, name = bn_name_base + '2b')(X)
X = Activation('relu')(X)
# Third component of main path (≈2 lines)
X = Conv2D(F3, (1, 1), strides = (1,1), padding='valid', name = conv_name_base + '2c', kernel_initializer = glorot_uniform(seed=0))(X)
X = BatchNormalization(axis = 3, name = bn_name_base + '2c')(X)
##### SHORTCUT PATH #### (≈2 lines)
X_shortcut = Conv2D(F3, (1, 1), strides = (s,s), padding='valid', name = conv_name_base + '1', kernel_initializer = glorot_uniform(seed=0))(X_shortcut)
X_shortcut = BatchNormalization(axis = 3, name = bn_name_base + '1')(X_shortcut)
# Final step: Add shortcut value to main path, and pass it through a RELU activation (≈2 lines)
X = Add()([X, X_shortcut])
X = Activation('relu')(X)
### END CODE HERE ###
return X
tf.reset_default_graph()
with tf.Session() as test:
np.random.seed(1)
A_prev = tf.placeholder("float", [3, 4, 4, 6])
X = np.random.randn(3, 4, 4, 6)
A = convolutional_block(A_prev, f = 2, filters = [2, 4, 6], stage = 1, block = 'a')
test.run(tf.global_variables_initializer())
out = test.run([A], feed_dict={A_prev: X, K.learning_phase(): 0})
print("out = " + str(out[0][1][1][0]))
```
**Expected Output**:
<table>
<tr>
<td>
**out**
</td>
<td>
[ 0.09018463 1.23489773 0.46822017 0.0367176 0. 0.65516603]
</td>
</tr>
</table>
## 3 - Building your first ResNet model (50 layers)
You now have the necessary blocks to build a very deep ResNet. The following figure describes in detail the architecture of this neural network. "ID BLOCK" in the diagram stands for "Identity block," and "ID BLOCK x3" means you should stack 3 identity blocks together.
<img src="images/resnet_kiank.png" style="width:850px;height:150px;">
<caption><center> <u> <font color='purple'> **Figure 5** </u><font color='purple'> : **ResNet-50 model** </center></caption>
The details of this ResNet-50 model are:
- Zero-padding pads the input with a pad of (3,3)
- Stage 1:
- The 2D Convolution has 64 filters of shape (7,7) and uses a stride of (2,2). Its name is "conv1".
- BatchNorm is applied to the 'channels' axis of the input.
- MaxPooling uses a (3,3) window and a (2,2) stride.
- Stage 2:
- The convolutional block uses three sets of filters of size [64,64,256], "f" is 3, "s" is 1 and the block is "a".
- The 2 identity blocks use three sets of filters of size [64,64,256], "f" is 3 and the blocks are "b" and "c".
- Stage 3:
- The convolutional block uses three sets of filters of size [128,128,512], "f" is 3, "s" is 2 and the block is "a".
- The 3 identity blocks use three sets of filters of size [128,128,512], "f" is 3 and the blocks are "b", "c" and "d".
- Stage 4:
- The convolutional block uses three sets of filters of size [256, 256, 1024], "f" is 3, "s" is 2 and the block is "a".
- The 5 identity blocks use three sets of filters of size [256, 256, 1024], "f" is 3 and the blocks are "b", "c", "d", "e" and "f".
- Stage 5:
- The convolutional block uses three sets of filters of size [512, 512, 2048], "f" is 3, "s" is 2 and the block is "a".
- The 2 identity blocks use three sets of filters of size [512, 512, 2048], "f" is 3 and the blocks are "b" and "c".
- The 2D Average Pooling uses a window of shape (2,2) and its name is "avg_pool".
- The 'flatten' layer doesn't have any hyperparameters or name.
- The Fully Connected (Dense) layer reduces its input to the number of classes using a softmax activation. Its name should be `'fc' + str(classes)`.
**Exercise**: Implement the ResNet with 50 layers described in the figure above. We have implemented Stages 1 and 2. Please implement the rest. (The syntax for implementing Stages 3-5 should be quite similar to that of Stage 2.) Make sure you follow the naming convention in the text above.
You'll need to use this function:
- Average pooling [see reference](https://keras.io/layers/pooling/#averagepooling2d)
Here are some other functions we used in the code below:
- Conv2D: [See reference](https://keras.io/layers/convolutional/#conv2d)
- BatchNorm: [See reference](https://keras.io/layers/normalization/#batchnormalization) (axis: Integer, the axis that should be normalized (typically the features axis))
- Zero padding: [See reference](https://keras.io/layers/convolutional/#zeropadding2d)
- Max pooling: [See reference](https://keras.io/layers/pooling/#maxpooling2d)
- Fully connected layer: [See reference](https://keras.io/layers/core/#dense)
- Addition: [See reference](https://keras.io/layers/merge/#add)
```
# GRADED FUNCTION: ResNet50
def ResNet50(input_shape = (64, 64, 3), classes = 6):
"""
Implementation of the popular ResNet50 the following architecture:
CONV2D -> BATCHNORM -> RELU -> MAXPOOL -> CONVBLOCK -> IDBLOCK*2 -> CONVBLOCK -> IDBLOCK*3
-> CONVBLOCK -> IDBLOCK*5 -> CONVBLOCK -> IDBLOCK*2 -> AVGPOOL -> TOPLAYER
Arguments:
input_shape -- shape of the images of the dataset
classes -- integer, number of classes
Returns:
model -- a Model() instance in Keras
"""
# Define the input as a tensor with shape input_shape
X_input = Input(input_shape)
# Zero-Padding
X = ZeroPadding2D((3, 3))(X_input)
# Stage 1
X = Conv2D(64, (7, 7), strides = (2, 2), name = 'conv1', kernel_initializer = glorot_uniform(seed=0))(X)
X = BatchNormalization(axis = 3, name = 'bn_conv1')(X)
X = Activation('relu')(X)
X = MaxPooling2D((3, 3), strides=(2, 2))(X)
# Stage 2
X = convolutional_block(X, f = 3, filters = [64, 64, 256], stage = 2, block='a', s = 1)
X = identity_block(X, 3, [64, 64, 256], stage=2, block='b')
X = identity_block(X, 3, [64, 64, 256], stage=2, block='c')
### START CODE HERE ###
# Stage 3 (≈4 lines)
X = convolutional_block(X, f = 3, filters = [128, 128, 512], stage = 3, block='a', s = 2)
X = identity_block(X, 3, [128, 128, 512], stage=3, block='b')
X = identity_block(X, 3, [128, 128, 512], stage=3, block='c')
X = identity_block(X, 3, [128, 128, 512], stage=3, block='d')
# Stage 4 (≈6 lines)
X = convolutional_block(X, f = 3, filters = [256, 256, 1024], stage = 4, block='a', s = 2)
X = identity_block(X, 3, [256, 256, 1024], stage=4, block='b')
X = identity_block(X, 3, [256, 256, 1024], stage=4, block='c')
X = identity_block(X, 3, [256, 256, 1024], stage=4, block='d')
X = identity_block(X, 3, [256, 256, 1024], stage=4, block='e')
X = identity_block(X, 3, [256, 256, 1024], stage=4, block='f')
# Stage 5 (≈3 lines)
X = convolutional_block(X, f = 3, filters = [512, 512, 2048], stage = 5, block='a', s = 2)
X = identity_block(X, 3, [512, 512, 2048], stage=5, block='b')
X = identity_block(X, 3, [512, 512, 2048], stage=5, block='c')
# AVGPOOL (≈1 line). Use "X = AveragePooling2D(...)(X)"
X = AveragePooling2D()(X)
### END CODE HERE ###
# output layer
X = Flatten()(X)
X = Dense(classes, activation='softmax', name='fc' + str(classes), kernel_initializer = glorot_uniform(seed=0))(X)
# Create model
model = Model(inputs = X_input, outputs = X, name='ResNet50')
return model
```
Run the following code to build the model's graph. If your implementation is not correct you will know it by checking your accuracy when running `model.fit(...)` below.
```
model = ResNet50(input_shape = (64, 64, 3), classes = 6)
```
As seen in the Keras Tutorial Notebook, prior training a model, you need to configure the learning process by compiling the model.
```
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
```
The model is now ready to be trained. The only thing you need is a dataset.
Let's load the SIGNS Dataset.
<img src="images/signs_data_kiank.png" style="width:450px;height:250px;">
<caption><center> <u> <font color='purple'> **Figure 6** </u><font color='purple'> : **SIGNS dataset** </center></caption>
```
X_train_orig, Y_train_orig, X_test_orig, Y_test_orig, classes = load_dataset()
# Normalize image vectors
X_train = X_train_orig/255.
X_test = X_test_orig/255.
# Convert training and test labels to one hot matrices
Y_train = convert_to_one_hot(Y_train_orig, 6).T
Y_test = convert_to_one_hot(Y_test_orig, 6).T
print ("number of training examples = " + str(X_train.shape[0]))
print ("number of test examples = " + str(X_test.shape[0]))
print ("X_train shape: " + str(X_train.shape))
print ("Y_train shape: " + str(Y_train.shape))
print ("X_test shape: " + str(X_test.shape))
print ("Y_test shape: " + str(Y_test.shape))
```
Run the following cell to train your model on 2 epochs with a batch size of 32. On a CPU it should take you around 5min per epoch.
```
model.fit(X_train, Y_train, epochs = 2, batch_size = 32)
```
**Expected Output**:
<table>
<tr>
<td>
** Epoch 1/2**
</td>
<td>
loss: between 1 and 5, acc: between 0.2 and 0.5, although your results can be different from ours.
</td>
</tr>
<tr>
<td>
** Epoch 2/2**
</td>
<td>
loss: between 1 and 5, acc: between 0.2 and 0.5, you should see your loss decreasing and the accuracy increasing.
</td>
</tr>
</table>
Let's see how this model (trained on only two epochs) performs on the test set.
```
preds = model.evaluate(X_test, Y_test)
print ("Loss = " + str(preds[0]))
print ("Test Accuracy = " + str(preds[1]))
```
**Expected Output**:
<table>
<tr>
<td>
**Test Accuracy**
</td>
<td>
between 0.16 and 0.25
</td>
</tr>
</table>
For the purpose of this assignment, we've asked you to train the model for just two epochs. You can see that it achieves poor performances. Please go ahead and submit your assignment; to check correctness, the online grader will run your code only for a small number of epochs as well.
After you have finished this official (graded) part of this assignment, you can also optionally train the ResNet for more iterations, if you want. We get a lot better performance when we train for ~20 epochs, but this will take more than an hour when training on a CPU.
Using a GPU, we've trained our own ResNet50 model's weights on the SIGNS dataset. You can load and run our trained model on the test set in the cells below. It may take ≈1min to load the model.
```
model = load_model('ResNet50.h5')
preds = model.evaluate(X_test, Y_test)
print ("Loss = " + str(preds[0]))
print ("Test Accuracy = " + str(preds[1]))
```
ResNet50 is a powerful model for image classification when it is trained for an adequate number of iterations. We hope you can use what you've learnt and apply it to your own classification problem to perform state-of-the-art accuracy.
Congratulations on finishing this assignment! You've now implemented a state-of-the-art image classification system!
## 4 - Test on your own image (Optional/Ungraded)
If you wish, you can also take a picture of your own hand and see the output of the model. To do this:
1. Click on "File" in the upper bar of this notebook, then click "Open" to go on your Coursera Hub.
2. Add your image to this Jupyter Notebook's directory, in the "images" folder
3. Write your image's name in the following code
4. Run the code and check if the algorithm is right!
```
img_path = 'images/my_image.jpg'
img = image.load_img(img_path, target_size=(64, 64))
x = image.img_to_array(img)
x = np.expand_dims(x, axis=0)
x = x/255.0
print('Input image shape:', x.shape)
my_image = scipy.misc.imread(img_path)
imshow(my_image)
print("class prediction vector [p(0), p(1), p(2), p(3), p(4), p(5)] = ")
print(model.predict(x))
```
You can also print a summary of your model by running the following code.
```
model.summary()
```
Finally, run the code below to visualize your ResNet50. You can also download a .png picture of your model by going to "File -> Open...-> model.png".
```
plot_model(model, to_file='model.png')
SVG(model_to_dot(model).create(prog='dot', format='svg'))
```
## What you should remember
- Very deep "plain" networks don't work in practice because they are hard to train due to vanishing gradients.
- The skip-connections help to address the Vanishing Gradient problem. They also make it easy for a ResNet block to learn an identity function.
- There are two main types of blocks: The identity block and the convolutional block.
- Very deep Residual Networks are built by stacking these blocks together.
### References
This notebook presents the ResNet algorithm due to He et al. (2015). The implementation here also took significant inspiration and follows the structure given in the GitHub repository of Francois Chollet:
- Kaiming He, Xiangyu Zhang, Shaoqing Ren, Jian Sun - [Deep Residual Learning for Image Recognition (2015)](https://arxiv.org/abs/1512.03385)
- Francois Chollet's GitHub repository: https://github.com/fchollet/deep-learning-models/blob/master/resnet50.py
| github_jupyter |
<a href="https://colab.research.google.com/github/williamsdoug/CTG_RP/blob/master/CTG_RP_Display_Denoised.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# CTG Display Denoised
Denoising using algorithm detailed in : [A Comprehensive Feature Analysis of the Fetal Heart Rate Signal for the Intelligent Assessment of Fetal State](https://www.mdpi.com/2077-0383/7/8/223)
### Excerpt:
In clinical practice, during the recording process using Doppler ultrasound, the FHR signal
contains many artifacts or spikes due to maternal and fetal movements or transducer displacement [1]. Therefore, before further analysis, we eliminated noise to obtain a relatively pure signal for more accurate results, as described in Reference [18].
In this work, we employed a preprocessing involving three steps. Assume x(i) is an FHR signal with unit of beats per min (bpm) and a frequency of 4 Hz, where i = 1,2, ..., N and N is the number of samples.
- A stable segment is chosen as the starting point; in such a segment, five adjacent samples do not differ by more than 10 bpm, and missing data are excluded when the length of x(i) = 0 is equal or more than 10 s.
- Values of x(i) ≤50 or x(i) ≥ 200 are considered data spikes and are removed using linear interpolation
- We extrapolate x(i) using spline interolation again when the difference between x(i) and x(i-1) exceed 25 bpm, a value used to define unstable segments
Twenty minutes (N = 4800 samples) of signal length was the target used for further continuous
processing in this paper. Taking the signal labeled No. 1001 as a typical example, the result of this artifact removal scheme is presented in Figure 3.
```
from config_local import *
#from config_common import * # used only for local laptop configuration
# get_github_files(['ctg_utils.py']) # for development purposes only
import wfdb
import os
from pprint import pprint
import numpy as np
import matplotlib.pyplot as plt
from ctg_utils import get_all_recno, parse_meta_comments
import basic_denoise
from basic_denoise import get_valid_segments
RECORDINGS_DIR
#!ls /content/ctu-uhb-ctgdb
#!ls /Users/Test/Documents/GitHub/CTG_Analysis/local_data/sample_physionet_ctb_uhb_recordings
```
# Code
## Process Recordings
```
nRecordings = 3
for recno in sorted(get_all_recno(RECORDINGS_DIR))[:nRecordings]:
recno_full = os.path.join(RECORDINGS_DIR, recno)
print('\nRecord: {}'.format(recno))
all_sig, meta = wfdb.io.rdsamp(recno_full)
print('nSamples: {}'.format(all_sig.shape[0]))
#pprint(meta['comments'])
orig_hr = all_sig[:, 0]
sig_hr = np.copy(orig_hr)
sig_uc = all_sig[:, 1]
ts = np.arange(len(sig_hr))/4.0
selected_segments = get_valid_segments(orig_hr, ts, recno, verbose=True,
#max_change=15, verbose_details=True
)
continue
```
| github_jupyter |
## Linear Regression With Python
```
import warnings
warnings.filterwarnings("ignore")
```
## Housing Dataset
Your neighbor is a real estate agent and wants some help predicting housing prices for regions in the USA. It would be great if you could somehow create a model for her that allows her to put in a few features of a house and returns back an estimate of what the house would sell for.
She has asked you if you could help her out with your new data science skills. You say yes, and decide that Linear Regression might be a good path to solve this problem!
Your neighbor then gives you some information about a bunch of houses in regions of the United States,it is all in the data set: USA_Housing.csv.
The data contains the following columns:
* 'Avg. Area Income': Avg. Income of residents of the city house is located in.
* 'Avg. Area House Age': Avg Age of Houses in same city
* 'Avg. Area Number of Rooms': Avg Number of Rooms for Houses in same city
* 'Avg. Area Number of Bedrooms': Avg Number of Bedrooms for Houses in same city
* 'Area Population': Population of city house is located in
* 'Price': Price that the house sold at
* 'Address': Address for the house
**Let's get started!**
## Check out the data
We've been able to get some data from your neighbor for housing prices as a csv set, let's get our environment ready with the libraries we'll need and then import the data!
### Import Libraries
```
import pandas as pd
import numpy as np
import matplotlib as plt
import seaborn as sns
# To see the visualisations within the notebook
%matplotlib inline
sns.set_style('darkgrid')
```
### Check out the Data
```
USA_Housing = pd.read_csv("USA_Housing.csv")
USA_Housing.head()
# Dataframe has some columns, every row represents different house with address in the end, there is price for each house
# and other columns represent different statistics for city or the area in which house is located such as Avg. Income of people
# in that area, Avg. Age of house in that area, Avg. # of rooms and bedrooms and population of that area.
USA_Housing.mean()
USA_Housing.info()
# Gives us info about total number of columns and entries, as well as info about type of objects in dataframe.
USA_Housing.describe()
# To reference column names
USA_Housing.columns # A list of column names
```
# Creating Some plots to check data
```
# A plot that we can do if data isn't extremely large is sns's pairplot, passing in the entire dataframe.
sns.pairplot(USA_Housing)
```
* From pairplot above we can see that everything is almost normally distributed, except for average number of bedrooms as they can be only discrete 2,3,4,5 or 6 there is some noise but still we can differentiate that there are 4 to 5 discrete entries in bedroom features which lines up with how we know bedrooms are discrete.
```
# Let's check distribution of one of the column, Price column
sns.distplot(USA_Housing['Price'])
# We are doing it for price as it is going to be our target column or what we will be predicting.
# Here we predict price of the house. From plot we can infer that average price falls somewhere
# between 1M to 1.5 Million mark and is normally distributed.
USA_Housing.corr()
# Plotting a heatmap of correlation between each of the columns
sns.heatmap(USA_Housing.corr(),annot=True)
```
## Training a Linear Regression Model
Let's now begin to train out regression model! We will need to first split up our data into an X array that contains the features to train on, and a y array with the target variable, in this case the Price column. We will toss out the Address column because it only has text info that the linear regression model can't use.
### X and y arrays
```
USA_Housing.head(1)
X = USA_Housing[['Avg. Area Income', 'Avg. Area House Age', 'Avg. Area Number of Rooms',
'Avg. Area Number of Bedrooms', 'Area Population']]
y = USA_Housing['Price']
```
## Train Test Split
Now let's split the data into a training set and a testing set. We will train out model on the training set and then use the test set to evaluate the model.
```
from sklearn.model_selection import train_test_split
# We pass in our x and y data and then specifying the test size
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.4, random_state=101)
```
## Creating and Training the Model
```
from sklearn.linear_model import LinearRegression
lm = LinearRegression()
lm.fit(X_train,y_train)
```
## Model Evaluation
Let's evaluate the model by checking out it's coefficients and how we can interpret them.
```
print(lm.intercept_)
lm.coef_ # Returns coefficient for each feature
# Let's create a dataframe based off of these coefficients, as each of these coefficients relates to columns
X_train.columns
coeff_df = pd.DataFrame(lm.coef_,X_train.columns,columns=['Coeff'])
coeff_df
```
Interpreting the coefficients:
- Holding all other features fixed, a 1 unit increase in **Avg. Area Income** is associated with an **increase of \$21.52 **.
- Holding all other features fixed, a 1 unit increase in **Avg. Area House Age** is associated with an **increase of \$164883.28 **.
- Holding all other features fixed, a 1 unit increase in **Avg. Area Number of Rooms** is associated with an **increase of \$122368.67 **.
- Holding all other features fixed, a 1 unit increase in **Avg. Area Number of Bedrooms** is associated with an **increase of \$2233.80 **.
- Holding all other features fixed, a 1 unit increase in **Area Population** is associated with an **increase of \$15.15 **.
Does this make sense? Probably not because Jose Portilla made up this data. If you want real data to repeat this sort of analysis, check out the [boston dataset](http://scikit-learn.org/stable/modules/generated/sklearn.datasets.load_boston.html):
- If the above coefficients were of real data then we would have seen negative coefficients in areas such as age of the house increases then its price goes down and much more.
```
from sklearn.datasets import load_boston
boston = load_boston()
boston.keys()
print(boston['DESCR'])
```
For Linear Regression on the above dataset check : https://towardsdatascience.com/linear-regression-on-boston-housing-dataset-f409b7e4a155
## Predictions from our Model
Let's grab predictions off our test set and see how well it did!
```
predictions = lm.predict(X_test)
# Asking our model to predict for features from test dataset which it hasn't seen before specifically during training period.
# Just printing the predictions won't make that much of difference so here we compare predictions against real y_test
plt.pyplot.scatter(y_test,predictions)
# This basically means that line of best fit is pretty close to y values and error would be less.
```
## Regression Evaluation Metrics
Here are three common evaluation metrics for regression problems:
**Mean Absolute Error** (MAE) is the mean of the absolute value of the errors:
$$\frac 1n\sum_{i=1}^n|y_i-\hat{y}_i|$$
**Mean Squared Error** (MSE) is the mean of the squared errors:
$$\frac 1n\sum_{i=1}^n(y_i-\hat{y}_i)^2$$
**Root Mean Squared Error** (RMSE) is the square root of the mean of the squared errors:
$$\sqrt{\frac 1n\sum_{i=1}^n(y_i-\hat{y}_i)^2}$$
Comparing these metrics:
- **MAE** is the easiest to understand, because it's the average error.
- **MSE** is more popular than MAE, because MSE "punishes" larger errors, which tends to be useful in the real world, as it squares the larger errors.
- **RMSE** is even more popular than MSE, because RMSE is interpretable in the "y" units.
All of these are **loss functions**, because we want to minimize them.
```
from sklearn import metrics
print('MAE:', metrics.mean_absolute_error(y_test, predictions))
print('MSE:', metrics.mean_squared_error(y_test, predictions))
print('RMSE:', np.sqrt(metrics.mean_squared_error(y_test, predictions)))
print('R2 Score:', np.sqrt(metrics.r2_score(y_test, predictions)))
```
| github_jupyter |
# 03 - Full-Waveform Inversion (FWI)
This notebook is the third in a series of tutorial highlighting various aspects of seismic inversion based on Devito operators. In this second example we aim to highlight the core ideas behind seismic inversion, where we create an image of the subsurface from field recorded data. This tutorial follows on the modelling tutorial and will reuse the modelling and velocity model.
## Inversion requirement
Seismic inversion relies on two known parameters:
- **Field data** - or also called **recorded data**. This is a shot record corresponding to the true velocity model. In practice this data is acquired as described in the first tutorial. In order to simplify this tutorial we will fake field data by modelling it with the true velocity model.
- **Initial velocity model**. This is a velocity model that has been obtained by processing the field data. This model is a rough and very smooth estimate of the velocity as an initial estimate for the inversion. This is a necessary requirement for any optimization (method).
## Inversion computational setup
In this tutorial, we will introduce the gradient operator. This operator corresponds to the imaging condition introduced in the previous tutorial with some minor modifications that are defined by the objective function (also referred to in the tutorial series as the *functional*, *f*) and its gradient, *g*. We will define these two terms in the tutorial too.
## Notes on the operators
As we have already described the creation of a forward modelling operator, we will only call a wrapper function here. This wrapper already contains all the necessary operators for seismic modeling, imaging and inversion. Operators introduced for the first time in this tutorial will be properly described.
```
import numpy as np
%matplotlib inline
from devito import configuration
configuration['log-level'] = 'WARNING'
```
## Computational considerations
As we will see, FWI is computationally extremely demanding, even more than RTM. To keep this tutorial as lightwight as possible we therefore again use a very small demonstration model. We also define here a few parameters for the final example runs that can be changed to modify the overall runtime of the tutorial.
```
nshots = 9 # Number of shots to create gradient from
nreceivers = 101 # Number of receiver locations per shot
fwi_iterations = 5 # Number of outer FWI iterations
```
# True and smooth velocity models
We will use a very simple model domain, consisting of a circle within a 2D domain. We will again use the "true" model to generate our synthetic shot data and use a "smooth" model as our initial guess. In this case the smooth model is very smooth indeed - it is simply a constant background velocity without any features.
```
#NBVAL_IGNORE_OUTPUT
from examples.seismic import demo_model, plot_velocity, plot_perturbation
# Define true and initial model
shape = (101, 101) # Number of grid point (nx, nz)
spacing = (10., 10.) # Grid spacing in m. The domain size is now 1km by 1km
origin = (0., 0.) # Need origin to define relative source and receiver locations
model = demo_model('circle-isotropic', vp_circle=3.0, vp_background=2.5,
origin=origin, shape=shape, spacing=spacing, nbl=40)
model0 = demo_model('circle-isotropic', vp_circle=2.5, vp_background=2.5,
origin=origin, shape=shape, spacing=spacing, nbl=40,
grid = model.grid)
plot_velocity(model)
plot_velocity(model0)
plot_perturbation(model0, model)
assert model.grid == model0.grid
assert model.vp.grid == model0.vp.grid
```
## Acquisition geometry
In this tutorial, we will use the easiest case for inversion, namely a transmission experiment. The sources are located on one side of the model and the receivers on the other side. This allows to record most of the information necessary for inversion, as reflections usually lead to poor inversion results.
```
#NBVAL_IGNORE_OUTPUT
# Define acquisition geometry: source
from examples.seismic import AcquisitionGeometry
t0 = 0.
tn = 1000.
f0 = 0.010
# First, position source centrally in all dimensions, then set depth
src_coordinates = np.empty((1, 2))
src_coordinates[0, :] = np.array(model.domain_size) * .5
src_coordinates[0, 0] = 20. # Depth is 20m
# Define acquisition geometry: receivers
# Initialize receivers for synthetic and imaging data
rec_coordinates = np.empty((nreceivers, 2))
rec_coordinates[:, 1] = np.linspace(0, model.domain_size[0], num=nreceivers)
rec_coordinates[:, 0] = 980.
# Geometry
geometry = AcquisitionGeometry(model, rec_coordinates, src_coordinates, t0, tn, f0=f0, src_type='Ricker')
# We can plot the time signature to see the wavelet
geometry.src.show()
#NBVAL_IGNORE_OUTPUT
# Plot acquisition geometry
plot_velocity(model, source=geometry.src_positions,
receiver=geometry.rec_positions[::4, :])
```
## True and smooth data
We can generate shot records for the true and smoothed initial velocity models, since the difference between them will again form the basis of our imaging procedure.
```
# Compute synthetic data with forward operator
from examples.seismic.acoustic import AcousticWaveSolver
solver = AcousticWaveSolver(model, geometry, space_order=4)
true_d, _, _ = solver.forward(vp=model.vp)
# Compute initial data with forward operator
smooth_d, _, _ = solver.forward(vp=model0.vp)
#NBVAL_IGNORE_OUTPUT
from examples.seismic import plot_shotrecord
# Plot shot record for true and smooth velocity model and the difference
plot_shotrecord(true_d.data, model, t0, tn)
plot_shotrecord(smooth_d.data, model, t0, tn)
plot_shotrecord(smooth_d.data - true_d.data, model, t0, tn)
```
# Full-Waveform Inversion
## Formulation
Full-waveform inversion (FWI) aims to invert an accurate model of the discrete wave velocity, $\mathbf{c}$, or equivalently the square slowness of the wave, $\mathbf{m} = \frac{1}{\mathbf{c}^2}$, from a given set of measurements of the pressure wavefield $\mathbf{u}$. This can be expressed as the following optimization problem [1, 2]:
\begin{aligned}
\mathop{\hbox{minimize}}_{\mathbf{m}} \Phi_s(\mathbf{m})&=\frac{1}{2}\left\lVert\mathbf{P}_r
\mathbf{u} - \mathbf{d}\right\rVert_2^2 \\
\mathbf{u} &= \mathbf{A}(\mathbf{m})^{-1} \mathbf{P}_s^T \mathbf{q}_s,
\end{aligned}
where $\mathbf{P}_r$ is the sampling operator at the receiver locations, $\mathbf{P}_s^T$ is the injection operator at the source locations, $\mathbf{A}(\mathbf{m})$ is the operator representing the discretized wave equation matrix, $\mathbf{u}$ is the discrete synthetic pressure wavefield, $\mathbf{q}_s$ is the corresponding pressure source and $\mathbf{d}$ is the measured data. It is worth noting that $\mathbf{m}$ is the unknown in this formulation and that multiple implementations of the wave equation operator $\mathbf{A}(\mathbf{m})$ are possible.
We have already defined a concrete solver scheme for $\mathbf{A}(\mathbf{m})$ in the first tutorial, including appropriate implementations of the sampling operator $\mathbf{P}_r$ and source term $\mathbf{q}_s$.
To solve this optimization problem using a gradient-based method, we use the
adjoint-state method to evaluate the gradient $\nabla\Phi_s(\mathbf{m})$:
\begin{align}
\nabla\Phi_s(\mathbf{m})=\sum_{\mathbf{t} =1}^{n_t}\mathbf{u}[\mathbf{t}] \mathbf{v}_{tt}[\mathbf{t}] =\mathbf{J}^T\delta\mathbf{d}_s,
\end{align}
where $n_t$ is the number of computational time steps, $\delta\mathbf{d}_s = \left(\mathbf{P}_r \mathbf{u} - \mathbf{d} \right)$ is the data residual (difference between the measured data and the modelled data), $\mathbf{J}$ is the Jacobian operator and $\mathbf{v}_{tt}$ is the second-order time derivative of the adjoint wavefield solving:
\begin{align}
\mathbf{A}^T(\mathbf{m}) \mathbf{v} = \mathbf{P}_r^T \delta\mathbf{d}.
\end{align}
We see that the gradient of the FWI function is the previously defined imaging condition with an extra second-order time derivative. We will therefore reuse the operators defined previously inside a Devito wrapper.
## FWI gradient operator
To compute a single gradient $\nabla\Phi_s(\mathbf{m})$ in our optimization workflow we again use `solver.forward` to compute the entire forward wavefield $\mathbf{u}$ and a similar pre-defined gradient operator to compute the adjoint wavefield `v`. The gradient operator provided by our `solver` utility also computes the correlation between the wavefields, allowing us to encode a similar procedure to the previous imaging tutorial as our gradient calculation:
- Simulate the forward wavefield with the background velocity model to get the synthetic data and save the full wavefield $\mathbf{u}$
- Compute the data residual
- Back-propagate the data residual and compute on the fly the gradient contribution at each time step.
This procedure is applied to multiple source positions and summed to obtain a gradient image of the subsurface. We again prepare the source locations for each shot and visualize them, before defining a single gradient computation over a number of shots as a single function.
```
#NBVAL_IGNORE_OUTPUT
# Prepare the varying source locations sources
source_locations = np.empty((nshots, 2), dtype=np.float32)
source_locations[:, 0] = 30.
source_locations[:, 1] = np.linspace(0., 1000, num=nshots)
plot_velocity(model, source=source_locations)
from devito import Eq, Operator
# Computes the residual between observed and synthetic data into the residual
def compute_residual(residual, dobs, dsyn):
if residual.grid.distributor.is_parallel:
# If we run with MPI, we have to compute the residual via an operator
# First make sure we can take the difference and that receivers are at the
# same position
assert np.allclose(dobs.coordinates.data[:], dsyn.coordinates.data)
assert np.allclose(residual.coordinates.data[:], dsyn.coordinates.data)
# Create a difference operator
diff_eq = Eq(residual, dsyn.subs({dsyn.dimensions[-1]: residual.dimensions[-1]}) -
dobs.subs({dobs.dimensions[-1]: residual.dimensions[-1]}))
Operator(diff_eq)()
else:
# A simple data difference is enough in serial
residual.data[:] = dsyn.data[:] - dobs.data[:]
return residual
# Create FWI gradient kernel
from devito import Function, TimeFunction, norm
from examples.seismic import Receiver
import scipy
def fwi_gradient(vp_in):
# Create symbols to hold the gradient
grad = Function(name="grad", grid=model.grid)
# Create placeholders for the data residual and data
residual = Receiver(name='residual', grid=model.grid,
time_range=geometry.time_axis,
coordinates=geometry.rec_positions)
d_obs = Receiver(name='d_obs', grid=model.grid,
time_range=geometry.time_axis,
coordinates=geometry.rec_positions)
d_syn = Receiver(name='d_syn', grid=model.grid,
time_range=geometry.time_axis,
coordinates=geometry.rec_positions)
objective = 0.
for i in range(nshots):
# Update source location
geometry.src_positions[0, :] = source_locations[i, :]
# Generate synthetic data from true model
_, _, _ = solver.forward(vp=model.vp, rec=d_obs)
# Compute smooth data and full forward wavefield u0
_, u0, _ = solver.forward(vp=vp_in, save=True, rec=d_syn)
# Compute gradient from data residual and update objective function
compute_residual(residual, d_obs, d_syn)
objective += .5*norm(residual)**2
solver.gradient(rec=residual, u=u0, vp=vp_in, grad=grad)
return objective, grad
```
Having defined our FWI gradient procedure we can compute the initial iteration from our starting model. This allows us to visualize the gradient alongside the model perturbation and the effect of the gradient update on the model.
```
# Compute gradient of initial model
ff, update = fwi_gradient(model0.vp)
assert np.isclose(ff, 57283, rtol=1e0)
#NBVAL_IGNORE_OUTPUT
from devito import mmax
from examples.seismic import plot_image
# Plot the FWI gradient
plot_image(-update.data, vmin=-1e4, vmax=1e4, cmap="jet")
# Plot the difference between the true and initial model.
# This is not known in practice as only the initial model is provided.
plot_image(model0.vp.data - model.vp.data, vmin=-1e-1, vmax=1e-1, cmap="jet")
# Show what the update does to the model
alpha = .5 / mmax(update)
plot_image(model0.vp.data + alpha*update.data, vmin=2.5, vmax=3.0, cmap="jet")
```
We see that the gradient and the true perturbation have the same sign, therefore, with an appropriate scaling factor, we will update the model in the correct direction.
```
from devito import Min, Max
# Define bounding box constraints on the solution.
def update_with_box(vp, alpha, dm, vmin=2.0, vmax=3.5):
"""
Apply gradient update in-place to vp with box constraint
Notes:
------
For more advanced algorithm, one will need to gather the non-distributed
velocity array to apply constrains and such.
"""
update = vp + alpha * dm
update_eq = Eq(vp, Max(Min(update, vmax), vmin))
Operator(update_eq)()
#NBVAL_SKIP
from devito import mmax
# Run FWI with gradient descent
history = np.zeros((fwi_iterations, 1))
for i in range(0, fwi_iterations):
# Compute the functional value and gradient for the current
# model estimate
phi, direction = fwi_gradient(model0.vp)
# Store the history of the functional values
history[i] = phi
# Artificial Step length for gradient descent
# In practice this would be replaced by a Linesearch (Wolfe, ...)
# that would guarantee functional decrease Phi(m-alpha g) <= epsilon Phi(m)
# where epsilon is a minimum decrease constant
alpha = .05 / mmax(direction)
# Update the model estimate and enforce minimum/maximum values
update_with_box(model0.vp , alpha , direction)
# Log the progress made
print('Objective value is %f at iteration %d' % (phi, i+1))
#NBVAL_IGNORE_OUTPUT
# Plot inverted velocity model
plot_velocity(model0)
#NBVAL_SKIP
import matplotlib.pyplot as plt
# Plot objective function decrease
plt.figure()
plt.loglog(history)
plt.xlabel('Iteration number')
plt.ylabel('Misift value Phi')
plt.title('Convergence')
plt.show()
```
## References
[1] _Virieux, J. and Operto, S.: An overview of full-waveform inversion in exploration geophysics, GEOPHYSICS, 74, WCC1–WCC26, doi:10.1190/1.3238367, http://library.seg.org/doi/abs/10.1190/1.3238367, 2009._
[2] _Haber, E., Chung, M., and Herrmann, F. J.: An effective method for parameter estimation with PDE constraints with multiple right hand sides, SIAM Journal on Optimization, 22, http://dx.doi.org/10.1137/11081126X, 2012._
<sup>This notebook is part of the tutorial "Optimised Symbolic Finite Difference Computation with Devito" presented at the Intel® HPC Developer Conference 2017.</sup>
| github_jupyter |
# Metrics Analysis Notebook (k8s)
#### Used to analyse / visualize the metrics, data fetched from prometheus (monitoring cluster)
### Contributor: Aditya Srivastava <adityasrivastava301199@gmail.com>
```
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
import numpy as np
import datetime
import time
import requests
from pprint import pprint
import json
from datetime import datetime
PROMETHEUS = 'http://10.10.120.211:30902/' #do not change, unless sure
```
## Helper Functions
```
#function to make DF out of query json
def convert_to_df(res_json):
data_list = res_json['data']['result']
res_df = pd.DataFrame()
if not data_list:
return res_df
# making colums
headers = data_list[0]
for data in data_list:
metrics = data['metric']
for metric in metrics.keys():
res_df[metric] = np.nan
res_df['value'] = 0
# filling the df
for data in data_list:
metrics = data['metric']
metrics['value'] = data['value'][-1]
res_df = res_df.append(metrics, ignore_index=True)
return res_df
def convert_to_df_range(res_json):
data_list = res_json['data']['result']
res_df = pd.DataFrame()
if not data_list:
return res_df
# filling the df
for data in data_list:
metrics = data['metric']
values = np.array(data['values'])
for time, value in values:
metrics['timestamp'] = time
metrics['value'] = value
res_df = res_df.append(metrics, ignore_index=True)
return res_df
# functions to query
def convert_to_timestamp(s):
return time.mktime(datetime.strptime(s, "%Y-%m-%d %H:%M:%S").timetuple())
def query_current(params={}):
# input: params
# type: dict
# Example: {'query': 'container_cpu_user_seconds_total'}
# Output: dict, loaded json response of the query
res = requests.get(PROMETHEUS + '/api/v1/query',
params=params)
return json.loads(res.text)
def query_range(start, end, params={}, steps = '30s'):
# input: params
# type: dict
# Example: {'query': 'container_cpu_user_seconds_total'}
# Output: dict, loaded json response of the query
params["start"] = convert_to_timestamp(start)
params["end"] = convert_to_timestamp(end)
params["step"] = steps
print(params)
res = requests.get(PROMETHEUS + '/api/v1/query_range',
params=params,
)
return json.loads(res.text)
```
## Analysis Function
#### CPU
```
# CPU Unused Cores
def unused_cores(start=None, end=None, node=None, steps='15s', csv=None, verbose=False):
if csv is not None:
df = pd.read_csv(csv)
return df
else:
if start is None or end is None or node is None:
return "Start, end and Node name required when fetching from prometheus"
params = {'query' : "collectd_cpu_percent{exported_instance='" + node + "'}"}
target_cpu_usage_range = query_range(start, end, params, steps)
df = convert_to_df_range(target_cpu_usage_range)
df = df.drop(['__name__', 'instance', 'job'], axis = 1)
groups = df.groupby(['cpu'])
if verbose: print("Unused Cores :")
unused_cores = []
for key, item in groups:
curr_df = item
idle_row = curr_df.loc[curr_df['type'] == 'idle']
if idle_row['value'].iloc[0] == '100':
if verbose: print("Core: ",key)
unused_cores.append(int(key))
print("Number of unused cores: ", len(unused_cores))
return unused_cores
#CPU fully used cores
def fully_used_cores(start=None, end=None, node=None, steps='15s', csv=None, verbose=False):
if csv is not None:
df = pd.read_csv(csv)
return df
else:
if start is None or end is None or node is None:
return "Start, end and Node name required when fetching from prometheus"
params = {'query' : "collectd_cpu_percent{exported_instance='" + node + "'}"}
target_cpu_usage_range = query_range(start, end, params, steps)
df = convert_to_df_range(target_cpu_usage_range)
df = df.drop(['__name__', 'instance', 'job'], axis = 1)
groups = df.groupby(['cpu'])
if verbose: print("Fully Used Cores :")
fully_used_cores = []
for key, item in groups:
curr_df = item
idle_row = curr_df.loc[curr_df['type'] == 'idle']
if idle_row['value'].iloc[0] == '0':
if verbose: print("Core: ",key)
fully_used_cores.append(int(key))
print("Number of fully used cores: ", len(fully_used_cores))
return fully_used_cores
# CPU used cores plots
def plot_used_cores(start=None, end=None, node=None, steps='15s', csv=None, verbose=False):
if csv is not None:
df = pd.read_csv(csv)
#
df['rate'] = df['value'].diff()
fig = plt.figure(figsize=(24,6), facecolor='oldlace', edgecolor='red')
ax1 = fig.add_subplot(111)
ax1.title.set_text('CPU usage')
ax1.plot(df['epoch'], df['rate'])
return df
else:
if start is None or end is None or node is None:
return "Start, end and Node name required when fetching from prometheus"
params = {'query' : "collectd_cpu_percent{exported_instance='" + node + "'}"}
target_cpu_usage_range = query_range(start, end, params, steps)
df = convert_to_df_range(target_cpu_usage_range)
df = df.drop(['__name__', 'instance', 'job'], axis = 1)
groups = df.groupby(['cpu'])
used_cores = []
for key, item in groups:
curr_df = item
idle_row = curr_df.loc[curr_df['type'] == 'idle']
if idle_row['value'].iloc[0] != '100':
used_cores.append(key)
type_grps = curr_df.groupby('type')
fig = plt.figure(figsize=(24,6), facecolor='oldlace', edgecolor='red')
for type_key, new_item in type_grps:
if type_key == 'system':
ax1 = fig.add_subplot(131)
ax1.title.set_text(type_key)
ax1.plot(new_item['timestamp'], new_item['value'])
elif type_key == 'user':
ax2 = fig.add_subplot(132)
ax2.title.set_text(type_key)
ax2.plot(new_item['timestamp'], new_item['value'])
elif type_key == 'wait':
ax3 = fig.add_subplot(133)
ax3.title.set_text(type_key)
ax3.plot(new_item['timestamp'], new_item['value'])
plt.suptitle('Used CPU Core {}'.format(key), fontsize=14)
plt.show()
print("Number of used cores: ", len(used_cores))
return used_cores
```
#### Interface
```
# Interface Dropped (both type 1 and 2, i.e rx and tx)
#TODO: Change this to separate functions later
def interface_dropped(start=None, end=None, node=None, steps='15s', csv=None, verbose=False):
if csv is not None:
df = pd.read_csv(csv)
df_0 = df #TODO: Change this
df_1 = df #TODO: Change this
else:
if start is None or end is None or node is None:
return "Start, end and Node name required when fetching from prometheus"
params = {'query' : "collectd_interface_if_dropped_0_total{exported_instance='" + node + "'}"}
interface_dropped_0 = query_range(start, end, params, steps)
df_0 = convert_to_df_range(interface_dropped_0)
params = {'query' : "collectd_interface_if_dropped_1_total{exported_instance='" + node + "'}"}
interface_dropped_1 = query_range(start, end, params, steps)
df_1 = convert_to_df_range(interface_dropped_1)
#df_0 : interfaces_dropped_0_df
df_0 = df_0.drop(['__name__', 'instance', 'job'], axis = 1)
#df_1 : interfaces_dropped_1_df
df_1 = df_1.drop(['__name__', 'instance', 'job'], axis = 1)
groups_0 = df_0.groupby(['interface'])
groups_1 = df_1.groupby(['interface'])
groups = [groups_0, groups_1]
dropped_interfaces= []
drop_type = 0
color = ['oldlace', 'mistyrose']
plot_iter = 111
for group in groups:
dropped = []
for key, item in group:
curr_df = item
if np.any(curr_df['value'] == '1'):
dropped_row = curr_df.loc[curr_df['value'] == '1']
dropped.append([key, dropped_row['timestamp'].iloc[0]])
fig = plt.figure(figsize=(24,6), facecolor=color[drop_type], edgecolor='red')
ax = fig.add_subplot(plot_iter)
ax.title.set_text("Interface: {}".format(key))
ax.plot(item['timestamp'], item['value'])
dropped_interfaces.append(dropped)
plt.suptitle('Interfaces Drop type {}'.format(drop_type), fontsize=14)
plt.show()
drop_type += 1
return dropped_interfaces
# Interface Errors (both type 1 and 2, i.e rx and tx)
#TODO: Change this to separate functions later
def interface_errors(start=None, end=None, node=None, steps='15s', csv=None, verbose=False):
if csv is not None:
df = pd.read_csv(csv)
df_0 = df #TODO: Change this
df_1 = df #TODO: Change this
else:
if start is None or end is None or node is None:
return "Start, end and Node name required when fetching from prometheus"
params = {'query' : "collectd_interface_if_errors_0_total{exported_instance='" + node + "'}"}
interfaces_errors_0 = query_range(start, end, params, steps)
df_0 = convert_to_df_range(interfaces_errors_0)
params = {'query' : "collectd_interface_if_errors_1_total{exported_instance='" + node + "'}"}
interface_errors_1 = query_range(start, end, params, steps)
df_1 = convert_to_df_range(interface_errors_1)
#df_0 : interfaces_errors_0_df
df_0 = df_0.drop(['__name__', 'instance', 'job'], axis = 1)
#df_1 : interfaces_dropped_1_df
df_1 = df_1.drop(['__name__', 'instance', 'job'], axis = 1)
groups_0 = df_0.groupby(['interface'])
groups_1 = df_1.groupby(['interface'])
groups = [groups_0, groups_1]
err_interfaces= []
err_type = 0
color = ['oldlace', 'mistyrose']
for group in groups:
errors = []
for key, item in group:
curr_df = item
if np.any(curr_df['value'] == '1'):
err_row = curr_df.loc[curr_df['value'] == '1']
erros.append([key, err_row['timestamp'].iloc[0]])
fig = plt.figure(figsize=(24,6), facecolor=color[err_type], edgecolor='red')
ax = fig.add_subplot(111)
ax.title.set_text("Interface: {}".format(key))
ax.plot(item['timestamp'], item['value'])
err_interfaces.append(errors)
plt.suptitle('Interfaces Error type {}'.format(err_type), fontsize=14)
plt.show()
err_type += 1
return err_interfaces
```
#### RDT
```
# L3 cache bytes
def plot_rdt_bytes(start=None, end=None, node=None, steps='15s', csv=None, verbose=False):
if csv is not None:
df = pd.read_csv(csv)
else:
if start is None or end is None or node is None:
return "Start, end and Node name required when fetching from prometheus"
params = {'query' : "collectd_intel_rdt_bytes{exported_instance='" + node + "'}"}
intel_rdt_bytes = query_range(start, end, params, steps)
df = convert_to_df_range(intel_rdt_bytes)
df = df.drop(['__name__', 'instance', 'job'], axis = 1)
groups = df.groupby(['intel_rdt'])
for key, item in groups:
curr_df = item
fig = plt.figure(figsize=(24,6), facecolor='oldlace', edgecolor='red')
ax1 = fig.add_subplot(111)
ax1.title.set_text("Intel RDT Number: {}".format(key))
ax1.plot(item['timestamp'], item['value'])
plt.show()
return
# L3 IPC values
def plot_rdt_ipc(start=None, end=None, node=None, steps='15s', csv=None, verbose=False):
if csv is not None:
df = pd.read_csv(csv)
else:
if start is None or end is None or node is None:
return "Start, end and Node name required when fetching from prometheus"
params = {'query' : "collectd_intel_rdt_ipc{exported_instance='" + node + "'}"}
intel_rdt_ipc = query_range(start, end, params, steps)
df = convert_to_df_range(intel_rdt_ipc)
df = df.drop(['__name__', 'instance', 'job'], axis = 1)
groups = df.groupby(['intel_rdt'])
for key, item in groups:
curr_df = item
fig = plt.figure(figsize=(24,6), facecolor='oldlace', edgecolor='red')
ax1 = fig.add_subplot(111)
ax1.title.set_text("Intel RDT Number: {}, IPC value".format(key))
ax1.plot(item['timestamp'], item['value'])
plt.show()
return
# memeory bandwidtdh
def get_rdt_memory_bandwidth(start=None, end=None, node=None, steps='15s', csv=None, verbose=False):
if csv is not None:
df = pd.read_csv(csv)
else:
if start is None or end is None or node is None:
return "Start, end and Node name required when fetching from prometheus"
params = {'query' : "collectd_intel_rdt_memory_bandwidth_total{exported_instance='" + node + "'}"}
intel_rdt_mem_bw = query_range(start, end, params, steps)
df = convert_to_df_range(intel_rdt_mem_bw)
df = df.drop(['__name__', 'instance', 'job'], axis = 1)
return df
```
#### Memory
```
def get_memory_usage(start=None, end=None, node=None, steps='15s', csv=None, verbose=False):
if csv is not None:
df = pd.read_csv(csv)
else:
if start is None or end is None or node is None:
return "Start, end and Node name required when fetching from prometheus"
params = {'query' : "collectd_memory{exported_instance='" + node + "'} / (1024*1024*1024) "}
target_memory_usage_range = query_range(start, end, params, steps)
df = convert_to_df_range(target_memory_usage_range)
df = df.drop(['instance', 'job'], axis = 1)
return df
```
## Testing Zone
```
# prom fetch
cores = unused_cores('2020-07-31 08:00:12', '2020-07-31 08:01:12', 'pod12-node4')
print(cores)
```
## Usage / Examples
##### CPU
- For calling cpu unsued cores
```py
# Fetching from prometheus
cores = unused_cores('2020-07-31 08:00:12', '2020-07-31 08:01:12', 'pod12-node4')
```
- For finding fully used cores
```py
# Fetching from prometheus
fully_used = fully_used_cores('2020-07-31 08:00:12', '2020-07-31 08:01:12', 'pod12-node4')
```
- Similarly for plotting used cores
```py
# Fetching
plot_used_cores('2020-07-31 08:00:12', '2020-07-31 08:01:12', 'pod12-node4')
#csv
# use Analysis-Monitoring-Local Notebook for correct analysis
plot_used_cores(csv='metrics_data/cpu-0/cpu-user-2020-06-02')
```
##### Interface
- Interface Dropped
```py
# Fetching from prom
dropped_interfaces = interface_dropped('2020-07-31 08:00:12', '2020-07-31 08:01:12', 'pod12-node4')
```
- Interface Errors
```py
# Fetching from prom
interface_errors('2020-07-31 08:00:12', '2020-07-31 08:01:12', 'pod12-node4')
```
##### RDT
- Plot bytes
```py
# fetch
plot_rdt_bytes('2020-07-31 08:00:12', '2020-07-31 08:01:12','pod12-node4')
```
- Plot ipc values
```py
#fetch
plot_rdt_ipc('2020-07-31 08:00:12', '2020-07-31 08:01:12', 'pod12-node4')
```
- Memory bandwidth
```py
#fetch
get_rdt_memory_bandwidth('2020-07-31 08:00:12', '2020-07-31 08:01:12', 'pod12-node4')
```
| github_jupyter |
# 2D Lists Indexing
**Remember:**
- Lists can be used to group different values together - it's just a collection of things.
- You can make a list in Python by putting different things in a box of brackets `[]` separated by commas
- You can make lists of lists, which are called 2D lists
Here we're going to learn how to index lists of lists. Let's recreate our 2D list of `food`:
```
# create 2D list
food = [['apple', 'banana', 'grape', 'mango'],
['lettuce','carrot','cucumber','beet'],
['chicken', 'beef', 'fish', 'pork']]
# print 2D list along with its length
print(food)
print(len(food))
```
Like we saw in the last lesson, the length of the `food` list (3) is the number of 1D lists that make up the 2D list. The fruit list is the first element in `food`. To get this first element from `food`, we do it the same way we would in a 1D list:
```
# get the fruit list from the food list
food[0]
```
In this way, we can think of the list `food` as a 1D list, where each element is another list. As we saw in the last command, we can reference the first sublist from `food` using `food[0]`.
Knowing this, how can we get the length of the first list within `food`?
```
# print length of the first index of food
len(food[0])
```
Thinking back to indexing 1D lists, how do you think we can get the lists fruit and vegetables from the list `food`?
```
# get fruit and vegetables from the list
food[0:2]
```
What if we want to get the value `'mango'` from our list? We can first save the first element from `food` to a variable `fruit`, and then index the `fruit` list:
```
# get the fruit list from food and save it in the variable fruit
fruit = food[0]
# print the fruit list to make sure you got what you wanted
print(fruit)
# get mango from the fruit list
fruit[3]
```
We can also get the value `'mango'` from our list in one line! To do this, we can just index `food` twice. The first index is to get the fruit list, and the second index is to get `'mango'` from the fruit list:
```
# get mango by indexing the food list twice
food[0][3]
```
How can you get `'carrot'` from the `food` 2D list?
```
# get carrot from the food 2D list
food[1][1]
```
Now, try getting `'fish'` and `'pork'` from the `food` 2D list:
```
# get fish and pork from the food 2D list
food[2][2:4]
```
Finally, let's make a new list called `favorites` with your favorite food within each list in `food`. Print it out afterwards to make sure you did what you want!
```
# make a list of your favorite fruit, vegetable, and meat
favorites = [food[0][3], food[0][2], food[2][1]]
# print favorite
print(favorites)
```
Nice job! You just learned how to index 2D lists in Python!
| github_jupyter |
# Imports
```
import gspread
import pandas as pd
import numpy as np
import datetime
from us_state_abbrev import abbrev_us_state
import re
from tqdm.notebook import tqdm
gc = gspread.oauth()
```
# Import data (1 team entry, 3 comparison datasets)
```
team_entry_sheet = gc.open("Mountains-Midwest COVID-19 by County (TEAM ENTRY) 10-28-2020 to present").sheet1
nyt_comparison_sheet = gc.open("QA TEAM ONLY - COVID-19 by County Comparison Data (NYT)").sheet1
team_entry = pd.DataFrame(team_entry_sheet.get_all_records(head = 2))
team_entry = team_entry.replace(r'^\s*$', np.nan, regex=True)
nyt_comparison = pd.DataFrame(nyt_comparison_sheet.get_all_records())
nyt_comparison = nyt_comparison.replace(r'^\s*$', np.nan, regex=True)
JHU_USAF_Cases_comparison_sheets = gc.open("COVID-19 by County Comparison Data (JHU/USAF) ")
USAF_Cases_comparison_sheet = JHU_USAF_Cases_comparison_sheets.sheet1
USAF_Deaths_comparison_sheet = JHU_USAF_Cases_comparison_sheets.get_worksheet(1)
JHU_comparison_sheet = JHU_USAF_Cases_comparison_sheets.get_worksheet(2)
USAF_Cases_comparison = pd.DataFrame(USAF_Cases_comparison_sheet.get_all_records())
USAF_Deaths_comparison = pd.DataFrame(USAF_Deaths_comparison_sheet.get_all_records())
JHU_comparison = pd.DataFrame(JHU_comparison_sheet.get_all_records())
USAF_Cases_comparison = USAF_Cases_comparison.replace(r'^\s*$', np.nan, regex=True)
USAF_Deaths_comparison = USAF_Deaths_comparison.replace(r'^\s*$', np.nan, regex=True)
JHU_comparison = JHU_comparison.replace(r'^\s*$', np.nan, regex=True)
# with pd.option_context("display.max_rows", 1000):
# display(team_entry[team_entry.state == 'TX'].loc[:, 'tstpos_101620':'pbmort_101620'])
# display(team_entry.loc[team_entry.state == 'TX', 'tstpos_101620':'pbmort_101620'])
# with pd.option_context('display.max_rows', None, 'display.max_columns', None): # more options can be specified also
# print(team_entry.pbmort_092520)
# team_entry.pbmort_092520.values[:] = 0
```
# Functions needed
```
# given a data frame and state, find out the latest updated time, formated in US time format
def lastTimeUpdated(state):
print(f'For {state}, last updated on:')
# team entry
yourstateDF = team_entry[team_entry.state == state]
nototalDF = yourstateDF.filter(regex=f"\d$", axis=1).iloc[1:]
first_col_empty = pd.isna(nototalDF).any().idxmax()
# print(first_col_empty) # show which column has empty values
nextdate_str = re.compile("\d.*").findall(first_col_empty)[0]
nextdate = datetime.datetime.strptime(nextdate_str, '%m%d%y')
te_date = nextdate - datetime.timedelta(days=1)
print(f"Team entry: {te_date.strftime('%m/%d/%y')}")
# nyt comparison dataset
nyt_date_str = nyt_comparison.date.iloc[-1]
try:
nyt_date = datetime.datetime.strptime(nyt_date_str, '%m/%d/%y')
except ValueError:
nyt_date = datetime.datetime.strptime(nyt_date_str, '%Y-%m-%d')
print(f"NYT comparison dataset: {nyt_date.strftime('%m/%d/%y')}")
# USAF
usaf_date_str = USAF_Cases_comparison.columns[-1]
usaf_date = datetime.datetime.strptime(usaf_date_str, '%m/%d/%Y')
print(f"USAF comparison dataset: {usaf_date.strftime('%m/%d/%y')}")
# JHU
jhu_date_str = JHU_comparison.columns[-1][1:-2]
jhu_date = datetime.datetime.strptime(jhu_date_str, '%Y%m%d')
print(f"JHU comparison dataset: {jhu_date.strftime('%m/%d/%y')} \n")
earliest_date = min([te_date, nyt_date, usaf_date, jhu_date])
print(f"Date you can work on: {earliest_date.strftime('%m/%d/%y')}")
return earliest_date
# given a state and a date (module datetime), output part of data frame you need to work on.
def partTeamEntry(state, date):
yourstateDF = team_entry[team_entry.state == state]
infoDF = yourstateDF.iloc[:, 0:5]
covidDF = yourstateDF.filter(regex=f"{date.strftime('%m%d%y')}$", axis=1)
return pd.concat([infoDF, covidDF], axis=1).iloc[1:,5:]
# return infoDF
def partNYT(state, date):
date_str = date.strftime('%m/%-d/%y')
# date_str = date.strftime('%Y-%m-%d')
return nyt_comparison[(nyt_comparison.date == date_str) & (nyt_comparison.state == abbrev_us_state[state])]
def partUSAFCases(state, date):
date_str = date.strftime('%-m/%-d/%Y')
return USAF_Cases_comparison[USAF_Cases_comparison.State == state].filter(items = [date_str])
def partUSAFDeaths(state, date):
date_str = date.strftime('%-m/%-d/%Y')
return USAF_Deaths_comparison[USAF_Deaths_comparison.State == state].filter(items = [date_str])
def partJHU(state, date):
date_str = [f"x{date.strftime('%Y%m%d')}_c", f"x{date.strftime('%Y%m%d')}_m"]
return JHU_comparison[JHU_comparison.state_ab == state].filter(items = date_str)
def letter_to_num(letter):
return [ord(char) - 97 for char in letter.lower()][0]
def inputData(comp_xlsx, state, date):
from openpyxl import load_workbook
book = load_workbook(comp_xlsx)
writer = pd.ExcelWriter(comp_xlsx, engine='openpyxl')
writer.book = book
## ExcelWriter for some reason uses writer.sheets to access the sheet.
## If you leave it empty it will not know that sheet Main is already there
## and will create a new sheet.
writer.sheets = dict((ws.title, ws) for ws in book.worksheets)
partTeamEntry(state,date).to_excel(writer,
startcol = letter_to_num('C'),
startrow = 3,
header=None, index=False,
sheet_name=list(writer.sheets.keys())[1])
partJHU(state,date).to_excel(writer,
startcol = letter_to_num('I'),
startrow = 3,
header=None, index=False,
sheet_name=list(writer.sheets.keys())[1])
partNYT(state,date).to_excel(writer,
startcol = letter_to_num('L'),
startrow = 3,
header=None, index=False,
sheet_name=list(writer.sheets.keys())[1])
partUSAFCases(state,date).to_excel(writer,
startcol = letter_to_num('V'),
startrow = 3,
header=None, index=False,
sheet_name=list(writer.sheets.keys())[1])
partUSAFDeaths(state,date).to_excel(writer,
startcol = 26,
startrow = 3,
header=None, index=False,
sheet_name=list(writer.sheets.keys())[1])
writer.save()
```
# Main Program
```
today = datetime.date.today()
print("Today's date is", today, '\n')
# state = input('Enter a state you working on: ').upper()
state = 'SD'
print('\n')
date = lastTimeUpdated(state)
# custom date
# last: 2020, 10 ,23
date = datetime.datetime(2020, 11, 25)
partNYT(state,date)
pre = './comparison sheets/'
if state == 'MN':
filename = "Minnesota - Mountains - County Comparison (with probable)"
inputData(f'{pre+filename}.xlsx', state, date)
if state == 'ND':
filename = "North Dakota - Mountains - County Comparison (with probable)"
inputData(f'{pre+filename}.xlsx', state, date)
if state == 'TX':
filename = "Texas - Mountains - County Comparison (with probable)"
inputData(f'{pre+filename}.xlsx', state, date)
if state == 'MT':
filename = "Montana - Mountains - County Comparison (with probable)"
inputData(f'{pre+filename}.xlsx', state, date)
if state == 'SD':
filename = "South Dakota - Mountains - County Comparison (with probable)"
inputData(f'{pre+filename}.xlsx', state, date)
from gspread.urls import SPREADSHEETS_API_V4_BASE_URL
def insert_note(worksheet, label, note):
"""
Insert note ito the google worksheet for a certain cell.
Compatible with gspread.
"""
spreadsheet_id = worksheet.spreadsheet.id
worksheet_id = worksheet.id
row, col = tuple(label) # [0, 0] is A1
url = f"{SPREADSHEETS_API_V4_BASE_URL}/{spreadsheet_id}:batchUpdate"
payload = {
"requests": [
{
"updateCells": {
"range": {
"sheetId": worksheet_id,
"startRowIndex": row,
"endRowIndex": row + 1,
"startColumnIndex": col,
"endColumnIndex": col + 1
},
"rows": [
{
"values": [
{
"note": note
}
]
}
],
"fields": "note"
}
}
]
}
worksheet.spreadsheet.client.request("post", url, json=payload)
```
Save as csv here!
```
read_comp_again = pd.read_csv(f'{pre+filename}.csv', header=1)
read_comp_again = read_comp_again.dropna(how='all')
part_read_again = read_comp_again.loc[:,['County', 'State', 'Unnamed: 21', 'Unnamed: 22']]
part_read_again
# change the order as team entry
part_read_again_wo_total = part_read_again.loc[1:,:]
unknown_row = part_read_again_wo_total.iloc[0,:]
part_read_again_wo_total
part_read_again_wo_total = part_read_again_wo_total.shift(-1)
part_read_again_wo_total.iloc[-1] = unknown_row.squeeze()
part_read_again_wo_total
part_reference = pd.concat([pd.DataFrame(part_read_again.loc[0,:]).transpose(), part_read_again_wo_total])
part_reference.columns = ['County', 'State', 'Case Comments', 'Death Comments']
part_reference
case_comments = part_reference.loc[:, 'Case Comments'].to_list()
death_comments = part_reference.loc[:, 'Death Comments'].to_list()
start_col_idx = [ i for i, col in enumerate(team_entry.columns) if col.endswith(date.strftime('%m%d%y')) ][::2]
if state == 'MT':
start_row_idx = 159 # MT
end_row_idx = 216 # MT
elif state == 'SD':
start_row_idx = 402 # SD
end_row_idx = 469 # SD
start_row_idx -= 1
end_row_idx -= 1
start_col_idx
from xlsxwriter.utility import xl_cell_to_rowcol
xl_cell_to_rowcol('CH159')
# Case
for i, ridx in enumerate(tqdm(range(start_row_idx, end_row_idx+1))):
if i == 0:
continue
if pd.isna(case_comments[i]):
continue
insert_note(team_entry_sheet, (ridx, start_col_idx[0]), case_comments[i])
# insert_note(team_entry, (ridx, cidx))
# Death
for i, ridx in enumerate(tqdm(range(start_row_idx, end_row_idx+1))):
if i == 0:
continue
if pd.isna(death_comments[i]):
continue
insert_note(team_entry_sheet, (ridx, start_col_idx[1]), death_comments[i])
print(date)
print('Case Comment for STATE TOTALS: ', case_comments[0])
print('Death Comment for STATE TOTALS: ', death_comments[0])
```
# Testing
```
test_sheet = gc.open("test").sheet1
test = pd.DataFrame(test_sheet.get_all_records())
insert_comment(test_sheet, (1,2), 'Hello comment')
def insert_comment(service, file_id, content):
"""
Insert a new document-level comment.
Args:
service: Drive API service instance.
file_id: ID of the file to insert comment for.
content: Text content of the comment.
Returns:
The inserted comment if successful, None otherwise.
"""
new_comment = {
'content': content
}
try:
return service.comments().insert(fileId=file_id, body=new_comment).execute()
except:
print('An error occurred: %s')
return None
```
| github_jupyter |
# Experiments
We ran experiments on MNIST with 10 different weight initializations using an overparameterization of $1000$ hidden units. The results are an average over 5 runs for each configuration.
The following command generates the bash commands that will run the experiments. If you do not have a High Performance Computing setup you can remove `--remote` to run them locally in sequence instead (be aware that this might take prohibitively long, however, since there are 50 experiment in total).
**Table of content**:
1. *Setup*.
Loads dependencies and defines helper to construct pandas dataframe.
2. *Train Teacher Network*.
Script for training the teacher network.
3. *Train Student Networks*.
Generates the scripts for running the student networks on a Slurm cluster.
4. *View Student Networks*.
Downloads the output from the cluster, aggregates the experiments in a pandas dataframe, and creates the Figure 1 in the paper.
## Setup
```
import os
import re
import math
import glob
import warnings
import subprocess
import pickle
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
from src.latexify import *
def construct_data_frame(exp_dirs):
"""Construct a pandas dataframe for a list of experiments
using their associated files.
"""
def get_last_entry(filename):
line = subprocess.check_output(['tail', '-1', filename])
try:
entry = line.split()[1:]
return entry
except ValueError:
warnings.warn(f"Experiment {filename} cannot be processed.")
return None
entries = []
for i, exp_dir in enumerate(exp_dirs):
args_fp = open(os.path.join(exp_dir, 'arguments.p'), 'rb')
args = pickle.load(args_fp)
w1 = args.w1
w2 = args.w2
args_fp.close()
print(exp_dir)
training_loss_filename = glob.glob(os.path.join(exp_dir, 'epoch-loss-*.txt'))[0]
test_loss_filename = glob.glob(os.path.join(exp_dir, 'test-loss-*.txt'))[0]
try:
train_loss, train_acc = get_last_entry(training_loss_filename)
test_loss, test_acc = get_last_entry(test_loss_filename)
train_loss = float(train_loss)
test_loss = float(test_loss)
train_acc = float(train_acc)
test_acc = float(test_acc)
except:
train_loss = np.inf
test_loss = np.inf
train_acc = np.inf
test_acc = np.inf
entry = {
'w1': w1,
'w2': w2,
'train_loss': train_loss,
'test_loss': test_loss,
'train_acc': train_acc,
'test_acc': test_acc,
'current_epoch': sum(1 for line in open(training_loss_filename)),
'args': args,
}
entries.append(entry)
df = pd.DataFrame(entries)
return df
```
### Setup He weights
```
# Generate the He initialization
input_dim = 784
output_dim = 10
network_width = 1000
he_w1 = np.sqrt(2 / (input_dim))
he_w2 = np.sqrt(2 / (network_width))
print(he_w1, he_w2)
```
## Train Teacher network
_Note: The resulting model is already stored in `teacher_mode.pth` so there is no need to recreate this._
First we generate the teacher network.
The teacher is trained with cross entropy. Here we generate the script to train it on the server and subsequently move it to the designated teacher network path `./teacher_model.pth`.
```
epoch = 300
batch_size = 128
lr = 0.01
name = 'mnist-he-teacher'
uuid = f"{name}-{he_w1}-{he_w2}"
print(f"python runner.py --exp-name {uuid} --d1 1000 --dataset MNIST --lr {lr} --w1 {he_w1} --w2 {he_w2} --epochs {epoch} --batch-size {batch_size} --remote")
```
_The above output can be run from the shell. Remove `--remote` to run the script locally_.
```
# Move the teacher model to the fixed reserved location
!make pull
subprocess.check_output(["cp", f"output/{uuid}/model.pth", "teacher_model.pth"])
!ls | grep 'teacher_model.pth'
```
## Train Student Networks
```
# Decides for what w1/w2 initialization we should conduct experiments
W1W2 = he_w1 * he_w2
line = np.linspace(0.002, 0.1, 10)
locations = np.array([
line[0],
line[0] + (line[1] - line[0]) / 2,
line[1],
line[1] + (line[2] - line[1]) / 2,
line[2],
line[3],
line[4],
line[5],
line[6],
line[7],
])
plt.scatter(locations, np.repeat(1,10), label="Evaluation locations")
plt.scatter(he_w2, 1, label="He")
plt.legend()
plt.xlabel('$\omega_2$')
None
# Generate the script for running the 50 experiments on the SLURM server
epoch = 300
batch_size = 128
lr = 0.01
W2 = locations
activation = 'gelu'
name = f'mnist-he-student-mse-{activation}'
for i in range(0, 5):
for w2 in W2:
w1 = W1W2 / w2
print(f"python runner.py --exp-name {name}-i{i}-{w1}-{w2} --d1 1000 --dataset MNIST --lr {lr} --w1 {w1} --w2 {w2} --epochs {epoch} --batch-size {batch_size} --activation {activation} --remote --teacher")
# Missing (and onwards): mnist-he-student-mse-gelu-i1-0.03354608550390784-0.06733333333333333
```
_The above output can be run from the shell. Remove `--remote` to run the scripts locally_.
## View Student Networks
```
# Pull the results from the server
!make pull
# Show all experiments
exp_dirs = glob.glob('output/mnist-he-student-mse-gelu-i*')
df = construct_data_frame(exp_dirs)
df = df[df['train_loss'] != np.inf]
df
# Show the aggregrated mean and std for both test and train
he_w2 = 0.044721359549995794
agg_df = df.groupby(np.round(df['w2'], 5)).agg({'test_acc': ['mean', 'std', 'count'], 'train_acc': ['mean', 'std', 'count']})
agg_df = agg_df.sort_values('w2', ascending=False)
agg_df = agg_df[agg_df.index <= 0.08]
agg_df = agg_df[~np.isclose(agg_df.index, he_w2, rtol=1e-04, atol=1e-08)]
agg_df.sort_index()
latexify(fig_width=3.5, fig_height=2)
#latexify(fig_width=7.2)
fig, ax = plt.subplots()
agg_df.train_acc['mean'].plot(yerr=2*agg_df.train_acc['std'], label="Training accuracy", logx=False, ax=ax, color='black', marker='.')
agg_df.test_acc['mean'].plot(yerr=2*agg_df.test_acc['std'], label="Test accuracy", logx=False, ax=ax, color='C0', marker='')
plt.axvline(he_w2, color='black', label="He initialization", zorder=3, linestyle=':')
ax.set_xlabel("$\omega_2$")
ax.set_ylabel("Accuracy")
ax.legend()
ax.set_xlim(0.000, 0.08)
ax.set_ylim(0.96, 1.002)
plt.tight_layout()
savefig(fig, 'mnist-he-generalization-error-mse-gelu.pdf', transparent=True)
```
| github_jupyter |

<table align="left">
<td>
<a target="_blank" href="https://colab.research.google.com/github/antmicro/tensorflow-arduino-examples/blob/master/examples/micro_speech/micro_speech_build_binary.ipynb"><img src="https://raw.githubusercontent.com/antmicro/tensorflow-arduino-examples/master/examples/.static/view-in-colab.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/antmicro/tensorflow-arduino-examples/blob/master/examples/micro_speech/micro_speech_build_binary.ipynb"><img src="https://raw.githubusercontent.com/antmicro/tensorflow-arduino-examples/master/examples/.static/view-ipynb.png" />View ipynb on GitHub</a>
</td>
<td>
<a target="_blank" href="https://github.com/antmicro/tensorflow-arduino-examples/blob/master/examples/micro_speech/micro_speech.py"><img src="https://raw.githubusercontent.com/antmicro/tensorflow-arduino-examples/master/examples/.static/view-source.png" />View Python source on GitHub</a>
</td>
</table>
## Install requirements
```
!pip install -q ffmpeg-python pyaudioconvert git+https://github.com/antmicro/pyrenode.git git+https://github.com/antmicro/renode-colab-tools.git
!apt install -q xxd sox
!mkdir -p renode && cd renode && wget https://dl.antmicro.com/projects/renode/builds/renode-latest.linux-portable.tar.gz && tar -xzf renode-latest.linux-portable.tar.gz --strip 1
!pip install -q -r renode/tests/requirements.txt
!wget -O binary_yes https://dl.antmicro.com/projects/renode/audio_yes_1s.s16le.pcm-s_32000-b69f5518615516f80ae0082fe9b5a5d29ffebce8
!wget -O binary_no https://dl.antmicro.com/projects/renode/audio_no_1s.s16le.pcm-s_32000-f36b60d425c9d5414a38658dbd096313d82d28c5
import os
from renode_colab_tools import audio, metrics
os.environ['PATH'] = os.getcwd()+"/renode:"+os.environ['PATH']
os.environ['TENSORFLOW_PATH'] = os.getcwd()+"/tensorflow-arduino-examples/tensorflow"
!git clone https://github.com/antmicro/tensorflow-arduino-examples.git
!
```
## Get audio
```
audio.audio_options()
```
## Run a micro_speech example in Renode
The command may not be recognized correctly due to https://github.com/tensorflow/tensorflow/pull/45878
```
import time
from pyrenode import *
shutdown_renode()
connect_renode() # this sets up a log file, and clears the simulation (just in case)
tell_renode('using sysbus')
tell_renode('mach create')
tell_renode('machine LoadPlatformDescription @platforms/boards/arduino_nano_33_ble.repl')
tell_renode('sysbus LoadELF @binaries/micro_speech/micro_speech.ino.elf')
tell_renode('logLevel 3')
tell_renode('pdm SetInputFile @audio_bin')
tell_renode('uart0 CreateFileBackend @uart.dump true')
tell_renode('machine EnableProfiler @metrics.dump')
tell_renode('s')
while not os.path.exists('renode/uart.dump'):
time.sleep(1) #waits for creating uart.dump
!timeout 60 tail -f renode/uart.dump | sed '/^.*Heard .*$/ q' | cut -c 2-
tell_renode('q')
expect_cli('Renode is quitting')
time.sleep(1) #wait not to kill Renode forcefully
shutdown_renode()
```
## Renode metrics analysis
```
from renode.tools.metrics_analyzer.metrics_parser import MetricsParser
metrics.init_notebook_mode(connected=False)
parser = MetricsParser('renode/metrics.dump')
metrics.configure_plotly_browser_state()
metrics.show_executed_instructions(parser)
metrics.configure_plotly_browser_state()
metrics.show_memory_access(parser)
metrics.configure_plotly_browser_state()
metrics.show_peripheral_access(parser)
metrics.configure_plotly_browser_state()
metrics.show_exceptions(parser)
```
| github_jupyter |
# Archive analysis
```
# Set root folder
import sys
import os
cwd = os.getcwd()
folder = os.path.basename(cwd)
cwd = os.path.dirname(cwd)
folder = os.path.basename(cwd)
ROOT = os.path.join(cwd)
sys.path.append(ROOT)
import numpy as np
from core.population import Archive
from environments.environments import *
from parameters import params
import pickle as pkl
import analysis.utils as utils
import matplotlib
import matplotlib.pyplot as plt
from mpl_toolkits.axes_grid1 import make_axes_locatable
import json
from scipy.stats.mstats import gmean
plt.rcParams['pdf.fonttype'] = 42
plt.rcParams['ps.fonttype'] = 42
font = {'size' : 18}
plt.rc('font', **font)
%matplotlib inline
```
### Get experiment data
```
env_type = 'cur'
if env_type == 'RW':
env = 'Walker2D'
elif env_type == 'Dummy':
env = 'Dummy'
elif env_type == 'CB':
env = 'CollectBall'
elif env_type == 'AM':
env = 'AntMaze'
elif env_type == 'arm':
env = 'NDofArm'
elif env_type == 'maze':
env = 'HardMaze'
elif env_type == 'cur':
env = 'Curling'
else:
raise ValueError('Wrong environment type given: {}'.format(env_type))
# EXP_ROOT = '/mnt/7e0bad1b-406b-4582-b7a1-84327ae60fc4/cmans/'
exp_path = os.path.join(ROOT, 'experiment_data')
# exp_path = EXP_ROOT
exp_types = [
'FitNS_all_gen',
'NS_all_gen',
'NSGA-II_all_gen',
'CMA-ME_all_gen',
'ME_all_gen',
'RND_all_gen',
]
paths = []
save_path = os.path.join(exp_path, "{}".format(env))
avail_exps = []
for exp_type in exp_types:
path = os.path.join(exp_path, "{}/{}_{}".format(env, env, exp_type))
# path = os.path.join(exp_path, "{}_{}".format(env, exp_type))
if not os.path.exists(path):
path = None
print("No {} experiment for {} env.".format(exp_type, env))
continue
paths.append(path)
avail_exps.append(exp_type)
exp_types = avail_exps
archive_gen = -1
max_len = registered_envs[env]['max_steps']
```
## Load analyzed data
```
def load_data(path):
data_path = os.path.join(path)
if os.path.exists(data_path):
try:
with open(data_path, 'rb') as file:
rd = pkl.load(file)
except Exception as e:
print('Error on {}'.format(data_path))
raise
else:
print("{} does not exists.".format(data_path))
return rd
cvgs = {}
unif = {}
final_gt_bd = {}
rew_eval = {}
recaps = {}
for exp, path in zip(exp_types, paths):
runs = utils.get_runs_list(path)
runs_cvg = []
runs_unif = []
runs_final_gt_bd = []
runs_rew_eval = []
runs_recaps = []
for run in runs:
if os.path.exists(os.path.join(path, run, 'analyzed_data/cvg_unif_by_eval.pkl')):
data = load_data(os.path.join(path, run, 'analyzed_data/cvg_unif_by_eval.pkl'))
runs_cvg.append(data['cvg'])
runs_unif.append(data['unif'])
if os.path.exists(os.path.join(path, run, 'analyzed_data/final_gt_bd.pkl')):
runs_final_gt_bd.append(load_data(os.path.join(path, run, 'analyzed_data/final_gt_bd.pkl')))
if os.path.exists(os.path.join(path, run, 'analyzed_data/rew_by_eval.pkl')):
runs_rew_eval.append(load_data(os.path.join(path, run, 'analyzed_data/rew_by_eval.pkl')))
if os.path.exists(os.path.join(path, run, 'recap.json')):
with open(os.path.join(path, run, 'recap.json'), 'rb') as f:
runs_recaps.append(json.load(f))
cvgs[exp] = np.array(runs_cvg);
unif[exp] = np.array(runs_unif);
final_gt_bd[exp] = np.array(runs_final_gt_bd);
rew_eval[exp] = np.array(runs_rew_eval);
recaps[exp] = runs_recaps
params.load(os.path.join(path, run, '_params.json'));
grid_parameters = registered_envs[env]['grid']
best_run = {}
for exp, path in zip(exp_types, paths):
if exp in cvgs:
try:
best_run[exp] = np.argmax(np.max(cvgs[exp], axis=1))
except ValueError:
best_run[exp] = 0
```
## Plot final BD
```
from matplotlib.patches import Rectangle
fig, axes = plt.subplots(1, len(exp_types), figsize=(len(exp_types)*10, 10))
fig.patch.set_facecolor('white')
axes = [axes]
if len(exp_types) == 1:
axes = np.swapaxes([axes], 0, 1)
for exp_idx, exp in enumerate(exp_types):
print("Working on exp: {}".format(exp))
name = exp.split('_')[0]
archive = final_gt_bd[exp][best_run[exp]]['archive']
rew_archive = final_gt_bd[exp][best_run[exp]]['rew archive']
axes[0][exp_idx].scatter(archive[:, 0], archive[:, 1])
if len(rew_archive)> 0:
axes[0][exp_idx].scatter(rew_archive[:, 0], rew_archive[:, 1])
axes[0][exp_idx].set_xticks(ticks=[]);
axes[0][exp_idx].set_yticks(ticks=[]);
axes[0][exp_idx].set_xlim(grid_parameters['min_coord'][0], grid_parameters['max_coord'][0]);
axes[0][exp_idx].set_ylim(grid_parameters['min_coord'][1], grid_parameters['max_coord'][1]);
axes[0][exp_idx].set_title('{} BD'.format(name), fontsize=22);
plt.subplots_adjust(wspace=.05, left=0.01, bottom=0.01, right=0.99, top=0.99)
with open(os.path.join(save_path, 'archive_points.png'), 'wb') as f:
plt.savefig(f, format='png');
plt.show();
```
## Plot CVG and REW
```
rew = {}
cvg = {}
areas = len(rew_eval[exp_types[0]][0])-1
fig, axes = plt.subplots(areas+1, 1, figsize=(4, 3*areas+1))
fig.patch.set_facecolor('white')
names = []
for exp_idx, exp in enumerate(exp_types):
linewidth = 1
name = exp.split('_')[0]
if name == 'FitNS':
name = 'SERENE'
linewidth = 2.5
names.append(name)
# Reward
# ----------------------------------------------------
rew[exp] = {i: [] for i in range(areas)}
for run in rew_eval[exp]:
# Reformat reward data as {exp: {area: [runs]}}
for idx, r_area in enumerate(run):
if r_area != 'eval':
rew[exp][idx].append(run[r_area])
else:
x_points = run['eval']
for idx in range(areas):
r_mean = np.mean(rew[exp][idx], axis=0)
r_std = np.std(rew[exp][idx], axis=0)
axes[idx].plot(x_points, r_mean, linewidth=linewidth)
axes[idx].fill_between(x_points, r_mean-r_std, r_mean+r_std, alpha=0.1)
axes[idx].set_title('Max Reward: Area {}'.format(idx+1))
axes[idx].set_xlabel('Evaluation steps x10^4')
axes[idx].set_ylabel('Reward')
# ----------------------------------------------------
# Coverage
# ----------------------------------------------------
cvg_mean = np.mean(cvgs[exp], axis=0)
cvg_std = np.std(cvgs[exp], axis=0)
axes[-1].plot(x_points, cvg_mean, linewidth=linewidth)
axes[-1].fill_between(x_points, cvg_mean-cvg_std, cvg_mean+cvg_std, alpha=0.1)
axes[-1].set_title('Coverage')
axes[-1].set_xlabel('Evaluation steps x10^4')
axes[-1].set_ylabel('Coverage %')
# ----------------------------------------------------
for ax in axes:
xtickslocs = ax.get_xticks()
# ax.set_xticklabels(np.array(xtickslocs/10000))
ax.set_ylim(-.1, 1.1)
for line, name in zip(ax.lines, names):
y = line.get_ydata()[-1]
ax.annotate(name, xy=(.97,y), xytext=(20,0), color=line.get_color(),
xycoords = ax.get_yaxis_transform(), textcoords="offset points",
size=14, va="center",
)
plt.tight_layout()
plt.savefig(os.path.join(save_path, 'perf_by_eval.pdf'), format='pdf')
plt.show();
```
## Plot Budget
```
solutions = {}
areas = len(rew_eval[exp_types[0]][0])-1
# Extract focus data from recaps
# ----------------------------------------------------
for exp in exp_types:
solutions[exp] = {i: [] for i in range(areas)}
for run in recaps[exp]:
for idx in range(areas):
try:
solutions[exp][idx].append(run['rew_area_{}'.format(idx+1)])
except:
solutions[exp][idx].append(0)
# ----------------------------------------------------
# Plot
# ----------------------------------------------------
fig, axes = plt.subplots(1, 1, figsize=(5, 5))
fig.patch.set_facecolor('white')
data = []
for pos, exp in enumerate(exp_types):
data.append([np.mean(solutions[exp][i]) for i in range(areas)])
data[-1].append(params.evaluation_budget - np.sum(data[-1]))
data = np.transpose(np.array(data))[::-1]/params.evaluation_budget
labels = ['Expl']
bar = []
for idx, dd in enumerate(data):
if idx == 0:
bottom = np.zeros_like(dd)
else:
bottom += data[idx-1]
labels.append('Area {}'.format(idx))
bar.append(axes.bar(range(len(exp_types)), dd, bottom=bottom, linewidth=0, alpha=.9))
axes.set_ylim(0, 1)
axes.set_xticks(np.array(range(len(exp_types))))
axes.set_xticklabels(names, rotation=45)
axes.set_title(env);
axes.set_ylabel('Budget %')
axes.legend(bar, labels);
axes.legend(bar, labels, loc="lower left");
with open(os.path.join(save_path, 'area_perf.svg'), 'w') as f:
plt.savefig(f, format='svg')
plt.show()
# ----------------------------------------------------
```
| github_jupyter |

In [part 1]({static}/ros-analysis-part-1.html) we looked at how to generate data
for the Bobble-Bot simulator. Either go back and read that post, or download the data
[here](https://github.com/super-owesome/bobble_controllers/raw/master/analysis/notebooks/RosJupyterAnalysis/data/sample_data.zip)
and keep reading. With your data ready, we can begin our analysis using Jupyter Notebook. First,
let's get our analysis environment setup.
All of the dependencies can be installed using pip.
```bash
sudo apt-get install python-pip python-dev build-essential
pip install pandas jupyter rosbag_pandas seaborn
```
Let's use the sample analysis script provided in the bobble_controllers repository
to test that we now have the needed packages.
```bash
cd ~/bobble_workspace
catkin build
source devel/setup.bash
cd ~/src/bobble_controllers/analysis
python make_plots.py -r impulse_force.bag
```
If all went well a TiltControl.png and VelocityControl.png file will be created in
the working directory. These plots should look like this:


The analysis script can be used to make these two plots for any of the runs (including
the data you produced in part 1). Here's how to use it:
```bash
python make_plots.py --help
usage: make_plots.py [-h] [-r RUN] [-o OUT]
optional arguments:
-h, --help show this help message and exit
-r RUN, --run RUN Bag file for simulation run of interest
-o OUT, --out OUT Output directory.
```
## Using the analysis_tools module
The [make_plots](https://github.com/super-owesome/bobble_controllers/blob/master/analysis/make_plots.py)
Python script provides a good introduction to making plots of
ROS data using [Pandas](https://pandas.pydata.org/) and
[Matplotlib](https://matplotlib.org/). For your convenience, the bobble_controllers
repository provides a helper Python module to facilitate making plots of the
Bobble-Bot simulation data. The [analysis_tools module](https://github.com/super-owesome/bobble_controllers/tree/master/src/analysis_tools)
is written to be fairly generic so that it can be adapted to more general ROS
data analysis. Check it out, and feel free to adapt it to your own project.
Our simulation data is stored in ROS bag format. Pandas DataFrames are a more
convenient data representation for a Python+Jupyter based analysis. Our bag files can be
quickly converted to this format by making use of the provided analysis_tools.parsing module.
This article discusses how the parsing module [loads a ROS bag file into Pandas](https://nimbus.unl.edu/2014/11/using-rosbag_pandas-to-analyze-rosbag-files/) in
more detail. Here's the relevant snippet of code that loads the simulation's bag
file into a Pandas DataFrame.
```python
from analysis_tools.parsing import parse_single_run
print "Loading sim data from run : " + str(run_name) + " ... "
df = {}
df[run_name] = parse_single_run(sim_data_bag_file)
```
Of course, this post is about a Jupyter Notebook based analysis. Let's see
how we can use the analysis_tools module to load data into a notebook.
## Loading data into Jupyter
The first step is to activate your environment and load Jupyter Notebook. To help
you follow along, the Jupyter Notebooks for this series can be found [here](https://github.com/super-owesome/bobble_controllers/tree/master/analysis/notebooks/RosJupyterAnalysis).
```bash
cd ~/bobble_workspace
source devel/setup.bash
jupyter
```
With Jupyter now open, navigate to the folder containing your data and .ipynb file.
In that folder, you will also find a file called nb_env.py. This file is used to import
the needed modules and load all the bag files found in the data directory. Feel free to
replace the contents of the data directory with the simulation data you generated from
[part 1]({static}/ros-analysis-part-1.html). To load this file and view the simulation runs available for analysis, use the cell shown below.
```
# Load anaylsis environment file. This file defines data directories
# and imports all needed Python packages for this notebook.
exec(open("nb_env.py").read())
# Print out the df dictionary keys (the test run names)
df.keys()
```
We now have all the data from the simulation runs generated in
[part 1]({static}/ros-analysis-part-1.html). Let's use Pandas to demonstrate some
basic manipulation of the simulation data.
### Print sim data in tabular form
We can print the first five rows from the 'balance' run in a nice tabular form like so:
```
n_rows = 5
df['balance'].loc[:, 'Tilt':'TurnRate'].head(n_rows)
```
### Search for a column
Here's how to search for a column(s) in a DataFrame:
```
search_string = 'Vel'
found_data = df['no_balance'].filter(regex=search_string)
found_data.head()
```
### Get all column names
You can view all of the data available in a given run like so:
```
df['balance'].dtypes.index
```
### Find the maximum value of a column
```
print "Max tilt no_balance run: %0.2f " % df['no_balance']['Tilt'].max()
print "Max tilt balance run: %0.2f " % df['balance']['Tilt'].max()
```
Many more data processing and manipulation functions are possible. Consult the
[Pandas documentation](https://pandas.pydata.org/pandas-docs/version/0.21/generated/pandas.DataFrame.html)
for more information.
## What's Next?
This post has demonstrated how to make a few plots of our simulation data from the command
line. It also covered how to load the simulation runs into Jupyter Notebook. Finally, we explored
using various Pandas functions to manipulate and print out this data within Jupyter.
In [part 3]({static}/ros-analysis-part-3), we will show how to post
process this data using [NumPy](http://www.numpy.org/), and then make some plots
that capture the Bobble-Bot balance controller's performance.
| github_jupyter |
# Инициализация
```
#@markdown - **Монтирование GoogleDrive**
from google.colab import drive
drive.mount('GoogleDrive')
# #@markdown - **Размонтирование**
# !fusermount -u GoogleDrive
```
# Область кодов
```
#@title Учебные пособия TensorFlow Dataset class { display-mode: "both" }
# Эта программа кратко представляет использование класса Dataset в TensorFlow
# Dataset tutorials
import tensorflow as tf
import tensorflow_datasets as tfds
import numpy as np
import os
#@markdown - **Настройка параметров**
tf.logging.set_verbosity(tf.logging.ERROR)
initial_rate = 1e-3 #@param {type: "number"}
scale_factor = 60000 #@param {type: "number"}, scale factor of l2 regularization, default is 3000
batchsize = 256 #@param {type: "integer"}, batch size, default is 128
num_epochs = 6000 #@param {type: "integer"}
event_path = './Tensorboard' #@param {type: "string"}
checkpoint_path = './Checkpoints' #@param {type: "string"}
# online_test = True #@param {type: "boolean"}
#@markdown - **Считывание данных изображений**
mnist_train = tfds.as_numpy(tfds.load("mnist", split=tfds.Split.TRAIN, batch_size=-1))
imgs_train, labels_train = mnist_train['image'] / 255., mnist_train['label']
mnist_test = tfds.as_numpy(tfds.load("mnist", split=tfds.Split.TEST, batch_size=-1))
imgs_test, labels_test = mnist_test['image'] / 255., mnist_test['label']
num_samples = labels_test.shape[0]
assert imgs_train.max() == 1., "warning: 'The value of the pixel is preferably between 0 and 1.'"
#@markdown - **Структура graph**
graph = tf.Graph()
with graph.as_default():
# learning rate с экспоненциальным затуханием
global_step = tf.Variable(0, name='global_step', trainable=False)
decay_steps = 400
decay_rate = 0.90
learning_rate = tf.train.exponential_decay(initial_rate,
global_step=global_step,
decay_steps=decay_steps,
decay_rate=decay_rate,
staircase=True,
name='learning_rate')
#@markdown - **Настройка итератора**
x_p = tf.placeholder(tf.float32, shape=[None, 28, 28, 1], name='input_images')
y_p = tf.placeholder(tf.int64, shape=[None, ], name='labels')
batch_size = tf.placeholder(tf.int64, name='batch_size')
data = tf.data.Dataset.from_tensor_slices((x_p, y_p))
data = data.repeat()
data_batch = data.shuffle(2).batch(batch_size).prefetch(batch_size)
iterator = data_batch.make_initializable_iterator()
with tf.name_scope('Input'):
x_input, y_input = iterator.get_next()
y = tf.one_hot(y_input, depth=10)
keep_pro = tf.placeholder(tf.float32)
with tf.name_scope('Conv1'):
w_1 = tf.Variable(tf.truncated_normal([3, 3, 1, 32], stddev=0.1), name='weights_conv1')
b_1 = tf.Variable(tf.constant(0.1, shape=[32]), name='bias_conv1')
h_conv1 = tf.nn.relu(tf.nn.conv2d(x_input, w_1, strides=[1, 1, 1, 1], padding='SAME') + b_1)
h_pool1 = tf.nn.max_pool(h_conv1, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='SAME')
with tf.name_scope('Conv2'):
w_2 = tf.Variable(tf.truncated_normal([3, 3, 32, 64], stddev=0.1), name='weights_conv2')
b_2 = tf.Variable(tf.constant(0.1, shape=[64]), name='bias_conv2')
h_conv2 = tf.nn.relu(tf.nn.conv2d(h_pool1, w_2, strides=[1, 1, 1, 1], padding='SAME') + b_2)
h_pool2 = tf.nn.max_pool(h_conv2, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='SAME')
with tf.name_scope('FC1'):
h_pool2_fla = tf.layers.flatten(h_pool2)
num_f = h_pool2_fla.get_shape().as_list()[-1]
w_fc1 = tf.Variable(tf.truncated_normal([num_f, 128], stddev=0.1), name='weights_fc1')
b_fc1 = tf.Variable(tf.constant(0.1, shape=[128]), name='bias_fc1')
h_fc1 = tf.nn.relu(tf.matmul(h_pool2_fla, w_fc1) + b_fc1)
h_drop1 = tf.nn.dropout(h_fc1, keep_prob=keep_pro, name='Dropout')
with tf.name_scope('Output'):
w_fc2 = tf.Variable(tf.truncated_normal([128, 10], stddev=0.1), name='weights_fc2')
b_fc2 = tf.Variable(tf.constant(0.1, shape=[10]), name='bias_fc2')
h_fc2 = tf.matmul(h_drop1, w_fc2) + b_fc2
#@markdown - **L2 регуляризация**
tf.add_to_collection(tf.GraphKeys.WEIGHTS, w_fc1)
tf.add_to_collection(tf.GraphKeys.WEIGHTS, w_fc2)
regularizer = tf.contrib.layers.l2_regularizer(scale=15./scale_factor)
reg_tem = tf.contrib.layers.apply_regularization(regularizer)
with tf.name_scope('loss'):
entropy_loss = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(labels=y, logits=h_fc2))
entropy_loss = tf.add(entropy_loss, reg_tem, name='l2_loss')
with tf.name_scope('accuracy'):
prediction = tf.cast(tf.equal(tf.arg_max(h_fc2, 1), tf.argmax(y, 1)), "float")
accuracy = tf.reduce_mean(prediction)
with tf.name_scope('train'):
optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate)
train_op = optimizer.minimize(entropy_loss, global_step=global_step)
#@markdown - **Summaries**
tf.summary.image('input_images', x_input, max_outputs=3, collections=['train'])
tf.summary.histogram('conv1_weights', w_1, collections=['train'])
tf.summary.histogram('conv1_bias', b_1, collections=['train'])
tf.summary.scalar('loss', entropy_loss, collections=['train'])
tf.summary.scalar('accuracy', accuracy, collections=['train'])
tf.summary.scalar('global_step', global_step, collections=['train'])
tf.summary.scalar('learning_rate', learning_rate, collections=['train'])
summ_train = tf.summary.merge_all('train')
#@markdown - **Обучение модели**
max_acc = 100.
min_loss = 0.1
with tf.Session(graph=graph) as sess:
sess.run(tf.global_variables_initializer())
sess.run(iterator.initializer, feed_dict={x_p: imgs_train, y_p: labels_train, batch_size: batchsize})
summ_train_dir = os.path.join(event_path, 'summaries','train')
summ_train_Writer = tf.summary.FileWriter(summ_train_dir)
summ_train_Writer.add_graph(sess.graph)
saver = tf.train.Saver(max_to_keep=1)
print('Training ========== (。・`ω´・) ========\n')
for num in range(num_epochs):
_, acc, loss, rs = sess.run([train_op, accuracy, entropy_loss, summ_train], feed_dict={keep_pro: 0.5,
batch_size: batchsize})
summ_train_Writer.add_summary(rs, global_step=num)
acc *= 100
num_e = str(num + 1)
print_list = [num_e, loss, acc]
if (num + 1) % 400 == 0:
print('Keep on training ========== (。・`ω´・) ========')
print('Epoch {0[0]}, train_loss is {0[1]:.4f}, accuracy is {0[2]:.2f}%.\n'.format(print_list))
print('\n', 'Training completed.')
#@markdown - **Набор тестовых данных**
print('Testing ========== (。・`ω´・) ========')
sess.run(iterator.initializer, feed_dict={x_p: imgs_test, y_p: labels_test, batch_size: num_samples})
acc, loss = sess.run([accuracy, entropy_loss], feed_dict={keep_pro: 1., batch_size: num_samples})
acc *= 100
print_list = [loss, acc]
print('Test_loss is {0[0]:.4f}, accuracy is {0[1]:.2f}%.\n'.format(print_list))
summ_train_Writer.close()
# sess.close()
```
| github_jupyter |
## MNIST Handwritten Digits Classification Experiment
This demo shows how you can use SageMaker Experiment Management Python SDK to organize, track, compare, and evaluate your machine learning (ML) model training experiments.
You can track artifacts for experiments, including data sets, algorithms, hyper-parameters, and metrics. Experiments executed on SageMaker such as SageMaker Autopilot jobs and training jobs are automatically tracked will be automatically tracked. You can also track artifacts for additional steps within an ML workflow that come before/after model training e.g. data pre-processing or post-training model evaluation.
The APIs also let you search and browse your current and past experiments, compare experiments, and identify best performing models.
Now we will demonstrate these capabilities through an MNIST handwritten digits classification example. The experiment will be organized as follow:
1. Download and prepare the MNIST dataset.
2. Train a Convolutional Neural Network (CNN) Model. Tune the hyper parameter that configures the number of hidden channels in the model. Track the parameter configurations and resulting model accuracy using SageMaker Experiments Python SDK.
3. Finally use the search and analytics capabilities of Python SDK to search, compare and evaluate the performance of all model versions generated from model tuning in Step 2.
4. We will also see an example of tracing the complete linage of a model version i.e. the collection of all the data pre-processing and training configurations and inputs that went into creating that model version.
Make sure you selected `Python 3 (Data Science)` kernel.
### Install Python SDKs
```
import sys
!{sys.executable} -m pip install sagemaker-experiments
```
### Install PyTroch
```
!{sys.executable} -m pip install torch
!{sys.executable} -m pip install torchvision
```
### Setup
```
import time
import boto3
import numpy as np
import pandas as pd
%config InlineBackend.figure_format = 'retina'
from matplotlib import pyplot as plt
from torchvision import datasets, transforms
import sagemaker
from sagemaker import get_execution_role
from sagemaker.session import Session
from sagemaker.analytics import ExperimentAnalytics
from smexperiments.experiment import Experiment
from smexperiments.trial import Trial
from smexperiments.trial_component import TrialComponent
from smexperiments.tracker import Tracker
sess = boto3.Session()
sm = sess.client('sagemaker')
role = get_execution_role()
```
### Create a S3 bucket to hold data
```
# create a s3 bucket to hold data, note that your account might already created a bucket with the same name
account_id = sess.client('sts').get_caller_identity()["Account"]
bucket = 'sagemaker-experiments-{}-{}'.format(sess.region_name, account_id)
prefix = 'mnist'
try:
if sess.region_name == "us-east-1":
sess.client('s3').create_bucket(Bucket=bucket)
else:
sess.client('s3').create_bucket(Bucket=bucket,
CreateBucketConfiguration={'LocationConstraint': sess.region_name})
except Exception as e:
print(e)
```
### Dataset
We download the MNIST hand written digits dataset, and then apply transformation on each of the image.
```
# download the dataset
# this will not only download data to ./mnist folder, but also load and transform (normalize) them
train_set = datasets.MNIST('mnist', train=True, transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.1307,), (0.3081,))]),
download=True)
test_set = datasets.MNIST('mnist', train=False, transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.1307,), (0.3081,))]),
download=False)
plt.imshow(train_set.data[2].numpy())
```
After transforming the images in the dataset, we upload it to s3.
```
inputs = sagemaker.Session().upload_data(path='mnist', bucket=bucket, key_prefix=prefix)
print('input spec: {}'.format(inputs))
```
Now lets track the parameters from the data pre-processing step.
```
with Tracker.create(display_name="Preprocessing", sagemaker_boto_client=sm) as tracker:
tracker.log_parameters({
"normalization_mean": 0.1307,
"normalization_std": 0.3081,
})
# we can log the s3 uri to the dataset we just uploaded
tracker.log_input(name="mnist-dataset", media_type="s3/uri", value=inputs)
```
### Step 1 - Set up the Experiment
Create an experiment to track all the model training iterations. Experiments are a great way to organize your data science work. You can create experiments to organize all your model development work for : [1] a business use case you are addressing (e.g. create experiment named “customer churn prediction”), or [2] a data science team that owns the experiment (e.g. create experiment named “marketing analytics experiment”), or [3] a specific data science and ML project. Think of it as a “folder” for organizing your “files”.
### Create an Experiment
```
mnist_experiment = Experiment.create(
experiment_name=f"mnist-hand-written-digits-classification-{int(time.time())}",
description="Classification of mnist hand-written digits",
sagemaker_boto_client=sm)
print(mnist_experiment)
```
### Step 2 - Track Experiment
### Now create a Trial for each training run to track the it's inputs, parameters, and metrics.
While training the CNN model on SageMaker, we will experiment with several values for the number of hidden channel in the model. We will create a Trial to track each training job run. We will also create a TrialComponent from the tracker we created before, and add to the Trial. This will enrich the Trial with the parameters we captured from the data pre-processing stage.
Note the execution of the following code takes a while.
```
from sagemaker.pytorch import PyTorch
hidden_channel_trial_name_map = {}
```
If you want to run the following training jobs asynchronously, you may need to increase your resource limit. Otherwise, you can run them sequentially.
```
preprocessing_trial_component = tracker.trial_component
for i, num_hidden_channel in enumerate([2, 5, 10, 20, 32]):
# create trial
trial_name = f"cnn-training-job-{num_hidden_channel}-hidden-channels-{int(time.time())}"
cnn_trial = Trial.create(
trial_name=trial_name,
experiment_name=mnist_experiment.experiment_name,
sagemaker_boto_client=sm,
)
hidden_channel_trial_name_map[num_hidden_channel] = trial_name
# associate the proprocessing trial component with the current trial
cnn_trial.add_trial_component(preprocessing_trial_component)
# all input configurations, parameters, and metrics specified in estimator
# definition are automatically tracked
estimator = PyTorch(
entry_point='./mnist.py',
role=role,
sagemaker_session=sagemaker.Session(sagemaker_client=sm),
framework_version='1.1.0',
train_instance_count=1,
train_instance_type='ml.c4.xlarge',
hyperparameters={
'epochs': 2,
'backend': 'gloo',
'hidden_channels': num_hidden_channel,
'dropout': 0.2,
'optimizer': 'sgd'
},
metric_definitions=[
{'Name':'train:loss', 'Regex':'Train Loss: (.*?);'},
{'Name':'test:loss', 'Regex':'Test Average loss: (.*?),'},
{'Name':'test:accuracy', 'Regex':'Test Accuracy: (.*?)%;'}
],
enable_sagemaker_metrics=True,
)
cnn_training_job_name = "cnn-training-job-{}".format(int(time.time()))
# Now associate the estimator with the Experiment and Trial
estimator.fit(
inputs={'training': inputs},
job_name=cnn_training_job_name,
experiment_config={
"TrialName": cnn_trial.trial_name,
"TrialComponentDisplayName": "Training",
},
wait=True,
)
# give it a while before dispatching the next training job
time.sleep(2)
```
### Compare the model training runs for an experiment
Now we will use the analytics capabilities of Python SDK to query and compare the training runs for identifying the best model produced by our experiment. You can retrieve trial components by using a search expression.
### Some Simple Analyses
```
search_expression = {
"Filters":[
{
"Name": "DisplayName",
"Operator": "Equals",
"Value": "Training",
}
],
}
trial_component_analytics = ExperimentAnalytics(
sagemaker_session=Session(sess, sm),
experiment_name=mnist_experiment.experiment_name,
search_expression=search_expression,
sort_by="metrics.test:accuracy.max",
sort_order="Descending",
metric_names=['test:accuracy'],
parameter_names=['hidden_channels', 'epochs', 'dropout', 'optimizer']
)
trial_component_analytics.dataframe()
```
To isolate and measure the impact of change in hidden channels on model accuracy, we vary the number of hidden channel and fix the value for other hyperparameters.
Next let's look at an example of tracing the lineage of a model by accessing the data tracked by SageMaker Experiments for `cnn-training-job-2-hidden-channels` trial
```
lineage_table = ExperimentAnalytics(
sagemaker_session=Session(sess, sm),
search_expression={
"Filters":[{
"Name": "Parents.TrialName",
"Operator": "Equals",
"Value": hidden_channel_trial_name_map[2]
}]
},
sort_by="CreationTime",
sort_order="Ascending",
)
lineage_table.dataframe()
```
| github_jupyter |
# Create a Target Forecast
* **About 40 mins may be elapsed**
```
import boto3
from time import sleep
import pandas as pd
import json
import time
import pprint
import numpy as np
# Recover variables stored by other notebooks
%store -r
session = boto3.Session(region_name=region)
forecast = session.client(service_name='forecast')
```
## Create Prophet and DeepAR+ Campaign
```
# Prophet
prophet_forecastName = project+'_prophet_algo_forecast' + target_suffix + suffix
prophet_create_forecast_response=forecast.create_forecast(
ForecastName=prophet_forecastName,
PredictorArn=target_prophet_predictorArn)
target_prophet_forecast_arn = prophet_create_forecast_response['ForecastArn']
forecast.describe_forecast(ForecastArn = target_prophet_forecast_arn)
# DeepAR+
deeparp_forecastName = project+'_deeparp_algo_forecast' + target_suffix + suffix
deeparp_create_forecast_response=forecast.create_forecast(
ForecastName=deeparp_forecastName,
PredictorArn=target_deepar_predictorArn)
target_deeparp_forecast_arn = deeparp_create_forecast_response['ForecastArn']
forecast.describe_forecast(ForecastArn = target_deeparp_forecast_arn)
%%time
# Check the Prophet status
while True:
createProphetStatus = forecast.describe_forecast(ForecastArn= target_prophet_forecast_arn)['Status']
createDeeparpStatus = forecast.describe_forecast(ForecastArn= target_deeparp_forecast_arn)['Status']
print("Prophet: ", createProphetStatus)
print("DeepARP: ", createProphetStatus)
if createProphetStatus != 'ACTIVE' and createProphetStatus != 'CREATE_FAILED':
sleep(60)
elif createDeeparpStatus != 'ACTIVE' and createDeeparpStatus != 'CREATE_FAILED':
sleep(60)
else:
break
```
## Upload forecast results to S3
```
target_prophet_path = "s3://" + bucket_name + "/" + bucket_folder + "/prophet_" + target_suffix + suffix + "/"
target_prophet_job_name = "ProphetExport1" + target_suffix + suffix
create_forecast_export_job_prophet_response = forecast.create_forecast_export_job(
ForecastExportJobName = target_prophet_job_name,
ForecastArn = target_prophet_forecast_arn,
Destination={
"S3Config" : {
"Path": target_prophet_path,
"RoleArn": role_arn
}
})
TargetForecastProphetExportJobArn = create_forecast_export_job_prophet_response["ForecastExportJobArn"]
forecast.describe_forecast_export_job(ForecastExportJobArn = TargetForecastProphetExportJobArn)
target_deeparp_path = "s3://" + bucket_name + "/" + bucket_folder + "/deeparp_" + target_suffix + suffix + "/"
target_deeparp_job_name = "DeepARPExport1" + target_suffix + suffix
create_forecast_export_job_deepar_response = forecast.create_forecast_export_job(
ForecastExportJobName = target_deeparp_job_name,
ForecastArn = target_deeparp_forecast_arn,
Destination={
"S3Config" : {
"Path": target_deeparp_path,
"RoleArn": role_arn
}
})
TargetForecastDeeparExportJobArn = create_forecast_export_job_deepar_response["ForecastExportJobArn"]
forecast.describe_forecast_export_job(ForecastExportJobArn = TargetForecastDeeparExportJobArn)
%%time
# Check the Prophet status
while True:
createProphetStatus = forecast.describe_forecast_export_job(ForecastExportJobArn= TargetForecastProphetExportJobArn)['Status']
createDeeparpStatus = forecast.describe_forecast_export_job(ForecastExportJobArn= TargetForecastDeeparExportJobArn)['Status']
print("Prophet: ", createProphetStatus)
print("DeepARP: ", createProphetStatus)
if createProphetStatus != 'ACTIVE' and createProphetStatus != 'CREATE_FAILED':
sleep(60)
elif createDeeparpStatus != 'ACTIVE' and createDeeparpStatus != 'CREATE_FAILED':
sleep(60)
else:
break
%store target_prophet_forecast_arn
%store target_deeparp_forecast_arn
%store TargetForecastProphetExportJobArn
%store TargetForecastDeeparExportJobArn
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib as mpl
import matplotlib.pyplot as plt
%matplotlib inline
#Graph Styling
# https://tonysyu.github.io/raw_content/matplotlib-style-gallery/gallery.html
plt.style.use('seaborn-darkgrid')
```
# Scatter Graphs
```
x1 = np.array([250,150,350,252,450,550,455,358,158,355])
y1 =np.array([40,50,80, 90, 100,50,60,88,54,45])
x2 = np.array([200,100,300,220,400,500,450,380,180,350])
y2 = np.array([400,500,800, 900, 1000,500,600,808,504,405])
#Graph - 1
plt.scatter(x1,y1)
plt.xlabel('$Time $ $ Spent$' , fontsize = 12)
plt.ylabel('$Score$' , fontsize = 12)
plt.title ('Scatter Graph')
plt.show()
#Graph - 2
plt.scatter(x2,y2 ,color = 'r')
plt.xlabel('$Time $ $ Spent$' , fontsize = 12)
plt.ylabel('$Score$' , fontsize = 12)
plt.title ('Scatter Graph')
plt.show()
#Graph - 3
plt.scatter(x1,y1 ,label = 'Class 1')
plt.scatter(x2,y2 ,label = 'Class 2',color ='r')
plt.xlabel('$Time $ $ Spent$' , fontsize = 12)
plt.ylabel('$Score$' , fontsize = 12)
plt.title ('Scatter Graph')
plt.legend()
plt.show()
#Graph - 4
plt.scatter(x1,y1 ,label = 'Class 1',marker='o' , color = 'm')
plt.scatter(x2,y2 ,label = 'Class 2',marker='v',color ='r')
plt.xlabel('$Time $ $ Spent$' , fontsize = 12)
plt.ylabel('$Score$' , fontsize = 12)
plt.title ('Scatter Graph')
plt.legend()
plt.show()
plt.figure(figsize=(10,6))
x = np.random.normal(0,10,1000)
y = np.random.normal(0,10,1000)
plt.scatter(x,y)
plt.show()
plt.figure(figsize=(8,6))
x = np.random.random(10)
y = np.random.random(10)
# "alpha" is used for softnening colors
plt.scatter(np.random.random(10),np.random.random(10),c='r', s=50 , alpha=0.6 , label = 'One' )
plt.scatter(np.random.random(10),np.random.random(10),c='b', s=100 , alpha=0.6 , label = 'Two')
plt.scatter(np.random.random(10),np.random.random(10),c='g', s=150 , alpha=0.6 , label = 'Three')
plt.scatter(np.random.random(10),np.random.random(10),c='y', s=200 , alpha=0.6 , label = 'Four')
plt.legend(bbox_to_anchor=(1.0, 1.0) , shadow=True, fontsize='x-large')
plt.show()
# Changing label color
plt.figure(figsize=(8,6))
x = np.random.random(10)
y = np.random.random(10)
# "alpha" is used for softnening colors
plt.rcParams['text.color'] = 'red' # Label Color
plt.scatter(np.random.random(10),np.random.random(10),c='r', s=50 , alpha=0.6 , label = 'One' )
plt.scatter(np.random.random(10),np.random.random(10),c='b', s=100 , alpha=0.6 , label = 'Two')
plt.scatter(np.random.random(10),np.random.random(10),c='g', s=150 , alpha=0.6 , label = 'Three')
plt.scatter(np.random.random(10),np.random.random(10),c='y', s=200 , alpha=0.6 , label = 'Four')
plt.legend(bbox_to_anchor=(1.0, 1.0) , shadow=True, fontsize='x-large')
plt.show()
```
| github_jupyter |
```
import os
import numpy as np
import pandas as pd
from datetime import datetime, timedelta
import datetime as dt
import xarray as xr
from math import atan2, log
import uuid
#read in email
text_file = open('C:/Users/gentemann/Google Drive/f_drive/docs/projects/saildrone/baja/docs/email_for_file.txt', "r")
email_str = text_file.readlines()
#definitions and things you might have to set differently for each file
itow_mask1=45 #see code just a bit below for figure to determine where to set flags here
itow_mask2=-110 #see code just a bit below for figure to determine where to set flags here
ISDP = 'Saildrone'
SST_type = 'SSTdepth'
Annex_version = '01.1'
File_version = '01.0'
astr_platform='SD1002'
astr_title = 'Data from Saildrone cruise from SF to Guadalupe Island April-June 2018'
astr_uuid = str(uuid.uuid4()) #'0f410de6-4ba5-4f79-af20-8a57a445f454'
#input filename
dir_in='f:/data/cruise_data/saildrone/baja-2018/daily_files/sd-1002/2018/'
dir_out='f:/data/cruise_data/saildrone/baja-2018/other_format_shared_data/'
droplist=['WWND_STDDEV', 'CHLOR_MEAN','RH_MEAN','WWND_MEAN','O2_CONC_STDDEV','CDOM_STDDEV',
'TEMP_O2_STDDEV','BARO_PRES_MEAN','TEMP_O2_MEAN','SAL_STDDEV','TEMP_AIR_MEAN',
'CDOM_MEAN','SAL_MEAN','O2_SAT_MEAN','CHLOR_STDDEV', 'COND_STDDEV', 'COND_MEAN',
'BKSCT_RED_MEAN', 'TEMP_IR_MEAN', 'O2_SAT_STDDEV','O2_CONC_MEAN', 'TEMP_AIR_STDDEV',
'BARO_PRES_STDDEV', 'TEMP_IR_STDDEV', 'VWND_STDDEV','RH_STDDEV', 'GUST_WND_STDDEV', 'GUST_WND_MEAN',
'BKSCT_RED_STDDEV', 'UWND_STDDEV','HDG_WING','WING_ANGLE']
#read data back in, into two arrays one with time encoding and one without
adir='F:/data/cruise_data/saildrone/baja-2018/'
filename_usv=adir+'saildrone-gen_5-baja_2018-sd1002-20180411T180000-20180611T055959-1_minutes-v2.1535585233403.nc'
with xr.open_dataset(filename_usv,drop_variables=droplist) as ds:
lons_usv=ds.longitude[0,:]
lats_usv=ds.latitude[0,:]
time_usv=ds.time[0,:]
with xr.open_dataset(filename_usv,drop_variables=droplist,decode_times=False) as ds_raw:
time_usv_raw=ds_raw.time[0,:]
mint=time_usv.min().data
maxt=time_usv.max().data
ds.attrs['time_coverage_start']=str(np.datetime64(mint,'ms'))+'Z'
ds.attrs['time_coverage_end']=str(np.datetime64(maxt,'ms'))+'Z'
ilen=(len(ds.latitude['obs']))
#dates_usv64=dataset_decodetime.TIME[0,:].values
#time_usv_pd=pd.to_datetime(time_usv, unit='ns')
print(ilen)
#calculate the average distance between observations as the spatial resolution global attribute
#import math
#from math import cos
# approximate radius of earth in km
R = 6373.0 #km
lat1 = np.deg2rad(lats_usv[1:ilen])
lon1 = np.deg2rad(lons_usv[1:ilen])
lat2 = np.deg2rad(lats_usv[0:ilen-1])
lon2 = np.deg2rad(lons_usv[0:ilen-1])
dlon = lon2 - lon1
dlat = lat2 - lat1
a = np.sin(dlat / 2)**2 + np.cos(lat1) * np.cos(lat2) * np.sin(dlon / 2)**2
c = 2 * np.arctan2(a**.5, (1 - a)**.5)
distance = R * c
#add global attributes that are missing
#some of these will need to be changed for new cruises
dataset=ds
gattrs = dataset.attrs.copy()
gattrs['title'] = astr_title
gattrs['summary'] = 'none'
gattrs['references'] = 'none'
gattrs['institution'] = 'Saildrone'
gattrs['history'] = 'Saildrone 6-hourly v1 files were used to create this file'
gattrs['comment'] = 'none'
gattrs['license'] = 'CC-BY-NC' #'free and open'
gattrs['id'] = 'SSTdepth'
gattrs['naming_authority'] = 'org.shipborne-radiometer'
gattrs['product_version'] = '1.0'
gattrs['uuid'] = astr_uuid
gattrs['l3r_version_id'] = '1.1'
gattrs['netcdf_version_id'] = '4.6.1'
gattrs['date_created'] = dt.datetime.now().strftime("%Y-%m-%dT%H:%M:%SZ") #yyyy-mm-ddThh:mm:ssZ
gattrs['file_quality_level'] = 3
gattrs['spatial_resolution'] = str(distance.mean().data*1000)+' m'
gattrs['start_time'] = ds.time_coverage_start
gattrs['stop_time'] = ds.time_coverage_end
gattrs['northernmost_latitude'] = lats_usv.max().data
gattrs['geospatial_lat_max'] = lats_usv.max().data
gattrs['southernmost_latitude'] = lats_usv.min().data
gattrs['geospatial_lat_min'] = lats_usv.min().data
gattrs['easternmost_longitude'] = lons_usv.max().data
gattrs['geospatial_lon_max'] = lons_usv.max().data
gattrs['westernmost_longitude'] = lons_usv.min().data
gattrs['geospatial_lon_min'] = lons_usv.min().data
gattrs['geospatial_lat_units'] = 'degrees_north'
gattrs['geospatial_lon_units'] = 'degrees_east'
gattrs['source'] = 'SSTdepth, wind_speed'
gattrs['platform'] = astr_platform
gattrs['sensor'] = str(dataset.TEMP_CTD_MEAN[0,0].device_name + ', ' + \
dataset.UWND_MEAN[0,0].device_name)
#Teledyne CTD (2632)_Teledyne CTD (2632)_2632, Gill Anemometer (172107)_Gill Anemometer (172107)_172107
#print(gattrs['sensor'])
gattrs['metadata_link'] = 'TBD'
gattrs['keywords'] = 'Oceans > Ocean Temperature > Sea Surface Temperature'
gattrs['keywords_vocabulary'] = 'NASA Global Change Master Directory (GCMD) Science Keywords'
gattrs['acknowledgment'] = 'The Schmidt Family Foundation, Saildrone, NASA Physical Oceanography'
gattrs['project'] = 'International Shipborne Radiometer Network'
gattrs['publisher_name'] = 'The ISRN Project Office'
gattrs['publisher_url'] = 'http://www.shipborne.radiometer.org'
gattrs['publisher_email'] = str(email_str[0])
#print(gattrs['publisher_email'])
gattrs['processing_level'] = '1.0'
#del dataset.attrs['nodc_template_version']
# convert names
COORD_ATTR = 'lat lon time'
#COORD_ATTR = 'time'
FLOAT_FILLVALUE = np.finfo(np.float32).min
DOUBLE_FILLVALUE = np.finfo(np.float64).min
VAR_TRANSLATE = {
'TEMP_CTD_MEAN': 'sea_water_temperature',
'COG': 'course_over_ground',
'latitude': 'lat',
'longitude': 'lon',
'HDG': 'true_bearing',
'ROLL': 'platform_roll',
'PITCH': 'platform_pitch',
'SOG': 'speed_over_ground',
'TEMP_CTD_STDDEV':'sst_total_uncertainty'
}
# copy variables from source dataset
varrays = {}
for v in dataset.data_vars:
if v in VAR_TRANSLATE:
# print(v)
# set attributes
vattrs = dataset[v].attrs.copy()
if v not in ['latitude', 'longitude', 'time']:
vattrs["coordinates"] = COORD_ATTR
vattrs["_FillValue"] = FLOAT_FILLVALUE
values = np.ma.fix_invalid(dataset[v].values[0,:])
if v == 'TEMP_CTD_MEAN':
# convert SST to Kelvin
values += 273.15
# create new data array
varrays[VAR_TRANSLATE[v]] = xr.DataArray(
values.filled(FLOAT_FILLVALUE).astype(np.float32,copy=False),
dims=['time'],
attrs=vattrs
)
for v in dataset.coords:
if v in VAR_TRANSLATE:
# print(v)
# set attributes
vattrs = dataset[v].attrs.copy()
if v not in ['latitude', 'longitude', 'time']:
vattrs["coordinates"] = COORD_ATTR
vattrs["_FillValue"] = FLOAT_FILLVALUE
values = np.ma.fix_invalid(dataset[v].values[0,:])
if v == 'TEMP_CTD_MEAN':
# convert SST to Kelvin
values += 273.15
# create new data array
varrays[VAR_TRANSLATE[v]] = xr.DataArray(
values.filled(FLOAT_FILLVALUE).astype(np.float32,copy=False),
dims=['time'],
attrs=vattrs
)
# 1. wind speed
vattrs = dataset.UWND_MEAN.attrs.copy()
vattrs['standard_name'] = 'wind_speed'
vattrs['long_name'] = 'wind_speed'
vattrs['valid_min'] = 0
vattrs['valid_max'] = 100
vattrs["_FillValue"] = FLOAT_FILLVALUE
vattrs['source'] = 'anemometer'
vattrs['comment'] = 'Instrument located at to of Saildrone mast at ' +\
str(dataset.UWND_MEAN.installed_height)+' m' + '. This was adjusted ' +\
'to 10 m as ws_10m = ws*log(10./1e-4))/log(WS_height/1e-4'
vattrs['height'] = '10 m' #str(str(dataset.UWND_MEAN.installed_height)+' m')
vattrs["coordinates"] = COORD_ATTR
WS=(dataset['UWND_MEAN'].values[0,:]**2 + dataset['VWND_MEAN'].values[0,:]**2)**.5
WS_height=int(dataset.UWND_MEAN.installed_height)
WS_10m = (WS*log(10./1e-4))/log(WS_height/1e-4)
varrays['wind_speed'] = xr.DataArray(
WS_10m.astype(np.float32,copy=False),
dims=['time'],
attrs=vattrs
)
# 2. wind direction
vattrs = dataset.UWND_MEAN.attrs.copy()
vattrs['standard_name'] = 'wind_to_direction'
vattrs['long_name'] = 'local wind direction'
vattrs['valid_min'] = 0
vattrs['valid_max'] = 360
vattrs["_FillValue"] = FLOAT_FILLVALUE
vattrs['units'] = 'degrees'
vattrs['source'] = 'anemometer'
vattrs['height'] = str(str(dataset.UWND_MEAN.installed_height)+' m')
vattrs["coordinates"] = COORD_ATTR
WD=np.arctan2(dataset.VWND_MEAN.values[0,:], dataset.UWND_MEAN.values[0,:])*180/3.14159
WD=np.where(WD<0,WD+360,WD)
varrays['wind_direction'] = xr.DataArray(
WD.astype(np.float32,copy=False),
dims=['time'],
attrs=vattrs
)
# 3. time
vattrs = ds_raw.time.attrs.copy()
vattrs["_FillValue"] = DOUBLE_FILLVALUE
values = np.ma.fix_invalid(ds_raw.time.values[0,:])
vtime = xr.DataArray(
values.filled(FLOAT_FILLVALUE).astype(np.float64,copy=False),
dims=['time'],
attrs=vattrs
)
# 4. Quality level
flag_bytes=np.byte((0,1,2,3,4,5)) #bytearray([0,1,2,3,4,5])
iqual_byte = np.ones(shape=time_usv.shape, dtype='b')*5 #change byte to b1
iqual_byte[:itow_mask1] = 2 #set at top of file from looking at data
iqual_byte[itow_mask2:] = 2
vattrs = {
'long_name': 'measurement quality value',
'flag_meanings': 'no_data bad_data worst_quality low_quality acceptable_quality best_quality',
'flag_values': flag_bytes,
'_FillValue': -128
}
varrays['quality_level'] = xr.DataArray(
iqual_byte,
dims=['time'],
attrs=vattrs
)
# create Dataset and save
l3r = xr.Dataset(
varrays,
coords = {
# 'time': time_usv
'time': vtime
},
attrs=gattrs
)
# embellishments
l3r.sea_water_temperature.attrs['valid_min']=260.0
l3r.sea_water_temperature.attrs['valid_max']=320.0
l3r.sea_water_temperature.attrs['units']='kelvin'
l3r.sea_water_temperature.attrs['long_name']='sea surface depth temperature at 0.6m'
l3r.time.attrs['standard_name']='time'
l3r.time.attrs['long_name']='time'
l3r.lon.attrs['standard_name']='longitude'
l3r.lon.attrs['long_name']='longitude'
l3r.lat.attrs['long_name']='latitude'
l3r.lat.attrs['standard_name']='latitude'
l3r.true_bearing.attrs['long_name']='platform true bearing'
l3r.true_bearing.attrs['standard_name']='platform_orientation'
l3r.speed_over_ground.attrs['long_name']='platform speed over ground'
l3r.sst_total_uncertainty.attrs['standard_name']='sea_water_temperature standard error'
l3r.sst_total_uncertainty.attrs['long_name']=' sea water temperature total uncertainty'
l3r.sst_total_uncertainty.attrs['valid_min']=0.0
l3r.sst_total_uncertainty.attrs['valid_max']=2.0
l3r.sst_total_uncertainty.attrs['units']='kelvin'
indicative_date_time=pd.to_datetime(str(time_usv[0].data)).strftime("%Y%m%d%H%M%S")
Product_string = astr_platform # str(ds.TEMP_CTD_MEAN.vendor_name) + '_' + str(ds.TEMP_CTD_MEAN.serial_number)
filename_L3R = dir_out + indicative_date_time + \
'-' + ISDP + '-' + 'L3R' + '-' + SST_type + '-' +Product_string+ '-v' +Annex_version+ '-fv' +File_version+ '.nc'
filename_L3R_test = dir_out + indicative_date_time + \
'-' + ISDP + '-' + 'L3R' + '-' + SST_type + '-' +Product_string+ '-v' +Annex_version+ '-fv' +File_version+ 'test.nc'
filename_L3R
l3r.to_netcdf(filename_L3R)
#for some reason the time not decoded is off by about 28 seconds so reset to original decoded time here
#l3r['time']=ds.time[0,:].data
#l3r.to_netcdf(filename_L3R)
#print(WD[100:800])
#print(WD.max())
#print(l3r.wind_direction[300:400])
#l3r.wind_direction[0:100].plot()
#filename_l3R_test=''
ds_test=xr.open_dataset(filename_L3R_test)
ds_test.time
```
| github_jupyter |
```
from geoscilabs.em.DipoleWidgetTD import DipoleWidgetTD, InteractiveDipoleProfileTD, InteractiveDipoleDecay
from geoscilabs.em.VolumeWidget import InteractivePlanes, plotObj3D
from matplotlib import rcParams
rcParams['font.size'] = 16
```
# Magnetic Dipole in a Whole-space (time domain)
## Purpose
By using an analytic solution electromagnetic (EM) fields from electrical dipole in a Whole-space, we present some fundamentals of EM responses in the context of crosswell EM survey.
# Set up
For time domain EM method using inductive source, we inject step-off currents to an induction coil, which will generate time varying magnetic field in the earth.
<img src="https://github.com/geoscixyz/geosci-labs/blob/main/images/em/stepoff_currents.png?raw=true"></img>
## Crosswell EM geometry
Here, we choose geometric parameters for a crosswell EM set-up having two boreholes for Tx and Rx. In the Tx hole, a VED source is located at (0m, 0m, 0m), and it is fixed. Horizontal location of the Rx hole is fixed to 50m apart from the source location in x-direction.
```
ax = plotObj3D()
```
## Backgrounds
When using crosswell electromagnetic (EM) survey, we inject step-off currents to the earth using induction coil, and measure magnetic field at receiver locations in the off-time, when DC effects are disappeared. A common goal here is imaging conductivity structure of the earth by interpreting measured voltages. However, to accomplish that task well, we first need to understand physical behavior of EM responses for the given survey set-up.
Assuming length of the area of the induction coil is small enough, this can be assumed as magnetic dipole (ED). For a croswell set-up, let we have a vertical magnetic dipole (VMD) source in a homogeneous earth with step-off currents, then we can have analytic solution of EM fields in time domain (WH1988). Solution of of arbitrary EM fields, $\mathbf{f}$, will be a function of
$$ \mathbf{f} (x, y, z; \sigma, t),$$
where $\sigma$ is conductivity of homogenous earth (S/m), and $f$ is transmitting frequency (Hz). Here $\mathbf{f}$ can be electic ($\mathbf{e}$) or magnetic field ($\mathbf{h}$), or current density ($\mathbf{j}$). <strong>Now, you will explore how EM responses behaves as a function of space, $\sigma$, and $t$ for the given crosswell EM set-up </strong>.
# Geometry app
Here, we choose geometric parameters for a crosswell EM set-up having two boreholes for Tx and Rx. In the Tx hole, a VED source is located at (0m, 0m, 0m), and it is fixed. Horizontal location of the Rx hole is fixed to 50m apart from the source location in x-direction.
## Parameters
- plane: Choose either "XZ" or "YZ" plane
- offset: Offset from a source plane (m)
- nRx: The number of receivers in the Rx hole
Chosen geometric parameters will be used in the Electric Dipole widget below.
```
Q0 = InteractivePlanes()
Q0
```
# Magnetic Dipole app
Explore behavior of EM fields, $\mathbf{f} (x, y, z; \sigma, t)$ on 2D plane chosen in the above app. And also at the receiver locations.
## Parameters:
- Field: Type of EM fields ("E": electric field, "H": magnetic field, "J": current density)
- AmpDir: Type of the vectoral EM fields
None: $f_x$ or $f_y$ or $f_z$
Amp: $\mathbf{f} \cdot \mathbf{f} = |\mathbf{f}|^2$
Dir: A vectoral EM fields, $\mathbf{f}$
- Comp.: Direction of $\mathbf{F}$ at Rx locations
- $t$: time after current switch-off
- $\sigma$: Conductivity of homogeneous earth (S/m)
- Offset: Offset from a source plane
- Scale: Choose "log" or "linear" scale
- Slider: When it is checked, it activates "flog" and "siglog" sliders above.
- TimeLog: A float slider for log10 time (only activated when slider is checked)
- SigLog: A float slider for log10 conductivity (only activated when slider is checked)
```
dwidget = DipoleWidgetTD()
Q1 = dwidget.InteractiveDipoleBH(nRx=Q0.kwargs["nRx"], plane=Q0.kwargs["Plane"], offset_plane=Q0.kwargs["Offset"], SrcType="MD", fieldvalue="H")
Q1
```
# Proflie app
Here we focuson data, which can be measured at receiver locations. We limit our attention to three different profile shown in **Geometry** app: Rxhole (red), Txhole (black), TxProfile (blue).
## Parameters:
- Comp.: Direction of $\mathbf{F}$ at Rx locations
- ComplexNumber: Type of complex data ("Re", "Im", "Amp", "Phase")
- $t_1$: Time (sec)
- $t_2$: Time (sec)
- $t_3$: Time (sec)
- Profile: Type of profile line ("Rxhole", "Txhole", "TxProfile")
- Scale: Choose "log" or "linear" scale
- Rx#: choice of Rx point for the following Sounding app
```
Q2 = InteractiveDipoleProfileTD(dwidget, Q1.kwargs["Sigma"], Q1.kwargs["Field"], Q1.kwargs["Component"], Q1.kwargs["Scale"])
Q2
```
# Sounding app
## Parameters:
- Comp.: Direction of $\mathbf{F}$ at Rx locations
- $\sigma$: Conductivity of homogeneous earth (S/m)
- Scale: Choose "log" or "linear" scale
```
InteractiveDipoleDecay(dwidget, dwidget.dataview.xyz_line[Q2.kwargs["irx"],:], Q1.kwargs["Field"], Q1.kwargs["Component"])
```
| github_jupyter |
# Load the Dataset
* annotated json files have to be stored in the `data/processed` dir.
```
%load_ext autoreload
%autoreload 2
import os
import sys
sys.path.append(os.path.abspath(os.path.join("..")))
import pandas as pd
import numpy as np
from scripts.features.reader import Reader
from scripts.features.post_feature import PostFeature
from scripts.features.length_extractor import TitleLengthExtractor, SectionCountExtractor, Header1MeanLengthExtractor, Header2MeanLengthExtractor, SentenceMeanLengthExtractor, SentenceMaxLengthExtractor, SentenceMinLengthExtractor
from scripts.features.charactor_extractor import RenderedBodyPreprocessor, KanjiRatioExtractor, HiraganaRatioExtractor, KatakanaRatioExtractor, NumberRatioExtractor, PunctuationRatioExtractor
from scripts.features.structure_extractor import ItemizationCountExtractor, FormulaCountExtractor, ImageCountExtractor, ItemizationRatioExtractor, ImageRatioExtractor, FormulaRatioExtractor
reader = Reader()
pf_dicts = []
for p in reader.post_iterator():
pf = PostFeature(p)
cleaned_rendered_body = RenderedBodyPreprocessor().clean_rendered_body(p.rendered_body)
pf.add(TitleLengthExtractor())
pf.add(SectionCountExtractor())
pf.add(KanjiRatioExtractor(cleaned_rendered_body))
pf.add(HiraganaRatioExtractor(cleaned_rendered_body))
pf.add(KatakanaRatioExtractor(cleaned_rendered_body))
pf.add(NumberRatioExtractor(cleaned_rendered_body))
pf.add(PunctuationRatioExtractor(cleaned_rendered_body))
pf.add(SentenceMeanLengthExtractor(cleaned_rendered_body))
# pf.add(Header1MeanLengthExtractor())
# pf.add(Header2MeanLengthExtractor())
pf.add(SentenceMeanLengthExtractor(cleaned_rendered_body))
pf.add(SentenceMaxLengthExtractor(cleaned_rendered_body))
# pf.add(SentenceMinLengthExtractor(cleaned_rendered_body))
# pf.add(ItemizationCountExtractor())
# pf.add(FormulaCountExtractor())
pf.add(ImageCountExtractor())
# pf.add(ItemizationRatioExtractor(cleaned_rendered_body))
# pf.add(FormulaRatioExtractor(cleaned_rendered_body))
pf.add(ImageRatioExtractor(cleaned_rendered_body))
pf_d = pf.to_dict(drop_disused_feature=False) # default True -> drop title, body etc fields
pf_dicts.append(pf_d)
pf_df = pd.DataFrame(pf_dicts)
pf_df.head(5)
# drop disused features
if "post_id" in pf_df.columns:
pf_df.drop(pf_df[["post_id", "title", "body", "url", "user_id", 'rendered_body']], axis=1, inplace=True)
quality = pf_df["quality"]
pf_df.drop("quality", axis=1, inplace=True)
# normalize
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
pf_df_n = scaler.fit_transform(pf_df)
import matplotlib.pyplot as plt
from sklearn.ensemble import RandomForestClassifier
from sklearn.svm import SVC
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report
# estimate accuracy by cross validation
clf = RandomForestClassifier(n_estimators=15, max_features=0.5)
scores = cross_val_score(clf, pf_df_n, quality, cv=2, scoring="f1")
print(scores)
# train and show score
train_f, test_f, train_lb, test_lb = train_test_split(pf_df_n, quality, test_size=0.2, random_state=42)
clf.fit(train_f, train_lb)
pred = clf.predict(test_f)
print(classification_report(test_lb, pred, target_names=["bad", "good"]))
# show feature importance
importances = clf.feature_importances_
std = np.std([tree.feature_importances_ for tree in clf.estimators_], axis=0)
indices = np.argsort(importances)[::-1]
# Plot the feature importances of the forest
labels = np.array(pf_df.columns.values.tolist())
plt.figure()
plt.title("Feature importances")
plt.bar(range(len(labels)), importances[indices], color="r", yerr=std[indices], align="center")
plt.xticks(range(len(labels)), labels[indices], rotation="vertical")
plt.xlim([-1, len(labels)])
plt.ylim([0, 1])
plt.tight_layout()
plt.show()
from scripts.models.save_models import SaveModelsScalor
# save model
SaveModelsScalor(clf, scaler, pf_df)
```
| github_jupyter |
# Generic LA tool
```
import numpy as np
import pandas as pd
import seaborn as sns
from tqdm import tqdm
from src.dlla.berg import make_mlp
from src.dlla.hw import dlla_p_gradient, prepare_traces_dl
from src.dlla.wegener import make_mlp_wegener, wegener_p_gradient
from src.pollution.tools import file_suffix
from src.tools.cache import cache_np
from src.tools.la import fixed_fixed
from src.tools.plotter import init_plots, plot_p_gradient, PALETTE_GRADIENT, store_sns, TVLA_PALETTE, DLLA_PALETTE
from src.trace_set.database import Database
from src.trace_set.set_hw import TraceSetHW
from src.trace_set.transform import reduce_fixed_fixed
from src.tvla.cri import tvla_t_test, rho_test
from src.tvla.tvla import prepare_tvla
init_plots()
# EDIT This line for other databases.
DB = Database.aisy
# ---
TRACE_SET = TraceSetHW(DB)
POLLUTION_TYPE = None
FILE_SUFFIX = file_suffix(POLLUTION_TYPE, 0)
NUM_CLASSES = 9
Y_LIM = 10 ** -40
TRACES_NAME = ""
if DB is Database.aisy:
TRACES_NAME = "AISy lab traces"
elif DB is Database.ascad_none:
TRACES_NAME = "ASCAD unprotected traces"
elif DB is Database.ascad:
TRACES_NAME = "ASCAD masked traces"
REPEAT_DLLA = 30
REPEAT_TVLA = 100
if DB is Database.aisy:
REPEAT_TVLA = 400
REPEAT_DLLA = 100
PROF_X, PROF_Y = TRACE_SET.profile()
PVS_T = tvla_t_test(*fixed_fixed(PROF_X, PROF_Y), progress=True)
np.min(PVS_T[1])
ORDER = 1 + (DB == Database.ascad)
G = sns.lineplot(data={f"TVLA $t$-test $\mu_{ORDER}$": PVS_T[ORDER]}, palette=[TVLA_PALETTE[-1]])
sns.lineplot(data= [10 ** -5] * len(PVS_T[ORDER]), color="#FF000080", linestyle="--", label="Threshold")
G.invert_yaxis()
G.set(yscale="log", ylabel="$p$-value", xlabel="Sample point index", title=f"{TRACES_NAME}, leakage location", ylim=(10 ** 0, 10 ** -128))
store_sns(G, f"{DB.name}-pvs-t-{FILE_SUFFIX}")
np.min(PVS_T)
WINDOW_SIZE = PROF_X.shape[1]
TITLE_SUB = f"{TRACES_NAME} with {WINDOW_SIZE} sample points"
def point_ixs(num_traces, min_traces=2, num_points=200, small_sample=1000):
# Large-scale point indexes (all traces)
point_ix_lg = np.linspace(min_traces, num_traces, num_points).astype(int)
# Small-scale point indexes (first traces)
point_ix_sm = np.linspace(min_traces, min(num_traces, small_sample), num_points).astype(int)
return np.union1d(point_ix_sm, point_ix_lg)
def shuffle(x, y):
ix = np.arange(len(x))
np.random.shuffle(ix)
return x[ix], y[ix]
def single_tvla_t_pg(x, y, min_pvs, order):
min_sp = min_pvs.argmin()
num_traces = len(x)
x_lim = np.array(x[:, min_sp:min_sp + 1])
ixs = point_ixs(num_traces)
pvs = np.ones(ixs.shape)
for ix, limit in enumerate(ixs):
pvs[ix] = np.min(tvla_t_test(x_lim[:limit], y[:limit], order, progress=False)[order])
return np.interp(np.arange(num_traces), ixs, pvs)
def tvla_t_p_gradient(x, y, min_pvs, order, times=REPEAT_TVLA):
res = []
for _ in tqdm(range(times), "TVLA t-test p-gradient"):
res.append(single_tvla_t_pg(*shuffle(x, y), min_pvs, order))
return np.median(res, axis=0)
LIM_MASK = np.ones(len(PROF_X)).astype(bool)
if DB is Database.ascad:
LIM_MASK[94463] = False
print(PROF_Y[94463])
PG2 = tvla_t_p_gradient(PROF_X[LIM_MASK], PROF_Y[LIM_MASK], PVS_T[2], 2, 1)
TVLA_X, TVLA_Y = prepare_tvla(PROF_X, PROF_Y)
TVLA_X_LIM, TVLA_Y_LIM = prepare_tvla(PROF_X[LIM_MASK], PROF_Y[LIM_MASK])
if DB is Database.ascad:
HIGH = TVLA_X[TVLA_Y]
HIGH_LIM = TVLA_X_LIM[TVLA_Y_LIM]
def with_and_without(b):
num_traces = len(b)
res = np.zeros(num_traces)
variance_all = b.var()
for ix in tqdm(range(num_traces)):
mask = np.ones(num_traces).astype(bool)
mask[ix] = False
var_diff = np.abs(b[mask].var() - variance_all)
res[ix] = var_diff
return res
ALL = cache_np("with_and_without", with_and_without, HIGH[:, np.argmin(PVS_T[2])])
MASK_LIM_TOP_2 = np.ones(len(HIGH_LIM)).astype(bool)
MASK_LIM_TOP_2[ALL.argsort()[-2]] = False
# It's in B!
G = sns.lineplot(data={"Trace 94463": abs(HIGH_LIM.var(axis=0) - HIGH.var(axis=0)), "Trace 0": abs(HIGH_LIM[MASK_LIM_TOP_2].var(axis=0) - HIGH_LIM.var(axis=0))})
G.set(title="Variance difference between trace sets\nwith and without a certain trace")
if DB is Database.ascad:
G = sns.lineplot(data={"Mean trace": np.mean(PROF_X[LIM_MASK], axis=0), "Trace 94463": PROF_X[94463]})
G.set(title="ASCAD database, most desynchronised trace.")
store_sns(G, "ascad-anomalous-trace")
PVS_T = tvla_t_test(TVLA_X_LIM, TVLA_Y_LIM, progress=True)
TVLA_PG_T_1 = cache_np(f"{DB.name}-tvla-t-1", tvla_t_p_gradient, TVLA_X_LIM, TVLA_Y_LIM, PVS_T[1], 1)
TVLA_PG_T_2 = cache_np(f"{DB.name}-tvla-t-2", tvla_t_p_gradient, TVLA_X_LIM, TVLA_Y_LIM, PVS_T[2], 2)
TVLA_PG_T_3 = cache_np(f"{DB.name}-tvla-t-3", tvla_t_p_gradient, TVLA_X_LIM, TVLA_Y_LIM, PVS_T[3], 3)
def single_tvla_rho_pg(x, y, min_pvs):
min_sp = min_pvs.argmin()
num_traces = len(x)
x_lim = np.array(x[:, min_sp:min_sp + 1])
ixs = point_ixs(num_traces)
pvs = np.ones(ixs.shape)
for ix, limit in enumerate(ixs):
pvs[ix] = np.min(rho_test(x_lim[:limit], y[:limit], progress=False))
return np.interp(np.arange(num_traces), ixs, pvs)
def tvla_rho_p_gradient(x, y, min_pvs, times=REPEAT_TVLA):
res = []
for _ in tqdm(range(times), "TVLA rho-test p-gradient"):
res.append(single_tvla_rho_pg(*shuffle(x, y), min_pvs))
return np.median(res, axis=0)
PVS_RHO = rho_test(TVLA_X_LIM, TVLA_Y_LIM, progress=True)[0]
TVLA_PG_RHO = cache_np(f"{DB.name}-tvla-rho", tvla_rho_p_gradient, TVLA_X_LIM, TVLA_Y_LIM, PVS_RHO)
if DB is Database.ascad:
PVS_T_WITH = tvla_t_test(TVLA_X, TVLA_Y, progress=True)
TVLA_PG_T_1_WITH = cache_np(f"{DB.name}-tvla-t-1-with", tvla_t_p_gradient, TVLA_X, TVLA_Y, PVS_T[1], 1)
TVLA_PG_T_2_WITH = cache_np(f"{DB.name}-tvla-t-2-with", tvla_t_p_gradient, TVLA_X, TVLA_Y, PVS_T[2], 2)
TVLA_PG_T_3_WITH = cache_np(f"{DB.name}-tvla-t-3-with", tvla_t_p_gradient, TVLA_X, TVLA_Y, PVS_T[3], 3)
PVS_RHO_WITH = rho_test(TVLA_X, TVLA_Y, progress=True)[0]
TVLA_PG_RHO_WITH = cache_np(f"{DB.name}-tvla-rho-with", tvla_rho_p_gradient, TVLA_X_LIM, TVLA_Y_LIM, PVS_RHO_WITH)
G = sns.lineplot(data={"$t$-test, $\mu_1$": TVLA_PG_T_1_WITH, "$t$-test, $\mu_2$": TVLA_PG_T_2_WITH, "$t$-test, $\mu_3$": TVLA_PG_T_3_WITH, "$\\rho$-test": TVLA_PG_RHO_WITH}, palette=np.array(TVLA_PALETTE)[[4, 3, 2, 1]])
sns.lineplot(data=[10 ** -5] * len(TVLA_PG_T_1), color="#FF000080", linestyle="--", label="Threshold")
G.set(yscale='log', ylim=(1, Y_LIM), xlabel="Number of traces", ylabel="$p$-value", title=f"TVLA, different kernel methods\n{TITLE_SUB}") #, xlim=(94314,94320))
store_sns(G, f"{DB.name}-tvla-with")
G = sns.lineplot(data={"$t$-test, $\mu_1$": TVLA_PG_T_1, "$t$-test, $\mu_2$": TVLA_PG_T_2, "$t$-test, $\mu_3$": TVLA_PG_T_3, "$\\rho$-test": TVLA_PG_RHO}, palette=np.array(TVLA_PALETTE)[[4, 3, 2, 1]])
sns.lineplot(data=[10 ** -5] * len(TVLA_PG_T_1), color="#FF000080", linestyle="--", label="Threshold")
G.set(yscale='log', ylim=(1, Y_LIM), xlabel="Number of traces", ylabel="$p$-value", title=f"TVLA, different kernel methods\n{TITLE_SUB}") #, xlim=(94314,94320))
store_sns(G, f"{DB.name}-tvla")
X, Y, X_ATT, Y_ATT = prepare_traces_dl(PROF_X, PROF_Y, *TRACE_SET.attack())
Y_RAND, Y_ATT_RAND = Y.copy(), Y_ATT.copy()
np.random.shuffle(Y_RAND), np.random.shuffle(Y_ATT_RAND)
def make_dlla9_p_gradient(x9, y9, x9_att, y9_att, trials=REPEAT_DLLA):
p_gradient = []
for _ in tqdm(range(trials)):
model_9 = make_mlp(*shuffle(x9, y9), progress=False)
pg = dlla_p_gradient(model_9, *shuffle(x9_att, y9_att))
p_gradient.append(pg)
return np.mean(p_gradient, axis=0)
PG_DLLA_9 = cache_np(f"{DB.name}_dlla9_pg", make_dlla9_p_gradient, X, Y, X_ATT, Y_ATT)
PG_DLLA_9_RANDOM = cache_np(f"{DB.name}_dlla9_pg_random", make_dlla9_p_gradient, X, Y_RAND, X_ATT, Y_ATT_RAND)
plot_p_gradient({
"A vs. B": np.array(PG_DLLA_9),
"FP check": np.array(PG_DLLA_9_RANDOM),
}, f"DL-LA performance validation (9-class)\n{TITLE_SUB}",
palette=PALETTE_GRADIENT,
)
def make_wegener_p_gradient(x, y, x_att, y_att, trials=REPEAT_DLLA):
p_gradient = []
x2, y2 = reduce_fixed_fixed(x, y)
x2_att, y2_att = reduce_fixed_fixed(x_att, y_att)
for _ in tqdm(range(trials)):
model_wegener = make_mlp_wegener(*shuffle(x2, y2), progress=False)
pg = wegener_p_gradient(model_wegener, *shuffle(x2_att, y2_att))
p_gradient.append(pg)
return np.mean(p_gradient, axis=0)
PG_DLLA_2_AB = cache_np(f"{DB.name}_dlla2_pg", make_wegener_p_gradient, X, Y, X_ATT, Y_ATT)
PG_DLLA_2_RANDOM = cache_np(f"{DB.name}_dlla2_pg_random", make_wegener_p_gradient, X, Y_RAND, X_ATT, Y_ATT_RAND)
plot_p_gradient({
"A vs. B": np.array(PG_DLLA_2_AB),
"FP check": np.array(PG_DLLA_2_RANDOM),
}, f"DL-LA performance validation (Wegener)\n{TITLE_SUB}",
palette=PALETTE_GRADIENT,
)
MAX_TRACE_SIZE = max(len(PG_DLLA_9), len(PG_DLLA_2_AB), len(TVLA_PG_T_1), len(TVLA_PG_RHO))
COLOR_PALETTE = [DLLA_PALETTE[4], DLLA_PALETTE[2], TVLA_PALETTE[4], TVLA_PALETTE[2]]
def expand(arr, max_len=MAX_TRACE_SIZE):
return np.pad(arr, (0, max_len - len(arr)), 'constant', constant_values=np.nan)
PGS = {
"DL-LA 9-class": expand(PG_DLLA_9),
"DL-LA Wegener": expand(PG_DLLA_2_AB),
"TVLA $t$-test, $\mu_1$": expand(TVLA_PG_T_1),
}
if DB is Database.ascad:
PGS["TVLA $t$-test, $\mu_2$"] = expand(TVLA_PG_T_2)
else:
PGS["TVLA $\\rho$-test"] = expand(TVLA_PG_RHO)
plot_p_gradient(PGS, f"Method performance comparison\n{TITLE_SUB}", file_name=f"{DB.name}-all{FILE_SUFFIX}", palette=COLOR_PALETTE)
DF = pd.DataFrame(PGS)
DF.to_csv(f"{DB.name}-p-gradients{FILE_SUFFIX}.csv")
plot_p_gradient(PGS, f"Method performance comparison\n{TITLE_SUB}", file_name=f"{DB.name}-all{FILE_SUFFIX}-1000", palette=COLOR_PALETTE, max_traces=1000)
```
| github_jupyter |
# IPython
**IPython** (Interactive Python) is an enhanced Python shell which provides a more robust and productive development environment for users. There are several key features that set it apart from the standard Python shell.
* Interactive data analysis and visualization
* Python kernel for Jupyter notebooks
* Easy parallel computation
### History
In IPython, all your inputs and outputs are saved. There are two variables named `In` and `Out` which are assigned as you work with your results. All outputs are saved automatically to variables of the form `_N`, where `N` is the prompt number, and inputs to `_iN`. This allows you to recover quickly the result of a prior computation by referring to its number even if you forgot to store it as a variable.
```
import numpy as np
np.sin(4)**2
_1
_i1
_1 / 4.
```
### Introspection
If you want details regarding the properties and functionality of any Python objects currently loaded into IPython, you can use the `?` to reveal any details that are available:
```
some_dict = {}
some_dict?
```
If available, additional detail is provided with two question marks, including the source code of the object itself.
```
from numpy.linalg import cholesky
cholesky??
```
This syntax can also be used to search namespaces with wildcards (`*`).
```
import numpy as np
np.random.rand*?
```
### Tab completion
Because IPython allows for introspection, it is able to afford the user the ability to tab-complete commands that have been partially typed. This is done by pressing the `<tab>` key at any point during the process of typing a command:
```
np.ar
```
This can even be used to help with specifying arguments to functions, which can sometimes be difficult to remember:
```
plt.hist
```
### System commands
In IPython, you can type `ls` to see your files or `cd` to change directories, just like you would at a regular system prompt:
```
ls /Users/fonnescj/repositories/scientific-python-workshop/
```
Virtually any system command can be accessed by prepending `!`, which passes any subsequent command directly to the OS.
```
!locate python | grep pdf
```
You can even use Python variables in commands sent to the OS:
```
file_type = 'csv'
!ls ../data/*$file_type
```
The output of a system command using the exclamation point syntax can be assigned to a Python variable.
```
data_files = !ls ../data/microbiome/
data_files
```
## Qt Console
If you type at the system prompt:
$ ipython qtconsole
instead of opening in a terminal, IPython will start a graphical console that at first sight appears just like a terminal, but which is in fact much more capable than a text-only terminal. This is a specialized terminal designed for interactive scientific work, and it supports full multi-line editing with color highlighting and graphical calltips for functions, it can keep multiple IPython sessions open simultaneously in tabs, and when scripts run it can display the figures inline directly in the work area.

# Jupyter Notebook
Over time, the IPython project grew to include several components, including:
* an interactive shell
* a REPL protocol
* a notebook document fromat
* a notebook document conversion tool
* a web-based notebook authoring tool
* tools for building interactive UI (widgets)
* interactive parallel Python
As each component has evolved, several had grown to the point that they warrented projects of their own. For example, pieces like the notebook and protocol are not even specific to Python. As the result, the IPython team created Project Jupyter, which is the new home of language-agnostic projects that began as part of IPython, such as the notebook in which you are reading this text.
The HTML notebook that is part of the Jupyter project supports **interactive data visualization** and easy high-performance **parallel computing**.
```
%matplotlib inline
import matplotlib.pyplot as plt
plt.style.use('fivethirtyeight')
def f(x):
return (x-3)*(x-5)*(x-7)+85
import numpy as np
x = np.linspace(0, 10, 200)
y = f(x)
plt.plot(x,y)
```
The notebook lets you document your workflow using either HTML or Markdown.
The Jupyter Notebook consists of two related components:
* A JSON based Notebook document format for recording and distributing Python code and rich text.
* A web-based user interface for authoring and running notebook documents.
The Notebook can be used by starting the Notebook server with the command:
$ ipython notebook
This initiates an **iPython engine**, which is a Python instance that takes Python commands over a network connection.
The **IPython controller** provides an interface for working with a set of engines, to which one or more **iPython clients** can connect.
The Notebook gives you everything that a browser gives you. For example, you can embed images, videos, or entire websites.
```
from IPython.display import IFrame
IFrame('https://jupyter.org', width='100%', height=350)
from IPython.display import YouTubeVideo
YouTubeVideo("rl5DaFbLc60")
```
## Markdown cells
Markdown is a simple *markup* language that allows plain text to be converted into HTML.
The advantages of using Markdown over HTML (and LaTeX):
- its a **human-readable** format
- allows writers to focus on content rather than formatting and layout
- easier to learn and use
For example, instead of writing:
```html
<p>In order to create valid
<a href="http://en.wikipedia.org/wiki/HTML">HTML</a>, you
need properly coded syntax that can be cumbersome for
“non-programmers” to write. Sometimes, you
just want to easily make certain words <strong>bold
</strong>, and certain words <em>italicized</em> without
having to remember the syntax. Additionally, for example,
creating lists:</p>
<ul>
<li>should be easy</li>
<li>should not involve programming</li>
</ul>
```
we can write the following in Markdown:
```markdown
In order to create valid [HTML], you need properly
coded syntax that can be cumbersome for
"non-programmers" to write. Sometimes, you just want
to easily make certain words **bold**, and certain
words *italicized* without having to remember the
syntax. Additionally, for example, creating lists:
* should be easy
* should not involve programming
```
### Emphasis
Markdown uses `*` (asterisk) and `_` (underscore) characters as
indicators of emphasis.
*italic*, _italic_
**bold**, __bold__
***bold-italic***, ___bold-italic___
*italic*, _italic_
**bold**, __bold__
***bold-italic***, ___bold-italic___
### Lists
Markdown supports both unordered and ordered lists. Unordered lists can use `*`, `-`, or
`+` to define a list. This is an unordered list:
* Apples
* Bananas
* Oranges
* Apples
* Bananas
* Oranges
Ordered lists are numbered lists in plain text:
1. Bryan Ferry
2. Brian Eno
3. Andy Mackay
4. Paul Thompson
5. Phil Manzanera
1. Bryan Ferry
2. Brian Eno
3. Andy Mackay
4. Paul Thompson
5. Phil Manzanera
### Links
Markdown inline links are equivalent to HTML `<a href='foo.com'>`
links, they just have a different syntax.
[Biostatistics home page](http://biostat.mc.vanderbilt.edu "Visit Biostat!")
[Biostatistics home page](http://biostat.mc.vanderbilt.edu "Visit Biostat!")
### Block quotes
Block quotes are denoted by a `>` (greater than) character
before each line of the block quote.
> Sometimes a simple model will outperform a more complex model . . .
> Nevertheless, I believe that deliberately limiting the complexity
> of the model is not fruitful when the problem is evidently complex.
> Sometimes a simple model will outperform a more complex model . . .
> Nevertheless, I believe that deliberately limiting the complexity
> of the model is not fruitful when the problem is evidently complex.
### Images
Images look an awful lot like Markdown links, they just have an extra
`!` (exclamation mark) in front of them.


### Remote Code
Use `%load` to add remote code
```
# %load http://matplotlib.org/mpl_examples/shapes_and_collections/scatter_demo.py
"""
Simple demo of a scatter plot.
"""
import numpy as np
import matplotlib.pyplot as plt
N = 50
x = np.random.rand(N)
y = np.random.rand(N)
colors = np.random.rand(N)
area = np.pi * (15 * np.random.rand(N))**2 # 0 to 15 point radiuses
plt.scatter(x, y, s=area, c=colors, alpha=0.5)
plt.show()
```
### Mathjax Support
Mathjax ia a javascript implementation $\alpha$ of LaTeX that allows equations to be embedded into HTML. For example, this markup:
"""$$ \int_{a}^{b} f(x)\, dx \approx \frac{1}{2} \sum_{k=1}^{N} \left( x_{k} - x_{k-1} \right) \left( f(x_{k}) + f(x_{k-1}) \right). $$"""
becomes this:
$$
\int_{a}^{b} f(x)\, dx \approx \frac{1}{2} \sum_{k=1}^{N} \left( x_{k} - x_{k-1} \right) \left( f(x_{k}) + f(x_{k-1}) \right).
$$
## SymPy Support
SymPy is a Python library for symbolic mathematics. It supports:
* polynomials
* calculus
* solving equations
* discrete math
* matrices
```
from sympy import *
init_printing()
x, y = symbols("x y")
eq = ((x+y)**2 * (x+1))
eq
expand(eq)
(1/cos(x)).series(x, 0, 6)
limit((sin(x)-x)/x**3, x, 0)
diff(cos(x**2)**2 / (1+x), x)
```
### Magic functions
IPython has a set of predefined ‘magic functions’ that you can call with a command line style syntax. These include:
* `%run`
* `%edit`
* `%debug`
* `%timeit`
* `%paste`
* `%load_ext`
```
%lsmagic
```
Timing the execution of code; the `timeit` magic exists both in line and cell form:
```
%timeit np.linalg.eigvals(np.random.rand(100,100))
%%timeit a = np.random.rand(100, 100)
np.linalg.eigvals(a)
```
IPython also creates aliases for a few common interpreters, such as bash, ruby, perl, etc.
These are all equivalent to `%%script <name>`
```
%%ruby
puts "Hello from Ruby #{RUBY_VERSION}"
%%bash
echo "hello from $BASH"
```
IPython has an `rmagic` extension that contains a some magic functions for working with R via rpy2. This extension can be loaded using the `%load_ext` magic as follows:
```
%load_ext rpy2.ipython
```
If the above generates an error, it is likely that you do not have the `rpy2` module installed. You can install this now via:
```
!pip install rpy2
```
or, if you are running Anaconda, via `conda`:
```
!conda install rpy2
x,y = np.arange(10), np.random.normal(size=10)
%R print(lm(rnorm(10)~rnorm(10)))
%%R -i x,y -o XYcoef
lm.fit <- lm(y~x)
par(mfrow=c(2,2))
print(summary(lm.fit))
plot(lm.fit)
XYcoef <- coef(lm.fit)
XYcoef
```
## Debugging
The `%debug` magic can be used to trigger the IPython debugger (`ipd`) for a cell that raises an exception. The debugger allows you to step through code line-by-line and inspect variables and execute code.
```
def div(x, y):
return x/y
div(1,0)
%debug
```
## Exporting and Converting Notebooks
In Jupyter, one can convert an `.ipynb` notebook document file into various static formats via the `nbconvert` tool. Currently, nbconvert is a command line tool, run as a script using Jupyter.
```
!jupyter nbconvert --to html "IPython and Jupyter.ipynb"
```
Currently, `nbconvert` supports HTML (default), LaTeX, Markdown, reStructuredText, Python and HTML5 slides for presentations. Some types can be post-processed, such as LaTeX to PDF (this requires [Pandoc](http://johnmacfarlane.net/pandoc/) to be installed, however).
```
!jupyter nbconvert --to pdf "Introduction to pandas.ipynb"
```
A very useful online service is the [IPython Notebook Viewer](http://nbviewer.ipython.org) which allows you to display your notebook as a static HTML page, which is useful for sharing with others:
```
IFrame("http://nbviewer.ipython.org/2352771", width='100%', height=350)
```
As of this year, GitHub supports the [rendering of Jupyter Notebooks](https://github.com/fonnesbeck/Bios8366/blob/master/notebooks/Section1_2-Programming-with-Python.ipynb) stored on its repositories.
## Reproducible Research
> reproducing conclusions from a single experiment based on the measurements from that experiment
The most basic form of reproducibility is a complete description of the data and associated analyses (including code!) so the results can be *exactly* reproduced by others.
Reproducing calculations can be onerous, even with one's own work!
Scientific data are becoming larger and more complex, making simple descriptions inadequate for reproducibility. As a result, most modern research is irreproducible without tremendous effort.
*** Reproducible research is not yet part of the culture of science in general, or scientific computing in particular. ***
## Scientific Computing Workflow
There are a number of steps to scientific endeavors that involve computing:

Many of the standard tools impose barriers between one or more of these steps. This can make it difficult to iterate, reproduce work.
The Jupyter notebook eliminates or reduces these barriers to reproducibility.
## Parallel IPython
The IPython architecture consists of four components, which reside in the `ipyparallel` package:
1. **Engine** The IPython engine is a Python instance that accepts Python commands over a network connection. When multiple engines are started, parallel and distributed computing becomes possible. An important property of an IPython engine is that it blocks while user code is being executed.
2. **Hub** The hub keeps track of engine connections, schedulers, clients, as well as persist all task requests and results in a database for later use.
3. **Schedulers** All actions that can be performed on the engine go through a Scheduler. While the engines themselves block when user code is run, the schedulers hide that from the user to provide a fully asynchronous interface to a set of engines.
4. **Client** The primary object for connecting to a cluster.

(courtesy Min Ragan-Kelley)
This architecture is implemented using the ØMQ messaging library and the associated Python bindings in `pyzmq`.
### Running parallel IPython
To enable the IPython Clusters tab in Jupyter Notebook:
ipcluster nbextension enable
When you then start a Jupyter session, you should see the following in your **IPython Clusters** tab:

Before running the next cell, make sure you have first started your cluster, you can use the [clusters tab in the dashboard](/#tab2) to do so.
Select the number if IPython engines (nodes) that you want to use, then click **Start**.
```
from ipyparallel import Client
client = Client()
dv = client.direct_view()
len(dv)
def where_am_i():
import os
import socket
return "In process with pid {0} on host: '{1}'".format(
os.getpid(), socket.gethostname())
where_am_i_direct_results = dv.apply(where_am_i)
where_am_i_direct_results.get()
```
Let's now consider a useful function that we might want to run in parallel. Here is a version of the approximate Bayesian computing (ABC) algorithm.
```
import numpy
def abc(y, N, epsilon=[0.2, 0.8]):
trace = []
while len(trace) < N:
# Simulate from priors
mu = numpy.random.normal(0, 10)
sigma = numpy.random.uniform(0, 20)
x = numpy.random.normal(mu, sigma, 50)
#if (np.linalg.norm(y - x) < epsilon):
if ((abs(x.mean() - y.mean()) < epsilon[0]) &
(abs(x.std() - y.std()) < epsilon[1])):
trace.append([mu, sigma])
return trace
y = numpy.random.normal(4, 2, 50)
```
Let's try running this on one of the cluster engines:
```
dv0 = client[0]
dv0.block = True
dv0.apply(abc, y, 10)
```
This fails with a NameError because NumPy has not been imported on the engine to which we sent the task. Each engine has its own namespace, so we need to import whatever modules we will need prior to running our code:
```
dv0.execute("import numpy")
dv0.apply(abc, y, 10)
```
An easier approach is to use the parallel cell magic to import everywhere:
```
%%px
import numpy
```
This magic can be used to execute the same code on all nodes.
```
%%px
import os
print(os.getpid())
%%px
%matplotlib inline
import matplotlib.pyplot as plt
import os
tsamples = numpy.random.randn(100)
plt.hist(tsamples)
_ = plt.title('PID %i' % os.getpid())
```
## Links and References
* [IPython Notebook Viewer](http://nbviewer.ipython.org) Displays static HTML versions of notebooks, and includes a gallery of notebook examples.
* [A Reference-Free Algorithm for Computational Normalization of Shotgun Sequencing Data](http://ged.msu.edu/papers/2012-diginorm/) A landmark example of reproducible research in genomics: Git repo, iPython notebook, data and scripts.
* Jacques Ravel and K Eric Wommack. 2014. [All Hail Reproducibility in Microbiome Research](http://www.microbiomejournal.com/content/pdf/2049-2618-2-8.pdf). Microbiome, 2:8.
* Benjamin Ragan-Kelley et al.. 2013. [Collaborative cloud-enabled tools allow rapid, reproducible biological insights](http://www.nature.com/ismej/journal/v7/n3/full/ismej2012123a.html). The ISME Journal, 7, 461–464; doi:10.1038/ismej.2012.123;
| github_jupyter |
```
import pickle
import requests
import json
```
## Call Expected timefraction
```
url = 'https://165.227.232.84:8000/predict/'
lower_bound = 1.1
upper_bound = 1.3
time_horizon = 100
inp = {
"prediction_type": "Expected_timefraction",
"lower_bound": lower_bound,
"upper_bound": upper_bound,
"time_fraction": 0.5,
"time_horizon": time_horizon
}
res = requests.post(url, headers = {'Content-type': 'application/json'}, json=inp)
#print(res)
print(json.loads(res.json()))
```
## Call Best range
```
lower_bound = 1.1
upper_bound = 1.3
time_horizon = 100
inp = {
"prediction_type": "best_range",
"lower_bound": lower_bound,
"upper_bound": upper_bound,
"time_fraction": 0.5,
"time_horizon": time_horizon
}
res = requests.post(url, headers = {'Content-type': 'application/json'}, json=inp)
#print(res)
print(json.loads(res.json()))
### Call Best range for timefraction 0,5 and 0.25 and plot some visuals
from src.models.import_ethereum_timeseries import ethereum_data
from matplotlib import pylab as plt
import matplotlib
f, ax = plt.subplots(1,1,figsize=(40,20))
time, eth_value = ethereum_data()
today = time[-1]
dt = time[-1] - time[-2]
inp["time_fraction"] = 0.5
inp['time_horizon'] = 30
res = requests.post(url, headers = {'Content-type': 'application/json'}, json=inp)
lower_range50, upper_range50 = json.loads(res.json())['Best range']
inp["time_fraction"] = 0.25
res = requests.post(url, headers = {'Content-type': 'application/json'}, json=inp)
lower_range25, upper_range25 = json.loads(res.json())['Best range']
font = {'family' : 'normal',
'weight' : 'normal',
'size' : 30}
matplotlib.rc('font', **font)
ax.plot(time, eth_value)
ax.plot([today, today + dt*time_horizon], [eth_value[-1], eth_value[-1]],
c='black', ls='--')
ax.plot([today, today + dt*time_horizon], [eth_value[-1]*lower_range50, eth_value[-1]*lower_range50],
c='orange', lw='3', label='50%')
ax.plot([today, today + dt*time_horizon], [eth_value[-1]*upper_range50, eth_value[-1]*upper_range50],
c='orange', lw='3', label='50%')
ax.plot([today, today + dt*time_horizon], [eth_value[-1]*lower_range25, eth_value[-1]*lower_range25],
c='green', lw='3', label='25%')
ax.plot([today, today + dt*time_horizon], [eth_value[-1]*upper_range25, eth_value[-1]*upper_range25],
c='green', lw='3', label='25%')
ax.legend()
ax.set_title("Time horizon " + str(dt*inp['time_horizon']) + "days",fontsize=30)
import datetime
import numpy as np
historic_point = np.argmin(abs(time - datetime.datetime(2020,12,23)))
cut_point = np.argmin(abs(time - datetime.datetime(2020,12,23)))
#metadata = {"cut_date": eth_value[cut_point]}
eth_value = eth_value[:cut_point+1]
f, ax = plt.subplots(1,1,figsize=(12,12))
time, eth_value = ethereum_data()
cut_point = np.argmin(abs(time - datetime.datetime(2020,12,23)))
#metadata = {"cut_date": eth_value[cut_point]}
eth_value = eth_value[:cut_point+1]
today = time[-1]
ax.plot(eth_value)
ax.plot()
inp = {}
inp['historic_point'] = 100
if hasattr(inp, "historic_point"):
print(inp["historic_point"])
```
| github_jupyter |
<a href="https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_13_04_retrain.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# T81-558: Applications of Deep Neural Networks
**Module 13: Advanced/Other Topics**
* Instructor: [Jeff Heaton](https://sites.wustl.edu/jeffheaton/), McKelvey School of Engineering, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
* For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
# Module 13 Video Material
* Part 13.1: Flask and Deep Learning Web Services [[Video]](https://www.youtube.com/watch?v=H73m9XvKHug&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_13_01_flask.ipynb)
* Part 13.2: Interrupting and Continuing Training [[Video]](https://www.youtube.com/watch?v=kaQCdv46OBA&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_13_02_checkpoint.ipynb)
* Part 13.3: Using a Keras Deep Neural Network with a Web Application [[Video]](https://www.youtube.com/watch?v=OBbw0e-UroI&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_13_03_web.ipynb)
* **Part 13.4: When to Retrain Your Neural Network** [[Video]](https://www.youtube.com/watch?v=K2Tjdx_1v9g&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_13_04_retrain.ipynb)
* Part 13.5: Tensor Processing Units (TPUs) [[Video]](https://www.youtube.com/watch?v=Ygyf3NUqvSc&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_13_05_tpu.ipynb)
# Part 13.4: When to Retrain Your Neural Network
Dataset drift is a problem frequently seen in real-world applications of machine learning. Academic problems that courses typically present in school assignments usually do not experience this problem. For a class assignment, your instructor provides a single data set representing all of the data you will ever see for a task. In the real world, you obtain initial data to train your model; then, you will acquire new data over time that you use your model to predict.
Consider this example. You create a startup company that develops a mobile application that helps people find jobs. To train your machine learning model, you collect attributes about people and their careers. Once you have your data, you can prepare your neural network to suggest the best jobs for individuals.
Once your application is released, you will hopefully obtain new data. This data will come from job seekers using your app. These people are your customers. You have x values (their attributes), but you do not have y-values (their jobs). Your customers have come to you to find out what their be jobs will be. You will provide the customer's attributes to the neural network, and then it will predict their jobs. Usually, companies develop neural networks on initial data than use the neural network to perform predictions on new data obtained over time from their customers.
However, as time passes, companies must look if their model is still relevant. Your job prediction model will become less relevant as industry introduces new job types and the demographics of your customers change. This change in your underlying data is called dataset drift. In this section, we will see ways that you can measure dataset drift.
You can present your model with new data and see how its accuracy changes over time. However, to calculate efficiency, you must know the expected outputs from the model (y-values). For new data that you are obtaining in real-time, you may not know the correct outcomes. Therefore, we will look at algorithms that examine the x-inputs and determine how much they have changed in distribution from the original x-inputs that we trained on. These changes are called dataset drift.
Let's begin by creating generated data that illustrates drift. We present the following code to create a chart that shows such drift.
```
import numpy as np
import matplotlib.pyplot as plot
from sklearn.linear_model import LinearRegression
def true_function(x):
x2 = (x*8) - 1
return ((np.sin(x2)/x2)*0.6)+0.3
#
x_train = np.arange(0, 0.6, 0.01)
x_test = np.arange(0.6, 1.1, 0.01)
x_true = np.concatenate( (x_train, x_test) )
#
y_true_train = true_function(x_train)
y_true_test = true_function(x_test)
y_true = np.concatenate( (y_true_train, y_true_test) )
#
y_train = y_true_train + (np.random.rand(*x_train.shape)-0.5)*0.4
y_test = y_true_test + (np.random.rand(*x_test.shape)-0.5)*0.4
#
lr_x_train = x_train.reshape((x_train.shape[0],1))
reg = LinearRegression().fit(lr_x_train, y_train)
reg_pred = reg.predict(lr_x_train)
print(reg.coef_[0])
print(reg.intercept_)
#
plot.xlim([0,1.5])
plot.ylim([0,1])
l1 = plot.scatter(x_train, y_train, c="g", label="Training Data")
l2 = plot.scatter(x_test, y_test, c="r", label="Testing Data")
l3, = plot.plot(lr_x_train, reg_pred, color='black', linewidth=3,
label="Trained Model")
l4, = plot.plot(x_true, y_true, label = "True Function")
plot.legend(handles=[l1, l2, l3, l4])
#
plot.title('Drift')
plot.xlabel('Time')
plot.ylabel('Sales')
plot.grid(True, which='both')
plot.show()
```
The true-function represents what the data does over time. Unfortunately, you only have the training portion of the data. Your model will do quite well on the data that you trained it trained with; however, it will be very inaccurate on the new test data presented to it. The prediction line for the model fits the training data well but does not fit the est data well.
### Preprocessing the Sberbank Russian Housing Market Data
The examples provided in this section use a Kaggle dataset named The Sberbank Russian Housing Market, which can be found at the following link.
* [Sberbank Russian Housing Market](https://www.kaggle.com/c/sberbank-russian-housing-market/data)
Kaggle datasets are already broken into training and test. We must load both of these files.
```
import os
import numpy as np
import pandas as pd
from sklearn.preprocessing import LabelEncoder
PATH = "/Users/jheaton/Downloads/sberbank-russian-housing-market"
train_df = pd.read_csv(os.path.join(PATH,"train.csv"))
test_df = pd.read_csv(os.path.join(PATH,"test.csv"))
```
I provide a simple preprocess function that converts all numerics to z-scores and all categoricals to dummies.
```
def preprocess(df):
for i in df.columns:
if df[i].dtype == 'object':
df[i] = df[i].fillna(df[i].mode().iloc[0])
elif (df[i].dtype == 'int' or df[i].dtype == 'float'):
df[i] = df[i].fillna(np.nanmedian(df[i]))
enc = LabelEncoder()
for i in df.columns:
if (df[i].dtype == 'object'):
df[i] = enc.fit_transform(df[i].astype('str'))
df[i] = df[i].astype('object')
```
Next, we run both the training and test datasets througn the preprocessing function.
```
preprocess(train_df)
preprocess(test_df)
```
Finally, we remove thr target variable. We are only looking for drift on the x (input data).
```
train_df.drop('price_doc',axis=1,inplace=True)
```
### KS-Statistic
We will use the KS-Statistic to determine the difference in distribution between columns in the training and test sets. Just as a baseline, consider if we compare the same field to itself. I this case, we are comparing the **kitch_sq** in the training set. Because there is no difference in distribution between a field in itself, the p-value is 1.0, and the KS-Statistic statistic is 0. The P-Value is the probability that there is no difference between the two distributions. Typically some lower threshold is used for how low a P-Value is needed to reject the null hypothesis and assume there is a difference. The value of 0.05 is a standard threshold for p-values. Because the p-value is NOT below 0.05, we can expect the two distributions are the same. If the p-value were below the threshold, then the **statistic** value becomes interesting. This value tells you how different the two distributions are. A value of 0.0, in this case, means no differences.
```
from scipy import stats
stats.ks_2samp(train_df['kitch_sq'], train_df['kitch_sq'])
```
Now let's do something more interesting. We will compare the same field **kitch_sq** between the test and training sets. In this case, the p-value is below 0.05, so the **statistic** value now contains the amount of difference detected.
```
stats.ks_2samp(train_df['kitch_sq'], test_df['kitch_sq'])
```
Next, we pull the KS-Stat for every field. We also establish a boundary for the maximum p-value to display and how much of a difference is needed before we display the column.
```
for col in train_df.columns:
ks = stats.ks_2samp(train_df[col], test_df[col])
if ks.pvalue < 0.05 and ks.statistic>0.1:
print(f'{col}: {ks}')
```
### Detecting Drift between Training and Testing Datasets by Training
Sample the training and test into smaller sets to train. We want 10K elements from each; however, the test set only has 7,662, so we only sample that amount from each side.
```
SAMPLE_SIZE = min(len(train_df),len(test_df))
SAMPLE_SIZE = min(SAMPLE_SIZE,10000)
print(SAMPLE_SIZE)
```
We take the random samples from the training and test sets and add a flag called **source_training** to tell the two apart.
```
training_sample = train_df.sample(SAMPLE_SIZE, random_state=49)
testing_sample = test_df.sample(SAMPLE_SIZE, random_state=48)
# Is the data from the training set?
training_sample['source_training'] = 1
testing_sample['source_training'] = 0
```
Next, we combine the data that we sampled from the training and test data sets and shuffle them.
```
# Build combined training set
combined = testing_sample.append(training_sample)
combined.reset_index(inplace=True, drop=True)
# Now randomize
combined = combined.reindex(np.random.permutation(combined.index))
combined.reset_index(inplace=True, drop=True)
```
We will now generate $x$ and $y$ to train. We are attempting to predict the **source_training** value as $y$, which indicates if the data came from the training or test set. If the model is very successful at using the data to predict if it came from training or testing, then there is likely drift. Ideally, the train and test data should be indistinguishable.
```
# Get ready to train
y = combined['source_training'].values
combined.drop('source_training',axis=1,inplace=True)
x = combined.values
y
```
We will consider anything above a 0.75 AUC as having a good chance of drift.
```
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import cross_val_score
model = RandomForestClassifier(n_estimators = 60, max_depth = 7,
min_samples_leaf = 5)
lst = []
for i in combined.columns:
score = cross_val_score(model,pd.DataFrame(combined[i]),y,cv=2,
scoring='roc_auc')
if (np.mean(score) > 0.75):
lst.append(i)
print(i,np.mean(score))
```
| github_jupyter |
<center>
<img src="../../img/ods_stickers.jpg" />
## [mlcourse.ai](mlcourse.ai) – Open Machine Learning Course
Authors: Yury Isakov, [Yury Kashnitskiy](https://yorko.github.io) (@yorko). Edited by Anna Tarelina (@feuerengel), and Kolchenko Sergey (@KolchenkoSergey). This material is subject to the terms and conditions of the [Creative Commons CC BY-NC-SA 4.0](https://creativecommons.org/licenses/by-nc-sa/4.0/) license. Free use is permitted for any non-commercial purpose.
# <center> Assignment #4
## <center> User Identification with Logistic Regression (beating baselines in the "Alice" competition)
Today we are going to practice working with sparse matrices, training Logistic Regression models, and doing feature engineering. We will reproduce a couple of baselines in the ["Catch Me If You Can: Intruder Detection through Webpage Session Tracking"](https://www.kaggle.com/c/catch-me-if-you-can-intruder-detection-through-webpage-session-tracking2) (a.k.a. "Alice") Kaggle inclass competition. More credits will be given for beating a stronger baseline.
**Your task:**
1. "Follow me". Complete the missing code and submit your answers via [the google-form](https://docs.google.com/forms/d/1V4lHXkjZvpDDvHAcnH6RuEQJecBaLo8zooxDl1_aP60). 14 credit max. for this part
2. "Freeride". Come up with good features to beat the baseline "A4 baseline 3". You need to name your [team](https://www.kaggle.com/c/catch-me-if-you-can-intruder-detection-through-webpage-session-tracking2/team) (out of 1 person) in full accordance with the course rating. You can think of it as a part of the assignment. 10 more credits for beating the mentioned baseline and correct team naming.
# Part 1. Follow me
<img src='../../img/followme_alice.png' width=50%>
*image credit [@muradosmann](https://www.instagram.com/muradosmann/?hl=en)*
```
# Import libraries and set desired options
import pickle
import numpy as np
import pandas as pd
from scipy.sparse import csr_matrix, hstack
from sklearn.preprocessing import StandardScaler
from sklearn.metrics import roc_auc_score
from sklearn.linear_model import LogisticRegression
from matplotlib import pyplot as plt
import seaborn as sns
sns.set()
```
##### Problem description
In this competition, we''ll analyze the sequence of websites consequently visited by a particular person and try to predict whether this person is Alice or someone else. As a metric we will use [ROC AUC](https://en.wikipedia.org/wiki/Receiver_operating_characteristic).
### 1. Data Downloading and Transformation
Register on [Kaggle](www.kaggle.com), if you have not done it before.
Go to the competition [page](https://inclass.kaggle.com/c/catch-me-if-you-can-intruder-detection-through-webpage-session-tracking2) and download the data.
First, read the training and test sets. Then we'll explore the data in hand and do a couple of simple exercises.
```
# Read the training and test data sets, change paths if needed
train_df = pd.read_csv('../../data/train_sessions.csv',
index_col='session_id')
test_df = pd.read_csv('../../data/test_sessions.csv',
index_col='session_id')
# Convert time1, ..., time10 columns to datetime type
times = ['time%s' % i for i in range(1, 11)]
train_df[times] = train_df[times].apply(pd.to_datetime)
test_df[times] = test_df[times].apply(pd.to_datetime)
# Sort the data by time
train_df = train_df.sort_values(by='time1')
# Look at the first rows of the training set
train_df.head()
```
The training data set contains the following features:
- **site1** – id of the first visited website in the session
- **time1** – visiting time for the first website in the session
- ...
- **site10** – id of the tenth visited website in the session
- **time10** – visiting time for the tenth website in the session
- **target** – target variable, 1 for Alice's sessions, and 0 for the other users' sessions
User sessions are chosen in the way that they are shorter than 30 min. long and contain no more than 10 websites. I.e. a session is considered over either if a user has visited 10 websites or if a session has lasted over 30 minutes.
There are some empty values in the table, it means that some sessions contain less than ten websites. Replace empty values with 0 and change columns types to integer. Also load the websites dictionary and check how it looks like:
```
# Change site1, ..., site10 columns type to integer and fill NA-values with zeros
sites = ['site%s' % i for i in range(1, 11)]
train_df[sites] = train_df[sites].fillna(0).astype(np.uint16)
test_df[sites] = test_df[sites].fillna(0).astype(np.uint16)
# Load websites dictionary
with open(r"../../data/site_dic.pkl", "rb") as input_file:
site_dict = pickle.load(input_file)
# Create dataframe for the dictionary
sites_dict = pd.DataFrame(list(site_dict.keys()), index=list(site_dict.values()), columns=['site'])
print(u'Websites total:', sites_dict.shape[0])
sites_dict.head()
```
#### 4.1. What are the dimensions of the training and test sets (in exactly this order)?
*For discussions, please stick to [ODS Slack](https://opendatascience.slack.com/), channel #mlcourse_ai, pinned thread __#a4_q1__*
- (82797, 20) and (253561, 20)
- (82797, 20) and (253561, 21)
- (253561, 21) and (82797, 20)
- (253561, 20) and (82797, 20)
```
# Your code is here
```
### 2. Brief Exploratory Data Analysis
Before we start training models, we have to perform Exploratory Data Analysis ([EDA](https://en.wikipedia.org/wiki/Exploratory_data_analysis)). Today, we are going to perform a shorter version, but we will use other techniques as we move forward. Let's check which websites in the training data set are the most visited. As you can see, they are Google services and a bioinformatics website (a website with 'zero'-index is our missed values, just ignore it):
```
# Top websites in the training data set
top_sites = pd.Series(train_df[sites].values.flatten()
).value_counts().sort_values(ascending=False).head(5)
print(top_sites)
sites_dict.loc[top_sites.drop(0).index]
```
##### 4.2. What kind of websites does Alice visit the most?
*For discussions, please stick to [ODS Slack](https://opendatascience.slack.com/), channel #mlcourse_ai, pinned thread __#a4_q2__*
- videohostings
- social networks
- torrent trackers
- news
```
# Your code is here
```
Now let us look at the timestamps and try to characterize sessions as timeframes:
```
# Create a separate dataframe where we will work with timestamps
time_df = pd.DataFrame(index=train_df.index)
time_df['target'] = train_df['target']
# Find sessions' starting and ending
time_df['min'] = train_df[times].min(axis=1)
time_df['max'] = train_df[times].max(axis=1)
# Calculate sessions' duration in seconds
time_df['seconds'] = (time_df['max'] - time_df['min']) / np.timedelta64(1, 's')
time_df.head()
```
In order to perform the next task, generate descriptive statistics as you did in the first assignment.
##### 4.3. Select all correct statements:
*For discussions, please stick to [ODS Slack](https://opendatascience.slack.com/), channel #mlcourse_ai, pinned thread __#a4_q3__*
- on average, Alice's session is shorter than that of other users
- more than 1% of all sessions in the dataset belong to Alice
- minimum and maximum durations of Alice's and other users' sessions are approximately the same
- variation about the mean session duration for all users (including Alice) is approximately the same
- less than a quarter of Alice's sessions are greater than or equal to 40 seconds
```
# Your code is here
```
In order to train our first model, we need to prepare the data. First of all, exclude the target variable from the training set. Now both training and test sets have the same number of columns, therefore aggregate them into one dataframe. Thus, all transformations will be performed simultaneously on both training and test data sets.
On the one hand, it leads to the fact that both data sets have one feature space (you don't have to worry that you forgot to transform a feature in some data sets). On the other hand, processing time will increase.
For the enormously large sets it might turn out that it is impossible to transform both data sets simultaneously (and sometimes you have to split your transformations into several stages only for train/test data set).
In our case, with this particular data set, we are going to perform all the transformations for the whole united dataframe at once, and before training the model or making predictions we will just take its appropriate part.
```
# Our target variable
y_train = train_df['target']
# United dataframe of the initial data
full_df = pd.concat([train_df.drop('target', axis=1), test_df])
# Index to split the training and test data sets
idx_split = train_df.shape[0]
```
For the very basic model, we will use only the visited websites in the session (but we will not take into account timestamp features). The point behind this data selection is: *Alice has her favorite sites, and the more often you see these sites in the session, the higher probability that this is Alice's session, and vice versa.*
Let us prepare the data, we will take only features `site1, site2, ... , site10` from the whole dataframe. Keep in mind that the missing values are replaced with zero. Here is how the first rows of the dataframe look like:
```
# Dataframe with indices of visited websites in session
full_sites = full_df[sites]
full_sites.head()
```
Sessions are sequences of website indices, and data in this representation is useless for machine learning method (just think, what happens if we switched all ids of all websites).
According to our hypothesis (Alice has favorite websites), we need to transform this dataframe so each website has a corresponding feature (column) and its value is equal to number of this website visits in the session. It can be done in two lines:
```
# sequence of indices
sites_flatten = full_sites.values.flatten()
# and the matrix we are looking for
# (make sure you understand which of the `csr_matrix` constructors is used here)
# a further toy example will help you with it
full_sites_sparse = csr_matrix(([1] * sites_flatten.shape[0],
sites_flatten,
range(0, sites_flatten.shape[0] + 10, 10)))[:, 1:]
full_sites_sparse.shape
```
If you understand what just happened here, then you can skip the next passage (perhaps, you can handle logistic regression too?), If not, then let us figure it out.
### Important detour #1: Sparse Matrices
Let us estimate how much memory it will require to store our data in the example above. Our united dataframe contains 336 thousand samples of 48 thousand integer features in each. It's easy to calculate the required amount of memory, roughly:
$$336K * 48K * 8 bytes = 16M * 8 bytes = 128 GB,$$
(that's the [exact](http://www.wolframalpha.com/input/?i=336358*48371*8+bytes) value). Obviously, ordinary mortals have no such volumes (strictly speaking, Python may allow you to create such a matrix, but it will not be easy to do anything with it). The interesting fact is that most of the elements of our matrix are zeros. If we count non-zero elements, then it will be about 1.8 million, i.е. slightly more than 10% of all matrix elements. Such a matrix, where most elements are zeros, is called sparse, and the ratio between the number of zero elements and the total number of elements is called the sparseness of the matrix.
For the work with such matrices you can use `scipy.sparse` library, check [documentation](https://docs.scipy.org/doc/scipy-0.18.1/reference/sparse.html) to understand what possible types of sparse matrices are, how to work with them and in which cases their usage is most effective. You can learn how they are arranged, for example, in Wikipedia [article](https://en.wikipedia.org/wiki/Sparse_matrix).
Note, that a sparse matrix contains only non-zero elements, and you can get the allocated memory size like this (significant memory savings are obvious):
```
# How much memory does a sparse matrix occupy?
print('{0} elements * {1} bytes = {2} bytes'.format(full_sites_sparse.count_nonzero(), 8,
full_sites_sparse.count_nonzero() * 8))
# Or just like this:
print('sparse_matrix_size = {0} bytes'.format(full_sites_sparse.data.nbytes))
```
Let us explore how the matrix with the websites has been formed using a mini example. Suppose we have the following table with user sessions:
| id | site1 | site2 | site3 |
|---|---|---|---|
| 1 | 1 | 0 | 0 |
| 2 | 1 | 3 | 1 |
| 3 | 2 | 3 | 4 |
There are 3 sessions, and no more than 3 websites in each. Users visited four different sites in total (there are numbers from 1 to 4 in the table cells). And let us assume that the mapping is:
1. vk.com
2. habrahabr.ru
3. yandex.ru
4. ods.ai
If the user has visited less than 3 websites during the session, the last few values will be zero. We want to convert the original dataframe in a way that each session has a corresponding row which shows the number of visits to each particular site. I.e. we want to transform the previous table into the following form:
| id | vk.com | habrahabr.ru | yandex.ru | ods.ai |
|---|---|---|---|---|
| 1 | 1 | 0 | 0 | 0 |
| 2 | 2 | 0 | 1 | 0 |
| 3 | 0 | 1 | 1 | 1 |
To do this, use the constructor: `csr_matrix ((data, indices, indptr))` and create a frequency table (see examples, code and comments on the links above to see how it works). Here we set all the parameters explicitly for greater clarity:
```
# data, create the list of ones, length of which equal to the number of elements in the initial dataframe (9)
# By summing the number of ones in the cell, we get the frequency,
# number of visits to a particular site per session
data = [1] * 9
# To do this, you need to correctly distribute the ones in cells
# Indices - website ids, i.e. columns of a new matrix. We will sum ones up grouping them by sessions (ids)
indices = [1, 0, 0, 1, 3, 1, 2, 3, 4]
# Indices for the division into rows (sessions)
# For example, line 0 is the elements between the indices [0; 3) - the rightmost value is not included
# Line 1 is the elements between the indices [3; 6)
# Line 2 is the elements between the indices [6; 9)
indptr = [0, 3, 6, 9]
# Aggregate these three variables into a tuple and compose a matrix
# To display this matrix on the screen transform it into the usual "dense" matrix
csr_matrix((data, indices, indptr)).todense()
```
As you might have noticed, there are not four columns in the resulting matrix (corresponding to number of different websites) but five. A zero column has been added, which indicates if the session was shorter (in our mini example we took sessions of three). This column is excessive and should be removed from the dataframe (do that yourself).
##### 4.4. What is the sparseness of the matrix in our small example?
*For discussions, please stick to [ODS Slack](https://opendatascience.slack.com/), channel #mlcourse_ai, pinned thread __#a4_q4__*
- 42%
- 47%
- 50%
- 53%
```
# Your code is here
```
Another benefit of using sparse matrices is that there are special implementations of both matrix operations and machine learning algorithms for them, which sometimes allows to significantly accelerate operations due to the data structure peculiarities. This applies to logistic regression as well. Now everything is ready to build our first model.
### 3. Training the first model
So, we have an algorithm and data for it. Let us build our first model, using [logistic regression](http://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LogisticRegression.html) implementation from ` Sklearn` with default parameters. We will use the first 90% of the data for training (the training data set is sorted by time), and the remaining 10% for validation. Let's write a simple function that returns the quality of the model and then train our first classifier:
```
def get_auc_lr_valid(X, y, C=1.0, seed=17, ratio = 0.9):
# Split the data into the training and validation sets
idx = int(round(X.shape[0] * ratio))
# Classifier training
lr = LogisticRegression(C=C, random_state=seed, solver='liblinear').fit(X[:idx, :], y[:idx])
# Prediction for validation set
y_pred = lr.predict_proba(X[idx:, :])[:, 1]
# Calculate the quality
score = roc_auc_score(y[idx:], y_pred)
return score
%%time
# Select the training set from the united dataframe (where we have the answers)
X_train = full_sites_sparse[:idx_split, :]
# Calculate metric on the validation set
print(get_auc_lr_valid(X_train, y_train))
```
The first model demonstrated the quality of 0.92 on the validation set. Let's take it as the first baseline and starting point. To make a prediction on the test data set **we need to train the model again on the entire training data set** (until this moment, our model used only part of the data for training), which will increase its generalizing ability:
```
# Function for writing predictions to a file
def write_to_submission_file(predicted_labels, out_file,
target='target', index_label="session_id"):
predicted_df = pd.DataFrame(predicted_labels,
index = np.arange(1, predicted_labels.shape[0] + 1),
columns=[target])
predicted_df.to_csv(out_file, index_label=index_label)
# Train the model on the whole training data set
# Use random_state=17 for repeatability
# Parameter C=1 by default, but here we set it explicitly
lr = LogisticRegression(C=1.0, random_state=17, solver='liblinear').fit(X_train, y_train)
# Make a prediction for test data set
X_test = full_sites_sparse[idx_split:,:]
y_test = lr.predict_proba(X_test)[:, 1]
# Write it to the file which could be submitted
write_to_submission_file(y_test, 'baseline_1.csv')
```
If you follow these steps and upload the answer to the competition [page](https://inclass.kaggle.com/c/catch-me-if-you-can-intruder-detection-through-webpage-session-tracking2), you will get `ROC AUC = 0.90812` on the public leaderboard ("A4 baseline 1").
### 4. Model Improvement: Feature Engineering
Now we are going to try to improve the quality of our model by adding new features to the data. But first, answer the following question:
##### 4.5. What years are present in the training and test datasets, if united?
*For discussions, please stick to [ODS Slack](https://opendatascience.slack.com/), channel #mlcourse_ai, pinned thread __#a4_q5__*
- 13 and 14
- 2012 and 2013
- 2013 and 2014
- 2014 and 2015
```
# Your code is here
```
Create a feature that will be a number in YYYYMM format from the date when the session was held, for example 201407 -- year 2014 and 7th month. Thus, we will take into account the monthly [linear trend](http://people.duke.edu/~rnau/411trend.htm) for the entire period of the data provided.
```
# Dataframe for new features
full_new_feat = pd.DataFrame(index=full_df.index)
# Add start_month feature
full_new_feat['start_month'] = full_df['time1'].apply(lambda ts:
100 * ts.year + ts.month).astype('float64')
```
##### 4.6. Plot the graph of the number of Alice sessions versus the new feature, start_month. Choose the correct statement:
*For discussions, please stick to [ODS Slack](https://opendatascience.slack.com/), channel #mlcourse_ai, pinned thread __#a4_q6__*
- Alice wasn't online at all for the entire period
- From the beginning of 2013 to mid-2014, the number of Alice's sessions per month decreased
- The number of Alice's sessions per month is generally constant for the entire period
- From the beginning of 2013 to mid-2014, the number of Alice's sessions per month increased
*Hint: the graph will be more explicit if you treat `start_month` as a categorical ordinal variable*.
```
# Your code is here
```
In this way, we have an illustration and thoughts about the usefulness of the new feature, add it to the training sample and check the quality of the new model:
```
# Add the new feature to the sparse matrix
tmp = full_new_feat[['start_month']].values
X_train = csr_matrix(hstack([full_sites_sparse[:idx_split,:], tmp[:idx_split,:]]))
# Compute the metric on the validation set
print(get_auc_lr_valid(X_train, y_train))
```
The quality of the model has decreased significantly. We added a feature that definitely seemed useful to us, but its usage only worsened the model. Why did it happen?
### Important detour #2: is it necessary to scale features?
Here we give an intuitive reasoning (a rigorous mathematical justification for one or another aspect in linear models you can easily find on the internet). Consider the features more closely: those of them that correspond to the number of visits to a particular web-site per session vary from 0 to 10. The feature `start_month` has a completely different range: from 201301 to 201412, this means the contribution of this variable is significantly greater than the others. It would seem that problem can be avoided if we put less weight in a linear combination of attributes in this case, but in our case logistic regression with regularization is used (by default, this parameter is `C = 1`), which penalizes the model the stronger the greater its weights are. Therefore, for linear methods with regularization, it is recommended to convert features to the same scale (you can read more about the regularization, for example, [here](https://habrahabr.ru/company/ods/blog/322076/)).
One way to do this is standardization: for each observation you need to subtract the average value of the feature and divide this difference by the standard deviation:
$$ x^{*}_{i} = \dfrac{x_{i} - \mu_x}{\sigma_x}$$
The following practical tips can be given:
- It is recommended to scale features if they have essentially different ranges or different units of measurement (for example, the country's population is indicated in units, and the country's GNP in trillions)
- Scale features if you do not have a reason/expert opinion to give a greater weight to any of them
- Scaling can be excessive if the ranges of some of your features differ from each other, but they are in the same system of units (for example, the proportion of middle-aged people and people over 80 among the entire population)
- If you want to get an interpreted model, then build a model without regularization and scaling (most likely, its quality will be worse)
- Binary features (which take only values of 0 or 1) are usually left without conversion, (but)
- If the quality of the model is crucial, try different options and select one where the quality is better
Getting back to `start_month`, let us rescale the new feature and train the model again. This time the quality has increased:
```
# Add the new standardized feature to the sparse matrix
tmp = StandardScaler().fit_transform(full_new_feat[['start_month']])
X_train = csr_matrix(hstack([full_sites_sparse[:idx_split,:], tmp[:idx_split,:]]))
# Compute metric on the validation set
print(get_auc_lr_valid(X_train, y_train))
```
##### 4.7. Add to the training set a new feature "n_unique_sites" – the number of the unique web-sites in a session. Calculate how the quality on the validation set has changed
*For discussions, please stick to [ODS Slack](https://opendatascience.slack.com/), channel #mlcourse_ai, pinned thread __#a4_q7__*
- It has decreased. It is better not to add a new feature.
- It has not changed
- It has decreased. The new feature should be scaled.
- I am confused, and I do not know if it's necessary to scale a new feature.
*Tips: use the nunique() function from `pandas`. Do not forget to include the start_month in the set. Will you scale a new feature? Why?*
```
# Your code is here
```
So, the new feature has slightly decreased the quality, so we will not use it. Nevertheless, do not rush to throw features out because they haven't performed well. They can be useful in a combination with other features (for example, when a new feature is a ratio or a product of two others).
##### 4.8. Add two new features: start_hour and morning. Calculate the metric. Which of these features gives an improvement?
The `start_hour` feature is the hour at which the session started (from 0 to 23), and the binary feature `morning` is equal to 1 if the session started in the morning and 0 if the session started later (we assume that morning means `start_hour` is equal to 11 or less).
Will you scale the new features? Make your assumptions and test them in practice.
*For discussions, please stick to [ODS Slack](https://opendatascience.slack.com/), channel #mlcourse_ai, pinned thread __#a4_q8__*
- None of the features gave an improvement :(
- `start_hour` feature gave an improvement, and `morning` did not
- `morning` feature gave an improvement, and `start_hour` did not
- Both features gave an improvement
*Tip: find suitable functions for working with time series data in [documentation](http://pandas.pydata.org/pandas-docs/stable/api.html). Do not forget to include the `start_month` feature.*
```
# Your code is here
```
### 5. Regularization and Parameter Tuning
We have introduced features that improve the quality of our model in comparison with the first baseline. Can we do even better? After we have changed the training and test sets, it almost always makes sense to search for the optimal hyperparameters - the parameters of the model that do not change during training.
For example, in week 3, you learned that, in decision trees, the depth of the tree is a hyperparameter, but the feature by which splitting occurs and its threshold is not.
In the logistic regression that we use, the weights of each feature are changing, and we find their optimal values during training; meanwhile, the regularization parameter remains constant. This is the hyperparameter that we are going to optimize now.
Calculate the quality on a validation set with a regularization parameter, which is equal to 1 by default:
```
# Compose the training set
tmp_scaled = StandardScaler().fit_transform(full_new_feat[['start_month',
'start_hour',
'morning']])
X_train = csr_matrix(hstack([full_sites_sparse[:idx_split,:],
tmp_scaled[:idx_split,:]]))
# Capture the quality with default parameters
score_C_1 = get_auc_lr_valid(X_train, y_train)
print(score_C_1)
```
We will try to beat this result by optimizing the regularization parameter. We will take a list of possible values of C and calculate the quality metric on the validation set for each of C-values:
```
from tqdm import tqdm
# List of possible C-values
Cs = np.logspace(-3, 1, 10)
scores = []
for C in tqdm(Cs):
scores.append(get_auc_lr_valid(X_train, y_train, C=C))
```
Plot the graph of the quality metric (AUC-ROC) versus the value of the regularization parameter. The value of quality metric corresponding to the default value of C=1 is represented by a horizontal dotted line:
```
plt.plot(Cs, scores, 'ro-')
plt.xscale('log')
plt.xlabel('C')
plt.ylabel('AUC-ROC')
plt.title('Regularization Parameter Tuning')
# horizontal line -- model quality with default C value
plt.axhline(y=score_C_1, linewidth=.5, color='b', linestyle='dashed')
plt.show()
```
##### 4.9. What is the value of parameter C (if rounded to 2 decimals) that corresponds to the highest model quality?
*For discussions, please stick to [ODS Slack](https://opendatascience.slack.com/), channel #mlcourse_ai, pinned thread __#a4_q9__*
- 0.17
- 0.46
- 1.29
- 3.14
```
# Your code is here
```
For the last task in this assignment: train the model using the optimal regularization parameter you found (do not round up to two digits like in the last question). If you do everything correctly and submit your solution, you should see `ROC AUC = 0.92784` on the public leaderboard ("A4 baseline 2"):
```
# Prepare the training and test data
tmp_scaled = StandardScaler().fit_transform(full_new_feat[['start_month', 'start_hour',
'morning']])
X_train = csr_matrix(hstack([full_sites_sparse[:idx_split,:],
tmp_scaled[:idx_split,:]]))
X_test = csr_matrix(hstack([full_sites_sparse[idx_split:,:],
tmp_scaled[idx_split:,:]]))
# Train the model on the whole training data set using optimal regularization parameter
lr = LogisticRegression(C=C, random_state=17, solver='liblinear').fit(X_train, y_train)
# Make a prediction for the test set
y_test = lr.predict_proba(X_test)[:, 1]
# Write it to the submission file
write_to_submission_file(y_test, 'baseline_2.csv')
```
In this part of the assignment, you have learned how to use sparse matrices, train logistic regression models, create new features and selected the best ones, learned why you need to scale features, and how to select hyperparameters. That's a lot!
# Part 2. Freeride
<img src='../../img/snowboard.jpg' width=70%>
*Yorko in Sheregesh, the best palce in Russia for snowboarding and skiing.*
In this part, you'll need to beat the "A4 baseline 3" baseline. No more step-by-step instructions. But it'll be very helpful for you to study the Kernel "[Correct time-aware cross-validation scheme](https://www.kaggle.com/kashnitsky/correct-time-aware-cross-validation-scheme)".
Here are a few tips for finding new features: think about what you can come up with using existing features, try multiplying or dividing two of them, justify or decline your hypotheses with plots, extract useful information from time series data (time1 ... time10), do not hesitate to convert an existing feature (for example, take a logarithm), etc. Checkout other [Kernels](https://www.kaggle.com/c/catch-me-if-you-can-intruder-detection-through-webpage-session-tracking2/kernels). We encourage you to try new ideas and models throughout the course and participate in the competitions - it's fun!
When you get into Kaggle and Xgboost, you'll feel like that, and it's OK :)
<img src='../../img/xgboost_meme.jpg' width=50%>
| github_jupyter |
# Methods to perform error analysis
# Given the result.txt file created by the official script, extract useful data
```
%load_ext autoreload
%autoreload
import os
from sys import path
import re
import pandas as pd
#path.append('..')
import numpy as np
from scipy.stats import ttest_rel
output_path = '/scratch/geeticka/relation-extraction/output/semeval2010/CrossValidation'
def res(path): return os.path.join(output_path, path)
result_file_location = res('cnn_72aaf050-50a5-42b1-b7d8-3893b12fd708_2018-12-16dataset_semeval2010-pos_embed_size_25-num_filters_100-filter_sizes_2,3,4,5-keep_prob_0.500000-early_stop_False-patience_100/Fold0')
```
## Below are the methods that gather the necessary information from the file
```
def read_confusion_matrix_per_line(cur_line):
if re.search(r'.*\|.*', cur_line): # only get those lines which have a pipe operator
splitted_line = cur_line.strip().split()
pipe_seen = 0 # the correct numbers are between two pipes
confusion_matrix_line = []
for val in splitted_line:
if val == '|':
pipe_seen += 1
if pipe_seen == 1 and val != '|': # keep collecting the values as you are
confusion_matrix_line.append(float(val))
return confusion_matrix_line
return None
def read_accuracy_per_line(cur_line):
if cur_line.startswith('Accuracy (calculated'):
accuracy = re.match(r'.*= (.*)%', cur_line).groups()[0]
accuracy = float(accuracy)
return accuracy
return None
def read_precision_recall_f1(cur_line): # assume that the mode is once we have read 'Results for the individual'
match = re.match(r'.*= (.*)%.*= (.*)%.*= (.*)%$', cur_line)
if match:
precision, recall, f1 = match.groups()
return float(precision), float(recall), float(f1)
else:
return None
#if not cur_line.startswith('Micro-averaged result'): # you want to read only up to the point when the relations
# will need to double check above
def get_file_metrics(result_file_location):
result_file = os.path.join(result_file_location, 'result.txt')
official_portion_file = False
individual_relations_f1_portion = False
micro_f1_portion = False
macro_f1_portion = False
confusion_matrix_official = [] # stores the official confusion matrix read from the file
accuracy = None
metrics_indiv_relations = [] # precision, recall and f1 for each relation
metrics_micro = [] # excluding the other relation
metrics_macro = [] # excluding the other relation
with open(result_file, 'r') as result_file:
for cur_line in result_file:
cur_line = cur_line.strip()
if cur_line.startswith('<<< (9+1)-WAY EVALUATION TAKING DIRECTIONALITY INTO ACCOUNT -- OFFICIAL >>>'):
official_portion_file = True
if official_portion_file is False:
continue
confusion_matrix_line = read_confusion_matrix_per_line(cur_line)
if confusion_matrix_line is not None: confusion_matrix_official.append(confusion_matrix_line)
acc = read_accuracy_per_line(cur_line)
if acc is not None: accuracy = acc
# figure out which sub portion of the official portion we are in
if cur_line.startswith('Results for the individual relations:'):
individual_relations_f1_portion = True
elif cur_line.startswith('Micro-averaged result (excluding Other):'):
micro_f1_portion = True
elif cur_line.startswith('MACRO-averaged result (excluding Other):'):
macro_f1_portion = True
# populate the precision, recall and f1 for the correct respective lists
if individual_relations_f1_portion is True and micro_f1_portion is False:
vals = read_precision_recall_f1(cur_line)
if vals is not None: metrics_indiv_relations.append([vals[0], vals[1], vals[2]])
elif micro_f1_portion is True and macro_f1_portion is False:
vals = read_precision_recall_f1(cur_line)
if vals is not None: metrics_micro.append([vals[0], vals[1], vals[2]])
elif macro_f1_portion is True:
vals = read_precision_recall_f1(cur_line)
if vals is not None: metrics_macro.append([vals[0], vals[1], vals[2]])
return confusion_matrix_official, accuracy, metrics_indiv_relations, metrics_micro, metrics_macro
```
## Get the file metrics
```
confusion_matrix_official, accuracy, \
metrics_indiv_relations, metrics_micro, metrics_macro = get_file_metrics(result_file_location)
```
### Generate the confusion matrix as a pandas dataframe
https://stackoverflow.com/questions/17091769/python-pandas-fill-a-dataframe-row-by-row
and https://stackoverflow.com/questions/35047842/how-to-store-the-name-of-rows-and-column-index-in-pandas-dataframe
```
relation_full_form_dictionary = {'C-E': 'Cause-Effect', 'C-W': 'Component-Whole', 'C-C': 'Content-Container',
'E-D': 'Entity-Destination', 'E-O': 'Entity-Origin', 'I-A': 'Instrument-Agency',
'M-C': 'Member-Collection', 'M-T': 'Message-Topic', 'P-P': 'Product-Producer',
'_O': 'Other'}
relation_as_short_list = ['C-E', 'C-W', 'C-C', 'E-D', 'E-O', 'I-A', 'M-C', 'M-T', 'P-P', '_O']
def get_confusion_matrix_as_df(confusion_matrix_official, relations_as_short_list):
index = pd.Index(relations_as_short_list, name='gold labels')
columns = pd.Index(relations_as_short_list, name='predicted')
confusion_matrix_df = pd.DataFrame(data=confusion_matrix_official, columns=columns,index=index)
return confusion_matrix_df
confusion_matrix_df = get_confusion_matrix_as_df(confusion_matrix_official, relation_as_short_list)
```
#### Give the confusions across each relation, with a special interest on other
```
# we want to go row by row, and get only those column names which have 0 values.
#and the number associated with that column name as a string.
# and we also want to get the correct values as a separate column
def generate_confused_with_string(index, row, relation_full_form_dictionary):
# index is the current relation that we are considering and row is all the predicted examples
confused_with_string = ""
num_of_columns = len(row.index)
for i in range(0, num_of_columns):
column_name = row.index[i]
column_value = int(row.loc[column_name])
if column_value > 0 and column_name != index:
confused_with_string += " " + relation_full_form_dictionary[column_name] + \
"(" + str(column_value) + ")"
return confused_with_string.strip()
#print(row.data[0])
#for val in row:
# print(val)
def generate_pretty_summary_confusion_matrix(confusion_matrix_df, relation_full_form_dictionary):
data = [] # index will be 0,1,2 and so on, but columns will be
# Actual label, confused with as a string, correct predictions as a number
for index, row in confusion_matrix_df.iterrows():
actual_label = relation_full_form_dictionary[index]
confused_with = generate_confused_with_string(index, row, relation_full_form_dictionary)
correct_predictions = row[index] # eg: gives the column value for C-E for an index C-E
if index != '_O': confused_with_other = row['_O'] # this is specific to semeval and will need to be changed
else: confused_with_other = None
data.append([actual_label, confused_with, confused_with_other, correct_predictions])
columns = pd.Index(['Gold Relation', 'Confused With(num_examples)', 'Confused with Other', 'Correct Predictions'], name='summary')
pretty_summary_confusion_matrix_df = pd.DataFrame(data=data, columns=columns)
return pretty_summary_confusion_matrix_df
```
#### Give the individual relation metrics as a dataframe
```
def create_metrics_indiv_relations_df(metrics_indiv_relations, relation_full_form_dictionary, relation_as_short_list):
index_list = relation_as_short_list
index_list_verbose = [relation_full_form_dictionary[x] for x in index_list]
index = pd.Index(index_list_verbose, name='labels')
columns = pd.Index(['Precision', 'Recall', 'F1'], name='metrics')
metrics_indiv_relations_df = pd.DataFrame(data=metrics_indiv_relations, columns=columns,index=index)
return metrics_indiv_relations_df
def create_metrics_macro_micro_df(metrics_macro, metrics_micro):
data = metrics_macro + metrics_micro
index = pd.Index(['macro', 'micro'], name='calculation type')
columns = pd.Index(['Precision', 'Recall', 'F1'], name='metrics')
metrics_macro_micro = pd.DataFrame(data=data, columns=columns,index=index)
return metrics_macro_micro
```
## Finally, create a large summary function
```
relation_full_form_dictionary = {'C-E': 'Cause-Effect', 'C-W': 'Component-Whole', 'C-C': 'Content-Container',
'E-D': 'Entity-Destination', 'E-O': 'Entity-Origin', 'I-A': 'Instrument-Agency',
'M-C': 'Member-Collection', 'M-T': 'Message-Topic', 'P-P': 'Product-Producer',
'_O': 'Other'}
relation_as_short_list = ['C-E', 'C-W', 'C-C', 'E-D', 'E-O', 'I-A', 'M-C', 'M-T', 'P-P', '_O']
def create_summary(result_file_location, relation_full_form_dictionary, relation_as_short_list):
if not os.path.exists(result_file_location):
print("Check your path first!")
return None
# get the file metrics
confusion_matrix_official, accuracy, \
metrics_indiv_relations, metrics_micro, metrics_macro = get_file_metrics(result_file_location)
# get the confusion matrix dataframe
confusion_matrix_df = get_confusion_matrix_as_df(confusion_matrix_official, relation_as_short_list)
# these are the summary information that will need to be returned
pretty_summary_confusion_matrix_df = generate_pretty_summary_confusion_matrix(confusion_matrix_df,
relation_full_form_dictionary)
total_correct_predictions = pretty_summary_confusion_matrix_df['Correct Predictions'].sum()
metrics_indiv_relations_df = create_metrics_indiv_relations_df(metrics_indiv_relations,
relation_full_form_dictionary,
relation_as_short_list)
metrics_macro_micro = create_metrics_macro_micro_df(metrics_macro, metrics_micro)
# report accuracy as well
return confusion_matrix_df, pretty_summary_confusion_matrix_df, total_correct_predictions, metrics_indiv_relations_df, \
metrics_macro_micro, accuracy
confusion_matrix_df, pretty_summary_confusion_matrix_df, \
total_correct_predictions, metrics_indiv_relations_df, \
metrics_macro_micro, accuracy = create_summary(result_file_location, relation_full_form_dictionary, relation_as_short_list)
# given the confusion matrix, return the sums of all the examples
def get_sum_confusion_matrix(confusion_matrix):
sum = 0
for column in confusion_matrix:
sum += confusion_matrix[column].sum()
return sum
# for each of the relations, do a t test between the two model metrics
def indiv_metric_comparison(metrics_i_model1, metrics_i_model2, model1_name, model2_name):
print("TTest from %s to %s"%(model1_name, model2_name))
print("Below is the metric comparsion across the two models" + \
"considering individual relations, excluding 'Other'")
for column in metrics_i_model1:
metric_model1 = metrics_i_model1[column].tolist()[:-1] # excluding "Other"
metric_model2 = metrics_i_model2[column].tolist()[:-1]
tt = ttest_rel(metric_model1, metric_model2)
print("Metric: %s \t statistic %.2f \t p_value %s"%
(column, tt.statistic, tt.pvalue))
def get_macro_micro_metric_comparison(metrics_ma_mi_model1, metrics_ma_mi_model2, model1_name, model2_name):
print("Macro - Micro for the %s model"%(model1_name))
for column in metrics_ma_mi_model1:
macro = metrics_ma_mi_model1[column].loc['macro']
micro = metrics_ma_mi_model1[column].loc['micro']
print("Metric: %s \t Macro-Micro %.2f"%(column, macro-micro))
print("\nMacro - Micro for the %s model"%(model2_name))
for column in metrics_ma_mi_model2:
macro = metrics_ma_mi_model2[column].loc['macro']
micro = metrics_ma_mi_model2[column].loc['micro']
print("Metric: %s \t Macro-Micro %.2f"%(column, macro-micro))
print("\nMacro_%s - Macro_%s"%(model1_name, model2_name))
for column in metrics_ma_mi_model1:
macro_model1 = metrics_ma_mi_model1[column].loc['macro']
macro_model2 = metrics_ma_mi_model2[column].loc['macro']
print("Metric: %s \t Difference %.2f"%(column, macro_model1-macro_model2))
print("\nMicro_%s - Micro_%s"%(model1_name, model2_name))
for column in metrics_ma_mi_model1:
micro_model1 = metrics_ma_mi_model1[column].loc['micro']
micro_model2 = metrics_ma_mi_model2[column].loc['micro']
print("Metric: %s \t Difference %.2f"%(column, micro_model1-micro_model2))
def get_accuracy_difference(accuracy_model1, accuracy_model2, model1_name, model2_name):
print("Accuracy_%s - Accuracy_%s %.2f"%(model1_name, model2_name, accuracy_model1 - accuracy_model2))
```
In nightingale, I created 2 folders inside
/scratch/geeticka/relation-extraction/output/semeval2010/CrossValidation/error-analysis
- One folder is baseline, the other is elmo-model
- Each folder has result.txt for each Fold
- I am going to generate a summary for each of the Folds
```
def print_full_summary(model1_loc, model2_loc, model1_name, model2_name, res,
relation_full_form_dictionary, relation_as_short_list):
model1 = res(model1_loc)
model2 = res(model2_loc)
cm_model1, summary_cm_model1, correct_pred_model1, metrics_i_model1, \
metrics_ma_mi_model1, accuracy_model1 \
= create_summary(model1, relation_full_form_dictionary, relation_as_short_list)
cm_model2, summary_cm_model2, correct_pred_model2, metrics_i_model2, \
metrics_ma_mi_model2, accuracy_model2 \
= create_summary(model2, relation_full_form_dictionary, relation_as_short_list)
# T Test of each metrics, across the relations not Other
indiv_metric_comparison(metrics_i_model1, metrics_i_model2, model1_name, model2_name)
# Get the difference in the macro and micro scores
get_macro_micro_metric_comparison(metrics_ma_mi_model1, metrics_ma_mi_model2, model1_name, model2_name)
# Print the accuracy difference as well
get_accuracy_difference(accuracy_model1, accuracy_model2, model1_name, model2_name)
return summary_cm_model1, summary_cm_model2
summary_cm_baseline, summary_cm_elmo = print_full_summary(
'error-analysis/baseline/Fold0', 'error-analysis/elmo-model/Fold0',
'Baseline','Elmo', res, relation_full_form_dictionary, relation_as_short_list)
summary_cm_baseline
summary_cm_elmo
```
| github_jupyter |
# Calibration
## Introduction
This tutorial describes how to use **skrf** to calibrate data taken from a VNA. For an introduction to VNA calibration see this article by Rumiantsev and Ridler [1]_, for an outline of different calibration algorithms see Doug Ryttings presentation [2]_. If you like to read books, then you may want to checkout [3]_ .
What follows is are various examples of how to calibrate a device under test (DUT), assuming you have measured an acceptable set of standards, and have a corresponding set ideal responses. This may be reffered to as *offline* calibration, because it is not occuring onboard the VNA itself. One benefit of this technique is that it provides maximum flexibilty for non-conventional calibrations, and preserves all raw data.
## Creating a Calibration
Calibrations are performed through a [Calibration](../api/calibration/index.rst) class. In General, [Calibration](../api/calibration/index.rst) objects require two arguments:
* a list of measured `Network`'s
* a list of ideal `Network`'s
The `Network` elements in each list must all be similar (same number of ports, frequency info, etc) and must be aligned to each other, meaning the first element of ideals list must correspond to the first element of measured list. Self calibration algorithms, such as TRL, do not require predefined ideal responses.
## One-Port Example
This example code is written to be instructive, not concise. To
construct a one-port calibration you need to have measured at least
three standards and have their known *ideal* responses in the form of
`Network`s. These `Network`
can be created from touchstone files, a format all modern VNA's support.
In the following script the measured and ideal touchstone files for
a conventional short-open-load (SOL) calkit are in folders `measured/`
and `ideal/`, respectively. These are used to create a OnePort Calibration and correct a measured DUT
import skrf as rf
from skrf.calibration import OnePort
## created necessary data for Calibration class
# a list of Network types, holding 'ideal' responses
my_ideals = [\
rf.Network('ideal/short.s1p'),
rf.Network('ideal/open.s1p'),
rf.Network('ideal/load.s1p'),
]
# a list of Network types, holding 'measured' responses
my_measured = [\
rf.Network('measured/short.s1p'),
rf.Network('measured/open.s1p'),
rf.Network('measured/load.s1p'),
]
## create a Calibration instance
cal = rf.OnePort(\
ideals = my_ideals,
measured = my_measured,
)
## run, and apply calibration to a DUT
# run calibration algorithm
cal.run()
# apply it to a dut
dut = rf.Network('my_dut.s1p')
dut_caled = cal.apply_cal(dut)
dut_caled.name = dut.name + ' corrected'
# plot results
dut_caled.plot_s_db()
# save results
dut_caled.write_touchstone()
## Concise One-port
This example achieves the same task as the one above except in a
more concise *programmatic* way.
import skrf as rf
from skrf.calibration import OnePort
my_ideals = rf.load_all_touchstones_in_dir('ideals/')
my_measured = rf.load_all_touchstones_in_dir('measured/')
## create a Calibration instance
cal = rf.OnePort(\
ideals = [my_ideals[k] for k in ['short','open','load']],
measured = [my_measured[k] for k in ['short','open','load']],
)
## what you do with 'cal' may may be similar to above example
## Two-port Calibrations
Naturally, two-port calibration is more involved than one-port. **skrf**
supports a few different two-port algorithms. The traditional `SOLT` algorithm uses the 12-term error model. This algorithms is straightforward, and similar to the OnePort example.
The `EightTerm` calibration is based on the algorithm described in [4]_, by R.A. Speciale. It can be constructed from any number of standards, providing that some fundamental constraints are met. In short, you need three two-port standards; one must be transmissive, and one must provide a known impedance and be reflective.
Note, that the word *8-term* is used in the literature to describe a specific error model used by a variety of calibration algorihtms, like TRL, LRM, etc. The `EightTerm` class, is an implementation of the algorithm cited above, which does not use any self-calibration.
One important detail of using the 8-term error model formulation is that switch-terms may need to be measured in order to achieve a high quality calibration (thanks to Dylan Williams for pointing this out). These are described next.
## Switch-terms
Originally described by Roger Marks [5]_ , switch-terms account for the fact that the 8-term (aka *error-box* ) model is overly simplified. The two error networks change slightly depending on which port is being excited. This is due to the internal switch within the VNA.
Switch terms can be directly measured with a custom measurement configuration on the VNA itself. **skrf** has support for measuring switch terms in the `skrf.vi.vna` module, see the HP8510's `skrf.vi.vna.HP8510C.switch_terms`, or PNA's `skrf.vi.vna.PNA.get_switch_terms` . Without switch-term measurements, your calibration quality will vary depending on properties of you VNA.
## Using one-port ideals in two-port Calibration
Commonly, you have data for ideal data for reflective standards in the form of one-port touchstone files (ie `.s1p`). To use this with skrf's two-port calibration method you need to create a two-port network that is a composite of the two networks. The function `skrf.network.two_port_reflect` does this
short = rf.Network('ideals/short.s1p')
shorts = rf.two_port_reflect(short, short)
## SOLT Example
Two-port calibration is accomplished in an identical way to one-port, except all the standards are two-port networks. This is even true of reflective standards (S21=S12=0). So if you measure reflective standards you must measure two of them simultaneously, and store information in a two-port. For example, connect a short to port-1 and a load to port-2, and save a two-port measurement as 'short,load.s2p' or similar
import skrf as rf
from skrf.calibration import SOLT
# a list of Network types, holding 'ideal' responses
my_ideals = [
rf.Network('ideal/thru.s2p'),
rf.Network('ideal/short, short.s2p'),
rf.Network('ideal/open, open.s2p'),
rf.Network('ideal/load, load.s2p'),
]
# a list of Network types, holding 'measured' responses
my_measured = [
rf.Network('measured/thru.s2p'),
rf.Network('measured/short, short.s2p'),
rf.Network('measured/open, open.s2p'),
rf.Network('measured/load, load.s2p'),
]
## create a SOLT instance
cal = SOLT(
ideals = my_ideals,
measured = my_measured,
)
## run, and apply calibration to a DUT
# run calibration algorithm
cal.run()
# apply it to a dut
dut = rf.Network('my_dut.s2p')
dut_caled = cal.apply_cal(dut)
# plot results
dut_caled.plot_s_db()
# save results
dut_caled.write_touchstone()
## Saving and Recalling a Calibration
[Calibration](../api/calibration/index.rst)'s can be written-to and
read-from disk using the temporary storage container of pickles. Writing can be accomplished by using `Calibration.write`, or `rf.write()`, and reading is done with `rf.read()`. Since these functions rely on pickling, they are not recomended for long-term data storage. Currently there is no way to achieve long term storage of a Calibration object, other than saving the script used to generate it.
| github_jupyter |
```
%pylab inline
import numpy as np
import pymc3 as pm
from scipy.special import expit
from sklearn.preprocessing import LabelBinarizer
import matplotlib.pyplot as plt
items = 400
k_mean = -7
k_sigma = 0.5
noise_sigma = 0.15
trials_lambda = 0.02
samples_per_item = 3
# drawing values at an item level
item_conversion_rates = np.random.normal(loc=k_mean, scale=k_sigma, size=items)
# values at a sample level
labels = np.repeat(np.arange(items), samples_per_item)
sample_conversion_rates = np.repeat(item_conversion_rates, samples_per_item)
trials = np.random.geometric(p=trials_lambda, size=samples_per_item * items)
adjusted_conversion_rate = expit(sample_conversion_rates)
conversions = np.random.binomial(trials, adjusted_conversion_rate)
plt.hist(adjusted_conversion_rate, 100);
with pm.Model() as indexed_model:
# Priors for unknown model parameters
k_mu = pm.Normal('c_mu', mu=0., sd=10.)
c_sigma = pm.HalfNormal('c_sigma', sd=5.)
# Matt trick
k_step = pm.Normal('k_step', mu=0, sd=1, shape=items)
k = k_mu + k_step * c_sigma
# Modeling conversions
k_indexed = k[labels]
conv_rate = pm.math.invlogit(k_indexed)
# Likelihood
product_conversions = pm.Binomial('product_conversion',
n=trials,
p=conv_rate,
observed=conversions)
indexed_trace = pm.sample(1000, tune=1000)
pm.traceplot(indexed_trace, varnames=['c_mu', 'c_sigma']);
indexed_trace.varnames
indexed_trace.stat_names
diverge = indexed_trace.get_sampler_stats('diverging', combine=True)
plt.scatter(indexed_trace['c_mu'], indexed_trace['c_sigma_log__'],
marker='.', alpha=0.1)
plt.scatter(indexed_trace['c_mu'][diverge], indexed_trace['c_sigma_log__'][diverge],
marker='.', alpha=0.5, color='r')
plt.show()
with pm.Model():
# Priors for unknown model parameters
k_mu = pm.Normal('c_mu', mu=0., sd=10.)
c_sigma = pm.HalfNormal('c_sigma', sd=5., transform=None)
# Matt trick
k_step = pm.Normal('k_step', mu=0, sd=1, shape=items)
k = k_mu + k_step * c_sigma
# Modeling conversions
k_indexed = k[labels]
conv_rate = pm.math.invlogit(k_indexed)
# Likelihood
product_conversions = pm.Binomial('product_conversion',
n=trials,
p=conv_rate,
observed=conversions)
indexed_trace_ = pm.sample(1000, tune=1000)
diverge = indexed_trace_.get_sampler_stats('diverging', combine=True)
mu_post = indexed_trace['c_mu']
sigma_post = indexed_trace['c_sigma']
plt.scatter(mu_post, sigma_post,
marker='.', alpha=0.1)
plt.scatter(mu_post[diverge], sigma_post[diverge],
marker='.', alpha=0.5, color='r')
plt.show()
```
| github_jupyter |
```
# Copyright 2021 NVIDIA Corporation. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
```
# <img src="http://developer.download.nvidia.com/compute/machine-learning/frameworks/nvidia_logo.png" style="width: 90px; float: right;">
# Creating Multi-Modal Movie Feature Store
Finally, with both the text and image features ready, we now put the multi-modal movie features into a unified feature store.
If you have downloaded the real data and proceeded through the feature extraction process in notebooks 03-05, then proceed to create the feature store. Else, skip to the `Synthetic data` section below to create random features.
## Real data
```
import pickle
with open('movies_poster_features.pkl', 'rb') as f:
poster_feature = pickle.load(f)["feature_dict"]
len(poster_feature)
with open('movies_synopsis_embeddings-1024.pkl', 'rb') as f:
text_feature = pickle.load(f)["embeddings"]
len(text_feature)
import pandas as pd
links = pd.read_csv("./data/ml-25m/links.csv", dtype={"imdbId": str})
links.shape
links.head()
poster_feature['0105812'].shape
import numpy as np
feature_array = np.zeros((len(links), 1+2048+1024))
for i, row in links.iterrows():
feature_array[i,0] = row['movieId']
if row['imdbId'] in poster_feature:
feature_array[i,1:2049] = poster_feature[row['imdbId']]
if row['movieId'] in text_feature:
feature_array[i,2049:] = text_feature[row['movieId']]
dtype= {**{'movieId': np.int64},**{x: np.float32 for x in ['poster_feature_%d'%i for i in range(2048)]+['text_feature_%d'%i for i in range(1024)]}}
len(dtype)
feature_df = pd.DataFrame(feature_array, columns=['movieId']+['poster_feature_%d'%i for i in range(2048)]+['text_feature_%d'%i for i in range(1024)])
feature_df.head()
feature_df.shape
!pip install pyarrow
feature_df.to_parquet('feature_df.parquet')
```
## Synthetic data
If you have not extrated image and text features from real data, proceed with this section to create synthetic features.
```
import pandas as pd
links = pd.read_csv("./data/ml-25m/links.csv", dtype={"imdbId": str})
import numpy as np
feature_array = np.random.rand(links.shape[0], 3073)
feature_array[:,0] = links['movieId'].values
feature_df = pd.DataFrame(feature_array, columns=['movieId']+['poster_feature_%d'%i for i in range(2048)]+['text_feature_%d'%i for i in range(1024)])
feature_df.to_parquet('feature_df.parquet')
feature_df.head()
```
| github_jupyter |
# Benchmarks
## Setup
We'll use the [credit card dataset](https://datahub.io/machine-learning/creditcard) that is commonly used as an imbalanced binary classification task.
```
!wget https://datahub.io/machine-learning/creditcard/r/creditcard.csv
```
Do a train/test split.
```
import pandas as pd
from sklearn import preprocessing
credit = pd.read_csv('creditcard.csv')
credit = credit.drop(columns=['Time'])
features = credit.columns.drop('Class')
credit.loc[:, features] = preprocessing.scale(credit.loc[:, features])
credit['Class'] = (credit['Class'] == "'1'").astype(int)
n_test = 40_000
credit[:-n_test].to_csv('train.csv', index=False)
credit[-n_test:].to_csv('test.csv', index=False)
```
While we're at it, let's look at the class distribution.
```
credit['Class'].value_counts(normalize=True)
```
Define a helper function to train a PyTorch model on an `IterableDataset`.
```
import torch
def train(net, optimizer, criterion, train_batches):
for x_batch, y_batch in train_batches:
optimizer.zero_grad()
y_pred = net(x_batch)
loss = criterion(y_pred[:, 0], y_batch.float())
loss.backward()
optimizer.step()
return net
```
Define a helper function to score a PyTorch model on a test set.
```
import numpy as np
def score(net, test_batches, metric):
y_true = []
y_pred = []
for x_batch, y_batch in test_batches:
y_true.extend(y_batch.detach().numpy())
y_pred.extend(net(x_batch).detach().numpy()[:, 0])
y_true = np.array(y_true)
y_pred = np.array(y_pred)
return metric(y_true, y_pred)
```
Let's also create an `IterableDataset` that reads from a CSV file. The following implementation is not very generic nor flexible, but it will do for this notebook.
```
import csv
class IterableCSV(torch.utils.data.IterableDataset):
def __init__(self, path):
self.path = path
def __iter__(self):
with open(self.path) as file:
reader = csv.reader(file)
header = next(reader)
for row in reader:
x = [float(el) for el in row[:-1]]
y = int(row[-1])
yield torch.tensor(x), y
```
We can now define the training and test set loaders.
```
train_set = IterableCSV(path='train.csv')
test_set = IterableCSV(path='test.csv')
```
## Vanilla
```
def make_net(n_features):
torch.manual_seed(0)
return torch.nn.Sequential(
torch.nn.Linear(n_features, 30),
torch.nn.Linear(30, 1)
)
%%time
net = make_net(len(features))
net = train(
net,
optimizer=torch.optim.SGD(net.parameters(), lr=1e-2),
criterion=torch.nn.BCEWithLogitsLoss(),
train_batches=torch.utils.data.DataLoader(train_set, batch_size=16)
)
from sklearn import metrics
score(
net,
test_batches=torch.utils.data.DataLoader(test_set, batch_size=16),
metric=metrics.roc_auc_score
)
```
## Under-sampling
```
import pytorch_resample
train_sample = pytorch_resample.UnderSampler(
train_set,
desired_dist={0: .8, 1: .2},
seed=42
)
%%time
net = make_net(len(features))
net = train(
net,
optimizer=torch.optim.SGD(net.parameters(), lr=1e-2),
criterion=torch.nn.BCEWithLogitsLoss(),
train_batches=torch.utils.data.DataLoader(train_sample, batch_size=16)
)
score(
net,
test_batches=torch.utils.data.DataLoader(test_set, batch_size=16),
metric=metrics.roc_auc_score
)
```
## Over-sampling
```
train_sample = pytorch_resample.OverSampler(
train_set,
desired_dist={0: .8, 1: .2},
seed=42
)
%%time
net = make_net(len(features))
net = train(
net,
optimizer=torch.optim.SGD(net.parameters(), lr=1e-2),
criterion=torch.nn.BCEWithLogitsLoss(),
train_batches=torch.utils.data.DataLoader(train_sample, batch_size=16)
)
score(
net,
test_batches=torch.utils.data.DataLoader(test_set, batch_size=16),
metric=metrics.roc_auc_score
)
```
## Hybrid method
```
train_sample = pytorch_resample.HybridSampler(
train_set,
desired_dist={0: .8, 1: .2},
sampling_rate=.5,
seed=42
)
%%time
net = make_net(len(features))
net = train(
net,
optimizer=torch.optim.SGD(net.parameters(), lr=1e-2),
criterion=torch.nn.BCEWithLogitsLoss(),
train_batches=torch.utils.data.DataLoader(train_sample, batch_size=16)
)
score(
net,
test_batches=torch.utils.data.DataLoader(test_set, batch_size=16),
metric=metrics.roc_auc_score
)
```
| github_jupyter |
```
import time
import cv2
import matplotlib.pyplot as plt
%matplotlib inline
```
# Haar Cascades
How does Haar cascade work?
<br>
- Trained on two image categories: one class with faecs and the other without.
1. Haar feature selection: slides windows from left to right, top to bottom to detect features of faces (combination of edge, lines and diagonals with specific spatial arrangement).
2. Compute integral image -- a faster way to compute feature values compared to convolution.
3. Model training using Adaboost: returns higher confidence score in the area where the model thinks it is a face. Remove redundant features and retain only the relevant ones using weak classifiers with weighted final score.
4. Cascading classifiers: if there are enough higher scores in a region, that becomes a detected face. To minimize false negative rate by rejecting non-face regions (decision tree).
- In-depth explanation: https://medium.com/geeky-bawa/face-identification-using-haar-cascade-classifier-af3468a44814
<br>
- Advantages: Lightweight, fast, small model size.
<br>
- Disadvantages: False positives, fair accuracy compared to deep learning models.
<br>
- Download the pre-trained face detector model on OpenCV GitHub: https://github.com/opencv/opencv/blob/master/data/haarcascades/haarcascade_frontalface_default.xml
# Detect Faces in an Image
```
# load classifier
face_detector = cv2.CascadeClassifier('./haarcascade_frontalface_default.xml')
# load image
img = cv2.imread('./group-photo.jpg')
# convert to RGB and greyscale
img_gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
img_RGB = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
plt.imshow(img_RGB)
```
Photo by Leah Hetteberg on Unsplash: https://unsplash.com/photos/Kb8_42IFMFk
```
# detect faces
detected_faces = face_detector.detectMultiScale(image=img_gray, scaleFactor=1.1, minNeighbors=20)
# draw a bounding box for every detected face
for (top_left_x, top_left_y, width, height) in detected_faces:
cv2.rectangle(img=img_RGB, pt1=(top_left_x, top_left_y), pt2=(top_left_x+width, top_left_y+height),
color=(255, 0, 0), thickness=30)
# display image
plt.imshow(img_RGB)
```
# Detect Faces in a Video
```
# capture video from default camera
video_frames = cv2.VideoCapture(0) # 0: default camera
# get frame size from camera
frame_width = int(video_frames.get(cv2.CAP_PROP_FRAME_WIDTH))
frame_height = int(video_frames.get(cv2.CAP_PROP_FRAME_HEIGHT))
# initialize writer
# fourcc: video codec. DIVX for windows, XVID for linux and macOS
# fps: 30
writer = cv2.VideoWriter('./my_video_face.mp4', cv2.VideoWriter_fourcc(*'DIVX'), 30, (frame_width, frame_height))
# loop: grab frame and display image
while True:
# read frames
ret, frame = video_frames.read()
# convert frame to greyscale
frame_gray = cv2.cvtColor(frame, cv2.COLOR_BGR2GRAY)
# detect faces
detected_faces = face_detector.detectMultiScale(image=frame_gray, scaleFactor=1.1, minNeighbors=20)
# draw a bounding box for every detected face
for (top_left_x, top_left_y, width, height) in detected_faces:
cv2.rectangle(img=frame, pt1=(top_left_x, top_left_y), pt2=(top_left_x+width, top_left_y+height),
color=(255, 0, 0), thickness=2)
# save video
writer.write(frame)
# show frame
cv2.imshow('frame', frame)
# if frame is display for more than 1 ms and ESC key is pressed, close display
if cv2.waitKey(1) & 0xFF == 27:
break
video_frames.release()
writer.release()
cv2.destroyAllWindows()
```
| github_jupyter |
```
import nltk
from nltk.corpus import twitter_samples
import matplotlib.pyplot as plt
import random
import re
import string
import numpy as np
from nltk.corpus import stopwords
from nltk.stem import PorterStemmer
from nltk.tokenize import TweetTokenizer
def process_tweet(tweet):
"""Process tweet function.
Input:
tweet: a string containing a tweet
Output:
tweets_clean: a list of words containing the processed tweet
"""
stemmer = PorterStemmer()
stopwords_english = stopwords.words('english')
# remove stock market tickers like $GE
tweet = re.sub(r'\$\w*', '', tweet)
# remove old style retweet text "RT"
tweet = re.sub(r'^RT[\s]+', '', tweet)
# remove hyperlinks
tweet = re.sub(r'https?:\/\/.*[\r\n]*', '', tweet)
# remove hashtags
# only removing the hash # sign from the word
tweet = re.sub(r'#', '', tweet)
# tokenize tweets
tokenizer = TweetTokenizer(preserve_case=False, strip_handles=True,
reduce_len=True)
tweet_tokens = tokenizer.tokenize(tweet)
tweets_clean = []
for word in tweet_tokens:
if (word not in stopwords_english and # remove stopwords
word not in string.punctuation): # remove punctuation
# tweets_clean.append(word)
stem_word = stemmer.stem(word) # stemming word
tweets_clean.append(stem_word)
return tweets_clean
def build_freqs(tweets, ys):
"""Build frequencies.
Input:
tweets: a list of tweets
ys: an m x 1 array with the sentiment label of each tweet
(either 0 or 1)
Output:
freqs: a dictionary mapping each (word, sentiment) pair to its
frequency
"""
# Convert np array to list since zip needs an iterable.
# The squeeze is necessary or the list ends up with one element.
# Also note that this is just a NOP if ys is already a list.
yslist = np.squeeze(ys).tolist()
# Start with an empty dictionary and populate it by looping over all tweets
# and over all processed words in each tweet.
freqs = {}
for y, tweet in zip(yslist, tweets):
for word in process_tweet(tweet):
pair = (word, y)
if pair in freqs:
freqs[pair] += 1
else:
freqs[pair] = 1
return freqs
#nltk.download('twitter_samples')
pos_tweets = twitter_samples.strings('positive_tweets.json')
neg_tweets = twitter_samples.strings('negative_tweets.json')
print(len(pos_tweets))
print(len(neg_tweets))
fig = plt.figure(figsize=(5,5))
labels = 'Pos','Neg'
sizes = [len(pos_tweets),len(neg_tweets)]
plt.pie(sizes, labels=labels,shadow=True)
plt.axis('equal')
plt.show()
# Preprocess
# Tokenize, lowercase, removing stopwords, stemming
tweet = pos_tweets [300]
print(tweet)
nltk.download('stopwords')
import re
import string
from nltk.corpus import stopwords
from nltk.stem import PorterStemmer
from nltk.tokenize import TweetTokenizer
# remove hyperlinks, marks, styles
print('\033[92m' + tweet)
print('\033[94m')
# remove old style retweet text "RT"
tweet2 = re.sub(r'^RT[\s]+', '', tweet)
# remove hyperlinks
tweet2 = re.sub(r'https?:\/\/.*[\r\n]*', '', tweet2)
# remove hashtags
# only removing the hash # sign from the word
tweet2 = re.sub(r'#', '', tweet2)
print(tweet2)
# Tokenize the string
print('\033[92m' + tweet2)
print('\033[94m')
tokenizer = TweetTokenizer(preserve_case=False, strip_handles=True, reduce_len=True)
tweet_tokens = tokenizer.tokenize(tweet2)
print()
print('Tokenize String')
print(tweet_tokens)
# remove stopwords
# import the english stop words list from nltk
stopwords_english = stopwords.words('english')
print ('Stop words')
print(stopwords_english)
print('\nPunctuation\n')
print(string.punctuation)
# clean up the text
print()
print('\033[92m')
print(tweet_tokens)
print('\033[94m')
tweets_clean = []
for word in tweet_tokens:
if(word not in stopwords_english and
word not in string.punctuation):
tweets_clean.append(word)
print('All clean')
print(tweets_clean)
# stemming
print()
print('\033[92m')
print(tweet_tokens)
print('\033[94m')
stemmer = PorterStemmer()
tweet_stem = []
for word in tweets_clean:
stem_word = stemmer.stem(word)
tweet_stem.append(stem_word)
print('stemmed word')
print(tweet_stem)
tweet = pos_tweets[455]
print()
print('\033[92m')
print(tweet)
print('\033[94m')
tweets_stem = process_tweet(tweet); # Preprocess a given tweet
print('preprocessed tweet:')
print(tweets_stem) # Print the result
# create frequency dictionary
freqs = build_freqs(tweet, labels)
# check data type
print(f'type(freqs) = {type(freqs)}')
# check length of the dictionary
print(f'len(freqs) = {len(freqs)}')
# select some words to appear in the report. we will assume that each word is unique (i.e. no duplicates)
keys = ['happi', 'merri', 'nice', 'good', 'bad', 'sad', 'mad', 'best', 'pretti',
'❤', ':)', ':(', '😒', '😬', '😄', '😍', '♛',
'song', 'idea', 'power', 'play', 'magnific']
# list representing our table of word counts.
# each element consist of a sublist with this pattern: [<word>, <positive_count>, <negative_count>]
data = []
# loop through our selected words
for word in keys:
# initialize positive and negative counts
pos = 0
neg = 0
# retrieve number of positive counts
if (word, 1) in freqs:
pos = freqs[(word, 1)]
# retrieve number of negative counts
if (word, 0) in freqs:
neg = freqs[(word, 0)]
# append the word counts to the table
data.append([word, pos, neg])
data
tweets = pos_tweets + neg_tweets
labels = np.append(np.ones((len(pos_tweets),1)),np.zeros((len(neg_tweets),1)), axis = 0)
train_pos = pos_tweets[:4000]
train_neg = neg_tweets[:4000]
train_x = train_pos + train_neg
print(len(train_x))
import pandas as pd
data = pd.read_csv('logistic_features.csv')
data.head()
X = data[['bias','positive','negative']].values
Y = data['sentiment'].values
print(X.shape)
theta = [7e-08, 0.0005239, -0.00055517]
# Equation for the separation plane
# It give a value in the negative axe as a function of a positive value
# f(pos, neg, W) = w0 + w1 * pos + w2 * neg = 0
# s(pos, W) = (w0 - w1 * pos) / w2
def neg(theta, pos):
return (-theta[0] - pos * theta[1]) / theta[2]
# Equation for the direction of the sentiments change
# We don't care about the magnitude of the change. We are only interested
# in the direction. So this direction is just a perpendicular function to the
# separation plane
# df(pos, W) = pos * w2 / w1
def direction(theta, pos):
return pos * theta[2] / theta[1]
```
| github_jupyter |
```
def my_function():
pass
def my_sq(x):
return x ** 2
my_sq(4)
assert my_sq(4) == 16
def avg_2(x, y):
"""Take the average of 2 numbers
"""
return (x + y) / 2
?avg_2
avg_2(10, 20)
import pandas as pd
# create an example dataframe from scratch
dat = pd.DataFrame({
'a': [10, 20, 30],
'b': [20, 30, 40]
})
dat
# square the a column
# math computations are vectorized/broadcast
dat["a"] ** 2
# using our function on a single value
my_sq(4)
# apply our function for each value in a column
dat["a"].apply(my_sq)
def my_exp(x, e):
return x ** e
assert my_exp(2, 10) == 1024
# pass in other function parameters into apply
dat["a"].apply(my_exp, e=3)
# if we want to pass in the value into something that is not the first argument
# one way is to write a wrapper function that will pass into first argument
def flip_exp(e, x):
return my_exp(x, e)
# apply function that uses column values as the exponent
dat["a"].apply(flip_exp, x=3)
# instead of re-writing a new function
# you can use lambda to write on the fly
dat["a"].apply(lambda pizza: my_exp(3, pizza))
def print_me(x):
print(x)
# applying functions on entire dataframes
# will work column by column
dat.apply(print_me)
def avg_3(x, y, z):
return (x + y + z) / 3
assert avg_3(1, 3, 5) == 3
# the entire column of values will be passed into the FIRST argument
dat.apply(avg_3)
import numpy as np
# some functions will automatically take in a vector/series of values
def avg_3_apply(col):
return (np.mean(col))
dat.apply(avg_3_apply)
# or we have to re-write and parse out the column values
def avg_3_apply(col):
x = col[0]
y = col[1]
z = col[2]
return (x + y + z) / 3
dat.apply(avg_3_apply)
dat
# axis-1 will work row by row
# you usually do not want to do this
# since there are also performance issues associated with this
dat.apply(avg_3_apply, axis=1)
dat
# again, moth math operations are already broadcast/vectorized
(dat["a"] + dat["b"]) / 2
def avg_2_mod(x, y):
if (x == 20):
return np.NaN
else:
return (x + y) / 2
# what if we wanted to pass columns of values and have our function work element-wise?
avg_2_mod(dat["a"], dat["b"])
# np.vectorize is a function that takes another function as input
# and it will output a vectorized version of that input function
avg_2_mod_vec = np.vectorize(avg_2_mod)
avg_2_mod_vec(dat["a"], dat["b"])
# we can also use the @ decorator to do the vectorization
# during function definition
# this way we don't need to create a new function
@np.vectorize
def v_avg_2_mod(x, y):
if (x == 20):
return np.NaN
else:
return (x + y) / 2
v_avg_2_mod(dat["a"], dat["b"])
import numba
@numba.vectorize
def v_avg_2_mod_numba(x, y):
if (x == 20):
return np.NaN
else:
return (x + y) / 2
# timing our functions
%%timeit
(dat["a"] + dat["b"]) / 2
%%timeit
avg_2(dat['a'], dat['b'])
%%timeit
v_avg_2_mod(dat['a'], dat['b'])
%%timeit
v_avg_2_mod_numba(dat['a'].values, dat['b'].values)
```
| github_jupyter |
# Testing Configurations
The behavior of a program is not only governed by its data. The _configuration_ of a program – that is, the settings that govern the execution of a program on its (regular) input data, as set by options or configuration files – just as well influences behavior, and thus can and should bne tested. In this chapter, we explore how to systematically _test_ and _cover_ software configurations. By _automatically inferring configuration options_, we can apply these techniques out of the box, with no need for writing a grammar. Finally, we show how to systematically cover _combinations_ of configuration options, quickly detecting unwanted interferences.
**Prerequisites**
* You should have read the [chapter on grammars](Grammars.ipynb).
* You should have read the [chapter on grammar coverage](GrammarCoverageFuzzer.ipynb).
## Configuration Options
When we talk about the input to a program, we usually think of the _data_ it processes. This is also what we have been fuzzing in the past chapters – be it with [random input](Fuzzer.ipynb), [mutation-based fuzzing](MutationFuzzer.ipynb), or [grammar-based fuzzing](GrammarFuzzer.ipynb). However, programs typically have several input sources, all of which can and should be tested – and included in test generation.
One important source of input is the program's _configuration_ – that is, a set of inputs that typically is set once when beginning to process data and then stays constant while processing data, while the program is running, or even while the program is deployed. Such a configuration is frequently set in _configuration files_ (for instance, as key/value pairs); the most ubiquitous method for command-line tools, though, are _configuration options_ on the command line.
As an example, consider the `grep` utility to find textual patterns in files. The exact mode by which `grep` works is governed by a multitude of options, which can be listed by providing a `--help` option:
```
!grep --help
```
All these options need to be tested for whether they operate correctly. In security testing, any such option may also trigger a yet unknown vulnerability. Hence, such options can become _fuzz targets_ on their own. In this chapter, we analyze how to systematically test such options – and better yet, how to extract possible configurations right out of given program files, such that we do not have to specify anything.
## Options in Python
Let us stick to our common programming language here and examine how options are processed in Python. The `argparse` module provides an parser for command-line arguments (and options) with great functionality – and great complexity. You start by defining a parser (`argparse.ArgumentParser()`) to which individual arguments with various features are added, one after another. Additional parameters for each argument can specify the type (`type`) of the argument (say, integers or strings), or the number of arguments (`nargs`).
By default, arguments are stored under their name in the `args` object coming from `parse_args()` – thus, `args.integers` holds the `integers` arguments added earlier. Special actions (`actions`) allow to store specific values in given variables; the `store_const` action stores the given `const` in the attribute named by `dest`. The following example takes a number of integer arguments (`integers`) as well as an operator (`--sum`, `--min`, or `--max`) to be applied on these integers. The operators all store a function reference in the `accumulate` attribute, which is finally invoked on the integers parsed:
```
import argparse
def process_numbers(args=[]):
parser = argparse.ArgumentParser(description='Process some integers.')
parser.add_argument('integers', metavar='N', type=int, nargs='+',
help='an integer for the accumulator')
group = parser.add_mutually_exclusive_group(required=True)
group.add_argument('--sum', dest='accumulate', action='store_const',
const=sum,
help='sum the integers')
group.add_argument('--min', dest='accumulate', action='store_const',
const=min,
help='compute the minimum')
group.add_argument('--max', dest='accumulate', action='store_const',
const=max,
help='compute the maximum')
args = parser.parse_args(args)
print(args.accumulate(args.integers))
```
Here's how `process_numbers()` works. We can, for instance, invoke the `--min` option on the given arguments to compute the minimum:
```
process_numbers(["--min", "100", "200", "300"])
```
Or compute the sum of three numbers:
```
process_numbers(["--sum", "1", "2", "3"])
```
When defined via `add_mutually_exclusive_group()` (as above), options are mutually exclusive. Consequently, we can have only one operator:
```
import fuzzingbook_utils
from ExpectError import ExpectError
with ExpectError(print_traceback=False):
process_numbers(["--sum", "--max", "1", "2", "3"])
```
## A Grammar for Configurations
How can we test a system with several options? The easiest answer is to write a grammar for it. The grammar `PROCESS_NUMBERS_EBNF_GRAMMAR` reflects the possible combinations of options and arguments:
```
from Grammars import crange, srange, convert_ebnf_grammar, is_valid_grammar, START_SYMBOL, new_symbol
PROCESS_NUMBERS_EBNF_GRAMMAR = {
"<start>": ["<operator> <integers>"],
"<operator>": ["--sum", "--min", "--max"],
"<integers>": ["<integer>", "<integers> <integer>"],
"<integer>": ["<digit>+"],
"<digit>": crange('0', '9')
}
assert is_valid_grammar(PROCESS_NUMBERS_EBNF_GRAMMAR)
PROCESS_NUMBERS_GRAMMAR = convert_ebnf_grammar(PROCESS_NUMBERS_EBNF_GRAMMAR)
```
We can feed this grammar into our [grammar coverage fuzzer](GrammarCoverageFuzzer.ipynb) and have it cover one option after another:
```
from GrammarCoverageFuzzer import GrammarCoverageFuzzer
f = GrammarCoverageFuzzer(PROCESS_NUMBERS_GRAMMAR, min_nonterminals=10)
for i in range(3):
print(f.fuzz())
```
Of course, we can also invoke `process_numbers()` with these very arguments. To this end, we need to convert the string produced by the grammar back into a list of individual arguments, using `split()`:
```
f = GrammarCoverageFuzzer(PROCESS_NUMBERS_GRAMMAR, min_nonterminals=10)
for i in range(3):
args = f.fuzz().split()
print(args)
process_numbers(args)
```
In a similar way, we can define grammars for any program to be tested; as well as define grammars for, say, configuration files. Yet, the grammar has to be updated with every change to the program, which creates a maintenance burden. Given that the information required for the grammar is already all encoded in the program, the question arises: _Can't we go and extract configuration options right out of the program in the first place?_
## Mining Configuration Options
In this section, we try to extract option and argument information right out of a program, such that we do not have to specify a configuration grammar. The aim is to have a configuration fuzzer that works on the options and arguments of an arbitrary program, as long as it follows specific conventions for processing its arguments. In the case of Python programs, this means using the `argparse` module.
Our idea is as follows: We execute the given program up to the point where the arguments are actually parsed – that is, `argparse.parse_args()` is invoked. Up to this point, we track all calls into the argument parser, notably those calls that define arguments and options (`add_argument()`). From these, we construct the grammar.
### Tracking Arguments
Let us illustrate this approach with a simple experiment: We define a trace function (see [our chapter on coverage](Coverage.ipynb) for details) that is active while `process_numbers` is invoked. If we have a call to a method `add_argument`, we access and print out the local variables (which at this point are the arguments to the method).
```
import sys
import string
def traceit(frame, event, arg):
if event != "call":
return
method_name = frame.f_code.co_name
if method_name != "add_argument":
return
locals = frame.f_locals
print(method_name, locals)
```
What we get is a list of all calls to `add_argument()`, together with the method arguments passed:
```
sys.settrace(traceit)
process_numbers(["--sum", "1", "2", "3"])
sys.settrace(None)
```
From the `args` argument, we can access the individual options and arguments to be defined:
```
def traceit(frame, event, arg):
if event != "call":
return
method_name = frame.f_code.co_name
if method_name != "add_argument":
return
locals = frame.f_locals
print(locals['args'])
sys.settrace(traceit)
process_numbers(["--sum", "1", "2", "3"])
sys.settrace(None)
```
We see that each argument comes as a tuple with one (say, `integers` or `--sum`) or two members (`-h` and `--help`), which denote alternate forms for the same option. Our job will be to go through the arguments of `add_arguments()` and detect not only the names of options and arguments, but also whether they accept additional parameters, as well as the type of the parameters.
### A Grammar Miner for Options and Arguments
Let us now build a class that gathers all this information to create a grammar.
We use the `ParseInterrupt` exception to interrupt program execution after gathering all arguments and options:
```
class ParseInterrupt(Exception):
pass
```
The class `OptionGrammarMiner` takes an executable function for which the grammar of options and arguments is to be mined:
```
class OptionGrammarMiner(object):
def __init__(self, function, log=False):
self.function = function
self.log = log
```
The method `mine_ebnf_grammar()` is where everything happens. It creates a grammar of the form
```
<start> ::= <option>* <arguments>
<option> ::= <empty>
<arguments> ::= <empty>
```
in which the options and arguments will be collected. It then sets a trace function (see [our chapter on coverage](Coverage.ipynb) for details) that is active while the previously defined `function` is invoked. Raising `ParseInterrupt` (when `parse_args()` is invoked) ends execution.
```
class OptionGrammarMiner(OptionGrammarMiner):
OPTION_SYMBOL = "<option>"
ARGUMENTS_SYMBOL = "<arguments>"
def mine_ebnf_grammar(self):
self.grammar = {
START_SYMBOL: ["(" + self.OPTION_SYMBOL + ")*" + self.ARGUMENTS_SYMBOL],
self.OPTION_SYMBOL: [],
self.ARGUMENTS_SYMBOL: []
}
self.current_group = self.OPTION_SYMBOL
old_trace = sys.settrace(self.traceit)
try:
self.function()
except ParseInterrupt:
pass
sys.settrace(old_trace)
return self.grammar
def mine_grammar(self):
return convert_ebnf_grammar(self.mine_ebnf_grammar())
```
The trace function checks for four methods: `add_argument()` is the most important function, resulting in processing arguments; `frame.f_locals` again is the set of local variables, which at this point is mostly the arguments to `add_argument()`. Since mutually exclusive groups also have a method `add_argument()`, we set the flag `in_group` to differentiate.
Note that we make no specific efforts to differentiate between multiple parsers or groups; we simply assume that there is one parser, and at any point at most one mutually exclusive group.
```
class OptionGrammarMiner(OptionGrammarMiner):
def traceit(self, frame, event, arg):
if event != "call":
return
if "self" not in frame.f_locals:
return
self_var = frame.f_locals["self"]
method_name = frame.f_code.co_name
if method_name == "add_argument":
in_group = repr(type(self_var)).find("Group") >= 0
self.process_argument(frame.f_locals, in_group)
elif method_name == "add_mutually_exclusive_group":
self.add_group(frame.f_locals, exclusive=True)
elif method_name == "add_argument_group":
# self.add_group(frame.f_locals, exclusive=False)
pass
elif method_name == "parse_args":
raise ParseInterrupt
return None
```
The `process_arguments()` now analyzes the arguments passed and adds them to the grammar:
* If the argument starts with `-`, it gets added as an optional element to the `<option>` list
* Otherwise, it gets added to the `<argument>` list.
The optional `nargs` argument specifies how many arguments can follow. If it is a number, we add the appropriate number of elements to the grammar; if it is an abstract specifier (say, `+` or `*`), we use it directly as EBNF operator.
Given the large number of parameters and optional behavior, this is a somewhat messy function, but it does the job.
```
class OptionGrammarMiner(OptionGrammarMiner):
def process_argument(self, locals, in_group):
args = locals["args"]
kwargs = locals["kwargs"]
if self.log:
print(args)
print(kwargs)
print()
for arg in args:
self.process_arg(arg, in_group, kwargs)
class OptionGrammarMiner(OptionGrammarMiner):
def process_arg(self, arg, in_group, kwargs):
if arg.startswith('-'):
if not in_group:
target = self.OPTION_SYMBOL
else:
target = self.current_group
metavar = None
arg = " " + arg
else:
target = self.ARGUMENTS_SYMBOL
metavar = arg
arg = ""
if "nargs" in kwargs:
nargs = kwargs["nargs"]
else:
nargs = 1
param = self.add_parameter(kwargs, metavar)
if param == "":
nargs = 0
if isinstance(nargs, int):
for i in range(nargs):
arg += param
else:
assert nargs in "?+*"
arg += '(' + param + ')' + nargs
if target == self.OPTION_SYMBOL:
self.grammar[target].append(arg)
else:
self.grammar[target].append(arg)
```
The method `add_parameter()` handles possible parameters of options. If the argument has an `action` defined, it takes no parameter. Otherwise, we identify the type of the parameter (as `int` or `str`) and augment the grammar with an appropriate rule.
```
import inspect
class OptionGrammarMiner(OptionGrammarMiner):
def add_parameter(self, kwargs, metavar):
if "action" in kwargs:
# No parameter
return ""
type_ = "str"
if "type" in kwargs:
given_type = kwargs["type"]
# int types come as '<class int>'
if inspect.isclass(given_type) and issubclass(given_type, int):
type_ = "int"
if metavar is None:
if "metavar" in kwargs:
metavar = kwargs["metavar"]
else:
metavar = type_
self.add_type_rule(type_)
if metavar != type_:
self.add_metavar_rule(metavar, type_)
param = " <" + metavar + ">"
return param
```
The method `add_type_rule()` adds a rule for parameter types to the grammar. If the parameter is identified by a meta-variable (say, `N`), we add a rule for this as well to improve legibility.
```
class OptionGrammarMiner(OptionGrammarMiner):
def add_type_rule(self, type_):
if type_ == "int":
self.add_int_rule()
else:
self.add_str_rule()
def add_int_rule(self):
self.grammar["<int>"] = ["(-)?<digit>+"]
self.grammar["<digit>"] = crange('0', '9')
def add_str_rule(self):
self.grammar["<str>"] = ["<char>+"]
self.grammar["<char>"] = srange(
string.digits
+ string.ascii_letters
+ string.punctuation)
def add_metavar_rule(self, metavar, type_):
self.grammar["<" + metavar + ">"] = ["<" + type_ + ">"]
```
The method `add_group()` adds a new mutually exclusive group to the grammar. We define a new symbol (say, `<group>`) for the options added to the group, and use the `required` and `exclusive` flags to define an appropriate expansion operator. The group is then prefixed to the grammar, as in
```
<start> ::= <group><option>* <arguments>
<group> ::= <empty>
```
and filled with the next calls to `add_argument()` within the group.
```
class OptionGrammarMiner(OptionGrammarMiner):
def add_group(self, locals, exclusive):
kwargs = locals["kwargs"]
if self.log:
print(kwargs)
required = kwargs.get("required", False)
group = new_symbol(self.grammar, "<group>")
if required and exclusive:
group_expansion = group
if required and not exclusive:
group_expansion = group + "+"
if not required and exclusive:
group_expansion = group + "?"
if not required and not exclusive:
group_expansion = group + "*"
self.grammar[START_SYMBOL][0] = group_expansion + \
self.grammar[START_SYMBOL][0]
self.grammar[group] = []
self.current_group = group
```
That's it! With this, we can now extract the grammar from our `process_numbers()` program. Turning on logging again reveals the variables we draw upon.
```
miner = OptionGrammarMiner(process_numbers, log=True)
ebnf_grammar = miner.mine_ebnf_grammar()
```
The grammar properly identifies the group found:
```
ebnf_grammar["<start>"]
ebnf_grammar["<group>"]
```
It also identifies a `--help` option provided not by us, but by the `argparse` module:
```
ebnf_grammar["<option>"]
```
The grammar also correctly identifies the types of the arguments:
```
ebnf_grammar["<arguments>"]
ebnf_grammar["<integers>"]
```
The rules for `int` are set as defined by `add_int_rule()`
```
ebnf_grammar["<int>"]
```
We can take this grammar and convert it to BNF, such that we can fuzz with it right away:
```
assert is_valid_grammar(ebnf_grammar)
grammar = convert_ebnf_grammar(ebnf_grammar)
assert is_valid_grammar(grammar)
f = GrammarCoverageFuzzer(grammar)
for i in range(10):
print(f.fuzz())
```
Each and every invocation adheres to the rules as set forth in the `argparse` calls. By mining options and arguments from existing programs, we can now fuzz these options out of the box – without having to specify a grammar.
## Testing Autopep8
Let us try out the option grammar miner on real-world Python programs. `autopep8` is a tool that automatically converts Python code to the [PEP 8 Style Guide for Python Code](https://www.python.org/dev/peps/pep-0008/). (Actually, all Python code in this book runs through `autopep8` during production.) `autopep8` offers a wide range of options, as can be seen by invoking it with `--help`:
```
!autopep8 --help
```
### Autopep8 Setup
We want to systematically test these options. In order to deploy our configuration grammar miner, we need to find the source code of the executable:
```
import os
def find_executable(name):
for path in os.get_exec_path():
qualified_name = os.path.join(path, name)
if os.path.exists(qualified_name):
return qualified_name
return None
autopep8_executable = find_executable("autopep8")
assert autopep8_executable is not None
autopep8_executable
```
Next, we build a function that reads the contents of the file and executes it.
```
def autopep8():
executable = find_executable("autopep8")
# First line has to contain "/usr/bin/env python" or like
first_line = open(executable).readline()
assert first_line.find("python") >= 0
contents = open(executable).read()
exec(contents)
```
### Mining an Autopep8 Grammar
We can use the `autopep8()` function in our grammar miner:
```
autopep8_miner = OptionGrammarMiner(autopep8)
```
and extract a grammar for it:
```
autopep8_ebnf_grammar = autopep8_miner.mine_ebnf_grammar()
```
This works because here, `autopep8` is not a separate process (and a separate Python interpreter), but we run the `autopep8()` function (and the `autopep8` code) in our current Python interpreter – up to the call to `parse_args()`, where we interrupt execution again. At this point, the `autopep8` code has done nothing but setting up the argument parser – which is what we are interested in.
The grammar options mined reflect precisely the options seen when providing `--help`:
```
print(autopep8_ebnf_grammar["<option>"])
```
Metavariables like `<n>` or `<line>` are placeholders for integers. We assume all metavariables of the same name have the same type:
```
autopep8_ebnf_grammar["<line>"]
```
The grammar miner has inferred that the argument to `autopep8` is a list of files:
```
autopep8_ebnf_grammar["<arguments>"]
```
which in turn all are strings:
```
autopep8_ebnf_grammar["<files>"]
```
As we are only interested in testing options, not arguments, we fix the arguments to a single mandatory input. (Otherwise, we'd have plenty of random file names generated.)
```
autopep8_ebnf_grammar["<arguments>"] = [" <files>"]
autopep8_ebnf_grammar["<files>"] = ["foo.py"]
assert is_valid_grammar(autopep8_ebnf_grammar)
```
### Creating Autopep8 Options
Let us now use the inferred grammar for fuzzing. Again, we convert the EBNF grammar into a regular BNF grammar:
```
autopep8_grammar = convert_ebnf_grammar(autopep8_ebnf_grammar)
assert is_valid_grammar(autopep8_grammar)
```
And we can use the grammar for fuzzing all options:
```
f = GrammarCoverageFuzzer(autopep8_grammar, max_nonterminals=4)
for i in range(20):
print(f.fuzz())
```
Let us apply these options on the actual program. We need a file `foo.py` that will serve as input:
```
def create_foo_py():
open("foo.py", "w").write("""
def twice(x = 2):
return x + x
""")
create_foo_py()
print(open("foo.py").read(), end="")
```
We see how `autopep8` fixes the spacing:
```
!autopep8 foo.py
```
Let us now put things together. We define a `ProgramRunner` that will run the `autopep8` executable with arguments coming from the mined `autopep8` grammar.
```
from Fuzzer import ProgramRunner
```
Running `autopep8` with the mined options reveals a surprising high number of passing runs. (We see that some options depend on each other or are mutually exclusive, but this is handled by the program logic, not the argument parser, and hence out of our scope.) The `GrammarCoverageFuzzer` ensures that each option is tested at least once. (Digits and letters, too, by the way.)
```
f = GrammarCoverageFuzzer(autopep8_grammar, max_nonterminals=5)
for i in range(20):
invocation = "autopep8" + f.fuzz()
print("$ " + invocation)
args = invocation.split()
autopep8 = ProgramRunner(args)
result, outcome = autopep8.run()
if result.stderr != "":
print(result.stderr, end="")
```
Our `foo.py` file now has been formatted in place a number of times:
```
print(open("foo.py").read(), end="")
```
We don't need it anymore, so we clean up things:
```
import os
os.remove("foo.py")
```
## Classes for Fuzzing Configuration Options
Let us now create reusable classes that we can use for testing arbitrary programs. (Okay, make that "arbitrary programs that are written in Python and use the `argparse` module to process command-line arguments.")
The class `OptionRunner` is a subclass of `ProgramRunner` that takes care of automatically determining the grammar, using the same steps we used for `autopep8`, above.
```
class OptionRunner(ProgramRunner):
def __init__(self, program, arguments=None):
if isinstance(program, str):
self.base_executable = program
else:
self.base_executable = program[0]
self.find_contents()
self.find_grammar()
if arguments is not None:
self.set_arguments(arguments)
super().__init__(program)
```
First, we find the contents of the Python executable:
```
class OptionRunner(OptionRunner):
def find_contents(self):
self._executable = find_executable(self.base_executable)
first_line = open(self._executable).readline()
assert first_line.find("python") >= 0
self.contents = open(self._executable).read()
def invoker(self):
exec(self.contents)
def executable(self):
return self._executable
```
Next, we determine the grammar using the `OptionGrammarMiner` class:
```
class OptionRunner(OptionRunner):
def find_grammar(self):
miner = OptionGrammarMiner(self.invoker)
self._ebnf_grammar = miner.mine_ebnf_grammar()
def ebnf_grammar(self):
return self._ebnf_grammar
def grammar(self):
return convert_ebnf_grammar(self._ebnf_grammar)
```
The two service methods `set_arguments()` and `set_invocation()` help us to change the arguments and program, respectively.
```
class OptionRunner(OptionRunner):
def set_arguments(self, args):
self._ebnf_grammar["<arguments>"] = [" " + args]
def set_invocation(self, program):
self.program = program
```
We can instantiate the class on `autopep8` and immediately get the grammar:
```
autopep8_runner = OptionRunner("autopep8", "foo.py")
print(autopep8_runner.ebnf_grammar()["<option>"])
```
An `OptionFuzzer` interacts with the given `OptionRunner` to obtain its grammar, which is then passed to its `GrammarCoverageFuzzer` superclass.
```
class OptionFuzzer(GrammarCoverageFuzzer):
def __init__(self, runner, *args, **kwargs):
assert issubclass(type(runner), OptionRunner)
self.runner = runner
grammar = runner.grammar()
super().__init__(grammar, *args, **kwargs)
```
When invoking `run()`, the `OptionFuzzer` creates a new invocation (using `fuzz()` from its grammar) and runs the now given (or previously set) runner with the arguments from the grammar. Note that the runner specified in `run()` can differ from the one set during initialization; this allows for mining options from one program and applying it in another context.
```
class OptionFuzzer(OptionFuzzer):
def run(self, runner=None, inp=""):
if runner is None:
runner = self.runner
assert issubclass(type(runner), OptionRunner)
invocation = runner.executable() + " " + self.fuzz()
runner.set_invocation(invocation.split())
return runner.run(inp)
```
### Example: Autopep8
Let us apply this on the `autopep8` runner:
```
autopep8_fuzzer = OptionFuzzer(autopep8_runner, max_nonterminals=5)
for i in range(3):
print(autopep8_fuzzer.fuzz())
```
We can now systematically test `autopep8` with these classes:
```
autopep8_fuzzer.run(autopep8_runner)
```
### Example: MyPy
We can extract options for the `mypy` static type checker for Python:
```
assert find_executable("mypy") is not None
mypy_runner = OptionRunner("mypy", "foo.py")
print(mypy_runner.ebnf_grammar()["<option>"])
mypy_fuzzer = OptionFuzzer(mypy_runner, max_nonterminals=5)
for i in range(10):
print(mypy_fuzzer.fuzz())
```
### Example: Notedown
Here's the configuration options for the `notedown` Notebook to Markdown converter:
```
assert find_executable("notedown") is not None
notedown_runner = OptionRunner("notedown")
print(notedown_runner.ebnf_grammar()["<option>"])
notedown_fuzzer = OptionFuzzer(notedown_runner, max_nonterminals=5)
for i in range(10):
print(notedown_fuzzer.fuzz())
```
## Combinatorial Testing
Our `CoverageGrammarFuzzer` does a good job in covering each and every option at least once, which is great for systematic testing. However, as we also can see in our examples above, some options require each other, while others interfere with each other. What we should do as good testers is not only to cover every option individually, but also _combinations_ of options.
The Python `itertools` module gives us means to create combinations from lists. We can, for instance, take the `notedown` options and create a list of all pairs.
```
from itertools import combinations
option_list = notedown_runner.ebnf_grammar()["<option>"]
pairs = list(combinations(option_list, 2))
```
There's quite a number of pairs:
```
len(pairs)
print(pairs[:20])
```
Testing every such pair of options frequently suffices to cover all interferences between options. (Programs rarely have conditions involving three or more configuration settings.) To this end, we _change_ the grammar from having a list of options to having a list of _option pairs_, such that covering these will automatically cover all pairs.
We create a function `pairwise()` that takes a list of options as occurring in our grammar and returns a list of _pairwise options_ – that is, our original options, but concatenated.
```
def pairwise(option_list):
return [option_1 + option_2
for (option_1, option_2) in combinations(option_list, 2)]
```
Here's the first 20 pairs:
```
print(pairwise(option_list)[:20])
```
The new grammar `pairwise_notedown_grammar` is a copy of the `notedown` grammar, but with the list of options replaced with the above pairwise option list.
```
from copy import deepcopy
notedown_grammar = notedown_runner.grammar()
pairwise_notedown_grammar = deepcopy(notedown_grammar)
pairwise_notedown_grammar["<option>"] = pairwise(notedown_grammar["<option>"])
assert is_valid_grammar(pairwise_notedown_grammar)
```
Using the "pairwise" grammar to fuzz now covers one pair after another:
```
notedown_fuzzer = GrammarCoverageFuzzer(
pairwise_notedown_grammar, max_nonterminals=4)
for i in range(10):
print(notedown_fuzzer.fuzz())
```
Can we actually test all combinations of options? Not in practice, as the number of combinations quickly grows as the length increases. It decreases again as the the number of options reach the maximum (with 20 options, there is only 1 combination involving _all_ options), but the absolute numbers are still staggering:
```
for combination_length in range(1, 20):
tuples = list(combinations(option_list, combination_length))
print(combination_length, len(tuples))
```
Formally, the number of combinations of length $k$ in a set of options of length $n$ is the binomial coefficient
$$
{n \choose k} = \frac{n!}{k!(n - k)!}
$$
which for $k = 2$ (all pairs) gives us
$$
{n \choose 2} = \frac{n!}{2(n - 2)!} = n \times (n - 1)
$$
For `autopep8` with its 29 options...
```
len(autopep8_runner.ebnf_grammar()["<option>"])
```
... we thus need 812 tests to cover all pairs:
```
len(autopep8_runner.ebnf_grammar()["<option>"]) * \
(len(autopep8_runner.ebnf_grammar()["<option>"]) - 1)
```
For `mypy` with its 110 options, though, we already end up with 11,990 tests to be conducted:
```
len(mypy_runner.ebnf_grammar()["<option>"])
len(mypy_runner.ebnf_grammar()["<option>"]) * \
(len(mypy_runner.ebnf_grammar()["<option>"]) - 1)
```
Even if each pair takes a second to run, we'd still be done in three hours of testing, though.
If your program has more options that you all want to get covered in combinations, it is advisable that you limit the number of configurations further – for instance by limiting combinatorial testing to those combinations that possibly can interact with each other; and covering all other (presumably orthogonal) options individually.
This mechanism of creating configurations by extending grammars can be easily extended to other configuration targets. One may want to explore a greater number of configurations, or expansions in specific contexts. The exercises, below, have a number of options ready for you.
## Lessons Learned
* Besides regular input data, program _configurations_ make an important testing target.
* For a given program using a standard library to parse command-line options and arguments, one can automatically extract these and convert them into a grammar.
* To cover not only single options, but combinations of options, one can expand the grammar to cover all pairs, or come up with even more ambitious targets.
## Next Steps
If you liked the idea of mining a grammar from a program, do not miss:
* [how to mine grammars for input data](GrammarMiner.ipynb)
Our next steps in the book focus on:
* [how to parse and recombine inputs](Parser.ipynb)
* [how to assign weights and probabilities to specific productions](ProbabilisticGrammarFuzzer.ipynb)
* [how to simply inputs that cause a failure](Reducer.ipynb)
## Background
Although configuration data is just as likely to cause failures as other input data, it has received relatively little attention in test generation – possibly because, unlike "regular" input data, configuration data is not so much under control of external parties, and because, again unlike regular data, there is little variance in configurations. Creating models for software configurations and using these models for testing is commonplace, as is the idea of pairwise testing. For an overview, see \cite{Pezze2008}; for a discussion and comparison of state-of-the-art techniques, see \cite{Petke2015}.
More specifically, \cite{Sutton2007} also discuss techniques to systematically cover command-line options. Dai et al. \cite{Dai2010} apply configuration fuzzing by changing variables associated with configuration files.
## Exercises
### Exercise 1: Configuration Files
Besides command-line options, a second important source of configurations are _configuration files_. In this exercise, we will consider the very simple configuration language provided by the Python `ConfigParser` module, which is very similar to what is found in Microsoft Windows _.ini_ files.
The following example for a `ConfigParser` input file stems right from [the ConfigParser documentation](https://docs.python.org/3/library/configparser.html):
```
[DEFAULT]
ServerAliveInterval = 45
Compression = yes
CompressionLevel = 9
ForwardX11 = yes
[bitbucket.org]
User = hg
[topsecret.server.com]
Port = 50022
ForwardX11 = no
```
The above `ConfigParser` file can be created programmatically:
```
import configparser
config = configparser.ConfigParser()
config['DEFAULT'] = {'ServerAliveInterval': '45',
'Compression': 'yes',
'CompressionLevel': '9'}
config['bitbucket.org'] = {}
config['bitbucket.org']['User'] = 'hg'
config['topsecret.server.com'] = {}
topsecret = config['topsecret.server.com']
topsecret['Port'] = '50022' # mutates the parser
topsecret['ForwardX11'] = 'no' # same here
config['DEFAULT']['ForwardX11'] = 'yes'
with open('example.ini', 'w') as configfile:
config.write(configfile)
with open('example.ini') as configfile:
print(configfile.read(), end="")
```
and be read in again:
```
config = configparser.ConfigParser()
config.read('example.ini')
topsecret = config['topsecret.server.com']
topsecret['Port']
```
#### Part 1: Read Configuration
Using `configparser`, create a program reading in the above configuration file and accessing the individual elements.
#### Part 2: Create a Configuration Grammar
Design a grammar that will automatically create configuration files suitable for your above program. Fuzz your program with it.
#### Part 3: Mine a Configuration Grammar
By dynamically tracking the individual accesses to configuration elements, you can again extract a basic grammar from the execution. To this end, create a subclass of `ConfigParser` with a special method `__getitem__`:
```
class TrackingConfigParser(configparser.ConfigParser):
def __getitem__(self, key):
print("Accessing", repr(key))
return super().__getitem__(key)
```
For a `TrackingConfigParser` object `p`, `p.__getitem__(key)` will be invoked whenever `p[key]` is accessed:
```
tracking_config_parser = TrackingConfigParser()
tracking_config_parser.read('example.ini')
section = tracking_config_parser['topsecret.server.com']
```
Using `__getitem__()`, as above, implement a tracking mechanism that, while your program accesses the read configuration, automatically saves options accessed and values read. Create a prototype grammar from these values; use it for fuzzing.
At the end, don't forget to clean up:
```
import os
os.remove("example.ini")
```
**Solution.** Left to the reader. Enjoy!
### Exercise 2: C Option Fuzzing
In C programs, the `getopt()` function are frequently used to process configuration options. A call
```
getopt(argc, argv, "bf:")
```
indicates that the program accepts two options `-b` and `-f`, with `-f` taking an argument (as indicated by the following colon).
#### Part 1: Getopt Fuzzing
Write a framework which, for a given C program, automatically extracts the argument to `getopt()` and derives a fuzzing grammar for it. There are multiple ways to achieve this:
1. Scan the program source code for occurrences of `getopt()` and return the string passed. (Crude, but should frequently work.)
2. Insert your own implementation of `getopt()` into the source code (effectively replacing `getopt()` from the runtime library), which outputs the `getopt()` argument and exits the program. Recompile and run.
3. (Advanced.) As above, but instead of changing the source code, hook into the _dynamic linker_ which at runtime links the program with the C runtime library. Set the library loading path (on Linux and Unix, this is the `LD_LIBRARY_PATH` environment variable) such that your own version of `getopt()` is linked first, and the regular libraries later. Executing the program (without recompiling) should yield the desired result.
Apply this on `grep` and `ls`; report the resulting grammars and results.
**Solution.** Left to the reader. Enjoy hacking!
#### Part 2: Fuzzing Long Options in C
Same as Part 1, but also hook into the GNU variant `getopt_long()`, which accepts "long" arguments with double dashes such as `--help`. Note that method 1, above, will not work here, since the "long" options are defined in a separately defined structure.
**Solution.** Left to the reader. Enjoy hacking!
### Exercise 3: Expansions in Context
In our above option configurations, we have multiple symbols which all expand to the same integer. For instance, the `--line-range` option of `autopep8` takes two `<line>` parameters which both expand into the same `<int>` symbol:
```
<option> ::= ... | --line-range <line> <line> | ...
<line> ::= <int>
<int> ::= (-)?<digit>+
<digit> ::= 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9
```
```
autopep8_runner.ebnf_grammar()["<line>"]
autopep8_runner.ebnf_grammar()["<int>"]
autopep8_runner.ebnf_grammar()["<digit>"]
```
Once the `GrammarCoverageFuzzer` has covered all variations of `<int>` (especially by covering all digits) for _one_ option, though, it will no longer strive to achieve such coverage for the next option. Yet, it could be desirable to achieve such coverage for each option separately.
One way to achieve this with our existing `GrammarCoverageFuzzer` is again to change the grammar accordingly. The idea is to _duplicate_ expansions – that is, to replace an expansion of a symbol $s$ with a new symbol $s'$ whose definition is duplicated from $s$. This way, $s'$ and $s$ are separate symbols from a coverage point of view and would be independently covered.
As an example, consider again the above `--line-range` option. If we want our tests to independently cover all elements of the two `<line>` parameters, we can duplicate the second `<line>` expansion into a new symbol `<line'>` with subsequent duplicated expansions:
```
<option> ::= ... | --line-range <line> <line'> | ...
<line> ::= <int>
<line'> ::= <int'>
<int> ::= (-)?<digit>+
<int'> ::= (-)?<digit'>+
<digit> ::= 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9
<digit'> ::= 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9
```
Design a function `inline(grammar, symbol)` that returns a duplicate of `grammar` in which every occurrence of `<symbol>` and its expansions become separate copies. The above grammar could be a result of `inline(autopep8_runner.ebnf_grammar(), "<line>")`.
When copying, expansions in the copy should also refer to symbols in the copy. Hence, when expanding `<int>` in
```<int> ::= <int><digit>```
make that
```<int> ::= <int><digit>
<int'> ::= <int'><digit'>
```
(and not `<int'> ::= <int><digit'>` or `<int'> ::= <int><digit>`).
Be sure to add precisely one new set of symbols for each occurrence in the original grammar, and not to expand further in the presence of recursion.
**Solution.** Again, left to the reader. Enjoy!
| github_jupyter |
# NAPARI visualization of UNet Training Data
You can use this notebook to view, modified and save out training data for UNet models
Labels:
+ 0 - background
+ 1 - GFP/Phase
+ 2 - RFP
Extra key bindings:
+ 'w' - calculate weightmap
+ 's' - save labels
+ 'o' - output all weightmaps and metadata for tfrecord creation
+ '\>' - grow the label under the mouse cursor
+ '\<' - shrink the label under the mouse cursor
```
import os
import re
import enum
import json
import napari
from skimage import io
import numpy as np
from scipy import ndimage
from scipy.ndimage.morphology import distance_transform_edt
from scipy.ndimage import gaussian_filter
@enum.unique
class Channels(enum.Enum):
BRIGHTFIELD = 0
GFP = 1
RFP = 2
IRFP = 3
PHASE = 4
WEIGHTS = 98
MASK = 99
```
---
## Set up the data path and channel used
```
# DATA_PATH = '/home/arl/Dropbox/Data/TrainingData/UNet_training_Ras'
# # DATA_PATH = '/home/arl/Dropbox/Data/TrainingData/UNet_training_Scribble'
# channels = [Channels.GFP, Channels.RFP]
DATA_PATH = '/Volumes/lowegrp/TrainingData/UNet_training_Brightfield_v2/'
channels = [Channels.BRIGHTFIELD]
WEIGHT_AMPLITUDE = 10.
```
---
## Code
```
def strip_modified_filename(filename):
if filename.endswith('.modified.tif'):
stripped_fn = filename[:-len('.modified.tif')]
return stripped_fn
return filename
def make_folder(foldername):
if os.path.exists(foldername):
return
os.mkdir(foldername)
def file_root(filename):
FILENAME_PATTERN = r'([a-zA-Z0-9]+)_([a-zA-Z0-9]+)_*.tif'
grps = re.search(FILENAME_PATTERN, filename)
return grps
def load_training_data(pth, channels=[Channels.GFP, Channels.RFP]):
""" load training data for visualisation with napari """
all_channels = [Channels.MASK]+channels
# find the sets and sort them
sets = [f for f in os.listdir(pth) if os.path.isdir(os.path.join(pth, f))]
sets.sort(key = lambda s: int(s[3:]))
def set_filename_format(filename):
grps = file_root(filename)
if grps.group(1) in [c.name.lower() for c in all_channels]:
FILENAME_FORMAT = 2
else:
FILENAME_FORMAT = 1
def filename_formatter(filename, channel):
assert(channel in [c.name.lower() for c in all_channels])
grps = file_root(filename)
return '{}_{}.tif'.format(*[grps.group(FILENAME_FORMAT), channel])
# return '{}_{}.tif'.format(*[channel, grps.group(FILENAME_FORMAT)])
return filename_formatter
files = {k:{'files':[], 'data':[], 'sets':[], 'path':[]} for k in all_channels}
for s in sets:
# root_folders
l_root = os.path.join(pth, s, 'labels')
# check that this folder exists
if not os.path.exists(l_root):
raise IOError('{} does not exist. Do you need to rename label -> labels?'.format(l_root))
# get the training label files
label_files = [f for f in os.listdir(l_root) if f.endswith('.tif')]
# sort to remove unmodified files and replace with the modified files
unmodified_files, modified_files = [], []
for i, f in enumerate(label_files):
if f.endswith('.modified.tif'):
modified_files.append(strip_modified_filename(f))
else:
unmodified_files.append(f)
unmodified_files = list(set(unmodified_files).difference(set(modified_files)))
label_files = unmodified_files + [f+'.modified.tif' for f in modified_files]
# print label_files
fnfmt = set_filename_format(label_files[0])
files[Channels.MASK]['path'] += [s+'/labels/'+f for f in label_files]
files[Channels.MASK]['files'] += [strip_modified_filename(f) for f in label_files]
files[Channels.MASK]['data'] += [io.imread(os.path.join(l_root, f)) for f in label_files]
files[Channels.MASK]['sets'] += [s] * len(label_files)
for channel in channels:
cfiles = [fnfmt(l, channel.name.lower()) for l in label_files]
files[channel]['path'] += [s+'/'+channel.name.lower()+'/'+f for f in cfiles]
files[channel]['files'] += cfiles
files[channel]['data'] += [io.imread(os.path.join(pth, s, channel.name.lower(), f)) for f in cfiles]
files[channel]['sets'] += [s] * len(label_files)
# now make image stacks
for channel in files.keys():
for i, im in enumerate(files[channel]['data']):
print(channel, files[channel]['path'][i], im.shape, im.dtype)
files[channel]['data'] = np.stack(files[channel]['data'], axis=0)
return files
data = load_training_data(DATA_PATH, channels)
def normalize_images(stack):
normed = stack.astype(np.float32)
for i in range(stack.shape[0]):
# normed[i,...] = (normed[i,...]-np.mean(normed[i,...])) / np.std(normed[i,...])
c = normed[i,...]
p_lo = np.percentile(c,5)
p_hi = np.percentile(c,99)
normed[i,...] = np.clip((c - p_lo) / p_hi, 0., 1.)
return normed
def bounding_boxes(seg):
lbl, nlbl = ndimage.label(seg)
class_label, _, minxy, maxxy = ndimage.extrema(seg, lbl, index=np.arange(1, nlbl+1))
return class_label, minxy, maxxy
seg = np.zeros(data[channels[0]]['data'].shape, dtype=np.uint8)
mask = data[Channels.MASK]['data']
if mask.ndim == 3:
seg = mask > 0
elif mask.ndim == 4:
seg[mask[:,0,...]>0] = 1
seg[mask[:,1,...]>0] = 2
def convert_to_mask(labels, unique_labels=range(1,len(channels)+1)):
print(unique_labels)
seg = np.zeros((len(unique_labels),)+labels.shape, dtype=np.uint8)
for i,l in enumerate(unique_labels):
seg[i,...] = labels==l
return np.squeeze(seg)
def save_labels(viewer):
# get the current image
current_slice = viewer.layers[viewer.active_layer].coordinates[0]
source_set = data[Channels.MASK]['sets'][current_slice]
source_file = data[Channels.MASK]['files'][current_slice]
source_fn = os.path.join(source_set, 'labels', source_file)
# get the current layer
current_labels = viewer.layers['labels'].data[current_slice,...]
current_mask = convert_to_mask(current_labels)
# write out the modified segmentation mask
new_file = os.path.join(DATA_PATH, source_fn+'.modified.tif')
print(new_file)
io.imsave(new_file, current_mask.astype('uint8'))
print(current_slice, current_labels.shape, new_file)
weightmaps = np.zeros((seg.shape), dtype=np.float32)
def calculate_weightmaps(viewer, weight=WEIGHT_AMPLITUDE, current_slice=0):
# get the current layer and make it binary
mask = viewer.layers['labels'].data[current_slice,...].astype(np.bool)
wmap = weight * (gaussian_filter(mask.astype(np.float32), sigma=5.) - gaussian_filter(mask.astype(np.float32), sigma=.1))
# normalize it
wmap += 1.
wmap[mask] = 1.
viewer.layers['weightmaps'].data[current_slice,...] = wmap.astype(np.float32)
viewer.layers['weightmaps'].contrast_limits = (np.min(wmap), np.max(wmap))
viewer.layers['weightmaps'].visible = True
return wmap
def grow_shrink_label(viewer, grow=True):
# get the current image
current_slice = viewer.layers[viewer.active_layer].coordinates[0]
current_labels = viewer.layers['labels'].data[current_slice,...]
cursor_coords = [int(p) for p in viewer.layers[viewer.active_layer].position]
labelled,_ = ndimage.label(current_labels.astype(np.bool))
real_label = current_labels[cursor_coords[0], cursor_coords[1]]
if real_label < 1: return
print(real_label)
mask = labelled == labelled[cursor_coords[0], cursor_coords[1]]
if grow:
mask = ndimage.morphology.binary_dilation(mask, iterations=3)
else:
current_labels[mask] = 0
mask = ndimage.morphology.binary_erosion(mask, iterations=3)
current_labels[mask] = real_label
viewer.layers['labels'].data[current_slice,...] = current_labels
viewer.layers['labels']._set_view_slice()
# start napari
with napari.gui_qt():
viewer = napari.Viewer()
if Channels.GFP in data:
gfp = normalize_images(data[Channels.GFP]['data'])
viewer.add_image(gfp, name='GFP', colormap='green', contrast_limits=(0.,1.))
if Channels.RFP in data:
rfp = normalize_images(data[Channels.RFP]['data'])
viewer.add_image(rfp, name='RFP', colormap='magenta', contrast_limits=(0.,1.))
viewer.layers['RFP'].blending = 'additive'
if Channels.PHASE in data:
phase = normalize_images(data[Channels.PHASE]['data'])
viewer.add_image(phase, name='Phase', colormap='gray')
if Channels.BRIGHTFIELD in data:
bf = normalize_images(data[Channels.BRIGHTFIELD]['data'])
viewer.add_image(bf, name='Brightfield', colormap='gray')
viewer.add_image(weightmaps, name='weightmaps', colormap='plasma', visible=False)
viewer.add_labels(seg, name='labels')
viewer.layers['labels'].opacity = 0.4
viewer.layers['weightmaps'].blending = 'additive'
@viewer.bind_key('s')
def k_save_labels(viewer):
save_labels(viewer)
@viewer.bind_key('w')
def k_calculate_weightmaps(viewer):
current_slice = viewer.layers[viewer.active_layer].coordinates[0]
calculate_weightmaps(viewer, current_slice=current_slice)
@viewer.bind_key('<')
def k_shrink_label(viewer):
print('shrink label')
grow_shrink_label(viewer, grow=False)
@viewer.bind_key('>')
def k_grow_label(viewer):
print('grow label')
grow_shrink_label(viewer, grow=True)
@viewer.bind_key('o')
def k_output(viewer):
print('output all with metadata')
data[Channels.WEIGHTS] = {'files':[], 'sets':[], 'path':[]}
for i in range(viewer.layers['weightmaps'].data.shape[0]):
wmap = calculate_weightmaps(viewer, current_slice=i)
source_set = data[Channels.MASK]['sets'][i]
source_file = data[Channels.MASK]['files'][i]
fn = file_root(source_file).group(1)
weight_folder = os.path.join(DATA_PATH, source_set, 'weights')
make_folder(weight_folder)
weight_fn = '{}_weights.tif'.format(fn)
io.imsave(os.path.join(weight_folder, weight_fn), wmap.astype(np.float32))
data[Channels.WEIGHTS]['files'].append(weight_fn)
data[Channels.WEIGHTS]['sets'].append(source_set)
data[Channels.WEIGHTS]['path'].append('{}/weights/{}'.format(source_set, weight_fn))
# write out a JSON file with the data
jfn = os.path.join(DATA_PATH, 'training_metadata.json')
jdata = {}
for channel in data.keys():
jdata[channel.name.lower()] = data[channel]['path']
with open(jfn, 'w') as json_file:
json.dump(jdata, json_file, indent=2, separators=(',', ': '))
# # convert segmentation output labels to multichannel stacks
# p = '/Users/arl/Dropbox/Data/TrainingData/set12'
# files = [f for f in os.listdir(os.path.join(p,'labels')) if f.endswith('.tif')]
# for f in files:
# mask = io.imread(os.path.join(p, 'labels', f))
# print(mask.shape)
# gfp = mask==1
# rfp = mask==2
# new_mask = np.stack([gfp, rfp], axis=0)
# io.imsave(os.path.join(p,f), new_mask.astype('uint8'))
```
| github_jupyter |
<a href="https://colab.research.google.com/github/yassir-a-p/blender-on-colab/blob/main/blender-on-colab.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Check GPU device
```
!nvidia-smi --query-gpu=gpu_name,driver_version,memory.total --format=csv
```
#Setup
Specify your desired blender version
```
#@title Setup
#@markdown Please configure your setup
blender_version = 'blender2.92.0' #@param ["blender2.92.0", "blender2.91.2", "blender2.90.1", "blender2.83.13", "blender2.82a", "blender2.81a", "blender2.80", "blender2.79b"] {allow-input: false}
gpu_enabled = True #@param {type:"boolean"}
cpu_enabled = True #@param {type:"boolean"}
#@markdown ---
use_drive = True #@param {type:"boolean"}
```
## Google Drive
Run this code if you want to connect to your Google Drive (check **use_drive** option above)<br><br>
There will be a link and an empty field.<br>Click the link, it will open a new tab, choose the account you want to access its GDrive, then click **Allow**. Finally, copy the verification code and paste it in the field below, then press **[Enter]**.
```
if use_drive:
from google.colab import drive
drive.mount('/content/drive')
```
## Blend file setup
Specify your blend file and the output path. Make sure the folder for the output exist, otherwise Blender will fail to write the file.
```
#@markdown You can right click a file and click copy path
path_to_blend = '/content/drive/your-folder/your-blend-file.blend' #@param {type: "string"}
output_path = '/content/drive/your-folder/output/frame_###' #@param {type: "string"}
```
# **Installing Blender in the cloud**
Specifying download location for chosen blender version
```
if blender_version == "blender2.79b":
download_path="https://ftp.halifax.rwth-aachen.de/blender/release/Blender2.79/blender-2.79b-linux-glibc219-x86_64.tar.bz2"
elif blender_version == "blender2.80":
download_path="https://ftp.halifax.rwth-aachen.de/blender/release/Blender2.80/blender-2.80-linux-glibc217-x86_64.tar.bz2"
elif blender_version == "blender2.81a":
download_path="https://ftp.halifax.rwth-aachen.de/blender/release/Blender2.81/blender-2.81a-linux-glibc217-x86_64.tar.bz2"
elif blender_version == "blender2.82":
download_path="https://ftp.halifax.rwth-aachen.de/blender/release/Blender2.82/blender-2.82a-linux64.tar.xz"
elif blender_version == "blender2.83.13":
download_path="https://ftp.halifax.rwth-aachen.de/blender/release/Blender2.83/blender-2.83.13-linux64.tar.xz"
elif blender_version == "blender2.90.1":
download_path="https://ftp.halifax.rwth-aachen.de/blender/release/Blender2.90/blender-2.90.1-linux64.tar.xz"
elif blender_version == "blender2.91.2":
download_path="https://ftp.halifax.rwth-aachen.de/blender/release/Blender2.91/blender-2.91.2-linux64.tar.xz"
elif blender_version == "blender2.92.0":
download_path="https://ftp.halifax.rwth-aachen.de/blender/release/Blender2.92/blender-2.92.0-linux64.tar.xz"
```
Download, unpack, and move blender to designated location
```
!mkdir $blender_version
if blender_version == "blender2.79b" or "blender2.80" or "blender2.81a":
!wget -O '{blender_version}.tar.bz2' -nc $download_path
!tar -xf '{blender_version}.tar.bz2' -C ./$blender_version --strip-components=1
else:
!wget -O '{blender_version}.tar.xz' -nc $download_path
!tar xf '{blender_version}.tar.xz' -C ./$blender_version --strip-components=1
```
*This block is required as some weird behaviors with libtcmalloc appeared in the colab VM*
```
import os
os.environ["LD_PRELOAD"] = ""
!apt update
!apt remove libtcmalloc-minimal4
!apt install libtcmalloc-minimal4
os.environ["LD_PRELOAD"] = "/usr/lib/x86_64-linux-gnu/libtcmalloc_minimal.so.4.3.0"
!echo $LD_PRELOAD
```
GPU Dependencies
```
!apt install libboost-all-dev
!apt install libgl1-mesa-dev
!apt install libglu1-mesa libsm-dev
```
# **Render**
```
#@title **Render Setup**
#@markdown Set **whatToRender** to:<br>
#@markdown + **renderOneFrame** : if you only want to render a single frame. Specify the frame on startFrame field below<br>
#@markdown + **renderAllFrame** : if you want to render all the frames as already specified in your .blend file.<br>
#@markdown + **renderWithinRange** : if you want to render specific frames within certain range], set the _**startFrame**_ dan _**endFrame**_ below
whatToRender = 'renderOneFrame' #@param ["renderOneFrame", "renderAllFrame", "renderWithinRange"] {allow-input: false}
#@markdown ---
startFrame = 1 #@param {type: "integer"}
endFrame = 250 #@param {type: "integer"}
#@markdown ---
override_certain_settings = False #@param {type:"boolean"}
#@markdown Override the settings of samples, resolution, dan compression<br>
#@markdown True Samples count, Squared Samples is disabled
samples = 500 #@param {type:"integer"}
resolution_x = 1280 #@param {type:"integer"}
resolution_y = 720 #@param {type:"integer"}
compression_level = 100 #@param ["0", "25", "50", "75", "100"] {allow-input: false}
#@markdown **!!! IMPORTANT !!!**<br>
#@markdown Make sure to rerun the code if you modify things above (frames to render, samples, etc)!
# Generating script to override Samples, Resolution, dan Compression settings
override = "# Override Samples, Resolution, dan Compression settings.\n"+\
"True samples count, Squared Samples is disabled\n"+\
"import bpy\n"+\
"scene = bpy.context.scene\n"+\
"scene.cycles.use_square_samples = False\n"+\
"scene.cycles.samples = "+str(samples)+"\n"+\
"scene.render.resolution_x = "+str(resolution_x)+"\n"+\
"scene.render.resolution_y = "+str(resolution_y)+"\n"+\
"scene.render.image_settings.compression = "+str(compression_level)+"\n"
with open('override_others.py', 'w') as f:
f.write(override)
```
## Generating script
Required for Blender to use the GPU as expected
```
data = "import re\n"+\
"import bpy\n"+\
"scene = bpy.context.scene\n"+\
"scene.cycles.device = 'GPU'\n"+\
"prefs = bpy.context.preferences\n"+\
"prefs.addons['cycles'].preferences.get_devices()\n"+\
"cprefs = prefs.addons['cycles'].preferences\n"+\
"print(cprefs)\n"+\
"# Attempt to set GPU device types if available\n"+\
"for compute_device_type in ('CUDA', 'OPENCL', 'NONE'):\n"+\
" try:\n"+\
" cprefs.compute_device_type = compute_device_type\n"+\
" print('Device found',compute_device_type)\n"+\
" break\n"+\
" except TypeError:\n"+\
" pass\n"+\
"#for scene in bpy.data.scenes:\n"+\
"# scene.render.tile_x = 64\n"+\
"# scene.render.tile_y = 64\n"+\
"# Enable all CPU and GPU devices\n"+\
"for device in cprefs.devices:\n"+\
" if not re.match('intel', device.name, re.I):\n"+\
" print('Activating',device)\n"+\
" device.use = "+str(gpu_enabled)+"\n"+\
" else:\n"+\
" device.use = "+str(cpu_enabled)+"\n"
with open('setgpu.py', 'w') as f:
f.write(data)
```
## **Render Cycles with CUDA**
```
if override_certain_settings == False:
if whatToRender == 'renderOneFrame':
!sudo ./$blender_version/blender -b '{path_to_blend}' -P './setgpu.py' -noaudio -E CYCLES -o '{output_path}' -F 'PNG' -f {startFrame}
elif whatToRender == 'renderAllFrame':
!sudo ./$blender_version/blender -b '{path_to_blend}' -P './setgpu.py' -noaudio -E CYCLES -o '{output_path}' -F 'PNG' -a
elif whatToRender == 'renderWithinRange':
!sudo ./$blender_version/blender -b '{path_to_blend}' -P './setgpu.py' -noaudio -E CYCLES -o '{output_path}' -F 'PNG' -s {startFrame} -e {endFrame} -a
else:
if whatToRender == 'renderOneFrame':
!sudo ./$blender_version/blender -b '{path_to_blend}' -P './setgpu.py' -P './override_others.py' -noaudio -E CYCLES -o '{output_path}' -F 'PNG' -f {startFrame}
elif whatToRender == 'renderAllFrame':
!sudo ./$blender_version/blender -b '{path_to_blend}' -P './setgpu.py' -P './override_others.py' -noaudio -E CYCLES -o '{output_path}' -F 'PNG' -a
elif whatToRender == 'renderWithinRange':
!sudo ./$blender_version/blender -b '{path_to_blend}' -P './setgpu.py' -P './override_others.py' -noaudio -E CYCLES -o '{output_path}' -F 'PNG' -s {startFrame} -e {endFrame} -a
```
## **Render Cycles with OPTIX**
not using setgpu.py
```
if override_certain_settings == False:
if whatToRender == 'renderOneFrame':
!sudo ./$blender_version/blender -b '{path_to_blend}' -noaudio -E CYCLES -o '{output_path}' -F 'PNG' -f {startFrame} -- --cycles-device OPTIX
elif whatToRender == 'renderAllFrame':
!sudo ./$blender_version/blender -b '{path_to_blend}' -noaudio -E CYCLES -o '{output_path}' -F 'PNG' -a -- --cycles-device OPTIX
elif whatToRender == 'renderWithinRange':
!sudo ./$blender_version/blender -b '{path_to_blend}' -noaudio -E CYCLES -o '{output_path}' -F 'PNG' -s {startFrame} -e {endFrame} -a -- --cycles-device OPTIX
else:
if whatToRender == 'renderOneFrame':
!sudo ./$blender_version/blender -b '{path_to_blend}' -P './override_others.py' -noaudio -E CYCLES -o '{output_path}' -F 'PNG' -f {startFrame} -- --cycles-device OPTIX
elif whatToRender == 'renderAllFrame':
!sudo ./$blender_version/blender -b '{path_to_blend}' -P './override_others.py' -noaudio -E CYCLES -o '{output_path}' -F 'PNG' -a -- --cycles-device OPTIX
elif whatToRender == 'renderWithinRange':
!sudo ./$blender_version/blender -b '{path_to_blend}' -P './override_others.py' -noaudio -E CYCLES -o '{output_path}' -F 'PNG' -s {startFrame} -e {endFrame} -a -- --cycles-device OPTIX
```
---
# **APPENDICES**
## Copyright, License and Modifications Notice
Copyright 2021 Yassir A. P.
Copyright 2021 github.com/donmahallem
Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License.
You may obtain a copy of the License at
http://www.apache.org/licenses/LICENSE-2.0
This file has been modified to improve workflow, use updated Blender versions, add support for OPTIX rendering, and add options to override settings of Samples, Resolution, and Compression.
## Questions? Suggestions?
If you have some suggestions, request, or questions, you're welcome to discuss things on [the repo's Discussion page](https://github.com/yassir-a-p/blender-on-colab/discussions). I don't promise anything, but I will try to reply and maintain this in the long run.
| github_jupyter |
# Week 2: Implementing Callbacks in TensorFlow using the MNIST Dataset
In the course you learned how to do classification using Fashion MNIST, a data set containing items of clothing. There's another, similar dataset called MNIST which has items of handwriting -- the digits 0 through 9.
Write an MNIST classifier that trains to 99% accuracy or above, and does it without a fixed number of epochs -- i.e. you should stop training once you reach that level of accuracy. In the lecture you saw how this was done for the loss but here you will be using accuracy instead.
Some notes:
1. Given the architecture of the net, it should succeed in less than 10 epochs.
2. When it reaches 99% or greater it should print out the string "Reached 99% accuracy so cancelling training!" and stop training.
3. If you add any additional variables, make sure you use the same names as the ones used in the class. This is important for the function signatures (the parameters and names) of the callbacks.
```
import os
import tensorflow as tf
from tensorflow import keras
```
Begin by loading the data. A couple of things to notice:
- The file `mnist.npz` is already included in the current workspace under the `data` directory. By default the `load_data` from Keras accepts a path relative to `~/.keras/datasets` but in this case it is stored somewhere else, as a result of this, you need to specify the full path.
- `load_data` returns the train and test sets in the form of the tuples `(x_train, y_train), (x_test, y_test)` but in this exercise you will be needing only the train set so you can ignore the second tuple.
```
# Load the data
# Get current working directory
current_dir = os.getcwd()
# Append data/mnist.npz to the previous path to get the full path
data_path = os.path.join(current_dir, "data/mnist.npz")
# Discard test set
(x_train, y_train), _ = tf.keras.datasets.mnist.load_data(path=data_path)
# Normalize pixel values
x_train = x_train / 255.0
```
Now take a look at the shape of the training data:
```
data_shape = x_train.shape
print(f"There are {data_shape[0]} examples with shape ({data_shape[1]}, {data_shape[2]})")
```
Now it is time to create your own custom callback. For this complete the `myCallback` class and the `on_epoch_end` method in the cell below. If you need some guidance on how to proceed, check out this [link](https://www.tensorflow.org/guide/keras/custom_callback).
```
# GRADED CLASS: myCallback
### START CODE HERE
# Remember to inherit from the correct class
class myCallback():
# Define the correct function signature for on_epoch_end
def on_epoch_end(None, None, None=None):
if logs.get('accuracy') is not None and logs.get('accuracy') > 0.99: # @KEEP
print("\nReached 99% accuracy so cancelling training!")
# Stop training once the above condition is met
None = None
### END CODE HERE
```
Now that you have defined your callback it is time to complete the `train_mnist` function below:
```
# GRADED FUNCTION: train_mnist
def train_mnist(x_train, y_train):
### START CODE HERE
# Instantiate the callback class
callbacks = None
# Define the model, it should have 3 layers:
# - A Flatten layer that receives inputs with the same shape as the images
# - A Dense layer with 512 units and ReLU activation function
# - A Dense layer with 10 units and softmax activation function
model = tf.keras.models.Sequential([
None,
None,
None
])
# Compile the model
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
# Fit the model for 10 epochs adding the callbacks
# and save the training history
history = model.fit(None, None, epochs=None, callbacks=[None])
### END CODE HERE
return history
```
Call the `train_mnist` passing in the appropiate parameters to get the training history:
```
hist = train_mnist(x_train, y_train)
```
If you see the message `Reached 99% accuracy so cancelling training!` printed out after less than 10 epochs it means your callback worked as expected.
**Congratulations on finishing this week's assignment!**
You have successfully implemented a callback that gives you more control over the training loop for your model. Nice job!
**Keep it up!**
| github_jupyter |
# Tutorial `hidrokit.prep.excel`
- **Kategori**: _data preparation_
- __Tujuan__: Memperoleh dataframe dari _excel file_.
- __Dokumentasi__: [readthedocs](https://github.com/taruma/hidrokit-nb/blob/master/notebook/taruma_hidrokit_prep_timeseries.ipynb)
## Informasi notebook
- __notebook name__: `taruma_hidrokit_prep_excel`
- __notebook version/date__: `1.0.2`/`20190713`
- __notebook server__: Google Colab
- __hidrokit version__: `0.2.0`
- **python version**: `3.7`
## CATATAN
Modul `.prep.excel` masih dalam tahap pengembangan sehingga fungsi yang akan ditampilkan pada tutorial ini adalah _private function_ yang ditandai dengan diawali `_` pada setiap fungsinya. Fungsi tersebut seharusnya tidak digunakan oleh _users_ dan hanya bertugas sebagai fungsi di balik layar. Notebook ini hanya melengkapi tutorial hidrokit.
**Disarankan tidak menggunakan _private function_, karena _private function_ akan mengalami perubahan signifikan selama pengembangan.**
## Instalasi hidrokit
```
### Instalasi melalui PyPI
!pip install hidrokit[excel] # Digunakan [excel] karena membutuhkan paket xlrd
### Instalasi melalui Github
# !pip install git+https://github.com/taruma/hidrokit.git
### Instalasi melalui Github (Latest)
# !pip install git+https://github.com/taruma/hidrokit.git@latest
```
## Import Library
```
import numpy as np
import pandas as pd
```
## Dataset
```
# Ambil dataset dari data test hidrokit
!wget -O '2006 data hujan stasiun A.xls' "https://github.com/taruma/hidrokit/blob/master/tests/data/excel/2006%20HUJAN%20DISNEY%20LAND.xls?raw=true"
# Menampilkan file excel dalam bentuk dataframe
filepath = '2006 data hujan stasiun A.xls'
year = 2006
pd.read_excel(filepath)
```
# Fungsi _private_ `prep.excel()`
- __Tujuan__: Memperoleh data dari _excel file_.
- __Dokumentasi__: [readthedocs](https://hidrokit.readthedocs.io/en/stable/prep.html#module-prep.excel)
```
from hidrokit.prep import excel
```
## `_file_single_pivot()`
Memperoleh tabel pivot dari _excel file_
```
tabel_pivot = excel._file_single_pivot(filepath)
tabel_pivot
```
## `_dataframe_table()`
Mengubah tabel pivot ke dataframe kolom tunggal.
```
tabel_tunggal = excel._dataframe_table(tabel_pivot, year, name='sta_a')
tabel_tunggal
```
# Changelog
```
- 20190713 - 1.0.2 - Informasi notebook
- 20190713 - 1.0.1 - Fix typo
- 20190713 - 1.0.0 - Initial
```
#### Copyright © 2019 [Taruma Sakti Megariansyah](https://taruma.github.io)
Source code in this notebook is licensed under a [MIT License](https://opensource.org/licenses/MIT). Data in this notebook is licensed under a [Creative Common Attribution 4.0 International](https://choosealicense.com/licenses/cc-by-4.0/).
| github_jupyter |
# Count Healthy Trees!
## OR: How can we use the Street Tree Census Data to more efficiently plan for the long term health of the urban forest
I celebrated National Day of Civic Hacking on June 4th 2016 by participating in TreesCount! Hackathon hosted by NYC Parks where I analyzed the health of the urban forest to facilitate more efficient disctribution of financial resources across the boroughs. My goal was to gain __actionable insights__ and:
* identify the key indicators of poor health
* forecast which areas in NYC are likely to need more funds in the future
### __To this aim I followed three steps__:
__[1. Data import and exploration](#first-bullet)__
__[2. Analysis and modeling](#second-bullet)__
__[3. Conclusions and future directions](#third-bullet)__
What drawn me to this project is that the data was collected with help of numerous volunteers during 2015 census. Those volunteers (and NYC Park staff) walked every street in NYC and laboriously recorded the GPS coordinates, species and health indicators of almost 500.000 trees. Isn't it amazing?
The dataset is available at https://nycopendata.socrata.com, feel free to play with it yourself, it's very well documented.
## 1. Data import and exploration <a class="anchor" id="first-bullet"></a>
List of Python libraries I'm gonna use:
```
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import ml_helper as mlhelp
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.cross_validation import train_test_split
from sklearn import metrics
from sklearn.cross_validation import cross_val_score
%matplotlib inline
import warnings
warnings.filterwarnings("ignore")
trees = pd.read_csv("/Users/zuzia/Desktop/Coding/2015_Street_Tree_Census.csv")
```
How does the data look like?
```
trees.head()
```
Taking a peek at data format:
Let's see how the distribution of samples looks like around the city. I used Carto to make the map, and you can see the original at https://zuzanna.carto.com/viz/7317a0b0-d77d-11e6-83c6-0e05a8b3e3d7/public_map.
<img src="map_trees.PNG">
Looks like we have a solid coverage across the city, with clear gaps where parks* (e.g. Central Park) or airports are - which is a good sanity check. For now, I'm not looking at the density of the trees in the city, just checking if there are any obvious flaws to the dataset.
*TreesCount dataset includes trees lining the streets of NYC, which excludes NYC Parks. Pretty confusing, but it sort of makes sense - those trees are under different management system.
In the first pass I will use just a subset of features choosing the ones that look like most likely candidates to influence trees health, such as trunk diameter, its health status, location or species.
Some of those variables are on categorical scale, so for the purpose of future analyses I will recode them as such.
```
trees_=pd.concat([trees.tree_dbh,
trees.status,
trees.nta,
trees.steward,
trees.user_type,
trees.spc_common,
trees.borocode,
trees.boroname,
trees.health,
trees.guards,
trees.longitude,
trees.latitude], axis=1)
trees_.columns=['diameter',
'status',
'neighborhood',
'stewardship',
'user',
'species',
'boro',
'boroname',
'health',
'guards',
'longitude',
'latitude']
trees_['status'] = trees_['status'].astype('category')
trees_['neighborhood'] = trees_['neighborhood'].astype('category')
trees_['stewardship'] = trees_['stewardship'].astype('category')
trees_['user'] = trees_['user'].astype('category')
trees_['species'] = trees_['species'].astype('category')
trees_['boro'] = trees_['boro'].astype('category')
trees_['boroname'] = trees_['boroname'].astype('category')
trees_['health'] = trees_['health'].astype('category')
trees_['guards'] = trees_['guards'].astype('category')
```
### Exploration: How healthy are the trees in NYC according to the census?
Does it look like someone is taking care of them? Let's look at the values in the 'stewardship' column
How healthy are the trees?
Do trees have guards (tiny fences around them that help to retain moisture)?
```
fig = plt.figure(figsize = [13,5])
plt.subplot(1,3,1)
h = sns.countplot(x="health", data=trees_, palette="husl")
plt.title('How healthy are the trees overall?', size=13)
plt.subplot(1,3,2)
f = sns.countplot(x="stewardship", data=trees_, palette="husl")
plt.title('How many stewardship signs trees have?', size=13)
plt.subplot(1,3,3)
g = sns.countplot(x="guards", data=trees_, palette="husl")
plt.title('How many trees have helpful guards around them?', size=13)
plt.tight_layout()
plt.show()
```
Is there a relationship between stewardship signs and health?
Is there a relationship between guards and health?
```
fig = plt.figure(figsize = [11,5])
plt.subplot(1,2,1)
f = sns.countplot(x="stewardship", data=trees_, palette="husl", hue='health')
plt.title('Stewardship signs and health', size=13)
plt.legend(loc=2)
plt.subplot(1,2,2)
h = sns.countplot(x="guards", data=trees_, palette="husl", hue='health')
plt.title('Guards around trunks and trees health', size=13)
plt.legend(loc=2)
plt.tight_layout()
plt.show()
```
Doesn't seem like it. Just from visual inspection seems like, proportionally, there are more healthy than in "poor condition" trees among those that have no guards whatsoever compared to those that do have (helpful or harmful).
How about trees' diameter? Does it look like there is a difference on any of the above features depending on trees' diameter?
```
fig = plt.figure(figsize = [11,5])
g = sns.PairGrid(trees_,
x_vars=['status','stewardship', 'user', 'boro','health'],
y_vars=["diameter"],
aspect=.75, size=3.5)
g.map(sns.stripplot, jitter=True, palette="Greens_d");
```
### Who collects the data? Are volunteers attracted more to certain boroughs?
This is an important question which explores how the community of volunteers was involved in the TreesCount! initiative. I expected that some boroughs, e.g. Manhattan, will be more attractive for volunteers to work (or more volunteers come from these areas).
```
from matplotlib.gridspec import GridSpec
fig = plt.figure(figsize = [13,8])
gs=GridSpec(2,2)
ax1=fig.add_subplot(gs[0,0])
#plt.subplot(2,2,1)
f = sns.countplot(x="user", data=trees_, palette="husl")
plt.title('Who collected the data?', size=15)
plt.ylabel('Number of entries', size=13)
#plt.subplot(2,2,2)
ax2=fig.add_subplot(gs[0,1])
h = sns.countplot(x="user", data=trees_, palette="husl", hue='health')
plt.title('Did it matter for health who collected the data?', size=15)
plt.ylabel('Number of entries', size=13)
#plt.subplot(2,2,3)
ax4=fig.add_subplot(gs[1,:])
g = sns.countplot(x="user", data=trees_, palette="husl", hue='boroname')
plt.title('Where did volunteers collect most data?', size=15)
plt.ylabel('Number of entries', size=13)
plt.tight_layout()
plt.show()
```
Interesting. Seems like we have a solid number of records entered by volunteers, about half. How does it look across the boroughs?
Looks like Manhattan, Brooklyn and Queens were favored by volunteer teams, with NYC Parks staff & TressCount staff leading the collection in the other boroughs. This came as a big surprise to me - in three out of five boroughs volunteers contributed between 45000 and 50000 entries! In Manhattan the difference is particurarly striking, volunteers entered approximately twice as many samples into the system as staff members. By contrast, in Bronx and Staten Island it was largely by staff efforts that we have a detailed map of the trees.
I suspected that if we look at the map it will be the farthest neighborhoods from Manhattan that were traversed by the staff. Mapping our data would also allow to quickly look at another variable which plays a role for volunteer programs: seasonality. For civic engagement time of the year definitely matters. For example, if a non-profit is planning an outdoor program, they will most likely avoid doing it in winter. Was it the case with the TreesCount! initiative?
### Does time of the year matter?
How did the data collection evolve over time? Are there any seasonal trends? Having data from just one year we can't use any reliable quantitative methods, such as ARIMA, so our observations are purely speculative.
The map below represents data entries within five boroughs recorded by three groups: staff (NYC Parks & TreesCount, in different shades of blue) and volunteers (green). Missing entries are plotted in grey. You can play around by changing the date of the records in the lower left corner.
If you're viewing this on Github the map will not be visible, you need to either download my Jupyter notebook or go to https://zuzanna.carto.com/viz/1583bbc6-d77f-11e6-af25-0e05a8b3e3d7/public_map to view the map. __The map below is a static screenshot, in order to see seasonal changes you need to click on the link.__
<img src="map_seasonality.PNG">
Few observations:
* The number of data entries, especially from volunteers, seems to be increasing with time. There could be a number of reasons. It could mean that either NYC trained volunteers / distributed the equipment later than they dispatched their own staff or that more volunteers signed up to do the data collection.
* in winter months it is usually the staff who collects the data - volunteers get more excited about being outdoors when it's warmer outside:)
* "null" values are samples from the most recent months, likely haven't been processed yet. Therefore, it would be unwise to get rid of them.
### Are certain species more likely to be unhealthy?
What are different species? What are the most numerous ones?
```
trees_['species'].value_counts().head()
```
Is there a relationship between species and health? Let's look at a plot of simple counts of trees within each specie that were categorized as "healthy" and those that were described as being in "fair" or "poor" health.
```
fig = plt.figure(figsize = [18,22])
ax = fig.add_subplot(411, title = 'Species of all trees')
ax.set_xticklabels(ax.xaxis.get_majorticklabels(), rotation = 90,fontsize = 10)
ax1 = fig.add_subplot(412, title = 'Species of trees in good health')
ax1.set_xticklabels(ax.xaxis.get_majorticklabels(), rotation = 90,fontsize = 10)
ax2 = fig.add_subplot(413, title = 'Species of trees in fair health')
ax2.set_xticklabels(ax.xaxis.get_majorticklabels(), rotation = 90,fontsize = 10)
ax3 = fig.add_subplot(414, title = 'Species of trees in poor health')
ax3.set_xticklabels(ax.xaxis.get_majorticklabels(), rotation = 90,fontsize = 10)
f = sns.countplot(ax=ax, x="species", data=trees_, palette="husl")
h = sns.countplot(ax=ax1, x="species", data=trees_[trees_['health'] == 'Good'], palette="husl")
g = sns.countplot(ax=ax2, x="species", data=trees_[trees_['health'] == 'Fair'], palette="husl", )
p = sns.countplot(ax=ax3, x="species", data=trees_[trees_['health'] == 'Poor'], palette="husl", )
plt.tight_layout()
plt.show()
```
Visual inspection suggests there aren't striking differences between distribution of trees labeled as "healthy" versus "in fair health" (compare second and third plots), however there are some species that seem to be more prone to fall into the unhealthy category. For example, Cherry, Ginkgo or Little Leaf Linden's bars seem to be higher in the third compared to second plot. This might be simply because it's easier to notice the difference in species that are common in the data set (and are subsequently represented with higher bars) or that other species are less likely to be labeled as unhealthy.
For analysis purposes, it will be useful to recode some values (e.g. strings into integers, or multiple classes into binary variable) and clean the dataset (e.g. remove dead trees).
```
cat_columns = trees_.select_dtypes(['category']).columns
trees_[cat_columns] = trees_[cat_columns].apply(lambda x: x.cat.codes)
#remove tree stumps
trees_ = trees_[trees_.stewardship != 0]
#remove an outlier that has diameter of 40 000 (most likely a typo)
trees_ = trees_[trees_.diameter <100000]
#recode guards variable
recode2={3:0}
trees_['guards'][trees_.guards==3]= trees_['guards'].map(recode2)
recode3={1:0}
trees_['guards'][trees_.guards==1]= trees_['guards'].map(recode3)
recode4={4:0}
trees_['guards'][trees_.guards==4]= trees_['guards'].map(recode4)
recode5={2:1}
trees_['guards'][trees_.guards==2]= trees_['guards'].map(recode5)
```
There is also the problem of a relatively small number of trees in poor health - around 20 000 out of 450 000. For the purpose of this analysis, I will add them to the trees in "fair" health to avoid __drastic class imbalance__ in the analysis.
```
#recode fair into poor health
recode0={2:0}
trees_['health'][trees_.health==2]= trees_['health'].map(recode0)
recode1={3:1}
trees_['health'][trees_.health==3]= trees_['health'].map(recode1)
```
Important thing to remeber is that label "1" means now that a tree is in __*poor*__ health health and "0" that it is in __*good*__ health. I changed it since __*the goal of this analysis is to determine the factors that predict poor health*__. It is an arbitrary decision, of course.
```
trees_['health'].value_counts()
```
Next, I create dummy variables and put them all in one matrix with the variable I want to predict: the health of a tree. Then we're finally ready to run an analysis:)
```
dummy_ranks_hood = pd.get_dummies(trees_['neighborhood'], prefix='neighborhood')
dummy_ranks_steward = pd.get_dummies(trees_['stewardship'], prefix='stewardship')
dummy_ranks_user = pd.get_dummies(trees_['user'], prefix='user')
dummy_ranks_species = pd.get_dummies(trees_['species'], prefix='species')
dummy_ranks_boro = pd.get_dummies(trees_['boro'], prefix='boro')
dummy_ranks_guards = pd.get_dummies(trees_['guards'], prefix='guards')
X_vec=pd.concat([trees_['diameter'],
dummy_ranks_steward,
dummy_ranks_user,
dummy_ranks_guards,
dummy_ranks_hood,
dummy_ranks_species],1)
X=X_vec
y=trees_['health']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.6, random_state=0)
print X_train.shape
print y_test.shape
```
# 2. Analysis and Modeling <a class="anchor" id="second-bullet"></a>
## Logistic regression
```
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.6, random_state=0)
logit = LogisticRegression()
logit = logit.fit(X_train, y_train)
logit_test_pred = logit.predict(X_test)
logit_test_score = logit.score(X_test, y_test)
logit_y_prob_test = logit.predict_proba(X_test)
logit_y_prob_train = logit.predict_proba(X_train)
logit_y = logit.predict_proba(X)
fpr_train, tpr_train, _ = metrics.roc_curve(y_train,logit_y_prob_train[:,1])
roc_auc_train = metrics.auc(fpr_train, tpr_train)
fpr_test, tpr_test, _ = metrics.roc_curve(y_test,logit_y_prob_test[:,1])
roc_auc_test = metrics.auc(fpr_test, tpr_test)
print"The accuracy of logistic regression is " + str(logit_test_score)
print "Chance level for this data is " + str(y.mean())
```
### Interpreting the results
The accuracy of our model is 78%, which means that in 78% of the cases we were able to predict the label of the case (healthy tree versus unhealthy one). That sounds good, right? Well, not really. When you take into account that "1" means that a tree is in poor health, you realize we should flip the chance level since it is a proportion of "1s" to "0s". When we do that:
```
print "Corrected chance level for this data set is " + str(1-y.mean())
```
Not so great anymore, right? When we look at the confusion matrix what's the likely culprit:
```
target_names = ['healthy trees', 'unhealthy trees']
print (metrics.confusion_matrix(y_test, logit_test_pred))
print (metrics.classification_report(y_test, logit_test_pred, target_names=target_names))
mlhelp.classification_accuracy(logit_test_pred, y_test);
```
For "1" label recall metric where \begin{equation*}recall = \frac{truepositives}{truepositives + falsepositives}\end{equation*} is merely 3%. It is a pretty terrible result!
As I mentioned before, there is big a class imbalance problem in our data set - only 20% of trees are labeled as in "poor" or "fair" health, which might have a __big__ influence on classification accuracy. Scikit learn actually has a way of dealing with this using class_weight="balanced" parameter.
## Improving logistic regression model
```
logitcv = LogisticRegression(class_weight='balanced')
logitcv.fit(X_train, y_train)
logitcv_test_pred = logitcv.predict(X_test)
print (metrics.confusion_matrix(y_test, logitcv_test_pred))
print (metrics.classification_report(y_test, logitcv_test_pred, target_names=target_names))
```
### __Taking into account class imbalance in the model does not increase our overall accuracy (in fact, it makes it worse) but it increases recall and precision for unhealthy trees (true positives) from 57% and 5% to 60% and 40%, respectively.__
This is great, because our goal here is to predict factors which help us detect unhealthy trees, not increase the average accuracy. See below the analysis with balanced class model parameter introduced.
```
logitcv_test_score = logitcv.score(X_test, y_test)
logitcv_y_prob_test = logitcv.predict_proba(X_test)
logitcv_y_prob_train = logitcv.predict_proba(X_train)
logitcv_y = logitcv.predict_proba(X)
fpr_cv_train, tpr_cv_train, _ = metrics.roc_curve(y_train,logitcv_y_prob_train[:,1])
roc_cv_auc_train = metrics.auc(fpr_cv_train, tpr_cv_train)
fpr_cv_test, tpr_cv_test, _ = metrics.roc_curve(y_test,logitcv_y_prob_test[:,1])
roc_cv_auc_test = metrics.auc(fpr_cv_test, tpr_cv_test)
print"The accuracy of logistic regression is " + str(logitcv_test_score)
print "Chance level for this data is " + str(1 - y.mean())
```
Now let's check what's the optimal regularization parameter using LogisticRegressionCV which looks for it via grid search using cross validation. Then we can feed it back to the model. We'll also change default scoring method which takes into account raw accuracy into one that optimizes for ROC / AUC.
```
from sklearn.linear_model import LogisticRegressionCV
lrcv = LogisticRegressionCV(scoring='roc_auc', class_weight='balanced')
lrcv.fit(X_train, y_train)
lrcv.C_
# Try again with the better regularisation parameter
learner = LogisticRegression(class_weight='balanced', C=lrcv.C_[0])
learner.fit(X_train, y_train)
y_test_predict = learner.predict(X_test)
y_test_probs = learner.predict_proba(X_test)
mlhelp.classification_accuracy(y_test_predict, y_test);
Xv = {'X_test': X_test, 'X_train': X_train}
yv = {'y_test': y_test, 'y_train': y_train}
metrics_ = mlhelp.print_cross_validation_metrics(Xv, yv, learner)
y_prob_val = learner.predict_proba(X)
precision_val, recall_val, thresholds_val = metrics.precision_recall_curve(y,y_prob_val[:,1])
thresholds_val = np.append(thresholds_val, 1)
queue_rate_val = []
for threshold in thresholds_val:
queue_rate_val.append((y_prob_val[:,1] >= threshold).mean())
plt.figure(figsize=(8,6))
plt.plot(thresholds_val, precision_val, color = 'b', linewidth=3.0)
plt.plot(thresholds_val, recall_val, color = 'g', linewidth=3.0)
plt.plot(thresholds_val, queue_rate_val, color = 'r', linewidth=3.0)
leg = plt.legend(('precision', 'recall', 'queue_rate'),loc="lower right")
plt.xlabel('threshold', size=13)
plt.xlim([0.0, 0.6])
plt.title('Precision, Recall and Queue Rate as a function of threshold')
```
## Feature importance
```
coeff = pd.DataFrame(zip(X.columns, np.transpose(learner.coef_)))
coeff.columns=['name','value']
newCoeffs = np.array([thisCoef[0] for thisCoef in coeff['value']])
odds = np.exp(newCoeffs)
odds = pd.Series(odds)
odds = pd.Series.to_frame(odds)
odds = odds - 1
odds.columns = ['value']
odds['name'] = coeff['name']
#most healthy
odds = odds.sort_values('value',ascending=False)
fig = plt.figure(figsize = [20,5])
ax = sns.barplot(x="name", y="value", data=odds[odds['value'] > 0], palette="Greens_d")
ax.set_xticklabels(ax.xaxis.get_majorticklabels(), rotation = 90,fontsize = 8)
ax.set_title('Odds for logistic regression coeffitients: factors that increase the risk of poor health')
#plt.tight_layout()
plt.show()
```
These are features that are most likely to increase the odds that a tree is in poor health.
```
high_odds = odds[odds['value'] > 1]
high_odds.head()
fig = plt.figure(figsize = [20,5])
ax = sns.barplot(x="name", y="value", data=odds[odds['value'] < 0], palette="Greens_d")
ax.set_xticklabels(ax.xaxis.get_majorticklabels(), rotation = 90,fontsize = 8)
ax.set_title('Odds for logistic regression coeffitients: factors that decrease the risk of poor health')
#plt.tight_layout()
plt.show()
low_odds = odds[odds['value'] < -0.5]
low_odds
y_prob_val = logit.predict_proba(X_vec)
trees_['score_logit'] = y_prob_val[:,1]
trees_.head()
```
# Analysis and modeling: random forests
```
forest = RandomForestClassifier(n_jobs=-1)
forest.fit(X_train,y_train)
forest_y_prob_train = forest.predict_proba(X_train)
forest_y_prob_test = forest.predict_proba(X_test)
fpr_train, tpr_train, _ = metrics.roc_curve(y_train,forest_y_prob_train[:,1])
roc_auc_train = metrics.auc(fpr_train, tpr_train)
fpr_test, tpr_test, _ = metrics.roc_curve(y_test,forest_y_prob_test[:,1])
roc_auc_test = metrics.auc(fpr_test, tpr_test)
rf_test_score = forest.score(X_test, y_test)
print "Chance level for this data is " + str(y.mean())
print "Accuracy of random forests is " + str(1-(rf_test_score))
import matplotlib.lines as mlines
plt.figure(figsize=(7,7))
plt.plot(fpr_train, tpr_train, color = 'r')
plt.plot(fpr_test, tpr_test, color = 'g')
plt.plot([0, 1], [0, 1], 'k--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.0])
plt.xlabel('False Positive Rate', size=13)
plt.ylabel('True Positive Rate', size=13)
plt.title('ROC curve for Random Forests', size=13)
green_line = mlines.Line2D([], [], color='green', label='ROC test (area = %0.2f)' % np.mean(roc_auc_test))
red_line = mlines.Line2D([], [], color='red', label='ROC train (area = %0.2f)' % roc_auc_train)
plt.legend(handles=[red_line, green_line],loc="lower right")
y_prob_val = forest.predict_proba(X)
precision_val, recall_val, thresholds_val = metrics.precision_recall_curve(y,y_prob_val[:,1])
thresholds_val = np.append(thresholds_val, 1)
queue_rate_val = []
for threshold in thresholds_val:
queue_rate_val.append((y_prob_val[:,1] >= threshold).mean())
plt.figure(figsize=(8,6))
plt.plot(thresholds_val, precision_val, color = 'b', linewidth=3.0)
plt.plot(thresholds_val, recall_val, color = 'g', linewidth=3.0)
plt.plot(thresholds_val, queue_rate_val, color = 'r', linewidth=3.0)
leg = plt.legend(('precision', 'recall', 'queue_rate'),loc="lower right")
plt.xlabel('threshold')
plt.xlim([0.0, 0.6])
```
Overfitting? Trying grid search with a subsample
```
import random
from sklearn import grid_search
num = 20000 #number of samples
sub_ind = random.sample(list(X_vec.index),num)
X_vec_200k = X_vec.loc[sub_ind]
y_200k = y.loc[sub_ind]
rf = RandomForestClassifier()
parameters = {'n_estimators': [10,30,100],'max_depth':[2,5,10],'min_samples_leaf':[5,10,20]}
model_cv_grid = grid_search.GridSearchCV(rf,parameters,scoring='roc_auc',verbose=2,n_jobs=-1)
model_cv_grid.fit(X_vec_10k,y_10k)
model_cv_grid.grid_scores_
model_cv_grid.best_estimator_
best_model = model_cv_grid.best_estimator_
model_tuned_200k = RandomForestClassifier(bootstrap=True, class_weight=None, criterion='gini',
max_depth=10, max_features='auto', max_leaf_nodes=None,
min_samples_leaf=10, min_samples_split=2,
min_weight_fraction_leaf=0.0, n_estimators=100, n_jobs=1,
oob_score=False, random_state=None, verbose=0)
#Check over-fitting after Grid Search¶
from sklearn import metrics, cross_validation
X_train_200k, X_test_200k, y_train_200k, y_test_200k = cross_validation.train_test_split(X_vec_200k,y_200k)
model_tuned_200k.fit(X_train_200k,y_train_200k)
y_prob_train_200k = model_tuned_200k.predict_proba(X_train_200k)
y_prob_test_200k = model_tuned_200k.predict_proba(X_test_200k)
fpr_train_200k, tpr_train_200k, _ = metrics.roc_curve(y_train_200k,y_prob_train_200k[:,1])
roc_auc_train_200k = metrics.auc(fpr_train_200k, tpr_train_200k)
fpr_test_200k, tpr_test_200k, _ = metrics.roc_curve(y_test_200k,y_prob_test_200k[:,1])
roc_auc_test_200k = metrics.auc(fpr_test_200k, tpr_test_200k)
plt.figure(figsize=(8,6))
plt.plot(fpr_train_200k, tpr_train_200k, color = 'r')
plt.plot(fpr_test_200k, tpr_test_200k, color = 'g')
plt.plot([0, 1], [0, 1], 'k--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.0])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('ROC after grid search 200k sample')
green_line = mlines.Line2D([], [], color='green', label='ROC test (area = %0.2f)' % np.mean(roc_auc_test_200k))
red_line = mlines.Line2D([], [], color='red', label='ROC train (area = %0.2f)' % roc_auc_train_200k)
plt.legend(handles=[red_line, green_line],loc="lower right")
#plt.savefig('ROC_best_model_10k.pdf')
```
What are the most important features from the model?
```
df_importance_200k = pd.DataFrame({'variable': X_vec.columns, 'importance' : best_model.feature_importances_})
df_importance_200k.sort_values('importance', ascending=False).head() #save the best estimators for later to plot
```
Save the dataset with new predicting variable
```
trees_.to_csv('Desktop/trees_.csv')
```
# 3. Conclusions and future directions <a class="anchor" id="third-bullet"></a>
### __My analysis indicates concrete actions NYC Parks should undertake. When planting new trees, they should take into account that: __
* __Freshly planted trees should be closely monitored for health issues__ as our analysis indicates that *small diameter* is related to poor health (and younger trees tend to have smaller diameter). Possibly, this is already being addressed with existing initiatives such as "I'm your new tree" tag with a QR code which aim to encourage New Yorkers to take care of the new neighbors:
<img src="new_tree.PNG">
It would be great to see if such initiatives work. Unfortunately, since it was introduced relatively recently, it is not possible with this data set.
* __Some species are more prone to health issues__ which should be taken into account when choosing species to be planted
* NYC Parks should direct resources towards neighborhoods where trees' health tends to be worse.
### __Conclusions about the analysis:__
* logistic regression results are comparable to those of random forests, and the results are much easier to interpret
| github_jupyter |
```
# -*- coding: utf-8 -*-
"""
Example script to serve as starting point for evaluating scatter results
The current script reads results from run_scatter_0 and displays them comparing
with the truth (i.e. simulation input and simulation scatter output)
run_scatter_0.sh run the standard SSS scatter estimation algorithm on the
simulated data, and then runs FBP. With this notebook you get an idea
of what the scatter looks like and how accurate the scatter estimation is
in the ideal case (i.e. where the model exactly matches the actual scatter generation).
Prerequisite:
You should have executed the following on your command prompt
./run_simulations_thorax.sh
./run_scatter_0.sh
Author: Kris Thielemans
"""
%matplotlib notebook
```
# Initial imports
```
import matplotlib.pyplot as plt
import stir
from stirextra import *
import os
```
# go to directory with input files
```
# adapt this path to your situation (or start everything in the exercises directory)
os.chdir(os.getenv('STIR_exercises_PATH'))
```
# change directory to where the output files are
```
os.chdir('working_folder/GATE1')
```
# read in data from GATE1
```
# original scatter as generated by the simulation
org_scatter=to_numpy(stir.ProjData.read_from_file('my_scatter_g1.hs'));
# estimated scatter
estimated_scatter=to_numpy(stir.ProjData.read_from_file('scatter_estimate_run0.hs'));
```
# Display bitmaps of a middle sinogram
```
maxforplot=org_scatter.max()*1.1;
plt.figure()
ax=plt.subplot(1,2,1);
plt.imshow(org_scatter[10,:,:,]);
plt.clim(0,maxforplot)
ax.set_title('Original simulated scatter');
plt.axis('off');
ax=plt.subplot(1,2,2);
plt.imshow(estimated_scatter[10,:,:,]);
plt.clim(0,maxforplot);
ax.set_title('estimated scatter');
plt.axis('off');
```
# Display central profiles through the sinogram
```
plt.figure()
plt.plot(org_scatter[10,:,192//2],'b');
#plt.hold(True)
plt.plot(estimated_scatter[10,:,192//2],'c');
plt.legend(('original','estimated'));
```
# Read in images
```
org_image=to_numpy(stir.FloatVoxelsOnCartesianGrid.read_from_file('FDG_g1.hv'));
fbp_result=to_numpy(stir.FloatVoxelsOnCartesianGrid.read_from_file('FBP_recon_with_scatter_correction_run0.hv'));
```
# bitmap display of images
```
maxforplot=org_image.max()*1.1;
slice=10;
plt.figure();
ax=plt.subplot(1,2,1);
plt.imshow(org_image[slice,:,:,]);
plt.colorbar();
plt.clim(0,maxforplot);
ax.set_title('input for simulation')
plt.axis('off');
ax=plt.subplot(1,2,2);
plt.imshow(fbp_result[slice,:,:,]);
plt.clim(0,maxforplot);
plt.colorbar();
ax.set_title('FBP after scatter correction')
plt.axis('off');
```
# horizontal profiles through images
```
plt.figure();
plt.plot(org_image[10,154//2,:],'b');
#plt.hold(True);
plt.plot(fbp_result[10,154//2,:],'c');
plt.legend(('Input for simulation','Reconstruction'));
```
| github_jupyter |
# Moist thermodynamics #
This notebook explores the effect of different processes on the thermodynamic structure of the troposphere. It uses a python thermodynamic libary (aes_thermo).
```
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import aes_thermo as mt
from scipy import interpolate, optimize
!%matplotlib inline
plot_dir = '/Users/m219063/Research/Projects/Thermodynamics/plots/'
print (mt.Rd,mt.cpd,mt.Rv,mt.sd00)
gravity = 9.8076
Rstar = 8.31446261815324
P0 = 100000. # Standard Pressure [Pa]
T0 = 273.15 # Standard Temperature [K]
#
# Based on Park et al (2004) Meteorlogia, O2 levels are declining as CO2 levels rise, but at a tiny arte.
#
x_ar = 9.332e-3
x_o2 = 0.20944
x_n2 = 0.78083
x_co2 = 0.415e-3
#
# Based on Chase (1998) J Phys Chem Ref Data
#
m_ar = 39.948
m_o2 = 15.9994 * 2
m_n2 = 14.0067 * 2
m_co2 = 44.011
m_h2o = 18.01528
cp_ar = 20.786 # 298.15K
cp_o2 = 29.376 # 298.15K or 29.126 @ 200K
cp_n2 = 29.124 # 298.15K or 29.107 @ 200K
cp_co2 = 37.129 # 298.15K or 32.359 @ 200K
cp_h2o = 33.349 + (33.590 - 33.349)/98.15 * (T0-200) # Interpolated to T0 from Chase values (but not used)
s0_ar = 154.845 # 298.15K
s0_o2 = 205.147 # 298.15K
s0_n2 = 191.609 # 298.15K
s0_co2 = 213.795 # 298.15K
s0_h2o = 188.854 # 298.15
md = x_ar*m_ar + x_o2*m_o2 + x_n2*m_n2 + x_co2*m_co2 # molar mass of dry air
q_ar = x_ar *m_ar /md
q_o2 = x_o2 *m_o2 /md
q_n2 = x_n2 *m_n2 /md
q_co2 = x_co2*m_co2/md
Rd = (Rstar/md)*(x_ar+x_o2+x_n2+x_co2) * 1000. #J/kg/K
cpd = ( 1./md)*(x_ar*cp_ar + x_o2*cp_o2 + x_n2*cp_n2 + x_co2*cp_co2) *1000. #J/kg/K
sd00= ( 1./md)*(x_ar*s0_ar + x_o2*s0_o2 + x_n2*s0_n2 + x_co2*s0_co2) * 1000. + cpd * np.log(T0/298.15) # Dry air entropy at P0, T0
cpv = 1865.01 # IAPWS97 at 273.15 , for this we could use the Chase values, but they are closer to 1861
cl = 4219.32 # ''
ci = 2096.70 # ''
lv0 = 2500.93e3 # ''
lf0 = 333.42e3 # ''
Rv = (Rstar/m_h2o) *1000. #J/kg/K
sv00 = (s0_h2o/m_h2o)*1000. + cpv * np.log(T0/298.15)
eps1 = Rd/Rv
eps2 = Rv/Rd -1.
PvC = 22.064e6 # Critical pressure [Pa] of water vapor
TvC = 647.096 # Critical temperature [K] of water vapor
TvT = 273.16 # Triple point temperature [K] of water
PvT = 611.655
print (sd00,sv00,Rd,mt.sd00,mt.sv00,mt.Rd)
```
## Calculating $T$ (or $T_\mathrm{v}$) for different thermodynamic processes ##
We start with a differential form, $\mathrm{d}X$ that allows us to construct how temperature changes with pressure following different processes. The form we construct is similar to $\theta_\mathrm{l}$ allows for a saturated isentrope allowing only for a liquid phase, a saturated isentrope assuming only a solidphase, whereby the saturated vapor pressure over ice is set to that over water at temperatures above 273.15 K. We also calculate the pseudo adiabats, one allowing only a liquid phase, and another that transitions to the ice-phase at temperatures below 273.15 K, and differs from the former by the additional fusion enthalpy. The true reversible isentrope would follow the saturated liquid phase isentrope at temperatures above the triple point and the saturated ice-phase isentrope at temperatures below the triple point.
The integration follows by virtue of $\mathrm{d}X = 0$ and $X=X(T,P,q_\mathrm{t}),$ hence we calculate $\partial X/\partial T$ and $\partial X/ \partial P$ following Stevens and Siebesma (2020), and integrate
$$ \mathrm{d}X = \frac{\partial X}{\partial T} \mathrm{d}T + \frac{\partial X}{\partial P} \mathrm{d}P = 0$$ to solve for $T$ given $P.$ That is $$ T(P) = T(P_0) - \int_{P_0}^P \left(\frac{\partial X}{\partial T} \right)^{-1} \frac{\partial X}{\partial P} \mathrm{d}P$$
```
def dX(TK,PPa,qt,es_formula=mt.es_default,pseudo=False) :
Psat = mt.es(TK,es_formula)
qs = (Psat/(PPa-Psat)) * mt.eps1 * (1-qt)
if (qs < qt):
if (pseudo):
qc = 0.
qd = 1.-qs
else:
qc = qt-qs
qd = 1.-qt
lv = mt.phase_change_enthalpy(TK)
if (es_formula != 'liq' and es_formula != 'analytic-liq'):
cp = qd * mt.cpd + qs * mt.cpv + qc * mt.ci
if ((not pseudo) or (TK < 273.15)): lv += mt.phase_change_enthalpy(TK,fusion=True)
else:
cp = qd * mt.cpd + qs * mt.cpv + qc * mt.cl
R = qd * mt.Rd + qs * mt.Rv
vol = R * TK/ PPa
if (pseudo):
beta_P = R/mt.Rd
else:
beta_P = R/(qd*mt.Rd)
beta_T = beta_P * lv/(mt.Rv * TK)
dX_dT = cp + lv * qs * beta_T/TK
dX_dP = vol * ( 1.0 + lv * qs * beta_P/(R*TK))
else:
cp = mt.cpd + qt * (mt.cpv - mt.cpd)
R = mt.Rd + qt * (mt.Rv - mt.Rd)
vol = R * TK/ PPa
dX_dT = cp
dX_dP = vol
return dX_dT, dX_dP;
```
### Comparing the different temperature changes ###
Here we simply calculate and plot some of the different temperature pressure diagrams associated with the different processes. We use the notation of $\theta_\mathrm{e}$ to denote the saturated isentope and the tilde for the pseudo adiabat. For the first comparision of $\theta_\mathrm{e}$ and $\theta_\mathrm{e,ice}$ we show by the grey line the $T=273.15~\mathrm{K}$ isotherm, at the pressure levels where it would connect the solid and liquid phase isentropes.
To get these results we numerically integrate along a constant $X$ which yields $T(P)$. Fot his we use a numerical ode solver form scipy, here the lsoda solver, but other solvers also work similarly well. These end up being more efficient than a simple integration using an Euler forward method.
```
from scipy.integrate import ode
def integrate_dTdP(T0,P0,P1,dP,qt,es_formula=mt.es_default,pseudo=False):
def f(P, T, qt):
dX_dT, dX_dP = dX(T,P,qt,es_formula,pseudo)
return (dX_dP/dX_dT)
r = ode(f).set_integrator('lsoda',atol=0.0001)
r.set_initial_value(T0, P0).set_f_params(qt)
t1 = P1
dt = dP
Te = []
Tx = []
Px = []
while r.successful() and r.t > t1:
r.integrate(r.t+dt)
Tx.append(r.y[0])
Px.append(r.t)
return (np.array(Tx),np.array(Px))
qt = 17.e-3
Tx1, Px = integrate_dTdP(300.,101000.,15000.,-10.,qt,es_formula='analytic-liq')
Tx2, Px = integrate_dTdP(300.,101000.,15000.,-10.,qt,es_formula='analytic-mxd')
Tx3, Px = integrate_dTdP(300.,101000.,15000.,-10.,qt,es_formula='analytic-liq',pseudo=True)
Tx4, Px = integrate_dTdP(300.,101000.,15000.,-10.,qt,es_formula='analytic-mxd',pseudo=True)
print (9.81/1.005 - (Tx1[90]-Tx1[101])/0.095679)
print ((Tx1[0]-Tx1[11])*1005/9.81)
fig = plt.figure(figsize=(15,5))
ax1 = plt.subplot(1,3,1)
ax1.set_xlabel('$T$ / K')
ax1.set_ylabel('$P$ / hPa')
plt.plot(Tx1,Px/100.,c='orange',label='$\\theta_\mathrm{e}$')
plt.plot(Tx2,Px/100.,c='orange',ls='dotted',label='$\\theta_\mathrm{e,ice}$')
plt.plot([273.15,273.15],[600,450], c='lightgrey')
plt.gca().invert_yaxis()
plt.legend(loc="lower left")
ax2 = plt.subplot(1,3,2)
ax2.set_xlabel('$T$ / K')
ax2.set_yticklabels([])
plt.plot(Tx1,Px,c='orange',label='$\\theta_\mathrm{e}$')
plt.plot(Tx3,Px,c='teal',label='$\\tilde{\\theta}_\mathrm{e}$')
plt.gca().invert_yaxis()
plt.legend(loc="lower left")
ax3 = plt.subplot(1,3,3)
ax3.set_xlabel('$T$ / K')
ax3.set_yticklabels([])
plt.plot(Tx3,Px/100.,c='teal',label='$\\tilde{\\theta}_\mathrm{e}$')
plt.plot(Tx4,Px/100.,c='teal',ls='dotted',label='$\\tilde{\\theta}_\mathrm{e,ice}$')
plt.gca().invert_yaxis()
plt.legend(loc="lower left")
sns.set_context("talk", font_scale=1.2)
sns.despine(offset=10)
```
### 1.2 Temperature Differences ###
The Temperature differences show the expected behavior. Ice has a substantial effect on the saturated isentrope, whereby here only the differences at temperatures below the triple point temperature are relevant, i.e., below about 525 hPa. Otherwise the differences between the saturated isentrope and the pseudo adiabat are a few kelvin in the upper troposphere, and the effect of ice processes on the pseudo adiabat contribute also a couple of kelvin near 300 hPa.
```
fig = plt.figure(figsize=(15,5))
ax1 = plt.subplot(1,1,1)
ax1.set_xlabel('$T$ / K')
ax1.set_ylabel('$P$ / hPa')
plt.plot(Tx2-Tx1,Px/100.,c='teal',label='$\\theta_\mathrm{e,ice} - \\theta_\mathrm{e}$')
plt.plot(Tx1-Tx3,Px/100.,c='grey',label='$\\theta_\mathrm{e} - \\tilde{\\theta}_\mathrm{e}$')
plt.plot(Tx4-Tx3,Px/100.,c='orange',label='$\\tilde{\\theta}_\mathrm{e,ice} - \\tilde{\\theta}_\mathrm{e}$')
plt.gca().invert_yaxis()
plt.legend(loc="lower right")
sns.set_context("talk", font_scale=1.2)
sns.despine(offset=10)
```
### Effects on Density ###
We repeat the above exercise but for the density temperature, $T_\rho = T(1+\epsilon_2 q_\mathrm{v} - q_\mathrm{c})$ including $q_\mathrm{c}$ in a manner consistent with the process yielding $T.$ What we find is that (as shown by the Teal curve) the effect of condensate loading increasingly compensates the compositional contribution to $T_\rho$ from water vapor, so that at pressures less than about 700 hPa it increasingly dominates. The grey curve compares the isentrope to the pseudo-adiabat, and shows that slower reduction in temperature of the former increasingly offests the effect of the condensate, so that its density temperature is higher in the upper troposphere. This makes sense because even if all of the vapor (in this case 17 g/kg) is converted to loading it makes a contribution that multiplies an increasingly small temperature, so that at 200K this corresponds to a reduction of 200(1-0.017) or about 4K. At colder temperatures the reduction is propotionally less, but the temperature difference between the saturated isentrope and the pseudo-adiabat is less.
```
Trho1 = np.zeros(len(Px))
Trho3 = np.zeros(len(Px))
for i,x in enumerate(Px):
Psat = mt.es(Tx1[i])
qs = (Psat/(x-Psat)) * (mt.Rd/mt.Rv) * (1-qt)
qv = np.min([qt,qs])
Trho1[i] = Tx1[i]*(1 + mt.eps2 * qv - (qt-qv))
Psat = mt.es(Tx3[i])
qs = (Psat/(x-Psat)) * (mt.Rd/mt.Rv) * (1-qt)
qv = np.min([qt,qs])
Trho3[i] = Tx3[i]*(1 + mt.eps2 * qv)
fig = plt.figure(figsize=(15,5))
ax1 = plt.subplot(1,1,1)
ax1.set_xlabel('$T$ / K')
ax1.set_ylabel('$P$ / hPa')
plt.plot(Trho1-Tx1,Px/100.,c='teal',label='$T_\\rho - T$')
plt.plot(Trho1-Trho3,Px/100.,c='grey',label='$T_\\rho - \\tilde{T}_\\rho$')
plt.gca().invert_yaxis()
plt.legend(loc="center right")
sns.set_context("talk", font_scale=1.2)
sns.despine(offset=10)
```
### Consistency Checks ###
Here we plot $\theta_\mathrm{e}$, $\theta_\mathrm{l}$, and $\theta_s$ calculated from the temperature, pressure and $q_\mathrm{t}$ profiles. These should, if properly defined, be constant, which they are, although not to machine precision, with differences being accumulated finite difference errors, at least they seem to go to zero as the dp is reduced in the numerical integration. For instance reducing dp by 100 eliminates the cloud base jump and the errors maximize at near 3 mK.
We can also compare the pseudo adiabatic temperature increase with the formula from Bolton. For this the differences (O(1K)) are substantial.
```
xfld = np.zeros((len(Px),4))
for i,x in enumerate(Px):
xfld[i,0] = mt.get_theta_e(Tx1[i],x,qt)
xfld[i,1] = mt.get_pseudo_theta_e(Tx3[i],x,qt)
fig = plt.figure(figsize=(10,5))
ax1 = plt.subplot(1,2,1)
ax1.set_xlabel('$\\theta_\mathrm{e}, \\tilde{\\theta}_\mathrm{e}$ / K')
ax1.set_ylabel('$P$ / hPa')
plt.plot(xfld[:,0]-xfld[0,0],Px/100.,c='teal',label='$\\Delta \\theta_\mathrm{e}$')
plt.plot(xfld[:,1]-xfld[0,1],Px/100.,c='orange',label='$\\Delta \\tilde{\\theta}_\mathrm{e}$')
plt.gca().invert_yaxis()
plt.legend(loc="lower left")
for i,x in enumerate(Px):
xfld[i,1] = mt.get_theta_l(Tx1[i],x,qt)
xfld[i,3] = mt.get_theta_s(Tx1[i],x,qt)
ax2 = plt.subplot(1,2,2)
ax2.set_xlabel('mK')
ax2.set_yticklabels([])
plt.plot((xfld[:,0]-xfld[0,0])*1000.,Px/100.,c='teal',label='$\\Delta \\theta_\mathrm{e}$')
plt.plot((xfld[:,1]-xfld[0,1])*1000.,Px/100.,c='dodgerblue',label='$\\Delta \\theta_\mathrm{l}$')
plt.plot((xfld[:,3]-xfld[0,3])*1000.,Px/100.,c='orange',ls='dashed',label='$\\Delta \\theta_s$')
plt.gca().invert_yaxis()
plt.legend(loc="lower left")
sns.set_context("talk", font_scale=1.2)
sns.despine(offset=10)
fig.savefig(plot_dir+'thetas-consistency.pdf')
fig = plt.figure(figsize=(6,6))
ax2 = plt.subplot(1,1,1)
ax2.set_xlabel('mK')
ax1.set_ylabel('$P$ / hPa')
plt.plot((xfld[:,0]-xfld[0,0])*1000.,Px/100.,c='orange',label='$\\Delta \\theta_\mathrm{e}$')
plt.plot((xfld[:,1]-xfld[0,1])*1000.,Px/100.,c='teal',label='$\\Delta \\theta_\mathrm{l}$')
plt.plot((xfld[:,3]-xfld[0,3])*1000.,Px/100.,c='dodgerblue',ls='dashed',label='$\\Delta \\theta_s$')
plt.gca().invert_yaxis()
plt.legend(loc="lower left")
plt.tight_layout()
sns.set_context("talk", font_scale=1.2)
sns.despine(offset=10)
fig.savefig(plot_dir+'thetas-consistency.pdf')
```
### Dependence of $\theta_e$ on $q_t$ ###
### Solving for $T$ from $T_\rho$ and comparing with data###
Here I take to temperature profiles composited from the DYAMOND ICON simulations for a wet 99.999 percentile sounding and a dry sounding. These we then use to look at what processes determine what I call the master temperature profile, i.e., the temperature profile in the convecting regions that is communicated through the tropical atmosphere.
```
Twet = np.array([
223.9353 , 221.97095, 220.03784, 218.19221, 216.31929, 214.27661,
211.73238, 208.92947, 206.17117, 203.89218, 202.05592, 200.44272,
198.63155, 196.21927, 194.47081, 195.16386, 197.48691, 200.50444,
203.81912, 207.28535, 210.82382, 214.43098, 218.06685, 221.70439,
225.32275, 228.9053 , 232.43118, 235.8637 , 239.20024, 242.42961,
245.54013, 248.53214, 251.42017, 254.20903, 256.89557, 259.48453,
261.9759 , 264.38116, 266.70425, 268.95264, 271.08478, 273.0942 ,
274.92133, 276.79697, 278.5961 , 280.27567, 281.83926, 283.30765,
284.69043, 285.99323, 287.22113, 288.37936, 289.47232, 290.50528,
291.48358, 292.41122, 293.28616, 294.0999 , 294.83893, 295.48620,
296.02863, 296.47165, 296.83948, 297.22357, 297.66333])
Pwet = np.array([
1781.3881, 2127.0454, 2521.3892, 2965.2852, 3458.2715,
4001.646 , 4597.039 , 5246.1895, 5949.923 , 6707.3438,
7515.943 , 8372.736 , 9275.428 , 10223.398 , 11213.994 ,
12235.951 , 13273.557 , 14314.763 , 15366.591 , 16440.645 ,
17551.559 , 18717.012 , 19938.434 , 21217.395 , 22555.648 ,
23955.1 , 25417.795 , 26946.035 , 28542.363 , 30209.469 ,
31950.23 , 33767.75 , 35665.18 , 37645.79 , 39712.992 ,
41870.387 , 44121.695 , 46470.805 , 48921.7 , 51478.516 ,
54104.145 , 56747.926 , 59396.582 , 62041.99 , 64675.426 ,
67288.7 , 69873.65 , 72421.9 , 74924.74 , 77373.4 ,
79758.9 , 82072.14 , 84303.92 , 86444.89 , 88485.586 ,
90416.375 , 92227.414 , 93908.73 , 95449.89 , 96839.95 ,
98067. , 99117.28 , 99973.836 , 100613.055 , 100989.66 ])
Qwet = np.array([
2.675785e-06, 2.653257e-06, 2.639634e-06, 2.634951e-06, 2.638471e-06,
2.649419e-06, 2.671635e-06, 2.722334e-06, 2.832603e-06, 3.010935e-06,
3.215084e-06, 3.322613e-06, 3.304213e-06, 3.132450e-06, 3.003545e-06,
3.306731e-06, 4.431343e-06, 6.642379e-06, 1.031196e-05, 1.604714e-05,
2.473562e-05, 3.774666e-05, 5.679549e-05, 8.410055e-05, 1.224644e-04,
1.757509e-04, 2.497731e-04, 3.527200e-04, 4.911225e-04, 6.678019e-04,
8.902289e-04, 1.161963e-03, 1.485134e-03, 1.860150e-03, 2.291250e-03,
2.778659e-03, 3.313594e-03, 3.888310e-03, 4.490240e-03, 5.116668e-03,
5.753872e-03, 6.392588e-03, 6.993116e-03, 7.623885e-03, 8.274390e-03,
8.951828e-03, 9.632755e-03, 1.030219e-02, 1.095124e-02, 1.157832e-02,
1.218259e-02, 1.276330e-02, 1.332142e-02, 1.385734e-02, 1.437104e-02,
1.486405e-02, 1.533483e-02, 1.577741e-02, 1.617590e-02, 1.651940e-02,
1.681609e-02, 1.709637e-02, 1.738570e-02, 1.766170e-02, 1.801199e-02])
Tdry = np.array([
260.46118, 258.63837, 255.8278 , 251.66296, 247.25632, 242.86287,
238.7334 , 235.24545, 232.37387, 230.01653, 227.97017, 226.12338,
224.44499, 222.37727, 220.10141, 218.00972, 216.08081, 214.18835,
212.19333, 210.03035, 207.1936 , 203.98264, 200.97385, 198.6071 ,
197.285 , 196.89725, 197.44398, 198.81459, 200.74696, 203.02396,
205.48637, 208.06682, 210.85538, 213.86523, 217.0021 , 220.21277,
223.46024, 226.73601, 230.04068, 233.36826, 236.69199, 239.98233,
243.23015, 246.4296 , 249.54622, 252.53671, 255.41533, 258.21182,
260.9037 , 263.49518, 266.03485, 268.47684, 270.7939 , 272.96356,
274.96674, 276.79706, 278.5634 , 280.32187, 282.00894, 283.64075,
285.23346, 286.68976, 287.80414, 288.41498, 288.44867, 288.2182 ,
288.21365, 288.74942, 289.6866 , 290.72845, 291.8448 , 292.96866,
294.00327, 294.89948, 295.63406, 296.19107, 296.5726 ])
Pdry = np.array([
141.82285, 179.46115, 225.8453 , 282.9698 , 353.243 ,
439.3996 , 544.55994, 672.13464, 825.7498 , 1009.26807,
1226.8136 , 1482.7734 , 1781.594 , 2126.512 , 2520.2896 ,
2964.1074 , 3457.401 , 4001.0703 , 4595.8213 , 5242.242 ,
5941.6104 , 6695.758 , 7504.999 , 8367.091 , 9276.7705 ,
10226.591 , 11207.969 , 12211.555 , 13228.347 , 14250.45 ,
15286.276 , 16347.967 , 17450.18 , 18610.117 , 19828.947 ,
21108.135 , 22449.408 , 23854.576 , 25325.535 , 26864.16 ,
28472.465 , 30152.693 , 31907.277 , 33738.805 , 35650.07 ,
37644.336 , 39725.1 , 41895.926 , 44160.445 , 46522.6 ,
48986.246 , 51555.383 , 54192.883 , 56847.95 , 59507.273 ,
62163.582 , 64808.98 , 67434.64 , 70031.65 , 72591.17 ,
75104.195 , 77561.9 , 79956.48 , 82281.22 , 84529.96 ,
86695.2 , 88766.5 , 90730.69 , 92574.266 , 94284.92 ,
95851.39 , 97262.33 , 98505.93 , 99568.93 , 100434.836 ,
101080.55 , 101460.92 ])
Qdry = np.array([
3.248840e-06, 3.216901e-06, 3.173897e-06, 3.116910e-06, 3.054899e-06,
3.004614e-06, 2.968617e-06, 2.941603e-06, 2.915867e-06, 2.880306e-06,
2.831446e-06, 2.763737e-06, 2.691604e-06, 2.622797e-06, 2.574177e-06,
2.552536e-06, 2.546951e-06, 2.549817e-06, 2.563278e-06, 2.589809e-06,
2.637191e-06, 2.734801e-06, 2.932778e-06, 3.241792e-06, 3.620764e-06,
4.022001e-06, 4.414107e-06, 4.910287e-06, 5.685483e-06, 6.925651e-06,
8.815266e-06, 1.150364e-05, 1.507771e-05, 1.948587e-05, 2.502891e-05,
3.231415e-05, 4.203726e-05, 5.449402e-05, 6.963828e-05, 8.651965e-05,
1.041388e-04, 1.225337e-04, 1.410077e-04, 1.571900e-04, 1.739966e-04,
1.960605e-04, 2.235830e-04, 2.576729e-04, 3.022493e-04, 3.610218e-04,
4.288332e-04, 5.132451e-04, 6.098776e-04, 7.398597e-04, 9.299355e-04,
1.195684e-03, 1.505343e-03, 1.819583e-03, 2.139176e-03, 2.443166e-03,
2.723889e-03, 3.043162e-03, 3.573373e-03, 4.478660e-03, 5.825936e-03,
7.405198e-03, 8.816672e-03, 9.857900e-03, 1.068776e-02, 1.156250e-02,
1.228730e-02, 1.265830e-02, 1.280781e-02, 1.288322e-02, 1.294559e-02,
1.303250e-02, 1.331252e-02])
Tnicam = np.array([
299.40314, 298.7981 , 298.28796, 297.81036, 297.33783, 296.8393 ,
296.31067, 295.73785, 295.1084 , 294.41476, 293.64764, 292.81082,
291.9038 , 290.89618, 289.79456, 288.55908, 287.19855, 285.6665 ,
283.97977, 282.13297, 280.28662, 278.352 , 276.38852, 274.12656,
272.58176, 270.62924, 268.55392, 266.41882, 264.21167, 261.90262,
259.50546, 257.01053, 254.39836, 251.67212, 248.8163 , 245.84465,
242.74887, 239.5574 , 236.26633, 232.89021, 229.4885 , 225.99297,
222.43375, 218.86075, 215.34106, 211.86104, 208.48413, 205.1801 ,
202.05127, 199.45296, 197.26627, 195.548 , 194.45282, 194.07564,
194.45062, 196.11362, 199.27036, 202.20082, 204.71103, 206.66414,
208.68393, 210.69957, 212.40833, 214.59325, 216.61465, 218.27762,
220.64337, 223.15 , 225.62079, 227.8689 , 230.6178 , 234.07594,
237.83073, 242.83904, 248.02869, 252.65822, 261.12402, 272.38043])
Pnicam = np.array([
1.005161e+05, 9.972547e+04, 9.885742e+04, 9.790580e+04, 9.686296e+04,
9.572198e+04, 9.447429e+04, 9.310973e+04, 9.162220e+04, 9.000211e+04,
8.823859e+04, 8.632358e+04, 8.424945e+04, 8.200543e+04, 7.958516e+04,
7.698066e+04, 7.418697e+04, 7.120077e+04, 6.802407e+04, 6.482330e+04,
6.175041e+04, 5.880122e+04, 5.596507e+04, 5.325968e+04, 5.068687e+04,
4.818499e+04, 4.579259e+04, 4.350231e+04, 4.130877e+04, 3.920805e+04,
3.719646e+04, 3.526966e+04, 3.342448e+04, 3.165760e+04, 2.996585e+04,
2.834629e+04, 2.679627e+04, 2.531299e+04, 2.389390e+04, 2.253645e+04,
2.123837e+04, 1.999742e+04, 1.881185e+04, 1.767964e+04, 1.659903e+04,
1.556874e+04, 1.458736e+04, 1.365369e+04, 1.276666e+04, 1.192616e+04,
1.113196e+04, 1.038355e+04, 9.680594e+03, 9.022758e+03, 8.409618e+03,
7.840943e+03, 7.289986e+03, 6.734334e+03, 6.176722e+03, 5.619943e+03,
5.067861e+03, 4.525567e+03, 3.997939e+03, 3.490395e+03, 3.008514e+03,
2.556543e+03, 2.139034e+03, 1.760290e+03, 1.422642e+03, 1.127076e+03,
8.734735e+02, 6.618490e+02, 4.893133e+02, 3.524920e+02, 2.471570e+02,
1.682373e+02, 1.110651e+02, 7.120367e+01])
Qnicam = np.array([
2.026752e-02, 2.010065e-02, 1.986337e-02, 1.962394e-02, 1.937968e-02,
1.910611e-02, 1.880108e-02, 1.844529e-02, 1.804634e-02, 1.760882e-02,
1.714808e-02, 1.665455e-02, 1.612683e-02, 1.555105e-02, 1.493348e-02,
1.425255e-02, 1.353385e-02, 1.273361e-02, 1.189201e-02, 1.101053e-02,
1.016815e-02, 9.324548e-03, 8.506890e-03, 7.746079e-03, 7.136990e-03,
6.482956e-03, 5.822762e-03, 5.181503e-03, 4.560578e-03, 3.964126e-03,
3.391524e-03, 2.852461e-03, 2.359560e-03, 1.915161e-03, 1.521001e-03,
1.185296e-03, 9.070947e-04, 6.793398e-04, 4.930609e-04, 3.497433e-04,
2.411266e-04, 1.664857e-04, 1.155010e-04, 7.917585e-05, 5.361325e-05,
3.618799e-05, 2.409760e-05, 1.597041e-05, 1.080024e-05, 7.683606e-06,
5.743596e-06, 4.677143e-06, 4.118659e-06, 3.998729e-06, 4.250445e-06,
4.740101e-06, 4.994502e-06, 4.879058e-06, 4.646197e-06, 4.473122e-06,
4.322985e-06, 4.202948e-06, 4.107990e-06, 4.017174e-06, 3.931252e-06,
3.852231e-06, 3.780950e-06, 3.727419e-06, 3.688840e-06, 3.669178e-06,
3.656065e-06, 3.645287e-06, 3.636165e-06, 3.628502e-06, 3.620473e-06,
3.623730e-06, 3.630907e-06, 3.632367e-06])
Soundings = {'dry' : (Tdry,Pdry,Qdry),'wet' : (Tdry,Pdry,Qdry),'nicam' : (Tnicam,Pnicam,Qnicam)}
sounding = 'nicam'
Tdat, Pdat, Qdat = Soundings[sounding]
xfld = np.zeros((len(Pdat),4))
qt = 20.3e-3
Trho = Tdat*(1+mt.eps2*Qdat)
def delta_Tv(TK,Trho,P,qt,loading=True):
# """ calculates difference between the virtual temperature in a saturated updraft with a
# specified qt and a given virtual temperture. Here qs<qt is enforced so that setting qt=0
# gets rid of the water loading effects.
# """
ps = mt.es(TK,es_formula='analytic-mxd')
if (loading):
qs = (ps/(P-ps)) * mt.Rd/mt.Rv * (1-qt)
qv = np.minimum.reduce([qt,qs])
err = (TK*(1. + mt.eps2*qv - (qt-qv)) - Trho)**2
else:
qs = (ps/(P-ps)) * mt.eps1 / (1. - ps*(1.-mt.eps1)/(P-ps))
qv = np.minimum.reduce([qt,qs])
err = (TK*(1. + mt.eps2*qv) - Trho)**2
return err;
Tc = np.zeros(len(Pdat))
Tc_nl = np.zeros(len(Pdat))
Qs = np.maximum((1/(Pdat/mt.es(Tdat,es_formula='analytic-mxd')-1)) * mt.Rd/mt.Rv * (1-qt),1.e-10) #Psat may be greater than Pdat in upper trop.
for i,x in enumerate(Pdat):
Tc[i] = optimize.minimize(delta_Tv, Trho[i], args=(Trho[i],x,qt)).x
if (np.abs(Tc[i]-Trho[i]) > 10.):
print('correcting T = %7.2f to Trho = %7.3f' % (Tc_nl[i],Trho[i],))
Tc[i] = Trho[i]
Tc_nl[i] = optimize.minimize(delta_Tv, Trho[i], args=(Trho[i],x,qt,False)).x
if (np.abs(Tc_nl[i]-Trho[i]) > 10.):
print('correcting T = %7.2f to Trho = %7.3f' % (Tc_nl[i],Trho[i],))
Tc_nl[i]=Trho[i]
Qcs = np.maximum((1/(Pdat/mt.es(Tc,es_formula='analytic-mxd')-1)) * mt.Rd/mt.Rv * (1-qt),1.e-10) #Psat may be greater than Pdat in upper trop.
Qcs_nl = np.maximum((1/(Pdat/mt.es(Tc_nl,es_formula='analytic-mxd')-1)) * mt.Rd/mt.Rv * (1-qt),1.e-10) #Psat may be greater than Pdat in upper trop.
for ipt in [11,21]:
print ('Point values: P at %3.2f hPa yields theta-e of %3.2f K'%(Pdat[ipt]/100.,mt.get_theta_e(Tc[ipt],Pdat[ipt],qt)))
# --------
fig = plt.figure(figsize=(10,5))
ax1 = plt.subplot(1,2,1)
ax1.set_xlabel('K')
ax1.set_ylabel('$P$ / hPa')
ax1.set_ylim(1020,100)
ax1.set_xlim(340,370)
for i,x in enumerate(Pdat):
xfld[i,0] = mt.get_theta_e(Tdat[i],Pdat[i],Qdat[i])
xfld[i,1] = mt.get_pseudo_theta_e(Tdat[i],Pdat[i],Qdat[i])
plt.plot(xfld[:,0],Pdat/100.,c='orange',label='$\\theta_\mathrm{e}$')
plt.plot(xfld[:,1],Pdat/100.,c='teal',label='$\\tilde{\\theta}_\mathrm{e}$')
plt.legend(loc="upper left")
ax2 = plt.subplot(1,2,2)
ax2.set_xlabel('K')
ax2.set_yticklabels([])
ax2.set_ylim(1020,100)
ax2.set_xlim(350,361)
for i,x in enumerate(Pdat):
xfld[i,0] = mt.get_theta_e(Tc[i],Pdat[i],qt)
xfld[i,1] = mt.get_pseudo_theta_e(Tc_nl[i],Pdat[i],Qcs_nl[i])
plt.plot(xfld[:,0],Pdat/100.,c='orange',label='$\\theta_\mathrm{e|\\rho}$')
plt.plot(xfld[:,1],Pdat/100.,c='teal',label='$\\tilde{\\theta}_\mathrm{e|\\rho}$')
#plt.plot(xfld[:,3],Pdat/100.,c='teal',ls='dotted',label='$\\tilde{\\theta}_\mathrm{e}$')
ax2.axvline(340,c='lightgrey',ls='dotted')
plt.legend(loc="upper left")
sns.set_context("talk", font_scale=1.2)
sns.despine(offset=10)
fig.savefig(plot_dir+'jiawei-profiles-'+sounding+'.pdf')
for ipt in [11,21]:
print ('Point values: P at %3.2f hPa yields estimate of delta T of %3.2f K'%(Pdat[ipt]/100.,Tdat[ipt]*(qt-Qdat[ipt])/(1+Qdat[ipt]/mt.eps1 - qt)))
print ('Point values: P at %3.2f hPa yields delta-T of %3.2f K'%(Pdat[ipt]/100.,Tc[ipt]-Tdat[ipt]))
print (2.33/3.1,0.72/1.08,mt.eps2-1)
#
print (np.max(Qdat))
fig = plt.figure(figsize=(10,10))
ax1 = plt.subplot(2,2,1)
ax1.set_xlabel('K')
ax1.set_ylabel('$P$ / hPa')
ax1.set_ylim(1020,100)
ax1.set_xscale('log')
ax1.set_xlim(1.e-5,0.1)
plt.plot(Qdat,Pdat/100.,c='teal',label='$q_\mathrm{v}$')
plt.plot(Qs,Pdat/100.,c='orange',label='$q_\mathrm{s}$')
plt.legend(loc="best")
ax2 = plt.subplot(2,2,2)
ax2.set_xlabel('K')
ax2.set_yticklabels([])
ax2.set_ylim(1020,100)
ax2.set_xlim(1.e-5,0.1)
ax2.set_xscale('log')
plt.plot(Qs-Qdat,Pdat/100.,c='orange',label='$q_\mathrm{s}-q_\mathrm{v}$')
plt.plot(Qdat-Qs,Pdat/100.,c='orange',ls='dashed',label='$q_\mathrm{v}-q_\mathrm{s}$')
plt.legend(loc="best")
ax3 = plt.subplot(2,2,3)
ax3.set_xlabel('K')
ax3.set_ylabel('$P$ / hPa')
ax3.set_ylim(1020,100)
ax3.set_xscale('log')
#ax3.set_xlim(1.e-5,0.1)
plt.plot(Tdat,Pdat/100.,c='teal',label='$T$')
plt.plot(Tc,Pdat/100.,c='orange',label='$T_\mathrm{c}$')
plt.plot(Tc_nl,Pdat/100.,c='orange',ls='dashed',label='$T_\mathrm{c,nl}$')
plt.legend(loc="best")
ax4 = plt.subplot(2,2,4)
ax4.set_xlabel('K')
ax4.set_yticklabels([])
ax4.set_ylim(1020,100)
ax4.set_xlim(-0.2,4.2)
#ax4.set_xscale('log')
plt.plot(Tc-Tdat,Pdat/100.,c='orange',label='$T_\mathrm{c}-T$')
plt.plot(Tc_nl-Tdat,Pdat/100.,c='orange',ls='dashed',label='$T_\mathrm{c,nl}-T$')
plt.legend(loc="best")
plt.tight_layout()
sns.set_context("talk", font_scale=1.2)
sns.despine(offset=10)
fig.savefig(plot_dir+'jiawei-profiles-'+sounding+'.pdf')
sounding = 'dry'
Tdat, Pdat, Qdat = Soundings[sounding]
xfld = np.zeros((len(Pdat),4))
qt = 18e-3
Trho = Tdat*(1+mt.eps2*Qdat)
Psat = mt.es(Tdat)
T = np.zeros(len(Pdat))
Tnl = np.zeros(len(Pdat))
Qs = np.maximum((Psat/(Pdat-Psat)) * mt.Rd/mt.Rv * (1-qt),1.e-10) #Psat may be greater than Pdat in upper trop.
for i,x in enumerate(Pdat):
T[i] = optimize.minimize(delta_Tv, Trho[i], args=(Trho[i],x,qt)).x
if (np.abs(T[i]-Trho[i]) > 10.):
print('correcting T = %7.2f to Trho = %7.3f' % (T[i],Trho[i],))
T[i]=Trho[i]
Tnl[i] = optimize.minimize(delta_Tv, Trho[i], args=(Trho[i],x,qt,False)).x
if (np.abs(T[i]-Trho[i]) > 10.):
print('correcting T = %7.2f to Trho = %7.3f' % (T[i],Trho[i],))
T[i]=Trho[i]
# --------
fig = plt.figure(figsize=(10,5))
ax1 = plt.subplot(1,2,1)
ax1.set_xlabel('K')
ax1.set_ylabel('$P$ / hPa')
ax1.set_ylim(1020,100)
ax1.set_xlim(295,355)
for i,x in enumerate(Pdat):
xfld[i,0] = mt.get_theta_e(Tdat[i],x,Qdat[i])
xfld[i,1] = mt.get_theta_l(Tdat[i],x,Qdat[i])
xfld[i,2] = mt.get_theta_s(Tdat[i],x,Qdat[i])
plt.plot(xfld[:,0],Pdat/100.,c='orange',label='$\\theta_\mathrm{e}$')
plt.plot(xfld[:,1],Pdat/100.,c='teal',label='$\\theta_\mathrm{l}$')
plt.plot(xfld[:,2],Pdat/100.,c='dodgerblue',ls='dashed',label='$\\theta_s$')
plt.legend(loc="upper left")
ax2 = plt.subplot(1,2,2)
ax2.set_xlabel('K')
ax2.set_yticklabels([])
ax2.set_ylim(1020,100)
ax2.set_xlim(295,355)
sounding = 'wet'
Tdat, Pdat, Qdat = Soundings[sounding]
xfld = np.zeros((len(Pdat),4))
qt = 18e-3
Trho = Tdat*(1+mt.eps2*Qdat)
Psat = mt.es(Tdat)
T = np.zeros(len(Pdat))
Tnl = np.zeros(len(Pdat))
Qs = np.maximum((Psat/(Pdat-Psat)) * mt.Rd/mt.Rv * (1-qt),1.e-10) #Psat may be greater than Pdat in upper trop.
for i,x in enumerate(Pdat):
T[i] = optimize.minimize(delta_Tv, Trho[i], args=(Trho[i],x,qt)).x
if (np.abs(T[i]-Trho[i]) > 10.):
print('correcting T = %7.2f to Trho = %7.3f' % (T[i],Trho[i],))
T[i]=Trho[i]
Tnl[i] = optimize.minimize(delta_Tv, Trho[i], args=(Trho[i],x,qt,False)).x
if (np.abs(T[i]-Trho[i]) > 10.):
print('correcting T = %7.2f to Trho = %7.3f' % (T[i],Trho[i],))
T[i]=Trho[i]
for i,x in enumerate(Pdat):
xfld[i,0] = mt.get_theta_e(Tdat[i],x,Qdat[i])
xfld[i,1] = mt.get_theta_l(Tdat[i],x,Qdat[i])
xfld[i,2] = mt.get_theta_s(Tdat[i],x,Qdat[i])
plt.plot(xfld[:,0],Pdat/100.,c='orange',label='$\\theta_\mathrm{e}$')
plt.plot(xfld[:,1],Pdat/100.,c='teal',label='$\\theta_\mathrm{l}$')
plt.plot(xfld[:,2],Pdat/100.,c='dodgerblue',ls='dashed',label='$\\theta_s$')
plt.legend(loc="upper left")
plt.tight_layout()
sns.set_context("talk", font_scale=1.2)
sns.despine(offset=10)
fig.savefig(plot_dir+'jiawei-profiles-marquet.pdf')
Ty1, Py = integrate_dTdP(300.,102000.,15000.,-100.,qt)
Ty2, Py = integrate_dTdP(300.,102000.,15000.,-100.,qt,pseudo=True)
Ty3, Py = integrate_dTdP(300.,102000.,15000.,-100.,qt,es_formula='analytic-mxd',pseudo=True)
Ty4, Py = integrate_dTdP(300.,102000.,15000.,-100.,qt,es_formula='analytic-mxd')
for i,x in enumerate(Py):
if (Ty1[i]<273.16 and Ty4[i] > 273.16): Ty4[i] = 273.16
if (Ty1[i]>273.16): Ty4[i] = Ty1[i]
yfld = np.zeros((len(Py),5))
# --------
fig = plt.figure(figsize=(5,5))
sns.set_context("paper",font_scale=1.2)
for i,x in enumerate(Py):
yfld[i,0] = mt.get_theta_e(Ty1[i],x,qt)
yfld[i,1] = mt.get_theta_e(Ty2[i],x,qt)
yfld[i,2] = mt.get_theta_e(Ty3[i],x,qt)
yfld[i,3] = mt.get_theta_e(Ty4[i],x,qt)
yfld[i,4] = yfld[i,0] - 5.* (1- np.exp((x-102000)/15000.))
ax1 = plt.subplot(1,1,1)
ax1.set_xlabel('K')
ax1.set_ylabel('$P$ / hPa')
ax1.set_ylim(1020,100)
ax1.set_xlim(335,365)
plt.plot(yfld[:,0],Py/100.,c='grey',label='isentrope')
plt.plot(yfld[:,1],Py/100.,c='lightgrey',label='pseudo')
plt.plot(yfld[:,2],Py/100.,c='lightgrey',ls='dashed',label='pseudo-ice')
plt.plot(yfld[:,3],Py/100.,c='grey',ls='dashed',label='isentrope-ice')
plt.plot(yfld[:,4],Py/100.,c='grey',ls='dashdot',label='entraining')
plt.legend(loc="lower right")
#
sns.despine(offset=10)
fig.savefig(plot_dir+'idealized-profiles.pdf')
```
### Checking sensitivities ###
The above showed how different processes affect the vertical structure of the atmosphere as realized in $\theta_\mathrm{e}$ space. Here I check the sensitivities to some assumptions.
1. What if we use a slightly different formulation for the vapor saturation.
2. what if the atmosphere follows a saturated isentrope with $q_\mathrm{t} = 17$ g/kg but we diagnose it assuming a value 20% higher or lower? How does this change the shape of the profile.
3. If we calculate $\theta_\mathrm{e}$ twice and average them how does this compare to the $\theta_\mathrm{e}$ calculated from the average sounding.
The first point was found to be entirely negligible, the difference between constructing the isentrope with the goff-gratch saturation pressure and diagnosing it with magnus is less than 0.1 K. The answer to the second and third questions are quantified by the calculations below with the answers presented quantiatively.
```
qt = 17.e-3
Ty1, Py = integrate_dTdP(300.,102000.,15000.,-10.,qt)
Ty2, Py = integrate_dTdP(300.,102000.,15000.,-10.,13.6e-3)
Ty3, Py = integrate_dTdP(300.,102000.,15000.,-10.,20.4e-3)
Trho = np.zeros(len(Py))
Tm = np.zeros(len(Py))
Tp = np.zeros(len(Py))
for i,x in enumerate(Py):
Ps = mt.es(Ty1[i])
qs = (Ps/(x-Ps)) * mt.Rd/mt.Rv * (1-qt)
if(qt > qs):
ql = qt-qs
qv = qs
else:
ql = 0.
qv = qt
Trho[i] = Ty1[i]*(1 + mt.eps2*np.min([qt,qs]) - ql)
Tm[i] = optimize.minimize(delta_Tv, Trho[i], args=(Trho[i],x,13.6e-3)).x
Tp[i] = optimize.minimize(delta_Tv, Trho[i], args=(Trho[i],x,20.4e-3)).x
yfld = np.zeros((len(Py),5))
# --------
fig = plt.figure(figsize=(5,5))
for i,x in enumerate(Py):
base = mt.get_theta_e(Ty1[i],x,17.0e-3)
yfld[i,0] = mt.get_theta_e(Tm[i],x,13.6e-3) - base
yfld[i,1] = mt.get_theta_e(Tp[i],x,20.4e-3) - base
yfld[i,2] = (mt.get_theta_e(Ty2[i],x,17.0e-3)+mt.get_theta_e(Ty3[i],x,17.0e-3))*0.5 -base
yfld[i,3] = mt.get_theta_e((Ty2[i]+Ty3[i])*0.5,x,17.0e-3) - base
ax1 = plt.subplot(1,1,1)
ax1.set_xlabel('$\\delta \\theta_\mathrm{e}$ / K')
ax1.set_ylabel('$P$ / hPa')
ax1.set_ylim(1010,100)
ax1.set_xlim(-6,3)
plt.plot(yfld[:,0],Py/100.,c='black',label='$q_\mathrm{t} = 13.6$ g/kg')
plt.plot(yfld[:,1],Py/100.,c='lightgrey',label='$q_\mathrm{t} = 20.4$ g/kg')
plt.plot(yfld[:,2],Py/100.,c='dodgerblue',label='$\\overline{\\theta}_\mathrm{e}(T)$')
plt.plot(yfld[:,3],Py/100.,c='orange',label='$\\theta_\mathrm{e}(\\overline{T})$')
plt.legend(loc="upper left")
sns.set_context("paper",font_scale=1.2)
sns.despine(offset=10)
fig.savefig(plot_dir+'sensitivities.pdf')
```
## Calculating the LCL ($P_\mathrm{lcl}$) and density potential temperatures. ##
Here we compare the LCL base predictions to those proposed by Romps and Bolton as well as the differences between density potential temperatures.
For the estimation of the LCL we modify the Romps expressions (using his code) to output pressure at the LCL, as this eliminates an assumption as to how pressure is distributed in the atmosphere, and thus only depends on the parcel state. What we find is that the much simpler Bolton expression is as good as the more complex expression by Romps, and differences between the two are commensurate with those arising from slight differences in how the saturation vapor pressure is calculated.
For the density potential temperatures were write $\theta_\rho$ as $\theta_\mathrm{l}(1+ q_\mathrm{v}/\epsilon_1 - q_\mathrm{t})$ and a naive implementation whereby $$\theta_\mathrm{v}^\mathrm{d} = \theta^\mathrm{d}(1 + \epsilon_2 q_\mathrm{v})$$ or
$$\theta_\mathrm{v}^\mathrm{m} = \theta^\mathrm{m}(1 + \epsilon_2 q_\mathrm{v}),$$ where superscript $m$ of $d$ refers to whether or not the dry or moist value of the specific heats and gas constants are used. These differences are trivial in the absence of saturation, as in that case $\theta_\rho = \theta_\mathrm{v}^\mathrm{m}$ identically. That said it is not clear to me what differences in $\theta_\rho$ tell you in the saturated, case, here one should better compare $T_\rho$ diagnosed at the desired pressure from $\theta_\mathrm{e}.$
```
# Version 1.0 released by David Romps on September 12, 2017.
#
# When using this code, please cite:
#
# @article{16lcl,
# Title = {Exact expression for the lifting condensation level},
# Author = {David M. Romps},
# Journal = {Journal of the Atmospheric Sciences},
# Year = {2017},
# Volume = {in press},
# }
#
# This lcl function returns the height of the lifting condensation level
# (LCL) in meters. The inputs are:
# - p in Pascals
# - T in Kelvins
# - Exactly one of rh, rhl, and rhs (dimensionless, from 0 to 1):
# * The value of rh is interpreted to be the relative humidity with
# respect to liquid water if T >= 273.15 K and with respect to ice if
# T < 273.15 K.
# * The value of rhl is interpreted to be the relative humidity with
# respect to liquid water
# * The value of rhs is interpreted to be the relative humidity with
# respect to ice
# - ldl is an optional logical flag. If true, the lifting deposition
# level (LDL) is returned instead of the LCL.
# - min_lcl_ldl is an optional logical flag. If true, the minimum of the
# LCL and LDL is returned.
def lcl(p,T,rh=None,rhl=None,rhs=None,return_ldl=False,return_min_lcl_ldl=False):
import math
import scipy.special
# Parameters
Ttrip = 273.16 # K
ptrip = 611.65 # Pa
E0v = 2.3740e6 # J/kg
E0s = 0.3337e6 # J/kg
ggr = 9.81 # m/s^2
rgasa = 287.04 # J/kg/K
rgasv = 461 # J/kg/K
cva = 719 # J/kg/K
cvv = 1418 # J/kg/K
cvl = 4119 # J/kg/K
cvs = 1861 # J/kg/K
cpa = cva + rgasa
cpv = cvv + rgasv
# The saturation vapor pressure over liquid water
def pvstarl(T):
return ptrip * (T/Ttrip)**((cpv-cvl)/rgasv) * math.exp( (E0v - (cvv-cvl)*Ttrip) / rgasv * (1/Ttrip - 1/T) )
# The saturation vapor pressure over solid ice
def pvstars(T):
return ptrip * (T/Ttrip)**((cpv-cvs)/rgasv) * math.exp( (E0v + E0s - (cvv-cvs)*Ttrip) / rgasv * (1/Ttrip - 1/T))
# Calculate pv from rh, rhl, or rhs
rh_counter = 0
if rh is not None:
rh_counter = rh_counter + 1
if rhl is not None:
rh_counter = rh_counter + 1
if rhs is not None:
rh_counter = rh_counter + 1
if rh_counter != 1:
print(rh_counter)
exit('Error in lcl: Exactly one of rh, rhl, and rhs must be specified')
if rh is not None:
# The variable rh is assumed to be
# with respect to liquid if T > Ttrip and
# with respect to solid if T < Ttrip
if T > Ttrip:
pv = rh * pvstarl(T)
else:
pv = rh * pvstars(T)
rhl = pv / pvstarl(T)
rhs = pv / pvstars(T)
elif rhl is not None:
pv = rhl * pvstarl(T)
rhs = pv / pvstars(T)
if T > Ttrip:
rh = rhl
else:
rh = rhs
elif rhs is not None:
pv = rhs * pvstars(T)
rhl = pv / pvstarl(T)
if T > Ttrip:
rh = rhl
else:
rh = rhs
if pv > p:
return N
# Calculate lcl_liquid and lcl_solid
qv = rgasa*pv / (rgasv*p + (rgasa-rgasv)*pv)
rgasm = (1-qv)*rgasa + qv*rgasv
cpm = (1-qv)*cpa + qv*cpv
if rh == 0:
return cpm*T/ggr
aL = -(cpv-cvl)/rgasv + cpm/rgasm
bL = -(E0v-(cvv-cvl)*Ttrip)/(rgasv*T)
cL = pv/pvstarl(T)*math.exp(-(E0v-(cvv-cvl)*Ttrip)/(rgasv*T))
aS = -(cpv-cvs)/rgasv + cpm/rgasm
bS = -(E0v+E0s-(cvv-cvs)*Ttrip)/(rgasv*T)
cS = pv/pvstars(T)*math.exp(-(E0v+E0s-(cvv-cvs)*Ttrip)/(rgasv*T))
X = bL/(aL*scipy.special.lambertw(bL/aL*cL**(1/aL),-1).real)
Y = bS/(aS*scipy.special.lambertw(bS/aS*cS**(1/aS),-1).real)
lcl = cpm*T/ggr*( 1 - X)
ldl = cpm*T/ggr*( 1 - Y)
# Modifications of the code to output Plcl or Pldl
Plcl = PPa * X**(cpm/rgasm)
Pldl = PPa * X**(cpm/rgasm)
# Return either lcl or ldl
if return_ldl and return_min_lcl_ldl:
exit('return_ldl and return_min_lcl_ldl cannot both be true')
elif return_ldl:
return Pldl
elif return_min_lcl_ldl:
return min(Plcl,Pldl)
else:
return Plcl
PPa = 1013.25
qt = np.arange(0.5,8,0.2)
TK = 285.
Plcl_X = mt.get_Plcl(TK,PPa,qt,iterate=True)
Plcl_B = mt.get_Plcl(TK,PPa,qt)
Plcl_R = np.zeros(len(Plcl_X))
for i,x in enumerate(qt):
if (x>0.1): x = x/1000.
RH = mt.mr2pp(x/(1.-x),PPa)/mt.es(TK)
Plcl_R[i] = lcl(PPa*100.,TK,RH)*100.
del1 = (Plcl_B-Plcl_X)/100.
del2 = (Plcl_R-Plcl_X)/100.
fig = plt.figure(figsize=(10,5))
ax1 = plt.subplot(1,2,1)
ax1.set_ylabel('$P$ / hPa')
ax1.set_xlabel('$q_\mathrm{t}$ / g/kg')
ax1.set_ylim(-1.2,1.2)
plt.plot(qt,del1,label='$\\delta_\mathrm{B}$, $T$=285K')
plt.plot(qt,del2,label='$\\delta_\mathrm{R}$, $T$=285K')
#plt.gca().invert_yaxis()
plt.legend(loc="best")
qt = np.arange(0.5,28,0.2)
TK = 310.
Plcl_X = mt.get_Plcl(TK,PPa,qt,iterate=True)
Plcl_B = mt.get_Plcl(TK,PPa,qt)
Plcl_R = np.zeros(len(Plcl_X))
for i,x in enumerate(qt):
if (x>0.1): x = x/1000.
RH = mt.mr2pp(x/(1.-x),PPa)/mt.es(TK)
Plcl_R[i] = lcl(PPa*100.,TK,RH)*100.
del1 = (Plcl_B-Plcl_X)/100.
del2 = (Plcl_R-Plcl_X)/100.
ax2 = plt.subplot(1,2,2)
ax2.set_xlabel('$q_\mathrm{t}$ / g/kg')
ax2.set_ylim(-1.2,1.2)
ax2.set_yticklabels([])
plt.plot(qt,del1,label='$\\delta_\mathrm{B}$, $T$=310K')
plt.plot(qt,del2,label='$\\delta_\mathrm{R}$, $T$=310K')
#plt.gca().invert_yaxis()
plt.legend(loc="best")
sns.set_context("talk", font_scale=1.2)
sns.despine(offset=10)
fig.savefig(plot_dir+'Plcl.pdf')
T = 300.
P = 1010.
qt = np.arange(0.5e-3,20e-3,0.2e-3)
theta_rho = mt.get_theta_rho(T,P,qt)
kappa = mt.Rd/mt.cpd
th_v1 = T*(1000./P)**kappa * (1 + mt.eps2*qt)
kappa = (mt.Rd + qt*(mt.Rv - mt.Rd))/(mt.cpd + qt*(mt.cpv - mt.cpd))
th_v2 = T*(1000./P)**kappa * (1 + mt.eps2*qt)
fig = plt.figure(figsize=(10,5))
ax1 = plt.subplot(1,2,1)
ax1.set_ylabel('$\delta \Theta_\mathrm{v}$ / K')
ax1.set_xlabel('$q_\mathrm{t}$ / g/kg')
ax1.set_ylim(-0.01,0.01)
plt.plot(qt*1000.,theta_rho-th_v1,label='$\\theta_\mathrm{v}^{d}(300,1010)$')
plt.plot(qt*1000.,theta_rho-th_v2,label='$\\theta_\mathrm{v}^{m}(300,1010)$')
plt.legend(loc="best")
T = np.arange(290,310,1)
P = 1010.
qt = 10.e-3
theta_rho = mt.get_theta_rho(T,P,qt)
kappa = mt.Rd/mt.cpd
th_v1 = T*(1000./P)**kappa * (1 + mt.eps2*qt)
kappa = (mt.Rd + qt*(mt.Rv - mt.Rd))/(mt.cpd + qt*(mt.cpv - mt.cpd))
th_v2 = T*(1000./P)**kappa * (1 + mt.eps2*qt)
ax2 = plt.subplot(1,2,2)
ax2.set_yticklabels([])
ax2.set_xlabel('$q_\mathrm{t}$ / g/kg')
ax2.set_ylim(-0.01,0.01)
plt.plot(T,theta_rho-th_v1,label='$\\theta_\mathrm{v}^{d}(300,1010)$')
plt.plot(T,theta_rho-th_v2,label='$\\theta_\mathrm{v}^{m}(300,1010)$')
plt.legend(loc="best")
sns.set_context("talk", font_scale=1.2)
sns.despine(offset=10)
fig.savefig(plot_dir+'Plcl.pdf')
```
## Leftovers ##
### Calculating $\theta_e$ and $T_\mathrm{sat}$ ###
Below solve for the saturation temperature and the equivalent potential temperature as a function of the saturation specific humidity. Two lines are shown, with the lcl differing by 50 hPa (925 hPa solid, 975 hPa dashed).
```
def delta_Tv(T,Tv,P,qt):
# """ calculates difference between the virtual temperature in a saturated updraft with a
# specified qt and a given virtual temperture. Here qs<qt is enforced so that setting qt=0
# gets rid of the water loading effects.
# """
ps = mt.es(T)
qs = (ps/(P-ps)) * (mt.Rd/mt.Rv) * (1. - qt)
if np.max(qs) > qt:
qt = qs
err = (T*(1. + mt.eps2*qs - (qt-qs)) - Tv)**2
return err;
def delta_Te(T,Te,P,qt):
# """ calculates difference between some specified theta_e (TE) and that corresponding to a
# given temperature, pressure and qt, assuming saturation allows us to neglect input of qv.
# """
err = (Te - mt.get_theta_e(T,P,qt))**2
return err;
def delta_qs_Te (P,Tguess,Te,qt):
# """ calculates difference between the saturation specific humidity and the total water specific
# humidity for air of a given theta_e (TE), total water, and pressure, which thus gives its
# temperature by minimizing delta_Te
# """
TK = optimize.minimize(delta_Te, Tguess, args=(Te,P,qt)).x
ps = mt.es(TK)
qs = (ps/(P-ps)) * (mt.eps1) * (1. - qt)
err = (qs - qt)**2
return err;
def delta_Ps_TK (TK,Ps):
# """ calculates difference between the saturation specific humidity and that associated with air at
# a given temperature
# """
x = np.asarray(TK)
if np.max(x) < 100:
x = x+273.15
err = (mt.es(x) - Ps)**2
return err;
def get_Tsat(Tguess,P,qs,qt=-999):
# """ calculates temperature at which air is saturated given a pressure and qt
# """
if np.max(qt < 0.0):
qt = qs
Ps = mt.mr2pp(qs/(1.0 - qt),P)
Ts = optimize.minimize(delta_Ps_TK, Tguess, args=(Ps)).x
return Ts;
ax1 = plt.subplot(1,1,1)
ax1.set_ylabel('K')
ax1.set_xlabel('$q_\mathrm{s}$ / gkg$^{-1}$')
P = 92500.
qs = np.arange(10.e-3,25.e-3,0.2e-3)
Ts = np.zeros(len(qs))
Te = np.zeros(len(qs))
for i,x in enumerate(qs):
Ts[i] = get_Tsat(300.,P,x)
Te[i] = mt.get_theta_e(Ts[i],P,x)
plt.plot(qs,Te,label='$\\theta_\mathrm{e}$',c='dodgerblue')
plt.plot(qs,Ts,label='$T_\mathrm{sat}$',c='orange')
#P = 97500.
for i,x in enumerate(qs):
Ts[i] = get_Tsat(300.,P,x)
Te[i] = mt.get_theta_e(Ts[i],P,x)
plt.plot(qs,Te,ls='dashed',c='dodgerblue')
plt.plot(qs,Ts,ls='dashed',c='orange')
plt.legend(loc="upper left")
sns.set_context("talk", font_scale=1.2)
sns.despine(offset=10)
T = 270.
P = 50000.
qt = 17.e-3
ps = mt.es(T)
qs = (ps/(P-ps)) * (mt.Rd/mt.Rv) * (1. - qt)
qt = np.arange(1.2*qs,3*qs, qs/4.)
theta_e = np.zeros(len(qt))
for i,x in enumerate(qt):
theta_e[i] = mt.get_theta_e(T,P,x)
ax1 = plt.subplot(1,1,1)
ax1.set_ylabel('$\Theta_e$ / K')
ax1.set_xlabel('$q_t$ / gkg$^{-1}$')
plt.plot(qt*1000.,theta_e)
sns.set_context("talk", font_scale=1.2)
sns.despine(offset=10)
def Theta(X,x):
if (x == 'e'):
Tx = mt.get_theta_e(X[0],X[1],X[2])
cp = mt.cpd*(1-X[2]) + mt.cl*X[2]
if (x == 'l'):
Tx = mt.get_theta_l(X[0],X[1],X[2])
cp = mt.cpd*(1-X[2]) + mt.cpv*X[2]
if (x == 's'):
Tx = mt.get_theta_s(X[0],X[1],X[2])
cp = mt.cpd
return Tx, cp
def T_from_Tx(Tx,P,q,x):
if (x == 'e'):
T = mt.T_from_Te(Tx,P,q)
if (x == 'l'):
T = mt.T_from_Tl(Tx,P,q)
if (x == 's'):
T = mt.T_from_Ts(Tx,P,q)
return T
def X(T,P,q):
cp = mt.cpd*(1-q) + mt.cl*q
return np.asarray([T,P,q])
kind = 's'
P = 100000
X1 = X(290,P,8.0e-3)
Tx1,cx1 = Theta(X1,kind)
X2 = X(300,P, .0e-3)
Tx2,cx2 = Theta(X2,kind)
qbar = 0.5*(X1[2] + X2[2])
cbar = 0.5*(cx1+cx2)
Txbar = 0.5*(Tx1*cx1 + Tx2*cx2)/cbar
Tbar = T_from_Tx(Txbar,mt.P0,qbar,kind)
print (Txbar,Theta(np.asarray([Tbar,P,qbar]),kind)[0])
Tsbar = mt.get_theta_s(Tbar,mt.P0,qbar)
sbar = mt.cpd*np.log(Tsbar)
Ts1,cs1 = Theta(X1,'s')
Ts2,cs2 = Theta(X2,'s')
s12 = mt.cpd*np.log(0.5*(Ts1+Ts2))
s1 = mt.cpd*np.log(Ts1)
s2 = mt.cpd*np.log(Ts2)
s1 = (mt.sv00 + mt.cpv * np.log(X1[0]/298.15) + mt.Rv * np.log(X1[1]/100000.)) * X1[2] + (mt.sd00 + mt.cpd * np.log(X1[0]/298.15) + mt.Rd * np.log(X1[1]/100000.)) * (1-X1[2])
s2 = (mt.sv00 + mt.cpv * np.log(X2[0]/298.15) + mt.Rv * np.log(X2[1]/100000.)) * X2[2] + (mt.sd00 + mt.cpd * np.log(X2[0]/298.15) + mt.Rd * np.log(X2[1]/100000.)) * (1-X2[2])
X3 = X2*1.
X3[2] = (X1[2]+X2[2])*0.5
X3[0] = ((mt.cpd * (1-X1[2]) + mt.cpv * X1[2])*X1[0] + (mt.cpd * (1-X2[2]) + mt.cpv * X2[2])*X2[0]) / (2.*(mt.cpd * (1-X3[2]) + mt.cpv * X3[2]))
s3 = (mt.sv00 + mt.cpv * np.log(X3[0]/298.15) + mt.Rv * np.log(X3[1]/100000.)) * X3[2] + (mt.sd00 + mt.cpd * np.log(X3[0]/298.15) + mt.Rd * np.log(X3[1]/100000.)) * (1-X3[2])
```
| github_jupyter |
# Doyle-Fuller-Newman Model (DFN)
## Dimensionless Model Equations
The DFN comprises equations for charge and mass conservation in the solid the solid and electrolyte, and also prescribes behaviour for the electrochemical reactions occurring on the interface between the solid an electrolyte. For more information please see [[2]](#References) or other standard texts.
Below we summarise the dimensionless form of the DFN, with all parameters give in the table at the end of this notebook. Here we use a roman subscript $\text{k} \in \text{n, s, p}$ is used to denote the regions negative electrode, separator, and positive electrode, respectively.
The model equations for the DFN read:
#### Charge conservation:
\begin{align}
\frac{\partial i_{\text{e,k}}}{\partial x} &= \begin{cases}
j_{\text{k}}, \quad &\text{k} = \text{n, p}\\
0, \qquad &\text{k} = \text{s}
\end{cases} , && \\
\mathcal{C}_{\text{e}} i_{\text{e,k}} &= \epsilon_{\text{k}}^{\text{b}} \gamma_{\text{e}} \kappa_{\text{e}}(c_{\text{e,k}}) \left( - \frac{\partial \phi_{\text{e,k}}}{\partial x} + 2(1-t^+)\frac{\partial}{\partial x}\left(\log(c_{\text{e,k}})\right)\right), && \text{k} \in \text{n, s, p}, \\
I-i_{\text{e,k}} &= - \sigma_{\text{k}} \frac{\partial \phi_{\text{e,k}}}{\partial x}, && \text{k} \in \text{n, s, p}.
\end{align}
#### Mass conservation:
\begin{align}
\mathcal{C}_{\text{e}} \epsilon_{\text{k}} \gamma_{\text{e}} \frac{\partial c_{\text{e,k}}}{\partial t} &= -\gamma_{\text{e}} \frac{\partial N_{\text{e,k}}}{\partial x} + \mathcal{C}_{\text{e}} \frac{\partial i_{\text{e,k}}}{\partial x}, && \text{k} \in \text{n, s, p},\\
N_{\text{e,k}} &= -\epsilon_{\text{k}}^{\text{b}} D_{\text{e}}(c_{\text{e,k}}) \frac{\partial c_{\text{e,k}}}{\partial x} + \frac{\mathcal{C}_{\text{e}} t^+}{\gamma_{\text{e}}} i_{\text{e,k}},
&& \text{k} \in \text{n, s, p}, \\
\mathcal{C}_{\text{k}} \frac{\partial c_{\text{s,k}}}{\partial t} &= -\frac{1}{r_{\text{k}}^2} \frac{\partial}{\partial r_{\text{k}}} \left(r_{\text{k}}^2 N_{\text{s,k}}\right), && \text{k} \in \text{n, p},\\
N_{\text{s,k}} &= -D_{\text{s,k}}(c_{\text{s,k}}) \frac{\partial c_{\text{s,k}}}{\partial r_{\text{k}}}, && \text{k} \in \text{n, p}.
\end{align}
#### Electrochemical reactions:
\begin{align}
j_{\text{k}} &= 2 j_{\text{0,k}} \sinh\left(\frac{ \eta_{\text{k}}}{2} \right), && \text{k} \in \text{n, p}, \\
j_{\text{0,k}} &= \frac{\gamma_{\text{k}}}{\mathcal{C}_{\text{r,k}}} c_{\text{s,k}}^{1/2} (1-c_{\text{s,k}})^{1/2}c_{\text{e,k}}^{1/2}\big|_{r_{\text{k}}=1}, && \text{k} \in \text{n, p}, \\
\eta_{\text{k}} &= \phi_{\text{s,k}} - \phi_{\text{e,k}} - U_{\text{k}}(c_{\text{s,k}}\big|_{r_{\text{k}}=1}), && \text{k} \in \text{n, p}.
\end{align}
These are to be solved subject to the following boundary conditions:
#### Current:
\begin{gather}
i_{\text{e,n}}\big|_{x=0} = 0, \quad i_{\text{e,p}}\big|_{x=1}=0, \\
\phi_{\text{e,n}}\big|_{x=L_{\text{n}}} = \phi_{\text{e,s}}\big|_{x=L_{\text{n}}}, \quad i_{\text{e,n}}\big|_{x=L_{\text{n}}} = i_{\text{e,s}}\big\vert_{x=L_{\text{n}}} = I, \\
\phi_{\text{e,s}}\big|_{x=1-L_{\text{p}}} = \phi_{\text{e,p}}\big|_{x=1-L_{\text{p}}}, \quad
i_{\text{e,s}}\big|_{x=1-L_{\text{p}}} = i_{\text{e,p}}\big|_{x=1-L_{\text{p}}} = I.
\end{gather}
#### Concentration in the electrolyte:
\begin{gather}
N_{\text{e,n}}\big|_{x=0} = 0, \quad N_{\text{e,p}}\big|_{x=1}=0,\\
c_{\text{e,n}}\big|_{x=L_{\text{n}}} = c_{\text{e,s}}|_{x=L_{\text{n}}}, \quad N_{\text{e,n}}\big|_{x=L_{\text{n}}}=N_{\text{e,s}}\big|_{x=L_{\text{n}}}, \\
c_{\text{e,s}}|_{x=1-L_{\text{p}}}=c_{\text{e,p}}|_{x=1-L_{\text{p}}}, \quad N_{\text{e,s}}\big|_{x=1-L_{\text{p}}}=N_{\text{e,p}}\big|_{x=1-L_{\text{p}}}. &&
\end{gather}
#### Concentration in the electrode active material:
\begin{gather}
N_{\text{s,k}}\big|_{r_{\text{k}}=0} = 0, \quad \text{k} \in \text{n, p}, \quad \ \ - \frac{a_{R, \text{k}}\gamma_{\text{k}}}{\mathcal{C}_{\text{k}}} N_{\text{s,k}}\big|_{r_{\text{k}}=1} = j_{\text{k}}, \quad \text{k} \in \text{n, p}.
\end{gather}
#### Reference potential:
$$\phi_{\text{s,cn}} = 0, \quad \boldsymbol{x} \in \partial \Omega_{\text{tab,n}}.$$
And the initial conditions:
\begin{align}
&c_{\text{s,k}}(x,r,0) = c_{\text{s,k,0}}, \quad \phi_{\text{s,n}}(x,0) = 0, \quad \phi_{\text{s,p}}(x,0) = \phi_{\text{s,p,0}}, && \text{k} \in \text{n, p},\\
&\phi_{\text{e,k}}(x,0) = \phi_{\text{e,0}}, \quad c_{\text{e,k}}(x,0) = 1, && \text{k} \in \text{n, s, p}.
\end{align}
## Example solving DFN using PyBaMM
Below we show how to solve the DFN model, using the default geometry, mesh, parameters, discretisation and solver provided with PyBaMM. For a more detailed example, see the notebook on the [SPM](https://github.com/pybamm-team/PyBaMM/blob/develop/examples/notebooks/models/SPM.ipynb).
In order to show off all the different points at which the process of setting up and solving a model in PyBaMM can be customised we explicitly handle the stages of choosing a geometry, setting parameters, discretising the model and solving the model. However, it is often simpler in practice to use the `Simulation` class, which handles many of the stages automatically, as shown [here](../simulation-class.ipynb).
First we need to import pybamm, along with numpy which we will use in this notebook.
```
%pip install pybamm -q # install PyBaMM if it is not installed
import pybamm
import numpy as np
```
We then load the DFN model and default geometry, and process them both using the default parameters.
```
# load model
model = pybamm.lithium_ion.DFN()
# create geometry
geometry = model.default_geometry
# load parameter values and process model and geometry
param = model.default_parameter_values
param.process_model(model)
param.process_geometry(geometry)
```
The next step is to set the mesh and discretise the model. Again, we choose the default settings.
```
# set mesh
mesh = pybamm.Mesh(geometry, model.default_submesh_types, model.default_var_pts)
# discretise model
disc = pybamm.Discretisation(mesh, model.default_spatial_methods)
disc.process_model(model);
```
The model is now ready to be solved. We select the default DAE solver for the DFN. Note that in order to successfully solve the system of DAEs we are required to give consistent initial conditions. This is handled automatically by PyBaMM during the solve operation.
```
# solve model
solver = model.default_solver
t_eval = np.linspace(0, 3600, 300) # time in seconds
solution = solver.solve(model, t_eval)
```
To get a quick overview of the model outputs we can use the QuickPlot class, which plots a common set of useful outputs. The method `Quickplot.dynamic_plot` makes a slider widget.
```
quick_plot = pybamm.QuickPlot(solution, ["Positive electrode interfacial current density [A.m-2]"])
quick_plot.dynamic_plot();
```
## Dimensionless Parameters
In the table below, we provide the dimensionless parameters in the DFN in terms of the dimensional parameters in mcmb2528_lif6-in-ecdmc_lico2_parameters_Dualfoil.csv. We use a superscript * to indicate dimensional quantities.
| Parameter | Expression |Interpretation |
|:--------------------------|:----------------------------------------|:------------------------------------------|
| $L_{\text{k}}$ | $L_{\text{k}}^*/L^*$ | Ratio of region thickness to cell thickness|
|$\sigma_{\text{k}}$ | $\sigma_{\text{k}}^*R^* T^*/(I^*F^*L^*)$| Dimensionless solid conductivity |
|$\mathcal{C}_{\text{k}}$ | $\tau_{\text{k}}^*/\tau_{\text{d}}^*$ | Ratio of solid diffusion and discharge timescales |
|$\mathcal{C}_{\text{e}}$ |$\tau_{\text{e}}^*/\tau_{\text{d}}^*$ |Ratio of electrolyte transport and discharge timescales|
|$\mathcal{C}_{\text{r,k}}$ |$\tau_{\text{r,k}}^*/\tau_{\text{d}}^*$ |Ratio of reaction and discharge timescales|
|$a_{R, \text{k}}$ |$a_{\text{k}}^* R_{\text{k}}^*$ | Product of particle radius and surface area to volume ratio|
|$\gamma_{\text{k}}$ |$c_{\text{k,max}}^*/c_{\text{n,max}}^*$ |Ratio of maximum lithium concentrations in solid|
|$\gamma_{\text{e}}$ |$c_{\text{e,typ}}^*/c_{\text{n,max}}^*$ |Ratio of maximum lithium concentration in the negative electrode solid and typical electrolyte concentration|
Note that the dimensionless parameters $\epsilon_{\text{k}}$, $\text{b}$, and $t^+$ are already provided in the parameter file mcmb2528_lif6-in-ecdmc_lico2_parameters_Dualfoil.csv
## References
The relevant papers for this notebook are:
```
pybamm.print_citations()
```
| github_jupyter |
```
import numpy as np
from random import shuffle
from math import log, floor
import pandas as pd
import tensorflow as tf
import tensorboard as tb
from keras import backend as K
from keras.models import *
from keras.layers import *
from keras.activations import *
from keras.callbacks import *
from keras.utils import *
from keras.layers.advanced_activations import *
# from keras.layers.advanced_activations import *
from keras import *
from keras.engine.topology import *
from keras.optimizers import *
import keras
# import pandas as pd
# import numpy as np
# import sklearn
import pickle
from keras.applications import *
from keras.preprocessing.image import *
from keras.losses import mse, binary_crossentropy
import pandas as pd # data frame
import numpy as np # matrix math
from scipy.io import wavfile # reading the wavfile
from sklearn.utils import shuffle # shuffling of data
from random import sample # random selection
from tqdm import tqdm # progress bar
import matplotlib.pyplot as plt # to view graphs
import wave
from math import log, floor
# audio processing
from scipy import signal # audio processing
from scipy.fftpack import dct
import librosa # library for audio processing
import numpy as np
import pandas as pd
from sklearn.decomposition import *
from sklearn.cluster import KMeans
import sys, os
import random,math
from tqdm import tqdm ##
from xgboost.sklearn import XGBClassifier
from sklearn.utils import shuffle # shuffling of data
from random import sample # random selection
from tqdm import tqdm # progress bar
# audio processing
from scipy import signal # audio processing
from scipy.fftpack import dct
import librosa # library for audio processing
import xgboost as xgb
import lightgbm as lgb
import catboost as ctb
from keras.utils import *
from sklearn.ensemble import *
import pickle
from bayes_opt import BayesianOptimization
from logHandler import Logger
from utils import readCSV, getPath, writePickle,readPickle
from keras.regularizers import l2
from keras.callbacks import History ,ModelCheckpoint, EarlyStopping
mean = np.load('feature/fbank4/mean.npy')
std = np.load('feature/fbank4/std.npy')
min_ = np.load('feature/fbank4/min.npy')
range_ = np.load('feature/fbank4/range.npy')
# autoencoder = load_model('model/lgd_dense_AE3.h5')
# X_un_dense = np.load('feature/fbank2/semi/X_un_dense.npy')
# Y_un_dense = np.load('feature/fbank2/semi/Y_un_dense.npy')
# X_test_dense = np.load('feature/fbank2/semi/X_test_dense.npy')
# Y_test_dense = np.load('feature/fbank2/semi/Y_test_dense.npy')
base_path = 'feature/fbank4/'#'/tmp2/b03902110/newphase1'
base_data_path = 'feature/fbank4/'#os.path.join(base_path, 'data')
num_fold = 10
def _shuffle(X, Y):
randomize = np.arange(len(X))
np.random.shuffle(randomize)
# print(X.shape, Y.shape)
return (X[randomize], Y[randomize])
def getTrainData():
X = []
y = []
for i in range(num_fold):
fileX = os.path.join(base_data_path, 'X/X' + str(i+1) + '.npy')
fileY = os.path.join(base_data_path, 'y/y' + str(i+1) + '.npy')
X.append(np.load(fileX))
y.append(np.load(fileY))
X = np.array(X)
y = np.array(y)
return X, y
def split_data(X, y, idx):
X_train = []
y_train = []
for i in range(num_fold):
if i == idx:
X_val = X[i]
y_val = y[i]
continue
if X_train == []:
X_train = X[i]
y_train = y[i]
else:
X_train = np.concatenate((X_train, X[i]))
y_train = np.concatenate((y_train, y[i]))
return X_train, y_train, X_val, y_val
def normalize(X_train, X_val):
X_train = (X_train - mean)/(std)
# X_train = (X_train - min_)/range_
X_val = (X_val - mean)/(std)
# X_val = (X_val - min_)/range_
return X_train, X_val
def get_model():
first_size = 11
second_size = 8
input_img = Input(shape=(X_train.shape[1],X_train.shape[2],X_train.shape[3]))
#block1
conv = Conv2D(32,(first_size,second_size),padding='same',kernel_initializer='glorot_normal')(input_img)
act = PReLU(alpha_initializer='zeros', alpha_regularizer=None, alpha_constraint=None,
shared_axes=None)(conv)
# pad = ZeroPadding2D(padding=(2, 2), data_format='channels_last')(act)
bn = BatchNormalization()(act)
dp1 = Dropout(0.25)(bn)
# add = Add()([])
conv = Conv2D(32,(first_size,second_size),padding='same',kernel_initializer='glorot_normal')(dp1)
act = LeakyReLU(alpha=0.05)(conv)
bn = BatchNormalization()(act)
# pad = ZeroPadding2D(padding=(1, 1), data_format='channels_last')(bn)
# maxpool = MaxPooling2D(pool_size=(2,2))(bn)
dp2 = Dropout(0.3)(bn)
# block2
add = Add()([dp1,dp2])
conv = Conv2D(32,(first_size,second_size),padding='same',kernel_initializer='glorot_normal')(add)
act = LeakyReLU(alpha=0.33)(conv)
bn = BatchNormalization()(act)
# pad = ZeroPadding2D(padding=(1, 1), data_format='channels_last')(bn)
dp1 = Dropout(0.35)(bn)
add = Add()([dp1,dp2])
conv = Conv2D(32,(first_size,second_size),padding='same',kernel_initializer='glorot_normal')(add)
act = LeakyReLU(alpha=0.33)(conv)
# pad = ZeroPadding2D(padding=(2, 2), data_format='channels_last')(act)
bn = BatchNormalization()(act)
# maxpool = MaxPooling2D(pool_size=(2,2))(bn)
dp2 = Dropout(0.4)(bn)
#block 3
add = Add()([dp1,dp2])
conv = Conv2D(32,(first_size,second_size),padding='same',kernel_initializer='glorot_normal')(add)
act = LeakyReLU(alpha=0.33)(conv)
bn = BatchNormalization()(act)
# pad = ZeroPadding2D(padding=(1, 1), data_format='channels_last')(bn)
dp1 = Dropout(0.45)(bn)
add = Add()([dp1,dp2])
conv = Conv2D(32,(first_size,second_size),padding='same',kernel_initializer='glorot_normal')(add)
act = LeakyReLU(alpha=0.33)(conv)
# pad = ZeroPadding2D(padding=(2, 2), data_format='channels_last')(act)
bn = BatchNormalization()(act)
# maxpool = MaxPooling2D(pool_size=(2,2))(bn)
dp2 = Dropout(0.5)(bn)
#block4
add = Add()([dp1,dp2])
conv = Conv2D(32,(int(first_size/2)+1,int(second_size/2)+1),padding='same',
kernel_initializer='glorot_normal')(add)
bn = BatchNormalization()(conv)
act = LeakyReLU(alpha=0.33)(bn)
# pad = ZeroPadding2D(padding=(2, 2), data_format='channels_last')(act)
dp1 = Dropout(0.55)(act)
add = Add()([dp1,dp2])
conv = Conv2D(32,(int(first_size/2)+1,int(second_size/2)+1),padding='same',
kernel_initializer='glorot_normal')(add)
bn = BatchNormalization()(conv)
act = LeakyReLU(alpha=0.33)(bn)
# pad = ZeroPadding2D(padding=(1, 1), data_format='channels_last')(bn)
# avgpool = AveragePooling2D(pool_size=(1,4))(act)
# maxpool = MaxPooling2D(pool_size=(2,2))(bn)
cnn = Dropout(0.6)(act)
shape = K.int_shape(cnn)
# clf block
x = Reshape((shape[1],shape[2]*shape[3]))(cnn)
# x = TimeDistributed(Dense(128),input_shape=(11,2,256))(cnn)
x = LSTM(1024,return_sequences=True,dropout=0.75,recurrent_dropout=0.75,
kernel_initializer='lecun_normal')(x)
res1 = GRU(512,return_sequences=False,dropout=0.55,recurrent_dropout=0.55,
kernel_initializer='lecun_normal')(x)
# x = Concatenate()([x,encoder])
x = BatchNormalization()(res1)
'''
for i in range(101):
x_ = Dense(512, activation='selu', kernel_initializer='lecun_normal',
kernel_regularizer=l2(1e-4))(x)
x_ = Dropout(0.3)(x_)
x = Add()([x_, x])
x = BatchNormalization()(x)
'''
for i in range(2):
res2 = Dense(256, activation='selu', kernel_initializer='lecun_normal',
kernel_regularizer=l2(3e-4))(x)
x = Dropout(0.45)(res2)
x = Concatenate()([x,res1])
x = BatchNormalization()(x)
res1 = Dense(256, activation='selu', kernel_initializer='lecun_normal',
kernel_regularizer=l2(3e-4))(x)
x = Dropout(0.45)(res1)
x = Concatenate()([x,res2])
x = BatchNormalization()(x)
# res2 = Dense(256, activation='selu', kernel_initializer='lecun_normal')(x)
# x = Dropout(0.25)(res2)
# x = Concatenate()([x,res1])
# x = BatchNormalization()(x)
# res1 = Dense(256, activation='selu', kernel_initializer='lecun_normal')(x)
# x = Dropout(0.25)(res1)
# x = Concatenate()([x,res2])
# x = BatchNormalization()(x)
# res2 = Dense(128, activation='selu', kernel_initializer='lecun_normal')(x)
# x = Dropout(0.25)(res2)
# x = Concatenate()([x,res1])
# x = BatchNormalization()(x)
x = Dense(64, activation='selu', kernel_initializer='lecun_normal')(x)
# x = Dropout(0.25)(x)
clf = Dense(41,activation='softmax',name='clf')(x)
model = Model(inputs=input_img, outputs=clf)
# model = Model(input_img,decoder)
model.summary()
return model
# val_set_num = [x+1 for x in range(num_fold)]#str(sys.argv[1])
val_set_num = [0]#str(sys.argv[1])
min_ = np.swapaxes(min_,0,1)
mean = np.swapaxes(mean,0,1)
range_ = np.swapaxes(range_,0,1)
std = np.swapaxes(std,0,1)
for fold in val_set_num:
X, y = getTrainData()
X = np.swapaxes(X,2,3)
X_train, Y_train, X_valid, Y_valid = split_data(X, y, fold) #fold
X_train, X_valid = normalize(X_train, X_valid)
print(X_train.shape, Y_train.shape)
model = get_model()
batchSize=[128,256]
batchSize = random.choice(batchSize)
patien=100
epoch=3000
saveP = 'model/LGD_ConvLSTM_clf_'+str(fold)+'.h5'
model.compile(loss=['categorical_crossentropy'],optimizer=Adam(lr=3e-3,decay=1e-8),
metrics=['acc'])
callback=[
ReduceLROnPlateau(monitor='val_loss', factor=0.5, patience=int(patien/2),
min_lr=1e-4,mode='min'),
EarlyStopping(patience=patien,monitor='val_acc',verbose=1,mode='max'),
ModelCheckpoint(saveP,monitor='val_acc',verbose=1,save_best_only=True,
save_weights_only=False,mode='max'),
]
model.fit(X_train,Y_train,
shuffle=True,
callbacks=callback,
class_weight='auto',
validation_data=(X_valid,Y_valid),
batch_size=batchSize,
epochs=epoch)
#1 =>0.41
#2 =>0.51482
#3 =>0.45822
#4 =>0.49596
#5 =>0.50674
#6 =>0.33154
#7 =>0.54717
#8 => 0.37736
#9 => 0.40431
#0 => 0.41240
first_size = 8
second_size = 8
input_img = Input(shape=(X_train.shape[1],X_train.shape[2],X_train.shape[3]))
#block1
conv = Conv2D(32,(first_size,second_size),padding='same',kernel_initializer='glorot_normal')(input_img)
act = PReLU(alpha_initializer='zeros', alpha_regularizer=None, alpha_constraint=None,
shared_axes=None)(conv)
bn = BatchNormalization()(act)
dp = Dropout(0.25)(bn)
conv = Conv2D(32,(first_size,second_size),padding='same',kernel_initializer='glorot_normal')(dp)
act = PReLU(alpha_initializer='zeros', alpha_regularizer=None, alpha_constraint=None,
shared_axes=None)(conv)
bn = BatchNormalization()(act)
maxpool = MaxPooling2D(pool_size=(2,2))(bn)
dp = Dropout(0.25)(maxpool)
# block2
conv = Conv2D(64,(first_size,second_size),padding='same',kernel_initializer='glorot_normal')(dp)
act = LeakyReLU(alpha=0.33)(conv)
bn = BatchNormalization()(act)
dp = Dropout(0.25)(bn)
conv = Conv2D(64,(first_size,second_size),padding='same',kernel_initializer='glorot_normal')(dp)
act = LeakyReLU(alpha=0.33)(conv)
bn = BatchNormalization()(act)
maxpool = MaxPooling2D(pool_size=(2,2))(bn)
dp = Dropout(0.25)(maxpool)
#block 3
conv = Conv2D(128,(first_size,second_size),padding='same',kernel_initializer='glorot_normal')(dp)
act = LeakyReLU(alpha=0.33)(conv)
bn = BatchNormalization()(act)
dp = Dropout(0.25)(bn)
conv = Conv2D(128,(first_size,second_size),padding='same',kernel_initializer='glorot_normal')(dp)
act = LeakyReLU(alpha=0.33)(conv)
bn = BatchNormalization()(act)
maxpool = MaxPooling2D(pool_size=(2,2))(bn)
dp = Dropout(0.25)(maxpool)
#block4
conv = Conv2D(256,(int(first_size/2),int(second_size/2)),padding='same',
kernel_initializer='glorot_normal')(dp)
bn = BatchNormalization()(conv)
act = LeakyReLU(alpha=0.33)(bn)
dp = Dropout(0.25)(act)
conv = Conv2D(256,(int(first_size/2),int(second_size/2)),padding='same',
kernel_initializer='glorot_normal')(dp)
bn = BatchNormalization()(conv)
act = LeakyReLU(alpha=0.33)(bn)
avgpool = AveragePooling2D(pool_size=(1,4))(act)
# maxpool = MaxPooling2D(pool_size=(2,2))(bn)
cnn = Dropout(0.25)(avgpool)
# AUTO ENCODER BLOCK
shape = K.int_shape(cnn)
flat = Flatten()(cnn)
x = Dense(4096)(flat)
x = BatchNormalization()(x)
x = LeakyReLU(alpha=0.33)(x)
encoder = Dense(2048,name='encoder',activation='selu')(x)
d = Dense(4096)(encoder)
d = LeakyReLU(alpha=0.33)(d)
d = Dense(shape[1] * shape[2] * shape[3])(d)
d = Reshape((shape[1], shape[2], shape[3]))(d)
d = UpSampling2D((1,4))(d)
d = Conv2DTranspose(256,(int(first_size/2),int(second_size/2)),padding='same',
kernel_initializer='glorot_normal')(d)
d = LeakyReLU(alpha=0.33)(d)
d = Conv2DTranspose(256,(int(first_size/2),int(second_size/2)),padding='same',
kernel_initializer='glorot_normal')(d)
d = LeakyReLU(alpha=0.33)(d)
d = UpSampling2D((2,2))(d)
d = Conv2DTranspose(128,(first_size,second_size),padding='same',
kernel_initializer='glorot_normal')(d)
d = LeakyReLU(alpha=0.33)(d)
d = Conv2DTranspose(128,(first_size,second_size),padding='same',
kernel_initializer='glorot_normal')(d)
d = LeakyReLU(alpha=0.33)(d)
d = UpSampling2D((2,2))(d)
d = Conv2DTranspose(64,(first_size,second_size),padding='same',
kernel_initializer='glorot_normal')(d)
d = LeakyReLU(alpha=0.33)(d)
d = Conv2DTranspose(64,(first_size,second_size),padding='same',
kernel_initializer='glorot_normal')(d)
d = LeakyReLU(alpha=0.33)(d)
d = UpSampling2D((2,2))(d)
d = Conv2DTranspose(32,(first_size,second_size),padding='same',
kernel_initializer='glorot_normal')(d)
d = LeakyReLU(alpha=0.33)(d)
d = Conv2DTranspose(32,(first_size,second_size),padding='same',
kernel_initializer='glorot_normal')(d)
d = LeakyReLU(alpha=0.33)(d)
decoder = Conv2DTranspose(1,(first_size,second_size),padding='same',
kernel_initializer='glorot_normal',name='AE')(d)
# = LeakyReLU(name='AE',alpha=0.33)(d)
# d = Conv2DTranspose(1,(first_size,second_size),padding='same',kernel_initializer='glorot_normal')(d)
# decoder = LeakyReLU(alpha=0.33)(d)
# clf block
x = Reshape((shape[1],shape[2]*shape[3]))(cnn)
# x = TimeDistributed(Dense(128),input_shape=(11,2,256))(cnn)
x = LSTM(512,return_sequences=True,dropout=0.55,recurrent_dropout=0.55,
kernel_initializer='lecun_normal')(x)
res1 = GRU(512,return_sequences=False,dropout=0.55,recurrent_dropout=0.55,
kernel_initializer='lecun_normal')(x)
# x = Concatenate()([x,encoder])
x = BatchNormalization()(res1)
'''
for i in range(101):
x_ = Dense(512, activation='selu', kernel_initializer='lecun_normal',
kernel_regularizer=l2(1e-4))(x)
x_ = Dropout(0.3)(x_)
x = Add()([x_, x])
x = BatchNormalization()(x)
'''
for i in range(16):
res2 = Dense(512, activation='selu', kernel_initializer='lecun_normal',
kernel_regularizer=l2(5e-5))(x)
x = Dropout(0.28)(res2)
x = Concatenate()([x,res1])
x = BatchNormalization()(x)
res1 = Dense(512, activation='selu', kernel_initializer='lecun_normal',
kernel_regularizer=l2(5e-5))(x)
x = Dropout(0.28)(res1)
x = Concatenate()([x,res2])
x = BatchNormalization()(x)
# res2 = Dense(256, activation='selu', kernel_initializer='lecun_normal')(x)
# x = Dropout(0.25)(res2)
# x = Concatenate()([x,res1])
# x = BatchNormalization()(x)
# res1 = Dense(256, activation='selu', kernel_initializer='lecun_normal')(x)
# x = Dropout(0.25)(res1)
# x = Concatenate()([x,res2])
# x = BatchNormalization()(x)
# res2 = Dense(128, activation='selu', kernel_initializer='lecun_normal')(x)
# x = Dropout(0.25)(res2)
# x = Concatenate()([x,res1])
# x = BatchNormalization()(x)
# x = Dense(128, activation='selu', kernel_initializer='lecun_normal')(x)
# x = Dropout(0.25)(x)
clf = Dense(41,activation='softmax',name='clf')(x)
model = Model(inputs=input_img, outputs=clf)
# model = Model(input_img,decoder)
model.summary()
model.compile(loss=['categorical_crossentropy'],optimizer=Adam(),metrics=['acc']) #,loss_weights=[0,1
model.fit(X_train,Y_train,
shuffle=True,
validation_data=(X_valid,Y_valid),
batch_size=128,epochs=500)
first_size = 8
second_size = 8
input_img = Input(shape=(X_train.shape[1],X_train.shape[2],X_train.shape[3]))
#block1
conv = Conv2D(32,(first_size,second_size),padding='same',activation='relu')(input_img)
conv = Conv2D(32,(first_size,second_size),padding='same',activation='relu')(conv)
maxpool = MaxPooling2D(pool_size=(2,2))(conv)
# block2
conv = Conv2D(64,(first_size,second_size),padding='same',activation='relu')(maxpool)
conv = Conv2D(64,(first_size,second_size),padding='same',activation='relu')(conv)
cnn = MaxPooling2D(pool_size=(2,2))(conv)
#block 3
# conv = Conv2D(128,(first_size,second_size),padding='same',activation='relu')(maxpool)
# conv = Conv2D(128,(first_size,second_size),padding='same',activation='relu')(conv)
# cnn = MaxPooling2D(pool_size=(2,2))(conv)
#block4
# conv = Conv2D(256,(int(first_size/2),int(second_size/2)),padding='same',activation='relu' )(maxpool)
# conv = Conv2D(256,(int(first_size/2),int(second_size/2)),padding='same',activation='relu')(conv)
# cnn = AveragePooling2D(pool_size=(1,4))(conv)
# AUTO ENCODER BLOCK
shape = K.int_shape(cnn)
flat = Flatten()(cnn)
x = Dense(4096,activation='relu')(flat)
encoder = Dense(2048,name='encoder')(x)
d = Dense(4096,activation='relu')(encoder)
d = Dense(shape[1] * shape[2] * shape[3])(d)
d = Reshape((shape[1], shape[2], shape[3]))(d)
# d = UpSampling2D((1,4))(d)
# d = Conv2DTranspose(256,(int(first_size/2),int(second_size/2)),padding='same',activation='relu')(d)
# d = Conv2DTranspose(256,(int(first_size/2),int(second_size/2)),padding='same',activation='relu')(d)
d = UpSampling2D((2,2))(d)
d = Conv2DTranspose(128,(first_size,second_size),padding='same',activation='relu')(d)
d = Conv2DTranspose(128,(first_size,second_size),padding='same',activation='relu')(d)
d = UpSampling2D((2,2))(d)
d = Conv2DTranspose(64,(first_size,second_size),padding='same',activation='relu')(d)
d = Conv2DTranspose(64,(first_size,second_size),padding='same',activation='relu')(d)
d = UpSampling2D((2,2))(d)
d = Conv2DTranspose(32,(first_size,second_size),padding='same',activation='relu')(d)
d = Conv2DTranspose(32,(first_size,second_size),padding='same',activation='relu')(d)
decoder = Conv2DTranspose(1,(first_size,second_size),padding='same',name='AE')(d)
# = LeakyReLU(name='AE',alpha=0.33)(d)
# d = Conv2DTranspose(1,(first_size,second_size),padding='same')(d)
# decoder = LeakyReLU(alpha=0.33)(d)
# clf block
# x = Reshape((shape[1]*shape[2]*shape[3]))(cnn)
x = Flatten()(cnn)
# x = TimeDistributed(Dense(128),input_shape=(11,2,256))(cnn)
# x = LSTM(512,return_sequences=True)(x)
# x = GRU(512,return_sequences=False)(x)
# x = Concatenate()([x,encoder])
x = Dense(2048, activation='relu')(x)
x = Dense(2048, activation='relu')(x)
x = Dense(256, activation='relu')(x)
x = Dense(256, activation='relu')(x)
x = Dense(128, activation='relu')(x)
x = Dense(128, activation='relu')(x)
clf = Dense(41,activation='softmax',name='clf')(x)
model = Model(inputs=input_img, outputs=clf)
# model = Model(input_img,decoder)
model.summary()
model.predict(X_valid)
Y_train
```
| github_jupyter |
```
# modules for structure decoration
import pandas as pd
import os
import itertools
from tqdm.notebook import tqdm
import networkx as nx
import glob
from glob import iglob
from copy import deepcopy
from collections import defaultdict
from pymatgen.core import Composition, Structure
from pymatgen.analysis import local_env
# print(f"pymatgen version: {pymatgen.__version__}")
import pip
pip.main(["show","pymatgen"])
os.chdir('..')
os.getcwd()
```
## Find the crystal structure with the maximum volume around the conducting ions
Action space:
1. Choose the elements desired for the battery material i.e., conducting ion, framework cation(s), and anion(s)
2. For a given combination of elements, randomly select a composition from one of the valence-balanced compounds available as a lookup table (see 3.1).
3. For the selected composition type, a number of prototype structures are available, which will be classified by their crystal system (cubic, hexagonal, ...). Choose a crystal system randomly.
4. For a chosen crystal system, consider all the prototypes and construct hypothetical decorated structures.
5. Compute the volume around the conducting ions
### Compositions
Elements commonly found in battery materials:
- Conducting ion (C): Li+, Na+, K+, Mg2+, Zn2+
- Anion (A): F-, Cl-, Br-, I-, O2-, S2-, N3-, P3-
- Framework cation (F): Sc3+, Y3+, La3+, Ti4+, Zr4+, Hf4+, W6+, Zn2+, Cd2+, Hg2+, B3+, Al3+, Si4+, Ge4+, Sn4+, P5+, Sb5+
Hypothetical compositions using combinations of C, F, and A are of the following forms:
1. Cx Az
2. Cx A1z1 A2z2
3. Cx Fy Az
4. Cx Fy A1z1 A2z2
5. Cx F1y1 F2y2 Az
6. Cx F1y1 F2y2 A1z1 A2z2
The following constraints are employed in generating the compositions:
1. A composition may contain only one C ion, up to two (0-2) F ions and at least one and up to two (1-2) A ions.
2. The sum of stoichiometric coefficients of the ions is less than or equal to ten, i.e., x + y1 + y2 + z1 + z2 ≤ 10 .
3. The generated compositions are valence-balanced, i.e., stoichiometric sum of oxidation states of ions equals to 0.
### Build Action space graph
We are going to build a networkx graph of all the possible actions, split into two parts. Actions 1-2, and actions 3-5.
```
# want to maximize the volume around only the conducting ions
conducting_ions = set(['Li', 'Na', 'K', 'Mg', 'Zn'])
anions = set(['F', 'Cl', 'Br', 'I', 'O', 'S', 'N', 'P'])
framework_cations = set(['Sc', 'Y', 'La', 'Ti', 'Zr', 'Hf', 'W', 'Zn', 'Cd', 'Hg', 'B', 'Al', 'Si', 'Ge', 'Sn', 'P', 'Sb'])
elements = conducting_ions | anions | framework_cations
# sort by the length of the string, so that multiple letter elements come first
elements = sorted(elements, key=len, reverse=True)
print(elements)
anions & framework_cations
conducting_ions & framework_cations
G = nx.DiGraph()
# the first state will just be the string 'root'
root_node = "root"
for c in conducting_ions:
G.add_edge(root_node, c)
```
For the reinforcement learning to better distinguish between the combination of elements, adding an element will be a specific action
For example:
1. Choose a conducting ion
2. Choose an anion
3. Possibly add a framework cation
4. Possibly add another anion
5. Possibly add another framework cation
```
def build_element_combination_actions(G):
"""
*G*: networkx DiGraph of crystal structure actions
"""
for c in conducting_ions:
# 1. Cx Az
for a1 in anions - {c}:
# since the ordering of the elements doesn't matter,
# only store the sorted version of the elements
c_a1 = tuple(sorted(set((c, a1))))
# add edge from c to (c a1)
G.add_edge(c, c_a1)
# 2. Cx A1z1 A2z2
for a2 in anions - set(c_a1):
c_a1_a2 = tuple(sorted(set((c, a1, a2))))
G.add_edge(c_a1, c_a1_a2)
# 4. Cx Fy A1z1 A2z2
for f1 in framework_cations - set(c_a1_a2):
c_f1_a1_a2 = tuple(sorted(set((c, f1, a1, a2))))
G.add_edge(c_a1_a2, c_f1_a1_a2)
# 6. Cx F1y1 F2y2 A1z1 A2z2
for f2 in framework_cations - set(c_f1_a1_a2):
c_f1_f2_a1_a2 = tuple(sorted(set((c, f1, f2, a1, a2))))
G.add_edge(c_f1_a1_a2, c_f1_f2_a1_a2)
# 3. Cx Fy Az
for f1 in framework_cations - set(c_a1):
c_f1_a1 = tuple(sorted(set((c, f1, a1))))
G.add_edge(c_a1, c_f1_a1)
# 5. Cx F1y1 F2y2 Az
for f2 in framework_cations - set(c_f1_a1):
c_f1_f2_a1 = tuple(sorted(set((c, f1, f2, a1))))
G.add_edge(c_f1_a1, c_f1_f2_a1)
build_element_combination_actions(G)
print(f'{G.number_of_nodes()} nodes, {G.number_of_edges()} edges')
# working_dir = "/projects/rlmolecule/shubham/file_transfer/decorations/jupyter_demo"
working_dir = "../../rlmolecule/crystal/inputs"
# read-in the dataframe containing all compositions to decorate
df_comp = pd.read_csv(f'{working_dir}/compositions.csv.gz')
df_comp
compositions = df_comp['composition'].to_list()
comp_types = set(df_comp['comp_type'].to_list())
comp_to_comp_type = dict(zip(df_comp['composition'], df_comp['comp_type']))
len(str(df_comp['stoichiometry'][0]))
# strip the compositions to just the atoms
comp_elements = defaultdict(set)
for c in compositions:
orig_c = c
ele_in_comp = []
# these elements are sorted such that the double letter elements come after single letter ones
for e in elements:
if e in c:
ele_in_comp.append(e)
# make sure a single letter element doesn't also match (e.g., Si vs S)
c = c.replace(e,'')
comp_elements[tuple(sorted(ele_in_comp))].add(orig_c)
# print(ele_in_comp)
# print(orig_c)
print(len(comp_elements))
# not all of the generated compositions are valence-balanced, i.e., stoichiometric sum of oxidation states of ions equals 0.
# Shubham already computed which combinations are valid, so limit those here
ele_combo_with_comp = set()
ele_combo_without_comp = set()
# the nodes that are element combinations are tuples
graph_ele_combos = [n for n in G.nodes() if isinstance(n, tuple)]
for ele_combo in graph_ele_combos:
if ele_combo in comp_elements:
ele_combo_with_comp.add(ele_combo)
else:
ele_combo_without_comp.add(ele_combo)
print(f"{len(ele_combo_with_comp)} out of {len(graph_ele_combos)} have a corresponding composition")
# delete the non-valid element combinations from the graph
G.remove_nodes_from(ele_combo_without_comp)
print(f"{len(G.nodes())} nodes remaining")
# Step 2: For a given combination of elements, randomly select a composition from one of the valence-balanced compounds available as a lookup table
# As a sanity check, see how many compositions there are for a given combination of elements (top 20):
print(sorted([len(vals) for vals in comp_elements.values()], reverse=True)[:20])
for eles, comps in comp_elements.items():
break
print(eles, comps)
# add the edges from the element combinations to the compositions
for ele_combo, comps in comp_elements.items():
for comp in comps:
G.add_edge(ele_combo, comp)
print(f'{G.number_of_nodes()} nodes, {G.number_of_edges()} edges')
# write this network as the action tree
out_file = f"{working_dir}/elements_to_compositions.edgelist.gz"
print(f"writing graph to {out_file}")
nx.write_edgelist(G, out_file, delimiter='\t', data=False)
# lets select the first one as an example:
curr_comp = next(iter(comps))
print(comp)
# get the comp_type for this comp
curr_comp_type = comp_to_comp_type[comp]
print(curr_comp_type)
```
## Second half of graph structure
- For the secod half of the search space, start from the composition type, and map out to the decorations.
- We will be able to use this graph to get to the final state after the composition (first graph) has been decided
### Step 3:
For the selected composition type, a number of prototype structures are available, which will be classified by their crystal system (e.g., cubic, hexagonal, ...). Select a crystal system.
The crystal systems are determined by "spacegroup numbers", which represents symmetry of a structure. There are 230 spacegroups (1-230).
Following is the classification of the 7 crystal systems by spacegroups (sg):
1. Triclinic: sg 1-2
2. Monoclinic: sg 3-15
3. Orthorhombic: sg 16-74
4. Tetragonal: 75-142
5. Trigonal: sg 143-167
6. Hexagonal: 168-194
7. Cubic: sg 195-230.
The spacegroup of a prototype structure is present in the structure id: sg33, sg225 etc.
For example, spacegroup of prototype structure "POSCAR_sg33_icsd_065132" is 33.
```
crystal_systems = {'triclinic': set([1,2]),
'monoclinic': set(range(3,16)),
'orthorhombic': set(range(16,75)),
'tetragonal': set(range(75,143)),
'trigonal': set(range(143,168)),
'hexagonal': set(range(168,195)),
'cubic': set(range(195,231)),
}
sg_num_to_crystal_sys = {n: crystal_sys for crystal_sys, nums in crystal_systems.items() for n in nums}
# how do I get the prototype structure?
# Check how many prototype structures there are for each
prototype_folder = '/projects/rlmolecule/shubham/icsd_prototypes_poscars/DB/'
comp_prototypes = defaultdict(set)
for comp_type in comp_types:
for poscar_file in glob.glob(f"{prototype_folder}/{comp_type}/*"):
file_name = os.path.basename(poscar_file)
comp_prototypes[comp_type].add(file_name)
print(list(comp_prototypes.items())[:2])
# head and tail of number of prototypes available to choose from per composition type
print(sorted([len(vals) for vals in comp_prototypes.values()], reverse=True)[:20])
print(sorted([len(vals) for vals in comp_prototypes.values()])[:20])
# total number of prototypes for these compositions
print(sum([len(vals) for vals in comp_prototypes.values()]))
# total number of unique prototypes
prototype_files = set(p for vals in comp_prototypes.values() for p in vals)
print(f'{len(prototype_files)} prototype files')
# map the poscar files to their crystal system
prototype_to_crystal_sys = {}
for poscar_file in prototype_files:
sg_num = poscar_file.split('_')[1].replace('sg','')
system = sg_num_to_crystal_sys[int(sg_num)]
prototype_to_crystal_sys[poscar_file] = system
crystal_sys_prototypes = defaultdict(set)
for p, c in prototype_to_crystal_sys.items():
crystal_sys_prototypes[c].add(p)
for c, p in crystal_sys_prototypes.items():
print(c, len(p))
print(list(prototype_to_crystal_sys.items())[:3])
# how many crystal systems are available for each comp_type?
comp_type_to_crystal_sys = defaultdict(set)
for comp_type, prototypes in comp_prototypes.items():
for p in prototypes:
comp_type_to_crystal_sys[comp_type].add(prototype_to_crystal_sys[p])
print(list(comp_type_to_crystal_sys.items())[:3])
print(f"histogram of number of crystal structures per comp_type (out of {len(comp_type_to_crystal_sys)} total):")
for i in range(1,8):
num_matching = len([x for x in comp_type_to_crystal_sys.values() if len(x) == i])
print(f"{num_matching} comp_types have {i} crystal_systems")
# # add the crystal system to the graph
# for comp in compositions:
# comp_type = comp_to_comp_type[comp]
# crystal_systems = comp_type_to_crystal_sys[comp_type]
# for crystal_sys in crystal_systems:
# G2.add_edge(comp, comp + '|' + crystal_sys)
# print(f'{G2.number_of_nodes()} nodes, {G2.number_of_edges()} edges')
curr_comp_type = comp_type
# choose a crystal system
print(curr_comp_type, comp_type_to_crystal_sys[curr_comp_type])
curr_crystal_sys = next(iter(comp_type_to_crystal_sys[curr_comp_type]))
print(curr_crystal_sys)
# now choose a prototype structure
# for comp_type, prototypes in comp_prototypes.items():
avail_prototypes = set()
for p in comp_prototypes[curr_comp_type]:
crystal_sys = prototype_to_crystal_sys[p]
if crystal_sys == curr_crystal_sys:
avail_prototypes.add(p)
print(f'{len(avail_prototypes)} prototypes are {curr_crystal_sys} out of {len(comp_prototypes[curr_comp_type])} possible for {curr_comp_type}:')
print(avail_prototypes)
curr_prototype = next(iter(avail_prototypes))
print(curr_prototype)
G2 = nx.DiGraph()
# For the selected composition type, a number of prototype structures are available,
# which will be classified by their crystal system (e.g., cubic, hexagonal, ...). Select a crystal system.
# 4. For a chosen crystal system, consider all the prototypes and construct hypothetical decorated structures.
for comp_type in comp_types:
# crystal_systems = comp_type_to_crystal_sys[comp_type]
# for crystal_sys in crystal_systems:
for proto in comp_prototypes[comp_type]:
crystal_sys = prototype_to_crystal_sys[proto]
n1 = comp_type
n2 = n1 + '|' + crystal_sys
n3 = n2 + '|' + proto
G2.add_edge(n1, n2)
G2.add_edge(n2, n3)
# also add the # decorations.
# Once we have the composition, we will generate the real decorations.
# For now, just use an integer placeholder for the decoration number.
comp_type_stoic = tuple([int(x) for x in comp_type.split('_')[1:]])
comp_type_permutations = list(itertools.permutations(comp_type_stoic))
num_decor = len([permu for permu in comp_type_permutations if permu == comp_type_stoic])
for i in range(1,num_decor+1):
n4 = n3 + '|' + str(i)
G2.add_edge(n3, n4)
print(f'{G2.number_of_nodes()} nodes, {G2.number_of_edges()} edges')
# write this network as the action tree
out_file = f"{working_dir}/comp_type_to_decorations.edgelist.gz"
print(f"writing graph to {out_file}")
nx.write_edgelist(G2, out_file, delimiter='\t', data=False)
# Also load all the prototypes and write those to a file
structures = {}
for comp_type, poscar_files in tqdm(comp_prototypes.items()):
for file_name in poscar_files:
prototype_file = f"{prototype_folder}/{comp_type}/{file_name}"
strc = Structure.from_file(prototype_file, primitive=False)
crystal_sys = prototype_to_crystal_sys[file_name]
# for the key, use the same str as the node would use in the action graph
action_node_key = '|'.join([comp_type, crystal_sys, file_name])
structures[action_node_key] = strc.as_dict()
# now store them in a single file
import json
import gzip
# https://pymatgen.org/usage.html#side-note-as-dict-from-dict
out_file = f"{working_dir}/icsd_prototypes.json.gz"
with gzip.open(out_file, 'w') as out:
out.write(json.dumps(structures, indent=2).encode())
# and write the prototype file names to a file
out_file = f"{working_dir}/icsd_prototype_filenames.txt"
with open(out_file, 'w') as out:
out.write('\n'.join(prototype_files)+'\n')
# read them back
with gzip.open(out_file, 'r') as f:
structures_dict = json.loads(f.read().decode())
new_structures = {}
for key, structure_dict in structures_dict.items():
new_structures[key] = Structure.from_dict(structure_dict)
```
### Below is how Shubam generated the decorations
```
# 4. For a chosen crystal system, consider all the prototypes and construct hypothetical decorated structures.
# structures = {}
for comp_type in tqdm(comp_types):
crystal_systems = comp_type_to_crystal_sys[comp_type]
# create permutations of order of elements within a composition
comp_type_permutations = list(itertools.permutations(int(x) for x in comp_type.split('_')[1:]))
# second loop over all prototypes for a given composition type
for proto in comp_prototypes[comp_type]:
crystal_sys = prototype_to_crystal_sys[proto]
# G2.add_edge(comp, comp + '|' + crystal_sys)
# G2.add_edge(comp + '|' + crystal_sys, comp + '|' + proto)
if proto in structures:
strc = structures[proto]
else:
fname = proto.replace('POSCAR_','')
strc_file = f"{prototype_folder}/{comp_to_comp_type[comp]}/{proto}"
strc = Structure.from_file(strc_file, primitive=False)
structures[proto] = strc
comp_proto = Composition(strc.formula).reduced_composition
proto_stoic = (int(p) for p in comp_proto.formula if p.isdigit())
c = 0
# third loop over all permutations of order of elements for a composition, created in the first loop
# that also match the stoichiometry of the given prototype structure
for permu in comp_type_permutations:
if permu
# strc_subs = deepcopy(strc)
separator = ' '
comp_permu = separator.join(list(permu))
comp_stoic = [int(p) for p in comp_permu if p.isdigit()]
# check if stoichiometric coefficients of the composition to decorate match with those of prototype structure
if comp_stoic == proto_stoic:
c += 1
# TODO rather than create a new structures file for every possible decoration,
# store them all in a single json file
# # define lists of elements for elemental substitution
# original_ele = ''.join(i for i in comp_proto.formula if not i.isdigit()).split(' ')
# replacement_ele = ''.join(i for i in comp_permu if not i.isdigit()).split(' ')
# # dictionary containing original elements as keys and new elements as values
# replacement = {original_ele[i]: replacement_ele[i] for i in range(len(original_ele))}
# # 'replace_species' function from pymatgen to replace original elements with new elements
# strc_subs.replace_species(replacement)
# new_strc_file = 'POSCAR' + '_' + comp + '_' + fname + '_' + str(c)
# # strc_subs.to(filename= dir + '/' + )
G2.add_edge(comp + '|' + proto, comp + '|' + proto + '|' + str(c))
# print(new_strc_file)
print(f'{G2.number_of_nodes()} nodes, {G2.number_of_edges()} edges')
# write this network as the action tree
out_file = "inputs/all_actions.edgelist"
print(f"writing graph to {out_file}")
nx.write_edgelist(G2, out_file, delimiter='\t', data=False)
```
| github_jupyter |
### Vignettes
http://atlas.gs.washington.edu/fly-atac/docs/
### Import packages
```
library(data.table)
library(dplyr)
library(Matrix)
library(BuenColors)
library(stringr)
library(cowplot)
library(RColorBrewer)
library(ggpubr)
library(irlba)
library(umap)
library(gplots)
```
### Preprocess
##### make windows
`bedtools makewindows -g ../input/hg19/hg19.chrom.sizes -w 5000 > hg19.windows.5kb.bed`
##### Count reads
`bsub < count_reads_windows.sh`
```
path = './count_reads_windows_output/'
files <- list.files(path,pattern = "\\.txt$")
length(files)
#assuming tab separated values with a header
datalist = lapply(files, function(x)fread(paste0(path,x))$V4)
#assuming the same header/columns for all files
datafr = do.call("cbind", datalist)
dim(datafr)
df_regions = read.csv("./hg19.windows.5kb.bed",
sep = '\t',header=FALSE,stringsAsFactors=FALSE)
dim(df_regions)
head(df_regions)
winnames = paste(df_regions$V1,df_regions$V2,df_regions$V3,sep = "_")
head(winnames)
head(sapply(strsplit(files,'\\.'),'[', 1))
colnames(datafr) = sapply(strsplit(files,'\\.'),'[', 1)
rownames(datafr) = winnames
dim(datafr)
head(datafr)
# saveRDS(datafr, file = './datafr.rds')
# datafr = readRDS('./datafr.rds')
elbow_plot <- function(mat,num_pcs=50,scale=FALSE,center=FALSE,title='',width=3,height=3){
set.seed(2019)
mat = data.matrix(mat)
SVD = irlba(mat, num_pcs, num_pcs,scale=scale,center=center)
options(repr.plot.width=width, repr.plot.height=height)
df_plot = data.frame(PC=1:num_pcs, SD=SVD$d);
# print(SVD$d[1:num_pcs])
p <- ggplot(df_plot, aes(x = PC, y = SD)) +
geom_point(col="#cd5c5c",size = 1) +
ggtitle(title)
return(p)
}
run_umap <- function(fm_mat){
umap_object = umap(t(fm_mat),random_state = 2019)
df_umap = umap_object$layout
return(df_umap)
}
plot_umap <- function(df_umap,labels,title='UMAP',colormap=colormap){
set.seed(2019)
df_umap = data.frame(cbind(df_umap,labels),stringsAsFactors = FALSE)
colnames(df_umap) = c('umap1','umap2','celltype')
df_umap$umap1 = as.numeric(df_umap$umap1)
df_umap$umap2 = as.numeric(df_umap$umap2)
options(repr.plot.width=4, repr.plot.height=4)
p <- ggplot(shuf(df_umap), aes(x = umap1, y = umap2, color = celltype)) +
geom_point(size = 1) + scale_color_manual(values = colormap,breaks=sort(unique(labels))) +
ggtitle(title)
return(p)
}
```
### Identify Clades
```
metadata <- read.table('../input/metadata.tsv',
header = TRUE,
stringsAsFactors=FALSE,quote="",row.names=1)
binary_mat = as.matrix((datafr > 0) + 0)
binary_mat = Matrix(binary_mat, sparse = TRUE)
num_cells_ncounted = rowSums(binary_mat)
ncounts = binary_mat[which(num_cells_ncounted >= num_cells_ncounted[order(num_cells_ncounted,decreasing=T)[20000]]),]
new_counts = colSums(ncounts)
# ncounts = ncounts[,new_counts >= quantile(new_counts,probs=0.1)]
ncounts = ncounts[rowSums(ncounts) > 0,]
options(repr.plot.width=8, repr.plot.height=4)
par(mfrow=c(1,2))
hist(log10(num_cells_ncounted),main="No. of Cells Each Site is Observed In",breaks=50)
abline(v=log10(num_cells_ncounted[order(num_cells_ncounted,decreasing=T)[20000]]),lwd=2,col="indianred")
hist(log10(new_counts),main="Number of Sites Each Cell Uses",breaks=50)
# abline(v=log10(quantile(new_counts,probs=0.1)),lwd=2,col="indianred")
dim(ncounts)
nfreqs = t(t(ncounts) / Matrix::colSums(ncounts))
idf = as(log(1 + ncol(ncounts) / Matrix::rowSums(ncounts)), "sparseVector")
tf_idf_counts = as(Diagonal(x=as.vector(idf)), "sparseMatrix") %*% nfreqs
dim(tf_idf_counts)
p_elbow <- elbow_plot(tf_idf_counts,num_pcs = 200, title = 'PCA')
p_elbow
set.seed(2019)
num_pcs = 150
SVD = irlba(tf_idf_counts, num_pcs, num_pcs)
sk_diag = matrix(0, nrow=num_pcs, ncol=num_pcs)
diag(sk_diag) = SVD$d
##remove component 1 as suggested
sk_diag[1,1] = 0
SVDumap_vd = t(sk_diag %*% t(SVD$v))
dim(SVDumap_vd)
LSI_out = t(t(sk_diag %*% t(SVD$v)) %*% t(SVD$u))
LSI_out = t(scale(t(LSI_out)))
LSI_out[LSI_out > 1.5] = 1.5
LSI_out[LSI_out < -1.5] = -1.5
dim(LSI_out)
#This step can take a minute too
hclust_cells = hclust(proxy::dist(t(sk_diag %*% t(SVD$v)), method="cosine"), method="ward.D2")
hclust_genes = hclust(proxy::dist(t(sk_diag %*% t(SVD$u)), method="cosine"), method="ward.D2")
color_pal = colorRampPalette(brewer.pal(8, "Set2"))(length(unique(metadata$label)))
hmcols = colorpanel(100, "steelblue", "white", "tomato")
cells_tree_cut = cutree(hclust_cells, length(unique(metadata$label)))
lsi_cells = cbind(colnames(ncounts),cells_tree_cut)
```
#### heatmap
```
options(repr.plot.width=4, repr.plot.height=6)
heatmap.2(LSI_out,
col=hmcols,
ColSideColors=color_pal[as.factor(cells_tree_cut)],
#RowSideColors=color_pal[as.factor(genes_tree_cut)],
Rowv = as.dendrogram(hclust_genes), Colv = as.dendrogram(hclust_cells),
labRow=FALSE, labCol=FALSE, trace="none", scale="none",
useRaster=TRUE)
str(hclust_cells)
dim(SVDumap_vd)
df_umap_LSI <- run_umap(t(SVDumap_vd))
df_umap_LSI = as.data.frame(df_umap_LSI)
row.names(df_umap_LSI) = colnames(datafr)
labels = metadata[colnames(datafr),'label']
colormap = c(jdb_color_maps, "UNK" = "#333333" )
p_LSI <- plot_umap(df_umap_LSI,labels = labels,colormap = colormap,title='Cusanovich2018')
p_LSI
labels2 = as.character(cells_tree_cut)
num_colors = length(unique(metadata$label))
colormap2= colorRampPalette(brewer.pal(8, "Set2"))(num_colors)
names(colormap2) = as.character(seq(1,num_colors))
p_LSI_cluster <- plot_umap(df_umap_LSI,labels = labels2,colormap = colormap2,title='Cusanovich2018')
p_LSI_cluster = p_LSI_cluster+labs(color='cluster')
p_LSI_cluster
options(repr.plot.width=4*2.5, repr.plot.height=4*1)
p_group = cowplot::plot_grid(p_LSI,p_LSI_cluster,
labels = "AUTO",ncol = 2)
p_group
cowplot::ggsave(p_group,filename = 'Cusanovich2018_clades_buenrostro2018.pdf',width = 4*2.5, height = 4*1)
save.image(file = 'Cusanovich2018_buenrostro2018_idenfify_clades.RData')
df_pseudobulk = cbind(cellid=colnames(datafr),clusterid=labels2)
write.table(df_pseudobulk,'./peak_calling/pseudobulk.tsv',quote = FALSE,sep='\t',row.names = FALSE)
```
| github_jupyter |
<center>
<img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/Logos/organization_logo/organization_logo.png" width="300" alt="cognitiveclass.ai logo" />
</center>
# Functions in Python
Estimated time needed: **40** minutes
## Objectives
After completing this lab you will be able to:
- Understand functions and variables
- Work with functions and variables
<h1>Functions in Python</h1>
<p><strong>Welcome!</strong> This notebook will teach you about the functions in the Python Programming Language. By the end of this lab, you'll know the basic concepts about function, variables, and how to use functions.</p>
<h2>Table of Contents</h2>
<div class="alert alert-block alert-info" style="margin-top: 20px">
<ul>
<li>
<a href="#func">Functions</a>
<ul>
<li><a href="content">What is a function?</a></li>
<li><a href="var">Variables</a></li>
<li><a href="simple">Functions Make Things Simple</a></li>
</ul>
</li>
<li><a href="pre">Pre-defined functions</a></li>
<li><a href="if">Using <code>if</code>/<code>else</code> Statements and Loops in Functions</a></li>
<li><a href="default">Setting default argument values in your custom functions</a></li>
<li><a href="global">Global variables</a></li>
<li><a href="scope">Scope of a Variable</a></li>
<li><a href="collec">Collections and Functions</a></li>
<li>
<a href="#quiz">Quiz on Loops</a>
</li>
</ul>
</div>
<hr>
<h2 id="func">Functions</h2>
A function is a reusable block of code which performs operations specified in the function. They let you break down tasks and allow you to reuse your code in different programs.
There are two types of functions :
- <b>Pre-defined functions</b>
- <b>User defined functions</b>
<h3 id="content">What is a Function?</h3>
You can define functions to provide the required functionality. Here are simple rules to define a function in Python:
- Functions blocks begin <code>def</code> followed by the function <code>name</code> and parentheses <code>()</code>.
- There are input parameters or arguments that should be placed within these parentheses.
- You can also define parameters inside these parentheses.
- There is a body within every function that starts with a colon (<code>:</code>) and is indented.
- You can also place documentation before the body.
- The statement <code>return</code> exits a function, optionally passing back a value.
An example of a function that adds on to the parameter <code>a</code> prints and returns the output as <code>b</code>:
```
# First function example: Add 1 to a and store as b
def add(a):
b = a + 1
print(a, "if you add one", b)
return(b)
```
The figure below illustrates the terminology:
<img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%203/Images/FuncsDefinition.png" width="500" />
We can obtain help about a function :
```
# Get a help on add function
help(add)
```
We can call the function:
```
# Call the function add()
add(1)
```
If we call the function with a new input we get a new result:
```
# Call the function add()
add(2)
```
We can create different functions. For example, we can create a function that multiplies two numbers. The numbers will be represented by the variables <code>a</code> and <code>b</code>:
```
# Define a function for multiple two numbers
def Mult(a, b):
c = a * b
return(c)
print('This is not printed')
result = Mult(12,2)
print(result)
```
The same function can be used for different data types. For example, we can multiply two integers:
```
# Use mult() multiply two integers
Mult(2, 3)
```
Note how the function terminates at the <code> return </code> statement, while passing back a value. This value can be further assigned to a different variable as desired.
<hr>
The same function can be used for different data types. For example, we can multiply two integers:
Two Floats:
```
# Use mult() multiply two floats
Mult(10.0, 3.14)
```
We can even replicate a string by multiplying with an integer:
```
# Use mult() multiply two different type values together
Mult(2, "Michael Jackson ")
```
<h3 id="var">Variables</h3>
The input to a function is called a formal parameter.
A variable that is declared inside a function is called a local variable. The parameter only exists within the function (i.e. the point where the function starts and stops).
A variable that is declared outside a function definition is a global variable, and its value is accessible and modifiable throughout the program. We will discuss more about global variables at the end of the lab.
```
# Function Definition
def square(a):
# Local variable b
b = 1
c = a * a + b
print(a, "if you square + 1", c)
return(c)
```
The labels are displayed in the figure:
<img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%203/Images/FuncsVar.png" width="500" />
We can call the function with an input of <b>3</b>:
```
# Initializes Global variable
x = 3
# Makes function call and return function a y
y = square(x)
y
```
We can call the function with an input of <b>2</b> in a different manner:
```
# Directly enter a number as parameter
square(2)
```
If there is no <code>return</code> statement, the function returns <code>None</code>. The following two functions are equivalent:
```
# Define functions, one with return value None and other without return value
def MJ():
print('Michael Jackson')
def MJ1():
print('Michael Jackson')
return(None)
# See the output
MJ()
# See the output
MJ1()
```
Printing the function after a call reveals a **None** is the default return statement:
```
# See what functions returns are
print(MJ())
print(MJ1())
```
Create a function <code>con</code> that concatenates two strings using the addition operation:
```
# Define the function for combining strings
def con(a, b):
return(a + b)
# Test on the con() function
con("This ", "is")
```
<hr/>
<div class="alert alert-success alertsuccess" style="margin-top: 20px">
<h4> [Tip] How do I learn more about the pre-defined functions in Python? </h4>
<p>We will be introducing a variety of pre-defined functions to you as you learn more about Python. There are just too many functions, so there's no way we can teach them all in one sitting. But if you'd like to take a quick peek, here's a short reference card for some of the commonly-used pre-defined functions: <a href="http://www.astro.up.pt/~sousasag/Python_For_Astronomers/Python_qr.pdf">Reference</a></p>
</div>
<hr/>
<h3 id="simple">Functions Make Things Simple</h3>
Consider the two lines of code in <b>Block 1</b> and <b>Block 2</b>: the procedure for each block is identical. The only thing that is different is the variable names and values.
<h4>Block 1:</h4>
```
# a and b calculation block1
a1 = 4
b1 = 5
c1 = a1 + b1 + 2 * a1 * b1 - 1
if(c1 < 0):
c1 = 0
else:
c1 = 5
c1
```
<h4>Block 2:</h4>
```
# a and b calculation block2
a2 = 0
b2 = 0
c2 = a2 + b2 + 2 * a2 * b2 - 1
if(c2 < 0):
c2 = 0
else:
c2 = 5
c2
```
We can replace the lines of code with a function. A function combines many instructions into a single line of code. Once a function is defined, it can be used repeatedly. You can invoke the same function many times in your program. You can save your function and use it in another program or use someone else’s function. The lines of code in code <b>Block 1</b> and code <b>Block 2</b> can be replaced by the following function:
```
# Make a Function for the calculation above
def Equation(a,b):
c = a + b + 2 * a * b - 1
if(c < 0):
c = 0
else:
c = 5
return(c)
```
This function takes two inputs, a and b, then applies several operations to return c.
We simply define the function, replace the instructions with the function, and input the new values of <code>a1</code>, <code>b1</code> and <code>a2</code>, <code>b2</code> as inputs. The entire process is demonstrated in the figure:
<img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%203/Images/FuncsPros.gif" width="850" />
Code **Blocks 1** and **Block 2** can now be replaced with code **Block 3** and code **Block 4**.
<h4>Block 3:</h4>
```
a1 = 4
b1 = 5
c1 = Equation(a1, b1)
c1
```
<h4>Block 4:</h4>
```
a2 = 0
b2 = 0
c2 = Equation(a2, b2)
c2
```
<hr>
<h2 id="pre">Pre-defined functions</h2>
There are many pre-defined functions in Python, so let's start with the simple ones.
The <code>print()</code> function:
```
# Build-in function print()
album_ratings = [10.0, 8.5, 9.5, 7.0, 7.0, 9.5, 9.0, 9.5]
print(album_ratings)
```
The <code>sum()</code> function adds all the elements in a list or tuple:
```
# Use sum() to add every element in a list or tuple together
sum(album_ratings)
```
The <code>len()</code> function returns the length of a list or tuple:
```
# Show the length of the list or tuple
len(album_ratings)
```
<h2 id="if">Using <code>if</code>/<code>else</code> Statements and Loops in Functions</h2>
The <code>return()</code> function is particularly useful if you have any IF statements in the function, when you want your output to be dependent on some condition:
```
# Function example
def type_of_album(artist, album, year_released):
print(artist, album, year_released)
if year_released > 1980:
return "Modern"
else:
return "Oldie"
x = type_of_album("Michael Jackson", "Thriller", 1980)
print(x)
```
We can use a loop in a function. For example, we can <code>print</code> out each element in a list:
```
# Print the list using for loop
def PrintList(the_list):
for element in the_list:
print(element)
# Implement the printlist function
PrintList(['1', 1, 'the man', "abc"])
```
<hr>
<h2 id="default">Setting default argument values in your custom functions</h2>
You can set a default value for arguments in your function. For example, in the <code>isGoodRating()</code> function, what if we wanted to create a threshold for what we consider to be a good rating? Perhaps by default, we should have a default rating of 4:
```
# Example for setting param with default value
def isGoodRating(rating=4):
if(rating < 7):
print("this album sucks it's rating is",rating)
else:
print("this album is good its rating is",rating)
# Test the value with default value and with input
isGoodRating()
isGoodRating(10)
```
<hr>
<h2 id="global">Global variables</h2>
So far, we've been creating variables within functions, but we have not discussed variables outside the function. These are called global variables.
<br>
Let's try to see what <code>printer1</code> returns:
```
# Example of global variable
artist = "Michael Jackson"
def printer1(artist):
internal_var1 = artist
print(artist, "is an artist")
printer1(artist)
# try runningthe following code
#printer1(internal_var1)
```
<b>We got a Name Error: <code>name 'internal_var' is not defined</code>. Why?</b>
It's because all the variables we create in the function is a <b>local variable</b>, meaning that the variable assignment does not persist outside the function.
But there is a way to create <b>global variables</b> from within a function as follows:
```
artist = "Michael Jackson"
def printer(artist):
global internal_var
internal_var= "Whitney Houston"
print(artist,"is an artist")
printer(artist)
printer(internal_var)
```
<h2 id="scope">Scope of a Variable</h2>
The scope of a variable is the part of that program where that variable is accessible. Variables that are declared outside of all function definitions, such as the <code>myFavouriteBand</code> variable in the code shown here, are accessible from anywhere within the program. As a result, such variables are said to have global scope, and are known as global variables.
<code>myFavouriteBand</code> is a global variable, so it is accessible from within the <code>getBandRating</code> function, and we can use it to determine a band's rating. We can also use it outside of the function, such as when we pass it to the print function to display it:
```
# Example of global variable
myFavouriteBand = "AC/DC"
def getBandRating(bandname):
if bandname == myFavouriteBand:
return 10.0
else:
return 0.0
print("AC/DC's rating is:", getBandRating("AC/DC"))
print("Deep Purple's rating is:",getBandRating("Deep Purple"))
print("My favourite band is:", myFavouriteBand)
```
Take a look at this modified version of our code. Now the <code>myFavouriteBand</code> variable is defined within the <code>getBandRating</code> function. A variable that is defined within a function is said to be a local variable of that function. That means that it is only accessible from within the function in which it is defined. Our <code>getBandRating</code> function will still work, because <code>myFavouriteBand</code> is still defined within the function. However, we can no longer print <code>myFavouriteBand</code> outside our function, because it is a local variable of our <code>getBandRating</code> function; it is only defined within the <code>getBandRating</code> function:
```
# Example of local variable
def getBandRating(bandname):
myFavouriteBand = "AC/DC"
if bandname == myFavouriteBand:
return 10.0
else:
return 0.0
print("AC/DC's rating is: ", getBandRating("AC/DC"))
print("Deep Purple's rating is: ", getBandRating("Deep Purple"))
print("My favourite band is", myFavouriteBand)
```
Finally, take a look at this example. We now have two <code>myFavouriteBand</code> variable definitions. The first one of these has a global scope, and the second of them is a local variable within the <code>getBandRating</code> function. Within the <code>getBandRating</code> function, the local variable takes precedence. **Deep Purple** will receive a rating of 10.0 when passed to the <code>getBandRating</code> function. However, outside of the <code>getBandRating</code> function, the <code>getBandRating</code> s local variable is not defined, so the <code>myFavouriteBand</code> variable we print is the global variable, which has a value of **AC/DC**:
```
# Example of global variable and local variable with the same name
myFavouriteBand = "AC/DC"
def getBandRating(bandname):
myFavouriteBand = "Deep Purple"
if bandname == myFavouriteBand:
return 10.0
else:
return 0.0
print("AC/DC's rating is:",getBandRating("AC/DC"))
print("Deep Purple's rating is: ",getBandRating("Deep Purple"))
print("My favourite band is:",myFavouriteBand)
```
<hr>
<h2 id ="collec"> Collections and Functions</h2>
When the number of arguments are unknown for a function, They can all be packed into a tuple as shown:
```
def printAll(*args): # All the arguments are 'packed' into args which can be treated like a tuple
print("No of arguments:", len(args))
for argument in args:
print(argument)
#printAll with 3 arguments
printAll('Horsefeather','Adonis','Bone')
#printAll with 4 arguments
printAll('Sidecar','Long Island','Mudslide','Carriage')
```
Similarly, The arguments can also be packed into a dictionary as shown:
```
def printDictionary(**args):
for key in args:
print(key + " : " + args[key])
printDictionary(Country='Canada',Province='Ontario',City='Toronto')
```
Functions can be incredibly powerful and versatile. They can accept (and return) data types, objects and even other functions as arguements. Consider the example below:
```
def addItems(list):
list.append("Three")
list.append("Four")
myList = ["One","Two"]
addItems(myList)
myList
```
Note how the changes made to the list are not limited to the functions scope. This occurs as it is the lists **reference** that is passed to the function - Any changes made are on the orignal instance of the list. Therefore, one should be cautious when passing mutable objects into functions.
<hr>
<h2>Quiz on Functions</h2>
Come up with a function that divides the first input by the second input:
```
# Write your code below and press Shift+Enter to execute
```
<details><summary>Click here for the solution</summary>
```python
def div(a, b):
return(a/b)
```
</details>
<hr>
Use the function <code>con</code> for the following question.
```
# Use the con function for the following question
def con(a, b):
return(a + b)
```
Can the <code>con</code> function we defined before be used to add two integers or strings?
```
# Write your code below and press Shift+Enter to execute
```
<details><summary>Click here for the solution</summary>
```python
Yes, for example:
con(2, 2)
```
</details>
<hr>
Can the <code>con</code> function we defined before be used to concatenate lists or tuples?
```
# Write your code below and press Shift+Enter to execute
```
<details><summary>Click here for the solution</summary>
```python
Yes, for example:
con(['a', 1], ['b', 1])
```
</details>
**Probability Bag**
You have been tasked with creating a lab that demonstrates the basics of probability by simulating a bag filled with colored balls. The bag is represented using a dictionary called "bag", where the key represents the color of the ball and the value represents the no of balls. The skeleton code has been made for you, do not add or remove any functions. Complete the following functions -
- fillBag - A function that packs it's arguments into a global dictionary "bag".
- totalBalls - returns the total no of balls in the bucket
- probOf - takes a color (string) as argument and returns probability of drawing the selected ball. Assume total balls are not zero and the color given is a valid key.
- probAll - returns a dictionary of all colors and their corresponding probability
```
def fillBag(**balls):
pass
def totalBalls():
pass
def probOf(color):
pass
def probAll():
pass
```
Run this snippet of code to test your solution.
Note: This is not a comprehensive test.
```
testBag = dict(red = 12, blue = 20, green = 14, grey = 10)
total = sum(testBag.values())
prob={}
for color in testBag:
prob[color] = testBag[color]/total;
def testMsg(passed):
if passed:
return 'Test Passed'
else :
return ' Test Failed'
print("fillBag : ")
try:
fillBag(**testBag)
print(testMsg(bag == testBag))
except NameError as e:
print('Error! Code: {c}, Message: {m}'.format(c = type(e).__name__, m = str(e)))
except:
print("An error occured. Recheck your function")
print("totalBalls : ")
try:
print(testMsg(total == totalBalls()))
except NameError as e:
print('Error! Code: {c}, Message: {m}'.format(c = type(e).__name__, m = str(e)))
except:
print("An error occured. Recheck your function")
print("probOf")
try:
passed = True
for color in testBag:
if probOf(color) != prob[color]:
passed = False
print(testMsg(passed) )
except NameError as e:
print('Error! Code: {c}, Message: {m}'.format(c = type(e).__name__, m = str(e)))
except:
print("An error occured. Recheck your function")
print("probAll")
try:
print(testMsg(probAll() == prob))
except NameError as e:
print('Error! Code: {c}, Message: {m}'.format(c = type(e).__name__, m = str(e)))
except:
print("An error occured. Recheck your function")
```
<details><summary>Click here for the solution</summary>
```python
def fillBag(**balls):
global bag
bag = balls
def totalBalls():
total = 0
for color in bag:
total += bag[color]
return total
# alternatively,
# return sum(bag.values())
def probOf(color):
return bag[color]/totalBalls()
def probAll():
probDict = {}
for color in bag:
probDict[color] = probOf(color)
return probDict
```
</details>
<hr>
<h2>The last exercise!</h2>
<p>Congratulations, you have completed your first lesson and hands-on lab in Python. However, there is one more thing you need to do. The Data Science community encourages sharing work. The best way to share and showcase your work is to share it on GitHub. By sharing your notebook on GitHub you are not only building your reputation with fellow data scientists, but you can also show it off when applying for a job. Even though this was your first piece of work, it is never too early to start building good habits. So, please read and follow <a href="https://cognitiveclass.ai/blog/data-scientists-stand-out-by-sharing-your-notebooks/" target="_blank">this article</a> to learn how to share your work.
<hr>
## Author
<a href="https://www.linkedin.com/in/joseph-s-50398b136/" target="_blank">Joseph Santarcangelo</a>
## Other contributors
<a href="www.linkedin.com/in/jiahui-mavis-zhou-a4537814a">Mavis Zhou</a>
## Change Log
| Date (YYYY-MM-DD) | Version | Changed By | Change Description |
| ----------------- | ------- | ---------- | ---------------------------------------------------------------------------------------------------------------------- |
| 2020-08-26 | 0.2 | Lavanya | Moved lab to course repo in GitLab |
| 2020 -09 -04 | 0.2 | Arjun | Under What is a function, added code/text to further demonstrate the functionality of the return statement |
| 2020 -09 -04 | 0.2 | Arjun | Under Global Variables, modify the code block to try and print ‘internal_var’ - So a nameError message can be observed |
| 2020 -09 -04 | 0.2 | Arjun | Added section Collections and Functions |
| 2020 -09 -04 | 0.2 | Arjun | Added exercise “Probability Bag” |
<hr/>
## <h3 align="center"> © IBM Corporation 2020. All rights reserved. <h3/>
| github_jupyter |
# Desafio - Módulo 4
O objetivo desse exercício é classificar imagens pelo uso do Deep Learning (Keras e Tensorflow).
```
#Libs nencessárias
import tensorflow as tf
import IPython.display as display
from PIL import Image
import numpy as np
import matplotlib.pyplot as plt
import os
AUTOTUNE = tf.data.experimental.AUTOTUNE
#Versão do tf:
tf.__version__
#Carregando o conjunto de dados
#import pathlib
#data_dir = tf.keras.utils.get_file(origin='http://download.tensorflow.org/example_images/flower_photos.tgz', fname='flower_photos', untar=True)
#Carregando o conjunto de dados
import pathlib
data_dir = tf.keras.utils.get_file(origin='https://storage.googleapis.com/download.tensorflow.org/example_images/flower', fname='flower_photos', untar=True)
data_dir = pathlib.Path(data_dir)
#Total de imagens
image_count = len(list(data_dir.glob('*/*.jpg')))
image_count
#Classes de imagens:
CLASS_NAMES = np.array([item.name for item in data_dir.glob('*') if item.name != 'LICENSE.txt'])
CLASS_NAMES
#Listando roses
roses = list(data_dir.glob('roses/*'))
for image_path in roses[:3]:
display.display(Image.open(str(image_path)))
```
# Desafio
Exploração de dados
```
module_selection = ("mobilenet_v2_100_224", 224)
handle_base, pixels = module_selection
MODULE_HANDLE ="https://tfhub.dev/google/imagenet/{}/feature_vector/4".format(handle_base)
IMAGE_SIZE = (pixels, pixels)
print("Using {} with input size {}".format(MODULE_HANDLE, IMAGE_SIZE))
BATCH_SIZE = 32
datagen_kwargs = dict(rescale=1./255, validation_split=.20)
dataflow_kwargs = dict(target_size=IMAGE_SIZE, batch_size=BATCH_SIZE,
interpolation="bilinear")
valid_datagen = tf.keras.preprocessing.image.ImageDataGenerator(
**datagen_kwargs)
valid_generator = valid_datagen.flow_from_directory(
data_dir, subset="validation", shuffle=False, **dataflow_kwargs)
do_data_augmentation = False
if do_data_augmentation:
train_datagen = tf.keras.preprocessing.image.ImageDataGenerator(
rotation_range=40,
horizontal_flip=True,
width_shift_range=0.2, height_shift_range=0.2,
shear_range=0.2, zoom_range=0.2,
**datagen_kwargs)
else:
train_datagen = valid_datagen
train_generator = train_datagen.flow_from_directory(
data_dir, subset="training", shuffle=True, **dataflow_kwargs)
```
Modelagem
```
do_fine_tuning = False
import tensorflow_hub as hub
print("Building model with", MODULE_HANDLE)
model = tf.keras.Sequential([
# Explicitly define the input shape so the model can be properly
# loaded by the TFLiteConverter
tf.keras.layers.InputLayer(input_shape=IMAGE_SIZE + (3,)),
hub.KerasLayer(MODULE_HANDLE, trainable=do_fine_tuning),
tf.keras.layers.Dropout(rate=0.2),
tf.keras.layers.Dense(train_generator.num_classes,
kernel_regularizer=tf.keras.regularizers.l2(0.0001))
])
model.build((None,)+IMAGE_SIZE+(3,))
model.summary()
```
Treinamento de modelo
```
#Compilando o modelo
model.compile(optimizer=tf.keras.optimizers.SGD(lr=0.005, momentum=0.9),
loss=tf.keras.losses.CategoricalCrossentropy(from_logits=True, label_smoothing=0.1),
metrics=['accuracy'])
#Definindo epochs de treinamento, quantidade de vezes que as images serão treinadas
steps_per_epoch = train_generator.samples // train_generator.batch_size
validation_steps = valid_generator.samples // valid_generator.batch_size
hist = model.fit(
train_generator,
epochs=5, steps_per_epoch=steps_per_epoch,
validation_data=valid_generator,
validation_steps=validation_steps).history
```
Avaliação do modelo (acurácia)
```
plt.figure()
plt.ylabel("Loss (training and validation)")
plt.xlabel("Training Steps")
plt.ylim([0,2])
plt.plot(hist["loss"], color='red', label='Training Loss')
plt.plot(hist["val_loss"], color='green', label='Validation Loss')
plt.legend(loc='upper right')
plt.figure()
plt.ylabel("Accuracy (training and validation)")
plt.xlabel("Training Steps")
plt.ylim([0,1])
plt.plot(hist["accuracy"], color = 'red', label='Acurracy')
plt.plot(hist["val_accuracy"], color = 'blue', label='Validation Acurracy')
plt.legend(loc='lower right')
#Testando o modelo em 1 imagem do dataset
def get_class_string_from_index(index):
for class_string, class_index in valid_generator.class_indices.items():
if class_index == index:
return class_string
x, y = next(valid_generator)
image = x[0, :, :, :]
true_index = np.argmax(y[0])
plt.imshow(image)
plt.axis('off')
plt.show()
prediction_scores = model.predict(np.expand_dims(image, axis=0))
predicted_index = np.argmax(prediction_scores)
print("True label: " + get_class_string_from_index(true_index))
print("Predicted label: " + get_class_string_from_index(predicted_index))
#Salvando modelo
saved_model_path = "/tmp/saved_flowers_model"
tf.saved_model.save(model, saved_model_path)
```
Predições
```
#Prevendo novos dados de fora do treinamento e teste
from tensorflow import keras
batch_size = 32
img_height = 180
img_width = 180
sunflower_url = "https://storage.googleapis.com/download.tensorflow.org/example_images/592px-Red_sunflower.jpg"
sunflower_path = tf.keras.utils.get_file('Red_sunflower', origin=sunflower_url)
img = keras.preprocessing.image.load_img(
sunflower_path, target_size=(img_height, img_width)
)
img_array = keras.preprocessing.image.img_to_array(img)
img_array = tf.expand_dims(img_array, 0) # Create a batch
predictions = model.predict(img_array)
score = tf.nn.softmax(predictions[0])
print(
"This image most likely belongs to {} with a {:.2f} percent confidence."
.format(class_names[np.argmax(score)], 100 * np.max(score)))
train_images = train_images / 255.0
test_images = test_images / 255.0
predictions = model.predict(test_images)
predictions[0]
rever com https://www.tensorflow.org/hub/tutorials/image_feature_vector
```
Saiba mais em:
https://www.tensorflow.org/tutorials/images/classification?hl=pt
https://www.tensorflow.org/hub/tutorials/tf2_image_retraining?hl=pt
Visite mau [Github](https://github.com/k3ybladewielder) :3
| github_jupyter |
# Making a Twitter Bot
This Jupyter Notebooks shows the basics of making a Twitter bot with Python.
This is a *very* heavily modified version of a [tutorial posted on the CUNY Academic Commons Blog](https://emerging.commons.gc.cuny.edu/2013/10/making-twitter-bot-python-tutorial/) by [Robin Camilla Davis](https://twitter.com/robincamille). I have taken some of the content and structure of the static blog post and ported it into a Jupyter Notebook Python environment.
Here is Robin's introduction:
> If there’s one thing this budding computational linguist finds delightful, it’s computers that talk to us. From [SmarterChild](http://en.wikipedia.org/wiki/SmarterChild) to [horse_ebooks](http://www.robincamille.com/2013-09-25-horse_ebooks/) to [Beetlejuice](http://muffinlabs.com/content/twitter-bot-info.html), I love the weirdness of machines that seem to have a voice, especially when it’s a Twitter bot that adds its murmur to a tweetstream of accounts mostly run by other humans.
> As fun midnight project a few weeks ago, I cobbled together [@MechanicalPoe](https://twitter.com/MechanicalPoe), a Twitter bot that tweets Poe works line by line on the hour from a long .txt file. This slow-tweeting of text is by no means new—[@SlowDante](https://twitter.com/slowdante) is pretty popular, and so is[@CDarwin](http://www.metaburbia.com/darwin/), among many others. In case you want to make your own, here are the quick ‘n’ easy steps I took. This is just one way of doing it—shop around and see what others have done, too.
## Contents
* [Steps Towards an Ecology of Bots](#Steps-Towards-an-Ecology-of-Bots)
* [Building a Basic Bot](#Building-a-Basic-Bot)
## Steps Towards an Ecology of Bots
Creating a Twitter bot means creating a Twitter account AND a Twitter app. The account will belong to the bot and the app will be the mechanism you use to access the Twitter API and control the bot **WITH PYTHON.**
Creating a Twitter bot is somewhat complicated because of quirks in the way Twitter manages verification. If you don’t already have a Twitter account, the process of creating a Twitter bot is pretty painless. If you already have a Twitter account, you have to do the *Twitter verification dance*. Managing more than one Twitter account is a huge *pain in the ass* because you need a unique email address and mobile phone number for each account.
**Step 0)** *Note: This step is only for people who already have Twitter accounts.* Log into Twitter and visit [your account's mobile devices settings page](https://twitter.com/settings/devices). If you have a phone number associated with that account click “Delete my phone” to remove your mobile from number from this account. You are going to need that phone number for creating your bot account. Alternatively, you can get a free phone number from [Google Voice](http://voice.google.com) and use that for all the mobile phone verification BS.
You are also going to need another email address for your new Twitter account. Twitter won’t let you create multiple accounts with the same email address. If you have a 2nd email address you’ll need to use it in step 1 below. If you don’t have another email address it is easy to [create one with gmail](https://accounts.google.com/signup). Alternatively, you can fake a new unique email address with Gmail by adding a `+` and something after your username. For example: `myemail+sometext@gmail.com`. Any messages addressed to this email will still show up in your inbox, but it behaves like a different email address. Give it a try!
**Step 1)** Create a new Twitter account by visiting the [Twitter signup page](https://twitter.com/signup). This will be the Twitter account for your bot, so pick your username wisely. You will need to pick a name that hasn’t already been selected. Twitter *should* ask you for a mobile phone verification when you are creating the account, if it doesn’t you’ll have to add the phone number to your profile in [the new account's mobile devices settings page](https://twitter.com/settings/devices).
**Step 2)** OK. Now that you have a Twitter **account** for your bot, you need to create a Twitter **application** so you can access the Twitter API **WITH PYTHON**. Twitter doesn’t allow people to connect to their API willy-nilly. Creating applications is Twitter's way of tracking (and controlling) who is programmatically accessing their network and data. In the world of big social media sites, heavily regulated access to APIs, networks, and data is a fact of life.
Fortunately, creating an application on Twitter is very easy. Visit Twitter’s [Application Management page](https://apps.twitter.com/) and click the “Create New App” button. You’ll be shown the form shown below. Pick a name for your application and a description, this information is just for you it should be named something different (and more descriptive) than the account you just created. For the website, put the name of the course `http://annettevee.com/2015fall_computationalmedia/` into the website field.

**Step 3)** The last step for this part of the Twitter bot making process is to create the Access Token so your Python code can access the API. With your app created, you should see a dashboard page will all kinds of technical jargon about your Twitter app. You can ignore most of it for now and instead focus on the tab that says “Keys and Access Tokens.” The keys and access tokens behave like passwords so your computer can talk to Twitter’s computer. Twitter uses a protocol called [OAuth](http://oauth.net) for [“providing authorized access to its API](https://dev.twitter.com/oauth). You don’t really need to wor
- Click the "Keys and Access Tokens” tab
- Click the “Create my access token button” at the bottom of the page.
Once you have created your access tokens there will be **four** pieces of information you will need to put into your Python script so it can access the Twitter API:
1. Consumer Key
2. Consumer Secret
3. Access Token
4. Access Token Secret

Some other things to consider:
- When you create a Twitter App you agree to the [developer agreement and policy](https://dev.twitter.com/overview/terms/agreement-and-policy) which has some complete bullshit within it. Basically, Twitter reserves the right to lock down your application and your account if their machine learning algorithms detect what they think is bad behavior.
- Creating to many Accounts can lock-out your mobile phone number and prevent you from creating new accounts or apps. This comes from **personal experience** and there is NO INDICATION if and when that happens. AND Twitter support is TOTALLY WORTHLESS. This is when creating another cell phone number with Google Voice is helpful.
- You are not alone in thinking this is a HUGE PAIN IN THE ASS.
## Building a Basic Bot
If you skim the code below, you'll notice *there isn't a lot of code* that is because a bulk of the work is being done by a Python library called [Tweepy](http://tweepy.readthedocs.org/en/latest/). Tweepy handles all the annoying plumbing work involved with connecting to Twitter and accessing specific functions like authentication, search or updating your status. You could access the Twitter API directly by issuing raw HTTP request to Twitters [REST endpoints](https://dev.twitter.com/rest/public), but why do that when someone has already a nice and easy to use interface to Twitter?
The code below is for a very simple Twitter bot that just Tweets lines of Edgar Allen Poe's poetry.
### Boilerplate
```
# Import the python libraries we need to
import time
import tweepy
# Be sure to paste your keys and tokens in here because it won't work otherwise!
CONSUMER_KEY = "INSERT KEY HERE"
CONSUMER_SECRET = "INSERT SECRET HERE"
ACCESS_KEY = "INSERT ACCESS KEY"
ACCESS_SECRET = "INSERT SECRET HERE"
```
### The Oauth Dance
This is the code the initates a connection to Twitter via the API using the Keys and Access Tokens we created earlier. Once we've gotten through the "Oauth Dance" we'll have an object, `the_twitter_api` that we can use to programmatically perform actions on Twitter.
```
# Send our keys and tokens to Twitter
credentials = tweepy.OAuthHandler(CONSUMER_KEY, CONSUMER_SECRET)
credentials.set_access_token(ACCESS_KEY, ACCESS_SECRET)
# Authenticate with Twitter to get access
the_twitter_api = tweepy.API(credentials)
```
### Tweeting
We are going to make a bot that tweets lines from classic texts.
> I chose some texts from [Project Gutenberg](http://www.gutenberg.org/) and copied them into separate .txt files. (Maybe don’t choose a long-winded writer.) I ran a script over them to split them up by sentence and mark sentences longer than 140 characters. ([Link to chunking script.](https://github.com/robincamille/kicks-and-giggles/blob/master/poe-twitterbot-chunker.py)) There are other scripts to break up long sentences intelligently, but I wanted to exert some editorial control over where the splits occurred in the texts, so the script I wrote writes ‘SPLIT’ next to long sentences to alert me as I went over the ~600 lines by hand. I copied my chunked texts into one .txt file and marked the beginnings and ends of each individual text. ([Link to the finalized .txt file.](https://github.com/robincamille/kicks-and-giggles/blob/master/poe-lines.txt))
So I am going to cheat a little bit here and just use the data files Robin already prepared. I downloaded Robin's pre-prepared text into a file in the `data/` directory.
```
# open Robin's Edgar Allen Poe data file and read every line into memory
with open('data/thats-so-raven.txt','r') as filename:
lines = filename.readlines()
# Tweet each line, then wait one minute and tweet another.
# Note: this design means the bot runs continuously
for line in lines:
the_twitter_api.update_status(status=line)
print(line)
time.sleep(60) # Sleep for 1 minute
```
That's it you're done! That's all it takes to create a very simple Twitter bot.
## Next Steps
Here are some other bot examples to play with:
* Copora Tutorial Bot - A bot that uses [Corpora](https://github.com/dariusk/corpora) to generate text to tweet. [TODO]
* Mayor of Pittsburgh Bot - A bot that translates the Mayor's tweets into Pittsburghese. [TODO]
To make your own bot, you can start with this [python code for a very minimal bot](minimal-bot.ipynb) and modify it to cause whatever havoc or happiness you'd like.
### Deploying you Botnet
Making a bot is the easy part, making it run all the time requires a server that can continually run the bots. We have been creating and modifying these bots in Jupyter Notebooks which are fun and easy to use, but not great for deploying bots in the wild.
If you have a server, the best way to deploy a bot you've written in a notebook is to save the Notebook as a python script (File -> Download As -> Python (.py) and run the scrip or set up a [CRON JOB](http://www.unixgeeks.org/security/newbie/unix/cron-1.html) to execute the python script periodically (if you've written the script to just execute one and quit rather than run continuously).
| github_jupyter |
```
import pandas as pd
import numpy as np
import os
import faiss
from sklearn.preprocessing import StandardScaler, MinMaxScaler, QuantileTransformer
from plotly import offline
from sklearn.decomposition import TruncatedSVD
from MulticoreTSNE import MulticoreTSNE as TSNE
import umap
import plotly.plotly as py
import plotly.graph_objs as go
from plotly.offline import init_notebook_mode, iplot
files = os.listdir("../data")
def fix_array(x):
x = np.fromstring(
x.replace('\n','')
.replace('[','')
.replace(']','')
.replace(' ',' '), sep=' ')
return x.reshape((1, 768))
qa = pd.read_csv("../data/" + files[0])
for file in files[1:]:
print(file)
qa = pd.concat([qa, pd.read_csv("../data/" + file)], axis = 0)
qa.drop(["answer_bert", "question_bert", "Unnamed: 0"], axis = 1, inplace = True)
qa["Q_FFNN_embeds"] = qa["Q_FFNN_embeds"].apply(fix_array)
qa["A_FFNN_embeds"] = qa["A_FFNN_embeds"].apply(fix_array)
for n_items in range(100, 5000, 500):
for perplexity in range(1, 40, 5):
n_iters = 5000
qa = qa.sample(frac = 1)
qa.reset_index(inplace = True, drop = True)
question_bert = np.concatenate(qa["Q_FFNN_embeds"].values, axis=0)
answer_bert = np.concatenate(qa["A_FFNN_embeds"].values, axis=0)
question_bert = question_bert.astype('float32')
answer_bert = answer_bert.astype('float32')
answer_index = faiss.IndexFlatIP(answer_bert.shape[-1])
answer_index.add(answer_bert)
question_index = faiss.IndexFlatIP(question_bert.shape[-1])
question_index.add(question_bert)
k = len(question_bert)
D1, I1 = answer_index.search(question_bert[0:1].astype('float32'), k)
D2, I2 = question_index.search(question_bert[0:1].astype('float32'), k)
QT = QuantileTransformer()
D2 = (QT.fit_transform(-D2.T)**4) #* 20
D1 = (QT.fit_transform(-D1.T)**4) #* 20
closest_ind_q = list(I2[0, :n_items]) +list(I2[0, -n_items:])
closest_ind_a = list(I1[0, :n_items]) +list(I1[0, -n_items:])
dist_answers = answer_bert[closest_ind_a, :]
dist_questions = question_bert[closest_ind_q, :]
D1_answers = D1[closest_ind_a, :]
D2_questions = D2[closest_ind_q, :]
reducer = TSNE(n_components = 3, perplexity=perplexity, n_iter = n_iters)
reduced_dimensions = reducer.fit_transform(np.concatenate([dist_questions, dist_answers, answer_bert[0:1]], axis = 0))
question_bert_3d_close = reduced_dimensions[:n_items]
question_bert_3d_far = reduced_dimensions[n_items:n_items*2]
answer_bert_3d_close = reduced_dimensions[n_items*2:n_items*3]
answer_bert_3d_far = reduced_dimensions[n_items*3:-1]
question_bert_dist_close = D2_questions[:n_items]
question_bert_dist_far = D2_questions[n_items:n_items*2]
answer_bert_dist_close = D1_answers[n_items*2:n_items*3]
answer_bert_dist_far = D1_answers[n_items*3:-1]
init_notebook_mode(connected=True)
orig_q = go.Scatter3d(
name = "Original Question",
x=question_bert_3d_close[0:1,0],
y=question_bert_3d_close[0:1,1],
z=question_bert_3d_close[0:1,2],
mode='markers',
text = qa["question"].loc[closest_ind_q[:1]],
marker=dict(
size=12,
line=dict(
color='rgba(255, 0, 0, 0.14)',
width=0.1
),
opacity=1.0
)
)
orig_a = go.Scatter3d(
name = "Original Answer",
x=reduced_dimensions[-1:,0],
y=reduced_dimensions[-1:,1],
z=reduced_dimensions[-1:,2],
mode='markers',
text = qa["answer"][0:1],
marker=dict(
size=12,
line=dict(
color='rgba(0, 255, 0, 0.14)',
width=0.1
),
opacity=1.0
)
)
recommended_a = go.Scatter3d(
name = "Recommended Answers",
x=answer_bert_3d_close[0:5,0],
y=answer_bert_3d_close[0:5,1],
z=answer_bert_3d_close[0:5,2],
mode='markers',
text = qa["answer"].loc[closest_ind_a[:5]],
marker=dict(
size=12,
line=dict(
color='rgba(0, 255, 0, 0.14)',
width=0.1
),
opacity=1.0
)
)
close_q = go.Scatter3d(
name = "Similar Questions",
x=question_bert_3d_close[:,0],
y=question_bert_3d_close[:,1],
z=question_bert_3d_close[:,2],
mode='markers',
text = qa["question"].loc[closest_ind_q],
marker=dict(
size=question_bert_dist_close*16,
line=dict(
color='rgba(217, 217, 217, 0.14)',
width=0.1
),
opacity=0.8
)
)
close_a = go.Scatter3d(
name = "Similar Answers",
x=answer_bert_3d_close[5:,0],
y=answer_bert_3d_close[5:,1],
z=answer_bert_3d_close[5:,2],
mode='markers',
text = qa["answer"].loc[closest_ind_a],
marker=dict(
size=answer_bert_dist_close*16,
line=dict(
color='rgba(244, 100, 40, 0.14)',
width=0.1
),
opacity=0.8
)
)
far_q = go.Scatter3d(
name = "Dissimilar Questions",
x=question_bert_3d_far[:,0],
y=question_bert_3d_far[:,1],
z=question_bert_3d_far[:,2],
mode='markers',
text = qa["question"].loc[closest_ind_q],
marker=dict(
size=question_bert_dist_far,
line=dict(
color='rgba(40, 100, 217, 0.14)',
width=0.1
),
opacity=0.8
)
)
far_a = go.Scatter3d(
name = "Dissimilar Answers",
x=answer_bert_3d_far[:,0],
y=answer_bert_3d_far[:,1],
z=answer_bert_3d_far[:,2],
mode='markers',
text = qa["answer"].loc[closest_ind_a],
marker=dict(
size=answer_bert_dist_far,
line=dict(
color='rgba(255, 40, 40, 0.14)',
width=0.1
),
opacity=0.8
)
)
data = [orig_q, orig_a, close_q, close_a,
#far_q, far_a,
recommended_a
]
layout = go.Layout(
margin=dict(
l=0,
r=0,
b=0,
t=0
)
)
fig = go.Figure(data=data, layout=layout)
#iplot(fig, filename='simple-3d-scatter')
offline.plot(fig, filename="./experiments/n_items_" + str(n_items) + "_perplexity_" + str(perplexity) + '.html', auto_open=False)
```
| github_jupyter |
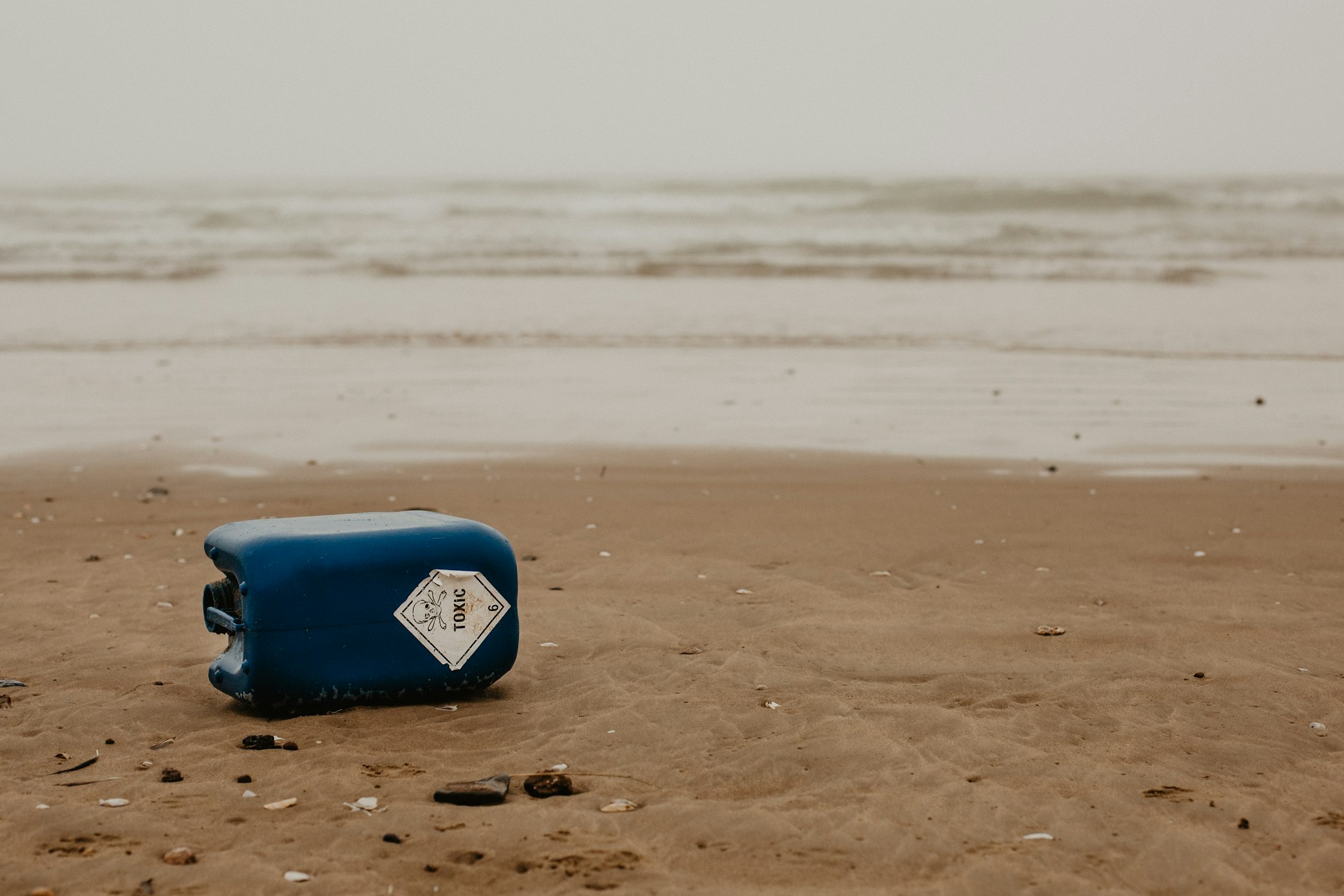

<div class = 'alert alert-block alert-info'
style = 'background-color:#4c1c84;
color:#eeebf1;
border-width:5px;
border-color:#4c1c84;
font-family:Comic Sans MS;
border-radius: 50px 50px'>
<p style = 'font-size:24px'>Exp 028</p>
<a href = "#Config"
style = "color:#eeebf1;
font-size:14px">1.Config</a><br>
<a href = "#Settings"
style = "color:#eeebf1;
font-size:14px">2.Settings</a><br>
<a href = "#Data-Load"
style = "color:#eeebf1;
font-size:14px">3.Data Load</a><br>
<a href = "#Pytorch-Settings"
style = "color:#eeebf1;
font-size:14px">4.Pytorch Settings</a><br>
<a href = "#Training"
style = "color:#eeebf1;
font-size:14px">5.Training</a><br>
</div>
<p style = 'font-size:24px;
color:#4c1c84'>
実施したこと
</p>
<li style = "color:#4c1c84;
font-size:14px">使用データ:Ruddit</li>
<li style = "color:#4c1c84;
font-size:14px">使用モデル:RoBERTa-Base</li>
<li style = "color:#4c1c84;
font-size:14px">New!! Attentionの可視化</li>
<br>
<h1 style = "font-size:45px; font-family:Comic Sans MS ; font-weight : normal; background-color: #4c1c84 ; color : #eeebf1; text-align: center; border-radius: 100px 100px;">
Config
</h1>
<br>
```
import sys
sys.path.append("../src/utils/iterative-stratification/")
sys.path.append("../src/utils/detoxify")
sys.path.append("../src/utils/coral-pytorch/")
import warnings
warnings.simplefilter('ignore')
import os
import gc
gc.enable()
import sys
import glob
import copy
import math
import time
import random
import string
import psutil
import pathlib
from pathlib import Path
from contextlib import contextmanager
from collections import defaultdict
from box import Box
from typing import Optional
from pprint import pprint
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import japanize_matplotlib
from tqdm.auto import tqdm as tqdmp
from tqdm.autonotebook import tqdm as tqdm
tqdmp.pandas()
## Model
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import StratifiedKFold, KFold
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.utils.data import Dataset, DataLoader
from transformers import AutoTokenizer, AutoModel, AdamW
from transformers import RobertaModel, RobertaForSequenceClassification
from transformers import RobertaTokenizer
from transformers import LukeTokenizer, LukeModel, LukeConfig
from transformers import get_linear_schedule_with_warmup, get_cosine_schedule_with_warmup
from transformers import BertTokenizer, BertForSequenceClassification
from transformers import RobertaTokenizer, RobertaForSequenceClassification
from transformers import XLMRobertaTokenizer, XLMRobertaForSequenceClassification
from transformers import DebertaTokenizer, DebertaModel
# Pytorch Lightning
import pytorch_lightning as pl
from pytorch_lightning.utilities.seed import seed_everything
from pytorch_lightning import callbacks
from pytorch_lightning.callbacks.progress import ProgressBarBase
from pytorch_lightning import LightningDataModule, LightningDataModule
from pytorch_lightning import Trainer
from pytorch_lightning.callbacks import ModelCheckpoint, EarlyStopping, LearningRateMonitor
from pytorch_lightning.loggers import WandbLogger
from pytorch_lightning.loggers.csv_logs import CSVLogger
from pytorch_lightning.callbacks import RichProgressBar
from sklearn.linear_model import Ridge
from sklearn.svm import SVC, SVR
from sklearn.feature_extraction.text import TfidfVectorizer
from scipy.stats import rankdata
from cuml.svm import SVR as cuml_SVR
from cuml.linear_model import Ridge as cuml_Ridge
import cudf
from detoxify import Detoxify
from iterstrat.ml_stratifiers import MultilabelStratifiedKFold
import torch
config = {
"exp_comment":"Jigsaw-UnbiasedをRoBERTaで学習",
"seed": 42,
"root": "/content/drive/MyDrive/kaggle/Jigsaw/raw",
"n_fold": 5,
"epoch": 5,
"max_length": 256,
"environment": "AWS",
"project": "Jigsaw",
"entity": "dataskywalker",
"exp_name": "028_exp",
"margin": 0.5,
"train_fold": [0],
"trainer": {
"gpus": 1,
"accumulate_grad_batches": 8,
"progress_bar_refresh_rate": 1,
"fast_dev_run": True,
"num_sanity_val_steps": 0,
},
"train_loader": {
"batch_size": 8,
"shuffle": True,
"num_workers": 1,
"pin_memory": True,
"drop_last": True,
},
"valid_loader": {
"batch_size": 8,
"shuffle": False,
"num_workers": 1,
"pin_memory": True,
"drop_last": False,
},
"test_loader": {
"batch_size": 8,
"shuffle": False,
"num_workers": 1,
"pin_memory": True,
"drop_last": False,
},
"backbone": {
"name": "roberta-base",
"output_dim": 1,
},
"optimizer": {
"name": "torch.optim.AdamW",
"params": {
"lr": 1e-6,
},
},
"scheduler": {
"name": "torch.optim.lr_scheduler.CosineAnnealingWarmRestarts",
"params": {
"T_0": 20,
"eta_min": 0,
},
},
"loss": "nn.BCEWithLogitsLoss",
}
config = Box(config)
config.tokenizer = RobertaTokenizer.from_pretrained(config.backbone.name)
config.model = RobertaModel.from_pretrained(config.backbone.name)
# pprint(config)
config.tokenizer.save_pretrained(f"../data/processed/{config.backbone.name}")
pretrain_model = RobertaModel.from_pretrained(config.backbone.name)
pretrain_model.save_pretrained(f"../data/processed/{config.backbone.name}")
# 個人的にAWSやKaggle環境やGoogle Colabを行ったり来たりしているのでまとめています
import os
import sys
from pathlib import Path
if config.environment == 'AWS':
INPUT_DIR = Path('/mnt/work/data/kaggle/Jigsaw/')
MODEL_DIR = Path(f'../models/{config.exp_name}/')
OUTPUT_DIR = Path(f'../data/interim/{config.exp_name}/')
UTIL_DIR = Path('/mnt/work/shimizu/kaggle/PetFinder/src/utils')
os.makedirs(MODEL_DIR, exist_ok=True)
os.makedirs(OUTPUT_DIR, exist_ok=True)
print(f"Your environment is 'AWS'.\nINPUT_DIR is {INPUT_DIR}\nMODEL_DIR is {MODEL_DIR}\nOUTPUT_DIR is {OUTPUT_DIR}\nUTIL_DIR is {UTIL_DIR}")
elif config.environment == 'Kaggle':
INPUT_DIR = Path('../input/*****')
MODEL_DIR = Path('./')
OUTPUT_DIR = Path('./')
print(f"Your environment is 'Kaggle'.\nINPUT_DIR is {INPUT_DIR}\nMODEL_DIR is {MODEL_DIR}\nOUTPUT_DIR is {OUTPUT_DIR}")
elif config.environment == 'Colab':
INPUT_DIR = Path('/content/drive/MyDrive/kaggle/Jigsaw/raw')
BASE_DIR = Path("/content/drive/MyDrive/kaggle/Jigsaw/interim")
MODEL_DIR = BASE_DIR / f'{config.exp_name}'
OUTPUT_DIR = BASE_DIR / f'{config.exp_name}/'
os.makedirs(MODEL_DIR, exist_ok=True)
os.makedirs(OUTPUT_DIR, exist_ok=True)
if not os.path.exists(INPUT_DIR):
print('Please Mount your Google Drive.')
else:
print(f"Your environment is 'Colab'.\nINPUT_DIR is {INPUT_DIR}\nMODEL_DIR is {MODEL_DIR}\nOUTPUT_DIR is {OUTPUT_DIR}")
else:
print("Please choose 'AWS' or 'Kaggle' or 'Colab'.\nINPUT_DIR is not found.")
# Seed固定
seed_everything(config.seed)
## 処理時間計測
@contextmanager
def timer(name:str, slack:bool=False):
t0 = time.time()
p = psutil.Process(os.getpid())
m0 = p.memory_info()[0] / 2. ** 30
print(f'<< {name} >> Start')
yield
m1 = p.memory_info()[0] / 2. ** 30
delta = m1 - m0
sign = '+' if delta >= 0 else '-'
delta = math.fabs(delta)
print(f"<< {name} >> {m1:.1f}GB({sign}{delta:.1f}GB):{time.time() - t0:.1f}sec", file=sys.stderr)
```
<br>
<h1 style = "font-size:45px; font-family:Comic Sans MS ; font-weight : normal; background-color: #4c1c84 ; color : #eeebf1; text-align: center; border-radius: 100px 100px;">
Data Load
</h1>
<br>
```
## Data Check
for dirnames, _, filenames in os.walk(INPUT_DIR):
for filename in filenames:
print(f'{dirnames}/{filename}')
val_df = pd.read_csv("/mnt/work/data/kaggle/Jigsaw/validation_data.csv")
test_df = pd.read_csv("/mnt/work/data/kaggle/Jigsaw/comments_to_score.csv")
display(val_df.head())
display(test_df.head())
```
<br>
<h2 style = "font-size:45px;
font-family:Comic Sans MS ;
font-weight : normal;
background-color: #eeebf1 ;
color : #4c1c84;
text-align: center;
border-radius: 100px 100px;">
Jigsaw Unbiased
</h2>
<br>
```
train_df = pd.read_csv("../data/external/jigsaw-unbiased/train.csv")
train_df = train_df.rename(columns={"target": "toxicity"})
display(train_df.head())
display(train_df.shape)
target_cols = [
"toxicity",
"severe_toxicity",
"identity_attack",
"insult",
"threat",
"sexual_explicit"
]
plt.figure(figsize=(12, 5))
sns.histplot(train_df["toxicity"], color="#4c1c84")
plt.grid()
plt.show()
def sample_df(df:pd.DataFrame, frac=0.2):
'''
train_dfからtoxicとnon_toxicを抽出
non_toxicの割合をfracで調整
'''
print(f"Before: {df.shape}")
label_cols = [
"toxicity",
"severe_toxicity",
"identity_attack",
"insult",
"threat",
"sexual_explicit"
]
df["y"] = df[label_cols].sum(axis=1)
df["y"] = df["y"]/df["y"].max()
toxic_df = df[df["y"]>0].reset_index(drop=True)
nontoxic_df = df[df["y"]==0].reset_index(drop=True)
nontoxic_df = nontoxic_df.sample(frac=frac, random_state=config.seed)
df = pd.concat([toxic_df, nontoxic_df], axis=0).sample(frac=1).reset_index(drop=True)
print(f"After: {df.shape}")
return df
with timer("sampling df"):
train_df = sample_df(train_df, frac=0.05)
display(train_df.head())
```
<br>
<h1 style = "font-size:45px; font-family:Comic Sans MS ; font-weight : normal; background-color: #4c1c84 ; color : #eeebf1; text-align: center; border-radius: 100px 100px;">
Pytorch Dataset
</h1>
<br>
```
class JigsawDataset:
def __init__(self, df, tokenizer, max_length, mode, target_cols):
self.df = df
self.max_len = max_length
self.tokenizer = tokenizer
self.mode = mode
self.target_cols = target_cols
if self.mode == "train":
self.text = df["comment_text"].values
self.target = df[target_cols].values
elif self.mode == "valid":
self.more_toxic = df["more_toxic"].values
self.less_toxic = df["less_toxic"].values
else:
self.text = df["text"].values
def __len__(self):
return len(self.df)
def __getitem__(self, index):
if self.mode == "train":
text = self.text[index]
target = self.target[index]
inputs_text = self.tokenizer.encode_plus(
text,
truncation=True,
return_attention_mask=True,
return_token_type_ids=True,
max_length = self.max_len,
padding="max_length",
)
text_ids = inputs_text["input_ids"]
text_mask = inputs_text["attention_mask"]
text_token_type_ids = inputs_text["token_type_ids"]
return {
'text_ids': torch.tensor(text_ids, dtype=torch.long),
'text_mask': torch.tensor(text_mask, dtype=torch.long),
'text_token_type_ids': torch.tensor(text_token_type_ids, dtype=torch.long),
'target': torch.tensor(target, dtype=torch.float)
}
elif self.mode == "valid":
more_toxic = self.more_toxic[index]
less_toxic = self.less_toxic[index]
inputs_more_toxic = self.tokenizer.encode_plus(
more_toxic,
truncation=True,
return_attention_mask=True,
return_token_type_ids=True,
max_length = self.max_len,
padding="max_length",
)
inputs_less_toxic = self.tokenizer.encode_plus(
less_toxic,
truncation=True,
return_attention_mask=True,
return_token_type_ids=True,
max_length = self.max_len,
padding="max_length",
)
target = 1
more_toxic_ids = inputs_more_toxic["input_ids"]
more_toxic_mask = inputs_more_toxic["attention_mask"]
more_token_type_ids = inputs_more_toxic["token_type_ids"]
less_toxic_ids = inputs_less_toxic["input_ids"]
less_toxic_mask = inputs_less_toxic["attention_mask"]
less_token_type_ids = inputs_less_toxic["token_type_ids"]
return {
'more_toxic_ids': torch.tensor(more_toxic_ids, dtype=torch.long),
'more_toxic_mask': torch.tensor(more_toxic_mask, dtype=torch.long),
'more_token_type_ids': torch.tensor(more_token_type_ids, dtype=torch.long),
'less_toxic_ids': torch.tensor(less_toxic_ids, dtype=torch.long),
'less_toxic_mask': torch.tensor(less_toxic_mask, dtype=torch.long),
'less_token_type_ids': torch.tensor(less_token_type_ids, dtype=torch.long),
'target': torch.tensor(target, dtype=torch.float)
}
else:
text = self.text[index]
inputs_text = self.tokenizer.encode_plus(
text,
truncation=True,
return_attention_mask=True,
return_token_type_ids=True,
max_length = self.max_len,
padding="max_length",
)
text_ids = inputs_text["input_ids"]
text_mask = inputs_text["attention_mask"]
text_token_type_ids = inputs_text["token_type_ids"]
return {
'text_ids': torch.tensor(text_ids, dtype=torch.long),
'text_mask': torch.tensor(text_mask, dtype=torch.long),
'text_token_type_ids': torch.tensor(text_token_type_ids, dtype=torch.long),
}
```
<br>
<h2 style = "font-size:45px;
font-family:Comic Sans MS ;
font-weight : normal;
background-color: #eeebf1 ;
color : #4c1c84;
text-align: center;
border-radius: 100px 100px;">
DataModule
</h2>
<br>
```
class JigsawDataModule(LightningDataModule):
def __init__(self, train_df, valid_df, test_df, cfg):
super().__init__()
self._train_df = train_df
self._valid_df = valid_df
self._test_df = test_df
self._cfg = cfg
def train_dataloader(self):
dataset = JigsawDataset(
df=self._train_df,
tokenizer=self._cfg.tokenizer,
max_length=self._cfg.max_length,
mode="train",
target_cols=target_cols
)
return DataLoader(dataset, **self._cfg.train_loader)
def val_dataloader(self):
dataset = JigsawDataset(
df=self._valid_df,
tokenizer=self._cfg.tokenizer,
max_length=self._cfg.max_length,
mode="valid",
target_cols=target_cols
)
return DataLoader(dataset, **self._cfg.valid_loader)
def test_dataloader(self):
dataset = JigsawDataset(
df=self._test_df,
tokenizer = self._cfg.tokenizer,
max_length=self._cfg.max_length,
mode="test",
target_cols=target_cols
)
return DataLoader(dataset, **self._cfg.test_loader)
## DataCheck
seed_everything(config.seed)
sample_dataloader = JigsawDataModule(train_df, val_df, test_df, config).train_dataloader()
for data in sample_dataloader:
break
print(data["text_ids"].size())
print(data["text_mask"].size())
print(data["text_token_type_ids"].size())
print(data["target"].size())
print(data["target"])
output = config.model(
data["text_ids"],
data["text_mask"],
data["text_token_type_ids"],
output_attentions=True
)
print(output["last_hidden_state"].size(), output["attentions"][-1].size())
print(output["last_hidden_state"][:, 0, :].size(), output["attentions"][-1].size())
```
<br>
<h2 style = "font-size:45px;
font-family:Comic Sans MS ;
font-weight : normal;
background-color: #eeebf1 ;
color : #4c1c84;
text-align: center;
border-radius: 100px 100px;">
LigitningModule
</h2>
<br>
```
class JigsawModel(pl.LightningModule):
def __init__(self, cfg, fold_num):
super().__init__()
self.cfg = cfg
self.__build_model()
self.criterion = eval(self.cfg.loss)()
self.save_hyperparameters(cfg)
self.fold_num = fold_num
def __build_model(self):
self.base_model = RobertaModel.from_pretrained(
self.cfg.backbone.name
)
print(f"Use Model: {self.cfg.backbone.name}")
self.norm = nn.LayerNorm(768)
self.drop = nn.Dropout(p=0.3)
self.head = nn.Linear(768, self.cfg.backbone.output_dim)
def forward(self, ids, mask, token_type_ids):
output = self.base_model(
input_ids=ids,
attention_mask=mask,
token_type_ids=token_type_ids,
output_attentions=True
)
feature = self.norm(output["last_hidden_state"][:, 0, :])
out = self.drop(feature)
out = self.head(out)
return {
"logits":out,
"attention":output["attentions"],
"mask":mask,
}
def training_step(self, batch, batch_idx):
text_ids = batch["text_ids"]
text_mask = batch['text_mask']
text_token_type_ids = batch['text_token_type_ids']
targets = batch['target']
outputs = self.forward(text_ids, text_mask, text_token_type_ids)
loss = self.criterion(outputs["logits"], targets)
return {
"loss":loss,
"targets":targets,
}
def training_epoch_end(self, training_step_outputs):
loss_list = []
for out in training_step_outputs:
loss_list.extend([out["loss"].cpu().detach().tolist()])
meanloss = sum(loss_list)/len(loss_list)
logs = {f"train_loss/fold{self.fold_num+1}": meanloss,}
self.log_dict(
logs,
on_step=False,
on_epoch=True,
prog_bar=True,
logger=True
)
def validation_step(self, batch, batch_idx):
more_toxic_ids = batch['more_toxic_ids']
more_toxic_mask = batch['more_toxic_mask']
more_text_token_type_ids = batch['more_token_type_ids']
less_toxic_ids = batch['less_toxic_ids']
less_toxic_mask = batch['less_toxic_mask']
less_text_token_type_ids = batch['less_token_type_ids']
targets = batch['target']
more_outputs = self.forward(
more_toxic_ids,
more_toxic_mask,
more_text_token_type_ids
)
less_outputs = self.forward(
less_toxic_ids,
less_toxic_mask,
less_text_token_type_ids
)
more_outputs = torch.sum(more_outputs["logits"], 1)
less_outputs = torch.sum(less_outputs["logits"], 1)
outputs = more_outputs - less_outputs
logits = outputs.clone()
logits[logits > 0] = 1
loss = self.criterion(logits, targets)
return {
"loss":loss,
"pred":outputs,
"targets":targets,
}
def validation_epoch_end(self, validation_step_outputs):
loss_list = []
pred_list = []
target_list = []
for out in validation_step_outputs:
loss_list.extend([out["loss"].cpu().detach().tolist()])
pred_list.append(out["pred"].detach().cpu().numpy())
target_list.append(out["targets"].detach().cpu().numpy())
meanloss = sum(loss_list)/len(loss_list)
pred_list = np.concatenate(pred_list)
pred_count = sum(x>0 for x in pred_list)/len(pred_list)
logs = {
f"valid_loss/fold{self.fold_num+1}":meanloss,
f"valid_acc/fold{self.fold_num+1}":pred_count,
}
self.log_dict(
logs,
on_step=False,
on_epoch=True,
prog_bar=True,
logger=True
)
def configure_optimizers(self):
optimizer = eval(self.cfg.optimizer.name)(
self.parameters(), **self.cfg.optimizer.params
)
self.scheduler = eval(self.cfg.scheduler.name)(
optimizer, **self.cfg.scheduler.params
)
scheduler = {"scheduler": self.scheduler, "interval": "step",}
return [optimizer], [scheduler]
```
<br>
<h2 style = "font-size:45px;
font-family:Comic Sans MS ;
font-weight : normal;
background-color: #eeebf1 ;
color : #4c1c84;
text-align: center;
border-radius: 100px 100px;">
Training
</h2>
<br>
```
skf = KFold(
n_splits=config.n_fold,
shuffle=True,
random_state=config.seed
)
for fold, (_, val_idx) in enumerate(skf.split(X=train_df, y=train_df[target_cols])):
train_df.loc[val_idx, "kfold"] = int(fold)
train_df["kfold"] = train_df["kfold"].astype(int)
train_df["kfold"] = -1
train_df.head()
## Debug
config.trainer.fast_dev_run = True
config.backbone.output_dim = len(target_cols)
for fold in config.train_fold:
print("★"*25, f" Fold{fold+1} ", "★"*25)
df_train = train_df[train_df.kfold != fold].reset_index(drop=True)
datamodule = JigsawDataModule(df_train, val_df, test_df, config)
sample_dataloader = JigsawDataModule(df_train, val_df, test_df, config).train_dataloader()
config.scheduler.params.T_0 = config.epoch * len(sample_dataloader)
model = JigsawModel(config, fold)
lr_monitor = callbacks.LearningRateMonitor()
loss_checkpoint = callbacks.ModelCheckpoint(
filename=f"best_acc_fold{fold+1}",
monitor=f"valid_acc/fold{fold+1}",
save_top_k=1,
mode="max",
save_last=False,
dirpath=MODEL_DIR,
save_weights_only=True,
)
wandb_logger = WandbLogger(
project=config.project,
entity=config.entity,
name = f"{config.exp_name}",
tags = ['RoBERTa-Base', "Jigsaw-Unbiased"]
)
lr_monitor = LearningRateMonitor(logging_interval='step')
trainer = pl.Trainer(
max_epochs=config.epoch,
callbacks=[loss_checkpoint, lr_monitor, RichProgressBar()],
# deterministic=True,
logger=[wandb_logger],
**config.trainer
)
trainer.fit(model, datamodule=datamodule)
## Training
config.trainer.fast_dev_run = False
config.backbone.output_dim = len(target_cols)
for fold in config.train_fold:
print("★"*25, f" Fold{fold+1} ", "★"*25)
df_train = train_df[train_df.kfold != fold].reset_index(drop=True)
datamodule = JigsawDataModule(df_train, val_df, test_df, config)
sample_dataloader = JigsawDataModule(df_train, val_df, test_df, config).train_dataloader()
config.scheduler.params.T_0 = config.epoch * len(sample_dataloader)
model = JigsawModel(config, fold)
lr_monitor = callbacks.LearningRateMonitor()
loss_checkpoint = callbacks.ModelCheckpoint(
filename=f"best_acc_fold{fold+1}",
monitor=f"valid_acc/fold{fold+1}",
save_top_k=1,
mode="max",
save_last=False,
dirpath=MODEL_DIR,
save_weights_only=True,
)
wandb_logger = WandbLogger(
project=config.project,
entity=config.entity,
name = f"{config.exp_name}",
tags = ['RoBERTa-Base', "Jigsaw-Unbiased"]
)
lr_monitor = LearningRateMonitor(logging_interval='step')
trainer = pl.Trainer(
max_epochs=config.epoch,
callbacks=[loss_checkpoint, lr_monitor, RichProgressBar()],
# deterministic=True,
logger=[wandb_logger],
**config.trainer
)
trainer.fit(model, datamodule=datamodule)
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
config.backbone.output_dim = len(target_cols)
print(f"Device == {device}")
MORE = np.zeros((len(val_df), config.backbone.output_dim))
LESS = np.zeros((len(val_df), config.backbone.output_dim))
PRED = np.zeros((len(test_df), config.backbone.output_dim))
attention_array = np.zeros((len(val_df), 256)) # attention格納
mask_array = np.zeros((len(val_df), 256)) # mask情報格納,後でattentionと掛け合わせる
for fold in config.train_fold:
pred_list = []
print("★"*25, f" Fold{fold+1} ", "★"*25)
valid_dataloader = JigsawDataModule(train_df, val_df, test_df, config).val_dataloader()
model = JigsawModel(config, fold)
loss_checkpoint = callbacks.ModelCheckpoint(
filename=f"best_acc_fold{fold+1}",
monitor=f"valid_acc/fold{fold+1}",
save_top_k=1,
mode="max",
save_last=False,
dirpath="../input/toxicroberta/",
)
model = model.load_from_checkpoint(MODEL_DIR/f"best_acc_fold{fold+1}.ckpt", cfg=config, fold_num=fold)
model.to(device)
model.eval()
more_list = []
less_list = []
for step, data in tqdm(enumerate(valid_dataloader), total=len(valid_dataloader)):
more_toxic_ids = data['more_toxic_ids'].to(device)
more_toxic_mask = data['more_toxic_mask'].to(device)
more_text_token_type_ids = data['more_token_type_ids'].to(device)
less_toxic_ids = data['less_toxic_ids'].to(device)
less_toxic_mask = data['less_toxic_mask'].to(device)
less_text_token_type_ids = data['less_token_type_ids'].to(device)
more_outputs = model(
more_toxic_ids,
more_toxic_mask,
more_text_token_type_ids,
)
less_outputs = model(
less_toxic_ids,
less_toxic_mask,
less_text_token_type_ids
)
more_list.append(more_outputs["logits"].detach().cpu().numpy())
less_list.append(less_outputs["logits"].detach().cpu().numpy())
MORE += np.concatenate(more_list)/len(config.train_fold)
LESS += np.concatenate(less_list)/len(config.train_fold)
# PRED += pred_list/len(config.train_fold)
plt.figure(figsize=(12, 5))
plt.scatter(LESS, MORE)
plt.xlabel("less-toxic")
plt.ylabel("more-toxic")
plt.grid()
plt.show()
val_df["less_attack"] = LESS.sum(axis=1)
val_df["more_attack"] = MORE.sum(axis=1)
val_df["diff_attack"] = val_df["more_attack"] - val_df["less_attack"]
attack_score = val_df[val_df["diff_attack"]>0]["diff_attack"].count()/len(val_df)
print(f"Wiki Attack Score: {attack_score:.6f}")
```
| github_jupyter |
# Linear Regression - Ecommerce Project
You just got some contract work with an Ecommerce company based in New York City that sells clothing online but they also have in-store style and clothing advice sessions. Customers come in to the store, have sessions/meetings with a personal stylist, then they can go home and order either on a mobile app or website for the clothes they want.
The company is trying to decide whether to focus their efforts on their mobile app experience or their website. They've hired you on contract to help them figure it out! Let's get started!
Note: Data are fake data.
## Imports
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
```
## Get the Data
We'll work with the Ecommerce Customers csv file from the company. It has Customer info, suchas Email, Address, and their color Avatar. Then it also has numerical value columns:
* Avg. Session Length: Average session of in-store style advice sessions.
* Time on App: Average time spent on App in minutes
* Time on Website: Average time spent on Website in minutes
* Length of Membership: How many years the customer has been a member.
```
df = pd.read_csv('Ecommerce Customers')
df.head()
df.info()
df.describe().T
```
-------
# Exploratory Data Analysis
**Let's explore the data!**
For the rest of the exercise we'll only be using the numerical data of the csv file.
___
**Use seaborn to create a jointplot to compare the Time on Website and Yearly Amount Spent columns. Does the correlation make sense?**
```
sns.jointplot(data=df, x='Time on Website', y='Yearly Amount Spent');
```
There seems to be no clear correlation between `Time on Website` and `Yearly Amount Spent`.
**Do the same but with the Time on App column instead.**
```
sns.jointplot(data=df, x='Time on App', y='Yearly Amount Spent');
```
We can see that there is some kind of positive correlation between `Time on App` and `Yearly Amount Spent`.
--------
**Use jointplot to create a 2D hex bin plot comparing Time on App and Length of Membership.**
```
df.columns
sns.jointplot(data=df, x='Time on App', y='Length of Membership', kind='hex');
```
There is some kind of cluster happening in the middle. Peoeple who have been members for between 3 to 4 tend to spend more time on the App.
--------
**Let's explore these types of relationships across the entire data set. Use [pairplot](https://stanford.edu/~mwaskom/software/seaborn/tutorial/axis_grids.html#plotting-pairwise-relationships-with-pairgrid-and-pairplot) to recreate the plot below.(Don't worry about the the colors)**
```
sns.pairplot(df);
```
**Based off this plot what looks to be the most correlated feature with Yearly Amount Spent?**
* We can see that most of the data are normallly distributed.
* There is some positive correlation between `Yearly Amount Spent` and feaures like `Lenght of Membership` and `Time on App`.
**Create a linear model plot (using seaborn's lmplot) of Yearly Amount Spent vs. Length of Membership.**
```
sns.lmplot(data=df, x='Length of Membership', y='Yearly Amount Spent');
```
# Training and Testing Data
Now that we've explored the data a bit, let's go ahead and split the data into training and testing sets.
**Set a variable X equal to the numerical features of the customers and a variable y equal to the "Yearly Amount Spent" column.**
```
df.columns
# separate features and labels
X = df[['Avg. Session Length', 'Time on App','Time on Website', 'Length of Membership']]
y = df[ 'Yearly Amount Spent']
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=101)
X_train.shape, X_test.shape, y_train.shape, y_test.shape
```
## Training the Model
```
from sklearn.linear_model import LinearRegression
lr = LinearRegression()
lr.fit(X_train, y_train)
```
**Print out the coefficients of the model**
```
lr.coef_
df_coef = pd.DataFrame(data=lr.coef_, index=X_train.columns, columns=['Coefficient'])
df_coef.sort_values(by='Coefficient', ascending=False)
```
### Intrepreting Coefficient
Holding all other features fixed,
* one unit increase in `Length of Membership` is associated with an increase of \$61.28 in `Yearly Amount Spent`.
* one unit increase in `Time on App` is associated with an increase of \$38.59 in `Yearly Amount Spent`.
* one unit increase in `TAvg. Session Length` is associated with an increase of \$25.98 in `Yearly Amount Spent`.
* one unit increase in `Time on Website` is associated with an increase of \$0.19 in `Yearly Amount Spent`.
## Predicting Test Data
Now that we have fit our model, let's evaluate its performance by predicting off the test values!
**Create a scatterplot of the real test values versus the predicted values.**
```
predictions = lr.predict(X_test)
plt.scatter(y_test, predictions);
```
## Evaluating the Model
Let's evaluate our model performance by calculating the residual sum of squares and the explained variance score (R^2).
```
from sklearn.metrics import mean_absolute_error, mean_squared_error, explained_variance_score
print('MAE : {}'.format(mean_absolute_error(y_test, predictions)))
print('MSE : {}'.format(mean_squared_error(y_test, predictions)))
print('RMSE : {}'.format(np.sqrt(mean_squared_error(y_test, predictions))))
print('Explained Variance Score: {}'.format(explained_variance_score(y_test, predictions)))
```
## Residuals
You should have gotten a very good model with a good fit. Let's quickly explore the residuals to make sure everything was okay with our data.
**Plot a histogram of the residuals and make sure it looks normally distributed. Use either seaborn distplot, or just plt.hist().**
```
sns.displot(y_test - predictions, kde=True, bins=50);
```
It seem a good Residual Historgram as the shape is pretty normally distributed.
## Conclusion
We still want to figure out the answer to the original question, do we focus our efforst on mobile app or website development? Or maybe that doesn't even really matter, and Membership Time is what is really important. Let's see if we can interpret the coefficients at all to get an idea.
**Recreate the dataframe below. **
```
df_coef.sort_values(by='Coefficient', ascending=False)
```
**How can you interpret these coefficients?**
Holding all other features fixed,
* one unit increase in `Length of Membership` is associated with an increase of \$61.28 in `Yearly Amount Spent`.
* one unit increase in `Time on App` is associated with an increase of \$38.59 in `Yearly Amount Spent`.
* one unit increase in `TAvg. Session Length` is associated with an increase of \$25.98 in `Yearly Amount Spent`.
* one unit increase in `Time on Website` is associated with an increase of \$0.19 in `Yearly Amount Spent`.
**Do you think the company should focus more on their mobile app or on their website?**
```
38.590159/0.190405
```
There are multiple suggestions.
* Currently,**Mobile App** because its coefficient yields nearly 200 times for `Yearly Amount Spent` than **Website**.
* So What we can do is Company focus on **Website** to make improvements so that it can catch up with Mobile app.
* On the other hand, if company doesn't want to make changes to website and want to focus on currently performing platform, then they should focus efforts more on **Mobile App**.
* Lastly, we might want to dig deeper for `Length of Membership` between its relationship with Mobile or Website.
## Great Job!
Congrats on your contract work! The company loved the insights! Let's move on.
| github_jupyter |
# Matrix generation
## Init symbols for *sympy*
```
from sympy import *
from geom_util import *
from sympy.vector import CoordSys3D
N = CoordSys3D('N')
alpha1, alpha2, alpha3 = symbols("alpha_1 alpha_2 alpha_3", real = True, positive=True)
init_printing()
%matplotlib inline
%reload_ext autoreload
%autoreload 2
%aimport geom_util
```
### Lame params
```
h1 = Function("H1")
h2 = Function("H2")
h3 = Function("H3")
H1 = h1(alpha1, alpha2, alpha3)
H2 = S(1
H3 = h3(alpha1, alpha2, alpha3)
```
### Metric tensor
${\displaystyle \hat{G}=\sum_{i,j} g^{ij}\vec{R}_i\vec{R}_j}$
```
G_up = getMetricTensorUpLame(H1, H2, H3)
```
${\displaystyle \hat{G}=\sum_{i,j} g_{ij}\vec{R}^i\vec{R}^j}$
```
G_down = getMetricTensorDownLame(H1, H2, H3)
```
### Christoffel symbols
```
GK = getChristoffelSymbols2(G_up, G_down, (alpha1, alpha2, alpha3))
```
### Gradient of vector
$
\left(
\begin{array}{c}
\nabla_1 u_1 \\ \nabla_2 u_1 \\ \nabla_3 u_1 \\
\nabla_1 u_2 \\ \nabla_2 u_2 \\ \nabla_3 u_2 \\
\nabla_1 u_3 \\ \nabla_2 u_3 \\ \nabla_3 u_3 \\
\end{array}
\right)
=
B \cdot
\left(
\begin{array}{c}
u_1 \\
\frac { \partial u_1 } { \partial \alpha_1} \\
\frac { \partial u_1 } { \partial \alpha_2} \\
\frac { \partial u_1 } { \partial \alpha_3} \\
u_2 \\
\frac { \partial u_2 } { \partial \alpha_1} \\
\frac { \partial u_2 } { \partial \alpha_2} \\
\frac { \partial u_2 } { \partial \alpha_3} \\
u_3 \\
\frac { \partial u_3 } { \partial \alpha_1} \\
\frac { \partial u_3 } { \partial \alpha_2} \\
\frac { \partial u_3 } { \partial \alpha_3} \\
\end{array}
\right)
= B \cdot D \cdot
\left(
\begin{array}{c}
u^1 \\
\frac { \partial u^1 } { \partial \alpha_1} \\
\frac { \partial u^1 } { \partial \alpha_2} \\
\frac { \partial u^1 } { \partial \alpha_3} \\
u^2 \\
\frac { \partial u^2 } { \partial \alpha_1} \\
\frac { \partial u^2 } { \partial \alpha_2} \\
\frac { \partial u^2 } { \partial \alpha_3} \\
u^3 \\
\frac { \partial u^3 } { \partial \alpha_1} \\
\frac { \partial u^3 } { \partial \alpha_2} \\
\frac { \partial u^3 } { \partial \alpha_3} \\
\end{array}
\right)
$
```
def row_index_to_i_j_grad(i_row):
return i_row // 3, i_row % 3
B = zeros(9, 12)
B[0,1] = S(1)
B[1,2] = S(1)
B[2,3] = S(1)
B[3,5] = S(1)
B[4,6] = S(1)
B[5,7] = S(1)
B[6,9] = S(1)
B[7,10] = S(1)
B[8,11] = S(1)
for row_index in range(9):
i,j=row_index_to_i_j_grad(row_index)
B[row_index, 0] = -GK[i,j,0]
B[row_index, 4] = -GK[i,j,1]
B[row_index, 8] = -GK[i,j,2]
B
```
### Strain tensor
$
\left(
\begin{array}{c}
\varepsilon_{11} \\
\varepsilon_{22} \\
\varepsilon_{33} \\
2\varepsilon_{12} \\
2\varepsilon_{13} \\
2\varepsilon_{23} \\
\end{array}
\right)
=
\left(E + E_{NL} \left( \nabla \vec{u} \right) \right) \cdot
\left(
\begin{array}{c}
\nabla_1 u_1 \\ \nabla_2 u_1 \\ \nabla_3 u_1 \\
\nabla_1 u_2 \\ \nabla_2 u_2 \\ \nabla_3 u_2 \\
\nabla_1 u_3 \\ \nabla_2 u_3 \\ \nabla_3 u_3 \\
\end{array}
\right)$
```
E=zeros(6,9)
E[0,0]=1
E[1,4]=1
E[2,8]=1
E[3,1]=1
E[3,3]=1
E[4,2]=1
E[4,6]=1
E[5,5]=1
E[5,7]=1
E
def E_NonLinear(grad_u):
N = 3
du = zeros(N, N)
# print("===Deformations===")
for i in range(N):
for j in range(N):
index = i*N+j
du[j,i] = grad_u[index]
# print("========")
a_values = S(1)/S(2) * du * G_up
E_NL = zeros(6,9)
E_NL[0,0] = a_values[0,0]
E_NL[0,3] = a_values[0,1]
E_NL[0,6] = a_values[0,2]
E_NL[1,1] = a_values[1,0]
E_NL[1,4] = a_values[1,1]
E_NL[1,7] = a_values[1,2]
E_NL[2,2] = a_values[2,0]
E_NL[2,5] = a_values[2,1]
E_NL[2,8] = a_values[2,2]
E_NL[3,1] = 2*a_values[0,0]
E_NL[3,4] = 2*a_values[0,1]
E_NL[3,7] = 2*a_values[0,2]
E_NL[4,0] = 2*a_values[2,0]
E_NL[4,3] = 2*a_values[2,1]
E_NL[4,6] = 2*a_values[2,2]
E_NL[5,2] = 2*a_values[1,0]
E_NL[5,5] = 2*a_values[1,1]
E_NL[5,8] = 2*a_values[1,2]
return E_NL
%aimport geom_util
#u=getUHat3D(alpha1, alpha2, alpha3)
u=getUHatU3Main(alpha1, alpha2, alpha3)
gradu=B*u
E_NL = E_NonLinear(gradu)*B
```
### Physical coordinates
$u_i=u_{[i]} H_i$
```
P=zeros(12,12)
P[0,0]=H1
P[1,0]=(H1).diff(alpha1)
P[1,1]=H1
P[2,0]=(H1).diff(alpha2)
P[2,2]=H1
P[3,0]=(H1).diff(alpha3)
P[3,3]=H1
P[4,4]=H2
P[5,4]=(H2).diff(alpha1)
P[5,5]=H2
P[6,4]=(H2).diff(alpha2)
P[6,6]=H2
P[7,4]=(H2).diff(alpha3)
P[7,7]=H2
P[8,8]=H3
P[9,8]=(H3).diff(alpha1)
P[9,9]=H3
P[10,8]=(H3).diff(alpha2)
P[10,10]=H3
P[11,8]=(H3).diff(alpha3)
P[11,11]=H3
P=simplify(P)
P
B_P = zeros(9,9)
for i in range(3):
for j in range(3):
ratio=1
if (i==0):
ratio = ratio*H1
elif (i==1):
ratio = ratio*H2
elif (i==2):
ratio = ratio*H3
if (j==0):
ratio = ratio*H1
elif (j==1):
ratio = ratio*H2
elif (j==2):
ratio = ratio*H3
row_index = i*3+j
B_P[row_index, row_index] = 1/ratio
Grad_U_P = simplify(B_P*B*P)
B_P
StrainL=simplify(E*Grad_U_P)
StrainL
%aimport geom_util
u=getUHatU3Main(alpha1, alpha2, alpha3)
gradup=B_P*B*P*u
E_NLp = E_NonLinear(gradup)*B*P*u
simplify(E_NLp)
```
### Tymoshenko theory
$u^1 \left( \alpha_1, \alpha_2, \alpha_3 \right)=u\left( \alpha_1 \right)+\alpha_3\gamma \left( \alpha_1 \right) $
$u^2 \left( \alpha_1, \alpha_2, \alpha_3 \right)=0 $
$u^3 \left( \alpha_1, \alpha_2, \alpha_3 \right)=w\left( \alpha_1 \right) $
$ \left(
\begin{array}{c}
u^1 \\
\frac { \partial u^1 } { \partial \alpha_1} \\
\frac { \partial u^1 } { \partial \alpha_2} \\
\frac { \partial u^1 } { \partial \alpha_3} \\
u^2 \\
\frac { \partial u^2 } { \partial \alpha_1} \\
\frac { \partial u^2 } { \partial \alpha_2} \\
\frac { \partial u^2 } { \partial \alpha_3} \\
u^3 \\
\frac { \partial u^3 } { \partial \alpha_1} \\
\frac { \partial u^3 } { \partial \alpha_2} \\
\frac { \partial u^3 } { \partial \alpha_3} \\
\end{array}
\right) = T \cdot
\left(
\begin{array}{c}
u \\
\frac { \partial u } { \partial \alpha_1} \\
\gamma \\
\frac { \partial \gamma } { \partial \alpha_1} \\
w \\
\frac { \partial w } { \partial \alpha_1} \\
\end{array}
\right) $
```
T=zeros(12,6)
T[0,0]=1
T[0,2]=alpha3
T[1,1]=1
T[1,3]=alpha3
T[3,2]=1
T[8,4]=1
T[9,5]=1
T
D_p_T = StrainL*T
simplify(D_p_T)
u = Function("u")
t = Function("theta")
w = Function("w")
u1=u(alpha1)+alpha3*t(alpha1)
u3=w(alpha1)
gu = zeros(12,1)
gu[0] = u1
gu[1] = u1.diff(alpha1)
gu[3] = u1.diff(alpha3)
gu[8] = u3
gu[9] = u3.diff(alpha1)
gradup=Grad_U_P*gu
# o20=(K*u(alpha1)-w(alpha1).diff(alpha1)+t(alpha1))/2
# o21=K*t(alpha1)
# O=1/2*o20*o20+alpha3*o20*o21-alpha3*K/2*o20*o20
# O=expand(O)
# O=collect(O,alpha3)
# simplify(O)
StrainNL = E_NonLinear(gradup)*gradup
simplify(StrainNL)
```
### Square theory
$u^1 \left( \alpha_1, \alpha_2, \alpha_3 \right)=u_{10}\left( \alpha_1 \right)p_0\left( \alpha_3 \right)+u_{11}\left( \alpha_1 \right)p_1\left( \alpha_3 \right)+u_{12}\left( \alpha_1 \right)p_2\left( \alpha_3 \right) $
$u^2 \left( \alpha_1, \alpha_2, \alpha_3 \right)=0 $
$u^3 \left( \alpha_1, \alpha_2, \alpha_3 \right)=u_{30}\left( \alpha_1 \right)p_0\left( \alpha_3 \right)+u_{31}\left( \alpha_1 \right)p_1\left( \alpha_3 \right)+u_{32}\left( \alpha_1 \right)p_2\left( \alpha_3 \right) $
$ \left(
\begin{array}{c}
u^1 \\
\frac { \partial u^1 } { \partial \alpha_1} \\
\frac { \partial u^1 } { \partial \alpha_2} \\
\frac { \partial u^1 } { \partial \alpha_3} \\
u^2 \\
\frac { \partial u^2 } { \partial \alpha_1} \\
\frac { \partial u^2 } { \partial \alpha_2} \\
\frac { \partial u^2 } { \partial \alpha_3} \\
u^3 \\
\frac { \partial u^3 } { \partial \alpha_1} \\
\frac { \partial u^3 } { \partial \alpha_2} \\
\frac { \partial u^3 } { \partial \alpha_3} \\
\end{array}
\right) = L \cdot
\left(
\begin{array}{c}
u_{10} \\
\frac { \partial u_{10} } { \partial \alpha_1} \\
u_{11} \\
\frac { \partial u_{11} } { \partial \alpha_1} \\
u_{12} \\
\frac { \partial u_{12} } { \partial \alpha_1} \\
u_{30} \\
\frac { \partial u_{30} } { \partial \alpha_1} \\
u_{31} \\
\frac { \partial u_{31} } { \partial \alpha_1} \\
u_{32} \\
\frac { \partial u_{32} } { \partial \alpha_1} \\
\end{array}
\right) $
```
L=zeros(12,12)
h=Symbol('h')
p0=1/2-alpha3/h
p1=1/2+alpha3/h
p2=1-(2*alpha3/h)**2
L[0,0]=p0
L[0,2]=p1
L[0,4]=p2
L[1,1]=p0
L[1,3]=p1
L[1,5]=p2
L[3,0]=p0.diff(alpha3)
L[3,2]=p1.diff(alpha3)
L[3,4]=p2.diff(alpha3)
L[8,6]=p0
L[8,8]=p1
L[8,10]=p2
L[9,7]=p0
L[9,9]=p1
L[9,11]=p2
L[11,6]=p0.diff(alpha3)
L[11,8]=p1.diff(alpha3)
L[11,10]=p2.diff(alpha3)
L
B_General = zeros(9, 12)
B_General[0,1] = S(1)
B_General[1,2] = S(1)
B_General[2,3] = S(1)
B_General[3,5] = S(1)
B_General[4,6] = S(1)
B_General[5,7] = S(1)
B_General[6,9] = S(1)
B_General[7,10] = S(1)
B_General[8,11] = S(1)
for row_index in range(9):
i,j=row_index_to_i_j_grad(row_index)
B_General[row_index, 0] = -Symbol("G_{{{}{}}}^1".format(i+1,j+1))
B_General[row_index, 4] = -Symbol("G_{{{}{}}}^2".format(i+1,j+1))
B_General[row_index, 8] = -Symbol("G_{{{}{}}}^3".format(i+1,j+1))
B_General
simplify(B_General*L)
simplify(E*B*D*P*L)
D_p_L = StrainL*L
K = Symbol('K')
D_p_L = D_p_L.subs(R, 1/K)
simplify(D_p_L)
h = 0.5
exp=(0.5-alpha3/h)*(1-(2*alpha3/h)**2)#/(1+alpha3*0.8)
p02=integrate(exp, (alpha3, -h/2, h/2))
integral = expand(simplify(p02))
integral
```
## Mass matrix
```
rho=Symbol('rho')
B_h=zeros(3,12)
B_h[0,0]=1
B_h[1,4]=1
B_h[1,8]=1
M=simplify(rho*P.T*B_h.T*G_up*B_h*P)
M
```
| github_jupyter |
# Progressive Distillation for Fast Sampling of Diffusion Models
Code for the <a href="https://openreview.net/forum?id=TIdIXIpzhoI">ICLR 2022 paper</a> by Tim Salimans and Jonathan Ho.
Model checkpoints to follow soon.
Make sure to use a TPU when running this notebook, enabled via Runtime -> Change runtime type -> Hardware accelerator
<a href="https://colab.research.google.com/github/google-research/google_research/diffusion_distillation/blob/master/diffusion_distillation.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
**abstract**:
Diffusion models have recently shown great promise for generative modeling, outperforming GANs on perceptual quality and autoregressive models at density estimation. A remaining downside is their slow sampling time: generating high quality samples takes many hundreds or thousands of model evaluations. Here we make two contributions to help eliminate this downside: First, we present new parameterizations of diffusion models that provide increased stability when using few sampling steps. Second, we present a method to distill a trained deterministic diffusion sampler, using many steps, into a new diffusion model that takes half as many sampling steps. We then keep progressively applying this distillation procedure to our model, halving the number of required sampling steps each time. On standard image generation benchmarks like CIFAR-10, ImageNet, and LSUN, we start out with state-of-the-art samplers taking as many as 8192 steps, and are able to distill down to models taking as few as 4 steps without losing much perceptual quality; achieving, for example, a FID of 3.0 on CIFAR-10 in 4 steps. Finally, we show that the full progressive distillation procedure does not take more time than it takes to train the original model, thus representing an efficient solution for generative modeling using diffusion at both train and test time.
This notebook is intended as an easy way to get started with the Progressive Distillation algorithm. Reproducing the results from the paper exactly can be done using the hyperparameters in the provided config files, but this requires running at a larger scale and for longer than is practical in a notebook. We hope to be able to release the checkpoints for the trained model at a later time.

```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# Download the diffusion_distillation repository
!apt-get -qq install subversion
!svn checkout https://github.com/google-research/google-research/trunk/diffusion_distillation
!pip install -r diffusion_distillation/requirements.txt --quiet
import os
import requests
import functools
import jax
from jax.config import config
import jax.numpy as jnp
import flax
from matplotlib import pyplot as plt
import numpy as onp
import tensorflow.compat.v2 as tf
tf.enable_v2_behavior()
from diffusion_distillation import diffusion_distillation
# configure JAX to use the TPU
if 'TPU_DRIVER_MODE' not in globals():
url = 'http://' + os.environ['COLAB_TPU_ADDR'].split(':')[0] + ':8475/requestversion/tpu_driver_nightly'
resp = requests.post(url)
TPU_DRIVER_MODE = 1
config.FLAGS.jax_xla_backend = "tpu_driver"
config.FLAGS.jax_backend_target = "grpc://" + os.environ['COLAB_TPU_ADDR']
print(config.FLAGS.jax_backend_target)
```
## Train a new diffusion model
```
# create model
config = diffusion_distillation.config.cifar_base.get_config()
model = diffusion_distillation.model.Model(config)
# init params
state = jax.device_get(model.make_init_state())
state = flax.jax_utils.replicate(state)
# JIT compile training step
train_step = functools.partial(model.step_fn, jax.random.PRNGKey(0), True)
train_step = functools.partial(jax.lax.scan, train_step) # for substeps
train_step = jax.pmap(train_step, axis_name='batch', donate_argnums=(0,))
# build input pipeline
total_bs = config.train.batch_size
device_bs = total_bs // jax.device_count()
train_ds = model.dataset.get_shuffled_repeated_dataset(
split='train',
batch_shape=(
jax.local_device_count(), # for pmap
config.train.substeps, # for lax.scan over multiple substeps
device_bs, # batch size per device
),
local_rng=jax.random.PRNGKey(0),
augment=True)
train_iter = diffusion_distillation.utils.numpy_iter(train_ds)
# run training
for step in range(10):
batch = next(train_iter)
state, metrics = train_step(state, batch)
metrics = jax.device_get(flax.jax_utils.unreplicate(metrics))
metrics = jax.tree_map(lambda x: float(x.mean(axis=0)), metrics)
print(metrics)
```
## Distill a trained diffusion model
```
# create model
config = diffusion_distillation.config.cifar_distill.get_config()
model = diffusion_distillation.model.Model(config)
# load the teacher params: todo
# model.load_teacher_state(config.distillation.teacher_checkpoint_path)
# init student state
init_params = diffusion_distillation.utils.copy_pytree(model.teacher_state.ema_params)
optim = model.make_optimizer_def().create(init_params)
state = diffusion_distillation.model.TrainState(
step=model.teacher_state.step,
optimizer=optim,
ema_params=diffusion_distillation.utils.copy_pytree(init_params),
num_sample_steps=model.teacher_state.num_sample_steps//2)
# build input pipeline
total_bs = config.train.batch_size
device_bs = total_bs // jax.device_count()
train_ds = model.dataset.get_shuffled_repeated_dataset(
split='train',
batch_shape=(
jax.local_device_count(), # for pmap
config.train.substeps, # for lax.scan over multiple substeps
device_bs, # batch size per device
),
local_rng=jax.random.PRNGKey(0),
augment=True)
train_iter = diffusion_distillation.utils.numpy_iter(train_ds)
steps_per_distill_iter = 10 # number of distillation steps per iteration of progressive distillation
end_num_steps = 4 # eventual number of sampling steps we want to use
while state.num_sample_steps >= end_num_steps:
# compile training step
train_step = functools.partial(model.step_fn, jax.random.PRNGKey(0), True)
train_step = functools.partial(jax.lax.scan, train_step) # for substeps
train_step = jax.pmap(train_step, axis_name='batch', donate_argnums=(0,))
# train the student against the teacher model
print('distilling teacher using %d sampling steps into student using %d steps'
% (model.teacher_state.num_sample_steps, state.num_sample_steps))
state = flax.jax_utils.replicate(state)
for step in range(steps_per_distill_iter):
batch = next(train_iter)
state, metrics = train_step(state, batch)
metrics = jax.device_get(flax.jax_utils.unreplicate(metrics))
metrics = jax.tree_map(lambda x: float(x.mean(axis=0)), metrics)
print(metrics)
# student becomes new teacher for next distillation iteration
model.teacher_state = jax.device_get(
flax.jax_utils.unreplicate(state).replace(optimizer=None))
# reset student optimizer for next distillation iteration
init_params = diffusion_distillation.utils.copy_pytree(model.teacher_state.ema_params)
optim = model.make_optimizer_def().create(init_params)
state = diffusion_distillation.model.TrainState(
step=model.teacher_state.step,
optimizer=optim,
ema_params=diffusion_distillation.utils.copy_pytree(init_params),
num_sample_steps=model.teacher_state.num_sample_steps//2)
```
## Load a distilled model checkpoint and sample from it
```
# list all available distilled checkpoints
# TODO: use cloud bucket in public version
# create imagenet model
config = diffusion_distillation.config.imagenet64_base.get_config()
model = diffusion_distillation.model.Model(config)
# load distilled checkpoint for 8 sampling steps
loaded_params = diffusion_distillation.checkpoints.restore_from_path('/todo/imagenet_8', target=None)['ema_params']
# fix possible flax version errors
ema_params = jax.device_get(model.make_init_state()).ema_params
loaded_params = flax.core.unfreeze(loaded_params)
loaded_params = jax.tree_map(
lambda x, y: onp.reshape(x, y.shape) if hasattr(y, 'shape') else x,
loaded_params,
flax.core.unfreeze(ema_params))
loaded_params = flax.core.freeze(loaded_params)
del ema_params
# sample from the model
imagenet_classes = {'malamute': 249, 'siamese': 284, 'great_white': 2,
'speedboat': 814, 'reef': 973, 'sports_car': 817,
'race_car': 751, 'model_t': 661, 'truck': 867}
labels = imagenet_classes['sports_car'] * jnp.ones((16,), dtype=jnp.int32)
samples = jax.device_get(model.samples_fn(rng=jax.random.PRNGKey(0), labels=labels, params=loaded_params, num_steps=8)).astype(onp.uint8)
# visualize samples
padded_samples = onp.pad(samples, ((0,0), (1,1), (1,1), (0,0)), mode='constant', constant_values=255)
nrows = int(onp.sqrt(padded_samples.shape[0]))
ncols = padded_samples.shape[0]//nrows
_, height, width, channels = padded_samples.shape
img_grid = padded_samples.reshape(nrows, ncols, height, width, channels).swapaxes(1,2).reshape(height*nrows, width*ncols, channels)
img = plt.imshow(img_grid)
plt.axis('off')
```
| github_jupyter |
```
import tweepy
import json
import pandas as pd
import csv
import mysql.connector
from mysql.connector import Error
#imports for catching the errors
from ssl import SSLError
from requests.exceptions import Timeout, ConnectionError
from urllib3.exceptions import ReadTimeoutError
#Twitter API credentials
consumer_key = "NDhGN1poxOV4el21shhNpFbbf"
consumer_secret = "xk8ZtyEh6Hpq2zucqvcrqSDXm7gTredBC0T8S6T9mSCPZJhEmx"
access_token = "825394860190470144-VmRgBQeYWF0MtoBYCrT4IA5ANzmDMwG"
access_token_secret = "gkHs9Beab9lPsJA9bMCwCUITRqsTLkGIwdYl6xlav0jIu"
#boundingbox ciudad de Madrid obtenido de https://boundingbox.klokantech.com/
campus = [-3.7395722584,40.4342310784,-3.7156005182,40.4612470501]
def connect(user_id, user_name, user_loc, user_follow_count,user_friends_count, user_fav_count,user_status_count,
tweet_id,text,created_at,source,
reply_id, reply_user_id,
retweet_id,retweet_user_id,
quote_id,quote_user_id,
reply_count,retweet_count,favorite_count,quote_count,
hashtags, mention_ids,
place_id, place_name, coord):
con = mysql.connector.connect(host = 'localhost',
database='twitterdb', user='david', password = 'password', charset = 'utf8mb4',auth_plugin='mysql_native_password')
cursor = con.cursor()
try:
if con.is_connected():
query = "INSERT INTO UsersCam (user_id, tweet_id,user_name, user_loc, user_follow_count,user_friends_count, user_fav_count,user_status_count) VALUES (%s,%s, %s, %s, %s, %s, %s, %s)"
cursor.execute(query, (user_id, tweet_id, user_name, user_loc, user_follow_count,user_friends_count, user_fav_count,user_status_count))
query2 = "INSERT INTO PostsCam (tweet_id,user_id,text,created_at,source,reply_id, reply_user_id,retweet_id, retweet_user_id,quote_id,quote_user_id,reply_count,retweet_count,favorite_count,quote_count,place_id, place_name, coord,hashtags, mention_ids) VALUES (%s,%s, %s, %s, %s, %s, %s, %s,%s, %s, %s, %s, %s, %s, %s, %s,%s, %s, %s, %s)"
cursor.execute(query2, (tweet_id,user_id,text,created_at,source,
reply_id, reply_user_id,
retweet_id, retweet_user_id,
quote_id,quote_user_id,
reply_count,retweet_count,favorite_count,quote_count,
place_id, place_name, coord,
hashtags, mention_ids))
con.commit()
except Error as e:
print(e)
print(text)
#Carlota: He dejado este print, porque no era capaz de almacenar emojis por la codificacion.
#Estoy casi segura de que se ha arreglado, pero por si acaso
cursor.close()
con.close()
return
class MyStreamListener(tweepy.StreamListener):
def on_data(self,data):
# Twitter returns data in JSON format - we need to decode it first
try:
decoded = json.loads(data)
except Exception as e:
print ("Error on_data: %s" % str(e)) #we don't want the listener to stop
return True
#LOCATION METADATA
#En caso de estar geolocalizado guardar la geolocalizacion
#Si esta geolocalizado dentro de un bounding box (no exacta)
if decoded.get('place') is not None:
place_id = decoded.get('place').get('id')
place_name =decoded.get('place').get('name')
else:
place_id = 'None'
place_name = 'None'
#Si es localizacion exacta
#Geo is deprecated, they suggest to use simply coordinates
if decoded.get('cordinates') is not None:
m_coord = decoded.get('coordinates')
c=0
coord=''
for i in range(0, len(m_coord)-1):
mc=m_coord[i]
m_coord=coord+mc+';'#use a different separator!
c=c+1
mc=m_coord[c]
m_coord=coord+mc
else:
coord = 'None'
#USER METADATA
user_name = '@' + decoded.get('user').get('screen_name') #nombre cuenta @itdUPM
user_id=decoded.get('user').get('id') #id de la cuenta (int)
user_loc=decoded.get('user').get('location')
user_follow_count=decoded.get('user').get('followers_count')
user_friends_count=decoded.get('user').get('friends_count')
user_fav_count=decoded.get('user').get('favourites_count')
user_status_count=decoded.get('user').get('statuses_count')
#POST METADATA
created_at = decoded.get('created_at') #Fecha
text = decoded['text'].replace('\n',' ') #Contenido tweet
tweet_id = decoded['id'] #tweet id (int64)
source = decoded['source'] #string source (web client, android, iphone) interesante???
#REPLY METADATA
reply_id=decoded['in_reply_to_status_id']
reply_user_id=decoded['in_reply_to_user_id']
#RETWEET
if decoded.get('retweeted_status') is not None:
retweet_id = decoded['retweeted_status'] ['id']
retweet_user_id = decoded['retweeted_status']['user']['id']
#Carlota: Si es un retweet los campos de nº de retweets favs etc vienen dentro de retweeted status
#David: ok bien visto, he añadido el id de usuario retweeteado
reply_count = decoded['retweeted_status']['reply_count'] #Number of times this Tweet has been replied to
retweet_count = decoded['retweeted_status']['retweet_count'] #Number of times this Tweet has been retweeted
favorite_count = decoded['retweeted_status']['favorite_count'] #how many times this Tweet has been liked by Twitter users.
quote_count = decoded['retweeted_status']['quote_count']
#hashtags_list=decoded.get('retweeted_status').get('entities').get('hashtags')
#mentions=decoded.get('retweeted_status').get('entities').get('user_mentions')
#David: para esto hay que crear una cadena de texto recorriendo la lista, el
#código estaba en la versión anterior...
hashtags_list=decoded['retweeted_status']['entities']['hashtags']
mentions=decoded['retweeted_status']['entities']['user_mentions']
hashtags=''
c=0
if len(hashtags_list)>0:
for i in range(0, len(hashtags_list)-1):
mh=hashtags_list[i].get('text')
hashtags=hashtags+mh+';'
c=c+1
mh=hashtags_list[c].get('text')
hashtags=hashtags+str(mh)
else:
hashtags='None'
mention_ids=''
c=0
if len(mentions)>0:
for i in range(0, len(mentions)-1):
mid=mentions[i].get('id_str')
mention_ids=mention_ids+mid+';'#use a different separator!
c=c+1
mid=mentions[c].get('id_str')
mention_ids=mention_ids+str(mid)
else:
mention_ids='None'
#David: esto no sé si haría falta... este justo es un retweet de un post que a su ves
#es un quote de una noticia, osea que hay dos pasos de conexión, pero el retweet
#con el quote ya existe... lo guardamos pero hay que tenerlo en cuenta que es redundante
#Carlota: Lo quito, porque tienes razon y no habia caido...
#David. lo podemos dejar porque no son campos adicionales
if decoded['retweeted_status']['is_quote_status']:
if 'quoted_status' not in decoded['retweeted_status']:
quote_id='None'
quote_user_id='None'
else:
quote_id=decoded['retweeted_status']['quoted_status']['id']
quote_user_id=decoded['retweeted_status']['quoted_status']['user']['id']
else:
quote_id='None'
quote_user_id='None'
else:
reply_count = decoded['reply_count'] #Number of times this Tweet has been replied to
retweet_count = decoded['retweet_count'] #Number of times this Tweet has been retweeted
favorite_count = decoded['favorite_count'] #how many times this Tweet has been liked by Twitter users.
quote_count = decoded['quote_count']
retweet_id = 'None'
retweet_user_id = 'None'
if decoded['is_quote_status']:
if 'quoted_status' not in decoded:
quote_id='None'
quote_user_id='None'
else:
quote_id=decoded['quoted_status']['id']
quote_user_id=decoded['quoted_status']['user']['id']
else:
quote_id='None'
quote_user_id='None'
hashtags_list=decoded.get('entities').get('hashtags')
mentions=decoded.get('entities').get('user_mentions')
hashtags=''
c=0
if len(hashtags_list)>0:
for i in range(0, len(hashtags_list)-1):
mh=hashtags_list[i].get('text')
hashtags=hashtags+mh+';'
c=c+1
mh=hashtags_list[c].get('text')
hashtags=hashtags+str(mh)
else:
hashtags='None'
mention_ids=''
c=0
if len(mentions)>0:
for i in range(0, len(mentions)-1):
mid=mentions[i].get('id_str')
mention_ids=mention_ids+mid+';'#use a different separator!
c=c+1
mid=mentions[c].get('id_str')
mention_ids=mention_ids+str(mid)
else:
mention_ids='None'
#insert data just collected into MySQL database
connect(user_id, user_name, user_loc, user_follow_count,user_friends_count, user_fav_count,user_status_count,
tweet_id,text,created_at,source,
reply_id, reply_user_id,
retweet_id,retweet_user_id,
quote_id,quote_user_id,
reply_count,retweet_count,favorite_count,quote_count,
hashtags, mention_ids,
place_id, place_name, coord)
#print("Tweet colleted at: {} ".format(str(created_at)))
def on_error(self, status_code):
if status_code == 420:
#returning False in on_error disconnects the stream
return False
# returning non-False reconnects the stream, with backoff.
while True:
if __name__ == '__main__':
try:
print ('Starting')
#authorize twitter, initialize tweepy
auth = tweepy.OAuthHandler(consumer_key, consumer_secret)
auth.set_access_token(access_token, access_token_secret)
api = tweepy.API(auth, wait_on_rate_limit=True)
#create the api and the stream object
myStreamListener = MyStreamListener()
myStream = tweepy.Stream(auth = api.auth, listener=myStreamListener)
#Filter the stream by the location of the campus
myStream.filter(locations = campus)
except (Timeout, SSLError, ReadTimeoutError, ConnectionError) as e:
#logging.warning("Network error occurred. Keep calm and carry on.", str(e))
print("Network error occurred. Keep calm and carry on.")
print(str(e))
continue
except Exception as e:
#logging.error("Unexpected error!", e)
print("Unexpected error!")
print(str(e))
continue
```
| github_jupyter |
```
% matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
from scipy.sparse import coo_matrix, eye
from scipy.sparse.linalg import svds
from sklearn.metrics import mean_squared_error
from sklearn import datasets
from sklearn.ensemble import RandomForestClassifier, ExtraTreesClassifier
from sklearn import tree
from sklearn.decomposition import PCA
from sklearn.cluster import KMeans, Birch, DBSCAN
genres = ["Action", "Adventure", "Animation", "Children", "Comedy", "Crime", "Documentary", "Drama", "Fantasy", "Film-Noir", "Horror", "Musical", "Mystery", "Romance", "Sci-Fi", "Thriller", "War", "Western"]
# 20 mln
# ratings = pd.read_csv("data/ml-20m/ratings.csv")
# movies = pd.read_csv("data/ml-20m/movies.csv")
# 1 mln
ratings = pd.read_csv("data/ml-1m/ratings.dat", sep="::", names=["userId", "movieId", "rating", "timestamp"], engine='python')
movies = pd.read_csv("data/ml-1m/movies.dat", sep="::", names=["movieId", "title", "genres"], engine='python')
```
# Macierz użyteczności
rzędy = użytkownicy,
kolumy = filmy
Może się okazać, że będzie trzeba zastosować inny rodzaj sparse_matrix
```
utility_matrix = coo_matrix((ratings['rating'], (ratings['userId'], ratings['movieId']))).asfptype()
utility_matrix.toarray(), utility_matrix.toarray().shape
# movies.head()
# ratings.head()
```
# Maicerz podobieństwa
```
%%time
A = utility_matrix @ utility_matrix.T
S = np.sqrt((A * eye(A.shape[0])).toarray() @ np.ones(A.shape))
Zero = np.logical_or(np.logical_or(S == 0, S.T == 0), np.eye(S.shape[0]))
S = S + np.logical_or(S == 0, S.T == 0)
similarity = (A.toarray() / S / S.T) * (1-Zero)
similarity.shape
def movie_values(user, n):
closest_users = np.argpartition((utility_matrix.toarray() > 0) * -similarity[:,[user]],kth=n, axis=0)[:n,]
closest_users_raitings = utility_matrix.toarray()[
closest_users,
np.arange(utility_matrix.shape[1])[None,:]
]
movie_val = np.sum(closest_users_raitings, axis=0)/n
return movie_val
#TODO: Wyzerwoać te które już ocenił i potem posortować
movie_values(user=1021,n=2)
```
# Wektory gatunków filmowych
```
for i, v in enumerate(genres):
movies[i] = movies['genres'].str.contains(v)
# tags = pd.read_csv("data/tags.csv")
movies = movies.merge(ratings.groupby('movieId')['rating'].agg([pd.np.mean]), how='left', on='movieId')
movies.head()
# ratings.head()
ratingsExt = pd.merge(ratings, movies, on='movieId')
ratingsExt.head()
for i in range(18):
ratingsExt[i] = ratingsExt[i] * (ratingsExt['rating'] - ratingsExt['mean'])
ratingsExt.head()
ratingsExt.groupby('userId')[list(range(18))].agg([pd.np.sum]).head()
data = ratingsExt.drop(["timestamp", "title", "genres"], axis = 1)
data.head()
```
## Przykłądowe dane:
```
ratingsExt.loc[(ratingsExt['userId'] == 1) & (ratingsExt['rating'] > 4)]
```
## Próba grupowania filmów według gatunków (w seumie bez sensu)
```
test = movies.drop(["movieId", 'genres'], axis = 1)
data = test.loc[:, range(18)]
print(data.shape)
pca_model = PCA(n_components=2)
pca = pca_model.fit_transform(data)
kmeans = KMeans(n_clusters=4, init='k-means++', n_init=100).fit(data)
birch = Birch(threshold=0.2, n_clusters=None).fit(data)
dbscan = DBSCAN(eps=0.999999, min_samples=25).fit(data)
x, y = pca[:,0], pca[:,1]
plt.axis('equal')
plt.scatter(x,y, c=kmeans.labels_)
plt.show()
plt.axis('equal')
plt.scatter(x,y, c=birch.labels_)
plt.show()data:image/png;base64,iVBORw0KGgoAAAANSUhEUgAAAX8AAAD8CAYAAACfF6SlAAAABHNCSVQICAgIfAhkiAAAAAlwSFlzAAALEgAACxIB0t1+/AAAADl0RVh0U29mdHdhcmUAbWF0cGxvdGxpYiB2ZXJzaW9uIDIuMi4zLCBodHRwOi8vbWF0cGxvdGxpYi5vcmcvIxREBQAAIABJREFUeJzs3Xd4FMUbwPHv7PVLgYRQQgm9NykKCIIFlK6AIhawFwSxYgMBQcXyU7AjClIUewMrKAoWQKpK772EEtKv7vz+uBAScpdcLndJkPk8Tx6SvdnZCUne3ZvyjpBSoiiKopxbtLJugKIoilL6VPBXFEU5B6ngryiKcg5SwV9RFOUcpIK/oijKOUgFf0VRlHOQCv6KoijnIBX8FUVRzkEq+CuKopyDjGXdgEASEhJknTp1yroZiqIoZ5XVq1cfk1JWLqpcuQ3+derUYdWqVWXdDEVRlLOKEGJPMOVUt4+iKMo5SAV/RVGUc1BYgr8QYqYQIlkIsT7A6xcLIVKFEOtyPsaF47qKoihKaMLV5z8LeB2YU0iZ36SUfcN0PUVRFKUEwvLkL6VcCpwIR12Kcq7wer0c3p1MZmpmWTdFOQeV5myfTkKIv4GDwMNSyg2leG1FKVeWfLqM10a8iyPLie710rFvO0a/NwJbtK2sm6acI0prwHcNUFtK2Rp4DfjKXyEhxJ1CiFVCiFVHjx4tpaYpSunauGwLL978OqnH0nBmOXE7PSz/ZjVPD5lS1k1TziGlEvyllGlSyoycz78DTEKIBD/lpksp20sp21euXOQaBUU5K338wte4HK58x9xOD+sWr+fYgeNl1CrlXFMqwV8IUU0IIXI+vyDnuuq3XDknHdpxBH9bZxvNRo4dUENnSukIS5+/EOJD4GIgQQixHxgPmACklNOAq4HhQggPkA0MkWrneOU/Std11i1ez7+/bSKuakUuHnIhsfExua+37NaMfVsO4HF7853ndXtJalqztJurnKPCEvyllNcV8frr+KaCKsp/mtvl5olez7J55XYcGQ4sdjPvPv4+z/0wlmadGgNQu1ktvB4933kWu4VrH70Se4wa8FVKh1rhqyhh9O30n9i0YiuODAcAziwX2ekOJl7zErqus+Lb1UwfPZu8b3yFEHTq344bx15dVs1WzkEq+CtKiJL3HmXyja8wqPKtDK03gs+nLGDhrF9wZrkKlM1My2LPhn28PXpugdellPz60Z/0jbqB6Y/MLa3mK+e4cpvVU1HKs5NHUxne/lEyUjLRvTppx9N578mPsNgtfss7s1zs/Gcv+zYfCFiny+Hm0//NJ3nvUcZ+9GCkmq4ogHryV5SQzH9rIY4MB7r3dN+9M8tFRkommrHgn5XUJdMenh1U3Us+WYYjyxG2tiqKPyr4K0oI1v+2CZfDXeC4NcqMweD/z+rkkdSg69+2ZlfIbVOUYKjgryghqNWkOgajocBxr0fHGmUtcf01GyWWuA5FKYwK/ooSggGj+mA05x8yM5qNNGhTjw592qL5efqPiY/GGuV/TCCvpKY1iatSMWxtVRR/VPBXlBDUbJjIs989QfX61TCajRjNRjr2bccz3zzGLU9fR3RcFCaLCQDNoGGxWxjz0QMkNqpSaL0xlaJ5bcWzpfEtKOc4NdtHUUJUuVYlBj7Qh9rNk9i7cR/rFq9nzlOf0veuHszYMIX5b/7IP0s2UrNRIv2GX8HdbUYXWWfGiUxSk9Owq+yeSoSp4K8oxeRyuRhadyQnDqUUeM1gNLDgrYXUaVELt9NN2+6tiI2PDirwg2/O/+9frOCah/uHu9mKko8K/opSTMPbPOo38AN4PV68Hi/bVu8EYPf6fcWuX9dV2isl8lSfv6IU095N+yNWt8Go0XnABRGrX1FOUcFfUcoJzaAxbMJgajZU0zyVyFPdPopSTEITyDB3zfS9uwcDRvUhqUmNsNarKIGoJ39FKaYBo3qHvc46zWupwK+UKhX8FaWY+o+4Iux1dujTLux1KkphVPBXlGKqXKNSsc+p3bwWFrvZ72v2WBvV6hS++EtRwk0Ff0UJQmZaFscOnkBKidlq5rxLmhfr/D0b9vnN8w8gkYztN1lt3q6UKlFet9Jt3769XLVqVVk3QznHZZzM5IWbX2flD+vwuDwAJDWtwdt//49e5kJ3Ly02TRO8sfoFGrSuE9Z6lXOLEGK1lLJ9UeXUk7+iFGJsv+dYNn9VbuAH2LvpQNgDP/gWd93b4XG+e+cnvF5v0ScoSgmo4K8oAezZtJ/ta3aW6jU9Lg9v3v8ezw19rVSvq5x7VPBXlACS9x4rkLa5NDizXSz7eiW7NxQ/NYSiBEsFf0UJoH7r2n536yoVAjb8sblsrq2cE1TwV5QA4qvF0fO2S8vk2prBQHxiXJlcWzk3qOCvKIUY+eqtVKgSW6rXFEJgtVs4v+d5pXpd5dyigr+iFELTND47PIMLerWO+LWsURYsNjNJzWry8pKnMJpU6i0lctRvl6IEIKVk4exf+eDpzwPm7w+XB6bfTfMLG2G2mkmsVzWi11IUUMFfKSMet4cF0xby3Ts/oXt1egztxoD7emOxFb3BeWn5fOq3zHryI5xZzoheR2jQ+/bLInoNRTmTCv5KqZNSMv6qF/h7ycbcwDp30mf88dVfTP3jaQwGQxm30HdzmvvUJxEP/ABSB4/Hg9Go/hyV0hOWPn8hxEwhRLIQYn2A14UQ4lUhxHYhxD9CiLbhuK5ydtq0Yhv/LN2YL7C6sl3s2bifv75bW4YtO+3k0TS87sisspWAu6EFaRQRqV9RghGuAd9ZQM9CXu8FNMz5uBN4K0zXVc5Cm5Zt9RtYszMc5WZue2ylGBCRC87SJEmfUwdvLRNA0E/9bt2DV+oRa5dy7ghL8JdSLgVOFFLkSmCO9FkOVBRCqL3qzlHxiXF+V85abGYSQkiXHAlmi4kBo3phsUdmDEJPNCNjDWSNqU6tp1rz2pYfSHNlBSy/I/0Ity2fxkULx3PRwvGMWfchae7siLRNOTeUVidjDSDvWvX9OccOldL1lXLkwivb8/q9JhyZDvImlTUYDVx6fZeya9gZbnn6OsxWE5+99A1Z6eELtAIwr8vGKUBPNLI+KYv1u5Yyd9dSxrUcRN8a7fj+wFqe3zifLK8To9AAgVd6kYAuvfx6ZCN7s44zp9MIRATfoSj/XaU1z9/fb2eBXNJCiDuFEKuEEKuOHj1aCs1SyoLFZuHlJU9Rq0kNLDYzFruFanUq8/yiJ33dLeWEpmkMHTeYL1NmcddLw8L6LkBk6IgjHjij33/Sv5/z2Z4VjP/3U7K8vjERj9Tx5AT+U9zSy97MY6xPVfl/lNCU1pP/fqBWnq9rAgfPLCSlnA5MB18+/9JpmhJOXq8XIQSaVvhzRe1mtZixYSqHdh3B69Gp0aBauX2C1TSNQff3JTM1m0//Nx9NE2RnOEpWqQ7YRIFxBQm8tGl+UFUIYG/mMVpWTCpZW5RzUmk9+c8HhuXM+ukIpEopVZfPf8jBHYcZ3f0peluvp7f1eiYOfonUY2kBy7vdbtxuN4l1q1KzYWK5DfynCCG4amRPegzrhtlqQjOE3l5pAE8TC7KC/2cvb8E3xX7pSBrGVAu5Hcq5LSxP/kKID4GLgQQhxH5gPGACkFJOA74DegPbgSzglnBcVykfMtOyuLfTE6QdT8/tzPvjy7/YvX4f765/Od+7gL1bDnB7iweQ3tMBrtvgToz96MHSbnaxZGc6uOf8Rzl+4AReT2izbU59xzJKUOWBpuwmI+T2mDUjLSrUolFs9ZDrUM5tYQn+UspCtzWSvr0iR4TjWkr5sW/LAcb0mcyhnUcKvKZ7dfZvOcCan/6h/eWnE5Td1vT+AmWXfLKMui0/54YxgyLa3uLoF3MjjszT6xA69G1L2rH0kAM/nB74EmmSOTc+TN9fnyc9OR3L3OOY/spE2jSa3tgSeVVlVp0suImMhsBsMGLWjPSr0Y67GvYIuS2KopYUKiHJysjmjpYP4fUEXgglJcx/88fc4P/ekx8GLDtnwiflJvj30K4pcGzFN2vCeo1dyQe40FqfP0ctgFQPaW1MMKYGK2Uq9r2ZxMbbSPOcnmFk0Yx8ftFDVLFVCGs7lHOXCv5KSGaP+6TQwH/KpuVbcz9ftmB1wHK6t3wsXPr81W8jfg0J3PzHFIy/eRDXVMTVv+LpF4Ugy6qDO5t+NdqSFJ1Aqwq1qRNTmU/2LGP1iV3Uslfi+jqdqa/6+5USUMFfCcnWVduDKpc3qHe+qj27/tkTsKzb5cZkNpW4bSUx7+kvIlp/br9/TDTuS3WwFJzxA4CABQfXIIBK5hg80kumx4lbevknZQ+LDv3D821uoFPlRhFtr/LfpfL5KyGp07xW0YWAtpe1zP38pglDApYzmg38/sVfJW5XSdVsFNl0yrlhXkowBwj8eUjgmCudk+4s3NL3TktH4tDdPLP+S6RUM6KV0Kjgr4TklmcKHePPNertu/J9fdtzN/gt53F5WbPo7xK3q6Re+f3ZiF9DWnICvlay6a2p7kySnYGn0ypKYVTwV0ISGx9DXNXAg49CE4x683ZiKkTlO163eS3sMbYC5U1mIwm1ykden2tG9y/R+fKMj3yvmQXuC6PCkjRORxJlKD/7HyhnFxX8lZBVqhEf8LWeN19Cv7uvQErJ2sX/8urId3l79Bziq8VhsVsKLOoyGA30urVsNks/053PD2WR/mlI53rsIDVwtrPhHFABPd6ANIAUoFcxkj08Acfokuc0NAkDHSs1JNpkLXFdyrlJDfgqIbt54hDG9p3s97Xl365GSslzw17jz6/+8s2ZF/D1Gz9w9QN9+OOrlRzZcxRN07DYzTz+/n1USapcyt+Bf8cPpfD9uz8V+zwJGHISc1pWZ8PqbETOcczguKYi7ityZvac9KAdcqM3sPr+CoN8J2A3mPFKSZPY6oxvVXBKqqIES5TXAaP27dvLVatWlXUzFD9+nL0Yk8XMpUO6MLTBCA7vTPZbzmA0BJwO2uv2y2h9cXPqtkiidvOa5WL3LoAtq3Yw+tIJIeXukfjPYJj39bR3ayGiTESP2INI8yJjDKTPqQtBpItoHluLuxt1p6q1AnWiqxS7fcq5QQixWkrZvshyKvgrwbq30xNsXrEt7PW+tGQirS5qGvZ6Q3F7ywfZsyEymTJP/aU5B1bAsiAVvOBpYyfr8apgMwT99J9gjmZmp+G8vOk73NLDPQ0vp2Gs2h5D8VHBXwmrxR/9zuTrX4lY/V+lzCLqjMHh0pZ2PJ1ra9yJx+WJSP0SkDaQcSYyroyB3vH5R92E8E0BDWEw+LyKtZne8a6iCyr/ecEGfzXgqwRl8g2RC/wAAyrdwvczfo7oNYrib3exYBX1CCWN4GltI/2zBmS8Uxv65gR+IU5/QMizgNad3MOC/ephSQmeCv5KcCL8BlHqkjdGzWTX+r2RvVAh7DE22lzSAoMptPGHAtM68U3tlGaBt4mV7CcS8wf7MKexfmd72d48lbOLCv5KUDRj5H9V3C4PP8xcHPHrFGb07JHUaFANW7QVk7VkqSYcN8aR+XR10t+qTeYLtZAxkR3Udnoj012l/DepqZ5KUN7550Vua/ZQRK+he3UyT2ZG9BpFiatSgXfXT2H975s5vDuZP79eye9frCj0nEBvijydY9Brl94irO6JLYsupCg51JO/EpSkJklc/2RkUy5boy1ceNUFEb1GMIQQtLyoKT2GdmPQg30Clsu7gjftYjvuC+z5bgSGjQ7wls6ECqtm4sHGgduqKGdSwV8pkjPbyZevfcfqH9YVOiha2IrfophtZlp2aUqHPm1DriMSnhr0UlDlon/NIntAFq6eMb4VvYB5QUrhE/9LyCQM2Axmeia25pfLxoVlrcSB7Yd4beS7PHTJeN597H2OH0oJQ0uV8kh1+yiFcjlc3Nd5LPu3HsSZ5Sq0bIN2tUk/kYEru/By/ggBT3z0QLlZ7HXKySOpAV/LG9c1IPZxSLOm41jQ0HcwxGmbwehetQXPtrk+qLJSSjYt38rh3Udp2LYutRrX8Ftu9oSPeX/SZ7lvZzb8sZlvpi/i9RXPUbOhWkfwX6OCv1KoxR/+wYFth4oM/AAr5q8N+TqaprHs65X0GNot5DoiQgOC2GfmVBqHWAfky7MZgRuAVTPRMcg8/iePpvJI94kc3pUMQuD1eOnQpy1j5t2PwXj6Rrtg2o98MOnzfAMYXo9O5sksnr72ZXrffhndh3XDHl0wKZ9ydlLdPkqhli1YmW8v20hxu9ycTC5/6Ykbt28QdNkCIT4C0zkBHLqbJUc2BlX2hZveYO+mA2RnOMhOz8aV7eKvb9fw2csLcsvous7MMR8G3Btgx7rdvDZyBlfGDmPupM/C8j0oZU8Ff6VQ8VUrohki/2uiGQy06lo+UjzkNfq9EWXdBL9WHd9RZJms9GzWLv63QH4lZ7aLBdMW5n6dcTIz6Bv8nPEfs3llcLu4KeWbCv5KofrefTkmS+R7B9tf3prG5wf/lF1aajetSbfBFwZV1reoK7LtOSVbdzN756+Flln7c8HAf4oz63Swj4q1Y7YEv6Zh1pMfBl1WKb9U8FcKVb91He6fdifWKAv2WFvIq18LE1e1AuM+i+wagpJ4/INR2GML9nXnnep56t+sJ0pvU/WPdy8L+Np7T37I5BteQeoFu3IMRgMd+7XP9/U1D/fDYg/uzpV6NL34jVXKHRX8lSJ1v7EbnyXPYOLXjzLp60ex2ML7eDv935fL3SyfvAwGA1+fnMMFvc7Ld1zHF/R1u8DT0Ezau7XxXhBTau2SAZaXHdlzlM9eWoDTz6wrk8VExSqx3DIp/37KN4y9mhvGXk1URTuaQcMWHXiTmPKwFkMpOZXVUym2ZQtW8ez1r+DILH7O+zNFxdr46uScMLSqdLlcLvpYb/Dl6P+6PhgjM7hbmLsb9ODWBpcUOP79jJ958/73/PbjN+vciGe/HUNUrN1vnbqu43K4sdjM3NLkPg5sO5Tv9ej4aD49/A5Go5ooWF6prJ5KxHTq157Pj86gXuvaJa6r27Wdw9Ci8HC53Mx56hMevXwSr4+aSWZaVsCytzS+D/DN8LFduQOOeXzTOk99RHhlr1kz+g38APZYO8LP5vAGk4FWXZsHDPy71u9l6t1vM67/c3z28jdM//clBj3Yl9hK0URVtHPFLZfw4d63VOD/j1BP/kqxpZ1I59am95N6tGRTM+2xNr44/l656PJJ3nuUmxvfh9vpzne82+BOjP3owQLle2gFt1DUAaI1pFsi61vIfL5mUDt0FZdRGHi8+VX0q9nO7+uOLCfXVr+DrLTsfMctNjPT1r5IzUbVc4+lpmayc80u9m3Zzxv3vofuPb2owRZtZcbGqVSuWSns34MSOcE++atbuBJQ6ol0nr/xVTat2EpsfCyj3rydLSu3M3v8x+hheLJ9aMZwNK18vPkc2//5AoEfYMkny9i6cgRzdryRe+zInqN+69AAMnzB0+OVaJuy0Vv4f8oOldVgok5UZa6o3trv61JKrHYLz3z7BE/2fy43mHvdXu5/+87cwJ+VlcXVlW7D7QycCTQ7w8GjPSYyc1Nk93JQyoYK/ko+Ho+HOeM+4Yf3fiHlyMnc4xkpWTx2xdNhvdbT106hcs1KjP34QZp2aBjWuotr9797Ar52aFcyf/24jguu8A34/jjrl4BlvVWNZE6ugUwwgYGwrfCtbU+girUCl1RrTq/E8zBrp/90XQ4X7zwylx/e+wVntosm5zeg+9CuPP7+KKSErat38PP7vzHv2S+Jq1qRdj1aM6jSbXgKCfyn7N92iOOHUqiUGFfi70EpX8IS/IUQPYFX8P26vyulfO6M128GXgQO5Bx6XUr5bjiurYSPrusMrTeSY/uPR6B2SWy8B6kL0k/6fu2kLknee4xHL5/I3B2vEmP9CLI+AJkF5i6ImNEIY60ItMVf6wr3yfNf5Qb/E2ckOzu1cbu3jpmMV2v5unrCPPi7J+sY8ZZoXt70Df/buIDz4urwRIsBJEUlMPGal1j787+4HL53LptWbGPTim0YjAakrqPnme752BVPU6NRYlCB/5SstCwV/P+DSvyeWwhhAN4AegHNgOuEEM38FP1YSnlezocK/OXQV699H6HADyBwuzQ+WLORK2/L322ie3QWv/cIZLwFejLIDHAuRB4fhNRPhL0lacfT+ei5L5kw8EVmT/iY44dSqNOs8JtMdJxvf+FH13zAh712kzqvLo5+scic4QoJZD5SLSKB/5S1KbvxSB0dydqU3dy6/C22bdrD2sXrcwN/Xl6PN1/gP+XA1kMFjgViMBqo3qD01i4opSccT/4XANullDsBhBAfAVcCwSUfUcqNxfN+i2Dtktg4Dxar5NbHD7F5dRRb1vn6w53ZLo7tPQjknTqqg8xGZn6AiLk3bK04tOsIIy94HEeW05fn5vu1fDH1W8Z+9ABj+z2Xb8Azr7SRlejwwxO+dwhmwGLEeXcVnHdXQVuTif1/R5A1TKU23VMicXk9fL38d0xmA67sos8JRWK9quViQF4Jv3CMttUA9uX5en/OsTMNEkL8I4T4TAhROu/llWLxt4o1nG565DAAJouk99BjucdtUV5adCz45ApOcIeeKdSfaQ/OJiPldNppt9NNVlo2c576lAXpc4mvXrB7wzmqGr87t5/uGsq72boQ6G2jyJhbJyIzewrj0N0sTNxJRquSbTdZmDaXtohY3UrZCkfw9/cbf+Z7zQVAHSllK+AnYLbfioS4UwixSgix6uhR/zMqlMgZOn5wmGuUuR9X3XaUywb5BpANBoiu4HvCNlt1ajd20P7iY37ON4ExvPl+Vi/6229XyNaV2xGa4OP903nnn5e4aGAH6p9XG72BFbEyHdL93ZxyCAEGLaIbtwRykmwyH66K66oKxTrv+icGULV25ULLaAaNW58Nbs8A5ewTjuC/H8j7JF8TOJi3gJTyuJTy1HLDdwC/E5SllNOllO2llO0rVy78F1MJv5ZdmnLVqF5BlR30UL8iy1ijPYx4Zj/DRh/CFq2zcZUdKSE7U7B+hZ3ajbMZ+tBhXvj0VIbKM9JGCBPCPrSY30XhAqWm0IyG3OylNRon8sf8lWxftwex3YFpWSZRYw6ibXP4Zu+UM14TuG+tWuSg9SlWu4Wh4wfzwPS7mPDFaL48MYt3N0yhcq3T8/krVI7l1WXPBFwQppz9SrzISwhhBLYCl+GbzbMSuF5KuSFPmUQp5aGczwcAj0opOxZWr1rkVXZOHE7h6zd+ZM+Gvaxb+i+ZJ0qSxkFiMEq8XoHFqnNhz1T63nScJm2yMObprXA6BFh6YRGLAB2M9RCxzyDM5wWsORTvPvY+X776Xb4BUpPZSNfBF/LYHN/Ywl1tHmbn3/mnfkoAC2TfURkZb8TTxg6WPM9OUoJDgq1s1i3YDRaaT3Oz9cstRZZ9eclTTBj4Ih63L+Onx+Vh+NRb6Hzl+Uwc/BKbV2zHaDRgi7Xx0LvD6dC7fG2tqRQu2EVeYVnhK4ToDUzFN9VzppTyGSHERGCVlHK+EGIy0B/wACeA4VLKzYXVqYJ/2RpQ6WYyUjLDXq9mkLS9KI1n5u3OPSYl/Ls8hnT9DboMbAvShdCiw35tAJfTzYSBL/LPrxswGA3ouk7dlrWZ/MPpfDf+Vu/mttUA0iwQQpA5LhFvK7vvG9jnQlQ0ImPLZnDUrBmx9t6ECGLXMbPV5Hd2kD8Wu4U3Vj5H7aY1S9hCpbSUavCPBBX8y86zN7zCLx/+HrH6LTYvX29fjxA5aXA8cMclLRn+XDMu6LYGhAmibkez9Y1YG3Zv2Mfu9Xup0TCRhm3r5XutsOCfl7QK0t6vi7bLiX30Abydo8l+qCqYREQGfwW+1A4SiUeejvJmzQhzj2D7ILhpsQaTAa/bf57/M2kGjT53dmfUG3eE0mSlDKjEbkrIln4WOE98yUncLsHB3WbSUjT++CGa27s14Lp7D9D+wo/Buw08GyH1QfTjtyHd25DujUgZxCNtMdRpXouLr+1cIPADxFWtGFQd7ua+tMd6MxsZ39THWVsj+pH9mJakY9iYDZ7wtrluVBXmdh7JY82vItpoxWYwY9IMXFi5MdYPg18PYTAG/2eve3UO7TwSSnOVck6ld1AK8LcBSDi1vjCDtb/HcP4lqXTumUHnnr5tAQtMkXf/hjz+FwgDCDtUnIowRz6X/JtrJnNdjeGFlnEMqoDz1pxJCTkN12+oQkarDCo8dhg9zkD6+3XD2q6dmcnUi65Kveiq9K7ehgPZKVQ02algttND/yroelzZwXX5AAhN0LZ7q1Caq5Rz6slfKaBR+/phrlGiGXQMRkl8NRdPf7CLPjcep0oNT+4e54HXRjl96R70Y8gTw9Az5wXcaDxcEhIT+CxlFrWaVA9YxnlL/sCf+2/LaFK/aUD6nDoRadvliyYBYNQM1I5KoILZN05x1QM9I3I9zaDR+/bLIlK3UrZUn79SgMPhoH/UsDAEWUm3K1NofkEWXreg/SXpJDUMbqPwwIxgvw4t9skS1hO8nRv2cFfLh3O/9lQ1kDmzrv871qlEbmFK6ObPXz2f9Xs82LGKYGkGwbWPXMmtz9wQ1nqVyFJ9/krIrFYr8zPncN6lLTCaDZisRgzGYGaxSKx2F5rBd9O4+bHDPP7mPq685TgD7zwWhsAP4IGsj5Hew2GoKzj1mtfm85MzsMXakBq44goJ6me+E4iAixdOwO3O33XjzHYSVSF8c/KtURZqNa7BkMcGhq1OpXxRwV/xy2q18uJP4/ne8RHfZs4joUZ8EGdJXv1+ByazL/j3uuF4ZGKgMIP77whUHNj6X7egu70IHWybPRiXZ4I7vAO6wcrSXXT9eUK+Y399vy4sYzVXjepFt2s6MeKVW3lz1fPYYyKb8kMpO2rAVylSZqYj4AYmp0nMdjexCS4mvLeLlx6ohckUqS5FHbSECNXt3zuPzM3dEF0AtqlHyJxcEz3R5DtgKd09fL1I1h7fRZtKvkHl7PTsEs2IEprgxcUTaN3VX0Je5b9IPfkrRXpr1MwgSglcWWaGNG/FpDtr8f6qTWgGPQLZEDTQKoOp+KtOpZQk7z1K6rHibz95cEf+6Y5auk70yL1EjT2AdVoyYp/BJrNaAAAgAElEQVSr1FM/fLV/Ze7nbS5ricfP1FJrtDWouqQuefji8Uy+Ue3ada5QwV8pkhb0giVfuaw0E799Z8LmS4GfGxNP7W0eegY0ExibIuLnIIr5lP33rxu4se493NL0fq6rdRcPXzqBE4dTij4xh7/NTARg3OTAsjAd01s5YxCnv8mI616tZe7nlWtW4rrHBmCxW3LfgFijLLn7EARr8bzfWTg78E5lyn+HCv5Kke6acksxSgssNp29W+LweiF5P3wzJ57De024HKDrUPS+WYGqjgJzRxCWYp12aNcRxvadTPLeY7iyXbidHtb/vonR3ScGPaNp2FOBM55KwNUjFsKwr3GwBDAxcRw9tGtyPzLTM5j8/Ri6D+3GRYM6MHzKzRzbF3hznrytlZrAnRiLBOZO+jzSzVfKARX8lSJFR9u45LouQZeXUlAh3oumQUJ1eP3xWtzUsRlTHq5FifYFkSchaw7y2ACknhH0aQveWpibxOwUr0fn6N5jbFq+Nag6et5yKRWrxRY4rlsEGS/XhEsqglEratFCWBgQRPfeVuD45y99x0/zltDnju6s+elfptz5dqH1CE4n3fZUiSatV1MyLmvIoZ1HmHzjK2Sl+3aIeXLWD7S/Zypth0+hw72vMGehmoL9X6AGfJWgPPHBfUHn+xECml+QiRCgafDl1r9xZBkwGMPxZOwGPQWZ9Qki+tagzji4/TAed8E9a4UQHN13HDoFd+VPD85gQKWbyEjJyn1qznyhBnoDa6kM9n7c5T6qWmKZ0O9/BNri5ru3f+b7dxYHPfPn1A0grXczMBlw1Y7HnVSRpZ8v58Shk6T2acaqbftzy7s9OlO//A2H282B42ksXrsdr67TuXkdHrr6YqrFx5T4+1RKhwr+StDa92rFqu//OePoqe3LfSw2Lx26p1Gv2ek00PZosEd7w9gV7gDXH0Bwwb/1Jc1ZtXAdzixXvuMet6fYq5m/PO7bh+jtbT8xY8fiYp1bEksvHofV6hu8XbvwzJ9Bfv4C/6kjAW9Rppy3ZEaN9O6NycxysW7pTo43jPF7Y5v2zfJ8X/+ybjvrdhzkywk3E20rXrecUjZUt48StMnfPpmn60Pm+4iv4qZByyzunniQx97cm3tO3rgRvodjAxiCTzF8xc2XUCEhFqPpdJ+TxW6h6zWdSKxXNaQW3FH/0pDOC9UNy18PSz1n3hakn2OYDOixVk70bBJ0vbqEzGwX367YVMIWKqVFPfkrxfLpwRkA6JlzIX0Sx48YuL5NM4bcd4j+N6fkC/CRy3BgQkQFv8OXPcbGGyuf44OnP+ePr/7CFmWl/8ie9L2rR8gt+Cd1LyYMuAkuNXJJnXRn5X7+/qHXuTFxZLHOFwIymlUhakNygWCfMqR5/oI5/xb3jZrD7WHDHpUB9GyhnvyVkGhRQ8F8EZNur4tmgDfHJJGVmX+mY/gDvxW0eETFVxDF3Nu3YuUKjHjlVubtmcaMjVO58p6eGEow+vzh7j/wlFLgBxjZ+HTitqpVqyICTL/91jEPs7Xghu4mk5HRr91J+j1dcFaPwWvUyGqUwIm7LoQK/vf/FUYDFlPwz4cGTdCgeqWiCyrlggr+il/b/95F3+gbuNw4mGENR+JwFNzK8c3xndi02o7u9f0aDWzUmj9+8L0W9qnuhtqIhK8Qlf9AWC8Jc+XFd9SRFuqE1ZAMSMqfynqh+xNeWjeuQLn+MTcya+tr1D+vjm8thIA6LZIYvegJnvl4MW6vTma/lqTe1pGO9/fCZg4c3K0WEw9ec1HQbTQbjfTv1Lzogkq5oIK/UsATfZ5heJtHcGa5kLrk0I4j9LMPZfvfu/B4PDw39FV6267n6zd+5MwhxEm3tebG9o1xZIfzBmAD+1CEsR5ChHebxLQT6Ux7aDY31BnOrc3u56vXv8frLfqJ/sLKjXw7aEVYVXNswCyerVq1LHDM69a5PuluWnRpgslqwmQxcTI5lYc+WFTgZvXr3ztwuArOggLfTKg2DWrQoUmdoJ7+7RYTcx4dQsVolQvobKH6/JUCVn6/zu/x4W0eoUbDRA5sOxTw3NYXZtBj8Am8OZkGSt7vbwStAri3oXuOoBlDG6AFSE/J4Oi+41SrWwV7jI3sTAcjzn+M4wdO4M4Jgu8+9gEb/tjMmA8fyD1PSsnG1P1sSz9MLXsl2sbXZXDtC/l6/yqOO9PzbakYLh92vo84cxTxlsB7GY++YkLA175+/Yfczw82KnyVr9lowOU5fcMTwMAuLXjk2kswGQxEW004z5gqazJoxMfasZqMNEmqQnqWi6lf/Ea/Ts3o3qYRmlZ6eY6U0Kh8/ko+b4yayVevfx/SubeNPUi/m45jteth6u+PAdLPOPSkb7yhGDxuD6+NnMGiOUswWYx4XB6admqE0AQbft+C25k/PbLZamLa2hep1bgGDq+LUatmsSl1PzoSqUuiTVZeajuMpKgEbln2Jvuzg99CMRgaAqvBhEfqdExoyFOtBhNlLDh9Mtj8/V6LgZND2xNohV33Ng1Z+u9OjAYNIQT3XtWFwd1aA9BnzDscOlFwQV3dahXp2LwOf3j/4aD1KFJK5H4r5p2xdGlan+dv71PsFBxKeKh8/kpIMtMyQzrPYvdy5S3HsEWFK/BDgcAPkD4JXc8qeLwQ7439kJ8/WIrb6SYrLRuXw83fv2xg3c/rCwR+AIPRwJaVOwDffP4NJ/fh1D24dS8edE66s7hjxdssSd5ItCm4xGnFoSPJ8rpw6R6WHd3KhH8+KVF9mtOLdeW+gK9PuPEKZj58Le0a1kR63Dz30WLaDp9C2+FT/AZ+gF2HU/jw57Xs/dWLd7cFXBrSLXBWyWTxmm38syvwu0OlfFDdPko+o6bdwaI5SwMXOLUklGy+2r4Va04Xr8tZikktM96C2IeCKiqlZMFbCwss8CqU8CVKA/jmwBrcsuAYgI7khY3zGVy7EzvSj/gtEw5u6WXZsW2kuDKIM+fvAopLrEDKodQi6xCAddtRHB1q++2D6/LQ65g08EqBHvQP8XQ9cnMU3i1RuQsGJJLbX/qElW/cH2RdSllQT/5KPlarlaSmNfy+9vCsEcTGuYFsfjjgC/ynUtmYLWAJ/0Owf9kfBZ2QzeP24MgKfgcxzaARV7UiLbs2BcCtBw7qRqFRN6oK8ZZoTGEeiD7zOimugu/IPjnwbtB1aFluYr/8Fy3D//+FW6cYgd8PKfDdEHwfXl0yd5Hqti3PVPBXCpixYSod+xfsMnx/4ic4szVm/r4L8L961+MuhXcA0gGejUEVNZlN1GhQrchyQghMFiPNOjXipV8moGm+P40uVRoHPgdBrMnGB51HcVv9S6hhi0MLOV11oa2jpt3//PlF+qdB1gDGYxnEfrMhAj8g/9/zbBX8yzUV/JUCdm/Yy/L5Bf9wD+88ijPbQPW6BfvJTxE5v1GRvQFooJ8MuvTI127DYjMHHIsw20zcOP4aPtgzjSlLJ5FQ43Sgva9xbyqY/O+NK5F0SGhArMnGrQ0u5cMu91PDHh/yu4BWFZKINlgx5PmztGom7m3cM+C00pTkVKo3qIYtxorJcnpxl7//fiFBZLkQTv/TO8NNzfgp31TwVwoYd+ULhb5eSE8Iug5Xt2xK6nEtgjcAD5haBXx186pt9DQPoYd2DZcbB3PiSAovLZlIx37tqZKUgNFsxGI3Y7IYsUZZaHNZK254YiBxVQqudK1sjeXrbqPpnNAYDYFRaFg1E1aDiRfa3IjVYM4tazWYmNXpHobU6UyitSJxAW4agbillw+63Eu/mu2oaa9Eu/i6PN/mBgYldQh4ztS73ubInqNkpztyB6/zzrLRLUa8dnOem4EIy16/wRje98JSuY4SGjXgqxRw/FDhUxenjU/knqf9z+aY8lB1Mk6YubZVS2o3ymb6r8Hlyy+W6IcQmv/Uwe+N+5B5T3+R+7XUJS8Me4PWFzfnf4snAOByuFg2fxXHD6bQvHNjGp/fgO3rdjH/jR8xWQwkNauF9EpaX9Kcui2SsBstvNRuKJM3fMW3B9bi0b1IJB/v/pN2cXXzpYmIMdnokNCAD3f9jofizf9P92STaIvjiRYDgirv9XpZ8e0avJ78d2MpJXq0mYxLGuKpGgNSomW5if51O8bk9AglXMqf3TWxUgwDuhRchKaUHyr4KwVUrpXAga2Bp+rNf68K/W85Ss36+bsPtv1j4efPKud+ffK4EV335fQPDwExY9CihgUskTfw5/X3rxtyPzdbzXQbfPqpdGz/yaz4Zk2+8gajhtFkpOvgToyeOYIZO37hx4N/48kzq+e3Y5u5cNE4Xmt/CxckNMDhcXHSncV9K9/DG0Lyh/1ZJ1h06G96JLYO7gSJ34HvU/n59QrW3P98PdZAWq+m2NbuB1vB3D8l1bJudfYlp2AyGrh/QBd6dVAbwZd3qttHKWDi148WWeb2rs255/IG7N5sweHw9fE3aOlk6COn0zlnZxj44zv/ScNCYrms0MBflN++Wl7g2NNDXi4Q+MG305cz28Vvny1nySd/Mm/37zj0gmMdEsm9q2bSdeF4uv40gf5LXggp8J8y7p9P0fXg3jEYjAZadWuGOKNv3V0tBj3GWvCua9TIPj8p5LYFYjJoTLrpCnqe35gR/TurwH+WCEvwF0L0FEJsEUJsF0I85ud1ixDi45zXVwgh6oTjukpkJDWuwaPvjyqy3M4NUdRu7MRqPT3l84b7Uvhu798AuJwaP39RMXwNc/6Entw95NOfuXYqn0/9JvfrmWPnseSTZYWe48h08t2Mn8nyBF4nIMHvjSEUXqnzx7Hgu8oenH43sZVisEb5VgAbrCa8lezgL+tnhLaYdHt1rpowi49+/ZsJcxfSdvgUlm3cHfbrKOFV4uAvfJm23gB6Ac2A64QQZ976bwNSpJQNgCnA8yW9rhJZ3a+/yM80wvxPtF/v+Ds3nuT90Aww+aON3D72ABUrhSco5tL3oh8Ofj/hvLxuL++N/ZCNy7YA8MmL84M+r4YtLqRrhsKbM6L+wa7fGLDkRfr/+jxvb/vJ7zuCxHpVmbvjdYa/fDMD7+/DHS/fRFbHuqWyrWRhRrz2ZZleXylaOPr8LwC2Syl3AgghPgKuBPJOxL4SmJDz+WfA60IIIctrYiElV94bwOOXXs6qX2M5NbBX2KKuNhe5aXPRsQi1Khnd7UYzFey7vv+du5l6x7SAZ7qy3SyYtpBG59fH6y56Va7QBD2GXUxMs+o8uHp2xNM4awhaVKjFTX+8zqb0g7nHZ+xYzLcH1vBV14dz1yCcYou20fuO0++IWuxrw3XPfhDhlhZt3i9ruf6SNmXdDCWAcHT71ADyJg7Zn3PMbxkppQdIBdSuD2eZZ+YdZNjow9iivSQk+rpB/D1gnvlOIDL8D0hv/Wt7oWdJKUk7noHRaAyqbSazke43XkTnyo35X5uhEVnClZdA0H/pC/kC/ymHHSf5cv/KIutoXKsKTwzsFonmFctGtatXuRaO4O/v7+HMB6RgyiCEuFMIsUoIsero0aNhaJoSLtfWuJPeSY2ZN7UqVWo6MFu9DGranKubN2fibbXZt720N+1O9Hv0x1m/FHqWNcpC16s7AtChX7tCy2qaRo+bumHMyWd/UdWmLLx0LJdXa0VFk514c3jTOgjAi15oiuhvD6wOqq7+3VoTH1O8dQbhNuyytmV6faVw4ej22Q/UyvN1TeDMx5ZTZfYLIYxABaDAZHIp5XRgOvhSOoehbUoYXGG6Ft2rc+pZYc/mU0HFd0//84cKrP0thjcXbSWxdjESqIUsym+XD1BoV47ZZiY+MY5Fc5eQdiKDp754hMd7Ps2an/4tUNZkNmKPtXHj2KvzHa9gtvP0eUMA8Ohehv/1LpvSDuDSS75qNphfeNMxne1rd5HUrCZmS+Apm4MmzuJEevGyn4ZTfIyNRrWqlNn1laKVOJ9/TjDfClwGHABWAtdLKTfkKTMCaCmlvFsIMQQYKKUcXFi9Kp9/+bD0sz+ZNHhKkeWEJml/cRpPv787wi3SoNK/AYN//wpDyU4vuOVkIB36tqVt91Y069gIl8PF/Dd+5PDuZM67tAWD7u9LXNXCZyu5dQ/fHljLdwfWsO7knmJ9J8UhUr3YnzmEdbsbo8GA7tUZ9EBfbnn6Or9589sOL/pnFikNa1Ti47GhT8lVSibYfP5h2cxFCNEbmAoYgJlSymeEEBOBVVLK+UIIKzAXaIPviX/IqQHiQFTwLx9ub/Uge9YHzgWfl8mss2DXvxGeaGJCq7Yh4KvfvL2IV4ZPL1aNRpOB2IRY3lj5HAnV44M65+2ti5i585dS28c36sF9mLY7kZ78V6zeoBqvLX+W2PjTK55dLi8d73u1lFqWn8VkYOHzdxFjK+1uQOWUUt3MRUr5nZSykZSyvpTymZxj46SU83M+d0gpr5FSNpBSXlBU4FfKj/ptagdZUhJdwVsKMwzzd684HA6uq303lxsG09t2HYkNqhJfvXjTMj1uLycOpfDOI3ODKv/53uXMKIXAbzOYuSapIwPFedj36AUCP8ChHYeZenf+m53ZHLn00oWxmo3c1aeTCvxnCbXCVynU47PvC7rszY+Vwu5N2ukn8+1/76KffSjH9h1HSonb6eGxHpM4cSglpKp//3JFUOWmbPo2pPqL69Gm/RndrD99o1piNPsfnpMSln29Ercr/3qK/p1Kb5WtJgT1EuOpUakC079bzmWjpzF7oXrXXt6p4K8UadQbtwd4RQISoUmuvTeZnteFFnSLJWZS7qfD2zwSuFkhOLWJe1FcEdq160xvb/8JgHqt6+AtpG26LvF68s8QmjDsCoZ1b4dWCou9JJKdh06w49BxHC4PKRnZvPLlbzz0dnCL6JSyoRK7KUXqN/wK+g2/gjlPfcy2tbsY8eotVEuqysiO9zL8yaU0bpMdxuRthYgZj2YLPb1DUWLjo4suBBjQ8BYzY2cojjl9exjbY2wMnTCYmU/My5l1lV/DtvWw2k93tZw52FurgpH9qZ6IdVMFGjb8Zd0OjqVmkFAhuP9XpXSpJ38laMPGX8ukrx6jWlJVAF5f/hpN28nSCfxo4A5ujjtAy4uaUP+8OhiMhqDHIcZ9/nBQ5W6oE1p6ieJKsJwexL129JU89O5wjKbT34/J4puO+uA7d+eW8zfLZ1+qB2sZZXtYvLbwRXdK2VFP/krJxDwB6eNK4UI6eIKfJ/Dsj2OwWn35J/Zs3MfIDo/jyAy8l2/jC+rTqktw/eQjm/Rkd+ZRlh7dFHR7QjGuZf41BpffdDGdrzqfH2f9yuYV26jWrAbNep9HtUa+BW8jXgm8pWN2Ga2aqZEQxqyuSliFZapnJKipnmcHKSXySOB9bsPHCLZr0SqMzz2y4vs1jO0zuUDJuKoV0AwamWnZNLmgAQ++czfpJzJ4++E5bFm5ndhKMdRtmcS21TsxWkyMeOUWOl95QbFbdDj7JP2XFL7rWSgMCB5q2o+ra3f0+7rHo3PXK5+ydvvptZSdmtVm2cbIrTMIhdVk5M9X7y3rZpxzSnWefySo4H920Q83imDtAkQUotJ8hLFmvlcO7T3KLQ1H4nUH7oPXDBozN0+lRv1EDu44zLxnv2Djn1uo3qAa1z8xkGadQr959Vz8LCdcGSGfn9tGBJ93fYhqlgr5dgbzZ+RrX/BnOQv0Z7KajcwaPYRGNSsXXVgJq2CDv+r2UUIi9Qxk1gfg/Bm0BDBeC56Pw3wVKwgTmM9HxIwuEPgB3nt8HrKIoUzdq/O/W9/i/ml3cm/Hx3FmudC9Ovu2HGTdL+t5bO4ougwIvE9uYWZ1uofr/niFTE/gLiV/BNAguhoZHgddKjdhROPLsRsLSZN66nvR9XIb+KOtZu4d0JlalePo2DTY9SFKWVHBXwmalBK8+5B6BpwcCfpRoHhBLzgCtEREwhcILfCK2xOHU/j9ixXo7qLfvW5duZ0Zj3+AI8OZb+tDZ5aLl+5+m6rn16NhCE+p1WwV+aX7eFYc28a29MN0rtyIutFVGfrbq2zJPMzlCc2Z2PY6Xtg0n+8PrMUjdVrG1eKpVtdS1Vr8/nCXJ/KzjEJhNGh8/+ztRKkFXmcN1e2jBEW6NyNPjgLvIcANkZjqGDsevMkI03lg6YooImPmxuVbeaLXM2SmBpfAzBJtxplRMPGcNAiybu5AnbrVqF4plkMn0mjXsCZDu7ejcsXyN03x/BFT8erl7+929iNDaFnXf7ZVpfSobh8lbKSehTxxI8i0iF5Hs99QrPI1Gybidga/U5i/wH+KQ0g270tm875kALYfOManS/+hVuUKVIy2cW2387i0TQO/SdRK281XnM+M7/8q62YUsHnXfhX8zyJqnr9SNOcPIEuesrhwgdMTBxJbKYbLb74k6PISyGyViCfOnu+Ys2ZFOGOQ1e3Vcbo9bD94nFVb9zNu9g9M+XxpsdsYCSP6d+b+gRdht5gQgKaV/Q0JYPKnv5d1E5RiUMFfKZr3KBB8muSQGBuGdNrI125F87dZeQDSZiJ1QEtctXJSNQtw1yk6k2e2y8MnS/4m+WTJZ/aEw7Ae7fl96khWv/UAMx4cjNVUPt7EX/f0nLJughIkFfyVopnbEvEeQsOZO38GeZrBwI/uTxBB3gCcDauAyUBm1/q+zERGA7rdHNS5JoOBtdsPhNTOSGpdvzpv3TeINvWrYzKW7Z/0lgPHSc+K8IOCEhYq+CtFM7UHYyS35LMjrD1LVMPnyTNpfXFzzFYTBlPBgWIJeOJtEOUL9LrViDfajDQb8NQsfMOW03VIKsWW7daIgbSuX50ZD19LjfjYsm4KgybO4VhqZlk3QymCCv5KkYQQiErvgvniYp5phugniy5magYlDP4xcdH8b/EEZmycyl0vDsWQ8wQsASnAWb8Sade0OX2CEOixVtL6t4Ag+sw1IYi1W2nboOBag/Ki75h32Z18sqybQUpGNtO/XVbWzVCKoIK/EhQhzGjx08E8KEAJIxCLb2FWNIgYsN8BGZMClM8rDiGKP+DrT7U6VajZuAaWnCyXXpMGEiw7jhP/9p9UnLncl4ZSStL7tUCPLbiwql5iJSYOu4Jom5koqxmLyUBslIXW9auzaPVW3J7SSelcXAdPpJd1EwDwenWW/KP2ayrvVPBXiiSlEz1tEvrhJuD6PEApDwgvaDEQ+yyIKpD1RnAXcC8KW1sBWnVtmruQy+DWEZD7obl14qYvA2NO19AZ61ysJiNjrr+Mvp2a8fOLd3P/gC5ICelZTn5ctYWJ7y/ihufmkeUojY3qg+d0RmKxXeiirMGNoyhlRwV/pUjy5L2Q9QlFLuySmb5Vv6mjQN9RrGvoeviCqcVmYcy8+wFfwM/r1E2AI6ng1dEynGjJ6Qinh/PqJfLW/YNo08A3+Kwh+N9nS3F5vLmLqrJdbvYcPsH7P68JW3vDwWIpPytrrWYjQy5pU3RBpUyp4K8USnp2g3M5kUnjkPdCpwdppXQhS7hbVqvurQrN+GPdeAQ0gXHnUeK+/JeaX/zL7Y3r07pe9dwyH/66Fqe74PoGt1fn+782l6h9/0VRVjNmo4Er2jfm6otalXVzlCKUj8nBSvnl2QXCGPLWiEHLmIy09UGmjQPPNsCItF2FiB2DELZiV/fy50sKfd3ROAEAV6uauA6kY091Yj6jq2LagsCDll5Z/nLs/Pr8PVz86Jtlcu2EWBuPX9edJklVSCwHM46UoqngrxTOWB9k8CkUQpb9ATL7Y06/w3BB9tdI7xFE/DvFru7bFZuw4ntrm7frJ/ceVv30wq70Ho3xHEzlge9+I/urxcRF2zEYBFmFpI7o1rJesdsUabGxFta89QBL/97O/dMWlNp1K0VbWfj83UUXVMoV1e2jFEoYk8DSBYh0n7KXgl1LTnAtR3r2FqsmKSUut5eTd12Ys8V8/o8TV5+X/wSTgeykODKdbnQJx9OzSD4ZeJ66AEZc1blYbSpNXVs34Ip2kdxfIb/UrPI1+K0ERwV/pUii4itgv8E3hbO0f2WEGbzFC/5CCNo3qokQkHLXhZwY0py0FtU4MbA1KXddCJX8LNQqRsK2JrWqYDWFZ2pqpEy+vU+pXcujl78uMKVoKvgrRRLCjBb7GFrVNWjVNoNtCFD0xiNhIZ1gbFDs0x4bcinRVgtmkwEqVMDTuR5UjiowtTMU9w28qMR1lIb/3XNVqVzHaFBh5GykfmpKsYnYcRB1M4hIpzqwgrUXwlCt2GfWqRbPV0/dzB29O3BZm4bcelk76q49gEhzlOgGYNAEx9LOjtQFl7asi9UU+YyfN14WydQfSqSozVyUkJVs8/Yo4MwgagZDNfAmgxYN9hsRUXciRMnnJdzb8XG2rdmF1+PF0SCBzK71EVJitltwFnPFrtVs5IFBXbmma+sStyvSdF1n/OyFfPvXprDXrWmCod3bcd+As+Od0Lki2M1c1JO/ErLQAz9QZRVYBwHmnLEEC9gGIhJ+QKv2D1qVP9Gi7wlL4N+/9SC71u/FmxPkrduPETd7JdE/beWCZCdfTbipWPU5XB7enP8nejncTetMo6d/E5HAD2A2GmjfqFZE6lYiT031VEKiHy7hbJLkZlBpERjqgmspGFtC9INhCfZnSjmSitFkyJ1LlN2sKtnnJyFNBn6T8NuE2X7Pa1Qzga37j/l9LcvhIiPbSWxUKY19BGHWjyv5YdVmOjerw70DLsLl8fDL38VbaV0cDpeHt79ZRufmdSJ2DSVySvSXJoSIBz4G6gC7gcFSyhQ/5bzAvzlf7pVS9i/JdZX/AgnHu5/+0v0XZM9Ar/g2mjX43bmCUa91bTwu31O/o1FlsjrWAT9pn88UKPADmE1GomzlI3/N0dQ0rnhsRu7XW/cf472Fq5g6PDx/Zt9MupVBE+f4Xe184FhqWK6hlL6Sdvs8BvwspWwI/JzztT/ZUsrzcj5U4FcCO3lX2PL8XDNpFu2GT+HysTMY9Eg/rFEWstvXCirwF2BM8MYAABLGSURBVMZqNnLT5e0xaBopGdls2ZdMZhkmeus75j2/xx94a35Y6t+dnBJwk5hGNSuH5RpK6Stp8L8SOPWeeTZQOnPLlHJgdOSqzppbotP3HjtG2+FT2HEwBQlkuz28dvgQFUddhowu+WK16y9ty9DL2jJm5vf0evwdbn/5U7o/8jZvzv+DsphA4fb6n2cfjpaYDBper87wvhdiNefvKLCajYy4svwudlMKV9LgX1VKeQgg598qAcpZhRCrhBDLhRDqBvEfoFW7I3KVew+W6PSrnvR/89hw4iRRtpIFf6vZyF19OvLSZ0tZvG47Lo+XTIcLp9vD+z+v4cvf15eo/vLG7dWZ+9NqLj6vPmOu705SlYrYLCbOq1+dt0YNokWd4k/DVcqHIvv8hRA/Af5+wmOKcZ0kKeVBIUQ9YLEQ4l8pZYGRKCHEncCdAElJScWoXikLWrWt6IffAV4Mb8W24HoGD2w7xPiBL7B34wEQ0KxjIybOf7TQczJK0D1jNRsZ1sM3g27B8g043fmniDpcHmYvWsXAi1qGfI3yaO32A9z8wkcsmHQrfTo0LevmKGFS5JO/lLK7lLKFn4+vgSNCiESAnH+TA9RxMOffncCvgN9k31LK6VLK9lLK9pUrq77Es4FW7Q6IGlncs8AUYG64oTGauej58xknM7it+QPs2bDft95Al2z4cwtD640oZluCE2u30KpeImu27efpD37Kze9/ppSMrIhcvzCDAtxswjULx6tLMh0ufv0ncjOHlNJX0m6f+cCpSdI3AV+fWUAIESeEsOR8ngB0BjaW8LpKeRI10jdlMwhata1o1TajVZoBlRaDoQm+N6A2sN0ElQr8Cvk14/F5ufP288pKy87dpjFcXh95FQ6Xh78272PV1v0sWL4Rj59+dgH59gMoLWOu784DA7tiyJOf6IJG1Xlt5ICwzfjJdrrVzJ7/mBKt8BVCVAI+AZKAvcA1UsoTQoj2wN1SytuFEBcCb+PbBkoDpkopZwSsNIda4Xv20bO/gewvQYuF6HvhWP5N2bVqW8N2rTtbPciu9fv8vpbZrAqOLvV9XwjhuxEIQbTNjIYgLbvgxjS1q1Zk1oNX02fc7EJTOZ9J4BtYNWgCi8nIrNFDaFAjIYTvqOQOHz5M76c+LHB8zVsP8MLHi9my7yhuj5f1e46EVH9CrJ1be57PewtX4fboXNG+MQ9f3RWDoWSzp5TwCnaFr0rvoJyVnrr6f/z+xYqAr2dXjyarVwsw5AR/LfCb3FMBPBQdmiRxJCWdFnUTub3XBSRViQuxppJrO3xKwNcGd2vN8k17OHAsNWCXVShsFiNLX7pH3QDKkWCDv1rhq5yV/t/enUdJVd0JHP/+ausNGhpooNlbQJTFBRBlMTEYEBmHTTmKCzgSHcUlxmQCjmNiyETGoMaoJOOWjCIqEHSGISAQoCdHMyCIICA74tCsLUgj0NBd3Xf+qNecpruqq4qq92r7fc6p01X1br/7q9vVv3p133333jfzrqDJ/0yPQk5f1RmT64WKKsj2hJ2uOZZU+NTE4bQpaBrDHuLj0KFDjW6f9z8bbam34qyfp99dyZN3DrNl/8o+OrePSklFxW34+YKfkFXnKtszFxdyavBFmDxfIOHnWj+jmKs/Gjk+T1IkfoBXPtyQsLo/XLuNap3TP+Vo8lcpa8jYq1l0ag5vfzmL2XtmUXFVp4ZX79qU+AEqKv08+cclCbmwq75/HHFF+EI2qaj0c9O/vMG+suMJi0FFT5O/SnltOrembZfW1OQ5P9fOnz/ZxpyV6x2vt762bRN7sVXZ8VM8FqfpJJQzNPmrtOFJ0EnH15eEPvHspPW//1HC6q4xhv1Hy/nqcIN5HVWS0hO+Km1M+N4VzF7h/FH4yYrkWcC87gfA+OlvsvvgMcfqdolQURn5MFmVWHrkr9LGj275LoX5di8t2VBx2xaO1xmJ+T+bRMdWzRyrz+N20a1dYq5xUNHTI3+VVspOODu9gkuEqbfGd/2BeFj4ty38eu5KKqr8NM/L5vipM7bV5XG78LhdTL97hC7mnkI0+au04hJwcnXFbK+Hvt06OFdhBP717WW8//GWc49jTfyFzfL4uvxUg+shcnweBvbqQrsW+dzyncsSeoGbip4mf5VWxg7uzQIHp1U+XVnFZ7v306978nwA1E388VBWfqrBc9k+D29NnUBX7eZJWfodTaWVJ+4YRm6Ws8c0y9ZtT5qLnHbsCzqxbtydqfQnbHSVig9N/irtfPTCw/xy0g3keJ35EFj4v1/w+BuLHakrnPy8HMfqWrRaJ+dNZZr8VVr6u2t6suLZB+hY2Nz2us5W+flo85fsKC2zva5w2rZoisu+i5rPU9gsz5mKlC00+au0le3zMHvqBEfqMsawYXdsy0/Gyys/vMX2OlwijBtyme31KPto8ldpLT8vO677GzOoNz5Pw75uj8tFywRcYxBMvx4d+eSlh7miaxF2fAlwu4TfThmNx6PpI5XpX0+lvR+PHRiX/Ywe2JMpowbhrrc2gAA+r4dre0e2mpkTPB4Pv7pnJN76E91dAJ/HTZfWBTw4ahC/mDicNS89wuAkeq3qwmjyV2nvjuHX8PFvz19n2HsBFyPt3H+UVs3yePGhMRQ2yyPH5yXb66FzmwJee2w8PodOMEeqqEU+U24aRJbXg9slUU1w6vO4KW7bgo6FzZg4rB+zH7+dyTdezd8P7IWrkYVxVOpIrnerUjaorPLzDzPnkuX1cLbKD0BVkDV4w7msaxEA/bp3YMnT97L38DE8bjedWtt/UvlCTRzen8G9u7B03XaqawzXX9mNO//t/KUeH58wlGfnlZxrkzYFTfiPf7otadYqUPbQZRxV2jnyzUkAWhc0AWDRmi+Y8c7KmCYdE+CTlx9J6+UKDx47QV62j/zc+J4nUc7SZRxVxln+6XYe/8MSaqz5HVwuYcY9N/K3zXujSvxulyAIfuvCrbwsL3/86W1pnfgh0E2kMocmf5UWyspPMvX18y+0qqkxTH19MTdf2zuqRdqrawz9urfntcfGxz1OpZKFnrlRaeHnby4NuW3djv1RL9L+2a79sQWkVJLT5K/SQmPrx/7fkehXl6oxhjOV/lhCUiqpafJXaaFnpzYht4Ua0+AJM/TRo0MaVRrTd7dKC09NGh7172Rl+ejRMfiUxNk+Dx9+uo0qf3WsoSmVlDT5q7SQ4/Mx78m7aJabFbasz+Mm2+thxuSRjBvcO+gUCGcq/Tw9ZwWTn5tHZZV2/6j0o6N9VNro1q4Vq56bwtCf/I7jp8422O71uLlzaF8K8nPp2bk1U15YQGUjF3udqfKzc38Zi9duY8yg3naGrpTj9MhfpZWSDTuCJn6AsYN68/DYIXy0eQ8/eG5+o4m/1tmqapat2x7vMJVKOL3CV6WVvg/8Ju77vLxrEVd0bU/Jxt00zcni9qFXckP/Hkg0k+Uo5ZBIr/CN6chfRMaLyBYRqRGRkJWJyAgR2S4iu0RkWix1KuW0TXsO8c7Kz/jq8Dds3nuI6W8v54X3/3pu+zcnKzhwtJxkPZBSKphY+/w3A+OAV0IVEBE3MAsYBpQCa0VkoTFG14BTKaHGGGrqjPo5U+lnbslGRg3sxcx5JXy2+wAuEfJzs3hq4g0M7Nk5gdEqFZmYjvyNMVuNMeE6RAcAu4wxe4wxlcB7wOhY6lUqlF6dCh2pp8YY7n1+Put3llLlr+ZslZ+y8lP8+JWFfHnomCMxKBULJ074tgf21Xlcaj2nVNzNfvxO3HFcxNYlBB0K6q+u4fipM/hrzu/qqfJX817JhrjVr5RdwiZ/EfmLiGwOcov06D3Y/07QzlERuU9E1onIurKyxC+GrVLT2lmPUvLM5Jj24fUIWV4PfYrbRbVIS3WNobSRqSaUShZh39XGmO/HWEcp0LHO4w5A0JWujTGvAq9CYLRPjPWqDJafn8+t113OBx9tpjLIVbptmudx+PipoL8rAi89OI52LfPpUNicko27+cVbyyg/fSZsvVleD1dd3DFsOaUSzYlun7VAdxEpFhEfcBuw0IF6VYZ7ZMy1ZIc4ag+V+AFWPHMfAy7pRIfCwApd113elUW/mhy2O8njdpGfm8XN1/a58KCVckisQz3HikgpMBD4s4gstZ5vJyKLAYwxfuAhYCmwFZhnjNkSW9hKhTf40Zc5URH8gq/G3D1zboPn8rJ9DOldjKfe2r9ul9C8SQ5tCpoybkgf3vnnO2iqK2GpFKAXeam0NGLaqxwpD310H87vH7mZqy/tdN5z33x7mnt/M59Dx77FGIMB+hQX8eKDY8hKssXbVebSZRxVRosl8QM88OICXntsPP26dzj3XEHTXOY/OZH1O/ez7+vjXNy+kJ6dQ08lrVQy07l9lAph+uzlDZ4TEfpd3IExg3pr4lcpTZO/Sku5Wd6Y97Gv7DjfRjDCR6lUpMlfpaXlM2Ib51/r3VV6wZZKT5r8VVrKycnh4+fvp1lebCNvVm7YFaeIlEoumvxV2srJyWHVsw/QK4a++eYxfngolaw0+au099Nbv0e2z4Mryjl/cnxeJgy90qaolEosTf4q7fUpLuLtabdzY/9L6NauJSMHXILP3bBccdsW5GR5aZLtw+d1c9ewfnz3sq7OB6yUA/QiL6Usxhi27TvCsROn6dWlLc2b5CQ6JKWiphd5KRUlEeHSTjp2X2UG7fZRSqkMpMlfKaUykCZ/pZTKQJr8lVIqA2nyV0qpDJS0Qz1FpAz4qpEirYCvHQonGhpXdDSu6Ghc0cnEuDobYwrDFUra5B+OiKyLZCyr0zSu6Ghc0dG4oqNxhabdPkoplYE0+SulVAZK5eT/aqIDCEHjio7GFR2NKzoaVwgp2+evlFLqwqXykb9SSqkLlDLJX0Rmisg2EflcRD4QkeYhyo0Qke0isktEpjkQ13gR2SIiNSIS8uy9iOwVkU0iskFEbJ+uNIq4nG6vFiKyXER2Wj8LQpSrttpqg4gstCmWRl+7iGSJyFxr+xoR6WJHHBcQ190iUlanfX7gUFx/EJEjIrI5xHYRkRetuD8Xkb5JEtd1IlJep71+5kBMHUVklYhstf4PfxikTELa6xxjTErcgOGAx7r/DPBMkDJuYDdwEeADNgI9bY7rUqAHUAL0b6TcXqCVg+0VNq4EtdevgWnW/WnB/o7WtpM2xxH2tQNTgH+37t8GzHXg7xZJXHcDLzv1XqpT73eAvsDmENtHAksAAa4B1iRJXNcBixxuqyKgr3W/KbAjyN8xIe1Ve0uZI39jzDJjjN96uBroEKTYAGCXMWaPMaYSeA8YbXNcW40x2+2s40JEGJfj7WXt/03r/pvAGJvrCyWS11431j8B14tIdMuB2RNXQhhj/goca6TIaOAtE7AaaC4iRUkQl+OMMQeNMeut+98CW4H29YolpL1qpUzyr+ceAp+Y9bUH9tV5XErDBk8UAywTkU9F5L5EB2NJRHu1McYchMA/CNA6RLlsEVknIqtFxI4PiEhe+7ky1oFHOdDShliijQvgZqur4E8i0tHmmCKVzP9/A0Vko4gsEZFeTlZsdRdeCayptymh7ZVUi7mIyF+AtkE2PWGM+S+rzBOAH5gTbBdBnot5OFMkcUVgsDHmgIi0BpaLyDbriCWRcTneXlHsppPVXhcBK0VkkzFmd6yx1RHJa7elfcKIpM7/Bt41xpwVkfsJfDsZanNckUhEe0ViPYEpD06KyEjgP4HuTlQsIk2ABcCjxpgT9TcH+RXH2iupkr8x5vuNbReRScBNwPXG6jSrpxSoexTUAThgd1wR7uOA9fOIiHxA4Ot9TMk/DnE53l4iclhEiowxB62vuEdC7KO2vfaISAmBI6d4Jv9IXnttmVIR8QDNsL97IWxcxpijdR6+RuAcWDKw5f0Uq7pJ1xizWER+JyKtjDG2zvkjIl4CiX+OMeb9IEUS2l4p0+0jIiOAqcAoY8zpEMXWAt1FpFhEfARO0tkyUiQaIpInIk1r7xM4eR10ZILDEtFeC4FJ1v1JQINvKCJSICJZ1v1WwGDgizjHEclrrxvrLcDKEAcdjsZVr194FIH+5GSwEJhojWK5Biiv7eJLJBFpW3uuRkQGEMh7Rxv/rZjrFOANYKsx5vkQxRLbXk6eXY7lBuwi0D+2wbrVjsJoByyuU24kgTPruwl0f9gd11gCn+BngcPA0vpxERi5sdG6bUmWuBLUXi2BFcBO62cL6/n+wOvW/UHAJqu9NgGTbYqlwWsHphM4wADIBuZb771PgIvsbp8I45phvY82AquASxyK613gIFBlvbcmA/cD91vbBZhlxb2JRka/ORzXQ3XaazUwyIGYhhDowvm8Ts4amQztVXvTK3yVUioDpUy3j1JKqfjR5K+UUhlIk79SSmUgTf5KKZWBNPkrpVQG0uSvlFIZSJO/UkplIE3+SimVgf4f4GyZxaoOIH4AAAAASUVORK5CYII=
plt.axis('equal')
plt.scatter(x,y, c=dbscan.labels_)
plt.show()
```
# Faktoryzacja macierzy
### Na razie na małych danych
```
u, s, vh = svds(utility_matrix, k = 1000)
u.shape, s.shape, vh.shape
mean_squared_error(utility_matrix.toarray()[utility_matrix.nonzero()], (u * s @ vh)[utility_matrix.nonzero()])
um_arr_non_zero = utility_matrix.nonzero()
um_arr = utility_matrix.toarray()[um_arr_non_zero]
f = open('MSEs.txt','w')
for k in range(3500, 3901, 100):
u, s, vh = svds(utility_matrix, k = k)
val = mean_squared_error(um_arr, (u * s @ vh)[um_arr_non_zero])
np.save(f"svd_table_{k}", u * s @ vh)
print(f"MSE dla k={k}: {val}")
f.write(f"MSE dla k={k}: {val}\n")
f.close()
arr = np.load('svd_table_3900.npy')
arr.shape
arr
u, s, vh = svds(utility_matrix, k = 10)
u.shape, s.shape, vh.shape
u * s @ vh
```
| github_jupyter |
```
import os, sys
import numpy as np
sys.path.insert(0, '/Users/kit/Documents/code/fair-classification/disparate_mistreatment/synthetic_data_demo')
sys.path.insert(0, '/Users/kit/Documents/code/fair-classification/fair_classification/')
from generate_synthetic_data import *
import utils as ut
import funcs_disp_mist as fdm
import plot_syn_boundaries as psb
def test_synthetic_data():
""" Generate the synthetic data """
data_type = 1
X, y, x_control = generate_synthetic_data(data_type=data_type, plot_data=False) # set plot_data to False to skip the data plot
sensitive_attrs = list(x_control.keys())
""" Split the data into train and test """
train_fold_size = 0.5
x_train, y_train, x_control_train, x_test, y_test, x_control_test = ut.split_into_train_test(X, y, x_control, train_fold_size)
cons_params = None # constraint parameters, will use them later
loss_function = "logreg" # perform the experiments with logistic regression
EPS = 1e-4
def train_test_classifier():
w = fdm.train_model_disp_mist(x_train, y_train, x_control_train, loss_function, EPS, cons_params)
train_score, test_score, cov_all_train, cov_all_test, s_attr_to_fp_fn_train, s_attr_to_fp_fn_test = fdm.get_clf_stats(w, x_train, y_train, x_control_train, x_test, y_test, x_control_test, sensitive_attrs)
# accuracy and FPR are for the test because we need of for plotting
return w, test_score, s_attr_to_fp_fn_test
""" Classify the data while optimizing for accuracy """
print()
print("== Unconstrained (original) classifier ==")
w_uncons, acc_uncons, s_attr_to_fp_fn_test_uncons = train_test_classifier()
print("\n-----------------------------------------------------------------------------------\n")
""" Now classify such that we optimize for accuracy while achieving perfect fairness """
print()
print( "== Classifier with fairness constraint ==")
print( "\n\n=== Constraints on FPR ===") # setting parameter for constraints
cons_type = 1 # FPR constraint -- just change the cons_type, the rest of parameters should stay the same
tau = 5.0
mu = 1.2
sensitive_attrs_to_cov_thresh = {"s1": {0:{0:0, 1:1e-6}, 1:{0:0, 1:1e-6}, 2:{0:0, 1:1e-6}}} # zero covariance threshold, means try to get the fairest solution
cons_params = {"cons_type": cons_type,
"tau": tau,
"mu": mu,
"sensitive_attrs_to_cov_thresh": sensitive_attrs_to_cov_thresh}
w_cons, acc_cons, s_attr_to_fp_fn_test_cons = train_test_classifier()
psb.plot_boundaries(X, y, x_control, [w_uncons, w_cons], [acc_uncons, acc_cons], [s_attr_to_fp_fn_test_uncons["s1"], s_attr_to_fp_fn_test_cons["s1"]], "img/syn_cons_dtype_%d_cons_type_%d.png"%(data_type, cons_type) )
print( "\n-----------------------------------------------------------------------------------\n")
print( "\n\n=== Constraints on FNR ===")
cons_type = 2
cons_params["cons_type"] = cons_type # FNR constraint -- just change the cons_type, the rest of parameters should stay the same
w_cons, acc_cons, s_attr_to_fp_fn_test_cons = train_test_classifier()
psb.plot_boundaries(X, y, x_control, [w_uncons, w_cons], [acc_uncons, acc_cons], [s_attr_to_fp_fn_test_uncons["s1"], s_attr_to_fp_fn_test_cons["s1"]], "img/syn_cons_dtype_%d_cons_type_%d.png"%(data_type, cons_type) )
print( "\n-----------------------------------------------------------------------------------\n")
print( "\n\n=== Constraints on both FPR and FNR ===")
cons_type = 4
cons_params["cons_type"] = cons_type # both FPR and FNR constraints
w_cons, acc_cons, s_attr_to_fp_fn_test_cons = train_test_classifier()
psb.plot_boundaries(X, y, x_control, [w_uncons, w_cons], [acc_uncons, acc_cons], [s_attr_to_fp_fn_test_uncons["s1"], s_attr_to_fp_fn_test_cons["s1"]], "img/syn_cons_dtype_%d_cons_type_%d.png"%(data_type, cons_type) )
print( "\n-----------------------------------------------------------------------------------\n")
return
test_synthetic_data()
```
## JOCO DATA TEST
```
import pandas as pd
FILE_PATH = '/Volumes/dbs/triage-working/matrices'
label_col = 'booking_view_warr_bw_1y'
demo_col = 'race_nonwhite'
exclude_cols = ['entity_id', 'as_of_date', label_col, demo_col]
def load_matrix(matrix_id, non_white):
df = pd.read_csv('%s/%s.csv' % (FILE_PATH, matrix_id))
df[demo_col] = non_white
df[label_col] = 2*(df[label_col] - 0.5)
return df
# training data
df_b_train = load_matrix('bf8f725453aebbcf3decbb72fcf95e6e', 1)
df_h_train = load_matrix('ee4a84ff473f0af6acf7d35432dcddfa', 1)
df_w_train = load_matrix('f6baf93649b912647017bb15b87131e0', 0)
df_train = pd.concat([df_b_train, df_h_train, df_w_train], axis=0)
x_train = df_train[[c for c in df_train.columns if c not in exclude_cols]].values
y_train = df_train[label_col].values
x_control_train = {demo_col: df_train[demo_col].values}
# test data
df_b_test = load_matrix('b6e936186bec0885640f5a0bb8ab787b', 1)
df_h_test = load_matrix('6b616fa0db246fc6222049e709efd430', 1)
df_w_test = load_matrix('0db1a017699f3d3142d7688128f844f1', 0)
df_test = pd.concat([df_b_test, df_h_test, df_w_test], axis=0)
x_test = df_test[[c for c in df_test.columns if c not in exclude_cols]].values
y_test = df_test[label_col].values
x_control_test = {demo_col: df_test[demo_col].values}
x_train.shape
df_train.shape
# TODO for JOCO Data:
# 1. load a training and test set
# 2. add columns to feature matrices for constant term (assumed to be in the X matrix here)
# 3. scale matrices as in scaled logit
# 4. read in race_3way and modify to race_2way for x_control
# 5. run a lasso logit as a means of feature selection
# 6. set up parameters for disparate mistreatment logit (cons_type=2 for FNR <=> recall)
# 7. run regressions, test on test set
# 8. iterate over full set of matrices/as_of_dates
# MAYBE modify the package with...
# - ability to use a ridge or lasso loss function
# - allow for multiple sensitive features (e.g., one-hot enconded categorical?)
loss_function = "logreg"
EPS = 1e-4
cons_params = None
def train_test_classifier():
w = fdm.train_model_disp_mist(x_train, y_train, x_control_train, loss_function, EPS, cons_params)
train_score, test_score, cov_all_train, cov_all_test, s_attr_to_fp_fn_train, s_attr_to_fp_fn_test = fdm.get_clf_stats(w, x_train, y_train, x_control_train, x_test, y_test, x_control_test, sensitive_attrs)
# accuracy and FPR are for the test because we need of for plotting
return w, test_score, s_attr_to_fp_fn_test
# Unconstrained classifier
cons_params = None
w_uncons, acc_uncons, s_attr_to_fp_fn_test_uncons = train_test_classifier()
# FNR fairness-constrained classifier
cons_type = 2 # FNR constraint -- same as recall
tau = 5.0
mu = 1.2
# note thresh for constraints will be cov_thresh[1] - cov_thresh[0]
# had trouble satisfying exact 0s here in synthetic data tests, so go with small epsilon, 1e-6...
sensitive_attrs_to_cov_thresh = {"s1": {0:{0:0, 1:1e-6}, 1:{0:0, 1:1e-6}, 2:{0:0, 1:1e-6}}} # zero covariance threshold, means try to get the fairest solution
cons_params = {"cons_type": cons_type,
"tau": tau,
"mu": mu,
"sensitive_attrs_to_cov_thresh": sensitive_attrs_to_cov_thresh}
w_cons, acc_cons, s_attr_to_fp_fn_test_cons = train_test_classifier()
x = x_train
y = y_train
x_control = x_control_train
max_iters = 150
max_iter_dccp = 75
num_points, num_features = x.shape
import cvxpy
w = cvxpy.Variable(num_features)
np.random.seed(112233)
w.value = np.random.rand(x.shape[1])
constraints = []
loss = cvxpy.sum( cvxpy.logistic( cvxpy.multiply(-y, x*w) ) ) / num_points
prob = cvxpy.Problem(cvxpy.Minimize(loss), constraints)
tau, mu = 0.005, 1.2
prob.solve(method='dccp', tau=tau, mu=mu, tau_max=1e10, solver=cvxpy.ECOS, verbose=True,
feastol=EPS, abstol=EPS, reltol=EPS, feastol_inacc=EPS, abstol_inacc=EPS, reltol_inacc=EPS,
max_iters=max_iters, max_iter=max_iter_dccp)
ret_w = np.array(w.value).flatten()
x_control_train = {demo_col: x_control_train}
x_control_test = {demo_col: x_control_test}
sensitive_attrs = list(x_control_train.keys())
train_score, test_score, cov_all_train, cov_all_test, s_attr_to_fp_fn_train, s_attr_to_fp_fn_test = fdm.get_clf_stats(ret_w, x_train, y_train, x_control_train, x_test, y_test, x_control_test, list(sensitive_attrs))
s_attr = sensitive_attrs[0]
distances_boundary_test = fdm.get_distance_boundary(ret_w, x_test, x_control_test[s_attr])
def label_top_k(distances, k):
sort_df = pd.DataFrame({'dist': distances})
sort_df.sort_values('dist', ascending=False, inplace=True)
sort_df['pred_label'] = -1
sort_df['orig_idx'] = sort_df.index
sort_df.reset_index(inplace=True)
i = k-1
sort_df.loc[:i,'pred_label'] = 1
sort_df.sort_values('orig_idx', inplace=True)
return sort_df['pred_label'].values
def calc_prec(pred_label, actual_label):
label_pos = sum((pred_label == 1).astype(int))
true_pos = sum(np.logical_and(pred_label == actual_label, pred_label == 1).astype(int))
return float(true_pos/label_pos)
k = 100
all_class_labels_assigned_test = label_top_k(distances_boundary_test, k)
#all_class_labels_assigned_test = np.sign(distances_boundary_test)
prec_k = calc_prec(all_class_labels_assigned_test, y_test)
print('prec@%s_abs: %.5f' % (k, prec_k))
s_attr_to_fp_fn_test = fdm.get_fpr_fnr_sensitive_features(y_test, all_class_labels_assigned_test, x_control_test, sensitive_attrs, False)
for s_attr in s_attr_to_fp_fn_test.keys():
for s_val in s_attr_to_fp_fn_test[s_attr].keys():
s_attr_to_fp_fn_test[s_attr][s_val]['recall'] = 1.000-s_attr_to_fp_fn_test[s_attr][s_val]['fnr']
recall_nonwhite = s_attr_to_fp_fn_test['race_nonwhite'][1]['recall']
recall_white = s_attr_to_fp_fn_test['race_nonwhite'][0]['recall']
print('recall non-white: %.6f' % recall_nonwhite)
print('recall white: %.6f' % recall_white)
print('recall ratio: %.6f' % float(recall_white/recall_nonwhite))
# 35% precision is incredibly low here -- likely because the logistic loss is totally unregularized,
# so presumably we're getting a massively overfit model...
# let's take a look at training set precision as an indication
k = 100
distances_boundary_train = fdm.get_distance_boundary(ret_w, x_train, x_control_train[s_attr])
all_class_labels_assigned_train = label_top_k(distances_boundary_train, k)
#all_class_labels_assigned_test = np.sign(distances_boundary_test)
prec_k = calc_prec(all_class_labels_assigned_train, y_train)
print('prec@%s_abs: %.5f' % (k, prec_k))
s_attr_to_fp_fn_train = fdm.get_fpr_fnr_sensitive_features(y_train, all_class_labels_assigned_train, x_control_train, sensitive_attrs, False)
for s_attr in s_attr_to_fp_fn_train.keys():
for s_val in s_attr_to_fp_fn_train[s_attr].keys():
s_attr_to_fp_fn_train[s_attr][s_val]['recall'] = 1.000-s_attr_to_fp_fn_train[s_attr][s_val]['fnr']
recall_nonwhite = s_attr_to_fp_fn_train['race_nonwhite'][1]['recall']
recall_white = s_attr_to_fp_fn_train['race_nonwhite'][0]['recall']
print('recall non-white: %.6f' % recall_nonwhite)
print('recall white: %.6f' % recall_white)
print('recall ratio: %.6f' % float(recall_white/recall_nonwhite))
```
yup -- training set precision at 100 is 100%, compared to 35% on the test set, so we're massively overfit.
Unfortunately, the best bet here is probably to scale the data and use a lasso to do feature selection, though perhaps we could simply pull the existing feature importances from the triage database as a first step here (scaling isn't strictly necessary since there's no regularization term here)
```
# load columns with non-zero lasso coefficients from one of the old models
from sklearn.externals import joblib
old_mod_hash = '/Users/kit/Documents/joco_working/5270794c51b1de17745de599be68e951'
old_mat_hash = '/Users/kit/Documents/joco_working/510a0fbff9a6f649ef881fe07b12b08e'
old_mod = joblib.load(open(old_mod_hash, 'rb'))
old_mat = pd.read_csv('%s.csv' % old_mat_hash, nrows=1)
old_exclude_cols = ['entity_id', 'as_of_date', 'booking_view_warr_bw_1y']
all_cols = [c for c in old_mat.columns if c not in old_exclude_cols]
keep_cols = []
for i, col in enumerate(all_cols):
if old_mod.coef_[0][i] != 0:
keep_cols.append(col)
keep_cols = keep_cols + ['intercept']
# forgot to do this above...
df_train['intercept'] = 1
df_test['intercept'] = 1
x_train = df_train[[c for c in df_train.columns if c in keep_cols]].values
y_train = df_train[label_col].values
x_control_train = {demo_col: df_train[demo_col].values}
x_test = df_test[[c for c in df_test.columns if c in keep_cols]].values
y_test = df_test[label_col].values
x_control_test = {demo_col: df_test[demo_col].values}
x = x_train
y = y_train
x_control = x_control_train
max_iters = 150
max_iter_dccp = 75
num_points, num_features = x.shape
import cvxpy
w = cvxpy.Variable(num_features)
np.random.seed(112233)
w.value = np.random.rand(x.shape[1])
constraints = []
loss = cvxpy.sum( cvxpy.logistic( cvxpy.multiply(-y, x*w) ) ) / num_points
prob = cvxpy.Problem(cvxpy.Minimize(loss), constraints)
tau, mu = 0.005, 1.2
prob.solve(method='dccp', tau=tau, mu=mu, tau_max=1e10, solver=cvxpy.ECOS, verbose=True,
feastol=EPS, abstol=EPS, reltol=EPS, feastol_inacc=EPS, abstol_inacc=EPS, reltol_inacc=EPS,
max_iters=max_iters, max_iter=max_iter_dccp)
ret_w_uncons = np.array(w.value).flatten()
train_score, test_score, cov_all_train, cov_all_test, s_attr_to_fp_fn_train, s_attr_to_fp_fn_test = fdm.get_clf_stats(ret_w_uncons, x_train, y_train, x_control_train, x_test, y_test, x_control_test, list(sensitive_attrs))
s_attr = sensitive_attrs[0]
distances_boundary_test = fdm.get_distance_boundary(ret_w_uncons, x_test, x_control_test[s_attr])
k = 100
all_class_labels_assigned_test = label_top_k(distances_boundary_test, k)
#all_class_labels_assigned_test = np.sign(distances_boundary_test)
prec_k = calc_prec(all_class_labels_assigned_test, y_test)
print('prec@%s_abs: %.5f' % (k, prec_k))
s_attr_to_fp_fn_test = fdm.get_fpr_fnr_sensitive_features(y_test, all_class_labels_assigned_test, x_control_test, sensitive_attrs, False)
for s_attr in s_attr_to_fp_fn_test.keys():
for s_val in s_attr_to_fp_fn_test[s_attr].keys():
s_attr_to_fp_fn_test[s_attr][s_val]['recall'] = 1.000-s_attr_to_fp_fn_test[s_attr][s_val]['fnr']
recall_nonwhite = s_attr_to_fp_fn_test['race_nonwhite'][1]['recall']
recall_white = s_attr_to_fp_fn_test['race_nonwhite'][0]['recall']
print('recall non-white: %.6f' % recall_nonwhite)
print('recall white: %.6f' % recall_white)
print('recall ratio: %.6f' % float(recall_white/recall_nonwhite))
s_attr = sensitive_attrs[0]
distances_boundary_test = fdm.get_distance_boundary(ret_w_uncons, x_test, x_control_test[s_attr])
k = 500
all_class_labels_assigned_test = label_top_k(distances_boundary_test, k)
#all_class_labels_assigned_test = np.sign(distances_boundary_test)
prec_k = calc_prec(all_class_labels_assigned_test, y_test)
print('prec@%s_abs: %.5f' % (k, prec_k))
s_attr_to_fp_fn_test = fdm.get_fpr_fnr_sensitive_features(y_test, all_class_labels_assigned_test, x_control_test, sensitive_attrs, False)
for s_attr in s_attr_to_fp_fn_test.keys():
for s_val in s_attr_to_fp_fn_test[s_attr].keys():
s_attr_to_fp_fn_test[s_attr][s_val]['recall'] = 1.000-s_attr_to_fp_fn_test[s_attr][s_val]['fnr']
recall_nonwhite = s_attr_to_fp_fn_test['race_nonwhite'][1]['recall']
recall_white = s_attr_to_fp_fn_test['race_nonwhite'][0]['recall']
print('recall non-white: %.6f' % recall_nonwhite)
print('recall white: %.6f' % recall_white)
print('recall ratio: %.6f' % float(recall_white/recall_nonwhite))
```
Ok -- that's at least a bit more in line with what we've seen with the regularized logistic regressions in general, so at least this makes a little more sense.
```
# let's take a look at training set precision as an indication
k = 100
distances_boundary_train = fdm.get_distance_boundary(ret_w_uncons, x_train, x_control_train[s_attr])
all_class_labels_assigned_train = label_top_k(distances_boundary_train, k)
#all_class_labels_assigned_test = np.sign(distances_boundary_test)
prec_k = calc_prec(all_class_labels_assigned_train, y_train)
print('prec@%s_abs: %.5f' % (k, prec_k))
s_attr_to_fp_fn_train = fdm.get_fpr_fnr_sensitive_features(y_train, all_class_labels_assigned_train, x_control_train, sensitive_attrs, False)
for s_attr in s_attr_to_fp_fn_train.keys():
for s_val in s_attr_to_fp_fn_train[s_attr].keys():
s_attr_to_fp_fn_train[s_attr][s_val]['recall'] = 1.000-s_attr_to_fp_fn_train[s_attr][s_val]['fnr']
recall_nonwhite = s_attr_to_fp_fn_train['race_nonwhite'][1]['recall']
recall_white = s_attr_to_fp_fn_train['race_nonwhite'][0]['recall']
print('recall non-white: %.6f' % recall_nonwhite)
print('recall white: %.6f' % recall_white)
print('recall ratio: %.6f' % float(recall_white/recall_nonwhite))
```
Admittedly, still seems a bit overfit and we could consider stronger regularization for the feature selection, but given the test set error in line with other models, I think we can forge ahead with this for now and see how it performs including the FNR fairness constraints.
```
loss_function = "logreg"
EPS = 1e-4
cons_params = None
def train_test_classifier():
w = fdm.train_model_disp_mist(x_train, y_train, x_control_train, loss_function, EPS, cons_params)
train_score, test_score, cov_all_train, cov_all_test, s_attr_to_fp_fn_train, s_attr_to_fp_fn_test = fdm.get_clf_stats(w, x_train, y_train, x_control_train, x_test, y_test, x_control_test, sensitive_attrs)
# accuracy and FPR are for the test because we need of for plotting
return w, test_score, s_attr_to_fp_fn_test
# FNR fairness-constrained classifier
cons_type = 2 # FNR constraint -- same as recall
tau = 5.0
mu = 1.2
# note thresh for constraints will be cov_thresh[1] - cov_thresh[0]
# had trouble satisfying exact 0s here in synthetic data tests, so go with small epsilon, 1e-6...
sensitive_attrs_to_cov_thresh = {"race_nonwhite": {0:{0:0, 1:1e-6}, 1:{0:0, 1:1e-6}, 2:{0:0, 1:1e-6}}} # zero covariance threshold, means try to get the fairest solution
cons_params = {"cons_type": cons_type,
"tau": tau,
"mu": mu,
"sensitive_attrs_to_cov_thresh": sensitive_attrs_to_cov_thresh}
ret_w, acc_cons, s_attr_to_fp_fn_test_cons = train_test_classifier()
```
A little surprising that this doesn't yield perfectly-balanced FNR at the original decision boundary, or even particularly close, but let's see what happens at the top 100...
```
s_attr = sensitive_attrs[0]
distances_boundary_test = fdm.get_distance_boundary(ret_w, x_test, x_control_test[s_attr])
k = 100
all_class_labels_assigned_test = label_top_k(distances_boundary_test, k)
#all_class_labels_assigned_test = np.sign(distances_boundary_test)
prec_k = calc_prec(all_class_labels_assigned_test, y_test)
print('prec@%s_abs: %.5f' % (k, prec_k))
s_attr_to_fp_fn_test = fdm.get_fpr_fnr_sensitive_features(y_test, all_class_labels_assigned_test, x_control_test, sensitive_attrs, False)
for s_attr in s_attr_to_fp_fn_test.keys():
for s_val in s_attr_to_fp_fn_test[s_attr].keys():
s_attr_to_fp_fn_test[s_attr][s_val]['recall'] = 1.000-s_attr_to_fp_fn_test[s_attr][s_val]['fnr']
recall_nonwhite = s_attr_to_fp_fn_test['race_nonwhite'][1]['recall']
recall_white = s_attr_to_fp_fn_test['race_nonwhite'][0]['recall']
print('recall non-white: %.6f' % recall_nonwhite)
print('recall white: %.6f' % recall_white)
print('recall ratio: %.6f' % float(recall_white/recall_nonwhite))
```
Not much change from the unconstrained model (in fact, a little worse) and the raw ratio is very high compared to what we get after choosing separate thresholds, while the precision is much lower that we can achieve with random forests.
```
s_attr = sensitive_attrs[0]
distances_boundary_test = fdm.get_distance_boundary(ret_w, x_test, x_control_test[s_attr])
k = 500
all_class_labels_assigned_test = label_top_k(distances_boundary_test, k)
#all_class_labels_assigned_test = np.sign(distances_boundary_test)
prec_k = calc_prec(all_class_labels_assigned_test, y_test)
print('prec@%s_abs: %.5f' % (k, prec_k))
s_attr_to_fp_fn_test = fdm.get_fpr_fnr_sensitive_features(y_test, all_class_labels_assigned_test, x_control_test, sensitive_attrs, False)
for s_attr in s_attr_to_fp_fn_test.keys():
for s_val in s_attr_to_fp_fn_test[s_attr].keys():
s_attr_to_fp_fn_test[s_attr][s_val]['recall'] = 1.000-s_attr_to_fp_fn_test[s_attr][s_val]['fnr']
recall_nonwhite = s_attr_to_fp_fn_test['race_nonwhite'][1]['recall']
recall_white = s_attr_to_fp_fn_test['race_nonwhite'][0]['recall']
print('recall non-white: %.6f' % recall_nonwhite)
print('recall white: %.6f' % recall_white)
print('recall ratio: %.6f' % float(recall_white/recall_nonwhite))
```
Almost identical story at k=500 as well.
Overall, this approach doesn't seem to working particularly well in the "top k" setting, which isn't terribly surprising since it wasn't designed for that (it's attempting to balance FNR based on the decision boundary from the underlying optimization problem, so there's no good reason to believe that simply shifting this boundary to yield a smaller set would preserve the fairness constraints as well). It wouldn't be fair to entirely discount this method because it doesn't work here, but we certainly do need methods that can work with the top k/resource constrained setting, as well as the flexibility to apply to a broader array of modeling methods than simple linear models as employed here.
Given this, it doesn't strike me as particularly worthwhile to clean up the code here to get this working robustly across every joco as_of_date / other datasets. Rather, I think we could provide it as a simple case study at the outset and then focus more deeply on the strategies that are better suited to the top k problem.
```
# let's pickle a couple of things for potential future use / figure creation...
import pickle
export_dict = {
'inputs': {
'x_train': x_train,
'x_control_train': x_control_train,
'y_train': y_train,
'x_test': x_test,
'x_control_test': x_control_test,
'y_test': y_test,
'sensitive_attrs': sensitive_attrs
},
'unconstrained': {
'model_output': {
'w': ret_w_uncons
},
'top_100_abs': {
'precision': 0.51000,
'recall_non_white': 0.010674,
'recall_white': 0.015916,
'recall_ratio': 1.491161
},
'top_500_abs': {
'precision': 0.52600,
'recall_non_white': 0.048032,
'recall_white': 0.086858,
'recall_ratio': 1.808328
}
},
'FNR constrined': {
'model_output': {
'w': ret_w
},
'top_100_abs': {
'precision': 0.53000,
'recall_non_white': 0.010674,
'recall_white': 0.016826,
'recall_ratio': 1.576370
},
'top_500_abs': {
'precision': 0.52400,
'recall_non_white': 0.049366,
'recall_white': 0.085493,
'recall_ratio': 1.731819
}
}
}
pickle.dump(export_dict, open('zafar_fair_classification.pkl', 'wb'))
del(export_dict)
```
## DEBUGGING....
```
data_type=1
X, y, x_control = generate_synthetic_data(data_type=data_type, plot_data=False)
train_fold_size = 0.5
x_train, y_train, x_control_train, x_test, y_test, x_control_test = ut.split_into_train_test(X, y, x_control, train_fold_size)
cons_params = None
loss_function = "logreg"
EPS = 1e-4
def train_test_classifier():
w = fdm.train_model_disp_mist(x_train, y_train, x_control_train, loss_function, EPS, cons_params)
train_score, test_score, cov_all_train, cov_all_test, s_attr_to_fp_fn_train, s_attr_to_fp_fn_test = fdm.get_clf_stats(w, x_train, y_train, x_control_train, x_test, y_test, x_control_test, sensitive_attrs)
# accuracy and FPR are for the test because we need of for plotting
return w, test_score, s_attr_to_fp_fn_test
w = fdm.train_model_disp_mist(x_train, y_train, x_control_train, loss_function, EPS, cons_params)
x = x_train
y = y_train
x_control = x_control_train
max_iters = 100
max_iter_dccp = 50
num_points, num_features = x.shape
import cvxpy
w = cvxpy.Variable(num_features)
np.random.seed(112233)
w.value = np.random.rand(x.shape[1])
constraints = []
loss = cvxpy.sum( cvxpy.logistic( cvxpy.multiply(-y, x*w) ) ) / num_points
prob = cvxpy.Problem(cvxpy.Minimize(loss), constraints)
prob.is_dcp()
from dccp.problem import is_dccp
is_dccp(prob)
tau, mu = 0.005, 1.2
prob.solve(method='dccp', tau=tau, mu=mu, tau_max=1e10, solver=cvxpy.ECOS, verbose=False,
feastol=EPS, abstol=EPS, reltol=EPS, feastol_inacc=EPS, abstol_inacc=EPS, reltol_inacc=EPS,
max_iters=max_iters, max_iter=max_iter_dccp)
prob.status
ret_w = np.array(w.value).flatten()
train_score, test_score, cov_all_train, cov_all_test, s_attr_to_fp_fn_train, s_attr_to_fp_fn_test = fdm.get_clf_stats(ret_w, x_train, y_train, x_control_train, x_test, y_test, x_control_test, list(sensitive_attrs))
w_uncons, acc_uncons, s_attr_to_fp_fn_test_uncons = ret_w, test_score, s_attr_to_fp_fn_test
psb.plot_boundaries(X, y, x_control, [w_uncons, w_uncons], [acc_uncons, acc_uncons],
[s_attr_to_fp_fn_test_uncons['s1'], s_attr_to_fp_fn_test_uncons['s1']],
"test_output.png")
print( "\n\n=== Constraints on FPR ===") # setting parameter for constraints
cons_type = 2 # FPR constraint -- just change the cons_type, the rest of parameters should stay the same
tau = 5.0
mu = 1.2
sensitive_attrs_to_cov_thresh = {"s1": {0:{0:0, 1:0.000001}, 1:{0:0, 1:0.000001}, 2:{0:0, 1:0.000001}}} # zero covariance threshold, means try to get the fairest solution
cons_params = {"cons_type": cons_type,
"tau": tau,
"mu": mu,
"sensitive_attrs_to_cov_thresh": sensitive_attrs_to_cov_thresh}
w_cons, acc_cons, s_attr_to_fp_fn_test_cons = train_test_classifier()
import cvxpy
x = x_train
y = y_train
x_control = x_control_train
max_iters = 100
max_iter_dccp = 50
num_points, num_features = x.shape
w = cvxpy.Variable(num_features) # this is the weight vector
np.random.seed(112233)
w.value = np.random.rand(x.shape[1])
constraints = fdm.get_constraint_list_cov(x, y, x_control, cons_params["sensitive_attrs_to_cov_thresh"], cons_params["cons_type"], w)
constraints
loss = cvxpy.sum( cvxpy.logistic( cvxpy.multiply(-y, x*w) ) ) / num_points
cons_params.get('take_initial_sol')
take_initial_sol = True
w.value
p = cvxpy.Problem(cvxpy.Minimize(loss), [])
p.solve()
w.value
prob = cvxpy.Problem(cvxpy.Minimize(loss), constraints)
# prob = cvxpy.Problem(cvxpy.Minimize(loss), [cvxpy.norm(w) >= 0.1])
cons_params.get('tau')
cons_params.get('mu')
tau = cons_params.get('tau')
mu = cons_params.get('mu')
prob.solve(method='dccp', tau=tau, mu=mu, tau_max=1e10, solver=cvxpy.ECOS, verbose=True,
feastol=EPS, abstol=EPS, reltol=EPS, feastol_inacc=EPS, abstol_inacc=EPS, reltol_inacc=EPS,
max_iters=max_iters, max_iter=max_iter_dccp)
print(constraints[1])
ut.get_one_hot_encoding(x_control_train['s1'])
sum(np.logical_and(x_control_train['s1'] == 0, y_train == -1))
cvxpy.norm(w)
xx = cvxpy.Parameter(value=1)
yy = cvxpy.Variable()
pp = cvxpy.Problem(cvxpy.Minimize(yy**2 + cvxpy.abs(xx)))
pp.solve(verbose=True)
xx.value
yy.value
pp.status
constraints[0].value()
constraints[0].args[0].value
constraints[0].args[1].value
```
| github_jupyter |
# Example: CanvasXpress heatmap Chart No. 9
This example page demonstrates how to, using the Python package, create a chart that matches the CanvasXpress online example located at:
https://www.canvasxpress.org/examples/heatmap-9.html
This example is generated using the reproducible JSON obtained from the above page and the `canvasxpress.util.generator.generate_canvasxpress_code_from_json_file()` function.
Everything required for the chart to render is included in the code below. Simply run the code block.
```
from canvasxpress.canvas import CanvasXpress
from canvasxpress.js.collection import CXEvents
from canvasxpress.render.jupyter import CXNoteBook
cx = CanvasXpress(
render_to="heatmap9",
data={
"z": {
"Type": [
"Pro",
"Tyr",
"Pho",
"Kin",
"Oth",
"Pro",
"Tyr",
"Pho",
"Kin",
"Oth",
"Pro",
"Tyr",
"Pho",
"Kin",
"Oth",
"Pro",
"Tyr",
"Pho",
"Kin",
"Oth",
"Pro",
"Tyr",
"Pho",
"Kin",
"Oth",
"Pro",
"Tyr",
"Pho",
"Kin",
"Oth",
"Pro",
"Tyr",
"Pho",
"Kin",
"Oth",
"Pro",
"Tyr",
"Pho",
"Kin",
"Oth"
],
"Sens": [
1,
2,
3,
4,
1,
2,
3,
4,
1,
2,
3,
4,
1,
2,
3,
4,
1,
2,
3,
4,
1,
2,
3,
4,
1,
2,
3,
4,
1,
2,
3,
4,
1,
2,
3,
4,
1,
2,
3,
4
]
},
"x": {
"Treatment": [
"Control",
"Control",
"Control",
"Control",
"Control",
"TreatmentA",
"TreatmentB",
"TreatmentA",
"TreatmentB",
"TreatmentA",
"TreatmentB",
"TreatmentA",
"TreatmentB",
"TreatmentA",
"TreatmentB",
"TreatmentA",
"TreatmentB",
"TreatmentA",
"TreatmentB",
"TreatmentA",
"TreatmentB",
"TreatmentA",
"TreatmentB",
"TreatmentA",
"TreatmentB"
],
"Site": [
"Site1",
"Site1",
"Site1",
"Site1",
"Site1",
"Site2",
"Site2",
"Site2",
"Site2",
"Site2",
"Site2",
"Site2",
"Site2",
"Site2",
"Site2",
"Site3",
"Site3",
"Site3",
"Site3",
"Site3",
"Site3",
"Site3",
"Site3",
"Site3",
"Site3"
],
"Dose-Type": [
"",
"",
"",
"",
"",
"Dose1",
"Dose1",
"Dose2",
"Dose2",
"Dose3",
"Dose3",
"Dose4",
"Dose4",
"Dose5",
"Dose5",
"Dose1",
"Dose1",
"Dose2",
"Dose2",
"Dose3",
"Dose3",
"Dose4",
"Dose4",
"Dose5",
"Dose5"
],
"Dose": [
0,
0,
0,
0,
0,
5,
5,
10,
10,
15,
15,
20,
20,
25,
25,
5,
5,
10,
10,
15,
15,
20,
20,
25,
25
]
},
"y": {
"smps": [
"S1",
"S2",
"S3",
"S4",
"S5",
"S6",
"S7",
"S8",
"S9",
"S10",
"S11",
"S12",
"S13",
"S14",
"S15",
"S16",
"S17",
"S18",
"S19",
"S20",
"S21",
"S22",
"S23",
"S24",
"S25"
],
"vars": [
"V1",
"V2",
"V3",
"V4",
"V5",
"V6",
"V7",
"V8",
"V9",
"V10",
"V11",
"V12",
"V13",
"V14",
"V15",
"V16",
"V17",
"V18",
"V19",
"V20",
"V21",
"V22",
"V23",
"V24",
"V25",
"V26",
"V27",
"V28",
"V29",
"V30",
"V31",
"V32",
"V33",
"V34",
"V35",
"V36",
"V37",
"V38",
"V39",
"V40"
],
"data": [
[
0.784,
1.036,
-0.641,
1.606,
2.208,
3.879,
0.333,
2.265,
-1.55,
1.678,
-0.639,
-0.533,
-0.078,
0.433,
0.391,
1.013,
0.928,
0.812,
0.072,
3.564,
0.47,
1.836,
0.351,
3.139,
-2.207
],
[
0.222,
0.716,
0.993,
-0.913,
0.996,
1.235,
1.396,
1.817,
0.162,
1.137,
-0.126,
1.56,
1.003,
1.86,
0.43,
0.696,
0.777,
1.6,
0.175,
2.423,
0.044,
3.881,
-0.757,
1.486,
0.01
],
[
0.486,
2.15,
-0.069,
-0.468,
0.402,
0.725,
-1.697,
0.653,
0.101,
2.852,
-0.27,
0.414,
-0.789,
1.877,
1.555,
2.511,
0.07,
0.244,
-0.41,
2.345,
2.401,
-0.033,
0.951,
2.053,
0.725
],
[
-1.857,
0.698,
0.528,
1.024,
-0.035,
2.181,
-0.015,
3.68,
-1.13,
-0.842,
-1.759,
1.784,
-0.673,
0.147,
0.765,
1.585,
0.33,
1.481,
-0.362,
1.456,
-0.719,
0.961,
1.296,
2.375,
0.208
],
[
-1.19,
1.564,
-2.407,
0.642,
-0.51,
4.116,
-0.379,
0.786,
1.508,
3.119,
1.011,
1.54,
1.184,
1.821,
-0.217,
2.752,
0.083,
1.663,
0.568,
2.48,
-1.207,
1.222,
0.296,
1.055,
1.078
],
[
0.256,
1.214,
1.919,
0.577,
1.07,
1.53,
1.537,
3.063,
0.481,
2.332,
-1.466,
0.167,
0.428,
1.401,
-1.716,
3.524,
-0.822,
1.073,
-1.825,
3.923,
-0.542,
2.637,
-1.296,
0.759,
0.836
],
[
-0.443,
0.286,
0.379,
1.076,
0.478,
3.443,
-0.287,
1.206,
-1.275,
2.275,
1.101,
2.821,
-0.638,
0.922,
-0.205,
2.318,
0.494,
1.648,
-0.585,
1.963,
-0.636,
1.229,
0.998,
1.523,
-1.01
],
[
1.023,
-0.417,
0.865,
1.645,
0.324,
1.94,
0.122,
-0.171,
0.352,
1.42,
-0.436,
3.076,
0.434,
0.986,
-1.912,
3.899,
-0.212,
0.716,
0.782,
0.534,
1.939,
1.374,
-0.083,
2.318,
0.982
],
[
-2.33,
0.575,
-0.543,
-0.38,
-2.153,
1.717,
-1.219,
0.725,
0.273,
1.908,
0.488,
1.426,
0.108,
2.586,
0.653,
0.317,
0.112,
3.138,
0.212,
1.393,
-0.506,
1.87,
0.332,
1.893,
1.017
],
[
0.841,
0.146,
0.716,
-0.233,
-0.206,
0.237,
-0.307,
2.499,
-1.619,
1.957,
-0.12,
3.058,
0.511,
3.598,
0.286,
0.922,
0.164,
0.782,
-3.468,
0.262,
0.812,
0.798,
1.209,
2.964,
-1.47
],
[
-0.099,
1.666,
-1.635,
1.475,
-0.186,
0.781,
-1.919,
1.472,
-0.109,
1.588,
-0.379,
0.862,
-1.455,
2.386,
2.783,
0.98,
-0.136,
1.042,
0.532,
1.778,
0.463,
0.647,
0.92,
2.427,
-0.07
],
[
0.663,
-1.411,
-0.69,
-0.216,
-0.735,
1.878,
-0.073,
1.568,
-1.254,
3.792,
-0.345,
3.384,
0.206,
1.572,
0.134,
2.035,
-0.26,
2.42,
0.437,
2.164,
-0.063,
5.027,
-0.166,
3.878,
-1.313
],
[
-0.647,
-1.152,
3.437,
-0.3,
0.358,
1.766,
0.067,
0.149,
-1.005,
1.191,
-1.299,
1.326,
-2.378,
1.8,
-0.858,
2.019,
-1.357,
2.278,
-0.711,
2.196,
-0.243,
3.326,
-0.215,
2.25,
-0.504
],
[
-0.264,
-1.328,
1.228,
1.247,
0.692,
1.763,
-0.978,
2.781,
-0.058,
2.223,
0.796,
2.414,
-1.834,
3.203,
0.459,
2.914,
0.375,
3.309,
0.946,
0.943,
-1.365,
2.452,
0.474,
0.503,
0.025
],
[
0.253,
-0.529,
-0.429,
-1.111,
0.398,
2.332,
-1.334,
2.202,
-0.585,
1.885,
0.398,
1.788,
0.972,
2.025,
-0.835,
0.622,
0.001,
0.837,
-0.776,
2.257,
0.682,
1.304,
2.407,
4.038,
0.518
],
[
-0.876,
-1.41,
0.538,
-1.04,
-0.717,
-0.889,
3.129,
1.202,
3.398,
0.398,
3.857,
1.372,
4.813,
-1.311,
4.029,
-0.432,
3.01,
0.756,
4.688,
0.294,
4.61,
0.859,
4.498,
1.794,
3.319
],
[
-0.363,
0.042,
-0.253,
-0.076,
-1.27,
-0.904,
2.931,
-0.119,
2.669,
-0.165,
6.023,
-0.65,
2.031,
1.424,
2.844,
-1.019,
4.062,
-0.025,
2.637,
-0.317,
4.228,
-0.142,
3.013,
0.611,
3.74
],
[
-1.674,
-0.318,
-0.726,
-1.271,
1.753,
-1.678,
3.341,
-1.772,
3.814,
-1.391,
2.622,
0.677,
3.307,
-0.92,
3.545,
0.305,
2.808,
0.836,
4.532,
-0.378,
4.87,
-0.044,
4.061,
1.684,
5.486
],
[
-0.288,
0.165,
-0.468,
1.219,
-3.353,
-0.578,
3.414,
-0.674,
4.755,
0.033,
4.025,
0.44,
4.186,
1.136,
2.505,
0.436,
3.293,
-0.868,
4.746,
-0.545,
3.666,
-0.295,
3.206,
-0.966,
4.678
],
[
-0.558,
-0.855,
-1.114,
-0.623,
0.368,
-0.182,
4.37,
0.563,
3.75,
0.189,
2.717,
-1.708,
5.274,
0.741,
2.537,
-1.583,
2.832,
-1.515,
3.829,
0.358,
5.306,
0.388,
3.284,
0.661,
3.804
],
[
1.693,
-1.53,
0.057,
-0.217,
0.511,
0.309,
3.998,
0.925,
1.045,
0.379,
2.024,
0.331,
3.612,
0.151,
5.808,
-1.429,
3.402,
-0.297,
4.692,
-0.439,
4.521,
-0.816,
4.693,
0.323,
2.869
],
[
-0.234,
1.999,
-1.603,
-0.292,
-0.91,
-0.766,
6.167,
1.242,
4.219,
-1.291,
6.974,
-0.443,
4.039,
0.72,
3.808,
1.465,
2.86,
2.736,
4.675,
-0.554,
3.091,
0.057,
4.311,
-0.005,
2.605
],
[
0.529,
-1.721,
2.207,
-0.873,
-1.364,
1.139,
3.146,
1.277,
3.963,
-0.234,
4.581,
-1.266,
3.743,
-0.84,
3.682,
-0.566,
4.249,
0.599,
4.202,
0.125,
4.136,
-0.67,
3.433,
-0.954,
3.97
],
[
-0.529,
0.375,
0.204,
-0.529,
1.001,
0.244,
3.922,
-0.904,
3.479,
-0.926,
4.171,
-0.047,
2.158,
0.467,
2.277,
0.429,
3.903,
-1.013,
3.182,
0.73,
3.318,
-1.663,
4.222,
0.264,
3.538
],
[
2.302,
-0.218,
-1.184,
-0.644,
0.118,
-1.35,
4.497,
1.262,
5.131,
-1.095,
4.354,
-1.364,
4.376,
-0.936,
3.278,
0.753,
4.903,
-2.193,
3.336,
0.722,
3.92,
-1.341,
4.762,
1.756,
4.032
],
[
0.957,
1.309,
-1.317,
1.254,
-0.397,
0.004,
3.34,
1.233,
4.681,
-0.875,
2.497,
0.207,
1.703,
-0.614,
3.171,
-0.034,
2.59,
0.968,
3.576,
0.946,
3.85,
1.128,
4.015,
0.633,
3.148
],
[
-0.789,
-1.139,
0.066,
0.418,
0.366,
-0.932,
3.982,
0.151,
4.018,
0.74,
5.374,
0.067,
6.07,
1.178,
6.316,
1.948,
3.389,
0.383,
5.084,
-0.251,
3.874,
-0.715,
3.101,
-0.172,
4.867
],
[
-0.26,
-0.005,
-0.12,
-0.422,
0.629,
1.242,
3.954,
-0.027,
4.352,
-0.074,
4.369,
0.196,
4.847,
-0.763,
3.042,
-1.643,
3.952,
-1.358,
4.105,
-0.257,
4.168,
0.047,
1.782,
-0.585,
5.465
],
[
1.882,
0.869,
-1.305,
1.095,
1.002,
-0.897,
3.248,
1.113,
5.83,
0.298,
4.811,
-0.128,
3.263,
0.186,
4.244,
1.314,
2.832,
0.222,
3.899,
-1.279,
4.133,
-1.523,
4.49,
0.966,
4.658
],
[
-1.052,
0.429,
0.646,
0.642,
1.037,
-1.046,
1.724,
-0.698,
5.316,
-0.403,
2.821,
-0.108,
5.52,
-0.352,
3.298,
-0.716,
2.672,
1.499,
3.919,
0.202,
3.409,
0.841,
5.47,
1.225,
1.988
],
[
-1.862,
-0.589,
0.205,
1.281,
-1.256,
0.924,
4.189,
-1.219,
3.137,
0.142,
5.869,
0.529,
2.138,
-0.034,
3.921,
-1.097,
5.402,
1.468,
5.034,
0.088,
3.055,
1.587,
3.374,
0.377,
2.939
],
[
-0.315,
-0.369,
0.634,
0.495,
-1.059,
-0.481,
1.329,
1.105,
5.3,
0.135,
6.515,
0.001,
4.161,
1.686,
4.747,
-0.911,
3.24,
-1.461,
4.64,
0.698,
5.006,
-1.072,
4.608,
-0.317,
5.208
],
[
0.558,
0.793,
-1.713,
0.055,
2.242,
0.588,
3.785,
2.949,
2.175,
2.055,
3.328,
0.236,
3.549,
-0.009,
1.477,
0.538,
3.116,
-0.621,
5.203,
0.736,
3.606,
-0.313,
4.402,
-1.039,
3.894
],
[
-1.332,
-1.134,
0.153,
0.66,
1.764,
-0.588,
3.417,
-0.547,
3.849,
-1.521,
3.332,
0.88,
5.449,
0.179,
4.596,
0.626,
4.006,
0.33,
2.969,
-0.42,
2.606,
-0.485,
4.581,
-0.199,
5.008
],
[
0.29,
0.228,
0.117,
-0.587,
-2.116,
0.188,
4.009,
0.551,
3.127,
0.682,
3.858,
-1.053,
4.388,
-1.46,
1.468,
0.434,
4.221,
0.782,
2.992,
0.056,
5.223,
-0.747,
6.549,
-0.959,
3.714
],
[
-0.015,
-1.665,
1.007,
0.278,
-0.091,
1.919,
3.861,
-0.318,
3.026,
-1.642,
5.379,
2.097,
4.396,
0.802,
3.66,
0.544,
2.156,
0.87,
4.044,
0.3,
4.422,
-0.788,
4.677,
-0.215,
4.643
],
[
-0.984,
0.915,
0.944,
-1.975,
-1.717,
0.16,
4.748,
1.521,
4.091,
-0.386,
3.802,
-1.134,
5.701,
-0.402,
5.682,
-0.987,
4.281,
0.844,
3.427,
1.368,
3.358,
-1.748,
3.792,
2.125,
5.137
],
[
-0.399,
-0.613,
2.211,
0.238,
2.799,
0.687,
5.522,
0.534,
5.233,
-0.395,
4.387,
-1.733,
4.494,
-1.26,
4.693,
1.679,
4.477,
0.399,
2.508,
1.683,
3.185,
0.865,
4.958,
0.602,
4.371
],
[
1.205,
-0.562,
1.134,
0.202,
0.209,
0.692,
2.419,
0.397,
2.429,
0.911,
6.341,
0.713,
4.548,
-0.688,
3.947,
0.439,
4.69,
-0.324,
3.07,
0.265,
3.757,
-1.535,
5.434,
-0.017,
4.125
],
[
-0.298,
0.118,
1.653,
1.519,
-0.821,
-0.85,
4.602,
1.073,
5.087,
0.155,
6.987,
-0.716,
2.912,
0.581,
2.112,
-0.426,
3.523,
0.188,
4.548,
0.155,
4.256,
0.775,
2.607,
-0.697,
5.338
]
]
}
},
config={
"colorKey": {
"Treatment": "Accent",
"Sens": [
"white",
"green",
"rgb(72,126,182)",
"rgb(167,206,49)",
"rgb(248,204,3)"
],
"Type": "YlGn",
"Site": "Pastel1"
},
"colorSpectrum": [
"magenta",
"blue",
"black",
"red",
"gold"
],
"colorSpectrumZeroValue": 0,
"graphType": "Heatmap",
"heatmapIndicatorHeight": 50,
"heatmapIndicatorHistogram": True,
"heatmapIndicatorPosition": "topLeft",
"heatmapIndicatorWidth": 60,
"samplesClustered": True,
"showTransition": False,
"smpOverlayProperties": {
"Treatment": {
"position": "right",
"type": "Default",
"color": "rgb(254,41,108)",
"spectrum": [
"rgb(255,0,255)",
"rgb(0,0,255)",
"rgb(0,0,0)",
"rgb(255,0,0)",
"rgb(255,215,0)"
],
"scheme": "User",
"showLegend": True,
"showName": True,
"showBox": True,
"rotate": False
},
"Site": {
"position": "left",
"type": "Default",
"color": "rgb(72,126,182)",
"spectrum": [
"rgb(255,0,255)",
"rgb(0,0,255)",
"rgb(0,0,0)",
"rgb(255,0,0)",
"rgb(255,215,0)"
],
"scheme": "User",
"showLegend": True,
"showName": True,
"showBox": True,
"rotate": False
},
"Dose": {
"thickness": 50,
"position": "right",
"type": "Dotplot",
"color": "rgb(167,206,49)",
"spectrum": [
"rgb(255,0,255)",
"rgb(0,0,255)",
"rgb(0,0,0)",
"rgb(255,0,0)",
"rgb(255,215,0)"
],
"scheme": "User",
"showLegend": True,
"showName": True,
"showBox": True,
"rotate": False
}
},
"smpOverlays": [
"Treatment",
"Site",
"Dose"
],
"title": "Advanced Overlays in Heatmaps",
"varOverlayProperties": {
"Sens": {
"type": "Bar",
"position": "bottom",
"thickness": 20,
"color": "red",
"spectrum": [
"rgb(255,0,255)",
"rgb(0,0,255)",
"rgb(0,0,0)",
"rgb(255,0,0)",
"rgb(255,215,0)"
],
"scheme": "User",
"showLegend": True,
"showName": True,
"showBox": True,
"rotate": False
},
"Type": {
"position": "top",
"type": "Default",
"color": "rgb(254,41,108)",
"spectrum": [
"rgb(255,0,255)",
"rgb(0,0,255)",
"rgb(0,0,0)",
"rgb(255,0,0)",
"rgb(255,215,0)"
],
"scheme": "User",
"showLegend": True,
"showName": True,
"showBox": True,
"rotate": False
}
},
"varOverlays": [
"Type",
"Sens"
],
"variablesClustered": True
},
width=613,
height=613,
events=CXEvents(),
after_render=[],
other_init_params={
"version": 35,
"events": False,
"info": False,
"afterRenderInit": False,
"noValidate": True
}
)
display = CXNoteBook(cx)
display.render(output_file="heatmap_9.html")
```
| github_jupyter |
```
import json
import requests
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
#from tensorflow.contrib import lite
from keras.models import Sequential
from keras.layers import Activation, Dense, Dropout, LSTM
from keras.callbacks import EarlyStopping
import matplotlib.pyplot as plt
import plotly.offline as py
import plotly.graph_objs as go
import numpy as np
import pandas as pd
import seaborn as sns
from sklearn.metrics import mean_absolute_error
%matplotlib inline
endpoint = 'https://min-api.cryptocompare.com/data/histoday'
res = requests.get(endpoint + '?fsym=BTC&tsym=USD&limit=500')
data = pd.DataFrame(json.loads(res.content)['Data'])
data = data.set_index('time')
data.index = pd.to_datetime(data.index, unit='s')
data.head(5)
target_col = data['close']
len(target_col)
target_col
target_col.describe()
plt.rcParams['figure.figsize'] = [10,6]
plt.plot(target_col)
from sklearn.preprocessing import MinMaxScaler
scaler=MinMaxScaler(feature_range=(0,1))
target_col=scaler.fit_transform(np.array(target_col).reshape(-1,1))
##splitting dataset into train and test split
training_size=int(len(target_col)*0.80)
test_size=len(target_col)-training_size
train_data,test_data=target_col[0:training_size,:],target_col[training_size:len(target_col),:1]
# convert an array of values into a dataset matrix
def create_dataset(dataset, time_step=1):
dataX, dataY = [], []
for i in range(len(dataset)-time_step-1):
a = dataset[i:(i+time_step), 0] ###i=0, 0,1,2,3-----99 100
dataX.append(a)
dataY.append(dataset[i + time_step, 0])
return np.array(dataX), np.array(dataY)
# reshape into X=t,t+1,t+2,t+3 and Y=t+4
time_step=1
X_train, y_train = create_dataset(train_data, time_step)
X_test, ytest = create_dataset(test_data, time_step)
print(X_train.shape), print(y_train.shape)
print(X_test.shape), print(ytest.shape)
# reshape input to be [samples, time steps, features] which is required for LSTM
X_train =X_train.reshape(X_train.shape[0],X_train.shape[1] , 1)
X_test = X_test.reshape(X_test.shape[0],X_test.shape[1] , 1)
model=keras.Sequential()
model.add(LSTM(500,return_sequences=True,input_shape=(X_train.shape[1], X_train.shape[2])))
model.add(LSTM(500,return_sequences=True))
model.add(LSTM(500,return_sequences=False))
model.add(Dropout(0.2))
model.add(Activation('sigmoid'))
model.add(Dense(1))
#opt = keras.optimizers.Adam(learning_rate=0.01)
#model.compile(loss='categorical_crossentropy', optimizer=opt, metrics=['mse'])
model.compile(loss='mean_squared_error',optimizer='adam', metrics=['mae'])
model.summary()
history=model.fit(X_train,y_train,validation_data=(X_test,ytest),epochs=20,batch_size=32,verbose=1)
history.history['loss']
trace1 = go.Scatter(
x = np.arange(0, len(history.history['loss']), 1),
y = history.history['loss'],
mode = 'lines',
name = 'Train loss',
line = dict(color=('rgb(255,0,0)'), width=2, dash='dash')
)
trace2 = go.Scatter(
x = np.arange(0, len(history.history['val_loss']), 1),
y = history.history['val_loss'],
mode = 'lines',
name = 'Test loss',
line = dict(color=('rgb(1, 0, 255)'), width=2)
)
data = [trace1, trace2]
layout = go.Layout(
autosize=False,
width=750,
height=600,
title = 'Train and Test Loss during training',
xaxis= go.layout.XAxis(linecolor = 'black',
linewidth = 1,
mirror = True,
title = "Epoch number",
color = 'black'
),
yaxis= go.layout.YAxis(linecolor = 'red',
linewidth = 1,
mirror = True,
title = "Loss",
color = 'black'),
margin=go.layout.Margin(
l=50,
r=50,
b=100,
t=100,
pad = 4
)
)
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='training_process')
trace1 = go.Scatter(
x = np.arange(0, len(history.history['mae']), 1),
y = history.history['mae'],
mode = 'lines',
name = 'Train Mean Absolute Error',
line = dict(color=('rgb(0,0,139)'), width=2, dash='dash')
)
trace2 = go.Scatter(
x = np.arange(0, len(history.history['val_mae']), 1),
y = history.history['val_mae'],
mode = 'lines',
name = 'Test Mean Absolute Error',
line = dict(color=('rgb(1, 0, 0)'), width=2)
)
data = [trace1, trace2]
layout = go.Layout(
autosize=False,
width=750,
height=600,
title = 'Train and Test Mean Squared Error during training',
xaxis= go.layout.XAxis(linecolor = 'black',
linewidth = 1,
mirror = True,
title = "Epoch number",
color = 'black'
),
yaxis= go.layout.YAxis(linecolor = 'red',
linewidth = 1,
mirror = True,
title = "Mean Absolute Error",
color = 'black'),
margin=go.layout.Margin(
l=50,
r=50,
b=100,
t=100,
pad = 4
)
)
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='training_process')
# add one additional data point to align shapes of the predictions and true labels
#X_test = np.append(X_test, scaler.transform(data.iloc[-1][0]))
#X_test = np.reshape(X_test, (len(X_test), 1, 1))
# get predictions and then make some transformations to be able to calculate RMSE properly in USD
prediction = model.predict(X_test)
prediction_inverse = scaler.inverse_transform(prediction.reshape(-1, 1))
Y_test_inverse = scaler.inverse_transform(ytest.reshape(-1, 1))
prediction2_inverse = np.array(prediction_inverse[:,0][1:])
Y_test2_inverse = np.array(Y_test_inverse[:,0])
trace1 = go.Scatter(
x = np.arange(0, len(prediction2_inverse), 1),
y = prediction2_inverse,
mode = 'lines',
name = 'Predicted Labels',
line = dict(color=('rgb(0,0,139)'), width=2)
)
trace2 = go.Scatter(
x = np.arange(0, len(Y_test2_inverse), 1),
y = Y_test2_inverse,
mode = 'lines',
name = 'True labels',
line = dict(color=('rgb(255, 0, 0)'), width=2)
)
data = [trace1, trace2]
layout = go.Layout(
autosize=False,
width=750,
height=600,
title = 'Prediction Graph',
xaxis= go.layout.XAxis(linecolor = 'black',
linewidth = 1,
mirror = True,
title = "Number of days",
color = 'black'
),
yaxis= go.layout.YAxis(linecolor = 'red',
linewidth = 1,
mirror = True,
title = "Price in USD",
color = 'black'),
margin=go.layout.Margin(
l=50,
r=50,
b=100,
t=100,
pad = 4
)
)
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='results_demonstrating0')
plt.savefig('demonstrating0.png', dpi=300, transparent=False, bbox_inches='tight')
```
| github_jupyter |
```
import numpy as np
import pandas as pd
from datetime import time
pd.set_option('html', False)
from IPython.core.display import Image
Image('http://akamaicovers.oreilly.com/images/0636920023784/lrg.jpg')
```
Important Features in pandas
===
```
import pandas as pd
import numpy as np
```
Fast tabular data IO
---
```
temp = '/Users/wesm/Downloads/minutebars/%s.csv'
path = temp % 'AAPL'
!wc -l $path
aapl_bars = pd.read_csv(temp % 'AAPL')
aapl_bars
%time _ = pd.read_csv(path)
```
Time series operations
---
```
aapl_bars.dt
aapl_bars.index = pd.to_datetime(aapl_bars.pop('dt'))
aapl_bars.head()
def load_bars(ticker):
bars = pd.read_csv(temp % ticker)
bars.index = pd.to_datetime(bars.pop('dt'))
return bars
aapl_bars.at_time(time(15, 0)).head(10)
aapl_bars.close_price['2009-10-15']
aapl_bars.close_price
mth_mean = aapl_bars.close_price.resample('M', how=['mean', 'median', 'std'])
mth_mean
mth_mean.plot()
close = aapl_bars.close_price
close / close.shift(1) - 1
minute_returns = aapl_bars.close_price.pct_change()
std_10day = pd.rolling_std(minute_returns, 390 * 10)
std_10day.resample('B').plot()
```
Data alignment
---
```
ts1 = pd.Series(np.random.randn(10),
index=pd.date_range('1/1/2000', periods=10))
ts1
ts2 = ts1[[0, 2, 4, 5, 6, 7, 8]]
ts2
ts1 + ts2
df = pd.DataFrame({'A': ts1, 'B': ts2})
df
ibm_bars = load_bars('IBM')
def subsample(frame, pct=0.9):
N = len(frame)
indexer = np.sort(np.random.permutation(N)[:pct*N])
return frame.take(indexer)
f1 = subsample(ibm_bars)
f2 = subsample(aapl_bars)
f1
both = pd.concat([f1, f2], axis=1, keys=['IBM', 'AAPL'])
both.head(20)
```
Missing data handling
---
```
df
df.count()
both.count()
df.sum()
df.mean(1)
df.dropna()
df.fillna(0)
df.fillna(method='ffill')
df.asfreq('4h')
df.asfreq('4h').ffill(limit=3)
```
Groupby operations
---
```
import random, string
import matplotlib as mpl
def rands(n):
choices = string.ascii_letters
return ''.join([random.choice(choices) for _ in xrange(n)])
mpl.rc('figure', figsize=(12, 8))
ind_names = np.array(['ENERGY', 'FINANCIAL', 'TECH',
'CONSDUR', 'SERVICES', 'UTILITIES'], dtype='O')
ccys = np.array(['USD', 'EUR'], dtype='O')
Nfull = 2000
tickers = np.array(sorted(rands(5).upper() for _ in xrange(Nfull)), dtype='O')
tickers = np.unique(tickers)
industries = pd.Series(ind_names.take(np.random.randint(0, 6, Nfull)),
index=tickers, name='industry')
ccy = pd.Series(ccys.take(np.random.randint(0, len(ccys), Nfull)),
index=tickers, name='ccy')
ccy
df = pd.DataFrame({'Momentum' : np.random.randn(1000) / 200 + 0.03,
'Value' : np.random.randn(1000) / 200 + 0.08,
'ShortInterest' : np.random.randn(1000) / 200 - 0.02},
index=tickers.take(np.random.permutation(Nfull)[:1000]))
df.head()
means = df.groupby(industries).mean()
means
means.plot(kind='barh')
means = df.groupby([industries, ccy]).mean()
means
keys = [industries, ccy]
zscore = lambda x: (x - x.mean()) / x.std()
normed = df.groupby(keys).apply(zscore)
normed.groupby(keys).agg(['mean', 'std'])
```
Hierarchical indexing
---
```
means
means['Momentum']
means.ix['TECH']
means.stack()
means.stack().unstack('industry')
```
Merging and joining
---
```
base = '/Users/wesm/Dropbox/book/svn/book_scripts/movielens/ml-1m'
get_path = lambda x: '%s/%s.dat' % (base, x)
unames = ['user_id', 'gender', 'age', 'occupation', 'zip']
users = pd.read_table(get_path('users'), sep='::', header=None, names=unames)
rnames = ['user_id', 'movie_id', 'rating', 'timestamp']
ratings = pd.read_table(get_path('ratings'), sep='::', header=None, names=rnames)
mnames = ['movie_id', 'title', 'genres']
movies = pd.read_table(get_path('movies'), sep='::', header=None, names=mnames)
movies.head()
ratings.head()
users.head()
data = pd.merge(pd.merge(ratings, users), movies)
data
rating_counts = data.groupby('title').size()
freq_titles = rating_counts.index[rating_counts > 1000]
freq_titles
highest_rated = data.groupby('title').rating.mean()[freq_titles].order()[-20:]
highest_rated
filtered = data[data.title.isin(highest_rated.index)]
filtered.title = filtered.title.str[:25]
filtered.groupby(['title', 'gender']).rating.count().unstack()
```
Pivot tables
---
```
mean_ratings = data.pivot_table('rating', rows='title',
cols='gender', aggfunc='mean')
mean_ratings.tail(20)
```
Data summary, statistics
---
summary, value_counts, etc.
```
data.title.value_counts()
data.rating.describe()
by_gender = data.groupby('gender').rating.describe()
by_gender
by_gender.unstack(0)
```
| github_jupyter |
```
%matplotlib inline
import py4j
py4j.__version__
from sagas.ofbiz.services import OfService as s, oc, track
from sagas.ofbiz.entities import OfEntity as e
from py4j.java_gateway import get_field
ok, r=track(lambda a: s().testScv(defaultValue=5.5, message="hello world"))
print(ok, r)
from sagas.ofbiz.entities import OfEntity as e, finder
from py4j.protocol import Py4JError
entity="Tests"
limit=1
start=5
try:
result = finder.find_list(entity, limit, start)
print(oc.j.ValueHelper.valueListToJson(result))
except Py4JError as e:
print(e.errmsg+e.java_exception.getMessage())
from sagas.ofbiz.services import OfService as s, oc, track
ok, r=await s('srv').testScv(defaultValue='x', message="hello world")
print(ok,r)
ok, r=await s('srv').testScv(defaultValue=6.6, message="hello world")
print(ok,r)
import asyncio
import uuid
import json
from sagas.ofbiz.srv_client import SrvClient
loop = asyncio.get_event_loop()
rpc = await SrvClient(loop).connect()
print(" [x] Requesting ...")
json_pars = json.dumps({'_service': 'testScv',
'defaultValue': 5.5,
'message': "hello world"})
response = await rpc.call(json_pars)
print(" [.] Got %s" % response)
json_pars = json.dumps({'_service': 'testScv',
'defaultValue': 5.5,
'message': "hello world"})
response = await rpc.call(json_pars)
print(" [.] Got %s" % response)
def params(**kwargs):
print(json.dumps(kwargs))
params(defaultValue=5.5, message="hello world")
import json
json_pars=json.dumps({'defaultValue':5.5, 'message':"hello world"})
srv=oc.j.ServiceInvoker(oc.dispatcher, oc.delegator, "testScv", json_pars)
ret=srv.invoke()
print(ret, ret.getID())
json_r=srv.getJsonResult()
print(json.dumps(json.loads(json_r), indent=2))
result=srv.getResult()
json_r=oc.j.ValueHelper.mapToJson(result)
# print(json_r)
print(json.dumps(json.loads(json_r), indent=2))
async def multiply(*, x, y):
return x * y
r=await multiply(x=5,y=5)
print(r)
# import pandas as pd
# df=pd.read_json(json_r, orient='index')
# df
from IPython.display import display, JSON
JSON(json.loads(json_r))
def service_gen(name):
serv_model=oc.service_model(name)
params=serv_model.getModelParamList()
for param in params:
internal = get_field(param, "internal")
p_name = get_field(param, "name")
p_type=get_field(param, 'type')
p_mode=get_field(param, 'mode')
# print(p_name, internal)
if internal:
pass
else:
print(p_mode, p_name, p_type)
service_gen('testScv')
from sagas.ofbiz.service_gen import get_all_regular_services
services=get_all_regular_services()
print(len(services))
from ruamel.yaml import YAML
yaml = YAML()
with open('.services.yml', 'w') as outfile:
yaml.dump(services, outfile)
with open('.services.yml') as fp:
str_data = fp.read()
data = yaml.load(str_data)
print(len(data), type(data))
print(data[:5])
print('hel'.find('x'))
print('oF'.lower())
def norm_loc(loc):
pkg_prefixes=['ofbiz-framework/applications',
'ofbiz-framework/framework',
'ofbiz-framework/plugins']
for pkg_prefix in pkg_prefixes:
idx=loc.find(pkg_prefix)
if idx!=-1:
return loc[idx+len(pkg_prefix)+1:].replace('servicedef/','').replace('.xml','').replace('/','_').lower()
raise ValueError('Cannot normalize the location '+loc)
def get_service_package(srv):
serv_model = oc.service_model(srv)
def_loc = get_field(serv_model, 'definitionLocation')
return norm_loc(def_loc)
# from sagas.ofbiz.service_gen import get_service_package
service_groups={}
for srv in services:
pkg=get_service_package(srv)
if pkg in service_groups:
grp=service_groups[pkg]
grp.append(srv)
else:
service_groups[pkg]=[srv]
print(len(service_groups))
for k in service_groups.keys():
print(k)
srvs=service_groups['humanres_services_position']
print(len(srvs))
from sagas.ofbiz.entities import OfEntity as e
from sagas.ofbiz.entity_gen import norm_package
def get_entity_package_def(entity_name):
model = oc.delegator.getModelEntity(entity_name)
return norm_package(model.getPackageName())
print(get_entity_package_def('Person'))
def proc_service_refs(serv_name, deps):
model_serv = oc.service_model(serv_name)
def_ent = get_field(model_serv, 'defaultEntityName')
if def_ent != "":
ent_pkg=get_entity_package_def(def_ent)
deps.add(ent_pkg)
serv_grp='humanres_services_position'
srvs=service_groups[serv_grp]
deps=set()
for srv in srvs:
proc_service_refs(srv, deps)
print(deps)
```
| github_jupyter |
Copyright (c) Microsoft Corporation. All rights reserved.
Licensed under the MIT License.
## AutoML Installation
**For Databricks non ML runtime 7.1(scala 2.21, spark 3.0.0) and up, Install AML sdk by running the following command in the first cell of the notebook.**
%pip install --upgrade --force-reinstall -r https://aka.ms/automl_linux_requirements.txt
**For Databricks non ML runtime 7.0 and lower, Install AML sdk using init script as shown in [readme](https://github.com/Azure/MachineLearningNotebooks/blob/master/how-to-use-azureml/azure-databricks/automl/README.md) before running this notebook.**
# AutoML : Classification with Local Compute on Azure DataBricks
In this example we use the scikit-learn's to showcase how you can use AutoML for a simple classification problem.
In this notebook you will learn how to:
1. Create Azure Machine Learning Workspace object and initialize your notebook directory to easily reload this object from a configuration file.
2. Create an `Experiment` in an existing `Workspace`.
3. Configure AutoML using `AutoMLConfig`.
4. Train the model using AzureDataBricks.
5. Explore the results.
6. Test the best fitted model.
Prerequisites:
Before running this notebook, please follow the readme for installing necessary libraries to your cluster.
## Register Machine Learning Services Resource Provider
Microsoft.MachineLearningServices only needs to be registed once in the subscription. To register it:
Start the Azure portal.
Select your All services and then Subscription.
Select the subscription that you want to use.
Click on Resource providers
Click the Register link next to Microsoft.MachineLearningServices
### Check the Azure ML Core SDK Version to Validate Your Installation
```
import azureml.core
print("SDK Version:", azureml.core.VERSION)
```
## Initialize an Azure ML Workspace
### What is an Azure ML Workspace and Why Do I Need One?
An Azure ML workspace is an Azure resource that organizes and coordinates the actions of many other Azure resources to assist in executing and sharing machine learning workflows. In particular, an Azure ML workspace coordinates storage, databases, and compute resources providing added functionality for machine learning experimentation, operationalization, and the monitoring of operationalized models.
### What do I Need?
To create or access an Azure ML workspace, you will need to import the Azure ML library and specify following information:
* A name for your workspace. You can choose one.
* Your subscription id. Use the `id` value from the `az account show` command output above.
* The resource group name. The resource group organizes Azure resources and provides a default region for the resources in the group. The resource group will be created if it doesn't exist. Resource groups can be created and viewed in the [Azure portal](https://portal.azure.com)
* Supported regions include `eastus2`, `eastus`,`westcentralus`, `southeastasia`, `westeurope`, `australiaeast`, `westus2`, `southcentralus`.
```
subscription_id = "<Your SubscriptionId>" #you should be owner or contributor
resource_group = "<Resource group - new or existing>" #you should be owner or contributor
workspace_name = "<workspace to be created>" #your workspace name
workspace_region = "<azureregion>" #your region
```
## Creating a Workspace
If you already have access to an Azure ML workspace you want to use, you can skip this cell. Otherwise, this cell will create an Azure ML workspace for you in the specified subscription, provided you have the correct permissions for the given `subscription_id`.
This will fail when:
1. The workspace already exists.
2. You do not have permission to create a workspace in the resource group.
3. You are not a subscription owner or contributor and no Azure ML workspaces have ever been created in this subscription.
If workspace creation fails for any reason other than already existing, please work with your IT administrator to provide you with the appropriate permissions or to provision the required resources.
**Note:** Creation of a new workspace can take several minutes.
```
# Import the Workspace class and check the Azure ML SDK version.
from azureml.core import Workspace
ws = Workspace.create(name = workspace_name,
subscription_id = subscription_id,
resource_group = resource_group,
location = workspace_region,
exist_ok=True)
ws.get_details()
```
## Configuring Your Local Environment
You can validate that you have access to the specified workspace and write a configuration file to the default configuration location, `./aml_config/config.json`.
```
from azureml.core import Workspace
ws = Workspace(workspace_name = workspace_name,
subscription_id = subscription_id,
resource_group = resource_group)
# Persist the subscription id, resource group name, and workspace name in aml_config/config.json.
ws.write_config()
write_config(path="/databricks/driver/aml_config/",file_name=<alias_conf.cfg>)
```
## Create an Experiment
As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments.
```
import logging
import os
import random
import time
import json
from matplotlib import pyplot as plt
from matplotlib.pyplot import imshow
import numpy as np
import pandas as pd
import azureml.core
from azureml.core.experiment import Experiment
from azureml.core.workspace import Workspace
from azureml.train.automl import AutoMLConfig
from azureml.train.automl.run import AutoMLRun
# Choose a name for the experiment and specify the project folder.
experiment_name = 'automl-local-classification'
experiment = Experiment(ws, experiment_name)
output = {}
output['SDK version'] = azureml.core.VERSION
output['Subscription ID'] = ws.subscription_id
output['Workspace Name'] = ws.name
output['Resource Group'] = ws.resource_group
output['Location'] = ws.location
output['Experiment Name'] = experiment.name
pd.set_option('display.max_colwidth', -1)
pd.DataFrame(data = output, index = ['']).T
```
## Load Training Data Using Dataset
Automated ML takes a `TabularDataset` as input.
You are free to use the data preparation libraries/tools of your choice to do the require preparation and once you are done, you can write it to a datastore and create a TabularDataset from it.
```
# The data referenced here was a 1MB simple random sample of the Chicago Crime data into a local temporary directory.
from azureml.core.dataset import Dataset
example_data = 'https://dprepdata.blob.core.windows.net/demo/crime0-random.csv'
dataset = Dataset.Tabular.from_delimited_files(example_data)
dataset.take(5).to_pandas_dataframe()
```
## Review the TabularDataset
You can peek the result of a TabularDataset at any range using `skip(i)` and `take(j).to_pandas_dataframe()`. Doing so evaluates only j records for all the steps in the TabularDataset, which makes it fast even against large datasets.
```
training_data = dataset.drop_columns(columns=['FBI Code'])
label = 'Primary Type'
```
## Configure AutoML
Instantiate an `AutoMLConfig` object to specify the settings and data used to run the experiment.
|Property|Description|
|-|-|
|**task**|classification or regression|
|**primary_metric**|This is the metric that you want to optimize. Classification supports the following primary metrics: <br><i>accuracy</i><br><i>AUC_weighted</i><br><i>average_precision_score_weighted</i><br><i>norm_macro_recall</i><br><i>precision_score_weighted</i>|
|**primary_metric**|This is the metric that you want to optimize. Regression supports the following primary metrics: <br><i>spearman_correlation</i><br><i>normalized_root_mean_squared_error</i><br><i>r2_score</i><br><i>normalized_mean_absolute_error</i>|
|**iteration_timeout_minutes**|Time limit in minutes for each iteration.|
|**iterations**|Number of iterations. In each iteration AutoML trains a specific pipeline with the data.|
|**spark_context**|Spark Context object. for Databricks, use spark_context=sc|
|**max_concurrent_iterations**|Maximum number of iterations to execute in parallel. This should be <= number of worker nodes in your Azure Databricks cluster.|
|**n_cross_validations**|Number of cross validation splits.|
|**training_data**|Input dataset, containing both features and label column.|
|**label_column_name**|The name of the label column.|
```
automl_config = AutoMLConfig(task = 'classification',
debug_log = 'automl_errors.log',
primary_metric = 'AUC_weighted',
iteration_timeout_minutes = 10,
iterations = 5,
n_cross_validations = 10,
max_concurrent_iterations = 2, #change it based on number of worker nodes
verbosity = logging.INFO,
spark_context=sc, #databricks/spark related
training_data=training_data,
label_column_name=label)
```
## Train the Models
Call the `submit` method on the experiment object and pass the run configuration. Execution of local runs is synchronous. Depending on the data and the number of iterations this can run for a while.
```
local_run = experiment.submit(automl_config, show_output = True)
```
## Explore the Results
#### Portal URL for Monitoring Runs
The following will provide a link to the web interface to explore individual run details and status. In the future we might support output displayed in the notebook.
```
displayHTML("<a href={} target='_blank'>Azure Portal: {}</a>".format(local_run.get_portal_url(), local_run.id))
```
## Deploy
### Retrieve the Best Model
Below we select the best pipeline from our iterations. The `get_output` method on `automl_classifier` returns the best run and the fitted model for the last invocation. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*.
```
best_run, fitted_model = local_run.get_output()
```
### Test the Best Fitted Model
#### Load Test Data - you can split the dataset beforehand & pass Train dataset to AutoML and use Test dataset to evaluate the best model.
```
dataset_test = Dataset.Tabular.from_delimited_files(path='https://dprepdata.blob.core.windows.net/demo/crime0-test.csv')
df_test = dataset_test.to_pandas_dataframe()
df_test = df_test[pd.notnull(df_test['Primary Type'])]
y_test = df_test[['Primary Type']]
X_test = df_test.drop(['Primary Type', 'FBI Code'], axis=1)
```
#### Testing Our Best Fitted Model
We will try to predict digits and see how our model works. This is just an example to show you.
```
fitted_model.predict(X_test)
```

| github_jupyter |
**[Advanced SQL Home Page](https://www.kaggle.com/learn/advanced-sql)**
---
# Introduction
Here, you'll use different types of SQL **JOINs** to answer questions about the [Stack Overflow](https://www.kaggle.com/stackoverflow/stackoverflow) dataset.
Before you get started, run the following cell to set everything up.
```
# Set up feedback system
from learntools.core import binder
binder.bind(globals())
from learntools.sql_advanced.ex1 import *
print("Setup Complete")
```
The code cell below fetches the `posts_questions` table from the `stackoverflow` dataset. We also preview the first five rows of the table.
```
from google.cloud import bigquery
# Create a "Client" object
client = bigquery.Client()
# Construct a reference to the "stackoverflow" dataset
dataset_ref = client.dataset("stackoverflow", project="bigquery-public-data")
# API request - fetch the dataset
dataset = client.get_dataset(dataset_ref)
# Construct a reference to the "posts_questions" table
table_ref = dataset_ref.table("posts_questions")
# API request - fetch the table
table = client.get_table(table_ref)
# Preview the first five lines of the table
client.list_rows(table, max_results=5).to_dataframe()
```
We also take a look at the `posts_answers` table.
```
# Construct a reference to the "posts_answers" table
table_ref = dataset_ref.table("posts_answers")
# API request - fetch the table
table = client.get_table(table_ref)
# Preview the first five lines of the table
client.list_rows(table, max_results=5).to_dataframe()
```
You will work with both of these tables to answer the questions below.
# Exercises
### 1) How long does it take for questions to receive answers?
You're interested in exploring the data to have a better understanding of how long it generally takes for questions to receive answers. Armed with this knowledge, you plan to use this information to better design the order in which questions are presented to Stack Overflow users.
With this goal in mind, you write the query below, which focuses on questions asked in January 2018. It returns a table with two columns:
- `q_id` - the ID of the question
- `time_to_answer` - how long it took (in seconds) for the question to receive an answer
Run the query below (without changes), and take a look at the output.
```
first_query = """
SELECT q.id AS q_id,
MIN(TIMESTAMP_DIFF(a.creation_date, q.creation_date, SECOND)) as time_to_answer
FROM `bigquery-public-data.stackoverflow.posts_questions` AS q
INNER JOIN `bigquery-public-data.stackoverflow.posts_answers` AS a
ON q.id = a.parent_id
WHERE q.creation_date >= '2018-01-01' and q.creation_date < '2018-02-01'
GROUP BY q_id
ORDER BY time_to_answer
"""
first_result = client.query(first_query).result().to_dataframe()
print("Percentage of answered questions: %s%%" % \
(sum(first_result["time_to_answer"].notnull()) / len(first_result) * 100))
print("Number of questions:", len(first_result))
first_result.head()
```
You're surprised at the results and strongly suspect that something is wrong with your query. In particular,
- According to the query, 100% of the questions from January 2018 received an answer. But, you know that ~80% of the questions on the site usually receive an answer.
- The total number of questions is surprisingly low. You expected to see at least 150,000 questions represented in the table.
Given these observations, you think that the type of **JOIN** you have chosen has inadvertently excluded unanswered questions. Using the code cell below, can you figure out what type of **JOIN** to use to fix the problem so that the table includes unanswered questions?
**Note**: You need only amend the type of **JOIN** (i.e., **INNER**, **LEFT**, **RIGHT**, or **FULL**) to answer the question successfully.
```
# Your code here
correct_query = """
SELECT q.id AS q_id,
MIN(TIMESTAMP_DIFF(a.creation_date, q.creation_date, SECOND)) as time_to_answer
FROM `bigquery-public-data.stackoverflow.posts_questions` AS q
LEFT JOIN `bigquery-public-data.stackoverflow.posts_answers` AS a
ON q.id = a.parent_id
WHERE q.creation_date >= '2018-01-01' and q.creation_date < '2018-02-01'
GROUP BY q_id
ORDER BY time_to_answer
"""
# Check your answer
q_1.check()
# Run the query, and return a pandas DataFrame
correct_result = client.query(correct_query).result().to_dataframe()
print("Percentage of answered questions: %s%%" % \
(sum(correct_result["time_to_answer"].notnull()) / len(correct_result) * 100))
print("Number of questions:", len(correct_result))
# Lines below will give you a hint or solution code
#q_1.hint()
#q_1.solution()
```
### 2) Initial questions and answers, Part 1
You're interested in understanding the initial experiences that users typically have with the Stack Overflow website. Is it more common for users to first ask questions or provide answers? After signing up, how long does it take for users to first interact with the website? To explore this further, you draft the (partial) query in the code cell below.
The query returns a table with three columns:
- `owner_user_id` - the user ID
- `q_creation_date` - the first time the user asked a question
- `a_creation_date` - the first time the user contributed an answer
You want to keep track of users who have asked questions, but have yet to provide answers. And, your table should also include users who have answered questions, but have yet to pose their own questions.
With this in mind, please fill in the appropriate **JOIN** (i.e., **INNER**, **LEFT**, **RIGHT**, or **FULL**) to return the correct information.
**Note**: You need only fill in the appropriate **JOIN**. All other parts of the query should be left as-is. (You also don't need to write any additional code to run the query, since the `cbeck()` method will take care of this for you.)
To avoid returning too much data, we'll restrict our attention to questions and answers posed in January 2019. We'll amend the timeframe in Part 2 of this question to be more realistic!
```
# Your code here
q_and_a_query = """
SELECT q.owner_user_id AS owner_user_id,
MIN(q.creation_date) AS q_creation_date,
MIN(a.creation_date) AS a_creation_date
FROM `bigquery-public-data.stackoverflow.posts_questions` AS q
FULL JOIN `bigquery-public-data.stackoverflow.posts_answers` AS a
ON q.owner_user_id = a.owner_user_id
WHERE q.creation_date >= '2019-01-01' AND q.creation_date < '2019-02-01'
AND a.creation_date >= '2019-01-01' AND a.creation_date < '2019-02-01'
GROUP BY owner_user_id
"""
# Check your answer
q_2.check()
# Lines below will give you a hint or solution code
#q_2.hint()
#q_2.solution()
```
### 3) Initial questions and answers, Part 2
Now you'll address a more realistic (and complex!) scenario. To answer this question, you'll need to pull information from *three* different tables! This syntax very similar to the case when we have to join only two tables. For instance, consider the three tables below.

We can use two different **JOINs** to link together information from all three tables, in a single query.

With this in mind, say you're interested in understanding users who joined the site in January 2019. You want to track their activity on the site: when did they post their first questions and answers, if ever?
Write a query that returns the following columns:
- `id` - the IDs of all users who created Stack Overflow accounts in January 2019 (January 1, 2019, to January 31, 2019, inclusive)
- `q_creation_date` - the first time the user posted a question on the site; if the user has never posted a question, the value should be null
- `a_creation_date` - the first time the user posted a question on the site; if the user has never posted a question, the value should be null
Note that questions and answers posted after January 31, 2019, should still be included in the results. And, all users who joined the site in January 2019 should be included (even if they have never posted a question or provided an answer).
The query from the previous question should be a nice starting point to answering this question! You'll need to use the `posts_answers` and `posts_questions` tables. You'll also need to use the `users` table from the Stack Overflow dataset. The relevant columns from the `users` table are `id` (the ID of each user) and `creation_date` (when the user joined the Stack Overflow site, in DATETIME format).
```
# Your code here
three_tables_query = """
SELECT u.id AS id,
MIN(q.creation_date) AS q_creation_date,
MIN(a.creation_date) AS a_creation_date
FROM `bigquery-public-data.stackoverflow.users` AS u
LEFT JOIN `bigquery-public-data.stackoverflow.posts_questions` AS q
ON u.id = q.owner_user_id
LEFT JOIN `bigquery-public-data.stackoverflow.posts_answers` AS a
ON u.id = a.owner_user_id
WHERE u.creation_date >= '2019-01-01' AND u.creation_date < '2019-02-01'
GROUP BY id
"""
# Check your answer
q_3.check()
# Lines below will give you a hint or solution code
#q_3.hint()
#q_3.solution()
```
### 4) How many distinct users posted on January 1, 2019?
In the code cell below, write a query that returns a table with a single column:
- `owner_user_id` - the IDs of all users who posted at least one question or answer on January 1, 2019. Each user ID should appear at most once.
In the `posts_questions` (and `posts_answers`) tables, you can get the ID of the original poster from the `owner_user_id` column. Likewise, the date of the original posting can be found in the `creation_date` column.
In order for your answer to be marked correct, your query must use a **UNION**.
```
# Your code here
all_users_query = """
SELECT q.owner_user_id AS owner_user_id
FROM `bigquery-public-data.stackoverflow.posts_questions` AS q
WHERE EXTRACT(DATE FROM q.creation_date) = '2019-01-01'
UNION DISTINCT
SELECT a.owner_user_id AS owner_user_id
FROM `bigquery-public-data.stackoverflow.posts_answers` AS a
WHERE EXTRACT(DATE FROM a.creation_date) = '2019-01-01'
"""
# Check your answer
q_4.check()
# Lines below will give you a hint or solution code
#q_4.hint()
#q_4.solution()
```
# Keep going
Learn how to use **[analytic functions](https://www.kaggle.com/alexisbcook/analytic-functions)** to perform complex calculations with minimal SQL code.
---
**[Advanced SQL Home Page](https://www.kaggle.com/learn/advanced-sql)**
*Have questions or comments? Visit the [Learn Discussion forum](https://www.kaggle.com/learn-forum) to chat with other Learners.*
| github_jupyter |
```
from google.colab import drive
drive.mount('/content/drive')
path="/content/drive/MyDrive/Research/cods_comad_plots/SDC/dataset_2/"
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
from tqdm import tqdm
%matplotlib inline
from torch.utils.data import Dataset, DataLoader
import torch
import torchvision
import torch.nn as nn
import torch.optim as optim
from torch.nn import functional as F
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
print(device)
```
# Generate dataset
```
np.random.seed(12)
y = np.random.randint(0,10,5000)
idx= []
for i in range(10):
print(i,sum(y==i))
idx.append(y==i)
x = np.zeros((5000,2))
np.random.seed(12)
x[idx[0],:] = np.random.multivariate_normal(mean = [5,5],cov=[[0.1,0],[0,0.1]],size=sum(idx[0]))
x[idx[1],:] = np.random.multivariate_normal(mean = [-6,7],cov=[[0.1,0],[0,0.1]],size=sum(idx[1]))
x[idx[2],:] = np.random.multivariate_normal(mean = [-5,-4],cov=[[0.1,0],[0,0.1]],size=sum(idx[2]))
x[idx[3],:] = np.random.multivariate_normal(mean = [-1,0],cov=[[0.1,0],[0,0.1]],size=sum(idx[3]))
x[idx[4],:] = np.random.multivariate_normal(mean = [0,2],cov=[[0.1,0],[0,0.1]],size=sum(idx[4]))
x[idx[5],:] = np.random.multivariate_normal(mean = [1,0],cov=[[0.1,0],[0,0.1]],size=sum(idx[5]))
x[idx[6],:] = np.random.multivariate_normal(mean = [0,-1],cov=[[0.1,0],[0,0.1]],size=sum(idx[6]))
x[idx[7],:] = np.random.multivariate_normal(mean = [0,0],cov=[[0.1,0],[0,0.1]],size=sum(idx[7]))
x[idx[8],:] = np.random.multivariate_normal(mean = [-0.5,-0.5],cov=[[0.1,0],[0,0.1]],size=sum(idx[8]))
x[idx[9],:] = np.random.multivariate_normal(mean = [0.4,0.2],cov=[[0.1,0],[0,0.1]],size=sum(idx[9]))
x[idx[0]][0], x[idx[5]][5]
for i in range(10):
plt.scatter(x[idx[i],0],x[idx[i],1],label="class_"+str(i))
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
bg_idx = [ np.where(idx[3] == True)[0],
np.where(idx[4] == True)[0],
np.where(idx[5] == True)[0],
np.where(idx[6] == True)[0],
np.where(idx[7] == True)[0],
np.where(idx[8] == True)[0],
np.where(idx[9] == True)[0]]
bg_idx = np.concatenate(bg_idx, axis = 0)
bg_idx.shape
np.unique(bg_idx).shape
x = x - np.mean(x[bg_idx], axis = 0, keepdims = True)
np.mean(x[bg_idx], axis = 0, keepdims = True), np.mean(x, axis = 0, keepdims = True)
x = x/np.std(x[bg_idx], axis = 0, keepdims = True)
np.std(x[bg_idx], axis = 0, keepdims = True), np.std(x, axis = 0, keepdims = True)
for i in range(10):
plt.scatter(x[idx[i],0],x[idx[i],1],label="class_"+str(i))
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
foreground_classes = {'class_0','class_1', 'class_2'}
background_classes = {'class_3','class_4', 'class_5', 'class_6','class_7', 'class_8', 'class_9'}
fg_class = np.random.randint(0,3)
fg_idx = np.random.randint(0,9)
a = []
for i in range(9):
if i == fg_idx:
b = np.random.choice(np.where(idx[fg_class]==True)[0],size=1)
a.append(x[b])
print("foreground "+str(fg_class)+" present at " + str(fg_idx))
else:
bg_class = np.random.randint(3,10)
b = np.random.choice(np.where(idx[bg_class]==True)[0],size=1)
a.append(x[b])
print("background "+str(bg_class)+" present at " + str(i))
a = np.concatenate(a,axis=0)
print(a.shape)
print(fg_class , fg_idx)
a.shape
np.reshape(a,(18,1)) #not required
a=np.reshape(a,(9,2))
plt.imshow(a)
desired_num = 2000
mosaic_list_of_images =[]
mosaic_label = []
fore_idx=[]
for j in range(desired_num):
np.random.seed(j)
fg_class = np.random.randint(0,3)
fg_idx = np.random.randint(0,9)
a = []
for i in range(9):
if i == fg_idx:
b = np.random.choice(np.where(idx[fg_class]==True)[0],size=1)
a.append(x[b])
# print("foreground "+str(fg_class)+" present at " + str(fg_idx))
else:
bg_class = np.random.randint(3,10)
b = np.random.choice(np.where(idx[bg_class]==True)[0],size=1)
a.append(x[b])
# print("background "+str(bg_class)+" present at " + str(i))
a = np.concatenate(a,axis=0)
mosaic_list_of_images.append(a)
mosaic_label.append(fg_class)
fore_idx.append(fg_idx)
# mosaic_list_of_images = np.concatenate(mosaic_list_of_images,axis=1).T
len(mosaic_list_of_images), mosaic_list_of_images[0]
mosaic_list_of_images_reshaped = np.reshape(mosaic_list_of_images, (2000,9,2))
mean_train = np.mean(mosaic_list_of_images_reshaped[0:1000], axis=0, keepdims= True)
print(mean_train.shape, mean_train)
std_train = np.std(mosaic_list_of_images_reshaped[0:1000], axis=0, keepdims= True)
print(std_train.shape, std_train)
mosaic_list_of_images = ( mosaic_list_of_images_reshaped - mean_train ) / std_train
print(np.mean(mosaic_list_of_images[0:1000], axis=0, keepdims= True))
print(np.std(mosaic_list_of_images[0:1000], axis=0, keepdims= True))
print(np.mean(mosaic_list_of_images[1000:2000], axis=0, keepdims= True))
print(np.std(mosaic_list_of_images[1000:2000], axis=0, keepdims= True))
mosaic_list_of_images.shape
class MosaicDataset(Dataset):
"""MosaicDataset dataset."""
def __init__(self, mosaic_list_of_images, mosaic_label, fore_idx):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.mosaic = mosaic_list_of_images
self.label = mosaic_label
self.fore_idx = fore_idx
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.mosaic[idx] , self.label[idx], self.fore_idx[idx]
batch = 250
msd1 = MosaicDataset(mosaic_list_of_images[0:1000], mosaic_label[0:1000] , fore_idx[0:1000])
train_loader = DataLoader( msd1 ,batch_size= batch ,shuffle=True)
batch = 250
msd2 = MosaicDataset(mosaic_list_of_images[1000:2000], mosaic_label[1000:2000] , fore_idx[1000:2000])
test_loader = DataLoader( msd2 ,batch_size= batch ,shuffle=True)
class Focus(nn.Module):
def __init__(self):
super(Focus, self).__init__()
self.fc1 = nn.Linear(2, 50, bias=False)
self.fc2 = nn.Linear(50, 10, bias=False)
self.fc3 = nn.Linear(10, 1, bias=False)
torch.nn.init.xavier_normal_(self.fc1.weight)
torch.nn.init.xavier_normal_(self.fc2.weight)
torch.nn.init.xavier_normal_(self.fc3.weight)
def forward(self,z): #y is avg image #z batch of list of 9 images
y = torch.zeros([batch,2], dtype=torch.float64)
x = torch.zeros([batch,9],dtype=torch.float64)
y = y.to("cuda")
x = x.to("cuda")
# print(x.shape, z.shape)
for i in range(9):
# print(z[:,i].shape)
# print(self.helper(z[:,i])[:,0].shape)
x[:,i] = self.helper(z[:,i])[:,0]
# print(x.shape, z.shape)
x = F.softmax(x,dim=1)
# print(x.shape, z.shape)
# x1 = x[:,0]
# print(torch.mul(x[:,0],z[:,0]).shape)
for i in range(9):
# x1 = x[:,i]
y = y + torch.mul(x[:,i,None],z[:,i])
# print(x.shape, y.shape)
return x, y
def helper(self, x):
x = x.view(-1, 2)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = (self.fc3(x))
return x
class Classification(nn.Module):
def __init__(self):
super(Classification, self).__init__()
self.fc1 = nn.Linear(2, 50)
self.fc2 = nn.Linear(50, 10)
self.fc3 = nn.Linear(10, 3)
torch.nn.init.xavier_normal_(self.fc1.weight)
torch.nn.init.zeros_(self.fc1.bias)
torch.nn.init.xavier_normal_(self.fc2.weight)
torch.nn.init.zeros_(self.fc2.bias)
torch.nn.init.xavier_normal_(self.fc3.weight)
torch.nn.init.zeros_(self.fc3.bias)
def forward(self, x):
x = x.view(-1, 2)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = (self.fc3(x))
return x
torch.manual_seed(12)
focus_net = Focus().double()
focus_net = focus_net.to("cuda")
torch.manual_seed(12)
classify = Classification().double()
classify = classify.to("cuda")
focus_net.fc2.weight.shape,focus_net.fc3.weight.shape
focus_net.fc2.weight.data[5:,:] = focus_net.fc2.weight.data[:5,:] #torch.nn.Parameter(torch.tensor([last_layer]) )
focus_net.fc2.weight[:5,:], focus_net.fc2.weight[5:,:]
focus_net.fc3.weight.data[:,5:] = -focus_net.fc3.weight.data[:,:5] #torch.nn.Parameter(torch.tensor([last_layer]) )
focus_net.fc3.weight
focus_net = focus_net.double().to("cuda")
focus_net.helper( torch.randn((1,9,2)).double().to("cuda") )
focus_net.fc1.weight
classify.fc1.weight, classify.fc1.bias, classify.fc1.weight.shape
import torch.optim as optim
criterion = nn.CrossEntropyLoss()
optimizer_classify = optim.Adam(classify.parameters(), lr=0.01 ) #, momentum=0.9)
optimizer_focus = optim.Adam(focus_net.parameters(), lr=0.01 ) #, momentum=0.9)
col1=[]
col2=[]
col3=[]
col4=[]
col5=[]
col6=[]
col7=[]
col8=[]
col9=[]
col10=[]
col11=[]
col12=[]
col13=[]
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs = inputs.double()
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
# print(outputs.shape)
_, predicted = torch.max(outputs.data, 1)
# print(predicted.shape)
for j in range(labels.size(0)):
count += 1
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
# print(focus, fore_idx[j], predicted[j])
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 1000 train images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
print(count)
print("="*100)
col1.append(0)
col2.append(argmax_more_than_half)
col3.append(argmax_less_than_half)
col4.append(focus_true_pred_true)
col5.append(focus_false_pred_true)
col6.append(focus_true_pred_false)
col7.append(focus_false_pred_false)
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in test_loader:
inputs, labels , fore_idx = data
inputs = inputs.double()
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 1000 test images: %d %%' % (
100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
col8.append(argmax_more_than_half)
col9.append(argmax_less_than_half)
col10.append(focus_true_pred_true)
col11.append(focus_false_pred_true)
col12.append(focus_true_pred_false)
col13.append(focus_false_pred_false)
nos_epochs = 1000
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
for epoch in range(nos_epochs): # loop over the dataset multiple times
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
running_loss = 0.0
epoch_loss = []
cnt=0
iteration = desired_num // batch
#training data set
for i, data in enumerate(train_loader):
inputs , labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
inputs = inputs.double()
# zero the parameter gradients
optimizer_focus.zero_grad()
optimizer_classify.zero_grad()
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
# print(outputs)
# print(outputs.shape,labels.shape , torch.argmax(outputs, dim=1))
loss = criterion(outputs, labels)
loss.backward()
optimizer_focus.step()
optimizer_classify.step()
running_loss += loss.item()
mini = 3
if cnt % mini == mini-1: # print every 40 mini-batches
print('[%d, %5d] loss: %.3f' %(epoch + 1, cnt + 1, running_loss / mini))
epoch_loss.append(running_loss/mini)
running_loss = 0.0
cnt=cnt+1
if epoch % 5 == 0:
for j in range (batch):
focus = torch.argmax(alphas[j])
if(alphas[j][focus] >= 0.5):
argmax_more_than_half +=1
else:
argmax_less_than_half +=1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true +=1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false +=1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false +=1
if(np.mean(epoch_loss) <= 0.001):
break;
if epoch % 5 == 0:
col1.append(epoch + 1)
col2.append(argmax_more_than_half)
col3.append(argmax_less_than_half)
col4.append(focus_true_pred_true)
col5.append(focus_false_pred_true)
col6.append(focus_true_pred_false)
col7.append(focus_false_pred_false)
# print("="*20)
# print("Train FTPT : ", col4)
# print("Train FFPT : ", col5)
#************************************************************************
#testing data set
# focus_net.eval()
with torch.no_grad():
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
for data in test_loader:
inputs, labels , fore_idx = data
inputs = inputs.double()
inputs, labels = inputs.to("cuda"), labels.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range (batch):
focus = torch.argmax(alphas[j])
if(alphas[j][focus] >= 0.5):
argmax_more_than_half +=1
else:
argmax_less_than_half +=1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true +=1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false +=1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false +=1
col8.append(argmax_more_than_half)
col9.append(argmax_less_than_half)
col10.append(focus_true_pred_true)
col11.append(focus_false_pred_true)
col12.append(focus_true_pred_false)
col13.append(focus_false_pred_false)
# print("Test FTPT : ", col10)
# print("Test FFPT : ", col11)
# print("="*20)
print('Finished Training')
df_train = pd.DataFrame()
df_test = pd.DataFrame()
columns = ["epochs", "argmax > 0.5" ,"argmax < 0.5", "focus_true_pred_true", "focus_false_pred_true", "focus_true_pred_false", "focus_false_pred_false" ]
df_train[columns[0]] = col1
df_train[columns[1]] = col2
df_train[columns[2]] = col3
df_train[columns[3]] = col4
df_train[columns[4]] = col5
df_train[columns[5]] = col6
df_train[columns[6]] = col7
df_test[columns[0]] = col1
df_test[columns[1]] = col8
df_test[columns[2]] = col9
df_test[columns[3]] = col10
df_test[columns[4]] = col11
df_test[columns[5]] = col12
df_test[columns[6]] = col13
df_train
# plt.figure(12,12)
plt.plot(col1,col2, label='argmax > 0.5')
plt.plot(col1,col3, label='argmax < 0.5')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("training data")
plt.title("On Training set")
plt.show()
plt.plot(col1,col4, label ="focus_true_pred_true ")
plt.plot(col1,col5, label ="focus_false_pred_true ")
plt.plot(col1,col6, label ="focus_true_pred_false ")
plt.plot(col1,col7, label ="focus_false_pred_false ")
plt.title("On Training set")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("training data")
plt.show()
plt.figure(figsize=(6,5))
plt.plot(col1,np.array(col4)/10, label ="FTPT")
plt.plot(col1,np.array(col5)/10, label ="FFPT")
plt.plot(col1,np.array(col6)/10, label ="FTPF")
plt.plot(col1,np.array(col7)/10, label ="FFPF")
plt.title("Dataset2 - SDC On Train set")
plt.grid()
# plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.legend()
plt.xlabel("epochs", fontsize=14, fontweight = 'bold')
plt.ylabel("percentage train data", fontsize=14, fontweight = 'bold')
plt.savefig(path+"ds2_train.png", bbox_inches="tight")
plt.savefig(path+"ds2_train.pdf", bbox_inches="tight")
plt.savefig("ds2_train.png", bbox_inches="tight")
plt.savefig("ds2_train.pdf", bbox_inches="tight")
plt.show()
df_test
# plt.figure(12,12)
plt.plot(col1,col8, label='argmax > 0.5')
plt.plot(col1,col9, label='argmax < 0.5')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("Testing data")
plt.title("On Testing set")
plt.show()
plt.plot(col1,col10, label ="focus_true_pred_true ")
plt.plot(col1,col11, label ="focus_false_pred_true ")
plt.plot(col1,col12, label ="focus_true_pred_false ")
plt.plot(col1,col13, label ="focus_false_pred_false ")
plt.title("On Testing set")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("Testing data")
plt.show()
plt.figure(figsize=(6,5))
plt.plot(col1,np.array(col10)/10, label ="FTPT")
plt.plot(col1,np.array(col11)/10, label ="FFPT")
plt.plot(col1,np.array(col12)/10, label ="FTPF")
plt.plot(col1,np.array(col13)/10, label ="FFPF")
plt.title("Dataset2 - SDC On Test set")
plt.grid()
# plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.legend()
plt.xlabel("epochs", fontsize=14, fontweight = 'bold')
plt.ylabel("percentage test data", fontsize=14, fontweight = 'bold')
plt.savefig(path+"ds2_test.png", bbox_inches="tight")
plt.savefig(path+"ds2_test.pdf", bbox_inches="tight")
plt.savefig("ds2_test.png", bbox_inches="tight")
plt.savefig("ds2_test.pdf", bbox_inches="tight")
plt.show()
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs = inputs.double()
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 1000 train images: %d %%' % (
100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in test_loader:
inputs, labels , fore_idx = data
inputs = inputs.double()
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 1000 test images: %d %%' % (
100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
correct = 0
total = 0
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs = inputs.double()
inputs, labels = inputs.to("cuda"), labels.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 1000 train images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
correct = 0
total = 0
with torch.no_grad():
for data in test_loader:
inputs, labels , fore_idx = data
inputs = inputs.double()
inputs, labels = inputs.to("cuda"), labels.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 1000 test images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
```
| github_jupyter |
# Classifying Fashion-MNIST
Now it's your turn to build and train a neural network. You'll be using the [Fashion-MNIST dataset](https://github.com/zalandoresearch/fashion-mnist), a drop-in replacement for the MNIST dataset. MNIST is actually quite trivial with neural networks where you can easily achieve better than 97% accuracy. Fashion-MNIST is a set of 28x28 greyscale images of clothes. It's more complex than MNIST, so it's a better representation of the actual performance of your network, and a better representation of datasets you'll use in the real world.
<img src='assets/fashion-mnist-sprite.png' width=500px>
In this notebook, you'll build your own neural network. For the most part, you could just copy and paste the code from Part 3, but you wouldn't be learning. It's important for you to write the code yourself and get it to work. Feel free to consult the previous notebook though as you work through this.
First off, let's load the dataset through torchvision.
```
import torch
from torchvision import datasets, transforms
import helper
# Define a transform to normalize the data
transform = transforms.Compose([transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
# Download and load the training data
trainset = datasets.FashionMNIST('F_MNIST_data/', download=True, train=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=64, shuffle=True)
# Download and load the test data
testset = datasets.FashionMNIST('F_MNIST_data/', download=True, train=False, transform=transform)
testloader = torch.utils.data.DataLoader(testset, batch_size=64, shuffle=True)
```
Here we can see one of the images.
```
image, label = next(iter(trainloader))
helper.imshow(image[0,:]);
```
With the data loaded, it's time to import the necessary packages.
```
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import matplotlib.pyplot as plt
import numpy as np
import time
import torch
from torch import nn
from torch import optim
import torch.nn.functional as F
from torchvision import datasets, transforms
import helper
```
## Building the network
Here you should define your network. As with MNIST, each image is 28x28 which is a total of 784 pixels, and there are 10 classes. You should include at least one hidden layer. We suggest you use ReLU activations for the layers and to return the logits from the forward pass. It's up to you how many layers you add and the size of those layers.
```
# TODO: Define your network architecture here
from collections import OrderedDict
input_units = 784
hidden_units = [600, 400, 100]
output_units = 10
model = nn.Sequential(OrderedDict([
('fc1', nn.Linear(input_units, hidden_units[0])),
('relu1', nn.ReLU()),
('fc2', nn.Linear(hidden_units[0], hidden_units[1])),
('relu2', nn.ReLU()),
('fc3', nn.Linear(hidden_units[1], hidden_units[2])),
('relu3', nn.ReLU()),
('logits', nn.Linear(hidden_units[2], output_units))]))
```
# Train the network
Now you should create your network and train it. First you'll want to define [the criterion](http://pytorch.org/docs/master/nn.html#loss-functions) ( something like `nn.CrossEntropyLoss`) and [the optimizer](http://pytorch.org/docs/master/optim.html) (typically `optim.SGD` or `optim.Adam`).
Then write the training code. Remember the training pass is a fairly straightforward process:
* Make a forward pass through the network to get the logits
* Use the logits to calculate the loss
* Perform a backward pass through the network with `loss.backward()` to calculate the gradients
* Take a step with the optimizer to update the weights
By adjusting the hyperparameters (hidden units, learning rate, etc), you should be able to get the training loss below 0.4.
```
# TODO: Create the network, define the criterion and optimizer
# model has already been created !
criterion = nn.CrossEntropyLoss()
optimiser = torch.optim.SGD(model.parameters(), lr=0.003)
# From Part 5 - Inference and Validation
#Implement a function for the validation pass
def validation(model, testloader, criterion):
test_loss = 0
accuracy = 0
for images, labels in testloader:
images.resize_(images.shape[0], 784)
output = model.forward(images)
test_loss += criterion(output, labels).item()
ps = torch.exp(output)
equality = (labels.data == ps.max(dim=1)[1])
accuracy += equality.type(torch.FloatTensor).mean()
return test_loss, accuracy
# TODO: Train the network here
epoch = 3
print_every = 40
steps = 0
total_loss = 0
for e in range(epoch):
for images,labels in iter(trainloader):
steps+=1
input_images = images.resize(images.size()[0],784)
optimiser.zero_grad()
outputs=model.forward(input_images)
loss = criterion(outputs,labels)
loss.backward()
optimiser.step()
total_loss+= loss.item()
if steps%print_every ==0:
print('item loss is {}. Avg loss is {}'.format(loss.item(), total_loss/print_every))
test_loss, accuracy = validation(model, testloader, criterion)
print ("test_loss {} accuracy {}".format(test_loss/len(testloader), accuracy/len(testloader)))
steps=0
total_loss=0
correct_values = {
0:'T-shirt/top',
1:'Trouser',
2:'Pullover',
3:'Dress',
4:'Coat',
5:'Sandal',
6:'Shirt',
7:'Sneaker',
8:'Bag',
9:'Ankle boot' }
# Test out your network!
dataiter = iter(testloader)
images, labels = dataiter.next()
img = images[0]
# Convert 2D image to 1D vector
img = img.resize_(1, 784)
# TODO: Calculate the class probabilities (softmax) for img
ps = F.softmax(model.forward(img))
# Plot the image and probabilities
helper.view_classify(img.resize_(1, 28, 28), ps, version='Fashion')
correct_values[labels[0].item()]
```
Now that your network is trained, you'll want to save it to disk so you can load it later instead of training it again. Obviously, it's impractical to train a network every time you need one. In practice, you'll train it once, save the model, then reload it for further training or making predictions. In the next part, I'll show you how to save and load trained models.
| github_jupyter |
# Making a particle distribution.
We are going to look at two classes in particula, for this notebook. The first is `particle_distribution` and the second (`dynamic_step`) we'll use to move forward in time.
So lets get those imported along with numpy and plotting.
```
# from particula import u
from particula import particle, dynamic_step
import numpy as np
import matplotlib
```
## ParticleDistribution
Lets just call this and see what we get.
```
default_distribution = particle.ParticleDistribution()
print("Radii")
default_distribution.pre_radius()[0:10]
```
At every radius listed in `_.pre_radius()` is a probability of the particles at that radius. Thus, we have probability distribution function (PDF) of particles.
```
print("Particles PFD given radii.")
default_distribution.pre_distribution()[0:10]
```
##Plot of particle PDF
Let's see what this particle PDF looks like now.
Note using `_.m` returns just the magnitude and drops the units.
```
fig, ax= matplotlib.pyplot.subplots(1,2,figsize=[14,6])
#linear scale
ax[0].plot(default_distribution.pre_radius().m, default_distribution.pre_distribution().m)
ax[0].set_ylabel(f"Number, {default_distribution.pre_distribution().u}")
ax[0].set_xlabel(f"Radius, {default_distribution.pre_radius().u}")
ax[0].grid(True, alpha=0.5)
#log x
ax[1].semilogx(default_distribution.pre_radius().m, default_distribution.pre_distribution().m)
ax[1].set_ylabel(f"Number, {default_distribution.pre_distribution().u}")
ax[1].set_xlabel(f"Radius, {default_distribution.pre_radius().u}")
ax[1].grid(True, alpha=0.5)
```
## Summary statistics of a particle PDF.
As the keen eyed reader may have notice this is not a typical PDF and the maximunm is very large ($1.75*10^{18}$). Lets define some diagnostic functions and see what we get for the integral area.
```
def pdf_total(radius, pdf_distribution):
return np.trapz(y=pdf_distribution, x=radius)
def pdf_volume_total(radius, pdf_distribution):
return np.trapz(y=pdf_distribution* 4/3 * np.pi * radius**3, x=radius)
print(f'Total number of the PDF, {pdf_total(default_distribution.pre_radius(), default_distribution.pre_distribution())}')
print(f'Total volume of the PDF, {pdf_volume_total(default_distribution.pre_radius(), default_distribution.pre_distribution())}')
```
* The integral area of the particle PDF is the total number of the distribution, $0.99*10^{12}~\#/m^3$.
* Converting to volume PDF, the integral is the total volume of the particle distribution, $5*10^{-10}~m^3/m^3$.
That was defaults but, those might not work for you case so you can change the distribution properties using keyword argument.
Note the line spacing is how the bin widths are space, for numerical efficiently in the coagulation calculation `linspace` is preferred over `logspace`.
For a complete list of the `kwargs` you can look at the XXX document.
```
# around here, just define a kwargs
fine_mode = {
"mode": 150e-9, # 200 nm median
"nbins": 1000, # 1000 bins
"nparticles": 1e8, # 1e4 #
"volume": 1e-6, # per 1e-6 m^3 (or 1 cc)
"gsigma": 1.25, # relatively narrow
"spacing": "linspace", # bin spacing,
}
coarse_mode = {
"mode": 2000e-9, # 200 nm median
"nbins": 1000, # 1000 bins
"nparticles": 1e5, # 1e4 #
"volume": 1e-6, # per 1e-6 m^3 (or 1 cc)
"gsigma": 1.4, # relatively narrow
"spacing": "linspace", # bin spacing,
}
multi_mode = {
"mode": [5000e-9, 50e-9, 300e-9], # 200 nm median
"nbins": 1000, # 1000 bins
"nparticles": [1e6,1e9,1e7], # 1e4 #
"volume": 1e-6, # per 1e-6 m^3 (or 1 cc)
"gsigma": [1.5,1.2,1.5], # relatively narrow
"spacing": "logspace", # bin spacing,
}
fine_mode_dist = particle.ParticleDistribution(**fine_mode) # pass the kwargs using ** prefix
coarse_mode_dist = particle.ParticleDistribution(**coarse_mode)
multi_mode = particle.ParticleDistribution(**multi_mode)
print(f'Total number of the PDF, {pdf_total(fine_mode_dist.pre_radius(), fine_mode_dist.pre_distribution())}')
print(f'Total volume of the PDF, {pdf_volume_total(fine_mode_dist.pre_radius(), fine_mode_dist.pre_distribution())}')
print(f'Total volume of the PDF, {pdf_volume_total(coarse_mode_dist.pre_radius(), coarse_mode_dist.pre_distribution())}')
print(f'Total volume of the PDF, {pdf_volume_total(multi_mode.pre_radius(), multi_mode.pre_distribution())}')
```
Let's see what we just made.
```
fig, ax= matplotlib.pyplot.subplots(1,2,figsize=[14,6])
ax[0].plot(fine_mode_dist.pre_radius().m, fine_mode_dist.pre_distribution().m)
ax[0].plot(coarse_mode_dist.pre_radius().m, coarse_mode_dist.pre_distribution().m)
ax[0].set_ylabel(f"Number, {fine_mode_dist.pre_distribution().u}")
ax[0].set_xlabel(f"Radius, {fine_mode_dist.pre_radius().u}")
ax[0].grid(True, alpha=0.5)
ax[1].loglog(fine_mode_dist.pre_radius().m, fine_mode_dist.pre_distribution().m)
ax[1].loglog(coarse_mode_dist.pre_radius().m, coarse_mode_dist.pre_distribution().m)
ax[1].set_ylabel(f"Number, {coarse_mode_dist.pre_distribution().u}")
ax[1].set_xlabel(f"Radius, {coarse_mode_dist.pre_radius().u}")
ax[1].grid(True, alpha=0.5)
fig, ax= matplotlib.pyplot.subplots(1,2,figsize=[14,6])
ax[0].plot(multi_mode.pre_radius().m, multi_mode.pre_distribution().m)
ax[0].set_ylabel(f"Number, {multi_mode.pre_distribution().u}")
ax[0].set_xlabel(f"Radius, {multi_mode.pre_radius().u}")
ax[0].grid(True, alpha=0.5)
ax[1].loglog(multi_mode.pre_radius().m, multi_mode.pre_distribution().m)
ax[1].set_ylabel(f"Number, {multi_mode.pre_distribution().u}")
ax[1].set_xlabel(f"Radius, {multi_mode.pre_radius().u}")
ax[1].grid(True, alpha=0.5)
```
## Coagulation Kernel
Now with particle number distribution we can calculated the coagulation rate, both the loss and gain in particle number.
```
coarse_coag_loss = dynamic_step.DynamicStep(**coarse_mode).coag_loss()
coarse_coag_gain = dynamic_step.DynamicStep(**coarse_mode).coag_gain()
```
Then plotting the results, we see that smaller particles are lost and larger particles gain number
```
fig, ax= matplotlib.pyplot.subplots(1,1,figsize=[9,6])
# (2*(peak of original distribution)**3)**(1/3)
ax.semilogx(coarse_mode_dist.pre_radius().m, -coarse_coag_loss.m, '-r', label='particles Lost')
ax.semilogx(coarse_mode_dist.pre_radius().m, coarse_coag_gain.m, '-b', label='particles Gained')
ax.semilogx(coarse_mode_dist.pre_radius().m, coarse_coag_gain.m-coarse_coag_loss.m, '--k', label= 'Net change')
ax.legend()
ax.set_xlabel(f"radius, {coarse_mode_dist.pre_radius().u}")
ax.set_ylabel(f"coagulation rates, {coarse_coag_gain.u}")
ax.grid(True, alpha=0.5)
```
# Time Steps
Now that we can make a distribution and calculate coagulation rates, lets move that forward in time.
```
from particula.util.coagulation_rate import CoagulationRate
from particula.util.dimensionless_coagulation import full_coag
from particula.util.input_handling import in_time
```
We can do this in a simple way by using a for loop to march through the time steps.
```
# Time steps
simple_dic_kwargs = {
"mode": 200e-9, # 200 nm median
"nbins": 500, # 1000 bins
"nparticles": 1e6, # 1e4 #
"volume": 1e-6, # per 1e-6 m^3 (or 1 cc)
"gsigma": 1.5, # relatively narrow
}
particle_dist2 = particle.ParticleDistribution(**simple_dic_kwargs) # pass the kwargs using ** prefix
#inital distribution
p_distribution_0 = particle_dist2.pre_distribution()
p_radius = particle_dist2.pre_radius()
coag_kernel = full_coag(radius=p_radius)
p_distribution_i = p_distribution_0
time_interval = in_time(10)
time_array = np.arange(0, 1000, time_interval.m)
distribution_time = np.zeros([len(time_array), len(p_distribution_0)])
for i, dpa in enumerate(time_array):
if i>0:
# calculate coagulations
coag_gain_i = CoagulationRate(p_distribution_i, p_radius, coag_kernel).coag_gain()
coag_loss_i = CoagulationRate(p_distribution_i, p_radius, coag_kernel).coag_loss()
net_change = (coag_gain_i-coag_loss_i)*time_interval
p_distribution_i = p_distribution_i+net_change
distribution_time[i,:] = p_distribution_i.m
```
## For loop Graph
```
fig, ax= matplotlib.pyplot.subplots(1,1,figsize=[9,6])
radius = p_radius.m
ax.semilogx(radius, p_distribution_0.m, '-b', label='Inital')
ax.semilogx(radius, distribution_time[49,:], '--', label='t=50')
ax.semilogx(radius, distribution_time[-1,:], '-r', label='t=end')
ax.legend()
ax.set_ylabel(f"Number, {particle_dist2.pre_distribution().u}")
ax.set_xlabel(f"Radius, {particle_dist2.pre_radius().u}")
ax.grid(True, alpha=0.5)
```
## ODE solver
Instead of prescribing the time steps, which can make the problem solution unstable, lets let the code handle that with an ODE solver.
Lets import that solver class
```
from particula.util.simple_solver import SimpleSolver
```
Then let us run the same dynamic problem again
```
simple_dic_kwargs = {
"mode": 200e-9, # 200 nm median
"nbins": 500, # 1000 bins
"nparticles": 1e6, # 1e4 #
"volume": 1e-6, # per 1e-6 m^3 (or 1 cc)
"gsigma": 1.5, # relatively narrow
}
particle_dist2 = particle.ParticleDistribution(**simple_dic_kwargs) # pass the kwargs using ** prefix
#inital distribution coag kernel
coag_kernel = full_coag(radius=particle_dist2.pre_radius())
time_array = np.arange(0, 1000, 10)
#setup the inital state of the distribution
problem = {
"distribution": particle_dist2.pre_distribution(),
"radius": particle_dist2.pre_radius(),
"kernel": coag_kernel,
"tspan": time_array
}
#call the solver
solution = SimpleSolver(**problem).solution()
#plot
fig, ax= matplotlib.pyplot.subplots(1,1,figsize=[9,6])
radius = p_radius.m
ax.semilogx(radius, particle_dist2.pre_distribution().m, '-b', label='Inital')
ax.semilogx(radius, solution.m[49,:], '--', label='t=50')
ax.semilogx(radius, solution.m[-1,:], '-r', label='t=end')
ax.legend()
ax.set_ylabel(f"Number, {particle_dist2.pre_distribution().u}")
ax.set_xlabel(f"Radius, {particle_dist2.pre_radius().u}")
ax.grid(True, alpha=0.5)
```
## summary on stepping
As we walked through, using the ODE solver is quite a nice way to get to the answer, without figuring out what time-step you might need.
# Fine + Coarse Mode
Usually in the ambient air there are at least two modes of aerosol particles. Fine mode (150 nm mode) generated from atmospheric chemistry and combustion emissions. Coarse mode (10 microns mode) generated from windblown dust, sea salt, or wildfire ash.
So lets make a multi-modal distribution an progress that forward in time.
## combined distributions
As we saw in the mulit-mode distribution call above, you can use keywards to get multiple distributions. Let's see how those fine and coarse mode distributions interact.
```
ambient_dist = {
"mode": [100e-9, 2000e-9], # 100 nm and 2000 nm
"nbins": 250, # cut the bins down for speed
"nparticles": [1e7, 1e4], # 1e4 #
"volume": 1e-6, # per 1e-6 m^3 (or 1 cc)
"gsigma": [1.25,1.4], # relatively narrow
"spacing": "logspace", # bin spacing,
}
ambient_dist = particle.ParticleDistribution(**ambient_dist)
#inital distribution coag kernel
coag_kernel = full_coag(radius=ambient_dist.pre_radius())
time_array = np.arange(0, 30*60, 1)
#setup the inital state of the distribution
problem = {
"distribution": ambient_dist.pre_distribution(),
"radius": ambient_dist.pre_radius(),
"kernel": coag_kernel,
"tspan": time_array
}
problem2 = {
"distribution": ambient_dist.pre_distribution(),
"radius": ambient_dist.pre_radius(),
"kernel": coag_kernel*10, # 10x increase to approximate soot
"tspan": time_array
}
#call the solver
solution = SimpleSolver(**problem).solution()
solution2 = SimpleSolver(**problem2).solution()
```
So we ran the simulation, and lets plot a couple slices for each. Solution 2 is an 10x increase in the coagulation kernel, which is a rough approximation for the coagulation enhancement due to a chain-like soot structure over the base spherical particle model.
```
#plot
fig, ax= matplotlib.pyplot.subplots(1,1,figsize=[10,10])
radius = ambient_dist.pre_radius().to('nm').m
ax.loglog(radius, ambient_dist.pre_distribution().m, '-k', label='Inital')
ax.loglog(radius, solution.m[29,:], '--b', label='t=30 sec spheres')
ax.loglog(radius, solution.m[-1,:], '-.b', label='t=30 min spheres')
ax.loglog(radius, solution2.m[29,:], '--r', label='t=30 sec soot')
ax.loglog(radius, solution2.m[-1,:], '-.r', label='t=30 min soot')
ax.legend()
ax.set_ylim(bottom=1e11)
ax.set_xlim(left=40, right= 10000)
ax.set_ylabel(f"Number, {ambient_dist.pre_distribution().u}")
ax.set_xlabel(f"Radius, {ambient_dist.pre_radius().u}")
ax.grid(True, which='both', alpha=0.5)
```
## Number vs Time
Now lets plot the integrated number vs time. We should see the fine mode decrease, and the coarse mode increase a little depending on how fast the coagulational growth is moving the distribution.
```
radius = ambient_dist.pre_radius().m
radius_withunits= ambient_dist.pre_radius()
split = 600e-9 # fine to coarse cut off we are using.
split_index = np.argmin(np.abs(radius-split))
print(split_index)
fine_mode = np.zeros_like(time_array)
coarse_mode = np.zeros_like(time_array)
fine_mode2 = np.zeros_like(time_array)
coarse_mode2 = np.zeros_like(time_array)
fine_mode_vol = np.zeros_like(time_array)
coarse_mode_vol = np.zeros_like(time_array)
for i, value in enumerate(time_array):
fine_mode[i] = pdf_total(radius[0:split_index], solution.m[i, 0:split_index] )
coarse_mode[i] = pdf_total(radius[split_index:-1], solution.m[i, split_index:-1] )
fine_mode2[i] = pdf_total(radius[0:split_index], solution2.m[i, 0:split_index] )
coarse_mode2[i] = pdf_total(radius[split_index:-1], solution2.m[i, split_index:-1] )
fine_mode_vol[i] = pdf_volume_total(radius_withunits[0:split_index], solution[i, 0:split_index] )
coarse_mode_vol[i] = pdf_volume_total(radius_withunits[split_index:-1], solution[i, split_index:-1] )
fig, ax= matplotlib.pyplot.subplots(1,1,figsize=[10,10])
ax.semilogy(time_array, fine_mode, '-b', label='fine sphere')
ax.semilogy(time_array, coarse_mode, '--b', label='coarse sphere')
ax.semilogy(time_array, fine_mode2, '-r', label='fine soot')
ax.semilogy(time_array, coarse_mode2, '--r', label='coarse soot')
ax.legend()
ax.set_xlim(left=0)
ax.set_ylabel(f"number, #/m3 ")
ax.set_xlabel(f"time, ")
ax.grid(True, which='both', alpha=0.5)
fig.savefig('coagulation_number.png')
fig.savefig('coagulation_number.pdf')
```
#Summary
So we covered how to make a distribution, and how to progress a distribution forward in time with coagulation. This is just one of the three key processes (nucleation, condensation, and coagulation) happening to particles in the atmosphere; all of which can be simulated with `Particula`!
| github_jupyter |
## Geomorphometry I: Terrain Modeling (data in ASCII files)
Resources: [
Download the ASCI (x,y,z) lidar bare earth data
[lid_be_pts.txt](data/lid_be_pts.txt)
Download the ASCI (x,y,z) lidar multiple return data
[lid_mr_pts.txt](data/lid_mr_pts.txt)
### Start GRASS GIS
Start GRASS - click on GRASS icon or type
```
# This is a quick introduction into Jupyter Notebook.
# Python code can be executed like this:
a = 6
b = 7
c = a * b
print("Answer is", c)
# Python code can be mixed with command line code (Bash).
# It is enough just to prefix the command line with an exclamation mark:
!echo "Answer is $c"
# Use Shift+Enter to execute this cell. The result is below.
# a proper directory is already set, download files
import urllib.request
urllib.request.urlretrieve("http://ncsu-geoforall-lab.github.io/geospatial-modeling-course/grass/data/lid_be_pts.txt", "lid_be_pts.txt")
urllib.request.urlretrieve("http://ncsu-geoforall-lab.github.io/geospatial-modeling-course/grass/data/lid_mr_pts.txt", "lid_mr_pts.txt")
urllib.request.urlretrieve("http://ncsu-geoforall-lab.github.io/geospatial-modeling-course/grass/data/lid_be_pts.txt", "lid_be_pts.txt")
import os
import sys
import subprocess
from IPython.display import Image
# create GRASS GIS runtime environment
gisbase = subprocess.check_output(["grass", "--config", "path"], text=True).strip()
os.environ['GISBASE'] = gisbase
sys.path.append(os.path.join(gisbase, "etc", "python"))
# do GRASS GIS imports
import grass.script as gs
import grass.script.setup as gsetup
# set GRASS GIS session data
rcfile = gsetup.init(gisbase, "/home/jovyan/grassdata", "nc_spm_08_grass7", "user1")
# default font displays
os.environ['GRASS_FONT'] = 'sans'
# overwrite existing maps
os.environ['GRASS_OVERWRITE'] = '1'
gs.set_raise_on_error(True)
gs.set_capture_stderr(True)
# set display modules to render into a file (named map.png by default)
os.environ['GRASS_RENDER_IMMEDIATE'] = 'cairo'
os.environ['GRASS_RENDER_FILE_READ'] = 'TRUE'
os.environ['GRASS_LEGEND_FILE'] = 'legend.txt'
```
In startup pannel set GRASS GIS Database Directory to path to datasets,
for example on MS Windows, `C:\Users\myname\grassdata`.
For GRASS Location select nc_spm_08_grass7 (North Carolina, State Plane, meters) and
for GRASS Mapset create a new mapset (called e.g. HW_terrain_modeling).
Click Start GRASS session.
Change working directory:
_Settings_ > _GRASS working environment_ > _Change working directory_ > select/create any directory
or type `cd` (stands for change directory) into the GUI
_Console_ and hit Enter:
Download all files (see above)
to the selected directory. Now you can use the commands from the assignment requiring the file
without the need to specify the full path to the file.
### Bare earth data
Analyze bare earth and multiple return lidar data properties.
First, download the ASCI (x,y,z) lidar data
[lid_be_pts.txt](data/lid_be_pts.txt)
and then compute a raster map representing number of points
per 2m and 6m grid cell to assess the point density.
If you are unsure about the current working directory and where your
text file with point is, run _r.in.xyz_ from GUI
to find the path to the external lid_be_pts.txt.
Do not forget to zoom to computational region
to check its extent and see the resulting data.
You can use horizontal legends by resizing legend into wide short
rectangle or by using at=6,10,2,30 in the command line.
You can resize the _Map Display_ to create white space above
and below the raster map image.
```
gs.parse_command('g.region', region="rural_1m", res="2", flags='pg')
gs.run_command('r.in.xyz', input="lid_be_pts.txt", output="lid_be_binn2m", method="n")
gs.run_command('r.in.xyz', input="lid_mr_pts.txt", output="lid_mr_binn2m", method="n")
gs.run_command('d.erase')
gs.run_command('d.rast', map="lid_mr_binn2m")
gs.run_command('d.legend', raster="lid_mr_binn2m", at="2,20,2,5")
Image(filename="map.png")
print(gs.read_command('r.report', map="lid_mr_binn2m", unit="p"))
gs.parse_command('r.univar', map="lid_mr_binn2m", flags='g')
gs.run_command('d.rast', map="lid_be_binn2m")
gs.run_command('d.legend', raster="lid_be_binn2m", at="2,20,2,5")
print(gs.read_command('r.report', map="lid_be_binn2m", unit="p"))
gs.parse_command('r.univar', map="lid_be_binn2m", flags='g')
Image(filename="map.png")
```
### Range of coordinates at lower resolution
Patch and overlay planimetry to see that there are
no points in areas with buildings:
```
gs.run_command('v.patch', input="P079214,P079215,P079218,P079219", out="planimetry_rural")
gs.run_command('d.vect', map="planimetry_rural")
Image(filename="map.png")
```
Decrease resolution and try the above steps with
the _r.in.xyz_ module again.
Again, run it from GUI, define full path, or manage your current
working directory.
```
gs.parse_command('g.region', region="rural_1m", res="6", flags='apg')
gs.run_command('r.in.xyz', input="lid_be_pts.txt", out="lid_be_binn6m", meth="n")
gs.run_command('d.erase')
gs.run_command('d.rast', map="lid_be_binn6m")
gs.run_command('d.legend', raster="lid_be_binn6m", at="2,20,2,5")
Image(filename="map.png")
print(gs.read_command('r.report', map="lid_be_binn6m", unit="p"))
gs.parse_command('r.univar', map="lid_be_binn6m", flags='g')
Image(filename="map.png")
```
Compute a raster map representing mean elevation for each 6m cell.
It will still have a few holes.
```
gs.run_command('r.in.xyz', input="lid_be_pts.txt", out="lid_be_binmean6m", meth="mean")
gs.run_command('r.colors', map="lid_be_binmean6m", color="elevation")
gs.run_command('d.rast', map="lid_be_binmean6m")
gs.run_command('d.legend', raster="lid_be_binmean6m", at="2,40,2,5")
gs.run_command('r.in.xyz', input="lid_mr_pts.txt", out="lid_mr_binmean6m", meth="mean")
gs.run_command('r.colors', map="lid_mr_binmean6m", co="elevation")
gs.run_command('d.rast', map="lid_mr_binmean6m")
gs.run_command('d.legend', raster="lid_mr_binmean6m", at="2,40,2,5")
Image(filename="map.png")
```
Compute range:
```
gs.run_command('r.in.xyz', input="lid_be_pts.txt", out="lid_be_binrange6m", meth="range")
gs.run_command('r.in.xyz', input="lid_mr_pts.txt", out="lid_mr_binrange6m", meth="range")
gs.run_command('d.erase')
gs.run_command('d.rast', map="lid_be_binrange6m")
gs.run_command('d.legend', raster="lid_be_binrange6m", at="2,40,2,5")
gs.run_command('d.rast', map="lid_mr_binrange6m")
gs.run_command('d.legend', raster="lid_mr_binrange6m", at="2,40,2,5")
Image(filename="map.png")
```
Identify the features that are associated with large range values.
Display with vector data.
```
gs.run_command('d.vect', map="planimetry_rural")
gs.run_command('d.vect', map="lakes", type="boundary", co="violet")
gs.run_command('d.vect', map="streams", co="blue")
Image(filename="map.png")
```
Display only the high values of range (0.5-20m) overlayed over orthophoto.
What landcover is associated with large range in multiple return data?
Which landscape features may be lost at 6m resolution?
```
gs.parse_command('g.region', region="rural_1m", flags='pg')
```
Do not forget to zoom/set the display to computational region
to display only selected interval of values in GUI.
Add raster > Required tab, select lid_be_binrange6m
In Selection tab, set List of values to be displayed to 0.5-20.
```
gs.run_command('d.erase')
gs.run_command('d.rast', map="ortho_2001_t792_1m")
gs.run_command('d.rast', map="lid_be_binrange6m", val="0.5-20.")
gs.run_command('d.erase')
gs.run_command('d.rast', map="ortho_2001_t792_1m")
gs.run_command('d.rast', map="lid_mr_binrange6m", val="0.5-20.")
Image(filename="map.png")
```
### Interpolation
We now know how dense the data are and what is the range within cell.
If we need 1m resolution DEM for our application
this analysis tells us that we need to interpolate.
Import the ascii lidar data as vector points.
Import only points in the rural area without building a table (-t flag in tab Points),
number of column used as z is 3 (third column),
and using z-coordinate for elevation (Tab Optional, Create 3D vector map).
We assume that the txt file is in your current working directory.
```
gs.parse_command('g.region', region="rural_1m", flags='pg')
gs.run_command('v.in.ascii', input="lid_be_pts.txt", out="elev_lid_bepts", z="3", flags='ztr')
```
Display bare ground and multiple return points over orthophoto:
```
gs.run_command('d.erase')
gs.run_command('d.rast', map="ortho_2001_t792_1m")
Image(filename="map.png")
```
Display our imported points:
```
gs.run_command('d.vect', map="elev_lid_bepts", size="2", col="red")
Image(filename="map.png")
```
Display points available in the data set
(elev_lid_bepts and elev_lid792_bepts should be identical):
```
gs.run_command('d.vect', map="elev_lidrural_mrpts", size="4", col="0:100:0")
gs.run_command('d.vect', map="elev_lid792_bepts", size="2", col="yellow")
Image(filename="map.png")
```
Extract first return to get points for DSM.
Interpolate DEM and DSM (more about interpolation in the interpolation assignment).
We use default parameters except for number of points used for
segmentation and interpolation - segmax and npmin
and higher tension for multiple return data:
```
gs.parse_command('g.region', region="rural_1m", flags='pg')
gs.run_command('v.extract', input="elev_lidrural_mrpts", out="elev_lidrur_first", type="point", where="Return=1")
gs.run_command('v.surf.rst', input="elev_lid792_bepts", elevation="elev_lidrural_1m", npmin="120", segmax="25")
gs.run_command('v.surf.rst', input="elev_lidrur_first", elevation="elev_lidrurfirst_1m", npmin="120", segmax="25", tension="100", smooth="0.5")
gs.run_command('d.erase')
gs.run_command('d.rast', map="elev_lidrural_1m")
gs.run_command('d.rast', map="elev_lidrurfirst_1m")
Image(filename="map.png")
```
Remove all layers except for elev_lidrural_1m and elev_lidrurfirst_1m and
switch to 3D view with cutting planes to compare the bare earth and terrain surface.
Make sure fine resolution is set to 1 for all surfaces.
Assign each surface constant color, add constant plane z=90 for reference.
Create crossections using cutting plane,
shade the crossection using the color by bottom surface option.
Save image for report.
If you don't remember this, see screen capture video for
[GRASS GIS 3D view](https://youtu.be/xo_jJHgtbR4).
### Multiple returns
Find out where we have multiple returns:
```
gs.parse_command('g.region', region="rural_1m", flags='pg')
gs.run_command('d.erase')
gs.run_command('d.rast', map="ortho_2001_t792_1m")
Image(filename="map.png")
```
Condition for subset in GUI:
Add vector > Selection > type return=1 into WHERE condition of SQL statement.
You need to add the points 4 times to create a map that will have
all sets of returns.
```
gs.run_command('d.vect', map="elev_lidrural_mrpts", where="return=1", col="red", size="2")
gs.run_command('d.vect', map="elev_lidrural_mrpts", where="return=2", col="green", size="3")
gs.run_command('d.vect', map="elev_lidrural_mrpts", where="return=3", col="blue")
gs.run_command('d.vect', map="elev_lidrural_mrpts", where="return=4", col="yellow")
Image(filename="map.png")
```
Can you guess what is in the area that does not have any 1st return points?
```
# end the GRASS session
os.remove(rcfile)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/arjunparmar/VIRTUON/blob/main/Harshit/Fashion_Synthesis_Dataset.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
from google.colab import drive
drive.mount('/content/drive')
!cp /drive/Shareddrives/Virtuon/Machine Learning Project/something.h5 something.h5
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import tqdm
import h5py
import time
import scipy.io
%matplotlib inline
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras.preprocessing.image import load_img,img_to_array
from tensorflow.keras import Sequential, Model
from tensorflow.keras.layers import Conv2D, Flatten, Dense, Input, Conv2DTranspose,MaxPool2D
from tensorflow.keras.layers import concatenate, Activation, MaxPooling2D, Dropout, BatchNormalization
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint, ReduceLROnPlateau
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.losses import binary_crossentropy
from tensorflow.keras.utils import Sequence,to_categorical
from tensorflow.keras.metrics import MeanIoU
import tensorflow.keras.backend as K
# tf.config.list_physical_devices('GPU')
dataset = h5py.File('/content/drive/Shareddrives/Virtuon/Machine Learning Project/G2.h5', 'r')
list(dataset)
dataset['b_']
dataset['ih']
dataset['ih_mean']
```
If U need to explore this dataset more read https://www.christopherlovell.co.uk/blog/2016/04/27/h5py-intro.html
First Modify Images by your own and the go down so that you will be clear of what I have done in dataset
```
test_index = 1000
fig, axes = plt.subplots(1,2, figsize = [10,50])
axes[0].imshow(dataset['b_'][test_index].T.reshape(128,128))
axes[1].imshow(dataset['ih'][test_index].T.reshape(128,128,3))
def preprocess(image, label):
image = tf.transpose(image)
label = tf.transpose(label)
label = tf.reshape(label, [128,128])
image = tf.image.convert_image_dtype(image, dtype = tf.uint8)
label = tf.image.convert_image_dtype(label, dtype = tf.uint8)
label = tf.one_hot(label, depth = 7, dtype = tf.uint8)
# print(label)
return image, label
train = tf.data.Dataset.from_tensor_slices((dataset['ih'][50000:60000], dataset['b_'][50000:60000]))
train = train.map(preprocess, num_parallel_calls = 4)
train = train.shuffle(10000)
train = train.batch(64).repeat()
train = train.prefetch(5)
valid = tf.data.Dataset.from_tensor_slices((dataset['ih'][69500:70000], dataset['b_'][69500:70000]))
valid = valid.map(preprocess, num_parallel_calls = 4)
valid = valid.shuffle(500)
valid = valid.batch(32)
def conv_2d_block(x,n_filters,k_size,batchnorm=False):
''' add two Conv layers with relu activation '''
#first layer
x = Conv2D(filters=n_filters,kernel_size=(k_size,k_size) ,
padding='same', kernel_initializer = 'he_normal')(x)
if batchnorm:
x = BatchNormalization()(x)
x = Activation('relu')(x)
# 2nd layer
x = Conv2D(filters=n_filters,kernel_size=(k_size,k_size) ,
padding='same', kernel_initializer = 'he_normal')(x)
if batchnorm:
x = BatchNormalization()(x)
x = Activation('relu')(x)
x = Conv2D(filters=n_filters,kernel_size=(k_size,k_size) ,
padding='same', kernel_initializer = 'he_normal')(x)
if batchnorm:
x = BatchNormalization()(x)
x = Activation('relu')(x)
return x
def get_u_net(input,n_filters=16,conv_k_size=3,pool_size=2,batchnorm=True,dropout=.2):
c1 = conv_2d_block(input,n_filters * 1 , conv_k_size,batchnorm)
p1 = MaxPool2D(pool_size=(pool_size,pool_size))(c1)
p1 = Dropout(dropout)(p1)
c2 = conv_2d_block(p1,n_filters * 2 , conv_k_size,batchnorm)
p2 = MaxPool2D(pool_size=(pool_size,pool_size))(c2)
p2 = Dropout(dropout)(p2)
c3 = conv_2d_block(p2,n_filters * 4 , conv_k_size,batchnorm)
p3 = MaxPool2D(pool_size=(pool_size,pool_size))(c3)
p3 = Dropout(dropout)(p3)
c4 = conv_2d_block(p3,n_filters * 8 , conv_k_size,batchnorm)
p4 = MaxPool2D(pool_size=(pool_size,pool_size))(c4)
p4 = Dropout(dropout)(p4)
c5 = conv_2d_block(p4,n_filters * 16 , conv_k_size,batchnorm)
#Up sampling
u6 = Conv2DTranspose(filters=n_filters * 8 ,kernel_size=(3,3), strides=(2,2),padding='same')(c5)
u6 = concatenate([u6,c4])
u6 = Dropout(dropout)(u6)
c7 = conv_2d_block(u6,n_filters * 8 , conv_k_size,batchnorm)
u8 = Conv2DTranspose(filters=n_filters * 4 ,kernel_size=(3,3), strides=(2,2),padding='same')(c7)
u8 = concatenate([u8,c3])
u8 = Dropout(dropout)(u8)
c9 = conv_2d_block(u8,n_filters * 4 , conv_k_size,batchnorm)
u10 = Conv2DTranspose(filters=n_filters * 2,kernel_size=(3,3) , strides=(2,2),padding='same')(c9)
u10 = concatenate([u10,c2])
u10 = Dropout(dropout)(u10)
c11 = conv_2d_block(u10,n_filters * 2 , conv_k_size,batchnorm)
u12 = Conv2DTranspose(filters=n_filters * 1 ,kernel_size=(3,3), strides=(2,2),padding='same')(c11)
u12 = concatenate([u12,c1])
u12 = Dropout(dropout)(u12)
c13 = conv_2d_block(u12,n_filters * 1 , conv_k_size,batchnorm)
output = Conv2D(filters=7 , kernel_size=(1,1),activation='softmax')(c13) # output layer
model = Model(inputs=input,outputs=output,name='classifier')
return model
def DiceLoss(targets, inputs):
smooth=1e-6
#flatten label and prediction tensors
inputs = K.flatten(inputs)
targets = K.flatten(targets)
BCE = K.binary_cross_entropy(inputs,targets, reduction='mean')
targets = tf.cast(targets, dtype = tf.float32)
inputs = tf.cast(inputs, dtype = tf.float32)
intersection = K.sum((targets * inputs))
union = K.sum(targets) + K.sum(inputs) - intersection
dice_loss = 1 - ((intersection + smooth) /(union + smooth))
Dice_BCE = dice_loss + BCE
return Dice_BCE
ALPHA = 1
BETA = 500
GAMMA = 1
def FocalTverskyLoss(targets, inputs, alpha=ALPHA, beta=BETA, gamma=GAMMA, smooth=1e-6):
#flatten label and prediction tensors
inputs = K.flatten(inputs)
targets = K.flatten(targets)
inputs = tf.cast(inputs, dtype = tf.float32)
targets = tf.cast(targets, dtype = tf.float32)
#True Positives, False Positives & False Negatives
TP = K.sum((inputs * targets))
FP = K.sum(((1-targets) * inputs))
FN = K.sum((targets * (1-inputs)))
Tversky = (TP + smooth) / (TP + alpha*FP + beta*FN + smooth)
FocalTversky = K.pow((1 - Tversky), gamma)
return FocalTversky
optimizer = Adam()
loss = FocalTverskyLoss
metrics = [MeanIoU(7), 'accuracy']
# Compile our model
# input = Input(shape=(128,128,3),name='img')
# model = get_u_net(input,n_filters=64)
# model.compile(optimizer=optimizer, loss=loss, metrics=metrics)
# model.summary()
callback = [
EarlyStopping(patience=10, verbose=1),
ReduceLROnPlateau(factor=0.1, patience=5, min_lr=1e-5, verbose=1)]
model = tf.keras.models.load_model('something.h5', compile = False)
model.compile(optimizer=optimizer, loss=loss, metrics=metrics)
results = model.fit(x=train,
epochs=25,
verbose=1,
callbacks=callback,
use_multiprocessing=True,
workers=6, steps_per_epoch = 157, validation_data = valid)
model.save('something.h5')
!cp something.h5 drive/MyDrive/Machine\ Learning\ Project/something.h5
test_index = 45000
fig, axes = plt.subplots(1,2, figsize = [10,50])
axes[0].imshow(dataset['b_'][test_index].T.reshape(128,128))
axes[1].imshow(dataset['ih'][test_index].T.reshape(128,128,3))
dataset['ih'][test_index:test_index + 1].shape
predict_ds = tf.data.Dataset.from_tensor_slices(dataset['ih'][test_index:test_index + 1]).map(tf.transpose).batch(1)
predict_ds.element_spec
result = model.predict(predict_ds)
temp = tf.one_hot(dataset['b_'][test_index].T.reshape(128,128), depth = 7, dtype = tf.uint8)
temp_1 = tf.cast(result, dtype = tf.uint8)
index = 3
np.unique(temp[:,:,index] == temp_1[0,:,:,index], return_counts=True)
from sklearn.metrics import confusion_matrix
confusion_matrix(K.flatten(temp[:,:,index]), K.flatten(temp_1[0,:,:,index]))
def one_hot_decoder(image):
temp = np.zeros((128,128))
for i in range(7):
temp += i*image[0,:,:,i]
return temp
plt.imshow(tf.cast(result[0,:,:,0], dtype = tf.uint8))
```
| github_jupyter |
# Deep Learning & Art: Neural Style Transfer
Welcome to the second assignment of this week. In this assignment, you will learn about Neural Style Transfer. This algorithm was created by Gatys et al. (2015) (https://arxiv.org/abs/1508.06576).
**In this assignment, you will:**
- Implement the neural style transfer algorithm
- Generate novel artistic images using your algorithm
Most of the algorithms you've studied optimize a cost function to get a set of parameter values. In Neural Style Transfer, you'll optimize a cost function to get pixel values!
```
import os
import sys
import scipy.io
import scipy.misc
import matplotlib.pyplot as plt
from matplotlib.pyplot import imshow
from PIL import Image
from nst_utils import *
import numpy as np
import tensorflow as tf
%matplotlib inline
```
## 1 - Problem Statement
Neural Style Transfer (NST) is one of the most fun techniques in deep learning. As seen below, it merges two images, namely, a "content" image (C) and a "style" image (S), to create a "generated" image (G). The generated image G combines the "content" of the image C with the "style" of image S.
In this example, you are going to generate an image of the Louvre museum in Paris (content image C), mixed with a painting by Claude Monet, a leader of the impressionist movement (style image S).
<img src="images/louvre_generated.png" style="width:750px;height:200px;">
Let's see how you can do this.
## 2 - Transfer Learning
Neural Style Transfer (NST) uses a previously trained convolutional network, and builds on top of that. The idea of using a network trained on a different task and applying it to a new task is called transfer learning.
Following the original NST paper (https://arxiv.org/abs/1508.06576), we will use the VGG network. Specifically, we'll use VGG-19, a 19-layer version of the VGG network. This model has already been trained on the very large ImageNet database, and thus has learned to recognize a variety of low level features (at the earlier layers) and high level features (at the deeper layers).
Run the following code to load parameters from the VGG model. This may take a few seconds.
```
model = load_vgg_model("pretrained-model/imagenet-vgg-verydeep-19.mat")
print(model)
```
The model is stored in a python dictionary where each variable name is the key and the corresponding value is a tensor containing that variable's value. To run an image through this network, you just have to feed the image to the model. In TensorFlow, you can do so using the [tf.assign](https://www.tensorflow.org/api_docs/python/tf/assign) function. In particular, you will use the assign function like this:
```python
model["input"].assign(image)
```
This assigns the image as an input to the model. After this, if you want to access the activations of a particular layer, say layer `4_2` when the network is run on this image, you would run a TensorFlow session on the correct tensor `conv4_2`, as follows:
```python
sess.run(model["conv4_2"])
```
## 3 - Neural Style Transfer
We will build the NST algorithm in three steps:
- Build the content cost function $J_{content}(C,G)$
- Build the style cost function $J_{style}(S,G)$
- Put it together to get $J(G) = \alpha J_{content}(C,G) + \beta J_{style}(S,G)$.
### 3.1 - Computing the content cost
In our running example, the content image C will be the picture of the Louvre Museum in Paris. Run the code below to see a picture of the Louvre.
```
content_image = scipy.misc.imread("images/louvre.jpg")
imshow(content_image)
```
The content image (C) shows the Louvre museum's pyramid surrounded by old Paris buildings, against a sunny sky with a few clouds.
** 3.1.1 - How do you ensure the generated image G matches the content of the image C?**
As we saw in lecture, the earlier (shallower) layers of a ConvNet tend to detect lower-level features such as edges and simple textures, and the later (deeper) layers tend to detect higher-level features such as more complex textures as well as object classes.
We would like the "generated" image G to have similar content as the input image C. Suppose you have chosen some layer's activations to represent the content of an image. In practice, you'll get the most visually pleasing results if you choose a layer in the middle of the network--neither too shallow nor too deep. (After you have finished this exercise, feel free to come back and experiment with using different layers, to see how the results vary.)
So, suppose you have picked one particular hidden layer to use. Now, set the image C as the input to the pretrained VGG network, and run forward propagation. Let $a^{(C)}$ be the hidden layer activations in the layer you had chosen. (In lecture, we had written this as $a^{[l](C)}$, but here we'll drop the superscript $[l]$ to simplify the notation.) This will be a $n_H \times n_W \times n_C$ tensor. Repeat this process with the image G: Set G as the input, and run forward progation. Let $$a^{(G)}$$ be the corresponding hidden layer activation. We will define as the content cost function as:
$$J_{content}(C,G) = \frac{1}{4 \times n_H \times n_W \times n_C}\sum _{ \text{all entries}} (a^{(C)} - a^{(G)})^2\tag{1} $$
Here, $n_H, n_W$ and $n_C$ are the height, width and number of channels of the hidden layer you have chosen, and appear in a normalization term in the cost. For clarity, note that $a^{(C)}$ and $a^{(G)}$ are the volumes corresponding to a hidden layer's activations. In order to compute the cost $J_{content}(C,G)$, it might also be convenient to unroll these 3D volumes into a 2D matrix, as shown below. (Technically this unrolling step isn't needed to compute $J_{content}$, but it will be good practice for when you do need to carry out a similar operation later for computing the style const $J_{style}$.)
<img src="images/NST_LOSS.png" style="width:800px;height:400px;">
**Exercise:** Compute the "content cost" using TensorFlow.
**Instructions**: The 3 steps to implement this function are:
1. Retrieve dimensions from a_G:
- To retrieve dimensions from a tensor X, use: `X.get_shape().as_list()`
2. Unroll a_C and a_G as explained in the picture above
- If you are stuck, take a look at [Hint1](https://www.tensorflow.org/versions/r1.3/api_docs/python/tf/transpose) and [Hint2](https://www.tensorflow.org/versions/r1.2/api_docs/python/tf/reshape).
3. Compute the content cost:
- If you are stuck, take a look at [Hint3](https://www.tensorflow.org/api_docs/python/tf/reduce_sum), [Hint4](https://www.tensorflow.org/api_docs/python/tf/square) and [Hint5](https://www.tensorflow.org/api_docs/python/tf/subtract).
```
# GRADED FUNCTION: compute_content_cost
def compute_content_cost(a_C, a_G):
"""
Computes the content cost
Arguments:
a_C -- tensor of dimension (1, n_H, n_W, n_C), hidden layer activations representing content of the image C
a_G -- tensor of dimension (1, n_H, n_W, n_C), hidden layer activations representing content of the image G
Returns:
J_content -- scalar that you compute using equation 1 above.
"""
### START CODE HERE ###
# Retrieve dimensions from a_G (≈1 line)
m, n_H, n_W, n_C = None
# Reshape a_C and a_G (≈2 lines)
a_C_unrolled = None
a_G_unrolled = None
# compute the cost with tensorflow (≈1 line)
J_content = None
### END CODE HERE ###
return J_content
tf.reset_default_graph()
with tf.Session() as test:
tf.set_random_seed(1)
a_C = tf.random_normal([1, 4, 4, 3], mean=1, stddev=4)
a_G = tf.random_normal([1, 4, 4, 3], mean=1, stddev=4)
J_content = compute_content_cost(a_C, a_G)
print("J_content = " + str(J_content.eval()))
```
**Expected Output**:
<table>
<tr>
<td>
**J_content**
</td>
<td>
6.76559
</td>
</tr>
</table>
<font color='blue'>
**What you should remember**:
- The content cost takes a hidden layer activation of the neural network, and measures how different $a^{(C)}$ and $a^{(G)}$ are.
- When we minimize the content cost later, this will help make sure $G$ has similar content as $C$.
### 3.2 - Computing the style cost
For our running example, we will use the following style image:
```
style_image = scipy.misc.imread("images/monet_800600.jpg")
imshow(style_image)
```
This painting was painted in the style of *[impressionism](https://en.wikipedia.org/wiki/Impressionism)*.
Lets see how you can now define a "style" const function $J_{style}(S,G)$.
### 3.2.1 - Style matrix
The style matrix is also called a "Gram matrix." In linear algebra, the Gram matrix G of a set of vectors $(v_{1},\dots ,v_{n})$ is the matrix of dot products, whose entries are ${\displaystyle G_{ij} = v_{i}^T v_{j} = np.dot(v_{i}, v_{j}) }$. In other words, $G_{ij}$ compares how similar $v_i$ is to $v_j$: If they are highly similar, you would expect them to have a large dot product, and thus for $G_{ij}$ to be large.
Note that there is an unfortunate collision in the variable names used here. We are following common terminology used in the literature, but $G$ is used to denote the Style matrix (or Gram matrix) as well as to denote the generated image $G$. We will try to make sure which $G$ we are referring to is always clear from the context.
In NST, you can compute the Style matrix by multiplying the "unrolled" filter matrix with their transpose:
<img src="images/NST_GM.png" style="width:900px;height:300px;">
The result is a matrix of dimension $(n_C,n_C)$ where $n_C$ is the number of filters. The value $G_{ij}$ measures how similar the activations of filter $i$ are to the activations of filter $j$.
One important part of the gram matrix is that the diagonal elements such as $G_{ii}$ also measures how active filter $i$ is. For example, suppose filter $i$ is detecting vertical textures in the image. Then $G_{ii}$ measures how common vertical textures are in the image as a whole: If $G_{ii}$ is large, this means that the image has a lot of vertical texture.
By capturing the prevalence of different types of features ($G_{ii}$), as well as how much different features occur together ($G_{ij}$), the Style matrix $G$ measures the style of an image.
**Exercise**:
Using TensorFlow, implement a function that computes the Gram matrix of a matrix A. The formula is: The gram matrix of A is $G_A = AA^T$. If you are stuck, take a look at [Hint 1](https://www.tensorflow.org/api_docs/python/tf/matmul) and [Hint 2](https://www.tensorflow.org/api_docs/python/tf/transpose).
```
# GRADED FUNCTION: gram_matrix
def gram_matrix(A):
"""
Argument:
A -- matrix of shape (n_C, n_H*n_W)
Returns:
GA -- Gram matrix of A, of shape (n_C, n_C)
"""
### START CODE HERE ### (≈1 line)
GA = None
### END CODE HERE ###
return GA
tf.reset_default_graph()
with tf.Session() as test:
tf.set_random_seed(1)
A = tf.random_normal([3, 2*1], mean=1, stddev=4)
GA = gram_matrix(A)
print("GA = " + str(GA.eval()))
```
**Expected Output**:
<table>
<tr>
<td>
**GA**
</td>
<td>
[[ 6.42230511 -4.42912197 -2.09668207] <br>
[ -4.42912197 19.46583748 19.56387138] <br>
[ -2.09668207 19.56387138 20.6864624 ]]
</td>
</tr>
</table>
### 3.2.2 - Style cost
After generating the Style matrix (Gram matrix), your goal will be to minimize the distance between the Gram matrix of the "style" image S and that of the "generated" image G. For now, we are using only a single hidden layer $a^{[l]}$, and the corresponding style cost for this layer is defined as:
$$J_{style}^{[l]}(S,G) = \frac{1}{4 \times {n_C}^2 \times (n_H \times n_W)^2} \sum _{i=1}^{n_C}\sum_{j=1}^{n_C}(G^{(S)}_{ij} - G^{(G)}_{ij})^2\tag{2} $$
where $G^{(S)}$ and $G^{(G)}$ are respectively the Gram matrices of the "style" image and the "generated" image, computed using the hidden layer activations for a particular hidden layer in the network.
**Exercise**: Compute the style cost for a single layer.
**Instructions**: The 3 steps to implement this function are:
1. Retrieve dimensions from the hidden layer activations a_G:
- To retrieve dimensions from a tensor X, use: `X.get_shape().as_list()`
2. Unroll the hidden layer activations a_S and a_G into 2D matrices, as explained in the picture above.
- You may find [Hint1](https://www.tensorflow.org/versions/r1.3/api_docs/python/tf/transpose) and [Hint2](https://www.tensorflow.org/versions/r1.2/api_docs/python/tf/reshape) useful.
3. Compute the Style matrix of the images S and G. (Use the function you had previously written.)
4. Compute the Style cost:
- You may find [Hint3](https://www.tensorflow.org/api_docs/python/tf/reduce_sum), [Hint4](https://www.tensorflow.org/api_docs/python/tf/square) and [Hint5](https://www.tensorflow.org/api_docs/python/tf/subtract) useful.
```
# GRADED FUNCTION: compute_layer_style_cost
def compute_layer_style_cost(a_S, a_G):
"""
Arguments:
a_S -- tensor of dimension (1, n_H, n_W, n_C), hidden layer activations representing style of the image S
a_G -- tensor of dimension (1, n_H, n_W, n_C), hidden layer activations representing style of the image G
Returns:
J_style_layer -- tensor representing a scalar value, style cost defined above by equation (2)
"""
### START CODE HERE ###
# Retrieve dimensions from a_G (≈1 line)
m, n_H, n_W, n_C = None
# Reshape the images to have them of shape (n_C, n_H*n_W) (≈2 lines)
a_S = None
a_G = None
# Computing gram_matrices for both images S and G (≈2 lines)
GS = None
GG = None
# Computing the loss (≈1 line)
J_style_layer = None
### END CODE HERE ###
return J_style_layer
tf.reset_default_graph()
with tf.Session() as test:
tf.set_random_seed(1)
a_S = tf.random_normal([1, 4, 4, 3], mean=1, stddev=4)
a_G = tf.random_normal([1, 4, 4, 3], mean=1, stddev=4)
J_style_layer = compute_layer_style_cost(a_S, a_G)
print("J_style_layer = " + str(J_style_layer.eval()))
```
**Expected Output**:
<table>
<tr>
<td>
**J_style_layer**
</td>
<td>
9.19028
</td>
</tr>
</table>
### 3.2.3 Style Weights
So far you have captured the style from only one layer. We'll get better results if we "merge" style costs from several different layers. After completing this exercise, feel free to come back and experiment with different weights to see how it changes the generated image $G$. But for now, this is a pretty reasonable default:
```
STYLE_LAYERS = [
('conv1_1', 0.2),
('conv2_1', 0.2),
('conv3_1', 0.2),
('conv4_1', 0.2),
('conv5_1', 0.2)]
```
You can combine the style costs for different layers as follows:
$$J_{style}(S,G) = \sum_{l} \lambda^{[l]} J^{[l]}_{style}(S,G)$$
where the values for $\lambda^{[l]}$ are given in `STYLE_LAYERS`.
We've implemented a compute_style_cost(...) function. It simply calls your `compute_layer_style_cost(...)` several times, and weights their results using the values in `STYLE_LAYERS`. Read over it to make sure you understand what it's doing.
<!--
2. Loop over (layer_name, coeff) from STYLE_LAYERS:
a. Select the output tensor of the current layer. As an example, to call the tensor from the "conv1_1" layer you would do: out = model["conv1_1"]
b. Get the style of the style image from the current layer by running the session on the tensor "out"
c. Get a tensor representing the style of the generated image from the current layer. It is just "out".
d. Now that you have both styles. Use the function you've implemented above to compute the style_cost for the current layer
e. Add (style_cost x coeff) of the current layer to overall style cost (J_style)
3. Return J_style, which should now be the sum of the (style_cost x coeff) for each layer.
!-->
```
def compute_style_cost(model, STYLE_LAYERS):
"""
Computes the overall style cost from several chosen layers
Arguments:
model -- our tensorflow model
STYLE_LAYERS -- A python list containing:
- the names of the layers we would like to extract style from
- a coefficient for each of them
Returns:
J_style -- tensor representing a scalar value, style cost defined above by equation (2)
"""
# initialize the overall style cost
J_style = 0
for layer_name, coeff in STYLE_LAYERS:
# Select the output tensor of the currently selected layer
out = model[layer_name]
# Set a_S to be the hidden layer activation from the layer we have selected, by running the session on out
a_S = sess.run(out)
# Set a_G to be the hidden layer activation from same layer. Here, a_G references model[layer_name]
# and isn't evaluated yet. Later in the code, we'll assign the image G as the model input, so that
# when we run the session, this will be the activations drawn from the appropriate layer, with G as input.
a_G = out
# Compute style_cost for the current layer
J_style_layer = compute_layer_style_cost(a_S, a_G)
# Add coeff * J_style_layer of this layer to overall style cost
J_style += coeff * J_style_layer
return J_style
```
**Note**: In the inner-loop of the for-loop above, `a_G` is a tensor and hasn't been evaluated yet. It will be evaluated and updated at each iteration when we run the TensorFlow graph in model_nn() below.
<!--
How do you choose the coefficients for each layer? The deeper layers capture higher-level concepts, and the features in the deeper layers are less localized in the image relative to each other. So if you want the generated image to softly follow the style image, try choosing larger weights for deeper layers and smaller weights for the first layers. In contrast, if you want the generated image to strongly follow the style image, try choosing smaller weights for deeper layers and larger weights for the first layers
!-->
<font color='blue'>
**What you should remember**:
- The style of an image can be represented using the Gram matrix of a hidden layer's activations. However, we get even better results combining this representation from multiple different layers. This is in contrast to the content representation, where usually using just a single hidden layer is sufficient.
- Minimizing the style cost will cause the image $G$ to follow the style of the image $S$.
</font color='blue'>
### 3.3 - Defining the total cost to optimize
Finally, let's create a cost function that minimizes both the style and the content cost. The formula is:
$$J(G) = \alpha J_{content}(C,G) + \beta J_{style}(S,G)$$
**Exercise**: Implement the total cost function which includes both the content cost and the style cost.
```
# GRADED FUNCTION: total_cost
def total_cost(J_content, J_style, alpha = 10, beta = 40):
"""
Computes the total cost function
Arguments:
J_content -- content cost coded above
J_style -- style cost coded above
alpha -- hyperparameter weighting the importance of the content cost
beta -- hyperparameter weighting the importance of the style cost
Returns:
J -- total cost as defined by the formula above.
"""
### START CODE HERE ### (≈1 line)
J = None
### END CODE HERE ###
return J
tf.reset_default_graph()
with tf.Session() as test:
np.random.seed(3)
J_content = np.random.randn()
J_style = np.random.randn()
J = total_cost(J_content, J_style)
print("J = " + str(J))
```
**Expected Output**:
<table>
<tr>
<td>
**J**
</td>
<td>
35.34667875478276
</td>
</tr>
</table>
<font color='blue'>
**What you should remember**:
- The total cost is a linear combination of the content cost $J_{content}(C,G)$ and the style cost $J_{style}(S,G)$
- $\alpha$ and $\beta$ are hyperparameters that control the relative weighting between content and style
## 4 - Solving the optimization problem
Finally, let's put everything together to implement Neural Style Transfer!
Here's what the program will have to do:
<font color='purple'>
1. Create an Interactive Session
2. Load the content image
3. Load the style image
4. Randomly initialize the image to be generated
5. Load the VGG16 model
7. Build the TensorFlow graph:
- Run the content image through the VGG16 model and compute the content cost
- Run the style image through the VGG16 model and compute the style cost
- Compute the total cost
- Define the optimizer and the learning rate
8. Initialize the TensorFlow graph and run it for a large number of iterations, updating the generated image at every step.
</font>
Lets go through the individual steps in detail.
You've previously implemented the overall cost $J(G)$. We'll now set up TensorFlow to optimize this with respect to $G$. To do so, your program has to reset the graph and use an "[Interactive Session](https://www.tensorflow.org/api_docs/python/tf/InteractiveSession)". Unlike a regular session, the "Interactive Session" installs itself as the default session to build a graph. This allows you to run variables without constantly needing to refer to the session object, which simplifies the code.
Lets start the interactive session.
```
# Reset the graph
tf.reset_default_graph()
# Start interactive session
sess = tf.InteractiveSession()
```
Let's load, reshape, and normalize our "content" image (the Louvre museum picture):
```
content_image = scipy.misc.imread("images/louvre_small.jpg")
content_image = reshape_and_normalize_image(content_image)
```
Let's load, reshape and normalize our "style" image (Claude Monet's painting):
```
style_image = scipy.misc.imread("images/monet.jpg")
style_image = reshape_and_normalize_image(style_image)
```
Now, we initialize the "generated" image as a noisy image created from the content_image. By initializing the pixels of the generated image to be mostly noise but still slightly correlated with the content image, this will help the content of the "generated" image more rapidly match the content of the "content" image. (Feel free to look in `nst_utils.py` to see the details of `generate_noise_image(...)`; to do so, click "File-->Open..." at the upper-left corner of this Jupyter notebook.)
```
generated_image = generate_noise_image(content_image)
imshow(generated_image[0])
```
Next, as explained in part (2), let's load the VGG16 model.
```
model = load_vgg_model("pretrained-model/imagenet-vgg-verydeep-19.mat")
```
To get the program to compute the content cost, we will now assign `a_C` and `a_G` to be the appropriate hidden layer activations. We will use layer `conv4_2` to compute the content cost. The code below does the following:
1. Assign the content image to be the input to the VGG model.
2. Set a_C to be the tensor giving the hidden layer activation for layer "conv4_2".
3. Set a_G to be the tensor giving the hidden layer activation for the same layer.
4. Compute the content cost using a_C and a_G.
```
# Assign the content image to be the input of the VGG model.
sess.run(model['input'].assign(content_image))
# Select the output tensor of layer conv4_2
out = model['conv4_2']
# Set a_C to be the hidden layer activation from the layer we have selected
a_C = sess.run(out)
# Set a_G to be the hidden layer activation from same layer. Here, a_G references model['conv4_2']
# and isn't evaluated yet. Later in the code, we'll assign the image G as the model input, so that
# when we run the session, this will be the activations drawn from the appropriate layer, with G as input.
a_G = out
# Compute the content cost
J_content = compute_content_cost(a_C, a_G)
```
**Note**: At this point, a_G is a tensor and hasn't been evaluated. It will be evaluated and updated at each iteration when we run the Tensorflow graph in model_nn() below.
```
# Assign the input of the model to be the "style" image
sess.run(model['input'].assign(style_image))
# Compute the style cost
J_style = compute_style_cost(model, STYLE_LAYERS)
```
**Exercise**: Now that you have J_content and J_style, compute the total cost J by calling `total_cost()`. Use `alpha = 10` and `beta = 40`.
```
### START CODE HERE ### (1 line)
J = None
### END CODE HERE ###
```
You'd previously learned how to set up the Adam optimizer in TensorFlow. Lets do that here, using a learning rate of 2.0. [See reference](https://www.tensorflow.org/api_docs/python/tf/train/AdamOptimizer)
```
# define optimizer (1 line)
optimizer = tf.train.AdamOptimizer(2.0)
# define train_step (1 line)
train_step = optimizer.minimize(J)
```
**Exercise**: Implement the model_nn() function which initializes the variables of the tensorflow graph, assigns the input image (initial generated image) as the input of the VGG16 model and runs the train_step for a large number of steps.
```
def model_nn(sess, input_image, num_iterations = 200):
# Initialize global variables (you need to run the session on the initializer)
### START CODE HERE ### (1 line)
None
### END CODE HERE ###
# Run the noisy input image (initial generated image) through the model. Use assign().
### START CODE HERE ### (1 line)
None
### END CODE HERE ###
for i in range(num_iterations):
# Run the session on the train_step to minimize the total cost
### START CODE HERE ### (1 line)
None
### END CODE HERE ###
# Compute the generated image by running the session on the current model['input']
### START CODE HERE ### (1 line)
generated_image = None
### END CODE HERE ###
# Print every 20 iteration.
if i%20 == 0:
Jt, Jc, Js = sess.run([J, J_content, J_style])
print("Iteration " + str(i) + " :")
print("total cost = " + str(Jt))
print("content cost = " + str(Jc))
print("style cost = " + str(Js))
# save current generated image in the "/output" directory
save_image("output/" + str(i) + ".png", generated_image)
# save last generated image
save_image('output/generated_image.jpg', generated_image)
return generated_image
```
Run the following cell to generate an artistic image. It should take about 3min on CPU for every 20 iterations but you start observing attractive results after ≈140 iterations. Neural Style Transfer is generally trained using GPUs.
```
model_nn(sess, generated_image)
```
**Expected Output**:
<table>
<tr>
<td>
**Iteration 0 : **
</td>
<td>
total cost = 5.05035e+09 <br>
content cost = 7877.67 <br>
style cost = 1.26257e+08
</td>
</tr>
</table>
You're done! After running this, in the upper bar of the notebook click on "File" and then "Open". Go to the "/output" directory to see all the saved images. Open "generated_image" to see the generated image! :)
You should see something the image presented below on the right:
<img src="images/louvre_generated.png" style="width:800px;height:300px;">
We didn't want you to wait too long to see an initial result, and so had set the hyperparameters accordingly. To get the best looking results, running the optimization algorithm longer (and perhaps with a smaller learning rate) might work better. After completing and submitting this assignment, we encourage you to come back and play more with this notebook, and see if you can generate even better looking images.
Here are few other examples:
- The beautiful ruins of the ancient city of Persepolis (Iran) with the style of Van Gogh (The Starry Night)
<img src="images/perspolis_vangogh.png" style="width:750px;height:300px;">
- The tomb of Cyrus the great in Pasargadae with the style of a Ceramic Kashi from Ispahan.
<img src="images/pasargad_kashi.png" style="width:750px;height:300px;">
- A scientific study of a turbulent fluid with the style of a abstract blue fluid painting.
<img src="images/circle_abstract.png" style="width:750px;height:300px;">
## 5 - Test with your own image (Optional/Ungraded)
Finally, you can also rerun the algorithm on your own images!
To do so, go back to part 4 and change the content image and style image with your own pictures. In detail, here's what you should do:
1. Click on "File -> Open" in the upper tab of the notebook
2. Go to "/images" and upload your images (requirement: (WIDTH = 300, HEIGHT = 225)), rename them "my_content.png" and "my_style.png" for example.
3. Change the code in part (3.4) from :
```python
content_image = scipy.misc.imread("images/louvre.jpg")
style_image = scipy.misc.imread("images/claude-monet.jpg")
```
to:
```python
content_image = scipy.misc.imread("images/my_content.jpg")
style_image = scipy.misc.imread("images/my_style.jpg")
```
4. Rerun the cells (you may need to restart the Kernel in the upper tab of the notebook).
You can share your generated images with us on social media with the hashtag #deeplearniNgAI or by direct tagging!
You can also tune your hyperparameters:
- Which layers are responsible for representing the style? STYLE_LAYERS
- How many iterations do you want to run the algorithm? num_iterations
- What is the relative weighting between content and style? alpha/beta
## 6 - Conclusion
Great job on completing this assignment! You are now able to use Neural Style Transfer to generate artistic images. This is also your first time building a model in which the optimization algorithm updates the pixel values rather than the neural network's parameters. Deep learning has many different types of models and this is only one of them!
<font color='blue'>
What you should remember:
- Neural Style Transfer is an algorithm that given a content image C and a style image S can generate an artistic image
- It uses representations (hidden layer activations) based on a pretrained ConvNet.
- The content cost function is computed using one hidden layer's activations.
- The style cost function for one layer is computed using the Gram matrix of that layer's activations. The overall style cost function is obtained using several hidden layers.
- Optimizing the total cost function results in synthesizing new images.
This was the final programming exercise of this course. Congratulations--you've finished all the programming exercises of this course on Convolutional Networks! We hope to also see you in Course 5, on Sequence models!
### References:
The Neural Style Transfer algorithm was due to Gatys et al. (2015). Harish Narayanan and Github user "log0" also have highly readable write-ups from which we drew inspiration. The pre-trained network used in this implementation is a VGG network, which is due to Simonyan and Zisserman (2015). Pre-trained weights were from the work of the MathConvNet team.
- Leon A. Gatys, Alexander S. Ecker, Matthias Bethge, (2015). A Neural Algorithm of Artistic Style (https://arxiv.org/abs/1508.06576)
- Harish Narayanan, Convolutional neural networks for artistic style transfer. https://harishnarayanan.org/writing/artistic-style-transfer/
- Log0, TensorFlow Implementation of "A Neural Algorithm of Artistic Style". http://www.chioka.in/tensorflow-implementation-neural-algorithm-of-artistic-style
- Karen Simonyan and Andrew Zisserman (2015). Very deep convolutional networks for large-scale image recognition (https://arxiv.org/pdf/1409.1556.pdf)
- MatConvNet. http://www.vlfeat.org/matconvnet/pretrained/
| github_jupyter |
## The `mne` package
It is possible to build chord diagrams from a connectivity matrix thanks to the neuroscience library [MNE](https://mne.tools/stable/index.html). It comes with a visual function called [plot_connectivity_circle()](https://mne.tools/stable/generated/mne.viz.plot_connectivity_circle.html#mne.viz.plot_connectivity_circle) that is pretty handy to get good-looking chord diagrams in minutes!
Let's load the library and see what it can make!
```
from mne.viz import plot_connectivity_circle
# only for the exemple
import numpy as np
```
## Most basic chord diagram with `mne`
Let's start with a basic examples. 20 nodes that are randomly connected. Two objects are created:
- `node_names` that is a list of 20 node names
- `con` that is an object containing some random links between nodes.
Both object are passed to the `plot_connectivity_circle()` function that automatically builds the chord diagram.
```
N = 20 # Number of nodes
node_names = [f"N{i}" for i in range(N)] # List of labels [N]
# Random connectivity
ran = np.random.rand(N,N)
con = np.where(ran > 0.9, ran, np.nan) # NaN so it doesn't display the weak links
fig, axes = plot_connectivity_circle(con, node_names)
```
## Split the chord
It is possible to split the chord diagram in several parts. It can be handy to build chord diagrams where nodes are split in 2 groups, like origin and destination for instance.
```
start, end = 45, 135
first_half = (np.linspace(start, end, len(node_names)//2) +90).astype(int)[::+1] %360
second_half = (np.linspace(start, end, len(node_names)//2) -90).astype(int)[::-1] %360
node_angles = np.array(list(first_half) + list(second_half))
fig, axes = plot_connectivity_circle(con, node_names,
node_angles=node_angles)
```
## Style: node customization
Pretty much all parts of the chord diagram can be customized. Let's start by changing the node width (with `node_width`) and filtering the links that are shown (with `vmin` and `vmax`)
```
fig, axes = plot_connectivity_circle(con, node_names,
node_width=2, vmin=0.97, vmax=0.98)
```
Now let's customize the nodes a bit more:
- `node_colors` for the fill color
- `node_edgecolor` for the edges
- `node_linewidth` for the width
```
node_edgecolor = N//2 * [(0,0,0,0.)] + N//2 * ['green']
node_colors = N//2 * ['crimson'] + N//2 * [(0,0,0,0.)]
fig, axes = plot_connectivity_circle(con, node_names,
node_colors=node_colors, node_edgecolor=node_edgecolor, node_linewidth=2)
```
## Style: labels and links
Now some customization for labels, links and background:
- `colormap`
- `facecolor`
- `textcolor`
- `colorbar`
- `linewidth`
```
fig, axes = plot_connectivity_circle(con, node_names,
colormap='Blues', facecolor='white', textcolor='black', colorbar=False,
linewidth=10)
```
## Brocoli
Let's get some fun and build a data art brocoli like chord diagram 😊 !
```
N = 200
node_names = N * ['']
ran = np.random.rand(N,N)
con = np.where(ran > 0.95, ran, np.nan)
first_half = (np.linspace(0, 180, len(node_names)//2)).astype(int)[::+1] %360
second_half = (np.linspace(70, 110, len(node_names)//2)-180).astype(int)[::-1] %360
node_angles = np.array(list(first_half) + list(second_half))
node_colors = node_edgecolor = N * ['green']
fig, axes = plot_connectivity_circle(con, node_names,
node_angles=node_angles,
colormap='Greens', facecolor='w', textcolor='k', colorbar=False,
node_colors=node_colors, node_edgecolor=node_edgecolor,
node_width=0.1, node_linewidth=1, linewidth=1)
```
| github_jupyter |
```
from statsmodels.stats.outliers_influence import variance_inflation_factor
from sklearn.model_selection import KFold
from sklearn.datasets import make_regression
from sklearn.linear_model import LinearRegression
from sklearn.metrics import r2_score
from sklearn.model_selection import cross_val_score
%matplotlib inline
%config InlineBackend.figure_formats = {'png', 'retina'}
pd.options.mode.chained_assignment = None # default='warn'?
# data_key DataFrame
data_key = pd.read_csv('key.csv')
# data_train DataFrame
data_train = pd.read_csv('train.csv')
# data_weather DataFrame
data_weather = pd.read_csv('weather.csv')
rain_text = ['FC', 'TS', 'GR', 'RA', 'DZ', 'SN', 'SG', 'GS', 'PL', 'IC', 'FG', 'BR', 'UP', 'FG+']
other_text = ['HZ', 'FU', 'VA', 'DU', 'DS', 'PO', 'SA', 'SS', 'PY', 'SQ', 'DR', 'SH', 'FZ', 'MI', 'PR', 'BC', 'BL', 'VC' ]
data_weather['codesum'].replace("+", "")
a = []
for i in range(len(data_weather['codesum'])):
a.append(data_weather['codesum'].values[i].split(" "))
for i_text in a[i]:
if len(i_text) == 4:
a[i].append(i_text[:2])
a[i].append(i_text[2:])
data_weather["nothing"] = 1
data_weather["rain"] = 0
data_weather["other"] = 0
b = -1
for ls in a:
b += 1
for text in ls:
if text in rain_text:
data_weather.loc[b, 'rain'] = 1
data_weather.loc[b, 'nothing'] = 0
elif text in other_text:
data_weather.loc[b,'other'] = 1
data_weather.loc[b, 'nothing'] = 0
# 모든 데이터 Merge
df = pd.merge(data_weather, data_key)
station_nbr = df['station_nbr']
df.drop('station_nbr', axis=1, inplace=True)
df['station_nbr'] = station_nbr
df = pd.merge(df, data_train)
# T 값 처리 하기. Remained Subject = > 'M' and '-'
df['snowfall'][df['snowfall'] == ' T'] = 0.05
df['preciptotal'][df['preciptotal'] == ' T'] = 0.005
# 주말과 주중 구분 작업 하기
df['date'] = pd.to_datetime(df['date'])
df['week7'] = df['date'].dt.dayofweek
df['weekend'] = 0
df.loc[df['week7'] == 5, 'weekend'] = 1
df.loc[df['week7'] == 6, 'weekend'] = 1
df1 = df[df['station_nbr'] == 1]; df11 = df[df['station_nbr'] == 11]
df2 = df[df['station_nbr'] == 2]; df12 = df[df['station_nbr'] == 12]
df3 = df[df['station_nbr'] == 3]; df13 = df[df['station_nbr'] == 13]
df4 = df[df['station_nbr'] == 4]; df14 = df[df['station_nbr'] == 14]
df5 = df[df['station_nbr'] == 5]; df15 = df[df['station_nbr'] == 15]
df6 = df[df['station_nbr'] == 6]; df16 = df[df['station_nbr'] == 16]
df7 = df[df['station_nbr'] == 7]; df17 = df[df['station_nbr'] == 17]
df8 = df[df['station_nbr'] == 8]; df18 = df[df['station_nbr'] == 18]
df9 = df[df['station_nbr'] == 9]; df19 = df[df['station_nbr'] == 19]
df10 = df[df['station_nbr'] == 10]; df20 = df[df['station_nbr'] == 20]
df17 = df17.apply(pd.to_numeric, errors = 'coerce')
df17.describe().iloc[:, :15]
# 없는 Column = date, codesum, depart, sunrise, sunset, station_nbr
df17['store_nbr'].unique()
df17_drop_columns = ['date', 'codesum', 'depart', 'sunrise', 'sunset', 'station_nbr']
df17 = df17.drop(columns = df17_drop_columns)
# np.nan를 포함하고 있는 변수(column)를 찾아서, 그 변수에 mean 값 대입해서 Frame의 모든 Value가 fill 되게 하기.
df17_columns = df17.columns
# Cateogry 값을 포함하는 변수는 np.nan에 mode값으로 대체하고, 나머지 실수 값을 포함한 변수는 np.nan에 mean값으로 대체
for i in df17_columns:
if (i == 'resultdir'):
df17[i].fillna(df17[i].mode()[0], inplace=True)
print(df17[i].mode()[0])
else:
df17[i].fillna(df17[i].mean(), inplace=True)
# 이제 모든 변수가 숫자로 표기 되었기 때문에, 가능 함.
# 상대 습도 추가 #
df17['relative_humility'] = 100*(np.exp((17.625*((df17['dewpoint']-32)/1.8))/(243.04+((df17['dewpoint']-32)/1.8)))/np.exp((17.625*((df17['tavg']-32)/1.8))/(243.04+((df17['tavg']-32)/1.8))))
# 체감온도 계산
df17["windchill"] = 35.74 + 0.6217*df17["tavg"] - 35.75*(df17["avgspeed"]**0.17) + 0.4275*df17["tavg"]*(df17["avgspeed"]**0.17)
df17 = df17[df17['units'] != 0]
# 'np.log1p(units) ~ tmax + tmin + tavg + depart + dewpoint + wetbulb + heat + cool + sunrise + sunset + codesume + \
# snowfall + preciptotal + stnpressure + sealevel + resultspeed + resultdir + avgspeed + nothing + rain + other + \
# store_nbr + station_nbr + item_nbr'
model_df17 = sm.OLS.from_formula('np.log1p(units) ~ tmax + tmin + tavg + dewpoint + heat + cool + preciptotal + \
resultspeed + sealevel + snowfall + resultdir + avgspeed + C(nothing) + C(rain) + C(other) + C(item_nbr) + C(week7) + \
C(weekend) + relative_humility + windchill + 0', data = df17)
result_df17 = model_df17.fit()
print(result_df17.summary())
model_df17 = sm.OLS.from_formula('np.log1p(units) ~ scale(tmax) + scale(tmin) + scale(tavg) + scale(dewpoint) + scale(heat) + scale(cool) + \
scale(preciptotal) + scale(resultspeed) + scale(sealevel) + scale(snowfall)+ scale(resultdir) + scale(avgspeed) + C(nothing) + C(rain) + \
C(other) + C(item_nbr) + C(week7) + C(weekend) + scale(relative_humility) + scale(windchill) + 0', data = df17)
result_df17 = model_df17.fit()
print(result_df17.summary())
anova_result_df17 = sm.stats.anova_lm(result_df17).sort_values(by=['PR(>F)'], ascending = False)
anova_result_df17[anova_result_df17['PR(>F)'] <= 0.05]
vif = pd.DataFrame()
vif["VIF Factor"] = [variance_inflation_factor(df17.values, i) for i in range(df17.shape[1])]
vif["features"] = df17.columns
vif = vif.sort_values("VIF Factor").reset_index(drop=True)
vif
# 10순위까지 겹치는것만 쓴다
# C(week7) + C(item_nbr)
# resultspeed, avgspeed, C(weekend), preciptotal, C(other), C(week7), C(item_nbr),
model_df17 = sm.OLS.from_formula('np.log1p(units) ~ C(week7) + C(item_nbr) + 0', data = df17)
result_df17 = model_df17.fit()
print(result_df17.summary())
# resultspeed, avgspeed, C(weekend), preciptotal, C(other), C(week7), C(item_nbr),
X17 = df17[['week7', 'item_nbr']]
y17 = df17['units']
model17 = LinearRegression()
cv17 = KFold(n_splits=17, shuffle=True, random_state=0)
cross_val_score(model17, X17, y17, scoring="r2", cv=cv17)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/ibrahimsesay/griddb/blob/master/Week_3_Assignment_3_Credit_Housing_Information_(1).ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# [Problem 1] Understanding the content of the competition
#### The competition's overview page and report what the "Home Credit Default Risk" competition looks like from the following perspectives
- <b>What kind of company is Home Credit?</b>
Home Credit is a non-banking financial institution, founded in 1997 in the Czech Republic.
The company operates in 14 countries (including United States, Russia, Kazahstan, Belarus, China, India) and focuses on lending primarily to people with little or no credit history which will either not obtain loans or became victims of untrustworthly lenders.
Home Credit group has over 29 million customers, total assests of 21 billions Euro, over 160 millions loans, with the majority in Asia and and almost half of them in China (as of 19-05-2018)
Home Credit is like a Micro-Finance company(<b><u>financial institution</u></b>) the gives or grant loan which </br>
strives to broaden financial inclusion for the unbanked population by providing a positive and safe borrowing experience
```
```
- <b>What is expected in this competition?</b>
Proper preditions are to be made with the uses of a variety of alternative data - including telco and transactional information in other to know how capable each applicant is of repaying a loan. </br>
- <b>What are the benefits companies can gain by predicting this</b>
This will help them unlock the full potential of their data. And also will ensure that clients capable of repayment are not rejected and that loans are given with a principal, maturity, and repayment calendar that will empower their clients to be successful.
# [Problem 2] Understanding the overview of data
#### Load the data
```
import pandas as pd
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
application = pd.read_csv("application_train.csv")
```
- <b>An Overview of the Data</b>
```
application.head()
```
##### The head function returns the object with the desired number of rows.
```
application.info()
```
##### The .info() method allows us to learn the shape of object types of our data
```
application.describe()
```
###### The describe() function is used for generating descriptive statistics of a dataset.
- This mean, it gives us summary statistics for numerical columns in our DataFrame
```
application.count()
```
##### The pandas count() function helps in counting non-NA cells of each column or row.
```
```
### Check missing data or values
```
def missing_data(data):
total = data.isnull().sum().sort_values(ascending = False)
percent = (data.isnull().sum()/data.isnull().count()*100).sort_values(ascending = False)
return pd.concat([total, percent], axis=1, keys=['Total', 'Percent'])
missing_data(application).head(10)
```
- <b>Number of columns with missing values or data</b>
```
application.isnull().any(axis=0).sum()
```
- <b>Number of rows with missing values or data</b>
```
application.isnull().any(axis=1).sum()
```
- <b>Draw a graph showing the percentage of classes</b>
```
X = application.drop(['TARGET'], axis=1)
y = application.loc[:, 'TARGET']
y.value_counts()
label_counts = y.value_counts()
labels = 'loan repaid or time', 'loan upaid'
plt.title("Loan repaid or not")
plt.pie(label_counts, labels=labels, autopct='%.1f%%', shadow=True, startangle=90)
plt.show()
```
##### TARGET value 0 means loan is repayed, value 1 means loan is not repayed
```
temp = application["TARGET"].value_counts()
df = pd.DataFrame({'labels': temp.index,
'values': temp.values
})
plt.figure(figsize = (6,6))
plt.title('Application loans repayed - train dataset')
sns.set_color_codes("pastel")
sns.barplot(x = 'labels', y="values", data=df)
locs, labels = plt.xticks()
plt.show()
```
# [Problem 3] Defining issues
- Please set multiple issues / questions of your own based on the overview of the data.
<ol>
<li>
Does the clients Single/not married and civil marriage?</li>
<li>Does the clients have children?
</li>
<li>What is the clients Education Level?</li>
<li>Does applicants for credits registered their housing as House/apartmen?</li>
<li>What is the type of income of the clients?</li>
<li>On which day of the week did the client apply for the loan?</li>
<li>Does the client owns a car or real estate?</li>
<li>What is the clients gender?</li>
<li>Which type of loan did the clients took?</li>
<li>How much loan did the clients take?</li>
<li>What is the Normalized population of region where client lives ?</li>
<li>How many family members does the clients have?</li>
<li>Who was accompanying client when he was applying for the loan?</li>
<li>How many rooms are in the house?</li>
<li>On which day of the week did the client apply for the loan?</li>
<li>Does the clients have a job?</li>
<li>What is the job/organization type of the clients?</li>
<li>What is the Days from birth distribution of the clients?</li>
<li>What is the distribution of total income for the clients?</li>
<li>What is the Clients income type (businessman, working, maternity leave,�)?</li>
<li>Approximately at what hour did the client apply for the loan?</li>
<li>In what year the house was built?</li>
<li>How many observation of client's social surroundings with observable 30 DPD ?</li>
<li>How many days before application did client change phone?</li>
<li>Did client provide documents ?</li>
</ol</ol>
# [Problem 4] Data exploration
- Which type of loan did the clients took?
Let's see the type of the loans taken and also, on a separate plot, the percent of the loans (by type of the loan) with TARGET value 1 (not returned loan)
```
def plot_stats(feature,label_rotation=False,horizontal_layout=True):
temp = application[feature].value_counts()
df1 = pd.DataFrame({feature: temp.index,'Number of contracts': temp.values})
# Calculate the percentage of target=1 per category value
cat_perc = application[[feature, 'TARGET']].groupby([feature],as_index=False).mean()
cat_perc.sort_values(by='TARGET', ascending=False, inplace=True)
if(horizontal_layout):
fig, (ax1, ax2) = plt.subplots(ncols=2, figsize=(12,6))
else:
fig, (ax1, ax2) = plt.subplots(nrows=2, figsize=(12,14))
sns.set_color_codes("pastel")
s = sns.barplot(ax=ax1, x = feature, y="Number of contracts",data=df1)
if(label_rotation):
s.set_xticklabels(s.get_xticklabels(),rotation=90)
s = sns.barplot(ax=ax2, x = feature, y='TARGET', order=cat_perc[feature], data=cat_perc)
if(label_rotation):
s.set_xticklabels(s.get_xticklabels(),rotation=90)
plt.ylabel('Percent of target with value 1 [%]', fontsize=10)
plt.tick_params(axis='both', which='major', labelsize=10)
plt.show();
def plot_distribution(var):
i = 0
t1 = application.loc[application['TARGET'] != 0]
t0 = application.loc[application['TARGET'] == 0]
sns.set_style('whitegrid')
plt.figure()
fig, ax = plt.subplots(2,2,figsize=(12,12))
for feature in var:
i += 1
plt.subplot(2,2,i)
sns.kdeplot(t1[feature], bw=0.5,label="TARGET = 1")
sns.kdeplot(t0[feature], bw=0.5,label="TARGET = 0")
plt.ylabel('Density plot', fontsize=12)
plt.xlabel(feature, fontsize=12)
locs, labels = plt.xticks()
plt.tick_params(axis='both', which='major', labelsize=12)
plt.show();
plot_stats('NAME_CONTRACT_TYPE')
```
- Does the clients have children?
```
plot_stats('CNT_CHILDREN')
```
- Does the clients Single/not married and civil marriage?
```
plot_stats('NAME_FAMILY_STATUS',True, True)
```
- What is the clients Education Level?
```
plot_stats('NAME_EDUCATION_TYPE',True)
```
- Does applicants for credits registered their housing as House/apartmen?
```
plot_stats('NAME_HOUSING_TYPE',True)
```
- Does the client owns a car or real estate?
```
plot_stats('FLAG_OWN_CAR')
plot_stats('FLAG_OWN_REALTY')
```
- What is the clients gender?
```
plot_stats('CODE_GENDER')
```
- What is the job type of the clients?
```
plot_stats('OCCUPATION_TYPE',True, False)
```
- How many family members does the clients have?
```
plot_stats('CNT_FAM_MEMBERS',True)
```
- What is the job/organization type of the clients?
```
plot_stats('ORGANIZATION_TYPE',True, False)
```
- Who was accompanying client when he was applying for the loan?
```
plot_stats('NAME_TYPE_SUITE',True)
```
- What is the Clients income type (businessman, working, maternity leave,�)?
```
plot_stats('NAME_INCOME_TYPE',False,False)
```
#### We will also consider the distribution of some features
```
# Plot distribution of one feature
def plot_distribution(feature,color):
plt.figure(figsize=(10,6))
plt.title("Distribution of %s" % feature)
sns.displot(application[feature].dropna(),color=color, kde=True,bins=100)
plt.show()
```
- <b>Note sns.distplot does not work well in my environment. Therefore sns.displot is used</b>
```
# Plot distribution of multiple features, with TARGET = 1/0 on the same graph
def plot_distribution_comp(var,nrow=2):
i = 0
t1 = application.loc[application['TARGET'] != 0]
t0 = application.loc[application['TARGET'] == 0]
sns.set_style('whitegrid')
plt.figure()
fig, ax = plt.subplots(nrow,2,figsize=(12,6*nrow))
for feature in var:
i += 1
plt.subplot(nrow,2,i)
sns.kdeplot(t1[feature], bw=0.5,label="TARGET = 1")
sns.kdeplot(t0[feature], bw=0.5,label="TARGET = 0")
plt.ylabel('Density plot', fontsize=12)
plt.xlabel(feature, fontsize=12)
locs, labels = plt.xticks()
plt.tick_params(axis='both', which='major', labelsize=12)
plt.show();
```
- What is the distribution of total income for the clients?
```
plot_distribution('AMT_INCOME_TOTAL','green')
```
- What is the Days from birth distribution of the clients?
```
plot_distribution('DAYS_BIRTH','red')
```
- On which day of the week did the client apply for the loan?
```
plot_stats('WEEKDAY_APPR_PROCESS_START','brown')
```
- Approximately at what hour did the client apply for the loan
```
plot_distribution('HOUR_APPR_PROCESS_START','green')
```
# [Problem 5] (Advanced task) Posting to Notebooks
```
```
| github_jupyter |
### 内容简介
特征选择的常用方法,包括Filter、Wrapper和Embedded方法。
Filter方法包括方差分析、相关系数法、卡方检验、F检验和互信息法。
Wrapper主要是递归特征消除法。
Embedded方法主要包括基于树模型的特征选择法和基于正则化的特征选择法。
```
import numpy as np
import pandas as pd
import pymysql
from sklearn.feature_selection import VarianceThreshold
from sklearn.preprocessing import StandardScaler, OneHotEncoder
from scipy.stats import pearsonr
from collections import Counter
from sklearn.feature_selection import chi2 # 卡方检验
from sklearn.feature_selection import SelectKBest # 根据 k个最高分选择功能。
from sklearn.feature_selection import mutual_info_classif
from sklearn.feature_selection import f_classif # F检验
from sklearn.feature_selection import RFE, RFECV
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import StratifiedKFold
from sklearn.feature_selection import SelectFromModel
from xgboost import XGBClassifier
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.svm import LinearSVC
import warnings
warnings.filterwarnings("ignore")
```
### 1、获取数据
```
connection = pymysql.Connect(
host="localhost",
port=3306,
user="root",
passwd="root",
charset="utf8",
db="project_researchers"
)
def get_data(connection):
"""
查询数据,包括特征和标签
:param connection:
:return:
"""
sql_select = """
SELECT bys_cn, hindex_cn,a_conf+a_journal as a_paper, b_conf + b_journal as b_paper,c_conf + c_journal as c_paper,papernum2017, papernum2016, papernum2015, papernum2014, papernum2013,num_journal,num_conference, project_num, degree, pagerank,degree_centrality,last_year - first_year as diff_year , coauthors_top10000, coauthors_top20000, coauthors_top30000, category, label
FROM classifier_isTeacher_label WHERE (label =1 or label = 0) and category is not null
"""
df = pd.read_sql_query(sql_select, connection)
all_features = ['bys_cn', 'hindex_cn', 'a_paper', 'b_paper', 'c_paper', 'papernum2017', 'papernum2016', 'papernum2015', 'papernum2014', 'papernum2013', 'num_journal', 'num_conference', 'degree', 'pagerank', 'degree_centrality', 'diff_year', 'coauthors_top10000', 'coauthors_top20000', 'coauthors_top30000', 'category', 'label']
data = df[all_features]
return data
data = get_data(connection)
print("shape of data:", data.shape)
print("data.info():", data.info())
```
### 2、数据处理
```
# 对缺失值进行处理
# Method1:直接将含有缺失字段的值去掉
data = data.dropna()
print("shape of data::", data.shape)
print("data.info()::", data.info())
# 将连续值和离散值以及y分开
continuous_features = ['bys_cn', 'hindex_cn', 'a_paper', 'b_paper', 'c_paper', 'papernum2017', 'papernum2016', 'papernum2015', 'papernum2014', 'papernum2013', 'num_journal', 'num_conference', 'degree', 'pagerank', 'degree_centrality', 'diff_year', 'coauthors_top10000', 'coauthors_top20000', 'coauthors_top30000']
discrete_features = ['category']
X_continous = data[continuous_features]
X_discrete = data[discrete_features]
y = data['label']
print("info of X_continous::", X_continous.info())
print("info of X_discrete::", X_discrete.info())
print("y::", Counter(y))
# 对连续值进行归一化处理,对离散值进行one-hot编码
# 暂时先不进行归一化处理,因为后面要寻找大方差的特征等
# ss = StandardScaler()
# X_continous = ss.fit_transform(X_continous)
# print("type of X_continous::", type(X_continous))
X_discrete_oneHot = OneHotEncoder(sparse=False).fit_transform(X_discrete)
print(X_discrete_oneHot)
X_all = np.hstack((X_continous, X_discrete_oneHot))
print("shape of X_all::", X_all.shape)
```
### 3、Filter方法
过滤法,按照发散性或者相关性对各个特征进行评分,设定阈值或者待选择阈值的个数,选择特征。
包括方差分析、相关系数法、卡方检验、F检验和互信息法。
#### (1)方差分析
方差较大的特征说明其取值发散,使用方差法,要先计算各个特征的方差,然后根据阈值,选择方差大于阈值的特征。
#### (2)相关系数法
皮尔逊系数只能衡量线性相关性,先要计算各个特征对目标值的相关系数以及相关系数的P值
#### (3)卡方检验
卡方检验只能用用于二分类。
#### (4)F检验
F检验和卡方检验都是检验的方法,f_classif用于分类模型,f_regression用于回归模型。
#### (5)互信息法
互信息稀疏反映相关性,互信息越大,说明越相关。
```
# 方差分析
vt = VarianceThreshold(threshold=2)
X_new = vt.fit_transform(X_all, y)
print(vt.variances_)
print(X_new)
print("shape of X_new::", X_new.shape)
# 相关系数法
columns = ['bys_cn', 'hindex_cn', 'a_paper', 'b_paper', 'c_paper', 'papernum2017', 'papernum2016', 'papernum2015', 'papernum2014', 'papernum2013', 'num_journal', 'num_conference', 'degree', 'pagerank', 'degree_centrality', 'diff_year', 'coauthors_top10000', 'coauthors_top20000', 'coauthors_top30000', 'category1', 'category2', 'category3']
X_all_df = pd.DataFrame(X_all, columns=columns)
# print("info of X_all_df::", X_all_df.info())
y_df = pd.DataFrame(y, columns = ['label'])
# print("info of y_df::", y_df.info())
X_y_all_df = pd.concat([X_all_df, y_df], axis=1)
# print("info of X_y_all_df::", X_y_all_df.info())
print(X_y_all_df.corr())
# 卡方检验
k_chi = SelectKBest(chi2, k=15)
X_chi = k_chi.fit_transform(X_all, data['label'])
print(X_chi)
print(k_chi.scores_)
print(k_chi.pvalues_)
# F检验
k_f = SelectKBest(f_classif, k=15)
X_f = k_f.fit_transform(X_all, data['label'])
print(X_f)
print(k_f.scores_)
print(k_f.pvalues_)
# 互信息法
k_info = SelectKBest(mutual_info_classif, k=15)
X_info = k_info.fit_transform(X_all, data['label'])
print(X_info)
print(k_info.scores_)
```
### 4、Wrapper方法
包装法,根据目标函数(通常是预测效果评分),每次选择若干特征,或者排除若干特征。包裹式特征选择直接把最终将要使用的模型的性能作为特征子集的评价标准,也就是说,包裹式特征选择的目的就是为给定的模型选择最有利于其性能的特征子集。
从模型的性能来看,包裹式特征选择比过滤式特征选择更好,但需要多次训练模型,因此计算开销较大。
包括递归特征消除法等。
#### (1)递归特征消除法
递归消除特征法使用一个基模型来进行多轮训练,每轮训练后,消除若干权值系数的特征,再基于新的特征集进行下一轮训练。
```
# RFE
model_lg = RFE(estimator=LogisticRegression(), n_features_to_select=15)
X_lg = model_lg.fit_transform(X_all, data['label'])
print(X_lg)
print(model_lg.n_features_)
print(model_lg.support_)
print(model_lg.ranking_)
# RFECV
model_lg_cv = RFECV(estimator=LogisticRegression(), step=1, cv=StratifiedKFold(n_splits=3), scoring="accuracy")
X_lg_cv = model_lg_cv.fit_transform(X_all, data['label'])
print(X_lg_cv)
print(model_lg_cv.n_features_)
print(model_lg_cv.support_)
print(model_lg_cv.ranking_)
```
### 5、Embedded方法
集成法,先使用某些机器学习的算法和模型进行训练,得到各个特征的权值系数,根据系数从大到小选择特征。类似于Filter方法,但是是通过训练来确定特征的优劣。
包括基于树模型的特征选择法、正则化方法等。
#### (1)基于树模型的特征选择法
树模型中GBDT也可用来作为基模型进行特征选择。
#### (2)基于L1的特征选择法
使用L1范数作为惩罚项的线性模型会得到稀疏解,可以起到特征选择的作用。
```
# 基于树模型的特征选择法
model_gdbc= SelectFromModel(GradientBoostingClassifier())
X_gdbc = model_gdbc.fit_transform(X_all, data['label'])
print(X_gdbc)
print("shape of X_gdbc::", X_gdbc.shape)
# L1正则化
model_lsvc = SelectFromModel(LinearSVC(C=0.01, penalty="l1", dual=False))
X_lsvc = model_lsvc.fit_transform(X_all, data['label'])
print(X_lsvc)
print("shape of X_lsvc::", X_lsvc.shape)
```
| github_jupyter |
```
import numpy as np
import json
with open("data/super_data.json", "r") as f:
super_data = json.load(f)
p_data=super_data['papers']
index_phrase=super_data['index_phrase']
# build pagerank graph
node_num=len(p_data)
bayes_graph=np.zeros((node_num,node_num))
bayes_rank=np.ones(node_num)
bayes_reserve=np.zeros(node_num)
markov_graph=np.zeros((node_num,node_num))
markov_rank=np.ones(node_num)
markov_reserve=np.zeros(node_num)
#node with less citations have high originality
originality=0.0000001
for p in p_data:
index=int(p['index'])
# undirected
markov_weight=np.array(p['all_cite_sim'])
z=originality
for cs in p['all_cite_sim']:
z+=cs
markov_weight/=float(z)
markov_reserve[index]=originality/float(z)
markov_graph[index][index]=originality/float(z)
i=0
for c in p['all_cite']:
markov_graph[int(c)][index]=markov_weight[i]
i+=1
# directed
bayes_weight=np.array(p['citations_sim'])
z=originality
for cs in p['citations_sim']:
z+=cs
bayes_weight/=float(z)
bayes_reserve[index]=originality/float(z)
bayes_graph[index][index]=originality/float(z)
i=0
for c in p['citations']:
bayes_graph[int(c)][index]=bayes_weight[i]
i+=1
bayes_rank=bayes_rank-bayes_reserve
markov_rank=markov_rank-markov_reserve
iterations=500
for i in range(iterations):
bayes_rank=np.dot(bayes_graph,bayes_rank)
for i in range(iterations):
markov_rank=np.dot(markov_graph,markov_rank)
bayes_rank+=bayes_reserve
markov_rank+=markov_reserve
bayes_rank[705]
#sanity check
count=0
for i in range(node_num):
if(bayes_rank[i]>2.6):
count+=1
print '#'+str(count)+' '+p_data[i]['index']
print p_data[i]['title']
print 'score: '+ str(bayes_rank[i])
print ' '
print p_data[i]['abstract']
print ' '
print ' '
for p in p_data:
idx=int(p['index'])
p['markov_rank']=markov_rank[idx]
p['bayes_rank']=bayes_rank[idx]
markov_ranks=np.argsort(markov_rank)[::-1]
bayes_ranks=np.argsort(bayes_rank)[::-1]
super_data['markov_ranks']=markov_ranks.tolist()
super_data['bayes_ranks']=bayes_ranks.tolist()
import networkx as nx
import community
G=nx.Graph()
for p in p_data:
idx=int(p['index'])
count=0
for i in p['citations']:
G.add_edge(idx, int(i),weight=p['citations_sim'][count])
count+=1
c_scores = nx.degree_centrality(G)
c_ranks=np.zeros(len(p_data))
for i in range (len(p_data)):
if i in c_scores:
c_ranks[i]=c_scores[i]
c_ranks=np.argsort(c_ranks)[::-1]
super_data['c_ranks']=c_ranks.tolist()
from operator import itemgetter
partition = community.best_partition(G)
check=set()
for key in partition:
check.add(partition[key])
label_num=len(check)
group=[[] for i in range(label_num)]
for i in range(len(p_data)):
if(i in partition):
p_data[i]['louvain_index']=partition[i]
group[partition[i]].append(i)
else:
p_data[i]['louvain_index']=-1
# build group info
final_group_info=[]
for i in range(label_num):
final_group_info.append({})
final_group_info[i]['nodes']=group[i]
final_group_info[i]['size']=len(group[i])
final_group_info[i]['index']=i
#Top phrase for group
for i in range(label_num):
top_phrase=[]
count=np.zeros(len(index_phrase))
for j in group[i]:
for key in p_data[j]['phrases']:
count[int(key)]+=p_data[j]['phrases'][key]
b=np.argsort(count)[::-1]
for k in range(30):
top_phrase.append(index_phrase[str(b[k])])
final_group_info[i]['top_phrase']=top_phrase
name_str = (top_phrase[0]+', '+top_phrase[1]+' and '+top_phrase[2])
final_group_info[i]['name']=name_str
#build connection
for i in range(label_num):
connected_group=set()
nodes=final_group_info[i]['nodes']
for node in nodes:
for c in p_data[node]['all_cite']:
out_index=p_data[int(c)]['louvain_index']
if(out_index!=index):
connected_group.add(out_index)
final_group_info[i]['connected_group']=connected_group
# get importer, exporter and contribution score
for group in final_group_info:
index=group['index']
#map group to node
importer={}
exporter={}
#map group to number
import_score={}
export_score={}
exchange_score={}
for cg in group['connected_group']:
importer[cg]=set()
exporter[cg]=set()
import_score[cg]=0
export_score[cg]=0
exchange_score[cg]=0
for node in group['nodes']:
for c in p_data[node]['citations']:
out_index=p_data[int(c)]['louvain_index']
if(out_index!=index):
importer[out_index].add(node)
import_score[out_index]+=1
exchange_score[out_index]+=1
for c in p_data[node]['cited_by']:
out_index=p_data[int(c)]['louvain_index']
if(out_index!=index):
exporter[out_index].add(node)
export_score[out_index]+=1
exchange_score[out_index]+=1
for cg in group['connected_group']:
importer[cg]=list(importer[cg])
exporter[cg]=list(exporter[cg])
import_list=[]
for key in import_score:
import_list.append([key, import_score[key]])
export_list=[]
for key in export_score:
export_list.append([key, export_score[key]])
exchange_list=[]
for key in exchange_score:
exchange_list.append([key, exchange_score[key]])
group['import_list']=sorted(import_list,key=itemgetter(1),reverse=True)
group['export_list']=sorted(export_list,key=itemgetter(1),reverse=True)
group['exchange_list']=sorted(exchange_list,key=itemgetter(1),reverse=True)
group['importer']=importer
group['exporter']=exporter
# turn set into list for storage
for group in final_group_info:
group['nodes']=list(group['nodes'])
group['connected_group']=list(group['connected_group'])
super_data['louvain_group']=final_group_info
import os
path = "data/super_data_2.json"
if(os.path.isfile(path)):
os.remove(path)
with open(path, "w") as f:
json.dump(super_data, f)
```
| github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc" style="margin-top: 1em;"><ul class="toc-item"><li><span><a href="#Algorithm-complexity" data-toc-modified-id="Algorithm-complexity-1"><span class="toc-item-num">1 </span>Algorithm complexity</a></span><ul class="toc-item"><li><span><a href="#Search-algorithm" data-toc-modified-id="Search-algorithm-1.1"><span class="toc-item-num">1.1 </span>Search algorithm</a></span><ul class="toc-item"><li><span><a href="#Linear-search" data-toc-modified-id="Linear-search-1.1.1"><span class="toc-item-num">1.1.1 </span>Linear search</a></span></li><li><span><a href="#Binary-search" data-toc-modified-id="Binary-search-1.1.2"><span class="toc-item-num">1.1.2 </span>Binary search</a></span></li></ul></li></ul></li><li><span><a href="#Sorting-algorithms" data-toc-modified-id="Sorting-algorithms-2"><span class="toc-item-num">2 </span>Sorting algorithms</a></span><ul class="toc-item"><li><ul class="toc-item"><li><span><a href="#Bubble-sort" data-toc-modified-id="Bubble-sort-2.0.1"><span class="toc-item-num">2.0.1 </span>Bubble sort</a></span></li><li><span><a href="#Selection-sort" data-toc-modified-id="Selection-sort-2.0.2"><span class="toc-item-num">2.0.2 </span>Selection sort</a></span></li><li><span><a href="#Insertion-sort" data-toc-modified-id="Insertion-sort-2.0.3"><span class="toc-item-num">2.0.3 </span>Insertion sort</a></span></li><li><span><a href="#Merge-Sort" data-toc-modified-id="Merge-Sort-2.0.4"><span class="toc-item-num">2.0.4 </span>Merge Sort</a></span></li><li><span><a href="#Quick-sort" data-toc-modified-id="Quick-sort-2.0.5"><span class="toc-item-num">2.0.5 </span>Quick sort</a></span></li></ul></li></ul></li></ul></div>
Discussion and implementation of basic sorting algorithms
Notebook is adapted from: https://github.com/shik3519/programming-concepts-for-data-science
Content is heavily derived from: http://interactivepython.org/runestone/static/pythonds/index.html
```
import pandas as pd
import numpy as np
import time
import matplotlib.pyplot as plt
%matplotlib inline
```
# Algorithm complexity
`Big O notation` is used in Computer Science to describe the performance or complexity of an algorithm. Big O specifically describes the worst-case scenario, and can be used to describe the execution time required or the space used.
To know more: http://interactivepython.org/runestone/static/pythonds/AlgorithmAnalysis/BigONotation.html

<img src="http://interactivepython.org/runestone/static/pythonds/_images/newplot.png">
Let's understand this through 2 different implementation of search algorithm
## Search algorithm
### Linear search
```
def linear_search(l, target):
for e in l:
if e == target:
return True
return False
l = np.arange(1000)
%%timeit
linear_search(l,999)
```
Time scales linearly with n. So Big-O is $O(n)$
### Binary search
Iterative algo
```
def binarySearchIterative(a, t):
upper = len(a) - 1
lower = 0
while lower <= upper:
middle = (lower + upper) // 2
if t == a[middle]:
return True
else:
if t < a[middle]:
upper = middle - 1
else:
lower = middle + 1
return False
l = np.arange(1000)
%%timeit
binarySearchIterative(l,999)
```
Time scales linearly with n. So Big-O is $O(log(n))$
We can see that binary search is almost 30x faster
We can do binary search in a recursive way too
```
def binarySearchRecursive(a, t):
upper = len(a) - 1
lower = 0
if upper >= 0:
middle = (lower + upper) // 2
if t == a[middle]: return True
if t < a[middle]: return binarySearchRecursive(a[:middle], t)
else: return binarySearchRecursive(a[middle + 1:], t)
return False
%%timeit
binarySearchRecursive(l,999)
```
# Sorting algorithms
```
# What is the worst type of sorting algorithm that you can think of ?
# https://en.wikipedia.org/wiki/Bogosort
# while not isInOrder(deck):
# shuffle(deck)
from random import shuffle
def is_sorted(data) -> bool:
"""Determine whether the data is sorted."""
# linear complexity
return all(data[i] <= data[i + 1] for i in range(len(data) - 1)) # go through all items in collection and check order
def bogosort(data) -> list:
"""Shuffle data until sorted."""
while not is_sorted(data): # well will we ever be done ? :)
shuffle(data) # in place
return data
mydata = [1,6,4,3,-7]
bogodata = bogosort(mydata)
bogodata
%%timeit
bogosort(mydata)
import random
rand100000 = [random.randint(1,1000000) for _ in range(100000)]
rand10000 = [random.randint(1,100000) for _ in range(10000)]
rand1000 = [random.randint(1,100000) for _ in range(1000)]
rand100 = [random.randint(1,1000) for _ in range(100)]
rand10 = [random.randint(1,1000) for _ in range(10)]
rand7 = [random.randint(1,1000) for _ in range(7)]
rand6 = [random.randint(1,1000) for _ in range(6)]
rand6sorted = bogosort(rand6)
rand6
%%timeit
bogosort(rand6)
rand7sorted = bogosort(rand7)
rand7
%%timeit
bogosort(rand7)
rand10sorted = bogosort(rand10)
rand10, rand10sorted
rand6
rand7
rand10
import math
math.factorial(10)
bogo7 = bogosort(rand7)
bogo7
%%timeit
bogosort(rand10)
# so time complexity for Bogosort is (n!) not very practical :)
```
Source: http://interactivepython.org/runestone/static/pythonds/SortSearch/toctree.html
### Bubble sort

$$Complexity: O(n^2)$$
A bubble sort is often considered the most inefficient sorting method since it must exchange items before the final location is known. These “wasted” exchange operations are very costly. However, because the bubble sort makes passes through the entire unsorted portion of the list, it has the capability to do something most sorting algorithms cannot. In particular, if during a pass there are no exchanges, then we know that the list must be sorted. A bubble sort can be modified to stop early if it finds that the list has become sorted. This means that for lists that require just a few passes, a bubble sort may have an advantage in that it will recognize the sorted list and stop.
```
l = [1,2,3,4,32,5,5,66,33,221,34,23,12]
def bubblesort(nums, debug = False):
n = len(nums)
exchange_cnt = 1
while exchange_cnt > 0: # outside loop
exchange_cnt = 0
for i in range(1, n): #inner loop
if nums[i] < nums[i - 1]:
exchange_cnt += 1
nums[i - 1], nums[i] = nums[i], nums[i - 1]
if debug:
print(nums)
return nums
bubblesort(l, debug=True)
ran
bubble1000 = bubblesort(rand1000)
%%timeit
bubblesort(rand1000)
%%timeit
bubblesort(rand10000)
%%time
bubblesort(rand100000)
10_000**2, 100_000**2
```
### Selection sort

$$Complexity: O(n^2)$$
The selection sort improves on the bubble sort by making only one exchange for every pass through the list. In order to do this, a selection sort looks for the largest value as it makes a pass and, after completing the pass, places it in the proper location. As with a bubble sort, after the first pass, the largest item is in the correct place. After the second pass, the next largest is in place. This process continues and requires n−1 passes to sort n items, since the final item must be in place after the (n−1) st pass.
```
l = [1,2,3,4,32,5,5,66,33,221,34,23,12]
def selectionSort(l, debug=False):
n = len(l)
end = n - 1
for j in range(n): ## outer loop
max_ = l[-1 - j]
max_idx = -1 - j
for i in range(end): ## inner loop
if l[i] > max_:
max_ = l[i]
max_idx = i
else:
continue
l[-1 - j], l[max_idx] = l[max_idx], l[-1 - j] # swapping items at the end so max value gets to the end
end = end - 1
if debug:
print(l)
return l
selectionSort(l, debug=True)
```
The benefit of selection over bubble sort is it does one exchange per pass whereas bubble sort can do multiple exchanges.
```
%%timeit
selectionSort(rand1000)
%%timeit
selectionSort(rand10000)
# so 10 x more items and we ge 10x10=100 slower speed
```
### Insertion sort

$$Complexity: O(n^2)$$
```
l = [1,2,3,4,32,5,5,66,33,221,34,23,12]
def insertionSort(l, debug=False):
for i in range(1, len(l)): # outer loop
cval = l[i]
pos = i
while pos > 0 and l[pos - 1] > cval: # inner loop
l[pos],l[pos-1] = l[pos - 1],l[pos]
pos = pos - 1
if debug:
print(l)
return l
insertionSort(l,debug=True)
%%timeit
insertionSort(rand1000)
%%timeit
insertionSort(rand10000)
rand20k = [random.randint(1,1000000) for _ in range(20_000)]
rand20k[:10]
%%timeit
insertionSort(rand20k)
rand20k[:10]
%%time
sorted20k = insertionSort(rand20k)
sorted20k[:5], rand20k[:5]
sorted20k[9000] = 777
sorted20k[12000] = 555
# so not sorted anymore
%%time
insertionSort(sorted20k)
```
### Merge Sort
```
# first implementation idea by https://en.wikipedia.org/wiki/John_von_Neumann
```


$$Complexity: O(nlog(n))$$
```
l = [1,2,3,4,32,5,5,66,33,221,34,23,12]
def mergeSort(alist):
# alist = olist[:] # should be a copy
# print("Splitting ", alist)
if len(alist) > 1:
mid = len(alist) // 2
lefthalf = alist[:mid]
righthalf = alist[mid:]
mergeSort(lefthalf)
mergeSort(righthalf)
# this could a seperate function the mergin part
i = 0
j = 0
k = 0
while i < len(lefthalf) and j < len(righthalf):
if lefthalf[i] < righthalf[j]:
alist[k] = lefthalf[i]
i = i + 1
else:
alist[k] = righthalf[j]
j = j + 1
k = k + 1
while i < len(lefthalf):
alist[k] = lefthalf[i]
i = i + 1
k = k + 1
while j < len(righthalf):
alist[k] = righthalf[j]
j = j + 1
k = k + 1
# print("Merging ", alist)
return alist
mergeSort(l)
rand1000[:10]
shuffle(rand1000)
rand1000[:10]
merge1000 = mergeSort(rand1000)
merge1000[:20]
merge10k = mergeSort(rand10000)
merge10k[:10]
%%timeit
mergeSort(rand1000)
%%timeit
mergeSort(rand10000)
r100_000 = [random.randint(1,1_000_000) for _ in range(100_000)]
r100_000[:5]
%%timeit
mergeSort(r100_000) # so here n log n starts to really shine
shuffle(r100_000)
%%timeit
sorted(r100_000) # sorted uses timsort which is a combination of insertion sort + merge sort
sorted100k = mergeSort(r100_000)
sorted100k[:5]
%%timeit
mergeSort(sorted100k) # so we gained nothing from being sorted
%%timeit
insertionSort(sorted100k) # insertion sort should be slow we shall see
sorted100k[50000:50000+10]
sorted100k[555] = 400000
sorted100k[9000] = 1333
%%timeit
insertionSort(sorted100k)
```
# Big O, Theta and Omega bounds
### Merge sort is defined by recurrence formula
T(n) = 2T(n/2) + n
So each problem has to be divided in two halfs and also we have linear(n) merging operation
```
# so how to prove that merge sort is really O(n log n) time complexity?
# in reality we are looking for tight bound the Θ(n log n) complexity
# so O is very loose, in every day usage when people say O they really mean Θ - theta
# O is showing that the algorith is no worse than some f(n)
# I could say that merge sort is O(n!) and it would still be correct but practically useless
# since most algorithms are O(n!)
# so Merge sort is O(n!), O(n^5),O(n^2) and so on and finally most crucially O(n log n)
# Merge sort is NOT O(n)
# so thats what Θ(n log n)
## So ideas on how to prove mergesort is O(n log n) ?
## Instinctively we see that we are dividing in halves and solving the problem for those
# There is something called Master Theorem which lets us quickly see the solution for most types of recurrence
# what is a reccurence relation then?
```
# given n is our data
# so merge sort the reccurence will be
T(n) = 2(T(n/2)) + n # because we have to merge in n time the halves
# so reccurence defines the recursive function
```
# so for next week we will look at solving this and also the Master Theorem on how to generally
# find the complexity
# we do not need the reccurrence if we have regular loops without recursion..
```
### Quick sort

$$Complexity: O(nlog(n))$$ $$Worst case : O(n^2)$$
```
def quickSort(alist):
quickSortHelper(alist, 0, len(alist) - 1)
def quickSortHelper(alist, first, last):
if first < last:
splitpoint = partition(alist, first, last)
quickSortHelper(alist, first, splitpoint - 1)
quickSortHelper(alist, splitpoint + 1, last)
def partition(alist, first, last):
pivotvalue = alist[first]
leftmark = first + 1
rightmark = last
done = False
while not done:
while leftmark <= rightmark and alist[leftmark] <= pivotvalue:
leftmark = leftmark + 1
while alist[rightmark] >= pivotvalue and rightmark >= leftmark:
rightmark = rightmark - 1
if rightmark < leftmark:
done = True
else:
temp = alist[leftmark]
alist[leftmark] = alist[rightmark]
alist[rightmark] = temp
temp = alist[first]
alist[first] = alist[rightmark]
alist[rightmark] = temp
return rightmark
alist = [54, 26, 93, 17, 77, 31, 44, 55, 20]
quickSort(alist)
print(alist)
```
# References and useful links:
* Visualization of these concepst : https://visualgo.net/en
```
```
| github_jupyter |
# Measures of Dispersion
By Evgenia "Jenny" Nitishinskaya, Maxwell Margenot, and Delaney Mackenzie.
Part of the Quantopian Lecture Series:
* [www.quantopian.com/lectures](https://www.quantopian.com/lectures)
* [github.com/quantopian/research_public](https://github.com/quantopian/research_public)
---
Dispersion measures how spread out a set of data is. This is especially important in finance because one of the main ways risk is measured is in how spread out returns have been historically. If returns have been very tight around a central value, then we have less reason to worry. If returns have been all over the place, that is risky.
Data with low dispersion is heavily clustered around the mean, while data with high dispersion indicates many very large and very small values.
Let's generate an array of random integers to work with.
```
# Import libraries
import numpy as np
np.random.seed(121)
# Generate 20 random integers < 100
X = np.random.randint(100, size=20)
# Sort them
X = np.sort(X)
print 'X: %s' %(X)
mu = np.mean(X)
print 'Mean of X:', mu
```
# Range
Range is simply the difference between the maximum and minimum values in a dataset. Not surprisingly, it is very sensitive to outliers. We'll use `numpy`'s peak to peak (ptp) function for this.
```
print 'Range of X: %s' %(np.ptp(X))
```
# Mean Absolute Deviation (MAD)
The mean absolute deviation is the average of the distances of observations from the arithmetic mean. We use the absolute value of the deviation, so that 5 above the mean and 5 below the mean both contribute 5, because otherwise the deviations always sum to 0.
$$ MAD = \frac{\sum_{i=1}^n |X_i - \mu|}{n} $$
where $n$ is the number of observations and $\mu$ is their mean.
```
abs_dispersion = [np.abs(mu - x) for x in X]
MAD = np.sum(abs_dispersion)/len(abs_dispersion)
print 'Mean absolute deviation of X:', MAD
```
# Variance and standard deviation
The variance $\sigma^2$ is defined as the average of the squared deviations around the mean:
$$ \sigma^2 = \frac{\sum_{i=1}^n (X_i - \mu)^2}{n} $$
This is sometimes more convenient than the mean absolute deviation because absolute value is not differentiable, while squaring is smooth, and some optimization algorithms rely on differentiability.
Standard deviation is defined as the square root of the variance, $\sigma$, and it is the easier of the two to interpret because it is in the same units as the observations.
```
print 'Variance of X:', np.var(X)
print 'Standard deviation of X:', np.std(X)
```
One way to interpret standard deviation is by referring to Chebyshev's inequality. This tells us that the proportion of samples within $k$ standard deviations (that is, within a distance of $k \cdot$ standard deviation) of the mean is at least $1 - 1/k^2$ for all $k>1$.
Let's check that this is true for our data set.
```
k = 1.25
dist = k*np.std(X)
l = [x for x in X if abs(x - mu) <= dist]
print 'Observations within', k, 'stds of mean:', l
print 'Confirming that', float(len(l))/len(X), '>', 1 - 1/k**2
```
The bound given by Chebyshev's inequality seems fairly loose in this case. This bound is rarely strict, but it is useful because it holds for all data sets and distributions.
# Semivariance and semideviation
Although variance and standard deviation tell us how volatile a quantity is, they do not differentiate between deviations upward and deviations downward. Often, such as in the case of returns on an asset, we are more worried about deviations downward. This is addressed by semivariance and semideviation, which only count the observations that fall below the mean. Semivariance is defined as
$$ \frac{\sum_{X_i < \mu} (X_i - \mu)^2}{n_<} $$
where $n_<$ is the number of observations which are smaller than the mean. Semideviation is the square root of the semivariance.
```
# Because there is no built-in semideviation, we'll compute it ourselves
lows = [e for e in X if e <= mu]
semivar = np.sum( (lows - mu) ** 2 ) / len(lows)
print 'Semivariance of X:', semivar
print 'Semideviation of X:', np.sqrt(semivar)
```
A related notion is target semivariance (and target semideviation), where we average the distance from a target of values which fall below that target:
$$ \frac{\sum_{X_i < B} (X_i - B)^2}{n_{<B}} $$
```
B = 19
lows_B = [e for e in X if e <= B]
semivar_B = sum(map(lambda x: (x - B)**2,lows_B))/len(lows_B)
print 'Target semivariance of X:', semivar_B
print 'Target semideviation of X:', np.sqrt(semivar_B)
```
# These are Only Estimates
All of these computations will give you sample statistics, that is standard deviation of a sample of data. Whether or not this reflects the current true population standard deviation is not always obvious, and more effort has to be put into determining that. This is especially problematic in finance because all data are time series and the mean and variance may change over time. There are many different techniques and subtleties here, some of which are addressed in other lectures in the [Quantopian Lecture Series](https://www.quantopian.com/lectures).
In general do not assume that because something is true of your sample, it will remain true going forward.
## References
* "Quantitative Investment Analysis", by DeFusco, McLeavey, Pinto, and Runkle
*This presentation is for informational purposes only and does not constitute an offer to sell, a solicitation to buy, or a recommendation for any security; nor does it constitute an offer to provide investment advisory or other services by Quantopian, Inc. ("Quantopian"). Nothing contained herein constitutes investment advice or offers any opinion with respect to the suitability of any security, and any views expressed herein should not be taken as advice to buy, sell, or hold any security or as an endorsement of any security or company. In preparing the information contained herein, Quantopian, Inc. has not taken into account the investment needs, objectives, and financial circumstances of any particular investor. Any views expressed and data illustrated herein were prepared based upon information, believed to be reliable, available to Quantopian, Inc. at the time of publication. Quantopian makes no guarantees as to their accuracy or completeness. All information is subject to change and may quickly become unreliable for various reasons, including changes in market conditions or economic circumstances.*
| github_jupyter |
# Introduction to Machine Learning
This lab introduces some basic concepts of machine learning with Python. In this lab you will use the K-Nearest Neighbor (KNN) algorithm to classify the species of iris flowers, given measurements of flower characteristics. By completing this lab you will have an overview of an end-to-end machine learning modeling process.
By the completion of this lab, you will:
1. Follow and understand a complete end-to-end machine learning process including data exploration, data preparation, modeling, and model evaluation.
2. Develop a basic understanding of the principles of machine learning and associated terminology.
3. Understand the basic process for evaluating machine learning models.
## Overview of KNN classification
Before discussing a specific algorithm, it helps to know a bit of machine learning terminology. In supervised machine learning a set of ***cases*** are used to ***train***, ***test*** and ***evaluate*** the model. Each case is comprised of the values of one or more ***features*** and a ***label*** value. The features are variables used by the model to ***predict** the value of the label. Minimizing the ***errors*** between the true value of the label and the prediction supervises the training of this model. Once the model is trained and tested, it can be evaluated based on the accuracy in predicting the label of a new set of cases.
In this lab you will use randomly selected cases to first train and then evaluate a k-nearest-neighbor (KNN) machine learning model. The goal is to predict the type or class of the label, which makes the machine learning model a ***classification*** model.
The k-nearest-neighbor algorithm is conceptually simple. In fact, there is no formal training step. Given a known set of cases, a new case is classified by majority vote of the K (where $k = 1, 2, 3$, etc.) points nearest to the values of the new case; that is, the nearest neighbors of the new case.
The schematic figure below illustrates the basic concepts of a KNN classifier. In this case there are two features, the values of one shown on the horizontal axis and the values of the other shown on the vertical axis. The cases are shown on the diagram as one of two classes, red triangles and blue circles. To summarize, each case has a value for the two features, and a class. The goal of the KNN algorithm is to classify cases with unknown labels.
Continuing with the example, on the left side of the diagram the $K = 1$ case is illustrated. The nearest neighbor is a red triangle. Therefore, this KNN algorithm will classify the unknown case, '?', as a red triangle. On the right side of the diagram, the $K = 3$ case is illustrated. There are three near neighbors within the circle. The majority of nearest neighbors for $K = 3$ are the blue circles, so the algorithm classifies the unknown case, '?', as a blue circle. Notice that class predicted for the unknown case changes as K changes. This behavior is inherent in the KNN method.

<center> **KNN for k = 1 and k = 3**</center>
There are some additional considerations in creating a robust KNN algorithm. These will be addressed later in this course.
## Examine the data set
In this lab you will work with the Iris data set. This data set is famous in the history of statistics. The first publication using these data in statistics by the pioneering statistician Ronald A Fisher was in 1936. Fisher proposed an algorithm to classify the species of iris flowers from physical measurements of their characteristics. The data set has been used as a teaching example ever since.
Now, you will load and examine these data which are in the statsmodels.api package. Execute the code in the cell below and examine the first few rows of the data frame.
```
import pandas as pd
#from statsmodels.api import datasets
from sklearn import datasets ## Get dataset from sklearn
## Import the dataset from sklearn.datasets
iris = datasets.load_iris()
## Create a data frame from the dictionary
species = [iris.target_names[x] for x in iris.target]
print(species)
iris = pd.DataFrame(iris['data'], columns = ['Sepal_Length', 'Sepal_Width', 'Petal_Length', 'Petal_Width'])
iris['Species'] = species
```
There are four features, containing the dimensions of parts of the iris flower structures. The label column is the Species of the flower. The goal is to create and test a KNN algorithm to correctly classify the species.
Next, you will execute the code in the cell below to show the data types of each column.
```
iris.dtypes
```
The features are all numeric, and the label is a categorical string variable.
Next, you will determine the number of unique categories, and number of cases for each category, for the label variable, Species. Execute the code in the cell below and examine the results.
```
iris['count'] = 1
iris[['Species', 'count']].groupby('Species').count()
```
You can see there are three species of iris, each with 50 cases.
Next, you will create some plots to see how the classes might, or might not, be well separated by the value of the features. In an ideal case, the label classes will be perfectly separated by one or more of the feature pairs. In the real-world this ideal situation will rarely, if ever, be the case.
There are six possible pair-wise scatter plots of these four features. For now, we will just create scatter plots of two variable pairs. Execute the code in the cell below and examine the resulting plots.
***
**Note:** Data visualization and the Seaborn package are covered in another lesson.
***
```
%matplotlib inline
def plot_iris(iris, col1, col2):
import seaborn as sns
import matplotlib.pyplot as plt
sns.lmplot(x = col1, y = col2,
data = iris,
hue = "Species", #hue is to denote in diff color and showing legends onto the graph
fit_reg = False)
plt.xlabel(col1)
plt.ylabel(col2)
plt.title('Iris species shown by color')
plt.show()
plot_iris(iris, 'Petal_Width', 'Sepal_Length')
plot_iris(iris, 'Sepal_Width', 'Sepal_Length')
```
Examine these results noticing the separation, or overlap, of the label values.
Then, answer **Question 1** on the course page.
## Prepare the data set
Data preparation is an important step before training any machine learning model. These data require only two preparation steps:
- Scale the numeric values of the features. It is important that numeric features used to train machine learning models have a similar range of values. Otherwise, features which happen to have large numeric values may dominate model training, even if other features with smaller numeric values are more informative. In this case Zscore normalization is used. This normalization process scales each feature so that the mean is 0 and the variance is 1.0.
- Split the dataset into randomly sampled training and evaluation data sets. The random selection of cases seeks to limit the leakage of information between the training and evaluation cases.
The code in the cell below normalizes the features by these steps:
- The scale function from scikit-learn.preprocessing is used to normalize the features.
- Column names are assigned to the resulting data frame.
- A statitical summary of the data frame is then printed.
***
**Note:** Data preparation with scikit-learn is covered in another lesson.
***
Execute this code and examine the results.
```
from sklearn.preprocessing import scale
import pandas as pd
num_cols = ['Sepal_Length', 'Sepal_Width', 'Petal_Length', 'Petal_Width']
iris_scaled = scale(iris[num_cols])
iris_scaled = pd.DataFrame(iris_scaled, columns = num_cols)
print(iris_scaled.describe().round(3))
```
Examine these results. The mean of each column is zero and the standard deviation is approximately 1.0.
The methods in the scikit-learn package requires numeric numpy arrays as arguments. Therefore, the strings indicting species must be re-coded as numbers. The code in the cell below does this using a dictionary lookup. Execute this code and examine the head of the data frame.
```
levels = {'setosa':0, 'versicolor':1, 'virginica':2}
iris_scaled['Species'] = [levels[x] for x in iris['Species']]
iris_scaled.head()
```
Now, you will split the dataset into a test and evaluation sub-sets. The code in the cell below randomly splits the dataset into training and testing subsets. The features and labels are then separated into numpy arrays. The dimension of each array is printed as a check. Execute this code to create these subsets.
***
**Note:** Splitting data sets for machine learning with scikit-learn is discussed in another lesson.
***
```
## Split the data into a training and test set by Bernoulli sampling
from sklearn.model_selection import train_test_split
import numpy as np
np.random.seed(3456)
iris_split = train_test_split(np.asmatrix(iris_scaled), test_size = 75)
iris_train_features = iris_split[0][:, :4]
iris_train_labels = np.ravel(iris_split[0][:, 4])
iris_test_features = iris_split[1][:, :4]
iris_test_labels = np.ravel(iris_split[1][:, 4])
print(iris_train_features.shape)
print(iris_train_labels.shape)
print(iris_test_features.shape)
print(iris_test_labels.shape)
```
## Train and evaluate the KNN model
With some understanding of the relationships between the features and the label and preparation of the data completed you will now train and evaluate a $K = 3$ model. The code in the cell below does the following:
- The KNN model is defined as having $K = 3$.
- The model is trained using the fit method with the feature and label numpy arrays as arguments.
- Displays a summary of the model.
Execute this code and examine the summary of these results.
***
**Note:** Constructing machine learning models with scikit-learn is covered in another lesson.
***
```
## Define and train the KNN model
from sklearn.neighbors import KNeighborsClassifier
KNN_mod = KNeighborsClassifier(n_neighbors = 3)
KNN_mod.fit(iris_train_features, iris_train_labels)
```
Next, you will evaluate this model using the accuracy statistic and a set of plots. The following steps create model predictions and compute accuracy:
- The predict method is used to compute KNN predictions from the model using the test features as an argument.
- The predictions are scored as correct or not using a list comprehension.
- Accuracy is computed as the percentage of the test cases correctly classified.
Execute this code, examine the results, and answer **Question 2** on the course page.
```
iris_test = pd.DataFrame(iris_test_features, columns = num_cols)
iris_test['predicted'] = KNN_mod.predict(iris_test_features)
iris_test['correct'] = [1 if x == z else 0 for x, z in zip(iris_test['predicted'], iris_test_labels)]
accuracy = 100.0 * float(sum(iris_test['correct'])) / float(iris_test.shape[0])
print(accuracy)
iris_test
```
The accuracy is pretty good.
Now, execute the code in the cell below and examine plots of the classifications of the iris species.
```
levels = {0:'setosa', 1:'versicolor', 2:'virginica'}
iris_test['Species'] = [levels[x] for x in iris_test['predicted']]
markers = {1:'^', 0:'o'}
colors = {'setosa':'blue', 'versicolor':'green', 'virginica':'red'}
def plot_shapes(df, col1,col2, markers, colors):
import matplotlib.pyplot as plt
import seaborn as sns
ax = plt.figure(figsize=(6, 6)).gca() # define plot axis
for m in markers: # iterate over marker dictionary keys
for c in colors: # iterate over color dictionary keys
df_temp = df[(df['correct'] == m) & (df['Species'] == c)]
print(df_temp)
sns.regplot(x = col1, y = col2,
data = df_temp,
fit_reg = False,
scatter_kws={'color': colors[c]},
marker = markers[m],
ax = ax)
plt.xlabel(col1)
plt.ylabel(col2)
plt.title('Iris species by color')
return 'Done'
plot_shapes(iris_test, 'Petal_Width', 'Sepal_Length', markers, colors)
plot_shapes(iris_test, 'Sepal_Width', 'Sepal_Length', markers, colors)
```
In the plots above color is used to show the predicted class. Correctly classified cases are shown by triangles and incorrectly classified cases are shown by circles.
Examine the plot and answer **Question 3** on the course page.
## Summary
In this lab you have created and evaluated a KNN machine learning classification model. Specifically you have:
1. Loaded and explored the data using visualization to determine if the features separate the classes.
2. Prepared the data by normalizing the numeric features and randomly sampling into training and testing subsets.
3. Constructing and evaluating the machine learning model. Evaluation was performed by statistically, with the accuracy metric, and with visualization.
| github_jupyter |
# Missing Value Treatment
```
import os
os.chdir(r'D:\D\Cognitior\Courses\Dataset\Python DataScience')
import pandas as pd
import numpy as np
import seaborn as sns
dataset = pd.read_excel('Employee_Data.xls')
dataset.isnull().sum()
sns.boxplot(y='Age',data=dataset)
sns.boxplot(y='Experience', data=dataset)
x=dataset.iloc[:,3:6].values
x
from sklearn.impute import SimpleImputer
imp =SimpleImputer( strategy = 'most_frequent')
x[:,0:1] = imp.fit_transform(x[:,0:1])
x
from sklearn.impute import SimpleImputer
imp =SimpleImputer( strategy = 'mean')
x[:,1:3] = imp.fit_transform(x[:,1:3])
x
pd.DataFrame(x)
dataset.isnull().sum()
dataset['Age'] = dataset['Age'].fillna(dataset['Age'].mean())
dataset['Experience'] = dataset['Experience'].fillna(dataset['Experience'].median())
dataset.groupby('Department').size()
dataset['Department'] = dataset['Department'].fillna('Sales and Marketing')
```
# Outlier Treatment
```
os.chdir(r'D:\D\Cognitior\Courses\Data Science - Course - Reusable component\Outlier Treatment')
dataset_ot = pd.read_excel('OutlierData.xlsx')
dataset_ot
Q1 = dataset_ot.quantile(0.25)
Q3 = dataset_ot.quantile(0.75)
Q1
Q3
IQR = Q3-Q1
IQR
dataset_ot[~((dataset_ot<(Q1-1.5*IQR))|
(dataset_ot>(Q3 + 1.5*IQR))).any(axis=1)]
```
# Feature Scaling
```
os.chdir(r'D:\D\Trainings\R and Python Classes\Machine Learning A-Z\Part 1 - Data Preprocessing\Section 2 -------------------- Part 1 - Data Preprocessing --------------------\Data_Preprocessing')
dataset_fs = pd.read_csv('data.csv')
x=dataset_fs.iloc[:].values
x
from sklearn.preprocessing import StandardScaler
sc_x = StandardScaler()
x[:,1:3] = sc_x.fit_transform(x[:,1:3])
x
from sklearn.preprocessing import Normalizer
nm_x = Normalizer()
x[:,1:3] = nm_x.fit_transform(x[:,1:3])
```
# Encoding
```
x
from sklearn.preprocessing import LabelEncoder, OneHotEncoder
labelencoder_x = LabelEncoder()
x[:,0] = labelencoder_x.fit_transform(x[:,0])
x[:,3] = labelencoder_x.fit_transform(x[:,3])
onehotencoder = OneHotEncoder(categorical_features=[0])
x = onehotencoder.fit_transform(x).toarray()
pd.DataFrame(x)
```
# Simple Linear Regression
```
os.chdir(r'D:\D\Edureka\Edureka - 24 June - Python\Class 9')
dataset=pd.read_csv('Salary_Data.csv')
x=dataset.iloc[:,0:1].values
y = dataset.iloc[:,1].values
y
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(x,y,test_size=0.3,
random_state=10)
from sklearn.linear_model import LinearRegression
regressor = LinearRegression()
regressor.fit(x_train, y_train)
regressor.coef_
regressor.intercept_
salary = 9303*Exp + 27206
y_pred=regressor.predict(x_test)
y_test
from sklearn.metrics import r2_score
r2_score(y_test,y_pred)
x_new = pd.read_excel('new.xlsx')
x_new = x_new.iloc[:,0:1].values
regressor.predict(x_new)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/mashnoor3/data-science-portfolio/blob/main/Super_Bowl_Exploratory_Data_Analysis.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Super Bowl - Exploratory Data Analysis
In this notebook we will look at historical data for superbowls, including game scores, tv ratings, and halftime performances. The goal is to understand how some of the elements interact with each other. Mainly we are going to answer questions like:
- What are the most extreme game outcomes?
- How does the game affect television viewership?
- How have viewership, TV ratings, and ad cost evolved over time?
- Who are the most prolific musicians in terms of halftime show performances?
```
import pandas as pd
from matplotlib import pyplot as plt
%matplotlib inline
import seaborn as sns
!pip -q install fsspec # File-system specification
!pip -q install gcsfs # Convenient Filesystem interface over GCS
gs_bucket = 'gs://mashnoor-data-science-portfolio/'
super_bowls = pd.read_csv(gs_bucket + 'super_bowls.csv', encoding='utf-8')
tv = pd.read_csv(gs_bucket + 'tv.csv', encoding='utf-8')
halftime_musicians = pd.read_csv(gs_bucket + 'halftime_musicians.csv', encoding='utf-8')
display(super_bowls.head())
display(tv.head())
display(halftime_musicians.head())
```
# Missing data
From looking at just the head of the super bowl dataset, we can note missing data for qb_winner_2 and qb_loser_2. There must be other missing data as well. Let's inspect the null data, and deal with them.
```
super_bowls.info()
print('\n')
tv.info()
print('\n')
halftime_musicians.info()
```
# Combined points distribution
For the TV data, the following columns have missing values and a lot of them:
- total_us_viewers (amount of U.S. viewers who watched at least some part of the broadcast)
- rating_18_49 (average % of U.S. adults 18-49 who live in a household with a TV that were watching for the entire broadcast)
- share_18_49 (average % of U.S. adults 18-49 who live in a household with a TV in use that were watching for the entire broadcast)
For the halftime musician data, there are missing numbers of songs performed (num_songs) for about a third of the performances.
Let's take note of where the dataset isn't perfect and start uncovering some insights.
Let's start by looking at combined points for each Super Bowl by visualizing the distribution. Let's also pinpoint the Super Bowls with the highest and lowest scores.
```
plt.style.use('seaborn')
# Plot a histogram of combined points
plt.hist(super_bowls['combined_pts'])
plt.xlabel('Combined Points')
plt.ylabel('Number of Super Bowls')
plt.show()
# Display the Super Bowls with the highest and lowest combined scores
display(super_bowls[super_bowls['combined_pts'] > 70])
display(super_bowls[super_bowls['combined_pts'] < 25])
```
# Point difference distribution
Most combined scores are around 40-50 points, with the extremes being roughly equal distance away in opposite directions. Going up to the highest combined scores at 74 and 75, we find two games featuring dominant quarterback performances. One even happened recently in 2018's Super Bowl LII where Tom Brady's Patriots lost to Nick Foles' underdog Eagles 41-33 for a combined score of 74.
Going down to the lowest combined scores, we have Super Bowl III and VII, which featured tough defenses that dominated. We also have Super Bowl IX in New Orleans in 1975, whose 16-6 score can be attributed to inclement weather. The field was slick from overnight rain, and it was cold at 46 °F (8 °C), making it hard for the Steelers and Vikings to do much offensively. This was the second-coldest Super Bowl ever and the last to be played in inclement weather for over 30 years. The NFL realized people like points, I guess.
UPDATE: In Super Bowl LIII in 2019, the Patriots and Rams broke the record for the lowest-scoring Super Bowl with a combined score of 16 points (13-3 for the Patriots).
Let's take a look at point difference now.
```
# Histogram of point differences
fig, ax = plt.subplots(figsize=(10, 10))
plt.hist(super_bowls.difference_pts)
plt.xlabel('Point Difference')
plt.ylabel('Number of Super Bowls')
plt.show()
# Displaying the closest game(s) and biggest blowouts
display(super_bowls.difference_pts.max())
display(super_bowls.difference_pts.min())
```
# Do large point differences (blowout games) lead to lost viewers?
Let's combine our game data and TV to see if this is the case. Do large point differences translate to lost viewers? We can plot household share (average percentage of U.S. households with a TV in use that were watching for the entire broadcast) vs. point difference to find out.
```
# Join game and TV data, filtering out super bowl 1 because it was split over two networks
games_tv = pd.merge(tv[tv['super_bowl'] > 1], super_bowls, on='super_bowl')
display(games_tv.head())
# Vislualizing with linear regression
# fig, ax = plt.subplots(figsize=(15, 10))
plt.figure(figsize = (15,10))
sns.regplot(x=games_tv['difference_pts'], y=games_tv['share_household'], data=games_tv)
plt.show()
```
# Viewership and the ad industry over time
The downward sloping regression line and the 95% confidence interval for that regression suggest that bailing on the game if it is a blowout is common. Though it matches our intuition, we must take it with a grain of salt because the linear relationship in the data is weak due to our small sample size of 52 games.
Regardless of the score though, I bet most people stick it out for the halftime show, which is good news for the TV networks and advertisers. A 30-second spot costs a pretty [$5 million](https://www.businessinsider.com/super-bowl-commercials-cost-more-than-eagles-quarterback-earns-2018-1) now, but has it always been that way? And how have number of viewers and household ratings trended alongside ad cost? We can find out using line plots that share a "Super Bowl" x-axis.
From the plots, below we can see that viewers increased before ad costs did. Maybe the networks weren't very data savvy and were slow to react?
```
plt.figure(figsize = (10,6))
# Create a figure with 3x1 subplot and activate the top subplot
plt.subplot(3, 1, 1)
plt.plot(games_tv['super_bowl'], games_tv['avg_us_viewers'], color='#648FFF')
plt.title('Average Number of US Viewers', fontsize=13)
# # Activate the middle subplot
plt.subplot(3, 1, 2)
plt.plot(games_tv['super_bowl'], games_tv['rating_household'], color='#DC267F')
plt.title('Household Rating', fontsize=13)
# Activate the bottom subplot
plt.subplot(3, 1, 3)
plt.plot(games_tv['super_bowl'], games_tv['ad_cost'], color='#FFB000')
plt.title('Ad Cost', fontsize=13)
plt.xlabel('SUPER BOWL')
# Improve the spacing between subplots
plt.tight_layout()
```
# Analysis of halftime shows
Another hypothesis: maybe halftime shows weren't that good in the earlier years? The modern spectacle of the Super Bowl has a lot to do with the cultural prestige of big halftime acts.
It turns out Michael Jackson's Super Bowl XXVII performance, one of the most watched events in American TV history, was when the NFL realized the value of Super Bowl airtime and decided they needed to sign big name acts from then on out. The halftime shows before MJ indeed weren't that impressive, which we can see by filtering our halftime_musician data.
```
display(halftime_musicians[halftime_musicians['super_bowl']<=27])
```
# Musicians with most halftime show appearances
Let's see all of the musicians that have done more than one halftime show, including their performance counts.
```
# Count halftime show appearances for each musician and sort them from most to least
halftime_appearances = halftime_musicians.groupby('musician').count()
halftime_appearances = halftime_appearances.sort_values('super_bowl', ascending=False)
display(halftime_appearances[halftime_appearances['super_bowl']>1])
```
# Who performed the most songs in a halftime show?
The world famous Grambling State University Tiger Marching Band takes the crown with six appearances. Beyoncé, Justin Timberlake, Nelly, and Bruno Mars are the only post-Y2K musicians with multiple appearances (two each).
From our previous inspections, the num_songs column has lots of missing values:
A lot of the marching bands don't have num_songs entries.
For non-marching bands, missing data starts occurring at Super Bowl XX.
Let's filter out marching bands by filtering out musicians with the word "Marching" in them and the word "Spirit" (a common naming convention for marching bands is "Spirit of [something]"). Then we'll filter for Super Bowls after Super Bowl XX to address the missing data issue, then let's see who has the most number of songs.
```
# Filter out most marching bands
no_bands = halftime_musicians[~halftime_musicians['musician'].str.contains('Marching')]
no_bands = no_bands[~no_bands['musician'].str.contains('Spirit')]
display(no_bands.head())
# Plot a histogram of number of songs per performance
most_songs = int(max(no_bands['num_songs'].values))
plt.figure(figsize = (15,10))
plt.hist(no_bands.num_songs.dropna(), bins=most_songs)
plt.xlabel('Number of Songs Per Halftime Show Performance')
plt.ylabel('Number of Musicians')
plt.title('Distribution of Number of Songs for Musicians', fontsize=20)
plt.show()
# Sort the non-band musicians by number of songs per appearance...
no_bands = no_bands.sort_values('num_songs', ascending=False)
# Display the top 15
display(no_bands.head(15))
```
# Conclusion
So most non-band musicians do 1-3 songs per halftime show. It's important to note that the duration of the halftime show is fixed (roughly 12 minutes) so songs per performance is more a measure of how many hit songs you have. JT went off in 2018, wow. 11 songs! Diana Ross comes in second with 10 in her medley in 1996.
In this notebook, we loaded, cleaned, then explored Super Bowl game, television, and halftime show data. We visualized the distributions of combined points, point differences, and halftime show performances using histograms. We used line plots to see how ad cost increases lagged behind viewership increases. And we discovered that blowouts do appear to lead to a drop in viewers.
| github_jupyter |
## AREBO Link Length Optimization
Notebook to carry out AREBO's link length optimization through a brute force parameter search.
```
%pylab inline
import itertools
import functools
import pandas as pd
import progressbar as pb
import sys
import json
import os
sys.path.append("scripts")
from ipywidgets import widgets
from ipywidgets import interactive
from mpl_toolkits.mplot3d import Axes3D
import areboopt as aropt
```
---
```
# Output data foler
outdir = "opt_data"
# Defining robot parameter ranges
d2r = np.pi / 180.
r2d = 180. / np.pi
robot_param = {'links': {'r1': [20., 30.], 'r2': [10., 20.],
'r3': [10., 15.], 'dr': 1.},
'angles': {'t1': [d2r * -180., d2r * 180.],
't2': [d2r * -60., d2r * 180.],
't3': [d2r * -180., d2r * 180.],
't4': [d2r * -180., d2r * 180.],
't5': [d2r * -180., d2r * 180.],
't6': [d2r * -180., d2r * 180.]}
}
# Save parameters
with open(f"{outdir}/robot_param_settings.json", "w") as f:
json.dump(robot_param, fp=f)
# Defining human parameter ranges
hlimb_param = {'links': {'uh': [15., 20.], 'dr': 2.5},
'angles': {'t1': [d2r * - 120, d2r * 0],
't2': [d2r * -50, d2r * 50],
'dt': 10 * np.pi / 180},
'loc': {'x': [-10., 0, 10.],
'y': [-10., 0, 10.],
'z': [10., 20., 30.]}
}
# Save parameters
# Save parameters
with open(f"{outdir}/hlimb_param_settings.json", "w") as f:
json.dump(hlimb_param, fp=f)
# Optimization summary array.
goodarray = pd.DataFrame(columns=("r1", "r2", "r3", "shx", "shy", "shz",
"uh", "g1", "g2", "g", "dataf"))
# All robot and human limb parametes.
all_hparam = np.array(aropt.iterate_over_hlimb_param(hlimb_param))
all_rparam = np.array(aropt.iterate_over_robotparam(robot_param))
# Save all robot and human parameters.
with open(f"{outdir}/all_hlimb_param.npy", 'wb') as f:
np.save(f, all_hparam)
with open(f"{outdir}/all_robot_param.npy", 'wb') as f:
np.save(f, all_rparam)
# Go through all human limb angles.
all_hangles, pha1, pha2 = aropt.all_hlimb_angles(hlimb_param)
Nha1, Nha2 = len(pha1), len(pha2)
# Go through each robot parameter combination
Nr = len(all_rparam)
Nh = len(all_hparam)
# Progress bar
# define progress bar
widgets = ['[', pb.Timer(), '] ', pb.Bar(), ' (', pb.ETA(), ')']
bar = pb.ProgressBar(widgets=widgets, maxval=Nr * Nh).start()
for ir, _rparam in enumerate(all_rparam):
for ih, _hparam in enumerate(all_hparam):
# Shoulder position and upper-arm length.
_shpos, _uh = _hparam[:3], _hparam[3]
# Base file name.
basename = "_".join(("r3s3l1",
f"{int(_rparam[0])}", f"{int(_rparam[1])}",
f"{int(_rparam[2])}", f"{int(_shpos[0])}",
f"{int(_shpos[0])}", f"{int(_shpos[0])}",
f"{int(_uh)}"))
# Get the inverse kinematic map
invkin_maps = aropt.get_invkinmaps(_rparam, _uh, _shpos, all_hangles,
Nha1, Nha2, robot_param)
# Compute "goodness" function
gvals = aropt.get_goodness_score(invkin_maps)
# Upate goodness array
dfname = f"{outdir}/invkinmap_{basename}.npy"
goodarray = goodarray.append({"r1": _rparam[0],
"r2": _rparam[1],
"r3": _rparam[2],
"shx": _shpos[0],
"shy": _shpos[1],
"shz": _shpos[2],
"uh": _uh,
"g1": gvals[0],
"g2": gvals[1],
"dataf": dfname},
ignore_index=True)
# Save invkinmaps data and images
with open(dfname, 'wb') as f:
np.save(f, np.array([invkin_maps['reachable'],
invkin_maps['fratio'],
invkin_maps['rangles'][:, :, 0],
invkin_maps['rangles'][:, :, 1],
invkin_maps['rangles'][:, :, 2],
invkin_maps['rangles'][:, :, 3],
invkin_maps['rangles'][:, :, 4],
invkin_maps['rangles'][:, :, 5]]))
# Update progress.
bar.update(ir * Nh + ih)
# Save goodarray
goodarray.to_csv(f"{outdir}/goodarray.csv", sep=",", index=False)
```
## Analyze goodness data
```
outdir = "opt_data"
# Read good array.
goodness = pd.read_csv(f"{outdir}/goodarray.csv")
# Read AREBO and human limb parameters
with open(f"{outdir}/robot_param_settings.json", "r") as f:
robot_param = json.load(fp=f)
with open(f"{outdir}/hlimb_param_settings.json", "r") as f:
hlimb_param = json.load(fp=f)
allgood = pd.DataFrame(columns=("r1", "r2", "r3", "O1", "O2", "O12sum", "O12prod"))
# Computer objective functions for each robot link combination.
all_rparam = np.array(aropt.iterate_over_robotparam(robot_param))
for _rparam in all_rparam:
# Find all data corresponding to the given robot link
# parameters.
_rinx = ((goodness['r1'] == _rparam[0]) &
(goodness['r2'] == _rparam[1]) &
(goodness['r3'] == _rparam[2]))
_O1 = np.mean(goodness[_rinx]['g1'])
_O2 = np.mean(goodness[_rinx]['g2'])
allgood = allgood.append({"r1": _rparam[0],
"r2": _rparam[1],
"r3": _rparam[2],
"O1": _O1,
"O2": _O2,
"O12sum": 0.5 * _O1 + 0.5 * _O2,
"O12prod": _O1 * _O2},
ignore_index=True)
# Save all good data.
allgood.to_csv(f"{outdir}/allgood.csv", sep=",", index=False)
fig = aropt.plot_objective_heatmap(allgood, robot_param, objective="O1", title="Norm. workspace");
fig.savefig(f"{outdir}/img/object1.svg", format="svg", dpi=300);
fig.savefig(f"{outdir}/img/object1.png", format="png", dpi=300);
fig = aropt.plot_objective_heatmap(allgood, robot_param, objective="O2", title="Force ratio");
fig.savefig(f"{outdir}/img/object2.svg", format="svg", dpi=300);
fig.savefig(f"{outdir}/img/object2.png", format="png", dpi=300);
fig = aropt.plot_objective_heatmap(allgood, robot_param, objective="O12sum", title="Overall Objective");
fig.savefig(f"{outdir}/img/object12sum.svg", format="svg", dpi=300);
fig.savefig(f"{outdir}/img/object12sum.png", format="png", dpi=300);
aropt.plot_objective_heatmap(allgood, robot_param, objective="O12prod", title="Overall Objective");
# Arrange the performance objective values in a descending order and
# find the link lengths at the top.
_cols = ['r1', 'r2', 'r3', 'O1', 'O2', 'O12sum']
_allgoodsum = allgood[_cols].sort_values(by="O12sum", ascending=False)
# First 10 % of the values.
_objrange = _allgoodsum['O12sum'].max() - _allgoodsum[ 'O12sum'].min()
prcnt = 10
th = _allgoodsum['O12sum'].max() - (prcnt / 100.) * _objrange
_inx = _allgoodsum['O12sum'] >= th
print(len(_allgoodsum[_inx]))
print(_allgoodsum[_inx])
print(_allgoodsum[_inx][['r1', 'r2', 'r3']].describe())
fig = figure(figsize=(12, 6.4))
# Different values of the objective function
ax = plt.subplot2grid((2, 6), loc=(0, 0), colspan=3)
ax.grid(color='0.85', linestyle='-', linewidth=0.5)
ax.set_xlim(0, len(allgood) - 1)
ax.plot(_allgoodsum['O12sum'].to_list(), lw=3)
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
ax.set_xlabel("Number of robot parameter combinations", fontsize=14)
ax.set_ylabel(r"$O\left(\mathbf{r}\right)$", fontsize=14)
ax.set_title(r"Descending values of $O\left(\mathbf{r}\right)$", fontsize=16)
# Historgram of objective function
ax = plt.subplot2grid((2, 6), loc=(0, 3), colspan=3)
ax.grid(color='0.85', linestyle='-', linewidth=0.5)
# ax.set_xlim(0, len(allgood) - 1)
ax.hist(allgood['O12sum'], lw=2, bins=50, histtype="step")
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
ax.set_xlabel(r"$O\left(\mathbf{r}\right)$", fontsize=14)
ax.set_ylabel(r"Frequency", fontsize=14)
ax.set_title(r"Historgram of $O\left(\mathbf{r}\right)$", fontsize=16)
# Objective function versus r1
ax = plt.subplot2grid((2, 6), loc=(1, 0), colspan=2)
ax.grid(color='0.85', linestyle='-', linewidth=0.5)
xvals, yvals = aropt.get_val_ranges(allgood, 'r1', 'O12sum')
ax.plot(xvals, yvals[:, 1], lw=1)
plt.fill_between(xvals, y1=yvals[:, 0], y2=yvals[:, 2], alpha=0.1)
ax.set_xlim(xvals[0],xvals[-1] )
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
ax.set_xlabel(r"$r_1$ (cm)", fontsize=16)
ax.set_ylabel(r"$O\left(\mathbf{r}\right)$", fontsize=14)
ax.set_title(r"$O\left(\mathbf{r}\right)$ vs. $r_1$", fontsize=16)
# Objective function versus r2
ax = plt.subplot2grid((2, 6), loc=(1, 2), colspan=2)
ax.grid(color='0.85', linestyle='-', linewidth=0.5)
xvals, yvals = aropt.get_val_ranges(allgood, 'r2', 'O12sum')
ax.plot(xvals, yvals[:, 1], lw=1)
plt.fill_between(xvals, y1=yvals[:, 0], y2=yvals[:, 2], alpha=0.1)
ax.set_xlim(xvals[0],xvals[-1] )
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
ax.set_xlabel(r"$r_2$ (cm)", fontsize=16)
ax.set_ylabel(r"$O\left(\mathbf{r}\right)$", fontsize=14)
ax.set_title(r"$O\left(\mathbf{r}\right)$ vs. $r_2$", fontsize=16)
# Objective function versus r3
ax = plt.subplot2grid((2, 6), loc=(1, 4), colspan=2)
ax.grid(color='0.85', linestyle='-', linewidth=0.5)
xvals, yvals = aropt.get_val_ranges(allgood, 'r3', 'O12sum')
ax.plot(xvals, yvals[:, 1], lw=1)
plt.fill_between(xvals, y1=yvals[:, 0], y2=yvals[:, 2], alpha=0.1)
ax.set_xlim(xvals[0],xvals[-1] )
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
ax.set_xlabel(r"$r_3$ (cm)", fontsize=16)
ax.set_ylabel(r"$O\left(\mathbf{r}\right)$", fontsize=14)
ax.set_title(r"$O\left(\mathbf{r}\right)$ vs. $r_3$", fontsize=16)
plt.tight_layout()
fig.savefig(f"{outdir}/img/objectsummary.svg", format="svg", dpi=300);
fig.savefig(f"{outdir}/img/objectsummary.png", format="png", dpi=300);
def group_data(df, grpcol, valcol, selinx):
grpcolvals = df[grpcol].unique()
valcolvals = [list(df[(df[grpcol] == _grpval) & selinx][valcol])
for _grpval in grpcolvals]
return grpcolvals, valcolvals
r1opt, r2opt, r3opt = 27, 20, 10
# Optimal link length selection index
_inxopt = ((goodness['r1'] == r1opt) &
(goodness['r2'] == r2opt) &
(goodness['r3'] == r3opt))
fig= figure(figsize=(14, 5.5))
# Plot the range of workspacce and force ratio values for different arm
# positions and lengths.
# Function of arm length.
uh, _g1 = group_data(goodness, grpcol='uh', valcol='g1', selinx=_inxopt)
_, _g2 = group_data(goodness, grpcol='uh', valcol='g2', selinx=_inxopt)
ax = fig.add_subplot(241)
ax.grid(color='0.85', linestyle='-', linewidth=0.5)
ax.boxplot(_g1, notch=True, sym="kx", whis=[5, 95],
positions=np.arange(1, len(uh) + 1));
ax.set_ylim(-0.1, 1.1);
plt.xticks(fontsize=16)
plt.yticks(fontsize=16)
ax.set_xticklabels(uh)
ax.set_ylabel(r"$\eta_1$", fontsize=18)
# ax.set_title(r"Normalized Workspace", fontsize=16)
ax = fig.add_subplot(245)
ax.grid(color='0.85', linestyle='-', linewidth=0.5)
ax.boxplot(_g2, notch=True, sym="kx", whis=[5, 95],
positions=np.arange(1, len(uh) + 1));
ax.set_ylim(-0.1, 1.1);
plt.xticks(fontsize=16)
plt.yticks(fontsize=16)
ax.set_xticklabels(uh)
ax.set_xlabel(r"$l$ (cm)", fontsize=18)
ax.set_ylabel(r"$\eta_2$", fontsize=18)
# ax.set_title(r"Normalized Force Ratio", fontsize=16)
# Function of shoulder x pos.
shx, _g1 = group_data(goodness, grpcol='shx', valcol='g1', selinx=_inxopt)
_, _g2 = group_data(goodness, grpcol='shx', valcol='g2', selinx=_inxopt)
ax = fig.add_subplot(242)
ax.grid(color='0.85', linestyle='-', linewidth=0.5)
ax.boxplot(_g1, notch=True, sym="kx", whis=[5, 95],
positions=np.arange(1, len(shx) + 1));
ax.set_ylim(-0.1, 1.1);
plt.xticks(fontsize=16)
plt.yticks(fontsize=16)
ax.set_xticklabels(shx)
# ax.set_title(r"Normalized Workspace", fontsize=16)
ax = fig.add_subplot(246)
ax.grid(color='0.85', linestyle='-', linewidth=0.5)
ax.boxplot(_g2, notch=True, sym="kx", whis=[5, 95],
positions=np.arange(1, len(shx) + 1));
ax.set_ylim(-0.1, 1.1);
plt.xticks(fontsize=16)
plt.yticks(fontsize=16)
ax.set_xticklabels(shx)
ax.set_xlabel(r"$p_x$ (cm)", fontsize=18)
# ax.set_title(r"Normalized Force Ratio", fontsize=16)
# Function of shoulder y pos.
shy, _g1 = group_data(goodness, grpcol='shy', valcol='g1', selinx=_inxopt)
_, _g2 = group_data(goodness, grpcol='shy', valcol='g2', selinx=_inxopt)
ax = fig.add_subplot(243)
ax.grid(color='0.85', linestyle='-', linewidth=0.5)
ax.boxplot(_g1, notch=True, sym="kx", whis=[5, 95],
positions=np.arange(1, len(shy) + 1));
ax.set_ylim(-0.1, 1.1);
plt.xticks(fontsize=16)
plt.yticks(fontsize=16)
ax.set_xticklabels(shy)
# ax.set_title(r"Normalized Workspace", fontsize=16)
ax = fig.add_subplot(247)
ax.grid(color='0.85', linestyle='-', linewidth=0.5)
ax.boxplot(_g2, notch=True, sym="kx", whis=[5, 95],
positions=np.arange(1, len(shy) + 1));
ax.set_ylim(-0.1, 1.1);
plt.xticks(fontsize=16)
plt.yticks(fontsize=16)
ax.set_xticklabels(shy)
ax.set_xlabel(r"$p_y$ (cm)", fontsize=18)
# ax.set_title(r"Normalized Force Ratio", fontsize=16)
# Function of shoulder z pos.
shz, _g1 = group_data(goodness, grpcol='shz', valcol='g1', selinx=_inxopt)
_, _g2 = group_data(goodness, grpcol='shz', valcol='g2', selinx=_inxopt)
ax = fig.add_subplot(244)
ax.grid(color='0.85', linestyle='-', linewidth=0.5)
ax.boxplot(_g1, notch=True, sym="kx", whis=[5, 95],
positions=np.arange(1, len(shz) + 1));
ax.set_ylim(-0.1, 1.1);
plt.xticks(fontsize=16)
plt.yticks(fontsize=16)
ax.set_xticklabels(shz)
# ax.set_title(r"Normalized Workspace", fontsize=16)
ax = fig.add_subplot(248)
ax.grid(color='0.85', linestyle='-', linewidth=0.5)
ax.boxplot(_g2, notch=True, sym="kx", whis=[5, 95],
positions=np.arange(1, len(shz) + 1));
ax.set_ylim(-0.1, 1.1);
plt.xticks(fontsize=16)
plt.yticks(fontsize=16)
ax.set_xticklabels(shz)
ax.set_xlabel(r"$p_z$ (cm)", fontsize=18)
# ax.set_title(r"Normalized Force Ratio", fontsize=16)
plt.tight_layout()
fig.savefig(f"{outdir}/img/optobject.svg", format="svg", dpi=300);
fig.savefig(f"{outdir}/img/optobject.png", format="png", dpi=300);
```
| github_jupyter |
# Logistic Regression
- To create a Logistic Regression, Sigmoid Function is need.
- Sigmoid Functin,
`p = 1 / ( 1 + e^-y)`
- Formula of Logistic Regression,
`ln ( p / (1 - p) ) = b0 + b1 * x `
- It is use to predict the probabilities which lies between 0 and 1.
**Importing Packages**
```
# Numpy allows us to work with array.
import numpy as np
# Maptplotlib which allows us to plot some chart.
import matplotlib.pyplot as plt
# Pandas allows us to not only import the datasets but also create the matrix of features(independent) and
# dependent variable.
import pandas as pd
```
**Importing Dataset**
- The independent variable usally in the first columns of dataset and dependent variable usally in the last columns of the data sets.
- X is Independent Variable.
- Y is Dependent Variable.
```
dataset = pd.read_csv('Social_Network_Ads.csv')
x = dataset.iloc[:, :-1].values
y = dataset.iloc[:, -1].values
print(x)
print(y)
```
**Splitting the dataset into the Training set and Test set**
```
# Importing Package
from sklearn.model_selection import train_test_split
# Dividing training and test set.
# The best ratio is 80 - 20 for trainging and testing respectively.
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size = 0.25, random_state = 0)
print(x_train)
print(y_train)
```
**Feature Scaling**
```
# Importing Package
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
# Fitting and Transforming
x_train = sc.fit_transform(x_train)
x_test = sc.transform(x_test)
print(x_train)
print(x_test)
```
**Training the Logistic Regression model on the Training set**
```
# Importing Package
from sklearn.linear_model import LogisticRegression
classifier = LogisticRegression(random_state = 0)
# Fitting
classifier.fit(x_train, y_train)
```
**Predicting a new result**
```
print(classifier.predict(sc.transform([[30, 87000]])))
```
**Predicting the Test set results**
```
# Predicting
y_pred = classifier.predict(x_test)
# Concatenating and Reshaping
print(np.concatenate((y_pred.reshape(len(y_pred),1), y_test.reshape(len(y_test),1)),1))
# First column is the predicted value and Second column is the real value.
```
**Making the confusion Matrix**
```
# Importing Package
from sklearn.metrics import confusion_matrix, accuracy_score
# Confusion Maxtrix
cm = confusion_matrix(y_test, y_pred)
print(cm)
# Accuracy Score
accuracy_score(y_test, y_pred)
```
**Visualising the Training Set results**
```
# Importing Package
from matplotlib.colors import ListedColormap
x_set, y_set = sc.inverse_transform(x_train), y_train
x1, x2 = np.meshgrid(np.arange(start = x_set[:, 0].min() - 10, stop = x_set[:, 0].max() + 10, step = 0.25),
np.arange(start = x_set[:, 1].min() - 1000, stop = x_set[:, 1].max() + 1000, step = 0.25))
plt.contourf(x1, x2, classifier.predict(sc.transform(np.array([x1.ravel(), x2.ravel()]).T)).reshape(x1.shape),
alpha = 0.75, cmap = ListedColormap(('red', 'green')))
plt.xlim(x1.min(), x1.max())
plt.ylim(x2.min(), x2.max())
for i, j in enumerate(np.unique(y_set)):
plt.scatter(x_set[y_set == j, 0], x_set[y_set == j, 1], c = ListedColormap(('red', 'green'))(i), label = j)
plt.title('Logistic Regression (Training set)')
plt.xlabel('Age')
plt.ylabel('Estimated Salary')
plt.legend()
plt.show()
```
**Visualising the Test Set results**
```
# Importing Package
from matplotlib.colors import ListedColormap
x_set, y_set = sc.inverse_transform(x_test), y_test
x1, x2 = np.meshgrid(np.arange(start = x_set[:, 0].min() - 10, stop = x_set[:, 0].max() + 10, step = 0.25),
np.arange(start = x_set[:, 1].min() - 1000, stop = x_set[:, 1].max() + 1000, step = 0.25))
plt.contourf(x1, x2, classifier.predict(sc.transform(np.array([x1.ravel(), x2.ravel()]).T)).reshape(x1.shape),
alpha = 0.75, cmap = ListedColormap(('red', 'green')))
plt.xlim(x1.min(), x1.max())
plt.ylim(x2.min(), x2.max())
for i, j in enumerate(np.unique(y_set)):
plt.scatter(x_set[y_set == j, 0], x_set[y_set == j, 1], c = ListedColormap(('red', 'green'))(i), label = j)
plt.title('Logistic Regression (Test set)')
plt.xlabel('Age')
plt.ylabel('Estimated Salary')
plt.legend()
plt.show()
```
| github_jupyter |
Yellowbrick可视化工具旨在指导模型选择过程。一般来说,模型选择是一个搜索问题,定义如下:给定N个由数值属性描述的实例和(可选)一个估计目标,找到一个由特征、算法和最适合数据的超参数组成的三元组描述的模型。在大多数情况下,“最佳”三元组是指收到模型类型的最佳交叉验证分数的三元组。
Yellowbrick.model_select包提供了可视化工具,用于检查交叉验证和超参数调优的性能。
许多可视化工具包装sklearn.model_select和其他工具中的功能,用于执行多模型比较。
当前实现的模型选择可视化器如下:
+ 验证曲线:可视化超参数的调整如何影响训练和测试分数,以调整偏差/方差。
+ 学习曲线:显示训练数据的大小如何影响模型,以诊断模型是否受方差误差和偏差误差的影响更大。
+ 交叉验证分数:将交叉验证的分数显示为条形图,平均值作为水平线。
+ 特征重要性:按模型中的相对重要性对特征进行排名
+ 递归特征消除:按重要性选择特征子集
模型选择大量使用交叉验证来评估估计器的性能。交叉验证将数据集分为训练数据集和测试数据集;该模型适合训练数据,并根据测试数据进行评估。这有助于避免常见的陷阱,过度拟合,因为模型只会记住训练数据,而不能很好地推广到新的或未知的输入中。
有很多方法可以定义如何拆分数据集以进行交叉验证。有关scikit-learn如何实现这些机制的更多信息,请查阅scikit-learn文档中的[交叉验证:评估估计器性能。](http://scikit-learn.org/stable/modules/cross_validation.html)
本文如果数据集下载不下来,查看下面地址,然后放入yellowbrick安装目录\datasets\fixtures文件夹:
```
{
"bikeshare": {
"url": "https://s3.amazonaws.com/ddl-data-lake/yellowbrick/v1.0/bikeshare.zip",
"signature": "4ed07a929ccbe0171309129e6adda1c4390190385dd6001ba9eecc795a21eef2"
},
"hobbies": {
"url": "https://s3.amazonaws.com/ddl-data-lake/yellowbrick/v1.0/hobbies.zip",
"signature": "6114e32f46baddf049a18fb05bad3efa98f4e6a0fe87066c94071541cb1e906f"
},
"concrete": {
"url": "https://s3.amazonaws.com/ddl-data-lake/yellowbrick/v1.0/concrete.zip",
"signature": "5807af2f04e14e407f61e66a4f3daf910361a99bb5052809096b47d3cccdfc0a"
},
"credit": {
"url": "https://s3.amazonaws.com/ddl-data-lake/yellowbrick/v1.0/credit.zip",
"signature": "2c6f5821c4039d70e901cc079d1404f6f49c3d6815871231c40348a69ae26573"
},
"energy": {
"url": "https://s3.amazonaws.com/ddl-data-lake/yellowbrick/v1.0/energy.zip",
"signature": "174eca3cd81e888fc416c006de77dbe5f89d643b20319902a0362e2f1972a34e"
},
"game": {
"url": "https://s3.amazonaws.com/ddl-data-lake/yellowbrick/v1.0/game.zip",
"signature": "ce799d1c55fcf1985a02def4d85672ac86c022f8f7afefbe42b20364fba47d7a"
},
"mushroom": {
"url": "https://s3.amazonaws.com/ddl-data-lake/yellowbrick/v1.0/mushroom.zip",
"signature": "f79fdbc33b012dabd06a8f3cb3007d244b6aab22d41358b9aeda74417c91f300"
},
"occupancy": {
"url": "https://s3.amazonaws.com/ddl-data-lake/yellowbrick/v1.0/occupancy.zip",
"signature": "0b390387584586a05f45c7da610fdaaf8922c5954834f323ae349137394e6253"
},
"spam": {
"url": "https://s3.amazonaws.com/ddl-data-lake/yellowbrick/v1.0/spam.zip",
"signature": "000309ac2b61090a3001de3e262a5f5319708bb42791c62d15a08a2f9f7cb30a"
},
"walking": {
"url": "https://s3.amazonaws.com/ddl-data-lake/yellowbrick/v1.0/walking.zip",
"signature": "7a36615978bc3bb74a2e9d5de216815621bd37f6a42c65d3fc28b242b4d6e040"
},
"nfl": {
"url": "https://s3.amazonaws.com/ddl-data-lake/yellowbrick/v1.0/nfl.zip",
"signature": "4989c66818ea18217ee0fe3a59932b963bd65869928c14075a5c50366cb81e1f"
}
}
```
```
import warnings
warnings.filterwarnings("ignore")
# 多行输出
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
```
# 1 验证曲线
模型验证用于确定估计器对经过训练的数据的有效性以及对新输入的通用性。为了衡量模型的性能,我们首先将数据集分为训练和测试分割,将模型拟合到训练数据上并在保留的测试数据上评分。
为了使评分最大化,必须选择模型的超参数,使模型在指定的特征空间内运行。大多数模型都有多个超参数,选择这些参数组合的最佳方法是使用网格搜索。然而,有时绘制单个超参数对训练和测试数据的影响,以确定估计器对某些超参数值是欠拟合还是过拟合是有用的。
|可视化器|ValidationCurve|
|-|-|
|快速使用方法|validation_curve()|
|模型|分类与回归|
|工作流程|选型|
## 1.1 基础使用
在第一个示例中,我们将使用ValidationCurve可视化工具探索回归数据集,在第二个示例中探索分类数据集。请注意,任何实现fit()且predict()具有适当评分机制的估算器均可与此可视化工具一起使用。
```
import numpy as np
from yellowbrick.datasets import load_energy
from yellowbrick.model_selection import ValidationCurve
from sklearn.tree import DecisionTreeRegressor
# Load a regression dataset
X, y = load_energy()
# param_name参数名称
# param_range参数范围
# cv交叉认证指定折数
# scoring评价指标
viz = ValidationCurve(
DecisionTreeRegressor(), param_name="max_depth",
param_range=np.arange(1, 11), cv=10, scoring="r2"
)
# Fit and show the visualizer
viz.fit(X, y)
viz.show();
```
在加载和修改数据之后,我们使用一个DecisionTreeRegressor初始化ValidationCurve。决策树越深越过适合,因为在树的每一层,分区处理的数据子集越小。处理这种过拟合过程的一种方法是限制树的深度。验证曲线探索了“max_depth”参数与R2评分之间的关系,并进行了10个shuffle拆分交叉验证。参数param_range指定max_depth的值,这里从1到10不等。
从可视化结果中我们可以看到,深度限制小于5个水平严重不适合这个数据集上的模型,因为训练分数和测试分数在这个参数范围内一起攀升,并且由于测试分数交叉验证的高可变性。深度为7之后,训练和测试的分数就会出现分歧,这是因为更深的树开始过度拟合训练数据,无法为模型提供普遍性。然而,由于交叉验证的分数不一定会减少,所以模型不会因为方差而产生很大的误差。
## 1.2 快速方法
使用关联的快速方法,可以在一行中实现与上述类似的功能validation_curve。此方法将实例化并适合ValidationCurve可视化器。
```
import numpy as np
from yellowbrick.datasets import load_energy
from yellowbrick.model_selection import validation_curve
from sklearn.tree import DecisionTreeRegressor
# Load a regression dataset
X, y = load_energy()
viz = validation_curve(
DecisionTreeRegressor(), X, y, param_name="max_depth",
param_range=np.arange(1, 11), cv=10, scoring="r2",
)
```
# 2 学习曲线
学习曲线显示了针对具有不同数量训练样本的估计量,训练分数与交叉验证的测试分数之间的关系。该可视化通常用于显示两件事:
估算器可从更多数据中受益多少(例如,我们是否有“足够的数据”,或者如果以在线方式使用,估算器会变得更好)。
如果估算器对因方差引起的误差与因偏差引起的误差更敏感。
|可视化器|LearningCurve|
|-|-|
|快速使用方法|learning_curve()|
|模型|分类,回归,聚类|
|工作流程|选型|
如果训练和交叉验证的分数随着更多数据的增加而聚合在一起(如左图所示),那么模型可能不会从更多的数据中获益。如果训练得分远大于验证得分,那么模型可能需要更多的训练实例,以便更有效地推广。
曲线用平均得分绘制,但是交叉验证过程中的可变性用阴影区域表示,所有交叉验证的平均值上下都有一个标准偏差。如果模型由于偏差而出现误差,那么在训练分数曲线附近可能会有更多的变化。如果模型由于方差而出现误差,那么在交叉验证得分附近会有更多的可变性。
## 2.1 分类
在下面的示例中,我们将展示如何可视化分类模型的学习曲线。
在加载DataFrame并执行分类编码之后,我们创建了一个StratifiedKFold交叉验证策略,以确保每个拆分中的所有类都以相同的比例表示。
然后,我们使用F1_加权评分度量(而不是默认度量准确度)来拟合可视化工具,以更好地了解分类器中的精确度和召回率之间的关系。
```
import numpy as np
from sklearn.model_selection import StratifiedKFold
from sklearn.naive_bayes import MultinomialNB
from sklearn.preprocessing import OneHotEncoder, LabelEncoder
from yellowbrick.datasets import load_game
from yellowbrick.model_selection import LearningCurve
# Load a classification dataset
X, y = load_game()
# Encode the categorical data
X = OneHotEncoder().fit_transform(X)
y = LabelEncoder().fit_transform(y)
# Create the learning curve visualizer
cv = StratifiedKFold(n_splits=12)
sizes = np.linspace(0.3, 1.0, 10)
# Instantiate the classification model and visualizer
model = MultinomialNB()
visualizer = LearningCurve(
model, cv=cv, scoring='f1_weighted', train_sizes=sizes, n_jobs=4
)
visualizer.fit(X, y) # Fit the data to the visualizer
visualizer.show();
```
这条学习曲线显示了高测试可变性和低分数,高达30000个实例,然而在这个水平之后,模型开始收敛到F1分数0.6左右。我们可以看到训练和测试的分数还没有收敛,所以这个模型可能会从更多的训练数据中获益。最后,该模型主要受方差误差的影响(测试数据的CV分数比训练数据的变异性更大),因此模型有可能过度拟合。
## 2.2 回归
为回归建立学习曲线非常简单,而且非常相似。在下面的示例中,在加载数据并选择目标之后,我们根据确定系数或R2分数探索学习曲线分数。
```
from sklearn.linear_model import RidgeCV
from yellowbrick.datasets import load_energy
from yellowbrick.model_selection import LearningCurve
# Load a regression dataset
X, y = load_energy()
# Instantiate the regression model and visualizer
model = RidgeCV()
visualizer = LearningCurve(model, scoring='r2')
visualizer.fit(X, y) # Fit the data to the visualizer
visualizer.show(); # Finalize and render the figure
```
该学习曲线显示出非常高的可变性,并且得分较低,直到大约350个实例为止。显然,该模型可以从更多数据中受益,因为它的得分很高。潜在地,随着更多的数据和更大的正则化alpha值,该模型在测试数据中的可变性将大大降低
## 2.3 聚类
学习曲线也适用于聚类模型,并且可以使用指定聚类的形状或组织的度量,例如轮廓分数或密度分数。如果预先知道membership,则可以使用rand得分来比较聚类性能,如下所示:
```
from sklearn.cluster import KMeans
from sklearn.datasets import make_blobs
from yellowbrick.model_selection import LearningCurve
# Generate synthetic dataset with 5 random clusters
X, y = make_blobs(n_samples=1000, centers=5, random_state=42)
# Instantiate the clustering model and visualizer
model = KMeans()
visualizer = LearningCurve(model, scoring="adjusted_rand_score", random_state=42)
visualizer.fit(X, y) # Fit the data to the visualizer
visualizer.show(); # Finalize and render the figure
```
不幸的是,对于随机数据,这些曲线变化很大,但是可以指出一些特定于聚类的项。首先,请注意y轴很窄,粗略地说,这些曲线是收敛的,并且实际上聚类算法的表现非常好。其次,对于集群而言,数据点的收敛不一定是一件坏事;实际上,我们希望确保在添加更多数据时,训练和交叉验证分数不会出现差异。
## 2.4 快速方法
使用关联的快速方法可以实现相同的功能learning_curve。此方法将LearningCurve使用关联的参数构建对象,将其拟合,然后(可选)立即显示可视化效果。
```
from sklearn.linear_model import RidgeCV
from yellowbrick.datasets import load_energy
from yellowbrick.model_selection import learning_curve
# Load a regression dataset
X, y = load_energy()
learning_curve(RidgeCV(), X, y, scoring='r2');
```
# 3 交叉验证分数
通常,我们通过查看给定模型的F1,精度,召回率和准确性(用于分类),或者确定系数(R2)和误差(用于回归)来确定其是否最优。但是,现实世界中的数据通常分布不均,这意味着拟合模型在数据的某些部分上的表现可能要好于其他部分。Yellowbrick的CVScores可视化工具使我们能够使用不同的交叉验证策略直观地探索这些性能差异。
|可视化器|CVScores|
|-|-|
|快速使用方法|cv_scores()|
|模型|分类,回归|
|工作流程|选型|
## 3.1 交叉验证介绍
交叉验证首先对数据进行无序处理(以防止意外的排序错误)并将其拆分为k个折叠。然后k模型适用于k-1/k的数据(称为训练分割),并对1/k的数据进行评估(称为测试分割)。每个评估的结果被平均在一起作为最终得分,然后最终模型适合于整个数据集进行操作。
在Yellowbrick中,CVScores可视化工具将交叉验证的分数显示为条形图(每折一个条形),所有折的平均分数绘制为水平虚线。
## 3.2 分类
在下面的示例中,我们显示了如何可视化分类模型的交叉验证得分。将占用数据加载为之后DataFrame,我们创建了StratifiedKFold交叉验证策略,以确保每个分组中的所有类均以相同的比例表示。然后,我们CVScores使用f1_weighted评分指标而不是默认指标“准确度”来拟合展示台,以便更好地了解分类器中所有折的精确度和召回率之间的关系。
```
from sklearn.model_selection import StratifiedKFold
from sklearn.naive_bayes import MultinomialNB
from yellowbrick.datasets import load_occupancy
from yellowbrick.model_selection import CVScores
# Load the classification dataset
X, y = load_occupancy()
# Create a cross-validation strategy
# 浇查验证策略
cv = StratifiedKFold(n_splits=12, random_state=42)
# Instantiate the classification model and visualizer
model = MultinomialNB()
# 交叉验证获得分数
visualizer = CVScores(model, cv=cv, scoring='f1_weighted')
visualizer.fit(X, y) # Fit the data to the visualizer
visualizer.show(); # Finalize and render the figure
```
我们得到的可视化结果表明,尽管我们的平均交叉验证得分非常高,但是在某些拆分中,我们的拟合MultinomialNB分类器的效果明显较差。
## 3.3 回归
在下一个示例中,我们显示如何可视化回归模型的交叉验证得分。在以加载enerygy数据,我们实例化了一个简单的KFold交叉验证策略。然后,我们通过CVScores使用r2计分指标对展示台进行拟合,以了解所有折处回归器的确定系数。
```
from sklearn.linear_model import Ridge
from sklearn.model_selection import KFold
from yellowbrick.datasets import load_energy
from yellowbrick.model_selection import CVScores
# Load the regression dataset
X, y = load_energy()
# Instantiate the regression model and visualizer
cv = KFold(n_splits=12, random_state=42)
model = Ridge()
visualizer = CVScores(model, cv=cv, scoring='r2')
visualizer.fit(X, y) # Fit the data to the visualizer
visualizer.show(); # Finalize and render the figure
```
与我们的分类CVScores可视化一样,我们的回归可视化表明我们的Ridge回归器几乎在所有折中都表现非常出色(例如,产生较高的确定系数),从而导致总体R2得分更高。
## 3.4 快速方法
上面的相同功能可以通过关联的快速方法来实现cv_scores。此方法将CVScores使用关联的参数构建对象,将其拟合,然后(可选)立即显示可视化效果。
```
from sklearn.linear_model import Ridge
from sklearn.model_selection import KFold
from yellowbrick.datasets import load_energy
from yellowbrick.model_selection import cv_scores
# Load the regression dataset
X, y = load_energy()
# Instantiate the regression model and visualizer
cv = KFold(n_splits=12, random_state=42)
model = Ridge()
visualizer = cv_scores(model, X, y, cv=cv, scoring='r2');
```
# 4 特征重要性
特征工程过程包括选择所需的最小特征来生成一个有效的模型,因为模型包含的特征越多,它就越复杂(数据越稀疏),因此模型对方差引起的误差就越敏感。消除特征的一种常见方法是描述它们对模型的相对重要性,然后消除薄弱的特征或特征的组合,然后重新评估,以确定在交叉验证期间模型是否更好。
许多模型形式描述了要素相对于彼此的潜在影响。在scikit-learn中,决策树模型和树的集合(例如,Random Forest,Gradient Boosting和Ada Boost)会feature_importances_在拟合时提供 属性。Yellowbrick FeatureImportances可视化工具利用此属性对相对重要性进行排名和绘图。
|可视化器|FeatureImportances|
|-|-|
|快速使用方法|feature_importances()|
|模型|分类,回归|
|工作流程|选型,特征提取|
## 4.1 基本使用
让我们从一个例子开始;首先加载分类数据集。
然后,我们可以创建一个新图形(这是可选的,如果Axes未指定,Yellowbrick将使用当前图形或创建一个图形)。然后,我们可以为FeatureImportances展示台添加,GradientBoostingClassifier以可视化排名特征。
```
from sklearn.ensemble import RandomForestClassifier
from yellowbrick.datasets import load_occupancy
from yellowbrick.model_selection import FeatureImportances
# Load the classification data set
X, y = load_occupancy()
model = RandomForestClassifier(n_estimators=10)
# 显示特征对于分类的重要程度
viz = FeatureImportances(model)
viz.fit(X, y)
viz.show();
```
相对于其相对重要性(即最重要特征的重要性百分比)绘制特征。可视化器还包含features_和 feature_importances_属性以获取排名的数值。
对于不支持feature_importances_属性的 模型,FeatureImportances可视化工具还将为coef_ 许多线性模型提供的属性绘制条形图。但是要注意的是系数重要性的解释取决于模型。
使用带有coef_属性的模型时,最好设置 relative=False为绘制系数的真实大小(可能为负)。如果数据集没有列名或打印更好的标题,我们也可以指定我们自己的标签集。在下面的示例中,我们为案例加了标题以提高可读性:
```
from sklearn.linear_model import Lasso
from yellowbrick.datasets import load_concrete
from yellowbrick.model_selection import FeatureImportances
# Load the regression dataset
dataset = load_concrete(return_dataset=True)
X, y = dataset.to_data()
# Title case the feature for better display and create the visualizer
labels = list(map(lambda s: s.title(), dataset.meta['features']))
viz = FeatureImportances(Lasso(), labels=labels, relative=False)
# Fit and show the feature importances
viz.fit(X, y)
viz.show();
```
## 4.2 堆叠重要性特征
有些估计器返回一个多维数组,用于feature_importances_x或coef_u属性。例如,在多类情况下,LogisticRegression分类器返回一个coef_u数组,其形状为(n_classes,n_features)。这些系数将特征的重要性映射到特定类别的概率预测中。虽然多维特征重要性的解释依赖于特定的估计器和模型族,但是在FeatureImportances visualizer中对数据的处理是相同的,即重要性被平均化。
出于几个原因,采用重要性的平均值可能是不可取的。例如,某个特性对于某些类来说可能比其他类更具信息量。在本质上是多个内部模型的情况下,多输出估计量也不会从中受益。在这种情况下,使用stack=True参数绘制重要程度的堆积条形图,如下所示:
```
from yellowbrick.model_selection import FeatureImportances
from sklearn.linear_model import LogisticRegression
from sklearn.datasets import load_iris
data = load_iris()
X, y = data.data, data.target
model = LogisticRegression(multi_class="auto", solver="liblinear")
viz = FeatureImportances(model, stack=True, relative=False)
viz.fit(X, y)
viz.show();
```
## 4.3 快速方法
可以通过关联的快速方法feature_importances实现上述相同功能。此方法将FeatureImportances使用关联的参数构建对象,将其拟合,然后
from sklearn.ensemble import AdaBoostClassifier(可选)立即显示它。
```
from sklearn.ensemble import AdaBoostClassifier
from yellowbrick.datasets import load_occupancy
from yellowbrick.model_selection import feature_importances
# Load dataset
X, y = load_occupancy()
# Use the quick method and immediately show the figure
feature_importances(AdaBoostClassifier(), X, y);
```
# 5 递归特征消除
递归特征消除(RFE)是一种特征选择方法,它适合模型并删除最弱的一个或多个特征,直到达到指定数量的特征为止。特征按模型coef_或feature_importances_属性进行排序,并且通过递归消除每个循环中的少量特征,RFE尝试消除模型中可能存在的依赖关系和共线性。
RFE需要保留指定数量的功能,但是通常事先不知道有多少有效功能。为了找到最佳数量的特征,将交叉验证与RFE一起使用,对不同的特征子集进行评分,并选择最佳的特征评分集合。该RFECV可视化工具绘制的模型特征的数量与他们的交叉验证测试分数和多变性和可视化选择的多项功能一起。
为了展示其在实际中的工作方式,我们将从一个人为设计的示例开始,该示例使用的数据集中只有25个中的3个信息功能。
|可视化器|RFECV|
|-|-|
|快速使用方法|rfecv()|
|模型|分类,回归|
|工作流程|选型|
## 5.1 基础使用
```
from sklearn.svm import SVC
from sklearn.datasets import make_classification
from yellowbrick.model_selection import RFECV
# Create a dataset with only 3 informative features
# 只有三个有效特征
X, y = make_classification(
n_samples=1000, n_features=25, n_informative=3, n_redundant=2,
n_repeated=0, n_classes=8, n_clusters_per_class=1, random_state=0
)
# Instantiate RFECV visualizer with a linear SVM classifier
visualizer = RFECV(SVC(kernel='linear', C=1))
visualizer.fit(X, y) # Fit the data to the visualizer
visualizer.show(); # Finalize and render the figure
```
该图显示了一条理想的RFECV曲线,当捕获了三个信息性特征时,该曲线跳至极佳的精度,然后随着将非信息性特征添加到模型中而逐渐降低了精度。阴影区域表示交叉验证的变异性,即曲线绘制的平均准确度得分之上和之下的一个标准偏差。
探索真实数据集,我们可以看到RFECV对credit默认二进制分类器的影响。
```
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import StratifiedKFold
from yellowbrick.model_selection import RFECV
from yellowbrick.datasets import load_credit
# Load classification dataset
X, y = load_credit()
cv = StratifiedKFold(2)
visualizer = RFECV(RandomForestClassifier(), cv=cv, scoring='f1_weighted')
visualizer.fit(X, y) # Fit the data to the visualizer
visualizer.show() # Finalize and render the figure
```
在此示例中,我们可以看到选择了15个特征,尽管在大约5个特征之后,模型的f1得分似乎没有太大改善。选择要消除的特征在确定每次递归的结果中起着很大的作用;修改step参数以在每个步骤中消除一个以上的特征可能有助于尽早消除最差的特征,从而增强其余特征(并且还可以用于加快具有大量特征的数据集的特征消除)。
## 5.2 快速方法
上面的相同功能可以通过关联的快速方法来实现rfecv。此方法将RFECV使用关联的参数构建对象,将其拟合,然后(可选)立即显示可视化效果。
```
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import StratifiedKFold
from yellowbrick.model_selection import rfecv
from yellowbrick.datasets import load_credit
# Load classification dataset
X, y = load_credit()
cv = StratifiedKFold(2)
visualizer = rfecv(RandomForestClassifier(), X=X, y=y, cv=cv, scoring='f1_weighted');
```
# 6 参考
[https://www.scikit-yb.org/en/latest/api/model_selection/validation_curve.html](https://www.scikit-yb.org/en/latest/api/model_selection/validation_curve.html)
[https://www.scikit-yb.org/en/latest/api/model_selection/learning_curve.html](https://www.scikit-yb.org/en/latest/api/model_selection/learning_curve.html)
[https://www.scikit-yb.org/en/latest/api/model_selection/cross_validation.html](https://www.scikit-yb.org/en/latest/api/model_selection/cross_validation.html)
[https://www.scikit-yb.org/en/latest/api/model_selection/importances.html](https://www.scikit-yb.org/en/latest/api/model_selection/importances.html)
[https://www.scikit-yb.org/en/latest/api/model_selection/rfecv.html](https://www.scikit-yb.org/en/latest/api/model_selection/rfecv.html)
| github_jupyter |
Copyright (c) Microsoft Corporation. All rights reserved.
Licensed under the MIT License.
# Tutorial: Train a classification model with automated machine learning
In this tutorial, you'll learn how to generate a machine learning model using automated machine learning (automated ML). Azure Machine Learning can perform algorithm selection and hyperparameter selection in an automated way for you. The final model can then be deployed following the workflow in the [Deploy a model](02.deploy-models.ipynb) tutorial.
[flow diagram](./imgs/flow2.png)
Similar to the [train models tutorial](01.train-models.ipynb), this tutorial classifies handwritten images of digits (0-9) from the [MNIST](http://yann.lecun.com/exdb/mnist/) dataset. But this time you don't to specify an algorithm or tune hyperparameters. The automated ML technique iterates over many combinations of algorithms and hyperparameters until it finds the best model based on your criterion.
You'll learn how to:
> * Set up your development environment
> * Access and examine the data
> * Train using an automated classifier locally with custom parameters
> * Explore the results
> * Review training results
> * Register the best model
## Prerequisites
Use [these instructions](https://aka.ms/aml-how-to-configure-environment) to:
* Create a workspace and its configuration file (**config.json**)
* Upload your **config.json** to the same folder as this notebook
### Start a notebook
To follow along, start a new notebook from the same directory as **config.json** and copy the code from the sections below.
## Set up your development environment
All the setup for your development work can be accomplished in the Python notebook. Setup includes:
* Import Python packages
* Configure a workspace to enable communication between your local computer and remote resources
* Create a directory to store training scripts
### Import packages
Import Python packages you need in this tutorial.
```
import azureml.core
import pandas as pd
from azureml.core.workspace import Workspace
from azureml.train.automl.run import AutoMLRun
import time
import logging
from sklearn import datasets
from matplotlib import pyplot as plt
from matplotlib.pyplot import imshow
import random
import numpy as np
```
### Configure workspace
Create a workspace object from the existing workspace. `Workspace.from_config()` reads the file **aml_config/config.json** and loads the details into an object named `ws`. `ws` is used throughout the rest of the code in this tutorial.
Once you have a workspace object, specify a name for the experiment and create and register a local directory with the workspace. The history of all runs is recorded under the specified experiment.
```
ws = Workspace.from_config()
# choose a name for the run history container in the workspace
experiment_name = 'automl-classifier'
# project folder
project_folder = './automl-classifier'
import os
output = {}
output['SDK version'] = azureml.core.VERSION
output['Subscription ID'] = ws.subscription_id
output['Workspace'] = ws.name
output['Resource Group'] = ws.resource_group
output['Location'] = ws.location
output['Project Directory'] = project_folder
pd.set_option('display.max_colwidth', -1)
pd.DataFrame(data=output, index=['']).T
```
## Explore data
The initial training tutorial used a high-resolution version of the MNIST dataset (28x28 pixels). Since auto training requires many iterations, this tutorial uses a smaller resolution version of the images (8x8 pixels) to demonstrate the concepts while speeding up the time needed for each iteration.
```
from sklearn import datasets
digits = datasets.load_digits()
# Exclude the first 100 rows from training so that they can be used for test.
X_train = digits.data[100:,:]
y_train = digits.target[100:]
```
### Display some sample images
Load the data into `numpy` arrays. Then use `matplotlib` to plot 30 random images from the dataset with their labels above them.
```
count = 0
sample_size = 30
plt.figure(figsize = (16, 6))
for i in np.random.permutation(X_train.shape[0])[:sample_size]:
count = count + 1
plt.subplot(1, sample_size, count)
plt.axhline('')
plt.axvline('')
plt.text(x = 2, y = -2, s = y_train[i], fontsize = 18)
plt.imshow(X_train[i].reshape(8, 8), cmap = plt.cm.Greys)
plt.show()
```
You now have the necessary packages and data ready for auto training for your model.
## Auto train a model
To auto train a model, first define settings for autogeneration and tuning and then run the automatic classifier.
### Define settings for autogeneration and tuning
Define the experiment parameters and models settings for autogeneration and tuning.
|Property| Value in this tutorial |Description|
|----|----|---|
|**primary_metric**|AUC Weighted | Metric that you want to optimize.|
|**max_time_sec**|12,000|Time limit in seconds for each iteration|
|**iterations**|20|Number of iterations. In each iteration, the model trains with the data with a specific pipeline|
|**n_cross_validations**|3|Number of cross validation splits|
|**exit_score**|0.9985|*double* value indicating the target for *primary_metric*. Once the target is surpassed the run terminates|
|**blacklist_algos**|['kNN','LinearSVM']|*Array* of *strings* indicating algorithms to ignore.
```
from azureml.train.automl import AutoMLConfig
##Local compute
Automl_config = AutoMLConfig(task = 'classification',
primary_metric = 'AUC_weighted',
max_time_sec = 12000,
iterations = 20,
n_cross_validations = 3,
exit_score = 0.9985,
blacklist_algos = ['kNN','LinearSVM'],
X = X_train,
y = y_train,
path=project_folder)
```
### Run the automatic classifier
Start the experiment to run locally. Define the compute target as local and set the output to true to view progress on the experiment.
```
from azureml.core.experiment import Experiment
experiment=Experiment(ws, experiment_name)
local_run = experiment.submit(Automl_config, show_output=True)
```
## Explore the results
Explore the results of automatic training with a Jupyter widget or by examining the experiment history.
### Jupyter widget
Use the Jupyter notebook widget to see a graph and a table of all results.
```
from azureml.widgets import RunDetails
RunDetails(local_run).show()
```
### Retrieve all iterations
View the experiment history and see individual metrics for each iteration run.
```
children = list(local_run.get_children())
metricslist = {}
for run in children:
properties = run.get_properties()
metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)}
metricslist[int(properties['iteration'])] = metrics
import pandas as pd
rundata = pd.DataFrame(metricslist).sort_index(1)
rundata
```
## Register the best model
Use the `local_run` object to get the best model and register it into the workspace.
```
# find the run with the highest accuracy value.
best_run, fitted_model = local_run.get_output()
# register model in workspace
description = 'Automated Machine Learning Model'
tags = None
local_run.register_model(description=description, tags=tags)
local_run.model_id # Use this id to deploy the model as a web service in Azure
```
## Test the best model
Use the model to predict a few random digits. Display the predicted and the image. Red font and inverse image (white on black) is used to highlight the misclassified samples.
Since the model accuracy is high, you might have to run the following code a few times before you can see a misclassified sample.
```
# find 30 random samples from test set
n = 30
X_test = digits.data[:100, :]
y_test = digits.target[:100]
sample_indices = np.random.permutation(X_test.shape[0])[0:n]
test_samples = X_test[sample_indices]
# predict using the model
result = fitted_model.predict(test_samples)
# compare actual value vs. the predicted values:
i = 0
plt.figure(figsize = (20, 1))
for s in sample_indices:
plt.subplot(1, n, i + 1)
plt.axhline('')
plt.axvline('')
# use different color for misclassified sample
font_color = 'red' if y_test[s] != result[i] else 'black'
clr_map = plt.cm.gray if y_test[s] != result[i] else plt.cm.Greys
plt.text(x = 2, y = -2, s = result[i], fontsize = 18, color = font_color)
plt.imshow(X_test[s].reshape(8, 8), cmap = clr_map)
i = i + 1
plt.show()
```
## Next steps
In this Azure Machine Learning tutorial, you used Python to:
> * Set up your development environment
> * Access and examine the data
> * Train using an automated classifier locally with custom parameters
> * Explore the results
> * Review training results
> * Register the best model
Learn more about [how to configure settings for automatic training](https://aka.ms/aml-how-to-configure-auto) or [how to use automatic training on a remote resource](https://aka.ms/aml-how-to-auto-remote).
| github_jupyter |
<a href="https://colab.research.google.com/github/alirezash97/Cardio/blob/master/Transfer_Learning.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
!pip install kaggle
!mkdir .kaggle
import json
token = {"username":"alirezashafaei97","key":"9cb262aa0c5658ffc4eb45857c41903c"}
with open('/content/.kaggle/kaggle.json', 'w') as file:
json.dump(token, file)
!mkdir ~/.kaggle
!cp /content/.kaggle/kaggle.json ~/.kaggle/kaggle.json
!kaggle config set -n path -v{/content}
!chmod 600 /root/.kaggle/kaggle.json
!kaggle datasets download -d shayanfazeli/heartbeat -p /content
!unzip /content/heartbeat.zip -d /content/heartbeat
import pandas as pd
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
from tensorflow.keras.models import load_model, Sequential, Model
from tensorflow.keras.layers import (Input, Dense, LeakyReLU, Softmax, InputLayer, concatenate, Conv1D, MaxPool1D
, Flatten, Dropout, ReLU, BatchNormalization, GlobalAveragePooling1D)
from tensorflow.keras import backend as K
from tensorflow.keras import regularizers
from random import uniform
import random
from sklearn.preprocessing import OneHotEncoder
from scipy.sparse import csr_matrix
train_df=pd.read_csv('/content/heartbeat/mitbih_train.csv',header=None)
test_df=pd.read_csv('/content/heartbeat/mitbih_test.csv',header=None)
train_df[187]=train_df[187].astype(int)
counter=train_df[187].value_counts()
print(counter)
trainset = train_df.values
testset = test_df.values
def split(dataset, number_of_sample_per_category):
new_dataset = np.zeros((np.sum(number_of_sample_per_category), dataset.shape[1]))
pointer = 0
for row in dataset :
row_label = int(row[-1])
if number_of_sample_per_category[row_label] > 0 :
number_of_sample_per_category[row_label] -= 1
new_dataset[pointer , :] = row
pointer += 1
else:
pass
return new_dataset
temp_trainset = split(trainset, [5500, 2223, 5500, 641, 5500])
temp_testset = split(testset, [500, 500, 500, 500, 500])
def data_augmentation(dataset, chance):
augmented = 0
number_of_rows = int(dataset.shape[0] + (dataset.shape[0] * (chance*2)))
new_dataset = np.zeros((number_of_rows, dataset.shape[1]))
pointer = 0
for row in dataset:
rand_num = random.uniform(0, 1)
if rand_num < chance:
augmented += 1
noise = np.random.normal(scale=0.01, size=187)
new_signal = np.zeros((1, 188))
new_signal[:, :187] = row[:187] + noise
new_signal[:, -1:] = row[-1:]
new_dataset[pointer:pointer+1, :] = new_signal
pointer += 1
else :
pass
new_dataset[pointer, :] = row
pointer += 1
return augmented, new_dataset
augmented, trainset = data_augmentation(temp_trainset, 0.08)
filled = augmented + temp_trainset.shape[0]
trainset = trainset[:filled , :]
trainset = np.take(trainset,np.random.permutation(trainset.shape[0]),axis=0,out=trainset)
testset = np.take(temp_testset,np.random.permutation(temp_testset.shape[0]),axis=0,out=temp_testset)
X_temp_train = trainset[:, :-1]
Y_train = trainset[:, -1:]
X_temp_test = testset[:, :-1]
Y_test = testset[:, -1:]
print("X_train : ", X_temp_train.shape)
print("Y_train : ", Y_train.shape)
print("X_test : ", X_temp_test.shape)
print("Y_test : ", Y_test.shape)
plt.plot(X_temp_train[7, :])
plt.show()
# One Hot enoding for target labels
ohe = OneHotEncoder()
Y_train = ohe.fit_transform(Y_train.reshape(-1,1))
Y_test = ohe.transform(Y_test.reshape(-1,1))
# handle sparse matrix for keras
Y_train = csr_matrix.toarray(Y_train)
Y_test = csr_matrix.toarray(Y_test)
print("X_train : ", X_temp_train.shape)
print("Y_train : ", Y_train.shape)
print("X_test : ", X_temp_test.shape)
print("Y_test : ", Y_test.shape)
Y_train[1000:1006, :]
# def number_generator(a, b, n):
# n = n+1
# Output = np.zeros((n-1))
# if a > b :
# s = (a - b) / n
# temp = b
# for i in range(n-1):
# temp += s
# Output[i] = temp
# elif a < b :
# s = (b - a) / n
# temp = a
# for i in range(n-1):
# temp += s
# Output[i] = temp
# else:
# for i in range(n-1):
# Output[i] = a
# return Output
# #### change X_train.shape from (1, 187) to (1, 748) for transfer learning ####
# X_train = np.zeros((X_temp_train.shape[0], 374)) #
# j = 1 #
# for i, signal in enumerate(X_temp_train):
# for index, item in enumerate(signal):
# if index != 186:
# temp = number_generator(signal[index], signal[index+1], 1) #
# if (signal[index+1]) > (signal[index]) :
# temp = np.sort(temp)
# else:
# temp = -np.sort(-temp)
# X_train[i, index*j:(index+1)*(j)] = temp
# else:
# pass
# plt.plot(X_temp_train[1, :])
# plt.show()
# plt.plot(X_train[1, :])
# plt.show()
#### change X_test.shape from (1, 187) to (1, 748) for transfer learning ####
# X_test = np.zeros((X_temp_test.shape[0], 374))
# j = 1
# for i, signal in enumerate(X_temp_test):
# for index, item in enumerate(signal):
# if index != 186:
# temp = number_generator(signal[index], signal[index+1], 1)
# if (signal[index+1]) > (signal[index]) :
# temp = np.sort(temp)
# else:
# temp = -np.sort(-temp)
# X_test[i, index*j:(index+1)*(j)] = temp
# else:
# pass
# plt.plot(X_temp_test[1, :])
# plt.show()
# plt.plot(X_test[1, :])
# plt.show()
# periodic signal extend
import pywt
XF_train = np.zeros((X_temp_train.shape[0], 9000))
XF_test = np.zeros((X_temp_test.shape[0], 9000))
for index, row in enumerate(X_temp_train):
XF_train[index, :-1] = pywt.pad(row, 4406, 'periodic')
XF_train[index, -1:] = 0
for index, row in enumerate(X_temp_test):
XF_test[index, :-1] = pywt.pad(row, 4406, 'periodic')
XF_test[index, -1:] = 0
XF_train = XF_train.reshape((20947, 9000, 1))
XF_test = XF_test.reshape((2500, 9000, 1))
print("X_train : ", XF_train.shape)
print("Y_train : ", Y_train.shape)
print("X_test : ", XF_test.shape)
print("Y_test : ", Y_test.shape)
# load model
model = load_model('/content/drive/My Drive/Cardio/AF_Classification.h5')
# summarize model.
model.summary()
X_input = model.layers[5].output
######
conv0 = Conv1D(filters=128, kernel_size= 9, strides = 2, padding='same')(X_input)
# Bn0 = BatchNormalization()(conv0)
Act0 = ReLU()(conv0)
conv1 = Conv1D(filters=256, kernel_size= 7, strides = 2, padding='same')(Act0)
# Bn1 = BatchNormalization()(conv1)
Act1 = ReLU()(conv1)
pool0=MaxPool1D(pool_size=(2), strides=(2), padding="same")(Act1)
flatten=Flatten()(pool0)
D0 = Dense(512)(flatten)
Bn2 = BatchNormalization()(D0)
Act2 = ReLU()(Bn2)
Drop0 = Dropout(0.5)(Act2)
######
D1 = Dense(128)(Drop0)
# AP1 = GlobalAveragePooling2D()(D1)
Bn3 = BatchNormalization()(D1)
Act3 = ReLU()(Bn3)
Drop1 = Dropout(0.3)(Act3)
#####
D2 = Dense(5)(Drop1)
Act4 = Softmax()(D2)
new_model = Model(inputs=model.inputs, outputs=Act4)
new_model.summary()
# compile model
new_model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
# Early Stopping
es_callback = tf.keras.callbacks.EarlyStopping(monitor='val_loss', patience=3)
# Fit the model
new_model.fit(XF_train, Y_train, epochs=10, batch_size=128, validation_data=(XF_test, Y_test), callbacks=[es_callback])
```
| github_jupyter |
<a href="https://colab.research.google.com/github/krakowiakpawel9/data-science-bootcamp/blob/master/05_p_stwo_statystyka/01_p_stwo_statystyka.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
* @author: krakowiakpawel9@gmail.com
* @site: e-smartdata.org
### Rachunek Prawdopodobieństwa i Statystyka
### Spis treści:
1. [Lekkie wprowadzenie](#1)
2. [Statystyka opisowa - miary tendencji centralnej](#2)
3. [Statystyka opisowa - miary rozrzutu](#3)
4. [Dystrybuanta empiryczna](#4)
5. [Przestrzeń probabilistyczna](#5)
6. [Przestrzeń klasyczna](#6)
7. [Niezależność zdarzeń](#7)
8. [Prawdopodobieństwo warunkowe](#8)
9. [Zmienna losowa](#9)
### <a name='1'></a> Lekkie wprowadzenie
Rozważmy populację składającą się dokładnie z $n$ elementów. Oznaczmy przez $X$ badaną cechę populacji oraz przez $x_1, \dots, x_n$ wartości cechy $X$. Cechę $X$ nazwyać będziemy zmienną losową.
---
**Przykład - dane dyskretne**
Populacja - grupa studentów (5 osób)
Badana cecha $X$- ocena z egzaminu
Realizacja zmiennej: 3, 4.5, 5, 4.5, 3
```
import numpy as np
X = np.array([3, 4.5, 5, 4.5, 3])
print(X)
```
**Przykład - dane ciągłe**
Populacja - grupa studentów (5 osób)
Badana cecha $X$- wzrost studenta
Realizacja zmiennej: 185.0, 179.5, 168.0, 192.0, 185.5
```
X = np.array([185.0, 179.5, 168.0, 192.0, 185.5])
print(X)
```
### <a name='2'></a> Statystyka opisowa - miary tendencji centralnej
**DEFINICJA 1.**
Średnią arytmetyczną ciągu wartości $x_1, \dots, x_n$ nazwyamy:
$$\bar{x}= \frac{x_{1} + x_{2} + \ \ldots \ + x_{n}}{n} = \frac{1}{n} \sum_{i=1}^{n} x_{i}$$
**PRZYKŁAD 2.**
Rozważmy ciąg: $3, 4.5, 5, 4.5, 3$
$$\bar{x}=\frac{3 + 4.5 + 5 + 4.5 + 3}{5} = 4.0$$
```
X = np.array([3, 4.5, 5, 4.5, 3])
X.mean()
```
**DEFINICJA 3.**
Medianą ciągu wartości $x_1, \dots, x_n$ nazywamy środkowy wyraz tego ciągu, gdy $n$ jest liczbą nieparzystą, w przeciwnym przypadku średnią arytmetyczną dwóch wyrazów środkowych.
$Mediana = Me = \left\{\begin{array} {lll} x_{(k+1)} & \hbox{ dla } & n=2k+1 \\[1mm] \frac{x_{(k)}+x_{(k+1)}}{2} & \hbox{ dla } & n=2k. \end{array} \right.$,
gdzie $x_{(1)} \le x_{(2)} \le \dots \le x_{(n)}.$
**PRZYKŁAD 4.**
Rozważmy ciąg: $3, 4.5, 5, 4.5, 3$
Porządkujemy ciąg: $3, 3, 4.5, 4.5, 5$
Ponieważ liczba elementów (=5) jest liczbą nieparzystą wybieramy środkowy wyraz tego ciągu czyli:
$$Mediana = 4.5$$
```
np.median(X)
```
**PRZYKŁAD 5.**
Rozważmy ciąg: $5, 4.5, 3, 5$
Porządkujemy ciąg: $3, 4.5, 5, 5$
Ponieważ liczba elementów (=4) jest liczbą parzystą obliczamy średnią arytmetyczną wyrazów środkowych:
$$Mediana = \frac{4.5 + 5}{2} = 4.75$$
**DEFINICJA 6.**
Moda (wartość modalna) to najczęściej występująca wartość w ciągu.
**PRZYKŁAD 7.**
Rozważmy ciąg: $3, 4.5, 5, 4.5, 2$
$$Moda = 4.5$$
### <a name='3'></a> Statystyka opisowa - miary rozrzutu
**DEFINICJA 8.**
**Średni błąd** ciągu wartości $x_1, \dots, x_n$ nazywamy wartość:
$$b=\frac{1}{n} \sum_{i=1}^{n} | x_{i}-\bar{x} |$$
Im mniejszy jest średni błąd tym zmienna $X$ ma mniejszy rozrzut.
**PRZYKŁAD 9.**
Rozważmy zmienną losową $X$ o realizacji $3.5, 4.0, 4.0$ oraz zmienną losową $Y$ o realizacji $2.0, 4.0, 5.0$.
Policzmy średni błąd.
$\bar{X} = \frac{3.5 + 4.0 + 4.0}{3} = 3.83$
$\bar{Y} = \frac{2.0 + 4.0 + 5.0}{3} = 3.67$
Zatem
$b_X = \frac{1}{3}(|3.5 - 3.83| + |4.0 - 3.83| + |4.0 - 3.83|) = $
```
X = [3.5, 4.0, 4.0]
Y = [2.0, 4.0, 5.0]
X_mean = np.mean(X)
Y_mean = np.mean(Y)
print(f'X_mean: {X_mean:.4f}')
print(f'Y_mean: {Y_mean:.4f}')
b_X = 1 / len(X) * (abs(X - X_mean).sum())
b_Y = 1 / len(Y) * (abs(Y - Y_mean).sum())
print(f'Średni błąd X: {b_X:.4f}')
print(f'Średni błąd Y: {b_Y:.4f}')
```
**DEFINICJA 10.**
Wariancją ciągu $x_1, \dots, x_n$ nazywamy wartość:
$$s_{n}^{2} = \frac{1}{n} \sum_{i=1}^{n} ( x_{i}-\bar{x})^{2}$$
**DEFINICJA 11.**
Odchyleniem standardowym ciągu $x_1, \dots, x_n$ nazywamy wartość:
$$s_n = \sqrt{\frac{1}{n} \sum_{i=1}^{n} ( x_{i}-\bar{x})^{2}}.$$
```
var_X = 1 / len(X) * ((X - X_mean)**2).sum()
var_Y = 1 / len(Y) * ((Y - Y_mean)**2).sum()
print(f'Wariancja zmiennej X: {var_X:.4f}')
print(f'Wariancja zmiennej Y: {var_Y:.4f}\n')
std_X = np.sqrt(var_X)
std_Y = np.sqrt(var_Y)
print(f'Odchylenie standardowe zmiennej X: {std_X:.4f}')
print(f'Odchylenie standardowe zmiennej Y: {std_Y:.4f}')
print(np.std(X))
print(np.std(Y))
```
### <a name='4'></a> Dystrybuanta empiryczna
**DEFINICJA 12.**
Kwantylem rzędu $p$ zmiennej losowej $X$ nazywamy wartość, która dzieli uporządkowany ciąg wartości $x_1, \dots, x_n$ na dwie części w proporcjach $p$ oraz $1-p$.
**PRZYKŁAD 13.**
Mediana jest kwantylem rzędu $p=\frac{1}{2}$.
**DEFINICJA 14.**
Dystrybuantą empiryczną ciągu $x_1, \dots, x_n$ nazywamy funkcję określoną wzorem:
$$F(x)=\frac{ \# \{x_{i}: \ x_{i} \leq x \} }{n} \;\; \textrm{dla}\;\; x\in {\Bbb R}$$
Przy pomocy dystrybuanty możemy w prosty sposób określić wzór na kwantyl rzędu $p$:
$$q_p = \min \{ x : F(x) \ge p \}$$
### <a name='5'></a> Przestrzeń probabilistyczna
**DEFINICJA 15.**
Przestrzenią probabilistyczną nazywamy trójkę $(\Omega, \mathscr{F}, P)$, gdzie $\Omega$ jest zbiorem niepustym (przestrzeń zdarzeń), $\mathscr{F}$ jest rodziną podzbiorów zbioru $\Omega$ (sigma ciało) oraz $P$ jest miarą probabilistyczną (prawdopodobieństwo):
$$P:\mathscr{F} \rightarrow R$$
Elementy zbioru $\Omega$ nazywamy zdarzeniami elementarnymi, elementy $\mathscr{F}$ zdarzeniami. Zbiór pusty reprezentuje zdarzenie niemożliwe, zaś zbiór $\Omega$ zdarzenie pewne. Zdarzenie $\Omega \backslash A$ nazywamy zdarzeniem przeciwnym do zdarzenia $A$ i wreszcie liczbę $P(A)$ nazywamy prawdopodobieństwem zdarzenia A.
### <a name='6'></a> Przestrzeń klasyczna
Niech $\Omega$ będzie zbiorem skończonym składającym się z $n$ jednakowo prawdopodobnych zdarzeń elementarnych:
$$\Omega = \{\omega_1, \dots, \omega_n\}$$
oraz niech $\mathscr{F}$ rodzina podzbiorów zbioru $\Omega$. Jeśli $A \in \mathscr{F}$, to:
$$P(A) = \frac{\#A}{n}$$
**PRZYKŁAD 16.**
Niech $\Omega = \{1, 2, 3, 4, 5, 6\}$, $A$ oznacza zdarzenie polegające na wyrzuceniu liczby parzystej, mamy zatem
$n=6$
$A=\{2, 4, 6\}$
oraz $\#A = 3$
Stąd $P(A) = \frac{\#A}{n} = \frac{3}{6} = \frac{1}{2}$
```
omega = {1, 2, 3, 4, 5, 6}
A = {item for item in omega if item % 2 == 0}
P_A = len(A) / len(omega)
print(f'Zbiór A: {A}')
print(f'Prawdopodobieństwo zdarzenia A: {P_A}')
```
**PRZYKŁAD 17.**
Niech $\Omega = \{1, 2, 3, 4, 5, 6\}$, $B$ oznacza zdarzenie polegające na wyrzuceniu liczby większej niż 4, mamy zatem
$n=6$
$B=\{5, 6\}$
oraz $\#B = 2$
Stąd $P(B) = \frac{\#A}{n} = \frac{2}{6} = \frac{1}{3}$
```
omega = {1, 2, 3, 4, 5, 6}
B = {item for item in omega if item > 4}
P_B = len(B) / len(omega)
print(f'Zbiór B: {B}')
print(f'Prawdopodobieństwo zdarzenia B: {P_B:.4f}')
```
### <a name='7'></a> Niezależność zdarzeń
**DEFINICJA 18.**
Dwa zdarzenia $A$ i $B$ są niezależne, gdy:
$$P(A\cap B) = P(A)\cdot P(B)$$
**PRZYKŁAD 17.**
Rozważmy dwa zdarzenia $A$ i $B$ z poprzednich przykładów.
```
intersection_AB = set.intersection(A, B)
P_intersection_AB = len(intersection_AB) / len(omega)
PA_PB = P_A * P_B
print(f'Przecięcie zbiorów A i B: {intersection_AB}\n')
print(f'Iloczyn prawdopodobieństw zdarzeń A i B: {PA_PB:.4f}')
print(f'Prawdopodobieństwo iloczynu zdarzeń A i B: {P_intersection_AB:.4f}\n')
check = 'Zdarzenia niezależne' if PA_PB == P_intersection_AB else 'Zdarzenia zależne'
print(check)
```
Rozważmy zdarzenie $C$ - wyrzucenie liczby oczek większą niż 1
```
C = {item for item in omega if item > 1}
P_C = len(C) / len(omega)
print(f'Zbiór C: {C}')
print(f'Prawdopodobieństwo zdarzenia C: {P_C:.4f}')
intersection_AC = set.intersection(A, C)
P_intersection_AC = len(intersection_AC) / len(omega)
PA_PC = P_A * P_C
print(f'Przecięcie zbiorów A i C: {intersection_AC}\n')
print(f'Iloczyn prawdopodobieństw zdarzeń A i C: {PA_PC:.4f}')
print(f'Prawdopodobieństwo iloczynu zdarzeń A i C: {P_intersection_AC:.4f}\n')
check = 'Zdarzenia niezależne' if PA_PC == P_intersection_AC else 'Zdarzenia zależne'
print(check)
```
### <a name='8'></a> Prawdopodobieństwo warunkowe
Prawdopodobieństwem warunkowym zdarzenia $A$ pod warunkiem zajścia zdarzenia $B$ przy zał. $P(B) > 0$ nazywamy:
$$P(A|B) = \frac{P(A\cap B)}{P(B)}.$$
```
print(f'A: {A}')
print(f'B: {B}')
print(f'Przecięcie A i B: {intersection_AB}')
PA_cond_B = P_intersection_AB / P_B
print(f'Prawdopodobieństwo wylosowania liczby parzystej pod warunkiem, że wylosowaliśmy liczbę większą niż 4 wynosi: {PA_cond_B}')
```
### <a name='9'> </a> Zmienna Losowa
### Dystrybuanta
Niech $X$ będzie zmienną losową z przestrzeni $R^n$. Niech $F(x)$ oznacza dystrybuantę zmiennej losowej $X$, tzn.
$$F(x) = P(X \leq x)$$
Jeśli dystrybuanta jest funkcją ciągłą to funkcja określona wzorem
$$f(x) = \frac{dF(x)}{dx} = F'(x)$$
jest nazywana **gęstością rozkładu** zmiennej losowej $X$.
Stąd mamy $$P(a < X \leq b) = \int_a^bf(x)\ dx$$
### Wartość oczekiwana
Wartością oczekiwaną zmiennej losowej ciągłej nazywamy wartość:
$$E(X) = \int_{-\infty}^{\infty}x \cdot f(x)\ dx$$
Wartością oczekiwaną zmiennej losowej dyskretnej nazywamy wartość:
$$E(X) = \sum_{i=1}^{n}x_i \cdot P(X=x_i)$$
Dla klasycznej przestrzeni probabilistycznej, gdzie każde zdarzenie ma jednakowe prawdopodobieństwo zajścia otrzymujemy klasyczny wzór:
$$E(X) = \frac{1}{n}\sum_{i=1}^{n}x_{i}$$
### Wariancja
Warjancją zmiennej losowej nazywamy wartość:
$$Var(X) = E(X-E(X))^2 = E(X^2) - (E(X))^2$$
Stąd wariancja zmiennej losowej ciągłęj ma postać:
$$Var(X) = D^2(X) = \int_{-\infty}^{\infty}(x - E(X))^2 \cdot f(x)\ dx$$
### Odchylenie Standardowe
Odchyleniem standardowym nazywamy pierwiastek z wariancji, tj.
$$\sigma(X) = \sqrt{D^2(X)}$$
### Kowariancja
$X, Y$ - zmienne losowe
$$Cov(X, Y) = E(X-E(X))(Y-E(Y))= E(XY) - E(X)E(Y)$$
### Korelacja
$$Cor(X, Y) = \frac{Cov(X, Y)}{\sigma(X)\cdot \sigma(Y)}$$
### Błąd standardowy
$$SE(X) = \frac{\sigma(X)}{\sqrt(n)}$$
### Przykład 1
| $X$ | 0 | 1 | 2 | 3 |
|----------|-----|-----|------|------|
| $P(X=x_i)$ | 0.5 | 0.3 | 0.15 | 0.05 |
$E(X) = 0 \cdot 0.5 + 1 \cdot 0.3 + 2 \cdot 0.15 + 3 \cdot 0.05$
```
EX = 0 * 0.5 + 1 * 0.3 + 2 * 0.15 + 3 * 0.05
EX
VAR_X = (0 - EX)**2 * 0.5 + (1 - EX)**2 * 0.3 + (2 - EX)**2 * 0.15 +(3 - EX)**2 * 0.05
VAR_X
import numpy as np
STD_X = np.sqrt(VAR_X)
STD_X
```
Przykład
```
x = np.random.randn(5)
y = np.random.randn(5)
print(x, '\n')
print(y)
x.mean()
x.std()
np.corrcoef(x, y)
```
### Rozkład normalny
Rozkład normalny $X \sim N(\mu, \sigma)$
```
import matplotlib.pyplot as plt
import seaborn as sns
from scipy.stats import norm
sns.set()
mu = 0 # średnia
sigma = 1 # odchylenie standardowe
x = np.linspace(mu - 3 * sigma, mu + 3 * sigma, 100)
plt.title('Rozkład normalny - funkcja gęstości')
_ = plt.plot(x, norm.pdf(x, mu, sigma))
x = np.linspace(mu - 5 * sigma, mu + 5 * sigma, 500)
params = [(0, 1), (0, 2), (0, 0.5)]
plt.figure(figsize=(8, 8))
for mu, sigma in params:
plt.plot(x, norm.pdf(x, mu, sigma), label=f'mu={mu}, sigma={sigma}')
plt.legend()
```
### Standaryzacja
Niech $X \sim N(\mu, \sigma)$. Wówczas
$$\frac{X - \mu}{\sigma} \sim N(0, 1)$$
$$E\left(\frac{X - \mu}{\sigma}\right) = \frac{E(X - \mu)}{\sigma} = \frac{E(X) - \mu}{\sigma} = \frac{\mu - \mu}{\sigma} = 0$$
$$D^2\left(\frac{X - \mu}{\sigma}\right) = \frac{1}{\sigma^2}\cdot D^2\left(X - \mu\right) = \frac{1}{\sigma^2} \cdot D^2(X) = \frac{1}{\sigma^2}\cdot \sigma^2 = 1$$
```
mu = 10
sigma = 3
x = np.linspace(mu - 8 * sigma, mu + 8 * sigma, 500)
plt.plot(x, norm.pdf(x, mu, sigma), label=f'mu={mu}, sigma={sigma}')
plt.plot(x, norm.pdf(x, mu - mu, sigma), label=f'mu={mu - mu}, sigma={sigma}')
plt.plot(x, norm.pdf(x, mu - mu, sigma / sigma), label=f'mu={mu - mu}, sigma={sigma / sigma}')
plt.legend()
```
### Wykres dystrybuanty
```
plt.title('Rozkład normalny - dystrybuanta')
_ = plt.plot(x, norm.cdf(x, mu, sigma))
plt.title('Rozkład normalny - funkcja przeżycia')
_ = plt.plot(x, norm.sf(x, mu, sigma))
from scipy import stats
dir(stats)
```
| github_jupyter |
# Importing Requires Libraries
```
import numpy as np
import seaborn as sns
import pandas as pd
import matplotlib.pyplot as plt
%matplotlib inline
from sklearn.linear_model import LogisticRegression
from sklearn.neighbors import KNeighborsClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split, cross_val_score
from sklearn.model_selection import RandomizedSearchCV, GridSearchCV
from sklearn.metrics import confusion_matrix, classification_report
from sklearn.metrics import precision_score, recall_score, f1_score
from sklearn.metrics import plot_roc_curve
```
# Loading Data
```
from google.colab import drive
drive.mount('/content/drive')
df = pd.read_csv("/content/drive/My Drive/DataSets/heart-disease.csv")
df
```
# Exploratory Data Analysis
```
df.head()
df.tail()
df["target"].value_counts()
df["target"].value_counts().plot(kind="bar", color=["salmon", "lightgreen"]);
df.info()
df.isna().sum()
df.describe()
df["sex"].value_counts()
pd.crosstab(df.target, df.sex)
pd.crosstab(df.target, df.sex).plot(kind="bar",
figsize=(10, 6),
color=["salmon", "lightblue"]);
plt.title("Heart Disease Frequency for Sex")
plt.xlabel("0 = No Disease, 1 = Disease")
plt.ylabel("Amount")
plt.legend(["Female", "Male"])
plt.xticks(rotation=0);
```
A women has more chances of getting a heart disease than a man according to this data
```
df["thalach"].value_counts()
plt.figure(figsize=(10, 8))
plt.scatter(df.age[df.target==1],
df.thalach[df.target==1],
c="salmon")
plt.scatter(df.age[df.target==0],
df.thalach[df.target==0],
c="lightblue");
plt.title("Heart Disease as a function of Age and Max Heart Rate")
plt.xlabel("Age")
plt.ylabel("Max Heart Rate")
plt.legend(["Disease", "No Disease"]);
```
As People get Older the Max Heart Rate decreases
```
df.age.plot.hist();
```
Most of the people lie in the range 50 - 65 years of age
```
pd.crosstab(df.cp, df.target)
pd.crosstab(df.cp, df.target).plot(kind="bar",
figsize=(10, 10),
color=["salmon", "lightblue"])
plt.title("Heart Disease Frequency per Chest Pain Type")
plt.xlabel("Chest Pain Type")
plt.ylabel("Amount")
plt.legend(["No Disease", "Disease"])
plt.xticks(rotation=0);
```
People having Chest Pain Type of 2 have more Chances of a Heart Disease than any other pain type
```
df.corr()
corr_matrix = df.corr()
fig, ax = plt.subplots(figsize=(15, 12))
ax = sns.heatmap(corr_matrix,
annot=True,
linewidths=0.5,
fmt=".2f",
cmap="YlGnBu");
```
* Thalach, CP, Slope - Have high postive correlation with target variable
* Thal, CA, OldPeak, Exang - Have high negative correlation with target variable
* Slope and Thalace have a significant Correlation value between them
# Modelling
```
df.head()
x = df.drop("target", axis=1)
y = df["target"]
x.head()
x = x.drop(["trestbps", "chol", "fbs", "restecg", "slope"], axis=1)
x.head()
y
np.random.seed(99)
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size = 0.2)
x_train.head(), len(x_train)
y_train.head(), len(y_train)
models = {"Logistic Regression": LogisticRegression(),
"KNN": KNeighborsClassifier(),
"Random Forest": RandomForestClassifier()}
def fit_and_score(models, x_train, x_test, y_train, y_test):
np.random.seed(99)
model_scores = {}
for name, model in models.items():
model.fit(x_train, y_train)
model_scores[name] = model.score(x_test, y_test)
return model_scores
model_scores = fit_and_score(models, x_train, x_test, y_train, y_test)
model_scores
model_compare = pd.DataFrame(model_scores, index=["accuracy"])
model_compare.T.plot.bar()
```
What else should we do to gurantee a CLASSIFICATION model's correctness and further improve it?
* Hyperparameter Tuning
* Feature Importance
* Confusion Matrix
* Cross-Validation
* Precision
* Recall
* F1 Score
* Classification Report
* ROC Curve
* Area under the curve (AUC)
## Tuning KNN
```
train_scores = []
test_scores = []
neighbors = range(1, 21)
knn = KNeighborsClassifier()
for i in neighbors:
knn.set_params(n_neighbors=i)
knn.fit(x_train, y_train)
train_scores.append(knn.score(x_train, y_train))
test_scores.append(knn.score(x_test, y_test))
train_scores
test_scores
plt.plot(neighbors, train_scores, label="Train score")
plt.plot(neighbors, test_scores, label="Test score")
plt.xticks(np.arange(1, 21, 1))
plt.xlabel("Number of neighbors")
plt.ylabel("Model Score")
plt.legend()
print(f"Maximum KNN score on the test data: {max(test_scores)*100:.2f}%");
```
## Tuning LogisticRegression and RandomForestClassifier with RandomizedSearchCV
```
# Parameter Grid For Logistic Regression
log_reg_grid = {"C": np.logspace(-4, 10, 20),
"solver": ["liblinear"]}
# Parameter Grid For RandomForest
rf_grid = {"n_estimators": np.arange(10, 1600, 90),
"max_depth": [None, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10],
"min_samples_split": np.arange(1, 22, 2),
"min_samples_leaf": np.arange(1, 22, 2)}
# Tuning LogisticRegression
np.random.seed(99)
rs_log_reg = RandomizedSearchCV(LogisticRegression(),
param_distributions=log_reg_grid,
cv=5,
n_iter=50,
verbose=True)
rs_log_reg.fit(x_train, y_train)
rs_log_reg.best_params_
rs_log_reg.score(x_test, y_test)
# Tuning RandomForestClassifier
np.random.seed(99)
rs_rf = RandomizedSearchCV(RandomForestClassifier(),
param_distributions=rf_grid,
cv=5,
n_iter=50,
verbose=True)
rs_rf.fit(x_train, y_train)
rs_rf.best_params_
rs_rf.score(x_test, y_test)
```
## Tuning Models with GridSearchCV
```
log_reg_grid = {"C": np.logspace(-4, 4, 30),
"solver": ["liblinear"]}
gs_log_reg = GridSearchCV(LogisticRegression(),
param_grid=log_reg_grid,
cv=5,
verbose=True)
gs_log_reg.fit(x_train, y_train);
gs_log_reg.best_params_
gs_log_reg.score(x_test, y_test)
gs_rf = GridSearchCV(RandomForestClassifier(),
param_grid=rf_grid,
cv=5,
verbose=True)
gs_rf.fit(x_train, y_train);
```
# Evaluation
```
y_preds = rs_rf.predict(x_test)
y_preds
y_test
#plot ROC curve and caluclate AUC metric
plot_roc_curve(rs_rf, x_test, y_test)
# confusion matrix
print(confusion_matrix(y_test, y_preds))
sns.set(font_scale=1.8)
def plot_conf_mat(y_test, y_preds):
fig, ax = plt.subplots(figsize=(3,3))
ax = sns.heatmap(confusion_matrix(y_test, y_preds),
annot=True,
cbar=False)
plt.xlabel("Predicted Label")
plt.ylabel("True Label")
plot_conf_mat(y_test, y_preds)
print(classification_report(y_test, y_preds))
```
* Recall - No false Negative => Recall=1
* Precision - No False Positive => Precision=1
Classification Report after performing Cross Validation -
```
rs_rf.best_params_
rf_clf = RandomForestClassifier(n_estimators=460,
max_depth=9,
min_samples_split=5,
min_samples_leaf=3)
cv_acc = cross_val_score(rf_clf, x, y, cv=5, scoring="accuracy")
cv_acc
cv_acc = np.mean(cv_acc)
cv_acc
cv_precision = cross_val_score(rf_clf, x, y, cv=5, scoring="precision")
cv_precision = np.mean(cv_precision)
cv_precision
cv_recall = cross_val_score(rf_clf, x, y, cv=5, scoring="recall")
cv_recall = np.mean(cv_recall)
cv_recall
cv_f1 = cross_val_score(rf_clf, x, y, cv=5, scoring="f1")
cv_f1 = np.mean(cv_f1)
cv_f1
cv_metrics = pd.DataFrame({"Accuracy": cv_acc,
"Precision": cv_precision,
"Recall": cv_recall,
"F1": cv_f1}, index=[0])
cv_metrics.T.plot.bar(title="Cross-Validated Classification Metrics", legend=False);
```
# Feature Importance
```
rf_clf.fit(x_train, y_train)
rf_clf.feature_importances_
feature_dict = dict(zip(df.columns, list(rf_clf.feature_importances_)))
feature_dict
feature_df = pd.DataFrame(feature_dict, index=[0])
feature_df.T.plot.bar(title="Feature Importance", legend=False);
import joblib
joblib.dump(rf_clf, 'heart_disease.mdl')
```
| github_jupyter |
### Summary:
This notebook contains the soft smoothing figures for Reed (Figure 3(b)).
## Load libraries
```
# import packages
from __future__ import division
import networkx as nx
import numpy as np
import os
from sklearn import metrics
from sklearn.preprocessing import label_binarize
from sklearn.metrics import f1_score
from sklearn.metrics import roc_auc_score
from sklearn.model_selection import StratifiedShuffleSplit
from sklearn.model_selection import ShuffleSplit
import matplotlib.pyplot as plt
import itertools
from numpy.linalg import inv
## function to create + save dictionary of features
def create_dict(key, obj):
return(dict([(key[i], obj[i]) for i in range(len(key))]))
```
## load helper functions and dataset
```
# set the working directory and import helper functions
#get the current working directory and then redirect into the functions under code
cwd = os.getcwd()
# parents working directory of the current directory: which is the code folder
parent_cwd = os.path.dirname(cwd)
# get into the functions folder
functions_cwd = parent_cwd + '/functions'
# change the working directory to be .../functions
os.chdir(functions_cwd)
# import all helper functions
exec(open('parsing.py').read())
exec(open('ZGL.py').read())
exec(open('create_graph.py').read())
exec(open('ZGL_softing_new_new.py').read())
exec(open('one_hop_majority_vote.py').read())
exec(open('iterative_method_test.py').read())
#exec(open('iterative_method_test_new.py').read())
exec(open('decoupling_prepare.py').read())
# import the data from the data folder
data_cwd = os.path.dirname(parent_cwd)+ '/data'
# change the working directory and import the fb dataset
fb100_file = data_cwd +'/Reed98'
A, metadata = parse_fb100_mat_file(fb100_file)
# change A(scipy csc matrix) into a numpy matrix
adj_matrix_tmp = A.todense()
#get the gender for each node(1/2,0 for missing)
gender_y_tmp = metadata[:,1]
# get the corresponding gender for each node in a disctionary form
gender_dict = create_dict(range(len(gender_y_tmp)), gender_y_tmp)
(graph, gender_y) = create_graph(adj_matrix_tmp,gender_dict,'gender',0,None,'yes')
```
## Setup
```
percent_initially_unlabelled = [0.99,0.95,0.9,0.8,0.7,0.6,0.5,0.4,0.3,0.2,0.1,0.05]
percent_initially_labelled = np.subtract(1, percent_initially_unlabelled)
n_iter = 10
cv_setup='stratified'
```
## 1-hop and 2-hop MV method
```
# 1-hop and 2-hop initialization
A = nx.adjacency_matrix(graph)
adj_matrix_gender = np.matrix(A.todense())
n = len(gender_y)
keys = list(graph.node)
# see how many classes are there and rearrange them
classes = np.sort(np.unique(gender_y))
class_labels = np.array(range(len(classes)))
# relabel membership class labels - for coding convenience
# preserve ordering of original class labels -- but force to be in sequential order now
gender_y_update = np.copy(gender_y)
for j in range(len(classes)):
gender_y_update[gender_y_update == classes[j]] = class_labels[j]
gender_y_update = np.array(gender_y_update)
mean_accuracy_one = []
se_accuracy_one = []
mean_micro_auc_one = []
se_micro_auc_one = []
mean_wt_auc_one = []
se_wt_auc_one = []
mean_accuracy_two = []
se_accuracy_two = []
mean_micro_auc_two = []
se_micro_auc_two = []
mean_wt_auc_two = []
se_wt_auc_two = []
for i in range(len(percent_initially_labelled)):
print(percent_initially_unlabelled[i])
(graph_new, gender_y_new) = create_graph(adj_matrix_tmp,gender_dict,'gender',0,None,'yes')
adj_matrix_gender = np.array(nx.adjacency_matrix(graph_new).todense())
gender_dict_input = create_dict(range(len(gender_y_new)), gender_y_new)
(graph_input, gender_y_input) = create_graph(adj_matrix_gender,gender_dict_input,'gender',5,None,'yes')
keys = list(graph_input.node())
bench_mark = np.mean(np.array(gender_y_input) == np.max(class_labels))
if cv_setup=='stratified':
k_fold = StratifiedShuffleSplit(n_splits=n_iter,test_size=percent_initially_unlabelled[i],
random_state=1)
else:
k_fold = cross_validation.ShuffleSplit(n_splits=n_iter,
test_size=percent_initially_unlabelled[i],
random_state=1)
accuracy_one = []
micro_auc_one = []
wt_auc_one = []
accuracy_two = []
micro_auc_two = []
wt_auc_two = []
# update rule for theta
for train_index, test_index in k_fold.split(keys, gender_y_input):
accuracy_score_benchmark = np.mean(np.array(gender_y_input)[train_index] == np.max(class_labels))
# get 1-hop result
(theta_one_tmp, micro_auc_one_tmp, wt_auc_one_tmp, accuracy_one_tmp) = one_hop_majority_vote(graph_input, gender_y_input,train_index, test_index)
accuracy_one.append(accuracy_one_tmp)
micro_auc_one.append(micro_auc_one_tmp)
wt_auc_one.append(wt_auc_one_tmp)
# get 2-hop result
bench_mark = gender_y_input/len(gender_y_input)
(theta_two_tmp, micro_auc_two_tmp, wt_auc_two_tmp, accuracy_two_tmp) = iterative_method_test(1, graph_input, gender_y_input, gender_y_input, train_index, test_index, bench_mark)
accuracy_two.append(accuracy_two_tmp)
micro_auc_two.append(micro_auc_two_tmp)
wt_auc_two.append(wt_auc_two_tmp)
# get the mean and standard deviation
mean_accuracy_one.append(np.mean(accuracy_one))
se_accuracy_one.append(np.std(accuracy_one))
mean_micro_auc_one.append(np.mean(micro_auc_one))
se_micro_auc_one.append(np.std(micro_auc_one))
mean_wt_auc_one.append(np.mean(wt_auc_one))
se_wt_auc_one.append(np.std(wt_auc_one))
# get the mean and standard deviation
mean_accuracy_two.append(np.mean(accuracy_two))
se_accuracy_two.append(np.std(accuracy_two))
mean_micro_auc_two.append(np.mean(micro_auc_two))
se_micro_auc_two.append(np.std(micro_auc_two))
mean_wt_auc_two.append(np.mean(wt_auc_two))
se_wt_auc_two.append(np.std(wt_auc_two))
```
## decoupled smoothing method (with parameter 0.1)
```
sigma_square = 0.1
(graph, gender_y) = create_graph(adj_matrix_tmp,gender_dict,'gender',0,None,'yes')
A_tilde = decoupling_prepare(graph,sigma_square)
(mean_accuracy_decoupling_Reed_01,se_accuracy_decoupling_Reed_01,mean_micro_auc_decoupling_Reed_01,se_micro_auc_decoupling_Reed_01,mean_wt_auc_decoupling_Reed_01,se_wt_auc_decoupling_Reed_01) = ZGL(np.array(A_tilde),gender_y,percent_initially_unlabelled, n_iter, cv_setup)
```
## hard smoothing method (ZGL)
```
adj_matrix_tmp_ZGL = adj_matrix_tmp
(graph, gender_y) = create_graph(adj_matrix_tmp_ZGL,gender_dict,'gender',0,None,'yes')
# run ZGL
(mean_accuracy_zgl_Reed, se_accuracy_zgl_Reed,
mean_micro_auc_zgl_Reed,se_micro_auc_zgl_Reed,
mean_wt_auc_zgl_Reed,se_wt_auc_zgl_Reed) =ZGL(np.array(adj_matrix_gender),
np.array(gender_y),percent_initially_unlabelled,
n_iter,cv_setup)
```
## Plot
AUC against Initial unlabled node precentage
```
%matplotlib inline
from matplotlib.ticker import FixedLocator,LinearLocator,MultipleLocator, FormatStrFormatter
fig = plt.figure()
#seaborn.set_style(style='white')
from mpl_toolkits.axes_grid1 import Grid
grid = Grid(fig, rect=111, nrows_ncols=(1,1),
axes_pad=0.1, label_mode='L')
for i in range(4):
if i == 0:
grid[i].xaxis.set_major_locator(FixedLocator([0,25,50,75,100]))
grid[i].yaxis.set_major_locator(FixedLocator([0.4, 0.5,0.6,0.7,0.8,0.9,1]))
grid[i].errorbar(percent_initially_labelled*100,mean_wt_auc_zgl_Reed,
yerr=se_wt_auc_zgl_Reed, fmt='--o', capthick=2,
alpha=1, elinewidth=3, color='orange')
grid[i].errorbar(percent_initially_labelled*100, mean_wt_auc_one,
yerr=se_wt_auc_one, fmt='--o', capthick=2,
alpha=1, elinewidth=3, color='red')
grid[i].errorbar(percent_initially_labelled*100, mean_wt_auc_two,
yerr=se_wt_auc_one, fmt='--o', capthick=2,
alpha=1, elinewidth=3, color='maroon')
grid[i].errorbar(percent_initially_labelled*100, mean_wt_auc_decoupling_Reed_01,
yerr=se_wt_auc_decoupling_Reed_01, fmt='--o', capthick=2,
alpha=1, elinewidth=3, color='black')
grid[i].set_ylim(0.4,1.1)
grid[i].set_xlim(0,101)
grid[i].annotate('1-hop MV', xy=(3, 0.80),
color='red', alpha=1, size=12)
grid[i].annotate('2-hop MV', xy=(3, 0.84),
color='maroon', alpha=1, size=12)
grid[i].annotate('hard smoothing', xy=(3, 0.88),
color='orange', alpha=1, size=12)
grid[i].annotate('decoupled smoothing', xy=(3, 0.92),
color='black', alpha=1, size=12)
grid[i].set_ylim(0.4,1.01)
grid[i].set_xlim(0,100)
grid[i].spines['right'].set_visible(False)
grid[i].spines['top'].set_visible(False)
grid[i].tick_params(axis='both', which='major', labelsize=13)
grid[i].tick_params(axis='both', which='minor', labelsize=13)
grid[i].set_xlabel('Percent of Nodes Initially Labeled').set_fontsize(15)
grid[i].set_ylabel('AUC').set_fontsize(15)
grid[0].set_xticks([0,25, 50, 75, 100])
grid[0].set_yticks([0.4,0.6,0.8,1])
grid[0].minorticks_on()
grid[0].tick_params('both', length=4, width=1, which='major', left=1, bottom=1, top=0, right=0)
from matplotlib.backends.backend_pdf import PdfPages
figure_cwd = os.path.dirname(parent_cwd) + '/figure'
pp = PdfPages.savefig(fname = figure_cwd + '/decoupled_smoothing_Reed.pdf')
pp.savefig(dpi = 300)
pp.close()
#fig.savefig('Sub Directory/graph.png')
# get into the functions folder
figure_cwd = os.path.dirname(parent_cwd) + '/figure'
figure_cwd
```
| github_jupyter |
<a href="https://colab.research.google.com/github/tensorflow/tpu/blob/master/tools/colab/shakespeare_with_tpu_and_keras.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
##### Copyright 2018 The TensorFlow Hub Authors.
Licensed under the Apache License, Version 2.0 (the "License");
```
# Copyright 2018 The TensorFlow Hub Authors. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
```
## Predict Shakespeare with Cloud TPUs and Keras
## Overview
This example uses [tf.keras](https://www.tensorflow.org/guide/keras) to build a *language model* and train it on a Cloud TPU. This language model predicts the next character of text given the text so far. The trained model can generate new snippets of text that read in a similar style to the text training data.
The model trains for 10 epochs and completes in approximately 5 minutes.
This notebook is hosted on GitHub. To view it in its original repository, after opening the notebook, select **File > View on GitHub**.
## Learning objectives
In this Colab, you will learn how to:
* Build a two-layer, forward-LSTM model.
* Use distribution strategy to produce a `tf.keras` model that runs on TPU version and then use the standard Keras methods to train: `fit`, `predict`, and `evaluate`.
* Use the trained model to make predictions and generate your own Shakespeare-esque play.
## Instructions
<h3> Train on TPU <a href="https://cloud.google.com/tpu/"><img valign="middle" src="https://raw.githubusercontent.com/GoogleCloudPlatform/tensorflow-without-a-phd/master/tensorflow-rl-pong/images/tpu-hexagon.png" width="50"></a></h3>
1. On the main menu, click Runtime and select **Change runtime type**. Set "TPU" as the hardware accelerator.
1. Click Runtime again and select **Runtime > Run All**. You can also run the cells manually with Shift-ENTER.
TPUs are located in Google Cloud, for optimal performance, they read data directly from Google Cloud Storage (GCS)
## Data, model, and training
In this example, you train the model on the combined works of William Shakespeare, then use the model to compose a play in the style of *The Great Bard*:
<blockquote>
Loves that led me no dumbs lack her Berjoy's face with her to-day.
The spirits roar'd; which shames which within his powers
Which tied up remedies lending with occasion,
A loud and Lancaster, stabb'd in me
Upon my sword for ever: 'Agripo'er, his days let me free.
Stop it of that word, be so: at Lear,
When I did profess the hour-stranger for my life,
When I did sink to be cried how for aught;
Some beds which seeks chaste senses prove burning;
But he perforces seen in her eyes so fast;
And _
</blockquote>
### Download data
Download *The Complete Works of William Shakespeare* as a single text file from [Project Gutenberg](https://www.gutenberg.org/). You use snippets from this file as the *training data* for the model. The *target* snippet is offset by one character.
```
!wget --show-progress --continue -O /content/shakespeare.txt http://www.gutenberg.org/files/100/100-0.txt
```
### Build the input dataset
We just downloaded some text. The following shows the start of the text and a random snippet so we can get a feel for the whole text.
```
!head -n5 /content/shakespeare.txt
!echo "..."
!shuf -n5 /content/shakespeare.txt
import numpy as np
import tensorflow as tf
import os
import distutils
if distutils.version.LooseVersion(tf.__version__) < '1.14':
raise Exception('This notebook is compatible with TensorFlow 1.14 or higher, for TensorFlow 1.13 or lower please use the previous version at https://github.com/tensorflow/tpu/blob/r1.13/tools/colab/shakespeare_with_tpu_and_keras.ipynb')
# This address identifies the TPU we'll use when configuring TensorFlow.
TPU_WORKER = 'grpc://' + os.environ['COLAB_TPU_ADDR']
SHAKESPEARE_TXT = '/content/shakespeare.txt'
def transform(txt):
return np.asarray([ord(c) for c in txt if ord(c) < 255], dtype=np.int32)
def input_fn(seq_len=100, batch_size=1024):
"""Return a dataset of source and target sequences for training."""
with tf.io.gfile.GFile(SHAKESPEARE_TXT, 'r') as f:
txt = f.read()
source = tf.constant(transform(txt), dtype=tf.int32)
ds = tf.data.Dataset.from_tensor_slices(source).batch(seq_len+1, drop_remainder=True)
def split_input_target(chunk):
input_text = chunk[:-1]
target_text = chunk[1:]
return input_text, target_text
BUFFER_SIZE = 10000
ds = ds.map(split_input_target).shuffle(BUFFER_SIZE).batch(batch_size, drop_remainder=True)
return ds.repeat()
```
### Build the model
The model is defined as a two-layer, forward-LSTM, the same model should work both on CPU and TPU.
Because our vocabulary size is 256, the input dimension to the Embedding layer is 256.
When specifying the arguments to the LSTM, it is important to note how the stateful argument is used. When training we will make sure that `stateful=False` because we do want to reset the state of our model between batches, but when sampling (computing predictions) from a trained model, we want `stateful=True` so that the model can retain information across the current batch and generate more interesting text.
```
EMBEDDING_DIM = 512
def lstm_model(seq_len=100, batch_size=None, stateful=True):
"""Language model: predict the next word given the current word."""
source = tf.keras.Input(
name='seed', shape=(seq_len,), batch_size=batch_size, dtype=tf.int32)
embedding = tf.keras.layers.Embedding(input_dim=256, output_dim=EMBEDDING_DIM)(source)
lstm_1 = tf.keras.layers.LSTM(EMBEDDING_DIM, stateful=stateful, return_sequences=True)(embedding)
lstm_2 = tf.keras.layers.LSTM(EMBEDDING_DIM, stateful=stateful, return_sequences=True)(lstm_1)
predicted_char = tf.keras.layers.TimeDistributed(tf.keras.layers.Dense(256, activation='softmax'))(lstm_2)
return tf.keras.Model(inputs=[source], outputs=[predicted_char])
```
### Train the model
First, we need to create a distribution strategy that can use the TPU. In this case it is TPUStrategy. You can create and compile the model inside its scope. Once that is done, future calls to the standard Keras methods `fit`, `evaluate` and `predict` use the TPU.
Again note that we train with `stateful=False` because while training, we only care about one batch at a time.
```
tf.keras.backend.clear_session()
resolver = tf.contrib.cluster_resolver.TPUClusterResolver(TPU_WORKER)
tf.contrib.distribute.initialize_tpu_system(resolver)
strategy = tf.contrib.distribute.TPUStrategy(resolver)
with strategy.scope():
training_model = lstm_model(seq_len=100, stateful=False)
training_model.compile(
optimizer=tf.keras.optimizers.RMSprop(learning_rate=0.01),
loss='sparse_categorical_crossentropy',
metrics=['sparse_categorical_accuracy'])
training_model.fit(
input_fn(),
steps_per_epoch=100,
epochs=10
)
training_model.save_weights('/tmp/bard.h5', overwrite=True)
```
### Make predictions with the model
Use the trained model to make predictions and generate your own Shakespeare-esque play.
Start the model off with a *seed* sentence, then generate 250 characters from it. The model makes five predictions from the initial seed.
The predictions are done on the CPU so the batch size (5) in this case does not have to be divisible by 8.
Note that when we are doing predictions or, to be more precise, text generation, we set `stateful=True` so that the model's state is kept between batches. If stateful is false, the model state is reset between each batch, and the model will only be able to use the information from the current batch (a single character) to make a prediction.
The output of the model is a set of probabilities for the next character (given the input so far). To build a paragraph, we predict one character at a time and sample a character (based on the probabilities provided by the model). For example, if the input character is "o" and the output probabilities are "p" (0.65), "t" (0.30), others characters (0.05), then we allow our model to generate text other than just "Ophelia" and "Othello."
```
BATCH_SIZE = 5
PREDICT_LEN = 250
# Keras requires the batch size be specified ahead of time for stateful models.
# We use a sequence length of 1, as we will be feeding in one character at a
# time and predicting the next character.
prediction_model = lstm_model(seq_len=1, batch_size=BATCH_SIZE, stateful=True)
prediction_model.load_weights('/tmp/bard.h5')
# We seed the model with our initial string, copied BATCH_SIZE times
seed_txt = 'Looks it not like the king? Verily, we must go! '
seed = transform(seed_txt)
seed = np.repeat(np.expand_dims(seed, 0), BATCH_SIZE, axis=0)
# First, run the seed forward to prime the state of the model.
prediction_model.reset_states()
for i in range(len(seed_txt) - 1):
prediction_model.predict(seed[:, i:i + 1])
# Now we can accumulate predictions!
predictions = [seed[:, -1:]]
for i in range(PREDICT_LEN):
last_word = predictions[-1]
next_probits = prediction_model.predict(last_word)[:, 0, :]
# sample from our output distribution
next_idx = [
np.random.choice(256, p=next_probits[i])
for i in range(BATCH_SIZE)
]
predictions.append(np.asarray(next_idx, dtype=np.int32))
for i in range(BATCH_SIZE):
print('PREDICTION %d\n\n' % i)
p = [predictions[j][i] for j in range(PREDICT_LEN)]
generated = ''.join([chr(c) for c in p]) # Convert back to text
print(generated)
print()
assert len(generated) == PREDICT_LEN, 'Generated text too short'
```
## What's next
* Learn about [Cloud TPUs](https://cloud.google.com/tpu/docs) that Google designed and optimized specifically to speed up and scale up ML workloads for training and inference and to enable ML engineers and researchers to iterate more quickly.
* Explore the range of [Cloud TPU tutorials and Colabs](https://cloud.google.com/tpu/docs/tutorials) to find other examples that can be used when implementing your ML project.
On Google Cloud Platform, in addition to GPUs and TPUs available on pre-configured [deep learning VMs](https://cloud.google.com/deep-learning-vm/), you will find [AutoML](https://cloud.google.com/automl/)*(beta)* for training custom models without writing code and [Cloud ML Engine](https://cloud.google.com/ml-engine/docs/) which will allows you to run parallel trainings and hyperparameter tuning of your custom models on powerful distributed hardware.
| github_jupyter |
# Denoising with a synthetic DNN prior
Suppose we are given a prior $G$ parameterized by an expansive DNN. Suppose the prior $G$ maps a $k$-dimensional input to a $n$-dimensional output, where $K \ll N$. Furthermore, we let the weights be Gaussian distributed, and the activation functions are ReLus.
We consider the classical image or data denoising problem, where the goal is to remove zero-mean white Gaussian noise from a given image or data point. In more detail, our goal is to obtain an estimate of a vector $y_0 \in \mathbb R^n$ from the noisy observation
$$
y = y_0 + \eta,
$$
where $\eta$ is zero-mean Gaussian noise with covariance matrix $\sigma^2/n I$, and $y_0$ lies in the range of the generator, i.e., $y_0=G(x_0)$.
We consider the following two-step denoising algorithm:
1. Obtaine an estimate $\hat x$ of the latent representation by minimizing the empirical loss
$$
f(x) = \|G(x) - y_0\|_2^2
$$
using gradient descent.
2. Obtain an estimate of the image as $\hat y = G(\hat x)$.
```
import torch
from torch.autograd import Variable
import torch.nn as nn
import torch.nn.functional as F
import torch.utils.data
import torch.optim as optim
import torchvision
from torchvision import datasets, transforms
import numpy as np
from IPython.core.debugger import Tracer
import matplotlib.pyplot as plt
```
# Denoising with a random prior
Suppose we add Gaussian noise to the output. What is the denoising rate if we reconstruct the input from the output, with empirical risk minimization?
```
def init_weights(m):
if type(m) == nn.Linear:
m.weight.data.normal_(0.0, 0.1)
def remove_grad(m):
if type(m) == nn.Linear:
m.weight.requires_grad = False
# generate a random DNN prior
def generate_random_prior(K,n_hidden = 500, n_out = 1500):
# define the network
net = nn.Sequential(
nn.Linear(in_features=K, out_features=n_hidden, bias=False),
nn.ReLU(),
nn.Linear(in_features=n_hidden, out_features=n_out, bias=False),
nn.ReLU(),
)
# initialize weights
net.apply(init_weights)
net.apply(remove_grad)
return net
# function to estimate the latent representation
def invert_prior(net, noisy_image,latent_param,learning_rate=0.1,num_steps=500):
optimizer = torch.optim.SGD([latent_param], lr=learning_rate)
iter_idx = 0
while iter_idx <= num_steps:
# correct the values of updated input signal
optimizer.zero_grad()
output = net(latent_param)
loss = F.mse_loss(output, noisy_image, size_average=False)
loss.backward()
optimizer.step()
iter_idx += 1
return latent_param
# denoise by recovering estimating a latent representation and passing that through the decoder
def denoise(net,noisy_image,K):
rec_rep = Variable(torch.randn(K), requires_grad=True)
rec_rep = invert_prior(net, noisy_image, rec_rep)
return net(rec_rep)
def sqnorm(a):
return sum(a.data.numpy()*a.data.numpy())
```
First, we demonstrate that in the noiseless case, we can perfectly recover the latent representation, provided $K$ is not too large:
```
n = 1500
K = 50
net = generate_random_prior(K)
# generate random latent representation
orig_rep = Variable( torch.randn(K) )
orig_img = net(orig_rep) # 784 tensor
sigma = 0.0
noise = Variable( torch.randn(n) )
noise = noise * (sigma*np.sqrt(sqnorm(orig_img) / sqnorm(noise)))
noisy_img = orig_img + noise
# recover image
rec_rep = Variable(torch.randn(K), requires_grad=True)
rec_rep = invert_prior(net, noisy_img, rec_rep, learning_rate=0.02,num_steps=500)
MSErep = sqnorm(orig_rep - rec_rep) / sqnorm(rec_rep)
# MSE in image space:
rec_img = net(rec_rep)
MSE = sqnorm(orig_img - rec_img) / sqnorm(orig_img)
print("Mean square error in latent space (0 if perfect recovery): ",MSErep)
print("Mean square recovery error (0 if perfect recovery): ",MSE)
```
Generate the plots in the paper on the mean squared error:
```
def sqnorm(a):
return np.sum(a.data.numpy()*a.data.numpy())
def simulate(Ks,numit,Sigmas):
MSEimg = np.zeros(len(Ks))
MSErep = np.zeros(len(Ks))
for i,(K,sigma) in enumerate(zip(Ks,Sigmas)):
for it in range(numit):
net = generate_random_prior(K)
# generate random input to the network
orig_rep = Variable( torch.randn(K) )
orig_img = net(orig_rep)
# add noise
noise = Variable( sigma * torch.randn(n) / np.sqrt(n) )
#noise = noise * (sigma*np.sqrt(sqnorm(orig_rep) / sqnorm(noise)))
noisy_img = orig_img + noise
if(sigma > 0):
SNR = sqnorm(orig_img) / sqnorm(noise)
SNRrep = sqnorm(orig_rep) / sqnorm(noise)
#print("SNR/signal: ", SNR, SNRrep )
# recover image
rec_rep = Variable(torch.randn(K), requires_grad=True)
rec_rep = invert_prior(net, noisy_img, rec_rep,0.02)
# MSE in image space:
rec_img = net(rec_rep)
MSE = sqnorm(orig_img - rec_img)
MSEimg[i] += MSE/numit
# MSE in latent space:
MSE = sqnorm(orig_rep - rec_rep)
MSErep[i] += MSE/numit
return MSErep,MSEimg
numit = 100
Ks = [5+10*i for i in range(6)]
Sigmas = [1.]*len(Ks)
MSErep,MSEimg = simulate(Ks,numit,Sigmas)
plt.plot(Ks,MSEimg)
plt.plot(Ks,10*MSErep)
plt.show()
mtx = np.array([Ks, MSEimg, 10*MSErep])
np.savetxt("./denoise_random_dnn_k.dat", mtx.T , delimiter='\t')
numit = 100
Sigmas = [0.5*i for i in range(10)]
Ks = [30]*len(Sigmas)
MSErep,MSEimg = simulate(Ks,numit,Sigmas)
Sigmas = np.array(Sigmas)
plt.plot(Sigmas**2,MSEimg)
plt.plot(Sigmas**2,10*MSErep)
plt.show()
mtx = np.array([Sigmas**2, MSEimg, 10*MSErep])
np.savetxt("./denoise_random_dnn_sigma.dat", mtx.T , delimiter='\t')
```
## Visualizing the loss surface
```
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import axes3d, Axes3D #<-- Note the capitalization!
def l1norm(a):
return sum(abs(a.data.numpy()))
def evaluate(z,y):
x = Variable( torch.ones(2) )
x.data[0] = z
x.data[1] = y
loss = l1norm(net(x) - orig_img)
K = 2
net = generate_random_prior(K,n_hidden = 150, n_out = 512)
# generate latent representation at (1,0)
orig_rep = Variable( torch.ones(2) )
orig_rep.data[1] = 0
orig_img = net(orig_rep) # 784 tensor
print(orig_rep.data)
# print coordinates plotted with pgfplots
def get_grid(zgrid,ygrid):
L = len(zgrid)
X = np.zeros((L,L))
Y = np.zeros((L,L))
Z = np.zeros((L,L))
for i,y in enumerate(ygrid):
for j,z in enumerate(zgrid):
x = Variable( torch.ones(2) )
x.data[0] = z
x.data[1] = y
loss = sqnorm(net(x) - orig_img)
#loss = l1norm(net(x) - orig_img)
X[i,j] = y
Y[i,j] = z
Z[i,j] = loss
# print(y,z,loss)
#print()
return X,Y,Z
```
#### Visualizaiton in relevant regime
```
N = 30
zgrid = [2*i/N for i in range(-N,N)]
ygrid = [i/N for i in range(-N,N)]
X,Y,Z = get_grid(zgrid,ygrid)
fig = plt.figure()
ax = Axes3D(fig) #<-- Note the difference from your original code...
cset = ax.plot_wireframe(X, -Y, Z)
ax.clabel(cset, fontsize=9, inline=1)
plt.show()
```
#### Visualizaiton around the critical point at $-\rho x_\ast$
```
N = 50
zgrid = [0.2*i/N - 0.2 for i in range(-N,N)]
ygrid = [0.5*i/N for i in range(-N,N)]
Xzoom,Yzoom,Zzoom = get_grid(zgrid,ygrid)
fig = plt.figure()
ax = Axes3D(fig)
cset = ax.plot_wireframe(Xzoom, -Yzoom, Zzoom)
ax.clabel(cset, fontsize=9, inline=1)
plt.show()
plt.contourf(Xzoom, -Yzoom, Zzoom,25)
plt.show()
```
| github_jupyter |
```
export_folder = "Data\Spine000\Test2"
filename_prefix = "Q000"
sequence_browser_name = "SegmentationBrowser"
num_previous = 2
from local_vars import root_folder
import os
export_fullpath = os.path.join(root_folder, export_folder)
if not os.path.exists(export_fullpath):
os.makedirs(export_fullpath)
print("Created folder: " + export_fullpath)
print "Export data to: " + export_fullpath
segmentation_name = "Segmentation"
image_name = "Image_Image"
segmentation_node = slicer.util.getFirstNodeByName(segmentation_name, className="vtkMRMLSegmentationNode")
print "Segmentation node: " + segmentation_node.GetName()
image_node = slicer.util.getFirstNodeByName(image_name, className="vtkMRMLScalarVolumeNode")
print "Ultrasound image node: " + image_node.GetName()
# sequence_browser_node = slicer.util.getFirstNodeByName('', className='vtkMRMLSequenceBrowserNode')
sequence_browser_node = slicer.util.getFirstNodeByName(sequence_browser_name, className='vtkMRMLSequenceBrowserNode')
print "Sequence browser node ID: " + str(sequence_browser_node.GetID())
print "Sequence browser node name: " + str(sequence_browser_node.GetName())
num_items = sequence_browser_node.GetNumberOfItems()
n = num_items
sequence_browser_node.SelectFirstItem()
print "Number of images: " + str(n)
original_sequence_browser_name = "SpineScan"
original_sequence_browser = slicer.util.getNode(original_sequence_browser_name)
labelmap_volume_node = slicer.mrmlScene.AddNewNodeByClass('vtkMRMLLabelMapVolumeNode')
ic = vtk.vtkImageCast()
ic.SetOutputScalarTypeToUnsignedChar()
ic.Update()
png_writer = vtk.vtkPNGWriter()
for i in range(n):
slicer.modules.segmentations.logic().ExportVisibleSegmentsToLabelmapNode(segmentation_node, labelmap_volume_node, image_node)
index = sequence_browser_node.GetMasterSequenceNode().GetNthDataNode(i).GetAttribute("SingleSliceSegmentation_OriginalImageIndex")
segmented_image = labelmap_volume_node.GetImageData()
input_image = image_node.GetImageData()
seg_file_name = filename_prefix + "_%04d_segmentation" % i + ".png"
img_file_name = filename_prefix + "_%04d_%d_ultrasound" % (i, num_previous+1) + ".png"
seg_fullname = os.path.join(export_fullpath, seg_file_name)
img_fullname = os.path.join(export_fullpath, img_file_name)
ic.SetInputData(segmented_image)
ic.Update()
png_writer.SetInputData(ic.GetOutput())
png_writer.SetFileName(seg_fullname)
png_writer.Update()
png_writer.Write()
ic.SetInputData(input_image)
ic.Update()
png_writer.SetInputData(ic.GetOutput())
png_writer.SetFileName(img_fullname)
png_writer.Update()
png_writer.Write()
for j in range(num_previous):
#Gets the previous images - must have images available or else kernel will crash - ie. cannot start at image 0 and ask for images preceeding it
prev = original_sequence_browser.GetMasterSequenceNode().GetNthDataNode(int(index) - (j + 1))
prev_image = prev.GetImageData()
prev_file_name = filename_prefix+ "_%04d_%d_ultrasound" % (i, num_previous - j) + ".png"
prev_fullname = os.path.join(export_fullpath, prev_file_name)
ic.SetInputData(prev_image)
ic.Update()
png_writer.SetInputData(ic.GetOutput())
png_writer.SetFileName(prev_fullname)
png_writer.Update()
png_writer.Write()
sequence_browser_node.SelectNextItem()
# slicer.app.processEvents()
```
| github_jupyter |
Bot testbed by Arseny
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from operator import itemgetter
from copy import deepcopy
import json
import draftsimtools as ds
# Load M19 drafts
raw_drafts = ds.load_drafts("../../data/m19_2.csv")
# Here other folks load card lists, but I grab them from json instead
# m19_set = ds.create_set("data/m19_rating.tsv", "data/m19_land_rating.tsv")
with open('../../data/Allsets.json', 'r',encoding='utf-8') as json_data:
mtgJSON = json.load(json_data)
jsonSubset = mtgJSON['M19']['cards']
thisSet = {card['name'] : card for card in jsonSubset}
# Another (fancier) way to create a list of names + lots of other useful stuff
nameList = pd.DataFrame.from_dict(thisSet, orient='index', columns=['colors','rarity','type','convertedManaCost'])
nameList['Name'] = nameList.index # We need names as a column, not an index
nameList['index'] = range(len(nameList))
nameList = nameList.set_index('index') # And we need a normal numerical index
nameList[1:5]
# Process names, then handle weird card names (those with commas)
nameList['Name'] = nameList.Name.str.replace(' ','_')
# This utility method searches for "Name" column in nameList that have commas
nameList, raw_drafts = ds.fix_commas(nameList, raw_drafts) # Returns a tuple, as it updates both
# nameList.Name[nameList.Name.str.find(',')!=-1] # There should be no longer any cards with commas
# Process the drafts, deconstructing packs (hands) at every turn of every draft
drafts = ds.process_drafts(raw_drafts)
```
### Make sure all cards are listed, even weird foils
```
# Make sure all cards are listed in the nameList; update if necessary
for iDraft in range(200): #range(len(subset_drafts)):
if iDraft>0 and iDraft % 100 == 0:
print("Draft #%d" % iDraft)
draft = drafts[iDraft]
for pack in draft:
for cardName in pack:
try:
pos = nameList[nameList.Name==cardName].index[0]
except:
print("---Unrecognized card: ",cardName) # All unrecognized cards here seem to be foil lands
# colors rarity type convertedManaCost Name
nameList = nameList.append({'colors':[],'rarity':'weird','type':'weird',
'convertedManaCost':0,'Name':cardName},ignore_index=True)
```
### Now this part below creates bots and tests them
Random bot, defined just in case we need it for benchmarking
```
class RandomBot(object):
def __init__(self,nameList):
self.nameList = nameList # a list with 'Name' column, containing the names
def rank_pack(self,longVector):
nCards = int(len(longVector)/2)
collection = longVector[0:nCards]
pack = longVector[nCards:]
# botCard = np.random.choice(np.nonzero(pack)[0])
# Create fake pick preferences (fake, because for this bot it's )
preferences = np.random.uniform(size=nCards)*(pack>0) # Mask a random vector with available cards
return preferences
pColl = np.loadtxt('bots_data/bayes_pCoDraft.csv', delimiter=",")
pPack = np.loadtxt('bots_data/bayes_pChoice.csv', delimiter=",")
pFull = np.loadtxt('bots_data/bayes_pFull.csv', delimiter=",")
names = pd.read_csv('bots_data/bayes_names.csv')
```
The Bayesian probability of drafting a card $i$ if we see card $j$ in the collection is: $P(i|j) = P_i P_c(j|i) \big/ P_j$ , where $P_i$ is the probability of eventually drafting card $i$, and $P_c(j|i)$ is the probability of seeing card $j$ in the collection, as the card $i$ is drafted.
$P_c(j|i)$ can be explicitly calculated from the draft statistics: $P_c(j|i) = p_{ij}/P_i$. Here $p_{ij}$ is the probability that in a randomly chosen moment, across all drafts, card $j$ is present in the collection, and card $i$ is being picked. Substituting $P_c(j|i)$ in the formula above, we get $P(i|j) = P_i \cdot p_{ij} \big/ (P_i \cdot P_j) = p_{ij}\big/ P_j$
As our goal is to find the card $i$ with highest probability of being drafted $i = \text{argmax}_i \big( P(i|j) \big)$, we can ignore $P_j$ in the denumenator, as it is independent of $i$. Therefore we have:
$i = \text{argmax}_i P(i|j) = \text{argmax}_i p_{ij} = \text{argmax}_i d_{ij}\big/n_{ij}$ , where $d_{ij}$ is the total number of time card $i$ was drafted when card $j$ was present in the collection, and $n_{ij}$ is the number of times card $i$ was in the pack, and could have been drafted, when $j$ was in the collection.
If we have several cards in the collection, and assuming independence of interactions between cards (Naive Bayes approach), we get:
$i= \text{argmax}_i \prod_j p_{ij}$
> Similar logic can be applied to the process of picking card $i$ when another card $k$ is also available in the pack. In this case, we can count all cases when card $i$ was preferred over card $k$ ($c_{ik}$), as well as the total number of cases when cards $i$ and $k$ were in the pack together, and so $i$ could have been picked over $k$ ($m_{ik}$), and include the related probability $c_{ik}\big/ m_{ij}$ in the formula.
In practice, after calculating $p_{ij}$, we move to log-likelihoods, to replace products by sums:
$i = \text{argmax}_i \sum_j \log(p_{ij})$, and this operation can be further vectorized as $Q\cdot c$ where $Q_{ij} = \log(p_{ij})$ is a matrix of log-transformed probabilities of co-drafting card $i$ towards card $j$ in the collection, and $c$ is an n-encoded vector of all cards in the collection: $c_j=$ the number of cards of type $j$ currently present in the collection.
```
minPEver = np.min(pColl[pColl>0])/2 # Even smaller than the smallest non-zero P observed
pF = pFull
pC = np.log(np.maximum(pColl,minPEver))-np.log(minPEver) # C from 'Collection'
pP = np.log(np.maximum(pPack,minPEver))-np.log(minPEver) # P from 'Pack'
```
Note that this formula for pC above is based on an approximation for the number of times i and j were seen together. For large numbers it could work, but it would be better to just keep track of it.
```
rating = np.matmul(pP,np.ones(len(names)))
order = np.argsort(rating)
rank = order.argsort()
d = pd.DataFrame({'rating': rating, 'rank': rank, 'name': nameList.Name, 'curm': nameList.rarity})
d = d.sort_values(by='rank',ascending=False)
# print(d.to_string()) # To print it in full
```
Here comes the main testing loop:
```
#bot = ds.RandomBot() # Doesn't work as Henry's bots take lists as arguments, not vectors
# bot = RaredraftBot(nameList)
# bot = RandomBot(nameList)
nCardsInSet = len(names)
pickCount = 0
accuracySimple = 0
accuracyRank = 0 # A difference in ranks
stats = [] # To track hits a bit better
for iDraft in range(500): #range(len(subset_drafts)): # <----- Uncomment this to ran it on a full dataset
if iDraft>0 and iDraft % 100 == 0:
print("Draft #%d" % iDraft)
draft = drafts[iDraft+2]
collection = np.zeros(nCardsInSet)
iPack = 0
for pack in draft:
packVector = np.zeros(nCardsInSet)
for cardName in pack:
#pos = nameList[nameList.Name==cardName].index[0]
pos = names[names.Name==cardName].index[0]
packVector[pos] += 1 # Mark the card
if cardName==pack[0]: # If the first card in the pack, note it, as the Human picked it
humanCard = pos
# ----- that's where the bot should be called, with current collection and pack
#ratings = bot.rank_pack(np.concatenate([collection, packVector]))
#ratings = pF + np.matmul(pC,collection) + np.matmul(pP,packVector)
if iPack==0: # First card
ratings = np.matmul(pP,packVector) # Only ratings
else:
ratings = np.matmul(pC,collection) # Only synergies
ratings = ratings*(packVector>0)
#ratings2 = np.matmul(pC,collection) # Alternative approach - synergy only
#ratings2 = ratings2*(packVector>0)
# print(pd.DataFrame({'rating': ratings, 'name': nameList.Name}).loc[ratings<0].to_string())
botCard = np.argmax(ratings)
#botCard2 = np.argmax(ratings2)
# --- Update accuracy measurements and the collection:
pickCount += 1
if(humanCard==botCard):
accuracySimple += 1 # Simple count of correctly picked cards
stats.append([1,iPack])
#if(humanCard==botCard2):
# stats.append([2,iPack])
# This debugging output uses nameList rather than names, works only if the sequence is the same
if 0:
print("%4d %2d %2d %20s %10s \t %20s %10s" % (iDraft,iPack,len(pack),
nameList.iloc[humanCard]['Name'],nameList.iloc[humanCard]['colors'],
nameList.iloc[botCard]['Name'], nameList.iloc[botCard]['colors']),end='')
if(humanCard==botCard):
print('\tyep',end='')
print()
ranks = len(ratings)-ratings.argsort().argsort()-1
accuracyRank += ranks[humanCard]
# print("%3d %3d %3d %3d" % (iDraft,iCard,sum(packVector),ranks[humanCard]))
collection[humanCard] += 1 # Update collection
iPack += 1
print("Simple accuracy: %4.2f" % (accuracySimple/pickCount))
print("Av. rank error: %4.2f" % (accuracyRank/pickCount))
data = pd.DataFrame(stats,columns=('type','y'))
data.to_csv('output_files/Arsenys_bots_summary.csv',index=False)
(y,x) = np.histogram(data[data.type==1].y,np.array(range(46))-0.5)
y = y/max(y)
plt.plot(range(0,15),y[0:15],'b.-',label="Full model");
plt.plot(range(15,30),y[15:30],'b.-');
plt.plot(range(30,45),y[30:45],'b.-');
(y,x) = np.histogram(data[data.type==2].y,np.array(range(46))-0.5)
y = y/max(y)
plt.plot(range(0,15),y[0:15],'r.-',label="Synergy only");
plt.plot(range(15,30),y[15:30],'r.-');
plt.plot(range(30,45),y[30:45],'r.-');
plt.legend(loc=4)
plt.show()
# Playground for ranking:
a = np.array([1,5,2,9,3])
b = len(b)-a.argsort().argsort()-1
b
# Playground for iterating through a list of lists
colorStats = {'W':0, 'U':0, 'B':0, 'R':0, 'G':0}
colorData = [['W'] , ['W','B'], ['U']]
for subList in colorData:
for item in subList:
colorStats[item] += 1
colorStats
# Playground for subsetting and quering
possible = np.array([3,5,15,28])
# nameList.iloc[possible].rarity.str[0]=='u' # A good stub to work with rarity
print(nameList.iloc[possible].colors)
condition = list('W' in color for color in nameList.iloc[possible].colors)
print(condition)
possible[condition]
# --- Naive bot (drafts like a 5-years-old)
class RaredraftBot(object):
def __init__(self,nameList):
self.nameList = nameList # a list with 'Name' column, containing the names
def rank_pack(self,longVector):
nCards = int(len(longVector)/2)
collection = longVector[0:nCards]
pack = longVector[nCards:]
# - Analyze collection:
colorStats = {'W':0, 'U':0, 'B':0, 'R':0, 'G':0}
for subList in list(color for color in self.nameList.iloc[np.nonzero(collection)[0]].colors):
for item in subList:
colorStats[item] += 1
currentColor = max(colorStats, key=colorStats.get)
# Comment for this notation of max: Passes each iterable to the key function,
# Returns largest iterable based on the return value of the key function
# print(colorStats, currentColor)
# - Make the pick:
possibleCards = np.nonzero(packVector)[0]
rating = np.ones(len(possibleCards)) # ones because existing cards still better than nonexisting
condition = self.nameList.iloc[possibleCards].rarity.str[0]=='m' # mythics
rating[condition] += 10 # Cecause rare+on color = 9, and 10 is greater
condition = self.nameList.iloc[possibleCards].rarity.str[0]=='r' # rares
rating[condition] += 6 # Because uncommon+oncolor = 5, and 6 is greater
condition = self.nameList.iloc[possibleCards].rarity.str[0]=='u' # uncommons
rating[condition] += 3 # because common+uncolor = 2, and 3 is greater
condition = list(currentColor in color
for color in self.nameList.iloc[possibleCards].colors) # Follow the color
rating[condition] += 2
#print(rating)
# botCard = possibleCards[np.argmax(rating)]
preferences = np.zeros(nCards)
preferences[possibleCards] = rating
return preferences
```
| github_jupyter |
```
import numpy as np
import random
from math import *
import time
import copy
import torch
from torch.autograd import Variable
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torch.optim.lr_scheduler import StepLR, MultiStepLR
torch.set_default_tensor_type('torch.DoubleTensor')
# defination of activation function
def activation(x):
return x * torch.sigmoid(x)
# build ResNet with one blocks
class Net(nn.Module):
def __init__(self,input_size,width):
super(Net,self).__init__()
self.layer_in = nn.Linear(input_size,width)
self.layer_1 = nn.Linear(width,width)
self.layer_2 = nn.Linear(width,width)
self.layer_out = nn.Linear(width,1)
def forward(self,x):
output = self.layer_in(x)
output = output + activation(self.layer_2(activation(self.layer_1(output)))) # residual block 1
output = self.layer_out(output)
return output
input_size = 1
width = 4
# exact solution
def u_ex(x):
return torch.sin(pi*x)
# f(x)
def f(x):
return pi**2 * torch.sin(pi*x)
grid_num = 200
x = torch.zeros(grid_num + 1, input_size)
for index in range(grid_num + 1):
x[index] = index * 1 / grid_num
net = Net(input_size,width)
def model(x):
return x * (x - 1.0) * net(x)
# loss function to DRM by auto differential
def loss_function(x):
h = 1 / grid_num
sum_0 = 0.0
sum_1 = 0.0
sum_2 = 0.0
sum_a = 0.0
sum_b = 0.0
for index in range(grid_num):
x_temp = x[index] + h / 2
x_temp.requires_grad = True
grad_x_temp = torch.autograd.grad(outputs = model(x_temp), inputs = x_temp, grad_outputs = torch.ones(model(x_temp).shape), create_graph = True)
sum_1 += (0.5*grad_x_temp[0]**2 - f(x_temp)[0]*model(x_temp)[0])
for index in range(1, grid_num):
x_temp = x[index]
x_temp.requires_grad = True
grad_x_temp = torch.autograd.grad(outputs = model(x_temp), inputs = x_temp, grad_outputs = torch.ones(model(x_temp).shape), create_graph = True)
sum_2 += (0.5*grad_x_temp[0]**2 - f(x_temp)[0]*model(x_temp)[0])
x_temp = x[0]
x_temp.requires_grad = True
grad_x_temp = torch.autograd.grad(outputs = model(x_temp), inputs = x_temp, grad_outputs = torch.ones(model(x_temp).shape), create_graph = True)
sum_a = 0.5*grad_x_temp[0]**2 - f(x_temp)[0]*model(x_temp)[0]
x_temp = x[grid_num]
x_temp.requires_grad = True
grad_x_temp = torch.autograd.grad(outputs = model(x_temp), inputs = x_temp, grad_outputs = torch.ones(model(x_temp).shape), create_graph = True)
sum_a = 0.5*grad_x_temp[0]**2 - f(x_temp)[0]*model(x_temp)[0]
sum_0 = h / 6 * (sum_a + 4 * sum_1 + 2 * sum_2 + sum_b)
return sum_0
def get_weights(net):
""" Extract parameters from net, and return a list of tensors"""
return [p.data for p in net.parameters()]
def set_weights(net, weights, directions=None, step=None):
"""
Overwrite the network's weights with a specified list of tensors
or change weights along directions with a step size.
"""
if directions is None:
# You cannot specify a step length without a direction.
for (p, w) in zip(net.parameters(), weights):
p.data.copy_(w.type(type(p.data)))
else:
assert step is not None, 'If a direction is specified then step must be specified as well'
if len(directions) == 2:
dx = directions[0]
dy = directions[1]
changes = [d0*step[0] + d1*step[1] for (d0, d1) in zip(dx, dy)]
else:
changes = [d*step for d in directions[0]]
for (p, w, d) in zip(net.parameters(), weights, changes):
p.data = w + torch.Tensor(d).type(type(w))
def set_states(net, states, directions=None, step=None):
"""
Overwrite the network's state_dict or change it along directions with a step size.
"""
if directions is None:
net.load_state_dict(states)
else:
assert step is not None, 'If direction is provided then the step must be specified as well'
if len(directions) == 2:
dx = directions[0]
dy = directions[1]
changes = [d0*step[0] + d1*step[1] for (d0, d1) in zip(dx, dy)]
else:
changes = [d*step for d in directions[0]]
new_states = copy.deepcopy(states)
assert (len(new_states) == len(changes))
for (k, v), d in zip(new_states.items(), changes):
d = torch.tensor(d)
v.add_(d.type(v.type()))
net.load_state_dict(new_states)
def get_random_weights(weights):
"""
Produce a random direction that is a list of random Gaussian tensors
with the same shape as the network's weights, so one direction entry per weight.
"""
return [torch.randn(w.size()) for w in weights]
def get_random_states(states):
"""
Produce a random direction that is a list of random Gaussian tensors
with the same shape as the network's state_dict(), so one direction entry
per weight, including BN's running_mean/var.
"""
return [torch.randn(w.size()) for k, w in states.items()]
def get_diff_weights(weights, weights2):
""" Produce a direction from 'weights' to 'weights2'."""
return [w2 - w for (w, w2) in zip(weights, weights2)]
def get_diff_states(states, states2):
""" Produce a direction from 'states' to 'states2'."""
return [v2 - v for (k, v), (k2, v2) in zip(states.items(), states2.items())]
def normalize_direction(direction, weights, norm='filter'):
"""
Rescale the direction so that it has similar norm as their corresponding
model in different levels.
Args:
direction: a variables of the random direction for one layer
weights: a variable of the original model for one layer
norm: normalization method, 'filter' | 'layer' | 'weight'
"""
if norm == 'filter':
# Rescale the filters (weights in group) in 'direction' so that each
# filter has the same norm as its corresponding filter in 'weights'.
for d, w in zip(direction, weights):
d.mul_(w.norm()/(d.norm() + 1e-10))
elif norm == 'layer':
# Rescale the layer variables in the direction so that each layer has
# the same norm as the layer variables in weights.
direction.mul_(weights.norm()/direction.norm())
elif norm == 'weight':
# Rescale the entries in the direction so that each entry has the same
# scale as the corresponding weight.
direction.mul_(weights)
elif norm == 'dfilter':
# Rescale the entries in the direction so that each filter direction
# has the unit norm.
for d in direction:
d.div_(d.norm() + 1e-10)
elif norm == 'dlayer':
# Rescale the entries in the direction so that each layer direction has
# the unit norm.
direction.div_(direction.norm())
def normalize_directions_for_weights(direction, weights, norm='filter', ignore='biasbn'):
"""
The normalization scales the direction entries according to the entries of weights.
"""
assert(len(direction) == len(weights))
for d, w in zip(direction, weights):
if d.dim() <= 1:
if ignore == 'biasbn':
d.fill_(0) # ignore directions for weights with 1 dimension
else:
d.copy_(w) # keep directions for weights/bias that are only 1 per node
else:
normalize_direction(d, w, norm)
def normalize_directions_for_states(direction, states, norm='filter', ignore='ignore'):
assert(len(direction) == len(states))
for d, (k, w) in zip(direction, states.items()):
if d.dim() <= 1:
if ignore == 'biasbn':
d.fill_(0) # ignore directions for weights with 1 dimension
else:
d.copy_(w) # keep directions for weights/bias that are only 1 per node
else:
normalize_direction(d, w, norm)
def ignore_biasbn(directions):
""" Set bias and bn parameters in directions to zero """
for d in directions:
if d.dim() <= 1:
d.fill_(0)
def create_random_direction(net, dir_type='weights', ignore='biasbn', norm='filter'):
"""
Setup a random (normalized) direction with the same dimension as
the weights or states.
Args:
net: the given trained model
dir_type: 'weights' or 'states', type of directions.
ignore: 'biasbn', ignore biases and BN parameters.
norm: direction normalization method, including
'filter" | 'layer' | 'weight' | 'dlayer' | 'dfilter'
Returns:
direction: a random direction with the same dimension as weights or states.
"""
# random direction
if dir_type == 'weights':
weights = get_weights(net) # a list of parameters.
direction = get_random_weights(weights)
normalize_directions_for_weights(direction, weights, norm, ignore)
elif dir_type == 'states':
states = net.state_dict() # a dict of parameters, including BN's running mean/var.
direction = get_random_states(states)
normalize_directions_for_states(direction, states, norm, ignore)
return direction
def tvd(m, l_i):
# load model parameters
pretrained_dict = torch.load('net_params_DRM_to_DGM.pkl')
# get state_dict
net_state_dict = net.state_dict()
# remove keys that does not belong to net_state_dict
pretrained_dict_1 = {k: v for k, v in pretrained_dict.items() if k in net_state_dict}
# update dict
net_state_dict.update(pretrained_dict_1)
# set new dict back to net
net.load_state_dict(net_state_dict)
weights_temp = get_weights(net)
states_temp = net.state_dict()
step_size = 2 * l_i / m
grid = np.arange(-l_i, l_i + step_size, step_size)
num_direction = 1
loss_matrix = torch.zeros((num_direction, len(grid)))
for temp in range(num_direction):
weights = weights_temp
states = states_temp
direction_temp = create_random_direction(net, dir_type='weights', ignore='biasbn', norm='filter')
normalize_directions_for_states(direction_temp, states, norm='filter', ignore='ignore')
directions = [direction_temp]
for dx in grid:
itemindex_1 = np.argwhere(grid == dx)
step = dx
set_states(net, states, directions, step)
loss_temp = loss_function(x)
loss_matrix[temp, itemindex_1[0]] = loss_temp
# clear memory
torch.cuda.empty_cache()
# get state_dict
net_state_dict = net.state_dict()
# remove keys that does not belong to net_state_dict
pretrained_dict_1 = {k: v for k, v in pretrained_dict.items() if k in net_state_dict}
# update dict
net_state_dict.update(pretrained_dict_1)
# set new dict back to net
net.load_state_dict(net_state_dict)
weights_temp = get_weights(net)
states_temp = net.state_dict()
interval_length = grid[-1] - grid[0]
TVD = 0.0
for temp in range(num_direction):
for index in range(loss_matrix.size()[1] - 1):
TVD = TVD + np.abs(float(loss_matrix[temp, index] - loss_matrix[temp, index + 1]))
Max = np.max(loss_matrix.detach().numpy())
Min = np.min(loss_matrix.detach().numpy())
TVD = TVD / interval_length / num_direction
TVD = TVD / interval_length / num_direction / (Max - Min)
return TVD, Max, Min
M = 100
m = 100
l_i = 0.01
TVD_DRM = 0.0
time_start = time.time()
Max = []
Min = []
Result = []
print('====================')
print('Result for l = 0.01.')
print('====================')
for count in range(M):
TVD_temp, Max_temp, Min_temp = tvd(m, l_i)
Max.append(Max_temp)
Min.append(Min_temp)
Result.append(TVD_temp)
print('Current direction TVD of DRM is: ', TVD_temp)
TVD_DRM = TVD_DRM + TVD_temp
print((count + 1) / M * 100, '% finished.')
TVD_DRM = TVD_DRM / M
print('All directions average TVD of DRM is: ', TVD_DRM)
# Result = Result / (max(Max) - min(Min))
print('Variance TVD of DRM is: ', np.sqrt(np.var(Result, ddof = 1)))
print('Roughness Index of DRM is: ', np.sqrt(np.var(Result, ddof = 1)) / TVD_DRM)
time_end = time.time()
print('Total time costs: ', time_end - time_start, 'seconds')
```
| github_jupyter |
In this notebook you can define your own configuration and run the model based on your custom configuration.
## Dataset
`dataset_path` shows the path to `data_paths` directory that contains every image and its pair path. The `resize` value selects the width, and the height dimensions that each image will be resized to.
```
dataset_path = '.'
resize = [128, 256]
```
## Model
`baseline_model` selects the compression model. The accepted models for this parameter are bmshj18 for [Variational image compression with a scale hyperprior](https://arxiv.org/abs/1802.01436) and bls17 for [End-to-end Optimized Image Compression](https://arxiv.org/abs/1611.01704). If `use_side_info` is set as `True`, then the baseline model is modified using our proposed method for using side information for compressing.
If `load_weight` is `True`, then in model initialization, the weight saved in `weight_path` is loaded to the model. You can also specify the experiment name in `experiment_name`.
```
baseline_model = 'bls17' # can be bmshj18 for Variational image compression with a scale hyperprior by Ballé, et al.
# or bls17 for End-to-end Optimized Image Compression by Ballé, et al.
use_side_info = True # if True then the modified version of baseline model for distributed compression is used.
num_filters = 192 # number of filters used in the baseline model network
cuda = True
load_weight = False
weight_path = './pretrained_weights/balle17+ours_MS-SSIM_lambda3e-05.pt' # weight path for loading the weight
# note that we provide some pretrained weights, accessible from the anonymous link provided in README.md
```
## Training
For training set `train` to be `True`. `lambda` shows the lambda value in the rate-distortion equation and `alpha` and `beta` correspond to the handles on the reconstruction of the correlated image and amount of common information extracted from the decoder-only side information, respectively. `distortion_loss` selects the distortion evaluating method. Its accepted values are MS-SSIM for the ms-ssim method or MSE for mean squared error.
`verbose_period: 50` indicates that every 50 epochs print the results of the validation dataset.
```
train = True
epochs = 50000
train_batch_size = 1
lr = 0.0001
lmbda = 0.00003 # the lambda value in rate-distortion equation
alpha = 1
beta = 1
distortion_loss = 'MS-SSIM' # can be MS-SSIM or MSE. selects the method by which the distortion is calculated during training
verbose_period = 50 # non-positive value indicates no verbose
```
## Weights and Results parameters
If you wish to save the model weights after training set `save_weights` `True`. `save_output_path` shows the directory path where the model weights are saved.
For the weights, in `save_output_path` a `weight` folder will be created, and the weights will be saved there with the name according to `experiment_name`.
```
save_weights = True
save_output_path = './outputs' # path where results and weights will be saved
experiment_name = 'bls17_with_side_info_MS-SSIM_lambda:3e-05'
```
## Test
If you wish to test the model and save the results set `test` to `True`. If `save_image` is set to `True` then a `results` folder will be created, and the reconstructed images will be saved in `save_output_path/results` during testing, with the results named according to `experiment_name`.
```
test = True
save_image = True
```
## Inference
In order to (only) carry out inference, please open `configs/config.yaml` and change the relevant lines as follows:
```
resize = [128, 256] # we used this crop size for our inference
dataset_path = '.'
train = False
load_weight = True
test = True
save_output_path = './inference'
save_image = True
```
Download the desired weights and put them in `pretrained_weights` folder and put the dataset folder in the root .
Based on the weight you chose, specify the weight name, and the experiment name in `configs/config.yaml`:
```
weight_path: './pretrained_weights/...' # load a specified pre-trained weight
experiment_name: '...' # a handle for the saved results of the inference
```
Also, change `baseline_model` and `use_side_info` parameters in `configs/config.yaml` accordingly.
For example, for the `balle2017+ours` weights, these parameters should be:
```
baseline_model: 'bls17'
use_side_info: True
```
After running the code using the commands in below section, the results will be saved in `inference` folder.
## Saving Custom Configuration
By running this piece of code you can save your configuration as a yaml file file in the configs folder. You can set your configuration file name by changing `config_name` variable.
```
import yaml
config = {
"dataset_path": dataset_path,
"resize": resize,
"baseline_model": baseline_model,
"use_side_info": use_side_info,
"num_filters": num_filters,
"cuda": cuda,
"load_weight": load_weight,
"weight_path": weight_path,
"experiment_name": experiment_name,
"train": train,
"epochs": epochs,
"train_batch_size": train_batch_size,
"lr": lr,
"lambda": lmbda,
"distortion_loss": distortion_loss,
"verbose_period": verbose_period,
"save_weights": save_weights,
"save_output_path": save_output_path,
"test": test,
"save_image": save_image
}
config_name = "CUSTOM_CONFIG_FILE_NAME.yaml"
with open('configs/' + config_name) + config_name, 'w') as outfile:
yaml.dump(config, outfile, default_flow_style=None, sort_keys=False)
```
## Running the Model
```
!python main.py --config=configs/$config_name
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.